text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
import parser
import json
import urllib
import pandas as pd
import imp
import numpy as np
import seaborn as sns
from tqdm import tqdm
import math
from sklearn.preprocessing import Imputer
from sklearn import ensemble
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from sklearn.cross_validation import cross_val_score
from PIL import Image
import matplotlib.pyplot as plt
import imdb_movie_content
from collections import Counter
%matplotlib inline
imp.reload(parser)
imp.reload(imdb_movie_content)
df = parser.create_dataframe("movie_contents.json", "movie_budget.json")
df.set_index('movie_title', inplace=True, drop=False)
df.sort_index(inplace=True)
awards_columns = [col for col in df.columns if 'nominated' in col or 'won' in col]
df_awards = df[awards_columns]
awards_per_movie = (len(awards_columns) - df_awards.isnull().sum(axis=1)).to_dict()
df['nb_awards'] = df['movie_title'].map(awards_per_movie)
df = df.drop_duplicates(['movie_title'])
df = df.rename(columns = {'director_fb_links': 'director_fb_likes'})
```
## Awards and Gross
```
d = df['nb_awards'].sort_values(ascending=False)[:15]
plt.figure(figsize=(20,5))
plot = sns.barplot(x=d.index, y=d)
_ = plot.set_xticklabels([elem[:17] for elem in d.index], rotation=15)
_ = plot.set_title('Most awarded (won and nominated) movies')
_ = plot.set_ylabel('Number of awards')
d = df.worldwide_gross.sort_values(ascending=False)[:17]
plt.figure(figsize=(20,5))
plot = sns.barplot(x=d.index, y=d)
_ = plot.set_xticklabels([elem[:20] for elem in d.index], rotation=15)
_ = plot.set_title('Most prolific movies')
_ = plot.set_ylabel('Gross (B$)')
sns.set()
d = df.worldwide_gross.sort_values(ascending=False)[:20]
e = df_awards[df_awards.index.isin(d.index)].isnull().sum(axis=1)
e = len(awards_columns) - e[~e.index.duplicated(keep='first')].reindex(d.index)
margin = 0.05
width = 4*(1.-2.*margin)/15
fig = plt.figure(figsize=(20,5))
ax = fig.add_subplot(111)
ax2 = ax.twinx()
d.plot(kind='bar', color='green', ax=ax, width=width, position=0)
e.plot(kind='bar', color='blue', ax=ax2, width=width, position=1)
ax.set_ylabel('Worldwide Gross (GREEN)')
ax2.set_ylabel('Awards (BLUE)')
ax.set_xlabel('')
ax.set_title('Comparaison between Worldwide Gross and Awards')
_ = ax.set_xticklabels([elem[:17] for elem in d.index], rotation = 30, ha='right')
ax2.grid(False)
```
## Facebook likes
```
d = df['total_cast_fb_likes'].sort_values(ascending=False)[:15]
e = df[df.index.isin(d.index)].num_facebook_like
plt.figure(figsize=(20,5))
plot = sns.barplot(x=d.index, y=d)
_ = plot.set_xticklabels([elem[:17] for elem in d.index], rotation=15)
_ = plot.set_title('Movies with the most "famous" casting')
_ = plot.set_ylabel('Total Facebook Likes (casting + director)')
sns.set()
d = df['total_cast_fb_likes'].sort_values(ascending=False)[:20]
e = df[df.index.isin(d.index)].num_facebook_like.reindex(d.index)
margin = 0.05
width = 4*(1.-2.*margin)/15
fig = plt.figure(figsize=(20,5))
ax = fig.add_subplot(111)
ax2 = ax.twinx()
d.plot(kind='bar', color='green', ax=ax, width=width, position=0)
e.plot(kind='bar', color='blue', ax=ax2, width=width, position=1)
ax.set_ylabel('Casting Likes (GREEN)')
ax2.set_ylabel('Movie Likes (BLUE)')
ax.set_xlabel('')
ax.set_title('Comparaison between Casting Likes and Movie Likes')
_ = ax.set_xticklabels([elem[:17] for elem in d.index], rotation = 30, ha='right')
ax2.grid(False)
```
## Best Actors
### Actor in movie
```
all_actors = [actor for actor in list(set(list(df.actor_1_name) + list(df.actor_2_name) + list(df.actor_3_name))) if pd.notnull(actor)]
imdb_score_per_actor = {}
for actor in all_actors:
imdb_score_per_actor[actor] = df[(df.actor_1_name == actor) | (df.actor_2_name == actor) | (df.actor_3_name == actor)].idmb_score.mean()
millnames = ['',' K',' M',' B']
def millify(n):
if pd.notnull(n):
n = float(n)
millidx = max(0,min(len(millnames)-1,
int(math.floor(0 if n == 0 else math.log10(abs(n))/3))))
return '{:.1f}{}'.format(n / 10**(3 * millidx), millnames[millidx])
else:
return n
gross_per_actor = {}
for actor in all_actors:
gross_per_actor[actor] = df[(df.actor_1_name == actor) | (df.actor_2_name == actor) | (df.actor_3_name == actor)].worldwide_gross.mean()
mini_movie = 3
top_k = 3
best_mini_gross = sorted([(k,v) for k,v in sorted(gross_per_actor.items(), key=lambda x:x[1], reverse=True) if len(df[(df.actor_1_name == k)
| (df.actor_2_name == k)
| (df.actor_3_name == k)]) >= mini_movie], key=lambda x:x[1], reverse=True)[:20]
best_mini_gross_str = [elem[0]+ ', %s (%s movie.s)' % (millify(elem[1]),len(df[(df.actor_1_name == elem[0])
| (df.actor_2_name == elem[0])
| (df.actor_3_name == elem[0])])) for elem in best_mini_gross][:top_k]
best_mini = [(k,v) for k,v in sorted(imdb_score_per_actor.items(), key=lambda x:x[1], reverse=True) if len(df[(df.actor_1_name == k)
| (df.actor_2_name == k)
| (df.actor_3_name == k)]) >= mini_movie][:20]
best_mini_str = [elem[0]+ ', %s (%s movie.s)' % (round(elem[1], 2),len(df[(df.actor_1_name == elem[0])
| (df.actor_2_name == elem[0])
| (df.actor_3_name == elem[0])])) for elem in best_mini][:top_k]
print('The {} best actors are (with minimum {} movies) : \n{}'.format(top_k, mini_movie,
'\n'.join(best_mini_str)))
print('\nThe {} most prolific actors are (with minimum {} movies) : \n{}'.format(top_k, mini_movie,
'\n'.join(best_mini_gross_str)))
plt.figure(figsize=(23,5))
plot = sns.barplot([elem[0] for elem in best_mini], [elem[1] for elem in best_mini])
_ = plot.set_xticklabels([elem[0] for elem in best_mini], rotation=15)
_ = plot.set_title('Most beneficial (IMDB score) actors')
_ = plot.set_ylabel('IMDB score')
plt.figure(figsize=(23,5))
plot = sns.barplot([elem[0] for elem in best_mini_gross], [elem[1] for elem in best_mini_gross])
_ = plot.set_xticklabels([elem[0] for elem in best_mini_gross], rotation=15)
_ = plot.set_title('Most prolific actors')
_ = plot.set_ylabel('Worldwide gross')
```
### First star in movie
```
big_star = df.groupby(['actor_1_name'])['idmb_score', 'worldwide_gross'].mean().sort_values(['idmb_score', 'worldwide_gross'], ascending=False)
big_star['nb_movies'] = big_star.index
big_star['nb_movies'] = big_star['nb_movies'].map(df.groupby(['actor_1_name'])['movie_title'].count().to_dict())
big_star['worldwide_gross'] = big_star['worldwide_gross'].apply(millify)
top_k = 7
print('The {} best actors as most famous actor are :'.format(top_k))
big_star[big_star.nb_movies >= 3].head(top_k)
big_star = df.groupby(['actor_1_name'])['idmb_score', 'worldwide_gross'].mean().sort_values(['worldwide_gross', 'idmb_score'], ascending=False)
big_star['nb_movies'] = big_star.index
big_star['nb_movies'] = big_star['nb_movies'].map(df.groupby(['actor_1_name'])['movie_title'].count().to_dict())
big_star['worldwide_gross'] = big_star['worldwide_gross'].apply(millify)
top_k = 7
print('The {} most prolific actors as most famous actor are :'.format(top_k))
big_star[big_star.nb_movies >= 3].head(top_k)
```
## IMDB rating and other variables
```
d = df['idmb_score'].apply(float).sort_values(ascending=False)[:12]
e = df[df.index.isin(d.index)].num_facebook_like.reindex(d.index)
f = df[df.index.isin(d.index)].worldwide_gross.reindex(d.index)
margin = 0.05
width = 4*(1.-2.*margin)/15
fig = plt.figure(figsize=(20,5))
ax = fig.add_subplot(111)
ax2 = ax.twinx()
ax3= ax2.twinx()
d.plot(kind='bar', color='green', ax=ax, width=width, position=0)
e.plot(kind='bar', color='blue', ax=ax2, width=width, position=1)
f.plot(kind='bar', color='purple', ax=ax3, width=width, position=2)
ax.set_ylabel('IMDB Score (GREEN)')
ax2.set_ylabel('Movie Likes(BLUE) and Gross(PURPLE)')
ax3.set_yticklabels('')
ax2.set_yticklabels('')
ax.set_xlabel('')
_ = ax.set_xticklabels([elem[:17] for elem in d.index], rotation = 30, ha='right')
ax3.grid(False)
ax2.grid(False)
ax.set_title('Gross and Movie Likes compared to IMDB score')
# Correlation Matrix
corr = df[['nb_awards', 'domestic_gross','worldwide_gross',
'total_cast_fb_likes','director_fb_likes', 'production_budget',
'num_critic_for_reviews', 'idmb_score', 'actor_1_fb_likes', 'actor_2_fb_likes', 'actor_3_fb_likes']].corr()
plt.figure(figsize=(8,8))
sns.heatmap(corr, mask=np.zeros_like(corr, dtype=np.bool), cmap=sns.diverging_palette(250, 10, as_cmap=True),
square=True)
plt.title('Correlation matrix for 7 variables and the IMDB Score')
corr
```
## Genres
```
with open('genre.json', 'r') as f:
genres = json.load(f)
imdb_score_per_genre = {}
gross_per_genre = {}
genre_columns = [col for col in df.columns if 'genre_' in col]
df_genres = df[genre_columns]
for genre, value in genres.items():
mask = np.column_stack([df_genres[col] == value for col in df_genres])
df_specific_genre = df.loc[mask.any(axis=1)][['genres', 'idmb_score', 'worldwide_gross']]
imdb_score_per_genre[genre] = df_specific_genre.idmb_score.mean()
gross_per_genre[genre] = df_specific_genre.worldwide_gross.mean()
gross_per_genre = {k:v for k,v in gross_per_genre.items() if pd.notnull(v)}
top_k = 5
print('The {} best genres (in terms of IMDB score) are : \n{}'.format(top_k,
'\n'.join(['%s (%s)' % (elem[0], round(elem[1], 1)) for elem in sorted(imdb_score_per_genre.items(), key=lambda x:x[1], reverse=True)][:top_k])))
print('\nThe {} most prolific genres are : \n{}'.format(top_k,
'\n'.join(['%s (%s)' % (elem[0], millify(elem[1])) for elem in sorted(gross_per_genre.items(), key=lambda x:x[1], reverse=True)][:top_k])))
margin = 0.05
width = 4*(1.-2.*margin)/15
fig = plt.figure(figsize=(20,5))
ax = fig.add_subplot(111)
ax2 = ax.twinx()
df_combine = pd.concat([pd.Series(gross_per_genre), pd.Series(imdb_score_per_genre)], axis=1)
df_combine = df_combine.sort_values(1, ascending=False)
df_combine.columns = ['Gross', 'Score']
df_combine.Gross.plot(kind='bar', color='green', ax=ax, width=width, position=0)
df_combine.Score.plot(kind='bar', color='blue', ax=ax2, width=width, position=1)
ax.set_ylabel('Worldwide Gross in M$ (green)')
ax2.set_ylabel('IMDB Score (blue)')
ax.set_xlabel('')
ax.set_title('Comparaison between Worldwide Gross and IMDB score per genre')
_ = ax.set_xticklabels(pd.Series(imdb_score_per_genre).index, rotation = 30)
ax2.grid(False)
```
## Prediction
### Preprocessing
```
## Fill NA for genres
df.genres = df.genres.fillna('')
## Mean Inputer
col_to_impute = ['actor_1_fb_likes', 'actor_2_fb_likes', 'actor_3_fb_likes',
'domestic_gross', 'duration_sec', 'num_critic_for_reviews', 'num_facebook_like', 'num_user_for_reviews',
'production_budget', 'total_cast_fb_likes', 'worldwide_gross', 'director_fb_likes']
for col in col_to_impute:
column = np.array(df[col]).reshape(1, -1)
imp = Imputer(missing_values='NaN', strategy='mean', axis=1)
df[col] = imp.fit_transform(column)[0]
numerical_cols = list(df.dtypes[df.dtypes != 'object'].index)
not_wanted_cols = ['title_year', 'storyline', 'release_date', 'image_urls', 'movie_title', 'keywords', 'movie_imdb_link', 'num_voted_users'] + genre_columns
df.country = df.country.apply(lambda x:x.split('|'))
df.language = df.language.apply(lambda x:x.split('|'))
list_cols = ['country', 'genres', 'language']
cols_to_transform = [cols for cols in df.columns if cols not in numerical_cols + not_wanted_cols + list_cols]
df2 = df[cols_to_transform]
## Dummies for columns with list
df_col_list = pd.DataFrame()
for col in list_cols:
df_col_list = pd.concat([df_col_list, pd.get_dummies(df[col].apply(pd.Series).stack()).sum(level=0)], axis=1)
## Dummies for columns with string
df_col_string = pd.get_dummies(df2, columns=cols_to_transform)
X_raw = pd.concat([df[numerical_cols], df_col_string, df_col_list], axis=1)
print('Columns dtypes :', Counter(X_raw.dtypes))
y = list(X_raw.idmb_score)
X = X_raw.drop('idmb_score', axis=1)
X_train, X_test, Y_train, Y_test = train_test_split(
X, y, test_size=0.20, random_state=42)
print('Train', X_train.shape, 'Test', X_test.shape)
```
### Choosing ML algorithm
```
gbr = ensemble.GradientBoostingRegressor(n_estimators=1000)
gbr.fit(X_train,Y_train)
print ("Training Score GradientBoosting: ", str(gbr.score(X_train,Y_train)))
print ("Test Score GradientBoosting: " , str(gbr.score(X_test,Y_test)))
abr = ensemble.AdaBoostRegressor(n_estimators=10, learning_rate=0.4, loss='linear')
abr.fit(X_train,Y_train)
print ("Training Score AdaBoostRegressor: ", str(abr.score(X_train,Y_train)))
print ("Test Score AdaBoostRegressor: " , str(abr.score(X_test,Y_test)))
rf=ensemble.RandomForestRegressor(n_estimators=500,oob_score=True, )
rf.fit(X,y)
print ("Training Score RandomForest: ", str(rf.score(X,y)))
print ("Cross Validation (10 fold) Score: " , np.mean(cross_val_score(rf, X_train, Y_train, cv=10)))
```
### Tuning
#### Cross Validation to choose n_estimators
```
rfs = {}
for k in [10, 20, 50, 70, 100, 120, 150, 200]:
rf=ensemble.RandomForestRegressor(n_estimators=k, oob_score=True)
rf.fit(X,y)
rfs[k] = np.mean(cross_val_score(rf, X_train, Y_train, cv=5))
x_plot = list(rfs.keys())
y_plot = list(rfs.values())
f, ax = plt.subplots()
ax.scatter(x_plot, y_plot)
ax.set_title('Variation of the Cross Validation score in function of the number of estimators')
ax.set_xlabel('Number of estimators')
ax.set_ylabel('Cross Validation score')
```
#### Min leaf
```
rfs2 = {}
for k in tqdm(list(range(1, 11, 2))+list(range(11,25,4))):
rf = ensemble.RandomForestRegressor(n_estimators=120, oob_score=True, min_samples_leaf=k)
rf.fit(X,y)
rfs2[k] = rf.oob_score_
x_plot = list(rfs2.keys())
y_plot = list(rfs2.values())
f, ax = plt.subplots()
ax.scatter(x_plot, y_plot)
ax.set_title('Variation of the Cross Validation score in function of the minimum of sample per leaf')
ax.set_xlabel('Minimum of Samples per leaf')
ax.set_ylabel('OOB score')
```
#### max_features
```
rfs2 = {}
for k in ["log2", "auto", "sqrt", 0.2, 0.1, 0.3] :
rf = ensemble.RandomForestRegressor(n_estimators=120, oob_score=True, min_samples_leaf= 1, max_features = k)
rf.fit(X,y)
rfs2[k] = rf.oob_score_
x_plot = range(len(rfs2))# list(rfs2.keys())
y_plot = list(rfs2.values())
print(list(rfs2.keys()))
f, ax = plt.subplots()
ax.scatter(x_plot, y_plot)
ax.set_title('Variation of the Cross Validation score in function of the minimum of sample per leaf')
ax.set_xlabel('Number of estimators')
ax.set_ylabel('Cross Validation score')
```
### Learning
```
rf = ensemble.RandomForestRegressor(n_estimators=120, oob_score=True, max_features=0.2, min_samples_leaf=5)
rf.fit(X,y)
print ("Training Score RandomForest: ", str(rf.score(X,y)))
print ("OOB Score RandomForest: " , str(rf.oob_score_))
```
### Most important features
```
top_k = 15
plt.figure(figsize=(20,5))
names = X_train.columns[np.argsort(rf.feature_importances_)[::-1][:top_k]]
values = np.sort(rf.feature_importances_)[::-1][:top_k]
plot = sns.barplot(x = names, y = values, order=names)
_ = plot.set_xticklabels(names, rotation=15)
_ = plot.set_title('Most important features')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NielsRogge/Transformers-Tutorials/blob/master/ImageGPT/(Un)conditional_image_generation_with_ImageGPT.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Setting-up environment
We first install HuggingFace Transformers.
```
!pip install -q git+https://github.com/huggingface/transformers.git
```
## Unconditional image generation
Next, we initialize the feature extractor and model, and put the model on the GPU.
```
from transformers import ImageGPTFeatureExtractor, ImageGPTForCausalImageModeling
import numpy as np
import torch
feature_extractor = ImageGPTFeatureExtractor.from_pretrained('openai/imagegpt-medium')
model = ImageGPTForCausalImageModeling.from_pretrained('openai/imagegpt-medium')
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model.to(device)
```
Here we only feed the start of sequence (SOS) special token to the model, and let it generate 32x32 = 1024 pixel values using the `generate()` method. Each pixel value is one of 512 possible color clusters.
```
# unconditional generation of 8 images
batch_size = 8
context = torch.full((batch_size, 1), model.config.vocab_size - 1) #initialize with SOS token (with ID 512)
context = torch.tensor(context).to(device)
output = model.generate(input_ids=context, max_length=model.config.n_positions + 1, temperature=1.0, do_sample=True, top_k=40)
#visualize samples with Image-GPT color palette.
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
clusters = feature_extractor.clusters
n_px = feature_extractor.size
samples = output[:,1:].cpu().detach().numpy()
samples_img = [np.reshape(np.rint(127.5 * (clusters[s] + 1.0)), [32, 32, 3]).astype(np.uint8) for s in samples] # convert color cluster tokens back to pixels
f, axes = plt.subplots(1, batch_size, dpi=300)
for img, ax in zip(samples_img, axes):
ax.axis('off')
ax.imshow(img)
```
# Tokenize Cropped Images for Image Completion
Given the upper half part of an image, it's interesting to see how ImageGPT would complete it.
Let's check 8 completions for a given image.
```
import requests
from PIL import Image
url = 'https://assetsnffrgf-a.akamaihd.net/assets/m/502013285/univ/art/502013285_univ_sqr_xl.jpg'
url = "https://avatars.githubusercontent.com/u/326577?v=4"
image = Image.open(requests.get(url, stream=True).raw).convert("RGB")
image
```
We prepare the images using ImageGPTFeatureExtractor, which will resize each image to 32x32x3, normalize it and then apply color clustering. Finally, it will flatten the pixel values out to a long list of 32x32 = 1024 values.
```
encoding = feature_extractor([image for _ in range(8)], return_tensors="pt")
print(encoding.keys())
encoding.pixel_values.shape
```
Next, we only keep the first 512 tokens (pixel values).
```
samples = encoding.pixel_values.numpy()
n_px_crop = 16
# crop top n_px_crop rows. These will be the conditioning tokens
primers = samples[:,:n_px_crop*n_px]
print(primers.shape)
```
We can visualize both the original (lower-resolution and color-clustered) images and the cropped ones:
```
#visualize samples and crops with Image-GPT color palette. Should look similar to original resized images
samples_img = [np.reshape(np.rint(127.5 * (clusters[s] + 1.0)), [n_px, n_px, 3]).astype(np.uint8) for s in samples] # convert color clusters back to pixels
primers_img = [np.reshape(np.rint(127.5 * (clusters[s] + 1.0)), [n_px_crop,n_px, 3]).astype(np.uint8) for s in primers] # convert color clusters back to pixels
f, axes = plt.subplots(1, batch_size, dpi=300)
for img,ax in zip(samples_img, axes):
ax.axis('off')
ax.imshow(img)
f, axes2 = plt.subplots(1, batch_size, dpi=300)
for img,ax in zip(primers_img, axes2):
ax.axis('off')
ax.imshow(img)
```
# Conditional Image Completion
Let's let ImageGPT complete the rest! For this, we also add the start token. Note that we can leverage all possibilities of HuggingFace's generate() method, which are explained in detail in [this blog post](https://huggingface.co/blog/how-to-generate).
```
context = np.concatenate((np.full((batch_size, 1), model.config.vocab_size - 1), primers), axis=1)
context = torch.tensor(context).to(device)
output = model.generate(input_ids=context, max_length=n_px*n_px + 1, temperature=1.0, do_sample=True, top_k=40)
#visualize samples with Image-GPT color palette.
samples = output[:,1:].cpu().detach().numpy()
samples_img = [np.reshape(np.rint(127.5 * (clusters[s] + 1.0)), [n_px, n_px, 3]).astype(np.uint8) for s in samples] # convert color cluster tokens back to pixels
f, axes = plt.subplots(1, batch_size, dpi=300)
for img,ax in zip(samples_img, axes):
ax.axis('off')
ax.imshow(img)
```
Let's combine them into a single image:
```
import numpy as np
row1 = np.hstack(samples_img[:4])
row2 = np.hstack(samples_img[4:])
result = np.vstack([row1, row2])
Image.fromarray(result)
```
## Gradio demo
```
!pip install -q gradio
url = 'https://upload.wikimedia.org/wikipedia/commons/thumb/6/6e/Football_%28soccer_ball%29.svg/1200px-Football_%28soccer_ball%29.svg.png'
image = Image.open(requests.get(url, stream=True).raw)
image
import os
os.system('pip install git+https://github.com/huggingface/transformers --upgrade')
import gradio as gr
from transformers import ImageGPTFeatureExtractor, ImageGPTForCausalImageModeling
import torch
import numpy as np
import requests
from PIL import Image
import matplotlib.pyplot as plt
feature_extractor = ImageGPTFeatureExtractor.from_pretrained("openai/imagegpt-medium")
model = ImageGPTForCausalImageModeling.from_pretrained("openai/imagegpt-medium")
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model.to(device)
# load image examples
urls = ['https://avatars.githubusercontent.com/u/326577?v=4',
'https://upload.wikimedia.org/wikipedia/commons/thumb/6/6e/Football_%28soccer_ball%29.svg/1200px-Football_%28soccer_ball%29.svg.png',
'https://i.imgflip.com/4/4t0m5.jpg',
]
for idx, url in enumerate(urls):
image = Image.open(requests.get(url, stream=True).raw)
image.save(f"image_{idx}.png")
def process_image(image):
# prepare 8 images, shape (8, 1024)
batch_size = 8
encoding = feature_extractor([image for _ in range(batch_size)], return_tensors="pt")
# create primers
samples = encoding.pixel_values.numpy()
n_px = feature_extractor.size
clusters = feature_extractor.clusters
n_px_crop = 16
primers = samples.reshape(-1,n_px*n_px)[:,:n_px_crop*n_px] # crop top n_px_crop rows. These will be the conditioning tokens
# generate (no beam search)
context = np.concatenate((np.full((batch_size, 1), model.config.vocab_size - 1), primers), axis=1)
context = torch.tensor(context).to(device)
output = model.generate(input_ids=context, max_length=n_px*n_px + 1, temperature=1.0, do_sample=True, top_k=40)
# decode back to images (convert color cluster tokens back to pixels)
samples = output[:,1:].cpu().detach().numpy()
samples_img = [np.reshape(np.rint(127.5 * (clusters[s] + 1.0)), [n_px, n_px, 3]).astype(np.uint8) for s in samples]
# stack images horizontally
row1 = np.hstack(samples_img[:4])
row2 = np.hstack(samples_img[4:])
result = np.vstack([row1, row2])
# return as PIL Image
completion = Image.fromarray(result)
return completion
title = "Interactive demo: ImageGPT"
description = "Demo for OpenAI's ImageGPT: Generative Pretraining from Pixels. To use it, simply upload an image or use the example image below and click 'submit'. Results will show up in a few seconds."
article = "<p style='text-align: center'><a href='https://arxiv.org/abs/2109.10282'>ImageGPT: Generative Pretraining from Pixels</a> | <a href='https://openai.com/blog/image-gpt/'>Official blog</a></p>"
examples =[f"image_{idx}.png" for idx in range(len(urls))]
iface = gr.Interface(fn=process_image,
inputs=gr.inputs.Image(type="pil"),
outputs=gr.outputs.Image(type="pil"),
title=title,
description=description,
article=article,
examples=examples)
iface.launch(debug=True)
```
## Verifying code examples
Here we verify whether the code examples in the docs of ImageGPT work fine.
```
from transformers import ImageGPTFeatureExtractor, ImageGPTForCausalImageModeling
import torch
import matplotlib.pyplot as plt
import numpy as np
feature_extractor = ImageGPTFeatureExtractor.from_pretrained('openai/imagegpt-small')
model = ImageGPTForCausalImageModeling.from_pretrained('openai/imagegpt-small')
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model.to(device)
# unconditional generation of 8 images
batch_size = 8
context = torch.full((batch_size, 1), model.config.vocab_size - 1) #initialize with SOS token
context = torch.tensor(context).to(device)
output = model.generate(input_ids=context, max_length=model.config.n_positions + 1, temperature=1.0, do_sample=True, top_k=40)
clusters = feature_extractor.clusters
n_px = feature_extractor.size
samples = output[:,1:].cpu().detach().numpy()
samples_img = [np.reshape(np.rint(127.5 * (clusters[s] + 1.0)), [n_px, n_px, 3]).astype(np.uint8) for s in samples] # convert color cluster tokens back to pixels
f, axes = plt.subplots(1, batch_size, dpi=300)
for img, ax in zip(samples_img, axes):
ax.axis('off')
ax.imshow(img)
from transformers import ImageGPTFeatureExtractor, ImageGPTForImageClassification
from PIL import Image
import requests
url = 'http://images.cocodataset.org/val2017/000000039769.jpg'
image = Image.open(requests.get(url, stream=True).raw)
feature_extractor = ImageGPTFeatureExtractor.from_pretrained('openai/imagegpt-small')
model = ImageGPTForImageClassification.from_pretrained('openai/imagegpt-small')
inputs = feature_extractor(images=image, return_tensors="pt")
outputs = model(**inputs)
logits = outputs.logits
logits.shape
from transformers import ImageGPTFeatureExtractor, ImageGPTModel
from PIL import Image
import requests
url = 'http://images.cocodataset.org/val2017/000000039769.jpg'
image = Image.open(requests.get(url, stream=True).raw)
feature_extractor = ImageGPTFeatureExtractor.from_pretrained('openai/imagegpt-small')
model = ImageGPTModel.from_pretrained('openai/imagegpt-small')
inputs = feature_extractor(images=image, return_tensors="pt")
outputs = model(**inputs)
outputs.last_hidden_state.shape
```
| github_jupyter |
# Stitch Two Networks with Different Spacing
This tutorial explains how to use the stitch function to not only combine two networks into a single domain, but to have OpenPNM automatically creat throats between the two domains based on the spatial proximity of pores on each network.
```
import numpy as np
import scipy as sp
import openpnm as op
import openpnm.models.geometry as gm
import openpnm.models.physics as pm
import openpnm.models.misc as mm
import matplotlib.pyplot as plt
np.set_printoptions(precision=4)
np.random.seed(10)
ws = op.Workspace()
ws.settings["loglevel"] = 40
%matplotlib inline
```
## Generate Two Networks with Different Spacing
```
spacing_lg = 0.00006
layer_lg = op.network.Cubic(shape=[10, 10, 1], spacing=spacing_lg)
spacing_sm = 0.00002
layer_sm = op.network.Cubic(shape=[30, 5, 1], spacing=spacing_sm)
```
## Position Networks Appropriately, then Stitch Together
```
# Start by assigning labels to each network for identification later
layer_sm.set_label("small", pores=layer_sm.Ps, throats=layer_sm.Ts)
layer_lg.set_label("large", pores=layer_lg.Ps, throats=layer_lg.Ts)
# Next manually offset CL one full thickness relative to the GDL
layer_sm['pore.coords'] -= [0, spacing_sm*5, 0]
layer_sm['pore.coords'] += [0, 0, spacing_lg/2 - spacing_sm/2] # And shift up by 1/2 a lattice spacing
# Finally, send both networks to stitch which will stitch CL onto GDL
from openpnm.topotools import stitch
stitch(network=layer_lg, donor=layer_sm,
P_network=layer_lg.pores('back'),
P_donor=layer_sm.pores('front'),
len_max=0.00005)
combo_net = layer_lg
combo_net.name = 'combo'
```
## Quickly Visualize the Network
Let's just make sure things are working as planned using OpenPNMs basic visualization tools:
```
fig = plt.figure(figsize=[10, 10])
fig = op.topotools.plot_connections(network=combo_net, fig=fig)
```
## Create Geometry Objects for Each Layer
```
Ps = combo_net.pores('small')
Ts = combo_net.throats('small')
geom_sm = op.geometry.GenericGeometry(network=combo_net, pores=Ps, throats=Ts)
Ps = combo_net.pores('large')
Ts = combo_net.throats('small', mode='not')
geom_lg = op.geometry.GenericGeometry(network=combo_net, pores=Ps, throats=Ts)
```
### Add Geometrical Properties to the *Small* Domain
The *small* domain will be treated as a continua, so instead of assigning pore sizes we want the 'pore' to be same size as the lattice cell.
```
geom_sm['pore.diameter'] = spacing_sm
geom_sm['pore.area'] = spacing_sm**2
geom_sm['throat.diameter'] = spacing_sm
geom_sm['throat.area'] = spacing_sm**2
geom_sm['throat.length'] = 1e-12 # A very small number to represent nearly 0-length
geom_sm.add_model(propname='throat.endpoints',
model=gm.throat_endpoints.circular_pores)
geom_sm.add_model(propname='throat.length',
model=gm.throat_length.piecewise)
geom_sm.add_model(propname='throat.conduit_lengths',
model=gm.throat_length.conduit_lengths)
```
### Add Geometrical Properties to the *Large* Domain
```
geom_lg['pore.diameter'] = spacing_lg*np.random.rand(combo_net.num_pores('large'))
geom_lg.add_model(propname='pore.area',
model=gm.pore_area.sphere)
geom_lg.add_model(propname='throat.diameter',
model=mm.from_neighbor_pores,
prop='pore.diameter', mode='min')
geom_lg.add_model(propname='throat.area',
model=gm.throat_area.cylinder)
geom_lg.add_model(propname='throat.endpoints',
model=gm.throat_endpoints.circular_pores)
geom_lg.add_model(propname='throat.length',
model=gm.throat_length.piecewise)
geom_lg.add_model(propname='throat.conduit_lengths',
model=gm.throat_length.conduit_lengths)
```
## Create Phase and Physics Objects
```
air = op.phases.Air(network=combo_net, name='air')
phys_lg = op.physics.GenericPhysics(network=combo_net, geometry=geom_lg, phase=air)
phys_sm = op.physics.GenericPhysics(network=combo_net, geometry=geom_sm, phase=air)
```
Add pore-scale models for diffusion to each Physics:
```
phys_lg.add_model(propname='throat.diffusive_conductance',
model=pm.diffusive_conductance.ordinary_diffusion)
phys_sm.add_model(propname='throat.diffusive_conductance',
model=pm.diffusive_conductance.ordinary_diffusion)
```
For the *small* layer we've used a normal diffusive conductance model, which when combined with the diffusion coefficient of air will be equivalent to open-air diffusion. If we want the *small* layer to have some tortuosity we must account for this:
```
porosity = 0.5
tortuosity = 2
phys_sm['throat.diffusive_conductance'] *= (porosity/tortuosity)
```
Note that this extra line is NOT a pore-scale model, so it will be over-written when the `phys_sm` object is regenerated.
### Add a Reaction Term to the Small Layer
A standard n-th order chemical reaction is $ r=k \cdot x^b $, or more generally: $ r = A_1 \cdot x^{A_2} + A_3 $. This model is available in `OpenPNM.Physics.models.generic_source_terms`, and we must specify values for each of the constants.
```
# Set Source Term
air['pore.A1'] = -1e-10 # Reaction pre-factor
air['pore.A2'] = 1 # Reaction order
air['pore.A3'] = 0 # A generic offset that is not needed so set to 0
phys_sm.add_model(propname='pore.reaction',
model=pm.generic_source_term.power_law,
A1='pore.A1', A2='pore.A2', A3='pore.A3',
X='pore.concentration',
regen_mode='deferred')
```
## Perform a Diffusion Calculation
```
Deff = op.algorithms.ReactiveTransport(network=combo_net, phase=air)
Ps = combo_net.pores(['large', 'front'], mode='intersection')
Deff.set_value_BC(pores=Ps, values=1)
Ps = combo_net.pores('small')
Deff.set_source(propname='pore.reaction', pores=Ps)
Deff.settings['conductance'] = 'throat.diffusive_conductance'
Deff.settings['quantity'] = 'pore.concentration'
Deff.run()
```
## Visualize the Concentration Distribution
And the result would look something like this:
```
fig = plt.figure(figsize=[10, 10])
fig = op.topotools.plot_coordinates(network=combo_net, c=Deff['pore.concentration'],
cmap='jet', markersize=40, fig=fig)
```
| github_jupyter |
# Lab 02 : Vanilla GAN with MLP - solution
The goal is to implement a GAN architecture with MLPs to generate new MNIST images.</br>
```
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
# mount google drive
from google.colab import drive
drive.mount('/content/gdrive')
path_to_file = '/content/gdrive/My Drive/CS4243_codes/codes/labs_lecture15/lab02_GAN_MLP'
print(path_to_file)
# move to Google Drive directory
os.chdir(path_to_file)
!pwd
import torch
import torch.nn as nn
import torch.optim as optim
import utils
import time
```
### GPU is required to train GAN
```
device= torch.device("cuda")
#device= torch.device("cpu")
print(device)
# Libraries
import matplotlib.pyplot as plt
import logging
logging.getLogger().setLevel(logging.CRITICAL) # remove warnings
```
### MNIST dataset
```
from utils import check_mnist_dataset_exists
data_path=check_mnist_dataset_exists()
train_data=torch.load(data_path+'mnist/train_data.pt')
train_label=torch.load(data_path+'mnist/train_label.pt')
print(train_data.size())
```
### Network architecture
```
# Global constants
# n : nb of pixels along each spatial dimension
# dz : latent dimension
# d : hidden dimension
# b : batch size
n = train_data.size(1)
dz = n
d = 32* (n//4)**2 # hidden dimension is a function of image size
b = 64
# Define the generator and discriminator networks
class generator(nn.Module):
def __init__(self):
super(generator, self).__init__()
# COMPLETE HERE
self.linear1 = nn.Linear(dz, d, bias=True)
self.bn1 = nn.BatchNorm1d(d)
self.linear2 = nn.Linear(d, d, bias=True)
self.bn2 = nn.BatchNorm1d(d)
self.linear3 = nn.Linear(d, n**2, bias=True)
def forward(self, z):
# COMPLETE HERE
h = self.linear1(z) # [b, d]
h = self.bn1(h)
h = torch.relu(h)
h = self.linear2(h) # [b, d]
h = self.bn2(h)
h = torch.relu(h)
h = self.linear3(h) # [b, n**2]
h = h.view(-1, 1, n, n) # [b, 1, n, n], nb colors = 1
#g_z = torch.tanh(h) # in general [-1,1]
g_z = torch.sigmoid(h) # for mnist [0,1]
return g_z
class discriminator(nn.Module):
def __init__(self):
super(discriminator, self).__init__()
# COMPLETE HERE
self.linear1 = nn.Linear(n**2, d, bias=True)
self.bn1 = nn.BatchNorm1d(d)
self.linear2 = nn.Linear(d, d, bias=True)
self.bn2 = nn.BatchNorm1d(d)
self.linear3 = nn.Linear(d, 1, bias=True)
def forward(self, h):
# COMPLETE HERE
h = h.view(-1, n**2) # [b, n**2]
h = self.linear1(h) # [b, d]
h = self.bn1(h)
h = torch.relu(h)
h = self.linear2(h) # [b, d]
h = self.bn2(h)
h = torch.relu(h)
h = self.linear3(h) # [b, 1], nb colors = 1
d_h = torch.sigmoid(h)
return d_h
# Instantiate the network
net_g = generator()
net_g = net_g.to(device)
print(net_g)
utils.display_num_param(net_g)
net_d = discriminator()
net_d = net_d.to(device)
print(net_d)
utils.display_num_param(net_d)
# Test the forward pass, backward pass and gradient update with a single batch
init_lr = 0.001
optimizer_g = torch.optim.Adam(net_g.parameters(), lr=init_lr)
optimizer_d = torch.optim.Adam(net_d.parameters(), lr=init_lr)
b = 10
idx = torch.LongTensor(b).random_(0,60000)
x_real = train_data[idx,:,:].view(b,-1).to(device) # [b, n**2]
print(x_real.size())
z = torch.rand(b, dz).to(device) # [b, dz]
print(z.size())
p_one = torch.ones(b, 1).to(device)
p_zero = torch.zeros(b, 1).to(device)
# update g
optimizer_g.zero_grad()
x_fake = net_g(z) # [b, 1, n, n]
p_fake = net_d(x_fake) # [b, 1]
print(x_fake.size(), p_fake.size())
loss_fake = nn.BCELoss()(p_fake, p_one)
loss = loss_fake
loss.backward()
optimizer_g.step()
# update d
optimizer_d.zero_grad()
x_fake = net_g(z) # [b, 1, n, n]
p_fake = net_d(x_fake) # [b, 1]
p_real = net_d(x_real.view(-1,n,n).unsqueeze(1)) # [b, 1]
print(x_fake.size(), p_fake.size(), p_real.size())
loss_real = nn.BCELoss()(p_real, p_one)
loss_fake = nn.BCELoss()(p_fake, p_zero)
loss = loss_real + loss_fake
loss.backward()
optimizer_d.step()
# Training loop
net_g = generator()
net_g = net_g.to(device)
print(net_g)
utils.display_num_param(net_g)
net_d = discriminator()
net_d = net_d.to(device)
print(net_d)
utils.display_num_param(net_d)
# Optimizer
init_lr = 0.0002
optimizer_g = torch.optim.Adam(net_g.parameters(), lr=init_lr, betas=(0.5, 0.999))
optimizer_d = torch.optim.Adam(net_d.parameters(), lr=init_lr, betas=(0.5, 0.999))
nb_batch = 200 # GPU # Nb of mini-batches per epoch
#nb_batch = 20 # CPU # Nb of mini-batches per epoch
b = 64 # Batch size
p_one = torch.ones(b, 1).to(device)
p_zero = torch.zeros(b, 1).to(device)
start=time.time()
for epoch in range(50):
running_loss_d = 0.0
running_loss_g = 0.0
num_batches = 0
for _ in range(nb_batch):
# FORWARD AND BACKWARD PASS
idx = torch.LongTensor(b).random_(0,60000)
x_real = train_data[idx,:,:].view(b,-1).to(device) # [b, n**2]
z = torch.rand(b, dz).to(device) # Uniform distribution # [b, dz]
# update d
optimizer_d.zero_grad()
x_fake = net_g(z) # [b, 1, n, n]
p_fake = net_d(x_fake) # [b, 1]
p_real = net_d(x_real.view(-1,n,n).unsqueeze(1)) # [b, 1]
loss_real = nn.BCELoss()(p_real, p_one)
loss_fake = nn.BCELoss()(p_fake, p_zero)
loss = loss_real + loss_fake
loss_d = loss.detach().item()
loss.backward()
optimizer_d.step()
# update g
optimizer_g.zero_grad()
x_fake = net_g(z) # [b, 1, n, n]
p_fake = net_d(x_fake) # [b, 1]
loss_fake = nn.BCELoss()(p_fake, p_one)
loss = loss_fake
loss_g = loss.detach().item()
loss.backward()
optimizer_g.step()
# COMPUTE STATS
running_loss_d += loss_d
running_loss_g += loss_g
num_batches += 1
# AVERAGE STATS THEN DISPLAY
total_loss_d = running_loss_d/ num_batches
total_loss_g = running_loss_g/ num_batches
elapsed = (time.time()-start)/60
print('epoch=',epoch, '\t time=', elapsed,'min', '\t lr=', init_lr ,'\t loss_d=', total_loss_d ,'\t loss_g=', total_loss_g )
if not epoch%5:
plt.imshow(x_fake.view(b,n,n).detach().cpu()[0,:,:], cmap='gray'); plt.show()
# Generate a few images
b = 10
z = torch.rand(b, dz) # Uniform distribution
z = z.to(device)
x_new = net_g(z).view(b,n,n).detach().cpu()
for k in range(b):
plt.imshow(x_new[k,:,:], cmap='gray'); plt.show()
```
| github_jupyter |
# Traversing and Analyzing Coronal Hole Connectivity Graph
```
import pickle
import networkx as nx
import os
import copy
import numpy as np
import scipy
from scipy import stats
import matplotlib.pyplot as plt
import matplotlib.colors as c
import matplotlib
import cv2
import chmap.database.db_classes as db_class
import chmap.database.db_funs as db_funs
import chmap.maps.magnetic.flux.br_flux as br_flux
import datetime as dt
plt.rcParams['savefig.facecolor']='white'
matplotlib.rcParams.update({'font.size': 12})
os.chdir(os.path.dirname('/Users/opalissan/PycharmProjects/CHDv1/chmap'))
from chmap.coronal_holes.tracking.tools.plots import plot_coronal_hole
```
### Read in Coronal Hole Connectivity Graph
Saved as a pickle file in DropBox.
```
res_dir = '/Users/opalissan/desktop/CHT_RESULTS/2010-12-29-2011-04-08c2hr'
map_dir = "/Users/opalissan/desktop/CH_DB"
user = "opalissan"
password = ""
db_session = db_funs.init_db_conn_old(db_name='mysql-Q', chd_base=db_class.Base,
user=user, password=password)
# define window that we are willing to look at for magnetic flux
window_half_width = dt.timedelta(hours=12)
pickle_file = os.path.join(res_dir + '/connectivity_graph.pkl')
graph = pickle.load(open(pickle_file, "rb"))
G = graph.G
```
### Analyze Properties of the Connectivity Graph
```
print("Total number of nodes: ", G.number_of_nodes())
print("Total number of edges: ", G.number_of_edges())
print("Total number of subplots: ", len(list(nx.connected_components(G))))
# order subgraphs based on average node area
subgraph_ordered = graph.order_subgraphs_based_on_area()
# save the total number of nodes in a subgraph
num_nodes_list = []
for ii, g in enumerate(subgraph_ordered):
subgraph = G.subgraph(g)
num_nodes_list.append(subgraph.number_of_nodes())
fig, ax = plt.subplots(figsize=(20, 6))
_ = ax.scatter(np.arange(len(num_nodes_list)), num_nodes_list)
_ = ax.set_ylabel("Number of Nodes")
_ = ax.set_xlabel("Subgraph Index")
_ = ax.set_title("Connected Subgraphs Total Number of Nodes \n Ordered by Average Node Area")
plt.savefig(res_dir + '/figures/subgraph_node_size.png')
```
#### *Note*
From the plot above, it is visible that the graph has three main subgraphs. *Assumption*: One of the main subgraphs is connected to the north pole and the other to the south. Lets examine the first and second subgraph (with the largest number of nodes $\approx$ 3000)
```
main_subgraph_1 = G.subgraph(subgraph_ordered[0])
main_subgraph_2 = G.subgraph(subgraph_ordered[1])
# list with all weights in G.
w_list = [w["weight"] for u,v,w in G.edges.data()]
# analyze the weight between nodes that have the same id and nodes that have a different id.
adj_diff_list = []
adj_same_list = []
# loop over all edges in G.
for u,v,w in G.edges.data():
# different id.
if G.nodes[u]["id"] != G.nodes[v]["id"]:
adj_diff_list.append(w["weight"])
# same id.
else:
adj_same_list.append(w["weight"])
print("Number of edges between nodes of the same ID = ", len(adj_same_list))
print("Number of edges between nodes with different ID = ", len(adj_diff_list))
fig, ax = plt.subplots(ncols=3, nrows=2, sharex=True, figsize=(12, 6))
mu, sigma = scipy.stats.norm.fit(w_list)
n , bins, _ = ax[0][0].hist(w_list, 25, density=True)
best_fit_line = stats.norm.pdf(bins, mu, sigma)
_ = ax[0][0].plot(bins, best_fit_line, color="red")
_ = ax[0][0].set_title("All Edge Weights \n $\mu$ = %.2f, median = %.2f, $\sigma$ = %.2f" % (mu, np.median(w_list), sigma))
_ = ax[0][0].set_ylabel("Probability density")
_ = ax[1][0].hist(w_list, 25)
_ = ax[1][0].set_xlabel("Weight")
_ = ax[1][0].set_ylabel("Count")
mu2, sigma2 = scipy.stats.norm.fit(adj_diff_list)
n , bins2, _ = ax[0][1].hist(adj_diff_list, 25, density=True, color="orange")
best_fit_line2 = stats.norm.pdf(bins2, mu2, sigma2)
_ = ax[0][1].plot(bins2, best_fit_line2, color="red")
_ = ax[0][1].set_title("Adjacent different IDs \n $\mu$ = %.2f, median = %.2f, $\sigma$ = %.2f" % (mu2, np.median(adj_diff_list), sigma2))
_ = ax[0][1].set_ylabel("Probability Density")
_ = ax[1][1].hist(adj_diff_list, 25, color="orange")
_ = ax[1][1].set_xlabel("Weight")
_ = ax[1][1].set_ylabel("Count")
mu3, sigma3 = scipy.stats.norm.fit(adj_same_list)
n , bins3, _ = ax[0][2].hist(adj_same_list, 25, density=True, color="green")
best_fit_line3 = stats.norm.pdf(bins3, mu3, sigma3)
_ = ax[0][2].plot(bins3, best_fit_line3, color="red")
_ = ax[0][2].set_title("Adjacent same ID \n $\mu$ = %.2f, median = %.2f, $\sigma$ = %.2f" % (mu3, np.median(adj_same_list), sigma3))
_ = ax[0][2].set_ylabel("Probability Density")
_ = ax[1][2].hist(adj_same_list, 25, color="green")
_ = ax[1][2].set_xlabel("Weight")
_ = ax[1][2].set_ylabel("Count")
fig.suptitle("Main Graph")
plt.tight_layout()
plt.savefig(res_dir + '/figures/edge_distribution_G.png')
```
# How to Traverse the Graph to explore all the nodes related to the North Pole Coronal Hole
```
for node in G.nodes:
if G.nodes[node]["id"] == 205 and G.nodes[node]["frame_num"] == 1205:
print(G.nodes[node])
print(node)
if G.nodes[node]["id"] == 8 and G.nodes[node]["frame_num"] == 1:
print(G.nodes[node])
print(node)
start = "1_8_0"
end = "1205_205_0"
```
# Create a *directed* version of our main Graph
```
DiG = G.to_directed()
```
# Shortest Path by Dijkstra's algorithm
```
# define the cost function
def func(n1, n2, d):
if G.nodes[n1]["id"] == DiG.nodes[n2]["id"]:
return 0
else:
weight = DiG.get_edge_data(n1, n2)["weight"]
return 1 - weight
# apply dijkstra's algorithm
path1 = nx.algorithms.shortest_paths.weighted.dijkstra_path(DiG, source=start, target=end, weight=func)
set_of_classes = set()
for node in path1:
#print(str(node) + ", id: " + str(G.nodes[node]["id"]) + ", frame: "+ str(G.nodes[node]["frame_num"]) )
set_of_classes.add(G.nodes[node]["id"])
list(set_of_classes)
path_g = G.subgraph(path1)
unique_id_class = []
for id_class in list(set_of_classes):
for node in path_g.nodes:
if path_g.nodes[node]["id"] == id_class:
unique_id_class.append(node)
break
unique_id_class
# set time array for plotting purposes
timearray = np.arange('2010-12-29T02', '2011-04-08T08',np.timedelta64(2,'h'), dtype='datetime64')
fig, ax = plt.subplots(figsize=(10, 10))
# draw graph, nodes positions are based on their count and frame_num.
# labels are the coronal hole id number.
pos, labels = graph.get_plot_features(sub_graph=path_g)
edge_weights = nx.get_edge_attributes(G=path_g, name='weight')
edges, weights = zip(*edge_weights.items())
# plot nodes and labels.
nx.draw(path_g, pos=pos, font_weight='bold', ax=ax, node_size=700,
node_color=[c.to_rgba(np.array(path_g.nodes[ch]["color"]) / 255)
for ch in path_g.nodes], edgelist=[])
#nx.draw_networkx_labels(G=path_g, pos=pos, labels=labels, ax=ax, font_size=8)
for ch in unique_id_class:
ax.scatter([],[], c=[c.to_rgba(np.array(path_g.nodes[ch]["color"]) / 255)], label='ID {}'.format(path_g.nodes[ch]["id"]))
nx.draw_networkx_edges(path_g, pos=pos, edge_color=weights, edgelist=edges,
edge_cmap=plt.cm.get_cmap('Greys'), edge_vmin=0,
edge_vmax=1, width=3, ax=ax)
nx.draw_networkx_edge_labels(G=path_g, pos=pos, edge_labels=edge_weights, ax=ax, alpha=1, font_size=10)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.set_xlim(tuple(sum(i) for i in zip(ax.get_xlim(), (-0.5, 0.5))))
# set y ticks
ax.yaxis.get_major_locator().set_params(integer=True)
ax.set_yticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_yticklabels(timearray[::150], fontsize=12)
ax.tick_params(axis='y', rotation=40)
ax.axis('on')
_ = ax.set_title("Coronal Hole Connectivity North")
_ = plt.gca().legend()
plt.savefig(res_dir + '/figures/north_ch_shortest_path_res_8_to_205.png')
```
# Spatio-temporal Analysis of the North pole
```
frame_array = np.arange(1, 1205)
fig, ax = plt.subplots(figsize=(15, 5))
for frame in frame_array:
holder = []
for node in path_g:
if path_g.nodes[node]["frame_num"] == frame:
holder.append(node)
area = 0
for node in holder:
area += path_g.nodes[node]["area"]
if area != 0:
ax.scatter(frame, area, c=[c.to_rgba(np.array(path_g.nodes[holder[0]]["color"]) / 255)])
for ch in unique_id_class:
ax.scatter([],[], c=[c.to_rgba(np.array(path_g.nodes[ch]["color"]) / 255)], label='ID {}'.format(path_g.nodes[ch]["id"]))
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150], fontsize=12)
ax.tick_params(axis='x', rotation=15)
# label axis
_ = ax.set_ylabel("Area ($R_{\odot}^2$)")
_ = ax.set_title("North Pole Coronal Hole Shortest Path Spatiotemporal Properties")
_ = plt.legend()
plt.savefig(res_dir + '/figures/north_ch_shortest_path_area.png')
fig, ax = plt.subplots(figsize=(15, 5))
for frame in frame_array:
holder = []
for node in path_g:
if path_g.nodes[node]["frame_num"] == frame:
holder.append(node)
area = 0
for node in holder:
area += path_g.nodes[node]["area"]
if area != 0:
ax.scatter(frame, area, c='b')
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
# label axis
_ = ax.set_ylabel("Area ($R_{\odot}^2$)")
_ = ax.set_title("North Pole Coronal Hole Shortest Path Spatiotemporal Properties")
plt.savefig(res_dir + '/figures/north_ch_shortest_path_area_all_blue.png')
net_flux_array = []
abs_flux_array = []
for frame in frame_array:
holder = []
for node in path_g:
if path_g.nodes[node]["frame_num"] == frame:
holder.append(node)
net_flux = 0
abs_flux = 0
if len(holder) > 0:
# get the node timestamp
timestamp = G.nodes[holder[0]]["frame_timestamp"]
# get pickle file with contour properties.
pickle_file = str(timestamp).replace(':', '-') + ".pkl"
pickle_file = pickle_file.replace(" ", "-")
frame_read = pickle.load(open(os.path.join(res_dir + "/pkl/" + pickle_file), "rb"))
ch_list = frame_read.contour_list
ii = 0
while ii < len(holder):
node = holder[ii]
for ch in ch_list:
if ch.id == path_g.nodes[node]["id"] and ch.count == path_g.nodes[node]["count"]:
nf, af = br_flux.coronal_flux(db_session, ch, timestamp,
map_dir, window_half_width=window_half_width)
ii +=1
net_flux += nf
abs_flux += af
net_flux_array.append(net_flux)
abs_flux_array.append(abs_flux)
fig, ax = plt.subplots(figsize=(15, 5))
ax.scatter(frame_array, net_flux_array, label="Net Flux")
ax.scatter(frame_array, abs_flux_array, label="Absolute Flux")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
plt.legend()
# label axis
_ = ax.set_ylabel("$\Phi_{B}$")
_ = ax.set_title("North Pole Coronal Hole Shortest Path Flux")
plt.savefig(res_dir + '/figures/north_ch_shortest_path_net_flux.png')
```
# North Shortest Path and *Neighbors*
```
THRESH = 0.5
north_and_ne = path_g.copy()
# add all nodes that are adjacent to nodes in shortest path
for node in path_g:
for adj_node in G[node]:
# if the edge weight is greater than threshold, then add the neighbor node.
if G[node][adj_node]['weight'] >= THRESH:
# check if the node already exists in shortest path
if adj_node not in north_and_ne.nodes:
# add the node.
north_and_ne.add_node(str(adj_node),
area=G.nodes[adj_node]["area"],
id=G.nodes[adj_node]["id"],
frame_num=G.nodes[adj_node]["frame_num"],
frame_timestamp=G.nodes[adj_node]["frame_timestamp"],
count=G.nodes[adj_node]["count"],
color=G.nodes[adj_node]["color"])
for u,v,w in G.edges.data():
# if the two nodes exist in the shorstest path and neighbors then add the edge.
if u in north_and_ne and v in north_and_ne:
if not north_and_ne.has_edge(u, v):
north_and_ne.add_edge(u, v, weight=w)
ne_classes = set()
for node in north_and_ne:
ne_classes.add(G.nodes[node]["id"])
str(ne_classes)
fig, ax = plt.subplots(figsize=(15, 5))
for frame in frame_array:
holder = []
for node in north_and_ne:
if north_and_ne.nodes[node]["frame_num"] == frame:
holder.append(node)
area = 0
for node in holder:
area += north_and_ne.nodes[node]["area"]
#if len(holder) == 0:
# print("no nodes in this frame", frame)
if area != 0:
ax.scatter(frame, area, c=['r' if len(holder) > 1 else 'b'])
# set color label
ax.scatter([], [], c="r", label="npf $>$ 1")
ax.scatter([], [], c="b", label="npf $\leq$ 1")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
_ = ax.set_ylabel("Area ($R_{\odot}^2$)")
# _ = ax.set_title("North Pole Coronal Hole Shortest Path and Neighbors Spatiotemporal Properties \n Edge Threshold is " + str(THRESH) + "\n " + str(ne_classes))
_ = ax.set_title("North Pole Coronal Hole Shortest Path and Neighbors Spatiotemporal Properties \n Edge Threshold is " + str(THRESH))
_ = plt.legend()
plt.savefig(res_dir + '/figures/north_ch_shortest_path_and_ne_area' + str(THRESH) + '.png', bbox_inches='tight')
net_flux_array_ne = []
abs_flux_array_ne = []
for frame in frame_array:
holder = []
for node in north_and_ne:
if north_and_ne.nodes[node]["frame_num"] == frame:
holder.append(node)
net_flux = 0
abs_flux = 0
if len(holder) > 0:
# get the node timestamp
timestamp = G.nodes[holder[0]]["frame_timestamp"]
# get pickle file with contour properties.
pickle_file = str(timestamp).replace(':', '-') + ".pkl"
pickle_file = pickle_file.replace(" ", "-")
frame_read = pickle.load(open(os.path.join(res_dir + "/pkl/" + pickle_file), "rb"))
ch_list = frame_read.contour_list
ii = 0
while ii < len(holder):
node = holder[ii]
for ch in ch_list:
if ch.id == north_and_ne.nodes[node]["id"] and ch.count == north_and_ne.nodes[node]["count"]:
nf, af = br_flux.coronal_flux(db_session, ch, timestamp,
map_dir, window_half_width=window_half_width)
ii +=1
net_flux += nf
abs_flux += af
net_flux_array_ne.append(net_flux)
abs_flux_array_ne.append(abs_flux)
fig, ax = plt.subplots(figsize=(15, 5))
ax.scatter(frame_array, net_flux_array_ne, label="Net Flux")
ax.scatter(frame_array, abs_flux_array_ne, label="Absolute Flux")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
plt.legend()
# label axis
_ = ax.set_ylabel("$\Phi_{B}$")
_ = ax.set_title("North Pole Coronal Hole Shortest Path and Neighbors Flux \n Edge Threshold is " + str(THRESH))
plt.savefig(res_dir + '/figures/north_ch_shortest_path_and_ne_flux' + str(THRESH) + '.png', bbox_inches='tight')
```
## shortest path + neighbors classes
```
north_and_ne_c = north_and_ne.copy()
# add all nodes that are adjacent to nodes in shortest path
for node in G:
if node not in north_and_ne_c.nodes and G.nodes[node]["id"] in ne_classes:
# add the node.
north_and_ne_c.add_node(str(node),
area=G.nodes[node]["area"],
id=G.nodes[node]["id"],
frame_num=G.nodes[node]["frame_num"],
frame_timestamp=G.nodes[node]["frame_timestamp"],
count=G.nodes[node]["count"],
color=G.nodes[node]["color"])
for u,v,w in G.edges.data():
# if the two nodes exist in the shorstest path and neighbors then add the edge.
if u in north_and_ne_c and v in north_and_ne_c:
if not north_and_ne_c.has_edge(u, v):
north_and_ne_c.add_edge(u, v, weight=w)
fig, ax = plt.subplots(figsize=(15, 5))
for frame in frame_array:
holder = []
for node in north_and_ne_c:
if north_and_ne_c.nodes[node]["frame_num"] == frame:
holder.append(node)
area = 0
for node in holder:
area += north_and_ne_c.nodes[node]["area"]
#if len(holder) == 0:
# print("no nodes in this frame", frame)
if area != 0:
ax.scatter(frame, area, c=['r' if len(holder) > 1 else 'b'])
ax.scatter([], [], c="r", label="npf $>$ 1")
ax.scatter([], [], c="b", label="npf $\leq$ 1")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
_ = ax.set_ylabel("Area ($R_{\odot}^2$)")
# _ = ax.set_title("North Pole Coronal Hole Shortest Path and Neighbors Spatiotemporal Properties \n Edge Threshold is " + str(THRESH) + "\n " + str(ne_classes))
_ = ax.set_title("North Pole Coronal Hole Shortest Path and Neighbors Spatiotemporal Properties \n Edge Threshold is " + str(THRESH))
_ = plt.legend()
plt.savefig(res_dir + '/figures/north_ch_shortest_path_and_ne_c_area' + str(THRESH) + '.png', bbox_inches='tight')
net_flux_array_ne_c = []
abs_flux_array_ne_c = []
for frame in frame_array:
holder = []
for node in north_and_ne_c:
if north_and_ne_c.nodes[node]["frame_num"] == frame:
holder.append(node)
net_flux = 0
abs_flux = 0
if len(holder) > 0:
# get the node timestamp
timestamp = G.nodes[holder[0]]["frame_timestamp"]
# get pickle file with contour properties.
pickle_file = str(timestamp).replace(':', '-') + ".pkl"
pickle_file = pickle_file.replace(" ", "-")
frame_read = pickle.load(open(os.path.join(res_dir + "/pkl/" + pickle_file), "rb"))
ch_list = frame_read.contour_list
ii = 0
while ii < len(holder):
node = holder[ii]
for ch in ch_list:
if ch.id == north_and_ne_c.nodes[node]["id"] and ch.count == north_and_ne_c.nodes[node]["count"]:
nf, af = br_flux.coronal_flux(db_session, ch, timestamp,
map_dir, window_half_width=window_half_width)
ii +=1
net_flux += nf
abs_flux += af
net_flux_array_ne_c.append(net_flux)
abs_flux_array_ne_c.append(abs_flux)
fig, ax = plt.subplots(figsize=(15, 5))
ax.scatter(frame_array, net_flux_array_ne, label="Net Flux")
ax.scatter(frame_array, abs_flux_array_ne, label="Absolute Flux")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
plt.legend()
# label axis
_ = ax.set_ylabel("$\Phi_{B}$")
_ = ax.set_title("North Pole Coronal Hole Shortest Path and Neighbors Classes Flux \n Edge Threshold is " + str(THRESH))
plt.savefig(res_dir + '/figures/north_ch_shortest_path_and_ne_c_flux' + str(THRESH) + '.png', bbox_inches='tight')
print("Total number of nodes shortest path + neighbors: ", north_and_ne.number_of_nodes())
print("Total number of edges: ", north_and_ne.number_of_edges())
print("Total number of nodes shortest path + neighbors and classes: ", north_and_ne_c.number_of_nodes())
print("Total number of edges: ", north_and_ne_c.number_of_edges())
```
# Create Video
```
# choose codec according to format needed
fourcc = cv2.VideoWriter_fourcc(*'mp4v')
video = cv2.VideoWriter(os.path.join(res_dir, "north_pole_ne_c" + str(THRESH) + ".mov"), fourcc, 1, (1200, 800))
for frame in frame_array:
holder = []
for node in north_and_ne_c:
if north_and_ne_c.nodes[node]["frame_num"] == frame:
holder.append(node)
else:
timestamp = None
for node in G:
if G.nodes[node]["frame_num"] == frame:
timestamp = G.nodes[node]["frame_timestamp"]
break
if timestamp is not None:
pickle_file = str(timestamp).replace(':', '-') + ".pkl"
pickle_file = pickle_file.replace(" ", "-")
frame_read = pickle.load(open(os.path.join(res_dir + "/pkl/" + pickle_file), "rb"))
ch_list = frame_read.contour_list
for ch in ch_list:
ch_id = str(frame) + "_" + str(ch.id) + "_" + str(ch.count)
if ch_id not in list(north_and_ne_c.nodes):
ch.color = (169,169,169)
plot_coronal_hole(ch_list, 160, 400, "North Coronal Hole \n Time: " + str(frame_read.timestamp),
filename=os.path.join(res_dir, "frames_north.png"),
plot_rect=False, plot_circle=True, circle_radius=50,
thickness_circle=1, thickness_rect=2, fontscale=0.3,
origin=None, dpi=200)
image_file_name = os.path.join(res_dir, "frames_north.png")
img = cv2.imread(image_file_name)
video.write(img)
cv2.destroyAllWindows()
video.release()
```
# Similar Analysis for the South Pole
```
for node in G.nodes:
if G.nodes[node]["id"] == 2 and G.nodes[node]["frame_num"] == 1205:
print(G.nodes[node])
print(node)
if G.nodes[node]["id"] == 2 and G.nodes[node]["frame_num"] == 1:
print(G.nodes[node])
print(node)
start = "1_2_0"
end = "1205_2_0"
path2 = nx.algorithms.shortest_paths.weighted.dijkstra_path(DiG, source=start, target=end, weight=func)
set_of_classes_south = set()
for node in path2:
#print(str(node) + ", id: " + str(G.nodes[node]["id"]) + ", frame: "+ str(G.nodes[node]["frame_num"]) )
set_of_classes_south.add(G.nodes[node]["id"])
set_of_classes_south
path_g_south = G.subgraph(path2)
unique_id_class_2 = []
for id_class in list(set_of_classes_south):
for node in path_g_south.nodes:
if path_g_south.nodes[node]["id"] == id_class:
unique_id_class_2.append(node)
break
unique_id_class_2
fig, ax = plt.subplots(figsize=(10, 10))
# draw graph, nodes positions are based on their count and frame_num.
# labels are the coronal hole id number.
pos, labels = graph.get_plot_features(sub_graph=path_g_south)
edge_weights = nx.get_edge_attributes(G=path_g_south, name='weight')
edges, weights = zip(*edge_weights.items())
# plot nodes and labels.
nx.draw(path_g_south, pos=pos, font_weight='bold', ax=ax, node_size=700,
node_color=[c.to_rgba(np.array(path_g_south.nodes[ch]["color"]) / 255)
for ch in path_g_south.nodes], edgelist=[])
#nx.draw_networkx_labels(G=path_g, pos=pos, labels=labels, ax=ax, font_size=8)
for ch in unique_id_class_2:
ax.scatter([],[], c=[c.to_rgba(np.array(path_g_south.nodes[ch]["color"]) / 255)], label='ID {}'.format(path_g_south.nodes[ch]["id"]))
nx.draw_networkx_edges(path_g_south, pos=pos, edge_color=weights, edgelist=edges,
edge_cmap=plt.cm.get_cmap('Greys'), edge_vmin=0,
edge_vmax=1, width=3, ax=ax)
nx.draw_networkx_edge_labels(G=path_g_south, pos=pos, edge_labels=edge_weights, ax=ax, alpha=1, font_size=10)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.set_xlim(tuple(sum(i) for i in zip(ax.get_xlim(), (-0.5, 0.5))))
# set y ticks
ax.yaxis.get_major_locator().set_params(integer=True)
ax.set_yticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_yticklabels(timearray[::150], fontsize=12)
ax.tick_params(axis='y', rotation=40)
ax.axis('on')
_ = ax.set_title("Coronal Hole Connectivity South")
_ = plt.gca().legend()
plt.savefig(res_dir + '/figures/south_ch_shortest_path_res_2_to_2.png')
```
# Spatiotemporal Analysis of the South Pole
```
set_of_classes_south
fig, ax = plt.subplots(figsize=(15, 5))
for frame in frame_array:
holder = []
for node in path_g_south:
if path_g_south.nodes[node]["frame_num"] == frame:
holder.append(node)
area = 0
for node in holder:
area += path_g_south.nodes[node]["area"]
#if len(holder) == 0:
# print("no nodes in this frame", frame)
if area != 0:
ax.scatter(frame, area, c=[c.to_rgba(np.array(path_g_south.nodes[holder[0]]["color"]) / 255)])
for ch in unique_id_class_2:
ax.scatter([],[], c=[c.to_rgba(np.array(path_g_south.nodes[ch]["color"]) / 255)], label='ID {}'.format(path_g_south.nodes[ch]["id"]))
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
_ = ax.set_ylabel("Area ($R_{\odot}^2$)")
_ = ax.set_title("South Pole Coronal Hole Shortest Path Spatiotemporal Properties")
plt.savefig(res_dir + '/figures/south_ch_shortest_path_area.png')
net_flux_array = []
abs_flux_array = []
for frame in frame_array:
holder = []
for node in path_g_south:
if path_g_south.nodes[node]["frame_num"] == frame:
holder.append(node)
net_flux = 0
abs_flux = 0
if len(holder) > 0:
# get the node timestamp
timestamp = G.nodes[holder[0]]["frame_timestamp"]
# get pickle file with contour properties.
pickle_file = str(timestamp).replace(':', '-') + ".pkl"
pickle_file = pickle_file.replace(" ", "-")
frame_read = pickle.load(open(os.path.join(res_dir + "/pkl/" + pickle_file), "rb"))
ch_list = frame_read.contour_list
ii = 0
while ii < len(holder):
node = holder[ii]
for ch in ch_list:
if ch.id == path_g_south.nodes[node]["id"] and ch.count == path_g_south.nodes[node]["count"]:
nf, af = br_flux.coronal_flux(db_session, ch, timestamp,
map_dir, window_half_width=window_half_width)
ii +=1
net_flux += nf
abs_flux += af
net_flux_array.append(net_flux)
abs_flux_array.append(abs_flux)
fig, ax = plt.subplots(figsize=(15, 5))
ax.scatter(frame_array, net_flux_array, label="Net Flux")
ax.scatter(frame_array, abs_flux_array, label="Absolute Flux")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
plt.legend()
# label axis
_ = ax.set_ylabel("$\Phi_{B}$")
_ = ax.set_title("South Pole Coronal Hole Shortest Path Flux")
plt.savefig(res_dir + '/figures/south_ch_shortest_path_net_flux.png')
```
# South Shortest Path and *Neighbors*
```
THRESH = 0.2
south_and_ne = path_g_south.copy()
# add all nodes that are adjacent to nodes in shortest path
for node in path_g_south:
for adj_node in G[node]:
# if the edge weight is greater than threshold, then add the neighbor node.
if G[node][adj_node]['weight'] >= THRESH:
# check if the node already exists in shortest path
if adj_node not in south_and_ne.nodes:
# add the node.
south_and_ne.add_node(str(adj_node),
area=G.nodes[adj_node]["area"],
id=G.nodes[adj_node]["id"],
frame_num=G.nodes[adj_node]["frame_num"],
frame_timestamp=G.nodes[adj_node]["frame_timestamp"],
count=G.nodes[adj_node]["count"],
color=G.nodes[adj_node]["color"])
for u,v,w in G.edges.data():
# if the two nodes exist in the shorstest path and neighbors then add the edge.
if u in south_and_ne and v in south_and_ne:
if not south_and_ne.has_edge(u, v):
south_and_ne.add_edge(u, v, weight=w)
ne_classes = set()
for node in south_and_ne:
ne_classes.add(G.nodes[node]["id"])
str(ne_classes)
fig, ax = plt.subplots(figsize=(15, 5))
for frame in frame_array:
holder = []
for node in south_and_ne:
if south_and_ne.nodes[node]["frame_num"] == frame:
holder.append(node)
area = 0
for node in holder:
area += south_and_ne.nodes[node]["area"]
#if len(holder) == 0:
# print("no nodes in this frame", frame)
if area != 0:
ax.scatter(frame, area, c=['r' if len(holder) > 1 else 'b'])
ax.scatter([], [], c="r", label="npf $>$ 1")
ax.scatter([], [], c="b", label="npf $\leq$ 1")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
_ = ax.set_ylabel("Area ($R_{\odot}^2$)")
#_ = ax.set_title("South Pole Coronal Hole Shortest Path and Neighbors Spatiotemporal Properties \n Edge Threshold is " + str(THRESH) + "\n " + str(ne_classes))
_ = ax.set_title("South Pole Coronal Hole Shortest Path and Neighbors Spatiotemporal Properties \n Edge Threshold is " + str(THRESH) + "\n ")
_ = plt.legend()
plt.savefig(res_dir + '/figures/south_ch_shortest_path_and_ne_area' + str(THRESH) + '.png', bbox_inches='tight')
```
## shortest path + neighbors classes
```
south_and_ne_c = south_and_ne.copy()
# add all nodes that are adjacent to nodes in shortest path
for node in G:
if node not in south_and_ne_c.nodes and G.nodes[node]["id"] in ne_classes:
# add the node.
south_and_ne_c.add_node(str(node),
area=G.nodes[node]["area"],
id=G.nodes[node]["id"],
frame_num=G.nodes[node]["frame_num"],
frame_timestamp=G.nodes[node]["frame_timestamp"],
count=G.nodes[node]["count"],
color=G.nodes[node]["color"])
for u,v,w in G.edges.data():
# if the two nodes exist in the shorstest path and neighbors then add the edge.
if u in south_and_ne_c and v in south_and_ne_c:
if not south_and_ne_c.has_edge(u, v):
south_and_ne_c.add_edge(u, v, weight=w)
fig, ax = plt.subplots(figsize=(15, 5))
for frame in frame_array:
holder = []
for node in south_and_ne_c:
if south_and_ne_c.nodes[node]["frame_num"] == frame:
holder.append(node)
area = 0
for node in holder:
area += south_and_ne_c.nodes[node]["area"]
#if len(holder) == 0:
# print("no nodes in this frame", frame)
if area != 0:
ax.scatter(frame, area, c=['r' if len(holder) > 1 else 'b'])
ax.scatter([], [], c="r", label="npf $>$ 1")
ax.scatter([], [], c="b", label="npf $\leq$ 1")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
_ = ax.set_ylabel("Area ($R_{\odot}^2$)")
#_ = ax.set_title("South Pole Coronal Hole Shortest Path and Neighbors Classes Spatiotemporal Properties \n Edge Threshold is " + str(THRESH) + "\n " + str(ne_classes))
_ = ax.set_title("South Pole Coronal Hole Shortest Path and Neighbors Classes Spatiotemporal Properties \n Edge Threshold is " + str(THRESH))
_ = plt.legend()
plt.savefig(res_dir + '/figures/south_ch_shortest_path_and_ne_c_area' + str(THRESH) + '.png', bbox_inches='tight')
print("Total number of nodes shortest path + neighbors: ", south_and_ne.number_of_nodes())
print("Total number of edges: ", south_and_ne.number_of_edges())
print("Total number of nodes shortest path + neighbors and classes: ", south_and_ne_c.number_of_nodes())
print("Total number of edges: ", south_and_ne_c.number_of_edges())
net_flux_array = []
abs_flux_array = []
for frame in frame_array:
holder = []
for node in south_and_ne_c:
if south_and_ne_c.nodes[node]["frame_num"] == frame:
holder.append(node)
net_flux = 0
abs_flux = 0
if len(holder) > 0:
# get the node timestamp
timestamp = G.nodes[holder[0]]["frame_timestamp"]
# get pickle file with contour properties.
pickle_file = str(timestamp).replace(':', '-') + ".pkl"
pickle_file = pickle_file.replace(" ", "-")
frame_read = pickle.load(open(os.path.join(res_dir + "/pkl/" + pickle_file), "rb"))
ch_list = frame_read.contour_list
ii = 0
while ii < len(holder):
node = holder[ii]
for ch in ch_list:
if ch.id == south_and_ne_c.nodes[node]["id"] and ch.count == south_and_ne_c.nodes[node]["count"]:
nf, af = br_flux.coronal_flux(db_session, ch, timestamp,
map_dir, window_half_width=window_half_width)
ii +=1
net_flux += nf
abs_flux += af
net_flux_array.append(net_flux)
abs_flux_array.append(abs_flux)
fig, ax = plt.subplots(figsize=(15, 5))
ax.scatter(frame_array, net_flux_array, label="Net Flux")
ax.scatter(frame_array, abs_flux_array, label="Absolute Flux")
# set x ticks to be timestamps
ax.set_xticks(np.linspace(0, len(timearray), len(timearray))[::150])
ax.set_xticklabels(timearray[::150])
ax.tick_params(axis='x', rotation=15)
plt.legend()
# label axis
_ = ax.set_ylabel("$\Phi_{B}$")
_ = ax.set_title("South Pole Coronal Hole Shortest Path and Neighbors Classes Flux \n Edge Threshold is " + str(THRESH))
plt.savefig(res_dir + '/figures/south_ch_shortest_path_and_ne_c_flux' + str(THRESH) + '.png', bbox_inches='tight')
```
# Create Video
```
# choose codec according to format needed
fourcc = cv2.VideoWriter_fourcc(*'mp4v')
video = cv2.VideoWriter(os.path.join(res_dir, "south_pole_ne_c" + str(THRESH) + ".mov"),
fourcc, 1, (1200, 800))
for frame in frame_array:
holder = []
for node in south_and_ne_c:
if south_and_ne_c.nodes[node]["frame_num"] == frame:
holder.append(node)
else:
timestamp = None
for node in G:
if G.nodes[node]["frame_num"] == frame:
timestamp = G.nodes[node]["frame_timestamp"]
break
if timestamp is not None:
pickle_file = str(timestamp).replace(':', '-') + ".pkl"
pickle_file = pickle_file.replace(" ", "-")
frame_read = pickle.load(open(os.path.join(res_dir + "/pkl/" + pickle_file), "rb"))
ch_list = frame_read.contour_list
for ch in ch_list:
ch_id = str(frame) + "_" + str(ch.id) + "_" + str(ch.count)
if ch_id not in list(south_and_ne_c.nodes):
ch.color = (169,169,169)
plot_coronal_hole(ch_list, 160, 400, "South Coronal Hole \n Time: " + str(frame_read.timestamp),
filename=os.path.join(res_dir, "frames_south.png"),
plot_rect=False, plot_circle=True, circle_radius=50,
thickness_circle=1, thickness_rect=2, fontscale=0.3,
origin=None)
image_file_name = os.path.join(res_dir, "frames_south.png")
img = cv2.imread(image_file_name)
video.write(img)
cv2.destroyAllWindows()
video.release()
```
| github_jupyter |
```
import gym
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.distributions import Normal
import matplotlib.pyplot as plt
from torch.utils.tensorboard import SummaryWriter
import time
import os
%matplotlib inline
device = torch.device("cuda:1" if torch.cuda.is_available() else "cpu")
import sys
sys.path.append("../utils/")
from replay_buffer import ReplayBuffer
from normalize_action import NormalizeActions
from noise import OUNoise
class ValueNetwork(nn.Module):
def __init__(self, state_dim, action_dim):
super(ValueNetwork, self).__init__()
self.linear_state = nn.Linear(state_dim, 64)
self.linear_action = nn.Linear(action_dim, 64)
self.linear2 = nn.Linear(128, 32)
self.linear3 = nn.Linear(32, 1)
def forward(self, state, action):
hidden_state = F.relu(self.linear_state(state))
hidden_action = F.relu(self.linear_action(action))
cat_state_action = torch.cat((hidden_action, hidden_state),dim=1)
hidden2 = F.relu(self.linear2(cat_state_action))
Q = self.linear3(hidden2)
return Q
class PolicyNetwork(nn.Module):
def __init__(self, in_dim, out_dim):
super(PolicyNetwork, self).__init__()
self.linear1 = nn.Linear(in_dim, 128)
self.linear2 = nn.Linear(128, 64)
self.linear3 = nn.Linear(64, out_dim) # (256, 1)
def forward(self, state):
x = F.relu(self.linear1(state))
x = F.relu(self.linear2(x))
x = torch.tanh(self.linear3(x))
return x
def get_action(self, state):
state = torch.tensor(state,dtype=torch.float).unsqueeze(0).to(device)
action = self.forward(state)
return action.detach().cpu().numpy()
class DDPG:
def __init__(self, ):
def ddpg_train(batch_size, gamma=0.99, soft_tau=1e-2):
samples = replay_buffer.sample()
state, action, next_state = samples['current_state'], samples['action'], samples['next_state']
reward, done = samples['reward'], samples['done']
target_value = reward + (1.0-done)*gamma*target_value_net(next_state, target_policy_net(next_state))
value = value_net(state, action)
value_loss = ((value - target_value.detach()).pow(2)).mean()
policy_loss = -value_net(state, policy_net(state)).mean()
value_optimizer.zero_grad()
value_loss.backward()
value_optimizer.step()
policy_optimizer.zero_grad()
policy_loss.backward()
policy_optimizer.step()
for target_param, param in zip(target_value_net.parameters(), value_net.parameters()):
target_param.data.copy_(target_param.data*(1.0-soft_tau) + param.data*soft_tau)
for target_param, param in zip(target_policy_net.parameters(), policy_net.parameters()):
target_param.data.copy_(target_param.data*(1.0-soft_tau) + param.data*soft_tau)
return value_loss.item(), policy_loss.item()
# env = NormalizeActions(gym.make("Pendulum-v0"))
env = NormalizeActions(gym.make("MountainCarContinuous-v0"))
ou_noise = OUNoise(env.action_space)
in_dim = env.observation_space.shape[0] # 3
out_dim = env.action_space.shape[0] # 1 连续动作空间
value_net = ValueNetwork(in_dim, out_dim).to(device)
policy_net = PolicyNetwork(in_dim, out_dim).to(device)
target_value_net = ValueNetwork(in_dim, out_dim).to(device)
target_policy_net = PolicyNetwork(in_dim, out_dim).to(device)
target_value_net.load_state_dict(value_net.state_dict())
target_policy_net.load_state_dict(policy_net.state_dict())
value_optimizer = optim.Adam(value_net.parameters())
policy_optimizer = optim.Adam(policy_net.parameters(), lr=1e-4)
train_episodes = 250
train_steps = 1000
test_episodes = int(train_episodes / 2)
test_steps = 100
buffer_size = 1000000
batch_size = 128
replay_buffer = ReplayBuffer(in_dim, batch_size, buffer_size)
test = True
def smooth_plot(factor, item, plot_decay):
item_x = np.arange(len(item))
item_smooth = [np.mean(item[i:i+factor]) if i > factor else np.mean(item[0:i+1])
for i in range(len(item))]
for i in range(len(item)// plot_decay):
item_x = item_x[::2]
item_smooth = item_smooth[::2]
return item_x, item_smooth
def plot(episode, rewards, value_losses, policy_losses, noise):
clear_output(True)
rewards_x, rewards_smooth = smooth_plot(10, rewards, 500)
value_losses_x, value_losses_smooth = smooth_plot(10, value_losses, 10000)
policy_losses_x, policy_losses_smooth = smooth_plot(10, policy_losses, 10000)
noise_x, noise_smooth = smooth_plot(10, noise, 100)
plt.figure(figsize=(18, 12))
plt.subplot(411)
plt.title('episode %s. reward: %s'%(episode, rewards_smooth[-1]))
plt.plot(rewards, label="Rewards", color='lightsteelblue', linewidth='1')
plt.plot(rewards_x, rewards_smooth, label='Smothed_Rewards', color='darkorange', linewidth='3')
plt.legend(loc='best')
plt.subplot(412)
plt.title('Value_Losses')
plt.plot(value_losses,label="Value_Losses",color='lightsteelblue',linewidth='1')
plt.plot(value_losses_x, value_losses_smooth,
label="Smoothed_Value_Losses",color='darkorange',linewidth='3')
plt.legend(loc='best')
plt.subplot(413)
plt.title('Policy_Losses')
plt.plot(policy_losses,label="Policy_Losses",color='lightsteelblue',linewidth='1')
plt.plot(policy_losses_x, policy_losses_smooth,
label="Smoothed_Policy_Losses",color='darkorange',linewidth='3')
plt.legend(loc='best')
plt.subplot(414)
plt.title('Noise')
plt.plot(noise,label="Noise",color='lightsteelblue',linewidth='1')
plt.plot(noise_x, noise_smooth,
label="Smoothed_Noise",color='darkorange',linewidth='3')
plt.legend(loc='best')
plt.show()
# value_losses = []
# policy_losses = []
# all_rewards = []
# updates = 0
# test = True
# for episode in range(train_episodes):
# state = env.reset()
# ou_noise.reset()
# episode_reward = 0
# noises = []
# for step in range(train_steps):
# action1 = policy_net.get_action(state)
# # action = ou_noise.get_action(action1, step)
# # noises.append(action[0][0]-action1[0][0])
# # 200 update in 10
# if step % 200 == 0:
# test = not test
# noise = abs(np.random.randn(1)) if test else -abs(np.random.randn(1))
# action = action1 + noise
# noises.append(noise)
# next_state, reward, done, _ = env.step(action.flatten())
# replay_buffer.store(state, action, next_state.flatten(), reward, done)
# if len(replay_buffer) > batch_size :
# value_loss, policy_loss = ddpg_train(batch_size)
# value_losses.append(value_loss)
# policy_losses.append(policy_loss)
# state = next_state
# episode_reward += reward
# if done:
# break
# updates += 1
# all_rewards.append(episode_reward)
# plot(episode, all_rewards, value_losses, policy_losses, noises[:200])
def run_ddpg(time, writer, update_step, noise_discount):
test = True
for episode in range(train_episodes):
state = env.reset()
episode_reward = 0
for step in range(train_steps):
action1 = policy_net.get_action(state)
if step % update_step == 0:
test = not test
noise_sample = abs(np.random.randn(1)) * noise_discount
noise = noise_sample if test else -noise_sample
action = action1 + noise
next_state, reward, done, _ = env.step(action.flatten())
replay_buffer.store(state, action, next_state.flatten(), reward, done)
if len(replay_buffer) > batch_size :
value_loss, policy_loss = ddpg_train(batch_size)
state = next_state
episode_reward += reward
if done:
break
writer.add_scalars("train_reward/update_step_{}".format(update_step),
{"noise_discount_{}".format(noise_discount):episode_reward}, episode)
torch.save(policy_net.state_dict(), "./test/Continue_time_{}/model/update_step_{}_noise_discount_{}.pth".format(time, update_step, noise_discount))
print("Train < update_step : {}, noise_discount : {} > finished !".format(update_step, noise_discount))
def test_ddpg(time, writer, update_step, noise_discount):
policy_net_1 = PolicyNetwork(in_dim, out_dim).to(device)
policy_net_1.load_state_dict(torch.load("./test/Continue_time_{}/model/update_step_{}_noise_discount_{}.pth".format(time, update_step, noise_discount)))
for test_episode in range(test_episodes):
state = env.reset()
rewards = 0
for _ in range(test_steps):
action = policy_net_1.get_action(state.flatten())
next_state, reward, done, info = env.step(action)
state = next_state
rewards += reward
if done: break
writer.add_scalars("test_reward/update_step_{}".format(update_step),
{"noise_discount_{}".format(noise_discount):rewards}, test_episode)
print("Test < update_step : {}, noise_discount : {} > finished !".format(update_step, noise_discount))
# torch.save(policy_net.state_dict(), "./model/DDPG_for_mountain_car.pth")
# policy_net_1 = PolicyNetwork(in_dim, out_dim).to(device)
# policy_net_1.load_state_dict(torch.load("./model/DDPG_for_mountain_car.pth"))
# policy_net_1.eval()
# import pdb
# import gym
# from IPython import display
# import matplotlib
# import matplotlib.pyplot as plt
# %matplotlib inline
# env = gym.make("MountainCarContinuous-v0")
# state = env.reset()
# img = plt.imshow(env.render(mode='rgb_array')) # only call this once
# for _ in range(1000):
# img.set_data(env.render(mode='rgb_array')) # just update the data
# display.display(plt.gcf())
# display.clear_output(wait=True)
# policy_net = policy_net.cpu()
# action = policy_net(torch.FloatTensor(state)).detach().numpy()
# # action = env.action_space.sample()
# next_state, _, done, _ = env.step(action)
# if done:
# state = env.reset()
# state = next_state
# from gym import wrappers
# env = gym.make("MountainCarContinuous-v0")
# env = wrappers.Monitor(env, "./gym-results/DDPG_mountaincar/", force=True)
```
## Test Results
#### Plot use tensorboard
```
time = time.strftime("%Y-%m-%d_%H:%M:%S", time.localtime())
writer = SummaryWriter(log_dir="test/Continue_time_{}/tensorboard/".format(time))
dirName = "./test/Continue_time_{}/model".format(time)
if not os.path.exists(dirName):
os.mkdir(dirName)
update_steps = [50, 100, 150, 200, 250, 300]
noise_discounts = [0.1, 0.5, 1.0, 1.5, 2.0, 3.0]
for update_step in update_steps:
for noise_discount in noise_discounts:
# train
run_ddpg(time, writer, update_step, noise_discount)
# test
test_ddpg(time, writer, update_step, noise_discount)
```
#### Plot use seaborn and matplotlib
```
# import pandas as pd
# train_reward = np.array([])
# test_reward = np.array([])
# for epoch in range(5):
# # train
# rewards = run_ddpg(epoch)
# train_reward = np.concatenate((train_reward, rewards))
# # test
# rewards = test_ddpg(epoch)
# test_reward = np.concatenate((test_reward, rewards))
# train_x = lambda : np.arange(1, train_episodes+1)
# train_list = np.stack((train_x() for _ in range(5)), axis=0).flatten()
# test_x = lambda : np.arange(1, test_episodes+1)
# test_list = np.stack((test_x() for _ in range(5)), axis=0).flatten()
# train_data = pd.DataFrame(dict(x=train_list, y=train_reward))
# test_data = pd.DataFrame(dict(x=test_list, y=test_reward))
# train_data.to_csv("./test/DDPG_OUnoise/trian_data.csv", index=False)
# test_data.to_csv("./test/DDPG_OUnoise/test_data.csv", index=False)
# import seaborn as sns; sns.set()
# import matplotlib.pyplot as plt
# import pandas as pd
# DDPG_NO_test_data = pd.read_csv("RL_notes_and_codes/algorithm_implement/test/DDPG_Normalnoise/test_data.csv")
# DDPG_NO_train_data = pd.read_csv("RL_notes_and_codes/algorithm_implement/test/DDPG_Normalnoise/trian_data.csv")
# DDPG_OU_test_data = pd.read_csv("RL_notes_and_codes/algorithm_implement/test/DDPG_OUnoise/test_data.csv")
# DDPG_OU_train_data = pd.read_csv("RL_notes_and_codes/algorithm_implement/test/DDPG_OUnoise/trian_data.csv")
# DDPG_NO_test_data["diff"] = "Normalnoise"
# DDPG_NO_train_data["diff"] = "Normalnoise"
# DDPG_OU_test_data["diff"] = "OUnoise"
# DDPG_OU_train_data["diff"] = "OUnoise"
# DDPG_test_data = pd.concat((DDPG_NO_test_data, DDPG_OU_test_data))
# DDPG_train_data = pd.concat((DDPG_NO_train_data, DDPG_OU_train_data))
# plt.figure(figsize=(16, 8))
# plt.subplot(211)
# ax = sns.lineplot(x="x", y="y", hue="diff", data=DDPG_test_data)
# plt.title("Test 5 times Reward of 100 steps for each episode, Avg: Normalnoise {}, OUnoise {}"
# .format(round(DDPG_test_data[DDPG_test_data['diff']=="Normalnoise"]['y'].mean(), 3),
# round(DDPG_test_data[DDPG_test_data['diff']=="OUnoise"]['y'].mean(), 3)))
# plt.xlabel("episodes")
# plt.ylabel("rewards")
# plt.subplot(212)
# ax = sns.lineplot(x="x", y="y", hue="diff", data=DDPG_train_data)
# plt.title("Train 5 times Reward of 1000000 steps for each episode, Avg: Normalnoise {}, OUnoise {}"
# .format(round(DDPG_train_data[DDPG_train_data['diff']=="Normalnoise"]['y'].mean(), 3),
# round(DDPG_train_data[DDPG_train_data['diff']=="OUnoise"]['y'].mean(), 3)))
# plt.xlabel("episodes")
# plt.ylabel("rewards")
# plt.savefig("RL_notes_and_codes/algorithm_implement/test/Noise_test.png")
# plt.show()
```

| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import os
os.chdir('drive/MyDrive/Dataset')
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.metrics import classification_report
def makedata(X):
for i in range(X.shape[0]):
if(X[i,1] == 0):
X[i,0] = 0
for j in range(7):
X[i,j+1] = 0
X = np.array(X)
X = np.average(X, axis = 0)
X = np.array(X.reshape(1, -1))
return X
dev = np.array(pd.read_csv('/content/drive/My Drive/Dataset/dev_split_Depression_AVEC2017.csv',delimiter=',',encoding='utf-8'))[:, 0:2]
test = np.array(pd.read_csv('/content/drive/My Drive/Dataset/full_test_split.csv',delimiter=',',encoding='utf-8'))[:, 0:2]
train = np.array(pd.read_csv('/content/drive/My Drive/Dataset/train_split_Depression_AVEC2017.csv',delimiter=',',encoding='utf-8'))[:, 0:2]
X_train = np.zeros((1,74))
Y_train = []
X_test = np.zeros((1,74))
Y_test = []
for i in range(len(test)):
data = pd.read_csv('/content/drive/My Drive/Dataset/test_data/'+str(int(test[i][0]))+"_COVAREP.csv",header = None)
X_temp = data.iloc[:,:].values
X_temp = makedata(X_temp)
X_test = np.concatenate((X_test,X_temp),0)
Y_test.append(test[i][1])
X_test = np.delete(X_test,0,0)
Y_test = np.array(Y_test)
count_0 = 0
count_1 = 0
for i in range(len(train)):
data = pd.read_csv('/content/drive/My Drive/Dataset/train_data/'+str(int(train[i][0]))+"_COVAREP.csv",header = None)
if(train[i][1] == 0):
count_0 +=1
if(count_0<27):
X_temp = data.iloc[:,:].values
X_temp = makedata(X_temp)
X_train = np.concatenate((X_train,X_temp),0)
Y_train.append(train[i][1])
else:
count_1 +=1
if(count_1<27):
X_temp = data.iloc[:,:].values
X_temp = makedata(X_temp)
X_train = np.concatenate((X_train,X_temp),0)
Y_train.append(train[i][1])
count_0 = 0
count_1 = 0
for i in range(len(dev)):
data = pd.read_csv('/content/drive/My Drive/Dataset/dev_data/'+str(int(dev[i][0]))+"_COVAREP.csv",header = None)
if(dev[i][1] == 0):
count_0 += 1
if(count_0<27):
X_temp = data.iloc[:,:].values
X_temp = makedata(X_temp)
X_train = np.concatenate((X_train,X_temp),0)
Y_train.append(dev[i][1])
else:
count_1+= 1
if(count_1<27):
X_temp = data.iloc[:,:].values
X_temp = makedata(X_temp)
X_train = np.concatenate((X_train,X_temp),0)
Y_train.append(dev[i][1])
X_train = np.delete(X_train,0,0)
Y_train = np.array(Y_train)
Y_test = np.delete(Y_test,15)
rfModel = RandomForestClassifier()
rfModel.fit(X_train, Y_train)
Y_pred1 = rfModel.predict(X_test)
print("Random forest: ")
print(classification_report(Y_test, Y_pred1))
svmModel = SVC(kernel="rbf",random_state=0)
svmModel.fit(X_train, Y_train)
Y_pred2 = svmModel.predict(X_test)
print("SVM: ")
print(classification_report(Y_test, Y_pred2))
```
| github_jupyter |
## Camvid segmentation
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
#export
from fastai import *
from fastai.vision import *
path = Path('data/camvid')
path.ls()
path_lbl = path/'labels'
path_img = path/'images'
```
## Data
```
fnames = get_image_files(path_img)
fnames[:5]
path_lbl.ls()[:5]
img_f = fnames[0]
img = open_image(img_f)
img.show(figsize=(5,5))
codes = np.loadtxt(path/'codes.txt', dtype=str)
codes
def get_y_fn(fn): return path_lbl/f'{fn.name[:-4]}_P.png'
mask = open_mask(get_y_fn(img_f))
mask.show(figsize=(5,5), alpha=1)
mask.data
```
## Datasets
```
valid_fns = np.loadtxt(path/'valid.txt', dtype=str)
valid_fns[:5]
valid_fns = [path_img/o for o in valid_fns]
train_fns = list(set(fnames)-set(valid_fns))
y_train_fns = [get_y_fn(o) for o in train_fns]
y_valid_fns = [get_y_fn(o) for o in valid_fns]
len(train_fns),len(valid_fns),len(y_train_fns),len(y_valid_fns)
size=128
bs=32
train_ds = SegmentationDataset(train_fns, y_train_fns, classes=codes)
valid_ds = SegmentationDataset(valid_fns, y_valid_fns, classes=codes)
train_tfms,valid_tfms = get_transforms()
train_tds = DatasetTfm(train_ds, train_tfms, size=size, tfm_y=True)
valid_tds = DatasetTfm(valid_ds, valid_tfms, size=size, tfm_y=True)
data = DataBunch.create(train_tds, valid_tds, bs=bs)
```
## Refactor
- valid set
- path
- fnames
- idxs
- min idx
- rand pct
- type of data
- source of labels
- get filenames
- get labels
- split data
- make datasets
- get tfms
- (make dls)
- (use device)
- databunch
```
#export
class ItemList():
"A collection of items with `__len__` and `__getitem__` with `ndarray` indexing semantics"
def __init__(self, items:Iterator): self.items = np.array(list(items))
def __len__(self)->int: return len(self.items)
def __getitem__(self,i:int)->Any: self.items[i]
def __repr__(self)->str: return f'{self.__class__.__name__} ({len(self)} items)\n{self.items}'
class PathItemList(ItemList):
def __init__(self, items:Iterator, path:PathOrStr='.'):
super().__init__(items)
self.path = Path(path)
def __repr__(self)->str: return f'{super().__repr__()}\nPath: {self.path}'
def join_path(fname:PathOrStr, path:PathOrStr='.')->Path:
"`Path(path)/Path(fname)`, `path` defaults to current dir"
return Path(path)/Path(fname)
def join_paths(fnames:FilePathList, path:PathOrStr='.')->Collection[Path]:
path = Path(path)
return [join_path(o,path) for o in fnames]
def loadtxt_str(path:PathOrStr)->np.ndarray:
"Return `ndarray` of `str` of lines of text from `path`"
return np.loadtxt(str(path), str)
#export
def _df_to_fns_labels(df:pd.DataFrame, fn_col:int=0, label_col:int=1,
label_delim:str=None, suffix:Optional[str]=None):
"""Get image file names in `fn_col` by adding `suffix` and labels in `label_col` from `df`.
If `label_delim` is specified, splits the values in `label_col` accordingly.
"""
if label_delim:
df.iloc[:,label_col] = list(csv.reader(df.iloc[:,label_col], delimiter=label_delim))
labels = df.iloc[:,label_col].values
fnames = df.iloc[:,fn_col].str.lstrip()
if suffix: fnames = fnames + suffix
return fnames.values, labels
#export
class ImageFileList(PathItemList):
@classmethod
def from_folder(cls, path:PathOrStr='.', check_ext:bool=True, recurse=True)->'ImageFileList':
return cls(get_image_files(path, check_ext=check_ext, recurse=recurse), path)
def label_from_func(self, func:Callable)->Collection:
return LabelList([(o,func(o)) for o in self.items], self.path)
def label_from_re(self, pat:str, full_path:bool=False)->Collection:
pat = re.compile(pat)
def _inner(o):
s = str(o if full_path else o.name)
res = pat.search(s)
assert res,f'Failed to find "{pat}" in "{s}"'
return res.group(1)
return self.label_from_func(_inner)
def label_from_df(self, df, fn_col:int=0, label_col:int=1, sep:str=None, folder:PathOrStr='.',
suffix:str=None)->Collection:
fnames, labels = _df_to_fns_labels(df, fn_col, label_col, sep, suffix)
fnames = join_paths(fnames, self.path/Path(folder))
return LabelList([(fn, np.array(lbl, dtype=np.object)) for fn, lbl in zip(fnames, labels) if fn in self.items],
self.path)
def label_from_csv(self, csv_fname, header:Optional[Union[int,str]]='infer', fn_col:int=0, label_col:int=1,
sep:str=None, folder:PathOrStr='.', suffix:str=None)->Collection:
df = pd.read_csv(self.path/csv_fname, header=header)
return self.label_from_df(df, fn_col, label_col, sep, folder, suffix)
def label_from_folder(self, classes:Collection[str]=None)->Collection:
labels = [fn.parent.parts[-1] for fn in self.items]
if classes is None: classes = uniqueify(labels)
return LabelList([(o,lbl) for o, lbl in zip(self.items, labels) if lbl in classes], self.path)
#export
class LabelList(PathItemList):
@property
def files(self): return self.items[:,0]
def split_by_files(self, valid_fnames:FilePathList)->'SplitData':
valid = [o for o in self.items if o[0] in valid_fnames]
train = [o for o in self.items if o[0] not in valid_fnames]
return SplitData(self.path, LabelList(train), LabelList(valid))
def split_by_fname_file(self, fname:PathOrStr, path:PathOrStr=None)->'SplitData':
path = Path(ifnone(path, self.path))
fnames = join_paths(loadtxt_str(self.path/fname), path)
return self.split_by_files(fnames)
def split_by_idx(self, valid_idx:Collection[int])->'SplitData':
valid = [o for i,o in enumerate(self.items) if i in valid_idx]
train = [o for i,o in enumerate(self.items) if i not in valid_idx]
return SplitData(self.path, LabelList(train), LabelList(valid))
def split_by_folder(self, train:str='train', valid:str='valid')->'SplitData':
n = len(self.path.parts)
folder_name = [o[0].parent.parts[n] for o in self.items]
valid = [o for o in self.items if o[0].parent.parts[n] == valid]
train = [o for o in self.items if o[0].parent.parts[n] == train]
return SplitData(self.path, LabelList(train), LabelList(valid))
def random_split_by_pct(self, valid_pct:float=0.2)->'SplitData':
rand_idx = np.random.permutation(range(len(self.items)))
cut = int(valid_pct * len(self.items))
return self.split_by_idx(rand_idx[:cut])
#export
@dataclass
class SplitData():
path:PathOrStr
train:LabelList
valid:LabelList
test: LabelList=None
def __post_init__(self): self.path = Path(self.path)
@property
def lists(self):
res = [self.train,self.valid]
if self.test is not None: res.append(self.test)
return res
def datasets(self, dataset_cls:type, **kwargs)->'SplitDatasets':
"Create datasets from the underlying data using `dataset_cls` and passing along the `kwargs`."
train = dataset_cls(*self.train.items.T, **kwargs)
dss = [train]
dss += [train.new(*o.items.T, **kwargs) for o in self.lists[1:]]
cls = getattr(train, '__splits_class__', SplitDatasets)
return cls(self.path, *dss)
#export
@dataclass
class SplitDatasets():
path:PathOrStr
train_ds:Dataset
valid_ds:Dataset
test_ds:Optional[Dataset] = None
@property
def datasets(self): return [self.train_ds,self.valid_ds]
def transform(self, tfms:TfmList, **kwargs)->'SplitDatasets':
assert not isinstance(self.train_ds, DatasetTfm)
self.train_ds = DatasetTfm(self.train_ds, tfms[0], **kwargs)
self.valid_ds = DatasetTfm(self.valid_ds, tfms[1], **kwargs)
if self.test_ds is not None:
self.test_ds = DatasetTfm(self.test_ds, tfms[1], **kwargs)
return self
def dataloaders(self, **kwargs):
return [DataLoader(o, **kwargs) for o in self.datasets]
def databunch(self, path=None, **kwargs):
path = Path(ifnone(path, self.path))
return ImageDataBunch.create(*self.datasets, path=path, **kwargs)
tfms = get_transforms()
ifl = ImageFileList.from_folder(path_img); ifl
ll = ifl.label_from_func(get_y_fn); ll
sd = ll.split_by_fname_file('../valid.txt')
tfms = get_transforms()
dss = sd.datasets(SegmentationDataset, classes=codes)
dss.train_ds.classes
tdss = dss.transform(tfms, size=128, tfm_y=True)
data = tdss.databunch()
data = (ImageFileList.from_folder(path_img)
.label_from_func(get_y_fn)
.split_by_fname_file('../valid.txt')
.datasets(SegmentationDataset, classes=codes)
.transform(tfms, size=128, tfm_y=True)
.databunch())
x,y = data.train_dl.one_batch()
show_xy_images(x,y,rows=3)
```
| github_jupyter |
# Viewing Molecules
[nglview](http://nglviewer.org/nglview/latest/api.html) is an extremely powerful and capable 3D molecule view that runs within a web browser. It supports complex visualisations of molecules from a range of file formats, and can even be used to view trajectories. It provides a full framework for building 3D molecular visualisation into your Jupyter notebooks or websites.
While nglview is very powerful, that power and flexibility can be a little daunting for newcomers. [BioSimSpace](https://biosimspace.org) is a project that provides easy-to-use wrappers around common molecular simulation tasks. One such task is viewing molecules. BioSimSpace provides the function `viewMolecules` that uses [nglview](http://nglviewer.org/nglview/latest/api.html) to do exactly that :-)
```
from BioSimSpace import viewMolecules
v = viewMolecules("data/dioxin.pdb")
```
The above code has use the molecule file parsers built into BioSimSpace to load the molecule contained in `dioxin.pdb`. This is then rendered using nglview. The above nglview interface allows you to rotate the molecule (left click and drag), zoom in and out (pinch or scroll up or down) and translate (right click and drag, or control+click on a Mac).
Try moving and rotating the molecule. If you lose the molecule, click the "Center" button in the General tab to recenter the molecule.

The BioSimSpace `viewMolecules` function has done two things:
* it first loaded the molecule(s) from the file,
* and it then rendered them
Loading molecules can take a long time and use a lot of memory. To prevent you from having to repeatedly load molecules, the `viewMolecules` function has returned a view object that can be re-used. To see how to use it, use python's help...
```
help(v)
```
As you can see, we can use `v.system()` to view all of the loaded molecules again, without having to reload them.
```
v.system()
```
You can change the representation of the molecule by clicking on the "Representation" tab. First click the "Remove" icon to remove the current representation. Then click the drop-down representation box to choose another representation (e.g. "spacefill"). Then click the "Add" icon to add that representation. Experiment with adding and removing different representations.

# Loading lots of molecules
nglview isn't just limited to viewing small molecules. It also works really well as a viewer for large molecular systems. It (sometimes) is sufficiently clever to select appropriate representations for the molecules being loaded.
For example, view the protein-ligand complex in `data/complex.pdb`
```
v = viewMolecules("data/complex.pdb")
```
In this case, nglview has automatically selected a cartoon representation for the protein and a ball and stick representation for the ligand.
You can achieve this yourself by using selections to set different representation for different molecules (or parts of molecules). First, delete the default representations by repeatedly clicking the "Remove" button in the representations tab. Once you have removed all of them, we will add a new representation. Select the type as surface, and then type "protein" into the selection box (which starts off with a "*" in it).

Click "Add". After some time thinking, nglview will show you a surface representation of the protein.
Next, add a "spacefill" representation to the ligand. The ligand residue is called "LIG", so to do this, select "spacefill", type "LIG" into the selection box, and then click add. You should now see the ligand neatly bound into the protein.

The selection box can be used to select proteins ("protein"), water ("water"), everything ("*") or residues by name (e.g. "LIG") or number (e.g. "35"). Play around creating different selections and representations. For example, create a "point" representation for water, a "tube" representation of the protein and a "licorice" representation of all alanine residues. Note - you can control the opacity (level of transparency) of a representation by selecting the representation in the drop down box and changing the "opacity" slider in the "Parameters" tab - see below. You can also change things like the colour scheme of the representation in this "Parameters" tab

# Viewing individual molecules
The view object returned by BioSimSpace can be used to view specific molecules from the file. To do this, use the `molecules` function. This takes a list of indicies of the molecules you want to view. For example, to view the first molecule (molecule at index 0) type;
```
v.molecules([0])
```
while to view molecules 100-1000 use the below code (noting that you may need to add a "ball and stick" represntation in case nglview automatically hides the water molecules).
```
v.molecules( range(100,1000) )
```
# Loading more complex files
BioSimSpace provides reader and writers for a variety of molecular file formats. Some of these split the molecular data over multiple files, e.g. a topology and coordinate file. To view these, pass all of the necessary files to `viewMolecules` in a list, e.g.
```
v = viewMolecules(["data/ala.top","data/ala.crd"])
```
This can be combined with molecule selection, e.g. to load and view only molecules 0-4 in the file pass the indicies of the molecules you want to view as a second argument to `viewMolecule`, e.g.
```
v = viewMolecules(["data/ala.top","data/ala.crd"], [0,1,2,3,4])
```
(in reality, all molecules are loaded, but only molecules specified by the indicies are viewed. You can still use `v.system()` to view all molecules)
```
v.system()
```
| github_jupyter |
```
!pip install morfessor
from collections import defaultdict
import nltk
from polyglot.text import Text
from nltk.tag import StanfordNERTagger
import os
CORPUS = os.path.join("../data", "item1")
## Encode UTF-u and remove non-printable characters
# document = filter(
# lambda char: char in string.printable,
# unicodedata.normalize('NFKD', document.decode('utf-8'))
# )
kddcorpus = nltk.corpus.PlaintextCorpusReader(CORPUS, '.*\.txt')
def polyglot_entities(fileids=None, section = None, corpus=kddcorpus):
"""
Extract entities from each file using polyglot
"""
results = defaultdict(lambda: defaultdict(list))
fileids = fileids or corpus.fileids()
for fileid in fileids:
if section is not None:
text = Text((list(sectpull([fileid],section=section))[0][1]))
else:
text = Text(corpus.raw(fileid))
for entity in text.entities:
etext = " ".join(entity)
if entity.tag == 'I-PER':
key = 'persons'
elif entity.tag == 'I-ORG':
key = 'organizations'
elif entity.tag == 'I-locations':
key = 'locations'
else:
key = 'other'
results[fileid][key].append(etext)
return results
def stanford_entities(model, jar, fileids=None, corpus=kddcorpus, section = None):
"""
Extract entities using the Stanford NER tagger.
Must pass in the path to the tagging model and jar as downloaded from the
Stanford Core NLP website.
"""
results = defaultdict(lambda: defaultdict(list))
fileids = fileids or corpus.fileids()
tagger = StanfordNERTagger(model, jar)
section = section
for fileid in fileids:
if section is not None:
text = nltk.word_tokenize(list(sectpull([fileid],section=section))[0][1])
else:
text = corpus.words(fileid)
chunk = []
for token, tag in tagger.tag(text):
if tag == 'O':
if chunk:
# Flush the current chunk
etext = " ".join([c[0] for c in chunk])
etag = chunk[0][1]
chunk = []
if etag == 'PERSON':
key = 'persons'
elif etag == 'ORGANIZATION':
key = 'organizations'
elif etag == 'LOCATION':
key = 'locations'
else:
key = 'other'
results[fileid][key].append(etext)
else:
# Build chunk from tags
chunk.append((token, tag))
return results
def nltk_entities(fileids=None, section = None,corpus=kddcorpus):
"""
Extract entities using the NLTK named entity chunker.
"""
results = defaultdict(lambda: defaultdict(list))
fileids = fileids or corpus.fileids()
for fileid in fileids:
if section is not None:
text = nltk.pos_tag(nltk.word_tokenize(list(sectpull([fileid],section=section))[0][1]))
else:
text = nltk.pos_tag(corpus.words(fileid))
for entity in nltk.ne_chunk(text):
if isinstance(entity, nltk.tree.Tree):
etext = " ".join([word for word, tag in entity.leaves()])
label = entity.label()
else:
continue
if label == 'PERSON':
key = 'persons'
elif label == 'ORGANIZATION':
key = 'organizations'
elif label == 'LOCATION':
key = 'locations'
elif label == 'GPE':
key = 'other'
else:
key = None
if key:
results[fileid][key].append(etext)
return results
# Only extract our annotated files.
fids = ['msft-item1-2018.txt', 'goog-item1-2016.txt']
# NLTK Entities
nltkents = nltk_entities(fids, section='top')
# Polyglot Entities
polyents = polyglot_entities(fids, section='top')
# Stanford Model Loading
root = os.path.expanduser('~/models/stanford-ner-2014-01-04/')
model = os.path.join(root, 'classifiers/english.muc.7class.distsim.crf.ser.gz')
jar = os.path.join(root, 'stanford-ner-2014-01-04.jar')
# Stanford Entities
stanents = stanford_entities(model, jar, fids, section='top')
##See https://www.districtdatalabs.com/named-entity-recognition-and-classification-for-entity-extraction
```
| github_jupyter |
# Land and Ocean Difference
```
import numpy as np
import pandas as pd
import xarray as xr
from tqdm import tqdm
import gc
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import matplotlib.ticker as mticker
from util import *
import geopandas as gpd
from geopandas import GeoDataFrame as gdf
def open_chi(path):
ds=(xr.open_dataset(path)*100)
ds=ds.assign_coords(lon=(((ds.lon + 180) % 360) - 180))
ds=ds.reindex(lon=sorted(ds.lon))
return ds
year = "2011"
method_ls = ["MAM4","ML"]
chi_ls = ["chi_b","chi_c","chi_h"]
file_path = {}
file_path["MAM4"] = "/data/keeling/a/zzheng25/d/mam4_paper_data/chi_only/mam4_chi/"
file_path["ML"] = "/data/keeling/a/zzheng25/d/mam4_paper_data/chi_only/ml_chi/"
file_path["diff"] = "/data/keeling/a/zzheng25/d/mam4_paper_data/chi_only/mam4_minus_ml_chi/"
file_path["diff_abs"] = "/data/keeling/a/zzheng25/d/mam4_paper_data/chi_only/mam4_minus_ml_chi/"
mask_path = "/data/keeling/a/zzheng25/d/mam4_paper_data/chi_only/mask/"
per_ls = [
"bc_a1_per","bc_a4_per",
"dst_a1_per","dst_a2_per",
"ncl_a1_per","ncl_a2_per",
"pom_a1_per","pom_a4_per",
"so4_a1_per","so4_a2_per",
"soa_a1_per","soa_a2_per"
]
comp = open_nc("/data/keeling/a/zzheng25/d/mam4_paper_data/chi_only/comp_analysis/"+str(year)+"_year_comp.nc")\
.to_dataframe()[per_ls].reset_index()
```
## load data
```
da={}
for chi in tqdm(chi_ls):
da[chi]={}
for method in method_ls:
if method=="diff_abs":
da_temp = open_chi(file_path[method]+str(year)+"_"+chi+"_mean_abs.nc")[chi]
mask = open_chi(mask_path+str(year)+"_"+chi+".nc")["mask"]
da[chi][method] = da_temp.where(mask)
else:
da_temp = open_chi(file_path[method]+str(year)+"_"+chi+"_mean.nc")[chi]
mask = open_chi(mask_path+str(year)+"_"+chi+".nc")["mask"]
da[chi][method] = da_temp.where(mask)
del da_temp, mask
gc.collect()
```
## Workflow
```
# get ocean and land data
def get_land_ocean(method, da):
chi_b=da["chi_b"][method].to_dataframe().reset_index()
chi_c=da["chi_c"][method].to_dataframe().reset_index()
chi_h=da["chi_h"][method].to_dataframe().reset_index()
merge_1=chi_b.merge(chi_c,on=["lat","lon"],how="outer")
df=merge_1.merge(chi_h,on=["lat","lon"],how="outer")
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
points= gdf(df.reset_index(), geometry=gpd.points_from_xy(df.reset_index().lon, df.reset_index().lat))
points.crs="EPSG:4326"
jps = gpd.sjoin(world, points, how='right', op='contains')
vari_ls_new = ["lat","lon","chi_b","chi_c","chi_h"]
land = jps[~jps["index_left"].isnull()][vari_ls_new]
ocean = jps[jps["index_left"].isnull()][vari_ls_new]
land_new = land.set_index(["lat","lon"]).stack()\
.reset_index(name="mixing_state_index")\
.rename(columns={'level_2':'mixing_state_type'})
print("land")
display(land_new.groupby(["mixing_state_type"])["mixing_state_index"].describe()["mean"])
ocean_new = ocean.set_index(["lat","lon"]).stack()\
.reset_index(name="mixing_state_index")\
.rename(columns={'level_2':'mixing_state_type'})
print("ocean")
display(ocean_new.groupby(["mixing_state_type"])["mixing_state_index"].describe()["mean"])
print("\n")
return land_new, ocean_new
# comp analysis for land or ocean
def comp_analysis(df, chi, lat_min=-90, lat_max=90, lon_min=-180, lon_max=180, comp=comp):
df_temp = df[(df["lat"]>=lat_min) & (df["lat"]<=lat_max) &
(df["lon"]>=lon_min) & (df["lon"]<=lon_max) &
(df["mixing_state_type"]==chi)]
# print(df_temp.shape)
df_temp_comp=df_temp.merge(comp, on=["lat","lon"], how="inner")
# print(df_temp_comp.shape)
display(df_temp_comp.describe().transpose()["mean"])
print("MAM4")
method="MAM4"
l_mam4, o_mam4 = get_land_ocean(method, da)
print("ML")
method="ML"
l_ml, o_ml = get_land_ocean(method, da)
```
## chi_opt1
### MAM4
```
chi = "chi_b"; lat_min = 45; lat_max = 60
comp_analysis(o_mam4, chi, lat_min, lat_max)
chi = "chi_b"; lat_min = -90; lat_max = -66.5
comp_analysis(l_mam4, chi, lat_min, lat_max)
```
| github_jupyter |
# Astronomy 8824 - Numerical and Statistical Methods in Astrophysics
## Introduction to Plotting
These notes are for the course Astronomy 8824: Numerical and Statistical Methods in Astrophysics.
#### Background reading:
- Statistics, Data Mining, and Machine Learning in Astronomy (SDMLA), Chapter 1
```
import math
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
import astropy
import sys, os
# matplotlib settings
SMALL_SIZE = 14
MEDIUM_SIZE = 16
BIGGER_SIZE = 18
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=BIGGER_SIZE) # fontsize of the x and y labels
plt.rc('lines', linewidth=2)
plt.rc('axes', linewidth=2)
plt.rc('xtick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=MEDIUM_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
def func8(x):
return 1./(x*x + x*x*x)
def func9(x):
return np.power(np.sin(x),2)/np.power(x,2)
```
### Create a simple plot
```
# Generate 100 random points on the interval (0,1)
np.random.seed(1216)
N = 100
bx = np.random.rand(N)
by = np.random.rand(N)
rx = np.random.rand(N)
ry = np.random.rand(N)
# Draw points of one color, circles of another
plt.figure(figsize=(8,8))
plt.plot(bx, by, 'b.')
plt.plot(rx, ry, 'ro')
plt.xlabel("X")
plt.ylabel("Y")
# Draw some plots with different line times based on functions in PS1
x = np.linspace(1,1000, 10000)
y8 = func8(x)
y9 = func9(x)
plt.figure(figsize=(10,7))
plt.plot(np.log(x), np.log(y8), 'k-', label="Func 8")
plt.plot(np.log(x), np.log(y9), "b:", label="Func 9")
plt.xlabel("X")
plt.ylabel("Y")
plt.legend()
```
### Plot a Gaussian distribution + histogram
```
# Here is a simple illustration of a histogram and a Gaussian distribution
def gaussian(x, mu, sigma):
return 1./(np.sqrt(2.*np.pi)*sigma)*np.exp(-np.power((x - mu)/sigma, 2.)/2)
plt.figure(figsize=(10, 6))
mu, sigma = 0, 1 # mean and standard deviation
g = np.random.normal(mu, sigma, 1000)
# Create a histogram of the random points
count, bins, ignored = plt.hist(g, 30, density=True, histtype='step', lw=2)
# Draw the gaussian
plt.plot(bins, gaussian(bins, mu, sigma), 'r', lw=2)
plt.xlim(-5, 5)
```
### subplot
This is a simple, multi-panel plot
```
Npts = 20
x = np.linspace(0, 5, Npts)
m = 2
b = 3
y = m*x + b
sig_y = np.random.normal(0, 1, Npts)
fx = y + sig_y
plt.figure(figsize=(10,7))
plt.subplot(211) # Note syntax is ncols nrols current
plt.plot(x, fx, 'bs', label="Data")
plt.plot(x, y, 'k:', label="Relation")
plt.ylabel("Y")
plt.legend(loc='upper left')
plt.subplot(212)
plt.plot(x, fx - y, 'bs', label="Residual")
plt.ylim(-5, 5)
plt.plot( [x[0], x[-1]], [0, 0], 'k:')
plt.ylabel("X")
plt.ylabel(r"$\Delta Y$")
plt.legend(loc='upper right')
```
### Create a plot with axes
axes can be used to explicitly plan out a figure, and offers greater flexibility for formatting
```
# This is a very simple application of axes
fig, ax = plt.subplots(figsize=(10,7))
ax.plot(x, fx, 'bs', label="Data")
ax.plot(x, y, 'k:', label="Relation")
ax.set_xlabel("X")
ax.set_ylabel("Y")
ax.legend()
```
### Create a multi-panel plot (subplots)
```
# Here is a slightly more intricate example
fig, axarr = plt.subplots(2, 1, figsize=(10,7))
axarr[0].plot(x, fx, 'bs', label="Data")
axarr[0].plot(x, y, 'k:', label="Relation")
axarr[0].set_xlabel("X")
axarr[0].set_ylabel("Y")
axarr[0].legend()
axarr[1].plot(x, fx - y, 'bs', label="Residual")
axarr[1].set_ylim(-5, 5)
axarr[1].plot( [x[0], x[-1]], [0, 0], 'k:')
axarr[1].set_ylabel(r"$\Delta Y$")
axarr[1].set_xlabel("X")
axarr[1].legend()
# Same plot with more sophistication
fig, axarr = plt.subplots(2, 1, figsize=(10,7), sharex=True)
axarr[0].plot(x, fx, 'bs', label="Data")
axarr[0].plot(x, y, 'k:', label="Relation")
axarr[0].set_ylabel("Y")
axarr[0].legend()
axarr[1].plot(x, fx - y, 'bs', label="Residual")
axarr[1].set_ylim(-5, 5)
axarr[1].plot( [x[0], x[-1]], [0, 0], 'k:')
axarr[1].set_ylabel(r"$\Delta Y$")
axarr[1].set_xlabel("X")
axarr[1].legend()
```
### Install astroML and healpy in ".local"
**astroML** is a package written by the authors of SDMLA and includes code to create all of the figures in the book
**healpy** is a module for using healpix, an acronym for _Hierarchical Equal Area isoLatitude Pixelization_ of a sphere. This subdivides the surface of a sphere (e.g. the sky) into pixels of equal area. Healpix representations are often used in CMB studies, large-scale clustering, among other applications.
```
! pip install astroml --user
! pip install healpy --user
# This is the 'user' installation path for OSC
newpath = os.path.join( os.getenv("HOME"), ".local/lib/python3.7/site-packages/")
sys.path.append(newpath)
```
### astroML
All of the code to construct the figures is available online:
https://www.astroml.org/book_figures/
Here are some examples from Chapter 1
```
import astroML
```
### Figure 1.1 from SDMLA
```
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from astroML.datasets import fetch_imaging_sample
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
if "setup_text_plots" not in globals():
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=16, usetex=True)
def get_stars_and_galaxies(Nstars=5000, Ngals=5000):
"""Get the subset of star/galaxy data to plot"""
data = fetch_imaging_sample()
objtype = data['type']
stars = data[objtype == 6][:Nstars]
galaxies = data[objtype == 3][:Ngals]
return stars, galaxies
def plot_stars_and_galaxies(stars, galaxies):
"""Plot the star and galaxy data"""
# Note: we use plot() rather than scatter() because it's more efficient
# for large numbers of points.
# Scatter should be used only when points need to be different colors
# and/or sizes
plot_kwargs = dict(color='k', linestyle='none', marker=',')
fig = plt.figure(figsize=(10, 7.5))
ax1 = fig.add_subplot(221)
ax1.plot(galaxies['gRaw'] - galaxies['rRaw'],
galaxies['rRaw'],
**plot_kwargs)
ax2 = fig.add_subplot(223, sharex=ax1)
ax2.plot(galaxies['gRaw'] - galaxies['rRaw'],
galaxies['rRaw'] - galaxies['iRaw'],
**plot_kwargs)
ax3 = fig.add_subplot(222, sharey=ax1)
ax3.plot(stars['gRaw'] - stars['rRaw'],
stars['rRaw'],
**plot_kwargs)
ax4 = fig.add_subplot(224, sharex=ax3, sharey=ax2)
ax4.plot(stars['gRaw'] - stars['rRaw'],
stars['rRaw'] - stars['iRaw'],
**plot_kwargs)
# set labels and titles
ax1.set_ylabel(r'${\rm r}$')
ax2.set_ylabel(r'${\rm r - i}$')
ax2.set_xlabel(r'${\rm g - r}$')
ax4.set_xlabel(r'${\rm g - r}$')
ax1.set_title('Galaxies')
ax3.set_title('Stars')
# set axis limits
ax2.set_xlim(-1, 3)
ax3.set_ylim(22.5, 14)
ax4.set_xlim(-1, 3)
ax4.set_ylim(-1, 2)
# adjust tick spacings on all axes
for ax in (ax1, ax2, ax3, ax4):
ax.xaxis.set_major_locator(plt.MultipleLocator(1))
ax.yaxis.set_major_locator(plt.MultipleLocator(1))
#------------------------------------------------------------
# Generate and show the plot
stars, galaxies = get_stars_and_galaxies()
plot_stars_and_galaxies(stars, galaxies)
plt.show()
```
### Figure 1.2 from SDMLA
```
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
from matplotlib import pyplot as plt
from astroML.datasets import fetch_sdss_spectrum
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
if "setup_text_plots" not in globals():
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=16, usetex=True)
#------------------------------------------------------------
# Fetch single spectrum
plate = 1615
mjd = 53166
fiber = 513
spec = fetch_sdss_spectrum(plate, mjd, fiber)
#------------------------------------------------------------
# Plot the resulting spectrum
fig, ax = plt.subplots(figsize=(10, 7.5))
ax.plot(spec.wavelength(), spec.spectrum, '-k', lw=1)
ax.set_xlim(3000, 10000)
ax.set_ylim(25, 300)
ax.set_xlabel(r'$\lambda {(\rm \AA)}$')
ax.set_ylabel('Flux')
ax.set_title('Plate = %(plate)i, MJD = %(mjd)i, Fiber = %(fiber)i' % locals())
plt.show()
```
### Figure 1.15 from SDMLA
```
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
from __future__ import print_function
import numpy as np
from matplotlib import pyplot as plt
# warning: due to a bug in healpy, importing it before pylab can cause
# a segmentation fault in some circumstances.
import healpy as hp
from astroML.datasets import fetch_wmap_temperatures
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
if "setup_text_plots" not in globals():
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=16, usetex=True)
#------------------------------------------------------------
# Next plot the wmap pixellization
wmap_unmasked = fetch_wmap_temperatures(masked=False)
# plot the unmasked map
fig = plt.figure(2, figsize=(10, 7.5))
hp.mollview(wmap_unmasked, min=-1, max=1, title='Raw WMAP data',
unit=r'$\Delta$T (mK)', fig=2)
fig.axes[1].texts[0].set_fontsize(16)
plt.show()
```
### Illustrate a few different healpy NSIDE values
For a healpy tutorial: https://healpy.readthedocs.io/en/latest/tutorial.html
For an introduction to healpix: https://healpix.jpl.nasa.gov/
```
nsides = np.power(2, np.arange(4))
#fig, axarr = plt.subplots(len(nsides), 1, figsize=(3*len(nsides),7))
# Prepare the healpix pixels
for i, nside in enumerate(nsides):
m = np.arange(hp.nside2npix(nside))
# Plot the pixelization
fig = plt.figure(i, figsize=(10, 7.5))
label = "HEALPix with NSIDE = {0} has {1} pixels".format(nside, len(m))
hp.mollview(m, nest=False, title=label, fig=i)
# remove colorbar: we don't need it for this plot
fig.delaxes(fig.axes[1])
# What is the pixel size for NSIDE = 64?
NSIDE = 64
print("Approximate resolution at NSIDE {} is {:.2} deg (per side)".format(
NSIDE, hp.nside2resol(NSIDE, arcmin=True) / 60) )
```
| github_jupyter |
# Concrete Feature Engineering
---
## Reference
> [What Is Feature Engineering](https://www.kaggle.com/ryanholbrook/what-is-feature-engineering)<br>
> [Data Source](https://www.kaggle.com/ryanholbrook/fe-course-data)
---
## Dependencies
```
import pandas as pd
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import cross_val_score
%load_ext autotime
```
---
## Import Dataset
To illustrate these ideas we'll see how adding a few synthetic features to a dataset can improve the predictive performance of a random forest model.
The [Concrete](https://www.kaggle.com/sinamhd9/concrete-comprehensive-strength) dataset contains a variety of concrete formulations and the resulting product's *compressive strength*, which is a measure of how much load that kind of concrete can bear. The task for this dataset is to predict a concrete's compressive strength given its formulation.
```
df = pd.read_csv("data/concrete.csv")
df.head()
```
---
## Baseline
You can see here the various ingredients going into each variety of concrete. We'll see in a moment how adding some additional synthetic features derived from these can help a model to learn important relationships among them.
We'll first establish a baseline by training the model on the un-augmented dataset. This will help us determine whether our new features are actually useful.
Establishing baselines like this is good practice at the start of the feature engineering process. A baseline score can help you decide whether your new features are worth keeping, or whether you should discard them and possibly try something else.
```
X = df.copy()
y = X.pop("CompressiveStrength")
# Train and score baseline model
baseline = RandomForestRegressor(criterion="mae", random_state=0)
baseline_score = cross_val_score(
baseline, X, y, cv=5, scoring="neg_mean_absolute_error"
)
baseline_score = -1 * baseline_score.mean()
print(f"MAE Baseline Score: {baseline_score:.4}")
```
If you ever cook at home, you might know that the ratio of ingredients in a recipe is usually a better predictor of how the recipe turns out than their absolute amounts. We might reason then that ratios of the features above would be a good predictor of `CompressiveStrength`.
The cell below adds three new ratio features to the dataset.
```
X = df.copy()
y = X.pop("CompressiveStrength")
# Create synthetic features
X["FCRatio"] = X["FineAggregate"] / X["CoarseAggregate"]
X["AggCmtRatio"] = (X["CoarseAggregate"] + X["FineAggregate"]) / X["Cement"]
X["WtrCmtRatio"] = X["Water"] / X["Cement"]
# Train and score model on dataset with additional ratio features
model = RandomForestRegressor(criterion="mae", random_state=0)
score = cross_val_score(
model, X, y, cv=5, scoring="neg_mean_absolute_error"
)
score = -1 * score.mean()
print(f"MAE Score with Ratio Features: {score:.4}")
```
And sure enough, performance improved! This is evidence that these new ratio features exposed important information to the model that it wasn't detecting before.
```
X.head()
```
| github_jupyter |
## Dependencies
```
!pip install --quiet efficientnet
# !pip install --quiet image-classifiers
import warnings, json, re, glob, math
from scripts_step_lr_schedulers import *
from melanoma_utility_scripts import *
from kaggle_datasets import KaggleDatasets
from sklearn.model_selection import KFold
import tensorflow.keras.layers as L
import tensorflow.keras.backend as K
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from tensorflow.keras import optimizers, layers, metrics, losses, Model
import efficientnet.tfkeras as efn
# from classification_models.tfkeras import Classifiers
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
## TPU configuration
```
strategy, tpu = set_up_strategy()
print("REPLICAS: ", strategy.num_replicas_in_sync)
AUTO = tf.data.experimental.AUTOTUNE
```
# Model parameters
```
config = {
"HEIGHT": 256,
"WIDTH": 256,
"CHANNELS": 3,
"BATCH_SIZE": 128,
"EPOCHS": 12,
"LEARNING_RATE": 3e-4,
"ES_PATIENCE": 10,
"N_FOLDS": 5,
"N_USED_FOLDS": 5,
"TTA_STEPS": 25,
"BASE_MODEL": 'EfficientNetB7',
"BASE_MODEL_WEIGHTS": 'noisy-student',
"DATASET_PATH": 'melanoma-256x256'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
config
```
# Load data
```
database_base_path = '/kaggle/input/siim-isic-melanoma-classification/'
k_fold = pd.read_csv(database_base_path + 'train.csv')
test = pd.read_csv(database_base_path + 'test.csv')
print('Train samples: %d' % len(k_fold))
display(k_fold.head())
print(f'Test samples: {len(test)}')
display(test.head())
GCS_PATH = 'gs://kds-65548a4c87d02212371fce6e9bd762100c34bf9b9ebbd04b0dd4b65b'# KaggleDatasets().get_gcs_path(config['DATASET_PATH'])
TRAINING_FILENAMES = tf.io.gfile.glob(GCS_PATH + '/train*.tfrec')
TEST_FILENAMES = tf.io.gfile.glob(GCS_PATH + '/test*.tfrec')
```
# Augmentations
```
def data_augment(image, label):
p_spatial = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_spatial2 = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_rotate = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_crop = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_pixel = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
### Spatial-level transforms
if p_spatial >= .2: # flips
image['input_image'] = tf.image.random_flip_left_right(image['input_image'])
image['input_image'] = tf.image.random_flip_up_down(image['input_image'])
if p_spatial >= .7:
image['input_image'] = tf.image.transpose(image['input_image'])
if p_rotate >= .8: # rotate 270º
image['input_image'] = tf.image.rot90(image['input_image'], k=3)
elif p_rotate >= .6: # rotate 180º
image['input_image'] = tf.image.rot90(image['input_image'], k=2)
elif p_rotate >= .4: # rotate 90º
image['input_image'] = tf.image.rot90(image['input_image'], k=1)
if p_spatial2 >= .6:
if p_spatial2 >= .9:
image['input_image'] = transform_rotation(image['input_image'], config['HEIGHT'], 180.)
elif p_spatial2 >= .8:
image['input_image'] = transform_zoom(image['input_image'], config['HEIGHT'], 8., 8.)
elif p_spatial2 >= .7:
image['input_image'] = transform_shift(image['input_image'], config['HEIGHT'], 8., 8.)
else:
image['input_image'] = transform_shear(image['input_image'], config['HEIGHT'], 2.)
if p_crop >= .6: # crops
if p_crop >= .8:
image['input_image'] = tf.image.random_crop(image['input_image'], size=[int(config['HEIGHT']*.8), int(config['WIDTH']*.8), config['CHANNELS']])
elif p_crop >= .7:
image['input_image'] = tf.image.random_crop(image['input_image'], size=[int(config['HEIGHT']*.9), int(config['WIDTH']*.9), config['CHANNELS']])
else:
image['input_image'] = tf.image.central_crop(image['input_image'], central_fraction=.8)
image['input_image'] = tf.image.resize(image['input_image'], size=[config['HEIGHT'], config['WIDTH']])
if p_pixel >= .6: # Pixel-level transforms
if p_pixel >= .9:
image['input_image'] = tf.image.random_hue(image['input_image'], 0.01)
elif p_pixel >= .8:
image['input_image'] = tf.image.random_saturation(image['input_image'], 0.7, 1.3)
elif p_pixel >= .7:
image['input_image'] = tf.image.random_contrast(image['input_image'], 0.8, 1.2)
else:
image['input_image'] = tf.image.random_brightness(image['input_image'], 0.1)
return image, label
```
## Auxiliary functions
```
# Datasets utility functions
def read_labeled_tfrecord(example, height=config['HEIGHT'], width=config['WIDTH'], channels=config['CHANNELS']):
example = tf.io.parse_single_example(example, LABELED_TFREC_FORMAT)
image = decode_image(example['image'], height, width, channels)
label = tf.cast(example['target'], tf.float32)
# meta features
data = {}
data['patient_id'] = tf.cast(example['patient_id'], tf.int32)
data['sex'] = tf.cast(example['sex'], tf.int32)
data['age_approx'] = tf.cast(example['age_approx'], tf.int32)
data['anatom_site_general_challenge'] = tf.cast(tf.one_hot(example['anatom_site_general_challenge'], 7), tf.int32)
return {'input_image': image, 'input_meta': data}, label # returns a dataset of (image, data, label)
def read_labeled_tfrecord_eval(example, height=config['HEIGHT'], width=config['WIDTH'], channels=config['CHANNELS']):
example = tf.io.parse_single_example(example, LABELED_TFREC_FORMAT)
image = decode_image(example['image'], height, width, channels)
label = tf.cast(example['target'], tf.float32)
image_name = example['image_name']
# meta features
data = {}
data['patient_id'] = tf.cast(example['patient_id'], tf.int32)
data['sex'] = tf.cast(example['sex'], tf.int32)
data['age_approx'] = tf.cast(example['age_approx'], tf.int32)
data['anatom_site_general_challenge'] = tf.cast(tf.one_hot(example['anatom_site_general_challenge'], 7), tf.int32)
return {'input_image': image, 'input_meta': data}, label, image_name # returns a dataset of (image, data, label, image_name)
def load_dataset(filenames, ordered=False, buffer_size=-1):
ignore_order = tf.data.Options()
if not ordered:
ignore_order.experimental_deterministic = False # disable order, increase speed
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=buffer_size) # automatically interleaves reads from multiple files
dataset = dataset.with_options(ignore_order) # uses data as soon as it streams in, rather than in its original order
dataset = dataset.map(read_labeled_tfrecord, num_parallel_calls=buffer_size)
return dataset # returns a dataset of (image, data, label)
def load_dataset_eval(filenames, buffer_size=-1):
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=buffer_size) # automatically interleaves reads from multiple files
dataset = dataset.map(read_labeled_tfrecord_eval, num_parallel_calls=buffer_size)
return dataset # returns a dataset of (image, data, label, image_name)
def get_training_dataset(filenames, batch_size, buffer_size=-1):
dataset = load_dataset(filenames, ordered=False, buffer_size=buffer_size)
dataset = dataset.map(data_augment, num_parallel_calls=AUTO)
dataset = dataset.repeat() # the training dataset must repeat for several epochs
dataset = dataset.shuffle(2048)
dataset = dataset.batch(batch_size, drop_remainder=True) # slighly faster with fixed tensor sizes
dataset = dataset.prefetch(buffer_size) # prefetch next batch while training (autotune prefetch buffer size)
return dataset
def get_validation_dataset(filenames, ordered=True, repeated=False, batch_size=32, buffer_size=-1):
dataset = load_dataset(filenames, ordered=ordered, buffer_size=buffer_size)
if repeated:
dataset = dataset.repeat()
dataset = dataset.shuffle(2048)
dataset = dataset.batch(batch_size, drop_remainder=repeated)
dataset = dataset.prefetch(buffer_size)
return dataset
def get_eval_dataset(filenames, batch_size=32, buffer_size=-1):
dataset = load_dataset_eval(filenames, buffer_size=buffer_size)
dataset = dataset.batch(batch_size, drop_remainder=False)
dataset = dataset.prefetch(buffer_size)
return dataset
# Test function
def read_unlabeled_tfrecord(example, height=config['HEIGHT'], width=config['WIDTH'], channels=config['CHANNELS']):
example = tf.io.parse_single_example(example, UNLABELED_TFREC_FORMAT)
image = decode_image(example['image'], height, width, channels)
image_name = example['image_name']
# meta features
data = {}
data['patient_id'] = tf.cast(example['patient_id'], tf.int32)
data['sex'] = tf.cast(example['sex'], tf.int32)
data['age_approx'] = tf.cast(example['age_approx'], tf.int32)
data['anatom_site_general_challenge'] = tf.cast(tf.one_hot(example['anatom_site_general_challenge'], 7), tf.int32)
return {'input_image': image, 'input_tabular': data}, image_name # returns a dataset of (image, data, image_name)
def load_dataset_test(filenames, buffer_size=-1):
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=buffer_size) # automatically interleaves reads from multiple files
dataset = dataset.map(read_unlabeled_tfrecord, num_parallel_calls=buffer_size)
# returns a dataset of (image, data, label, image_name) pairs if labeled=True or (image, data, image_name) pairs if labeled=False
return dataset
def get_test_dataset(filenames, batch_size=32, buffer_size=-1, tta=False):
dataset = load_dataset_test(filenames, buffer_size=buffer_size)
if tta:
dataset = dataset.map(data_augment, num_parallel_calls=AUTO)
dataset = dataset.batch(batch_size, drop_remainder=False)
dataset = dataset.prefetch(buffer_size)
return dataset
# Advanced augmentations
def transform_rotation(image, height, rotation):
# input image - is one image of size [dim,dim,3] not a batch of [b,dim,dim,3]
# output - image randomly rotated
DIM = height
XDIM = DIM%2 #fix for size 331
rotation = rotation * tf.random.normal([1],dtype='float32')
# CONVERT DEGREES TO RADIANS
rotation = math.pi * rotation / 180.
# ROTATION MATRIX
c1 = tf.math.cos(rotation)
s1 = tf.math.sin(rotation)
one = tf.constant([1],dtype='float32')
zero = tf.constant([0],dtype='float32')
rotation_matrix = tf.reshape( tf.concat([c1,s1,zero, -s1,c1,zero, zero,zero,one],axis=0),[3,3] )
# LIST DESTINATION PIXEL INDICES
x = tf.repeat( tf.range(DIM//2,-DIM//2,-1), DIM )
y = tf.tile( tf.range(-DIM//2,DIM//2),[DIM] )
z = tf.ones([DIM*DIM],dtype='int32')
idx = tf.stack( [x,y,z] )
# ROTATE DESTINATION PIXELS ONTO ORIGIN PIXELS
idx2 = K.dot(rotation_matrix,tf.cast(idx,dtype='float32'))
idx2 = K.cast(idx2,dtype='int32')
idx2 = K.clip(idx2,-DIM//2+XDIM+1,DIM//2)
# FIND ORIGIN PIXEL VALUES
idx3 = tf.stack( [DIM//2-idx2[0,], DIM//2-1+idx2[1,]] )
d = tf.gather_nd(image, tf.transpose(idx3))
return tf.reshape(d,[DIM,DIM,3])
def transform_shear(image, height, shear):
# input image - is one image of size [dim,dim,3] not a batch of [b,dim,dim,3]
# output - image randomly sheared
DIM = height
XDIM = DIM%2 #fix for size 331
shear = shear * tf.random.normal([1],dtype='float32')
shear = math.pi * shear / 180.
# SHEAR MATRIX
one = tf.constant([1],dtype='float32')
zero = tf.constant([0],dtype='float32')
c2 = tf.math.cos(shear)
s2 = tf.math.sin(shear)
shear_matrix = tf.reshape( tf.concat([one,s2,zero, zero,c2,zero, zero,zero,one],axis=0),[3,3] )
# LIST DESTINATION PIXEL INDICES
x = tf.repeat( tf.range(DIM//2,-DIM//2,-1), DIM )
y = tf.tile( tf.range(-DIM//2,DIM//2),[DIM] )
z = tf.ones([DIM*DIM],dtype='int32')
idx = tf.stack( [x,y,z] )
# ROTATE DESTINATION PIXELS ONTO ORIGIN PIXELS
idx2 = K.dot(shear_matrix,tf.cast(idx,dtype='float32'))
idx2 = K.cast(idx2,dtype='int32')
idx2 = K.clip(idx2,-DIM//2+XDIM+1,DIM//2)
# FIND ORIGIN PIXEL VALUES
idx3 = tf.stack( [DIM//2-idx2[0,], DIM//2-1+idx2[1,]] )
d = tf.gather_nd(image, tf.transpose(idx3))
return tf.reshape(d,[DIM,DIM,3])
def transform_shift(image, height, h_shift, w_shift):
# input image - is one image of size [dim,dim,3] not a batch of [b,dim,dim,3]
# output - image randomly shifted
DIM = height
XDIM = DIM%2 #fix for size 331
height_shift = h_shift * tf.random.normal([1],dtype='float32')
width_shift = w_shift * tf.random.normal([1],dtype='float32')
one = tf.constant([1],dtype='float32')
zero = tf.constant([0],dtype='float32')
# SHIFT MATRIX
shift_matrix = tf.reshape( tf.concat([one,zero,height_shift, zero,one,width_shift, zero,zero,one],axis=0),[3,3] )
# LIST DESTINATION PIXEL INDICES
x = tf.repeat( tf.range(DIM//2,-DIM//2,-1), DIM )
y = tf.tile( tf.range(-DIM//2,DIM//2),[DIM] )
z = tf.ones([DIM*DIM],dtype='int32')
idx = tf.stack( [x,y,z] )
# ROTATE DESTINATION PIXELS ONTO ORIGIN PIXELS
idx2 = K.dot(shift_matrix,tf.cast(idx,dtype='float32'))
idx2 = K.cast(idx2,dtype='int32')
idx2 = K.clip(idx2,-DIM//2+XDIM+1,DIM//2)
# FIND ORIGIN PIXEL VALUES
idx3 = tf.stack( [DIM//2-idx2[0,], DIM//2-1+idx2[1,]] )
d = tf.gather_nd(image, tf.transpose(idx3))
return tf.reshape(d,[DIM,DIM,3])
def transform_zoom(image, height, h_zoom, w_zoom):
# input image - is one image of size [dim,dim,3] not a batch of [b,dim,dim,3]
# output - image randomly zoomed
DIM = height
XDIM = DIM%2 #fix for size 331
height_zoom = 1.0 + tf.random.normal([1],dtype='float32')/h_zoom
width_zoom = 1.0 + tf.random.normal([1],dtype='float32')/w_zoom
one = tf.constant([1],dtype='float32')
zero = tf.constant([0],dtype='float32')
# ZOOM MATRIX
zoom_matrix = tf.reshape( tf.concat([one/height_zoom,zero,zero, zero,one/width_zoom,zero, zero,zero,one],axis=0),[3,3] )
# LIST DESTINATION PIXEL INDICES
x = tf.repeat( tf.range(DIM//2,-DIM//2,-1), DIM )
y = tf.tile( tf.range(-DIM//2,DIM//2),[DIM] )
z = tf.ones([DIM*DIM],dtype='int32')
idx = tf.stack( [x,y,z] )
# ROTATE DESTINATION PIXELS ONTO ORIGIN PIXELS
idx2 = K.dot(zoom_matrix,tf.cast(idx,dtype='float32'))
idx2 = K.cast(idx2,dtype='int32')
idx2 = K.clip(idx2,-DIM//2+XDIM+1,DIM//2)
# FIND ORIGIN PIXEL VALUES
idx3 = tf.stack( [DIM//2-idx2[0,], DIM//2-1+idx2[1,]] )
d = tf.gather_nd(image, tf.transpose(idx3))
return tf.reshape(d,[DIM,DIM,3])
```
## Learning rate scheduler
```
lr_min = 1e-6
lr_start = 5e-6
lr_max = config['LEARNING_RATE']
steps_per_epoch = 24844 // config['BATCH_SIZE']
total_steps = config['EPOCHS'] * steps_per_epoch
warmup_steps = steps_per_epoch * 5
hold_max_steps = 0
step_decay = .8
step_size = steps_per_epoch * 1
rng = [i for i in range(0, total_steps, 32)]
y = [step_schedule_with_warmup(tf.cast(x, tf.float32), step_size=step_size,
warmup_steps=warmup_steps, hold_max_steps=hold_max_steps,
lr_start=lr_start, lr_max=lr_max, step_decay=step_decay) for x in rng]
sns.set(style="whitegrid")
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
```
# Model
```
# Initial bias
pos = len(k_fold[k_fold['target'] == 1])
neg = len(k_fold[k_fold['target'] == 0])
initial_bias = np.log([pos/neg])
print('Bias')
print(pos)
print(neg)
print(initial_bias)
# class weights
total = len(k_fold)
weight_for_0 = (1 / neg)*(total)/2.0
weight_for_1 = (1 / pos)*(total)/2.0
class_weight = {0: weight_for_0, 1: weight_for_1}
print('Class weight')
print(class_weight)
def model_fn(input_shape):
input_image = L.Input(shape=input_shape, name='input_image')
base_model = efn.EfficientNetB7(weights=config['BASE_MODEL_WEIGHTS'],
include_top=False)
x = base_model(input_image)
x = L.GlobalAveragePooling2D()(x)
output = L.Dense(1, activation='sigmoid', name='output',
bias_initializer=tf.keras.initializers.Constant(initial_bias))(x)
model = Model(inputs=input_image, outputs=output)
return model
```
# Training
```
# Evaluation
eval_dataset = get_eval_dataset(TRAINING_FILENAMES, batch_size=config['BATCH_SIZE'], buffer_size=AUTO)
image_names = next(iter(eval_dataset.unbatch().map(lambda data, label, image_name: image_name).batch(count_data_items(TRAINING_FILENAMES)))).numpy().astype('U')
image_data = eval_dataset.map(lambda data, label, image_name: data)
# Test
NUM_TEST_IMAGES = len(test)
test_preds = np.zeros((NUM_TEST_IMAGES, 1))
test_preds_tta = np.zeros((NUM_TEST_IMAGES, 1))
test_preds_last = np.zeros((NUM_TEST_IMAGES, 1))
test_preds_tta_last = np.zeros((NUM_TEST_IMAGES, 1))
test_dataset = get_test_dataset(TEST_FILENAMES, batch_size=config['BATCH_SIZE'], buffer_size=AUTO)
test_dataset_tta = get_test_dataset(TEST_FILENAMES, batch_size=config['BATCH_SIZE'], buffer_size=AUTO, tta=True)
image_names_test = next(iter(test_dataset.unbatch().map(lambda data, image_name: image_name).batch(NUM_TEST_IMAGES))).numpy().astype('U')
test_image_data = test_dataset.map(lambda data, image_name: data)
test_tta_image_data = test_dataset_tta.map(lambda data, image_name: data)
history_list = []
k_fold_best = k_fold.copy()
kfold = KFold(config['N_FOLDS'], shuffle=True, random_state=SEED)
for n_fold, (trn_idx, val_idx) in enumerate(kfold.split(TRAINING_FILENAMES)):
if n_fold < config['N_USED_FOLDS']:
n_fold +=1
print('\nFOLD: %d' % (n_fold))
tf.tpu.experimental.initialize_tpu_system(tpu)
K.clear_session()
### Data
train_filenames = np.array(TRAINING_FILENAMES)[trn_idx]
valid_filenames = np.array(TRAINING_FILENAMES)[val_idx]
steps_per_epoch = count_data_items(train_filenames) // config['BATCH_SIZE']
# Train model
model_path = f'model_fold_{n_fold}.h5'
es = EarlyStopping(monitor='val_auc', mode='max', patience=config['ES_PATIENCE'],
restore_best_weights=False, verbose=1)
checkpoint = ModelCheckpoint(model_path, monitor='val_auc', mode='max',
save_best_only=True, save_weights_only=True)
with strategy.scope():
model = model_fn((config['HEIGHT'], config['WIDTH'], config['CHANNELS']))
lr = lambda: step_schedule_with_warmup(tf.cast(optimizer.iterations, tf.float32),
step_size=step_size, warmup_steps=warmup_steps,
hold_max_steps=hold_max_steps, lr_start=lr_start,
lr_max=lr_max, step_decay=step_decay)
optimizer = optimizers.Adam(learning_rate=lr)
model.compile(optimizer, loss=losses.BinaryCrossentropy(label_smoothing=0.05),
metrics=[metrics.AUC()])
history = model.fit(get_training_dataset(train_filenames, batch_size=config['BATCH_SIZE'], buffer_size=AUTO),
validation_data=get_validation_dataset(valid_filenames, ordered=True, repeated=False,
batch_size=config['BATCH_SIZE'], buffer_size=AUTO),
epochs=config['EPOCHS'],
steps_per_epoch=steps_per_epoch,
callbacks=[checkpoint, es],
class_weight=class_weight,
verbose=2).history
# save last epoch weights
model.save_weights('last_' + model_path)
history_list.append(history)
# Get validation IDs
valid_dataset = get_eval_dataset(valid_filenames, batch_size=config['BATCH_SIZE'], buffer_size=AUTO)
valid_image_names = next(iter(valid_dataset.unbatch().map(lambda data, label, image_name: image_name).batch(count_data_items(valid_filenames)))).numpy().astype('U')
k_fold[f'fold_{n_fold}'] = k_fold.apply(lambda x: 'validation' if x['image_name'] in valid_image_names else 'train', axis=1)
k_fold_best[f'fold_{n_fold}'] = k_fold_best.apply(lambda x: 'validation' if x['image_name'] in valid_image_names else 'train', axis=1)
##### Last model #####
print('Last model evaluation...')
preds = model.predict(image_data)
name_preds_eval = dict(zip(image_names, preds.reshape(len(preds))))
k_fold[f'pred_fold_{n_fold}'] = k_fold.apply(lambda x: name_preds_eval[x['image_name']], axis=1)
print('Last model inference...')
test_preds_last += model.predict(test_image_data)
# TTA preds
print(f'Running TTA (last) {config["TTA_STEPS"]} steps...')
for step in range(config['TTA_STEPS']):
test_preds_tta_last += model.predict(test_tta_image_data)
##### Best model #####
print('Best model evaluation...')
model.load_weights(model_path)
preds = model.predict(image_data)
name_preds_eval = dict(zip(image_names, preds.reshape(len(preds))))
k_fold_best[f'pred_fold_{n_fold}'] = k_fold_best.apply(lambda x: name_preds_eval[x['image_name']], axis=1)
print('Best model inference...')
test_preds += model.predict(test_image_data)
# TTA preds
print(f'Running TTA (best) {config["TTA_STEPS"]} steps...')
for step in range(config['TTA_STEPS']):
test_preds_tta += model.predict(test_tta_image_data)
# normalize preds
test_preds /= config['N_USED_FOLDS']
test_preds_tta /= (config['N_USED_FOLDS'] * config['TTA_STEPS'])
test_preds_last /= config['N_USED_FOLDS']
test_preds_tta_last /= (config['N_USED_FOLDS'] * config['TTA_STEPS'])
name_preds = dict(zip(image_names_test, test_preds.reshape(NUM_TEST_IMAGES)))
name_preds_tta = dict(zip(image_names_test, test_preds_tta.reshape(NUM_TEST_IMAGES)))
name_preds_last = dict(zip(image_names_test, test_preds_last.reshape(NUM_TEST_IMAGES)))
name_preds_tta_last = dict(zip(image_names_test, test_preds_tta_last.reshape(NUM_TEST_IMAGES)))
test['target'] = test.apply(lambda x: name_preds[x['image_name']], axis=1)
test['target_tta'] = test.apply(lambda x: name_preds_tta[x['image_name']], axis=1)
test['target_last'] = test.apply(lambda x: name_preds_last[x['image_name']], axis=1)
test['target_tta_last'] = test.apply(lambda x: name_preds_tta_last[x['image_name']], axis=1)
```
## Model loss graph
```
for n_fold in range(config['N_USED_FOLDS']):
print(f'Fold: {n_fold + 1}')
plot_metrics(history_list[n_fold])
```
## Model loss graph aggregated
```
plot_metrics_agg(history_list, config['N_USED_FOLDS'])
```
# Model evaluation (last)
```
display(evaluate_model(k_fold, config['N_USED_FOLDS']).style.applymap(color_map))
display(evaluate_model_Subset(k_fold, config['N_USED_FOLDS']).style.applymap(color_map))
```
# Model evaluation (best)
```
display(evaluate_model(k_fold_best, config['N_USED_FOLDS']).style.applymap(color_map))
display(evaluate_model_Subset(k_fold_best, config['N_USED_FOLDS']).style.applymap(color_map))
```
# Confusion matrix
```
for n_fold in range(config['N_USED_FOLDS']):
n_fold += 1
pred_col = f'pred_fold_{n_fold}'
train_set = k_fold_best[k_fold_best[f'fold_{n_fold}'] == 'train']
valid_set = k_fold_best[k_fold_best[f'fold_{n_fold}'] == 'validation']
print(f'Fold: {n_fold}')
plot_confusion_matrix(train_set['target'], np.round(train_set[pred_col]),
valid_set['target'], np.round(valid_set[pred_col]))
```
# Visualize predictions
```
k_fold['pred'] = 0
for n_fold in range(config['N_USED_FOLDS']):
k_fold['pred'] += k_fold[f'pred_fold_{n_fold+1}'] / config['N_FOLDS']
print('Label/prediction distribution')
print(f"Train positive labels: {len(k_fold[k_fold['target'] > .5])}")
print(f"Train positive predictions: {len(k_fold[k_fold['pred'] > .5])}")
print(f"Train positive correct predictions: {len(k_fold[(k_fold['target'] > .5) & (k_fold['pred'] > .5)])}")
print('Top 10 samples')
display(k_fold[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'diagnosis',
'target', 'pred'] + [c for c in k_fold.columns if (c.startswith('pred_fold'))]].head(10))
print('Top 10 positive samples')
display(k_fold[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'diagnosis',
'target', 'pred'] + [c for c in k_fold.columns if (c.startswith('pred_fold'))]].query('target == 1').head(10))
print('Top 10 predicted positive samples')
display(k_fold[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'diagnosis',
'target', 'pred'] + [c for c in k_fold.columns if (c.startswith('pred_fold'))]].query('pred > .5').head(10))
```
# Visualize test predictions
```
print(f"Test predictions {len(test[test['target'] > .5])}|{len(test[test['target'] <= .5])}")
print(f"Test predictions (last) {len(test[test['target_last'] > .5])}|{len(test[test['target_last'] <= .5])}")
print(f"Test predictions (tta) {len(test[test['target_tta'] > .5])}|{len(test[test['target_tta'] <= .5])}")
print(f"Test predictions (last tta) {len(test[test['target_tta_last'] > .5])}|{len(test[test['target_tta_last'] <= .5])}")
print('Top 10 samples')
display(test[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'target', 'target_last',
'target_tta', 'target_tta_last'] + [c for c in test.columns if (c.startswith('pred_fold'))]].head(10))
print('Top 10 positive samples')
display(test[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'target', 'target_last',
'target_tta', 'target_tta_last'] + [c for c in test.columns if (c.startswith('pred_fold'))]].query('target > .5').head(10))
print('Top 10 positive samples (last)')
display(test[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'target', 'target_last',
'target_tta', 'target_tta_last'] + [c for c in test.columns if (c.startswith('pred_fold'))]].query('target_last > .5').head(10))
```
# Test set predictions
```
submission = pd.read_csv(database_base_path + 'sample_submission.csv')
submission['target'] = test['target']
submission['target_last'] = test['target_last']
submission['target_blend'] = (test['target'] * .5) + (test['target_last'] * .5)
submission['target_tta'] = test['target_tta']
submission['target_tta_last'] = test['target_tta_last']
submission['target_tta_blend'] = (test['target_tta'] * .5) + (test['target_tta_last'] * .5)
display(submission.head(10))
display(submission.describe())
### BEST ###
submission[['image_name', 'target']].to_csv('submission.csv', index=False)
### LAST ###
submission_last = submission[['image_name', 'target_last']]
submission_last.columns = ['image_name', 'target']
submission_last.to_csv('submission_last.csv', index=False)
### BLEND ###
submission_blend = submission[['image_name', 'target_blend']]
submission_blend.columns = ['image_name', 'target']
submission_blend.to_csv('submission_blend.csv', index=False)
### TTA ###
submission_tta = submission[['image_name', 'target_tta']]
submission_tta.columns = ['image_name', 'target']
submission_tta.to_csv('submission_tta.csv', index=False)
### TTA LAST ###
submission_tta_last = submission[['image_name', 'target_tta_last']]
submission_tta_last.columns = ['image_name', 'target']
submission_tta_last.to_csv('submission_tta_last.csv', index=False)
### TTA BLEND ###
submission_blend_tta = submission[['image_name', 'target_tta_blend']]
submission_blend_tta.columns = ['image_name', 'target']
submission_blend_tta.to_csv('submission_blend_tta.csv', index=False)
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# `FishboneMoncriefID`: An Einstein Toolkit Initial Data Thorn for Fishbone-Moncrief initial data
## Author: Zach Etienne
### Formatting improvements courtesy Brandon Clark
[comment]: <> (Abstract: TODO)
**Notebook Status:** <font color='green'><b> Validated </b></font>
**Validation Notes:** Agrees with trusted Fishbone-Moncrief initial data module in HARM3D. Also generates results in agreement with trusted version sent to Event Horizon Telescope (EHT) GRMHD code comparison project collaborators. This thorn was used for the [IllinoisGRMHD](http://illinoisgrmhd.net) contribution to the [EHT GRMHD code comparison project](https://arxiv.org/abs/1904.04923).
### NRPy+ Source Code for this module: [FishboneMoncriefID/FishboneMoncriefID.py](../edit/FishboneMoncriefID/FishboneMoncriefID.py) [\[tutorial\]](Tutorial-FishboneMoncriefID.ipynb) Constructs SymPy expressions for [Fishbone-Moncrief initial data](Tutorial-FishboneMoncriefID.ipynb)
## Introduction:
In this part of the tutorial, we will construct an Einstein Toolkit (ETK) thorn (module) that will set up Fishbone-Moncrief initial data. In the [Tutorial-FishboneMoncriefID](Tutorial-FishboneMoncriefID.ipynb) tutorial notebook, we used NRPy+ to construct the SymPy expressions for Fishbone-Moncrief initial data.
We will construct this thorn in two steps.
1. Call on NRPy+ to convert the SymPy expressions for the initial data into one C-code kernel.
1. Write the C code and linkages to the Einstein Toolkit infrastructure (i.e., the .ccl files) to complete this Einstein Toolkit module.
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#initializenrpy): Call on NRPy+ to convert the SymPy expression for the Fishbone-Moncrief initial data into a C-code kernel
1. [Step 2](#einstein): Interfacing with the Einstein Toolkit
1. [Step 2.a](#einstein_c): Constructing the Einstein Toolkit C-code calling functions that include the C code kernels
1. [Step 2.b](#einstein_ccl): CCL files - Define how this module interacts and interfaces with the larger Einstein Toolkit infrastructure
1. [Step 2.c](#einstein_list): Add the C code to the Einstein Toolkit compilation list
1. [Step 3](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initializenrpy'></a>
# Step 1: Call on NRPy+ to convert the SymPy expression for the Fishbone-Moncrief initial data into a C-code kernel \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
After importing the core modules, we will set `GridFuncMemAccess` to `ETK`. SymPy expressions for Fishbone-Moncrief initial data are written inside [FishboneMoncriefID/FishboneMoncriefID.py](../edit/FishboneMoncriefID/FishboneMoncriefID.py), and we simply import them for use here.
```
# Step 1: Call on NRPy+ to convert the SymPy expression for the
# Fishbone-Moncrief initial data into a C-code kernel
# Step 1a: Import needed NRPy+ core modules:
import NRPy_param_funcs as par
import indexedexp as ixp
import grid as gri
import finite_difference as fin
from outputC import *
import loop
# Step 1b: This is an Einstein Toolkit (ETK) thorn. Here we
# tell NRPy+ that gridfunction memory access will
# therefore be in the "ETK" style.
par.set_parval_from_str("grid::GridFuncMemAccess","ETK")
par.set_parval_from_str("grid::DIM", 3)
DIM = par.parval_from_str("grid::DIM")
# Step 1c: Call the FishboneMoncriefID() function from within the
# FishboneMoncriefID/FishboneMoncriefID.py module.
import FishboneMoncriefID.FishboneMoncriefID as fmid
# Step 1d: Within the ETK, the 3D gridfunctions x, y, and z store the
# Cartesian grid coordinates. Setting the gri.xx[] arrays
# to point to these gridfunctions forces NRPy+ to treat
# the Cartesian coordinate gridfunctions properly --
# reading them from memory as needed.
xcoord,ycoord,zcoord = gri.register_gridfunctions("AUX",["xcoord","ycoord","zcoord"])
gri.xx[0] = xcoord
gri.xx[1] = ycoord
gri.xx[2] = zcoord
# Step 1e: Set up the Fishbone-Moncrief initial data. This sets all the ID gridfunctions.
fmid.FishboneMoncriefID()
Valencia3velocityU = ixp.register_gridfunctions_for_single_rank1("EVOL","Valencia3velocityU")
# -={ Spacetime quantities: Generate C code from expressions and output to file }=-
KerrSchild_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","alpha"),rhs=fmid.IDalpha),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU0"),rhs=fmid.IDbetaU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU1"),rhs=fmid.IDbetaU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU2"),rhs=fmid.IDbetaU[2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD00"),rhs=fmid.IDgammaDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD01"),rhs=fmid.IDgammaDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD02"),rhs=fmid.IDgammaDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD11"),rhs=fmid.IDgammaDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD12"),rhs=fmid.IDgammaDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD22"),rhs=fmid.IDgammaDD[2][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD00"),rhs=fmid.IDKDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD01"),rhs=fmid.IDKDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD02"),rhs=fmid.IDKDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD11"),rhs=fmid.IDKDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD12"),rhs=fmid.IDKDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD22"),rhs=fmid.IDKDD[2][2]),\
]
# Force outCverbose=False for this module to avoid gigantic C files
# filled with the non-CSE expressions for the Weyl scalars.
KerrSchild_CcodeKernel = fin.FD_outputC("returnstring",KerrSchild_to_print,params="outCverbose=False")
# -={ GRMHD quantities: Generate C code from expressions and output to file }=-
FMdisk_GRHD_rho_initial_to_print = [lhrh(lhs=gri.gfaccess("out_gfs","rho_initial"),rhs=fmid.rho_initial)]
FMdisk_GRHD_rho_initial_CcodeKernel = fin.FD_outputC("returnstring",FMdisk_GRHD_rho_initial_to_print)
FMdisk_GRHD_velocities_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","Valencia3velocityU0"),rhs=fmid.IDValencia3velocityU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","Valencia3velocityU1"),rhs=fmid.IDValencia3velocityU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","Valencia3velocityU2"),rhs=fmid.IDValencia3velocityU[2]),\
]
FMdisk_GRHD_velocities_CcodeKernel = fin.FD_outputC("returnstring",FMdisk_GRHD_velocities_to_print)
#KerrSchild_looped = loop.loop(["i2","i1","i0"],["0","0","0"],["cctk_lsh[2]","cctk_lsh[1]","cctk_lsh[0]"],\
# ["1","1","1"],["#pragma omp parallel for","",""],"",\
# KerrSchild_CcodeKernel.replace("time","cctk_time"))
#FMdisk_GRHD_velocities_looped = loop.loop(["i2","i1","i0"],["0","0","0"],["cctk_lsh[2]","cctk_lsh[1]","cctk_lsh[0]"],\
# ["1","1","1"],["#pragma omp parallel for","",""],"",\
# FMdisk_GRHD_velocities_CcodeKernel.replace("time","cctk_time"))
#FMdisk_GRHD_rho_initial_looped = loop.loop(["i2","i1","i0"],["0","0","0"],["cctk_lsh[2]","cctk_lsh[1]","cctk_lsh[0]"],\
# ["1","1","1"],["#pragma omp parallel for","",""],"",\
# FMdisk_GRHD_rho_initial_CcodeKernel.replace("time","cctk_time"))
# Step 1f: Create directories for the thorn if they don't exist.
!mkdir FishboneMoncriefID 2>/dev/null # 2>/dev/null: Don't throw an error if the directory already exists.
!mkdir FishboneMoncriefID/src 2>/dev/null # 2>/dev/null: Don't throw an error if the directory already exists.
# Step 1g: Write the C code kernel to file.
with open("FishboneMoncriefID/src/KerrSchild.h", "w") as file:
file.write(str(KerrSchild_CcodeKernel.replace("time","cctk_time")))
with open("FishboneMoncriefID/src/FMdisk_GRHD_velocities.h", "w") as file:
file.write(str(FMdisk_GRHD_velocities_CcodeKernel.replace("time","cctk_time")))
with open("FishboneMoncriefID/src/FMdisk_GRHD_rho_initial.h", "w") as file:
file.write(str(FMdisk_GRHD_rho_initial_CcodeKernel.replace("time","cctk_time")))
hm1string = outputC(fmid.hm1,"hm1",filename="returnstring")
with open("FishboneMoncriefID/src/FMdisk_GRHD_hm1.h", "w") as file:
file.write(str(hm1string))
```
<a id='einstein'></a>
# Step 2: Interfacing with the Einstein Toolkit \[Back to [top](#toc)\]
$$\label{einstein}$$
<a id='einstein_c'></a>
## Step 2.a: Constructing the Einstein Toolkit C-code calling functions that include the C code kernels \[Back to [top](#toc)\]
$$\label{einstein_c}$$
We will write another C file with the functions we need here.
```
%%writefile FishboneMoncriefID/src/InitialData.c
#include <math.h>
#include <stdio.h>
#include <stdbool.h>
#include <stdlib.h> // Needed for rand()
#include "cctk.h"
#include "cctk_Parameters.h"
#include "cctk_Arguments.h"
// Alias for "vel" vector gridfunction:
#define velx (&vel[0*cctk_lsh[0]*cctk_lsh[1]*cctk_lsh[2]])
#define vely (&vel[1*cctk_lsh[0]*cctk_lsh[1]*cctk_lsh[2]])
#define velz (&vel[2*cctk_lsh[0]*cctk_lsh[1]*cctk_lsh[2]])
void FishboneMoncrief_KerrSchild(const cGH* restrict const cctkGH,const CCTK_INT *cctk_lsh,
const CCTK_INT i0,const CCTK_INT i1,const CCTK_INT i2,
const CCTK_REAL *xcoordGF,const CCTK_REAL *ycoordGF,const CCTK_REAL *zcoordGF,
CCTK_REAL *alphaGF,CCTK_REAL *betaU0GF,CCTK_REAL *betaU1GF,CCTK_REAL *betaU2GF,
CCTK_REAL *gammaDD00GF,CCTK_REAL *gammaDD01GF,CCTK_REAL *gammaDD02GF,CCTK_REAL *gammaDD11GF,CCTK_REAL *gammaDD12GF,CCTK_REAL *gammaDD22GF,
CCTK_REAL *KDD00GF,CCTK_REAL *KDD01GF,CCTK_REAL *KDD02GF,CCTK_REAL *KDD11GF,CCTK_REAL *KDD12GF,CCTK_REAL *KDD22GF)
{
DECLARE_CCTK_PARAMETERS
#include "KerrSchild.h"
}
void FishboneMoncrief_FMdisk_GRHD_velocities(const cGH* restrict const cctkGH,const CCTK_INT *cctk_lsh,
const CCTK_INT i0,const CCTK_INT i1,const CCTK_INT i2,
const CCTK_REAL *xcoordGF,const CCTK_REAL *ycoordGF,const CCTK_REAL *zcoordGF,
CCTK_REAL *Valencia3velocityU0GF, CCTK_REAL *Valencia3velocityU1GF, CCTK_REAL *Valencia3velocityU2GF)
{
DECLARE_CCTK_PARAMETERS
#include "FMdisk_GRHD_velocities.h"
}
void FishboneMoncrief_ET_GRHD_initial(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
CCTK_VINFO("Fishbone-Moncrief Disk Initial data.");
CCTK_VINFO("Using input parameters of\n a = %e,\n M = %e,\nr_in = %e,\nr_at_max_density = %e\nkappa = %e\ngamma = %e",a,M,r_in,r_at_max_density,kappa,gamma);
// First compute maximum density
CCTK_REAL rho_max;
{
CCTK_REAL hm1;
CCTK_REAL xcoord = r_at_max_density;
CCTK_REAL ycoord = 0.0;
CCTK_REAL zcoord = 0.0;
{
#include "FMdisk_GRHD_hm1.h"
}
rho_max = pow( hm1 * (gamma-1.0) / (kappa*gamma), 1.0/(gamma-1.0) );
}
#pragma omp parallel for
for(CCTK_INT k=0;k<cctk_lsh[2];k++) for(CCTK_INT j=0;j<cctk_lsh[1];j++) for(CCTK_INT i=0;i<cctk_lsh[0];i++) {
CCTK_INT idx = CCTK_GFINDEX3D(cctkGH,i,j,k);
CCTK_REAL xcoord = x[idx];
CCTK_REAL ycoord = y[idx];
CCTK_REAL zcoord = z[idx];
CCTK_REAL rr = r[idx];
FishboneMoncrief_KerrSchild(cctkGH,cctk_lsh,
i,j,k,
x,y,z,
alp,betax,betay,betaz,
gxx,gxy,gxz,gyy,gyz,gzz,
kxx,kxy,kxz,kyy,kyz,kzz);
CCTK_REAL hm1;
bool set_to_atmosphere=false;
if(rr > r_in) {
{
#include "FMdisk_GRHD_hm1.h"
}
if(hm1 > 0) {
rho[idx] = pow( hm1 * (gamma-1.0) / (kappa*gamma), 1.0/(gamma-1.0) ) / rho_max;
press[idx] = kappa*pow(rho[idx], gamma);
// P = (\Gamma - 1) rho epsilon
eps[idx] = press[idx] / (rho[idx] * (gamma - 1.0));
FishboneMoncrief_FMdisk_GRHD_velocities(cctkGH,cctk_lsh,
i,j,k,
x,y,z,
velx,vely,velz);
} else {
set_to_atmosphere=true;
}
} else {
set_to_atmosphere=true;
}
// Outside the disk? Set to atmosphere all hydrodynamic variables!
if(set_to_atmosphere) {
// Choose an atmosphere such that
// rho = 1e-5 * r^(-3/2), and
// P = k rho^gamma
// Add 1e-100 or 1e-300 to rr or rho to avoid divisions by zero.
rho[idx] = 1e-5 * pow(rr + 1e-100,-3.0/2.0);
press[idx] = kappa*pow(rho[idx], gamma);
eps[idx] = press[idx] / ((rho[idx] + 1e-300) * (gamma - 1.0));
w_lorentz[idx] = 1.0;
velx[idx] = 0.0;
vely[idx] = 0.0;
velz[idx] = 0.0;
}
}
CCTK_INT final_idx = CCTK_GFINDEX3D(cctkGH,cctk_lsh[0]-1,cctk_lsh[1]-1,cctk_lsh[2]-1);
CCTK_VINFO("===== OUTPUTS =====");
CCTK_VINFO("betai: %e %e %e \ngij: %e %e %e %e %e %e \nKij: %e %e %e %e %e %e\nalp: %e\n",betax[final_idx],betay[final_idx],betaz[final_idx],gxx[final_idx],gxy[final_idx],gxz[final_idx],gyy[final_idx],gyz[final_idx],gzz[final_idx],kxx[final_idx],kxy[final_idx],kxz[final_idx],kyy[final_idx],kyz[final_idx],kzz[final_idx],alp[final_idx]);
CCTK_VINFO("rho: %.15e\nPressure: %.15e\nvx: %.15e\nvy: %.15e\nvz: %.15e",rho[final_idx],press[final_idx],velx[final_idx],vely[final_idx],velz[final_idx]);
}
void FishboneMoncrief_ET_GRHD_initial__perturb_pressure(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
for(CCTK_INT k=0;k<cctk_lsh[2];k++) for(CCTK_INT j=0;j<cctk_lsh[1];j++) for(CCTK_INT i=0;i<cctk_lsh[0];i++) {
CCTK_INT idx = CCTK_GFINDEX3D(cctkGH,i,j,k);
// Generate random number in range [0,1),
// snippet courtesy http://daviddeley.com/random/crandom.htm
CCTK_REAL random_number_between_0_and_1 = ( (double)rand() / ((double)(RAND_MAX)+(double)(1)) );
CCTK_REAL random_number_between_min_and_max = random_min + (random_max - random_min)*random_number_between_0_and_1;
press[idx] = press[idx]*(1.0 + random_number_between_min_and_max);
// Add 1e-300 to rho to avoid division by zero when density is zero.
eps[idx] = press[idx] / ((rho[idx] + 1e-300) * (gamma - 1.0));
}
}
```
<a id='einstein_ccl'></a>
## Step 2.b: CCL files - Define how this module interacts and interfaces with the larger Einstein Toolkit infrastructure \[Back to [top](#toc)\]
$$\label{einstein_ccl}$$
Writing a module ("thorn") within the Einstein Toolkit requires that three "ccl" files be constructed, all in the root directory of the thorn:
1. `interface.ccl}`: defines the gridfunction groups needed, and provides keywords denoting what this thorn provides and what it should inherit from other thorns. Specifically, this file governs the interaction between this thorn and others; more information can be found in the [official Einstein Toolkit documentation](http://cactuscode.org/documentation/referencemanual/ReferenceManualch8.html#x12-260000C2.2).
With "implements", we give our thorn its unique name. By "inheriting" other thorns, we tell the Toolkit that we will rely on variables that exist and are declared "public" within those functions.
```
%%writefile FishboneMoncriefID/interface.ccl
implements: FishboneMoncriefID
inherits: admbase grid hydrobase
```
2. `param.ccl`: specifies free parameters within the thorn, enabling them to be set at runtime. It is required to provide allowed ranges and default values for each parameter. More information on this file's syntax can be found in the [official Einstein Toolkit documentation](http://cactuscode.org/documentation/referencemanual/ReferenceManualch8.html#x12-265000C2.3).
```
%%writefile FishboneMoncriefID/param.ccl
shares: grid
shares: ADMBase
USES CCTK_INT lapse_timelevels
USES CCTK_INT shift_timelevels
USES CCTK_INT metric_timelevels
USES KEYWORD metric_type
EXTENDS KEYWORD initial_data
{
"FishboneMoncriefID" :: "Initial data from FishboneMoncriefID solution"
}
EXTENDS KEYWORD initial_lapse
{
"FishboneMoncriefID" :: "Initial lapse from FishboneMoncriefID solution"
}
EXTENDS KEYWORD initial_shift
{
"FishboneMoncriefID" :: "Initial shift from FishboneMoncriefID solution"
}
EXTENDS KEYWORD initial_dtlapse
{
"FishboneMoncriefID" :: "Initial dtlapse from FishboneMoncriefID solution"
}
EXTENDS KEYWORD initial_dtshift
{
"FishboneMoncriefID" :: "Initial dtshift from FishboneMoncriefID solution"
}
shares: HydroBase
EXTENDS KEYWORD initial_hydro
{
"FishboneMoncriefID" :: "Initial GRHD data from FishboneMoncriefID solution"
}
#["r_in","r_at_max_density","a","M"] A_b, kappa, gamma
restricted:
CCTK_REAL r_in "Fixes the inner edge of the disk"
{
0.0:* :: "Must be positive"
} 6.0
restricted:
CCTK_REAL r_at_max_density "Radius at maximum disk density. Needs to be > r_in"
{
0.0:* :: "Must be positive"
} 12.0
restricted:
CCTK_REAL a "The spin parameter of the black hole"
{
0:1.0 :: "Positive values, up to 1. Negative disallowed, as certain roots are chosen in the hydro fields setup. Check those before enabling negative spins!"
} 0.9375
restricted:
CCTK_REAL M "Kerr-Schild BH mass. Probably should always set M=1."
{
0.0:* :: "Must be positive"
} 1.0
restricted:
CCTK_REAL A_b "Scaling factor for the vector potential"
{
*:* :: ""
} 1.0
restricted:
CCTK_REAL kappa "Equation of state: P = kappa * rho^gamma"
{
0.0:* :: "Positive values"
} 1.0e-3
restricted:
CCTK_REAL gamma "Equation of state: P = kappa * rho^gamma"
{
0.0:* :: "Positive values"
} 1.3333333333333333333333333333
##################################
# PRESSURE PERTURBATION PARAMETERS
private:
CCTK_REAL random_min "Floor value of random perturbation to initial pressure, where perturbed pressure = pressure*(1.0 + (random_min + (random_max-random_min)*RAND[0,1)))"
{
*:* :: "Any value"
} -0.02
private:
CCTK_REAL random_max "Ceiling value of random perturbation to initial pressure, where perturbed pressure = pressure*(1.0 + (random_min + (random_max-random_min)*RAND[0,1)))"
{
*:* :: "Any value"
} 0.02
```
3. `schedule.ccl`: allocates storage for gridfunctions, defines how the thorn's functions should be scheduled in a broader simulation, and specifies the regions of memory written to or read from gridfunctions. $\text{schedule.ccl}$'s official documentation may be found [here](http://cactuscode.org/documentation/referencemanual/ReferenceManualch8.html#x12-268000C2.4).
We specify here the standardized ETK "scheduling bins" in which we want each of our thorn's functions to run.
```
%%writefile FishboneMoncriefID/schedule.ccl
STORAGE: ADMBase::metric[metric_timelevels], ADMBase::curv[metric_timelevels], ADMBase::lapse[lapse_timelevels], ADMBase::shift[shift_timelevels]
schedule FishboneMoncrief_ET_GRHD_initial IN HydroBase_Initial
{
LANG: C
READS: grid::x(Everywhere)
READS: grid::y(Everywhere)
READS: grid::y(Everywhere)
WRITES: admbase::alp(Everywhere)
WRITES: admbase::betax(Everywhere)
WRITES: admbase::betay(Everywhere)
WRITES: admbase::betaz(Everywhere)
WRITES: admbase::kxx(Everywhere)
WRITES: admbase::kxy(Everywhere)
WRITES: admbase::kxz(Everywhere)
WRITES: admbase::kyy(Everywhere)
WRITES: admbase::kyz(Everywhere)
WRITES: admbase::kzz(Everywhere)
WRITES: admbase::gxx(Everywhere)
WRITES: admbase::gxy(Everywhere)
WRITES: admbase::gxz(Everywhere)
WRITES: admbase::gyy(Everywhere)
WRITES: admbase::gyz(Everywhere)
WRITES: admbase::gzz(Everywhere)
WRITES: hydrobase::velx(Everywhere)
WRITES: hydrobase::vely(Everywhere)
WRITES: hydrobase::velz(Everywhere)
WRITES: hydrobase::rho(Everywhere)
WRITES: hydrobase::eps(Everywhere)
WRITES: hydrobase::press(Everywhere)
} "Set up general relativistic hydrodynamic (GRHD) fields for Fishbone-Moncrief disk"
schedule FishboneMoncrief_ET_GRHD_initial__perturb_pressure IN CCTK_INITIAL AFTER Seed_Magnetic_Fields BEFORE IllinoisGRMHD_ID_Converter
{
LANG: C
} "Add random perturbation to initial pressure, after seed magnetic fields have been set up (in case we'd like the seed magnetic fields to depend on the pristine pressures)"
```
<a id='einstein_list'></a>
## Step 2.c: Add the C code to the Einstein Toolkit compilation list \[Back to [top](#toc)\]
$$\label{einstein_list}$$
We will also need `make.code.defn`, which indicates the list of files that need to be compiled. This thorn only has the one C file to compile.
```
%%writefile FishboneMoncriefID/src/make.code.defn
SRCS = InitialData.c
```
<a id='latex_pdf_output'></a>
# Step 3: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-ETK_thorn-FishboneMoncriefID.pdf](Tutorial-ETK_thorn-FishboneMoncriefID.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
!jupyter nbconvert --to latex --template latex_nrpy_style.tplx --log-level='WARN' Tutorial-ETK_thorn-FishboneMoncriefID.ipynb
!pdflatex -interaction=batchmode Tutorial-ETK_thorn-FishboneMoncriefID.tex
!pdflatex -interaction=batchmode Tutorial-ETK_thorn-FishboneMoncriefID.tex
!pdflatex -interaction=batchmode Tutorial-ETK_thorn-FishboneMoncriefID.tex
!rm -f Tut*.out Tut*.aux Tut*.log
```
| github_jupyter |
# Developing, Training, and Deploying a TensorFlow model on Google Cloud Platform (completely within Jupyter)
In this notebook, we will develop a Keras model to predict flight delays using TensorFlow 2.0 as the backend.
### Make sure you are using compatible pip and TensorFlow 2.0
You need pip 19.1 or higher and TensorFlow 2.0 post Aug 2019. If not, run the necessary update scripts and check the versions again.
```
%pip --version
import tensorflow as tf
print(tf.version.VERSION)
%pip install --user --upgrade --quiet pip
%pip install --user --upgrade --quiet tf_nightly-2.0-preview
```
### Setup
```
# change these to try this notebook out
# In "production", these will be replaced by the parameters passed to papermill
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
DEVELOP_MODE = True
NBUCKETS = 5 # for embeddings
NUM_EXAMPLES = 1000*1000 # assume 1 million examples
TRAIN_BATCH_SIZE = 64
DNN_HIDDEN_UNITS = '64,32'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
%%bash
gcloud config set project $PROJECT
gcloud config set compute/region $REGION
```
## Creating the input data pipeline
```
DATA_BUCKET = "gs://{}/flights/chapter8/output/".format(BUCKET)
TRAIN_DATA_PATTERN = DATA_BUCKET + "train*"
EVAL_DATA_PATTERN = DATA_BUCKET + "test*"
!gsutil ls $DATA_BUCKET
```
### Use tf.data to read the CSV files
```
import os, json, math, shutil
import numpy as np
import tensorflow as tf
print("Tensorflow version " + tf.__version__)
CSV_COLUMNS = ('ontime,dep_delay,taxiout,distance,avg_dep_delay,avg_arr_delay' + \
',carrier,dep_lat,dep_lon,arr_lat,arr_lon,origin,dest').split(',')
LABEL_COLUMN = 'ontime'
DEFAULTS = [[0.0],[0.0],[0.0],[0.0],[0.0],[0.0],\
['na'],[0.0],[0.0],[0.0],[0.0],['na'],['na']]
def load_dataset(pattern, batch_size=1):
return tf.data.experimental.make_csv_dataset(pattern, batch_size, CSV_COLUMNS, DEFAULTS)
if DEVELOP_MODE:
dataset = load_dataset(TRAIN_DATA_PATTERN)
for n, data in enumerate(dataset):
numpy_data = {k: v.numpy() for k, v in data.items()} # .numpy() works only in eager mode
print(numpy_data)
if n>3: break
def features_and_labels(features):
label = features.pop('ontime') # this is what we will train for
return features, label
def read_dataset(pattern, batch_size, mode=tf.estimator.ModeKeys.TRAIN, truncate=None):
dataset = load_dataset(pattern, batch_size)
dataset = dataset.map(features_and_labels)
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.shuffle(batch_size*10)
dataset = dataset.repeat()
dataset = dataset.prefetch(1)
if truncate is not None:
dataset = dataset.take(truncate)
return dataset
if DEVELOP_MODE:
print("Checking input pipeline")
one_item = read_dataset(TRAIN_DATA_PATTERN, batch_size=2, truncate=1)
print(list(one_item)) # should print one batch of 2 items
```
## Create TensorFlow wide-and-deep model
We'll create feature columns, and do some discretization and feature engineering.
See the book for details.
```
import tensorflow as tf
real = {
colname : tf.feature_column.numeric_column(colname)
for colname in
('dep_delay,taxiout,distance,avg_dep_delay,avg_arr_delay' +
',dep_lat,dep_lon,arr_lat,arr_lon').split(',')
}
sparse = {
'carrier': tf.feature_column.categorical_column_with_vocabulary_list('carrier',
vocabulary_list='AS,VX,F9,UA,US,WN,HA,EV,MQ,DL,OO,B6,NK,AA'.split(',')),
'origin' : tf.feature_column.categorical_column_with_hash_bucket('origin', hash_bucket_size=1000),
'dest' : tf.feature_column.categorical_column_with_hash_bucket('dest', hash_bucket_size=1000)
}
inputs = {
colname : tf.keras.layers.Input(name=colname, shape=(), dtype='float32')
for colname in real.keys()
}
inputs.update({
colname : tf.keras.layers.Input(name=colname, shape=(), dtype='string')
for colname in sparse.keys()
})
```
### Feature engineering
```
latbuckets = np.linspace(20.0, 50.0, NBUCKETS).tolist() # USA
lonbuckets = np.linspace(-120.0, -70.0, NBUCKETS).tolist() # USA
disc = {}
disc.update({
'd_{}'.format(key) : tf.feature_column.bucketized_column(real[key], latbuckets)
for key in ['dep_lat', 'arr_lat']
})
disc.update({
'd_{}'.format(key) : tf.feature_column.bucketized_column(real[key], lonbuckets)
for key in ['dep_lon', 'arr_lon']
})
# cross columns that make sense in combination
sparse['dep_loc'] = tf.feature_column.crossed_column([disc['d_dep_lat'], disc['d_dep_lon']], NBUCKETS*NBUCKETS)
sparse['arr_loc'] = tf.feature_column.crossed_column([disc['d_arr_lat'], disc['d_arr_lon']], NBUCKETS*NBUCKETS)
sparse['dep_arr'] = tf.feature_column.crossed_column([sparse['dep_loc'], sparse['arr_loc']], NBUCKETS ** 4)
#sparse['ori_dest'] = tf.feature_column.crossed_column(['origin', 'dest'], hash_bucket_size=1000)
# embed all the sparse columns
embed = {
'embed_{}'.format(colname) : tf.feature_column.embedding_column(col, 10)
for colname, col in sparse.items()
}
real.update(embed)
# one-hot encode the sparse columns
sparse = {
colname : tf.feature_column.indicator_column(col)
for colname, col in sparse.items()
}
if DEVELOP_MODE:
print(sparse.keys())
print(real.keys())
```
## Train the model and evaluate once in a while
Also checkpoint
```
output_dir='gs://{}/flights/trained_model'.format(BUCKET)
os.environ['OUTDIR'] = output_dir # needed for deployment
print('Writing trained model to {}'.format(output_dir))
!gsutil -m rm -rf $OUTDIR
# Build a wide-and-deep model.
def wide_and_deep_classifier(inputs, linear_feature_columns, dnn_feature_columns, dnn_hidden_units):
deep = tf.keras.layers.DenseFeatures(dnn_feature_columns, name='deep_inputs')(inputs)
layers = [int(x) for x in dnn_hidden_units.split(',')]
for layerno, numnodes in enumerate(layers):
deep = tf.keras.layers.Dense(numnodes, activation='relu', name='dnn_{}'.format(layerno+1))(deep)
wide = tf.keras.layers.DenseFeatures(linear_feature_columns, name='wide_inputs')(inputs)
both = tf.keras.layers.concatenate([deep, wide], name='both')
output = tf.keras.layers.Dense(1, activation='sigmoid', name='pred')(both)
model = tf.keras.Model(inputs, output)
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
return model
model = wide_and_deep_classifier(
inputs,
linear_feature_columns = sparse.values(),
dnn_feature_columns = real.values(),
dnn_hidden_units = DNN_HIDDEN_UNITS)
tf.keras.utils.plot_model(model, 'flights_model.png', show_shapes=False, rankdir='LR')
# training and evaluation dataset
train_batch_size = TRAIN_BATCH_SIZE
if DEVELOP_MODE:
eval_batch_size = 100
steps_per_epoch = 3
epochs = 2
else:
eval_batch_size = 10000
steps_per_epoch = NUM_EXAMPLES // train_batch_size
epochs = 10
train_dataset = prepare_dataset(TRAIN_DATA_PATTERN, train_batch_size)
eval_dataset = prepare_dataset(EVAL_DATA_PATTERN, eval_batch_size, tf.estimator.ModeKeys.EVAL, eval_batch_size*10)
checkpoint_path = '{}/checkpoints/flights.cpt'.format(output_dir)
shutil.rmtree(checkpoint_path, ignore_errors=True)
cp_callback = tf.keras.callbacks.ModelCheckpoint(checkpoint_path,
save_weights_only=True,
verbose=1)
history = model.fit(train_dataset,
validation_data=eval_dataset,
epochs=epochs,
steps_per_epoch=steps_per_epoch,
callbacks=[cp_callback])
print(history.history.keys())
import matplotlib.pyplot as plt
nrows = 1
ncols = 2
fig = plt.figure(figsize=(10, 5))
for idx, key in enumerate(['loss', 'accuracy']):
ax = fig.add_subplot(nrows, ncols, idx+1)
plt.plot(history.history[key])
plt.plot(history.history['val_{}'.format(key)])
plt.title('model {}'.format(key))
plt.ylabel(key)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left');
```
## Export and deploy the trained model
```
import time
export_dir = '{}/export/flights_{}'.format(output_dir, time.strftime("%Y%m%d-%H%M%S"))
print('Exporting to {}'.format(export_dir))
tf.saved_model.save(model, export_dir)
%%bash
model_dir=$(gsutil ls ${OUTDIR}/export | tail -1)
echo $model_dir
saved_model_cli show --tag_set serve --signature_def serving_default --dir $model_dir
%%bash
PROJECT=cloud-training-demos
BUCKET=${PROJECT}-ml
REGION=us-east1
MODEL_NAME=flights
VERSION_NAME=kfp
EXPORT_PATH=$(gsutil ls ${OUTDIR}/export | tail -1)
if [[ $(gcloud ai-platform models list --format='value(name)' | grep $MODEL_NAME) ]]; then
echo "$MODEL_NAME already exists"
else
# create model
echo "Creating $MODEL_NAME"
gcloud ai-platform models create --regions=$REGION $MODEL_NAME
fi
if [[ $(gcloud ai-platform versions list --model $MODEL_NAME --format='value(name)' | grep $VERSION_NAME) ]]; then
echo "Deleting already existing $MODEL_NAME:$VERSION_NAME ... "
gcloud ai-platform versions delete --model=$MODEL_NAME $VERSION_NAME
echo "Please run this cell again if you don't see a Creating message ... "
sleep 10
fi
# create model
echo "Creating $MODEL_NAME:$VERSION_NAME"
gcloud ai-platform versions create --model=$MODEL_NAME $VERSION_NAME --async \
--framework=tensorflow --python-version=3.5 --runtime-version=1.14 \
--origin=$EXPORT_PATH --staging-bucket=gs://$BUCKET
!gcloud ai-platform predict --model=flights --version=kfp --json-instances=example_input.json
```
Copyright 2016 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
```
import os
import pandas as pd
from scipy.integrate import odeint
from scipy.interpolate import interp1d
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import time
import matplotlib.pyplot as plt
import sys
sys.path.append("..")
from symbolic_RLC import fxu_ODE, fxu_ODE_mod, A_nominal, B_nominal
from torchid.ssfitter import NeuralODE
from torchid.ssmodels import StateSpaceModelLin, NeuralStateSpaceModelLin
Ts = 2e-7
A_lin = A_nominal * Ts
B_lin = B_nominal * Ts
A_lin = np.array([[10, 100], [1000, 10000]])
#ss_model = StateSpaceModelLin(A_lin, B_lin)
ss_model = NeuralStateSpaceModelLin(A_lin, B_lin)
# In[Linearization time - 1 a time]
nx = 2
nu = 1
# Linearization variable
VAR = []
for idx_var in range(nx):
var = np.zeros((1,nx)).astype(np.float32)
var[0,idx_var] = 1.0 # differentiate w.r.t the nth variable
VAR.append(torch.tensor(var))
# Random linearization point
x_arr = np.random.rand(nx).astype(np.float32)
x_torch = torch.tensor(x_arr, requires_grad=True)
u_batch = np.random.rand(nu).astype(np.float32)
u_torch = torch.tensor(u_batch, requires_grad=True)
%%timeit -n 1000
# Linearization around the random point
F_xu = ss_model(x_torch,u_torch)
A = np.empty((nx,nx))
B = np.empty((nx,nu))
for idx_var in range(nx):
var = VAR[idx_var]
#var = np.zeros((1,nx)).astype(np.float32)
#var[0,idx_var] = 1.0 # differentiate w.r.t the nth variable
F_xu.backward(var, retain_graph=True)
A[idx_var,:] = np.array(x_torch.grad)
B[idx_var,:] = np.array(u_torch.grad)
x_torch.grad.data.zero_()
u_torch.grad.data.zero_()
# Linearization around the random point
F_xu = ss_model(x_torch,u_torch)
A = np.empty((nx,nx))
B = np.empty((nx,nu))
for idx_var in range(nx):
var = VAR[idx_var]
F_xu.backward(var, retain_graph=True)
A[idx_var,:] = np.array(x_torch.grad)
B[idx_var,:] = np.array(u_torch.grad)
x_torch.grad.data.zero_()
u_torch.grad.data.zero_()
print(A)
print(B)
#A[0,0]-10
# Batch linearization variable
batch_size = 128
VAR = []
for idx_var in range(nx):
var = np.zeros((batch_size,nx)).astype(np.float32)
var[:,idx_var] = 1.0 # differentiate w.r.t the nth variable
VAR.append(torch.tensor(var))
# Random batch of linearization point
x_batch = np.random.rand(batch_size, nx).astype(np.float32)
x_torch = torch.tensor(x_batch, requires_grad=True)
u_batch = np.random.rand(batch_size, nu).astype(np.float32)
u_torch = torch.tensor(u_batch, requires_grad=True)
%%timeit -n 1000
# In[Linearization time - batched]
F_xu = ss_model(x_torch,u_torch)
A = np.empty((batch_size,nx,nx))
B = np.empty((batch_size,nx,nu))
for idx_var in range(nx):
var = VAR[idx_var]
F_xu.backward(var, retain_graph=True)
A[:,idx_var,:] = np.array(x_torch.grad)
B[:,idx_var,:] = np.array(u_torch.grad)
x_torch.grad.data.zero_()
u_torch.grad.data.zero_()
F_xu = ss_model(x_torch,u_torch)
A = np.empty((batch_size,nx,nx))
B = np.empty((batch_size,nx,nu))
for idx_var in range(nx):
var = VAR[idx_var]
F_xu.backward(var, retain_graph=True)
A[:,idx_var,:] = np.array(x_torch.grad)
B[:,idx_var,:] = np.array(u_torch.grad)
x_torch.grad.data.zero_()
u_torch.grad.data.zero_()
A
```
| github_jupyter |
# Inference Example
#### Table of Contents
<a id='toc'></a>
- [Setup](#setup)
- [Imputation](#imputation)
- [Create Features/Labels](#createfeatures)
- [Logistic Regression](#logreg)
- [Machine Learning](#machinelearning)
- [Adding Group Membership](#groupmember)
- [Logistic with Group Feature](#grouplog)
- [ML with Group Features](#groupml)
During the lecture today we discussed different factors that can affect inference. As a result of this notebook you will learn how a "grouping" variable can impact the fit of a model. We'll also be reviewing some of the ML code you saw during the last session.
The dataset used in this notebook is class2.for_inference_example and was developed for this purpose. It is a subset of household benefit spells reported as ending within 2013. There are over 200k rows in the dataset.
Our goal with this analysis is to predict individuals returning to benefits within 1 year after the end of a benefit spell. Variables that will be used in this analysis are listed below. The majority of the features will be what we will consider "individual variables," and we'll consider the *district* variable a group variable. We will first run models with the individual features, and then see if the inclusion of the group variable changes our model output/prediction.
We will first look at the more familiar Logistic Regression, then at ML models we learned in the last class session.
## Important Variables
---
### Identification Variables
* receptno = IDHS provided receipt number
* ch_dpa_caseid = Chapin Hall Case ID number
* new_id = unique row id number created for this dataset
* start_date
* end_date
### Features/Grouping Variables
*note that variables from the case records may have missingness*
* benefit_type
* sex
* rac
* rootrace
* foreignbrn
* edlevel
* health
* martlst
* workexp
* district
* homeless
#### Features Developed From Wage Table
* has_job_win1yr = 0/1 indicating any wage table employment within one year
* lose_job_win1yr = 0/1 indicating that had wage table employment and then did not within one year
* has_job_q(1-4) = 0/1 variable for each of the 4 quarters in the year after end_date indicating employment
* wage_q(1-4) = wage total for each quarter (could be from multiple jobs)
* total_wage_1yr = sum of all the wages within the year following end_date
### Labels/Outcome Variables
* new_spell_win1yr = 0/1 variable indicating a new spell within one year of end date of spell
* new_spell_win1yr_benefit = same as above, but has to be same benefit_type (tanf, foodstamps, etc.)
# Setup
<a id='setup'></a>
- Return to [Table of Contents](#toc)
```
%pylab inline
import pandas as pd
import numpy as np
import psycopg2
import psycopg2.extras
import sklearn
import seaborn as sns
from sklearn.cross_validation import train_test_split
from sklearn.ensemble import (RandomForestClassifier, ExtraTreesClassifier,
GradientBoostingClassifier,
AdaBoostClassifier)
from sklearn.linear_model import LogisticRegression, SGDClassifier
from sklearn.metrics import precision_recall_curve, auc
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
import sqlalchemy
sns.set_style("white")
db_name = "appliedda"
hostname = "10.10.2.10"
conn = psycopg2.connect(database=db_name, host = hostname) #database connection
#load some of Avishek's function defintions to help with model comparison
def plot_precision_recall_n(y_true, y_prob, model_name):
"""
y_true: ls
ls of ground truth labels
y_prob: ls
ls of predic proba from model
model_name: str
str of model name (e.g, LR_123)
"""
from sklearn.metrics import precision_recall_curve
y_score = y_prob
precision_curve, recall_curve, pr_thresholds = precision_recall_curve(y_true, y_score)
precision_curve = precision_curve[:-1]
recall_curve = recall_curve[:-1]
pct_above_per_thresh = []
number_scored = len(y_score)
for value in pr_thresholds:
num_above_thresh = len(y_score[y_score>=value])
pct_above_thresh = num_above_thresh / float(number_scored)
pct_above_per_thresh.append(pct_above_thresh)
pct_above_per_thresh = np.array(pct_above_per_thresh)
plt.clf()
fig, ax1 = plt.subplots()
ax1.plot(pct_above_per_thresh, precision_curve, 'b')
ax1.set_xlabel('percent of population')
ax1.set_ylabel('precision', color='b')
ax1.set_ylim(0,1.05)
ax2 = ax1.twinx()
ax2.plot(pct_above_per_thresh, recall_curve, 'r')
ax2.set_ylabel('recall', color='r')
ax2.set_ylim(0,1.05)
name = model_name
plt.title(name)
plt.show()
plt.clf()
def precision_at_k(y_true, y_scores,k):
threshold = np.sort(y_scores)[::-1][int(k*len(y_scores))]
y_pred = np.asarray([1 if i >= threshold else 0 for i in y_scores ])
return precision_score(y_true, y_pred)
select_statement = """SELECT new_id, sex, rootrace, edlevel, workexp, martlst, homeless, benefit_type,
has_job_q1, has_job_q2, has_job_q3, has_job_q4, wage_q1, wage_q2, wage_q3, wage_q4,
has_job_win1yr, lose_job_win1yr, total_wage_1yr, new_spell_win1yr, new_spell_win1yr_benefit,
district FROM class2.for_inference_example WHERE total_wage_1yr IS NOT NULL;"""
df = pd.read_sql( select_statement, conn )
print df.shape
df.head()
```
## Imputation
<a id='imputation'></a>
- Return to [Table of Contents](#toc)
We can see from just the few rows above that there is clearly missing data. Because our variables are categorical, we cannot simply impute the mean for the missing values. Instead, we will add a 0/1 variable for missing in each variable that will be another feature/predictor used in the model.
```
df['sex_miss'] = (df['sex'] == 0)
df['race_miss'] = (df['rootrace'] == 0)
df['ed_miss'] = (df['edlevel'] == None)
df['mar_miss'] = (df['martlst'] == 0)
df['home_miss'] = (df['homeless'] == None)
df.head()
```
## Create Features
<a id='createfeatures'></a>
- Return to [Table of Contents](#toc)
We need to create 0/1 "dummy" variables/features for the rest of the different levels of the predictors we want to include in our model. At the end of this code block we set up two different feature tables - one with the yearly employment features and the other with the quarterly employment features.
```
#sex
df['male'] = (df['sex'] == 1)
df['female'] = (df['sex'] == 2)
#rootrace
df['nhwhite'] = (df['rootrace'] == 1)
df['nhblack'] = (df['rootrace'] == 2)
df['native'] = (df['rootrace'] == 3)
df['hispanic'] = (df['rootrace'] == 6)
df['asian'] = (df['rootrace'] == 7)
#edlevel
less_list = ['A', 'B', 'C', 'D', 1, 2, 3]
somehs_list = ['E', 'F', 4]
hsgrad_list = ['G', 'H', 'V', 5]
somecoll_list = ['W', 'X', 'Y', 6]
collgrad_list = ['Z', 'P', 7]
df['lessthanhs'] = (df['edlevel'].isin(less_list))
df['somehs'] = (df['edlevel'].isin(somehs_list))
df['hsgrad'] = (df['edlevel'].isin(hsgrad_list))
df['somecoll'] = (df['edlevel'].isin(somecoll_list))
df['collgrad'] = (df['edlevel'].isin(collgrad_list))
#workexp
df['noattach'] = (df['workexp'] == 0)
df['nowkexp'] = (df['workexp'] == 1)
df['prof'] = (df['workexp'] == 2)
df['othermgr'] = (df['workexp'] == 3)
df['clerical'] = (df['workexp'] == 4)
df['sales'] = (df['workexp'] == 5)
df['crafts'] = (df['workexp'] == 6)
df['oper'] = (df['workexp'] == 7)
df['service'] = (df['workexp'] == 8)
df['labor'] = (df['workexp'] == 9)
#martlst
df['nvrmar'] = (df['martlst'] == 1)
df['marwspouse'] = (df['martlst'] == 2)
df['marwospouse'] = (df['martlst'].isin([3,4,6]))
df['sepordiv'] = (df['martlst'].isin([5,7]))
df['widow'] = (df['martlst'] == 8)
#homeless
df['nothomeless'] = (df['homeless'] == 'N')
df['ishomeless'] = (df['homeless'].isin(['1','2','3','4','Y']))
#benefit_type
df['foodstamp'] = (df['benefit_type'] == 'foodstamp')
df['tanf'] = (df['benefit_type'] == 'tanf46')
df['grant'] = (df['benefit_type'] == 'grant')
#create features df
df_features = df[['male', 'female', 'nhwhite', 'nhblack', 'native', 'hispanic', 'asian', 'lessthanhs',
'somehs', 'hsgrad', 'somecoll', 'collgrad', 'noattach', 'nowkexp', 'prof', 'othermgr',
'clerical', 'sales', 'crafts', 'oper', 'service', 'labor', 'nvrmar', 'marwspouse',
'sepordiv', 'widow', 'nothomeless', 'ishomeless', 'foodstamp', 'tanf', 'grant']].copy()
# features df with qtr based job variables
df_features_wjobqtr = df[['has_job_q1', 'has_job_q2', 'has_job_q3', 'has_job_q4',
'wage_q1', 'wage_q2', 'wage_q3', 'wage_q4']].copy()
df_features_wjobqtr.join(df_features)
# features df with year based job variables
df_features_wjobyr = df[['has_job_win1yr', 'lose_job_win1yr', 'total_wage_1yr']].copy()
df_features_wjobyr = df_features_wjobyr.join(df_features)
```
### Create Labels
The labels dataframes each have just one column, the outcome variable for your model. Below we've created two, one for returns to any benefit within the year after a spell, the other is an indicator for return to the same benefit within the year. We'll be using the first in the examples below.
```
df_label_returnany = df[['new_spell_win1yr']].copy()
df_label_returnsame = df[['new_spell_win1yr_benefit']].copy()
```
### Start with a more familiar model - Logistic Regression
<a id='logreg'></a>
- Return to [Table of Contents](#toc)
We'll first fit a logistic regression model to predict return to benefits based on the individual predictors we have selected. Typically when a social scientist uses this type of model it is to describe the individual predictors' effects on some outcome, therefore the entire dataset is used to fit the model.
Unlike in the ML class, we will be using the package *statsmodels* to run our Logistic Regression models, since it provides many more output options/functions vs. sci-kit learn.
```
import statsmodels.api as sm
import statsmodels.formula.api as smf
# create one df and set up listings of variables for below
#drop reference categories (missing, male)
df_features_forlr = df_features_wjobyr[['female', 'nhwhite', 'nhblack', 'native', 'hispanic', 'asian', 'lessthanhs',
'somehs', 'hsgrad', 'somecoll', 'collgrad', 'noattach', 'nowkexp', 'prof', 'othermgr',
'clerical', 'sales', 'crafts', 'oper', 'service', 'labor', 'nvrmar', 'marwspouse',
'sepordiv', 'widow', 'nothomeless', 'ishomeless', 'foodstamp', 'tanf', 'grant',
'has_job_win1yr', 'lose_job_win1yr', 'total_wage_1yr']].copy()
df_lrmodel = df_features_forlr.join(df_label_returnany)
```
*statsmodels* takes as an argument a string with your model specification, in a format you may be familiar with if you have used R in the past. In order to build that list I'll employ some code to avoid typing all the variable names.
```
#get list of features to build model statement
feat_list = list(df_features_forlr)
print feat_list
print len(feat_list)
length = len(feat_list)
#create string of features with plus signs
count = 0
feat_string = ''
for feature in feat_list:
count += 1
if count < (length):
feat_string += feature
feat_string += ' + '
else:
feat_string += feature
## END FOR BLOCK
print feat_string
formula = "new_spell_win1yr ~ " + feat_string
print (formula)
#fit model - note the procedure is glm so you have to specify binomial in the family argument to get LR
model =smf.glm(formula=formula, data=df_lrmodel, family=sm.families.Binomial())
result = model.fit()
print (result.summary())
```
It's no surprise that the majority of the coefficients are significant - with Ns this large significance becomes pretty meaningless. Looking at the direction of the coefficients, we start to see which variables impact the probability of going back on benefits either positive or negatively. For female and nhblack (Non-Hispanic Blacks) we see they are more likely to return to benefits (reference values are male and missing-race). On the other hand nhwhite (Non-Hispanic Whites) are less likely to return.
This package also has the benefit of returning a model output summary familiar to any SAS or stata user. We can also get just the values using .params or .pvalues (for example result.params contains just the coefficients for this model
```
print (result.params)
```
However, we have run this model how a statistician would typically - using all of the data to fit the model. In our ML notebook we learned about using training and test sets of data to show the strength of the model in predicting a subsequent outcome. This can also be done with the logistic regression as we saw in the ML notebook. Below we'll work through it in *statsmodels* with our same data from above. So we can cut our dataframe into train and test sets. Here I'll use the index (row numbers) to cut the dataframe into roughly 80/20 train/test.
```
df_lrtrain = df_lrmodel[:201540]
df_lrtest = df_lrmodel[201540:]
print df_lrtrain.shape
print df_lrtest.shape
#fit model - this time we're only fitting the model to the training set.
model =smf.glm(formula=formula, data=df_lrtrain, family=sm.families.Binomial())
result = model.fit()
print (result.summary())
```
This is fitting the model to just `%%%` of the data that we used above, however, as to be expected, the resulting coefficients are largely similar.
Compared to the first model, we get:
- female - this model coefficient = `####`, first model coefficient = `####`
- nhwhite - this model coefficient = `####`, first model coefficient = `####`
- nhblack - this model coefficient = `####`, first model coefficient = `####`
Any coefficients with larger changes between this model (like native which went from `####` to `####`) is because we cut data out of our dataset in a not exactly random way and those are rarer instances.
Now we can use these coefficients to make predictions on the test set and see how well our model works for prediction.
```
from sklearn.metrics import confusion_matrix, classification_report
predictions = result.predict(df_lrtest)
pred_binary = (predictions > 0.5)
print confusion_matrix(df_lrtest['new_spell_win1yr'], pred_binary )
print classification_report(df_lrtest['new_spell_win1yr'], pred_binary, digits=3 )
plot_precision_recall_n(df_lrtest['new_spell_win1yr'], predictions, "Logistic Regression")
```
## Moving to Machine Learning
<a id='machinelearning'></a>
- Return to [Table of Contents](#toc)
### Create Testing/Training sets of both Features and Labels sets
We're going to start with using the features table with the yearly job variables and the label that indicates a return to any form of benefits.
This is the same code as we used in the ML notebook.
```
# create train/test sets
X_train, X_test, y_train, y_test = train_test_split(df_features_wjobyr, df_label_returnany, test_size = 0.2)
print X_train.shape, y_train.shape
print X_test.shape, y_test.shape
sel_features = list(X_train)
```
### Running through the ML models
Below is again code familiar from the ML notebook. We set up our arguments in a dictionary so that we can loop through the models and print the results using the same code.
Again we're using the full set of features to predict returning to benefits within 1 year of the end of a spell.
```
clfs = {'RF': RandomForestClassifier(n_estimators=50, n_jobs=-1),
'ET': ExtraTreesClassifier(n_estimators=10, n_jobs=-1, criterion='entropy'),
'LR': LogisticRegression(penalty='l1', C=1e5),
'SGD':SGDClassifier(loss='log'),
'GB': GradientBoostingClassifier(learning_rate=0.05, subsample=0.5, max_depth=6, random_state=17, n_estimators=10),
'NB': GaussianNB()}
sel_clfs = ['RF', 'ET', 'LR', 'SGD', 'GB', 'NB']
max_p_at_k = 0
for clfNM in sel_clfs:
clf = clfs[clfNM]
clf.fit( X_train, y_train )
print clf
y_score = clf.predict_proba(X_test)[:,1]
predicted = np.array(y_score)
expected = np.array(y_test)
plot_precision_recall_n(expected,predicted, clfNM)
p_at_1 = precision_at_k(expected,y_score, 0.05) ## note that i changed the k value here from 0.01 to 0.05
print('Precision at 5%: {:.2f}'.format(p_at_1))
```
To summarize, the Precision at %5 values for the 6 models are:
* Random Forest: XX
* Extra Trees: XX
* Logistic Regression: XX
* SGD Classifier: XX
* Gradient Boosting: XX
* GaussianNB: XX
To get the best prediction for our money, we'll probably want to choose a model with high precision at 5% (the level of funding we can afford).
Focusing on the ML models with highest precision at %5, we'll look at the feature importances for these two ML models (Random Forest and Gradient Boosting).
Feature importances indicate which variables are most important to the prediction/classification of cases as the trees split. Features at earlier/higher splits are more important than those at lower levels in the tree.
```
#select only the two ML models
sel_clfs = ['RF', 'GB']
#here I've adapted the model loop from above to print feature importances instead of the precision/recall graph
for clfNM in sel_clfs:
clf = clfs[clfNM]
clf.fit( X_train, y_train )
print clf
y_score = clf.predict_proba(X_test)[:,1]
predicted = np.array(y_score)
expected = np.array(y_test)
var_names = list(X_train) # get a list of variable names
importances = clf.feature_importances_ # get the feature importances
indices = np.argsort(importances)[::-1] # sort the list to get the highest importance first
for f in range(X_train.shape[1]):
print ("%d. feature (%s) importance = %f" % (f + 1, var_names[indices[f]], importances[indices[f]]))
p_at_1 = precision_at_k(expected,y_score, 0.05)
print('Precision at 5%: {:.2f}'.format(p_at_1))
print
```
The most important features in the Random Forest Model are:
1. total_wage_1yr
2. nothomeless
3. nvrmar
4. hsgrad
5. noattach
This makes sense - wages and homelessness status likely have a huge impact on whether a person returns to benefits within the year or not.
The most important features in the Gradient Boosting Model are:
1. nothomeless
2. nvrmar
3. total_wage_1yr
4. foodstamp
5. hsgrad
Again we see most of the same variables. The slight differences are due to the different ways in which these models are processing the same data.
# Will Adding a Group Membership Variable make a difference?
<a id='groupmember'></a>
- Return to [Table of Contents](#toc)
We'll add in the district variable and see if it makes a difference in either our prediction or our coefficients/feature importances. Adding this variable could improve prediction to the extent that it is new information not already contained within the other features. It can also change the coefficients in a logistic model or the feature importances in a ML model if there are clusters of individuals in the groups who are alike on that particular feature. For example, if there are more individuals of a particular race collected in one of the groups, not including the group variable meant that the race variable was partially serving as "proxy" for the group membership. Adding the group membership to the model lets you see the effect of race above and beyond the clustering.
We will use the *district* variable and split it into two groupings - downstate (codes 10-115) and Cook County (codes 200-294).
```
df['cookcty'] = ((df['district'] >= 200) & (df['district'] <= 294))
df['downstate'] = ((df['district'] >= 10) & (df['district'] <= 115))
```
### Return to Logistic Regression
<a id='grouplog'></a>
- Return to [Table of Contents](#toc)
Adding in our new variable.
```
# add new cookcty to df for logistic regression
df_lrmodel['cookcty'] = df['cookcty']
df_lrtrain = df_lrmodel[:201540]
df_lrtest = df_lrmodel[201540:]
print df_lrtrain.shape
print df_lrtest.shape
formula += " + cookcty"
model =smf.glm(formula=formula, data=df_lrtrain, family=sm.families.Binomial())
result = model.fit()
print (result.summary())
```
Compared to our previous LR model, we get:
- female - previous model coefficient = XX, this model coefficient = XX
- nhwhite - prevous model coefficient = XX, this model coefficient = XX
- nhblack - previous model coefficient = XX, this model coefficient = XX
So adding the group variable did account for some of the race effects we were seeing in the previous model.
We also see cookcty has a coefficient of XX, reflecting the power that group membership has on the prediction
```
from sklearn.metrics import confusion_matrix, classification_report
predictions = result.predict(df_lrtest)
pred_binary = (predictions > 0.5)
print confusion_matrix(df_lrtest['new_spell_win1yr'], pred_binary )
print classification_report(df_lrtest['new_spell_win1yr'], pred_binary, digits=3 )
plot_precision_recall_n(df_lrtest['new_spell_win1yr'], predictions, "Logistic Regression")
```
The slight improvement to the precision is reflected in the graph above.
### Will ML Models be changed?
<a id='groupml'></a>
- Return to [Table of Contents](#toc)
We saw an impact of adding this grouping variable in the Logistic Regression. Will it improve the predictive power of our ML models? Will it be an important feature?
None of our model fitting code is changed below - we're just adding two new features and rerunning.
```
df_features_wjobyr['cookcty'] = df['cookcty']
df_features_wjobyr['downstate'] =df['downstate']
X_train, X_test, y_train, y_test = train_test_split(df_features_wjobyr, df_label_returnany, test_size = 0.2)
print X_train.shape, y_train.shape
print X_test.shape, y_test.shape
sel_features = list(X_train)
sel_clfs = ['RF', 'ET', 'LR', 'SGD', 'GB', 'NB']
max_p_at_k = 0
for clfNM in sel_clfs:
clf = clfs[clfNM]
clf.fit( X_train, y_train )
print clf
y_score = clf.predict_proba(X_test)[:,1]
predicted = np.array(y_score)
expected = np.array(y_test)
plot_precision_recall_n(expected,predicted, clfNM)
p_at_1 = precision_at_k(expected,y_score, 0.05)
print('Precision at 5%: {:.2f}'.format(p_at_1))
```
The Precision at %5 did not substantially improve with any of the models. And the prediction power of the SGD model was decreased.
* Random Forest: both XX
* Extra Trees: both XX
* Logistic Regression: now XX, was XX
* SGD Classifier: now XX, was XX
* Gradient Boosting: now XX, was XX
* GaussianNB: now XX, was XX
We'll again look at the feature importances to see if there are any substantial changes.
```
sel_clfs = ['RF', 'GB']
for clfNM in sel_clfs:
clf = clfs[clfNM]
clf.fit( X_train, y_train )
print clf
y_score = clf.predict_proba(X_test)[:,1]
predicted = np.array(y_score)
expected = np.array(y_test)
var_names = list(X_train)
importances = clf.feature_importances_
indices = np.argsort(importances)[::-1]
for f in range(X_train.shape[1]):
print ("%d. feature (%s) importance = %f" % (f + 1, var_names[indices[f]], importances[indices[f]]))
p_at_1 = precision_at_k(expected,y_score, 0.05)
print('Precision at 5%: {:.2f}'.format(p_at_1))
print
```
Our most important features in each model are largely unchanged. *cookcty* and *downstate* show up as 16th and 21st most important features in the Random Forest model, respectively. In the Gradient Boosing model, though, *cookcty* shows up 5th, indicating it was an important classifier, however the resulting output slightly lowered our precision at 5%.
| github_jupyter |
# Simulation of Sequences
```
from pulser import Pulse, Sequence, Register, Simulation
from pulser.waveforms import BlackmanWaveform, RampWaveform
from pulser.devices import MockDevice
import numpy as np
import qutip
import matplotlib.pyplot as plt
```
To illustrate the simulation of sequences, let us study a simple one-dimensional system with periodic boundary conditions (a ring of atoms):
```
# Setup
L = 14
Omega_max = 2.3 * 2*np.pi
U = Omega_max / 2.3
delta_0 = -3 * U
delta_f = 1 * U
t_rise = 2000
t_fall = 2000
t_sweep = (delta_f - delta_0)/(2 * np.pi * 10) * 5000
# Define a ring of atoms distanced by a blockade radius distance:
R_interatomic = MockDevice.rydberg_blockade_radius(U)
coords = R_interatomic/(2*np.tan(np.pi/L)) * np.array([(np.cos(theta*2*np.pi/L), np.sin(theta*2*np.pi/L)) for theta in range(L)])
reg = Register.from_coordinates(coords, prefix='atom')
reg.draw(blockade_radius=R_interatomic, draw_half_radius=True, draw_graph = True)
```
We use the drawing capabilites of the `Register` class to highlight the area **half** the blockade radius away from each atom, which makes it so that overlapping circles correspond to interacting atoms. This is further fleshed out by the graph edges drawn using the `draw_graph` option.
In this register, we shall act with the following pulser sequence, which is designed to reach a state with *antiferromagnetic order*:
```
rise = Pulse.ConstantDetuning(RampWaveform(t_rise, 0., Omega_max), delta_0, 0.)
sweep = Pulse.ConstantAmplitude(Omega_max, RampWaveform(t_sweep, delta_0, delta_f), 0.)
fall = Pulse.ConstantDetuning(RampWaveform(t_fall, Omega_max, 0.), delta_f, 0.)
seq = Sequence(reg, MockDevice)
seq.declare_channel('ising', 'rydberg_global')
seq.add(rise, 'ising')
seq.add(sweep, 'ising')
seq.add(fall, 'ising')
seq.draw()
```
## 1. Running a Simulation
First we define our `Simulation` object, which creates an internal respresentation of the quantum system, including the Hamiltonian which will drive the evolution:
```
sim = Simulation(seq, sampling_rate=0.01)
```
Notice we have included the parameter `sampling_rate` which allows us to determine how many samples from the pulse sequence we wish to simulate. In the case of the simple shapes in our sequence, only a very small fraction is needed. This largely accelerates the simulation time in the solver.
To run the simulation we simply apply the method `run()`. At the time of writing of this notebook, the method uses a series of routines from **QuTiP** for solving the Schröedinger equation of the system. It returns a `SimulationResults` object, which will allow the study or post-processing of the states for each time step in our simulation. Additionally, we can include a progress bar to have an estimate of how the simulation is advancing:
```
results = sim.run(progress_bar=True)
```
## 2. Using the `SimulationResults` object
The `SimulationResults` object that we created contains the quantum state at each time step. We can call them using the `states` attribute:
```
results.states[23] # Given as a `qutip.Qobj` object
```
We can sample the final state directly, using the `sample_final_state()` method from the `SimulationResults` object. We try it with $1000$ samples and discard the less frequent bitstrings:
```
counts = results.sample_final_state(meas_basis='ground-rydberg', N_samples=1000)
large_counts = {k:v for k,v in counts.items() if v > 5}
plt.figure(figsize=(15,4))
plt.xticks(rotation=90, fontsize=14)
plt.title("Most frequent observations")
plt.bar(large_counts.keys(), large_counts.values())
```
Notice how the most frequent bitstrings correspond to the antiferromagnetic order states.
We can also compute the expectation values of operators for the states in the evolution, using the `expect()` method, which takes a list of operators (in this case, the local magnetization acting on the $j$-th spin):
```
def magnetization(j, total_sites):
prod = [qutip.qeye(2) for _ in range(total_sites)]
prod[j] = qutip.sigmaz()
return qutip.tensor(prod)
magn_list = [magnetization(j, L) for j in range(L)]
expect_magnetization = results.expect(magn_list)
for data in expect_magnetization:
plt.plot(sim._times, data)
```
Notice how the local magnetization on *each* atom goes in the same way from $-1$ (which corresponds to the ground state) to $0$. This is expected since as we saw above, the state after the evolution has antiferromagnetic-order, so at each site, there is a compensation of magnetization. The parity (even) and the boundary conditions (periodic) allow for two lowest-energy states, whose superposition is similar to that of the perfectly antiferromagnetic state: $\Big(|grgr\cdots \rangle + |rgrg\cdots \rangle\Big)/\sqrt{2}$
| github_jupyter |
```
def normalize(x):
"""utility function to normalize a tensor.
# Arguments
x: An input tensor.
# Returns
The normalized input tensor.
"""
return x / (K.sqrt(K.mean(K.square(x))) + K.epsilon())
def deprocess_image(x):
"""utility function to convert a float array into a valid uint8 image.
# Arguments
x: A numpy-array representing the generated image.
# Returns
A processed numpy-array, which could be used in e.g. imshow.
"""
# normalize tensor: center on 0., ensure std is 0.25
x -= x.mean()
x /= (x.std() + K.epsilon())
x *= 0.25
# clip to [0, 1]
x += 0.5
x = np.clip(x, 0, 1)
# convert to RGB array
x *= 255
if K.image_data_format() == 'channels_first':
x = x.transpose((1, 2, 0))
x = np.clip(x, 0, 255).astype('uint8')
return x
def process_image(x, former):
"""utility function to convert a valid uint8 image back into a float array.
Reverses `deprocess_image`.
# Arguments
x: A numpy-array, which could be used in e.g. imshow.
former: The former numpy-array.
Need to determine the former mean and variance.
# Returns
A processed numpy-array representing the generated image.
"""
if K.image_data_format() == 'channels_first':
x = x.transpose((2, 0, 1))
return (x / 255 - 0.5) * 4 * former.std() + former.mean()
def visualize_layer(model,
layer_name,
step=1.,
epochs=15,
upscaling_steps=9,
upscaling_factor=1.2,
output_dim=(412, 412),
filter_range=(0, None)):
"""Visualizes the most relevant filters of one conv-layer in a certain model.
# Arguments
model: The model containing layer_name.
layer_name: The name of the layer to be visualized.
Has to be a part of model.
step: step size for gradient ascent.
epochs: Number of iterations for gradient ascent.
upscaling_steps: Number of upscaling steps.
Starting image is in this case (80, 80).
upscaling_factor: Factor to which to slowly upgrade
the image towards output_dim.
output_dim: [img_width, img_height] The output image dimensions.
filter_range: Tupel[lower, upper]
Determines the to be computed filter numbers.
If the second value is `None`,
the last filter will be inferred as the upper boundary.
"""
def _generate_filter_image(input_img,
layer_output,
filter_index):
"""Generates image for one particular filter.
# Arguments
input_img: The input-image Tensor.
layer_output: The output-image Tensor.
filter_index: The to be processed filter number.
Assumed to be valid.
#Returns
Either None if no image could be generated.
or a tuple of the image (array) itself and the last loss.
"""
s_time = time.time()
# we build a loss function that maximizes the activation
# of the nth filter of the layer considered
if K.image_data_format() == 'channels_first':
loss = K.mean(layer_output[:, filter_index, :, :])
else:
loss = K.mean(layer_output[:, :, :, filter_index])
# we compute the gradient of the input picture wrt this loss
grads = K.gradients(loss, input_img)[0]
# normalization trick: we normalize the gradient
grads = normalize(grads)
# this function returns the loss and grads given the input picture
iterate = K.function([input_img], [loss, grads])
# we start from a gray image with some random noise
intermediate_dim = tuple(
int(x / (upscaling_factor ** upscaling_steps)) for x in output_dim)
if K.image_data_format() == 'channels_first':
input_img_data = np.random.random(
(1, 3, intermediate_dim[0], intermediate_dim[1]))
else:
input_img_data = np.random.random(
(1, intermediate_dim[0], intermediate_dim[1], 3))
input_img_data = (input_img_data - 0.5) * 20 + 128
# Slowly upscaling towards the original size prevents
# a dominating high-frequency of the to visualized structure
# as it would occur if we directly compute the 412d-image.
# Behaves as a better starting point for each following dimension
# and therefore avoids poor local minima
for up in reversed(range(upscaling_steps)):
# we run gradient ascent for e.g. 20 steps
for _ in range(epochs):
loss_value, grads_value = iterate([input_img_data])
input_img_data += grads_value * step
# some filters get stuck to 0, we can skip them
if loss_value <= K.epsilon():
return None
# Calculate upscaled dimension
intermediate_dim = tuple(
int(x / (upscaling_factor ** up)) for x in output_dim)
# Upscale
img = deprocess_image(input_img_data[0])
img = np.array(pil_image.fromarray(img).resize(intermediate_dim,
pil_image.BICUBIC))
input_img_data = np.expand_dims(
process_image(img, input_img_data[0]), 0)
# decode the resulting input image
img = deprocess_image(input_img_data[0])
e_time = time.time()
print('Costs of filter {:3}: {:5.0f} ( {:4.2f}s )'.format(filter_index,
loss_value,
e_time - s_time))
return img, loss_value
def _draw_filters(filters, n=None):
"""Draw the best filters in a nxn grid.
# Arguments
filters: A List of generated images and their corresponding losses
for each processed filter.
n: dimension of the grid.
If none, the largest possible square will be used
"""
if n is None:
n = int(np.floor(np.sqrt(len(filters))))
# the filters that have the highest loss are assumed to be better-looking.
# we will only keep the top n*n filters.
filters.sort(key=lambda x: x[1], reverse=True)
filters = filters[:n * n]
# build a black picture with enough space for
# e.g. our 8 x 8 filters of size 412 x 412, with a 5px margin in between
MARGIN = 5
width = n * output_dim[0] + (n - 1) * MARGIN
height = n * output_dim[1] + (n - 1) * MARGIN
stitched_filters = np.zeros((width, height, 3), dtype='uint8')
# fill the picture with our saved filters
for i in range(n):
for j in range(n):
img, _ = filters[i * n + j]
width_margin = (output_dim[0] + MARGIN) * i
height_margin = (output_dim[1] + MARGIN) * j
stitched_filters[
width_margin: width_margin + output_dim[0],
height_margin: height_margin + output_dim[1], :] = img
# save the result to disk
save_img('vgg_{0:}_{1:}x{1:}.png'.format(layer_name, n), stitched_filters)
# this is the placeholder for the input images
assert len(model.inputs) == 1
input_img = model.inputs[0]
# get the symbolic outputs of each "key" layer (we gave them unique names).
layer_dict = dict([(layer.name, layer) for layer in model.layers[1:]])
output_layer = layer_dict[layer_name]
assert isinstance(output_layer, layers.Conv2D)
# Compute to be processed filter range
filter_lower = filter_range[0]
filter_upper = (filter_range[1]
if filter_range[1] is not None
else len(output_layer.get_weights()[1]))
assert(filter_lower >= 0
and filter_upper <= len(output_layer.get_weights()[1])
and filter_upper > filter_lower)
print('Compute filters {:} to {:}'.format(filter_lower, filter_upper))
# iterate through each filter and generate its corresponding image
processed_filters = []
for f in range(filter_lower, filter_upper):
img_loss = _generate_filter_image(input_img, output_layer.output, f)
if img_loss is not None:
processed_filters.append(img_loss)
print('{} filter processed.'.format(len(processed_filters)))
# Finally draw and store the best filters to disk
_draw_filters(processed_filters)
"""
#Visualization of the filters of VGG16, via gradient ascent in input space.
This script can run on CPU in a few minutes.
Results example: 
"""
from __future__ import print_function
import time
import numpy as np
from PIL import Image as pil_image
from keras.preprocessing.image import save_img
from keras import layers
from keras.applications import vgg16
from keras import backend as K
print("libraries imported")
if __name__ == '__main__':
# the name of the layer we want to visualize
# (see model definition at keras/applications/vgg16.py)
LAYER_NAME = 'block5_conv1'
# build the VGG16 network with ImageNet weights
vgg = vgg16.VGG16(weights='imagenet', include_top=False)
print('Model loaded.')
vgg.summary()
# example function call
visualize_layer(vgg, LAYER_NAME)
```
| github_jupyter |
```
import sys
import warnings
if not sys.warnoptions:
warnings.simplefilter('ignore')
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from datetime import datetime
from datetime import timedelta
from tqdm import tqdm
sns.set()
tf.compat.v1.random.set_random_seed(1234)
df = pd.read_csv('../dataset/GOOG-year.csv')
df.head()
minmax = MinMaxScaler().fit(df.iloc[:, 4:5].astype('float32')) # Close index
df_log = minmax.transform(df.iloc[:, 4:5].astype('float32')) # Close index
df_log = pd.DataFrame(df_log)
df_log.head()
```
## Split train and test
I will cut the dataset to train and test datasets,
1. Train dataset derived from starting timestamp until last 30 days
2. Test dataset derived from last 30 days until end of the dataset
So we will let the model do forecasting based on last 30 hours, and we will going to repeat the experiment for 10 times. You can increase it locally if you want, and tuning parameters will help you by a lot.
```
test_size = 30
simulation_size = 10
df_train = df_log.iloc[:-test_size]
df_test = df_log.iloc[-test_size:]
df.shape, df_train.shape, df_test.shape
class Model:
def __init__(
self,
learning_rate,
num_layers,
size,
size_layer,
output_size,
forget_bias = 0.1,
):
def lstm_cell(size_layer):
return tf.nn.rnn_cell.GRUCell(size_layer)
rnn_cells = tf.nn.rnn_cell.MultiRNNCell(
[lstm_cell(size_layer) for _ in range(num_layers)],
state_is_tuple = False,
)
self.X = tf.placeholder(tf.float32, (None, None, size))
self.Y = tf.placeholder(tf.float32, (None, output_size))
drop = tf.contrib.rnn.DropoutWrapper(
rnn_cells, output_keep_prob = forget_bias
)
self.hidden_layer = tf.placeholder(
tf.float32, (None, num_layers * size_layer)
)
self.outputs, self.last_state = tf.nn.dynamic_rnn(
drop, self.X, initial_state = self.hidden_layer, dtype = tf.float32
)
self.logits = tf.layers.dense(self.outputs[-1], output_size)
self.cost = tf.reduce_mean(tf.square(self.Y - self.logits))
self.optimizer = tf.train.AdamOptimizer(learning_rate).minimize(
self.cost
)
def calculate_accuracy(real, predict):
real = np.array(real) + 1
predict = np.array(predict) + 1
percentage = 1 - np.sqrt(np.mean(np.square((real - predict) / real)))
return percentage * 100
def anchor(signal, weight):
buffer = []
last = signal[0]
for i in signal:
smoothed_val = last * weight + (1 - weight) * i
buffer.append(smoothed_val)
last = smoothed_val
return buffer
num_layers = 1
size_layer = 128
timestamp = 5
epoch = 300
dropout_rate = 0.8
future_day = test_size
learning_rate = 0.01
def forecast():
tf.reset_default_graph()
modelnn = Model(
learning_rate, num_layers, df_log.shape[1], size_layer, df_log.shape[1], dropout_rate
)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
date_ori = pd.to_datetime(df.iloc[:, 0]).tolist()
pbar = tqdm(range(epoch), desc = 'train loop')
for i in pbar:
init_value = np.zeros((1, num_layers * size_layer))
total_loss, total_acc = [], []
for k in range(0, df_train.shape[0] - 1, timestamp):
index = min(k + timestamp, df_train.shape[0] - 1)
batch_x = np.expand_dims(
df_train.iloc[k : index, :].values, axis = 0
)
batch_y = df_train.iloc[k + 1 : index + 1, :].values
logits, last_state, _, loss = sess.run(
[modelnn.logits, modelnn.last_state, modelnn.optimizer, modelnn.cost],
feed_dict = {
modelnn.X: batch_x,
modelnn.Y: batch_y,
modelnn.hidden_layer: init_value,
},
)
init_value = last_state
total_loss.append(loss)
total_acc.append(calculate_accuracy(batch_y[:, 0], logits[:, 0]))
pbar.set_postfix(cost = np.mean(total_loss), acc = np.mean(total_acc))
future_day = test_size
output_predict = np.zeros((df_train.shape[0] + future_day, df_train.shape[1]))
output_predict[0] = df_train.iloc[0]
upper_b = (df_train.shape[0] // timestamp) * timestamp
init_value = np.zeros((1, num_layers * size_layer))
for k in range(0, (df_train.shape[0] // timestamp) * timestamp, timestamp):
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(
df_train.iloc[k : k + timestamp], axis = 0
),
modelnn.hidden_layer: init_value,
},
)
init_value = last_state
output_predict[k + 1 : k + timestamp + 1] = out_logits
if upper_b != df_train.shape[0]:
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(df_train.iloc[upper_b:], axis = 0),
modelnn.hidden_layer: init_value,
},
)
output_predict[upper_b + 1 : df_train.shape[0] + 1] = out_logits
future_day -= 1
date_ori.append(date_ori[-1] + timedelta(days = 1))
init_value = last_state
for i in range(future_day):
o = output_predict[-future_day - timestamp + i:-future_day + i]
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(o, axis = 0),
modelnn.hidden_layer: init_value,
},
)
init_value = last_state
output_predict[-future_day + i] = out_logits[-1]
date_ori.append(date_ori[-1] + timedelta(days = 1))
output_predict = minmax.inverse_transform(output_predict)
deep_future = anchor(output_predict[:, 0], 0.3)
return deep_future[-test_size:]
results = []
for i in range(simulation_size):
print('simulation %d'%(i + 1))
results.append(forecast())
accuracies = [calculate_accuracy(df['Close'].iloc[-test_size:].values, r) for r in results]
plt.figure(figsize = (15, 5))
for no, r in enumerate(results):
plt.plot(r, label = 'forecast %d'%(no + 1))
plt.plot(df['Close'].iloc[-test_size:].values, label = 'true trend', c = 'black')
plt.legend()
plt.title('average accuracy: %.4f'%(np.mean(accuracies)))
plt.show()
```
| github_jupyter |
# Enrichment of TFs and CSMs among DE genes (Fig 7)
```
from __future__ import division
import sys
import random
import copy
import math
import json
import numpy as np
import pandas as pd
import scipy
%matplotlib inline
from matplotlib import pyplot as plt
import matplotlib as mpl
import seaborn as sns
sys.path.append("../resources/")
import sct
reload(sct)
sns.set_style("ticks")
sns.set_context("talk")
output_dir = "out/"
output_suffix = ""
output_formats = [".png", ".pdf"]
def save_figure(fig, name):
for output_format in output_formats:
fig.savefig(output_dir + "/" + name + output_suffix + output_format)
return None
mpl.rc('savefig', dpi=300)
pd.options.mode.chained_assignment = None # default='warn'
```
# Load data
```
# Gene expression
df = pd.read_csv("../data/df_GH146_Fig2.csv.gz", header=0, index_col=0)
# Cluster labels
df_labels = pd.read_csv("../data/labels_HDBSCAN.csv", sep="\t", header=0, index_col=0)
# Load names of TFs and CSM genes
def load_list(infile):
X = []
with open(infile) as f:
for line in f:
X.append(line.rstrip())
return X
def write_list(X, outfile):
with open(outfile, 'w') as out:
for x in X:
out.write(str(x) + "\n")
CSMs = load_list("../resources/CSMs.txt")
TFs = load_list("../resources/TFs.txt")
genes_genome_noTFs_noCSMs = list(set(list(df.index)) - set(TFs) - set(CSMs))
```
# Calculate and plot enrichment of TFs, CSMs among top DE genes between pairs of clusters
```
# Results of differential expression analysis between each pair of clusters
import pickle
import gzip
dfs_DE_labels = pickle.load(gzip.open("../data/dfs_DE_labels.pickle.gz"))
def calc_frac_top_index_in_list(df, n, L):
# Returns fraction of top indexes that are in the list
x = len(set(df.head(n=n).index).intersection(L))
return x / float(df.head(n=n).shape[0])
# Aggregate data into dataframe
ranks = []
frac_TFs = []
frac_CSMs = []
labels = []
for num_top_genes in [30, 50, 100, 200, 500, 1000]:
for label, df_DE in dfs_DE_labels.items():
x = df_DE.sort_values("pvalue")
my_frac_TFs = calc_frac_top_index_in_list(x, num_top_genes, TFs)
my_frac_CSMs = calc_frac_top_index_in_list(x, num_top_genes, CSMs)
frac_TFs.append(my_frac_TFs)
frac_CSMs.append(my_frac_CSMs)
labels.append(label)
ranks.extend([num_top_genes]*len(dfs_DE_labels.keys()))
df_DE_gene_enrichment = pd.DataFrame({"rank": ranks,
"labels": labels,
"frac_TFs": frac_TFs,
"frac_CSMs": frac_CSMs})
frac_CSMs_genome = len(CSMs)/float(15525)
frac_TFs_genome = len(TFs)/float(15525)
df_DE_gene_enrichment["fold_enrichment_CSMs"] = df_DE_gene_enrichment["frac_CSMs"] / frac_CSMs_genome
df_DE_gene_enrichment["fold_enrichment_TFs"] = df_DE_gene_enrichment["frac_TFs"] / frac_TFs_genome
# Plot as violins
fig, ax = plt.subplots(1, 1, figsize=(8,4))
sns.violinplot(x="rank", y="fold_enrichment_CSMs", ax=ax, data=df_DE_gene_enrichment,
cut=0, scale="width", palette="Oranges")
ax.axhline(1, color="k")
ax.set_xlabel("Top differentially expressed genes between classes")
ax.set_ylabel("Fold enrichment of CSMs")
fig, ax = plt.subplots(1, 1, figsize=(8,4))
sns.violinplot(x="rank", y="fold_enrichment_TFs", ax=ax, data=df_DE_gene_enrichment,
cut=0, scale="width", palette="Greens")
ax.axhline(1, color="k")
ax.set_xlabel("Top differentially expressed genes between classes")
ax.set_ylabel("Fold enrichment of TFs")
# Report fraction of genes in genome that are TFs or CSMs
print "CSMs", len(CSMs)
print "Genes in genome", 15525
print "% CSMs in genome", len(CSMs)/float(15525) * 100
print
print "TFs", len(TFs)
print "Genes in genome", 15525
print "% TFs in genome", len(TFs)/float(15525) * 100
```
| github_jupyter |
# The following are the parameter spaces used for evaluations
## The parameter search space used by Tuner for xgboost, KNN and SVM in the Section IV-A2.
```
from scipy.stats import uniform
from mango.domain.distribution import loguniform
param_svm = {'gamma':uniform(0.1, 4),
'C':loguniform(-7, 10)}
param_xgboost = {'learning_rate':uniform(0, 1),
'gamma':uniform(0, 5),
'max_depth':range(1,11),
'n_estimators':range(1,301),
'booster':['gbtree','gblinear','dart']}
param_knn = {'n_neighbors':range(1, 51),
'algorithm':['auto','ball_tree','kd_tree',
'brute']}
```
## The parameter search space used by MetaTuner for neural network, decision tree, xgboost, KNN and SVM in the Section IV-A3. The neural network has one hidden layer with number of hidden nodes specified in the search space.
```
from scipy.stats import uniform
from mango.domain.distribution import loguniform
param_nn = {'type':'clf_nn',
'num_of_nodes':range(10, 101)}
param_dtree = {'type':'clf_dtree',
'max_features':['auto', 'sqrt', 'log2'],
'max_depth':range(1,21),
'splitter':['best','random'],
'criterion':['gini','entropy']}
param_svm = {'type':'clf_svm',
'gamma':uniform(0.1, 4),
'C':loguniform(-7, 10)}
param_xgboost = {'type':'clf_xgboost',
'learning_rate':uniform(0, 1),
'gamma':uniform(0, 5),
'max_depth':range(1,21),
'n_estimators':range(1,11),
'booster':['gbtree','gblinear','dart']}
param_knn = {'type':'clf_knn',
'n_neighbors': range(1, 51),
'algorithm':['auto','ball_tree','kd_tree',
'brute']}
```
# The parameter space used for the TCN network in the Section IV:D Case Study: Network Architecture Search for TinyML Platforms
```
import itertools
import numpy as np
min_layer = 3
max_layer = 8
a_list = [1,2,4,8,16,32,64,128,256]
all_combinations = []
dil_list = []
for r in range(len(a_list) + 1):
combinations_object = itertools.combinations(a_list, r)
combinations_list = list(combinations_object)
all_combinations += combinations_list
all_combinations = all_combinations[1:]
for item in all_combinations:
if(len(item) >= min_layer and len(item) <= max_layer):
dil_list.append(list(item))
param_dict = {
'nb_filters': range(2,64),
'kernel_size': range(2,16),
'dropout_rate': np.arange(0.0,0.5,0.1),
'use_skip_connections': [True, False],
'norm_flag': np.arange(0,1),
'dil_list': dil_list
}
# A sample TCN is constructed using the above space using Keras TCN library as:
def objective_NN(epochs=500,nb_filters=32,kernel_size=7,dilations=[1, 2, 4, 8, 16, 32, 64, 128],dropout_rate=0,
use_skip_connections=False,norm_flag=0):
batch_size, timesteps, input_dim = 256, window_size, 6
i = Input(shape=(timesteps, input_dim))
if(norm_flag==1):
m = TCN(nb_filters=nb_filters,kernel_size=kernel_size,dilations=dilations,dropout_rate=dropout_rate,
use_skip_connections=use_skip_connections,use_batch_norm=True)(i)
else:
m = TCN(nb_filters=nb_filters,kernel_size=kernel_size,dilations=dilations,dropout_rate=dropout_rate,
use_skip_connections=use_skip_connections)(i)
m = tf.reshape(m, [-1, nb_filters, 1])
m = MaxPooling1D(pool_size=(2))(m)
m = Flatten()(m)
m = Dense(32, activation='linear', name='pre')(m)
output1 = Dense(1, activation='linear', name='velx')(m)
output2 = Dense(1, activation='linear', name='vely')(m)
model = Model(inputs=[i], outputs=[output1, output2])
opt = tf.keras.optimizers.Adam()
model.compile(loss={'velx': 'mse','vely':'mse'},optimizer=opt)
return model
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
f, ax = plt.subplots()
```
# Create Figure and Axes
figsize parameters list?
```
f, ax = plt.subplots(figsize=[12, 6])
plt.close(f)
f, ax_array = plt.subplots(1, 4, figsize=[12, 6])
print(ax_array.shape)
plt.close(f)
f, ax_array = plt.subplots(2, 2, figsize=[12, 6])
print(ax_array.shape)
plt.close(f)
```
# Plot
```
def test_plot(ax):
ax.plot([1, 2, 3, 2, 3], 'r--', linewidth=5)
ax.set_title('line plot') # set ax property explicitly
ax.set_xlabel('x', fontsize=24, color='red')
ax.set_ylabel('y', fontsize=24, color='blue')
ax.set_xlim(1, 5)
ax.set_ylim(1, 5)
# ax.axis([1, 5, 1, 5]) # axis range
ax.grid(True)
ax.text(3, 3, r'$\mu=100,\ \sigma=15$')
ax.annotate('local maximum', xy=(2, 3), xytext=(3, 4),
arrowprops=dict(facecolor='black', shrink=0.1),
)
# marker list?
f, ax_array = plt.subplots(2, 2, figsize=[10, 10])
for ax in ax_array.flatten():
test_plot(ax)
f.subplots_adjust(top=0.8, bottom=0.2, left=0.20, right=0.8, hspace=0.2,
wspace=0.35)
```
# Set lines
* use param
* use plt.setp
```
f, ax = plt.subplots()
line = ax.plot([1, 20, 300, 200, 3], 'r--', linewidth=5) # get objects
plt.setp(line, color='blue')
ax.set_title('line plot') # set ax property explicitly
ax.set_xlabel('x', fontsize=24, color='red')
ax.set_ylabel('y', fontsize=24, color='blue')
ax.set_xlim(1, 5)
ax.set_ylim(1, 400)
ax.set_yscale('log')
# ax.axis([1, 5, 1, 5]) # axis range
ax.grid(True)
ax.text(3, 3, r'$\mu=100,\ \sigma=15$')
ax.annotate('local maximum', xy=(2, 3), xytext=(3, 4),
arrowprops=dict(facecolor='black', shrink=0.1),
)
```
# line property
[line property](https://matplotlib.org/tutorials/introductory/pyplot.html#sphx-glr-tutorials-introductory-pyplot-py)
```
plt.plot([1, 2, 3, 4])
plt.ylabel('some numbers')
plt.show()
```
# Coding styles
Use the pyplot interface for creating figures, and then use the object methods for the rest:
```
x = np.arange(0, 10, 0.2)
y = np.sin(x)
fig, ax = plt.subplots()
ax.plot(x, y)
plt.show()
# function to reuse plotting should receive a ax object
# recommended function signature
def my_plotter(ax, data1, data2, param_dict):
pass
# Create ax outside
fig, ax = plt.subplots(1, 1)
my_plotter(ax, data1, data2, {'marker': 'x'})
%matplotlib widget
# Jupyter widget ecosystem
import matplotlib.pyplot as plt
plt.plot([1.6, 2.7])
plt.title("interactive test")
plt.xlabel("index")
ax = plt.gca()
ax.plot([3.1, 2.2])
# close to release memory
mu, sigma = 100, 15
x = mu + sigma * np.random.randn(10000)
# the histogram of the data
n, bins, patches = plt.hist(x, 50, density=1, facecolor='g', alpha=0.75)
plt.xlabel('Smarts')
plt.ylabel('Probability')
plt.title('Histogram of IQ')
plt.text(60, .025, r'$\mu=100,\ \sigma=15$')
plt.axis([40, 160, 0, 0.03])
plt.grid(True)
plt.show()
figure = plt.figure()
ax = plt.subplot(111)
t = np.arange(0.0, 5.0, 0.01)
s = np.cos(2*np.pi*t)
line, = plt.plot(t, s, lw=2)
plt.annotate('local max', xy=(2, 1), xytext=(3, 1.5),
arrowprops=dict(facecolor='black', shrink=0.05),
)
plt.ylim(-2, 2)
plt.show()
```
# Plot Samples
```
t = np.arange(0.0, 2.0, 0.01)
s = 1 + np.sin(2 * np.pi * t)
fig, ax = plt.subplots()
ax.plot(t, s)
ax.set(xlabel='time (s)', ylabel='voltage (mV)',
title='About as simple as it gets, folks')
ax.grid()
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.colors import BoundaryNorm
from matplotlib.ticker import MaxNLocator
import numpy as np
# make these smaller to increase the resolution
dx, dy = 0.05, 0.05
# generate 2 2d grids for the x & y bounds
y, x = np.mgrid[slice(1, 5 + dy, dy),
slice(1, 5 + dx, dx)]
sl = slice(1, 5 + dy, dy)
z = np.sin(x)**10 + np.cos(10 + y*x) * np.cos(x)
z = z[:-1, :-1]
levels = MaxNLocator(nbins=100).tick_values(z.min(), z.max())
# pick the desired colormap, sensible levels, and define a normalization
# instance which takes data values and translates those into levels.
cmap = plt.get_cmap('PiYG')
norm = BoundaryNorm(levels, ncolors=cmap.N, clip=True)
b = BoundaryNorm(levels, ncolors=cmap.N, clip=True)
fig, (ax0, ax1) = plt.subplots(nrows=2)
im = ax0.pcolormesh(x, y, z, cmap=cmap, norm=norm)
fig.colorbar(im, ax=ax0)
ax0.set_title('pcolormesh with levels')
# contours are *point* based plots, so convert our bound into point
# centers
cf = ax1.contourf(x[:-1, :-1] + dx/2.,
y[:-1, :-1] + dy/2., z, levels=levels,
cmap=cmap)
cf = ax1.contourf(x[:-1, :-1].,
y[:-1, :-1] + dy/2., z, levels=levels,
cmap=cmap)
fig.colorbar(cf, ax=ax1)
ax1.set_title('contourf with levels')
# adjust spacing between subplots so `ax1` title and `ax0` tick labels
# don't overlap
fig.tight_layout()
plt.show()
```
## Histogram
```
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(19680801)
# example data
mu = 100 # mean of distribution
sigma = 15 # standard deviation of distribution
x = mu + sigma * np.random.randn(437)
num_bins = 50
fig, ax = plt.subplots()
# the histogram of the data
n, bins, patches = ax.hist(x, num_bins, density=1)
# add a 'best fit' line
y = ((1 / (np.sqrt(2 * np.pi) * sigma)) *
np.exp(-0.5 * (1 / sigma * (bins - mu))**2))
ax.plot(bins, y, '--')
ax.set_xlabel('Smarts')
ax.set_ylabel('Probability density')
ax.set_title(r'Histogram of IQ: $\mu=100$, $\sigma=15$')
# Tweak spacing to prevent clipping of ylabel
fig.tight_layout()
plt.show()
## 3D surface
# This import registers the 3D projection, but is otherwise unused.
from mpl_toolkits.mplot3d import Axes3D # noqa: F401 unused import
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import numpy as np
fig = plt.figure()
ax = fig.gca(projection='3d')
# Make data.
X = np.arange(-5, 5, 0.25)
Y = np.arange(-5, 5, 0.25)
X, Y = np.meshgrid(X, Y)
R = np.sqrt(X**2 + Y**2)
Z = np.sin(R)
# Plot the surface.
surf = ax.plot_surface(X, Y, Z, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
# Customize the z axis.
ax.set_zlim(-1.01, 1.01)
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=10)
plt.show()
```
## Bar
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import MaxNLocator
from collections import namedtuple
n_groups = 5
means_men = (20, 35, 30, 35, 27)
std_men = (2, 3, 4, 1, 2)
means_women = (25, 32, 34, 20, 25)
std_women = (3, 5, 2, 3, 3)
fig, ax = plt.subplots()
index = np.arange(n_groups)
bar_width = 0.35
opacity = 0.4
error_config = {'ecolor': '0.3'}
rects1 = ax.bar(index, means_men, bar_width,
alpha=opacity, color='b',
yerr=std_men, error_kw=error_config,
label='Men')
rects2 = ax.bar(index + bar_width, means_women, bar_width,
alpha=opacity, color='r',
yerr=std_women, error_kw=error_config,
label='Women')
ax.set_xlabel('Group')
ax.set_ylabel('Scores')
ax.set_title('Scores by group and gender')
ax.set_xticks(index + bar_width / 2)
ax.set_xticklabels(('A', 'B', 'C', 'D', 'E'))
ax.legend()
fig.tight_layout()
plt.show()
## Stacked Bars
import numpy as np
import matplotlib.pyplot as plt
N = 5
menMeans = (20, 35, 30, 35, 27)
womenMeans = (25, 32, 34, 20, 25)
menStd = (2, 3, 4, 1, 2)
womenStd = (3, 5, 2, 3, 3)
ind = np.arange(N) # the x locations for the groups
width = 0.35 # the width of the bars: can also be len(x) sequence
p1 = plt.bar(ind, menMeans, width, yerr=menStd)
p2 = plt.bar(ind, womenMeans, width,
bottom=menMeans, yerr=womenStd)
plt.ylabel('Scores')
plt.title('Scores by group and gender')
plt.xticks(ind, ('G1', 'G2', 'G3', 'G4', 'G5'))
plt.yticks(np.arange(0, 81, 10))
plt.legend((p1[0], p2[0]), ('Men', 'Women'))
plt.show()
## Horizontal Bar
import matplotlib.pyplot as plt
import numpy as np
# Fixing random state for reproducibility
np.random.seed(19680801)
plt.rcdefaults()
fig, ax = plt.subplots()
# Example data
people = ('Tom', 'Dick', 'Harry', 'Slim', 'Jim')
y_pos = np.arange(len(people))
performance = 3 + 10 * np.random.rand(len(people))
error = np.random.rand(len(people))
ax.barh(y_pos, performance, xerr=error, align='center',
color='green', ecolor='black')
ax.set_yticks(y_pos)
ax.set_yticklabels(people)
ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel('Performance')
ax.set_title('How fast do you want to go today?')
plt.show()
## Scatter Plot
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cbook as cbook
# Load a numpy record array from yahoo csv data with fields date, open, close,
# volume, adj_close from the mpl-data/example directory. The record array
# stores the date as an np.datetime64 with a day unit ('D') in the date column.
with cbook.get_sample_data('goog.npz') as datafile:
price_data = np.load(datafile)['price_data'].view(np.recarray)
price_data = price_data[-250:] # get the most recent 250 trading days
delta1 = np.diff(price_data.adj_close) / price_data.adj_close[:-1]
# Marker size in units of points^2
volume = (15 * price_data.volume[:-2] / price_data.volume[0])**2
close = 0.003 * price_data.close[:-2] / 0.003 * price_data.open[:-2]
fig, ax = plt.subplots()
ax.scatter(delta1[:-1], delta1[1:], c=close, s=volume, alpha=0.5)
ax.set_xlabel(r'$\Delta_i$', fontsize=15)
ax.set_ylabel(r'$\Delta_{i+1}$', fontsize=15)
ax.set_title('Volume and percent change')
ax.grid(True)
fig.tight_layout()
plt.show()
## import numpy as np
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import matplotlib.cbook as cbook
years = mdates.YearLocator() # every year
months = mdates.MonthLocator() # every month
yearsFmt = mdates.DateFormatter('%Y')
# Load a numpy record array from yahoo csv data with fields date, open, close,
# volume, adj_close from the mpl-data/example directory. The record array
# stores the date as an np.datetime64 with a day unit ('D') in the date column.
with cbook.get_sample_data('goog.npz') as datafile:
r = np.load(datafile)['price_data'].view(np.recarray)
fig, ax = plt.subplots()
ax.plot(r.date, r.adj_close)
# format the ticks
ax.xaxis.set_major_locator(years)
ax.xaxis.set_major_formatter(yearsFmt)
ax.xaxis.set_minor_locator(months)
# round to nearest years...
datemin = np.datetime64(r.date[0], 'Y')
datemax = np.datetime64(r.date[-1], 'Y') + np.timedelta64(1, 'Y')
ax.set_xlim(datemin, datemax)
# format the coords message box
def price(x):
return '$%1.2f' % x
ax.format_xdata = mdates.DateFormatter('%Y-%m-%d')
ax.format_ydata = price
ax.grid(True)
# rotates and right aligns the x labels, and moves the bottom of the
# axes up to make room for them
fig.autofmt_xdate()
plt.show()
# Style
print(plt.style.available)
```
# Customizing the plot
```
# sphinx_gallery_thumbnail_number = 10
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import FuncFormatter
plt.rcParams.update({'figure.autolayout': True})
data = {'Barton LLC': 109438.50,
'Frami, Hills and Schmidt': 103569.59,
'Fritsch, Russel and Anderson': 112214.71,
'Jerde-Hilpert': 112591.43,
'Keeling LLC': 100934.30,
'Koepp Ltd': 103660.54,
'Kulas Inc': 137351.96,
'Trantow-Barrows': 123381.38,
'White-Trantow': 135841.99,
'Will LLC': 104437.60}
group_data = list(data.values())
group_names = list(data.keys())
group_mean = np.mean(group_data)
fig, ax = plt.subplots(figsize=[6,3])
ax.barh(group_names, group_data)
labels = ax.get_xticklabels()
plt.setp(labels, rotation=45, horizontalalignment='right')
fig.show()
# Temporary styling
with plt.style.context(('dark_background')):
plt.plot(np.sin(np.linspace(0, 2 * np.pi)), 'r-o')
plt.show()
with plt.xkcd():
plt.plot(np.sin(np.linspace(0, 2 * np.pi)))
plt.show()
# Dynamic styling
import matplotlib as mpl
mpl.rcParams['lines.linewidth'] = 1
plt.plot([1, 2, 3])
mpl.rc('lines', linewidth=8, color='g')
plt.plot([1,2,3], 'g')
f, ax = plt.subplots()
n, bins, patches = ax.hist(np.random.randn(1000), 50,
facecolor='yellow', edgecolor='yellow')
plt.setp(patches, facecolor='red')
ax.set_xlabel('time (s)')
plt.show()
mpl.artist.getp(patches[0])
```
## Artist Property
[Artist Property](https://matplotlib.org/tutorials/intermediate/artists.html#sphx-glr-tutorials-intermediate-artists-py)
```
fig, ax = plt.subplots()
axis = ax.xaxis
axis.get_ticklocs()
axis.get_ticklabels()
axis.get_ticklines()
# Artist Hiarachy
```
# Layout
```
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
fig1, f1_axes = plt.subplots(ncols=2, nrows=2, constrained_layout=True)
fig1, f1_axes = plt.subplots(ncols=2, nrows=2)
fig2 = plt.figure(constrained_layout=True)
spec2 = gridspec.GridSpec(ncols=2, nrows=2, figure=fig2)
f2_ax1 = fig2.add_subplot(spec2[0, 0])
f2_ax2 = fig2.add_subplot(spec2[0, 1])
f2_ax3 = fig2.add_subplot(spec2[1, 0])
f2_ax4 = fig2.add_subplot(spec2[1, 1])
fig3 = plt.figure(constrained_layout=True)
gs = fig3.add_gridspec(3, 3)
f3_ax1 = fig3.add_subplot(gs[0, :])
f3_ax1.set_title('gs[0, :]')
f3_ax2 = fig3.add_subplot(gs[1, :-1])
f3_ax2.set_title('gs[1, :-1]')
f3_ax3 = fig3.add_subplot(gs[1:, -1])
f3_ax3.set_title('gs[1:, -1]')
f3_ax4 = fig3.add_subplot(gs[-1, 0])
f3_ax4.set_title('gs[-1, 0]')
f3_ax5 = fig3.add_subplot(gs[-1, -2])
f3_ax5.set_title('gs[-1, -2]')
fig5 = plt.figure(constrained_layout=True)
widths = [2, 3, 1.5]
heights = [1, 3, 2]
spec5 = fig5.add_gridspec(ncols=3, nrows=3, width_ratios=widths,
height_ratios=heights)
for row in range(3):
for col in range(3):
ax = fig5.add_subplot(spec5[row, col])
label = 'Width: {}\nHeight: {}'.format(widths[col], heights[row])
ax.annotate(label, (0.1, 0.5), xycoords='axes fraction', va='center')
gs_kw = dict(width_ratios=widths, height_ratios=heights)
fig6, f6_axes = plt.subplots(ncols=3, nrows=3, constrained_layout=True,
gridspec_kw=gs_kw)
fig7, f7_axs = plt.subplots(ncols=3, nrows=3)
gs = f7_axs[1, 2].get_gridspec()
# remove the underlying axes
for ax in f7_axs[1:, -1]:
ax.remove()
axbig = fig7.add_subplot(gs[1:, -1])
axbig.annotate('Big Axes \nGridSpec[1:, -1]', (0.1, 0.5),
xycoords='axes fraction', va='center')
fig7.tight_layout()
fig8 = plt.figure(constrained_layout=False)
gs1 = fig8.add_gridspec(nrows=3, ncols=3, left=0.1, right=0.48, wspace=0.5, hspace=0.5)
f8_ax1 = fig8.add_subplot(gs1[:-1, :])
f8_ax2 = fig8.add_subplot(gs1[-1, :-1])
f8_ax3 = fig8.add_subplot(gs1[-1, -1])
fig8 = plt.figure(constrained_layout=False)
gs1 = fig8.add_gridspec(nrows=3, ncols=3, left=0.32, right=0.48, wspace=0.5, hspace=0.5)
f8_ax1 = fig8.add_subplot(gs1[:-1, :])
f8_ax2 = fig8.add_subplot(gs1[-1, :-1])
f8_ax3 = fig8.add_subplot(gs1[-1, -1])
import warnings
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.colors as mcolors
import matplotlib.gridspec as gridspec
import matplotlib._layoutbox as layoutbox
plt.rcParams['savefig.facecolor'] = "0.8"
plt.rcParams['figure.figsize'] = 4.5, 4.
def example_plot(ax, fontsize=12, nodec=False):
ax.plot([1, 2])
ax.locator_params(nbins=3)
if not nodec:
ax.set_xlabel('x-label', fontsize=fontsize)
ax.set_ylabel('y-label', fontsize=fontsize)
ax.set_title('Title', fontsize=fontsize)
else:
ax.set_xticklabels('')
ax.set_yticklabels('')
fig, ax = plt.subplots(constrained_layout=False)
example_plot(ax, fontsize=24)
fig, axs = plt.subplots(2, 2, constrained_layout=False)
for ax in axs.flatten():
example_plot(ax)
fig, axs = plt.subplots(2, 2, constrained_layout=True)
for ax in axs.flatten():
example_plot(ax)
```
| github_jupyter |
# Introduction
This notebook demonstrates how to generate various slabs for a material using surface.py. In addition, it will also demonstrate how to obtain the Wulff shape with a set of surface energies associated with Miller indices.
# References
R. Tran, Z. Xu, B. Radhakrishnan, D. Winston, W. Sun, K. A. Persson, S. P. Ong, "Surface Energies of Elemental Crystals", Scientific Data, 2016, 3:160080, doi:10.1038/sdata.2016.80.
Sun, W.; Ceder, G. Efficient creation and convergence of surface slabs, Surface Science, 2013, 617, 53–59, doi:10.1016/j.susc.2013.05.016.
```
# Import the neccesary tools to generate surfaces
from pymatgen.core.surface import SlabGenerator, generate_all_slabs, Structure, Lattice
# Import the neccesary tools for making a Wulff shape
from pymatgen.analysis.wulff import WulffShape
import os
# Let's start with fcc Ni
lattice = Lattice.cubic(3.508)
Ni = Structure(lattice, ["Ni", "Ni", "Ni", "Ni"],
[[0,0,0], [0,0.5,0],
[0.5,0,0], [0,0,0.5]
])
# We'll use the SlabGenerator class to get a single slab. We'll start with the
# (111) slab of Ni. Plug in the CONVENTIONAL unit cell of your structure, the
# maximum Miller index value to generate the different slab orientations along
# with the minimum slab and vacuum size in Angstroms
slabgen = SlabGenerator(Ni, (1,1,1), 10, 10)
# If we want to find all terminations for a particular Miller index orientation,
# we use the get_slabs() method. This returns a LIST of slabs rather than a single
# slab. When generating a slab for a particular orientation, there are sometimes
# more than one location we can terminate or cut the structure to create a slab. The
# simplest example of this would be the Si(Fd-3m) (111) slab which can be cut or
# terminated in two different locations along the vector of the Miller index. For a
# fcc structure such as Ni however, there should only be one way to cut a (111) slab.
all_slabs = slabgen.get_slabs()
print("The Ni(111) slab only has %s termination." %(len(all_slabs)))
# Let's try this for a diamond Silicon structure
lattice = Lattice.cubic(5.46873)
Si = Structure(lattice, ["Si", "Si", "Si", "Si",
"Si", "Si", "Si", "Si"],
[[0.00000, 0.00000, 0.50000],
[0.75000, 0.75000, 0.75000],
[0.00000, 0.50000, 0.00000],
[0.75000, 0.25000, 0.25000],
[0.50000, 0.00000, 0.00000],
[0.25000, 0.75000, 0.25000],
[0.50000, 0.50000, 0.50000],
[0.25000, 0.25000, 0.75000]])
slabgen = SlabGenerator(Si, (1,1,1), 10, 10)
print("Notice now there are actually now %s terminations that can be \
generated in the (111) direction for diamond Si" %(len(slabgen.get_slabs())))
# The simplest way to do this is to just use generate_all_slabs which finds all the unique
# Miller indices for a structure and uses SlabGenerator to create all terminations for all of them.
all_slabs = generate_all_slabs(Si, 3, 10, 10)
print("%s unique slab structures have been found for a max Miller index of 3" %(len(all_slabs)))
# What are the Miller indices of these slabs?
for slab in all_slabs:
print(slab.miller_index)
print("Notice some Miller indices are repeated. Again, this is due to there being more than one termination")
```
# Calculating the surface energy
To do this, we actually need to calculate (from first principles) the total energy of two structures. The total energy of the bulk ($E_{bulk}$) (preferably the oriented unit cell corresponding to the slab) and the total energy of the slab($E_{slab}$) which is several layers of atoms interfaced with a vacuum. Using the following equation, we can get the surface energy $\gamma = \frac{E_{slab}-E_{bulk}}{2A}$ where both slab and bulk systems have the same number of atoms and species (you can multiply $E_{bulk}$ where n is the number of oriented unit cell layers contained in your slab system relative ot your bulk system). The factor of 1/2 accounts for the two surfaces in our slab model so $\gamma$ is the surface energy of one surface.
# Some basic properties of the Wulff shape
Using the WulffShape class, there are a couple of properties we can get. Here we will explain what some of them are.
weighted_surface_energy:
Surface Gibbs free energy for a crystal is given by $\Delta G=\sum\limits_{hkl}\gamma_{hkl}A_{hkl}$. Where $\gamma_{hkl}$ is the surface energy of facet (hkl) and $A_{hkl}$ is the surface area of that particular facet that occupies the Wulff shape. We can normalize this value with the total surface area of the Wulff shape to get the weighted (average) surface energy for a particular material $\bar{\gamma}=\frac{\Delta G}{\sum\limits_{hkl}A_{hkl}}$
anisotropy:
Typically in the literature when discussing surface anisotropy, we would only look at the ratios of 2 surface energies when talking about anisotropy. eg. the ratio of a generic fcc (111) to (100) surface energy should be less than 1 as the (111) facet is the closest packed surface of an fcc structure and should have the lowest surface energy. However this method of determining surface anisotropy does not allow us to determine an overall anisotropy of a material, ie. how different are all the surface energies for a material. As such, we used the Coefficient of Variation from the weighted surface energy. For reference, an ideal sphere Wulff shape (eg. completely isotropic) has a anisotropy of 0.
shape_factor:
An alternative to anisotropy. This is useful for determining the critical nucleus size. A large shape factor indicates great anisotropy.
See Ballufi, R. W., Allen, S. M. & Carter, W. C. Kinetics of Materials. (John Wiley & Sons, 2005), p.461
```
# Now let's assume that we then calculated the surface energies for these slabs
# Surface energy values in J/m^2
surface_energies_Ni = {(3, 2, 0): 2.3869, (1, 1, 0): 2.2862,
(3, 1, 0): 2.3964, (2, 1, 0): 2.3969,
(3, 3, 2): 2.0944, (1, 0, 0): 2.2084,
(2, 1, 1): 2.2353, (3, 2, 2): 2.1242,
(3, 2, 1): 2.3183, (2, 2, 1): 2.1732,
(3, 3, 1): 2.2288, (3, 1, 1): 2.3039,
(1, 1, 1): 1.9235}
miller_list = surface_energies_Ni.keys()
e_surf_list = surface_energies_Ni.values()
# We can now construct a Wulff shape with an accuracy up to a max Miller index of 3
wulffshape = WulffShape(Ni.lattice, miller_list, e_surf_list)
# Let's get some useful information from our wulffshape object
print("shape factor: %.3f, anisotropy: \
%.3f, weighted surface energy: %.3f J/m^2" %(wulffshape.shape_factor,
wulffshape.anisotropy,
wulffshape.weighted_surface_energy))
# If we want to see what our Wulff shape looks like
wulffshape.show()
# Lets try something a little more complicated, say LiFePO4
from pymatgen.util.testing import PymatgenTest
# Get the LiFePO4 structure
LiFePO4 = PymatgenTest.get_structure("LiFePO4")
# Let's add some oxidation states to LiFePO4, this will be
# important when we want to take surface polarity into consideration
LiFePO4.add_oxidation_state_by_element({"Fe": 2, "Li": 1, "P": 5, "O": -2})
slabgen = SlabGenerator(LiFePO4, (0,0,1), 10, 10)
```
When generating a slab of LiFePO4, we also want to be careful
not break any of the P-O bonds. These bonds are strong
enough that they will result in a significantly high surface
energy when broken so its reasonable to say that any terminations
with such broken bonds will not yield the lowest surface energy. To
implement this, we add the bonds parameter to get_slabs, a dictionary
where the key will be two atoms whose bonds we do not want to break
and the element of that value would be their maximum bond length.
```
all_slabs = slabgen.get_slabs(bonds={("P", "O"): 2})
# any bond between P and O less than 2 Angstroms cannot be broken when generating slabs
print("For the (001) slab of LiFePO4, there are %s terminations." %(len(all_slabs)))
```
There are a couple of rules before we actually run calculations on some of these
slab models. First off, we need to ensure that all slabs we will be calculating have
the same surface on both sides. To do this, we need to ensure the slab model has Laue
point group symmetry ie. contains inversion symmetry. We use the is_symmetric()
property of our slab object to check this. It's important that both surfaces are the
same as the above equation for surface energy is used to get the energy of one surface,
hence the 1/2 factor in the equation. If the surfaces are different (the slab is not
symmetric), we would be calculating the average surface energy of two different surfaces
in our slab rather than the surface energy for one slab in our calculation. Secondly,
for structures containing oxidation states, we need to ensure that our surfaces are
nonpolar. A polar termination will lead to a very high surface energy, so we can skip
those particular structures. We can check polarity using the is_polar() property of
our slab object. Both these criterias (nonpolar and symmetric) should be satisfied
before calculating a particular slab model.
```
for slab in all_slabs:
print(slab.is_polar(), slab.is_symmetric())
# Notice that none of the terminations in the (001) direction do not simultaneously satisfy
# our two criteria so a (001) surface with a reasonable surface energy cannot be calculated.
# In such cases, we need to modify the surfaces of our slabs. A future release of surface.py
# will implement such modification techniques for these cases.
# Now let's generate all possible slabs for a max Miller index of 2 for LiFePO4 and see if
# any of these surfaces can be calculated to yield reasonable and accurate surface energy
# values. This may take a while.
all_slabs = generate_all_slabs(LiFePO4, 2, 10, 10, bonds={("P", "O"): 2})
print("There is a total of %s slabs generated including polar, asymmetric, and \
P-O terminated slabs" %(len(all_slabs)))
# store any slabs for calculation that satisfies our criterias
valid_slabs = []
for slab in all_slabs:
if not slab.is_polar() and slab.is_symmetric():
print(slab.miller_index)
valid_slabs.append(slab)
print("Number of slabs that are nonpolar, symmetric and do not terminate P-O bonds: %s" %(len(valid_slabs)))
```
| github_jupyter |
# Numpy Basics
Numpy is a core library of many scientific packages in Python. Its highly optimized and written largely in C and is critical for performing complex calculations as quckly as possible. It provides some basic data structures, like the Numpy Array, that we'll be using extensively (or other libraries will be using under the hood).
The numpy array is similar to a python list, save that is is wrapped in a numpy class called `ndarray`.
```
import numpy as np
array = np.array([1,2,3,4,5])
# Numpy supports many attributes on the ndarray
print(type(array))
print(array)
print(array.shape)
print(array.size)
print(array.ndim)
print(array.dtype)
```
By default numpy is giving me an array of type int64. Assuming I wanted something else, you can force the type with the `dtype` kwarg.
```
array = np.array([1,2,3,4,5], dtype=np.float64)
print(array)
print(array.dtype)
array = np.array([1.0, 2.2, 3.3], dtype=np.int8)
print(array)
```
You can also change the type of an ndarray after the fact
```
array = array.astype(np.float64)
print(array)
```
Ndarrays can also be used for more interesting things. The `nd` after all stands for __N-Dimensional__, so we can have much more complicated matrices
```
array = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
print(type(array))
print(array)
print(array.shape)
print(array.size)
print(array.ndim)
print(array.dtype)
```
Like Python's lists, Numpy's arrays can be accessed by index, are iterable, and are __sliceable__
```
for i in range(array.shape[0]):
for j in range(array.shape[1]):
print(array[i][j])
print("\n\n")
for row in array:
for element in row:
print(element)
print("\n\n")
# Note the difference between the slicing styles!
print(array[1:][:3])
print(array[1:, :3])
```
### Operators and Numpy
Ndarrays supporrt most python operators, and will usually do what you expect
```
array + array
array * 2
array - 1
array % 2
array == 3
array != 1
```
Numpy also supports vector and matrix operations
```
a = np.array([[1,2], [2,3]])
b = np.array([[5,6], [2,3]])
a.dot(b)
np.matmul(a, b)
# Transpose
np.array([[1,1], [2,3]]).T
```
You can use the fact that numpy works with boolean operators to index into numpy arrays and return subsets of the data based on specific criteria.
```
index = array < 5
print(array[index])
print(array[(array < 5) & (array > 2)])
print(array[(array < 5) | (array > 10)])
```
### Ndarray Methods
Ndarrays supports a myriad of methods, a few of the most useful are listed below:
* sum
* mean
* std
* min
* max
* min
* max
* sqrt
```
array = np.ones(5)
print(array)
array.sum()
array.mean()
array = np.array([1,2,3,4,5])
print(array.min())
print(array.max())
array.std()
```
Numpy also supports an extensive library of other methods designed for linear algebra and scientific computing. A very useful one is `Polynomial` which can be used for fitting a line. Lets generate a random data set, and attempt to fit a line to it.
```
import matplotlib.pyplot as plt
X = np.linspace(0.0, 20.0, 100)
Y = 0.2 + 0.05 * X + 0.4 * np.sin(X) + 0.1 * np.random.normal(size = (100))
plt.figure(1)
plt.clf()
plt.plot(X, Y, 'o-')
plt.draw()
```
This data plot might be best described as a noisy sine wave.
```
import numpy.polynomial.polynomial as poly
# Polyfit will generate coefficients, the 3rd argument is degrees of fitting
coefficients = poly.polyfit(X, Y, 2)
fit = poly.polyval(X, coefficients)
plt.figure(1)
plt.clf()
plt.plot(X, Y, 'o-', X, fit, '--')
plt.draw()
```
We'll need to scale up the number of degrees, but keep in mind, the higher the number, the longer the operation can take. Fitting the line too precisely can also lead to overfitting.
```
coefficients = poly.polyfit(X, Y, 10)
fit = poly.polyval(X, coefficients)
plt.figure(1)
plt.clf()
plt.plot(X, Y, 'o-', X, fit, '--')
plt.draw()
```
### Working with Missing Data
Many datasets are incomplete - that is, they have missing data. There are many ways we can handle missing data - we can normalize it, ignore it, or drop it completely. Let's look at some examples of how numpy represents missing data.
```
data = np.array([1,2,3,4, None])
print(data)
print(data.dtype)
```
Generally you do not want to be working with object data, unless the data in question is string, or stringlike. Numpy typically represents missing data with the NaN identifier (not a number). None, or empty string, might be a good representation for missing data in a string array, but for numeric vectors, we'll want to make sure the missing data is NaN. This will force the type of the data to float. You cannot have an array with NaN in it of type float.
```
data = np.array([1,2,3,4, None], dtype=np.float64)
print(data)
# this will cause an exception
data = np.array([1,2,3,4, None], dtype=np.int64)
print(data)
```
Missing data can be filtered out many ways. An easy choice is to use boolean indexing
```
data = np.array([1,2,3,4, np.nan])
print(data[~np.isnan(data)])
```
You can also replace missing values with something else, like the mean of the data.
```
data = np.array([4.2,2.7,3.4,4.9, np.nan])
data[np.isnan(data)] = data.mean()
print(data)
```
It didnt work! What happened? `mean` by dedfault does not ignore NaNs. Most of the numpy methods will not. Numpy does have a special method we can use here called `nanmean` if you want to find the mean before removing the missing data.
```
data = np.array([4.2,2.7,3.4,4.9, np.nan])
data[np.isnan(data)] = np.nanmean(data)
print(data)
```
We can also opt to remove entire rows that have missing data
```
data = np.array([[1,2,4], [3,5,6], [4, np.nan, 5], [1,2,3]])
print(data)
print()
print(data[~np.isnan(data).any(axis=1)])
```
We can use a mask and then apply the `any` method on it to remove any rows that are not true in every column (hence the axis=1 argument).
| github_jupyter |
# Carry com DI Futuro
### Bibliotecas
```
import pandas as pd
import numpy as np
from dataapi import DBConnect, SGS
import matplotlib.pyplot as plt
from calendars import DayCounts
from tqdm import tqdm
%matplotlib inline
```
### Dados do DI Futuro
Servidor AWS do FinanceHub.
```
dbc = DBConnect(username='fhreadonly', password='finquant')
query = "select * from \"B3futures\" where contract='DI1'"
df = pd.read_sql(query, dbc.connection)
df.tail(10)
```
### Cálculo dos dias úteis até o vencimento de um contrato
```
df['time_stamp'] = pd.to_datetime(df['time_stamp'])
mat_dict = {'01': 'F',
'02': 'G',
'03': 'H',
'04': 'J',
'05': 'K',
'06': 'M',
'07': 'N',
'08': 'Q',
'09': 'U',
'10': 'V',
'11': 'X',
'12': 'Z'}
mat_dict = {v: k for k, v in mat_dict.items()}
mat_month = df['maturity_code'].str[0].map(mat_dict)
def convert_year(year2digits):
if int(year2digits) >= 91:
year4digits = '19' + year2digits
else:
year4digits = '20' + year2digits
return year4digits
mat_year = df['maturity_code'].str[-2:].apply(convert_year)
dates = pd.to_datetime(mat_year + mat_month + '01')
dc = DayCounts('BUS/252', calendar='anbima')
dates = dc.busdateroll(dates, 'following')
dates
df['maturity_date'] = dates
df['DU'] = dc.days(df['time_stamp'], df['maturity_date'])
df = df[df['DU'] >= 0]
# Utilizando apenas PUs não nulos
df = df[df.settlement_price != 0]
# Apenas Open Interest > 1
df = df[df['trading_volume'] != 0]
df.tail(35)
```
### Série do CDI de um dia
API do [Sistema Gerenciador de Series](https://www3.bcb.gov.br/sgspub/localizarseries/localizarSeries.do?method=prepararTelaLocalizarSeries) do BCB.
```
sgs = SGS()
df_cdi = sgs.fetch({12: 'CDI'})/100
df_cdi.tail(5)
```
# (1) Montando a Curva de Juros
## Cálculo do PU dos DIs
No DataFrame `df`, a coluna `settlement_price` é o PU (ou notional) do contrato de DI. A coluna `DU` é o número de dias úteis, de acordo com o calendário da anbima, entre a data de hoje e o vencimento do contrato.
$$
DI_{t}=\left(\frac{100000}{PU_{t}}\right)^{\frac{252}{bus}}-1
$$
```
df['settlement_rate'] = (100000/df['settlement_price'])**(252/df['DU'])-1
df
df_yield_curve = pd.pivot_table(df, values='settlement_rate', index=['time_stamp'], columns=['DU'], aggfunc=np.sum)
df_yield_curve.tail(5)
# Taxa CDI anualizada
cdi_series = pd.Series(((1+df_cdi['CDI'])**(252))-1)
# Dateframe atualizado com taxa CDI anualizada
df_yield_curve[1] = df_yield_curve[1].fillna(value=cdi_series)
df_yield_curve.tail(95)
```
**Interpolação cúbica**
O método de interpolação escolhido foi o método cúbico, que garante uma aproximação melhor que os métodos quadráticos e lineares, porém mantém o comportamento da curva, diferentemente do Spline.
A desvantagem são os "efeitos de borda", pois não é realizada a interpolação para pontos além da última medição, o que leva a uma redução das variáveis em algumas situações.
```
df_yield_curve_int = df_yield_curve.interpolate(method ='cubic', axis=1)
df_yield_curve_int
int_plot = df_yield_curve_int.iloc[4535]
int_plot.plot(figsize=(12,8), color = 'black')
#int_plot.savefig('int curve.png', dpi=400)
```
---
# (2) Estratégia de Carry
## Carry
Cálculo do carry anualizado de se segurar um contrato por 3 meses, para os vértices da curva de 6 meses em 6 meses.
```
# Dataframe com yield curve para último dia útil da série
df_yield_curve_day = df_yield_curve.loc['2020-05-15'].dropna()
df_yield_curve_int_day = df_yield_curve_int.loc['2020-05-15'].dropna()
df_yield_curve_day
# Vetor de vértices de DIs disponíveis na data analisada
vert = df_yield_curve_day.index.values
vert = vert[vert>63]
vert
# Dataframe com yield curve para os vértices de DIs disponíveis
y_ini = df_yield_curve_int_day[vert].values
# Dataframe com yield curve, começando 3 meses antes do y_ini
y_fim = df_yield_curve_int_day[vert-63].values
# Vetor com o cálculo do carry
carry = (((1+y_ini)**(vert/63))/((1+y_fim)**((vert-63)/63)))-1
# Dataframe do vetor carry
df_carry = pd.DataFrame(carry, index=vert, columns =['Carry'])
df_carry
```
## Carry ajustado por DV01
```
PU = 100000/(1+df_yield_curve_int_day[vert])**(vert/252)
dv01 =(vert/252)*PU.values*0.01
dv01
carry_ajust_dv01 = carry/dv01
df_carry_ajust_dv01 = pd.DataFrame(carry_ajust_dv01, index=vert, columns =['Carry Ajust DV01'])
df_carry_ajust_dv01
```
## Carry Ajustado por Duration
```
carry_ajust_dur = carry/(vert/252)
df_carry_ajust_dur = pd.DataFrame(carry_ajust_dur, index=vert, columns =['Carry Ajust Dur'])
df_carry_ajust_dur
```
## Estratégia
Tracker dos carry trades ao longo das datas em 2020 e avaliação da performance através do Sharpe.
```
df_yield_curve_int_tracker = df_yield_curve_int.loc['2020-05-15':]
# Melhor Sharpe
j = 76
df_retorno = pd.DataFrame(np.nan, index = df_yield_curve_int_tracker.index, columns =['Retorno'])
for i in np.arange(0,len(df_yield_curve_int_tracker.index)-1):
y_ant = df_yield_curve_int_tracker.iloc[i,j]
y = df_yield_curve_int_tracker.iloc[i+1,j-1]
pu_ant = 100000/((1+y_ant)**(j/252))
pu = 100000/((1+y)**((j-1)/252))
df_retorno.iloc[i+1,0] = pu/pu_ant - 1
j = j - 1
if j <= 63:
j = 126
print(df_retorno)
print(df_retorno.cumsum())
print(df_cdi.loc['2020-01-02':'2020-05-15'].cumsum())
df_cdi_bench = df_cdi.loc['2020-01-02':'2020-05-15']
X1 = (1 + np.nanmean(df_retorno.values))**252-1
X2 = (1 + np.nanmean(df_cdi_bench.values))**252-1
X3 = np.nanstd(df_retorno.values, ddof=1)*np.sqrt(252)
Sharpe = (X1-X2)/X3
print(Sharpe)
# Maior Retorno Acumulado
j = 76
df_retorno_2 = pd.DataFrame(np.nan, index = df_yield_curve_int_tracker.index, columns =['Retorno'])
for i in np.arange(0,len(df_yield_curve_int_tracker.index)-1):
y_ant = df_yield_curve_int_tracker.iloc[i,j]
y = df_yield_curve_int_tracker.iloc[i+1,j-1]
pu_ant = 100000/((1+y_ant)**(j/252))
pu = 100000/((1+y)**((j-1)/252))
df_retorno_2.iloc[i+1,0] = pu/pu_ant - 1
j = j - 1
if j <= 1575:
j = 1638
print(df_retorno_2)
print(df_retorno_2.cumsum())
print(df_cdi.loc['2020-01-02':'2020-05-15'].cumsum())
df_cdi_bench = df_cdi.loc['2020-01-02':'2020-05-15']
Z1 = (1 + np.nanmean(df_retorno_2.values))**252-1
Z2 = (1 + np.nanmean(df_cdi_bench.values))**252-1
Z3 = np.nanstd(df_retorno_2.values, ddof=1)*np.sqrt(252)
Sharpe_2 = (Z1-Z2)/Z3
print(Sharpe_2)
```
| github_jupyter |
# Scientific Computing with Python
## Scientific Computing
Scientific computing refers to the use of computational tools to solve scientific problems. It is an umbrella term for multiple subfields of computer science and other scientific disciplines, e.g. machine learning, bioinformatics, simulation, etc.
## Python for Scientific Computing
For a long time, MATLAB was the *lingua franca* of scientific computing; however, recently things have changed dramatically. New languages for scientific computing have burst on to the scene (e.g. Julia) and older programming languages were supplemented with new libraries that gave them the capabilities to handle this programming paradigm. In the case of Python, the Scipy stack was critical in making Python a huge player in the scientific computing world.
The Scipy stack is a collection of open source libraries that enable easy scientific computing in Python. The most popular of these libraries are:
* **Numpy**: a fast matrix maths library for Python
* **Matplotlib**: a mature plotting library for Python (Note that this is the same as the library used in Julia)
* **Scipy**: a collection of utilities for scientific computing
* **pandas**: implements data structures for processing and manipulating data
* **Sympy**: a symbol maths library for Python
* **scikit-learn**: a machine learning library for Python
## Numpy and Matplotlib
In this tutorial, we are going to look into two of the above libraries, namely Numpy and Matplotlib. We will motivate this by a simple machine learning / data analysis example.
## Limitations of Python Lists
Python lists can act like vectors in a scientific implementation; however, operating on them is a big hassle. Firstly, in Python there is no such thing as a matrix. To implement a matrix, we need to create a list of lists.
```
list_of_lists = [list(range(5)) for i in range(5)] # 5x5 matrix
list_of_lists
print(list_of_lists[0][3]) # Access element (0, 3) (or (1, 4) in mathematical indexing)
```
To do operations with such a data structure requires writing lots of computer code. For instance, if we are to add 3 to every element in this matrix we need to loop over all the elements and add 3 to each.
```
for i in range(5):
for j in range(5):
list_of_lists[i][j] = list_of_lists[i][j] + 3
list_of_lists
```
As you can imagine, the more complex the operation that we want to perform, the harder it gets to implement and the more code we need to write (which increases the probability of making mistakes). Hence, we need a tool to abstract these operations away: Enter Numpy! Note, we use `import` to call a library and `as` to give the library a _nickname_.
```
import numpy as np
numpy_matrix = np.array([range(5) for i in range(5)])
numpy_matrix
numpy_matrix[0, 3] # Note how the syntax differs between Numpy arrays and Python lists of lists
```
To add a scalar to every element in the matrix, simply do:
```
numpy_matrix = numpy_matrix + 3
numpy_matrix
```
## Elementwise Operations
The above is an example of an elementwise operation (applying an operation to every element). Numpy is very efficient at those and has an intuitive syntax for them. Numpy uses vectorization (look that up!) to perform these operations efficiently. Here is an example of elementwise multiplication of two matrices.
```
# Example of array creation in Numpy
a = np.ones((5, 5))
a
# Multiplication by Scaler
a = a * 3
a
b = np.array([range(5, 10) for j in range(5)])
b
# Elementwise multiplication of two arrays
c = a * b
c
```
### Task 1
Look up and try some other elementwise operations in Numpy.
## Speed Up
Due to vectorization, Numpy provides a considerable amount of speed-up in elementwise operations compared to standard Python, even for one-dimensional arrays (lists). Here is an example of squaring each element in the array.
```
big_list = range(10000)
big_list
big_array = np.arange(10000)
big_array
%%timeit
# Standard Python
[x**2 for x in big_list]
%%timeit
# Numpy
big_array**2
```
Notice the massive speed-up that Numpy offers in comparison to standard Python. Note that `%%timeit` is called a **magic command**, and this specific magic command is a shortcut for the timing function **timeit**. You can look into magic commands if you want.
## Multi-Dimensional Arrays
Numpy is not just restricted to vectors and matrices - it can handle arrays of arbitrary dimensions (as long as you have enough memory!).
```
lots_of_zeroes = np.zeros((10, 100, 11))
lots_of_zeroes
lots_of_zeroes.shape
```
Numpy arrays can be easily manipulated. For instance, we can easily reshape an array.
```
different_zeroes = lots_of_zeroes.reshape(1000, 11)
different_zeroes.shape
```
We can also select subranges of the array.
```
# Select first row
lots_of_zeroes[0,:,:]
lots_of_zeroes[0,:,:].shape
# Select first 100th column
lots_of_zeroes[:,99,:]
lots_of_zeroes[:,99,:].shape
# Select first 5 rows and first 50 columns
lots_of_zeroes[:5,:50,:]
lots_of_zeroes[:5,:50,:].shape
```
We can also transpose the array.
```
lots_of_zeroes.transpose()
# All axes
lots_of_zeroes.transpose().shape
# Last two axes
lots_of_zeroes.transpose([0,2,1]).shape
```
### Task 2
Look up **broadcasting** in Numpy.
## Example: Linear Regression
The following is a simple example for a linear regression in Numpy. We have the following model:
$$
y = 3x + 5 + \epsilon \\
\epsilon \sim N(0, 0.04)
$$
```
X = np.random.uniform(size=(20, 1)) # Generate the Xs uniformly at random
Y = 3* X + 5. + np.random.normal(scale=0.2, size=(20, 1)) # Generate the Ys according to the equation above and add noise
```
Now we plot the generated data. For this we use Matplotlib, a plotting library in the Scipy stack.
```
import matplotlib.pyplot as plt # Import the library
%matplotlib inline
plt.scatter(X, Y) # Generate a scatter plot
plt.xlabel('X') # Label X axis
plt.ylabel('Y') # Label Y axis
plt.title('X vs Y')
```
We define $\tilde{X}$ as the design matrix with the following form $[1, X]$, i.e. the first column is ones and the second are the $x$ locations. Hence, the solution to the regression is given by:
$$
\hat{W} = (\tilde{X}^T\tilde{X})^{-1}\tilde{X}^TY
$$
```
X_tilde = np.hstack([np.ones((20, 1)), X])
XT_X = np.dot(X_tilde.transpose(), X_tilde)
XT_X_inv = np.linalg.inv(XT_X)
XT_Y = np.dot(X_tilde.transpose(), Y)
W_hat = np.dot(XT_X_inv, XT_Y)
print(W_hat) # Pretty close to the generative model!
```
Now we plot the results
```
xx = np.linspace(0, 1, 100)
yy = W_hat[1]*xx + W_hat[0]
plt.scatter(X, Y)
plt.plot(xx, yy, c='r')
plt.xlabel('X')
plt.ylabel('Y')
plt.title('X vs Y')
```
## Capstone Project
Generate data and fit it to a linear model of the form:
$$
y = a + bx_1 + cx+2 + \epsilon \\
\epsilon \sim N(0, 0.04)
$$
You are free to pick what $a$, $b$ and $c$ are.
If this was too easy, fit the data to a ridge regression model:
$$
\hat{W} = (\tilde{X}^T\tilde{X} + \lambda I)^{-1}\tilde{X}^TY
$$
You are free to choose what $\lambda$ is.
| github_jupyter |
# Learning Objectives
- [ ] 1.2.1 Implement sort algorithms.
- Insertion sort
- Bubble sort
- Quicksort
- Merge sort
- [ ] 1.2.2 Use examples to explain sort algorithms.
- [ ] 1.2.3 Implement search algorithms.
- Linear search
- Binary search
- Hash table search (after Abstract Data Type)
- [ ] 1.2.4 Use Examples to explain search algorithms.
- [ ] 1.2.5 Compare and describe the efficiencies of the sort and search algorithms using Big-$O$ notation for time complexity (worst case). Exclude: space complexity
- [ ] 2.3.1 Implement sort programs.
- Insertion sort
- Bubble sort
- Quicksort
- Merge sort
- [ ] 2.3.2 Implement search programs.
- Linear search
- Binary search
- Hash table search (after Abstract Data Type)
# References
1. Leadbetter, C., Blackford, R., & Piper, T. (2012). Cambridge international AS and A level computing coursebook. Cambridge: Cambridge University Press.
2. https://www.sparknotes.com/cs/sorting/bubble/section1/#:~:text=The%20total%20number%20of%20comparisons,since%20no%20swaps%20were%20made.
3. https://visualgo.net/en
# 10.1 Search Algorithm
A search algorithm is an algorithm to retrieve information from some data structure. Some examples include:
- Finding the maximum or minimum value in a list or array
- Checking to see if a given value is present in a set of values
- Retrieving a record from a database
## 10.1.1 Linear Search
A **linear search**, also called **serial** or **sequential** searches an item in a given array sequentially till the end of the collection. It does not require the data to be in any particular order.
To find the position of a particular value involves looking at each value in turn – starting with the first – and comparing it with the value you are looking for. When the value is found, you need to note its position. You must also be able to report the special case that a value has not been found. This last part only becomes apparent when the search has reached the final data item without finding the required value.
### Example
In this example, you have the array `[10,14,19,26,27,31,33,35,42,44]` and you are looking for the value `33` in the array.
<center>
<img src="images/algorithm_linear_search.gif" width="400" align="center"/>
</center>
The pseudocode for linear search function is given below. It returns the index of the searched value in the array if it exists. In the case that the value is not in the array, the function returns `-1`.
```
FUNCTION LINEARSEARCH(A: ARRAY of INTEGER, t: INTEGER) RETURNS INTEGER
DECLARE index: INTEGER
index ← -1
FOR i = 1 TO A.SIZE
IF A[i] = t THEN
index ← i
BREAK
ENDIF
ENDFOR
RETURN index
ENDFUNCTION
```
### Exercise
Implement a function `linear_search(array, val)` which searches the list `array` for a value `val` using the linear search algorithm.
Test your function with the following list
> `
[39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]`
with the values `9` and `2`. What do you observe for the latter value?
```
#YOUR_CODE_HERE
def linear_search(array,val):
index = -1
for i in range(0,len(array)):
if array[i] == val:
index = i
break
return index
array = [39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]
print(linear_search(array,9))
print(linear_search(array,2))
```
In linear search, all items are searched one-by-one to find the required item.
If the array has $n$ elements to be compared to,
- The best-case lookup to find an item is $1$ comparison, i.e., the item is at the head of the array.
- The worst-case lookup to find an item is $n$ comparisons, i.e. the item is at the end of the array.
- The average lookup to find an item is approximately $\frac{n}{2}$ comparisons.
Clearly, if $n$ is large, this can be a very large number of comparisons and the serial search algorithm can take a long time.
Consequently, we have for serial search,
- Advantage:
- algorithm is straightforward and easy to implement,
- data need not be in any particular order,
- works well if there is a small number of data item.
- Disadvantage:
- search can take a long time if value of $n$ is large, i.e. inefficient if there is a large number of data items.
- Variations:
- Search target requires a different criteria (not just object existence).
- Must find all instances of target.
- Must find particular instance of target (first, last, etc.).
- Must find object just greater/smaller than target.
## 10.1.2 Binary Search
In the previous section, we looked at linear search where the data is not required to be stored in any particular order. On the other hand, if we know that the data is stored in an ascending order, we can utilize the another algorithm called the **binary search**.
Workings of binary search algorithm:
- First check the MIDDLE element in the list.
- If it is the value we want, we can stop.
- If it is HIGHER than the value we want, we repeat the search process with the portion of the list BEFORE the middle element.
- If it is LOWER than the value we want, we repeat the search process with the portion of the list AFTER the middle element.
The following example illustrates the case where we're looking for the value `19` in the following sorted array.
<center>
<img src="images/algorithm_binary_search.jpg" height="400" align="center"/>
</center>
Note that if there is an even number of values in the array, dividing by two gives a whole number and we split the array there. However, if the array consists of an odd number of values we need to find the integer part of it, as an array index must be an integer.
The pseudocode for binary search function is given below. It returns the index of the searched value in the array if it exists. In the case that the value is not in the array, the function returns `-1`.
### Example
In this example, you have the array `[3,4,5,7,8,9,10,12,15,19,20,21,22,24,25]` and you are looking for the value `22` in the array.
<center>
<img src="images/algorithm_binary_search.gif" height="150" align="center"/>
</center>
```
FUNCTION BinarySearch(A: ARRAY of INTEGER, t: INTEGER) RETURNS INTEGER
DECLARE start, mid, end: INTEGER
start ← 1
end ← A.SIZE
WHILE start <= end DO
mid ← (start + end) DIV 2
IF t = A[mid] THEN
RETURN mid
ENDIF
IF t < A[mid] THEN
end ← mid – 1
ELSE
start ← mid + 1
ENDIF
ENDWHILE
RETURN -1
ENDFUNCTION
```
### Exercise
Implement a function `binary_search(array, val)` which searches the list `array` for a value `val` using the binary search algorithm described above.
Test your function with the following list
> `
[39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]`
with the values `9` and `2`.
```
#YOUR_CODE_HERE
def binary_search(array,val):
start = 0
end = len(array)
while start <end :
mid = (start + end ) //2
if val == array[mid]:
return mid
if val < array[mid]:
end = mid - 1
else:
start = mid + 1
return -1
array = [39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]
sorted_array = sorted(array)
print(sorted_array)
print(binary_search(sorted_array,9))
print(binary_search(sorted_array,2))
def binary_search_rec(array, low, high,val):
if high >= low:
mid = (low+high)//2
if array[mid]==val:
return mid
elif array[mid]<val:
return binary_search_rec(array,mid+1,high,val)
elif val<array[mid]:
return binary_search_rec(array,low,mid-1,val)
else:
return -1
```
If the array has $n$ elements to be compared to,
- The best-case lookup to find an item is $1$ comparison, i.e., the item is at the middle of the array.
- The worst-case lookup to find an item is approximately $\log_2{n}$ comparisons.
```
#YOUR_CODE_HERE
```
Jupyter Notebook provides a magic function `%timeit` and `%%timeit` to time a code execution.
* `%timeit` is used to time a single line of statement
* `%%timeit` is used to time all codes in a cell. `%%timeit` must be placed at first line of cell.
### Exercise
Use `%timeit` to time the code executions for both the functions:
- `linear_search`,
- `binary_search`
that you have coded in the previous exercise, using the
> `
[39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]`
with the search value `9`.
```
%%timeit
#YOUR_CODE_HERE
array = [39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]
linear_search(array,98)
%%timeit
#YOUR_CODE_HERE
array = [39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]
binary_search(sorted(array),98)
%%timeit
#YOUR_CODE_HERE
array = [39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]
binary_search_rec(sorted(array),0,len(98)
```
# 10.2 Sorting Algorithms
Sorting refers to arranging a fixed set of data in a particular order. Sorting orders could be numerical (`1`,`2`, `3`, ...), lexicographical/dictionary (`AA`, `AB`, `AC`, ...) or custom ('Mon', 'Tue', 'Wed', ...).
Sorting algorithms specify ways to arrange data in particular ways to put the data in order. In this section, it is assumed that the sorted data is in ascending order.
## 10.1 Insertion Sort
In insertion sort algorithm, we compare each element, termed `key` element, in turn with the elements before it in the array. We then insert the `key` element into its correct position in the array.
### Example
In this example, the array `[6,5,3,1,8,7,2,4]` is sorted with insertion sort.
<center>
<img src="images/algorithm_insertion_sort.gif" height="250" align="center"/>
</center>
The pseudocode for insertion sort function for an array containing integer elements is given below:
```
FUNCTION InsertionSort(A: ARRAY of INTEGER) RETURNS ARRAY of INTEGER
DECLARE j, temp: INTEGER
FOR i = 2 to A.SIZE
j ← i
WHILE j > 1 AND A[j] < A[j – 1] DO
temp ← A[j]
A[j] ← A[j - 1]
A[j - 1] ← temp
j ← j - 1
ENDWHILE
ENDFOR
RETURN A
ENDFUNCTION
```
### Exercise
Implement a function `insertion_sort(array)` which sorts the list `array` in the ascending order according to the insertion algorithm given above.
Test your function with the following list
> `
[39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]`.
```
#YOUR_CODE_HERE
arr = [6,5,3,1,8,7,2,4]
print(insertion_sort(arr))
arr = [6,5,3,1,8,7,2,4]
print(insertion_sort_ori(arr))
```
Note:
- The outer for-loop in Insertion Sort function always iterates $n-1$ times.
- The inner while-loop will make $1 + 2 + 3 ... + (n-1)=\frac{n(n-1)}{2}$ comparisons in worst case.
# 10.2 Bubble Sort
The next sorting algorithm iterates over an array multiple times.
* In each iteration, it takes 2 consecutive elements and compare them.
* It swaps the smaller value to the left and larger value to the right.
* It repeats until the larger elements "bubble up" to the end of the list, and the smaller elements moves to the "bottom". This is the reason for the naming of the algorithm.
* The right-hand side of the array are sorted.
### Example
In this example, the array `[6,5,3,1,8,7,2,4]` is sorted with bubble sort.
<center>
<img src="images/algorithm_bubble_sort.gif" height="250" align="center"/>
</center>
We see that
* For 1st iteration, we need to make $n-1$ comparisons. It will bring the largest value to the extreme right.
* For 2nd iteration, we need to make $n-2$ comparisons. It will bring 2nd largest value to the 2nd extreme right.
* And so on...
Consequently, we need a nested loops to make multiple iterations.
The pseudocode for bubble sort function for an array containing integer elements is given below:
```
FUNCTION BubbleSort(A: ARRAY of INTEGER) RETURNS ARRAY of INTEGER
DECLARE swap: BOOLEAN
DECLARE temp: INTEGER
FOR i = 1 to (A.SIZE – 1)
swap ← FALSE
FOR j = 1 to (A.SIZE – i)
IF A[j] > A[j + 1] THEN
temp ← A[j]
A[j] ← A[j + 1]
A[j + 1] ← temp
swap ← TRUE
ENDIF
ENDFOR
IF NOT swap THEN
BREAK
ENDIF
ENDFOR
RETURN A
ENDFUNCTION
a = [6,5,3,1,8,7,2,4]
def BubbleSort(a):
for i in range(0,len(a)-1):
swap = False
for j in range(0,len(a)-i-1):
if a[j] > a[j+1]:
temp = a[j]
a[j] = a[j+1]
a[j+1] = temp
swap = True
if swap == False:
break
return a
BubbleSort(a)
```
### Exercise
Implement a function `bubble_sort(array)` which sorts the list `array` in the ascending order according to the bubble algorithm given above.
Test your function with the following list
> `
[39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]`.
```
#YOUR_CODE_HERE
```
Note:
- The amount of comparisons in Bubble Sort algorithm is $(n - 1) + (n - 2) + ... + 1=\frac{n(n-1)}{2}$ comparisons,
- Best case is when the array is already sorted and bubble sort will terminate after the first iterations.
- Bubble sort is also efficient when one random element needs to be sorted into a sorted array, provided that new element is placed at the beginning and not at the end.
- The absolute worst case for bubble sort is when the smallest element of the array is the last element in the end of the array. Because in each iteration only the largest unsorted element gets put in its proper location, when the smallest element is at the end, it will have to be swapped each time through the array, and it wont get to the front of the list until all $n$ iterations have occurred.
# 10.3 Quicksort
Quicksort is a sorting technique based on divide and conquer technique. Quicksort first selects an element, termed the `pivot`, and partitions the array around the pivot, putting every smaller element into a low array and every larger element into a high array.
* The `pivot` element can selected randomly, but one way to select the pivot is to use the element in the middle of the array as the pivot
* The first pass partitions data into 3 sub-arrays, `lesser` (less than pivot), `equal` (equal to pivot) and `greater` (greater than pivot).
* The process repeats for `lesser` array and `greater` array.
<center>
<img src="images/algorithm_quick_sort.gif" height="250" align="center"/>
</center>
The pseudocode for quicksort function for an array containing $N$ elements is given below:
```
PROCEDURE QuickSort(Arr : ARRAY OF INTEGERS):
IF Arr.Size = 0:
RETURN []
ENDIF
Mid ← (1+Arr.Size) //2
Pivot ← Arr[Mid]
DECLARE Equal : Integer
DECLARE Lesser : Integer
DECLARE More : Integer
Equal = 0
Lesser = 0
More = 0
For i = 1 to Arr.Size
IF Arr[i] = Pivot THEN
Equal = Equal + 1
ENDIF
IF Arr[i] < Pivot THEN
Lesser = Lesser + 1
ENDIF
IF Arr[i] > Pivot THEN
More = More + 1
ENDIF
ENDFOR
DECLARE EqualArray[1:Equal] : ARRAY OF INTEGER
DECLARE LesserArray[1:Lesser] : ARRAY OF INTEGER
DECLARE MoreArray[1:More] : ARRAY OF INTEGER
DECLARE EqualIndex : Integer
DECLARE LesserIndex : Integer
DECLARE MoreIndex : Integer
EqualIndex = 1
LesserIndex = 1
MoreIndex = 1
For i = 1 to Arr.Size
IF Arr[i] = Pivot THEN
EqualArray[EqualIndex] ← Arr[i]
EqualIndex ← EqualIndex + 1
ENDIF
IF Arr[i] < Pivot THEN
LesserArray[LesserIndex] ← Arr[i]
LesserIndex ← LesserIndex + 1
ENDIF
IF Arr[i] > Pivot THEN
MoreArray[MoreIndex] ← Arr[i]
MoreIndex ← MoreIndex + 1
ENDIF
ENDFOR
RETURN QuickSort(LesserArray) + EqualArray + QuickSort(MoreArray) //Assuming `+` is array concatenation
END PROCEDURE
TYPE List
Buffer: ARRAY[1:N] OF OBJECT
Size: N
ENDTYPE
CREATE_LIST(N) RETURN List
APPEND(List, d: OBJECT) RETURNS List // appends d to end of List
SUB_LIST(LIST, start:INTEGER, end:INTEGER) RETURNS List //return sub-list from start to end
CONCATENATE(A:List, B:List) RETURNS List //concatenates A and B into a new List
LEN(L) RETURNS INTEGER //size of List
FUNCTION QS(L: List )
DECLARE
Lesser : List
Greater : List
Pivot: OBJECT
Lesser ← CREATE_LIST(0)
Greater ← CREATE_LIST(0)
Pivot ← L[1]
IF L.Size = 0 THEN
RETURN CREATE_LIST(0)
ENDIF
FOR i = 2 TO LEN(L)
IF L[i] <= Pivot THEN
APPEND(Lesser, L[i])
ELSE:
APPEND(Greater, L[i])
ENDIF
ENDFOR
RETURN CONCATENATE( APPEND(QS(Lesser),Pivot),QS(Greater))
ENDFUNCTION
def quicksort(arr):
if arr==[]:
return []
pivot = arr[len(arr)//2]
equal=[x for x in arr if x == pivot ]
lesser=[x for x in arr if x < pivot ]
more=[x for x in arr if x > pivot ]
return quicksort(lesser)+equal+quicksort(more)
arr = [6]
quicksort(arr)
```
### Exercise
Implement a function `quicksort(array)` which sorts the list `array` in the ascending order according to the quicksort algorithm given above.
Test your function with the following list
> `
[39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]`.
```
#YOUR_CODE_HERE
```
Note:
- The worst case scenario is when the smallest or largest element is always selected as the pivot. This would create partitions of size $n-1$, causing recursive calls $n-1$ times. And such, if the first element of partition is always chosen to be the first element and the array is already sorted.
- With a good pivot, the input list is partitioned in linear time, $O(n)$, and this process repeats recursively an average of $\log_2{n}$ times.
- This leads to a final complexity of $O(n \log_2n)$.
- The above implementation of the quicksort algorithm does not sort “in place”, and has a high space complexity. In order to overcome this, you need to change the algorithm slightly – i.e., use a variant that does not create new linked lists to store elements greater/less than the pivot. The in-place version of the pseudocode is given below.
```
PROCEDURE QuickSort(MyList, LB, UB)
IF LB <> UB THEN
#there is more than one element in MyList
LeftP ← LB #Left pointer
RightP ← UB #Right pointer
REPEAT
WHILE LeftP <> RightP AND MyList[LeftP] < MyList[RightP] DO
#move right pointer left
RightP ← RightP — l
ENDWHILE
IF LeftP <> RightP THEN
swap MyList[LeftP] and MyList[J]
WHILE LeftP <> RightP AND MyList[LeftP] < MyList[RightP] DO
#move left pointer right
LeftP ← LeftP + 1
ENDWHILE
IF LeftP <> RightP THEN
swap MyList[LeftP] and MyList[RightP]
UNTIL LeftP = RightP
#value now in correct position so sort left sub-list
QuickSort(MyList, LB, LeftP — 1)
#now sort right sub-list
QuickSort(MyList, LeftP + l, UB)
ENDIF
END PROCEDURE
```
# 10.4 Merge Sort
Similar to Quicksort, Merge sort is a sorting technique based on divide and conquer technique. It first divides the array into equal halves and then combines them in a sorted manner.
- if it is only one element in the list, it is already sorted. Return the list.
- divide the list recursively into two halves until it can no more be divided.
- merge the smaller lists into new list in sorted order.
<center>
<img src="images/algorithm_merge_sort.gif" height="250" align="center"/>
</center>
The important subroutine `merge` : Given two sorted array, $A$ and $B$ of size $n_1$ and $n_2$, `merge(A,B)` returns a sorted array of size $n_1+n_2$ whose elements come from $A$ and $B$.
The pseudocode for merge sort function for an array containing $N$ elements is given below:
```
FUNCTION Merge(A:ARRAY of INTEGER, B: ARRAY of Integer) RETURNS ARRAY of INTEGER
DECLARE C: ARRAY [1: A.SIZE + B.SIZE] of INTEGER
DECLARE J: INTEGER
J = 1
WHILE ( A.SIZE >= 1 AND B.SIZE >= 1)
IF ( A[1] > B[1] )
THEN
C[J] ← B[1]
J ← J + 1
B ← B [2 : B.SIZE]
ELSE
C[J] ← A[1]
J ← J + 1
A ← A [2 : A.SIZE]
ENDIF
ENDWHILE
WHILE ( A.SIZE >= 1 )
C[J] ← A[1]
J ← J + 1
A ← A [2 : A.SIZE]
ENDWHILE
WHILE ( B.SIZE >= 1 )
C[J] ← B[1]
J ← J + 1
B ← B [2 : B.SIZE]
ENDWHILE
RETURN C
ENDFUNCTION
FUNCTION MergeSort(A: ARRAY of INTEGER) RETURNS ARRAY of INTEGER
// Base case. A list of zero or one elements is sorted, by definition.
IF A.SIZE <= 1
THEN
RETURN A
ENDIF
// Recursive case. First, divide the list into roughly equal-sized subarrays
// consisting of the first half and second half of the array A.
DECLARE Left: ARRAY [1: A.SIZE/2] of INTEGER
DECLARE Right: ARRAY [1: A.SIZE/2] of INTEGER
FOR i = 1 to A.SIZE
IF i <= A.SIZE/2
THEN
Left[i] ← A[i]
ELSE
Right[i - A.SIZE/2] ← A[i]
// Recursively sort both subarray.
Left ← MergeSort(Left)
Right ← MergeSort(Right)
// Then merge the now-sorted sublists.
RETURN Merge(Left, Right)
ENDFUNCTION
```
Note:
- In sorting $n$ objects, merge sort has an average and worst-case performance of $O(n \log n)$. If the running time of merge sort for a list of length $n$ is $T(n)$, then the recurrence relation $T(n) = 2T(\frac{n}{2}) + n$ follows from the definition of the algorithm (apply the algorithm to two lists of half the size of the original list, and add the $n$ steps taken to merge the resulting two lists). The closed form follows from the master theorem for divide-and-conquer recurrences.
https://en.wikipedia.org/wiki/Master_theorem_(analysis_of_algorithms)
### Exercise
Implement a function `merge_sort(array)` which sorts the list `array` in the ascending order according to the merge sort algorithm given above.
Test your function with the following list
> `
[39, 96, 51, 20, 42, 42, 74, 28, 66, 16, 10, 86, 6, 43, 67, 98, 32, 73, 99, 7, 80, 88, 57, 83, 1, 64, 33, 38, 38, 8, 68, 38, 42, 80, 71, 82, 25, 29, 2, 85, 2, 96, 34, 14, 9, 65, 50, 63, 99, 94, 5, 93, 84, 46, 64, 22, 59, 31, 74, 13, 93, 13, 98, 93]`.
```
#YOUR_CODE_HERE
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import math
import os
def printCategory(category_dict):
for category in category_dict:
print(category)
if category_dict[category] is not None:
sub_dic = category_dict[category]
printCategory(sub_dic)
def createLevelColumns(df, levels, new_columns):
for i in range(len(df)):
df_level1, df_level2 = "NULL","NULL"
df_level1, nextLevels = findNotNullLevel(df, i, levels)
if nextLevels is not None:
df_level2, nextLevels = findNotNullLevel(df, i, nextLevels)
most_specific_category = "NULL"
if df_level2 != "NULL":
df_level2 = df_level2 + "__" + df_level1
if df_level2 != "NULL":
most_specific_category = df_level2
elif df_level1 != "NULL":
most_specific_category = df_level1
df.iloc[i, df.columns.get_loc('level1')] = df_level1
df.iloc[i, df.columns.get_loc('level2')] = df_level2
df.iloc[i, df.columns.get_loc('mostSpecificCategory')] = most_specific_category
def findNotNullLevel(df, i, levels):
for level in levels:
if df.iloc[i][level] == "NULL":
continue
else:
return df.iloc[i][level], levels[level]
return "NULL", None
def unionTwoLists(list1, list2):
for category in list1:
if category not in list2:
list2.append(category)
return list2
def checkNULL(checked_list):
for item in checked_list:
if item == "NULL":
print("Contains NULL")
return
print("Not contains NULL")
def create_name_id_dictionary(name_list):
count = 1
dic_name_id = {}
for name in name_list:
if name not in dic_name_id and name != "NULL":
dic_name_id[name] = count
count = count + 1
return dic_name_id
path = os.path.abspath(os.getcwd())
df = pd.read_csv(path + "/Data2/Last Names.csv")
df = df.dropna(axis = 0, how = 'all')
df = df.where(df.notnull(), "NULL")
print(df.columns)
df.head()
# {'马': {'id': 1, 'pinyin':'Ma'}, '李': ...}
name_pingying_id = {}
count = 1
name_heading = list(df.columns)[3:8]
pinyin_heading = list(df.columns)[8:13]
for i in range(len(df)):
for j in range(len(name_heading)):
name = df[name_heading[j]][i]
pinyin = df[pinyin_heading[j]][i]
if name not in name_pingying_id:
name_pingying_id[name]['id'] = count
name_pingying_id[name]['pinyin'] = pinyin
count = count + 1
df_last_name = pd.DataFrame(columns = ['gazetteerId', 'totalNumber', 'firstNameId', 'secondNameId', 'thirdNameId', 'fourthNameId','fifthNameId'])
dic_last_name = {'gazetteerId':[], 'totalNumber':[] 'firstNameId':[], 'secondNameId':[], 'thirdNameId':[], 'fourthNameId':[], 'fifthNameId':[]}
def create_name_id_dictionary(name_list):
count = 1
dic_name_id = {}
for name in name_list:
if name not in dic_name_id and name != "NULL":
dic_name_id[name] = count
count = count + 1
return dic_name_id
# create dict "dic_category_id" store { category_name : id}
total_categories = ['海拔 (米) Altitude', '平均降水量 Average Yearly Precipitation Amount', '平均温度 Average Yearly Temperature']
dic_category_id = create_name_id_dictionary(total_categories)
# create dict "dic_unit_id" store { unit_name : id}
unit_list = [cat for cat in df_total['unit'].astype('category').unique()]
dic_unit_id = create_name_id_dictionary(unit_list)
print(dic_category_id)
print(dic_unit_id)
for category in dic_category_id:
df_total = df_total.replace(category, dic_category_id[category])
for unit in dic_unit_id:
df_total = df_total.replace(unit, dic_unit_id[unit])
# create natural_environment_df
natural_environment_df = df_total.copy()
natural_environment_df.columns = columns = ['gazetteerId', 'category', 'data', 'unitId']
natural_environment_df.head()
# create economyCategory_df
category_df = pd.DataFrame(columns = ['id', 'name', 'parentId'])
dic_for_category_df = {'id':[], 'name':[], 'parentId':[]}
for category in dic_category_id:
child_parent = category.split('__', 1)
name = child_parent[0]
if len(child_parent) == 1:
dic_for_category_df['id'].append(dic_category_id[category])
dic_for_category_df['name'].append(name)
dic_for_category_df['parentId'].append("NULL")
else:
parentId = dic_category_id[child_parent[1]]
dic_for_category_df['id'].append(dic_category_id[category])
dic_for_category_df['name'].append(name)
dic_for_category_df['parentId'].append(parentId)
for attribute in category_df.columns:
category_df[attribute] = dic_for_category_df[attribute]
category_df.head()
dic_unit_df = {'id':[], 'name':[]}
for unit in dic_unit_id:
dic_unit_df['name'].append(unit)
dic_unit_df['id'].append(dic_unit_id[unit])
unit_id_df = pd.DataFrame(dic_unit_df,columns = ['id', 'name'])
unit_id_df.head()
unit_id_df.to_csv('natural_env_unit_df.csv',index = False, na_rep = "NULL")
natural_environment_df.to_csv('natural_env_df.csv', index = False, na_rep = "NULL")
category_df.to_csv('natural_env_category_df.csv', index = False, na_rep = "NULL")
```
| github_jupyter |
# Training a second model
In this notebook, I train a second model using features in order to address the first model's shortcomings.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pathlib import Path
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score, classification_report
from sklearn.externals import joblib
import sys
sys.path.append("..")
import warnings
warnings.filterwarnings('ignore')
from ml_editor.data_processing import (
format_raw_df,
get_split_by_author,
train_vectorizer,
get_vectorized_series,
get_feature_vector_and_label
)
from ml_editor.model_evaluation import (
get_feature_importance,
get_roc_plot,
get_confusion_matrix_plot,
get_calibration_plot
)
from ml_editor.model_v2 import (
add_char_count_features,
get_word_stats,
get_sentiment_score,
POS_NAMES,
get_question_score_from_input
)
%load_ext autoreload
%autoreload 2
np.random.seed(35)
data_path = Path('../data/writers.csv')
df = pd.read_csv(data_path)
df = format_raw_df(df.copy())
df = df.loc[df["is_question"]].copy()
df["full_text"] = df["Title"].str.cat(df["body_text"], sep=" ", na_rep="")
```
Let's add new features we've identified as potential candidates in our new model.
```
train_df, test_df = get_split_by_author(df, test_size=0.2, random_state=40)
vectorizer = train_vectorizer(train_df)
df["vectors"] = get_vectorized_series(df["full_text"].copy(), vectorizer)
```
Check out the ml_editor source code to see more about what these functions are doing!
```
df = add_char_count_features(df.copy())
df = get_word_stats(df.copy())
df = get_sentiment_score(df.copy())
feature_arr = ["num_questions",
"num_periods",
"num_commas",
"num_exclam",
"num_quotes",
"num_colon",
"num_stops",
"num_semicolon",
"num_words",
"num_chars",
"num_diff_words",
"avg_word_len",
"polarity"
]
feature_arr.extend(POS_NAMES.keys())
```
# Model
Now that we've added new features, let's train a new model. We'll use the same model as before, only the features are different.
```
# We split again since we have now added all features.
train_df, test_df = get_split_by_author(df, test_size=0.2, random_state=40)
X_train, y_train = get_feature_vector_and_label(train_df, feature_arr)
X_test, y_test = get_feature_vector_and_label(test_df, feature_arr)
y_train.value_counts()
X_test.shape
clf = RandomForestClassifier(n_estimators=100, class_weight='balanced', oob_score=True)
clf.fit(X_train, y_train)
y_predicted = clf.predict(X_test)
y_predicted_proba = clf.predict_proba(X_test)
```
Now, we can measure performance as we saw in the first training notebook.
```
def get_metrics(y_test, y_predicted):
# true positives / (true positives+false positives)
precision = precision_score(y_test, y_predicted, pos_label=True,
average='binary')
# true positives / (true positives + false negatives)
recall = recall_score(y_test, y_predicted, pos_label=True,
average='binary')
# harmonic mean of precision and recall
f1 = f1_score(y_test, y_predicted, pos_label=True, average='binary')
# true positives + true negatives/ total
accuracy = accuracy_score(y_test, y_predicted)
return accuracy, precision, recall, f1
# Training accuracy
# Thanks to https://datascience.stackexchange.com/questions/13151/randomforestclassifier-oob-scoring-method
y_train_pred = np.argmax(clf.oob_decision_function_,axis=1)
accuracy, precision, recall, f1 = get_metrics(y_train, y_train_pred)
print("Training accuracy = %.3f, precision = %.3f, recall = %.3f, f1 = %.3f" % (accuracy, precision, recall, f1))
accuracy, precision, recall, f1 = get_metrics(y_test, y_predicted)
print("Validation accuracy = %.3f, precision = %.3f, recall = %.3f, f1 = %.3f" % (accuracy, precision, recall, f1))
```
Fortunately, this model shows stronger aggregate performance than our previous model! Let's save our new model and vectorizer to disk so we can use them later.
```
model_path = Path("../models/model_2.pkl")
vectorizer_path = Path("../models/vectorizer_2.pkl")
joblib.dump(clf, model_path)
joblib.dump(vectorizer, vectorizer_path)
```
## Validating that features are useful
Next, we'll use the method described in the feature importance [notebook](https://github.com/hundredblocks/ml-powered-applications/blob/master/notebooks/feature_importance.ipynb) to validate that our new features are being used by the new model.
```
w_indices = vectorizer.get_feature_names()
w_indices.extend(feature_arr)
all_feature_names = np.array(w_indices)
k = 20
print("Top %s importances:\n" % k)
print('\n'.join(["%s: %.2g" % (tup[0], tup[1]) for tup in get_feature_importance(clf, all_feature_names)[:k]]))
print("\nBottom %s importances:\n" % k)
print('\n'.join(["%s: %.2g" % (tup[0], tup[1]) for tup in get_feature_importance(clf, all_feature_names)[-k:]]))
```
Our new features are amongst the most predictive! On the flip side, we can see that the word vectors from the TF-IDF vectorization approach don't seem to be particularly helpful. In a following [notebook](https://github.com/hundredblocks/ml-powered-applications/blob/master/notebooks/third_model.ipynb), we will train a third model without these features and see how well it performs.
## Comparing predictions to data
This section uses the evaluation methods described in the Comparing Data To Predictions [notebook](https://github.com/hundredblocks/ml-powered-applications/blob/master/notebooks/comparing_data_to_predictions.ipynb), but on our new model.
```
get_roc_plot(y_predicted_proba[:,1], y_test, figsize=(10,10))
get_confusion_matrix_plot(y_predicted, y_test, figsize=(9, 9))
get_calibration_plot(y_predicted_proba[:,1], y_test, figsize=(9,9))
```
## Inference Function
Just like for our first model, we define an inference function that takes in an arbitrary question and outputs an estimated probability of it receiving a high score according to our model.
```
pos_prob = get_question_score_from_input("""
When quoting a person's informal speech, how much liberty do you have to make changes to what they say?
""")
print("%s probability of the question receiving a high score according to our model" % (pos_prob))
```
| github_jupyter |
# Syllabus (https://bit.ly/intro_python_00)
# Lecture 1 - Intro (https://bit.ly/intro_python_01)
* What's a program?
* Hello World!
* Basic Input and Output
* Interpreted vs. compiled languages
* Ways to run Python (notebooks, via the interactive console, via an editor (PyCharm))
* The debug loop
* Basic input and output
* Comments
* Basic types
* integers (ints)
* strings
* floating point numbers (floats)
* ZyBook: Chapter 1
# Lecture 2 - Variables and Expressions (https://bit.ly/intro_python_02)
* Objects
* Variables
* Assignment is not equals
* Copying References
* Variable Names
* Expressions
* Statements
* Operators
* Abbreviated assignment
* Logical operators
* Order of operations
* ZyBook: Chapter 2
# Lecture 3 - More Types (https://bit.ly/intro_python_03)
* More strings
* Type Conversions
* Automatic Type Conversion
* More Input
* List basics
* Dictionary Basics
* ZyBook: Chapter 3
# Lecture 4 - Conditionals and Branching (https://bit.ly/intro_python_04)
* If statements
* Else
* Elif
* Pass
* First looping control flow:
* While loops
* ZyBook: Chapter 4
# Lecture 5 - Loops (https://bit.ly/intro_python_05)
* More control flow:
* For loops and list iteration
* Nested loops
* Break statement
* Continue statement
* Look at some complex examples of control flow that use control statements to get more comfortable
* ZyBook: Chapter 5
# Lecture 6 - Functions (https://bit.ly/intro_python_06)
* Function definitions and function calls
* Return values are optional
* Docstrings
* Return can be used for control flow
* None and the default return value
* Return can often sub for break
* Functions can call other functions - the stack
* ZyBook: Chapter 6
# Lecture 7 - Functions Continued (https://bit.ly/intro_python_07)
* Functions continued:
* Functions, namespaces and score
* Optional functional arguments and other tricks
* A few common mistakes about functions
* Some examples
# Lecture 8 - Strings (https://bit.ly/intro_python_08)
* Strings:
* String operators
* Length function
* Slicing
* Immutability
* String comparison
* For loops
* In operator
* Convenience functions
* Find
* Split
* Join
* etc.
* Format method
* Formatting numbers
* ZyBook: Chapter 7
# Lecture 9 - Tuples, Lists and Dictionaries (https://bit.ly/intro_python_09)
* Tuples (recap many things introduced with respect to strings)
* Lists
* List comprehensions
* More Dictionaries
* Sets
* ZyBook: Chapter 8
# Midterm in class (everything in lectures 1 - 9)
# Lecture 10 - Modules (https://bit.ly/intro_python_10)
* Finishing up sequences:
* Iterators vs. lists
* Generators and the yield keyword
* Modules:
* Some useful modules
* Hierarchical namespaces
* Making your own modules
* The main() function
* Parsing user arguments with the argparse module
* PEP 8 (v. briefly)
* A revisit to debugging, now that we're writing longer programs:
* Looking at different error types (syntax, runtime, logical)
* ZyBook: Chapter 11
# Lecture 11 - Files (https://bit.ly/intro_python_11)
* Writing Files
* Reading files
* With keyword
* For loops on file handles
* Binary files
* Large worked example (Fasta file processing)
* Directories
* Web file retrieval
* ZyBook: Chapter 12
# Lecture 12 - Classes and Objects (https://bit.ly/intro_python_12)
* The basics of Python classes and objects:
* Classes and Objects
* The \__init__ constructor method
* Object membership: Dot notation and classes
* Everything is an object in Python!
* Methods: adding functions to class and the self argument
* Object vs. class variables
* Objects Mutability
* Is vs. ==
* ZyBook: Chapter 9
# Lecture 13 - Objects and Polymorphism (https://bit.ly/intro_python_13)
* Python objects:
* Polymorphism
* Polymorphism by reimplementing basic object functionality:
* Redefining the \__str__ method
* How Python implements ''==" for objects
* General operator overloading, including comparison operators
* Copying objects
* No Zybooks!
# Lecture 14 - Inheritance (https://bit.ly/intro_python_14)
* Inheritance
* Object composition
* Inheritance vs. composition: aka: 'Is a' vs. 'has a'.
* An Example of OOP: Implementing the card game Old Maid
* ZyBook: Chapter 13
# Lecture 15 - More Functions and Recursion (https://bit.ly/intro_python_15)
* Recursion
* Consolidate: Look at some more complex examples of recursive control flow that use functions and control statements to get more comfortable
* Lambda functions
* Functions as arguments
* ZyBook: Chapter 14
# Lecture 16 - Exceptions and Unit Testing (https://bit.ly/intro_python_16)
* Unit-testing with unitest
* Exceptions and error handling
* ZyBook: Chapter 10
# Lecture 17 - Search Algorithms (https://bit.ly/intro_python_17)
* A couple of simple list algorithms:
* Linear search
* Binary search
* A very brief introduction to runtime analysis and Big O Notation
* Word counting with dictionaries
* Zybook Chapter 16
# Lecture 18 - Intro to data science (https://bit.ly/intro_python_18)
Worked example intro to Python data science
* NumPy
* MatPlotLib (to make plots)
* Pandas
* Sklearn
* Zybook Chapter 15
# Revision for final (everything!)
| github_jupyter |
```
# from google.colab import drive
# drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 125
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=64, out_channels=64, kernel_size=3, padding=0)
self.fc1 = nn.Linear(216, 128)
self.fc2 = nn.Linear(128, 64)
self.fc3 = nn.Linear(64, 10)
self.fc4 = nn.Linear(10,1)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
y = y.to("cuda")
x = x.to("cuda")
for i in range(9):
x[:,i] = self.helper(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
return x, y
def helper(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=64, out_channels=64, kernel_size=3, padding=0)
self.fc1 = nn.Linear(216, 128)
self.fc2 = nn.Linear(128, 64)
self.fc3 = nn.Linear(64, 10)
self.fc4 = nn.Linear(10,3)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
classify = Classification().double()
classify = classify.to("cuda")
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.SGD(classify.parameters(), lr=0.01, momentum=0.9)
optimizer_focus = optim.SGD(focus_net.parameters(), lr=0.01, momentum=0.9)
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
nos_epochs = 200
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 60
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.05):
break;
if epoch % 5 == 0:
# focus_net.eval()
# classify.eval()
col1.append(epoch+1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
print('Finished Training')
# torch.save(focus_net.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_focus_net.pt")
# torch.save(classify.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_classify.pt")
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig("train_ftpt.pdf", bbox_inches='tight')
plt.show()
df_test
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig("test_ftpt.pdf", bbox_inches='tight')
plt.show()
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
max_alpha =[]
alpha_ftpt=[]
argmax_more_than_half=0
argmax_less_than_half=0
for i, data in enumerate(test_loader):
inputs, labels,_ = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
alphas, avg = focus_net(inputs)
outputs = classify(avg)
mx,_ = torch.max(alphas,1)
max_alpha.append(mx.cpu().detach().numpy())
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if (focus == fore_idx[j] and predicted[j] == labels[j]):
alpha_ftpt.append(alphas[j][focus].item())
max_alpha = np.concatenate(max_alpha,axis=0)
print(max_alpha.shape)
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(max_alpha,bins=50,color ="c")
plt.title("alpha values histogram")
plt.savefig("alpha_hist.pdf")
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(np.array(alpha_ftpt),bins=50,color ="c")
plt.title("alpha values in ftpt")
plt.savefig("alpha_hist_ftpt.pdf")
```
| github_jupyter |
# Analyzing wind and stress measurements from IRGASON
## Experiment parameters
* Date: 2019-12-03
* Tank: SUSTAIN
* Start time: 17:30 UTC
* Wind only, no paddle
* Fan from 0 to 60 Hz in 5 Hz intervals, each run 600 s (10 minutes)
* Fresh water
* Mean water depth: 0.8 m
**Note: Experiment stopped, last run being 50 Hz.**
## Loading the data
```
from datetime import datetime, timedelta
import glob
import matplotlib.pyplot as plt
import numpy as np
from scipy.signal import detrend
from sustain_drag_2020.irgason import read_irgason_from_toa5
import warnings
warnings.filterwarnings('ignore')
# data parameters
DATAPATH = '/home/milan/Work/sustain/data/sustain-nsf-2019/20191203'
irgason_files = glob.glob(DATAPATH + '/TOA5_SUSTAIN_Wind.FAST*.dat')
# experiment parameters
start_time = datetime(2019, 12, 3, 17, 30)
fan = range(0, 55, 5)
run_seconds = 600
# read IRGASON data from TOA5 files
time, irg1, irg2 = read_irgason_from_toa5(irgason_files)
irg1
irg2
```
## Raw velocity data
```
fig = plt.figure(figsize=(12, 6))
plt.plot(time, irg1['u'], 'b-', lw=0.1)
plt.grid()
plt.xlabel('Time [UTC]')
plt.ylabel('Velocity [m/s]')
plt.title('u-component of wind, IRGASON 1')
```
This is raw u-velocity (along-tank direction), from IRGASON 1 (short fetch). Comments:
* Some dropouts in data first appear at the beginning of the 50 Hz run, and some more at 55 and 60 Hz
* Mean wind seems to have a small negative trend, especially in higher winds. Perhaps due to water loss in the tank -- less water -> more air -> wind weakens over time?
```
fig = plt.figure(figsize=(12, 6))
plt.plot(time, irg1['v'], 'g-', lw=0.1)
plt.grid()
plt.xlabel('Time [UTC]')
plt.ylabel('Velocity [m/s]')
plt.ylim(-10, 10)
plt.title('v-component of wind, IRGASON 1')
```
Raw values of cross-tank velocity.
* Mean is biased and positive, which means the instrument is not perfectly aligned in the along-tank direction. This is fine -- cross-tank velocity will go into the horizontal velocity.
* Yes, some dropouts in the data here as well, although the values are less extreme than in the along-tank component.
```
fig = plt.figure(figsize=(12, 6))
plt.plot(time, irg1['w'], 'r-', lw=0.1)
plt.grid()
plt.xlabel('Time [UTC]')
plt.ylabel('Velocity [m/s]')
plt.ylim(-5, 5)
plt.title('w-component of wind, IRGASON 1')
```
## Cleaning up and rotating the data
We can perform some basic cleaning of the data by setting some a priori maximum gust values and limiting the data at each fan speed to +/- gust range from the mean. This takes care of extreme velocities due to spray but is not ideal.
```
def rotate(u, w, th):
"""Rotates the vector (u, w) by angle th."""
ur = np.cos(th) * u + np.sin(th) * w
wr = -np.sin(th) * u + np.cos(th) * w
return ur, wr
def momentum_flux(irg, time, t0, t1):
U, Ustd, Wstd, uw = [], [], [], []
max_u_gust = 10
max_w_gust = 5
for n in range(len(fan)):
mask = (time >= t0[n]) & (time <= t1[n])
u, v, w = irg['u'][mask][:], irg['v'][mask][:], irg['w'][mask][:]
# clean up
um, vm, wm = np.nanmean(u), np.nanmean(v), np.nanmean(w)
u[u > um + max_u_gust] = um + max_u_gust
u[u < um - max_u_gust] = um - max_u_gust
v[v > vm + max_u_gust] = vm + max_u_gust
v[v < vm - max_u_gust] = vm - max_u_gust
w[w > wm + max_w_gust] = wm + max_w_gust
w[w < wm - max_w_gust] = wm - max_w_gust
# horizontal velocity
u = np.sqrt(u**2 + v**2)
# rotate
angle = np.arctan2(np.nanmean(w), np.nanmean(u))
u, w = rotate(u, w, angle)
# time average
um, wm = np.nanmean(u), np.nanmean(w)
up, wp = u - um, w - wm
U.append(um)
Ustd.append(np.nanstd(u))
Wstd.append(np.nanstd(w))
uw.append(np.nanmean(up * wp))
return np.array(U), np.array(Ustd), np.array(Wstd), np.array(uw)
# 9-minute time windows for each run;
# we exclude the first minute (thus 9 and not 10) due to fan spinup
t0 = [start_time + timedelta(seconds=n * run_seconds + 60)
for n in range(len(fan))]
t1 = [start_time + timedelta(seconds=(n + 1) * run_seconds)
for n in range(len(fan))]
U1, Ustd1, Wstd1, uw1 = momentum_flux(irg1, time, t0, t1)
U2, Ustd2, Wstd2, uw2 = momentum_flux(irg2, time, t0, t1)
fig = plt.figure(figsize=(8, 6))
plt.plot(fan, U1, color='tab:blue', marker='o', label='IRGASON 1')
for n in range(U1.size):
plt.plot([fan[n], fan[n]], [U1[n]-Ustd1[n], U1[n]+Ustd1[n]], color='tab:blue')
plt.plot(fan, U2, color='tab:orange', marker='o', label='IRGASON 2')
for n in range(U2.size):
plt.plot([fan[n], fan[n]], [U2[n]-Ustd2[n], U2[n]+Ustd2[n]], color='tab:orange')
plt.legend(loc='upper left', fancybox=True, shadow=True)
plt.grid()
plt.xlabel('Fan [Hz]')
plt.ylabel('Wind speed [m/s]')
plt.title('Mean wind speed vs. fan')
# exclude 0 Hz data and some questionable data in high winds
uw1[0] = np.nan
uw2[0] = np.nan
uw2[-1:] = np.nan
fig = plt.figure(figsize=(8, 6))
plt.plot(U1, uw1, color='tab:blue', marker='o', label='IRGASON 1')
plt.plot(U2, uw2, color='tab:orange', marker='o', label='IRGASON 2')
plt.legend(loc='upper left', fancybox=True, shadow=True)
plt.grid()
plt.xlim(0, 40)
plt.ylim(0, 0.4)
plt.xlabel('Wind speed [m/s]')
plt.ylabel(r"$-\overline{u'w'}$ [$m^2/s^2$]")
plt.title('Stress vs. mean wind speed')
```
| github_jupyter |
# 📝 Exercise M4.01
The aim of this exercise is two-fold:
* understand the parametrization of a linear model;
* quantify the fitting accuracy of a set of such models.
We will reuse part of the code of the course to:
* load data;
* create the function representing a linear model.
## Prerequisites
### Data loading
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">If you want a deeper overview regarding this dataset, you can refer to the
Appendix - Datasets description section at the end of this MOOC.</p>
</div>
```
import pandas as pd
penguins = pd.read_csv("../datasets/penguins_regression.csv")
feature_name = "Flipper Length (mm)"
target_name = "Body Mass (g)"
data, target = penguins[[feature_name]], penguins[target_name]
```
### Model definition
```
def linear_model_flipper_mass(
flipper_length, weight_flipper_length, intercept_body_mass
):
"""Linear model of the form y = a * x + b"""
body_mass = weight_flipper_length * flipper_length + intercept_body_mass
return body_mass
```
## Main exercise
Define a vector `weights = [...]` and a vector `intercepts = [...]` of
the same length. Each pair of entries `(weights[i], intercepts[i])` tags a
different model. Use these vectors along with the vector
`flipper_length_range` to plot several linear models that could possibly
fit our data. Use the above helper function to visualize both the models and
the real samples.
```
import numpy as np
flipper_length_range = np.linspace(data.min(), data.max(), num=300)
# Write your code here.
```
In the previous question, you were asked to create several linear models.
The visualization allowed you to qualitatively assess if a model was better
than another.
Now, you should come up with a quantitative measure which indicates the
goodness of fit of each linear model and allows you to select the best model.
Define a function `goodness_fit_measure(true_values, predictions)` that takes
as inputs the true target values and the predictions and returns a single
scalar as output.
```
# Write your code here.
```
You can now copy and paste the code below to show the goodness of fit for
each model.
```python
for model_idx, (weight, intercept) in enumerate(zip(weights, intercepts)):
target_predicted = linear_model_flipper_mass(data, weight, intercept)
print(f"Model #{model_idx}:")
print(f"{weight:.2f} (g / mm) * flipper length + {intercept:.2f} (g)")
print(f"Error: {goodness_fit_measure(target, target_predicted):.3f}\n")
```
```
# Write your code here.
```
| github_jupyter |
# Defining Disruption Budgets for Seldon Deployments
## Prerequisites
* A kubernetes cluster with kubectl configured
* pygmentize
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Setup-Cluster) with [Ambassador Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Ambassador) and [Install Seldon Core](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Install-Seldon-Core). Instructions [also online](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html).
```
!kubectl create namespace seldon
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
```
## Create model with Pod Disruption Budget
To create a model with a Pod Disruption Budget, it is first important to understand how you would like your application to respond to [voluntary disruptions](https://kubernetes.io/docs/concepts/workloads/pods/disruptions/#voluntary-and-involuntary-disruptions). Depending on the type of disruption budgeting your application needs, you will either define either of the following:
* `minAvailable` which is a description of the number of pods from that set that must still be available after the eviction, even in the absence of the evicted pod. `minAvailable` can be either an absolute number or a percentage.
* `maxUnavailable` which is a description of the number of pods from that set that can be unavailable after the eviction. It can be either an absolute number or a percentage.
The full SeldonDeployment spec is shown below.
```
!pygmentize model_with_pdb.yaml
!kubectl apply -f model_with_pdb.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=seldon-model -o jsonpath='{.items[0].metadata.name}')
```
## Validate Disruption Budget Configuration
```
import json
def getPdbConfig():
dp=!kubectl get pdb seldon-model-example-0-classifier -o json
dp=json.loads("".join(dp))
return dp["spec"]["maxUnavailable"]
assert getPdbConfig() == 2
!kubectl get pods,deployments,pdb
```
## Update Disruption Budget and Validate Change
Next, we'll update the maximum number of unavailable pods and check that the PDB is properly updated to match.
```
!pygmentize model_with_patched_pdb.yaml
!kubectl apply -f model_with_patched_pdb.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=seldon-model -o jsonpath='{.items[0].metadata.name}')
assert getPdbConfig() == 1
```
## Clean Up
```
!kubectl get pods,deployments,pdb
!kubectl delete -f model_with_patched_pdb.yaml
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import os
import GPy
import matplotlib.pyplot as plt
%matplotlib inline
%load_ext autoreload
%autoreload 2
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import VarianceThreshold
from sklearn.model_selection import train_test_split
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF, RationalQuadratic, DotProduct, ConstantKernel as C
from sklearn.ensemble import GradientBoostingRegressor
DATADIR = '../data'
TRAIN_SIZE = 0.1
import sys
sys.path.append('../')
from dispersant_screener.utils import plot_parity
from dispersant_screener.gp import build_model, _get_matern_52_kernel, _get_matern_32_kernel
from dispersant_screener.definitions import FEATURES
df_full_factorial_feat = pd.read_csv(os.path.join(DATADIR, 'new_features_full_random.csv'))[FEATURES].values
a2 = pd.read_csv(os.path.join(DATADIR, 'b1-b21_random_virial_large_fit2.csv'))['A2_normalized'].values
deltaGMax = pd.read_csv(os.path.join(DATADIR, 'b1-b21_random_virial_large_new.csv'))['A2_normalized'].values
gibbs = pd.read_csv(os.path.join(DATADIR, 'b1-b21_random_deltaG.csv'))['deltaGmin'].values
gibbs_max = pd.read_csv(os.path.join(DATADIR, 'b1-b21_random_virial_large_fit2.csv'))['deltaGmax'].values
force_max = pd.read_csv(os.path.join(DATADIR, 'b1-b21_random_virial_large_fit2.csv'))['F_repel_max'].values
rg = pd.read_csv(os.path.join(DATADIR, 'rg_results.csv'))['Rg'].values
y = np.hstack([rg.reshape(-1,1), gibbs.reshape(-1,1), gibbs_max.reshape(-1,1), force_max.reshape(-1,1), a2.reshape(-1,1)])
assert len(df_full_factorial_feat) == len(a2) == len(gibbs) == len(y)
X_train, X_test, y_train, y_test = train_test_split(df_full_factorial_feat, y, train_size=TRAIN_SIZE)
vt = VarianceThreshold(0.1)
X_train = vt.fit_transform(X_train)
X_test = vt.transform(X_test)
feat_scaler = StandardScaler()
X_train = feat_scaler.fit_transform(X_train)
X_test = feat_scaler.transform(X_test)
NFEAT = df_full_factorial_feat.shape[1]
model = build_model(X_train, y_train)
model.optimize()
prediction_test, var = model.predict(X_test)
prediction_train, vartrain = model.predict(X_train)
plot_parity([(y_test[:,0], prediction_test, var),
(y_train[:,0], prediction_train, vartrain)])
model1 = build_model(X_train, y_train, 1)
model1.optimize()
prediction_test1, var1 = model1.predict(X_test)
prediction_train1, vartrain1 = model1.predict(X_train)
plot_parity([(y_test[:,1], prediction_test1, var1),
(y_train[:,1], prediction_train1, vartrain1)])
kern = GPy.kern.RatQuad(X_train.shape[1], ARD=True)
model2 = build_model(X_train, y_train, 2, kern)
model2.optimize()
prediction_test2, var2 = model2.predict(X_test)
prediction_train2, vartrain2 = model2.predict(X_train)
plot_parity([(y_test[:,2], prediction_test2, var2),
(y_train[:,2], prediction_train2, vartrain2)])
model3 = build_model(X_train, y_train, 3)
model3.optimize()
prediction_test3, var3 = model3.predict(X_test)
prediction_train3, vartrain3 = model3.predict(X_train)
plot_parity([(y_test[:,3], prediction_test3, var3),
(y_train[:,3], prediction_train3, vartrain3)])
kern = GPy.kern.Matern32(X_train.shape[1], ARD=True)
model4 = build_model(X_train, y_train, 4, kernel=kern)
model4.optimize()
prediction_test4, var4 = model4.predict(X_test)
prediction_train4, vartrain4 = model4.predict(X_train)
plot_parity([(y_test[:,4], prediction_test4, var4),
(y_train[:,4], prediction_train4, vartrain4)])
```
## Get a baseline
### Gibbs
```
adaboost = GradientBoostingRegressor(n_estimators=500)
adaboost.fit(X_train, y_train[:,1])
prediction_test = adaboost.predict(X_test)
prediction_train = adaboost.predict(X_train)
plot_parity(prediction_test, y_test[:,1])
plot_parity(prediction_train, y_train[:,1])
```
## Virial
```
adaboost = GradientBoostingRegressor(n_estimators=500)
adaboost.fit(X_train, y_train[:,0])
prediction_test = adaboost.predict(X_test)
prediction_train = adaboost.predict(X_train)
plot_parity(prediction_test, y_test[:,0])
plot_parity(prediction_train, y_train[:,0])
```
## GP coregionalized
```
import GPy
K = GPy.kern.Matern32(NFEAT)
icm = GPy.util.multioutput.ICM(input_dim=NFEAT,num_outputs=2,kernel=K)
m = GPy.models.GPCoregionalizedRegression([X_train, X_train],
[y_train[:,0].reshape(-1,1), y_train[:,1].reshape(-1,1)],
kernel=icm)
m.optimize()
import matplotlib as mpl
newX = np.hstack([X_test,0 * np.ones_like(X_test)])
mu_c0, var_c0 =m.predict(newX,Y_metadata={'output_index':0 * np.ones((newX.shape[0],1)).astype(int)})
newX = np.hstack([X_test, 1 * np.ones_like(X_test)])
mu_c1, var_c1 =m.predict(newX,Y_metadata={'output_index': 1 * np.ones((newX.shape[0],1)).astype(int)})
get_metrics(mu_c0, y_test[:,0])
get_metrics(mu_c1, y_test[:,1])
def plot_parity(y_pred0, y_true0, var0, y_pred1, y_true1, var1, outname=None):
fig, ax = plt.subplots(1,2)
# now plot both limits against eachother
ax[0].scatter(y_true0, y_pred0, c=var0, cmap=plt.cm.coolwarm, s=.3)
ax[1].scatter(y_true1, y_pred1, c=var1, cmap=plt.cm.coolwarm, s=.3)
for a in ax:
a.spines['left'].set_smart_bounds(True)
a.spines['bottom'].set_smart_bounds(True)
a.set_xlabel(r'$y_{true}$')
lims = [
np.min([a.get_xlim(), a.get_ylim()]), # min of both axes
np.max([a.get_xlim(), a.get_ylim()]), # max of both axes
]
a.plot(lims, lims, 'k-', alpha=0.75, zorder=0)
ax[0].set_ylabel(r'$\hat{y}$')
fig.tight_layout()
if outname is not None:
fig.savefig(outname, bbox_inches='tight')
plot_parity(mu, y_test[:,0], var, mu2, y_test[:,1], var1, 'test.pdf')
```
## Compare against simple GP models
```
m_0 = GPy.models.GPRegression(X_train, y_train[:,0].reshape(-1,1),
kernel=K)
m_0.optimize()
m_1 = GPy.models.GPRegression(X_train, y_train[:,1].reshape(-1,1),
kernel=K)
m_1.optimize()
mu1, var1 = m_1.predict(X_test)
mu0, var0 = m_0.predict(X_test)
plot_parity(mu0, y_test[:,0], var0, mu1, y_test[:,1], var1, 'test.pdf')
import sys
sys.path.append('..')
from dispersant_screener.utils import get_metrics
get_metrics(mu0, y_test[:,0])
get_metrics(mu1, y_test[:,1])
def get_variance_descriptors(var):
return {
'max_var': np.max(var),
'min_var': np.min(var),
'mean_var': np.mean(var),
'median_var': np.median(var),
'std_var': np.std(var)
}
get_variance_descriptors(var0)
get_variance_descriptors(var_c0)
get_variance_descriptors(var1)
get_variance_descriptors(var_c1)
```
| github_jupyter |
**Copyright 2018 The TensorFlow Authors.**
Licensed under the Apache License, Version 2.0 (the "License");
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Get Started with Eager Execution
Note: you can run **[this notebook, live in Google Colab](https://colab.research.google.com/github/tensorflow/models/blob/master/samples/core/get_started/eager.ipynb)** with zero setup.
This tutorial describes how to use machine learning to *categorize* Iris flowers by species. It uses [TensorFlow](https://www.tensorflow.org)'s eager execution to (1) build a *model*, (2) *train* the model on example data, and (3) use the model to make *predictions* on unknown data. Machine learning experience isn't required to follow this guide, but you'll need to read some Python code.
## TensorFlow programming
There many [TensorFlow APIs](https://www.tensorflow.org/api_docs/python/) available, but we recommend starting with these high-level TensorFlow concepts:
* Enable an [eager execution](https://www.tensorflow.org/programmers_guide/eager) development environment,
* Import data with the [Datasets API](https://www.tensorflow.org/programmers_guide/datasets),
* Build models and layers with TensorFlow's [Keras API](https://keras.io/getting-started/sequential-model-guide/).
This tutorial shows these APIs and is structured like many other TensorFlow programs:
1. Import and parse the data sets.
2. Select the type of model.
3. Train the model.
4. Evaluate the model's effectiveness.
5. Use the trained model to make predictions.
To learn more about using TensorFlow, see the [Getting Started guide](https://www.tensorflow.org/get_started/) and the [example tutorials](https://www.tensorflow.org/tutorials/). If you'd like to learn about the basics of machine learning, consider taking the [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/).
## Run the notebook
This tutorial is available as an interactive [Colab notebook](https://colab.research.google.com) for you to run and change the Python code directly in the browser. The notebook handles setup and dependencies while you "play" cells to execute the code blocks. This is a fun way to explore the program and test ideas. If you are unfamiliar with Python notebook environments, there are a couple of things to keep in mind:
1. Executing code requires connecting to a runtime environment. In the Colab notebook menu, select *Runtime > Connect to runtime...*
2. Notebook cells are arranged sequentially to gradually build the program. Typically, later code cells depend on prior code cells, though you can always rerun a code block. To execute the entire notebook in order, select *Runtime > Run all*. To rerun a code cell, select the cell and click the *play icon* on the left.
## Setup program
### Install the latest version of TensorFlow
This tutorial uses eager execution, which is available in [TensorFlow 1.7](https://www.tensorflow.org/install/). (You may need to restart the runtime after upgrading.)
```
!pip install --upgrade tensorflow
```
### Configure imports and eager execution
Import the required Python modules, including TensorFlow, and enable eager execution for this program. Eager execution makes TensorFlow evaluate operations immediately, returning concrete values instead of creating a [computational graph](https://www.tensorflow.org/programmers_guide/graphs) that is executed later. If you are used to a REPL or the `python` interactive console, you'll feel at home.
Once eager execution is enabled, it *cannot* be disabled within the same program. See the [eager execution guide](https://www.tensorflow.org/programmers_guide/eager) for more details.
```
from __future__ import absolute_import, division, print_function
import os
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow.contrib.eager as tfe
tf.enable_eager_execution()
print("TensorFlow version: {}".format(tf.VERSION))
print("Eager execution: {}".format(tf.executing_eagerly()))
```
## The Iris classification problem
Imagine you are a botanist seeking an automated way to categorize each Iris flower you find. Machine learning provides many algorithms to statistically classify flowers. For instance, a sophisticated machine learning program could classify flowers based on photographs. Our ambitions are more modest—we're going to classify Iris flowers based on the length and width measurements of their [sepals](https://en.wikipedia.org/wiki/Sepal) and [petals](https://en.wikipedia.org/wiki/Petal).
The Iris genus entails about 300 species, but our program will classify only the following three:
* Iris setosa
* Iris virginica
* Iris versicolor
<table>
<tr><td>
<img src="https://www.tensorflow.org/images/iris_three_species.jpg"
alt="Petal geometry compared for three iris species: Iris setosa, Iris virginica, and Iris versicolor">
</td></tr>
<tr><td align="center">
<b>Figure 1.</b> <a href="https://commons.wikimedia.org/w/index.php?curid=170298">Iris setosa</a> (by <a href="https://commons.wikimedia.org/wiki/User:Radomil">Radomil</a>, CC BY-SA 3.0), <a href="https://commons.wikimedia.org/w/index.php?curid=248095">Iris versicolor</a>, (by <a href="https://commons.wikimedia.org/wiki/User:Dlanglois">Dlanglois</a>, CC BY-SA 3.0), and <a href="https://www.flickr.com/photos/33397993@N05/3352169862">Iris virginica</a> (by <a href="https://www.flickr.com/photos/33397993@N05">Frank Mayfield</a>, CC BY-SA 2.0).<br/>
</td></tr>
</table>
Fortunately, someone has already created a [data set of 120 Iris flowers](https://en.wikipedia.org/wiki/Iris_flower_data_set) with the sepal and petal measurements. This is a classic dataset that is popular for beginner machine learning classification problems.
## Import and parse the training dataset
We need to download the dataset file and convert it to a structure that can be used by this Python program.
### Download the dataset
Download the training dataset file using the [tf.keras.utils.get_file](https://www.tensorflow.org/api_docs/python/tf/keras/utils/get_file) function. This returns the file path of the downloaded file.
```
train_dataset_url = "http://download.tensorflow.org/data/iris_training.csv"
train_dataset_fp = tf.keras.utils.get_file(fname=os.path.basename(train_dataset_url),
origin=train_dataset_url)
print("Local copy of the dataset file: {}".format(train_dataset_fp))
```
### Inspect the data
This dataset, `iris_training.csv`, is a plain text file that stores tabular data formatted as comma-separated values (CSV). Use the `head -n5` command to take a peak at the first five entries:
```
!head -n5 {train_dataset_fp}
```
From this view of the dataset, we see the following:
1. The first line is a header containing information about the dataset:
* There are 120 total examples. Each example has four features and one of three possible label names.
2. Subsequent rows are data records, one *[example](https://developers.google.com/machine-learning/glossary/#example)* per line, where:
* The first four fields are *[features](https://developers.google.com/machine-learning/glossary/#feature)*: these are characteristics of an example. Here, the fields hold float numbers representing flower measurements.
* The last column is the *[label](https://developers.google.com/machine-learning/glossary/#label)*: this is the value we want to predict. For this dataset, it's an integer value of 0, 1, or 2 that corresponds to a flower name.
Each label is associated with string name (for example, "setosa"), but machine learning typically relies on numeric values. The label numbers are mapped to a named representation, such as:
* `0`: Iris setosa
* `1`: Iris versicolor
* `2`: Iris virginica
For more information about features and labels, see the [ML Terminology section of the Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/framing/ml-terminology).
### Parse the dataset
Since our dataset is a CSV-formatted text file, we'll parse the feature and label values into a format our Python model can use. Each line—or row—in the file is passed to the `parse_csv` function which grabs the first four feature fields and combines them into a single tensor. Then, the last field is parsed as the label. The function returns *both* the `features` and `label` tensors:
```
def parse_csv(line):
example_defaults = [[0.], [0.], [0.], [0.], [0]] # sets field types
parsed_line = tf.decode_csv(line, example_defaults)
# First 4 fields are features, combine into single tensor
features = tf.reshape(parsed_line[:-1], shape=(4,))
# Last field is the label
label = tf.reshape(parsed_line[-1], shape=())
return features, label
```
### Create the training tf.data.Dataset
TensorFlow's [Dataset API](https://www.tensorflow.org/programmers_guide/datasets) handles many common cases for feeding data into a model. This is a high-level API for reading data and transforming it into a form used for training. See the [Datasets Quick Start guide](https://www.tensorflow.org/get_started/datasets_quickstart) for more information.
This program uses [tf.data.TextLineDataset](https://www.tensorflow.org/api_docs/python/tf/data/TextLineDataset) to load a CSV-formatted text file and is parsed with our `parse_csv` function. A [tf.data.Dataset](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) represents an input pipeline as a collection of elements and a series of transformations that act on those elements. Transformation methods are chained together or called sequentially—just make sure to keep a reference to the returned `Dataset` object.
Training works best if the examples are in random order. Use `tf.data.Dataset.shuffle` to randomize entries, setting `buffer_size` to a value larger than the number of examples (120 in this case). To train the model faster, the dataset's *[batch size](https://developers.google.com/machine-learning/glossary/#batch_size)* is set to `32` examples to train at once.
```
train_dataset = tf.data.TextLineDataset(train_dataset_fp)
train_dataset = train_dataset.skip(1) # skip the first header row
train_dataset = train_dataset.map(parse_csv) # parse each row
train_dataset = train_dataset.shuffle(buffer_size=1000) # randomize
train_dataset = train_dataset.batch(32)
# View a single example entry from a batch
features, label = tfe.Iterator(train_dataset).next()
print("example features:", features[0])
print("example label:", label[0])
```
## Select the type of model
### Why model?
A *[model](https://developers.google.com/machine-learning/crash-course/glossary#model)* is the relationship between features and the label. For the Iris classification problem, the model defines the relationship between the sepal and petal measurements and the predicted Iris species. Some simple models can be described with a few lines of algebra, but complex machine learning models have a large number of parameters that are difficult to summarize.
Could you determine the relationship between the four features and the Iris species *without* using machine learning? That is, could you use traditional programming techniques (for example, a lot of conditional statements) to create a model? Perhaps—if you analyzed the dataset long enough to determine the relationships between petal and sepal measurements to a particular species. And this becomes difficult—maybe impossible—on more complicated datasets. A good machine learning approach *determines the model for you*. If you feed enough representative examples into the right machine learning model type, the program will figure out the relationships for you.
### Select the model
We need to select the kind of model to train. There are many types of models and picking a good one takes experience. This tutorial uses a neural network to solve the Iris classification problem. *[Neural networks](https://developers.google.com/machine-learning/glossary/#neural_network)* can find complex relationships between features and the label. It is a highly-structured graph, organized into one or more *[hidden layers](https://developers.google.com/machine-learning/glossary/#hidden_layer)*. Each hidden layer consists of one or more *[neurons](https://developers.google.com/machine-learning/glossary/#neuron)*. There are several categories of neural networks and this program uses a dense, or *[fully-connected neural network](https://developers.google.com/machine-learning/glossary/#fully_connected_layer)*: the neurons in one layer receive input connections from *every* neuron in the previous layer. For example, Figure 2 illustrates a dense neural network consisting of an input layer, two hidden layers, and an output layer:
<table>
<tr><td>
<img src="https://www.tensorflow.org/images/custom_estimators/full_network.png"
alt="A diagram of the network architecture: Inputs, 2 hidden layers, and outputs">
</td></tr>
<tr><td align="center">
<b>Figure 2.</b> A neural network with features, hidden layers, and predictions.<br/>
</td></tr>
</table>
When the model from Figure 2 is trained and fed an unlabeled example, it yields three predictions: the likelihood that this flower is the given Iris species. This prediction is called *[inference](https://developers.google.com/machine-learning/crash-course/glossary#inference)*. For this example, the sum of the output predictions are 1.0. In Figure 2, this prediction breaks down as: `0.03` for *Iris setosa*, `0.95` for *Iris versicolor*, and `0.02` for *Iris virginica*. This means that the model predicts—with 95% probability—that an unlabeled example flower is an *Iris versicolor*.
### Create a model using Keras
The TensorFlow [tf.keras](https://www.tensorflow.org/api_docs/python/tf/keras) API is the preferred way to create models and layers. This makes it easy to build models and experiment while Keras handles the complexity of connecting everything together. See the [Keras documentation](https://keras.io/) for details.
The [tf.keras.Sequential](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) model is a linear stack of layers. Its constructor takes a list of layer instances, in this case, two [Dense](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense) layers with 10 nodes each, and an output layer with 3 nodes representing our label predictions. The first layer's `input_shape` parameter corresponds to the amount of features from the dataset, and is required.
```
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation="relu", input_shape=(4,)), # input shape required
tf.keras.layers.Dense(10, activation="relu"),
tf.keras.layers.Dense(3)
])
```
The *[activation function](https://developers.google.com/machine-learning/crash-course/glossary#activation_function)* determines the output of a single neuron to the next layer. This is loosely based on how brain neurons are connected. There are many [available activations](https://www.tensorflow.org/api_docs/python/tf/keras/activations), but [ReLU](https://developers.google.com/machine-learning/crash-course/glossary#ReLU) is common for hidden layers.
The ideal number of hidden layers and neurons depends on the problem and the dataset. Like many aspects of machine learning, picking the best shape of the neural network requires a mixture of knowledge and experimentation. As a rule of thumb, increasing the number of hidden layers and neurons typically creates a more powerful model, which requires more data to train effectively.
## Train the model
*[Training](https://developers.google.com/machine-learning/crash-course/glossary#training)* is the stage of machine learning when the model is gradually optimized, or the model *learns* the dataset. The goal is to learn enough about the structure of the training dataset to make predictions about unseen data. If you learn *too much* about the training dataset, then the predictions only work for the data it has seen and will not be generalizable. This problem is called *[overfitting](https://developers.google.com/machine-learning/crash-course/glossary#overfitting)*—it's like memorizing the answers instead of understanding how to solve a problem.
The Iris classification problem is an example of *[supervised machine learning](https://developers.google.com/machine-learning/glossary/#supervised_machine_learning)*: the model is trained from examples that contain labels. In *[unsupervised machine learning](https://developers.google.com/machine-learning/glossary/#unsupervised_machine_learning)*, the examples don't contain labels. Instead, the model typically finds patterns among the features.
### Define the loss and gradient function
Both training and evaluation stages need to calculate the model's *[loss](https://developers.google.com/machine-learning/crash-course/glossary#loss)*. This measures how off a model's predictions are from the desired label, in other words, how bad the model is performing. We want to minimize, or optimize, this value.
Our model will calculate its loss using the [tf.losses.sparse_softmax_cross_entropy](https://www.tensorflow.org/api_docs/python/tf/losses/sparse_softmax_cross_entropy) function which takes the model's prediction and the desired label. The returned loss value is progressively larger as the prediction gets worse.
```
def loss(model, x, y):
y_ = model(x)
return tf.losses.sparse_softmax_cross_entropy(labels=y, logits=y_)
def grad(model, inputs, targets):
with tfe.GradientTape() as tape:
loss_value = loss(model, inputs, targets)
return tape.gradient(loss_value, model.variables)
```
The `grad` function uses the `loss` function and the [tfe.GradientTape](https://www.tensorflow.org/api_docs/python/tf/contrib/eager/GradientTape) to record operations that compute the *[gradients](https://developers.google.com/machine-learning/crash-course/glossary#gradient)* used to optimize our model. For more examples of this, see the [eager execution guide](https://www.tensorflow.org/programmers_guide/eager).
### Create an optimizer
An *[optimizer](https://developers.google.com/machine-learning/crash-course/glossary#optimizer)* applies the computed gradients to the model's variables to minimize the `loss` function. You can think of a curved surface (see Figure 3) and we want to find its lowest point by walking around. The gradients point in the direction of steepest ascent—so we'll travel the opposite way and move down the hill. By iteratively calculating the loss and gradient for each batch, we'll adjust the model during training. Gradually, the model will find the best combination of weights and bias to minimize loss. And the lower the loss, the better the model's predictions.
<table>
<tr><td>
<img src="https://cs231n.github.io/assets/nn3/opt1.gif" width="70%"
alt="Optimization algorthims visualized over time in 3D space.">
</td></tr>
<tr><td align="center">
<b>Figure 3.</b> Optimization algorthims visualized over time in 3D space. (Source: <a href="http://cs231n.github.io/neural-networks-3/">Stanford class CS231n</a>, MIT License)<br/>
</td></tr>
</table>
TensorFlow has many [optimization algorithms](https://www.tensorflow.org/api_guides/python/train) available for training. This model uses the [tf.train.GradientDescentOptimizer](https://www.tensorflow.org/api_docs/python/tf/train/GradientDescentOptimizer) that implements the *[stochastic gradient descent](https://developers.google.com/machine-learning/crash-course/glossary#gradient_descent)* (SGD) algorithm. The `learning_rate` sets the step size to take for each iteration down the hill. This is a *hyperparameter* that you'll commonly adjust to achieve better results.
```
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
```
### Training loop
With all the pieces in place, the model is ready for training! A training loop feeds the dataset examples into the model to help it make better predictions. The following code block sets up these training steps:
1. Iterate each epoch. An epoch is one pass through the dataset.
2. Within an epoch, iterate over each example in the training `Dataset` grabbing its *features* (`x`) and *label* (`y`).
3. Using the example's features, make a prediction and compare it with the label. Measure the inaccuracy of the prediction and use that to calculate the model's loss and gradients.
4. Use an `optimizer` to update the model's variables.
5. Keep track of some stats for visualization.
6. Repeat for each epoch.
The `num_epochs` variable is the amount of times to loop over the dataset collection. Counter-intuitively, training a model longer does not guarantee a better model. `num_epochs` is a *[hyperparameter](https://developers.google.com/machine-learning/glossary/#hyperparameter)* that you can tune. Choosing the right number usually requires both experience and experimentation.
```
## Note: Rerunning this cell uses the same model variables
# keep results for plotting
train_loss_results = []
train_accuracy_results = []
num_epochs = 201
for epoch in range(num_epochs):
epoch_loss_avg = tfe.metrics.Mean()
epoch_accuracy = tfe.metrics.Accuracy()
# Training loop - using batches of 32
for x, y in tfe.Iterator(train_dataset):
# Optimize the model
grads = grad(model, x, y)
optimizer.apply_gradients(zip(grads, model.variables),
global_step=tf.train.get_or_create_global_step())
# Track progress
epoch_loss_avg(loss(model, x, y)) # add current batch loss
# compare predicted label to actual label
epoch_accuracy(tf.argmax(model(x), axis=1, output_type=tf.int32), y)
# end epoch
train_loss_results.append(epoch_loss_avg.result())
train_accuracy_results.append(epoch_accuracy.result())
if epoch % 50 == 0:
print("Epoch {:03d}: Loss: {:.3f}, Accuracy: {:.3%}".format(epoch,
epoch_loss_avg.result(),
epoch_accuracy.result()))
```
### Visualize the loss function over time
While it's helpful to print out the model's training progress, it's often *more helpful* to see this progress. [TensorBoard](https://www.tensorflow.org/programmers_guide/summaries_and_tensorboard) is a nice visualization tool that is packaged with TensorFlow, but we can create basic charts using the `mathplotlib` module.
Interpreting these charts takes some experience, but you really want to see the *loss* go down and the *accuracy* go up.
```
fig, axes = plt.subplots(2, sharex=True, figsize=(12, 8))
fig.suptitle('Training Metrics')
axes[0].set_ylabel("Loss", fontsize=14)
axes[0].plot(train_loss_results)
axes[1].set_ylabel("Accuracy", fontsize=14)
axes[1].set_xlabel("Epoch", fontsize=14)
axes[1].plot(train_accuracy_results)
plt.show()
```
## Evaluate the model's effectiveness
Now that the model is trained, we can get some statistics on its performance.
*Evaluating* means determining how effectively the model makes predictions. To determine the model's effectiveness at Iris classification, pass some sepal and petal measurements to the model and ask the model to predict what Iris species they represent. Then compare the model's prediction against the actual label. For example, a model that picked the correct species on half the input examples has an *[accuracy](https://developers.google.com/machine-learning/glossary/#accuracy)* of `0.5`. Figure 4 shows a slightly more effective model, getting 4 out of 5 predictions correct at 80% accuracy:
<table cellpadding="8" border="0">
<colgroup>
<col span="4" >
<col span="1" bgcolor="lightblue">
<col span="1" bgcolor="lightgreen">
</colgroup>
<tr bgcolor="lightgray">
<th colspan="4">Example features</th>
<th colspan="1">Label</th>
<th colspan="1" >Model prediction</th>
</tr>
<tr>
<td>5.9</td><td>3.0</td><td>4.3</td><td>1.5</td><td align="center">1</td><td align="center">1</td>
</tr>
<tr>
<td>6.9</td><td>3.1</td><td>5.4</td><td>2.1</td><td align="center">2</td><td align="center">2</td>
</tr>
<tr>
<td>5.1</td><td>3.3</td><td>1.7</td><td>0.5</td><td align="center">0</td><td align="center">0</td>
</tr>
<tr>
<td>6.0</td> <td>3.4</td> <td>4.5</td> <td>1.6</td> <td align="center">1</td><td align="center" bgcolor="red">2</td>
</tr>
<tr>
<td>5.5</td><td>2.5</td><td>4.0</td><td>1.3</td><td align="center">1</td><td align="center">1</td>
</tr>
<tr><td align="center" colspan="6">
<b>Figure 4.</b> An Iris classifier that is 80% accurate.<br/>
</td></tr>
</table>
### Setup the test dataset
Evaluating the model is similar to training the model. The biggest difference is the examples come from a separate *[test set](https://developers.google.com/machine-learning/crash-course/glossary#test_set)* rather than the training set. To fairly assess a model's effectiveness, the examples used to evaluate a model must be different from the examples used to train the model.
The setup for the test `Dataset` is similar to the setup for training `Dataset`. Download the CSV text file and parse that values, then give it a little shuffle:
```
test_url = "http://download.tensorflow.org/data/iris_test.csv"
test_fp = tf.keras.utils.get_file(fname=os.path.basename(test_url),
origin=test_url)
test_dataset = tf.data.TextLineDataset(test_fp)
test_dataset = test_dataset.skip(1) # skip header row
test_dataset = test_dataset.map(parse_csv) # parse each row with the function created earlier
test_dataset = test_dataset.shuffle(1000) # randomize
test_dataset = test_dataset.batch(32) # use the same batch size as the training set
```
### Evaluate the model on the test dataset
Unlike the training stage, the model only evaluates a single [epoch](https://developers.google.com/machine-learning/glossary/#epoch) of the test data. In the following code cell, we iterate over each example in the test set and compare the model's prediction against the actual label. This is used to measure the model's accuracy across the entire test set.
```
test_accuracy = tfe.metrics.Accuracy()
for (x, y) in tfe.Iterator(test_dataset):
prediction = tf.argmax(model(x), axis=1, output_type=tf.int32)
test_accuracy(prediction, y)
print("Test set accuracy: {:.3%}".format(test_accuracy.result()))
```
## Use the trained model to make predictions
We've trained a model and "proven" that it's good—but not perfect—at classifying Iris species. Now let's use the trained model to make some predictions on [unlabeled examples](https://developers.google.com/machine-learning/glossary/#unlabeled_example); that is, on examples that contain features but not a label.
In real-life, the unlabeled examples could come from lots of different sources including apps, CSV files, and data feeds. For now, we're going to manually provide three unlabeled examples to predict their labels. Recall, the label numbers are mapped to a named representation as:
* `0`: Iris setosa
* `1`: Iris versicolor
* `2`: Iris virginica
```
class_ids = ["Iris setosa", "Iris versicolor", "Iris virginica"]
predict_dataset = tf.convert_to_tensor([
[5.1, 3.3, 1.7, 0.5,],
[5.9, 3.0, 4.2, 1.5,],
[6.9, 3.1, 5.4, 2.1]
])
predictions = model(predict_dataset)
for i, logits in enumerate(predictions):
class_idx = tf.argmax(logits).numpy()
name = class_ids[class_idx]
print("Example {} prediction: {}".format(i, name))
```
These predictions look good!
To dig deeper into machine learning models, take a look at the TensorFlow [Programmer's Guide](https://www.tensorflow.org/programmers_guide/) and check out the [community](https://www.tensorflow.org/community/).
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 自动将代码升级到 TensorFlow 2
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://tensorflow.google.cn/guide/upgrade"> <img src="https://tensorflow.google.cn/images/tf_logo_32px.png"> View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/guide/upgrade.ipynb"> <img src="https://tensorflow.google.cn/images/colab_logo_32px.png"> Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/guide/upgrade.ipynb"> <img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png"> View source on GitHub</a>
</td>
<td>
<a target="_blank" href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/guide/upgrade.ipynb"> <img src="https://tensorflow.google.cn/images/download_logo_32px.png"> Download notebook</a>
</td>
</table>
TensorFlow 2.0 包含许多 API 变更,例如重新排序了参数,重命名了符号,更改了参数的默认值。手动执行所有这些修改可能很乏味,而且很容易出错。为了简化更改,尽可能地让您无缝过渡到 TF 2.0,TensorFlow 团队创建了 `tf_upgrade_v2` 实用工具,帮助您将旧版代码转换至新的 API。
注:TensorFlow 1.13 和更高版本(包括所有 TF 2.0 版本)会自动安装 `tf_upgrade_v2`。
典型的用法如下:
<pre class="devsite-terminal devsite-click-to-copy prettyprint lang-bsh">tf_upgrade_v2 \
--intree my_project/ \
--outtree my_project_v2/ \
--reportfile report.txt
</pre>
将现有 TensorFlow 1.x Python 脚本转换为 TensorFlow 2.0 脚本可以加快升级流程。
转换脚本会尽可能实现自动化处理,但仍有一些语法和样式变更无法通过脚本执行转换。
## 兼容性模块
某些 API 符号无法通过简单的字符串替换进行升级。为了确保代码在 TensorFlow 2.0 中仍受支持,升级脚本包含了一个 `compat.v1` 模块。该模块可将 TF 1.x 符号(如 `tf.foo`)替换为等效的 `tf.compat.v1.foo` 引用。虽然该兼容性模块效果不错,但我们仍建议人工校对替换,并尽快将代码迁移到 `tf.*` 命名空间(而不是 `tf.compat.v1` 命名空间)中的新 API。
由于 TensorFlow 2.x 模块弃用(例如,`tf.flags` 和 `tf.contrib`),切换到 `compat.v1` 无法解决某些更改。升级此代码可能需要其他库(例如,[`absl.flags`](https://github.com/abseil/abseil-py))或切换到 [tensorflow/addons](http://www.github.com/tensorflow/addons) 中的软件包。
## 推荐的升级流程
本指南的剩余部分演示如何使用升级脚本。虽然升级脚本的使用非常简单,我们仍强烈建议在以下流程中使用脚本:
1. **单元测试**:确保要升级的代码包含具有合理覆盖范围的单元测试套件。这是 Python 代码,该语言并不会帮助您避免各种类型的错误。同时为了与 TensorFlow 2.0 兼容,还要确保升级所有依赖项。
2. **安装 TensorFlow 1.14**:将 TensorFlow 升级到最新的 TensorFlow 1.x 版本(最低为 1.14 版本)。其中包括 `tf.compat.v2` 中的最终 TensorFlow 2.0 API。
3. **通过 1.14 版本进行测试**:确保此时可通过单元测试。在升级过程中,您将反复进行测试,因此,从无错误的代码开始非常重要。
4. **运行升级脚本**:对整个源代码树运行 `tf_upgrade_v2`(已包含测试)。这样可将代码升级为仅使用 TensorFlow 2.0 中所提供的符号的格式。被弃用的符号将通过 `tf.compat.v1` 进行访问。最终需要人工检查这些升级,但不是现在。
5. **通过 TensorFlow 1.14 运行转换的测试**:代码在 TensorFlow 1.14 中应该仍可以正常运行。再次运行单元测试。测试在此时产生任何错误都意味着升级脚本存在错误。[请通知我们](https://github.com/tensorflow/tensorflow/issues)。
6. **检查更新报告中的警告和错误**:该脚本会编写一个对需要复查的转换或需要执行的人工操作进行解释的报告文件。例如:contrib 的所有剩余实例需要通过人工操作删除。请查阅 [RFC 中的详细说明](https://github.com/tensorflow/community/blob/master/rfcs/20180907-contrib-sunset.md)。
7. **安装 TensorFlow 2.0**:此时应该可以安全切换到 TensorFlow 2.0
8. **使用 `v1.disable_v2_behavior` 进行测试**:使用测试主函数中的 `v1.disable_v2_behavior()` 重新运行测试产生的结果应与在 1.14 下运行时产生的结果相同。
9. **启用 V2 行为**:现在,使用 v2 API 已经成功通过了测试,不妨开始考虑启用 v2 行为。这可能需要执行一些更改,具体取决于代码编写方式。有关详细信息,请参阅[迁移指南](migrate.ipynb)。
## 使用升级脚本
### 设置
开始之前,请确保已安装 TensorlFlow 2.0。
```
import tensorflow as tf
print(tf.__version__)
```
克隆 [tensorflow/models](https://github.com/tensorflow/models) git 仓库,以便获得一些要测试的代码:
```
!git clone --branch r1.13.0 --depth 1 https://github.com/tensorflow/models
```
### 读取帮助
脚本应当随 TensorFlow 安装。下面是内置帮助命令:
```
!tf_upgrade_v2 -h
```
### TF1 代码示例
下面是一个简单的 TensorFlow 1.0 脚本示例:
```
!head -n 65 models/samples/cookbook/regression/custom_regression.py | tail -n 10
```
对于安装的 TensorFlow 2.0,它不会运行:
```
!(cd models/samples/cookbook/regression && python custom_regression.py)
```
### 单个文件
升级脚本可以在单个 Python 文件上运行:
```
!tf_upgrade_v2 \
--infile models/samples/cookbook/regression/custom_regression.py \
--outfile /tmp/custom_regression_v2.py
```
如果无法找到解决代码问题的方法,该脚本会打印错误消息。
### 目录树
典型项目(包括下面的简单示例)会使用远不止一个文件。通常需要升级整个软件包,所以该脚本也可以在目录树上运行:
```
# upgrade the .py files and copy all the other files to the outtree
!tf_upgrade_v2 \
--intree models/samples/cookbook/regression/ \
--outtree regression_v2/ \
--reportfile tree_report.txt
```
注意关于 `dataset.make_one_shot_iterator` 函数的一条警告。
现在,对于 TensorFlow 2.0,该脚本已经可以发挥作用:
请注意,凭借 `tf.compat.v1` 模块,转换的脚本在 TensorFlow 1.14 中也可以运行。
```
!(cd regression_v2 && python custom_regression.py 2>&1) | tail
```
## 详细报告
该脚本还会报告一个详细更改列表。在本例中,它发现了一个可能不安全的转换,因此在文件顶部包含了一条警告:
```
!head -n 20 tree_report.txt
```
再次注意关于 `Dataset.make_one_shot_iterator` 函数的一条警告。
在其他情况下,对于非常用更改,输出会解释原因:
```
%%writefile dropout.py
import tensorflow as tf
d = tf.nn.dropout(tf.range(10), 0.2)
z = tf.zeros_like(d, optimize=False)
!tf_upgrade_v2 \
--infile dropout.py \
--outfile dropout_v2.py \
--reportfile dropout_report.txt > /dev/null
!cat dropout_report.txt
```
以下是经过修改的文件内容,请注意脚本如何通过添加参数名来处理移动和重命名的参数:
```
!cat dropout_v2.py
```
更大的项目可能会包含一些错误,例如转换 DeepLab 模型:
```
!tf_upgrade_v2 \
--intree models/research/deeplab \
--outtree deeplab_v2 \
--reportfile deeplab_report.txt > /dev/null
```
它会生成输出文件:
```
!ls deeplab_v2
```
但是其中包含错误。该报告会帮助您找到确保代码可以正常运行所需要解决的错误。下面是前三个错误:
```
!cat deeplab_report.txt | grep -i models/research/deeplab | grep -i error | head -n 3
```
## “安全”模式
该转换脚本还有一种介入度相对较低的 `SAFETY` 模式。在此模式下,只需更改导入来使用 `tensorflow.compat.v1` 模块:
```
!cat dropout.py
!tf_upgrade_v2 --mode SAFETY --infile dropout.py --outfile dropout_v2_safe.py > /dev/null
!cat dropout_v2_safe.py
```
如您所见,这不会升级代码,但允许 TensorFlow 1 代码在 TensorFlow 2 中运行
## 注意事项
- 在运行此脚本之前,不要手动更新代码的某些部分。尤其是更改了参数顺序的函数(如 `tf.argmax` 或 `tf.batch_to_space`),否则会导致代码无法正确添加与现有代码匹配的关键字参数。
- 该脚本假定使用 `import tensorflow as tf` 导入 `tensorflow`。
- 该脚本不会更改参数顺序,但是会将关键字参数添加到本身已更改参数顺序的函数。
- 请查阅 [tf2up.ml](https://github.com/lc0/tf2up),找到一款方便的工具来升级 GitHub 仓库中的 Jupyter 笔记本和 Python 文件。
要报告升级脚本错误或提出功能请求,请在 [GitHub](https://github.com/tensorflow/tensorflow/issues) 上提交问题。如果您在测试 TensorFlow 2.0,我们非常希望了解您的反馈意见!请加入 [TF 2.0 测试社区](https://groups.google.com/a/tensorflow.org/forum/#!forum/testing),将您的问题和讨论发送到 [testing@tensorflow.org](mailto:testing@tensorflow.org)。
| github_jupyter |
```
import pandas as pd
from py2neo import Graph
graph = Graph('bolt://127.0.0.1:7687/db/data')
def to_df(result):
return pd.DataFrame([r.values() for r in result], columns=result.keys())
```
## Data exploration
```
to_df(graph.run("MATCH (n) RETURN distinct labels(n), count(*)"))
to_df(graph.run("MATCH (p:Publication) RETURN p.id AS id, p.title AS title"))
```
## Query relationships
```
pub = 'http://arxiv.org/abs/1906.02739v1'
query = """
MATCH (p1:Publication)-[r]-(n)
WHERE p1.id = {pub}
RETURN p1.title, type(r), n.name
"""
to_df(graph.run(query, pub=pub))
query = """
MATCH (p1:Publication)-[]-(rn)-[]-(p2:Publication)
WHERE p1.id = {pub}
RETURN p1.id, p1.title, count(rn) as n_relations, p2.id, p2.title
ORDER BY n_relations DESC
LIMIT 10
"""
to_df(graph.run(query, pub=pub))
```
## Compute similarities
```
query = """
MATCH (pub:Publication)-[]-(rel)
WITH {item: id(pub), categories: collect(id(rel))} as pubData
WITH collect(pubData) as data
CALL algo.similarity.jaccard(data, {concurrency:1, similarityCutoff: 0.1, write:false})
YIELD nodes, similarityPairs, write, writeRelationshipType, writeProperty, min, max, mean, stdDev, p25, p50, p75, p90, p95, p99, p999, p100
RETURN nodes, similarityPairs, write, writeRelationshipType, writeProperty, min, max, mean, p95
"""
to_df(graph.run(query, pub=pub))
query = """
MATCH (pub:Publication)-[]-(rel), (from:Publication {id: {pub}})
WITH {item: id(pub), categories: collect(id(rel))} as pubData, [id(from)] as sourceIds
WITH collect(pubData) as data, sourceIds
CALL algo.similarity.overlap.stream(data, {sourceIds: sourceIds, concurrency:1, similarityCutoff: 0.1})
YIELD item1, item2, similarity
WITH algo.getNodeById(item1) AS from, algo.getNodeById(item2) AS to, similarity
RETURN from.id, from.title, to.id, to.title, similarity
ORDER BY similarity DESC
LIMIT 100
"""
to_df(graph.run(query, pub=pub))
compared_to = [
'http://arxiv.org/abs/1906.01792v1',
'http://arxiv.org/abs/1906.02698v1',
'http://arxiv.org/abs/1812.01601v3',
'http://arxiv.org/abs/1906.02739v1',
]
query = """
MATCH (pub:Publication)-[]-(rel),
(from:Publication),
(to:Publication)
WHERE from.id = {pub} AND to.id IN {to}
WITH {item: id(pub), categories: collect(id(rel))} as pubData, [id(from)] as sourceIds, [id(to)] as targetIds
WITH collect(pubData) as data, sourceIds, targetIds
CALL algo.similarity.overlap.stream(data, {sourceIds: sourceIds, targetIds: targetIds, concurrency:1, similarityCutoff: 0.1})
YIELD item1, item2, similarity
WITH algo.getNodeById(item1) AS from, algo.getNodeById(item2) AS to, similarity
RETURN from.id, from.title, to.id, to.title, similarity
ORDER BY similarity DESC
LIMIT 100
"""
to_df(graph.run(query, pub=pub, to=compared_to))
```
| github_jupyter |
# Get The Top Subreddits By Subscriber Count
The top Subreddits does not include any that are private/quarantined. This does include NSFW subreddits.
Data:
* From: 2021-11-19
* Source: https://frontpagemetrics.com/list-all-subreddits
New Data Stored In: top_subreddits.json
```
import gzip
import praw
from config import *
from csv import reader
from collections import defaultdict
from prawcore.exceptions import Forbidden
# Will map a subreddit to its popularity
subreddit_popularity = defaultdict(int)
# The total amount of subreddits that will be returned
n = 500
# Open the subreddit popularity data
with gzip.open("../data/2021-11-19.csv.gz", "rt", errors="ignore") as read_obj:
# pass the file object to reader() to get the reader object
csv_reader = reader(read_obj)
# Skip the header
next(csv_reader)
# Iterate over each row in the csv using reader object
for row in csv_reader:
subreddit = row[0]
subscribers = row[-1]
subreddit_popularity[subreddit] = subscribers
# Sort the subreddits by their subscriber count
subreddit_popularity = sorted(subreddit_popularity.items(), key=lambda x: int(x[1]), reverse=True)
# Initialize PRAW API
reddit = praw.Reddit(
client_id=CLIENT_ID,
client_secret=CLIENT_SECRET,
user_agent=USER_AGENT,
username=REDDIT_USERNAME,
password=REDDIT_PASSWORD,
)
# Remove any private subreddits from the list
remove_list = []
# Only remove subreddits until n public subreddits are in the top
m = 0
for i in range(len(subreddit_popularity)):
# Edge Case: Stop once n amount of subreddits are in the top
if m == n:
break
subreddit = subreddit_popularity[i][0]
# Edge Case: Remove r/announcements due to barely 5 posts this year
if subreddit == "announcements":
remove_list.append(subreddit_popularity[i])
continue
# Try to find subreddits that are locked/quarantined
try:
for submission in reddit.subreddit(subreddit).top('year', limit=5):
pass
m += 1
except Forbidden:
print("Locked/Quarantined Subreddit: {}".format(subreddit))
remove_list.append(subreddit_popularity[i])
for subreddit in remove_list:
subreddit_popularity.remove(subreddit)
file = open("../data/top_subreddits.json", "w")
file.write("{\n")
for i, subreddit in enumerate(subreddit_popularity[:n]):
if i < (n - 1):
file.write("\t\"{}\":{},\n".format(subreddit[0], subreddit[1]))
else:
file.write("\t\"{}\":{}\n".format(subreddit[0], subreddit[1]))
file.write("}\n")
file.close()
```
| github_jupyter |
```
import numpy as np
import progressbar
import random
import os
import cv2
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import LabelBinarizer
import keras
from keras.applications import imagenet_utils
from keras.preprocessing.image import img_to_array
from keras.preprocessing.sequence import pad_sequences
import matplotlib.pyplot as plt
%matplotlib inline
from helpers import HDF5DatasetWriter
from helpers import Utils
output_path = "datasets/UCF-101-hdf5/fixed-sequence.hdf5"
root = 'datasets/UCF-101-frames-resize/'
include = ['ApplyEyeMakeup', 'ApplyLipstick', 'Archery', 'BabyCrawling',
'BalanceBeam', 'BandMarching', 'BaseballPitch', 'Basketball']
full_categories = [x[0] for x in os.walk(root) if x[0]][1:]
categories = [c for c in full_categories if c in [os.path.join(root, e) for e in include]]
categories
def get_image(img_path):
img = cv2.imread(img_path)
return img
img = get_image("datasets/UCF-101-frames-resize/ApplyEyeMakeup/v_ApplyEyeMakeup_g06_c04_3.jpg")
img.shape
data = []
for c, category in enumerate(categories):
images = [os.path.join(dp, f) for dp, dn, filenames
in os.walk(category) for f in filenames
if os.path.splitext(f)[1].lower() in ['.jpg','.png','.jpeg']]
current_squence_name = []
next_squence_name = images[0].split('/')[2].split('_')[1:3]
img_sequence = []
img_count = 0
img_max = 0
img_min = 60
for img_path in images:
squence_name = img_path.split('/')[2].split('_')[1:3]
current_squence_name = squence_name
if current_squence_name == next_squence_name:
img = get_image(img_path)
img_count = img_count + 1
if img_count <= 30:
img_sequence.append(img)
else:
next_squence_name = current_squence_name
data.append({'imgs':np.array(img_sequence), 'label':c})
img_sequence = []
if img_count < img_min:
img_min = img_count
if img_count > img_max:
img_max = img_count
img_count = 0
data.append({'imgs':np.array(img_sequence), 'label':c})
print(img_min, img_max)
data = np.array(data)
data[4]['imgs'].shape
x, y = np.array([t["imgs"] for t in data]), [t["label"] for t in data]
lb = LabelBinarizer()
y = lb.fit_transform(y)
def export_data(x, y, output_path, batch_size, buffer_size):
dataset = HDF5DatasetWriter(x.shape, y.shape, output_path, bufSize = buffer_size)
widgets = ["Exporting Data: ", progressbar.Percentage(), " ", progressbar.Bar(), " ", progressbar.ETA()]
pbar = progressbar.ProgressBar(maxval=x.shape[0], widgets=widgets).start()
for i in np.arange(0, x.shape[0], batch_size):
batchData = x[i:i + batch_size]
batchLabels = y[i:i + batch_size]
dataset.add(batchData, batchLabels)
pbar.update(i)
dataset.close()
pbar.finish()
export_data(x, y, output_path, 32, 1000)
```
| github_jupyter |
```
from google.colab import files
#
uploaded=files.upload()
for fn in uploaded.keys():
print('user uploaded file "{name}" with length {length} bytes'.format(name=fn,length=len(uploaded[fn])))
ls
import zipfile
zip_ref = zipfile.ZipFile('Images.zip', 'r')
zip_ref.extractall()
zip_ref.close()
!ls
import numpy as np
from scipy import misc
import glob
import cv2
TCClass = 0.0 ## non asd
TSClass = 1.0 ## asd
imgDimension = 100
f_handle = open('ASDDataset_Augmented.csv', 'a')#The CSV file where data records are appended to
from scipy.misc import imread
from scipy.misc import imresize
import os
ls=[]
folder='Images/TCImages'
files=os.listdir(folder)
files=list(map(lambda x: os.path.join(folder,x),files))
a=(len(files))
for i in range(a):
im = cv2.imread(files[i])
im=imresize(im,[imgDimension,imgDimension,3])
im = cv2.cvtColor(im, cv2.COLOR_BGR2GRAY)
im=im.flatten()
ls.append(im)
print(len(ls))
folder='Images/TSImages'
files=os.listdir(folder)
files=list(map(lambda x: os.path.join(folder,x),files))
a=(len(files))
for i in range(a):
im = cv2.imread(files[i])
im=imresize(im,[imgDimension,imgDimension,3])
im = cv2.cvtColor(im, cv2.COLOR_BGR2GRAY)
im=im.flatten()
ls.append(im)
import numpy as np
raw_data=np.array(ls,dtype=object)
print(raw_data.shape)
label_train=np.zeros(shape=[547,1])
label_train[329:547,:]=1
print(label_train[400])
import keras
from keras.datasets import cifar10
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras.models import model_from_json
num_classes=2
label_train= keras.utils.to_categorical(label_train, num_classes)
print(label_train.shape)
a=(raw_data.shape[0])
i=np.random.permutation(a)
x_train,y_train=raw_data[i],label_train[i]
print(y_train.shape,x_train.shape)
x_train=x_train.reshape(-1,100,100,1)
x_train.shape
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same',
input_shape=x_train.shape[1:]))
model.add(Activation('relu'))
#model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.1))
model.add(Conv2D(64, (3, 3), padding='same'))
model.add(Activation('relu'))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.1))
model.add(Flatten())
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
opt = keras.optimizers.rmsprop(lr=0.0001, decay=1e-6)
model.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
batch_size=16
epochs=15
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
shuffle=True)
img='TS024_19.png'
img = cv2.imread(img)
img=imresize(img,[imgDimension,imgDimension,3])
img = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
img=img.reshape(1,100,100,1)
#print(img.shape)
score = model.predict_classes(img, verbose=1)
model_json = model.to_json()
with open("model.json", "w") as json_file:
json_file.write(model_json)
model.save_weights("model.h5")
print("Saved model to disk")
json_file = open('model.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model1 = model_from_json(loaded_model_json)
loaded_model1.load_weights("model.h5")
print("Loaded model from disk")
loaded_model1.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
score = loaded_model.predict_classes(img, verbose=1)
from google.colab import files
files.download("model.h5")
files.download("model.json")
files.download("model.json")
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
%cd /content/drive/MyDrive/Neural_Tangent_Kernel/
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
from myrmsprop import MyRmsprop
from utils import plot_decision_boundary,attn_avg,plot_analysis
from synthetic_dataset import MosaicDataset1
from eval_model import calculate_attn_loss,analyse_data
%matplotlib inline
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
train_data = np.load("train_type4_data.npy",allow_pickle=True)
test_data = np.load("test_type4_data.npy",allow_pickle=True)
mosaic_list_of_images = train_data[0]["mosaic_list"]
mosaic_label = train_data[0]["mosaic_label"]
fore_idx = train_data[0]["fore_idx"]
test_mosaic_list_of_images = test_data[0]["mosaic_list"]
test_mosaic_label = test_data[0]["mosaic_label"]
test_fore_idx = test_data[0]["fore_idx"]
batch = 3000
train_dataset = MosaicDataset1(mosaic_list_of_images, mosaic_label, fore_idx)
train_loader = DataLoader( train_dataset,batch_size= batch ,shuffle=False)
#batch = 2000
#test_dataset = MosaicDataset1(test_mosaic_list_of_images, test_mosaic_label, test_fore_idx)
#test_loader = DataLoader(test_dataset,batch_size= batch ,shuffle=False)
```
# NTK
```
data = np.load("NTK_1.npy",allow_pickle=True)
# H = data[0]
print(data[0].keys())
H = torch.tensor(data[0]["NTK"])
lr_1 = 1/1470559.2
# p_vec = nn.utils.parameters_to_vector(where_func.parameters())
# p, = p_vec.shape
# n_m, n_obj,_ = inputs.shape # number of mosaic images x number of objects in each mosaic x d
# # this is the transpose jacobian (grad y(w))^T)
# features = torch.zeros(n_m*n_obj, p, requires_grad=False)
# k = 0
# for i in range(27000):
# out = where_func(inpp[i])
# where_func.zero_grad()
# out.backward(retain_graph=False)
# p_grad = torch.tensor([], requires_grad=False)
# for p in where_func.parameters():
# p_grad = torch.cat((p_grad, p.grad.reshape(-1)))
# features[k,:] = p_grad
# k = k+1
# tangent_kernel = features@features.T
# class Module1(nn.Module):
# def __init__(self):
# super(Module1, self).__init__()
# self.linear1 = nn.Linear(2,100)
# self.linear2 = nn.Linear(100,1)
# def forward(self,x):
# x = F.relu(self.linear1(x))
# x = self.linear2(x)
# return x
# from tqdm import tqdm as tqdm
# inputs,_,_ = iter(train_loader).next()
# inputs = torch.reshape(inputs,(27000,2))
# inputs = (inputs - torch.mean(inputs,dim=0,keepdims=True) )/torch.std(inputs,dim=0,keepdims=True)
# where_net = Module1()
# outputs = where_net(inputs)
# feature1 = torch.zeros((27000,200))
# feature2 = torch.zeros((27000,100))
# for i in tqdm(range(27000)):
# where_net.zero_grad()
# outputs[i].backward(retain_graph=True)
# par = []
# j = 0
# for p in where_net.parameters():
# if j%2 == 0:
# vec = torch.nn.utils.parameters_to_vector(p)
# p_grad = p.grad.reshape(-1)
# par.append(p_grad)
# j = j+1
# feature1[i,:] = par[0]
# feature2[i,:] = par[1]
# H = feature1@feature1.T + feature2@feature2.T
```
# Models
```
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
self.linear1 = nn.Linear(2,100)
self.linear2 = nn.Linear(100,3)
def forward(self,x):
x = F.relu(self.linear1(x))
x = self.linear2(x)
return x
print(H)
torch.manual_seed(1234)
what_net = Module2().double()
#what_net.load_state_dict(torch.load("type4_what_net.pt"))
what_net = what_net.to("cuda")
n_batches = 3000//batch
bg = []
for i in range(n_batches):
torch.manual_seed(i)
betag = torch.randn(3000,9)#torch.ones((250,9))/9
bg.append( betag.requires_grad_() )
```
# training
```
criterion = nn.CrossEntropyLoss()
optim1 = []
H= H.to("cpu")
for i in range(n_batches):
optim1.append(MyRmsprop([bg[i]],H=H,lr=1))
# instantiate what net optimizer
optimizer_what = optim.RMSprop(what_net.parameters(), lr=0.0001)#, momentum=0.9)#,nesterov=True)
acti = []
analysis_data_tr = []
analysis_data_tst = []
loss_curi_tr = []
loss_curi_tst = []
epochs = 2500
# calculate zeroth epoch loss and FTPT values
running_loss,anlys_data,correct,total,accuracy = calculate_attn_loss(train_loader,bg,what_net,criterion)
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(0,running_loss,correct,total,accuracy))
loss_curi_tr.append(running_loss)
analysis_data_tr.append(anlys_data)
# training starts
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
what_net.train()
for i, data in enumerate(train_loader, 0):
# get the inputs
inputs, labels,_ = data
inputs = inputs.double()
beta = bg[i] # alpha for ith batch
#print(labels)
inputs, labels,beta = inputs.to("cuda"),labels.to("cuda"),beta.to("cuda")
# zero the parameter gradients
optimizer_what.zero_grad()
optim1[i].zero_grad()
# forward + backward + optimize
avg,alpha = attn_avg(inputs,beta)
outputs = what_net(avg)
loss = criterion(outputs, labels)
# print statistics
running_loss += loss.item()
#alpha.retain_grad()
loss.backward(retain_graph=False)
optimizer_what.step()
optim1[i].step()
running_loss_tr,anls_data,correct,total,accuracy = calculate_attn_loss(train_loader,bg,what_net,criterion)
analysis_data_tr.append(anls_data)
loss_curi_tr.append(running_loss_tr) #loss per epoch
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(epoch+1,running_loss_tr,correct,total,accuracy))
if running_loss_tr<=0.08:
break
print('Finished Training run ')
analysis_data_tr = np.array(analysis_data_tr)
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = np.arange(0,epoch+2)
df_train[columns[1]] = analysis_data_tr[:,-2]/30+
df_train[columns[2]] = analysis_data_tr[:,-1]/30
df_train[columns[3]] = analysis_data_tr[:,0]/30
df_train[columns[4]] = analysis_data_tr[:,1]/30
df_train[columns[5]] = analysis_data_tr[:,2]/30
df_train[columns[6]] = analysis_data_tr[:,3]/30
df_train
%cd /content/
plot_analysis(df_train,columns,[0,500,1000,1500,2000,2500])
aph = []
for i in bg:
aph.append(F.softmax(i,dim=1).detach().numpy())
aph = np.concatenate(aph,axis=0)
# torch.save({
# 'epoch': 500,
# 'model_state_dict': what_net.state_dict(),
# #'optimizer_state_dict': optimizer_what.state_dict(),
# "optimizer_alpha":optim1,
# "FTPT_analysis":analysis_data_tr,
# "alpha":aph
# }, "type4_what_net_500.pt")
aph[0]
avrg = []
avrg_lbls = []
with torch.no_grad():
for i, data1 in enumerate(train_loader):
inputs , labels , fore_idx = data1
inputs = inputs.double()
inputs = inputs.to("cuda")
beta = bg[i]
beta = beta.to("cuda")
avg,alpha = attn_avg(inputs,beta)
avrg.append(avg.detach().cpu().numpy())
avrg_lbls.append(labels.numpy())
avrg= np.concatenate(avrg,axis=0)
avrg_lbls = np.concatenate(avrg_lbls,axis=0)
%cd /content/drive/MyDrive/Neural_Tangent_Kernel/
data = np.load("type_4_data.npy",allow_pickle=True)
%cd /content/
plot_decision_boundary(what_net,[1,8,2,9],data,bg,avrg,avrg_lbls)
plt.plot(loss_curi_tr)
```
```
```
```
```
| github_jupyter |
```
import pickle
import pandas as pd
from collections import OrderedDict
# reading csv file
df=pd.read_csv("Data.csv")
from datetime import datetime
year = lambda x: datetime.strptime(x, "%Y-%m-%d" ).year
df['year'] = df['Date_of_Booking'].map(year)
df.head()
month = lambda x: datetime.strptime(x, "%Y-%m-%d" ).month
df['month'] = df['Date_of_Booking'].map(month)
df.head()
day = lambda x: datetime.strptime(x, "%Y-%m-%d" ).day
df['day'] = df['Date_of_Booking'].map(day)
df.head(25)
day = lambda x: datetime.strptime(x, "%Y-%m-%d" )
df['DateofBooking'] = df['Date_of_Booking'].map(day)
#df['DateofServiceRequested']=df['Date_of_Service_Requested'].map(day)
df.head()
df=df.sort_values(by=['Profile ID', 'Date_of_Booking'],ascending=True)
df.tail(25)
date_carts= ( df.groupby('Profile ID',as_index=False)
.agg({'DateofBooking':(lambda x: list(x))})
.rename(columns={'DateofBooking':'latest_cart'}))
date_carts.head(40)
#df['days_since_last_event'] = df['Date_of_Booking'].diff()
#df.head()
df['diff_minutes'] = df['DateofServiceRequested'] - df['DateofBooking']
df.head()
df['days_since_last_event'] = df.groupby('Profile ID')['DateofBooking'].diff()
df['days_sincelast_event'] = df.groupby('Profile ID')['DateofBooking'].diff().apply(lambda x: x.days)
df.head()
df = df.merge(date_carts, on='Profile ID')
df.head(20)
# get total past orders per user from max order number
df_date = (
df.sort_values(by=['Profile ID', 'Date_of_Booking'],ascending=False)
.groupby('Profile ID')
['Date_of_Booking'].nth(0)
)
df_date.tail(20)
x=df['latest_cart'].iloc[2]
print(x)
#date1=datetime.strptime(x[0], "%Y-%m-%d" )
test=df['latest_cart'].iloc[0]
#type(test)
df['user_total_orders'] = (
df.sort_values(by=['Profile ID', 'Date_of_Booking'],ascending=False)
.groupby('user_id',as_index=False)
['order_number'].nth(0)
)
df_user =df.groupby('Profile ID')
df_user.get_group(14)
df_user.head()
user_df = df.groupby(['Profile ID'],as_index=False).agg(OrderedDict(
[('Transaction_ID','nunique'),
('Date_of_Booking','count')]))
user_df.head(5)
df.head()
df.drop('latest_cart',axis=1,inplace=True)
df.drop('DateofBooking_x',axis=1,inplace=True)
df.drop('DateofBooking_y',axis=1,inplace=True)
df.drop('dateofbooking',axis=1,inplace=True)
df.drop('days_since_prior_order',axis=1,inplace=True)
df.drop('apr',axis=1,inplace=True)
df[df['Profile ID']==14]
values = {'days_sincelast_event': 0}
df.fillna(value=values, inplace=True)
df.drop('labels',axis=1,inplace=True)
df.head()
#df['days_sincelast_event'] = df.groupby('Profile ID')['DateofBooking'].diff().apply(lambda x: x.days)
#df['labels']=df.groupby('Profile ID')['days_sincelast_event'].diff()
df['labels']=df.groupby('Profile ID')['days_sincelast_event'].shift(periods=-1, fill_value=0)
df.head(24)
def func(x):
if(x<31 and x>0):
return 1
else:
return 0
df['thirty']=df['labels'].map(func)
df.head(24)
def func(x):
if(x<91 and x>0):
return 1
else:
return 0
df['ninty']=df['labels'].map(func)
df.head(24)
df.tail()
df.to_pickle('features.pickle')
!ls
```
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex AI client library: AutoML text entity extraction model for batch prediction
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/community/gapic/automl/showcase_automl_text_entity_extraction_batch.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/community/gapic/automl/showcase_automl_text_entity_extraction_batch.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex AI Python client library to create text entity extraction models and do batch prediction using Google Cloud's [AutoML](https://cloud.google.com/ai-platform-unified/docs/start/automl-users).
### Dataset
The dataset used for this tutorial is the [NCBI Disease Research Abstracts dataset](https://www.ncbi.nlm.nih.gov/CBBresearch/Dogan/DISEASE/) from [National Center for Biotechnology Information](https://www.ncbi.nlm.nih.gov/). The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket.
### Objective
In this tutorial, you create an AutoML text entity extraction model from a Python script, and then do a batch prediction using the Vertex AI client library. You can alternatively create and deploy models using the `gcloud` command-line tool or online using the Google Cloud Console.
The steps performed include:
- Create a Vertex AI `Dataset` resource.
- Train the model.
- View the model evaluation.
- Make a batch prediction.
There is one key difference between using batch prediction and using online prediction:
* Prediction Service: Does an on-demand prediction for the entire set of instances (i.e., one or more data items) and returns the results in real-time.
* Batch Prediction Service: Does a queued (batch) prediction for the entire set of instances in the background and stores the results in a Cloud Storage bucket when ready.
### Costs
This tutorial uses billable components of Google Cloud (GCP):
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/ai-platform-unified/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Installation
Install the latest version of Vertex AI client library.
```
import sys
if "google.colab" in sys.modules:
USER_FLAG = ""
else:
USER_FLAG = "--user"
! pip3 install -U google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
### Restart the kernel
Once you've installed the Vertex AI client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex AI APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Vertex AI Notebooks.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex AI. Not all regions provide support for all Vertex AI services. For the latest support per region, see the [Vertex AI locations documentation](https://cloud.google.com/ai-platform-unified/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Vertex AI Notebooks**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex AI" into the filter box, and select **Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on AI Platform, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
This tutorial is designed to use training data that is in a public Cloud Storage bucket and a local Cloud Storage bucket for your batch predictions. You may alternatively use your own training data that you have stored in a local Cloud Storage bucket.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
#### Import Vertex AI client library
Import the Vertex AI client library into our Python environment.
```
import os
import sys
import time
import google.cloud.aiplatform_v1 as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
```
#### Vertex AI constants
Setup up the following constants for Vertex AI:
- `API_ENDPOINT`: The Vertex AI API service endpoint for dataset, model, job, pipeline and endpoint services.
- `PARENT`: The Vertex AI location root path for dataset, model, job, pipeline and endpoint resources.
```
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex AI location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
```
#### AutoML constants
Set constants unique to AutoML datasets and training:
- Dataset Schemas: Tells the `Dataset` resource service which type of dataset it is.
- Data Labeling (Annotations) Schemas: Tells the `Dataset` resource service how the data is labeled (annotated).
- Dataset Training Schemas: Tells the `Pipeline` resource service the task (e.g., classification) to train the model for.
```
# Text Dataset type
DATA_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/metadata/text_1.0.0.yaml"
# Text Labeling type
LABEL_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/ioformat/text_extraction_io_format_1.0.0.yaml"
# Text Training task
TRAINING_SCHEMA = "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_text_extraction_1.0.0.yaml"
```
#### Hardware Accelerators
Set the hardware accelerators (e.g., GPU), if any, for prediction.
Set the variable `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Telsa K80 GPUs allocated to each VM, you would specify:
(aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
For GPU, available accelerators include:
- aip.AcceleratorType.NVIDIA_TESLA_K80
- aip.AcceleratorType.NVIDIA_TESLA_P100
- aip.AcceleratorType.NVIDIA_TESLA_P4
- aip.AcceleratorType.NVIDIA_TESLA_T4
- aip.AcceleratorType.NVIDIA_TESLA_V100
Otherwise specify `(None, None)` to use a container image to run on a CPU.
```
if os.getenv("IS_TESTING_DEPOLY_GPU"):
DEPLOY_GPU, DEPLOY_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_DEPOLY_GPU")),
)
else:
DEPLOY_GPU, DEPLOY_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
```
#### Container (Docker) image
For AutoML batch prediction, the container image for the serving binary is pre-determined by the Vertex AI prediction service. More specifically, the service will pick the appropriate container for the model depending on the hardware accelerator you selected.
#### Machine Type
Next, set the machine type to use for prediction.
- Set the variable `DEPLOY_COMPUTE` to configure the compute resources for the VM you will use for prediction.
- `machine type`
- `n1-standard`: 3.75GB of memory per vCPU.
- `n1-highmem`: 6.5GB of memory per vCPU
- `n1-highcpu`: 0.9 GB of memory per vCPU
- `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
*Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*
```
if os.getenv("IS_TESTING_DEPLOY_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_DEPLOY_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
```
# Tutorial
Now you are ready to start creating your own AutoML text entity extraction model.
## Set up clients
The Vertex AI client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex AI server.
You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
- Dataset Service for `Dataset` resources.
- Model Service for `Model` resources.
- Pipeline Service for training.
- Job Service for batch prediction and custom training.
```
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_dataset_client():
client = aip.DatasetServiceClient(client_options=client_options)
return client
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_pipeline_client():
client = aip.PipelineServiceClient(client_options=client_options)
return client
def create_job_client():
client = aip.JobServiceClient(client_options=client_options)
return client
clients = {}
clients["dataset"] = create_dataset_client()
clients["model"] = create_model_client()
clients["pipeline"] = create_pipeline_client()
clients["job"] = create_job_client()
for client in clients.items():
print(client)
```
## Dataset
Now that your clients are ready, your first step in training a model is to create a managed dataset instance, and then upload your labeled data to it.
### Create `Dataset` resource instance
Use the helper function `create_dataset` to create the instance of a `Dataset` resource. This function does the following:
1. Uses the dataset client service.
2. Creates an Vertex AI `Dataset` resource (`aip.Dataset`), with the following parameters:
- `display_name`: The human-readable name you choose to give it.
- `metadata_schema_uri`: The schema for the dataset type.
3. Calls the client dataset service method `create_dataset`, with the following parameters:
- `parent`: The Vertex AI location root path for your `Database`, `Model` and `Endpoint` resources.
- `dataset`: The Vertex AI dataset object instance you created.
4. The method returns an `operation` object.
An `operation` object is how Vertex AI handles asynchronous calls for long running operations. While this step usually goes fast, when you first use it in your project, there is a longer delay due to provisioning.
You can use the `operation` object to get status on the operation (e.g., create `Dataset` resource) or to cancel the operation, by invoking an operation method:
| Method | Description |
| ----------- | ----------- |
| result() | Waits for the operation to complete and returns a result object in JSON format. |
| running() | Returns True/False on whether the operation is still running. |
| done() | Returns True/False on whether the operation is completed. |
| canceled() | Returns True/False on whether the operation was canceled. |
| cancel() | Cancels the operation (this may take up to 30 seconds). |
```
TIMEOUT = 90
def create_dataset(name, schema, labels=None, timeout=TIMEOUT):
start_time = time.time()
try:
dataset = aip.Dataset(
display_name=name, metadata_schema_uri=schema, labels=labels
)
operation = clients["dataset"].create_dataset(parent=PARENT, dataset=dataset)
print("Long running operation:", operation.operation.name)
result = operation.result(timeout=TIMEOUT)
print("time:", time.time() - start_time)
print("response")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" metadata_schema_uri:", result.metadata_schema_uri)
print(" metadata:", dict(result.metadata))
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
print(" etag:", result.etag)
print(" labels:", dict(result.labels))
return result
except Exception as e:
print("exception:", e)
return None
result = create_dataset("biomedical-" + TIMESTAMP, DATA_SCHEMA)
```
Now save the unique dataset identifier for the `Dataset` resource instance you created.
```
# The full unique ID for the dataset
dataset_id = result.name
# The short numeric ID for the dataset
dataset_short_id = dataset_id.split("/")[-1]
print(dataset_id)
```
### Data preparation
The Vertex AI `Dataset` resource for text has a couple of requirements for your text entity extraction data.
- Text examples must be stored in a JSONL file. Unlike text classification and sentiment analysis, a CSV index file is not supported.
- The examples must be either inline text or reference text files that are in Cloud Storage buckets.
#### JSONL
For text entity extraction, the JSONL file has a few requirements:
- Each data item is a separate JSON object, on a separate line.
- The key/value pair `text_segment_annotations` is a list of character start/end positions in the text per entity with the corresponding label.
- `display_name`: The label.
- `start_offset/end_offset`: The character offsets of the start/end of the entity.
- The key/value pair `text_content` is the text.
{'text_segment_annotations': [{'end_offset': value, 'start_offset': value, 'display_name': label}, ...], 'text_content': text}
*Note*: The dictionary key fields may alternatively be in camelCase. For example, 'display_name' can also be 'displayName'.
#### Location of Cloud Storage training data.
Now set the variable `IMPORT_FILE` to the location of the JSONL index file in Cloud Storage.
```
IMPORT_FILE = "gs://ucaip-test-us-central1/dataset/ucaip_ten_dataset.jsonl"
```
#### Quick peek at your data
You will use a version of the NCBI Biomedical dataset that is stored in a public Cloud Storage bucket, using a JSONL index file.
Start by doing a quick peek at the data. You count the number of examples by counting the number of objects in a JSONL index file (`wc -l`) and then peek at the first few rows.
```
if "IMPORT_FILES" in globals():
FILE = IMPORT_FILES[0]
else:
FILE = IMPORT_FILE
count = ! gsutil cat $FILE | wc -l
print("Number of Examples", int(count[0]))
print("First 10 rows")
! gsutil cat $FILE | head
```
### Import data
Now, import the data into your Vertex AI Dataset resource. Use this helper function `import_data` to import the data. The function does the following:
- Uses the `Dataset` client.
- Calls the client method `import_data`, with the following parameters:
- `name`: The human readable name you give to the `Dataset` resource (e.g., biomedical).
- `import_configs`: The import configuration.
- `import_configs`: A Python list containing a dictionary, with the key/value entries:
- `gcs_sources`: A list of URIs to the paths of the one or more index files.
- `import_schema_uri`: The schema identifying the labeling type.
The `import_data()` method returns a long running `operation` object. This will take a few minutes to complete. If you are in a live tutorial, this would be a good time to ask questions, or take a personal break.
```
def import_data(dataset, gcs_sources, schema):
config = [{"gcs_source": {"uris": gcs_sources}, "import_schema_uri": schema}]
print("dataset:", dataset_id)
start_time = time.time()
try:
operation = clients["dataset"].import_data(
name=dataset_id, import_configs=config
)
print("Long running operation:", operation.operation.name)
result = operation.result()
print("result:", result)
print("time:", int(time.time() - start_time), "secs")
print("error:", operation.exception())
print("meta :", operation.metadata)
print(
"after: running:",
operation.running(),
"done:",
operation.done(),
"cancelled:",
operation.cancelled(),
)
return operation
except Exception as e:
print("exception:", e)
return None
import_data(dataset_id, [IMPORT_FILE], LABEL_SCHEMA)
```
## Train the model
Now train an AutoML text entity extraction model using your Vertex AI `Dataset` resource. To train the model, do the following steps:
1. Create an Vertex AI training pipeline for the `Dataset` resource.
2. Execute the pipeline to start the training.
### Create a training pipeline
You may ask, what do we use a pipeline for? You typically use pipelines when the job (such as training) has multiple steps, generally in sequential order: do step A, do step B, etc. By putting the steps into a pipeline, we gain the benefits of:
1. Being reusable for subsequent training jobs.
2. Can be containerized and ran as a batch job.
3. Can be distributed.
4. All the steps are associated with the same pipeline job for tracking progress.
Use this helper function `create_pipeline`, which takes the following parameters:
- `pipeline_name`: A human readable name for the pipeline job.
- `model_name`: A human readable name for the model.
- `dataset`: The Vertex AI fully qualified dataset identifier.
- `schema`: The dataset labeling (annotation) training schema.
- `task`: A dictionary describing the requirements for the training job.
The helper function calls the `Pipeline` client service'smethod `create_pipeline`, which takes the following parameters:
- `parent`: The Vertex AI location root path for your `Dataset`, `Model` and `Endpoint` resources.
- `training_pipeline`: the full specification for the pipeline training job.
Let's look now deeper into the *minimal* requirements for constructing a `training_pipeline` specification:
- `display_name`: A human readable name for the pipeline job.
- `training_task_definition`: The dataset labeling (annotation) training schema.
- `training_task_inputs`: A dictionary describing the requirements for the training job.
- `model_to_upload`: A human readable name for the model.
- `input_data_config`: The dataset specification.
- `dataset_id`: The Vertex AI dataset identifier only (non-fully qualified) -- this is the last part of the fully-qualified identifier.
- `fraction_split`: If specified, the percentages of the dataset to use for training, test and validation. Otherwise, the percentages are automatically selected by AutoML.
```
def create_pipeline(pipeline_name, model_name, dataset, schema, task):
dataset_id = dataset.split("/")[-1]
input_config = {
"dataset_id": dataset_id,
"fraction_split": {
"training_fraction": 0.8,
"validation_fraction": 0.1,
"test_fraction": 0.1,
},
}
training_pipeline = {
"display_name": pipeline_name,
"training_task_definition": schema,
"training_task_inputs": task,
"input_data_config": input_config,
"model_to_upload": {"display_name": model_name},
}
try:
pipeline = clients["pipeline"].create_training_pipeline(
parent=PARENT, training_pipeline=training_pipeline
)
print(pipeline)
except Exception as e:
print("exception:", e)
return None
return pipeline
```
### Construct the task requirements
Next, construct the task requirements. Unlike other parameters which take a Python (JSON-like) dictionary, the `task` field takes a Google protobuf Struct, which is very similar to a Python dictionary. Use the `json_format.ParseDict` method for the conversion.
The minimal fields you need to specify are:
- `multi_label`: Whether True/False this is a multi-label (vs single) classification.
- `budget_milli_node_hours`: The maximum time to budget (billed) for training the model, where 1000 = 1 hour.
- `model_type`: The type of deployed model:
- `CLOUD`: For deploying to Google Cloud.
- `disable_early_stopping`: Whether True/False to let AutoML use its judgement to stop training early or train for the entire budget.
Finally, you create the pipeline by calling the helper function `create_pipeline`, which returns an instance of a training pipeline object.
```
PIPE_NAME = "biomedical_pipe-" + TIMESTAMP
MODEL_NAME = "biomedical_model-" + TIMESTAMP
task = json_format.ParseDict(
{
"multi_label": False,
"budget_milli_node_hours": 8000,
"model_type": "CLOUD",
"disable_early_stopping": False,
},
Value(),
)
response = create_pipeline(PIPE_NAME, MODEL_NAME, dataset_id, TRAINING_SCHEMA, task)
```
Now save the unique identifier of the training pipeline you created.
```
# The full unique ID for the pipeline
pipeline_id = response.name
# The short numeric ID for the pipeline
pipeline_short_id = pipeline_id.split("/")[-1]
print(pipeline_id)
```
### Get information on a training pipeline
Now get pipeline information for just this training pipeline instance. The helper function gets the job information for just this job by calling the the job client service's `get_training_pipeline` method, with the following parameter:
- `name`: The Vertex AI fully qualified pipeline identifier.
When the model is done training, the pipeline state will be `PIPELINE_STATE_SUCCEEDED`.
```
def get_training_pipeline(name, silent=False):
response = clients["pipeline"].get_training_pipeline(name=name)
if silent:
return response
print("pipeline")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" state:", response.state)
print(" training_task_definition:", response.training_task_definition)
print(" training_task_inputs:", dict(response.training_task_inputs))
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", dict(response.labels))
return response
response = get_training_pipeline(pipeline_id)
```
# Deployment
Training the above model may take upwards of 120 minutes time.
Once your model is done training, you can calculate the actual time it took to train the model by subtracting `end_time` from `start_time`. For your model, you will need to know the fully qualified Vertex AI Model resource identifier, which the pipeline service assigned to it. You can get this from the returned pipeline instance as the field `model_to_deploy.name`.
```
while True:
response = get_training_pipeline(pipeline_id, True)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
model_to_deploy_id = None
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
raise Exception("Training Job Failed")
else:
model_to_deploy = response.model_to_upload
model_to_deploy_id = model_to_deploy.name
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
print("model to deploy:", model_to_deploy_id)
```
## Model information
Now that your model is trained, you can get some information on your model.
## Evaluate the Model resource
Now find out how good the model service believes your model is. As part of training, some portion of the dataset was set aside as the test (holdout) data, which is used by the pipeline service to evaluate the model.
### List evaluations for all slices
Use this helper function `list_model_evaluations`, which takes the following parameter:
- `name`: The Vertex AI fully qualified model identifier for the `Model` resource.
This helper function uses the model client service's `list_model_evaluations` method, which takes the same parameter. The response object from the call is a list, where each element is an evaluation metric.
For each evaluation -- you probably only have one, we then print all the key names for each metric in the evaluation, and for a small set (`confusionMatrix` and `confidenceMetrics`) you will print the result.
```
def list_model_evaluations(name):
response = clients["model"].list_model_evaluations(parent=name)
for evaluation in response:
print("model_evaluation")
print(" name:", evaluation.name)
print(" metrics_schema_uri:", evaluation.metrics_schema_uri)
metrics = json_format.MessageToDict(evaluation._pb.metrics)
for metric in metrics.keys():
print(metric)
print("confusionMatrix", metrics["confusionMatrix"])
print("confidenceMetrics", metrics["confidenceMetrics"])
return evaluation.name
last_evaluation = list_model_evaluations(model_to_deploy_id)
```
## Model deployment for batch prediction
Now deploy the trained Vertex AI `Model` resource you created for batch prediction. This differs from deploying a `Model` resource for on-demand prediction.
For online prediction, you:
1. Create an `Endpoint` resource for deploying the `Model` resource to.
2. Deploy the `Model` resource to the `Endpoint` resource.
3. Make online prediction requests to the `Endpoint` resource.
For batch-prediction, you:
1. Create a batch prediction job.
2. The job service will provision resources for the batch prediction request.
3. The results of the batch prediction request are returned to the caller.
4. The job service will unprovision the resoures for the batch prediction request.
## Make a batch prediction request
Now do a batch prediction to your deployed model.
### Make test items
You will use synthetic data as a test data items. Don't be concerned that we are using synthetic data -- we just want to demonstrate how to make a prediction.
```
test_item_1 = 'Molecular basis of hexosaminidase A deficiency and pseudodeficiency in the Berks County Pennsylvania Dutch.\tFollowing the birth of two infants with Tay-Sachs disease ( TSD ) , a non-Jewish , Pennsylvania Dutch kindred was screened for TSD carriers using the biochemical assay . A high frequency of individuals who appeared to be TSD heterozygotes was detected ( Kelly et al . , 1975 ) . Clinical and biochemical evidence suggested that the increased carrier frequency was due to at least two altered alleles for the hexosaminidase A alpha-subunit . We now report two mutant alleles in this Pennsylvania Dutch kindred , and one polymorphism . One allele , reported originally in a French TSD patient ( Akli et al . , 1991 ) , is a GT-- > AT transition at the donor splice-site of intron 9 . The second , a C-- > T transition at nucleotide 739 ( Arg247Trp ) , has been shown by Triggs-Raine et al . ( 1992 ) to be a clinically benign " pseudodeficient " allele associated with reduced enzyme activity against artificial substrate . Finally , a polymorphism [ G-- > A ( 759 ) ] , which leaves valine at codon 253 unchanged , is described'
test_item_2 = "Analysis of alkaptonuria (AKU) mutations and polymorphisms reveals that the CCC sequence motif is a mutational hot spot in the homogentisate 1,2 dioxygenase gene (HGO). We recently showed that alkaptonuria ( AKU ) is caused by loss-of-function mutations in the homogentisate 1 , 2 dioxygenase gene ( HGO ) . Herein we describe haplotype and mutational analyses of HGO in seven new AKU pedigrees . These analyses identified two novel single-nucleotide polymorphisms ( INV4 + 31A-- > G and INV11 + 18A-- > G ) and six novel AKU mutations ( INV1-1G-- > A , W60G , Y62C , A122D , P230T , and D291E ) , which further illustrates the remarkable allelic heterogeneity found in AKU . Reexamination of all 29 mutations and polymorphisms thus far described in HGO shows that these nucleotide changes are not randomly distributed ; the CCC sequence motif and its inverted complement , GGG , are preferentially mutated . These analyses also demonstrated that the nucleotide substitutions in HGO do not involve CpG dinucleotides , which illustrates important differences between HGO and other genes for the occurrence of mutation at specific short-sequence motifs . Because the CCC sequence motifs comprise a significant proportion ( 34 . 5 % ) of all mutated bases that have been observed in HGO , we conclude that the CCC triplet is a mutational hot spot in HGO ."
```
### Make the batch input file
Now make a batch input file, which you will store in your local Cloud Storage bucket. The batch input file can only be in JSONL format. For JSONL file, you make one dictionary entry per line for each data item (instance). The dictionary contains the key/value pairs:
- `content`: The Cloud Storage path to the file with the text item.
- `mime_type`: The content type. In our example, it is an `text` file.
For example:
{'content': '[your-bucket]/file1.txt', 'mime_type': 'text'}
```
import json
import tensorflow as tf
gcs_test_item_1 = BUCKET_NAME + "/test1.txt"
with tf.io.gfile.GFile(gcs_test_item_1, "w") as f:
f.write(test_item_1 + "\n")
gcs_test_item_2 = BUCKET_NAME + "/test2.txt"
with tf.io.gfile.GFile(gcs_test_item_2, "w") as f:
f.write(test_item_2 + "\n")
gcs_input_uri = BUCKET_NAME + "/test.jsonl"
with tf.io.gfile.GFile(gcs_input_uri, "w") as f:
data = {"content": gcs_test_item_1, "mime_type": "text/plain"}
f.write(json.dumps(data) + "\n")
data = {"content": gcs_test_item_2, "mime_type": "text/plain"}
f.write(json.dumps(data) + "\n")
print(gcs_input_uri)
! gsutil cat $gcs_input_uri
```
### Compute instance scaling
You have several choices on scaling the compute instances for handling your batch prediction requests:
- Single Instance: The batch prediction requests are processed on a single compute instance.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to one.
- Manual Scaling: The batch prediction requests are split across a fixed number of compute instances that you manually specified.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to the same number of nodes. When a model is first deployed to the instance, the fixed number of compute instances are provisioned and batch prediction requests are evenly distributed across them.
- Auto Scaling: The batch prediction requests are split across a scaleable number of compute instances.
- Set the minimum (`MIN_NODES`) number of compute instances to provision when a model is first deployed and to de-provision, and set the maximum (`MAX_NODES) number of compute instances to provision, depending on load conditions.
The minimum number of compute instances corresponds to the field `min_replica_count` and the maximum number of compute instances corresponds to the field `max_replica_count`, in your subsequent deployment request.
```
MIN_NODES = 1
MAX_NODES = 1
```
### Make batch prediction request
Now that your batch of two test items is ready, let's do the batch request. Use this helper function `create_batch_prediction_job`, with the following parameters:
- `display_name`: The human readable name for the prediction job.
- `model_name`: The Vertex AI fully qualified identifier for the `Model` resource.
- `gcs_source_uri`: The Cloud Storage path to the input file -- which you created above.
- `gcs_destination_output_uri_prefix`: The Cloud Storage path that the service will write the predictions to.
- `parameters`: Additional filtering parameters for serving prediction results.
The helper function calls the job client service's `create_batch_prediction_job` metho, with the following parameters:
- `parent`: The Vertex AI location root path for Dataset, Model and Pipeline resources.
- `batch_prediction_job`: The specification for the batch prediction job.
Let's now dive into the specification for the `batch_prediction_job`:
- `display_name`: The human readable name for the prediction batch job.
- `model`: The Vertex AI fully qualified identifier for the `Model` resource.
- `dedicated_resources`: The compute resources to provision for the batch prediction job.
- `machine_spec`: The compute instance to provision. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
- `starting_replica_count`: The number of compute instances to initially provision, which you set earlier as the variable `MIN_NODES`.
- `max_replica_count`: The maximum number of compute instances to scale to, which you set earlier as the variable `MAX_NODES`.
- `model_parameters`: Additional filtering parameters for serving prediction results. *Note*, text models do not support additional parameters.
- `input_config`: The input source and format type for the instances to predict.
- `instances_format`: The format of the batch prediction request file: `jsonl` only supported.
- `gcs_source`: A list of one or more Cloud Storage paths to your batch prediction requests.
- `output_config`: The output destination and format for the predictions.
- `prediction_format`: The format of the batch prediction response file: `jsonl` only supported.
- `gcs_destination`: The output destination for the predictions.
- `dedicated_resources`: The compute resources to provision for the batch prediction job.
- `machine_spec`: The compute instance to provision. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
- `starting_replica_count`: The number of compute instances to initially provision.
- `max_replica_count`: The maximum number of compute instances to scale to. In this tutorial, only one instance is provisioned.
This call is an asychronous operation. You will print from the response object a few select fields, including:
- `name`: The Vertex AI fully qualified identifier assigned to the batch prediction job.
- `display_name`: The human readable name for the prediction batch job.
- `model`: The Vertex AI fully qualified identifier for the Model resource.
- `generate_explanations`: Whether True/False explanations were provided with the predictions (explainability).
- `state`: The state of the prediction job (pending, running, etc).
Since this call will take a few moments to execute, you will likely get `JobState.JOB_STATE_PENDING` for `state`.
```
BATCH_MODEL = "biomedical_batch-" + TIMESTAMP
def create_batch_prediction_job(
display_name,
model_name,
gcs_source_uri,
gcs_destination_output_uri_prefix,
parameters=None,
):
if DEPLOY_GPU:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_type": DEPLOY_GPU,
"accelerator_count": DEPLOY_NGPU,
}
else:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_count": 0,
}
batch_prediction_job = {
"display_name": display_name,
# Format: 'projects/{project}/locations/{location}/models/{model_id}'
"model": model_name,
"model_parameters": json_format.ParseDict(parameters, Value()),
"input_config": {
"instances_format": IN_FORMAT,
"gcs_source": {"uris": [gcs_source_uri]},
},
"output_config": {
"predictions_format": OUT_FORMAT,
"gcs_destination": {"output_uri_prefix": gcs_destination_output_uri_prefix},
},
"dedicated_resources": {
"machine_spec": machine_spec,
"starting_replica_count": MIN_NODES,
"max_replica_count": MAX_NODES,
},
}
response = clients["job"].create_batch_prediction_job(
parent=PARENT, batch_prediction_job=batch_prediction_job
)
print("response")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" model:", response.model)
try:
print(" generate_explanation:", response.generate_explanation)
except:
pass
print(" state:", response.state)
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", response.labels)
return response
IN_FORMAT = "jsonl"
OUT_FORMAT = "jsonl" # [jsonl]
response = create_batch_prediction_job(
BATCH_MODEL, model_to_deploy_id, gcs_input_uri, BUCKET_NAME, None
)
```
Now get the unique identifier for the batch prediction job you created.
```
# The full unique ID for the batch job
batch_job_id = response.name
# The short numeric ID for the batch job
batch_job_short_id = batch_job_id.split("/")[-1]
print(batch_job_id)
```
### Get information on a batch prediction job
Use this helper function `get_batch_prediction_job`, with the following paramter:
- `job_name`: The Vertex AI fully qualified identifier for the batch prediction job.
The helper function calls the job client service's `get_batch_prediction_job` method, with the following paramter:
- `name`: The Vertex AI fully qualified identifier for the batch prediction job. In this tutorial, you will pass it the Vertex AI fully qualified identifier for your batch prediction job -- `batch_job_id`
The helper function will return the Cloud Storage path to where the predictions are stored -- `gcs_destination`.
```
def get_batch_prediction_job(job_name, silent=False):
response = clients["job"].get_batch_prediction_job(name=job_name)
if silent:
return response.output_config.gcs_destination.output_uri_prefix, response.state
print("response")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" model:", response.model)
try: # not all data types support explanations
print(" generate_explanation:", response.generate_explanation)
except:
pass
print(" state:", response.state)
print(" error:", response.error)
gcs_destination = response.output_config.gcs_destination
print(" gcs_destination")
print(" output_uri_prefix:", gcs_destination.output_uri_prefix)
return gcs_destination.output_uri_prefix, response.state
predictions, state = get_batch_prediction_job(batch_job_id)
```
### Get the predictions
When the batch prediction is done processing, the job state will be `JOB_STATE_SUCCEEDED`.
Finally you view the predictions stored at the Cloud Storage path you set as output. The predictions will be in a JSONL format, which you indicated at the time you made the batch prediction job, under a subfolder starting with the name `prediction`, and under that folder will be a file called `predictions*.jsonl`.
Now display (cat) the contents. You will see multiple JSON objects, one for each prediction.
The first field `text_snippet` is the text file you did the prediction on, and the second field `annotations` is the prediction, which is further broken down into:
- `text_extraction`: The extracted entity from the text.
- `display_name`: The predicted label for the extraction entity.
- `score`: The confidence level between 0 and 1 in the prediction.
- `startOffset`: The character offset in the text of the start of the extracted entity.
- `endOffset`: The character offset in the text of the end of the extracted entity.
```
def get_latest_predictions(gcs_out_dir):
""" Get the latest prediction subfolder using the timestamp in the subfolder name"""
folders = !gsutil ls $gcs_out_dir
latest = ""
for folder in folders:
subfolder = folder.split("/")[-2]
if subfolder.startswith("prediction-"):
if subfolder > latest:
latest = folder[:-1]
return latest
while True:
predictions, state = get_batch_prediction_job(batch_job_id, True)
if state != aip.JobState.JOB_STATE_SUCCEEDED:
print("The job has not completed:", state)
if state == aip.JobState.JOB_STATE_FAILED:
raise Exception("Batch Job Failed")
else:
folder = get_latest_predictions(predictions)
! gsutil ls $folder/prediction*.jsonl
! gsutil cat $folder/prediction*.jsonl
break
time.sleep(60)
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex AI fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex AI fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex AI fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex AI fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex AI fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex AI fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex AI fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from sklearn import preprocessing
import matplotlib.pyplot as plt
plt.rc("font", size=14)
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
import seaborn as sns
sns.set(style="white")
sns.set(style="whitegrid", color_codes=True)
```
## Data
The data is related with direct marketing campaigns (phone calls) of a Portuguese banking institution. The classification goal is to predict if the client will subscribe (1/0) a term deposit (variable y).
This dataset provides the customer information. It includes 41188 records and 21 fields.
```
data = pd.read_csv('data/bank.csv', header=0, delimiter=';')
data = data.dropna()
print(data.shape)
print(list(data.columns))
data.head()
```
#### Input variables
1 - age (numeric)
2 - job : type of job (categorical: 'admin.','blue-collar','entrepreneur','housemaid','management','retired','self-employed','services','student','technician','unemployed','unknown')
3 - marital : marital status (categorical: 'divorced','married','single','unknown'; note: 'divorced' means divorced or widowed)
4 - education (categorical: 'basic.4y','basic.6y','basic.9y','high.school','illiterate','professional.course','university.degree','unknown')
5 - default: has credit in default? (categorical: 'no','yes','unknown')
6 - housing: has housing loan? (categorical: 'no','yes','unknown')
7 - loan: has personal loan? (categorical: 'no','yes','unknown')
8 - contact: contact communication type (categorical: 'cellular','telephone')
9 - month: last contact month of year (categorical: 'jan', 'feb', 'mar', ..., 'nov', 'dec')
10 - day_of_week: last contact day of the week (categorical: 'mon','tue','wed','thu','fri')
11 - duration: last contact duration, in seconds (numeric). Important note: this attribute highly affects the output target (e.g., if duration=0 then y='no'). Yet, the duration is not known before a call is performed. Also, after the end of the call y is obviously known. Thus, this input should only be included for benchmark purposes and should be discarded if the intention is to have a realistic predictive model.
12 - campaign: number of contacts performed during this campaign and for this client (numeric, includes last contact)
13 - pdays: number of days that passed by after the client was last contacted from a previous campaign (numeric; 999 means client was not previously contacted)
14 - previous: number of contacts performed before this campaign and for this client (numeric)
15 - poutcome: outcome of the previous marketing campaign (categorical: 'failure','nonexistent','success')
16 - emp.var.rate: employment variation rate - (numeric)
17 - cons.price.idx: consumer price index - (numeric)
18 - cons.conf.idx: consumer confidence index - (numeric)
19 - euribor3m: euribor 3 month rate - (numeric)
20 - nr.employed: number of employees - (numeric)
#### Predict variable (desired target):
y - has the client subscribed a term deposit? (binary: '1','0')
The education column of the dataset has many categories and we need to reduce the categories for a better modelling. The education column has the following categories:
```
data['education'].unique()
```
Let us group "basic.4y", "basic.9y" and "basic.6y" together and call them "basic".
```
data['education']=np.where(data['education'] =='basic.9y', 'Basic', data['education'])
data['education']=np.where(data['education'] =='basic.6y', 'Basic', data['education'])
data['education']=np.where(data['education'] =='basic.4y', 'Basic', data['education'])
```
After grouping, this is the columns
```
data['education'].unique()
```
### Data exploration
```
data['y'].value_counts()
sns.countplot(x='y',data=data, palette='hls')
plt.show()
plt.savefig('count_plot')
```
There are 36548 no's and 4640 yes's in the outcome variables.
Let's get a sense of the numbers across the two classes
```
data.groupby('y').mean()
```
Observations:
The average age of customers who bought the term deposit is higher than that of the customers who didn't.
The pdays (days since the customer was last contacted) is understandably lower for the customers who bought it. The lower the pdays, the better the memory of the last call and hence the better chances of a sale.
Surprisingly, campaigns (number of contacts or calls made during the current campaign) are lower for customers who bought the term deposit.
We can calculate categorical means for other categorical variables such as education and marital status to get a more detailed sense of our data.
```
data.groupby('job').mean()
data.groupby('marital').mean()
data.groupby('education').mean()
```
Visualizations
```
%matplotlib inline
pd.crosstab(data.job,data.y).plot(kind='bar')
plt.title('Purchase Frequency for Job Title')
plt.xlabel('Job')
plt.ylabel('Frequency of Purchase')
plt.savefig('purchase_fre_job')
```
The frequency of purchase of the deposit depends a great deal on the job title. Thus, the job title can be a good predictor of the outcome variable.
```
table=pd.crosstab(data.marital,data.y)
table.div(table.sum(1).astype(float), axis=0).plot(kind='bar', stacked=True)
plt.title('Stacked Bar Chart of Marital Status vs Purchase')
plt.xlabel('Marital Status')
plt.ylabel('Proportion of Customers')
plt.savefig('mariral_vs_pur_stack')
```
Hard to see, but the marital status does not seem a strong predictor for the outcome variable.
```
table=pd.crosstab(data.education,data.y)
table.div(table.sum(1).astype(float), axis=0).plot(kind='bar', stacked=True)
plt.title('Stacked Bar Chart of Education vs Purchase')
plt.xlabel('Education')
plt.ylabel('Proportion of Customers')
plt.savefig('edu_vs_pur_stack')
```
Education seems a good predictor of the outcome variable.
```
pd.crosstab(data.day_of_week,data.y).plot(kind='bar')
plt.title('Purchase Frequency for Day of Week')
plt.xlabel('Day of Week')
plt.ylabel('Frequency of Purchase')
plt.savefig('pur_dayofweek_bar')
```
Day of week may not be a good predictor of the outcome
```
pd.crosstab(data.month,data.y).plot(kind='bar')
plt.title('Purchase Frequency for Month')
plt.xlabel('Month')
plt.ylabel('Frequency of Purchase')
plt.savefig('pur_fre_month_bar')
```
Month might be a good predictor of the outcome variable
```
data.age.hist()
plt.title('Histogram of Age')
plt.xlabel('Age')
plt.ylabel('Frequency')
plt.savefig('hist_age')
```
The most of the customers of the bank in this dataset are in the age range of 30-40.
```
pd.crosstab(data.poutcome,data.y).plot(kind='bar')
plt.title('Purchase Frequency for Poutcome')
plt.xlabel('Poutcome')
plt.ylabel('Frequency of Purchase')
plt.savefig('pur_fre_pout_bar')
```
Poutcome seems to be a good predictor of the outcome variable.
### Create dummy variables
```
cat_vars=['job','marital','education','default','housing','loan','contact','month','day_of_week','poutcome']
for var in cat_vars:
cat_list='var'+'_'+var
cat_list = pd.get_dummies(data[var], prefix=var)
data1=data.join(cat_list)
data=data1
cat_vars=['job','marital','education','default','housing','loan','contact','month','day_of_week','poutcome']
data_vars=data.columns.values.tolist()
to_keep=[i for i in data_vars if i not in cat_vars]
data_final=data[to_keep]
data_final.columns.values
data_final_vars=data_final.columns.values.tolist()
y=['y']
X=[i for i in data_final_vars if i not in y]
```
### Feature Selection
```
from sklearn import datasets
from sklearn.feature_selection import RFE
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression()
rfe = RFE(logreg, 18)
rfe = rfe.fit(data_final[X], data_final[y] )
print(rfe.support_)
print(rfe.ranking_)
```
The Recursive Feature Elimination (RFE) has helped us select the following features: "previous", "euribor3m", "job_blue-collar", "job_retired", "job_services", "job_student", "default_no", "month_aug", "month_dec", "month_jul", "month_nov", "month_oct", "month_sep", "day_of_week_fri", "day_of_week_wed", "poutcome_failure", "poutcome_nonexistent", "poutcome_success".
```
cols=["previous", "euribor3m", "job_blue-collar", "job_retired", "job_services", "job_student", "default_no",
"month_aug", "month_dec", "month_jul", "month_nov", "month_oct", "month_sep", "day_of_week_fri", "day_of_week_wed",
"poutcome_failure", "poutcome_nonexistent", "poutcome_success"]
X=data_final[cols]
y=data_final['y']
```
### Implementing the model
```
import statsmodels.api as sm
logit_model=sm.Logit(y,X)
result=logit_model.fit()
print(result.summary())
```
The p-values for most of the variables are very small, therefore, most of them are significant to the model.
### Logistic Regression Model Fitting
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
```
#### Predicting the test set results and caculating the accuracy
```
y_pred = logreg.predict(X_test)
print('Accuracy of logistic regression classifier on test set: {:.2f}'.format(logreg.score(X_test, y_test)))
```
### Cross Validation
```
from sklearn import model_selection
from sklearn.model_selection import cross_val_score
kfold = model_selection.KFold(n_splits=10, random_state=7)
modelCV = LogisticRegression()
scoring = 'accuracy'
results = model_selection.cross_val_score(modelCV, X_train, y_train, cv=kfold, scoring=scoring)
print("10-fold cross validation average accuracy: %.3f" % (results.mean()))
```
### Confusion Matrix
```
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
```
The result is telling us that we have 10872+254 correct predictions and 1122+109 incorrect predictions.
#### Accuracy
```
print('Accuracy of logistic regression classifier on test set: {:.2f}'.format(classifier.score(X_test, y_test)))
```
#### Compute precision, recall, F-measure and support
The precision is the ratio tp / (tp + fp) where tp is the number of true positives and fp the number of false positives. The precision is intuitively the ability of the classifier not to label as positive a sample that is negative.
The recall is the ratio tp / (tp + fn) where tp is the number of true positives and fn the number of false negatives. The recall is intuitively the ability of the classifier to find all the positive samples.
The F-beta score can be interpreted as a weighted harmonic mean of the precision and recall, where an F-beta score reaches its best value at 1 and worst score at 0.
The F-beta score weights recall more than precision by a factor of beta. beta == 1.0 means recall and precision are equally important.
The support is the number of occurrences of each class in y_test.
```
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
```
#### Interpretation:
Of the entire test set, 88% of the promoted term deposit were the term deposit that the customers liked. Of the entire test set, 90% of the customer's preferred term deposit were promoted.
### ROC Curvefrom sklearn import metrics
from ggplot import *
prob = clf1.predict_proba(X_test)[:,1]
fpr, sensitivity, _ = metrics.roc_curve(Y_test, prob)
df = pd.DataFrame(dict(fpr=fpr, sensitivity=sensitivity))
ggplot(df, aes(x='fpr', y='sensitivity')) +\
geom_line() +\
geom_abline(linetype='dashed')
```
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
logit_roc_auc = roc_auc_score(y_test, logreg.predict(X_test))
fpr, tpr, thresholds = roc_curve(y_test, logreg.predict_proba(X_test)[:,1])
plt.figure()
plt.plot(fpr, tpr, label='Logistic Regression (area = %0.2f)' % logit_roc_auc)
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.savefig('Log_ROC')
plt.show()
```
| github_jupyter |
```
from __future__ import print_function
import os
import sys
import time
import argparse
import datetime
import math
import pickle
import torchvision
import torchvision.transforms as transforms
from utils.autoaugment import CIFAR10Policy
import torch
import torch.utils.data as data
import numpy as np
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
from torch.autograd import Variable
from torchvision import datasets
from torch.utils.data.sampler import SubsetRandomSampler
# from utils.BBBlayers import GaussianVariationalInference
# from utils.BayesianModels.Bayesian3Conv3FC import BBB3Conv3FC
from utils.BayesianModels.BayesianAlexNet import BBBAlexNet
# from utils.BayesianModels.BayesianLeNet import BBBLeNet
# from utils.BayesianModels.BayesianSqueezeNet import BBBSqueezeNet
net_type = 'alexnet'
dataset = 'CIFAR10'
outputs = 10
inputs = 3
resume = False
n_epochs = 150
lr = 0.01
weight_decay = 0.0005
num_samples = 1
beta_type = "Standard"
resize=32
# Hyper Parameter settings
use_cuda = torch.cuda.is_available()
torch.cuda.set_device(0)
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 256
# percentage of training set to use as validation
valid_size = 0.2
# convert data to a normalized torch.FloatTensor
transform = transforms.Compose([
transforms.RandomHorizontalFlip(), # randomly flip and rotate
transforms.RandomRotation(10),
transforms.ToTensor(),
# transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
# choose the training and test datasets
train_data = datasets.CIFAR10('data', train=True,
download=True, transform=transform)
test_data = datasets.CIFAR10('data', train=False,
download=True, transform=transform)
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders (combine dataset and sampler)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
# specify the image classes
classes = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
import matplotlib.pyplot as plt
%matplotlib inline
# helper function to un-normalize and display an image
def imshow(img):
# Uncomment if normalizing the data
#img = img / 2 + 0.5 # unnormalize
plt.imshow(np.transpose(img, (1, 2, 0))) # convert from Tensor image
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
# display 20 images
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
ax.set_title(classes[labels[idx]])
rgb_img = np.squeeze(images[3])
channels = ['red channel', 'green channel', 'blue channel']
fig = plt.figure(figsize = (36, 36))
for idx in np.arange(rgb_img.shape[0]):
ax = fig.add_subplot(1, 3, idx + 1)
img = rgb_img[idx]
ax.imshow(img, cmap='gray')
ax.set_title(channels[idx])
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center', size=8,
color='white' if img[x][y]<thresh else 'black')
# Architecture
if (net_type == 'lenet'):
net = BBBLeNet(outputs,inputs)
elif (net_type == 'alexnet'):
net = BBBAlexNet( outputs, inputs)
#net = BBBAlexNet(outputs,inputs)
elif (net_type == '3conv3fc'):
net = BBB3Conv3FC(outputs,inputs)
else:
print('Error : Network should be either [LeNet / AlexNet / 3Conv3FC')
if use_cuda:
net.cuda()
ckpt_name = f'model_{net_type}_{dataset}_bayesian.pt'
ckpt_name
def get_beta(batch_idx, m, beta_type):
if beta_type == "Blundell":
beta = 2 ** (m - (batch_idx + 1)) / (2 ** m - 1)
elif beta_type == "Soenderby":
beta = min(epoch / (num_epochs // 4), 1)
elif beta_type == "Standard":
beta = 1 / m
else:
beta = 0
return beta
def elbo(out, y, kl, beta):
loss = F.cross_entropy(out, y)
return loss + beta * kl
def train(epoch):
print('Epoch: %d' % epoch)
net.train()
train_loss = 0
correct = 0
total = 0
for batch_idx, (inputs, targets) in enumerate(train_loader):
inputs, targets = inputs.cuda(), targets.cuda()
optimizer.zero_grad()
outputs, kl = net.probforward(inputs)
loss = elbo(outputs, targets, kl, get_beta(epoch, len(train_data), beta_type))
loss.backward()
optimizer.step()
pred = torch.max(outputs, dim=1)[1]
correct += torch.sum(pred.eq(targets)).item()
total += targets.numel()
print(f'[TRAIN] Acc: {100.*correct/total:.3f}')
def test(epoch):
net.eval()
test_loss = 0
correct = 0
total = 0
accuracy_max = 0
with torch.no_grad():
for batch_idx, (inputs, targets) in enumerate(test_loader):
inputs, targets = inputs.cuda(), targets.cuda()
outputs, _ = net.probforward(inputs)
_, predicted = outputs.max(1)
total += targets.size(0)
correct += predicted.eq(targets).sum().item()
accuracy = 100.*correct/total
print(f'[TEST] Acc: {accuracy:.3f}')
torch.save(net.state_dict(), ckpt_name)
epochs = [80, 60, 40, 20]
count = 0
from torch.optim import Adam
for epoch in epochs:
optimizer = Adam(net.parameters(), lr=lr)
for _ in range(epoch):
train(count)
test(count)
count += 1
lr /= 10
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
images.numpy()
# move model inputs to cuda, if GPU available
if use_cuda:
images = images.cuda()
# get sample outputs
output, kl = net.probforward(images)
# convert output probabilities to predicted class
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.numpy()) if not use_cuda else np.squeeze(preds_tensor.cpu().numpy())
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images.cpu()[idx])
ax.set_title("{} ({})".format(classes[preds[idx]], classes[labels[idx]]),
color=("green" if preds[idx]==labels[idx].item() else "red"))
```
| github_jupyter |
```
import panel as pn
pn.extension('plotly')
```
The ``Plotly`` pane renders Plotly plots inside a panel. It optimizes the plot rendering by using binary serialization for any array data found on the Plotly object, providing efficient updates. Note that to use the Plotly pane in a Jupyter notebook, the Panel extension has to be loaded with 'plotly' as an argument to ensure that Plotly.js is initialized.
#### Parameters:
For layout and styling related parameters see the [customization user guide](../../user_guide/Customization.ipynb).
* **``object``** (object): The Plotly figure being displayed
* **``click_data``** (dict): Click callback data
* **``clickannotation_data``** (dict): Clickannotation callback data
* **``config``** (dict): Config data
* **``hover_data``** (dict): Hover callback data
* **``relayout_data``** (dict): Relayout callback data
* **``restyle_data``** (dict): Restyle callback data
* **``selected_data``** (dict): Selected callback data
* **``viewport``** (dict): Current viewport state
* **``viewport_update_policy``** (str, default = 'mouseup'): Policy by which the viewport parameter is updated during user interactions
* ``mouseup``: updates are synchronized when mouse button is released after panning
* ``continuoius``: updates are synchronized continually while panning
* ``throttle``: updates are synchronized while panning, at intervals determined by the viewport_update_throttle parameter
* **``viewport_update_throttle``** (int, default = 200, bounds = (0, None)): Time interval in milliseconds at which viewport updates are synchronized when viewport_update_policy is "throttle".
___
As with most other types ``Panel`` will automatically convert a Plotly figure to a ``Plotly`` pane if it is passed to the ``pn.panel`` function or a panel layout, but a ``Plotly`` pane can be constructed directly using the ``pn.pane.Plotly`` constructor:
```
import numpy as np
import plotly.graph_objs as go
xx = np.linspace(-3.5, 3.5, 100)
yy = np.linspace(-3.5, 3.5, 100)
x, y = np.meshgrid(xx, yy)
z = np.exp(-(x-1)**2-y**2)-(x**3+y**4-x/5)*np.exp(-(x**2+y**2))
surface = go.Surface(z=z)
layout = go.Layout(
title='Plotly 3D Plot',
autosize=False,
width=500,
height=500,
margin=dict(t=50, b=50, r=50, l=50)
)
fig = dict(data=[surface], layout=layout)
plotly_pane = pn.pane.Plotly(fig)
plotly_pane
```
Once created the plot can be updated by modifying the Plotly traces and then triggering an update by setting or triggering an event on the pane ``object``. Note that this only works if the ``Figure`` is defined as a dictionary, since Plotly will make copies of the traces, which means that modifying them in place has no effect. Modifying an array will send just the array using a binary protocol, leading to fast and efficient updates.
```
surface.z = np.sin(z+1)
plotly_pane.object = fig
```
Similarly, modifying the plot ``layout`` will only modify the layout, leaving the traces unaffected.
```
fig['layout']['width'] = 800
plotly_pane.object = fig
```
The Plotly pane supports layouts and subplots of arbitrary complexity, allowing even deeply nested Plotly figures to be displayed:
```
from plotly import subplots
heatmap = go.Heatmap(
z=[[1, 20, 30],
[20, 1, 60],
[30, 60, 1]],
showscale=False)
y0 = np.random.randn(50)
y1 = np.random.randn(50)+1
box_1 = go.Box(y=y0)
box_2 = go.Box(y=y1)
data = [heatmap, box_1, box_2]
fig = subplots.make_subplots(
rows=2, cols=2, specs=[[{}, {}], [{'colspan': 2}, None]],
subplot_titles=('First Subplot','Second Subplot', 'Third Subplot')
)
fig.append_trace(box_1, 1, 1)
fig.append_trace(box_2, 1, 2)
fig.append_trace(heatmap, 2, 1)
fig['layout'].update(height=600, width=600, title='i <3 subplots')
fig = fig.to_dict()
subplot_panel = pn.pane.Plotly(fig)
subplot_panel
```
Just like in the single-subplot case we can modify just certain aspects of a plot and then trigger an update. E.g. here we replace the overall title text:
```
fig['layout']['title']['text'] = 'i <3 updating subplots'
subplot_panel.object = fig
```
Lastly, Plotly plots can be made responsive using the `autosize` option on a Plotly layout:
```
import pandas as pd
import plotly.express as px
data = pd.DataFrame([
('Monday', 7), ('Tuesday', 4), ('Wednesday', 9), ('Thursday', 4),
('Friday', 4), ('Saturday', 4), ('Sunay', 4)], columns=['Day', 'Orders']
)
fig = px.line(data, x="Day", y="Orders")
fig.update_traces(mode="lines+markers", marker=dict(size=10), line=dict(width=4))
fig.layout.autosize = True
responsive = pn.pane.Plotly(fig, config={'responsive': True})
pn.Column('# A responsive plot', responsive, sizing_mode='stretch_width')
```
### Controls
The `Plotly` pane exposes a number of options which can be changed from both Python and Javascript try out the effect of these parameters interactively:
```
pn.Row(responsive.controls(jslink=True), responsive)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/afeld/python-public-policy/blob/main/lecture_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# a bit of setup
import numpy as np
np.random.seed(5)
```
# **NYU Wagner - Python Coding for Public Policy**
# Class 1: Intro to Pandas
# LECTURE
# Pandas
- A Python package (bundled up code that you can reuse)
- Very common for data science in Python
- [A lot like R](https://pandas.pydata.org/docs/getting_started/comparison/comparison_with_r.html), in their core concept being data frames
## Start by importing necessary packages
_Hold off on coding until the in-class exercise._
```
import pandas as pd
```
## Read and save 311 Service Requests dataset as a pandas dataframe
This will take a while (~30 seconds).
```
df = pd.read_csv('https://storage.googleapis.com/python-public-policy/data/311_Service_Requests_2018-19.csv.zip')
```
Ignore the `DtypeWarning` for now; we'll come back to it.
## Today's goal
Learn which 311 complaints are most common and which agencies are responsible for handling them. But first, let's take a look at the data, then clean it up!
## Preview the data contents
```
df.head() # defaults to providing the first 5 if you don't specify a number
df.tail(10) # last 10 records in the dataframe
df.sample(5) # random sample of size determined by you
```
## Learning more about the data
[Source page](https://data.cityofnewyork.us/Social-Services/311-Service-Requests-from-2010-to-Present/erm2-nwe9)
## Pandas data structures
<!-- source: https://docs.google.com/document/d/1HGw2BdbuXSIwcgDWXkzZGPXYr5yJ_WEM3Gw-nLoHzCo/edit#heading=h.7z4rqdvodt9j -->

## How many records are in the dataset?
### `size` method
How many cells are there in the data table?
```
df.size
```
What if I only care about how many rows there are? The columns in the dataframe are like a list. You can use a column name as an index to get one column from the dataframe.
`.size` includes `null` (empty) values.
```
df['Facility Type'].size
```
### `count()` method
You can also use the `count()` function, which gives the count of values per column. `count()` doesn't include `null` (empty) values.
```
df.count()
```
To just get the count in the "unique key" column:
```
df['Unique Key'].count()
```
### `info()` method
```
df.info()
```
## What are the distinct sets of values in columns that seem most useful?
### [`unique()`](https://pandas.pydata.org/docs/reference/api/pandas.Series.unique.html) method for getting list of unique values
Let's look at the "status" column. What are the status options for these 311 complaints?
```
df['Status'].unique()
df['Open Data Channel Type'].unique()
df['Agency'].unique()
df['Complaint Type'].unique()
```
## In-class exercise
[Create a copy of the Homework 1 starter notebook.](https://colab.research.google.com/github/afeld/python-public-policy/blob/main/hw_1.ipynb)
## Excluding bad records from the dataframe
First, let's refresh ourselves on what the invalid complaint types are, by getting the distinct list of all complaint types
```
df['Complaint Type'].unique()
```
Let's make that a bit easier to read:
```
complaints = df['Complaint Type'].unique()
complaints.sort()
list(complaints)
```
Let's see how frequently these invalid Complaint Type values appear in the data.
Use `.groupby().size()` to get the count of 311 requests per complaint type value. This is very similar to [pivot tables](https://support.google.com/docs/answer/1272900) in spreadsheets.
```
# remember .size gives you the count of cells across all columns in the dataframe
df.size
# to just get the total count of records in the dataset, we should get the size of the 'Unique Key' column
df['Unique Key'].size
with pd.option_context("display.max_rows", None):
display(df.groupby('Complaint Type').size())
```
```python
with pd.option_context("display.max_rows", None):
display(...)
```
What this code is doing: showing all cells in a DataFrame with [rich output](https://ipython.readthedocs.io/en/stable/interactive/plotting.html#rich-outputs).
```python
df.groupby('Complaint Type').size()
```
What this code is doing:
1. Group the records in the dataset based on their `Complaint Type` value
1. Count the records that have been grouped together by their shared `Complaint Type` value
Watch out! `.groupby().size()` function doesn't work the same way as `.size`. The former gets the count of number of rows in each group.
It looks like most invalid complaint types only have a few records. Try excluding all complaint type categories with < 4 records, assuming that all complaint type categories with < 4 instances in the data are bad data entries.
Why 4? It's arbitrary. We're looking for trends in the data in this case don't care about low frequency entries.
Create a dataframe that captures the count of records per `Complaint Type` value.
```
counts = df.groupby('Complaint Type').size().reset_index(name='count')
counts
# .reset_index(name='count') allows us to name the new column that contains the count of rows
```
You can also use `.count()` but [the output is a little different](https://stackoverflow.com/questions/33346591/what-is-the-difference-between-size-and-count-in-pandas).
Create a "series" that only lists the `Complaint Type` values that have record counts > 4. (Remember: A single column from a pandas dataframe is called a series. It's essentially a list containing all the values in the column.)
```
valid_complaint_types = counts['Complaint Type'][counts['count'] > 4]
valid_complaint_types
```
Filter our `df` dataframe to only keep the rows where the `Complaint Type` value is in the `valid_complaint_types` series we created in the previous step. Save the result in a new dataframe.
```
df_cleaned = df[df['Complaint Type'].isin(valid_complaint_types)]
```
How can we make sure this worked? Let's check how many records there were originally in `df` vs how many are in `df_cleaned`.
Before:
```
df['Unique Key'].size
```
After:
```
df_cleaned['Unique Key'].size
```
We can also print the set of complaint_type values from our cleaned dataframe to make sure they look correct.
```
df_cleaned['Complaint Type'].sample(10)
```
Great, now those invalid records will be excluded from our analysis!
Another approach to excluding those invalid records would be to use ["regex" (regular expressions)](https://www.w3schools.com/python/python_regex.asp) to find records with weird characters.
## Filtering rows
Slicing and dicing is done through [indexing](https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html).

### Boolean indexing

#### How it works
```
%%html
<video controls width="700">
<source src="https://github.com/afeld/python-public-policy/raw/main/img/boolean-indexing.mp4" type="video/mp4">
</video>
```
## Done with clean up! Time for the actual analysis:
### Which 311 complaints are most common and which agencies are responsible for handling them?
#### Which complaints are the most common?
```
df_cleaned.groupby('Complaint Type').size().reset_index(name='count')
# .reset_index(name='count') isn't necessary but it's helpful to include because it allows us to name the new column that contains the count of rows
```
#### Which agencies are responsible for handling these complaint categories?
```
df_cleaned.groupby(['Agency', 'Complaint Type']).size().reset_index(name='count')
```
#### Which agencies receive the most total 311 requests?
```
df_cleaned.groupby('Agency').size().reset_index(name='count')
```
#### What is the most frequent request per agency?
First, create a dataframe that contains the count of complaints per `Agency` per `Complaint Type`.
```
agency_counts = df_cleaned.groupby(['Agency', 'Complaint Type']).size().reset_index(name='count')
agency_counts.head(20)
```
Use `drop_duplicates()` to keep the row with the highest value per `Agency`.
```
agency_counts.sort_values('count', ascending=False).drop_duplicates('Agency').sort_values('Agency')
```
Another way, only sorting it once:
```
agency_counts.sort_values(['Agency', 'count']).drop_duplicates('Agency', keep='last')
```
# HOMEWORK 1
Continue in the notebook you created.
| github_jupyter |
# Solving combinatorial optimization problems using QAOA
In this tutorial, we introduce combinatorial optimization problems, explain approximate optimization algorithms, explain how the Quantum Approximate Optimization Algorithm (QAOA) works and present the implementation of an example that can be run on a simulator or on a 5 qubit quantum chip
## Contents
1. [Introduction](#introduction)
2. [Examples](#examples)
3. [Approximate optimization algorithms](#approximateOPT)
4. [The QAOA algorithm](#QAOA)
5. [Qiskit Implementation](#implementation)
5.1 [Running QAOA on a simulator](#implementationsim)
5.2 [Running QAOA on a real quantum device](#implementationdev)
6. [Problems](#problems)
7. [References](#references)
## 1. Introduction <a id='introduction'></a>
Combinatorial optimization [1](#references) means searching for an optimal solution in a finite or countably infinite set of potential solutions. Optimality is defined with respect to some criterion function, which is to be minimized or maximized, which is typically called the cost function.
There are various types of optimization problems. These include Minimization: cost, distance, length of a traversal, weight, processing time, material, energy consumption, number of objects. Maximization: profit, value, output, return, yield, utility, efficiency, capacity, number of objects. Any maximization problem can be cast in terms of a minimization problem and vice versa. Hence the general form a combinatorial optimization problem is given by
$$ \text{maximize } \;\; C(x)$$
$$ \text{subject to } \;\; x \in S $$
where $x \in S$, is a discrete variable and $C : D \rightarrow \mathbb{R}$ is the cost function, that maps from some domain $S$ in to the real numbers $\mathbb{R}$. The variable $x$ can be subject to a set of constraints and lies within the set $S \subset D$ of feasible points.
In binary combinatorial optimization problems, the cost function $C$ can typically be expressed as a sum of terms that only involve a subset $Q \subset[n]$ of the $n$ bits in the string $x \in \{0,1\}^n$ and is written in the canonical form
$$ C(x) = \sum_{(Q,\overline{Q}) \subset [n]} w_{(Q,\overline{Q})} \; \prod_{i\in Q} x_i \; \prod_{j\in \overline{Q}} (1- x_j), $$
where $x_i \in \{0,1\}$ and $w_{(Q,\overline{Q})}\in \mathbb{R}$. We want to find the n-bit string $x$ for which $C(x)$ is the maximal.
### 1.1 Diagonal Hamiltonians
This cost function can be mapped to a Hamiltonian that is diagonal in the computational basis. Given the cost-function $C$ this Hamiltonian is then written as
$$ H = \sum_{x \in \{0,1\}^n} C(x) |x \rangle\langle x| $$
where $x \in \{0,1\}^n$ labels the computational basis states $|x \rangle \in \mathbb{C}^{2^n}$. If the cost function only has at most weight $k$ terms, i.e. when only $Q$ contribute that involve at most $Q \leq k$ bits, then this diagonal Hamiltonian is also only a sum of weight $k$ Pauli $Z$ operators.
The expansion of $H$ in to Pauli $Z$ operators can be obtained from the canonical expansion of the cost-function $C$ by substituting for every binary variable $x_i \in \{0,1\}$ the matrix $x_i \rightarrow 2^{-1}(1 - Z_i)$. Here $Z_i$ is read as the Pauli $Z$ operator that acts on qubit $i$ and trivial on all others, i.e.
$$ Z_i = \left(\begin{array}{cc} 1 & 0 \\ 0 & -1 \end{array}\right). $$
This means that the spin Hamiltonian encoding the classical cost function is written as a $|Q|$ - local quantum spin Hamiltonian only involving Pauli $Z$- operators.
$$ H = \sum_{(Q,\overline{Q}) \subset [n]} w_{(Q,\overline{Q})} \; \frac{1}{2^{|Q| + |\overline{Q}|}}\prod_{i\in Q} \left(1 - Z_i\right) \; \prod_{j\in \overline{Q}} \left(1 + Z_j\right).$$
Now, we will assume that only a few (polynomially many in $n$) $w_{(Q,\overline{Q})}$ will be non-zero. Moreover we will assume that the set $|(Q,\overline{Q})|$ is bounded and not too large. This means we can write the cost function as well as the Hamiltonian $H$ as the sum of $m$ local terms $\hat{C}_k$,
$$ H = \sum_{k = 1}^m \hat{C}_k, $$
where both $m$ and the support of $\hat{C}_k$ is reasonably bounded.
## 2 Examples: <a id='examples'></a>
We consider 2 examples to illustrate combinatorial optimization problems. We will only implement the first example as in Qiskit, but provide a sequence of exercises that give the instructions to implement the second example as well.
### 2.1 (weighted) $MAXCUT$
Consider an $n$-node non-directed graph *G = (V, E)* where *|V| = n* with edge weights $w_{ij}>0$, $w_{ij}=w_{ji}$, for $(j,k)\in E$. A cut is defined as a partition of the original set V into two subsets. The cost function to be optimized is in this case the sum of weights of edges connecting points in the two different subsets, *crossing* the cut. By assigning $x_i=0$ or $x_i=1$ to each node $i$, one tries to maximize the global profit function (here and in the following summations run over indices 0,1,...n-1)
$$C(\textbf{x}) = \sum_{i,j = 1}^n w_{ij} x_i (1-x_j).$$
To simplify notation, we assume uniform weights $ w_{ij} = 1$ for $(i,j) \in E$. In order to find a solution to this problem on a quantum computer, one needs first to map it to a diagonal Hamiltonian as discussed above. We write the sum as a sum over edges in the set $(i,j) = E$
$$C(\textbf{x}) = \sum_{i,j = 1}^n w_{ij} x_i (1-x_j) = \sum_{(i,j) \in E} \left( x_i (1-x_j) + x_j (1-x_i)\right)$$
To map it to a spin Hamiltonian, we make the assignment $x_i\rightarrow (1-Z_i)/2$, where $Z_i$ is the Pauli Z operator that has eigenvalues $\pm 1$ and obtain $X \rightarrow H$
$$ H = \sum_{(j,k) \in E} \frac{1}{2}\left(1 - Z_j Z_k \right).$$
This means that the Hamiltonian can be written as a sum of $m = |E|$ local terms $\hat{C}_e = \frac{1}{2}\left(1 - Z_{e1}Z_{e2}\right)$ with $e = (e1,e2) \in E$.
### 2.2 Constraint satisfaction problems and $MAX \; 3-SAT$.
Another example of a combinatorial optimization problem is $3-SAT$. Here the cost function $C(\textbf{x}) = \sum_{k = 1}^m c_k(\textbf{x})$ is a sum of clauses $c_k(\textbf{x})$ that constrain the values of $3$ bits of some $\textbf{x} \in \{0,1\}^n$ that participate in the clause. Consider for instance this example of a $3-SAT$ clause
$$ c_1(\textbf{x}) = (1-x_1)(1-x_3)x_{132} $$
for a bit string $\textbf{x} \in \{0,1\}^{133}$. The clause can only be satisfied by setting the bits $x_1 = 0$,$x_3 = 0$ and $x_{132} = 1$. The $3-SAT$ problem now asks whether there is a bit string that satisfies all of the $m$ clauses or whether no such string exists. This decision problem is the prime example of a problem that is $NP$-complete.
The closely related optimization problem $MAX \; 3-SAT$ asks to find the bit string $\textbf{x}$ that satisfies the maximal number of of clauses in $C(\textbf{x})$. This can of course be turned again in to a decision problem if we ask where there exists a bit string that satisfies more than $\tilde{m}$ of the $m$ clauses, which is again $NP$-complete.
## 3. Approximate optimization algorithms <a id='approximateOPT'></a>
Both the previously considered problems $MAXCUT$ and $MAX \; 3-SAT$ are actually known to be a NP-hard problems [1](#references). In fact it turns out that many combinatorial optimization problems are computationally hard to solve in general. In light of this fact, we can't expect to find a provably efficient algorithm, i.e. an algorithm with polynomial runtime in the problem size, that solves these problems. This also applies to quantum algorithms. There are two main approaches to dealing with such problems. First approach is approximation algorithms that are guaranteed to find solution of specified quality in polynomial time. The second approach are heuristic algorithms that don't have a polynomial runtime guarantee but appear to perform well on some instances of such problems.
Approximate optimization algorithms are efficient and provide a provable guarantee on how close the approximate solution is to the actual optimum of the problem. The guarantee typically comes in the form of an approximation ratio, $\alpha \leq 1$. A probabilistic approximate optimization algorithm guarantees that it produces a bit-string $\textbf{x}^* \in \{0,1\}^n$ so that *with high probability* we have that with a positive $C_{max} = \max_{\textbf{x}}C(\textbf{x})$
$$ C_{max} \geq C(\textbf{x}^*) \geq \alpha C_{max}. $$
For the $MAXCUT$ problem there is a famous approximate algorithm due to Goemans and Williamson [2](#references) . This algorithm is based on an SDP relaxation of the original problem combined with a probabilistic rounding technique that yields an with high probability approximate solution $\textbf{x}^*$ that has an approximation ratio of $\alpha \approx 0.878$. This approximation ratio is actually believed to optimal so we do not expect to see an improvement by using a quantum algorithm.
## 4. The QAOA algorithm <a id="QAOA"></a>
The Quantum approximate optimization algorithm (QAOA) by Farhi, Goldstone and Gutmann [3](#references) is an example of a heuristic algorithm. Unlike Goemans-Williamson algorithm, QAOA does not come with performance guarantees.
QAOA takes the approach of classical approximate algorithms and looks for a quantum analogue that will likewise produce a classical bit string $x^*$ that with high probability is expected to have a good approximation ratio $\alpha$. Before discussing the details, let us first present the general idea of this approach.
### 4.1 Overview:
We want to find a quantum state $|\psi_p(\vec{\gamma},\vec{\beta})\rangle$, that depends on some real parameters $\vec{\gamma},\vec{\beta} \in \mathbb{R}^p$, which has the property that it maximizes the expectation value with respect to the problem Hamiltonian $H$. Given this trial state we search for parameters $\vec{\gamma}^*,\vec{\beta}^*$ that maximize $F_p(\vec{\gamma},\vec{\beta}) = \langle \psi_p(\vec{\gamma},\vec{\beta})|H|\psi_p(\vec{\gamma},\vec{\beta})\rangle$.
Once we have such a state and the corresponding parameters we prepare the state $|\psi_p(\vec{\gamma}^*,\vec{\beta}^*)\rangle$ on a quantum computer and measure the state in the $Z$ basis $|x \rangle = |x_1,\ldots x_n \rangle$ to obtain a random outcome $x^*$.
We will see that this random $x^*$ is going to be a bit string that is with high probability close to the expected value $M_p = F_p(\vec{\gamma}^*,\vec{\beta}^*)$. Hence, if $M_p$ is close to $C_{max}$ so is $C(x^*)$.
### 4.2 The components of the QAOA algorithm.
### 4.2.1 The QAOA trial state <a id="section_421"></a>
Central to QAOA is the trial state $|\psi_p(\vec{\gamma},\vec{\beta})\rangle$ that will be prepared on the quantum computer. Ideally we want this state to give rise to a large expectation value $F_p(\vec{\gamma},\vec{\beta}) = \langle \psi_p(\vec{\gamma},\vec{\beta})|H|\psi_p(\vec{\gamma},\vec{\beta})\rangle$ with respect to the problem Hamiltonian $H$. In Farhi [3](#references), the trial states $|\psi_p(\vec{\gamma},\vec{\beta})\rangle$ are constructed from the problem Hamiltonian $H$ together with single qubit Pauli $X$ rotations. That means, given a problems Hamiltonian
$$ H = \sum_{k = 1}^m \hat{C}_k $$
diagonal in the computational basis and a transverse field Hamiltonian
$$ B = \sum_{i = 1}^n X_i $$
the trial state is prepared by applying $p$ alternating unitaries
$$ |\psi_p(\vec{\gamma},\vec{\beta})\rangle = e^{ -i\beta_p B } e^{ -i\gamma_p H } \ldots e^{ -i\beta_1 B } e^{ -i\gamma_1 H } |+\rangle^n $$
to the product state $|+\rangle^n$ with $ X |+\rangle = |+\rangle$.
This particular ansatz has the advantage that there exists an explicit choice for the vectors $\vec{\gamma}^*,\vec{\beta}^*$ such that for $M_p = F_p(\vec{\gamma}^*,\vec{\beta}^*)$ when we take the limit $\lim_{p \rightarrow \infty} M_p = C_{max}$. This follows by viewing the trial state $|\psi_p(\vec{\gamma},\vec{\beta}) \rangle$ as the state that follows from trotterizing the adiabatic evolution with respect to $H$ and the transverse field Hamiltonian $B$, c.f. Ref [3](#references).
Conversely the disadvantage of this trial state is one would typically want a state that has been generated from a quantum circuit that is not too deep. Here depth is measured with respect to the gates that can be applied directly on the quantum chip. Hence there are other proposals that suggest using Ansatz trial state that are more tailored to the Hardware of the quantum chip Ref. [4](#references), Ref. [5](#references).
### 4.2.2 Computing the expectation value <a id="section_422"></a>
An important component of this approach is that we will have to compute or estimate the expectation value
$$
F_p(\vec{\gamma},\vec{\beta}) = \langle \psi_p(\vec{\gamma},\vec{\beta})|H|\psi_p(\vec{\gamma},\vec{\beta})\rangle
$$
so we can optimize the parameters $\vec{\gamma},\vec{\beta}$. We will be considering two scenarios here.
#### Classical evaluation
Note that when the circuit to prepare $|\psi_p(\vec{\gamma},\vec{\beta})\rangle$ is not too deep it may be possible to evaluate the expectation value $F_p$ classically.
This happens for instance when one considers $MAXCUT$ for graphs with bounded degree and one considers a circuit with $p=1$. We will see an example of this in the Qiskit implementation below (section 5.2) and provide an exercise to compute the expectation value.
To illustrate the idea, recall that the Hamiltonian can be written as a sum of individual terms $H = \sum_{k = 1}^m \hat{C}_k$. Due to the linearity of the expectation value, it is sufficient to consider the expectation values of the individual summands. For $p = 1$ one has that
$$ \langle \psi_1(\vec{\gamma},\vec{\beta})|\hat{C}_k|\psi_1(\vec{\gamma},\vec{\beta})\rangle = \langle +^n | e^{ i\gamma_1 H } e^{ i\beta_1 B } | \hat{C}_k | e^{ -i\beta_1 B } e^{ -i\gamma_1 H } |+^n\rangle.$$
Observe that with $B = \sum_{i = 1}^n X_i$ the unitary $e^{ -i\beta_1 B }$ is actually a product of single qubit rotations about $X$ with an angle $\beta$ for which we will write $X(\beta)_k = \exp(i\beta X_k)$.
All the individual rotations that don't act on the qubits where $\hat{C}_k$ is supported commute with $\hat{C}_k$ and therefore cancel. This does not increase the support of the operator $\hat{C}_k$. This means that the second set of unitary gates $e^{ -i\gamma_1 H } = \prod_{l=1}^m U_l(\gamma)$ have a large set of gates $U_l(\gamma) = e^{ -i\gamma_1 \hat{C}_l }$ that commute with the operator $e^{ i\beta_1 B } \hat{C}_k e^{ -i\beta_1 B }$. The only gates $U_l(\gamma) = e^{ -i\gamma_1 \hat{C}_l }$ that contribute to the expectation value are those which involve qubits in the support of the original $\hat{C}_k$.
Hence, for bounded degree interaction the support of $e^{ i\gamma_1 H } e^{ i\beta_1 B } \hat{C}_k e^{ -i\beta_1 B } e^{ -i\gamma_1 H }$ only expands by an amount given by the degree of the interaction in $H$ and is therefore independent of the system size. This means that for these smaller sub problems the expectation values are independent of $n$ and can be evaluated classically. The case of a general degree $3$ is considered in [3](#references).
This is a general observation, which means that if we have a problem where the circuit used for the trial state preparation only increases the support of each term in the Hamiltonian by a constant amount the cost function can be directly evaluated.
When this is the case, and only a few parameters $\beta, \gamma$ are needed in the preparation of the trial state,
these can be found easily by a simple grid search. Furthermore, an exact optimal value of $M_p$ can be used to bound the approximation ratio
$$ \frac{M_p}{C_{max}} \geq \alpha $$
to obtain an estimate of $\alpha$. For this case the QAOA algorithm has the same characteristics as a conventional approximate optimization algorithm that comes with a guaranteed approximation ratio that can be obtained with polynomial efficiency in the problem size.
#### Evaluation on a quantum computer
When the quantum circuit becomes too deep to be evaluated classically, or when the connectivity of the Problem Hamiltonian is too high we can resort to other means of estimating the expectation value. This involves directly estimating $F_p(\vec{\gamma},\vec{\beta})$ on the quantum computer. The approach here follows the path of the conventional expectation value estimation as used in VQE [4](#references), where a trial state $| \psi_p(\vec{\gamma},\vec{\beta}) \rangle$ is prepared directly on the quantum computer and the expectation value is obtained from sampling.
Since QAOA has a diagonal Hamiltonian $H$ it is actually straight forward to estimate the expectation value. We only need to obtain samples from the trial state in the computational basis. Recall that $H = \sum_{x \in \{0,1\}^n} C(x) |x \rangle\langle x|$ so that we can obtain the sampling estimate of
$$ \langle \psi_p(\vec{\gamma},\vec{\beta})|H|\psi_p(\vec{\gamma},\vec{\beta})\rangle = \sum_{x \in \{0,1\}^n} C(x) |\langle x| \psi_p(\vec{\gamma},\vec{\beta}) \rangle |^2$$
by repeated single qubit measurements of the state $| \psi_p(\vec{\gamma},\vec{\beta}) \rangle $ in the $Z$ basis. For every bit string $x$ obtained from the distribution $|\langle x| \psi_p(\vec{\gamma},\vec{\beta}) \rangle |^2$ we evaluate the cost function $C(x)$ and average it over the total number of samples. The resulting empirical average approximates the expectation value up to an additive sampling error that lies within the variance of the state. The variance will be discussed below.
With access to the expectation value, we can now run a classical optimization algorithm, such as [6](#references),
to optimize the $F_p$.
While this approach does not lead to an a-priori approximation guarantee for $x^*$, the optimized function value
can be used later to provide an estimate for the approximation ratio $\alpha$.
### 4.3.3 Obtaining a solution with a given approximation ratio with high probability
The algorithm is probabilistic in nature and produces random bit strings from the distribution $|\langle x| \psi_p(\vec{\gamma},\vec{\beta}) \rangle |^2$. So how can we be sure that we will sample an approximation $x^*$ that is close to the value of the optimized expectation value $M_p$? Note that this question is also relevant to the estimation of $M_p$ on a quantum computer in the first place. If the samples drawn from $|\langle x| \psi_p(\vec{\gamma},\vec{\beta}) \rangle |^2$ have too much variance, many samples are necessary to determine the mean.
We will draw a bit string $x^*$ that is close to the mean $M_p$ with high probability when the energy as variable has little variance.
Note that the number of terms in the Hamiltonian $H = \sum_{k=1}^m \hat{C}_k$ are bounded by $m$. Say each individual
summand $\hat{C}_k$ has an operator norm that can be bounded by a universal constant $\|\hat{C}_k\| \leq \tilde{C}$ for all $k = 1\ldots m$. Then consider
$$
\begin{eqnarray}
\langle \psi_p(\vec{\gamma},\vec{\beta})|H^2|\psi_p(\vec{\gamma},\vec{\beta})\rangle - \langle \psi_p(\vec{\gamma},\vec{\beta})|H|\psi_p(\vec{\gamma},\vec{\beta})\rangle^2 &\leq & \langle \psi_p(\vec{\gamma},\vec{\beta})|H^2|\psi_p(\vec{\gamma},\vec{\beta})\rangle \\\nonumber
&=& \sum_{k,l =1}^m \langle \psi_p(\vec{\gamma},\vec{\beta})|\hat{C}_k \hat{C}_l |\psi_p(\vec{\gamma},\vec{\beta})\rangle \\\nonumber
&\leq& m^2 \tilde{C}^2 \\\nonumber
\end{eqnarray}
$$
where we have used that
$\langle \psi_p(\vec{\gamma},\vec{\beta})|\hat{C}_k \hat{C}_l |\psi_p(\vec{\gamma},\vec{\beta})\rangle \leq \tilde{C}^2$.
This means that the variance of any expectation $F_p(\vec{\gamma},\vec{\beta})$ is bounded by $m^2 \tilde{C}^2$. Hence this in particular applies for $M_p$. Furthermore if $m$ only grows polynomially in the number of qubits $n$, we know that taking polynomially growing number of samples $s = O\left(\frac{\tilde{C}^2 m^2}{\epsilon^2}\right)$ from $|\langle x| \psi_p(\vec{\gamma},\vec{\beta}) \rangle |^2$ will be sufficient to obtain a $x^*$ that leads to an $C(x^*)$ that will be close to $M_p$.
## 5. Qiskit Implementation<a id='implementation'></a>
As the example implementation we consider the $MAXCUT$ problem on the butterfly graph of the openly available IBMQ 5-qubit chip. The graph will be defined below and corresponds to the native connectivity of the device. This allows us to implement the original version of the $QAOA$ algorithm, where the cost function $C$ and the Hamiltonian $H$ that is used to generate the state coincide. Moreover, for such a simple graph the exact cost function can be calculated analytically, avoiding the need to find optimal parameters variationally [7](#references). To implement the circuit, we follow the notation and gate definitions from the [Qiskit Documentation](https://qiskit.org/documentation/).
As the first step will will load Qiskit and additional python packages.
```
import numpy as np
import networkx as nx # tool to handle general Graphs
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
from qiskit import Aer, IBMQ
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, transpile, assemble
from qiskit.providers.ibmq import least_busy
from qiskit.tools.monitor import job_monitor
from qiskit.visualization import plot_histogram
```
### 5.1 Problem definition
We define the cost function in terms of the butterfly graph of the superconducting chip. The graph has $n = 5$ vertices $ V = \{0,1,2,3,4\}$ and six edges $E = \{(0,1),(0,2),(1,2),(3,2),(3,4),(4,2)\}$, which will all carry the same unit weight $w_{ij} = 1$. We load an additional network package to encode the graph and plot connectivity below.
```
# Generating the butterfly graph with 5 nodes
n = 5
V = np.arange(0,n,1)
E =[(0,1,1.0),(0,2,1.0),(1,2,1.0),(3,2,1.0),(3,4,1.0),(4,2,1.0)]
G = nx.Graph()
G.add_nodes_from(V)
G.add_weighted_edges_from(E)
# Generate plot of the Graph
colors = ['r' for node in G.nodes()]
default_axes = plt.axes(frameon=True)
pos = nx.spring_layout(G)
nx.draw_networkx(G, node_color=colors, node_size=600, alpha=1, ax=default_axes, pos=pos)
```
### 5.2 Optimal trial state parameters<a id="implementation_sec52"></a>
In this example we consider the case for $p = 1$, i.e. only layer of gates. The expectation value $F_1(\gamma,\beta) = \langle \psi_1(\gamma,\beta))|H|\psi_1(\gamma,\beta) \rangle$ can be calculated analytically for this simple setting. Let us discuss the steps explicitly for the Hamiltonian $H = \sum_{(j,k) \in E} \frac{1}{2}\left(1 - Z_i Z_k\right)$. Due to the linearity of the expectation value we can compute the expectation value for the edges individually
$$f_{(i,k)}(\gamma,\beta) = \langle \psi_1(\gamma,\beta)|\;\frac{1}{2}\left(1 - Z_i Z_k\right)\;|\psi_1(\gamma,\beta)\rangle. $$
For the butterfly graph as plotted above, we observe that there are only two kinds of edges $A = \{(0,1),(3,4)\}$ and
$B = \{(0,2),(1,2),(2,3),(2,4)\}$. The edges in $A$ only have two neighboring edges, while the edges in $B$ have four. You can convince yourself that we only need to compute the expectation of a single edge in each set since the other expectation values will be the same. This means that we can compute $F_1(\gamma,\beta) = 2 f_A(\gamma,\beta) + 4f_B(\gamma,\beta)$ by evaluating only computing two expectation values. Note, that following the argument as outlined in [section 4.2.2](#section_422), all the gates that do not intersect with the Pauli operator $Z_0Z_1$ or $Z_0Z_2$ commute and cancel out so that we only need to compute
$$f_A(\gamma,\beta) = \frac{1}{2}\left(1 - \langle +^3|U_{21}(\gamma)U_{02}(\gamma)U_{01}(\gamma)X_{0}(\beta)X_{1}(\beta)\;Z_0Z_1\; X^\dagger_{1}(\beta)X^\dagger_{0}(\beta)U^\dagger_{01}(\gamma)U^\dagger_{02}(\gamma)U^\dagger_{12}(\gamma) | +^3 \rangle \right)$$
and
$$f_B(\gamma,\beta) = \frac{1}{2}\left(1 - \langle +^5|U_{21}(\gamma)U_{24}(\gamma)U_{23}(\gamma)U_{01}(\gamma)U_{02}(\gamma)X_{0}(\beta)X_{2}(\beta)\;Z_0Z_2\; X^\dagger_{0}(\beta)X^\dagger_{2}(\beta)U^\dagger_{02}(\gamma)U^\dagger_{01}(\gamma)U^\dagger_{12}(\gamma)U^\dagger_{23}(\gamma)U^\dagger_{24}(\gamma) | +^5 \rangle \right)$$
How complex these expectation values become in general depend only on the degree of the graph we are considering and is independent of the size of the full graph if the degree is bounded. A direct evaluation of this expression with $U_{k,l}(\gamma) = \exp\frac{i\gamma}{2}\left(1 - Z_kZ_l\right)$ and
$X_k(\beta) = \exp(i\beta X_k)$ yields
$$f_A(\gamma,\beta) = \frac{1}{2}\left(sin(4\gamma)sin(4\beta) + sin^2(2\beta)sin^2(2\gamma)\right)$$
and
$$f_B(\gamma,\beta) = \frac{1}{2}\left(1 - sin^2(2\beta)sin^2(2\gamma)cos^2(4\gamma) - \frac{1}{4}sin(4\beta)sin(4\gamma)(1+cos^2(4\gamma))\right) $$
These results can now be combined as described above, and the expectation value is therefore given by
$$ F_1(\gamma,\beta) = 3 - \left(sin^2(2\beta)sin^2(2\gamma)- \frac{1}{2}sin(4\beta)sin(4\gamma)\right)\left(1 + cos^2(4\gamma)\right),$$
We plot the function $F_1(\gamma,\beta)$ and use a simple grid search to find the parameters $(\gamma^*,\beta^*)$
that maximize the expectation value.
```
# Evaluate the function
step_size = 0.1;
a_gamma = np.arange(0, np.pi, step_size)
a_beta = np.arange(0, np.pi, step_size)
a_gamma, a_beta = np.meshgrid(a_gamma,a_beta)
F1 = 3-(np.sin(2*a_beta)**2*np.sin(2*a_gamma)**2-0.5*np.sin(4*a_beta)*np.sin(4*a_gamma))*(1+np.cos(4*a_gamma)**2)
# Grid search for the minimizing variables
result = np.where(F1 == np.amax(F1))
a = list(zip(result[0],result[1]))[0]
gamma = a[0]*step_size;
beta = a[1]*step_size;
# Plot the expetation value F1
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_surface(a_gamma, a_beta, F1, cmap=cm.coolwarm, linewidth=0, antialiased=True)
ax.set_zlim(1,4)
ax.zaxis.set_major_locator(LinearLocator(3))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
plt.show()
#The smallest parameters and the expectation can be extracted
print('\n --- OPTIMAL PARAMETERS --- \n')
print('The maximal expectation value is: M1 = %.03f' % np.amax(F1))
print('This is attained for gamma = %.03f and beta = %.03f' % (gamma,beta))
```
### 5.3 Quantum circuit<a id="implementation_sec53"></a>
With these parameters we can now construct the circuit that prepares the trial state for the Graph
or the Graph $G = (V,E)$ described above with vertex set $V = \{0,1,2,3,4\}$ and the edges are $E = \{(0,1),(0,2),(1,2),(3,2),(3,4),(4,2)\}$. The circuit is going to require $n = 5$ qubits and we prepare the state
$$ |\psi_1(\gamma ,\beta)\rangle = e^{ -i\beta B } e^{ -i\gamma H } |+\rangle^n. $$
Recall that the terms are given by $B = \sum_{k \in V} X_k$ and $H = \sum_{(k,m) \in E} \frac{1}{2}\left(1 - Z_kZ_m\right)$. To generate the circuit we follow these steps:
- We first implement 5 Hadamard $H$ gates to generate the uniform superposition.
- This is follow by $6$ Ising type gates $U_{k,l}(\gamma)$ with angle $\gamma$ along the edges $(k,l) \in E$. This gate can be expressed in terms of the native Qiskit gates as
$$ U_{k,l}(\gamma) = C_{u1}(-2\gamma)_{k,l}u1(\gamma)_k u1(\gamma)_l$$
- Lastly we apply single qubit $X$ rotations $X_k(\beta)$ for every vertex $k \in V$ with $\beta$ as angle. This gate directly parametrized as $X_k(\beta) = R_x(2\beta)_k$ in Qiskit.
- In the last step we measure the qubits in the computational basis, i.e. we perform a $Z$ measurement and record the resulting bit-string $x \in \{0,1\}^5$.
```
# prepare the quantum and classical resisters
QAOA = QuantumCircuit(len(V), len(V))
# apply the layer of Hadamard gates to all qubits
QAOA.h(range(len(V)))
QAOA.barrier()
# apply the Ising type gates with angle gamma along the edges in E
for edge in E:
k = edge[0]
l = edge[1]
QAOA.cp(-2*gamma, k, l)
QAOA.p(gamma, k)
QAOA.p(gamma, l)
# then apply the single qubit X rotations with angle beta to all qubits
QAOA.barrier()
QAOA.rx(2*beta, range(len(V)))
# Finally measure the result in the computational basis
QAOA.barrier()
QAOA.measure(range(len(V)),range(len(V)))
### draw the circuit for comparison
QAOA.draw()
```
### 5.4 Cost function evaluation<a id="implementation_sec54"></a>
Finally, we need a routine to compute the cost function value from the bit string.
This is necessary to decide whether we have found a "good candidate" bit string $x$ but could also
be used to estimate the expectation value $F_1(\gamma,\beta)$ in settings where the expectation value can not be evaluated directly.
```
# Compute the value of the cost function
def cost_function_C(x,G):
E = G.edges()
if( len(x) != len(G.nodes())):
return np.nan
C = 0;
for index in E:
e1 = index[0]
e2 = index[1]
w = G[e1][e2]['weight']
C = C + w*x[e1]*(1-x[e2]) + w*x[e2]*(1-x[e1])
return C
```
## 5a. Running QAOA on a simulator<a id="implementationsim"></a>
We first run the algorithm on a local QASM simulator.
```
# run on local simulator
backend = Aer.get_backend("aer_simulator")
shots = 10000
TQAOA = transpile(QAOA, backend)
qobj = assemble(TQAOA, shots=shots)
QAOA_results = backend.run(qobj).result()
plot_histogram(QAOA_results.get_counts(),figsize = (8,6),bar_labels = False)
```
#### Evaluate the data from the simulation
Let us now proceed to calculate the relevant information from the simulated data. We will use the
obtained results to
- Compute the mean energy and check whether it agrees with the theoretical prediction
- Report the sampled bit string $x^*$ with the largest observed cost function $C(x^*)$
- Plot the Histogram of the energies to see whether it indeed concentrates around the predicted mean
```
# Evaluate the data from the simulator
counts = QAOA_results.get_counts()
avr_C = 0
max_C = [0,0]
hist = {}
for k in range(len(G.edges())+1):
hist[str(k)] = hist.get(str(k),0)
for sample in list(counts.keys()):
# use sampled bit string x to compute C(x)
x = [int(num) for num in list(sample)]
tmp_eng = cost_function_C(x,G)
# compute the expectation value and energy distribution
avr_C = avr_C + counts[sample]*tmp_eng
hist[str(round(tmp_eng))] = hist.get(str(round(tmp_eng)),0) + counts[sample]
# save best bit string
if( max_C[1] < tmp_eng):
max_C[0] = sample
max_C[1] = tmp_eng
M1_sampled = avr_C/shots
print('\n --- SIMULATION RESULTS ---\n')
print('The sampled mean value is M1_sampled = %.02f while the true value is M1 = %.02f \n' % (M1_sampled,np.amax(F1)))
print('The approximate solution is x* = %s with C(x*) = %d \n' % (max_C[0],max_C[1]))
print('The cost function is distributed as: \n')
plot_histogram(hist,figsize = (8,6),bar_labels = False)
```
## 5b. Running QAOA on a real quantum device<a id="implementationdev"></a>
We then see how the same circuit can be executed on real-device backends.
```
provider = IBMQ.load_account()
backend = provider.get_backend('ibmq_santiago')
shots = 2048
TQAOA = transpile(QAOA, backend)
job_exp = backend.run(TQAOA, shots=shots)
job_monitor(job_exp)
exp_results = job_exp.result()
plot_histogram(exp_results.get_counts(),figsize = (10,8),bar_labels = False)
```
#### Evaluate the data from the experiment
We can now repeat the same analysis as before and compare the experimental result.
```
# Evaluate the data from the experiment
counts = exp_results.get_counts()
avr_C = 0
max_C = [0,0]
hist = {}
for k in range(len(G.edges())+1):
hist[str(k)] = hist.get(str(k),0)
for sample in list(counts.keys()):
# use sampled bit string x to compute C(x)
x = [int(num) for num in list(sample)]
tmp_eng = cost_function_C(x,G)
# compute the expectation value and energy distribution
avr_C = avr_C + counts[sample]*tmp_eng
hist[str(round(tmp_eng))] = hist.get(str(round(tmp_eng)),0) + counts[sample]
# save best bit string
if( max_C[1] < tmp_eng):
max_C[0] = sample
max_C[1] = tmp_eng
M1_sampled = avr_C/shots
print('\n --- EXPERIMENTAL RESULTS ---\n')
print('The sampled mean value is M1_sampled = %.02f while the true value is M1 = %.02f \n' % (M1_sampled,np.amax(F1)))
print('The approximate solution is x* = %s with C(x*) = %d \n' % (max_C[0],max_C[1]))
print('The cost function is distributed as: \n')
plot_histogram(hist,figsize = (8,6),bar_labels = False)
```
## 6. Problems<a id="problems"></a>
0. The QAOA algorithm produces a bit string, is this string the optimal solution for this graph? Compare the experimental results from the superconducting chip with the results from the local QASM simulation.
1. We have computed the cost function $F_1$ analytically in [section 5.2](#implementation_sec52). Verify the steps and compute $f_A(\gamma,\beta)$ as well $f_B(\gamma,\beta)$.
2. We have given an exact expression for $F_1$ in the Qiskit implementation.
-Write a routine to estimate the expectation value $F_1(\gamma,\beta)$ from the samples obtained in the result (hint: use the function cost_function_C(x,G) from [section 5.4](#implementation_sec54) and the evaluation of the data in both section [5.a / 5.b](#implementationsim))
-Use an optimization routine,e.g. SPSA from the VQE example in this tutorial, to optimize the parameters in the sampled $F_1(\gamma,\beta)$ numerically. Do you find the same values for $\gamma^*,\beta^*$ ?
3. The Trial circuit in [section 5.3](#implementation_sec53) corresponds to depth $p=1$ and was directly aimed at being compatible with the Hardware.
-Use the routine from exercise 2 to evaluate the cost functions $F_p(\gamma,\beta)$ for $p=2,3$. What do you expect to see in the actual Hardware?
-Generalize this class of trial state to other candidate wave functions, such as the Hardware efficient ansatz of Ref. [4](#references).
4. Consider an example of $MAX \;\; 3-SAT$ as discussed in the example section and modify the function cost_function_C(c,G) from [section 5.4](#implementation_sec54) you have used to compute $F_p$ accordingly. Run the QAOA algorithm for this instance of $MAX \; 3-SAT$ using the hardware efficient algorithm and analyze the results.
## 7. References<a id="references"></a>
1. Garey, Michael R.; David S. Johnson (1979). Computers and Intractability: A Guide to the Theory of NP-Completeness. W. H. Freeman. ISBN 0-7167-1045-5
2. Goemans, Michel X., and David P. Williamson. [Journal of the ACM (JACM) 42.6 (1995): 1115-1145](http://www-math.mit.edu/~goemans/PAPERS/maxcut-jacm.pdf).
3. Farhi, Edward, Jeffrey Goldstone, and Sam Gutmann. "A quantum approximate optimization algorithm." arXiv preprint [arXiv:1411.4028 (2014)](https://arxiv.org/abs/1411.4028).
4. Kandala, Abhinav, et al. "Hardware-efficient variational quantum eigensolver for small molecules and quantum magnets." [Nature 549.7671 (2017): 242](https://www.nature.com/articles/nature23879).
5. Farhi, Edward, et al. "Quantum algorithms for fixed qubit architectures." arXiv preprint [arXiv:1703.06199 (2017)](https://arxiv.org/abs/1703.06199).
6. Spall, J. C. (1992), [IEEE Transactions on Automatic Control, vol. 37(3), pp. 332–341](https://ieeexplore.ieee.org/document/119632).
7. Michael Streif and Martin Leib "Training the quantum approximate optimization algorithm without access to a quantum processing unit" (2020) [Quantum Sci. Technol. 5 034008](https://doi.org/10.1088/2058-9565/ab8c2b)
```
import qiskit.tools.jupyter
%qiskit_version_table
```
| github_jupyter |
```
%pylab inline
from galaxy2galaxy import problems
from astropy.visualization import make_lupton_rgb
import tensorflow as tf
Modes = tf.estimator.ModeKeys
# To list the problems currently available in G2G
problems.available()
# Let's create an instance of the hsc_problem
problem128 = problems.problem('attrs2img_cosmos_psf_euclide')
def wiener_tf(image, psf, balance):
trans_func = tf.signal.rfft2d(tf.signal.ifftshift(psf))
wiener_filter = tf.math.conj(trans_func) / (tf.dtypes.cast(tf.math.abs(trans_func),'complex64') ** 2 + balance)
deconv = tf.signal.irfft2d(wiener_filter * tf.signal.rfft2d(image),fft_length=image.shape)
deconv = tf.keras.backend.clip(deconv, -1, 1)
return deconv
def tikhonov_tf(img, psf, tau):
if len(img.shape) == 2:
return(wiener_tf(img, psf, tau))
tikho_list = []
for i in range(img.shape[0]):
deconvolved = wiener_tf(img[i], psf[i], tau[i])
tikho_list += [deconvolved]
return tikho_list
dset = problem128.dataset(Modes.TRAIN, data_dir='/sparseastro/Euclid/ShapeDeconv/attrs2img_cosmos_psf_euclide')
dset2 = problem128.dataset(Modes.TRAIN, data_dir='/sparseastro/Euclid/ShapeDeconv/attrs2img_cosmos_psf_euclide')
#from tensorflow.compat.v1.spectral import irfft2d
def pre_proc_unet(dico):
print(dico['inputs'].shape)
print(dico['psf'].shape)
dico['psf'] = tf.signal.irfft2d(tf.cast(dico['psf'], complex64)[...,0])
print(dico['psf'].shape)
dico['psf'] = tf.signal.fftshift(dico['psf'])
dico['inputs'] = wiener_tf(tf.compat.v1.image.resize_image_with_pad(dico['inputs'], 128, 128)[...,0], dico['psf'], 0.05)
dico['psf'] = tf.expand_dims(dico['psf'], axis=-1)
dico['inputs'] = tf.expand_dims(dico['inputs'], axis=-1)
dico['inputs'] = tf.image.crop_to_bounding_box(dico['inputs'],33,33, 64,64)
print(dico['psf'].shape)
print(dico['inputs'].shape)
return dico
dset = dset.map(pre_proc_unet)
dset2 = dset2.batch(1)
dset = dset.batch(1)
dset
# Build an iterator over this dataset, and extract a batch
it = dset.make_one_shot_iterator().get_next()
it2 = dset2.make_one_shot_iterator().get_next()
sess = tf.Session()
batch = sess.run(it)
batch2 = sess.run(it2)
type(it['inputs'])
#it.keys()
figure(figsize=(10,10))
for i in range(1):
subplot(141)
imshow(batch2['inputs'][i])
subplot(142)
imshow(batch['inputs2'][i])
subplot(143)
imshow(batch['inputs'][i])
subplot(144)
imshow(batch['psf'][i])
axis('off')
figure(figsize=(10,10))
imshow(batch['inputs'][0])
```
## Test de raccordement avec le UNet
```
def DenseBlock(n_layers, n_kernels, input_layer, activation_function='swish',
axis_concat=3, concat_input=True,dilation_rate=1):
#concat_input: implies that we have a skip concatenation between input and output of block
connect_input = input_layer
for n in range(n_layers):
bn = tf.keras.layers.BatchNormalization()(connect_input)
act = tf.keras.layers.Activation(activation_function)(bn)
conv = tf.keras.layers.SeparableConv2D(n_kernels, 3, padding='same', use_bias=False,
kernel_initializer='he_uniform')(act)
if n == 0:
concat = conv
else:
concat = tf.keras.layers.Concatenate(axis = axis_concat)([concat, conv])
connect_input = concat
if concat_input:
return tf.keras.layers.Concatenate(axis = axis_concat)([input_layer, concat])
return concat
# Define the modified U-net
img_rows = img_cols = 64
nb_scales = 4
growth_rate = 12
nb_layers_per_block = [4,5,6,7]
activation_function= 'relu'
gamma=1
inputs = tf.keras.Input(shape=(64,64,1), name="inputs")
targets = tf.keras.Input(shape=(64,64,1), name="inputs2")
#INPUT CONV
net = tf.keras.layers.Conv2D(32, 3, padding='same', use_bias=False, kernel_initializer='he_uniform')(inputs)
#CONTRACTING PATH
skip_connect = []
for scale in range(nb_scales-1):
block = DenseBlock(n_layers=nb_layers_per_block[scale],
n_kernels=growth_rate,
input_layer=net,
activation_function=activation_function,
concat_input=True)
skip_connect.append(block)
batch_norm = tf.keras.layers.BatchNormalization()(block)
activation = tf.keras.layers.Activation(activation_function)(batch_norm)
conv_transi = tf.keras.layers.Conv2D(32+np.sum(nb_layers_per_block[:scale+1])*growth_rate, 1,
padding='same', use_bias=False, kernel_initializer='he_uniform')(activation)
net = tf.keras.layers.AveragePooling2D(pool_size=(2, 2), strides=(2, 2), padding='same')(conv_transi)
#BOTTLENECK
block = DenseBlock(n_layers=nb_layers_per_block[-1],
n_kernels=growth_rate,
input_layer=net,
activation_function=activation_function,
concat_input=False)
#EXPANSIVE PATH
for scale in range(nb_scales-2, -1, -1):
up = tf.keras.layers.Conv2D(nb_layers_per_block[scale+1]*growth_rate, 2, activation = activation_function,
padding = 'same', kernel_initializer = 'he_normal')(tf.keras.layers.UpSampling2D(size = (2,2))(block))
net = tf.keras.layers.Concatenate(axis = 3)([up, skip_connect[scale]])
block = DenseBlock(n_layers=nb_layers_per_block[scale],
n_kernels=growth_rate,
input_layer=net,
activation_function=activation_function,
concat_input=False)
#FUSION AND SKIP CONNECT
print("chat")
batch_norm = tf.keras.layers.BatchNormalization()(block)
activation = tf.keras.layers.Activation(activation_function)(batch_norm)
outputs = tf.keras.layers.Conv2D(1, 1, activation = 'linear')(activation)
print("chien")
model = tf.keras.Model(inputs=[inputs, targets], outputs=outputs)
model.compile(optimizer = tf.keras.optimizers.Adam(lr=1e-3), loss = 'mse', metrics=['accuracy'])
print("ours")
history = model.fit(dset, steps_per_epoch=1,epochs=1, max_queue_size=1, shuffle= False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/keivanipchihagh/Intro_To_MachineLearning/blob/master/Models/Movie_Classification_with_IMDB.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Movie Classification with IMDB
##### Imports
```
import numpy as np # numpy
import pandas as pd # Pandas
from keras.datasets import imdb # IMDB Dataset
from tensorflow import keras # Keras
from matplotlib import pyplot as plt # Matplotlib
```
##### Loading Data
```
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words = 10000) # Load the top 10000 frequently data into numpy arrays
print('Training data [0]:', train_data[0])
print('Training Label [0]:', train_labels[0])
print('Legnth:', len(train_data))
```
##### Get the feel of the data
```
def decode(index): # Decoding the sequential integers into the corresponding words
word_index = imdb.get_word_index()
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
decoded_review = ' '.join([reverse_word_index.get(i - 3, '?') for i in train_data[index]])
return decoded_review
print('Training data [0]:', decode(0))
```
##### Data Prep (One-Hot Encoding)
```
def vectorize_sequences(sequences, dimension = 10000): # Encoding the integer sequences into a binary matrix
results = np.zeros((len(sequences), dimension)) # Creating an all-zero matrix
for i, sequence in enumerate(sequences):
results[i, sequence] = 1.
return results
train_data = vectorize_sequences(train_data)
test_data = vectorize_sequences(test_data)
train_labels = np.asarray(train_labels).astype('float32')
test_labels = np.asarray(test_labels).astype('float32')
print("Vectorized training data: ", train_data, sep = '\n');
```
##### Building the model
```
model = keras.models.Sequential()
model.add(keras.layers.Dense(units = 16, activation = 'relu', input_shape = (10000,)))
model.add(keras.layers.Dense(units = 16, activation = 'relu'))
model.add(keras.layers.Dense(1, activation = 'sigmoid'))
model.compile(optimizer = keras.optimizers.RMSprop(0.001), loss = keras.losses.binary_crossentropy, metrics = [keras.metrics.binary_accuracy, 'acc'])
model.summary()
```
##### Training the moel
```
x_val = train_data[:10000]
train_data = train_data[10000:]
y_val = train_labels[:10000]
train_labels = train_labels[10000:]
history = model.fit(train_data, train_labels, batch_size = 512, epochs = 5, verbose = False, validation_data = (x_val, y_val))
```
##### Evalucating The Model
```
test_loss, test_binary_acc, test_acc = model.evaluate(test_data, test_labels)
print('Loss:', test_loss)
print('Binary Accuracy:', test_binary_acc * 100)
print('Accuracy:', test_acc * 100)
```
##### Statistics
```
# Draw the statistice for the training & validation process
epochs = range(1, len(history.history['acc']) + 1)
plt.plot(epochs, history.history['loss'], 'b', label = 'Training loss')
plt.plot(epochs, history.history['val_loss'], 'r', label = 'Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
plt.clf()
plt.plot(epochs, history.history['acc'], 'b', label='Training acc')
plt.plot(epochs, history.history['val_acc'], 'r', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
```
| github_jupyter |
# The Autodiff Cookbook
*alexbw@, mattjj@*
JAX has a pretty general automatic differentiation system. In this notebook, we'll go through a whole bunch of neat autodiff ideas that you can cherry pick for your own work, starting with the basics.
```
import jax.numpy as np
from jax import grad, jit, vmap
from jax import random
key = random.PRNGKey(0)
```
## Gradients
### Starting with `grad`
You can differentiate a function with `grad`:
```
grad_tanh = grad(np.tanh)
print(grad_tanh(2.0))
```
`grad` takes a function and returns a function. If you have a Python function `f` that evaluates the mathematical function $f$, then `grad(f)` is a Python function that evaluates the mathematical function $\nabla f$. That means `grad(f)(x)` represents the value $\nabla f(x)$.
Since `grad` operates on functions, you can apply it to its own output to differentiate as many times as you like:
```
print(grad(grad(np.tanh))(2.0))
print(grad(grad(grad(np.tanh)))(2.0))
```
Let's look at computing gradients with `grad` in a linear logistic regression model. First, the setup:
```
def sigmoid(x):
return 0.5 * (np.tanh(x / 2) + 1)
# Outputs probability of a label being true.
def predict(W, b, inputs):
return sigmoid(np.dot(inputs, W) + b)
# Build a toy dataset.
inputs = np.array([[0.52, 1.12, 0.77],
[0.88, -1.08, 0.15],
[0.52, 0.06, -1.30],
[0.74, -2.49, 1.39]])
targets = np.array([True, True, False, True])
# Training loss is the negative log-likelihood of the training examples.
def loss(W, b):
preds = predict(W, b, inputs)
label_probs = preds * targets + (1 - preds) * (1 - targets)
return -np.sum(np.log(label_probs))
# Initialize random model coefficients
key, W_key, b_key = random.split(key, 3)
W = random.normal(W_key, (3,))
b = random.normal(b_key, ())
```
Use the `grad` function with its `argnums` argument to differentiate a function with respect to positional arguments.
```
# Differentiate `loss` with respect to the first positional argument:
W_grad = grad(loss, argnums=0)(W, b)
print('W_grad', W_grad)
# Since argnums=0 is the default, this does the same thing:
W_grad = grad(loss)(W, b)
print('W_grad', W_grad)
# But we can choose different values too, and drop the keyword:
b_grad = grad(loss, 1)(W, b)
print('b_grad', b_grad)
# Including tuple values
W_grad, b_grad = grad(loss, (0, 1))(W, b)
print('W_grad', W_grad)
print('b_grad', b_grad)
```
This `grad` API has a direct correspondence to the excellent notation in Spivak's classic *Calculus on Manifolds* (1965), also used in Sussman and Wisdom's [*Structure and Interpretation of Classical Mechanics*](http://mitpress.mit.edu/sites/default/files/titles/content/sicm_edition_2/book.html) (2015) and their [*Functional Differential Geometry*](https://mitpress.mit.edu/books/functional-differential-geometry) (2013). Both books are open-access. See in particular the "Prologue" section of *Functional Differential Geometry* for a defense of this notation.
Essentially, when using the `argnums` argument, if `f` is a Python function for evaluating the mathematical function $f$, then the Python expression `grad(f, i)` evaluates to a Python function for evaluating $\partial_i f$.
### Differentiating with respect to nested lists, tuples, and dicts
Differentiating with respect to standard Python containers just works, so use tuples, lists, and dicts (and arbitrary nesting) however you like.
```
def loss2(params_dict):
preds = predict(params_dict['W'], params_dict['b'], inputs)
label_probs = preds * targets + (1 - preds) * (1 - targets)
return -np.sum(np.log(label_probs))
print(grad(loss2)({'W': W, 'b': b}))
```
You can [register your own container types](https://github.com/google/jax/issues/446#issuecomment-467105048) to work with not just `grad` but all the JAX transformations (`jit`, `vmap`, etc.).
### Evaluate a function and its gradient using `value_and_grad`
Another convenient function is `value_and_grad` for efficiently computing both a function's value as well as its gradient's value:
```
from jax import value_and_grad
loss_value, Wb_grad = value_and_grad(loss, (0, 1))(W, b)
print('loss value', loss_value)
print('loss value', loss(W, b))
```
### Checking against numerical differences
A great thing about derivatives is that they're straightforward to check with finite differences:
```
# Set a step size for finite differences calculations
eps = 1e-4
# Check b_grad with scalar finite differences
b_grad_numerical = (loss(W, b + eps / 2.) - loss(W, b - eps / 2.)) / eps
print('b_grad_numerical', b_grad_numerical)
print('b_grad_autodiff', grad(loss, 1)(W, b))
# Check W_grad with finite differences in a random direction
key, subkey = random.split(key)
vec = random.normal(subkey, W.shape)
unitvec = vec / np.sqrt(np.vdot(vec, vec))
W_grad_numerical = (loss(W + eps / 2. * unitvec, b) - loss(W - eps / 2. * unitvec, b)) / eps
print('W_dirderiv_numerical', W_grad_numerical)
print('W_dirderiv_autodiff', np.vdot(grad(loss)(W, b), unitvec))
```
JAX provides a simple convenience function that does essentially the same thing, but checks up to any order of differentiation that you like:
```
from jax.test_util import check_grads
check_grads(loss, (W, b), order=2) # check up to 2nd order derivatives
```
### Hessian-vector products with `grad`-of-`grad`
One thing we can do with higher-order `grad` is build a Hessian-vector product function. (Later on we'll write an even more efficient implementation that mixes both forward- and reverse-mode, but this one will use pure reverse-mode.)
A Hessian-vector product function can be useful in a [truncated Newton Conjugate-Gradient algorithm](https://en.wikipedia.org/wiki/Truncated_Newton_method) for minimizing smooth convex functions, or for studying the curvature of neural network training objectives (e.g. [1](https://arxiv.org/abs/1406.2572), [2](https://arxiv.org/abs/1811.07062), [3](https://arxiv.org/abs/1706.04454), [4](https://arxiv.org/abs/1802.03451)).
For a scalar-valued function $f : \mathbb{R}^n \to \mathbb{R}$, the Hessian at a point $x \in \mathbb{R}^n$ is written as $\partial^2 f(x)$. A Hessian-vector product function is then able to evaluate
$\qquad v \mapsto \partial^2 f(x) \cdot v$
for any $v \in \mathbb{R}^n$.
The trick is not to instantiate the full Hessian matrix: if $n$ is large, perhaps in the millions or billions in the context of neural networks, then that might be impossible to store.
Luckily, `grad` already gives us a way to write an efficient Hessian-vector product function. We just have to use the identity
$\qquad \partial^2 f (x) v = \partial [x \mapsto \partial f(x) \cdot v] = \partial g(x)$,
where $g(x) = \partial f(x) \cdot v$ is a new scalar-valued function that dots the gradient of $f$ at $x$ with the vector $v$. Notice that we're only ever differentiating scalar-valued functions of vector-valued arguments, which is exactly where we know `grad` is efficient.
In JAX code, we can just write this:
```
def hvp(f, x, v):
return grad(lambda x: np.vdot(grad(f)(x), v))
```
This example shows that you can freely use lexical closure, and JAX will never get perturbed or confused.
We'll check this implementation a few cells down, once we see how to compute dense Hessian matrices. We'll also write an even better version that uses both forward-mode and reverse-mode.
### Jacobians and Hessians using `jacfwd` and `jacrev`
You can compute full Jacobian matrices using the `jacfwd` and `jacrev` functions:
```
from jax import jacfwd, jacrev
# Isolate the function from the weight matrix to the predictions
f = lambda W: predict(W, b, inputs)
J = jacfwd(f)(W)
print("jacfwd result, with shape", J.shape)
print(J)
J = jacrev(f)(W)
print("jacrev result, with shape", J.shape)
print(J)
```
These two functions compute the same values (up to machine numerics), but differ in their implementation: `jacfwd` uses forward-mode automatic differentiation, which is more efficient for "tall" Jacobian matrices, while `jacrev` uses reverse-mode, which is more efficient for "wide" Jacobian matrices. For matrices that are near-square, `jacfwd` probably has an edge over `jacrev`.
You can also use `jacfwd` and `jacrev` with container types:
```
def predict_dict(params, inputs):
return predict(params['W'], params['b'], inputs)
J_dict = jacrev(predict_dict)({'W': W, 'b': b}, inputs)
for k, v in J_dict.items():
print("Jacobian from {} to logits is".format(k))
print(v)
```
For more details on forward- and reverse-mode, as well as how to implement `jacfwd` and `jacrev` as efficiently as possible, read on!
Using a composition of two of these functions gives us a way to compute dense Hessian matrices:
```
def hessian(f):
return jacfwd(jacrev(f))
H = hessian(f)(W)
print("hessian, with shape", H.shape)
print(H)
```
This shape makes sense: if we start with a function $f : \mathbb{R}^n \to \mathbb{R}^m$, then at a point $x \in \mathbb{R}^n$ we expect to get the shapes
* $f(x) \in \mathbb{R}^m$, the value of $f$ at $x$,
* $\partial f(x) \in \mathbb{R}^{m \times n}$, the Jacobian matrix at $x$,
* $\partial^2 f(x) \in \mathbb{R}^{m \times n \times n}$, the Hessian at $x$,
and so on.
To implement `hessian`, we could have used `jacrev(jacrev(f))` or `jacrev(jacfwd(f))` or any other composition of the two. But forward-over-reverse is typically the most efficient. That's because in the inner Jacobian computation we're often differentiating a function wide Jacobian (maybe like a loss function $f : \mathbb{R}^n \to \mathbb{R}$), while in the outer Jacobian computation we're differentiating a function with a square Jacobian (since $\nabla f : \mathbb{R}^n \to \mathbb{R}^n$), which is where forward-mode wins out.
## How it's made: two foundational autodiff functions
### Jacobian-Vector products (JVPs, aka forward-mode autodiff)
JAX includes efficient and general implementations of both forward- and reverse-mode automatic differentiation. The familiar `grad` function is built on reverse-mode, but to explain the difference in the two modes, and when each can be useful, we need a bit of math background.
#### JVPs in math
Mathematically, given a function $f : \mathbb{R}^n \to \mathbb{R}^m$, the Jacobian matrix of $f$ evaluated at an input point $x \in \mathbb{R}^n$, denoted $\partial f(x)$, is often thought of as a matrix in $\mathbb{R}^m \times \mathbb{R}^n$:
$\qquad \partial f(x) \in \mathbb{R}^{m \times n}$.
But we can also think of $\partial f(x)$ as a linear map, which maps the tangent space of the domain of $f$ at the point $x$ (which is just another copy of $\mathbb{R}^n$) to the tangent space of the codomain of $f$ at the point $f(x)$ (a copy of $\mathbb{R}^m$):
$\qquad \partial f(x) : \mathbb{R}^n \to \mathbb{R}^m$.
This map is called the [pushforward map](https://en.wikipedia.org/wiki/Pushforward_(differential)) of $f$ at $x$. The Jacobian matrix is just the matrix for this linear map in a standard basis.
If we don't commit to one specific input point $x$, then we can think of the function $\partial f$ as first taking an input point and returning the Jacobian linear map at that input point:
$\qquad \partial f : \mathbb{R}^n \to \mathbb{R}^n \to \mathbb{R}^m$.
In particular, we can uncurry things so that given input point $x \in \mathbb{R}^n$ and a tangent vector $v \in \mathbb{R}^n$, we get back an output tangent vector in $\mathbb{R}^m$. We call that mapping, from $(x, v)$ pairs to output tangent vectors, the *Jacobian-vector product*, and write it as
$\qquad (x, v) \mapsto \partial f(x) v$
#### JVPs in JAX code
Back in Python code, JAX's `jvp` function models this transformation. Given a Python function that evaluates $f$, JAX's `jvp` is a way to get a Python function for evaluating $(x, v) \mapsto (f(x), \partial f(x) v)$.
```
from jax import jvp
# Isolate the function from the weight matrix to the predictions
f = lambda W: predict(W, b, inputs)
key, subkey = random.split(key)
v = random.normal(subkey, W.shape)
# Push forward the vector `v` along `f` evaluated at `W`
y, u = jvp(f, (W,), (v,))
```
In terms of Haskell-like type signatures, we could write
```haskell
jvp :: (a -> b) -> a -> T a -> (b, T b)
```
where we use `T a` to denote the type of the tangent space for `a`. In words, `jvp` takes as arguments a function of type `a -> b`, a value of type `a`, and a tangent vector value of type `T a`. It gives back a pair consisting of a value of type `b` and an output tangent vector of type `T b`.
The `jvp`-transformed function is evaluated much like the original function, but paired up with each primal value of type `a` it pushes along tangent values of type `T a`. For each primitive numerical operation that the original function would have applied, the `jvp`-transformed function executes a "JVP rule" for that primitive that both evaluates the primitive on the primals and applies the primitive's JVP at those primal values.
That evaluation strategy has some immediate implications about computational complexity: since we evaluate JVPs as we go, we don't need to store anything for later, and so the memory cost is independent of the depth of the computation. In addition, the FLOP cost of the `jvp`-transformed function is about 2x the cost of just evaluating the function. Put another way, for a fixed primal point $x$, we can evaluate $v \mapsto \partial f(x) \cdot v$ for about the same cost as evaluating $f$.
That memory complexity sounds pretty compelling! So why don't we see forward-mode very often in machine learning?
To answer that, first think about how you could use a JVP to build a full Jacobian matrix. If we apply a JVP to a one-hot tangent vector, it reveals one column of the Jacobian matrix, corresponding to the nonzero entry we fed in. So we can build a full Jacobian one column at a time, and to get each column costs about the same as one function evaluation. That will be efficient for functions with "tall" Jacobians, but inefficient for "wide" Jacobians.
If you're doing gradient-based optimization in machine learning, you probably want to minimize a loss function from parameters in $\mathbb{R}^n$ to a scalar loss value in $\mathbb{R}$. That means the Jacobian of this function is a very wide matrix: $\partial f(x) \in \mathbb{R}^{1 \times n}$, which we often identify with the Gradient vector $\nabla f(x) \in \mathbb{R}^n$. Building that matrix one column at a time, with each call taking a similar number of FLOPs to evaluating the original function, sure seems inefficient! In particular, for training neural networks, where $f$ is a training loss function and $n$ can be in the millions or billions, this approach just won't scale.
To do better for functions like this, we just need to use reverse-mode.
### Vector-Jacobian products (VJPs, aka reverse-mode autodiff)
Where forward-mode gives us back a function for evaluating Jacobian-vector products, which we can then use to build Jacobian matrices one column at a time, reverse-mode is a way to get back a function for evaluating vector-Jacobian products (equivalently Jacobian-transpose-vector products), which we can use to build Jacobian matrices one row at a time.
#### VJPs in math
Let's again consider a function $f : \mathbb{R}^n \to \mathbb{R}^m$.
Starting from our notation for JVPs, the notation for VJPs is pretty simple:
$\qquad (x, v) \mapsto v \partial f(x)$,
where $v$ is an element of the cotangent space of $f$ at $x$ (isomorphic to another copy of $\mathbb{R}^m$). When being rigorous, we should think of $v$ as a linear map $v : \mathbb{R}^m \to \mathbb{R}$, and when we write $v \partial f(x)$ we mean function composition $v \circ \partial f(x)$, where the types work out because $\partial f(x) : \mathbb{R}^n \to \mathbb{R}^m$. But in the common case we can identify $v$ with a vector in $\mathbb{R}^m$ and use the two almost interchageably, just like we might sometimes flip between "column vectors" and "row vectors" without much comment.
With that identification, we can alternatively think of the linear part of a VJP as the transpose (or adjoint conjugate) of the linear part of a JVP:
$\qquad (x, v) \mapsto \partial f(x)^\mathsf{T} v$.
For a given point $x$, we can write the signature as
$\qquad \partial f(x)^\mathsf{T} : \mathbb{R}^m \to \mathbb{R}^n$.
The corresponding map on cotangent spaces is often called the [pullback](https://en.wikipedia.org/wiki/Pullback_(differential_geometry))
of $f$ at $x$. The key for our purposes is that it goes from something that looks like the output of $f$ to something that looks like the input of $f$, just like we might expect from a transposed linear function.
#### VJPs in JAX code
Switching from math back to Python, the JAX function `vjp` can take a Python function for evaluating $f$ and give us back a Python function for evaluating the VJP $(x, v) \mapsto (f(x), v^\mathsf{T} \partial f(x))$.
```
from jax import vjp
# Isolate the function from the weight matrix to the predictions
f = lambda W: predict(W, b, inputs)
y, vjp_fun = vjp(f, W)
key, subkey = random.split(key)
u = random.normal(subkey, y.shape)
# Pull back the covector `u` along `f` evaluated at `W`
v = vjp_fun(u)
```
In terms of Haskell-like type signatures, we could write
```haskell
vjp :: (a -> b) -> a -> (b, CT b -> CT a)
```
where we use `CT a` to denote the type for the cotangent space for `a`. In words, `vjp` takes as arguments a function of type `a -> b` and a point of type `a`, and gives back a pair consisting of a value of type `b` and a linear map of type `CT b -> CT a`.
This is great because it lets us build Jacobian matrices one row at a time, and the FLOP cost for evaluating $(x, v) \mapsto (f(x), v^\mathsf{T} \partial f(x))$ is only about twice the cost of evaluating $f$. In particular, if we want the gradient of a function $f : \mathbb{R}^n \to \mathbb{R}$, we can do it in just one call. That's how `grad` is efficient for gradient-based optimization, even for objectives like neural network training loss functions on millions or billions of parameters.
There's a cost, though: though the FLOPs are friendly, memory scales with the depth of the computation. Also, the implementation is traditionally more complex than that of forward-mode, though JAX has some tricks up its sleeve (that's a story for a future notebook!).
For more on how reverse-mode works, see [this tutorial video from the Deep Learning Summer School in 2017](http://videolectures.net/deeplearning2017_johnson_automatic_differentiation/).
### Hessian-vector products using both forward- and reverse-mode
In a previous section, we implemented a Hessian-vector product function just using reverse-mode:
```
def hvp(f, x, v):
return grad(lambda x: np.vdot(grad(f)(x), v))
```
That's efficient, but we can do even better and save some memory by using forward-mode together with reverse-mode.
Mathematically, given a function $f : \mathbb{R}^n \to \mathbb{R}$ to differentiate, a point $x \in \mathbb{R}^n$ at which to linearize the function, and a vector $v \in \mathbb{R}^n$, the Hessian-vector product function we want is
$(x, v) \mapsto \partial^2 f(x) v$
Consider the helper function $g : \mathbb{R}^n \to \mathbb{R}^n$ defined to be the derivative (or gradient) of $f$, namely $g(x) = \partial f(x)$. All we need is its JVP, since that will give us
$(x, v) \mapsto \partial g(x) v = \partial^2 f(x) v$.
We can translate that almost directly into code:
```
from jax import jvp, grad
# forward-over-reverse
def hvp(f, primals, tangents):
return jvp(grad(f), primals, tangents)[1]
```
Even better, since we didn't have to call `np.dot` directly, this `hvp` function works with arrays of any shape and with arbitrary container types (like vectors stored as nested lists/dicts/tuples), and doesn't even have a dependence on `jax.numpy`.
Here's an example of how to use it:
```
def f(X):
return np.sum(np.tanh(X)**2)
key, subkey1, subkey2 = random.split(key, 3)
X = random.normal(subkey1, (30, 40))
V = random.normal(subkey2, (30, 40))
ans1 = hvp(f, (X,), (V,))
ans2 = np.tensordot(hessian(f)(X), V, 2)
print(np.allclose(ans1, ans2, 1e-4, 1e-4))
```
Another way you might consider writing this is using reverse-over-forward:
```
# reverse-over-forward
def hvp_revfwd(f, primals, tangents):
g = lambda primals: jvp(f, primals, tangents)[1]
return grad(g)(primals)
```
That's not quite as good, though, because forward-mode has less overhead than reverse-mode, and since the outer differentiation operator here has to differentiate a larger computation than the inner one, keeping forward-mode on the outside works best:
```
# reverse-over-reverse, only works for single arguments
def hvp_revrev(f, primals, tangents):
x, = primals
v, = tangents
return grad(lambda x: np.vdot(grad(f)(x), v))(x)
print("Forward over reverse")
%timeit -n10 -r3 hvp(f, (X,), (V,))
print("Reverse over forward")
%timeit -n10 -r3 hvp_revfwd(f, (X,), (V,))
print("Reverse over reverse")
%timeit -n10 -r3 hvp_revrev(f, (X,), (V,))
print("Naive full Hessian materialization")
%timeit -n10 -r3 np.tensordot(hessian(f)(X), V, 2)
```
## Composing VJPs, JVPs, and `vmap`
### Jacobian-Matrix and Matrix-Jacobian products
Now that we have `jvp` and `vjp` transformations that give us functions to push-forward or pull-back single vectors at a time, we can use JAX's [`vmap` transformation](https://github.com/google/jax#auto-vectorization-with-vmap) to push and pull entire bases at once. In particular, we can use that to write fast matrix-Jacobian and Jacobian-matrix products.
```
# Isolate the function from the weight matrix to the predictions
f = lambda W: predict(W, b, inputs)
# Pull back the covectors `m_i` along `f`, evaluated at `W`, for all `i`.
# First, use a list comprehension to loop over rows in the matrix M.
def loop_mjp(f, x, M):
y, vjp_fun = vjp(f, x)
return np.vstack([vjp_fun(mi) for mi in M])
# Now, use vmap to build a computation that does a single fast matrix-matrix
# multiply, rather than an outer loop over vector-matrix multiplies.
def vmap_mjp(f, x, M):
y, vjp_fun = vjp(f, x)
return vmap(vjp_fun)(M)
key = random.PRNGKey(0)
num_covecs = 128
U = random.normal(key, (num_covecs,) + y.shape)
loop_vs = loop_mjp(f, W, M=U)
print('Non-vmapped Matrix-Jacobian product')
%timeit -n10 -r3 loop_mjp(f, W, M=U)
print('\nVmapped Matrix-Jacobian product')
vmap_vs = vmap_mjp(f, W, M=U)
%timeit -n10 -r3 vmap_mjp(f, W, M=U)
assert np.allclose(loop_vs, vmap_vs), 'Vmap and non-vmapped Matrix-Jacobian Products should be identical'
def loop_jmp(f, x, M):
# jvp immediately returns the primal and tangent values as a tuple,
# so we'll compute and select the tangents in a list comprehension
return np.vstack([jvp(f, (W,), (si,))[1] for si in S])
def vmap_jmp(f, x, M):
_jvp = lambda s: jvp(f, (W,), (s,))[1]
return vmap(_jvp)(M)
num_vecs = 128
S = random.normal(key, (num_vecs,) + W.shape)
loop_vs = loop_jmp(f, W, M=S)
print('Non-vmapped Jacobian-Matrix product')
%timeit -n10 -r3 loop_jmp(f, W, M=S)
vmap_vs = vmap_jmp(f, W, M=S)
print('\nVmapped Jacobian-Matrix product')
%timeit -n10 -r3 vmap_jmp(f, W, M=S)
assert np.allclose(loop_vs, vmap_vs), 'Vmap and non-vmapped Jacobian-Matrix products should be identical'
```
### The implementation of `jacfwd` and `jacrev`
Now that we've seen fast Jacobian-matrix and matrix-Jacobian products, it's not hard to guess how to write `jacfwd` and `jacrev`. We just use the same technique to push-forward or pull-back an entire standard basis (isomorphic to an identity matrix) at once.
```
from jax import jacrev as builtin_jacrev
def our_jacrev(f):
def jacfun(x):
y, vjp_fun = vjp(f, x)
# Use vmap to do a matrix-Jacobian product.
# Here, the matrix is the Euclidean basis, so we get all
# entries in the Jacobian at once.
J, = vmap(vjp_fun, in_axes=0)(np.eye(len(y)))
return J
return jacfun
assert np.allclose(builtin_jacrev(f)(W), our_jacrev(f)(W)), 'Incorrect reverse-mode Jacobian results!'
from jax import jacfwd as builtin_jacfwd
def our_jacfwd(f):
def jacfun(x):
_jvp = lambda s: jvp(f, (x,), (s,))[1]
Jt =vmap(_jvp, in_axes=1)(np.eye(len(x)))
return np.transpose(Jt)
return jacfun
assert np.allclose(builtin_jacfwd(f)(W), our_jacfwd(f)(W)), 'Incorrect forward-mode Jacobian results!'
```
Interestingly, [Autograd](https://github.com/hips/autograd) couldn't do this. Our [implementation of reverse-mode `jacobian` in Autograd](https://github.com/HIPS/autograd/blob/96a03f44da43cd7044c61ac945c483955deba957/autograd/differential_operators.py#L60) had to pull back one vector at a time with an outer-loop `map`. Pushing one vector at a time through the computation is much less efficient than batching it all together with `vmap`.
Another thing that Autograd couldn't do is `jit`. Interestingly, no matter how much Python dynamism you use in your function to be differentiated, we could always use `jit` on the linear part of the computation. For example:
```
def f(x):
try:
if x < 3:
return 2 * x ** 3
else:
raise ValueError
except ValueError:
return np.pi * x
y, f_vjp = vjp(f, 4.)
print(jit(f_vjp)(1.))
```
## Complex numbers and differentiation
JAX is great at complex numbers and differentiation. To support both [holomorphic and non-holomorphic differentiation](https://en.wikipedia.org/wiki/Holomorphic_function), JAX follows [Autograd's convention](https://github.com/HIPS/autograd/blob/master/docs/tutorial.md#complex-numbers) for encoding complex derivatives.
Consider a complex-to-complex function $f: \mathbb{C} \to \mathbb{C}$ that we break down into its component real-to-real functions:
```
def f(z):
x, y = real(z), imag(z)
return u(x, y), v(x, y) * 1j
```
That is, we've decomposed $f(z) = u(x, y) + v(x, y) i$ where $z = x + y i$. We define `grad(f)` to correspond to
```
def grad_f(z):
x, y = real(z), imag(z)
return grad(u, 0)(x, y) + grad(u, 1)(x, y) * 1j
```
In math symbols, that means we define $\partial f(z) \triangleq \partial_0 u(x, y) + \partial_1 u(x, y)$. So we throw out $v$, ignoring the complex component function of $f$ entirely!
This convention covers three important cases:
1. If `f` evaluates a holomorphic function, then we get the usual complex derivative, since $\partial_0 u = \partial_1 v$ and $\partial_1 u = - \partial_0 v$.
2. If `f` is evaluates the real-valued loss function of a complex parameter `x`, then we get a result that we can use in gradient-based optimization by taking steps in the direction of the conjugate of `grad(f)(x)`.
3. If `f` evaluates a real-to-real function, but its implementation uses complex primitives internally (some of which must be non-holomorphic, e.g. FFTs used in convolutions) then we get the same result that an implementation that only used real primitives would have given.
By throwing away `v` entirely, this convention does not handle the case where `f` evaluates a non-holomorphic function and you want to evaluate all of $\partial_0 u$, $\partial_1 u$, $\partial_0 v$, and $\partial_1 v$ at once. But in that case the answer would have to contain four real values, and so there's no way to express it as a single complex number.
You should expect complex numbers to work everywhere in JAX. Here's differentiating through a Cholesky decomposition of a complex matrix:
```
A = np.array([[5., 2.+3j, 5j],
[2.-3j, 7., 1.+7j],
[-5j, 1.-7j, 12.]])
def f(X):
L = np.linalg.cholesky(X)
return np.sum((L - np.sin(L))**2)
grad(f, holomorphic=True)(A)
```
For primitives' JVP rules, writing the primals as $z = a + bi$ and the tangents as $t = c + di$, we define the Jacobian-vector product $t \mapsto \partial f(z) \cdot t$ as
$t \mapsto
\begin{matrix} \begin{bmatrix} 1 & 1 \end{bmatrix} \\ ~ \end{matrix}
\begin{bmatrix} \partial_0 u(a, b) & -\partial_0 v(a, b) \\ - \partial_1 u(a, b) i & \partial_1 v(a, b) i \end{bmatrix}
\begin{bmatrix} c \\ d \end{bmatrix}$.
See Chapter 4 of [Dougal's PhD thesis](https://dougalmaclaurin.com/phd-thesis.pdf) for more details.
## More advanced autodiff
In this notebook, we worked through some easy, and then progressively more complicated, applications of automatic differentiation in JAX. We hope you now feel that taking derivatives in JAX is easy and powerful.
There's a whole world of other autodiff tricks and functionality out there. Topics we didn't cover, but hope to in a "Advanced Autodiff Cookbook" include:
- Gauss-Newton Vector Products, linearizing once
- Custom VJPs and JVPs
- Efficient derivatives at fixed-points
- Estimating the trace of a Hessian using random Hessian-vector products.
- Forward-mode autodiff using only reverse-mode autodiff.
- Taking derivatives with respect to custom data types.
- Checkpointing (binomial checkpointing for efficient reverse-mode, not model snapshotting).
- Optimizing VJPs with Jacobian pre-accumulation.
| github_jupyter |
```
import panel as pn
import pyvista as pv
from pyvista import examples
pn.extension('vtk', sizing_mode="stretch_width")
```
For this example we use the pyvista library to load a dataset and generate easily a VTK scene
```
m = examples.download_st_helens().warp_by_scalar()
# default camera position
cpos = [(567000.9232163235, 5119147.423216323, 6460.423216322832),
(562835.0, 5114981.5, 2294.5),
(-0.4082482904638299, -0.40824829046381844, 0.8164965809277649)]
# pyvista plotter
pl = pv.Plotter(notebook=True);
actor = pl.add_mesh(m, smooth_shading=True, lighting=True)
pl.camera_position = cpos #set camera position
# save initial camera properties
renderer = list(pl.ren_win.GetRenderers())[0]
initial_camera = renderer.GetActiveCamera()
initial_camera_pos = {"focalPoint": initial_camera.GetFocalPoint(),
"position": initial_camera.GetPosition(),
"viewUp": initial_camera.GetViewUp()}
# Panel creation using the VTK Scene created by the plotter pyvista
orientation_widget = True
enable_keybindings = True
vtkpan = pn.panel(pl.ren_win, sizing_mode='stretch_both', orientation_widget=orientation_widget,
enable_keybindings=enable_keybindings, height=600)
vtkpan
```
WidgetBox with colorbars and actor selection
```
# Creation of a mapping between Custom name and the VTK object reference
actor_ref = ["None", actor.__this__]
actor_names = ["None", 'St Helen']
actor_opts = {k:v for k,v in zip(actor_names, actor_ref)}
options = {}
actor_selection = pn.widgets.Select(value=None, options = actor_opts , name="Actor Selection")
actor_selection
```
WidgetBoxes with general parameters of the vtk Scene (Widgets, Background, Lights,...)
```
# Scene Layout
scene_props = pn.WidgetBox()
bind_and_orient = pn.widgets.CheckBoxGroup(value=['Orientation Widget', 'Key Bindings'], options=['Orientation Widget', 'Key Bindings']) #initialisation => coherence with panel params
reset_camera = pn.widgets.Button(name='Reset Camera')
background_color = pn.widgets.ColorPicker(value=''.join(['#'] + ['{:02x}'.format(int(v*255)) for v in pl.background_color]),
name='Background Color')
[scene_props.append(w) for w in [bind_and_orient, reset_camera, background_color]]
# Light properties
light_props = pn.WidgetBox()
light_box_title = pn.widgets.StaticText(value='Light properties')
light_type = pn.widgets.Select(value='HeadLight', options=['HeadLight','SceneLight','CameraLight'])
light_intensity = pn.widgets.FloatSlider(start=0, end=1, value=1, name="Intensity")
[light_props.append(w) for w in [light_box_title, light_type, light_intensity]]
pn.Row(scene_props, light_props)
```
WidgetBox with properties of the Actors
```
#layout actor props
actor_props = pn.WidgetBox()
opacity = pn.widgets.FloatSlider(value=1, start=0, end=1, name='Opacity', disabled=True)
lighting = pn.widgets.Toggle(value=True, name='Lighting', disabled=True)
interpolation = pn.widgets.Select(value='Phong', options=['Flat','Phong'], name='Interpolation', disabled=True)
edges = pn.widgets.Toggle(value=False, name='Show Edges', disabled=True)
edges_color = pn.widgets.ColorPicker(value='#ffffff', name='Edges Color', disabled=True)
representation = pn.widgets.Select(value='Surface', options=['Points','Wireframe','Surface'], name='Representation', disabled=True)
frontface_culling = pn.widgets.Toggle(value=False, name='Frontface Culling', disabled=True)
backface_culling = pn.widgets.Toggle(value=False, name='Backface Culling', disabled=True)
ambient = pn.widgets.FloatSlider(value=0, start=-1, end=1, name='Ambient', disabled=True)
diffuse = pn.widgets.FloatSlider(value=1, start=0, end=2, name='Diffuse', disabled=True)
specular = pn.widgets.FloatSlider(value=0, start=0, end=10, name='Specular', disabled=True)
specular_power = pn.widgets.FloatSlider(value=100, start=0, end=100, name='Specular Power', disabled=True)
[actor_props.append(w) for w in [opacity, lighting, interpolation, edges, edges_color,
representation,frontface_culling,backface_culling, ambient,
diffuse, specular, specular_power]]
actor_props
```
Linking all widgets together using jslinks
```
#Linking
light_type.jslink(vtkpan, code={'value':"""
const light = target.renderer_el.getRenderer().getLights()[0]
if (source.value == 'HeadLight')
light.setLightTypeToHeadLight()
else if (source.value == 'CameraLight')
light.setLightTypeToCameraLight()
else if (source.value == 'SceneLight')
light.setLightTypeToSceneLight()
target.renderer_el.getRenderWindow().render()
"""})
light_intensity.jslink(vtkpan, code={'value':"""
const light = target.renderer_el.getRenderer().getLights()[0]
light.setIntensity(source.value)
target.renderer_el.getRenderWindow().render()
"""})
bind_and_orient.jslink(vtkpan, code = {'active':"""
target.orientation_widget = source.active.includes(0)
target.enable_keybindings = source.active.includes(1)
"""})
reset_camera.js_on_click(args={'target': vtkpan, 'initial_camera':initial_camera_pos},
code="target.camera = initial_camera");
background_color.jslink(vtkpan, code={'value':"""
const hextoarr = (color) => {return [parseInt(color.slice(1,3),16)/255, parseInt(color.slice(3,5),16)/255, parseInt(color.slice(5,7),16)/255]}
target.renderer_el.getRenderer().setBackground(hextoarr(source.color))
target.renderer_el.getRenderWindow().render()
"""});
opacity.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setOpacity(source.value)
target.renderer_el.getRenderWindow().render()
}
""")
lighting.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setLighting(source.active)
target.renderer_el.getRenderWindow().render()
}
""")
edges.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setEdgeVisibility(source.active)
target.renderer_el.getRenderWindow().render()
}
""")
interpolation.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
if(source.value=="Flat"){
actor.getProperty().setInterpolationToFlat()
}else{
actor.getProperty().setInterpolationToPhong()
}
target.renderer_el.getRenderWindow().render()
}
""")
edges_color.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const hextoarr = (color) => {return [parseInt(color.slice(1,3),16)/255, parseInt(color.slice(3,5),16)/255, parseInt(color.slice(5,7),16)/255]}
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setEdgeColor(hextoarr(source.color))
target.renderer_el.getRenderWindow().render()
}
""")
representation.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
if(source.value=="Points"){
actor.getProperty().setRepresentationToPoints()
}else if(source.value=="Wireframe"){
actor.getProperty().setRepresentationToWireframe()
}else if(source.value=="Surface"){
actor.getProperty().setRepresentationToSurface()
}
target.renderer_el.getRenderWindow().render()
}
""")
frontface_culling.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setFrontfaceCulling(source.active)
target.renderer_el.getRenderWindow().render()
}
""")
backface_culling.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setBackfaceCulling(source.active)
target.renderer_el.getRenderWindow().render()
}
""")
ambient.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setAmbient(source.value)
target.renderer_el.getRenderWindow().render()
}
""")
diffuse.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setDiffuse(source.value)
target.renderer_el.getRenderWindow().render()
}
""")
specular.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setSpecular(source.value)
target.renderer_el.getRenderWindow().render()
}
""")
specular_power.jscallback(args={"target":vtkpan, "actor_selection":actor_selection}, value="""
if (actor_selection.value!="None"){
const actor = target.getActors(actor_selection.value)[0]
actor.getProperty().setSpecularPower(source.value)
target.renderer_el.getRenderWindow().render()
}
""")
actor_selection.jslink(target=vtkpan, code = {'value' : """
if (source.value!="None"){
const actor = target.getActors(source.value)[0]
target.outline.setInputData(actor.getMapper().getInputData())
target.renderer_el.getRenderer().addActor(target.outline_actor)
//synchronize actor props and widgets values
const properties = actor.getProperty()
opacity.setv({value: properties.getOpacity()}, {silent: true})
lighting.setv({active: !!properties.getLighting()}, {silent: true})
edges.active = !!properties.getEdgeVisibility()
const actor_color = "#" + properties.getEdgeColor().map((c) => ("0" + Math.round(255*c).toString(16,2)).slice(-2)).join('')
edges_color.setv({color: actor_color}, {silent: true})
const interp_string = properties.getInterpolationAsString()
interpolation.setv({value: interp_string[0] + interp_string.slice(1).toLocaleLowerCase()}, {silent: true})
const repr_string = properties.getRepresentationAsString()
representation.setv({value: repr_string[0] + repr_string.slice(1).toLocaleLowerCase()}, {silent: true})
frontface_culling.setv({active: !!properties.getFrontfaceCulling()}, {silent: true})
backface_culling.setv({active: !!properties.getBackfaceCulling()}, {silent: true})
ambient.setv({value: properties.getAmbient()}, {silent: true})
diffuse.setv({value: properties.getDiffuse()}, {silent: true})
specular.setv({value: properties.getSpecular()}, {silent: true})
specular_power.setv({value: properties.getSpecularPower()}, {silent: true})
//enable widgets modifications
opacity.disabled = false
lighting.disabled = false
interpolation.disabled = false
edges.disabled = false
edges_color.disabled = false
representation.disabled = false
frontface_culling.disabled = false
backface_culling.disabled = false
ambient.disabled = false
diffuse.disabled = false
specular.disabled = false
specular_power.disabled = false
} else {
target.renderer_el.getRenderer().removeActor(target.outline_actor)
opacity.disabled = true
lighting.disabled = true
interpolation.disabled = true
edges.disabled = true
edges_color.disabled = true
representation.disabled = true
frontface_culling.disabled = true
backface_culling.disabled = true
ambient.disabled = true
diffuse.disabled = true
specular.disabled = true
specular_power.disabled = true
}
target.renderer_el.getRenderWindow().render()
"""}, args={"opacity":opacity, "lighting":lighting, "interpolation": interpolation, "edges": edges, "edges_color": edges_color,
"representation": representation, "frontface_culling": frontface_culling, "backface_culling": backface_culling,
"ambient": ambient, "diffuse": diffuse, "specular": specular, "specular_power": specular_power});
```
Display all together
```
settings = pn.Column(actor_selection,pn.Tabs(('Scene controller', pn.Column(scene_props, light_props)), ('Actor properties',actor_props)))
pn.Row(vtkpan, settings)
```
## App
Lets wrap it into nice template that can be served via `panel serve VTKInteractive.ipynb`
```
description="This example demonstrates the use of **VTK and pyvista** to display a *scene*"
pn.template.FastListTemplate(site="Panel", title="VTK Interactive", sidebar=[settings], main=[description, vtkpan]).servable();
```
| github_jupyter |
# Download ImageCollection
Here we download an imagecollection using the multiprocessing API
> Note: this module uses the python multiprocessing API stopping the multiprocessing API can cause the kernel to become
> buggy, due to the worker pool not being closed properly. Stopping the IPython kernel and starting it again after a short
> wait can resolve these issues
```
! pip install retry
from pathlib import Path
import sys
import ee
import geemap
from geojson import Polygon
# sys.path.append(str(Path.cwd().parent.parent.parent / "ee-packages-py"))
from eepackages.applications import bathymetry
from eepackages.utils import download_image_collection, download_image_collection_thumb
Map = geemap.Map(center=(52.97, 4.74), zoom=11)
bounds = ee.Geometry(Polygon([[
(4.574518432617185, 52.91662291147939),
(4.90548156738281, 52.91662291147939),
(4.90548156738281, 53.02331125248889),
(4.574518432617185, 53.02331125248889),
(4.574518432617185, 52.91662291147939)
]]))
scale = Map.getScale()
# scale = 30
sdb = bathymetry.Bathymetry()
Map.addLayer(bounds)
scale = Map.getScale()
# Parameters
start_date = '2016-01-01'
stop_date = '2018-01-01'
image = sdb.compute_intertidal_depth(
bounds=bounds,
start=start_date,
stop=stop_date,
scale=scale,
missions=['S2', 'L8'],
# filter: ee.Filter.dayOfYear(7*30, 9*30), # summer-only
filter_masked=False,
# filterMasked: true,
# filterMaskedFraction: 0.5,
skip_scene_boundary_fix=False,
skip_neighborhood_search=False,
neighborhood_search_parameters={"erosion": 0, "dilation": 0, "weight": 50},
bounds_buffer=0,
water_index_min=-0.05,
water_index_max=0.15,
# lowerCdfBoundary: 45,
# upperCdfBoundary: 50
) # .reproject(ee.Projection("EPSG:3857").atScale(90))
min = 0.04
max = 0.2
Map.addLayer(sdb.composite, { "min": min, "max": max }, 'red-green-blue')
Map
help(download_image_collection)
# Adding logging to see debug logs from the image downloads, some warnings will appear.
import logging
logging.basicConfig()
path = Path.cwd() / "output"
ic = sdb._raw_images.map(lambda img: img.clip(bounds))
download_image_collection(ic, out_dir=path, download_kwargs={"format": "GEO_TIFF", "scale": 30})
import os
path = Path.cwd() / "output"
downloads = os.listdir(path)
print(downloads)
!pip install rasterio
from matplotlib import pyplot
import numpy as np
import rasterio
from rasterio.merge import merge
from rasterio.plot import show
import glob
example_image_path = str(path / downloads[0])
array = rasterio.open(example_image_path).read()
print(f"shape: {array.shape}")
print(f"band names: {ee.Image(sdb._raw_images.toList(sdb._raw_images.size()).get(0)).bandNames().getInfo()}")
blue = array[2,:,:]
green = array[3,:,:]
red = array[4,:,:]
# Function to normalize the grid values
def normalize(array):
"""Normalizes numpy arrays into scale 0.0 - 1.0"""
array_min, array_max = array.min(), array.max()
return ((array - array_min)/(array_max - array_min))
redn = normalize(red)
greenn = normalize(green)
bluen = normalize(blue)
print("Normalized bands")
print(redn.min(), '-', redn.max(), 'mean:', redn.mean())
print(greenn.min(), '-', greenn.max(), 'mean:', greenn.mean())
print(bluen.min(), '-', bluen.max(), 'mean:', bluen.mean())
# Create RGB natural color composite
rgb = np.dstack((redn, greenn, bluen))
# Let's see how our color composite looks like
pyplot.imshow(rgb)
help(download_image_collection_thumb)
# Adding logging to see debug logs from the image downloads, some warnings will appear.
import logging
logging.basicConfig()
thumb_path = Path.cwd() / "out_thumb"
ic = sdb._raw_images.select("red", "green", "blue").map(lambda img: img.clip(bounds))
download_image_collection_thumb(ic, out_dir=thumb_path, download_kwargs={
"format": "PNG",
"min": [0, 0, 0],
"max": [1, 1, 1],
"scale": 30
})
import os
downloads_thumb = os.listdir(thumb_path)
print(downloads_thumb)
%pylab inline
import matplotlib.image as mpimg
img = mpimg.imread(str(thumb_path / 'ic_1497524105260.png'))
imgplot = pyplot.imshow(img)
pyplot.show()
```
| github_jupyter |
```
import pandas as pd
df_news = pd.read_csv("https://github.com/roccqqck/news_bert/raw/master/data/2015_Company.csv", encoding="utf-8")
df_news
df_news.shape
df_news['風險類型'].value_counts()
# column name rename "編號":"id", "風險類型": "label", "段落摘要": "text"
df_news = df_news.rename(columns={"編號":"id", "風險類型": "label", "段落摘要": "text"})
df_news.head(2)
# df_news['風險事件區間'] = pd.to_datetime(df_news['風險事件區間'])
df_news['text'] = df_news['text'].astype(str).str.replace('\t' , ' ').str.replace("\n", " ") # tab都換成空格
# 只取特定欄位
df_news2 = df_news.loc[:, ['id', 'label', 'text']]
df_news2.head(2)
from snownlp import SnowNLP
def sentiments_all(text):
s = SnowNLP(u""+text) # 宣告物件 s = SnowNLP(u"这个东西真心很不错呀")
text2 = s.han # snownlp 要簡體中文效果比較好
s = SnowNLP(u""+text2)
sentiments_score = s.sentiments # 得到 0 ~ 1的sentiments分數
return sentiments_score
# text rank 演算法 死出top5的關鍵字
def keywords_all(text):
s = SnowNLP(u""+text) # 宣告物件 s = SnowNLP(u"这个东西真心很不错呀")
text2 = s.han # snownlp 要簡體中文效果比較好
s = SnowNLP(u""+text2)
top_list = s.keywords(5) # 得到 top5的關鍵字
return top_list
# text rank 演算法 死出top5的關鍵字
def text_add_keywords(text):
s = SnowNLP(u""+text) # 宣告物件 s = SnowNLP(u"这个东西真心很不错呀")
text2 = s.han # snownlp 要簡體中文效果比較好
s = SnowNLP(u""+text2)
top_list = s.keywords(5) # top5的關鍵字
text2 = "".join(top_list) + text # top5的關鍵字加到文章的最前面
return text2
text = "「陽光普照」一舉拿下最佳劇情長片、最佳導演獎、最佳男主角獎、最佳男配角獎、最佳剪輯獎、觀眾票選獎。"
sentiments_all(text)
top_list = keywords_all(text) # 把文章丟進去 回傳top5的關鍵字
print(type(top_list))
top_list
text2 = "".join(top_list) + text # top5的關鍵字加到文章的最前面
text2
text_add_keywords(text)
%%time
df_news2['keywords'] = df_news2['text'].apply(keywords_all)
df_news2['text2'] = df_news2['text'].apply(text_add_keywords)
df_news2['sentiments'] = df_news2['text'].apply(sentiments_all) # 得到 0 ~ 1的分數
df_news2['sentiments'] = 2*(df_news2['sentiments'] - 0.5) # 轉成 -1 ~ 1的分數
df_news2
text = "辭舊迎新之際,你卻在遺憾2018年沒有等到能夠打動自己的黑科技手機?"
print(len(text))
text[:10] # 只取前10個字
# 文章字數 > 510了話 去尾
# 字數小於512-2 因為還有CLS SEP
def remove_510(text):
if len(text) > 510:
text = text[:510] # 只取前510個字
return text
# # 只留字數 <= 510的文章
# # 字數小於512-2 因為還有CLS SEP
# MAX_LENGTH = 510
# # df_news2 = df_news2[~(df_news2["text"].apply(lambda x : len(x)) > MAX_LENGTH)]
# df_news2 = df_news2[df_news2["text"].apply(lambda x : len(x) <= MAX_LENGTH) ] # 去掉字數>510的文章
# df_news2 = df_news2.reset_index(drop=True) # 重設index
# df_news2['id'] = df_news2.index # index 變成 id column
# df_news2
df_news2["text"] = df_news2["text"].apply(remove_510)
df_news2["text2"] = df_news2["text2"].apply(remove_510)
df_news2.head(2)
df_news2["label"].unique()
df_news2.loc[df_news2['label'] == "財務營運風險", 'label2'] = 0
df_news2.loc[df_news2['label'] == "法律風險", 'label2'] = 1
df_news2.loc[df_news2['label'] == "資安風險", 'label2'] = 2
df_news2.loc[df_news2['label'] == "勞動風險", 'label2'] = 3
df_news2.loc[df_news2['label'] == "管理風險", 'label2'] = 4
df_news2.head(2)
df_news2["label2"].unique()
df_news2["label"] = df_news2["label2"]
df_news2["label"] = df_news2["label"].astype(int)
df_news2 = df_news2.drop(columns=['label2'])
df_news2.head(2)
df_news3 = df_news2.drop(columns=['label2'])
df_news3.head(2)
# idempotence, 將處理結果另存成 tsv 供 PyTorch 使用 # header=False會沒有column名 # \t 是tab
df_news3.to_csv("data/2015_Company_textrank_sentiments.tsv", sep="\t", index=False)
df_news3 = df_news3.drop(columns=['sentiments', 'text2'])
df_news3.head(2)
# idempotence, 將處理結果另存成 tsv 供 PyTorch 使用 # header=False會沒有column名 # \t 是tab
df_news3.to_csv("data/2015_Company.tsv", sep="\t", index=False)
# # csv 用,區隔
# df_news2.to_csv("data/2015_Company.2.csv", index=False)
# 讀取tsv測試
df_news3 = pd.read_csv("data/2015_Company.tsv", sep="\t", encoding="utf-8")
df_news3
# 讀取tsv測試
df_news3 = pd.read_csv("data/2015_Company_textrank_sentiments.tsv", sep="\t", encoding="utf-8")
df_news3
```
| github_jupyter |
# Train a Simple Audio Recognition model for microcontroller use
This notebook demonstrates how to train a 20kb [Simple Audio Recognition](https://www.tensorflow.org/tutorials/sequences/audio_recognition) model for [TensorFlow Lite for Microcontrollers](https://tensorflow.org/lite/microcontrollers/overview). It will produce the same model used in the [micro_speech](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/lite/micro/examples/micro_speech) example application.
The model is designed to be used with [Google Colaboratory](https://colab.research.google.com).
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/micro_speech/train_speech_model.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/micro_speech/train_speech_model.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
The notebook runs Python scripts to train and freeze the model, and uses the TensorFlow Lite converter to convert it for use with TensorFlow Lite for Microcontrollers.
**Training is much faster using GPU acceleration.** Before you proceed, ensure you are using a GPU runtime by going to **Runtime -> Change runtime type** and selecting **GPU**. Training 18,000 iterations will take 1.5-2 hours on a GPU runtime.
## Configure training
The following `os.environ` lines can be customized to set the words that will be trained for, and the steps and learning rate of the training. The default values will result in the same model that is used in the micro_speech example. Run the cell to set the configuration:
```
import os
# A comma-delimited list of the words you want to train for.
# The options are: yes,no,up,down,left,right,on,off,stop,go
# All other words will be used to train an "unknown" category.
os.environ["WANTED_WORDS"] = "yes,no"
# The number of steps and learning rates can be specified as comma-separated
# lists to define the rate at each stage. For example,
# TRAINING_STEPS=15000,3000 and LEARNING_RATE=0.001,0.0001
# will run 18,000 training loops in total, with a rate of 0.001 for the first
# 15,000, and 0.0001 for the final 3,000.
os.environ["TRAINING_STEPS"]="15000,3000"
os.environ["LEARNING_RATE"]="0.001,0.0001"
# Calculate the total number of steps, which is used to identify the checkpoint
# file name.
total_steps = sum(map(lambda string: int(string),
os.environ["TRAINING_STEPS"].split(",")))
os.environ["TOTAL_STEPS"] = str(total_steps)
# Print the configuration to confirm it
!echo "Training these words: ${WANTED_WORDS}"
!echo "Training steps in each stage: ${TRAINING_STEPS}"
!echo "Learning rate in each stage: ${LEARNING_RATE}"
!echo "Total number of training steps: ${TOTAL_STEPS}"
```
## Install dependencies
Next, we'll install a GPU build of TensorFlow, so we can use GPU acceleration for training.
```
# Replace Colab's default TensorFlow install with an older
# build that contains the operations that are needed for training
!pip uninstall -y tensorflow tensorflow_estimator tensorboard
!pip install -q tensorflow==1.15
```
We'll also clone the TensorFlow repository, which contains the scripts that train and freeze the model.
```
# Clone the repository from GitHub
!git clone -q https://github.com/tensorflow/tensorflow
```
## Load TensorBoard
Now, set up TensorBoard so that we can graph our accuracy and loss as training proceeds.
```
# Delete any old logs from previous runs
!rm -rf /content/retrain_logs
# Load TensorBoard
%load_ext tensorboard
%tensorboard --logdir /content/retrain_logs
```
## Begin training
Next, run the following script to begin training. The script will first download the training data:
```
!python tensorflow/tensorflow/examples/speech_commands/train.py \
--model_architecture=tiny_conv --window_stride=20 --preprocess=micro \
--wanted_words=${WANTED_WORDS} --silence_percentage=25 --unknown_percentage=25 \
--quantize=1 --verbosity=WARN --how_many_training_steps=${TRAINING_STEPS} \
--learning_rate=${LEARNING_RATE} --summaries_dir=/content/retrain_logs \
--data_dir=/content/speech_dataset --train_dir=/content/speech_commands_train
```
## Freeze the graph
Once training is complete, run the following cell to freeze the graph.
```
!python tensorflow/tensorflow/examples/speech_commands/freeze.py \
--model_architecture=tiny_conv --window_stride=20 --preprocess=micro \
--wanted_words=${WANTED_WORDS} --quantize=1 --output_file=/content/tiny_conv.pb \
--start_checkpoint=/content/speech_commands_train/tiny_conv.ckpt-${TOTAL_STEPS}
```
## Convert the model
Run this cell to use the TensorFlow Lite converter to convert the frozen graph into the TensorFlow Lite format, fully quantized for use with embedded devices.
```
!toco \
--graph_def_file=/content/tiny_conv.pb --output_file=/content/tiny_conv.tflite \
--input_shapes=1,49,40,1 --input_arrays=Reshape_2 --output_arrays='labels_softmax' \
--inference_type=QUANTIZED_UINT8 --mean_values=0 --std_dev_values=9.8077
```
The following cell will print the model size, which will be under 20 kilobytes.
```
import os
model_size = os.path.getsize("/content/tiny_conv.tflite")
print("Model is %d bytes" % model_size)
```
Finally, we use xxd to transform the model into a source file that can be included in a C++ project and loaded by TensorFlow Lite for Microcontrollers.
```
# Install xxd if it is not available
!apt-get -qq install xxd
# Save the file as a C source file
!xxd -i /content/tiny_conv.tflite > /content/tiny_conv.cc
# Print the source file
!cat /content/tiny_conv.cc
```
| github_jupyter |
Task:
PFA
A person’s signature is representative of his identity. For us at the Bank, a signed document by a customer is an instruction from him for carrying out an approved transaction for him.
On on-boarding a customer we capture an image of his signature in our systems, and on receiving a signed document (Cheques, DDs, and others) from him we match the signature on the document with the one recorded in the database before proceeding with the instruction.
In the case of skilled forgeries, it becomes very difficult to verify the identity of the customer.
We want you to build a system that can help us distinguish forgeries from actual signatures. This system should be able to study signature parameters as strokes, curves, dots, dashes, writing fluidity & style, in a Writer-Independent manner and create features for identification of the signature.
The system should not use any existing APIs and should be completely self-developed.
How should it work?
The system shall work in 2 steps:
Step 1: Accept & Store Genuine Signature Image: Take actual signature scanned image of the onboarding customer and store it in a database against a unique Customer ID
Step 2: Accept & Compare Signature Images: Accept inputs of Customer ID and corresponding signature image. Compare with the signature stored in DB against the given Customer ID, and return a Confidence Match Score between the two signature images.
This is the code for training the model (Deep learning model)
For this project i used Siamese Network for creating deep learninf model,
Description is given in reference of GIT documentation, Readme
```
# Import all the necessary Library
import torchvision
import torch.utils.data as utils
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data import DataLoader,Dataset
from torch.autograd import Variable
import matplotlib.pyplot as plt
import torchvision.utils
import numpy as np
import time
import copy
from torch.optim import lr_scheduler
import os
from PIL import Image
import torch
from torch.autograd import Variable
import PIL.ImageOps
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
import pandas as pd
from pathlib import Path
from dataset import SignaturesDataset
from model import SignaturesNetwork, SiameseNetwork
from loss import ContrastiveLoss
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
if not use_cuda:
print('No GPU found. Please use a GPU to train your neural network.')
def imshow(img,text=None,should_save=False):
npimg = img.numpy()
plt.axis("off")
if text:
plt.text(75, 8, text, style='italic',fontweight='bold',
bbox={'facecolor':'white', 'alpha':0.8, 'pad':10})
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
def show_plot(iteration,loss):
plt.plot(iteration,loss)
plt.show()
genuine_path = Path('sample_Signature/genuine')
forged_path = Path('sample_Signature/forged')
path = Path('sample_Signature')
train = pd.read_csv(path/"train_final.csv")
display(train.head())
# Load the the dataset from raw image folders
#an = SignaturesDataset(train, genuine_path)
#attrs = vars(an)
#print(attrs)
signatures_dataset = SignaturesDataset(train, genuine_path, training_dir=path, transform="train")
signatures_dataset_test = SignaturesDataset(train, genuine_path, training_dir=path, transform="test")
# Viewing the sample of images and to check whether its loading properly
vis_dataloader = DataLoader(signatures_dataset,
shuffle=True,
batch_size=8)
dataiter = iter(vis_dataloader)
example_batch = next(dataiter)
concatenated = torch.cat((example_batch[0], example_batch[1]), 0)
imshow(torchvision.utils.make_grid(concatenated))
print(example_batch[2].numpy())
batch_size = 32
epochs = 40
learning_rate = 1e-4
alpha = 0.99
show_every_n_batches = 5
# Load the dataset as pytorch tensors using dataloader
train_dataloader = DataLoader(signatures_dataset,
shuffle=True,
num_workers=8,
batch_size=batch_size)
test_dataloader = DataLoader(signatures_dataset_test, num_workers=6, batch_size=1, shuffle=True)
def init_weights(m):
if type(m) == nn.Linear:
torch.nn.init.xavier_uniform(m.weight)
m.bias.data.fill_(0.01)
if type(m) == nn.Conv2d:
torch.nn.init.xavier_uniform(m.weight)
m.bias.data.fill_(0.01)
# Declare Siamese Network
net = SignaturesNetwork()
#net = SiameseNetwork()
net.apply(init_weights)
if use_cuda:
net = net.cuda()
net = torch.nn.DataParallel(net)
net.load_state_dict(torch.load("model/model_v4.pt"))
# Decalre Loss Function
margin = 2
criterion = ContrastiveLoss(margin)
# Declare Optimizer
optimizer = optim.RMSprop(net.parameters(), lr=learning_rate, alpha=alpha, eps=1e-8, weight_decay=0.0005, momentum=0.9)
#optimizer = optim.Adam(net.parameters(),lr = learning_rate )
def train():
counter = []
loss_history = []
iteration_number= 0
original_loss = np.inf
valid_loss = 0
total_loss = 0
for epoch in range(epochs):
for i, (img0, img1 , label) in enumerate(train_dataloader):
#img0, img1 , label = data
img0, img1 , label = img0.cuda(), img1.cuda() , label.cuda()
optimizer.zero_grad()
output1, output2 = net(img0, img1)
loss_contrastive = criterion(output1, output2, label)
loss_contrastive.backward()
optimizer.step()
total_loss += loss_contrastive.item() * img0.size(0)
#valid_corrects += torch.sum(preds == target.data)
iteration_number += 1
total_loss = total_loss/len(train_dataloader.dataset)
if epoch%show_every_n_batches == 0:
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(epoch, total_loss, valid_loss))
counter.append(iteration_number)
loss_history.append(total_loss)
if total_loss < original_loss:
print('Training loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(original_loss,
total_loss))
with open('model/model.pt', 'wb') as pickle_file:
torch.save(net.state_dict(), pickle_file)
original_loss = total_loss
total_loss = 0
show_plot(counter, loss_history)
return net
model = train()
#torch.save(model.state_dict(), "model/model.pt")
print("Model Saved Successfully")
model = net
model.load_state_dict(torch.load("model/model.pt"))
accuracy=0
counter=0
correct=0
for i, data in enumerate(test_dataloader,0):
x0, x1, label = data
# onehsot applies in the output of 128 dense vectors which is then converted to 2 dense vectors
output1, output2 = model(x0.to(device),x1.to(device))
res = nn.Softmax(dim=1)(output1.cuda() - output2.cuda())
#res = torch.abs(output1.cuda() - output2.cuda())
label = label[0].tolist()
label = int(label[0])
result=torch.max(res,1)[1].data[0].tolist()
if label == result:
correct = correct+1
counter=counter+1
accuracy=(correct/len(test_dataloader))*100
print("Accuracy:{}%".format(accuracy))
# Print the sample outputs to view its dissimilarity
counter=0
list_1 = torch.FloatTensor([[1]])
list_0 = torch.FloatTensor([[0]])
for i, data in enumerate(test_dataloader,0):
x0, x1, label = data
concatenated = torch.cat((x0,x1),0)
output1, output2 = model(Variable(x0).to(device),Variable(x1).to(device))
res = nn.Softmax(dim=1)(output1.cuda() - output2.cuda())
eucledian_distance_1 = F.pairwise_distance(output2, output1)
#eucledian_distance_2 = (output2 - output1).pow(2).sum(1)
result=torch.max(res,1)[1].data[0].tolist()
print(res)
#print(eucledian_distance_1.item())
#print(eucledian_distance_2)
if result==0:
pred_label="Orginial"
else:
pred_label="Forged"
if label==list_0:
label="Orginial"
else:
label="Forged"
imshow(torchvision.utils.make_grid(concatenated),'Dissimilarity: {:.2f} Label: {} predicted: {}'.format(eucledian_distance_1.item(),label, pred_label))
counter = counter + 1
if counter ==20:
break
```
| github_jupyter |
```
import keras
import keras2onnx
import onnx
# from tensorflow.keras.models import load_model
from keras.models import load_model
# import tensorflow as tf
import keras
import keras2onnx
import onnx
# from tensorflow.keras.models import load_model
from keras.models import load_model
# model = keras.applications.resnet50.ResNet50(include_top=True)
# model.summary()
# model = keras.models.Model(inputs=model.input, outputs=model.get_layer('avg_pool').output)
# model.save('./res_top.h5')
# onnx_model = keras2onnx.convert_keras(model, model.name) #, channel_first_inputs=(1,3,224,224), target_opset=11)
# onnx.save_model(onnx_model, './res_top.onnx')
# model = keras.applications.resnet50.ResNet50(include_top=False)#, input_shape=(224,224,3))
# model = keras.models.Model(inputs=model.input, outputs=model.get_layer('avg_pool').output)
# model.save('./res_notop.h5')
model = load_model('./res_notop.h5')
# model.summary()
onnx_model = keras2onnx.convert_keras(model, model.name)#, channel_first_inputs=[None, 224, 224, 3])#224,224,3))
h = onnx_model.graph.input[0].type.tensor_type.shape.dim[1]
w = onnx_model.graph.input[0].type.tensor_type.shape.dim[2]
h.dim_value = 224
w.dim_value = 224
c = onnx_model.graph.input[0].type.tensor_type.shape.dim[3]
print(type(c.dim_param))
print(c)
print(c.dim_param)
print(onnx_model.graph.input[0].type.tensor_type.shape.dim[2])
# onnx_model.graph.input[0].type.tensor_type.shape = [None,224,224,3]
onnx.save_model(onnx_model, './res_notop.onnx')
import keras
import keras2onnx
import onnx
# from tensorflow.keras.models import load_model
from keras.models import load_model
import tensorflow as tf
model = load_model('./res.h5')
onnx_model = keras2onnx.convert_keras(model, model.name)#, channel_first_inputs=[None, 224, 224, 3])#224,224,3))
h = onnx_model.graph.input[0].type.tensor_type.shape.dim[1]
w = onnx_model.graph.input[0].type.tensor_type.shape.dim[2]
h.dim_value = 1024
w.dim_value = 1024
print(w, h)
onnx.save_model(onnx_model, './res_notop.onnx')
import keras
import keras2onnx
import onnx
# from tensorflow.keras.models import load_model
from keras.models import load_model
# model = keras.applications.resnet50.ResNet50(include_top=True)
model = load_model('./res.h5')
#model = load_model('./resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5')
model.summary()
#model.load_weights("./res.h5")
# inputs = keras.models.Input(shape=(224,224,3))
# out = model.get_layer('input_image')(inputs)
# model1 = keras.models.Model(inputs=inputs, outputs=model.get_layer('activation_2').output)
# model2 = keras.models.Model(inputs=model.input, outputs=model.get_layer('bn2a_branch2a').output)
# model1.summary()
# model2.summary()
# model3 = keras.models.Model(inputs=inputs, outputs=model.get_layer('bn2a_branch2a').output)
# model3.summary()
# onnx_model = keras2onnx.convert_keras(model, model.name, channel_first_inputs=(1,3,224,224), target_opset=11)
onnx_model = keras2onnx.convert_keras(model, model.name, channel_first_inputs=(1,3,224,224), target_opset=11)
onnx.save_model(onnx_model, './res.onnx')
import onnx
# Load the ONNX model
onnx_model = onnx.load("/home/e00064/imagination/icleague_demo/networks/maskrcnn/mask_rcnn_R_50_FPN_1x.onnx")
h = onnx_model.graph.input[0].type.tensor_type.shape.dim[1]
w = onnx_model.graph.input[0].type.tensor_type.shape.dim[2]
h.dim_value = 1024
w.dim_value = 1024
# onnx_model.graph.input[0].type.tensor_type.shape = [None,224,224,3]
onnx.save_model(onnx_model, './res_notop.onnx')
import numpy as np
def clip_boxes_graph(boxes, window):
"""
boxes: [N, (y1, x1, y2, x2)]
window: [4] in the form y1, x1, y2, x2
"""
# Split
wy1, wx1, wy2, wx2 = tf.split(window, 4)
y1, x1, y2, x2 = tf.split(boxes, 4, axis=1)
# Clip
y1 = tf.maximum(tf.minimum(y1, wy2), wy1)
x1 = tf.maximum(tf.minimum(x1, wx2), wx1)
y2 = tf.maximum(tf.minimum(y2, wy2), wy1)
x2 = tf.maximum(tf.minimum(x2, wx2), wx1)
clipped = tf.concat([y1, x1, y2, x2], axis=1, name="clipped_boxes")
clipped.set_shape((clipped.shape[0], 4))
return clipped
boxes = [10,20, 100, 200]
window = [20, 30, 90, 100]
clip_boxes_graph(boxes, window)
```
| github_jupyter |
```
import os
dirpath,dirnames,filenames = next(os.walk('functions'))
dirpath
try:
%load_ext autoreload
%autoreload 2
except:
pass
%matplotlib inline
import functions.add_path
import functions.plotnine_theme
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from plotnine import *
from plotnine.data import *
import os
print(os.getcwd())
if os.getcwd()[-4:] != 'code':
try:
os.chdir('./code')
except:
os.chdir('/net/store/nbp/projects/IntoTheWild/ET_Analysis/etcomp/code')
import av # import to import before any pupillabs libraries
from lib.pupil.pupil_src.shared_modules import file_methods as pl_file_methods
from functions import et_import
#from functions.nbp_recalib import sort_pupil
import av
# Pretty serious workaround. Ignores errors in imports :S
import builtins
from types import ModuleType
class DummyModule(ModuleType):
def __getattr__(self, key):
return None
__all__ = [] # support wildcard imports
def tryimport(name, *args, **kwargs):
try:
imp = realimport(name, *args,**kwargs)
#print('success: '+name)
return imp
except Exception as e:
print('reached exception:' + name)
if name =='cPickle': # because how they import cPickle/Pickle
return realimport('pickle', *args,**kwargs)
#print(e)
return DummyModule(name)
realimport, builtins.__import__ = builtins.__import__, tryimport
try:
import lib.pupil.pupil_src.shared_modules.calibration_routines.manual_marker_calibration as mc
except Exception as e:
print('-------------------')
print(e)
pass
tryimport, builtins.__import__ = builtins.__import__, realimport
import pl_anna_tools
from functions.et_helper import tic,toc
video_file_path = '/net/store/nbp/projects/IntoTheWild/Daten/Eyetracking/Wild/VP21/2018_05_24/001'
cap = pl_anna_tools.init_playback(video_name = 'world.mp4',video_file_path = video_file_path)
pupil_data = pl_file_methods.load_object(os.path.join(video_file_path,'pupil_data'))
#pupil_data2 = pl_file_methods.load_object('/net/store/nbp/projects/IntoTheWild/Daten/Eyetracking/Wild/VP21/2018_05_24/001/offline_data/offline_pupil.pldata')
pupil_data2 = pl_file_methods.load_pldata_file(directory='/net/store/nbp/projects/IntoTheWild/Daten/Eyetracking/Wild/VP21/2018_05_24/001/offline_data/',topic='offline_pupil')
pupil_data['pupil_position'] = pupil_data2.data
world_timestamps= np.load(os.path.join(video_file_path,'world_timestamps.npy'))
ind = np.digitize([p['timestamp'] for p in pupil_data['pupil_positions']],world_timestamps)-1
class Global_Container():
pass
calib = mc.Manual_Marker_Calibration(Global_Container)
# we cant use the calib.start() because of the super-inherited function that wants to update the gui
import types
calib.active = True
calib.ref_list = []
calib.pupil_list = []
calib.button = Global_Container()
calib.g_pool.get_timestamp = lambda:None
calib.trackerid = 0
def notify_capture_complete(self,x):
if x['subject'] == 'calibration.marker_sample_completed':
self.trackerid +=1
print(self.trackerid)
for i,r in enumerate(self.ref_list):
if 'trackerid' not in r.keys():
r['trackerid'] = self.trackerid
self.ref_list[i] = r
calib.notify_all = types.MethodType(notify_capture_complete,calib)
calib.button = Global_Container()
calib.button.status_text = ''
calibrange = range(29724,37479)
calib.circle_tracker._wait_count = 0
tic()
for k in calibrange:
if np.mod(k-min(calibrange),500) == 0:
print('Progress: %.1f%%'%((float(k-min(calibrange))/(max(calibrange)-min(calibrange)))*100))
cap.seek_to_frame(k)
frame= cap.get_frame()
pupil_idx = np.where(ind == cap.get_frame_index())[0]
pupil_in_bound = [pupil_data['pupil_positions'][i] for i in pupil_idx]
# look at the current frame
# pupil_position for pupil <v1.8 , pupil for pupil >=v1.8
calib.recent_events({'frame':frame,'pupil':pupil_in_bound,'pupil_positions':pupil_in_bound})
#if calib.sample_ref_dist >0.1:
# print('marker found: %i, smooth_vel:%.3f, sample_ref_dist:%.3f'%(len(calib.markers),calib.smooth_vel,calib.sample_ref_dist))
toc()
trackerids = [r['trackerid'] for r in calib.ref_list]
# for jedes trackerid timestamp set, find pupil positions that fall into that set
from functions.et_helper import mad
pupil_ts = np.array([c['timestamp'] for c in calib.pupil_list])
ref_pupil_spread = pd.DataFrame(columns=['trackerid','mad'])
for ref_id in np.unique(trackerids):
ref_ts = [r['timestamp'] for r in calib.ref_list if r['trackerid']==ref_id]
ref_ts_min,ref_ts_max = min(ref_ts),max(ref_ts)
ix = np.where((pupil_ts>ref_ts_min) & (pupil_ts<=ref_ts_max))[0]
pos2d_1 = np.array([calib.pupil_list[i]['norm_pos'] for i in ix if calib.pupil_list[i]['id']==0 and calib.pupil_list[i]['confidence']>0.8])
pos2d_2 = np.array([calib.pupil_list[i]['norm_pos'] for i in ix if calib.pupil_list[i]['id']==1 and calib.pupil_list[i]['confidence']>0.8])
plt.figure()
if pos2d_1.size>0:
plt.plot(pos2d_1[:,0],pos2d_1[:,1],'or')
if pos2d_2.size>0:
plt.plot(pos2d_2[:,0],pos2d_2[:,1],'og')
plt.xlim([0,1])
plt.ylim([0,1])
m = np.mean(np.array([calib.pupil_list[i]['norm_pos'] for i in ix]),axis=0)
#sd2d = np.mean(np.sqrt(np.square(pos2d[:,0]-m[0]) + np.square(pos2d[:,1]-m[1])))
#ref_pupil_spread = ref_pupil_spread.append(pd.DataFrame([[ref_id,sd2d]],columns=['trackerid','mad']))
ref_pupil_spread
pos2d = np.array([calib.pupil_list[i]['norm_pos'] for i in ix])
m = np.mean(np.array([calib.pupil_list[i]['norm_pos'] for i in ix]),axis=0)
mad2d = np.mean(np.sqrt(np.square(pos2d[:,0]-m[0]) + np.square(pos2d[:,1]-m[1])))
ref_x = [p['norm_pos'][0] for p in calib.ref_list]
ref_y = [p['norm_pos'][1] for p in calib.ref_list]
pup_x = [p['norm_pos'][0] for p in calib.pupil_list]
pup_y = [p['norm_pos'][1] for p in calib.pupil_list]
%matplotlib inline
#plt.plot(np.array(pup_x),np.array(pup_y),'ob')
plt.plot(ref_x,ref_y,'sr')
plt.xlim([0,1])
plt.ylim([0,1])
plt.figure
plt.plot([p['timestamp'] for p in calib.ref_list])
640/frame.gray.shape[0]
calib.markers
import lib.pupil.pupil_src.shared_modules.circle_detector as cd
cd.find_pupil_circle_marker(frame.gray,0.5)
```
| github_jupyter |
# Day 3: Using Pandas Data Frames to analyze single cell electrophysiology data
<img src="http://www.zocalopublicsquare.org/wp-content/uploads/2016/12/Mathews-on-US-China.jpg" width="300" height="300" />
Today we will analyze patch-clamp data from a single PV+ neuron in a mouse cortical slice. The experiment was performed in current clamp. The experimenter injected 20 different square pulses of current and recorded the voltage response of the neuron to each pulse. Using the data from this experiment, we will create and F-I (frequency-current) curve.
## Outline of this notebook
[3.0 Import statements](#3.0-Import-statements)
[3.1 More on for-loops and if-else statements](#3.1-for-loops-and-if-else-statements)
* A review on for-loops and if/else statements, and some new tricks for for-loops
[3.2 Load single cell electrophysiology data from csv file](#3.2-Load-single-cell-electrophysiology-data-from-.csv-file)
* Inspect data using pandas
* Visualize data with matplotlib
[3.3 Analyze a single sweep of electrophysiology data](#3.3-Analyze-a-single-sweep-of-electrophysiology-data)
* Calculate the firing rate of the neuron during a single sweep of data
[3.4 Create an F-I (frequency - current) curve](#3.4-Create-an-F-I-curve-for-a-single-cell)
* Write a function to calculate firing frequency during one sweep (3.4)
* Employ function in a for-loop to perform the operation for all sweeps
[3.5 Bonus exercises](#3.5-Bonus-exercises)
* Calculate the input resistance of a cell
* Extract spike cutouts
* Create a phase plot for a spike (rate of change of voltage vs. voltage)
[3.6 Appendix](#Appendix)
* Additional information about loading binary file formats
# 3.0 Import statements
```
import os
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import scipy.signal as ss
## Note: the following line is only needed inside the
## Jupyter notebook, it is not a Python statement
%matplotlib inline
```
# 3.1 for-loops and if-else statements
Yesterday you learned how to use a for loop to iterate over the elements of a list (refer to section 2.7). Today we'll learn a couple additional tricks you can use when constructing for-loops.
Often we only want to perform an operation on the current element when a specific condition is met. For this, we use **if-else statements**.
Though it's nice to loop over the elements in a list, sometimes we also would like the index where that element occurs. For this we can use the function **`enumerate()`**.
Let's create some dummy data in a Pandas `DataFrame` to illustrate these points:
```
# Let's use the numpy's rand() function to create an array of random floats with
# shape (100,10)
data = np.random.rand(100,10) # creates a 100 x 10 numpy array of random numbers between 0 and 1
# then create column names for the DataFrame
cols = ['one','two','three','four','five','six','seven','eight','nine','ten']
# and convert our data into a DataFrame and cacluate indices
df = pd.DataFrame(columns = cols, index = range(0,data.shape[0]), data = data)
df.head(5)
```
### Exercise 1 - Review from day 2
> * Refer to section 2.7 from yesterday to write a for loop that loops over all the column names in `df` and prints them out.
```
# save list of all column names in the df above into the variable: cols
cols =
# Print each col name using a loop.
# Your loop goes here:
```
Sometimes, we want to have access not only to the column names ('one', 'two' etc.), but also to the correspondings indexes for these elements (0, 1, 2 etc.). We can achieve this by using a built-in function [enumerate()](http://book.pythontips.com/en/latest/enumerate.html).
One example of why this might be useful is if you only want to perform an operation on the columns of **odd** indexes. With only the column name, we can't do this. Using a combination of the enumerate and if-else statements (refer to section 1.14 of Day 1) we can do this:
```
for i, col in enumerate(cols):
if (i+1)%2 == 0: # notice that i starts counting from 0, so we must add 1
pass # "pass" tells python to skip this if-else statement, finish the code block,
# and move on to the next iteration in the loop
else:
print("index: "+str(i))
print("column name: "+col)
```
### Exercise 2
>* Use what you just learned about for-loops to write a loop that prints out only the names of the even numbered columns.
```
# Your code goes here
```
Yesterday you also learned a little bit about how to write your own functions. Below is an illustration of how functions, loops, and if-else statements can be combined in a useful way. It's a lot, but we will use these concepts later so make sure you have a good grasp on the code below.
```
def print_even_or_odd_column(dataframe, cols='even'): # note that the default value of columns is even
columns=dataframe.columns
# code to be executed if you tell the function to print even columns
if cols=='even':
for i, col in enumerate(columns):
if (i+1)%2==0:
print(col)
else:
pass
# code to be executed if you tell the function to print off columns
elif cols=='odd':
for i, col in enumerate(columns):
if (i+1)%2!=0:
print(col)
else:
pass
# ==== Call the function ====
print_even_or_odd_column(df,cols='even') # can set the last argument as: cols='odd' or cols='even'
```
# 3.2 Load single cell electrophysiology data from .csv file
**Set file path to day-2 of the python_neurobootcamp (concepts from Day 2 - section 2.1)**
```
#print "Current Working Directory (cwd)" to check we're in the right place and don't need absolute path
os.getcwd()
path = 'csv_data/'
```
#### The data:
As those of you who do patch clamp know, e-phys data doesn't come nicely packaged into a simple spreadhsheet format. Therefore, we have written a function that converts axon binary files (.abf) into spreadheet format (.csv) for the purposes of this course.
If you are interested in exactly how this process works, look into the file: `binary_file_loading_tools.py` in the `day-3` directory or see Appendix section at the bottom of this notebook. It's a little bit rough, but will help give you an idea of how you can convert binary files to spreadsheets. After completing this course, you should be able to modify this file on your own to suit your needs!
A reminder about csv files: CSV = Comma-Separated Value file that stores tabular data (numbers and text) in plain text. You can save an excel spreadsheet in .csv and then import it in Python. Let's look at an example. (open up csv file from the day-3/csv_data folder in excel)
**Load spreadsheet data into pandas**
Note the use of `index_col=0`. Try to get rid of this and see what happens when you load based on default settings. For more information: https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html
```
meta = pd.read_csv("csv_data/meta_data_PV_3_10_03_2014.csv", index_col=0)
data = pd.read_csv("csv_data/data_PV_3_10_03_2014.csv",index_col=0)
print(meta.shape)
meta.head()
```
Notice that in our meta data, we have information about the sampling rate (fs), cell type, date of recording, units for ch1 data, and units for ch2 data for each sweep in the binary file that we loaded. This information is particulalrly useful for analyses in which, for example, we want to group anlayses by cell type. Today we won't use it a whole lot, but it's a common way this type of information is stored.
```
print(data.shape)
data.head()
data.index.name='Time'
data.head()
```
Though the channels aren't labeled, we can determine what each represents by the their units (pico-amps or milli-volts in this case) which we can get above (in meta data).
Notice that data contains the time series for each channel on each sweep. The indexes here represent the time points that each value was acquired at.
### Visualization using matplotlib and seaborn
We learned a bit about seaborn yesterday. Matplotlib is another widely used plotting library in Python. Everyone has their own preference for which they like to use. Here we will go through examples of each, though we will focus on matplotlib. You can decide for yourself which you prefer.
Before we get into analysis, it's always a good idea to inspect your raw data to make sure you know what's there, and that it was loaded properly.
First let's use Pandas built-in `filter()` function to assign all ch1 traces to a `DataFrame` called `voltage_traces` and all ch2 traces to a `DataFrame` called `current_traces`. For basics of filtering, refer back to section 2.9.
Here we use regular expressions to filter the data. These can be very confusing at first, but don't worry. Regular expression are very powerful and worth spending some time getting used to. For more information: https://docs.python.org/2/library/re.html
```
# Based on meta data, we can tell that ch1 refers to voltage and ch2 refers to current
voltage_traces = data.filter(regex="ch1.", axis=1) # to filter on rows instead of columns, use axis=0
current_traces = data.filter(regex="ch2.",axis=1)
voltage_traces.head()
```
### Plot using matplotlib
```
fig, ax = plt.subplots(2,1,sharex=True) # Create a figure and an axis. Specify the layout: (2 rows, 1 column)
# there are now two axes for your plots (ax[0] and ax[1])
# plot the voltages traces on the first subplot
ax[0].plot(voltage_traces)
# Add a figure legend (this is easy because pandas and matplotlib play nice together)
ax[0].legend(voltage_traces,loc='upper right')
# If data wasn't in a pandas frame, could also do something more explicit, like pass it a list of names:
# ax[0].legend(['ch1_sweep1', 'ch2_sweep2'...], loc='upper right')
# set the title of the first subplot
ax[0].set_title('Voltage traces')
# set the ylabel of the first subplot (using the meta data to get units)
ax[0].set_ylabel(meta['ch1_units'][0])
# Create the same plot in your second set of axes for current_traces
ax[1].plot(current_traces)
ax[1].legend(current_traces,loc='upper right')
ax[1].set_title('Current traces')
ax[1].set_xlabel('Time (s)')
ax[1].set_ylabel(meta['ch2_units'][0])
plt.tight_layout() # trick to make formatting look a little nicer
```
### Plot using Pandas
Pandas offers wrapper functions around matplotlib that make the above code a bit more concise. We'll use this (plotting with Pandas) quite a bit in this notebook.
Just note that what it's really doing is written out explicity above. See [here](https://pandas.pydata.org/pandas-docs/stable/visualization.html) for a little more explanation.
```
fig, ax = plt.subplots(2,1,sharex=True)
# Call plot. plot is a dataframe method that in turn calls the matplotlib function plot
voltage_traces.plot(title='Voltage Traces', ax=ax[0])
current_traces.plot(title='Current Traces', ax=ax[1])
## Position the legends
ax[0].legend(loc='upper right')
ax[1].legend(loc='upper right')
## Set X and Y axis labels
ax[0].set(ylabel=meta['ch1_units'][0])
ax[1].set(ylabel=meta['ch2_units'][0], xlabel='Time (s)')
plt.tight_layout()
```
### Plot using seaborn
Note that there are some slight tweaks you need to make to the data in this case in order to use seaborn, including downsampling the data (otherwise it takes A LONG time to run). Mostly for this reason, we'll stick with using matplotlib for the rest of today. Just be aware that you can also use seaborn if you prefer.
Reformat (melt) the data into "long form" from "wide from". This is a step that will be necessary with any data in wideform that you want to plot in this way - using tsplot.
```
## Reformat the dataframe to work with a Seaborn time-series plot
voltage_ts = voltage_traces.iloc[:,:5].copy()
voltage_ts['Time'] = voltage_ts.index
voltage_ts = pd.melt(voltage_ts, id_vars='Time', var_name='Sweep', value_name='mV')
voltage_ts['Dummy'] = 0
```
Downsample the data so that this runs faster. This is quirk for this type of data that is sampled at a very high sampling rate making seaborn run rather slowly.
```
## Downsample the data (take every 50th row)
voltage_ts = voltage_ts.iloc[::50,:]
voltage_ts.head()
```
Perform the same operations on the current trace data.
```
## Do the same for the current traces
## Reformat the dataframe to work with a Seaborn time-series plot
current_ts = current_traces.iloc[:,:5].copy()
current_ts['Time'] = current_ts.index
current_ts = pd.melt(current_ts, id_vars='Time', var_name='Sweep', value_name='pA')
current_ts['Dummy'] = 0
## Downsample the data (take every 50th row)
current_ts = current_ts.iloc[::50,:]
```
Now, plot the data using the seaborn function `tsplot` [time-series plot](http://seaborn.pydata.org/generated/seaborn.tsplot.html).
Note in particular the argument "unit". We must pass it something otherwise seaborn will run into issues when it tries to calculate confidence intervals for this data. Therefore we just pass a "Dummy" unit to make seaborn happy since we don't really care about the confidence interval in this case.
```
# This will take a minute to load
fig, ax = plt.subplots(2,1,sharex=True)
sns.tsplot(data=voltage_ts, time='Time', condition='Sweep', unit='Dummy', value='mV', err_style=None, ax=ax[0])
sns.tsplot(data=current_ts, time='Time', condition='Sweep', unit='Dummy', value='pA', err_style=None, ax=ax[1])
# Position the legends
ax[0].legend(loc='upper right')
ax[1].legend(loc='upper right');
```
These are all pretty messy and don't tell us a ton about the data. Can we make it better?
### Exercise 3
* Try plotting only the last sweep of both current and voltage so that we can see what's going on more easily.
* Use the code from above as a starting point for how to create your own plot (you should be able to get most of the
way there with copy and paste). Ask a TA or instructor if you need help.
**Step 1)** Find the column id for the last sweep (there are multiple ways to do this).
Hint: As you might remember from Day-1, we can use "-1" to grab the last index of something. This is equivalent to using "end" in Matlab.
Hint: we can return a list of data frame columns by using our_data_frame.columns
```
lastSweep_voltage =
lastSweep_current =
print(lastSweep_voltage)
print(lastSweep_current)
```
**Step 2)** Now, use these column id's to select only the columns containing data for the last sweep and assigne these to new variables called: last_voltage and last_current
```
last_voltage =
last_current =
# Plot with Pandas
# Create a figure with two subplots that share the x axis (like we did above)
# plot the last voltage trace and last current trace
## Position the legends
## Set X and Y axis labels
```
That looks better, but can we zoom in on some of the spikes in the voltage traces?
### Exercise 4
* Plot only time points from 0.02 to 0.10 seconds:
** Step 1) ** Define the list of indexes (times) that are greater than 0.02 and less than 0.1. Create a mask (array of True and False values) where the condition is true. Use the mask to select only the appropriate indexes (time values).
```
# Make a mask
mask =
print(mask)
# Use mask to select the appropriate indexes (time values) ***remember these are stored in the index of our data frame
indexes =
print(indexes)
```
** Step 2) ** Use the indexes you just defined to select only the rows of last_voltage and last_current where time > 0.02 and time < 0.1. Save these results into two new data frames called: **last_volt_short** and **last_curr_short**
```
last_volt_short=
last_curr_short=
```
** Step 3) ** Plot the result with pandas
* Note: I've filled in the plotting commands to save time, which means this next cell will only work if you correctly defined ** last_volt_short ** and **last_curr_short** in the cell above
```
## Plot with Pandas
## Create a figure and plot voltage and current traces
fig, ax = plt.subplots(2,1,sharex=True)
last_volt_short.plot(title='Voltage Traces', ax=ax[0])
last_curr_short.plot(title='Current Traces', ax=ax[1])
## Position the legends
ax[0].legend(loc='upper right')
ax[1].legend(loc='upper right')
## Set X and Y axis labels
ax[0].set(ylabel=meta['ch1_units'][0])
ax[1].set(ylabel=meta['ch2_units'][0], xlabel='Time (s)')
plt.tight_layout();
```
# 3.3 Analyze a single sweep of electrophysiology data
A common way to show a neuron's response to stimulation is to make a frequency-intensity curve (F-I curve). The frequency of firing is plotted on the y-axis and the amplitude of the current injection that caused the spikes is plotted on the x-axis.
From looking at these F-I curves you can identify if the cell is transiently firing, whether the spike rate reaches a maximum at some high current step, the gain of firing (the slope of the line, in Hz/pA), etc. To construct a F-I curve we need to first find the number of spikes per unit time in each sweep and the current injection that drove those spikes. Then plot it.
First, we will perform the analysis for just one sweep of data. Then, if there is time, we will define a function to help automate the process. We will run the function on all sweeps using a for-loop, and plot the results in a F-I curve.
Becasue we have already pulled out the last voltage trace, let's go ahead and analyze this time-series first.
The analysis will consist of a few key steps:
* Step 1: Locate the time points of all spikes
* Step 2: Find the time window and magnitude of current injection
* Step 3: Calculate the firing rate
### Step 1: Locate the time points of all spikes
* There are a couple of ways to do this. We will do it by locating the relative maxima of the voltage trace during the current injection
* We will also set a threshold above which to detect maxima (to avoid counting local maxima in the baseline (noise) as a spikes)
```
threshold = -20
last_voltage_thresholded = last_voltage[last_voltage > threshold]
```
Let's take a look at what happened when we thresholded the voltage trace:
```
## Plot with Pandas
## Note: X-axis is different for each subplot (what times were cut out by thesholding)
fig, ax = plt.subplots(2,1)
last_voltage_thresholded.plot(title='Thresholded Trace', ax=ax[0])
last_voltage.plot(title='Raw Trace', ax=ax[1])
## Position the legends
ax[0].legend(loc='upper right')
ax[1].legend(loc='upper right')
## Set X and Y axis labels
ax[0].set(ylabel=meta['ch1_units'][0], xlabel='Time (s)')
ax[1].set(ylabel=meta['ch1_units'][0], xlabel='Time (s)')
plt.tight_layout();
```
Using a a threshold of -20, we have selected all the voltage points above -20 mV. In other words, these are all points during which an action potential was happening.
*Scipy*, another useful Python package, has a function that finds the indices where relative extrema occur: `argrelextrema`
This function takes two required arguments: data and comparator (for details see [documentation](https://docs.scipy.org/doc/scipy-0.19.0/reference/generated/scipy.signal.argrelextrema.html)).
* Note: `last_voltage_thresholded` is a Pandas DataFrame. `argrelextrema` expects data to be an Array, not a DataFrame.
* Note: `argrelextrema` also takes an optional argument: "order" that specifies the amount of points on each size to use for the comparison.
```
indexes_of_maxima = ss.argrelextrema(data=last_voltage_thresholded.values, comparator= np.greater, order=1)
np.greater(4,3) # np.greater is a function that compares two input arguments
```
The Scipy function returns indexes, *i.e.* 1,2 3... BUT we want the time points (which are stored in the index of our DataFrame)
```
spike_times = last_voltage_thresholded.index[indexes_of_maxima]
```
It's possible that the voltage trace is a little noisy, even during a spike. This might cause a spike to be double counted by the Scipy function, which is only locating relative/local maxima. To fix these cases, we will impose a constraint based on the refractory period of neuron. To do this, we'll first define a constant refractory period limit:
```
refractory_period_low_limit = 0.002 # 2ms is a lower limit on the refracotry period for a neuron
```
Now, let's calculate the interspike intervals for all the maxima we located (using numpy's ediff1d function) and exclude any spikes that violate the refractory period.
* np.ediff1d: Calculates the difference between all adjancent elements in a 1d array: https://docs.scipy.org/doc/numpy/reference/generated/numpy.ediff1d.html
* np.arghwere: Locates the indexes where the given condition is met: https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.argwhere.html
```
interspike_interval = np.ediff1d(spike_times)
```
Remove the refractory period violations
```
VIOLATION_indexes = np.argwhere(interspike_interval<refractory_period_low_limit)
spike_times = spike_times.delete(VIOLATION_indexes)
print(spike_times)
```
We now have an array of time points where we think spikes occured. Let's plot them to see if we were successful
```
## Plot with Pandas
fig, ax = plt.subplots(1,1)
ax = last_voltage.plot(title='Raw Trace')
ax.set(ylabel=meta['ch1_units'][0], xlabel='Time (s)')
# plot a point at each time point where a spike was detected
ax.plot(spike_times, last_voltage[spike_times], ".", markersize=20);
```
This is looking pretty good.
Now, let's zoom in on some spikes to make sure we really are counting each individual spike.
```
## Plot with Pandas
fig, ax = plt.subplots(1,1)
ax = last_volt_short.plot(title='Raw Trace')
ax.set(ylabel=meta['ch1_units'][0], xlabel='Time (s)')
ax.plot(last_volt_short[spike_times], ".", markersize=9); # plot a point at each time point where a spike was detected
```
Great, looks like we are capturing all the spikes, not double counting, and not counting any noise as spikes.
One final check we can perform is just to look at all the raw ISI (interspike interval) values to make sure there wasn't a change in frequency over the course of the stimulation (high firing at the beginnning, slower at the end). To do this, let's define a variable called ISI_check using numpy's ediff1d funciton:
```
ISI_check = np.ediff1d(spike_times)
plt.figure()
plt.hist(ISI_check*1000,bins=50) # convert to ms by multiplying by 1000
plt.ylabel('Count')
plt.xlabel('Interspike interval (ms)');
```
### Exercise 5
* Go back and re-run everything from "Step 1" but this time, DO NOT run the cells that corrected for refractory period violations (the cell that begins with: VIOLATION_indexes...) This should illustrate why this final histogram can be a good check on your analysis.
### Step 2: Find out the time window of current injection
This is pretty straightforward for this experiment because as we saw earlier, it's just square pulse injections. Therefore, we can just find the time points where `current == max(current).
```
current_magnitude = max(last_current.values)
current_inj_times = last_current.index[last_current==current_magnitude] # time points when current is being injected
current_inj_length = max(current_inj_times) - min(current_inj_times) # duration in seconds
print("Current injection: "+str(current_magnitude)+" pA")
print("Current injection times:")
print(current_inj_times)
print('Current injection length: '+str(current_inj_length)+" s")
```
Before we calculate the firing frequecny, there is one last check we must perform. It is possile that some spontaneous spikes occured outside the current injection window (it didn't happen in this case, but it could and that could change our results). To handle this, we remove all spikes times outside of the current injection window using a boolean mask (list of `True` and `False`) which we can then use to only grab indixes where there is a one:
```
boolean_array = (max(current_inj_times)>= spike_times) & (spike_times >= min(current_inj_times)) # boolean array (0's and 1's)
print(boolean_array)
```
Now, use the boolean mask to get only the spike times inside the appropriate time window
```
spike_times = spike_times[boolean_array] # only saving spike times where boolean_array=True e.g. during current injection
```
### Step 3: Calculate the firing rate
There are couple of different ways to calculate firing frequency. They are both useful and tell you different things.
The most straighforward way is to simply count the number of spikes and divide by the amout of time (in seconds) that the current was being injected.
```
n_spikes = len(spike_times) # count the number of spikes
spike_freq = n_spikes/current_inj_length # divide the number of spikes by the curent injection time
print("spiking frequency = " + str(round(spike_freq,2))+" Hz")
```
The second way to go about this is to calculate the average interspike interval. This is particularly useful if the neuron does not continue to fire or adapts during the current injection.
```
ISI = np.ediff1d(spike_times) # interspike interval (ISI) (as we did for the histogram)
meanISI = np.mean(ISI) # calculate the mean ISI
spike_freq = 1/meanISI # ISI is in units of seconds, do: 1s/meanISI to convert to mean firing frequency
print("spiking frequency = " +str(round(spike_freq,2))+"Hz")
```
For this case, both methods give us basially the same result because as we saw above, this neuron fires pretty regularly during the entire stimulation. This isn't necesarily the case, so it's good to check.
#### Summary of single sweep analysis
Although what we really want is the F-I curve for the cell, analyzing one sweep at a time, like we did here, is often a good idea. This way, you can make sure that the code you write is really analyzing the data in the way you intend. Plus, at this point we have basically written all the code we need to analyze any sweep of data. In order to perform the analysis for the whole experiment, we just need to wrap all the code we wrote into a function that can be call iterativley from a for-loop!
# 3.4 Create an F-I curve for a single cell
To do this, we will perform the above analysis many times. Once for each current injection/sweep.
Because we are doing the same thing many times, rather than write all of the above code 20 times (once for each sweep), we'll write a for-loop that performs the same operation many times
When you are performing an operation many times, it is often nice to write function to perform this operation. We will write a function that returns the firing frequency for any sweep.
**Let's write a function that returns the firing rate for a single sweep of data:**
Note: this looks like a lot of code, but, everything inside this function is stuff we already wrote above!
```
def get_firing_rate(v, threshold, tstart, tend, method="ISI"):
'''
================ This is called a doc string ====================
========== It tells you how to use the function =================
======= It is what is printed when you call "help" ==============
Arguments:
v (data frame): array of voltage values during one sweep with the index being the time series
threshold (float): voltage cut-off to count spikes
tstart (float): time current injection begins
tend (float): time current injection ends
method (string, optional): method for calculating the firing rate
ISI: use interspike interval
AVG: use average over whole current injection window
Output:
firing_rate (float): firing rate during the period defined by tstart and tend
'''
refractory_limit=0.002 # set refractory period limit (to avoid double counting spikes)
current_duration = tend-tstart
v_thresh = v[v>threshold]
spike_indexes = ss.argrelextrema(v_thresh.values, np.greater,order=1) # indices of spikes (0,1,2...)
spike_times = v_thresh.index[spike_indexes] # convert to time (in seconds)
# get rid of refractory violations
interspike_interval = np.ediff1d(spike_times)
VIOLATION_indexes = np.argwhere(interspike_interval<refractory_limit)
spike_times = spike_times.delete(VIOLATION_indexes) # delete any spike times that occured too soon after a previous spike
# get rid of spikes outside the current injection window
tf = (tend >= spike_times) & (spike_times >= tstart) # boolean array (0's and 1's)
spike_times = spike_times[tf] # only saving spike times where tf==1 e.g. during current injection
if method=="ISI":
# calculate the firing frequency using interspike interval
# in case there were no spikes, set firing rate to 0 Hz
if spike_times.size==0:
firing_rate=0 # there were no spikes, so firing rate is 0
else:
ISI = np.mean(np.ediff1d(spike_times))
firing_rate = 1/ISI
elif method=="AVG":
firing_rate = len(spike_times)/current_duration
return firing_rate
```
There is alot going on in that function. If you are confused on any part of it, please ask the instructor or TAs for assitance.
* You should understand how the function works, and how to modify it if you need/want to
* You should understand the difference between required and optional function arguments
* You should understand the if-else statements at the end of the function definition
Note: check out what happens when you call "help" on the function we just wrote:
```
help(get_firing_rate)
```
**Write a for-loop to calculate the firing rate for every sweep of data**
First, let's intitalize the constants that we will use throughout the analysis
* threshold
* current_duration
```
# set threshold above which to detect spikes
threshold = -20
# Find current duration (same for all sweeps so we can just do this once)
current_magnitude = max(last_current.values)
current_inj_times = last_current.index[last_current==current_magnitude] # time points when current is being injected
I_start = min(current_inj_times)
I_end = max(current_inj_times)
current_duration = max(current_inj_times) - min(current_inj_times) # duration in seconds
```
Loop over all the sweeps, find the current magnitude and firing rate, print them out
```
sweeps = meta.index # define the list of sweeps to loop over
for i, sweep in enumerate(sweeps):
# Call our function that we wrote above (get_firing_rate)
firing_rate = get_firing_rate(data['ch1_'+sweep], threshold, I_start, I_end, method="ISI")
# get current injection magnitude
# Note how we deal with negative current injections using if-else statements
I_mag = max(data['ch2_'+sweep])
if I_mag == 0 and min(data['ch2_'+sweep])==0: # case where current = 0
I_mag = 0
elif I_mag == 0 and min(data['ch2_'+sweep]) != 0: # case where current < 0
I_mag = min(data['ch2_'+sweep])
# print the result
print(sweep+": "+"I: "+str(I_mag)+" "+meta['ch2_units'][i]+ ", Firing rate: "+str(firing_rate)+" Hz")
```
There's a lot of stuff going on here as well. Some key takeaways:
* Make sure you understand the for loop, what it's doing, and why it's useful
* Make sure that the if-else statements inside the for loop are making sense
* If you are confused on these points, ask the instructor or a TA
Now that we understand how the code above is working, let's run it again but this time actually save the results instead of just printing them out. To save our results, let's first create a new data frame called "results" to store everything.
```
results = pd.DataFrame(columns=["Current Injection", "Firing Rate"],index = meta.index)
results.head()
```
We've now created an empty data frame to store the results for all data sweeps ("NaN" means "empty" or "undefined"). Using a for loop, let's again calculate the current injection and firing rate for each sweep and fill up the empty results DataFrame.
```
# copy and paste the code from above, but save results this time
for i, sweep in enumerate(sweeps):
firing_rate = get_firing_rate(data['ch1_'+sweep],threshold,I_start,I_end,method="ISI")
I_mag = max(data['ch2_'+sweep])
if I_mag == 0 and min(data['ch2_'+sweep])==0:
I_mag = 0
elif I_mag == 0 and min(data['ch2_'+sweep]) != 0:
I_mag = min(data['ch2_'+sweep])
results['Current Injection'][sweep]=I_mag
results['Firing Rate'][sweep]=firing_rate
```
Now, we have stored all our results into a new data frame.
```
results
```
And we can create our F-I plot for this neuron
```
## Plot with Pandas
plt.figure()
ax = results.plot('Current Injection', 'Firing Rate', title='F-I Curve', marker=".", color="r", lw=2, markersize=5, alpha=0.6)
ax.set(xlabel='Current '+meta['ch2_units'][0], ylabel='Firing rate (Hz)');
```
Cool, we now have an F-I curve for this PV neuron.
### Final Excercise: do the same analysis but calculate frequency as n_spikes/time instead of using the ISI as we just did
> * Add the result to the DataFrame called `results` and name this new column `"Firing rate (avg)"`
**Hint**: one way to add a column to a DataFrame is to make a series and then add it to the original DataFrame
* newcolumn = pd.Series(index=, name=)
* pd.concat([results,newcolumn],axis=1)
* Once you've accomplished all this, plot the results alongside the trace above
[More on how to create new column using Pandas series](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.html)
```
# Create a new, empty column with the correct number of rows using pd.Series()
newColumn = pd.Series()
```
Now loop over all the data and fill in the new column.
** Remember: ** Useful variables (like threshold, I_start and I_end) are already defined above so we can use them again here!
```
# Write a loop to calculate firing rate and save it into the the new column
for in :
firing_rate =
newColumn[sweep]=
```
Concatenate `newColumn` with results
Check out [this](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.concat.html) for help. Or recall how we concatenated data frames during yesterday's lesson.
```
# concatenate data frame. What axis do you want to concatenate on?
results_big = pd.concat()
results_big
```
Plot the results for both methods on the same figure. You can steal (copy and paste) a lot of code from above to do this. You want to create a single plot with two lines on it. Using [this documentaion](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.plot.html) might also be helpful.
```
# Use examples of plotting from above, and follow the commented suggestions
# Create a figure
# plot the ISI results in red (using color='red')
# plot the AVG results
# label axes
# set legend
# set title
```
Basically, we get the same result. However, we do see a clear difference for the first positive current step, when the cell did not continue to fire over the course of the current injection but only fired a burst of spikes at the beginning of stimulation
# 3.5 Bonus exercises
### 1. Calculate the input resistance of the cell (Easier)
Recall Ohm's law: voltage = current * resistance (V=IR). We know the current and we know the voltage so we can solve for R, the cell's input resistance. Using ohms law and the hints below, solve for this cell's input resistance.
* Hint 1: Input resistance is a passive property of a neuron, therefore we'll have to measure it when voltage activated channels are not opening/closing. Remember from our analysis above, there were a bunch of negative current steps that elicited no action potentials.
* Hint 2: Even though the negative current steps didn't cause action potentials to fire there could still have been some channels opened, changing the resistance of the cell (for example, Ih currents). Therefore, it's probably best to use voltage measurements when the cell was in a steady state (later on in the current injection).
* Hint 3: Ohms law is a linear relationship. In other words, if we plot voltage vs. current we should see a straight line (if no channels were opening and closing). The slope of this line is R. Plot voltage vs. current for all negative current injections and meausre the slope at the point where the line is most linear.
```
# ============================== Solution ========================================
```
### 2. Create a phase-plot for this cell (More difficult)
A phase plot is a plot of voltage vs. the derivative of the voltage (rate of change) during an action potential. Here's an example:
<img src = 'phase_plot.png'>
Your goal is to create a phase plot for the action potentials fired during the 500pA current injection. There a few steps involved to do this:
* Step 1: Generate an array of "spike cutouts" during this current step. In other words, make a 2D numpy array where the rows are time points and the columns are individual action potentials. Do this by taking 1ms of data before an AP peak and 2ms of data after a peak. Hint: remember how we found the time points of AP peaks earlier...
* Step 2: Generate an equivalent numpy array except in this array, each column is the derivative of the action potential in the array from step 1
* Step 3: Take the mean of all columns in both arrays and then plot action potential mean on the x-axis vs. the derivative mean on the y-axis
```
# ============================== Solution =======================================
```
# 3.6 Appendix
### Loading binary files (.abf, .dat, etc.) using the python package, neo
*neo* is an open source python package, like numpy or pandas. It was written specifically to help load binary file formats from electrophysiology data into python. We have written a function that uses this package to load .abf files into a .csv data format (spreadsheet). If you are interested in learning more about this, ask one of your TA's or intsructors, or just go look at the file binary_file_loading_tools.py that is located in the day 3 folder. Note, in order to run the loading function, you will need neo installed on your machine. To do this, type: "pip install neo" into a terminal window on your machine. For more information on neo, see: http://neuralensemble.org/neo/
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from mpes import fprocessing as fp, visualization as vis
import matplotlib.cm as cm
fdir = r'../data/data_20180605_119.h5'
axes = ['X', 'Y', 't']
# Pick very odd number of bins to create the artefact
bins = [91, 91, 230]
ranges = [(300, 1700), (300, 1700), (68500, 72500)]
enum_max = 4e7
hp = fp.hdf5Processor(fdir)
```
### Binning without data jittering
```
binned_nojitter = hp.localBinning(axes=axes, nbins=bins, ranges=ranges, amax=enum_max)
binned_nj = binned_nojitter['binned']
```
### Binning with data jittering
Jitter amplitude used = 0.1 * bin size
```
binned_jitter = hp.localBinning(axes, bins, ranges, jittered=True, \
jitter_amplitude=[1e-1]*3, amax=enum_max);
binned_jt = binned_jitter['binned']
```
### Comparison plots
#### (1) 2D cuts of the band structure
```
f, axs = plt.subplots(1, 2, figsize=(10,5))
vis.colormesh2d((binned_nj[50,::-1,::-1]).T, colormap='terrain_r',\
vmin=0, cbar=True, plotaxes=axs[0])
vis.colormesh2d(binned_jt[50,::-1,::-1].T, colormap='terrain_r',\
vmin=0, cbar=True, plotaxes=axs[1])
axs[0].set_title('Without jittering', fontsize=20)
axs[1].set_title('With jittering', fontsize=20);
#plt.savefig('Jittering_2D.png', bbox_inches='tight', dpi=300)
fd, axd = plt.subplots(1, 1, figsize=(5,5))
vis.colormesh2d(binned_nj[50,::-1,::-1].T - binned_jt[50,::-1,::-1].T, colormap='RdBu_r', \
cscale={'vmin':-20, 'vmax':20}, plotaxes=axd);
# plt.savefig('Jittering_2D_diff.png', bbox_inches='tight', dpi=300)
```
#### (2) Angle-integrated energy dispersion curve (EDC)
```
nj_1d_EDC = binned_nj.sum(axis=(0,1))
jt_1d_EDC = binned_jt.sum(axis=(0,1))
fedc, axedc = plt.subplots(figsize=(10,6))
plt.plot(binned_nojitter['t'], nj_1d_EDC, lw=2, label='without jittering')
plt.plot(binned_jitter['t'], jt_1d_EDC, 'r', lw=2, label='with jittering')
plt.plot(binned_jitter['t'], nj_1d_EDC - jt_1d_EDC, 'k', lw=2, label='difference')
axedc.set_xlabel('Energy (a.u.)', fontsize=15)
axedc.set_ylabel('Photoemission intensity (a.u.)', fontsize=15)
plt.legend();
#plt.savefig('Jittering_1D.png', bbox_inches='tight', dpi=300)
```
#### (3) Energy-integrated momentum dispersion curve (MDC)
```
nj_1d_MDC = binned_nj[:,:,100:].sum(axis=(0,2))
jt_1d_MDC = binned_jt[:,:,100:].sum(axis=(0,2))
fmdc, axmdc = plt.subplots(figsize=(10,6))
plt.plot(binned_nojitter['Y'], nj_1d_MDC, lw=2, \
label='without jittering')
plt.plot(binned_jitter['Y'], jt_1d_MDC, 'r', lw=2, \
label='with jittering')
plt.plot(binned_jitter['Y'], nj_1d_MDC - jt_1d_MDC, 'k', lw=2, label='difference')
axmdc.set_xlabel('Momentum k$_y$ (a.u.)', fontsize=15)
axmdc.set_ylabel('Photoemission intensity (a.u.)', fontsize=15)
plt.legend();
```
Combine the plots
```
# rect [left, bottom, width, height]
f, ax = plt.subplots(figsize=(5, 5))
vis.colormesh2d(binned_nj[50,::-1,::-1].T - binned_jt[50,::-1,::-1].T, colormap='RdBu_r', \
cscale={'vmin':-20, 'vmax':20}, plotaxes=ax);
ax.set_xticks([])
ax.set_yticks([])
ax.xaxis.set_label_position("top")
ax.set_xlabel('Momentum ($\AA^{-1}$)', fontsize=15)
ax.set_ylabel('Energy (eV)', fontsize=15)
ax_edc = f.add_axes([0.98, 0.08, 0.25, 0.88])
ax_edc.plot(nj_1d_EDC[::-1], binned_nojitter['t'], lw=2, label='without jittering')
ax_edc.plot(jt_1d_EDC[::-1], binned_jitter['t'], 'r', lw=2, label='with jittering')
ax_edc.plot(nj_1d_EDC[::-1] - jt_1d_EDC[::-1], binned_jitter['t'], 'k', lw=2, label='difference')
ax_edc.set_xticks([])
ax_edc.set_yticks([])
ax_edc.set_xlabel('Intensity', fontsize=15)
ax_edc.xaxis.set_label_position("top")
ax_mdc = f.add_axes([0.1, -0.2, 0.85, 0.25])
ax_mdc.plot(binned_nojitter['Y'], nj_1d_MDC, lw=2, \
label='w/o jittering')
ax_mdc.plot(binned_jitter['Y'], jt_1d_MDC, 'r', lw=2, \
label='w/ jittering')
ax_mdc.plot(binned_jitter['Y'], nj_1d_MDC - jt_1d_MDC, 'k', lw=2, label='difference')
ax_mdc.set_xticks([])
ax_mdc.set_yticks([])
ax_mdc.set_ylabel('Intensity', fontsize=15)
# ax_lg = f.add_axes([0.98, -0.2, 0.25, 0.25])
ax_mdc.legend(loc='lower right', bbox_to_anchor=[1.1, 0, 0.25, 0.25], frameon=False, labelspacing=1.5)
# ax_lg.set_xticks([])
# ax_lg.set_yticks([])
# plt.savefig('./Tutfiles/Jittered_Panel.png', dpi=300, bbox_inches='tight')
```
### Parallel version
```
import glob as g
from time import time
hfiles = g.glob(r'../data/data_114_parts/data*.h5')
_, hfsorted = fp.sortNamesBy(hfiles, r'\d+', slicerange=(-10, None))
parp = fp.parallelHDF5Processor(hfsorted[:5])
tstart = time()
parp.parallelBinning(axes=axes, nbins=bins, ranges=ranges, scheduler='threads',
binning_kwds={'jittered':True, 'jitter_amplitude':[2e-1]*3}, ret=False)
tend = time()
print('Binning took about {} s.'.format(int(tend - tstart)))
result = parp.combineResults()
plt.figure(figsize=(10,6))
plt.imshow(result['binned'][50,...].T, cmap='terrain_r', aspect=0.4)
plt.plot(result['binned'].sum(axis=(0,1)))
plt.plot(result['binned'][:,:,100:].sum(axis=(0,2)))
```
| github_jupyter |
```
#import packages
import datetime as dat
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pylab
import scipy.stats as stats
import seaborn as sns
from sklearn.ensemble import RandomForestClassifier
import time
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import StratifiedKFold
#load data
df = pd.read_csv('Data/dataframe.csv')
#drop the unamed column
df = df.drop(['Unnamed: 0'], axis=1)
#view data
df.head()
#deal with class imblanance
#first divide into target and features
# Specify the name of the target
target = 'EV'
# Get the target vector
y= df[target]
features = df.drop('dataid', axis=1)
# Specify the name of the features
features = list(features.drop(target,axis=1).columns)
# Get the feature vector
x= df[features]
from sklearn.model_selection import train_test_split
# Randomly choose 30% of the data for testing (using random_state = 0)
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.3, random_state=0, stratify=y)
from imblearn.under_sampling import RandomUnderSampler
rus = RandomUnderSampler(random_state=0)
X_resampled, y_resampled = rus.fit_resample(X_train, y_train)
#choose the best n_neighbors for KNN
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import cross_val_score
#try to plot KNN
#creating list of K for KNN
k_list = list(range(1,50,2))
# create a new list called cv_scores
cv_scores = []
# perform 10-fold cross validation
for k in k_list:
knn = KNeighborsClassifier(n_neighbors=k)
scores = cross_val_score(knn, X_resampled, y_resampled, cv=10, scoring='f1')
cv_scores.append(scores.mean())
# find MSE
MSE = [1 - x for x in cv_scores]
#make a plot
plt.figure()
plt.figure(figsize=(15,10))
plt.title('Identifying the number of neighbors', fontsize=20, fontweight='bold')
plt.xlabel('Number of Neighbors', fontsize=15)
plt.ylabel('MSE', fontsize=15)
sns.set_style("whitegrid")
plt.plot(k_list, MSE)
plt.show()
#This pattern shows that less numbers may reduce MSE
# Delcare the model
rf = RandomForestClassifier(criterion='entropy',
min_samples_leaf=2, min_samples_split=8,
n_estimators=40, n_jobs=1)
# Train the model
# use the fit method
result = rf.fit(X_resampled, y_resampled)
# Print the accuracy
print('Accuracy on train: ' + str(rf.score(X_resampled, y_resampled)), 'Accuracy on test: ' + str(rf.score(X_test, y_test)))
traindataframe = pd.DataFrame(X_resampled, columns=[features])
import matplotlib.pyplot as plt
# Convert the importances into one-dimensional 1darray with corresponding df column names as axis labels
# Implement me
f_importances = pd.Series(rf.feature_importances_, traindataframe.columns)
# Sort the array in descending order of the importances
# Implement me
f_importances = f_importances.sort_values(ascending=False)
# Draw the bar Plot from f_importances
f_importances.plot(x='Features', y='Importance', kind='bar', figsize=(16,9), rot=45, fontsize=20)
# Show the plot
plt.tight_layout()
plt.show()
# models to try
#Logistic Regression
#K Nearest Neighbor
#Perceptron
#Decision Tree
#Support Vector Machine
#Random Forest
#import machine learning packages
from sklearn.linear_model import LogisticRegression
#from sklearn.neighbors import KNeighborsClassifier
from sklearn.neural_network import MLPClassifier
#from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
#from sklearn.ensemble import RandomForestClassifier
#create clfs
clfs = {'lr': LogisticRegression(random_state=0),
'knn':KNeighborsClassifier(),
'mlp': MLPClassifier(random_state=0)}
'rf': RandomForestClassifier(random_state=0),
'svc': SVC(random_state=0)}
#create pipline
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import SelectFromModel
thresholds = [0, .05, .2]
pipe_clfs = {}
for name, clf in clfs.items():
pipe_clfs[name] = {}
for threshold in thresholds:
pipe_clfs[name][threshold] = Pipeline([('StandardScaler', StandardScaler()),
('SelectFromModel', SelectFromModel(estimator=RandomForestClassifier(random_state=0),
threshold=threshold)),
('clf', clf)])
#create paramter grids
param_grids = {}
#Logistic Regression
param_grid = [{'clf__multi_class': ['ovr'],
'clf__solver': ['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga']}]
param_grids['lr'] = param_grid
#KNN
#create parameter grid for KNN
param_grid = [{'clf__n_neighbors': [2,10,11,12, 15, 30, 40],
'clf__algorithm':['auto','brute','kd_tree','ball_tree']}]
param_grids['knn'] = param_grid
#perceptron
param_grid = [{'clf__hidden_layer_sizes': [50, 100, 150],
'clf__activation': ['identity', 'logistic', 'tanh', 'relu']}]
param_grids['mlp'] = param_grid
#create parameter grid for SVC
param_grid = [{'clf__C': [0.1, 1, 10, 20, 50],
'clf__gamma': [0.1, 1, 10, 20, 50],
'clf__kernel': ['rbf', 'sigmoid']}]
param_grids['svc'] = param_grid
#create parameter grid for random forest
param_grid = [{'clf__n_estimators': [5, 10, 20, 30, 40],
'clf__min_samples_split': [10, 15, 20, 30, 40, 50],
'clf__min_samples_leaf': [ 3, 4, 5, 7, 10, 15, 20]}]
param_grids['rf'] = param_grid
# The start time
start = time.time()
# The list of [best_score_, best_params_, best_estimator_]
best_score_param_estimators = []
#scoring = ['accuracy', 'precision', 'recall', 'f1']
# For each classifier
for name in pipe_clfs.keys():
for threshold in thresholds:
# GridSearchCV
gs = GridSearchCV(estimator=pipe_clfs[name][threshold],
scoring='f1',
param_grid=param_grids[name],
n_jobs=1,
cv=StratifiedKFold(n_splits=5,
shuffle=False,
random_state=0))
# Fit the pipeline
gs = gs.fit(X_resampled, y_resampled)
# Update best_score_param_estimators
best_score_param_estimators.append([gs.best_score_, gs.best_params_, gs.best_estimator_])
# The end time
end = time.time()
# Print the Run time
print('Run time: ' + str(end - start))
# Sort best_score_param_estimators in descending order of the best_score_
best_score_param_estimators = sorted(best_score_param_estimators, key=lambda x : x[0], reverse=True)
# Print the best_estimator
print('best_estimator:', end=' ')
print(best_score_param_estimators[0][2].named_steps['clf'], end='\n\n')
# Print the threshold used by the best_estimator
print('threshold:', end=' ')
print(best_score_param_estimators[0][2].named_steps['SelectFromModel'].threshold_)
#print the score
print('score:', end='')
print(best_score_param_estimators[0][0])
print(len(best_score_param_estimators))
#confusion matrix
i = [0,1]
from sklearn.metrics import confusion_matrix
y_pred = best_score_param_estimators[0][2].predict(X_resampled)
cm = confusion_matrix(y_resampled,y_pred)
print(cm)
TN = cm[0][0]
FP = cm[0][1]
FN = cm[1][0]
TP = cm[1][1]
print(TN, FP, FN, TP)
#create recall, precision, F1, Sensitivyt and specificty from matrix
recall = TP/(TP+FN)
precision = TP/(TP+FP)
Sensitivity = TP/(TP+FN)
Specificity = TN/ (TN+FP)
Accuracy = (TN + TN) / (TP + TN + FP + FN)
f1 = 2*((recall*precision)/(precision+recall))
print('recall is', recall)
print('precision is', precision)
print('sensitivity is', Sensitivity)
print('specificity is', Specificity)
print('Accuracy is', Accuracy)
print('f1 is', f1)
#predict target based on best model
y_pred_test = best_score_param_estimators[0][2].predict(X_test)
cm_test = confusion_matrix(y_test,y_pred_test)
print(cm_test)
TrueN = cm_test[0][0]
FalseP = cm_test[0][1]
FalseN = cm_test[1][0]
TrueP = cm_test[1][1]
print(TrueN, FalseP, FalseN, TrueP)
#create recall, precision, F1, Sensitivyt and specificty from matrix
recall_test = TrueP/(TrueP+FalseN)
precision_test = TrueP/(TrueP+FalseP)
Sensitivity_test = TrueP/(TrueP+FalseN)
Specificity_test = TrueN/ (TrueN+FalseP)
Accuracy = (TrueP + TrueN) / (TrueP + TrueN + FalseP + FalseN)
f1 = 2*((recall*precision)/(precision+recall))
print('recall is', recall_test)
print('precision is', precision_test)
print('sensitivity is', Sensitivity_test)
print('specificity is', Specificity_test)
print('Accuracy is', Accuracy)
print('f1 is', f1)
```
| github_jupyter |
This tutorial guides you on how to generate text using a character-based RNN model. This model helps to generate a character one by one and a longer sequence can execute it repeatedly. **However, such a model can't understand the meanings of word, how to spell them as well, and what a role plays in a sentence.**
On the contrary, this model is simple to capture the details on the character levels, such as capitalization, the conversation format, etc.
```
!pip install -q tf-nightly
import tensorflow as tf
import numpy as np
import os
import time
print("Tensorflow Version: {}".format(tf.__version__))
print("GPU {} available.".format("is" if tf.config.experimental.list_physical_devices("GPU") else "not"))
```
# Data Preprocessing
Here we are going to use the dataset of Shakespeare's writing from [The Unreasonable Effectiveness of Recurrent Neural Networks](http://karpathy.github.io/2015/05/21/rnn-effectiveness/).
```
path_to_file = tf.keras.utils.get_file('shakespeare.txt',
'https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt')
path_to_file
```
## Read the Data
```
text = open(path_to_file, 'rb').read().decode('UTF-8')
print("Length of text: {} characters".format(len(text)))
```
Let's take a look at first few characters.
```
print(text[:200])
```
Let's calculate the unique characters.
```
vocab = sorted(set(text))
print("Unique characters: {}".format(len(vocab)))
```
## Vectorize the text
Before training, we have to convert the strings to the numerical representation.
```
char2idx = {u:i for i, u in enumerate(vocab)}
idx2char = np.array(vocab)
# transform the text with integers
text_as_int = [char2idx[c] for c in text]
for char, _ in zip(char2idx, range(20)):
print('{:4s}: {:3d}'.format(repr(char), char2idx[char]))
```
Show the first few characters with their mapping integers.
```
print("{} <---> {}".format(repr(text[:20]), text_as_int[:20]))
```
## Prediction
The purpose of the prediction is to predict the character given an input one. We are going to train a model for the above purpose. The input of the model is a sequence of characters and the prediction is the next character of the same sequence. The attributes of RNNs are memorizing the previously seen data, a stateful concept. The prediction would be what the next character is, given all the characters until this moment.
### Training Examples and Targets
The basic operation idea is to break down the same text into the inputs and outputs. The output is shifted forward one character of the input. For example, if the `sequence_len` is 4 and the sentence is `hello`, the input sequence becomes `hell` and the output (or target) one becomes `ello`.
```
seq_length = 100
examples_per_epoch = len(text) // (seq_length + 1)
char_dataset = tf.data.Dataset.from_tensor_slices(text_as_int)
for c in char_dataset.take(10):
print(idx2char[c.numpy()], end=" ")
```
Here we use `batch()` to split the characters to sequences of the desired size.
```
sequences = char_dataset.batch(seq_length + 1, drop_remainder=True)
for seq in sequences.take(5):
print(repr(''.join(idx2char[seq.numpy()])))
```
For each sequence, we need to split it into the training and the target sequence.
```
def split_input_target(data):
inputs = data[:-1]
target = data[1:]
return inputs, target
dataset = sequences.map(split_input_target)
for input_example, target_example in dataset.take(1):
print(repr(''.join(idx2char[input_example.numpy()])))
print(repr(''.join(idx2char[target_example.numpy()])))
```
Each index of these sequence vectors represents one time step. For the input character at time 0 is `F` and the target to be predicted is `i`. The RNN model considers the previous step status in addition to the current input character.
```
for i, (input_idx, target_idx) in enumerate(zip(input_example[:5], target_example[:5])):
print("Step {}: input {:s}, target {:s}".format(
i, repr(idx2char[input_idx.numpy()]), repr(idx2char[target_idx.numpy()])))
```
### Creating Training Batches
```
BATCH_SIZE = 64
BUFFER_SIZE = 10000
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
dataset
```
# Build the Model
```
vocab_size = len(vocab)
embedding_dim = 256
rnn_units = 1024 # for gru cell
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
def _model(inputs):
embed = tf.keras.layers.Embedding(input_dim=vocab_size, output_dim=embedding_dim)(inputs) # [batch_size, None, 256]
# set `return_sequences=True` for predicting the next character
gru = tf.keras.layers.GRU(units=rnn_units, return_sequences=True, stateful=True,
recurrent_initializer='glorot_uniform')(embed) # [None, None, 1024]
outputs = tf.keras.layers.Dense(units=vocan_size)(gru)
return outputs
# if the stateful is True, it requires the batch size
inputs = tf.keras.Input(shape=(None, ), batch_size=batch_size)
generator = _model(inputs)
return tf.keras.Model(inputs, generator)
model = build_model(vocab_size=vocab_size, embedding_dim=embedding_dim,
rnn_units=rnn_units, batch_size=BATCH_SIZE)
model.summary()
```
The above model can be translated as the below model architecture.

Refer to Tensorflow.org (2020).
# Try the Model
Let's try the model first to check the result.
```
for input_example_batch, target_example_batch in dataset.take(1):
prediction_example_batch = model(input_example_batch)
print(prediction_example_batch.shape) # [batch_size, sequence_length, vocab_size]
```
To get the actual predictions from the model, we need to sample from the output distribution to get the actual indices.
```
sampled_indices = tf.random.categorical(logits=prediction_example_batch[0], num_samples=1)
sampled_indices = tf.squeeze(sampled_indices, axis=-1).numpy()
sampled_indices
print("Input: {}".format(repr(''.join(idx2char[input_example_batch[0].numpy()]))))
print("Output: {}".format(repr(''.join(idx2char[sampled_indices]))))
```
# Train the Model
Before training, we have to decide how to train the model. To this problem, we can regard it as a classification problem. Given the previous RNN state, and the input this timestamp, predict the class of the next character.
## Attach an Optimizer, and Define a Loss Function
The `tf.keras.losses.sparse_categorical_crossentropy` API works in this case because it is applied across the last dimension of the predictions. For example, the target shape is `[batch_size, sequence_length]` and the prediction shape is `[batch_size, sequence_length, vocab_size]`. This API helps you calculate the loss alongside the last dimension of the predictions.
```
def loss(labels, logits):
return tf.reduce_mean(
tf.keras.losses.sparse_categorical_crossentropy(labels, logits, from_logits=True))
example_batch_loss = loss(target_example_batch, prediction_example_batch)
print("Prediction Shape: {}".format(prediction_example_batch.shape))
print("Scalar Loss: {}".format(example_batch_loss.numpy().mean()))
print("Target shapes: {}".format(target_example_batch.shape))
print("Prediction shape: {}".format(prediction_example_batch.shape))
def accuracy(labels, logits):
logits = tf.cast(tf.argmax(logits, axis=-1), tf.int32)
labels = tf.cast(labels, tf.int32)
return tf.reduce_mean(tf.keras.metrics.binary_accuracy(labels, logits))
accuracy(target_example_batch, prediction_example_batch)
model.compile(loss=loss, optimizer=tf.keras.optimizers.Adam(), metrics=[accuracy])
```
## Configure the Checkpoints
```
ckpt_dir = "./train_ckpt"
ckpt_prefix = os.path.join(ckpt_dir, 'ckpt_{epoch}')
ckpt_callbacks = tf.keras.callbacks.ModelCheckpoint(
filepath=ckpt_prefix, save_weights_only=True
)
```
## Start Training
```
history = model.fit(dataset, epochs=10, callbacks=[ckpt_callbacks])
```
# Generate Text
## Restore the Latest Checkpoint
To keep the prediction simple, use a batch size of 1. Because of the way the RNN state passed from timestamp to timestamp (stateful=True), the model only accepts the same batch size once it was built.
```
latest_ckpt = tf.train.latest_checkpoint(ckpt_dir)
latest_ckpt
loaded = build_model(vocab_size, embedding_dim, rnn_units, 1)
loaded.load_weights(latest_ckpt)
# the batch size was not reset
loaded.summary()
```
## The Prediction Loop

Refer to Tensorflow.org (2020).
```
def generate_text(model, start_string):
# the number of characters to generate
num_generate = 1000
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, axis=0)
# for the generated characters
text_generated = []
# low temperature results in more predictable characters
# high temperature results in more surprising characters
# this value requires experimenting
temperature = 1.0
# now let's reset state for the first character
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
# remove the batch dimension
predictions = tf.squeeze(predictions, axis=0)
# using a categorical distribution to predict the character
predictions = predictions / temperature
# [-1, 0]: the output of the final character, select the first character
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1, 0].numpy()
text_generated.append(idx2char[predicted_id])
input_eval = tf.expand_dims([predicted_id], axis=0)
return start_string + ''.join(text_generated)
print(generate_text(loaded, start_string=u"ROMEO: "))
```
You can train the model with more epochs, stack RNN layers to the model, or change the temperature value to check the model performance.
# Customized Training
These advanced operations help you in controlling the training step and help stabilize the model's open-loop output.
```
adv_model = build_model(vocab_size, embedding_dim, rnn_units, BATCH_SIZE)
optimizer = tf.keras.optimizers.Adam()
@tf.function
def train_step(inputs, targets):
with tf.GradientTape() as tape:
predictions = adv_model(inputs)
loss = tf.reduce_mean(
tf.keras.losses.sparse_categorical_crossentropy(targets, predictions, from_logits=True)
)
grads = tape.gradient(loss, adv_model.trainable_variables)
optimizer.apply_gradients(zip(grads, adv_model.trainable_variables))
return loss
def train(epochs=10):
for epoch in range(epochs):
# initializing the hidden state at the beginning of every epoch
# initial hidden state is None
hidden = adv_model.reset_states()
for (batch_n, (inputs, targets)) in enumerate(dataset):
loss = train_step(inputs, targets)
if batch_n % 100 == 0:
template = "Epoch {} Batch {} Loss {}"
print(template.format(epoch, batch_n, loss))
# save the checkpoints
if (epoch + 1) % 5 == 0:
adv_model.save_weights(ckpt_prefix.format(epoch=epoch))
print("Epoch {} Loss {:.4f}".format(epoch + 1, loss))
adv_model.save_weights(ckpt_prefix.format(epoch=epoch))
train(epochs=10)
```
| github_jupyter |
# 化合物表示学习和性质预测
在这篇教程中,我们将介绍如何运用图神经网络(GNN)模型来预测化合物的性质。具体来说,我们将演示如何对其进行预训练(pretrain),如何针对下游任务进行模型微调(finetune),并利用最终的模型进行推断(inference)。如果你想了解更多细节,请查阅 "[info graph](https://github.com/PaddlePaddle/PaddleHelix/apps/pretrained_compound/info_graph/README_cn.md)" 和 "[pretrained GNN](https://github.com/PaddlePaddle/PaddleHelix/apps/pretrained_compound/pretrain_gnns/README_cn.md)" 的详细解释.
# 第一部分:预训练
在这一部分,我们将展示如何预训练一个化合物 GNN 模型。本文中的预训练技术是在预训练 GNN 的基础上发展起来的,包括 attribute masking、context prediction 和有监督预训练。
更多细节请查看文件:`pretrain_attrmask.py`,`pretrain_contextpred.py` 和 `pretrain_supervised.py`。
```
import os
import numpy as np
import sys
sys.path.insert(0, os.getcwd() + "/..")
os.chdir("../apps/pretrained_compound/pretrain_gnns")
```
PaddleHelix 是构建于 PaddlePaddle 之上的生物计算深度学习框架。
```
import paddle
import paddle.fluid as fluid
from paddle.fluid.incubate.fleet.collective import fleet
from pahelix.datasets import load_zinc_dataset
from pahelix.featurizers import PreGNNAttrMaskFeaturizer
from pahelix.utils.compound_tools import CompoundConstants
from pahelix.model_zoo import PreGNNAttrmaskModel
# switch to paddle static graph mode.
paddle.enable_static()
```
## 构建静态图
通常情况下,我们用 Paddle 提供的 `Program` 和 `Executor` 来构建静态图。这里,我们使用 `model_config` 保存模型配置。`PreGNNAttrmaskModel` 是一种无监督的预训练模型,它随机地对某个节点的原子类型进行 mask,然后再尝试去预测这个原子的类型。同时,我们使用 Adam 优化器并将学习率(learning rate)设置为 0.001。
若要使用GPU进行训练,请取消注释行 `exe = fluid.Executor(fluid.CUDAPlace(0))`。当 `fluid.CPUPlace()` 用于CPU训练。
```
model_config = {
"dropout_rate": 0.5,# dropout rate
"gnn_type": "gin", # other choices like "gat", "gcn".
"layer_num": 5, # the number of gnn layers.
}
train_prog = fluid.Program()
startup_prog = fluid.Program()
with fluid.program_guard(train_prog, startup_prog):
with fluid.unique_name.guard():
model = PreGNNAttrmaskModel(model_config=model_config)
model.forward()
opt = fluid.optimizer.Adam(learning_rate=0.001)
opt.minimize(model.loss)
exe = fluid.Executor(fluid.CPUPlace())
# exe = fluid.Executor(fluid.CUDAPlace(0))
exe.run(startup_prog)
print(model.loss)
```
## 数据集加载和特征提取
我们首先使用 `wget` 来下载一个小型的测试数据集,如果你的本地计算机上没有 `wget`,你也可以复制下面的链接到你的浏览器中来下载数据。但是请注意你需要把数据包移动到这个路径:"../apps/pretrained_compound/pretrain_gnns/"。
```
### Download a toy dataset for demonstration:
!wget "https://baidu-nlp.bj.bcebos.com/PaddleHelix%2Fdatasets%2Fcompound_datasets%2Fchem_dataset_small.tgz" --no-check-certificate
!tar -zxf "PaddleHelix%2Fdatasets%2Fcompound_datasets%2Fchem_dataset_small.tgz"
!ls "./chem_dataset_small"
### Download the full dataset as you want:
# !wget "http://snap.stanford.edu/gnn-pretrain/data/chem_dataset.zip" --no-check-certificate
# !unzip "chem_dataset.zip"
# !ls "./chem_dataset"
```
使用 `PreGNNAttrMaskFeaturizer` 来配合模型 `PreGNNAttrmaskModel`。它继承了用于特征提取的超类 `Featurizer`。`Featurizer` 有两个功能:`gen_features` 用于将一条原始 SMILES 转换为图数据,而 `collate_fn` 用于将图数据的子列表聚合为一个 batch。这里我们采用 Zinc 数据集来进行预训练。
```
featurizer = PreGNNAttrMaskFeaturizer(
model.graph_wrapper,
atom_type_num=len(CompoundConstants.atom_num_list),
mask_ratio=0.15)
### Load the first 1000 of the toy dataset for speed up
dataset = load_zinc_dataset("./chem_dataset_small/zinc_standard_agent/raw", featurizer=featurizer)
dataset = dataset[:1000]
### Load the full dataset:
# dataset = load_zinc_dataset("./chem_dataset/zinc_standard_agent/raw", featurizer=featurizer)
print("dataset num: %s" % (len(dataset)))
```
## 启动训练
现在我们开始训练 Attrmask 模型。我们仅训练两个 epoch 作为演示,数据加载的过程通过4个 `workers` 进行了加速。然后我们将预训练后的模型保存到 "./model/pretrain_attrmask",作为下游任务的初始模型。
```
def train(exe, train_prog, model, dataset, featurizer):
data_gen = dataset.iter_batch(
batch_size=256, num_workers=4, shuffle=True, collate_fn=featurizer.collate_fn)
list_loss = []
for batch_id, feed_dict in enumerate(data_gen):
train_loss, = exe.run(train_prog,
feed=feed_dict, fetch_list=[model.loss], return_numpy=False)
list_loss.append(np.array(train_loss).mean())
return np.mean(list_loss)
for epoch_id in range(2):
train_loss = train(exe, train_prog, model, dataset, featurizer)
print("epoch:%d train/loss:%s" % (epoch_id, train_loss))
fluid.io.save_params(exe, './model/pretrain_attrmask', train_prog)
```
模型预训练的内容到此为止,你可以根据自己的需要对上面的参数进行调整。
# 第二部分:下游任务模型微调(fintune)
下面我们将介绍如何对预训练的模型进行微调来适应下游任务。
更多细节参见 `finetune.py` 文件中的内容。
```
from pahelix.utils.paddle_utils import load_partial_params
from pahelix.utils.splitters import \
RandomSplitter, IndexSplitter, ScaffoldSplitter, RandomScaffoldSplitter
from pahelix.datasets import *
from model import DownstreamModel
from featurizer import DownstreamFeaturizer
from utils import calc_rocauc_score
```
下游任务的数据集通常规模很小,并且面向不同的任务。例如,BBBP 数据集用于预测化合物的血脑屏障通透性;Tox21 数据集用于预测化合物的毒性等。这里我们使用 Tox21 数据集进行演示。
```
task_names = get_default_tox21_task_names()
print(task_names)
```
## 构建静态图
这里我们采用和之前一样的方式构建一个静态图模型。注意这里的模型结构的设置应该和预训练模型中的设置保持一致,否则模型加载将会失败。`DownstreamModel` 是一个有监督的 GNN 模型,用于上述 `task_names` 中定义的预测任务。
我们使用 `train_prog` 和 `test_prog` 来保存静态图,用于后续的训练和测试。它们具有相同的网络架构,但某些操作符的功能将发生变化,例如 `Dropout` 和 `BatchNorm`。
```
model_config = {
"dropout_rate": 0.5,# dropout rate
"gnn_type": "gin", # other choices like "gat", "gcn".
"layer_num": 5, # the number of gnn layers.
"num_tasks": len(task_names), # number of targets to predict for the downstream task.
}
train_prog = fluid.Program()
test_prog = fluid.Program()
startup_prog = fluid.Program()
with fluid.program_guard(train_prog, startup_prog):
with fluid.unique_name.guard():
model = DownstreamModel(model_config=model_config)
model.train()
opt = fluid.optimizer.Adam(learning_rate=0.001)
opt.minimize(model.loss)
with fluid.program_guard(test_prog, fluid.Program()):
with fluid.unique_name.guard():
model = DownstreamModel(model_config=model_config)
model.train(is_test=True)
exe = fluid.Executor(fluid.CPUPlace())
# exe = fluid.Executor(fluid.CUDAPlace(0))
exe.run(startup_prog)
```
## 加载预训练模型
加载预训练阶段得到的模型。这里我们加载模型 "pretrain_attrmask" 作为一个例子。
```
load_partial_params(exe, './model/pretrain_attrmask', train_prog)
```
## 数据加载和特征提取
将 `DownstreamFeaturizer` 与 `DownstreamModel` 一起使用。它继承自用于特征提取的超类 `featureizer`。`featureizer` 有两个功能:`gen_features` 用于将一条原始 SMILES 转换为单个图数据,而 `collate_fn` 用于将图数据的子列表聚合为一个 batch。
Tox21 数据集用作下游任务数据集,我们使用 `ScaffoldSplitter` 将数据集拆分为训练/验证/测试集。`ScaffoldSplitter` 首先根据 Bemis-Murcko scaffold 对化合物进行排序,然后从前到后,将参数 `frac_train` 定义的比例的数据作为训练集,将 `frac_valid` 定义的比例的数据作为验证集,其余的作为测试集。`ScaffoldSplitter` 能更好地评价模型对非同分布样本的泛化能力。这里也可以使用其他的拆分器,如 `RandomSplitter`、`RandomScaffoldSplitter` 和 `IndexSplitter`。
```
featurizer = DownstreamFeaturizer(model.graph_wrapper)
### Load the toy dataset:
dataset = load_tox21_dataset(
"./chem_dataset_small/tox21/raw", task_names, featurizer=featurizer)
### Load the full dataset:
# dataset = load_tox21_dataset(
# "./chem_dataset/tox21/raw", task_names, featurizer=featurizer)
# splitter = RandomSplitter()
splitter = ScaffoldSplitter()
train_dataset, valid_dataset, test_dataset = splitter.split(
dataset, frac_train=0.8, frac_valid=0.1, frac_test=0.1)
print("Train/Valid/Test num: %s/%s/%s" % (
len(train_dataset), len(valid_dataset), len(test_dataset)))
```
## 启动训练
出于演示的目的,这里我们只将 attrmask 模型训练了4轮。由于每个下游任务都包含了多个子任务,我们分别计算了每个子任务的 roc-auc,在求其均值作为最后的评估标准。
```
def train(exe, train_prog, model, train_dataset, featurizer):
data_gen = train_dataset.iter_batch(
batch_size=64, num_workers=4, shuffle=True, collate_fn=featurizer.collate_fn)
list_loss = []
for batch_id, feed_dict in enumerate(data_gen):
train_loss, = exe.run(train_prog, feed=feed_dict, fetch_list=[model.loss], return_numpy=False)
list_loss.append(np.array(train_loss).mean())
return np.mean(list_loss)
def evaluate(exe, test_prog, model, test_dataset, featurizer):
"""
In the dataset, a proportion of labels are blank. So we use a `valid` tensor
to help eliminate these blank labels in both training and evaluation phase.
Returns:
the average roc-auc of all sub-tasks.
"""
data_gen = test_dataset.iter_batch(
batch_size=64, num_workers=4, shuffle=False, collate_fn=featurizer.collate_fn)
total_pred = []
total_label = []
total_valid = []
for batch_id, feed_dict in enumerate(data_gen):
pred, = exe.run(test_prog, feed=feed_dict, fetch_list=[model.pred], return_numpy=False)
total_pred.append(np.array(pred))
total_label.append(feed_dict['finetune_label'])
total_valid.append(feed_dict['valid'])
total_pred = np.concatenate(total_pred, 0)
total_label = np.concatenate(total_label, 0)
total_valid = np.concatenate(total_valid, 0)
return calc_rocauc_score(total_label, total_pred, total_valid)
for epoch_id in range(4):
train_loss = train(exe, train_prog, model, train_dataset, featurizer)
val_auc = evaluate(exe, test_prog, model, valid_dataset, featurizer)
test_auc = evaluate(exe, test_prog, model, test_dataset, featurizer)
print("epoch:%s train/loss:%s" % (epoch_id, train_loss))
print("epoch:%s val/auc:%s" % (epoch_id, val_auc))
print("epoch:%s test/auc:%s" % (epoch_id, test_auc))
fluid.io.save_params(exe, './model/tox21', train_prog)
```
# 第三部分:下游任务模型预测
在这部分,我们将简单介绍如何利用训好的下游任务模型来对给定的 SMILES 序列做预测。
## 构建静态图
这部分跟第二部分的建图部分基本相同。
```
model_config = {
"dropout_rate": 0.5,# dropout rate
"gnn_type": "gin", # other choices like "gat", "gcn".
"layer_num": 5, # the number of gnn layers.
"num_tasks": len(task_names), # number of targets to predict for the downstream task.
}
inference_prog = fluid.Program()
startup_prog = fluid.Program()
with fluid.program_guard(inference_prog, startup_prog):
with fluid.unique_name.guard():
model = DownstreamModel(model_config=model_config)
model.inference()
exe = fluid.Executor(fluid.CPUPlace())
# exe = fluid.Executor(fluid.CUDAPlace(0))
exe.run(startup_prog)
```
## 加载训练好的下游任务模型
加载在第二部分中训练好的下游任务模型。
```
load_partial_params(exe, './model/tox21', inference_prog)
```
## 开始预测
对给定的 SMILES 序列进行预测。我们直接调用 `DownstreamFeaturizer` 的 `gen_features` 和 `collate_fn` 函数将原始的 SMILES 序列转化为模型的输入。
以 Tox21 数据集为例,我们的下游任务模型可以给出 Tox21 里面的12个子任务的预测。
```
SMILES="O=C1c2ccccc2C(=O)C1c1ccc2cc(S(=O)(=O)[O-])cc(S(=O)(=O)[O-])c2n1"
featurizer = DownstreamFeaturizer(model.graph_wrapper, is_inference=True)
feed_dict = featurizer.collate_fn([featurizer.gen_features({'smiles': SMILES})])
pred, = exe.run(inference_prog, feed=feed_dict, fetch_list=[model.pred], return_numpy=False)
probs = np.array(pred)[0]
print('SMILES:%s' % SMILES)
print('Predictions:')
for name, prob in zip(task_names, probs):
print(" %s:\t%s" % (name, prob))
```
| github_jupyter |
# 循环神经网络
### 从零开始实现
```
import torch
import torch.nn as nn
import torch.nn.functional as F
X, W_xh = torch.randn(1, 5), torch.randn(5, 4) #X为5维数据,时序为1
H, W_hh = torch.randn(1, 4), torch.randn(4, 4)
b = torch.ones(1,1)
h_t = F.relu(torch.matmul(X, W_xh) + torch.matmul(H, W_hh) + b)
h_t
```
### Pytorch模块定义
```
class RnnNet(nn.Module):
def __init__(self, dim_input, dim_hidden, dim_output):
super(RnnNet, self).__init__()
self.fc_x2h = nn.Linear(dim_input, dim_hidden)
self.fc_h2h = nn.Linear(dim_hidden, dim_hidden, bias = False)
self.fc_h2y = nn.Linear(dim_hidden, dim_output) #4x1
self.dim_hidden = dim_hidden
def forward(self, x):
h = x.new_zeros(1, self.dim_hidden)
for t in range(x.size(0)):
h = F.relu(self.fc_x2h(x[t:t+1]) + self.fc_h2h(h)) #
return self.fc_h2y(h)
rnn = RnnNet(5, 20, 10)
t = torch.randn(20, 5) #时序长为20
rnn(t)
rnn = nn.RNN(10, 20, 2, batch_first=True) # inputsize, hidden size, num_layers
input = torch.randn(3, 5, 10) # batchsize 3 时序长度为5, 10:特征维度
h0 = torch.randn(2, 3, 20) # 层数,batchsize, hiddensize
output, hn = rnn(input, h0) #
print(hn.shape)
output.shape # W_hy x H 输出size默认为h的维度
```
### 使用RNNCell进行单个样本运算
```
rnn = nn.RNNCell(10, 20)
input = torch.randn(6, 3, 10) # (time_steps, batch, input_size)
hx = torch.randn(3, 20)
output = []
for i in range(input.size(0)):
hx = rnn(input[i], hx)
output.append(hx)
output[0].shape
```
### gating RNN
```
class RecNetWithGating(nn.Module):
def __init__(self, dim_input, dim_recurrent, dim_output):
super(RecNetWithGating, self).__init__()
self.fc_x2h = nn.Linear(dim_input, dim_recurrent)
self.fc_h2h = nn.Linear(dim_recurrent, dim_recurrent, bias = False)
self.fc_x2z = nn.Linear(dim_input, dim_recurrent)
self.fc_h2z = nn.Linear(dim_recurrent, dim_recurrent, bias = False)
self.fc_h2y = nn.Linear(dim_recurrent, dim_output)
self.dim_hidden = dim_recurrent
def forward(self, input):
h = input.new_zeros(1, self.dim_hidden)
for t in range(input.size(0)):
z = torch.sigmoid(self.fc_x2z(input[t:t+1]) + self.fc_h2z(h))
hb = F.relu(self.fc_x2h(input[t:t+1]) + self.fc_h2h(h))
h = z * h + (1 - z) * hb
return self.fc_h2y(h)
rnn = RecNetWithGating(5, 4, 4)
t = torch.randn(20, 5) #时序长为20
rnn(t)
```
### Pytorch LSTM
```
lstm = nn.LSTMCell(10, 20) #input_dim, recurrent dim
input = torch.randn(2, 3, 10) # (time_steps, batch, input_size)
hx = torch.randn(3, 20) # (batch, hidden_size)
cx = torch.randn(3, 20)
output = []
for i in range(input.size()[0]):
hx, cx = lstm(input[i], (hx, cx)) # 每次输入一个时间样本
output.append(hx)
output = torch.stack(output, dim=0)
output.shape
class lstmNet(nn.Module):
def __init__(self, dim_input, dim_recurrent, num_layers, dim_output):
super(lstmNet, self).__init__()
self.lstm = nn.LSTM(dim_input, dim_recurrent, num_layers)
self.fc = nn.Linear(dim_recurrent, dim_output)
def forward(self, x):
hx, cx = self.lstm(x)
o = hx[-1,:,:]
o = o.squeeze(axis=0)
return self.fc(o)
input = torch.randn(2, 3, 10) #T N C
lstm = lstmNet(10, 20, 1, 7)
output = lstm(input)
output.shape
```
### GRU
```
class gruNet(nn.Module):
def __init__(self, dim_input, dim_recurrent, num_layers, dim_output):
super(gruNet, self).__init__()
self.gru = nn.GRU(dim_input, dim_recurrent, num_layers)
self.fc = nn.Linear(dim_recurrent, dim_output)
def forward(self, x):
hx, cx = self.gru(x)
o = hx[-1,:,:]
o = o.squeeze(axis=0)
return self.fc(o)
input = torch.randn(2, 3, 10) #T N C
gru = gruNet(10, 20, 1, 7)
output = gru(input)
output.shape
```
### IMDB 文本情感分类
```
import torch
from torch.utils.data import DataLoader,Dataset
import os
import re
# 路径需要根据情况修改,文件太大的时候可以引用绝对路径
data_base_path = r"F:\SZTU-教学文件\UG-深度学习方法与应用\examples\data\aclImdb_v1\aclImdb"
#1. 定义tokenize的方法,对评论文本分词
def tokenize(text):
# fileters = '!"#$%&()*+,-./:;<=>?@[\\]^_`{|}~\t\n'
fileters = ['!','"','#','$','%','&','\(','\)','\*','\+',',','-','\.','/',':',';','<','=','>','\?','@'
,'\[','\\','\]','^','_','`','\{','\|','\}','~','\t','\n','\x97','\x96','”','“',]
# sub方法是替换
text = re.sub("<.*?>"," ",text,flags=re.S)# 去掉<...>中间的内容,主要是文本内容中存在<br/>等内容
text = re.sub("|".join(fileters)," ",text,flags=re.S)# 替换掉特殊字符,'|'是把所有要匹配的特殊字符连在一起
return [i.strip() for i in text.split()]# 去掉前后多余的空格
#2. 准备dataset
class ImdbDataset(Dataset):
def __init__(self,mode):
super(ImdbDataset,self).__init__()
# 读取所有的训练文件夹名称
if mode=="train":
text_path = [os.path.join(data_base_path,i) for i in ["train/neg","train/pos"]]
else:
text_path = [os.path.join(data_base_path,i) for i in ["test/neg","test/pos"]]
self.total_file_path_list = []
# 进一步获取所有文件的名称
for i in text_path:
self.total_file_path_list.extend([os.path.join(i,j) for j in os.listdir(i)])
def __getitem__(self, idx):
cur_path = self.total_file_path_list[idx]
cur_filename = os.path.basename(cur_path)
# 标题的形式是:3_4.txt 前面的3是索引,后面的4是分类
# 原本的分类是1-10,现在变为0-9
label = int(cur_filename.split("_")[-1].split(".")[0]) -1 #处理标题,获取label,-1是因为要转化为[0-9]
text = tokenize(open(cur_path).read().strip()) #直接按照空格进行分词
return label,text
def __len__(self):
return len(self.total_file_path_list)
# 测试是否能成功获取数据
dataset = ImdbDataset(mode="train")
print(dataset[0])
# Word2Sequence
class Word2Sequence:
# 未出现过的词
UNK_TAG = "UNK"
PAD_TAG = "PAD"
# 填充的词
UNK = 0
PAD = 1
def __init__(self):
self.dict = {
self.UNK_TAG: self.UNK,
self.PAD_TAG: self.PAD
}
self.count = {}
def to_index(self, word):
"""word -> index"""
return self.dict.get(word, self.UNK)
def to_word(self, index):
"""index -> word"""
if index in self.inversed_dict:
return self.inversed_dict[index]
return self.UNK_TAG
def __len__(self):
return len(self.dict)
def fit(self, sentence):
"""count字典中存储每个单词出现的次数"""
for word in sentence:
self.count[word] = self.count.get(word, 0) + 1
def build_vocab(self, min_count=None, max_count=None, max_feature=None):
"""
构建词典
只筛选出现次数在[min_count,max_count]之间的词
词典最大的容纳的词为max_feature,按照出现次数降序排序,要是max_feature有规定,出现频率很低的词就被舍弃了
"""
if min_count is not None:
self.count = {word: count for word, count in self.count.items() if count >= min_count}
if max_count is not None:
self.count = {word: count for word, count in self.count.items() if count <= max_count}
if max_feature is not None:
self.count = dict(sorted(self.count.items(), lambda x: x[-1], reverse=True)[:max_feature])
# 给词典中每个词分配一个数字ID
for word in self.count:
self.dict[word] = len(self.dict)
# 构建一个数字映射到单词的词典,方法反向转换,但程序中用不太到
self.inversed_dict = dict(zip(self.dict.values(), self.dict.keys()))
def transform(self, sentence, max_len=None):
"""
根据词典给每个词分配的数字ID,将给定的sentence(字符串序列)转换为数字序列
max_len:统一文本的单词个数
"""
if max_len is not None:
r = [self.PAD] * max_len
else:
r = [self.PAD] * len(sentence)
# 截断文本
if max_len is not None and len(sentence) > max_len:
sentence = sentence[:max_len]
for index, word in enumerate(sentence):
r[index] = self.to_index(word)
return np.array(r, dtype=np.int64)
def inverse_transform(self, indices):
"""数字序列-->单词序列"""
sentence = []
for i in indices:
word = self.to_word(i)
sentence.append(word)
return sentence
# 自定义的collate_fn方法
def collate_fn(batch):
# 手动zip操作,并转换为list,否则无法获取文本和标签了
batch = list(zip(*batch))
labels = torch.tensor(batch[0], dtype=torch.int32)
texts = batch[1]
texts = torch.tensor([ws.transform(i, max_len) for i in texts])
del batch
# 注意这里long()不可少,否则会报错
return labels.long(), texts.long()
train_batch_size = 64
test_batch_size = 500
max_len = 50
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
def get_dataloader(train=True):
if train:
mode = 'train'
else:
mode = "test"
dataset = ImdbDataset(mode)
batch_size = train_batch_size if train else test_batch_size
return DataLoader(dataset, batch_size=batch_size, shuffle=True, collate_fn=collate_fn)
dataloader = get_dataloader()
for idx,(label,text) in enumerate(dataloader):
print("idx:",idx)
print("label:",label)
print("text:",text)
# 建立词表
def fit_save_word_sequence():
word_to_sequence = Word2Sequence()
train_path = [os.path.join(data_base_path, i) for i in ["train/neg", "train/pos"]]
# total_file_path_list存储总的需要读取的txt文件
total_file_path_list = []
for i in train_path:
total_file_path_list.extend([os.path.join(i, j) for j in os.listdir(i)])
# tqdm是显示进度条的
for cur_path in tqdm(total_file_path_list, ascii=True, desc="fitting"):
word_to_sequence.fit(tokenize(open(cur_path, encoding="utf-8").read().strip()))
word_to_sequence.build_vocab()
# 对wordSequesnce进行保存
pickle.dump(word_to_sequence, open("model/ws.pkl", "wb"))
ws = pickle.load(open("./model/ws.pkl", "rb"))
```
| github_jupyter |
```
import numpy as np
import xray
import dask.array as daskarray
from matplotlib import pyplot as plt
%matplotlib inline
import resource
resource.setrlimit(resource.RLIMIT_NOFILE, (4096 ,4096 ))
resource.getrlimit(resource.RLIMIT_NOFILE)
import xgcm
iters = range(2073840, 2384880, 480)
ddir = '/data/scratch/rpa/channel_moc/GCM/run'
ds = xgcm.open_mdsdataset(ddir, iters, deltaT=900)
ds
def pow_spec_2d(x, axis=[-1,-2]):
# x and time
f = np.fft.fftn(x, axes=axis)
return np.real(f * f.conj())
# operate on a dataarray
def xray_isotropic_power_spectrum(da, kiso=None, axis=('Y','X')):
axis_num = [da.get_axis_num(a) for a in axis]
# see if there are other coords to leave alone
other_axis = set(da.dims).difference(axis)
if other_axis:
other_axis_num = [da.get_axis_num for a in other_axis]
# calcualte wavenumbers from axes
N = [da.shape[n] for n in axis_num]
delta_x = [np.diff(da[a])[0] for a in axis]
k = [ np.fft.fftfreq(Nx, dx) for (Nx, dx) in zip(N, delta_x)]
kk = np.array(np.meshgrid(k[1], k[0]))
k2 = (kk**2).sum(axis=0)
# set up wavenumber range
if kiso is None:
# no isotropic wavenumber grid specified
kidx = np.argmin( np.array([l.max() for l in k]) )
kiso = k[kidx][:N[kidx]/2]
Niso = len(kiso)
bins = np.digitize(k2.ravel(), kiso**2)
# do fft
#def iso_ps(q):
f = np.fft.fftn(da.values, axes=axis_num)
# sum isotropically
fiso = np.bincount(bins,
weights=np.real(f*f.conj()).ravel(),
minlength=Niso)[:Niso]
# replace zeros with nans
fiso[fiso==0.] = np.nan
# normalize properly
#count = np.bincount(bins, minlength=Niso)[:Niso]
#fiso *= (kiso / count)
# (didn't work like I hoped)
return xray.DataArray(fiso, coords={'k_iso': kiso})
def ts_iso_power_spec(sst):
kiso = np.linspace(0,(5000.)**-1 / np.sqrt(2), 100)
xvars = set(['X', 'Xp1'])
yvars = set(['Y', 'Yp1'])
xvar = xvars.intersection(sst.dims).pop()
yvar = yvars.intersection(sst.dims).pop()
ssta = sst - sst.mean(dim=xvar)
hwin = np.hanning(ds[yvar].shape[0])
window = xray.DataArray(daskarray.from_array(hwin, hwin.shape), coords={yvar: ds[yvar]})
sst_win = ssta * window
return sst_win.groupby('time').apply( lambda q: xray_isotropic_power_spectrum(q, kiso=kiso, axis=(yvar,xvar)) )
%time t_ps = ts_iso_power_spec(ds['THETA'].isel(Z=0))
%time u_ps = ts_iso_power_spec(ds['UVEL'].isel(Z=0))
%time v_ps = ts_iso_power_spec(ds['VVEL'].isel(Z=0))
%time w_ps = ts_iso_power_spec(ds['WVEL'].isel(Zl=5))
eke_ps = 0.5 * (u_ps + v_ps)
from matplotlib.colors import LogNorm
plt.figure(figsize=(12,8))
pc = xray.plot.pcolormesh(eke_ps.T, norm=LogNorm())
plt.figure(figsize=(12,5))
ax = plt.subplot(111)
xray.plot.plot(10*eke_ps.mean(dim='time'))
xray.plot.plot(t_ps.mean(dim='time'))
xray.plot.plot(1e5*w_ps.mean(dim='time'))
ax.set_xscale('log')
ax.set_yscale('log')
#ax.set_ylim([1e4,1e9])
x0 = np.array([1e-5, 1e-4])
plt.plot(x0, 1e-14*x0**(-3), 'k-')
plt.plot(x0, 1e-7*x0**(-5./3), 'k--')
plt.legend(['EKE', 'SST', 'W', r'$k^{-3}$', r'$k^{-5/3}$'])
plt.grid()
plt.title('Isotropic Power Spectra')
plt.figure(figsize=(7,10))
pc = xray.plot.pcolormesh(ds['WVEL'].isel(time=0,Zl=5))
pc.set_clim([-1e-3,1e-3])
plt.title('W')
Ny,Nx = (32,16)
a = np.random.rand(Nx)
afft = np.fft.fftn(a)
b = np.random.rand(Ny,Nx)
bfft = np.random.rand()
print (a**2).mean()
print np.real(a*a.conj()).sum()
print (b**2).mean()
print np.real(b*b.conj()).sum()
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# 02. Train locally
* Create or load workspace.
* Create scripts locally.
* Create `train.py` in a folder, along with a `my.lib` file.
* Configure & execute a local run in a user-managed Python environment.
* Configure & execute a local run in a system-managed Python environment.
* Configure & execute a local run in a Docker environment.
* Query run metrics to find the best model
* Register model for operationalization.
## Prerequisites
Make sure you go through the [00. Installation and Configuration](00.configuration.ipynb) Notebook first if you haven't.
```
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
```
## Initialize Workspace
Initialize a workspace object from persisted configuration.
```
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\n')
```
## Create An Experiment
**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments.
```
from azureml.core import Experiment
experiment_name = 'train-on-local'
exp = Experiment(workspace=ws, name=experiment_name)
```
## View `train.py`
`train.py` is already created for you.
```
with open('./train.py', 'r') as f:
print(f.read())
```
Note `train.py` also references a `mylib.py` file.
```
with open('./mylib.py', 'r') as f:
print(f.read())
```
## Configure & Run
### User-managed environment
Below, we use a user-managed run, which means you are responsible to ensure all the necessary packages are available in the Python environment you choose to run the script.
```
from azureml.core.runconfig import RunConfiguration
# Editing a run configuration property on-fly.
run_config_user_managed = RunConfiguration()
run_config_user_managed.environment.python.user_managed_dependencies = True
# You can choose a specific Python environment by pointing to a Python path
#run_config.environment.python.interpreter_path = '/home/johndoe/miniconda3/envs/sdk2/bin/python'
```
#### Submit script to run in the user-managed environment
Note whole script folder is submitted for execution, including the `mylib.py` file.
```
from azureml.core import ScriptRunConfig
src = ScriptRunConfig(source_directory='./', script='train.py', run_config=run_config_user_managed)
run = exp.submit(src)
```
#### Get run history details
```
run
```
Block to wait till run finishes.
```
run.wait_for_completion(show_output=True)
```
### System-managed environment
You can also ask the system to build a new conda environment and execute your scripts in it. The environment is built once and will be reused in subsequent executions as long as the conda dependencies remain unchanged.
```
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
run_config_system_managed = RunConfiguration()
run_config_system_managed.environment.python.user_managed_dependencies = False
run_config_system_managed.auto_prepare_environment = True
# Specify conda dependencies with scikit-learn
cd = CondaDependencies.create(conda_packages=['scikit-learn'])
run_config_system_managed.environment.python.conda_dependencies = cd
```
#### Submit script to run in the system-managed environment
A new conda environment is built based on the conda dependencies object. If you are running this for the first time, this might take up to 5 mninutes. But this conda environment is reused so long as you don't change the conda dependencies.
```
src = ScriptRunConfig(source_directory="./", script='train.py', run_config=run_config_system_managed)
run = exp.submit(src)
```
#### Get run history details
```
run
```
Block and wait till run finishes.
```
run.wait_for_completion(show_output = True)
```
### Docker-based execution
**IMPORTANT**: You must have Docker engine installed locally in order to use this execution mode. If your kernel is already running in a Docker container, such as **Azure Notebooks**, this mode will **NOT** work.
You can also ask the system to pull down a Docker image and execute your scripts in it.
```
run_config_docker = RunConfiguration()
run_config_docker.environment.python.user_managed_dependencies = False
run_config_docker.auto_prepare_environment = True
run_config_docker.environment.docker.enabled = True
run_config_docker.environment.docker.base_image = azureml.core.runconfig.DEFAULT_CPU_IMAGE
# Specify conda dependencies with scikit-learn
cd = CondaDependencies.create(conda_packages=['scikit-learn'])
run_config_docker.environment.python.conda_dependencies = cd
src = ScriptRunConfig(source_directory="./", script='train.py', run_config=run_config_docker)
```
Submit script to run in the system-managed environment
A new conda environment is built based on the conda dependencies object. If you are running this for the first time, this might take up to 5 mninutes. But this conda environment is reused so long as you don't change the conda dependencies.
```
import subprocess
# Check if Docker is installed and Linux containers are enables
if subprocess.run("docker -v", shell=True) == 0:
out = subprocess.check_output("docker system info", shell=True, encoding="ascii").split("\n")
if not "OSType: linux" in out:
print("Switch Docker engine to use Linux containers.")
else:
run = exp.submit(src)
else:
print("Docker engine not installed.")
#Get run history details
run
run.wait_for_completion(show_output=True)
```
## Query run metrics
```
# get all metris logged in the run
run.get_metrics()
metrics = run.get_metrics()
```
Let's find the model that has the lowest MSE value logged.
```
import numpy as np
best_alpha = metrics['alpha'][np.argmin(metrics['mse'])]
print('When alpha is {1:0.2f}, we have min MSE {0:0.2f}.'.format(
min(metrics['mse']),
best_alpha
))
```
You can also list all the files that are associated with this run record
```
run.get_file_names()
```
We know the model `ridge_0.40.pkl` is the best performing model from the eariler queries. So let's register it with the workspace.
```
# supply a model name, and the full path to the serialized model file.
model = run.register_model(model_name='best_ridge_model', model_path='./outputs/ridge_0.40.pkl')
print(model.name, model.version, model.url)
```
Now you can deploy this model following the example in the 01 notebook.
| github_jupyter |
     
     
     
     
     
  
[Home Page](Start_Here.ipynb)
[Previous Notebook](Introduction_to_Deepstream_and_Gstreamer.ipynb)
     
     
     
    
[1](Introduction_to_Deepstream_and_Gstreamer.ipynb#)
[2]
[3](Introduction_to_Multi-DNN_pipeline.ipynb)
[4](Multi-stream_pipeline.ipynb)
[5](Multi-stream_Multi_DNN.ipynb)
     
     
     
     
[Next Notebook](Introduction_to_Multi-DNN_pipeline.ipynb)
# Getting started with Deepstream pipeline
In this notebook, you will be get started with DeepStream's Python Bindings ,it's workflow and build a 4-class object detection pipeline.
**Contents of this Notebook :**
- [NVIDIA DeepStream Plugins](#NVIDIA-DeepStream-Plugins)
- [Nvinfer](#Nvinfer)
- [Nvvidconv](#Nvvidconv)
- [Nvosd](#Nvosd)
- [Building the pipeline](#Building-the-pipeline)
- [Understanding the configuration file](#Understanding-the-configuration-file)
- [Working with the Metadata](#Working-with-the-Metadata)
We will be building a 4-class object detection pipeline as shown in the illustration below.

We notice there are multiple DeepStream plugins used in the pipeline , Let us have a look at them and try to understand them.
## NVIDIA DeepStream Plugins
### Nvinfer
The nvinfer plugin provides [TensorRT](https://docs.nvidia.com/deeplearning/tensorrt/developer-guide/index.html)-based inference for detection and tracking. The lowlevel library (libnvds_infer) operates either on float RGB or BGR planar data with dimensions of Network Height and Network Width. The plugin accepts NV12/RGBA data from upstream components like the decoder, muxer, and dewarper.
The Gst-nvinfer plugin also performs preprocessing operations like format conversion, scaling, mean subtraction, and produces final float RGB/BGR planar data which is passed to the low-level library. The low-level library uses the TensorRT engine for inferencing. It outputs each classified object’s class and each detected object’s bounding boxes (Bboxes) after clustering.

### Nvvidconv
We create the nvvidconv plugin that performs color format conversions, which is required to make data ready for the nvosd plugin.

### Nvosd
The nvosd plugin draws bounding boxes, text, and RoI (Regions of Interest) polygons (Polygons are presented as a set of lines). The plugin accepts an RGBA buffer with attached metadata from the upstream component. It
draws bounding boxes, which may be shaded depending on the configuration (e.g. width, color, and opacity) of a given bounding box. It also draws text and RoI polygons at specified locations in the frame. Text and polygon parameters are configurable through metadata.

Now with this idea , let us get started into building the pipeline.
# Building the pipeline

```
# Import Required Libraries
import sys
sys.path.append('../source_code')
import gi
import time
gi.require_version('Gst', '1.0')
from gi.repository import GObject, Gst
from common.bus_call import bus_call
import pyds
# Defining the Class Labels
PGIE_CLASS_ID_VEHICLE = 0
PGIE_CLASS_ID_BICYCLE = 1
PGIE_CLASS_ID_PERSON = 2
PGIE_CLASS_ID_ROADSIGN = 3
# Defining the input output video file
INPUT_VIDEO_NAME = '/opt/nvidia/deepstream/deepstream-5.0/samples/streams/sample_720p.h264'
OUTPUT_VIDEO_NAME = "../source_code/N1/ds_out.mp4"
```
We define a function `make_elm_or_print_err()` to create our elements and report any errors if the creation fails.
Elements are created using the `Gst.ElementFactory.make()` function as part of Gstreamer library.
```
## Make Element or Print Error and any other detail
def make_elm_or_print_err(factoryname, name, printedname, detail=""):
print("Creating", printedname)
elm = Gst.ElementFactory.make(factoryname, name)
if not elm:
sys.stderr.write("Unable to create " + printedname + " \n")
if detail:
sys.stderr.write(detail)
return elm
```
#### Initialise GStreamer and Create an Empty Pipeline
```
# Standard GStreamer initialization
GObject.threads_init()
Gst.init(None)
# Create gstreamer elements
# Create Pipeline element that will form a connection of other elements
print("Creating Pipeline \n ")
pipeline = Gst.Pipeline()
if not pipeline:
sys.stderr.write(" Unable to create Pipeline \n")
```
#### Create Elements that are required for our pipeline
```
########### Create Elements required for the Pipeline ###########
# Source element for reading from the file
source = make_elm_or_print_err("filesrc", "file-source","Source")
# Since the data format in the input file is elementary h264 stream we need a h264parser
h264parser = make_elm_or_print_err("h264parse", "h264-parser","h264 parse")
# Use nvdec_h264 for hardware accelerated decode on GPU
decoder = make_elm_or_print_err("nvv4l2decoder", "nvv4l2-decoder","Nvv4l2 Decoder")
# Create nvstreammux instance to form batches from one or more sources.
streammux = make_elm_or_print_err("nvstreammux", "Stream-muxer",'NvStreamMux')
# Use nvinfer to run inferencing on decoder's output,behaviour of inferencing is set through config file
pgie = make_elm_or_print_err("nvinfer", "primary-inference" ,"pgie")
# Use convertor to convert from NV12 to RGBA as required by nvosd
nvvidconv = make_elm_or_print_err("nvvideoconvert", "convertor","nvvidconv")
# Create OSD to draw on the converted RGBA buffer
nvosd = make_elm_or_print_err("nvdsosd", "onscreendisplay","nvosd")
# Finally encode and save the osd output
queue = make_elm_or_print_err("queue", "queue", "Queue")
# Use convertor to convert from NV12 to RGBA as required by nvosd
nvvidconv2 = make_elm_or_print_err("nvvideoconvert", "convertor2","nvvidconv2")
# Place an encoder instead of OSD to save as video file
encoder = make_elm_or_print_err("avenc_mpeg4", "encoder", "Encoder")
# Parse output from Encoder
codeparser = make_elm_or_print_err("mpeg4videoparse", "mpeg4-parser", 'Code Parser')
# Create a container
container = make_elm_or_print_err("qtmux", "qtmux", "Container")
# Create Sink for storing the output
sink = make_elm_or_print_err("filesink", "filesink", "Sink")
```
Now that we have created the elements ,we can now set various properties for out pipeline at this point.
### Understanding the configuration file
We set an `config-file-path` for our nvinfer ( Interference plugin ) and it points to the file `dstest1_pgie_config.txt`
You can have a have a look at the [file](../source_code/N1/dstest1_pgie_config.txt)
Here are some parts of the configuration file :
```
# Copyright (c) 2020 NVIDIA Corporation. All rights reserved.
#
# NVIDIA Corporation and its licensors retain all intellectual property
# and proprietary rights in and to this software, related documentation
# and any modifications thereto. Any use, reproduction, disclosure or
# distribution of this software and related documentation without an express
# license agreement from NVIDIA Corporation is strictly prohibited.
[property]
gpu-id=0
net-scale-factor=0.0039215697906911373
model-file=/opt/nvidia/deepstream/deepstream-5.0/samples/models/Primary_Detector/resnet10.caffemodel
proto-file=/opt/nvidia/deepstream/deepstream-5.0/samples/models/Primary_Detector/resnet10.prototxt
#model-engine-file=/opt/nvidia/deepstream/deepstream-5.0/samples/models/Primary_Detector/resnet10.caffemodel_b1_gpu0_fp32.engine
labelfile-path=/opt/nvidia/deepstream/deepstream-5.0/samples/models/Primary_Detector/labels.txt
int8-calib-file=/opt/nvidia/deepstream/deepstream-5.0/samples/models/Primary_Detector/cal_trt.bin
force-implicit-batch-dim=1
batch-size=1
network-mode=1
process-mode=1
model-color-format=0
num-detected-classes=4
interval=0
gie-unique-id=1
output-blob-names=conv2d_bbox;conv2d_cov/Sigmoid
[class-attrs-all]
pre-cluster-threshold=0.2
eps=0.2
group-threshold=1
```
Here we define all the parameters of our model. In this example we use model-file `resnet10`. `Nvinfer` creates an TensorRT Engine specific to the Host GPU to accelerate it's inference performance.
```
############ Set properties for the Elements ############
print("Playing file ",INPUT_VIDEO_NAME)
# Set Input File Name
source.set_property('location', INPUT_VIDEO_NAME)
# Set Input Width , Height and Batch Size
streammux.set_property('width', 1920)
streammux.set_property('height', 1080)
streammux.set_property('batch-size', 1)
# Timeout in microseconds to wait after the first buffer is available
# to push the batch even if a complete batch is not formed.
streammux.set_property('batched-push-timeout', 4000000)
# Set Congifuration file for nvinfer
pgie.set_property('config-file-path', "../source_code/N1/dstest1_pgie_config.txt")
# Set Encoder bitrate for output video
encoder.set_property("bitrate", 2000000)
# Set Output file name and disable sync and async
sink.set_property("location", OUTPUT_VIDEO_NAME)
sink.set_property("sync", 0)
sink.set_property("async", 0)
```
We now link all the elements in the order we prefer and create Gstreamer bus to feed all messages through it.
```
########## Add and Link ELements in the Pipeline ##########
print("Adding elements to Pipeline \n")
pipeline.add(source)
pipeline.add(h264parser)
pipeline.add(decoder)
pipeline.add(streammux)
pipeline.add(pgie)
pipeline.add(nvvidconv)
pipeline.add(nvosd)
pipeline.add(queue)
pipeline.add(nvvidconv2)
pipeline.add(encoder)
pipeline.add(codeparser)
pipeline.add(container)
pipeline.add(sink)
# We now link the elements together
# file-source -> h264-parser -> nvh264-decoder -> nvinfer -> nvvidconv ->
# queue -> nvvidconv2 -> encoder -> parser -> container -> sink -> output-file
print("Linking elements in the Pipeline \n")
source.link(h264parser)
h264parser.link(decoder)
##### Creating Sink pad and source pads and linking them together
# Create Sinkpad to Streammux
sinkpad = streammux.get_request_pad("sink_0")
if not sinkpad:
sys.stderr.write(" Unable to get the sink pad of streammux \n")
# Create source pad from Decoder
srcpad = decoder.get_static_pad("src")
if not srcpad:
sys.stderr.write(" Unable to get source pad of decoder \n")
srcpad.link(sinkpad)
streammux.link(pgie)
pgie.link(nvvidconv)
nvvidconv.link(nvosd)
nvosd.link(queue)
queue.link(nvvidconv2)
nvvidconv2.link(encoder)
encoder.link(codeparser)
codeparser.link(container)
container.link(sink)
# create an event loop and feed gstreamer bus mesages to it
loop = GObject.MainLoop()
bus = pipeline.get_bus()
bus.add_signal_watch()
bus.connect ("message", bus_call, loop)
```
## Working with the Metadata
Our pipeline now carries the metadata forward but we have not done anything with it until now, but as mentoioned in the above pipeline diagram , we will now create a callback function to write relevant data on the frame once called and create a sink pad in the `nvosd` element to call the function.
```
############## Working with the Metadata ################
def osd_sink_pad_buffer_probe(pad,info,u_data):
#Intiallizing object counter with 0.
obj_counter = {
PGIE_CLASS_ID_VEHICLE:0,
PGIE_CLASS_ID_PERSON:0,
PGIE_CLASS_ID_BICYCLE:0,
PGIE_CLASS_ID_ROADSIGN:0
}
# Set frame_number & rectangles to draw as 0
frame_number=0
num_rects=0
gst_buffer = info.get_buffer()
if not gst_buffer:
print("Unable to get GstBuffer ")
return
# Retrieve batch metadata from the gst_buffer
# Note that pyds.gst_buffer_get_nvds_batch_meta() expects the
# C address of gst_buffer as input, which is obtained with hash(gst_buffer)
batch_meta = pyds.gst_buffer_get_nvds_batch_meta(hash(gst_buffer))
l_frame = batch_meta.frame_meta_list
while l_frame is not None:
try:
# Note that l_frame.data needs a cast to pyds.NvDsFrameMeta
frame_meta = pyds.NvDsFrameMeta.cast(l_frame.data)
except StopIteration:
break
# Get frame number , number of rectables to draw and object metadata
frame_number=frame_meta.frame_num
num_rects = frame_meta.num_obj_meta
l_obj=frame_meta.obj_meta_list
while l_obj is not None:
try:
# Casting l_obj.data to pyds.NvDsObjectMeta
obj_meta=pyds.NvDsObjectMeta.cast(l_obj.data)
except StopIteration:
break
# Increment Object class by 1 and Set Box border to Red color
obj_counter[obj_meta.class_id] += 1
obj_meta.rect_params.border_color.set(0.0, 0.0, 1.0, 0.0)
try:
l_obj=l_obj.next
except StopIteration:
break
################## Setting Metadata Display configruation ###############
# Acquiring a display meta object.
display_meta=pyds.nvds_acquire_display_meta_from_pool(batch_meta)
display_meta.num_labels = 1
py_nvosd_text_params = display_meta.text_params[0]
# Setting display text to be shown on screen
py_nvosd_text_params.display_text = "Frame Number={} Number of Objects={} Vehicle_count={} Person_count={}".format(frame_number, num_rects, obj_counter[PGIE_CLASS_ID_VEHICLE], obj_counter[PGIE_CLASS_ID_PERSON])
# Now set the offsets where the string should appear
py_nvosd_text_params.x_offset = 10
py_nvosd_text_params.y_offset = 12
# Font , font-color and font-size
py_nvosd_text_params.font_params.font_name = "Serif"
py_nvosd_text_params.font_params.font_size = 10
# Set(red, green, blue, alpha); Set to White
py_nvosd_text_params.font_params.font_color.set(1.0, 1.0, 1.0, 1.0)
# Text background color
py_nvosd_text_params.set_bg_clr = 1
# Set(red, green, blue, alpha); set to Black
py_nvosd_text_params.text_bg_clr.set(0.0, 0.0, 0.0, 1.0)
# Using pyds.get_string() to get display_text as string to print in notebook
print(pyds.get_string(py_nvosd_text_params.display_text))
pyds.nvds_add_display_meta_to_frame(frame_meta, display_meta)
############################################################################
try:
l_frame=l_frame.next
except StopIteration:
break
return Gst.PadProbeReturn.OK
# Lets add probe to get informed of the meta data generated, we add probe to the sink pad
# of the osd element, since by that time, the buffer would have had got all the metadata.
osdsinkpad = nvosd.get_static_pad("sink")
if not osdsinkpad:
sys.stderr.write(" Unable to get sink pad of nvosd \n")
osdsinkpad.add_probe(Gst.PadProbeType.BUFFER, osd_sink_pad_buffer_probe, 0)
```
Now with everything defined , we can start the playback and listen the events.
```
# start play back and listen to events
print("Starting pipeline \n")
start_time = time.time()
pipeline.set_state(Gst.State.PLAYING)
try:
loop.run()
except:
pass
# cleanup
pipeline.set_state(Gst.State.NULL)
print("--- %s seconds ---" % (time.time() - start_time))
# Convert video profile to be compatible with Jupyter notebook
!ffmpeg -loglevel panic -y -an -i ../source_code/N1/ds_out.mp4 -vcodec libx264 -pix_fmt yuv420p -profile:v baseline -level 3 ../source_code/N1/output.mp4
# Display the Output
from IPython.display import HTML
HTML("""
<video width="640" height="480" controls>
<source src="../source_code/N1/output.mp4"
</video>
""".format())
```
In the next notebook , we will learn object tracking and build an attribute classification pipeline along with the primary inference build in the notebook.
## Licensing
This material is released by NVIDIA Corporation under the Creative Commons Attribution 4.0 International (CC BY 4.0).
[Previous Notebook](Introduction_to_Deepstream_and_Gstreamer.ipynb)
     
     
     
    
[1](Introduction_to_Deepstream_and_Gstreamer.ipynb#)
[2]
[3](Introduction_to_Multi-DNN_pipeline.ipynb)
[4](Multi-stream_pipeline.ipynb)
[5](Multi-stream_Multi_DNN.ipynb)
     
     
     
     
[Next Notebook](Introduction_to_Multi-DNN_pipeline.ipynb)
     
     
     
     
     
  
[Home Page](Start_Here.ipynb)
| github_jupyter |
# Voice Identification
### n-shot learning
The idea of one shot learning is to train an algorithm so that after only `n` examples, an algorithm can identify a example again.
Siamese networks attempt to do this by training a model to learn how to take a high dimensional input an generate a feature vector. The network is trained by taking two examples either similar or different and training them to reduce/increase the distance of the feature vector generated by the output.
### Applying to Voice Identification
The end goal of this model is to take a audio sample that has undergone speech diarihsation and identify each speaker in the set.
### Datasets
| Name | Speakers | Min | Max
| ------------- |-------------|-----|-----|
[VoxCeleb](http://www.robots.ox.ac.uk/~vgg/data/voxceleb/)| 7000+| 3s | 3s
[10 English Speakers](http://www.openslr.org/resources/45/ST-AEDS-20180100_1-OS) | 10 | ? | ?
### Articles
- https://github.com/zdmc23/oneshot-audio/blob/master/OneShot.ipynb
```
## Notebook settings
%reload_ext autoreload
%autoreload 2
%matplotlib inline
from IPython.core.debugger import set_trace
import sys
sys.path.append("..")
from audio import *
np.random.seed(2)
```
## 10 Speakers
Lets get a feel for the architecture by creating it and training on a dataset that we know we can do well on using standard classification techiques
```
## The actual url is http://www.openslr.org/resources/45/ST-AEDS-20180100_1-OS.tgz
## but we need to strip off the extension otherwise fastai gets confused.
data_url = 'http://www.openslr.org/resources/45/ST-AEDS-20180100_1-OS'
## Need this because the source tar file doesn't extract to its own folder
data_folder = datapath4file(url2name(data_url))
untar_data(data_url, dest=data_folder)
label_pattern = r'_([mf]\d+)_'
audios = AudioList.from_folder(data_folder).split_none().label_from_re(label_pattern)
audios.train.x.tfm_args = tfm_params
audios.valid.x.tfm_args = tfm_params
```
## Data Loading
Generic for any size number of tuple meaning that experiments with Triplet Loss are possible
```
class ItemTuple(ItemBase):
def __init__(self, items):
# Warn if the items gets larger than intended
self.size = len(items)
self.items = items
self.data = torch.cat([x.data.unsqueeze(0) for x in items])
def show(self):
[x.show() for x in self.items]
def apply_tfms(self, tfms):
for tfm in tfms:
self.data = torch.stack([tfm(x) for x in self.data])
return self
def __repr__(self):
return ''.join([str(x) for x in self.items]) + '\n'
def __len__(self):
return self.size
class SiameseList(ItemList):
def __init__(self, *args, open_fn=lambda x:x, **kwargs):
super().__init__(*args, **kwargs)
self.open = open_fn
def get(self, i):
item = super().get(i)
if isinstance(item, ItemTuple):
return item
return ItemTuple([self.open(x) for x in item])
from copy import deepcopy
class SiameseDataset(LabelList):
def show_similar(self):
pass
def show_different(self):
pass
@classmethod
def from_label_list(cls, ll:LabelList, hidden_classes=None, train_num=20, valid_num=10, use_all=False):
# Use random chunk of the classes for validation if none specified
if hidden_classes is None: hidden_classes = list(range(ll.c))[-int(ll.c*0.2)]
if hasattr(hidden_classes[0], 'data'):
hidden_classes = [x.data for x in hidden_classes]
if len(hidden_classes) < 2:
raise Exception("Must be atleast 2 classes")
train_cls = [i for i in range(ll.c - 1) if i not in hidden_classes]
def mask(i,c): i.y.items == c
# Copy to get transforms
tll = deepcopy(ll)
# Generate Items and Labels
tll.x.items = np.concatenate([ll.x.items[mask(ll,c)] for c in train_cls])
tll.y.items = np.concatenate([ll.y.items[mask(ll,c)] for c in train_cls])
vll = deepcopy(ll)
vll.x.items = np.concatenate([ll.x.items[mask(ll,c)] for c in hidden_classes])
vll.y.items = np.concatenate([ll.y.items[mask(ll,c)] for c in hidden_classes])
tll = cls._from_label_list(tll, train_num, train_num//2, use_all)
vll = cls._from_label_list(vll, valid_num, valid_num//2, use_all)
return LabelLists(ll.x.path, tll, vll)
@classmethod
def _from_label_list(cls, ll:LabelList, num_same=20, num_diff=30, use_all=False):
x = ll.x
y = ll.y
# Seperate into lists of individual classes
# Some of these may be empty because of the spit
seperated = [x.items[y.items==c] for c in range(ll.c) if len(x.items[y.items==c]) > 0]
# Create sets of same pairs
# TODO Don't create the same pairs
same_pairs = np.empty((0,2))
for cis in seperated:
r = np.array([np.random.choice(cis, num_same), np.random.choice(cis, num_same)]).T
same_pairs = np.concatenate([same_pairs, r])
# Create pairs of different items
diff_pairs = np.empty((0,2))
for i, cis in enumerate(seperated):
other = [k for k in range(len(seperated)) if k != i]
for i in other:
ocis = seperated[i]
dps = np.array([np.random.choice(cis, num_diff), np.random.choice(ocis, num_diff)]).T
diff_pairs = np.concatenate([diff_pairs, dps])
# Combine together
al = np.concatenate([same_pairs, diff_pairs])
# Generate labels
labels = np.concatenate([np.ones(len(same_pairs), dtype=np.int8), np.zeros(len(diff_pairs), dtype=np.int8)])
inst = cls(SiameseList(al, open_fn=ll.x.open), CategoryList(labels, ['different', 'similar']))
return inst
# We choose to keep one of each gender hidden in our validation set
sd = SiameseDataset.from_label_list(audios.train, [0, 6], 100, 100)
sd.train.x.tfm_args = tfm_params
sd.valid.x.tfm_args = tfm_params
tfm_sg_shift.use_on_y = False # Library broken
tfm_sg_aug.use_on_y = False # Library broken
tfms = [tfm_sg_shift, tfm_sg_aug], []
data = sd.transform(tfms).databunch(bs=30)
x, y = data.train_ds[-159]
print(y)
x.show()
```
Now that we have the data loaded and paired up, we can start to see whether the torch module that we've created works with the data we have.
```
class SiameseNetwork(nn.Module):
def __init__(self, encoder=models.resnet18, s_out=512):
# TODO warn is s_out is to large
super().__init__()
self.body = create_body(encoder, cut=-2)
self.head = create_head(1024, 1, [s_out])[:5]
def forward(self, items):
# Get the batch size in the correct place
ins = items.transpose(1, 0)
outs = [self.body(x) for x in ins]
outs = [self.head(x) for x in outs]
outs = torch.stack(outs)
return outs
```
## Loss Functions
```
def hinge_loss(x, y, m=1):
# Was getting autograd errors if I didn't clone
diff = torch.sqrt(torch.pow(x[0]-x[1], 2)).clone()**2
diff[y==0] = m**2 - diff[y==0]
diff[diff < 0] = 0
return diff
def gen_loss_m(loss_func):
return lambda x,y: loss_func(x, y).mean()
def loss_less_acc(loss_func, l):
return lambda x,y: (loss_func(x,y) < l**2).float().mean()
def create_loss_acc(loss_func, l):
return gen_loss_m(loss_func), loss_less_acc(loss_func, l)
```
## Creating a Learner
Here we create a loss function and accuracy metric to help track how many of the items in the
validation set are below the threshold number as the loss is continous
```
def siamese_learner(data: DataBunch, encoder:nn.Module=models.resnet18, s_out=512, loss_func=None, loss_size=None, m=3):
if loss_func is None: loss_func = partial(hinge_loss, m=m)
# m/2 is the middle of confidence so if we're bellow it that means we've guessed correctly
# although with very low confidence
if loss_size is None: loss_size = m/2
loss, acc = create_loss_acc(loss_func, loss_size)
learner = Learner(data, SiameseNetwork(encoder, s_out), loss_func=loss, metrics=acc)
learner.encode = lambda x: x # TODO create a simple way to get a vector of a piece of data
return learner
# Here we decide the length of the output vector.
# This will be dependent on your encoder arch (bigger means bigger s_out)
learn = siamese_learner(data, encoder=models.resnet18, s_out=256, m=4)
# Audio Specific
learn.model.body[0] = nn.Conv2d(1, 64, kernel_size=(7, 7), stride=(2, 2), padding=(3, 3), bias=False).cuda()
learn.lr_find()
learn.recorder.plot()
learn.fit_one_cycle(20, 1e-3)
```
# TODO
- More reasonable split of validation and training set
- Duplicate pairs potentially being added.
- Use on image problem
- Show predictions
- Compare the results of having k classes with n examples of a specific class
| github_jupyter |
```
%matplotlib inline
import matplotlib.cm as cm
```
### Grab a random latin hypercube grid of $U(\beta_{\rm early})$, $U(\beta_{\rm late})$ and $U(\log_{10}\tau_{\rm c})$
```
from conc_pop_model import get_u_param_grids
N_GRID = 250
u_be_grid, u_lgtc_bl_grid = get_u_param_grids(N_GRID)
```
### Calculate the PDF weight associated with each grid point and each value of $p_{50\%}$
```
from conc_pop_model import DEFAULT_PARAMS, get_pdf_weights_on_grid
from diffprof.nfw_evolution import DEFAULT_CONC_PARAMS
conc_k = DEFAULT_CONC_PARAMS['conc_k']
params_p50 = np.array(list(DEFAULT_PARAMS.values()))
N_P50 = 25
p50_arr = np.linspace(0.1, 0.9, N_P50)
_res = get_pdf_weights_on_grid(p50_arr, u_be_grid, u_lgtc_bl_grid, conc_k, params_p50)
u_be_weights, u_lgtc_bl_weights = _res
print(u_be_weights.shape, u_lgtc_bl_weights.shape)
```
### Generate a prediction for ${\rm conc}(t)$ for each point on the grid and each value of $p_{50\%}$
```
from conc_pop_model import lgc_pop_vs_lgt_and_p50, get_param_grids_from_u_param_grids
N_TIMES = 60
tarr = np.linspace(2, 13.8, N_TIMES)
lgtarr = np.log10(tarr)
be_grid, lgtc_bl_grid = get_param_grids_from_u_param_grids(u_be_grid, u_lgtc_bl_grid)
_res = lgc_pop_vs_lgt_and_p50(lgtarr, p50_arr, be_grid, lgtc_bl_grid, conc_k)
lgc_p50_pop = _res
print(lgc_p50_pop.shape)
```
### Calculate $\langle{\rm conc}(t)\vert p_{50\%}\rangle$ as the PDF-weighted sum
```
avg_log_conc_p50 = np.sum(u_be_weights.reshape((N_GRID, N_P50, 1))*lgc_p50_pop, axis=0)
avg_log_conc_p50.shape
```
### Plot the results
```
colormap_seq=np.linspace(1, 0, N_P50)
colors=cm.coolwarm(colormap_seq)
fig, ax = plt.subplots(1, 1)
yscale = ax.set_yscale('log')
ylim = ax.set_ylim(1, 90)
xlim = ax.set_xlim(1.5, 13.8)
for ip50 in range(N_P50):
__=ax.plot(tarr, 10**avg_log_conc_p50[ip50, :], color=colors[ip50])
```
### Hmmm...
There is very little, if any, $p_{50\%}-$dependence that shows up in that plot. This _could_ just mean that the model parameters are poorly tuned (which is my guess), or it could mean there is a bug somewhere, I haven't had a chance to do more testing. For now I would proceed with next steps with existing code, but just remember that there could be some more digging to do here to finish the single-mass implementation.
## Next steps
### Step 1
For starters, in order to make sure everything is working correctly, pick a single mass and use gradient descent to fit the parameters of this model to capture the usual summary statistics:
1. $\langle{\rm log_{10}(conc(t))}\vert p_{50\%},M_0\rangle$
2. $\langle{\rm log_{10}(conc(t))}\vert M_0\rangle$
3. $\sigma({\rm log_{10}(conc(t))}\vert M_0)$
Calculating (1) is demonstrated above in this notebook. You will calculate (2) by averaging over $p_{50\%}$. You will calculate (3) by _first_ squaring the result of (2), and then in a separate step, calculating $\langle{\rm log_{10}(conc(t))^{2}}\vert M_0\rangle$, and then finally taking the square root of their difference. See the ${\tt diffmah}$ source code for an example of how to differentiably compute the stddev of a logarithmic variable, which for you is $x={\rm log_{10}(conc(t))},$ whereas in ${\tt diffmah}$ was $x=\log_{10}(M_{\rm halo}(t)).$
When you do the fitting, for your target data try using the target data model function, hopefully it will be less noisy than the actual simulated data. Try and use the "batch-training" trick we discussed on slack, such that for each step of the gradient descent, generate some new target data based on a newly generated array of grid parameters and $p_{50\%$ values. The notebook I shared with you on Slack shows how to do this using a latin hypercube to randomly generate the arrays. Repeat this exercise for a few different values of $M_0$ and verify that the differentiable model has been formulated with sufficient flexibility to give an accurate approximation.
### Step 2
Once you have a working differentiable version of the single-mass model, then it will be time to adapt the code to build a differentiable mass-dependent model. When you train the model, again use the batch-training tricks, this time selecting different parameter grids and different $p_{50\%}$ and different $M_0$ grids each time. Once you have an accurate fit of the population-level model across $M_0$ and $p_{50\%},$ that will be the principal result of the project and it will be time to write up the results and submit for publication.
| github_jupyter |
```
import importlib
from library import data_preprocess as dp
import matplotlib.pyplot as plt
importlib.reload(dp)
import pickle
#https://www.kaggle.com/kazanova/sentiment140/data
file_path1 = "./data/training.1600000.processed.noemoticon.csv"
tweets = list(set(dp.load_data(file_path1, header=False, sep=',', usecols=[5])[5].str.lower().values))
print(len(tweets))
print(tweets[:2])
unique_chars = dp.get_unique_chars(tweets)
print(len(unique_chars))
print(unique_chars)
dp.get_char_presence(quotes)
print("Total number of tweets = ", len(tweets))
filter_tweets = dp.filter_data(tweets, "^[a-zA-Z !\"',.?\-\:\*]+$")
print(len(filter_tweets))
print(filter_tweets[:10])
pad_chars = [' ', '!', '"', '*', ',', '-', ':', '?']
clean_tweets = dp.clean_data(tweets, "^[a-zA-Z !\"'*,\-.:?]+$", pad_chars)
unique_chars = dp.get_unique_chars(clean_tweets)
print(len(unique_chars))
print(unique_chars)
clean_tweets = dp.tokenize(clean_tweets)
print(len(clean_tweets))
print(clean_tweets[:2])
# stats of number of tokens in the sentence
dp.get_len_stats(clean_tweets)
MIN_NUM_TOKENS = 10
MAX_NUM_TOKENS = 40
clean_tweets = dp.filter_data_on_length(clean_tweets, MIN_NUM_TOKENS, MAX_NUM_TOKENS)
print(len(clean_tweets))
word_freqs = dp.get_vocabulary(clean_tweets)
dp.get_vocabulary_stats(word_freqs)
filtered_words = dp.filter_words(word_freqs, 20)
print("Number of filtered words = ", len(filtered_words))
sents = dp.get_sentences_with_words(clean_tweets, filtered_words)
print(len(sents))
print(sents[:5])
# Analysis for choosing optimum word freq cut off
freqsL = list(range(1, 80, 5))
num_words = []
num_sents = []
for x in freqsL:
filtered_words = dp.filter_words(word_freqs, x)
num_words.append(len(filtered_words))
num_sents.append(len(dp.get_sentences_with_words(clean_tweets, filtered_words)))
plt.plot(num_words, num_sents)
plt.show()
# let's consider tweets whose vocabulary matches with that of jokes and quotes dataset
with open('./datasets/jokes_vocabulary.pickle', 'rb') as pickleFile:
jokes_vocabulary = pickle.load(pickleFile)
with open('./datasets/quotes_vocabulary.pickle', 'rb') as pickleFile:
quotes_vocabulary = pickle.load(pickleFile)
final_vocabulary = jokes_vocabulary
final_vocabulary.extend(quotes_vocabulary)
final_vocabulary = list(set(final_vocabulary))
print("vocabulary size = ", len(final_vocabulary))
final_clean_tweets = dp.get_sentences_with_words(clean_tweets, final_vocabulary)
print("Number of sentences: ", len(final_clean_tweets))
print(final_clean_tweets[:10])
# Let's append sos and eos tokens to all sentences
clean_tweets_final = [["sos"] + tokens + ["eos"] for tokens in final_clean_tweets]
clean_tweets_final = [' '.join(tokens) for tokens in clean_tweets_final]
clean_tweets_final = list(set(clean_tweets_final))
vocabulary_final = dp.filter_words(dp.get_vocabulary(dp.tokenize(clean_tweets_final)), 0)
print("Final vocabulary size: ", len(vocabulary_final))
print("Number of sentences: ", len(clean_tweets_final))
print(clean_tweets_final[:5])
print(vocabulary_final[:10])
with open("./datasets/tweets.pickle","wb") as fop:
pickle.dump(clean_tweets_final, fop)
with open("./datasets/tweets_vocabulary.pickle","wb") as fop:
pickle.dump(vocabulary_final, fop)
print(clean_tweets_final[:15])
```
| github_jupyter |
```
import re
import urllib
import urllib3
import requests
from bs4 import BeautifulSoup
urllib3.disable_warnings()
headers = {'User-Agent':'Mozilla/5.0'}
len(dataset1)
384-290
58
94
https://www.botresell.com/product-category/wordpress-plugin/ 81
qq
mainlink
for aa in range(0,7):
mainlink='https://www.botresell.com/product-category/wordpress-plugin/page/'+str(aa)
page1 = requests.get(mainlink, headers=headers, verify=False)
data1 = page1.content
soup1 = BeautifulSoup(data1, "html.parser")
categories=soup1.find_all('h1',class_="page-title")[0].text
# print categories
products=soup1.find_all('a',class_="woocommerce-LoopProduct-link")
for qq in range(0,len(products)):
dataset0=['-']*20
products_link=products[qq].get('href')
# print products_link
dataset0[0]=categories
dataset0[1]=products_link
page = requests.get(products_link, headers=headers, verify=False)
data = page.content
soup = BeautifulSoup(data, "html.parser")
title=soup.find_all('h1',class_="product_title entry-title")[0].text
dataset0[2]=title
pricet=soup.find_all('p',class_="price")[0]
price_real0=pricet.find_all('del')
if len(price_real0)>0:
price_real=price_real0[0].text
dataset0[3]=price_real
price_disc0=pricet.find_all('ins')
if len(price_disc0)>0:
price_disc=price_disc0[0].text
dataset0[4]=price_disc
# print price_real,price_disc
logolink0=soup.find_all('div',class_="images")[0].find_all('a')
if len(logolink0)>0:
logolink=logolink0[0].get('href')
print 'copying logo'
# print logolink
phot_name=title+'.png'
dataset0[5]=phot_name
urllib.urlretrieve(logolink,phot_name)
product_summery=soup.find_all('div',class_="product_summery")
for jj in range(0,len(product_summery)):
otherimages=product_summery[jj].find_all('img')
print len(otherimages)
for ss in range(0,len(otherimages)):
other_images_link=otherimages[ss].get('src')
# print other_images_link
other_images_name=title+'_'+str(ss)+'.png'
dataset0[6+ss]=other_images_name
urllib.urlretrieve(other_images_link,other_images_name)
textp=product_summery[jj].find_all('p')
dataset01=[]
for pp in range(0,len(textp)):
if len(textp[pp].text)>1:
# print textp[pp].text
dataset0.append(textp[pp].text)
print dataset0
dataset1.append(dataset0)
len(dataset1)
print products_link
from openpyxl import load_workbook
import warnings
from openpyxl import Workbook
wb = Workbook(write_only=True)
ws = wb.create_sheet()
# now we'll fill it with 100 rows x 200 columns
for irow in dataset1:
ws.append(irow)
# save the file
wb.save('first1.xlsx')
```
| github_jupyter |
```
#akyork
#written to analyze the 16 HMO glycans provided by Ben
import sys
import __init__
import json_utility
from glypy.io import glycoct, iupac
import extract_motif
import customize_motif_vec
import glycan_io
import pandas as pd
import numpy as np
import plot_glycan_utilities
import seaborn as sns
import motif_class
from importlib import reload
import clustering_analysis_pip
import matplotlib.pyplot as plt
import ndex
from ndex.networkn import NdexGraph
import networkx as nx
import warnings
warnings.filterwarnings('ignore')
import json_utility
%matplotlib inline
root_addr = '/Users/apple/Desktop/NathanLab/CHO_Anders/lib_cho'
__init__.exact_Ture=True
glycan_dict_addr = root_addr+"glycan_cho_dict.json"
glycan_dict = glycan_io.load_glycan_obj_from_dir('/Users/apple/Desktop/NathanLab/CHO_Anders/GlycanDef/')
name_list = ['A4FG4S4', 'M9']
for i in name_list:
plot_glycan_utilities.plot_glycan(glycan_dict[i], i)
libDict = {}
libstrDict = {}
for i in name_list:
libDict[i] = glycan_dict[i]
libstrDict[i] = str(glycan_dict[i])
json_utility.store_json(glycan_dict_addr, libstrDict)
#run motif search on 16 HMO glycans
#loads the json into dictionary form
#gets motifs and saves to json file
glycan_motif_dict = root_addr+"glycan_motif_dict.json"
choMotif_dict = extract_motif.get_motif_pip(libDict, gly_len=22, output_file=glycan_motif_dict)
# hmoMotif_dict = glycan_io.glycan_str_to_glycan(json_utility.load_json(root_addr+"glycanHMO_motif_dict.json"))
# glycan_dict_addr = root_addr+"glycan_dict.json"
output_merged_motif_addr = root_addr+"lib_motif_dict.json"
master_motif_dict = customize_motif_vec.merge_motif_dict_pipe(choMotif_dict, output_merged_motif_addr, glycan_dict_addr)
motif_vector = glycan_io.motif_dict_to_motif_vec(master_motif_dict)
reload(motif_class)
_motif_lab = motif_class.MotifLabwithCore(motif_vector, glycan_core=motif_class.tri_glycan_core) # unicarbkb_motifs_12259.json
motif_vec = [_motif_lab.motif_vec[i] for i in _motif_lab.motif_with_core_list]
{MonosaccharideResidue(Glc2NAc): 6,
MonosaccharideResidue(Fuc): 1,
MonosaccharideResidue(Man): 3,
MonosaccharideResidue(Neu5Ac): 4,
MonosaccharideResidue(Gal): 4}
import nglycan_composition
reload(nglycan_composition)
lib_dict = {}
count = 0
lib_vec = []
for i in motif_vec:
_glycan_comp = nglycan_composition.nglycan_composition(i)
if _glycan_comp.possible:
lib_vec.append(i)
print(_glycan_comp.mono_composition())
print(_glycan_comp.composition())
count+=1
if _glycan_comp.shorthand() not in lib_dict:
lib_dict[_glycan_comp.shorthand()] = [i]
else:
lib_dict[_glycan_comp.shorthand()].append(i)
count
_glycan_comp = nglycan_composition.nglycan_composition(lib_vec[89])
plot_glycan_utilities.plot_glycan(lib_vec[89])
print(_glycan_comp.mono_composition())
print(_glycan_comp.composition())
print(_glycan_comp.shorthand())
print(_glycan_comp.shorthand_dict)
plot_glycan_utilities.plot_glycan_list(lib_vec, range(len(lib_vec)), )
plot_glycan_utilities.plot_glycan_list(lib_vec[84:87])
lib_dict.keys()
a_glycan
a_glycan[0].total_composition()
glycan.
```
| github_jupyter |
# Plot the AR6 SSP ERF time series 1750-2500
Theme Song: Endless Ways<br>
Artist: Anathema<br>
Album: The Optimist<br>
Released: 2017
```
import matplotlib.pyplot as pl
import numpy as np
import pandas as pd
from matplotlib import gridspec, rc
pl.rcParams['figure.figsize'] = (18/2.54, 22/2.54)
pl.rcParams['font.size'] = 9
pl.rcParams['font.family'] = 'Arial'
pl.rcParams['xtick.direction'] = 'out'
pl.rcParams['xtick.minor.visible'] = True
pl.rcParams['ytick.minor.visible'] = True
pl.rcParams['ytick.right'] = True
pl.rcParams['xtick.top'] = True
scenarios = ['ssp119','ssp126','ssp245','ssp370','ssp370-lowNTCF','ssp370-lowNTCFCH4','ssp434','ssp460','ssp534-over','ssp585']
# put in a module
colors = {
'ssp119': '#1e9583',
'ssp126': '#1d3354',
'ssp245': '#e9dc3d',
'ssp370': '#f11111',
'ssp370-lowNTCF': '#f11111',
'ssp370-lowNTCFCH4': '#f11111',
'ssp434': '#63bce4',
'ssp460': '#e78731',
'ssp534-over': '#996dc8',
'ssp585': '#830b22',
}
ls = {
'ssp119': '-',
'ssp126': '-',
'ssp245': '-',
'ssp370': '-',
'ssp370-lowNTCF': '--',
'ssp370-lowNTCFCH4': ':',
'ssp434': '-',
'ssp460': '-',
'ssp534-over': '-',
'ssp585': '-',
}
forcing = {}
forcing_p05 = {}
forcing_p95 = {}
for scenario in scenarios:
forcing[scenario] = pd.read_csv('../data_output/SSPs/ERF_%s_1750-2500.csv' % scenario)
forcing_p05[scenario] = pd.read_csv('../data_output/SSPs/ERF_%s_1750-2500_pc05.csv' % scenario)
forcing_p95[scenario] = pd.read_csv('../data_output/SSPs/ERF_%s_1750-2500_pc95.csv' % scenario)
pl.figure()
gs = gridspec.GridSpec(6,4)
ax_main = pl.subplot(gs[0:3, 0:])
for scenario in scenarios:
if scenario=='ssp534-over':
zorder=3
else:
zorder=4
if 'lowNTCF' not in scenario:
ax_main.fill_between(np.arange(1750, 2501), forcing_p05[scenario]['total'], forcing_p95[scenario]['total'], color=colors[scenario], lw=0, alpha=0.2, zorder=zorder)
ax_main.plot(np.arange(1750,2501), forcing[scenario]['total'], label=scenario, color=colors[scenario], ls=ls[scenario], zorder=zorder)
ax_main.set_ylim(0,15)
ax_main.set_xlim(2000,2500)
#ax_main.legend()
ax_main.set_title('Total Effective Radiative Forcing in SSP scenarios, 2000-2500')
ax_main.set_ylabel('W m$^{-2}$')
ax_co2 = pl.subplot(gs[3, 0])
ax_ch4 = pl.subplot(gs[3, 1])
ax_n2o = pl.subplot(gs[3, 2])
ax_oth = pl.subplot(gs[3, 3])
ax_ozo = pl.subplot(gs[4, 0])
ax_h2o = pl.subplot(gs[4, 1])
ax_con = pl.subplot(gs[4, 2])
ax_ari = pl.subplot(gs[4, 3])
ax_aci = pl.subplot(gs[5, 0])
ax_bcs = pl.subplot(gs[5, 1])
ax_lnd = pl.subplot(gs[5, 2])
ax_nat = pl.subplot(gs[5, 3])
ax_bcs.axhline(0, ls=':', color='k', lw=0.5)
ax_aci.axhline(0, ls=':', color='k', lw=0.5)
ax_ozo.axhline(0, ls=':', color='k', lw=0.5)
for scenario in scenarios:
ax_co2.plot(np.arange(1750,2501),forcing[scenario]['co2'], color=colors[scenario], ls=ls[scenario])
ax_ch4.plot(np.arange(1750,2501),forcing[scenario]['ch4'], color=colors[scenario], ls=ls[scenario])
ax_n2o.plot(np.arange(1750,2501),forcing[scenario]['n2o'], color=colors[scenario], ls=ls[scenario])
ax_oth.plot(np.arange(1750,2501),forcing[scenario]['other_wmghg'], color=colors[scenario], ls=ls[scenario])
ax_ozo.plot(np.arange(1750,2501),forcing[scenario]['o3'], color=colors[scenario], ls=ls[scenario])
ax_h2o.plot(np.arange(1750,2501),forcing[scenario]['h2o_stratospheric'], color=colors[scenario], ls=ls[scenario])
ax_con.plot(np.arange(1750,2501),forcing[scenario]['contrails'], color=colors[scenario], ls=ls[scenario])
ax_ari.plot(np.arange(1750,2501),forcing[scenario]['aerosol-radiation_interactions'], color=colors[scenario], ls=ls[scenario])
ax_aci.plot(np.arange(1750,2501),forcing[scenario]['aerosol-cloud_interactions'], color=colors[scenario], ls=ls[scenario])
ax_bcs.plot(np.arange(1750,2501),forcing[scenario]['bc_on_snow'], color=colors[scenario], ls=ls[scenario])
ax_lnd.plot(np.arange(1750,2501),forcing[scenario]['land_use'], color=colors[scenario], ls=ls[scenario])
#ax_nat.plot(np.arange(1750,2501),forcing[scenario]['volcanic'], color='k', ls=ls[scenario])
ax_nat.plot(np.arange(1750,2501),forcing[scenario]['solar'], color='k', ls=ls[scenario])
ax_co2.set_xlim(2000, 2500)
ax_ch4.set_xlim(2000, 2500)
ax_n2o.set_xlim(2000, 2500)
ax_oth.set_xlim(2000, 2500)
ax_ozo.set_xlim(2000, 2500)
ax_h2o.set_xlim(2000, 2500)
ax_con.set_xlim(2000, 2500)
ax_ari.set_xlim(2000, 2500)
ax_aci.set_xlim(2000, 2500)
ax_bcs.set_xlim(2000, 2500)
ax_lnd.set_xlim(2000, 2500)
ax_nat.set_xlim(2000, 2500)
ax_co2.set_ylim(0, 13)
ax_ch4.set_ylim(0, 1.1)
ax_n2o.set_ylim(0, 0.8)
ax_oth.set_ylim(0, 0.7)
ax_ozo.set_ylim(-0.2, 0.8)
ax_h2o.set_ylim(0, 0.1)
ax_con.set_ylim(0, 0.2)
ax_ari.set_ylim(-0.4, 0)
ax_aci.set_ylim(-1.2, 0.2)
ax_bcs.set_ylim(-0.02, 0.14)
ax_lnd.set_ylim(-0.4, 0)
ax_nat.set_ylim(-0.1, 0.1)
ax_co2.text(0.02, 0.98, 'CO$_2$', va='top', ha='left', transform=ax_co2.transAxes)
ax_ch4.text(0.98, 0.98, 'CH$_4$', va='top', ha='right', transform=ax_ch4.transAxes)
ax_n2o.text(0.02, 0.98, 'N$_2$O', va='top', ha='left', transform=ax_n2o.transAxes)
ax_oth.text(0.98, 0.98, 'Halogenated\ngases', va='top', ha='right', transform=ax_oth.transAxes)
ax_ozo.text(0.98, 0.98, 'O$_3$', va='top', ha='right', transform=ax_ozo.transAxes)
ax_h2o.text(0.98, 0.98, 'Strat. H$_2$O', va='top', ha='right', transform=ax_h2o.transAxes)
ax_con.text(0.98, 0.98, 'Contrails and\naviation-\ninduced\ncirrus', va='top', ha='right', transform=ax_con.transAxes)
ax_ari.text(0.98, 0.02, 'Aerosol-radiation\ninteractions', va='bottom', ha='right', transform=ax_ari.transAxes)
ax_aci.text(0.98, 0.02, 'Aerosol-cloud\ninteractions', va='bottom', ha='right', transform=ax_aci.transAxes)
ax_bcs.text(0.98, 0.98, 'Light absorbing\nparticles on\nsnow and\nice', va='top', ha='right', transform=ax_bcs.transAxes)
ax_lnd.text(0.02, 0.98, 'Land use', va='top', ha='left', transform=ax_lnd.transAxes)
ax_nat.text(0.98, 0.98, 'Solar', va='top', ha='right', transform=ax_nat.transAxes)
ax_co2.set_ylabel('W m$^{-2}$')
ax_ozo.set_ylabel('W m$^{-2}$')
ax_aci.set_ylabel('W m$^{-2}$')
ax_main.text(2506, forcing['ssp585']['total'][750], 'SSP5-8.5', color=colors['ssp585'], va='center', ha='left')
ax_main.text(2506, forcing['ssp370']['total'][750], 'SSP3-7.0', color=colors['ssp370'], va='center', ha='left')
ax_main.text(2499, forcing['ssp370-lowNTCF']['total'][750]+.2, 'SSP3-7.0-lowNTCF', color=colors['ssp370'], va='bottom', ha='right', rotation=-3)
ax_main.text(2499, forcing['ssp370-lowNTCFCH4']['total'][750]+0.05, 'SSP3-7.0-lowNTCFCH4', color=colors['ssp370'], va='top', ha='right', rotation=-3)
ax_main.text(2506, forcing['ssp460']['total'][750], 'SSP4-6.0', color=colors['ssp460'], va='center', ha='left')
ax_main.text(2506, forcing['ssp245']['total'][750], 'SSP2-4.5', color=colors['ssp245'], va='center', ha='left')
ax_main.text(2506, forcing['ssp434']['total'][750], 'SSP4-3.4', color=colors['ssp434'], va='center', ha='left')
#ax_main.text(2499, forcing['ssp534-over']['total'][700], 'SSP5-3.4-over', color=colors['ssp534-over'], va='bottom', ha='right')
ax_main.text(2506, forcing['ssp126']['total'][750], 'SSP1-2.6', color=colors['ssp126'], va='center', ha='left')
ax_main.text(2506, forcing['ssp119']['total'][750], 'SSP1-1.9', color=colors['ssp119'], va='center', ha='left')
ax_main.annotate('SSP5-3.4-over', arrowprops={'arrowstyle':'->'}, xy=(2075, forcing['ssp534-over']['total'][325]), xytext=(2130, forcing['ssp534-over']['total'][325]), color=colors['ssp534-over'], va='center')
#ax_main.annotate('ssp370-lowNTCFCH4', arrowprops={'arrowstyle':'->'}, xy=(2400, forcing['ssp370-lowNTCFCH4']['total'][650]), xytext=(2400, 9), color=colors['ssp370-lowNTCFCH4'], va='center', ha='center')
#ax_main.annotate('ssp370-lowNTCF', arrowprops={'arrowstyle':'->'}, xy=(2100, forcing['ssp370-lowNTCF']['total'][350]), xytext=(2050, 12), color=colors['ssp370-lowNTCF'], va='center', ha='center')
pl.tight_layout()
pl.savefig('../figures/fig7.SM.1.png', dpi=300)
pl.savefig('../figures/fig7.SM.1.pdf')
#pl.savefig('/nfs/see-fs-02_users/mencsm/ssp_erf/total.png')
```
| github_jupyter |
```
import os
import glob
import numpy as np
from shutil import copyfile
symlink = True # If this is false the files are copied instead
combine_train_valid = False # If this is true, the train and valid sets are ALSO combined
```
# CIFAR-10 constituent samples' extraction
This notebook shows how to construct a dataset that has only CIFAR samples. This can be used for other tasks or for assessment of models trained on the imagenet constituents, to understand how well these models deal with distribution shift.
#### ENSURE THAT CINIC-10 IS DOWNLOADED AND STORED IN ../data/cinic-10
```
cinic_directory = "../data/cinic-10"
cifar_directory = "../data/cinic-10-cifar"
classes = ["airplane", "automobile", "bird", "cat", "deer", "dog", "frog", "horse", "ship", "truck"]
sets = ['train', 'valid', 'test']
if not os.path.exists(cifar_directory):
os.makedirs(cifar_directory)
if not os.path.exists(cifar_directory + '/train'):
os.makedirs(cifar_directory + '/train')
if not os.path.exists(cifar_directory + '/test'):
os.makedirs(cifar_directory + '/test')
for c in classes:
if not os.path.exists('{}/train/{}'.format(cifar_directory, c)):
os.makedirs('{}/train/{}'.format(cifar_directory, c))
if not os.path.exists('{}/test/{}'.format(cifar_directory, c)):
os.makedirs('{}/test/{}'.format(cifar_directory, c))
if not combine_train_valid:
if not os.path.exists('{}/valid/{}'.format(cifar_directory, c)):
os.makedirs('{}/valid/{}'.format(cifar_directory, c))
for s in sets:
for c in classes:
source_directory = '{}/{}/{}'.format(cinic_directory, s, c)
filenames = glob.glob('{}/*.png'.format(source_directory))
for fn in filenames:
dest_fn = fn.split('/')[-1]
if (s == 'train' or s == 'valid') and combine_train_valid and 'cifar' in fn.split('/')[-1]:
dest_fn = '{}/train/{}/{}'.format(cifar_directory, c, dest_fn)
if symlink:
if not os.path.islink(dest_fn):
os.symlink(fn, dest_fn)
else:
copyfile(fn, dest_fn)
elif (s == 'train') and 'cifar' in fn.split('/')[-1]:
dest_fn = '{}/train/{}/{}'.format(cifar_directory, c, dest_fn)
if symlink:
if not os.path.islink(dest_fn):
os.symlink(fn, dest_fn)
else:
copyfile(fn, dest_fn)
elif (s == 'valid') and 'cifar' in fn.split('/')[-1]:
dest_fn = '{}/valid/{}/{}'.format(cifar_directory, c, dest_fn)
if symlink:
if not os.path.islink(dest_fn):
os.symlink(fn, dest_fn)
else:
copyfile(fn, dest_fn)
elif s == 'test' and 'cifar' in fn.split('/')[-1]:
dest_fn = '{}/test/{}/{}'.format(cifar_directory, c, dest_fn)
if symlink:
if not os.path.islink(dest_fn):
os.symlink(fn, dest_fn)
else:
copyfile(fn, dest_fn)
```
| github_jupyter |
# 用每日新闻预测金融市场变化(标准版)
TF-IDF + SVM 是文本分类问题的基准线
这篇教程我直接用最简单直接的方式处理。
高级版本的教程会在日后的课程中放出。
```
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
import pandas as pd
import numpy as np
from sklearn.svm import SVC
from sklearn.metrics import roc_auc_score
from datetime import date
```
### 监视数据
我们先读入数据。这里我提供了一个已经combine好了的数据。
```
data = pd.read_csv('../input/Combined_News_DJIA.csv')
```
这时候,我们可以看一下数据长什么样子
```
data.head()
```
其实看起来特别的简单直观。如果是1,那么当日的DJIA就提高或者不变了。如果是1,那么DJIA那天就是跌了。
接下来我们把headlines先合并起来。因为我们显然是需要考虑所有的news的。
```
data["combined_news"] = data.filter(regex=("Top.*")).apply(lambda x: ''.join(str(x.values)), axis=1)
```
### 分割测试/训练集
这下,我们可以把数据给分成Training/Testing data
```
train = data[data['Date'] < '2015-01-01']
test = data[data['Date'] > '2014-12-31']
```
### 提取features
也就是文本中的特征
这里一定要注意,fit你的model的时候,要用training set,不能一股脑的把所有的数据都放进来。(当然,现实中你可以这么做)因为我们要假设testing set我们在训练的时候是完全接触不到的,是不可知的。
```
feature_extraction = TfidfVectorizer()
X_train = feature_extraction.fit_transform(train["combined_news"].values)
```
所以*X_train* 搞完以后,直接给 *X_test* 做个Transform
```
X_test = feature_extraction.transform(test["combined_news"].values)
```
同理,y就是我们已经准备好的label
```
y_train = train["Label"].values
y_test = test["Label"].values
```
### 训练模型
```
clf = SVC(probability=True, kernel='rbf')
```
把你的*X_train*和*y_train*给fit进去
```
clf.fit(X_train, y_train)
```
### 预测
```
predictions = clf.predict_proba(X_test)
```
### 验证准确度
按照我之前给的要求,用AUC作为binary classification的Metrics
```
print('ROC-AUC yields ' + str(roc_auc_score(y_test, predictions[:,1])))
```
## 进阶版
### 文本预处理
我们这样直接把文本放进TF-IDF,虽然简单方便,但是还是不够严谨的。
我们可以把原文本做进一步的处理。
+ 小写 / 分成小tokens
```
X_train = train["combined_news"].str.lower().str.replace('"', '').str.replace("'", '').str.split()
X_test = test["combined_news"].str.lower().str.replace('"', '').str.replace("'", '').str.split()
print(X_test[1611])
```
+ 删减停止词
```
from nltk.corpus import stopwords
stop = stopwords.words('english')
```
+ 删除数字
```
import re
def hasNumbers(inputString):
return bool(re.search(r'\d', inputString))
```
+ lemma
```
from nltk.stem import WordNetLemmatizer
wordnet_lemmatizer = WordNetLemmatizer()
```
我们把这些元素全都合成一个func
```
def check(word):
"""
如果需要这个单词,则True
如果应该去除,则False
"""
if word in stop:
return False
elif hasNumbers(word):
return False
else:
return True
```
然后我们把整个流程放进我们的DF中处理
```
X_train = X_train.apply(lambda x: [wordnet_lemmatizer.lemmatize(item) for item in x if check(item)])
X_test = X_test.apply(lambda x: [wordnet_lemmatizer.lemmatize(item) for item in x if check(item)])
print(X_test[1611])
```
因为外部库,比如sklearn 只支持string输入,所以我们把调整后的list再变回string
```
X_train = X_train.apply(lambda x: ' '.join(x))
X_test = X_test.apply(lambda x: ' '.join(x))
print(X_test[1611])
```
重新Fit一遍我们的clf
```
feature_extraction = TfidfVectorizer(lowercase=False)
X_train = feature_extraction.fit_transform(X_train.values)
X_test = feature_extraction.transform(X_test.values)
```
再跑一遍
```
clf = SVC(probability=True, kernel='rbf')
clf.fit(X_train, y_train)
predictions = clf.predict_proba(X_test)
print('ROC-AUC yields ' + str(roc_auc_score(y_test, predictions[:,1])))
```
不小心发现,折腾一圈以后,这个结果还不如之前的简单版。
这个故事告诉我们,**逼装得太多,不一定有卵用。**
造成如此的原因有几种:
+ 数据点太少
在大量的数据下,标准的文本预处理流程还是需要的,以提高机器学习的准确度。
+ One-Off result
我们到现在都只是跑了一次而已。如果我们像前面的例子一样,用Cross Validation来玩这组数据,说不定我们会发现,分数高的clf其实是overfitted了的。
所以,我们在做kaggle竞赛的时候,最好是要给自己的clf做好CV验证。不要贻笑大方。
| github_jupyter |
# Segmentation of Road from Satellite imagery
## Importing Libraries
```
import warnings
warnings.filterwarnings('ignore')
import os
import cv2
#from google.colab.patches import cv2_imshow
import numpy as np
import tensorflow as tf
import pandas as pd
from keras.models import Model, load_model
from skimage.morphology import label
import pickle
from keras import backend as K
from matplotlib import pyplot as plt
from tqdm import tqdm_notebook
import random
from skimage.io import imread, imshow, imread_collection, concatenate_images
from matplotlib import pyplot as plt
import h5py
seed = 56
!pip install tensorflow==1.14.0
!pip install -U segmentation-models
from google.colab import drive
drive.mount('/content/gdrive/')
base_path = "gdrive/My\ Drive/MapSegClean/"
%cd gdrive/My\ Drive/MapSegClean/
```
## Defining Custom Loss functions and accuracy Metric.
```
#Source: https://towardsdatascience.com/metrics-to-evaluate-your-semantic-segmentation-model-6bcb99639aa2
from keras import backend as K
def iou_coef(y_true, y_pred, smooth=1):
intersection = K.sum(K.abs(y_true * y_pred), axis=[1,2,3])
union = K.sum(y_true,[1,2,3])+K.sum(y_pred,[1,2,3])-intersection
iou = K.mean((intersection + smooth) / (union + smooth), axis=0)
return iou
def dice_coef(y_true, y_pred, smooth = 1):
y_true_f = K.flatten(y_true)
y_pred_f = K.flatten(y_pred)
intersection = K.sum(y_true_f * y_pred_f)
return (2. * intersection + smooth) / (K.sum(y_true_f) + K.sum(y_pred_f) + smooth)
def soft_dice_loss(y_true, y_pred):
return 1-dice_coef(y_true, y_pred)
```
## Defining Our Model
```
from segmentation_models import PSPNet
from segmentation_models import get_preprocessing
from segmentation_models.losses import bce_jaccard_loss
from segmentation_models.metrics import iou_score
model = PSPNet(input_shape=(240, 240, 3), encoder_weights=None, classes=1, psp_use_batchnorm=True, psp_pooling_type='max', activation='sigmoid')
#model = Unet(input_shape=(256, 256, 3), weights=None, activation='elu')
model.summary()
```
### On Test Images
```
model.load_weights("/content/gdrive/My Drive/MapSegClean/Models/Final_PSPNet_road_weights.h5")
import cv2
import glob
import numpy as np
import h5py
#test_images = np.array([cv2.imread(file) for file in glob.glob("/home/bisag/Desktop/Road-Segmentation/I/")])
#test_masks = np.array([cv2.imread(file) for file in glob.glob("/home/bisag/Desktop/Road-Segmentation/M/")])
test_masks = []
test_images = []
files = glob.glob ("TestI/*.png")
for myFile in files:
print(myFile)
image = cv2.imread (myFile)
test_images.append (image[0:240,0:240])
myFile = 'TestM' + myFile[5:len(myFile)]
image = cv2.cvtColor(cv2.imread (myFile), cv2.COLOR_BGR2GRAY)
test_masks.append (image[0:240,0:240])
#files = glob.glob ("TestM/*.png")
#for myFile in files:
# print(myFile)
#test_images = cv2.imread("/home/bisag/Desktop/Road-Segmentation/I/1.png")
#test_masks = cv2.imread("/home/bisag/Desktop/Road-Segmentation/M/1.png")
test_images = np.array(test_images)
test_masks = np.array(test_masks)
test_masks = np.expand_dims(test_masks, -1)
print("Unique elements in the train mask:", np.unique(test_masks))
print(test_images.shape)
print(test_masks.shape)
predictions = model.predict(test_images, verbose=1)*255
thresh_val = 0.1
predicton_threshold = (predictions > thresh_val).astype(np.uint8)
import numpy as np
from google.colab.patches import cv2_imshow
cv2_imshow(np.squeeze(test_images[14]))
cv2_imshow(np.squeeze(predictions[14]))
cv2_imshow(np.squeeze(test_masks[14]))
import matplotlib
for i in range(len(predictions)):
print(i)
cv2.imwrite( "Results/" + str(i) + "Image.png" , test_images[i])
cv2.imwrite( "Results/" + str(i) + "GroundTruth.png" , test_masks[i])
#cv2.imwrite( "/home/bisag/Desktop/Road-Segmentation/Results/" + str(i) + "Prediction.png" , np.squeeze(predictions[i][:,:,0]))
#cv2.imwrite( "/home/bisag/Desktop/Road-Segmentation/Results/" + str(i) + "Prediction_Threshold.png" , np.squeeze(predicton_threshold[i][:,:,0]))
#matplotlib.image.imsave('/home/bisag/Desktop/Road-Segmentation/Results/000.png', np.squeeze(predicton_threshold[0][:,:,0]))
matplotlib.image.imsave("Results/" + str(i) + "Prediction.png" , np.squeeze(predictions[i][:,:,0]))
matplotlib.image.imsave( "Results/" + str(i) + "Prediction_Threshold.png" , np.squeeze(predicton_threshold[i][:,:,0]))
#imshow(np.squeeze(predictions[0][:,:,0]))
#import scipy.misc
#scipy.misc.imsave('/home/bisag/Desktop/Road-Segmentation/Results/00.png', np.squeeze(predictions[0][:,:,0]))
```
| github_jupyter |
# Inference and Validation
Now that you have a trained network, you can use it for making predictions. This is typically called **inference**, a term borrowed from statistics. However, neural networks have a tendency to perform *too well* on the training data and aren't able to generalize to data that hasn't been seen before. This is called **overfitting** and it impairs inference performance. To test for overfitting while training, we measure the performance on data not in the training set called the **validation** set. We avoid overfitting through regularization such as dropout while monitoring the validation performance during training. In this notebook, I'll show you how to do this in PyTorch.
As usual, let's start by loading the dataset through torchvision. You'll learn more about torchvision and loading data in a later part. This time we'll be taking advantage of the test set which you can get by setting `train=False` here:
```python
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
```
The test set contains images just like the training set. Typically you'll see 10-20% of the original dataset held out for testing and validation with the rest being used for training.
```
import torch
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# Download and load the training data
trainset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
```
Here I'll create a model like normal, using the same one from my solution for part 4.
```
from torch import nn, optim
import torch.nn.functional as F
class Classifier(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(784, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 64)
self.fc4 = nn.Linear(64, 10)
def forward(self, x):
# make sure input tensor is flattened
x = x.view(x.shape[0], -1)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = F.log_softmax(self.fc4(x), dim=1)
return x
```
The goal of validation is to measure the model's performance on data that isn't part of the training set. Performance here is up to the developer to define though. Typically this is just accuracy, the percentage of classes the network predicted correctly. Other options are [precision and recall](https://en.wikipedia.org/wiki/Precision_and_recall#Definition_(classification_context)) and top-5 error rate. We'll focus on accuracy here. First I'll do a forward pass with one batch from the test set.
```
model = Classifier()
images, labels = next(iter(testloader))
# Get the class probabilities
ps = torch.exp(model(images))
# Make sure the shape is appropriate, we should get 10 class probabilities for 64 examples
print(ps.shape)
```
With the probabilities, we can get the most likely class using the `ps.topk` method. This returns the $k$ highest values. Since we just want the most likely class, we can use `ps.topk(1)`. This returns a tuple of the top-$k$ values and the top-$k$ indices. If the highest value is the fifth element, we'll get back 4 as the index.
```
top_p, top_class = ps.topk(1, dim=1)
# Look at the most likely classes for the first 10 examples
print(top_class[:10,:])
```
Now we can check if the predicted classes match the labels. This is simple to do by equating `top_class` and `labels`, but we have to be careful of the shapes. Here `top_class` is a 2D tensor with shape `(64, 1)` while `labels` is 1D with shape `(64)`. To get the equality to work out the way we want, `top_class` and `labels` must have the same shape.
If we do
```python
equals = top_class == labels
```
`equals` will have shape `(64, 64)`, try it yourself. What it's doing is comparing the one element in each row of `top_class` with each element in `labels` which returns 64 True/False boolean values for each row.
```
equals = top_class == labels.view(*top_class.shape)
```
Now we need to calculate the percentage of correct predictions. `equals` has binary values, either 0 or 1. This means that if we just sum up all the values and divide by the number of values, we get the percentage of correct predictions. This is the same operation as taking the mean, so we can get the accuracy with a call to `torch.mean`. If only it was that simple. If you try `torch.mean(equals)`, you'll get an error
```
RuntimeError: mean is not implemented for type torch.ByteTensor
```
This happens because `equals` has type `torch.ByteTensor` but `torch.mean` isn't implemented for tensors with that type. So we'll need to convert `equals` to a float tensor. Note that when we take `torch.mean` it returns a scalar tensor, to get the actual value as a float we'll need to do `accuracy.item()`.
```
accuracy = torch.mean(equals.type(torch.FloatTensor))
print(f'Accuracy: {accuracy.item()*100}%')
```
The network is untrained so it's making random guesses and we should see an accuracy around 10%. Now let's train our network and include our validation pass so we can measure how well the network is performing on the test set. Since we're not updating our parameters in the validation pass, we can speed up our code by turning off gradients using `torch.no_grad()`:
```python
# turn off gradients
with torch.no_grad():
# validation pass here
for images, labels in testloader:
...
```
>**Exercise:** Implement the validation loop below and print out the total accuracy after the loop. You can largely copy and paste the code from above, but I suggest typing it in because writing it out yourself is essential for building the skill. In general you'll always learn more by typing it rather than copy-pasting. You should be able to get an accuracy above 80%.
```
model = Classifier()
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
epochs = 30
steps = 0
train_losses, test_losses = [], []
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
optimizer.zero_grad()
log_ps = model(images)
loss = criterion(log_ps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
test_loss = 0
accuracy = 0
# Turn off gradients for validation, saves memory and computations
with torch.no_grad():
for images, labels in testloader:
log_ps = model(images)
test_loss += criterion(log_ps, labels)
ps = torch.exp(log_ps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor))
train_losses.append(running_loss/len(trainloader))
test_losses.append(test_loss/len(testloader))
print("Epoch: {}/{}.. ".format(e+1, epochs),
"Training Loss: {:.3f}.. ".format(running_loss/len(trainloader)),
"Test Loss: {:.3f}.. ".format(test_loss/len(testloader)),
"Test Accuracy: {:.3f}".format(accuracy/len(testloader)))
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
plt.plot(train_losses, label='Training loss')
plt.plot(test_losses, label='Validation loss')
plt.legend(frameon=False)
```
## Overfitting
If we look at the training and validation losses as we train the network, we can see a phenomenon known as overfitting.
<img src='assets/overfitting.png' width=450px>
The network learns the training set better and better, resulting in lower training losses. However, it starts having problems generalizing to data outside the training set leading to the validation loss increasing. The ultimate goal of any deep learning model is to make predictions on new data, so we should strive to get the lowest validation loss possible. One option is to use the version of the model with the lowest validation loss, here the one around 8-10 training epochs. This strategy is called *early-stopping*. In practice, you'd save the model frequently as you're training then later choose the model with the lowest validation loss.
The most common method to reduce overfitting (outside of early-stopping) is *dropout*, where we randomly drop input units. This forces the network to share information between weights, increasing it's ability to generalize to new data. Adding dropout in PyTorch is straightforward using the [`nn.Dropout`](https://pytorch.org/docs/stable/nn.html#torch.nn.Dropout) module.
```python
class Classifier(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(784, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 64)
self.fc4 = nn.Linear(64, 10)
# Dropout module with 0.2 drop probability
self.dropout = nn.Dropout(p=0.2)
def forward(self, x):
# make sure input tensor is flattened
x = x.view(x.shape[0], -1)
# Now with dropout
x = self.dropout(F.relu(self.fc1(x)))
x = self.dropout(F.relu(self.fc2(x)))
x = self.dropout(F.relu(self.fc3(x)))
# output so no dropout here
x = F.log_softmax(self.fc4(x), dim=1)
return x
```
During training we want to use dropout to prevent overfitting, but during inference we want to use the entire network. So, we need to turn off dropout during validation, testing, and whenever we're using the network to make predictions. To do this, you use `model.eval()`. This sets the model to evaluation mode where the dropout probability is 0. You can turn dropout back on by setting the model to train mode with `model.train()`. In general, the pattern for the validation loop will look like this, where you turn off gradients, set the model to evaluation mode, calculate the validation loss and metric, then set the model back to train mode.
```python
# turn off gradients
with torch.no_grad():
# set model to evaluation mode
model.eval()
# validation pass here
for images, labels in testloader:
...
# set model back to train mode
model.train()
```
> **Exercise:** Add dropout to your model and train it on Fashion-MNIST again. See if you can get a lower validation loss or higher accuracy.
```
## TODO: Define your model with dropout added
class Classifier(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(784, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 64)
self.fc4 = nn.Linear(64, 10)
# Dropout module with 0.2 drop probability
self.dropout = nn.Dropout(p=0.2)
def forward(self, x):
# make sure input tensor is flattened
x = x.view(x.shape[0], -1)
# Now with dropout
x = self.dropout(F.relu(self.fc1(x)))
x = self.dropout(F.relu(self.fc2(x)))
x = self.dropout(F.relu(self.fc3(x)))
# output so no dropout here
x = F.log_softmax(self.fc4(x), dim=1)
return x
## TODO: Train your model with dropout, and monitor the training progress with the validation loss and accuracy
model = Classifier()
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
epochs = 30
steps = 0
train_losses, test_losses = [], []
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
optimizer.zero_grad()
log_ps = model(images)
loss = criterion(log_ps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
test_loss = 0
accuracy = 0
# Turn off gradients for validation, saves memory and computations
with torch.no_grad():
model.eval()
for images, labels in testloader:
log_ps = model(images)
test_loss += criterion(log_ps, labels)
ps = torch.exp(log_ps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor))
model.train()
train_losses.append(running_loss/len(trainloader))
test_losses.append(test_loss/len(testloader))
print("Epoch: {}/{}.. ".format(e+1, epochs),
"Training Loss: {:.3f}.. ".format(running_loss/len(trainloader)),
"Test Loss: {:.3f}.. ".format(test_loss/len(testloader)),
"Test Accuracy: {:.3f}".format(accuracy/len(testloader)))
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
plt.plot(train_losses, label='Training loss')
plt.plot(test_losses, label='Validation loss')
plt.legend(frameon=False)
```
## Inference
Now that the model is trained, we can use it for inference. We've done this before, but now we need to remember to set the model in inference mode with `model.eval()`. You'll also want to turn off autograd with the `torch.no_grad()` context.
```
# Import helper module (should be in the repo)
import helper
# Test out your network!
model.eval()
dataiter = iter(testloader)
images, labels = dataiter.next()
img = images[0]
# Convert 2D image to 1D vector
img = img.view(1, 784)
# Calculate the class probabilities (softmax) for img
with torch.no_grad():
output = model.forward(img)
ps = torch.exp(output)
# Plot the image and probabilities
helper.view_classify(img.view(1, 28, 28), ps, version='Fashion')
```
## Next Up!
In the next part, I'll show you how to save your trained models. In general, you won't want to train a model everytime you need it. Instead, you'll train once, save it, then load the model when you want to train more or use if for inference.
| github_jupyter |
# Programming with Python
## Control Flow Statements
The control flow of a program determines the order in which lines of code are executed. All else being equal, Python code is executed linearly, in the order that lines appear in the program. However, all is not usually equal, and so the appropriate control flow is frequently specified with the help of control flow statements. These include loops, conditional statements and calls to functions. Let’s look at a few of these here.
### for statements
One way to repeatedly execute a block of statements (*i.e.* loop) is to use a `for` statement. These statements iterate over the number of elements in a specified sequence, according to the following syntax:
```
for letter in 'ciao':
print('give me a {0}'.format(letter.upper()))
```
Recall that strings are simply regarded as sequences of characters. Hence, the above `for` statement loops over each letter, converting each to upper case with the `upper()` method and printing it.
Similarly, as shown in the introduction, **list comprehensions** may be constructed using `for` statements:
```
[i**2 for i in range(10)]
```
Here, the expression loops over `range(10)` -- the sequence from 0 to 9 -- and squares each before placing it in the returned list.
### if statements
As the name implies, `if` statements execute particular sections of code depending on some tested **condition**. For example, to code an absolute value function, one might employ conditional statements:
```
def absval(some_list):
# Create empty list
absolutes = []
# Loop over elements in some_list
for value in some_list:
# Conditional statement
if value<0:
# Negative value
absolutes.append(-value)
else:
# Positive value
absolutes.append(value)
return absolutes
```
Here, each value in `some_list` is tested for the condition that it is negative, in which case it is multiplied by -1, otherwise it is appended as-is.
For conditions that have more than two possible values, the `elif` clause can be used:
```
x = 5
if x < 0:
print('x is negative')
elif x % 2:
print('x is positive and odd')
else:
print('x is even and non-negative')
```
### while statements
A different type of conditional loop is provided by the `while` statement. Rather than iterating a specified number of times, according to a given sequence, `while` executes its block of code repeatedly, until its condition is no longer true.
For example, suppose we want to sample from a truncated normal distribution, where we are only interested in positive-valued samples. The following function is one solution:
```
# Import function
from numpy.random import normal
def truncated_normals(how_many, l):
# Create empty list
values = []
# Loop until we have specified number of samples
while (len(values) < how_many):
# Sample from standard normal
x = normal(0,1)
# Append if not truncateed
if x > l: values.append(x)
return values
truncated_normals(15, 0)
```
This function iteratively samples from a standard normal distribution, and appends it to the output array if it is positive, stopping to return the array once the specified number of values have been added.
Obviously, the body of the `while` statement should contain code that eventually renders the condition false, otherwise the loop will never end! An exception to this is if the body of the statement contains a `break` or `return` statement; in either case, the loop will be interrupted.
## Generators
When a Python functions is called, it creates a namespace for the function, executes the code that comprises the function (creating objects inside the namespace as required), and returns some result to its caller. After the return, everything inside the namespace (including the namespace itself) is gone, and is created anew when the function is called again.
However, one particular class of functions in Python breaks this pattern, returning a value to the caller while still active, and able to return subsequent values as needed. Python ***generators*** employ `yield` statements in place of `return`, allowing a sequence of values to be generated without having to create a new function namespace each time. In other languages, this construct is known as a *coroutine*.
For example, we may want to have a function that returns a sequence of values; let's consider, for a simple illustration, the Fibonacci sequence:
$$F_i = F_{i-2} + F_{i-1}$$
its certaintly possible to write a standard Python function that returns however many Fibonacci numbers that we need:
```
import numpy as np
def fibonacci(size):
F = np.empty(size, 'int')
a, b = 0, 1
for i in range(size):
F[i] = a
a, b = b, a + b
return F
```
and this works just fine:
```
fibonacci(20)
```
However, what if we need one number at a time, or if we need a million or 10 million values? In the first case, you would somehow have to store the values from the last iteration, and restore the state to the function each time it is called. In the second case, you would have to generate and then store a very large number of values, most of which you may not need right now.
A more sensible solution is to create a `generator`, which calculates a single value in the sequence, then *returns control back to the caller*. This allows the generator to be called again, resuming the sequence generation where it left off. Here's the Fibonacci function, implemented as a generator:
```
def gfibonacci(size):
a, b = 0, 1
for _ in range(size):
yield a
a, b = b, a + b
```
Notice that there is no `return` statement at all; just `yield`, which is where a value is returned each time one is requested. The `yield` statement is what defines a generator.
When we call our generator, rather than a sequence of Fibonacci numbers, we get a generator object:
```
f = gfibonacci(100)
f
```
A generator has a `__next__()` method that can be called via the builtin function `next()`. The call to `next` executes the generator until the `yield` statement is reached, returning the next generated value, and then pausing until another call to `next` occurs:
```
next(f), next(f), next(f)
```
A generator is a type of `iterator`. If we call a function that supports iterables using a generator as an argument, it will know how to use the generator.
```
np.array(list(f))
```
What happens when we reach the "end" of a generator?
```
a_few_fibs = gfibonacci(2)
next(a_few_fibs)
next(a_few_fibs)
next(a_few_fibs)
```
Thus, generators signal when there are no further values to generate by throwing a `StopIteration` exception. We must either handle this exception, or create a generator that is infinite, which we can do in this example by replacing a `for` loop with a `while` loop:
```
def infinite_fib():
a, b = 0, 1
while True:
yield a
a, b = b, a + b
f = infinite_fib()
vals = [next(f) for _ in range(10000)]
vals[-1]
```
## Error Handling
Inevitably, some code you write will generate errors, at least in some situations. Unless we explicitly anticipate and **handle** these errors, they will cause your code to halt (sometimes this is a good thing!). Errors are handled using `try/except` blocks.
If code executed in the `try` block generates an error, code execution moves to the `except` block. If the exception that is specified corresponsd to that which has been raised, the code in the `except` block is executed before continuing; otherwise, the exception is carried out and the code is halted.
```
absval(-5)
```
In the call to `absval`, we passed a single negative integer, whereas the function expects some sort of iterable data structure. Other than changing the function itself, we can avoid this error using exception handling.
```
x = -5
try:
print(absval(x))
except TypeError:
print('The argument to absval must be iterable!')
x = -5
try:
print(absval(x))
except TypeError:
print(absval([x]))
```
We can raise exceptions manually by using the `raise` expression.
```
raise ValueError('This is the wrong value')
```
## Importing and Manipulating Data
Python includes operations for importing and exporting data from files and binary objects, and third-party packages exist for database connectivity. The easiest way to import data from a file is to parse **delimited** text file, which can usually be exported from spreadsheets and databases. In fact, file is a built-in type in python. Data may be read from and written to regular files by specifying them as file objects:
```
microbiome = open('../data/microbiome.csv')
```
Here, a file containing microbiome data in a comma-delimited format is opened, and assigned to an object, called `microbiome`. The next step is to transfer the information in the file to a usable data structure in Python. Since this dataset contains four variables, the name of the taxon, the patient identifier (de-identified), the bacteria count in tissue and the bacteria count in stool, it is convenient to use a dictionary. This allows each variable to be specified by name.
First, a dictionary object is initialized, with appropriate keys and corresponding lists, initially empty. Since the file has a header, we can use it to generate an empty dict:
```
column_names = next(microbiome).rstrip('\n').split(',')
column_names
```
> **Compatibility Corner**: In Python 2, `open` would not return a generator, but rather a `file` object with a `next` method. In Python 3, an generator is returned, which requires the use of the built-in function `next`.
```
mb_dict = {name:[] for name in column_names}
mb_dict
for line in microbiome:
taxon, patient, group, tissue, stool = line.rstrip('\n').split(',')
mb_dict['Taxon'].append(taxon)
mb_dict['Patient'].append(int(patient))
mb_dict['Group'].append(int(group))
mb_dict['Tissue'].append(int(tissue))
mb_dict['Stool'].append(int(stool))
```
For each line in the file, data elements are split by the comma delimiter, using the `split` method that is built-in to string objects. Each datum is subsequently appended to the appropriate list stored in the dictionary. After all the data is parsed, it is polite to close the file:
```
microbiome.close()
```
The data can now be readily accessed by indexing the appropriate variable by name:
```
mb_dict['Tissue'][:10]
```
A second approach to importing data involves interfacing directly with a relational database management system. Relational databases are far more efficient for storing, maintaining and querying data than plain text files or spreadsheets, particularly for large datasets or multiple tables. A number of third parties have created packages for database access in Python. For example, `sqlite3` is a package that provides connectivity for SQLite databases:
```
import sqlite3
db = sqlite3.connect(database='../data/baseball-archive-2011.sqlite')
# create a cursor object to communicate with database
cur = db.cursor()
# run query
cur.execute('SELECT playerid, HR, SB FROM Batting WHERE yearID=1970')
# fetch data, and assign to variable
baseball = cur.fetchall()
baseball[:10]
```
## Functions
Python uses the `def` statement to encapsulate code into a callable function. Here again is a very simple Python function:
```
# Function for calulating the mean of some data
def mean(data):
# Initialize sum to zero
sum_x = 0.0
# Loop over data
for x in data:
# Add to sum
sum_x += x
# Divide by number of elements in list, and return
return sum_x / len(data)
```
As we can see, arguments are specified in parentheses following the function name. If there are sensible "default" values, they can be specified as a **keyword argument**.
```
def var(data, sample=True):
# Get mean of data from function above
x_bar = mean(data)
# Do sum of squares in one line
sum_squares = sum([(x - x_bar)**2 for x in data])
# Divide by n-1 and return
if sample:
return sum_squares/(len(data)-1)
return sum_squares/len(data)
```
Non-keyword arguments must always predede keyword arguments, and must always be presented in order; order is not important for keyword arguments.
Arguments can also be passed to functions as a `tuple`/`list`/`dict` using the asterisk notation.
```
def some_computation(a=-1, b=4.3, c=7):
return (a + b) / float(c)
args = (5, 4, 3)
some_computation(*args)
kwargs = {'b':4, 'a':5, 'c':3}
some_computation(**kwargs)
```
The `lambda` statement creates anonymous one-line functions that can simply be assigned to a name.
```
import numpy as np
normalize = lambda data: (np.array(data) - np.mean(data)) / np.std(data)
```
or not:
```
(lambda data: (np.array(data) - np.mean(data)) / np.std(data))([5,8,3,8,3,1,2,1])
```
Python has several built-in, higher-order functions that are useful.
```
list(filter(lambda x: x > 5, range(10)))
abs([5,-6])
list(map(abs, [5, -6]))
```
## Example: Least Squares Estimation
Lets try coding a statistical function. Suppose we want to estimate the parameters of a simple linear regression model. The objective of regression analysis is to specify an equation that will predict some response variable $Y$ based on a set of predictor variables $X$. This is done by fitting parameter values $\beta$ of a regression model using extant data for $X$ and $Y$. This equation has the form:
$$Y = X\beta + \epsilon$$
where $\epsilon$ is a vector of errors. One way to fit this model is using the method of *least squares*, which is given by:
$$\hat{\beta} = (X^{\prime} X)^{-1}X^{\prime} Y$$
We can write a function that calculates this estimate, with the help of some functions from other modules:
```
from numpy.linalg import inv
from numpy import transpose, array, dot
```
We will call this function `solve`, requiring the predictor and response variables as arguments. For simplicity, we will restrict the function to univariate regression, whereby only a single slope and intercept are estimated:
```
def solve(x,y):
'Estimates regession coefficents from data'
'''
The first step is to specify the design matrix. For this,
we need to create a vector of ones (corresponding to the intercept term,
and along with x, create a n x 2 array:
'''
X = array([[1]*len(x), x])
'''
An array is a data structure from the numpy package, similar to a list,
but allowing for multiple dimensions. Next, we calculate the transpose of x,
using another numpy function, transpose
'''
Xt = transpose(X)
'''
Finally, we use the matrix multiplication function dot, also from numpy
to calculate the dot product. The inverse function is provided by the LinearAlgebra
package. Provided that x is not singular (which would raise an exception), this
yields estimates of the intercept and slope, as an array
'''
b_hat = dot(inv(dot(X,Xt)), dot(X,y))
return b_hat
```
Here is solve in action:
```
solve((10,5,10,11,14),(-4,3,0,23,0.6))
```
## Object-oriented Programming
As previously stated, Python is an object-oriented programming (OOP) language, in contrast to procedural languages like FORTRAN and C. As the name implies, object-oriented languages employ objects to create convenient abstractions of data structures. This allows for more flexible programs, fewer lines of code, and a more natural programming paradigm in general. An object is simply a modular unit of data and associated functions, related to the state and behavior, respectively, of some abstract entity. Object-oriented languages group similar objects into classes. For example, consider a Python class representing a bird:
```
class Bird:
# Class representing a bird
name = 'bird'
def __init__(self, sex):
# Initialization method
self.sex = sex
def fly(self):
# Makes bird fly
print('Flying!')
def nest(self):
# Makes bird build nest
print('Building nest ...')
@classmethod
def get_name(cls):
# Class methods are shared among instances
return cls.name
```
You will notice that this `bird` class is simply a container for two functions (called *methods* in Python), `fly` and `nest`, as well as one attribute, `name`. The methods represent functions in common with all members of this class. You can run this code in Python, and create birds:
```
Tweety = Bird('male')
Tweety.name
Tweety.fly()
Foghorn = Bird('male')
Foghorn.nest()
```
A `classmethod` can be called without instantiating an object.
```
Bird.get_name()
```
Whereas standard methods cannot:
```
Bird.fly()
```
As many instances of the `bird` class can be generated as desired, though it may quickly become boring. One of the important benefits of using object-oriented classes is code re-use. For example, we may want more specific kinds of birds, with unique functionality:
```
class Duck(Bird):
# Duck is a subclass of bird
name = 'duck'
def swim(self):
# Ducks can swim
print('Swimming!')
def quack(self,n):
# Ducks can quack
print('Quack! ' * n)
```
Notice that this new `duck` class refers to the `bird` class in parentheses after the class declaration; this is called **inheritance**. The subclass `duck` automatically inherits all of the variables and methods of the superclass, but allows new functions or variables to be added. In addition to flying and best-building, our duck can also swim and quack:
```
Daffy = Duck('male')
Daffy.swim()
Daffy.quack(3)
Daffy.nest()
```
Along with adding new variables and methods, a subclass can also override existing variables and methods of the superclass. For example, one might define `fly` in the `duck` subclass to return an entirely different string. It is easy to see how inheritance promotes code re-use, sometimes dramatically reducing development time. Classes which are very similar need not be coded repetitiously, but rather, just extended from a single superclass.
This brief introduction to object-oriented programming is intended only to introduce new users of Python to this programming paradigm. There are many more salient object-oriented topics, including interfaces, composition, and introspection. I encourage interested readers to refer to any number of current Python and OOP books for a more comprehensive treatment.
## In Python, everything is an object
Everything (and I mean *everything*) in Python is an object, in the sense that they possess attributes, such as methods and variables, that we usually associate with more "structured" objects like those we created above.
Check it out:
```
dir(1)
(1).bit_length()
```
This has implications for how assignment works in Python.
Let's create a trivial class:
```
class Thing: pass
```
and instantiate it:
```
x = Thing()
x
```
Here, `x` is simply a "label" for the object that we created when calling `Thing`. That object resides at the memory location that is identified when we print `x`. Notice that if we create another `Thing`, we create an new object, and give it a label. We know it is a new object because it has its own memory location.
```
y = Thing()
y
```
What if we assign `x` to `z`?
```
z = x
z
```
We see that the object labeled with `z` is the same as the object as that labeled with `x`. So, we say that `z` is a label (or name) with a *binding* to the object created by `Thing`.
So, there are no "variables", in the sense of a container for values, in Python. There are only labels and bindings.
```
x.name = 'thing x'
z.name
```
This can get you into trouble. Consider the following (seemingly inoccuous) way of creating a dictionary of emtpy lists:
```
evil_dict = dict.fromkeys(column_names, [])
evil_dict
evil_dict['Tissue'].append(5)
evil_dict
```
Why did this happen?
## References
* [Learn Python the Hard Way](http://learnpythonthehardway.org/book/)
* [Learn X in Y Minutes (where X=Python)](http://learnxinyminutes.com/docs/python/)
* [29 common beginner Python errors on one page](http://pythonforbiologists.com/index.php/29-common-beginner-python-errors-on-one-page/)
* [Understanding Python's Execution Model](http://www.jeffknupp.com/blog/2013/02/14/drastically-improve-your-python-understanding-pythons-execution-model/)
| github_jupyter |
# Statistics for Hackers Created by Github@croach
### An Exploration of Statistics Through Computational Simulation
#### A [talk][video] by [Jake VanDerPlas][jakevdp] for PyCon 2016
#### [Slides][slides] available on speakerdeck
## Motivation
There's no shortage of absolutely magnificent material out there on the topics of data science and machine learning for an autodidact, such as myself, to learn from. In fact, so many great resources exist that an individual can be forgiven for not knowing where to begin their studies, or for getting distracted once they're off the starting block. I honestly can't count the number of times that I've started working through many of these online courses and tutorials only to have my attention stolen by one of the multitudes of amazing articles on data analysis with Python, or some great new [MOOC][mooc] on Deep Learning. But this year is different! This year, for one of my new year's resolutions, I've decided to create a personalized data science curriculum and stick to it. This year, I promise not to just casually sign up for another course, or start reading yet another textbook to be distracted part way through. This year, I'm sticking to the plan.
As part of my personalized program of study, I've chosen to start with [Harvard's Data Science course][cs109]. I'm currently on week 3 and one of the suggested readings for this week is [Jake VanderPlas'][jakevdp] talk from PyCon 2016 titled "Statistics for Hackers". As I was watching the [video][video] and following along with the [slides][slides], I wanted to try out some of the examples and create a set of notes that I could refer to later, so I figured why not create a Jupyter notebook. Once I'd finished, I realized I'd created a decently-sized resource that could be of use to others working their way through the talk. The result is the article you're reading right now, the remainder of which contains my notes and code examples for Jake's excellent talk.
So, enjoy the article, I hope you find this resource useful, and if you have any problems or suggestions of any kind, the full [notebook][notebook] can be found on [github][github], so please send me a [pull request][github_pulls], or submit an [issue][github_issues], or just message me directly on [Twitter][twitter].
[mooc]: https://en.wikipedia.org/wiki/Massive_open_online_course
[cs109]: http://cs109.github.io/2015/index.html
[video]: https://youtu.be/Iq9DzN6mvYA
[slides]: https://speakerdeck.com/jakevdp/statistics-for-hackers
[jakevdp]: https://staff.washington.edu/jakevdp/
[notebook]: http://nbviewer.jupyter.org/github/croach/statistics-for-hackers/blob/master/statistics-for-hackers.ipynb
[github]: https://github.com/croach/statistics-for-hackers
[github_pulls]: https://github.com/croach/statistics-for-hackers/pulls
[github_issues]: https://github.com/croach/statistics-for-hackers/issues
[twitter]: https://twitter.com/vthakr
## Preliminaries
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# Suppress all warnings just to keep the notebook nice and clean.
# This must happen after all imports since numpy actually adds its
# RankWarning class back in.
import warnings
warnings.filterwarnings("ignore")
# Setup the look and feel of the notebook
sns.set_context("notebook",
font_scale=1.5,
rc={"lines.linewidth": 2.5})
sns.set_style('whitegrid')
sns.set_palette('deep')
# Create a couple of colors to use throughout the notebook
red = sns.xkcd_rgb['vermillion']
blue = sns.xkcd_rgb['dark sky blue']
from IPython.display import display
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
```
## Warm-up
The talk starts off with a motivating example that asks the question "If you toss a coin 30 times and see 22 heads, is it a fair coin?"
We all know that a fair coin should come up heads roughly 15 out of 30 tosses, give or take, so it does seem unlikely to see so many heads. However, the skeptic might argue that even a fair coin could show 22 heads in 30 tosses from time-to-time. This could just be a chance event. So, the question would then be "how can you determine if you're tossing a fair coin?"
### The Classic Method
The classic method would assume that the skeptic is correct and would then test the hypothesis (i.e., the [*Null Hypothesis*][null_hypothesis]) that the observation of 22 heads in 30 tosses could happen simply by chance. Let's start by first considering the probability of a single coin flip coming up heads and work our way up to 22 out of 30.
$$
P(H) = \frac{1}{2}
$$
As our equation shows, the probability of a single coin toss turning up heads is exactly 50% since there is an equal chance of either heads or tails turning up. Taking this one step further, to determine the probability of getting 2 heads in a row with 2 coin tosses, we would need to multiply the probability of getting heads by the probability of getting heads again since the two events are independent of one another.
$$
P(HH) = P(H) \cdot P(H) = P(H)^2 = \left(\frac{1}{2}\right)^2 = \frac{1}{4}
$$
From the equation above, we can see that the probability of getting 2 heads in a row from a total of 2 coin tosses is 25%. Let's now take a look at a slightly different scenario and calculate the probability of getting 2 heads and 1 tails with 3 coin tosses.
$$
P(HHT) = P(H)^2 \cdot P(T) = \left(\frac{1}{2}\right)^2 \cdot \frac{1}{2} = \left(\frac{1}{2}\right)^3 = \frac{1}{8}
$$
The equation above tells us that the probability of getting 2 heads and 1 tails in 3 tosses is 12.5%. This is actually the exact same probability as getting heads in all three tosses, which doesn't sound quite right. The problem is that we've only calculated the probability for a single permutation of 2 heads and 1 tails; specifically for the scenario where we only see tails on the third toss. To get the actual probability of tossing 2 heads and 1 tails we will have to add the probabilities for all of the possible permutations, of which there are exactly three: HHT, HTH, and THH.
$$
P(2H,1T) = P(HHT) + P(HTH) + P(THH) = \frac{1}{8} + \frac{1}{8} + \frac{1}{8} = \frac{3}{8}
$$
Another way we could do this is to calculate the total number of permutations and simply multiply that by the probability of each event happening. To get the total number of permutations we can use the [binomial coefficient][binom_coeff]. Then, we can simply calculate the probability above using the following equation.
$$
P(2H,1T) = \binom{3}{2} \left(\frac{1}{2}\right)^{3} = 3 \left(\frac{1}{8}\right) = \frac{3}{8}
$$
While the equation above works in our particular case, where each event has an equal probability of happening, it will run into trouble with events that have an unequal chance of taking place. To deal with those situations, you'll want to extend the last equation to take into account the differing probabilities. The result would be the following equation, where $N$ is number of coin flips, $N_H$ is the number of expected heads, $N_T$ is the number of expected tails, and $P_H$ is the probability of getting heads on each flip.
$$
P(N_H,N_T) = \binom{N}{N_H} \left(P_H\right)^{N_H} \left(1 - P_H\right)^{N_T}
$$
Now that we understand the classic method, let's use it to test our null hypothesis that we *are* actually tossing a fair coin, and that this is just a chance occurrence. The following code implements the equations we've just discussed above.
<!--
The following code sets up a handful of helper functions that we'll use below to create the probability distribution for a fair coin being flipped 30 times using the equations we just discussed above.
-->
[null_hypothesis]: https://en.wikipedia.org/wiki/Null_hypothesis
[binom_coeff]: https://en.wikipedia.org/wiki/Binomial_coefficient
```
def factorial(n):
"""Calculates the factorial of `n`
"""
vals = list(range(1, n + 1))
if len(vals) <= 0:
return 1
prod = 1
for val in vals:
prod *= val
return prod
def n_choose_k(n, k):
"""Calculates the binomial coefficient
"""
return factorial(n) / (factorial(k) * factorial(n - k))
def binom_prob(n, k, p):
"""Returns the probability of see `k` heads in `n` coin tosses
Arguments:
n - number of trials
k - number of trials in which an event took place
p - probability of an event happening
"""
return n_choose_k(n, k) * p**k * (1 - p)**(n - k)
```
Now that we have a method that will calculate the probability for a specific event happening (e.g., 22 heads in 30 coin tosses), we can calculate the probability for every possible outcome of flipping a coin 30 times, and if we plot these values we'll get a visual representation of our coin's probability distribution.
```
# Calculate the probability for every possible outcome of tossing
# a fair coin 30 times.
probabilities = [binom_prob(30, k, 0.5) for k in range(1, 31)]
# Plot the probability distribution using the probabilities list
# we created above.
plt.step(range(1, 31), probabilities, where='mid', color=blue)
plt.xlabel('number of heads')
plt.ylabel('probability')
plt.plot((22, 22), (0, 0.1599), color=red);
plt.annotate('0.8%',
xytext=(25, 0.08),
xy=(22, 0.08),
multialignment='right',
va='center',
color=red,
size='large',
arrowprops={'arrowstyle': '<|-',
'lw': 2,
'color': red,
'shrinkA': 10});
```
The visualization above shows the probability distribution for flipping a fair coin 30 times. Using this visualization we can now determine the probability of getting, say for example, 12 heads in 30 flips, which looks to be about 8%. Notice that we've labeled our example of 22 heads as 0.8%. If we look at the probability of flipping exactly 22 heads, it looks likes to be a little less than 0.8%, in fact if we calculate it using the `binom_prob` function from above, we get 0.5%
```
print("Probability of flipping 22 heads: %0.1f%%" % (binom_prob(30, 22, 0.5) * 100))
```
So, then why do we have 0.8% labeled in our probability distribution above? Well, that's because we are showing the probability of getting **at least** 22 heads, which is also known as the p-value.
#### What's a p-value?
In [statistical hypothesis testing][hypothesis_test] we have an idea that we want to test, but considering that it's very hard to prove something to be true beyond doubt, rather than test our hypothesis directly, we formulate a competing hypothesis, called a [null hypothesis][null_hypothesis], and then try to disprove it instead. The null hypothesis essentially assumes that the effect we're seeing in the data could just be due to chance.
In our example, the null hypothesis assumes we have a fair coin, and the way we determine if this hypothesis is true or not is by calculating how often flipping this fair coin 30 times would result in 22 or more heads. If we then take the number of times that we got 22 or more heads and divide that number by the total of all possible permutations of 30 coin tosses, we get the probability of tossing 22 or more heads with a fair coin. This probability is what we call the [p-value][p_value].
The p-value is used to check the validity of the null hypothesis. The way this is done is by agreeing upon some predetermined upper limit for our p-value, below which we will assume that our null hypothesis is false. In other words, if our null hypothesis were true, and 22 heads in 30 flips could happen often enough by chance, we would expect to see it happen more often than the given threshold percentage of times. So, for example, if we chose 10% as our threshold, then we would expect to see 22 or more heads show up at least 10% of the time to determine that this is a chance occurrence and not due to some bias in the coin. Historically, the generally accepted threshold has been 5%, and so if our p-value is less than 5%, we can then make the assumption that our coin may not be fair.
The `binom_prob` function from above calculates the probability of a single event happening, so now all we need for calculating our p-value is a function that adds up the probabilities of a given event, or a more extreme event happening. So, as an example, we would need a function to add up the probabilities of getting 22 heads, 23 heads, 24 heads, and so on. The next bit of code creates that function and uses it to calculate our p-value.
[null_hypothesis]: https://en.wikipedia.org/wiki/Null_hypothesis
[hypothesis_test]: https://en.wikipedia.org/wiki/Statistical_hypothesis_testing
[p_value]: https://www.statisticsdonewrong.com/data-analysis.html#the-power-of-p-values
```
def p_value(n, k, p):
"""Returns the p-value for the given the given set
"""
return sum(binom_prob(n, i, p) for i in range(k, n+1))
print("P-value: %0.1f%%" % (p_value(30, 22, 0.5) * 100))
```
Running the code above gives us a p-value of roughly 0.8%, which matches the value in our probability distribution above and is also less than the 5% threshold needed to reject our null hypothesis, so it does look like we may have a biased coin.
### The Easier Method
That's an example of using the classic method for testing if our coin is fair or not. However, if you don't happen to have at least some background in statistics, it can be a little hard to follow at times, but luckily for us, there's an easier method...
Simulation!
The code below seeks to answer the same question of whether or not our coin is fair by running a large number of simulated coin flips and calculating the proportion of these experiments that resulted in at least 22 heads or more.
```
M = 0
n = 50000
for i in range(n):
trials = np.random.randint(2, size=30)
if (trials.sum() >= 22):
M += 1
p = M / n
print("Simulated P-value: %0.1f%%" % (p * 100))
```
The result of our simulations is 0.8%, the exact same result we got earlier when we calculated the p-value using the classical method above. So, it definitely looks like it's possible that we have a biased coin since the chances of seeing 22 or more heads in 30 tosses of a fair coin is less than 1%.
## Four Recipes for Hacking Statistics
We've just seen one example of how our hacking skills can make it easy for us to answer questions that typically only a statistician would be able to answer using the classical methods of statistical analysis. This is just one possible method for answering statistical questions using our coding skills, but Jake's talk describes four recipes in total for "hacking statistics", each of which is listed below. The rest of this article will go into each of the remaining techniques in some detail.
1. [Direct Simulation](#Warm-up)
2. [Shuffling](#Shuffling)
3. [Bootstrapping](#Bootstrapping)
4. [Cross Validation](#Cross-Validation)
In the [Warm-up](#Warm-up) section above, we saw an example direct simulation, the first recipe in our tour of statistical hacks. The next example uses the Shuffling method to figure out if there's a statistically significant difference between two different sample populations.
### Shuffling
In this example, we look at the Dr. Seuss story about the Star-belly Sneetches. In this Seussian world, a group of creatures called the Sneetches are divided into two groups: those with stars on their bellies, and those with no "stars upon thars". Over time, the star-bellied sneetches have come to think of themselves as better than the plain-bellied sneetches. As researchers of sneetches, it's our job to uncover whether or not star-bellied sneetches really are better than their plain-bellied cousins.
The first step in answering this question will be to create our experimental data. In the following code snippet we create a dataframe object that contains a set of test scores for both star-bellied and plain-bellied sneetches.
```
import pandas as pd
df = pd.DataFrame({'star': [1, 1, 1, 1, 1, 1, 1, 1] +
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
'score': [84, 72, 57, 46, 63, 76, 99, 91] +
[81, 69, 74, 61, 56, 87, 69, 65, 66, 44, 62, 69]})
df
```
If we then take a look at the average scores for each group of sneetches, we will see that there's a difference in scores of **6.6** between the two groups. So, on average, the star-bellied sneetches performed better on their tests than the plain-bellied sneetches. But, the real question is, is this a significant difference?
```
star_bellied_mean = df[df.star == 1].score.mean()
plain_bellied_mean = df[df.star == 0].score.mean()
print("Star-bellied Sneetches Mean: %2.1f" % star_bellied_mean)
print("Plain-bellied Sneetches Mean: %2.1f" % plain_bellied_mean)
print("Difference: %2.1f" % (star_bellied_mean - plain_bellied_mean))
```
To determine if this is a signficant difference, we could perform a [t-test][t_test] on our data to compute a p-value, and then just make sure that the p-value is less than the target 0.05. Alternatively, we could use simulation instead.
Unlike our first example, however, we don't have a generative function that we can use to create our probability distribution. So, how can we then use simulation to solve our problem?
Well, we can run a bunch of simulations where we randomly shuffle the labels (i.e., star-bellied or plain-bellied) of each sneetch, recompute the difference between the means, and then determine if the proportion of simulations in which the difference was at least as extreme as 6.6 was less than the target 5%. If so, we can conclude that the difference we see is, in fact, one that doesn't occur strictly by chance very often and so the difference is a significant one. In other words, if the proportion of simulations that have a difference of 6.6 or greater is less than 5%, we can conclude that the labels really do matter, and so we can conclude that star-bellied sneetches are "better" than their plain-bellied counterparts.
[t_test]: https://en.wikipedia.org/wiki/Student's_t-test
```
df['label'] = df['star']
np.random.shuffle(df['label'])
df
df['label'] = df['star']
num_simulations = 10000
differences = []
for i in range(num_simulations):
np.random.shuffle(df['label'])
star_bellied_mean = df[df.label == 1].score.mean()
plain_bellied_mean = df[df.label == 0].score.mean()
differences.append(star_bellied_mean - plain_bellied_mean)
```
Now that we've ran our simulations, we can calculate our p-value, which is simply the proportion of simulations that resulted in a difference greater than or equal to 6.6.
$$
p = \frac{N_{>6.6}}{N_{total}} = \frac{1512}{10000} = 0.15
$$
```
p_value = sum(diff >= 6.6 for diff in differences) / num_simulations
print("p-value: %2.2f" % p_value)
```
<!--
Since our p-value is greater than 0.05, we can conclude that the difference in test scores between the two groups is not a significant one. In other words, if having a star on your belly actually mattered, we wouldn't expect to see so many simulations result in the same, or greater, difference as the one in the real sample population.
-->
The following code plots the distribution of the differences we found by running the simulations above. We've also added an annotation that marks where the difference of 6.6 falls in the distribution along with its corresponding p-value.
```
plt.hist(differences, bins=50, color='blue')
plt.xlabel('score difference')
plt.ylabel('number')
plt.plot((6.6, 6.6), (0, 700), color='red');
plt.annotate('%2.f%%' % (p_value * 100),
xytext=(15, 350),
xy=(6.6, 350),
multialignment='right',
va='center',
color='red',
size='large',
arrowprops={'arrowstyle': '<|-',
'lw': 2,
'color': 'red',
'shrinkA': 10});
```
We can see from the histogram above---and from our simulated p-value, which was greater than 5%---that the difference that we are seeing between the populations can be explained by random chance, so we can effectively dismiss the difference as not statistically significant. In short, star-bellied sneetches are no better than the plain-bellied ones, at least not from a statistical point of view.
For further discussion on this method of simulation, check out John Rauser's keynote talk ["Statistics Without the Agonizing Pain"][rauser] from Strata + Hadoop 2014. Jake mentions that he drew inspiration from it in his talk, and it is a really excellent talk as well; I wholeheartedly recommend it.
[rauser]: https://youtu.be/5Dnw46eC-0o
### Bootstrapping
In this example, we'll be using the story of Yertle the Turtle to explore the bootstrapping recipe. As the story goes, in the land of Sala-ma-Sond, Yertle the Turtle was the king of the pond and he wanted to be the most powerful, highest turtle in the land. To achieve this goal, he would stack turtles as high as he could in order to stand upon their backs. As observers of this curious behavior, we've recorded the heights of 20 turtle towers and we've placed them in a dataframe in the following bit of code.
```
df = pd.DataFrame({'heights': [48, 24, 51, 12, 21,
41, 25, 23, 32, 61,
19, 24, 29, 21, 23,
13, 32, 18, 42, 18]})
df
```
The questions we want to answer in this example are: what is the mean height of Yertle's turtle stacks, and what is the uncertainty of this estimate?
#### The Classic Method
The classic method is simply to calculate the sample mean...
$$
\bar{x} = \frac{1}{N} \sum_{i=1}^{N} x_i = 28.9
$$
...and the standard error of the mean.
$$
\sigma_{\bar{x}} = \frac{1}{
\sqrt{N}}\sqrt{\frac{1}{N - 1}
\sum_{i=1}^{N} (x_i - \bar{x})^2
} = 3.0
$$
But, being hackers, we'll be using simulation instead.
Just like in our last example, we are once again faced with the problem of not having a generative model, but unlike the last example, we're not comparing two groups, so we can't just shuffle around labels here, instead we'll use something called [bootstrap][bootstrap] [resampling][resampling].
Bootstrap resampling is a method that simulates several random sample distributions by drawing samples from the current distribution with replacement, i.e., we can draw the same data point more than once. Luckily, pandas makes this super easy with its `sample` function. We simply need to make sure that we pass in `True` for the `replace` argument to sample from our dataset with replacement.
[bootstrap]: https://en.wikipedia.org/wiki/Bootstrapping_(statistics)
[resampling]: https://en.wikipedia.org/wiki/Resampling_(statistics)
```
sample = df.sample(20, replace=True)
display(sample)
print("Mean: %2.2f" % sample.heights.mean())
print("Standard Error: %2.2f" % (sample.heights.std() / np.sqrt(len(sample))))
```
More than likely the mean and standard error from our freshly drawn sample above didn't exactly match the one that we calculated using the classic method beforehand. But, if we continue to resample several thousand times and take a look at the average (mean) of all those sample means and their standard deviation, we should have something that very closely approximates the mean and standard error derived from using the classic method above.
```
xbar = []
for i in range(10000):
sample = df.sample(20, replace=True)
xbar.append(sample.heights.mean())
print("Mean: %2.1f" % np.mean(xbar))
print("Standard Error: %2.1f" % np.std(xbar))
```
### Cross Validation
For the final example, we dive into the world of the Lorax. In the story of the Lorax, a faceless creature sells an item that (presumably) all creatures need called a Thneed. Our job as consultants to Onceler Industries is to project Thneed sales. But, before we can get started forecasting the sales of Thneeds, we'll first need some data.
Lucky for you, I've already done the hard work of assembling that data in the code below by "eyeballing" the data in the scatter plot from the slides of the talk. So, it may not be exactly the same, but it should be close enough for our example analysis.
```
df = pd.DataFrame({
'temp': [22, 36, 36, 38, 44, 45, 47,
43, 44, 45, 47, 49,
52, 53, 53, 53, 54, 55, 55, 55, 56, 57, 58, 59,
60, 61, 61.5, 61.7, 61.7, 61.7, 61.8, 62, 62, 63.4, 64.6,
65, 65.6, 65.6, 66.4, 66.9, 67, 67, 67.4, 67.5, 68, 69,
70, 71, 71, 71.5, 72, 72, 72, 72.7, 73, 73, 73, 73.3, 74, 75, 75,
77, 77, 77, 77.4, 77.9, 78, 78, 79,
80, 82, 83, 84, 85, 85, 86, 87, 88,
90, 90, 91, 93, 95, 97,
102, 104],
'sales': [660, 433, 475, 492, 302, 345, 337,
479, 456, 440, 423, 269,
331, 197, 283, 351, 470, 252, 278, 350, 253, 253, 343, 280,
200, 194, 188, 171, 204, 266, 275, 171, 282, 218, 226,
187, 184, 192, 167, 136, 149, 168, 218, 298, 199, 268,
235, 157, 196, 203, 148, 157, 213, 173, 145, 184, 226, 204, 250, 102, 176,
97, 138, 226, 35, 190, 221, 95, 211,
110, 150, 152, 37, 76, 56, 51, 27, 82,
100, 123, 145, 51, 156, 99,
147, 54]
})
```
Now that we have our sales data in a pandas dataframe, we can take a look to see if any trends show up. Plotting the data in a scatterplot, like the one below, reveals that a relationship does seem to exist between temperature and Thneed sales.
```
# Grab a reference to fig and axes object so we can reuse them
fig, ax = plt.subplots()
# Plot the Thneed sales data
ax.scatter(df.temp, df.sales)
ax.set_xlim(xmin=20, xmax=110)
ax.set_ylim(ymin=0, ymax=700)
ax.set_xlabel('temperature (F)')
ax.set_ylabel('thneed sales (daily)');
```
We can see what looks like a relationship between the two variables temperature and sales, but how can we best model that relationship so we can accurately predict sales based on temperature?
Well, one measure of a model's accuracy is the [Root-Mean-Square Error (RMSE)][rmse]. This metric represents the sample standard deviation between a set of predicted values (from our model) and the actual observed values.
[rmse]: https://en.wikipedia.org/wiki/Root-mean-square_deviation
```
def rmse(predictions, targets):
return np.sqrt(((predictions - targets)**2).mean())
```
We can now use our `rmse` function to measure how well our models' accurately represent the Thneed sales dataset. And, in the next cell, we'll give it a try by creating two different models and seeing which one does a better job of fitting our sales data.
```
# 1D Polynomial Fit
d1_model = np.poly1d(np.polyfit(df.temp, df.sales, 1))
d1_predictions = d1_model(range(111))
ax.plot(range(111), d1_predictions,
color=blue, alpha=0.7)
# 2D Polynomial Fit
d2_model = np.poly1d(np.polyfit(df.temp, df.sales, 2))
d2_predictions = d2_model(range(111))
ax.plot(range(111), d2_predictions,
color=red, alpha=0.5)
ax.annotate('RMS error = %2.1f' % rmse(d1_model(df.temp), df.sales),
xy=(75, 650),
fontsize=20,
color=blue,
backgroundcolor='w')
ax.annotate('RMS error = %2.1f' % rmse(d2_model(df.temp), df.sales),
xy=(75, 580),
fontsize=20,
color=red,
backgroundcolor='w')
display(fig);
```
In the figure above, we plotted our sales data along with the two models we created in the previous step. The first model (in blue) is a simple linear model, i.e., a [first-degree polynomial][degree]. The second model (in red) is a second-degree polynomial, so rather than a straight line, we end up with a slight curve.
We can see from the RMSE values in the figure above that the second-degree polynomial performed better than the simple linear model. Of course, the question you should now be asking is, is this the best possible model that we can find?
To find out, let's take a look at the RMSE of a few more models to see if we can do any better.
[degree]: https://en.wikipedia.org/wiki/Degree_of_a_polynomial
```
rmses = []
for deg in range(15):
model = np.poly1d(np.polyfit(df.temp, df.sales, deg))
predictions = model(df.temp)
rmses.append(rmse(predictions, df.sales))
plt.plot(range(15), rmses)
plt.ylim(45, 70)
plt.xlabel('number of terms in fit')
plt.ylabel('rms error')
plt.annotate('$y = a + bx$',
xytext=(14.2, 70),
xy=(1, rmses[1]),
multialignment='right',
va='center',
arrowprops={'arrowstyle': '-|>',
'lw': 1,
'shrinkA': 10,
'shrinkB': 3})
plt.annotate('$y = a + bx + cx^2$',
xytext=(14.2, 64),
xy=(2, rmses[2]),
multialignment='right',
va='top',
arrowprops={'arrowstyle': '-|>',
'lw': 1,
'shrinkA': 35,
'shrinkB': 3})
plt.annotate('$y = a + bx + cx^2 + dx^3$',
xytext=(14.2, 58),
xy=(3, rmses[3]),
multialignment='right',
va='top',
arrowprops={'arrowstyle': '-|>',
'lw': 1,
'shrinkA': 12,
'shrinkB': 3});
```
We can see, from the plot above, that as we increase the number of terms (i.e., the degrees of freedom) in our model we decrease the RMSE, and this behavior can continue indefinitely, or until we have as many terms as we do data points, at which point we would be fitting the data perfectly.
The problem with this approach though, is that as we increase the number of terms in our equation, we simply match the given dataset closer and closer, but what if our model were to see a data point that's not in our training dataset?
As you can see in the plot below, the model that we've created, though it has a very low RMSE, it has so many terms that it matches our current dataset too closely.
```
# Remove everything but the datapoints
ax.lines.clear()
ax.texts.clear()
# Changing the y-axis limits to match the figure in the slides
ax.set_ylim(0, 1000)
# 14 Dimensional Model
model = np.poly1d(np.polyfit(df.temp, df.sales, 14))
ax.plot(range(20, 110), model(range(20, 110)),
color=sns.xkcd_rgb['sky blue'])
display(fig)
```
The problem with fitting the data too closely, is that our model is so finely tuned to our specific dataset, that if we were to use it to predict future sales, it would most likely fail to get very close to the actual value. This phenomenon of too closely modeling the training dataset is well known amongst machine learning practitioners as [overfitting][overfitting] and one way that we can avoid it is to use [cross-validation][cv].
Cross-validation avoids overfitting by splitting the training dataset into several subsets and using each one to train and test multiple models. Then, the RMSE's of each of those models are averaged to give a more likely estimate of how a model of that type would perform on unseen data.
So, let's give it a try by splitting our data into two groups and randomly assigning data points into each one.
[overfitting]: https://en.wikipedia.org/wiki/Overfitting
[cv]: https://en.wikipedia.org/wiki/Cross-validation_(statistics)
```
df_a = df.sample(n=len(df)//2)
df_b = df.drop(df_a.index)
```
We can get a look at the data points assigned to each subset by plotting each one as a different color.
```
plt.scatter(df_a.temp, df_a.sales, color='red')
plt.scatter(df_b.temp, df_b.sales, color='blue')
plt.xlim(0, 110)
plt.ylim(0, 700)
plt.xlabel('temprature (F)')
plt.ylabel('thneed sales (daily)');
```
Then, we'll find the best model for each subset of data. In this particular example, we'll fit a second-degree polynomial to each subset and plot both below.
```
# Create a 2-degree model for each subset of data
m1 = np.poly1d(np.polyfit(df_a.temp, df_a.sales, 2))
m2 = np.poly1d(np.polyfit(df_b.temp, df_b.sales, 2))
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2,
sharex=False, sharey=True,
figsize=(12, 5))
x_min, x_max = 20, 110
y_min, y_max = 0, 700
x = range(x_min, x_max + 1)
# Plot the df_a group
ax1.scatter(df_a.temp, df_a.sales, color='red')
ax1.set_xlim(xmin=x_min, xmax=x_max)
ax1.set_ylim(ymin=y_min, ymax=y_max)
ax1.set_xlabel('temprature (F)')
ax1.set_ylabel('thneed sales (daily)')
ax1.plot(x, m1(x),
color=sns.xkcd_rgb['sky blue'],
alpha=0.7)
# Plot the df_b group
ax2.scatter(df_b.temp, df_b.sales, color='blue')
ax2.set_xlim(xmin=x_min, xmax=x_max)
ax2.set_ylim(ymin=y_min, ymax=y_max)
ax2.set_xlabel('temprature (F)')
ax2.plot(x, m2(x),
color=sns.xkcd_rgb['rose'],
alpha=0.5);
```
Finally, we'll compare models across subsets by calculating the RMSE for each model using the training set for the other model. This will give us two RMSE scores which we'll then average to get a more accurate estimate of how well a second-degree polynomial will perform on any unseen data.
```
print("RMS = %2.1f" % rmse(m1(df_b.temp), df_b.sales))
print("RMS = %2.1f" % rmse(m2(df_a.temp), df_a.sales))
print("RMS estimate = %2.1f" % np.mean([rmse(m1(df_b.temp), df_b.sales),
rmse(m2(df_a.temp), df_a.sales)]))
```
Then, we simply repeat this process for as long as we so desire.
The following code repeats the process described above for polynomials up to 14 degrees and plots the average RMSE for each one against the non-cross-validated RMSE's that we calculated earlier.
```
rmses = []
cross_validated_rmses = []
for deg in range(15):
# df_a the model on the whole dataset and calculate its
# RMSE on the same set of data
model = np.poly1d(np.polyfit(df.temp, df.sales, deg))
predictions = model(df.temp)
rmses.append(rmse(predictions, df.sales))
# Use cross-validation to create the model and df_a it
m1 = np.poly1d(np.polyfit(df_a.temp, df_a.sales, deg))
m2 = np.poly1d(np.polyfit(df_b.temp, df_b.sales, deg))
p1 = m1(df_b.temp)
p2 = m2(df_a.temp)
cross_validated_rmses.append(np.mean([rmse(p1, df_b.sales),
rmse(p2, df_a.sales)]))
plt.plot(range(15), rmses, color=blue,
label='RMS')
plt.plot(range(15), cross_validated_rmses, color=red,
label='cross validated RMS')
plt.ylim(45, 70)
plt.xlabel('number of terms in fit')
plt.ylabel('rms error')
plt.legend(frameon=True)
plt.annotate('Best model minimizes the\ncross-validated error.',
xytext=(7, 60),
xy=(2, cross_validated_rmses[2]),
multialignment='center',
va='top',
color='blue',
size=25,
backgroundcolor='w',
arrowprops={'arrowstyle': '-|>',
'lw': 3,
'shrinkA': 12,
'shrinkB': 3,
'color': 'blue'});
```
According to the graph above, going from a 1-degree to a 2-degree polynomial gives us quite a large improvement overall. But, unlike the RMSE that we calculated against the training set, when using cross-validation we can see that adding more degrees of freedom to our equation quickly reduces the effectiveness of the model against unseen data. This is overfitting in action! In fact, from the looks of the graph above, it would seem that a second-degree polynomial is actually our best bet for this particular dataset.
#### 2-Fold Cross-Validation
Several different methods for performing cross-validation exist, the one we've just seen is called [2-fold cross-validation][2-fold-cv] since the data is split into two subsets. Another close relative is a method called [$k$-fold cross-validation][k-fold-cv]. It differs slightly in that the original dataset is divided into $k$ subsets (instead of just 2), one of which is reserved strictly for testing and the other $k - 1$ subsets are used for training models. This is just one example of an alternate cross-validation method, but more do exist and each one has advantages and drawbacks that you'll need to consider when deciding which method to use.
[2-fold-cv]: https://en.wikipedia.org/wiki/Cross-validation_(statistics)#2-fold_cross-validation
[k-fold-cv]: https://en.wikipedia.org/wiki/Cross-validation_(statistics)#k-fold_cross-validation
## Conclusion
Ok, so that covers nearly everything that Jake covered in his talk. The end of the talk contains a short overview of some important areas that he didn't have time to cover, and there's a nice set of Q&A at the end, but I'll simply direct you to the [video][video] for those parts of the talk. Hopefully, this article/notebook has been helpful to anyone working their way through Jake's talk, and if for some reason, you've read through this entire article and haven't watched the video of the talk yet, I encourage you to take 40 minutes out of your day and go [watch it now][video]---it really is a fantastic talk!
[video]: https://youtu.be/Iq9DzN6mvYA
| github_jupyter |
# Section 4: Feeding the model - parameters estimation and inference
While defining a reasonable architecture for the model(s) is guided by the general principles discussed in the notebooks Section 0 - Section 1, being able to make testable predictions requires to determine, or at least reasonably guess, model parameters. Parameter estimation is a key challenge in computational biomedicine, in particular because only a small fraction of the parameters may be measured “directly”, while others are obtained indirectly from data (“fitted”). The purpose of this section is to go over the important questions that arise when it comes to parameter estimation, and get a qualitative understanding of some common methods used to address these questions, and the associated vocabulary. Rather than a substitute to a full course in data analysis, this notebook shall be viewed as a general introduction to data analysis in the context of mathematical/computational modeling of biological and biomedical systems. It is a collection of tips and thoughts which should help you to handle model parameters in an optimal way.
If the model has been constructed along the guidelines provided in this course, the model structure encompasses a basis of “known” interactors and interactions, and a subset of hypotheses on either new interactors or new interactions which the model allows to test. The model parameters related to the former category of interactors/interactions are generally constrained by published data, and reasonable ranges of values for such parameters can therefore be pulled from the literature, hence we will focus on how to get information on the <b> other parameters, related to the hypothesized interactions </b>.
## The difficulty of parameter measurement and estimation: what a <i> measurement </i> is
Model parameters depend on the type of modeling (see Figure below). Some might be obtained directly or indirectly from data (green boxes). Some might also be inferred from certain types of (microscopic) modeling, to feed higher-level models (blue arrows).

As a crucial introductory point, we stress that <b> the inference of any parameter from a biological measurement is tied to a model, or at least a theoretical framework in which the measurement is interpreted. </b>
This is obvious when a parameter is determined indirectly, by fitting experimental data to a model. Remember the road traffic example where we are interested in knowing the parameter “n=number of cars on highway #1 (Espoo-Helsinki) per hour”, but instead of measuring it directly, we count the number of cars $N$ that leave Espoo towards Helsinki during an hour with no information on the itinerary. If this data is fitted with a road network model including only the main highway #1, then the fit will yield $n=N$ because in the model, all cars leaving Espoo towards Helsinki (the variable measured) take highway #1. If the same data is fitted with another road network model that includes all secondary routes, the model will include the parameter $n$ but the fit will yield $n<N$ since many cars will use other routes within the network. Thus, the <i> same </i> parameter extracted from the <i> same </i> dataset can be <i> very </i> different depending on the model used to fit the data. Therefore, using published model parameters requires caution, and a deep understanding of the theoretical framework in which this parameter has been interpreted from measurements.
But the statement that <b> the inference of any parameter from a biological measurement is tied to a theoretical framework in which the measurement is interpreted </b> is true for <b> any </b> parameter measurement, including <b> what we believe to be direct measurements </b> in targeted experiments. Indeed, we almost <i> never </i> measure directly any biological quantity. We measure, with physical detectors, physical quantities that we believe are associated with the biological quantity, and to quantify this association we use …. <b> models </b>. Let’s go through a few classical examples:
- when we measure a growth curve with a spectrophotometer, we directly measure the amount of light of a certain wavelength absorbed by the cell solution (Optical Density, OD). Then, this amount of light is <b> fitted with a model where the OD is proportional to cell mass </b>. But 1) the constant of proportionality might depend on the type of cells, their chemical content… 2) cell mass is <i> not </i> cell volume, neither cell density, nor cell number, and does not distinguish live and dead cells, and 3) the <b> model itself might become untrue </b> if for instance, throughout its growth, the cell synthesizes a molecule that absorbs light at the same wavelength.
- when we measure concentrations of fluorescently-tagged proteins with fluorescence microscopy, what we directly measure is either photons (if out microscope is equipped with single photon detectors, and we are working in the right illumination regime), or electric signals that are triggered by the interaction of photons with semi-conductors (e.g. avalanche photodiodes). In any case, we’re very far from directly measuring proteins concentrations. But we fit this data with <b> a model where the recorded signal is proportional to protein concentration </b>. Again, the constant of proportionality depends on a lot of things (pH, type of fluorescent tag, existence of previous illumination, protein degradation rate…). And again, the model itself can fail, for instance if we are saturating detectors, or confocal conditions are not ensured…
- the same is true when quantifying proteins using “quantitative” Western blots. What we actually measure is either an optical signal (staining or fluorescent secondary antibodies) or a radioactive signal, but not a total number of proteins. This signal is fit with a <b> model where the signal measured is proportional to the total protein content of the extract loaded on the gel within each band </b>, which itself is believed to be proportional to the amount of proteins in cells…
- when we aim to infer a substrate-enzyme binding affinity $K_d$ from <i> in vitro </i> measurements of the reaction velocity (i.e., the rate at which substrates are converted to products in a controlled reaction), we generally fit the reaction velocity data with a Michaelis-Menten equation for the reaction kinetics, and identify $K_d$ with the Michaelis-Menten constant. Doing this, we are <b> fitting the data with a Michaelis-Menten model for enzymatic kinetics </b>, and 1) there are situations where the Michaelis-Menten constant deviates from the $K_d$ and 2) there are situations where the Michaelis-Menten approximation is not even valid, and the entire model falls down.
Hence, in short, <b> any experimental measurement of a parameter is in fact a fitting of a direct measurement to a model of how the biological parameter affects this measurement </b>. So when using data, including what looks like a direct measurement, to infer a model parameter, the very first question to ask is: <b> can any of the other model parameters or variables influence the result of my experiment? </b> If the answer is no, then it means that all the things (e.g. pH, temperature… ) that influence the measurement are not included in the model. Hence, in this scenario the measured parameter can be used directly to feed the model, with the restriction that the model prediction will be “valid” in the same conditions (pH, temperature… ). If the answer is yes, then the next one is <b> how much do other model parameters influence the measurement? </b> Answering this question in order to design strategies to improve parameter measurements is the focus of this section.
In the context of mathematical modeling of biological systems, the analysis of the influence of parameters on experimentally measurable (directly or not) model outputs relates to two distinct categories of situations, which will be discussed separately: the case where <b> we have experimental data from which we want to obtain model parameters - Parameter Inference - </b> and the situation where we want to <b> modify parameters and predict the changes in measurable biological quantities - Model Predictions </b>. Both are in fact not mutually exclusive, since accurately measuring non-controllable model parameters may be an important pre-requisite to accurately predict the system’s behavior upon changes of experimentally controllable parameters.
## Parameter inference
As illustrated in the examples above, the purpose of parameter inference is to use data to measure (or at least, get quantitative information on) a parameter <b> within the framework of a given model </b>. Biological data is inherently stochastic, but follows deterministic trends, hence any measurement is a mix of a deterministic part and a stochastic part. Therefore, biological data can be interpreted in the frameworks of both deterministic and stochastic models. But in both cases, because the data itself is stochastic, the parameter values that we infer from the data are <b> estimations </b> of the underlying parameter values (one would need an infinite number of an infinite number of different measurements to actually <b> measure </b> the parameter). So the adequate language in parameter inference problems is the language of statistics, and the parameter values we obtain are <b> statistical estimators </b> of the underlying parameters. This explains <b> why there are so many different ways of inferring parameters from limited datasets </b>, because there always exist many different statistical estimators that all have their pros and cons and range of “optimal validity”.
The take-home message is therefore that, together with any estimation of a parameter, we shall always provide the uncertainty with which this parameter is determined
### 1) Principles of data fitting (deterministic models)
In a deterministic model, the output variables (for instance, time-course of a protein phosphorylation) are functions of the input variables (e.g. concentrations of kinases and phosphatases) and the model parameters (e.g., rates). Formally, this can be written as:
$$outputs=F(inputs,\alpha_1,\alpha_2 … \alpha_N)$$
where we have assumed our model has $N$ parameters $\alpha_1,\alpha_2 … \alpha_N$. If we have been able to solve the model analytically, we are able to <b> write an explicit mathematical formula for the function F as a function of its parameters </b>, and thus we can plot curves of $outputs$ as a function of any $input$ or $parameter$. If we haven’t found such an analytical solution, we can “integrate” our model (generally, ODEs/PDEs or algebraic equations) numerically and instead of finding a closed form mathematical formula, we get the values $output_i=F_i$ for a discrete number of input values $input_i$, for any possible values of the model parameters. This is the case, for instance, when we integrate an ODE/PDE model: for any set of parameters that we freely choose, we obtain the solution $F$ at discrete input times (and for PDEs, discrete input positions in space). And we can almost always choose which discrete times and positions, so in short we are able to plot the function $outputs=F(inputs, \{\alpha_i\})$, for any set of parameter values that we want.
Parametric fitting is the process by which we try to find the parameter values $\alpha_1^0,\alpha_2^0 … \alpha_N^0$ that minimize the deviation (or error) between a set of outputs $ O^i_{meas}$ measured for a given set of input values, and a set of output values $ O^i_{pred}$ predicted by the model for the <b> same inputs values </b>. There are many different fitting procedures and the purpose here is not to list all of them, however they generally differ on <b> how the “error” is defined </b> and/or what <b> minimize </b> means in the context of a particular procedure.
A very popular method is least-squares fitting, where the error function is defined as:
$$E(\alpha_1,\alpha_2 … \alpha_N) = \sum_{i=1}^K \displaystyle \Bigg( O^i_{meas}- O^i_{pred}(\alpha_1,\alpha_2 … \alpha_N) \Bigg)^2 $$
$K$ is the total number of measurements of the output $O$ and the minimization is understood as finding the smallest value of the function $E$ of several variables $\alpha_1,\alpha_2 … \alpha_N$, i.e. finding the values $\alpha_1^0,\alpha_2^0 … \alpha_N^0$ for which $E$ is minimal.
One immediate problem we can see with such an approach is that with such a definition of the error function, each measurement is equally weighted in the total error and thus contributes to the same extent to the estimation of the parameters minimizing $E$. In practice, some measurements might be more reliable than other, or we might even want to <b> choose </b> that some measurements are more important. An alternative is then to define an error function with different <b> weights </b> for all residuals $ O^i_{meas}- O^i_{pred}(\alpha_1,\alpha_2 … \alpha_N)$:
$$E(\alpha_1,\alpha_2 … \alpha_N) = \sum_{i=1}^K w_i*\displaystyle \Bigg( O^i_{meas}- O^i_{pred}(\alpha_1,\alpha_2 … \alpha_N) \Bigg)^2 $$
where the choice of the weights $w_i$ is completely free. A popular choice is to use $w_i=1/\sigma_i^2$ where $\sigma_i^2$ is the variance of the i-th measurement. This choice reduces the weight of the less-reliable measurements in the estimation of the parameters, and happens to be the best choice when the different measurements are independent and uncorrelated. However, we can imagine to chose weight coefficients that scale with the typical scale of the measurement $O^i$ if it happens that the $O^i$ are measurements of different biological quantities… this provides a way to compare apples and oranges.
When the model is stochastic, in principle any set of observations $O_i$ can be explained by any set of parameters, just because any set of parameters can generate any output (remember: outputs are random variables). Any set of observations $O_i$ can be explained by any set of parameters? Yes. But not with the same probability. When fitting a dataset with a stochastic model, we therefore define the likelihood function, which is the probability to observe what we have observed knowing the underlying model and its parameters $\alpha_1,\alpha_2 … \alpha_N$ :
$$ L(\alpha_1,\alpha_2 … \alpha_N) = Proba(observing \, \{O^i_{meas} \}|\alpha_1,\alpha_2 … \alpha_N)$$
This probability is then computed in the framework of the stochastic model, for instance by simulating the model a large number of times for each choice of parameters and computing the number of time $O_i$ is obtained, relative to the total number of simulations, and then reiterating for a new choice of parameters. The optimal parameters $\alpha_1^0,\alpha_2^0 … \alpha_N^0$ are then obtained upon <b> maximization </b> of the likelihood function with respect of the parameters.
We stress here that regardless of how the error function $E$ is defined, “how much minimization” we can achieve (how small it can get) depends on the number of model parameters, and generally the more parameters we have, the smaller $E$ can be. In fact, any function $ O^i_{pred}(\alpha_1,\alpha_2 … \alpha_N)$ that goes exactly through the observed points $O^i_{meas}$ at the sampled input values will lead to a least square error of exactly 0, the smallest we can get. And mathematics tell us that there is always a polynomial function that go through exactly all points, for any discrete number of points. Hence, there always exist a model curve that fits exactly all data points. This of course seems ridiculous, but it is a true and universal statement: <b> more complex models with more parameters are susceptible of producing better fits </b>. This effect is called <b> over-fitting </b>, and is nicely illustrated by this schematic picture from Wikipedia (no further explanation is needed, isn’t it?):

This is also true for stochastic models and using of the likelihood function for parameter fitting. Too many parameters in a model, especially if they provide an excellent fit of a dataset, generally also restrict a lot the ability of the model to accurately <b> predict </b> other observations, and there is always a trade-off between how well we can fit data and how predictive a model is.
This statement is illustrated by the famous quote from the French poet Paul Valéry: <b> “Ce qui est simple est toujours faux. Ce qui ne l'est pas est inutilisable. </b> which could be translated as <b> “What is simple is always wrong. What is not is useless. ” </b> Simple models with few parameters have the potential to guide scientific thinking on a broad range of questions, but are generally poor as accurately describing data. This duality in model choice is discussed below (model selection).
But before going further, it is important to understand what a <b> good/bad fit </b> is, and how reliable the parameter estimations we get from the fit are.
### 2) Goodness of fit, sensitivity analysis, parameter identifiability and reliability of the parameter estimation
How good a fit is quantified by statistical tests, the choice of which depends on the type of data, the information we have on their underlying probability distribution… and are covered in the data analysis and statistics courses. The Goodness of Fit is generally a number ($\chi^2$, $R^2$ …) which indicates how well the given model, with its “optimized” parameters (see 1), describes the data. Thus, we might be inclined to <b> blindly trust </b> parameter estimations when this goodness-of-fit number is good (e.g., $\chi^2 \approx 1$). However not all these estimations are equally reliable, and to understand it we have to take a step back and thing again about our model.
Indeed, not all model parameters equally affect the output that we have observed, $O$. And parameters that have little effect on $O$ are potentially badly determined, even if the fit is “good”. The analysis of the impact of parameters on model outputs is called <b> sensitivity analysis </b>.
If we solved the model analytically, we have an explicit formula for the solution $F$ of our model
$outputs=F(inputs,\alpha_1,\alpha_2 … \alpha_N)$
and we can perform a variational analysis and formally write the difference in model output following small changes in parameters (but no changes in input) as :
$$d(outputs)=dF=\frac{\partial F}{\partial inputs} d(inputs) +\frac{\partial F}{\partial \alpha_1} d\alpha_1 +\frac{\partial F}{\partial \alpha_2} d\alpha_2+ … +\frac{\partial F}{\partial \alpha_N} d\alpha_N $$
This is nothing else than the expression of the total differential of the function $F$ of several “variables”, the model parameters. Because we study how $F$ changes with respect with parameters at constant input, we set $ d(inputs)=0$ and we get:
$$d(outputs)=\frac{\partial F}{\partial \alpha_1} d\alpha_1 +\frac{\partial F}{\partial \alpha_2} d\alpha_2+ … +\frac{\partial F}{\partial \alpha_N} d\alpha_N $$
What does that mean? It means that if you slightly modify the value of one parameter $\alpha_i$ from its “optimal” value obtained with the fit $\alpha_i^0$, for instance $\alpha_i=\alpha_i^0+d\alpha$ where $d\alpha$ is small compared to $\alpha_i^0$ you can expect a change in the output that is:
$$d(outputs)=d\alpha * \frac{\partial F}{\partial \alpha_i} (\alpha_1^0, …\alpha_i^0, …\alpha_N^0) $$
where the partial derivative is computed for the optimal parameters that yielded the best fit, $\alpha_1^0,\alpha_2^0 …\alpha_i^0, …\alpha_N^0$.
But conversely, it also means that <b> if you know the output with a precision </b> $d(output)$, for instance due to an uncertainty in the experimental measurement of the output, then the parameter $\alpha_i$ will be determined with an accuracy $d\alpha$ that depends on those partial derivatives. When dealing with uncertainties, we always deal with positive $d(something)$ but in the total differential of the outputs, the partial derivatives with respect to some parameters can be negative. Hence, in terms of positive-only quantities we can rewrite the total differential as an inequality:
$$d(outputs)\leq|\frac{\partial F}{\partial \alpha_1} |d\alpha_1 +|\frac{\partial F}{\partial \alpha_2}| d\alpha_2+ … +|\frac{\partial F}{\partial \alpha_N}|d\alpha_N $$
where $||$ denotes the absolute value. This is nothing else than the formula for the propagation of uncertainties. In a context of parameter estimation, if the measured variable is known with an uncertainty $d(output)$, the parameter $\alpha_i$ would be known with an accuracy
$$d\alpha \geq \frac{ d(outputs)}{ |\frac{\partial F}{\partial \alpha_i} |(\alpha_1^0, …\alpha_i^0, …\alpha_N^0)},$$
which means that </b> the lower </b> $ \frac{\partial F}{\partial \alpha_i} (\alpha_1^0, …\alpha_i^0, …\alpha_N^0)$ <b>, the less accurate the determination of the parameter </b> $\alpha_i$. But if this is true for one parameter, i.e. if the optimal fit gave a ridiculously wrong value $\alpha_i^0$ for the i-th parameter, how reliable is the value of the partial derivative $ \frac{\partial F}{\partial \alpha_j} (\alpha_1^0, …\alpha_i^0, …\alpha_N^0)$ with respect to all other parameters $\alpha_j$? If it is not reliable, how reliable is the estimation of those parameters itself?
This example teaches us that the estimation of the uncertainty of parameters fitted from experimental values is a complex problem.
The approach we have used here is called a “one-at-a-time” sensitivity analysis method. If the model is linear or weakly non-linear, if parameters are statistically independent (non-correlated), we can assume as a first approximation that the variance in the measured quantity (or its “uncertainty”) is equally distributed between all model parameters, and in this case the uncertainty of each parameter can be estimated as
$$d\alpha \approx |\frac{ d(outputs)}{ \frac{\partial F}{\partial \alpha_i} |(\alpha_1^0, …\alpha_i^0, …\alpha_N^0)},$$
in order of magnitude. We stress that if an explicit analytical formula for the model outputs as a function of the parameters is not available (e.g., model has been solved numerically), we cannot compute analytically the partial derivatives, however we can compute them numerically as “ratios of small differences”, obtained by comparing the values of the outputs in two numerical resolutions of the model for slightly different values of the parameter $\alpha_i$:
$$\frac{ d(outputs)}{ \frac{\partial F}{\partial \alpha_i} (\alpha_1^0, …\alpha_i^0, …\alpha_N^0)} = \frac{outputs(\alpha_1^0, …\alpha_i^0+\delta \alpha, …\alpha_N^0)-outputs(\alpha_1^0, …\alpha_i^0, …\alpha_N^0)}{\delta \alpha}$$
where $\delta \alpha$ is small compared to $\alpha_i^0$. With this way of computing the partial derivative, all the discussion above applies equally to models solved numerically.
This simplistic approach of one-at-a-time sensitivity analysis might provide information on the accuracy of model parameters estimation, with the following restrictions/limitations/caution when using:
- combined effects of parameters often happen, in which case analyzing the sensitivity with respect to individual parameters leads to erroneous interpretations.
- similar effects on the outputs from different parameters: this is particularly frequent if the model has a large number of parameters; this situation will be discussed later more extensively.
- analysis of individual outputs separately when multiple outputs are available: it is still possible to analyze both outputs simultaneously, but if they are somehow correlated through some - but not all - of the parameters, the sensitivity with respect to those will be artefactually emphasized.
We have introduced it here for pedagogical purpose, because it illustrates the general rule of thumb: the lower $|\frac{\partial F}{\partial \alpha_i} |$ for the optimal, fitted parameter values (the <b> sensitivity </b> of the outputs to the parameter $\alpha_i$), the less accurate the estimation of the parameter $\alpha_i$. We will use this rule of thumb to discuss more complex notions.
In practice, because models include multiple parameters, <b> global sensitivity analysis </b> approaches are more reliable. <b> Global sensitivity analysis </b> refers to the analysis of output variations when <b> all parameters can be changed at the same time </b>. Those techniques are powerful because they measure sensitivity across the entire parameter space, and therefore perform well even in presence of strong non-linear effects, non-independence of parameters (i.e. change in one parameter can always be compensated by a change in another or a few others), interactions in parameters … They can be applied to deterministic models with or without an analytical solution, but also to stochastic models. The downside of such global approaches is that they usually have a high computational cost.
One of such methods is the <b> Sobol variance-based sensitivity analysis method </b>. The principle of the method is to generate model outputs with a large number of randomly picked parameter values, and to analyze the resulting <b> variance </b> in the output. Then, a statistical decomposition method is used to separate the contributions to the output variance in terms due to the variance of all parameters independently of each other, + terms due to the cross variance of all pairs of parameters, + terms due to the cross variance of triplets of parameters… and so on.
Such global approaches that screen a large number of parameter values are also very useful to infer parameters of a stochastic model from data. Indeed, in a stochastic model any variation in a parameter results in a new stochastic trajectory of the system. But not changing anything also results in a new trajectory. Hence, there is an output variance associated with the stochasticity of the model even without changing any parameter. The actual spread of this variance is also in general <b> dependent on the parameters </b>. Hence, the variance in model outputs that are compared with data contain information on the model parameters as well. So, biological noise perturbs parameter analysis, but also offers an <b> additional tool for parameter inference .</b>
### 3) Parameter identifiability and reliability of the parameter estimation
Another limitation of sensitivity analysis stems from the fact that even though biomedical data available to compare with model output tends to become richer and richer, it cannot be </b> unlimited </b>. Hence, the situation can happen where a given parameter has <b> no influence at all on any of the outputs we have measured </b>, but yet it affects either other model outputs, of the model behavior in different dynamical regimes and then getting information on this parameter is important to make accurate model predictions. For instance, assume we model a biochemical system that accounts for some fast processes on the millisecond scale, and we want to infer parameters from data acquired in response to a stimulus with an experimental time resolution of 1s (due e.g. to a technical limitation). By the time we acquire the first time point following stimulation, all fast processes have reached a steady state, regardless of the exact values of the parameters describing the fast processes. Hence, our experiment won’t be sensitive to those parameters. Yet, they influence the dynamics at the millisecond timescale, and our model will be unable to provide any insight on another experiment that has a faster time resolution.
Another common situation is the case where several parameter changes, or changes in pairs, triplets… of parameters have exactly the same effect on our model outputs. In this case, the model output is still sensitive to the parameters, but is not able to <b> discriminate between them </b>. In both cases, we say that the parameters are <b> not identifiable </b>.
And in both cases, regardless of how sophisticated our sensitivity analysis method is, <b> we are facing a wall </b> because the measurements we have performed are inherently <b> incapable </b> of distinguishing the influence of two (or more!) parameters.
The <b> identifiability </b> of a model parameter is a crucial notion one often encounters in the context of parameter inference. It is linked to the question: Can <i> any </i> available data provide <i> any </i> insight on the value of the parameter? If the answer is “no”, then the parameter is said <b> non-identifiable </b>. In this situation, we should try to formulate the model without the model interaction related to this parameter. If this is impossible, we should try to <b> design an experiment that could turn this no into a yes </b>. If this is not possible, for instance due to current technical limitations, then the dynamical behavior of the model shall be studied <b> for any possible value of this parameter </b> and conclusions shall be robust to changes in this parameter.
There are generally speaking two categories of situations in which we answer “no” to the question: can <i> any </i> measurement provide <i> any </i> insight on the value of the parameter? The first category is when variations in this parameter (which we denote $\alpha$ do not affect any measurable output $Y$ of the system: for any $Y$,
$$\frac{\partial Y}{\partial \alpha }=0.$$
In other words, the output variables that we have measured are “insensitive” to the parameter $\alpha$, in the sense of parameter sensitivity analysis. The second category is when the effects of variations in this parameter cannot be distinguished from variations caused by another parameter $\beta$ on all measurable outputs $Y$ of the model :
$$\frac{\partial Y}{\partial \alpha }=\frac{\partial Y}{\partial \beta }.$$
In this case, any measurement we perform can be equally attributed to $\alpha$ or $\beta$ and we can’t get insight on those parameters: $\alpha$ and $\beta$ are <b> not independent </b>, or rather we should say that $Y$ does not depend independently on $\alpha$ and $\beta$.
Let’s look at an example to understand this notion. We assume a simple gene transcription model where a pre-formed RNA Pol II complex binds a gene promoter with rate $k_{on}$ (unbinds with rate $k_{off}$, forming a pre-initiation complex, which eventually enters an active mRNA elongation phase with rate $k_{elong}$ and completes this phase with rate $k_{term}$, releasing the RNA Pol II complex in the nucleoplasm and releasing one full-length mRNA molecule. In addition, we assume that the elongating polymerase complex can enter a “paused” state (with rate $k_{p}$) where it stops elongating the mRNA, and leaves this paused state with a rate $k_{q}$. This model can be represented with the following graph:

where $P_c$ is the number of nucleoplasmic complexes, $P_{pic}$ the number of pre-initiation complexes, $P_{e}$ the number of actively elongating polymerases and $P_{p}$ the number of paused ones.
The ODEs representing the variations of the number of polymerase complex at each stage of the process are (good exercise to re-derive them):
$$\frac{dP_c}{dt}=k_{off}*P_{pic}+k_{term}*P_e-k_{on}*P_c$$
$$\frac{dP_{pic}}{dt}=-k_{off}*P_{pic}-k_{elong}*P_{pic}+k_{on}*P_c$$
$$\frac{dP_e}{dt}=k_{elong}*P_{pic}-k_{term}*P_e-k_p*P_e+k_q*P_p$$
$$\frac{dP_p}{dt}=k_p*P_e-k_q*P_p$$
from which we get (by summing all 4 equations)
$$\frac{d(P_{pic}+P_c+P_e+P_p)}{dt}=0$$
which expresses the fact that the total pool of polymerase complexes $P_{tot}= P_{pic}+P_c+P_e+P_p$ is conserved over time by this process.
In the steady state of this ODE model, $d/dt=0$ and all the left hand sides of the equations above are =0. Hence, in Eq. 2 we can get $P_{pic}$ as a function of $P_c$ and parameters, and in Eq. 3 $P_e$ as a function of $P_{pic}$ and parameters, and in Eq. 4 $P_p$ as a function of $P_e$ and parameters:
$$P_{pic} = P_c*\frac{k_{on}}{ k_{off}+ k_{elong}}$$
$$P_e=P_{pic}*\frac{ k_{elong}}{ k_{term}}= P_c*\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}$$
$$P_p=P_e*\frac{k_p}{k_q}= P_c*\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}$$
and from the conservation of the total number of polymerase complexes, we get:
$$P_c=\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}$$
and using this expression for $P_c$ we can compute the steady state number of polymerases in all other stages of the process:
$$P_{pic} = \frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}$$
$$P_e=\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}$$
$$P_p=\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}*\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}.$$
And the rate of mRNA production in the steady state is therefore:
$$\frac{dmRNA}{dt}=k_{term}*P_e=k_{term}*\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}$$
<b> Question 1: </b> We assume we can tag a protein component of the RNA Pol II complex with a fluorophore, and monitor the variations of the fluorescence signal over time. Can we infer some of the parameters of this model with this data?
```
#%run solutionQuestion1.py
```
<b> Question 2: </b> We now measure the rate of production of the mRNA molecules. After a transient regime, this rate becomes stable in time and we measure it with good accuracy. In addition, we have a probe that gives a signal from which we can compute the number of highly mobile pool of polymerases (the nucleoplasmic pool - this can be achieved by modern microscopy techniques). Can we infer some of the parameters of this model with this data? Is there a particular parameter regime in which this experiment might be even more informative?
```
#%run solutionQuestion2.py
```
<b> Question 3: </b> What ratio of measurable quantities should we measure to get information on the pausing dynamics k_p, k_q? What should we do to get both k_p and k_q? Is that always possible experimentally?
```
#%run solutionQuestion3.py
```
In complex biomedical research models, we have access to a limited amount of data (though it can be tremendous - still not “infinite”), and it is possible that not all parameters are identifiable with the available datasets. More model parameters generally reduce the identifiability of each of them, unless datasets are enriched as well. In the previous example of the gene transcription model, the experiments suggested in the question are insufficient to identify all parameters, and the model should be simplified, or extensively analyzed before conclusions are drawn (see below, parameter space analysis). The extent to which we need more data to preserve parameter identifiability in a more complex model depends on the model, there is no general rule. However, there are techniques to compare the performance of models of different complexity in fitting datasets (termed model selection methods, see below).
In practice, we also don’t have often a simple solution of the model like in our previous example. Hence, the identification of which parameters are identifiable and which are not is not immediate, and requires a proper methodology. This methodology is termed <b> a (structural) identifiability test </b>. The development of identifiability tests is still an active topic of research.
<b> Further recommended reading: </b> an intuitive way to test parameter identification in systems biology-like models has been derived by Clemens Kreutz (https://doi.org/10.1093/bioinformatics/bty035). The method is based on the addition to the square error $E(\alpha_1,\alpha_2 … \alpha_N) = \sum_{i=1}^K w_i*\displaystyle \Bigg( O^i_{meas}- O^i_{pred}(\alpha_1,\alpha_2 … \alpha_N) \Bigg)^2 $ of a penalty term :
$$P(\alpha_1,\alpha_2 … \alpha_N)=\lambda * \displaystyle \Bigg( \sqrt{(\alpha_1-\alpha_2^0)^2+(\alpha_1-\alpha_2^0)^2+…(\alpha_N-\alpha_N^0)^2 } - R \Bigg)^2 $$
where $R$ is a parameter, $\alpha_i^0$ are the values of the parameters that minimize the error function (in other words, the fitted parameters), and $\lambda$ is the penalty coefficient and is chosen as $1/R^2 $ in the basic version of the method. This term quantifies <b> how much it costs to drag the parameters “away from their optimal value” </b> (specifically, at a “distance” $R$ on average).
And the principle is intuitive: if the parameters are identifiable, there is no other parameter set that can minimize the error function in a satisfactory way, and therefore we can’t cancel the “penalty” part without increasing the “data” part ($E$). If the parameters are non-identifiable, it is possible. Hence, we know if parameters are collectively identifiable by trying to minimize both the error $E$ and the penalty $P$. If we succeed, they cannot be identified in a unique way.
We stress that there is no general consensus on the best tests for parameter identifiability in a general model. But when inferring parameters from data, it is advised to try one identifiability approach to make sure that a given dataset is sufficient to provide model parameter values in a <b> unique </b> way. And if it is not, using sensitivity analysis allows to guess which parameters are reasonably well identified and which are not.
### 4) model validation, model comparison: the trade-off between parameters and information
An important aspect of mathematical modeling related to parameter estimation is the possibility to describe a given biological system (and the associated data) with several different models. We’re not talking here about the possibility to choose a mathematical formulation, e.g. ODE vs PDE, steady state or fully time-dependent, stochastic vs deterministic… but really about <b> the choice of the objects and features to include in the model </b>.
Let’s look again at the gene transcription model we studied in the previous section, and assume we have access to Global Run On Sequencing data (GRO-seq) (that provides an estimation of the ratio of actively elongating polymerase complexes to “paused” complexes, $P_e/P_p$), access to mobility data that yields the number of nucleoplasmic complexes $P_c$ and access to RNAseq data that allows to count the mRNA rate transcription ($k_{term}*P_e$). As discussed in the previous section, several parameters of this model remain non-identifiable with these two datasets in hand. The question is: is it worth describing this data with such a complicated model?
The following model:

and where the steady state is
$$P_p=\frac{k_p}{k_q}*\frac{k_{on}}{k_{term}}*\frac{P_{tot}}{1+\frac{k_{on}}{k_{term}}+\frac{k_{on}}{k_{term}}*\frac{k_p}{k_q}}$$
$$P_e=\frac{k_{on}}{k_{term}}*\frac{P_{tot}}{1+\frac{k_{on}}{k_{term}}+\frac{k_{on}}{k_{term}}*\frac{k_p}{k_q}}$$
$$P_c=\frac{P_{tot}}{1+\frac{k_{on}}{k_{term}}+\frac{k_{on}}{k_{term}}*\frac{k_p}{k_q}}$$
is also a possible description of the full biological process. A noticeable difference with the previous model is that it does not account explicitly for the pre-initiation complex stage, and therefore that the rate $k_{on}$ integrates the binding/unbinding and initiation of the elongation (i.e., the rates denoted as $k_{on}$, $k_{off}$ and $k_{elong}$ in the previous model). Thus, there are less parameters (4 instead of 6).
In the framework of this new model, the mRNA production rate in the steady state is:
$$\frac{dmRNA}{dt}=k_{term}*P_e=k_{on}*\frac{P_{tot}}{1+\frac{k_{on}}{k_{term}}+\frac{k_{on}}{k_{term}}*\frac{k_p}{k_q}}= k_{on}*P_c$$
In this model, there are still independent parameters that characterize the GRO-seq data ($k_p/k_q=P_p/P_e$), and the mRNA-seq data normalized to the mobility data ($(dmRNA/dt)/P_c$, characterized with the parameter $k_{on}$), see formulas above). Hence, there is <b> enough flexibility </b> in the model to account for all datasets available, and the fit would be as good as a fit with the more complex model we discussed first.
Hence, which model should we choose?
The question we are asking here is the question of <b> model selection </b>. In this simple example, the two models are qualitatively very close, and the processes of polymerase complex binding to promoter, spontaneous unbinding, elongation initiation and abortion… are rather well understood. So there is no new biology, new putative hypotheses to test included in this “initial step” of the two models. But in the general situation, different models that explain equally well the data might rely on different biological hypotheses and in this case which model we choose means <b> how do we interpret the data biologically </b> and this is a crucial point, if not the <b> most important point when doing an experiment. </b>.
This is Ockham’s principle (or “razor”): of competing hypotheses (different models) that make the same prediction (explain equally well the available data), one should select the solution with the fewest assumptions (and/or parameters). Simpler is best if, of course, it fits well the data!
Intuitively, the ideal model should 1) describe reasonably well the data available, 2) contain all the objects on which we intend to make predictions, and 3) contain as few additional things as possible, since any additional ingredient makes the model more complex and won’t help to describe the data or to make predictions since those are covered by 1) and 2). Because additional/unnecessary model objects and/or model interactions come with additional model parameters, the <b> number of parameters is an intuitive measure of the complexity of a model </b>. So, the trade-off between the ability of a model to explain (fit) the observations and its complexity should be well represented by the difference between the goodness of fit and the number of parameters.
This intuitive idea of balancing model complexity with the ability to describe/fit/reproduce existing data is the basis of model selection procedures. But a model should also have a <b> predictive power </b>, i.e. be able to describe/fit/reproduce data that is yet not acquired. Indeed, selecting among competing hypotheses to understand the biological interactions and processes underlying the generation of the data we are looking at is not the only purpose of a model: another equally important purpose is to predict future observations. And this second aspect is much more complicated to quantify reasonably, for obvious reasons. This is why, again, there is no general consensus on which model selection procedure is the best. Below, we’ll describe some commonly used, with their pros and cons.
<b> Akaike Information Criterion (AIC): </b>
$$AIC_M=2*K_M-2*\ln{L_M}$$
where $K_M$ is the number of parameters in the model $M$, and $L_M$ is the maximum of the likelihood function (i.e., the value of the likelihood that the data is explained by the model $M$ when the model parameters are the best possible, those which maximize this likelihood). $L_M$ measures then how good the data fit is with the model $M$ once model parameters have been optimized. If we have in hand competing models $M_1,M_2…$ to analyze the data, we compute the AIC number for all models, and get a list of a handful of AIC numbers. The “best” model according to AIC is the one that <b> minimizes the AIC </b>. Let’s denote $M_0$ this model, and by definition we have
$$AIC_{M_0}<AIC_{M_1}, AIC_{M_1}, …$$
In practice, it is frequent that a couple of models give values of the AIC which are very close, and it might be informative to quantify <b> how likely the other models are, relative to the “best” model </b>. In the Akaike theory, this relative likelihood is (for the model $M_j$):
$$r=e^{ \frac{AIC_{M_0}-AIC_{M_j}}{2}}<1$$
Let’s compare the two gene transcription models we used previously. Let’s call $M_1$ the first, complete model with 6 parameters, and $M_2$ the shortened model with only 4 parameters. As discussed above, because both models have enough free independent parameters to describe our datasets, in practice we would find $ L_{M_1}= L_{M_2}$ (the models would perform equally well in fitting the data) and therefore $\ln{ L_{M_1}}=\ln{ L_{M_2}}$. Let’s denote $\mu$ this common value. Hence,
$$ AIC_{M_1} = 2*6-2*\ln{ L_{M_1}}=12-2*\mu$$
$$ AIC_{M_2} = 2*4-2*\ln{ L_{M_1}}=8-2*\mu < AIC_{M_1},$$
therefore we would pick the simplest model $M_2$ as the “best” model. The relative likelihood of the other model would be:
$$r=e^{ \frac{(8-2*\mu)-( 12-2*\mu)}{2}}= e^{-2}=0.135:$$
the model 2 is 0.135 times as likely as the model 1 to be the right model underlying the data.
Given its definition, the Akaike Information criterion does not account for the predictive power of the model, and is well adapted to situation where the data is fit to the models in terms of “probability to observe what we observed given the underlying model is … ” (i.e., when data is fitted using maximum likelihood estimators, and not squared error minimization). Hence, it is strictly speaking a criterion to compare among competing <b> statistical models </b> (termed “stochastic” models in this course), rather than deterministic mathematical models (e.g. ODE/PDE-based). This criterion appears simple and intuitive, yet it has solid mathematical foundations, and is strictly valid only in the asymptotic limit where we have an infinite number of observations. When we have to base model selection on a small number of observations, the AIC needs to be corrected, and the mathematical expression of the corrected criterion AICc depends on statistical properties of the model and of the data.
<b> Bayes Information Criterion (BIC): </b>
$$BIC_M=K_M*\ln{Q}-2*\ln{L_M}$$
where $K_M$ is the number of parameters in the model $M$, and $L_M$ is the maximum of the likelihood function (as in the AIC), and $Q$ is the number of observations available. The “best” model $M_0$ is the one that minimizes the BIC. Hence, when compared to the AIC, when using the BIC <b> the penalty associated with excess parameters increases with the number of observations </b>. In other words, the more observations we are accounting for when fitting for model parameters, the larger the fitting error we can accept from simple models with less parameters. Using the BIC, the relative likelihood of another model $M_j$ is:
$$r=e^{ \frac{BIC_{M_0}-BIC_{M_j}}{2}}<1$$
In our example where we compared the two gene transcription models, we were using only two measurements to fit the parameters: the ratio $P_p/P_e$ provided by GRO-seq data, and the ratio $(dmRNA/dt)/P_c$ provided by RNA-seq and the mobility data. Hence, $Q=2$. Without surprise, the simplest model minimizes the BIC but the relative likelihood of the more complex one becomes
$$r= e^{-\ln{2}}=0.5,$$
i.e. because we didn’t fit that many observations with the simple model, it’s “advantage” over the more complex model is less obvious than in the AIC. Model selection using AIC or BIC do not always agree, because those apparently similar formulas stem from completely different mathematical concepts.
Similar to the AIC, the BIC does not account for the predictive power of the model, is well adapted to the comparison of statistical/stochastic models, and becomes more and more acceptable when the number of observations $Q$ increases.
We stress that both the AIC and BIC, which are in principle defined for stochastic models, can be somehow extended for deterministic models, replacing the maximum of the likelihood function $L_M$ by its equivalent in deterministic model parameter fitting, the error function $E$. In this case, the error function (called the residual variance) is the minimal one, computed at the values of the parameters that minimize it $E_0 =E(\alpha_1^0, \alpha_2^0 …\alpha_N^0 )$, leading to a criterion C that the “best” model will minimize:
$$C=\lambda*K_M+\ln{ E(\alpha_1^0, \alpha_2^0 …\alpha_N^0 )} .$$
The “+” sign comes from the fact the in this context we need to minimize the error, not maximize the likelihood. There is no consensus on what the constant $\lambda$ should be, because it depends on the relative weights we intend to give to the “exactness” of fitting and the number of parameters (aren’t BIC and AIC different in this respect?). Research works have derived different forms for $\lambda$ in different purpose, and depending on the statistical properties of the residuals that compose the total error. If the latter are normally distributed, then $\lambda=1/Q$ where $Q$ is the number of observations is somehow equivalent to the Akaike criterion. Hence, $\lambda = \ln{Q}/Q$ would penalize complex models the same way as the BIC does. Another popular choice is to choose $\lambda=\sigma^2/Q$ where $\sigma^2$ is the data variance, or uncertainty. In this case, simple models are even more favored than complex ones if they describe reasonably well data that is very noisy. But, again, it depends on the purpose and those techniques for model selection should be approached with both flexibility and caution.
<b> Minimum Message Length - Minimum Description Length criterion (MML/MDL): </b>
One common limitation of both the AIC and BIC is that they represent model complexity by its number of parameters. The Ockham’s razor principle is slightly more general, in that it states we should be looking for the <b> simplest explanation of the data </b>. Simpler does not necessarily mean less parameters (and conversely), even though <i> in general </i> it is the case.
In addition to AIC/BIC and similar criterion, computer scientists often use the concepts of <b> message length and description length </b> to estimate how well a model explains a given dataset. The message length ML is the length (in bits) of a code that explains both the content of the model and describes the data (obviously, the shortest explanation/description possible). The description length DL, even if it looks similar, is a fundamentally different concept: it is the length of a model-based description that <b> encodes </b> the data (for instance, “logistic growth with rate 1 and saturation 4” is a description that encodes the entire logistic curve $P(t)$ with parameters $r=1$ and $P_{max}=4$, and therefore encodes any sampling of datapoints long this curve). In both cases, the selected model is the one that minimizes the ML or the DL, hence the MML/MDL nomenclature to classify those methods. Because there are several metrics for ML and DL depending on the contexts, there are several MML/MDL methods.
<b> Selecting for predictive power: </b>
One common limitation of both the AIC/BIC and MML/MDL approaches is that the model selection is based on <b> existing </b> datasets, and therefore does not explicitly accounts for the predictive power of a model, i.e. the ability of a model to <b> accurately describe experiments to come </b>. The universe of the possible experiments “to come” is unbounded, therefore there is no - and there will never be - a generic statistical method that is able to predict how well a model will perform in predicting a certain type of experiment that does not yet exist. In the next section, we describe general guidelines to design model validation experiments.
However, there are techniques that allows to estimate how good a model is in predicting some data of the same type as the data on which the model has been optimized. Such techniques are called <b> cross-validation (CV) </b> techniques and are based on the split of the total dataset (generally) in two groups, the <b> training </b> dataset and the <b> validation </b> dataset. The model parameters are estimated from the minimization of the error function on the training dataset (observations $O_i$ with $i=1..i_0$ for each model independently (or the maximization of the likelihood function for stochastic models):
$$E(\alpha_1,\alpha_2 … \alpha_N) = \sum_{i=1}^{i_0} w_i*\displaystyle \Bigg( O^i_{meas}- O^i_{pred}(\alpha_1,\alpha_2 … \alpha_N) \Bigg)^2 $$
and once the parameters are chosen, we compute the total error (a number) on the rest of the dataset assuming that the model parameters are the ones we just picked:
$$E_{sel} = \sum_{i=i_0+1}^{K} w_i*\displaystyle \Bigg( O^i_{meas}- O^i_{pred}(\alpha_1^0,\alpha_2^0 … \alpha_N^0) \Bigg)^2 $$
It somehow means that we are looking at that this second part of the dataset as a <b> future experiment </b>, and we are measuring the total error that the models yielded on this experiment. Hence, in the purpose of selecting models for their predictive power, we would choose the model that minimizes this residual error on the “future” experiment, $E_{sel}$. We could, then, add a BIC/AIC-like penalty term, to balance the choice…. We note that once the model has been selected, it is recommended to re-estimate the parameters on the entire dataset.
In practice, of course the model choice could be influenced by how exactly we split the data. It is therefore recommended to partition the data randomly several times, in order to generate <b> many training and validation sets </b>. For each partition of the data, we compute one residual error $E_{sel}$, and we select the model that minimizes the average error over all the training+validation partitions.
In practice also, one key parameter in this procedure is the relative size of the training and validation datasets. Having a large training dataset allows more accurate parameter estimation, hence predictions that are closer to what the model is capable of achieving. But that leaves the validation set with less datapoints to really correctly estimate this prediction, and therefore to discriminate competing models. Depending on the type of model, the optimal ratio is different. But as a rule of thumb, the validation dataset should be quite larger than the training dataset, e.g. 80% of the total dataset.
<b> Careful with model selection! </b>
We stress that noise in data, which is unavoidable in biology, has the potential to increase the score of some models compared to other models, regardless of the selection criteria chosen. Therefore, the more models are compared, the more likely one of them “benefits” from the particular data sample, increasing the odds that we eventually select a winner model by “chance” rather than because it is indeed the best model. Hence, model selection should not be followed blindly: estimating the likelihood of other models, comparing different selections methods, studying how sensitive the selection is to a particular data sample by, for instance, randomly splitting the dataset in two or more datasets and performing parameter optimization and model selection separately on each of them, all are possible approaches to “temper” the conclusions of a model selection procedure. And, eventually, picking the right model. Or even better, <b> NOT </b> picking a wrong one and convincing all the community that the Graal has been found.
## Parameters and model predictions
Once we have optimized (some) parameters of a model on some experimental dataset, and used model selection procedures to eliminate all models that are highly unlikely, we are generally left with 1-2 models which we <b> know </b> describe well what has been observed. At this stage, the model might already be able to provide insight on the biological mechanisms underlying some observations, since the biological hypotheses underlying the models excluded by the model selection are now part of the “unlikely” world. But another fundamental aspect of modeling is the ability to make testable predictions, in order to:
- either select among the remainder plausible models
- validate or invalidate the winner model
- or explore new biology! Which is the most exciting part.
How the validation experiments are designed, and how the prediction is shaped depends on the particular purpose of a prediction. But the very first step is to be able to make a prediction. And in mathematical modeling, making a prediction is answering the question: <b> if I change this and/or this parameter experimentally, what change, if any, can I expect from my biological system ?</b>
This is the reason why the very first task to perform after having optimized a model to recapitulate available data, and before making plans for validation experiments, is to explore the parameter space.
### 1) Parameter space analysis
Exploring the parameter space means computing the changes in model output(s) associated with any possible changes in parameter values. This includes single parameter changes, but also multiple simultaneous parameter changes… Of course, we can left aside the parameters which have been constrained (if not accurately determined) by the data-based parameter optimization. But even those parameters might be tuned experimentally (e.g., even if we have fitted a complex dissociation constant $K_d$ to some data doesn’t mean it’s not interesting to study what happens in presence of proteins that are mutated to decrease their relative affinity, yielding a larger $K_d$...). Generally speaking, in principle all parameters are worth of interest. So the exploration of the entire parameter space rapidly become intractable.
This is why it is strongly recommended to <b> incorporate as many parameters as possible in re-normalized variables </b>, as we did for the space-dependent diffusive-logistic growth equation in which all 3 parameters (diffusion coefficient, growth rate and saturation value) have been incorporated in renormalized time, space, and variable amplitude. Indeed, the overall effect of such parameters now becomes trivial, and they might be excluded from the parameter space analysis. For instance, if we have a model $M$ where one parameter $k$ is “absorbed” in a pseudo-time $\tau = k*t$, and we have explored the parameter space defined from all other and found the system’s dynamical regimes as a function of $\tau$, then in “real” time $t$ the same regimes are accessed at time $\tau/k$. Period.
Renormalizing time, space, and all other biological variables will considerably reduce the amount of other parameters, and generally will be associated with renormalized version of those parameters. Hence, “clusters” of parameters shall be treated as single parameters, this will make the parameter space exploration easier. For instance, in the gene transcription model the steady state is:
$$P_c=\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}$$
$$P_{pic} = \frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}$$
$$P_e=\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}$$
$$P_p=\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}*\frac{P_{tot}}{1+\frac{k_{on}}{ k_{off}+ k_{elong}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}+\frac{k_{on}}{ k_{off}+ k_{elong}}*\frac{ k_{elong}}{ k_{term}}*\frac{k_p}{k_q}}.$$
And in fact, there are only 4 parameters in the steady state, and as long as we are performing parameter space exploration for steady state variables, we can define only 4 parameters:
$$P_{tot},\frac{k_{on}}{ k_{off}+ k_{elong}},\frac{ k_{elong}}{ k_{term}},\frac{k_p}{k_q}$$
The first one $ P_{tot}$ can be absorbed in the renormalization of all polymerase populations to the total population, which leaves us with “only” 3 effective parameters, $K_1=\frac{k_{on}}{ k_{off}+ k_{elong}},K_2=\frac{ k_{elong}}{ k_{term}},K_3=\frac{k_p}{k_q}$, and with the following expressions for the renormalized polymerase populations in each stage of the process, in terms of these effective, independent parameters:
$$\frac{P_c}{P_{tot}}=\frac{1}{1+K_1+K_1*K_2+ K_1*K_2*K_3}$$
$$\frac{P_{pic}}{P_{tot}} = \frac{K_1}{1+K_1+K_1*K_2+ K_1*K_2*K_3}$$
$$\frac{P_e}{P_{tot}}=\frac{K_1*K_2}{1+K_1+K_1*K_2+ K_1*K_2*K_3}$$
$$\frac{P_p}{P_{tot}}=\frac{K_1*K_2*K_3}{1+K_1+K_1*K_2+ K_1*K_2*K_3}.$$
We could even make the equations look simpler, by renaming $K_1*K_2$ and $K_1*K_2*K_3$. But for the purpose of reducing the extent of the parameter space, this is not useful.
Following this preliminary reduction of the parameter space to a minimal number of <b> free, independent effective parameters, </b> the outputs of the model need to be evaluated for all biologically plausible values of these parameters. Each parameter can span orders of magnitude, and in non-linear models or models with complex dynamics, some parameters can “interact” in a way that interesting responses of the system are achieved only upon a coordinated change in several parameters. Thus, studying independently the consequences of the parameter variations one at a time is generally insufficient. 3D-dimensional plots can be used to represent graphically outputs with two varying parameters, but that can be insufficient.
A way to circumvent this limitation of the physical space to 3-dimensions, which allows only to represent graphically in a exhaustive way the variations with respect to 2 parameters is to generate output model data using <b> a large number of sets of randomly chosen parameters </b>. In this dataset, each set of parameters will be associated with predicted values for all model outputs. Then, the profiles of output data can be classified/sorted in different classes of model response regimes, and for each response, the parameter ranges yielding this response are automatically identified. This technique has the advantage to rapidly explore the entire parameter space, and to reveal particular combinations of values/ranges of different parameters associated with a given set of model outputs values/ranges. This approach can be complemented with a standard model output plotting in function of parameters, which is made easier once the different regions of the parameter space and their association with model response regimes have been identified. This method has the advantage to have no limitation in the number of original parameters, and to be equally applicable to models where an explicit analytical solution has been found, and to models solved or simulated numerically (with a computational cost for the latter).
### 2) Sensitivity, identifiability, and validation experiments design
The exploration of the parameter space allows to identify parameter ranges where new biology happens, and to suggest experiments which have never been attempted before. This is the <b> final purpose of mathematical modeling in biology and biomedical sciences.</b> Once such a parameter range has been identified, it is time to plan the experiment.
Any model is a simplification of the reality and comes with assumptions, among which are the “known” parameters, for instance pulled from the literature or inferred from previous datasets. Paraphrasing the famous economist Edward Leamer, model predictions are robust if they are valid over a range of assumptions and parameter values “wide enough to be credible”, and are themselves associated with a confidence interval “narrow enough to be useful”. If predictions are too sensitive to the uncertainty in the assumptions or “known” input parameters, they are unlikely to ever happen. If they are not accurate enough, if they don’t provide enough “contrast”, then the experimental signature of our prediction will be drowned in biological noise and we’ll struggle to convince the community that we have found something new.
Imagine we are modeling a biological mechanism, and our model predicts that a certain parameter $\alpha_0$ is a key determinant of the mechanism, in a parameter regime where the other parameters $\alpha_1,\alpha_2…$ are within biologically reasonable ranges. Then, the condition that “the prediction must not be too sensitive to the uncertainty in the assumptions or “known” input parameters” can be summarized as:
$$\frac{\partial Y}{\partial \alpha_1}\approx \frac{\partial Y}{\partial \alpha_2}\approx … \approx 0$$
and the condition “ the prediction is accurate enough, provides enough contrast” can be summarized as:
$$\frac{\partial Y}{\partial \alpha_0}=maximal.$$
Hence, the problem of finding the ideal experiment to demonstrate that $\alpha_0$ is a key determinant of the mechanism is to <b> find a biological variable </b> $Y$ <b> that can be expressed in terms of the model variable </b> and which is such that $\frac{\partial Y}{\partial \alpha_1}, \frac{\partial Y}{\partial \alpha_2} … $ are as small as possible, and $\frac{\partial Y}{\partial \alpha_0}$ as large as possible (in absolute value). This is a sensitivity problem: we seek for a variable which is very sensitive to the parameter of interest, and not sensitive to others, and these conditions have to be realized in the range of parameter values where the experiment will be performed.
Let’s illustrate this on our gene transcription example: imagine we want to demonstrate that RNA Pol II pauses when transcribing certain genes, and not pausing when transcribing others. Which variable should we measure?
Because we want to compare across different samples (i.e., different genes) what the pausing situation is (pausing rate $k_p$), we seek a model variable which is very sensitive to $k_p$ (or, equivalently, the renormalized parameter $K_3$, i.e. a model output for which the difference between the low $K_3$ and high $K_3$ regimes is emphasized.
Below is a Python code that plots the 4 output variables $P_c, P_{pic},P_e,P_p$ as a function of $K_1$ for $K_2$ and $K_3$ fixed, $K_2$ for $K_1$ and $K_3$ fixed, $K_3$ for $K_1$ and $K_2$ fixed. Using (and modifying!) this script to explore the parameter space, answer the following questions:
<b> Exercise 1: </b> Which model output seems to provide the best contrast between the low and high $K_3$ regimes, for arbitrary values of $K_1$ and $K_2$ ? Is there a particular region of the parameter space where this sensitivity is optimal? Is this variable sensitive also to $K_1$ and $K_2$, in the different $K_3$ regimes? Finally, how would you try to implement the experiment?
```
# %load solutionCodeExercise1.py
#%run solutionExercise1.py
```
Of course, this model is too simplistic to fully explain transcription, and the predicted parameter regimes for the optimization of the experimental detection of the pausing kinetics might be not so accurate. But yet, this analysis predicts which trends to follow when preparing the prediction validation experiments.
We stress that even if the optimal conditions for sensitivity cannot be achieved in practice, at least the <b> planned measurement should ensure the identifiability of the model parameters we want to access by the experiments, and compare across samples </b>. The (anticipated) identifiability or parameters in the planned experiments can be estimated following the identifiability test procedures mentioned above on simulated experimental data that resembles the kind of data we expect from the experiment.
We also stress that biological noise tends to reduce the accuracy to which observations might fit predictions. We all know this. But if we are working with a stochastic model, the <b> predicted biological noise i.e. the model output variability from simulation to simulation can be itself considered as an output variable </b>. As such, the predicted output variance is an output that might be the most sensitive output so certain parameters, and might also not be sensitive to other parameters so it might be the <b> best choice of model output variable to measure </b>. We can fit for model parameters using an error-function based on the comparison of the <b> predicted variance </b> of a given (stochastic) output $X$. In this case,
$$O^i_{pred}=\sigma^2_X(\alpha_1,\alpha_2 … \alpha_N)$$
and depends on model parameters $(\alpha_1,\alpha_2 … \alpha_N)$, and the measured data we compare this predicted output is the <b> measured variance </b> $O^i_{meas}=\sigma^2_X$ computed from the different measurements of $X$. The error function read:
$$E(\alpha_1,\alpha_2 … \alpha_N) = \sum_{i=1}^{i_0} w_i*\displaystyle \Bigg( O^i_{meas}- O^i_{pred}(\alpha_1,\alpha_2 … \alpha_N) \Bigg)^2 $$
and we can redo all we have done so far with this new “observable”. And in fact, all moments of the distribution of a model output $X$, not only the variance (which is a second order moment), can be considered as model outputs and compared with similar measurements.
Finally, the strategy underlined above to optimize the design of a validation to access a particular model parameter also applies to the design of an experiment that intends to distinguish between two models. In this situation, we know the data generated by the test experiment will be analyzed in the framework of the two models and the ability of the two models to fit the data will be assessed by model selection criterions (AIC/BIC… with a penalty to model complexity). In this scenario, we will look for an output variable $Y$ that <b> maximizes the difference in AIC/BIC between the two models </b>, and this will be equivalent to <b> maximizing the difference between the value of the error function (or the likelihood function) between the two models </b>. This can be achieved either by finding an output that the model 1 will be able to fit well but not the model 2, or conversely, or just an output that will be fitted a tiny bit better by model 1 and a tiny bit less well by model 2.
If we look one last time at our two models for single gene transcription, one such experiment could include for instance the ratio of elongating polymerases to the nucleoplasmic, unbound polymerases. Indeed, in the most simple model this ratio depends only on one ratio of rates (one independent parameter), whereas in the more complex model, it depends on two ratios (2 independent parameters). If for some reasons there one of the parameters in constrained in each case, the model 2 is susceptible to perform better in the fitting of this output because it has an extra, possibly unconstrained independent parameter to optimize the fit.
| github_jupyter |
<a href="https://colab.research.google.com/github/bs3537/dengueAI/blob/master/V11_San_Juan_XGB_environmental_and_time_features_PDP_Plot.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
#https://www.drivendata.org/competitions/44/dengai-predicting-disease-spread/page/80/
#Your goal is to predict the total_cases label for each (city, year, weekofyear) in the test set.
#Performance metric = mean absolute error
```
##LIST OF FEATURES:
You are provided the following set of information on a (year, weekofyear) timescale:
(Where appropriate, units are provided as a _unit suffix on the feature name.)
###City and date indicators
1. city – City abbreviations: sj for San Juan and iq for Iquitos
2. week_start_date – Date given in yyyy-mm-dd format
###NOAA's GHCN daily climate data weather station measurements
1. station_max_temp_c – Maximum temperature
2. station_min_temp_c – Minimum temperature
3. station_avg_temp_c – Average temperature
4. station_precip_mm – Total precipitation
5. station_diur_temp_rng_c – Diurnal temperature range
###PERSIANN satellite precipitation measurements (0.25x0.25 degree scale)
6. precipitation_amt_mm – Total precipitation
###NOAA's NCEP Climate Forecast System Reanalysis measurements (0.5x0.5 degree scale)
7. reanalysis_sat_precip_amt_mm – Total precipitation
8. reanalysis_dew_point_temp_k – Mean dew point temperature
9. reanalysis_air_temp_k – Mean air temperature
10. reanalysis_relative_humidity_percent – Mean relative humidity
11. reanalysis_specific_humidity_g_per_kg – Mean specific humidity
12. reanalysis_precip_amt_kg_per_m2 – Total precipitation
13. reanalysis_max_air_temp_k – Maximum air temperature
14. reanalysis_min_air_temp_k – Minimum air temperature
15. reanalysis_avg_temp_k – Average air temperature
16. reanalysis_tdtr_k – Diurnal temperature range
###Satellite vegetation - Normalized difference vegetation index (NDVI) - NOAA's CDR Normalized Difference Vegetation Index (0.5x0.5 degree scale) measurements
17. ndvi_se – Pixel southeast of city centroid
18. ndvi_sw – Pixel southwest of city centroid
19. ndvi_ne – Pixel northeast of city centroid
20. ndvi_nw – Pixel northwest of city centroid
####TARGET VARIABLE = total_cases label for each (city, year, weekofyear)
```
import sys
#Load train features and labels datasets
train_features = pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_features_train.csv')
train_features.head()
train_features.shape
train_labels = pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_labels_train.csv')
train_labels.head()
train_labels.shape
#Merge train features and labels datasets
train = pd.merge(train_features, train_labels)
train.head()
train.shape
#city, year and week of year columns are duplicate in train_features and train_labels datasets so the total_cases column is added to the features dataset
train.dtypes
#Data rows for San Juan
train.city.value_counts()
#San Juan has 936 rows which we can isolate and analyze separately
train = train[train['city'].str.match('sj')]
train.head(5)
train.shape
#Thus, we have isolated the train dataset with only city data for San Juan
#Distribution of the target
import seaborn as sns
sns.distplot(train['total_cases'])
#The target distribution is skewed
#Find outliers
train['total_cases'].describe()
#Remove outliers
train = train[(train['total_cases'] >= np.percentile(train['total_cases'], 0.5)) &
(train['total_cases'] <= np.percentile(train['total_cases'], 99.5))]
train.shape
sns.distplot(train['total_cases'])
#Do train, val split
from sklearn.model_selection import train_test_split
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
random_state=42)
train.shape, val.shape
#Load test features dataset (for the competition)
test = pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_features_test.csv')
#Pandas Profiling
```
#####Baseline statistics (mean and MAE) for the target variable total_cases in train dataset and baseline validation MAE
```
train['total_cases']. describe()
#Baseline mean and mean absolute error
guess = train['total_cases'].mean()
print(f'At the baseline, the mean total number of dengue cases in a year is: {guess:.2f}')
#If we had just guessed that the total number of dengue cases was 31.58 for a city in a particular year, we would be off by how much?
from sklearn.metrics import mean_absolute_error
# Arrange y target vectors
target = 'total_cases'
y_train = train[target]
y_val = val[target]
# Get mean baseline
print('Mean Baseline (using 0 features)')
guess = y_train.mean()
# Train Error
y_pred = [guess] * len(y_train)
mae = mean_absolute_error(y_train, y_pred)
print(f'Train mean absolute error: {mae:.2f} dengue cases per year')
# Test Error
y_pred = [guess] * len(y_val)
mae = mean_absolute_error(y_val, y_pred)
print(f'Validation mean absolute error: {mae:.2f} dengue cases per year')
#we need to convert week_start_date to numeric form uisng pd.to_dateime function
#wrangle function
def wrangle(X):
X = X.copy()
# Convert week_start_date to numeric form
X['week_start_date'] = pd.to_datetime(X['week_start_date'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['week_start_date'].dt.year
X['month_recorded'] = X['week_start_date'].dt.month
#X['day_recorded'] = X['week_start_date'].dt.day
X = X.drop(columns='week_start_date')
X = X.drop(columns='year')
X = X.drop(columns='station_precip_mm')
#I engineered few features which represent standing water, high risk feature for mosquitos
#1. X['standing water feature 1'] = X['station_precip_mm'] / X['station_max_temp_c']
#Standing water features
X['total satellite vegetation index of city'] = X['ndvi_se'] + X['ndvi_sw'] + X['ndvi_ne'] + X['ndvi_nw']
#Standing water features
#Standing water feature 1 = 'NOAA GCN precipitation amount in kg per m2 reanalyzed' * (total vegetation, sum of all 4 parts of the city)
X['standing water feature 1'] = X['reanalysis_precip_amt_kg_per_m2'] * X['total satellite vegetation index of city']
#Standing water feature 2: 'NOAA GCN precipitation amount in kg per m2 reanalyzed'} * 'NOAA GCN mean relative humidity in pct reanalyzed'}
X['standing water feature 2'] = X['reanalysis_precip_amt_kg_per_m2'] * X['reanalysis_relative_humidity_percent']
#Standing water feature 3: 'NOAA GCN precipitation amount in kg per m2 reanalyzed'} * 'NOAA GCN mean relative humidity in pct reanalyzed'} * (total vegetation)
X['standing water feature 3'] = X['reanalysis_precip_amt_kg_per_m2'] * X['reanalysis_relative_humidity_percent'] * X['total satellite vegetation index of city']
#Standing water feature 4: 'NOAA GCN precipitation amount in kg per m2 reanalyzed'} / 'NOAA GCN max air temp reanalyzed'
X['standing water feature 4'] = X['reanalysis_precip_amt_kg_per_m2'] / X['reanalysis_max_air_temp_k']
#Standing water feature 5: ['NOAA GCN precipitation amount in kg per m2 reanalyzed'} * 'NOAA GCN mean relative humidity in pct reanalyzed'} * (total vegetation)]/['NOAA GCN max air temp reanalyzed']
X['standing water feature 5'] = X['reanalysis_precip_amt_kg_per_m2'] * X['reanalysis_relative_humidity_percent'] * X['total satellite vegetation index of city'] / X['reanalysis_max_air_temp_k']
#Rename columns
X.rename(columns= {'reanalysis_air_temp_k':'Mean air temperature in K'}, inplace=True)
X.rename(columns= {'reanalysis_min_air_temp_k':'Minimum air temperature in K'}, inplace=True)
X.rename(columns= {'weekofyear':'Week of Year'}, inplace=True)
X.rename(columns= {'station_diur_temp_rng_c':'Diurnal temperature range in C'}, inplace=True)
X.rename(columns= {'reanalysis_precip_amt_kg_per_m2':'Total precipitation kg/m2'}, inplace=True)
X.rename(columns= {'reanalysis_tdtr_k':'Diurnal temperature range in K'}, inplace=True)
X.rename(columns= {'reanalysis_max_air_temp_k':'Maximum air temperature in K'}, inplace=True)
X.rename(columns= {'year_recorded':'Year recorded'}, inplace=True)
X.rename(columns= {'reanalysis_relative_humidity_percent':'Mean relative humidity'}, inplace=True)
X.rename(columns= {'month_recorded':'Month recorded'}, inplace=True)
X.rename(columns= {'reanalysis_dew_point_temp_k':'Mean dew point temp in K'}, inplace=True)
X.rename(columns= {'precipitation_amt_mm':'Total precipitation in mm'}, inplace=True)
X.rename(columns= {'station_min_temp_c':'Minimum temp in C'}, inplace=True)
X.rename(columns= {'ndvi_se':'Southeast vegetation index'}, inplace=True)
X.rename(columns= {'ndvi_ne':'Northeast vegetation index'}, inplace=True)
X.rename(columns= {'ndvi_nw':'Northwest vegetation index'}, inplace=True)
X.rename(columns= {'ndvi_sw':'Southwest vegetation index'}, inplace=True)
X.rename(columns= {'reanalysis_avg_temp_k':'Average air temperature in K'}, inplace=True)
X.rename(columns= {'reanalysis_sat_precip_amt_mm':'Total precipitation in mm (2)'}, inplace=True)
X.rename(columns= {'reanalysis_specific_humidity_g_per_kg':'Mean specific humidity'}, inplace=True)
X.rename(columns= {'station_avg_temp_c':'Average temp in C'}, inplace=True)
X.rename(columns= {'station_max_temp_c':'Maximum temp in C'}, inplace=True)
X.rename(columns= {'total_cases':'Total dengue cases in the week'}, inplace=True)
#Drop columns
X = X.drop(columns='Total precipitation in mm (2)')
X = X.drop(columns='Average temp in C')
X = X.drop(columns='Maximum temp in C')
X = X.drop(columns='Minimum temp in C')
X = X.drop(columns='Diurnal temperature range in C')
X = X.drop(columns='Average air temperature in K')
X = X.drop(columns='city')
# return the wrangled dataframe
return X
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
train.head().T
train.dtypes
#Before we build the model to train on train dataset, log transform target variable due to skew
import numpy as np
target_log = np.log1p(train['Total dengue cases in the week'])
sns.distplot(target_log)
plt.title('Log-transformed target');
target_log_series = pd.Series(target_log)
train = train.assign(log_total_cases = target_log_series)
#drop total_cases target column while training the model
train = train.drop(columns='Total dengue cases in the week')
#Do the same log transformation with validation dataset
target_log_val = np.log1p(val['Total dengue cases in the week'])
target_log_val_series = pd.Series(target_log_val)
val = val.assign(log_total_cases = target_log_val_series)
val = val.drop(columns='Total dengue cases in the week')
#Fitting XGBoost Regresser model
#Define target and features
# The status_group column is the target
target = 'log_total_cases'
# Get a dataframe with all train columns except the target
train_features = train.drop(columns=[target])
# Get a list of the numeric features
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
# Combine the lists
features = numeric_features
# Arrange data into X features matrix and y target vector
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
pip install category_encoders
from sklearn.pipeline import make_pipeline
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import OneHotEncoder
import xgboost as xgb
from xgboost import XGBRegressor
from sklearn import model_selection, preprocessing
pipeline = make_pipeline(
SimpleImputer(strategy='mean'),
XGBRegressor(n_estimators=137, objective='reg:squarederror', n_jobs=-1)
)
pipeline.fit(X_train, y_train)
#predict on X_val
y_pred = pipeline.predict(X_val)
print('XGBoost Validation Mean Absolute Error, log transformed)', mean_absolute_error(y_val, y_pred))
#Transform y_pred back to original units from log transformed
y_pred_original = np.expm1(y_pred)
y_val_original = np.expm1(y_val)
print('XGBoost Validation Mean Absolute Error (non-log transformed)', mean_absolute_error(y_val_original, y_pred_original))
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 72
pip install pdpbox
from pdpbox.pdp import pdp_isolate, pdp_plot
feature='Week of Year'
isolated = pdp_isolate(
model = pipeline,
dataset = X_val,
model_features = X_val.columns,
feature = feature
)
pdp_plot(isolated, feature_name=feature)
```
| github_jupyter |
```
// #r ".\binaries2\bossspad.dll"
// #r ".\binaries2\XNSEC.dll"
// #r "C:\BoSSS_Binaries\bossspad.dll"
// #r "C:\BoSSS_Binaries\XNSEC.dll"
#r "C:\BoSSS\experimental\public\src\L4-application\BoSSSpad\bin\Release\net5.0\bossspad.dll"
#r "C:\BoSSS\experimental\public\src\L4-application\BoSSSpad\bin\Release\net5.0\XNSEC.dll"
using System;
using System.Collections.Generic;
using System.Linq;
using System.IO;
using System.Data;
using System.Globalization;
using System.Threading;
using ilPSP;
using ilPSP.Utils;
using BoSSS.Platform;
using BoSSS.Foundation;
using BoSSS.Foundation.Grid;
using BoSSS.Foundation.Grid.Classic;
using BoSSS.Foundation.IO;
using BoSSS.Solution;
using BoSSS.Solution.Control;
using BoSSS.Solution.GridImport;
using BoSSS.Solution.Statistic;
using BoSSS.Solution.Utils;
using BoSSS.Solution.Gnuplot;
using BoSSS.Application.BoSSSpad;
using static BoSSS.Application.BoSSSpad.BoSSSshell;
using BoSSS.Foundation.Grid.RefElements;
using BoSSS.Platform.LinAlg;
using BoSSS.Solution.NSECommon;
using BoSSS.Application.XNSEC;
Init();
```
## Case configuration
```
int[] dgDegree = new int[]{1,2,3,4};
int[] Resolutions = new int[]{3,4,5,6,7,8,9};
int numberOfMpiCores =4;
var NofCells = Resolutions.ToList().Select(r=> (int)Math.Pow(2,r)).ToArray();
string baseJobName = "ChamberedFlame";
NofCells
```
## Open Database
```
// static var myDb = OpenOrCreateDatabase(@"C:\Databases\ChamberedDiffFlame5");
static var myDb = OpenOrCreateDatabase(@"\\hpccluster\hpccluster-scratch\gutierrez\UnstrainedFlame_100ItMax_unitylew_noVariableKin_ConstantCP_highervel");
// static string dirname ="CounterDiffusionFlame_HLLC222";
// static string winpath = @"S:\work\scratch\jg11bano\"+dirname;
// static string remotepath = @"/work/scratch/jg11bano/"+dirname;
// static var myDb = OpenOrCreateDatabase(@"S:\work\scratch\jg11bano\CDF_FirstTryHLLC");
// static var myDb = OpenOrCreateDatabase(winpath);
myDb.Summary()
BoSSSshell.WorkflowMgm.Init("ChamberedFlame4");
// BoSSSshell.WorkflowMgm.SetNameBasedSessionJobControlCorrelation();
BoSSSshell.WorkflowMgm.Sessions
var myBatch = BoSSSshell.ExecutionQueues[3];
// MiniBatchProcessor.Server.StartIfNotRunning(true);
myBatch.AllowedDatabasesPaths.Add(new AllowedDatabasesPair(myDb.Path,""));
// ==================================
// setup Client & Workflow & Database
// ==================================
// var myBatch = (SlurmClient)ExecutionQueues[1];
// var AddSbatchCmds = new List<string>();
// AddSbatchCmds.AddRange(new string[]{"#SBATCH -N 4","#SBATCH -p test24", "#SBATCH -C avx512", "#SBATCH --mem-per-cpu="+2000});
// myBatch.AllowedDatabasesPaths.Add(new AllowedDatabasesPair(myDb.Path,""));
// myBatch.AdditionalBatchCommands = AddSbatchCmds.ToArray();
// myBatch.AdditionalBatchCommands
```
## Create grid
```
public static class GridFactory {
public static Grid2D GenerateGrid(int nCells) {
double sepAdim = 20 ; // 20 is already ok!
double xleft = -sepAdim;
double xright = 0.0;
double R = sepAdim / 10;
var _xNodes = GenericBlas.Linspace(xleft, xright, nCells + 1);
double[] _yNodes = GenericBlas.Linspace(-R, R, 3 + 1);
string BC = "ScalarDirichlet_PressureOutlet";
Console.WriteLine("Number of cells in the X direction: {0}", _xNodes.Length);
Console.WriteLine("Number of cells in the Y direction: {0}", _yNodes.Length);
var grd = Grid2D.Cartesian2DGrid(_xNodes, _yNodes, periodicY: true);
grd.EdgeTagNames.Add(1, "Velocity_Inlet_CH4");
grd.EdgeTagNames.Add(2, BC);
grd.DefineEdgeTags(delegate (double[] X) {
double x = X[0];
double y = X[1];
if (Math.Abs(x - xleft) < 1e-8) { // Left boundary
return 1; // Left
} else if (Math.Abs(x - xright) < 1e-8) { // right boundary
return 2; // right
} else {
return 3;
// throw new Exception("Problem while setting GridEdgeTagFunc");
}
}
);
myDb.SaveGrid(ref grd, true);
return grd;
}
}
public static class BoundaryValueFactory {
public static string GetPrefixCode(double ConstVal) {
using(var stw = new System.IO.StringWriter()) {
stw.WriteLine("static class BoundaryValues {");
stw.WriteLine(" static public double ConstantValue(double[] X) {");
stw.WriteLine(" return "+ ConstVal +";");
stw.WriteLine(" }");
stw.WriteLine("}");
return stw.ToString();
}
}
static public Formula Get_ConstantValue(double ConstVal){
return new Formula("BoundaryValues.ConstantValue", AdditionalPrefixCode:GetPrefixCode(ConstVal));
}
}
```
## Create base control file
In this ControlFile basic configuration of the ChamberedDiffusionFlame is defined.
```
static XNSEC_Control GiveMeTheCtrlFile(int dg, int nCells, bool isMF) {
var CC = new ChemicalConstants();
var C = isMF ? new XNSEC_MF_Control() : new XNSEC_Control();
// C.AlternateDbPaths = new[] {
// (@"S:\work\scratch\jg11bano\"+dirname, "PCMIT30"),
// (@"/work/scratch/jg11bano/"+dirname,"")};
// C.AlternateDbPaths = new[] {
// (winpath, ""),
// (dirname,"")};
// C.AlternateDbPaths = new[]{
// new ValueTuple<string,string>(@"/work/scratch/jg11bano/bosss_db", ""),
// new ValueTuple<string,string>(@"Y:\bosss_db", "")
// };
C.NumberOfChemicalSpecies = 4;
C.SetDGdegree(dg); //
C.SetGrid(GridFactory.GenerateGrid(nCells)); //
C.MatParamsMode = MaterialParamsMode.Sutherland; //
// Problem Definition
//===================
double TemperatureIn = 300;
double massFuelIn = 0.24 * 0.1; //kg/m2s
double AtmPressure = 101325; // Pa
double[] FuelInletConcentrations = new double[] { 0.2, 0.0, 0.0, 0.0, 0.8 };
double[] OxidizerInletConcentrations = new double[] { 0.0, 0.23, 0.0, 0.0, 0.77 };
double[] MWs = new double[] { CC.MW_CH4, CC.MW_O2, CC.MW_CO2, CC.MW_H2O, CC.MW_N2 };
double mwFuel = CC.getAvgMW(MWs, FuelInletConcentrations);
double mwAir = CC.getAvgMW(MWs, OxidizerInletConcentrations);
double densityFuelIn = AtmPressure * mwFuel / (CC.R_gas * TemperatureIn * 1000); // Kg/m3. ok
double uInFuel = massFuelIn / densityFuelIn; //
Console.WriteLine("Velocity is "+uInFuel);
// Reference values
//===================
// Basic units to be used: Kg, m, s, mol, pa,
double TRef = TemperatureIn;// Reference temperature is the inlet temperature, (K)
double pRef = AtmPressure; // Pa
double uRef = uInFuel; // m/s
double rhoRef = pRef * mwAir / (8.314 * TRef * 1000); // Kg/m3. ok ;
double DRef = 2.2133029473872009E-05 / rhoRef;
double LRef = DRef / uRef;
C.GravityDirection = new double[] { 0.0, 0.0, 0.0 }; //No gravity.
// Solver configuration
// =======================
C.smoothingFactor = 80*0-1*1;
// C.NonLinearSolver.ConvergenceCriterion = 1e-8;
// C.LinearSolver.ConvergenceCriterion = 1e-10;
C.NonLinearSolver.verbose = true;
C.NonLinearSolver.SolverCode = NonLinearSolverCode.Newton;
C.NonLinearSolver.MaxSolverIterations = 10;
C.LinearSolver.SolverCode = LinearSolverCode.classic_pardiso;
C.LinearSolver.verbose = false;
C.TimesteppingMode = AppControl._TimesteppingMode.Steady;
C.saveperiod = 1;
C.PenaltyViscMomentum = 1.0; ///////////////////////////////////////////////////////////////////////////////////////////////////////////////////////
C.PenaltyHeatConduction = 1.0;///////////////////////////////////////////////////////////////////////////////////////////////////////////////////////
C.YFuelInlet = FuelInletConcentrations[0];
C.YOxInlet = OxidizerInletConcentrations[1];
C.FuelInletConcentrations = FuelInletConcentrations;
C.OxidizerInletConcentrations = OxidizerInletConcentrations;
C.TFuelInlet = 1.0;
C.TOxInlet = 1.0;
C.PhysicalParameters.IncludeConvection = true;
// Chemical related parameters
double s = (CC.nu_O2 * CC.MW_O2) / (CC.nu_CH4 * CC.MW_CH4);
C.phi = s * C.YFuelInlet / C.YOxInlet;
C.zSt = 1.0 / (1.0 + C.phi);
var MLC = new MaterialLawCombustion(300, new double[] { }, C.MatParamsMode, C.rhoOne, true, 1.0, 1, 1, C.YOxInlet, C.YFuelInlet, C.zSt, CC, 0.75);
var ThermoProperties = new ThermodynamicalProperties();
//==========================
//Derived reference values
//==========================
C.uRef = uRef; // Reference velocity
C.LRef = LRef; // reference length
C.pRef = AtmPressure; // reference pressure
C.TRef = TRef;// reference temperature
C.MWRef = MLC.getAvgMW(MWs, C.OxidizerInletConcentrations); // Air mean molecular weight
C.rhoRef = C.pRef * C.MWRef / (8.314 * C.TRef * 1000); // Kg/m3. ok ;
C.cpRef = 1.3;//ThermoProperties.Calculate_Cp_Mixture(new double[] { 0.23, 0.77 }, new string[] { "O2", "N2" }, 300); // 1.219185317353029;// Representative value, KJ/Kg K ========> 1.31 for the one-step kinetic model
C.muRef = MLC.getViscosityDim(300);
C.MolarMasses = new double[] { C.CC.MW_CH4, C.CC.MW_O2, C.CC.MW_CO2, C.CC.MW_H2O, C.CC.MW_N2 };
C.MolarMasses.ScaleV(1.0 / C.MWRef); //NonDimensionalized Molar masses
C.T_ref_Sutherland = 300;
double heatRelease_Ref = (C.TRef * C.cpRef);
C.HeatRelease = C.CC.HeatReleaseMass / heatRelease_Ref;
C.B = CC.PreExponentialFactor;
C.StoichiometricCoefficients = new double[] { -1, -2, 1, 2, 0 };
C.Damk = C.rhoRef * C.LRef * C.B / (C.uRef * C.MWRef);
C.Reynolds = C.rhoRef * C.uRef * C.LRef / C.muRef;
C.Prandtl = 0.75;////////////////////0.75;
C.Schmidt = C.Prandtl; // Because Lewis number is assumed as 1.0 (Le = Pr/Sc)
// // C.Lewis = new double[] { 0.97, 1.11, 1.39, 0.83, 1.0 };
C.Lewis = new double[] {1.0, 1.0, 1.0,1.0, 1.0 };
double g = 9.8; // m/s2
C.Froude = Math.Sqrt(uRef * uRef / (C.LRef * g)); // Not used
C.ReactionRateConstants = new double[] { C.Damk, CC.Ta / TRef, 1.0, 1.0 }; // NOTE! activation temperature is also nondimensional
//==========================
// Initial conditions
//==========================
double dummy = 0;
C.AddInitialValue(VariableNames.VelocityX, BoundaryValueFactory.Get_ConstantValue(0.0));
C.AddInitialValue(VariableNames.VelocityY, BoundaryValueFactory.Get_ConstantValue(0.0));
C.AddInitialValue(VariableNames.Pressure, BoundaryValueFactory.Get_ConstantValue(0.0));
//==========================
// Boundary conditions
//==========================
C.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.Velocity_d(0), BoundaryValueFactory.Get_ConstantValue(uInFuel / C.uRef));
C.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.Velocity_d(1), BoundaryValueFactory.Get_ConstantValue(0.0));
return C;
}
```
## Starting the MixtureFraction simulation
Configuration for the simulation using the mixture fraction approach, where an infinite reaction rate is assumed. Used to find adequate starting solution for the full problem.
```
static XNSEC_Control GiveMeTheMixtureFractionCtrlFile(int dg, int nCells){
var C_MixtureFraction = GiveMeTheCtrlFile(dg, nCells, true);
C_MixtureFraction.physicsMode = PhysicsMode.MixtureFraction;
C_MixtureFraction.ProjectName = "ChamberedFlame";
string name = C_MixtureFraction.ProjectName + "P" + dg + "K" + nCells;
C_MixtureFraction.SessionName = "FS_" + name;
C_MixtureFraction.UseSelfMadeTemporalOperator = false;
C_MixtureFraction.ChemicalReactionActive = false;
C_MixtureFraction.physicsMode = PhysicsMode.MixtureFraction;
C_MixtureFraction.NonLinearSolver.MaxSolverIterations = 50;
// Boundary and initial conditions
C_MixtureFraction.AddInitialValue(VariableNames.MixtureFraction,BoundaryValueFactory.Get_ConstantValue(1.0));
C_MixtureFraction.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.MixtureFraction, BoundaryValueFactory.Get_ConstantValue(1.0));
C_MixtureFraction.AdaptiveMeshRefinement = false;
C_MixtureFraction.TimesteppingMode = BoSSS.Solution.Control.AppControl._TimesteppingMode.Steady;
return C_MixtureFraction;
}
```
## Send and run jobs
```
bool HLLRCalculation = true;
foreach(int nCells in NofCells){
int dg = 2;
Type solver_MF = typeof(BoSSS.Application.XNSEC.XNSEC_MixtureFraction);
var C_MixtureFraction = GiveMeTheMixtureFractionCtrlFile(dg, nCells);
string jobName = C_MixtureFraction.SessionName;
Console.WriteLine(jobName);
var oneJob = new Job(jobName, solver_MF);
oneJob.NumberOfMPIProcs = 4;
// oneJob.UseComputeNodesExclusive = true;
oneJob.SetControlObject(C_MixtureFraction);
oneJob.Activate(myBatch);
}
BoSSSshell.WorkflowMgm.BlockUntilAllJobsTerminate();
```
## Starting the finite-rate chemistry simulation
Now that the simulation for an "infinite" reaction rate is done, we use it for initializing the system with finite reaction rate. The goal is to obtain solutions of the counter difussion flame for increasing strain values. We start with a low strain (bigger Dahmkoehler number), which is increased until extintion is (hopefully) found
```
static XNSEC_Control GiveMeTheFullCtrlFile(int dg, int nCells, ISessionInfo SessionToRestart) {
var C_OneStep = GiveMeTheCtrlFile(dg, nCells, false);
C_OneStep.physicsMode = PhysicsMode.Combustion;
C_OneStep.ProjectName = "ChamberedFlame";
string name = C_OneStep.ProjectName + "P" + dg + "K" + nCells;
C_OneStep.SessionName = "Full_" + name;
C_OneStep.VariableOneStepParameters = false; ///////////////////////
// C_OneStep.Tags.Add("VelocityMultiplier" + mult);
C_OneStep.UseSelfMadeTemporalOperator = false;
C_OneStep.myThermalWallType = SIPDiffusionTemperature.ThermalWallType.Adiabatic;
C_OneStep.Timestepper_LevelSetHandling = BoSSS.Solution.XdgTimestepping.LevelSetHandling.None;
C_OneStep.UseMixtureFractionsForCombustionInitialization = true;
C_OneStep.LinearSolver.SolverCode = LinearSolverCode.exp_Kcycle_schwarz;
C_OneStep.LinearSolver.NoOfMultigridLevels = 5;
C_OneStep.ChemicalReactionActive = true;
C_OneStep.AdaptiveMeshRefinement = false;
C_OneStep.HeatCapacityMode = MaterialLaw_MultipleSpecies.CpCalculationMode.constant;
C_OneStep.NoOfTimesteps = 1; // The steady solution will be calculated again and do AMR
C_OneStep.NonLinearSolver.MaxSolverIterations = 50;
// C_OneStep.NonLinearSolver.MaxSolverIterations = 10;
// limiting of variable values
Dictionary<string, Tuple<double, double>> Bounds = new Dictionary<string, Tuple<double, double>>();
double eps = 1e-2;
Bounds.Add(VariableNames.Temperature, new Tuple<double, double>(1.0 - eps, 10)); // Min temp should be the inlet temperature.
Bounds.Add(VariableNames.MassFraction0, new Tuple<double, double>(0.0 - 1e-1, 1.0 + 1e-1)); // Between 0 and 1 per definition
Bounds.Add(VariableNames.MassFraction1, new Tuple<double, double>(0.0 - 1e-1, 1.0 + 1e-1));
Bounds.Add(VariableNames.MassFraction2, new Tuple<double, double>(0.0 - 1e-1, 1.0 + 1e-1));
Bounds.Add(VariableNames.MassFraction3, new Tuple<double, double>(0.0 - 1e-1, 1.0 + 1e-1));
C_OneStep.VariableBounds = Bounds;
// Boundary conditions
double dummy = 0;
if(SessionToRestart != null) {
C_OneStep.SetRestart(SessionToRestart);
} else {
C_OneStep.AddInitialValue(VariableNames.Temperature, BoundaryValueFactory.Get_ConstantValue(1.0));
C_OneStep.AddInitialValue(VariableNames.MassFraction0, BoundaryValueFactory.Get_ConstantValue(0.0));
C_OneStep.AddInitialValue(VariableNames.MassFraction1, BoundaryValueFactory.Get_ConstantValue(0.23));
C_OneStep.AddInitialValue(VariableNames.MassFraction2, BoundaryValueFactory.Get_ConstantValue(0.0));
C_OneStep.AddInitialValue(VariableNames.MassFraction3, BoundaryValueFactory.Get_ConstantValue(0.0));
}
C_OneStep.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.Temperature, BoundaryValueFactory.Get_ConstantValue(1.0));
C_OneStep.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.MassFraction0, BoundaryValueFactory.Get_ConstantValue(C_OneStep.FuelInletConcentrations[0]));
C_OneStep.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.MassFraction1, BoundaryValueFactory.Get_ConstantValue(C_OneStep.FuelInletConcentrations[1]));
C_OneStep.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.MassFraction2, BoundaryValueFactory.Get_ConstantValue(C_OneStep.FuelInletConcentrations[2]));
C_OneStep.AddBoundaryValue("Velocity_Inlet_CH4", VariableNames.MassFraction3, BoundaryValueFactory.Get_ConstantValue(C_OneStep.FuelInletConcentrations[3]));
C_OneStep.AddBoundaryValue("ScalarDirichlet_PressureOutlet", VariableNames.Temperature, BoundaryValueFactory.Get_ConstantValue(1.0));
C_OneStep.AddBoundaryValue("ScalarDirichlet_PressureOutlet", VariableNames.MassFraction0, BoundaryValueFactory.Get_ConstantValue(C_OneStep.OxidizerInletConcentrations[0]));
C_OneStep.AddBoundaryValue("ScalarDirichlet_PressureOutlet", VariableNames.MassFraction1, BoundaryValueFactory.Get_ConstantValue(C_OneStep.OxidizerInletConcentrations[1]));
C_OneStep.AddBoundaryValue("ScalarDirichlet_PressureOutlet", VariableNames.MassFraction2, BoundaryValueFactory.Get_ConstantValue(C_OneStep.OxidizerInletConcentrations[2]));
C_OneStep.AddBoundaryValue("ScalarDirichlet_PressureOutlet", VariableNames.MassFraction3, BoundaryValueFactory.Get_ConstantValue(C_OneStep.OxidizerInletConcentrations[3]));
return C_OneStep;
}
Type solver = typeof(BoSSS.Application.XNSEC.XNSEC);NofCells
```
Calculate the full solution for the initial value
```
int counter = 0;
foreach (int nCells in NofCells) {
foreach (int dg in dgDegree) {
// var sess =(myDb.Sessions.Where(s=>Convert.ToInt64(s.Tags.ToArray()[0]) == i)).FirstOrDefault();
var sess = (myDb.Sessions.Where(s => s.Name == "FS_ChamberedFlameP" + 2 + "K" + nCells)).FirstOrDefault();
var C = GiveMeTheFullCtrlFile(dg, nCells, sess);
string jobName = C.SessionName + "AllOn";
Console.WriteLine(jobName);
var oneJob = new Job(jobName, solver);
oneJob.NumberOfMPIProcs = nCells > 32? 12:4 ;
oneJob.SetControlObject(C);
oneJob.Activate(myBatch);
counter++;
}
}
BoSSSshell.WorkflowMgm.BlockUntilAllJobsTerminate();
```
## Postprocessing
```
// Now the flame sheet estimations should be deleted
BoSSSshell.WorkflowMgm.Sessions.Where(s => s.Name.StartsWith("FS_")).ForEach(x=>x.Delete(true));
//WorkflowMgm.ResetSessionsCache();
string basepath = @"C:\tmp\UnstrainedFlame_ConvStudy";
System.IO.Directory.CreateDirectory(basepath); // Create folder for storing data
string[] varnames = new string[]{"VelocityX","VelocityY","Temperature", "Pressure",
"MassFraction0","MassFraction1","MassFraction2","MassFraction3" };
foreach (var varname in varnames) {
foreach (int pDeg in dgDegree) {
var pDegSessions = myDb.Sessions.Where(
Si => (Convert.ToInt32(Si.KeysAndQueries["DGdegree:Velocity*"]) == pDeg
&& (Convert.ToInt32(Si.KeysAndQueries["Grid:NoOfCells"]))/ 3 > 31
) // function which is true on all sessions we are interested in:
).ToArray();
Plot2Ddata pDegPlot =
pDegSessions.ToEstimatedGridConvergenceData(varname,
xAxis_Is_hOrDof: true, // false selects DOFs for x-axis
normType: NormType.L2_embedded );
//Saving to a txt file
pDegPlot.SaveTextFileToPublish(basepath+"\\"+ varname + "DG" + pDeg,false);
}
}
myDb.Sessions
```
| github_jupyter |
# Classification model to detect pneumonia
## Context
This model detects pneumonia based on chest x-ray images. It is a convolutional neural network, with transfer learning using VGG16. Its final accuracy on the test set is 89%. The data was taken from Kaggle [Chest X-Ray Images (Pneumonia)](https://www.kaggle.com/paultimothymooney/chest-xray-pneumonia).
The project was inspired by the article on AI detection of COVID-19 pneumonia, available on [Medical News](https://www.news-medical.net/news/20200409/Ai-used-to-detect-COVID-19-pneumonia.aspx). Majority of people with COVID-19 have mild symptoms such as coughing and fever. In some cases, however, severe pneumonia in both lungs can develop, which can be deadly. There is a significant pressure on the hospital staff during the outbreak, as well as the shortage of the the medical equipment and PPE. Potentially AI could allow the doctors to determine whether the patients require supportive care or could recover at home.
_"Pneumonia can be subtle, especially if it’s not your average bacterial pneumonia, and if we could identify those patients early before you can even detect it with a stethoscope, we might be better positioned to treat those at highest risk for severe disease and death",_ said Dr. Albert Hsiao in the article.
## Data
<img src='assets/intro_01.png' />
*Illustrative examples of chest x-rays in patients with pneumonia. The normal chest X-ray (left panel) depicts clear lungs without any areas of abnormal opacification in the image. Bacterial pneumonia (middle) typically exhibits a focal lobar consolidation, in this case in the right upper lobe (white arrows), whereas viral pneumonia (right) manifests with a more diffuse ‘‘interstitial’’ pattern in both lungs* [(Kermany DS, Goldbaum M, Cai W, et al. (2018))]('http://www.cell.com/cell/fulltext/S0092-86741830154-5).
Chest X-ray images (anterior-posterior) were selected from retrospective cohorts of pediatric patients of one to five years old from Guangzhou Women and Children’s Medical Center, Guangzhou. All chest X-ray imaging was performed as part of patients’ routine clinical care.
For the analysis of chest x-ray images, all chest radiographs were initially screened for quality control by removing all low quality or unreadable scans. The diagnoses for the images were then graded by two expert physicians before being cleared for training the AI system. In order to account for any grading errors, the evaluation set was also checked by a third expert.
```
import numpy as np
import pandas as pd
import os
from glob import glob
```
First, let's inspect the data that is available.
```
# loading the directories
trai_dir = '../input/chest-xray-pneumonia/chest_xray/train/'
val_dir = '../input/chest-xray-pneumonia/chest_xray/val/'
test_dir = '../input/chest-xray-pneumonia/chest_xray/test/'
# getting the number of classes
folders = glob(trai_dir + '/*')
num_classes = len(folders)
class_labels=os.listdir(trai_dir)
print ('Total number of classes = ' + str(num_classes))
print('Class names: {0}'.format(class_labels))
# getting number of files
train_files = np.array(glob(trai_dir+"*/*"))
val_files = np.array(glob(val_dir+"*/*"))
test_files = np.array(glob(test_dir+"*/*"))
# print number of images in each dataset
print('There are %d total train images.' % len(train_files))
print('There are %d total validation images.' % len(val_files))
print('There are %d total test images.' % len(test_files))
train_normal = np.array(glob(trai_dir+"NORMAL/*"))
val_normal = np.array(glob(val_dir+"NORMAL/*"))
test_normal = np.array(glob(test_dir+"NORMAL/*"))
train_pneumonia = np.array(glob(trai_dir+"PNEUMONIA/*"))
val_pneumonia = np.array(glob(val_dir+"PNEUMONIA/*"))
test_pneumonia = np.array(glob(test_dir+"PNEUMONIA/*"))
print('There are %d total normal train images.' % len(train_normal))
print('There are %d total normal validation images.' % len(val_normal))
print('There are %d total normal test images.' % len(test_normal))
print('There are %d total pneumonia train images.' % len(train_pneumonia))
print('There are %d total pneumonia validation images.' % len(val_pneumonia))
print('There are %d total pneumonia test images.' % len(test_pneumonia))
```
Data distribution is very uneven. There is signinficantly less pneumonia images (3875) than normal ones (14341). Moreover, the validation set is particularly small, only 8 images for both normal and pneumonia classes.
Let's display some sample unprocessed images of healthy and infected lungs.
```
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
file_path = np.concatenate((train_normal[0:3],train_pneumonia[0:3]))
fig = plt.figure(figsize=(13, 9))
for i in range(len(file_path)):
ax = fig.add_subplot(2,3,i+1, xticks=[], yticks=[])
img=mpimg.imread(file_path[i])
ax.set_title(file_path[i].split('/')[-2])
imgplot = plt.imshow(img, cmap='gray', vmin=0, vmax=255)
plt.show()
```
## Specify Data Loaders
Three separate data loaders for the training, validation and test datasets are created. Train and validation sets were resized to 256, randomly flipped and rotated up to 20 degrees. Next, all data sets have been cropped to 224 x 224 size images, similarly to other pretrained models available in torchvision.models module. Finally, the images were transformed to tensors and their channels were normalised with means of [0.485, 0.456, 0.406] and standard deviations of [0.229, 0.224, 0.225].
```
import torch
import torchvision.transforms as transforms
from torchvision import datasets
# Create training and test dataloaders
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 32
# resize the picture
size = 256
data_transform_train = transforms.Compose([
transforms.RandomHorizontalFlip(), # randomly flip and rotate
transforms.RandomRotation(20),
transforms.Resize(size),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406),
(0.229, 0.224, 0.225))])
data_transform_test = transforms.Compose([
transforms.Resize(size),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406),
(0.229, 0.224, 0.225))])
dog_image_dir = '/data/dog_images'
# choose the training and test datasets
train_data = datasets.ImageFolder(trai_dir, transform=data_transform_train)
valid_data = datasets.ImageFolder(val_dir, transform=data_transform_train)
test_data = datasets.ImageFolder(test_dir, transform=data_transform_test)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, shuffle=True, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(valid_data, batch_size=batch_size, shuffle=True, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size, shuffle=True, num_workers=num_workers)
data_loader = dict(train=train_loader, valid=valid_loader, test=test_loader)
# Let's verify that the data was loaded correctly by printing out data stats
print('Num training images: ', len(train_data))
print('Num validation images: ', len(valid_data))
print('Num test images: ', len(test_data))
# Visualize some sample train data
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# denormalize the image
def denormalise(image):
image = np.transpose(image, (1, 2, 0))
mean = [0.485, 0.456, 0.406]
std = [0.229, 0.224, 0.225]
image = (image * std + mean).clip(0, 1)
return image
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 9))
no_vis_imag = 14
for idx in np.arange(no_vis_imag):
ax = fig.add_subplot(2, no_vis_imag/2, idx+1, xticks=[], yticks=[])
plt.imshow(denormalise(images[idx]))
ax.set_title(class_labels[labels[idx]])
```
## Create a CNN using Transfer Learning
VGG16 is used with all the convolutional layers obtained from the pretrained model. The final fully-connected layer is replaced with a new classifier to match the two lung classes.
```
import torchvision.models as models
#import torch.nn.functional as F
import torch.nn as nn
# Load the pretrained model from pytorch
VGG16 = models.vgg16(pretrained=True)
# Modify the last layer
n_inputs = VGG16.classifier[6].in_features
last_layer = nn.Linear(n_inputs, len(class_labels))
VGG16.classifier[6] = last_layer
## Specify Loss Function and Optimizer
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(VGG16.parameters(), lr=0.01)
# check if CUDA is available
use_cuda = torch.cuda.is_available()
if use_cuda:
VGG16 = VGG16.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
#from PIL import ImageFile
#ImageFile.LOAD_TRUNCATED_IMAGES = True
def train(n_epochs, loaders, model, optimizer, criterion, use_cuda, save_path):
"""returns trained model with the possibility to resume analysis and load validation parameters"""
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# initialize variables to monitor training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
# set model to train mode by using dropout to prevent overfitting
model.train()
for batch_idx, (data, target) in enumerate(loaders['train']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
#train_loss += loss.item()*data.size(0)
train_loss = train_loss + ((1 / (batch_idx + 1)) * (loss.data - train_loss))
######################
# validate the model #
######################
model.eval()
for batch_idx, (data, target) in enumerate(loaders['valid']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
## update the average validation loss
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss = valid_loss + ((1 / (batch_idx + 1)) * (loss.data - valid_loss))
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min, valid_loss))
torch.save(model.state_dict(), save_path)
valid_loss_min = valid_loss
# return trained model
return model
# train the model
n_epoch = 8
model = train(n_epoch, data_loader, VGG16, optimizer,
criterion, use_cuda, 'model.pt')
# load the model that got the best validation accuracy
model.load_state_dict(torch.load('model.pt'))
```
## Test the Trained Network
Test the trained model on previously unseen data.
```
def test(loaders, model, criterion, use_cuda):
# monitor test loss and accuracy
test_loss = 0.
correct = 0.
total = 0.
model.eval()
#for batch_idx, (data, target) in enumerate(loaders['test']):
for batch_idx, (data, target) in enumerate(test_loader):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update average test loss
test_loss = test_loss + ((1 / (batch_idx + 1)) * (loss.data - test_loss))
# convert output probabilities to predicted class
pred = output.data.max(1, keepdim=True)[1]
# compare predictions to true label
correct += np.sum(np.squeeze(pred.eq(target.data.view_as(pred))).cpu().numpy())
total += data.size(0)
print('Test Loss: {:.6f}\n'.format(test_loss))
print('\nTest Accuracy: %2d%% (%2d/%2d)' % (
100. * correct / total, correct, total))
# call test function
test(data_loader, model, criterion, use_cuda)
```
## Visualize Sample Test Results
The first 16 images from the test loader are displayed below. The caption colour indicates whether the scan was <span style="color:green">correctly</span> or <span style="color:red">incorrectly</span> classified. The first label indicates the model prediction and the second one in brackets shows the true label.
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
#images = images.numpy() # convert images to numpy for display
images_np = images.cpu().numpy() if not use_cuda else images.numpy()
# move model inputs to cuda, if GPU available
if use_cuda:
images = images.cuda()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.numpy()) if not use_cuda else np.squeeze(preds_tensor.cpu().numpy())
#preds = np.squeeze(preds_tensor.cpu().numpy()) # if not use_cuda else np.squeeze(preds_tensor.cpu().numpy())
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
no_vis_imag = 16
for idx in np.arange(no_vis_imag):
ax = fig.add_subplot(2, no_vis_imag/2, idx+1, xticks=[], yticks=[])
plt.imshow(denormalise(images_np[idx]))
ax.set_title(class_labels[labels[idx]])
ax.set_title("{} ({})".format(class_labels[preds[idx]], class_labels[labels[idx]]),
color=("green" if preds[idx]==labels[idx].item() else "red"))
```
| github_jupyter |
<table width="100%"> <tr>
<td style="background-color:#ffffff;">
<a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="35%" align="left"> </a></td>
<td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
prepared by Özlem Salehi
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
<h2>Resources</h2>
Below is a list of useful resources on quantum computing.
<li><a href="https://www.kuantumturkiye.org" target="_blank"> Kuantum Türkiye Platformu</a>
<li><a href="https://www.duzensiz.org" target="_blank"> Düzensiz</a>
<li><a href="https://arxiv.org/pdf/1804.03719.pdf" target="_blank"> Quantum algorithm implementations for beginners</a>
An article about implementation of the well known quantum algorithms
<li><a href="https://github.com/msramalho/Teach-Me-Quantum" target="_blank">
Course on Quantum Information Science and Quantum Computing </a>
Gitlab page of the winner of the competition Teach me Qiskit by IBM
<li><a href="https://www.quantum-quest.nl/quantumquest.pdf" target="_blank">Quantum Quest</a>
Lecture notes of the 4-week long course for high school students </li>
<li><a href="http://www.michaelnielsen.org/qcqi/" target="_blank">Quantum Computation and Quantum Information</a>
Book by Nielsen and Chuang
<li><a href="https://www.scottaaronson.com/" target="_blank">Scott Aaaronson</a>
Web site of Scott Aaronson. You can find links to his lecture notes and blog.
<li><a href="http://play.quantumgame.io/" target="_blank"> Quantum Game</a>
An online quantum game about quantum optics
<li><a href="https://medium.com/@jonathan_hui/qc-what-is-a-quantum-computer-222edc3a887d" target="_blank"> Quantum Computing Series</a>
A series of notes for quantum computing
<li><a href="https://quantumalgorithmzoo.org/" target="_blank"> Quantum Algorithm Zoo</a>
A list of quantum algorithms and their complexities.
<li><a href="https://quantum.country/qcvc" target="_blank"> Quantum Computing for the Very Curious</a>
A nice website to start learning about quantum computing
<li><a href="https://www.smbc-comics.com/comic/the-talk-3" target="blank">Comics</a>
A comics about quantum computing
| github_jupyter |
TSG097 - Get BDC stateful sets (Kubernetes)
===========================================
Description
-----------
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
def run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False, regex_mask=None):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportability, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
# Display an install HINT, so the user can click on a SOP to install the missing binary
#
if which_binary == None:
print(f"The path used to search for '{cmd_actual[0]}' was:")
print(sys.path)
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
cmd_display = cmd
if regex_mask is not None:
regex = re.compile(regex_mask)
cmd_display = re.sub(regex, '******', cmd)
print(f"START: {cmd_display} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
# Hints for tool retry (on transient fault), known errors and install guide
#
retry_hints = {'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond', ], 'python': [ ], 'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use', 'Login timeout expired (0) (SQLDriverConnect)', 'SSPI Provider: No Kerberos credentials available', ], }
error_hints = {'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb'], ], 'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb'], ], 'azdata': [['Please run \'azdata login\' to first authenticate', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Can\'t open lib \'ODBC Driver 17 for SQL Server', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb'], ['NameError: name \'azdata_login_secret_name\' is not defined', 'SOP013 - Create secret for azdata login (inside cluster)', '../common/sop013-create-secret-for-azdata-login.ipynb'], ['ERROR: No credentials were supplied, or the credentials were unavailable or inaccessible.', 'TSG124 - \'No credentials were supplied\' error from azdata login', '../repair/tsg124-no-credentials-were-supplied.ipynb'], ['Please accept the license terms to use this product through', 'TSG126 - azdata fails with \'accept the license terms to use this product\'', '../repair/tsg126-accept-license-terms.ipynb'], ], }
install_hint = {'kubectl': [ 'SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb' ], 'azdata': [ 'SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb' ], }
print('Common functions defined successfully.')
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the Big Data Cluster use the kubectl command line
interface .
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Run kubectl to display the Stateful sets
```
run(f"kubectl get statefulset -n {namespace} -o wide")
print("Notebook execution is complete.")
```
| github_jupyter |
# Compare similarity search strategies (OLD)
In this notebook I am comparing the following similarity search strategies for finding genomic regions with similar peaks:
1. kNN search in the autoencoded latent space
(1b. kNN search in the UMAP 2D embedding of the autoencoded latent space)
2. kNN search in the Euclidean space of the original data
3. kNN search in the DTW space of the original data
4. kNN search in the 2D UMAP embedding of the original data
5. Template-matching with normalized cross correlation
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import numpy as np
import os
import sys
import warnings
# Ignore warnings as they just pollute the output
warnings.filterwarnings('ignore')
# Enable importing modules from the parent directory
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
module_path = os.path.abspath(os.path.join('../experiments'))
if module_path not in sys.path:
sys.path.append(module_path)
# DNase-seq 2011, hg19
bw = 'data/ENCFF158GBQ.bigWig'
```
## Load and normalize the data
#### Download the data
```
from download import download_encode_file
from pathlib import Path
Path('data').mkdir(parents=True, exist_ok=True)
download_encode_file('ENCFF158GBQ.bigWig')
```
#### Load models, extract genomic windows, and normalize data
```
# Load model and data and normalize data
from keras.models import load_model
from server import bigwig
encoder_2_4kb = load_model('../../examples/encode-gm12878-dnase-seq-2011-encoder-2_4kb.h5')
decoder_2_4kb = load_model('../../examples/encode-gm12878-dnase-seq-2011-decoder-2_4kb.h5')
data_2_4kb = bigwig.chunk(
bw,
2400,
100,
2400 / 2,
['chr22'],
verbose=True,
)
data_2_4kb_b = bigwig.chunk(
bw,
2400,
100,
2400 / 2,
['chr1', 'chr2', 'chr3'],
verbose=True,
)
```
## Encode data
```
from ae.utils import predict
predicted_2_4kb, _, latent_2_4kb = predict(
encoder_2_4kb,
decoder_2_4kb,
data_2_4kb.reshape(data_2_4kb.shape[0], data_2_4kb.shape[1], 1)
)
```
## Helper methods
#### Simple kNN search
```
from scipy.spatial.distance import cdist
def knn(data, target_idx, k, metric='euclidean', sax = None, ignore: int = 0):
"""K nearest neighbors
Find the `k` nearest neighbors of a
"""
target = data[target_idx]
if sax is None:
dist = cdist(data, target.reshape((1, target.size)), metric='euclidean').flatten()
else:
N = data.shape[0]
dist = np.zeros(N)
for i in range(N):
dist[i] = sax.distance_sax(target, data[i])
# Ensure that the target is always first
dist[target_idx] = -1
for i in range(1, ignore + 1):
dist[min(target_idx + i, data.shape[0] - 1)] = -1
dist[max(target_idx - i, 0)] = -1
return np.argsort(dist)[1 + (2 * ignore):k + 1 + (2 * ignore)]
```
#### DTW distance
```
from scipy.spatial.distance import euclidean
from fastdtw import fastdtw
from multiprocessing import Pool
def dtw(data, target_idx: int, n: int, target = None, return_all = False, print_progress: bool = False):
N = data.shape[0]
dist = np.zeros(N)
if target is None:
target = data[target_idx]
p = ((np.arange(4) + 1) * (N // 4)).astype(int)
for i in np.arange(N):
if i in p and print_progress:
print('.', end='', flush=True)
d, _ = fastdtw(data[i], target, dist=euclidean)
dist[i] = d
if return_all:
return dist
return np.argsort(dist)[:n]
def pooled_dtw(data, target_idx: int, n: int, print_progress: bool = False, ignore: int = 0):
target = data[target_idx]
is_not_target = np.arange(data.shape[0]) != target_idx
for i in range(0, ignore + 1):
is_not_target[min(target_idx + i, data.shape[0] - 1)] = False
is_not_target[max(target_idx - i, 0)] = False
with Pool() as pool:
args = [[d, 0, n, target, True, print_progress] for d in np.array_split(data[is_not_target], pool._processes)]
dist = np.concatenate(pool.starmap(dtw, args))
return np.argsort(dist)[:n]
```
#### Normalized cross correlation search
```
from scipy.signal import correlate
def norm(data, zero_norm: bool = False):
mean = np.mean(data) if zero_norm else 0
return (data - mean) / np.std(data)
def norm2d(data, zero_norm: bool = False):
mean = np.mean(data, axis=1).reshape(-1, 1) if zero_norm else np.zeros((data.shape[0], 1))
std = np.std(data, axis=1).reshape(-1, 1)
return (data - mean) / std
def xcorrelation(data, template_idx, n, normalize=False, zero_normalize=False, ignore: int = 0):
unknown = data
template = data[template_idx]
if norm:
unknown = norm2d(unknown, zero_norm=zero_normalize)
template = norm(template, zero_norm=zero_normalize)
xcorr = np.apply_along_axis(lambda m: correlate(m, template, mode='full'), axis=1, arr=unknown)
xcorr[np.where(np.isnan(xcorr))] = 0
max_xcorr = np.nanmax(xcorr, axis=1)
# Ensure that the target is always last
max_xcorr[template_idx] = -1
for i in range(1, ignore + 1):
max_xcorr[min(template_idx + i, data.shape[0] - 1)] = -1
max_xcorr[max(template_idx - i, 0)] = -1
return np.argsort(max_xcorr)[::-1][:n]
```
## Generate UMAP embeddings of the latent space and original data space
```
import umap
umap_embedding_2_4kb = umap.UMAP(
n_neighbors=10,
min_dist=0.01,
metric='l2',
).fit_transform(data_2_4kb)
```
## 2.4kb Search
```
# Some genomic windows with interesting peaks
targets = [20196, 26570, 26576, 26790, 26791, 31670, 25513]
k = 5
knn_ae = []
knn_eq = []
knn_dtw = []
knn_umap = []
top_xcross = []
for target in targets:
print('Search for window #{}'.format(target), end='', flush=True)
knn_ae.append(knn(latent, target, k))
print('.', end='', flush=True)
knn_eq.append(knn(data_test, target, k))
print('.', end='', flush=True)
knn_dtw.append(pooled_dtw(data_test, target, k))
print('.', end='', flush=True)
knn_umap.append(knn(umap_embedding, target, k))
print('.', end='', flush=True)
top_xcross.append(xcorrelation(data_test, target, k, normalize=True, zero_normalize=True))
print('. done!')
```
## 2.4kb visual comparison of search strategies
Compare encoding-based knn similarity search strategies against normalize cross correlation.
| Rows | Description |
|-------|--------------------------------------------------|
| 1 | Search target. Manually picked. |
| 2-6 | k-NN in the CAE-based latent space. |
| 7-11 | k-NN in the Euclide space. |
| 11-16 | k-NN on the dynamic time warping space. |
| 17-21 | k-NN on the UMAP embedded space. |
| 21-26 | Zero-normalized cross correlation. |
```
import matplotlib.pyplot as plt
import numpy as np
N = (k + 1) * 6
T = len(targets)
sz = data_test[0].size
plt.figure(figsize=(5 * T, N))
ymax = 1.0
show_predictions = False
for i, target in enumerate(targets):
ax = plt.subplot(N, T, (i + 1))
ax.set_facecolor("#D9FAFF")
if show_predictions:
plt.bar(np.arange(sz), data_test[target], color='black', alpha=0.33)
plt.bar(np.arange(sz), data_test[target], color='black', alpha=0.33)
else:
plt.bar(np.arange(sz), data_test[target], color='#004754')
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_ae[i]):
plt.subplot(N, T, ((j + 1) * T) + (i + 1))
if show_predictions:
plt.bar(np.arange(sz), predicted[hit], color='green', alpha=0.33)
plt.bar(np.arange(sz), data_test[hit], color='black', alpha=0.33)
else:
plt.bar(np.arange(sz), data_test[hit], color='#d24f00')
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
plt.subplots_adjust(top=0.9)
for j, hit in enumerate(knn_eq[i]):
plt.subplot(N, T, ((j + 6) * T) + (i + 1))
if show_predictions:
plt.bar(np.arange(sz), predicted[hit], color='green', alpha=0.33)
plt.bar(np.arange(sz), data_test[hit], color='black', alpha=0.33)
else:
plt.bar(np.arange(sz), data_test[hit], color='#008ca8')
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_dtw[i]):
plt.subplot(N, T, ((j + 11) * T) + (i + 1))
if show_predictions:
plt.bar(np.arange(sz), predicted[hit], color='red', alpha=0.33)
plt.bar(np.arange(sz), data_test[hit], color='black', alpha=0.33)
else:
plt.bar(np.arange(sz), data_test[hit], color='#93209e')
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_umap[i]):
plt.subplot(N, T, ((j + 16) * T) + (i + 1))
if show_predictions:
plt.bar(np.arange(sz), predicted[hit], color='yellow', alpha=0.33)
plt.bar(np.arange(sz), data_test[hit], color='black')
else:
plt.bar(np.arange(sz), data_test[hit], color='#209e4e')
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(top_xcross[i]):
plt.subplot(N, T, ((j + 21) * T) + (i + 1))
plt.bar(np.arange(sz), data_test[hit], color='#cc9900')
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/intel-analytics/analytics-zoo/blob/master/docs/docs/colab-notebook/chronos/chronos_autots_nyc_taxi.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

---
##### Copyright 2018 Analytics Zoo Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
```
## **Environment Preparation**
**Install Analytics Zoo**
You can install the latest pre-release version with chronos support using `pip install --pre --upgrade analytics-zoo[automl]`.
```
# Install latest pre-release version of Analytics Zoo
# Installing Analytics Zoo from pip will automatically install pyspark, bigdl, and their dependencies.
!pip install --pre --upgrade analytics-zoo[automl]
exit() # restart the runtime to refresh installed pkg
```
### **Step 0: Download & prepare dataset**
We used NYC taxi passengers dataset in [Numenta Anomaly Benchmark (NAB)](https://github.com/numenta/NAB) for demo, which contains 10320 records, each indicating the total number of taxi passengers in NYC at a corresonponding time spot.
```
# download the dataset
!wget https://raw.githubusercontent.com/numenta/NAB/v1.0/data/realKnownCause/nyc_taxi.csv
# load the dataset. The downloaded dataframe contains two columns, "timestamp" and "value".
import pandas as pd
df = pd.read_csv("nyc_taxi.csv", parse_dates=["timestamp"])
```
## **Time series forecasting using Chronos Forecaster**
### Forecaster Step1. Data transformation and feature engineering using Chronos TSDataset
[TSDataset](https://analytics-zoo.readthedocs.io/en/latest/doc/PythonAPI/Chronos/tsdataset.html) is our abstract of time series dataset for data transformation and feature engineering. Here we use it to preprocess the data.
```
from zoo.chronos.data import TSDataset
from sklearn.preprocessing import StandardScaler
```
Initialize train, valid and test tsdataset from raw pandas dataframe.
```
tsdata_train, tsdata_valid, tsdata_test = TSDataset.from_pandas(df, dt_col="timestamp", target_col="value",
with_split=True, val_ratio=0.1, test_ratio=0.1)
```
Preprocess the datasets. Here we perform:
- deduplicate: remove those identical data records
- impute: fill the missing values
- gen_dt_feature: generate feature from datetime (e.g. month, day...)
- scale: scale each feature to standard distribution.
- roll: sample the data with sliding window.
For forecasting task, we will look back 3 hours' historical data (6 records) and predict the value of next 30 miniutes (1 records).
We perform the same transformation processes on train, valid and test set.
```
lookback, horizon = 6, 1
scaler = StandardScaler()
for tsdata in [tsdata_train, tsdata_valid, tsdata_test]:
tsdata.deduplicate()\
.impute()\
.gen_dt_feature()\
.scale(scaler, fit=(tsdata is tsdata_train))\
.roll(lookback=lookback, horizon=horizon)
```
### Forecaster Step 2: Time series forecasting using Chronos Forecaster
After preprocessing the datasets. We can use [Chronos Forecaster](https://analytics-zoo.readthedocs.io/en/latest/doc/PythonAPI/Chronos/forecasters.html) to handle the forecasting tasks.
Transform TSDataset to sampled numpy ndarray and feed them to forecaster.
```
from zoo.chronos.forecast.tcn_forecaster import TCNForecaster
x, y = tsdata_train.to_numpy()
# x.shape = (num of sample, lookback, num of input feature)
# y.shape = (num of sample, horizon, num of output feature)
forecaster = TCNForecaster(past_seq_len=lookback, # number of steps to look back
future_seq_len=horizon, # number of steps to predict
input_feature_num=x.shape[-1], # number of feature to use
output_feature_num=y.shape[-1], # number of feature to predict
seed=0)
res = forecaster.fit((x, y), epochs=3)
```
### Forecaster Step 3: Further deployment with fitted forecaster
Use fitted forecaster to predict test data and plot the result
```
x_test, y_test = tsdata_test.to_numpy()
pred = forecaster.predict(x_test)
pred_unscale, groundtruth_unscale = tsdata_test.unscale_numpy(pred), tsdata_test.unscale_numpy(y_test)
import matplotlib.pyplot as plt
plt.figure(figsize=(24,6))
plt.plot(pred_unscale[:,:,0])
plt.plot(groundtruth_unscale[:,:,0])
plt.legend(["prediction", "ground truth"])
```
Save & restore the forecaster.
```
forecaster.save("nyc_taxi.fxt")
forecaster.load("nyc_taxi.fxt")
```
| github_jupyter |
```
test_index = 0
```
#### testing
```
from load_data import *
# load_data()
```
## Loading the data
```
from load_data import *
X_train,X_test,y_train,y_test = load_data()
len(X_train),len(y_train)
len(X_test),len(y_test)
```
## Test Modelling
```
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
class Test_Model(nn.Module):
def __init__(self) -> None:
super().__init__()
self.c1 = nn.Conv2d(1,64,5)
self.c2 = nn.Conv2d(64,128,5)
self.c3 = nn.Conv2d(128,256,5)
self.fc4 = nn.Linear(256*10*10,256)
self.fc6 = nn.Linear(256,128)
self.fc5 = nn.Linear(128,4)
def forward(self,X):
preds = F.max_pool2d(F.relu(self.c1(X)),(2,2))
preds = F.max_pool2d(F.relu(self.c2(preds)),(2,2))
preds = F.max_pool2d(F.relu(self.c3(preds)),(2,2))
# print(preds.shape)
preds = preds.view(-1,256*10*10)
preds = F.relu(self.fc4(preds))
preds = F.relu(self.fc6(preds))
preds = self.fc5(preds)
return preds
device = torch.device('cuda')
BATCH_SIZE = 32
IMG_SIZE = 112
model = Test_Model().to(device)
optimizer = optim.SGD(model.parameters(),lr=0.1)
criterion = nn.CrossEntropyLoss()
EPOCHS = 125
from tqdm import tqdm
PROJECT_NAME = 'Weather-Clf'
import wandb
# test_index += 1
# wandb.init(project=PROJECT_NAME,name=f'test')
# for _ in tqdm(range(EPOCHS)):
# for i in range(0,len(X_train),BATCH_SIZE):
# X_batch = X_train[i:i+BATCH_SIZE].view(-1,1,112,112).to(device)
# y_batch = y_train[i:i+BATCH_SIZE].to(device)
# model.to(device)
# preds = model(X_batch.float())
# preds.to(device)
# loss = criterion(preds,torch.tensor(y_batch,dtype=torch.long))
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':loss.item()})
# wandb.finish()
# for index in range(10):
# print(torch.argmax(preds[index]))
# print(y_batch[index])
# print('\n')
class Test_Model(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1,16,5)
self.conv2 = nn.Conv2d(16,32,5)
self.conv3 = nn.Conv2d(32,64,5)
self.fc1 = nn.Linear(64*10*10,16)
self.fc2 = nn.Linear(16,32)
self.fc3 = nn.Linear(32,64)
self.fc4 = nn.Linear(64,32)
self.fc5 = nn.Linear(32,6)
def forward(self,X):
preds = F.max_pool2d(F.relu(self.conv1(X)),(2,2))
preds = F.max_pool2d(F.relu(self.conv2(preds)),(2,2))
preds = F.max_pool2d(F.relu(self.conv3(preds)),(2,2))
# print(preds.shape)
preds = preds.view(-1,64*10*10)
preds = F.relu(self.fc1(preds))
preds = F.relu(self.fc2(preds))
preds = F.relu(self.fc3(preds))
preds = F.relu(self.fc4(preds))
preds = F.relu(self.fc5(preds))
return preds
model = Test_Model().to(device)
optimizer = optim.SGD(model.parameters(),lr=0.1)
criterion = nn.CrossEntropyLoss()
# test_index += 1
# wandb.init(project=PROJECT_NAME,name=f'test-{test_index}')
# for _ in tqdm(range(EPOCHS)):
# for i in range(0,len(X_train),BATCH_SIZE):
# X_batch = X_train[i:i+BATCH_SIZE].view(-1,1,112,112).to(device)
# y_batch = y_train[i:i+BATCH_SIZE].to(device)
# model.to(device)
# preds = model(X_batch.float())
# preds.to(device)
# loss = criterion(preds,torch.tensor(y_batch,dtype=torch.long))
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':loss.item()})
# wandb.finish()
```
## Modelling
```
class Test_Model(nn.Module):
def __init__(self,conv1_output=16,conv2_output=32,conv3_output=64,fc1_output=16,fc2_output=32,fc3_output=64,activation=F.relu):
super().__init__()
self.conv3_output = conv3_output
self.conv1 = nn.Conv2d(1,conv1_output,5)
self.conv2 = nn.Conv2d(conv1_output,conv2_output,5)
self.conv3 = nn.Conv2d(conv2_output,conv3_output,5)
self.fc1 = nn.Linear(conv3_output*10*10,fc1_output)
self.fc2 = nn.Linear(fc1_output,fc2_output)
self.fc3 = nn.Linear(fc2_output,fc3_output)
self.fc4 = nn.Linear(fc3_output,fc2_output)
self.fc5 = nn.Linear(fc2_output,6)
self.activation = activation
def forward(self,X):
preds = F.max_pool2d(self.activation(self.conv1(X)),(2,2))
preds = F.max_pool2d(self.activation(self.conv2(preds)),(2,2))
preds = F.max_pool2d(self.activation(self.conv3(preds)),(2,2))
# print(preds.shape)
preds = preds.view(-1,self.conv3_output*10*10)
preds = self.activation(self.fc1(preds))
preds = self.activation(self.fc2(preds))
preds = self.activation(self.fc3(preds))
preds = self.activation(self.fc4(preds))
preds = self.activation(self.fc5(preds))
return preds
# conv1_output = 32
# conv2_output = 8
# conv3_output = 64
# fc1_output = 512
# fc2_output = 512
# fc3_output = 256
# activation
# optimizer
# loss
# lr
# num of epochs
def get_loss(criterion,y,model,X):
model.to('cpu')
preds = model(X.view(-1,1,112,112).to('cpu').float())
preds.to('cpu')
loss = criterion(preds,torch.tensor(y,dtype=torch.long).to('cpu'))
loss.backward()
return loss.item()
def test(net,X,y):
device = 'cpu'
net.to(device)
correct = 0
total = 0
net.eval()
with torch.no_grad():
for i in range(len(X)):
real_class = torch.argmax(y[i]).to(device)
net_out = net(X[i].view(-1,1,112,112).to(device).float())
net_out = net_out[0]
predictied_class = torch.argmax(net_out)
if predictied_class == real_class:
correct += 1
total += 1
net.train()
net.to('cuda')
return round(correct/total,3)
EPOCHS = 3
optimizers = [torch.optim.Adam,torch.optim.AdamW,torch.optim.Adamax,torch.optim.SGD]
for optimizer in optimizers:
model = Test_Model(conv1_output=32,conv2_output=8,conv3_output=64,fc1_output=512,fc3_output=256,fc2_output=512,activation=nn.CELU()).to(device)
optimizer = optimizer(model.parameters(),lr=0.1)
criterion = nn.CrossEntropyLoss()
wandb.init(project=PROJECT_NAME,name=f'optimizer-{optimizer}')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,1,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds.to(device)
loss = criterion(preds,torch.tensor(y_batch,dtype=torch.long))
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':get_loss(criterion,y_train,model,X_train),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test,y_test),'val_loss':get_loss(criterion,y_test,model,X_test)})
for index in range(10):
print(torch.argmax(preds[index]))
print(y_batch[index])
print('\n')
wandb.finish()
# fc2_outputs = [64,128,256,512]
# for fc2_output in fc2_outputs:
# wandb.init(project=PROJECT_NAME,name=f'fc2_output-{fc2_output}')
# model = Test_Model(conv1_output=32,conv2_output=8,conv3_output=64,fc1_output=512,fc2_output=fc2_output).to(device)
# optimizer = optim.SGD(model.parameters(),lr=0.1)
# criterion = nn.CrossEntropyLoss()
# for _ in tqdm(range(EPOCHS)):
# for i in range(0,len(X_train),BATCH_SIZE):
# X_batch = X_train[i:i+BATCH_SIZE].view(-1,1,112,112).to(device)
# y_batch = y_train[i:i+BATCH_SIZE].to(device)
# model.to(device)
# preds = model(X_batch.float())
# preds.to(device)
# loss = criterion(preds,torch.tensor(y_batch,dtype=torch.long))
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':get_loss(criterion,y_train,model,X_train),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test,y_test),'val_loss':get_loss(criterion,y_test,model,X_test)})
# for index in range(10):
# print(torch.argmax(preds[index]))
# print(y_batch[index])
# print('\n')
# wandb.finish()
```
| github_jupyter |
## TVP-VAR, MCMC, and sparse simulation smoothing
```
%matplotlib inline
from importlib import reload
import numpy as np
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
from scipy.stats import invwishart, invgamma
# Get the macro dataset
dta = sm.datasets.macrodata.load_pandas().data
dta.index = pd.date_range('1959Q1', '2009Q3', freq='QS')
```
### Background
Bayesian analysis of linear Gaussian state space models via Markov chain Monte Carlo (MCMC) methods has become both commonplace and relatively straightforward in recent years, due especially to advances in sampling from the joint posterior of the unobserved state vector conditional on the data and model parameters (see especially Carter and Kohn (1994), de Jong and Shephard (1995), and Durbin and Koopman (2002)). This is particularly useful for Gibbs sampling MCMC approaches.
While these procedures make use of the forward/backward application of the recursive Kalman filter and smoother, another recent line of research takes a different approach and constructs the posterior joint distribution of the entire vector of states at once - see in particular Chan and Jeliazkov (2009) for an econometric time series treatment and McCausland et al. (2011) for a more general survey. In particular, the posterior mean and precision matrix are constructed explicitly, with the latter a sparse band matrix. Advantage is then taken of efficient algorithms for Cholesky factorization of sparse band matrices; this reduces memory costs and can improve performance. Following McCausland et al. (2011), we refer to this method as the "Cholesky Factor Algorithm" (CFA) approach.
The CFA-based simulation smoother has some advantages and some drawbacks compared to that based on the more typical Kalman filter and smoother (KFS).
**Advantages of CFA**:
- Derivation of the joint posterior distribution is relatively straightforward and easy to understand.
- In some cases can be both faster and less memory-intensive than the KFS approach
- In the Appendix at the end of this notebook, we briefly discuss the performance of the two simulation smoothers for the TVP-VAR model. In summary: simple tests on a single machine suggest that for the TVP-VAR model, the CFA and KFS implementations in Statsmodels have about the same runtimes, while both implementations are about twice as fast as the replication code, written in Matlab, provided by Chan and Jeliakov (2009).
**Drawbacks of CFA**:
The main drawback is that this method has not (at least so far) reached the generality of the KFS approach. For example:
- It can not be used with models that have reduced-rank error terms in the observation or state equations.
- One implication of this is that the typical state space model trick of including identities in the state equation to accomodate, for example, higher-order lags in autoregressive models is not applicable. These models can still be handled by the CFA approach, but at the cost of requiring a slightly different implementation for each lag that is included.
- As an example, standard ways of representing ARMA and VARMA processes in state space form do include identities in the observation and/or state equations, and so the basic formulas presented in Chan and Jeliazkov (2009) do not apply immediately to these models.
- Less flexibility is available in the state initialization / prior.
### Implementation in Statsmodels
A CFA simulation smoother along the lines of the basic formulas presented in Chan and Jeliazkov (2009) has been implemented in Statsmodels.
**Notes**:
- Therefore, the CFA simulation smoother in Statsmodels so-far only supports the case that the state transition is truly a first-order Markov process (i.e. it does not support a p-th order Markov process that has been stacked using identities into a first-order process).
- By contrast, the KFS smoother in Statsmodels is fully general any can be used for any state space model, including those with stacked p-th order Markov processes or other identities in the observation and state equations.
Either a KFS or the CFA simulation smoothers can be constructed from a state space model using the `simulation_smoother` method. To show the basic idea, we first consider a simple example.
#### Local level model
A local level model decomposes an observed series $y_t$ into a persistent trend $\mu_t$ and a transitory error component
$$
\begin{aligned}
y_t & = \mu_t + \varepsilon_t, \qquad \varepsilon_t \sim N(0, \sigma_\text{irregular}^2) \\
\mu_t & = \mu_{t-1} + \eta_t, \quad ~ \eta_t \sim N(0, \sigma_\text{level}^2)
\end{aligned}
$$
This model satisfies the requirements of the CFA simulation smoother because both the observation error term $\varepsilon_t$ and the state innovation term $\eta_t$ are non-degenerate - that is, their covariance matrices are full rank.
We apply this model to inflation, and consider simulating draws from the posterior of the joint state vector. That is, we are interested in sampling from
$$p(\mu^t \mid y^t, \sigma_\text{irregular}^2, \sigma_\text{level}^2)$$
where we define $\mu^t \equiv (\mu_1, \dots, \mu_T)'$ and $y^t \equiv (y_1, \dots, y_T)'$.
In Statsmodels, the local level model falls into the more general class of "unobserved components" models, and can be contructed as follows:
```
# Construct a local level model for inflation
mod = sm.tsa.UnobservedComponents(dta.infl, 'llevel')
# Fit the model's parameters (sigma2_varepsilon and sigma2_eta)
# via maximum likelihood
res = mod.fit()
print(res.params)
# Create simulation smoother objects
sim_kfs = mod.simulation_smoother() # default method is KFS
sim_cfa = mod.simulation_smoother(method='cfa') # can specify CFA method
```
The simulation smoother objects `sim_kfs` and `sim_cfa` have `simulate` methods that perform simulation smoothing. Each time that `simulate` is called, the `simulated_state` attribute will be re-populated with a new simulated draw from the posterior.
Below, we construct 20 simulated paths for the trend, using the KFS and CFA approaches, where the simulation is at the maximum likelihood parameter estimates.
```
nsimulations = 20
simulated_state_kfs = pd.DataFrame(
np.zeros((mod.nobs, nsimulations)), index=dta.index)
simulated_state_cfa = pd.DataFrame(
np.zeros((mod.nobs, nsimulations)), index=dta.index)
for i in range(nsimulations):
# Apply KFS simulation smoothing
sim_kfs.simulate()
# Save the KFS simulated state
simulated_state_kfs.iloc[:, i] = sim_kfs.simulated_state[0]
# Apply CFA simulation smoothing
sim_cfa.simulate()
# Save the CFA simulated state
simulated_state_cfa.iloc[:, i] = sim_cfa.simulated_state[0]
```
Plotting the observed data and the simulations created using each method below, it is not too hard to see that these two methods are doing the same thing.
```
# Plot the inflation data along with simulated trends
fig, axes = plt.subplots(2, figsize=(15, 6))
# Plot data and KFS simulations
dta.infl.plot(ax=axes[0], color='k')
axes[0].set_title('Simulations based on KFS approach, MLE parameters')
simulated_state_kfs.plot(ax=axes[0], color='C0', alpha=0.25, legend=False)
# Plot data and CFA simulations
dta.infl.plot(ax=axes[1], color='k')
axes[1].set_title('Simulations based on CFA approach, MLE parameters')
simulated_state_cfa.plot(ax=axes[1], color='C0', alpha=0.25, legend=False)
# Add a legend, clean up layout
handles, labels = axes[0].get_legend_handles_labels()
axes[0].legend(handles[:2], ['Data', 'Simulated state'])
fig.tight_layout();
```
#### Updating the model's parameters
The simulation smoothers are tied to the model instance, here the variable `mod`. Whenever the model instance is updated with new parameters, the simulation smoothers will take those new parameters into account in future calls to the `simulate` method.
This is convenient for MCMC algorithms, which repeatedly (a) update the model's parameters, (b) draw a sample of the state vector, and then (c) draw new values for the model's parameters.
Here we will change the model to a different parameterization that yields a smoother trend, and show how the simulated values change (for brevity we only show the simulations from the KFS approach, but simulations from the CFA approach would be the same).
```
fig, ax = plt.subplots(figsize=(15, 3))
# Update the model's parameterization to one that attributes more
# variation in inflation to the observation error and so has less
# variation in the trend component
mod.update([4, 0.05])
# Plot simulations
for i in range(nsimulations):
sim_kfs.simulate()
ax.plot(dta.index, sim_kfs.simulated_state[0],
color='C0', alpha=0.25, label='Simulated state')
# Plot data
dta.infl.plot(ax=ax, color='k', label='Data', zorder=-1)
# Add title, legend, clean up layout
ax.set_title('Simulations with alternative parameterization yielding a smoother trend')
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles[-2:], labels[-2:])
fig.tight_layout();
```
### Application: Bayesian analysis of a TVP-VAR model by MCMC
One of the applications that Chan and Jeliazkov (2009) consider is the time-varying parameters vector autoregression (TVP-VAR) model, estimated with Bayesian Gibb sampling (MCMC) methods. They apply this to model the co-movements in four macroeconomic time series:
- Real GDP growth
- Inflation
- Unemployment rate
- Short-term interest rates
We will replicate their example, using a very similar dataset that is included in Statsmodels.
```
# Subset to the four variables of interest
y = dta[['realgdp', 'cpi', 'unemp', 'tbilrate']].copy()
y.columns = ['gdp', 'inf', 'unemp', 'int']
# Convert to real GDP growth and CPI inflation rates
y[['gdp', 'inf']] = np.log(y[['gdp', 'inf']]).diff() * 100
y = y.iloc[1:]
fig, ax = plt.subplots(figsize=(15, 5))
y.plot(ax=ax)
ax.set_title('Evolution of macroeconomic variables included in TVP-VAR exercise');
```
#### TVP-VAR model
**Note**: this section is based on Chan and Jeliazkov (2009) section 3.1, which can be consulted for additional details.
The usual (time-invariant) VAR(1) model is typically written:
$$
\begin{aligned}
y_t & = \mu + \Phi y_{t-1} + \varepsilon_t, \qquad \varepsilon_t \sim N(0, H)
\end{aligned}
$$
where $y_t$ is a $p \times 1$ vector of variables observed at time $t$ and $H$ is a covariance matrix.
The TVP-VAR(1) model generalizes this to allow the coefficients to vary over time according. Stacking all the parameters into a vector according to $\alpha_t = \text{vec}([\mu_t : \Phi_t])$, where $\text{vec}$ denotes the operation that stacks columns of a matrix into a vector, we model their evolution over time according to:
$$\alpha_{i,t+1} = \alpha_{i, t} + \eta_{i,t}, \qquad \eta_{i, t} \sim N(0, \sigma_i^2)$$
In other words, each parameter evolves independently according to a random walk.
Note that there are $p$ coefficients in $\mu_t$ and $p^2$ coefficients in $\Phi_t$, so the full state vector $\alpha$ is shaped $p * (p + 1) \times 1$.
Putting the TVP-VAR(1) model into state-space form is relatively straightforward, and in fact we just have to re-write the observation equation into SUR form:
$$
\begin{aligned}
y_t & = Z_t \alpha_t + \varepsilon_t, \qquad \varepsilon_t \sim N(0, H) \\
\alpha_{t+1} & = \alpha_t + \eta_t, \qquad \eta_t \sim N(0, \text{diag}(\{\sigma_i^2\}))
\end{aligned}
$$
where
$$
Z_t = \begin{bmatrix}
1 & y_{t-1}' & 0 & \dots & & 0 \\
0 & 0 & 1 & y_{t-1}' & & 0 \\
\vdots & & & \ddots & \ddots & \vdots \\
0 & 0 & 0 & 0 & 1 & y_{t-1}' \\
\end{bmatrix}
$$
As long as $H$ is full rank and each of the variances $\sigma_i^2$ is non-zero, the model satisfies the requirements of the CFA simulation smoother.
We also need to specify the initialization / prior for the initial state, $\alpha_1$. Here we will follow Chan and Jeliazkov (2009) in using $\alpha_1 \sim N(0, 5 I)$, although we could also model it as diffuse.
Aside from the time-varying coefficients $\alpha_t$, the other parameters that we will need to estimate are terms in the covariance matrix $H$ and the random walk variances $\sigma_i^2$.
#### TVP-VAR model in Statsmodels
Constructing this model programatically in Statsmodels is also relatively straightforward, since there are basically four steps:
1. Create a new `TVPVAR` class as a subclass of `sm.tsa.statespace.MLEModel`
2. Fill in the fixed values of the state space system matrices
3. Specify the initialization of $\alpha_1$
4. Create a method for updating the state space system matrices with new values of the covariance matrix $H$ and the random walk variances $\sigma_i^2$.
To do this, first note that the general state space representation used by Statsmodels is:
$$
\begin{aligned}
y_t & = d_t + Z_t \alpha_t + \varepsilon_t, \qquad \varepsilon_t \sim N(0, H_t) \\
\alpha_{t+1} & = c_t + T_t \alpha_t + R_t \eta_t, \qquad \eta_t \sim N(0, Q_t) \\
\end{aligned}
$$
Then the TVP-VAR(1) model implies the following specializations:
- The intercept terms are zero, i.e. $c_t = d_t = 0$
- The design matrix $Z_t$ is time-varying but its valeus are fixed as described above (i.e. its values contain ones and lags of $y_t$)
- The observation covariance matrix is not time-varying, i.e. $H_t = H_{t+1} = H$
- The transition matrix is not time-varying and is equal to the identity matrix, i.e. $T_t = T_{t+1} = I$
- The selection matrix $R_t$ is not time-varying and is also equal to the identity matrix, i.e. $R_t = R_{t+1} = I$
- The state covariance matrix $Q_t$ is not time-varying and is diagonal, i.e. $Q_t = Q_{t+1} = \text{diag}(\{\sigma_i^2\})$
```
# 1. Create a new TVPVAR class as a subclass of sm.tsa.statespace.MLEModel
class TVPVAR(sm.tsa.statespace.MLEModel):
# Steps 2-3 are best done in the class "constructor", i.e. the __init__ method
def __init__(self, y):
# Create a matrix with [y_t' : y_{t-1}'] for t = 2, ..., T
augmented = sm.tsa.lagmat(y, 1, trim='both', original='in', use_pandas=True)
# Separate into y_t and z_t = [1 : y_{t-1}']
p = y.shape[1]
y_t = augmented.iloc[:, :p]
z_t = sm.add_constant(augmented.iloc[:, p:])
# Recall that the length of the state vector is p * (p + 1)
k_states = p * (p + 1)
super().__init__(y_t, exog=z_t, k_states=k_states)
# Note that the state space system matrices default to contain zeros,
# so we don't need to explicitly set c_t = d_t = 0.
# Construct the design matrix Z_t
# Notes:
# -> self.k_endog = p is the dimension of the observed vector
# -> self.k_states = p * (p + 1) is the dimension of the observed vector
# -> self.nobs = T is the number of observations in y_t
self['design'] = np.zeros((self.k_endog, self.k_states, self.nobs))
for i in range(self.k_endog):
start = i * (self.k_endog + 1)
end = start + self.k_endog + 1
self['design', i, start:end, :] = z_t.T
# Construct the transition matrix T = I
self['transition'] = np.eye(k_states)
# Construct the selection matrix R = I
self['selection'] = np.eye(k_states)
# Step 3: Initialize the state vector as alpha_1 ~ N(0, 5I)
self.ssm.initialize('known', stationary_cov=5 * np.eye(self.k_states))
# Step 4. Create a method that we can call to update H and Q
def update_variances(self, obs_cov, state_cov_diag):
self['obs_cov'] = obs_cov
self['state_cov'] = np.diag(state_cov_diag)
# Finally, it can be convenient to define human-readable names for
# each element of the state vector. These will be available in output
@property
def state_names(self):
state_names = np.empty((self.k_endog, self.k_endog + 1), dtype=object)
for i in range(self.k_endog):
endog_name = self.endog_names[i]
state_names[i] = (
['intercept.%s' % endog_name] +
['L1.%s->%s' % (other_name, endog_name) for other_name in self.endog_names])
return state_names.ravel().tolist()
```
The above class defined the state space model for any given dataset. Now we need to create a specific instance of it with the dataset that we created earlier containing real GDP growth, inflation, unemployment, and interest rates.
```
# Create an instance of our TVPVAR class with our observed dataset y
mod = TVPVAR(y)
```
#### Preliminary investigation with ad-hoc parameters in H, Q
In our analysis below, we will need to begin our MCMC iterations with some initial parameterization. Following Chan and Jeliazkov (2009) we will set $H$ to be the sample covariance matrix of our dataset, and we will set $\sigma_i^2 = 0.01$ for each $i$.
Before discussing the MCMC scheme that will allow us to make inferences about the model, first we can consider the output of the model when simply plugging in these initial parameters. To fill in these parameters, we use the `update_variances` method that we defined earlier and then perform Kalman filtering and smoothing conditional on those parameters.
**Warning: This exercise is just by way of explanation - we must wait for the output of the MCMC exercise to study the actual implications of the model in a meaningful way**.
```
initial_obs_cov = np.cov(y.T)
initial_state_cov_diag = [0.01] * mod.k_states
# Update H and Q
mod.update_variances(initial_obs_cov, initial_state_cov_diag)
# Perform Kalman filtering and smoothing
# (the [] is just an empty list that in some models might contain
# additional parameters. Here, we don't have any additional parameters
# so we just pass an empty list)
initial_res = mod.smooth([])
```
The `initial_res` variable contains the output of Kalman filtering and smoothing, conditional on those initial parameters. In particular, we may be interested in the "smoothed states", which are $E[\alpha_t \mid y^t, H, \{\sigma_i^2\}]$.
First, lets create a function that graphs the coefficients over time, separated into the equations for equation of the observed variables.
```
def plot_coefficients_by_equation(states):
fig, axes = plt.subplots(2, 2, figsize=(15, 8))
# The way we defined Z_t implies that the first 5 elements of the
# state vector correspond to the first variable in y_t, which is GDP growth
ax = axes[0, 0]
states.iloc[:, :5].plot(ax=ax)
ax.set_title('GDP growth')
ax.legend()
# The next 5 elements correspond to inflation
ax = axes[0, 1]
states.iloc[:, 5:10].plot(ax=ax)
ax.set_title('Inflation rate')
ax.legend();
# The next 5 elements correspond to unemployment
ax = axes[1, 0]
states.iloc[:, 10:15].plot(ax=ax)
ax.set_title('Unemployment equation')
ax.legend()
# The last 5 elements correspond to the interest rate
ax = axes[1, 1]
states.iloc[:, 15:20].plot(ax=ax)
ax.set_title('Interest rate equation')
ax.legend();
return ax
```
Now, we are interested in the smoothed states, which are available in the `states.smoothed` attribute out our results object `initial_res`.
As the graph below shows, the initial parameterization implies substantial time-variation in some of the coefficients.
```
# Here, for illustration purposes only, we plot the time-varying
# coefficients conditional on an ad-hoc parameterization
# Recall that `initial_res` contains the Kalman filtering and smoothing,
# and the `states.smoothed` attribute contains the smoothed states
plot_coefficients_by_equation(initial_res.states.smoothed);
```
#### Bayesian estimation via MCMC
We will now implement the Gibbs sampler scheme described in Chan and Jeliazkov (2009), Algorithm 2.
We use the following (conditionally conjugate) priors:
$$
\begin{aligned}
H & \sim \mathcal{IW}(\nu_1^0, S_1^0) \\
\sigma_i^2 & \sim \mathcal{IG} \left ( \frac{\nu_{i2}^0}{2}, \frac{S_{i2}^0}{2} \right )
\end{aligned}
$$
where $\mathcal{IW}$ denotes the inverse-Wishart distribution and $\mathcal{IG}$ denotes the inverse-Gamma distribution. We set the prior hyperparameters as:
$$
\begin{aligned}
v_1^0 = T + 3, & \quad S_1^0 = I \\
v_{i2}^0 = 6, & \quad S_{i2}^0 = 0.01 \qquad \text{for each} ~ i\\
\end{aligned}
$$
```
# Prior hyperparameters
# Prior for obs. cov. is inverse-Wishart(v_1^0=k + 3, S10=I)
v10 = mod.k_endog + 3
S10 = np.eye(mod.k_endog)
# Prior for state cov. variances is inverse-Gamma(v_{i2}^0 / 2 = 3, S+{i2}^0 / 2 = 0.005)
vi20 = 6
Si20 = 0.01
```
Before running the MCMC iterations, there are a couple of practical steps:
1. Create arrays to store the draws of our state vector, observation covariance matrix, and state error variances.
2. Put the initial values for H and Q (described above) into the storage vectors
3. Construct the simulation smoother object associated with our `TVPVAR` instance to make draws of the state vector
```
# Gibbs sampler setup
niter = 11000
nburn = 1000
# 1. Create storage arrays
store_states = np.zeros((niter + 1, mod.nobs, mod.k_states))
store_obs_cov = np.zeros((niter + 1, mod.k_endog, mod.k_endog))
store_state_cov = np.zeros((niter + 1, mod.k_states))
# 2. Put in the initial values
store_obs_cov[0] = initial_obs_cov
store_state_cov[0] = initial_state_cov_diag
mod.update_variances(store_obs_cov[0], store_state_cov[0])
# 3. Construct posterior samplers
sim = mod.simulation_smoother(method='cfa')
```
As before, we could have used either the simulation smoother based on the Kalman filter and smoother or that based on the Cholesky Factor Algorithm.
```
for i in range(niter):
mod.update_variances(store_obs_cov[i], store_state_cov[i])
sim.simulate()
# 1. Sample states
store_states[i + 1] = sim.simulated_state.T
# 2. Simulate obs cov
fitted = np.matmul(mod['design'].transpose(2, 0, 1), store_states[i + 1][..., None])[..., 0]
resid = mod.endog - fitted
store_obs_cov[i + 1] = invwishart.rvs(v10 + mod.nobs, S10 + resid.T @ resid)
# 3. Simulate state cov variances
resid = store_states[i + 1, 1:] - store_states[i + 1, :-1]
sse = np.sum(resid**2, axis=0)
for j in range(mod.k_states):
rv = invgamma.rvs((vi20 + mod.nobs - 1) / 2, scale=(Si20 + sse[j]) / 2)
store_state_cov[i + 1, j] = rv
```
After removing a number of initial draws, the remaining draws from the posterior allow us to conduct inference. Below, we plot the posterior mean of the time-varying regression coefficients.
(**Note**: these plots are different from those in Figure 1 of the published version of Chan and Jeliakov (2009), but they are very similar to those produced by the Matlab replication code available at http://joshuachan.org/code/code_TVPVAR.html)
```
# Collect the posterior means of each time-varying coefficient
states_posterior_mean = pd.DataFrame(
np.mean(store_states[nburn + 1:], axis=0),
index=mod._index, columns=mod.state_names)
# Plot these means over time
plot_coefficients_by_equation(states_posterior_mean);
```
Python also has a number of libraries to assist with exploring Bayesian models. Here we'll just use the [arviz](https://arviz-devs.github.io/arviz/index.html) package to explore the credible intervals of each of the covariance and variance parameters, although it makes available a much wider set of tools for analysis.
```
import arviz as az
# Collect the observation error covariance parameters
az_obs_cov = az.convert_to_inference_data({
('Var[%s]' % mod.endog_names[i] if i == j else
'Cov[%s, %s]' % (mod.endog_names[i], mod.endog_names[j])):
store_obs_cov[nburn + 1:, i, j]
for i in range(mod.k_endog) for j in range(i, mod.k_endog)})
# Plot the credible intervals
az.plot_forest(az_obs_cov, figsize=(8, 7));
# Collect the state innovation variance parameters
az_state_cov = az.convert_to_inference_data({
r'$\sigma^2$[%s]' % mod.state_names[i]: store_state_cov[nburn + 1:, i]
for i in range(mod.k_states)})
# Plot the credible intervals
az.plot_forest(az_state_cov, figsize=(8, 7));
```
### Appendix: performance
Finally, we run a few simple tests to compare the performance of the KFS and CFA simulation smoothers by using the `%timeit` Jupytor notebook magic.
One caveat is that the KFS simulation smoother can produce a variety of output beyond just simulations of the posterior state vector, and these additional computations could bias the results. To make the results comparable, we will tell the KFS simulation smoother to only compute simulations of the state by using the `simulation_output` argument.
```
from statsmodels.tsa.statespace.simulation_smoother import SIMULATION_STATE
sim_cfa = mod.simulation_smoother(method='cfa')
sim_kfs = mod.simulation_smoother(simulation_output=SIMULATION_STATE)
```
Then we can use the following code to perform a basic timing exercise:
```python
%timeit -n 10000 -r 3 sim_cfa.simulate()
%timeit -n 10000 -r 3 sim_kfs.simulate()
```
On the machine this was tested on, this resulted in the following:
```
2.06 ms ± 26.5 µs per loop (mean ± std. dev. of 3 runs, 10000 loops each)
2.02 ms ± 68.4 µs per loop (mean ± std. dev. of 3 runs, 10000 loops each)
```
These results suggest that - at least for this model - there are not noticeable computational gains from the CFA approach relative to the KFS approach. However, this does not rule out the following:
1. The Statsmodels implementation of the CFA simulation smoother could possibly be further optimized
2. The CFA apporach may only show improvement for certain models (for example with a large number of `endog` variables)
One simple way to take a first pass at assessing the first possibility is to compare the runtime of the Statsmodels implementation of the CFA simulation smoother to the Matlab implementation in the replication codes of Chan and Jeliakov (2009), available at http://joshuachan.org/code/code_TVPVAR.html.
While the Statsmodels version of the CFA simulation smoother is written in Cython and compiled to C code, the Matlab version takes advantage of the Matlab's sparse matrix capabilities. As a result, even though it is not compiled code, we might expect it to have relatively good performance.
On the machine this was tested on, the Matlab version typically ran the MCMC loop with 11,000 iterations in 70-75 seconds, while the MCMC loop in this notebook using the Statsmodels CFA simulation smoother (see above), also with 11,0000 iterations, ran in 40-45 seconds. This is some evidence that the Statsmodels implementation of the CFA smoother already performs relatively well (although it doesn't rule out that there are additional gains possible).
### Bibliography
Carter, Chris K., and Robert Kohn. "On Gibbs sampling for state space models." Biometrika 81, no. 3 (1994): 541-553.
Chan, Joshua CC, and Ivan Jeliazkov. "Efficient simulation and integrated likelihood estimation in state space models." International Journal of Mathematical Modelling and Numerical Optimisation 1, no. 1-2 (2009): 101-120.
De Jong, Piet, and Neil Shephard. "The simulation smoother for time series models." Biometrika 82, no. 2 (1995): 339-350.
Durbin, James, and Siem Jan Koopman. "A simple and efficient simulation smoother for state space time series analysis." Biometrika 89, no. 3 (2002): 603-616.
McCausland, William J., Shirley Miller, and Denis Pelletier. "Simulation smoothing for state–space models: A computational efficiency analysis." Computational Statistics & Data Analysis 55, no. 1 (2011): 199-212.
| github_jupyter |
# Research Notes:
Here, we implement empirical measurements of graph embedding algorithms. We implement multiple methods:
1) Link Prediction. This is done for all graphs. We remove a fixed % of edges in the graph and predict missing links in the graph. We train a logistic regression and LightGBM gradient boosted decision tree to predict the edges and report their AUC, accuracy and F1 score.
2) Clustering (on graphs with community/cluster labels). We use hierarchical agglomerative clustering on the graph embeddings and measure overlap of the embedding with the real-world communities in the network. Agglomerative Hierarchical clustering is chosen because it is deterministic and not sensitive to cluster shape or scaling or embedding metric space. We measure overlap with RAND index, mutual information score and Fowlkes-Mallows score.
3) Label prediction. This can be multilabel classifications (for graphs where a node can be in multiple groups) or regular classification performance. We train a logistic regression and LightGBM gradient boosted decision tree to predict the labels and report their AUC, accuracy and F1 score.
### First vs Second Order:
We see a sharp divide in empirical performance along first-order and higher-order graph embedding methods.
**First order** methods directly minimize the distance between nodes and their neighbours. Most graph embedding methods that are based around adjacency matrix factorization (laplacian eigenmaps, SVD, etc.) are first order.
**Higher order** methods take in account deeper graph structure. For instance, a *second order* method would account for neighbors-of-neighbors in each node's embeddings. 3rd and higher order deepen this relationship.
Note that you can do higher order embedding through graph factorization algorithms by augmenting the graph adjacency matrix.
For instance, one of the methods tested below samples random walks on the graph and generates a co-occurence matrix from these samples to train GLoVe (a first order algorithm). GGVec and GraRep generate higher-order adjacency matrices by taking the dot product of the graph's random walk markov chain transition matrix with itself, then taking a first order embedding method on that.
Methods based around random walks + word2vec (deepwalk, node2vec, etc.) are naturally higher order methods. The order can be constrained by reducing the word2vec window, or restricting walk length to be extremely short.
### Findings
We find that first-order methods generally perform better on clustering and label prediction than higher-order methods. Recommended first order methods are GGVec and ProNE.
On the other hand, higher order methods perform better on the link prediction task. Interestingly, the gap in link prediction performance is inexistant for artificially created graphs. This suggests higher order methods do learn some of the structure intrinsic to real world graphs.
These results put in context that it's important to have a diversity of downstream tasks when evaluating embedding models.
Moreover, we find that neural-net based methods (deepwalk, node2vec and descendants) are extremely inefficient with respect to output dimensions. They tend to perform much worse with smaller number of output dimensions.
### GGVec
We develop GGVec, a first (and higher) order embedding algorithm.
This method is very fast and scalable for large graphs by directly minimizing distange between nodes with edges (like GLoVe). It is naturally a first-order method but can be made higher order through the method mentionned above (dot product of graph transition matrix). Scaling of higher-order is worse however.
Moreover, this is the first algorithm that can embed directly from an edgelist file (because minimization loop is per-edge). This remains to be implemented.
```
import gc
import networkx as nx
import numpy as np
import os
import pandas as pd
import time
import scipy
import sklearn
from sklearn import cluster, linear_model
from sklearn.decomposition import TruncatedSVD
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.model_selection import train_test_split
from sklearn.multiclass import OneVsRestClassifier
import sys
import warnings # Silence perf warning
sys.path.append(os.path.realpath('..'))
import nodevectors
import csrgraph as cg
from csrgraph import methods
from nodevectors.evaluation import link_pred
from nodevectors.evaluation import graph_eval
# UMAP to test (on pip)
import umap
warnings.simplefilter("ignore")
def nx_node_weights(G, method, **kwargs):
"""Node Weights through networkX API"""
pr = np.zeros(len(G))
prdict = method(G, **kwargs)
for i in G.nodes:
pr[i] = prdict[i]
return pr
#### CONFIG
N_COMPONENTS = 32 # resulting embedding dim
SEED = 42 # RNG Seed
TEST_SIZE = 0.2
# For resampling tests
RESAMPLE_WALKS = 10
RESAMPLE_LEN = 6
```
# Data Availability
Data for these notebooks can be found here: https://github.com/VHRanger/Graph-Data
Just download it and point the graph generation methods below to it
The data is in a different repo to avoid polluting the pip package.
```
#### GRAPHS
#### Uncomment one to choose which graph to run evaluation on
#### Artificial random graphs
# G = nx.binomial_graph(700, 0.6)
# G, labels = graph_eval.make_cluster_graph(n_nodes=820, n_clusters=18, connections=1000, drop_pct=0.5)
# G, labels = graph_eval.make_weighed_cluster_graph(n_nodes=500, n_clusters=6, connections=1500, drop_pct=0.2, max_edge_weight=15)
#### Social graphs
# G, labels = graph_eval.make_blogcatalog(dedupe=True)
# G, mlabels = graph_eval.make_blogcatalog(dedupe=False)
G, labels = graph_eval.make_email()
# G, labels = graph_eval.get_karateclub("facebook") # twitch, github, facebook, wikipedia
# G = graph_eval.get_from_snap(url="http://snap.stanford.edu/data/facebook_combined.txt.gz", sep=' ', header=None, comment='#')
#### Biology Graphs
# G, mlabels = graph_eval.get_n2v_ppi("../data/bioNEV/node2vec_PPI")
#### Needs OutOfBounds Nodes support from CSRGraphs to work
# G = graph_eval.get_drugbank_ddi("../data/bioNEV/DrugBank_DDI")
# G, mlabels = graph_eval.get_mashup_ppi("../data/bioNEV/Mashup_PPI")
#### For Link Prediction: Split graph into train and test edge sets
#### (All nodes are still present in both)
G_train, testing_pos_edges = link_pred.split_train_test_graph(G, testing_ratio=TEST_SIZE)
#### Lazy way to set up evaluation
try:
y = labels.label
n_clusters = y.nunique()
HAS_LABELS = True
print(f"clusters: {n_clusters}")
except:
try: # Multilabels
y = MultiLabelBinarizer().fit_transform(mlabels.mlabels)
HAS_LABELS = True
print(f"multilabels: {y.shape[1]}")
except: # No Labels
HAS_LABELS = False
print("No Labels")
NNODES = len(G)
print(f"Nodes: {NNODES}\nEdges: {len(G.edges)}\nconnected: {nx.is_connected(G_train)}")
ggvec_params = dict(
n_components=N_COMPONENTS,
order=2,
tol=0.07,
tol_samples=10,
max_epoch=6_000,
learning_rate=0.1,
negative_ratio=0.15,
exponent=0.33,
verbose=True,
)
start_t = time.time()
w_train = nodevectors.GGVec(**ggvec_params).fit_transform(G_train)
print(f"Time: {time.time() - start_t :.4f}")
result = link_pred.LinkPrediction(w_train, G, G_train, testing_pos_edges)
time.sleep(0.1)
if HAS_LABELS:
w = nodevectors.GGVec(**ggvec_params).fit_transform(G)
graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
n2v_params = dict(
n_components=N_COMPONENTS,
epochs=20,
walklen=60,
return_weight=1.,
neighbor_weight=1.,
w2vparams={
"window":3,
"negative":5,
"iter":2,
"batch_words":128}
)
start_t = time.time()
w_train = nodevectors.Node2Vec(**n2v_params).fit_transform(G_train)
print(f"Time: {time.time() - start_t :.4f}")
result = link_pred.LinkPrediction(w_train, G, G_train, testing_pos_edges)
if HAS_LABELS:
w = nodevectors.Node2Vec(**n2v_params).fit_transform(G)
graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
pne_params = dict(
n_components=N_COMPONENTS,
step=5,
mu=0.2,
theta=0.5,
)
start_t = time.time()
pne = nodevectors.ProNE(**pne_params)
w_train = pne.fit_transform(G_train)
print(f"Time: {time.time() - start_t :.4f}")
result = link_pred.LinkPrediction(w_train, G, G_train, testing_pos_edges)
if HAS_LABELS:
pne = nodevectors.ProNE(**pne_params)
w = pne.fit_transform(G)
graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
grarep_params = dict(
n_components=N_COMPONENTS,
order=1,
embedder=TruncatedSVD(
n_iter=10,
random_state=42),
# merger=(lambda x : np.sum(x, axis=0)),
merger=lambda x : x[-1]
)
start_t = time.time()
w_train = nodevectors.GraRep(**grarep_params).fit_transform(G_train)
print(f"Time: {time.time() - start_t :.4f}")
result = link_pred.LinkPrediction(w_train, G, G_train, testing_pos_edges)
time.sleep(0.1)
if HAS_LABELS:
w = nodevectors.GraRep(**grarep_params).fit_transform(G)
graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
ump_params = dict(
embedder=umap.UMAP,
n_neighbors=3,
min_dist=0.,
metric='cosine',
normalize_graph=True,
n_components=N_COMPONENTS,
)
start_t = time.time()
w_train = nodevectors.SKLearnEmbedder(**ump_params).fit_transform(G_train)
print(f"Time: {time.time() - start_t :.4f}")
result = link_pred.LinkPrediction(w_train, G, G_train, testing_pos_edges)
if HAS_LABELS:
w = nodevectors.SKLearnEmbedder(**ump_params).fit_transform(G)
graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
### GLoVe with random walks ###
glove_params = dict(
n_components=N_COMPONENTS,
tol=0.0005,
max_epoch=6_000,
learning_rate=0.02,
max_loss=10.,
max_count=50,
exponent=0.5,
)
start_t = time.time()
wg = cg.csrgraph(G_train).random_walk_resample(walklen=RESAMPLE_LEN, epochs=RESAMPLE_WALKS)
w_train = nodevectors.Glove(**glove_params).fit_transform(wg)
print(f"Time: {time.time() - start_t :.4f}")
print(f"Virtual edges: {wg.dst.size}")
result = link_pred.LinkPrediction(w_train, G, G_train, testing_pos_edges)
if HAS_LABELS:
wg = cg.csrgraph(G).random_walk_resample(walklen=RESAMPLE_LEN, epochs=RESAMPLE_WALKS)
w = nodevectors.Glove(**glove_params).fit_transform(wg)
graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
# From the related karateclub lib (on pip)
# https://github.com/benedekrozemberczki/KarateClub
# from karateclub.node_embedding.neighbourhood import NodeSketch, Walklets
###### Slooooowwwwwww ########
# walklets_params = dict(
# walk_number=10,
# walk_length=30,
# dimensions=N_COMPONENTS,
# window_size=4,
# epochs=1,
# learning_rate=0.05
# )
# try: # Karateclub models don't handle certain graphs
# start_t = time.time()
# model = Walklets(**walklets_params)
# model.fit(G_train)
# print(f"Time: {time.time() - start_t :.3f}")
# w_train = model.get_embedding()
# result = link_pred.LinkPrediction(w_train, G, G_train, testing_pos_edges)
# if HAS_LABELS:
# model = Walklets(**walklets_params)
# model.fit(G)
# w = model.get_embedding()
# graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
# except: pass
### Completely random baseline ###
w = np.random.randn(len(G), N_COMPONENTS)
result = link_pred.LinkPrediction(w, G, G_train, testing_pos_edges)
try:
graph_eval.print_labeled_tests(w, y, test_size=TEST_SIZE, seed=SEED)
except: pass
```
| github_jupyter |
```
%autosave 60
%load_ext autoreload
%autoreload 2
%matplotlib inline
import json
import os
import pickle
import sys
from collections import Counter
from io import BytesIO
from pathlib import Path
from typing import Dict, List, Optional, Tuple, Union, cast
import cv2
import matplotlib as plt
import numpy as np
import pandas as pd
import PIL.Image as pil_img
import seaborn as sns
import sklearn as skl
from IPython.display import Image, display
from matplotlib.patches import Rectangle
from matplotlib_inline.backend_inline import set_matplotlib_formats
from pycocotools.helpers import CocoClassDistHelper as COCO
from pycocotools.helpers import CocoJsonBuilder, check_boxes, check_stats
from tqdm import tqdm
from geoscreens.consts import VIDEO_PATH
from geoscreens.data.metadata import get_all_metadata, get_geoguessr_split_metadata, load_metadata
from geoscreens.data.splitting import get_images_with_metadata, generate_train_val_splits, get_metadata_df
pd.set_option("display.max_colwidth", None)
pd.set_option("display.max_columns", 15)
pd.set_option("display.max_rows", 50)
# Suitable default display for floats
pd.options.display.float_format = "{:,.2f}".format
plt.rcParams["figure.figsize"] = (12, 10)
# This one is optional -- change graphs to SVG only use if you don't have a
# lot of points/lines in your graphs. Can also just use ['retina'] if you
# don't want SVG.
%config InlineBackend.figure_formats = ["retina"]
set_matplotlib_formats("pdf", "png")
coco_path = Path("/home/gbiamby/proj/geoscreens/datasets/geoscreens_012/geoscreens_012.json")
gs = COCO(coco_path)
all_metadata = get_all_metadata()
df_meta = get_metadata_df(all_metadata)
img_data = get_images_with_metadata(gs, df_meta)
# df_images = pd.DataFrame(img_data)
# df_images.rename(columns={"id": "image_id"}, inplace=True)
# df_images
check_boxes(coco_path)
# check_stats(coco_path)
```
---
```
pd.DataFrame(gs.get_cat_counts().values())
```
# Anchor Boxes
```
from pycocotools.helpers.anchors import compute_anchors
results = compute_anchors(coco_path, coco_path.parent / "anchors", [5, 60], stop_at_iou=0.90)
nc = 20
print(results[nc]["iou"])
df = pd.DataFrame(results[nc]["clusters"], columns=["width", "height"])
df["ratio"] = df.height / df.width
df["base_size"] = df.height / 8.0
display(df)
print(",".join([str(bs) for bs in [4,5,6,8,15,26,31,48,65,69]]))
print(",".join([f"{bs:0.02f}" for bs in df.ratio.values]))
print(",".join([str(bs) for bs in [4,6,23,52,63]]))
print(",".join([str(bs) for bs in [3,4,5,22,52,64]]))
print(",".join([f"{bs:0.02f}" for bs in df.ratio.values]))
print(",".join([str(bs) for bs in [4,5,6,8,15,26,31,48,65,69]]))
print(",".join([str(8*bs) for bs in [4,5,6,8,15,26,31,48,65,69]]))
ratios = [0.03232666, 0.32466155, 0.16829857, 0.06960362, 0.13733681, 0.26874476
, 0.21486479, 0.12025294, 0.10558306, 0.0686041, 0.05882353, 0.095259
, 0.10972197, 0.03576037, 0.05837962, 0.05233987, 0.16004019, 0.15555059
, 0.72779865, 0.56002639, 0.67524641, 0.74870974, 0.6576581, 0.41884503
, 0.57760766, 0.17744202, 0.7537875, 0.39796285, 0.25761093, 0.2220346
, 0.76633415, 0.36426718, 0.26349288, 0.52401231, 1.35222693, 0.27887195
, 0.71730647, 0.52501349, 0.76417979, 1.00861367, 2.57618869, 0.65038579
, 1.00513278, 1.82245827, 0.73063804, 2.34746611, 3.2583827, 3.12290856
, 3.47869761, 0.7429066 ]
sorted(ratios)
dims = [[836.08333333, 27.02777778],
[94.10889292, 30.55353902],
[202.06970509, 34.0080429, ],
[494.24817518, 34.40145985],
[253.79518072, 34.85542169],
[130.28324946, 35.01294033],
[166.73361082, 35.8251821, ],
[301.01072961, 36.19742489],
[347.28537736, 36.66745283],
[572.12323944, 39.25, ],
[685.93023256, 40.34883721],
[440.12784091, 41.92613636],
[393.22346369, 43.1452514, ],
[1276.47058824, 45.64705882],
[963.40540541, 56.24324324],
[1092.05263158, 57.15789474],
[554.37025316, 88.72151899],
[680.89130435, 105.91304348],
[166.62612613, 121.27027027],
[239.36842105, 134.05263158],
[204.12301587, 137.83333333],
[247.88268156, 185.59217877],
[289.76056338, 190.56338028],
[478.18852459, 200.28688525],
[360.89830508, 208.45762712],
[1278.13978495, 226.79569892],
[303.87195122, 229.05487805],
[578.85714286, 230.36363636],
[936.975, 241.375, ],
[1266.69444444, 281.25, ],
[378.425, 290., ],
[797.65116279, 290.55813953],
[1136.76470588, 299.52941176],
[592.925, 310.7, ],
[238.8375, 322.9625, ],
[1273.18867925, 355.05660377],
[552.18309859, 396.08450704],
[794.42857143, 417.08571429],
[597.62068966, 456.68965517],
[464.37837838, 468.37837838],
[200.325, 516.075, ],
[832.27710843, 541.30120482],
[560.125, 563., ],
[309.53030303, 564.10606061],
[836.28421053, 611.02105263],
[262.75438596, 616.80701754],
[189.66292135, 617.99438202],
[204.75925926, 639.44444444],
[199.86923077, 695.28461538],
[949.07142857, 705.07142857],]
def scale_dim(w, h, from_w=1280, from_h=720, dim_to=640):
return (
w * (dim_to / from_w),
h * (dim_to / from_h),
(h * (dim_to / from_h)) / (w * (dim_to / from_w)),
)
# [
# tf_dim(dim[0], dim[1])
# for dim in [
# [1280, 720],
# [1280/2, 720],
# [1280/4, 720],
# [1280, 720/2],
# [1280, 720/4],
# ]
# ]
centroids_scaled = [tf_dim(dim[0], dim[1]) for dim in dims]
sorted(centroids_scaled, key=lambda x: x[2])
[0.05,0.11,0.16,0.19,0.27,0.39,0.]
from mmdet.core.anchor import AnchorGenerator
ag = AnchorGenerator(
strides=[2],
scales=[4,8],
ratios=[1.0],
base_sizes=[2],
)
print(ag.grid_priors([(2, 2)], device="cpu"))
# base size gets multiplied by scales, and that gives the anchor box sizes
# strides don't get multiplied
# print([[s] for s in [4, 8, 16, 32, 64]])
ag = AnchorGenerator(
strides=[4, 8, 16, 32, 64],
# ratios=[0.08, 0.16, 0.25, 0.36, 0.5, 0.7, 1, 2],
ratios=[1.0],
scales=[8],
)
ag.grid_priors([(2, 2)])
```
| github_jupyter |
<img src="https://raw.githubusercontent.com/dimitreOliveira/MachineLearning/master/Kaggle/Microsoft%20Malware%20Prediction/Microsoft_logo.png" width="600" height="170">
<h1><center>Microsoft Malware Prediction</center></h1>
<h2><center>Malware detection - Encoding evaluation</center></h2>
```
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import gc
import category_encoders as ce
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
dtypes = {
'ProductName': 'category',
'EngineVersion': 'category',
'AvSigVersion': 'category',
'IsSxsPassiveMode': 'int8',
'CountryIdentifier': 'int16',
'OrganizationIdentifier': 'float16',
'GeoNameIdentifier': 'float16',
'LocaleEnglishNameIdentifier': 'int8',
'Platform': 'category',
'Processor': 'category',
'OsVer': 'category',
'OsSuite': 'int16',
'OsPlatformSubRelease': 'category',
'SkuEdition': 'category',
'IsProtected': 'float16',
'IeVerIdentifier': 'float16',
'SmartScreen': 'category',
'UacLuaenable': 'float32',
'Census_MDC2FormFactor': 'category',
'Census_DeviceFamily': 'category',
'Census_ProcessorCoreCount': 'float16',
'Census_ProcessorManufacturerIdentifier': 'float16',
'Census_PrimaryDiskTypeName': 'category',
'Census_SystemVolumeTotalCapacity': 'float32',
'Census_TotalPhysicalRAM': 'float32',
'Census_ChassisTypeName': 'category',
'Census_PowerPlatformRoleName': 'category',
'Census_OSVersion': 'category',
'Census_OSArchitecture': 'category',
'Census_OSBranch': 'category',
'Census_OSBuildRevision': 'int32',
'Census_OSEdition': 'category',
'Census_OSSkuName': 'category',
'Census_OSInstallTypeName': 'category',
'Census_OSInstallLanguageIdentifier': 'float16',
'Census_OSUILocaleIdentifier': 'int16',
'Census_OSWUAutoUpdateOptionsName': 'category',
'Census_GenuineStateName': 'category',
'Census_ActivationChannel': 'category',
'Census_FlightRing': 'category',
'Census_IsVirtualDevice': 'float16',
'Census_IsTouchEnabled': 'int8',
'Census_IsAlwaysOnAlwaysConnectedCapable': 'float16',
'Wdft_IsGamer': 'float16',
'Wdft_RegionIdentifier': 'float16',
'HasDetections': 'int8'
}
cols = ['Census_OSInstallTypeName', 'Census_PowerPlatformRoleName', 'Census_OSSkuName',
'Wdft_RegionIdentifier', 'ProductName', 'Census_InternalBatteryType', 'Census_DeviceFamily',
'Census_OSBranch', 'OsPlatformSubRelease', 'Census_ActivationChannel', 'SmartScreen',
'Census_ProcessorManufacturerIdentifier', 'UacLuaenable', 'Census_OSInstallLanguageIdentifier',
'Census_OSArchitecture', 'Census_OSEdition', 'SkuEdition', 'Census_OSWUAutoUpdateOptionsName',
'Census_MDC2FormFactor', 'OsSuite', 'Census_ChassisTypeName', 'Census_PrimaryDiskTypeName',
'Census_GenuineStateName', 'OrganizationIdentifier', 'Census_FlightRing']
#### Encoding ###
def mean_encoding(dataset, columns, target):
for feature in columns:
dataset_target_mean = dataset.groupby(feature)[target].mean() # calculate mean
enc_name = feature #('%s_target_enc' % feature) # new variable name
dataset[enc_name] = dataset[feature].map(dataset_target_mean) #assign new values
return(dataset)
def frequency_encoding(dataset, columns):
for feature in columns:
dataset_target_mean = dataset[feature].value_counts() # calculate count
enc_name = feature # ('%s_target_enc' % feature) # new variable name
dataset[enc_name] = dataset[feature].map(dataset_target_mean) #assign new values
return(dataset)
def binary_encoding(dataset, columns):
encoder = ce.BinaryEncoder(cols=columns)
dataset = encoder.fit_transform(dataset)
return(dataset)
def onehot_encoding(dataset, columns):
encoder = ce.OneHotEncoder(cols=columns)
dataset = encoder.fit_transform(dataset)
return(dataset)
def ordinal_encoding(dataset, columns):
encoder = ce.OrdinalEncoder(cols=columns)
dataset = encoder.fit_transform(dataset)
return(dataset)
def loo_encoding(dataset, columns, target):
# separete datasets
y = dataset[target]
X = dataset.drop(target, axis = 1)
encoder = ce.LeaveOneOutEncoder(cols=columns)
encoder.fit(X, y)
dataset = encoder.transform(X, y)
dataset[target] = y
return(dataset)
def hash_encoding(dataset, columns):
encoder = ce.HashingEncoder(cols=columns, drop_invariant=True)
dataset = encoder.fit_transform(dataset)
return(dataset)
### Correlation ###
def get_correlation(dataframe, col, target):
encoded_feature = dataframe[col].values
cor = abs(np.corrcoef(dataframe[target].values, encoded_feature)[0][1]) # getting abs value here so it will be easier to compare
return(cor)
def generate_corr_list(dataframe, columns, method, df_name, tag, target):
for feature in columns:
try:
enc_name = feature #'%s%s' % (feature, tag) # new variable name
# saving correlation
# corr_features.append([df_name, method, ('%s_target_enc' % feature), get_correlation(dataframe, enc_name)])
corr_features.append([df_name, method, enc_name, get_correlation(dataframe, enc_name, target)])
enc_columns.append(enc_name)
except:
print('ERROR')
pass
def generate_corr_df():
# creating dataframe
return pd.DataFrame(corr_features, columns=['Dataset', 'Method', 'Feature', 'Correlation'])
### Utils ###
def has_high_cardinality(dataset, columns):
CARDINALITY_THRESHOLD = 100
high_cardinality_columns = [c for c in columns if dataset[c].nunique() >= CARDINALITY_THRESHOLD]
return high_cardinality_columns
def only_numbers(dataset, columns):
for feature in columns:
enc_name = feature # ('%s_target_enc' % feature) # new variable name
dataset[enc_name] = dataset[feature].str.extractall('(\d+)').unstack().apply(''.join, axis=1)
return dataset
```
# **Data**
```
target = 'HasDetections'
df = pd.read_csv('../input/train.csv', dtype=dtypes, usecols=(cols + [target]))
# Would treat differently columns with high cardinality if we had any
has_high_cardinality(df, cols)
# creating correlation list
corr_features = []
enc_columns = []
tag = '_target_enc'
```
## **Correlation without encode**
```
generate_corr_list(df, cols, 'without_encode', 'low', '', target)
```
## **Mean**
```
mean_df = mean_encoding(df, cols, target)
generate_corr_list(mean_df, cols, 'mean', 'low', tag, target)
# freeing space
del mean_df
gc.collect()
```
## **Frequency**
```
frequency_df = frequency_encoding(df, cols)
generate_corr_list(frequency_df, cols, 'frequency', 'low', tag, target)
# freeing space
del frequency_df
gc.collect()
```
## **Bynary**
```
# binary_df = binary_encoding(df.astype(str), cols)
# binary_df.head()
```
## **One hot**
```
# onehot_df = onehot_encoding(df.astype(str), cols)
# onehot_df.head()
```
## **Leave one out**
```
lov_df = loo_encoding(df, cols, target)
generate_corr_list(lov_df, cols, 'leave_one_out', 'low', tag, target)
# freeing space
del lov_df
gc.collect()
```
## **Hashing**
```
# hash_df = hash_encoding(df, cols)
# hash_df.head()
```
# **Result**
```
corr_df = generate_corr_df()
sns.set(rc={'figure.figsize':(8.7,20.27)})
ax = sns.barplot(x="Correlation", y="Feature", hue="Method", data=corr_df)
```
### Conclusion
There is a couple of different encoder that should be used in the model, they are: mean, leave one out, one hot and hash encode. Mean and leave one have higher correlation for some features, while one hot and hash can't be tested against the former two encoders.
OBS: some of the encodes where left out in the commit because they were incapabilitating the kernel commit.
| github_jupyter |
# Maze Solver
In this notebook, we write a maze solver by solving the Poisson equation with two Dirichlet boundary conditions imposed on the two faces that correspond to the start and end of the maze, respectively.
The logic is pretty simple: once we have the solution, we just need to start off from one face and follow the gradient. Since the gradient in the deadends is almost close to 0, following the nonzero gradient should guide us toward the other side of the maze.
We implement two different approaches:
1. Direct numerical simulation
Here, we first convert the image into a Cubic network, trim the pores that correspond to the walls, and finally run a basic `OhmicConduction` (or `FickianDiffusion`) on the resulting trimmed network.
2. Network extraction
Here, we first use the SNOW algorithm to extract the equivalent network of the maze. Note that the nodes in the equivalent network will not exactly give us the corners of the maze, but at least it gives us a rough idea, enough for solving the maze! Then, like the first approach, we run a basic `OhmicConduction` on the extracted network. The advantage of this approach is that it's way faster due to much fewer unknowns.
Note: Inspired by this post by Jeremy Theler https://www.linkedin.com/posts/jeremytheler_how-to-solve-a-maze-without-ai-use-laplaces-activity-6831291311832760320-x9d5
```
# Install the required pmeal packages in the current Jupyter kernel
import sys
try:
import openpnm as op
except:
!{sys.executable} -m pip install openpnm
import openpnm as op
try:
import porespy as ps
except:
!{sys.executable} -m pip install porespy
import porespy as ps
import requests
import numpy as np
from scipy import ndimage
import matplotlib.pyplot as plt
import porespy as ps
import openpnm as op
from openpnm.utils import tic, toc
from PIL import Image
from io import BytesIO
%config InlineBackend.figure_formats = ['svg']
ws = op.Workspace()
ws.settings["loglevel"] = 60
```
## Load maze samples
```
im_size = 'medium'
if im_size == 'small':
url = 'https://imgur.com/ZLbV4eh.png'
elif im_size == 'medium':
url = 'https://imgur.com/A3Jx8SJ.png'
else:
url = 'https://imgur.com/FLJ21e5.png'
response = requests.get(url)
img = Image.open(BytesIO(response.content))
im = np.array(img.getdata()).reshape(img.size[0], img.size[1], 4)[:, :, 0]
im = im == 255
Nx, Ny, = im.shape
fig, ax = plt.subplots(figsize=(5, 5))
ax.imshow(im, cmap='Blues', interpolation="none")
ax.axis("off");
```
# Approach A: Direct numerical simulation
## Thicken the walls to reduce number of unknowns
```
# Structuring element for thickening walls
strel = np.array([[1, 1, 1],
[1, 1, 1],
[1, 1, 1]])
# Save some computation by thickening the walls
def thicken_wall(im):
return ~ndimage.morphology.binary_dilation(~im, structure=strel)
for _ in range(5):
im = thicken_wall(im)
fig, ax = plt.subplots(figsize=(5, 5))
ax.imshow(im, cmap='Blues', interpolation="none")
ax.axis("off")
```
## Convert the maze into a Cubic network
```
# Get top and bottom boundaries
BP_top = np.zeros_like(im)
BP_bot = np.zeros_like(im)
BP_top[0, :] = True
BP_bot[-1, :] = True
BP_top *= im
BP_bot *= im
# Make a cubis network with same dimensions as image and assign the props
net = op.network.Cubic(shape=[Nx, Ny, 1])
net['pore.index'] = np.arange(0, net.Np)
net['pore.BP_top'] = BP_top.flatten()
net['pore.BP_bot'] = BP_bot.flatten()
# Trim wall pores
op.topotools.trim(network=net, pores=~im.flatten())
```
## Solve the Poisson equation ($\nabla^2 \phi = 0$) on the maze
```
# Set up a dummy phase and apply uniform arbitrary conductance
phase = op.phases.GenericPhase(network=net)
phase['throat.electrical_conductance'] = 1.0
# Run algorithm
alg = op.algorithms.OhmicConduction(network=net, phase=phase)
alg.set_value_BC(pores=net.pores('BP_top'), values=0.0)
alg.set_value_BC(pores=net.pores('BP_bot'), values=1.0)
tic()
alg.run()
dt = toc(quiet=True);
print(f'Solve time: {dt:.3f} s')
```
## Follow the gradient!
```
# Calculate flux in throats and show in pores
# Note: No need to calculate pore.rate as it auto interpolates from throat values
phase['throat.rate'] = alg.rate(throats=net.Ts, mode='single')
rate_im = np.ones([Nx, Ny]).flatten() * np.nan
rate_im[net['pore.index']] = phase['pore.rate']
rate_im = rate_im.reshape([Nx, Ny])
# Plot the maze solution
fig, ax = plt.subplots(figsize=(5, 5))
ax.imshow(rate_im, cmap='jet', interpolation="none")
ax.axis("off");
```
# Approach B: Network extraction
## Network extraction using SNOW algorithm
```
# We need to pass image transpose since matrix xy coords is inverted
# i.e., row is x and col is y, whereas in Cartesian convention, it's the opposite.
out = ps.networks.snow2(im.T)
proj = op.io.PoreSpy.import_data(out.network)
net = proj.network
```
## Solve the Poisson equation ($\nabla^2 \phi = 0$) on the extracted network
```
# Set up a dummy phase and apply uniform arbitrary conductance
phase = op.phases.GenericPhase(network=net)
phase['throat.electrical_conductance'] = 1.0
# Run algorithm
alg = op.algorithms.OhmicConduction(network=net, phase=phase)
alg.set_value_BC(pores=net.pores('ymin'), values=0.0)
alg.set_value_BC(pores=net.pores('ymax'), values=1.0)
tic()
alg.run()
dt = toc(quiet=True);
print(f'Solve time: {dt:.3f} s')
```
## Follow the gradient!
```
# Get throat rate values
phase['throat.rate'] = alg.rate(throats=net.Ts, mode='single')
# Plot the maze solution (i.e., throat rates!)
fig, ax = plt.subplots(figsize=(5, 5))
op.topotools.plot_connections(net, ax=ax,
color_by=phase["throat.rate"],
linewidth=2, cmap="Wistia")
ax.imshow(im, interpolation="none", cmap='Blues');
ax.axis("off");
```
| github_jupyter |
# Evolution of $M_{\rm D}$ and $\omega$ over Time
This notebook aims to reproduce Fig. 2 from [Gibson et al. (2017)](https://arxiv.org/abs/1706.04802) which examines how the disc mass $M_{\rm D}$ and the angular frequency $\omega$ evolve over time after `odeint` has solved the ODEs and how the evolution is affected by different values of timescale ratio $\epsilon$ and mass ratio $\delta$. By also examining the fastness parameter evolution, we can observe how the variation in $\epsilon$ and $\delta$ affects the "switch-on" of the propeller regime ($\Omega > 1$).
## 1. Module Imports
Here we import standard libraries `numpy` and `matplotlib` and import `odeint` from `scipy`. Rather than redefine `init_conds` and `ODEs` again, they have been exported to a package called `magnetar` and can also be imported.
```
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
from magnetar.funcs import init_conds, ODEs
%matplotlib inline
```
## 2. Variable Set-up
Here we set up global constants required for the calculation and the values of $\epsilon$ and $\delta$ we'd like to test.
```
# Constants
G = 6.674e-8 # Gravitational constant - cgs units
Msol = 1.99e33 # Solar mass - grams
M = 1.4 * Msol # Magnetar mass - grams
R = 1.0e6 # Magnetar radius - cm
alpha = 0.1 # Sound speed prescription
cs7 = 1.0 # Sound speed - cm/s
GM = G * M
B = 1.0 # Magnetic Field Strength - 10^15 Gauss
P = 5.0 # Initial spin period - milliseconds
MdiscI = 0.001 # Initial disc mass - Solar masses
RdiscI = 100.0 # Disc radius - km
mu = 1.0e15 * B * (R ** 3.0) # Magnetic Dipole Moment
Rdisc = RdiscI * 1.0e5 # Convert disc radius to cm
tvisc = Rdisc / (alpha * cs7 * 1.0e7) # Viscous timescale - seconds
t = np.logspace(0.0, 6.0, base=10.0, num=10001) # Time array
y = init_conds(P, MdiscI) # Initial conditions
eps_vals = [1.0, 1.0, 10.0, 10.0] # Epsilon values
delt_vals = [1.0, 10.0, 1.0, 10.0] # Delta values
```
## 3. Solve the Equations and Plot the Results
Here we loop over the desired $\epsilon$ and $\delta$ values and plot the figure.
```
# Set up a figure with 3 subplots
fig, (ax1, ax2, ax3) = plt.subplots(3, 1, sharex=True, figsize=(6,7))
ax3.axhline(1.0, ls='--', c='k') # Marks the w > 1 condition
colours = ['r', 'r', 'g', 'g'] # Line colours
lines = ['-', '--', '-', '--'] # Linestyles
# Loop over the epsilon and delta values
for i, vals in enumerate(zip(eps_vals, delt_vals)):
epsilon, delta = vals
# Solve the ODEs and split the solution
# n value defaults to 1 when absent from args
soln = odeint(ODEs, y, t, args=(B, MdiscI, RdiscI, epsilon, delta))
Mdisc = soln[:,0]
omega = soln[:,1]
# Radii
Rm = ((mu ** (4.0 / 7.0)) * (GM ** (-1.0 / 7.0)) * (Mdisc / tvisc) **
(-2.0 / 7.0))
Rc = (GM / (omega ** 2.0)) ** (1.0 / 3.0)
# Plotting
ax1.loglog(t, Mdisc/Msol, ls=lines[i], c=colours[i],
label='$\epsilon$ = {0}; $\delta$ = {1}'.format(
int(epsilon), int(delta)))
ax2.semilogx(t, omega, ls=lines[i], c=colours[i])
ax3.semilogx(t, Rm/Rc, ls=lines[i], c=colours[i])
# Plot formatting
ax1.set_xlim(1.0e0, 1.0e6)
ax1.set_ylim(bottom=1.0e-8)
ax2.set_xlim(1.0e0, 1.0e6)
ax3.set_xlim(1.0e0, 1.0e6)
ax3.set_ylim(0.0, 2.0)
ax1.tick_params(axis='both', which='major', labelsize=8)
ax2.tick_params(axis='both', which='major', labelsize=8)
ax3.tick_params(axis='both', which='major', labelsize=8)
ax1.legend(loc='upper right', fontsize=8)
ax3.set_xlabel('Time (s)', fontsize=10)
ax1.set_ylabel('$M_{\\rm D}~\left({\\rm M}_{\odot}\\right)$', fontsize=10)
ax2.set_ylabel('$\omega~\left({\\rm s}^{-1}\\right)$', fontsize=10)
ax3.set_ylabel('$r_{\\rm m}/r_{\\rm c}$', fontsize=10);
```
<div class="alert alert-block alert-warning">
<b>Warning:</b> This plot is currently not reproducible from the paper.
</div>
| github_jupyter |
# Matrix Multiplication
:label:`ch_matmul_cpu`
We saw the NumPy `dot` operator nearly reaches the peak performance of our CPU (the Xeon E5-2686 v4) in :numref:`ch_cpu_arch`. In this section, we will investigate multiple scheduling strategies for this operator in TVM.
## Setup
```
%matplotlib inline
import d2ltvm
import numpy as np
import timeit
import tvm
from tvm import te
target = 'llvm -mcpu=skylake-avx512'
```
As we did in :numref:`ch_vector_add_cpu`, we first define a method to measure NumPy performance as our baseline.
```
# Save to the d2ltvm package.
def np_matmul_timer(n):
timer = timeit.Timer(setup='import numpy as np\n'
'import d2ltvm\n'
'a, b, c = d2ltvm.get_abc(%s)' % str((n,n)),
stmt = 'np.dot(a, b, out=c)')
return timer.timeit
sizes = 2**np.arange(5, 12, 1)
exe_times = [d2ltvm.bench_workload(np_matmul_timer(n)) for n in sizes]
np_gflops = 2 * sizes **3 / 1e9 / np.array(exe_times)
```
## Default Schedule
The default schedule consists of three nested for-loops.
```
def default(n):
A, B, C = d2ltvm.matmul(n, n, n)
return te.create_schedule(C.op), (A, B, C)
s, args = default(64)
print(tvm.lower(s, args, simple_mode=True))
```
To benchmark its performance, we also define a reusable method as we did in :numref:`ch_vector_add_cpu`.
```
# Save to the d2ltvm package.
def bench_matmul_tvm(func, sizes, target):
def workload(nrepeats):
timer = mod.time_evaluator(mod.entry_name, ctx=ctx, number=nrepeats)
return timer(a, b, c).mean * nrepeats
times = []
for n in sizes:
s, (A, B, C) = func(int(n))
mod = tvm.build(s, [A, B, C], target)
ctx = tvm.context(target, 0)
a, b, c = d2ltvm.get_abc((n, n), lambda x: tvm.nd.array(x, ctx=ctx))
times.append(d2ltvm.bench_workload(workload))
return 2 * sizes**3 / 1e9 / np.array(times)
```
The default schedule follows the computation illustrated in :numref:`fig_matmul_default`.
It's not surprising to see that the default schedule doesn't perform well, especially for large matrices that cannot fit into the cache.
```
default_gflops = bench_matmul_tvm(default, sizes, target)
d2ltvm.plot_gflops(sizes, [np_gflops, default_gflops], ['numpy', 'default'])
```
## Reordering Axes
The first problem we can see from :numref:`fig_matmul_default` is that matrix $B$ is accessed column by column while its elements are stored by rows (i.e. matrix $B$ is in [row-major](https://en.wikipedia.org/wiki/Row-_and_column-major_order)). In other words, in the pseudo code above, we iterate axis `y` before axis `k`. Simply switching these two for-loops will make all elements read and written sequentially. :numref:`fig_matmul_reorder` illustrates the changed the data access pattern.

:label:`fig_matmul_reorder`
To implement it, we change the axes order from (`x`, `y`, `k`) to (`x`, `k`, `y`) by the `reorder` primitive. The corresponding pseudo code verifies that we are processing all matrices row by row now.
```
def reorder(n):
s, (A, B, C) = default(n)
(x, y), (k,) = C.op.axis, C.op.reduce_axis
s[C].reorder(x, k, y)
return s, (A, B, C)
s, args = reorder(64)
print(tvm.lower(s, args, simple_mode=True))
```
We can see that the reordering significantly improves the performance compared to the default schedule.
```
reorder_gflops = bench_matmul_tvm(reorder, sizes, target)
d2ltvm.plot_gflops(sizes, [np_gflops, default_gflops, reorder_gflops],
['numpy', 'default', 'reorder'])
```
## Parallelization
Let's revisit the pseudo code above. In the outermost for-loop `for (x, 0, 64)`, each time we compute the results of a row in $C$. Each row can be computed in parallel, so we can make the schedule parallelize axis `x`.
```
def parallel(n):
s, (A, B, C) = reorder(n)
s[C].parallel(C.op.axis[0])
return s, (A, B, C)
s, args = parallel(64)
print(tvm.lower(s, args, simple_mode=True))
parallel_gflops = bench_matmul_tvm(parallel, sizes, target)
d2ltvm.plot_gflops(sizes, [np_gflops, default_gflops, reorder_gflops, parallel_gflops],
['numpy', 'default', 'reorder', 'parallel'])
```
Parallelization improves the performance again. But we can see that there is still a gap compared to NumPy on large matrices, specially when they cannot fit into the L2 cache. We will try to resolve it in the next section.
## Summary
- Reordering the for-loops in matrix multiplication properly improves the performance.
- Proper thread-level parallelization also improves the performance.
## Exercises
1. Change the number of threads
1. Try to order the axes in method `parallel` differently
1. Benchmark matrix multiplication in larger sizes
| github_jupyter |
# A Comprehensive Data Analysis on a WhatsApp Group Chat
##### *Author*: [Tushar Nankani](https://www.linkedin.com/in/tusharnankani/)
## *Overview*
- Introduction
- Data Retrieval & Preprocessing
- Exploratory Data Analysis
- Data Visualization
- Data Interpretation
- Summarizing the Inferences
- Conclusion
# *Introduction*:
Whatsapp has quickly become the world’s most popular text and voice messaging application. Specializing in cross-platform messaging with over 1.5 billion monthly active users, this makes it the most popular mobile messenger app worldwide.
- I thought of various projects on which I could analyse data like - *Air Quality Index* or The *cliched* *Covid-19 Data Analysis*.
- But I thought why not do **Data Analysis on a WhatsApp group chat** of *college students* and find out interesting insights about *who is most active, who are ghosts (the ones who do not reply), my sleep schedule,* *the most used emoji, the sentiment score of each person, who swears the most, the most actives times of the day, or does the group use phones during college teaching hours?*
- These would be some interesting insights for sure, more for me than for you, since the people in this chat are people I know personally.
## Beginning. How do I export my conversations? From Where To Obtain Data?
- The first step is **Data Retrieval & Preprocessing**, that is to **gather the data**. WhatsApp allows you to **export your chats** through a **.txt format**.
- Go to the respective chat, which you want to export!
<img src="https://imgur.com/lYd4A4J.png" width=200 length=200 align="center">
<!--  -->
- Tap on **options**, click on **More**, and **Export Chat.**
<img src="https://imgur.com/KD0vCs1.png" width=200 length=200 align="center">
<!--  -->
- I will be Exporting **Without Media.**
#### NOTE:
- Without media: exports about **40k messages **
- With media: exports about *10k messages along with pictures/videos*
- While exporting data, *avoid including media files* because if the number of media files is greater than certain figure then not all the media files are exported.
<img src="https://imgur.com/BlQx2dl.png" width=300 length=300 align="center">
<!--  -->
## Opening this .txt file up, you get messages in a format that looks like this:

# *Importing Necessary Libraries*
We will be using :
1. **Regex (re)** to extract and manipulate strings based on specific patterns.
- References:
- [Regex - Python Docs](https://docs.python.org/3/library/re.html)
- [Regex cheatsheet](https://www.rexegg.com/regex-quickstart.html)
- [Regex Test - live](https://regexr.com/)
- [Datetime Format](http://strftime.org/)
2. **pandas** for analysis.
3. **matlotlib** and **seaborn** for visualization.
4. **emoji** to deal with emojis.
- References:
- [Python Docs](https://pypi.org/project/emoji/)
- [Emoji](https://github.com/carpedm20/emoji)
- [EMOJI CHEAT SHEET](https://www.webfx.com/tools/emoji-cheat-sheet/)
5. **wordcloud** for the most used words.
```
import re
import datetime
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from wordcloud import WordCloud, STOPWORDS
import emoji
import itertools
from collections import Counter
import warnings
%matplotlib inline
warnings.filterwarnings('ignore')
```
##### NOTE:
- This special command `%matplotlib inline` to ensure that plots are shown and embedded within the Jupyter notebook itself. Without this command, sometimes plots may show up in pop-up windows.
- `warnings.filterwarnings('ignore')`: The warnings filter controls whether warnings are ignored, displayed, or turned into errors (raising an exception). [REFERENCE](https://docs.python.org/3/library/warnings.html)
## *Preparation and reading data*
```
def rawToDf(file, key):
'''Converts raw .txt file into a Data Frame'''
split_formats = {
'12hr' : '\d{1,2}/\d{1,2}/\d{2,4},\s\d{1,2}:\d{2}\s[APap][mM]\s-\s',
'24hr' : '\d{1,2}/\d{1,2}/\d{2,4},\s\d{1,2}:\d{2}\s-\s',
'custom' : ''
}
datetime_formats = {
'12hr' : '%d/%m/%Y, %I:%M %p - ',
'24hr' : '%d/%m/%Y, %H:%M - ',
'custom': ''
}
with open(file, 'r', encoding='utf-8') as raw_data:
# print(raw_data.read())
raw_string = ' '.join(raw_data.read().split('\n')) # converting the list split by newline char. as one whole string as there can be multi-line messages
user_msg = re.split(split_formats[key], raw_string) [1:] # splits at all the date-time pattern, resulting in list of all the messages with user names
date_time = re.findall(split_formats[key], raw_string) # finds all the date-time patterns
df = pd.DataFrame({'date_time': date_time, 'user_msg': user_msg}) # exporting it to a df
# converting date-time pattern which is of type String to type datetime,
# format is to be specified for the whole string where the placeholders are extracted by the method
df['date_time'] = pd.to_datetime(df['date_time'], format=datetime_formats[key])
# split user and msg
usernames = []
msgs = []
for i in df['user_msg']:
a = re.split('([\w\W]+?):\s', i) # lazy pattern match to first {user_name}: pattern and spliting it aka each msg from a user
if(a[1:]): # user typed messages
usernames.append(a[1])
msgs.append(a[2])
else: # other notifications in the group(eg: someone was added, some left ...)
usernames.append("group_notification")
msgs.append(a[0])
# creating new columns
df['user'] = usernames
df['message'] = msgs
# dropping the old user_msg col.
df.drop('user_msg', axis=1, inplace=True)
return df
```
#### *NOTE*:
- Since WhatsApp texts are *multi-line*, you cannot just read the file line by line and get each message that you want. Instead, you need a way to *identify* if a line is a new message or part of an old message. You could use regular expressions.
- While reading each line, I split it based on a comma and take the first item returned from the `split()` function. If the line is a new message, the first item would be a valid date, and it will be appended as a new message to the list of messages. If it’s not, the message is part of the previous message, and hence, will be appended to the end of the previous message as one continuous message.
```
df = rawToDf('whatsapp-chat-data.txt', '12hr')
```
## *Prior Information of my Whatsapp Data*
- This is my most active college coding group.
- It has **235+** participants; 237 to be precise.
- This group was made in **January 2020.**
## Pre-Processing
```
df.info()
```
### The dataset contains 3 rows, and 13655 respective entries.
- Here is how it looks like:
```
df.sample(10)
```
### An important observation
- One might wonder there are no NaNs, but as you can see, there are some rows, where messages are an *empty string.*
- This can be because, the message might just contain **emojis.**
#### To see, how many such rows are there, we can use `df[df['message'] == ""].shape[0]`
```
df[df['message'] == ""].shape[0]
```
## Adding extra *helper columns for analysis and visualization*
```
df['day'] = df['date_time'].dt.strftime('%a')
df['month'] = df['date_time'].dt.strftime('%b')
df['year'] = df['date_time'].dt.year
df['date'] = df['date_time'].apply(lambda x: x.date())
```
# Now that we have a clean DataFrame to work with, it’s time to perform analysis on it.
### Final Data Frame
```
df
```
# *Data Analysis*
**1. Overall frequency of total messages on the group.**
**2. Top 10 most active days.**
**3. Top 10 active users on the group (with a twist).**
- Ghosts present in the group. (shocking results.)
**4. Top 10 users most sent media.**
**5. Top 10 most used emojis.**
**6. Most active hours and days.**
- Heatmaps of weekdays and months.
- Most active hours, weekdays, and months.
**7. Most used words - WordCloud**
# 1. Overall frequency of total messages on the group.
- I will first do this to get a look at overall data.
I will plot a simple line graph to see the frequency of messages over the months.
##### I expect to see a nice line graph with crests and troughs in odd places.
```
df1 = df.copy() # I will be using a copy of the original data frame everytime, to avoid loss of data!
df1['message_count'] = [1] * df1.shape[0] # adding extra helper column --> message_count.
df1.drop(columns='year', inplace=True) # dropping unnecessary columns, using `inplace=True`, since this is copy of the DF and won't affect the original DataFrame.
df1 = df1.groupby('date').sum().reset_index() # grouping by date; since plot is of frequency of messages --> no. of messages / day.
df1
# Improving Default Styles using Seaborn
sns.set_style("darkgrid")
# For better readablity;
import matplotlib
matplotlib.rcParams['font.size'] = 20
matplotlib.rcParams['figure.figsize'] = (27, 6) # Same as `plt.figure(figsize = (27, 6))`
# A basic plot
plt.plot(df1.date, df1.message_count)
plt.title('Messages sent per day over a time period');
# Could have used Seaborn's lineplot as well.
# sns.lineplot(df1.date, df1.message_count);
# Saving the plots
plt.savefig('msg_plots.svg', format = 'svg')
```
# 2 *Top 10 most active days.*
```
top10days = df1.sort_values(by="message_count", ascending=False).head(10) # Sort values according to the number of messages per day.
top10days.reset_index(inplace=True) # reset index in order.
top10days.drop(columns="index", inplace=True) # dropping original indices.
top10days
# Improving Default Styles using Seaborn
sns.set_style("darkgrid")
# For better readablity;
import matplotlib
matplotlib.rcParams['font.size'] = 10
matplotlib.rcParams['figure.figsize'] = (12, 8)
# A bar plot for top 10 days
sns.barplot(top10days.date, top10days.message_count, palette="hls");
# Saving the plots
plt.savefig('top10_days.svg', format = 'svg')
```
#### Apparently, the group was very active on 13th Spetember'20
- Because we were discussing fundamental yet tricky and brain-wracking "Guess the Output" Java questions!
# 3. Top 10 active users on the group.
### Before, analysing that, we will see the *number of Ghosts* in the group.
```
# Total number of people who have sent at least one message on the group;
print(f"Total number of people who have sent at least one message on the group are {len(df.user.unique()) - 1}") # `-1` because excluding "group_notficiation"
print(f"Number of people who haven't sent even a single message on the group are {237 - len(df.user.unique()) - 1}")
```
### *Shocking Result*
- Total number of people who have sent at least one message on the group are **154**.
- BUT, the total number of participants were **237**.
- **That means 81 people in the group have not sent even a single message throughout these 9 months and 13500+ messages.**
## Now, pre-processing top 10 active users.
```
df2 = df.copy()
df2 = df2[df2.user != "group_notification"]
top10df = df2.groupby("user")["message"].count().sort_values(ascending=False)
# Final Data Frame
top10df = top10df.head(10).reset_index()
top10df
```
## Now, visualizing top 10 active users.
### *Replacing names with initials for better visualization*
```
top10df['initials'] = ''
for i in range(10):
top10df.initials[i] = top10df.user[i].split()[0][0] + top10df.user[i].split()[1][0]
top10df.initials[7] = "Me" # That's me
top10df.initials[8] = "DT"
```
### Starting with a basic plot.
- [Color References for Matplotlib](https://matplotlib.org/3.1.0/gallery/color/named_colors.html)
- **Improving Default Styles using Seaborn**
- [References](https://seaborn.pydata.org/generated/seaborn.set_style.html)
```
# For better readablity;
import matplotlib
matplotlib.rcParams['font.size'] = 14
matplotlib.rcParams['figure.figsize'] = (9, 5)
matplotlib.rcParams['figure.facecolor'] = '#00000000'
```
## *Now, I will be trying different visualization methods.*
##### Plotting a simple line graph.
```
# Improving Default Styles using Seaborn
sns.set_style("whitegrid")
# Increasing the figure size
plt.figure(figsize=(12, 6))
# plt.plot(top10df.initials, top10df.message, marker='o', ls='--', c='cyan')
# BETTER IMPLEMENTATION using the `fmt` argument;
plt.plot(top10df.initials, top10df.message, 'o--c')
# Labels and Title
plt.xlabel('Users')
plt.ylabel('Total number of messages')
plt.title("Number of messages sent by group members.")
plt.legend(['Messages']);
# Saving the plots
# plt.savefig('msg_plots.svg', format = 'svg')
```
#### *Plotting a bar chart and line graph together.*
```
# Improving Default Styles using Seaborn
sns.set_style("whitegrid")
# Increasing the figure size
plt.figure(figsize=(12, 6))
plt.bar(top10df.initials, top10df.message) # basic bar chart
plt.plot(top10df.initials, top10df.message, 'o--c'); # line chart
```
### *Beautifying Default Styles using Seaborn*
```
# Beautifying Default Styles using Seaborn
sns.set_style("darkgrid")
sns.barplot(top10df.initials, top10df.message, data=top10df);
```
# *Important Note*
- Since almost all the plots will be ***comparing* one person with another**, I’ll assign a **specific colour to each person** so that it becomes **easy to identify** each person among *multiple plots*.
- I could've used *seaborn's color palette* but:
- Seaborn assigns default colors itself, but i wanted the color of a **certain person remains the same, no matter the plot**;
- Also, I wanted to try some different colors so I grabbed my **colour palette** from https://coolors.co/
- Next, I made a dictionary where each key is the name and the value for each would be their assigned colour. I create a function which reorders colours given a list of names to match the ordering of the plot. This function takes the ordered names as input and returns a reordered list of colours. This list has to be passed into the `pallete` argument in a seaborn plotting function.
## Defining a function to tackle the problem.
I'm defining the following function to ***maintain consistent colors for each person across all plots***. Since the order will vary depending on the plot, this is passed to the function which will reorder colors in a particular order so that **the color of a certain person remains the same no matter the plot**. This will help maintain **consistency and readability** amongst the many graphs I will be plotting.
```
def get_colors_of_certain_order(names_in_certain_order):
'''the color of a certain person remains the same, no matter the plot'''
order = list(names_in_certain_order)
return_list = []
for name in order:
return_list.append(color_dict[name])
return return_list
```
### Now, we have a really nice set of colours for each person, with which we can visualize using `sns.palplot`.
```
colors = ['#F94144', '#F3722C', '#F8961E', '#FDC500', '#F9C74F', '#90BE6D', '#43AA8B', '#577590', '#6D597A','#003F88']
sns.palplot(colors) # visualizing the colors' list
names = top10df.initials
color_dict = {}
for name, color in zip(names, colors):
color_dict[name] = color
color_dict
```
## Now, finding the *average message length* of the 10 most active users of the group.
```
# Adding another column for message length; using the apply method;
df2['message_length'] = df2['message'].apply(lambda x: len(x))
# Creating another dataframe for average length per user;
avg_msg_lengths = df2.groupby(df2.user).mean().reset_index().sort_values(by = 'message_length', ascending = False)
# Creating helper columns;
top10df['avg_message_length'] = [0] * 10
i, j = 0, 0
while i < 10:
if top10df['user'][i] == avg_msg_lengths['user'][j]:
top10df['avg_message_length'][i] = avg_msg_lengths['message_length'][j]
i += 1
j = -1
j += 1
# Sorting the average message lengths of the same to 10 active users;
top10df_msg = top10df.sort_values(by = "avg_message_length", ascending=False)
```
## Now, we will be plotting *most sent messages* and respective *average message lengths* simultaneously, to see some interesting results.
- Plotting multiple charts in a grid
- Matplotlib and Seaborn also support plotting multiple charts in a grid, using `plt.subplots`, which returns a set of axes that can be used for plotting.
```
# plotting multiple charts in a grid
fig, axes = plt.subplots(1, 2, figsize=(16, 6))
sns.set_style("darkgrid")
# Plot 1 - Countplot of total messages sent
sns.barplot(top10df.initials, top10df.message, data=top10df, ax = axes[0], palette=get_colors_of_certain_order(top10df.initials)); # Note: the palette argument;
axes[0].set_title('Total Messages Sent ')
axes[0].set_xlabel('User')
axes[0].set_ylabel('Number of Messages Sent')
# Plot 2 - Barplot of those top 10 users' average message lengths
sns.barplot(top10df_msg.initials, top10df_msg.avg_message_length, ax = axes[1], palette = get_colors_of_certain_order(top10df_msg.initials)) # Note: the respective palette argument;
axes[1].set_title('Average Message Lengths')
axes[1].set_xlabel('User');
axes[1].set_ylabel('Average Messages Length');
# Saving the plots
plt.savefig('top10_msg_plots_diff.svg', format = 'svg')
```
### It’s really interesting to see plots like this *side by side*, because here comes the twist:
- Ironically, **TK**, the person who sent the **most amount of texts** (2000+), has ***least*** messages' length on *average*. This means this person sends broken and many WhatsApp messages in one go.
- We can see that, **I** send less number of messages while having a **relatively longer message length**, followed by **KS**.
- Here's a **snippet** of how TK sends messages:
<img src="https://imgur.com/xznZczG.png">
#### "*Things aren't always the way they seem like.*"
# 4. Top 10 users most sent media
- The exported chats were exported without any media files. Any message that contained media was indicated with `‘<Media Omitted> ’`.
### *Pre-processing*
**We can use this to filter out and see who sends the most media.**
```
# Using `groupby`, `count` and `sort_values` attributes.
top10media = df[df.message == '<Media omitted> '].groupby('user').count().sort_values(by="message", ascending = False).head(10)
# Dropping unused column;
top10media.drop(columns=['date_time', 'day', 'month', 'year', 'date'], inplace=True)
# Renaming column name for visualization;
top10media.rename(columns={"message": "media_sent"}, inplace=True)
# resetting index;
top10media.reset_index(inplace=True)
top10media['initials'] = ''
for i in range(10):
top10media.initials[i] = top10media.user[i].split()[0][0] + top10media.user[i].split()[1][0]
top10media.initials[2] = "Me" # That's me
top10media.initials[9] = "VR"
```
### *Visualization using different Seaborn's Color Palettes*
- [Seaborn References](http://seaborn.pydata.org/tutorial/color_palettes.html#using-circular-color-systems)
- [Seaborn's Different Colors](https://medium.com/@andykashyap/top-5-tricks-to-make-plots-look-better-9f6e687c1e08)
- [Seaborn's Color Visualization](https://python-graph-gallery.com/197-available-color-palettes-with-matplotlib/)
###### QUICK HACK
- to get **all possible Seaborn's color palettes**:
- Just put a random input `palette="xyz"`
- It will then show an error, showing all possible palettes you can try out from!
### Which user sends the most media?
```
# Increasing the figure size
plt.figure(figsize=(15, 6))
# Beautifying Default Styles using Seaborn
sns.set_style("darkgrid")
# Plotting a bar graph;
sns.barplot(top10media.initials, top10media.media_sent, palette="CMRmap");
plt.title('Most Sent Media')
plt.xlabel('User')
plt.ylabel('Total Media Sent');
# Saving the plots
plt.savefig('top10media.svg', format = 'svg')
```
### **TK** is beating everyone by a mile, followed by **DL**. He also ranks the **top** in total messages, *though **last** in average message length*. Most dedicated contributor award goes to **TK**!
# 5. Top 10 most used Emojis
- Will be using the `emoji` module, that was imported earlier.
```
emoji_ctr = Counter()
emojis_list = map(lambda x: ''.join(x.split()), emoji.UNICODE_EMOJI.keys())
r = re.compile('|'.join(re.escape(p) for p in emojis_list))
for idx, row in df.iterrows():
emojis_found = r.findall(row["message"])
for emoji_found in emojis_found:
emoji_ctr[emoji_found] += 1
```
#### Will create another helper column using `emoji.demojize("<emoji>")`, since emojis will not rendered.
```
top10emojis = pd.DataFrame()
# top10emojis = pd.DataFrame(data, columns={"emoji", "emoji_description", "emoji_count"})
top10emojis['emoji'] = [''] * 10
top10emojis['emoji_count'] = [0] * 10
top10emojis['emoji_description'] = [''] * 10
i = 0
for item in emoji_ctr.most_common(10):
# will be using another helper column, since during visualization, the emojis won't be rendered.
description = emoji.demojize(item[0])[1:-1] # using `[1:-1]` to remove the colons ':' at the end of the demojized strin
# appending top 10 data of emojis. # Loading into a DataFrame.
top10emojis.emoji[i] = item[0]
top10emojis.emoji_count[i] = int(item[1])
top10emojis.emoji_description[i] = description
i += 1
top10emojis
```
### Which Emoji is the most used in the chat?
```
# Increasing the figure size
plt.figure(figsize=(15, 6))
# Better Readablity
import matplotlib
matplotlib.rcParams['font.size'] = 15
# Beautifying Default Styles using Seaborn
sns.set_style("darkgrid")
# Plotting;
sns.barplot(top10emojis.emoji_count, top10emojis.emoji_description, palette = "Paired_r")
plt.title('Most Used Emoji')
plt.xlabel('Emoji Count')
plt.ylabel('Emoji Used');
# Saving the plots
plt.savefig('top10emoji.svg', format = 'svg')
```
#### Not that it is worth anything, but "😂" beats everyone by a huge margin!
# 6. Most active days, most active hours, most active months.
### *Pre-processing*
```
df3 = df.copy()
df3['message_count'] = [1] * df.shape[0] # helper column to keep a count.
df3['hour'] = df3['date_time'].apply(lambda x: x.hour)
grouped_by_time = df3.groupby('hour').sum().reset_index().sort_values(by = 'hour')
```
### Which hour of the day are most messages exchanged?
```
# Better Readablity
import matplotlib
matplotlib.rcParams['font.size'] = 16
matplotlib.rcParams['figure.figsize'] = (20, 8)
# Beautifying Default Styles using Seaborn
sns.set_style("darkgrid")
# PLOT: grouped by hour
sns.barplot(grouped_by_time.hour, grouped_by_time.message_count)
plt.title('Most Active Hours');
# Saving the plots;
plt.savefig('most_active_hours.svg', format = 'svg')
```
#### Intrestingly, the group is *most active around midnight*, followed by *afternoon*.
### *Pre-processing weekdays and months*
```
# specific `order` to be printed in;
days = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
# grouping by day;
grouped_by_day = df3.groupby('day').sum().reset_index()[['day', 'message_count']]
# specific `order` to be printed in;
months = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep'] # till Sept, since chats are till Septemeber
# grouping by month;
grouped_by_month = df3.groupby('month').sum().reset_index()[['month', 'message_count']]
```
## *Visualization*
### Now, we will be plotting *grouped by day* and respective *group by month* simultaneously, to see some interesting results.
- Using `plt.subplots` to plot multiple charts in a grid.
```
fig, axs = plt.subplots(1, 2, figsize = (24, 6))
# Better Readablity
import matplotlib
matplotlib.rcParams['font.size'] = 20
# Beautifying Default Styles using Seaborn
sns.set_style("darkgrid")
# Plotting;
# PLOT 1: Messages grouped by weekday
sns.barplot(grouped_by_day.day, grouped_by_day.message_count, order=days, ax = axs[0], palette='Pastel2_r')
axs[0].set_title('Total messages sent grouped by day')
# PLOT 2: Messages grouped by months
sns.barplot(y = grouped_by_month.month, x=grouped_by_month.message_count, order = months, ax = axs[1], palette='Pastel1_d')
axs[1].set_title('Total messages sent grouped by month');
# Saving the plots;
plt.savefig('days_and_month.svg', format = 'svg')
```
##### The group is most active on Sundays, and least active on Mondays (probably *Monday Blues*)
##### It has been recently very active, in September.
### Now, we will plot a *heatmap*, combining the above to bar plots, for a better understanding!
```
# Better Readablity
import matplotlib
matplotlib.rcParams['font.size'] = 14
matplotlib.rcParams['figure.figsize'] = (18, 6)
# Beautifying Default Styles using Seaborn,
sns.set_style("darkgrid")
# Pre-Processing by month and day,
grouped_by_month_and_day = df3.groupby(['month', 'day']).sum().reset_index()[['month', 'day', 'message_count']]
# creating a pivot table,
pt = grouped_by_month_and_day.pivot_table(index = 'month', columns = 'day', values = 'message_count').reindex(index = months, columns = days)
# PLOT: heatmap.
sns.heatmap(pt, cmap = 'cividis');
plt.title('Heatmap of Month sent and Day sent');
# Saving the plots;
plt.savefig('month_day_heatmap.svg', format = 'svg')
```
# *Inferences*
- The group is more active on weekends, throughout the months.
- September has the most lighter blue shades and more yellow gradients.
- This gives a combined analysis, which is really helpful in real-time projects.
# Most used words in the chat.
### Finally, I will be ending with *the most used words*, using `WordCloud` module in Python.
```
comment_words = ' '
# stopwords --> Words to be avoided while forming the WordCloud,
# removed group_notifications like 'joined', 'deleted';
# removed really common words like "yeah" and "okay".
stopwords = STOPWORDS.update(['group', 'link', 'invite', 'joined', 'message', 'deleted', 'yeah', 'hai', 'yes', 'okay', 'ok', 'will', 'use', 'using', 'one', 'know', 'guy', 'group', 'media', 'omitted'])
# iterate through the DataFrame.
for val in df3.message.values:
# typecaste each val to string.
val = str(val)
# split the value.
tokens = val.split()
# Converts each token into lowercase.
for i in range(len(tokens)):
tokens[i] = tokens[i].lower()
for words in tokens:
comment_words = comment_words + words + ' '
wordcloud = WordCloud(width = 600, height = 600,
background_color ='white',
stopwords = stopwords,
min_font_size = 8).generate(comment_words)
wordcloud.to_image()
```
# *Conclusion*
- The insights were really interesting to look at!
- We first loaded the data as a .txt file coverted it using `RawtoDF` function.
- Then we added helper columns, manipulated datetime entries.
- Then, we started analysing our whatsapp data!
Here is what we looked at!
**1. Overall frequency of total messages on the group.**
**2. Top 10 most active days.**
**3. Top 10 active users on the group (with a twist - Most active user had the least average message length ).**
- Ghosts present in the group. (shocking results - 80+ participants who haven't even sent a single message!)
**4. Top 10 users most sent media.**
- *TK* beats everyone by a mile!
**5. Top 10 most used emojis.**
- using the `emoji` module!
**6. Most active hours and weekdays.**
- Heatmaps of weekdays and months.
- Most active hours, weekdays, and months.
**7. Most used words - WordCloud**
### That's it from my end! I hope you *learnt and enjoyed* a lot!
# *Where to go from here?*
- Extending this **Whatsapp Ananlysis Project**!
- Adding a **Sentiment Analyser to the texts** - Swear Words & Sentiments!
- Libraries called `profanity_check`, `TextBlob`
- VADER (Valence Aware Dictionary and Sentiment Reasoner) from NTLK and TextBlob.
- Instagram Data Analysis
- Play Store Data Analysis
- Exploring more datasets! It is *never-ending*, as there can be ***Infinite Stories with Data***!
- Once, done with enough Data Analysis, try making Datasets, scraping Data from websites and creating Interesting Datasets!
# *Thank You*
- [Tushar Nankani](https://www.linkedin.com/in/tusharnankani/)
| github_jupyter |
# Machine Learning untuk Aplikasi Geospasial
Bagian kedua dari tiga seri Geospatial Data Crash Course ini akan membahas contoh-contoh aplikasi Machine Learning untuk bidang Geospasial. Pada [bagian pertama](https://colab.research.google.com/drive/1MyjuxE2O-3YX1Q0m6EMA6xrKeum6sETE?usp=sharing) telah dibahas mengenai dasar Geospatial Data Science dan penggunaan Python untuk keperluan pengolahan data spasial. Bagian kedua ini membahas mengenai *machine learning* yang diterapkan untuk beberapa contoh kasus pada bidang Geospasial.
## Pengantar
Machine learning adalah *'buzzword of the century'*. Beragam aplikasi yang kita saksikan sehari-hari saat ini adalah bagian dari produk yang dihasilkan oleh machine learning. Machine learning merupakan bagian dari bidang Artificial Intelligence yang memberikan komputer kemampuan untuk belajar dari data yang dimasukkan kepada mesin untuk dipelajari. Selanjutnya, mesin dapat menentukan pola dan memuat jawaban dari pertanyaan yang diajukan.

Machine learning saat ini diterapkan pada hampir semua bidang, termasuk pada bidang geospasial. Pada notebook ini akan diberikan contoh mengenai bagaimana machine learning dapat digunakan dalam keperluan analisis spasial, khususnya ketika mengolah data dalam jumlah besar.
## Studi Kasus
Pada bagian ini akan diberikan beberapa Contoh studi kasus untuk dilakukan. Penjelasan untuk tiap kode dapat dijumpai sebagai komentar maupun baris paragraf yang diberikan sebelumnya.
### Teachable Machine
Teachable Machine merupakan antarmuka untuk mesin Machine Learning yang dibuat oleh Google untuk memudahkan pemahaman mengenai cara kerja Machine Learning. Di belakang layar, Teachable Machine menggunakan Tensorflow untuk membuat model dari data yang dimasukkan oleh pengguna.
Pada latihan berikut kita akan menggunakan Teachable Machine untuk melatin mesin ini mengenali beberapa kelas data yang berbeda. Prosedurnya cukup mudah:
1. Unduh data untuk latihan klasifikasi. Anda dapat menggunakan data berikut:
> https://drive.google.com/file/d/1fgtwpGIh5lWv0H4FOwmFm_dtdURqSzax/view?usp=sharing
2. Buka website Teachable Machine: https://teachablemachine.withgoogle.com/
3. Buat kelas. Kelas-kelas ini dibuat sesuai dengan jenis hewan, misalnya ayam, sapi, dst.
4. Lakukan training untuk membuat model.
5. Gunakan gambar baru untuk melakukan prediksi. Misalnya gambar seperti berikut:
{}(https://static.dw.com/image/47863948_303.jpg)
Lihat apakah mesin berhasil mendeteksi objek tersebut sebagai seekor sapi.
### Menggunakan Scikit-Learn
Scikit-Learn merupakan modul Python yang digunakan untuk keperluan pemodelan, analisis statistik serta pembuatan machine learning. Scikit-Learn memiliki modul yang sangat lengkap untuk keperluan machine learning, sekaligus menyediakan kemampuan untuk melakukan modifikasi dan tuning parameter model machine learning yang dikembangkan sendiri.
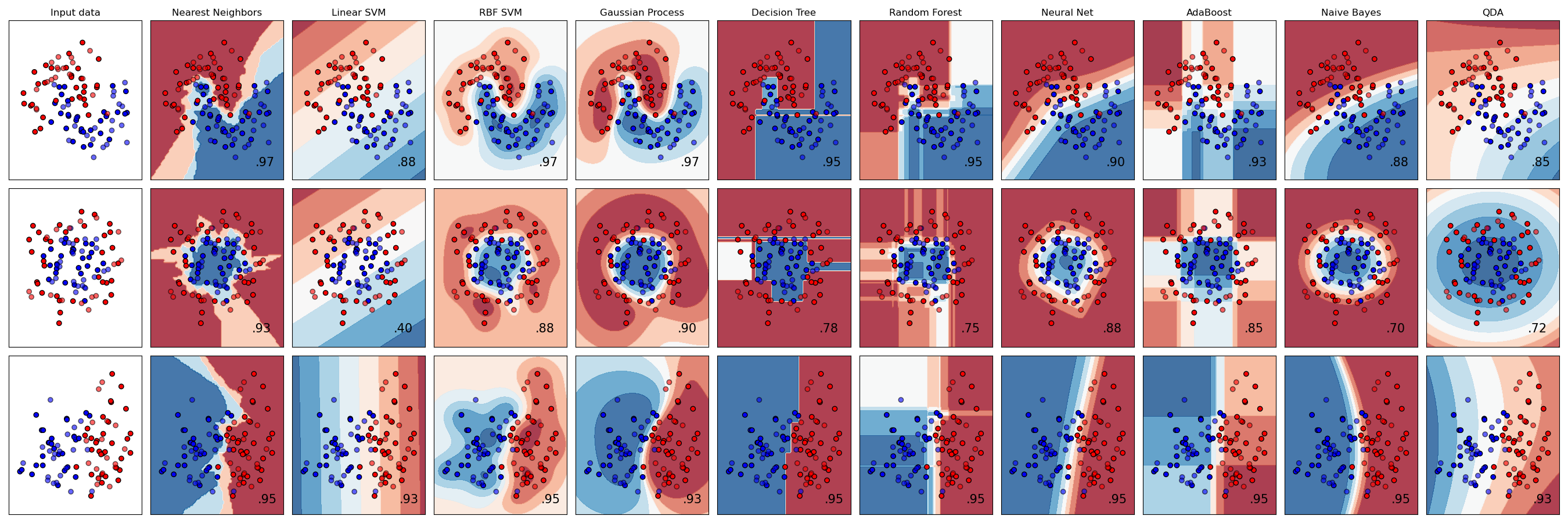
Penjelasan mengenai Scikit Learn sendiri dapat dijumpai lebih lengkap pada dokumentasi yang tersedia. Pada contoh di bawah ini, kita akan melakukan klasifikasi sederhana menggunakan classifier MeanShift. Detilnya dapat dilihat pada:
https://scikit-learn.org/stable/auto_examples/cluster/plot_mean_shift.html
```
import numpy as np
from sklearn.cluster import MeanShift, estimate_bandwidth
from sklearn.datasets import make_blobs
```
Pertama, kita akan buat data random sebagai masukan untuk klasifikasi. Data ini dapat dibuat menggunakan fungsi `make_blobs` pada scikit learn, seperti berikut:
```
# membuat data sampel
centers = [[1, 1], [-1, -1], [1, -1]]
X, _ = make_blobs(n_samples=10000, centers=centers, cluster_std=0.6)
```
Pada data di atas dapat kita lihat bahwa kita meminta scikit learn untuk memuat 10000 data dengan standar deviasi 0.6. Kita dapat mengganti data ini dengan data lainnya, misalnya data covid yang kita gunakan di minggu sebelumnya.
MeanShift merupakan salah satu metode untuk melakukan klasifikasi dari serangkaian data menggunakan metode unsupervised learning. Terlebih dahulu kita perlu membuat bandwith yang menyatakan nilai kuantil dan besaran sampel yang akan digunakan pada tiap klasifikasi, seperti berikut:
```
# menghitung clustering dengan MeanShift
# Menghitung Bandwith
bandwidth = estimate_bandwidth(X, quantile=0.2, n_samples=500)
# unsupervised machine learning
ms = MeanShift(bandwidth=bandwidth, bin_seeding=True)
ms.fit(X)
labels = ms.labels_
cluster_centers = ms.cluster_centers_
labels_unique = np.unique(labels)
n_clusters_ = len(labels_unique)
print("number of estimated clusters : %d" % n_clusters_)
```
Selanjutnya, kita dapat mencoba menampilkan hasil klustering yang kita buat. Kita gunakan Matplotlib untuk keperluan plotting dan melihat bagaimana hasil klasifikasi yang dibuat oleh SKLearn.
```
import matplotlib.pyplot as plt
from itertools import cycle
plt.figure(1)
plt.clf()
colors = cycle('bgrcmykbgrcmykbgrcmykbgrcmyk')
for k, col in zip(range(n_clusters_), colors):
my_members = labels == k
cluster_center = cluster_centers[k]
plt.plot(X[my_members, 0], X[my_members, 1], col + '.')
plt.plot(cluster_center[0], cluster_center[1], 'o', markerfacecolor=col,
markeredgecolor='k', markersize=14)
plt.title('Estimasi jumlah kluster: %d' % n_clusters_)
plt.show()
```
Dari contoh di atas, kita dapat gunakan model klasifikasi unsupervised ini pada berbagai data lain. Klasifikasi jenis ini banyak digunakan sebagai data masukan pada klasifikasi supervised.
**Sebagai latihan**, gunakan metode MeanShift di atas untuk membuat klasifikasi dari berbagai data lain yang dapat Anda temukan, misalnya data Covid atau data gempa bumi dari latihan sebelumnya.
```
```
### Melakukan Analisis Spasial dengan Machine Learning
Pada latihan ini, kita akan lakukan contoh analisis spasial yang sederhana dengan menggunakan Machine Learning, khususnya modul Scikit-Learn pada Python. Kita akan gunakan metode [Ball Tree](https://towardsdatascience.com/tree-algorithms-explained-ball-tree-algorithm-vs-kd-tree-vs-brute-force-9746debcd940) yang merupakan salah satu algoritma Nearest Neighbor untuk analisis titik terdekat.
Pada latihan ini kita akan banyak menggunakan modul Python dengan nama [OSMNX](https://osmnx.readthedocs.io/en/stable/osmnx.html). Modul ini digunakan untuk berinteraksi dengan data OSM agar dapat langsung digunakan dan dianalisis di Python. Untuk itu kita perlu melakukan instalasi modul ini terlebih dahulu.
> Analisisnya: Carilah bangunan (rumah) yang paling dekat dengan titik-titik sekolah di Kota Sleman
```
!pip install osmnx
```
Selanjutnya, kita akan gunakan modul ini untuk memanggil titik-titik POI dari OSM ke dalam Google Colab:
```
# memanggil modul
import osmnx as ox
all_pois = ox.geometries_from_place(
'Sleman, ID',
tags = {'amenity':True})
all_pois['amenity'].value_counts()
```
Dari data di atas, kita akan coba panggil titik-titik sekolah:
```
# Run query
sekolah = ox.geometries_from_place(
'Sleman, ID',
tags = {'amenity':'school'},
which_result=1)
sekolah.head()
# berapa jumlah sekolah di Sleman?
sekolah['osmid'].count()
```
Demikian pula, kita panggil bangunan. Kali ini kita panggil sebagai poligon:
```
bangunan_gdf = ox.geometries_from_place(
'Sleman, ID',
tags = {'building':True})
bangunan_gdf.head()
```
Berapa jumlahnya?
```
bangunan_gdf['osmid'].count()
```
Terdapat 29.880 bangunan yang kita peroleh. Kita coba tampilkan data tersebut, bagaimana distribusi datanya?
```
# Download London's Boundary
sleman = ox.geocoder.geocode_to_gdf('Sleman, ID')
# Set up a plot axis
fig, ax = plt.subplots(figsize = (15,10))
# Visualise both on the plot
sleman.plot(ax = ax, alpha = 0.5)
sekolah.plot(ax = ax, markersize = 8, color = 'red', alpha = 0.8, label = 'Lokasi Sekolah')
bangunan_gdf.plot(ax = ax, markersize = 1, color = 'blue', alpha = 0.8, label = 'Lokasi Bangunan')
plt.legend()
```
Saatnya melakukan analisis Machine Learning. Fungsi berikut kita gunakan untuk melakukan analisis Nearest Neighbor pada titik bangunan dan sekolah:
```
from sklearn.neighbors import BallTree
import numpy as np
def get_nearest(src_points, candidates, k_neighbors=1):
"""Menentukan NN dari dua"""
# Create tree from the candidate points
tree = BallTree(candidates, leaf_size=15, metric='haversine')
# Find closest points and distances
distances, indices = tree.query(src_points, k=k_neighbors)
# Transpose to get distances and indices into arrays
distances = distances.transpose()
indices = indices.transpose()
# Get closest indices and distances (i.e. array at index 0)
# note: for the second closest points, you would take index 1, etc.
closest = indices[0]
closest_dist = distances[0]
# Return indices and distances
return (closest, closest_dist)
def nearest_neighbor(left_gdf, right_gdf, return_dist=False):
"""
For each point in left_gdf, find closest point in right GeoDataFrame and return them.
NOTICE: Assumes that the input Points are in WGS84 projection (lat/lon).
"""
left_geom_col = left_gdf.geometry.name
right_geom_col = right_gdf.geometry.name
# Ensure that index in right gdf is formed of sequential numbers
right = right_gdf.copy().reset_index(drop=True)
# Parse coordinates from points and insert them into a numpy array as RADIANS
left_radians = np.array(left_gdf[left_geom_col].apply(lambda geom: (geom.centroid.x * np.pi / 180, geom.centroid.y * np.pi / 180)).to_list())
right_radians = np.array(right[right_geom_col].apply(lambda geom: (geom.centroid.x * np.pi / 180, geom.centroid.y * np.pi / 180)).to_list())
# Find the nearest points
# -----------------------
# closest ==> index in right_gdf that corresponds to the closest point
# dist ==> distance between the nearest neighbors (in meters)
closest, dist = get_nearest(src_points=left_radians, candidates=right_radians)
# Return points from right GeoDataFrame that are closest to points in left GeoDataFrame
closest_points = right.loc[closest]
# Ensure that the index corresponds the one in left_gdf
closest_points = closest_points.reset_index(drop=True)
# Add distance if requested
# --ini adalah haversine distance--
if return_dist:
# Convert to meters from radians
earth_radius = 6371000 # meters
closest_points['distance'] = dist * earth_radius
return closest_points
```
Kita coba tampilkan waktu yang digunakan oleh mesin ini untuk menganalisis menggunakan fungsi di atas. Perintah `%%time` digunakan untuk keperluan ini
```
# mulai analisis. berapa waktunya?
%%time
closest_points = nearest_neighbor(bangunan_gdf, sekolah, return_dist=True)
```
Hasilnya?
```
# Melihat hasilnya:
closest_points
```
Scroll ke kanan untuk melihat bahwa tiap data telah dihitung jaraknya, antara sekolah dan bangunan di Sleman. Kita bisa cek apakah seluruh titik sudah dihitung:
```
print(len(closest_points), '==', len(bangunan_gdf))
```
Gabungkan hasil ini menggunakan Left Join. Geopandas dapat kita gunakan untuk keperluan ini:
```
# Rename the geometry of closest stops gdf so that we can easily identify it
closest_points = closest_points.rename(columns={'geom': 'closest_stop_geom'})
# Merge the datasets by index (for this, it is good to use '.join()' -function)
bangunan_gdf = bangunan_gdf.join(closest_points, how = 'left', lsuffix = '_left', rsuffix = '_right')
# Let's see what we have
bangunan_gdf.head()
```
Kita juga dapat menerapkan statistik sederhana untuk menghitung berbagai parameter:
```
bangunan_gdf['distance'].describe()
```
Terakhir, kita akan gunakan beberapa modul untuk menampilkan klasifikasi data. Mapclassify digunakan untuk keperluan ini, sehingga perlu kita install terlebih dahulu:
```
## install mapclassify
!pip install mapclassify
```
Kemudian, kita tampilkan seperti berikut:
```
closest_points.plot(column='distance', markersize=0.2, alpha=0.5, figsize=(10,10), scheme='quantiles', k=4,
legend=True)
```
Dengan demikian kita telah melakukan analisis untuk melihat bagaimana machine learning dapat kita gunakan untuk melakukan analisis pencarian lokasi dengan lebih mudah dan lebih cepat.
**Sebagai Latihan**, lakukan analisis yang sama pada berbagai data yang berbeda: di kota lain dengan jenis 'amenities' OSM yang berbeda. Bagaimana hasilnya?
## Rujukan
* https://towardsdatascience.com/exploring-classifiers-with-python-scikit-learn-iris-dataset-2bcb490d2e1b
* https://medium.com/spatial-data-science/deep-learning-for-geospatial-data-applications-multi-label-classification-2b0a1838fcf3
* https://automating-gis-processes.github.io/site/notebooks/L3/nearest-neighbor-faster.html
* https://colab.research.google.com/github/sentinel-hub/eo-learn/blob/master/examples/land-cover-map/SI_LULC_pipeline.ipynb
```
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.