text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Query Classifier Tutorial
[](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial14_Query_Classifier.ipynb)
In this tutorial we introduce the query classifier the goal of introducing this feature was to optimize the overall flow of Haystack pipeline by detecting the nature of user queries. Now, the Haystack can detect primarily three types of queries using both light-weight SKLearn Gradient Boosted classifier or Transformer based more robust classifier. The three categories of queries are as follows:
### 1. Keyword Queries:
Such queries don't have semantic meaning and merely consist of keywords. For instance these three are the examples of keyword queries.
* arya stark father
* jon snow country
* arya stark younger brothers
### 2. Interrogative Queries:
In such queries users usually ask a question, regardless of presence of "?" in the query the goal here is to detect the intent of the user whether any question is asked or not in the query. For example:
* who is the father of arya stark ?
* which country was jon snow filmed ?
* who are the younger brothers of arya stark ?
### 3. Declarative Queries:
Such queries are variation of keyword queries, however, there is semantic relationship between words. Fo example:
* Arya stark was a daughter of a lord.
* Jon snow was filmed in a country in UK.
* Bran was brother of a princess.
In this tutorial, you will learn how the `TransformersQueryClassifier` and `SklearnQueryClassifier` classes can be used to intelligently route your queries, based on the nature of the user query. Also, you can choose between a lightweight Gradients boosted classifier or a transformer based classifier.
Furthermore, there are two types of classifiers you can use out of the box from Haystack.
1. Keyword vs Statement/Question Query Classifier
2. Statement vs Question Query Classifier
As evident from the name the first classifier detects the keywords search queries and semantic statements like sentences/questions. The second classifier differentiates between question based queries and declarative sentences.
### Prepare environment
#### Colab: Enable the GPU runtime
Make sure you enable the GPU runtime to experience decent speed in this tutorial.
**Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
<img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/_src/img/colab_gpu_runtime.jpg">
These lines are to install Haystack through pip
```
# Install the latest release of Haystack in your own environment
#! pip install farm-haystack
# Install the latest master of Haystack
!pip install grpcio-tools==1.34.1
!pip install --upgrade git+https://github.com/deepset-ai/haystack.git
# Install pygraphviz
!apt install libgraphviz-dev
!pip install pygraphviz
# If you run this notebook on Google Colab, you might need to
# restart the runtime after installing haystack.
# In Colab / No Docker environments: Start Elasticsearch from source
! wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.9.2-linux-x86_64.tar.gz -q
! tar -xzf elasticsearch-7.9.2-linux-x86_64.tar.gz
! chown -R daemon:daemon elasticsearch-7.9.2
import os
from subprocess import Popen, PIPE, STDOUT
es_server = Popen(['elasticsearch-7.9.2/bin/elasticsearch'],
stdout=PIPE, stderr=STDOUT,
preexec_fn=lambda: os.setuid(1) # as daemon
)
# wait until ES has started
! sleep 30
```
If running from Colab or a no Docker environment, you will want to start Elasticsearch from source
## Initialization
Here are some core imports
Then let's fetch some data (in this case, pages from the Game of Thrones wiki) and prepare it so that it can
be used indexed into our `DocumentStore`
```
from haystack.utils import print_answers, fetch_archive_from_http, convert_files_to_dicts, clean_wiki_text, launch_es
from haystack.pipelines import Pipeline, RootNode
from haystack.document_stores import ElasticsearchDocumentStore
from haystack.nodes import ElasticsearchRetriever, DensePassageRetriever, FARMReader, TransformersQueryClassifier, SklearnQueryClassifier
#Download and prepare data - 517 Wikipedia articles for Game of Thrones
doc_dir = "data/article_txt_got"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/documents/wiki_gameofthrones_txt.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# convert files to dicts containing documents that can be indexed to our datastore
got_dicts = convert_files_to_dicts(
dir_path=doc_dir,
clean_func=clean_wiki_text,
split_paragraphs=True
)
# Initialize DocumentStore and index documents
launch_es()
document_store = ElasticsearchDocumentStore()
document_store.delete_documents()
document_store.write_documents(got_dicts)
# Initialize Sparse retriever
es_retriever = ElasticsearchRetriever(document_store=document_store)
# Initialize dense retriever
dpr_retriever = DensePassageRetriever(document_store)
document_store.update_embeddings(dpr_retriever, update_existing_embeddings=False)
reader = FARMReader(model_name_or_path="deepset/roberta-base-squad2")
```
## Keyword vs Question/Statement Classifier
The keyword vs question/statement query classifier essentially distinguishes between the keyword queries and statements/questions. So you can intelligently route to different retrieval nodes based on the nature of the query. Using this classifier can potentially yield the following benefits:
* Getting better search results (e.g. by routing only proper questions to DPR / QA branches and not keyword queries)
* Less GPU costs (e.g. if 50% of your traffic is only keyword queries you could just use elastic here and save the GPU resources for the other 50% of traffic with semantic queries)

Below, we define a `SklQueryClassifier` and show how to use it:
Read more about the trained model and dataset used [here](https://ext-models-haystack.s3.eu-central-1.amazonaws.com/gradboost_query_classifier/readme.txt)
```
# Here we build the pipeline
sklearn_keyword_classifier = Pipeline()
sklearn_keyword_classifier.add_node(component=SklearnQueryClassifier(), name="QueryClassifier", inputs=["Query"])
sklearn_keyword_classifier.add_node(component=dpr_retriever, name="DPRRetriever", inputs=["QueryClassifier.output_1"])
sklearn_keyword_classifier.add_node(component=es_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"])
sklearn_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "DPRRetriever"])
sklearn_keyword_classifier.draw("pipeline_classifier.png")
# Run only the dense retriever on the full sentence query
res_1 = sklearn_keyword_classifier.run(
query="Who is the father of Arya Stark?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_1, details="minimum")
# Run only the sparse retriever on a keyword based query
res_2 = sklearn_keyword_classifier.run(
query="arya stark father"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_2, details="minimum")
# Run only the dense retriever on the full sentence query
res_3 = sklearn_keyword_classifier.run(
query="which country was jon snow filmed ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_3, details="minimum")
# Run only the sparse retriever on a keyword based query
res_4 = sklearn_keyword_classifier.run(
query="jon snow country"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_4, details="minimum")
# Run only the dense retriever on the full sentence query
res_5 = sklearn_keyword_classifier.run(
query="who are the younger brothers of arya stark ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_5, details="minimum")
# Run only the sparse retriever on a keyword based query
res_6 = sklearn_keyword_classifier.run(
query="arya stark younger brothers"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_6, details="minimum")
```
## Transformer Keyword vs Question/Statement Classifier
Firstly, it's essential to understand the trade-offs between SkLearn and Transformer query classifiers. The transformer classifier is more accurate than SkLearn classifier however, it requires more memory and most probably GPU for faster inference however the transformer size is roughly `50 MBs`. Whereas, SkLearn is less accurate however is much more faster and doesn't require GPU for inference.
Below, we define a `TransformersQueryClassifier` and show how to use it:
Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/bert-mini-finetune-question-detection)
```
# Here we build the pipeline
transformer_keyword_classifier = Pipeline()
transformer_keyword_classifier.add_node(component=TransformersQueryClassifier(), name="QueryClassifier", inputs=["Query"])
transformer_keyword_classifier.add_node(component=dpr_retriever, name="DPRRetriever", inputs=["QueryClassifier.output_1"])
transformer_keyword_classifier.add_node(component=es_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"])
transformer_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "DPRRetriever"])
transformer_keyword_classifier.draw("pipeline_classifier.png")
# Run only the dense retriever on the full sentence query
res_1 = transformer_keyword_classifier.run(
query="Who is the father of Arya Stark?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_1, details="minimum")
# Run only the sparse retriever on a keyword based query
res_2 = transformer_keyword_classifier.run(
query="arya stark father"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_2, details="minimum")
# Run only the dense retriever on the full sentence query
res_3 = transformer_keyword_classifier.run(
query="which country was jon snow filmed ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_3, details="minimum")
# Run only the sparse retriever on a keyword based query
res_4 = transformer_keyword_classifier.run(
query="jon snow country"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_4, details="minimum")
# Run only the dense retriever on the full sentence query
res_5 = transformer_keyword_classifier.run(
query="who are the younger brothers of arya stark ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_5, details="minimum")
# Run only the sparse retriever on a keyword based query
res_6 = transformer_keyword_classifier.run(
query="arya stark younger brothers"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_6, details="minimum")
```
## Question vs Statement Classifier
One possible use case of this classifier could be to route queries after the document retrieval to only send questions to QA reader and in case of declarative sentence, just return the DPR/ES results back to user to enhance user experience and only show answers when user explicitly asks it.

Below, we define a `TransformersQueryClassifier` and show how to use it:
Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/question-vs-statement-classifier)
```
# Here we build the pipeline
transformer_question_classifier = Pipeline()
transformer_question_classifier.add_node(component=dpr_retriever, name="DPRRetriever", inputs=["Query"])
transformer_question_classifier.add_node(component=TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier"), name="QueryClassifier", inputs=["DPRRetriever"])
transformer_question_classifier.add_node(component=reader, name="QAReader", inputs=["QueryClassifier.output_1"])
transformer_question_classifier.draw("question_classifier.png")
# Run only the QA reader on the question query
res_1 = transformer_question_classifier.run(
query="Who is the father of Arya Stark?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_1, details="minimum")
# Show only DPR results
res_2 = transformer_question_classifier.run(
query="Arya Stark was the daughter of a Lord."
)
print("ES Results" + "\n" + "="*15)
print_answers(res_2, details="minimum")
```
## Standalone Query Classifier
Below we run queries classifiers standalone to better understand their outputs on each of the three types of queries
```
# Here we create the keyword vs question/statement query classifier
from haystack.pipelines import TransformersQueryClassifier
queries = ["arya stark father","jon snow country",
"who is the father of arya stark","which country was jon snow filmed?"]
keyword_classifier = TransformersQueryClassifier()
for query in queries:
result = keyword_classifier.run(query=query)
if result[1] == "output_1":
category = "question/statement"
else:
category = "keyword"
print(f"Query: {query}, raw_output: {result}, class: {category}")
# Here we create the question vs statement query classifier
from haystack.pipelines import TransformersQueryClassifier
queries = ["Lord Eddard was the father of Arya Stark.","Jon Snow was filmed in United Kingdom.",
"who is the father of arya stark?","Which country was jon snow filmed in?"]
question_classifier = TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier")
for query in queries:
result = question_classifier.run(query=query)
if result[1] == "output_1":
category = "question"
else:
category = "statement"
print(f"Query: {query}, raw_output: {result}, class: {category}")
```
## Conclusion
The query classifier gives you more possibility to be more creative with the pipelines and use different retrieval nodes in a flexible fashion. Moreover, as in the case of Question vs Statement classifier you can also choose the queries which you want to send to the reader.
Finally, you also have the possible of bringing your own classifier and plugging it into either `TransformersQueryClassifier(model_name_or_path="<huggingface_model_name_or_file_path>")` or using the `SklearnQueryClassifier(model_name_or_path="url_to_classifier_or_file_path_as_pickle", vectorizer_name_or_path="url_to_vectorizer_or_file_path_as_pickle")`
## About us
This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
We bring NLP to the industry via open source!
Our focus: Industry specific language models & large scale QA systems.
Some of our other work:
- [German BERT](https://deepset.ai/german-bert)
- [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
- [FARM](https://github.com/deepset-ai/FARM)
Get in touch:
[Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
By the way: [we're hiring!](https://www.deepset.ai/jobs)
| github_jupyter |
# Data description & Problem statement:
I will use the Yelp Review Data Set from Kaggle. Each observation in this dataset is a review of a particular business by a particular user. The "stars" column is the number of stars (1 through 5) assigned by the reviewer to the business. (Higher stars is better.) In other words, it is the rating of the business by the person who wrote the review. The "cool" column is the number of "cool" votes this review received from other Yelp users. The "useful" and "funny" columns are similar to the "cool" column. Here, the goal is to model/clusterize the topics of Yelp reviews.
# Workflow:
- Load the dataset
- Data cleaning (e.g. remove formats and punctuations)
- Basic data exploration
- Text vectorization, using "Bag of Words" technique
- Use "Latent Dirichlet Allocation" for document clustering (i.e. topic modeling)
- Determine, sort and print most important words/features for each topic
```
import sklearn
import numpy as np
import scipy as sc
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import preprocessing
%matplotlib inline
from scipy import stats
import warnings
warnings.filterwarnings("ignore")
# we insatll and import spacy package for some advanced tokenizaion techniques:
import spacy
# we also install and import mglearn package (using !pip install mglearn) for some interesting visualization of results:
import mglearn
ls
```
# load and prepare the text data:
```
reviews = pd.read_csv('yelp_review.csv')
from sklearn.feature_extraction.text import CountVectorizer
vect = CountVectorizer(max_features=500,
stop_words="english",
ngram_range=(1, 1),
max_df=0.3)
X = vect.fit_transform(reviews['text'][0:100000])
```
# document clustering with Latent Dirichlet Allocation: LDA
```
from sklearn.decomposition import LatentDirichletAllocation
lda = LatentDirichletAllocation(n_topics=5,
learning_method="batch",
max_iter=24,
random_state=42)
# We build the model and transform the data in one step
document_topics = lda.fit_transform(X)
# For each topic (a row in the components_), sort the features (ascending)
sorting = np.argsort(lda.components_, axis=1)[:, ::-1]
# Get the feature names from the vectorizer
feature_names = np.array(vect.get_feature_names())
# Print out the 5 topics:
mglearn.tools.print_topics(topics=range(5), feature_names=feature_names,
sorting=sorting, topics_per_chunk=5, n_words=10)
```
| github_jupyter |
# Create journals
We need some more ways to create the journals for the batch runs. Currently, these methods are supported:
- b.from_db()
- b.from_file(filename)
## Within the Journal class
```python
def from_file(self, file_name=None):
"""Loads a DataFrame with all the needed info about the experiment"""
file_name = self._check_file_name(file_name)
with open(file_name, "r") as infile:
top_level_dict = json.load(infile)
pages_dict = top_level_dict["info_df"]
pages = pd.DataFrame(pages_dict)
pages.cellpy_file_names = pages.cellpy_file_names.apply(self._fix_cellpy_paths)
self.pages = pages
self.file_name = file_name
self._prm_packer(top_level_dict["metadata"])
self.generate_folder_names()
self.paginate()
```
## Within the Batch class
```python
def create_journal(self, description=None, from_db=True):
logging.debug("Creating a journal")
logging.debug(f"description: {description}")
logging.debug(f"from_db: {from_db}")
# rename to: create_journal (combine this with function above)
logging.info(f"name: {self.experiment.journal.name}")
logging.info(f"project: {self.experiment.journal.project}")
if description is not None:
from_db = False
if from_db:
self.experiment.journal.from_db()
self.experiment.journal.to_file()
else:
# TODO: move this into the bacth journal class
if description is not None:
print(f"Creating from {type(description)} is not implemented yet")
logging.info("Creating an empty journal")
logging.info(f"name: {self.experiment.journal.name}")
logging.info(f"project: {self.experiment.journal.project}")
self.experiment.journal.pages = pd.DataFrame(
columns=[
"filenames",
"masses",
"total_masses",
"loadings",
"fixed",
"labels",
"cell_type",
"raw_file_names",
"cellpy_file_names",
"groups",
"sub_groups",
]
)
self.experiment.journal.pages.set_index("filenames", inplace=True)
self.experiment.journal.generate_folder_names()
self.experiment.journal.paginate()
```
```
%load_ext autoreload
%autoreload 2
import os
from pathlib import Path
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import cellpy
from cellpy import prms
from cellpy import prmreader
from cellpy.utils import batch
import holoviews as hv
%matplotlib inline
hv.extension('bokeh')
name = "first"
project = "ocv_tests"
print(" INITIALISATION OF BATCH ".center(80, "="))
b = batch.init(name, project, default_log_level="INFO")
p = b.experiment.journal.create_empty_pages()
filename = "20190204_FC_snx012_01_cc_03"
mass = 0.5
total_mass = 1.0
loading = 0.1
fixed = False
label = "fc_snx012_01"
cell_type = "full_cell"
raw_file_name = [Path(".") / "20190204_FC_snx012_01_cc_01.res"]
cellpy_file_name = Path(".") / "20190204_FC_snx012_01_cc_01.h5"
group = 1
sub_group = 1
p.loc[filename] = [
mass,
total_mass,
loading,
fixed,
label,
cell_type,
raw_file_name,
cellpy_file_name,
group,
sub_group,
]
p
b.pages = p
b.pages
```
### Checking the ``create_journal`` function in ``Batch``
```
b2 = batch.init(default_log_level="DEBUG")
b2.experiment.journal.name = "first"
b2.experiment.journal.project = "ocv_tests"
# see if it finds files (str)
b2.create_journal("creating_journals_by_different_methods.ipynb")
# see if it finds files (pathlib.Path)
b2.create_journal(Path("creating_journals_by_different_methods.ipynb"))
```
### different methods
#### dataframe
```
filename = "20190204_FC_snx012_01_cc_03"
mass = 0.5
total_mass = 1.0
loading = 0.1
fixed = False
label = "fc_snx012_01"
cell_type = "full_cell"
raw_file_name = [Path(".") / "20190204_FC_snx012_01_cc_01.res"]
cellpy_file_name = Path(".") / "20190204_FC_snx012_01_cc_01.h5"
group = 1
sub_group = 1
d = {
"filenames": filename,
"masses": mass,
"total_masses": total_mass,
"loadings": loading,
"fixed": fixed,
"labels": label,
"cell_type": cell_type,
"raw_file_names": raw_file_name,
"cellpy_file_names": cellpy_file_name,
"groups": group,
"sub_groups": sub_group,
}
d2 = {
"filenames": [filename],
"masses": [mass],
"total_masses": [total_mass],
"loadings": [loading],
"fixed": [fixed],
"labels": [label],
"cell_type": [cell_type],
"raw_file_names": [raw_file_name],
"cellpy_file_names": [cellpy_file_name],
"groups": [group],
"sub_groups": [sub_group],
}
d3 = {
"filenames": [filename, filename+"b"],
"masses": [mass, 0.4],
"total_masses": [total_mass, 1.0],
"loadings": [loading, 0.2],
"fixed": [fixed, 1],
"labels": [label, "JPM"],
"cell_type": [cell_type, "anode"],
"raw_file_names": [raw_file_name, raw_file_name],
"cellpy_file_names": [cellpy_file_name, cellpy_file_name],
"groups": [group, 2],
"sub_groups": [sub_group, 1],
}
d4 = {
"filenames": [filename, filename+"b"],
"masses": [mass], # Different length
"total_masses": [total_mass, 1.0],
"loadings": [loading, 0.2],
"fixed": [fixed, 1],
"labels": [label, "JPM"],
"cell_type": [cell_type, "anode"],
"raw_file_names": [raw_file_name, raw_file_name],
"cellpy_file_names": [cellpy_file_name, cellpy_file_name],
"groups": [group, 2],
"sub_groups": [sub_group, 1],
} # this should fail
d5 = {
"filenames": [filename, filename+"b"],
"masses": [mass, 0.2], # Different length
"total_masses": [total_mass, 1.0],
"loadings": [loading, 0.2],
# "fixed": [fixed, 1],
"labels": [label, "JPM"],
"cell_type": [cell_type, "anode"],
"raw_file_names": [raw_file_name, raw_file_name],
"cellpy_file_names": [cellpy_file_name, cellpy_file_name],
"groups": [group, 2],
"sub_groups": [sub_group, 1],
}
p = pd.DataFrame(d)
p2 = pd.DataFrame(d2)
p3 = pd.DataFrame(d3)
p5 = pd.DataFrame(d5)
b3 = batch.init(name, project, default_log_level="DEBUG")
b3.create_journal(d)
b3.pages
b3.create_journal(d2)
b3.pages
b3.create_journal(d3)
b3.pages
b3.create_journal(d5)
b3.pages
b3.create_journal(p)
b3.pages
b3.create_journal(p2)
b3.pages
b3.create_journal(p3)
b3.pages
b3.create_journal(p5)
b3.pages
```
| github_jupyter |
# After creating a script to download the data, and running it, I will look at the data and test some of the functions that I implemented for its analysis (most of them were implemented to solve the Machine Learning for Trading assignments).
```
import os
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import datetime as dt
import scipy.optimize as spo
import sys
%matplotlib inline
%pylab inline
pylab.rcParams['figure.figsize'] = (20.0, 10.0)
%load_ext autoreload
%autoreload 2
sys.path.append('../../')
data_df = pd.read_pickle('../../data/data_df.pkl')
print(data_df.shape)
data_df.head(25)
```
## What if I only want the 'Close' value, maybe in a range, for only some symbols?
```
data_df.xs('Close', level='feature')
data_df.loc[dt.datetime(1993,2,4):dt.datetime(1993,2,7)]
symbols = ['SPY', 'AMD', 'IBM']
data_df.xs('Close', level='feature').loc[dt.datetime(1993,2,4):dt.datetime(1993,2,7),symbols]
```
## Let's test the function to fill the missing data
```
from utils import preprocessing
select = ['SPY', 'GOOG', 'GM']
selected_df = data_df.xs('Close', level='feature').loc[:,select]
selected_df.plot()
selected_df = preprocessing.fill_missing(selected_df)
selected_df.plot()
```
## A useful function to show the evolution of a portfolio value
```
from utils import analysis
analysis.assess_portfolio(start_date = dt.datetime(2006,1,22),
end_date = dt.datetime(2016,12,31),
symbols = ['GOOG','AAPL','AMD','XOM'],
allocations = [0.1,0.2,0.3,0.4],
initial_capital = 1000,
risk_free_rate = 0.0,
sampling_frequency = 252.0,
data = data_df,
gen_plot=True,
verbose=True)
from utils import marketsim
```
### Limit leverage to 2.0
```
value_df, constituents_df = marketsim.simulate_orders('../../data/orders.csv',
data_df,
initial_cap=100000,
leverage_limit=2.0,
from_csv=True)
analysis.value_eval(value_df, verbose=True, graph=True, data_df=data_df)
constituents_df.plot()
```
### No leverage limit
```
value_df, constituents_df = marketsim.simulate_orders('../../data/orders.csv',
data_df,
initial_cap=100000,
leverage_limit=None,
from_csv=True)
analysis.value_eval(value_df, verbose=True, graph=True, data_df=data_df)
constituents_df.plot()
analysis.assess_portfolio(start_date = dt.datetime(1993,1,22),
end_date = dt.datetime(2016,12,31),
symbols = ['SPY'],
allocations = [1.0],
initial_capital = 1000,
risk_free_rate = 0.0,
sampling_frequency = 252.0,
data = data_df,
gen_plot=True,
verbose=True)
```
| github_jupyter |
# Deep Learning with Python
## 6.2 Understanding recurrent neural networks
> 理解循环神经网络
之前我们用的全连接网络和卷积神经网络都有是被叫做 feedforward networks (前馈网络) 的。这种网络是无记忆的,也就是说,它们单独处理每个输入,在输入与输入之间没有保存任何状态。在这种网络中,我们要处理时间/文本等序列,就必须把一个完整的序列处理成一个大张量,整个的传到网络中,让模型一次看完整个序列。
这个显然和我们人类阅读、学习文本等信息的方式有所区别。我们不是一眼看完整本书的,我们要一个词一个词地看,眼睛不停移动获取新的数据的同时,记住之前的内容,将新的、旧的内容联系在一起来理解整句话的意思。说抽象一些,我们会保存一个关于所处理内容的内部模型,这个模型根据过去的信息构建,并随着新信息的进入而不断更新。我们都是以这种渐进的方式处理信息的。
按照这种思想,我们又得到一种新的模型,叫做**循环神经网络**(recurrent neural network, RNN),这网络会遍历处理所有序列元素,并保存一个记录已查看内容相关信息的状态(state)。而在处理下一条序列之时,RNN 状态会被重置。使用 RNN 时,我们仍可以将一个序列整个的输出网络,不过在网络内部,数据不再是直接被整个处理,而是自动对序列元素进行遍历。

为了理解循环神经网络,我们用 Numpy 手写一个玩具版的 RNN 前向传递。考虑处理形状为 `(timesteps, input_features)` 的一条序列,RNN 在 timesteps 上做迭代,将当前 timestep 的 input_features 与前一步得到的状态结合算出这一步的输出,然后将这个输出保存为新的状态供下一步使用。第一步时,没有状态,因此将状态初始化为一个全零向量,称为网络的初始状态。
伪代码:
```python
state_t = 0
for input_t in input_sequence:
output_t = f(input_t, state_t)
state_t = output_t
```
这里的 `f(...)` 其实和我们的 Dense 层比较类似,但这里不仅处理输出,还要同时加入状态的影响。所以它就需要包含 3 个参数:分别作用与输出和状态的矩阵 W、U,以及偏移向量 b:
```python
def f(input_t, state_t):
return activation(
dot(W, input_t) + dot(U, state_t) + b
)
```
画个图来表示这个程序:

下面把它写成真实的代码:
```
import numpy as np
# 定义各种维度大小
timesteps = 100
input_features = 32
output_features = 64
inputs = np.random.random((timesteps, input_features))
state_t = np.zeros((output_features))
W = np.random.random((output_features, input_features))
U = np.random.random((output_features, output_features))
b = np.random.random((output_features))
successive_outputs = []
for input_t in inputs: # input_t: (input_features, )
output_t = np.tanh( # output_t: (output_features, )
np.dot(W, input_t) + np.dot(U, state_t) + b
)
successive_outputs.append(output_t)
state_t = output_t
final_output_sequence = np.stack(successive_outputs, axis=0) # (timesteps, output_features)
print(successive_outputs[-1].shape)
print(final_output_sequence.shape)
```
在这里,我们最终输出是一个形状为 (timesteps, output_features) ,是所有 timesteps 的结果拼起来的。但实际上,我们一般只用最后一个结果 `successive_outputs[-1]` 就行了,这个里面已经包含了之前所有步骤的结果,即包含了整个序列的信息。
### Keras 中的循环层
把刚才这个玩具版本再加工一下,让它能接收形状为 `(batch_size, timesteps, input_features)` 的输入,批量去处理,就得到了 keras 中的 `SimpleRNN` 层:
```python
from tensorflow.keras.layers import SimpleRNN
```
这个 SimpleRNN 层和 keras 中的其他循环层都有两种可选的输出模式:
| 输出形状 | 说明 | 使用 |
| --- | --- | --- |
| `(batch_size, timesteps, output_features)` | 输出每个 timestep 输出的完整序列 | return_sequences=True |
| `(batch_size, output_features)` | 只返回每个序列的最终输出 | return_sequences=False (默认) |
```
# 只返回最后一个时间步的输出
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Embedding, SimpleRNN
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32))
model.summary()
# 返回完整的状态序列
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32, return_sequences=True))
model.summary()
```
如果我们要堆叠使用多个 RNN 层的时候,中间的层必须返回完整的状态序列:
```
# 堆叠多个 RNN 层,中间层返回完整的状态序列
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32)) # 最后一层要最后一个输出就行了
model.summary()
```
接下来,我们尝试用 RNN 再次处理 IMDB 问题。首先,准备数据:
```
# 准备 IMDB 数据
from tensorflow.keras.datasets import imdb
from tensorflow.keras.preprocessing import sequence
max_features = 10000
maxlen = 500
batch_size = 32
print('Loading data...')
(input_train, y_train), (input_test, y_test) = imdb.load_data(num_words=max_features)
print(len(input_train), 'train sequences')
print(len(input_test), 'test sequences')
print('Pad sequences (samples x time)')
input_train = sequence.pad_sequences(input_train, maxlen=maxlen)
input_test = sequence.pad_sequences(input_test, maxlen=maxlen)
print('input_train shape:', input_train.shape)
print('input_train shape:', input_test.shape)
```
构建并训练网络:
```
# 用 Embedding 层和 SimpleRNN 层来训练模型
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Embedding, SimpleRNN, Dense
model = Sequential()
model.add(Embedding(max_features, 32))
model.add(SimpleRNN(32))
model.add(Dense(1, activation='sigmoid'))
model.summary()
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(input_train, y_train,
epochs=10,
batch_size=128,
validation_split=0.2)
```
绘制训练过程看看:
```
# 绘制结果
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo-', label='Training acc')
plt.plot(epochs, val_acc, 'rs-', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'rs-', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
```
Emmmm,其实吧,这个模型的结果还没有第三章里面的用几个全连接层堆叠起来的模型好。原因有好几个,一个是我们这里只考虑了每个序列的前 500 个单词,还有一个是 SimpleRNN 其实并不擅长处理很长的序列。接下来,我们会看几个能表现的更好的循环层。
#### LSTM 层和 GRU 层
在 Keras 中的循环层,除了 SimpleRNN,还有更“不simple”一些的 LSTM 层和 GRU 层,后面这两种会更常用。
SimpleRNN 是有一些问题的,理论上,在遍历到时间步 t 的时候,它应该是能留存着之前许多步以来见过的信息的,但实际的应用中,由于某种叫做 vanishing gradient problem(梯度消失问题)的现象,它并不能学到这种长期依赖。
梯度消失问题其实在层数比较多的前馈网络里面也会有发生,主要表现就是随着层数多了之后,网络无法训练了。LSTM 层和 GRU 层就是对抗这种问题而生的。
**LSTM** 层是基于 LSTM (长短期记忆,long short-term memory) 算法的,这算法就是专门研究了处理梯度消失问题的。其实它的核心思想就是要保存信息以便后面使用,防止前面得到的信息在后面的处理中逐渐消失。
LSTM 在 SimpleRNN 的基础上,增加了一种跨越多个时间步传递信息的方法。这个新方法做的事情就像一条在序列旁边的辅助传送带,序列中的信息可以在任意位置跳上传送带, 然后被传送到更晚的时间步,并在需要时原封不动地跳回来。

这里把之前 SimpleRNN 里面的权重 W、U 重命名为 Wo、Uo 了(o 表示 output)。然后加了一个“携带轨道”数据流,这个携带轨道就是用来携带信息跨越时间步的。这个携带轨道上面放着时间步 t 的 ct 信息(c 表示 carry),这些信息将与输入、状态一起进行运算,而影响传递到下一个时间步的状态:
```pythoon
output_t = activation(dot(state_t, Uo) + dot(input_t, Wo) + dot(C_t, Vo) + bo)
i_t = activation(dot(state_t, Ui) + dot(input_t, Wi) + bi)
f_t = activation(dot(state_t, Uf) + dot(input_t, Wf) + bf)
k_t = activation(dot(state_t, Uk) + dot(input_t, Wk) + bk)
c_t_next = i_t * k_t + c_t * f_t
```
关于 LSTM 更多的细节、内部实现就不介绍了。咱完全不需要理解关于 LSTM 单元的具体架构,理解这东西就不是人干的事。我们只需要记住 LSTM 单元的作用: 允许把过去的信息稍后再次拿进来用,从而对抗梯度消失问题。
(P.S. 作者说这里是玄学,信他就行了。🤪 Emmm,这句是我胡翻的,原话是:“it may seem a bit arbitrary, but bear with me.”)
**GRU**(Gated Recurrent Unit, 门控循环单元),书上提的比较少,参考这篇 《[人人都能看懂的GRU](https://zhuanlan.zhihu.com/p/32481747)》,说 GRU 大概是 LSTM 的一种变种吧,二者原理区别不大、实际效果上也差不多。但 GRU 比 LSTM 新一些,它做了一些简化,更容易计算一些,但相应表示能力可能稍差一点点。
#### Keras 中使用 LSTM
我们还是继续用之前处理好的的 IMDB 数据来跑一个 LSTM:
```
from tensorflow.keras.layers import LSTM
model = Sequential()
model.add(Embedding(max_features, 32))
model.add(LSTM(32))
model.add(Dense(1, activation='sigmoid'))
model.summary()
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(input_train, y_train,
epochs=10,
batch_size=128,
validation_split=0.2)
# 绘制结果
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo-', label='Training acc')
plt.plot(epochs, val_acc, 'rs-', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'rs-', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
```
比 SimpleRNN 好多了。但也没比以前那种用全连接层的网络好多少,而且还比较慢(计算代价大),其实主要是由于情感分析这样的问题,用 LSTM 去分析全局的长期性结构帮助并不是很大,LSTM 擅长的是更复杂的自然语言处理问题,比如机器翻译。用全连接的方法,其实是做了看出现了哪些词及其出现频率,这个对这种简单问题还比较有效。
然后,我们再试试书上没提的 GRU:
```
# 把 LSTM 改成用 GRU
from tensorflow.keras.layers import GRU
model = Sequential()
model.add(Embedding(max_features, 32))
model.add(GRU(32))
model.add(Dense(1, activation='sigmoid'))
model.summary()
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(input_train, y_train,
epochs=10,
batch_size=128,
validation_split=0.2)
# 绘制结果
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo-', label='Training acc')
plt.plot(epochs, val_acc, 'rs-', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'rs-', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
```
区别不大。
| github_jupyter |
# Description
This notebook trains and evaluates the **inceptionv3** model.
# MOUNT GOOGLE Drive
```
from google.colab import drive
drive.mount('/content/gdrive')
```
# Change your working directory
```
cd /content/gdrive/My\ Drive/WIRE_DETECTION/TPU_COLAB/
```
# MODEL SPEC
```
model_name='inceptionv3'
iden='model2'
```
# TPU CHECK
The model trains faster in TPU (approximately 17 times)
```
%tensorflow_version 2.x
import tensorflow as tf
print("Tensorflow version " + tf.__version__)
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver() # TPU detection
print('Running on TPU ', tpu.cluster_spec().as_dict()['worker'])
except ValueError:
raise BaseException('ERROR: Not connected to a TPU runtime;')
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
tpu_strategy = tf.distribute.experimental.TPUStrategy(tpu)
```
# FIXED PARAMETERS
```
from glob import glob
import os
BUCKET='tfalldata' # @param
TFIDEN='WireDTF' # @param
IMG_DIM=256 # @param
NB_CHANNEL=3 # @param
BATCH_SIZE=128 # @param
BUFFER_SIZE=2048 # @param
TRAIN_DATA=1024*21 # @param
EVAL_DATA=1024*2 # @param
EPOCHS=250 # @param
TOTAL_DATA=TRAIN_DATA+EVAL_DATA
STEPS_PER_EPOCH = TOTAL_DATA//BATCH_SIZE
EVAL_STEPS = EVAL_DATA//BATCH_SIZE
GCS_PATH='gs://{}/{}'.format(BUCKET,TFIDEN)
print(GCS_PATH)
WEIGHT_PATH=os.path.join(os.getcwd(),'model_weights','{}.h5'.format(iden))
if os.path.exists(WEIGHT_PATH):
print('FOUND PRETRAINED WEIGHTS')
LOAD_WEIGHTS=True
else:
print('NO PRETRAINED WEIGHTS FOUND')
LOAD_WEIGHTS=False
```
# Dataset wrapper with tf.data api
```
def data_input_fn(mode,BUFFER_SIZE,BATCH_SIZE,img_dim):
def _parser(example):
feature ={ 'image' : tf.io.FixedLenFeature([],tf.string) ,
'target' : tf.io.FixedLenFeature([],tf.string)
}
parsed_example=tf.io.parse_single_example(example,feature)
image_raw=parsed_example['image']
image=tf.image.decode_png(image_raw,channels=3)
image=tf.cast(image,tf.float32)/255.0
image=tf.reshape(image,(img_dim,img_dim,3))
target_raw=parsed_example['target']
target=tf.image.decode_png(target_raw,channels=1)
target=tf.cast(target,tf.float32)/255.0
target=tf.reshape(target,(img_dim,img_dim,1))
return image,target
gcs_pattern=os.path.join(GCS_PATH,mode,'*.tfrecord')
file_paths = tf.io.gfile.glob(gcs_pattern)
dataset = tf.data.TFRecordDataset(file_paths)
dataset = dataset.map(_parser)
dataset = dataset.shuffle(BUFFER_SIZE,reshuffle_each_iteration=True)
dataset = dataset.repeat()
dataset = dataset.batch(BATCH_SIZE,drop_remainder=True)
dataset = dataset.prefetch(tf.data.experimental.AUTOTUNE)
return dataset
eval_ds = data_input_fn("Eval",BUFFER_SIZE,BATCH_SIZE,IMG_DIM)
train_ds = data_input_fn("Train",BUFFER_SIZE,BATCH_SIZE,IMG_DIM)
for x,y in eval_ds.take(1):
print(x.shape)
print(y.shape)
```
# install segmentation-models
```
!pip3 install segmentation-models
```
# framework setup
```
import segmentation_models as sm
sm.set_framework('tf.keras')
```
# model creation
```
def ssim(y_true, y_pred):
return tf.reduce_mean(tf.image.ssim(y_true, y_pred, 1.0))
with tpu_strategy.scope():
model = sm.Unet(model_name,input_shape=(IMG_DIM,IMG_DIM,NB_CHANNEL), encoder_weights=None)
model.compile(optimizer="Adam",
loss=tf.keras.losses.mean_squared_error,
metrics=[ssim])
if LOAD_WEIGHTS:
model.load_weights(WEIGHT_PATH)
model.summary()
```
# Training
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# reduces learning rate on plateau
lr_reducer = tf.keras.callbacks.ReduceLROnPlateau(factor=0.1,
cooldown= 10,
patience=10,
verbose =1,
min_lr=0.1e-5)
mode_autosave = tf.keras.callbacks.ModelCheckpoint(WEIGHT_PATH,
monitor='val_ssim',
mode = 'max',
save_best_only=True,
verbose=1,
period =10)
# stop learining as metric on validatopn stop increasing
early_stopping = tf.keras.callbacks.EarlyStopping(patience=15,
verbose=1,
mode = 'auto')
callbacks = [mode_autosave, lr_reducer,early_stopping ]
history = model.fit(train_ds,
steps_per_epoch=STEPS_PER_EPOCH,
epochs=EPOCHS,
verbose=1,
validation_data=eval_ds,
validation_steps=EVAL_STEPS,
callbacks=callbacks)
# save model
model.save_weights(WEIGHT_PATH)
def plot_history(history):
"""
Plots model training history
"""
fig, (ax_loss, ax_acc) = plt.subplots(1, 2, figsize=(15,5))
ax_loss.plot(history.epoch, history.history["loss"], label="Train loss")
ax_loss.plot(history.epoch, history.history["val_loss"], label="Validation loss")
ax_loss.legend()
ax_acc.plot(history.epoch, history.history["ssim"], label="Train ssim")
ax_acc.plot(history.epoch, history.history["val_ssim"], label="Validation ssim")
ax_acc.legend()
plt.show()
# show history
plot_history(history)
```
# Model Predictions and Scores
```
from skimage.measure import compare_ssim
from sklearn.metrics import jaccard_similarity_score
from glob import glob
from PIL import Image as imgop
import cv2
import imageio
img_dir = os.path.join(os.getcwd(),'test','images')
tgt_dir = os.path.join(os.getcwd(),'test','masks')
def create_dir(base_dir,ext_name):
'''
creates a new dir with ext_name in base_dir and returns the path
'''
new_dir=os.path.join(base_dir,ext_name)
if not os.path.exists(new_dir):
os.mkdir(new_dir)
return new_dir
pred_path=create_dir(os.path.join(os.getcwd(),'test'),'preds')
pred_dir=create_dir(pred_path,iden)
# preprocess data
def get_img(_path):
data=imgop.open(_path)
data=data.resize((IMG_DIM,IMG_DIM))
data=np.array(data)
data=data.astype('float32')/255.0
data=np.expand_dims(data,axis=0)
return data
def get_gt(_path):
# test folder mask path
_mpath=str(_path).replace("images","masks")
# ground truth
gt=cv2.imread(_mpath,0)
# resize
gt= cv2.resize(gt,(IMG_DIM,IMG_DIM), interpolation = cv2.INTER_AREA)
# Otsu's thresholding after Gaussian filtering
blur = cv2.GaussianBlur(gt,(5,5),0)
_,gt = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
return gt
def get_pred(model,img,_path):
_ppath=str(_path).replace('images','preds/{}'.format(iden))
pred=model.predict([img])
pred =np.squeeze(pred)*255.0
pred=pred.astype('uint8')
imageio.imsave(_ppath,pred)
pred=cv2.imread(_ppath,0)
pred= cv2.resize(pred,(IMG_DIM,IMG_DIM), interpolation = cv2.INTER_AREA)
# Otsu's thresholding after Gaussian filtering
blur = cv2.GaussianBlur(pred,(5,5),0)
_,pred = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
return pred
def get_overlay(pred,img):
img=np.squeeze(img)
overlay=img*0.2
xs,ys=np.nonzero(pred)
for x,y in zip(xs,ys):
overlay[x,y,:]=img[x,y,:]
return overlay
def get_score(pred,gt):
(ssim_score,_) = compare_ssim(gt,pred,full=True)
iou = jaccard_similarity_score(gt.flatten(), pred.flatten())
return ssim_score,iou
def score_summary(arr,model_name,score_iden):
print(model_name,':',score_iden)
print('max:',np.amax(arr))
print('mean:',np.mean(arr))
print('min:',np.amin(arr))
# plotting data
def plot_data(img,gt,pred,overlay):
fig, (ax1, ax2, ax3,ax4) = plt.subplots(1, 4,figsize=(20,20))
ax1.imshow(np.squeeze(img))
ax1.title.set_text('image')
ax2.imshow(np.squeeze(gt))
ax2.title.set_text('ground truth')
ax3.imshow(np.squeeze(pred))
ax3.title.set_text('prediction')
ax4.imshow(np.squeeze(overlay))
ax4.title.set_text('Overlay')
plt.show()
img_paths=glob(os.path.join(img_dir,'*.*'))
SSIM=[]
IOU=[]
# inference model
model_infer = sm.Unet(model_name,input_shape=(IMG_DIM,IMG_DIM,NB_CHANNEL), encoder_weights=None)
model_infer.load_weights(WEIGHT_PATH)
print('Loaded inference weights')
for _path in img_paths:
# ground truth
gt=get_gt(_path)
# image
img=get_img(_path)
# prediction
pred=get_pred(model_infer,img,_path)
# overlay
overlay=get_overlay(pred,img)
# scores
ssim_score,iou=get_score(pred,gt)
SSIM.append(ssim_score)
IOU.append(iou)
plot_data(img,gt,pred,overlay)
```
# Evaluation Scores
```
score_summary(np.array(SSIM),model_name,'ssim')
score_summary(np.array(IOU),model_name,'IoU/F1')
```
| github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import os
import glob
import pickle
import numpy as np
import numpy.lib.recfunctions as rfn
import matplotlib.pyplot as plt
from matplotlib.lines import Line2D
import scipy.integrate, scipy.interpolate, scipy.stats
from astropy.table import Table
from sklearn.linear_model import LinearRegression
from ltsfit import lts_linefit
from colossus.halo import mass_defs
from colossus.halo import concentration
from colossus.cosmology import cosmology
```
- Some of the calculation here also depends on the `awesome_cluster_finder` package by Christopher Bradshaw
- It will be available [in this Github repo](https://github.com/Christopher-Bradshaw/awesome_cluster_finder)
- If you don't have access to `acf` or don't have space for downloading the data, you can load the saved data in this folder to reproduce the figure.
```
import awesome_cluster_finder as acf
import jianbing
from jianbing import scatter
fig_dir = jianbing.FIG_DIR
data_dir = jianbing.DATA_DIR
sim_dir = jianbing.SIM_DIR
bin_dir = jianbing.BIN_DIR
res_dir = jianbing.RES_DIR
```
### Estimate the $\sigma_{\mathcal{O}}$
#### M*100
```
# Assuming alpha=0.2
# Bin 1
print(np.round(scatter.sigo_to_sigm(0.20, alpha=0.35), 2))
# Bin 2
print(np.round(scatter.sigo_to_sigm(0.35, alpha=0.35), 2))
# Bin 3
print(np.round(scatter.sigo_to_sigm(0.35, alpha=0.35), 2))
```
#### M*[50, 100]
```
# Bin 1
print(np.round(scatter.sigo_to_sigm(0.30, alpha=0.66), 2))
# Bin 2
print(np.round(scatter.sigo_to_sigm(0.42, alpha=0.66), 2))
# Bin 2
print(np.round(scatter.sigo_to_sigm(0.44, alpha=0.66), 2))
```
#### Richness
```
# Bin 1
print(np.round(scatter.sigo_to_sigm(0.26, alpha=0.86), 2))
# Bin 2
print(np.round(scatter.sigo_to_sigm(0.38, alpha=0.86), 2))
```
### UniverseMachine: logMvir v.s. in-situ & ex-situ stellar mass
```
um_cat = np.load(
'/Users/song/Dropbox/work/project/asap/data/umachine/um_smdpl_insitu_exsitu_0.7124_basic_logmp_11.5.npy')
um_cen = um_cat[um_cat['upid'] == -1]
logm_ins = um_cen['logms_gal']
logm_exs = um_cen['logms_icl']
logmh_vir = um_cen['logmh_vir']
```
#### Ex-situ
```
mask = um_cen['logmh_vir'] >= 13.6
x_err = np.full(len(um_cen[mask]), 0.03)
y_err = np.full(len(um_cen[mask]), 0.04)
w = 1. / (y_err ** 2.)
x_arr, y_arr = um_cen[mask]['logmh_vir'], um_cen[mask]['logms_icl']
reg = LinearRegression().fit(
x_arr.reshape(-1, 1), y_arr, sample_weight=w)
print(reg.coef_, reg.intercept_)
plt.scatter(x_arr, y_arr, s=2)
x_grid = np.linspace(13.6, 15.2, 100)
plt.plot(x_grid, reg.coef_ * x_grid + reg.intercept_, linewidth=2.0, linestyle='--', c='k')
lts_linefit.lts_linefit(x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
scatter.sigo_to_sigm(0.25, alpha=0.867)
```
#### In-situ
```
mask = um_cen['logmh_vir'] >= 14.0
x_err = np.full(len(um_cen[mask]), 0.03)
y_err = np.full(len(um_cen[mask]), 0.04)
w = 1. / (y_err ** 2.)
x_arr, y_arr = um_cen[mask]['logmh_vir'], um_cen[mask]['logms_gal']
reg = LinearRegression().fit(
x_arr.reshape(-1, 1), y_arr, sample_weight=w)
print(reg.coef_, reg.intercept_)
plt.scatter(x_arr, y_arr, s=2)
x_grid = np.linspace(13.8, 15.2, 100)
plt.plot(x_grid, reg.coef_ * x_grid + reg.intercept_, linewidth=2.0, linestyle='--', c='k')
plt.ylim(8.9, 12.2)
lts_linefit.lts_linefit(x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
scatter.sigo_to_sigm(0.37, alpha=0.141)
```
### Ex-situ mass in Illustris and IllustrisTNG
```
def exsitu_frac_sum(gal):
"""Summarize the ex-situ fraction of a galaxy."""
summary = {}
# Central flag
summary['cen'] = gal['info']['cen_flag']
# Total stellar mass
summary['logms'] = gal['info']['logms']
# Total halo mass
summary['logm_200c'] = gal['info']['logm200c']
c_200c = concentration.concentration(
10.0 ** summary['logm_200c'], '200c', 0.4, model='diemer19')
mvir, rvir, cvir = mass_defs.changeMassDefinition(
10.0 ** summary['logm_200c'], c_200c, 0.4, '200c', 'vir')
summary['logm_vir'] = np.log10(mvir)
summary['logm_ins'] = gal['info']['logms_map_ins']
summary['logm_exs'] = gal['info']['logms_map_exs']
# Total ex-situ fraction
summary['fexs_tot'] = (10.0 ** gal['info']['logms_map_exs'] / 10.0 ** gal['info']['logms_map_gal'])
# 5kpc, 10kpc, 100kpc stellar mass
summary['logms_5'] = np.log10(gal['aper']['maper_gal'][6])
summary['logms_10'] = np.log10(gal['aper']['maper_gal'][9])
summary['logms_30'] = np.log10(gal['aper']['maper_gal'][12])
summary['logms_40'] = np.log10(gal['aper']['maper_gal'][13])
summary['logms_60'] = np.log10(gal['aper']['maper_gal'][14])
summary['logms_100'] = np.log10(gal['aper']['maper_gal'][16])
summary['logms_150'] = np.log10(gal['aper']['maper_gal'][17])
summary['logms_30_100'] = np.log10(gal['aper']['maper_gal'][16] - gal['aper']['maper_gal'][12])
summary['logms_40_100'] = np.log10(gal['aper']['maper_gal'][16] - gal['aper']['maper_gal'][13])
summary['logms_60_100'] = np.log10(gal['aper']['maper_gal'][16] - gal['aper']['maper_gal'][14])
summary['logms_30_150'] = np.log10(gal['aper']['maper_gal'][17] - gal['aper']['maper_gal'][12])
summary['logms_40_150'] = np.log10(gal['aper']['maper_gal'][17] - gal['aper']['maper_gal'][13])
summary['logms_60_150'] = np.log10(gal['aper']['maper_gal'][17] - gal['aper']['maper_gal'][14])
# Mass fraction in 5 and 10 kpc
summary['fmass_5'] = gal['aper']['maper_gal'][6] / gal['aper']['maper_gal'][16]
summary['fmass_10'] = gal['aper']['maper_gal'][9] / gal['aper']['maper_gal'][16]
# Ex-situ fraction within 5, 10, 100 kpc
summary['fexs_5'] = gal['aper']['maper_exs'][6] / gal['aper']['maper_gal'][6]
summary['fexs_10'] = gal['aper']['maper_exs'][9] / gal['aper']['maper_gal'][9]
summary['fexs_100'] = gal['aper']['maper_exs'][9] / gal['aper']['maper_gal'][9]
# In-situ and ex-situ mass profile
summary['rad'] = gal['aper']['rad_mid']
summary['mprof_ins'] = gal['aper']['mprof_ins']
summary['mprof_exs'] = gal['aper']['mprof_exs']
return summary
```
#### Illustris @ z=0.4
```
data_dir = '/Volumes/astro6/massive/simulation/riker/ori/sum'
xy_list = glob.glob(os.path.join(data_dir, '*xy_sum.npy'))
xy_sum = [np.load(gal, allow_pickle=True) for gal in xy_list]
print("# There are %d Illustris massive galaxies" % len(xy_list))
ori_cat = Table([exsitu_frac_sum(gal) for gal in xy_sum])
mask = ori_cat['cen'] & (ori_cat['logm_vir'] >= 13.2)
x_arr = ori_cat['logm_vir'][mask]
y_arr = ori_cat['logm_exs'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.01)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = ori_cat['cen'] & (ori_cat['logm_vir'] >= 13.2)
x_arr = ori_cat['logm_vir'][mask]
y_arr = ori_cat['logm_ins'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.01)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = ori_cat['cen'] & (ori_cat['logm_vir'] >= 13.2)
x_arr = ori_cat['logm_vir'][mask]
y_arr = ori_cat['logms_40_100'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.02)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = ori_cat['cen'] & (ori_cat['logm_vir'] >= 13.2)
x_arr = ori_cat['logm_vir'][mask]
y_arr = ori_cat['logms_100'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.02)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = ori_cat['cen'] & (ori_cat['logm_vir'] >= 13.2)
x_arr = ori_cat['logm_vir'][mask]
y_arr = ori_cat['logms_10'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.02)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
```
#### TNG100 @ z=0.4
```
data_dir = '/Volumes/astro6/massive/simulation/riker/tng/sum'
xy_list = glob.glob(os.path.join(data_dir, '*xy_sum.npy'))
xy_sum = [np.load(gal, allow_pickle=True) for gal in xy_list]
print("# There are %d TNG massive galaxies" % len(xy_list))
tng_cat = Table([exsitu_frac_sum(gal) for gal in xy_sum])
mask = tng_cat['cen'] & (tng_cat['logm_vir'] >= 13.2)
x_arr = tng_cat['logm_vir'][mask]
y_arr = tng_cat['logm_exs'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.01)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = tng_cat['cen'] & (tng_cat['logm_vir'] >= 13.2)
x_arr = tng_cat['logm_vir'][mask]
y_arr = tng_cat['logm_ins'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.01)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = tng_cat['cen'] & (tng_cat['logm_vir'] >= 13.2)
x_arr = tng_cat['logm_vir'][mask]
y_arr = tng_cat['logms_40_100'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.02)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = tng_cat['cen'] & (tng_cat['logm_vir'] >= 13.2)
x_arr = tng_cat['logm_vir'][mask]
y_arr = tng_cat['logms_100'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.02)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
mask = tng_cat['cen'] & (tng_cat['logm_vir'] >= 13.2)
x_arr = tng_cat['logm_vir'][mask]
y_arr = tng_cat['logms_10'][mask]
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.02)
lts_linefit.lts_linefit(
x_arr, y_arr, x_err, y_err, pivot=np.nanmedian(x_arr), clip=4.0)
# Illustris
logm_exs_ori = 11.528 + 0.800 * (logmh_grid - 13.44)
sigm_exs_ori = 0.21
logm_ins_ori = 11.384 + 0.360 * (logmh_grid - 13.44)
sigm_ins_ori = 0.14
logm_out_ori = 10.994 + 0.920 * (logmh_grid - 13.44)
sigm_out_ori = 0.21
logm_100_ori = 11.740 + 0.594 * (logmh_grid - 13.44)
sigm_100_ori = 0.14
logm_10_ori = 11.444 + 0.389 * (logmh_grid - 13.44)
sigm_10_ori = 0.12
# TNG
logm_exs_tng = 11.437 + 0.784 * (logmh_grid - 13.48)
sigm_exs_tng = 0.19
logm_ins_tng = 11.182 + 0.543 * (logmh_grid - 13.48)
sigm_ins_tng = 0.16
logm_out_tng = 10.860 + 0.921 * (logmh_grid - 13.48)
sigm_out_tng = 0.21
logm_100_tng = 11.610 + 0.660 * (logmh_grid - 13.48)
sigm_100_tng = 0.15
logm_10_tng = 11.296 + 0.541 * (logmh_grid - 13.48)
sigm_10_tng = 0.13
```
### Assign CAMIRA richness to ASAP halos
- Based on the Mvir-richness relation of CAMIRA clusters from Murata et al. (2019)
- $P(\ln N \mid M, z)=\frac{1}{\sqrt{2 \pi} \sigma_{\ln N \mid M, z}} \exp \left(-\frac{x^{2}(N, M, z)}{2 \sigma_{\ln N \mid M, z}^{2}}\right)$
- $\begin{aligned} x(N, M, z) & \equiv \ln N-\left[A+B \ln \left(\frac{M}{M_{\text {pivot }}}\right)\right.\\ & \left.+B_{z} \ln \left(\frac{1+z}{1+z_{\text {pivot }}}\right)+C_{z}\left[\ln \left(\frac{1+z}{1+z_{\text {pivot }}}\right)\right]^{2}\right] \end{aligned}$
- $\begin{aligned} \sigma_{\ln N \mid M, z} &=\sigma_{0}+q \ln \left(\frac{M}{M_{\text {pivot }}}\right) \\ &+q_{z} \ln \left(\frac{1+z}{1+z_{\text {pivot }}}\right)+p_{z}\left[\ln \left(\frac{1+z}{1+z_{\text {pivot }}}\right)\right]^{2} \end{aligned}$
- Parameters for low-z ($0.1 < z < 0.4$) clusters using Planck cosmology:
- $A = 3.34^{+0.25}_{-0.20}$
- $B = 0.85^{+0.08}_{-0.07}$
- $\sigma_0 = 0.36^{+0.07}_{-0.21}$
- $q = -0.06^{0.09}_{-0.11}$
- Parameters for full redshift range using Planck cosmology:
- $A = 3.15^{+0.07}_{-0.08}$
- $B = 0.86^{+0.05}_{-0.05}$
- $B_{z} = -0.21^{+0.35}_{-0.42}$
- $C_{z} = 3.61^{+1.96}_{-2.23}$
- $\sigma_0 = 0.32^{+0.06}_{-0.06}$
- $q = -0.06^{+0.09}_{-0.11}$
- $q_{z} = 0.03^{+0.31}_{-0.30}$
- $p_{z} = 0.70^{+1.71}_{-1.60}$
- Pivot redshift and mass
- $M_{\rm Pivot} = 3\times 10^{14} h^{-1} M_{\odot}$
- $z_{\rm Pivot} = 0.6$
- Here, $M \equiv M_{200m}$ and $h=0.68$.
```
def mean_ln_N(m200m, z=None, m_pivot=3e14, h=0.68, A=3.15, B=0.86,
z_pivot=0.6, B_z=-0.21, C_z=3.61):
"""
Estimate the mean ln(N) for CAMIRA clusters based on the halo mass-richness
relation calibrated by Murata et al. (2019).
"""
lnN = A + B * np.log(m200m / m_pivot / h)
if z is None:
return lnN
z_term = np.log((1 + z) / (1 + z_pivot))
return lnN + B_z * z_term + C_z * (z_term ** 2)
def sig_ln_N(m200m, z=None, m_pivot=3e14, h=0.68, sig0=0.32, z_pivot=0.6,
q=-0.06, q_z=0.03, p_z=0.70):
"""
Estimate the scatter of ln(N) for CAMINRA clusters based on the halo mass-richness
relation calibrated by Murata et al. (2019).
"""
sig_lnN = sig0 + q * np.log(m200m / m_pivot / h)
if z is None:
return sig_lnN
z_term = np.log((1 + z) / (1 + z_pivot))
return sig_lnN + q_z * z_term + p_z * (z_term ** 2)
lnN = np.random.normal(
loc=mean_ln_N(um_cen['m200b_hlist'], z=0.4),
scale=sig_ln_N(um_cen['m200b_hlist'], z=0.4))
log10_N = np.log10(np.exp(lnN))
x_arr = np.log10(um_cen['mvir'])
y_arr = log10_N
mask = x_arr >= 14.0
x_err = np.full(len(x_arr), 0.02)
y_err = np.full(len(y_arr), 0.01)
reg = LinearRegression().fit(
x_arr[mask].reshape(-1, 1), y_arr[mask])
print(reg.coef_, reg.intercept_)
plt.scatter(np.log10(um_cen['mvir']), log10_N, s=2, alpha=0.1)
plt.xlim(13.8, 15.2)
plt.ylim(0.1, 2.6)
lts_linefit.lts_linefit(
x_arr[mask], y_arr[mask], x_err[mask], y_err[mask], pivot=np.nanmedian(x_arr[mask]), clip=4.0)
logmh_grid = np.linspace(13.6, 15.3, 30)
# Relation for M*ex-situ
logm_exs = 11.7441 + 0.867 * (logmh_grid - 14.17)
sigm_exs = 0.25
# Relation for M*in-situ
logm_ins = 11.0242 + 0.141 * (logmh_grid - 14.17)
sigm_ins = 0.37
# Relation for M*[50-100]
logm_out = 10.7474 + 0.66 * (logmh_grid - 13.77)
sigm_out = 0.3
# Relation for richness
nmem_cam = 1.1615 + 0.864 * (logmh_grid - 14.17)
sign_cam = 0.16
fig = plt.figure(figsize=(7.2, 10))
fig.subplots_adjust(
left=0.175, bottom=0.09, right=0.855, top=0.99, wspace=0, hspace=0)
ax1 = fig.add_subplot(2, 1, 1)
ax1.fill_between(
logmh_grid, logm_exs - sigm_exs, logm_exs + sigm_exs,
alpha=0.3, edgecolor='none', linewidth=1.0,
label=r'__no_label__', facecolor='skyblue', linestyle='-', rasterized=True)
l1 = ax1.plot(logmh_grid, logm_exs, linestyle='-', linewidth=5, alpha=0.7,
label=r'$\rm UM\ ex\ situ$', color='dodgerblue', zorder=100)
ax1.fill_between(
logmh_grid, logm_out - sigm_out + 0.52, logm_out + sigm_out + 0.52,
alpha=0.3, edgecolor='none', linewidth=1.0,
label=r'__no_label__', facecolor='grey', linestyle='-', rasterized=True)
l2 = ax1.plot(logmh_grid, logm_out + 0.52, linestyle='-.', linewidth=5, alpha=0.8,
label=r'$\rm M_{\star,[50,100]} + 0.5\ \rm dex$', color='grey', zorder=100)
ax1.set_ylabel(r'$\log_{10} (M_{\star}/M_{\odot})$', fontsize=32)
#------------------------------------------------------------------------------------#
ax2=ax1.twinx()
ax2.yaxis.label.set_color('orangered')
ax2.tick_params(axis='y', colors='orangered', which='both')
ax2.spines['right'].set_color('red')
ax2.set_ylabel(r"$\log_{10} N_{\rm CAMIRA}$", color="red", fontsize=32)
ax2.fill_between(
logmh_grid, nmem_cam - sign_cam, nmem_cam + sign_cam,
alpha=0.2, edgecolor='none', linewidth=2.0, zorder=0,
label=r'__no_label__', facecolor='red', linestyle='-', rasterized=True)
l3 = ax2.plot(logmh_grid, nmem_cam, linestyle='--', linewidth=5, alpha=0.7,
label=r'$\rm N_{\rm CAMIRA}$', color='red', zorder=100)
ax2.set_ylim(0.45, 2.3)
ax1.set_ylim(11.01, 12.95)
ax1.set_xticklabels([])
custom_lines = [Line2D([0], [0], color="dodgerblue", lw=4, ls='-'),
Line2D([0], [0], color="grey", lw=4, ls='-.'),
Line2D([0], [0], color="red", lw=4, ls='--')]
ax1.legend(custom_lines,
[r'$\rm UM\ ex\ situ$', r'$M_{\star,[50,100]} + 0.5\ \rm dex$',
r'$\rm N_{\rm CAMIRA}$'],
loc='best', fontsize=18)
#------------------------------------------------------------------------------------#
ax3 = fig.add_subplot(2, 1, 2)
# Universe Machine
ax3.fill_between(
logmh_grid, logm_ins - sigm_ins, logm_ins + sigm_ins,
alpha=0.1, edgecolor='none', linewidth=1.0, zorder=0,
label=r'__no_label__', facecolor='orange', linestyle='-', rasterized=True)
l1 = ax3.plot(logmh_grid, logm_ins, linestyle='-', linewidth=5, alpha=0.7,
label=r'$\rm UM\ ins$', color='orangered', zorder=100)
ax3.fill_between(
logmh_grid, logm_exs - sigm_exs, logm_exs + sigm_exs,
alpha=0.1, edgecolor='none', linewidth=1.0, zorder=0,
label=r'__no_label__', facecolor='dodgerblue', linestyle='-', rasterized=True)
l2 = ax3.plot(logmh_grid, logm_exs, linestyle='-', linewidth=5, alpha=0.7,
label=r'$\rm UM\ exs$', color='dodgerblue')
# Illustris
ax3.fill_between(
logmh_grid, logm_ins_ori - sigm_ins_ori, logm_ins_ori + sigm_ins_ori,
alpha=0.1, edgecolor='none', linewidth=1.0, zorder=0,
label=r'__no_label__', facecolor='coral', linestyle='--', rasterized=True)
l3 = ax3.plot(logmh_grid, logm_ins_ori, linestyle='-.', linewidth=6, alpha=0.6,
label=r'$\rm Illustris\ ins$', color='coral', zorder=100)
ax3.fill_between(
logmh_grid, logm_exs_ori - sigm_exs_ori, logm_exs_ori + sigm_exs_ori,
alpha=0.05, edgecolor='none', linewidth=1.0, zorder=0,
label=r'__no_label__', facecolor='steelblue', linestyle='--', rasterized=True)
l4 = ax3.plot(logmh_grid, logm_exs_ori, linestyle='-.', linewidth=5, alpha=0.7,
label=r'$\rm Illustris\ exs$', color='steelblue', zorder=100)
# TNG
ax3.fill_between(
logmh_grid, logm_ins_tng - sigm_ins_tng, logm_ins_tng + sigm_ins_tng,
alpha=0.1, edgecolor='none', linewidth=1.0, zorder=0,
label=r'__no_label__', facecolor='goldenrod', linestyle='-', rasterized=True)
l3 = ax3.plot(logmh_grid, logm_ins_tng, linestyle='--', linewidth=6, alpha=0.7,
label=r'$\rm TNG\ ins$', color='goldenrod')
ax3.fill_between(
logmh_grid, logm_exs_tng - sigm_exs_tng, logm_exs_tng + sigm_exs_tng,
alpha=0.05, edgecolor='grey', linewidth=1.0,
label=r'__no_label__', facecolor='royalblue', linestyle='--', rasterized=True)
l4 = ax3.plot(logmh_grid, logm_exs_tng, linestyle='--', linewidth=6, alpha=0.5,
label=r'$\rm TNG\ exs$', color='royalblue')
ax3.legend(loc='best', ncol=2, fontsize=16, handletextpad=0.5, labelspacing=0.3)
ax1.set_xlim(13.59, 15.25)
ax2.set_xlim(13.59, 15.25)
ax3.set_xlim(13.59, 15.25)
ax3.set_ylim(10.61, 13.45)
ax3.set_xlabel(r'$\log_{10} (M_{\rm vir}/ M_{\odot})$', fontsize=32)
ax3.set_ylabel(r"$\log_{10} (M_{\star}/ M_{\odot})$", fontsize=32)
fig.savefig(os.path.join(fig_dir, 'fig_13.png'), dpi=120)
fig.savefig(os.path.join(fig_dir, 'fig_13.pdf'), dpi=120)
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive/')
# Import required Libraries
!pip install fastparquet
import math
import pandas_datareader as web
import numpy as np
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, LSTM
import matplotlib.pyplot as plt
import datetime
stocks = pd.read_parquet('https://github.com/alik604/CMPT-419/blob/master/data/wsb.daily.joined.parquet.gz?raw=true', engine="fastparquet")
stocks.head(5)
stocks.tail(5)
# Drop created date
data = stocks.drop(['created_utc', 'AAPL_CPrc','AMZN_CPrc','BA_CPrc', 'SPY_CPrc', 'TSLA_CPrc' ] , axis=1)
# Closing Price dataset
plotData = stocks.filter(['created_utc','AAPL_CPrc', 'AMZN_CPrc', 'BA_CPrc', 'SPY_CPrc', 'TSLA_CPrc'])
dataCPrc = stocks.filter(['AAPL_CPrc', 'AMZN_CPrc', 'BA_CPrc', 'SPY_CPrc', 'TSLA_CPrc'])
datasetCPrc = dataCPrc.values
dataset = data.values
print(dataset)
# set training set len
training_data_len = math.ceil(len(dataset) * 0.8)
training_data_len
# Scale data
scaler = MinMaxScaler(feature_range=(0,1))
scaled_data = scaler.fit_transform(dataset)
scalerCPrc = MinMaxScaler(feature_range=(0,1))
scaledCPrc_data = scalerCPrc.fit_transform(datasetCPrc)
scaledCPrc_data
# Create training set from scaled data
train_data = scaled_data[0:training_data_len, :]
train_y = scaledCPrc_data[0:training_data_len, :]
# Split data into x_train, y_train
x_train = []
y_train = []
for i in range(60, len(train_data)):
x_train.append(train_data[i-60:i, :])
y_train.append(train_y[i:i+1,:][0].tolist())
if(i < 65):
print(train_y[i:i+1,:][0], type(train_y[i:i+1,:][0]))
# Convert both x, y training sets to np array
x_train, y_train = np.array(x_train), np.array(y_train)
print(y_train, type(y_train))
# Reshape the data // LSTM network expects 3 dimensional input in the form of
# (number of samples, number of timesteps, number of features)
x_train = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 33))
x_train.shape
# Declare the LSTM model architecture
model = Sequential()
# Add layers , input shape expected is (number of timesteps, number of features)
model.add(LSTM(75, return_sequences=True, input_shape=(x_train.shape[1], 33)))
model.add(LSTM(50, return_sequences=False))
model.add(Dense(25))
model.add(Dense(5))
# Compile Model
model.compile(optimizer='adam', loss='mean_squared_error')
```
```
# Train the model
model.fit(x_train, y_train, batch_size=16, epochs=100)
# Create the testing data set
test_data = scaled_data[training_data_len - 60: , :]
test_y = scaledCPrc_data[training_data_len-60:,:]
# Create the data sets x_test and y_test
x_test = []
y_test = []
for i in range(60, len(test_data)):
x_test.append(test_data[i-60:i, :])
y_test.append(train_y[i:i+1,:][0].tolist())
print(len(test))
# Convert the data into a np array
x_test = np.array(x_test)
# Reshape data into 3 dimensions ( num samples, timesteps, num features )
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1], 33))
# Get the models predicted price values
predictions = model.predict(x_test)
# Inverse transform
predictions = scalerCPrc.inverse_transform(predictions)
# Evaluate model with root mean square error (RMSE)
rmse = np.sqrt(np.mean((predictions-y_test)**2))
rmse
# Set up ploting dataset
plotData = stocks.filter(['created_utc','AAPL_CPrc', 'AMZN_CPrc', 'BA_CPrc', 'SPY_CPrc', 'TSLA_CPrc'])
plotData = plotData.set_index("created_utc")
print(plotData, type(plotData))
# This is so it graphs nicely with dates
AAPL = plotData["AAPL_CPrc"]
AAPL = AAPL.to_frame()
AMZN = plotData["AMZN_CPrc"]
AMZN = AMZN.to_frame()
BA = plotData["BA_CPrc"]
BA = BA.to_frame();
SPY = plotData["SPY_CPrc"]
SPY = SPY.to_frame();
TSLA = plotData["TSLA_CPrc"]
TSLA = TSLA.to_frame()
appleTest = AAPL[0:training_data_len]
amazonTest = AMZN[0:training_data_len]
boeTest = BA[0:training_data_len]
spyTest = SPY[0: training_data_len]
teslaTest = TSLA[0: training_data_len]
appleValid = AAPL[training_data_len:]
amazonValid = AMZN[training_data_len:]
boeValid = BA[training_data_len:]
spyValid = SPY[training_data_len:]
teslaValid = TSLA[training_data_len:]
# print(appleValid)
appleValid['Predictions'] = predictions[:,:1]
amazonValid["Predictions"] = predictions[:,1:2]
boeValid['Predictions'] = predictions[:, 2:3]
spyValid["Predictions"] = predictions[:, 3:4]
teslaValid["Predictions"] = predictions[:, 4:5]
print(len(appleValid), len(predictions))
# Visualize data
plt.figure(figsize=(16,8))
plt.title('Prediction with reddit glove vectors')
plt.xlabel('Date', fontsize=18)
plt.ylabel('Close Price USD ($)', fontsize=18)
plt.plot(appleTest)
plt.plot(appleValid)
plt.plot(appleValid['Predictions'])
# plt.plot(amazonTest)
# plt.plot(amazonValid)
# plt.plot(amazonValid['Predictions'])
# plt.plot(boeTest)
# plt.plot(boeValid)
# plt.plot(boeValid['Predictions'])
# plt.plot(spyTest)
# plt.plot(spyValid)
# plt.plot(spyValid['Predictions'])
# plt.plot(teslaTest)
# plt.plot(teslaValid)
# plt.plot(teslaValid['Predictions'])
plt.legend(['Train', 'Validation', 'Prediction'])
plt.show()
# Lets train a model without reddit posts to see how it performs
opening_prices = stocks.filter(['created_utc','AAPL_OPrc', 'AMZN_OPrc', 'BA_OPrc', 'SPY_OPrc', 'TSLA_OPrc'])
opening_prices = opening_prices.set_index('created_utc')
# PlotData contains closing prices
opening_dataset = opening_prices.values
closing_dataset = plotData.values
print(opening_dataset)
print(closing_dataset)
# set training set len
train_data_len = math.ceil(len(opening_dataset) * 0.80)
openScaler = MinMaxScaler(feature_range=(0,1))
scaled_open_prices = openScaler.fit_transform(opening_dataset)
closeScaler = MinMaxScaler(feature_range=(0,1))
scaled_close_prices = closeScaler.fit_transform(closing_dataset)
o_train_data = scaled_open_prices[0:train_data_len, :]
c_train_data = scaled_close_prices[0:train_data_len,:]
print(o_train_data, len(o_train_data))
c_train_data
o_x_train = []
c_y_train = []
print(len(o_x_train))
for i in range(60, len(o_train_data)):
o_x_train.append(o_train_data[i-60:i, :])
c_y_train.append(c_train_data[i,:].tolist())
if(i<62):
print(c_train_data[i,:].tolist())
x_train2, y_train2 = np.array(o_x_train), np.array(c_y_train)
x_train2 = np.reshape(x_train2, (x_train2.shape[0], x_train2.shape[1], 5))
x_train2.shape
# Declare the LSTM model architecture
model2 = Sequential()
# Add layers , input shape expected is (number of timesteps, number of features)
model2.add(LSTM(75, return_sequences=True, input_shape=(x_train2.shape[1], 5)))
model2.add(LSTM(50, return_sequences=False))
model2.add(Dense(25))
model2.add(Dense(5))
# Compile Model
model2.compile(optimizer='adam', loss='mean_squared_error')
# Train the model
model2.fit(x_train2, y_train2, batch_size=16, epochs=100)
x_test_data = scaled_open_prices[train_data_len - 60:, :]
y_test_data = scaled_close_prices[train_data_len - 60:, :]
x_test2 = []
y_test2 = []
for i in range(60, len(x_test_data)):
x_test2.append(x_test_data[i-60:i, :])
y_test2.append(y_test_data[i:i+1,:][0].tolist())
x_test2 = np.array(x_test2)
x_test2 = np.reshape(x_test2, (x_test2.shape[0], x_test2.shape[1], 5))
# Get the models predicted price values
predictions2 = model2.predict(x_test2)
# Inverse transform
predictions2 = closeScaler.inverse_transform(predictions2)
# Evaluate model with root mean square error (RMSE)
rmse2 = np.sqrt(np.mean((predictions2-y_test2)**2))
rmse2
appleValid['Predictions2'] = predictions2[:,:1]
amazonValid["Predictions2"] = predictions2[:,1:2]
boeValid['Predictions2'] = predictions2[:, 2:3]
spyValid["Predictions2"] = predictions2[:, 3:4]
teslaValid["Predictions2"] = predictions2[:, 4:5]
# Visualize data
plt.figure(figsize=(16,8))
plt.title('Stock Prediction with and without Reddit word vector data \n (TSLA)')
plt.xlabel('Date', fontsize=18)
plt.ylabel('Close Price USD ($)', fontsize=18)
# plt.plot(appleTest)
# plt.plot(appleValid)
# plt.plot(appleValid['Predictions'], color='red')
# plt.plot(appleValid['Predictions2'], color='green')
# plt.plot(amazonTest)
# plt.plot(amazonValid)
# plt.plot(amazonValid['Predictions'], color='red')
# plt.plot(amazonValid['Predictions2'], color='green')
# plt.plot(boeTest)
# plt.plot(boeValid)
# plt.plot(boeValid['Predictions'], color='red')
# plt.plot(boeValid['Predictions2'], color='green')
# plt.plot(spyTest, label='Train')
# plt.plot(spyValid, label='Actual')
# plt.plot(spyValid['Predictions'], label='Prediction with vector', color='red')
# plt.plot(spyValid['Predictions2'], label='Prediction without vector', color='green')
plt.plot(teslaTest, label='Train')
plt.plot(teslaValid, label="Actual")
plt.plot(teslaValid['Predictions'], label="vector prediction", color='red')
plt.plot(teslaValid['Predictions2'], label="without vector prediction", color="green")
plt.legend(('Train', 'Validation', 'Prediction with vector', 'Prediction without'), title="Legend",)
plt.show()
```
| github_jupyter |
# Exploring the BBBC021 dataset
## A little background on high content imaging / screening
In a high-content screening / imaging assay, a cell line is treated with a number of different compounds (often on the order of 10k, 100k, or more molecules) for a given period of time,
and then the cells are [fixed](https://en.wikipedia.org/wiki/Fixation_(histology)) and stained with fluorescent dyes which visualize important cellular structures that are then imaged under a microscope.
Through this procedure, we can directly observe the impact of the given (drug) molecules on cellular morphology -
changes in cell and subcellular shape and structure.
The biophysical interaction by which a bioactive molecule exerts its effects on cells is known as its [mechanism of action (MoA)](https://en.wikipedia.org/wiki/Mechanism_of_action).
Different compounds with the same MoA will have similar effects on cellular morphology, which we should be able to detect in our screen.
Note that a molecule in fact may have more than one MoA - these ["dirty drugs"](https://en.wikipedia.org/wiki/Dirty_drug) may exhibit multiple effects on cellular processes in the assay simultaneously,
or effects may change based on dosage.
## Our dataset: BBBC021 from the Broad Bioimage Benchmark Collection
The [Broad Bioimage Benchmark Collection](https://bbbc.broadinstitute.org/) is a collection of open microscopy imaging datasets published by the [Broad Institute](https://www.broadinstitute.org/),
an MIT- and Harvard-affiliated research institute in Cambridge, MA, USA.
The [BBBC021 dataset](https://bbbc.broadinstitute.org/BBBC021) comprises a [high-content screening](https://en.wikipedia.org/wiki/High-content_screening) assay of [Human MCF-7 cells](https://en.wikipedia.org/wiki/MCF-7),
a very commonly used breast cancer cell line in biomedical research.
In the BBBC021 dataset, 3 structures have been stained: DNA, and the cytoskeletal proteins F-actin and B-tubulin,
which comprise actin filaments and microtubules, respectively.
```
%load_ext autoreload
%autoreload 2
import holoviews as hv
import numpy as np
from pybbbc import BBBC021
hv.extension("bokeh")
im_opts = hv.opts.Image(
aspect="equal",
tools=["hover"],
active_tools=["wheel_zoom"],
colorbar=True,
cmap="fire",
normalize=False,
)
rgb_opts = hv.opts.RGB(
aspect="equal",
active_tools=["wheel_zoom"],
)
hv.opts.defaults(im_opts, rgb_opts)
```
### Working with pybbbc
#### Constructing the BBBC021 object
When you create the `BBBC021` object, you can choose which images to include by selecting subsets with keyword arguments. For example:
```
from pybbbc import BBBC021
# Entire BBBC021 dataset, including unknown MoA
bbbc021_all = BBBC021()
# Just the images with known MoA
bbbc021_moa = BBBC021(moa=[moa for moa in BBBC021.MOA if moa != "null"])
```
`BBBC021` has a number of useful constant class attributes that describe the entirety of the dataset
(and can be accessed without creating an object):
* `IMG_SHAPE`
* `CHANNELS`
* `PLATES`
* `COMPOUNDS`
* `MOA`
These don't change with the subset of BBBC021 you have selected. On other other hand, these do:
* `moa`
* `compounds`
* `plates`
* `sites`
* `wells`
For example, `BBBC021.MOA` will give you a list of all the MoAs in the full dataset:
```
BBBC021.MOA
```
### Access an image and its metadata
Your initialized `BBBC021` object is indexable and has a length.
An index is the integer offset into the subset of BBBC021 you have selected.
```
print(f'Number of images in BBBC021: {len(bbbc021_all)}')
print(f'Number of images with known MoA: {len(bbbc021_moa)}')
```
What you get back from the object is a `tuple` of the given image followed by its associated metadata
in the form of a `namedtuple`:
```
image, metadata = bbbc021_moa[0]
plate, compound, image_idx = metadata # it can be unpacked like a regular `tuple`
print(f'{metadata=}\n\n{metadata.plate=}\n\n{metadata.compound=}')
```
### View the metadata `DataFrame`s
The metadata is compiled into two Pandas `DataFrame`s, `image_df` and `moa_df`,
which contain only metadata from the selected subset of the BBBC021 dataset.
`image_df` contains metadata information on an individual image level.
Each row corresponds to an image in the subset of BBBC021 you selected:
```
bbbc021_moa.image_df
```
`image_idx` corresponds to the absolute index of the image in the full BBBC021 dataset.
`relative_image_idx` is the index you would use to access the given image as in:
`image, metadata = your_bbbc021_obj[relative_image_idx]`
`moa_df` is a metadata `DataFrame` which provides you with all the compound-concentration pairs in the selected BBBC021 subset:
```
bbbc021_moa.moa_df
```
# Visualize all BBBC021 images
```
def make_layout(image_idx):
image, metadata = bbbc021_all[image_idx]
prefix = f"{metadata.compound.compound} @ {metadata.compound.concentration:.2e} μM, {metadata.compound.moa}"
plots = []
cmaps = ["fire", "kg", "kb"]
for channel_idx, im_channel in enumerate(image):
plot = hv.Image(
im_channel,
bounds=(0, 0, im_channel.shape[1], im_channel.shape[0]),
label=f"{prefix} | {bbbc021_all.CHANNELS[channel_idx]}",
).opts(cmap=cmaps[channel_idx])
plots.append(plot)
plots.append(
hv.RGB(
image.transpose(1, 2, 0),
bounds=(0, 0, im_channel.shape[1], im_channel.shape[0]),
label="Channel overlay",
)
)
return hv.Layout(plots).cols(2)
hv.DynamicMap(make_layout, kdims="image").redim.range(image=(0, len(bbbc021_all) - 1))
```
| github_jupyter |
## Binary structure classification used in tree building: Step 2. Feature-rich approach
Train models, save the best one.
Output:
- ``models/structure_predictor_baseline/*``
```
%load_ext autoreload
%autoreload 2
import os
import glob
import pandas as pd
import pickle
from utils.file_reading import read_edus, read_gold, read_negative, read_annotation
from utils.prepare_sequence import _prepare_sequence
from tqdm import tqdm_notebook as tqdm
random_state = 45
```
### Make a directory
```
import os
model_path = 'models/structure_predictor_baseline'
if not os.path.isdir(model_path):
os.path.mkdir(model_path)
```
### Prepare train/test sets
```
IN_PATH = 'data_structure'
train_samples = pd.read_pickle(os.path.join(IN_PATH, 'train_samples.pkl'))
dev_samples = pd.read_pickle(os.path.join(IN_PATH, 'dev_samples.pkl'))
test_samples = pd.read_pickle(os.path.join(IN_PATH, 'test_samples.pkl'))
drop_columns = ['snippet_x', 'snippet_y', 'category_id',
'snippet_x_tmp', 'snippet_y_tmp',
'filename', 'order', 'postags_x', 'postags_y',
'is_broken', 'tokens_x', 'tokens_y']
y_train, X_train = train_samples['relation'].to_frame(), train_samples.drop('relation', axis=1).drop(
columns=drop_columns + ['category_id'])
y_dev, X_dev = dev_samples['relation'].to_frame(), dev_samples.drop('relation', axis=1).drop(
columns=drop_columns + ['category_id'])
y_test, X_test = test_samples['relation'].to_frame(), test_samples.drop('relation', axis=1).drop(
columns=drop_columns + ['category_id'])
constants = [c for c in X_train.columns if len(set(X_train[c])) == 1]
X_train = X_train.drop(columns=constants)
X_dev = X_dev.drop(columns=constants)
X_test = X_test.drop(columns=constants)
```
### Classifiers training
```
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler, RobustScaler
std_scaler = MinMaxScaler().fit(X_train.values)
X_train = pd.DataFrame(std_scaler.transform(X_train.values), index=X_train.index, columns=X_train.columns)
X_dev = pd.DataFrame(std_scaler.transform(X_dev.values), index=X_dev.index, columns=X_dev.columns)
X_test = pd.DataFrame(std_scaler.transform(X_test.values), index=X_test.index, columns=X_test.columns)
X_train.shape
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
model = LogisticRegression(solver='lbfgs', C=0.0005, n_jobs=4, class_weight='balanced', random_state=random_state)
model.fit(X_train, y_train)
from sklearn import metrics
predicted = model.predict(X_test)
print('pr:', metrics.precision_score(y_test, predicted))
print('re:', metrics.recall_score(y_test, predicted))
print('f1:', metrics.f1_score(y_test, predicted))
print()
print(metrics.classification_report(y_test, predicted))
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
params = {
'C':[0.01, 0.1, 0.5, 1.,]
}
model = GridSearchCV(estimator=LinearSVC(random_state=random_state, class_weight='balanced'),
param_grid=params,
scoring = 'f1_macro',
n_jobs=-1, cv=10, verbose=2)
model.fit(X_train, y_train)
model.best_params_, model.best_score_
from sklearn.svm import LinearSVC
model = LinearSVC(random_state=random_state, C=0.1, class_weight='balanced')
model.fit(X_train, y_train)
from sklearn import metrics
predicted = model.predict(X_dev)
print('f1: %.2f'%(metrics.f1_score(y_dev, predicted)*100.))
print('pr: %.2f'%(metrics.precision_score(y_dev, predicted)*100.))
print('re: %.2f'%(metrics.recall_score(y_dev, predicted)*100.))
print()
print(metrics.classification_report(y_dev, predicted, digits=4))
predicted = model.predict(X_test)
print('f1: %.2f'%(metrics.f1_score(y_test, predicted)*100.))
print('pr: %.2f'%(metrics.precision_score(y_test, predicted)*100.))
print('re: %.2f'%(metrics.recall_score(y_test, predicted)*100.))
print()
print(metrics.classification_report(y_test, predicted, digits=4))
model.labels = ["0", "1"]
pickle.dump(model, open(os.path.join(model_path, 'model.pkl'), 'wb'))
pickle.dump(std_scaler, open(os.path.join(model_path, 'scaler.pkl'), 'wb'))
pickle.dump(constants+drop_columns, open(os.path.join(model_path, 'drop_columns.pkl'), 'wb'))
```
| github_jupyter |
```
!pip install google-compute-engine
%%writefile spark_analysis.py
import matplotlib
matplotlib.use('agg')
import argparse
parser = argparse.ArgumentParser()
parser.add_argument("--bucket", help="bucket for input and output")
args = parser.parse_args()
BUCKET = args.bucket
```
## Migrating from Spark to BigQuery via Dataproc -- Part 1
* [Part 1](01_spark.ipynb): The original Spark code, now running on Dataproc (lift-and-shift).
* [Part 2](02_gcs.ipynb): Replace HDFS by Google Cloud Storage. This enables job-specific-clusters. (cloud-native)
* [Part 3](03_automate.ipynb): Automate everything, so that we can run in a job-specific cluster. (cloud-optimized)
* [Part 4](04_bigquery.ipynb): Load CSV into BigQuery, use BigQuery. (modernize)
* [Part 5](05_functions.ipynb): Using Cloud Functions, launch analysis every time there is a new file in the bucket. (serverless)
### Copy data to HDFS
The Spark code in this notebook is based loosely on the [code](https://github.com/dipanjanS/data_science_for_all/blob/master/tds_spark_sql_intro/Working%20with%20SQL%20at%20Scale%20-%20Spark%20SQL%20Tutorial.ipynb) accompanying [this post](https://opensource.com/article/19/3/apache-spark-and-dataframes-tutorial) by Dipanjan Sarkar. I am using it to illustrate migrating a Spark analytics workload to BigQuery via Dataproc.
The data itself comes from the 1999 KDD competition. Let's grab 10% of the data to use as an illustration.
### Reading in data
The data are CSV files. In Spark, these can be read using textFile and splitting rows on commas.
```
%%writefile -a spark_analysis.py
from pyspark.sql import SparkSession, SQLContext, Row
gcs_bucket='qwiklabs-gcp-ffc84680e86718f5'
spark = SparkSession.builder.appName("kdd").getOrCreate()
sc = spark.sparkContext
data_file = "gs://"+gcs_bucket+"//kddcup.data_10_percent.gz"
raw_rdd = sc.textFile(data_file).cache()
raw_rdd.take(5)
%%writefile -a spark_analysis.py
csv_rdd = raw_rdd.map(lambda row: row.split(","))
parsed_rdd = csv_rdd.map(lambda r: Row(
duration=int(r[0]),
protocol_type=r[1],
service=r[2],
flag=r[3],
src_bytes=int(r[4]),
dst_bytes=int(r[5]),
wrong_fragment=int(r[7]),
urgent=int(r[8]),
hot=int(r[9]),
num_failed_logins=int(r[10]),
num_compromised=int(r[12]),
su_attempted=r[14],
num_root=int(r[15]),
num_file_creations=int(r[16]),
label=r[-1]
)
)
parsed_rdd.take(5)
```
### Spark analysis
One way to analyze data in Spark is to call methods on a dataframe.
```
%%writefile -a spark_analysis.py
sqlContext = SQLContext(sc)
df = sqlContext.createDataFrame(parsed_rdd)
connections_by_protocol = df.groupBy('protocol_type').count().orderBy('count', ascending=False)
connections_by_protocol.show()
```
Another way is to use Spark SQL
```
%%writefile -a spark_analysis.py
df.registerTempTable("connections")
attack_stats = sqlContext.sql("""
SELECT
protocol_type,
CASE label
WHEN 'normal.' THEN 'no attack'
ELSE 'attack'
END AS state,
COUNT(*) as total_freq,
ROUND(AVG(src_bytes), 2) as mean_src_bytes,
ROUND(AVG(dst_bytes), 2) as mean_dst_bytes,
ROUND(AVG(duration), 2) as mean_duration,
SUM(num_failed_logins) as total_failed_logins,
SUM(num_compromised) as total_compromised,
SUM(num_file_creations) as total_file_creations,
SUM(su_attempted) as total_root_attempts,
SUM(num_root) as total_root_acceses
FROM connections
GROUP BY protocol_type, state
ORDER BY 3 DESC
""")
attack_stats.show()
%%writefile -a spark_analysis.py
# %matplotlib inline
ax = attack_stats.toPandas().plot.bar(x='protocol_type', subplots=True, figsize=(10,25))
%%writefile -a spark_analysis.py
ax[0].get_figure().savefig('report.png');
%%writefile -a spark_analysis.py
import google.cloud.storage as gcs
bucket = gcs.Client().get_bucket(BUCKET)
for blob in bucket.list_blobs(prefix='sparktobq/'):
blob.delete()
bucket.blob('sparktobq/report.pdf').upload_from_filename('report.png')
%%writefile -a spark_analysis.py
connections_by_protocol.write.format("csv").mode("overwrite").save(
"gs://{}/sparktobq/connections_by_protocol".format(BUCKET))
BUCKET_list = !gcloud info --format='value(config.project)'
BUCKET=BUCKET_list[0]
print('Writing to {}'.format(BUCKET))
!python spark_analysis.py --bucket=$BUCKET
!gsutil ls gs://$BUCKET/sparktobq/**
!gsutil cp spark_analysis.py gs://$BUCKET/sparktobq/spark_analysis.py
```
Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
```
import pandas as pd
import numpy as np
def calculaPorDia(x):
#print(x[3]/x[6].days)
#print(x[3],'',x[7])
return x[3]/x[7].days
hotels = []
#filename = 'hotels-in-Veneza.txt'
#filename = 'hotels-in-Zurich.txt'
filename = 'hotels-in-Munich.txt'
#filename = 'hotels-in-Lucerna.txt'
with open(filename) as f:
for line in f:
if len(line)>10:
hotels.append([str(n.replace('\xa0','').replace('"','').replace('[','').replace(']','')) for n in line.strip().split(';')])
#print(line)
import datetime
hotelDict = []
for i in hotels:
#print(i[0][3:5])
dtIni = datetime.datetime(int(i[0][6:10]), int(i[0][3:5]), int(i[0][0:2]), 0, 0, 0)
dtEnd = datetime.datetime(int(i[1][6:10]), int(i[1][3:5]), int(i[1][0:2]), 0, 0, 0)
hotelDict.append({'de':dtIni,'ate':dtEnd,'nome':i[2],'preco':i[3],'nota':i[4],'distancia':i[5],'url':i[6]})
df = pd.DataFrame(hotelDict)
df.drop_duplicates(inplace = True)
df = df[df['preco']>'-1']
df['preco'].replace(regex=True,inplace=True,to_replace=r'\D',value=r'')
df['preco'] = pd.to_numeric(df['preco'])
df['dias'] = (df['ate']-df['de']).replace(regex=True,to_replace=r'\D',value=r'')
df["precoPorNoite"] = df.apply(calculaPorDia,axis=1)
display(df.describe())
datetimeFormat = '%Y-%m-%d %H:%M:%S.%f'
#df['diasteste'] = datetime.datetime.strptime(df['ate'],datetimeFormat) - datetime.datetime.strptime(df['de'],datetimeFormat)
df['dias'] = (df['ate']-df['de']).replace(regex=True,to_replace=r'\D',value=r'')
df["nota"] = pd.to_numeric(df["nota"])
df['distanciaMedida'] = pd.np.where(df.distancia.str.contains("km"),"km","m")
df["distanciaNumerica"] = pd.to_numeric(df['distancia'].replace(regex=True,to_replace=r'\D',value=r''))#, downcast='float')
df["distanciaMetros"] = pd.np.where(df.distancia.str.contains("km") & df.distancia.str.contains("."),df.distanciaNumerica*1000,df["distanciaNumerica"])
df["distanciaMetros"] = pd.np.where(df.distanciaMetros > 9000,(df.distanciaNumerica/10),df["distanciaNumerica"])
df["distanciaMetros"] = df["distanciaMetros"].apply(lambda x: x*1000 if x < 100 else x)
#df["precoPorNoite"] = df.apply(calculaPorDia,axis=1)
df = df[df['precoPorNoite']<1000]
df = df[df['nota']>7.5]
df = df[df['distanciaMetros']<2000]
df = df[~df["nome"].str.contains('Hostel')]
del df['distanciaMedida']
del df['distanciaNumerica']
df.drop_duplicates(inplace = True)
display(df.describe())
display(df)
import plotly.express as px
#fig = px.scatter(df, x="precoPorNoite", y="nota",color="nome")
#fig = px.scatter(df, x="preco", y="nota",color="nome")
#fig.show()
import plotly.express as px
fig = px.scatter(df, x="precoPorNoite", y="nota", size="distanciaMetros", color="nome",hover_name="nome",log_x=True)
fig.show()
fig.write_html(filename+".html")
df['link'] ='<a href="http://'+df['url']+'">Clique</a>'
df= df.drop_duplicates(subset=['nome', 'preco'])
dfa = df.copy()
del dfa['url']
del dfa['de']
del dfa['ate']
del dfa['dias']
del dfa['distanciaMetros']
dfa = dfa.to_html()
print('de: ',df['de'].max())
print('até: ',df['ate'].max())
#display(dfa)
#display(df.drop_duplicates().sort_values(by=['precoPorNoite'],ascending=False))
from IPython.core.display import display, HTML
display(HTML(dfa))
```
| github_jupyter |
```
import neutromeratio
from neutromeratio.constants import kT
from simtk import unit
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sb
import pickle
from collections import defaultdict
from neutromeratio.analysis import bootstrap_rmse_r
# defining the plotting function
def plot_correlation_analysis(
names: list,
x_: list,
y_: list,
title: str,
x_label: str,
y_label: str,
fontsize: int = 15,
nsamples: int = 5000,
yerror: list = [],
mark_point_by_name=[],
):
"""Plot correlation between x and y.
Parameters
----------
df : pd.Dataframe
the df contains columns with colum names 'names', 'x', 'y', 'y-error'
title : str
to put above plot. use '' (empty string) for no title.
nsamples : int
number of samples to draw for bootstrap
"""
plt.figure(figsize=[8, 8], dpi=300)
ax = plt.gca()
ax.set_title(title, fontsize=fontsize)
rmse, mae, r = bootstrap_rmse_r(np.array(x_), np.array(y_), 1000)
plt.text(-9.0, 20.0, r"""MAE$ = {}$ kcal/mol
RMSE$ = {}$ kcal/mol
tautomer pairs$ = {}$""".format(mae,rmse, len(names)), fontsize=fontsize-4, bbox={'facecolor': 'grey', 'alpha': 0.7, 'pad': 5})
if yerror:
for X, Y, name, error in zip(x_, y_, names, yerror):
ax.errorbar(
X,
Y,
yerr=error,
mfc="blue",
mec="blue",
ms=4,
fmt="o",
capthick=2,
capsize=2,
alpha=0.6,
ecolor="red",
)
else:
for X, Y, name in zip(x_, y_, names):
if name in mark_point_by_name:
ax.scatter(X, Y, color="red", s=13, alpha=0.6)
else:
ax.scatter(X, Y, color="blue", s=13, alpha=0.6)
# draw lines +- 1kcal/mol
ax.plot((-10.0, 30.0), (-10.0, 30.0), "k--", zorder=-1, linewidth=1.0, alpha=0.5)
ax.plot((-9.0, 30.0), (-10.0, 29.0), "gray", zorder=-1, linewidth=1.0, alpha=0.5)
ax.plot((-10.0, 29.0), (-9.0, 30.0), "gray", zorder=-1, linewidth=1.0, alpha=0.5)
ax.plot((-10.0, 30.0), (0.0, 0.0), "r--", zorder=-1, linewidth=1.0, alpha=0.5)
ax.plot((0.0, 0.0), (-10.0, 30.0), "r--", zorder=-1, linewidth=1.0, alpha=0.5)
ax.set_xlabel(x_label, fontsize=fontsize)
ax.set_ylabel(y_label, fontsize=fontsize)
plt.tight_layout()
plt.fill(
0.0,
)
plt.subplots_adjust(bottom=0.3), # left=1.3, right=0.3)
# make sure that we plot a square
ax.set_aspect("equal", "box")
# color quadrants
x = np.arange(0.01, 30, 0.1)
y = -30 # np.arange(0.01,30,0.1)
plt.fill_between(x, y, color="#539ecd", alpha=0.2)
x = -np.arange(0.01, 30, 0.1)
y = 30 # np.arange(0.01,30,0.1)
plt.fill_between(x, y, color="#539ecd", alpha=0.2)
plt.axvspan(-10, 0, color="grey")
plt.setp(ax.get_xticklabels(), fontsize=20)
plt.setp(ax.get_yticklabels(), fontsize=20)
ax.set_xlim([-10, 30])
ax.set_ylim([-10, 30])
return plt.gca()
# load the experimental and RAFE results
exp_results = pickle.load(open(f'../data/input/exp_results.pickle', 'rb'))
# which potential
potential = 'ANI2x' # or 'ANI1ccx'
csv_file = f'../calculated/Alchemical{potential}_kT.csv'
results = defaultdict(list)
error = defaultdict(list)
exp = defaultdict(list)
with open(csv_file, 'r') as f:
for line in f.readlines():
name, ddG, dddG = (line.split(','))
results[name] = (float(ddG) *kT).value_in_unit(unit.kilocalorie_per_mole)
error[name] = (float(dddG) *kT).value_in_unit(unit.kilocalorie_per_mole)
exp[name]= exp_results[name]['energy [kcal/mol]'].value_in_unit(unit.kilocalorie_per_mole)
# comparing exp and AFE resutls
import seaborn as sns
sns.set_context('paper')
sns.set(color_codes=True)
x_list = []
y_list = []
yerror_list = []
names = []
for name in exp:
x, y, y_error = exp[name], results[name], error[name]
if x < 0.0:
x *= -1.
y *= -1.
x_list.append(x)
y_list.append(y)
yerror_list.append(y_error)
names.append(name)
assert(len(x_list) == len(y_list))
f = plot_correlation_analysis(names, x_list, y_list, yerror=yerror_list, title='Alchemical free energy\n calculations with ANI-1ccx', x_label='$\Delta_{t} G_{solv}^{exp}$ [kcal/mol]', y_label='$\Delta_{t} G_{solv}^{calc}$ [kcal/mol]', fontsize=25)
f.plot()
```
| github_jupyter |
```
!curl -L https://raw.githubusercontent.com/facebookresearch/habitat-sim/master/examples/colab_utils/colab_install.sh | NIGHTLY=true bash -s
%cd /content/habitat-sim
## [setup]
import os
import random
import sys
import git
import magnum as mn
import numpy as np
import habitat_sim
from habitat_sim.utils import viz_utils as vut
if "google.colab" in sys.modules:
os.environ["IMAGEIO_FFMPEG_EXE"] = "/usr/bin/ffmpeg"
repo = git.Repo(".", search_parent_directories=True)
dir_path = repo.working_tree_dir
%cd $dir_path
data_path = os.path.join(dir_path, "data")
output_path = os.path.join(
dir_path, "examples/tutorials/managed_rigid_object_tutorial_output/"
)
def place_agent(sim):
# place our agent in the scene
agent_state = habitat_sim.AgentState()
agent_state.position = [-0.15, -0.7, 1.0]
agent_state.rotation = np.quaternion(-0.83147, 0, 0.55557, 0)
agent = sim.initialize_agent(0, agent_state)
return agent.scene_node.transformation_matrix()
def make_configuration():
# simulator configuration
backend_cfg = habitat_sim.SimulatorConfiguration()
backend_cfg.scene_id = os.path.join(
data_path, "scene_datasets/habitat-test-scenes/apartment_1.glb"
)
assert os.path.exists(backend_cfg.scene_id)
backend_cfg.enable_physics = True
# sensor configurations
# Note: all sensors must have the same resolution
# setup 2 rgb sensors for 1st and 3rd person views
camera_resolution = [544, 720]
sensor_specs = []
rgba_camera_1stperson_spec = habitat_sim.CameraSensorSpec()
rgba_camera_1stperson_spec.uuid = "rgba_camera_1stperson"
rgba_camera_1stperson_spec.sensor_type = habitat_sim.SensorType.COLOR
rgba_camera_1stperson_spec.resolution = camera_resolution
rgba_camera_1stperson_spec.position = [0.0, 0.6, 0.0]
rgba_camera_1stperson_spec.orientation = [0.0, 0.0, 0.0]
rgba_camera_1stperson_spec.sensor_subtype = habitat_sim.SensorSubType.PINHOLE
sensor_specs.append(rgba_camera_1stperson_spec)
depth_camera_1stperson_spec = habitat_sim.CameraSensorSpec()
depth_camera_1stperson_spec.uuid = "depth_camera_1stperson"
depth_camera_1stperson_spec.sensor_type = habitat_sim.SensorType.DEPTH
depth_camera_1stperson_spec.resolution = camera_resolution
depth_camera_1stperson_spec.position = [0.0, 0.6, 0.0]
depth_camera_1stperson_spec.orientation = [0.0, 0.0, 0.0]
depth_camera_1stperson_spec.sensor_subtype = habitat_sim.SensorSubType.PINHOLE
sensor_specs.append(depth_camera_1stperson_spec)
rgba_camera_3rdperson_spec = habitat_sim.CameraSensorSpec()
rgba_camera_3rdperson_spec.uuid = "rgba_camera_3rdperson"
rgba_camera_3rdperson_spec.sensor_type = habitat_sim.SensorType.COLOR
rgba_camera_3rdperson_spec.resolution = camera_resolution
rgba_camera_3rdperson_spec.position = [0.0, 1.0, 0.3]
rgba_camera_3rdperson_spec.orientation = [-45, 0.0, 0.0]
rgba_camera_3rdperson_spec.sensor_subtype = habitat_sim.SensorSubType.PINHOLE
sensor_specs.append(rgba_camera_3rdperson_spec)
# agent configuration
agent_cfg = habitat_sim.agent.AgentConfiguration()
agent_cfg.sensor_specifications = sensor_specs
return habitat_sim.Configuration(backend_cfg, [agent_cfg])
def simulate(sim, dt=1.0, get_frames=True):
# simulate dt seconds at 60Hz to the nearest fixed timestep
print("Simulating " + str(dt) + " world seconds.")
observations = []
start_time = sim.get_world_time()
while sim.get_world_time() < start_time + dt:
sim.step_physics(1.0 / 60.0)
if get_frames:
observations.append(sim.get_sensor_observations())
return observations
# [/setup]
if __name__ == "__main__":
import argparse
parser = argparse.ArgumentParser()
parser.add_argument("--no-show-video", dest="show_video", action="store_false")
parser.add_argument("--no-make-video", dest="make_video", action="store_false")
parser.set_defaults(show_video=True, make_video=True)
args, _ = parser.parse_known_args()
show_video = args.show_video
make_video = args.make_video
if make_video and not os.path.exists(output_path):
os.mkdir(output_path)
# [initialize]
# create the simulators AND resets the simulator
cfg = make_configuration()
try: # Got to make initialization idiot proof
sim.close()
except NameError:
pass
sim = habitat_sim.Simulator(cfg)
agent_transform = place_agent(sim)
# get the primitive assets attributes manager
prim_templates_mgr = sim.get_asset_template_manager()
# get the physics object attributes manager
obj_templates_mgr = sim.get_object_template_manager()
# get the rigid object manager
rigid_obj_mgr = sim.get_rigid_object_manager()
# [/initialize]
# [basics]
# load some object templates from configuration files
sphere_template_id = obj_templates_mgr.load_configs(
str(os.path.join(data_path, "test_assets/objects/sphere"))
)[0]
# add a sphere to the scene, returns the object
sphere_obj = rigid_obj_mgr.add_object_by_template_id(sphere_template_id)
# move sphere
sphere_obj.translation = [2.50, 0.0, 0.2]
# simulate
observations = simulate(sim, dt=1.5, get_frames=make_video)
if make_video:
vut.make_video(
observations,
"rgba_camera_1stperson",
"color",
output_path + "sim_basics",
open_vid=show_video,
)
# [/basics]
rigid_obj_mgr.remove_all_objects()
# [object_user_configurations]
# modify an object's user-defined configurations
# load some object templates from configuration files
sphere_template_id = obj_templates_mgr.load_configs(
str(os.path.join(data_path, "test_assets/objects/sphere"))
)[0]
# add a sphere to the scene, returns the object
sphere_obj = rigid_obj_mgr.add_object_by_template_id(sphere_template_id)
# set user-defined configuration values
user_attributes_dict = {
"obj_val_0": "This is a sphere object named " + sphere_obj.handle,
"obj_val_1": 17,
"obj_val_2": False,
"obj_val_3": 19.7,
"obj_val_4": [2.50, 0.0, 0.2],
"obj_val_5": mn.Quaternion.rotation(mn.Deg(90.0), [-1.0, 0.0, 0.0]),
}
for k, v in user_attributes_dict.items():
sphere_obj.user_attributes.set(k, v)
for k, _ in user_attributes_dict.items():
print(
"Sphere Object user attribute : {} : {}".format(
k, sphere_obj.user_attributes.get_as_string(k)
)
)
# [/object_user_configurations]
rigid_obj_mgr.remove_all_objects()
# [dynamic_control]
observations = []
obj_templates_mgr.load_configs(
str(os.path.join(data_path, "objects/example_objects/"))
)
# search for an object template by key sub-string
cheezit_template_handle = obj_templates_mgr.get_template_handles(
"data/objects/example_objects/cheezit"
)[0]
# build multiple object initial positions
box_positions = [
[2.39, -0.37, 0.0],
[2.39, -0.64, 0.0],
[2.39, -0.91, 0.0],
[2.39, -0.64, -0.22],
[2.39, -0.64, 0.22],
]
box_orientation = mn.Quaternion.rotation(mn.Deg(90.0), [-1.0, 0.0, 0.0])
# instance and place the boxes
boxes = []
for b in range(len(box_positions)):
boxes.append(
rigid_obj_mgr.add_object_by_template_handle(cheezit_template_handle)
)
boxes[b].translation = box_positions[b]
boxes[b].rotation = box_orientation
# anti-gravity force f=m(-g) using first object's mass (all objects have the same mass)
anti_grav_force = -1.0 * sim.get_gravity() * boxes[0].mass
# throw a sphere at the boxes from the agent position
sphere_template = obj_templates_mgr.get_template_by_id(sphere_template_id)
sphere_template.scale = [0.5, 0.5, 0.5]
obj_templates_mgr.register_template(sphere_template)
# create sphere
sphere_obj = rigid_obj_mgr.add_object_by_template_id(sphere_template_id)
sphere_obj.translation = sim.agents[0].get_state().position + [0.0, 1.0, 0.0]
# get the vector from the sphere to a box
target_direction = boxes[0].translation - sphere_obj.translation
# apply an initial velocity for one step
sphere_obj.linear_velocity = target_direction * 5
sphere_obj.angular_velocity = [0.0, -1.0, 0.0]
start_time = sim.get_world_time()
dt = 3.0
while sim.get_world_time() < start_time + dt:
# set forces/torques before stepping the world
for box in boxes:
box.apply_force(anti_grav_force, [0.0, 0.0, 0.0])
box.apply_torque([0.0, 0.01, 0.0])
sim.step_physics(1.0 / 60.0)
observations.append(sim.get_sensor_observations())
if make_video:
vut.make_video(
observations,
"rgba_camera_1stperson",
"color",
output_path + "dynamic_control",
open_vid=show_video,
)
# [/dynamic_control]
rigid_obj_mgr.remove_all_objects()
# [kinematic_interactions]
chefcan_template_handle = obj_templates_mgr.get_template_handles(
"data/objects/example_objects/chefcan"
)[0]
chefcan_obj = rigid_obj_mgr.add_object_by_template_handle(chefcan_template_handle)
chefcan_obj.translation = [2.4, -0.64, 0.0]
# set object to kinematic
chefcan_obj.motion_type = habitat_sim.physics.MotionType.KINEMATIC
# drop some dynamic objects
chefcan_obj_2 = rigid_obj_mgr.add_object_by_template_handle(chefcan_template_handle)
chefcan_obj_2.translation = [2.4, -0.64, 0.28]
chefcan_obj_3 = rigid_obj_mgr.add_object_by_template_handle(chefcan_template_handle)
chefcan_obj_3.translation = [2.4, -0.64, -0.28]
chefcan_obj_4 = rigid_obj_mgr.add_object_by_template_handle(chefcan_template_handle)
chefcan_obj_4.translation = [2.4, -0.3, 0.0]
# simulate
observations = simulate(sim, dt=1.5, get_frames=True)
if make_video:
vut.make_video(
observations,
"rgba_camera_1stperson",
"color",
output_path + "kinematic_interactions",
open_vid=show_video,
)
# [/kinematic_interactions]
rigid_obj_mgr.remove_all_objects()
# [kinematic_update]
observations = []
clamp_template_handle = obj_templates_mgr.get_template_handles(
"data/objects/example_objects/largeclamp"
)[0]
clamp_obj = rigid_obj_mgr.add_object_by_template_handle(clamp_template_handle)
clamp_obj.motion_type = habitat_sim.physics.MotionType.KINEMATIC
clamp_obj.translation = [0.8, 0.2, 0.5]
start_time = sim.get_world_time()
dt = 1.0
while sim.get_world_time() < start_time + dt:
# manually control the object's kinematic state
clamp_obj.translation += [0.0, 0.0, 0.01]
clamp_obj.rotation = (
mn.Quaternion.rotation(mn.Rad(0.05), [-1.0, 0.0, 0.0]) * clamp_obj.rotation
)
sim.step_physics(1.0 / 60.0)
observations.append(sim.get_sensor_observations())
if make_video:
vut.make_video(
observations,
"rgba_camera_1stperson",
"color",
output_path + "kinematic_update",
open_vid=show_video,
)
# [/kinematic_update]
# [velocity_control]
# get object VelocityControl structure and setup control
vel_control = clamp_obj.velocity_control
vel_control.linear_velocity = [0.0, 0.0, -1.0]
vel_control.angular_velocity = [4.0, 0.0, 0.0]
vel_control.controlling_lin_vel = True
vel_control.controlling_ang_vel = True
observations = simulate(sim, dt=1.0, get_frames=True)
# reverse linear direction
vel_control.linear_velocity = [0.0, 0.0, 1.0]
observations += simulate(sim, dt=1.0, get_frames=True)
if make_video:
vut.make_video(
observations,
"rgba_camera_1stperson",
"color",
output_path + "velocity_control",
open_vid=show_video,
)
# [/velocity_control]
# [local_velocity_control]
vel_control.linear_velocity = [0.0, 0.0, 2.3]
vel_control.angular_velocity = [-4.3, 0.0, 0.0]
vel_control.lin_vel_is_local = True
vel_control.ang_vel_is_local = True
observations = simulate(sim, dt=1.5, get_frames=True)
# video rendering
if make_video:
vut.make_video(
observations,
"rgba_camera_1stperson",
"color",
output_path + "local_velocity_control",
open_vid=show_video,
)
# [/local_velocity_control]
rigid_obj_mgr.remove_all_objects()
# [embodied_agent]
# load the lobot_merged asset
locobot_template_id = obj_templates_mgr.load_configs(
str(os.path.join(data_path, "objects/locobot_merged"))
)[0]
# add robot object to the scene with the agent/camera SceneNode attached
locobot = rigid_obj_mgr.add_object_by_template_id(
locobot_template_id, sim.agents[0].scene_node
)
locobot.translation = [1.75, -1.02, 0.4]
vel_control = locobot.velocity_control
vel_control.linear_velocity = [0.0, 0.0, -1.0]
vel_control.angular_velocity = [0.0, 2.0, 0.0]
# simulate robot dropping into place
observations = simulate(sim, dt=1.5, get_frames=make_video)
vel_control.controlling_lin_vel = True
vel_control.controlling_ang_vel = True
vel_control.lin_vel_is_local = True
vel_control.ang_vel_is_local = True
# simulate forward and turn
observations += simulate(sim, dt=1.0, get_frames=make_video)
vel_control.controlling_lin_vel = False
vel_control.angular_velocity = [0.0, 1.0, 0.0]
# simulate turn only
observations += simulate(sim, dt=1.5, get_frames=make_video)
vel_control.angular_velocity = [0.0, 0.0, 0.0]
vel_control.controlling_lin_vel = True
vel_control.controlling_ang_vel = True
# simulate forward only with damped angular velocity (reset angular velocity to 0 after each step)
observations += simulate(sim, dt=1.0, get_frames=make_video)
vel_control.angular_velocity = [0.0, -1.25, 0.0]
# simulate forward and turn
observations += simulate(sim, dt=2.0, get_frames=make_video)
vel_control.controlling_ang_vel = False
vel_control.controlling_lin_vel = False
# simulate settling
observations += simulate(sim, dt=3.0, get_frames=make_video)
# remove the agent's body while preserving the SceneNode
rigid_obj_mgr.remove_object_by_id(locobot.object_id, delete_object_node=False)
# demonstrate that the locobot object does not now exist'
print("Locobot is still alive : {}".format(locobot.is_alive))
# video rendering with embedded 1st person view
if make_video:
vut.make_video(
observations,
"rgba_camera_1stperson",
"color",
output_path + "robot_control",
open_vid=show_video,
)
# [/embodied_agent]
# [embodied_agent_navmesh]
# load the lobot_merged asset
locobot_template_id = obj_templates_mgr.load_configs(
str(os.path.join(data_path, "objects/locobot_merged"))
)[0]
# add robot object to the scene with the agent/camera SceneNode attached
locobot = rigid_obj_mgr.add_object_by_template_id(
locobot_template_id, sim.agents[0].scene_node
)
initial_rotation = locobot.rotation
# set the agent's body to kinematic since we will be updating position manually
locobot.motion_type = habitat_sim.physics.MotionType.KINEMATIC
# create and configure a new VelocityControl structure
# Note: this is NOT the object's VelocityControl, so it will not be consumed automatically in sim.step_physics
vel_control = habitat_sim.physics.VelocityControl()
vel_control.controlling_lin_vel = True
vel_control.lin_vel_is_local = True
vel_control.controlling_ang_vel = True
vel_control.ang_vel_is_local = True
vel_control.linear_velocity = [0.0, 0.0, -1.0]
# try 2 variations of the control experiment
for iteration in range(2):
# reset observations and robot state
observations = []
locobot.translation = [1.75, -1.02, 0.4]
locobot.rotation = initial_rotation
vel_control.angular_velocity = [0.0, 0.0, 0.0]
video_prefix = "robot_control_sliding"
# turn sliding off for the 2nd pass
if iteration == 1:
sim.config.sim_cfg.allow_sliding = False
video_prefix = "robot_control_no_sliding"
# manually control the object's kinematic state via velocity integration
start_time = sim.get_world_time()
last_velocity_set = 0
dt = 6.0
time_step = 1.0 / 60.0
while sim.get_world_time() < start_time + dt:
previous_rigid_state = locobot.rigid_state
# manually integrate the rigid state
target_rigid_state = vel_control.integrate_transform(
time_step, previous_rigid_state
)
# snap rigid state to navmesh and set state to object/agent
end_pos = sim.step_filter(
previous_rigid_state.translation, target_rigid_state.translation
)
locobot.translation = end_pos
locobot.rotation = target_rigid_state.rotation
# Check if a collision occured
dist_moved_before_filter = (
target_rigid_state.translation - previous_rigid_state.translation
).dot()
dist_moved_after_filter = (end_pos - previous_rigid_state.translation).dot()
# NB: There are some cases where ||filter_end - end_pos|| > 0 when a
# collision _didn't_ happen. One such case is going up stairs. Instead,
# we check to see if the the amount moved after the application of the filter
# is _less_ than the amount moved before the application of the filter
EPS = 1e-5
collided = (dist_moved_after_filter + EPS) < dist_moved_before_filter
# run any dynamics simulation
sim.step_physics(time_step)
# render observation
observations.append(sim.get_sensor_observations())
# randomize angular velocity
last_velocity_set += time_step
if last_velocity_set >= 1.0:
vel_control.angular_velocity = [0.0, (random.random() - 0.5) * 2.0, 0.0]
last_velocity_set = 0
# video rendering with embedded 1st person views
if make_video:
sensor_dims = (
sim.get_agent(0).agent_config.sensor_specifications[0].resolution
)
overlay_dims = (int(sensor_dims[1] / 4), int(sensor_dims[0] / 4))
overlay_settings = [
{
"obs": "rgba_camera_1stperson",
"type": "color",
"dims": overlay_dims,
"pos": (10, 10),
"border": 2,
},
{
"obs": "depth_camera_1stperson",
"type": "depth",
"dims": overlay_dims,
"pos": (10, 30 + overlay_dims[1]),
"border": 2,
},
]
vut.make_video(
observations=observations,
primary_obs="rgba_camera_3rdperson",
primary_obs_type="color",
video_file=output_path + video_prefix,
fps=60,
open_vid=show_video,
overlay_settings=overlay_settings,
depth_clip=10.0,
)
# [/embodied_agent_navmesh]
```
| github_jupyter |
### OpenAI Gym
1. Gym is a toolkit for developing and comparing reinforcement learning algorithms.
2. It makes no assumptions about the structure of your agent, and is compatible with any numerical computation library, such as TensorFlow or Theano.
```
import gym
import numpy as np
```
#### CartPole Env
1. CartPole is the most basic environment of all types.
2. The problem consists of balancing a pole connected with one joint on top of a moving cart. The only actions are to add a force of -1 or +1 to the cart, pushing it left or right.
3. In CartPole's environment, there are four observations at any given state, representing information such as the angle of the pole and the position of the cart.
4. Using these observations, the agent needs to decide on one of two possible actions: move the cart left or right.
```
env = gym.make('CartPole-v0')
def run_episode(env, parameters, play=False):
observation = env.reset()
totalreward = 0
for _ in range(200):
action = 0 if np.matmul(parameters,observation) < 0 else 1
observation, reward, done, info = env.step(action)
totalreward += reward
if done:
if play: env.close()
break
else:
if play: env.render()
return totalreward
```
#### Strategy 1 : Random Search
1. One fairly straightforward strategy is to keep trying random weights, and pick the one that performs the best.
2. Since the CartPole environment is relatively simple, with only 4 observations, this basic method works surprisingly well.
```
#look what taking an action returns
print(env.step(0))
env.reset()
best_parameters = np.random.rand(4) * 2 - 1
best_reward = run_episode(env,best_parameters,False)
iters = 0
for _ in range(1000):
parameters = np.random.rand(4) * 2 - 1
reward = run_episode(env,parameters,False)
if reward > best_reward:
best_parameters = parameters
best_reward = reward
iters+=1
if best_reward==200:
break
print("Highest Reward : {} in {} iterations".format(best_reward,iters))
```
#### Strategy 2 : Hill Climbing
1. We start with some randomly chosen initial weights. Every episode, add some noise to the weights, and keep the new weights if the agent improves.
2. Idea here is to gradually improve the weights, rather than keep jumping around and hopefully finding some combination that works. If noise_scaling is high enough in comparison to the current weights, this algorithm is essentially the same as random search.
3. If the weights are initialized badly, adding noise may have no effect on how well the agent performs, causing it to get stuck.
```
noise_scaling = 0.1
best_parameters = np.random.rand(4)*2 - 1
best_reward = 0
last_update = 0
iters = 0
while last_update < 100:
parameters = best_parameters + (np.random.rand(4)*2 - 1)*noise_scaling
reward = run_episode(env,parameters,False)
if reward > best_reward:
best_reward = reward
best_parameters = parameters
last_update = 0
else:
last_update+=1
#incrementing iterations
iters += 1
if best_reward==200:
break
print("Highest Reward : {} in {} iterations".format(best_reward,iters))
```
| github_jupyter |
# Performance of LeNet-5
```
% matplotlib inline
import os
import numpy as np
import matplotlib.pyplot as plt
from scripts.utils import ExpResults
plotproperties = {'font.size': 13,
'axes.titlesize': 'xx-large',
'axes.labelsize': 'xx-large',
'xtick.labelsize': 'xx-large',
'xtick.major.size': 7,
'xtick.minor.size': 5,
'ytick.labelsize': 'xx-large',
'ytick.major.size': 7,
'ytick.minor.size': 5,
'legend.fontsize': 'x-large',
'figure.figsize': (7, 6),
'savefig.dpi': 300,
'savefig.format': 'pdf'}
import matplotlib as mpl
mpl.rcParams.update(plotproperties)
```
## Parameters and definitions
```
base_dir = '/home/rbodo/.snntoolbox/data/mnist/cnn/lenet5/keras'
log_dir = os.path.join(base_dir, '32bit', 'log', 'gui')
runlabel_rate = '08'
runlabel_ttfs = '19' # With softmax: '18'
runlabel_dyn_thresh = '20' # With softmax: '24'
runlabel_clamped = '04' # With softmax: '03'
runlabel_dyn_thresh2 = '21' # With softmax: '25'. Includes operations due to threshold updates
path_rate = os.path.join(log_dir, runlabel_rate)
path_ttfs = os.path.join(log_dir, runlabel_ttfs)
path_dyn_thresh = os.path.join(log_dir, runlabel_dyn_thresh)
path_dyn_thresh2 = os.path.join(log_dir, runlabel_dyn_thresh2)
path_clamped = os.path.join(base_dir, 'clamped_relu', 'log', 'gui', runlabel_clamped)
exp_rate = ExpResults(path_rate, 'rate-code', 'D', 'gold', markersize=7)
exp_ttfs = ExpResults(path_ttfs, 'TTFS base', 'o', 'blue', markersize=7)
exp_dyn_thresh = ExpResults(path_dyn_thresh, 'TTFS dyn thresh', '^', 'red', markersize=7)
# exp_dyn_thresh2 = ExpResults(path_dyn_thresh2, 'TTFS dyn thresh\nincl thresh updates', 'v', 'red')
exp_clamped = ExpResults(path_clamped, 'TTFS clamped', 's', 'green', markersize=7)
experiments = [exp_rate, exp_ttfs, exp_dyn_thresh, exp_clamped]
```
## Plot error vs operations
```
from mpl_toolkits.axes_grid1.inset_locator import zoomed_inset_axes, InsetPosition, mark_inset
show_numbers = False
fig, ax = plt.subplots()
axins = zoomed_inset_axes(ax, 5)
for exp in experiments:
ax.errorbar(exp.mean_computations_t, exp.e1_mean, xerr=exp.std_computations_t, yerr=exp.e1_confidence95,
fmt=exp.marker, markersize=exp.markersize, label='SNN '+exp.label, capsize=0, elinewidth=0.1, color=exp.color1)
axins.errorbar(exp.mean_computations_t, exp.e1_mean, markersize=exp.markersize,
fmt=exp.marker, label='SNN '+exp.label, capsize=0, elinewidth=0.1, color=exp.color1)
axins.axis([0.2, 0.4, 0, 5])
ip = InsetPosition(axins, [0.25, 0.075, .65, .65])
axins.set_axes_locator(ip)
axins.set_xticks([])
axins.set_yticks([])
mark_inset(ax, axins, loc1=2, loc2=4, fc="none", ec="0.5")
exp = experiments[0]
ax.scatter(exp.operations_ann, exp.e1_ann, marker='*', s=200, label='ANN', color='black', alpha=1, linewidths=3, zorder=10)
ax.annotate('ANN', xy=(exp.operations_ann, exp.e1_ann + 3), xytext=(exp.operations_ann - 0.11, exp.e1_ann + 16),
fontsize=16, arrowprops=dict(color='black', shrink=0.05, width=5, headwidth=10))
if show_numbers:
plt.annotate('({:.2f} MOps, {:.2f}%)'.format(exp.operations_ann, exp.e1_ann), xy=(exp.operations_ann - 0.8, exp.e1_ann + 7), fontsize=16, color=exp.color1)
plt.annotate('({:.2f} MOps, {:.2f}%)'.format(exp.op1_0, exp.e1_0), xy=(exp.op1_0, exp.e1_0),
xytext=(exp.op1_0, exp.e1_0 + 13), fontsize=16,
arrowprops=dict(color=exp.color1, shrink=0.05, width=5, headwidth=10), color=exp.color1)
ax.set_xlim(0, 2.5)
ax.set_ylim(0, 100)
ax.set_xlabel('MOps')
ax.set_ylabel('Error [%]')
leg = ax.legend(loc='upper right')
leg.get_frame().set_alpha(1)
# plt.title('ANN vs SNN performance')
fig.savefig(os.path.join(log_dir, 'err_vs_ops_mnist'), bbox_inches='tight')
for exp in experiments:
print(exp.label)
print('ANN top-1: ({:.5f} MOps/frame, {:.2f} %)'.format(exp.operations_ann, exp.e1_ann))
print('SNN top-1 best error: ({:.5f} MOps/frame, {:.2f} %)'.format(exp.op1_0, exp.e1_0))
print('SNN top-1 converged: ({:.5f} MOps/frame, {:.2f} %)'.format(exp.op1_1, exp.e1_1))
print('')
exp = experiments[0]
print(exp.e1_mean[-150:])
t = np.min(np.nonzero(exp.e1_mean <= 1.61))
ops = exp.mean_computations_t[t]
print(ops)
exp = experiments[3]
print(exp.e1_mean[-150:])
t = np.min(np.nonzero(exp.e1_mean <= 1.5))
ops = exp.mean_computations_t[t]
print(ops)
experiments[0].mean_computations_t
19/50
```
| github_jupyter |
# Mountain Car with Amazon SageMaker RL
---
Mountain Car is a classic control Reinforcement Learning problem that was first introduced by A. Moore in 1991 [1]. An under-powered car is tasked with climbing a steep mountain, and is only successful when it reaches the top. Luckily there's another mountain on the opposite side which can be used to gain momentum, and launch the car to the peak. It can be tricky to find this optimal solution due to the sparsity of the reward. Complex exploration strategies can be used to incentivise exploration of the mountain, but to keep things simple in this example we extend the amount of time in each episode from Open AI Gym's default of 200 environment steps to 10,000 steps, showing how to customise environments. We consider two variants in this example: `PatientMountainCar` for discrete actions and `PatientContinuousMountainCar` for continuous actions.
<img src="./successful_policy.gif">
### `PatientMountainCar`
> 1. Objective: Get the car to the top of the right hand side mountain.
> 2. Environment(s): Open AI Gym's `MountainCar-v0` that is extended to 10,000 steps per episode.
> 3. State: Car's horizontal position and velocity (can be negative).
> 4. Action: Direction of push (left, nothing or right).
> 5. Reward: -1 for every environment step until success, which incentivises quick solutions.
### `PatientContinuousMountainCar`
> 1. Objective: Get the car to the top of the right hand side mountain.
> 2. Environment(s): Open AI Gym's `MountainCarContinuous-v0` that is extended to 10,000 steps per episode.
> 3. State: Car's horizontal position and velocity (can be negative).
> 4. Action: Mmagnitude of push (if negative push to left, if positive push to right).
> 5. Reward: +100 for reaching top of the right hand side mountain, minus the squared sum of actions from start to end.
[1] A. Moore, Efficient Memory-Based Learning for Robot Control, PhD thesis, University of Cambridge, November 1990.
## Pre-requisites
### Imports
To get started, we'll import the Python libraries we need, set up the environment with a few prerequisites for permissions and configurations.
```
import sagemaker
import boto3
import sys
import os
import glob
import re
import subprocess
import numpy as np
from IPython.display import HTML
import time
from time import gmtime, strftime
sys.path.append("common")
from misc import get_execution_role, wait_for_s3_object
from sagemaker.rl import RLEstimator, RLToolkit, RLFramework
```
### Setup S3 bucket
Set up the linkage and authentication to the S3 bucket that you want to use for checkpoint and the metadata.
```
sage_session = sagemaker.session.Session()
s3_bucket = sage_session.default_bucket()
s3_output_path = 's3://{}/'.format(s3_bucket)
print("S3 bucket path: {}".format(s3_output_path))
```
### Define Variables
We define variables such as the job prefix for the training jobs *and the image path for the container (only when this is BYOC).*
```
# create unique job name
job_name_prefix = 'rl-mountain-car'
```
### Configure settings
You can run your RL training jobs on a SageMaker notebook instance or on your own machine. In both of these scenarios, you can run in either 'local' (where you run the commands) or 'SageMaker' mode (on SageMaker training instances). 'local' mode uses the SageMaker Python SDK to run your code in Docker containers locally. It can speed up iterative testing and debugging while using the same familiar Python SDK interface. Just set `local_mode = True`. And when you're ready move to 'SageMaker' mode to scale things up.
```
# run in local mode?
local_mode = False
```
### Create an IAM role
Either get the execution role when running from a SageMaker notebook instance `role = sagemaker.get_execution_role()` or, when running from local notebook instance, use utils method `role = get_execution_role()` to create an execution role.
```
try:
role = sagemaker.get_execution_role()
except:
role = get_execution_role()
print("Using IAM role arn: {}".format(role))
```
### Install docker for `local` mode
In order to work in `local` mode, you need to have docker installed. When running from you local machine, please make sure that you have docker or docker-compose (for local CPU machines) and nvidia-docker (for local GPU machines) installed. Alternatively, when running from a SageMaker notebook instance, you can simply run the following script to install dependenceis.
Note, you can only run a single local notebook at one time.
```
# only run from SageMaker notebook instance
if local_mode:
!/bin/bash ./common/setup.sh
```
## Setup the environment
We create a file called `src/patient_envs.py` for our modified environments. We can create a custom environment class or create a function that returns our environment. Since we're using Open AI Gym environment and wrappers, we just create functions that take the classic control environments `MountainCarEnv` and `Continuous_MountainCarEnv` and wrap them with a `TimeLimit` where we specify the `max_episode_steps` to 10,000.
```
!pygmentize src/patient_envs.py
```
## Configure the presets for RL algorithm
The presets that configure the RL training jobs are defined in the "preset-mountain-car-continuous-clipped-ppo.py" file which is also uploaded on the /src directory. Also see "preset-mountain-car-dqn.py" for the discrete environment case. Using the preset file, you can define agent parameters to select the specific agent algorithm. You can also set the environment parameters, define the schedule and visualization parameters, and define the graph manager. The schedule presets will define the number of heat up steps, periodic evaluation steps, training steps between evaluations.
These can be overridden at runtime by specifying the RLCOACH_PRESET hyperparameter. Additionally, it can be used to define custom hyperparameters.
```
!pygmentize src/preset-mountain-car-continuous-clipped-ppo.py
```
## Write the Training Code
The training code is written in the file “train-coach.py” which is uploaded in the /src directory.
We create a custom `SageMakerCoachPresetLauncher` which sets the default preset, maps and ties hyperparameters.
```
!pygmentize src/train-coach.py
```
## Train the RL model using the Python SDK Script mode
If you are using local mode, the training will run on the notebook instance. When using SageMaker for training, you can select a GPU or CPU instance. The RLEstimator is used for training RL jobs.
1. Specify the source directory where the environment, presets and training code is uploaded.
2. Specify the entry point as the training code
3. Specify the choice of RL toolkit and framework. This automatically resolves to the ECR path for the RL Container.
4. Define the training parameters such as the instance count, job name, S3 path for output and job name.
5. Specify the hyperparameters for the RL agent algorithm. The RLCOACH_PRESET can be used to specify the RL agent algorithm you want to use.
6. Define the metrics definitions that you are interested in capturing in your logs. These can also be visualized in CloudWatch and SageMaker Notebooks.
We use a variant of Proximal Policy Optimization (PPO) called Clipped PPO, which removes the need for complex KL divergence calculations.
```
if local_mode:
instance_type = 'local'
else:
instance_type = "ml.m4.4xlarge"
estimator = RLEstimator(entry_point="train-coach.py",
source_dir='src',
dependencies=["common/sagemaker_rl"],
toolkit=RLToolkit.COACH,
toolkit_version='0.11.0',
framework=RLFramework.MXNET,
role=role,
instance_type=instance_type,
instance_count=1,
output_path=s3_output_path,
base_job_name=job_name_prefix,
hyperparameters = {
"RLCOACH_PRESET": "preset-mountain-car-continuous-clipped-ppo", # "preset-mountain-car-dqn",
"discount": 0.995,
"gae_lambda": 0.997,
"evaluation_episodes": 3,
# approx 100 episodes
"improve_steps": 100000,
# approx 5 episodes to start with
"training_freq_env_steps": 75000,
"training_learning_rate": 0.004,
"training_batch_size": 256,
# times number below by training_freq_env_steps to get total samples per policy training
"training_epochs": 15,
'save_model': 1
}
)
estimator.fit(wait=local_mode)
```
## Store intermediate training output and model checkpoints
The output from the training job above is stored on S3. The intermediate folder contains gifs and metadata of the training.
```
job_name=estimator._current_job_name
print("Job name: {}".format(job_name))
s3_url = "s3://{}/{}".format(s3_bucket,job_name)
if local_mode:
output_tar_key = "{}/output.tar.gz".format(job_name)
else:
output_tar_key = "{}/output/output.tar.gz".format(job_name)
intermediate_folder_key = "{}/output/intermediate/".format(job_name)
output_url = "s3://{}/{}".format(s3_bucket, output_tar_key)
intermediate_url = "s3://{}/{}".format(s3_bucket, intermediate_folder_key)
print("S3 job path: {}".format(s3_url))
print("Output.tar.gz location: {}".format(output_url))
print("Intermediate folder path: {}".format(intermediate_url))
tmp_dir = "/tmp/{}".format(job_name)
os.system("mkdir {}".format(tmp_dir))
print("Create local folder {}".format(tmp_dir))
```
## Visualization
### Plot metrics for training job
We can pull the reward metric of the training and plot it to see the performance of the model over time.
```
%matplotlib inline
import pandas as pd
csv_file_name = "worker_0.simple_rl_graph.main_level.main_level.agent_0.csv"
key = os.path.join(intermediate_folder_key, csv_file_name)
wait_for_s3_object(s3_bucket, key, tmp_dir)
csv_file = "{}/{}".format(tmp_dir, csv_file_name)
df = pd.read_csv(csv_file)
df = df.dropna(subset=['Training Reward'])
x_axis = 'Episode #'
y_axis = 'Training Reward'
if len(df) > 0:
plt = df.plot(x=x_axis,y=y_axis, figsize=(12,5), legend=True, style='b-')
plt.set_ylabel(y_axis)
plt.set_xlabel(x_axis)
```
### Visualize the rendered gifs
The latest gif file found in the gifs directory is displayed. You can replace the tmp.gif file below to visualize other files generated.
```
key = os.path.join(intermediate_folder_key, 'gifs')
wait_for_s3_object(s3_bucket, key, tmp_dir)
print("Copied gifs files to {}".format(tmp_dir))
glob_pattern = os.path.join("{}/*.gif".format(tmp_dir))
gifs = [file for file in glob.iglob(glob_pattern, recursive=True)]
extract_episode = lambda string: int(re.search('.*episode-(\d*)_.*', string, re.IGNORECASE).group(1))
gifs.sort(key=extract_episode)
print("GIFs found:\n{}".format("\n".join([os.path.basename(gif) for gif in gifs])))
# visualize a specific episode
gif_index = -1 # since we want last gif
gif_filepath = gifs[gif_index]
gif_filename = os.path.basename(gif_filepath)
print("Selected GIF: {}".format(gif_filename))
os.system("mkdir -p ./src/tmp/ && cp {} ./src/tmp/{}.gif".format(gif_filepath, gif_filename))
HTML('<img src="./src/tmp/{}.gif">'.format(gif_filename))
```
## Evaluation of RL models
We use the last checkpointed model to run evaluation for the RL Agent.
### Load checkpointed model
Checkpointed data from the previously trained models will be passed on for evaluation / inference in the checkpoint channel. In local mode, we can simply use the local directory, whereas in the SageMaker mode, it needs to be moved to S3 first.
```
wait_for_s3_object(s3_bucket, output_tar_key, tmp_dir)
if not os.path.isfile("{}/output.tar.gz".format(tmp_dir)):
raise FileNotFoundError("File output.tar.gz not found")
os.system("tar -xvzf {}/output.tar.gz -C {}".format(tmp_dir, tmp_dir))
if local_mode:
checkpoint_dir = "{}/data/checkpoint".format(tmp_dir)
else:
checkpoint_dir = "{}/checkpoint".format(tmp_dir)
print("Checkpoint directory {}".format(checkpoint_dir))
if local_mode:
checkpoint_path = 'file://{}'.format(checkpoint_dir)
print("Local checkpoint file path: {}".format(checkpoint_path))
else:
checkpoint_path = "s3://{}/{}/checkpoint/".format(s3_bucket, job_name)
if not os.listdir(checkpoint_dir):
raise FileNotFoundError("Checkpoint files not found under the path")
os.system("aws s3 cp --recursive {} {}".format(checkpoint_dir, checkpoint_path))
print("S3 checkpoint file path: {}".format(checkpoint_path))
```
### Run the evaluation step
Use the checkpointed model to run the evaluation step.
```
estimator_eval = RLEstimator(role=role,
source_dir='src/',
dependencies=["common/sagemaker_rl"],
toolkit=RLToolkit.COACH,
toolkit_version='0.11.0',
framework=RLFramework.MXNET,
entry_point="evaluate-coach.py",
instance_count=1,
instance_type=instance_type,
hyperparameters = {
"RLCOACH_PRESET": "preset-mountain-car-continuous-clipped-ppo",
"evaluate_steps": 10000*2 # evaluate on 2 episodes
}
)
estimator_eval.fit({'checkpoint': checkpoint_path})
```
### Visualize the output
Optionally, you can run the steps defined earlier to visualize the output
## Model deployment
Since we specified MXNet when configuring the RLEstimator, the MXNet deployment container will be used for hosting.
```
predictor = estimator.deploy(initial_instance_count=1,
instance_type=instance_type,
entry_point='deploy-mxnet-coach.py')
```
We can test the endpoint with 2 samples observations. Starting with the car on the right side, but starting to fall back down the hill.
```
output = predictor.predict(np.array([0.5, -0.5]))
action = output[1][0]
action
```
We see the policy decides to move the car to the left (negative value). And similarly in the other direction.
```
output = predictor.predict(np.array([-0.5, 0.5]))
action = output[1][0]
action
```
### Clean up endpoint
```
predictor.delete_endpoint()
```
| github_jupyter |
```
# Import the modules
import sqlite3
import spiceypy
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
# Establish a connection to the comet database
CON = sqlite3.connect('../_databases/_comets/mpc_comets.db')
# Extract information about the comet 67P
COMET_67P_FROM_DB = pd.read_sql('SELECT NAME, PERIHELION_AU, ' \
'SEMI_MAJOR_AXIS_AU, ' \
'APHELION_AU, ECCENTRICITY, ' \
'ARG_OF_PERIH_DEG, LONG_OF_ASC_NODE_DEG ' \
'FROM comets_main WHERE NAME LIKE "67P%"', CON)
# Print the orbital elements of 67P
print(f'{COMET_67P_FROM_DB.iloc[0]}')
# Load SPICE kernels (meta file)
spiceypy.furnsh('kernel_meta.txt')
# Get the G*M value for the Sun
_, GM_SUN_PRE = spiceypy.bodvcd(bodyid=10, item='GM', maxn=1)
GM_SUN = GM_SUN_PRE[0]
# Set an initial and end time
INI_DATETIME = pd.Timestamp('2004-01-01')
END_DATETIME = pd.Timestamp('2014-01-01')
# Create a numpy array with 1000 timesteps between the initial and end time
datetime_range = np.linspace(INI_DATETIME.value, END_DATETIME.value, 1000)
# Convert the numpy arraay to a pandas date-time object
datetime_range = pd.to_datetime(datetime_range)
# Set an initial dataframe for the 67P computations
comet_67p_df = pd.DataFrame([])
# Set the UTC date-times
comet_67p_df.loc[:, 'UTC'] = datetime_range
# Convert the UTC date-time strings to ET
comet_67p_df.loc[:, 'ET'] = comet_67p_df['UTC'].apply(lambda x: \
spiceypy.utc2et(x.strftime('%Y-%m-%dT%H:%M:%S')))
# Compute the ET corresponding state vectors
comet_67p_df.loc[:, 'STATE_VEC'] = \
comet_67p_df['ET'].apply(lambda x: spiceypy.spkgeo(targ=1000012, \
et=x, \
ref='ECLIPJ2000', \
obs=10)[0])
# Compute the state vectors corresponding orbital elements
comet_67p_df.loc[:, 'STATE_VEC_ORB_ELEM'] = \
comet_67p_df.apply(lambda x: spiceypy.oscltx(state=x['STATE_VEC'], \
et=x['ET'], \
mu=GM_SUN), \
axis=1)
# Assign miscellaneous orbital elements as individual columns
# Set the perihelion. Convert km to AU
comet_67p_df.loc[:, 'PERIHELION_AU'] = \
comet_67p_df['STATE_VEC_ORB_ELEM'].apply(lambda x: \
spiceypy.convrt(x[0], \
inunit='km', \
outunit='AU'))
# Set the eccentricity
comet_67p_df.loc[:, 'ECCENTRICITY'] = \
comet_67p_df['STATE_VEC_ORB_ELEM'].apply(lambda x: x[1])
# Set the inclination in degrees
comet_67p_df.loc[:, 'INCLINATION_DEG'] = \
comet_67p_df['STATE_VEC_ORB_ELEM'].apply(lambda x: np.degrees(x[2]))
# Set the longitude of ascending node in degrees
comet_67p_df.loc[:, 'LONG_OF_ASC_NODE_DEG'] = \
comet_67p_df['STATE_VEC_ORB_ELEM'].apply(lambda x: np.degrees(x[3]))
# Set the argument of perihelion in degrees
comet_67p_df.loc[:, 'ARG_OF_PERIH_DEG'] = \
comet_67p_df['STATE_VEC_ORB_ELEM'].apply(lambda x: np.degrees(x[4]))
# Set the semi-major axis in AU
comet_67p_df.loc[:, 'SEMI_MAJOR_AXIS_AU'] = \
comet_67p_df['STATE_VEC_ORB_ELEM'].apply(lambda x: \
spiceypy.convrt(x[-2], \
inunit='km', \
outunit='AU'))
# Compute the aphelion, based on the semi-major axis and eccentricity
comet_67p_df.loc[:, 'APHELION_AU'] = \
comet_67p_df.apply(lambda x: x['SEMI_MAJOR_AXIS_AU'] \
* (1.0 + x['ECCENTRICITY']), \
axis=1)
# Let's plot the perihelion, eccentricity and argument of perihelion
# Let's set a dark background
plt.style.use('dark_background')
# Set a default font size for better readability
plt.rcParams.update({'font.size': 14})
# We plot the data dynamically in a for loop. col_name represents the column
# name for both dataframes; ylabel_name is used to change the y label.
for col_name, ylabel_name in zip(['PERIHELION_AU', \
'ECCENTRICITY', \
'ARG_OF_PERIH_DEG'], \
['Perihelion in AU', \
'Eccentricity', \
'Arg. of. peri. in degrees']):
# Set a figure with a certain figure size
fig, ax = plt.subplots(figsize=(12, 8))
# Line plot of the parameter vs. the UTC date-time from the SPICE data
ax.plot(comet_67p_df['UTC'], \
comet_67p_df[col_name], \
color='tab:orange', alpha=0.7, label='SPICE Kernel')
# As a guideline, plot the parameter data from the MPC data set as a
# horizontal line
ax.hlines(y=COMET_67P_FROM_DB[col_name], \
xmin=INI_DATETIME, \
xmax=END_DATETIME, \
color='tab:orange', linestyles='dashed', label='MPC Data')
# Set a grid for better readability
ax.grid(axis='both', linestyle='dashed', alpha=0.2)
# Set labels for the x and y axis
ax.set_xlabel('Time in UTC')
ax.set_ylabel(ylabel_name)
# Now we set a legend. However, the marker opacity in the legend has the
# same value as in the plot ...
leg = ax.legend(fancybox=True, loc='upper right', framealpha=1)
# ... thus, we set the markers' opacity to 1 with this small code
for lh in leg.legendHandles:
lh.set_alpha(1)
# Save the plot in high quality
plt.savefig(f'67P_{col_name}.png', dpi=300)
# Assignments:
# 1. Does the Tisserand Parameter w.r.t. Jupiter change over time?
# 2. Visualise the distance between Jupiter and 67P between the initial and
# end time. Use spiceypy.spkgps for this purpose and think about the
# targ and obs parameters. Convert the results in AU.
```
| github_jupyter |
# Navigation
---
You are welcome to use this coding environment to train your agent for the project. Follow the instructions below to get started!
### 1. Start the Environment
Run the next code cell to install a few packages. This line will take a few minutes to run!
```
!pip -q install ../python
```
The environment is already saved in the Workspace and can be accessed at the file path provided below. Please run the next code cell without making any changes.
```
from unityagents import UnityEnvironment
import numpy as np
# please do not modify the line below
# env = UnityEnvironment(file_name="/data/Banana_Linux_NoVis/Banana.x86_64")
env = UnityEnvironment(file_name="./Banana.app")
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
### 2. Examine the State and Action Spaces
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
Note that **in this coding environment, you will not be able to watch the agent while it is training**, and you should set `train_mode=True` to restart the environment.
```
env_info = env.reset(train_mode=True)[brain_name] # reset the environment
state = env_info.vector_observations[0] # get the current state
score = 0 # initialize the score
while True:
action = np.random.randint(action_size) # select an action
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
```
When finished, you can close the environment.
```
env.close()
```
### 4. It's Your Turn!
Now it's your turn to train your own agent to solve the environment! A few **important notes**:
- When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
- To structure your work, you're welcome to work directly in this Jupyter notebook, or you might like to start over with a new file! You can see the list of files in the workspace by clicking on **_Jupyter_** in the top left corner of the notebook.
- In this coding environment, you will not be able to watch the agent while it is training. However, **_after training the agent_**, you can download the saved model weights to watch the agent on your own machine!
# Train
```
!pip -q install ./python
from unityagents import UnityEnvironment
import numpy as np
env = UnityEnvironment(file_name="/data/Banana_Linux_NoVis/Banana.x86_64")
brain_name = env.brain_names[0]
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
state_size = len(state)
brain = env.brains[brain_name]
action_size = brain.vector_action_space_size
import model
import dqn_agent
from importlib import reload
reload(model)
reload(dqn_agent)
import torch
import dqn_agent
seed = 1234
agent = dqn_agent.Agent(state_size, action_size, seed)
eps_decay = 0.999
eps_min = 0.02
n_epi = 2000
i_save = 1000
window_size = 100
scores = []
save_dir = 'ckpts'
import os
os.makedirs(save_dir, exist_ok=True)
eps = 1.
for i_epi in range(1, n_epi + 1):
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
score = 0
while True:
action = agent.act(state)
env_info = env.step(action)[brain_name]
next_state = env_info.vector_observations[0]
reward = env_info.rewards[0]
done = env_info.local_done[0]
agent.step(state, action, reward, next_state, done)
score += reward
state = next_state
if done:
break
eps = max(eps * eps_decay, eps_min)
scores.append(score)
mean_score = np.mean(scores[-window_size:])
print("[{}/{}] eps: {}, score: {}, mean_score: {}".format(
i_epi, n_epi, eps, score, mean_score), end='\r')
if i_epi % i_save == 0:
torch.save(agent.qnetwork_local.state_dict(),
os.path.join(save_dir, '{}_{}'.format(i_epi, int(10000 * mean_score))))
if mean_score >= 15:
torch.save(agent.qnetwork_local.state_dict(),
os.path.join(save_dir, '{}_{}'.format(i_epi, int(10000 * mean_score))))
break
torch.save(agent.qnetwork_local.state_dict(), 'model.ckpt')
print(scores[-window_size:])
import matplotlib.pyplot as plt
%matplotlib inline
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.plot(np.arange(window_size, len(scores)+1),
[np.mean(scores[i-window_size:i]) for i in range(window_size, len(scores)+1)])
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# env.close()
```
# Evaluation
```
!pip -q install ./python
from unityagents import UnityEnvironment
import numpy as np
env = UnityEnvironment(file_name="./Banana.app")
brain_name = env.brain_names[0]
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
state_size = len(state)
brain = env.brains[brain_name]
action_size = brain.vector_action_space_size
import dqn_agent
import torch
seed = 1234
agent = dqn_agent.Agent(state_size, action_size, seed)
agent.qnetwork_local.load_state_dict(torch.load('model.ckpt'))
brain_name = env.brain_names[0]
scores = []
for i in range(100):
print("[%f%%]" % (i+1), end='\r')
score = 0
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
while True:
action = agent.act(state)
env_info = env.step(action)[brain_name]
next_state = env_info.vector_observations[0]
reward = env_info.rewards[0]
done = env_info.local_done[0]
score += reward
state = next_state
if done:
break
scores.append(score)
print("Mean score: {}".format(sum(scores) / len(scores)))
import time
score = 0
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
while True:
action = agent.act(state)
env_info = env.step(action)[brain_name]
next_state = env_info.vector_observations[0]
reward = env_info.rewards[0]
done = env_info.local_done[0]
score += reward
state = next_state
if done:
break
time.sleep(0.033)
env.close()
```
| github_jupyter |
**Before you start**
Make sure you have completed "Tutorial 1 - Exploring the data" before starting this one as it creates a data file that will be used below.
## **Preparing the Discovery data for Machine Learning**
Having performed our initial data analysis and produced a filtered dataset to work with, we will now move on to preparing the data for machine learning.
The first step is to create a numerical representation of the data. We will start with the simple way, and then a slightly more sophisticated way which is very common in data science.
Then we can look at an approach called stemming which deals with things like pluralisation of words (you may want to treat 'machines' and 'machine' as the same thing in your model).
At the end of this tutorial we will have 3 datasets which we will experiment with in the next one.
```
import sys
import os
# Where do you want to get data from - Google Drive or Github
data_source = "Github" # Change to either Github or Drive - if Drive, copy the data into a folder called "Data" within your "My Drive folder"
if data_source == "Github":
!git clone https://github.com/nationalarchives/MachineLearningClub.git
sys.path.insert(0, 'MachineLearningClub')
data_folder = "MachineLearningClub/Session 3/Data/"
os.listdir(data_folder)
else:
# Connect to gdrive
from google.colab import drive
drive.mount('/content/gdrive')
data_folder = "/content/gdrive/My Drive/Data/"
os.listdir(data_folder)
```
This piece of code imports the libraries that we will be using in the first part of this notebook.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
import sklearn # The most common Python Machine Learning library - scikit learn
from sklearn.model_selection import train_test_split # Used to create training and test data
from sklearn.metrics import confusion_matrix, accuracy_score, f1_score, balanced_accuracy_score
import seaborn as sns
from sklearn import svm
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.naive_bayes import GaussianNB, MultinomialNB
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import SGDClassifier, LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.metrics import precision_score, recall_score, confusion_matrix, classification_report, accuracy_score, f1_score
from operator import itemgetter
from nltk.stem.porter import *
import pickle
from nltk import toktok
```
Run the following code to load the reduced dataset you created in the previous tutorial.
```
descriptions = pd.read_csv(data_folder + 'topN_taxonomy.csv',
delimiter="|", header=None, lineterminator='\n')
descriptions = descriptions.drop(descriptions.columns[[0]], axis=1)
descriptions.columns = ["IAID","TAXID","Description"]
descriptions.count()
```
This time there are no rows with blanks.
We will also load up the table of taxonomy category names again which will be useful for understanding the various categories.
```
taxonomy = pd.read_csv(data_folder + 'taxonomyids_and_names.txt',
delimiter="|", header=None, lineterminator='\n')
taxonomy.columns = ["TAXID","TaxonomyCategory"]
```
### **Bag-of-Words**
Do you remember the Bag-of-Words technique from the first session? This is a way of identifying the top words or phrases in a corpus.
We will now go into more detail of what was happening behind the scenes. There are a number of ways of turning text into numbers. One is simply to create a table with a column for every word that appears in the corpus and then for each record, count how many times each word occurs in the text for that row.
For example, imagine we have two sentences:
* the dog laughed
* the dog walked the dog
Our table would have have a column for each unique word: 'the', 'dog', 'laughed', 'walked', a row for each sentence. **Run the code below to see what it looks like**
```
pd.DataFrame({'text' : ['the dog laughed','the dog walked the dog'], 'the' : [1,2], 'dog' : [1,2], 'laughed' : [1,0], 'walked' : [0,1]})
```
The following code does the same thing creating a function which will convert text into a 'vector'. (There's no need to run the code, it is just here for discussion purposes). A vector is just a list of numbers, and in this situation is analogous to a row in a spreadsheet. The column headings are set by the vocabulary in the text we used to set up the function (which was our table of descriptions). The MAX_FEATURES variable defines how many columns there will be - it is the size of the vocabulary (in the example above our vocabulary was 4 words).
This is just what we used in the previous tutorial to get the top 10 words.
```
MAX_FEATURES = 500
count_vectorizer = CountVectorizer(max_features = MAX_FEATURES) # A library function to turn text into a vector of length 'MAX_FEATURES'
word_counts = count_vectorizer.fit_transform(descriptions.Description) # Use data to generate the column headings
count_vocab = count_vectorizer.get_feature_names() # This is just the names of the columns
```
Why not set MAX_FEATURES large enough to use all of the words in the corpus? In fact, why don't we use a giant dictionary to create all the column headings rather than restricting it to the words in our corpus?
In answer to the first question, we can do that but the primary reason for restricting the vocabulary is the amount of memory and computing power you have available. If you remember back to session 1, we restricted this variable for the Amazon data because the Colab could potentially crash. Another reason is that a lot of words may only appear once in the text and so they might not be particularly useful for machine learning. Often a machine learning algorithm will peform better with fewer features to work with.
In answer to the second question, there is no point in having columns outside of your corpus. When you fit an algorithm it will attach some weight to each word depending on the category of the row. Any word that doesn't appear in the corpus will therefore not pass through the algorithm and so that column will always get zero weighting. In other words it's just a waste of a column.
To illustrate the points above we will experiment with building a 'vectorizer' of different sizes and passing a new sentence into it to see what happens.
Run the code below. **Are all of the words in the sentence printed out?**
**What happens if you change MAX_FEATURES to 1000? Or 2000?**
**Try changing the sentence too**
```
MAX_FEATURES = 500
test_sentence = 'my records are about design and food and the war and design during the war'
count_vectorizer = CountVectorizer(max_features = MAX_FEATURES) # A library function to turn text into a vector of length 'MAX_FEATURES'
word_counts = count_vectorizer.fit_transform(descriptions.Description) # Use data to generate the column headings
count_vocab = count_vectorizer.get_feature_names() # This is just the names of the columns
sentence_counts = count_vectorizer.transform([test_sentence])
nz = sentence_counts.nonzero()
ft_names = count_vectorizer.get_feature_names()
pd.DataFrame([(ft_names[i], sentence_counts[0, i]) for i in nz[1]])
```
While this approach is simple to implement it has some issues. Firstly, long documents could end up with disproportionally high scores. For example, an essay about dogs could have a 20 in the 'dog' column. Is it more about dogs than the sentences we had at the beginning?
Secondly, words that appear regularly in the corpus will have high values but may not be especially meaningful. Think of how often 'record' might appear in a set of documents about archiving. It is effectively a stop word!
The most common approach for turning text into numbers is to use a score called the TF-IDF score. The TF stands for Term Frequency, which is just the same as counting words. The IDF stands for Inverse Document Frequency, and this is what makes it a better system. IDF is a count of the number of documents that a word appears in. (If you're interested in how the calculation is made, see http://www.tfidf.com/)
As a quick intuition though, if a word appears in lots of documents it will end up with a low score. If it appears several times in one document but is rare across the corpus it will have a high score.
Let's run the previous example again but this time using the TF-IDF scorer.
**Compare the scores for 'the', 'and', and 'are' to the CountVectorizer results above**
**Which is the top scoring word?**
**Why do you think 'design' has scored so low, despite appearing twice?**
**Where is food in the scoring?**
Note: You may notice a new command in the code below - pickle.dump
This is a method that Python uses for saving complex objects that can't be easily output into a CSV file. The TfidfVectorizer counts as a complex object. The reason we are using this here is to save this TFIDF representation of the text for the next tutorial. It is also a format for the more technically minded to consider when thinking about how to archive an ML model in the final session.
```
MAX_FEATURES = 2000
test_sentence = 'my records are about design and food and the war and design during the war'
tfidf_vectorizer = TfidfVectorizer(max_features = MAX_FEATURES)
word_tfidf = tfidf_vectorizer.fit_transform(descriptions.Description)
pickle.dump(tfidf_vectorizer, open(data_folder + 'word_tfidf.pck','wb'))
tfidf_vocab = tfidf_vectorizer.get_feature_names()
sentence_counts = tfidf_vectorizer.transform([test_sentence])
nz = sentence_counts.nonzero()
ft_names = tfidf_vectorizer.get_feature_names()
pd.DataFrame([(ft_names[i], sentence_counts[0, i]) for i in nz[1]])
```
If you think back to session 1, you may remember we used this approach to find the 2 and 3 word phrases (n-grams) in the Amazon reviews. We will do that again for this dataset. We're not going to analyze the phrases in our data now (although if you want to, you can go back to Tutorial 1 and add the "ngram_range=[1,3]" into the code where we printed out the top 10 words), but looking at our test sentence again is informative.
First we rebuild the TfidfVectorizer with phrases between 1 and 3 words long. Then pass in the test sentence again. As you can see only one two word phrase ("and the") is found in the feature set. If you **change the MAX_FEATURES to 25000**, you will get some more 2 word ones but still no triplets. This is a feature of language. There may be lots of common words but they mostly used in unique combinations, unless you're working with a large corpus. This suggests that when we work with phrases rather than words we need far more features, so again we have to decide on what is feasible given our computational resources.
**Note: using 20000 features made the machine crash in the 3rd tutorial, which demonstrates this point**
```
MAX_FEATURES = 5000
test_sentence = 'my records are about design and food and the war and design during the war'
tfidf_vectorizer = TfidfVectorizer(max_features = MAX_FEATURES, ngram_range=[1,3])
tfidf_vectorizer.fit_transform(descriptions.Description)
pickle.dump(tfidf_vectorizer, open(data_folder + 'ngram_tfidf.pck','wb'))
tfidf_vocab = tfidf_vectorizer.get_feature_names()
sentence_counts = tfidf_vectorizer.transform([test_sentence])
nz = sentence_counts.nonzero()
ft_names = tfidf_vectorizer.get_feature_names()
pd.DataFrame([(ft_names[i], sentence_counts[0, i]) for i in nz[1]])
```
So we'll start with 5000 features for the n-gram dataset that we're about to create, and if that works ok feel free to return here and bump up the value.
**Re-run the code above with MAX_FEATURES set to 5000 before continuing**
**Stemming**
Stemming is another technique we can use to prepare our data. It is used to standardise words by removing suffixes, such as the 's' or 'es' at the end of a plural form. It is used, for example, in Discovery's search system. There are several stemming algorithms available but we're going to choose the popular Porter stemmer from the NLTK (Natural Language Tool Kit) library.
Best way to see what it does is to try it out. Change the words in the list to try different endings, and see if you can get a feel for what it is and isn't good at.
```
word_list = ['grows','grow','growing','leave','leaves','leaf','fairly']
ps = PorterStemmer()
for w in word_list:
print(w,"becomes",ps.stem(w))
```
We won't delve into the results of applying this to our corpus but we are going to create a third dataset so that we can see what (if any) difference stemming makes to the results. One observation is that stemming reduces the vocabulary size (because variations of words become one) so we should need fewer features. We will start with 4000 (this is the n-gram version) and work from there.
This bit of code is a little different to the earlier ones as we have to create a function which does the stemming and then pass it to the TFIDFVectorizer. This is because the library we're using to do the vectorising, sklearn, doesn't do stemming because it is a machine learning library not a Natural Language Processing (NLP) one. To do the stemming we are using the NLTK library. (**Note: this one might take a little while to run as the stemmer isn't super fast.)
**Compare the numbers output by this function to the ones in the other two versions**
**Why do you think they are different?**
```
MAX_FEATURES = 4000
test_sentence = 'my records are about design and food and the war and design during the war'
class PorterTokenizer:
def __init__(self):
self.porter = PorterStemmer()
def __call__(self, doc):
ttt = toktok.ToktokTokenizer()
return [self.porter.stem(t) for t in ttt.tokenize(doc)]
tfidf_vectorizer = TfidfVectorizer(max_features = MAX_FEATURES, ngram_range=[1,3], tokenizer=PorterTokenizer())
tfidf_vectorizer.fit_transform(descriptions.Description)
pickle.dump(tfidf_vectorizer, open(data_folder + 'stemmed_ngram_tfidf.pck','wb'))
tfidf_vocab = tfidf_vectorizer.get_feature_names()
sentence_counts = tfidf_vectorizer.transform([test_sentence])
nz = sentence_counts.nonzero()
ft_names = tfidf_vectorizer.get_feature_names()
pd.DataFrame([(ft_names[i], sentence_counts[0, i]) for i in nz[1]])
```
That's the end of this tutorial in which we have:
1. Learned about TF-IDF
2. And why it is better than just counting words
1. Learned about using stemming to standardise words
2. Created 3 datasets for use in the next tutorial
We're now ready to start Machine Learning with Discovery data!
| github_jupyter |
+++
title = "Working with (Vanilla) PyTorch"
+++
This guide goes through how to use this package with the standard PyTorch workflow.
```
!pip3 install tabben torch
```
## Loading the data
For this example, we'll use the poker hand prediction dataset.
```
from tabben.datasets import OpenTabularDataset
ds = OpenTabularDataset('./data/', 'poker')
```
And let's just look at the input and output attributes:
```
print('Input Attribute Names:')
print(ds.input_attributes)
print('Output Attribute Names:')
print(ds.output_attributes)
```
Since we're working with PyTorch, the `OpenTabularDataset` object above is a PyTorch `Dataset` object that we can directly feed into a `DataLoader`.
```
from torch.utils.data import DataLoader
dl = DataLoader(ds, batch_size=8)
example_batch = next(iter(dl))
print(f'Input Shape (Batched): {example_batch[0].shape}')
print(f'Output Shape (Batched): {example_batch[1].shape}')
```
## Setting up a basic model
First, we'll create a basic model in PyTorch, just for illustration (you can replace this with whatever model you're trying to train/evaluate). It'll just be a feedforward neural network with a couple dense/linear layers (this probably won't perform well).
```
from torch import nn
import torch.nn.functional as F
class ShallowClassificationNetwork(nn.Module):
def __init__(self, num_inputs, num_classes):
super().__init__()
self.linear1 = nn.Linear(num_inputs, 32)
self.linear2 = nn.Linear(32, 32)
self.linear3 = nn.Linear(32, num_classes)
def forward(self, inputs):
# [b, num_inputs] -> [b, 32]
x = F.relu(self.linear1(inputs))
# [b, 32] -> [b, 32]
x = F.relu(self.linear2(x))
# [b, 32] -> [b, num_classes] (log(softmax(.)) computed over each row)
x = F.log_softmax(self.linear3(x), dim=1)
return x
def predict(self, inputs):
x = F.relu(self.linear1(inputs))
x = F.relu(self.linear2(x))
return F.softmax(self.linear3(x), dim=1)
import torch
device = torch.device('cpu') # change this to 'cuda' if you have access to a CUDA GPU
model = ShallowClassificationNetwork(ds.num_inputs, ds.num_classes).to(device)
```
## Training the model
Now that we have a basic model and a training dataset (the default split for `OpenTabularDataset` is the train split), we can train our simple network using a PyTorch training loop.
```
from torch import optim
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.05, momentum=0.9)
from tqdm import trange
model.train()
training_progress = trange(30, desc='Train epoch')
for epoch in training_progress:
running_loss = 0.
running_acc = 0.
running_count = 0
for batch_input, batch_output in dl:
optimizer.zero_grad()
outputs = model(batch_input.float().to(device))
preds = outputs.argmax(dim=1)
loss = criterion(outputs, batch_output)
loss.backward()
optimizer.step()
running_loss += loss.item()
running_acc += (preds == batch_output).sum()
running_count += batch_input.size(0)
training_progress.set_postfix({
'train loss': f'{running_loss / running_count:.4f}',
'train acc': f'{100 * running_acc / running_count:.2f}%',
})
```
You can play around with the hyperparameters, but the model isn't likely to get particularly good performance like this. But, we'll go ahead and evaluate the final model (we ignored having a validation set in this guide) on the test set.
## Evaluating on the test set
```
test_ds = OpenTabularDataset('./data/', 'poker', split='test')
test_dl = DataLoader(test_ds, batch_size=16)
print(len(test_ds))
```
Let's run the model and save its outputs for later evaluation (since we left the softmax operation for the loss when we defined the model above, we'll need to softmax the outputs to get probabilities).
```
model.eval()
pred_outputs = []
gt_outputs = []
for test_inputs, test_outputs in test_dl:
batch_outputs = model(test_inputs.float().to(device))
pred_outputs.append(batch_outputs.detach().cpu())
gt_outputs.append(test_outputs)
test_pred_outputs = torch.softmax(torch.vstack(pred_outputs).detach().cpu(), axis=1)
test_gt_outputs = torch.hstack(gt_outputs).detach().cpu()
```
We can get the standard set of metrics and then evaluate the outputs of the test set on them.
```
from tabben.evaluators import get_metrics
eval_metrics = get_metrics(ds.task, classes=ds.num_classes)
for metric in eval_metrics:
print(f'{metric}: {metric(test_gt_outputs, test_pred_outputs)}')
```
---
This code was last run with the following versions (if you're looking at the no-output webpage, see the notebook in the repository for versions):
```
from importlib.metadata import version
packages = ['torch', 'tabben']
for pkg in packages:
print(f'{pkg}: {version(pkg)}')
```
| github_jupyter |
```
# default_exp processing
#hide
%load_ext autoreload
%autoreload 2
```
# processing
> Processing the different stream of data to calculate responses of retinal cells
```
#export
from functools import partial
import numpy as np
from sklearn.decomposition import PCA
from sklearn import cluster
import scipy.ndimage as ndimage
import scipy.signal as signal
import scipy as sp
from cmath import *
import itertools
import random
from theonerig.core import *
from theonerig.utils import *
from theonerig.modelling import *
from theonerig.leddome import *
```
# Spatial stimulus (checkerboard)
```
#export
def eyetrack_stim_inten(stim_inten, eye_track,
upsampling=2,
eye_calib=[[94 ,8], [ 18, 59]],
box_w=None, box_h=None, stim_axis="x"):
"""
From stimulus data and eye tracking, returns a corrected and upsampled stimulus data.
Calibration corresponds to the width and height of the stimulus screen, in
terms of pixels in the eye video: [[WIDTH_x, WIDTH_y], [HEIGHT_x, HEIGHT_y]]
params:
- stim_inten: Stimulus intensity matrix of shape (t, y, x), or (t, x) or (t, y) depending on stim_axis
- eye_track: Eye tracking data of shape (t, x_pos, y_pos, ...)
- upsampling: Factor for the upsampling (2 will multiply by 2 number of box in width and height)
- eye_calib: Calibration matrix of shape (2,2)
- box_w: Width of a block in pixel (40px in case of a 32 box in width of a checkerboard on a 1280px width)
- box_h: Height of a block in pixel. Both box_x and box_h are calculated from a 1280x720 screen if None
- stim_axis: Specify which direction to shift in case of stim shape different than (t, y, x)
return:
- Upsampled and shift corrected stimulus intensity
"""
eye_x, eye_y = eye_track[:,0], eye_track[:,1]
shape_y, shape_x = 1, 1
if len(stim_inten.shape)==2:
if stim_axis=="x":
shape_x = stim_inten.shape[1]
elif stim_axis=="y":
shape_y = stim_inten.shape[1]
elif len(stim_inten.shape)==3:
shape_y = stim_inten.shape[1]
shape_x = stim_inten.shape[2]
if box_w is None:
box_w = 1280//shape_x
if box_h is None:
box_h = 720//shape_y
if shape_y>1 and shape_x>1:
box_w, box_h = int(box_w/upsampling), int(box_h/upsampling)
elif shape_x > 1:
box_w, box_h = int(box_w/upsampling), box_h
elif shape_y > 1:
box_w, box_h = box_w , int(box_h/upsampling)
eye_transfo_f = _eye_to_stim_f(eye_calib=eye_calib,
box_width=box_w,
box_height=box_h)
if shape_y>1 and shape_x>1:
stim_inten = stim_inten.repeat(upsampling,axis=1).repeat(upsampling,axis=2)
else:
stim_inten = stim_inten.repeat(upsampling,axis=1)
xpos_avg = np.mean(eye_x)
ypos_avg = np.mean(eye_y)
mean_stim_inten = int((np.max(stim_inten)+np.min(stim_inten))/2)
#After getting the shift of the matrix to apply, we roll the matrix instead of extending it to the shifts
#This seems strange, but from the cell point of view, that is potentially looking at no stimulus,
# the response it gives are uncorrelated with the stimulus, and so shouldn't impact further analysis
# Advantage is that it keeps the data small enough, without loosing regions of the stimulus.
for i in range(len(stim_inten)):
x_eyeShift = eye_x[i]-xpos_avg
y_eyeShift = eye_y[i]-ypos_avg
stim_shift_x, stim_shift_y = eye_transfo_f(x_eyeShift=x_eyeShift,
y_eyeShift=y_eyeShift)
if shape_y>1 and shape_x>1:
rolled_stim = np.roll(stim_inten[i],stim_shift_y,axis=0)
rolled_stim = np.roll(rolled_stim ,stim_shift_x,axis=1)
else:
if stim_axis=="x":
rolled_stim = np.roll(stim_inten[i],stim_shift_x,axis=0)
else:
rolled_stim = np.roll(stim_inten[i],stim_shift_y,axis=0)
stim_inten[i] = rolled_stim
return stim_inten
def saccade_distances(eye_track):
"""
Create a mask for the eye position timeserie that indicate how far was the last saccade.
The eye positions need to be smoothed with smooth_eye_position.
params:
- eye_track: Eye tracking data of shape (t, x_pos, y_pos, ...)
return:
- Distance to last saccade array
"""
x_pos, y_pos = eye_track[:,0], eye_track[:,1]
saccade_pos = np.where((x_pos[1:] != x_pos[:-1]) & (y_pos[1:] != y_pos[:-1]))[0] + 1
len_chunks = [saccade_pos[0]]+list(saccade_pos[1:]-saccade_pos[:-1])
len_chunks.append(len(x_pos) - saccade_pos[-1])
saccade_mask = []
for len_chunk in len_chunks:
saccade_mask.extend(list(range(len_chunk)))
return np.array(saccade_mask)
def smooth_eye_position(eye_track, threshold=2):
"""
Smooth the eye positions. Uses clustering of eye position epochs to smooth.
Clustering is made with sklearn.cluster.dbscan.
params:
- eye_track: Eye tracking data of shape (t, x_pos, y_pos, ...)
- threshold: Distance threshold in the position clustering
return:
- Distance to last saccade array
"""
x_pos, y_pos = eye_position[:,0], eye_position[:,1]
X = np.stack((x_pos,y_pos, np.linspace(0, len(x_pos)/2, len(x_pos)))).T
clusters = cluster.dbscan(X, eps=threshold, min_samples=3, metric='minkowski', p=2)
move_events = np.where(clusters[1][1:] > clusters[1][:-1])[0] + 1
len_chunks = [move_events[0]]+list(move_events[1:]-move_events[:-1])
len_chunks.append(len(x_pos) - move_events[-1])
eye_x_positions = np.split(x_pos, move_events)
eye_y_positions = np.split(y_pos, move_events)
mean_x_pos = np.array(list(map(np.mean, eye_x_positions)))
mean_y_pos = np.array(list(map(np.mean, eye_y_positions)))
x_pos_smooth = np.concatenate([[x_pos]*len_chunk for x_pos,len_chunk in zip(mean_x_pos, len_chunks)])
y_pos_smooth = np.concatenate([[y_pos]*len_chunk for y_pos,len_chunk in zip(mean_y_pos, len_chunks)])
return np.stack((x_pos_smooth, y_pos_smooth)).T
def _eye_to_stim_f(eye_calib, box_width, box_height):
"""
Initialise stimulus shift function parameters and returns a partial function.
params:
- eye_calib: Calibration matrix of shape (2,2)
- box_width: Width of the elementary block in pixel
- box_height: Width of the elementary block in pixel
return:
- Partial function to obtain stimulus shift from eye positions
"""
eye_to_stim = np.linalg.inv(eye_calib)
box_dim = np.array([1280/(box_width), 720/(box_height)])
return partial(_linear_transform, box_dim=box_dim, transfo_matrix=eye_to_stim)
def _linear_transform(box_dim, transfo_matrix, x_eyeShift, y_eyeShift):
"""
Computes the shift of the stimulus from eye position shifts.
params:
- box_dim: Size in pixel of the box [width, height]
- transfo_matrix: Inverse matrix of the calibration matrix described in eyetrack_stim_inten
- x_eyeShift: Eye shift in x
- y_eyeShift: Eye shift in y
return:
- Stimulus shift tuple (delta_x, delta_y)
"""
transform_coord = np.dot(transfo_matrix, np.array([x_eyeShift, y_eyeShift]).T)
stim_vec = np.round(transform_coord * box_dim).astype(int)
return stim_vec[0], -stim_vec[1]
#export
def process_sta_batch(stim_inten, spike_counts, Hw=30, Fw=2, return_pval=False, normalisation="abs"):
"""
Computes the STA and associated pvalues in parallel for a batch of cells.
params:
- stim_inten: stimulus intensity matrix of shape (t, ...)
- spike_counts: cells activity matrix of shape (t, n_cell)
- Hw: Lenght in frames of the history window, including the 0 timepoint
- Fw: Lenght in frames of the forward window
- return_pval: Flag to signal whether or not to return the pvalues
- normalisation: Normalization applied to the STA. One of ["abs", "L2", None]
return:
- stas of shape (n_cell, Hw+Fw, ...)
- stas and pvalues if return_pval=True, both of shape (n_cell, Hw+Fw, ...)
"""
assert normalisation in ["abs", "L2", None], "normalisation must be one of ['abs', 'L2', None]"
#Preparing the stimulus
orig_shape = stim_inten.shape
stim_inten = stim_inten_norm(stim_inten)
sum_spikes = np.sum(spike_counts, axis=0)
len_stim = len(stim_inten)
#We just have to calculate one STA over the whole record
stim_inten = np.reshape(stim_inten, (len(stim_inten),-1))
stim_inten = np.transpose(stim_inten)
allCells_sta = staEst_fromBins(stim_inten, spike_counts, Hw, Fw=Fw)
if len(orig_shape)==3:
allCells_sta = allCells_sta.reshape((len(allCells_sta),Hw+Fw, orig_shape[-2], orig_shape[-1]))
elif len(orig_shape)==2:
allCells_sta = allCells_sta.reshape((len(allCells_sta),Hw+Fw,-1))
else:
allCells_sta = allCells_sta.reshape((len(allCells_sta),Hw+Fw))
if allCells_sta.shape[0]==1: #Only one cell, but we need to keep the axis
allCells_sta = np.squeeze(allCells_sta)
allCells_sta = np.expand_dims(allCells_sta, axis=0)
else:
allCells_sta = np.squeeze(allCells_sta)
if return_pval:
p_values = np.empty(allCells_sta.shape)
for k, cell_sta in enumerate(allCells_sta): #Easy way to do normalization for each cell that works for all possible shapes
if return_pval:
z_scores = cell_sta/ np.sqrt(1/sum_spikes[k]) #Standard score is calculated as (x-mean)/std
p_values[k] = sp.stats.norm.sf(abs(z_scores))*2
if normalisation is None:
continue
elif normalisation=="abs":
allCells_sta[k] = np.nan_to_num(cell_sta/np.max(np.abs(cell_sta)))
elif normalisation=="L2":
allCells_sta[k] = np.nan_to_num(cell_sta/np.sqrt(np.sum(np.power(cell_sta, 2))))
if return_pval:
return allCells_sta, p_values
else:
return allCells_sta
def staEst_fromBins(stim, spike_counts, Hw, Fw=0):
"""
Matrix mutliplication to compute the STA. Use the wrapper process_sta_batch to avoid bugs.
params:
- stim_inten: stimulus intensity matrix of shape (flattened_frame, t)
- spike_counts: cells activity matrix of shape (t, n_cell)
- Hw: Lenght in frames of the history window, including the 0 timepoint
- Fw: Lenght in frames of the forward window
return:
- STA of shape (n_cell, Hw+Fw, flattened_frame)
"""
spike_counts[:Hw] = 0
spike_counts = np.nan_to_num(spike_counts / np.sum(spike_counts,axis=0))
spike_counts = spike_counts - np.mean(spike_counts, axis=0) #Center to 0 the spike counts to include "inhibitory" stimulus
sta = np.zeros((Hw+Fw, stim.shape[0], spike_counts.shape[-1]))
for i in range(Hw): #Does the dot product of cell activity and frame intensity matrix
sta[(Hw-1-i),:,:] = np.dot(stim, spike_counts)
spike_counts = np.roll(spike_counts, -1, axis=0) #And shift the activity to compute the next frame
spike_counts = np.roll(spike_counts, Hw, axis=0)
if Fw != 0:
spike_counts[-Fw:] = 0
for i in range(Fw): #Same thing for Fw
spike_counts = np.roll(spike_counts, 1, axis=0)
sta[Hw+i,:,:] = np.dot(stim, spike_counts)
spike_counts = np.roll(spike_counts, -Fw, axis=0)
return np.transpose(sta, (2,0,1))
def process_sta_batch_large(stim_inten, spike_counts, Hw=30, Fw=2, return_pval=False, normalisation="abs", bs=1000):
"""
Computes the STA and associated pvalues in parallel for a batch of cells, for a large stimulus.
params:
- stim_inten: stimulus intensity matrix of shape (t, ...)
- spike_counts: cells activity matrix of shape (t, n_cell)
- Hw: Lenght in frames of the history window, including the 0 timepoint
- Fw: Lenght in frames of the forward window
- return_pval: Flag to signal whether or not to return the pvalues
- normalisation: Normalization applied to the STA. One of ["abs", "L2", None]
- bs: batch size to compute partial STA
return:
- stas of shape (n_cell, Hw+Fw, ...)
- stas and pvalues if return_pval=True, both of shape (n_cell, Hw+Fw, ...)
"""
orig_shape = stim_inten.shape
n_spatial_dim = orig_shape[1]*orig_shape[2]
sum_spikes = np.sum(spike_counts, axis=0)
len_stim = len(stim_inten)
allCells_sta = np.zeros((n_spatial_dim, spike_counts.shape[1], Hw+Fw))
stim_inten = stim_inten.reshape((len_stim,-1))
print("Computing the STA part by part:")
for i, batch_pos in enumerate(range(0, n_spatial_dim, bs)):
print(str(round(100*batch_pos/n_spatial_dim,2))+"% ", end="\r", flush=True)
#Computing STA on partial portions of the screen sequentially
stim_part = stim_inten_norm(stim_inten[:, batch_pos:batch_pos+bs]).T
sub_sta = staEst_fromBins(stim_part, spike_counts, Hw, Fw=Fw)
allCells_sta[batch_pos:batch_pos+bs] = np.transpose(sub_sta, (2,0,1))#(ncell,Hw,stim_len) to (stim_len,ncell,Hw)
allCells_sta = np.transpose(allCells_sta, (1,2,0))
print("100% ")
if len(orig_shape)==3:
allCells_sta = allCells_sta.reshape((len(allCells_sta),Hw+Fw, orig_shape[-2], orig_shape[-1]))
elif len(orig_shape)==2:
allCells_sta = allCells_sta.reshape((len(allCells_sta),Hw+Fw,-1))
else:
allCells_sta = allCells_sta.reshape((len(allCells_sta),Hw+Fw))
if allCells_sta.shape[0]==1: #Only one cell, but we need to keep the axis
allCells_sta = np.squeeze(allCells_sta)
allCells_sta = np.expand_dims(allCells_sta, axis=0)
else:
allCells_sta = np.squeeze(allCells_sta)
if return_pval:
p_values = np.empty(allCells_sta.shape)
for k, cell_sta in enumerate(allCells_sta): #Easy way to do normalization for each cell that works for all possible shapes
if return_pval:
z_scores = cell_sta/ np.sqrt(1/sum_spikes[k]) #Standard score is calculated as (x-mean)/std
p_values[k] = sp.stats.norm.sf(abs(z_scores))*2
if normalisation is None:
continue
elif normalisation=="abs":
allCells_sta[k] = np.nan_to_num(cell_sta/np.max(np.abs(cell_sta)))
elif normalisation=="L2":
allCells_sta[k] = np.nan_to_num(cell_sta/np.sqrt(np.sum(np.power(cell_sta, 2))))
if return_pval:
return allCells_sta, p_values
else:
return allCells_sta
#export
def cross_correlation(spike_counts, tail_len=100):
"""
From calculate the cross correlation of the cells over a time window.
params:
- spike_counts of shape (t, n_cell)
- tail_len: time correlation window size
return:
- cross correlation between the cells
"""
n_dpoints, n_cell = spike_counts.shape
corr_arr = np.zeros((n_cell,n_cell,tail_len+1))
spike_counts = (spike_counts / np.max(spike_counts, axis=0)) #Independant normalization of the cells
spike_counts_edged = np.concatenate((np.zeros((tail_len,n_cell)),
spike_counts,
np.zeros((tail_len,n_cell)))) #Creating an array with 0 tails on both side to use the valid mode
# of numpy.correlate
for i in range(n_cell):
for j in range(i, n_cell):
corr_arr[i,j] = np.correlate(spike_counts_edged[:,i],
spike_counts[:,j],
mode="valid")
corr_arr[j,i] = corr_arr[i,j]
return corr_arr/n_dpoints
#export
def corrcoef(spike_counts):
"""
Computes correlation coefficient between the cells
params:
- spike_counts: Cells activity of shape (t, n_cell)
return:
- Correlation matrix
"""
return np.corrcoef(spike_counts.T)
def flatten_corrcoef(corrcoef_matrix):
"""
Retrieves a one dimensional array of the cells correlations
params:
- corrcoef_matrix: Correlation matrix from `corrcoef`
return:
- flattened correlation matrix"""
shp = corrcoef_matrix.shape
return np.array([corrcoef_matrix[i,j] for i in range(shp[0]) for j in range(i+1, shp[0])])
#export
def stimulus_ensemble(stim_inten, Hw=30, x=0, y=0, w=None, h=None):
"""
Generate the stimulus ensemble used to compute the nonlinearity
params:
- stim_inten: stimulus intensity matrix of shape (t, ...)
- Hw: Lenght in frames of the history window, including the 0 timepoint
- x: Left position of the window where to get the ensemble from
- y: Up position of the window where to get the ensemble from
- w: Width of the window where to get the ensemble from. If None, is set to stim_inten.shape[2]
- h: Height of the window where to get the ensemble from. If None, is set to stim_inten.shape[1]
return:
- Flatten stimulus ensemble of size (len(stim_inten)-(Hw-1), w*h*Hw). To obtain the corresponding cell activity,
slice it like so: slice(Hw-1, None)
"""
stim_inten = stim_inten_norm(stim_inten)
if len(stim_inten.shape) == 1:
stim_inten = stim_inten[..., np.newaxis, np.newaxis]
elif len(stim_inten.shape) == 2:
stim_inten = stim_inten[..., np.newaxis]
if w is None:
w = stim_inten.shape[2]
if h is None:
h = stim_inten.shape[1]
xmin, xmax = max(0,x-w), min(stim_inten.shape[2], x+w+1)
ymin, ymax = max(0,y-h), min(stim_inten.shape[1], y+h+1)
dtype = stim_inten.dtype
if np.all(np.in1d(stim_inten, [-1,0,1])):
dtype = "int8"
stim_ensmbl = np.zeros((len(stim_inten)-(Hw-1), (xmax-xmin)*(ymax-ymin)*Hw), dtype=dtype)
for i in range(0, len(stim_inten)-(Hw-1)):
flat_stim = np.ndarray.flatten(stim_inten[i:i+Hw,
ymin:ymax,
xmin:xmax])
stim_ensmbl[i] = flat_stim
return stim_ensmbl
def process_nonlinearity(stim_inten, spike_counts, bins, stas, p_norm=2):
"""
Computes the nonlinearity of a single cell. The STA of the cell is in L2 normalization, which
should restrict the histogram values.
params:
- stim_inten: stimulus intensity in shape (t, y, x)
- spike_counts: cells activity in shape (t, n_cell)
- bins: bins in which the transformed stimuli ensembles are set. (usually between -6 and 6)
- stas: The STAs to convolve with stim_inten in shape (n_cell, Hw, ...)
- p_norm: Power for the normalization. https://en.wikipedia.org/wiki/Norm_(mathematics)#p-norm :
1 -> can compare nonlinearites of stimuli with different dimensionality
2 -> Common L2 normalization for STAs
return:
- nonlinearity of the cell.
"""
assert len(stim_inten)==len(spike_counts)
stim_inten = stim_inten_norm(np.array(stim_inten))
nonlins = np.empty((len(stas), len(bins)-1))
compute_with_dotprod = False
try:
stim_ensemble = stimulus_ensemble(stim_inten, Hw=stas.shape[1])
compute_with_dotprod = True
except MemoryError as err:
print("Not enough memory to generate the stimulus ensemble, "+
"computing nonlinearity with cross-correlation (will be slower)")
#In the case of calcium imaging, we have probabilities, not spike counts. Need to make it integers
# The discretisation of the calcium imaging is done here globally (for all cells together)
# If it's not what you want, either do the discretisation inside the loop bellow, or discretise the S_matrix
# before passing it to this function
if np.max(spike_counts)<1:
mask = np.where(spike_counts > 0)
nonzero_min = np.min(spike_counts[mask])
discretized = spike_counts/nonzero_min
spike_counts = ((10*discretized)/(np.max(discretized))).astype(int)
spike_counts = np.array(spike_counts)[stas.shape[1]-1:].astype(int)
for i, (sta, sp_count) in enumerate(zip(stas, spike_counts.T)):
sta /= np.power(np.sum(np.power(np.abs(sta), p_norm)), 1/p_norm) # p-norm
if not sp_count.any(): #No spikes
continue
if compute_with_dotprod:
#This one is faster, but requires the stim_ensemble to fit the computer memory
filtered_stim = stim_ensemble@sta.reshape(-1)
else:
filtered_stim = np.squeeze(sp.signal.correlate(stim_inten, sta, mode="valid"))
filtered_sptrigg = np.repeat(filtered_stim, sp_count)
hist_all = np.histogram(filtered_stim, bins=bins)[0]
hist_trigg = np.histogram(filtered_sptrigg, bins=bins)[0]
nonlin = hist_trigg/hist_all
if np.count_nonzero(~np.isnan(nonlin))<2: #Less than two values in the nonlin, cannot fill the gaps
nonlin = np.nan_to_num(nonlin)
else:
nonlin = np.nan_to_num(fill_nan(nonlin))
nonlins[i] = nonlin
return nonlins
#export
def activity_histogram(spike_counts):
"""
Retrieve an histogram of the individual cells activity.
params:
- spike_counts: cells activity matrix of shape (t, n_cell)
return:
- Cells activity histogram
"""
flat_spikes = spike_counts.reshape(-1)
flat_cell = np.array([[i]*spike_counts.shape[0] for i in range(spike_counts.shape[1])]).reshape(-1)
hist = np.histogram2d(flat_spikes, flat_cell, bins=[100,spike_counts.shape[1]])[0] / spike_counts.shape[0]
return hist
#export
def cross_distances(masks):
"""
Computes cross distances from the center of mass of a list of mask.
params:
- masks: cells mask of shape (n_mask, y, x)
return:
- cross distances matrix
"""
center_mass = np.array([ndimage.measurements.center_of_mass(mask) for mask in masks])
cross_distances = np.zeros((len(masks),len(masks)))
for i in range(len(masks)):
for j in range(i,len(masks)):
cross_distances[i,j] = np.linalg.norm(center_mass[i]-center_mass[j])
cross_distances[j,i] = cross_distances[i,j]
return cross_distances
def cross_distances_sta(fits, sta_shape, f):
"""
Computes cross distances between STAs
params:
- fits: fits of STA to compare
- sta_shape: shape of the STA
- f: function used to compute the stas from the fits parameters
return:
- cross distances matrix
"""
sta_masks = np.array([img_2d_fit(sta_shape, fit, f) for fit in fits])
for i,sta_mask in enumerate(sta_masks):
if abs(np.min(sta_mask)) > np.max(sta_mask):
sta_masks[i] = sta_mask < -.5
else:
sta_masks[i] = sta_mask > .5
return cross_distances(sta_masks)
def paired_distances(masks_1, masks_2):
"""
Compute the distance between two masks
params:
- masks_1: Mask of the first cell
- masks_2: Mask of the second cell
return:
- distance between the two masks
"""
center_mass_1 = np.array([ndimage.measurements.center_of_mass(mask) for mask in masks_1])
center_mass_2 = np.array([ndimage.measurements.center_of_mass(mask) for mask in masks_2])
paired_distances = np.zeros(len(masks_1))
for i, (center_1, center_2) in enumerate(zip(masks_1, masks_2)):
paired_distances[i] = np.linalg.norm(center_1-center_2)
return paired_distances
def paired_distances_sta(sta_fits_1, sta_fits_2, sta_shape, f):
"""
Compute the distance between two STAs.
params:
- sta_fits_1: fits of STA of the first cell
- sta_fits_2: fits of STA of the second cell
- sta_shape: shape of the STA
- f: function used to compute the stas from the fits parameters
return:
- distance between the two STAs
"""
sta_masks_1 = np.array([img_2d_fit(sta_shape, fit, f) for fit in sta_fits_1])
for i,sta_mask in enumerate(sta_masks_1):
if abs(np.min(sta_mask)) > np.max(sta_mask):
sta_masks_1[i] = sta_mask < -.5
else:
sta_masks_1[i] = sta_mask > .5
sta_masks_2 = np.array([img_2d_fit(sta_shape, fit, f) for fit in sta_fits_2])
for i,sta_mask in enumerate(sta_masks_2):
if abs(np.min(sta_mask)) > np.max(sta_mask):
sta_masks_2[i] = sta_mask < -.5
else:
sta_masks_2[i] = sta_mask > .5
return paired_distances(sta_masks_1, sta_masks_2)
```
# Direction and orientation selectivity with imaginary numbers
We have 8 angles
```
n_angle = 8
x = np.linspace(0, (n_angle-1)/4*np.pi, num=n_angle)
print(x)
```
Exponentiating those and multiplying with 0+1i will wind up the angles around the origin in imag number space
```
vectors = np.exp(x*1j)
print(vectors)
```
And so multiplying by two before exponentiation double the angle of the arrow. It gives to originally opposite vectors the same direction -> good to calculate orientation selectivity
```
import matplotlib.pyplot as plt
colors = plt.rcParams["axes.prop_cycle"].by_key()['color']
fig, axs = plt.subplots(1,2)
vectors = np.exp(x*1j)
for dx,dy, c in zip (vectors.real, vectors.imag, colors):
axs[0].arrow(0, 0, dx, dy, head_width=0.2, head_length=0.1,length_includes_head=True, color=c, width=.04)
axs[0].set_xlim(-1.5,1.5)
axs[0].set_ylim(-1.5,1.5)
axs[0].set_title("DS vectors")
vectors = np.exp(x*1j*2)
for i,(dx,dy,c) in enumerate(zip(vectors.real, vectors.imag, colors)):
if i>=4:
dx = dx*0.8
dy = dy*0.8
axs[1].arrow(0, 0, dx, dy, head_width=0.2, head_length=0.1, length_includes_head=True, color=c, width=.04)
axs[1].set_xlim(-1.5,1.5)
axs[1].set_ylim(-1.5,1.5)
axs[1].set_title("OS vectors")
#export
def direction_selectivity(grouped_spikes_d, n_bootstrap=1000):
"""
Compute the direction selectivity index of cells in the given dict containing for each condition as
the keys, an array of shape (n_angle, n_repeat, trial_len, n_cell). Such dictionnary can be obtained
by using `utils.group_direction_response`.
params:
- grouped_spikes_d: Results of the group_direction_response of shape (n_angle, n_repeat, t, n_cell)
- n_bootstrap: Number of bootstrap iteration to calculate the p-value
return:
- A dictionnary with a key for each condition retrieving a tuple containing list of the cells
(spike sum, direction pref, DS idx, orientation pref, OS idx, pval DS, pval OS)
"""
res_d = {}
for cond, sp_count in grouped_spikes_d.items():
n_angle = sp_count.shape[0]
sum_rep_spike = np.sum(sp_count, axis=(1,2)).T
x = np.linspace(0, (n_angle-1)/4*np.pi, num=n_angle)
#Direction selectivity
vect_dir = np.exp(x*1j)#np.array([np.cos(x) + np.sin(x)*1j])
dir_pref = np.nan_to_num((vect_dir * sum_rep_spike).sum(axis=1) / sum_rep_spike.sum(axis=1))
ds_idx = abs(dir_pref)
#Orientation selectivity
vect_ori = np.exp(x*1j*2)#np.concatenate((vect_dir[:,:n_angle//2], vect_dir[:,:n_angle//2]), axis=1)
ori_pref = np.nan_to_num((vect_ori * sum_rep_spike).sum(axis=1) / sum_rep_spike.sum(axis=1))
ori_idx = abs(ori_pref)
#Generating direction and orientation index from shuffled trials
axtup_l = list(itertools.product(range(sp_count.shape[0]), range(sp_count.shape[1])))
random.seed(1)
axtup_l_shuffled = axtup_l.copy()
rand_ori_idx_l = np.empty((n_bootstrap, sp_count.shape[3]))
rand_dir_idx_l = np.empty((n_bootstrap, sp_count.shape[3]))
for i in range(n_bootstrap):
random.shuffle(axtup_l_shuffled)
shuffled_sp_count = np.empty(sp_count.shape)
for axtup, axtup_shuff in zip(axtup_l, axtup_l_shuffled):
shuffled_sp_count[axtup] = sp_count[axtup_shuff]
rand_sum_rep_spike = np.sum(shuffled_sp_count, axis=(1,2)).T
rand_dir_pref = np.nan_to_num((vect_dir * rand_sum_rep_spike).sum(axis=1) / rand_sum_rep_spike.sum(axis=1))
rand_dir_idx_l[i] = abs(rand_dir_pref)
rand_ori_pref = np.nan_to_num((vect_ori * rand_sum_rep_spike).sum(axis=1) / rand_sum_rep_spike.sum(axis=1))
rand_ori_idx_l[i] = abs(rand_ori_pref)
#Same calculation of pval as in Baden et al 2016
p_val_dir = 1 - (np.sum(rand_dir_idx_l<ds_idx, axis=0)/n_bootstrap)
p_val_ori = 1 - (np.sum(rand_ori_idx_l<ori_idx, axis=0)/n_bootstrap)
#Finally we have to transform the orientation selectivity vectors to put them back in their
# original orientation, by divinding the phase of the vector by two
tau = np.pi*2
polar_ori_pref = np.array(list((map(polar, ori_pref))))
polar_ori_pref[:,1] = ((polar_ori_pref[:,1]+tau)%tau)/2 #Convert to positive radian angle and divide by two
ori_pref = np.array([rect(pol[0], pol[1]) for pol in polar_ori_pref])
res_d[cond] = (sum_rep_spike, dir_pref, ds_idx, ori_pref, ori_idx, p_val_dir, p_val_ori)
return res_d
#export
def wave_direction_selectivity(wave_array, spike_counts, moving_distance_th=1, looming_distance_th=.3, n_bootstrap=1000):
"""
Computes the direction, orientation and looming/shrinking indexes of the cells in response to the wave stimulus (in LED dome).
params:
- wave_array: The indexes of the waves from the record master.
- spike_counts: The cells response to the waves from the record master.
- moving_distance_th: Distance threshold in radians above which a stimulus can be considered for the OS and DS.
- looming_distance_th: Distance threshold in radians bellow which a stimulus can be considered for the looming/shrinking response
- n_bootstrap: Number of repetition in the bootstraping method to compute the pvalues
return:
- A tuple containing lists of the cells response, in order:
* [0] summed_responses: Summed response of a cell to each wave condition
* [1] dir_pref_l: Direction preference vector
* [2] dir_idx_l : Direction indexes
* [3] dir_pval_l: Direction p_values
* [4] ori_pref_l: Orientation preference vector
* [5] ori_idx_l : Orientation indexes
* [6] ori_pval_l: Orientation p_values
* [7] loom_idx_l : Looming indexes
* [8] loom_pval_l: Looming p_values
* [9] stas_position_l: Position (theta, phi) of the cells receptive fields with the water stimulus
* [10] waves_position_l: Relative positions of the waves to the cells STA.
"""
tau = np.pi*2
indexes, order = np.unique(wave_array, return_index=True)
epoch_sequence = indexes[1:][np.argsort(order[1:])]
wave_inten = build_wave_stimulus_array(epoch_sequence)
stas_wave = process_sta_batch(wave_inten, spike_counts, Hw=1, Fw=0, return_pval=False)
#Hw of 1 because there is a high temporal correlation in the stimulus, so that's enough to find the RF
summed_responses = np.zeros((100, spike_counts.shape[1]))
for i in indexes[1:]: #Iterate from 0 to n_wave-1
where = np.where(wave_array==i)[0]
summed_responses[i,:] = np.sum(spike_counts[where,:], axis=0)
summed_responses = summed_responses.T
dome_positions = get_dome_positions(mode="spherical")
ori_pref_l, dir_pref_l = [], []
ori_idx_l, dir_idx_l, loom_idx_l = [], [], []
ori_pval_l, dir_pval_l, loom_pval_l = [], [], []
stas_position_l, waves_position_l = [], []
for sta, cell_responses in zip(stas_wave, summed_responses):
maxidx_sta = np.argmax(np.abs(sta))
theta_led = dome_positions[maxidx_sta//237,maxidx_sta%237,1]
phi_led = dome_positions[maxidx_sta//237,maxidx_sta%237,2]
relative_waves = get_waves_relative_position((theta_led, phi_led), mode="spherical")
stas_position_l.append((theta_led, phi_led))
waves_position_l.append(relative_waves)
waves_distance = relative_waves[:,1]
waves_angle = (relative_waves[:,2]+tau)%(tau) #Set the angle in the (0,2pi) range
looming_mask = (waves_distance<looming_distance_th)
shrink_mask = (waves_distance>np.pi-looming_distance_th)
waves_mask = (waves_distance>moving_distance_th) & (waves_distance<np.pi-moving_distance_th)
vectors_dir = np.exp(waves_angle*1j) #Create vectors using imaginary numbers
vectors_ori = np.exp(waves_angle*1j*2)#x2 gather the vectors with opposite directions
dir_pref = np.nan_to_num((vectors_dir[waves_mask] * cell_responses[waves_mask]).sum() / cell_responses[waves_mask].sum())
ori_pref = np.nan_to_num((vectors_ori[waves_mask] * cell_responses[waves_mask]).sum() / cell_responses[waves_mask].sum())
ds_idx = abs(dir_pref)
os_idx = abs(ori_pref)
looming_response = (cell_responses[looming_mask]).sum()
shrinking_response = (cell_responses[shrink_mask]).sum()
looming_idx = (looming_response-shrinking_response)/(looming_response+shrinking_response)
ori_pref_l.append(ori_pref); dir_pref_l.append(dir_pref)
ori_idx_l.append(os_idx); dir_idx_l.append(ds_idx)
loom_idx_l.append(looming_idx)
np.random.seed(1)
rand_ori_idx_l = np.empty(n_bootstrap)
rand_dir_idx_l = np.empty(n_bootstrap)
rand_loom_idx_l = np.empty(n_bootstrap)
for i in range(n_bootstrap):
shuffled_response = cell_responses.copy()
np.random.shuffle(shuffled_response)
rand_dir_pref = np.nan_to_num((vectors_dir[waves_mask] * shuffled_response[waves_mask]).sum() / shuffled_response[waves_mask].sum())
rand_ori_pref = np.nan_to_num((vectors_ori[waves_mask] * shuffled_response[waves_mask]).sum() / shuffled_response[waves_mask].sum())
rand_dir_idx_l[i] = abs(rand_dir_pref)
rand_ori_idx_l[i] = abs(rand_ori_pref)
rand_looming_response = (shuffled_response[looming_mask]).sum()
rand_shrinking_response = (shuffled_response[shrink_mask]).sum()
rand_loom_idx_l[i] = (rand_looming_response-rand_shrinking_response)/(rand_looming_response+rand_shrinking_response)
#Same calculation of pval as in Baden et al 2016
p_val_dir = 1 - (np.sum(rand_dir_idx_l<ds_idx)/n_bootstrap)
p_val_ori = 1 - (np.sum(rand_ori_idx_l<os_idx)/n_bootstrap)
p_val_loom = 1 - (np.sum(np.abs(rand_loom_idx_l)<abs(looming_idx))/n_bootstrap)
ori_pval_l.append(p_val_ori); dir_pval_l.append(p_val_dir); loom_pval_l.append(p_val_loom)
# original orientation, by divinding the phase of the vector by two
polar_ori_pref = polar(ori_pref)
new_vector = ((polar_ori_pref[1]+tau)%tau)/2 #Convert to positive radian angle and divide by two
ori_pref = rect(polar_ori_pref[0], new_vector)
return (summed_responses,
dir_pref_l, dir_idx_l, dir_pval_l,
ori_pref_l, ori_idx_l, ori_pval_l,
loom_idx_l, loom_pval_l,
stas_position_l, waves_position_l)
#export
def peri_saccadic_response(spike_counts, eye_track, motion_threshold=5, window=15):
"""
Computes the cell average response around saccades.
params:
- spike_counts: cells activity matrix of shape (t, n_cell)
- eye_track: Eye tracking data of shape (t, x_pos, y_pos, ...)
- motion_threshold: Amount of motion in pixel to account for a saccade
- window: Size of the window before and after the saccade on which to average the cell response
return:
- peri saccadic response of cells of shape (n_cell, window*2+1)
"""
eye_shifts = np.concatenate(([0],
np.linalg.norm(eye_tracking[1:,:2]-eye_tracking[:-1,:2], axis=1)))
#Because eye tracking is usually upsampled from 15 to 60Hz, it sums the shift, and smooth the peak
# detection
summed_shifts = np.convolve(eye_shifts, [1,1,1,1,1,1,1], mode="same")
peaks, res = signal.find_peaks(eye_shifts, height=motion_threshold, distance=10)
heights = res["peak_heights"] #Not used for now
psr = np.zeros((window*2, spike_bins.shape[1]))
for peak in peaks:
if peak<window or (peak+window)>len(spike_bins):
continue #Just ignoring peaks too close to the matrix edges
psr += spike_bins[peak-window:peak+window]
psr /= len(peaks)
return psr
#hide
from nbdev.export import *
notebook2script()
```
| github_jupyter |
# Model Priors
In this notebook I will compare the different model priors that are imposed on inferred physical properties: ${\rm SSFR}$ and $Z_{\rm MW}$. The models we will compare are:
- $\tau$ model
- delayed $\tau$ model
- Rita's 4 component NMF SFH basis
- updated TNG 4 component NMF SFH basis
- updated TNG 4 component NMF SFH basis with burst
```
import numpy as np
from provabgs import infer as Infer
from provabgs import models as Models
from astropy.cosmology import Planck13
# --- plotting ---
import corner as DFM
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
np.random.seed(0)
def prior_tau(tage):
'''model priors for tau and delayed-tau models depend on the age.
We set the fraction of mass from constant SFH to 0 and from bursts
to 0 (i.e. I use the simple dust model)
'''
prior = Infer.load_priors([
Infer.UniformPrior(0., 0.),
Infer.UniformPrior(0.3, 1e1), # tau SFH
Infer.UniformPrior(0., 0.), # constant SFH
Infer.UniformPrior(0., tage-1.), # start time
Infer.UniformPrior(0., 0.), # fburst
Infer.UniformPrior(1e-2, tage), # tburst
Infer.UniformPrior(4.5e-5, 4.5e-2), # metallicity
Infer.UniformPrior(0., 4.)
])
return prior
def prior_nmf(tage):
return Infer.load_priors([
Infer.UniformPrior(0, 0, label='sed'), # uniform priors on logM*
Infer.FlatDirichletPrior(4, label='sed'), # flat dirichilet priors
Infer.LogUniformPrior(1e-5, 1.5e-2, label='sed'),# uniform priors on ZH coeff
Infer.LogUniformPrior(1e-5, 1.5e-2, label='sed'),# uniform priors on ZH coeff
#Infer.LogUniformPrior(4.5e-5, 4.5e-2, label='sed'),# uniform priors on ZH coeffb
#Infer.LogUniformPrior(4.5e-5, 4.5e-2, label='sed'),# uniform priors on ZH coeff
Infer.UniformPrior(0., 3., label='sed'), # uniform priors on dust1
Infer.UniformPrior(0., 3., label='sed'), # uniform priors on dust2
Infer.UniformPrior(-2.2, 0.4, label='sed') # uniform priors on dust_index
])
def prior_nmf_burst(tage):
return Infer.load_priors([
Infer.UniformPrior(0, 0, label='sed'), # uniform priors on logM*
Infer.FlatDirichletPrior(4, label='sed'), # flat dirichilet priors
Infer.UniformPrior(0., 1.), # fburst
Infer.UniformPrior(1e-2, tage), # tburst
Infer.LogUniformPrior(1e-5, 1.5e-2, label='sed'),# uniform priors on ZH coeff
Infer.LogUniformPrior(1e-5, 1.5e-2, label='sed'),# uniform priors on ZH coeff
#Infer.LogUniformPrior(4.5e-5, 4.5e-2, label='sed'),# uniform priors on ZH coeffb
#Infer.LogUniformPrior(4.5e-5, 4.5e-2, label='sed'),# uniform priors on ZH coeff
Infer.UniformPrior(0., 3., label='sed'), # uniform priors on dust1
Infer.UniformPrior(0., 3., label='sed'), # uniform priors on dust2
Infer.UniformPrior(-2.2, 0.4, label='sed') # uniform priors on dust_index
])
models = [
Models.Tau(), # tau model
Models.Tau(delayed=True), # delayed tau
Models.NMF(burst=False), # 4 basis SFH
Models.NMF(burst=True)
]
```
# Comparison at a single redshift $z=0.01$
```
zred = 0.01
tage = Planck13.age(zred).value
priors = [
prior_tau(tage),
prior_tau(tage),
prior_nmf(tage),
prior_nmf_burst(tage)
]
n_sample = 10000 # draw n_sample samples from the prior
SSFRs_1gyr, Zmws, ts_sfh, sfhs = [], [], [], []
for i, model, prior in zip(range(len(models)), models, priors):
_thetas = np.array([prior.sample() for i in range(n_sample)])
thetas = prior.transform(_thetas)
_ssfr_1gyr = model.avgSFR(thetas, zred=zred, dt=1.)
_zmw = model.Z_MW(thetas, zred=zred)
SSFRs_1gyr.append(_ssfr_1gyr)
Zmws.append(_zmw)
print([_ssfr.max() for _ssfr in SSFRs_1gyr])
print([_zmw.max() for _ssfr in SSFRs_1gyr])
model_labels = [r'$\tau$', r'delayed $\tau$', 'NMF 4 comp', 'NMF w/ burst']
props = [SSFRs_1gyr, Zmws]
lbls = [r'$\log {\rm SSFR}_{\rm 1 Gyr}$', r'$\log Z_{\rm MW}$']
rngs = [(-14, -7), (-5, -1)]
fig = plt.figure(figsize=(12,5))
for i, prop in enumerate(props):
sub = fig.add_subplot(1,2,i+1)
for ii, prop_i in enumerate(prop):
sub.hist(np.log10(prop_i), density=True, range=rngs[i], bins=100, alpha=0.5, label=model_labels[ii])
if i == 0:
sub.legend(loc='upper left', handletextpad=0, fontsize=20)
sub.axvline(-9., color='k', ls='--')
elif i == 1:
sub.axvline(np.log10(4.5e-2), color='k', ls='--')
sub.set_xlabel(lbls[i], fontsize=25)
sub.set_xlim(rngs[i])
fig = None
for i in range(len(models)):
fig = DFM.corner(np.log10(np.array(props)[:,i,:].T),
levels=[0.68, 0.95],
smooth=False,
color='C%i' % i,
range=rngs,
labels=lbls,
label_kwargs={'fontsize': 20},
fig=fig,
plot_datapoints=False, fill_contours=False, plot_density=False
)
```
Overall the NMF and compressed NMF models impose more restrictive priors on $\log {\rm SSFR}$ and $\log Z_{\rm MW}$
# redshift dependence of model priors
```
prior_fns = [prior_tau, prior_tau, prior_nmf, prior_nmf_burst]
for model, prior_fn, lbl in zip(models, prior_fns, model_labels):
fig = None
for i_z, zred in enumerate([0.01, 0.05, 0.1, 0.2, 0.4]):
tage = Planck13.age(zred).value
prior = prior_fn(tage)
_thetas = np.array([prior.sample() for i in range(n_sample)])
thetas = prior.transform(_thetas)
_ssfr_1gyr = model.avgSFR(thetas, zred=zred, dt=1.)
_zmw = model.Z_MW(thetas, zred=zred)
fig = DFM.corner(np.log10(np.array([_ssfr_1gyr, _zmw])).T,
levels=[0.68, 0.95],
smooth=True,
range=rngs,
labels=lbls,
label_kwargs={'fontsize': 20},
color='C%i' % i_z,
fig=fig,
plot_datapoints=False, fill_contours=False, plot_density=False
)
fig.suptitle(lbl+' model', fontsize=25)
```
None of the models have significant redshift dependence.
## prior on recovered SFH
```
zred = 0.01
tage = Planck13.age(zred).value
recov_sfhs = []
for model, prior_fn in zip(models, prior_fns):
prior = prior_fn(tage)
_thetas = np.array([prior.sample() for i in range(n_sample)])
thetas = prior.transform(_thetas)
recov_sfh = [model.avgSFR(thetas, zred=zred, dt=1.)]
for tlb in np.arange(1, 11):
recov_sfh.append(model.avgSFR(thetas, zred=zred, dt=1., t0=tlb))
recov_sfhs.append(recov_sfh)
fig = plt.figure(figsize=(20,5))
for i, recov_sfh in enumerate(recov_sfhs):
sub = fig.add_subplot(1, len(recov_sfhs), i+1)
for tlb, rsfh in zip(np.arange(11) + 0.5, recov_sfh):
lll, ll, l, m, h, hh, hhh = np.quantile(rsfh, [0.005, 0.025, 0.16, 0.5, 0.84, 0.975, 0.995])
sub.fill_between([tlb-0.5, tlb+0.5], [lll, lll], [hhh, hhh], color='C0', alpha=0.1, linewidth=0)
sub.fill_between([tlb-0.5, tlb+0.5], [ll, ll], [hh, hh], color='C0', alpha=0.25, linewidth=0)
sub.fill_between([tlb-0.5, tlb+0.5], [l, l], [h, h], color='C0', alpha=0.5, linewidth=0)
sub.plot([tlb-0.5, tlb+0.5], [m, m], c='C0', ls='-')
if i == 0: sub.set_ylabel('SSFR', fontsize=25)
else: sub.set_yticklabels([])
sub.set_yscale('log')
sub.set_ylim(1e-12, 1e-9)
if i == 2: sub.set_xlabel(r'$t_{\rm lookback}$', fontsize=25)
sub.set_xlim(0, tage)
sub.set_title(model_labels[i], fontsize=20)
```
| github_jupyter |
# Exp 123 analysis
See `./informercial/Makefile` for experimental
details.
```
import os
import numpy as np
from IPython.display import Image
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import seaborn as sns
sns.set_style('ticks')
matplotlib.rcParams.update({'font.size': 16})
matplotlib.rc('axes', titlesize=16)
from infomercial.exp import meta_bandit
from infomercial.exp import epsilon_bandit
from infomercial.exp import beta_bandit
from infomercial.exp import softbeta_bandit
from infomercial.local_gym import bandit
from infomercial.exp.meta_bandit import load_checkpoint
import gym
def plot_meta(env_name, result):
"""Plots!"""
# episodes, actions, scores_E, scores_R, values_E, values_R, ties, policies
episodes = result["episodes"]
actions =result["actions"]
bests =result["p_bests"]
scores_E = result["scores_E"]
scores_R = result["scores_R"]
values_R = result["values_R"]
values_E = result["values_E"]
ties = result["ties"]
policies = result["policies"]
# -
env = gym.make(env_name)
best = env.best
print(f"Best arm: {best}, last arm: {actions[-1]}")
# Plotz
fig = plt.figure(figsize=(6, 14))
grid = plt.GridSpec(6, 1, wspace=0.3, hspace=0.8)
# Arm
plt.subplot(grid[0, 0])
plt.scatter(episodes, actions, color="black", alpha=.5, s=2, label="Bandit")
plt.plot(episodes, np.repeat(best, np.max(episodes)+1),
color="red", alpha=0.8, ls='--', linewidth=2)
plt.ylim(-.1, np.max(actions)+1.1)
plt.ylabel("Arm choice")
plt.xlabel("Episode")
# Policy
policies = np.asarray(policies)
episodes = np.asarray(episodes)
plt.subplot(grid[1, 0])
m = policies == 0
plt.scatter(episodes[m], policies[m], alpha=.4, s=2, label="$\pi_E$", color="purple")
m = policies == 1
plt.scatter(episodes[m], policies[m], alpha=.4, s=2, label="$\pi_R$", color="grey")
plt.ylim(-.1, 1+.1)
plt.ylabel("Controlling\npolicy")
plt.xlabel("Episode")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
_ = sns.despine()
# score
plt.subplot(grid[2, 0])
plt.scatter(episodes, scores_E, color="purple", alpha=0.4, s=2, label="E")
plt.plot(episodes, scores_E, color="purple", alpha=0.4)
plt.scatter(episodes, scores_R, color="grey", alpha=0.4, s=2, label="R")
plt.plot(episodes, scores_R, color="grey", alpha=0.4)
plt.plot(episodes, np.repeat(tie_threshold, np.max(episodes)+1),
color="violet", alpha=0.8, ls='--', linewidth=2)
plt.ylabel("Score")
plt.xlabel("Episode")
# plt.semilogy()
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
_ = sns.despine()
# Q
plt.subplot(grid[3, 0])
plt.scatter(episodes, values_E, color="purple", alpha=0.4, s=2, label="$Q_E$")
plt.scatter(episodes, values_R, color="grey", alpha=0.4, s=2, label="$Q_R$")
plt.plot(episodes, np.repeat(tie_threshold, np.max(episodes)+1),
color="violet", alpha=0.8, ls='--', linewidth=2)
plt.ylabel("Value")
plt.xlabel("Episode")
# plt.semilogy()
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
_ = sns.despine()
# Ties
plt.subplot(grid[4, 0])
plt.scatter(episodes, bests, color="red", alpha=.5, s=2)
plt.ylabel("p(best)")
plt.xlabel("Episode")
plt.ylim(0, 1)
# Ties
plt.subplot(grid[5, 0])
plt.scatter(episodes, ties, color="black", alpha=.5, s=2, label="$\pi_{tie}$ : 1\n $\pi_\pi$ : 0")
plt.ylim(-.1, 1+.1)
plt.ylabel("Ties index")
plt.xlabel("Episode")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
def plot_epsilon(env_name, result):
"""Plots!"""
# episodes, actions, scores_E, scores_R, values_E, values_R, ties, policies
episodes = result["episodes"]
actions =result["actions"]
bests =result["p_bests"]
scores_R = result["scores_R"]
values_R = result["values_R"]
epsilons = result["epsilons"]
# -
env = gym.make(env_name)
best = env.best
print(f"Best arm: {best}, last arm: {actions[-1]}")
# Plotz
fig = plt.figure(figsize=(6, 14))
grid = plt.GridSpec(6, 1, wspace=0.3, hspace=0.8)
# Arm
plt.subplot(grid[0, 0])
plt.scatter(episodes, actions, color="black", alpha=.5, s=2, label="Bandit")
plt.plot(episodes, np.repeat(best, np.max(episodes)+1),
color="red", alpha=0.8, ls='--', linewidth=2)
plt.ylim(-.1, np.max(actions)+1.1)
plt.ylabel("Arm choice")
plt.xlabel("Episode")
# score
plt.subplot(grid[1, 0])
plt.scatter(episodes, scores_R, color="grey", alpha=0.4, s=2, label="R")
plt.ylabel("Score")
plt.xlabel("Episode")
# plt.semilogy()
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
_ = sns.despine()
# Q
plt.subplot(grid[2, 0])
plt.scatter(episodes, values_R, color="grey", alpha=0.4, s=2, label="$Q_R$")
plt.ylabel("Value")
plt.xlabel("Episode")
# plt.semilogy()
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
_ = sns.despine()
# best
plt.subplot(grid[3, 0])
plt.scatter(episodes, bests, color="red", alpha=.5, s=2)
plt.ylabel("p(best)")
plt.xlabel("Episode")
plt.ylim(0, 1)
# Decay
plt.subplot(grid[4, 0])
plt.scatter(episodes, epsilons, color="black", alpha=.5, s=2)
plt.ylabel("$\epsilon_R$")
plt.xlabel("Episode")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
def plot_beta(env_name, result):
"""Plots!"""
# episodes, actions, scores_E, scores_R, values_E, values_R, ties, policies
episodes = result["episodes"]
actions =result["actions"]
bests =result["p_bests"]
scores_R = result["scores_R"]
values_R = result["values_R"]
beta = result["beta"]
# -
env = gym.make(env_name)
best = env.best
print(f"Best arm: {best}, last arm: {actions[-1]}")
# Plotz
fig = plt.figure(figsize=(6, 14))
grid = plt.GridSpec(6, 1, wspace=0.3, hspace=0.8)
# Arm
plt.subplot(grid[0, 0])
plt.scatter(episodes, actions, color="black", alpha=.5, s=2, label="Bandit")
plt.plot(episodes, np.repeat(best, np.max(episodes)+1),
color="red", alpha=0.8, ls='--', linewidth=2)
plt.ylim(-.1, np.max(actions)+1.1)
plt.ylabel("Arm choice")
plt.xlabel("Episode")
# score
plt.subplot(grid[1, 0])
plt.scatter(episodes, scores_R, color="grey", alpha=0.4, s=2, label="R")
plt.ylabel("Score")
plt.xlabel("Episode")
# plt.semilogy()
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
_ = sns.despine()
# Q
plt.subplot(grid[2, 0])
plt.scatter(episodes, values_R, color="grey", alpha=0.4, s=2, label="$Q_R$")
plt.ylabel("Value")
plt.xlabel("Episode")
# plt.semilogy()
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
_ = sns.despine()
# best
plt.subplot(grid[3, 0])
plt.scatter(episodes, bests, color="red", alpha=.5, s=2)
plt.ylabel("p(best)")
plt.xlabel("Episode")
plt.ylim(0, 1)
def plot_critic(critic_name, env_name, result):
# -
env = gym.make(env_name)
best = env.best
# Data
critic = result[critic_name]
arms = list(critic.keys())
values = list(critic.values())
# Plotz
fig = plt.figure(figsize=(8, 3))
grid = plt.GridSpec(1, 1, wspace=0.3, hspace=0.8)
# Arm
plt.subplot(grid[0])
plt.scatter(arms, values, color="black", alpha=.5, s=30)
plt.plot([best]*10, np.linspace(min(values), max(values), 10), color="red", alpha=0.8, ls='--', linewidth=2)
plt.ylabel("Value")
plt.xlabel("Arm")
```
# Load and process data
```
data_path ="/Users/qualia/Code/infomercial/data/"
exp_name = "exp123"
sorted_params = load_checkpoint(os.path.join(data_path, f"{exp_name}_sorted.pkl"))
# print(sorted_params.keys())
best_params = sorted_params[0]
beta = best_params['beta']
sorted_params
```
# Performance
of best parameters
```
env_name = 'BanditHardAndSparse10-v0'
num_episodes = 50000*10
# Run w/ best params
result = beta_bandit(
env_name=env_name,
num_episodes=num_episodes,
lr_R=best_params["lr_R"],
beta=best_params["beta"],
seed_value=2,
)
print(best_params)
plot_beta(env_name, result=result)
plot_critic('critic', env_name, result)
```
# Sensitivity
to parameter choices
```
total_Rs = []
betas = []
lrs_R = []
lrs_E = []
trials = list(sorted_params.keys())
for t in trials:
total_Rs.append(sorted_params[t]['total_R'])
lrs_R.append(sorted_params[t]['lr_R'])
betas.append(sorted_params[t]['beta'])
# Init plot
fig = plt.figure(figsize=(5, 18))
grid = plt.GridSpec(6, 1, wspace=0.3, hspace=0.8)
# Do plots:
# Arm
plt.subplot(grid[0, 0])
plt.scatter(trials, total_Rs, color="black", alpha=.5, s=6, label="total R")
plt.xlabel("Sorted params")
plt.ylabel("total R")
_ = sns.despine()
plt.subplot(grid[1, 0])
plt.scatter(trials, lrs_R, color="black", alpha=.5, s=6, label="total R")
plt.xlabel("Sorted params")
plt.ylabel("lr_R")
_ = sns.despine()
plt.subplot(grid[2, 0])
plt.scatter(lrs_R, total_Rs, color="black", alpha=.5, s=6, label="total R")
plt.xlabel("lrs_R")
plt.ylabel("total_Rs")
_ = sns.despine()
plt.subplot(grid[3, 0])
plt.scatter(betas, total_Rs, color="black", alpha=.5, s=6, label="total R")
plt.xlabel("beta")
plt.ylabel("total_Rs")
_ = sns.despine()
```
# Parameter correlations
```
from scipy.stats import spearmanr
spearmanr(total_Rs, lrs_R)
spearmanr(betas, total_Rs)
spearmanr(betas, lrs_R)
```
# Distributions
of parameters
```
# Init plot
fig = plt.figure(figsize=(5, 6))
grid = plt.GridSpec(3, 1, wspace=0.3, hspace=0.8)
plt.subplot(grid[0, 0])
plt.hist(betas, color="black")
plt.xlabel("beta")
plt.ylabel("Count")
_ = sns.despine()
plt.subplot(grid[1, 0])
plt.hist(lrs_R, color="black")
plt.xlabel("lr_R")
plt.ylabel("Count")
_ = sns.despine()
```
of total reward
```
# Init plot
fig = plt.figure(figsize=(5, 2))
grid = plt.GridSpec(1, 1, wspace=0.3, hspace=0.8)
plt.subplot(grid[0, 0])
plt.hist(total_Rs, color="black", bins=50)
plt.xlabel("Total reward")
plt.ylabel("Count")
# plt.xlim(0, 10)
_ = sns.despine()
```
| github_jupyter |
```
# import datetime
import os
import pandas
import numpy as np
import matplotlib.pyplot as plt
from datlib.FRED import *
from datlib.plots import *
import pandas_datareader.data as web
#FRED.py
#. . .
def bil_to_mil(series):
return series* 10**3
# . . .
#fedProject.py
# . . .
data_codes = {# Assets
"Balance Sheet: Total Assets ($ Mil)": "WALCL",
"Balance Sheet Securities, Prem-Disc, Repos, and Loans ($ Mil)": "WSRLL",
"Balance Sheet: Securities Held Outright ($ Mil)": "WSHOSHO",
### breakdown of securities holdings ###
"Balance Sheet: U.S. Treasuries Held Outright ($ Mil)":"WSHOTSL",
"Balance Sheet: Federal Agency Debt Securities ($ Mil)" : "WSHOFADSL",
"Balance Sheet: Mortgage-Backed Securities ($ Mil)": "WSHOMCB",
# other forms of lending
"Balance Sheet: Repos ($ Mil)": "WORAL",
"Balance Sheet: Central Bank Liquidity Swaps ($ Mil)" : "SWPT",
"Balance Sheet: Direct Lending ($ Mil)" : "WLCFLL",
# unamortized value of securities held (due to changes in interest rates)
"Balance Sheet: Unamortized Security Premiums ($ Mil)": "WUPSHO",
# Liabilities
"Balance Sheet: Total Liabilities ($ Mil)" : "WLTLECL",
"Balance Sheet: Federal Reserve Notes Outstanding ($ Mil)" : "WLFN",
"Balance Sheet: Reverse Repos ($ Mil)": "WLRRAL",
### Major share of deposits
"Balance Sheet: Deposits from Dep. Institutions ($ Mil)":"WLODLL",
"Balance Sheet: U.S. Treasury General Account ($ Mil)": "WDTGAL",
"Balance Sheet: Other Deposits ($ Mil)": "WOTHLB",
"Balance Sheet: All Deposits ($ Mil)": "WLDLCL",
# Capital
"Balance Sheet: Total Capital": "WCTCL",
# Interest Rates
"Unemployment Rate": "UNRATE",
"Nominal GDP ($ Bil)":"GDP",
"Real GDP ($ Bil)":"GDPC1",
"GDP Deflator":"GDPDEF",
"CPI":"CPIAUCSL",
"Core PCE":"PCEPILFE",
"Private Investment":"GPDI",
"Base: Total ($ Mil)": "BOGMBASE",
"Base: Currency in Circulation ($ Bil)": "WCURCIR",
"1 Month Treasury Rate (%)": "DGS1MO",
"3 Month Treasury Rate (%)": "DGS3MO",
"1 Year Treasury Rate (%)": "DGS1",
"2 Year Treasury Rate (%)": "DGS2",
"10 Year Treasury Rate (%)": "DGS10",
"30 Year Treasury Rate (%)": "DGS30",
"Effective Federal Funds Rate (%)": "DFF",
"Federal Funds Target Rate (Pre-crisis)":"DFEDTAR",
"Federal Funds Upper Target":"DFEDTARU",
"Federal Funds Lower Target":"DFEDTARL",
"Interest on Reserves (%)": "IOER",
"VIX": "VIXCLS",
"5 Year Forward Rate": "T5YIFR"
}
inflation_target = .02
unemployment_target = .04
# Select start and end dates
start = datetime.datetime(2001, 12, 1)
end = datetime.datetime.today()
annual_div = {"Q":4,
"W":52,
"M":12}
# freq refers to data frequency. Choose "D", "W", "M", "Q", "A"
# a number may also be place in front of a letter. "2D" indicates
# alternating days
if "data_gathered" not in locals():
freq = "M"
year = annual_div[freq]
data = gather_data(data_codes, start,
end = end, freq = freq)
data.fillna(0, inplace=True)
for key in data.keys():
data["Log " + key]= np.log(data[key])
# Create new variables
data_gathered = True
ticker = "^GSPC"
data["Base: Currency in Circulation ($ Mil)"] = data["Base: Currency in Circulation ($ Bil)"].mul(1000)
data["Base: Currency not in Circulation ($ Mil)"] = data["Base: Total ($ Mil)"].sub(data["Base: Currency in Circulation ($ Mil)"])
data["Currency in Circulation Growth Rate (%)"] = data["Base: Currency in Circulation ($ Mil)"].pct_change(year) * 100
data["% Currency not in Circulation"] = data["Base: Currency not in Circulation ($ Mil)"].div(data["Base: Total ($ Mil)"]) * 100
data["% Currency in Circulation"] = data["Base: Currency in Circulation ($ Mil)"].div(data["Base: Total ($ Mil)"]) * 100
data["Base: Total Growth Rate (%)"] = data["Base: Total ($ Mil)"]
data["Change % Currency not in Circulation"] = data["% Currency not in Circulation"].diff(year)
data["Currency not in Circulation Growth Rate (%)"] = data["Base: Currency not in Circulation ($ Mil)"].pct_change(year) * 100
data["Inflation (CPI)"] = web.DataReader("CPIAUCSL", "fred", start, end).resample(freq).mean().pct_change(year).mul(100).dropna()
data["Inflation (PCE)"] = web.DataReader("PCEPILFE", "fred", start, end).resample(freq).mean().pct_change(year).mul(100).dropna()
data["Effective Federal Funds Rate Diff (%)"] = data["Effective Federal Funds Rate (%)"].diff(year)
data["1 Year Treasury Rate (%; diff)"] = data["1 Year Treasury Rate (%)"].diff(year)
data["2 Year Treasury Rate (%; diff)"] = data["2 Year Treasury Rate (%)"].diff(year)
data["10 Year Treasury Rate (%; diff)"] = data["10 Year Treasury Rate (%)"].diff(year)
data["30 Year Treasury Rate (%; diff)"] = data["30 Year Treasury Rate (%)"].diff(year)
data["Unemployment Rate Diff"] = data["Unemployment Rate"].diff(year)
data["Nominal GDP ($ Mil)"] = data["Nominal GDP ($ Bil)"].mul(1000)
data["Nominal GDP Growth Rate (%)"] = data["Nominal GDP ($ Bil)"].pct_change(year) * 100
data["Real GDP ($ Mil)"] = data["Real GDP ($ Bil)"].mul(1000)
data["Real GDP Growth Rate (%)"] = data["Real GDP ($ Bil)"].pct_change(year) * 100
data["Inflation (GDPDEF)"] = data["GDP Deflator"].pct_change(year) * 100
data["Real Currency in Circulation Growth Rate (%)"] = data["Currency in Circulation Growth Rate (%)"].sub(data["Inflation (GDPDEF)"])
data["Currency in Circulation Velocity"] = data["Nominal GDP ($ Mil)"].div(data["Base: Currency in Circulation ($ Mil)"])
data["Currency in Circulation % Change Velocity"] = data["Currency in Circulation Velocity"].pct_change(year)
data["Inflation Loss"]= data["Inflation (PCE)"].sub(inflation_target)
data["Unemployment Loss"]= data["Unemployment Rate"].sub(unemployment_target)
data["Inflation Loss Sq"]= data["Inflation (PCE)"].sub(inflation_target).pow(2)
data["Inflation Loss Sq"][data["Inflation Loss"] < 0] = data["Inflation Loss Sq"].mul(-1)
data["Unemployment Loss Sq"]= data["Unemployment Rate"].sub(unemployment_target).pow(2)
data["Unemployment Loss Sq"][data["Unemployment Loss"] < 0] = data["Unemployment Loss Sq"].mul(-1)
data["Inflation Loss Diff"]= data["Inflation Loss"].diff(year)
data["Unemployment Loss Diff"]= data["Unemployment Loss"].diff(year)
data["Inflation Loss Sq Diff"]= data["Inflation Loss Sq"].diff(year)
data["Unemployment Loss Sq Diff"]= data["Unemployment Loss Sq"].diff(year)
data["Linear Loss"] = data["Inflation Loss"].sub(data["Unemployment Loss"])
data["Loss Function"] = data["Inflation Loss Sq"].sub(data["Unemployment Loss Sq"])
data["Linear Loss Diff"] = data["Linear Loss"].diff(year)
data["Loss Function Diff"] = data["Loss Function"].diff(year)
data["Real 1 Year Treasury Rate"] = data["1 Year Treasury Rate (%)"].sub(data["Inflation (CPI)"])
data["Real 3 Month Treasury Rate"] = data["3 Month Treasury Rate (%)"].sub(data["Inflation (CPI)"])
data["Real 1 Month Treasury Rate"] = data["1 Month Treasury Rate (%)"].sub(data["Inflation (CPI)"])
data["Real Effective Federal Funds Rate"] = data['Effective Federal Funds Rate (%)'].sub(data["Inflation (CPI)"])
data["30 Year Minus 1 Year (%)"] = data["30 Year Treasury Rate (%)"].sub(data["1 Year Treasury Rate (%)"])
data["30 Year Minus 3 Month (%)"] = data["30 Year Treasury Rate (%)"].sub(data["3 Month Treasury Rate (%)"])
data["30 Year Minus 1 Month (%)"] = data["30 Year Treasury Rate (%)"].sub(data["1 Month Treasury Rate (%)"])
data["30 Year Minus Effective Federal Funds Rate"] = data["30 Year Treasury Rate (%)"].sub(data['Effective Federal Funds Rate (%)'])
data["10 Year Minus 2 Year (%)"] = data["10 Year Treasury Rate (%)"].sub(data["2 Year Treasury Rate (%)"])
data["10 Year Minus 1 Year (%)"] = data["10 Year Treasury Rate (%)"].sub(data["1 Year Treasury Rate (%)"])
data["10 Year Minus 3 Month (%)"] = data["10 Year Treasury Rate (%)"].sub(data["3 Month Treasury Rate (%)"])
data["10 Year Minus 1 Month (%)"] = data["10 Year Treasury Rate (%)"].sub(data["1 Month Treasury Rate (%)"])
data["10 Year Minus Effective Federal Funds Rate"] = data["10 Year Treasury Rate (%)"].sub(data['Effective Federal Funds Rate (%)'])
keys = list(data.keys())
keys = ["Date"] + keys
data["Date"] = data.index.astype(str)
import pandas as pd
sheet_names = pd.ExcelFile('BEAGrossOutputbyindustryQuarterly.xlsx').sheet_names # see all sheet names
info = pd.read_excel("BEAGrossOutputbyindustryQuarterly.xlsx", sheet_name = sheet_names[0], header = [3]).dropna(axis = 1)
key_names = {}
for ix in info.index:
row = info.loc[ix]
key, val = row.values[0], row.values[1]
key_names[key] = val
key_names
sector_data = {}
for sheet_name, variable in key_names.items():
sector_data[variable] = pd.read_excel("BEAGrossOutputbyindustryQuarterly.xlsx",
index_col = [1],
sheet_name=sheet_name+"-Q",
header = [7], parse_dates= True).T.iloc[2:].dropna(axis = 1)
sector_data[variable].rename(index = {ix:ix.replace("Q","-") for ix in sector_data[variable].index},
inplace = True)
sector_data[variable].rename(index = {ix:ix[:-1] + ("0" + str(int(ix[-1]) * 3))[-2:] for ix in sector_data[variable].index},
inplace = True)
sector_data[variable].index = pd.DatetimeIndex(sector_data[variable].index)
sector_data.keys()
indices = ['Chain-Type Quantity Indexes for Gross Output by Industry',
'Chain-Type Price Indexes for Gross Output by Industry',
'Gross Output by Industry',
'Real Gross Output by Industry']
layers = {}
for index in indices:
data = sector_data[index]
layers[index] = {0:[],
1:{"Private industries":[], "Government":[]},
2:{}}
sector = "Private industries"
for key in sector_data[index].keys():
if key == "Government":
sector = "Government"
if key == key.strip():
layers[index][0].append(key)
subsector = key
elif key[:2] == " " and key[:3] != " ":
layers[index][1][layers[index][0][-1]].append(key)
# elif key[:4] == " " and key[:5] != " ":
# layers[index][2][layers[index][0][-1]].append(key)
layers
# elif "All industries" not in key:
# layers[2][sector]
# .append(layers[0][-1] + ": "+ layers[1][-1] + ": " + key)
private_and_public = layers['Chain-Type Quantity Indexes for Gross Output by Industry'][1]["Private industries"] + layers['Chain-Type Quantity Indexes for Gross Output by Industry'][1]["Government"]
private_and_public
df.keys()
df["Date"].astype(str).str[:4].astype(int)
import matplotlib.pyplot as plt
from matplotlib.backends.backend_pdf import PdfPages
import pingouin
plt.rcParams.update({"font.size":24})
vals = private_and_public
for group in layers:
pp = PdfPages(group + "LinePlots.pdf")
df = sector_data[group]
keys = layers[group][1]["Private industries"] + layers[group][1]["Government"]
df = df[keys]
i = 0
for key in keys:
color = "C" + str(i)
color_df = df[[key]]
other_keys = [k for k in keys if k != key]
grey_df = df[other_keys]
fig, ax = plt.subplots(figsize = (30,30))
grey_df.plot.line(ax=ax,
color = "k",
alpha = .3,
linewidth = 5,
legend = False)
for other_key in other_keys:
plt.text(grey_df.index[-1], grey_df[other_key].iloc[-1],
other_key, color = "k", alpha= .3)
color_df.plot.line(ax = ax, color = color, linewidth = 10, legend = False)
plt.text(color_df.index[-1], color_df[key].iloc[-1], key, color = color, fontsize= 30)
i += 1
plt.title(group)
pp.savefig(fig, bbox_inches = "tight")
plt.close()
pp.close()
pp = PdfPages(group + "ScatterPlots.pdf")
df["Date"] = df.index.astype(str).str[:4].astype(int)
for key1 in keys:
for key2 in keys:
if key1 != key2:
fig, ax = plt.subplots(figsize= (20,20))
im = ax.scatter(x = df[key1], y = df[key2], c = df["Date"],
cmap = "viridis", s = 50)
ax.set_xlabel(key1)
ax.set_ylabel(key2)
cbar = plt.colorbar(im)
cbar.ax.set_ylabel('Date')
pp.savefig(fig, bbox_inches = "tight")
plt.title(group)
# plt.show()
plt.close()
pp.close()
corr_keys = list(df.keys())[:-1]
for key in corr_keys:
df[key] = df[key].astype(float)
df.corr().dropna().to_csv(group + "CorrMatrix.csv")
df.pcorr().dropna().to_csv(group + "PcorrMatrix.csv")
# series.plot(ax = ax, logy = False, legend = False, linewidth = 10)
# j = 0
# for key in series.keys():
# color = "C" + str(j)
# plt.text(series.index[-1], series[key].iloc[-1], key, color = color)
# j += 1
# fig, ax = plt.subplots(figsize = (30,30))
# i = 0
# df = sector_data[group]
# print(val)
# # print(val)
# keys = layers[group][1]
# for key, vals in keys.items():
# for val in vals:
# series = df[val]
# j = i % 10
# series.plot(ax = ax, logy = False, legend = False, linewidth = 10)
# plt.text(series.index[-1], series.iloc[-1], series.index.name, color = "C" + str(j))
# i+=1
# plt.title(group)
# plt.show()
# plt.close()
# i = 0
# for key, val in layers[group][1].items():
# for series in val:
# fig, ax = plt.subplots(figsize = (30,30))
# for series2 in val:
# if series2.keys() != series.keys():
# series2.plot(ax = ax, logy = False, legend = False, color = "k", alpha = .3, linewidth = 5)
# plt.text(series2.index[-1], series2.iloc[-1], series2.keys()[0], color = "k", alpha = .4)
# j = i % 10
# color = "C" + str(j)
# series.plot(ax = ax, logy = False, legend = False, color = color, linewidth = 10)
# plt.text(series.index[-1], series.iloc[-1], series.keys()[0], color = color, size = 50)
# i+=1
# plt.title(group)
# plt.show()
# plt.close()
corr_keys = list(df.keys())[:-1]
for key in corr_keys:
df[key] = df[key].astype(float)
df.pcorr()
# import pandas_datareader.data as web
# start = datetime.datetime(2001, 12, 1)
# end = datetime.datetime.today()
# x = web.DataReader("CPIAUCSL", "fred", start, end).resample(freq).mean().pct_change(year).mul(100).dropna()
# x
import yfinance as yfin
yfin.pdr_override()
data["S&P"]= web.get_data_yahoo(ticker, start = start, end = end).resample(freq).mean()["Close"].iloc[:-2]
data["S&P Growth Rate (%)"] = data["S&P"].pct_change(year)
data["S&P Growth Rate Change Diff (%)"] = data["S&P Growth Rate (%)"].diff(year)
data["Real S&P Growth Rate (%)"] = data["S&P Growth Rate (%)"].sub(data["Inflation (CPI)"])
data["VIX Diff"] = data["VIX"].diff(year)
data
#fedProject
# . . .
data["Balance Sheet: Direct Lending and Central Bank Liquidity Swaps"] =\
data["Balance Sheet: Central Bank Liquidity Swaps ($ Mil)"].add(
data["Balance Sheet: Direct Lending ($ Mil)"])
data["Balance Sheet: Other Securities"] = data["Balance Sheet: Securities Held Outright ($ Mil)"].sub(
data["Balance Sheet: U.S. Treasuries Held Outright ($ Mil)"]).sub(
data["Balance Sheet: Mortgage-Backed Securities ($ Mil)"])
data["Balance Sheet: Other Assets"] = data["Balance Sheet: Total Assets ($ Mil)"].sub(
data["Balance Sheet: Securities Held Outright ($ Mil)"]).sub(
data["Balance Sheet: Direct Lending and Central Bank Liquidity Swaps"]).sub(
data["Balance Sheet: Repos ($ Mil)"]).sub(
data["Balance Sheet: Unamortized Security Premiums ($ Mil)"])
data["Balance Sheet: Other Deposits ($ Mil)"] = data["Balance Sheet: All Deposits ($ Mil)"].sub(
data["Balance Sheet: U.S. Treasury General Account ($ Mil)"]).sub(
data["Balance Sheet: Deposits from Dep. Institutions ($ Mil)"])
data["Balance Sheet: Other Liabilities"]= data["Balance Sheet: Total Liabilities ($ Mil)"].sub(
data["Balance Sheet: Federal Reserve Notes Outstanding ($ Mil)"]).sub(
data["Balance Sheet: U.S. Treasury General Account ($ Mil)"]).sub(
data["Balance Sheet: Deposits from Dep. Institutions ($ Mil)"]).sub(
data["Balance Sheet: Other Deposits ($ Mil)"]).sub(
data["Balance Sheet: Reverse Repos ($ Mil)"])
plt.rcParams.update({"font.size":32})
interest_vars = ["Effective Federal Funds Rate (%)",
"Interest on Reserves (%)",
"1 Month Treasury Rate (%)"]
fig, ax = plt.subplots(figsize = (30,20))
data[interest_vars].plot.line(legend=True, linewidth = 3, ax = ax)
y_vals = ax.get_yticks()
ax.set_yticklabels([str(round(y,2))+ "%" for y in y_vals])
ax.set_title("Short Term Interest Rates", fontsize = 48)
data[["Date"] + interest_vars].to_json("shortTermRates.json", orient="records")
data = data.rename(columns = {
key: key.replace("Balance Sheet: ", "").replace(" ($ Mil)", "").replace("Base: ","") for key in data.keys()})
keys = list(data.keys())
interest_rates = ["Effective Federal Funds Rate (%)",
"Interest on Reserves (%)",
"1 Month Treasury Rate (%)"]
plot_data = data.copy()
for key, val in data.items():
if key not in interest_rates:
try:
plot_data[key] = val.div(10**6)
except:
continue
account_vars = ["U.S. Treasuries Held Outright",
"Mortgage-Backed Securities",
"Other Securities",
"Direct Lending and Central Bank Liquidity Swaps",
"Repos",
"Unamortized Security Premiums",
"Other Assets"]
figsize= (36,18)
fig, ax = plt.subplots(figsize = figsize)
plot_data[account_vars].plot.area(stacked = True, linewidth = 3,
ax = ax)
# change y vals from mil to tril
total_var = "Total Assets"
plot_data[total_var].plot.line(linewidth = 1,
ax = ax, c = "k",
label = total_var, ls = "--")
plt.xticks(rotation = 90)
ax.legend(loc=2, ncol = 2)
ax.set_ylabel("$ Trillion", fontsize = 40)
ax.set_title("Federal Reserve Balance Sheet: Assets", fontsize = 50)
plot_data[[total_var] + account_vars].to_csv("FederalReserveAssets.csv")
plot_data[["Date"] + account_vars].to_json("fedAssets.json", orient = "records")
account_vars = ["Federal Reserve Notes Outstanding",
"U.S. Treasury General Account",
"Deposits from Dep. Institutions",
"Other Deposits",
"Reverse Repos",
"Other Liabilities"]
total_var = "Total Liabilities"
plot_stacked_lines(
plot_data,
account_vars, linewidth = 1,
total_var = "Total Liabilities")
figsize= (36,18)
fig, ax = plt.subplots(figsize = figsize)
plot_data[account_vars].plot.area(stacked = True, linewidth = 3,
ax = ax)
# change y vals from mil to tril
total_var = "Total Liabilities"
plot_data[total_var].plot.line(linewidth = 2,
ax = ax, c = "k",
label = total_var, ls = "--")
plt.xticks(rotation = 90)
ax.legend(loc=2, ncol = 2)
ax.set_ylabel("$ Trillion", fontsize = 40)
ax.set_title("Federal Reserve Balance Sheet: Liabilities", fontsize = 50)
plot_data[[total_var] + account_vars].to_csv("FederalReserveLiabilities.csv")
plt.rcParams.update({"font.size":32})
df = plot_data.copy()
for key in account_vars:
df[key] = df[key].div(df["Total Liabilities"])
figsize= (36,18)
fig, ax = plt.subplots(figsize = figsize)
df[account_vars].plot.area(stacked = True, linewidth = 3,
ax = ax)
plt.xticks(rotation= 90, fontsize = 45)
ax.set_yticklabels([str(int(val * 100)) + "% " for val in ax.get_yticks()], fontsize = 45)
#plt.yticks([int(tick * 100) for tick in ax.get_yticks() if tick <1.01])
plt.title("Liabilities as a Proportion of the\nFederal Reserve's Balance Sheet", fontsize =45)
plt.show()
plot_data["Net Effect on Overnight Lending Market"] =\
plot_data["Repos"].sub(plot_data["Reverse Repos"])
overnight_vars = ["Repos",
"Reverse Repos",
"Net Effect on Overnight Lending Market"]
fig, ax = plt.subplots(figsize = (30,20))
plot_data[overnight_vars].plot.line(legend=True, linewidth = 3, ax = ax)
ax.set_ylabel("$ Trillion", fontsize = 40)
ax.set_title("Federal Reserve Activity\nin Overnight Lending Market", fontsize = 48)
plot_data[["Date"] + overnight_vars].to_json("overnightLending.json", orient="records")
data["Total Liabilities Growth Rate (%)"] = data["Total Liabilities"].pct_change(year)
data["Total Assets Growth Rate (%)"] = data["Total Assets"].pct_change(year)
data["Total Liabilities / Currency in Circulation"] = data["Total Liabilities"].div(data["Currency in Circulation ($ Bil)"].mul(1000))
data["Total Assets / Currency in Circulation"] = data["Total Assets"].div(data["Currency in Circulation ($ Bil)"].mul(1000))
data["Currency in Circulation / Total Assets"] = data["Currency in Circulation ($ Bil)"].mul(1000).div(data["Total Assets"])
data["Currency in Circulation / Total Assets Diff"] = data["Currency in Circulation / Total Assets"].diff(year)
data["Currency in Circulation / Total Liabilities"] = data["Currency in Circulation ($ Bil)"].mul(1000).div(data["Total Liabilities"])
data["Currency in Circulation / Total Liabilities Diff"] = data["Currency in Circulation / Total Liabilities"].diff(year)
data["Log Total Liabilities"] = np.log(data["Total Liabilities"])
data["Log Total Assets"] = np.log(data["Total Assets"])
#data["Currency in Circulation / Total Liabilities %"] = data["Currency in Circulation / Total Liabilities"].pct_change()
# pd.set_option('display.max_rows', None)
# pd.set_option('display.max_columns', None)
data
import pingouin
plot_vars = ["Inflation (PCE)",
"Unemployment Rate",
"Inflation Loss",
"Unemployment Loss",
"Linear Loss",
"Loss Function",
"Effective Federal Funds Rate (%)",
"Currency in Circulation Growth Rate (%)",
"Currency in Circulation / Total Assets",
"Total Assets Growth Rate (%)"]
#data[plot_vars].describe().T.to_excel("C:\\Users\\JLCat\\OneDrive\\Documents\\For NDSU\\Projects\\Sound Money Project\\Frederal Reserve QE Framework\\DescriptionStatistics.xls")
#data[plot_vars].describe().corr()
#data[plot_vars].describe().pcorr()
plot_vars = [#Currency in Circulation Growth Rate (%)",
#"Currency not in Circulation Growth Rate (%)",
#"5 Year Forward Rate",
#"Inflation (CPI)",
#"Unemployment Rate Diff",
#"Loss Function Diff",
#"Unemployment Loss Diff",
#"Inflation Loss Diff",
#"Linear Loss Diff",
#"Unemployment Loss Sq Diff",
#"Inflation Loss Sq Diff",
"Linear Loss Diff",
#"Currency in Circulation % Change Velocity",
#"Nominal GDP Growth Rate (%)",
#"Real GDP Growth Rate (%)",
"Effective Federal Funds Rate Diff (%)",
#"30 Year Treasury Rate (%)",
#"1 Month Treasury Rate (%)",
#"S&P Growth Rate (%)",
"Currency in Circulation / Total Assets Diff",
#Total Assets Growth Rate (%)",
"VIX Diff"
]
dag_df = data[plot_vars].dropna().rename(columns={key: key.replace(" ", "\n") for key in plot_vars}).loc["2008-10-01":"2020-08-01"]
dag_df.pcorr().round(2)
from pgmpy.estimators import PC
from pgmpy.base import DAG
import matplotlib.pyplot as plt
from matplotlib.patches import ArrowStyle
import networkx as nx
plt.rcParams.update({"font.size":20})
def graph_DAG(edges, df, title = ""):
pcorr = df.pcorr()
graph = nx.DiGraph()
edge_labels = {}
for edge in edges:
edge_labels[edge] = str(round(pcorr[edge[0]].loc[edge[1]],2))
graph.add_edges_from(edges)
color_map = ["C7" for g in graph]
fig, ax = plt.subplots(figsize = (15,20))
graph.nodes()
plt.tight_layout()
pos = nx.spring_layout(graph)#, k = 5/(len(sig_corr.keys())**.5))
plt.title(title, fontsize = 30)
nx.draw_networkx(graph, pos, node_color=color_map, node_size = 1200,
with_labels=True, arrows=True,
#font_color = "white",
font_size = 26, alpha = 1,
width = 1, edge_color = "C1",
arrowstyle=ArrowStyle("Fancy, head_length=3, head_width=1.5, tail_width=.1"), ax = ax)
nx.draw_networkx_edge_labels(graph,pos,
edge_labels=edge_labels,
font_color='green',
font_size=20)
plot_vars_dct = {0: ["Loss Function Diff",
"Effective Federal Funds Rate Diff (%)",
"Currency in Circulation / Total Assets Diff"],
# "VIX Diff"],
1:["Currency in Circulation Growth Rate (%)",
"Loss Function Diff",
"Effective Federal Funds Rate Diff (%)",
"Total Assets Growth Rate (%)"],
# "VIX Diff"],
2:["Unemployment Loss Sq Diff",
"Inflation Loss Sq Diff",
"Effective Federal Funds Rate Diff (%)",
"Currency in Circulation / Total Assets Diff"],
# "VIX Diff"],
3:["Currency in Circulation Growth Rate (%)",
"Unemployment Loss Sq Diff",
"Inflation Loss Sq Diff",
"Effective Federal Funds Rate Diff (%)",
"Total Assets Growth Rate (%)"],
#"VIX Diff"]
}
sig = 0.01
variant = "parallel"
ci_test = "pearsonr"
for key, plot_vars in plot_vars_dct.items():
dfs = {"2008-10-31 to 2020-07-31": data[plot_vars].dropna().rename(columns={key: key.replace(" ", "\n") for key in plot_vars}).loc["2008-10-31":"2020-08-31"],
"2003-12-31 to 2008-09-30": data[plot_vars].dropna().rename(columns={key: key.replace(" ", "\n") for key in plot_vars}).loc[:"2008-09-30"]}
for dates, dag_df in dfs.items():
keys = dag_df.keys()
c = PC(dag_df.dropna())
max_cond_vars = len(keys) - 2
# print(dag_df.index)
model = c.estimate(return_type = "dag",variant= variant,
significance_level = sig,
max_cond_vars = max_cond_vars, ci_test = ci_test)
edges = model.edges()
pcorr = dag_df.pcorr()
weights = {}
# for edge in edges:
# print(edge, ":",pcorr[edge[0]].loc[edge[1]])
# skel, sep_sets = c.build_skeleton(variant = variant, ci_test = ci_test, significance_level = sig,
# max_cond_vars = max_cond_vars)
graph_DAG(edges, dag_df, title = dates)
from pgmpy.estimators import PC
from pgmpy.base import DAG
keys = dag_df.keys()
c = PC(dag_df.dropna())
max_cond_vars = len(keys) - 2
sig = 0.05
variant = "parallel"
ci_test = "pearsonr"
model = c.estimate(return_type = "dag",variant= variant,
significance_level = sig,
max_cond_vars = max_cond_vars, ci_test = ci_test)
edges = model.edges()
pcorr = dag_df.pcorr()
weights = {}
for edge in edges:
print(edge, ":",pcorr[edge[0]].loc[edge[1]])
skel, sep_sets = c.build_skeleton(variant = variant, ci_test = ci_test, significance_level = 0.01,
max_cond_vars = max_cond_vars)
sep_sets
import matplotlib.pyplot as plt
from matplotlib.patches import ArrowStyle
import networkx as nx
plt.rcParams.update({"font.size":20})
def graph_DAG(edges, df, title = ""):
pcorr = df.pcorr()
graph = nx.DiGraph()
edge_labels = {}
for edge in edges:
edge_labels[edge] = str(round(pcorr[edge[0]].loc[edge[1]],2))
graph.add_edges_from(edges)
color_map = ["C0" for g in graph]
fig, ax = plt.subplots(figsize = (20,20))
graph.nodes()
plt.tight_layout()
pos = nx.spring_layout(graph)#, k = 5/(len(sig_corr.keys())**.5))
plt.title(title, fontsize = 30)
nx.draw_networkx(graph, pos, node_color=color_map, node_size = 1200,
with_labels=True, arrows=True,
#font_color = "white",
font_size = 26, alpha = 1,
width = 1, edge_color = "C1",
arrowstyle=ArrowStyle("Fancy, head_length=3, head_width=1.5, tail_width=.1"), ax = ax)
nx.draw_networkx_edge_labels(graph,pos,
edge_labels=edge_labels,
font_color='green',
font_size=20)
graph_DAG(edges, dag_df)
from pgmpy.estimators import PC
from pgmpy.base import DAG
keys = dag_df.keys()
c = PC(dag_df.dropna())
max_cond_vars = len(keys) - 2
variant = "parallel"
ci_test = "pearsonr"
model = c.estimate(return_type = "dag",variant= variant,
significance_level = sig,
max_cond_vars = max_cond_vars, ci_test = ci_test)
edges = model.edges()
pcorr = dag_df.pcorr()
weights = {}
for edge in edges:
print(edge, ":",pcorr[edge[0]].loc[edge[1]])
skel, sep_sets = c.build_skeleton(variant = variant, ci_test = ci_test, significance_level = 0.01,
max_cond_vars = max_cond_vars)
sep_sets
graph_DAG(edges, dag_df)
from pgmpy.estimators import PC
from pgmpy.base import DAG
keys = dag_df.keys()
c = PC(dag_df.dropna())
max_cond_vars = len(keys) - 2
variant = "parallel"
ci_test = "pearsonr"
model = c.estimate(return_type = "dag",variant= variant,
significance_level = sig,
max_cond_vars = max_cond_vars, ci_test = ci_test)
edges = model.edges()
pcorr = dag_df.pcorr()
weights = {}
for edge in edges:
print(edge, ":",pcorr[edge[0]].loc[edge[1]])
skel, sep_sets = c.build_skeleton(variant = variant, ci_test = ci_test, significance_level = 0.01,
max_cond_vars = max_cond_vars)
sep_sets
graph_DAG(edges, dag_df)
from pgmpy.estimators import PC
from pgmpy.base import DAG
keys = dag_df.keys()
c = PC(dag_df.dropna())
max_cond_vars = len(keys) - 2
variant = "parallel"
ci_test = "pearsonr"
model = c.estimate(return_type = "dag",variant= variant,
significance_level = sig,
max_cond_vars = max_cond_vars, ci_test = ci_test)
edges = model.edges()
pcorr = dag_df.pcorr()
weights = {}
for edge in edges:
print(edge, ":",pcorr[edge[0]].loc[edge[1]])
skel, sep_sets = c.build_skeleton(variant = variant, ci_test = ci_test, significance_level = 0.01,
max_cond_vars = max_cond_vars)
sep_sets
for key in sep_sets:
print(list(sep_sets[key]))
graph_DAG(edges, dag_df)
import matplotlib.pyplot as plt
from matplotlib.patches import ArrowStyle
import networkx as nx
plt.rcParams.update({"font.size":20})
def graph_DAG(edges, df,sep_sets, title = ""):
graph = nx.DiGraph()
edge_labels = {}
pcorr = df.pcorr()
for edge in sep_sets:
sep_set = list(sep_sets[edge])
key = list(edge)
# pcorr = df[key + sep_set].pcorr()
edge_labels[edge] = str(round(pcorr[key[0]].loc[key[1]],2))
print(edge_labels)
graph.add_edges_from(sep_sets.keys())
color_map = ["C0" for g in graph]
fig, ax = plt.subplots(figsize = (20,20))
graph.nodes()
plt.tight_layout()
pos = nx.spring_layout(graph)#, k = 5/(len(sig_corr.keys())**.5))
plt.title(title, fontsize = 30)
nx.draw_networkx(graph, pos, node_color=color_map, node_size = 1200,
with_labels=True, arrows=True,
#font_color = "white",
font_size = 26, alpha = 1,
width = 1, edge_color = "C1",
arrowstyle=ArrowStyle("Fancy, head_length=3, head_width=1.5, tail_width=.1"), ax = ax)
nx.draw_networkx_edge_labels(graph,pos,
edge_labels=edge_labels,
font_color='green',
font_size=20)
graph_DAG(edges, dag_df, sep_sets)
dag_df
figsize = (20,12)
fig,ax = plt.subplots(figsize = figsize)
ax2 = ax.twinx()
ax3 = ax.twinx()
data[["Currency in Circulation / Total Assets"]].plot(legend = False,
linewidth = 5,
alpha = .8,
ax = ax)
data[["Effective Federal Funds Rate (%)"]].plot(legend = False,
linewidth = 5,
alpha = .8,
ax = ax2,
c = "C1")
data[["Linear Loss"]].iloc[:-1].plot(legend = False,
linewidth = 5,
alpha = .1,
ax = ax3,
c = "k")
ax.set_yticklabels([str(int(val * 100)) + "%" for val in ax.get_yticks()], c = "C0")
ax2.set_yticklabels([str(int(val)) + "%" for val in ax2.get_yticks()], c = "C1")
ax3.set_yticklabels(ax3.get_yticks(), alpha = .2, c = "k")
ax3.set_yticks([])
ax3.axhline(0, c = "k", alpha = .1, linewidth = 5, ls = "--")
plt.axvline(datetime.datetime(2008,10,1), c = "k", ls = "--", alpha = .9)
ax.text(datetime.datetime(2013,3,1), .42, "Currency in Circulation\n/ \nTotal Assets", fontsize = 25, c = "C0",
ha = "center")
ax.text(datetime.datetime(2018,5,1), .58, "Effective\nFederal\nFunds\nRate", fontsize = 25, c = "C1",
ha = "center")
ax.text(datetime.datetime(2014,6,1), .78, "Linear\nLoss\nFunction", fontsize = 25, c = "k", alpha = .3,
ha = "center")
ax.text(datetime.datetime(2002,10,1), .955, "0", fontsize = 30, c = "k", alpha = .3,
ha = "right")
ax.text(datetime.datetime(2022,1,1), .955, "0 ", fontsize = 30, c = "k", alpha = .3,
ha = "right")
fig, ax = plt.subplots(figsize = (30,15))
data[["Linear Loss Diff", "Total Assets Growth Rate (%)"]].plot.scatter(x = "Linear Loss Diff", y = "Total Assets Growth Rate (%)", ax = ax)
plt.xticks(rotation=90)
fig, ax = plt.subplots(figsize = (30,15))
data.iloc[:-1].plot.scatter(x = "Linear Loss", y = "Currency in Circulation / Total Assets", ax = ax)
plt.xticks(rotation=90)
```
| github_jupyter |
```
import csv
import pandas as pd
test = pd.read_csv("../input/bert-18000-128/dataset/test.csv", escapechar = "\\", quoting = csv.QUOTE_NONE)
from transformers import BertTokenizerFast
tokenizer = BertTokenizerFast.from_pretrained('bert-base-uncased')
import pickle
file = open("../input/bert-18000-128/lable_map.pickle",'rb')
label_map = pickle.load(file)
file.close()
label_to_id = {}
for key, value in label_map.items():
label_to_id[value] = key
import torch
class Dataset(torch.utils.data.Dataset):
def __init__(self, df, tokenizer, is_train=True, label_map={}, max_length=128, load_desc = True, load_bullets = True):
self.df = df
self.load_desc = load_desc
self.load_bullets = load_bullets
self.title = df.TITLE.values
self.desc = df.DESCRIPTION.values
self.bullets = df.BULLET_POINTS.apply(lambda x: x[1:-1] if len(x)>0 and x[0]=='[' else x).values
self.tokenizer = tokenizer
if is_train:
self.labels = df.BROWSE_NODE_ID.apply(lambda x: label_map[x]).values
self.label_map = label_map
self.is_train = is_train
self.max_length = max_length
def __getitem__(self, idx):
req_string = self.title[idx] + ' ~ '
if self.load_desc:
req_string += self.desc[idx]
req_string += ' ~ '
if self.load_bullets:
req_string += self.bullets[idx]
tokenized_data = tokenizer.tokenize(req_string)
to_append = ["[CLS]"] + tokenized_data[:self.max_length - 2] + ["[SEP]"]
input_ids = tokenizer.convert_tokens_to_ids(to_append)
input_mask = [1] * len(input_ids)
padding = [0] * (self.max_length - len(input_ids))
input_ids += padding
input_mask += padding
item = {
"input_ids": torch.tensor(input_ids),
"attention_mask": torch.tensor(input_mask),
"token_type_ids": torch.tensor([0]*self.max_length)
}
if self.is_train:
item['labels'] = torch.tensor(self.labels[idx])
return item
def __len__(self):
return len(self.df)
from transformers import BertPreTrainedModel, Trainer, BertModel
from transformers.modeling_outputs import SequenceClassifierOutput
from torch import nn
class BertForSequenceClassification(BertPreTrainedModel):
def __init__(self, config, num_labels=len(label_map)):
super().__init__(config)
self.num_labels = num_labels
self.config = config
self.bert = BertModel(config)
self.dropout = nn.Dropout(config.hidden_dropout_prob)
self.classifier = nn.Linear(config.hidden_size, self.num_labels)
self.init_weights()
def forward(
self,
input_ids=None,
attention_mask=None,
token_type_ids=None,
position_ids=None,
head_mask=None,
inputs_embeds=None,
labels=None,
output_attentions=None,
output_hidden_states=None,
return_dict=None,
):
r"""
labels (:obj:`torch.LongTensor` of shape :obj:`(batch_size,)`, `optional`):
Labels for computing the sequence classification/regression loss. Indices should be in :obj:`[0, ...,
config.num_labels - 1]`. If :obj:`config.num_labels == 1` a regression loss is computed (Mean-Square loss),
If :obj:`config.num_labels > 1` a classification loss is computed (Cross-Entropy).
"""
return_dict = return_dict if return_dict is not None else self.config.use_return_dict
outputs = self.bert(
input_ids,
attention_mask=attention_mask,
token_type_ids=token_type_ids,
position_ids=position_ids,
head_mask=head_mask,
inputs_embeds=inputs_embeds,
output_attentions=output_attentions,
output_hidden_states=output_hidden_states,
return_dict=return_dict,
)
pooled_output = outputs[1]
pooled_output = self.dropout(pooled_output)
logits = self.classifier(pooled_output)
loss = None
if labels is not None:
if self.config.problem_type is None:
if self.num_labels == 1:
self.config.problem_type = "regression"
elif self.num_labels > 1 and (labels.dtype == torch.long or labels.dtype == torch.int):
self.config.problem_type = "single_label_classification"
else:
self.config.problem_type = "multi_label_classification"
if self.config.problem_type == "regression":
loss_fct = MSELoss()
if self.num_labels == 1:
loss = loss_fct(logits.squeeze(), labels.squeeze())
else:
loss = loss_fct(logits, labels)
elif self.config.problem_type == "single_label_classification":
loss_fct = CrossEntropyLoss()
loss = loss_fct(logits.view(-1, self.num_labels), labels.view(-1))
elif self.config.problem_type == "multi_label_classification":
loss_fct = BCEWithLogitsLoss()
loss = loss_fct(logits, labels)
if not return_dict:
output = (logits,) + outputs[2:]
return ((loss,) + output) if loss is not None else output
return SequenceClassifierOutput(
loss=loss,
logits=logits,
hidden_states=outputs.hidden_states,
attentions=outputs.attentions,
)
model = BertForSequenceClassification.from_pretrained("../input/bert-18000-128/results/checkpoint-18000")
test_dataset = Dataset(test.fillna(""), tokenizer, is_train=False, load_desc = False, load_bullets = True)
from torch.utils.data import DataLoader
from tqdm.auto import tqdm
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
model.eval()
test_loader = DataLoader(test_dataset, batch_size=16, shuffle=False)
logits = []
with torch.no_grad():
for batch in tqdm(test_loader):
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
outputs = model(input_ids, attention_mask=attention_mask)
logits.append(outputs["logits"])
labels = torch.cat(logits).argmax(1).cpu()
data_dict = {
"PRODUCT_ID": range(1, 110776),
"BROWSE_NODE_ID": labels
}
submit = pd.DataFrame.from_dict(data_dict)
submit["BROWSE_NODE_ID"] = submit["BROWSE_NODE_ID"].apply(lambda x: label_to_id[x])
submit.to_csv("BERTbase_18000_no_desc.csv", index=False)
submit.head()
logits = torch.cat(logits).cpu()
with open('logits_bert_18000_no_desc.pickle', 'wb') as handle:
pickle.dump(logits, handle, protocol=pickle.HIGHEST_PROTOCOL)
test_dataset = Dataset(test.fillna(""), tokenizer, is_train=False, load_desc = True, load_bullets = False)
from torch.utils.data import DataLoader
from tqdm.auto import tqdm
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
model.eval()
test_loader = DataLoader(test_dataset, batch_size=16, shuffle=False)
logits = []
with torch.no_grad():
for batch in tqdm(test_loader):
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
outputs = model(input_ids, attention_mask=attention_mask)
logits.append(outputs["logits"])
labels = torch.cat(logits).argmax(1).cpu()
data_dict = {
"PRODUCT_ID": range(1, 110776),
"BROWSE_NODE_ID": labels
}
submit = pd.DataFrame.from_dict(data_dict)
submit["BROWSE_NODE_ID"] = submit["BROWSE_NODE_ID"].apply(lambda x: label_to_id[x])
submit.to_csv("BERTbase_18000_no_bullet.csv", index=False)
submit.head()
logits = torch.cat(logits).cpu()
with open('logits_bert_18000_no_bullet.pickle', 'wb') as handle:
pickle.dump(logits, handle, protocol=pickle.HIGHEST_PROTOCOL)
test_dataset = Dataset(test.fillna(""), tokenizer, is_train=False, load_desc = False, load_bullets = False)
from torch.utils.data import DataLoader
from tqdm.auto import tqdm
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
model.eval()
test_loader = DataLoader(test_dataset, batch_size=16, shuffle=False)
logits = []
with torch.no_grad():
for batch in tqdm(test_loader):
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
outputs = model(input_ids, attention_mask=attention_mask)
logits.append(outputs["logits"])
labels = torch.cat(logits).argmax(1).cpu()
data_dict = {
"PRODUCT_ID": range(1, 110776),
"BROWSE_NODE_ID": labels
}
submit = pd.DataFrame.from_dict(data_dict)
submit["BROWSE_NODE_ID"] = submit["BROWSE_NODE_ID"].apply(lambda x: label_to_id[x])
submit.to_csv("BERTbase_18000_no_desc_no_bullet.csv", index=False)
submit.head()
logits = torch.cat(logits).cpu()
with open('logits_bert_18000_no_desc_no_bullet.pickle', 'wb') as handle:
pickle.dump(logits, handle, protocol=pickle.HIGHEST_PROTOCOL)
test_dataset = Dataset(test.fillna(""), tokenizer, is_train=False, load_desc = True, load_bullets = True)
from torch.utils.data import DataLoader
from tqdm.auto import tqdm
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
model.eval()
test_loader = DataLoader(test_dataset, batch_size=16, shuffle=False)
logits = []
with torch.no_grad():
for batch in tqdm(test_loader):
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
outputs = model(input_ids, attention_mask=attention_mask)
logits.append(outputs["logits"])
labels = torch.cat(logits).argmax(1).cpu()
data_dict = {
"PRODUCT_ID": range(1, 110776),
"BROWSE_NODE_ID": labels
}
submit = pd.DataFrame.from_dict(data_dict)
submit["BROWSE_NODE_ID"] = submit["BROWSE_NODE_ID"].apply(lambda x: label_to_id[x])
submit.to_csv("BERTbase_18000.csv", index=False)
submit.head()
logits = torch.cat(logits).cpu()
with open('logits_bert_18000.pickle', 'wb') as handle:
pickle.dump(logits, handle, protocol=pickle.HIGHEST_PROTOCOL)
```
| github_jupyter |
# Step 3 - Snap CXB OD Estimates
This notebook is the final step of calculating the base OD matrices which underlie all GOSTNets-derived accessibility analysis. It's also the slowest step
```
import geopandas as gpd
import pandas as pd
import os, sys, time, importlib
# add to your system path the location of the LoadOSM.py and GOSTnet.py scripts
# sys.path.append("../../../GOSTnets/GOSTnets")
import GOSTnets as gn
import importlib
# import Network_Clean as gnClean
importlib.reload(gn)
import networkx as nx
import osmnx
from shapely.geometry import Point
import numpy as np
import rasterio
from rasterio import features
from shapely.wkt import loads
import ipyparallel as ipp
```
### Setup
paths
```
input_pth = r'inputs\\dests'
geo_pth = r'../../../GEO'
fin_pth = 'final'
```
pickles
```
fin_pckle = r'final_current_G.pickle'
upgr_all = r'final_upgrade_all_G.pickle'
upgr_nosouth = r'final_upgrade_nosouth_G.pickle'
upgr_noferry = r'final_upgrade_noferry_G.pickle'
G_current = nx.read_gpickle(os.path.join(fin_pth, fin_pckle))
G_upgr_all = nx.read_gpickle(os.path.join(fin_pth, upgr_all))
G_upgr_nosouth = nx.read_gpickle(os.path.join(fin_pth, upgr_nosouth))
G_upgr_noferry = nx.read_gpickle(os.path.join(fin_pth, upgr_noferry))
```
origins
```
# grid_name = r'hrsl_2018_cxb_pts_snapped.csv'
grid_name = r'growth_center_origins_snapped.csv'
origins = pd.read_csv(os.path.join(fin_pth,grid_name))
grid = origins
```
destinations
```
cxb_fil = r'cxb_ctr.shp'
chitt_fil = r'chittagong.shp'
health_fil = r'hc_merge_200324_4326.shp'
primary_fil = r'schools/school_category_primary.gpkg'
secondary_fil = r'schools/school_category_secondary.gpkg'
tertiary_fil = r'schools/school_category_tertiary.gpkg'
matar_fil = r'martarbari.shp'
mkts_fil = r'mkts_merge_4326.shp'
gc_fil = r'cxb_lged_gc_moved_4326.shp'
dests = {"CXB" : cxb_fil, "Chittagong" : chitt_fil, "Health" : health_fil, \
"Primary_education" : primary_fil, "Secondary_education" : secondary_fil, "Tertiary_education" : tertiary_fil, \
"Martarbari" : matar_fil, "All_markets" : mkts_fil, "Growth_centers" : gc_fil}
# note you can use smaller / larger dest dictionaries as needed
# This is helpful for going back and re-running only certain destinations, or adding in new ones.
# dests = {"CXB" : cxb_fil}
```
scenarios (for looping)
```
# the dict here allows us to match a label to a pickle
scenarios = {'current' : [G_current,'_current_snapped.csv'],\
'upgrade_all' : [G_upgr_all,'_ua_snapped.csv'],\
'upgrade_nosouth' : [G_upgr_nosouth,'_uns_snapped.csv'],\
'upgrade_noferry' : [G_upgr_noferry,'_unf_snapped.csv']}
```
Settings
```
walk_speed = 4.5
WGS = {'epsg':'4326'}
measure_crs = {'epsg':'32646'}
# date = 'May2020'
```
#### Troubleshooting
```
# dests = {"All_markets" : mkts_fil, "Growth_centers" : gc_fil}
# cur = {'current' : [G_current,'_current_snapped.csv']}
# for scen, values in cur.items():
# # namestr(values, globals())
# od_routine(values[0],scen,values[1])
# for scen, values in cur.items():
# add_walking_time(scen,values[1])
# for u, v, data in G_upgr_all.edges(data=True):
# if data['osm_id'] == '244861071':
# print(data['infra_type'])
# for u, v, data in G_upgr_nosouth.edges(data=True):
# if data['osm_id'] == '244861071':
# print(data['infra_type'])
```
#### Define functions that will generate OD matrices, and then generate direct walking times from origins to destinations
The benefit of functions is they allow us to loop the analysis over the dictionary items
These functions are big, messy, and adapted from old GOSTNets code. Though functional, the code could likely be streamlined as it's tricky to troubleshoot.
```
def od_routine(G_input,scenario='',snap_ending='_snapped.csv'):
origins_pop_nodes = list(set(origins.NN)) # consolidates by shared nodes.
for dest_type, fpth in dests.items():
snapfile = dest_type + snap_ending
print(dest_type)
dest = pd.read_csv(os.path.join(fin_pth,snapfile))
dest_nodes = list(set(dest.NN))
print(len(list(set(dest.NN))))
od_time = gn.calculate_OD(G_input, origins=origins_pop_nodes,
destinations=dest_nodes, fail_value=99999999, weight='time')
od_time_df = pd.DataFrame(od_time, index=origins_pop_nodes, columns=dest_nodes)
print(od_time_df.shape)
# Add walking time (from origin to NN) for each OD
# origins_join = origins_pop_snapped.merge(od_time_df, how='left', on='NN')
origins['NN_dist_seconds'] = ((origins.NN_dist / 1000) / walk_speed) * 60 * 60
origins_join = origins.join(od_time_df, on='NN', rsuffix="dist_")
# print(origins_join.head())
origins_join.columns[6:len(origins_join.columns)]
origins_join.to_csv(os.path.join(fin_pth,'origins_walktime_{}_NN_{}.csv'.format(scenario,dest_type)))
od_time_df.to_csv(os.path.join(fin_pth,'OD_matrix_{}_NN_{}.csv'.format(scenario,dest_type)))
def add_walking_time(scenario='',snap_ending='_snapped.csv'):
# Main method
print(scenario)
for dest_type, fpth in dests.items():
snapfile = dest_type + snap_ending
print(dest_type)
OD_name = r'OD_matrix_{}_NN_{}.csv'.format(scenario,dest_type)
OD = pd.read_csv(os.path.join(fin_pth, OD_name))
# OD = od_time_df
OD = OD.rename(columns = {'Unnamed: 0':'O_ID'})
OD = OD.set_index('O_ID')
OD = OD.replace([np.inf, -np.inf], np.nan)
# # Filtering by only desired destination in an all-destination OD matrix.
# # Skipping for now
od_dest_df = pd.read_csv(os.path.join(fin_pth,snapfile))
od_dest_df['geometry'] = od_dest_df['geometry'].apply(loads)
# od_dest_gdf = gpd.GeoDataFrame(od_dest_df, crs = {'init':'epsg:4326'}, geometry = 'geometry')
# accepted_facilities = list(set(list(acceptable_df.NN)))
# accepted_facilities_str = [str(i) for i in accepted_facilities]
# print(accepted_facilities)
# print(accepted_facilities_str)
# # OD = OD_original[accepted_facilities_str] # not necessary, already done
# # acceptable_df.to_csv(os.path.join(basepth,'Output','%s.csv' % subset))
# Computing walk time from network to destination
dest = pd.read_csv(os.path.join(fin_pth,snapfile))
dest_df = dest[['NN','NN_dist']]
dest_df = dest_df.set_index('NN')
dest_df['NN_dist'] = dest_df['NN_dist'] / 1000 * 3600 / walk_speed
dest_df.index = dest_df.index.map(str)
d_f = OD.transpose()
for i in d_f.columns:
dest_df[i] = d_f[i]
for i in dest_df.columns:
if i == 'NN_dist':
pass
else:
dest_df[i] = dest_df[i] + dest_df['NN_dist']
dest_df = dest_df.drop('NN_dist', axis = 1)
dest_df = dest_df.transpose()
dest_df['min_time'] = dest_df.min(axis = 1)
# dest_df['geometry'] = dest_df['geometry'].apply(loads)
dest_gdf = gpd.GeoDataFrame(od_dest_df, geometry = 'geometry', crs = {'init':'epsg:4326'})
# Add walk time from origin to network
grid = pd.read_csv(os.path.join(fin_pth, grid_name))
grid = grid.rename(columns = {'NN':'O_ID','NN_dist':'walk_to_road_net'})
grid = grid.set_index(grid['O_ID'])
grid['on_network_time'] = dest_df['min_time']
grid['walk_to_road_net'] = grid['walk_to_road_net'] / 1000 * 3600 / walk_speed
grid['total_time_net'] = grid['on_network_time'] + grid['walk_to_road_net']
# print(grid.head())
grid['geometry'] = grid['geometry'].apply(loads)
o_2_d = gpd.GeoDataFrame(grid, crs = {'init':'epsg:4326'}, geometry = 'geometry')
# Snapping!
print('start of snapping: %s\n' % time.ctime())
o_2_d = gn.pandana_snap_points(o_2_d,
dest_gdf, # eventually just use dest_gdf
source_crs='epsg:4326',
target_crs='epsg:32646',
add_dist_to_node_col = True)
print('\nend of snapping: %s' % time.ctime())
print('\n--- processing complete')
# Recalculating the resulting walking times into seconds and minutes.
# Make sure that if walking is faster than on-network travel, it prefers walking
o_2_d['walk_time_direct'] = o_2_d['idx_dist'] / 1000 * 3600 / walk_speed
grid['walk_time_direct'] = o_2_d['walk_time_direct']
# grid['PLOT_TIME_SECS'] = grid[['total_time_net']].min(axis = 1)
# grid['PLOT_TIME_SECS'] = grid[['walk_to_road_net','total_time_net']].min(axis = 1)
# The city locations / port location don't have walk_time_direct values so we use if/else logic to work around them.
if 'walk_time_direct' in grid.columns:
grid['PLOT_TIME_SECS'] = grid[['walk_time_direct','total_time_net']].min(axis = 1)
else:
grid['PLOT_TIME_SECS'] = grid[['total_time_net']]
grid['PLOT_TIME_MINS'] = grid['PLOT_TIME_SECS'] / 60
if 'walk_time_direct' in grid.columns:
def choice(x):
if x.walk_time_direct < x.total_time_net:
return 'walk'
else:
return 'net'
grid['choice'] = grid.apply(lambda x: choice(x), axis = 1)
grid['choice'].value_counts()
# print(grid.head())
# Export
grid.to_csv(os.path.join(fin_pth,'final_cxb_{}_od_grid_{}.csv'.format(scenario,dest_type)))
```
### Load origins and destinations, get unique origin nodes, run OD, export
Run the OD routine function on all destinations for the 3 different scenarios
```
for scen, values in scenarios.items():
od_routine(values[0],scen,values[1])
```
### Import completed OD matrix, calcualte walking times from origins to the destinations
```
for scen, values in scenarios.items():
add_walking_time(scen,values[1])
```
### That's it
You now have completed OD matrices for everything
| github_jupyter |
# Interactive Machine Learning Demo - Logistic Regression
```
from ipywidgets import interact, interactive, IntSlider, Layout
import ipywidgets as widgets
from IPython.display import display
import numpy as np
import matplotlib.pyplot as plt
#%matplotlib inline
import pandas as pd
```
## Logistic Regression and Classficiation
```
from sklearn.linear_model import logistic
from sklearn.model_selection import train_test_split
```
### Function definition generating data points with noise and classifying
```
def func_log(N_samples,x_min,x_max,noise_magnitude,noise_sd,noise_mean):
x= np.linspace(x_min,x_max,N_samples*5)
x1= np.random.choice(x,size=N_samples)
x2= np.random.choice(x,size=N_samples)
y=1.5*x1-2*x2
yn= y+noise_magnitude*np.random.normal(loc=noise_mean,scale=noise_sd,size=N_samples)
plt.figure(figsize=(8,5))
c = [i < 0 for i in y]
f, ax = plt.subplots(figsize=(8, 6))
#ax.contour(xx, yy, probs, levels=[.5], cmap="Greys", vmin=0, vmax=.6)
ax.set_xlim(left=x_min*1.2,right=x_max*1.2)
ax.set_ylim(bottom=x_min*1.2,top=x_max*1.2)
ax.scatter(x1,x2,c=c,edgecolors='k',s=80)
plt.grid(True)
plt.show()
return (np.array(x),np.array(y),np.array(yn),np.array(x1),np.array(x2))
```
### Plot and interactively control the data points by calling ipywidget object
```
pl=interactive(func_log,N_samples={'Low (50 samples)':50,'High (200 samples)':200},x_min=(-5,0,1), x_max=(0,5,1),
noise_magnitude=(0,5,1),noise_sd=(0.1,1,0.1),noise_mean=(-2,2,0.5))
display(pl)
```
### Store the generated data
```
xl,yl,ynl,x1l,x2l = pl.result
dic = {'X1':x1l,'X2':x2l,'out':ynl}
df = pd.DataFrame(data=dic)
def boolean (x):
return int(x>0)
df['y']=df['out'].apply(boolean)
df.head()
```
### Logistic regression encapsulated within a function
```
def log_fit(C,test_size,penalty):
X_train, X_test, y_train, y_test = train_test_split(df[['X1','X2']],df['y'],test_size=test_size,random_state=101)
df_test=pd.DataFrame(X_test,columns=['X1','X2'])
df_test['y']=y_test
df_train=pd.DataFrame(X_train,columns=['X1','X2'])
df_train['y']=y_train
if (penalty=='L1 norm'):
logm = logistic.LogisticRegressionCV(Cs=[np.power(10,-C)],penalty='l1',solver='saga')
if (penalty=='L2 norm'):
logm = logistic.LogisticRegressionCV(Cs=[np.power(10,-C)],penalty='l2',solver='lbfgs')
logm.fit(X_train,y_train)
train_score = logm.score(X_train,y_train)
test_score = logm.score(X_test,y_test)
xx, yy = np.mgrid[-5:5:.01, -5:5:.01]
grid = np.c_[xx.ravel(), yy.ravel()]
probs = logm.predict_proba(grid)[:, 1].reshape(xx.shape)
c_test = [i for i in df_test['y']]
c_train = [i for i in df_train['y']]
f, ax = plt.subplots(nrows=1,ncols=2,figsize=(15,7))
contour = ax[0].contourf(xx, yy, probs, 25, cmap="RdBu",vmin=0, vmax=1)
ax1_c = f.colorbar(contour)
ax1_c.set_label("$Prob\ (y = 1)$",fontsize=15)
ax1_c.set_ticks([0, .25, .5, .75, 1])
ax[0].set_xlabel("$X_1$",fontsize=15)
ax[0].set_ylabel("$X_2$",fontsize=15)
ax[0].contour(xx, yy, probs, levels=[.5], cmap="Greys", vmin=0, vmax=0.6,lw=3)
ax[0].scatter(df_test['X1'],df_test['X2'],c=c_test,edgecolors='k',s=100)
ax[0].set_title("\nTest score: %.3f\n"%(test_score),fontsize=20)
contour = ax[1].contourf(xx, yy, probs, 25, cmap="RdBu",vmin=0, vmax=1)
#ax2_c = f.colorbar(contour)
#ax2_c.set_label("$Prob\ (y = 1)$",fontsize=15)
#ax2_c.set_ticks([0, .25, .5, .75, 1])
ax[1].set_xlabel("$X_1$",fontsize=15)
ax[1].set_ylabel("$X_2$",fontsize=15)
ax[1].contour(xx, yy, probs, levels=[.5], cmap="Greys", vmin=0, vmax=0.6,lw=3)
ax[1].scatter(df_train['X1'],df_train['X2'],c=c_train,edgecolors='k',s=100)
ax[1].set_title("\nTraining score: %.3f\n"%(train_score),fontsize=20)
plt.grid(True)
plt.tight_layout()
plt.show()
return (train_score,test_score)
```
### Run the encapsulated ML function with ipywidget interactive
```
from ipywidgets import HBox, Label, FloatSlider
style = {'description_width': 'initial'}
lb1=Label (value="Play with options and watch the probability space evolve dynamically. \
Remember, smaller the value of $C$, stronger the regularization strength",fontsize=15)
logp=interactive(log_fit,C=FloatSlider(value=0,min=0,max=2,step=0.1,
description='Regularization ($10^{-C}$), $C$', style=style,continuous_update=False),
test_size=FloatSlider(value=0.1,min=0.1,max=0.5,step=0.1,
description = 'Test fraction ($X_{test}$)', style=style),
penalty=widgets.RadioButtons(options=["L1 norm","L2 norm"], description = 'Penalty norm'))
# Set the height of the control.children[-1] so that the output does not jump and flicker
output = logp.children[-1]
output.layout.height = '500px'
output.layout.width = '1000px'
display(lb1)
display(logp)
```
| github_jupyter |
## 1. The most Nobel of Prizes
<p><img style="float: right;margin:5px 20px 5px 1px; max-width:250px" src="https://assets.datacamp.com/production/project_441/img/Nobel_Prize.png"></p>
<p>The Nobel Prize is perhaps the world's most well known scientific award. Except for the honor, prestige and substantial prize money the recipient also gets a gold medal showing Alfred Nobel (1833 - 1896) who established the prize. Every year it's given to scientists and scholars in the categories chemistry, literature, physics, physiology or medicine, economics, and peace. The first Nobel Prize was handed out in 1901, and at that time the Prize was very Eurocentric and male-focused, but nowadays it's not biased in any way whatsoever. Surely. Right?</p>
<p>Well, we're going to find out! The Nobel Foundation has made a dataset available of all prize winners from the start of the prize, in 1901, to 2016. Let's load it in and take a look.</p>
```
# Loading in required libraries
import pandas as pd
import seaborn as sns
import numpy as np
# Reading in the Nobel Prize data
nobel = pd.read_csv('datasets/nobel.csv')
# Taking a look at the first several winners
nobel.head(6)
```
## 2. So, who gets the Nobel Prize?
<p>Just looking at the first couple of prize winners, or Nobel laureates as they are also called, we already see a celebrity: Wilhelm Conrad Röntgen, the guy who discovered X-rays. And actually, we see that all of the winners in 1901 were guys that came from Europe. But that was back in 1901, looking at all winners in the dataset, from 1901 to 2016, which sex and which country is the most commonly represented? </p>
<p>(For <em>country</em>, we will use the <code>birth_country</code> of the winner, as the <code>organization_country</code> is <code>NaN</code> for all shared Nobel Prizes.)</p>
```
# Display the number of (possibly shared) Nobel Prizes handed
# out between 1901 and 2016
display(len(nobel[(nobel['year'] >= 1901) & (nobel['year'] <= 2016)]))
# Display the number of prizes won by male and female recipients.
display(nobel['sex'].value_counts())
# Display the number of prizes won by the top 10 nationalities.
nobel['birth_country'].value_counts().head(10)
```
## 3. USA dominance
<p>Not so surprising perhaps: the most common Nobel laureate between 1901 and 2016 was a man born in the United States of America. But in 1901 all the winners were European. When did the USA start to dominate the Nobel Prize charts?</p>
```
# Calculating the proportion of USA born winners per decade
nobel['usa_born_winner'] = nobel['birth_country']=='United States of America'
nobel['decade'] = (np.floor(nobel['year'] / 10) * 10).astype(int)
prop_usa_winners = nobel.groupby('decade', as_index=False)['usa_born_winner'].mean()
# Display the proportions of USA born winners per decade
prop_usa_winners
```
## 4. USA dominance, visualized
<p>A table is OK, but to <em>see</em> when the USA started to dominate the Nobel charts we need a plot!</p>
```
# Setting the plotting theme
sns.set()
# and setting the size of all plots.
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [11, 7]
# Plotting USA born winners
ax = sns.lineplot(x='decade',
y='usa_born_winner',
data=prop_usa_winners)
# Adding %-formatting to the y-axis
from matplotlib.ticker import PercentFormatter
ax.yaxis.set_major_formatter(PercentFormatter(1.0))
```
## 5. What is the gender of a typical Nobel Prize winner?
<p>So the USA became the dominating winner of the Nobel Prize first in the 1930s and had kept the leading position ever since. But one group that was in the lead from the start, and never seems to let go, are <em>men</em>. Maybe it shouldn't come as a shock that there is some imbalance between how many male and female prize winners there are, but how significant is this imbalance? And is it better or worse within specific prize categories like physics, medicine, literature, etc.?</p>
```
# Calculating the proportion of female laureates per decade
nobel['female_winner'] = nobel['sex']=='Female'
prop_female_winners = nobel.groupby(['decade','category'],as_index=False)['female_winner'].mean()
# Plotting Female winners
plt.rcParams['figure.figsize'] = [11, 7]
ax = sns.lineplot(x='decade',
y='female_winner',
data=prop_female_winners,
hue='category')
# Adding %-formatting to the y-axis
ax.yaxis.set_major_formatter(PercentFormatter(1.0))
```
## 6. The first woman to win the Nobel Prize
<p>The plot above is a bit messy as the lines are overplotting. But it does show some interesting trends and patterns. Overall the imbalance is pretty large with physics, economics, and chemistry having the largest imbalance. Medicine has a somewhat positive trend, and since the 1990s the literature prize is also now more balanced. The big outlier is the peace prize during the 2010s, but keep in mind that this just covers the years 2010 to 2016.</p>
<p>Given this imbalance, who was the first woman to receive a Nobel Prize? And in what category?</p>
```
# Picking out the first woman to win a Nobel Prize
nobel[nobel['female_winner']==True].nsmallest(1, 'year', keep='first')['full_name']
```
## 7. Repeat laureates
<p>For most scientists/writers/activists a Nobel Prize would be the crowning achievement of a long career. But for some people, one is just not enough, and few have gotten it more than once. Who are these lucky few? (Having won no Nobel Prize myself, I'll assume it's just about luck.)</p>
```
# Selecting the laureates that have received 2 or more prizes.
nobel.groupby('full_name').filter(lambda group: len(group) >= 2)
```
## 8. How old are you when you get the prize?
<p>The list of repeat winners contains some illustrious names! We again meet Marie Curie, who got the prize in physics for discovering radiation and in chemistry for isolating radium and polonium. John Bardeen got it twice in physics for transistors and superconductivity, Frederick Sanger got it twice in chemistry, and Linus Carl Pauling got it first in chemistry and later in peace for his work in promoting nuclear disarmament. We also learn that organizations also get the prize as both the Red Cross and the UNHCR have gotten it twice.</p>
<p>But how old are you generally when you get the prize?</p>
```
# Converting birth_date from String to datetime
nobel['birth_date'] = pd.to_datetime(nobel['birth_date'])
# Calculating the age of Nobel Prize winners
nobel['age'] = nobel['year'] - nobel['birth_date'].dt.year
# Plotting the age of Nobel Prize winners
sns.lmplot(x='year', y='age', data=nobel, lowess=True,
aspect=2, line_kws={'color' : 'black'})
```
## 9. Age differences between prize categories
<p>The plot above shows us a lot! We see that people use to be around 55 when they received the price, but nowadays the average is closer to 65. But there is a large spread in the laureates' ages, and while most are 50+, some are very young.</p>
<p>We also see that the density of points is much high nowadays than in the early 1900s -- nowadays many more of the prizes are shared, and so there are many more winners. We also see that there was a disruption in awarded prizes around the Second World War (1939 - 1945). </p>
<p>Let's look at age trends within different prize categories.</p>
```
# Same plot as above, but separate plots for each type of Nobel Prize
sns.lmplot(data=nobel, x='year', y='age', row='category', lowess=True,
aspect=2, line_kws={'color' : 'black'})
```
## 10. Oldest and youngest winners
<p>More plots with lots of exciting stuff going on! We see that both winners of the chemistry, medicine, and physics prize have gotten older over time. The trend is strongest for physics: the average age used to be below 50, and now it's almost 70. Literature and economics are more stable. We also see that economics is a newer category. But peace shows an opposite trend where winners are getting younger! </p>
<p>In the peace category we also a winner around 2010 that seems exceptionally young. This begs the questions, who are the oldest and youngest people ever to have won a Nobel Prize?</p>
```
# The oldest winner of a Nobel Prize as of 2016
display(nobel.nlargest(1, 'age', keep='first')[['full_name','age']])
# The youngest winner of a Nobel Prize as of 2016
nobel.nsmallest(1, 'age', keep='first')[['full_name','age']]
```
## 11. You get a prize!
<p><img style="float: right;margin:20px 20px 20px 20px; max-width:200px" src="https://assets.datacamp.com/production/project_441/img/paint_nobel_prize.png"></p>
<p>Hey! You get a prize for making it to the very end of this notebook! It might not be a Nobel Prize, but I made it myself in paint so it should count for something. But don't despair, Leonid Hurwicz was 90 years old when he got his prize, so it might not be too late for you. Who knows.</p>
<p>Before you leave, what was again the name of the youngest winner ever who in 2014 got the prize for "[her] struggle against the suppression of children and young people and for the right of all children to education"?</p>
```
# The name of the youngest winner of the Nobel Prize as of 2016
youngest_winner = '...'
```
| github_jupyter |
# ibm_db.prepare()
## Purpose:
Send an SQL statement to a Db2 server or database to have it prepared for execution.
## Syntax:
`IBM_DBStatement ibm_db.prepare( IBM_DBConnection `*`connection,`*` string `*`SQLstatement`*` [, dictionary `*`options`*`] )`
## Parameters:
* __*connection*__ : A valid Db2 server or database connection.
* __*SQLstatement*__ : A valid, executable SQL statement (that may or may not contain one or more parameter markers).
* __options__ : A dictionary containing key-value pairs for the statement attributes that are to be set when the statement provided in the __*SQLstatement*__ parameter is prepared. Valid keys and values are:<p>
* `ibm_db.SQL_ATTR_CURSOR_TYPE` : Specifies the type of cursor that is to be used for processing result sets. Valid values are:
* `ibm_db.SQL_CURSOR_FORWARD_ONLY` : A forward only cursor should be used (i.e., a cursor that only scrolls forward).
* `ibm_db. SQL_CURSOR_KEYSET_DRIVEN` : A keyset-driven cursor should be used.
* `ibm_db.SQL_CURSOR_STATIC` : A static cursor should be used.
* `ibm_db.SQL_CURSOR_DYNAMIC` : A dynamic, scrollable cursor that detects all changes to the result set, should be used. Dynamic cursors are only supported by Db2 for z/OS servers.
* `ibm_db.SQL_ATTR_ROWCOUNT_PREFETCH` : Enables Db2 to determine the number of rows that are returned by a query (so the entire result set can be prefetched into memory, when possible); `ibm_db.SQL_ROWCOUNT_PREFETCH_ON` will turn this behavior __ON__ and `ibm_db.SQL_ROWCOUNT_PREFETCH_OFF` will turn it __OFF__.
* `ibm_db.SQL_ATTR_QUERY_TIMEOUT` : The number of seconds to wait for an SQL statement to execute before aborting and returning to the application. This option can be used to terminate long running queries — the default value of __0__ means an application will wait indefinitely for the server to complete execution.<p>
## Return values:
* If __successful__, a valid `IBM_DBStatement` object.
* If __unsuccessful__, the value `None`.
## Description:
The __ibm_db.prepare()__ API is used to send an SQL statement to a Db2 server or database to have it prepared for execution — once an SQL statement has been prepared, it can be submitted for execution multiple times without having to be re-prepared. This benefits applications in two ways:<p>
* __Improved performance__: when an SQL statement is prepared, the database server creates an optimized access plan for executing that statement. Subsequent executions of the prepared statement (using the __ibm_db.execute()__ or __ibm_db.execute_many()__ API) will reuse that access plan, eliminating the overhead of dynamically creating new access plans each time the statement is executed.<p>
* __Greater flexibility__: when an SQL statement is prepared, literal values hard-coded in the statement can be replaced with parameter markers — depicted by question mark (?) characters. An SQL statement can contain zero or more parameter markers, each representing a variable or value that is to be provided at run time. Values for parameter markers can be supplied via variables (using the __ibm_db.bind_param()__ API), or for input parameters only, in a tuple (if the __ibm_db.execute()__ API is used) or a tuple of tuples (if the __ibm_db.execute_many()__ API is used). Regardless of which method used, when the prepared statement is executed, the database server will check each input value supplied to ensure the appropriate data type is used.<p>
It is important to note that parameter markers are only allowed in certain places in an SQL statement. For example, they cannot be used to represent one or more columns in the result set reurned by a __SELECT__ statement, nor can they be used as the operand of a binary operator like the equal sign (=).
## Example:
```
#----------------------------------------------------------------------------------------------#
# NAME: ibm_db-prepare.py #
# #
# PURPOSE: This program is designed to illustrate how to use the ibm_db.prepare() API. #
# #
# Additional APIs used: #
# ibm_db.execute() #
# ibm_db.fetch_assoc() #
# #
#----------------------------------------------------------------------------------------------#
# DISCLAIMER OF WARRANTIES AND LIMITATION OF LIABILITY #
# #
# (C) COPYRIGHT International Business Machines Corp. 2018, 2019 All Rights Reserved #
# Licensed Materials - Property of IBM #
# #
# US Government Users Restricted Rights - Use, duplication or disclosure restricted by GSA #
# ADP Schedule Contract with IBM Corp. #
# #
# The following source code ("Sample") is owned by International Business Machines #
# Corporation ("IBM") or one of its subsidiaries and is copyrighted and licensed, not sold. #
# You may use, copy, modify, and distribute the Sample in any form without payment to IBM, #
# for the purpose of assisting you in the creation of Python applications using the ibm_db #
# library. #
# #
# The Sample code is provided to you on an "AS IS" basis, without warranty of any kind. IBM #
# HEREBY EXPRESSLY DISCLAIMS ALL WARRANTIES, EITHER EXPRESS OR IMPLIED, INCLUDING, BUT NOT #
# LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE. #
# Some jurisdictions do not allow for the exclusion or limitation of implied warranties, so #
# the above limitations or exclusions may not apply to you. IBM shall not be liable for any #
# damages you suffer as a result of using, copying, modifying or distributing the Sample, #
# even if IBM has been advised of the possibility of such damages. #
#----------------------------------------------------------------------------------------------#
# Load The Appropriate Python Modules
import sys # Provides Information About Python Interpreter Constants And Functions
import ibm_db # Contains The APIs Needed To Work With Db2 Databases
#----------------------------------------------------------------------------------------------#
# Import The Db2ConnectionMgr Class Definition, Attributes, And Methods That Have Been Defined #
# In The File Named "ibm_db_tools.py"; This Class Contains The Programming Logic Needed To #
# Establish And Terminate A Connection To A Db2 Server Or Database #
#----------------------------------------------------------------------------------------------#
from ibm_db_tools import Db2ConnectionMgr
#----------------------------------------------------------------------------------------------#
# Import The ipynb_exit Class Definition, Attributes, And Methods That Have Been Defined In #
# The File Named "ipynb_exit.py"; This Class Contains The Programming Logic Needed To Allow #
# "exit()" Functionality To Work Without Raising An Error Or Stopping The Kernel If The #
# Application Is Invoked In A Jupyter Notebook #
#----------------------------------------------------------------------------------------------#
from ipynb_exit import exit
# Define And Initialize The Appropriate Variables
dbName = "SAMPLE"
userID = "db2inst1"
passWord = "Passw0rd"
dbConnection = None
preparedStmt = False
returnCode = False
dataRecord = None
# Create An Instance Of The Db2ConnectionMgr Class And Use It To Connect To A Db2 Database
conn = Db2ConnectionMgr('DB', dbName, '', '', userID, passWord)
conn.openConnection()
if conn.returnCode is True:
dbConnection = conn.connectionID
else:
conn.closeConnection()
exit(-1)
# Define The SQL Statement That Is To Be Executed
sqlStatement = "SELECT lastname, firstnme FROM employee WHERE job = 'PRES'"
# Prepare The SQL Statement Just Defined
print("Preparing the SQL statement \"" + sqlStatement + "\" ... ", end="")
try:
preparedStmt = ibm_db.prepare(dbConnection, sqlStatement)
except Exception:
pass
# If The SQL Statement Could Not Be Prepared By Db2, Display An Error Message And Exit
if preparedStmt is False:
print("\nERROR: Unable to prepare the SQL statement specified.")
conn.closeConnection()
exit(-1)
# Otherwise, Complete The Status Message
else:
print("Done!\n")
# Execute The SQL Statement Just Prepared
print("Executing the prepared SQL statement ... ", end="")
try:
returnCode = ibm_db.execute(preparedStmt)
except Exception:
pass
# If The SQL Statement Could Not Be Executed, Display An Error Message And Exit
if returnCode is False:
print("\nERROR: Unable to execute the SQL statement specified.")
conn.closeConnection()
exit(-1)
# Otherwise, Complete The Status Message
else:
print("Done!\n")
# Retrieve The Data Produced By The SQL Statement And Store It In A Python Dictionary
try:
dataRecord = ibm_db.fetch_assoc(preparedStmt)
except:
pass
# If The Data Could Not Be Retrieved, Display An Error Message And Exit
if returnCode is False:
print("\nERROR: Unable to retrieve the data produced by the SQL statement.")
conn.closeConnection()
exit(-1)
# If The Data Could be Retrieved, Display It
else:
print("Query results:\n")
print("FIRSTNME LASTNAME")
print("__________ ________")
print("{:10} {:<24}\n" .format(dataRecord['FIRSTNME'], dataRecord['LASTNAME']))
# Close The Database Connection That Was Opened Earlier
conn.closeConnection()
# Return Control To The Operating System
exit()
```
| github_jupyter |
# Calculate MI for each unit/language
1. load datasets
2. calculate MI
```
import pandas as pd
import numpy as np
from parallelspaper.config.paths import DATA_DIR, FIGURE_DIR
from parallelspaper.speech_datasets import LCOL_DICT
from parallelspaper import information_theory as it
from parallelspaper.quickplots import plot_model_fits
from tqdm.autonotebook import tqdm
from parallelspaper import model_fitting as mf
from parallelspaper.utils import save_fig
import matplotlib.pyplot as plt
%matplotlib inline
MI_DF = pd.read_pickle((DATA_DIR / 'MI_DF/language/language_MI_DF_fitted-utterance.pickle'))
```
### Plot shuffling analysis within vs between for utterances in japanese and english
```
fontsize=18
yoff=-.20
ncol = 4
nrow = len(MI_DF)//ncol
zoom = 5
fig, axs = plt.subplots(ncols=ncol, nrows=nrow, figsize=zoom*np.array([ncol,nrow]))
for axi, (idx, row) in enumerate(MI_DF.sort_values(by=['analysis', 'language', 'unit']).iterrows()):
ax = axs.flatten()[axi]
color = LCOL_DICT[row.language]
sig = np.array(row.MI-row.MI_shuff)
distances = row.distances
sig = sig
distances = distances
# get signal limits
sig_lims = np.log([np.min(sig[sig>0]), np.nanmax(sig)])
sig_lims = [sig_lims[0] - (sig_lims[1]-sig_lims[0])/10,
sig_lims[1] + (sig_lims[1]-sig_lims[0])/10]
if axi%ncol == 0:
ax.set_ylabel('Mutual Information (bits)', labelpad=5, fontsize=fontsize)
ax.yaxis.set_label_coords(yoff,0.5)
if axi >= (nrow-1)*ncol:
ax.set_xlabel('Distance (phones)', labelpad=5, fontsize=fontsize)
# plot real data
ax.scatter(distances, sig, alpha = 1, s=40, color=color)
best_fit_model = np.array(['exp','pow','pow_exp'])[np.argmin(row[['AICc_exp', 'AICc_power', 'AICc_concat']].values)]
# set title
analysis = 'within utt.' if row.analysis == 'shuffled_within_utterance' else 'between utt.'
model_type = {'pow_exp': 'comp.', 'exp': 'exp.', 'pow':'power law'}[best_fit_model]
ax.set_title(' | '.join([row.language.capitalize(), analysis, model_type]), fontsize=16)
# plot model
distances_model = np.logspace(0,np.log10(distances[-1]), base=10, num=1000)
if best_fit_model == 'pow_exp':
ax.axvline(distances_model[int(row.min_peak)], lw=3,alpha=0.5, color=color, ls='dashed')
if best_fit_model == 'pow_exp':
# model data
#row.concat_results.params.intercept = 0
y_model = mf.get_y(mf.pow_exp_decay, row.concat_results, distances_model)
y_pow = mf.get_y(mf.powerlaw_decay, row.concat_results, distances_model)
y_exp = mf.get_y(mf.exp_decay, row.concat_results, distances_model)
ax.plot(distances_model, y_pow, ls='dotted', color= 'k', lw=5, alpha=0.5)
ax.plot(distances_model, y_exp-row.concat_results.params['intercept'].value, ls='dashed', color= 'k', lw=5, alpha=0.5)
# plot modelled data
ax.plot(distances_model, y_model, alpha = 0.5, lw=10, color=color)
elif best_fit_model == 'pow':
y_model = mf.get_y(mf.powerlaw_decay, row.pow_results, distances_model)
# plot modelled data
ax.plot(distances_model, y_model, alpha = 0.5, lw=10, color=color)
elif best_fit_model == 'exp':
y_model = mf.get_y(mf.exp_decay, row.exp_results, distances_model)
# plot modelled data
ax.plot(distances_model, y_model, alpha = 0.5, lw=10, color=color)
# axis params
ax.set_xlim([distances[0], distances[-1]])
sig_lims[0] = np.log(10e-6)
ax.set_ylim(np.exp(sig_lims))
ax.tick_params(which='both', direction='in', labelsize=14, pad=10)
ax.tick_params(which='major', length=10, width =3)
ax.tick_params(which='minor', length=5, width =2)
ax.set_xscale( "log" , basex=10)
ax.set_yscale( "log" , basey=10)
ax.set_xticks([])
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(3)
ax.spines[axis].set_color('k')
ax.set_xlim([1,100])
ax.set_xticks([1,10,100])
ax.set_xticklabels(['1','10','100'])
save_fig(FIGURE_DIR/'speech_shuffle_utterance')
```
| github_jupyter |
```
%matplotlib inline
```
# Colormap reference
Reference for colormaps included with Matplotlib.
A reversed version of each of these colormaps is available by appending
``_r`` to the name, e.g., ``viridis_r``.
See :doc:`/tutorials/colors/colormaps` for an in-depth discussion about
colormaps, including colorblind-friendliness.
```
import numpy as np
import matplotlib.pyplot as plt
cmaps = [('Perceptually Uniform Sequential', [
'viridis', 'plasma', 'inferno', 'magma', 'cividis']),
('Sequential', [
'Greys', 'Purples', 'Blues', 'Greens', 'Oranges', 'Reds',
'YlOrBr', 'YlOrRd', 'OrRd', 'PuRd', 'RdPu', 'BuPu',
'GnBu', 'PuBu', 'YlGnBu', 'PuBuGn', 'BuGn', 'YlGn']),
('Sequential (2)', [
'binary', 'gist_yarg', 'gist_gray', 'gray', 'bone', 'pink',
'spring', 'summer', 'autumn', 'winter', 'cool', 'Wistia',
'hot', 'afmhot', 'gist_heat', 'copper']),
('Diverging', [
'PiYG', 'PRGn', 'BrBG', 'PuOr', 'RdGy', 'RdBu',
'RdYlBu', 'RdYlGn', 'Spectral', 'coolwarm', 'bwr', 'seismic']),
('Cyclic', ['twilight', 'twilight_shifted', 'hsv']),
('Qualitative', [
'Pastel1', 'Pastel2', 'Paired', 'Accent',
'Dark2', 'Set1', 'Set2', 'Set3',
'tab10', 'tab20', 'tab20b', 'tab20c']),
('Miscellaneous', [
'flag', 'prism', 'ocean', 'gist_earth', 'terrain', 'gist_stern',
'gnuplot', 'gnuplot2', 'CMRmap', 'cubehelix', 'brg',
'gist_rainbow', 'rainbow', 'jet', 'nipy_spectral', 'gist_ncar'])]
gradient = np.linspace(0, 1, 256)
gradient = np.vstack((gradient, gradient))
def plot_color_gradients(cmap_category, cmap_list):
# Create figure and adjust figure height to number of colormaps
nrows = len(cmap_list)
figh = 0.35 + 0.15 + (nrows + (nrows-1)*0.1)*0.22
fig, axs = plt.subplots(nrows=nrows, figsize=(6.4, figh))
fig.subplots_adjust(top=1-.35/figh, bottom=.15/figh, left=0.2, right=0.99)
axs[0].set_title(cmap_category + ' colormaps', fontsize=14)
for ax, name in zip(axs, cmap_list):
ax.imshow(gradient, aspect='auto', cmap=plt.get_cmap(name))
ax.text(-.01, .5, name, va='center', ha='right', fontsize=10,
transform=ax.transAxes)
# Turn off *all* ticks & spines, not just the ones with colormaps.
for ax in axs:
ax.set_axis_off()
for cmap_category, cmap_list in cmaps:
plot_color_gradients(cmap_category, cmap_list)
plt.show()
```
------------
References
""""""""""
The use of the following functions, methods, classes and modules is shown
in this example:
```
import matplotlib
matplotlib.colors
matplotlib.axes.Axes.imshow
matplotlib.figure.Figure.text
matplotlib.axes.Axes.set_axis_off
```
| github_jupyter |
# Load Dataset and Pre-Processing
```
import re
import pandas as pd
from tqdm import tqdm_notebook as tqdm
import matplotlib.pyplot as plt
import torch
from torch.utils.tensorboard import SummaryWriter
```
# Read Files and Fill NaN
```
lines_path = 'data/movie_lines.txt'
convs_path = 'data/movie_conversations.txt'
lines_df = pd.read_csv(lines_path,
sep='\s*\+\+\+\$\+\+\+\s*',
engine='python',
header=None,
index_col=0)
lines_df.head()
lines_df.isna().sum()
lines_df.iloc[:, -1].fillna('', inplace=True)
lines_df.isna().sum()
convs_df = pd.read_csv(convs_path,
sep='\s*\+\+\+\$\+\+\+\s*',
engine='python',
header=None)
convs_df.head()
convs_df.isna().sum()
```
# Obtain Conversations
```
convs_data = []
for line_ids in tqdm(convs_df.iloc[:, -1], desc='Build'):
conv = []
for line_id in eval(line_ids):
conv.append(lines_df.loc[line_id].iloc[-1])
convs_data.append(conv)
```
## (Check Length of Conversations)
```
lens = [len(conv) for conv in convs_data]
plt.hist(lens, bins=30)
plt.ylim(0, 100)
plt.show()
```
# Clean Each Sentence and Tokenize
```
def clean(sent):
sent = sent.lower().strip()
sent = re.sub(r'[^a-zA-Z.!?]+', r' ', sent)
sent = re.sub(r'([.!?])\1*', r' \1', sent)
sent = re.sub(r'\s+', r' ', sent)
sent = sent.strip().split()
return sent
convs = []
for conv in tqdm(convs_data, desc='Clean'):
convs.append(list(map(clean, conv)))
```
## (Check Length of Each Sentence )
```
lens = []
for conv in convs:
for sent in conv:
lens.append(len(sent))
plt.hist(lens, bins=50)
plt.ylim(0, 5000)
plt.show()
lens_short = [x for x in lens if x <= 30]
print(len(lens_short), len(lens), len(lens_short) / len(lens))
```
# Make Sentence Pairs and Filter
```
pairs = []
for conv in convs:
for i in range(len(conv) - 1):
pairs.append((conv[i], conv[i + 1]))
print(len(pairs))
def filter_pairs(pairs, min_length):
pairs_short = []
for a, b in pairs:
if len(a) > 0 and len(a) <= MIN_LENGTH \
and len(b) > 0 and len(b) <= MIN_LENGTH:
pairs_short.append((a, b))
return pairs_short
MIN_LENGTH = 30
pairs = filter_pairs(pairs, MIN_LENGTH)
print(len(pairs))
print(pairs[0][0])
print(pairs[0][1])
convs_data[0]
```
# Make Vocabulary
```
class Vocab:
def __init__(self):
self.special_tokens = ['<pad>', '<eos>', '<unk>']
self.word_count = {}
self.index2word = []
self.word2index = {}
def add_word(self, word):
if word in self.word_count:
self.word_count[word] += 1
else:
self.word_count[word] = 1
def make_vocab(self, size):
self.word_count = sorted(self.word_count, key=self.word_count.get, reverse=True)
for w in self.special_tokens + self.word_count[:size]:
self.index2word.append(w)
self.word2index[w] = len(self.index2word) - 1
def __len__(self):
return len(self.index2word)
def __getitem__(self, query):
if isinstance(query, int):
return self.index2word[query]
if isinstance(query, str):
if query in self.word2index:
return self.word2index[query]
else:
return self.word2index['<unk>']
```
## (Test Vocab)
```
a = ['a', 'boy', 'cow', 'and', 'a', 'boy', 'a']
vocab = Vocab()
for word in a:
vocab.add_word(word)
print(vocab.word_count)
vocab.make_vocab(size=3)
print(vocab.index2word)
print(vocab.word2index)
print(len(vocab))
print(vocab[4])
print(vocab['<eos>'])
print(vocab['abc'])
```
# Build Vocabuary
```
vocab = Vocab()
for i in range(len(pairs) - 1):
if pairs[i][1] == pairs[i + 1][0]:
words = pairs[i][0]
else:
words = pairs[i][0] + pairs[i][1]
for word in words:
vocab.add_word(word)
print(len(vocab.word_count))
```
## (Check Word Count)
```
a = sorted(vocab.word_count.values(), reverse=True)
plt.plot(a)
plt.ylim(0, 50)
plt.show()
print(a[10000])
VOCAB_SIZE = 10000
vocab.make_vocab(size=VOCAB_SIZE)
len(vocab)
```
# Build Dataset
```
class Dataset(torch.utils.data.Dataset):
def __init__(self, pairs, vocab):
self.pairs = pairs
self.vocab = vocab
def __getitem__(self, index):
x, y = self.pairs[index]
x = [vocab[w] for w in x]
y = [vocab[w] for w in y] + [vocab['<eos>']]
return x, y
def __len__(self):
return len(self.pairs)
dataset = Dataset(pairs, vocab)
print(len(dataset))
print(pairs[0][0])
print(dataset[0][0])
print(pairs[0][1])
print(dataset[0][1])
print(vocab.index2word[:20])
```
# Collate Function
```
def collate_fn(batch):
batch_x, batch_y = [], []
for x, y in batch:
batch_x.append(torch.tensor(x))
batch_y.append(torch.tensor(y))
batch_x = torch.nn.utils.rnn.pad_sequence(batch_x, padding_value=0)
batch_y = torch.nn.utils.rnn.pad_sequence(batch_y, padding_value=0)
return batch_x, batch_y
batch = [
([1, 2, 3], [1, 2]),
([1], [1, 2, 3]),
]
batch_x, batch_y = collate_fn(batch)
print(batch_x)
print(batch_y)
x = batch_x
((x != 0).sum(dim=0) <= 2).int()
```
# DataLoader
```
data_loader = torch.utils.data.DataLoader(
dataset,
batch_size=4,
collate_fn=collate_fn,
)
a = next(iter(data_loader))
print(a[0].shape)
print(a[1].shape)
```
# Model
```
class Encoder(torch.nn.Module):
def __init__(self, embed, hidden_size, pad_value):
super(Encoder, self).__init__()
self.pad_value = pad_value
self.embed = embed
self.lstm = torch.nn.LSTM(
input_size=embed.embedding_dim,
hidden_size=hidden_size,
num_layers=1,
bidirectional=True,
)
def forward(self, x):
lengths = (x != self.pad_value).sum(dim=0)
x = self.embed(x)
x = torch.nn.utils.rnn.pack_padded_sequence(
x, lengths, enforce_sorted=False)
output, (h, c) = self.lstm(x)
output, _ = torch.nn.utils.rnn.pad_packed_sequence(
output, padding_value=self.pad_value)
return output, (h.mean(dim=0), c.mean(dim=0))
vocab_size = 10
embed_size = 5
batch_size = 4
hidden_size = 4
embed = torch.nn.Embedding(len(vocab), embed_size)
encoder = Encoder(embed, hidden_size, pad_value=0)
batch_x, batch_y = next(iter(data_loader))
contex, (h, c) = encoder(batch_x)
print(contex.shape, h.shape, c.shape)
class Decoder(torch.nn.Module):
def __init__(self, embed, hidden_size, start_token):
super(Decoder, self).__init__()
self.start_token = start_token
self.embed = embed
self.lstm_cell = torch.nn.LSTMCell(
embed.embedding_dim, hidden_size)
self.out = torch.nn.Sequential(
torch.nn.Linear(2 * hidden_size, hidden_size),
torch.nn.ReLU(),
torch.nn.Linear(hidden_size, embed.num_embeddings),
)
def forward(self, hidden, contex, y):
y_preds = []
h, c = hidden
x = torch.empty(*y.size()[1:], dtype=torch.long)
x = x.fill_(self.start_token)
for i in range(len(y)):
x = self.embed(x)
h, c = self.lstm_cell(x, (h, c))
y_pred = self.out(torch.cat([h, c], dim=1))
y_preds.append(y_pred)
x = y[i]
return torch.stack(y_preds)
class Seq2Seq(torch.nn.Module):
def __init__(self, vocab_size, embed_size, hidden_size,
pad_value, start_token):
super(Seq2Seq, self).__init__()
self.embed = torch.nn.Embedding(vocab_size, embed_size)
self.encoder = Encoder(embed, hidden_size, pad_value)
self.decoder = Decoder(embed, hidden_size, start_token)
def forward(self, x, y):
contex, (h, c) = self.encoder(x)
y_preds = self.decoder((h, c), contex, y)
return y_preds
class Trainer:
def __init__(self, model, train_dataloader, n_epoch, optim, tb_dir=None,
case_interval=None, vocab=None,
valid_dataloader=None, test_dataloader=None):
self.model = model
self.train_dataloader = train_dataloader
self.valid_dataloader = valid_dataloader
self.test_dataloader = test_dataloader
self.n_epoch = n_epoch
self.optim = optim
self.case_interval = case_interval
self.vocab = vocab
self.writer = SummaryWriter(tb_dir, flush_secs=1) if tb_dir else None
def loss_fn(self, input, target):
input = input.reshape(-1, input.size(-1))
target = target.reshape(-1)
loss = torch.nn.functional.cross_entropy(
input=input, target=target,
ignore_index=0, reduction='mean')
return loss
def batch2sents(self, batch):
sents = []
for data in batch.tolist():
for _ in range(data.count(0)):
data.remove(0)
if data == []:
data.append('<pad> ...')
sent = [self.vocab[x] for x in data]
sents.append(' '.join(sent))
return sents
def show_case(self, x, y, y_preds, step):
post = self.batch2sents(x.T)[1]
targ = self.batch2sents(y.T)[1]
pred = y_preds.argmax(dim=2)
pred = self.batch2sents(pred.T)[1]
texts = [
f'[Post] {post}',
f'[Targ] {targ}',
f'[Pred] {pred}'
]
self.writer.add_text('case', '\n\n'.join(texts), step)
def train_batch(self, batch, batch_idx):
x, y = batch
y_preds = self.model(x, y)
loss = self.loss_fn(input=y_preds, target=y)
self.model.zero_grad()
loss.backward()
self.optim.step()
if batch_idx % 10 == 0:
self.show_case(x, y, y_preds, batch_idx)
return {'loss': loss.item()}
def overfit_one_batch(self, n_step):
self.model.train()
batch = next(iter(self.train_dataloader))
pbar = tqdm(range(n_step), desc='Overfit')
for i in pbar:
state = self.train_batch(batch, i)
pbar.set_postfix(state)
if self.writer is not None:
self.writer.add_scalars('overfit', state, i)
def fit(self):
for epoch in tqdm(range(self.n_epoch), desc='Total'):
self.train_epoch(epoch)
if self.valid_dataloader is not None:
self.valid_epoch(epoch)
if self.test_dataloader is not None:
self.test_epoch(epoch)
def train_epoch(self, epoch):
self.model.train()
pbar = tqdm(self.train_dataloader, desc=f'Train Epoch {epoch}')
for idx, batch in enumerate(pbar):
state = self.train_batch(batch, idx)
pbar.set_postfix(state)
if self.writer is not None:
self.writer.add_scalars('train', state, idx)
train_dataloader = torch.utils.data.DataLoader(dataset,
batch_size=4,
collate_fn=collate_fn)
model = Seq2Seq(
vocab_size=len(vocab),
embed_size=100,
hidden_size=100,
pad_value=0,
start_token=1,
)
adam = torch.optim.Adam(model.parameters())
trainer = Trainer(
model=model,
train_dataloader=train_dataloader,
n_epoch=2,
optim=adam,
tb_dir='runs/train7',
case_interval=10,
vocab=vocab,
)
# trainer.overfit_one_batch(800)
trainer.fit()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
data = pd.read_csv('https://raw.githubusercontent.com/PacktWorkshops/The-Data-Analysis-Workshop/master/Chapter09/Datasets/energydata_complete.csv')
data.head()
data.isnull().sum()
df1 = data.rename(columns = {
'date' : 'date_time',
'Appliances' : 'a_energy',
'lights' : 'l_energy',
'T1' : 'kitchen_temp',
'RH_1' : 'kitchen_hum',
'T2' : 'liv_temp',
'RH_2' : 'liv_hum',
'T3' : 'laun_temp',
'RH_3' : 'laun_hum',
'T4' : 'off_temp',
'RH_4' : 'off_hum',
'T5' : 'bath_temp',
'RH_5' : 'bath_hum',
'T6' : 'out_b_temp',
'RH_6' : 'out_b_hum',
'T7' : 'iron_temp',
'RH_7' : 'iron_hum',
'T8' : 'teen_temp',
'RH_8' : 'teen_hum',
'T9' : 'par_temp',
'RH_9' : 'par_hum',
'T_out' : 'out_temp',
'Press_mm_hg' : 'out_press',
'RH_out' : 'out_hum',
'Windspeed' : 'wind',
'Visibility' : 'visibility',
'Tdewpoint' : 'dew_point',
'rv1' : 'rv1',
'rv2' : 'rv2'
})
df1.head()
df1.tail()
df1.describe()
lights_box = sns.boxplot(df1.l_energy)
l = [0, 10, 20, 30, 40, 50, 60, 70]
counts = []
for i in l:
a = (df1.l_energy == i).sum()
counts.append(a)
counts
lights = sns.barplot(x = l, y = counts)
lights.set_xlabel('Energy Consumed by Lights')
lights.set_ylabel('Number of Lights')
lights.set_title('Distribution of Energy Consumed by Lights')
((df1.l_energy == 0).sum() / (df1.shape[0])) * 100
new_data = df1
new_data.drop(['l_energy'], axis = 1, inplace = True)
new_data.head()
app_box = sns.boxplot(new_data.a_energy)
out = (new_data['a_energy'] > 200).sum()
out
(out/19735) * 100
out_e = (new_data['a_energy'] > 950).sum()
out_e
(out_e/19735) * 100
energy = new_data[(new_data['a_energy'] <= 200)]
energy.describe()
new_en = energy
new_en['date_time'] = pd.to_datetime(new_en.date_time, format = '%Y-%m-%d %H:%M:%S')
new_en.head()
new_en.insert(loc = 1, column = 'month', value = new_en.date_time.dt.month)
new_en.insert(loc = 2, column = 'day', value = (new_en.date_time.dt.dayofweek)+1)
new_en.head()
```
**Exercise 9.04: Visualizing the Dataset**
```
import plotly.graph_objs as go
app_date = go.Scatter(x = new_en.date_time, mode = "lines", y = new_en.a_energy)
layout = go.Layout(title = 'Appliance Energy Consumed by Date', xaxis = dict(title='Date'), yaxis = dict(title='Wh'))
fig = go.Figure(data = [app_date], layout = layout)
fig.show()
app_mon = new_en.groupby(by = ['month'], as_index = False)['a_energy'].sum()
app_mon
app_mon.sort_values(by = 'a_energy', ascending = False).head()
plt.subplots(figsize = (15, 6))
am = sns.barplot(app_mon.month, app_mon.a_energy)
plt.xlabel('Month')
plt.ylabel('Energy Consumed by Appliances')
plt.title('Total Energy Consumed by Appliances per Month')
plt.show()
```
| github_jupyter |
# Study of perfect weights recall
```
import pprint
import subprocess
import sys
sys.path.append('../')
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import matplotlib.gridspec as gridspec
from mpl_toolkits.axes_grid1 import make_axes_locatable
import seaborn as sns
%matplotlib inline
np.set_printoptions(suppress=True, precision=2)
sns.set(font_scale=2.0)
from network import Protocol, BCPNNFast, NetworkManager
from connectivity_functions import artificial_connectivity_matrix, create_artificial_manager
from analysis_functions import calculate_angle_from_history, calculate_winning_pattern_from_distances
from analysis_functions import calculate_patterns_timings, calculate_recall_success
from analysis_functions import calculate_recall_success_sequences
from plotting_functions import plot_weight_matrix
```
## An example
```
# Patterns parameters
hypercolumns = 4
minicolumns = 20
dt = 0.001
# Recall
n = 1
T_cue = 0.100
T_recall = 4.0
# Artificial matrix
beta = False
value = 3.0
inhibition = -1.0
extension = 1
decay_factor = 0.0
sequence_decay = 0.0
# Sequence structure
overlap = 3
number_of_sequences = 2
half_width = 3
# Build chain protocol
chain_protocol = Protocol()
units_to_overload = [i for i in range(overlap)]
sequences = chain_protocol.create_overload_chain(number_of_sequences, half_width, units_to_overload)
manager = create_artificial_manager(hypercolumns, minicolumns, sequences, value=value,
inhibition=inhibition,
extension=extension, decay_factor=decay_factor,
sequence_decay=sequence_decay,
dt=dt, BCPNNFast=BCPNNFast, NetworkManager=NetworkManager, ampa=True,
beta=beta, values_to_save=['o', 's', 'z_pre'])
plot_weight_matrix(manager.nn)
sequences
tau_z_pre = 0.150
manager.nn.tau_z_pre = tau_z_pre
manager.nn.sigma = 1.0
manager.nn.g_w = 1.0
manager.nn.G = 1.0
n = 1
successes = calculate_recall_success(manager, T_recall=T_recall, I_cue=3, T_cue=T_cue,
n=n, patterns_indexes=sequences[0])
angles = calculate_angle_from_history(manager)
winning_patterns = calculate_winning_pattern_from_distances(angles)
timings = calculate_patterns_timings(winning_patterns, manager.dt, remove=0.010)
timings
plt.imshow(angles, aspect='auto')
plt.grid()
plt.colorbar()
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
cmap = matplotlib.cm.terrain
norm = matplotlib.colors.Normalize(vmin=0, vmax=15)
for index, angle in enumerate(angles.T):
if index < 15:
if index == 6:
ax.plot(angle + 0.1, color=cmap(norm(index)), label='pattern =' + str(index))
else:
ax.plot(angle, color=cmap(norm(index)), label='pattern =' + str(index))
ax.legend();
plt.plot(winning_patterns)
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
cmap = matplotlib.cm.terrain
norm = matplotlib.colors.Normalize(vmin=0, vmax=15)
for index, angle in enumerate(angles.T):
if index < 15:
ax.plot(angle, color=cmap(norm(index)), label='pattern =' + str(index))
ax.set_xlim([2000, 3000])
ax.set_ylim([0.7, 0.71]);
s = manager.history['s']
o = manager.history['o']
z_pre = manager.history['z_pre']
start = 0
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(211)
ax.plot(o[start:, 6], label='o 6')
ax.plot(o[start:, 12], label='o 12')
ax.plot(z_pre[start:, 5], label='z_pre 5')
ax.plot(z_pre[start:, 11], label='z_pre 11')
ax.legend()
ax = fig.add_subplot(212)
ax.plot(s[start:, 6], label='s6')
ax.plot(s[start:, 12], label='s12')
ax.legend()
plt.show()
n = 1
successes = calculate_recall_success_sequences(manager, T_recall=T_recall, T_cue=T_cue, n=n, sequences=sequences)
print(successes)
```
## Real training
```
# Patterns parameters
hypercolumns = 4
minicolumns = 20
dt = 0.001
values_to_save = ['o', 'z_pre', 's']
# Recall
n = 10
T_cue = 0.100
T_recall = 4.0
# Protocol
training_time = 0.1
inter_sequence_interval = 1.0
inter_pulse_interval = 0.0
epochs = 3
# Build the network
tau_z_pre = 0.050
tau_p = 100.0
nn = BCPNNFast(hypercolumns, minicolumns, tau_z_pre=tau_z_pre, tau_p=tau_p)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# Build the protocol for training
overlap = 3
number_of_sequences = 2
half_width = 3
units_to_overload = [i for i in range(overlap)]
protocol = Protocol()
units_to_overload = [i for i in range(overlap)]
sequences = protocol.create_overload_chain(number_of_sequences, half_width, units_to_overload)
sequences = [sequences[1], sequences[0]]
protocol.cross_protocol(sequences, training_time=training_time,
inter_sequence_interval=inter_sequence_interval, epochs=epochs)
manager.run_network_protocol(protocol=protocol, verbose=True, reset=True, empty_history=False)
# manager.run_network_protocol(protocol=protocol, verbose=True, reset=True, empty_history=False)
# manager.run_network_protocol(protocol=protocol, verbose=True, reset=True, empty_history=False)
plot_weight_matrix(manager.nn, one_hypercolum=True)
plot_weight_matrix(manager.nn, ampa=True, one_hypercolum=True)
w = nn.w
print(w[6, 5])
print(w[12, 5])
print('-----')
print(w[6, 11])
print(w[12, 11])
print('------')
print('overlap')
print(w[6, 2])
print(w[12, 2])
print('-----')
print(w[6, 1])
print(w[12, 1])
print('----')
print(w[6, 0])
print(w[12, 0])
print('----')
print(w[6, minicolumns + 6])
print(w[12, minicolumns + 12])
print('----')
print(w[1, 0]) * hypercolumns
print(w[0, minicolumns]) * (hypercolumns - 1)
sequences
tau_z_pre = 0.050
manager.nn.tau_z_pre = tau_z_pre
manager.nn.sigma = 0.0
manager.nn.g_w = 5.0
manager.nn.g_w_ampa = 1.0
manager.nn.G = 1.0
if True:
for i in range(3):
for k in range(hypercolumns):
index_1 = k * minicolumns
print('------')
print(index_1)
print('-------')
for l in range(hypercolumns):
index_2 = l * minicolumns
print(index_2)
if True:
aux1 = nn.w[6 + index_1, i + index_2]
aux2 = nn.w[12 + index_1, i + index_2]
nn.w[12 + index_1, i + index_2] = aux1
nn.w[6 + index_1, i + index_2] = aux2
if True:
aux1 = nn.w_ampa[6 + index_1, i + index_2]
aux2 = nn.w_ampa[12 + index_1, i + index_2]
nn.w_ampa[12 + index_1, i + index_2] = aux1
nn.w_ampa[6 + index_1, i + index_2] = aux2
# nn.w[12, 2] = nn.w[6, 2]
#nn.w[12, 1] = nn.w[6, 1]
# nn.w[12, 0] = nn.w[6, 0]
n = 1
T_recall = 3.0
successes = calculate_recall_success(manager, T_recall=T_recall, I_cue=3, T_cue=T_cue,
n=n, patterns_indexes=sequences[0])
angles = calculate_angle_from_history(manager)
winning_patterns = calculate_winning_pattern_from_distances(angles)
timings = calculate_patterns_timings(winning_patterns, manager.dt, remove=0.010)
timings
plt.imshow(angles, aspect='auto')
plt.grid()
plt.colorbar()
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
ax.plot(nn.w[6, :minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w[12, :minicolumns:], '*-', markersize=15, label='12')
if False:
ax.plot(nn.w[6, minicolumns:2 * minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w[12, minicolumns:2 * minicolumns], '*-', markersize=15, label='12')
ax.axhline(0, ls='--', color='black')
ax.legend();
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
ax.plot(nn.w_ampa[6, :minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w_ampa[12, :minicolumns:], '*-', markersize=15, label='12')
if False:
ax.plot(nn.w_ampa[6, minicolumns:2 * minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w_ampa[12, minicolumns:2 * minicolumns], '*-', markersize=15, label='12')
ax.axhline(0, ls='--', color='black')
ax.legend();
# swap(w[6, 5] - w[12, 5])
# swap(w[6, 11] = w[12, 11])
# aux1 = w[12, 5]
# aux2 = w[12, 11]
# w[12, 5] = w[6, 5]
# w[12, 11] = w[6, 11]
# w[6, 5] = aux1
# w[6, 11] = aux2
for i in range(3):
for k in range(hypercolumns):
index_1 = k * minicolumns
for l in range(hypercolumns):
index_2 = l * minicolumns
if True:
aux1 = nn.w[6 + index_1, i + index_2]
nn.w[12 + index_1, i + index_2] = aux1
nn.w[6 + index_1, i + index_2] = aux1
if True:
aux1 = nn.w_ampa[6 + index_1, i + index_2]
nn.w_ampa[12 + index_1, i + index_2] = aux1
nn.w_ampa[6 + index_1, i + index_2] = aux1
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
ax.plot(nn.w[6, :minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w[12, :minicolumns:], '*-', markersize=15, label='12')
if False:
ax.plot(nn.w[6, minicolumns:2 * minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w[12, minicolumns:2 * minicolumns], '*-', markersize=15, label='12')
ax.axhline(0, ls='--', color='black')
ax.legend();
tau_z_pre = 0.050
manager.nn.tau_z_pre = tau_z_pre
manager.nn.sigma = 0.0
manager.nn.g_w = 5.0
manager.nn.g_w_ampa = 1.0
manager.nn.G = 1.0
n = 1
T_recall = 3.0
successes = calculate_recall_success(manager, T_recall=T_recall, I_cue=3, T_cue=T_cue,
n=n, patterns_indexes=sequences[0])
angles = calculate_angle_from_history(manager)
winning_patterns = calculate_winning_pattern_from_distances(angles)
timings = calculate_patterns_timings(winning_patterns, manager.dt, remove=0.010)
timings
plt.imshow(angles, aspect='auto')
plt.grid()
plt.colorbar()
sequences
past_list = [(5, 11), (4, 10), (3, 9)]
for past1, past2 in past_list:
for k in range(hypercolumns):
index_1 = k * minicolumns
for l in range(hypercolumns):
index_2 = l * minicolumns
if False:
aux1 = nn.w[12 + index_1, past1 + index_2]
aux2 = nn.w[12 + index_1, past2 + index_2]
nn.w[12 + index_1, past1 + index_1] = nn.w[6 + index_1, past1 + index_2]
nn.w[12 + index_1, past2 + index_1] = nn.w[6 + index_1, past2 + index_2]
nn.w[6 + index_1, past1 + index_2] = aux1
nn.w[6 + index_1, past2 + index_2] = aux2
if True:
aux1 = nn.w_ampa[12 + index_1, past1 + index_2]
aux2 = nn.w_ampa[12 + index_1, past2 + index_2]
nn.w_ampa[12 + index_1, past1 + index_1] = nn.w_ampa[6 + index_1, past1 + index_2]
nn.w_ampa[12 + index_1, past2 + index_1] = nn.w_ampa[6 + index_1, past2 + index_2]
nn.w_ampa[6 + index_1, past1 + index_2] = aux1
nn.w_ampa[6 + index_1, past2 + index_2] = aux2
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
ax.plot(nn.w[6, :minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w[12, :minicolumns:], '*-', markersize=15, label='12')
if False:
ax.plot(nn.w[6, minicolumns:2 * minicolumns], '*-', markersize=15, label='6')
ax.plot(nn.w[12, minicolumns:2 * minicolumns], '*-', markersize=15, label='12')
ax.axhline(0, ls='--', color='black')
ax.legend();
tau_z_pre = 0.250
manager.nn.tau_z_pre = tau_z_pre
manager.nn.sigma = 0.0
manager.nn.g_w = 5.0
manager.nn.g_w_ampa = 1.0
manager.nn.G = 1.0
n = 1
T_recall = 3.0
successes = calculate_recall_success(manager, T_recall=T_recall, I_cue=3, T_cue=T_cue,
n=n, patterns_indexes=sequences[0])
angles = calculate_angle_from_history(manager)
winning_patterns = calculate_winning_pattern_from_distances(angles)
timings = calculate_patterns_timings(winning_patterns, manager.dt, remove=0.010)
timings
plt.imshow(angles, aspect='auto')
plt.grid()
plt.colorbar()
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
cmap = matplotlib.cm.terrain
norm = matplotlib.colors.Normalize(vmin=0, vmax=15)
for index, angle in enumerate(angles.T):
if index < 15:
if index == 6:
ax.plot(angle, color=cmap(norm(index)), label='pattern =' + str(index))
else:
ax.plot(angle, color=cmap(norm(index)), label='pattern =' + str(index))
ax.legend();
s = manager.history['s']
o = manager.history['o']
z_pre = manager.history['z_pre']
start = 0
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(211)
ax.plot(o[start:, 6], label='o 6')
ax.plot(o[start:, 12], label='o 12')
ax.plot(z_pre[start:, 5], label='z_pre 5')
ax.plot(z_pre[start:, 11], label='z_pre 11')
ax.legend()
ax = fig.add_subplot(212)
ax.plot(s[start:, 6], label='s6')
ax.plot(s[start:, 12], label='s12')
ax.legend()
plt.show()
n = 20
manager.nn_w_ampa = 0.0
manager.nn.tau_z_pre = 0.010
successes = calculate_recall_success_sequences(manager, T_recall=T_recall, T_cue=T_cue, n=n, sequences=sequences)
successes
```
| github_jupyter |
# T1078 - Valid Accounts
Adversaries may obtain and abuse credentials of existing accounts as a means of gaining Initial Access, Persistence, Privilege Escalation, or Defense Evasion. Compromised credentials may be used to bypass access controls placed on various resources on systems within the network and may even be used for persistent access to remote systems and externally available services, such as VPNs, Outlook Web Access and remote desktop. Compromised credentials may also grant an adversary increased privilege to specific systems or access to restricted areas of the network. Adversaries may choose not to use malware or tools in conjunction with the legitimate access those credentials provide to make it harder to detect their presence.
The overlap of permissions for local, domain, and cloud accounts across a network of systems is of concern because the adversary may be able to pivot across accounts and systems to reach a high level of access (i.e., domain or enterprise administrator) to bypass access controls set within the enterprise. (Citation: TechNet Credential Theft)
## Atomic Tests:
Currently, no tests are available for this technique.
## Detection
Configure robust, consistent account activity audit policies across the enterprise and with externally accessible services. (Citation: TechNet Audit Policy) Look for suspicious account behavior across systems that share accounts, either user, admin, or service accounts. Examples: one account logged into multiple systems simultaneously; multiple accounts logged into the same machine simultaneously; accounts logged in at odd times or outside of business hours. Activity may be from interactive login sessions or process ownership from accounts being used to execute binaries on a remote system as a particular account. Correlate other security systems with login information (e.g., a user has an active login session but has not entered the building or does not have VPN access).
Perform regular audits of domain and local system accounts to detect accounts that may have been created by an adversary for persistence. Checks on these accounts could also include whether default accounts such as Guest have been activated. These audits should also include checks on any appliances and applications for default credentials or SSH keys, and if any are discovered, they should be updated immediately.
## Shield Active Defense
### Decoy Account
Create an account that is used for active defense purposes.
A decoy account is one that is created specifically for defensive or deceptive purposes. It can be in the form of user accounts, service accounts, software accounts, etc. The decoy account can be used to make a system, service, or software look more realistic or to entice an action.
#### Opportunity
There is an opportunity to introduce user accounts that are used to make a system look more realistic.
#### Use Case
A defender can create decoy user accounts which are used to make a decoy system or network look more realistic.
#### Procedures
Create a user account with a specified job function. Populate the user account's groups, description, logon hours, etc., with decoy data that looks normal in the environment.
Create a user that has a valid email account. Use this account in such a way that the email address could be harvested by the adversary. This can be monitored to see if it is used in future attacks.
| github_jupyter |
NOTE: for the most up to date version of this notebook, please copy from
[](https://colab.research.google.com/drive/1ntAL_zI68xfvZ4uCSAF6XT27g0U4mZbW#scrollTo=VHS_o3KGIyXm)
## **Training YOLOv3 object detection on a custom dataset**
### **Overview**
This notebook walks through how to train a YOLOv3 object detection model on your own dataset using Roboflow and Colab.
In this specific example, we'll training an object detection model to recognize chess pieces in images. **To adapt this example to your own dataset, you only need to change one line of code in this notebook.**

### **Our Data**
Our dataset of 289 chess images (and 2894 annotations!) is hosted publicly on Roboflow [here](https://public.roboflow.ai/object-detection/chess-full).
### **Our Model**
We'll be training a YOLOv3 (You Only Look Once) model. This specific model is a one-shot learner, meaning each image only passes through the network once to make a prediction, which allows the architecture to be very performant, viewing up to 60 frames per second in predicting against video feeds.
The GitHub repo containing the majority of the code we'll use is available [here](https://github.com/roboflow-ai/yolov3).
### **Training**
Google Colab provides free GPU resources. Click "Runtime" → "Change runtime type" → Hardware Accelerator dropdown to "GPU."
Colab does have memory limitations, and notebooks must be open in your browser to run. Sessions automatically clear themselves after 12 hours.
### **Inference**
We'll leverage the `detect.py --weights weights/last.pt` script to produce predictions. Arguments are specified below.
```
import os
import torch
from IPython.display import Image, clear_output
print('PyTorch %s %s' % (torch.__version__, torch.cuda.get_device_properties(0) if torch.cuda.is_available() else 'CPU'))
!git clone https://github.com/roboflow-ai/yolov3 # clone
%ls
```
## Get Data from Roboflow
Create an export from Roboflow. **Select "YOLO Darknet" as the export type.**
Our labels will be formatted to our model's architecture.
```
# REPLACE THIS LINK WITH YOUR OWN
!curl -L "https://public.roboflow.ai/ds/3103AzYDyI?key=0MAoJ714iA" > roboflow.zip; unzip roboflow.zip; rm roboflow.zip
```
## Organize data and labels for Ultralytics YOLOv3 Implementation
Ultalytics's implemention of YOLOv3 calls for [a specific file management](https://github.com/ultralytics/yolov3/wiki/Train-Custom-Data) where our images are in a folder called `images` and corresponding labels in a folder called `labels`. The image and label names must match identically. Fortunately, our files are named appropriately from Roboflow.
We need to reorganize the folder structure slightly.
```
%cd train
%ls
%mkdir labels
%mkdir images
%mv *.jpg ./images/
%mv *.txt ./labels/
%cd images
# create Ultralytics specific text file of training images
file = open("train_images_roboflow.txt", "w")
for root, dirs, files in os.walk("."):
for filename in files:
# print("../train/images/" + filename)
if filename == "train_images_roboflow.txt":
pass
else:
file.write("../train/images/" + filename + "\n")
file.close()
%cat train_images_roboflow.txt
%cd ../../valid
%mkdir labels
%mkdir images
%mv *.jpg ./images/
%mv *.txt ./labels/
%cd images
# create Ultralytics specific text file of validation images
file = open("valid_images_roboflow.txt", "w")
for root, dirs, files in os.walk("."):
for filename in files:
# print("../train/images/" + filename)
if filename == "valid_images_roboflow.txt":
pass
else:
file.write("../valid/images/" + filename + "\n")
file.close()
%cat valid_images_roboflow.txt
```
## Set up model config
We should configure our model for training.
This requires editing the `roboflow.data` file, which tells our model where to find our data, our numbers of classes, and our class label names.
Our paths for our labels and images are correct.
But we need to update our class names. That's handled below..
```
%cd ../../yolov3/data
# display class labels imported from Roboflow
%cat ../../train/_darknet.labels
# convert .labels to .names for Ultralytics specification
%cat ../../train/_darknet.labels > ../../train/roboflow_data.names
def get_num_classes(labels_file_path):
classes = 0
with open(labels_file_path, 'r') as f:
for line in f:
classes += 1
return classes
# update the roboflow.data file with correct number of classes
import re
num_classes = get_num_classes("../../train/_darknet.labels")
with open("roboflow.data") as f:
s = f.read()
with open("roboflow.data", 'w') as f:
# Set number of classes num_classes.
s = re.sub('classes=[0-9]+',
'classes={}'.format(num_classes), s)
f.write(s)
# display updated number of classes
%cat roboflow.data
```
## Training our model
Once we have our data prepped, we'll train our model using the train script.
By default, this script trains for 300 epochs.
```
%cd ../
!python3 train.py --data data/roboflow.data --epochs 300
```
## Display training performance
We'll use a default provided script to display image results. **For example:**

```
from utils import utils; utils.plot_results()
```
## Conduct inference and display results
### Conduct inference
The below script has a few key arguments we're using:
- **Weights**: we're specifying the weights to use for our model should be those that we most recently used in training
- **Source**: we're specifying the source images we want to use for our predictions
- **Names**: we're defining the names we want to use. Here, we're referencing `roboflow_data.names`, which we created from our Roboflow `_darknet.labels`italicized text above.
```
!python3 detect.py --weights weights/last.pt --source='/content/gdrive/My Drive/chess_practice_yolo/chess_yolo_v3_pytorch/dataset/train/0115e4df73475b550e5c6f7a88b2474f_jpg.rf.dfa577bd4af5440d689046c2f48bc48e.jpg' --names=../train/roboflow_data.names
```
### Displaying our results
Ultralytics generates predictions which include the labels and bounding boxes "printed" directly on top of our images. They're saved in our `output` directory within the YOLOv3 repo we cloned above.
```
# import libraries for display
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import display, Image
from glob import glob
import random
import PIL
# plot just one random image prediction
filename = random.choice(os.listdir('./output'))
print(filename)
Image('./output/' + filename)
# grab all images from our output directory
images = [ PIL.Image.open(f) for f in glob('./output/*') ]
# convert images to numPy
def img2array(im):
if im.mode != 'RGB':
im = im.convert(mode='RGB')
return np.fromstring(im.tobytes(), dtype='uint8').reshape((im.size[1], im.size[0], 3))
# create array of numPy images
np_images = [ img2array(im) for im in images ]
# plot ALL results in test directory (NOTE: adjust figsize as you please)
for img in np_images:
plt.figure(figsize=(8, 6))
plt.imshow(img)
```
## Save Our Weights
We can save the weights of our model to use them for inference in the future, or pick up training where we left off.
We can first save them locally. We'll connect our Google Drive, and save them there.
```
# save locally
from google.colab import files
files.download('./weights/last.pt')
# connect Google Drive
from google.colab import drive
drive.mount('/content/gdrive')
%pwd
# create a copy of the weights file with a datetime
# and move that file to your own Drive
%cp ./weights/last.pt ./weights/last_copy.pt
%mv ./weights/last_copy.pt /content/gdrive/My\ Drive/chess_practice_yolo/
```
| github_jupyter |
```
import SNPLIB
import math
import numpy as np
import numpy.linalg as npl
import matplotlib.pyplot as plt
from numpy import random
from scipy import stats
Fst = 0.1
num_generations = 200
effective_sample_size = math.floor(num_generations/2/(1-math.exp(-Fst)))
num_snps = 100000
num_causal_snps = 10000
num_samples = 4000
num_traits = 10000
num_pairs = num_traits//2
aaf = 0.05+0.9*np.random.rand(num_snps)
pop_af = SNPLIB.UpdateAf(aaf,2,num_generations,effective_sample_size)
pop = np.zeros((num_samples,2),dtype='double',order='F')
pop[:,0] = np.sort(np.random.beta(0.1,0.1,num_samples))
pop[:,1] = 1.0-pop[:,0]
af = pop@pop_af
obj = SNPLIB.SNPLIB()
obj.GenerateIndividuals(af)
geno_d = obj.UnpackGeno()*random.randn(1,num_snps)
"""
Same additive effects
Same direction
"""
true_genetic_corr = np.zeros(num_pairs)
true_env_corr = np.zeros(num_pairs)
sim_traits = np.zeros((num_samples, num_traits), dtype='double', order='F')
for i in range(num_pairs):
all_ind = np.arange(num_snps)
additive_snp = random.choice(all_ind,replace=False)
all_ind = np.setdiff1d(all_ind,additive_snp)
snp_ind1 = random.choice(all_ind,size=(num_causal_snps,1),replace=False).squeeze()
all_ind = np.setdiff1d(all_ind,snp_ind1)
num_shared_snps = random.randint(num_causal_snps)
snp_ind2 = random.choice(snp_ind1, size=(num_shared_snps,1),replace=False).squeeze()
snp_ind2 = np.concatenate((snp_ind2, random.choice(all_ind, size=(num_causal_snps-num_shared_snps,1),replace=False).squeeze()))
rho = random.rand(1)*2-1
env_effects = random.multivariate_normal([0.,0.],[[1,rho],[rho,1]],num_samples)
effects_1 = np.zeros((num_samples,3),dtype='double',order='F')
effects_1[:,0] = geno_d[:,additive_snp]
effects_1[:,1] = np.sum(geno_d[:,snp_ind1], axis=1)
effects_1[:,2] = env_effects[:,0]
effects_1 = stats.zscore(effects_1,axis=0,ddof=1)
effects_1,_ = npl.qr(effects_1)
effects_1 *= math.sqrt(num_samples-1)
effects_1[:,0] *= math.sqrt(0.02)
effects_1[:,1] *= 0.7
effects_1[:,2] *= 0.7
sim_traits[:,2*i] = np.sum(effects_1, axis=1);
effects_2 = np.zeros((num_samples,3),dtype='double',order='F')
effects_2[:,0] = geno_d[:,additive_snp]
effects_2[:,1] = np.sum(geno_d[:,snp_ind2], axis=1)
effects_2[:,2] = env_effects[:,1]
effects_2 = stats.zscore(effects_2,axis=0,ddof=1)
effects_2,_ = npl.qr(effects_2)
effects_2 *= math.sqrt(num_samples-1)
effects_2[:,0] *= math.sqrt(0.02)
effects_2[:,1] *= 0.7
effects_2[:,2] *= 0.7
sim_traits[:,2*i+1] = np.sum(effects_2, axis=1);
true_genetic_corr[i],_ = stats.pearsonr(effects_2[:,1],effects_1[:,1])
true_env_corr[i],_ = stats.pearsonr(effects_2[:,2],effects_1[:,2])
"""
Same additive effects
Opposite direction
"""
true_genetic_corr = np.zeros(num_pairs)
true_env_corr = np.zeros(num_pairs)
sim_traits = np.zeros((num_samples, num_traits), dtype='double', order='F')
for i in range(num_pairs):
all_ind = np.arange(num_snps)
additive_snp = random.choice(all_ind,replace=False)
all_ind = np.setdiff1d(all_ind,additive_snp)
snp_ind1 = random.choice(all_ind,size=(num_causal_snps,1),replace=False).squeeze()
all_ind = np.setdiff1d(all_ind,snp_ind1)
num_shared_snps = random.randint(num_causal_snps)
snp_ind2 = random.choice(snp_ind1, size=(num_shared_snps,1),replace=False).squeeze()
snp_ind2 = np.concatenate((snp_ind2, random.choice(all_ind, size=(num_causal_snps-num_shared_snps,1),replace=False).squeeze()))
rho = random.rand(1)*2-1
env_effects = random.multivariate_normal([0.,0.],[[1,rho],[rho,1]],num_samples)
effects_1 = np.zeros((num_samples,3),dtype='double',order='F')
effects_1[:,0] = geno_d[:,additive_snp]
effects_1[:,1] = np.sum(geno_d[:,snp_ind1], axis=1)
effects_1[:,2] = env_effects[:,0]
effects_1 = stats.zscore(effects_1,axis=0,ddof=1)
effects_1,_ = npl.qr(effects_1)
effects_1 *= math.sqrt(num_samples-1)
effects_1[:,0] *= math.sqrt(0.02)
effects_1[:,1] *= 0.7
effects_1[:,2] *= 0.7
sim_traits[:,2*i] = np.sum(effects_1, axis=1);
effects_2 = np.zeros((num_samples,3),dtype='double',order='F')
effects_2[:,0] = -geno_d[:,additive_snp]
effects_2[:,1] = np.sum(geno_d[:,snp_ind2], axis=1)
effects_2[:,2] = env_effects[:,1]
effects_2 = stats.zscore(effects_2,axis=0,ddof=1)
effects_2,_ = npl.qr(effects_2)
effects_2 *= math.sqrt(num_samples-1)
effects_2[:,0] *= math.sqrt(0.02)
effects_2[:,1] *= 0.7
effects_2[:,2] *= 0.7
sim_traits[:,2*i+1] = np.sum(effects_2, axis=1);
true_genetic_corr[i],_ = stats.pearsonr(effects_2[:,1],effects_1[:,1])
true_env_corr[i],_ = stats.pearsonr(effects_2[:,2],effects_1[:,2])
"""
Difference additive effects
"""
true_genetic_corr = np.zeros(num_pairs)
true_env_corr = np.zeros(num_pairs)
sim_traits = np.zeros((num_samples, num_traits), dtype='double', order='F')
for i in range(num_pairs):
all_ind = np.arange(num_snps)
additive_snp = random.choice(all_ind,replace=False)
all_ind = np.setdiff1d(all_ind,additive_snp)
snp_ind1 = random.choice(all_ind,size=(num_causal_snps,1),replace=False).squeeze()
all_ind = np.setdiff1d(all_ind,snp_ind1)
num_shared_snps = random.randint(num_causal_snps)
snp_ind2 = random.choice(snp_ind1, size=(num_shared_snps,1),replace=False).squeeze()
snp_ind2 = np.concatenate((snp_ind2, random.choice(all_ind, size=(num_causal_snps-num_shared_snps,1),replace=False).squeeze()))
rho = random.rand(1)*2-1
env_effects = random.multivariate_normal([0.,0.],[[1,rho],[rho,1]],num_samples)
effects_1 = np.zeros((num_samples,3),dtype='double',order='F')
effects_1[:,0] = geno_d[:,additive_snp]
effects_1[:,1] = np.sum(geno_d[:,snp_ind1], axis=1)
effects_1[:,2] = env_effects[:,0]
effects_1 = stats.zscore(effects_1,axis=0,ddof=1)
effects_1,_ = npl.qr(effects_1)
effects_1 *= math.sqrt(num_samples-1)
effects_1[:,0] *= math.sqrt(0.02)
effects_1[:,1] *= 0.7
effects_1[:,2] *= 0.7
sim_traits[:,2*i] = np.sum(effects_1, axis=1);
effects_2 = np.zeros((num_samples,3),dtype='double',order='F')
effects_2[:,0] = geno_d[:,additive_snp]
effects_2[:,1] = np.sum(geno_d[:,snp_ind2], axis=1)
effects_2[:,2] = env_effects[:,1]
effects_2 = stats.zscore(effects_2,axis=0,ddof=1)
effects_2,_ = npl.qr(effects_2)
effects_2 *= math.sqrt(num_samples-1)
effects_2[:,0] *= math.sqrt(0.02)
effects_2[:,1] *= 0.7
effects_2[:,2] *= 0.7
sim_traits[:,2*i+1] = np.sum(effects_2, axis=1);
true_genetic_corr[i],_ = stats.pearsonr(effects_2[:,1],effects_1[:,1])
true_env_corr[i],_ = stats.pearsonr(effects_2[:,2],effects_1[:,2])
```
### plt.plot(additive_effect2,'.')
plt.show()
```
rho = random.rand(1)*2-1
env_effects = random.multivariate_normal([0.,0.],[[1,rho],[rho,1]],num_samples)
effects_1 = np.zeros((num_samples,3),dtype='double',order='F')
effects_1[:,0] = geno_d[:,additive_snp]
effects_1[:,1] = np.sum(geno_d[:,snp_ind1], axis=1)
effects_1[:,2] = env_effects[:,0]
effects_1 = stats.zscore(effects_1,axis=0,ddof=1)
effects_1,_ = npl.qr(effects_1)
effects_1 *= math.sqrt(num_samples-1)
effects_1[:,0] *= math.sqrt(0.02)
effects_1[:,1] *= 0.7
effects_1[:,2] *= 0.7
a = np.sum(effects_1,axis=1)
np.std(a,ddof=1)
num_shared_snps
```
| github_jupyter |
# Map partition
```
from dask.distributed import Client, LocalCluster
cluster = LocalCluster(n_workers=4)
client = Client(cluster)
client
import dask.dataframe as dd
import dask.dataframe as dd
path = "../Chapter2/nyctaxi/part_parquet_stage1/year=2016/"
ddf = dd.read_parquet(path)
ddf.head()
ddf.dtypes.to_dict()
ddf.astype(ddf.dtypes.to_dict()).head()
dtypes = ddf.dtypes.to_dict()
df = ddf.get_partition(0).compute()
df[["pickup_latitude", "pickup_longitude",
"dropoff_latitude", "dropoff_longitude"]].head()
import numpy as np
import numpy as np
df = df.assign(
pu_lat_rad=np.deg2rad(df["pickup_latitude"].values),
pu_lon_rad=np.deg2rad(df["pickup_longitude"].values),
do_lat_rad=np.deg2rad(df["dropoff_latitude"].values),
do_lon_rad=np.deg2rad(df["dropoff_longitude"].values))
df = df.assign(
pu_lat_rad=np.deg2rad(df["pickup_latitude"].values),
pu_lon_rad=np.deg2rad(df["pickup_longitude"].values),
do_lat_rad=np.deg2rad(df["dropoff_latitude"].values),
do_lon_rad=np.deg2rad(df["dropoff_longitude"].values))
meta_1 = df.dtypes.to_dict()
meta_1
df[df.columns[-4:]].head()
df = df.assign(
pu_x=np.cos(df["pu_lat_rad"].values) *
np.cos(df["pu_lon_rad"].values),
pu_y=np.cos(df["pu_lat_rad"].values) *
np.sin(df["pu_lon_rad"].values),
pu_z=np.sin(df["pu_lat_rad"].values),
do_x=np.cos(df["do_lat_rad"].values) *
np.cos(df["do_lon_rad"].values),
do_y=np.cos(df["do_lat_rad"].values) *
np.sin(df["do_lon_rad"].values),
do_z=np.sin(df["do_lat_rad"].values)
)
meta2 = df.dtypes.to_dict()
meta = df.dtypes.to_dict()
def lat_lon_to_cartesian(df):
df = df.assign(
pu_lat_rad=np.deg2rad(df["pickup_latitude"].values),
pu_lon_rad=np.deg2rad(df["pickup_longitude"].values),
do_lat_rad=np.deg2rad(df["dropoff_latitude"].values),
do_lon_rad=np.deg2rad(df["dropoff_longitude"].values)
)
df = df.assign(
pu_x=np.cos(df["pu_lat_rad"].values) *
np.cos(df["pu_lon_rad"].values),
pu_y=np.cos(df["pu_lat_rad"].values) *
np.sin(df["pu_lon_rad"].values),
pu_z=np.sin(df["pu_lat_rad"].values),
do_x=np.cos(df["do_lat_rad"].values) *
np.cos(df["do_lon_rad"].values),
do_y=np.cos(df["do_lat_rad"].values) *
np.sin(df["do_lon_rad"].values),
do_z=np.sin(df["do_lat_rad"].values)
)
return df
df = ddf.get_partition(0).compute()
lat_lon_to_cartesian(df)
ddf = ddf.map_partitions(lambda df:
lat_lon_to_cartesian(df),
meta=meta)
%%time
path = "2016nyc_taxi_cartesian/"
ddf.to_parquet(path, write_index=False)
```
# Data Analisys
```
from dask.distributed import Client, LocalCluster
cluster = LocalCluster(n_workers=4)
client = Client(cluster)
client
import dask.dataframe as dd
cols_to_read = ["VendorID", "passenger_count",
"trip_distance", "fare_amount",
"tip_amount", "holiday"]
path = "../Chapter2/nyctaxi/2018_trips_holidays/"
ddf = dd.read_parquet(path,
columns=cols_to_read)
ddf
ddf = ddf[ddf["passenger_count"].between(1, 11) &
ddf["trip_distance"].between(0.1, 150) &
ddf["fare_amount"].between(2.5, 375) &
ddf["tip_amount"].ge(0)]
%%time
amnt_by_vndr = ddf.groupby("VendorID")[["fare_amount",
"tip_amount"]].sum().compute()
amnt_by_vndr["percentage"] = amnt_by_vndr["tip_amount"]/\
amnt_by_vndr["fare_amount"]
amnt_by_vndr
tips_fare = ddf.groupby("holiday")[["fare_amount",
"tip_amount"]].sum().compute()
tips_fare["percentage"] = tips_fare["tip_amount"]/\
tips_fare["fare_amount"]
tips_fare
```
| github_jupyter |
# 911 Calls Capstone Project - Solutions
For this capstone project we will be analyzing some 911 call data from [Kaggle](https://www.kaggle.com/mchirico/montcoalert). The data contains the following fields:
* lat : String variable, Latitude
* lng: String variable, Longitude
* desc: String variable, Description of the Emergency Call
* zip: String variable, Zipcode
* title: String variable, Title
* timeStamp: String variable, YYYY-MM-DD HH:MM:SS
* twp: String variable, Township
* addr: String variable, Address
* e: String variable, Dummy variable (always 1)
Just go along with this notebook and try to complete the instructions or answer the questions in bold using your Python and Data Science skills!
## Data and Setup
____
** Import numpy and pandas **
```
import numpy as np
import pandas as pd
```
** Import visualization libraries and set %matplotlib inline. **
```
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
%matplotlib inline
```
** Read in the csv file as a dataframe called df **
```
df = pd.read_csv('911.csv')
```
** Check the info() of the df **
```
df.info()
```
** Check the head of df **
```
df.head(3)
```
## Basic Questions
** What are the top 5 zipcodes for 911 calls? **
```
df['zip'].value_counts().head(5)
```
** What are the top 5 townships (twp) for 911 calls? **
```
df['twp'].value_counts().head(5)
```
** Take a look at the 'title' column, how many unique title codes are there? **
```
df['title'].nunique()
```
## Creating new features
** In the titles column there are "Reasons/Departments" specified before the title code. These are EMS, Fire, and Traffic. Use .apply() with a custom lambda expression to create a new column called "Reason" that contains this string value.**
**For example, if the title column value is EMS: BACK PAINS/INJURY , the Reason column value would be EMS. **
```
df['Reason'] = df['title'].apply(lambda title: title.split(':')[0])
```
** What is the most common Reason for a 911 call based off of this new column? **
```
df['Reason'].value_counts()
```
** Now use seaborn to create a countplot of 911 calls by Reason. **
```
sns.countplot(x='Reason',data=df,palette='viridis')
```
___
** Now let us begin to focus on time information. What is the data type of the objects in the timeStamp column? **
```
type(df['timeStamp'].iloc[0])
```
** You should have seen that these timestamps are still strings. Use [pd.to_datetime](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.to_datetime.html) to convert the column from strings to DateTime objects. **
```
df['timeStamp'] = pd.to_datetime(df['timeStamp'])
```
** You can now grab specific attributes from a Datetime object by calling them. For example:**
time = df['timeStamp'].iloc[0]
time.hour
**You can use Jupyter's tab method to explore the various attributes you can call. Now that the timestamp column are actually DateTime objects, use .apply() to create 3 new columns called Hour, Month, and Day of Week. You will create these columns based off of the timeStamp column, reference the solutions if you get stuck on this step.**
```
df['Hour'] = df['timeStamp'].apply(lambda time: time.hour)
df['Month'] = df['timeStamp'].apply(lambda time: time.month)
df['Day of Week'] = df['timeStamp'].apply(lambda time: time.dayofweek)
```
** Notice how the Day of Week is an integer 0-6. Use the .map() with this dictionary to map the actual string names to the day of the week: **
dmap = {0:'Mon',1:'Tue',2:'Wed',3:'Thu',4:'Fri',5:'Sat',6:'Sun'}
```
dmap = {0:'Mon',1:'Tue',2:'Wed',3:'Thu',4:'Fri',5:'Sat',6:'Sun'}
df['Day of Week'] = df['Day of Week'].map(dmap)
```
** Now use seaborn to create a countplot of the Day of Week column with the hue based off of the Reason column. **
```
sns.countplot(x='Day of Week',data=df,hue='Reason',palette='viridis')
# To relocate the legend
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
```
** Now do the same for Month:**
```
sns.countplot(x='Month',data=df,hue='Reason',palette='viridis')
# To relocate the legend
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
```
** Did you notice something strange about the Plot? **
```
# It is missing some months! 9,10, and 11 are not there.
```
** You should have noticed it was missing some Months, let's see if we can maybe fill in this information by plotting the information in another way, possibly a simple line plot that fills in the missing months, in order to do this, we'll need to do some work with pandas...**
** Now create a gropuby object called byMonth, where you group the DataFrame by the month column and use the count() method for aggregation. Use the head() method on this returned DataFrame. **
```
byMonth = df.groupby('Month').count()
byMonth.head()
```
** Now create a simple plot off of the dataframe indicating the count of calls per month. **
```
# Could be any column
byMonth['twp'].plot()
```
** Now see if you can use seaborn's lmplot() to create a linear fit on the number of calls per month. Keep in mind you may need to reset the index to a column. **
```
sns.lmplot(x='Month',y='twp',data=byMonth.reset_index())
```
**Create a new column called 'Date' that contains the date from the timeStamp column. You'll need to use apply along with the .date() method. **
```
df['Date']=df['timeStamp'].apply(lambda t: t.date())
```
** Now groupby this Date column with the count() aggregate and create a plot of counts of 911 calls.**
```
df.groupby('Date').count()['twp'].plot()
plt.tight_layout()
```
** Now recreate this plot but create 3 separate plots with each plot representing a Reason for the 911 call**
```
df[df['Reason']=='Traffic'].groupby('Date').count()['twp'].plot()
plt.title('Traffic')
plt.tight_layout()
df[df['Reason']=='Fire'].groupby('Date').count()['twp'].plot()
plt.title('Fire')
plt.tight_layout()
df[df['Reason']=='EMS'].groupby('Date').count()['twp'].plot()
plt.title('EMS')
plt.tight_layout()
```
____
** Now let's move on to creating heatmaps with seaborn and our data. We'll first need to restructure the dataframe so that the columns become the Hours and the Index becomes the Day of the Week. There are lots of ways to do this, but I would recommend trying to combine groupby with an [unstack](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.unstack.html) method. Reference the solutions if you get stuck on this!**
```
dayHour = df.groupby(by=['Day of Week','Hour']).count()['Reason'].unstack()
dayHour.head()
```
** Now create a HeatMap using this new DataFrame. **
```
plt.figure(figsize=(12,6))
sns.heatmap(dayHour,cmap='viridis')
```
** Now create a clustermap using this DataFrame. **
```
sns.clustermap(dayHour,cmap='viridis')
```
** Now repeat these same plots and operations, for a DataFrame that shows the Month as the column. **
```
dayMonth = df.groupby(by=['Day of Week','Month']).count()['Reason'].unstack()
dayMonth.head()
plt.figure(figsize=(12,6))
sns.heatmap(dayMonth,cmap='viridis')
sns.clustermap(dayMonth,cmap='viridis')
```
**Continue exploring the Data however you see fit!**
# Great Job!
| github_jupyter |
```
import re
# xtract all twitter handles from following text. Twitter handle is the text that appears after https://twitter.com/ and is a single word. Also it contains only alpha numeric characters i.e. A-Z a-z , o to 9 and underscore _
text = '''
Follow our leader Elon musk on twitter here: https://twitter.com/elonmusk, more information
on Tesla's products can be found at https://www.tesla.com/. Also here are leading influencers
for tesla related news,
https://twitter.com/teslarati
https://twitter.com/dummy_tesla
https://twitter.com/dummy_2_tesla\
'''
def get_pattern_information(pattern, text):
matches = re.findall(pattern, text)
if matches:
return matches
else:
return "not a match"
get_pattern_information('https:\/\/twitter\.com\/([a-zA-Z0-9_]*)', text)
# Extract Concentration Risk Types. It will be a text that appears after "Concentration Risk:", In below example, your regex should extract these two strings
text2 = '''
Concentration of Risk: Credit Risk
Financial instruments that potentially subject us to a concentration of credit risk consist of cash, cash equivalents, marketable securities,
restricted cash, accounts receivable, convertible note hedges, and interest rate swaps. Our cash balances are primarily invested in money market funds
or on deposit at high credit quality financial institutions in the U.S. These deposits are typically in excess of insured limits. As of September 30, 2021
and December 31, 2020, no entity represented 10% or more of our total accounts receivable balance. The risk of concentration for our convertible note
hedges and interest rate swaps is mitigated by transacting with several highly-rated multinational banks.
Concentration of Risk: Supply Risk
We are dependent on our suppliers, including single source suppliers, and the inability of these suppliers to deliver necessary components of our
products in a timely manner at prices, quality levels and volumes acceptable to us, or our inability to efficiently manage these components from these
suppliers, could have a material adverse effect on our business, prospects, financial condition and operating results.
'''
get_pattern_information('Concentration of Risk: ([^\n]*)', text2)
#Companies in europe reports their financial numbers of semi annual basis and you can have a document like this. To exatract quarterly and semin annual period you can use a regex as shown below
text3 = '''
Tesla's gross cost of operating lease vehicles in FY2021 Q1 was $4.85 billion.
BMW's gross cost of operating vehicles in FY2021 S1 was $8 billion.
'''
get_pattern_information('FY(\d{4} (?:Q[1-4]|S[1-2]))', text3)
```
| github_jupyter |
# Exemplary end-to-end causal analysis with ```cause2e```
This notebook shows an example of how ```cause2e``` can be used as a standalone package for end-to-end causal analysis. It illustrates how we can proceed in stringing together many causal techniques that have previously required fitting together various algorithms from separate sources with unclear interfaces. Hopefully, you will find this notebook helpful in guiding you through the process of setting up your own causal analyses for custom problems. The overall structure should always be the same regardless of the application domain. Some of cause2e's functionality is only hinted at and is explained more thoroughly in other notebooks. If you are looking for a less detailed version, we recommend the minimal end to end analysis in another notebook for getting started.
### Imports
By the end of this notebook, you will probably be pleasantly surprised by the fact that we did not have to import lots of different packages to perform a complex analysis consisting of many different subtasks.
```
import os
from numpy.random import seed
from cause2e import path_mgr, data_generator, knowledge, discovery, estimator
```
## Set up paths to data and output directories
This step is conveniently handled by the ```PathManager``` class, which avoids having to wrestle with paths throughout the multistep causal analysis. If we want to perform the analysis in a directory ```'dirname'``` that contains ```'dirname/data'``` and ```'dirname/output'``` as subdirectories, we can also use ```PathManagerQuick``` for an even easier setup. The experiment name is used for generating output files with meaningful names, in case we want to study multiple scenarios (e.g. with varying model parameters).
```
cwd = os.getcwd()
wd = os.path.dirname(cwd)
paths = path_mgr.PathManagerQuick(experiment_name='linear_test',
data_name='linear_test.csv',
directory=wd
)
```
## Generate data
For checking if our algorithms can recover known causal graphs and effects, it can be helpful to use synthetic data. This data generator is based on a class from the ```DoWhy``` package. Setting the seed only serves the purpose of providing consistent results over several runs of this notebook. Feel free to change it and experiment with the different outcomes!
```
seed(4)
generator = data_generator.DataGenerator(paths)
generator.generate_linear_dataset(beta=2,
n_common_causes=4,
nrows=25000
)
```
In this example, we want to estimate the causal effect that a variable ```v0``` has on another variable ```y```. Note that we have prescribed the causal effect to be ```beta=2```. There are 4 common causes acting as confounders. In causal inference, it can be very helpful to visualize these assumptions in a causal graph.
```
generator.display_graph()
```
In order to imitate in the remainder of the notebook how we would analyze real-world data, the data is written to a csv file.
```
generator.write_csv()
```
## Learn the causal graph from data and domain knowledge
Model-based causal inference leverages qualitative knowledge about pairwise causal connections to obtain unbiased estimates of quantitative causal effects. The qualitative knowledge is encoded in the causal graph, so we must recover this graph before we can start actually estimating the desired effect. For learning the graph from data and domain knowledge, we use the ```StructureLearner``` class. Note that we can reuse the ```PathManager``` from before to set the right paths.
```
learner = discovery.StructureLearner(paths)
```
### Read the data
The ```StructureLearner``` has reading methods for csv and parquet files.
```
learner.read_csv(index_col=0)
```
Supposing that we had not generated the data ourselves, the first step in the analysis should be an assessment of which variables we are dealing with.
```
learner.variables
```
### Preprocess the data
The raw data is often not suited as immediate input to machine learning algorithms, so we provide some preprocessing facilities: Adding, deleting, renaming and recombining variables is possible. In this example, we demonstrate the possibility to rename some variables, which is necessary to meet the naming constraints of the internally used ```py-causal``` package.
```
name_change = {'v0': 'v',
'W0': 'W_zero',
'W1': 'W_one',
'W2': 'W_two',
'W3': 'W_three'
}
for current_name, new_name in name_change.items():
learner.rename_variable(current_name, new_name)
```
Once we have our data in the final form, it will be necessary to communicate to the ```StructureLearner``` if the variables are discrete, continuous, or both. We check how many unique values each variable takes on in our sample and deduce that all variables except for ```v``` are continuous.
```
learner.data.nunique()
```
This information is passed to the StructureLearner by indicating the exact sets of discrete and continuous variables.
```
learner.discrete = {'v'}
learner.continuous = learner.variables - learner.discrete
```
### Provide domain knowledge
Humans can often infer parts of the causal graph from domain knowledge. The nodes are always just the variables in the data, so the problem of finding the right graph comes down to selecting the right edges between them.
There are three ways of passing domain knowledge:
- Indicate which edges must be present in the causal graph.
- Indicate which edges must not be present in the causal graph.
- Indicate a temporal order in which the variables have been created. This is then used to generate forbidden edges, since the future can never influence the past.
In our first attempt to recover the graph from data and domain knowledge, we only assume that ```y``` is directly influenced by ```v```.
```
edge_creator = knowledge.EdgeCreator()
edge_creator.require_edge('v', 'y')
```
We pass the knowledge to the StructureLearner and check if it has been correctly received.
```
learner.set_knowledge(edge_creator)
learner.show_knowledge()
```
### Select a structure learning algorithm
```Cause2e``` provides some informative methods for finding the right structure learning algorithm from the internally called ```py-causal``` package. In order to spare users from the pain of going through all the necessary reading whenever they want to perform just a quick exploratory analysis, we have provided default arguments (FGES with CG-BIC score for possibly mixed datatypes and respecting domain knowledge) that let you start the search without any finetuning. Just call ```run_quick_search``` and you are good to go. For algorithm selection and finetuning of the parameters, check out the related notebook.
### Apply the selected structure learning algorithm
The output of the search is a proposed causal graph. We can ignore the warning about stopping the Java Virtual Machine (needed by ```py-causal``` which is a wrapper around the ```TETRAD``` software that is written in Java) if we do not run into any problems. If the algorithm cannot orient all edges, we need to do this manually. Therefore, the output includes a list of all undirected edges, so we do not miss them in complicated graphs with many variables and edges.
```
learner.run_quick_search(save_graph=False)
```
Are we done after some calls to ```learner.add_edge```? No! If we compare the output of the search algorithm to the true causal graph that was used for generating the data, we see that it is not right.
This can happen for several reasons:
- Insufficient data
- Bad choice of search algorithm, hyperparameters, ...
- Several Markov-equivalent graphs fit the data equally well
If we cannot get more data and we have no way of choosing a better search algorithm, the key to good search results is passing comprehensive domain knowledge. This is especially helpful for the problem of choosing between equivalent graphs that are impossible to distinguish using purely statistical methods without domain knowledge.
As a quick check, let us see what happens if we do not pass any domain knowledge.
```
learner.erase_knowledge()
learner.show_knowledge()
```
The graph is the same as above, meaning that our sparse domain knowledge about only the edge from ```v``` to ```y``` was basically worthless.
```
learner.run_quick_search(save_graph=False)
```
Now, let us see if we can come closer to the true causal graph by nurturing our search algorithm with more valuable domain knowledge. Suppose that we know that the confounders themselves are not influenced by any variables. This short and compact sentence contains a lot of constraints on the edges in the graph, as you can see from the output of the next code cell. In principle, we could have passed the set of forbidden edges explicitly after manually enumerating them, but it is often more convenient to directly reason about groups of variables, as is illustrated in the code.
```
edge_creator = knowledge.EdgeCreator()
w_group = {'W_zero', 'W_one', 'W_two', 'W_three'}
edge_creator.forbid_edges_from_groups(w_group, incoming=learner.variables)
edge_creator.require_edge('v', 'y')
learner.set_knowledge(edge_creator)
learner.show_knowledge()
```
Repeating the search with the additional domain knowledge gives a causal graph that is already very close to the true graph, showing the importance of explicitly using our knowledge in the search process.
```
learner.run_quick_search(save_graph=False)
```
### Manually postprocess the graph
It often happens that some edges (present or absent) in the resulting graph feel "odd" and we want to change them manually and compare the results. The possible postprocessing operations are adding, removing and reversing edges.
\[Note: If there is one edge that absolutely should not (or absolutely should) be present in the graph, please consider adding it to the domain knowledge prior to starting the search, instead of making the search unnecessarily hard for the algorithm.\]
In our case, we might find it weird that all the ```W``` variables except for ```W_zero``` have edges into both ```v``` and ```y```, so let us add the "missing" edge.
```
learner.add_edge('W_zero', 'v')
```
### Saving the graph
```Cause2e``` allows us to save the result of our search to different file formats. The name of the file is determined by the ```experiment_name``` parameter from the ```PathManager```.
```
learner.save_graphs(['dot', 'png', 'svg'])
```
It is worth noting that the saving procedure includes checking that the graph
- is fully directed
- does not contain any cycles
- respects our domain knowledge
This ensures that it can be used in the further causal analysis without any unwanted problems. Just to see this check in action, we add an edge that creates a cycle.
```
learner.add_edge('y', 'W_one')
```
When trying to save now, we get an error because the graph is not acyclic.
```
learner.save_graphs(['dot', 'png', 'svg'])
```
You can also check these conditions explicitly outside of a saving attempt. This time, we get a message that one of the edges conflicts with our domain knowledge.
```
learner.is_acyclic()
learner.has_undirected_edges()
learner.respects_knowledge()
```
If we are not interested in feeding the search result into other algorithms for further causal analysis, but rather in just recording what the search procedure has delivered, we can also switch off the checks.
```
learner.save_graphs(['dot', 'png', 'svg'], strict=False)
```
We end this excursion by removing the faulty edge and restoring the true causal graph.
```
learner.remove_edge('y', 'W_one')
learner.save_graphs(['dot', 'png', 'svg'])
```
## Estimate causal effects from the graph and the data
After we have successfully recovered the causal graph from data and domain knowledge, we can use it to estimate quantitative causal effects between the variables in the graph. It is pleasant that we can use the same graph and data to estimate multiple causal effects, e.g. the one that ```v``` has on ```y```, as well as the one that ```W_one``` has on ```v```, without having to repeat the previous steps. Once we have managed to qualitatively model the data generating process, we are already in a very good position. The remaining challenges can be tackled with the core functionality from the DoWhy package and some helper methods that make the transition from learning the graph to using it for estimation less cumbersome.
### Create an Estimator
The ```Estimator``` class is mainly a wrapper around methods of the ```DoWhy``` package. It helps us in performing a principled and modular estimation of causal effects from data and a given causal graph. An exemplary added feature is the possibility to pass a list of transformations that the ```StructureLearner``` has applied in preprocessing the data, which makes it easy to cast your data into the same form for estimation.
```
estim = estimator.Estimator(paths, learner.transformations)
```
### Read the data
Just like the ```StructureLearner```, the ```Estimator``` also has methods for reading data. Having two independent reading steps instead of passing the data directly comes in handy when we want to use different sample sizes for causal discovery and estimation, when we are dealing with big data that cannot be fully stored in RAM, or when we are dealing with two entirely different datasets that only have the qualitative graph structure in common.
```
estim.read_csv(nrows=7000, index_col=0)
```
### Imitate preprocessing steps
With our built-in method, it is effortless to get the data in the same format that the causal graph is referring to. In our example, this just means that some variables will be renamed, but the principle stays the same for more involved preprocessing pipelines.
```
estim.imitate_data_trafos()
```
### Initialize the causal model
This is the step where we decide which specific effect we want to analyze. We are interested in estimating the effect of ```v``` on ```y```. ```DoWhy``` conveniently lets us choose between different types of causal effects (overall average treatment effect vs. more specific direct and indirect treatment effects in a mediation analysis).
```
treatment = 'v'
outcome = 'y'
estim.initialize_model(treatment,
outcome,
estimand_type="nonparametric-ate"
)
```
### Identify estimands for the causal effect
The big advantage of model-based causal inference is that once we have the causal graph, a statistical estimand for a specified causal effect (e.g. ```v``` on ```y```) can be derived by purely algebraic methods (Pearl's do-calculus). The estimand (verbally translated: "the thing that is to be estimated") does not tell us in numbers what the causal effect is, but it is more like an algebraic recipe for estimating the quantity of interest. In our case, the do-calculus tells us that a certain type of backdoor adjustment is the way to go, and the key assumptions for drawing this conclusion are explicitly stated right after a mathematical expression of the estimand. Depending on the graph structure, instrumental variable or frontdoor adjustment strategies can also work, but in this case no valid estimand can be derived using these strategies. The sole argument of the method serves to silence ```DoWhy```'s prompting of a user input that must be answered with 'yes' anyway and causes problems in non-interactive development settings.
```
estim.identify_estimand(proceed_when_unidentifiable=True)
```
### Estimate the causal effect
Now that we know what to estimate, we can focus on the how. For this example, we demonstrate the use of a simple linear regression estimator, but there are many more possibilities documented in [this notebook](https://microsoft.github.io/dowhy/example_notebooks/dowhy_estimation_methods.html) provided by the creators of ```DoWhy```. Using the estimand from the last step, the package knows which covariates should be included as inputs of the linear regression. The result shows that the estimate is very close to the true value of the causal effect (```beta=2```). The additional output summarizes all decisions that we have taken in order to arrive at this value.
```
estim.estimate_effect(verbose=True,
method_name='backdoor.linear_regression'
)
```
### Perform robustness checks
After we have estimated the causal effect, performing robustness checks can lend additional credibility to our final estimate. DoWhy provides several such checks called 'refuters' that are explained [here](https://microsoft.github.io/dowhy/readme.html#iv-refute-the-obtained-estimate). As an example, we check that the estimate is robust to adding an independent random variable as a common cause to the dataset.
```
estim.check_robustness(verbose=True,
method_name='random_common_cause'
)
```
### Compare the result to a non-causal regression analysis
Without the causal framework, causal effects are often estimated by running a linear regression with the outcome (```y```) as target variable and the treatment (```v```) and possibly other covariates as input variables. The causal effect is then assumed to be the coefficient associated with the treatment variable in the resulting regression equation. In order to assess what we have actually gained by applying all the causal machinery in this notebook, it can be helpful to compare our causal estimate of the effect to the regression-based method. We see that a regression analysis with only ```v``` as input fails to estimate the correct causal effect. This is not surprising, given that it neglects all 4 confounders.
```
estim.compare_to_noncausal_regression(input_cols={'v'})
```
The situation changes if we select all variables except for ```y``` as input to the linear regression. Up to a small numerical error, the result is identical to the causal estimate that we have gained after lots of hard work. Again, this is not a surprise if we look at the "Realized estimand" in the causal estimation step, showing that we use the exact same linear regression in both approaches.
```
estim.compare_to_noncausal_regression(input_cols={'y'}, drop_cols=True)
```
## Final thoughts
After this comparison, we might ask ourselves: Was it worth all the trouble to arrive at the same result that we could have gotten with a simple linear regression? The answer is subtle: In principle, we can always guess the correct input variables for a (possibly nonlinear) regression that will allow us to estimate the desired causal effect.
The problem is that
- we must be at the same time domain experts and statistical experts to take the right decisions
- there is no principled way to decide if a variable should be a regression input or not, so it will ultimatively always come down to justifying it with "gut feeling". There are many situations where simply including all the covariates as inputs will be as misleading as not including any covariates except for the treatment itself and only a carefully chosen subset of the variables will lead to the correct causal estimate.
- many underlying causal graphs are just too complex to be disentangled by humans. How would you choose correct backdoor (let alone frontdoor) adjustment sets in a problem with 100 variables without even having the causal graph as an explicit representation?
The causal framework on the other hand provides
- a modular structure with clear separation between all the methodological and knowledge-oriented decisions that need to be taken
- a principled way of analysis with firm theoretical backing (e.g. the mathematics behind the do-calculus)
- representations of otherwise implicit domain knowledge that make graph learning and causal identification algorithmically treatable and therefore also possible in highly complex scenarios
| github_jupyter |
**Math - Linear Algebra**
*Linear Algebra is the branch of mathematics that studies [vector spaces](https://en.wikipedia.org/wiki/Vector_space) and linear transformations between vector spaces, such as rotating a shape, scaling it up or down, translating it (ie. moving it), etc.*
*Machine Learning relies heavily on Linear Algebra, so it is essential to understand what vectors and matrices are, what operations you can perform with them, and how they can be useful.*
# Vectors
## Definition
A vector is a quantity defined by a magnitude and a direction. For example, a rocket's velocity is a 3-dimensional vector: its magnitude is the speed of the rocket, and its direction is (hopefully) up. A vector can be represented by an array of numbers called *scalars*. Each scalar corresponds to the magnitude of the vector with regards to each dimension.
For example, say the rocket is going up at a slight angle: it has a vertical speed of 5,000 m/s, and also a slight speed towards the East at 10 m/s, and a slight speed towards the North at 50 m/s. The rocket's velocity may be represented by the following vector:
**velocity** $= \begin{pmatrix}
10 \\
50 \\
5000 \\
\end{pmatrix}$
Note: by convention vectors are generally presented in the form of columns. Also, vector names are generally lowercase to distinguish them from matrices (which we will discuss below) and in bold (when possible) to distinguish them from simple scalar values such as ${meters\_per\_second} = 5026$.
A list of N numbers may also represent the coordinates of a point in an N-dimensional space, so it is quite frequent to represent vectors as simple points instead of arrows. A vector with 1 element may be represented as an arrow or a point on an axis, a vector with 2 elements is an arrow or a point on a plane, a vector with 3 elements is an arrow or point in space, and a vector with N elements is an arrow or a point in an N-dimensional space… which most people find hard to imagine.
## Purpose
Vectors have many purposes in Machine Learning, most notably to represent observations and predictions. For example, say we built a Machine Learning system to classify videos into 3 categories (good, spam, clickbait) based on what we know about them. For each video, we would have a vector representing what we know about it, such as:
**video** $= \begin{pmatrix}
10.5 \\
5.2 \\
3.25 \\
7.0
\end{pmatrix}$
This vector could represent a video that lasts 10.5 minutes, but only 5.2% viewers watch for more than a minute, it gets 3.25 views per day on average, and it was flagged 7 times as spam. As you can see, each axis may have a different meaning.
Based on this vector our Machine Learning system may predict that there is an 80% probability that it is a spam video, 18% that it is clickbait, and 2% that it is a good video. This could be represented as the following vector:
**class_probabilities** $= \begin{pmatrix}
0.80 \\
0.18 \\
0.02
\end{pmatrix}$
## Vectors in python
In python, a vector can be represented in many ways, the simplest being a regular python list of numbers:
```
[10.5, 5.2, 3.25, 7.0]
```
Since we plan to do quite a lot of scientific calculations, it is much better to use NumPy's `ndarray`, which provides a lot of convenient and optimized implementations of essential mathematical operations on vectors (for more details about NumPy, check out the [NumPy tutorial](tools_numpy.ipynb)). For example:
```
import numpy as np
video = np.array([10.5, 5.2, 3.25, 7.0])
video
```
The size of a vector can be obtained using the `size` attribute:
```
video.size
```
The $i^{th}$ element (also called *entry* or *item*) of a vector $\textbf{v}$ is noted $\textbf{v}_i$.
Note that indices in mathematics generally start at 1, but in programming they usually start at 0. So to access $\textbf{video}_3$ programmatically, we would write:
```
video[2] # 3rd element
```
## Plotting vectors
To plot vectors we will use matplotlib, so let's start by importing it (for details about matplotlib, check the [matplotlib tutorial](tools_matplotlib.ipynb)):
```
%matplotlib inline
import matplotlib.pyplot as plt
```
### 2D vectors
Let's create a couple very simple 2D vectors to plot:
```
u = np.array([2, 5])
v = np.array([3, 1])
```
These vectors each have 2 elements, so they can easily be represented graphically on a 2D graph, for example as points:
```
x_coords, y_coords = zip(u, v)
plt.scatter(x_coords, y_coords, color=["r","b"])
plt.axis([0, 9, 0, 6])
plt.grid()
plt.show()
```
Vectors can also be represented as arrows. Let's create a small convenience function to draw nice arrows:
```
def plot_vector2d(vector2d, origin=[0, 0], **options):
return plt.arrow(origin[0], origin[1], vector2d[0], vector2d[1],
head_width=0.2, head_length=0.3, length_includes_head=True,
**options)
```
Now let's draw the vectors **u** and **v** as arrows:
```
plot_vector2d(u, color="r")
plot_vector2d(v, color="b")
plt.axis([0, 9, 0, 6])
plt.grid()
plt.show()
```
### 3D vectors
Plotting 3D vectors is also relatively straightforward. First let's create two 3D vectors:
```
a = np.array([1, 2, 8])
b = np.array([5, 6, 3])
```
Now let's plot them using matplotlib's `Axes3D`:
```
from mpl_toolkits.mplot3d import Axes3D
subplot3d = plt.subplot(111, projection='3d')
x_coords, y_coords, z_coords = zip(a,b)
subplot3d.scatter(x_coords, y_coords, z_coords)
subplot3d.set_zlim3d([0, 9])
plt.show()
```
It is a bit hard to visualize exactly where in space these two points are, so let's add vertical lines. We'll create a small convenience function to plot a list of 3d vectors with vertical lines attached:
```
def plot_vectors3d(ax, vectors3d, z0, **options):
for v in vectors3d:
x, y, z = v
ax.plot([x,x], [y,y], [z0, z], color="gray", linestyle='dotted', marker=".")
x_coords, y_coords, z_coords = zip(*vectors3d)
ax.scatter(x_coords, y_coords, z_coords, **options)
subplot3d = plt.subplot(111, projection='3d')
subplot3d.set_zlim([0, 9])
plot_vectors3d(subplot3d, [a,b], 0, color=("r","b"))
plt.show()
```
## Norm
The norm of a vector $\textbf{u}$, noted $\left \Vert \textbf{u} \right \|$, is a measure of the length (a.k.a. the magnitude) of $\textbf{u}$. There are multiple possible norms, but the most common one (and the only one we will discuss here) is the Euclidian norm, which is defined as:
$\left \Vert \textbf{u} \right \| = \sqrt{\sum_{i}{\textbf{u}_i}^2}$
We could implement this easily in pure python, recalling that $\sqrt x = x^{\frac{1}{2}}$
```
def vector_norm(vector):
squares = [element**2 for element in vector]
return sum(squares)**0.5
print("||", u, "|| =")
vector_norm(u)
```
However, it is much more efficient to use NumPy's `norm` function, available in the `linalg` (**Lin**ear **Alg**ebra) module:
```
import numpy.linalg as LA
LA.norm(u)
```
Let's plot a little diagram to confirm that the length of vector $\textbf{v}$ is indeed $\approx5.4$:
```
radius = LA.norm(u)
plt.gca().add_artist(plt.Circle((0,0), radius, color="#DDDDDD"))
plot_vector2d(u, color="red")
plt.axis([0, 8.7, 0, 6])
plt.grid()
plt.show()
```
Looks about right!
## Addition
Vectors of same size can be added together. Addition is performed *elementwise*:
```
print(" ", u)
print("+", v)
print("-"*10)
u + v
```
Let's look at what vector addition looks like graphically:
```
plot_vector2d(u, color="r")
plot_vector2d(v, color="b")
plot_vector2d(v, origin=u, color="b", linestyle="dotted")
plot_vector2d(u, origin=v, color="r", linestyle="dotted")
plot_vector2d(u+v, color="g")
plt.axis([0, 9, 0, 7])
plt.text(0.7, 3, "u", color="r", fontsize=18)
plt.text(4, 3, "u", color="r", fontsize=18)
plt.text(1.8, 0.2, "v", color="b", fontsize=18)
plt.text(3.1, 5.6, "v", color="b", fontsize=18)
plt.text(2.4, 2.5, "u+v", color="g", fontsize=18)
plt.grid()
plt.show()
```
Vector addition is **commutative**, meaning that $\textbf{u} + \textbf{v} = \textbf{v} + \textbf{u}$. You can see it on the previous image: following $\textbf{u}$ *then* $\textbf{v}$ leads to the same point as following $\textbf{v}$ *then* $\textbf{u}$.
Vector addition is also **associative**, meaning that $\textbf{u} + (\textbf{v} + \textbf{w}) = (\textbf{u} + \textbf{v}) + \textbf{w}$.
If you have a shape defined by a number of points (vectors), and you add a vector $\textbf{v}$ to all of these points, then the whole shape gets shifted by $\textbf{v}$. This is called a [geometric translation](https://en.wikipedia.org/wiki/Translation_%28geometry%29):
```
t1 = np.array([2, 0.25])
t2 = np.array([2.5, 3.5])
t3 = np.array([1, 2])
x_coords, y_coords = zip(t1, t2, t3, t1)
plt.plot(x_coords, y_coords, "c--", x_coords, y_coords, "co")
plot_vector2d(v, t1, color="r", linestyle=":")
plot_vector2d(v, t2, color="r", linestyle=":")
plot_vector2d(v, t3, color="r", linestyle=":")
t1b = t1 + v
t2b = t2 + v
t3b = t3 + v
x_coords_b, y_coords_b = zip(t1b, t2b, t3b, t1b)
plt.plot(x_coords_b, y_coords_b, "b-", x_coords_b, y_coords_b, "bo")
plt.text(4, 4.2, "v", color="r", fontsize=18)
plt.text(3, 2.3, "v", color="r", fontsize=18)
plt.text(3.5, 0.4, "v", color="r", fontsize=18)
plt.axis([0, 6, 0, 5])
plt.grid()
plt.show()
```
Finally, substracting a vector is like adding the opposite vector.
## Multiplication by a scalar
Vectors can be multiplied by scalars. All elements in the vector are multiplied by that number, for example:
```
print("1.5 *", u, "=")
1.5 * u
```
Graphically, scalar multiplication results in changing the scale of a figure, hence the name *scalar*. The distance from the origin (the point at coordinates equal to zero) is also multiplied by the scalar. For example, let's scale up by a factor of `k = 2.5`:
```
k = 2.5
t1c = k * t1
t2c = k * t2
t3c = k * t3
plt.plot(x_coords, y_coords, "c--", x_coords, y_coords, "co")
plot_vector2d(t1, color="r")
plot_vector2d(t2, color="r")
plot_vector2d(t3, color="r")
x_coords_c, y_coords_c = zip(t1c, t2c, t3c, t1c)
plt.plot(x_coords_c, y_coords_c, "b-", x_coords_c, y_coords_c, "bo")
plot_vector2d(k * t1, color="b", linestyle=":")
plot_vector2d(k * t2, color="b", linestyle=":")
plot_vector2d(k * t3, color="b", linestyle=":")
plt.axis([0, 9, 0, 9])
plt.grid()
plt.show()
```
As you might guess, dividing a vector by a scalar is equivalent to multiplying by its multiplicative inverse (reciprocal):
$\dfrac{\textbf{u}}{\lambda} = \dfrac{1}{\lambda} \times \textbf{u}$
Scalar multiplication is **commutative**: $\lambda \times \textbf{u} = \textbf{u} \times \lambda$.
It is also **associative**: $\lambda_1 \times (\lambda_2 \times \textbf{u}) = (\lambda_1 \times \lambda_2) \times \textbf{u}$.
Finally, it is **distributive** over addition of vectors: $\lambda \times (\textbf{u} + \textbf{v}) = \lambda \times \textbf{u} + \lambda \times \textbf{v}$.
## Zero, unit and normalized vectors
* A **zero-vector ** is a vector full of 0s.
* A **unit vector** is a vector with a norm equal to 1.
* The **normalized vector** of a non-null vector $\textbf{u}$, noted $\hat{\textbf{u}}$, is the unit vector that points in the same direction as $\textbf{u}$. It is equal to: $\hat{\textbf{u}} = \dfrac{\textbf{u}}{\left \Vert \textbf{u} \right \|}$
```
plt.gca().add_artist(plt.Circle((0,0),1,color='c'))
plt.plot(0, 0, "ko")
plot_vector2d(v / LA.norm(v), color="k")
plot_vector2d(v, color="b", linestyle=":")
plt.text(0.3, 0.3, "$\hat{u}$", color="k", fontsize=18)
plt.text(1.5, 0.7, "$u$", color="b", fontsize=18)
plt.axis([-1.5, 5.5, -1.5, 3.5])
plt.grid()
plt.show()
```
## Dot product
### Definition
The dot product (also called *scalar product* or *inner product* in the context of the Euclidian space) of two vectors $\textbf{u}$ and $\textbf{v}$ is a useful operation that comes up fairly often in linear algebra. It is noted $\textbf{u} \cdot \textbf{v}$, or sometimes $⟨\textbf{u}|\textbf{v}⟩$ or $(\textbf{u}|\textbf{v})$, and it is defined as:
$\textbf{u} \cdot \textbf{v} = \left \Vert \textbf{u} \right \| \times \left \Vert \textbf{v} \right \| \times cos(\theta)$
where $\theta$ is the angle between $\textbf{u}$ and $\textbf{v}$.
Another way to calculate the dot product is:
$\textbf{u} \cdot \textbf{v} = \sum_i{\textbf{u}_i \times \textbf{v}_i}$
### In python
The dot product is pretty simple to implement:
```
def dot_product(v1, v2):
return sum(v1i * v2i for v1i, v2i in zip(v1, v2))
dot_product(u, v)
```
But a *much* more efficient implementation is provided by NumPy with the `dot` function:
```
np.dot(u,v)
```
Equivalently, you can use the `dot` method of `ndarray`s:
```
u.dot(v)
```
**Caution**: the `*` operator will perform an *elementwise* multiplication, *NOT* a dot product:
```
print(" ",u)
print("* ",v, "(NOT a dot product)")
print("-"*10)
u * v
```
### Main properties
* The dot product is **commutative**: $\textbf{u} \cdot \textbf{v} = \textbf{v} \cdot \textbf{u}$.
* The dot product is only defined between two vectors, not between a scalar and a vector. This means that we cannot chain dot products: for example, the expression $\textbf{u} \cdot \textbf{v} \cdot \textbf{w}$ is not defined since $\textbf{u} \cdot \textbf{v}$ is a scalar and $\textbf{w}$ is a vector.
* This also means that the dot product is **NOT associative**: $(\textbf{u} \cdot \textbf{v}) \cdot \textbf{w} ≠ \textbf{u} \cdot (\textbf{v} \cdot \textbf{w})$ since neither are defined.
* However, the dot product is **associative with regards to scalar multiplication**: $\lambda \times (\textbf{u} \cdot \textbf{v}) = (\lambda \times \textbf{u}) \cdot \textbf{v} = \textbf{u} \cdot (\lambda \times \textbf{v})$
* Finally, the dot product is **distributive** over addition of vectors: $\textbf{u} \cdot (\textbf{v} + \textbf{w}) = \textbf{u} \cdot \textbf{v} + \textbf{u} \cdot \textbf{w}$.
### Calculating the angle between vectors
One of the many uses of the dot product is to calculate the angle between two non-zero vectors. Looking at the dot product definition, we can deduce the following formula:
$\theta = \arccos{\left ( \dfrac{\textbf{u} \cdot \textbf{v}}{\left \Vert \textbf{u} \right \| \times \left \Vert \textbf{v} \right \|} \right ) }$
Note that if $\textbf{u} \cdot \textbf{v} = 0$, it follows that $\theta = \dfrac{π}{2}$. In other words, if the dot product of two non-null vectors is zero, it means that they are orthogonal.
Let's use this formula to calculate the angle between $\textbf{u}$ and $\textbf{v}$ (in radians):
```
def vector_angle(u, v):
cos_theta = u.dot(v) / LA.norm(u) / LA.norm(v)
return np.arccos(np.clip(cos_theta, -1, 1))
theta = vector_angle(u, v)
print("Angle =", theta, "radians")
print(" =", theta * 180 / np.pi, "degrees")
```
Note: due to small floating point errors, `cos_theta` may be very slightly outside of the $[-1, 1]$ interval, which would make `arccos` fail. This is why we clipped the value within the range, using NumPy's `clip` function.
### Projecting a point onto an axis
The dot product is also very useful to project points onto an axis. The projection of vector $\textbf{v}$ onto $\textbf{u}$'s axis is given by this formula:
$\textbf{proj}_{\textbf{u}}{\textbf{v}} = \dfrac{\textbf{u} \cdot \textbf{v}}{\left \Vert \textbf{u} \right \| ^2} \times \textbf{u}$
Which is equivalent to:
$\textbf{proj}_{\textbf{u}}{\textbf{v}} = (\textbf{v} \cdot \hat{\textbf{u}}) \times \hat{\textbf{u}}$
```
u_normalized = u / LA.norm(u)
proj = v.dot(u_normalized) * u_normalized
plot_vector2d(u, color="r")
plot_vector2d(v, color="b")
plot_vector2d(proj, color="k", linestyle=":")
plt.plot(proj[0], proj[1], "ko")
plt.plot([proj[0], v[0]], [proj[1], v[1]], "b:")
plt.text(1, 2, "$proj_u v$", color="k", fontsize=18)
plt.text(1.8, 0.2, "$v$", color="b", fontsize=18)
plt.text(0.8, 3, "$u$", color="r", fontsize=18)
plt.axis([0, 8, 0, 5.5])
plt.grid()
plt.show()
```
# Matrices
A matrix is a rectangular array of scalars (ie. any number: integer, real or complex) arranged in rows and columns, for example:
\begin{bmatrix} 10 & 20 & 30 \\ 40 & 50 & 60 \end{bmatrix}
You can also think of a matrix as a list of vectors: the previous matrix contains either 2 horizontal 3D vectors or 3 vertical 2D vectors.
Matrices are convenient and very efficient to run operations on many vectors at a time. We will also see that they are great at representing and performing linear transformations such rotations, translations and scaling.
## Matrices in python
In python, a matrix can be represented in various ways. The simplest is just a list of python lists:
```
[
[10, 20, 30],
[40, 50, 60]
]
```
A much more efficient way is to use the NumPy library which provides optimized implementations of many matrix operations:
```
A = np.array([
[10,20,30],
[40,50,60]
])
A
```
By convention matrices generally have uppercase names, such as $A$.
In the rest of this tutorial, we will assume that we are using NumPy arrays (type `ndarray`) to represent matrices.
## Size
The size of a matrix is defined by its number of rows and number of columns. It is noted $rows \times columns$. For example, the matrix $A$ above is an example of a $2 \times 3$ matrix: 2 rows, 3 columns. Caution: a $3 \times 2$ matrix would have 3 rows and 2 columns.
To get a matrix's size in NumPy:
```
A.shape
```
**Caution**: the `size` attribute represents the number of elements in the `ndarray`, not the matrix's size:
```
A.size
```
## Element indexing
The number located in the $i^{th}$ row, and $j^{th}$ column of a matrix $X$ is sometimes noted $X_{i,j}$ or $X_{ij}$, but there is no standard notation, so people often prefer to explicitely name the elements, like this: "*let $X = (x_{i,j})_{1 ≤ i ≤ m, 1 ≤ j ≤ n}$*". This means that $X$ is equal to:
$X = \begin{bmatrix}
x_{1,1} & x_{1,2} & x_{1,3} & \cdots & x_{1,n}\\
x_{2,1} & x_{2,2} & x_{2,3} & \cdots & x_{2,n}\\
x_{3,1} & x_{3,2} & x_{3,3} & \cdots & x_{3,n}\\
\vdots & \vdots & \vdots & \ddots & \vdots \\
x_{m,1} & x_{m,2} & x_{m,3} & \cdots & x_{m,n}\\
\end{bmatrix}$
However in this notebook we will use the $X_{i,j}$ notation, as it matches fairly well NumPy's notation. Note that in math indices generally start at 1, but in programming they usually start at 0. So to access $A_{2,3}$ programmatically, we need to write this:
```
A[1,2] # 2nd row, 3rd column
```
The $i^{th}$ row vector is sometimes noted $M_i$ or $M_{i,*}$, but again there is no standard notation so people often prefer to explicitely define their own names, for example: "*let **x**$_{i}$ be the $i^{th}$ row vector of matrix $X$*". We will use the $M_{i,*}$, for the same reason as above. For example, to access $A_{2,*}$ (ie. $A$'s 2nd row vector):
```
A[1, :] # 2nd row vector (as a 1D array)
```
Similarly, the $j^{th}$ column vector is sometimes noted $M^j$ or $M_{*,j}$, but there is no standard notation. We will use $M_{*,j}$. For example, to access $A_{*,3}$ (ie. $A$'s 3rd column vector):
```
A[:, 2] # 3rd column vector (as a 1D array)
```
Note that the result is actually a one-dimensional NumPy array: there is no such thing as a *vertical* or *horizontal* one-dimensional array. If you need to actually represent a row vector as a one-row matrix (ie. a 2D NumPy array), or a column vector as a one-column matrix, then you need to use a slice instead of an integer when accessing the row or column, for example:
```
A[1:2, :] # rows 2 to 3 (excluded): this returns row 2 as a one-row matrix
A[:, 2:3] # columns 3 to 4 (excluded): this returns column 3 as a one-column matrix
```
## Square, triangular, diagonal and identity matrices
A **square matrix** is a matrix that has the same number of rows and columns, for example a $3 \times 3$ matrix:
\begin{bmatrix}
4 & 9 & 2 \\
3 & 5 & 7 \\
8 & 1 & 6
\end{bmatrix}
An **upper triangular matrix** is a special kind of square matrix where all the elements *below* the main diagonal (top-left to bottom-right) are zero, for example:
\begin{bmatrix}
4 & 9 & 2 \\
0 & 5 & 7 \\
0 & 0 & 6
\end{bmatrix}
Similarly, a **lower triangular matrix** is a square matrix where all elements *above* the main diagonal are zero, for example:
\begin{bmatrix}
4 & 0 & 0 \\
3 & 5 & 0 \\
8 & 1 & 6
\end{bmatrix}
A **triangular matrix** is one that is either lower triangular or upper triangular.
A matrix that is both upper and lower triangular is called a **diagonal matrix**, for example:
\begin{bmatrix}
4 & 0 & 0 \\
0 & 5 & 0 \\
0 & 0 & 6
\end{bmatrix}
You can construct a diagonal matrix using NumPy's `diag` function:
```
np.diag([4, 5, 6])
```
If you pass a matrix to the `diag` function, it will happily extract the diagonal values:
```
D = np.array([
[1, 2, 3],
[4, 5, 6],
[7, 8, 9],
])
np.diag(D)
```
Finally, the **identity matrix** of size $n$, noted $I_n$, is a diagonal matrix of size $n \times n$ with $1$'s in the main diagonal, for example $I_3$:
\begin{bmatrix}
1 & 0 & 0 \\
0 & 1 & 0 \\
0 & 0 & 1
\end{bmatrix}
Numpy's `eye` function returns the identity matrix of the desired size:
```
np.eye(3)
```
The identity matrix is often noted simply $I$ (instead of $I_n$) when its size is clear given the context. It is called the *identity* matrix because multiplying a matrix with it leaves the matrix unchanged as we will see below.
## Adding matrices
If two matrices $Q$ and $R$ have the same size $m \times n$, they can be added together. Addition is performed *elementwise*: the result is also a $m \times n$ matrix $S$ where each element is the sum of the elements at the corresponding position: $S_{i,j} = Q_{i,j} + R_{i,j}$
$S =
\begin{bmatrix}
Q_{11} + R_{11} & Q_{12} + R_{12} & Q_{13} + R_{13} & \cdots & Q_{1n} + R_{1n} \\
Q_{21} + R_{21} & Q_{22} + R_{22} & Q_{23} + R_{23} & \cdots & Q_{2n} + R_{2n} \\
Q_{31} + R_{31} & Q_{32} + R_{32} & Q_{33} + R_{33} & \cdots & Q_{3n} + R_{3n} \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
Q_{m1} + R_{m1} & Q_{m2} + R_{m2} & Q_{m3} + R_{m3} & \cdots & Q_{mn} + R_{mn} \\
\end{bmatrix}$
For example, let's create a $2 \times 3$ matrix $B$ and compute $A + B$:
```
B = np.array([[1,2,3], [4, 5, 6]])
B
A
A + B
```
**Addition is *commutative***, meaning that $A + B = B + A$:
```
B + A
```
**It is also *associative***, meaning that $A + (B + C) = (A + B) + C$:
```
C = np.array([[100,200,300], [400, 500, 600]])
A + (B + C)
(A + B) + C
```
## Scalar multiplication
A matrix $M$ can be multiplied by a scalar $\lambda$. The result is noted $\lambda M$, and it is a matrix of the same size as $M$ with all elements multiplied by $\lambda$:
$\lambda M =
\begin{bmatrix}
\lambda \times M_{11} & \lambda \times M_{12} & \lambda \times M_{13} & \cdots & \lambda \times M_{1n} \\
\lambda \times M_{21} & \lambda \times M_{22} & \lambda \times M_{23} & \cdots & \lambda \times M_{2n} \\
\lambda \times M_{31} & \lambda \times M_{32} & \lambda \times M_{33} & \cdots & \lambda \times M_{3n} \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
\lambda \times M_{m1} & \lambda \times M_{m2} & \lambda \times M_{m3} & \cdots & \lambda \times M_{mn} \\
\end{bmatrix}$
A more concise way of writing this is:
$(\lambda M)_{i,j} = \lambda (M)_{i,j}$
In NumPy, simply use the `*` operator to multiply a matrix by a scalar. For example:
```
2 * A
```
Scalar multiplication is also defined on the right hand side, and gives the same result: $M \lambda = \lambda M$. For example:
```
A * 2
```
This makes scalar multiplication **commutative**.
It is also **associative**, meaning that $\alpha (\beta M) = (\alpha \times \beta) M$, where $\alpha$ and $\beta$ are scalars. For example:
```
2 * (3 * A)
(2 * 3) * A
```
Finally, it is **distributive over addition** of matrices, meaning that $\lambda (Q + R) = \lambda Q + \lambda R$:
```
2 * (A + B)
2 * A + 2 * B
```
## Matrix multiplication
So far, matrix operations have been rather intuitive. But multiplying matrices is a bit more involved.
A matrix $Q$ of size $m \times n$ can be multiplied by a matrix $R$ of size $n \times q$. It is noted simply $QR$ without multiplication sign or dot. The result $P$ is an $m \times q$ matrix where each element is computed as a sum of products:
$P_{i,j} = \sum_{k=1}^n{Q_{i,k} \times R_{k,j}}$
The element at position $i,j$ in the resulting matrix is the sum of the products of elements in row $i$ of matrix $Q$ by the elements in column $j$ of matrix $R$.
$P =
\begin{bmatrix}
Q_{11} R_{11} + Q_{12} R_{21} + \cdots + Q_{1n} R_{n1} &
Q_{11} R_{12} + Q_{12} R_{22} + \cdots + Q_{1n} R_{n2} &
\cdots &
Q_{11} R_{1q} + Q_{12} R_{2q} + \cdots + Q_{1n} R_{nq} \\
Q_{21} R_{11} + Q_{22} R_{21} + \cdots + Q_{2n} R_{n1} &
Q_{21} R_{12} + Q_{22} R_{22} + \cdots + Q_{2n} R_{n2} &
\cdots &
Q_{21} R_{1q} + Q_{22} R_{2q} + \cdots + Q_{2n} R_{nq} \\
\vdots & \vdots & \ddots & \vdots \\
Q_{m1} R_{11} + Q_{m2} R_{21} + \cdots + Q_{mn} R_{n1} &
Q_{m1} R_{12} + Q_{m2} R_{22} + \cdots + Q_{mn} R_{n2} &
\cdots &
Q_{m1} R_{1q} + Q_{m2} R_{2q} + \cdots + Q_{mn} R_{nq}
\end{bmatrix}$
You may notice that each element $P_{i,j}$ is the dot product of the row vector $Q_{i,*}$ and the column vector $R_{*,j}$:
$P_{i,j} = Q_{i,*} \cdot R_{*,j}$
So we can rewrite $P$ more concisely as:
$P =
\begin{bmatrix}
Q_{1,*} \cdot R_{*,1} & Q_{1,*} \cdot R_{*,2} & \cdots & Q_{1,*} \cdot R_{*,q} \\
Q_{2,*} \cdot R_{*,1} & Q_{2,*} \cdot R_{*,2} & \cdots & Q_{2,*} \cdot R_{*,q} \\
\vdots & \vdots & \ddots & \vdots \\
Q_{m,*} \cdot R_{*,1} & Q_{m,*} \cdot R_{*,2} & \cdots & Q_{m,*} \cdot R_{*,q}
\end{bmatrix}$
Let's multiply two matrices in NumPy, using `ndarray`'s `dot` method:
$E = AD = \begin{bmatrix}
10 & 20 & 30 \\
40 & 50 & 60
\end{bmatrix}
\begin{bmatrix}
2 & 3 & 5 & 7 \\
11 & 13 & 17 & 19 \\
23 & 29 & 31 & 37
\end{bmatrix} =
\begin{bmatrix}
930 & 1160 & 1320 & 1560 \\
2010 & 2510 & 2910 & 3450
\end{bmatrix}$
```
D = np.array([
[ 2, 3, 5, 7],
[11, 13, 17, 19],
[23, 29, 31, 37]
])
E = A.dot(D)
E
```
Let's check this result by looking at one element, just to be sure: looking at $E_{2,3}$ for example, we need to multiply elements in $A$'s $2^{nd}$ row by elements in $D$'s $3^{rd}$ column, and sum up these products:
```
40*5 + 50*17 + 60*31
E[1,2] # row 2, column 3
```
Looks good! You can check the other elements until you get used to the algorithm.
We multiplied a $2 \times 3$ matrix by a $3 \times 4$ matrix, so the result is a $2 \times 4$ matrix. The first matrix's number of columns has to be equal to the second matrix's number of rows. If we try to multiple $D$ by $A$, we get an error because D has 4 columns while A has 2 rows:
```
try:
D.dot(A)
except ValueError as e:
print("ValueError:", e)
```
This illustrates the fact that **matrix multiplication is *NOT* commutative**: in general $QR ≠ RQ$
In fact, $QR$ and $RQ$ are only *both* defined if $Q$ has size $m \times n$ and $R$ has size $n \times m$. Let's look at an example where both *are* defined and show that they are (in general) *NOT* equal:
```
F = np.array([
[5,2],
[4,1],
[9,3]
])
A.dot(F)
F.dot(A)
```
On the other hand, **matrix multiplication *is* associative**, meaning that $Q(RS) = (QR)S$. Let's create a $4 \times 5$ matrix $G$ to illustrate this:
```
G = np.array([
[8, 7, 4, 2, 5],
[2, 5, 1, 0, 5],
[9, 11, 17, 21, 0],
[0, 1, 0, 1, 2]])
A.dot(D).dot(G) # (AB)G
A.dot(D.dot(G)) # A(BG)
```
It is also ***distributive* over addition** of matrices, meaning that $(Q + R)S = QS + RS$. For example:
```
(A + B).dot(D)
A.dot(D) + B.dot(D)
```
The product of a matrix $M$ by the identity matrix (of matching size) results in the same matrix $M$. More formally, if $M$ is an $m \times n$ matrix, then:
$M I_n = I_m M = M$
This is generally written more concisely (since the size of the identity matrices is unambiguous given the context):
$MI = IM = M$
For example:
```
A.dot(np.eye(3))
np.eye(2).dot(A)
```
**Caution**: NumPy's `*` operator performs elementwise multiplication, *NOT* a matrix multiplication:
```
A * B # NOT a matrix multiplication
```
**The @ infix operator**
Python 3.5 [introduced](https://docs.python.org/3/whatsnew/3.5.html#pep-465-a-dedicated-infix-operator-for-matrix-multiplication) the `@` infix operator for matrix multiplication, and NumPy 1.10 added support for it. If you are using Python 3.5+ and NumPy 1.10+, you can simply write `A @ D` instead of `A.dot(D)`, making your code much more readable (but less portable). This operator also works for vector dot products.
```
import sys
print("Python version: {}.{}.{}".format(*sys.version_info))
print("Numpy version:", np.version.version)
# Uncomment the following line if your Python version is ≥3.5
# and your NumPy version is ≥1.10:
#A @ D
```
Note: `Q @ R` is actually equivalent to `Q.__matmul__(R)` which is implemented by NumPy as `np.matmul(Q, R)`, not as `Q.dot(R)`. The main difference is that `matmul` does not support scalar multiplication, while `dot` does, so you can write `Q.dot(3)`, which is equivalent to `Q * 3`, but you cannot write `Q @ 3` ([more details](http://stackoverflow.com/a/34142617/38626)).
## Matrix transpose
The transpose of a matrix $M$ is a matrix noted $M^T$ such that the $i^{th}$ row in $M^T$ is equal to the $i^{th}$ column in $M$:
$ A^T =
\begin{bmatrix}
10 & 20 & 30 \\
40 & 50 & 60
\end{bmatrix}^T =
\begin{bmatrix}
10 & 40 \\
20 & 50 \\
30 & 60
\end{bmatrix}$
In other words, ($A^T)_{i,j}$ = $A_{j,i}$
Obviously, if $M$ is an $m \times n$ matrix, then $M^T$ is an $n \times m$ matrix.
Note: there are a few other notations, such as $M^t$, $M′$, or ${^t}M$.
In NumPy, a matrix's transpose can be obtained simply using the `T` attribute:
```
A
A.T
```
As you might expect, transposing a matrix twice returns the original matrix:
```
A.T.T
```
Transposition is distributive over addition of matrices, meaning that $(Q + R)^T = Q^T + R^T$. For example:
```
(A + B).T
A.T + B.T
```
Moreover, $(Q \cdot R)^T = R^T \cdot Q^T$. Note that the order is reversed. For example:
```
(A.dot(D)).T
D.T.dot(A.T)
```
A **symmetric matrix** $M$ is defined as a matrix that is equal to its transpose: $M^T = M$. This definition implies that it must be a square matrix whose elements are symmetric relative to the main diagonal, for example:
\begin{bmatrix}
17 & 22 & 27 & 49 \\
22 & 29 & 36 & 0 \\
27 & 36 & 45 & 2 \\
49 & 0 & 2 & 99
\end{bmatrix}
The product of a matrix by its transpose is always a symmetric matrix, for example:
```
D.dot(D.T)
```
## Converting 1D arrays to 2D arrays in NumPy
As we mentionned earlier, in NumPy (as opposed to Matlab, for example), 1D really means 1D: there is no such thing as a vertical 1D-array or a horizontal 1D-array. So you should not be surprised to see that transposing a 1D array does not do anything:
```
u
u.T
```
We want to convert $\textbf{u}$ into a row vector before transposing it. There are a few ways to do this:
```
u_row = np.array([u])
u_row
```
Notice the extra square brackets: this is a 2D array with just one row (ie. a 1x2 matrix). In other words it really is a **row vector**.
```
u[np.newaxis, :]
```
This quite explicit: we are asking for a new vertical axis, keeping the existing data as the horizontal axis.
```
u[np.newaxis]
```
This is equivalent, but a little less explicit.
```
u[None]
```
This is the shortest version, but you probably want to avoid it because it is unclear. The reason it works is that `np.newaxis` is actually equal to `None`, so this is equivalent to the previous version.
Ok, now let's transpose our row vector:
```
u_row.T
```
Great! We now have a nice **column vector**.
Rather than creating a row vector then transposing it, it is also possible to convert a 1D array directly into a column vector:
```
u[:, np.newaxis]
```
## Plotting a matrix
We have already seen that vectors can been represented as points or arrows in N-dimensional space. Is there a good graphical representation of matrices? Well you can simply see a matrix as a list of vectors, so plotting a matrix results in many points or arrows. For example, let's create a $2 \times 4$ matrix `P` and plot it as points:
```
P = np.array([
[3.0, 4.0, 1.0, 4.6],
[0.2, 3.5, 2.0, 0.5]
])
x_coords_P, y_coords_P = P
plt.scatter(x_coords_P, y_coords_P)
plt.axis([0, 5, 0, 4])
plt.show()
```
Of course we could also have stored the same 4 vectors as row vectors instead of column vectors, resulting in a $4 \times 2$ matrix (the transpose of $P$, in fact). It is really an arbitrary choice.
Since the vectors are ordered, you can see the matrix as a path and represent it with connected dots:
```
plt.plot(x_coords_P, y_coords_P, "bo")
plt.plot(x_coords_P, y_coords_P, "b--")
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
Or you can represent it as a polygon: matplotlib's `Polygon` class expects an $n \times 2$ NumPy array, not a $2 \times n$ array, so we just need to give it $P^T$:
```
from matplotlib.patches import Polygon
plt.gca().add_artist(Polygon(P.T))
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
## Geometric applications of matrix operations
We saw earlier that vector addition results in a geometric translation, vector multiplication by a scalar results in rescaling (zooming in or out, centered on the origin), and vector dot product results in projecting a vector onto another vector, rescaling and measuring the resulting coordinate.
Similarly, matrix operations have very useful geometric applications.
### Addition = multiple geometric translations
First, adding two matrices together is equivalent to adding all their vectors together. For example, let's create a $2 \times 4$ matrix $H$ and add it to $P$, and look at the result:
```
H = np.array([
[ 0.5, -0.2, 0.2, -0.1],
[ 0.4, 0.4, 1.5, 0.6]
])
P_moved = P + H
plt.gca().add_artist(Polygon(P.T, alpha=0.2))
plt.gca().add_artist(Polygon(P_moved.T, alpha=0.3, color="r"))
for vector, origin in zip(H.T, P.T):
plot_vector2d(vector, origin=origin)
plt.text(2.2, 1.8, "$P$", color="b", fontsize=18)
plt.text(2.0, 3.2, "$P+H$", color="r", fontsize=18)
plt.text(2.5, 0.5, "$H_{*,1}$", color="k", fontsize=18)
plt.text(4.1, 3.5, "$H_{*,2}$", color="k", fontsize=18)
plt.text(0.4, 2.6, "$H_{*,3}$", color="k", fontsize=18)
plt.text(4.4, 0.2, "$H_{*,4}$", color="k", fontsize=18)
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
If we add a matrix full of identical vectors, we get a simple geometric translation:
```
H2 = np.array([
[-0.5, -0.5, -0.5, -0.5],
[ 0.4, 0.4, 0.4, 0.4]
])
P_translated = P + H2
plt.gca().add_artist(Polygon(P.T, alpha=0.2))
plt.gca().add_artist(Polygon(P_translated.T, alpha=0.3, color="r"))
for vector, origin in zip(H2.T, P.T):
plot_vector2d(vector, origin=origin)
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
Although matrices can only be added together if they have the same size, NumPy allows adding a row vector or a column vector to a matrix: this is called *broadcasting* and is explained in further details in the [NumPy tutorial](tools_numpy.ipynb). We could have obtained the same result as above with:
```
P + [[-0.5], [0.4]] # same as P + H2, thanks to NumPy broadcasting
```
### Scalar multiplication
Multiplying a matrix by a scalar results in all its vectors being multiplied by that scalar, so unsurprisingly, the geometric result is a rescaling of the entire figure. For example, let's rescale our polygon by a factor of 60% (zooming out, centered on the origin):
```
def plot_transformation(P_before, P_after, text_before, text_after, axis = [0, 5, 0, 4], arrows=False):
if arrows:
for vector_before, vector_after in zip(P_before.T, P_after.T):
plot_vector2d(vector_before, color="blue", linestyle="--")
plot_vector2d(vector_after, color="red", linestyle="-")
plt.gca().add_artist(Polygon(P_before.T, alpha=0.2))
plt.gca().add_artist(Polygon(P_after.T, alpha=0.3, color="r"))
plt.text(P_before[0].mean(), P_before[1].mean(), text_before, fontsize=18, color="blue")
plt.text(P_after[0].mean(), P_after[1].mean(), text_after, fontsize=18, color="red")
plt.axis(axis)
plt.grid()
P_rescaled = 0.60 * P
plot_transformation(P, P_rescaled, "$P$", "$0.6 P$", arrows=True)
plt.show()
```
### Matrix multiplication – Projection onto an axis
Matrix multiplication is more complex to visualize, but it is also the most powerful tool in the box.
Let's start simple, by defining a $1 \times 2$ matrix $U = \begin{bmatrix} 1 & 0 \end{bmatrix}$. This row vector is just the horizontal unit vector.
```
U = np.array([[1, 0]])
```
Now let's look at the dot product $U \cdot P$:
```
U.dot(P)
```
These are the horizontal coordinates of the vectors in $P$. In other words, we just projected $P$ onto the horizontal axis:
```
def plot_projection(U, P):
U_P = U.dot(P)
axis_end = 100 * U
plot_vector2d(axis_end[0], color="black")
plt.gca().add_artist(Polygon(P.T, alpha=0.2))
for vector, proj_coordinate in zip(P.T, U_P.T):
proj_point = proj_coordinate * U
plt.plot(proj_point[0][0], proj_point[0][1], "ro")
plt.plot([vector[0], proj_point[0][0]], [vector[1], proj_point[0][1]], "r--")
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
plot_projection(U, P)
```
We can actually project on any other axis by just replacing $U$ with any other unit vector. For example, let's project on the axis that is at a 30° angle above the horizontal axis:
```
angle30 = 30 * np.pi / 180 # angle in radians
U_30 = np.array([[np.cos(angle30), np.sin(angle30)]])
plot_projection(U_30, P)
```
Good! Remember that the dot product of a unit vector and a matrix basically performs a projection on an axis and gives us the coordinates of the resulting points on that axis.
### Matrix multiplication – Rotation
Now let's create a $2 \times 2$ matrix $V$ containing two unit vectors that make 30° and 120° angles with the horizontal axis:
$V = \begin{bmatrix} \cos(30°) & \sin(30°) \\ \cos(120°) & \sin(120°) \end{bmatrix}$
```
angle120 = 120 * np.pi / 180
V = np.array([
[np.cos(angle30), np.sin(angle30)],
[np.cos(angle120), np.sin(angle120)]
])
V
```
Let's look at the product $VP$:
```
V.dot(P)
```
The first row is equal to $V_{1,*} P$, which is the coordinates of the projection of $P$ onto the 30° axis, as we have seen above. The second row is $V_{2,*} P$, which is the coordinates of the projection of $P$ onto the 120° axis. So basically we obtained the coordinates of $P$ after rotating the horizontal and vertical axes by 30° (or equivalently after rotating the polygon by -30° around the origin)! Let's plot $VP$ to see this:
```
P_rotated = V.dot(P)
plot_transformation(P, P_rotated, "$P$", "$VP$", [-2, 6, -2, 4], arrows=True)
plt.show()
```
Matrix $V$ is called a **rotation matrix**.
### Matrix multiplication – Other linear transformations
More generally, any linear transformation $f$ that maps n-dimensional vectors to m-dimensional vectors can be represented as an $m \times n$ matrix. For example, say $\textbf{u}$ is a 3-dimensional vector:
$\textbf{u} = \begin{pmatrix} x \\ y \\ z \end{pmatrix}$
and $f$ is defined as:
$f(\textbf{u}) = \begin{pmatrix}
ax + by + cz \\
dx + ey + fz
\end{pmatrix}$
This transormation $f$ maps 3-dimensional vectors to 2-dimensional vectors in a linear way (ie. the resulting coordinates only involve sums of multiples of the original coordinates). We can represent this transformation as matrix $F$:
$F = \begin{bmatrix}
a & b & c \\
d & e & f
\end{bmatrix}$
Now, to compute $f(\textbf{u})$ we can simply do a matrix multiplication:
$f(\textbf{u}) = F \textbf{u}$
If we have a matric $G = \begin{bmatrix}\textbf{u}_1 & \textbf{u}_2 & \cdots & \textbf{u}_q \end{bmatrix}$, where each $\textbf{u}_i$ is a 3-dimensional column vector, then $FG$ results in the linear transformation of all vectors $\textbf{u}_i$ as defined by the matrix $F$:
$FG = \begin{bmatrix}f(\textbf{u}_1) & f(\textbf{u}_2) & \cdots & f(\textbf{u}_q) \end{bmatrix}$
To summarize, the matrix on the left hand side of a dot product specifies what linear transormation to apply to the right hand side vectors. We have already shown that this can be used to perform projections and rotations, but any other linear transformation is possible. For example, here is a transformation known as a *shear mapping*:
```
F_shear = np.array([
[1, 1.5],
[0, 1]
])
plot_transformation(P, F_shear.dot(P), "$P$", "$F_{shear} P$",
axis=[0, 10, 0, 7])
plt.show()
```
Let's look at how this transformation affects the **unit square**:
```
Square = np.array([
[0, 0, 1, 1],
[0, 1, 1, 0]
])
plot_transformation(Square, F_shear.dot(Square), "$Square$", "$F_{shear} Square$",
axis=[0, 2.6, 0, 1.8])
plt.show()
```
Now let's look at a **squeeze mapping**:
```
F_squeeze = np.array([
[1.4, 0],
[0, 1/1.4]
])
plot_transformation(P, F_squeeze.dot(P), "$P$", "$F_{squeeze} P$",
axis=[0, 7, 0, 5])
plt.show()
```
The effect on the unit square is:
```
plot_transformation(Square, F_squeeze.dot(Square), "$Square$", "$F_{squeeze} Square$",
axis=[0, 1.8, 0, 1.2])
plt.show()
```
Let's show a last one: reflection through the horizontal axis:
```
F_reflect = np.array([
[1, 0],
[0, -1]
])
plot_transformation(P, F_reflect.dot(P), "$P$", "$F_{reflect} P$",
axis=[-2, 9, -4.5, 4.5])
plt.show()
```
## Matrix inverse
Now that we understand that a matrix can represent any linear transformation, a natural question is: can we find a transformation matrix that reverses the effect of a given transformation matrix $F$? The answer is yes… sometimes! When it exists, such a matrix is called the **inverse** of $F$, and it is noted $F^{-1}$.
For example, the rotation, the shear mapping and the squeeze mapping above all have inverse transformations. Let's demonstrate this on the shear mapping:
```
F_inv_shear = np.array([
[1, -1.5],
[0, 1]
])
P_sheared = F_shear.dot(P)
P_unsheared = F_inv_shear.dot(P_sheared)
plot_transformation(P_sheared, P_unsheared, "$P_{sheared}$", "$P_{unsheared}$",
axis=[0, 10, 0, 7])
plt.plot(P[0], P[1], "b--")
plt.show()
```
We applied a shear mapping on $P$, just like we did before, but then we applied a second transformation to the result, and *lo and behold* this had the effect of coming back to the original $P$ (we plotted the original $P$'s outline to double check). The second transformation is the inverse of the first one.
We defined the inverse matrix $F_{shear}^{-1}$ manually this time, but NumPy provides an `inv` function to compute a matrix's inverse, so we could have written instead:
```
F_inv_shear = LA.inv(F_shear)
F_inv_shear
```
Only square matrices can be inversed. This makes sense when you think about it: if you have a transformation that reduces the number of dimensions, then some information is lost and there is no way that you can get it back. For example say you use a $2 \times 3$ matrix to project a 3D object onto a plane. The result may look like this:
```
plt.plot([0, 0, 1, 1, 0, 0.1, 0.1, 0, 0.1, 1.1, 1.0, 1.1, 1.1, 1.0, 1.1, 0.1],
[0, 1, 1, 0, 0, 0.1, 1.1, 1.0, 1.1, 1.1, 1.0, 1.1, 0.1, 0, 0.1, 0.1],
"r-")
plt.axis([-0.5, 2.1, -0.5, 1.5])
plt.show()
```
Looking at this image, it is impossible to tell whether this is the projection of a cube or the projection of a narrow rectangular object. Some information has been lost in the projection.
Even square transformation matrices can lose information. For example, consider this transformation matrix:
```
F_project = np.array([
[1, 0],
[0, 0]
])
plot_transformation(P, F_project.dot(P), "$P$", "$F_{project} \cdot P$",
axis=[0, 6, -1, 4])
plt.show()
```
This transformation matrix performs a projection onto the horizontal axis. Our polygon gets entirely flattened out so some information is entirely lost and it is impossible to go back to the original polygon using a linear transformation. In other words, $F_{project}$ has no inverse. Such a square matrix that cannot be inversed is called a **singular matrix** (aka degenerate matrix). If we ask NumPy to calculate its inverse, it raises an exception:
```
try:
LA.inv(F_project)
except LA.LinAlgError as e:
print("LinAlgError:", e)
```
Here is another example of a singular matrix. This one performs a projection onto the axis at a 30° angle above the horizontal axis:
```
angle30 = 30 * np.pi / 180
F_project_30 = np.array([
[np.cos(angle30)**2, np.sin(2*angle30)/2],
[np.sin(2*angle30)/2, np.sin(angle30)**2]
])
plot_transformation(P, F_project_30.dot(P), "$P$", "$F_{project\_30} \cdot P$",
axis=[0, 6, -1, 4])
plt.show()
```
But this time, due to floating point rounding errors, NumPy manages to calculate an inverse (notice how large the elements are, though):
```
LA.inv(F_project_30)
```
As you might expect, the dot product of a matrix by its inverse results in the identity matrix:
$M \cdot M^{-1} = M^{-1} \cdot M = I$
This makes sense since doing a linear transformation followed by the inverse transformation results in no change at all.
```
F_shear.dot(LA.inv(F_shear))
```
Another way to express this is that the inverse of the inverse of a matrix $M$ is $M$ itself:
$((M)^{-1})^{-1} = M$
```
LA.inv(LA.inv(F_shear))
```
Also, the inverse of scaling by a factor of $\lambda$ is of course scaling by a factor or $\frac{1}{\lambda}$:
$ (\lambda \times M)^{-1} = \frac{1}{\lambda} \times M^{-1}$
Once you understand the geometric interpretation of matrices as linear transformations, most of these properties seem fairly intuitive.
A matrix that is its own inverse is called an **involution**. The simplest examples are reflection matrices, or a rotation by 180°, but there are also more complex involutions, for example imagine a transformation that squeezes horizontally, then reflects over the vertical axis and finally rotates by 90° clockwise. Pick up a napkin and try doing that twice: you will end up in the original position. Here is the corresponding involutory matrix:
```
F_involution = np.array([
[0, -2],
[-1/2, 0]
])
plot_transformation(P, F_involution.dot(P), "$P$", "$F_{involution} \cdot P$",
axis=[-8, 5, -4, 4])
plt.show()
```
Finally, a square matrix $H$ whose inverse is its own transpose is an **orthogonal matrix**:
$H^{-1} = H^T$
Therefore:
$H \cdot H^T = H^T \cdot H = I$
It corresponds to a transformation that preserves distances, such as rotations and reflections, and combinations of these, but not rescaling, shearing or squeezing. Let's check that $F_{reflect}$ is indeed orthogonal:
```
F_reflect.dot(F_reflect.T)
```
## Determinant
The determinant of a square matrix $M$, noted $\det(M)$ or $\det M$ or $|M|$ is a value that can be calculated from its elements $(M_{i,j})$ using various equivalent methods. One of the simplest methods is this recursive approach:
$|M| = M_{1,1}\times|M^{(1,1)}| - M_{2,1}\times|M^{(2,1)}| + M_{3,1}\times|M^{(3,1)}| - M_{4,1}\times|M^{(4,1)}| + \cdots ± M_{n,1}\times|M^{(n,1)}|$
* Where $M^{(i,j)}$ is the matrix $M$ without row $i$ and column $j$.
For example, let's calculate the determinant of the following $3 \times 3$ matrix:
$M = \begin{bmatrix}
1 & 2 & 3 \\
4 & 5 & 6 \\
7 & 8 & 0
\end{bmatrix}$
Using the method above, we get:
$|M| = 1 \times \left | \begin{bmatrix} 5 & 6 \\ 8 & 0 \end{bmatrix} \right |
- 2 \times \left | \begin{bmatrix} 4 & 6 \\ 7 & 0 \end{bmatrix} \right |
+ 3 \times \left | \begin{bmatrix} 4 & 5 \\ 7 & 8 \end{bmatrix} \right |$
Now we need to compute the determinant of each of these $2 \times 2$ matrices (these determinants are called **minors**):
$\left | \begin{bmatrix} 5 & 6 \\ 8 & 0 \end{bmatrix} \right | = 5 \times 0 - 6 \times 8 = -48$
$\left | \begin{bmatrix} 4 & 6 \\ 7 & 0 \end{bmatrix} \right | = 4 \times 0 - 6 \times 7 = -42$
$\left | \begin{bmatrix} 4 & 5 \\ 7 & 8 \end{bmatrix} \right | = 4 \times 8 - 5 \times 7 = -3$
Now we can calculate the final result:
$|M| = 1 \times (-48) - 2 \times (-42) + 3 \times (-3) = 27$
To get the determinant of a matrix, you can call NumPy's `det` function in the `numpy.linalg` module:
```
M = np.array([
[1, 2, 3],
[4, 5, 6],
[7, 8, 0]
])
LA.det(M)
```
One of the main uses of the determinant is to *determine* whether a square matrix can be inversed or not: if the determinant is equal to 0, then the matrix *cannot* be inversed (it is a singular matrix), and if the determinant is not 0, then it *can* be inversed.
For example, let's compute the determinant for the $F_{project}$, $F_{project\_30}$ and $F_{shear}$ matrices that we defined earlier:
```
LA.det(F_project)
```
That's right, $F_{project}$ is singular, as we saw earlier.
```
LA.det(F_project_30)
```
This determinant is suspiciously close to 0: it really should be 0, but it's not due to tiny floating point errors. The matrix is actually singular.
```
LA.det(F_shear)
```
Perfect! This matrix *can* be inversed as we saw earlier. Wow, math really works!
The determinant can also be used to measure how much a linear transformation affects surface areas: for example, the projection matrices $F_{project}$ and $F_{project\_30}$ completely flatten the polygon $P$, until its area is zero. This is why the determinant of these matrices is 0. The shear mapping modified the shape of the polygon, but it did not affect its surface area, which is why the determinant is 1. You can try computing the determinant of a rotation matrix, and you should also find 1. What about a scaling matrix? Let's see:
```
F_scale = np.array([
[0.5, 0],
[0, 0.5]
])
plot_transformation(P, F_scale.dot(P), "$P$", "$F_{scale} \cdot P$",
axis=[0, 6, -1, 4])
plt.show()
```
We rescaled the polygon by a factor of 1/2 on both vertical and horizontal axes so the surface area of the resulting polygon is 1/4$^{th}$ of the original polygon. Let's compute the determinant and check that:
```
LA.det(F_scale)
```
Correct!
The determinant can actually be negative, when the transformation results in a "flipped over" version of the original polygon (eg. a left hand glove becomes a right hand glove). For example, the determinant of the `F_reflect` matrix is -1 because the surface area is preserved but the polygon gets flipped over:
```
LA.det(F_reflect)
```
## Composing linear transformations
Several linear transformations can be chained simply by performing multiple dot products in a row. For example, to perform a squeeze mapping followed by a shear mapping, just write:
```
P_squeezed_then_sheared = F_shear.dot(F_squeeze.dot(P))
```
Since the dot product is associative, the following code is equivalent:
```
P_squeezed_then_sheared = (F_shear.dot(F_squeeze)).dot(P)
```
Note that the order of the transformations is the reverse of the dot product order.
If we are going to perform this composition of linear transformations more than once, we might as well save the composition matrix like this:
```
F_squeeze_then_shear = F_shear.dot(F_squeeze)
P_squeezed_then_sheared = F_squeeze_then_shear.dot(P)
```
From now on we can perform both transformations in just one dot product, which can lead to a very significant performance boost.
What if you want to perform the inverse of this double transformation? Well, if you squeezed and then you sheared, and you want to undo what you have done, it should be obvious that you should unshear first and then unsqueeze. In more mathematical terms, given two invertible (aka nonsingular) matrices $Q$ and $R$:
$(Q \cdot R)^{-1} = R^{-1} \cdot Q^{-1}$
And in NumPy:
```
LA.inv(F_shear.dot(F_squeeze)) == LA.inv(F_squeeze).dot(LA.inv(F_shear))
```
## Singular Value Decomposition
It turns out that any $m \times n$ matrix $M$ can be decomposed into the dot product of three simple matrices:
* a rotation matrix $U$ (an $m \times m$ orthogonal matrix)
* a scaling & projecting matrix $\Sigma$ (an $m \times n$ diagonal matrix)
* and another rotation matrix $V^T$ (an $n \times n$ orthogonal matrix)
$M = U \cdot \Sigma \cdot V^{T}$
For example, let's decompose the shear transformation:
```
U, S_diag, V_T = LA.svd(F_shear) # note: in python 3 you can rename S_diag to Σ_diag
U
S_diag
```
Note that this is just a 1D array containing the diagonal values of Σ. To get the actual matrix Σ, we can use NumPy's `diag` function:
```
S = np.diag(S_diag)
S
```
Now let's check that $U \cdot \Sigma \cdot V^T$ is indeed equal to `F_shear`:
```
U.dot(np.diag(S_diag)).dot(V_T)
F_shear
```
It worked like a charm. Let's apply these transformations one by one (in reverse order) on the unit square to understand what's going on. First, let's apply the first rotation $V^T$:
```
plot_transformation(Square, V_T.dot(Square), "$Square$", "$V^T \cdot Square$",
axis=[-0.5, 3.5 , -1.5, 1.5])
plt.show()
```
Now let's rescale along the vertical and horizontal axes using $\Sigma$:
```
plot_transformation(V_T.dot(Square), S.dot(V_T).dot(Square), "$V^T \cdot Square$", "$\Sigma \cdot V^T \cdot Square$",
axis=[-0.5, 3.5 , -1.5, 1.5])
plt.show()
```
Finally, we apply the second rotation $U$:
```
plot_transformation(S.dot(V_T).dot(Square), U.dot(S).dot(V_T).dot(Square),"$\Sigma \cdot V^T \cdot Square$", "$U \cdot \Sigma \cdot V^T \cdot Square$",
axis=[-0.5, 3.5 , -1.5, 1.5])
plt.show()
```
And we can see that the result is indeed a shear mapping of the original unit square.
## Eigenvectors and eigenvalues
An **eigenvector** of a square matrix $M$ (also called a **characteristic vector**) is a non-zero vector that remains on the same line after transformation by the linear transformation associated with $M$. A more formal definition is any vector $v$ such that:
$M \cdot v = \lambda \times v$
Where $\lambda$ is a scalar value called the **eigenvalue** associated to the vector $v$.
For example, any horizontal vector remains horizontal after applying the shear mapping (as you can see on the image above), so it is an eigenvector of $M$. A vertical vector ends up tilted to the right, so vertical vectors are *NOT* eigenvectors of $M$.
If we look at the squeeze mapping, we find that any horizontal or vertical vector keeps its direction (although its length changes), so all horizontal and vertical vectors are eigenvectors of $F_{squeeze}$.
However, rotation matrices have no eigenvectors at all (except if the rotation angle is 0° or 180°, in which case all non-zero vectors are eigenvectors).
NumPy's `eig` function returns the list of unit eigenvectors and their corresponding eigenvalues for any square matrix. Let's look at the eigenvectors and eigenvalues of the squeeze mapping matrix $F_{squeeze}$:
```
eigenvalues, eigenvectors = LA.eig(F_squeeze)
eigenvalues # [λ0, λ1, …]
eigenvectors # [v0, v1, …]
```
Indeed the horizontal vectors are stretched by a factor of 1.4, and the vertical vectors are shrunk by a factor of 1/1.4=0.714…, so far so good. Let's look at the shear mapping matrix $F_{shear}$:
```
eigenvalues2, eigenvectors2 = LA.eig(F_shear)
eigenvalues2 # [λ0, λ1, …]
eigenvectors2 # [v0, v1, …]
```
Wait, what!? We expected just one unit eigenvector, not two. The second vector is almost equal to $\begin{pmatrix}-1 \\ 0 \end{pmatrix}$, which is on the same line as the first vector $\begin{pmatrix}1 \\ 0 \end{pmatrix}$. This is due to floating point errors. We can safely ignore vectors that are (almost) colinear (ie. on the same line).
## Trace
The trace of a square matrix $M$, noted $tr(M)$ is the sum of the values on its main diagonal. For example:
```
D = np.array([
[100, 200, 300],
[ 10, 20, 30],
[ 1, 2, 3],
])
np.trace(D)
```
The trace does not have a simple geometric interpretation (in general), but it has a number of properties that make it useful in many areas:
* $tr(A + B) = tr(A) + tr(B)$
* $tr(A \cdot B) = tr(B \cdot A)$
* $tr(A \cdot B \cdot \cdots \cdot Y \cdot Z) = tr(Z \cdot A \cdot B \cdot \cdots \cdot Y)$
* $tr(A^T \cdot B) = tr(A \cdot B^T) = tr(B^T \cdot A) = tr(B \cdot A^T) = \sum_{i,j}X_{i,j} \times Y_{i,j}$
* …
It does, however, have a useful geometric interpretation in the case of projection matrices (such as $F_{project}$ that we discussed earlier): it corresponds to the number of dimensions after projection. For example:
```
np.trace(F_project)
```
# What next?
This concludes this introduction to Linear Algebra. Although these basics cover most of what you will need to know for Machine Learning, if you wish to go deeper into this topic there are many options available: Linear Algebra [books](http://linear.axler.net/), [Khan Academy](https://www.khanacademy.org/math/linear-algebra) lessons, or just [Wikipedia](https://en.wikipedia.org/wiki/Linear_algebra) pages.
| github_jupyter |
# User's Guide, Chapter 4: Lists, Streams (I) and Output
In the last two chapters, I introduced the concept of
:class:`~music21.note.Note` objects which are made up of
:class:`~music21.pitch.Pitch` and :class:`~music21.duration.Duration`
objects, and we even displayed a Note on a staff and played it
via MIDI. But unless you're challenging Cage and Webern for the
status of least musical material, you will probably want to
analyze, manipulate, or create more than one Note.
Python has ways of working with multiple objects and `music21`
extends those ways to be more musical. Let's look at how
Python does it first and then see how `music21` extends these
ways. (If you've been programming for a bit, or especially
if you have Python experience, skip to the section on
Streams below after creating the objects `note1`, `note2` and
`note3` described below).
Say you have two notes, a C and an F# in the middle of the treble staff.
(If the concept of working with a tritone bothers you, go ahead and make
the second note a G; we won't mind; we'll just call you Pope
Gregory from now on). Lets create those notes:
## Working with multiple objects via Lists
```
from music21 import *
note1 = note.Note("C4")
note2 = note.Note("F#4")
```
Let's make the first note a half note by modifying its duration (by default
all `Note` objects are quarter notes):
```
note1.duration.type = 'half'
note1.duration.quarterLength
note2.duration.quarterLength
```
To print the `step` (that is, the name without any octave or accidental
information) of each of these notes, you could do something like this:
```
print(note1.step)
print(note2.step)
```
But suppose you had thirty notes? Then it'd be a pain to type "`print(noteX.step)`"
thirty times. Fortunately, there's a solution: we can put each of the
note objects into a `List` which is a built in Python object that stores multiple
other objects (like Notes or Chords, or even things like numbers). To create
a list in Python, put square brackets (`[]`) around the things that you want
to put in the list, separated by commas. Let's create a list called `noteList`
that contains note1 and note2:
```
noteList = [note1, note2]
```
We can check that `noteList` contains our Notes by printing it:
```
print(noteList)
```
The list is represented by the square brackets around the end with the comma in between them, just like how they were created originally. The act of creation is mirrored in the representation. That's nice. Medieval philosophers would approve.
Now we can write a two-line program that will print the step of each note in noteList. Most modern languages have a way of doing some action for each member ("element") in a list (also called an "array" or sometimes "row"). In Python this is the "for" command. When you type these lines, make sure to type the spaces at the start of the second line. (When you're done typing `print(thisNote.step)`, you'll probably have to hit enter twice to see the results.)
```
for thisNote in noteList:
print(thisNote.step)
```
What's happening here? What ``for thisNote in noteList:`` says is that Python should
take each note in noteList in order and temporarily call that note "`thisNote`" (you
could have it called anything you want; `myNote`, `n`, `currentNote` are all good
names, but `note` is not because `note` is the name of a module). Then the ":" at
the end of the line indicates that everything that happens for a bit will apply
to every `Note` in noteList one at a time. How does Python know when "a bit" is
over? Simple: every line that is a part of the loop needs to be indented by putting
in some spaces. (I usually use four spaces or hit tab. Some people use two spaces. Just be consistent.)
Loops don't save much time here, but imagine if noteList had dozens or hundreds
of Notes in it? Then the ability to do something to each object becomes more and
more important.
Let's add another note to noteList. First let's create another note, a low B-flat:
```
note3 = note.Note("B-2")
```
Then we'll append that note to the end of noteList:
```
noteList.append(note3)
```
Lists can be manipulated or changed. They are called "mutable" objects (we'll learn about immutable objects later). Streams, as we will see, can be manipulated the same way through `.append()`.
We can see that the length of noteList is now 3 using the `len()` function:
```
len(noteList)
```
And if we write our looping function again, we will get a third note:
```
for thisNote in noteList:
print(thisNote.step)
```
We can find out what the first note of noteList is by writing:
```
noteList[0]
```
Notice that in a list, the first element is `[0]`, not `[1]`. There are all
sorts of historical reasons why computers start counting lists with zero
rather than one--some good, some obsolete--but we need to live with this
if we're going to get any work done. Think of it like how floors are numbered
in European buildings compared to American buildings. If we go forward one note,
to the second note, we write:
```
noteList[1]
```
We can also ask `noteList` where is `note2` within it, using the `index()` method:
```
noteList.index(note2)
```
If we want to get the last element of a list, we can write:
```
noteList[-1]
```
Which is how basements are numbered in Europe as well. This is the same
element as `noteList[2]` (our third Note), as we can have Python prove:`
```
noteList[-1] is noteList[2]
```
Lists will become important tools in your programming, but they don't know
anything about music. To get some intelligence into our music we'll need to
know about a `music21` object similar to lists, called a :class:`~music21.stream.Stream`.
## Introduction to Streams
The :class:`~music21.stream.Stream` object and its subclasses (Score,
Part, Measure) are the fundamental containers for music21 objects such
as :class:`~music21.note.Note`, :class:`~music21.chord.Chord`,
:class:`~music21.clef.Clef`, :class:`~music21.meter.TimeSignature` objects.
A container is like a Python list (or an array in some languages).
Objects stored in a Stream are generally spaced in time; each stored object has
an offset usually representing how many quarter notes it lies from the beginning
of the Stream. For instance in a 4/4 measure of two half notes, the first note
will be at offset 0.0, and the second at offset 2.0.
Streams, further, can store other Streams, permitting a wide variety of nested,
ordered, and timed structures. These stored streams also have offsets. So if
we put two 4/4 Measure objects (subclasses of Stream) into a Part (also a
type of Stream), then the first measure will be at offset 0.0 and the second
measure will be at offset 4.0.
Commonly used subclasses of Streams include the :class:`~music21.stream.Score`,
:class:`~music21.stream.Part`, and :class:`~music21.stream.Measure`. It is
important to grasp that any time we want to collect and contain a group of
music21 objects, we put them into a Stream. Streams can also be used for
less conventional organizational structures. We frequently will build and pass
around short-lived, temporary Streams, since doing this opens up a wide variety
of tools for extracting, processing, and manipulating objects on the Stream.
For instance, if you are looking at only notes on beat 2 of any measure, you'll
probably want to put them into a Stream as well.
A critical feature of music21's design that distinguishes it from other
music analysis frameworks is that one music21 object can be
simultaneously stored (or, more accurately, referenced) in more than one Stream.
For examples, we might have numerous :class:`~music21.stream.Measure` Streams
contained in a :class:`~music21.stream.Part` Stream. If we extract a region of
this Part (using the :meth:`~music21.stream.Stream.measures` method), we get a
new Stream containing the specified Measures and the contained notes. We have
not actually created new
notes within these extracted measures; the output Stream simply has references
to the
same objects. Changes made to Notes in this output Stream will be simultaneously
reflected in Notes in the source Part. There is one limitation though:
the same object should not appear twice in one hierarchical structure of Streams.
For instance, you should not put a note object in both measure 3 and measure 5
of the same piece -- it can appear in measure 3 of one piece and measure 5 of
another piece. (For instance, if you wanted to track a particular note's context
in an original version of a score and an arrangement). Most users will never
need to worry about these details: just know that this feature lets music21
do some things that no other software package can do.
## Creating simple Streams
Objects stored in Streams are called elements and must be some type of Music21Object (don’t worry, almost everything in music21 is a Music21Object, such as Note, Chord, TimeSignature, etc.).
(If you want to put an object that's not a Music21Object in a Stream,
put it in an :class:`~music21.base.ElementWrapper`.)
Streams are similar to Python lists in that they hold individual elements
in order. They're different in that they can only hold `music21` objects
such as Notes or :class:`~music21.clef.Clef` objects. But they're a lot
smarter and more powerful.
To create a Stream you'll need to type `stream.Stream()` and assign it to
a variable using the equal sign. Let's call our Stream `stream1`:
```
stream1 = stream.Stream()
```
Notice that just like how the (capital) `Note` object lives in a module
called (lowercase) `note`, the (capital) `Stream` object lives in a module
called (lowercase) `stream`. Variable names, like `stream1` can be either
uppercase or lowercase, but I tend to use lowercase variable names (or camelCase
like we did with `noteList`).
The most common use of Streams is as places to store Notes. So let's do just that: we can add the three `Note` objects we created above by using the `append` method of `Stream`:
```
stream1.append(note1)
stream1.append(note2)
stream1.append(note3)
```
Of course, this would be a pain to type for hundreds of `Notes`, so we could also use the Stream method :meth:`~music21.stream.Stream.repeatAppend` to add a number of independent, unique copies of the same Note. This creates independent copies (using Python's `copy.deepcopy` function) of the supplied object, not references.
```
stream2 = stream.Stream()
n3 = note.Note('D#5') # octave values can be included in creation arguments
stream2.repeatAppend(n3, 4)
stream2.show()
```
But let's worry about that later. Going back to our first stream, we can see that it has three notes using the same `len()` function that we used before:
```
len(stream1)
```
Alternatively, we can use the :meth:`~music21.base.Music21Object.show` method
called as `show('text')` to see what is in the Stream and what its offset
is (here 0.0, since we put it at the end of an empty stream).
```
stream1.show('text')
```
If you’ve setup your environment properly, then calling show with the `musicxml` argument should open up Finale, or Sibelius, or MuseScore or some music notation software and display the notes below.
```
stream1.show()
```
## Accessing Streams
We can also dive deeper into streams. Let's get the `step` of each `Note` using the `for thisNote in ...:`
command. But now we'll use `stream1` instead of `noteList`:
```
for thisNote in stream1:
print(thisNote.step)
```
And we can get the first and the last `Note` in a `Stream` by using the [X] form, just like other Python list-like objects:
```
stream1[0]
stream1[-1].nameWithOctave
```
While full list-like functionality of the Stream isn't there, some additional methods familiar to users of Python lists are also available. The Stream :meth:`~music21.stream.Stream.index` method can be used to get the first-encountered index of a supplied object.
```
note3Index = stream1.index(note3)
note3Index
```
Given an index, an element from the Stream can be removed with the :meth:`~music21.stream.Stream.pop` method.
```
stream1.pop(note3Index)
stream1.show()
```
Since we removed `note3` from `stream1` with the the :meth:`~music21.stream.Stream.pop` method, let's add `note3` back into `stream1` so that we can continue with the examples below using `stream1` as we originally created it.
```
stream1.append(note3)
stream1.show()
```
### Separating out elements by class with `.getElementsByClass()`
We can also gather elements based on the class (object type) of the element, by offset range, or by specific identifiers attached to the element. Gathering elements from a Stream based on the class of the element provides a way to filter the Stream for desired types of objects. The :meth:`~music21.stream.Stream.getElementsByClass` method iterates over a Stream of elements that are instances or subclasses of the provided classes. The example below gathers all :class:`~music21.note.Note` objects and then all :class:`~music21.note.Rest` objects. The easiest way to do this is to use `for` loops with `.getElementsByClass()`:
```
for thisNote in stream1.getElementsByClass(note.Note):
print(thisNote, thisNote.offset)
```
If you want instead of passing the class `note.Note` you could instead pass the string `"Note"`.
```
for thisNote in stream1.getElementsByClass('Note'):
print(thisNote, thisNote.offset)
```
It is also possible to pass in a list of classes or strings of class names to `.getElementsByClass()` which will return anything that matches any of the classes. Notice the `[]` marks in the next call, indicating that we are creating a list to pass to `.getElementsByClass()`:
```
for thisNote in stream1.getElementsByClass(['Note', 'Rest']):
print(thisNote, thisNote.offset)
```
Since there are no `note.Rest` objects, it's the same as above. Oh well...
`music21` has a couple of shortcuts that are equivalent to `.getElementsByClass`. For instance `.notes` is equivalent to `.getElementsByClass(['Note', 'Chord'])` (we'll get to chords soon):
```
for thisNote in stream1.notes:
print(thisNote)
```
And `.notesAndRests` is equivalent to `.getElementsByClass(['Note', 'Chord', 'Rest'])`.
```
for thisNote in stream1.notesAndRests:
print(thisNote)
```
Finally, there's something slightly different. `.pitches` begins with a call to `.notes`, but then returns a list of all the pitches from every `Note` or `Chord` in the Stream:
```
listOut = stream1.pitches
listOut
```
The result of a `.getElementsByClass` are not technically streams, but you can still call `.show()` on it:
```
sOut = stream1.getElementsByClass(note.Note)
sOut.show('text')
```
But if you want to be absolutely sure, put the expression `.stream()` after it:
```
sOut = stream1.getElementsByClass(note.Note).stream() # <-- different
sOut.show('text')
```
### Separating out elements by offset with `.getElementsByOffset()`
The :meth:`~music21.stream.Stream.getElementsByOffset` method returns a Stream of all elements that fall either at a single offset or within a range of two offsets provided as an argument. In both cases a Stream is returned.
```
sOut = stream1.getElementsByOffset(3)
len(sOut)
sOut[0]
```
Like with `.getElementsByClass()` if you want a `Stream` from `.getElementsByOffset()`, add `.stream()` to the end of it.
```
sOut = stream1.getElementsByOffset(2, 3).stream()
sOut.show('text')
```
We will do more with `.getElementsByOffset()` later when we also talk about :meth:`~music21.stream.Stream.getElementAtOrBefore` and :meth:`~music21.stream.Stream.getElementAfterElement`
## More Stream Features
Okay, so far we've seen that `Streams` can do the same things as lists, but
can they do more? Let's call the analyze method on stream to get the
ambitus (that is, the range from the lowest note to the highest note) of
the `Notes` in the `Stream`:
```
stream1.analyze('ambitus')
```
Let's take a second to check this. Our lowest note is `note3` (B-flat in octave 2)
and our highest note is `note2` (F-sharp in octave 4). From B-flat to the F-sharp
above it, is an augmented fifth. An augmented fifth plus an octave is an augmented
twelfth. So we're doing well so far. (We'll get to other things we can analyze in
chapter 18 and we'll see what an :class:`~music21.interval.Interval` object can do
in chapter 15).
As we mentioned earlier, when placed in a Stream, Notes and other elements also have an offset (stored in .offset) that describes their position from the beginning of the stream. These offset values are also given in quarter-lengths (QLs).
Once a Note is in a Stream, we can ask for the `offset` of the `Notes` (or
anything else) in it. The `offset` is the position of a Note relative to the start
of the `Stream` measured in quarter notes. So note1's offset will be 0.0,
since it's at the start of the Stream:
```
note1.offset
```
`note2`'s offset will be 2.0, since `note1` is a half note, worth two quarter notes:
```
note2.offset
```
And `note3`, which follows the quarter note `note2` will be at offset 3.0:
```
note3.offset
```
(If we made `note2` an eighth note, then `note3`'s offset would be the floating point
[decimal] value 2.5. But we didn't.) So now when we're looping we can see the offset of
each note. Let's print the note's offset followed by its name by putting .offset and .name
in the same line, separated by a comma:
```
for thisNote in stream1:
print(thisNote.offset, thisNote.name)
```
(**Digression**: It's probably not too early to learn that a safer form of `.offset` is `.getOffsetBySite(stream1)`:
```
note2.offset
note2.getOffsetBySite(stream1)
```
What's the difference? Remember how I said that `.offset` refers to the number of quarter notes
that the `Note` is from the front of a `Stream`? Well, eventually you may put the same `Note` in
different places in multiple `Streams`, so the `.getOffsetBySite(X)` command is a safer way that
specifies exactly which Stream we are talking about. End of digression...)
As a final note about offsets, the :attr:`~music21.stream.Stream.lowestOffset` property returns the minimum of all offsets for all elements on the Stream.
```
stream1.lowestOffset
```
So, what else can we do with Streams? Like `Note` objects, we can `show()` them in a couple of different
ways. Let's hear these three Notes as a MIDI file:
```
stream1.show('midi')
```
Or let's see them as a score:
```
stream1.show()
```
You might ask why is the piece in common-time (4/4)? This is just the default for new pieces, which is in the
`defaults` module:
```
defaults.meterNumerator
defaults.meterDenominator
```
(Some of these examples use a system that automatically tries to get an appropriate time signature and appropriate clef; in this case, `music21` figured out that that low B-flat would be easier to see in bass clef than treble.)
We'll learn how to switch the :class:`~music21.meter.TimeSignature` soon enough.
If you don't have MIDI or MusicXML configured yet (we'll get to it in a second) and you don't want to have
other programs open up, you can show a `Stream` in text in your editor:
```
stream1.show('text')
```
This display shows the `offset` for each element (that is, each object in the Stream) along with
what class it is, and a little bit more helpful information. The information is the same as
what's called the ``__repr__`` (representation) of the object, which is what you get if you type
its variable name at the prompt:
```
note1
```
By the way, Streams have a ``__repr__`` as well:
```
stream1
```
that number at the end is the hex form of the `.id` of the `Stream`, which is a way of identifying it. Often
the `.id` of a Stream will be the name of the `Part` ("Violin II"), but if it's undefined
then a somewhat random number is used (actually the location of the Stream in your computer's
memory). We can change the `.id` of a Stream:
```
stream1.id = 'some_notes'
stream1
```
We could have also changed the `.id` of any of our `Note` objects, but it doesn't show up in
the `Note`'s ``__repr__``:
```
note1.id = 'my_favorite_C'
note1
```
Now, a `Stream` is a :class:`~music21.base.Music21Object` just like a `Note` is. This is why
it has an `.id` attribute and, more importantly, why you can call `.show()` on it.
What else makes a `Music21Object` what it is?
It has a `.duration` attribute which stores a `Duration` object:
```
stream1.duration
stream1.duration.type
stream1.duration.quarterLength
```
(Notice that the `len()` of a `Stream`, which stands for "length", is not the same as the duration.
the `len()` of a Stream is the number of objects stored in it, so `len(stream1)` is 3).
A related concept to the `.duration` of a Stream is its `.highestTime`, which is the time at which the latest element in the `Stream` ends. Usually this is the last element of the stream's `.offset` plus its `.quarterLength`.
```
stream1.highestTime
```
## Streams within Streams
And, as a `Music21Object`, a `Stream` can be placed inside of another `Stream` object. Let's create
a stream, called biggerStream (for reasons that will become obvious), that holds a `Note` D# at the
beginning
```
biggerStream = stream.Stream()
note2 = note.Note("D#5")
biggerStream.insert(0, note2)
```
Now we use the `.append` functionality to put `stream1` at the end of `biggerStream`:
```
biggerStream.append(stream1)
```
Notice that when we call `.show('text')` on biggerStream, we see not only the presence of `note2`
and `stream1` but also all the contents of `stream1` as well:
```
biggerStream.show('text')
```
Notice though that the offsets, the little numbers inside curly brackets, for the elements of
`stream1` ("some notes") relate only to their positions within `stream1`, not to their position
within `biggerStream`. This is because each `Music21Object` knows its offset only in relation
to its containing `Stream`, not necessarily to the `Stream` containing *that* `Stream`.
Also notice that `note1` knows that it is in `stream1` but doesn't know that it is somewhere inside
`biggerStream`:
```
note1 in stream1
note1 in biggerStream
```
All this might not seem like much of a big deal, until we tell you that in music21, `Scores` are
made up of `Streams` within `Streams` within `Streams`. So if you have an orchestral score, it is
a `Stream`, and the viola part is a `Stream` in that `Stream`, and measure 5 of the viola part is a
`Stream` within that `Stream`, and, if there were a ''divisi'', then each ''diviso'' voice would be
a `Stream` within that `Stream`. Each of these `Streams` has a special name and its own class
(:class:`~music21.stream.Score`, :class:`~music21.stream.Part`, :class:`~music21.stream.Measure`,
and :class:`~music21.stream.Voice`), but they are all types of `Streams`.
So how do we find `note1` inside `biggerStream`? That's what the next two chapters are about.
:ref:`Chapter 5 covers Lists of Lists <usersGuide_05_listsOfLists>`. Those with programming experience who have familiarity with
lists of lists and defining functions might want to skip to :ref:`Chapter 6 Streams of Streams <usersGuide_06_stream2>`.
| github_jupyter |
```
# importing libs
import numpy as np
import tensorflow as tf
import keras
from keras.layers import Input, Dense, GaussianNoise,Lambda
from keras.models import Model
from keras import regularizers
from keras.layers.normalization import BatchNormalization
from keras.optimizers import Adam,SGD
from keras import backend as K
# for reproducing reslut
from numpy.random import seed
seed(1)
from tensorflow import set_random_seed
set_random_seed(3)
# defining parameters
M = 4
k = np.log2(M)
k = int(k)
R = 1
n_channel = 2
print ('M:',M,'k:',k,'n:',n_channel)
#generating data of size N
N = 8000
label = np.random.randint(M,size=N)
# creating one hot encoded vectors
data = []
for i in label:
temp = np.zeros(M)
temp[i] = 1
data.append(temp)
data = np.array(data)
print (data.shape)
temp_check = [17,23,45,67,89,96,72,250,350]
for i in temp_check:
print(label[i],data[i])
def antirectifier(x):
y = x/K.l2_normalize(x,axis=0)
return
def antirectifier_output_shape(input_shape):
return input_shape
input_signal = Input(shape=(M,))
encoded = Dense(M, activation='relu')(input_signal)
encoded1 = Dense(n_channel, activation='linear')(encoded)
encoded2 = Lambda(lambda x: x/tf.norm(x))(encoded1)
EbNo_train = 5.01187 # coverted 7 db of EbNo
encoded3 = GaussianNoise(np.sqrt(1/(2*R*EbNo_train)))(encoded2)
decoded = Dense(M, activation='relu')(encoded3)
decoded1 = Dense(M, activation='softmax')(decoded)
autoencoder = Model(input_signal, decoded1)
adam = Adam(lr=0.01)
autoencoder.compile(optimizer=adam, loss='categorical_crossentropy')
print (autoencoder.summary())
tbCallBack = keras.callbacks.TensorBoard(log_dir='./logs', histogram_freq=0, batch_size=32, write_graph=True, write_grads=True, write_images=False, embeddings_freq=0, embeddings_layer_names=None, embeddings_metadata=None)
autoencoder.fit(data, data,
epochs=100,
batch_size=256)
from keras.models import load_model
#autoencoder.save('2_2_symbol_autoencoder_v_best.model')
#autoencoder_loaded = load_model('4_7_symbol_autoencoder_v_best.model')
encoder = Model(input_signal, encoded2)
encoded_input = Input(shape=(n_channel,))
deco = autoencoder.layers[-2](encoded_input)
deco = autoencoder.layers[-1](deco)
# create the decoder model
decoder = Model(encoded_input, deco)
N = 5000
test_label = np.random.randint(M,size=N)
test_data = []
for i in test_label:
temp = np.zeros(M)
temp[i] = 1
test_data.append(temp)
test_data = np.array(test_data)
temp_test = 6
print (test_data[temp_test][test_label[temp_test]],test_label[temp_test])
print (encoder.predict(np.expand_dims([0,0,0,1],axis=0)))
print (encoder.predict(np.expand_dims([0,0,1,0],axis=0)))
print (encoder.predict(np.expand_dims([0,1,0,0],axis=0)))
print (encoder.predict(np.expand_dims([1,0,0,0],axis=0)))
def frange(x, y, jump):
while x < y:
yield x
x += jump
EbNodB_range = list(frange(-2,10.5,0.5))
ber = [None]*len(EbNodB_range)
for n in range(0,len(EbNodB_range)):
EbNo=10.0**(EbNodB_range[n]/10.0)
noise_std = np.sqrt(1/(2*R*EbNo))
noise_mean = 0
no_errors = 0
nn = N
noise = noise_std * np.random.randn(nn,n_channel)
encoded_signal = encoder.predict(test_data)
final_signal = encoded_signal + noise
pred_final_signal = decoder.predict(final_signal)
pred_output = np.argmax(pred_final_signal,axis=1)
no_errors = (pred_output != test_label)
no_errors = no_errors.astype(int).sum()
ber[n] = no_errors / nn
print ('SNR:',EbNodB_range[n],'BER:',ber[n])
import matplotlib.pyplot as plt
from scipy import interpolate
plt.plot(EbNodB_range, ber, 'bo',label='Autoencoder(2,2)')
#tck = interpolate.splrep(EbNodB_range, ber, s=0)
#xnew = np.arange(-2,8.8, 0.6)
#ynew = interpolate.splev(xnew, tck, der=0)
#plt.plot(xnew,ynew,'y')
#plt.plot(list(EbNodB_range), ber_theory, 'ro-',label='BPSK BER')
plt.yscale('log')
plt.xlabel('SNR Range')
plt.ylabel('Block Error Rate')
plt.grid()
plt.legend(loc='upper right',ncol = 1)
plt.savefig('AutoEncoder_2_2_BER_matplotlib')
plt.show()
```
| github_jupyter |
Thank you to Alex's Lemonade Stand and the Seurat team for their teaching resources, from which much of the material below is adapted.
## scRNA-seq preprocessing and normalization
It's always important to do rigorous preprocessing before analyzing our datasets, and more so for noisy scRNA-seq data. These steps are essential for producing interpretable results in the downstream analysis. In this class, we’ll perform quality control and normalization of scRNA-seq count data with Seurat.
### Setup
Many tools have been developed for the analysis of scRNA-seq data - we'll focus on one of the most widely used and well-maintained packages. Seurat is an R package that can perform QC, clustering, exploration, and visualization of scRNA-seq data, and they are regularly adding more features. It's a good place to start for us!
If you're interested, they have many more useful vigenettes on their website: https://satijalab.org/seurat/get_started.html
We installed Seurat for you, but you need to access it by going to **Project settings --> Project control --> switch to "Experimental" software environment --> Save and Restart**
```
library(Seurat)
packageVersion("Seurat")
library(ggplot2)
```
### Load in Darmanis et al. dataset
We're going to continue using the dataset we started working on in the prelab.
This dataset is generated from human primary glioblastoma tumor cells with the Smart-seq2 protocol (https://www.ncbi.nlm.nih.gov/pubmed/29091775). We're using a subset of this dataset (n=1854 cells). The raw count matrix provided has been quantified using Salmon.
```
# Load in raw counts from Darmanis et al.
sc_data <- read.table("data/unfiltered_darmanis_counts.tsv", header=T, sep="\t", row.names=1)
```
Take a look at the data. How many genes and how many cells are there in the raw dataset? What is the prefix of cell names?
What percentage of the data matrix is 0? Is this surprising?
### Seurat can do our QC and normalization
Recall from the prelab that we looked at two QC metrics of interest: 1. number of reads detected per cell and 2. number of genes detected per cell. Now that we have a better understanding of these QC metrics, we can make our lives easier by using Seurat to visualize these QC stats (and more!) and filter out cells and features that don't pass QC metrics.
If you're interested, you can get a lot more information about the Seurat package and its capabilities here: https://satijalab.org/seurat/get_started.html
Before we start Seurat, we first want to convert Ensembl gene ids to HGNC symbols for easier interpretation. Run the code below to replace Ensembl ids with hgnc symbols in rownames of our data matrix.
```
library(biomaRt)
ensembl <- useMart("ensembl", dataset="hsapiens_gene_ensembl")
bm <- getBM(attributes=c("ensembl_gene_id", "hgnc_symbol"), values=rownames(sc_data), mart=ensembl)
hgnc.symbols <- bm$hgnc_symbol[match(rownames(sc_data), bm$ensembl_gene_id)]
sc_data <- as.matrix(sc_data)
rownames(sc_data) <- hgnc.symbols
#Filter out any rows where the HGNC symbol is blank or NA
sc_data <- subset(sc_data, rownames(sc_data) != "" & !is.na(rownames(sc_data)))
```
###### 1) We'll start by reading in our count data as a Seurat object. This object will hold our count matrix, as well as data for all downstream processing steps or analyses (normalization, scaling, PCA, clustering results, etc.). We can specify extra parameters to take only the features that are present in *min.cells* and the cells that have *min.features*.
```
sc <- CreateSeuratObject(counts=sc_data, project="Darmanis", min.cells=5, min.features=200)
sc
```
The original count matrix is stored in *sc[["RNA"]]@counts*
Print the raw counts for the first 6 genes of the first 6 cells below:
###### 2) Seurat automatically generates the number of reads (nCount_RNA) and number of genes (nFeature_RNA) detected per cell. We can access this data in the *sc@meta.data* slot.
Generate a density plot of nCount_RNA - it should exactly match the density plot we produced in the prelab for total read count!
```
head(sc@meta.data)
qplot(sc@meta.data$nCount_RNA, xlab="Total read counts", geom="density") +
geom_vline(xintercept=25000, color="red") +
theme_classic()
```
###### 3) One final QC metric we're often interested in is the percentage of reads mapping to the mitochondrial genome. A high percentage often indicates low-quality or dying cells. Seurat allows us to search for mitochondrial gene names and calculate the percentage of reads mapping to those genes. We can then stash these stats into our Seurat object's metadata by assigning *sc[[<metadata_feature_name>]]*.
```
sc[["percent.mito"]] <- PercentageFeatureSet(object=sc, pattern="^MT-")
```
What would you have to change in the code above if we were working with mouse data?
###### 4) How is the quality of this experiment? How do you know? We can visualize some QC metrics of interest in a violin plot. We can also check that the number of genes detected correlates with read count.
```
VlnPlot(object=sc, features=c("nCount_RNA", "nFeature_RNA", "percent.mito"), ncol=3, pt.size=0.5)
FeatureScatter(object=sc, feature1="nCount_RNA", feature2="nFeature_RNA")
```
###### 5) Remove low-quality cells (high mitochondrial content), empty droplets (low number of genes), doublets/multiplets (abnormally high number of genes).
Seurat lets us easily apply QC filters based on our custom criteria. In this case, we want cells with >250 genes, but less than 2500, and <10% mitochondrial content.
```
sc <- subset(sc, subset = nFeature_RNA > 250 & nFeature_RNA < 2500 & percent.mito < 10)
sc
```
How many genes and cells remain after QC?
###### 6) Recover cell type annotations, stash in sc@meta.data
Run the following code to add cell type annotations to our Seurat object metadata. This will be useful later when we're visualizing the different cell populations.
```
sc_metadata <- read.table("data/unfiltered_darmanis_metadata.tsv", sep="\t", header=T)
celltypes <- sc_metadata$cell.type.ch1[match(rownames(sc@meta.data), sc_metadata$geo_accession)]
sc@meta.data$celltype <- celltypes
```
Print the metadata for the first 6 cells below:
###### 7) Normalization
Seurat v3 implements a more sophisticated normalization method, SCTransform, but it is still a work in progress and doesn't work well on large datasets yet. Thus, we will use their original method that normalizes by sequencing depth and log transforms the result. We should note that normalization of scRNA-seq data is an active field of research with many distinct approaches and little benchmarking.
```
sc <- NormalizeData(object=sc, normalization.method = "LogNormalize", scale.factor=10000)
```
###### 8) Identify highly variable features
We'll specify that we want the top 2000 highly variable genes with nfeatures. These will be used as input genes for downstream dimensionality reduction (PCA).
```
sc <- FindVariableFeatures(object=sc, selection.method="vst", nfeatures=2000)
# Plot top 10 most variable genes
top10 <- head(x=VariableFeatures(object=sc), 10)
plot1 <- VariableFeaturePlot(object=sc)
plot2 <- LabelPoints(plot=plot1, points=top10, repel=TRUE)
plot2
```
###### 9) Scale data
Scaling is an essential pre-processing step for dimensionality reduction methods like PCA. Here, we not only scale the data, but also regress out unwanted sources of variation such as mitochondrial contamination (we could also do cell cycle stage!).
We scale only the top 2000 highly variable genes for speed. This won't affect our PCA result, as PCA will use only the top variable genes as input anyway. However, if you need to plot heatmaps, you'll want to run ScaleData() on *all* the genes.
```
sc <- ScaleData(object=sc, vars.to.regress="percent.mito")
```
Print the scaled counts for the first 6 genes of the first 6 cells below:
###### 10) Compare normalized data to raw count data
Let's look at how proper data processing impacts our ability to draw interpretable conclusions about the biology. We'll generate PCA plots for both the raw count data and the normalized count data. What do you notice?
```
# Run PCA on raw count data and normalized count data
raw_pca <- prcomp(t(as.matrix(sc[["RNA"]]@counts)))
norm_pca <- prcomp(t((sc[["RNA"]]@scale.data)))
# Retrieve PCA loading scores
raw_pca_scores <- data.frame(raw_pca$x[,1:2], cell_type=sc@meta.data$celltype)
norm_pca_scores <- data.frame(norm_pca$x[,1:2], cell_type=sc@meta.data$celltype)
# Plot PCA with cell labels
ggplot(raw_pca_scores, aes(x=PC1, y=PC2, color=cell_type)) +
geom_point() +
ggtitle("Raw counts PCA") +
theme_classic()
# colorblindr::scale_color_OkabeIto()
# Plot PCA with cell labels
ggplot(norm_pca_scores, aes(x=PC1, y=PC2, color=cell_type)) +
geom_point() +
ggtitle("Normalized counts PCA") +
theme_classic()
# colorblindr::scale_color_OkabeIto()
```
###### 11) Finally, save your processed Seurat object for future downstream analysis.
```
saveRDS(sc, file="sc_Darmanis_normalized.rds")
```
## Homework (10 pts)
###### We're going to be analyzing the Zheng et al. dataset in the next homework (https://www.nature.com/articles/ncomms14049). This dataset consists of 68k FACS-sorted immune cells from peripheral blood. We'll use a small downsampled subset to save time and memory.
###### Using what you've learned above, process the Zheng et al. dataset. You DON'T need to rerun every step, only the ones essential for producing the final processed Seurat object.
###### These steps include:
1. Visualize QC metrics
2. Filter out low-quality cells and lowly expressed genes. Criteria: nFeature_RNA > 500, nFeature_RNA < 2500, percent.mito < 10
3. Normalize
4. Scale, regress out mitochondrial content variable
5. Save the filtered, processed Seurat object
###### Make sure to include all essential code!
```
# We loaded in the data for you
Zheng_data <- read.table("data/Zheng_pbmc_downsample300_C9_filt.txt", sep="\t", header=T, check.names=F)
geneids <- as.character(Zheng_data[,ncol(Zheng_data)])
Zheng_data$gene_symbols <- NULL
Zheng_data <- as.matrix(Zheng_data)
rownames(Zheng_data) <- geneids
# Create Seurat object
sc <- CreateSeuratObject(counts=Zheng_data, project="Zheng2017")
# Store type information in meta data object
celltype <- sapply(strsplit(rownames(sc@meta.data), split="_"), function(x) x[[2]])
sc@meta.data$celltype <- celltype
```
###### Q1 (4 pts). Visualize QC metrics of interest and filter out poor-quality cells.
###### Q2 (4 pts). Normalize the data, identify the top 2000 highly variable features, and scale the data based on those features. Make sure to regress out mitochondrial contamination in the scale step. Finally, use saveRDS() to save the Seurat object as an .rds object named *sc_Zheng_normlized.rds*.
###### Q3 (2 pts). What are the primary sources of technical variability and biases you would be worried about in this experiment? See Zheng et al. for information about the experiment and Ziegenhain et al. for an overview of scRNA-seq technical biases (both papers are in your directory).
| github_jupyter |
## Q. Register at www.kaggle.com / Download csv files / Store folder Dataset root Jupyter
```
import pandas as pd
import matplotlib.pyplot as pl
data = pd.read_csv('covid-19.csv')
df = pd.DataFrame(data)
print(df)
df_pivot =pd.pivot_table(data,index="State/UnionTerritory",values = ["Confirmed","Cured","Deaths"]).sort_values(by="Confirmed",ascending=False).astype(int)
df_pivot
df_pivot.to_excel("Covid19_infected.xlsx")
df_pivot.plot(kind='bar')
pl.show()
```
## Q.Tkinter Pandemic Analyser
```
from tkinter import *
import tkinter
root = Tk()
root.title("Pandemic Analytics Engine")
root.geometry("608x275")
def calculation():
total_ppl = entry_2.get()
int_total_ppl = int(total_ppl)
infected = entry_3.get()
int_infected = int(infected)
deceased = entry_4.get()
int_deceased = int(deceased)
ifr = (int_deceased / int_infected) * 100
cmr = (int_deceased / int_total_ppl)
#root = Tk()
#root.title("Report")
label_1 = Label(root, font=("helvetica", 22, "bold"), text="Result")
label_1.grid(row=7, columnspan=1)
label_2 = Label(root, font=("courier", 12, "bold"), text="IFR : ")
label_2.grid(row=9, column=1)
label_3 = Label(root, text=ifr)
label_3.grid(row=9, column=2)
label_4 = Label(root, font=("courier", 12, "bold"), text="CMR : ")
label_4.grid(row=10, column=1)
label_5 = Label(root, text=cmr)
label_5.grid(row=10, column=2)
options = ["Maharashtra","Tamil Nadu","Delhi","Gujarat","Uttar Pradesh","West Bengal","Rajasthan","Telengana","Karnataka","Andhra Pradesh","Haryana","Madhya Pradesh","Bihar","Assam","Jammu and Kashmir","Odisha","Punjab","Kerala","Chhattisgarh","Uttarakhand","Jharkhand","Goa","Tripura","Manipur","Himachal Pradesh","Ladakh","Puducherry","Nagaland","Chandigarh","Dadra and Nagar Haveli","Arunachal Pradesh","Mizoram","Andaman and Nicobar","Sikkim","Meghalaya","Lakshadweep"]
select = StringVar()
select.set(options[0])
label_1 = Label(root,font=("helvetica", 22, "bold"), text="Pandemic Analytics Engine")
label_1.grid(row=1,columnspan = 1)
label_2 = Label(root, text="Enter State ")
label_2.grid(row=2)
drop = OptionMenu(root,select,*options)
drop.grid(row=2,column=2)
label_3 = Label(root, text="Total number of ppl")
label_3.grid(row=3)
entry_2 = Entry(root)
entry_2.grid(row=3, column=2)
label_4 = Label(root, text="Infected")
label_4.grid(row=4)
entry_3 = Entry(root)
entry_3.grid(row=4, column=2)
label_5 = Label(root, text="Deceased")
label_5.grid(row=5)
entry_4 = Entry(root)
entry_4.grid(row=5, column=2)
button1 = Button(root, text="Calculate IFR & CMR ",command=calculation)
button1.grid(column=2)
root.mainloop()
```
| github_jupyter |
<img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
<img src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/3f/HubSpot_Logo.svg/220px-HubSpot_Logo.svg.png" alt="drawing" width="200" align='left'/>
# Hubspot - Update contact
<a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Hubspot/Hubspot_update_contact.ipynb" target="_parent"><img src="https://img.shields.io/badge/-Open%20in%20Naas-success?labelColor=000000&logo=data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iMTAyNHB4IiBoZWlnaHQ9IjEwMjRweCIgdmlld0JveD0iMCAwIDEwMjQgMTAyNCIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayIgdmVyc2lvbj0iMS4xIj4KIDwhLS0gR2VuZXJhdGVkIGJ5IFBpeGVsbWF0b3IgUHJvIDIuMC41IC0tPgogPGRlZnM+CiAgPHRleHQgaWQ9InN0cmluZyIgdHJhbnNmb3JtPSJtYXRyaXgoMS4wIDAuMCAwLjAgMS4wIDIyOC4wIDU0LjUpIiBmb250LWZhbWlseT0iQ29tZm9ydGFhLVJlZ3VsYXIsIENvbWZvcnRhYSIgZm9udC1zaXplPSI4MDAiIHRleHQtZGVjb3JhdGlvbj0ibm9uZSIgZmlsbD0iI2ZmZmZmZiIgeD0iMS4xOTk5OTk5OTk5OTk5ODg2IiB5PSI3MDUuMCI+bjwvdGV4dD4KIDwvZGVmcz4KIDx1c2UgaWQ9Im4iIHhsaW5rOmhyZWY9IiNzdHJpbmciLz4KPC9zdmc+Cg=="/></a>
#hubspot #crm #sales
## Input
### Import library
```
from naas_drivers import hubspot
```
### Enter your Hubspot api key
```
auth_token = "YOUR_HUBSPOT_API_KEY"
```
### Connect to Hubspot
```
hs = hubspot.connect(auth_token)
```
## Model
### Enter contact parameters
```
contact_id = "280751"
email = "test@cashstory.com"
firstname = "Jean test"
lastname ='CASHSTOrY'
phone = "+336.00.00.00.00"
jobtitle = "Consultant"
website = "www.cashstory.com"
company = 'CASHSTORY'
hubspot_owner_id = None
```
## Output
### Using patch method
```
update_contact = {"properties":
{
"email": email,
"firstname": firstname,
"lastname": lastname,
"phone": phone,
"jobtitle": jobtitle,
"website": website,
"company": company,
"url": "test3",
"hubspot_owner_id": hubspot_owner_id,
}
}
hs.contacts.patch(contact_id, update_contact)
```
### Using update method
```
hs.contacts.update(contact_id,
email,
firstname,
lastname,
phone,
jobtitle,
website,
company,
hubspot_owner_id)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NielsRogge/Transformers-Tutorials/blob/master/.ipynb_checkpoints/Custom_Named_Entity_Recognition_with_BERT.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## **Fine-tuning BERT for named-entity recognition**
In this notebook, we are going to use **BertForTokenClassification** which is included in the [Transformers library](https://github.com/huggingface/transformers) by HuggingFace. This model has BERT as its base architecture, with a token classification head on top, allowing it to make predictions at the token level, rather than the sequence level. Named entity recognition is typically treated as a token classification problem, so that's what we are going to use it for.
This tutorial uses the idea of **transfer learning**, i.e. first pretraining a large neural network in an unsupervised way, and then fine-tuning that neural network on a task of interest. In this case, BERT is a neural network pretrained on 2 tasks: masked language modeling and next sentence prediction. Now, we are going to fine-tune this network on a NER dataset. Fine-tuning is supervised learning, so this means we will need a labeled dataset.
If you want to know more about BERT, I suggest the following resources:
* the original [paper](https://arxiv.org/abs/1810.04805)
* Jay Allamar's [blog post](http://jalammar.github.io/illustrated-bert/) as well as his [tutorial](http://jalammar.github.io/a-visual-guide-to-using-bert-for-the-first-time/)
* Chris Mccormick's [Youtube channel](https://www.youtube.com/channel/UCoRX98PLOsaN8PtekB9kWrw)
* Abbishek Kumar Mishra's [Youtube channel](https://www.youtube.com/user/abhisheksvnit)
The following notebook largely follows the same structure as the tutorials by Abhishek Kumar Mishra. For his tutorials on the Transformers library, see his [Github repository](https://github.com/abhimishra91/transformers-tutorials).
NOTE: this notebook assumes basic knowledge about deep learning, BERT, and native PyTorch. If you want to learn more Python, deep learning and PyTorch, I highly recommend cs231n by Stanford University and the FastAI course by Jeremy Howard et al. Both are freely available on the web.
Now, let's move on to the real stuff!
#### **Importing Python Libraries and preparing the environment**
This notebook assumes that you have the following libraries installed:
* pandas
* numpy
* sklearn
* pytorch
* transformers
* seqeval
As we are running this in Google Colab, the only libraries we need to additionally install are transformers and seqeval (GPU version):
```
#!pip install transformers seqeval[gpu]
import pandas as pd
import numpy as np
from sklearn.metrics import accuracy_score
import torch
from torch.utils.data import Dataset, DataLoader
from transformers import BertTokenizer, BertConfig, BertForTokenClassification
```
As deep learning can be accellerated a lot using a GPU instead of a CPU, make sure you can run this notebook in a GPU runtime (which Google Colab provides for free! - check "Runtime" - "Change runtime type" - and set the hardware accelerator to "GPU").
We can set the default device to GPU using the following code (if it prints "cuda", it means the GPU has been recognized):
```
from torch import cuda
device = 'cuda' if cuda.is_available() else 'cpu'
print(device)
```
#### **Downloading and preprocessing the data**
Named entity recognition (NER) uses a specific annotation scheme, which is defined (at least for European languages) at the *word* level. An annotation scheme that is widely used is called **[IOB-tagging](https://en.wikipedia.org/wiki/Inside%E2%80%93outside%E2%80%93beginning_(tagging))**, which stands for Inside-Outside-Beginning. Each tag indicates whether the corresponding word is *inside*, *outside* or at the *beginning* of a specific named entity. The reason this is used is because named entities usually comprise more than 1 word.
Let's have a look at an example. If you have a sentence like "Barack Obama was born in Hawaï", then the corresponding tags would be [B-PERS, I-PERS, O, O, O, B-GEO]. B-PERS means that the word "Barack" is the beginning of a person, I-PERS means that the word "Obama" is inside a person, "O" means that the word "was" is outside a named entity, and so on. So one typically has as many tags as there are words in a sentence.
So if you want to train a deep learning model for NER, it requires that you have your data in this IOB format (or similar formats such as [BILOU](https://stackoverflow.com/questions/17116446/what-do-the-bilou-tags-mean-in-named-entity-recognition)). There exist many annotation tools which let you create these kind of annotations automatically (such as Spacy's [Prodigy](https://prodi.gy/), [Tagtog](https://docs.tagtog.net/) or [Doccano](https://github.com/doccano/doccano)). You can also use Spacy's [biluo_tags_from_offsets](https://spacy.io/api/goldparse#biluo_tags_from_offsets) function to convert annotations at the character level to IOB format.
Here, we will use a NER dataset from [Kaggle](https://www.kaggle.com/namanj27/ner-dataset) that is already in IOB format. One has to go to this web page, download the dataset, unzip it, and upload the csv file to this notebook. Let's print out the first few rows of this csv file:
```
data = pd.read_csv("ner_datasetreference.csv", encoding='unicode_escape')
data.head()
```
Let's check how many sentences and words (and corresponding tags) there are in this dataset:
```
data.count()
```
As we can see, there are approximately 48,000 sentences in the dataset, comprising more than 1 million words and tags (quite huge!). This corresponds to approximately 20 words per sentence.
Let's have a look at the different NER tags, and their frequency:
```
print("Number of tags: {}".format(len(data.Tag.unique())))
frequencies = data.Tag.value_counts()
frequencies
```
There are 8 category tags, each with a "beginning" and "inside" variant, and the "outside" tag. It is not really clear what these tags mean - "geo" probably stands for geographical entity, "gpe" for geopolitical entity, and so on. They do not seem to correspond with what the publisher says on Kaggle. Some tags seem to be underrepresented. Let's print them by frequency (highest to lowest):
```
tags = {}
for tag, count in zip(frequencies.index, frequencies):
if tag != "O":
if tag[2:5] not in tags.keys():
tags[tag[2:5]] = count
else:
tags[tag[2:5]] += count
continue
print(sorted(tags.items(), key=lambda x: x[1], reverse=True))
```
Let's remove "art", "eve" and "nat" named entities, as performance on them will probably be not comparable to the other named entities.
```
entities_to_remove = ["B-art", "I-art", "B-eve", "I-eve", "B-nat", "I-nat"]
data = data[~data.Tag.isin(entities_to_remove)]
data.head()
```
Now, we have to ask ourself the question: what is a training example in the case of NER, which is provided in a single forward pass? A training example is typically a **sentence**, with corresponding IOB tags. Let's group the words and corresponding tags by sentence:
```
# pandas has a very handy "forward fill" function to fill missing values based on the last upper non-nan value
data = data.fillna(method='ffill')
data.head()
# let's create a new column called "sentence" which groups the words by sentence
data['sentence'] = data[['Sentence #','Word','Tag']].groupby(['Sentence #'])['Word'].transform(lambda x: ' '.join(x))
# let's also create a new column called "word_labels" which groups the tags by sentence
data['word_labels'] = data[['Sentence #','Word','Tag']].groupby(['Sentence #'])['Tag'].transform(lambda x: ','.join(x))
data.head()
```
Let's have a look at the different NER tags.
We create 2 dictionaries: one that maps individual tags to indices, and one that maps indices to their individual tags. This is necessary in order to create the labels (as computers work with numbers = indices, rather than words = tags) - see further in this notebook.
```
labels_to_ids = {k: v for v, k in enumerate(data.Tag.unique())}
ids_to_labels = {v: k for v, k in enumerate(data.Tag.unique())}
labels_to_ids
```
As we can see, there are now only 10 different tags.
Let's only keep the "sentence" and "word_labels" columns, and drop duplicates:
```
data = data[["sentence", "word_labels"]].drop_duplicates().reset_index(drop=True)
data.head()
len(data)
```
Let's verify that a random sentence and its corresponding tags are correct:
```
data.iloc[41].sentence
data.iloc[41].word_labels
```
#### **Preparing the dataset and dataloader**
Now that our data is preprocessed, we can turn it into PyTorch tensors such that we can provide it to the model. Let's start by defining some key variables that will be used later on in the training/evaluation process:
```
MAX_LEN = 128
TRAIN_BATCH_SIZE = 4
VALID_BATCH_SIZE = 2
EPOCHS = 1
LEARNING_RATE = 1e-05
MAX_GRAD_NORM = 10
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
```
A tricky part of NER with BERT is that BERT relies on **wordpiece tokenization**, rather than word tokenization. This means that we should also define the labels at the wordpiece-level, rather than the word-level!
For example, if you have word like "Washington" which is labeled as "b-gpe", but it gets tokenized to "Wash", "##ing", "##ton", then we will have to propagate the word’s original label to all of its wordpieces: "b-gpe", "b-gpe", "b-gpe". The model should be able to produce the correct labels for each individual wordpiece. The function below (taken from [here](https://github.com/chambliss/Multilingual_NER/blob/master/python/utils/main_utils.py#L118)) implements this.
```
def tokenize_and_preserve_labels(sentence, text_labels, tokenizer):
"""
Word piece tokenization makes it difficult to match word labels
back up with individual word pieces. This function tokenizes each
word one at a time so that it is easier to preserve the correct
label for each subword. It is, of course, a bit slower in processing
time, but it will help our model achieve higher accuracy.
"""
tokenized_sentence = []
labels = []
sentence = sentence.strip()
for word, label in zip(sentence.split(), text_labels.split(",")):
# Tokenize the word and count # of subwords the word is broken into
tokenized_word = tokenizer.tokenize(word)
n_subwords = len(tokenized_word)
# Add the tokenized word to the final tokenized word list
tokenized_sentence.extend(tokenized_word)
# Add the same label to the new list of labels `n_subwords` times
labels.extend([label] * n_subwords)
return tokenized_sentence, labels
```
Note that this is a **design decision**. You could also decide to only label the first wordpiece of each word and let the model only learn this (this is what was done in the original BERT paper, see Github discussion [here](https://github.com/huggingface/transformers/issues/64#issuecomment-443703063)). Another design decision could be to give the first wordpiece of each word the original word label, and then use the label “X” for all subsequent subwords of that word.
All of them lead to good performance.
Next, we define a regular PyTorch [dataset class](https://pytorch.org/docs/stable/data.html) (which transforms examples of a dataframe to PyTorch tensors). Here, each sentence gets tokenized, the special tokens that BERT expects are added, the tokens are padded or truncated based on the max length of the model, the attention mask is created and the labels are created based on the dictionary which we defined above.
For more information about BERT's inputs, see [here](https://huggingface.co/transformers/glossary.html).
```
class dataset(Dataset):
def __init__(self, dataframe, tokenizer, max_len):
self.len = len(dataframe)
self.data = dataframe
self.tokenizer = tokenizer
self.max_len = max_len
def __getitem__(self, index):
# step 1: tokenize (and adapt corresponding labels)
sentence = self.data.sentence[index]
word_labels = self.data.word_labels[index]
tokenized_sentence, labels = tokenize_and_preserve_labels(sentence, word_labels, self.tokenizer)
# step 2: add special tokens (and corresponding labels)
tokenized_sentence = ["[CLS]"] + tokenized_sentence + ["[SEP]"] # add special tokens
labels.insert(0, "O") # add outside label for [CLS] token
labels.insert(-1, "O") # add outside label for [SEP] token
# step 3: truncating/padding
maxlen = self.max_len
if (len(tokenized_sentence) > maxlen):
# truncate
tokenized_sentence = tokenized_sentence[:maxlen]
labels = labels[:maxlen]
else:
# pad
tokenized_sentence = tokenized_sentence + ['[PAD]'for _ in range(maxlen - len(tokenized_sentence))]
labels = labels + ["O" for _ in range(maxlen - len(labels))]
# step 4: obtain the attention mask
attn_mask = [1 if tok != '[PAD]' else 0 for tok in tokenized_sentence]
# step 5: convert tokens to input ids
ids = self.tokenizer.convert_tokens_to_ids(tokenized_sentence)
label_ids = [labels_to_ids[label] for label in labels]
# the following line is deprecated
#label_ids = [label if label != 0 else -100 for label in label_ids]
return {
'ids': torch.tensor(ids, dtype=torch.long),
'mask': torch.tensor(attn_mask, dtype=torch.long),
#'token_type_ids': torch.tensor(token_ids, dtype=torch.long),
'targets': torch.tensor(label_ids, dtype=torch.long)
}
def __len__(self):
return self.len
```
Now, based on the class we defined above, we can create 2 datasets, one for training and one for testing. Let's use a 80/20 split:
```
train_size = 0.8
train_dataset = data.sample(frac=train_size,random_state=200)
test_dataset = data.drop(train_dataset.index).reset_index(drop=True)
train_dataset = train_dataset.reset_index(drop=True)
print("FULL Dataset: {}".format(data.shape))
print("TRAIN Dataset: {}".format(train_dataset.shape))
print("TEST Dataset: {}".format(test_dataset.shape))
training_set = dataset(train_dataset, tokenizer, MAX_LEN)
testing_set = dataset(test_dataset, tokenizer, MAX_LEN)
```
Let's have a look at the first training example:
```
training_set[0]
```
Let's verify that the input ids and corresponding targets are correct:
```
for token, label in zip(tokenizer.convert_ids_to_tokens(training_set[0]["ids"]), training_set[0]["targets"]):
print('{0:10} {1}'.format(token, label))
```
Now, let's define the corresponding PyTorch dataloaders:
```
train_params = {'batch_size': TRAIN_BATCH_SIZE,
'shuffle': True,
'num_workers': 0
}
test_params = {'batch_size': VALID_BATCH_SIZE,
'shuffle': True,
'num_workers': 0
}
training_loader = DataLoader(training_set, **train_params)
testing_loader = DataLoader(testing_set, **test_params)
```
#### **Defining the model**
Here we define the model, BertForTokenClassification, and load it with the pretrained weights of "bert-base-uncased". The only thing we need to additionally specify is the number of labels (as this will determine the architecture of the classification head).
Note that only the base layers are initialized with the pretrained weights. The token classification head of top has just randomly initialized weights, which we will train, together with the pretrained weights, using our labelled dataset. This is also printed as a warning when you run the code cell below.
Then, we move the model to the GPU.
```
model = BertForTokenClassification.from_pretrained('bert-base-uncased', num_labels=len(labels_to_ids))
model.to(device)
```
#### **Training the model**
Before training the model, let's perform a sanity check, which I learned thanks to Andrej Karpathy's wonderful [cs231n course](http://cs231n.stanford.edu/) at Stanford (see also his [blog post about debugging neural networks](http://karpathy.github.io/2019/04/25/recipe/)). The initial loss of your model should be close to -ln(1/number of classes) = -ln(1/17) = 2.83.
Why? Because we are using cross entropy loss. The cross entropy loss is defined as -ln(probability score of the model for the correct class). In the beginning, the weights are random, so the probability distribution for all of the classes for a given token will be uniform, meaning that the probability for the correct class will be near 1/17. The loss for a given token will thus be -ln(1/17). As PyTorch's [CrossEntropyLoss](https://pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html) (which is used by `BertForTokenClassification`) uses *mean reduction* by default, it will compute the mean loss for each of the tokens in the sequence (in other words, for all of the 512 tokens). The mean of 512 times -log(1/17) is, you guessed it, -log(1/17).
Let's verify this:
```
ids = training_set[0]["ids"].unsqueeze(0)
mask = training_set[0]["mask"].unsqueeze(0)
targets = training_set[0]["targets"].unsqueeze(0)
ids = ids.to(device)
mask = mask.to(device)
targets = targets.to(device)
outputs = model(input_ids=ids, attention_mask=mask, labels=targets)
initial_loss = outputs[0]
initial_loss
```
This looks good. Let's also verify that the logits of the neural network have a shape of (batch_size, sequence_length, num_labels):
```
tr_logits = outputs[1]
tr_logits.shape
```
Next, we define the optimizer. Here, we are just going to use Adam with a default learning rate. One can also decide to use more advanced ones such as AdamW (Adam with weight decay fix), which is [included](https://huggingface.co/transformers/main_classes/optimizer_schedules.html) in the Transformers repository, and a learning rate scheduler, but we are not going to do that here.
```
optimizer = torch.optim.Adam(params=model.parameters(), lr=LEARNING_RATE)
```
Now let's define a regular PyTorch training function. It is partly based on [a really good repository about multilingual NER](https://github.com/chambliss/Multilingual_NER/blob/master/python/utils/main_utils.py#L344).
```
# Defining the training function on the 80% of the dataset for tuning the bert model
def train(epoch):
tr_loss, tr_accuracy = 0, 0
nb_tr_examples, nb_tr_steps = 0, 0
tr_preds, tr_labels = [], []
# put model in training mode
model.train()
for idx, batch in enumerate(training_loader):
ids = batch['ids'].to(device, dtype = torch.long)
mask = batch['mask'].to(device, dtype = torch.long)
targets = batch['targets'].to(device, dtype = torch.long)
loss, tr_logits = model(input_ids=ids, attention_mask=mask, labels=targets)
tr_loss += loss.item()
nb_tr_steps += 1
nb_tr_examples += targets.size(0)
if idx % 100==0:
loss_step = tr_loss/nb_tr_steps
print(f"Training loss per 100 training steps: {loss_step}")
# compute training accuracy
flattened_targets = targets.view(-1) # shape (batch_size * seq_len,)
active_logits = tr_logits.view(-1, model.num_labels) # shape (batch_size * seq_len, num_labels)
flattened_predictions = torch.argmax(active_logits, axis=1) # shape (batch_size * seq_len,)
# now, use mask to determine where we should compare predictions with targets (includes [CLS] and [SEP] token predictions)
active_accuracy = mask.view(-1) == 1 # active accuracy is also of shape (batch_size * seq_len,)
targets = torch.masked_select(flattened_targets, active_accuracy)
predictions = torch.masked_select(flattened_predictions, active_accuracy)
tr_preds.extend(predictions)
tr_labels.extend(targets)
tmp_tr_accuracy = accuracy_score(targets.cpu().numpy(), predictions.cpu().numpy())
tr_accuracy += tmp_tr_accuracy
# gradient clipping
torch.nn.utils.clip_grad_norm_(
parameters=model.parameters(), max_norm=MAX_GRAD_NORM
)
# backward pass
optimizer.zero_grad()
loss.backward()
optimizer.step()
epoch_loss = tr_loss / nb_tr_steps
tr_accuracy = tr_accuracy / nb_tr_steps
print(f"Training loss epoch: {epoch_loss}")
print(f"Training accuracy epoch: {tr_accuracy}")
```
And let's train the model!
```
for epoch in range(EPOCHS):
print(f"Training epoch: {epoch + 1}")
train(epoch)
```
#### **Evaluating the model**
Now that we've trained our model, we can evaluate its performance on the held-out test set (which is 20% of the data). Note that here, no gradient updates are performed, the model just outputs its logits.
```
def valid(model, testing_loader):
# put model in evaluation mode
model.eval()
eval_loss, eval_accuracy = 0, 0
nb_eval_examples, nb_eval_steps = 0, 0
eval_preds, eval_labels = [], []
with torch.no_grad():
for idx, batch in enumerate(testing_loader):
ids = batch['ids'].to(device, dtype = torch.long)
mask = batch['mask'].to(device, dtype = torch.long)
targets = batch['targets'].to(device, dtype = torch.long)
loss, eval_logits = model(input_ids=ids, attention_mask=mask, labels=targets)
eval_loss += loss.item()
nb_eval_steps += 1
nb_eval_examples += targets.size(0)
if idx % 100==0:
loss_step = eval_loss/nb_eval_steps
print(f"Validation loss per 100 evaluation steps: {loss_step}")
# compute evaluation accuracy
flattened_targets = targets.view(-1) # shape (batch_size * seq_len,)
active_logits = eval_logits.view(-1, model.num_labels) # shape (batch_size * seq_len, num_labels)
flattened_predictions = torch.argmax(active_logits, axis=1) # shape (batch_size * seq_len,)
# now, use mask to determine where we should compare predictions with targets (includes [CLS] and [SEP] token predictions)
active_accuracy = mask.view(-1) == 1 # active accuracy is also of shape (batch_size * seq_len,)
targets = torch.masked_select(flattened_targets, active_accuracy)
predictions = torch.masked_select(flattened_predictions, active_accuracy)
eval_labels.extend(targets)
eval_preds.extend(predictions)
tmp_eval_accuracy = accuracy_score(targets.cpu().numpy(), predictions.cpu().numpy())
eval_accuracy += tmp_eval_accuracy
#print(eval_labels)
#print(eval_preds)
labels = [ids_to_labels[id.item()] for id in eval_labels]
predictions = [ids_to_labels[id.item()] for id in eval_preds]
#print(labels)
#print(predictions)
eval_loss = eval_loss / nb_eval_steps
eval_accuracy = eval_accuracy / nb_eval_steps
print(f"Validation Loss: {eval_loss}")
print(f"Validation Accuracy: {eval_accuracy}")
return labels, predictions
```
As we can see below, performance is quite good! Accuracy on the test test is > 93%.
```
labels, predictions = valid(model, testing_loader)
```
However, the accuracy metric is misleading, as a lot of labels are "outside" (O), even after omitting predictions on the [PAD] tokens. What is important is looking at the precision, recall and f1-score of the individual tags. For this, we use the seqeval Python library:
```
from seqeval.metrics import classification_report
print(classification_report(labels, predictions))
```
#### **Inference**
The fun part is when we can quickly test the model on new, unseen sentences.
Here, we use the prediction of the **first word piece of every word**. Note that the function we used to train our model (`tokenze_and_preserve_labels`) propagated the label to all subsequent word pieces (so you could for example also perform a majority vote on the predicted labels of all word pieces of a word).
*In other words, the code below does not take into account when predictions of different word pieces that belong to the same word do not match.*
```
sentence = "India has a capital called Mumbai. On wednesday, the president will give a presentation"
inputs = tokenizer(sentence, padding='max_length', truncation=True, max_length=MAX_LEN, return_tensors="pt")
# move to gpu
ids = inputs["input_ids"].to(device)
mask = inputs["attention_mask"].to(device)
# forward pass
outputs = model(ids, mask)
logits = outputs[0]
active_logits = logits.view(-1, model.num_labels) # shape (batch_size * seq_len, num_labels)
flattened_predictions = torch.argmax(active_logits, axis=1) # shape (batch_size*seq_len,) - predictions at the token level
tokens = tokenizer.convert_ids_to_tokens(ids.squeeze().tolist())
token_predictions = [ids_to_labels[i] for i in flattened_predictions.cpu().numpy()]
wp_preds = list(zip(tokens, token_predictions)) # list of tuples. Each tuple = (wordpiece, prediction)
word_level_predictions = []
for pair in wp_preds:
if (pair[0].startswith(" ##")) or (pair[0] in ['[CLS]', '[SEP]', '[PAD]']):
# skip prediction
continue
else:
word_level_predictions.append(pair[1])
# we join tokens, if they are not special ones
str_rep = " ".join([t[0] for t in wp_preds if t[0] not in ['[CLS]', '[SEP]', '[PAD]']]).replace(" ##", "")
print(str_rep)
print(word_level_predictions)
```
#### **Saving the model for future use**
Finally, let's save the vocabulary, model weights and the model's configuration file to a directory, so that they can be re-loaded using the `from_pretrained()` class method.
```
import os
directory = "./model"
if not os.path.exists(directory):
os.makedirs(directory)
# save vocabulary of the tokenizer
tokenizer.save_vocabulary(directory)
# save the model weights and its configuration file
model.save_pretrained(directory)
print('All files saved')
print('This tutorial is completed')
```
## Legacy
```
def prepare_sentence(sentence, tokenizer, maxlen):
# step 1: tokenize the sentence
tokenized_sentence = tokenizer.tokenize(sentence)
# step 2: add special tokens
tokenized_sentence = ["[CLS]"] + tokenized_sentence + ["[SEP]"]
# step 3: truncating/padding
if (len(tokenized_sentence) > maxlen):
# truncate
tokenized_sentence = tokenized_sentence[:maxlen]
else:
# pad
tokenized_sentence = tokenized_sentence + ['[PAD]'for _ in range(maxlen - len(tokenized_sentence))]
# step 4: obtain the attention mask
attn_mask = [1 if tok != '[PAD]' else 0 for tok in tokenized_sentence]
# step 5: convert tokens to input ids
ids = tokenizer.convert_tokens_to_ids(tokenized_sentence)
return {
'ids': torch.tensor(ids, dtype=torch.long),
'mask': torch.tensor(attn_mask, dtype=torch.long),
#'token_type_ids': torch.tensor(token_ids, dtype=torch.long),
}
```
| github_jupyter |
# The JAX emulator: Greybody prototype
In this notebook, I will prototype my idea for emulating radiative transfer codes with a Deepnet in order for it to be used inside xidplus. As `numpyro` uses JAX, the Deepnet wil ideally be trained with a JAX network. As a proof of concept, I will use a greybody rather than a radiative transfer code.
```
import fitIR
import fitIR.models as models
import fitIR.analyse as analyse
from astropy.cosmology import WMAP9 as cosmo
import jax
import numpy as onp
import pylab as plt
import astropy.units as u
import scipy.integrate as integrate
%matplotlib inline
import jax.numpy as np
from jax import grad, jit, vmap, value_and_grad
from jax import random
from jax import vmap # for auto-vectorizing functions
from functools import partial # for use with vmap
from jax import jit # for compiling functions for speedup
from jax.experimental import stax # neural network library
from jax.experimental.stax import Conv, Dense, MaxPool, Relu, Flatten, LogSoftmax, LeakyRelu # neural network layers
from jax.experimental import optimizers
from jax.tree_util import tree_multimap # Element-wise manipulation of collections of numpy arrays
import matplotlib.pyplot as plt # visualization
# Generate key which is used to generate random numbers
key = random.PRNGKey(1)
```
The first step is to create a training and validation dataset. To do this I will randomly sample from parameter space (rather than a grid). I will create a function to do the sampling. I will also define function to do the transform and inverse_transform from standardised values to physical values.
```
def standarise_uniform(lims):
param_sd=(lims[1]-lims[0])/np.sqrt(12.0)
param_mean=0.5*(lims[1]+lims[0])
return param_sd,param_mean
def generate_samples(size=100,lims=np.array([[6,16],[0,7],[20,80]])):
"""Sample from uniform space"""
#get parameter values from uniform distribution
LIR=onp.random.uniform(low=lims[0,0],high=lims[0,1],size=size)
#sample in log10 space for redshift
redshift=onp.random.uniform(low=lims[1,0],high=lims[1,1],size=size)
#sample in log10 space
temperature=onp.random.uniform(low=lims[2,0],high=lims[2,1],size=size)
#get standard deviation and mean for uniform dist
LIR_sd,LIR_mean=standarise_uniform(lims[0,:])
red_sd, red_mean=standarise_uniform(lims[1,:])
temp_sd,temp_mean=standarise_uniform(lims[2,:])
return onp.vstack((LIR,redshift,temperature)).T,onp.vstack(((LIR-LIR_mean)/LIR_sd,(redshift-red_mean)/red_sd,(temperature-temp_mean)/temp_sd)).T
def transform_parameters(param,lims=np.array([[6,16],[0,7],[20,80]])):
"""transform from physical values to standardised values"""
LIR_sd,LIR_mean=standarise_uniform(lims[0,:])
red_sd, red_mean=standarise_uniform(lims[1,:])
temp_sd,temp_mean=standarise_uniform(lims[2,:])
LIR_norm=(param[0]-LIR_mean)/LIR_sd
red_norm=(param[1]-red_mean)/red_sd
temp_norm=(param[2]-temp_mean)/temp_sd
return np.vstack((LIR_norm,red_norm,temp_norm)).T
def inverse_transform_parameters(param,lims=np.array([[6,16],[0,7],[20,80]])):
""" Transform from standardised parameters to physical values
function works with posterior samples"""
LIR_sd,LIR_mean=standarise_uniform(lims[0,:])
red_sd, red_mean=standarise_uniform(lims[1,:])
temp_sd,temp_mean=standarise_uniform(lims[2,:])
LIR=param[...,0]*LIR_sd+LIR_mean
red=param[...,1]*red_sd+red_mean
temp=param[...,2]*temp_sd+temp_mean
return np.stack((LIR.T,red.T,temp.T)).T
```
I need to convolve the grebody with the relevant filters. I will use the code I already wrote in xidplus for the original SED work
```
import xidplus
from xidplus import filters
filter_=filters.FilterFile(file=xidplus.__path__[0]+'/../test_files/filters.res')
SPIRE_250=filter_.filters[215]
SPIRE_350=filter_.filters[216]
SPIRE_500=filter_.filters[217]
MIPS_24=filter_.filters[201]
PACS_100=filter_.filters[250]
PACS_160=filter_.filters[251]
bands=[SPIRE_250,SPIRE_350,SPIRE_500]#,PACS_100,PACS_160]
eff_lam=[250.0,350.0,500.0]#, 100.0,160.0]
from scipy.interpolate import interp1d
def get_fluxes(samples):
measured=onp.empty((samples.shape[0],len(bands)))
val = onp.linspace(onp.log10(3E8/8E-6),onp.log10(3E8/1E-3),1000)
val = 10**val
for i,s in enumerate(samples):
z=s[1]
prior = {}
prior['z'] = s[1]
prior['log10LIR'] = s[0]
prior['T'] = s[2]
prior['emissivity'] = 1.5
source = models.greybody(prior)
nu,lnu = source.generate_greybody(val,z)
wave = 3E8/nu*1E6
sed=interp1d(wave,lnu)
dist = cosmo.luminosity_distance(z).to(u.cm).value
for b in range(0,len(bands)):
measured[i,b]=(1.0+z)*filters.fnu_filt(sed(bands[b].wavelength/1E4),
3E8/(bands[b].wavelength/1E10),
bands[b].transmission,
3E8/(eff_lam[b]*1E-6),
sed(eff_lam[b]))/(4*onp.pi*dist**2)
return measured/10**(-26)
```
## DeepNet building
I will build a multi input, multi output deepnet model as my emulator, with parameters as input and the observed flux as outputs. I will train on log10 flux to make the model easier to train, and have already standarised the input parameters. I wilkl be using `stax` which can be thought of as the `Keras` equivalent for `JAX`. This [blog](https://blog.evjang.com/2019/02/maml-jax.html) was useful starting point.
```
import torch
from torch.utils.data import Dataset, DataLoader
## class for sed using the torch dataset class
class sed_data(Dataset):
def __init__(self,params,fluxes):
self.X=params
self.y=fluxes
def __len__(self):
return len(self.X)
def __getitem__(self,idx):
return self.X[idx],self.y[idx]
```
I will use batches to help train the network
```
batch_size=10
## generate random SED samples
samp_train,samp_stand_train=generate_samples(2000)
## Use Steve's code and xidplus filters to get fluxes
measured_train=get_fluxes(samp_train)
## use data in SED dataclass
ds = sed_data(samp_stand_train,measured_train)
## use torch DataLoader
train_loader = DataLoader(ds, batch_size=batch_size,)
## do same but for test set
samp_test,samp_stand_test=generate_samples(500)
measured_test=get_fluxes(samp_test)
ds = sed_data(samp_stand_test,measured_test)
test_loader = DataLoader(ds, batch_size=batch_size)
# Use stax to set up network initialization and evaluation functions
net_init, net_apply = stax.serial(
Dense(128), LeakyRelu,
Dense(128), LeakyRelu,
Dense(len(bands))
)
in_shape = (-1, 3,)
out_shape, net_params = net_init(key,in_shape)
def loss(params, inputs, targets):
# Computes average loss for the batch
predictions = net_apply(params, inputs)
return np.mean((targets - predictions)**2)
def batch_loss(p,x_b,y_b):
loss_b=vmap(partial(loss,p))(x_b,y_b)
return np.mean(loss_b)
def sample_batch(outer_batch_size,inner_batch_size):
def get_batch():
xs, ys = [], []
for i in range(0,outer_batch_size):
samp_train,samp_stand_train=generate_samples(inner_batch_size)
## Use Steve's code and xidplus filters to get fluxes
measured_train=get_fluxes(samp_train)
xs.append(samp_stand_train)
ys.append(np.log(measured_train))
return np.stack(xs), np.stack(ys)
x1, y1 = get_batch()
return x1, y1
opt_init, opt_update, get_params= optimizers.adam(step_size=1e-3)
out_shape, net_params = net_init(key,in_shape)
opt_state = opt_init(net_params)
@jit
def step(i, opt_state, x1, y1):
p = get_params(opt_state)
g = grad(batch_loss)(p, x1, y1)
loss_tmp=batch_loss(p,x1,y1)
return opt_update(i, g, opt_state),loss_tmp
np_batched_loss_1 = []
valid_loss=[]
K=40
for i in range(4000):
# sample random batchs for training
x1_b, y1_b = sample_batch(10, K)
# sample random batches for validation
x2_b,y2_b = sample_batch(1,K)
opt_state, l = step(i, opt_state, x1_b, y1_b)
p = get_params(opt_state)
valid_loss.append(batch_loss(p,x2_b,y2_b))
np_batched_loss_1.append(l)
if i % 100 == 0:
print(i)
net_params = get_params(opt_state)
opt_init, opt_update, get_params= optimizers.adam(step_size=1e-4)
for i in range(5000):
# sample random batchs for training
x1_b, y1_b = sample_batch(10, K)
# sample random batches for validation
x2_b,y2_b = sample_batch(1,K)
opt_state, l = step(i, opt_state, x1_b, y1_b)
p = get_params(opt_state)
valid_loss.append(batch_loss(p,x2_b,y2_b))
np_batched_loss_1.append(l)
if i % 100 == 0:
print(i)
net_params = get_params(opt_state)
plt.figure(figsize=(10,5))
plt.semilogy(np_batched_loss_1,label='Training loss')
plt.semilogy(valid_loss,label='Validation loss')
plt.xlabel('Iteration')
plt.ylabel('Loss (MSE)')
plt.legend()
```
## Investigate performance of each band of emulator
To visulise performance of the trainied emulator, I will show the difference between real and emulated for each band.
```
x,y=sample_batch(100,100)
predictions = net_apply(net_params,x)
res=(predictions-y)/(y)
fig,axes=plt.subplots(1,3,figsize=(50,10))
for i in range(0,3):
axes[i].hist(res[:,:,i].flatten()*100.0,np.arange(-20,20,0.5))
axes[i].set_title(bands[i].name)
axes[i].set_xlabel(r'$\frac{f_{pred} - f_{True}}{f_{True}} \ \%$ error')
plt.subplots_adjust(wspace=0.5)
```
## Save network
Having trained and validated network, I need to save the network and relevant functions
```
import cloudpickle
with open('GB_emulator_20210324_notlog10z_T.pkl', 'wb') as f:
cloudpickle.dump({'net_init':net_init,'net_apply': net_apply,'params':net_params,'transform_parameters':transform_parameters,'inverse_transform_parameters':inverse_transform_parameters}, f)
net_init, net_apply
transform_parameters
from xidplus.numpyro_fit.misc import load_emulator
obj=load_emulator('GB_emulator_20210323.pkl')
```
| github_jupyter |
<a href="/assets/tutorial06_example.ipynb" class="link-button">Download</a>
<a href="https://colab.research.google.com/github/technion046195/technion046195/blob/master/content/tutorial06/example.ipynb" target="_blank">
<img src="../assets/colab-badge.svg" style="display:inline"/>
</a>
<center><h1>
תרגול 6 - דוגמא מעשית
</h1></center>
## Setup
```
## Importing packages
import os # A build in package for interacting with the OS. For example to create a folder.
import numpy as np # Numerical package (mainly multi-dimensional arrays and linear algebra)
import pandas as pd # A package for working with data frames
import matplotlib.pyplot as plt # A plotting package
import imageio # A package to read and write image (is used here to save gif images)
import tabulate # A package from pretty printing tables
from graphviz import Digraph # A package for plothing graphs (of nodes and edges)
## Setup matplotlib to output figures into the notebook
## - To make the figures interactive (zoomable, tooltip, etc.) use ""%matplotlib notebook" instead
%matplotlib inline
## Setting some nice matplotlib defaults
plt.rcParams['figure.figsize'] = (4.5, 4.5) # Set default plot's sizes
plt.rcParams['figure.dpi'] = 120 # Set default plot's dpi (increase fonts' size)
plt.rcParams['axes.grid'] = True # Show grid by default in figures
## Auxiliary function for prining equations, pandas tables and images in cells output
from IPython.core.display import display, HTML, Latex, Markdown
## Create output folder
if not os.path.isdir('./output'):
os.mkdir('./output')
```
### Data Inspection
```
data_file = 'https://technion046195.netlify.app/datasets/nyc_taxi_rides.csv'
## Loading the data
dataset = pd.read_csv(data_file)
## Print the number of rows in the data set
number_of_rows = len(dataset)
display(Markdown(f'Number of rows in the dataset: $N={number_of_rows}$'))
## Show the first 10 rows
dataset.head(10)
x = dataset['duration'].values
```
## Train-test split
```
n_samples = len(x)
## Generate a random generator with a fixed seed
rand_gen = np.random.RandomState(0)
## Generating a vector of indices
indices = np.arange(n_samples)
## Shuffle the indices
rand_gen.shuffle(indices)
## Split the indices into 80% train / 20% test
n_samples_train = int(n_samples * 0.8)
train_indices = indices[:n_samples_train]
test_indices = indices[n_samples_train:]
x_train = x[train_indices]
x_test = x[test_indices]
x_grid = np.arange(-10, 60 + 0.1, 0.1)
```
## Attempt 1 : Normal Distribution
Calculating models parameters:
$$
\mu=\displaystyle{\frac{1}{N}\sum_i x_i} \\
\sigma=\sqrt{\displaystyle{\frac{1}{N}\sum_i\left(x_i-\mu\right)^2}} \\
$$
```
# Normal distribution parameters
mu = np.sum(x) / len(x)
sigma = np.sqrt(np.sum((x - mu) ** 2) / len(x))
display(Latex('$\\mu = {:.01f}\\ \\text{{min}}$'.format(mu)))
display(Latex('$\\sigma = {:.01f}\\ \\text{{min}}$'.format(sigma)))
```
From here on we will use [np.mean](http://lagrange.univ-lyon1.fr/docs/numpy/1.11.0/reference/generated/numpy.mean.html) and [np.std](http://lagrange.univ-lyon1.fr/docs/numpy/1.11.0/reference/generated/numpy.std.html) functions to calculate the mean and standard deviation.
In addition [scipy.stats](https://docs.scipy.org/doc/scipy/reference/stats.html) has a wide range of distribution models. Each model comes with a set of methods for calculating the CDF, PDF, performing MLE fit, generate samples and more.
```
## Import the normal distribution model from SciPy
from scipy.stats import norm
## Define the normal distribution object
norm_dist = norm(mu, sigma)
## Calculate the normal distribution PDF over the grid
norm_pdf = norm_dist.pdf(x_grid)
## Prepare the figure
fig, ax = plt.subplots()
ax.hist(dataset['duration'].values, bins=300 ,density=True, label='Histogram')
ax.plot(x_grid, norm_pdf, label='Normal')
ax.set_title('Distribution of Durations')
ax.set_ylabel('PDF')
ax.set_xlabel('Duration [min]')
ax.legend();
fig.savefig('./output/nyc_duration_normal.png')
```
### Attempt 2 : Rayleigh Distribution
Calculating models parameters:
$$
\Leftrightarrow \sigma = \sqrt{\frac{1}{2N}\sum_i x^2}
$$
```
## Import the normal distribution model from SciPy
from scipy.stats import rayleigh
## Find the model's parameters using SciPy
_, sigma = rayleigh.fit(x, floc=0) ## equivalent to running: sigma = np.sqrt(np.sum(x ** 2) / len(x) / 2)
display(Latex('$\\sigma = {:.01f}$'.format(sigma)))
## Define the Rayleigh distribution object
rayleigh_dist = rayleigh(0, sigma)
## Calculate the Rayleigh distribution PDF over the grid
rayleigh_pdf = rayleigh_dist.pdf(x_grid)
## Prepare the figure
fig, ax = plt.subplots()
ax.hist(dataset['duration'].values, bins=300 ,density=True, label='Histogram')
ax.plot(x_grid, norm_pdf, label='Normal')
ax.plot(x_grid, rayleigh_pdf, label='Rayleigh')
ax.set_title('Distribution of Durations')
ax.set_ylabel('PDF')
ax.set_xlabel('Duration [min]')
ax.legend();
fig.savefig('./output/nyc_duration_rayleigh.png')
```
### Attempt 3 : Generalized Gamma Distribution
Numerical solution
```
## Import the normal distribution model from SciPy
from scipy.stats import gengamma
## Find the model's parameters using SciPy
a, c, _, sigma = gengamma.fit(x, floc=0)
display(Latex('$a = {:.01f}$'.format(a)))
display(Latex('$c = {:.01f}$'.format(c)))
display(Latex('$\\sigma = {:.01f}$'.format(sigma)))
## Define the generalized gamma distribution object
gengamma_dist = gengamma(a, c, 0, sigma)
## Calculate the generalized gamma distribution PDF over the grid
gengamma_pdf = gengamma_dist.pdf(x_grid)
## Prepare the figure
fig, ax = plt.subplots()
ax.hist(dataset['duration'].values, bins=300 ,density=True, label='Histogram')
ax.plot(x_grid, norm_pdf, label='Normal')
ax.plot(x_grid, rayleigh_pdf, label='Rayleigh')
ax.plot(x_grid, gengamma_pdf, label='Generalized Gamma')
ax.set_title('Distribution of Durations')
ax.set_ylabel('PDF')
ax.set_xlabel('Duration [min]')
ax.legend();
fig.savefig('./output/nyc_duration_generalized_gamma.png')
```
| github_jupyter |
## Part 1: Predicting IMDB movie review polarity
Sentiment analysis is a hot topic in data science right now due to the immense amount of user-generated text data being created every day online. Businesses can now look at what is being said about them on review sites to get an idea of how well they are liked, how much they are disliked, and what they can do to improve. While most of this data is unlabeled, some sites also ask users to provide a numerical or star rating. This allows us to build a classifier for positive/negative reviews using the star rating as a label, which could then hypothetically applied to unlabeled text.
IMDB collects information about movies and lets users write their own reviews, as well as provide a 1-10 numerical rating. The data for this assignment can be found in hw4_IMDB.csv. It consists of 12,500 positive and 12,500 negative reviews collected from IMDB. The ratings have been binarized by labeling anything with score between 7 and 10 as “P” and anything between 1 and 4 with “N” (there are no “neutral” reviews in the data). We will build and evaluate a system that classifies these movie reviews as positive or negative.
```
import pandas as pd
url = "hw4_IMDB.csv"
df = pd.read_csv(url)
```
### 1.1 Preprocessing our data
Before we build a classifier, we normally do some text pre-processing: text is an unstructured form of data---at least more unstructured than the feature vectors we used in the previous exercises. By pre-processing we can "clean our textual data"
Do the following preprocessing on the textual data:
- Convert all upper case characters to lower case
- Remove all non alphanumeric characters
- Remove stopwords
Why is each of these steps beneficial? Explain.
### 1.2 Building a Naive Bayes classifier
We will now build our predictive model
- First, turn the text into a features vector by using 1-grams, also called bag-of-words. This can be done with the CountVectorizer function in SKLearn. What is the shape of the resulting table? What does the number of rows and what do the number of columns mean?
- Measure the performance of a Naive Bayes classifier using 3-fold CV, and report the accuract of the classifier across each fold, as well as the average accuracy. Is accuracy a good measure of predictive performance here? If yes, why? If no, what measure would you instead use?
### 1.3 Interpreting the results of the classifier
Get the cross-validation predictions. Pick some instances that were incorrectly classified and read them through. Are there any words in these misclassified reviews that may have misled the classifier? Explain with at least three examples for each type of error you can find.
### 1.4 Improving the performance of your classifier
This is an open ended exercise. How far can you push the performance of your classifier? You can try some of the following:
- Use 2-grams (ordered pairs of 2 words), or higher degree n-grams
- Preproces your data differently. For example, you may choose to not remove some punctuation marks, not lowercasing the words, use a stemmer, or not remove stopwords
- Use other predictive algorithms, as you see fit.
## Part 2: Predicting Yelp star rating
Yelp is a very popular website that collects reviews of restaurants and other local businesses, along with star ratings from 1-5. Instead of classifying these reviews as positive or negative, we’ll now build a classifier to predict the star rating of a Yelp review from its text. Star rating prediction can be viewed either as a multiclass classification problem or a regression problem. For now we’ll treat it as multiclass classification. This is our first problem that is not simple binary classification, and will come with its own set of issues, which we will delve into below.
### 2.1 Interpreting the new accuracies
Read the data from "hw4_Yelp.csv" and then perform the preprocessing steps, Naive Bayes classifier fitting, and evaluation as in Part 1.
Why are the accuracies lower?
```
import pandas as pd
url = "hw4_Yelp.csv"
df = pd.read_csv(url)
```
### 2.2 Confusion and cost matrices
Use the confusion_matrix function from sklearn.metrics to get the confusion matrix for your classifier.
We have provided two cost matrices below. Apply them to your confusion matrix and report the total cost of using this classifier under each cost scheme. Which one of these (a or b) makes more sense in the context of this multiclass classification problem, and why?
**Table 1**
| a | b | c | d | e | |
| --- | --- | --- | --- | --- | --- |
| 0 | 2 | 2 | 2 | 2 | a=1 ||
| 2 | 0 | 2 | 2 | 2 | b=2 |
| 2 | 2 | 0 | 2 | 2 | c=3 |
| 2 | 2 | 2 | 0 | 2 | d=4 |
| 2 | 2 | 2 | 2 | 0 | e=5 |
**Table 2**
| a | b | c | d | e | |
| --- | --- | --- | --- | --- | --- |
| 0 | 1 | 2 | 3 | 4 | a=1 ||
| 1 | 0 | 1 | 2 | 3 | b=2 |
| 2 | 1 | 0 | 1 | 2 | c=3 |
| 3 | 2 | 1 | 0 | 1 | d=4 |
| 4 | 3 | 2 | 1 | 0 | e=5 |
### 2.3 (bonus question) Using regression
Could we instead use a regression predictive model for this problem? Fit a simple linear regression to the training data, using the same pipeline as before. What are some advantages and disadvantages in using a regression instead of multi-class classification
| github_jupyter |
```
import pandas as pd
import numpy as np
import sklearn
import subprocess
import warnings
pd.set_option('display.max_columns', None)
source_path = "/Users/sandrapietrowska/Documents/Trainings/luigi/data_source/"
# coding: ISO-8859-1
```
# Import data
```
raw_dataset = pd.read_csv(source_path + "Speed_Dating_Data.csv",encoding = "ISO-8859-1")
```
# Data exploration
### Shape, types, distribution, modalities and potential missing values
```
raw_dataset.head(3)
raw_dataset_copy = raw_dataset
check1 = raw_dataset_copy[raw_dataset_copy["iid"] == 1]
check1_sel = check1[["iid", "pid", "match","gender","date","go_out","sports","tvsports","exercise","dining",
"museums","art","hiking","gaming","clubbing","reading","tv","theater","movies",
"concerts","music","shopping","yoga"]]
check1_sel.drop_duplicates().head(20)
#merged_datasets = raw_dataset.merge(raw_dataset_copy, left_on="pid", right_on="iid")
#merged_datasets[["iid_x","gender_x","pid_y","gender_y"]].head(5)
#same_gender = merged_datasets[merged_datasets["gender_x"] == merged_datasets["gender_y"]]
#same_gender.head()
columns_by_types = raw_dataset.columns.to_series().groupby(raw_dataset.dtypes).groups
raw_dataset.dtypes.value_counts()
raw_dataset.isnull().sum().head(3)
summary = raw_dataset.describe() #.transpose()
print (summary.head())
#raw_dataset.groupby("gender").agg({"iid": pd.Series.nunique})
raw_dataset.groupby('gender').iid.nunique()
raw_dataset.groupby('career').iid.nunique().sort_values(ascending=False).head(5)
raw_dataset.groupby(["gender","match"]).iid.nunique()
```
# Data processing
```
local_path = "/Users/sandrapietrowska/Documents/Trainings/luigi/data_source/"
local_filename = "Speed_Dating_Data.csv"
my_variables_selection = ["iid", "pid", "match","gender","date","go_out","sports","tvsports","exercise","dining",
"museums","art","hiking","gaming","clubbing","reading","tv","theater","movies",
"concerts","music","shopping","yoga"]
class RawSetProcessing(object):
"""
This class aims to load and clean the dataset.
"""
def __init__(self,source_path,filename,features):
self.source_path = source_path
self.filename = filename
self.features = features
# Load data
def load_data(self):
raw_dataset_df = pd.read_csv(self.source_path + self.filename,encoding = "ISO-8859-1")
return raw_dataset_df
# Select variables to process and include in the model
def subset_features(self, df):
sel_vars_df = df[self.features]
return sel_vars_df
@staticmethod
# Remove ids with missing values
def remove_ids_with_missing_values(df):
sel_vars_filled_df = df.dropna()
return sel_vars_filled_df
@staticmethod
def drop_duplicated_values(df):
df = df.drop_duplicates()
return df
# Combine processing stages
def combiner_pipeline(self):
raw_dataset = self.load_data()
subset_df = self.subset_features(raw_dataset)
subset_no_dup_df = self.drop_duplicated_values(subset_df)
subset_filled_df = self.remove_ids_with_missing_values(subset_no_dup_df)
return subset_filled_df
raw_set = RawSetProcessing(local_path, local_filename, my_variables_selection)
dataset_df = raw_set.combiner_pipeline()
dataset_df.head(3)
# Number of unique participants
dataset_df.iid.nunique()
dataset_df.shape
```
# Feature engineering
```
suffix_me = "_me"
suffix_partner = "_partner"
def get_partner_features(df, suffix_1, suffix_2, ignore_vars=True):
#print df[df["iid"] == 1]
df_partner = df.copy()
if ignore_vars is True:
df_partner = df_partner.drop(['pid','match'], 1).drop_duplicates()
else:
df_partner = df_partner.copy()
#print df_partner.shape
merged_datasets = df.merge(df_partner, how = "inner",left_on="pid", right_on="iid",suffixes=(suffix_1,suffix_2))
#print merged_datasets[merged_datasets["iid_me"] == 1]
return merged_datasets
feat_eng_df = get_partner_features(dataset_df,suffix_me,suffix_partner)
feat_eng_df.head(3)
```
# Modelling
This model aims to answer the questions what is the profile of the persons regarding interests that got the most matches.
Variables:
* gender
* date (In general, how frequently do you go on dates?)
* go out (How often do you go out (not necessarily on dates)?
* sports: Playing sports/ athletics
* tvsports: Watching sports
* excersice: Body building/exercising
* dining: Dining out
* museums: Museums/galleries
* art: Art
* hiking: Hiking/camping
* gaming: Gaming
* clubbing: Dancing/clubbing
* reading: Reading
* tv: Watching TV
* theater: Theater
* movies: Movies
* concerts: Going to concerts
* music: Music
* shopping: Shopping
* yoga: Yoga/meditation
```
import sklearn
print (sklearn.__version__)
from sklearn import tree
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
import subprocess
```
## Variables selection
```
#features = list(["gender","age_o","race_o","goal","samerace","imprace","imprelig","date","go_out","career_c"])
features = list(["gender","date","go_out","sports","tvsports","exercise","dining","museums","art",
"hiking","gaming","clubbing","reading","tv","theater","movies","concerts","music",
"shopping","yoga"])
label = "match"
#add suffix to each element of list
def process_features_names(features, suffix_1, suffix_2):
features_me = [feat + suffix_1 for feat in features]
features_partner = [feat + suffix_2 for feat in features]
features_all = features_me + features_partner
print (features_all)
return features_all
features_model = process_features_names(features, suffix_me, suffix_partner)
feat_eng_df.head(5)
explanatory = feat_eng_df[features_model]
explained = feat_eng_df[label]
```
#### Decision Tree
```
clf = tree.DecisionTreeClassifier(min_samples_split=20,min_samples_leaf=10,max_depth=4)
clf = clf.fit(explanatory, explained)
# Download http://www.graphviz.org/
with open("data.dot", 'w') as f:
f = tree.export_graphviz(clf, out_file=f, feature_names= features_model, class_names="match")
subprocess.call(['dot', '-Tpdf', 'data.dot', '-o' 'data.pdf'])
```
## Tuning Parameters
```
# Split the dataset in two equal parts
X_train, X_test, y_train, y_test = train_test_split(explanatory, explained, test_size=0.3, random_state=0)
parameters = [
{'criterion': ['gini','entropy'], 'max_depth': [4,6,10,12,14],
'min_samples_split': [10,20,30], 'min_samples_leaf': [10,15,20]
}
]
scores = ['precision', 'recall']
dtc = tree.DecisionTreeClassifier()
clf = GridSearchCV(dtc, parameters,n_jobs=3, cv=5, refit=True)
warnings.filterwarnings("ignore")
for score in scores:
print("# Tuning hyper-parameters for %s" % score)
print("")
clf = GridSearchCV(dtc, parameters, cv=5,
scoring='%s_macro' % score)
clf.fit(X_train, y_train)
print("Best parameters set found on development set:")
print("")
print(clf.best_params_)
print("")
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print("")
best_param_dtc = tree.DecisionTreeClassifier(criterion="entropy",min_samples_split=10,min_samples_leaf=10,max_depth=14)
best_param_dtc = best_param_dtc.fit(explanatory, explained)
best_param_dtc.feature_importances_
raw_dataset.rename(columns={"age_o":"age_of_partner","race_o":"race_of_partner"},inplace=True)
```
# Test
```
import unittest
from pandas.util.testing import assert_frame_equal
```
There is a weird thing, with self.XX the code does not work. I tried self.assertEqual
```
class FeatureEngineeringTest(unittest.TestCase):
def test_get_partner_features(self):
"""
:return:
"""
# Given
raw_data_a = {
'iid': ['1', '2', '3', '4', '5','6'],
'first_name': ['Sue', 'Maria', 'Sandra', 'Bill', 'Brian','Bruce'],
'sport':['foot','run','volley','basket','swim','tv'],
'pid': ['4', '5', '6', '1', '2','3'],}
df_a = pd.DataFrame(raw_data_a, columns = ['iid', 'first_name', 'sport','pid'])
expected_output_values = pd.DataFrame({
'iid_me': ['1', '2', '3', '4', '5','6'],
'first_name_me': ['Sue', 'Maria', 'Sandra', 'Bill', 'Brian','Bruce'],
'sport_me': ['foot','run','volley','basket','swim','tv'],
'pid_me': ['4', '5', '6', '1', '2','3'],
'iid_partner': ['4', '5', '6', '1', '2','3'],
'first_name_partner': ['Bill', 'Brian','Bruce','Sue', 'Maria', 'Sandra'],
'sport_partner': ['basket','swim','tv','foot','run','volley'],
'pid_partner':['1', '2', '3', '4', '5','6']
}, columns = ['iid_me','first_name_me','sport_me','pid_me',
'iid_partner','first_name_partner','sport_partner','pid_partner'])
# When
output_values = get_partner_features(df_a, "_me","_partner",ignore_vars=False)
# Then
assert_frame_equal(output_values, expected_output_values)
suite = unittest.TestLoader().loadTestsFromTestCase(FeatureEngineeringTest)
unittest.TextTestRunner(verbosity=2).run(suite)
```
| github_jupyter |
<img style="float: right;" src="figures/research_commons_logo.png">
# WORKSHOP: INTRODUCTION TO R FOR STATISTICAL ANALYSIS
* Arthur Marques (GAA)
* Minjeong Park
* Mohammad Zubair
# Learning Goals
<img src="figures/learning_objectives.png">
* Understand the basic commands of R and how to use RStudio environment
* Learn how to load, preprocess, and explore datasets
* Learn the basic commands for descriptive statistics
* Learn basic commands for inference statistics
***
## Workshop materials
### Access the UBC research commons R workshop
https://guides.library.ubc.ca/library_research_commons/rworkshop
<img src="figures/path_to_files.png">
***
### Download the files for this workshop
<img src="figures/save_to_desktop.png">
***
## Overview of quantitative research
<img src="figures/overview.png">
***
<img src="figures/comparison.png">
***
## R environment
### Open RStudio
<img src="figures/R_environment.png">
***
### Create a new R script
#### File > New file > R Script
<img src="figures/R_Studio.png">
***
### Change your working directory
Set up a folder/path where data are located as a working directory
<img src="figures/working_directory.png">
***
# Basic Commands
<img src="figures/learning_objectives.png">
### R packages
R package is a library of prewritten code designed for a particular task or a collection of tasks
<img src="figures/R_packages.png">
***
### Installing a new package (2 options)
#### Under Tools -> Packages tab -> Search for “psych” and “dplyr”
<img src="figures/install_packages.png">
***
#### Using code: install.packages( ) and library( ) functions
```
install.packages(c("ggplot2", "dplyr", "readr", "psych"))
library("ggplot2")
library("dplyr")
library("readr")
library("psych")
```
```
library("ggplot2")
library("dplyr")
library("readr")
```
#### Tips
**On the editor**
<kbd>CTRL</kbd>+<kbd>enter</kbd> runs the command in the current line (<kbd>cmd</kbd>+<kbd>enter</kbd> on MacOS)
<kbd>CTRL</kbd>+<kbd>w</kbd> closes current tab (<kbd>cmd</kbd>+<kbd>w</kbd> on MacOS)
After typing the first few lines of a command, <kbd>tab</kbd> auto-completes the command
**On the console**
<kbd>↑</kbd> shows the last command (useful to rerun the command of fix errors)
***
# Data Manipulation
<img src="figures/learning_objectives.png">
#### Our data
<img src="figures/global_warming.jpeg">
***
### Import data from excel
<img src="figures/data_import_from_excel.png">
***
### Preview data and give it a nice name
<img src="figures/data_import_preview.png">
***
### You can also load your data with the read_csv command
```
mydata <- read_csv("GlobalLandTemperaturesByCountry.csv")
```
***
## Basic data commands
### names( ): check variable names
```
names(mydata)
```
***
### View first n lines of the table
```
head(mydata, n = 10)
```
***
### table( ): check variable values and frequency
```
table(mydata$Country)
```
```
head(table(mydata$Country), n=10)
```
***
## Basic data management commands
### is.factor( ): check if the variable is defined as categorical
```
is.factor(mydata$Country)
```
### as.factor( ): changes variable to categorical format
```
mydata$Country <- as.factor(mydata$Country)
is.factor(mydata$Country)
```
### numeric( ): check if the variable is defined as numerical
```
is.numeric(mydata$AverageTemperature)
```
***
## Data management with "dplyr" package
### Removing empty cells/rows
```
head(mydata, n = 10)
mydata <- na.omit(mydata)
head(mydata, n = 10)
```
***
### select ( ): selects columns based on columns names
```
head(select(mydata, dt, Country), n=5)
```
***
### filter ( ): selects cases based on conditions
```
head(filter(mydata, Country=="Canada"))
```
### filter may accept more than one condition
```
head(filter(mydata, Country=="Canada" | Country == "China"))
head(filter(mydata, Country=="Canada" & AverageTemperature > 12))
```
***
### mutate( ): adds new variables
#### Adding column representing the year
```
mydata <- mutate(mydata, year = as.numeric(format(dt, "%Y")))
head(mydata)
is.numeric(mydata$year)
```
**Whait? What?**
Command breakdown:
The commands below should be run on RStudio:
```
format(mydata$dt, "%Y")
as.numeric(format(mydata$dt, "%Y"))
mutate(mydata, year = as.numeric(format(dt, "%Y")))
mydata <- mutate(mydata, year = as.numeric(format(dt, "%Y")))
```
***
#### Adding column representing the industrial era
```
mydata <- mutate(mydata, era=if_else(
year <= 1969, "gas & oil", "electronic",
))
head(mydata, n=5)
tail(mydata, n=5)
```
***
# Descriptive Statistics
<img src="figures/learning_objectives.png">
### Descriptive stat with basic summary function
```
summary(mydata)
```
***
### Descriptive statistics with visualization
#### Create histogram for “year”
```
hist(mydata$year)
```
***
#### Create histogram for “Countries”
```
barplot(table(mydata$era))
```
***
### More visualization functions with “ggplot2”
Try to run this command:
```
ggplot(data=mydata, aes(x=year, y=AverageTemperature)) + geom_line()
```
<img src="figures/staring.jpg">
### Can we do better?
What if we can create smaller data with the year and the average temperature of that year?
```
grouped_data <- mydata %>%
group_by(year) %>%
summarise(avg_temp = mean(AverageTemperature))
head(grouped_data)
ggplot(data=grouped_data, aes(x=year, y=avg_temp)) +
geom_line()
```
# Exercise
## If we are on track, try to:
### 1. load carbon dioxide data
### 2. remove NA
### 3. Change column "CarbonDioxide" to numeric
### 4. Change column "year" to numeric
### view the data using the head( ) and tail( ) commands
<br><br><br><br>
## Don't spoil the fun. The stick figure is watching you
<br><br><br><br>
<img src="figures/watch.png" width="30%">
### Answer:
```
carbon <- read_csv("CarbonDioxideEmission.csv")
carbon <- na.omit(carbon)
carbon$CarbonDioxide <- as.numeric(carbon$CarbonDioxide)
carbon$year <- as.numeric(carbon$year)
head(carbon)
tail(carbon)
```
# Inference Statistics
<img src="figures/learning_objectives.png">
## Some housekeeping and background
### Copy & run the code below
```
carbon <- group_by(carbon, year) %>%
summarise(AverageCarbonEmission = mean(CarbonDioxide))
newdata <- group_by(mydata, year, era) %>%
summarise(AverageTemperature = mean(AverageTemperature))
carbon <- merge(newdata, carbon[, c("year", "AverageCarbonEmission")], by="year")
head(carbon)
```
### You may want to review this code at home, but for now let's just consider that it merges two datasets
A detailed explanation for later reference explained line-by-line:
1. the `carbon` data will be updated with the result of the right-hand operation. The right-hand operation groups the dataset by year
<br>
2. after grouping, we apply the `summarise` to create a column named `carbon` the value of the column is the `mean` of the `CarbonDioxide` emission
<br>
3. we do a similar grouping operation creating a `newdata` data.
<br>
4. this time, we are interested in the mean of the `AverageTemperature`
<br>
5. we now have two variables, one with the `year` and `AverageCarbonEmission` and another with year and `AverageTemperature`. Let's `merge` these two tables in a final `carbon` table. Our merging criteria is the `year` and we will copy two columns from the initial carbon table: `c("year", "AverageCarbonEmission")`
<br>
### Hypothesis testing
From [Wikipedia](https://en.wikipedia.org/wiki/Statistical_significance):
### To determine whether a result is statistically significant, a researcher calculates a p-value, which is the probability of observing an effect of the same magnitude or more extreme given that the null hypothesis is true.
### The null hypothesis is rejected if the p-value is less than a predetermined level, α.
### α is called the significance level, and is the probability of rejecting the null hypothesis given that it is true (a type I error).
### α is usually set at or below 5%.
<img src="figures/normal_curve.png">
## Our null hypotheses
### There is no difference in the Average temperature in the `gas & oil` era and the `electronic` era
```
head(mydata)
```
#### Independent T-test compares means between two groups
It is often used to see whether there is a group difference in continuous data **between two groups**
*Model assumptions*
(1) Independence, (2) Normality, (3) Equal variance
```
t.test(AverageTemperature ~ era, data=carbon, var.eq=TRUE)
```
Interpreting the results:
* `t` value guides our analysis. Read more at this [link](https://blog.minitab.com/blog/adventures-in-statistics-2/understanding-t-tests-t-values-and-t-distributions)
* `df = 54` degrees of freedom
* `p-value < 0.0004415` pretty low, so that means that we can reject the null hypothesis
<br>
* Which one seems higher?
* mean in group `gas & oil` = `18.74364`
* mean in group `eletronics` = `19.13249`
### Is there any association between the `AverageTemperature` and the `AverageCarbonEmission` ?
#### Pearson’s correlation is used to examine associations between variables (represented by continuous data) by looking at the direction and strength of the associations
```
cor.test(carbon$AverageTemperature, carbon$AverageCarbonEmission, method="pearson")
```
Interpreting the results:
* `p-value < 2.2e-16` pretty low, so that means that there is statistically significant correlation between `temperature` and `carbon emission`
<br>
* How strong is the correlation `cor` = `0.8970832`
Interpretation varies by research field so results should be interpreted with caution
`cor` varies from `-1` to `1` positive values indicate that an increase in the `x` variable increases the `y` variable. In this case, a value closer to `1` means a strong positive correlation
***
### Conclusions from our analysis
<img src="figures/this_is_fine.png">
***
# Research Commons: An interdisciplinary research-driven learning environment
### * Literature review
### * Systematic Review Search Methodology
### * Citation Management
### * Thesis Formatting
### * Nvivo Software Support
### * SPSS Software Support
<img style="float: right;" src="figures/research_commons_logo.png">
### * R Group
### * Multi-Disciplinary Graduate Student Writing Group
***
<img style="position: relative;" src="figures/background.png">
<img style="position: absolute; top: 50px; left: 200px; background: rgba(255,255,255, 0.5);" src="figures/thanks.png">
<center> <h1><a src="http://bit.ly/RCfeedbackwinter2018">Feedback</a></h1> </center>
| github_jupyter |
```
import numpy as np
def sigmoid(x):
"""
Calculate sigmoid
"""
return 1 / (1 + np.exp(-x))
x = np.array([0.5, 0.1, -0.2])
target = 0.6
learnrate = 0.5
weights_input_hidden = np.array([[0.5, -0.6],
[0.1, -0.2],
[0.1, 0.7]])
weights_hidden_output = np.array([0.1, -0.3])
## Forward pass
hidden_layer_input = np.dot(x, weights_input_hidden)
hidden_layer_output = sigmoid(hidden_layer_input)
output_layer_in = np.dot(hidden_layer_output, weights_hidden_output)
output = sigmoid(output_layer_in)
## Backwards pass
## TODO: Calculate error
error = target - output
# TODO: Calculate error gradient for output layer
del_err_output = error * output * (1 - output)
# TODO: Calculate error gradient for hidden layer
del_err_hidden = np.dot(del_err_output, weights_hidden_output) * \
hidden_layer_output * (1 - hidden_layer_output)
# TODO: Calculate change in weights for hidden layer to output layer
delta_w_h_o = learnrate * del_err_output * hidden_layer_output
# TODO: Calculate change in weights for input layer to hidden layer
delta_w_i_h = learnrate * del_err_hidden * x[:, None]
print('Change in weights for hidden layer to output layer:')
print(delta_w_h_o)
print('Change in weights for input layer to hidden layer:')
print(delta_w_i_h)
import numpy as np
import pandas as pd
data=pd.read_csv("binary.csv")
data['rank'].astype(int)
def rank1_std(x):
if x['rank'] == 1:
return 1
else:
return 0
def rank2_std(x):
if x['rank'] == 2:
return 1
else:
return 0
def rank3_std(x):
if x['rank'] == 3:
return 1
else:
return 0
def rank4_std(x):
if x['rank'] == 4:
return 1
else:
return 0
def admit_std(x):
if x['admit']:
return True
else:
return False
data['rank_1']=data.apply(rank1_std,axis=1)
data['rank_2']=data.apply(rank2_std,axis=1)
data['rank_3']=data.apply(rank3_std,axis=1)
data['rank_4']=data.apply(rank4_std,axis=1)
data['admit']=data.apply(admit_std,axis=1)
gre_mean=data['gre'].mean()
gre_max=data['gre'].max()
gre_min=data['gre'].min()
gre_std=data['gre'].std()
gpa_mean=data['gpa'].mean()
gpa_max=data['gpa'].max()
gpa_min=data['gpa'].min()
gpa_std=data['gpa'].std()
data['gre']=data['gre'].map(lambda x: (x-gre_mean)/gre_std)
data['gpa']=data['gpa'].map(lambda x: (x-gpa_mean)/gpa_std)
del data['rank']
data.head(20)
import numpy as np
import pandas as pd
admissions = pd.read_csv("binary.csv")
# Make dummy variables for rank
data = pd.concat([admissions, pd.get_dummies(admissions['rank'], prefix='rank')], axis=1)
data = data.drop('rank', axis=1)
# Standarize features
for field in ['gre', 'gpa']:
mean, std = data[field].mean(), data[field].std()
data.loc[:,field] = (data[field]-mean)/std
# Split off random 10% of the data for testing
np.random.seed(21)
sample = np.random.choice(data.index, size=int(len(data)*0.9), replace=False)
data, test_data = data.ix[sample], data.drop(sample)
# Split into features and targets
features, targets = data.drop('admit', axis=1), data['admit']
features_test, targets_test = test_data.drop('admit', axis=1), test_data['admit']
# features=data[['gre','gpa','rank_1','rank_2','rank_3','rank_4']][:390]
# targets=data['admit'][:390]
# features_test=data[['gre','gpa','rank_1','rank_2','rank_3','rank_4']][390:]
# targets_test=data['admit'][390:]
# features.head(10)
np.random.seed(21)
def sigmoid(x):
"""
Calculate sigmoid
"""
return 1 / (1 + np.exp(-x))
# Hyperparameters
n_hidden = 2 # number of hidden units
epochs = 900
learnrate = 0.005
n_records, n_features = features.shape
last_loss = None
# Initialize weights
weights_input_hidden = np.random.normal(scale=1 / n_features ** .5,
size=(n_features, n_hidden))
weights_hidden_output = np.random.normal(scale=1 / n_features ** .5,
size=n_hidden)
for e in range(epochs):
del_w_input_hidden = np.zeros(weights_input_hidden.shape)
del_w_hidden_output = np.zeros(weights_hidden_output.shape)
for x, y in zip(features.values, targets):
## Forward pass ##
# TODO: Calculate the output
hidden_input = np.dot(x, weights_input_hidden)
hidden_output = sigmoid(hidden_input)
output = sigmoid(np.dot(hidden_output,
weights_hidden_output))
## Backward pass ##
# TODO: Calculate the error
error = y - output
# TODO: Calculate error gradient in output unit
output_error = error * output * (1 - output)
# TODO: propagate errors to hidden layer
hidden_error = np.dot(output_error, weights_hidden_output) * \
hidden_output * (1 - hidden_output)
# TODO: Update the change in weights
del_w_hidden_output += output_error * hidden_output
del_w_input_hidden += hidden_error * x[:, None]
# TODO: Update weights
weights_input_hidden += learnrate * del_w_input_hidden / n_records
weights_hidden_output += learnrate * del_w_hidden_output / n_records
# Printing out the mean square error on the training set
if e % (epochs / 10) == 0:
hidden_output = sigmoid(np.dot(x, weights_input_hidden))
out = sigmoid(np.dot(hidden_output,
weights_hidden_output))
loss = np.mean((out - targets) ** 2)
if last_loss and last_loss < loss:
print("Train loss: ", loss, " WARNING - Loss Increasing")
else:
print("Train loss: ", loss)
last_loss = loss
# Calculate accuracy on test data
hidden = sigmoid(np.dot(features_test, weights_input_hidden))
out = sigmoid(np.dot(hidden, weights_hidden_output))
predictions = out > 0.5
print(out)
print(predictions)
print(targets_test)
accuracy = np.mean(predictions == targets_test)
print("Prediction accuracy: {:.3f}".format(accuracy))
# for x, y in zip(features.values, targets):
# print(x)
# print(y)
print(True-0.5)
print(False-0.5)
```
| github_jupyter |
# Table of Contents
<div class="toc" style="margin-top: 1em;"><ul class="toc-item" id="toc-level0"></ul></div>
Superceded by Make Climatology
```
import netCDF4 as NC
import numpy as np
import matplotlib.pyplot as plt
from salishsea_tools import rivertools, river_201702, nc_tools
%matplotlib inline
import importlib
importlib.reload(rivertools)
#what type of file would you like to create?
#e.g. 'constant' - yearly average used for all months
# 'monthly' - monthly averages used for each month
rivertype = 'monthly'
bathy = '../../../nemo-forcing/grid/bathymetry_201702.nc'
# leave temperature at -99, I'm not setting it in this file
#get watershed fluxes from netcdf file
if rivertype == 'constant':
fluxfile = NC.Dataset('../../../nemo-forcing/rivers/Salish_allrivers_cnst.nc','r')
#inialise the runoff and run_depth arrays
runoff, run_depth, run_temp = rivertools.init_runoff_array(bathy=bathy, init_depth=3)
if rivertype == 'monthly':
fluxfile = NC.Dataset('../../../nemo-forcing/rivers/Salish_allrivers_monthly.nc','r')
#inialise the runoff and run_depth arrays
runoff, run_depth, run_temp = rivertools.init_runoff_array_monthly(bathy=bathy,
init_depth=3)
print (run_depth.max(), run_temp.max())
#get dimensions for netcdf files
fB = NC.Dataset(bathy, 'r')
lat = fB.variables['nav_lat']
lon = fB.variables['nav_lon']
D = fB.variables['Bathymetry'][:]
ymax, xmax = D.shape
print (lat[:].shape)
mesh = NC.Dataset('../../../NEMO-forcing/grid/mesh_mask201702.nc')
e1t = mesh.variables['e1t'][0,:]
e2t = mesh.variables['e2t'][0,:]
mesh.close()
#list of watersheds we are including
names = ['skagit', 'fraser', 'evi_n', 'howe', 'bute', 'puget', 'jdf', 'evi_s', 'jervis', 'toba']
prop_dict = river_201702.prop_dict
for name in range(0,len(names)):
watershedname = names[name]
print (watershedname)
Flux = fluxfile.variables[watershedname][:]
if rivertype == 'constant':
Flux = float(Flux)
runoff_orig = np.copy(runoff)
runoff, run_depth, run_temp = rivertools.put_watershed_into_runoff(rivertype,
watershedname, Flux, runoff, run_depth, run_temp,
use_prop_dict=True, prop_dict=prop_dict[watershedname])
if rivertype == 'constant':
rivertools.check_sum(runoff_orig, runoff, Flux)
if rivertype == 'monthly':
rivertools.check_sum_monthly(runoff_orig, runoff, Flux)
if rivertype == 'constant':
nemo = NC.Dataset('../../../NEMO-forcing/rivers/rivers_cnst_201702.nc', 'w')
# dimensions
nemo.createDimension('x', xmax)
nemo.createDimension('y', ymax)
nemo.createDimension('time_counter', None)
# variables
# latitude and longitude
nav_lat = nemo.createVariable('nav_lat', 'float32', ('y', 'x'), zlib=True)
nav_lat.setncattr('units', 'degrees_north')
nav_lat.setncattr('long_name', 'latitude')
nav_lat = lat
nav_lon = nemo.createVariable('nav_lon','float32',('y','x'),zlib=True)
nav_lon.setncattr('units', 'degrees_east')
nav_lon.setncattr('long_name', 'longitude')
nav_lon = lon
# time
time_counter = nemo.createVariable('time_counter', 'float32', ('time_counter'),zlib=True)
time_counter.setncattr('units', '1')
time_counter.setncattr('long_name', 'time')
time_counter[0] = 1
# runoff
rorunoff = nemo.createVariable('rorunoff', 'float32', ('time_counter','y','x'), zlib=True)
rorunoff.setncattr('units', 'kg m-2 s-1')
rorunoff.setncattr('long_name', 'runoff_flux')
rorunoff[0,:] = runoff
# depth
rodepth = nemo.createVariable('rodepth','float32',('y','x'),zlib=True)
rodepth.setncattr('units', 'm')
rodepth.setncattr('long_name', 'maximum_runoff_depth')
rodepth[:] = run_depth
nc_tools.init_dataset_attrs(nemo, 'Constant Rivers for Bathymetry 201702', 'Add Rivers Month and Constant-B201702',
'NEMO-forcing/rivers/rivers_cnst_201702.nc',
'Mean yearly flow for Rivers for Bathymetry 201702')
nc_tools.show_dataset_attrs(nemo)
nemo.setncattr('references', 'NEMO-forcing/rivers/rivers_cnst_201702.nc')
nc_tools.check_dataset_attrs(nemo)
nemo.close()
if rivertype == 'monthly':
nemo = NC.Dataset('rivers_month_201702.nc', 'w')
nc_tools.init_dataset_attrs(nemo, 'Monthly Rivers for Bathymetry 201702', 'Add Rivers Month and Constant-B201702',
'NEMO-forcing/rivers/rivers_month_201702.nc',
'Monthly Averages, Bathy 201702, all Four Fraser Arms from River Head')
# dimensions
nemo.createDimension('x', xmax)
nemo.createDimension('y', ymax)
nemo.createDimension('time_counter', None)
# variables
# latitude and longitude
nav_lat = nemo.createVariable('nav_lat', 'float32', ('y', 'x'), zlib=True)
nav_lat.setncattr('units', 'degrees_north')
nav_lat.setncattr('long_name', 'latitude')
nav_lat = lat
nav_lon = nemo.createVariable('nav_lon','float32',('y','x'),zlib=True)
nav_lon.setncattr('units', 'degrees_east')
nav_lon.setncattr('long_name', 'longitude')
nav_lon = lon
# time
time_counter = nemo.createVariable('time_counter', 'float32', ('time_counter'),zlib=True)
time_counter.setncattr('units', 'month')
time_counter.setncattr('long_name', 'time')
time_counter[0:12] = range(1,13)
# runoff
rorunoff = nemo.createVariable('rorunoff', 'float32', ('time_counter','y','x'), zlib=True)
rorunoff.setncattr('units', 'kg m-2 s-1')
rorunoff.setncattr('long_name', 'runoff_flux')
rorunoff[0:12,:] = runoff
# depth
rodepth = nemo.createVariable('rodepth','float32',('y','x'),zlib=True)
rodepth.setncattr('units', 'm')
rodepth.setncattr('long_name', 'maximum_runoff_depth')
rodepth[:] = run_depth[0,:,:]
nc_tools.show_dataset_attrs(nemo)
nemo.setncattr('references', 'NEMO-forcing/rivers/rivers_month_201702.nc')
nc_tools.check_dataset_attrs(nemo)
nemo.close()
%matplotlib inline
Fraser = 8.10384 * e1t[500, 395] * e2t[500, 395]
test = NC.Dataset('../../../NEMO-forcing/rivers/rivers_cnst_201702.nc','r')
test2 = NC.Dataset('../../../NEMO-forcing/rivers/rivers_cnst_extended.nc', 'r')
test3 = NC.Dataset('../../../NEMO-forcing/rivers/rivers_cnst.nc', 'r')
plotting = test.variables['rorunoff'][0, :, :]
plot2 = test2.variables['rorunoff'][0, :, :]
orig = test3.variables['rorunoff'][0, :, :]
plt.figure(figsize=(19, 19))
plt.pcolormesh(lon[:], lat[:], D, cmap='cool')
sumit1 = 0.
sumit2 = 0.
sumit3 = 0.
for i in range(898):
for j in range(398):
if plot2[i, j] > 0:
plt.plot(lon[i,j], lat[i,j],'ys') #down by one two
if plotting[i, j] > 0:
plt.plot(lon[i,j], lat[i,j], 'ko') # new
if orig[i, j] > 0:
plt.plot(lon[i,j], lat[i,j], 'r+') # orig
# if plot2[i, j] != plotting[i, j] or plot2[i, j] != orig[i, j] or plotting[i, j] != orig[i, j]:
# sumit1 += e1t[i, j] * e2t[i, j] * plotting[i, j]/Fraser
# sumit2 += e1t[i, j] * e2t[i, j] * plot2[i, j]/Fraser
# sumit3 += e1t[i, j] * e2t[i, j] * orig[i ,j]/Fraser
# print (i, j, e1t[i, j] * e2t[i, j] * plotting[i, j]/Fraser,
# e1t[i, j] * e2t[i, j] * plot2[i, j]/Fraser,
# e1t[i, j] * e2t[i, j] * orig[i, j]/Fraser, sumit1, sumit2, sumit3)
plt.colorbar()
plt.ylabel('Latitude')
plt.xlabel('Longitude')
plt.title('Location of rivers included, with depth contours [m]')
plt.savefig('river_locations.pdf')
6.07788/8.10384
temp = test2.variables['rotemper'][:]
roro = test2.variables['rorunoff'][:]
print (temp.shape)
plt.plot(temp[:,351,345])
print (temp.max())
if rivertype == 'monthly':
depths = D[roro[0] > 0.]
ii, jj = np.where(roro[0] > 0.)
elif rivertype == 'constant':
depths = D[roro > 0.]
ii, jj = np.where(roro > 0.)
print (ii[depths.mask], jj[depths.mask])
ip = ii[depths.mask]; jp = jj[depths.mask]
fig, ax = plt.subplots(1,1,figsize=(5,5))
ax.pcolormesh(D)
ax.plot(jp[0], ip[0],'mo')
ax.set_xlim((35, 65))
ax.set_ylim((870, 898))
ax.plot(45+0.5, 891+0.5,'r*')
fig, ax = plt.subplots(1,1,figsize=(5,5))
ax.pcolormesh(D)
ax.plot(jp[:3], ip[:3],'mo')
ax.set_xlim((300, 320))
ax.set_ylim((635, 655))
ax.plot(307+0.5, 651+0.5,'r*')
ax.plot(309+0.5, 650+0.5,'r*')
ax.plot(310+0.5, 649+0.5,'r*')
test.close()
test2.close()
nemo.close()
test = NC.Dataset('/results/forcing/rivers/RLonFraCElse_y2017m05d13.nc.aside')
test2 = NC.Dataset('/results/forcing/rivers/junk_RLonFraCElse_y2017m05d13.nc')
runoff = test.variables['rorunoff'][:]
new_runoff = test2.variables['rorunoff'][:]
print(np.max(runoff), np.max(new_runoff))
print(np.min(runoff-new_runoff), np.max(runoff-new_runoff))
print (runoff.shape)
print(runoff[0, 500, 395]-22.9068)
print(new_runoff[0, 500, 395]-22.9068)
!ls -l /results/forcing/rivers/RLonFraCElse_y2017m05d13.nc.aside
!ls -l /results/forcing/rivers/junk_RLonFraCElse_y2017m05d13.nc
```
| github_jupyter |
# Superruns
### Basic concept of a superrun:
A superrun is made up of many regular runs and helps us therefore to organize data in logic units and to load it faster. In the following notebook we will give some brief examples how superruns work and can be used to make analysts lives easier.
Lets get started how we can define superruns. The example I demonstrate here is based on some dummy Record and Peak plugins. But it works in the same way for regular data.
```
import strax
import straxen
```
### Define context and create some dummy data:
In the subsequent cells I create a dummy context and write some dummy-data. You can either read through it if you are interested or skip until **Define a superrun**. For the working examples on superruns you only need to know:
* Superruns can be created with any of our regular online and offline contexts.
* In the two cells below I define 3 runs and records for the run_ids 0, 1, 2.
* The constituents of a superrun are called subruns which we call runs.
```
from strax.testutils import Records, Peaks
superrun_name='_superrun_test'
st = strax.Context(storage=[strax.DataDirectory('./strax_data',
provide_run_metadata=True,
readonly=False,
deep_scan=True)],
register=[Records, Peaks],
config={'bonus_area': 42}
)
st.set_context_config({ 'use_per_run_defaults': False})
import datetime
import pytz
import numpy as np
import json
from bson import json_util
def _write_run_doc(context, run_id, time, endtime):
"""Function which writes a dummy run document.
"""
run_doc = {'name': run_id, 'start': time, 'end': endtime}
with open(context.storage[0]._run_meta_path(str(run_id)), 'w') as fp:
json.dump(run_doc, fp,sort_keys=True, indent=4, default=json_util.default)
offset_between_subruns = 10
now = datetime.datetime.now()
now.replace(tzinfo=pytz.utc)
subrun_ids = [str(r) for r in range(3)]
for run_id in subrun_ids:
rr = st.get_array(run_id, 'records')
time = np.min(rr['time'])
endtime = np.max(strax.endtime(rr))
_write_run_doc(st,
run_id,
now + datetime.timedelta(0, int(time)),
now + datetime.timedelta(0, int(endtime)),
)
st.set_config({'secret_time_offset': endtime + offset_between_subruns}) # untracked option
assert st.is_stored(run_id, 'records')
```
If we print now the lineage and hash for the three runs you will see it is equivalent to our regular data.
```
print(st.key_for('2', 'records'))
st.key_for('2', 'records').lineage
```
### Metadata of our subruns:
To understand a bit better how our dummy data looks like we can have a look into the metadata for a single run. Each subrun is made of 10 chunks each containing 10 waveforms in 10 different channels.
```
st.get_meta('2', 'records')
```
### Define a superrun:
Defining a superrun is quite simple one has to call:
```
st.define_run(superrun_name, subrun_ids)
print('superrun_name: ', superrun_name, '\n'
'subrun_ids: ', subrun_ids
)
```
where the first argument is a string specifying the name of the superrun e.g. `_Kr83m_20200816`. Please note that superrun names must start with an underscore.
The second argument is a list of run_ids of subruns the superrun should be made of. Please note that the definition of a superrun does not need any specification of a data_kind like peaks or event_info because it is a "run".
By default it is only allowed to store new runs under the usere's specified strax_data directory. In this example it is simply `./strax_data` and the run_meta data can be looked at via:
```
st.run_metadata('_superrun_test')
```
The superrun-metadata contains a list of all subruns making up the superrun, the start and end time (in milliseconds) of the corresponding collections of runs and its naive livetime in nanoseconds without any corrections for deadtime.
Please note that in the presented example the time difference between start and end time is 50 s while the live time is only about 30 s. This comes from the fact that I defined the time between two runs to be 10 s. It should be always kept in mind for superruns that livetime is not the same as the end - start of the superrun.
The superun will appear in the run selection as any other run:
```
st.select_runs()
```
### Loading data with superruns:
Loading superruns can be done in two different ways. Lets try first the already implemented approach and compare the data with loading the individual runs separately:
```
sub_runs = st.get_array(subrun_ids, 'records') # Loading all subruns individually like we are used to
superrun = st.get_array(superrun_name, 'records') # Loading the superrun
assert np.all(sub_runs['time'] == superrun['time']) # Comparing if the data is the same
```
To increase the loading speed it can be allowed to skip the lineage check of the individual subruns:
```
sub_runs = st.get_array(subrun_ids, 'records')
superrun = st.get_array(superrun_name, 'records', _check_lineage_per_run_id=False)
assert np.all(sub_runs['time'] == superrun['time'])
```
So how does this magic work? Under the hood a superrun first checks if the data of the different subruns has been created before. If not it will make the data for you. After that the data of the individual runs is loaded.
The loading speed can be further increased if we rechunk and write the data of our superrun as "new" data to disk. This can be done easily for light weight data_types like peaks and above. Further, this allows us to combine multiple data_types if the same data_kind, like for example `event_info` and `cuts`.
### Writing a "new" superrun:
To write a new superrun one has to set the corresponding context setting to true:
```
st.set_context_config({'write_superruns': True})
st.is_stored(superrun_name, 'records')
st.make(superrun_name, 'records')
st.is_stored(superrun_name, 'records')
```
Lets see if the data is the same:
```
sub_runs = st.get_array(subrun_ids, 'records')
superrun = st.get_array(superrun_name, 'records', _check_lineage_per_run_id=False)
assert np.all(sub_runs['time'] == superrun['time'])
```
And the data will now shown as available in select runs:
```
st.select_runs(available=('records', ))
```
If a some data does not exist for a super run we can simply created it via the superrun_id. This will not only create the data of the rechunked superrun but also the data of the subrungs if not already stored:
```
st.is_stored(subrun_ids[0], 'peaks')
st.make(superrun_name, 'peaks')
st.is_stored(subrun_ids[0], 'peaks')
peaks = st.get_array(superrun_name, 'peaks')
```
**Some developer information:**
In case of a stored and rechunked superruns every chunk has also now some additional information about the individual subruns it is made of:
```
for chunk in st.get_iter(superrun_name, 'records'):
chunk
chunk.subruns, chunk.run_id
```
The same goes for the meta data:
```
st.get_meta(superrun_name, 'records')['chunks']
```
| github_jupyter |
## q<sup>2</sup> Tutorial - 1. Introduction
Welcome to the q<sup>2</sup> Python package!
*Before you begin this Tutorial, make sure that you have MOOGSILENT installed; it is assumed that you have some experience using <a href="http://www.as.utexas.edu/~chris/moog.html">MOOG</a>. If you are not familiar with Python, know that this package has a number of dependencies such as numpy, scipy, matplotlib, and bokeh. The easiest way to satisfy all these requirements is to install the <a href="https://www.continuum.io/downloads">Anaconda</a> Python distribution. Also, it is best to follow this Tutorial on an IPython notebook. Finally, you should have the directory in which q<sup>2</sup> is located in your PYTHONPATH environment variable.*
Begin by importing the q<sup>2</sup> package:
```
import q2
```
For any given q<sup>2</sup> project, two csv (comma-separated-values) files are required: one with star data such as Teff, logg, etc., and another one with spectral line data, i.e., a line list with atomic data and measured equivalent widths (EWs).
Take a look at the two files used in this example: "standard_stars.csv" and "standards_lines.csv", to have an idea of the format (you should be able to open these files with MS Excel, LibreOffice, or any text editor). Column order is not important, but a few columns are required: id, teff, logg, feh, and vt in the "stars" file, and wavelength, species, ep, and gf in the "lines" file. The EWs should be given in the "lines" file with a column header corresponding to the id of the star.
Here, the data from the csv files are loaded into an object called data (nothing will happen to the csv files from now on; in fact, q<sup>2</sup> will never modify them):
```
data = q2.Data('standards_stars.csv', 'standards_lines.csv')
```
We can inspect our data by calling the object attributes "star_data" and "lines". For example, let's check the stars' ids, Teffs, and the first five EWs of the Sun (compare this output with the data in the csv files to understand what you are doing here):
```
print(data.star_data['id'])
print(data.star_data['teff'])
print(data.lines['Sun'][:5])
```
To begin using q<sup>2</sup>'s tools we need to create Star objects (note that we create the star objects with the star's names as the first parameter and exacly as listed in the csv files, or the data object; if you create the star object with a star name that is not in the data object, e.g., "q2.Star('Sol')" or "q2.Star('arcturus')", q<sup>2</sup> will not be able to figure out what to do with that star):
```
sun = q2.Star('Sun')
arcturus = q2.Star('Arcturus')
print(sun)
```
At this point, the sun object has no parameters, spectral lines, or a model atmosphere; it is just a bare star object (the same is true for arcturus).
To asign the attributes to these stars' objects we use the "get_data_from" method. Our csv files are no longer being used to get this information; the data object is instead employed for that:
```
sun.get_data_from(data)
arcturus.get_data_from(data)
print(sun)
print("")
print(arcturus)
```
Since we also uploaded a master line list (the "lines" csv file), the star objects now also have their own linelist data. These data were also asigned by the "get_data_from" method. We can inspect these linelist details as follows:
```
print("Wavelength column (first five):")
print(sun.linelist['wavelength'][:5])
print("Equivalent width column (first five):")
print(sun.linelist['ew'][:5])
```
Our stars have all fundamental atmospheric parameters defined (they can be input guess values at this point). Therefore, we can use the "get_model_atmosphere" method to look for models or interpolate them from the available grids. Here we will use MARCS models, hence the 'marcs' argument that is passed to this method. For Kurucz models we can use 'odfnew' (the default), 'aodfnew', 'over', or 'nover'.
```
sun.get_model_atmosphere('marcs')
arcturus.get_model_atmosphere('marcs')
print(sun)
```
We just added an attribute model_atmosphere to our star objects sun and arcturus. At any point, we can check what grid the model attached to a given Star object was taken or interpolated from by looking at the model_atmosphere_grid attribute (or just by printing the object as done above for sun). The code below makes the classic RHOX vs T graph (after importing matplotlib; the "%matplotlib inline" command allows the images created by q<sup>2</sup> to be shown on the IPython notebook):
```
import matplotlib.pylab as plt
%matplotlib inline
plt.plot(sun.model_atmosphere['RHOX'], sun.model_atmosphere['T'])
plt.xlabel("RHOX")
plt.ylabel("T (K)")
plt.title("Interpolated solar model atmosphere. Grid: {}".\
format(sun.model_atmosphere_grid));
```
Note that the "model_atmosphere" attribute/dictionary has all the necessary physical information to run a MOOG abfind calculation (and our star objects have the linelist data as well). The q2.moog module can help us create input files for a typical MOOG abfind run:
```
md = q2.moog.Driver()
md.create_file() #Creates a simple MOOGSILENT abfind driver
q2.moog.create_model_in(sun) #creates the model atmosphere file
q2.moog.create_lines_in(sun); #creates the linelist file
```
Have a look at the files you just created and inspect them. The files created (batch.par, model.in, lines.in) should have the familiar MOOG formats. At this point, you could go to your terminal and run MOOGSILENT (or MOOG and then enter batch.par as the driver). Of course, it is not terribly helpful if we are still going to have to call MOOG manually from the terminal. Instead, we can use q<sup>2</sup>'s "moog.abfind" function to calculate abundances running MOOG under the hood (let's keep our fingers crossed; this is when q<sup>2</sup> actually talks to MOOG!). In this example, we are computing line-by-line "FeII abundances" for the Sun:
```
q2.moog.abfind(sun, 26.1, 'fe2')
print(sun.fe2.keys())
print(sun.fe2['ww'])
print(sun.fe2['ab'])
```
The "moog.abfind" function ran MOOGSILENT for us, creating the MOOG input files and deleting them when done (all within a hidden folder .q2). The sun object in this example now has an fe2 attribute, which is a dictionary containing the MOOG outputs: ww=wavelength, ab=abundance, etc.
A common exercise in stellar spectroscopy is to derive iron abundances from FeI and FeII lines separately, in addition to calculating some statistics like abundance versus EP or REW slopes. This can be done with the "specpars" module as follows:
```
q2.specpars.iron_stats(sun, plot='sun')
```
The q2.specpars.iron_stats function makes a figure with the standard abundance versus EP/REW/wavelength plots. Look for the "sun.png" file for a hardcopy of this figure; the name of this file is set by the "plot" argument. In the figure, blue crosses are the FeI lines and green circles are the FeII lines. The solid blue lines are linear fits to the FeI data and the black line in the bottom panel is a horizontal line at the average value of the iron abundance.
The sun object now has an "iron_stats" attribute with the results of this calculation, so we can use them for further analysis. For example, let's have a look at the important results from the calculation above, namely average iron abundances from FeI and FeII lines and the EP/REW slopes:
```
print("A(Fe I) = {0:5.3f} +/- {1:5.3f}".\
format(sun.iron_stats['afe1'], sun.iron_stats['err_afe1']))
print("A(Fe II) = {0:5.3f} +/- {1:5.3f}".\
format(sun.iron_stats['afe2'], sun.iron_stats['err_afe2']))
print("A(FeI) vs. EP slope = {0:.4f} +/- {1:.4f}".\
format(sun.iron_stats['slope_ep'], sun.iron_stats['err_slope_ep']))
print("A(FeI) vs. REW slope = {0:.4f} +/- {1:.4f}".\
format(sun.iron_stats['slope_rew'], sun.iron_stats['err_slope_rew']))
```
There is some flexibility to modify the plots generated by q<sup>2</sup> with a PlotPars object that we will call "pp" here. This object can be passed on to q2.specpars.iron_stats. For example, we can force the y-axis to go from 7.1 to 7.7 using the "afe" controller. We can also change the default title and add an inside title as shown below. You could use these options to get rid of the titles as well (e.g., pp.title = '', pp.title_inside = ''):
```
pp = q2.specpars.PlotPars()
pp.afe = [7.1, 7.7]
pp.title = 'Abundances computed with q$^2$'
pp.title_inside = 'Sun'
q2.specpars.iron_stats(sun, plot='sun', PlotPars=pp)
```
The pp.afe controller sets the range of the y-axis: [ymin, ymax]. However, when ymin is set to -1000, q<sup>2</sup> makes the y-axis centered around the derived mean iron abundance and extends it to +/-ymax. Next we are going to redo the plots above with the default PlotPars parameters, except that the y-axis range is set to +/-0.25 from the average iron abundance:
```
pp = q2.specpars.PlotPars()
pp.afe = [-1000, 0.25]
q2.specpars.iron_stats(sun, plot='sun', PlotPars=pp)
```
Let's do the same thing for Arcturus (note that we are controlling the plots' appearance with pp as well so they are consistent with the Sun's plots, particularly the y-axis range; look for the "arcturus.png" file after running this command):
```
q2.specpars.iron_stats(arcturus, plot='arcturus', PlotPars=pp)
```
We have used q<sup>2</sup> to calculate "absolute" iron abundances of the Sun and Arcturus. Another common practice in this field is to compute "differential" abundances, i.e., abundances in a given problem star relative to a reference. In these cases, it is common to use the Sun as the reference star.
To calculate differential abundances we can also use q2.specpars.iron_stats. For example, if we want to calculate the differential iron abundances of Arcturus relative to the Sun all we have to do is pass the object sun as Ref(erence):
```
q2.specpars.iron_stats(arcturus, Ref=sun,
plot='arcturus_sun', PlotPars=pp)
```
This created a figure "arcturus_sun.png". Note that the y-axis label is not A(Fe), but [Fe/H], which is the standard notation for differential abundance.
Note that we didn't have to deal with the MOOG inputs or outputs, match line lists, create tables and do the statistics separately. All that was handled by q<sup>2</sup>. The single-line command above did the following:
- Created MOOG inputs for the Sun, ran MOOG, read the outputs, and attached the results to the sun object as sun.fe1, sun.fe2, and sun.iron_stats. Deleted the MOOG input and output files.
- Did the same thing for Arcturus.
- Matched the linelists of the Sun and Arcturus (sun.fe1 to arcturus fe1, etc.), subtracted the abundances measured on a line-by-line basis, and added a column "difab' to the arcturus.fe1 and arcturus.fe2 attributes, which contains these results.
- Calculated the slopes of 'difab' versus EP, REW and other common statistics.
- Made the figure.
Below we can see the abundances measured from FeII lines for Arcturus, first the absolute ones, and then the differential ones. You can always check what the reference star was to calculate those 'difab' values by looking at the 'reference' column of the iron_stats attribute:
```
print(arcturus.fe2['ab'])
print(arcturus.fe2['difab'])
print(arcturus.iron_stats['reference'])
```
This concludes the Introduction. So far we have calculated iron abundances using input parameters as given in our csv files. As shown in the last figure we made, the iron abundances of Arcturus show trends with excitation potential and line-strength. Also, the mean FeI and FeII abundances are a bit different. This could be indicating that the input parameters used are not correct (within our 1D-LTE model assumptions). Let's move on to Part 2 of this Tutorial and use q<sup>2</sup> to derive "better" spectroscopic stellar parameters of stars.
| github_jupyter |
# Initialization
```
from fnmatch import filter
import matplotlib.pyplot as plt
from matplotlib import rc, rcParams, dates, lines
import numpy as np
import datetime as dt
import spacepy.pycdf as cdf
import spacepy.toolbox as tb
import bisect as bi
import seaborn as sns
import os
from scipy.integrate import simps
import scipy.constants as c
from multilabel import multilabel
sns.set_context('talk')
sns.set_style('ticks')
sns.set_palette('muted', color_codes=True)
rc('text', usetex=True)
rc('font', family='Mono')
rcParams['text.latex.preamble']=[r'\usepackage{amsmath}']
```
# Code
## Input
```
# spe = 'H'
# start = dt.datetime(2013, 7, 11, 13, 0)
# stop = dt.datetime(2013, 7, 11, 16, 0)
# craft = 'A'
# spe = 'H'
# start = dt.datetime(2013, 10, 10, 0, 0)
# stop = dt.datetime(2013, 10, 10, 3, 0)
# craft = 'B'
spe = 'H'
start = dt.datetime(2013, 7, 6, 11, 0)
stop = dt.datetime(2013, 7, 6, 14, 0)
craft = 'B'
# spe = 'H'
# start = dt.datetime(2013, 7, 6, 2, 0)
# stop = dt.datetime(2013, 7, 6, 5, 0)
# craft = 'B'
# spe = 'H'
# start = dt.datetime(2014, 2, 20, 4, 0)
# stop = dt.datetime(2014, 2, 20, 7, 0)
# craft = 'B'
# spe = 'H'
# start = dt.datetime(2014, 2, 15, 9, 0)
# stop = dt.datetime(2014, 2, 15, 12, 0)
# craft = 'A'
# spe = 'H'
# start = dt.datetime(2014, 2, 10, 9, 0)
# stop = dt.datetime(2014, 2, 10, 12, 0)
# craft = 'B'
```
## Processing
```
#Load data from RBSPICE
if spe == 'H':
sp = 'P'
else:
sp = spe
Hpath = r'C:\Users\Rjc12\WorkSchool\Work\CDFs'+'\\'+craft+r'\TOFxE'+spe+'\\'
Hfname = filter(os.listdir(Hpath), '*'+start.strftime('%Y%m%d')+'*')[-1]
Hcdf = cdf.CDF(Hpath+Hfname)
epoch = Hcdf['Epoch'][...]
Flux = Hcdf['F'+sp+'DU'][...]*(6.242e12)*10000
Energies = Hcdf['F'+sp+'DU_Energy'][...]*(1.60218e-13)
L = Hcdf['L'][...]
MLT = Hcdf['MLT'][...]
PAs = np.radians(Hcdf['PA_Midpoint'][...])
Hcdf.close()
for dater in [start + dt.timedelta(days=x) for x in range(1, (stop.day-start.day)+1)]:
Hpath = r'C:\Users\Rjc12\WorkSchool\Work\CDFs'+'\\'+craft+r'\TOFxEH\\'
Hfname = filter(os.listdir(Hpath), '*'+dater.strftime('%Y%m%d')+'*')[0]
Hcdf = cdf.CDF(Hpath+Hfname)
epoch = np.append(epoch, Hcdf['Epoch'][...])
Flux = np.append(Flux, Hcdf['F'+sp+'DU'][...]*(6.242e12)*10000, axis=0)
L = np.append(L, Hcdf['L'][...])
MLT = np.append(MLT, Hcdf['MLT'][...])
Hcdf.close()
#Load data from HOPE
Hopath = r'C:\Users\Rjc12\WorkSchool\Work\CDFs'+'\\'+craft+r'\HOPE\\'
Hofname = filter(os.listdir(Hopath), '*'+start.strftime('%Y%m%d')+'*')[-1]
Hocdf = cdf.CDF(Hopath+Hofname)
Hoepoch = Hocdf['Epoch_Ion'][...]
HoFlux = Hocdf['F'+sp+'DU'][...]*(6.242e15)*10000
HoEnergies = Hocdf['HOPE_ENERGY_Ion'][...]
HoPAs = np.radians(Hocdf['PITCH_ANGLE'][...])
Hocdf.close()
for dater in [start + dt.timedelta(days=x) for x in range(1, (stop.day-start.day)+1)]:
Hopath = r'C:\Users\Rjc12\WorkSchool\Work\CDFs'+'\\'+craft+r'\HOPE\\'
Hofname = filter(os.listdir(Hopath), '*'+dater.strftime('%Y%m%d')+'*')[0]
Hocdf = cdf.CDF(Hopath+Hofname)
Hoepoch = np.append(Hoepoch, Hocdf['Epoch_Ion'][...])
HoFlux = np.append(HoFlux, Hocdf['F'+sp+'DU'][...]*(6.242e15)*10000, axis=0)
Hocdf.close()
#Load data from EMFISIS
Bpath = r'C:\Users\Rjc12\WorkSchool\Work\CDFs'+'\\'+craft+r'\EMFISIS'+'\\'
Bfname = filter(os.listdir(Bpath), '*'+start.strftime('%Y%m%d')+'*')[-1]
Bcdf = cdf.CDF(Bpath+Bfname)
Bepoch = Bcdf['Epoch'][...]
Bmag = Bcdf['Magnitude'][...]
Binvalid = Bcdf['magInvalid'][...]
Bcdf.close()
for dater in [start + dt.timedelta(days=x) for x in range(1, (stop.day-start.day)+1)]:
Bpath = r'C:\Users\Rjc12\WorkSchool\Work\CDFs'+'\\'+craft+r'\EMFISIS\\'
Bfname = filter(os.listdir(Bpath), '*'+dater.strftime('%Y%m%d')+'*')[0]
Bcdf = cdf.CDF(Bpath+Bfname)
Bepoch = np.append(Bepoch, Bcdf['Epoch'][...])
Bmag = np.append(Bmag, Bcdf['Magnitude'][...])
Bcdf.close()
#Get array indices that match time window
fidx = bi.bisect_left(epoch, start)
lidx = bi.bisect_left(epoch, stop)
Hofidx = bi.bisect_left(Hoepoch, start)
Holidx = bi.bisect_left(Hoepoch, stop)
Bfidx = bi.bisect(Bepoch, start)
Blidx = bi.bisect(Bepoch, stop)
#Make sure that B is valid
if np.any(Binvalid[Bfidx:Blidx]):
raise ValueError('Bmag is not valid for this time period!')
#Make sure HOPE energy bins don't change in the time window
if np.any(HoEnergies[Hofidx:Holidx,:] != HoEnergies[Hofidx,:]):
raise ValueError('HOPE energies change!')
else:
HoEnergies = HoEnergies[Hofidx,:]
#Discard low energies from HOPE
LowEidx = bi.bisect(HoEnergies, 300)
HoFlux = HoFlux[:,:,LowEidx:]
HoEnergies = HoEnergies[LowEidx:]*(1.60218e-19)
#Convert datetimes into a format that simps can use (seconds since the epoch)
epsec = [(ep-dt.datetime(1970,1,1)).total_seconds() for ep in epoch[fidx:lidx]]
Hoepsec = [(ep-dt.datetime(1970,1,1)).total_seconds() for ep in Hoepoch[Hofidx:Holidx]]
Bepsec = [(ep-dt.datetime(1970,1,1)).total_seconds() for ep in Bepoch[Bfidx:Blidx]]
#Replace bad data with the nearest valid data (going backwards in time)
for i in range(len(PAs)):
for j in range(len(Energies)):
bads = []
for k in range(fidx, lidx):
if Flux[k,j,i]<0 or np.isnan(Flux[k,j,i]):
for l in range(1, k+2):
if l == k+1:
Flux[k,j,i] = 0
if l in bads:
continue
if Flux[k-l,j,i] >= 0:
Flux[k,j,i] = Flux[k-l, j, i]
break
bads.append(l)
for j in range(len(HoPAs)):
for i in range(len(HoEnergies)):
bads = []
for k in range(Hofidx, Holidx):
if HoFlux[k,j,i]<0 or np.isnan(HoFlux[k,j,i]):
for l in range(1, k+2):
if l == k+1:
HoFlux[k,j,i] = 0
if l in bads:
continue
if HoFlux[k-l, j,i] >= 0:
HoFlux[k,j,i] = HoFlux[k-l, j, i]
break
bads.append(l)
#Resample B so that it matches RBSPICE samples
Binter = np.interp(epsec, Bepsec, Bmag[Bfidx:Blidx])
Bpres = ((Binter*c.nano)**2)/(2*c.mu_0)
#Calculate Beta/eV
if spe == 'H':
m = c.m_p
elif spe == 'He':
m = 4*c.m_p
elif spe == 'O':
m = 16*c.m_p
TBeta = np.zeros((len(Energies), len(PAs)))
#First integrate over the time window while dividing by B
for i in range(len(Energies)):
for j in range(len(PAs)):
TBeta[i,j] = simps(Flux[fidx:lidx,i,j]/Bpres, x=epsec)
Betaperp = np.zeros(len(Energies))
Betapara = np.zeros(len(Energies))
#Integrate over the PAs to get Beta/eV, perpendicular and parallel
for i, E in enumerate(Energies):
Betaperptemp = np.zeros(len(PAs))
for j, PA in enumerate(PAs):
Betaperptemp[j] = np.pi*np.sqrt(2*m*E)*(np.sin(PA)**3)*TBeta[i,j]
Betaperp[i] = simps(Betaperptemp, x=PAs)
for i, E in enumerate(Energies):
Betaparatemp = np.zeros(len(PAs))
for j, PA in enumerate(PAs):
Betaparatemp[j] = 2*np.pi*np.sqrt(2*m*E)*(np.sin(PA)*np.cos(PA)**2)*TBeta[i,j]
Betapara[i] = simps(Betaperptemp, x=PAs)
#Calculate total Beta/eV
Beta = (1/3)*(Betapara+2*Betaperp)
#Calculate Beta/eV from HOPE in the same manner
HoBinter = np.interp(Hoepsec, Bepsec, Bmag[Bfidx:Blidx])
HoBpres = ((HoBinter*c.nano)**2)/(2*c.mu_0)
THoBeta = np.zeros((len(HoPAs), len(HoEnergies)))
for i in range(len(HoPAs)):
for j in range(len(HoEnergies)):
THoBeta[i,j] = simps(HoFlux[Hofidx:Holidx,i,j]/HoBpres, x=Hoepsec)
HoBetaperp = np.zeros(len(HoEnergies))
HoBetapara = np.zeros(len(HoEnergies))
for i, E in enumerate(HoEnergies):
HoBetaperptemp = np.zeros(len(HoPAs))
for j, PA in enumerate(HoPAs):
HoBetaperptemp[j] = np.pi*np.sqrt(2*m*E)*(np.sin(PA)**3)*THoBeta[j,i]
HoBetaperp[i] = simps(HoBetaperptemp, x=HoPAs)
for i, E in enumerate(HoEnergies):
HoBetaparatemp = np.zeros(len(HoPAs))
for j, PA in enumerate(HoPAs):
HoBetaparatemp[j] = 2*np.pi*np.sqrt(2*m*E)*(np.sin(PA)*np.cos(PA)**2)*THoBeta[j,i]
HoBetapara[i] = simps(HoBetaperptemp, x=HoPAs)
HoBeta = (1/3)*(HoBetapara+2*HoBetaperp)
#Calculate Beta from first energy bin up to x-th energy bin
#use even='first', so that simpsons rule is always used on the last interval, which is changing
BetaCDF = []
OBetaCDF = []
TBeta = np.append(HoBeta, Beta[1:])
TEnergies = np.append(HoEnergies, Energies[1:])
for x in range(2, len(TEnergies)):
BetaCDF = np.append(BetaCDF, simps(TBeta[:x], x=TEnergies[:x], even='first'))
for x in range(2, len(Energies)):
OBetaCDF = np.append(OBetaCDF, simps(Beta[:x], x=Energies[:x], even='first'))
#Average HOPE flux to smooth zig-zags
HosBeta = [HoBeta[0]]
for i in range(len(HoBeta)-2):
HosBeta = np.append(HosBeta, np.average([HoBeta[i], HoBeta[i+1]]))
HosBeta = np.append(HosBeta, HoBeta[-1])
```
## Plotting
```
plt.close('all')
fig, ax = plt.subplots()
ax2 = ax.twinx()
ax.semilogx(HoEnergies*(6.242e18), HosBeta, 'k', label='HOPE', marker='o', markerfacecolor='r', markeredgewidth=1)
ax.semilogx(HoEnergies*(6.242e18), HosBeta*3, 'k', label='HOPE*3', marker='o', markerfacecolor='g', markeredgewidth=1)
ax.semilogx(Energies*(6.242e18), Beta, 'k', label='RBSPICE TOFxE', marker='o', markerfacecolor='b', markeredgewidth=1)
Width = np.diff(TEnergies*(6.242e18))
NBetaCDF = BetaCDF/np.amax(BetaCDF)
NOBetaCDF = OBetaCDF/np.amax(BetaCDF)
Hob = ax2.bar((TEnergies*(6.242e18))[1:len(HoEnergies)], NBetaCDF[:len(HoEnergies)-1],
width=Width[1:len(HoEnergies)], color='r', label='HOPE')
RBHob = ax2.bar((TEnergies*(6.242e18))[len(HoEnergies):-1], NBetaCDF[len(HoEnergies)-1:],
width=Width[len(HoEnergies):], color='m', label='HOPE and RBSPICE\nw/o RBSPICE first energy channel')
RBb = ax2.bar((Energies*(6.242e18))[1:-1], NOBetaCDF,
width=Width[len(HoEnergies):], color='b', label='RBSPICE TOFxE')
Ho = lines.Line2D([], [], color='k', marker='o', markerfacecolor='r', markeredgewidth=1, label='HOPE')
Ho3 = lines.Line2D([], [], color='k', marker='o', markerfacecolor='g', markeredgewidth=1, label='HOPE*3')
RB = lines.Line2D([], [], color='k', marker='o', markerfacecolor='b', markeredgewidth=1, label='RBSPICE TOFxE')
hands = [Ho, Ho3, RB, Hob, RBHob, RBb]
labs = ['HOPE', 'HOPE*3', 'RBSPICE TOFxE', 'HOPE', 'HOPE and RBSPICE\nw/o RBSPICE first energy channel', 'RBSPICE TOFxE']
ax.legend(hands, labs, loc=2)
ax.set_zorder(2)
ax.patch.set_visible(False)
fig.suptitle(r'H$^+ \boldsymbol{\beta}/eV$ Contribution vs Energy for '+start.strftime('%m/%d/%y, %H:%M-')+stop.strftime('%H:%M'))
ax.set_ylabel(r'Total $\boldsymbol{\beta}/eV$ Contribution for Time Period')
ax2.set_ylabel(r'Normalized Cumulative $\boldsymbol{\beta}$ Contribution for Time Period')
ax.set_xlabel('Energy (eV)')
# plt.savefig(craft+start.strftime('%Y%m%d-%H%M-')+stop.strftime('%H%M-')
# +'timeIntegrated.png', format='png', dpi=100)
plt.show()
#Create a similar graph, using HOPE*3 for the Beta calculation
BetaCDF = []
OBetaCDF = []
TBeta = np.append(HoBeta*3, Beta[1:])
TEnergies = np.append(HoEnergies, Energies[1:])
for x in range(2, len(TEnergies)):
BetaCDF = np.append(BetaCDF, simps(TBeta[:x], x=TEnergies[:x], even='first'))
for x in range(2, len(Energies)):
OBetaCDF = np.append(OBetaCDF, simps(Beta[:x], x=Energies[:x], even='first'))
HosBeta = [HoBeta[0]]
for i in range(len(HoBeta)-2):
HosBeta = np.append(HosBeta, np.average([HoBeta[i], HoBeta[i+1]]))
HosBeta = np.append(HosBeta, HoBeta[-1])
plt.close('all')
fig, ax = plt.subplots()
ax2 = ax.twinx()
ax.semilogx(HoEnergies*(6.242e18), HosBeta, 'k', label='HOPE', marker='o', markerfacecolor='r', markeredgewidth=1)
ax.semilogx(HoEnergies*(6.242e18), HosBeta*3, 'k', label='HOPE*3', marker='o', markerfacecolor='g', markeredgewidth=1)
ax.semilogx(Energies*(6.242e18), Beta, 'k', label='RBSPICE TOFxE', marker='o', markerfacecolor='b', markeredgewidth=1)
Width = np.diff(TEnergies*(6.242e18))
NBetaCDF = BetaCDF/np.amax(BetaCDF)
NOBetaCDF = OBetaCDF/np.amax(BetaCDF)
Hob = ax2.bar((TEnergies*(6.242e18))[1:len(HoEnergies)], NBetaCDF[:len(HoEnergies)-1],
width=Width[1:len(HoEnergies)], color='r', label='HOPE')
RBHob = ax2.bar((TEnergies*(6.242e18))[len(HoEnergies):-1], NBetaCDF[len(HoEnergies)-1:],
width=Width[len(HoEnergies):], color='m', label='HOPE and RBSPICE\nw/o RBSPICE first energy channel')
RBb = ax2.bar((Energies*(6.242e18))[1:-1], NOBetaCDF,
width=Width[len(HoEnergies):], color='b', label='RBSPICE TOFxE')
Ho = lines.Line2D([], [], color='k', marker='o', markerfacecolor='r', markeredgewidth=1, label='HOPE')
Ho3 = lines.Line2D([], [], color='k', marker='o', markerfacecolor='g', markeredgewidth=1, label='HOPE*3')
RB = lines.Line2D([], [], color='k', marker='o', markerfacecolor='b', markeredgewidth=1, label='RBSPICE TOFxE')
hands = [Ho, Ho3, RB, Hob, RBHob, RBb]
labs = ['HOPE', 'HOPE*3', 'RBSPICE TOFxE', 'HOPE',
'HOPE and RBSPICE\nw/o RBSPICE first energy channel', 'RBSPICE TOFxE']
ax.legend(hands, labs, loc=2)
ax.set_zorder(2)
ax.patch.set_visible(False)
fig.suptitle(r'H$^+ \boldsymbol{\beta}/eV$ Contribution vs Energy for '+start.strftime('%m/%d/%y, %H:%M-')+stop.strftime('%H:%M'))
ax.set_ylabel(r'Total $\boldsymbol{\beta}/eV$ Contribution for Time Period')
ax2.set_ylabel(r'Normalized Cumulative $\boldsymbol{\beta}$ Contribution for Time Period')
ax.set_xlabel('Energy (eV)')
# plt.savefig(craft+start.strftime('%Y%m%d-%H%M-')+stop.strftime('%H%M-')
# +'timeIntegrated3.png', format='png', dpi=100)
plt.show()
```
| github_jupyter |
# Reliability Diagram
```
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from os.path import join
from cal_methods import HistogramBinning, TemperatureScaling
from betacal import BetaCalibration
from sklearn.isotonic import IsotonicRegression
from sklearn.linear_model import LogisticRegression
# Imports to get "utility" package
import sys
from os import path
sys.path.append( path.dirname( path.dirname( path.abspath("utility") ) ) )
from utility.unpickle_probs import unpickle_probs
from utility.evaluation import get_bin_info, softmax
```
Script for plotting reliability diagrams.
## Load in the data
```
PATH = join('..', '..', 'logits')
files = (
'probs_resnet110_c10_logits.p', 'probs_resnet110_c100_logits.p',
'probs_densenet40_c10_logits.p', 'probs_densenet40_c100_logits.p',
'probs_resnet_wide32_c10_logits.p', 'probs_resnet_wide32_c100_logits.p',
'probs_resnet50_birds_logits.p',
'probs_resnet110_SD_c10_logits.p', 'probs_resnet110_SD_c100_logits.p',
'probs_resnet152_SD_SVHN_logits.p',
'probs_resnet152_imgnet_logits.p', 'probs_densenet161_imgnet_logits.p'
)
```
### Reliability diagrams as subgraph
```
# reliability diagram plotting for subplot case.
def rel_diagram_sub(accs, confs, ax, M = 10, name = "Reliability Diagram", xname = "", yname=""):
acc_conf = np.column_stack([accs,confs])
acc_conf.sort(axis=1)
outputs = acc_conf[:, 0]
gap = acc_conf[:, 1]
bin_size = 1/M
positions = np.arange(0+bin_size/2, 1+bin_size/2, bin_size)
# Plot gap first, so its below everything
gap_plt = ax.bar(positions, gap, width = bin_size, edgecolor = "red", color = "red", alpha = 0.3, label="Gap", linewidth=2, zorder=2)
# Next add error lines
#for i in range(M):
#plt.plot([i/M,1], [0, (M-i)/M], color = "red", alpha=0.5, zorder=1)
#Bars with outputs
output_plt = ax.bar(positions, outputs, width = bin_size, edgecolor = "black", color = "blue", label="Outputs", zorder = 3)
# Line plot with center line.
ax.set_aspect('equal')
ax.plot([0,1], [0,1], linestyle = "--")
ax.legend(handles = [gap_plt, output_plt])
ax.set_xlim(0,1)
ax.set_ylim(0,1)
ax.set_title(name, fontsize=24)
ax.set_xlabel(xname, fontsize=22, color = "black")
ax.set_ylabel(yname, fontsize=22, color = "black")
def get_pred_conf(y_probs, normalize = False):
y_preds = np.argmax(y_probs, axis=1) # Take maximum confidence as prediction
if normalize:
y_confs = np.max(y_probs, axis=1)/np.sum(y_probs, axis=1)
else:
y_confs = np.max(y_probs, axis=1) # Take only maximum confidence
return y_preds, y_confs
```
## Calibration methods for both 1-vs-rest and mutliclass approach
```
def cal_res(method, path, file, M = 15, name = "", approach = "single", m_kwargs = {}):
bin_size = 1/M
FILE_PATH = join(path, file)
(y_logits_val, y_val), (y_logits_test, y_test) = unpickle_probs(FILE_PATH)
y_probs_val = softmax(y_logits_val) # Softmax logits
y_probs_test = softmax(y_logits_test)
if approach == "single":
K = y_probs_test.shape[1]
# Go through all the classes
for k in range(K):
# Prep class labels (1 fixed true class, 0 other classes)
y_cal = np.array(y_val == k, dtype="int")[:, 0]
# Train model
model = method(**m_kwargs)
model.fit(y_probs_val[:, k], y_cal) # Get only one column with probs for given class "k"
y_probs_val[:, k] = model.predict(y_probs_val[:, k]) # Predict new values based on the fittting
y_probs_test[:, k] = model.predict(y_probs_test[:, k])
# Replace NaN with 0, as it should be close to zero # TODO is it needed?
idx_nan = np.where(np.isnan(y_probs_test))
y_probs_test[idx_nan] = 0
idx_nan = np.where(np.isnan(y_probs_val))
y_probs_val[idx_nan] = 0
y_preds_val, y_confs_val = get_pred_conf(y_probs_val, normalize = True)
y_preds_test, y_confs_test = get_pred_conf(y_probs_test, normalize = True)
else:
model = method(**m_kwargs)
model.fit(y_logits_val, y_val)
y_probs_val = model.predict(y_logits_val)
y_probs_test = model.predict(y_logits_test)
y_preds_val, y_confs_val = get_pred_conf(y_probs_val, normalize = False)
y_preds_test, y_confs_test = get_pred_conf(y_probs_test, normalize = False)
accs_val, confs_val, len_bins_val = get_bin_info(y_confs_val, y_preds_val, y_val, bin_size = bin_size)
accs_test, confs_test, len_bins_test = get_bin_info(y_confs_test, y_preds_test, y_test, bin_size = bin_size)
return (accs_test, confs_test, len_bins_test), (accs_val, confs_val, len_bins_val)
def get_uncalibrated_res(path, file, M = 15):
bin_size = 1/M
FILE_PATH = join(path, file)
(y_logits_val, y_val), (y_logits_test, y_test) = unpickle_probs(FILE_PATH)
y_probs_test = softmax(y_logits_test)
y_preds_test, y_confs_test = get_pred_conf(y_probs_test, normalize = False)
return get_bin_info(y_confs_test, y_preds_test, y_test, bin_size = bin_size)
import pickle
def gen_plots(files, plot_names = [], M = 15, val_set = False):
if val_set: # Plot Reliability diagrams for validation set
k = 1
else:
k = 0
for i, file in enumerate(files):
bin_info_uncal = get_uncalibrated_res(PATH, file, M)
accs_confs = []
accs_confs.append(cal_res(TemperatureScaling, PATH, file, M, "", "multi"))
accs_confs.append(cal_res(HistogramBinning, PATH, file, M, "", "single", {'M':M}))
accs_confs.append(cal_res(IsotonicRegression, PATH, file, M, "", "single", {'y_min':0, 'y_max':1}))
accs_confs.append(cal_res(BetaCalibration, PATH, file, M, "", "single", {'parameters':"abm"}))
with open(plot_names[i] + "_bin_info.p", "wb") as f:
pickle.dump(accs_confs, f)
plt.style.use('ggplot')
fig, ax = plt.subplots(nrows=1, ncols=5, figsize=(22.5, 4), sharex='col', sharey='row')
names = [" (Uncal)", " (Temp)", " (Histo)", " (Iso)", " (Beta)"]
# Uncalibrated
rel_diagram_sub(bin_info_uncal[0], bin_info_uncal[1], ax[0] , M = M, name = "\n".join(plot_names[i].split()) + names[0], xname="Confidence")
# Calibrated
for j in range(4):
rel_diagram_sub(accs_confs[j][k][0], accs_confs[j][k][1], ax[j+1] , M = M, name = "\n".join(plot_names[i].split()) + names[j+1], xname="Confidence")
ax[0].set_ylabel("Accuracy", color = "black")
for ax_temp in ax:
plt.setp(ax_temp.get_xticklabels(), rotation='horizontal', fontsize=18)
plt.setp(ax_temp.get_yticklabels(), fontsize=18)
plt.savefig("_".join(plot_names[i].split()) + ".pdf", format='pdf', dpi=1000, bbox_inches='tight', pad_inches=0.2)
plt.show()
gen_plots(files[:2], plot_names = ["ResNet-110(SD) CIFAR-10", "ResNet-110(SD) CIFAR-100",
#"DenseNet-40 CIFAR-10", "DenseNet-40 CIFAR-100",
#"WideNet-32 CIFAR-10", "WideNet-32 CIFAR-100",
#"ResNet-50 Birds", "ResNet-110(SD) CIFAR-10",
#"ResNet-110(SD) CIFAR-100",
#"ResNet-152(SD) SVHN",
#"ResNet-152 ImageNet", "DenseNet-161 ImageNet"
], M = 10, val_set=False)
gen_plots(files, plot_names = ["ResNet-110 CIFAR-10", "ResNet-110 CIFAR-100",
"DenseNet-40 CIFAR-10", "DenseNet-40 CIFAR-100",
"WideNet-32 CIFAR-10", "WideNet-32 CIFAR-100",
"ResNet-50 Birds",
"ResNet-110(SD) CIFAR-10", "ResNet-110(SD) CIFAR-100",
"ResNet-152(SD) SVHN",
#"ResNet-152 ImageNet", "DenseNet-161 ImageNet"
], val_set = False)
```
| github_jupyter |
## CIFAR 10
```
%matplotlib inline
%reload_ext autoreload
%autoreload 2
```
You can get the data via:
wget http://pjreddie.com/media/files/cifar.tgz
```
from resources.conv_learner import *
import pickle
PATH = "../datasets/yeast_v4/"
os.makedirs(PATH,exist_ok=True)
def get_data(sz,bs):
# load stats dictionary
with open('./norm_stats.dict','rb') as file:
stats = pickle.load(file)
return ImageClassifierData.from_paths_and_stats(stats,sz,PATH, val_name='test',bs=bs)
### the eventual sub-function of ImageClassifierData (read_dirs) expects subdirectories for each class:
### e.g. all "test/cat.png" images should be in a "cat" folder.
```
### Look at data
```
batch_size = 4
data = get_data(200, batch_size)
x,y=next(iter(data.trn_dl))
print(y)
imdenorm = data.trn_ds.denorm(x, 1) #denorm function called has a rollaxis() hence indexing changes.
imdenorm.shape
plt.imshow(imdenorm[0,:,:,1])
```
## Fully connected model
```
data = get_data(200,bs)
lr=1e-5
```
From [this notebook](https://github.com/KeremTurgutlu/deeplearning/blob/master/Exploring%20Optimizers.ipynb) by our student Kerem Turgutlu:
```
class SimpleNet(nn.Module):
def __init__(self, layers):
super().__init__()
self.layers = nn.ModuleList([
nn.Linear(layers[i], layers[i + 1]) for i in range(len(layers) - 1)])
def forward(self, x):
x = x.view(x.size(0), -1)
for l in self.layers:
l_x = l(x)
x = F.relu(l_x)
return F.log_softmax(l_x, dim=-1)
learn = ConvLearner.from_model_data(SimpleNet([200*200*2, 40,2]), data) #(!) change channel-number & classes accordingly
learn, [o.numel() for o in learn.model.parameters()]
learn.summary()
learn.lr_find2()
learn.sched.plot()
%time learn.fit(lr, 2)
%time learn.fit(lr, 200, cycle_len=1)
learn.sched.plot()
```
## CNN
```
class ConvNet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.layers = nn.ModuleList([
nn.Conv2d(layers[i], layers[i + 1], kernel_size=3, stride=2)
for i in range(len(layers) - 1)])
self.pool = nn.AdaptiveMaxPool2d(1)
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
for l in self.layers: x = F.relu(l(x))
x = self.pool(x)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvNet([2, 20, 40, 80], 2), data)
### .from_model_data triggers basicmodel() which forwards to a pytorch's .cuda function to run model on GPU. Hence
### an explicit model.cuda is not necessary here.
learn.summary() ### learner.summary is hardcording the number of channels = 3
learn.lr_find(end_lr=100)
learn.sched.plot()
%time learn.fit(1e-5, 2)
%time learn.fit(1e-5, 4, cycle_len=1)
```
## Refactored
```
class ConvLayer(nn.Module):
def __init__(self, ni, nf):
super().__init__()
self.conv = nn.Conv2d(ni, nf, kernel_size=3, stride=2, padding=1)
def forward(self, x): return F.relu(self.conv(x))
class ConvNet2(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.layers = nn.ModuleList([ConvLayer(layers[i], layers[i + 1])
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
for l in self.layers: x = l(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvNet2([3, 20, 40, 80], 2), data)
learn.summary()
%time learn.fit(1e-1, 2)
%time learn.fit(1e-1, 2, cycle_len=1)
```
## BatchNorm
```
class BnLayer(nn.Module):
def __init__(self, ni, nf, stride=2, kernel_size=3):
super().__init__()
self.conv = nn.Conv2d(ni, nf, kernel_size=kernel_size, stride=stride,
bias=False, padding=1)
self.a = nn.Parameter(torch.zeros(nf,1,1))
self.m = nn.Parameter(torch.ones(nf,1,1))
def forward(self, x):
x = F.relu(self.conv(x))
x_chan = x.transpose(0,1).contiguous().view(x.size(1), -1)
if self.training:
self.means = x_chan.mean(1)[:,None,None]
self.stds = x_chan.std (1)[:,None,None]
return (x-self.means) / self.stds *self.m + self.a
class ConvBnNet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(2, 10, kernel_size=5, stride=1, padding=2) #(!) changed 3 to 2
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i + 1])
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l in self.layers: x = l(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvBnNet([10, 20, 40, 80, 160], 2), data)
learn.summary()
%time learn.fit(3e-2, 2)
%time learn.fit(1e-4, 20, cycle_len=1)
learn.save('tmp_batchNorm_aug')
```
## Deep BatchNorm
```
class ConvBnNet2(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([BnLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l,l2 in zip(self.layers, self.layers2):
x = l(x)
x = l2(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvBnNet2([10, 20, 40, 80, 160], 10), data)
%time learn.fit(1e-2, 2)
%time learn.fit(1e-2, 2, cycle_len=1)
```
## Resnet
```
class ResnetLayer(BnLayer):
def forward(self, x): return x + super().forward(x)
class Resnet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.layers3 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l,l2,l3 in zip(self.layers, self.layers2, self.layers3):
x = l3(l2(l(x)))
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(Resnet([10, 20, 40, 80, 160], 2), data)
wd=1e-5
%time learn.fit(1e-5, 2, wds=wd, cycle_len=1, use_clr=(20,8, 0.95, 0.85))
%time learn.fit(1e-5, 3, cycle_len=1, cycle_mult=2, wds=wd)
%time learn.fit(1e-5, 20, cycle_len=4, wds=wd)
```
## Resnet 2
```
class Resnet2(nn.Module):
def __init__(self, layers, c, p=0.5):
super().__init__()
self.conv1 = BnLayer(3, 16, stride=1, kernel_size=7)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.layers3 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
self.drop = nn.Dropout(p)
def forward(self, x):
x = self.conv1(x)
for l,l2,l3 in zip(self.layers, self.layers2, self.layers3):
x = l3(l2(l(x)))
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
x = self.drop(x)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(Resnet2([16, 32, 64, 128, 256], 10, 0.2), data)
wd=1e-6
%time learn.fit(1e-2, 2, wds=wd)
%time learn.fit(1e-2, 3, cycle_len=1, cycle_mult=2, wds=wd)
%time learn.fit(1e-2, 8, wds=wd, cycle_len=4, use_clr=(20,8, 0.95, 0.85))
learn.save('tmp4_clr')
log_preds,y = learn.TTA()
preds = np.mean(np.exp(log_preds),0)
metrics.log_loss(y,preds), accuracy(preds,y)
metrics.log_loss(y,preds)
```
### End
| github_jupyter |
# Example: CanvasXpress density Chart No. 1
This example page demonstrates how to, using the Python package, create a chart that matches the CanvasXpress online example located at:
https://www.canvasxpress.org/examples/density-1.html
This example is generated using the reproducible JSON obtained from the above page and the `canvasxpress.util.generator.generate_canvasxpress_code_from_json_file()` function.
Everything required for the chart to render is included in the code below. Simply run the code block.
```
from canvasxpress.canvas import CanvasXpress
from canvasxpress.js.collection import CXEvents
from canvasxpress.render.jupyter import CXNoteBook
cx = CanvasXpress(
render_to="density1",
data={
"y": {
"smps": [
"weight"
],
"data": [
[
49
],
[
56
],
[
60
],
[
43
],
[
57
],
[
58
],
[
52
],
[
52
],
[
52
],
[
51
],
[
53
],
[
50
],
[
51
],
[
55
],
[
60
],
[
54
],
[
52
],
[
50
],
[
51
],
[
67
],
[
56
],
[
53
],
[
53
],
[
57
],
[
52
],
[
48
],
[
58
],
[
50
],
[
55
],
[
50
],
[
61
],
[
53
],
[
51
],
[
52
],
[
47
],
[
49
],
[
44
],
[
48
],
[
54
],
[
53
],
[
62
],
[
50
],
[
51
],
[
54
],
[
50
],
[
50
],
[
49
],
[
49
],
[
52
],
[
53
],
[
46
],
[
52
],
[
49
],
[
50
],
[
54
],
[
58
],
[
63
],
[
51
],
[
63
],
[
49
],
[
58
],
[
68
],
[
55
],
[
52
],
[
55
],
[
64
],
[
49
],
[
62
],
[
62
],
[
57
],
[
55
],
[
53
],
[
53
],
[
58
],
[
65
],
[
54
],
[
48
],
[
51
],
[
56
],
[
53
],
[
54
],
[
54
],
[
48
],
[
54
],
[
59
],
[
58
],
[
58
],
[
53
],
[
54
],
[
49
],
[
55
],
[
56
],
[
64
],
[
60
],
[
53
],
[
57
],
[
49
],
[
59
],
[
60
],
[
66
],
[
57
],
[
53
],
[
55
],
[
52
],
[
51
],
[
56
],
[
51
],
[
56
],
[
57
],
[
55
],
[
54
],
[
52
],
[
49
],
[
59
],
[
55
],
[
59
],
[
49
],
[
56
],
[
58
],
[
55
],
[
54
],
[
51
],
[
65
],
[
59
],
[
64
],
[
55
],
[
52
],
[
47
],
[
52
],
[
56
],
[
60
],
[
56
],
[
49
],
[
58
],
[
47
],
[
53
],
[
53
],
[
45
],
[
60
],
[
52
],
[
53
],
[
62
],
[
58
],
[
54
],
[
58
],
[
57
],
[
63
],
[
56
],
[
58
],
[
57
],
[
53
],
[
55
],
[
63
],
[
51
],
[
56
],
[
62
],
[
54
],
[
50
],
[
51
],
[
53
],
[
51
],
[
54
],
[
53
],
[
54
],
[
57
],
[
58
],
[
63
],
[
55
],
[
53
],
[
62
],
[
64
],
[
55
],
[
53
],
[
46
],
[
62
],
[
51
],
[
49
],
[
70
],
[
56
],
[
55
],
[
41
],
[
55
],
[
60
],
[
57
],
[
60
],
[
65
],
[
61
],
[
52
],
[
59
],
[
54
],
[
52
],
[
41
],
[
51
],
[
57
],
[
66
],
[
58
],
[
58
],
[
50
],
[
56
],
[
45
],
[
67
],
[
68
],
[
66
],
[
69
],
[
67
],
[
69
],
[
74
],
[
71
],
[
65
],
[
59
],
[
67
],
[
63
],
[
72
],
[
57
],
[
63
],
[
67
],
[
64
],
[
62
],
[
63
],
[
68
],
[
69
],
[
68
],
[
76
],
[
71
],
[
66
],
[
62
],
[
80
],
[
68
],
[
62
],
[
66
],
[
63
],
[
64
],
[
60
],
[
66
],
[
67
],
[
60
],
[
49
],
[
64
],
[
65
],
[
68
],
[
65
],
[
67
],
[
60
],
[
69
],
[
69
],
[
66
],
[
72
],
[
67
],
[
66
],
[
66
],
[
67
],
[
70
],
[
67
],
[
68
],
[
59
],
[
63
],
[
72
],
[
59
],
[
66
],
[
67
],
[
70
],
[
63
],
[
66
],
[
56
],
[
67
],
[
62
],
[
64
],
[
59
],
[
67
],
[
68
],
[
63
],
[
74
],
[
68
],
[
70
],
[
75
],
[
62
],
[
69
],
[
70
],
[
65
],
[
67
],
[
60
],
[
67
],
[
61
],
[
69
],
[
61
],
[
67
],
[
61
],
[
64
],
[
57
],
[
66
],
[
70
],
[
66
],
[
56
],
[
62
],
[
73
],
[
74
],
[
59
],
[
63
],
[
67
],
[
67
],
[
62
],
[
60
],
[
64
],
[
70
],
[
65
],
[
62
],
[
62
],
[
73
],
[
63
],
[
69
],
[
72
],
[
67
],
[
63
],
[
65
],
[
63
],
[
71
],
[
64
],
[
73
],
[
62
],
[
62
],
[
66
],
[
65
],
[
62
],
[
57
],
[
65
],
[
61
],
[
70
],
[
60
],
[
71
],
[
62
],
[
66
],
[
69
],
[
62
],
[
68
],
[
65
],
[
59
],
[
64
],
[
73
],
[
64
],
[
61
],
[
65
],
[
67
],
[
70
],
[
71
],
[
66
],
[
71
],
[
61
],
[
53
],
[
63
],
[
62
],
[
53
],
[
68
],
[
61
],
[
64
],
[
57
],
[
68
],
[
74
],
[
61
],
[
64
],
[
75
],
[
70
],
[
75
],
[
65
],
[
64
],
[
62
],
[
72
],
[
59
],
[
67
],
[
65
],
[
76
],
[
62
],
[
57
],
[
66
],
[
65
],
[
61
],
[
66
],
[
64
],
[
62
],
[
68
],
[
63
],
[
56
],
[
52
],
[
62
],
[
72
],
[
69
],
[
71
],
[
70
],
[
67
],
[
57
],
[
66
],
[
73
],
[
48
],
[
61
],
[
71
],
[
68
],
[
69
],
[
67
],
[
68
],
[
65
],
[
60
]
],
"vars": [
"var1",
"var2",
"var3",
"var4",
"var5",
"var6",
"var7",
"var8",
"var9",
"var10",
"var11",
"var12",
"var13",
"var14",
"var15",
"var16",
"var17",
"var18",
"var19",
"var20",
"var21",
"var22",
"var23",
"var24",
"var25",
"var26",
"var27",
"var28",
"var29",
"var30",
"var31",
"var32",
"var33",
"var34",
"var35",
"var36",
"var37",
"var38",
"var39",
"var40",
"var41",
"var42",
"var43",
"var44",
"var45",
"var46",
"var47",
"var48",
"var49",
"var50",
"var51",
"var52",
"var53",
"var54",
"var55",
"var56",
"var57",
"var58",
"var59",
"var60",
"var61",
"var62",
"var63",
"var64",
"var65",
"var66",
"var67",
"var68",
"var69",
"var70",
"var71",
"var72",
"var73",
"var74",
"var75",
"var76",
"var77",
"var78",
"var79",
"var80",
"var81",
"var82",
"var83",
"var84",
"var85",
"var86",
"var87",
"var88",
"var89",
"var90",
"var91",
"var92",
"var93",
"var94",
"var95",
"var96",
"var97",
"var98",
"var99",
"var100",
"var101",
"var102",
"var103",
"var104",
"var105",
"var106",
"var107",
"var108",
"var109",
"var110",
"var111",
"var112",
"var113",
"var114",
"var115",
"var116",
"var117",
"var118",
"var119",
"var120",
"var121",
"var122",
"var123",
"var124",
"var125",
"var126",
"var127",
"var128",
"var129",
"var130",
"var131",
"var132",
"var133",
"var134",
"var135",
"var136",
"var137",
"var138",
"var139",
"var140",
"var141",
"var142",
"var143",
"var144",
"var145",
"var146",
"var147",
"var148",
"var149",
"var150",
"var151",
"var152",
"var153",
"var154",
"var155",
"var156",
"var157",
"var158",
"var159",
"var160",
"var161",
"var162",
"var163",
"var164",
"var165",
"var166",
"var167",
"var168",
"var169",
"var170",
"var171",
"var172",
"var173",
"var174",
"var175",
"var176",
"var177",
"var178",
"var179",
"var180",
"var181",
"var182",
"var183",
"var184",
"var185",
"var186",
"var187",
"var188",
"var189",
"var190",
"var191",
"var192",
"var193",
"var194",
"var195",
"var196",
"var197",
"var198",
"var199",
"var200",
"var201",
"var202",
"var203",
"var204",
"var205",
"var206",
"var207",
"var208",
"var209",
"var210",
"var211",
"var212",
"var213",
"var214",
"var215",
"var216",
"var217",
"var218",
"var219",
"var220",
"var221",
"var222",
"var223",
"var224",
"var225",
"var226",
"var227",
"var228",
"var229",
"var230",
"var231",
"var232",
"var233",
"var234",
"var235",
"var236",
"var237",
"var238",
"var239",
"var240",
"var241",
"var242",
"var243",
"var244",
"var245",
"var246",
"var247",
"var248",
"var249",
"var250",
"var251",
"var252",
"var253",
"var254",
"var255",
"var256",
"var257",
"var258",
"var259",
"var260",
"var261",
"var262",
"var263",
"var264",
"var265",
"var266",
"var267",
"var268",
"var269",
"var270",
"var271",
"var272",
"var273",
"var274",
"var275",
"var276",
"var277",
"var278",
"var279",
"var280",
"var281",
"var282",
"var283",
"var284",
"var285",
"var286",
"var287",
"var288",
"var289",
"var290",
"var291",
"var292",
"var293",
"var294",
"var295",
"var296",
"var297",
"var298",
"var299",
"var300",
"var301",
"var302",
"var303",
"var304",
"var305",
"var306",
"var307",
"var308",
"var309",
"var310",
"var311",
"var312",
"var313",
"var314",
"var315",
"var316",
"var317",
"var318",
"var319",
"var320",
"var321",
"var322",
"var323",
"var324",
"var325",
"var326",
"var327",
"var328",
"var329",
"var330",
"var331",
"var332",
"var333",
"var334",
"var335",
"var336",
"var337",
"var338",
"var339",
"var340",
"var341",
"var342",
"var343",
"var344",
"var345",
"var346",
"var347",
"var348",
"var349",
"var350",
"var351",
"var352",
"var353",
"var354",
"var355",
"var356",
"var357",
"var358",
"var359",
"var360",
"var361",
"var362",
"var363",
"var364",
"var365",
"var366",
"var367",
"var368",
"var369",
"var370",
"var371",
"var372",
"var373",
"var374",
"var375",
"var376",
"var377",
"var378",
"var379",
"var380",
"var381",
"var382",
"var383",
"var384",
"var385",
"var386",
"var387",
"var388",
"var389",
"var390",
"var391",
"var392",
"var393",
"var394",
"var395",
"var396",
"var397",
"var398",
"var399",
"var400"
]
},
"z": {
"sex": [
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"F",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M",
"M"
]
}
},
config={
"graphType": "Scatter2D",
"hideHistogram": True,
"showHistogramDensity": True,
"theme": "CanvasXpress"
},
width=613,
height=613,
events=CXEvents(),
after_render=[
[
"createHistogram",
[
False,
None,
None
]
]
],
other_init_params={
"version": 35,
"events": False,
"info": False,
"afterRenderInit": False,
"noValidate": True
}
)
display = CXNoteBook(cx)
display.render(output_file="density_1.html")
```
| github_jupyter |
# SUMO DEMAND GENERATION
## Overview
Author: Joseph Severino
Contact: joseph.severino@nrel.gov
This notebook is a configuration notebook for building demand in XML format. SUMO uses XML files as input
files for simulation purposes. The beginning part of this notebook is clean and process the data for
exploration so that a user understands the data they working with. The analysis is basic VDA to help guide
which days to investigate.
## Inputs
This notebook only takes the operational model input (i.e. "athena_sumo_v1.csv"). This data was generated based on flight arrivals and departures along with the predictive analysis the operational modelling team worked on.
## Outputs
There are two XML files that this notebook generates: a Trips file and an additional file. The trips file conatins all the entry, stopping and exit routes for all the patron vehicles at DFW.
```
import xml.etree.ElementTree as ET
from xml.etree.ElementTree import Element
from lxml import etree
from copy import copy
import os
import inspect
from xml.dom import minidom
import pandas as pd
import numpy as np
import random
import matplotlib.pyplot as plt
import seaborn as sns
import DFW_gen_flow as gf
this_folder = os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe())))
this_folder
```
# Generating Demand from flight data model
```
demand = pd.read_csv('../Example_Files/athena_sumo_v1.csv')
demand.head()
gf.addDayOfWeek(demand)
demand['total_pass'] = demand[['parking','pass','A','B','C','D','E']].sum(axis=1) # aggregating across the terminals
demand.head()
gf.extractDate(demand)
sorted_top10, sorted_bot10, median_days, sort_all = gf.buildTopBottomMedianDays(demand)
```
# Top 10 highest Volume days - Aggregated over the day
```
sorted_top10[['parking','pass','A','B','C','D','E','total_pass']]
```
# Bottom 10 Volume Days
```
sorted_bot10[['parking','pass','A','B','C','D','E','total_pass']]
```
# Middle 10 days
```
median_days[['parking','pass','A','B','C','D','E','total_pass']]
```
## Distributions of Number of Vehicles/Day
```
plt.figure(figsize=(15,4))
plt.subplot(1, 2, 1)
plt.hist(sort_all['total_pass'],bins=15,color='#074659')
plt.title("Distribution of Volume")
plt.xlabel("Number of Vehicles in Day")
plt.ylabel("Frequency (number of days)")
plt.subplot(1, 2, 2)
sort_all.reset_index(inplace=True, drop=True)
sort_all['percentile'] = np.round(100*sort_all.index.values/sort_all.shape[0],3)
plt.plot(sort_all['percentile'],sort_all['total_pass'],color='#074659')
plt.fill_between(sort_all['percentile'],sort_all['total_pass'],color='#074659')
plt.title("Cumulative Distribution Total Volume")
plt.xlabel("Percent of Days")
plt.ylabel("Number Of Vehicles/Day")
plt.show()
day_df = demand.groupby("date").sum()
```
# Showing all data
Below you can see how volume varies through time over the past year. Notice one day that is exceptionally large compared to all the other days ("Mother's Day)
```
plt.figure(figsize=(20,8))
sns.set_style("whitegrid")
ax = day_df['total_pass'].plot(color="#074659")
max_value = np.max(day_df['total_pass'])
string_max = day_df['total_pass'].idxmax() + ": Mother's Day"
index_max = np.argmax(list(day_df['total_pass'])) +1
plt.text(index_max, max_value, string_max, color="#aa2200")
min_value = np.min(day_df['total_pass'])
string_min = day_df['total_pass'].idxmin()
index_min = np.argmin(list(day_df['total_pass'])) +1
plt.text(index_min, min_value, string_min, color="#aa2200")
plt.legend()
plt.title("Vehicle Volume over the Year")
plt.xlabel("Time")
plt.ylabel("Number of Vehicles")
plt.show()
```
# Looking for highest and lowest volume by Day of the week
* Monday, Thursday and Friday are highest days
* Saturday and Sunday are the lowest days followed by Tuesday Wednesday
```
col = 'day_of_week'
fig, [ax,ax2] = plt.subplots(nrows=2,ncols=1,figsize=(14,8),sharey=True)
demand[['parking','pass','A','B','C','D','E','day_of_week']].groupby(col).mean().plot(kind="bar",ax=ax,stacked=True)
demand[['total_pass','day_of_week']].groupby(col).mean().plot(kind="bar",ax=ax2, color="#074659")
ax.set_ylabel("Number of Vehicles")
ax2.set_ylabel("Number of Vehicles")
```
* May, June July are highest months
* January, Febuary, December are lowest months
```
col = 'month'
fig, [ax,ax2] = plt.subplots(nrows=2,ncols=1,figsize=(14,8),sharey=True)
demand[['parking','pass','A','B','C','D','E','month']].groupby(col).mean().plot(kind="bar",ax=ax,stacked=True)
demand[['total_pass','month']].groupby(col).mean().plot(kind="bar",ax=ax2, color="#074659")
ax.set_ylabel("Number of Vehicles")
ax2.set_ylabel("Number of Vehicles")
```
# Sampling Day by Day Of Week and Month Based on Quantile/Mean
```
# Below is
d = "Monday"
m = "June"
df = gf.top_day_month(demand,d,m,.95)
df.head(5)
d = "Monday"
m = "June"
df = gf.avg_day_month(demand,d,m)
df.head()
```
# Pick your day of Interest
```
day, date = gf.pick_day(demand,'2017-12-31')
day.head()
```
* May 13th, 2018 was the highest volume day from the data we recieved
* Choose another day of interest from above to study or generate demand
# Plotting the Volume by terminal over time
* A and C tend ot have the most volume through the day
* E always seems to have far less than all the rest. May need to look into validating this?
* B and D are similar in volume
* Parking tends to always have it's largeest spike in AM around 4:30 am and tends to diminish through the day.
* Parking also has second peak around 12:30pm
* Pass through traffic typically mimics am and pm peaks that are common is traffic. Most likely occers on high volume days were regional traffic uses DFW as shortcut during am and pm peak hours
```
gf.plot_vol_by_type(day)
day['terminal_tot'] = day[['A','B','C','D','E']].sum(axis=1)
day.head()
gf.plot_tot_vs_term(day)
```
## Pie Chart to show statistics about this days distribution of volume
```
gf.plot_pie_all(day)
```
## Number of vehicles on this day:
```
day['total_pass'].sum()
terminal_dict = {
"A":['A_top_1','A_top_2','A_top_3','A_bot_1','A_bot_2','A_bot_3'],
"B":['B_top_1','B_top_2','B_top_3','B_bot_1','B_bot_2','B_bot_3'],
"C":['C_top_1','C_top_2','C_top_3','C_bot_1','C_bot_2','C_bot_3'],
"D":['D_depart_1','D_depart_2','D_arrive_1','D_arrive_2','D_service'],
"E":['E_top_1','E_top_2','E_top_3','E_bot_1','E_bot_2','E_bot_3'],
"parking":['park_south_term_1','park_south_term_2','park_south_term_3','park_south_term_4',
'park_south_emp_1','park_south_emp_2','park_south_emp_3','park_north_emp_1',
'park_north_emp_2','park_north_emp_3','park_north_emp_4','park_north_emp_5',
'park_north_emp_6'],
"pass":"pass_through"
}
stop_dict = {
"A":['A_top_1','A_top_2','A_top_3','A_bot_1','A_bot_2','A_bot_3'],
"B":['B_top_1','B_top_2','B_top_3','B_bot_1','B_bot_2','B_bot_3'],
"C":['C_top_1','C_top_2','C_top_3','C_bot_1','C_bot_2','C_bot_3'],
"D":['D_depart_1','D_depart_2','D_arrive_1','D_arrive_2','D_service'],
"E":['E_top_1','E_top_2','E_top_3','E_bot_1','E_bot_2','E_bot_3'],
"others":['park_south_term_1','park_south_term_2','park_south_term_3','park_south_term_4',
'park_south_emp_1','park_south_emp_2','park_south_emp_3','park_north_emp_1',
'park_north_emp_2','park_north_emp_3','park_north_emp_4','park_north_emp_5',
'park_north_emp_6']
}
```
# GENERATE XML DEMAND FILE FOR SUMO
```
gf.model_to_sumo(this_folder,day,date,False)
```
# GENERATE ADDITIONAL FILES FOR SUMO
```
gf.create_additional_file(this_folder,stop_dict,date)
```
# You now have all the inputs to run SUMO for DFW
| github_jupyter |
# scikit-learn for NLP -- Part 1 Introductory Tutorial
[scikit-learning](http://scikit-learn.org/stable/index.html) is an open-sourced simple and efficient tools for data mining, data analysis and machine learning in Python. It is built on NumPy, SciPy and matplotlib. There are built-in classification, regression, and clustering models, as well as useful features like dimensionality reduction, evaluation and preprocessing.
These tutorials are specifically tailored for NLP, i.e. working with text data. Part one covers the following topics: loading data, simple preprocessing, training (supervised, semi-supervised, and unsupervised), and evaluation. Part two focuses on feature engineering and covers more advanced topics like feature extraction, building a pipeline, creating custom transformers, feature union, dimensionality reduction, etc.
For this tutorial, we will use the [20 Newsgroups data set](http://qwone.com/~jason/20Newsgroups/) and perform topic classification. For the sake of time, I converted all the data into a CSV file.
Note: apparently Jupyter disables spell-checker (or I just don't know how to enable it), so I'm only partially responsible for the typos in this tutorial.
### 1. Loading Dataset
For the sake of convenience, we will use pandas to read CSV file. (You may do so with numpy as well; there is a `loadtext()` function, but you might encounter encoding issues when using it. Before you begin, the dataset in the repo is a tarball, so in the terminal run ``tar xfz 20news-18828.csv.tar.gz`` to uncompress it.
```
import pandas as pd
dataset = pd.read_csv('20news-18828.csv', header=None, delimiter=',', names=['label', 'text'])
```
Sanity check on the dataset.
```
print("There are 20 categories: %s" % (len(dataset.label.unique()) == 20))
print("There are 18828 records: %s" % (len(dataset) == 18828))
```
Now we need to split it to train set and test set. To do so, we can use the `train_test_split()` function. In scikit-learn's convention, X indicates data (yeah, uppercase X), and y indicates truths (and yeah, lowercase y). I mean, it doesn't really matter, as long as you remember what is what.
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(dataset.text, dataset.label, train_size=0.8)
```
### 2. A Simple Example
Before going too much into preprocessing, feature extraction and other more complicated tasks, we will do a relatively simple but complete example. In this example, we will use bag-of-words as features, and Naive Bayes as classifier to establish our baseline.
There are some built-in vectorizers, `CountVectorizer` and `TfidfVectorizer` that we can use to vectorizer our raw data and perform preprocessing and feature exctration on it. First, we will experiment with `CountVectorizer` which basically makes a token/ngram a feature and stores its count in the corresponding feature space. The `fit_transform()` function is the combination of `fit()` and `transform()`, and it's a more efficient implementation. `fit()` indexes the vocabulary/features, and `transform()` transforms the dataset into a matrix.
```
from sklearn.feature_extraction.text import CountVectorizer
# initialize a CountVectorizer
cv = CountVectorizer()
# fit the raw data into the vectorizer and tranform it into a series of arrays
X_train_counts = cv.fit_transform(X_train)
X_train_counts.shape
```
Similar thing needs to be done for the test set, but we only need to use the `transform()` function to transform the test data into a matrix.
```
X_test_counts = cv.transform(X_test)
X_test_counts.shape
```
Then, we fit our features and labels into a Naive Bayes classifier, which basically trains a model (if you fit the data more than once, it overwrites the parameters the model learns previously). After training, we can use it to perform prediction.
```
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB().fit(X_train_counts, y_train)
predicted = clf.predict(X_test_counts)
# sample some of the predictions against the ground truths
for prediction, truth in zip(predicted[:10], y_test[:10]):
print(prediction, truth)
```
Let's do some legit evaluation. The `classification_report()` function gives you precison, recall and f1 scores for each label, and their average. If you want to calculate overall macro-averaged, micro-averaged or weighted performance, you can use the `precision_recall_fscore_support`. Finally, the `confusion_matrix()` can show you which labels are confusing to the model, but unfortunately, it does not include the labels. I think that's why they call it a confusion matrix.
```
from sklearn import metrics
print(metrics.classification_report(y_test, predicted, labels=dataset.label.unique()))
p, r, f1, _ = metrics.precision_recall_fscore_support(y_test, predicted, labels=dataset.label.unique(), average='micro')
print("Micro-averaged Performance:\nPrecision: {0}, Recall: {1}, F1: {2}".format(p, r, f1))
print(metrics.confusion_matrix(y_test, predicted, labels=dataset.label.unique()))
```
### 3. Preprocessing & Feature Extraction
One may ask, "how do I remove stop words, tokenize the texts differently, or use bigrams/trigrams as features?"
The answer is you can do all that with a [`CountVectorizer`](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) object, i.e. you can pass various arguments to the constructor.
Here are some of them: `ngram_range` takes in a tuple (n_min, n_max). For example, `(2,2)` means only use bigrams, and `(1,3)` means use unigrams, bigrams, and trigrams. `stop_words` takes in a list of stopwords that you'd like to remove. If you want to use default stopword list in scikit-learn, pass in the string `'english'`. `tokenizer` is a function that takes in a string and returns a string, inside that function, you can define how to tokenize your text. By default, scikit-learn tokenization pattern is `u'(?u)\b\w\w+\b'`. Finally, `preprocessor` takes in a function of which the argument is a string and the output is a string. You can use it to perform more customized preprocessing. For more detail, please checkout the documentation for `CountVectorizer` or `TfidfVectorizer`.
Let's start with defining a preprocessor to normalize all the numeric values, i.e. replacing numbers with the string `NUM`. Then, we construct a new `CountVectorizer`, and then use unigrams, bigrams, and trigrams as features, and remove stop words.
```
import re
def normalize_numbers(s):
return re.sub(r'\b\d+\b', 'NUM', s)
cv = CountVectorizer(preprocessor=normalize_numbers, ngram_range=(1,3), stop_words='english')
```
Let's fit and transform the train data and transform the test data. The speed of preprocessing and feature extraction depends on the running time of each step. For example, the running time of stopword removal is O(N * M), where N is the vocabulary size of the document, and M is the stopword list size.
```
# fit the raw data into the vectorizer and tranform it into a series of arrays
X_train_counts = cv.fit_transform(X_train)
X_test_counts = cv.transform(X_test)
```
Let's use the Naive Bayes classifier to train a new model and see if it works better. From the last section with out preprocessing or feature engineering, our precison, recall and F1 are in the mid 80s, but now we got 90 for each score.
```
clf = MultinomialNB().fit(X_train_counts, y_train)
predicted = clf.predict(X_test_counts)
print(metrics.classification_report(y_test, predicted, labels=dataset.label.unique()))
```
Do you remember there are other vecotrizers that you can use? Walla, one of them is `TfidfVecotrizer`. LOL, what is tf-idf? It's on [Wikipedia](https://en.wikipedia.org/wiki/Tf%E2%80%93idf). It basically reflects how important a word/phrase is to a document in a corpus. The constructor of `TfidfVectorizer` takes in the same parameters as that of `CountVectorizer`, so you can perfrom the same preprocessing/feature extraction. Try to run the following block of code and see if using tf-idf will help improve the performance. There are some other parameters in the constructor that you can tweak when initializing the object, and they could affect the performance as well.
```
from sklearn.feature_extraction.text import TfidfVectorizer
tv = TfidfVectorizer(preprocessor=normalize_numbers, ngram_range=(1,3), stop_words='english')
X_train_tf = tv.fit_transform(X_train)
X_test_tf = tv.transform(X_test)
clf2 = MultinomialNB().fit(X_train_tf, y_train)
predicted = clf2.predict(X_test_tf)
print(metrics.classification_report(y_test, predicted, labels=dataset.label.unique()))
```
Alternatively, if you like typing a longer block of code, you can use the `TfidfTransformer` to transform a word count matrix created by `CountVectorizer` into a tf-idf matrix.
```
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
X_test_tfidf = tfidf_transformer.transform(X_test_counts)
```
### 4. Model Selection
Depending on the size of your data and the nature your task, some classifiers might perform better than others. These days, Maximum Entropy is a very popular classifier for many machine learning tasks, and every often, it is used to establish the baseline for a task. So let's try the [logistic regression classifier](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html) in scikit-learn (Maxent and logistic regression are virtually the same thing). Please note that this classifier is a lot slower than Naive Bayes, because of [math and whatever](https://en.wikipedia.org/wiki/Multinomial_logistic_regression) (JK, Maxent updates the weights for each feature for some iterations, whereas Naive Bayes only calculates the probabilities of each feature at once).
```
from sklearn.linear_model import LogisticRegression
# for the sake of speed, we will just use all the default value of the constructor
cv = CountVectorizer()
X_train_counts = cv.fit_transform(X_train)
X_test_counts = cv.transform(X_test)
clf = LogisticRegression(solver='liblinear', max_iter=500, n_jobs=4)
clf.fit(X_train_counts, y_train)
predicted = clf.predict(X_test_counts)
print(metrics.classification_report(y_test, predicted, labels=dataset.label.unique()))
```
There are many other supervised, semi-supervised and unsupervised algorithms, and many of them work almost the same: (1) initialize a classifier, (2) fit the feature matrix and labels of the train set, and (3) pass in the feature matrix of test set to perform prediction.
The following is an example of semi-supervised algorithm, label spreading, which uses both labeled and unlabeled data to train a classifier. It is especially nice when you don't have too much annotated data.
```
from sklearn.semi_supervised import LabelSpreading
from sklearn import preprocessing
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn import metrics
label_prop_model = LabelSpreading()
# randomly select some of the training data and use them as unlabeled data
random_unlabeled_points = np.where(np.random.randint(0, 2, size=len(y_train)))
labels = np.copy(y_train)
# convert all labels into indecies, since we'll assign -1 to the unlabeled dataset
le = preprocessing.LabelEncoder()
le.fit(labels)
# transforms labels to numeric values
labels = le.transform(labels)
labels[random_unlabeled_points] = -1
labels_test = le.transform(y_test)
# here we will need to limit the features space, because we are going to convert the sparse
# matrices into dense ones required by the implementation of LabelSpreading.
# Using dense matrices in general slows down training and takes up a lot of memory; therefore,
# in most cases, it is not recommended.
cv = CountVectorizer(max_features=100)
X_train_counts = cv.fit_transform(X_train)
X_test_counts = cv.transform(X_test)
# toarray() here is to convert a sparse matrix into a dense matrix, as the latter is required
# by the algorithm.
label_prop_model.fit(X_train_counts.toarray(), labels)
predicted = label_prop_model.predict(X_test_counts.toarray())
# For this experiment, we are only using 100 most frequent words as features in the train set,
# so the test results will be lousy.
p, r, f1, _ = metrics.precision_recall_fscore_support(labels_test, predicted, average='macro')
print("Macro-averaged Performance:\nPrecision: {0}, Recall: {1}, F1: {2}".format(p, r, f1))
```
Finally, we are going to experiment a clustering (unsupervised) aglorithm, K-means clustering. Generally speaking, it's one of the fastest and popular clustering algorithms.
```
from sklearn.cluster import KMeans
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn import metrics
from sklearn import preprocessing
X_train, X_test, y_train, y_test = train_test_split(dataset.text, dataset.label, train_size=0.8)
# I reduced the number of features here just to make training a little faster
cv = CountVectorizer(max_features=200)
X_train_counts = cv.fit_transform(X_train)
X_test_counts = cv.transform(X_test)
kmeans = KMeans(n_clusters=20, random_state=0)
kmeans.fit(X_train_counts)
# see the cluster label for each instance
print(kmeans.labels_)
print(kmeans.predict(X_test_counts))
le = preprocessing.LabelEncoder()
le.fit(y_train)
# you can also pass in the labels, if you have some annotated data
kmeans.fit(X_train_counts, le.transform(y_train))
predicted = kmeans.predict(X_test_counts)
p, r, f1, _ = metrics.precision_recall_fscore_support(le.transform(y_test), predicted, average='macro')
print("Macro-averaged Performance:\nPrecision: {0}, Recall: {1}, F1: {2}".format(p, r, f1))
```
### 5. More on Evaluation
In some examples above, we have talked about the ways to calculate precision, recall and F1; therefore, we are not going to repeat that for this section. Here, we are going to conclude this introductory tutorial with [cross-validation](https://www.youtube.com/watch?v=TIgfjmp-4BA). There are many strategies for cross validation, and scikit-learn has [a rich selection for them](http://scikit-learn.org/stable/modules/cross_validation.html); for the purpose of demonstration, we will using K-fold cross validation (and it's widely used in NLP).
```
from sklearn.model_selection import KFold
from sklearn.naive_bayes import MultinomialNB
dataset = pd.read_csv('20news-18828.csv', header=None, delimiter=',', names=['label', 'text'])
X = dataset.text
y = dataset.label
# the train_test_split() function shuffles the dataset under the hood, but the KFold object
# does not; therefore, if your dataset is sorted, make sure to shuffle it. For the sake of time,
# we are doing 5 fold
kf = KFold(n_splits=5, shuffle=True)
for train_index, test_index in kf.split(X):
X_train, X_test, y_train, y_test = X[train_index], X[test_index], y[train_index], y[test_index]
cv = CountVectorizer()
X_train_counts = cv.fit_transform(X_train)
X_test_counts = cv.transform(X_test)
clf = MultinomialNB().fit(X_train_counts, y_train)
predicted = clf.predict(X_test_counts)
p, r, f1, _ = metrics.precision_recall_fscore_support(y_test, predicted, average='macro')
print("Macro-averaged Performance:\nPrecision: {0}, Recall: {1}, F1: {2}".format(p, r, f1))
```
### 6. Conclusion
In this tutorial, we went through some basic features of scikit-learn that allow us to perform some straightforward NLP/ML tasks. In this tutorial, we only use texts or bag of words as features; what if we want to incorporate some other features, such as document length, overall document sentiment, etc? In the next tutorial, we will address those issues and focus more on using scikit-learn for feature engineering.
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Automated Machine Learning
_**AutoML Model Testing for Classification**_
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Results Analysis and Test](#Results-Analysis-and-Test)
1. [Acknowledgements](#Acknowledgements)
## Introduction
In this example we use the associated credit card dataset to showcase how you can do model training, selection and test with AutoML for classification tasks. Besides this example notebook, you can find two other example notebooks with model testing for [regression](../regression/regression-TSI.ipynb) and [forecasting](../forecasting/forecasting-TSI.ipynb). Also, these [docs](https://docs.microsoft.com/en-us/azure/machine-learning/concept-automated-ml#training-validation-and-test-data) and [example code snippets](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-configure-cross-validation-data-splits#provide-test-data-preview) can give you more useful information.
This notebook is using remote compute to do all the computation.
If you are using an Azure Machine Learning Compute Instance, you are all set. Otherwise, go through the [configuration](../configuration.ipynb) notebook first if you haven't already to establish your connection to the AzureML Workspace.
In this notebook you will learn how to:
1. Setup Workspace and Compute Instance.
2. Prepare data and configure AutoML using `AutoMLConfig` to enable test run
3. Retrieve and explore the results.
4. Start remote test runs on demand
## Setup
As part of the setup you have already created an Azure ML `Workspace` object. For Automated ML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
```
import logging
from matplotlib import pyplot as plt
import pandas as pd
import os
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.core.dataset import Dataset
from azureml.train.automl import AutoMLConfig
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
```
print("This notebook was created using version 1.36.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
ws = Workspace.from_config()
# choose a name for experiment
experiment_name = "test-set-support-automl-classification-ccard-remote"
experiment = Experiment(ws, experiment_name)
output = {}
output["Subscription ID"] = ws.subscription_id
output["Workspace"] = ws.name
output["Resource Group"] = ws.resource_group
output["Location"] = ws.location
output["Experiment Name"] = experiment.name
pd.set_option("display.max_colwidth", -1)
outputDf = pd.DataFrame(data=output, index=[""])
outputDf.T
```
A compute target is required to execute the Automated ML run. In this tutorial, you create AmlCompute as your training compute resource.
If the AmlCompute with that name is already in your workspace this code will skip the creation process.
As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "cpu-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print("Found existing cluster, use it.")
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(
vm_size="STANDARD_DS12_V2", max_nodes=6
)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
## Data
Load the credit card dataset from a csv file containing both training features and labels. The features are inputs to the model, while the training labels represent the expected output of the model.
```
data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/creditcard.csv"
dataset = Dataset.Tabular.from_delimited_files(data)
label_column_name = "Class"
```
## Train
Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|classification or regression|
|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>average_precision_score_weighted</i><br><i>norm_macro_recall</i><br><i>precision_score_weighted</i>|
|**enable_early_stopping**|Stop the run if the metric score is not showing improvement.|
|**n_cross_validations**|Number of cross validation splits.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
**_You can find more information about primary metrics_** [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train#primary-metric)
### Data Splitting
Next, we'll do train/validation/test split to the data by specifying parameters in `AutoMLConfig`.
You can use `test_size` to determine what percentage of the data to be the test set, which will be used for getting predictions and metrics for the best model after all model training has completed. This parameter must be a floating point value between 0.0 and 1.0. For regression based tasks, random sampling is used. For classification tasks, stratified sampling is used. **Forecasting does not currently support specifying a test dataset using a train/test split**.
**Note that** if `test_size` and `validation_size` are both set, like the example below, the provided dataset will be first splitted into a training set and a test set. Then the training set will be splitted again into a new training set and a validation set. For example, if the original dataset has 1000 samples, `validation_size = 0.2` and `test_size = 0.1`, the final test set will have `1000 * 0.1 = 100` samples, training set will have `1000 * (1 - 0.1) * (1 - 0.2) = 720` samples and validation set will have `1000 * (1 - 0.1) * 0.2 = 180` samples
If you would prefer provide test data directly instead of using a train/test split, use the `test_data` parameter when creating the `AutoMLConfig`. In the [example notebook for regression](../regression/regression-TSI.ipynb) you can find an example using `test_data`. Test data in the form of a `Dataset` can be passed in to `AutoMLConfig` using the `test_data` parameter. When this parameter is specified, a test run will be started after all training runs have completed and the best model has been identified.
Note, the `test_data` and `test_size` `AutoMLConfig` parameters are mutually exclusive and can not be specified at the same time.
```
# Use test_size for train/validation/test split
automl_settings = {
"primary_metric": "average_precision_score_weighted",
"enable_early_stopping": True,
"max_concurrent_iterations": 6, # This is a limit for testing purpose, please increase it as per cluster size
"iterations": 10,
"experiment_timeout_hours": 0.25, # This is a time limit for testing purposes, remove it for real use cases, this will drastically limit ablity to find the best model possible
"verbosity": logging.INFO,
}
automl_config = AutoMLConfig(
task="classification",
compute_target=compute_target,
training_data=dataset,
label_column_name=label_column_name,
validation_size=0.2, # Use validation set split. This will be used to select the "winner" model
test_size=0.1, # Use test set split. This will be used only for the Winner model Test metrics
**automl_settings
)
```
Call the `submit` method on the experiment object and pass the run configuration. Depending on the data and the number of iterations this can run for a while. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
```
remote_run = experiment.submit(automl_config, show_output=True)
# If you need to retrieve a run that already started, use the following code
# from azureml.train.automl.run import AutoMLRun
# remote_run = AutoMLRun(experiment = experiment, run_id = '<replace with your run id>')
```
## Results Analysis and Test
### Widget for Monitoring Runs
The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details
```
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
remote_run.wait_for_completion(show_output=False)
```
### Explain model
Automated ML models can be explained and visualized using the SDK Explainability library.
### Retrieve the Best Model/Run
Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
```
best_run, fitted_model = remote_run.get_output()
fitted_model
```
#### Get the Predictions and Metrics Generated by the Test Run
Get the test run associated with the best run. When a remote test run is requested by passing in a value for `test_data` or `test_size` to `AutoMLConfig`, only the best training run will have a test run associated with it. To start a test run for models which are not associated with the best run or to start another best run test run, use `ModelProxy`. See the example later in the notebook in the __Test the fitted model remotely__ section for more details.
To see more details about the test run in Azure Machine Learning Studio (view its metrics, get a preview of the predictions, etc...) follow the link to the __Details Page__ listed in the next cells output.
```
test_run = next(best_run.get_children(type="automl.model_test"))
test_run.wait_for_completion(show_output=False, wait_post_processing=True)
test_run
```
Get the __metrics__ from the test run.
```
test_run_metrics = test_run.get_metrics()
for name, value in test_run_metrics.items():
print(f"{name}: {value}")
```
Get the __predictions__ from the test run.
```
test_run_details = test_run.get_details()
test_run_predictions = Dataset.get_by_id(
ws, test_run_details["outputDatasets"][0]["identifier"]["savedId"]
)
test_run_predictions.to_pandas_dataframe().head()
```
### Test the fitted model remotely (On-demand)
This is testing a model the same way it was tested at the end of the training. However, in this case it's performed on-demand, meaning that you can test the model at any time, not just right after training.
```
from azureml.train.automl.model_proxy import ModelProxy
# get test data from the full dataset
training_data, test_data = dataset.random_split(percentage=0.8, seed=223)
model_proxy = ModelProxy(best_run)
predictions, test_run_metrics = model_proxy.test(test_data)
predictions.to_pandas_dataframe().head()
pd.DataFrame.from_dict(test_run_metrics, orient="index", columns=["Value"])
```
## Acknowledgements
This Credit Card fraud Detection dataset is made available under the Open Database License: http://opendatacommons.org/licenses/odbl/1.0/. Any rights in individual contents of the database are licensed under the Database Contents License: http://opendatacommons.org/licenses/dbcl/1.0/ and is available at: https://www.kaggle.com/mlg-ulb/creditcardfraud
The dataset has been collected and analysed during a research collaboration of Worldline and the Machine Learning Group (http://mlg.ulb.ac.be) of ULB (Université Libre de Bruxelles) on big data mining and fraud detection.
More details on current and past projects on related topics are available on https://www.researchgate.net/project/Fraud-detection-5 and the page of the DefeatFraud project
Please cite the following works:
Andrea Dal Pozzolo, Olivier Caelen, Reid A. Johnson and Gianluca Bontempi. Calibrating Probability with Undersampling for Unbalanced Classification. In Symposium on Computational Intelligence and Data Mining (CIDM), IEEE, 2015
Dal Pozzolo, Andrea; Caelen, Olivier; Le Borgne, Yann-Ael; Waterschoot, Serge; Bontempi, Gianluca. Learned lessons in credit card fraud detection from a practitioner perspective, Expert systems with applications,41,10,4915-4928,2014, Pergamon
Dal Pozzolo, Andrea; Boracchi, Giacomo; Caelen, Olivier; Alippi, Cesare; Bontempi, Gianluca. Credit card fraud detection: a realistic modeling and a novel learning strategy, IEEE transactions on neural networks and learning systems,29,8,3784-3797,2018,IEEE
Dal Pozzolo, Andrea Adaptive Machine learning for credit card fraud detection ULB MLG PhD thesis (supervised by G. Bontempi)
Carcillo, Fabrizio; Dal Pozzolo, Andrea; Le Borgne, Yann-Aël; Caelen, Olivier; Mazzer, Yannis; Bontempi, Gianluca. Scarff: a scalable framework for streaming credit card fraud detection with Spark, Information fusion,41, 182-194,2018,Elsevier
Carcillo, Fabrizio; Le Borgne, Yann-Aël; Caelen, Olivier; Bontempi, Gianluca. Streaming active learning strategies for real-life credit card fraud detection: assessment and visualization, International Journal of Data Science and Analytics, 5,4,285-300,2018,Springer International Publishing
Bertrand Lebichot, Yann-Aël Le Borgne, Liyun He, Frederic Oblé, Gianluca Bontempi Deep-Learning Domain Adaptation Techniques for Credit Cards Fraud Detection, INNSBDDL 2019: Recent Advances in Big Data and Deep Learning, pp 78-88, 2019
Fabrizio Carcillo, Yann-Aël Le Borgne, Olivier Caelen, Frederic Oblé, Gianluca Bontempi Combining Unsupervised and Supervised Learning in Credit Card Fraud Detection Information Sciences, 2019
| github_jupyter |

### ODPi Egeria Hands-On Lab
# Welcome to the Configuring Discovery Servers Lab
## Introduction
ODPi Egeria is an open source project that provides open standards and implementation libraries to connect tools,
catalogues and platforms together so they can share information about data and technology (called metadata).
In this hands-on lab you will get a chance to run an Egeria metadata server, configure discovery services in a discovery engine and run the discovery engine in a discovery server.
## What is open discovery?
Open discovery is the ability to automatically analyse and create metadata about assets. ODPi Egeria provides an Open Discovery Framework (ODF) that defines open interfaces for components that provide specific types of discovery capability so that they can be called from discovery platforms offered by different vendors.
The Open Discovery Framework (ODF) provides standard interfaces for building **discovery services** and grouping them together into a useful collection of capability called a **discovery engine**.
ODPi Egeria then provides a governance server called the **discovery server** that can host one or more discovery engines.
## The scenario
Gary Geeke is the IT Infrastructure leader at Coco Pharmaceuticals. He has set up a number of OMAG Server Platforms and
is configuring the servers to run on them.

In this hands-on lab Gary is setting up a discovery server for the data lake. Gary's userId is `garygeeke`.
```
import requests
adminUserId = "garygeeke"
organizationName = "Coco Pharmaceuticals"
serverType = "Open Discovery Server"
```
In the **Metadata Server Configuration** lab, Gary configured servers for the OMAG Server Platforms shown in Figure 1:

> **Figure 1:** Coco Pharmaceuticals' OMAG Server Platforms
Below are the host name and port number where the core, data lake and development platforms will run. The discovery server will run on the Data Lake OMAG Server Platform.
```
import os
corePlatformURL = os.environ.get('corePlatformURL','http://localhost:8080')
dataLakePlatformURL = os.environ.get('dataLakePlatformURL','http://localhost:8081')
devPlatformURL = os.environ.get('devPlatformURL','http://localhost:8082')
```
There are two parts to setting up a discovery server. First the configuration for a discovery engine needs to be created and added to a metadata server. In this example, this will be the Data Lake Operations Metadata Server called `cocoMDS1`.
```
mdrServerName = "cocoMDS1"
```
Then the discovery server is configured with the location of the metadata server and the identity of one or more discovery engines.
When the discovery server starts it contacts the metadata server and retrieves the configuration for the discovery engine(s).
## Exercise 1 - Configuring the Discovery Engine
The discovery engine is configured using calls to the Discovery Engine OMAS. The commands all begin with this root.
```
configCommandURLRoot = dataLakePlatformURL + "/servers/" + mdrServerName + "/open-metadata/access-services/discovery-engine/users/" + adminUserId
```
The first configuration call is to create the discovery engine.
```
createDiscoveryEngineURL = configCommandURLRoot + '/discovery-engines'
print (createDiscoveryEngineURL)
jsonHeader = {'content-type':'application/json'}
body = {
"class" : "NewDiscoveryEngineRequestBody",
"qualifiedName" : "data-lake-discovery-engine",
"displayName" : "Data Lake Discovery Engine",
"description" : "Discovery engine used for onboarding assets."
}
response=requests.post(createDiscoveryEngineURL, json=body, headers=jsonHeader)
response.json()
discoveryEngineGUID=response.json().get('guid')
print (" ")
print ("The guid for the discovery engine is: " + discoveryEngineGUID)
print (" ")
createDiscoveryServiceURL = configCommandURLRoot + '/discovery-services'
print (createDiscoveryServiceURL)
jsonHeader = {'content-type':'application/json'}
body = {
"class" : "NewDiscoveryServiceRequestBody",
"qualifiedName" : "csv-asset-discovery-service",
"displayName" : "CSV Asset Discovery Service",
"description" : "Discovers columns for CSV Files.",
"connection" : {
"class": "Connection",
"type": {
"class": "ElementType",
"elementTypeId": "114e9f8f-5ff3-4c32-bd37-a7eb42712253",
"elementTypeName": "Connection",
"elementTypeVersion": 1,
"elementTypeDescription": "A set of properties to identify and configure a connector instance.",
"elementOrigin": "CONFIGURATION"
},
"guid": "1111abc7-2b13-4c4e-b840-97c4282f7416",
"qualifiedName": "csv-asset-discovery-service-implementation",
"displayName": "CSV Discovery Service Implementation Connector",
"description": "Connector to discover a CSV File.",
"connectorType": {
"class": "ConnectorType",
"type": {
"class": "ElementType",
"elementTypeId": "954421eb-33a6-462d-a8ca-b5709a1bd0d4",
"elementTypeName": "ConnectorType",
"elementTypeVersion": 1,
"elementTypeDescription": "A set of properties describing a type of connector.",
"elementOrigin": "LOCAL_COHORT"
},
"guid": "1111f73d-e343-abcd-82cb-3918fed81da6",
"qualifiedName": "CSVDiscoveryServiceProvider",
"displayName": "CSV File Discovery Service Provider Implementation",
"description": "This connector explores the content of a CSV File.",
"connectorProviderClassName": "org.odpi.openmetadata.accessservices.discoveryengine.samples.discoveryservices.CSVDiscoveryServiceProvider"
}
}
}
response=requests.post(createDiscoveryServiceURL, json=body, headers=jsonHeader)
response.json()
discoveryServiceGUID=response.json().get('guid')
print (" ")
print ("The guid for the discovery service is: " + discoveryServiceGUID)
print (" ")
```
Finally the discovery service is registered with the discovery engine.
```
registerDiscoveryServiceURL = configCommandURLRoot + '/discovery-engines/' + discoveryEngineGUID + '/discovery-services'
print (registerDiscoveryServiceURL)
jsonHeader = {'content-type':'application/json'}
body = {
"class" : "DiscoveryServiceRegistrationRequestBody",
"discoveryServiceGUID" : discoveryServiceGUID,
"assetTypes" : [ "small-csv" ]
}
response=requests.post(registerDiscoveryServiceURL, json=body, headers=jsonHeader)
response.json()
```
## Exercise 2 - Configuring the Discovery Server
The discovery server is to be called `discoDL01`.
```
organizationName = "Coco Pharmaceuticals"
serverType = "Open Discovery Server"
discoServerName = "discoDL01"
```
The code below sets up the basic properties of a governance server.
```
adminPlatformURL = dataLakePlatformURL
adminCommandURLRoot = adminPlatformURL + '/open-metadata/admin-services/users/' + adminUserId + '/servers/'
print (" ")
print ("Configuring the platform that the server will run on ...")
url = adminCommandURLRoot + discoServerName + '/server-url-root?url=' + dataLakePlatformURL
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
maxPageSize = '100'
print (" ")
print ("Configuring the maximum page size ...")
url = adminCommandURLRoot + discoServerName + '/max-page-size?limit=' + maxPageSize
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
print (" ")
print ("Configuring the server's type ...")
url = adminCommandURLRoot + discoServerName + '/server-type?typeName=' + serverType
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
print (" ")
print ("Configuring the server's owning organization ...")
url = adminCommandURLRoot + discoServerName + '/organization-name?name=' + organizationName
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
discoServerUserId = "discoDL01npa"
discoServerPassword = "discoDL01passw0rd"
print (" ")
print ("Configuring the server's userId ...")
url = adminCommandURLRoot + discoServerName + '/server-user-id?id=' + discoServerUserId
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
print (" ")
print ("Configuring the server's password (optional) ...")
url = adminCommandURLRoot + discoServerName + '/server-user-password?password=' + discoServerPassword
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
print (" ")
print ("Configuring the server's security connector ...")
url = adminCommandURLRoot + discoServerName + '/security/connection'
print ("POST " + url)
header={'content-type':'application/json'}
body = {
"class": "Connection",
"type": {
"class": "ElementType",
"elementTypeId": "114e9f8f-5ff3-4c32-bd37-a7eb42712253",
"elementTypeName": "Connection",
"elementTypeVersion": 1,
"elementTypeDescription": "A set of properties to identify and configure a connector instance.",
"elementOrigin": "CONFIGURATION"
},
"guid": "1213abc7-2b13-4c4e-b840-97c4282f7416",
"qualifiedName": "CocoPharmaceuticalsMetadataServerSecurityConnector",
"displayName": "Metadata Server Security Connector",
"description": "Connector to enforce authorization rules for accessing and updating metadata.",
"connectorType": {
"class": "ConnectorType",
"type": {
"class": "ElementType",
"elementTypeId": "954421eb-33a6-462d-a8ca-b5709a1bd0d4",
"elementTypeName": "ConnectorType",
"elementTypeVersion": 1,
"elementTypeDescription": "A set of properties describing a type of connector.",
"elementOrigin": "LOCAL_COHORT"
},
"guid": "1851f73d-e343-abcd-82cb-3918fed81da6",
"qualifiedName": "CocoPharmaServerSecurityConnectorType",
"displayName": "Coco Pharmaceuticals Server Security Connector Implementation",
"description": "This connector ensures only valid and authorized people can access the metadata.",
"connectorProviderClassName": "org.odpi.openmetadata.metadatasecurity.samples.CocoPharmaServerSecurityProvider"
}
}
response=requests.post(url, json=body, headers=header)
print ("Response: ")
print (response.json())
```
----
Next the discovery engine services need to be enabled.
```
print (" ")
print ("Configuring the access service URL and Server Name ...")
url = adminCommandURLRoot + discoServerName + '/discovery-server/access-service-root-url?accessServiceRootURL=' + dataLakePlatformURL
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
url = adminCommandURLRoot + discoServerName + '/discovery-server/access-service-server-name?accessServiceServerName=' + mdrServerName
print ("POST " + url)
response=requests.post(url)
print ("Response: ")
print (response.json())
print (" ")
print ("Configuring the server's discovery engine ...")
url = adminCommandURLRoot + discoServerName + '/discovery-server/set-discovery-engines'
print ("POST " + url)
header={'content-type':'application/json'}
body = [ discoveryEngineGUID ]
response=requests.post(url, json=body, headers=header)
print ("Response: ")
print (response.json())
```
## Exercise 3 - Discovering Assets
The final exercise is to run metadata discovery on a new asset.
```
petersUserId = "peterprofile"
serverAssetOwnerURL = dataLakePlatformURL + '/servers/' + mdrServerName + '/open-metadata/access-services/asset-owner/users/' + petersUserId
createAssetURL = serverAssetOwnerURL + '/assets/files/csv'
print (createAssetURL)
jsonHeader = {'content-type':'application/json'}
body = {
"class" : "NewFileAssetRequestBody",
"displayName" : "Week 1: Drop Foot Clinical Trial Measurements",
"description" : "One week's data covering foot angle, hip displacement and mobility measurements.",
"fullPath" : "file://secured/research/clinical-trials/drop-foot/DropFootMeasurementsWeek1.csv"
}
response=requests.post(createAssetURL, json=body, headers=jsonHeader)
response.json()
asset1guid=response.json().get('guid')
print (" ")
print ("The guid for asset 1 is: " + asset1guid)
print (" ")
discoveryCommandRootURL = dataLakePlatformURL + '/servers/' + discoServerName + '/open-metadata/discovery-server/users/' + petersUserId + '/discovery-engine/' + discoveryEngineGUID
assetType = "small-csv"
discoverAssetURL = discoveryCommandRootURL + '/asset-types/' + assetType + '/assets/' + asset1guid
response=requests.post(discoverAssetURL)
print ("Returns:")
print (response.json())
print (" ")
```
| github_jupyter |
```
import ftfy
ftfy.__version__
```
# Mojibake!
## What the h—ck happened to this text?
Robyn Speer
## Mojibake is when text ends up with the wrong Unicode characters due to an encoding mistake.
- It's Japanese for "ghost characters"
- ftfy is my Python library that fixes them. (get it with `pip install ftfy`)
```
ftfy.fix_text("merci de télécharger le plug-in")
```
## The mascot of ftfy
```
ftfy.fix_text("(Ã\xa0¸‡'̀⌣'ÃŒÂ\x81)Ã\xa0¸‡")
```
This little guy, and in fact every example in this talk, comes from mojibake I found in the wild -- usually on Twitter or in the OSCAR Web Corpus.
> **Side note**: there are a lot of tangents I would like to go off on, but this is a 10-minute talk and there's no time for tangents. So I'll be sprinkling in these side notes that I'll be skipping over as I present the talk.
>
> If you're interested in them, I suggest pausing the video, or reading the Jupyter Notebook version of this talk that I'll be linking later.
It used to be that 1 byte = 1 character, so there were at most 256 possible characters of text that could be shown on a computer. Here's some quick code to see them:
```
import blessings
term = blessings.Terminal() # enable colorful text
def displayable_codepoint(codepoint, encoding):
char = bytes([codepoint]).decode(encoding, 'replace')
if char == '�':
return '▓▓'
elif not char.isprintable():
return '░░'
else:
return char
def show_char_table(encoding):
print(f"encoding: {encoding}\n 0 1 2 3 4 5 6 7 8 9 a b c d e f\n")
for row in range(16):
print(f"{row*16:>02x}", end=" ")
if row == 0:
print(ftfy.formatting.display_center(term.green(" control characters "), 32, "░"))
elif row == 8 and encoding == 'latin-1':
print(ftfy.formatting.display_center(term.green(" here be dragons "), 32, "░"))
else:
for col in range(16):
char = displayable_codepoint(row * 16 + col, encoding)
print(f"{char:<2}", end="")
print()
```
# ASCII
- In the '60s, we agreed that 128 of these bytes should have well-defined meanings as characters of text. That's ASCII
- It worked pretty well for monolingual Americans
```
show_char_table("ascii")
```
- Basically everyone decided the other 128 bytes should be characters too
- So people started talking about "extended ASCII" which means "whatever my computer does with the other bytes"
- A different computer, in a different country or with a different OS, would do something different
- This is how mojibake started
Let's take a look at the different ways that ASCII got "extended", which are now called _codepages_.
Here's Latin-1. This encoding was used on a lot of UNIX-like systems before they switched to UTF-8, but probably the biggest reason you see it is that it's the first 256 characters of Unicode.
If you don't know about decoding text and you just replace each byte with the Unicode character with the same number, Latin-1 is the encoding you get by accident.
```
show_char_table('latin-1')
```
## Dragons? 🐉
I labeled rows 80 and 90 as "here be dragons" because they're full of control characters, but nobody ever agreed on what they do.
Text that includes these codepoints can be shown as nearly anything, including little boxes, characters from the Windows-1252 character set, or just messing up your whole terminal window.
> Side note: Control character 85 was an attempt to resolve the war between Unix line breaks (`0a`), Windows line breaks (`0d 0a`), and Mac Classic line breaks (`0d`), by introducing an _entirely new_ line break that wasn't even in ASCII.
>
> This is hilarious in retrospect. Clearly nobody would ever repeat that terrible idea, except the Unicode Consortium, who did it twice more.
>
> Anyway, ftfy has a function that turns all of these into byte `0a`.
```
show_char_table('windows-1252')
show_char_table('cp437')
show_char_table('macroman')
show_char_table("windows-1251")
```
## So text wasn't portable between different computers
```
phrase = "Plus ça change, plus c’est la même chose"
phrase.encode('windows-1252').decode('macroman')
```
## Vintage '90s mojibake
Maybe you've seen a ".NFO" file like this, telling you about the nice people who provided you with a DOS game for free:
```
crack_nfo = r"""
───── ────────────── ───────────────── ────────────── ─────────────── ────
▄█████▄ ▀█████▄ ████▄ ▀█████████ ████▄ ▄█████▄▀████▄ ▀██████▄ ▀████▄ ▄██▄
████████ █████▀ ██████ ▀████████ ██████ ▀███████▌██████ ███████ ████▄█████
███ ▀███▌█▀ ▄▄▄█▌▀█████ ▌████ ▄▄▄█▀▐████ ███▀▌▀█▌█▌ ███▌ ██▀▐████ ██████████
███ ▐██▌▌ ████ ▌████ ████ ████ ████ ██▌ ▌▐▌█▌ ████ ██ ████ ██▌▐█▌▐███
███ ▄███▌▄▄ ████ ████ ████ ████ ▄████ ██▌ █▄▄█▌ ████ ██ ▄████ ██ ▐█▌ ███
████████ ██ ████ ████ ████ ████ █████ ██▌ ▀▀██▌▐███▀ ██▐█████ ██ █ ███
██████▀ ▄▀▀ ████ ████ ████ ████ ▀████ ██▌███▄▐██▄▀▀ ▄███ ▀████ ██ ▄ ███
███▀ ▄▄██ ████ ▐████ ████ ████ ████ ██▌▐▐██▌█▀██▄ ████ ████ ██ ███
███ █████▄ ▀▀█ ████▀ ▐████ ░███ ▐████ ███▄▐██▌█▌▐██▌▐███ ▐███░ ██▌ r███
░██ █████████▄ ▐██▌▄██████ ▒░██ █████ ███████▌█▌▐███ ███ ███░▒ ███ o██░
▒░█ ▀███████▀ ▀ ███▐████▀ ▓▒░█ ▐███▌ ▀██████▐█▌▐███ ███ ▐█░▒▓ ██▌ y█░▒
- ▌─────▐▀─ ▄▄▄█ ── ▀▀ ───── ────── ▀▀▀ ─ ▐▀▀▀▀ ▀▀ ████ ──── ▀█▀ ─ ▀ ────▐ ─
╓────────────────────────[ RELEASE INFORMATION ]───────────────────────╖
╓────────────────────────────────────────────────────────────────────────────╖
║ -/\- THE EVEN MORE INCREDIBLE MACHINE FOR *DOS* FROM SIERRA/DYNAMIX -/\- ║
╙────────────────────────────────────────────────────────────────────────────╜
"""
```
> **Side note**: It's funny that most of the things we call "ASCII art" aren't ASCII.
```
print(crack_nfo.encode('cp437').decode('windows-1252'))
```
## Here comes UTF-8
Instead of using the other 128 bytes for 128 more characters, what if we used them as a variable-length encoding for the whole rest of Unicode?
There were other variable-length encodings, but UTF-8 is well-designed.
- It leaves ASCII as ASCII
- It doesn't overlap with ASCII _ever_: ASCII bytes always exclusively stand for the characters you expect
- It's self-synchronizing, so you can tell where each character starts and ends even with no context
Everyone recognized how good this idea was and switched every system to UTF-8. Encoding problems were solved forever, the end.
I'm kidding. UTF-8 is great but not everyone adopted it, especially not Microsoft who had just done _all that work_ to switch Windows APIs to UTF-16, which nobody likes. So now we have more kinds of mojibake.
```
# Code to look at the encoding of each character in UTF-8
def show_utf8(text):
for char in text:
char_bytes = char.encode('utf-8')
byte_sequence = ' '.join([f"{byte:>02x}" for byte in char_bytes])
print(f"{char} = {byte_sequence}")
text = "l’Hôpital"
show_utf8(text)
```
## What happens when not everyone is on board with UTF-8
```
text.encode('utf-8')
print(text.encode('utf-8').decode('windows-1252'))
print(text.encode('utf-8').decode('windows-1252').encode('utf-8').decode('windows-1252'))
EXAMPLES = [
"Merci de télécharger le plug-in",
"The Mona Lisa doesn’t have eyebrows.",
"I just figured out how to tweet emojis! â\x9a½í\xa0½í¸\x80í\xa0½í¸\x81í\xa0½í¸"
"\x82í\xa0½í¸\x86í\xa0½í¸\x8eí\xa0½í¸\x8eí\xa0½í¸\x8eí\xa0½í¸\x8e",
]
```
## We can recognize UTF-8 mojibake by its distinct patterns
The pattern of bytes that makes UTF-8 self-synchronizing is a pattern we can recognize even via a different encoding.
The example `doesn’t` is recognizable as UTF-8 / Windows-1252 mojibake, for example. `t‚Äö√†√∂¬¨¬©l‚Äö√†√∂¬¨¬©charger` is recognizable as UTF-8 / MacRoman mojibake.
When we see such a pattern, we encode as the appropriate other encoding, then decode as UTF-8.
```
EXAMPLES[0].encode('macroman').decode('utf-8')
_.encode('macroman').decode('utf-8')
_.encode('macroman').decode('utf-8')
```
## This is a job that the computer can do for us
```
# Some code to format the output of ftfy
from ftfy import fix_and_explain
from pprint import pprint
def show_explanation(text):
print(f"Original: {text}")
fixed, expl = fix_and_explain(text)
print(f" Fixed: {fixed}\n")
pprint(expl)
show_explanation(EXAMPLES[0])
show_explanation(EXAMPLES[1])
show_explanation(EXAMPLES[2])
```
> **Side note**: ftfy adds encodings to Python like `sloppy-windows-1252` and `utf-8-variants` when it's imported. Python is very strict about encoding standards, but to deal with real mojibake, we have to be very loose about them. For example, this tweet requires us to recognize and simulate a broken implementation of UTF-8.
## Avoiding false positives
ftfy only changes text that trips its mojibake-detector regex.
Here are some examples that ftfy _could_ consider to be UTF-8 mojibake and try to "fix", but thankfully it doesn't:
```
NEGATIVE_EXAMPLES = [
"Con il corpo e lo spirito ammaccato,\u00a0è come se nel cuore avessi un vetro conficcato.",
"2012—∞",
"TEM QUE SEGUIR, SDV SÓ…",
"Join ZZAJÉ’s Official Fan List",
"(-1/2)! = √π",
"OK??:( `¬´ ):"
]
for example in NEGATIVE_EXAMPLES:
# ftfy doesn't "fix" these because they're not broken, but we can manually try fixes
try:
print(example.encode('sloppy-windows-1252').decode('utf-8'))
except UnicodeError:
print(example.encode('macroman').decode('utf-8'))
assert ftfy.fix_encoding(example) == example
```
## The "badness" metric
ftfy doesn't just look for UTF-8-like patterns of characters, it also makes sure they are unlikely to be the intended text.
- Improbable combinations: accented letters next to currency signs, math symbols next to console line art
- Lots of things involving capital à and Â, where it doesn't look like they're being used for real in a capitalized word
- Unloved characters like `¶` PILCROW SIGN, `‡` DOUBLE DAGGER, `◊` LOZENGE next to other mojibake-related characters
...and many more cases that it looks for in a big regex.
Strings that match the regex can be re-decoded. Specific character sequences that match the regex can be reinterpreted even if they're inconsistent with the rest of the string.
> **Side note**: We used to try to categorize every Unicode character to find "badness". Now we only categorize the 400 or so characters that actually can appear in UTF-8 mojibake, because ftfy wouldn't have a reason to replace the other characters anyway.
```
text = "à perturber la réflexion des théologiens jusqu'à nos jours"
# We want to highlight the matches to this regular expression:
ftfy.badness.BADNESS_RE.findall(text)
# We'll just highlight it manually:
term = blessings.Terminal()
highlighted_text = term.on_yellow("à ") + "perturber la r" + term.on_yellow("é") + "flexion des th" + term.on_yellow("é") + "ologiens jusqu'à nos jours"
# Highlighted text shows matches for the 'badness' expression.
# If we've confirmed from them that this is mojibake, and there's a consistent fix, we
# can fix even text in contexts that were too unclear for the regex, such as the final Ã.
print(highlighted_text)
print(ftfy.fix_text(highlighted_text))
```
## ftfy is a hand-tuned heuristic. Why doesn't it use machine learning?
I don't want ftfy to have false positives. It does, but every one of them is a bug I should fix. The actual rate of false positives should be once in several gigabytes of natural text.
- Machine learning techniques aren't designed to have error rates this low
- Machine learning would have a tendency to make its output look like what a language model "expects" even if the text didn't say that
## Why does mojibake keep happening?
The top 3 root causes:
1. Microsoft Excel
2. Programming language APIs that let you confuse bytes and text
3. An outdated heuristic called `chardet`
## chardet
`chardet` is a heuristic that takes in unlabeled bytes and tries to guess what encoding they're in. It was designed as part of Netscape Navigator in 1998, then ported to Python and other languages.
It doesn't know that the correct answer to "what encoding is this?" is usually "UTF-8", and it thinks emoji are some kind of Turkish.
> **Side note**: I recognize that we could be looking at ftfy the same way one day, particularly if I stop updating ftfy's heuristic. But chardet is fundamentally built on assumptions that aren't true anymore, and its original developer decided long ago that he'd written enough Python for a lifetime and he was going to do something else.
>
> It was an okay idea at the time, but we should have been able to replace it in two decades.
## Microsoft Excel and the Three Decades of Backward Compatibility
- Excel gives you lots of text formats you can export in, and _every one of them is wrong in 2021_
- Even the one that says "Unicode text" is not what you want
- The format of "CSV" depends on your OS, language, and region, like pre-Unicode days, and it will almost certainly mojibake your text
I know you probably have to use Excel sometimes, but my recommendation is to make CSVs with LibreOffice or Google Spreadsheets.

## How to avoid mojibake
The bad news is that English speakers can go for a long time without noticing mojibake.
The good news is that emoji are *everywhere*, people expect them to work, and they quickly reveal if you're doing Unicode right.
- Use lots of emoji! 👍💚🌠
- Use non-ASCII text, such as emoji, in your code, your UIs, your test cases
- This is similar to how Web frameworks used to pass around parameters like `&snowman=☃` and `&utf8=✔` to make sure browsers used UTF-8 and detect if they came out wrong
> **Side note**: Services such as Slack and Discord don't use Unicode for their emoji. They use ASCII strings like `:green-heart:` and turn them into images. These won't help you test anything. I recommend getting emoji for your test cases by copy-pasting them from emojipedia.org.
## Thanks!
- ftfy documentation: https://ftfy.readthedocs.org
- My e-mail address: rspeer@arborelia.net
- I'm @r_speer on Twitter
- I'll link from my Twitter to the notebook version of this talk
- BTW, I'm on the job market right now
Fonts I used for code in the presentation:
- Input: Fantasque Sans Mono
- Output: Fira Code
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Resuming Tensorflow training from previous run
In this tutorial, you will resume a mnist model in TensorFlow from a previously submitted run.
## Prerequisites
* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning (AML)
* Go through the [configuration notebook](../../../configuration.ipynb) to:
* install the AML SDK
* create a workspace and its configuration file (`config.json`)
* Review the [tutorial](../train-hyperparameter-tune-deploy-with-tensorflow/train-hyperparameter-tune-deploy-with-tensorflow.ipynb) on single-node TensorFlow training using the SDK
```
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
```
## Diagnostics
Opt-in diagnostics for better experience, quality, and security of future releases.
```
from azureml.telemetry import set_diagnostics_collection
set_diagnostics_collection(send_diagnostics=True)
```
## Initialize workspace
Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`.
```
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep='\n')
```
## Create or Attach existing AmlCompute
You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.
**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.
As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# choose a name for your cluster
cluster_name = "gpu-cluster"
try:
compute_target = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing compute target.')
except ComputeTargetException:
print('Creating a new compute target...')
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6',
max_nodes=4)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# use get_status() to get a detailed status for the current cluster.
print(compute_target.get_status().serialize())
```
The above code creates a GPU cluster. If you instead want to create a CPU cluster, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`.
## Create a Dataset for Files
A Dataset can reference single or multiple files in your datastores or public urls. The files can be of any format. Dataset provides you with the ability to download or mount the files to your compute. By creating a dataset, you create a reference to the data source location. If you applied any subsetting transformations to the dataset, they will be stored in the dataset as well. The data remains in its existing location, so no extra storage cost is incurred. [Learn More](https://aka.ms/azureml/howto/createdatasets)
```
#initialize file dataset
from azureml.core.dataset import Dataset
web_paths = ['http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz'
]
dataset = Dataset.File.from_files(path = web_paths)
```
you may want to register datasets using the register() method to your workspace so they can be shared with others, reused across various experiments, and referred to by name in your training script.
You can try get the dataset first to see if it's already registered.
```
dataset_registered = False
try:
temp = Dataset.get_by_name(workspace = ws, name = 'mnist-dataset')
dataset_registered = True
except:
print("The dataset mnist-dataset is not registered in workspace yet.")
if not dataset_registered:
#register dataset to workspace
dataset = dataset.register(workspace = ws,
name = 'mnist-dataset',
description='training and test dataset',
create_new_version=True)
# list the files referenced by dataset
dataset.to_path()
```
## Train model on the remote compute
### Create a project directory
Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on.
```
import os
script_folder = './tf-resume-training'
os.makedirs(script_folder, exist_ok=True)
```
Copy the training script `tf_mnist_with_checkpoint.py` into this project directory.
```
import shutil
# the training logic is in the tf_mnist_with_checkpoint.py file.
shutil.copy('./tf_mnist_with_checkpoint.py', script_folder)
# the utils.py just helps loading data from the downloaded MNIST dataset into numpy arrays.
shutil.copy('./utils.py', script_folder)
```
### Create an experiment
Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed TensorFlow tutorial.
```
from azureml.core import Experiment
experiment_name = 'tf-resume-training'
experiment = Experiment(ws, name=experiment_name)
```
### Create a TensorFlow estimator
The AML SDK's TensorFlow estimator enables you to easily submit TensorFlow training jobs for both single-node and distributed runs. For more information on the TensorFlow estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-tensorflow).
The TensorFlow estimator also takes a `framework_version` parameter -- if no version is provided, the estimator will default to the latest version supported by AzureML. Use `TensorFlow.get_supported_versions()` to get a list of all versions supported by your current SDK version or see the [SDK documentation](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.dnn?view=azure-ml-py) for the versions supported in the most current release.
```
from azureml.train.dnn import TensorFlow
script_params={
'--data-folder': dataset.as_named_input('mnist').as_mount()
}
estimator= TensorFlow(source_directory=script_folder,
compute_target=compute_target,
script_params=script_params,
entry_script='tf_mnist_with_checkpoint.py',
use_gpu=True,
pip_packages=['azureml-dataset-runtime[pandas,fuse]'])
```
In the above code, we passed our training data reference `ds_data` to our script's `--data-folder` argument. This will 1) mount our datastore on the remote compute and 2) provide the path to the data zip file on our datastore.
### Submit job
### Run your experiment by submitting your estimator object. Note that this call is asynchronous.
```
run = experiment.submit(estimator)
print(run)
```
### Monitor your run
You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
```
from azureml.widgets import RunDetails
RunDetails(run).show()
```
Alternatively, you can block until the script has completed training before running more code.
```
run.wait_for_completion(show_output=True)
```
# Now let's resume the training from the above run
First, we will get the DataPath to the outputs directory of the above run which
contains the checkpoint files and/or model
```
model_location = run._get_outputs_datapath()
```
Now, we will create a new TensorFlow estimator and pass in the model location. On passing 'resume_from' parameter, a new entry in script_params is created with key as 'resume_from' and value as the model/checkpoint files location and the location gets automatically mounted on the compute target.
```
from azureml.train.dnn import TensorFlow
script_params={
'--data-folder': dataset.as_named_input('mnist').as_mount()
}
estimator2 = TensorFlow(source_directory=script_folder,
compute_target=compute_target,
script_params=script_params,
entry_script='tf_mnist_with_checkpoint.py',
resume_from=model_location,
use_gpu=True,
pip_packages=['azureml-dataset-runtime[pandas,fuse]'])
```
Now you can submit the experiment and it should resume from previous run's checkpoint files.
```
run2 = experiment.submit(estimator2)
print(run2)
run2.wait_for_completion(show_output=True)
```
| github_jupyter |
This notebook investigates the relative performance of scipy, numpy, and pyculib for FFTs
```
import time
import numpy as np
from numpy.fft import fft2 as nfft2, fftshift as nfftshift
from scipy.fftpack import fft2 as sfft2, fftshift as sfftshift
from prysm.mathops import cu_fft2
from matplotlib import pyplot as plt
%matplotlib inline
def npfft2(array):
return nfftshift(nfft2(nfftshift(array)))
def spfft2(array):
return sfftshift(sfft2(sfftshift(array)))
def cufft2(array):
return nfftshift(cu_fft2(nfftshift(array)))
arr_sizes = [16, 32, 128, 256, 512, 1024, 2048, 4048, 8096] # 8096x8096 arrays will require ~6GB of RAM to run
def test_algorithm_speed(function):
data = [np.random.rand(size, size) for size in arr_sizes]
times = []
for dat in data:
t0 = time.time()
try:
function(dat)
t1 = time.time()
times.append(t1-t0)
except Exception as e:
# probably cuFFT error -- array too big to fit in GPU memory
times.append(np.nan)
return times
ntrials = 5
results_np = np.empty((len(arr_sizes),ntrials), dtype='float64')
results_sp = np.empty((len(arr_sizes),ntrials), dtype='float64')
results_cu = np.empty((len(arr_sizes),ntrials), dtype='float64')
for t in range(ntrials):
results_np[:,t] = test_algorithm_speed(npfft2)
results_sp[:,t] = test_algorithm_speed(spfft2)
results_cu[:,t] = test_algorithm_speed(cufft2)
results_np *= 1e3
results_sp *= 1e3
results_cu *= 1e3
cpu = 'Intel i7-7700HQ 4c/8t @ 3.20Ghz\nnVidia GTX 1050 4GB'
plt.style.use('ggplot')
fig, (ax1, ax2, ax3) = plt.subplots(ncols=3, figsize=(16,4))
ax1.plot(arr_sizes, results_np.mean(axis=1), lw=3, label='numpy')
ax1.plot(arr_sizes, results_sp.mean(axis=1), lw=3, label='scipy')
ax1.plot(arr_sizes, results_sp.mean(axis=1), lw=3, label='cuda')
ax1.legend()
ax1.set(xscale='log', xlabel='Linear Array Size',
yscale='log', ylabel='Execution Time [ms, log]')
ax2.plot(arr_sizes, results_np.mean(axis=1), lw=3, label='numpy')
ax2.plot(arr_sizes, results_sp.mean(axis=1), lw=3, label='scipy')
ax2.plot(arr_sizes, results_sp.mean(axis=1), lw=3, label='cuda')
ax2.legend()
ax2.set(xlabel='Linear Array Size', ylabel='Execution Time [ms, linear]')
ax3.plot(arr_sizes, results_np.mean(axis=1), lw=3, label='numpy')
ax3.plot(arr_sizes, results_sp.mean(axis=1), lw=3, label='scipy')
ax3.plot(arr_sizes, results_sp.mean(axis=1), lw=3, label='cuda')
ax3.legend()
ax3.set(xlabel='Linear Array Size', ylabel='Execution Time [ms, linear]', xlim=(0,2048), ylim=(0,500))
plt.suptitle(cpu + ' FFT performance');
```
note that GPU performance is currently severely handicapped by transfer from CPU<-->GPU, and fftshifts being done in numpy and not on the GPU itself.
| github_jupyter |
# DAT257x: Reinforcement Learning Explained
## Lab 5: Temporal Difference Learning
### Exercise 5.4: Q-Learning Agent
```
import numpy as np
import sys
if "../" not in sys.path:
sys.path.append("../")
from lib.envs.simple_rooms import SimpleRoomsEnv
from lib.envs.windy_gridworld import WindyGridworldEnv
from lib.envs.cliff_walking import CliffWalkingEnv
from lib.simulation import Experiment
class Agent(object):
def __init__(self, actions):
self.actions = actions
self.num_actions = len(actions)
def act(self, state):
raise NotImplementedError
class QLearningAgent(Agent):
def __init__(self, actions, epsilon=0.01, alpha=0.5, gamma=1):
super(QLearningAgent, self).__init__(actions)
## TODO 1
## Initialize empty dictionary here
## In addition, initialize the value of epsilon, alpha and gamma
def stateToString(self, state):
mystring = ""
if np.isscalar(state):
mystring = str(state)
else:
for digit in state:
mystring += str(digit)
return mystring
def act(self, state):
stateStr = self.stateToString(state)
action = np.random.randint(0, self.num_actions)
## TODO 2
## Implement epsilon greedy policy here
return action
def learn(self, state1, action1, reward, state2, done):
state1Str = self.stateToString(state1)
state2Str = self.stateToString(state2)
## TODO 3
## Implement the q-learning update here
"""
Q-learning Update:
Q(s,a) <- Q(s,a) + alpha * (reward + gamma * max(Q(s') - Q(s,a))
or
Q(s,a) <- Q(s,a) + alpha * (td_target - Q(s,a))
or
Q(s,a) <- Q(s,a) + alpha * td_delta
"""
interactive = True
%matplotlib nbagg
env = SimpleRoomsEnv()
agent = QLearningAgent(range(env.action_space.n))
experiment = Experiment(env, agent)
experiment.run_qlearning(10, interactive)
interactive = False
%matplotlib inline
env = SimpleRoomsEnv()
agent = QLearningAgent(range(env.action_space.n))
experiment = Experiment(env, agent)
experiment.run_qlearning(50, interactive)
interactive = True
%matplotlib nbagg
env = CliffWalkingEnv()
agent = QLearningAgent(range(env.action_space.n))
experiment = Experiment(env, agent)
experiment.run_qlearning(10, interactive)
interactive = False
%matplotlib inline
env = CliffWalkingEnv()
agent = QLearningAgent(range(env.action_space.n))
experiment = Experiment(env, agent)
experiment.run_qlearning(100, interactive)
interactive = False
%matplotlib inline
env = WindyGridworldEnv()
agent = QLearningAgent(range(env.action_space.n))
experiment = Experiment(env, agent)
experiment.run_qlearning(50, interactive)
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
```
First, define a function that takes two inputs (a mean and a standard deviation) and spits out a random number.
The number should be generated randomly from a normal distribution with that mean and that standard deviation. See [numpy.random.normal](https://docs.scipy.org/doc/numpy/reference/generated/numpy.random.normal.html), though, implementing the PDF for a normal distribution is a good exercise if you finish early.
```
def rand_val(mean, stdev):
##################
# Your Code Here #
##################
return np.random.normal(mean, stdev)
```
Now define a function that takes in a mean, a standard deviation, and a number n, and returns a numpy vector with n components.
The components should be randomly generated from a normal distribution with that mean and standard deviation
```
def rand_vec(mean, stdev, n):
##################
# Your Code Here #
##################
return np.random.normal(mean, stdev, n)
```
Recall that there are many ways to compute the distance between two vectors. However you wish to do so, write a function that takes in
two vectors and returns the distance between them. Since this function only makes sense if the two vectors have the same number of
components, this function should return -1 if the two vectors live in different numbers of dimensions
```
def vec_dist(vecA, vecB):
##################
# Your Code Here #
##################
if vecA.shape != vecB.shape:
return -1
return np.linalg.norm(vecA - vecB)
```
From here on out, every component you randomly generate should come from a normal distribution centered at 0 with standard
deviation 1. With that in mind, write a function that takes in a number n (the number of dimensions), creates two random
vectors in n-dimensional space, calculates the distance between them, and returns that distance.
```
def rand_vec_dist(n):
##################
# Your Code Here #
##################
vecA, vecB = rand_vec(0, 1, n), rand_vec(0, 1, n)
return vec_dist(vecA, vecB)
```
Using what you've made so far, what is the average distance between two points in 1-dimensional space? In other words, between
two regular old floating point numbers with mean 0 and standard deviation 1. After you calculate this, store it as the first
component in a Pythin list named average_dist_list.
```
def average_dist_calculator(n, trials):
##################
# Your Code Here #
##################
return np.mean([rand_vec_dist(n) for _ in range(trials)])
average_dist_list = [average_dist_calculator(1, 50)]
print(average_dist_list)
```
What's the average distance between two points in 2-dimensional space? What about 3? Keep calculating these average distances and
appending them to average_dist_list up through 200-dimensional space.
```
##################
# Your Code Here #
##################
average_dist_list = [average_dist_calculator(n, 50) for n in range(0, 201)]
print(average_dist_list)
n = np.arange(0, 201)
plt.plot(n,average_dist_list);
```
As the number of dimensions n grows larger and larger, approximately how does the average distance grow?
It should be well-approximated by the function f(n) = c*n^p for some numbers c and p. However you choose to do so, find the values of c and p.
Using my reasoning from the [Vector Magnitude Coding Challenge](https://github.com/rayheberer/LambdaSchoolDataScience/blob/master/Week%201%20Mathematical%20Foundations/Code%20Challenges/Day%204%20Vector%20Magnitude.ipynb), I expect that the average distance between points will be proportional to the square root of the dimension of the space.
I will first create square-root features, and then fit a linear estimator to them.
```
from sklearn.linear_model import LinearRegression
root_n = np.sqrt(n)
plt.plot(root_n,average_dist_list, 'bo');
# the y-intercept should be zero, since it is not meaningful for the average distance between objects in the zero vector space to be nonzero
model = LinearRegression(fit_intercept=False)
root_n = root_n.reshape((-1, 1))
average_dist_list = np.array(average_dist_list).reshape((-1, 1))
model.fit(root_n, average_dist_list);
c = model.coef_
print(c)
```
So our function is about $f(n) = 1.41\sqrt{n}$
```
line = c*np.sqrt(n)
line = line.reshape(-1,)
plt.scatter(n,average_dist_list, s=15)
plt.plot(n, line, color='red');
```
| github_jupyter |
```
import argparse
import logging
import os
import torch
import sys
sys.path.append(os.path.join(os.path.dirname("__file__"), '../multimodal_seq2seq_gSCAN/'))
import random
import copy
from seq2seq.gSCAN_dataset import GroundedScanDataset
from seq2seq.model import Model
from seq2seq.train import train
from seq2seq.predict import predict_and_save
from tqdm import tqdm, trange
from GroundedScan.dataset import GroundedScan
from typing import List
from typing import Tuple
from collections import defaultdict
from collections import Counter
import json
import numpy as np
from seq2seq.gSCAN_dataset import Vocabulary
from seq2seq.helpers import sequence_accuracy
from experiments_utils import *
FORMAT = "%(asctime)-15s %(message)s"
logging.basicConfig(format=FORMAT, level=logging.DEBUG,
datefmt="%Y-%m-%d %H:%M")
logger = logging.getLogger(__name__)
def isnotebook():
try:
shell = get_ipython().__class__.__name__
if shell == 'ZMQInteractiveShell':
return True # Jupyter notebook or qtconsole
elif shell == 'TerminalInteractiveShell':
return False # Terminal running IPython
else:
return False # Other type (?)
except NameError:
return False # Probably standard Python interpreter
use_cuda = True if torch.cuda.is_available() and not isnotebook() else False
device = "cuda" if use_cuda else "cpu"
if use_cuda:
logger.info("Using CUDA.")
logger.info("Cuda version: {}".format(torch.version.cuda))
def evaluate_syntactic_dependency(flags):
for argument, value in flags.items():
logger.info("{}: {}".format(argument, value))
# 1. preparing datasets
logger.info("Loading datasets.")
compositional_splits_data_path = os.path.join(flags["data_directory"], "dataset.txt")
compositional_splits_preprocessor = DummyGroundedScanDataset(compositional_splits_data_path,
flags["data_directory"],
input_vocabulary_file=flags["input_vocab_path"],
target_vocabulary_file=flags["target_vocab_path"],
generate_vocabulary=False,
k=flags["k"])
compositional_splits_dataset = \
GroundedScan.load_dataset_from_file(
compositional_splits_data_path,
save_directory=flags["output_directory"],
k=flags["k"])
logger.info("Loading models.")
# 2. load up models
raw_example = None
for _, example in enumerate(compositional_splits_dataset.get_examples_with_image(flags["split"], True)):
raw_example = example
break
single_example = compositional_splits_preprocessor.process(raw_example)
model = Model(input_vocabulary_size=compositional_splits_preprocessor.input_vocabulary_size,
target_vocabulary_size=compositional_splits_preprocessor.target_vocabulary_size,
num_cnn_channels=compositional_splits_preprocessor.image_channels,
input_padding_idx=compositional_splits_preprocessor.input_vocabulary.pad_idx,
target_pad_idx=compositional_splits_preprocessor.target_vocabulary.pad_idx,
target_eos_idx=compositional_splits_preprocessor.target_vocabulary.eos_idx,
**input_flags)
model = model.cuda() if use_cuda else model
_ = model.load_model(flags["resume_from_file"])
# TODO: let us enable multi-gpu settings here to save up times
logger.info("Starting evaluations.")
input_levDs = []
pred_levDs = []
accuracies = []
corrupt_accuracies = []
example_count = 0
limit = flags["max_testing_examples"]
split = flags["split"]
dataloader = [example for example in compositional_splits_dataset.get_examples_with_image(split, True)]
random.shuffle(dataloader) # shuffle this to get a unbiased estimate of accuracies
dataloader = dataloader[:limit] if limit else dataloader
for _, example in enumerate(tqdm(dataloader, desc="Iteration")):
# non-corrupt
single_example = compositional_splits_preprocessor.process(example)
output = predict_single(single_example, model=model,
max_decoding_steps=30,
pad_idx=compositional_splits_preprocessor.target_vocabulary.pad_idx,
sos_idx=compositional_splits_preprocessor.target_vocabulary.sos_idx,
eos_idx=compositional_splits_preprocessor.target_vocabulary.eos_idx,
device=device)
pred_command = compositional_splits_preprocessor.array_to_sentence(output[3], vocabulary="target")
accuracy = sequence_accuracy(output[3], output[4][0].tolist()[1:-1])
accuracies += [accuracy]
# corrupt
corrupt_example = make_corrupt_example(example, flags["corrupt_methods"])
corrupt_single_example = compositional_splits_preprocessor.process(corrupt_example)
corrupt_output = predict_single(corrupt_single_example, model=model,
max_decoding_steps=30,
pad_idx=compositional_splits_preprocessor.target_vocabulary.pad_idx,
sos_idx=compositional_splits_preprocessor.target_vocabulary.sos_idx,
eos_idx=compositional_splits_preprocessor.target_vocabulary.eos_idx,
device=device)
corrupt_pred_command = compositional_splits_preprocessor.array_to_sentence(corrupt_output[3], vocabulary="target")
corrupt_accuracy = sequence_accuracy(corrupt_output[3], corrupt_output[4][0].tolist()[1:-1])
corrupt_accuracies += [corrupt_accuracy]
input_levD = levenshteinDistance(example['input_command'], corrupt_example['input_command'])
pred_levD = levenshteinDistance(pred_command, corrupt_pred_command)
input_levDs.append(input_levD)
pred_levDs.append(pred_levD)
example_count += 1
exact_match = 0
for acc in accuracies:
if acc == 100:
exact_match += 1
exact_match = exact_match * 1.0 / len(accuracies)
corrupt_exact_match = 0
for acc in corrupt_accuracies:
if acc == 100:
corrupt_exact_match += 1
corrupt_exact_match = corrupt_exact_match * 1.0 / len(corrupt_accuracies)
logger.info("Eval Split={}, Original Exact Match %={}, Corrupt Exact Match %={}".format(split, exact_match, corrupt_exact_match))
return {"input_levDs" : input_levDs,
"pred_levDs" : pred_levDs,
"accuracies" : accuracies,
"corrupt_accuracies" : corrupt_accuracies}
if __name__ == "__main__":
input_flags = vars(get_gSCAN_parser().parse_args())
saved_to_dict = evaluate_syntactic_dependency(flags=input_flags)
split = input_flags["split"]
corrupt_methods = input_flags["corrupt_methods"]
if input_flags["save_eval_result_dict"]:
torch.save(saved_to_dict,
os.path.join(
input_flags["output_directory"],
f"eval_result_split_{split}_corrupt_{corrupt_methods}_dict.bin")
)
else:
logger.info("Skip saving results.")
```
| github_jupyter |
```
from collections import Counter
import numpy as np
import pandas as pd
import re
import glob
import gzip
import json
from sklearn.linear_model import LogisticRegression
from sklearn.feature_extraction import DictVectorizer
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score
def load_data(datafile):
"""
Read your data into a single pandas dataframe where
- each row is an instance to be classified
(this could be a tweet, user, or news article, depending on your project)
- there is a column called `label` which stores the class label (e.g., the true
category for this row)
"""
bots = []
humans = []
folder = ['/bots', '/humans']
name = '/*.json.gz'
for f in folder:
paths = glob.glob(datafile + f + name)
for p in paths:
with gzip.open(p, 'r') as file:
for line in file:
if f == folder[0]:
bots.append(json.loads(line))
elif f == folder[1]:
humans.append(json.loads(line))
df_bots = pd.DataFrame(bots)[['screen_name','tweets','listed_count']]
df_bots['label'] = 'bot'
df_humans = pd.DataFrame(humans)[['screen_name','tweets','listed_count']]
df_humans['label'] = 'human'
frames = [df_bots, df_humans]
df = pd.concat(frames)
users = bots + humans
tweets = [u['tweets'] for u in users]
text = [d['full_text'] for t in tweets for d in t]
# tweets_avg_len = []
tweets_avg_mentions = []
tweets_avg_urls = []
factor = 100
for u in users:
tweets = u['tweets'] # a list of dicts
texts = [t['full_text'] for t in tweets]
# avg_len = sum(map(len, texts))/len(texts)
# tweets_avg_len.append(int(avg_len))
count_mention = 0
count_url = 0
for s in texts:
if 'http' in s:
count_url+=1
if '@' in s:
count_mention+=1
tweets_avg_urls.append(100 * count_url / len(texts))
tweets_avg_mentions.append(100 * count_mention / len(texts))
# df['tweets_avg_len'] = tweets_avg_len
df['tweets_avg_urls'] = tweets_avg_urls
df['tweets_avg_mentions'] = tweets_avg_mentions
return df
# df = load_data('~/Dropbox/elevate/harassment/training_data/data.csv.gz')
df = load_data('/Users/lcj/small')
df
# what is the distribution over class labels?
df.label.value_counts()
df.dtypes
def make_features(df):
vec = DictVectorizer()
feature_dicts = []
labels_to_track = ['tweets_avg_urls', 'tweets_avg_mentions','listed_count']
for i, row in df.iterrows():
features = {}
features['tweets_avg_urls'] = row['tweets_avg_urls']
features['tweets_avg_mentions'] = row['tweets_avg_mentions']
features['listed_count'] = row['listed_count']
feature_dicts.append(features)
X = vec.fit_transform(feature_dicts)
# print(X)
return X, vec
X, vec = make_features(df)
# what are dimensions of the feature matrix?
X.shape
# what are the feature names?
# vocabulary_ is a dict from feature name to column index
vec.vocabulary_
# how often does each word occur?
for word, idx in vec.vocabulary_.items():
print('%20s\t%d' % (word, X[:,idx].sum()))
# can also get a simple list of feature names:
vec.get_feature_names()
# e.g., first column is 'hate', second is 'love', etc.
# we'll first store the classes separately in a numpy array
y = np.array(df.label)
Counter(y)
# to find the row indices with hostile label
np.where(y=='bot')[0]
# np.where(y=='human')[0]
# store the class names
class_names = set(df.label)
# how often does each word appear in each class?
for word, idx in vec.vocabulary_.items():
for class_name in class_names:
class_idx = np.where(y==class_name)[0]
print('%20s\t%20s\t%d' % (word, class_name, X[class_idx, idx].sum()))
a= [10,50,100,200]
b = [ 0.001, 0.0001, 0.00001]
for hidden_layer_sizes in a:
for alpha in b:
clf = MLPClassifier(hidden_layer_sizes = (50,) , activation='relu', solver='adam', alpha=0.0001)
clf.fit(X, y)
kf = KFold(n_splits=5, shuffle=True, random_state=42)
accuracies = []
for train, test in kf.split(X):
clf.fit(X[train], y[train])
pred = clf.predict(X[test])
accuracies.append(accuracy_score(y[test], pred))
clf.coefs_
print('accuracy over all cross-validation folds: %s' % str(accuracies))
print('mean=%.2f std=%.2f' % (np.mean(accuracies), np.std(accuracies)))
c= [1, 3, 5]
d = [100, 200, 300]
for min_samples_leaf in c:
for n_estimators in d:
rand = RandomForestClassifier(n_estimators=n_estimators, min_samples_leaf= min_samples_leaf)
rand.fit(X, y)
kf = KFold(n_splits=5, shuffle=True, random_state=42)
accuracies = []
for train, test in kf.split(X):
rand.fit(X[train], y[train])
pred = rand.predict(X[test])
accuracies.append(accuracy_score(y[test], pred))
print(rand.feature_importances_)
print('accuracy over all cross-validation folds: %s' % str(accuracies))
print('mean=%.2f std=%.2f' % (np.mean(accuracies), np.std(accuracies)))
```
| github_jupyter |
# Jupyter Notebook problems in the Essentials of Paleomagnetism Textbook by L. Tauxe
## Problems in Chapter 12
## Problem 1a
We can look at the data from within this notebook in the same way we did for Chapter 11. Let's dispense with the preliminaries:
```
# import PmagPy functions
import pmagpy.pmag as pmag
import pmagpy.ipmag as ipmag
import numpy as np # while we are at it, let's import numpy
import matplotlib.pyplot as plt # set up for plotting
%matplotlib inline
```
and get to it using **ipmag.plot_net()** with **ipmag.plot_di( )**.
```
data=np.loadtxt('Chapter_12/ps12-1a.di')# read in data
ipmag.plot_net(1) # plot the equal area net
ipmag.plot_di(di_block=data,title="Problem 12.1a directions") # put on the directions
```
This looks Fisherian to me! See how the data are pretty much symmetrical about the mean direction? But let's check with either **ipmag.fishqq( )**.
```
help(ipmag.fishqq)
ipmag.fishqq(di_block=data) # use data read in problem 1a.
```
And it looks like at the 95% confidence level, this data set IS Fisherian.
## Problem 1b
Now let's try the other data set.
```
data=np.loadtxt('Chapter_12/ps12-1b.di')# read in data
ipmag.plot_net(1) # plot the equal area net
ipmag.plot_di(di_block=data,title="Problem 12.1b directions") # put on the directions
```
Whoa! This one does not look Fisherian. See how there are 'outliers' and the data are spread out more in inclination than in declination? So let's check it.
```
ipmag.fishqq(di_block=data) # use data read in problem 1a.
```
And, yeah, this one totally failed.
Let's take a look at the data in Chapter_12/ps12-1c.di
```
data=np.loadtxt('Chapter_12/ps12-1c.di')# read in data
ipmag.plot_net(1) # plot the equal area net
ipmag.plot_di(di_block=data,title="Problem 12.1c directions") # put on the directions
```
And now get the Fisher mean using **ipmag.fisher_mean**.
```
ipmag.fisher_mean(di_block=data)
```
Now we rotate the data to the mean declination and inclination using **pmag.dodirot()**. But first, a little help message would be great.
```
help(pmag.dodirot_V)
rotdata=pmag.dodirot_V(data,1.8,58)
ipmag.plot_net(1)
ipmag.plot_di(di_block=rotdata)
```
It looks like the inclinations are spread out too much. Let's see what **ipmag.fishqq()** has to say.
```
ipmag.fishqq(di_block=rotdata)
```
And sure enough... the inclinations are too spread out. They are not exponentially distributed.
## Problem 2
The best way to do a foldtest in a Jupyter Notebook is to use **ipmag.bootstrap_fold_test( )**:
```
help(ipmag.bootstrap_fold_test)
```
So we first have to read in the data and make sure it is in a suitable array, then call **ipmag.bootstrap_fold_test( )**.
```
fold_data=np.loadtxt('Chapter_12/ps12-2.dat')
fold_data
ipmag.bootstrap_fold_test(fold_data)
```
These data are much better grouped in geographic coordinates and much worse after tilt correction. So these were magnetized after tilting.
## Problem 3a
We know what to do here:
```
di_block=np.loadtxt('Chapter_12/ps12-3.dat')# read in data
ipmag.plot_net(1) # plot the equal area net
ipmag.plot_di(di_block=di_block,title="Problem 12.3 directions") # put on the directions
```
## Problem 3b
To separate by polarity, one could just sort by inclination and put all the negative ones in one group and all the positive ones in the other. But this is dangerous for low inclination data (because you could easily have negative inclinations pointing north). A more general approach (which would allow southward directed declinations with positive inclinations, for example), would be to read the data into a Pandas DataFrame, calculate the principal direction of the entire dataset using **pmag.doprinc()** and calculate the directions rotated to that reference. Then all positive inclinations would be one polarity (say, normal) and the other would all be the reverse polarity. So, here goes.
```
help(pmag.doprinc)
# calculate the principle direction for the data set
principal=pmag.doprinc(di_block)
print ('Principal direction declination: ' + '%7.1f'%(principal['dec']))
print ('Principal direction inclination: ' + '%7.1f'%(principal['inc']))
```
Now we can use some nice **Pandas** functionality to assign polarity:
```
import pandas as pd
# make a dataframe
df=pd.DataFrame(di_block)
df.columns=['dec','inc']
# make a column with the principal dec and inc
df['principal_dec'] = principal['dec'] # assign these to the dataframe
df['principal_inc'] = principal['inc']
# make a principal block for comparison
principal_block=df[['principal_dec','principal_inc']].values
# get the angle from each data point to the principal direction
df['angle'] = pmag.angle(di_block,principal_block)
# assign polarity
df.loc[df.angle>90,'polarity'] = 'Reverse'
df.loc[df.angle<=90,'polarity'] = 'Normal'
```
Now that polarity is assigned using the angle from the principal component, let's filter the data by polarity and then put the data back into arrays for calculating stuff.
```
normal_data = df[df.polarity=='Normal'].reset_index(drop=True)
reverse_data = df[df.polarity=='Reverse'].reset_index(drop=True)
NormBlock=np.array([normal_data["dec"],normal_data["inc"]]).transpose()
RevBlock=np.array([reverse_data["dec"],reverse_data["inc"]]).transpose()
help(pmag.fisher_mean)
norm_fpars= pmag.fisher_mean(NormBlock)
print ('Mean normal declination: ' + '%7.1f'%(norm_fpars['dec']))
print ('Mean normal inclination: ' + '%7.1f'%(norm_fpars['inc']))
rev_fpars= pmag.fisher_mean(RevBlock)
print ('Mean reverse declination: ' + '%7.1f'%(rev_fpars['dec']))
print ('Mean reverse inclination: ' + '%7.1f'%(rev_fpars['inc']))
```
Now let's check if the data are Fisher distributed using our old friend **ipmag.fishqq( )**
```
ipmag.fishqq(di_block=di_block)
```
The uniform null hypothesis fails at the 95\% confidence level for the normal data. So lets try **pmag.dobingham()** and **pmag.dokent( )** for the whole dataset and the norm and reverse data, respectively. Note that dokent has a different syntax.
```
norm_kpars=pmag.dokent(NormBlock,len(NormBlock))
print (norm_kpars)
rev_kpars=pmag.dokent(RevBlock,len(RevBlock))
print (rev_kpars)
bpars=pmag.dobingham(di_block)
print (bpars)
```
And finally the bootstrapped means:
```
help(pmag.di_boot)
BnDIs=pmag.di_boot(NormBlock)
BrDIs=pmag.di_boot(RevBlock)
norm_boot_kpars=pmag.dokent(BnDIs,NN=1)
rev_boot_kpars=pmag.dokent(BrDIs,NN=1)
```
And now for the plots, starting with the Fisher confidence ellipses using **ipmag.plot_di_mean**.
```
help(ipmag.plot_di_mean)
ipmag.plot_net(1)
ipmag.plot_di(di_block=di_block)
ipmag.plot_di_mean(dec=norm_fpars['dec'],inc=norm_fpars['inc'],a95=norm_fpars['alpha95'],\
marker='*',color='blue',markersize=50)
ipmag.plot_di_mean(dec=rev_fpars['dec'],inc=rev_fpars['inc'],a95=rev_fpars['alpha95'],\
marker='*',color='blue',markersize=50)
```
The other ellipses get plotted with a different function, **ipmag.plot_di_mean_ellipse**.
```
help(ipmag.plot_di_mean_ellipse)
```
Bingham:
```
ipmag.plot_net(1)
ipmag.plot_di(di_block=di_block)
ipmag.plot_di_mean_ellipse(bpars,marker='*',color='cyan',markersize=20)
```
Kent:
```
ipmag.plot_net(1)
ipmag.plot_di(di_block=di_block)
ipmag.plot_di_mean_ellipse(norm_kpars,marker='*',color='cyan',markersize=20)
ipmag.plot_di_mean_ellipse(rev_kpars,marker='*',color='cyan',markersize=20)
```
And bootstrapped
```
ipmag.plot_net(1)
ipmag.plot_di(di_block=di_block)
ipmag.plot_di_mean_ellipse(norm_boot_kpars,marker='*',color='cyan',markersize=20)
ipmag.plot_di_mean_ellipse(rev_boot_kpars,marker='*',color='cyan',markersize=20)
```
The Kent ellipses do a pretty good job - as well as the bootstrapped ones.
## Problem 3c
For this problem, we have to flip the reverse mode to the antipodes and then try the Watson's V test.
```
help(pmag.flip)
norms,rev_antis=pmag.flip(di_block)
help(ipmag.common_mean_watson)
ipmag.common_mean_watson(norms,rev_antis,plot='yes')
```
Accoding to Dr. Watson, these two modes are not drawn from the same distribution (according to watson's Vw criterion), so they fail the reversals test. Now let's try the function **ipmag.reversal_test_bootstrap** instead.
```
help(ipmag.reversal_test_bootstrap)
ipmag.reversal_test_bootstrap(di_block=di_block)
```
The Y components are significantly different in the reverse mode (after flipping) with respect to the normal mode. Therefore, this data set fails the reversals test. And you should definately NOT use Bingham statistics on this data set!
| github_jupyter |
<a href="https://colab.research.google.com/github/Dewanik/RNN_from_scratch/blob/master/Recurrent_Neural_Network.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Reccurent Neural Network from scratch**
**Understanding the mathematics behind RNNs**
Importing Numpy for matrix math
```
import copy, numpy as np
```
**Reading Data From Kafka.txt creating a dictinoary map of characters to integer(or their position) in total number of unique characters in data and
and integer(or their position) in total number of data to its accosiated characters and one_hot_encoding of the character**
```
data = open('kafka.txt','r').read()
chars = list(set(data))
n = 2
data_size , unique_size = len(data),len(chars)
char_ix = {ch:i for i,ch in enumerate(chars)}
ix_char = {i:ch for i,ch in enumerate(chars)}
```
Weight matrix Initialization for :
1. Input to Current Hidden Layer
2. Current Hidden to Previous Hidden Layer
3. Merged Current Hidden to output
and biases
```
hidden_size = 100
seq_length = 500
wxh = np.random.randn(hidden_size, len(chars)) * 0.01
whh = np.random.randn(hidden_size, hidden_size) * 0.01
why = np.random.randn(len(chars), hidden_size) * 0.01 #input to hidden
bh = np.zeros((hidden_size,1))
by = np.zeros((len(chars),1))
print(wxh.shape)
```
Example of One_Hot_Encoding Of Character to Vectors
```
vector_for_a = np.zeros((len(chars),1))
vector_for_a[char_ix['a']] = 1
```
Feed Forward of RNN
Creating a dataset of the position of characters mapping it to next upcoming character in kafka.txt dataset
```
def train(x,y,hprev):
xs , hs , ys , ps = {}, {} , {} , {}
hs[-1] = np.copy(hprev)
loss = 0
#Forward Propagation
for i in range(seq_length):
xs[i] = np.zeros((len(chars),1))
xs[i][x[i]] = 1
hs[i] = np.tanh(np.dot(wxh,xs[i]) + np.dot(whh,hs[i-1]) + bh)
ys[i] = np.dot(why,hs[i])+ by
ps[i] = np.exp(ys[i])/np.sum(np.exp(ys[i]))
loss += -np.log(ps[i][y[i]])
#Back Propagation Through Time (BPTT)
dwxh , dwhh , dwhy , dbh , dby = np.zeros_like(wxh) , np.zeros_like(whh) , np.zeros_like(why) , np.zeros_like(bh) ,np.zeros_like(by)
dhnext = np.zeros_like(hs[0])
for i in reversed(range(len(x))):
#Updating Gradients calculating small gradients at every time step
dy = np.copy(ps[i])
dy[y[i]] -= 1
dwhy += np.dot(dy,hs[i].T)
dby += dy
dh = np.dot(why.T,dy) + dhnext
dhraw = (1 - hs[i] * hs[i]) * dh
dbh += dhraw
dwxh += np.dot(dhraw,xs[i].T)
dwhh += np.dot(dhraw,hs[i-1].T)
dhnext = np.dot(whh.T,dhraw)
for dparam in [dwxh, dwhh, dwhy, dbh, dby]:
np.clip(dparam, -5, 5, out=dparam) # clip to mitigate exploding gradients
return loss, dwxh, dwhh, dwhy, dbh, dby, hs[seq_length-1]
def predict(seed_x,h):
x = np.zeros((len(chars),1))
x[seed_x] = 1
vec_outputs = list()
yguess = list()
for i in range(seq_length):
#Input * whx + prev_hidden * whh + bh ==> new_merged_hidden
h = np.tanh(np.dot(wxh,x) + np.dot(whh,h) + bh)
#new_merged_hidden * why + by ==> yhat
y = np.dot(why,h) + by
#normalized(yhat) ==> normalized_output
p = np.exp(y)/np.sum(np.exp(y))
#new_vecx == normalized_output == new_x
#Choose with highest probability
ix = np.argmax(p)
x = np.zeros((len(chars),1))
x[ix] = 1
yguess.append(ix)
print(''.join([ix_char[c] for c in yguess]))
x = [char_ix[c] for c in data[0:seq_length]]
hprev = np.zeros((hidden_size,1))
predict(x[0],hprev)
x = [char_ix[c] for c in data[0:seq_length]]
y = [char_ix[c] for c in data[1:seq_length+1]]
print('Given X:')
print(x[0])
print(ix_char[x[0]])
print('Known Y:')
print(y)
print(len(y))
print(''.join([ix_char[c] for c in y]))
print('\n')
nses , pses = 0,0
lr = 1e-1
mWxh, mWhh, mWhy = np.zeros_like(wxh), np.zeros_like(whh), np.zeros_like(why)
mbh, mby = np.zeros_like(bh), np.zeros_like(by) # memory variables for Adagrad
smooth_loss = -np.log(1.0/len(chars))*seq_length # loss at iteration 0
for i in range(1000*100):
if pses+seq_length+1 >= len(data) or nses == 0:
hprev = np.zeros((hidden_size,1)) # reset RNN memory
pses = 0 # go from start of data
x = [char_ix[c] for c in data[0:seq_length]]
y = [char_ix[c] for c in data[1:seq_length+1]]
loss, dwxh, dwhh, dwhy, dbh, dby, hprev = train(x,y,hprev)
smooth_loss = smooth_loss * 0.999 + loss * 0.001
if i % 1000 == 0:
predict(x[0],hprev)
print('Iter:{1} Loss:{0}'.format(smooth_loss,i))
for param, dparam, mem in zip([wxh, whh, why, bh, by],
[dwxh, dwhh, dwhy, dbh, dby],
[mWxh, mWhh, mWhy, mbh, mby]):
mem += dparam * dparam
param += -lr * dparam / np.sqrt(mem + 1e-8) # adagrad update
length = 600
x = [char_ix[c] for c in data[506:seq_length]]
y = [char_ix[c] for c in data[507:seq_length+1]]
print(ix_char[x[1]])
print(y)
hprev = np.zeros((hidden_size,1))
predict(x[1],hprev)
```
| github_jupyter |
```
%matplotlib inline
```
# Fitting TESS FFIs
```
%run notebook_setup
```
This tutorial is nearly identical to the :ref:`tess` tutorial with added support for the TESS full frame images using [tesscut](https://mast.stsci.edu/tesscut/).
First, we query the TESS input catalog for the coordinates and properties of this source:
```
import numpy as np
from astroquery.mast import Catalogs
ticid = 49899799
tic = Catalogs.query_object("TIC {0}".format(ticid), radius=0.2, catalog="TIC")
star = tic[np.argmin(tic["dstArcSec"])]
tic_mass = float(star["mass"]), float(star["e_mass"])
tic_radius = float(star["rad"]), float(star["e_rad"])
```
Then we download the data from tesscut.
This is similar to what we did in the :ref:`tess` tutorial, but we need to do some background subtraction because the pipeline doesn't seem to do too well for the official TESS FFIs.
```
from io import BytesIO
from zipfile import ZipFile
from astropy.io import fits
from astropy.utils.data import download_file
# Download the cutout
url = "https://mast.stsci.edu/tesscut/api/v0.1/astrocut?ra={0[ra]}&dec={0[dec]}&y=15&x=15&units=px§or=All".format(star)
fn = download_file(url, cache=True)
with ZipFile(fn, "r") as f:
with fits.open(BytesIO(f.read(f.namelist()[0]))) as hdus:
tpf = hdus[1].data
tpf_hdr = hdus[1].header
texp = tpf_hdr["FRAMETIM"] * tpf_hdr["NUM_FRM"]
texp /= 60.0 * 60.0 * 24.0
time = tpf["TIME"]
flux = tpf["FLUX"]
m = np.any(np.isfinite(flux), axis=(1, 2)) & (tpf["QUALITY"] == 0)
ref_time = 0.5 * (np.min(time[m])+np.max(time[m]))
time = np.ascontiguousarray(time[m] - ref_time, dtype=np.float64)
flux = np.ascontiguousarray(flux[m], dtype=np.float64)
# Compute the median image
mean_img = np.median(flux, axis=0)
# Sort the pixels by median brightness
order = np.argsort(mean_img.flatten())[::-1]
# Choose a mask for the background
bkg_mask = np.zeros_like(mean_img, dtype=bool)
bkg_mask[np.unravel_index(order[-100:], mean_img.shape)] = True
flux -= np.median(flux[:, bkg_mask], axis=-1)[:, None, None]
```
Everything below this line is the same as the :ref:`tess` tutorial.
```
import matplotlib.pyplot as plt
from scipy.signal import savgol_filter
# A function to estimate the windowed scatter in a lightcurve
def estimate_scatter_with_mask(mask):
f = np.sum(flux[:, mask], axis=-1)
smooth = savgol_filter(f, 1001, polyorder=5)
return 1e6 * np.sqrt(np.median((f / smooth - 1)**2))
# Loop over pixels ordered by brightness and add them one-by-one
# to the aperture
masks, scatters = [], []
for i in range(1, 100):
msk = np.zeros_like(mean_img, dtype=bool)
msk[np.unravel_index(order[:i], mean_img.shape)] = True
scatter = estimate_scatter_with_mask(msk)
masks.append(msk)
scatters.append(scatter)
# Choose the aperture that minimizes the scatter
pix_mask = masks[np.argmin(scatters)]
# Plot the selected aperture
plt.imshow(mean_img.T, cmap="gray_r")
plt.imshow(pix_mask.T, cmap="Reds", alpha=0.3)
plt.title("selected aperture")
plt.xticks([])
plt.yticks([]);
plt.figure(figsize=(10, 5))
sap_flux = np.sum(flux[:, pix_mask], axis=-1)
sap_flux = (sap_flux / np.median(sap_flux) - 1) * 1e3
plt.plot(time, sap_flux, "k")
plt.xlabel("time [days]")
plt.ylabel("relative flux [ppt]")
plt.title("raw light curve")
plt.xlim(time.min(), time.max());
# Build the first order PLD basis
X_pld = np.reshape(flux[:, pix_mask], (len(flux), -1))
X_pld = X_pld / np.sum(flux[:, pix_mask], axis=-1)[:, None]
# Build the second order PLD basis and run PCA to reduce the number of dimensions
X2_pld = np.reshape(X_pld[:, None, :] * X_pld[:, :, None], (len(flux), -1))
U, _, _ = np.linalg.svd(X2_pld, full_matrices=False)
X2_pld = U[:, :X_pld.shape[1]]
# Construct the design matrix and fit for the PLD model
X_pld = np.concatenate((np.ones((len(flux), 1)), X_pld, X2_pld), axis=-1)
XTX = np.dot(X_pld.T, X_pld)
w_pld = np.linalg.solve(XTX, np.dot(X_pld.T, sap_flux))
pld_flux = np.dot(X_pld, w_pld)
# Plot the de-trended light curve
plt.figure(figsize=(10, 5))
plt.plot(time, sap_flux-pld_flux, "k")
plt.xlabel("time [days]")
plt.ylabel("de-trended flux [ppt]")
plt.title("initial de-trended light curve")
plt.xlim(time.min(), time.max());
from astropy.stats import BoxLeastSquares
period_grid = np.exp(np.linspace(np.log(1), np.log(15), 50000))
bls = BoxLeastSquares(time, sap_flux - pld_flux)
bls_power = bls.power(period_grid, 0.1, oversample=20)
# Save the highest peak as the planet candidate
index = np.argmax(bls_power.power)
bls_period = bls_power.period[index]
bls_t0 = bls_power.transit_time[index]
bls_depth = bls_power.depth[index]
transit_mask = bls.transit_mask(time, bls_period, 0.2, bls_t0)
fig, axes = plt.subplots(2, 1, figsize=(10, 10))
# Plot the periodogram
ax = axes[0]
ax.axvline(np.log10(bls_period), color="C1", lw=5, alpha=0.8)
ax.plot(np.log10(bls_power.period), bls_power.power, "k")
ax.annotate("period = {0:.4f} d".format(bls_period),
(0, 1), xycoords="axes fraction",
xytext=(5, -5), textcoords="offset points",
va="top", ha="left", fontsize=12)
ax.set_ylabel("bls power")
ax.set_yticks([])
ax.set_xlim(np.log10(period_grid.min()), np.log10(period_grid.max()))
ax.set_xlabel("log10(period)")
# Plot the folded transit
ax = axes[1]
x_fold = (time - bls_t0 + 0.5*bls_period)%bls_period - 0.5*bls_period
m = np.abs(x_fold) < 0.4
ax.plot(x_fold[m], sap_flux[m] - pld_flux[m], ".k")
# Overplot the phase binned light curve
bins = np.linspace(-0.41, 0.41, 32)
denom, _ = np.histogram(x_fold, bins)
num, _ = np.histogram(x_fold, bins, weights=sap_flux - pld_flux)
denom[num == 0] = 1.0
ax.plot(0.5*(bins[1:] + bins[:-1]), num / denom, color="C1")
ax.set_xlim(-0.3, 0.3)
ax.set_ylabel("de-trended flux [ppt]")
ax.set_xlabel("time since transit");
m = ~transit_mask
XTX = np.dot(X_pld[m].T, X_pld[m])
w_pld = np.linalg.solve(XTX, np.dot(X_pld[m].T, sap_flux[m]))
pld_flux = np.dot(X_pld, w_pld)
x = np.ascontiguousarray(time, dtype=np.float64)
y = np.ascontiguousarray(sap_flux-pld_flux, dtype=np.float64)
plt.figure(figsize=(10, 5))
plt.plot(time, y, "k")
plt.xlabel("time [days]")
plt.ylabel("de-trended flux [ppt]")
plt.title("final de-trended light curve")
plt.xlim(time.min(), time.max());
import exoplanet as xo
import pymc3 as pm
import theano.tensor as tt
def build_model(mask=None, start=None):
if mask is None:
mask = np.ones(len(x), dtype=bool)
with pm.Model() as model:
# Parameters for the stellar properties
mean = pm.Normal("mean", mu=0.0, sd=10.0)
u_star = xo.distributions.QuadLimbDark("u_star")
# Stellar parameters from TIC
M_star_huang = 1.094, 0.039
R_star_huang = 1.10, 0.023
m_star = pm.Normal("m_star", mu=tic_mass[0], sd=tic_mass[1])
r_star = pm.Normal("r_star", mu=tic_radius[0], sd=tic_radius[1])
# Prior to require physical parameters
pm.Potential("m_star_prior", tt.switch(m_star > 0, 0, -np.inf))
pm.Potential("r_star_prior", tt.switch(r_star > 0, 0, -np.inf))
# Orbital parameters for the planets
logP = pm.Normal("logP", mu=np.log(bls_period), sd=1)
t0 = pm.Normal("t0", mu=bls_t0, sd=1)
b = pm.Uniform("b", lower=0, upper=1, testval=0.5)
logr = pm.Normal("logr", sd=1.0,
mu=0.5*np.log(1e-3*np.array(bls_depth))+np.log(tic_radius[0]))
r_pl = pm.Deterministic("r_pl", tt.exp(logr))
ror = pm.Deterministic("ror", r_pl / r_star)
# This is the eccentricity prior from Kipping (2013):
# https://arxiv.org/abs/1306.4982
ecc = pm.Beta("ecc", alpha=0.867, beta=3.03, testval=0.1)
omega = xo.distributions.Angle("omega")
# Transit jitter & GP parameters
logs2 = pm.Normal("logs2", mu=np.log(np.var(y[mask])), sd=10)
logw0_guess = np.log(2*np.pi/10)
logw0 = pm.Normal("logw0", mu=logw0_guess, sd=10)
# We'll parameterize using the maximum power (S_0 * w_0^4) instead of
# S_0 directly because this removes some of the degeneracies between
# S_0 and omega_0
logpower = pm.Normal("logpower",
mu=np.log(np.var(y[mask]))+4*logw0_guess,
sd=10)
logS0 = pm.Deterministic("logS0", logpower - 4 * logw0)
# Tracking planet parameters
period = pm.Deterministic("period", tt.exp(logP))
# Orbit model
orbit = xo.orbits.KeplerianOrbit(
r_star=r_star, m_star=m_star,
period=period, t0=t0, b=b,
ecc=ecc, omega=omega)
pm.Deterministic("a", orbit.a_planet)
pm.Deterministic("incl", orbit.incl)
# Compute the model light curve using starry
light_curves = xo.StarryLightCurve(u_star).get_light_curve(
orbit=orbit, r=r_pl, t=x[mask], texp=texp)*1e3
light_curve = pm.math.sum(light_curves, axis=-1) + mean
pm.Deterministic("light_curves", light_curves)
# GP model for the light curve
kernel = xo.gp.terms.SHOTerm(log_S0=logS0, log_w0=logw0, Q=1/np.sqrt(2))
gp = xo.gp.GP(kernel, x[mask], tt.exp(logs2) + tt.zeros(mask.sum()), J=2)
pm.Potential("transit_obs", gp.log_likelihood(y[mask] - light_curve))
pm.Deterministic("gp_pred", gp.predict())
# Fit for the maximum a posteriori parameters, I've found that I can get
# a better solution by trying different combinations of parameters in turn
if start is None:
start = model.test_point
map_soln = xo.optimize(start=start, vars=[b])
map_soln = xo.optimize(start=map_soln, vars=[logs2, logpower, logw0])
map_soln = xo.optimize(start=map_soln, vars=[logr])
map_soln = xo.optimize(start=map_soln)
return model, map_soln
model0, map_soln0 = build_model()
def plot_light_curve(soln, mask=None):
if mask is None:
mask = np.ones(len(x), dtype=bool)
fig, axes = plt.subplots(3, 1, figsize=(10, 7), sharex=True)
ax = axes[0]
ax.plot(x[mask], y[mask], "k", label="data")
gp_mod = soln["gp_pred"] + soln["mean"]
ax.plot(x[mask], gp_mod, color="C2", label="gp model")
ax.legend(fontsize=10)
ax.set_ylabel("relative flux [ppt]")
ax = axes[1]
ax.plot(x[mask], y[mask] - gp_mod, "k", label="de-trended data")
for i, l in enumerate("b"):
mod = soln["light_curves"][:, i]
ax.plot(x[mask], mod, label="planet {0}".format(l))
ax.legend(fontsize=10, loc=3)
ax.set_ylabel("de-trended flux [ppt]")
ax = axes[2]
mod = gp_mod + np.sum(soln["light_curves"], axis=-1)
ax.plot(x[mask], y[mask] - mod, "k")
ax.axhline(0, color="#aaaaaa", lw=1)
ax.set_ylabel("residuals [ppt]")
ax.set_xlim(x[mask].min(), x[mask].max())
ax.set_xlabel("time [days]")
return fig
plot_light_curve(map_soln0);
mod = map_soln0["gp_pred"] + map_soln0["mean"] + np.sum(map_soln0["light_curves"], axis=-1)
resid = y - mod
rms = np.sqrt(np.median(resid**2))
mask = np.abs(resid) < 5 * rms
plt.figure(figsize=(10, 5))
plt.plot(x, resid, "k", label="data")
plt.plot(x[~mask], resid[~mask], "xr", label="outliers")
plt.axhline(0, color="#aaaaaa", lw=1)
plt.ylabel("residuals [ppt]")
plt.xlabel("time [days]")
plt.legend(fontsize=12, loc=3)
plt.xlim(x.min(), x.max());
model, map_soln = build_model(mask, map_soln0)
plot_light_curve(map_soln, mask);
np.random.seed(12345)
sampler = xo.PyMC3Sampler(window=100, start=300, finish=500)
with model:
burnin = sampler.tune(tune=3500, start=map_soln,
step_kwargs=dict(target_accept=0.9),
chains=4)
with model:
trace = sampler.sample(draws=1000, chains=4)
pm.summary(trace, varnames=["logw0", "logS0", "logs2", "omega", "ecc", "r_pl", "b", "t0", "logP", "r_star", "m_star", "u_star", "mean"])
# Compute the GP prediction
gp_mod = np.median(trace["gp_pred"] + trace["mean"][:, None], axis=0)
# Get the posterior median orbital parameters
p = np.median(trace["period"])
t0 = np.median(trace["t0"])
# Plot the folded data
x_fold = (x[mask] - t0 + 0.5*p) % p - 0.5*p
plt.plot(x_fold, y[mask] - gp_mod, ".k", label="data", zorder=-1000)
# Plot the folded model
inds = np.argsort(x_fold)
inds = inds[np.abs(x_fold)[inds] < 0.3]
pred = trace["light_curves"][:, inds, 0]
pred = np.percentile(pred, [16, 50, 84], axis=0)
plt.plot(x_fold[inds], pred[1], color="C1", label="model")
art = plt.fill_between(x_fold[inds], pred[0], pred[2], color="C1", alpha=0.5,
zorder=1000)
art.set_edgecolor("none")
# Annotate the plot with the planet's period
txt = "period = {0:.5f} +/- {1:.5f} d".format(
np.mean(trace["period"]), np.std(trace["period"]))
plt.annotate(txt, (0, 0), xycoords="axes fraction",
xytext=(5, 5), textcoords="offset points",
ha="left", va="bottom", fontsize=12)
plt.legend(fontsize=10, loc=4)
plt.xlim(-0.5*p, 0.5*p)
plt.xlabel("time since transit [days]")
plt.ylabel("de-trended flux")
plt.xlim(-0.2, 0.2);
import corner
import astropy.units as u
varnames = ["period", "b", "ecc", "r_pl"]
samples = pm.trace_to_dataframe(trace, varnames=varnames)
# Convert the radius to Earth radii
samples["r_pl"] = (np.array(samples["r_pl"]) * u.R_sun).to(u.R_jupiter).value
corner.corner(
samples[["period", "r_pl", "b", "ecc"]],
labels=["period [days]", "radius [Jupiter radii]", "impact param", "eccentricity"]);
aor = -trace["a"] / trace["r_star"]
e = trace["ecc"]
w = trace["omega"]
i = trace["incl"]
b = trace["b"]
k = trace["r_pl"] / trace["r_star"]
P = trace["period"]
T_tot = P/np.pi * np.arcsin(np.sqrt(1 - b**2) / np.sin(i) / aor)
dur = T_tot * np.sqrt(1 - e**2) / (1 + e * np.sin(w))
samples = pm.trace_to_dataframe(trace, varnames=["r_pl", "t0", "b", "ecc", "omega"])
samples["duration"] = dur
corner.corner(samples);
```
| github_jupyter |
# Imputação de valores faltantes - Experimento
Este é um componente para imputação de valores faltantes usando média, mediana ou mais frequente. Faz uso da implementação do [Scikit-learn](https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html#sklearn.impute.SimpleImputer). <br>
Scikit-learn é uma biblioteca open source de machine learning que suporta apredizado supervisionado e não supervisionado. Também provê várias ferramentas para montagem de modelo, pré-processamento de dados, seleção e avaliação de modelos, e muitos outros utilitários.
## Declaração de parâmetros e hiperparâmetros
Declare parâmetros com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKBqlpGMRcVMdkUMvKID3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMOEYuFFpZbmBUNlXiKOKyoGuULGZcVzluc1VKFNe7JXxjMacsprtMcRAyLiCMBETIq2EAJFiK0aqSYSNJ+1MM/4PgT5JLJtQFGjnmUoUJy/OB/8LtbMz854SYFo0D7i21/DAOBXaBete3vY9uunwD+Z+BKa/rLNWDmk/RqUwsfAb3bwMV1U5P3gMsdoP9JlwzJkfw0hXweeD+jb8oCfbdA16rbW2Mfpw9AmrpaugEODoGRAmWveby7s7W3f880+vsBocZyukMJsmwAAAAGYktHRAD/AP8A/6C9p5MAAAAJcEhZcwAADdcAAA3XAUIom3gAAAAHdElNRQfkBgsMIwnXL7c0AAACDUlEQVQ4y92UP4gTQRTGf29zJxhJZ2NxbMBKziYWlmJ/ile44Nlkd+dIYWFzItiNgoIEtFaTzF5Ac/inE/urtLWxsMqmUOwCEpt1Zmw2xxKi53XitPO9H9978+aDf/3IUQvSNG0450Yi0jXG7C/eB0cFeu9viciGiDyNoqh2KFBrHSilWstgnU7nFLBTgl+ur6/7PwK11kGe5z3n3Hul1MaiuCgKDZwALHA7z/Oe1jpYCtRaB+PxuA8kQM1aW68Kt7e3zwBp6a5b1ibj8bhfhQYVZwMRiQHrvW9nWfaqCrTWPgRWvPdvsiy7IyLXgEJE4slk8nw+T5nDgDbwE9gyxryuwpRSF5xz+0BhrT07HA4/AyRJchUYASvAbhiGaRVWLIMBYq3tAojIszkMoNRulbXtPM8HwV/sXSQi54HvQRDcO0wfhGGYArvAKjAq2wAgiqJj3vsHpbtur9f7Vi2utLx60LLW2hljEuBJOYu9OI6vAzQajRvAaeBLURSPlsBelA+VhWGYaq3dwaZvbm6+m06noYicE5ErrVbrK3AXqHvvd4bD4Ye5No7jSERGwKr3Pms2m0pr7Rb30DWbTQWYcnFvAieBT7PZbFB1V6vVfpQaU4UtDQetdTCZTC557/eA48BlY8zbRZ1SqrW2tvaxCvtt2iRJ0i9/xb4x5uJRwmNlaaaJ3AfqIvKY/+78Av++6uiSZhYMAAAAAElFTkSuQmCC" /> na barra de ferramentas.<br>
A variável `dataset` possui o caminho para leitura do arquivos importados na tarefa de "Upload de dados".<br>
Você também pode importar arquivos com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKBqlpGMRcVMdkUMvKID3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMOEYuFFpZbmBUNlXiKOKyoGuULGZcVzluc1VKFNe7JXxjMacsprtMcRAyLiCMBETIq2EAJFiK0aqSYSNJ+1MM/4PgT5JLJtQFGjnmUoUJy/OB/8LtbMz854SYFo0D7i21/DAOBXaBete3vY9uunwD+Z+BKa/rLNWDmk/RqUwsfAb3bwMV1U5P3gMsdoP9JlwzJkfw0hXweeD+jb8oCfbdA16rbW2Mfpw9AmrpaugEODoGRAmWveby7s7W3f880+vsBocZyukMJsmwAAAAGYktHRAD/AP8A/6C9p5MAAAAJcEhZcwAADdcAAA3XAUIom3gAAAAHdElNRQfkBgsOBy6ASTeXAAAC/0lEQVQ4y5WUT2gcdRTHP29m99B23Uiq6dZisgoWCxVJW0oL9dqLfyhCvGWY2YUBI95MsXgwFISirQcLhS5hfgk5CF3wJIhFI7aHNsL2VFZFik1jS1qkiZKdTTKZ3/MyDWuz0fQLc/m99/vMvDfv+4RMlUrlkKqeAAaBAWAP8DSgwJ/AXRG5rao/WWsvTU5O3qKLBMD3fSMiPluXFZEPoyj67PGAMzw83PeEMABHVT/oGpiamnoAmCcEWhH5tFsgF4bh9oWFhfeKxeJ5a+0JVT0oImWgBPQCKfAQuAvcBq67rltX1b+6ApMkKRcKhe9V9QLwbavV+qRer692Sx4ZGSnEcXw0TdP3gSrQswGYz+d/S5IkVtXTwOlCoZAGQXAfmAdagAvsAErtdnuXiDy6+023l7qNRsMODg5+CawBzwB9wFPA7mx8ns/KL2Tl3xCRz5eWlkabzebahrHxPG+v4zgnc7ncufHx8Z+Hhoa29fT0lNM03Q30ikiqqg+ttX/EcTy3WTvWgdVqtddaOw/kgXvADHBHROZVNRaRvKruUNU+EdkPfGWM+WJTYOaSt1T1LPDS/4zLWWPMaLVaPWytrYvIaBRFl/4F9H2/JCKvGmMu+76/X0QOqGoZKDmOs1NV28AicMsYc97zvFdc1/0hG6kEeNsY83UnsCwivwM3VfU7YEZE7lhr74tIK8tbnJiYWPY8b6/ruleAXR0ftQy8boyZXi85CIIICDYpc2ZgYODY3NzcHmvt1eyvP64lETkeRdE1yZyixWLx5U2c8q4x5mIQBE1g33/0d3FlZeXFR06ZttZesNZejuO4q1NE5CPgWVV9E3ij47wB1IDlJEn+ljAM86urq7+KyAtZTgqsO0VV247jnOnv7/9xbGzMViqVMVX9uANYj6LonfVtU6vVkjRNj6jqGeCXzGrPAQeA10TkuKpOz87ONrayhnIA2Qo7BZwKw3B7kiRloKSqO13Xja21C47jPNgysFO1Wi0GmtmzQap6DWgD24A1Vb3SGf8Hfstmz1CuXEIAAAAASUVORK5CYII=" /> na barra de ferramentas.
```
# parâmetros
dataset = "/tmp/data/iris.csv" #@param {type:"string"}
target = None #@param {type:"feature", label:"Atributo alvo", description: "Esse valor será utilizado para garantir que o alvo não seja removido."}
strategy_num = "mean" #@param ["mean", "median", "most_frequent", "constant"] {type:"string", label:"Estratégia de atribuição de valores para colunas numéricas", description:"Os valores 'mean' e 'median' são utilizados apenas para atributos numéricos,'most_frequent' é para atributos numéricos e categóricos e 'constant' substituirá os valores ausentes por fill_value"}
strategy_cat = "most_frequent" #@param ["most_frequent", "constant"] {type:"string", label:"Estratégia de atribuição de valores para colunas categóricas", description:"O valor 'most_frequent' é para atributos numéricos e categóricos e 'constant' substituirá os valores ausentes por fill_value"}
fillvalue_num = 0 #@param {type:"number", label:"Valor de preenchimento para colunas numéricas (utilizado apenas com estratégia 'constant')", description:"Valor a ser preenchido no lugar de valores nulos"}
fillvalue_cat = "" #@param {type:"string", label:"Valor de preenchimento para colunas categóricas (utilizado apenas com estratégia 'constant')", description:"Valor a ser preenchido no lugar de valores nulos"}
```
## Acesso ao conjunto de dados
O conjunto de dados utilizado nesta etapa será o mesmo carregado através da plataforma.<br>
O tipo da variável retornada depende do arquivo de origem:
- [pandas.DataFrame](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html) para CSV e compressed CSV: .csv .csv.zip .csv.gz .csv.bz2 .csv.xz
- [Binary IO stream](https://docs.python.org/3/library/io.html#binary-i-o) para outros tipos de arquivo: .jpg .wav .zip .h5 .parquet etc
```
import pandas as pd
df = pd.read_csv(dataset)
has_target = True if target is not None and target in df.columns else False
X = df.copy()
if has_target:
X = df.drop(target, axis=1)
y = df[target]
```
## Acesso aos metadados do conjunto de dados
Utiliza a função `stat_dataset` do [SDK da PlatIAgro](https://platiagro.github.io/sdk/) para carregar metadados. <br>
Por exemplo, arquivos CSV possuem `metadata['featuretypes']` para cada coluna no conjunto de dados (ex: categorical, numerical, or datetime).
```
import numpy as np
from platiagro import stat_dataset
metadata = stat_dataset(name=dataset)
featuretypes = metadata["featuretypes"]
columns = df.columns.to_numpy()
featuretypes = np.array(featuretypes)
if has_target:
target_index = np.argwhere(columns == target)
columns = np.delete(columns, target_index)
featuretypes = np.delete(featuretypes, target_index)
```
## Configuração das features
```
from platiagro.featuretypes import NUMERICAL
# Selects the indexes of numerical
numerical_indexes = np.where(featuretypes == NUMERICAL)[0]
non_numerical_indexes = np.where(~(featuretypes == NUMERICAL))[0]
```
## Treina um modelo usando sklearn.impute.SimpleImputer
```
# from platiagro.featuretypes import NUMERICAL
from sklearn.compose import make_column_transformer
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
pipeline = make_pipeline(
make_column_transformer(
(
SimpleImputer(strategy=strategy_num, fill_value=fillvalue_num),
numerical_indexes,
),
(
SimpleImputer(strategy=strategy_cat, fill_value=fillvalue_cat),
non_numerical_indexes,
),
remainder="passthrough",
)
)
# Train model and transform dataset
X = pipeline.fit_transform(X)
features_after_pipeline = np.concatenate(
(columns[numerical_indexes], columns[non_numerical_indexes])
)
# Put data back in a pandas.DataFrame
df = pd.DataFrame(data=X, columns=features_after_pipeline)
if has_target:
df[target] = y
```
## Cria visualização do resultado
Cria visualização do resultado como uma planilha.
```
import matplotlib.pyplot as plt
from platiagro.plotting import plot_data_table
ax = plot_data_table(df)
plt.show()
```
## Salva alterações no conjunto de dados
O conjunto de dados será salvo (e sobrescrito com as respectivas mudanças) localmente, no container da experimentação, utilizando a função `pandas.DataFrame.to_csv`.<br>
```
# save dataset changes
df.to_csv(dataset, index=False)
```
## Salva resultados da tarefa
A plataforma guarda o conteúdo de `/tmp/data/` para as tarefas subsequentes.
```
from joblib import dump
artifacts = {
"pipeline": pipeline,
"columns": columns,
"features_after_pipeline": features_after_pipeline,
}
dump(artifacts, "/tmp/data/imputer.joblib")
```
| github_jupyter |
# qecsim demos
## Simulating error correction with a planar stabilizer code
This demo shows verbosely how to simulate one error correction run.
| For normal use, the code in this demo is encapsulated in the function:
| `qecsim.app.run_once(code, error_model, decoder, error_probability)`,
| and the simulation of many error correction runs is encapsulated in the function:
| `qecsim.app.run(code, error_model, decoder, error_probability, max_runs, max_failures)`.
Notes:
* Operators can be visualised in binary symplectic form (bsf) or Pauli form, e.g. `[1 1 0|0 1 0] = XYI`.
* The binary symplectic product is denoted by $\odot$ and defined as $A \odot B \equiv A \Lambda B \bmod 2$ where $\Lambda = \left[\begin{matrix} 0 & I \\ I & 0 \end{matrix}\right]$.
* Binary addition is denoted by $\oplus$ and defined as addition modulo 2, or equivalently exclusive-or.
### Initialise the models
```
%run qsu.ipynb # color-printing functions
import numpy as np
from qecsim import paulitools as pt
from qecsim.models.generic import DepolarizingErrorModel
from qecsim.models.planar import PlanarCode, PlanarMWPMDecoder
# initialise models
my_code = PlanarCode(5, 5)
my_error_model = DepolarizingErrorModel()
my_decoder = PlanarMWPMDecoder()
# print models
print(my_code)
print(my_error_model)
print(my_decoder)
```
### Generate a random error
```
# set physical error probability to 10%
error_probability = 0.1
# seed random number generator for repeatability
rng = np.random.default_rng(59)
# error: random error based on error probability
error = my_error_model.generate(my_code, error_probability, rng)
qsu.print_pauli('error:\n{}'.format(my_code.new_pauli(error)))
```
### Evaluate the syndrome
The syndrome is a binary array indicating the stabilizers with which the error does not commute. It is calculated as $syndrome = error \odot stabilisers^T$.
```
# syndrome: stabilizers that do not commute with the error
syndrome = pt.bsp(error, my_code.stabilizers.T)
qsu.print_pauli('syndrome:\n{}'.format(my_code.ascii_art(syndrome)))
```
### Find a recovery operation
In this case, the recovery operation is found by a minimum weight perfect matching decoder that finds the recovery operation as follows:
* The syndrome is resolved to plaquettes using: `PlanarCode.syndrome_to_plaquette_indices`.
* For each plaquette, the nearest corresponding off-boundary plaquette is found using: `PlanarCode.virtual_plaquette_index`.
* A graph between plaquettes is built with weights given by: `PlanarMWPMDecoder.distance`.
* A MWPM algorithm is used to match plaquettes into pairs.
* A recovery operator is constructed by applying the shortest path between matching plaquette pairs using:
`PlanarPauli.path`.
```
# recovery: best match recovery operation based on decoder
recovery = my_decoder.decode(my_code, syndrome)
qsu.print_pauli('recovery:\n{}'.format(my_code.new_pauli(recovery)))
```
As a sanity check, we expect $recovery \oplus error$ to commute with all stabilizers, i.e. $(recovery \oplus error) \odot stabilisers^T = 0$.
```
# check recovery ^ error commutes with stabilizers (by construction)
print(pt.bsp(recovery ^ error, my_code.stabilizers.T))
```
### Visualise $recovery \oplus error$
Just out of curiosity, we can see what $recovery \oplus error$ looks like. If successful, it should be a product of stabilizer plaquette / vertex operators.
```
# print recovery ^ error (out of curiosity)
qsu.print_pauli('recovery ^ error:\n{}'.format(my_code.new_pauli(recovery ^ error)))
```
### Test if the recovery operation is successful
The recovery operation is successful iff $recovery \oplus error$ commutes with all logical operators, i.e. $(recovery \oplus error) \odot logicals^T = 0.$
```
# success iff recovery ^ error commutes with logicals
print(pt.bsp(recovery ^ error, my_code.logicals.T))
```
Note: The decoder is not guaranteed to find a successful recovery operation. The planar 5 x 5 code has distance $d = 5$ so we can only guarantee to correct errors up to weight $(d - 1)/2=2$.
### Equivalent code in single call
The above demo is equivalent to the following code.
```
# repeat demo in single call
from qecsim import app
print(app.run_once(my_code, my_error_model, my_decoder, error_probability))
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Using Azure Machine Learning Pipelines for Batch Inference
In this notebook, we will demonstrate how to make predictions on large quantities of data asynchronously using the ML pipelines with Azure Machine Learning. Batch inference (or batch scoring) provides cost-effective inference, with unparalleled throughput for asynchronous applications. Batch prediction pipelines can scale to perform inference on terabytes of production data. Batch prediction is optimized for high throughput, fire-and-forget predictions for a large collection of data.
> **Tip**
If your system requires low-latency processing (to process a single document or small set of documents quickly), use [real-time scoring](https://docs.microsoft.com/azure/machine-learning/service/how-to-consume-web-service) instead of batch prediction.
In this example will be take a digit identification model already-trained on MNIST dataset using the [AzureML training with deep learning example notebook](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/ml-frameworks/keras/train-hyperparameter-tune-deploy-with-keras/train-hyperparameter-tune-deploy-with-keras.ipynb), and run that trained model on some of the MNIST test images in batch.
The input dataset used for this notebook differs from a standard MNIST dataset in that it has been converted to PNG images to demonstrate use of files as inputs to Batch Inference. A sample of PNG-converted images of the MNIST dataset were take from [this repository](https://github.com/myleott/mnist_png).
The outline of this notebook is as follows:
- Create a DataStore referencing MNIST images stored in a blob container.
- Register the pretrained MNIST model into the model registry.
- Use the registered model to do batch inference on the images in the data blob container.
## Prerequisites
If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first. This sets you up with a working config file that has information on your workspace, subscription id, etc.
### Connect to workspace
Create a workspace object from the existing workspace. Workspace.from_config() reads the file config.json and loads the details into an object named ws.
```
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
from azureml.core import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
```
### Create or Attach existing compute resource
By using Azure Machine Learning Compute, a managed service, data scientists can train machine learning models on clusters of Azure virtual machines. Examples include VMs with GPU support. In this tutorial, you create Azure Machine Learning Compute as your training environment. The code below creates the compute clusters for you if they don't already exist in your workspace.
> Note that if you have an AzureML Data Scientist role, you will not have permission to create compute resources. Talk to your workspace or IT admin to create the compute targets described in this section, if they do not already exist.
**Creation of compute takes approximately 5 minutes. If the AmlCompute with that name is already in your workspace the code will skip the creation process.**
```
import os
from azureml.core.compute import AmlCompute, ComputeTarget
# choose a name for your cluster
compute_name = os.environ.get("AML_COMPUTE_CLUSTER_NAME", "cpu-cluster")
compute_min_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MIN_NODES", 0)
compute_max_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MAX_NODES", 4)
# This example uses CPU VM. For using GPU VM, set SKU to STANDARD_NC6
vm_size = os.environ.get("AML_COMPUTE_CLUSTER_SKU", "STANDARD_D2_V2")
if compute_name in ws.compute_targets:
compute_target = ws.compute_targets[compute_name]
if compute_target and type(compute_target) is AmlCompute:
print('found compute target. just use it. ' + compute_name)
else:
print('creating a new compute target...')
provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size,
min_nodes = compute_min_nodes,
max_nodes = compute_max_nodes)
# create the cluster
compute_target = ComputeTarget.create(ws, compute_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it will use the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current AmlCompute status, use get_status()
print(compute_target.get_status().serialize())
```
### Create a datastore containing sample images
The input dataset used for this notebook differs from a standard MNIST dataset in that it has been converted to PNG images to demonstrate use of files as inputs to Batch Inference. A sample of PNG-converted images of the MNIST dataset were take from [this repository](https://github.com/myleott/mnist_png).
We have created a public blob container `sampledata` on an account named `pipelinedata`, containing these images from the MNIST dataset. In the next step, we create a datastore with the name `images_datastore`, which points to this blob container. In the call to `register_azure_blob_container` below, setting the `overwrite` flag to `True` overwrites any datastore that was created previously with that name.
This step can be changed to point to your blob container by providing your own `datastore_name`, `container_name`, and `account_name`.
```
from azureml.core.datastore import Datastore
account_name = "pipelinedata"
datastore_name = "mnist_datastore"
container_name = "sampledata"
mnist_data = Datastore.register_azure_blob_container(ws,
datastore_name=datastore_name,
container_name=container_name,
account_name=account_name,
overwrite=True)
```
Next, let's specify the default datastore for the outputs.
```
def_data_store = ws.get_default_datastore()
```
### Create a FileDataset
A [FileDataset](https://docs.microsoft.com/python/api/azureml-core/azureml.data.filedataset?view=azure-ml-py) references single or multiple files in your datastores or public urls. The files can be of any format. FileDataset provides you with the ability to download or mount the files to your compute. By creating a dataset, you create a reference to the data source location. If you applied any subsetting transformations to the dataset, they will be stored in the dataset as well. The data remains in its existing location, so no extra storage cost is incurred.
You can use dataset objects as inputs. Register the datasets to the workspace if you want to reuse them later.
```
from azureml.core.dataset import Dataset
mnist_ds_name = 'mnist_sample_data'
path_on_datastore = mnist_data.path('mnist')
input_mnist_ds = Dataset.File.from_files(path=path_on_datastore, validate=False)
```
The input dataset can be specified as a pipeline parameter, so that you can pass in new data when rerun the PRS pipeline.
```
from azureml.data.dataset_consumption_config import DatasetConsumptionConfig
from azureml.pipeline.core import PipelineParameter
pipeline_param = PipelineParameter(name="mnist_param", default_value=input_mnist_ds)
input_mnist_ds_consumption = DatasetConsumptionConfig("minist_param_config", pipeline_param).as_mount()
```
### Intermediate/Output Data
Intermediate data (or output of a Step) is represented by [PipelineData](https://docs.microsoft.com/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelinedata?view=azure-ml-py) object. PipelineData can be produced by one step and consumed in another step by providing the PipelineData object as an output of one step and the input of one or more steps.
```
from azureml.pipeline.core import Pipeline, PipelineData
output_dir = PipelineData(name="inferences", datastore=def_data_store)
```
### Download the Model
Download and extract the model from https://pipelinedata.blob.core.windows.net/mnist-model/mnist-tf.tar.gz to "models" directory
```
import tarfile
import urllib.request
# create directory for model
model_dir = 'models'
if not os.path.isdir(model_dir):
os.mkdir(model_dir)
url="https://pipelinedata.blob.core.windows.net/mnist-model/mnist-tf.tar.gz"
response = urllib.request.urlretrieve(url, "model.tar.gz")
tar = tarfile.open("model.tar.gz", "r:gz")
tar.extractall(model_dir)
os.listdir(model_dir)
```
### Register the model with Workspace
A registered model is a logical container for one or more files that make up your model. For example, if you have a model that's stored in multiple files, you can register them as a single model in the workspace. After you register the files, you can then download or deploy the registered model and receive all the files that you registered.
Using tags, you can track useful information such as the name and version of the machine learning library used to train the model. Note that tags must be alphanumeric. Learn more about registering models [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-deploy-and-where#registermodel)
```
from azureml.core.model import Model
# register downloaded model
model = Model.register(model_path="models/",
model_name="mnist-prs", # this is the name the model is registered as
tags={'pretrained': "mnist"},
description="Mnist trained tensorflow model",
workspace=ws)
```
### Using your model to make batch predictions
To use the model to make batch predictions, you need an **entry script** and a list of **dependencies**:
#### An entry script
This script accepts requests, scores the requests by using the model, and returns the results.
- __init()__ - Typically this function loads the model into a global object. This function is run only once at the start of batch processing per worker node/process. Init method can make use of following environment variables (ParallelRunStep input):
1. AZUREML_BI_OUTPUT_PATH – output folder path
- __run(mini_batch)__ - The method to be parallelized. Each invocation will have one minibatch.<BR>
__mini_batch__: Batch inference will invoke run method and pass either a list or Pandas DataFrame as an argument to the method. Each entry in min_batch will be - a filepath if input is a FileDataset, a Pandas DataFrame if input is a TabularDataset.<BR>
__run__ method response: run() method should return a Pandas DataFrame or an array. For append_row output_action, these returned elements are appended into the common output file. For summary_only, the contents of the elements are ignored. For all output actions, each returned output element indicates one successful inference of input element in the input mini-batch.
User should make sure that enough data is included in inference result to map input to inference. Inference output will be written in output file and not guaranteed to be in order, user should use some key in the output to map it to input.
#### Dependencies
Helper scripts or Python/Conda packages required to run the entry script.
```
scripts_folder = "Code"
script_file = "digit_identification.py"
# peek at contents
with open(os.path.join(scripts_folder, script_file)) as inference_file:
print(inference_file.read())
```
## Build and run the batch inference pipeline
The data, models, and compute resource are now available. Let's put all these together in a pipeline.
### Specify the environment to run the script
Specify the conda dependencies for your script. This will allow us to install pip packages as well as configure the inference environment.
* Always include **azureml-core** and **azureml-dataset-runtime\[fuse\]** in the pip package list to make ParallelRunStep run properly.
If you're using custom image (`batch_env.python.user_managed_dependencies = True`), you need to install the package to your image.
```
from azureml.core import Environment
from azureml.core.runconfig import CondaDependencies, DEFAULT_CPU_IMAGE
batch_conda_deps = CondaDependencies.create(pip_packages=["tensorflow==1.15.2", "pillow",
"azureml-core", "azureml-dataset-runtime[fuse]"])
batch_env = Environment(name="batch_environment")
batch_env.python.conda_dependencies = batch_conda_deps
batch_env.docker.base_image = DEFAULT_CPU_IMAGE
```
### Create the configuration to wrap the inference script
```
from azureml.pipeline.steps import ParallelRunStep, ParallelRunConfig
parallel_run_config = ParallelRunConfig(
source_directory=scripts_folder,
entry_script=script_file,
mini_batch_size=PipelineParameter(name="batch_size_param", default_value="5"),
error_threshold=10,
output_action="append_row",
append_row_file_name="mnist_outputs.txt",
environment=batch_env,
compute_target=compute_target,
process_count_per_node=PipelineParameter(name="process_count_param", default_value=2),
node_count=2
)
```
### Create the pipeline step
Create the pipeline step using the script, environment configuration, and parameters. Specify the compute target you already attached to your workspace as the target of execution of the script. We will use ParallelRunStep to create the pipeline step.
```
parallelrun_step = ParallelRunStep(
name="predict-digits-mnist",
parallel_run_config=parallel_run_config,
inputs=[ input_mnist_ds_consumption ],
output=output_dir,
allow_reuse=False
)
```
### Run the pipeline
At this point you can run the pipeline and examine the output it produced. The Experiment object is used to track the run of the pipeline
```
from azureml.core import Experiment
pipeline = Pipeline(workspace=ws, steps=[parallelrun_step])
experiment = Experiment(ws, 'digit_identification')
pipeline_run = experiment.submit(pipeline)
```
### Monitor the run
The pipeline run status could be checked in Azure Machine Learning portal (https://ml.azure.com). The link to the pipeline run could be retrieved by inspecting the `pipeline_run` object.
```
# This will output information of the pipeline run, including the link to the details page of portal.
pipeline_run
```
### Optional: View detailed logs (streaming)
```
# Wait the run for completion and show output log to console
pipeline_run.wait_for_completion(show_output=True)
```
### View the prediction results per input image
In the digit_identification.py file above you can see that the ResultList with the filename and the prediction result gets returned. These are written to the DataStore specified in the PipelineData object as the output data, which in this case is called *inferences*. This containers the outputs from all of the worker nodes used in the compute cluster. You can download this data to view the results ... below just filters to the first 10 rows
```
import pandas as pd
import tempfile
batch_run = pipeline_run.find_step_run(parallelrun_step.name)[0]
batch_output = batch_run.get_output_data(output_dir.name)
target_dir = tempfile.mkdtemp()
batch_output.download(local_path=target_dir)
result_file = os.path.join(target_dir, batch_output.path_on_datastore, parallel_run_config.append_row_file_name)
df = pd.read_csv(result_file, delimiter=":", header=None)
df.columns = ["Filename", "Prediction"]
print("Prediction has ", df.shape[0], " rows")
df.head(10)
```
### Resubmit a with different dataset
Since we made the input a `PipelineParameter`, we can resubmit with a different dataset without having to create an entirely new experiment. We'll use the same datastore but use only a single image.
```
path_on_datastore = mnist_data.path('mnist/0.png')
single_image_ds = Dataset.File.from_files(path=path_on_datastore, validate=False)
pipeline_run_2 = experiment.submit(pipeline,
pipeline_parameters={"mnist_param": single_image_ds,
"batch_size_param": "1",
"process_count_param": 1}
)
# This will output information of the pipeline run, including the link to the details page of portal.
pipeline_run_2
# Wait the run for completion and show output log to console
pipeline_run_2.wait_for_completion(show_output=True)
```
## Cleanup Compute resources
For re-occurring jobs, it may be wise to keep compute the compute resources and allow compute nodes to scale down to 0. However, since this is just a single-run job, we are free to release the allocated compute resources.
```
# uncomment below and run if compute resources are no longer needed
# compute_target.delete()
```
| github_jupyter |
```
import numpy as np
import time
import pandas as pd
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
datasets = ['data/fdata1','data/fdata2','data/fdata3','data/fdata4']
# import os
# data0 = []
# for dataset in datasets:
# metadata = os.path.join(dataset, 'fdata.csv')
# pic_path = os.path.join(dataset, 'pic')
# df = pd.read_csv(metadata, names=['name','state'])
# df['name'] = [os.path.join(pic_path, pic) for pic in df['name']]
# data0.append(df)
import os
import copy
# MouseMode = False
MouseMode = True
data0 = []
for dataset in datasets:
metadata = os.path.join(dataset, 'fdata.csv')
pic_path = os.path.join(dataset, 'pic')
df = pd.read_csv(metadata, names=['name','state'])
if MouseMode: dfv = copy.copy(df)
df['name'] = [os.path.join(pic_path, pic) for pic in df['name']]
data0.append(df)
if MouseMode:
pic_pathv = pic_path + 'v' # picv
if not os.path.exists(pic_pathv):
os.makedirs(pic_pathv)
for pic in dfv['name']:
# make flip image
imgFile = os.path.join(pic_path, pic)
img = cv2.imread(imgFile)
img = copy.deepcopy(img)
img = cv2.flip(img, 1) # flip horizontally
# save flip image
imgFilev = os.path.join(pic_pathv, 'v_' + pic)
cv2.imwrite(imgFilev, img)
dfv['name'] = [os.path.join(pic_pathv, 'v_' + pic) for pic in dfv['name']]
dfv['state'] = [ 'v' + state[-1] for state in dfv['state']]
data0.append(dfv)
# join into one
joined_data = pd.concat(data0)
joined_data.head()
joined_data.to_csv('full_data.csv', index=False)
num_class = len(joined_data['state'].unique())
ges_to_num = dict({(g,i) for i, g in enumerate(joined_data['state'].unique())})
num_to_ges = dict({(i,g) for i, g in enumerate(joined_data['state'].unique())})
import json
with open('ges2num.json', 'w') as fout:
json.dump(ges_to_num, fout)
with open('num2ges.json', 'w') as fout:
json.dump(num_to_ges, fout)
num_class, ges_to_num
joined_data = joined_data.replace({'state':ges_to_num})
joined_data.head()
# labels = np.empty((joined_data.shape[0]))
# res_width, res_height = 200, 200
# imgs = np.empty(shape=(joined_data.shape[0],2,res_width,res_height))
# imgs.shape, labels.shape
from collections import defaultdict
processed_hand = defaultdict(list)
processed_mask = defaultdict(list)
for i, (im_path, state) in enumerate(joined_data.values):
im_name = im_path.split('/')[-1]
hand_outdir = os.path.join(im_path.split('/')[0], im_path.split('/')[1], 'hand_pic')
mask_outdir = os.path.join(im_path.split('/')[0], im_path.split('/')[1], 'mask_pic')
if not os.path.exists(hand_outdir):
os.makedirs(hand_outdir)
if not os.path.exists(mask_outdir):
os.makedirs(mask_outdir)
im = cv2.imread(im_path)
im_ycrcb = cv2.cvtColor(im, cv2.COLOR_BGR2YCR_CB)
skin_ycrcb_mint = np.array((0, 133, 77))
skin_ycrcb_maxt = np.array((255, 173, 127))
skin_ycrcb = cv2.inRange(im_ycrcb, skin_ycrcb_mint, skin_ycrcb_maxt)
_, contours, _ = cv2.findContours(skin_ycrcb, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
# fimd max contour
max_area=100
ci=0
for i in range(len(contours)):
cnt=contours[i]
area = cv2.contourArea(cnt)
if(area>max_area):
max_area=area
ci=i
cnts = contours[ci]
(x,y,w,h) = cv2.boundingRect(cnts)
moments = cv2.moments(cnts)
# Central mass of first order moments
if moments['m00']!=0:
cx = int(moments['m10']/moments['m00']) # cx = M10/M00
cy = int(moments['m01']/moments['m00']) # cy = M01/M00
centerMass=(cx,cy)
#Draw center mass
cv2.circle(im,centerMass,1,[100,0,255],1)
crop = im[y:y+h, x:x+w]
f_im = cv2.resize(crop,(200, 200), interpolation=cv2.INTER_CUBIC)
hand_out_path = os.path.join(hand_outdir, im_name)
mask_out_path = os.path.join(mask_outdir, im_name)
cv2.imwrite(hand_out_path, f_im)
cv2.imwrite(mask_out_path, skin_ycrcb)
processed_hand['name'].append(hand_out_path)
processed_hand['state'].append(state)
processed_mask['name'].append(mask_out_path)
processed_mask['state'].append(state)
print hand_out_path, mask_out_path
pd.DataFrame(processed_hand).to_csv('full_hand_data.csv', index=False)
pd.DataFrame(processed_mask).to_csv('full_mask_data.csv', index=False)
```
### This outputs "full_data.csv", "full_hand_data.csv", "full_mask_data.csv"
### And gesture to number mappings: "ges2num.json" and "num2ges.json
| github_jupyter |
# Watch Me Build an AI Startup (10 Steps)

- **Step 1** List Personal Problems
- **Step 2 ** Market Research (competing products)
- **Step 3** Buy Domain
- **Step 4** Create Landing Page
- **Step 5** Share Landing Page
- **Step 6** Create Business Plan
- **Step 7** Design Pipeline
- **Step 8** Transfer Learning (!)
- **Step 9** Create Web App
- **Step 10** Deploy!
# Step 1 - List Personal Problems
## The star is your next startup idea

## This has actually been codified by the Japanese 100+ years ago. It's called "Ikigai"

### What Industry excites you?

### My 3 passions (human enhancement)
- Healthcare - Physical Enhancement via AI
- Education - Intellectual Enhancement via AI
- Scientific Research - Increasing human knowledge of this reality
## Thought exercise
1) Hey, this problem sucks!
2) How many people have this problem?
3) How many other ways are there to solve it or live with it?
4) What would it cost to offer a solution?
5) How does the cost compare with the pain of the problem, and the cost of other options (including 'doing nothing'!)?
# Still can't find a problem you're passionate about? Live abroad for at least a month (increase training data via new culture, values, language, perspectives)
# Step 2 - Market Research (competing products)
### AI can do so much! This is a subset of possibilities

### Lets do Automated Diagnosis using image classification technology
### Competitor Landscape

### The Lung Imaging Landscape looks easily disruptable (not specialized enough, pricing out lower end hospitals)
# Step 3 - Buy Domain
1. Go to Godaddy.com
2. Find an unused domain name (don't buy an existing one, not worth it)
3. Buy it!
# Order of Preference
1. .com
2. .ai
# Step 4 - Create Landing Page
## We need a logo and style guide! Lets use AI to generate one (i used this for School of AI as well)
[www.brandmark.io](https://www.brandmark.io)

## Now we can instantly create an email landing page with Mailchimp
[www.mailchimp.com](https://www.mailchimp.com)
## Step 5 - Share Landing Page
- Reach out to Social Networks to see if we get any signups
- Reach out to nearby hospitals and other medical facitilies

# Step 6 - Create Business Plan

- Step 1 - Get 25 signups
- Step 2 - Create SaaS
- Step 3 - Convert signups to paid clients
- Step 4 - Collect feedback
- Step 5 - Improve app using feedback
- Step 6 - Collect 100 more paid clients
- Step 7 - Repeat Steps 4-6 until I hit a scaling limit
- Step 8 - Make 2 hires (full stack engineer, sales)
- Step 9 - Create more generalized offering
- Step 10 - Raise a first round
- Step 11 - Hire a larger team
- Step 12 - ???
- Step 13 - Profit
- Step 14 - Exit Strategy (Google buys us out. Actually, we'll buy Google out :) )

# Step 7 Design Pipeline
## Features we need
- User Authentication (SQL)
- Database (SQL)
- Payment (Stripe)
- Uploads (Flask native)
- Inference (Keras)

#### OK so...
- Authentication + Database + Payment functionality https://github.com/alectrocute/flasksaas
- Upload functionality https://github.com/bboe/flask-image-uploader
- Inference? Keras
##### basically..
If upload && paid == true, run inference, display result!
### Step 8 Transfer Learning (!)
## Convolutional Networks do really well at Image classification
## So we just need to train a model on labeled data
## But that already exists!
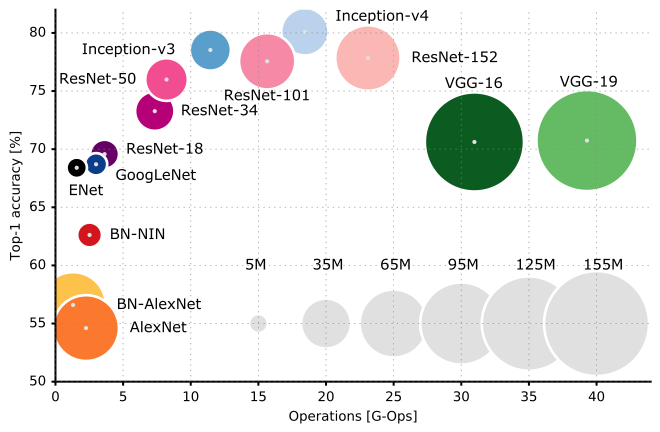
## So we'll retrain an existing model (ResNet) on new data to leverage existing learnings and improve overall accuracy (transfer learning!)
- 1: There is no need for an extremely large training dataset.
- 2: Not much computational power is required. As we are using pre-trained weights and only have to learn the weights of the last few layers.
# Step 9 - Write Web App
# Step 10 - Deploy!
- Mobile app?
- Continous Serving via Tensorflow Serving
- Batch inference
- Better Design
| github_jupyter |
In this notebook, we play a bit with the visualization and sampling of a Gaussian mixture. We first sample from a two-dimensional Gaussian mixture. But before we start, we need to import a few packages and define some helper functions.
```
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats
##############################################
# Draw from a finite distribution defined
# by a vector p with elements adding up
# to one. We return a number between
# 1 and the number of elements of p
# and i is returned with probability p[i-1]
##############################################
def draw(p):
u = np.random.uniform()
x = 0
n = 1
for i in p:
x = x + i
if x >= u:
return n
n += 1
return i
#############################################
#
# Utility function to create a sample
#
# This will create a sample of data points
# distributed over K clusters. The sample
# will have size points.
# The clusters are Gaussian normal
# distributions with the given centres
# and standard deviations
#
# The centre points are assumed to be given
# as a matrix with K rows. The vector p is
# a vector with non-negative elements that
# sum up to one
#
# Returns the sample and the true cluster
#############################################
def get_sample(M, p, cov, size=10):
#
# Number of clusters
#
X = []
T = []
for i in range(size):
k = draw(p) - 1
T.append(k)
#
# Then draw from the normal distribution with mean M[k,:]
#
rv = scipy.stats.multivariate_normal(mean=M[k], cov=cov[k])
_X = [rv.rvs()]
X.append(_X)
return np.concatenate(X), T
#############################################
#
# Plot a distribution
#
#############################################
def plot_clusters(S, T, axis):
for i in range(S.shape[0]):
x = S[i,0]
y = S[i,1]
if T[i] == 0:
axis.plot([x],[y],marker="o", color="red")
else:
axis.plot([x],[y],marker="o", color="blue")
_M = np.array([[5,1], [1,4]])
fig = plt.figure(figsize=(20,8))
ax1 = fig.add_subplot(1,2,2)
S, T = get_sample(_M,p=[0.5, 0.5], size=200, cov=[[0.5, 0.5], [0.5,0.5]])
plot_clusters(S,T, ax1)
```
Now let us try something else. We sample from a one-dimensional distribution and create a histogramm
```
_M = np.array([2, -3])
ax2 = fig.add_subplot(1,2,1)
cov = [0.5, 0.5]
p = [0.7, 0.3]
S, T = get_sample(_M,p=p, size=500, cov=cov)
ax2.hist(S, 100, rwidth=0.8, normed=True);
x = np.arange(-5, 5, 0.1)
for k in range(2):
pdf = p[k]*scipy.stats.norm.pdf(x, loc=_M[k], scale=cov[k])
ax2.plot(x, pdf, "g")
plt.savefig("/tmp/GaussianMixture.png")
plt.show()
```
| github_jupyter |
# Zajęcia 1 (część 2)
## Wczytanie danych
Bedziemy wykorzystywać danye z ankiety StackOverflow z 2020.
https://insights.stackoverflow.com/survey/
Dane sa dostepne na google drive. Skorzystamy z modułu GoogleDriveDownloader, ktory pozwala pobrac dokument o podanym id.
```
from google_drive_downloader import GoogleDriveDownloader as gdd
from pathlib import Path
path_dir = str(Path.home()) + "/data/2020/" # ustawmy sciezke na HOME/data/2020
archive_dir = path_dir + "survey.zip" # plik zapiszemy pod nazwa survey.zip
path_dir
archive_dir
# sciagniecie pliku we wskazane miejsce
gdd.download_file_from_google_drive(file_id='1dfGerWeWkcyQ9GX9x20rdSGj7WtEpzBB',
dest_path=archive_dir,
unzip=True)
```
## <span style='background:yellow'> ZADANIE 1 </span>
Zapoznaj sie z plikami tekstowymi (survey_results_public.csv oraz survey_results_schema.csv). Podejrzyj ich zawartość, sprawdź ich wielkość (liczba liniii oraz rozmiar). Wgraj plik do swojego kubełka na GCS do survey/2020/ gs://bdg-lab-$USER/survey/2020/. Jesli nie masz kubełka stwórz go.
## Podłączenie do sesji Spark na GKE
#### WAZNE
jesli w poprzednim notatniku masz aktywną sesję Spark zakończ ją (w poprzednim notatniku) poleceniem spark.stop()
```
from pyspark.sql import SparkSession
spark = SparkSession \
.builder \
.config('spark.driver.memory','1g')\
.config('spark.executor.memory', '1g') \
.getOrCreate()
```
## Dostęp do danych na GCS
```
import os
user_name = os.environ.get('USER')
print(user_name)
# ścieżka dostępu do pliku na GCS
gs_path = f'gs://bdg-lab-{user_name}/survey/2020/survey_results_public.csv'
gs_path
```
## Spark SQL
Platforma Apache Spark posiada komponent Spark SQL, który pozwala traktować dane jak tabele w bazie danych. Można zakładać swoje schematy baz danych oraz korzystać z języka SQL.
```
table_name = "survey_2020" # nazwa tabeli ktora bedziemy chcieli stworzyc
spark.sql(f'DROP TABLE IF EXISTS {table_name}') # usun te tabele jesli istniala wczesniej
# stworz tabele korzystajac z danych we wskazanej lokalizacji
spark.sql(f'CREATE TABLE IF NOT EXISTS {table_name} \
USING csv \
OPTIONS (HEADER true, INFERSCHEMA true) \
LOCATION "{gs_path}"')
```
## Weryfikacja danych
Sprawdzmy zaczytane dane.
```
spark.sql(f"describe {table_name}").show() # nie wszystkie dane ...
spark.sql(f"describe {table_name}").show(100, truncate=False) # niepoprawne typy danych... "NA"
spark.sql(f"SELECT DISTINCT Age FROM {table_name} ORDER BY Age DESC").show()
```
## Obsługa wartosci 'NA' - ponowne stworzenie tabeli
```
spark.sql(f'DROP TABLE IF EXISTS {table_name}')
# wykorzystujemy dodatkową opcję: NULLVALUE "NA"
spark.sql(f'CREATE TABLE IF NOT EXISTS {table_name} \
USING csv \
OPTIONS (HEADER true, INFERSCHEMA true, NULLVALUE "NA") \
LOCATION "{gs_path}"')
spark.sql(f"DESCRIBE {table_name}").show(100)
spark.sql(f"SELECT DISTINCT Age FROM {table_name} ORDER BY Age DESC").show()
# sprawdzenie liczności tabeli
spark.sql(f"select count(*) from {table_name}").show()
spark.sql(f"select count(*) from {table_name}").explain() # tak jak na poprzednich zajeciach mozemy wygenerowac plany wykonania polecenia
df = spark.sql(f"select * from {table_name}") # mozemy tez w zapytaniu SELECT pobrac wszystkie wiersze i kolumny i zapisac je jako DF (jak na poprzednich zajeciach)
df
```
## Podgląd danych
```
spark.sql(f"select * from {table_name}").show()
```
## Biblioteka Pandas
https://pandas.pydata.org/
Moduł Pandas jest biblioteką Pythonową do manipulacji danymi. W szczegolnosci w pandas mozemy stworzyc ramki danych i wykonywac na niej analize, agregacje oraz wizualizacje danych.
Przy nieduzych zbiorach danych i prostych operacjach to doskonała biblioteka. Jednak kiedy zbior danych sie rozrasta lub kiedy wymagane sa zlozone transformacje to operacje moga byc wolne.
Operacje na rozproszonych danych sa szybsze. Ale tu takze napotykamy ograniczenia np trudność w wizualizacji danych.
```
import pandas as pd
spark.sql(f"select * from {table_name} limit 10").toPandas()
```
**Ważne**
Metoda toPandas() na ramce pyspark, konwertuje ramkę pyspark do ramki pandas. Wykonuje akcje pobrania (collect) wszystkich danych z executorów (z JVM) i transfer do programu sterujacego (driver) i konwersje do typu Pythonowego w notatniku. Ze względu na ograniczenia pamięciowe w programie sterującym należy to wykonywać na podzbiorach danych.
**DataFrame.collect() collects the distributed data to the driver side as the local data in Python. Note that this can throw an out-of-memory error when the dataset is too large to fit in the driver side because it collects all the data from executors to the driver side.**
**Note that DataFrame.toPandas() results in the collection of all records in the DataFrame to the driver program and should be done on a small subset of the data.**
```
dist_df = spark.sql(f"select * from {table_name} LIMIT 10")
local_df = spark.sql(f"select * from {table_name} LIMIT 10").toPandas()
type(dist_df) # dataframe Sparkowy ("przepis na dane, rozproszony, leniwy")
type(local_df) # dataframe Pandasowy (lokalny, sciągnięty do pamięci operacyjnej)
dist_df.show()
local_df
pd.set_option('max_columns', None) # pokazuj wszystkie kolumny
# pd.reset_option(“max_columns”)
```
## <span style='background:yellow'> ZADANIE 2 </span>
Napisz w Spark SQL zapytanie które zwróci średnią liczbę godzin przepracowywanych przez z respondentów pogrupowanych ze względu na kraj. Następnie przekształć wynik do ramki pandasowej i ją wyświetl.
## Wizualizacje
Do wizualizacji będziemy się posługiwać modułami matplotlib (https://matplotlib.org/) i seaborn (https://seaborn.pydata.org/). Do bardzo rozbudowane moduły, zachęcamy do eksploracji oficjalnych dokumentacji. Na zajęciach zrealizujemy następujące wykresy:
* histogramy
* liniowe
* wiolinowe
* kołowe
Moduły wizualizacyjne wymagają danych na lokalnej maszynie. Mogą być to natywne typy danych Pythonowe (słowniki, listy) ale także np ramki danych pandasowe. ~~Nie działa wizualizacja na ramkach danych Sparkowych.~~
```
import matplotlib.pyplot as plt
import seaborn as sns
```
## Narysuj histogram wieku respondentów
```
# przygotowanie danych
# przycinamy dane tylko do zakresu ktory jest potrzebny do realizacji polecenia
ages = spark.sql(f"SELECT CAST (Age AS INT) \
FROM {table_name} \
WHERE age IS NOT NULL \
AND age BETWEEN 10 AND 80").toPandas()
ages
ages.hist("Age", bins=10)
sns.displot(ages, bins=10, rug=True, kde=False)
```
## Jaki jest udział programistów hobbistów? Narysuj wykres kołowy
Będzie nas interesowała ta proporcja ze względu na płeć.
```
# przygotowanie (filtrowanie, grupowanie i zliczenie) danych na rozproszonych danych (spark sql)
# pozniej pobranie do pandasowej ramki
hobby_all = spark.sql(f"SELECT Hobbyist, COUNT(*) AS cnt FROM {table_name} WHERE Hobbyist IS NOT NULL GROUP BY Hobbyist").toPandas()
hobby_men = spark.sql(f"SELECT Hobbyist, COUNT(*) AS cnt FROM {table_name} WHERE Hobbyist IS NOT NULL AND Gender='Man' GROUP BY Hobbyist").toPandas()
hobby_women = spark.sql(f"SELECT Hobbyist, COUNT(*) AS cnt FROM {table_name} WHERE Hobbyist IS NOT NULL AND Gender='Woman' GROUP BY Hobbyist").toPandas()
hobby_all.plot.pie(y='cnt', labels=hobby_all['Hobbyist'], title="All", autopct='%.0f')
plt.legend(loc="lower center")
plt.show()
hobby_men.plot.pie(y='cnt', labels=hobby_men['Hobbyist'], title="Men", autopct='%.0f')
plt.legend(loc="lower center")
plt.show()
hobby_women.plot.pie(y='cnt', labels=hobby_women['Hobbyist'], title="Women", autopct='%.0f')
plt.legend(loc="lower center")
plt.show()
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(15,5))
hobby_all.plot.pie(y='cnt', labels=hobby_all['Hobbyist'], title="All", ax=axes[0], autopct='%.0f')
hobby_men.plot.pie(y='cnt', labels=hobby_men['Hobbyist'], title="Men", ax=axes[1], autopct='%.0f')
hobby_women.plot.pie(y='cnt', labels=hobby_men['Hobbyist'], title="Women", ax=axes[2], autopct='%.0f')
plt.show()
```
## Wykres liniowy. Zależność między wiekiem a liczbą przepracowanych godzin
Interesują nas dla developerzy profesjonaliści (nie hobbiści) w przedziale wiekowym 18-65.
```
# przygotowanie (filtrowanie, grupowanie, wyliczenie sredniej oraz sortowanie) danych na rozproszonych danych (spark sql)
# pozniej pobranie do pandasowej ramki
age_work = spark.sql(f"SELECT age, CAST (avg(WorkWeekHrs) AS INT) AS avg FROM {table_name} \
WHERE WorkWeekHrs IS NOT NULL AND age BETWEEN 18 AND 65 AND hobbyist = 'No' \
GROUP BY age \
ORDER BY age ASC").toPandas()
age_work
age_work.plot(x='age', y='avg', kind='line')
sns.relplot(x="age", y="avg", kind="line", data=age_work);
```
## Wykres słupkowy. Pokaż liczbę respondentów na kraj
Interesuje nas tylko 10 krajów o najwyższej liczbie respondentow.
```
# przygotowanie (grupowanie, zliczenie, sortowanie oraz przyciecie do 10 wyników) danych na rozproszonych danych (spark sql)
# pozniej pobranie do pandasowej ramki
max_countries = spark.sql(f"SELECT country, COUNT(*) AS cnt \
FROM {table_name} \
GROUP BY country \
ORDER BY cnt DESC \
LIMIT 10 ").toPandas()
max_countries.plot.bar(y='cnt', x='country')
plt.show()
sns.catplot(x="country", y="cnt", kind="bar",\
data=max_countries).set_xticklabels(rotation=65)
plt.show()
```
## Wykres słupkowy. Średnie zarobki w krajach w ktorych jest powyzej 1000 respondentów
```
# przygotowanie (filtrowanie, grupowanie, wyliczenie sredniej, filtrowanie po liczności i grup oraz sortowanie) danych na rozproszonych danych (spark sql)
# pozniej pobranie do pandasowej ramki
country_salary = spark.sql(f"SELECT country, \
CAST (avg(ConvertedComp) AS INT) as avg \
FROM {table_name} \
WHERE country IS NOT NULL \
GROUP BY country \
HAVING COUNT(*) > 1000 \
ORDER BY avg DESC ").toPandas()
country_salary.plot.barh(("country"))
plt.show()
```
## Boxplot. Pokaz rozklad pensji w krajach gdzie jest powyzej 1000 respondentów
Tutaj będziemy musieli skorzystać z podzapytania.
```
# przygotowanie danych na rozproszonych danych (spark sql). Mamy tu do czynienia z podzapytaniem
# pozniej pobranie do pandasowej ramki
country_comp = spark.sql(f"SELECT country, CAST(ConvertedComp AS INT) \
FROM {table_name} \
WHERE country IN (SELECT country FROM {table_name} GROUP BY country HAVING COUNT(*) > 1000) \
AND ConvertedComp IS NOT NULL AND ConvertedComp > 0 \
ORDER BY ConvertedComp desc").toPandas()
country_comp
country_comp.boxplot(column="ConvertedComp", by="country", \
showfliers=False, rot=60, meanline=True)
plt.show()
sns.catplot(x="country", y="ConvertedComp", kind="box", \
showfliers=False, data=country_comp, palette="Blues")\
.set_xticklabels(rotation=65)
plt.show()
```
## <span style='background:yellow'> ZADANIE 3 </span>
Narysuj rozklad pensji w zaleznosci od plci.
## Narysuj wykres popularnosci jezykow programowania
```
spark.sql(f"select LanguageWorkedWith from {table_name} where LanguageWorkedWith IS NOT NULL").show(truncate=False)
```
Języki programowania są zapisane w pojedynczej komórce. Będzie trzeba je rozdzielić i zliczyć. Tak przygotowane dane dopiero posłużą nam do narysowania wykresu. Wykorzystamy funkcję `posexplode`.
```
langs = spark.sql(f"select LanguageWorkedWith from {table_name} where LanguageWorkedWith IS NOT NULL")
from pyspark.sql.functions import *
langs.select(
posexplode(split("LanguageWorkedWith", ";")).alias("pos", "language")
).show()
langs.select(
posexplode(split("LanguageWorkedWith", ";")).alias("pos", "language")).groupBy("language").count().orderBy("count").show()
langs_pd = langs.select(
posexplode(split("LanguageWorkedWith", ";")).alias("pos", "language")).groupBy("language").count().orderBy("count").toPandas()
langs_pd
from matplotlib.pyplot import figure
figure(figsize=(8, 9))
plt.barh(width=langs_pd["count"], y=langs_pd["language"])
plt.show()
```
## Narysuj wykres popularnosci jezykow wsrod Data Scientists
Zdefiniujmy sobie funkcję, która przekształca nam języki do wymaganej przez nas postaci
```
from pyspark.sql.functions import *
def prepare_lang(df, colName='LanguageWorkedWith'):
summary = df.select(posexplode(split(colName, ";")).alias("pos", "language")).groupBy("language").count().orderBy("count")
return summary
langs_ds = spark.sql(f"SELECT LanguageWorkedWith \
FROM {table_name} \
WHERE DevType LIKE '%Data scientist%'")
sum_lang = prepare_lang(langs_ds).toPandas()
figure(figsize=(8, 9))
plt.barh(width=sum_lang["count"], y=sum_lang["language"])
plt.show()
```
## Narysuj wykres którego chcą wykorzystywać w przyszłości Data Scientists
```
lang_desired = spark.sql(f"select LanguageDesireNextYear \
from {table_name} \
where DevType like '%Data scientist%'")
sum_lang = prepare_lang(lang_desired, 'LanguageDesireNextYear').toPandas()
figure(num=None, figsize=(8, 9), dpi=80, facecolor='w', edgecolor='k')
plt.barh(width=sum_lang["count"], y=sum_lang["language"])
plt.show()
```
## Narysuj wykres prezentujący liczbę godzin na pracy w zależności od wykształcenia
```
spark.sql(f"select distinct EdLevel from {table_name}").show(truncate=False) # jakie są wartości wykształcenia
ed_pandas = spark.sql(f"SELECT EdLevel, WorkWeekHrs FROM {table_name} \
WHERE WorkWeekHrs BETWEEN 10 AND 80 \
AND (EdLevel LIKE '%Bachelor%' OR EdLevel LIKE '%Master%' OR EdLevel LIKE '%Other doctoral%')").toPandas()
ed_pandas['EdLevel'] = ed_pandas['EdLevel'].replace('Bachelor’s degree (B.A., B.S., B.Eng., etc.)','Bachelor')
ed_pandas['EdLevel'] = ed_pandas['EdLevel'].replace('Master’s degree (M.A., M.S., M.Eng., MBA, etc.)','Master')
ed_pandas['EdLevel'] = ed_pandas['EdLevel'].replace('Other doctoral degree (Ph.D., Ed.D., etc.)','Doctor')
ed_pandas
sns.catplot(x="EdLevel", y="WorkWeekHrs", data=ed_pandas)
```
## Narysuj wykres wiolinowy pokazujacy rozkład dochodów w zależności od wykształcenia
```
ed_pay = spark.sql(f"SELECT EdLevel, CAST (CompTotal AS INT) AS CompTotal FROM {table_name} \
WHERE CompTotal BETWEEN 0 AND 1000000 \
AND (EdLevel LIKE '%Bachelor%' OR EdLevel LIKE '%Master%' OR EdLevel LIKE '%Other doctoral%')").toPandas()
ed_pay['EdLevel'] = ed_pay['EdLevel'].replace('Bachelor’s degree (B.A., B.S., B.Eng., etc.)','Bachelor')
ed_pay['EdLevel'] = ed_pay['EdLevel'].replace('Master’s degree (M.A., M.S., M.Eng., MBA, etc.)','Master')
ed_pay['EdLevel'] = ed_pay['EdLevel'].replace('Other doctoral degree (Ph.D., Ed.D., etc.)','Doctor')
ed_pay
sns.catplot(x="EdLevel", y="CompTotal", kind="boxen",
data=ed_pay);
```
## ⭐ Narysuj heatmape odwiedzin na StackOverflow dla wybranych krajów ⭐
```
spark.sql(f"SELECT DISTINCT SOVisitFreq FROM {table_name}").show(truncate=False)
so_v = spark.sql(f"SELECT SOVisitFreq, t1.country, COUNT(*)/first(t2.t) AS cnt from {table_name} t1 \
JOIN (SELECT country, COUNT(*) as t FROM {table_name} GROUP BY country) t2 \
ON t1.country = t2.country \
WHERE t1.country IS NOT NULL AND SOVisitFreq IS NOT NULL \
AND t1.country IN ('Poland', 'United States', 'Russian Federation', 'China', 'India', 'Germany', 'Japan') \
GROUP BY t1.country, SOVisitFreq").toPandas()
so_v['SOVisitFreq'] = pd.Categorical(so_v['SOVisitFreq'], ["I have never visited Stack Overflow (before today)", "Less than once per month or monthly", "A few times per month or weekly", "A few times per week", "Daily or almost daily", "Multiple times per day"])
# so_v.sort_values['SOVisitFreq']
heatmap2_data = pd.pivot_table(so_v, values='cnt', index=['country'], columns='SOVisitFreq')
sns.heatmap(heatmap2_data, cmap="BuGn")
```
## <span style='background:yellow'> ZADANIE 4 </span>
* Narysuj wykres słupkowy popularności wykorzystywanych baz danych przez profesjonalnych programistów.
* Narysuj wykres kołowy przedstawiający procentowy udział poziomu wykształcenia inz, mgr i dr w grupie respondentów.
```
spark.stop()
```
| github_jupyter |
**NB:** This notebook was created using **Google Colab**
[](https://colab.research.google.com/github/A2-insights/Covid19-Data-Insights/blob/master/notebooks/A2_insights_Lab_Covid19_EDA.ipynb)
# **$A^{2}$ Insights Lab** *presents*
The COVID-19 informative analysis crafted using python, pandas and other modern technologies.
---
## **Disclaimer**
The information contained in this Notebook is for **Educational purposes only**. The Data Analysis and code is provided by **$A^{2}$ Insights Lab**, Conclusions or point of views made from this notebook should be interpreted as Opinoins and not as health professional advice. **Any reliance you place on such information is therefore strictly at your own risk.**
**We Do not Claim** ownership nor correctness of the data used, therefore
In no event will we be liable for any violation of the government Regulations or damage including without limitation, indirect or consequential loss or damage whatsoever arising from loss of data or misleading information, or in connection with, the use of this information.
<br>
# **Imports**
```
# Data manipulation
import pandas as pd
import numpy as np
import requests
# Data Visualization
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
# Extas
import warnings
# Disabling library Warnings
warnings.filterwarnings('ignore')
# Out Put Settings, precision
pd.set_option('precision', 3)
```
# **Loading Data**
---
**API starter:**
>Use this notebook to connect to the COVID-19 database for access to our datasources.
>Use [these API's](https://ijvcpr9af1.execute-api.eu-west-1.amazonaws.com/api/) to access the data.
---
## Build a dataframe using the following code:
```url = "INSERT API URL"``` <br>
```headers = {'x-api-key': "INSERT API KEY HERE"}```<br>
```response = requests.request("GET", url, headers=headers)```<br>
```x = response.json()```<br>
```df = pd.DataFrame(x)```
## **Call the following function to read the data To a Dataframe**
```
load_covid19(url, API_KEY)
```
<br>
<br>
## CONSTANTS
```
# CONSTANTS
API_KEY = 'WVllUkRA01awNNgKxGg607vl5qIvuOAN3pW9HXmD'
# External Data Sources URLs
SYMPTOMS_URL = 'https://en.wikipedia.org/wiki/Coronavirus'
# FULL GLOBAL DATASETS
GLOBAL_URL = 'https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/ecdc/full_data.csv'
LOCATIONS_URL = 'https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/ecdc/locations.csv'
# GLOBAL DATA TO BE MERGED
CONFIRMED_URL = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv'
RECOVERED_URL = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv'
DEATHS_URL = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv'
# SOUTH AFRICA DATASETS
RSA_URL = 'https://raw.githubusercontent.com/dsfsi/covid19za/master/data/covid19za_provincial_cumulative_timeline_confirmed.csv'
# DISTRICT DATA
WESTERN_CAPE_URL = 'https://raw.githubusercontent.com/dsfsi/covid19za/master/data/district_data/provincial_wc_cumulative.csv'
GAUTENG_URL = 'https://raw.githubusercontent.com/dsfsi/covid19za/master/data/district_data/provincial_gp_cumulative.csv'
# API URLs
url_CasesGlobalView = 'https://9gnht4xyvf.execute-api.eu-west-1.amazonaws.com/api/get_table/CasesGlobalView'
url_CasesLocalView = 'https://9gnht4xyvf.execute-api.eu-west-1.amazonaws.com/api/get_table/CasesLocalView'
ulr_CounterMeasureView = 'https://9gnht4xyvf.execute-api.eu-west-1.amazonaws.com/api/get_table/CounterMeasureView'
```
## **GATHER EXTERNAL DATA**
---
Functions to gather data from other sources[link text](https://)
```
def scrape_covid19_symptoms(url):
"""scrape_covid19_symptoms(url)
Scrapes covid 19 symptoms from wikipedia coronavirus pages
PARAMETERS:
url (string): wikipedia coronavirus url page
"""
# get all tables and extract covid 19 symptoms table
df = pd.read_html(url, header=None)
df = df[1][11:17].loc[: , ['Unnamed: 0', 'SARS-CoV-2'] ].reset_index().drop('index', axis=1)
df.rename(columns={'Unnamed: 0': 'symptoms', 'SARS-CoV-2': 'covid19_percentage'}, inplace= True)
return df
def load_covid19(url, API_KEY = 'WVllUkRA01awNNgKxGg607vl5qIvuOAN3pW9HXmD' ):
"""load_covid19(url)
DESCRIPTION:
The function reads in an API URL for COVID-19 Data and returns a pandas DataFrame.
PARAMETERS:
url (string): API URL.
RETURNS:
(DataFrame): Data in a form of a pandas dataframe.
"""
headers = {
'x-api-key': API_KEY
}
response = requests.request("GET", url, headers=headers)
x = response.json()
return pd.DataFrame(x)
```
## **Load Data to DataFrames**
---
Calling functions to lead data
```
# Reading Data to DataFrames
global_view = load_covid19(url_CasesGlobalView)
local_view = load_covid19(url_CasesLocalView)
counterMeasure_view = load_covid19(ulr_CounterMeasureView)
# loading covid 19 symptoms dataframe
symptoms_df = scrape_covid19_symptoms(SYMPTOMS_URL)
# loading external sources
global_confirmed_ = pd.read_csv(CONFIRMED_URL)
global_recorvered_ = pd.read_csv(RECOVERED_URL)
global_deaths_ = pd.read_csv(DEATHS_URL)
```
# **Domain Knowledge & Few Data Checks**
*Quick Descriptive Analysis, Data Values & Data Types*
---
Lets get to understand the **Features** and **Categorical** values.
- The **cardinality of Categorical Features** might not change as the data increases.
- **investigate Categorical Features :**
- country
- province/state/region
- measures
- and more if any
---
- **investigate the Data type per Feature:**
- **Date** (format)
- **floats** stored as **objects** or vice versa
- **Down cast data types** to **pandas data types** which **reduce memory usage**.
---
- **Investigate Feature Values:**
- **Missing Data values**
- missing at Random
- missing by aggregation
- indicated as missing but the actual value is Zero.
- **non-atomic Features & format:**
- Create a new Feature
- Remove part of the data
- Use **iso_** features to correct cuntries.
---
## **CasesGlobalView**
```
# Dimension of the global_view data
print(f'The global Instances are : {global_view.shape[0]}')
print(f'The global Features are : {global_view.shape[1]}')
# view 5 samples of the data
global_view.sample(5)
# understanding global view columns
global_cols = global_view.columns.to_list()
global_cols
# data types per Feature, Memeory size
global_view.info()
# get count for null values
global_view.isnull().mean()
```
**Notes 2** Missing Data:
> We need to understand the Nature of missing values before we can drop the records or impute the values.
> NB: Imputaion / droping records with missing values may yeild misleading results and may affect the interpretation of the pandemic outbreak.
The following columns have more less **10%** missing values out of current loaded observations :
* confirmed_daily
* deaths_daily
* recovered_daily
* daily_change_in_active_cases
* active_dailiy_growth_rate
* active_rolling_3_day_growth_rate
```
# Eliminate the Logitude and latitude the view the summary statistics
global_view.drop(['lat','long'], axis=1).describe()
```
**Notes 3**
> Understanding the **mean**, **std** and **MAx** for the following features/ Variables:
- **confirmed**
- **deaths**
- **recovered**
> **Ideal Scenario**
* The **std** and **mean** for **confirmed** and **deaths** are expected to have a very high variation, this will mean that the virus **preading-rate** and **death-rate** are droping(flaterning the curve).
* The **std** and **mean** for **Recovered** are expected to have a less to no variation and this will mean that more people are recovering at a short period of time.[[1]('')]
```
# get number of countries
global_view['country'].nunique()
```
**notes #**
we got **185 countries**, this should be considered when ploting vs country.
## **CasesLocalView**
```
# Dimension of the global_view data
print(f'The local Instances are : {local_view.shape[0]}')
print(f'The local Features are : {local_view.shape[1]}')
# view 5 samples of the local view data
local_view.sample(5)
# data types
local_view.info()
# get count for Missing Data
local_view.isnull().mean()
# Remove the id & country_id Features since they are not informative on the summary statistics
local_view.drop(['id','country_id'], axis=1).describe()
# group by location and get the size
local_view.groupby('location').size()
```
## **CounterMeasureView**
```
# Dimension of the global_view data
print(f'The Counter Measure Instances are : {counterMeasure_view.shape[0]}')
print(f'The Counter Measure Features are : {counterMeasure_view.shape[1]}')
# view 5 samples of the Counter Measure View data
counterMeasure_view.sample(5)
# data type per Feature and memory size
counterMeasure_view.info()
# cheget the % of missing values
counterMeasure_view.isnull().mean()
# Summary statisctics
counterMeasure_view.describe()
# investigate the number of measures
counterMeasure_view['measure'].nunique()
```
**Notes #**
we got **23 measures**,So need to be careful when using scatter plots and not cluster by measures.
## **Symptoms Data**
```
# Dimension of the symptoms_df data
print(f'The Symptoms Instances are : {symptoms_df.shape[0]}')
print(f'The Symptoms Features are : {symptoms_df.shape[1]}')
# view data samples
symptoms_df
```
clean up the % and extra numbers
```
# data types
symptoms_df.info()
```
cast **percentage** to be a float not object
# **Data Preprocessing**
* Cast the DATATYPES to reduce memory usage
* Create pipeline To clean the Data
* Pass all the data Stream through the pipeline
## **LOGIC**
---
custorm Functions to preprocess all the data
```
def preprocess_symptoms_df(df):
# preprocening the dataframe
df.replace(regex=['%\[69]', '%\[71]'], value='', inplace=True)
# cast object to float and reduce memory usage
df.covid19_percentage = df.covid19_percentage.astype('float')
return df
```
## **PREPROCESING**
---
```
symptoms_df = preprocess_symptoms_df(symptoms_df)
symptoms_df
```
# **Visualizations**
*Lets plan the charts so as to better group them based on the information then convey.*
---
**Groups of Plots:**
1. Summary plots e.g pie charts
2. Trends plots (Time series)
2. Distribution plots
3. Relationship plots
---
1. **Trends:**
- **investigate by date of report** (chart Type = ?) time series
- Total deaths, recovered and confirmed vs date of report
- daily deaths, recovered and confirmed vs date of report
---
2. **Distribution plots:**
- **investigate by country**
- deaths vs country
- recovered vs country
- confirmed vs country
- **investigate by province**
- **investigate by measures**
---
```
## plotting the symptoms VS covid19_percentage
plt.title('plotting the symptoms VS covid19_percentage')
sns.barplot(x=symptoms_df['symptoms'],y=symptoms_df['covid19_percentage']);
plt.xticks(rotation=60)
```
**Interpretation:**
---
*According to our dataset, a patiant suffering from fever has a high probability of having covid19.*
# **Future Tasks**
- creating a PipeLine Api to pull all the data to our open source covid 19 data repo.
- creating dynamic visuals (power Bi/ Data Studio/ tablue)
- hosting them on the internet
- creating ML and DL models to continuesly train and predict the trend of covid 19
- perform batch and online training for our model.
- plot and publish the model results and its logs.
- Automating the pipeline API for CI and CD life cycle.
- contribute to UCI data portal or any other portal
- Helping people Study the data easy
### You're Welcome to Join Our GitHub Community.
# **Credits & Sources**
*We would like to Acknowledge the sources listed below for data, Inpiration and domain knowledge.*
---
### **Data Sources** with data updates time
1. [westerncape.gov.za](https://www.westerncape.gov.za/department-of-health/coronavirus)
2. [Data Science for Social Impact research group](https://dsfsi.github.io/)
3. [CSSE at Johns Hopkins University](https://systems.jhu.edu/)
4. [ourworldindata.org](https://ourworldindata.org/coronavirus)
---
### **Blogs**
1. [The National Center for Health Statistics]('https://nces.ed.gov/')
---
### **Books**
1. books here.
---
### **Research papers**
1. papers here.
---
# THANK YOU
Please post any questions on our github project by creating an **issue** and if you found this project informative, give us a **star** and **follow** our ***github organization & profiles***.

---
| github_jupyter |
```
# -*- coding: utf-8 -*-
import re
from tqdm import tqdm
import time
from datetime import datetime
import sqlite3
import sys
import os
import pandas as pd
import unify
def connect(file_path, primary, columns):
con = sqlite3.connect(file_path)
cur = con.cursor()
cols = ", ".join([c + ' Varchar' for c in columns])
cur.execute("create table meta ("+primary+" Varchar PRIMARY KEY, "+cols+" )")
cur.execute("CREATE INDEX log on meta (id);")
cur.execute("create table plain_texts (id Varchar(128) NOT NULL PRIMARY KEY, text Varchar NOT NULL);")
cur.execute("create table tagged_texts (id Varchar(128) NOT NULL PRIMARY KEY, text Varchar NOT NULL );")
con.commit()
return con, cur
workdir = r'/home/tari/Загрузки/taiga/Fontanka/Fontanka'
filename = 'Fontanka.db'
file_path = os.path.join(workdir, filename)
metatablepathlist = sorted([os.path.join(workdir,file) for file in os.listdir(workdir) if 'metatable' in file])
years = [str(y) for y in range(2005,2018)]
tagged = os.path.join(workdir,'texts_tagged')
plain = os.path.join(workdir,'texts')
meta = pd.read_csv(metatablepathlist[0], sep='\t', encoding='utf8')
meta = meta.fillna('')
meta.head()
meta.columns
if not os.path.exists(filename):
con, cur = connect(filename, meta.columns[1], [meta.columns[0]]+list(meta.columns[2:]))
else:
con = sqlite3.connect(filename, meta.columns[1], [meta.columns[0]]+list(meta.columns[2:]))
cur = con.cursor()
meta.iloc[7].to_dict()
for i in tqdm(range(len(meta))):
values = meta.iloc[i].to_dict()
columns = ', '.join(values.keys())
#print(list(values.values()))
placeholders = ', '.join('?' * len(values))
sql = 'INSERT INTO meta ({}) VALUES ({})'.format(columns, placeholders)
#print(sql)
try:
cur.execute(sql, list(values.values()))
valuest = {'id': values['textid'], 'text': unify.open_text(os.path.join(os.path.join(plain,years[0]), str(values['textid'])+".txt"))}
columns = ', '.join(valuest.keys())
placeholders = ', '.join('?' * len(valuest))
sql2 = 'INSERT INTO plain_texts ({}) VALUES ({})'.format(columns, placeholders)
cur.execute(sql2, list(valuest.values()))
try:
valuest2 = {'id': values['textid'], 'text': unify.open_text(open(os.path.join(os.path.join(tagged,years[0]), str(values['textid'])+".txt"),'r', encoding='utf8').read())}
columns = ', '.join(valuest2.keys())
placeholders = ', '.join('?' * len(valuest2))
sql3 = 'INSERT INTO tagged_texts ({}) VALUES ({})'.format(columns, placeholders)
cur.execute(sql3, list(valuest2.values()))
except:
valuest2 = {'id': values['textid'], 'text': ""}
columns = ', '.join(valuest2.keys())
placeholders = ', '.join('?' * len(valuest2))
sql3 = 'INSERT INTO tagged_texts ({}) VALUES ({})'.format(columns, placeholders)
cur.execute(sql3, list(valuest2.values()))
con.commit()
except:
continue
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import xarray as xr
import netCDF4
```
## Read paraminfo csv file
```
csvfile = '../parameter_sets/OAAT_csv_testing.csv'
data = pd.read_csv(csvfile, names=['paramkey', 'paramname', 'perturblev', 'paramval'])
data
```
## Testing out reading parameter/ensemble information from saved csv file
```
# paramval entries for single values
data.paramval[data.paramname=='jmaxb1']
# specify parameter and perturb level
jmaxb1min = data.paramval[(data.paramname=='jmaxb1') & (data.perturblev=='min')]
jmaxb1min
# extra the data (is there another way?)
# convert from str to float
float(jmaxb1min.iloc[0])
# paramval entries for PFT-varying
data.paramval[data.paramname=='dleaf']
# paramval entry for dleaf min
dleafmin = data.paramval[(data.paramname=='dleaf') & (data.perturblev=='min')]
dleafmin.iloc[0]
# trying to convert string to np
# need to work on this...there is weird spacing and line breaks due to the way this data was saved out
np.fromstring(dleafmin.iloc[0], dtype='float', sep=' ')
# paramnval entries for dependencies
data.paramval[data.paramname=="['lf_fcel', 'lf_flab', 'lf_flig']"]
# paramval entry for LF (flag) min
lfmin = data.paramval[(data.paramname=="['lf_fcel', 'lf_flab', 'lf_flig']") & (data.perturblev=='min')]
lfmin.iloc[0]
# trying to convert string to np
# what should be the sep argument in order to parse?
np.fromstring(lfmin.iloc[0], dtype='float', sep='"["')
# paramval entry for KCN (flag) min
# even more complex because the individual parameters are PFT-varying
kcnmin = data.paramval[(data.paramname=="['kc_nonmyc', 'kn_nonmyc', 'akc_active', 'akn_active', 'ekc_active', 'ekn_active']") & (data.perturblev=='min')]
kcnmin.iloc[0]
# for these parameters, value somehow has units attached (??)
#UPDATE: fixed 4/9/21
data.paramval[data.paramname=='ndays_off']
data.paramval[data.paramname=='ndays_on']
# let's check how the default value for this param is read in
basepftfile='/glade/p/cgd/tss/people/oleson/modify_param/ctsm51_params.c210217_kwo.c210222.nc'
def_params = xr.open_dataset(basepftfile)
# these parameters are interpreted by xarray as times in nanoseconds!
ndays_off_def = def_params['ndays_off'].values
ndays_off_def
# the other parameters are just arrays
def_params['som_diffus'].values
# use decode_times=False to get rid of this behavior
def_params_2 = xr.open_dataset(basepftfile, decode_times=False)
def_params_2['ndays_off'].values
# netCDF4 doesn't seem to have this problem (must be xarray interpretation)
dset = netCDF4.Dataset(basepftfile,'r')
dset['ndays_off'][:]
# get the shape (non-variant params)
dset['kc_nonmyc'][:]
len(dset['kc_nonmyc'][:].shape)
# get the shape (variant params)
dset['rootprof_beta'][:]
dset['rootprof_beta'][0,:]
len(dset['rootprof_beta'][:].shape)
# look at the perturbed file
ndays_off_paramfile = '/glade/scratch/kdagon/CLM5PPE/paramfiles/wrongndays/OAAT0245.nc'
ndays_off_params = xr.open_dataset(ndays_off_paramfile)
ndays_off_params['ndays_off'].values
ndays_off_params_2 = netCDF4.Dataset(ndays_off_paramfile,'r')
ndays_off_params_2['ndays_off'][:]
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Name" data-toc-modified-id="Name-1"><span class="toc-item-num">1 </span>Name</a></span></li><li><span><a href="#Search" data-toc-modified-id="Search-2"><span class="toc-item-num">2 </span>Search</a></span><ul class="toc-item"><li><span><a href="#Load-Cached-Results" data-toc-modified-id="Load-Cached-Results-2.1"><span class="toc-item-num">2.1 </span>Load Cached Results</a></span></li><li><span><a href="#Build-Model-From-Google-Images" data-toc-modified-id="Build-Model-From-Google-Images-2.2"><span class="toc-item-num">2.2 </span>Build Model From Google Images</a></span></li></ul></li><li><span><a href="#Analysis" data-toc-modified-id="Analysis-3"><span class="toc-item-num">3 </span>Analysis</a></span><ul class="toc-item"><li><span><a href="#Gender-cross-validation" data-toc-modified-id="Gender-cross-validation-3.1"><span class="toc-item-num">3.1 </span>Gender cross validation</a></span></li><li><span><a href="#Face-Sizes" data-toc-modified-id="Face-Sizes-3.2"><span class="toc-item-num">3.2 </span>Face Sizes</a></span></li><li><span><a href="#Screen-Time-Across-All-Shows" data-toc-modified-id="Screen-Time-Across-All-Shows-3.3"><span class="toc-item-num">3.3 </span>Screen Time Across All Shows</a></span></li><li><span><a href="#Appearances-on-a-Single-Show" data-toc-modified-id="Appearances-on-a-Single-Show-3.4"><span class="toc-item-num">3.4 </span>Appearances on a Single Show</a></span></li></ul></li><li><span><a href="#Persist-to-Cloud" data-toc-modified-id="Persist-to-Cloud-4"><span class="toc-item-num">4 </span>Persist to Cloud</a></span><ul class="toc-item"><li><span><a href="#Save-Model-to-Google-Cloud-Storage" data-toc-modified-id="Save-Model-to-Google-Cloud-Storage-4.1"><span class="toc-item-num">4.1 </span>Save Model to Google Cloud Storage</a></span></li><li><span><a href="#Save-Labels-to-DB" data-toc-modified-id="Save-Labels-to-DB-4.2"><span class="toc-item-num">4.2 </span>Save Labels to DB</a></span><ul class="toc-item"><li><span><a href="#Commit-the-person-and-labeler" data-toc-modified-id="Commit-the-person-and-labeler-4.2.1"><span class="toc-item-num">4.2.1 </span>Commit the person and labeler</a></span></li><li><span><a href="#Commit-the-FaceIdentity-labels" data-toc-modified-id="Commit-the-FaceIdentity-labels-4.2.2"><span class="toc-item-num">4.2.2 </span>Commit the FaceIdentity labels</a></span></li></ul></li></ul></li></ul></div>
```
from esper.prelude import *
from esper.identity import *
from esper import embed_google_images
```
# Name
Please add the person's name and their expected gender below (Male/Female).
```
name = 'Elizabeth Prann'
gender = 'Female'
```
# Search
## Load Cached Results
Reads cached identity model from local disk. Run this if the person has been labelled before and you only wish to regenerate the graphs. Otherwise, if you have never created a model for this person, please see the next section.
```
assert name != ''
results = FaceIdentityModel.load(name=name)
imshow(tile_imgs([cv2.resize(x[1][0], (200, 200)) for x in results.model_params['images']], cols=10))
plt.show()
plot_precision_and_cdf(results)
```
## Build Model From Google Images
Run this section if you do not have a cached model and precision curve estimates. This section will grab images using Google Image Search and score each of the faces in the dataset. We will interactively build the precision vs score curve.
It is important that the images that you select are accurate. If you make a mistake, rerun the cell below.
```
assert name != ''
# Grab face images from Google
img_dir = embed_google_images.fetch_images(name)
# If the images returned are not satisfactory, rerun the above with extra params:
# query_extras='' # additional keywords to add to search
# force=True # ignore cached images
face_imgs = load_and_select_faces_from_images(img_dir)
face_embs = embed_google_images.embed_images(face_imgs)
assert(len(face_embs) == len(face_imgs))
reference_imgs = tile_imgs([cv2.resize(x[0], (200, 200)) for x in face_imgs if x], cols=10)
def show_reference_imgs():
print('User selected reference images for {}.'.format(name))
imshow(reference_imgs)
plt.show()
show_reference_imgs()
# Score all of the faces in the dataset (this can take a minute)
face_ids_by_bucket, face_ids_to_score = face_search_by_embeddings(face_embs)
precision_model = PrecisionModel(face_ids_by_bucket)
```
Now we will validate which of the images in the dataset are of the target identity.
__Hover over with mouse and press S to select a face. Press F to expand the frame.__
```
show_reference_imgs()
print(('Mark all images that ARE NOT {}. Thumbnails are ordered by DESCENDING distance '
'to your selected images. (The first page is more likely to have non "{}" images.) '
'There are a total of {} frames. (CLICK THE DISABLE JUPYTER KEYBOARD BUTTON '
'BEFORE PROCEEDING.)').format(
name, name, precision_model.get_lower_count()))
lower_widget = precision_model.get_lower_widget()
lower_widget
show_reference_imgs()
print(('Mark all images that ARE {}. Thumbnails are ordered by ASCENDING distance '
'to your selected images. (The first page is more likely to have "{}" images.) '
'There are a total of {} frames. (CLICK THE DISABLE JUPYTER KEYBOARD BUTTON '
'BEFORE PROCEEDING.)').format(
name, name, precision_model.get_lower_count()))
upper_widget = precision_model.get_upper_widget()
upper_widget
```
Run the following cell after labelling to compute the precision curve. Do not forget to re-enable jupyter shortcuts.
```
# Compute the precision from the selections
lower_precision = precision_model.compute_precision_for_lower_buckets(lower_widget.selected)
upper_precision = precision_model.compute_precision_for_upper_buckets(upper_widget.selected)
precision_by_bucket = {**lower_precision, **upper_precision}
results = FaceIdentityModel(
name=name,
face_ids_by_bucket=face_ids_by_bucket,
face_ids_to_score=face_ids_to_score,
precision_by_bucket=precision_by_bucket,
model_params={
'images': list(zip(face_embs, face_imgs))
}
)
plot_precision_and_cdf(results)
```
The next cell persists the model locally.
```
results.save()
```
# Analysis
## Gender cross validation
Situations where the identity model disagrees with the gender classifier may be cause for alarm. We would like to check that instances of the person have the expected gender as a sanity check. This section shows the breakdown of the identity instances and their labels from the gender classifier.
```
gender_breakdown = compute_gender_breakdown(results)
print('Expected counts by gender:')
for k, v in gender_breakdown.items():
print(' {} : {}'.format(k, int(v)))
print()
print('Percentage by gender:')
denominator = sum(v for v in gender_breakdown.values())
for k, v in gender_breakdown.items():
print(' {} : {:0.1f}%'.format(k, 100 * v / denominator))
print()
```
Situations where the identity detector returns high confidence, but where the gender is not the expected gender indicate either an error on the part of the identity detector or the gender detector. The following visualization shows randomly sampled images, where the identity detector returns high confidence, grouped by the gender label.
```
high_probability_threshold = 0.8
show_gender_examples(results, high_probability_threshold)
```
## Face Sizes
Faces shown on-screen vary in size. For a person such as a host, they may be shown in a full body shot or as a face in a box. Faces in the background or those part of side graphics might be smaller than the rest. When calculuating screentime for a person, we would like to know whether the results represent the time the person was featured as opposed to merely in the background or as a tiny thumbnail in some graphic.
The next cell, plots the distribution of face sizes. Some possible anomalies include there only being very small faces or large faces.
```
plot_histogram_of_face_sizes(results)
```
The histogram above shows the distribution of face sizes, but not how those sizes occur in the dataset. For instance, one might ask why some faces are so large or whhether the small faces are actually errors. The following cell groups example faces, which are of the target identity with probability, by their sizes in terms of screen area.
```
high_probability_threshold = 0.8
show_faces_by_size(results, high_probability_threshold, n=10)
```
## Screen Time Across All Shows
One question that we might ask about a person is whether they received a significantly different amount of screentime on different shows. The following section visualizes the amount of screentime by show in total minutes and also in proportion of the show's total time. For a celebrity or political figure such as Donald Trump, we would expect significant screentime on many shows. For a show host such as Wolf Blitzer, we expect that the screentime be high for shows hosted by Wolf Blitzer.
```
screen_time_by_show = get_screen_time_by_show(results)
plot_screen_time_by_show(name, screen_time_by_show)
```
## Appearances on a Single Show
For people such as hosts, we would like to examine in greater detail the screen time allotted for a single show. First, fill in a show below.
```
show_name = 'Americas News HQ'
# Compute the screen time for each video of the show
screen_time_by_video_id = compute_screen_time_by_video(results, show_name)
```
One question we might ask about a host is "how long they are show on screen" for an episode. Likewise, we might also ask for how many episodes is the host not present due to being on vacation or on assignment elsewhere. The following cell plots a histogram of the distribution of the length of the person's appearances in videos of the chosen show.
```
plot_histogram_of_screen_times_by_video(name, show_name, screen_time_by_video_id)
```
For a host, we expect screentime over time to be consistent as long as the person remains a host. For figures such as Hilary Clinton, we expect the screentime to track events in the real world such as the lead-up to 2016 election and then to drop afterwards. The following cell plots a time series of the person's screentime over time. Each dot is a video of the chosen show. Red Xs are videos for which the face detector did not run.
```
plot_screentime_over_time(name, show_name, screen_time_by_video_id)
```
We hypothesized that a host is more likely to appear at the beginning of a video and then also appear throughout the video. The following plot visualizes the distibution of shot beginning times for videos of the show.
```
plot_distribution_of_appearance_times_by_video(results, show_name)
```
In the section 3.3, we see that some shows may have much larger variance in the screen time estimates than others. This may be because a host or frequent guest appears similar to the target identity. Alternatively, the images of the identity may be consistently low quality, leading to lower scores. The next cell plots a histogram of the probabilites for for faces in a show.
```
plot_distribution_of_identity_probabilities(results, show_name)
```
# Persist to Cloud
The remaining code in this notebook uploads the built identity model to Google Cloud Storage and adds the FaceIdentity labels to the database.
## Save Model to Google Cloud Storage
```
gcs_model_path = results.save_to_gcs()
```
To ensure that the model stored to Google Cloud is valid, we load it and print the precision and cdf curve below.
```
gcs_results = FaceIdentityModel.load_from_gcs(name=name)
imshow(tile_imgs([cv2.resize(x[1][0], (200, 200)) for x in gcs_results.model_params['images']], cols=10))
plt.show()
plot_precision_and_cdf(gcs_results)
```
## Save Labels to DB
If you are satisfied with the model, we can commit the labels to the database.
```
from django.core.exceptions import ObjectDoesNotExist
def standardize_name(name):
return name.lower()
person_type = ThingType.objects.get(name='person')
try:
person = Thing.objects.get(name=standardize_name(name), type=person_type)
print('Found person:', person.name)
except ObjectDoesNotExist:
person = Thing(name=standardize_name(name), type=person_type)
print('Creating person:', person.name)
labeler = Labeler(name='face-identity:{}'.format(person.name), data_path=gcs_model_path)
```
### Commit the person and labeler
The labeler and person have been created but not set saved to the database. If a person was created, please make sure that the name is correct before saving.
```
person.save()
labeler.save()
```
### Commit the FaceIdentity labels
Now, we are ready to add the labels to the database. We will create a FaceIdentity for each face whose probability exceeds the minimum threshold.
```
commit_face_identities_to_db(results, person, labeler, min_threshold=0.001)
print('Committed {} labels to the db'.format(FaceIdentity.objects.filter(labeler=labeler).count()))
```
| github_jupyter |
# Stochastic Block Models (SBM)
Let's imagine that we have $100$ students, each of whom can go to one of two possible schools: school one or school two. Our network has $100$ nodes, and each node represents a single student. The edges of this network represent whether a pair of students are friends. Intuitively, if two students go to the same school, the probably have a higher chance of being friends than if they do not go to the same school. If we were to try to characterize this using an ER random network, we would run into a problem: we have no way to capture the impact that school has on friendships. Intuitively, there must be a better way!
The Stochastic Block Model, or SBM, captures this idea by assigning each of the $n$ nodes in the network to one of $K$ communities. A **community** is a group of nodes within the network. In our example case, the communities would represent the schools that students are able to attend. We use $K$ here to just denote an integer greater than $1$ (for example, in the school example we gave above, $K$ is $2$) for the number of *possible* communities that nodes could be members of. In an SBM, instead of describing all pairs of nodes with a fixed probability like with the ER model, we instead describe properties that hold for edges between *pairs of communities*. In our example, what this means is that if two students go to school one, the probability that they are friends might be different than if the two students went to school two, or if one student went to school one and the other to school two. Let's take a look at what a realization of this setup we have described might look like:
```
from graphbook_code import draw_multiplot
from graspologic.simulations import sbm
ns = [50, 50] # network with 50 nodes
B = [[0.6, 0.2], [0.2, 0.6]] # probability of an edge existing is .3
# sample a single simple adjacency matrix from ER(50, .3)
A = sbm(n=ns, p=B, directed=False, loops=False)
ys = [1 for i in range(0, 50)] + [2 for i in range(0, 50)]
draw_multiplot(A, labels=ys, title="$SBM_n(\\tau, B)$ Simulation");
```
To describe an SBM random network, we proceed very similarly to an ER random network, with a twist. An SBM random network has a parameter, $\vec\tau$, which has a single element for each of the node. We call $\vec\tau$ the **community assignment vector**, which means that for each node of our random network, $\tau_i$ tells us which community the node is in. To state this another way, $\vec\tau$ is a vector where each element $\tau_i$ can take one of $K$ possible values, where $K$ is the total number of communities in the network. For example, if we had an SBM random network with four nodes in total, and two total communities, each element $\tau_i$ can be either $1$ or $2$. If the first two nodes were in community $1$, and the second two in community $2$, we would say that $\tau_1 = 1$, $\tau_2 = 1$, $\tau_3 = 2$, and $\tau_4 = 2$, which means that $\vec\tau$ looks like:
\begin{align*}
\vec\tau^\top &= \begin{bmatrix}1 & 1 & 2 & 2\end{bmatrix}
\end{align*}
The other parameter for an SBM random network is called the block matrix, for which we will use the capital letter $B$. If there are $K$ communities in the SBM random network, then $B$ is a $K \times K$ matrix, with one entry for each pair of communities. For instance, if $K$ were two like above, $B$ would be a $2 \times 2$ matrix, and would look like this:
\begin{align*}
B &= \begin{bmatrix}
b_{11} & b_{12} \\ b_{21} & b_{22}
\end{bmatrix}
\end{align*}
Each of the entries of $B$, which we denote as $b_{kl}$ in the above matrix, is a probability of an edge existing between a node in community $k$ and a node in community $l$.
Fortunately, we can also think of this formulation of a random network using coin flips. In our mini example above, if node $1$ is in community $1$ (since $\tau_1 = 1$) and node $2$ is in community $1$ (since $\tau_2 = 1$), we have a weighted coin which has a probability $b_{11}$ (the first row, first column of the block matrix above) of landing on heads, and a $1 - b_{11}$ chance of landing on tails. An edge between nodes one and two exists if the weighted coin lands on heads, and does not exist if that weighted coin lands on tails. If we wanted to describe an edge between nodes one and three instead, note that $\tau_3 = 2$. Therefore, we use the entry $b_{12}$ as the probability of obtaining a heads for the weighted coin we flip this time. In the general case, to use the block matrix to obtain the probability of an edge $(i, j)$ existing between any pair of nodes $i$ and $j$ in our network, we will flip a coin with probability $b_{\tau_i \tau_j}$, where $\tau_i$ is the community assignment for the $i^{th}$ node and $\tau_j$ is the community assignment for the $j^{th}$ node.
If $\mathbf A$ is a random network which is an $SBM_n(\vec \tau, B)$ with $n$ nodes, the community vector $\vec\tau$, and the block matrix $B$, we say that $\mathbf A$ is an $SBM_n(\vec\tau, B)$ random network.
The procedure below will produce for us a network $A$, which has nodes and edges, where the underlying random network $\mathbf A$ is an $SBM_n(\vec\tau, B)$ random network:
```{admonition}
1. Determine a community assignment vector, $\vec\tau$, for each of the $n$ nodes. Each node should be assigned to one of $K$ communities.
2. Determine a block matrix, $B$, for each pair of the $K$ communities.
3. For each pair of communities $k$ and $l$, obtain a weighted coin (which we will call the $(k,l)$ coin) which as a $b_{kl}$ chance of landing on heads, and a $1 - b_{kl}$ chance of landing on tails.
4. For each pair of nodes $i$ and $j$:
a. Denote $\tau_i$ to be the community assignment of node $i$, and $\tau_j$ to be the community assignment of node $j$.
b. Flip the $(\tau_i, \tau_j)$ coin, and if it lands on heads, the corresponding entry $a_{ij}$ in the adjacency matrix is $1$. If it lands on tails, the corresponding entry $a_{ij}$ in the adjacency matrix is $0$.
5. The adjacency matrix we produce, $A$, is a realization of an $SBM_n(\vec\tau, B)$ random network.
```
### Code Examples
We just covered a lot of intuition! This intuition will come in handy later, but let's take a break from the theory by working through an example. Let's use the school example we started above. Say we have $100$ students, and we know that each student goes to one of two possible schools. Remember that we already know the community assignment vector $\vec{\tau}$ ahead of time. We don't really care too much about the ordering of the students for now, so let's just assume that the first $50$ students all go to the first school, and the second $50$ students all go to the second school.
```{admonition} Thought Exercise
Before you read on, try to think to yourself about what the node-assignment vector $\vec \tau$ looks like.
```
Next, let's plot what $\vec \tau$ look like:
```
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import matplotlib
def plot_tau(tau, title="", xlab="Node"):
cmap = matplotlib.colors.ListedColormap(["skyblue", 'blue'])
fig, ax = plt.subplots(figsize=(10,2))
with sns.plotting_context("talk", font_scale=1):
ax = sns.heatmap((tau - 1).reshape((1,tau.shape[0])), cmap=cmap,
ax=ax, cbar_kws=dict(shrink=1), yticklabels=False,
xticklabels=False)
ax.set_title(title)
cbar = ax.collections[0].colorbar
cbar.set_ticks([0.25, .75])
cbar.set_ticklabels(['School 1', 'School 2'])
ax.set(xlabel=xlab)
ax.set_xticks([.5,49.5,99.5])
ax.set_xticklabels(["1", "50", "100"])
cbar.ax.set_frame_on(True)
return
n = 100 # number of students
# tau is a column vector of 150 1s followed by 50 2s
# this vector gives the school each of the 300 students are from
tau = np.vstack((np.ones((int(n/2),1)), np.full((int(n/2),1), 2)))
plot_tau(tau, title="Tau, Node Assignment Vector",
xlab="Student")
```
So as we can see, the first $50$ students are from the first school, and the second $50$ students are from second school.
Let's assume that the students from the first school are better friends in general than the students from the second school, so we'll say that the probability of two students who both go to the first school being friends is $0.5$, and the probability of two students who both go to school $2$ being friends is $0.3$. Finally, let's assume that if one student goes to the first school and the other student goes to school $2$, that the probability that they are friends is $0.2$.
```{admonition} Thought Exercise
Before you read on, try to think to yourself about what the block matrix $B$ looks like.
```
Next, let's look at the block matrix $B$:
```
K = 2 # 2 communities in total
# construct the block matrix B as described above
B = [[0.6, 0.2],
[0.2, 0.4]]
def plot_block(X, title="", blockname="School", blocktix=[0.5, 1.5],
blocklabs=["School 1", "School 2"]):
fig, ax = plt.subplots(figsize=(8, 6))
with sns.plotting_context("talk", font_scale=1):
ax = sns.heatmap(X, cmap="Purples",
ax=ax, cbar_kws=dict(shrink=1), yticklabels=False,
xticklabels=False, vmin=0, vmax=1, annot=True)
ax.set_title(title)
cbar = ax.collections[0].colorbar
ax.set(ylabel=blockname, xlabel=blockname)
ax.set_yticks(blocktix)
ax.set_yticklabels(blocklabs)
ax.set_xticks(blocktix)
ax.set_xticklabels(blocklabs)
cbar.ax.set_frame_on(True)
return
plot_block(B, title="Block Matrix")
plt.show()
```
As we can see, the matrix $B$ is a symmetric block matrix, since our network is undirected. Finally, let's sample a single network from the $SBM_n(\vec\tau, B)$ with parameters $\vec \tau$ and $B$:
```
from graspologic.simulations import sbm
from graphbook_code import draw_multiplot
import pandas as pd
# sample a graph from SBM_{300}(tau, B)
A = sbm(n=[int(n/2), int(n/2)], p=B, directed=False, loops=False)
ys = [1 for i in range(0, 50)] + [2 for i in range(0, 50)]
draw_multiplot(A, labels=ys, title="$SBM_n(\\tau, B)$ Simulation");
```
The above network shows students, ordered by the school they are in (first school and the second school, respectively). As we can see in the above network, people from the first school are more connected than people from school $2$. Also, the connections between people from different schools (the *off-diagonal* blocks of the adjacency matrix, the lower left and upper right blocks) appear to be a bit *more sparse* (fewer edges) than connections betwen schools (the *on-diagonal* blocks of the adjacency matrix, the upper left and lower right blocks). The above heatmap can be described as **modular**: it has clear communities. Remember that the connections for each node are indicated by a single row, or a single column, of the adjacency matrix. The first half of the rows have strong connections with the first half of the columns, which indicates that the first half of students tend to be better friends with other students in the first half. We can duplicate this argument for the second half of students ot see that it seems reasonable to conclude that there are two communities of students here.
Something easy to mistake about a realization of an $SBM_n(\vec\tau, B)$ is that the realizations will *not always* have the obvious modular structure we can see above when we look at a heatmap. Rather, this modular structure is *only* made obvious because the students are ordered according to the school in which they are in. What do you think will happen if we look at the students in a random order? Do you think that he structure that exists in this network will be obvious?
The answer is: *No!* Let's see what happens when we reorder the nodes from the network into a random order, and pretend we don't know the true community labels ahead of time:
```
import numpy as np
# generate a reordering of the n nodes
vtx_perm = np.random.choice(n, size=n, replace=False)
Aperm = A[tuple([vtx_perm])] [:,vtx_perm]
yperm = np.array(ys)[vtx_perm]
draw_multiplot(Aperm, title="$SBM_n(\\tau, B)$ Simulation");
```
Now, the students are *not* organized according to school, because they have been randomly reordered. It becomes pretty tough to figure out whether there are communities just by looking at an adjacency matrix, unless you are looking at a network in which the nodes are *already arranged* in an order which respects the community structure. By an *order that respects the community structure*, we mean that the community assignment vector $\vec\tau$ is arranged so that all of the nodes in the first community come first, followed by all of the nodes in the second community, followed by all of the nodes in the third community, so on and so forth up to the nodes of the community $K$.
In practice, this means that if you know ahead of time what natural groupings of the nodes might be (such as knowing which school each student goes to) by way of your node attributes, you can visualize your data according to that grouping. If you don't know anything about natural groupings of nodes, however, we are left with the problem of *estimating community structure*. A later method, called the *spectral embedding*, will be paired with clustering techniques to allow us to estimate node assignment vectors.
| github_jupyter |
<a href="https://cognitiveclass.ai"><img src = "https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/Logos/organization_logo/organization_logo.png" width = 400> </a>
<h1 align=center><font size = 5>Pre-Trained Models</font></h1>
```
from google.colab import drive
drive.mount('/content/drive')
cd /content/drive/My Drive/AI Capestone
```
## Introduction
In this lab, you will learn how to leverage pre-trained models to build image classifiers instead of building a model from scratch.
## Table of Contents
<div class="alert alert-block alert-info" style="margin-top: 20px">
<font size = 3>
1. <a href="#item31">Import Libraries and Packages</a>
2. <a href="#item32">Download Data</a>
3. <a href="#item33">Define Global Constants</a>
4. <a href="#item34">Construct ImageDataGenerator Instances</a>
5. <a href="#item35">Compile and Fit Model</a>
</font>
</div>
<a id='item31'></a>
## Import Libraries and Packages
Let's start the lab by importing the libraries that we will be using in this lab.
First, we will import the ImageDataGenerator module since we will be leveraging it to train our model in batches.
```
from keras.preprocessing.image import ImageDataGenerator
```
In this lab, we will be using the Keras library to build an image classifier, so let's download the Keras library.
```
import keras
from keras.models import Sequential
from keras.layers import Dense
```
Finally, we will be leveraging the ResNet50 model to build our classifier, so let's download it as well.
```
from keras.applications import ResNet50
from keras.applications.resnet50 import preprocess_input
```
<a id='item32'></a>
```
print("hi")
```
## Download Data
For your convenience, I have placed the data on a server which you can retrieve easily using the **wget** command. So let's run the following line of code to get the data. Given the large size of the image dataset, it might take some time depending on your internet speed.
```
## get the data
!wget https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0321EN/data/concrete_data_week3.zip
```
And now if you check the left directory pane, you should see the zipped file *concrete_data_week3.zip* appear. So, let's go ahead and unzip the file to access the images. Given the large number of images in the dataset, this might take a couple of minutes, so please be patient, and wait until the code finishes running.
```
!unzip concrete_data_week3.zip
```
Now, you should see the folder *concrete_data_week3* appear in the left pane. If you open this folder by double-clicking on it, you will find that it contains two folders: *train* and *valid*. And if you explore these folders, you will find that each contains two subfolders: *positive* and *negative*. These are the same folders that we saw in the labs in the previous modules of this course, where *negative* is the negative class and it represents the concrete images with no cracks and *positive* is the positive class and it represents the concrete images with cracks.
**Important Note**: There are thousands and thousands of images in each folder, so please don't attempt to double click on the *negative* and *positive* folders. This may consume all of your memory and you may end up with a **50*** error. So please **DO NOT DO IT**.
<a id='item33'></a>
## Define Global Constants
Here, we will define constants that we will be using throughout the rest of the lab.
1. We are obviously dealing with two classes, so *num_classes* is 2.
2. The ResNet50 model was built and trained using images of size (224 x 224). Therefore, we will have to resize our images from (227 x 227) to (224 x 224).
3. We will training and validating the model using batches of 100 images.
```
num_classes = 2
image_resize = 224
batch_size_training = 100
batch_size_validation = 100
```
<a id='item34'></a>
## Construct ImageDataGenerator Instances
In order to instantiate an ImageDataGenerator instance, we will set the **preprocessing_function** argument to *preprocess_input* which we imported from **keras.applications.resnet50** in order to preprocess our images the same way the images used to train ResNet50 model were processed.
```
data_generator = ImageDataGenerator(
preprocessing_function=preprocess_input,
)
p
```
Next, we will use the *flow_from_directory* method to get the training images as follows:
```
train_generator = data_generator.flow_from_directory(
'concrete_data_week3/train',
target_size=(image_resize, image_resize),
batch_size=batch_size_training,
class_mode='categorical')
```
**Your Turn**: Use the *flow_from_directory* method to get the validation images and assign the result to **validation_generator**.
```
## Type your answer here
validation_generator=data_generator.flow_from_directory('concrete_data_week3/valid',target_size=(224,224),batch_size=100,class_mode='categorical')
```
Double-click __here__ for the solution.
<!-- The correct answer is:
validation_generator = data_generator.flow_from_directory(
'concrete_data_week3/valid',
target_size=(image_resize, image_resize),
batch_size=batch_size_validation,
class_mode='categorical')
-->
<a id='item35'></a>
## Build, Compile and Fit Model
In this section, we will start building our model. We will use the Sequential model class from Keras.
```
model = Sequential()
```
Next, we will add the ResNet50 pre-trained model to out model. However, note that we don't want to include the top layer or the output layer of the pre-trained model. We actually want to define our own output layer and train it so that it is optimized for our image dataset. In order to leave out the output layer of the pre-trained model, we will use the argument *include_top* and set it to **False**.
```
model.add(ResNet50(
include_top=False,
pooling='avg',
weights='imagenet',
))
```
Then, we will define our output layer as a **Dense** layer, that consists of two nodes and uses the **Softmax** function as the activation function.
```
model.add(Dense(num_classes, activation='softmax'))
```
You can access the model's layers using the *layers* attribute of our model object.
```
model.layers
```
You can see that our model is composed of two sets of layers. The first set is the layers pertaining to ResNet50 and the second set is a single layer, which is our Dense layer that we defined above.
You can access the ResNet50 layers by running the following:
```
model.layers[0].layers
```
Since the ResNet50 model has already been trained, then we want to tell our model not to bother with training the ResNet part, but to train only our dense output layer. To do that, we run the following.
```
model.layers[0].trainable = False
```
And now using the *summary* attribute of the model, we can see how many parameters we will need to optimize in order to train the output layer.
```
model.summary()
```
Next we compile our model using the **adam** optimizer.
```
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
```
Before we are able to start the training process, with an ImageDataGenerator, we will need to define how many steps compose an epoch. Typically, that is the number of images divided by the batch size. Therefore, we define our steps per epoch as follows:
```
steps_per_epoch_training = len(train_generator)
steps_per_epoch_validation = len(validation_generator)
num_epochs = 2
```
Finally, we are ready to start training our model. Unlike a conventional deep learning training were data is not streamed from a directory, with an ImageDataGenerator where data is augmented in batches, we use the **fit_generator** method.
```
fit_history = model.fit_generator(
train_generator,
steps_per_epoch=steps_per_epoch_training,
epochs=num_epochs,
validation_data=validation_generator,
validation_steps=steps_per_epoch_validation,
verbose=1,
)
fit_history = model.fit_generator(
train_generator,
steps_per_epoch=steps_per_epoch_training,
epochs=2,
validation_data=validation_generator,
validation_steps=steps_per_epoch_validation,
verbose=1,
)
```
Now that the model is trained, you are ready to start using it to classify images.
Since training can take a long time when building deep learning models, it is always a good idea to save your model once the training is complete if you believe you will be using the model again later. You will be using this model in the next module, so go ahead and save your model.
```
model.save('classifier_resnet_model.h5')
```
Now, you should see the model file *classifier_resnet_model.h5* apprear in the left directory pane.
### Thank you for completing this lab!
This notebook was created by Alex Aklson. I hope you found this lab interesting and educational.
This notebook is part of a course on **Coursera** called *AI Capstone Project with Deep Learning*. If you accessed this notebook outside the course, you can take this course online by clicking [here](https://cocl.us/DL0321EN_Coursera_Week3_LAB1).
<hr>
Copyright © 2020 [IBM Developer Skills Network](https://cognitiveclass.ai/?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
| github_jupyter |
# Introduction to a numpy API for ONNX: CustomClassifier
This notebook shows how to write python classifier using similar functions as numpy offers and get a class which can be inserted into a pipeline and still be converted into ONNX.
```
from jyquickhelper import add_notebook_menu
add_notebook_menu()
%load_ext mlprodict
```
## A custom binary classifier
Let's imagine a classifier not that simple about simple but not that complex about predictions. It does the following:
* compute the barycenters of both classes,
* determine an hyperplan containing the two barycenters of the clusters,
* train a logistic regression on both sides.
Some data first...
```
from sklearn.datasets import make_classification
from pandas import DataFrame
X, y = make_classification(200, n_classes=2, n_features=2, n_informative=2,
n_redundant=0, n_clusters_per_class=2, hypercube=False)
df = DataFrame(X)
df.columns = ['X1', 'X2']
df['y'] = y
ax = df[df.y == 0].plot.scatter(x="X1", y="X2", color="blue", label="y=0")
df[df.y == 1].plot.scatter(x="X1", y="X2", color="red", label="y=1", ax=ax);
```
Split into train and test as usual.
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y)
```
The model...
```
import numpy
from sklearn.base import ClassifierMixin, BaseEstimator
from sklearn.linear_model import LogisticRegression
class TwoLogisticRegression(ClassifierMixin, BaseEstimator):
def __init__(self):
ClassifierMixin.__init__(self)
BaseEstimator.__init__(self)
def fit(self, X, y, sample_weights=None):
if sample_weights is not None:
raise NotImplementedError("weighted sample not implemented in this example.")
# Barycenters
self.weights_ = numpy.array([(y==0).sum(), (y==1).sum()])
p1 = X[y==0].sum(axis=0) / self.weights_[0]
p2 = X[y==1].sum(axis=0) / self.weights_[1]
self.centers_ = numpy.vstack([p1, p2])
# A vector orthogonal
v = p2 - p1
v /= numpy.linalg.norm(v)
x = numpy.random.randn(X.shape[1])
x -= x.dot(v) * v
x /= numpy.linalg.norm(x)
self.hyperplan_ = x.reshape((-1, 1))
# sign
sign = ((X - p1) @ self.hyperplan_ >= 0).astype(numpy.int64).ravel()
# Trains models
self.lr0_ = LogisticRegression().fit(X[sign == 0], y[sign == 0])
self.lr1_ = LogisticRegression().fit(X[sign == 1], y[sign == 1])
return self
def predict_proba(self, X):
sign = self.predict_side(X).reshape((-1, 1))
prob0 = self.lr0_.predict_proba(X)
prob1 = self.lr1_.predict_proba(X)
prob = prob1 * sign - prob0 * (sign - 1)
return prob
def predict(self, X):
prob = self.predict_proba(X)
return prob.argmax(axis=1)
def predict_side(self, X):
return ((X - self.centers_[0]) @ self.hyperplan_ >= 0).astype(numpy.int64).ravel()
model = TwoLogisticRegression()
model.fit(X_train, y_train)
model.predict(X_test)
```
Let's compare the model a single logistic regression. It shouuld be better. The same logistic regression applied on both sides is equivalent a single logistic regression and both half logistic regression is better on its side.
```
from sklearn.metrics import accuracy_score
lr = LogisticRegression().fit(X_train, y_train)
accuracy_score(y_test, lr.predict(X_test)), accuracy_score(y_test, model.predict(X_test))
```
However, this is true in average but not necessarily true for one particular datasets. But that's not the point of this notebook.
```
model.centers_
model.hyperplan_
model.lr0_.coef_, model.lr1_.coef_
```
Let's draw the model predictions. Colored zones indicate the predicted class, green line indicates the hyperplan splitting the features into two. A different logistic regression is applied on each side.
```
import matplotlib.pyplot as plt
def draw_line(ax, v, p0, rect, N=50, label=None, color="black"):
x1, x2, y1, y2 = rect
v = v / numpy.linalg.norm(v) * (x2 - x1)
points = [p0 + v * ((i * 2. / N - 2) + (x1 - p0[0]) / v[0]) for i in range(0, N * 4 + 1)]
arr = numpy.vstack(points)
arr = arr[arr[:, 0] >= x1]
arr = arr[arr[:, 0] <= x2]
arr = arr[arr[:, 1] >= y1]
arr = arr[arr[:, 1] <= y2]
ax.plot(arr[:, 0], arr[:, 1], '.', label=label, color=color)
def zones(ax, model, X):
r = (X[:, 0].min(), X[:, 0].max(), X[:, 1].min(), X[:, 1].max())
h = .02 # step size in the mesh
xx, yy = numpy.meshgrid(numpy.arange(r[0], r[1], h), numpy.arange(r[2], r[3], h))
Z = model.predict(numpy.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
return ax.pcolormesh(xx, yy, Z, cmap=plt.cm.Paired)
fig, ax = plt.subplots(1, 1)
zones(ax, model, X)
df[df.y == 0].plot.scatter(x="X1", y="X2", color="blue", label="y=0", ax=ax)
df[df.y == 1].plot.scatter(x="X1", y="X2", color="red", label="y=1", ax=ax);
rect = (df.X1.min(), df.X1.max(), df.X2.min(), df.X2.max())
draw_line(ax, model.centers_[1] - model.centers_[0], model.centers_[0],
rect, N=100, label="hyperplan", color="green")
ax.legend();
```
## Conversion to ONNX = second implementation
The conversion fails as expected because there is no registered converter for this new model.
```
from skl2onnx import to_onnx
one_row = X_train[:1].astype(numpy.float32)
try:
to_onnx(model, one_row)
except Exception as e:
print(e.__class__.__name__)
print("---")
print(e)
```
Writing a converter means implementing the prediction methods with ONNX operators. That's very similar to learning a new mathematical language even if this language is very close to *numpy*. Instead of having a second implementation of the predictions, why not having a single one based on ONNX? That way the conversion to ONNX would be obvious. Well do you know ONNX operators? Not really... Why not using then numpy functions implemented with ONNX operators? Ok! But how?
## A single implementation with ONNX operators
A classifier needs two pethods, `predict` and `predict_proba` and one graph is going to produce both of them. The user need to implement the function producing this graph, a decorator adds the two methods based on this graph.
```
from mlprodict.npy import onnxsklearn_class
from mlprodict.npy.onnx_variable import MultiOnnxVar
import mlprodict.npy.numpy_onnx_impl as nxnp
import mlprodict.npy.numpy_onnx_impl_skl as nxnpskl
@onnxsklearn_class('onnx_graph')
class TwoLogisticRegressionOnnx(ClassifierMixin, BaseEstimator):
def __init__(self):
ClassifierMixin.__init__(self)
BaseEstimator.__init__(self)
def fit(self, X, y, sample_weights=None):
if sample_weights is not None:
raise NotImplementedError("weighted sample not implemented in this example.")
# Barycenters
self.weights_ = numpy.array([(y==0).sum(), (y==1).sum()])
p1 = X[y==0].sum(axis=0) / self.weights_[0]
p2 = X[y==1].sum(axis=0) / self.weights_[1]
self.centers_ = numpy.vstack([p1, p2])
# A vector orthogonal
v = p2 - p1
v /= numpy.linalg.norm(v)
x = numpy.random.randn(X.shape[1])
x -= x.dot(v) * v
x /= numpy.linalg.norm(x)
self.hyperplan_ = x.reshape((-1, 1))
# sign
sign = ((X - p1) @ self.hyperplan_ >= 0).astype(numpy.int64).ravel()
# Trains models
self.lr0_ = LogisticRegression().fit(X[sign == 0], y[sign == 0])
self.lr1_ = LogisticRegression().fit(X[sign == 1], y[sign == 1])
return self
def onnx_graph(self, X):
h = self.hyperplan_.astype(X.dtype)
c = self.centers_.astype(X.dtype)
sign = ((X - c[0]) @ h) >= numpy.array([0], dtype=X.dtype)
cast = sign.astype(X.dtype).reshape((-1, 1))
prob0 = nxnpskl.logistic_regression( # pylint: disable=E1136
X, model=self.lr0_)[1]
prob1 = nxnpskl.logistic_regression( # pylint: disable=E1136
X, model=self.lr1_)[1]
prob = prob1 * cast - prob0 * (cast - numpy.array([1], dtype=X.dtype))
label = nxnp.argmax(prob, axis=1)
return MultiOnnxVar(label, prob)
model = TwoLogisticRegressionOnnx()
model.fit(X_train, y_train)
model.predict(X_test.astype(numpy.float32))
model.predict_proba(X_test.astype(numpy.float32))[:5]
```
It works with double too.
```
model.predict_proba(X_test.astype(numpy.float64))[:5]
```
And now the conversion to ONNX.
```
onx = to_onnx(model, X_test[:1].astype(numpy.float32),
options={id(model): {'zipmap': False}})
```
Let's check the output.
```
from mlprodict.onnxrt import OnnxInference
oinf = OnnxInference(onx)
oinf.run({'X': X_test[:5].astype(numpy.float32)})
```
| github_jupyter |
# Getting Started
### Libraries Required
* numpy
* pandas
* matplotlib
* seaborn
* wordcloud
* emoji
* jovian (optional)
Install all the above libraries using the command :
```
pip install numpy pandas matplotlib seaborn wordcloud emoji jovian --upgrade
```
```
import re
import jovian
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from wordcloud import WordCloud, STOPWORDS
import emoji
from collections import Counter
```
### WhatsApp Chat Data
* Open WhatsApp
* Open a Group/Inbox
* Click on the 3 dotted options button
* Click on More
* Click on Export Chat
* Click on without media (we are analyzing only text messages here)
* Export via Email/other IM's/....
* Download to your system, rename to chat.txt and put it in a folder.
### Data Processing
```
def rawToDf(file, key):
split_formats = {
'12hr' : '\d{1,2}/\d{1,2}/\d{2,4},\s\d{1,2}:\d{2}\s[APap][mM]\s-\s',
'24hr' : '\d{1,2}/\d{1,2}/\d{2,4},\s\d{1,2}:\d{2}\s-\s',
'custom' : ''
}
datetime_formats = {
'12hr' : '%m/%d/%y, %I:%M %p - ',
'24hr' : '%m/%d/%y, %H:%M - ',
'custom': ''
}
with open(file, 'r', encoding="utf8") as raw_data:
raw_string = ' '.join(raw_data.read().split('\n')) # converting the list split by newline char. as one whole string as there can be multi-line messages
user_msg = re.split(split_formats[key], raw_string) [1:] # splits at all the date-time pattern, resulting in list of all the messages with user names
date_time = re.findall(split_formats[key], raw_string) # finds all the date-time patterns
df = pd.DataFrame({'date_time': date_time, 'user_msg': user_msg}) # exporting it to a df
# converting date-time pattern which is of type String to type datetime,
# format is to be specified for the whole string where the placeholders are extracted by the method
df['date_time'] = pd.to_datetime(df['date_time'], format=datetime_formats[key])
# split user and msg
usernames = []
msgs = []
for i in df['user_msg']:
a = re.split('([\w\W]+?):\s', i) # lazy pattern match to first {user_name}: pattern and spliting it aka each msg from a user
if(a[1:]): # user typed messages
usernames.append(a[1])
msgs.append(a[2])
else: # other notifications in the group(eg: someone was added, some left ...)
usernames.append("grp_notif")
msgs.append(a[0])
# creating new columns
df['user'] = usernames
df['msg'] = msgs
# dropping the old user_msg col.
df.drop('user_msg', axis=1, inplace=True)
return df
```
### Import Data
```
df = rawToDf('chat-data.txt', '12hr')
df.tail()
# no. of msgs
df.shape
me = "Ashutosh Krishna"
```
### Data Cleaning
Let's delete the messages having media. We can see above the media part is omitted.
```
media = df[df['msg']=="<Media omitted> "] #no. of images, images are represented by <media omitted>
media.shape
df["user"].unique()
grp_notif = df[df['user']=="grp_notif"] #no. of grp notifications
grp_notif.shape
df.drop(media.index, inplace=True) #removing images
df.drop(grp_notif.index, inplace=True) #removing grp_notif
df.tail()
df.reset_index(inplace=True, drop=True)
df.shape
```
# Let's Answer Some Questions
## Q. Who are the least active and most active persons in the group?
```
df.groupby("user")["msg"].count().sort_values(ascending=False)
```
## Q. How many emojis I have used?
```
emoji_ctr = Counter()
emojis_list = map(lambda x: ''.join(x.split()), emoji.UNICODE_EMOJI.keys())
r = re.compile('|'.join(re.escape(p) for p in emojis_list))
for idx, row in df.iterrows():
if row["user"] == me:
emojis_found = r.findall(row["msg"])
for emoji_found in emojis_found:
emoji_ctr[emoji_found] += 1
for item in emoji_ctr.most_common(10):
print(item[0] + " - " + str(item[1]))
```
## Q. What does my WhatsApp activity tell about my sleep cycle?
```
df['hour'] = df['date_time'].apply(lambda x: x.hour)
df[df['user']==me].groupby(['hour']).size().sort_index().plot(x="hour", kind='bar')
```
## Let's take Week Days and Weekends into consideration
## Q. How many words do I type on average on weekday vs weekend?
```
df['weekday'] = df['date_time'].apply(lambda x: x.day_name()) # can use day_name or weekday from datetime
df['is_weekend'] = df.weekday.isin(['Sunday', 'Saturday'])
msgs_per_user = df['user'].value_counts(sort=True)
msgs_per_user
```
Who are the top 5 message senders?
```
top5_users = msgs_per_user.index.tolist()[:5]
top5_users
df_top5 = df.copy()
df_top5 = df_top5[df_top5.user.isin(top5_users)]
df_top5.head()
plt.figure(figsize=(30,10))
sns.countplot(x="user", hue="weekday", data=df)
df_top5['is_weekend'] = df_top5.weekday.isin(['Sunday', 'Saturday'])
plt.figure(figsize=(20,10))
sns.countplot(x="user", hue="is_weekend", data=df_top5)
def word_count(val):
return len(val.split())
df['no_of_words'] = df['msg'].apply(word_count)
df_top5['no_of_words'] = df_top5['msg'].apply(word_count)
```
Total words used in Weekdays
```
total_words_weekday = df[df['is_weekend']==False]['no_of_words'].sum()
total_words_weekday
```
Total words used in Weekends
```
total_words_weekend = df[df['is_weekend']]['no_of_words'].sum()
total_words_weekend
# average words on a weekday
total_words_weekday/5
# average words on a weekend
total_words_weekend/2
```
Number of words used by users in descending order
```
df.groupby('user')['no_of_words'].sum().sort_values(ascending=False)
(df_top5.groupby('user')['no_of_words'].sum()/df_top5.groupby('user').size()).sort_values(ascending=False)
wordPerMsg_weekday_vs_weekend = (df_top5.groupby(['user', 'is_weekend'])['no_of_words'].sum()/df_top5.groupby(['user', 'is_weekend']).size())
wordPerMsg_weekday_vs_weekend
wordPerMsg_weekday_vs_weekend.plot(kind='barh')
```
## Q. At what time of day do I use WhatsApp most?
```
x = df.groupby(['hour', 'weekday'])['msg'].size().reset_index()
x2 = x.pivot("hour", 'weekday', 'msg')
x2.head()
days = ["Monday", 'Tuesday', "Wednesday", "Thursday", "Friday", "Saturday", "Sunday"]
sns.heatmap(x2[days].fillna(0), robust=True)
```
## Let's know whom did I respond the most in the group?
```
my_msgs_index = np.array(df[df['user']==me].index)
print(my_msgs_index, my_msgs_index.shape)
prev_msgs_index = my_msgs_index - 1
print(prev_msgs_index, prev_msgs_index.shape)
df_replies = df.iloc[prev_msgs_index].copy()
df_replies.shape
df_replies.groupby(["user"])["msg"].size().sort_values().plot(kind='barh')
comment_words = ' '
# stopwords = STOPWORDS.update([])
for val in df.msg.values:
val = str(val)
tokens = val.split()
for i in range(len(tokens)):
tokens[i] = tokens[i].lower()
for words in tokens:
comment_words = comment_words + words + ' '
wordcloud = WordCloud(width = 800, height = 800,
background_color ='black',
# stopwords = stopwords,
min_font_size = 10).generate(comment_words)
wordcloud.to_image()
```
| github_jupyter |
# 3.4. Set cardinality with HyperLogLog
## Set cardinality
Again, let's revisit our betalactamase genes:
```
import nbimporter
import Background as utils
# These AMR genes are downloaded from the ResFinder database (accessed 05/2017):
## blaIMP-1_1_DQ522237
geneA = "ATGAGCAAGTTATCTGTATTCTTTATATTTTTGTTTTGCAGCATTGCTACCGCAGCAGAGTCTTTGCCAGATTTAAAAATTGAAAAGCTTGATGAAGGCGTTTATGTTCATACTTCGTTTAAAGAAGTTAACGGGTGGGGCGTTGTTCCTAAACATGGTTTGGTGGTTCTTGTAAATGCTGAGGCTTACCTAATTGACACTCCATTTACGGCTAAAGATACTGAAAAGTTAGTCACTTGGTTTGTGGAGCGTGGCTATAAAATAAAAGGCAGCATTTCCTCTCATTTTCATAGCGACAGCACGGGCGGAATAGAGTGGCTTAATTCTCGATCTATCCCCACGTATGCATCTGAATTAACAAATGAACTGCTTAAAAAAGACGGTAAGGTTCAAGCCACAAATTCATTTAGCGGAGTTAACTATTGGCTAGTTAAAAATAAAATTGAAGTTTTTTATCCAGGCCCGGGACACACTCCAGATAACGTAGTGGTTTGGTTGCCTGAAAGGAAAATATTATTCGGTGGTTGTTTTATTAAACCGTACGGTTTAGGCAATTTGGGTGACGCAAATATAGAAGCTTGGCCAAAGTCCGCCAAATTATTAAAGTCCAAATATGGTAAGGCAAAACTGGTTGTTCCAAGTCACAGTGAAGTTGGAGACGCATCACTCTTGAAACTTACATTAGAGCAGGCGGTTAAAGGGTTAAACGAAAGTAAAAAACCATCAAAACCAAGCAACTAA"
## blaIMP-2_1_AJ243491
geneB = "ATGAAGAAATTATTTGTTTTATGTGTATGCTTCCTTTGTAGCATTACTGCCGCGGGAGCGCGTTTGCCTGATTTAAAAATCGAGAAGCTTGAAGAAGGTGTTTATGTTCATACATCGTTCGAAGAAGTTAACGGTTGGGGTGTTGTTTCTAAACACGGTTTGGTGGTTCTTGTAAACACTGACGCCTATCTGATTGACACTCCATTTACTGCTACAGATACTGAAAAGTTAGTCAATTGGTTTGTGGAGCGCGGCTATAAAATCAAAGGCACTATTTCCTCACATTTCCATAGCGACAGCACAGGGGGAATAGAGTGGCTTAATTCTCAATCTATTCCCACGTATGCATCTGAATTAACAAATGAACTTCTTAAAAAAGACGGTAAGGTGCAAGCTAAAAACTCATTTAGCGGAGTTAGTTATTGGCTAGTTAAAAATAAAATTGAAGTTTTTTATCCCGGCCCGGGGCACACTCAAGATAACGTAGTGGTTTGGTTACCTGAAAAGAAAATTTTATTCGGTGGTTGTTTTGTTAAACCGGACGGTCTTGGTAATTTGGGTGACGCAAATTTAGAAGCTTGGCCAAAGTCCGCCAAAATATTAATGTCTAAATATGTTAAAGCAAAACTGGTTGTTTCAAGTCATAGTGAAATTGGGGACGCATCACTCTTGAAACGTACATGGGAACAGGCTGTTAAAGGGCTAAATGAAAGTAAAAAACCATCACAGCCAAGTAACTAA"
# get the canonical k-mers for each gene
kmersA = utils.getKmers(geneA, 7)
kmersB = utils.getKmers(geneB, 7)
# how many k-mers do we have
print("blaIMP-1 is {} bases long and has {} k-mers (k=7)" .format(len(geneA), len(kmersA)))
print("blaIMP-2 is {} bases long and has {} k-mers (k=7)" .format(len(geneA), len(kmersB)))
```
* we counted the number of k-mers by using the length function in python. However, this will count duplicate k-mers. * we will now use HyperLogLog sketch to approximate the cardinality for each set of k-mers:
```
from datasketch import HyperLogLog
sketch = HyperLogLog(p=4)
for kmer in kmersA:
sketch.update(kmer.encode('utf8'))
print("Estimated cardinality of 7-mers in blaIMP-1 is {0:.0f}" .format(sketch.count()))
```
* let's work out the actual cardinality and compare this to our estimate:
```
setA = set(kmersA)
print("Actual cardinality of 7-mers in blaIMP-1 is {}" .format(len(setA)))
```
* we are quite far off with this estimate
* let's increase the accuracy of the sketch by adding more registers
* change the number of registers from 4 to 14:
```
sketch = HyperLogLog(p=14)
for kmer in kmersA:
sketch.update(kmer.encode('utf8'))
print("Estimated cardinality of 7-mers in blaIMP-1 is {0:.0f}" .format(sketch.count()))
```
| github_jupyter |
# Connect-4 Classification
_____
Dataset : https://archive.ics.uci.edu/ml/datasets/Connect-4
This database contains all legal 8-ply positions in the game of connect-4 in which neither player has won yet, and in which the next move is not forced.
x is the first player; o the second.
The outcome class is the game theoretical value for the first player.
Attribute Information: (x=player x has taken, o=player o has taken, b=blank)
The board is numbered like:
<pre>
6 . . . . . . .
5 . . . . . . .
4 . . . . . . .
3 . . . . . . .
2 . . . . . . .
1 . . . . . . .
a b c d e f g
1. a1: {x,o,b}
2. a2: {x,o,b}
3. a3: {x,o,b}
4. a4: {x,o,b}
5. a5: {x,o,b}
6. a6: {x,o,b}
7. b1: {x,o,b}
8. b2: {x,o,b}
9. b3: {x,o,b}
10. b4: {x,o,b}
11. b5: {x,o,b}
12. b6: {x,o,b}
13. c1: {x,o,b}
14. c2: {x,o,b}
15. c3: {x,o,b}
16. c4: {x,o,b}
17. c5: {x,o,b}
18. c6: {x,o,b}
19. d1: {x,o,b}
20. d2: {x,o,b}
21. d3: {x,o,b}
22. d4: {x,o,b}
23. d5: {x,o,b}
24. d6: {x,o,b}
25. e1: {x,o,b}
26. e2: {x,o,b}
27. e3: {x,o,b}
28. e4: {x,o,b}
29. e5: {x,o,b}
30. e6: {x,o,b}
31. f1: {x,o,b}
32. f2: {x,o,b}
33. f3: {x,o,b}
34. f4: {x,o,b}
35. f5: {x,o,b}
36. f6: {x,o,b}
37. g1: {x,o,b}
38. g2: {x,o,b}
39. g3: {x,o,b}
40. g4: {x,o,b}
41. g5: {x,o,b}
42. g6: {x,o,b}
43. Class: {win,loss,draw}
</pre>
```
import os
from sklearn.tree import DecisionTreeClassifier, export_graphviz
import pandas as pd
import numpy as np
from sklearn.cross_validation import train_test_split
from sklearn import cross_validation, metrics
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import BernoulliNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from time import time
from sklearn import preprocessing
from sklearn.pipeline import Pipeline
from sklearn.metrics import roc_auc_score , classification_report
from sklearn.grid_search import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.metrics import precision_score, recall_score, accuracy_score, classification_report
index=[]
board = ['a','b','c','d','e','f','g']
for i in board:
for j in range(6):
index.append(i + str(j+1))
column_names = index +['Class']
# read .csv from provided dataset
csv_filename="connect-4.data"
# df=pd.read_csv(csv_filename,index_col=0)
df=pd.read_csv(csv_filename,
names= column_names)
df.head()
df['Class'].unique()
#Convert animal labels to numbers
le = preprocessing.LabelEncoder()
for col in df.columns:
df[col] = le.fit_transform(df[col])
df.head()
for col in df.columns:
df[col] = pd.get_dummies(df[col])
df.head()
X = df[index]
y = df['Class']
X.head()
# split dataset to 60% training and 40% testing
X_train, X_test, y_train, y_test = cross_validation.train_test_split(X, y, test_size=0.4, random_state=0)
print (X_train.shape, y_train.shape)
```
### Feature importances with forests of trees
This examples shows the use of forests of trees to evaluate the importance of features on an artificial classification task. The red bars are the feature importances of the forest, along with their inter-trees variability.
```
features = index
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from sklearn.ensemble import ExtraTreesClassifier
# Build a classification task using 3 informative features
# Build a forest and compute the feature importances
forest = ExtraTreesClassifier(n_estimators=250,
random_state=0)
forest.fit(X, y)
importances = forest.feature_importances_
std = np.std([tree.feature_importances_ for tree in forest.estimators_],
axis=0)
indices = np.argsort(importances)[::-1]
# Print the feature ranking
print("Feature ranking:")
for f in range(X.shape[1]):
print("%d. feature %d - %s (%f) " % (f + 1, indices[f], features[indices[f]], importances[indices[f]]))
# Plot the feature importances of the forest
plt.figure(num=None, figsize=(14, 10), dpi=80, facecolor='w', edgecolor='k')
plt.title("Feature importances")
plt.bar(range(X.shape[1]), importances[indices],
color="r", yerr=std[indices], align="center")
plt.xticks(range(X.shape[1]), indices)
plt.xlim([-1, X.shape[1]])
plt.show()
importances[indices[:5]]
for f in range(5):
print("%d. feature %d - %s (%f)" % (f + 1, indices[f], features[indices[f]] ,importances[indices[f]]))
best_features = []
for i in indices[:5]:
best_features.append(features[i])
# Plot the top 5 feature importances of the forest
plt.figure(num=None, figsize=(8, 6), dpi=80, facecolor='w', edgecolor='k')
plt.title("Feature importances")
plt.bar(range(5), importances[indices][:5],
color="r", yerr=std[indices][:5], align="center")
plt.xticks(range(5), best_features)
plt.xlim([-1, 5])
plt.show()
```
# Decision Tree accuracy and time elapsed caculation
```
t0=time()
print ("DecisionTree")
dt = DecisionTreeClassifier(min_samples_split=20,random_state=99)
# dt = DecisionTreeClassifier(min_samples_split=20,max_depth=5,random_state=99)
clf_dt=dt.fit(X_train,y_train)
print ("Acurracy: ", clf_dt.score(X_test,y_test))
t1=time()
print ("time elapsed: ", t1-t0)
```
## cross validation for DT
```
tt0=time()
print ("cross result========")
scores = cross_validation.cross_val_score(dt, X, y, cv=5)
print (scores)
print (scores.mean())
tt1=time()
print ("time elapsed: ", tt1-tt0)
```
### Tuning our hyperparameters using GridSearch
```
from sklearn.metrics import classification_report
pipeline = Pipeline([
('clf', DecisionTreeClassifier(criterion='entropy'))
])
parameters = {
'clf__max_depth': (5, 25 , 50),
'clf__min_samples_split': (2, 5, 10),
'clf__min_samples_leaf': (1, 2, 3)
}
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1, scoring='f1')
grid_search.fit(X_train, y_train)
print ('Best score: %0.3f' % grid_search.best_score_)
print ('Best parameters set:')
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print ('\t%s: %r' % (param_name, best_parameters[param_name]))
predictions = grid_search.predict(X_test)
print (classification_report(y_test, predictions))
```
# Exporting Decision Tree to an Image
```
t0=time()
print ("DecisionTree")
dt1 = DecisionTreeClassifier(min_samples_split=2,max_depth=25,min_samples_leaf=1, random_state=99)
clf_dt1=dt1.fit(X_train,y_train)
print ("Acurracy: ", clf_dt1.score(X_test,y_test))
t1=time()
print ("time elapsed: ", t1-t0)
export_graphviz(clf_dt1,
out_file='tree.dot',
feature_names=features)
```
After we have installed GraphViz on our computer, we can convert the tree.dot file into a PNG file by executing the following command from the command line in the location where we saved the tree.dot file:
> dot -Tpng tree.dot -o tree.png
# Random Forest accuracy and time elapsed caculation
```
t2=time()
print ("RandomForest")
rf = RandomForestClassifier(n_estimators=100,n_jobs=-1)
clf_rf = rf.fit(X_train,y_train)
print ("Acurracy: ", clf_rf.score(X_test,y_test))
t3=time()
print ("time elapsed: ", t3-t2)
```
## cross validation for RF
```
tt2=time()
print ("cross result========")
scores = cross_validation.cross_val_score(rf, X, y, cv=5)
print (scores)
print (scores.mean())
tt3=time()
print ("time elapsed: ", tt3-tt2)
```
### Tuning Models using GridSearch
```
pipeline2 = Pipeline([
('clf', RandomForestClassifier(criterion='entropy'))
])
parameters = {
'clf__n_estimators': (5, 25, 50, 100),
'clf__max_depth': (5, 25 , 50),
'clf__min_samples_split': (2, 5, 10),
'clf__min_samples_leaf': (1, 2, 3)
}
grid_search = GridSearchCV(pipeline2, parameters, n_jobs=-1, verbose=1, scoring='accuracy', cv=3)
grid_search.fit(X_train, y_train)
print ('Best score: %0.3f' % grid_search.best_score_)
print ('Best parameters set:')
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print ('\t%s: %r' % (param_name, best_parameters[param_name]))
predictions = grid_search.predict(X_test)
print ('Accuracy:', accuracy_score(y_test, predictions))
print (classification_report(y_test, predictions))
```
# Naive Bayes accuracy and time elapsed caculation
```
t4=time()
print ("NaiveBayes")
nb = BernoulliNB()
clf_nb=nb.fit(X_train,y_train)
print ("Acurracy: ", clf_nb.score(X_test,y_test))
t5=time()
print ("time elapsed: ", t5-t4)
```
## cross-validation for NB
```
tt4=time()
print ("cross result========")
scores = cross_validation.cross_val_score(nb, X,y, cv=3)
print (scores)
print (scores.mean())
tt5=time()
print ("time elapsed: ", tt5-tt4)
```
# KNN accuracy and time elapsed caculation
```
t6=time()
print( "KNN")
# knn = KNeighborsClassifier(n_neighbors=3)
knn = KNeighborsClassifier(n_neighbors=3)
clf_knn=knn.fit(X_train, y_train)
print ("Acurracy: ", clf_knn.score(X_test,y_test) )
t7=time()
print ("time elapsed: ", t7-t6)
```
## cross validation for KNN
```
tt6=time()
print ("cross result========")
scores = cross_validation.cross_val_score(knn, X,y, cv=5)
print (scores)
print (scores.mean())
tt7=time()
print ("time elapsed: ", tt7-tt6)
```
### Fine tuning the model using GridSearch
```
from sklearn.svm import SVC
from sklearn.cross_validation import cross_val_score
from sklearn.pipeline import Pipeline
from sklearn import grid_search
knn = KNeighborsClassifier()
parameters = {'n_neighbors':[1,10]}
grid = grid_search.GridSearchCV(knn, parameters, n_jobs=-1, verbose=1, scoring='accuracy')
grid.fit(X_train, y_train)
print ('Best score: %0.3f' % grid.best_score_)
print ('Best parameters set:')
best_parameters = grid.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print ('\t%s: %r' % (param_name, best_parameters[param_name]))
predictions = grid.predict(X_test)
print (classification_report(y_test, predictions))
```
# SVM accuracy and time elapsed caculation
```
t7=time()
print ("SVM")
svc = SVC(kernel='rbf')
clf_svc=svc.fit(X_train, y_train)
print ("Acurracy: ", clf_svc.score(X_test,y_test) )
t8=time()
print ("time elapsed: ", t8-t7)
```
## cross validation for SVM
```
tt7=time()
print ("cross result========")
scores = cross_validation.cross_val_score(svc,X,y, cv=5)
print (scores)
print (scores.mean())
tt8=time()
print ("time elapsed: ", tt7-tt6)
```
____
| github_jupyter |
# Web Scraping using Selenium and Beautiful Soup
<table align="left"><td>
<a target="_blank" href="https://colab.research.google.com/github/TannerGilbert/Tutorials/blob/master/Web%20Scraping%20using%20Selenium%20and%20BeautifulSoup/Web%20Scraping%20using%20Selenium%20and%20Beautiful%20Soup.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab
</a>
</td><td>
<a target="_blank" href="https://github.com/TannerGilbert/Tutorials/blob/master/Web%20Scraping%20using%20Selenium%20and%20BeautifulSoup/Web%20Scraping%20using%20Selenium%20and%20Beautiful%20Soup.ipynb">
<img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td></table>
Selenium is a browser automation tool that can not only be used for testing, but also for many other purposes. It's especially useful because using it we can also scrape data that are client side rendered.
Installation:
```
!pip install selenium
```
or
```
!conda install selenium
```
To use Selenuim a WebDriver for your favorite web browser must also be installed. The Firefox WebDriver(GeckoDriver) can be installed by going to [this page](https://github.com/mozilla/geckodriver/releases/) and downloading the appropriate file for your operating system. After the download has finished the file has to be extracted.
Now the file can either be [added to path](https://www.architectryan.com/2018/03/17/add-to-the-path-on-windows-10/) or copied into the working directory. I chose to copy it to my working directory because I’m not using it that often.
Importing:
```
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import WebDriverWait
from bs4 import BeautifulSoup
import pandas as pd
import re
import os
# website url
base_url = "https://programmingwithgilbert.firebaseapp.com/"
videos_url = "https://programmingwithgilbert.firebaseapp.com/videos/keras-tutorials"
```
Trying to load the data using urllib. This won't get any data because it can't load data which is loaded after the document.onload function
```
import urllib.request
page = urllib.request.urlopen(videos_url)
soup = BeautifulSoup(page, 'html.parser')
soup
```
Now we can create a firefox session and navigate to the base url of the video section
```
# Firefox session
driver = webdriver.Firefox()
driver.get(videos_url)
driver.implicitly_wait(100)
```
To navigate to the specifc pages we nned to get the buttons which a a text of "Watch" and then navigate to each side, scrape the data, save it and go back to the main page
```
num_links = len(driver.find_elements_by_link_text('Watch'))
code_blocks = []
for i in range(num_links):
# navigate to link
button = driver.find_elements_by_class_name("btn-primary")[i]
button.click()
# get soup
element = WebDriverWait(driver, 10).until(lambda x: x.find_element_by_id('iframe_container'))
tutorial_soup = BeautifulSoup(driver.page_source, 'html.parser')
tutorial_code_soup = tutorial_soup.find_all('div', attrs={'class': 'code-toolbar'})
tutorial_code = [i.getText() for i in tutorial_code_soup]
code_blocks.append(tutorial_code)
# go back to initial page
driver.execute_script("window.history.go(-1)")
code_blocks
code_blocks[1]
```
After scraping all the needed data we can close the browser session and save the results into .txt files
```
driver.quit()
for i, tutorial_code in enumerate(code_blocks):
with open('code_blocks{}.txt'.format(i), 'w') as f:
for code_block in tutorial_code:
f.write(code_block+"\n")
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import os, sys
from tqdm import tqdm
import numpy as np
import pandas as pd
DELQSAR_ROOT = os.getcwd() + '/../../'
sys.path += [DELQSAR_ROOT + '/../']
if not os.path.isdir('DD1S_CAIX_cycle2_distrib_shift'):
os.mkdir('DD1S_CAIX_cycle2_distrib_shift')
def pathify(fname):
return os.path.join('DD1S_CAIX_cycle2_distrib_shift', fname)
LOG_FILE = os.path.join(DELQSAR_ROOT, 'experiments', 'visualizations',
'DD1S_CAIX_cycle2_distrib_shift', 'DD1S_CAIX_cycle2_distrib_shift.log')
from del_qsar import splitters
from del_qsar.enrichments import R_ranges
import matplotlib
import matplotlib.pyplot as plt
plt.style.use('seaborn-paper')
matplotlib.rc('font', family='sans-serif')
matplotlib.rc('font', serif='Arial')
matplotlib.rc('text', usetex='false')
df_data = pd.read_csv(os.path.join(DELQSAR_ROOT, 'experiments', 'datasets', 'DD1S_CAIX_QSAR.csv'))
exp_counts = np.array(df_data[['exp_tot']], dtype='int')
bead_counts = np.array(df_data[['beads_tot']], dtype='int')
exp_tot = np.sum(exp_counts, axis=0) # column sums
bead_tot = np.sum(bead_counts, axis=0)
splitter = splitters.OneCycleSplitter(['cycle2'], LOG_FILE)
test_slices = [splitter(_, df_data, seed=i)[2] for i in tqdm(range(5))]
[len(ts) for ts in test_slices]
def make_plot_calc_enrichments(eval_slices, seeds, zoomIn):
fig = plt.figure(figsize=(3.5, 2), dpi=300)
for seed in seeds:
R, R_lb, R_ub = R_ranges(bead_counts[eval_slices[seed], 0], bead_tot[0],
exp_counts[eval_slices[seed], 0], exp_tot[0])
bins = np.arange(0, max(R_lb)+0.001, 0.06)
_, bins, patches = plt.hist(
np.clip(R_lb, 0, bins[-1]),
bins=bins,
density=True,
zorder=2,
alpha=0.4,
label=f'seed {seed}',
)
plt.legend(fontsize=7)
fig.canvas.draw()
ax = plt.gca()
ax.tick_params(labelsize=9)
ax.set_xlim([0, 2])
if zoomIn:
ax.set_ylim([0, 0.4])
ax.grid(zorder=1)
ax.set_xlabel('calculated enrichment (lower bound)', fontsize=9)
ax.set_ylabel('probability density', fontsize=9)
plt.tight_layout()
if zoomIn:
plt.savefig(pathify(f'DD1S_CAIX_cycle2_calculated_enrichments_LB_seeds_{str(seeds)}_y-axis_zoomed_in.png'))
else:
plt.savefig(pathify(f'DD1S_CAIX_cycle2_calculated_enrichments_LB_seeds_{str(seeds)}.png'))
plt.show()
make_plot_calc_enrichments(test_slices, [0,1,2,3,4], zoomIn=False)
make_plot_calc_enrichments(test_slices, [0,1,2,3,4], zoomIn=True)
make_plot_calc_enrichments(test_slices, [0,4], False)
make_plot_calc_enrichments(test_slices, [0,4], True)
make_plot_calc_enrichments(test_slices, [1,4], False)
make_plot_calc_enrichments(test_slices, [1,4], True)
make_plot_calc_enrichments(test_slices, [2,4], False)
make_plot_calc_enrichments(test_slices, [2,4], True)
make_plot_calc_enrichments(test_slices, [3,4], False)
make_plot_calc_enrichments(test_slices, [3,4], True)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import gensim
import re
import nltk
import json
import sys
import datetime
import operator
import matplotlib.pyplot as plt
import math
import csv
import timeit
from collections import Counter
from nltk.corpus import stopwords
from nltk import word_tokenize, ngrams
from gensim import corpora, models, similarities
from gensim.models.doc2vec import Doc2Vec
from gensim.models import doc2vec
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
from pylab import plot, show, subplot, specgram, imshow, savefig
from tqdm import tqdm
%matplotlib inline
def cosine(v1, v2):
v1 = np.array(v1)
v2 = np.array(v2)
return np.dot(v1, v2) / (np.sqrt(np.sum(v1**2)) * np.sqrt(np.sum(v2**2)))
def concatenate(data):
X_set1 = data['question1']
X_set2 = data['question2']
X = X_set1.append(X_set2, ignore_index=True)
return X
class LabeledLineSentence(object):
def __init__(self, doc_list, labels_list):
self.labels_list = labels_list
self.doc_list = doc_list
def __iter__(self):
for idx, doc in enumerate(self.doc_list):
yield doc2vec.TaggedDocument(words=word_tokenize(doc),
tags=[self.labels_list[idx]])
def get_dists_doc2vec(data):
docvec1s = []
docvec2s = []
for i in tqdm(range(data.shape[0])):
doc1 = word_tokenize(data.iloc[i, -2])
doc2 = word_tokenize(data.iloc[i, -1])
docvec1 = model.infer_vector(doc1, alpha=start_alpha, steps=infer_epoch)
docvec2 = model.infer_vector(doc2, alpha=start_alpha, steps=infer_epoch)
docvec1s.append(docvec1)
docvec2s.append(docvec2)
return docvec1s, docvec2s
src_train = 'df_train_spacylemmat_fullclean.csv'
src_test = 'df_test_spacylemmat_fullclean.csv'
df_train = pd.read_csv(src_train)
df_test = pd.read_csv(src_test)
df_train = df_train[['id', 'question1', 'question2']]
df_test = df_test[['test_id', 'question1', 'question2']]
df_train.fillna('NULL', inplace = True)
df_test.fillna('NULL', inplace = True)
df_test.rename(columns = {'test_id': 'id'}, inplace = True)
data = pd.concat([df_train, df_test], ignore_index = True)
X_train = data[['id', 'question1', 'question2']]
X = concatenate(X_train)
labels = []
for label in X_train['id'].tolist():
labels.append('SENT_%s_1' % label)
for label in X_train['id'].tolist():
labels.append('SENT_%s_2' % label)
docs = LabeledLineSentence(X.tolist(), labels)
it = docs.__iter__()
model = Doc2Vec(size=100, window=10, min_count=2, sample=1e-5, workers=8, iter = 20)
model.build_vocab(docs)
print('Model built.')
model.train(docs, total_examples=model.corpus_count, epochs=model.iter)
print('Model trained.')
start_alpha = 0.01
infer_epoch = 10
results = get_dists_doc2vec(data)
docvec1s, docvec2s = results[0], results[1]
docvec1s = np.array(docvec1s)
docvec1s_tr = docvec1s[:df_train.shape[0]]
docvec1s_te = docvec1s[df_train.shape[0]:]
docvec2s = np.array(docvec2s)
docvec2s_tr = docvec2s[:df_train.shape[0]]
docvec2s_te = docvec2s[df_train.shape[0]:]
np.save('train_q1_doc2vec_vectors_trainquora', docvec1s_tr)
np.save('test_q1_doc2vec_vectors_trainquora', docvec1s_te)
np.save('train_q2_doc2vec_vectors_trainquora', docvec2s_tr)
np.save('test_q2_doc2vec_vectors_trainquora', docvec2s_te)
```
| github_jupyter |
# Tutorial Part 5: Putting Multitask Learning to Work
This notebook walks through the creation of multitask models on MUV [1]. The goal is to demonstrate that multitask methods outperform singletask methods on MUV.
## Colab
This tutorial and the rest in this sequence are designed to be done in Google colab. If you'd like to open this notebook in colab, you can use the following link.
[](https://colab.research.google.com/github/deepchem/deepchem/blob/master/examples/tutorials/05_Putting_Multitask_Learning_to_Work.ipynb)
## Setup
To run DeepChem within Colab, you'll need to run the following cell of installation commands. This will take about 5 minutes to run to completion and install your environment.
```
!curl -Lo conda_installer.py https://raw.githubusercontent.com/deepchem/deepchem/master/scripts/colab_install.py
import conda_installer
conda_installer.install()
!/root/miniconda/bin/conda info -e
!pip install --pre deepchem
import deepchem
deepchem.__version__
```
The MUV dataset is a challenging benchmark in molecular design that consists of 17 different "targets" where there are only a few "active" compounds per target. The goal of working with this dataset is to make a machine learnign model which achieves high accuracy on held-out compounds at predicting activity. To get started, let's download the MUV dataset for us to play with.
```
import os
import deepchem as dc
current_dir = os.path.dirname(os.path.realpath("__file__"))
dataset_file = "medium_muv.csv.gz"
full_dataset_file = "muv.csv.gz"
# We use a small version of MUV to make online rendering of notebooks easy. Replace with full_dataset_file
# In order to run the full version of this notebook
dc.utils.download_url("https://s3-us-west-1.amazonaws.com/deepchem.io/datasets/%s" % dataset_file,
current_dir)
dataset = dc.utils.save.load_from_disk(dataset_file)
print("Columns of dataset: %s" % str(dataset.columns.values))
print("Number of examples in dataset: %s" % str(dataset.shape[0]))
```
Now, let's visualize some compounds from our dataset
```
from rdkit import Chem
from rdkit.Chem import Draw
from itertools import islice
from IPython.display import Image, display, HTML
def display_images(filenames):
"""Helper to pretty-print images."""
for filename in filenames:
display(Image(filename))
def mols_to_pngs(mols, basename="test"):
"""Helper to write RDKit mols to png files."""
filenames = []
for i, mol in enumerate(mols):
filename = "MUV_%s%d.png" % (basename, i)
Draw.MolToFile(mol, filename)
filenames.append(filename)
return filenames
num_to_display = 12
molecules = []
for _, data in islice(dataset.iterrows(), num_to_display):
molecules.append(Chem.MolFromSmiles(data["smiles"]))
display_images(mols_to_pngs(molecules))
```
There are 17 datasets total in MUV as we mentioned previously. We're going to train a multitask model that attempts to build a joint model to predict activity across all 17 datasets simultaneously. There's some evidence [2] that multitask training creates more robust models.
As fair warning, from my experience, this effect can be quite fragile. Nonetheless, it's a tool worth trying given how easy DeepChem makes it to build these models. To get started towards building our actual model, let's first featurize our data.
```
MUV_tasks = ['MUV-692', 'MUV-689', 'MUV-846', 'MUV-859', 'MUV-644',
'MUV-548', 'MUV-852', 'MUV-600', 'MUV-810', 'MUV-712',
'MUV-737', 'MUV-858', 'MUV-713', 'MUV-733', 'MUV-652',
'MUV-466', 'MUV-832']
featurizer = dc.feat.CircularFingerprint(size=1024)
loader = dc.data.CSVLoader(
tasks=MUV_tasks, smiles_field="smiles",
featurizer=featurizer)
dataset = loader.featurize(dataset_file)
```
We'll now want to split our dataset into training, validation, and test sets. We're going to do a simple random split using `dc.splits.RandomSplitter`. It's worth noting that this will provide overestimates of real generalizability! For better real world estimates of prospective performance, you'll want to use a harder splitter.
```
# splitter = dc.splits.RandomSplitter(dataset_file)
# train_dataset, valid_dataset, test_dataset = splitter.train_valid_test_split(
# dataset)
# #NOTE THE RENAMING:
# valid_dataset, test_dataset = test_dataset, valid_dataset
```
Let's now get started building some models! We'll do some simple hyperparameter searching to build a robust model.
```
# import numpy as np
# import numpy.random
# params_dict = {"activation": ["relu"],
# "momentum": [.9],
# "batch_size": [50],
# "init": ["glorot_uniform"],
# "data_shape": [train_dataset.get_data_shape()],
# "learning_rate": [1e-3],
# "decay": [1e-6],
# "nb_epoch": [1],
# "nesterov": [False],
# "dropouts": [(.5,)],
# "nb_layers": [1],
# "batchnorm": [False],
# "layer_sizes": [(1000,)],
# "weight_init_stddevs": [(.1,)],
# "bias_init_consts": [(1.,)],
# "penalty": [0.],
# }
# n_features = train_dataset.get_data_shape()[0]
# def model_builder(model_params, model_dir):
# model = dc.models.MultitaskClassifier(
# len(MUV_tasks), n_features, **model_params)
# return model
# metric = dc.metrics.Metric(dc.metrics.roc_auc_score, np.mean)
# optimizer = dc.hyper.HyperparamOpt(model_builder)
# best_dnn, best_hyperparams, all_results = optimizer.hyperparam_search(
# params_dict, train_dataset, valid_dataset, [], metric)
```
# Congratulations! Time to join the Community!
Congratulations on completing this tutorial notebook! If you enjoyed working through the tutorial, and want to continue working with DeepChem, we encourage you to finish the rest of the tutorials in this series. You can also help the DeepChem community in the following ways:
## Star DeepChem on [GitHub](https://github.com/deepchem/deepchem)
This helps build awareness of the DeepChem project and the tools for open source drug discovery that we're trying to build.
## Join the DeepChem Gitter
The DeepChem [Gitter](https://gitter.im/deepchem/Lobby) hosts a number of scientists, developers, and enthusiasts interested in deep learning for the life sciences. Join the conversation!
# Bibliography
[1] https://pubs.acs.org/doi/10.1021/ci8002649
[2] https://pubs.acs.org/doi/abs/10.1021/acs.jcim.7b00146
| github_jupyter |
# CME 193 - Pandas Exercise Supplement
In this extended exercise, you'll load and play with CO2 data collected at the Mauna Loa observatory over the last 60 years.
* NOAA Website: https://www.esrl.noaa.gov/gmd/ccgg/trends/full.html
* NOAA data: https://www.esrl.noaa.gov/gmd/ccgg/trends/data.html
The monthly data can be found at this [link](ftp://aftp.cmdl.noaa.gov/products/trends/co2/co2_mm_mlo.txt)
```
import numpy as np
import scipy
import pandas as pd
import matplotlib.pyplot as plt
```
Reads the data from the ftp server directly.
```
df = pd.read_csv('ftp://aftp.cmdl.noaa.gov/products/trends/co2/co2_mm_mlo.txt',
delim_whitespace=True,
comment='#',
names=["year", "month", "decdate", "co2", "co2interp", "trend", "days"],
index_col=False)
pd.set_option('display.max_rows', 10)
df
# copies the original data.
orig = df.copy()
```
## Part 1 - Normalize the Date
1. create a new column for the dataframe called 'day' that is set to be 1 in every entry
```
# your code here
```
2. The dataframe now has columns for 'day', 'month', and 'year'. Use `pd.to_datetime()` to create a new series of dates
`dates = pd.to_datetime(...)`
```
# your code here
```
3. set a new column of the dataframe to hold this series. Call the column `'date'`
```
# your code here
```
4. set the index of the dataframe to be the `'date'` column using the `set_index()` method.
```
# your code here
```
5. Now let's remove the old columns with date information. Use the `drop()` method to remove the 'day', 'month', 'year', and 'decdate' columns. Hint: `df.drop(..., axis=1, inplace=True)`
5a. Go ahead and drop the 'days' column as well, since we're not going to use it.
```
# your code here
```
## Part 2 - deal with missing values
1. First, use the `plot()` method to visualize the contents of your dataframe. What do you see?
```
# your code here
```
if you read the header for the file we used to load the dataframe, you'll see that missing values take the value -99.99.
2. Set values that are `-99.99` to `None` (this indicates a missing value in Pandas).
Hint: use the `applymap()` method, and the lambda function
```python
lambda x: None if x == -99.99 else x
```
If you're familiar with [ternary operators](https://en.wikipedia.org/wiki/%3F:), this is the equivalent of
```
x == -99.99 ? None : x
```
Note that you may need to make a new assignment e.g., `df = df.applymap(...)`
```
# your code here
```
3. Plot your dataframe again. What do you see now?
3a. Try plotting just the 'co2' series. What do you see?
```
# your code here
```
## Part 3 - Create New DataFrames with rows that meet conditions
1. Create new dataframe called `recent` that contains all rows of the previous dataframe since 2007. Plot it.
```
# your code here
```
2. Create a new dataframe called `old` that contains all rows of the dataframe before 1990. Plot it.
```
# your code here
```
##### At this point, by inspection, you might be convinced there is further analysis to be done
```
np.var(old['trend']), np.var(recent['trend'])
```
## Part 4 - Create some groups
Let's go back to the original data that we loaded
```
df = orig
df
```
Suppose that we want to look at co2 averages by year instead of by month.
1. drop rows with missing values
1a. apply the map that sends -99.99 to none
1b. use the `dropna()` method to remove rows with missing values: `df = df.dropna()`
```
# your code here
```
2. Create a group for each year (use key 'year')
```
# your code here
```
3. Aggregate the groups into a new dataframe, `df2`, using `np.mean`
3a. you can drop all the columns except `'co2'` if you'd like
```
# your code here
```
4. make a plot of the `'co2'` series
```
# your code here
```
| github_jupyter |
## Imports
The plotting utilities and various visualization tools for this notebook depend on [degas](https://github.com/williamgilpin/degas) and [tsfresh](https://tsfresh.readthedocs.io/en/latest/)
```
import sys
import matplotlib.pyplot as plt
import json
import pandas as pd
import dysts
from dysts.datasets import *
from dysts.utils import *
try:
from degas import *
except:
pass
import degas as dg
FIGURE_PATH = "../private_writing/fig_resources/"
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
# Featurize many random initial conditions
```
# chunk and featurize several time windows
# EXPENSIVE: can bypass this and just import the features I found in the cell below
all_feature_dfs = list()
for i in np.linspace(0, 8000, 40).astype(int):
dataset = TimeSeriesDataset("../dysts/data/large_univariate__pts_per_period_100__periods_100.json")
dataset.trim_series(i, i + 2000)
feature_df = featurize_timeseries(dataset)
all_feature_dfs.append(feature_df)
print(i)
## Prune features not detected across all replicates
all_feature_lists = [set(item.columns.to_list()) for item in all_feature_dfs]
common_features = np.array(list(all_feature_lists[0].intersection(*all_feature_lists[1:])))
print(f"{len(common_features)} common features found.")
for i in range(len(all_feature_dfs)):
all_feature_dfs[i] = all_feature_dfs[i][common_features]
rep_stds = np.std(np.dstack([np.array(df) for df in all_feature_dfs]), axis=-1)
topk_feature_inds = np.squeeze(np.array([np.argsort(np.median(rep_stds, axis=0))]))[:100]
feat_arr_all = np.dstack([np.array(df)[:, topk_feature_inds] for df in all_feature_dfs])
feat_arr_all = np.transpose(feat_arr_all, (2, 0, 1))
# feat_arr_all.dump("benchmarks/resources/feat_arr_all.pkl")
# Load output of cell above
dataset = TimeSeriesDataset("../dysts/data/large_univariate__pts_per_period_100__periods_100.json")
feat_arr_all = np.load("./resources/feat_arr_all.pkl", allow_pickle=True)
features_mean = np.median(feat_arr_all, axis=0)
feat_arr_all_flat = np.reshape(feat_arr_all, (-1, feat_arr_all.shape[-1]))
feat_arr = features_mean
```
# Create an embedding using the centroids
```
import umap
# model = PCA(n_components=2, random_state=0)
# model = umap.UMAP(random_state=0, densmap=True)
# from umap.parametric_umap import ParametricUMAP
# model = ParametricUMAP(densmap=False)
model = umap.UMAP(random_state=15, n_neighbors=5)
embedding_mean = model.fit_transform(features_mean)
embedding_all = model.transform(feat_arr_all_flat)
plt.plot(embedding_all[:, 0], embedding_all[:, 1], '.', color=[0.6, 0.6, 0.6], markersize=1)
plt.plot(embedding_mean[:, 0], embedding_mean[:, 1], '.k')
## Visualize single clusters
plt.plot(
embedding_mean[:, 0],
embedding_mean[:, 1],
".k"
)
view_ind = 2
plt.plot(
embedding_mean[:, 0][labels_mean == view_ind],
embedding_mean[:, 1][labels_mean == view_ind],
".",
color = dg.pastel_rainbow[view_ind]
)
import hdbscan
from sklearn.cluster import AffinityPropagation
clusterer = AffinityPropagation()
labels_mean = clusterer.fit_predict(embedding_mean)
labels_all = clusterer.predict(embedding_all)
all_name_clusters = list()
print("Number of classes: ", len(np.unique(labels_mean)) - 1)
for label in np.unique(labels_mean):
#if label >= 0:
all_name_clusters.append(dataset.names[labels_mean == label])
if label >= 0:
# color_val = pastel_rainbow_interpolated[label]
color_val = dg.pastel_rainbow[label]
# color_val = dg.pastel_rainbow[label]
else:
color_val = (0.6, 0.6, 0.6)
plt.plot(
embedding_all[labels_all == label, 0],
embedding_all[labels_all == label, 1],
'.',
markersize=2,
color=dg.lighter(color_val, 0.5)
)
plt.plot(
embedding_mean[labels_mean == label, 0],
embedding_mean[labels_mean == label, 1],
'.',
color=color_val,
markersize=8
)
fixed_aspect_ratio(1)
dg.vanish_axes()
# dg.better_savefig(FIGURE_PATH + "clustered_umap2.png")
import seaborn as sns
ax = sns.kdeplot(x=embedding_all[:, 0], y=embedding_all[:, 1], hue=labels_all,
palette=sns.color_palette(dg.pastel_rainbow[:len(np.unique(labels_all))]),
shade=True, bw_adjust=1.3, levels=5)
for contour in ax.collections[:]:
contour.set_alpha(0.3)
ax.grid(False)
ax.legend_.remove()
for label in np.unique(labels_mean):
#if label >= 0:
all_name_clusters.append(dataset.names[labels_mean == label])
if label >= 0:
# color_val = pastel_rainbow_interpolated[label]
color_val = dg.pastel_rainbow[label]
# color_val = dg.pastel_rainbow[label]
else:
color_val = (0.6, 0.6, 0.6)
# plt.plot(
# embedding_all[labels_all == label, 0],
# embedding_all[labels_all == label, 1],
# '.',
# markersize=2,
# color=dg.lighter(color_val, 0.5)
# )
plt.plot(
embedding_mean[labels_mean == label, 0],
embedding_mean[labels_mean == label, 1],
'.',
color=color_val,
markersize=8
)
dg.fixed_aspect_ratio(1)
dg.vanish_axes()
# dg.better_savefig(FIGURE_PATH + "clustered_umap.png")
import dysts.flows
for label, cluster in zip(np.unique(labels_mean), all_name_clusters):
if label < 0:
continue
#print(label, cluster)
color_val = dg.pastel_rainbow[label]
plt.figure()
for item in cluster:
model_dyn = getattr(dysts.flows, item)()
sol = model_dyn.make_trajectory(1000, resample=True, standardize=True, pts_per_period=500)
#plt.plot(sol[:, 0], sol[:, 1], color=color_val)
plt.plot(sol[:, 0], color=color_val, linewidth=3)
fixed_aspect_ratio(1/3)
dg.vanish_axes()
dg.better_savefig(FIGURE_PATH + f"ts{label}.png")
plt.show()
print(label, " ", cluster, "\n--------------\n", flush=True)
```
# Plot example dynamics
```
## Who is in each category?
np.array(all_name_clusters)[[0, 1, 2, 4, 5]]
from dysts.flows import *
# eq = Lorenz84()
# eq = SprottA() # relaxation oscillator dynamics / spiking
# eq = ForcedFitzHughNagumo() # nearly-quasiperiodic
# eq = Lorenz()
# eq = ArnoldBeltramiChildress() # ABC Turbulence
for clr_ind, equation_name in zip([0, 1, 2, 4, 5], ["Chua", "Rossler", "Lorenz", "Lorenz96", "MackeyGlass"]):
# equation_name = all_name_clusters[clr_ind][3]
eq = getattr(dysts.flows, equation_name)()
# eq = HindmarshRose(); clr = dg.pastel_rainbow[0 % 8] # HindmarshRose neuron
# eq = Colpitts(); clr = dg.pastel_rainbow[3 % 8] #Colpitts LC Circuit
# eq = CaTwoPlus(); clr = dg.pastel_rainbow[5 % 8] # Quasiperiodic family
clr = dg.pastel_rainbow[clr_ind]
sol = eq.make_trajectory(8000, resample=True, pts_per_period=200)
if equation_name == "Lorenz96":
sol = eq.make_trajectory(2000, resample=True, pts_per_period=200)
if equation_name == "MackeyGlass":
sol = eq.make_trajectory(4000, resample=True, pts_per_period=200)
plt.figure()
plt.plot(sol[:, 0], 'k')
plt.figure()
plt.plot(sol[:, 0], sol[:, 1], color=clr, linewidth=3)
vanish_axes()
if equation_name == "Lorenz":
dg.better_savefig(FIGURE_PATH + f"sample_{equation_name}.png")
```
## Make histograms of attractor properties
```
import pandas as pd
import seaborn as sns
from dysts.base import get_attractor_list
attributes = ['maximum_lyapunov_estimated', 'kaplan_yorke_dimension', 'pesin_entropy', 'correlation_dimension', "multiscale_entropy"]
all_properties = dict()
for equation_name in get_attractor_list():
eq = getattr(dysts.flows, equation_name)()
attr_vals = [getattr(eq, item, np.nan) for item in attributes]
all_properties[equation_name] = dict(zip(attributes, attr_vals))
all_properties[equation_name]["dynamics_dimension"] = int(len(eq.ic))
all_properties[equation_name]["lyapunov_scaled"] = all_properties[equation_name]["maximum_lyapunov_estimated"] * eq.period
all_properties = pd.DataFrame(all_properties).transpose()
def mirror_df(df, mirror_val=0):
"""
Create a mirrored augmented dataframe. Used
for setting the right boundary conditions on kernel
density plots
"""
return pd.concat([df, mirror_val - df])
sns.set_style()
dg.set_style()
for i, key in enumerate(all_properties.columns):
reduced_df = all_properties[np.logical_not(np.isnan(all_properties[key]))]
reduced_df = reduced_df[reduced_df[key] < np.percentile(reduced_df[key], 95)]
reduced_df = mirror_df(reduced_df)
sns.displot(reduced_df,
x=key,
kde=True,
stat="probability",
color=(0.5, 0.5, 0.5),
linewidth=0,
bins=20,
#clip=(0.0, np.nanpercentile(all_properties[key], 99.5)),
#kde=True, cut=0,
kde_kws={"bw_method" : 0.2}
)
plt.xlim([0, np.max(reduced_df[key])])
dg.fixed_aspect_ratio(1/3)
dg.better_savefig(FIGURE_PATH + f"histogram_{key}.png")
```
| github_jupyter |
```
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
from importlib import reload
import mathutils
import fitsio
import setcover
import archespec
import scipy.optimize as op
from scipy.ndimage import gaussian_filter1d
import cookb_signalsmooth
from matplotlib.colors import LogNorm
plt.rcParams["figure.figsize"]=(10,8)
plt.rcParams['axes.linewidth'] = 2
plt.rcParams['xtick.major.size'] = 15
plt.rcParams['xtick.major.width'] = 2
plt.rcParams['xtick.minor.size'] = 10
plt.rcParams['xtick.minor.width'] = 2
plt.rcParams['xtick.labelsize'] = 25
plt.rcParams['ytick.major.size'] = 15
plt.rcParams['ytick.major.width'] = 2
plt.rcParams['ytick.minor.size'] = 10
plt.rcParams['ytick.minor.width'] = 2
plt.rcParams['ytick.labelsize'] = 25
masterwave, allflux, allivar = archespec.rest_allspec_readin()
objs_ori = archespec.arche_readin()
nobj = objs_ori.size
z = objs_ori['Z']
mass = objs_ori['MASS']
sfr = objs_ori['SFR']
index_wave_all = np.searchsorted(masterwave, [3605., 6997.])
tmpflux = allflux[index_wave_all[0]:index_wave_all[1],:]
tmpivar = allivar[index_wave_all[0]:index_wave_all[1],:]
tmpwave = masterwave[index_wave_all[0]:index_wave_all[1]]
tmploglam = np.log10(tmpwave)
print(tmploglam.size, tmploglam.size//15*15)
itmp = (np.where(np.logical_and(np.logical_and(\
np.logical_and(np.logical_and(mass>8, mass<12), sfr>1e-4),\
z<0.3), z>0.05)))[0]
iuse = np.random.choice(itmp, 1000)
print(iuse.shape)
chi2 = np.zeros((iuse.size, iuse.size))
A = np.zeros((iuse.size, iuse.size))
newwave = np.median(tmpwave.reshape(tmploglam.size//15, 15), axis=1)
newflux = np.sum(tmpflux.reshape(tmploglam.size//15, 15, tmpflux.shape[1]), axis=1)
newivar = 1./np.sum(1./tmpivar.reshape(tmploglam.size//15, 15, tmpflux.shape[1]), axis=1)
tmp_yerr = 1./np.sqrt(newivar[:, iuse].T.reshape(iuse.size, newwave.size))
tmp_y = newflux[:,iuse].T
for i in np.arange(iuse.size):
tmp_x = newflux[:, iuse[i]].T.reshape(1,newwave.size)
tmp_xerr = 1./np.sqrt(newivar[:, iuse[i]].T.reshape(1,newwave.size))
A_tmp, chi2_tmp = mathutils.quick_amplitude(tmp_x, tmp_y, tmp_xerr, tmp_yerr)
A[i,:] = A_tmp
chi2[i,:] = chi2_tmp
a_matrix = chi2/newwave.size<3.5
cost = np.ones(iuse.size)
plt.hist(np.ravel(chi2/newwave.size), 40, range=[0,100])
g = setcover.SetCover(a_matrix, cost)
g.CFT()
iarchetype = np.nonzero(g.s)[0]
print(iarchetype.shape)
tmpmedian = tmpflux[:,iuse[iarchetype]]
# Calculate [O II] EW
wave_oii = 3728.
index_oii = np.searchsorted(tmpwave, wave_oii)
flux_oii = np.sum(tmpmedian[index_oii-5:index_oii+5,:], axis=0)
flux_left = np.sum(tmpmedian[index_oii-15:index_oii-5,:], axis=0)
flux_right = np.sum(tmpmedian[index_oii+5:index_oii+15,:], axis=0)
ew_oii = flux_oii/(flux_left+flux_right)*2.
isort_oii = np.argsort(ew_oii)[::-1]
wave_ha = 6563.
index_ha = np.searchsorted(tmpwave, wave_ha)
flux_ha = np.sum(tmpmedian[index_ha-5:index_ha+5,:], axis=0)
flux_haleft = np.sum(tmpmedian[index_ha-115:index_ha-105,:], axis=0)
flux_haright = np.sum(tmpmedian[index_ha+105:index_ha+115,:], axis=0)
ew_ha = flux_ha/(flux_haleft+flux_haright)*2.
isort_ha = np.argsort(ew_ha)[::-1]
wave_norm = 4600.
index_norm = np.searchsorted(tmpwave, wave_norm)
flux_norm = np.median(tmpmedian[index_norm-5:index_norm+5,:], axis=0)
wave_left = 4200.
index_left = np.searchsorted(tmpwave, wave_left)
flux_left = np.sum(tmpmedian[index_left-15:index_left+15,:], axis=0)
wave_right = 5500.
index_right = np.searchsorted(tmpwave, wave_right)
flux_right = np.sum(tmpmedian[index_right-15:index_right+15,:], axis=0)
color_cont = flux_right/flux_left
isort_cont = np.argsort(color_cont)
#i = -1
i += 1
fig1 = plt.figure(figsize=(20,10))
ax1 = fig1.add_subplot(111)
fig2 = plt.figure(figsize=(20,10))
ax2 = fig2.add_subplot(111)
#x = gaussian_filter1d(tmpwave, 1.5)
x = tmpwave
for i in np.arange(iarchetype.size):
#y = gaussian_filter1d(tmpmedian[:,isort_oii[i]]/flux_norm[isort_oii[i]], 1.5)
#y = cookb_signalsmooth.smooth(tmpmedian[:,isort_oii[i]]/flux_norm[isort_oii[i]], 20)
#iwave_use = np.where(tmpmedian[:,isort_cont[i]]>1.E-5)[0]
iwave_use = np.where(tmpmedian[:,isort_ha[i]]>1.E-5)[0]
y = cookb_signalsmooth.smooth(tmpmedian[iwave_use,isort_ha[i]]/flux_norm[isort_ha[i]], 10)
#ax.plot(x, y)
ax1.plot(x[iwave_use], y)
ax2.loglog(x[iwave_use], y)
ax1.set_xlim(3600, 7200)
ax2.set_xlim(3600, 7200)
ax1.set_xticks((np.arange(4)+1.)*1000.+3000.)
ax2.set_xticks((np.arange(4)+1.)*1000.+3000.)
ax1.set_ylim(1E-1, 5E0)
ax2.set_ylim(1E-1, 2E1)
filefile1 = "/Users/Benjamin/Dropbox/Zhu_Projects/Archetype/Movie/Full50_"+'%03d'%i+".jpg"
filefile2 = "/Users/Benjamin/Dropbox/Zhu_Projects/Archetype/Movie/Loglog_Full50_"+'%03d'%i+".jpg"
fig1.savefig(filefile1)
fig2.savefig(filefile2)
ax1.clear()
ax2.clear()
#print(objs_ori['RA'][iuse[iarchetype[isort_oii[i]]]], objs_ori['DEC'][iuse[iarchetype[isort_oii[i]]]])
#print(objs_ori['RA'][iuse[iarchetype]])
for i in (np.arange(iarchetype.size))[::-1]:
print(objs_ori['RA'][iuse[iarchetype[isort_ha[i]]]], objs_ori['DEC'][iuse[iarchetype[isort_ha[i]]]])
for i in (np.arange(iarchetype.size))[::-1]:
print(objs_ori['RA'][iuse[iarchetype[isort_oii[i]]]], objs_ori['DEC'][iuse[iarchetype[isort_oii[i]]]])
150./360.*24
print(isort_oii)
a_school = np.zeros((11,11), dtype=bool)
cost_school = np.ones(11)
# 1
a_school[0:4, 0] = True
# 2
a_school[0:3, 1] = True
a_school[4, 1] = True
# 3
a_school[0:6, 2] = True
# 4
a_school[0, 3] = True
a_school[2:4, 3] = True
a_school[5:7, 3] = True
# 5
a_school[1:3, 4] = True
a_school[4:6, 4] = True
a_school[7:9, 4] = True
# 6
a_school[2:8, 5] = True
# 7
a_school[3, 6] = True
a_school[5:8, 6] = True
# 8
a_school[4:10, 7] = True
# 9
a_school[4, 8] = True
a_school[7:10, 8] = True
# 10
a_school[7:11, 9] = True
# 11
a_school[9:11, 10] = True
print(a_school*0.1)
fig = plt.figure(figsize=(11,11))
ax = fig.add_subplot(111)
mat = ax.imshow(a_school, cmap='hot_r', interpolation='nearest', vmax=3., vmin=0)
#plt.yticks(np.arange(a_school.shape[0]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
#plt.xticks(np.arange(a_school.shape[1]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
plt.yticks(np.arange(a_school.shape[0]-1)+0.5, [])#+1)
plt.xticks(np.arange(a_school.shape[1]-1)+0.5, [])#+1)
#plt.xticks(rotation=30)
#plt.xlabel('Concert Dates')
# this places 0 or 1 centered in the individual squares
for x in np.arange(a_school.shape[0]):
for y in np.arange(a_school.shape[1]):
ax.annotate(str(a_school[x, y].astype(int))[0], xy=(y, x),
horizontalalignment='center', verticalalignment='center', fontsize=20)
ax.grid(True)
#plt.show()
fig = plt.figure(figsize=(11,11))
ax = fig.add_subplot(111)
#ax.hist2d(dr7_mass['MEDIAN'][iarche], np.log10(dr7_sfr['MEDIAN'][iarche]), bins=160)
#ax.plot(dr7_mass['MEDIAN'][iarche], np.log10(dr7_sfr['MEDIAN'][iarche]), '.')
ax.plot(dr7_mass['MEDIAN'][iarche], dr7_sfr['MEDIAN'][iarche], '.')
ax.set_ylim(2, -2)
dr7_info = fitsio.read('/Users/Benjamin/AstroData/Garching/gal_info_dr7_v5_2.fit.gz', ext=1)
dr7_ssfr = fitsio.read('/Users/Benjamin/AstroData/Garching/gal_totspecsfr_dr7_v5_2.fits.gz', ext=1)
dr7_mass = fitsio.read('/Users/Benjamin/AstroData/Garching/totlgm_dr7_v5_2.fit.gz', ext=1)
idr7 = np.nonzero(np.logical_and(np.logical_and(np.logical_and(np.logical_and(np.logical_and(np.logical_and(
dr7_info['Z']>0.02, dr7_info['Z']<0.30),
dr7_mass['MEDIAN']>7.), dr7_mass['MEDIAN']<13.),
dr7_ssfr['MEDIAN']<-1), dr7_ssfr['MEDIAN']>-20),
dr7_info['SN_MEDIAN']>3.))[0]
iarche_dr7 = np.random.choice(idr7, 10000)
print(iarche_dr7.shape)
print(dr7_info.size, dr7_mass.size, dr7_sfr.size)
dr7_sfr.dtype
fig = plt.figure(figsize=(11,11))
ax = fig.add_subplot(111)
ax.hist2d(dr7_mass['MEDIAN'][idr7], dr7_ssfr['MEDIAN'][idr7], bins=80, cmin=3, cmap='Greys', norm=LogNorm())
#ax.plot(dr7_mass['MEDIAN'][iarche], np.log10(dr7_sfr['MEDIAN'][iarche]), '.')
#ax.plot(dr7_mass[1000:11000]['MEDIAN'], dr7_sfr[1000:11000]['MEDIAN'], '.')
ax.plot(mass[iarchetype], sfr[iarchetype]-mass[iarchetype], '*', ms=15)
ax.set_ylim(-7, -14)
ax.set_xlim(7, 13.)
ax.set_xlabel(r'$\log_{10}$ $M_*$ [M$_\odot$]', fontsize=22)
ax.set_ylabel(r'sSFR [yr$^{-1}$]', fontsize=22)
print(iarchetype.size)
fig = plt.figure(figsize=(11,11))
ax = fig.add_subplot(111)
mat = ax.imshow(a_matrix[0:100, 0:100], cmap='hot_r', interpolation='nearest', vmax=3., vmin=0)
#plt.yticks(np.arange(a_school.shape[0]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
#plt.xticks(np.arange(a_school.shape[1]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
#plt.yticks(np.arange(a_matrix.shape[0]-1)+0.5, [])#+1)
#plt.xticks(np.arange(a_matrix.shape[1]-1)+0.5, [])#+1)
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_xticks([])
ax.set_yticks([])
#plt.xticks(rotation=30)
#plt.xlabel('Concert Dates')
# this places 0 or 1 centered in the individual squares
#for x in np.arange(a_school.shape[0]):
# for y in np.arange(a_school.shape[1]):
# ax.annotate(str(a_school[x, y].astype(int))[0], xy=(y, x),
# horizontalalignment='center', verticalalignment='center', fontsize=20)
#ax.grid(True)
#plt.show()
fig = plt.figure(figsize=(11,11))
ax = fig.add_subplot(111)
mat = ax.imshow(a_matrix, cmap='hot_r', interpolation='nearest', vmax=2.6, vmin=0)
#plt.yticks(np.arange(a_school.shape[0]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
#plt.xticks(np.arange(a_school.shape[1]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
#plt.yticks(np.arange(a_matrix.shape[0]-1)+0.5, [])#+1)
#plt.xticks(np.arange(a_matrix.shape[1]-1)+0.5, [])#+1)
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_xticks([])
ax.set_yticks([])
#plt.xticks(rotation=30)
#plt.xlabel('Concert Dates')
# this places 0 or 1 centered in the individual squares
#for x in np.arange(a_school.shape[0]):
# for y in np.arange(a_school.shape[1]):
# ax.annotate(str(a_school[x, y].astype(int))[0], xy=(y, x),
# horizontalalignment='center', verticalalignment='center', fontsize=20)
#ax.grid(True)
#plt.show()
fig = plt.figure(figsize=(11,11))
ax = fig.add_subplot(111)
mat = ax.imshow(chi2[0:11, 0:11]/newwave.size, cmap='hot', interpolation='nearest', vmax=8., vmin=-2)
#plt.yticks(np.arange(a_school.shape[0]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
#plt.xticks(np.arange(a_school.shape[1]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
plt.yticks(np.arange(11-1)+0.5, [])#+1)
plt.xticks(np.arange(11-1)+0.5, [])#+1)
#ax.set_xticklabels([])
#ax.set_yticklabels([])
#ax.set_xticks([])
#ax.set_yticks([])
#plt.xticks(rotation=30)
#plt.xlabel('Concert Dates')
# this places 0 or 1 centered in the individual squares
#for x in np.arange(a_school.shape[0]):
# for y in np.arange(a_school.shape[1]):
# ax.annotate(str(a_school[x, y].astype(int))[0], xy=(y, x),
# horizontalalignment='center', verticalalignment='center', fontsize=20)
ax.grid(True)
#plt.show()
fig = plt.figure(figsize=(11,11))
ax = fig.add_subplot(111)
mat = ax.imshow(a_matrix[0:11, 0:11], cmap='hot_r', interpolation='nearest', vmax=3., vmin=0)
#plt.yticks(np.arange(a_school.shape[0]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
#plt.xticks(np.arange(a_school.shape[1]-1)+0.5, np.arange(a_school.shape[0]-1)+1)#+1)
plt.yticks(np.arange(11-1)+0.5, [])#+1)
plt.xticks(np.arange(11-1)+0.5, [])#+1)
ax.set_xticklabels([])
ax.set_yticklabels([])
#ax.set_xticks([])
#ax.set_yticks([])
#plt.xticks(rotation=30)
#plt.xlabel('Concert Dates')
# this places 0 or 1 centered in the individual squares
#for x in np.arange(a_school.shape[0]):
# for y in np.arange(a_school.shape[1]):
# ax.annotate(str(a_school[x, y].astype(int))[0], xy=(y, x),
# horizontalalignment='center', verticalalignment='center', fontsize=20)
ax.grid(True)
#plt.show()
fig = plt.figure(figsize=(12,6))
ax = fig.add_subplot(111)
x = tmpwave
iwave_use = np.where(tmpmedian[:,4]>1.E-5)[0]
y = cookb_signalsmooth.smooth(tmpmedian[iwave_use,4], 10)
ax.plot(x[iwave_use], y)
ax.set_xticks([])
ax.set_yticks([])
fig = plt.figure(figsize=(12,6))
ax = fig.add_subplot(111)
x = tmpwave
iwave_use = np.where(tmpmedian[:,8]>1.E-5)[0]
y = cookb_signalsmooth.smooth(tmpmedian[iwave_use,8], 10)
ax.plot(x[iwave_use], y)
ax.set_xticks([])
ax.set_yticks([])
g_school = setcover.SetCover(a_school, cost_school)
g_school.CFT()
np.nonzero(g_school.s)
print(a_school)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.