text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
## RIHAD VARIAWA, Data Scientist - Who has fun LEARNING, EXPLORING & GROWING
In workspaces like this one, you will be able to practice visualization techniques you've seen in the course materials. In this particular workspace, you'll practice creating single-variable plots for categorical data.
```
# prerequisite package imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
%matplotlib inline
# solution script imports
from solutions_univ import bar_chart_solution_1, bar_chart_solution_2
```
In this workspace, you'll be working with this dataset comprised of attributes of creatures in the video game series Pokémon. The data was assembled from the database of information found in [this GitHub repository](https://github.com/veekun/pokedex/tree/master/pokedex/data/csv).
```
pokemon = pd.read_csv('./data/pokemon.csv')
pokemon.head()
```
**Task 1**: There have been quite a few Pokémon introduced over the series' history. How many were introduced in each generation? Create a _bar chart_ of these frequencies using the 'generation_id' column.
```
base_color = sb.color_palette()[0]
sb.countplot(data = pokemon, x = 'generation_id', color = base_color);
```
Once you've created your chart, run the cell below to check the output from our solution. Your visualization does not need to be exactly the same as ours, but it should be able to come up with the same conclusions.
```
bar_chart_solution_1()
```
**Task 2**: Each Pokémon species has one or two 'types' that play a part in its offensive and defensive capabilities. How frequent is each type? The code below creates a new dataframe that puts all of the type counts in a single column.
```
pkmn_types = pokemon.melt(id_vars = ['id','species'],
value_vars = ['type_1', 'type_2'],
var_name = 'type_level', value_name = 'type').dropna()
pkmn_types.head()
```
Your task is to use this dataframe to create a _relative frequency_ plot of the proportion of Pokémon with each type, _sorted_ from most frequent to least. **Hint**: The sum across bars should be greater than 100%, since many Pokémon have two types. Keep this in mind when considering a denominator to compute relative frequencies.
```
pkmn_cnts = pkmn_types['type'].value_counts(sort=True)
pkmn_labels = pkmn_cnts.index
pkmn_max = max(pkmn_cnts) / pokemon.shape[0]
ticks=np.arange(0,pkmn_max,.02)
ticks_names = ['{:.2f}'.format(x) for x in ticks]
sb.countplot(data = pkmn_types, y = 'type',color = base_color, order = pkmn_labels);
plt.xticks(ticks * pokemon.shape[0],ticks_names);
bar_chart_solution_2()
```
If you're interested in seeing the code used to generate the solution plots, you can find it in the `solutions_univ.py` script in the workspace folder. You can navigate there by clicking on the Jupyter icon in the upper left corner of the workspace. Spoiler warning: the script contains solutions for all of the workspace exercises in this lesson, so take care not to spoil your practice!
| github_jupyter |
```
import panel as pn
import numpy as np
import pandas as pd
pn.extension()
```
The ``Bokeh`` pane allows displaying any displayable [Bokeh](http://bokeh.org) model inside a Panel app. Since Panel is built on Bokeh internally, the Bokeh model is simply inserted into the plot. Since Bokeh models are ordinarily only displayed once, some Panel-related functionality such as syncing multiple views of the same model may not work. Nonetheless this pane type is very useful for combining raw Bokeh code with the higher-level Panel API.
When working in a notebook any changes to a Bokeh objects may not be synced automatically requiring an explicit call to `pn.state.push_notebook` with the Panel component containing the Bokeh object.
#### Parameters:
For the ``theme`` parameter, see the [bokeh themes docs](https://docs.bokeh.org/en/latest/docs/reference/themes.html).
* **``object``** (bokeh.layouts.LayoutDOM): The Bokeh model to be displayed
* **``theme``** (bokeh.themes.Theme): The Bokeh theme to apply
___
```
from math import pi
from bokeh.palettes import Category20c, Category20
from bokeh.plotting import figure
from bokeh.transform import cumsum
x = {
'United States': 157,
'United Kingdom': 93,
'Japan': 89,
'China': 63,
'Germany': 44,
'India': 42,
'Italy': 40,
'Australia': 35,
'Brazil': 32,
'France': 31,
'Taiwan': 31,
'Spain': 29
}
data = pd.Series(x).reset_index(name='value').rename(columns={'index':'country'})
data['angle'] = data['value']/data['value'].sum() * 2*pi
data['color'] = Category20c[len(x)]
p = figure(plot_height=350, title="Pie Chart", toolbar_location=None,
tools="hover", tooltips="@country: @value", x_range=(-0.5, 1.0))
r = p.wedge(x=0, y=1, radius=0.4,
start_angle=cumsum('angle', include_zero=True), end_angle=cumsum('angle'),
line_color="white", fill_color='color', legend_field='country', source=data)
p.axis.axis_label=None
p.axis.visible=False
p.grid.grid_line_color = None
bokeh_pane = pn.pane.Bokeh(p, theme="dark_minimal")
bokeh_pane
```
To update a plot with a live server, we can simply modify the underlying model. If we are working in a Jupyter notebook we also have to call the `pn.io.push_notebook` helper function on the component or explicitly trigger an event with `bokeh_pane.param.trigger('object')`:
```
r.data_source.data['color'] = Category20[len(x)]
pn.io.push_notebook(bokeh_pane)
```
Alternatively the model may also be replaced entirely, in a live server:
```
from bokeh.models import Div
bokeh_pane.object = Div(text='<h2>This text replaced the pie chart</h2>')
```
The other nice feature when using Panel to render bokeh objects is that callbacks will work just like they would on the server. So you can simply wrap your existing bokeh app in Panel and it will render and work out of the box both in the notebook and when served as a standalone app:
```
from bokeh.layouts import column, row
from bokeh.models import ColumnDataSource, Slider, TextInput
# Set up data
N = 200
x = np.linspace(0, 4*np.pi, N)
y = np.sin(x)
source = ColumnDataSource(data=dict(x=x, y=y))
# Set up plot
plot = figure(plot_height=400, plot_width=400, title="my sine wave",
tools="crosshair,pan,reset,save,wheel_zoom",
x_range=[0, 4*np.pi], y_range=[-2.5, 2.5])
plot.line('x', 'y', source=source, line_width=3, line_alpha=0.6)
# Set up widgets
text = TextInput(title="title", value='my sine wave')
offset = Slider(title="offset", value=0.0, start=-5.0, end=5.0, step=0.1)
amplitude = Slider(title="amplitude", value=1.0, start=-5.0, end=5.0, step=0.1)
phase = Slider(title="phase", value=0.0, start=0.0, end=2*np.pi)
freq = Slider(title="frequency", value=1.0, start=0.1, end=5.1, step=0.1)
# Set up callbacks
def update_title(attrname, old, new):
plot.title.text = text.value
text.on_change('value', update_title)
def update_data(attrname, old, new):
# Get the current slider values
a = amplitude.value
b = offset.value
w = phase.value
k = freq.value
# Generate the new curve
x = np.linspace(0, 4*np.pi, N)
y = a*np.sin(k*x + w) + b
source.data = dict(x=x, y=y)
for w in [offset, amplitude, phase, freq]:
w.on_change('value', update_data)
# Set up layouts and add to document
inputs = column(text, offset, amplitude, phase, freq)
bokeh_app = pn.pane.Bokeh(row(inputs, plot, width=800))
bokeh_app
```
### Controls
The `Bokeh` pane exposes a number of options which can be changed from both Python and Javascript. Try out the effect of these parameters interactively:
```
pn.Row(bokeh_app.controls(jslink=True), bokeh_app)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/finlytics-hub/LTV_predictions/blob/master/LTV_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Environment Setup
```
# import and install all required libraries
import pandas as pd
import seaborn as sns
import datetime as dt
import matplotlib.pyplot as plt
import numpy as np
! pip install lifetimes==0.11.3
from lifetimes.plotting import *
from lifetimes.utils import *
from lifetimes import BetaGeoFitter
from lifetimes import GammaGammaFitter
# Load data
data = pd.read_csv('.../OnlineRetail_2yrs.csv')
```
# EDA and Data Preprocessing
```
data.head(10)
data.info()
# Convert InvoiceDate into DateTime format and extract the date values
data['InvoiceDate'] = pd.to_datetime(data['InvoiceDate']).dt.date
# drop rows with missing CustomerID as our analysis will be at the individual customer level
data.dropna(axis = 0, subset = ['Customer ID'], inplace = True)
# filter out the negative values from Quantity field as these could relate to returns that are not relevant to LTV predictions
data = data[(data['Quantity'] > 0)]
# create a new column for Sales per invoice and filter out only the required columns for the Lifetimes package
data['Sales'] = data['Quantity'] * data['Price']
data_final = data[['Customer ID', 'InvoiceDate', 'Sales']]
data_final.head()
data_final.info()
# transform our transaction level data into the required summary form for Lifetimes
data_summary = summary_data_from_transaction_data(data_final, customer_id_col = 'Customer ID', datetime_col = 'InvoiceDate', monetary_value_col = 'Sales', freq = 'D')
# used freq = 'D' since we have a daily transactions log
data_summary.head(10)
# retain only those customers with frequency > 0
data_summary = data_summary[data_summary['frequency'] > 0]
# Some descriptive statistics of the summary data
data_summary.describe()
```
# BG/NBD Model Training & Visualisation
```
# fit the BG/NBD model to our data_summary
bgf = BetaGeoFitter()
bgf.fit(data_summary['frequency'], data_summary['recency'], data_summary['T'])
# plot the estimated gamma distribution of λ (customers' propensities to purchase)
plot_transaction_rate_heterogeneity(bgf);
# plot the estimated beta distribution of p, a customers' probability of dropping out immediately after a transaction
plot_dropout_rate_heterogeneity(bgf);
# visualize our frequency/recency matrix
fig = plt.figure(figsize=(12,8))
plot_frequency_recency_matrix(bgf, T = 7);
# Now let's visualise the probability of a customer being alive
fig = plt.figure(figsize=(12,8))
plot_probability_alive_matrix(bgf);
```
# Model Validation
```
# partition the dataset into a calibration and a holdout dataset
summary_cal_holdout = calibration_and_holdout_data(data_final, 'Customer ID', 'InvoiceDate', freq = "D", monetary_value_col = 'Sales', calibration_period_end='2011-06-30')
summary_cal_holdout.head()
# again, retain only the +ve frequency_cal values
summary_cal_holdout = summary_cal_holdout[summary_cal_holdout['frequency_cal'] > 0]
# compare the predicted # of repeat puchases with actual repeat purchases during the holdout period
bgf_cal = BetaGeoFitter()
bgf_cal.fit(summary_cal_holdout['frequency_cal'], summary_cal_holdout['recency_cal'], summary_cal_holdout['T_cal'])
plot_calibration_purchases_vs_holdout_purchases(bgf_cal, summary_cal_holdout, kind = 'frequency_cal', n = int(summary_cal_holdout['frequency_holdout'].max()), figsize = (12,8));
```
# Gamma-Gamma Model Fitting
```
# Let's fit the Gamma-Gamma model to our data_summary
ggf = GammaGammaFitter()
ggf.fit(frequency = data_summary['frequency'], monetary_value = data_summary['monetary_value'])
```
# Prediction Time!
```
# Calculate the expected number of repeat purchases up to time t for a randomly chosen individual from the population
t = 30 # to calculate the number of expected repeat purchases over the next 30 days
data_summary['predicted_purchases'] = bgf.conditional_expected_number_of_purchases_up_to_time(t, data_summary['frequency'], data_summary['recency'], data_summary['T'])
data_summary.sort_values(by='predicted_purchases').tail(5)
# Calculate probability of being currently alive and assign to each CustomerID
data_summary['p_alive'] = bgf.conditional_probability_alive(data_summary['frequency'], data_summary['recency'], data_summary['T'])
data_summary.sort_values(by='predicted_purchases').tail(5)
sns.distplot(data_summary['p_alive']);
data_summary['churn'] = ['churned' if p_alive < 0.5 else 'not churned' for p_alive in data_summary['p_alive']]
data_summary['churn'][(data_summary['p_alive'] >= 0.5) & (data_summary['p_alive'] < 0.75)] = "high risk"
data_summary['churn'].value_counts()
# After applying Gamma-Gamma model, now we can estimate average transaction value for each customer over his/her lifetime
data_summary['predicted_Sales'] = ggf.conditional_expected_average_profit(data_summary['frequency'], data_summary['monetary_value'])
data_summary.head()
# Final piece of the puzzle - calculate LTV for each customer over the next 12 months with an assumed monthly discount rate of 0.01%
data_summary['LTV'] = ggf.customer_lifetime_value(
bgf, #the model to use to predict the number of future transactions
data_summary['frequency'], data_summary['recency'], data_summary['T'], data_summary['monetary_value'],
time = 12, # number of months to predict LTV for
discount_rate = 0.01 # monthly discount rate ~ 12.7% annually
)
data_summary.head()
# Let's identify our top 20 customers based on LTV
best_projected_cust_LTV = data_summary.sort_values('LTV').tail(20)
best_projected_cust_LTV
```
| github_jupyter |
```
#hide
from nbdev.showdoc import *
```
# 1 - SC2 Training Grounds Computer Model
In this deliverable, I focus on developing the computer model of _SC2 Training Grounds_. This model is based on the application's conceptual model defined in the project's design brief.
> Note: see this project's dissertation document (Chapter 2, Human-Computer Interaction section). There, I explore the idea of design as the process of connecting an artifact's computer and user models.
To this end, I first use the activity diagrams shown below, which illustrates the application's three primary processes. I separate the processes, given that they are meant to be triggered by different events and run parallel to each other. However, they also interact with each other in different ways at different moments.
## Primary User Interaction
First, the following diagram shows the actions that define the users' primary interaction with the application. Through these actions, the application responds to two goals: handling the player profile classification process and offering recommendations based on the progress and results of that process.
The profile classification process, in turn, has three goals:
1. Check if the player's profile has been classified into one of the categories built in the application's clustering process (see Bellow).
2. Remind and update the player about their progress in this process.
3. Trigger the reclassification process once a player completes a training regime.
> Note: The process proposed in the diagram asks players to play five classification online matches every time a season starts or when they complete a training regime. I settled on five matches since it's the number of matches that the game already asks players to complete to rank them in the online competitive ladders every season.
Meanwhile, _SC2 Training Grounds_ centres its recommendation process on two tasks. First, it maintains a set of similarity matrixes that it uses to generate recommendations based on Item-to-Item and User-to-User comparisons. Second, it determines what recommendations to offer to the players based on their classification and current training regime status.
It maintains the similarity matrixes by assigning positive or negative points to recommendations similarly to how other recommender systems award points through ranking or rating systems. However, in this case, the system rewards regimes and challenges chosen by the players, to regimes carried out to completion and regimes that elicit profile changes according to the reclassification. Meanwhile, the process punishes regimes that players abandon.
> Note: see dissertation document Section 4.3 for a review of collaborative filtering recommendation systems.
<img alt="Primary UX Diagram" caption="Sc2 Training Grouns' Primary User Interaction Diagram" id="interaction_diagram" height="900" src="images/Final_Activity_diagram.png">
## Clustering and Classification Model
The next diagram shows the _clustering_ and _classification model training_ process that runs parallel and in the background of the users' interaction.
This process is conceptualised to match StarCraft 2's ranked ladder season cycles. In other words, since players are being asked to get ranked every season, I propose that _SC2 Training Grounds_ recalculates its classification clusters at that same pace.
In this case, the clustering is meant as a form of **Player Experience Modelling** (see Dissertation Document Chapters 2.1.2, 2.3.5 and 3.3.3). Here, the system responds to the changing tactics and strategies of the game to classify players dividing players into clusters based on their performance indicators every season. In addition, these cluster features can also be used as a source of information to communicate the state of the population's characteristics (see Dissertation Document Chapters 3.3.6).
Clustering also tags the replay batch it uses to create its classification. These tags can then be used to train an efficient classification model that can be used during the Primary Interaction Loop described above.
Since all other processes in the system are contingent upon this classification process, I develop the components that run this process in Section 1 of this deliverable.
<img alt="Clustering process diagram" caption="Clustering and classification model training process diagram" width="300" id="cluster_diagram" src="images/Clustering_diagram.png">
## Collaborative Filtering
> Warning: To be contimnued
| github_jupyter |
# Analyzing the flexible path length simulation
```
from __future__ import print_function
%matplotlib inline
import openpathsampling as paths
import numpy as np
import matplotlib.pyplot as plt
import os
import openpathsampling.visualize as ops_vis
from IPython.display import SVG
```
Load the file, and from the file pull our the engine (which tells us what the timestep was) and the move scheme (which gives us a starting point for much of the analysis).
```
# note that this log will overwrite the log from the previous notebook
#import logging.config
#logging.config.fileConfig("logging.conf", disable_existing_loggers=False)
%%time
flexible = paths.AnalysisStorage("ad_tps.nc")
# opening as AnalysisStorage is a little slower, but speeds up the move_summary
engine = flexible.engines[0]
flex_scheme = flexible.schemes[0]
print("File size: {0} for {1} steps, {2} snapshots".format(
flexible.file_size_str,
len(flexible.steps),
len(flexible.snapshots)
))
```
That tell us a little about the file we're dealing with. Now we'll start analyzing the contents of that file. We used a very simple move scheme (only shooting), so the main information that the `move_summary` gives us is the acceptance of the only kind of move in that scheme. See the MSTIS examples for more complicated move schemes, where you want to make sure that frequency at which the move runs is close to what was expected.
```
flex_scheme.move_summary(flexible.steps)
```
## Replica history tree and decorrelated trajectories
The `ReplicaHistoryTree` object gives us both the history tree (often called the "move tree") and the number of decorrelated trajectories.
A `ReplicaHistoryTree` is made for a certain set of Monte Carlo steps. First, we make a tree of only the first 25 steps in order to visualize it. (All 10000 steps would be unwieldy.)
After the visualization, we make a second `PathTree` of all the steps. We won't visualize that; instead we use it to count the number of decorrelated trajectories.
```
replica_history = ops_vis.ReplicaEvolution(replica=0)
tree = ops_vis.PathTree(
flexible.steps[0:25],
replica_history
)
tree.options.css['scale_x'] = 3
SVG(tree.svg())
# can write to svg file and open with programs that can read SVG
with open("flex_tps_tree.svg", 'w') as f:
f.write(tree.svg())
print("Decorrelated trajectories:", len(tree.generator.decorrelated_trajectories))
%%time
full_history = ops_vis.PathTree(
flexible.steps,
ops_vis.ReplicaEvolution(
replica=0
)
)
n_decorrelated = len(full_history.generator.decorrelated_trajectories)
print("All decorrelated trajectories:", n_decorrelated)
```
## Path length distribution
Flexible length TPS gives a distribution of path lengths. Here we calculate the length of every accepted trajectory, then histogram those lengths, and calculate the maximum and average path lengths.
We also use `engine.snapshot_timestep` to convert the count of frames to time, including correct units.
```
path_lengths = [len(step.active[0].trajectory) for step in flexible.steps]
plt.hist(path_lengths, bins=40, alpha=0.5);
print("Maximum:", max(path_lengths),
"("+(max(path_lengths)*engine.snapshot_timestep).format("%.3f")+")")
print ("Average:", "{0:.2f}".format(np.mean(path_lengths)),
"("+(np.mean(path_lengths)*engine.snapshot_timestep).format("%.3f")+")")
```
## Path density histogram
Next we will create a path density histogram. Calculating the histogram itself is quite easy: first we reload the collective variables we want to plot it in (we choose the phi and psi angles). Then we create the empty path density histogram, by telling it which CVs to use and how to make the histogram (bin sizes, etc). Finally, we build the histogram by giving it the list of active trajectories to histogram.
```
from openpathsampling.numerics import HistogramPlotter2D
psi = flexible.cvs['psi']
phi = flexible.cvs['phi']
deg = 180.0 / np.pi
path_density = paths.PathDensityHistogram(cvs=[phi, psi],
left_bin_edges=(-180/deg,-180/deg),
bin_widths=(2.0/deg,2.0/deg))
path_dens_counter = path_density.histogram([s.active[0].trajectory for s in flexible.steps])
```
Now we've built the path density histogram, and we want to visualize it. We have a convenient `plot_2d_histogram` function that works in this case, and takes the histogram, desired plot tick labels and limits, and additional `matplotlib` named arguments to `plt.pcolormesh`.
```
tick_labels = np.arange(-np.pi, np.pi+0.01, np.pi/4)
plotter = HistogramPlotter2D(path_density,
xticklabels=tick_labels,
yticklabels=tick_labels,
label_format="{:4.2f}")
ax = plotter.plot(cmap="Blues")
```
## Convert to MDTraj for analysis by external tools
The trajectory can be converted to an MDTraj trajectory, and then used anywhere that MDTraj can be used. This includes writing it to a file (in any number of file formats) or visualizing the trajectory using, e.g., NGLView.
```
ops_traj = flexible.steps[1000].active[0].trajectory
traj = ops_traj.to_mdtraj()
traj
# Here's how you would then use NGLView:
#import nglview as nv
#view = nv.show_mdtraj(traj)
#view
flexible.close()
```
| github_jupyter |
# OpenACC Interoperability
This lab is intended for C/C++ programmers. If you prefer to use Fortran, click [this link.](../Fortran/README.ipynb)
---
## Introduction
The primary goal of this lab is to cover how to write an OpenACC code to work alongside other CUDA codes and accelerated libraries. There are several ways to make an OpenACC/CUDA interoperable code, and we will go through them one-by-one, with a short exercise for each.
When programming in OpenACC, the distinction between CPU/GPU memory is abstracted. For the most part, you do not need to worry about explicitly differentiating between CPU and GPU pointers; the OpenACC runtime handles this for you. However, in CUDA, you do need to differentiate between these two types of pointers. Let's start with using CUDA allocated GPU data in our OpenACC code.
---
## OpenACC Deviceptr Clause
The OpenACC `deviceptr` clause is used with the `data`, `parallel`, or `kernels` directives. It can be used in the same way as other data clauses such as `copyin`, `copyout`, `copy`, or `present`. The `deviceptr` clause is used to specify that a pointer is not a host pointer but rather a device pointer.
This clause is important when working with OpenACC + CUDA interoperability because it is one way we can operate on CUDA allocated device data within an OpenACC code. Take the following example:
**Allocation with CUDA**
```c++
double *cuda_allocate(int size) {
double *ptr;
cudaMalloc((void**) &ptr, size * sizeof(double));
return ptr;
}
```
**Parallel Loop with OpenACC**
```c++
int main() {
double *cuda_ptr = cuda_allocate(100); // Allocated on the device, but not the host!
#pragma acc parallel loop deviceptr(cuda_ptr)
for(int i = 0; i < 100; i++) {
cuda_ptr[i] = 0.0;
}
}
```
Normally, the OpenACC runtime expects to be given a host pointer, which will then be translated to some associated device pointer. However, when using CUDA to do our data management, we do not have that connection between host and device. The `deviceptr` clause is a way to tell the OpenACC runtime that a given pointer should not be translated since it is already a device pointer.
To practice using the `deviceptr` clause, we have a short exercise. We will examine two functions, both compute a dot product. The first code is [dot.c](/edit/C/deviceptr/dot.c), which is a serial dot product. Next is [dot_acc.c](/edit/C/deviceptr/dot_acc.c), which is an OpenACC parallelized version of dot. Both dot and dot_acc are called from [main.cu](/edit/C/deviceptr/main.cu) (*note: .cu is the conventional extension for a CUDA C++ source file*). In main.cu, we use host pointers to call dot, and device pointers to call dot_acc. Let's quickly run the code, it will produce an error.
```
!make -C deviceptr
```
To fix the error, we must tell the OpenACC runtime in the dot_acc function that our pointers are device pointers. Edit the [dot_acc.c](/edit/C/deviceptr/dot_acc.c) file using the deviceptr clause to get the code working. When you think you have it, run the code below and see if the error is fixed.
```
!make -C deviceptr
```
Next, let's do the opposite. Let's take data that was allocated with OpenACC, and use it in a CUDA function.
---
## OpenACC host_data directive
The `host_data` directive is used to make the OpenACC mapped device address available to the host. There are a few clauses that can be used with host_data, but the one that we are interested in using is `use_device`. We will use the `host_data` directive with the `use_device` clause to grab the underlying device pointer that OpenACC usually abstracts for us. Then we can use this device pointer to pass to CUDA kernels or to use accelerated libraries. Let's look at a code example:
**Inside CUDA Code**
```c++
__global__
void example_kernel(int *A, int size) {
// Kernel Code
}
extern "C" void example_cuda(int *A, int size) {
example_kernel<<<512,128>>>(A, size);
}
```
**Inside OpenACC Code**
```c++
extern void example_cuda(int*, int);
int main() {
int *A = (int*) malloc(100*sizeof(int));
#pragma acc data create(A[0:100])
{
#pragma acc host_data use_device(A)
{
example_cuda(A, 100);
}
}
}
```
A brief rundown of what is actually happening under-the-hood: the `data` directive creates a device copy of the array `A`, and the host pointer of `A` is linked to the device pointer of `A`. This is typical OpenACC behavior. Next, the `host_data use_device` translates the `A` variable on the host to the device pointer so that we can pass it to our CUDA function.
To practice this, let's work on another code. We still have [dot.c](/edit/C/host_data/dot.c) for our serial code. But instead of an OpenACC version of dot, we have a CUDA version in [dot_kernel.cu](/edit/C/host_data/dot_kernel.cu). Both of these functions are called in [main.c](/edit/C/host_data/main.c). First, let's run the code and see the error.
```
!make -C host_data
```
Now edit [main.c](/edit/C/host_data/main.c) and use the `host_data` and `use_device` to pass device pointers when calling our CUDA function. When you're ready, rerun the code below, and see if the error is fixed.
```
!make -C host_data
```
---
## Using cuBLAS with OpenACC
We are also able to use accelerated libraries with `host_data` and `use_device` as well. Just like the previous section, we can allocate the data with OpenACC using either the `data` or `enter data` directives. Then, pass that data to a cuBLAS call with `host_data`. This code is slightly different than before; we will be working on a matrix multiplication code. The serial code is found in [matmult.c](/edit/C/cublas/matmult.c). The cuBLAS code is in [matmult_cublas.cu](/edit/C/cublas/matmult_cublas.cu). Both of these are called from [main.c](/edit/C/cublas/main.c). Let's try running the code and seeing the error.
```
!make -C cublas
```
Now, edit [main.c](/edit/C/cublas/main.c) and use host_data/use_device on the cublas call (similar to what you did in the previous exercise). Rerun the code below when you're ready, and see if the error is fixed.
```
!make -C cublas
```
Next we will learn how make CUDA allocated memory behave like OpenACC allocated memory.
---
## OpenACC map_data
We briefly mentioned earlier about how OpenACC creates a mapping between host and device memory. When using CUDA allocated memory within OpenACC, that mapping is not created automatically, but it can be created manually. We are able to map a host pointer to a device pointer by using the OpenACC `acc_map_data` function. Then, before the data is unallocated, you will use `acc_unmap_data` to undo the mapping. Let's look at a quick example.
**Inside CUDA Code**
```c++
int *cuda_allocate(int size) {
int *ptr;
cudaMalloc((void**) &ptr, size*sizeof(int));
return ptr;
}
void cuda_deallocate(int* ptr) {
cudaFree(ptr);
}
```
**Inside OpenACC Code**
```c++
int main() {
int *A = (int*) malloc(100 * sizeof(int));
int *A_device = cuda_allocate(100);
acc_map_data(A, A_device, 100*sizeof(int));
#pragma acc parallel loop present(A[0:100])
for(int i = 0; i < 100; i++) {
// Computation
}
acc_unmap_data(A);
cuda_deallocate(A_device);
free(A);
}
```
To practice, we have another example code which uses the `dot` functions again. Serial `dot` is in [dot.c](/edit/C/map/dot.c). OpenACC `dot` is in [dot_acc.c](/edit/C/map/dot_acc.c). Both of them are called from [main.cu](/edit/C/map/main.cu). Since main is a CUDA code, we have placed the OpenACC map/unmap in a separate file [map.c](/edit/C/map/map.c). Try running the code and seeing the error.
```
!make -C map
```
Now, edit [map.c](/edit/C/map/map.c) and add the OpenACC mapping functions. When you're ready, rerun the code below and see if the error is fixed.
```
!make -C map
```
---
## Routine
The last topic to discuss is using CUDA device functions within OpenACC `parallel` and `kernels` regions. These are functions that are compiled to be called from the accelerator within a GPU kernel or OpenACC region.
If you want to compile an OpenACC function to be used on the device, you will use the `routine` directive with the following syntax:
```c++
#pragma acc routine seq
int func() {
return 0;
}
```
You can also have a function with a loop you want to parallelize like so:
```c++
#pragma acc routine vector
int func() {
int sum = 0;
#pragma acc loop vector
for(int i = 0; i < 100; i++) {
sum += i;
}
return sum;
}
```
To use CUDA `__device__` functions within our OpenACC loops, we can also use the `routine` directive. See the following example:
**In CUDA Code**
```c++
extern "C" __device__
int cuda_func(int x) {
return x*x;
}
```
**In OpenACC Code**
```c++
#pragma acc routine seq
extern int cuda_func(int);
...
int main() {
A = (int*) malloc(100 * sizeof(int));
#pragma acc parallel loop copyout(A[:100])
for(int i = 0; i < 100; i++) {
A[i] = cuda_func(i);
}
}
```
To practice, we have one last code to try out. Our main function is in [main.c](/edit/C/routine/main.c), and our serial code is in [distance_map.c](/edit/C/routine/distance_map.c). Our parallel loop is in [distance_map_acc.c](/edit/C/routine/distance_map_acc.c). Note that the parallel loop is trying to use a CUDA `__device__` function without including any routine information. The CUDA function is in [dist_cuda.cu](/edit/C/routine/dist_cuda.cu). Let's run the code and see the error.
```
!make -C routine
```
Now, edit [distance_map_acc.c](/edit/C/routine/distance_map_acc.c) and include the routine directive. When you're ready, rerun the code below and see if the error is fixed.
```
!make -C routine
```
---
## Bonus Task
Here are some additional resources for OpenACC/CUDA interoperability:
[This is an NVIDIA devblog about some common techniques for implementing OpenACC + CUDA](https://devblogs.nvidia.com/3-versatile-openacc-interoperability-techniques/)
[This is a github repo with some additional code examples demonstrating the lessons covered in this lab](https://github.com/jefflarkin/openacc-interoperability)
---
## Post-Lab Summary
If you would like to download this lab for later viewing, it is recommend you go to your browsers File menu (not the Jupyter notebook file menu) and save the complete web page. This will ensure the images are copied down as well.
You can also execute the following cell block to create a zip-file of the files you've been working on, and download it with the link below.
```
%%bash
rm -f openacc_files.zip
zip -r openacc_files.zip *
```
**After** executing the above zip command, you should be able to download the zip file [here](files/openacc_files.zip)
| github_jupyter |
*This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by Jake VanderPlas; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
# Introduction to machine learning

*image from Deep Learning with Javascript*
# Act 1

```
def decide(income, criminal_record, years_job, credit_payments):
if income < 30000:
if criminal_record:
return 1
else:
return 0
elif income <= 70000:
if years_job < 1:
return 0
elif years_job <= 5:
if credit_payments:
return 1
else:
return 0
else:
return 1
else:
if criminal_record:
return 0
else:
return 1
decide(income=20000, criminal_record=1, years_job=3, credit_payments=1)
import random
import pandas as pd
random.seed(333)
data = []
for i in range(100):
income = random.randint(0, 100000)
criminal_record = random.randint(0, 1)
years_job = random.randint(0, 10)
credit_payments = random.randint(0, 1)
decision = decide(income, criminal_record, years_job, credit_payments)
data.append({'income':income, 'criminal_record':criminal_record, 'years_job':years_job,
'credit_payments':credit_payments, 'decision':decision})
df = pd.DataFrame(data)
df.head(20)
```
# Act 2
```
import pandas as pd
from sklearn import tree
from sklearn.tree import export_text
dtree = tree.DecisionTreeClassifier().fit(
df[['income','criminal_record','years_job','credit_payments']], df['decision'])
print(export_text(dtree, feature_names=['income','criminal_record','years_job','credit_payments']))
```
# What Is Machine Learning?
*building models of data*
Fundamentally, machine learning involves building mathematical models to help understand data.
"Learning" enters the fray when we give these models *tunable parameters* that can be adapted to observed data; in this way the program can be considered to be "learning" from the data.
Once these models have been fit to previously seen data, they can be used to predict and understand aspects of newly observed data.
## Categories of Machine Learning
- *Supervised learning*: Models that can predict labels based on labeled training data
- *Classification*: Models that predict labels as two or more discrete categories
- *Regression*: Models that predict continuous labels
- *Unsupervised learning*: Models that identify structure in unlabeled data
- *Clustering*: Models that detect and identify distinct groups in the data
- *Dimensionality reduction*: Models that detect and identify lower-dimensional structure in higher-dimensional data
### Classification: Predicting discrete labels
We will first take a look at a simple *classification* task, in which you are given a set of labeled points and want to use these to classify some unlabeled points.
Imagine that we have the data shown in this figure:

[figure source in Appendix](06.00-Figure-Code.ipynb#Classification-Example-Figure-1)

[figure source in Appendix](06.00-Figure-Code.ipynb#Classification-Example-Figure-2)
Now that this model has been trained, it can be generalized to new, unlabeled data.
In other words, we can take a new set of data, draw this model line through it, and assign labels to the new points based on this model.
This stage is usually called *prediction*. See the following figure:

[figure source in Appendix](06.00-Figure-Code.ipynb#Classification-Example-Figure-3)
### Regression: Predicting continuous labels
In contrast with the discrete labels of a classification algorithm, we will next look at a simple *regression* task in which the labels are continuous quantities.
Consider the data shown in the following figure, which consists of a set of points each with a continuous label:

[figure source in Appendix](06.00-Figure-Code.ipynb#Regression-Example-Figure-1)
As with the classification example, we have two-dimensional data: that is, there are two features describing each data point.
The color of each point represents the continuous label for that point.
There are a number of possible regression models we might use for this type of data, but here we will use a simple linear regression to predict the points.
This simple linear regression model assumes that if we treat the label as a third spatial dimension, we can fit a plane to the data.
This is a higher-level generalization of the well-known problem of fitting a line to data with two coordinates.
We can visualize this setup as shown in the following figure:

[figure source in Appendix](06.00-Figure-Code.ipynb#Regression-Example-Figure-2)
This plane of fit gives us what we need to predict labels for new points.
Visually, we find the results shown in the following figure:

[figure source in Appendix](06.00-Figure-Code.ipynb#Regression-Example-Figure-4)
### Clustering: Inferring labels on unlabeled data
The classification and regression illustrations we just looked at are examples of supervised learning algorithms, in which we are trying to build a model that will predict labels for new data.
Unsupervised learning involves models that describe data without reference to any known labels.
One common case of unsupervised learning is "clustering," in which data is automatically assigned to some number of discrete groups.
For example, we might have some two-dimensional data like that shown in the following figure:

[figure source in Appendix](06.00-Figure-Code.ipynb#Clustering-Example-Figure-2)
By eye, it is clear that each of these points is part of a distinct group.
Given this input, a clustering model will use the intrinsic structure of the data to determine which points are related.
Using the very fast and intuitive *k*-means algorithm (see [In Depth: K-Means Clustering](05.11-K-Means.ipynb)), we find the clusters shown in the following figure:

[figure source in Appendix](06.00-Figure-Code.ipynb#Clustering-Example-Figure-2)
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_02_5_pandas_features.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 2: Python for Machine Learning**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 2 Material
Main video lecture:
* Part 2.1: Introduction to Pandas [[Video]](https://www.youtube.com/watch?v=bN4UuCBdpZc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_1_python_pandas.ipynb)
* Part 2.2: Categorical Values [[Video]](https://www.youtube.com/watch?v=4a1odDpG0Ho&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_2_pandas_cat.ipynb)
* Part 2.3: Grouping, Sorting, and Shuffling in Python Pandas [[Video]](https://www.youtube.com/watch?v=YS4wm5gD8DM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_3_pandas_grouping.ipynb)
* Part 2.4: Using Apply and Map in Pandas for Keras [[Video]](https://www.youtube.com/watch?v=XNCEZ4WaPBY&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_4_pandas_functional.ipynb)
* **Part 2.5: Feature Engineering in Pandas for Deep Learning in Keras** [[Video]](https://www.youtube.com/watch?v=BWPTj4_Mi9E&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_5_pandas_features.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
try:
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
```
# Part 2.5: Feature Engineering
Feature engineering is an essential part of machine learning. For now, we will manually engineer features. However, later in this course, we will see some techniques for automatic feature engineering.
## Calculated Fields
It is possible to add new fields to the data frame that your program calculates from the other fields. We can create a new column that gives the weight in kilograms. The equation to calculate a metric weight, given weight in pounds, is:
$ m_{(kg)} = m_{(lb)} \times 0.45359237 $
The following Python code performs this transformation:
```
import os
import pandas as pd
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA', '?'])
df.insert(1, 'weight_kg', (df['weight'] * 0.45359237).astype(int))
pd.set_option('display.max_columns', 6)
pd.set_option('display.max_rows', 5)
df
```
## Google API Keys
Sometimes you will use external API's to obtain data. The following examples show how to use the Google API keys to encode addresses for use with neural networks. To use these, you will need your own Google API key. The key I have below is not a real key; you need to put your own in there. Google will ask for a credit card, but unless you use a huge number of lookups, there will be no actual cost. YOU ARE NOT required to get a Google API key for this class; this only shows you how. If you would like to get a Google API key, visit this site and obtain one for **geocode**.
[Google API Keys](https://developers.google.com/maps/documentation/embed/get-api-key)
```
GOOGLE_KEY = 'REPLACE WITH YOUR GOOGLE API KEY'
```
# Other Examples: Dealing with Addresses
Addresses can be difficult to encode into a neural network. There are many different approaches, and you must consider how you can transform the address into something more meaningful. Map coordinates can be a good approach. [latitude and longitude](https://en.wikipedia.org/wiki/Geographic_coordinate_system) can be a useful encoding. Thanks to the power of the Internet, it is relatively easy to transform an address into its latitude and longitude values. The following code determines the coordinates of [Washington University](https://wustl.edu/):
```
import requests
address = "1 Brookings Dr, St. Louis, MO 63130"
response = requests.get(
'https://maps.googleapis.com/maps/api/geocode/json?key={}&address={}' \
.format(GOOGLE_KEY,address))
resp_json_payload = response.json()
if 'error_message' in resp_json_payload:
print(resp_json_payload['error_message'])
else:
print(resp_json_payload['results'][0]['geometry']['location'])
```
If latitude and longitude are fed into the neural network as two features, they might not be overly helpful. These two values would allow your neural network to cluster locations on a map. Sometimes cluster locations on a map can be useful. Figure 2.SMK shows the percentage of the population that smokes in the USA by state.
**Figure 2.SMK: Smokers by State**

The above map shows that certain behaviors, like smoking, can be clustered by the global region.
However, often you will want to transform the coordinates into distances. It is reasonably easy to estimate the distance between any two points on Earth by using the [great circle distance](https://en.wikipedia.org/wiki/Great-circle_distance) between any two points on a sphere:
The following code implements this formula:
$\Delta\sigma=\arccos\bigl(\sin\phi_1\cdot\sin\phi_2+\cos\phi_1\cdot\cos\phi_2\cdot\cos(\Delta\lambda)\bigr)$
$d = r \, \Delta\sigma$
```
from math import sin, cos, sqrt, atan2, radians
# Distance function
def distance_lat_lng(lat1,lng1,lat2,lng2):
# approximate radius of earth in km
R = 6373.0
# degrees to radians (lat/lon are in degrees)
lat1 = radians(lat1)
lng1 = radians(lng1)
lat2 = radians(lat2)
lng2 = radians(lng2)
dlng = lng2 - lng1
dlat = lat2 - lat1
a = sin(dlat / 2)**2 + cos(lat1) * cos(lat2) * sin(dlng / 2)**2
c = 2 * atan2(sqrt(a), sqrt(1 - a))
return R * c
# Find lat lon for address
def lookup_lat_lng(address):
response = requests.get(
'https://maps.googleapis.com/maps/api/geocode/json?key={}&address={}' \
.format(GOOGLE_KEY,address))
json = response.json()
if len(json['results']) == 0:
print("Can't find: {}".format(address))
return 0,0
map = json['results'][0]['geometry']['location']
return map['lat'],map['lng']
# Distance between two locations
import requests
address1 = "1 Brookings Dr, St. Louis, MO 63130"
address2 = "3301 College Ave, Fort Lauderdale, FL 33314"
lat1, lng1 = lookup_lat_lng(address1)
lat2, lng2 = lookup_lat_lng(address2)
print("Distance, St. Louis, MO to Ft. Lauderdale, FL: {} km".format(
distance_lat_lng(lat1,lng1,lat2,lng2)))
```
Distances can be a useful means to encode addresses. It would help if you considered what distance might be helpful for your dataset. Consider:
* Distance to a major metropolitan area
* Distance to a competitor
* Distance to a distribution center
* Distance to a retail outlet
The following code calculates the distance between 10 universities and Washington University in St. Louis:
```
# Encoding other universities by their distance to Washington University
schools = [
["Princeton University, Princeton, NJ 08544", 'Princeton'],
["Massachusetts Hall, Cambridge, MA 02138", 'Harvard'],
["5801 S Ellis Ave, Chicago, IL 60637", 'University of Chicago'],
["Yale, New Haven, CT 06520", 'Yale'],
["116th St & Broadway, New York, NY 10027", 'Columbia University'],
["450 Serra Mall, Stanford, CA 94305", 'Stanford'],
["77 Massachusetts Ave, Cambridge, MA 02139", 'MIT'],
["Duke University, Durham, NC 27708", 'Duke University'],
["University of Pennsylvania, Philadelphia, PA 19104",
'University of Pennsylvania'],
["Johns Hopkins University, Baltimore, MD 21218", 'Johns Hopkins']
]
lat1, lng1 = lookup_lat_lng("1 Brookings Dr, St. Louis, MO 63130")
for address, name in schools:
lat2,lng2 = lookup_lat_lng(address)
dist = distance_lat_lng(lat1,lng1,lat2,lng2)
print("School '{}', distance to wustl is: {}".format(name,dist))
```
| github_jupyter |
# SSNet Predictions
This notebook is meant for hands-on interaction with the code and data used in `SSNet_predictions.py`. Annotations explaining the general functioning of each section and the other modules they reference are provided. Similar notebooks may be added for individual models and combiners in the future. Note that the code shown here does not necessarily reflect the content of the script version.
This cell can be run to easily convert this notebook to a Python script:
```
!jupyter nbconvert --to script SSNet_predictions_notebook.ipynb
```
## License
```
'''
==================================================LICENSING TERMS==================================================
This code and data was developed by employees of the National Institute of Standards and Technology (NIST), an agency of the Federal Government. Pursuant to title 17 United States Code Section 105, works of NIST employees are not subject to copyright protection in the United States and are considered to be in the public domain. The code and data is provided by NIST as a public service and is expressly provided "AS IS." NIST MAKES NO WARRANTY OF ANY KIND, EXPRESS, IMPLIED OR STATUTORY, INCLUDING, WITHOUT LIMITATION, THE IMPLIED WARRANTY OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE, NON-INFRINGEMENT AND DATA ACCURACY. NIST does not warrant or make any representations regarding the use of the data or the results thereof, including but not limited to the correctness, accuracy, reliability or usefulness of the data. NIST SHALL NOT BE LIABLE AND YOU HEREBY RELEASE NIST FROM LIABILITY FOR ANY INDIRECT, CONSEQUENTIAL, SPECIAL, OR INCIDENTAL DAMAGES (INCLUDING DAMAGES FOR LOSS OF BUSINESS PROFITS, BUSINESS INTERRUPTION, LOSS OF BUSINESS INFORMATION, AND THE LIKE), WHETHER ARISING IN TORT, CONTRACT, OR OTHERWISE, ARISING FROM OR RELATING TO THE DATA (OR THE USE OF OR INABILITY TO USE THIS DATA), EVEN IF NIST HAS BEEN ADVISED OF THE POSSIBILITY OF SUCH DAMAGES.
To the extent that NIST may hold copyright in countries other than the United States, you are hereby granted the non-exclusive irrevocable and unconditional right to print, publish, prepare derivative works and distribute the NIST data, in any medium, or authorize others to do so on your behalf, on a royalty-free basis throughout the world.
You may improve, modify, and create derivative works of the code or the data or any portion of the code or the data, and you may copy and distribute such modifications or works. Modified works should carry a notice stating that you changed the code or the data and should note the date and nature of any such change. Please explicitly acknowledge the National Institute of Standards and Technology as the source of the code or the data: Citation recommendations are provided below. Permission to use this code and data is contingent upon your acceptance of the terms of this agreement and upon your providing appropriate acknowledgments of NIST's creation of the code and data.
Paper Title:
SSNet: a Sagittal Stratum-inspired Neural Network Framework for Sentiment Analysis
SSNet authors and developers:
Apostol Vassilev:
Affiliation: National Institute of Standards and Technology
Email: apostol.vassilev@nist.gov
Munawar Hasan:
Affiliation: National Institute of Standards and Technology
Email: munawar.hasan@nist.gov
Jin Honglan
Affiliation: National Institute of Standards and Technology
Email: honglan.jin@nist.gov
====================================================================================================================
'''
'''
This is master file that runs all the three combiners proposed in the paper.
Use following snippet to run all the three combiners: python SSNet_predictions.py
Please note that this code has tensorflow dependencies.
'''
```
## Imports/Dependencies
TensorFlow is the main machine learning framework used to implement, train, and apply the models. Pandas and NumPy are used for general data preprocessing and manipulation. Components from Matplotlib/Pyplot and IPython which are absent from `SSNet_predictions.py` are utilized here to provide enhanced interactivity and visualization. Functions from the following scripts (corresponding to the combiner models described in the paper) are imported:
- `SSNet_Neural_Network.py`
- `SSNet_Bayesian_Decision.py`
- `SSNet_Heuristic_Hybrid.py`
```
import tensorflow as tf
import math
import re
import pandas as pd
import numpy as np
import random
import os
import csv
import matplotlib.pyplot as plt
from IPython.display import JSON
import itertools
from SSNet_Neural_Network import nn
from SSNet_Bayesian_Decision import bayesian_decision
from SSNet_Heuristic_Hybrid import heuristic_hybrid
imdb_5ktr = 'imdb_train_5k.csv'
model_a_tr = 'model_1_5ktrain.csv'
model_b_tr = 'model_2_5ktrain.csv'
# model_c_tr = 'model_3_bert_result_train_5k.csv'
# model_d_tr = 'model_4_use_result_train_5k.csv'
model_c_tr = 'model_3_5ktrain.csv'
model_d_tr = 'model_4_5ktrain.csv'
model_a_te = 'model_1_25ktest.csv'
model_b_te = 'model_2_25ktest.csv'
# model_c_te = 'model_3_bert_result_test_25k.csv'
# model_d_te = 'model_4_use_result_test_25k.csv'
model_c_te = 'model_3_25ktest.csv'
model_d_te = 'model_4_25ktest.csv'
```
## Utilities
### Training Dict Threshold
```
def get_training_dict_threshold(split):
training_dict = dict()
if split == "5K":
training_dict["5K"] = [
[model_a_tr, model_b_tr, model_c_tr, model_d_tr], [
model_a_te, model_b_te, model_c_te, model_d_te]
]
return training_dict
list(itertools.combinations('1234', 2))
```
### Training Dictionary
```
def get_training_dict(split):
training_dict = dict()
# Store a running list of model combinations
# included_models = []
if split == "5K":
# Loop through integers 2 to 4 (inclusive); the number of component models in each combination
for r in range(2, 5):
# Generate combinations of model indices (without repetition)
for c in itertools.combinations(map(str, range(1, 5)), r):
training_dict['model_{{{}}}'.format(','.join(c))] = [
# Generate the file names corresponding to each model's output on both the training and testing data
[f'model_{m}_{n}{t}.csv' for m in c] for n, t in [
('5k', 'train'),
('25k', 'test')
]
]
return training_dict
# Test the function
JSON(get_training_dict("5K"))
# Compile a list of text files containing reviews and their corresponding sentiment labels
imdb_25k_list = list()
data_dir = 'models/train'
for file_name in os.listdir(f'../{data_dir}/pos'):
if file_name != '.DS_Store':
imdb_25k_list.append([file_name, str(1)])
for file_name in os.listdir(f'../{data_dir}/neg'):
if file_name != '.DS_Store':
imdb_25k_list.append([file_name, str(0)])
SAMPLE_SPLIT = ["5K"]
len(imdb_25k_list)
model_weights = []
```
## Train Predictors
Train the predictors and return the results.
```
def train_predictor():
for split in SAMPLE_SPLIT:
print("Sample Split: ", split)
imdb_list = list()
training_dict = None
training_dict_threshold = None
if split == "5K":
df_imdb_tr = pd.read_csv(imdb_5ktr)
for index in df_imdb_tr.index:
file_name = str(df_imdb_tr['file'][index])
label = int(df_imdb_tr['label'][index])
imdb_list.append([file_name, str(label)])
random.shuffle(imdb_list)
training_dict = get_training_dict(split)
training_dict_threshold = get_training_dict_threshold(split)
random.shuffle(imdb_list)
training_dict = get_training_dict(split)
training_dict_threshold = get_training_dict_threshold(split)
acc_dict_nn = dict()
acc_dict_bdc = dict()
for k, v in training_dict.items():
tr_list = list()
te_list = list()
for i in range(len(v[0])):
df = pd.read_csv(v[0][i])
df_dict = dict()
for idx in df.index:
file_name = str(df['file'][idx])
proba = float(df['prob'][idx])
df_dict[file_name] = proba
tr_list.append(df_dict)
for i in range(len(v[1])):
df = pd.read_csv(v[1][i])
df_dict = dict()
for idx in df.index:
file_name = str(df['file'][idx])
proba = float(df['prob'][idx])
df_dict[file_name] = proba
te_list.append(df_dict)
assert len(tr_list) == len(te_list), "train and test samples mismatch ...."
tr_acc = -1.
te_acc = -1.
while True:
tr_acc, te_acc, weights = nn(tr_list=tr_list, imdb_tr_list=imdb_list,
te_list=te_list, imdb_te_list=imdb_25k_list)
model_weights.append(weights)
if weights[0][0] == 0. or weights[0][1] == 0.:
print("bad event ...., training again")
print("\t" +k)
else:
break
acc_dict_nn[k] = [tr_acc, te_acc]
acc_dict_bdc[k] = bayesian_decision(tr_list=tr_list, imdb_tr_list=imdb_list,
te_list=te_list, imdb_te_list=imdb_25k_list)
for k, v in training_dict_threshold.items():
tr_list = list()
te_list = list()
for i in range(len(v[0])):
df = pd.read_csv(v[0][i])
df_dict = dict()
for idx in df.index:
file_name = str(df['file'][idx])
proba = float(df['prob'][idx])
df_dict[file_name] = proba
tr_list.append(df_dict)
for i in range(len(v[1])):
df = pd.read_csv(v[1][i])
df_dict = dict()
for idx in df.index:
file_name = str(df['file'][idx])
proba = float(df['prob'][idx])
df_dict[file_name] = proba
te_list.append(df_dict)
hh_dict = heuristic_hybrid(tr_list=tr_list, imdb_tr_list=imdb_list,
te_list=te_list, imdb_te_list=imdb_25k_list)
nn_metrics = []
# Print summaries of the results for each combiner
#print("Training Complete: ")
print("Neural Network Combiner: ")
for k, v in acc_dict_nn.items():
print("\t" +k +": training accuracy = " +str(v[0]) + ", test accuracy = " +str(v[1]))
nn_metrics.append(v)
bdr_metrics = []
print("\n")
print("Bayesian Decision Rule Combiner: ")
for k, v in acc_dict_bdc.items():
print("\t" +k)
for i, j in v.items():
print("\t\t" +i +": training accuracy = " +str(j[0]) +", test accuracy = " +str(j[1]))
bdr_metrics.append(v)
hh_metrics = []
print("\n")
print("Heuristic-Hybrid Combiner: ")
for k, v in hh_dict.items():
print("Base:", k)
for index in range(len(v)):
print("\t\t", v[index])
print("\n")
hh_metrics.append(v)
return nn_metrics, bdr_metrics, hh_metrics
W = [m[0] for m in model_weights[:7]]
print(W)
# plt.pcolormesh(model_weights)
```
## Result Aggregation
Runs the predictor training script and displays the results
```
trials = []
for i in range(1):
results = train_predictor()
trials.append(results)
def to_list(d):
L = [v for k, v in d.items()]
return L
# Convert result dictionaries to lists
for k, t in enumerate(trials):
for i in range(len(trials[k])):
for j, m in enumerate(trials[k][i]):
# for k, v in m.items():
if type(m) is dict:
trials[k][i][j] = to_list(m)
for j2, m2 in enumerate(m):
if type(m2) is dict:
trials[k][i][j][j2] = to_list(m2)
JSON(list(trials))
```
## Visualization of Results
```
use_scienceplots = True
b = trials[0][0]
# print(trials[0][0])
l = ['Train', 'Test']
combiner_names = [
'Neural Network',
'Bayesian Decision Rule',
'Heuristic-Hybrid'
]
bdr_props = ['Max', 'Avg', 'Sum', 'Majority']
hh_props = l + ['Threshold']
label_data = [l, bdr_props, hh_props]
# plot_style = 'ggplot'
if use_scienceplots:
try:
plt.style.use('science')
plt.style.use(['science','no-latex'])
except:
pass
plt.style.use('ggplot')
def grouped_plot(C=0, sections=2, reduce=0):
labels = [x.split('_')[-1] for x in get_training_dict('5K').keys()]
values = trials[0][C]
# print(values)
# valshape = np.array(values).shape
# if len(valshape) == 3:
# values
reduced = False
combiner_type = combiner_names[C]
if 'Bayesian' in combiner_type:
for i, v in enumerate(values):
for j, vi in enumerate(v):
if type(vi) in [list, tuple]:
values[i][j] = vi[reduce]
elif 'Heuristic' in combiner_type:
# print(values)
values = values[reduce]
# print(values)
if any(g in combiner_type for g in ['Bayesian', 'Heuristic']):
reduced = True
# plt.bar(labels, trials[0][0])
pos = np.arange(len(labels))
fig, ax = plt.subplots(figsize=(10, 5))
# sections = min(len(values[0]), sections)
sections = min(len(label_data[C]), sections)
for n in range(sections):
sections_ = min(len(values[0]), sections)
barwidth = (0.5/sections_)
# V = [x[n] if n <= len(x) else None for x in values]
V = []
for x in values:
try:
V.append(x[n])
except:
V.append(0)
# print(C, n, min(len(label_data[C]), n))
section_labels = label_data[C][min(len(label_data[C])-1, n)]
section_pos = pos+(barwidth*n)
# print(V, section_labels, section_pos, labels)
ax.bar(section_pos, V, barwidth, label=section_labels)
# ticks = pos-(barwidth*sections/4)
ticks = pos+(barwidth*round((sections-2)/2))
# print(ticks)
ax.set_xticks(ticks)
ax.set_xticklabels(labels)
ax.set_ylabel('Accuracy (%)')
ax.set_xlabel('Contributing Models')
s = l[reduce] if reduced else ''
title = f'{combiner_names[C]} Combiner | {s} Accuracy by Model'
# title += ' - '+l[reduce]
ax.set_title(title, pad=30)
# ax.margins(0.5)
# plt.subplots_adjust(left=0.3)
ax.legend(loc='upper right', bbox_to_anchor=(1.2, 1))
# fig.tight_layout(pad=1.5)
return ax
```
### Performance
Visualizes each model/combiner's loss values on the training and testing datasets
```
# Loop through the combiner indices and generate the graph for each one
for i in range(2):
grouped_plot(C=i, sections=4, reduce=1)
# Read a CSV file and return a (nested) list of its rows (split on commas)
# Params:
# filename: the name of the CSV file
# *or* [a and b]; the model number (e.g., "2") and dataset (e.g., "5ktrain")
def read_csv(filename=None, a=None, b=None, s=1):
if not filename:
filename = f'./model_{int(a)}_{b}.csv'
# If file extension is missing, assume CSV
if not filename.endswith('.csv'):
filename += '.csv'
with open(filename) as predictions:
reader = csv.reader(predictions)
data = list(reader)
data.sort(key=lambda d: d[s])
return data
imdb_data = []
for f in [5]:
imdb_data.extend(read_csv(f'imdb_train_{f}k.csv', s=2))
imdb_data.sort(key=lambda d: d[2])
print([d[2] for d in imdb_data[:3]])
imdb_data = [d[1:] for d in imdb_data]
```
### Predictions
```
p = []
# for f in range(2):
dataset = '5ktrain'
plot_size = (12, 6)
fig = plt.figure(figsize=plot_size)
plt.rcParams["figure.figsize"] = plot_size
fig, ax = plt.subplots()
# ax.set_xscale('log')
# ax.set_yscale('log')
# Load predictions for each model
for f in list('1234'):
p.append(read_csv(a=f, b=dataset))
# p.append(read_csv('imdb_train_5k'))
p.append(imdb_data)
# p.append(read_csv(a='2', b=dataset))
# Convert strings in data to floating-point numbers
def prep_data(n):
return [float(m[0]) for m in n if '.' in m[0]]
# Prepare the data to graph
points = [prep_data(n) for n in p]
# ax.scatter(*points)
# Draw the plot
ax.scatter(*points[:2], alpha=0.5, s=5, c=points[4], cmap='inferno')
ax.axis('off')
```
### Losses
Plot the distribution of loss values (absolute value of difference between predicted and actual value) for each model.
```
target = [0, 1, 2, 3]
A = 1 / len(target)
# losses = np.abs(np.array(points[1]) - np.array(points[2]))
losses = []
for t in target:
ys = np.array(prep_data(imdb_data))
# ys = np.random.randint(0, 2, ys.shape)
loss = np.abs(np.array(prep_data(p[t])) - np.array(ys))
losses.append(loss)
x = plt.hist(losses, bins=15, alpha=1)
np.array(prep_data(p[0])).shape
```
### Prediction Distribution
Plot a histogram of the models' predictions.
```
target = [0, 1, 2, 3]
A = 1 / len(target)
# for t in target:
# plt.hist(points[t], bins=50, alpha=A)
plt.style.use('seaborn-deep')
x = plt.hist([points[t] for t in target], bins=15)
p[3][:10]
a = {
'a': 5,
'b': 35,
'c': 2.8
}
# plt.plot(a)
```
| github_jupyter |
<a href="https://colab.research.google.com/gist/justheuristic/4c82ef4d448ce62cb5459484f66f56aa/practice.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
### Practice 1: Parallel GloVe
In this assignment we'll build parallel GloVe training from scratch. Well, almost from scratch:
* we'll use python's builtin [`multiprocessing`](https://docs.python.org/3/library/multiprocessing.html) library
* and learn to access numpy arrays from multiple processes!

```
%env MKL_NUM_THREADS=1
%env NUMEXPR_NUM_THREADS=1
%env OMP_NUM_THREADS=1
# set numpy to single-threaded mode for benchmarking
!pip install --upgrade nltk datasets tqdm
!wget https://raw.githubusercontent.com/mryab/efficient-dl-systems/main/week02_distributed/utils.py -O utils.py
import time, random
import multiprocessing as mp
import numpy as np
from tqdm import tqdm, trange
from IPython.display import clear_output
import matplotlib.pyplot as plt
%matplotlib inline
```
### Multiprocessing basics
```
def foo(i):
""" Imagine particularly computation-heavy function... """
print(end=f"Began foo({i})...\n")
result = np.sin(i)
time.sleep(abs(result))
print(end=f"Finished foo({i}) = {result:.3f}.\n")
return result
%%time
results_naive = [foo(i) for i in range(10)]
```
Same, but with multiple processes
```
%%time
processes = []
for i in range(10):
proc = mp.Process(target=foo, args=[i])
processes.append(proc)
print(f"Created {len(processes)} processes!")
# start in parallel
for proc in processes:
proc.start()
# wait for everyone finish
for proc in processes:
proc.join() # wait until proc terminates
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
Great! But how do we collect the values?
__Solution 1:__ with pipes!
Two "sides", __one__ process from each side
* `pipe_side.send(data)` - throw data into the pipe (do not wait for it to be read)
* `data = pipe_side.recv()` - read data. If there is none, wait for someone to send data
__Rules:__
* each side should be controlled by __one__ process
* data transferred through pipes must be serializable
* if `duplex=True`, processes can communicate both ways
* if `duplex=False`, "left" receives and "right" side sends
```
side_A, side_B = mp.Pipe()
side_A.send(123)
side_A.send({'ololo': np.random.randn(3)})
print("side_B.recv() -> ", side_B.recv())
print("side_B.recv() -> ", side_B.recv())
# note: calling recv a third will hang the process (waiting for someone to send data)
def compute_and_send(i, output_pipe):
print(end=f"Began compute_and_send({i})...\n")
result = np.sin(i)
time.sleep(abs(result))
print(end=f"Finished compute_and_send({i}) = {result:.3f}.\n")
output_pipe.send(result)
%%time
result_pipes = []
for i in range(10):
side_A, side_B = mp.Pipe(duplex=False)
# note: duplex=False means that side_B can only send
# and side_A can only recv. Otherwise its bidirectional
result_pipes.append(side_A)
proc = mp.Process(target=compute_and_send, args=[i, side_B])
proc.start()
print("MAIN PROCESS: awaiting results...")
for pipe in result_pipes:
print(f"MAIN_PROCESS: received {pipe.recv()}")
print("MAIN PROCESS: done!")
```
__Solution 2:__ with multiprocessing templates
Multiprocessing contains some template data structures that help you communicate between processes.
One such structure is `mp.Queue` a Queue that can be accessed by multiple processes in parallel.
* `queue.put` adds the value to the queue, accessible by all other processes
* `queue.get` returns the earliest added value and removes it from queue
```
queue = mp.Queue()
def func_A(queue):
print("A: awaiting queue...")
print("A: retreived from queue:", queue.get())
print("A: awaiting queue...")
print("A: retreived from queue:", queue.get())
print("A: done!")
def func_B(i, queue):
value = np.random.rand()
time.sleep(value)
print(f"proc_B{i}: putting more stuff into queue!")
queue.put(value)
proc_A = mp.Process(target=func_A, args=[queue])
proc_A.start();
proc_B1 = mp.Process(target=func_B, args=[1, queue])
proc_B2 = mp.Process(target=func_B, args=[2, queue])
proc_B1.start(), proc_B2.start();
```
__Important note:__ you can see that the two values above are identical.
This is because proc_B1 and proc_B2 were forked (cloned) with __the same random state!__
To mitigate this issue, run `np.random.seed()` in each process (same for torch, tensorflow).
<details>
<summary>In fact, please go and to that <b>right now!</b></summary>
<img src='https://media.tenor.com/images/32c950f36a61ec7e5060f5eee9140396/tenor.gif' height=200px>
</details>
```
```
__Less important note:__ `mp.Queue vs mp.Pipe`
- pipes are much faster for 1v1 communication
- queues support arbitrary number of processes
- queues are implemented with pipes
### GloVe preprocessing
Before we can train GloVe, we must first construct the co-occurence
```
import datasets
data = datasets.load_dataset('wikitext', 'wikitext-103-raw-v1')
# for fast debugging, you can temporarily use smaller data: 'wikitext-2-raw-v1'
print("Example:", data['train']['text'][5])
```
__First,__ let's build a vocabulary:
```
from collections import Counter
from nltk.tokenize import NLTKWordTokenizer
tokenizer = NLTKWordTokenizer()
def count_tokens(lines, top_k=None):
""" Tokenize lines and return top_k most frequent tokens and their counts """
sent_tokens = tokenizer.tokenize_sents(map(str.lower, lines))
token_counts = Counter([token for sent in sent_tokens for token in sent])
return Counter(dict(token_counts.most_common(top_k)))
count_tokens(data['train']['text'][:100], top_k=10)
# sequential algorithm
texts = data['train']['text'][:100_000]
vocabulary_size = 32_000
batch_size = 10_000
token_counts = Counter()
for batch_start in trange(0, len(texts), batch_size):
batch_texts = texts[batch_start: batch_start + batch_size]
batch_counts = count_tokens(batch_texts, top_k=vocabulary_size)
token_counts += Counter(batch_counts)
# save for later
token_counts_reference = Counter(token_counts)
```
### Let's parallelize (20% points)
__Your task__ is to speed up the code above using using multiprocessing with queues and/or pipes _(or [shared memory](https://docs.python.org/3/library/multiprocessing.shared_memory.html) if you're up to that)_.
__Kudos__ for implementing some form of global progress tracker (like progressbar above)
Please do **not** use task executors (e.g. mp.pool, joblib, ProcessPoolExecutor), we'll get to them soon!
```
texts = data['train']['text'][:100_000]
vocabulary_size = 32_000
batch_size = 10_000
<YOUR CODE HERE>
token_counts = <...>
assert len(token_counts) == len(token_counts_reference)
for token, ref_count in token_counts_reference.items():
assert token in token_counts, token
assert token_counts[token] == ref_count, token
token_counts = Counter(dict(token_counts.most_common(vocabulary_size)))
vocabulary = sorted(token_counts.keys())
token_to_index = {token: i for i, token in enumerate(vocabulary)}
assert len(vocabulary) == vocabulary_size, len(vocabulary)
print("Well done!")
```
### Part 2: Construct co-occurence matrix (10% points)
__Your task__ is to count co-occurences of all words in a 5-token window. Please use the same preprocessing and tokenizer as above.
__Also:__ please only count words that are in the vocabulary defined above.

__Note:__ this task and everything below has no instructions/interfaces. We will design those interfaces __together on the seminar.__
The detailed instructions will appear later this night after the seminar is over.
However, if you want to write the code from scratch, feel free to ignore these instructions.
```
import scipy
def count_token_cooccurences(lines, vocabulary_size: int, window_size: int):
""" Tokenize lines and return top_k most frequent tokens and their counts """
cooc = Counter()
for line in lines:
tokens = tokenizer.tokenize(line.lower())
token_ix = [token_to_index[token] for token in tokens
if token in token_to_index]
for i in range(len(token_ix)):
for j in range(max(i - window_size, 0),
min(i + window_size + 1, len(token_ix))):
if i != j:
cooc[token_ix[i], token_ix[j]] += 1 / abs(i - j)
return counter_to_matrix(cooc, vocabulary_size)
def counter_to_matrix(counter, vocabulary_size):
keys, values = zip(*counter.items())
ii, jj = zip(*keys)
return scipy.sparse.csr_matrix((values, (ii, jj)), dtype='float32',
shape=(vocabulary_size, vocabulary_size))
texts = data['train']['text'][:100_000]
batch_size = 10_000
window_size = 5
cooc = scipy.sparse.csr_matrix((vocabulary_size, vocabulary_size), dtype='float32')
for batch_start in trange(0, len(texts), batch_size):
batch_texts = texts[batch_start: batch_start + batch_size]
batch_cooc = count_token_cooccurences(batch_texts, vocabulary_size, window_size)
cooc += batch_cooc
# This cell will run for a couple minutes, go get some tea!
reference_cooc = cooc
```
__Simple parallelism with `mp.Pool`__
Many standard parallel tasks, such as applying the same function to an array of inputs, can be automated by using prebuilt primitives such as Pool.
```
def foo(i):
print(f'Began foo({i})', flush=True)
time.sleep(1)
print(f'Done foo({i})', flush=True)
return i ** 2
with mp.Pool(processes=8) as pool:
results = pool.map(foo, range(5))
# or use iterators:
# for result in pool.imap(foo, range(5)):
# print('Got', result)
```
__Our next step__ is to implement a parallel version of co-occurence computation using the process pool functionality.
There are multiple alternatives to mp.Pool: [joblib.Parallel](https://joblib.readthedocs.io/en/latest/), [ProcessPoolExecutor](https://docs.python.org/3/library/concurrent.futures.html#concurrent.futures.ProcessPoolExecutor), [ipyparallel](https://github.com/ipython/ipyparallel), etc. Feel free to use whichever one you prefer.
```
texts = data['train']['text'][:100_000]
batch_size = 10_000
window_size = 5
<YOUR CODE HERE>
cooc = <...>
assert isinstance(cooc, scipy.sparse.csr_matrix)
assert cooc.nnz == reference_cooc.nnz
for _ in trange(100_000):
i, j = np.random.randint(0, vocabulary_size, size=2)
assert np.allclose(cooc[i, j], reference_cooc[i, j])
print("Perfect!")
```
__Preprocess and save the full data__
Finally, let's run the preprocessing code for the entire dataset and save the results for training.
```
texts = data['train']['text']
vocabulary_size = 32_000
batch_size = 10_000
window_size = 5
# YOUR CODE: compute both vocabulary and cooc on the entire training corpora and save the results
<A WHOLE LOT OF YOUR CODE>
token_counts = <...>
cooc = <...>
assert len(vocabulary) == vocabulary_size
assert cooc.shape == (vocabulary_size, vocabulary_size)
assert 440_000_000 < np.sum(cooc) < 450_000_000
assert 0.05 < cooc.nnz / vocabulary_size ** 2 < 0.06
import pickle
with open('preprocessed_data.pcl', 'wb') as f:
pickle.dump((vocabulary, cooc.tocoo()), f)
```
### Finally, GloVe! (20% points)
```
import pickle
with open('preprocessed_data.pcl', 'rb') as f:
vocabulary, cooc = pickle.load(f)
```
### Weight function

```
def compute_loss_weights(counts_ij):
""" Compute GloVe weights """
<YOUR CODE HERE>
return <...>
dummy_weights = compute_loss_weights(np.arange(0, 200, 30))
dummy_reference_weights = [0. , 0.40536, 0.681731, 0.92402, 1. , 1. , 1.]
assert np.allclose(dummy_weights, dummy_reference_weights, rtol=1e-4, atol=1e-3)
```
### Loss function

__The goal__ is to compute the loss function as per formula above. The only difference is that you should take _mean_ over batch instead of sum.
```
def compute_loss(emb_ii, emb_jj, bias_ii, bias_jj, counts_ij):
"""
Compute GloVe loss function given embeddings, biases and targets
:param emb_ii, emb_jj: vectors of left- and right-side words, shape: [batch_size, embedding_dimension]
:param bias_ii, bias_jj: biases for left- and right-side words, shape: [batch_size]
:param counts_ij: values from co-occurence matrix, shape: [batch_size]
:returns: mean GloVe loss over batch, shape: scalar
"""
weights = compute_loss_weights(counts_ij)
target = np.log(counts_ij)
<YOUR CODE>
return <...>
dummy_emb_ii = np.sin(np.linspace(0, 10, 40)).reshape(4, 10)
dummy_emb_jj = np.cos(np.linspace(10, 20, 40)).reshape(4, 10)
dummy_bias_ii = np.linspace(-3, 2, 4)
dummy_bias_jj = np.linspace(4, -1, 4)
dummy_counts_ij = np.abs(np.sin(np.linspace(1, 100, 4)) * 150)
dummy_loss = compute_loss(dummy_emb_ii, dummy_emb_jj, dummy_bias_ii, dummy_bias_jj, dummy_counts_ij)
assert np.shape(dummy_loss) == ()
assert np.allclose(dummy_loss, 1.84289356)
def compute_grads(emb_ii, emb_jj, bias_ii, bias_jj, counts_ij):
"""
Compute gradients of GloVe loss with respect to emb_ii/jj and bias_ii/jj
Assume the same parameter shapes as above
:returns: (grad_wrt_emb_ii, grad_wrt_emb_jj, grad_wrt_bias_ii, grad_wrt_bias_jj)
"""
<YOUR CODE>
return <...>, <...>, <...>, <...>
grad_emb_ii, grad_emb_jj, grad_bias_ii, grad_bias_jj = compute_grads(
dummy_emb_ii, dummy_emb_jj, dummy_bias_ii, dummy_bias_jj, dummy_counts_ij)
assert np.shape(grad_emb_ii) == np.shape(grad_emb_jj) == np.shape(dummy_emb_ii)
assert np.shape(grad_bias_ii) == np.shape(grad_bias_jj) == np.shape(dummy_bias_ii)
from utils import eval_numerical_gradient
reference_grad_bias_ii = eval_numerical_gradient(
lambda x: compute_loss(dummy_emb_ii, dummy_emb_jj, x, dummy_bias_jj, dummy_counts_ij),
x=dummy_bias_ii)
assert np.allclose(reference_grad_bias_ii, grad_bias_ii, rtol=1e-4, atol=1e-3)
print("dL/db[ii] OK")
reference_grad_bias_jj = eval_numerical_gradient(
lambda x: compute_loss(dummy_emb_ii, dummy_emb_jj, dummy_bias_ii, x, dummy_counts_ij),
x=dummy_bias_jj)
assert np.allclose(reference_grad_bias_jj, grad_bias_jj, rtol=1e-4, atol=1e-3)
print("dL/db[jj] OK")
reference_grad_emb_ii = eval_numerical_gradient(
lambda x: compute_loss(x, dummy_emb_jj, dummy_bias_ii, dummy_bias_jj, dummy_counts_ij),
x=dummy_emb_ii)
assert np.allclose(reference_grad_emb_ii, grad_emb_ii, rtol=1e-4, atol=1e-3)
print("dL/dEmb[ii] OK")
reference_grad_emb_jj = eval_numerical_gradient(
lambda x: compute_loss(dummy_emb_ii, x, dummy_bias_ii, dummy_bias_jj, dummy_counts_ij),
x=dummy_emb_jj)
assert np.allclose(reference_grad_emb_jj, grad_emb_jj, rtol=1e-4, atol=1e-3)
print("dL/dEmb[ii] OK")
print("All tests passed!")
```
### Part 3: Parallel GloVe training (50% points)
Finally, let's write the actual parameter server for parallel GloVe training. In order to do so efficiently, we shall use shared memory instead of pipes.
You can find an example of how shared memory works below:
### Demo: shared memory
```
def make_shared_array(shape, dtype, fill=None, lock=True):
""" Create a numpy array that is shared across processes. """
size = int(np.prod(shape))
ctype = np.ctypeslib.as_ctypes_type(dtype)
if lock:
x_mp = mp.Array(ctype, size, lock=True).get_obj()
else:
x_mp = mp.Array(ctype, size, lock=False)
array = np.ctypeslib.as_array(x_mp)
if fill is not None:
array[...] = fill
return np.reshape(array, shape)
shared_array = make_shared_array((5, 5), 'float32', fill=1)
normal_array = np.ones((5, 5), 'float32')
def proc_A():
time.sleep(0.5)
print("A: setting value at [2, 3]")
shared_array[2, 3] = 42
normal_array[2, 3] = 42
time.sleep(1)
print(f"A: value after 1.5s: normal = {normal_array[2, 3]}\t shared = {shared_array[2, 3]}")
def proc_B():
print(f"B: initial value: normal = {normal_array[2, 3]}\t shared = {shared_array[2, 3]}")
time.sleep(1)
print(f"B: value after 1s: normal = {normal_array[2, 3]}\t shared = {shared_array[2, 3]}")
print("B: dividing value at [2, 3] by 2")
shared_array[2, 3] /= 2
normal_array[2, 3] /= 2
mp.Process(target=proc_A).start()
mp.Process(target=proc_B).start()
# the same can be done with individual values:
x = mp.Value(np.ctypeslib.as_ctypes_type(np.int32))
x.value += 1 # shared across all processes
```
__So, let's put all trainable parameters in shared memory!__
```
class SharedEmbeddings:
"""
Word embeddings trainable parameters, allocated in shared memory
"""
def __init__(self, vocabulary_size: int, embedding_dimension: int, init_scale: float = 0.01):
self.embeddings = make_shared_array([vocabulary_size, embedding_dimension], np.float32, lock=False)
self.embeddings[...] = np.random.randn(*self.embeddings.shape) * init_scale
self.biases = make_shared_array([vocabulary_size], np.float32, fill=0.0, lock=False)
```
### Training (single-core baseline)
```
batch_size = 64
learning_rate = 0.01
max_steps = 10 ** 6
start_time = time.perf_counter()
timestep_history = []
loss_history = []
model = SharedEmbeddings(vocabulary_size, embedding_dimension=256)
for t in trange(max_steps):
batch_ix = np.random.randint(0, len(cooc.row), size=batch_size)
ii, jj, counts_ij = cooc.row[batch_ix], cooc.col[batch_ix], cooc.data[batch_ix]
# Compute gradients
emb_ii, emb_jj, bias_ii, bias_jj = \
model.embeddings[ii], model.embeddings[jj], model.biases[ii], model.biases[jj]
grad_emb_ii, grad_emb_jj, grad_bias_ii, grad_bias_jj = compute_grads(
emb_ii, emb_jj, bias_ii, bias_jj, counts_ij)
# SGD step
model.embeddings[ii] -= learning_rate * grad_emb_ii
model.embeddings[jj] -= learning_rate * grad_emb_jj
model.biases[ii] -= learning_rate * grad_bias_ii
model.biases[jj] -= learning_rate * grad_bias_jj
if t % 10_000 == 0:
batch_ix = np.random.randint(0, len(cooc.row), size=4096)
ii, jj, counts_ij = cooc.row[batch_ix], cooc.col[batch_ix], cooc.data[batch_ix]
emb_ii, emb_jj, bias_ii, bias_jj = \
model.embeddings[ii], model.embeddings[jj], model.biases[ii], model.biases[jj]
timestep_history.append(time.perf_counter() - start_time)
loss_history.append(compute_loss(emb_ii, emb_jj, bias_ii, bias_jj, counts_ij))
clear_output(True)
plt.plot(timestep_history, loss_history)
plt.xlabel('training time(seconds)')
plt.grid()
plt.show()
```
__Now let's parallelize it!__
The code above is cute, but it only uses one CPU core. Surely we can go faster!
The main challenge in this week's seminar is to speed up GloVe training by all means necessary.
Here's what you should do:
* make multiple parallel workers, each training your model on different random data,
* build some centralized means of progress tracking: track the average loss and the number of training steps,
* implement workers in such a way that no process is left hanging after the training is over.
Finally, please compare the loss / training time plot of your algorithm against the baseline.
_Notes:_
* Remember to set a different np.random.seed in each worker!
* You can track the training progress either via mp.Pipe or via shared variables
* It is better to separate training and plotting into different processes
* If you want to prevent concurrent updates to shared memory, you can use [mp.Lock](https://docs.python.org/3/library/multiprocessing.html#multiprocessing.Lock) or similar.
```
batch_size = 64
learning_rate = 0.01
max_steps = 10 ** 6
start_time = time.perf_counter()
timestep_history = []
loss_history = []
model = SharedEmbeddings(vocabulary_size, embedding_dimension=256)
# <YOUR CODE HERE> - optional preparations, auxiliary functions, locks, pipes, etc.
# <YOUR CODE HERE> - actually train the model, track performance, and clean up at the end
```
Hello, I'm `______` and here's what i've done:
* something
| github_jupyter |
# Compas Analysis
What follows are the calculations performed for ProPublica's analaysis of the COMPAS Recidivism Risk Scores. It might be helpful to open [the methodology](https://www.propublica.org/article/how-we-analyzed-the-compas-recidivism-algorithm/) in another tab to understand the following.
## Loading the Data
We select fields for severity of charge, number of priors, demographics, age, sex, compas scores, and whether each person was accused of a crime within two years.
```
# filter dplyr warnings
%load_ext rpy2.ipython
import warnings
warnings.filterwarnings('ignore')
%%R
library(dplyr)
library(ggplot2)
raw_data <- read.csv("./compas-scores-two-years.csv")
nrow(raw_data)
```
However not all of the rows are useable for the first round of analysis.
There are a number of reasons remove rows because of missing data:
* If the charge date of a defendants Compas scored crime was not within 30 days from when the person was arrested, we assume that because of data quality reasons, that we do not have the right offense.
* We coded the recidivist flag -- `is_recid` -- to be -1 if we could not find a compas case at all.
* In a similar vein, ordinary traffic offenses -- those with a `c_charge_degree` of 'O' -- will not result in Jail time are removed (only two of them).
* We filtered the underlying data from Broward county to include only those rows representing people who had either recidivated in two years, or had at least two years outside of a correctional facility.
```
%%R
df <- dplyr::select(raw_data, age, c_charge_degree, race, age_cat, score_text, sex, priors_count,
days_b_screening_arrest, decile_score, is_recid, two_year_recid, c_jail_in, c_jail_out) %>%
filter(days_b_screening_arrest <= 30) %>%
filter(days_b_screening_arrest >= -30) %>%
filter(is_recid != -1) %>%
filter(c_charge_degree != "O") %>%
filter(score_text != 'N/A')
nrow(df)
```
Higher COMPAS scores are slightly correlated with a longer length of stay.
```
%%R
df$length_of_stay <- as.numeric(as.Date(df$c_jail_out) - as.Date(df$c_jail_in))
cor(df$length_of_stay, df$decile_score)
```
After filtering we have the following demographic breakdown:
```
%%R
summary(df$age_cat)
%%R
summary(df$race)
print("Black defendants: %.2f%%" % (3175 / 6172 * 100))
print("White defendants: %.2f%%" % (2103 / 6172 * 100))
print("Hispanic defendants: %.2f%%" % (509 / 6172 * 100))
print("Asian defendants: %.2f%%" % (31 / 6172 * 100))
print("Native American defendants: %.2f%%" % (11 / 6172 * 100))
%%R
summary(df$score_text)
%%R
xtabs(~ sex + race, data=df)
%%R
summary(df$sex)
print("Men: %.2f%%" % (4997 / 6172 * 100))
print("Women: %.2f%%" % (1175 / 6172 * 100))
%%R
nrow(filter(df, two_year_recid == 1))
%%R
nrow(filter(df, two_year_recid == 1)) / nrow(df) * 100
```
Judges are often presented with two sets of scores from the Compas system -- one that classifies people into High, Medium and Low risk, and a corresponding decile score. There is a clear downward trend in the decile scores as those scores increase for white defendants.
```
%%R -w 900 -h 363 -u px
library(grid)
library(gridExtra)
pblack <- ggplot(data=filter(df, race =="African-American"), aes(ordered(decile_score))) +
geom_bar() + xlab("Decile Score") +
ylim(0, 650) + ggtitle("Black Defendant's Decile Scores")
pwhite <- ggplot(data=filter(df, race =="Caucasian"), aes(ordered(decile_score))) +
geom_bar() + xlab("Decile Score") +
ylim(0, 650) + ggtitle("White Defendant's Decile Scores")
grid.arrange(pblack, pwhite, ncol = 2)
%%R
xtabs(~ decile_score + race, data=df)
```
## Racial Bias in Compas
After filtering out bad rows, our first question is whether there is a significant difference in Compas scores between races. To do so we need to change some variables into factors, and run a logistic regression, comparing low scores to high scores.
```
%%R
df <- mutate(df, crime_factor = factor(c_charge_degree)) %>%
mutate(age_factor = as.factor(age_cat)) %>%
within(age_factor <- relevel(age_factor, ref = 1)) %>%
mutate(race_factor = factor(race)) %>%
within(race_factor <- relevel(race_factor, ref = 3)) %>%
mutate(gender_factor = factor(sex, labels= c("Female","Male"))) %>%
within(gender_factor <- relevel(gender_factor, ref = 2)) %>%
mutate(score_factor = factor(score_text != "Low", labels = c("LowScore","HighScore")))
model <- glm(score_factor ~ gender_factor + age_factor + race_factor +
priors_count + crime_factor + two_year_recid, family="binomial", data=df)
summary(model)
```
Black defendants are 45% more likely than white defendants to receive a higher score correcting for the seriousness of their crime, previous arrests, and future criminal behavior.
```
%%R
control <- exp(-1.52554) / (1 + exp(-1.52554))
exp(0.47721) / (1 - control + (control * exp(0.47721)))
```
Women are 19.4% more likely than men to get a higher score.
```
%%R
exp(0.22127) / (1 - control + (control * exp(0.22127)))
```
Most surprisingly, people under 25 are 2.5 times as likely to get a higher score as middle aged defendants.
```
%%R
exp(1.30839) / (1 - control + (control * exp(1.30839)))
```
### Risk of Violent Recidivism
Compas also offers a score that aims to measure a persons risk of violent recidivism, which has a similar overall accuracy to the Recidivism score. As before, we can use a logistic regression to test for racial bias.
```
%%R
raw_data <- read.csv("./compas-scores-two-years-violent.csv")
nrow(raw_data)
%%R
df <- dplyr::select(raw_data, age, c_charge_degree, race, age_cat, v_score_text, sex, priors_count,
days_b_screening_arrest, v_decile_score, is_recid, two_year_recid) %>%
filter(days_b_screening_arrest <= 30) %>%
filter(days_b_screening_arrest >= -30) %>%
filter(is_recid != -1) %>%
filter(c_charge_degree != "O") %>%
filter(v_score_text != 'N/A')
nrow(df)
%%R
summary(df$age_cat)
%%R
summary(df$race)
%%R
summary(df$v_score_text)
%%R
nrow(filter(df, two_year_recid == 1)) / nrow(df) * 100
%%R
nrow(filter(df, two_year_recid == 1))
%%R -w 900 -h 363 -u px
library(grid)
library(gridExtra)
pblack <- ggplot(data=filter(df, race =="African-American"), aes(ordered(v_decile_score))) +
geom_bar() + xlab("Violent Decile Score") +
ylim(0, 700) + ggtitle("Black Defendant's Violent Decile Scores")
pwhite <- ggplot(data=filter(df, race =="Caucasian"), aes(ordered(v_decile_score))) +
geom_bar() + xlab("Violent Decile Score") +
ylim(0, 700) + ggtitle("White Defendant's Violent Decile Scores")
grid.arrange(pblack, pwhite, ncol = 2)
%%R
df <- mutate(df, crime_factor = factor(c_charge_degree)) %>%
mutate(age_factor = as.factor(age_cat)) %>%
within(age_factor <- relevel(age_factor, ref = 1)) %>%
mutate(race_factor = factor(race,
labels = c("African-American",
"Asian",
"Caucasian",
"Hispanic",
"Native American",
"Other"))) %>%
within(race_factor <- relevel(race_factor, ref = 3)) %>%
mutate(gender_factor = factor(sex, labels= c("Female","Male"))) %>%
within(gender_factor <- relevel(gender_factor, ref = 2)) %>%
mutate(score_factor = factor(v_score_text != "Low", labels = c("LowScore","HighScore")))
model <- glm(score_factor ~ gender_factor + age_factor + race_factor +
priors_count + crime_factor + two_year_recid, family="binomial", data=df)
summary(model)
```
The violent score overpredicts recidivism for black defendants by 77.3% compared to white defendants.
```
%%R
control <- exp(-2.24274) / (1 + exp(-2.24274))
exp(0.65893) / (1 - control + (control * exp(0.65893)))
```
Defendands under 25 are 7.4 times as likely to get a higher score as middle aged defendants.
```
%%R
exp(3.14591) / (1 - control + (control * exp(3.14591)))
```
## Predictive Accuracy of COMPAS
In order to test whether Compas scores do an accurate job of deciding whether an offender is Low, Medium or High risk, we ran a Cox Proportional Hazards model. Northpointe, the company that created COMPAS and markets it to Law Enforcement, also ran a Cox model in their [validation study](http://cjb.sagepub.com/content/36/1/21.abstract).
We used the counting model and removed people when they were incarcerated. Due to errors in the underlying jail data, we need to filter out 32 rows that have an end date more than the start date. Considering that there are 13,334 total rows in the data, such a small amount of errors will not affect the results.
```
%%R
library(survival)
library(ggfortify)
data <- filter(filter(read.csv("./cox-parsed.csv"), score_text != "N/A"), end > start) %>%
mutate(race_factor = factor(race,
labels = c("African-American",
"Asian",
"Caucasian",
"Hispanic",
"Native American",
"Other"))) %>%
within(race_factor <- relevel(race_factor, ref = 3)) %>%
mutate(score_factor = factor(score_text)) %>%
within(score_factor <- relevel(score_factor, ref=2))
grp <- data[!duplicated(data$id),]
nrow(grp)
%%R
summary(grp$score_factor)
%%R
summary(grp$race_factor)
%%R
f <- Surv(start, end, event, type="counting") ~ score_factor
model <- coxph(f, data=data)
summary(model)
```
People placed in the High category are 3.5 times as likely to recidivate, and the COMPAS system's concordance 63.6%. This is lower than the accuracy quoted in the Northpoint study of 68%.
```
%%R
decile_f <- Surv(start, end, event, type="counting") ~ decile_score
dmodel <- coxph(decile_f, data=data)
summary(dmodel)
```
COMPAS's decile scores are a bit more accurate at 66%.
We can test if the algorithm is behaving differently across races by including a race interaction term in the cox model.
```
%%R
f2 <- Surv(start, end, event, type="counting") ~ race_factor + score_factor + race_factor * score_factor
model <- coxph(f2, data=data)
print(summary(model))
```
The interaction term shows a similar disparity as the logistic regression above.
High risk white defendants are 3.61 more likely than low risk white defendants, while High risk black defendants are 2.99 more likely than low.
```
import math
print("Black High Hazard: %.2f" % (math.exp(-0.18976 + 1.28350)))
print("White High Hazard: %.2f" % (math.exp(1.28350)))
print("Black Medium Hazard: %.2f" % (math.exp(0.84286-0.17261)))
print("White Medium Hazard: %.2f" % (math.exp(0.84286)))
%%R -w 900 -h 563 -u px
fit <- survfit(f, data=data)
plotty <- function(fit, title) {
return(autoplot(fit, conf.int=T, censor=F) + ggtitle(title) + ylim(0,1))
}
plotty(fit, "Overall")
```
Black defendants do recidivate at higher rates according to race specific Kaplan Meier plots.
```
%%R -w 900 -h 363 -u px
white <- filter(data, race == "Caucasian")
white_fit <- survfit(f, data=white)
black <- filter(data, race == "African-American")
black_fit <- survfit(f, data=black)
grid.arrange(plotty(white_fit, "White defendants"),
plotty(black_fit, "Black defendants"), ncol=2)
%%R
summary(fit, times=c(730))
%%R
summary(black_fit, times=c(730))
%%R
summary(white_fit, times=c(730))
```
Race specific models have similar concordance values.
```
%%R
summary(coxph(f, data=white))
%%R
summary(coxph(f, data=black))
```
Compas's violent recidivism score has a slightly higher overall concordance score of 65.1%.
```
%%R
violent_data <- filter(filter(read.csv("./cox-violent-parsed.csv"), score_text != "N/A"), end > start) %>%
mutate(race_factor = factor(race,
labels = c("African-American",
"Asian",
"Caucasian",
"Hispanic",
"Native American",
"Other"))) %>%
within(race_factor <- relevel(race_factor, ref = 3)) %>%
mutate(score_factor = factor(score_text)) %>%
within(score_factor <- relevel(score_factor, ref=2))
vf <- Surv(start, end, event, type="counting") ~ score_factor
vmodel <- coxph(vf, data=violent_data)
vgrp <- violent_data[!duplicated(violent_data$id),]
print(nrow(vgrp))
summary(vmodel)
```
In this case, there isn't a significant coefficient on African American's with High Scores.
```
%%R
vf2 <- Surv(start, end, event, type="counting") ~ race_factor + race_factor * score_factor
vmodel <- coxph(vf2, data=violent_data)
summary(vmodel)
%%R
summary(coxph(vf, data=filter(violent_data, race == "African-American")))
%%R
summary(coxph(vf, data=filter(violent_data, race == "Caucasian")))
%%R -w 900 -h 363 -u px
white <- filter(violent_data, race == "Caucasian")
white_fit <- survfit(vf, data=white)
black <- filter(violent_data, race == "African-American")
black_fit <- survfit(vf, data=black)
grid.arrange(plotty(white_fit, "White defendants"),
plotty(black_fit, "Black defendants"), ncol=2)
```
## Directions of the Racial Bias
The above analysis shows that the Compas algorithm does overpredict African-American defendant's future recidivism, but we haven't yet explored the direction of the bias. We can discover fine differences in overprediction and underprediction by comparing Compas scores across racial lines.
```
from truth_tables import PeekyReader, Person, table, is_race, count, vtable, hightable, vhightable
from csv import DictReader
people = []
with open("./cox-parsed.csv") as f:
reader = PeekyReader(DictReader(f))
try:
while True:
p = Person(reader)
if p.valid:
people.append(p)
except StopIteration:
pass
pop = list(filter(lambda i: ((i.recidivist == True and i.lifetime <= 730) or
i.lifetime > 730), list(filter(lambda x: x.score_valid, people))))
recid = list(filter(lambda i: i.recidivist == True and i.lifetime <= 730, pop))
rset = set(recid)
surv = [i for i in pop if i not in rset]
print("All defendants")
table(list(recid), list(surv))
print("Total pop: %i" % (2681 + 1282 + 1216 + 2035))
import statistics
print("Average followup time %.2f (sd %.2f)" % (statistics.mean(map(lambda i: i.lifetime, pop)),
statistics.stdev(map(lambda i: i.lifetime, pop))))
print("Median followup time %i" % (statistics.median(map(lambda i: i.lifetime, pop))))
```
Overall, the false positive rate is 32.35%.
```
print("Black defendants")
is_afam = is_race("African-American")
table(list(filter(is_afam, recid)), list(filter(is_afam, surv)))
```
That number is higher for African Americans at 44.85%.
```
print("White defendants")
is_white = is_race("Caucasian")
table(list(filter(is_white, recid)), list(filter(is_white, surv)))
```
And lower for whites at 23.45%.
```
44.85 / 23.45
```
Which means under COMPAS black defendants are 91% more likely to get a higher score and not go on to commit more crimes than white defendants after two year.
COMPAS scores misclassify white reoffenders as low risk at 70.4% more often than black reoffenders.
```
47.72 / 27.99
hightable(list(filter(is_white, recid)), list(filter(is_white, surv)))
hightable(list(filter(is_afam, recid)), list(filter(is_afam, surv)))
```
## Risk of Violent Recidivism
Compas also offers a score that aims to measure a persons risk of violent recidivism, which has a similar overall accuracy to the Recidivism score.
```
vpeople = []
with open("./cox-violent-parsed.csv") as f:
reader = PeekyReader(DictReader(f))
try:
while True:
p = Person(reader)
if p.valid:
vpeople.append(p)
except StopIteration:
pass
vpop = list(filter(lambda i: ((i.violent_recidivist == True and i.lifetime <= 730) or
i.lifetime > 730), list(filter(lambda x: x.vscore_valid, vpeople))))
vrecid = list(filter(lambda i: i.violent_recidivist == True and i.lifetime <= 730, vpeople))
vrset = set(vrecid)
vsurv = [i for i in vpop if i not in vrset]
print("All defendants")
vtable(list(vrecid), list(vsurv))
```
Even moreso for Black defendants.
```
print("Black defendants")
is_afam = is_race("African-American")
vtable(list(filter(is_afam, vrecid)), list(filter(is_afam, vsurv)))
print("White defendants")
is_white = is_race("Caucasian")
vtable(list(filter(is_white, vrecid)), list(filter(is_white, vsurv)))
```
Black defendants are twice as likely to be false positives for a Higher violent score than white defendants.
```
38.14 / 18.46
```
White defendants are 63% more likely to get a lower score and commit another crime than Black defendants.
```
62.62 / 38.37
```
## Gender differences in Compas scores
In terms of underlying recidivism rates, we can look at gender specific Kaplan Meier estimates. There is a striking difference between women and men.
```
%%R
female <- filter(data, sex == "Female")
male <- filter(data, sex == "Male")
male_fit <- survfit(f, data=male)
female_fit <- survfit(f, data=female)
%%R
summary(male_fit, times=c(730))
%%R
summary(female_fit, times=c(730))
%%R -w 900 -h 363 -u px
grid.arrange(plotty(female_fit, "Female"), plotty(male_fit, "Male"),ncol=2)
```
As these plots show, the Compas score treats a High risk women the same as a Medium risk man.
| github_jupyter |
<a href="https://colab.research.google.com/github/ArpitaChatterjee/ANN-Datasets/blob/main/Ann.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#Artificial Neural Network
from google.colab import drive
drive.mount('/content/drive')
#Part1--Data Processing
#import the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
#import the dataset
dataset = pd.read_csv('/content/drive/My Drive/Colab Notebooks/ANN (DL)/Churn_Modelling.csv')
x=dataset.iloc[:, 3:13]
y=dataset.iloc[:, 13]
#create dummy variables...here its the ones which is of no use
geography=pd.get_dummies(x["Geography"], drop_first=True)
gender=pd.get_dummies(x["Gender"], drop_first=True)
#concatenate the data frames..i.e join the dummy variables created in one folder
x=pd.concat([x,geography,gender], axis=1)
#drop the unnecessary columns since they r already concated with x
x=x.drop(["Geography", "Gender"], axis=1)
#Splitting the dataset into training set and test set
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size= 0.2, random_state =0)
#feature Scaling.. it is necessary for every test cases to
from sklearn.preprocessing import StandardScaler#its a libraray for feature scaling
sc = StandardScaler()
x_train = sc.fit_transform(x_train)
x_test = sc.transform(x_test)
#Part2-- Lets make the ANN!
#import keras libraries and packages
from tensorflow import keras
from keras.models import Sequential#every nn we create ANN, RNN,CNN, need seq. library,its resposible for creating nn
from keras.layers import Dense#to create hiddenlayer we import this
#from keras.layers import LeakyRelu, PRelu, ELU#tocreate diff kind of layers
from keras.layers import Dropout#regularization parameter, used more frequently for deep NN
#initializing the ANN
classifier = Sequential()#empty nn to initialize NN, how many neurons r present there
#Adding the input layer and the first hidden layer
classifier.add(Dense(units = 6, kernel_initializer = 'he_uniform', activation='relu', input_dim = 11 ))
#to create first hl,1st= Output-dimension--no of neurons we consider for 1st hl--units,
#init--how our wt.should be initialized,for relu avtivation fn wt.initializatn=heuniform/henormal--kernel_initialization
# input-dimension--no of input layers connected to the hl, total no of col=11 in the xtest, trains.
#here we just used 6=hl neurons but using hyperoptimization technique well come to actual sol
#how many hl neurons r used how many hl are used, to calculate no of nurons in exact
#Adding the second hiddenlayer
classifier.add(Dense(units = 6, kernel_initializer ='he_uniform', activation='relu'))
#Adding the output layer
classifier.add(Dense(units = 1, kernel_initializer='glorot_uniform', activation='sigmoid'))
#lst layer will be 1dimension, with 1 hl neuron since its a binary classification and
#based on it well be able to predict if its >.5 ans=1 for this sigmoid activatn fn is used
#for all hl should have relu/leakyrelu as our Activatn fn
#as it solves Vanishing Gradient Problem unlike sigmoid hence used 1dim
#when we run classifier.summary we see how many output layers n neurons we have;params calc
# how many wt. we have initialized,n no of bias which we can calc mannually as well with the help of hl
#compiling the ANN..fitting up my optimizer
classifier.compile(optimizer = 'Adamax', loss= 'binary_crossentropy', metrics=['accuracy'])
#while compiling we determine type of optimizer, loss fn n matrix we r looking at
#fitting the ANN into the training dataset
model_history=classifier.fit(x_train, y_train,validation_split=0.33, batch_size = 10, nb_epoch= 100)
#we fit the xtrain, ytrain, validationsplit for testing the modules separately for test dataset
#batchsize--computational power bcomes less, more amt of data can beloaded at a time and our system is free for further computational
#no of epoch--=100, the accuracy will comme to given amt with time due to use of correct kernal initializer
#list all data in history
print(model_history.history.keys())
#summarize history for accuracy
plt.plot(model_history.history['acc'])
plt.plot(model_history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
#Part3--Making predictions and evaluating the models
#predicting the test set results
y_pred =classifier.predict(x_test)
y_pred = (y_pred >0.5)
#Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
#calculate the accuracy
from sklearn.metrics import accuracy_score
score=accuracy_score(y_pred, y_test)
```
| github_jupyter |
```
import torch
import torch.nn as nn
from torch.distributions import Categorical
import gym, os
from itertools import count
import torch.optim as optim
import torch.nn.functional as F
from torch.autograd import Variable
import matplotlib.pyplot as plt
import numpy as np
from math import log2
import pdb
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
class Model(nn.Module):
def __init__(self, state_dim, action_dim, n_latent_var):
super(Model, self).__init__()
self.affine = nn.Linear(state_dim, n_latent_var)
# actor
self.action_layer = nn.Sequential(
nn.Linear(state_dim, n_latent_var),
nn.Tanh(),
nn.Linear(n_latent_var, n_latent_var),
nn.Tanh(),
nn.Linear(n_latent_var, action_dim),
nn.Softmax(dim = -1)
)
# critic
self.value_layer = nn.Sequential(
nn.Linear(state_dim, n_latent_var),
nn.Tanh(),
nn.Linear(n_latent_var, n_latent_var),
nn.Tanh(),
nn.Linear(n_latent_var, 1)
)
# Memory:
self.actions = []
self.states = []
self.logprobs = []
self.state_values = []
self.rewards = []
def forward(self, state, action=None, evaluate=False):
# if evaluate is True then we also need to pass an action for evaluation
# else we return a new action from distribution
if not evaluate:
state = torch.from_numpy(state).float().to(device)
state_value = self.value_layer(state)
action_probs = self.action_layer(state)
action_distribution = Categorical(action_probs)
if not evaluate:
action = action_distribution.sample()
self.actions.append(action)
self.logprobs.append(action_distribution.log_prob(action))
self.state_values.append(state_value)
if evaluate:
return action_distribution.entropy().mean()
if not evaluate:
return action.item()
def clearMemory(self):
del self.actions[:]
del self.states[:]
del self.logprobs[:]
del self.state_values[:]
del self.rewards[:]
class PPO:
def __init__(self, state_dim, action_dim, n_latent_var, lr, betas, gamma, K_epochs, eps_clip):
self.lr = lr
self.betas = betas
self.gamma = gamma
self.eps_clip = eps_clip
self.K_epochs = K_epochs
self.policy = Model(state_dim, action_dim, n_latent_var).to(device)
self.optimizer = torch.optim.Adam(self.policy.parameters(),
lr=lr, betas=betas)
self.policy_old = Model(state_dim, action_dim, n_latent_var).to(device)
self.MseLoss = nn.MSELoss()
self.kl = 0
def update(self):
# Monte Carlo estimate of state rewards:
rewards = []
discounted_reward = 0
for reward in reversed(self.policy_old.rewards):
discounted_reward = reward + (self.gamma * discounted_reward)
rewards.insert(0, discounted_reward)
# Normalizing the rewards:
rewards = torch.tensor(rewards).to(device)
rewards = (rewards - rewards.mean()) / (rewards.std() + 1e-5)
# convert list in tensor
old_states = torch.tensor(self.policy_old.states).to(device).detach()
old_actions = torch.tensor(self.policy_old.actions).to(device).detach()
old_logprobs = torch.tensor(self.policy_old.logprobs).to(device).detach()
# Optimize policy for K epochs:
for _ in range(self.K_epochs):
# Evaluating old actions and values :
dist_entropy = self.policy(old_states, old_actions, evaluate=True)
# Finding the ratio (pi_theta / pi_theta__old):
logprobs = self.policy.logprobs[0].to(device)
ratios = torch.exp(logprobs - old_logprobs.detach())
self.kl =F.kl_div(logprobs - old_logprobs.detach())
# Finding Surrogate Loss:
state_values = self.policy.state_values[0].to(device)
advantages = rewards - state_values.squeeze().detach()
surr1 = ratios * advantages
surr2 = torch.clamp(ratios, 1-self.eps_clip, 1+self.eps_clip) * advantages
loss = -torch.min(surr1, surr2) + 0.5*self.MseLoss(state_values, rewards) - 0.01*dist_entropy
# take gradient step
self.optimizer.zero_grad()
loss.mean().backward()
self.optimizer.step()
self.policy.clearMemory()
self.policy_old.clearMemory()
# Copy new weights into old policy:
self.policy_old.load_state_dict(self.policy.state_dict())
import fruit.envs.games.deep_sea_treasure.engine as dst
from fruit.envs.juice import FruitEnvironment
game = dst.DeepSeaTreasure(width=5, seed=100, render=False, max_treasure=100, speed=1000)
env = FruitEnvironment(game)
print(env.get_number_of_objectives())
print(env.get_number_of_agents())
state_dim = 2
action_dim = 4
n_obj = env.get_number_of_objectives()
n_episodes = 100
max_timesteps = 500
kl_param = 0.1
log_interval = 10
n_latent_var = 64 # number of variables in hidden layer
lr = 0.0007
betas = (0.9, 0.999)
gamma = 0.99 # discount factor
K_epochs = 4 # update policy for K epochs
eps_clip = 0.2 # clip parameter for PPO
random_seed = None
if random_seed:
torch.manual_seed(random_seed)
env.seed(random_seed)
filename = "PPO_DeepSeaTreasure.pth"
directory = "./preTrained/"
env.get_state_space().get_shape()
ppo = PPO(state_dim, action_dim, n_latent_var, lr, betas, gamma, K_epochs, eps_clip)
running_reward = 0
rewards = np.zeros((n_episodes,n_obj))
for ep in range(1, n_episodes+1):
state = np.array([0.0,0.0])
for t in range(max_timesteps):
# Running policy_old:
action = ppo.policy_old(state)
reward = env.step(action)[0]
state_n = env.get_state()
done = env.is_terminal()
# Saving state and reward:
ppo.policy_old.states.append(state)
ppo.policy_old.rewards.append(reward)
state = np.array([state_n,0.0])
running_reward +=reward
if done:
# save model
torch.save(ppo.policy.state_dict(), directory+filename+str(0))
rewards[ep-1,0]= running_reward
break
running_reward = 0
for k in range(1,n_obj):
state = np.array([0.0,0.0])
for t in range(max_timesteps):
# Running policy_old:
action = ppo.policy_old(state)
reward = env.step(action)[k]
state_n = env.get_state()
done = env.is_terminal()
kl = ppo.kl
new_reward = reward + (kl_param*kl)
# Saving state and reward:
ppo.policy_old.states.append(state)
ppo.policy_old.rewards.append(new_reward)
state = np.array([state_n,0.0])
running_reward += new_reward
if done:
# save model
torch.save(ppo.policy.state_dict(), directory+filename+str(k))
rewards[ep-1,k]= running_reward
break
print('Episode: {}\tReward: {}'.format(ep, rewards[ep-1,:]))
running_reward = 0
plt.plot(np.arange(len(rewards[:,0])), rewards[:,0], label = 'reward1')
plt.plot(np.arange(len(rewards[:,1])), rewards[:,1], label = 'reward2')
plt.ylabel('Total Reward')
plt.xlabel('Episode')
plt.savefig('dst',bbox_inches='tight',facecolor="#FFFFFF")
plt.show()
ppo = PPO(state_dim, action_dim, n_latent_var, lr, betas, gamma, K_epochs, eps_clip)
running_reward = 0
rewards = []
for ep in range(1, n_episodes+1):
state = np.array([0.0,0.0])
for t in range(max_timesteps):
# Running policy_old:
action = ppo.policy_old(state)
reward = env.step(action)[0] + env.step(action)[1]
state_n = env.get_state()
done = env.is_terminal()
# Saving state and reward:
ppo.policy_old.states.append(state)
ppo.policy_old.rewards.append(reward)
state = np.array([state_n,0.0])
running_reward +=reward
if done:
# save model
torch.save(ppo.policy.state_dict(), directory+filename)
rewards.append(running_reward)
break
print('Episode: {}\tReward: {}'.format(ep, int(running_reward)))
running_reward = 0
plt.plot(np.arange(len(rewards)), rewards, label = 'reward1')
plt.ylabel('Total Reward')
plt.xlabel('Episode')
plt.savefig('dst_base',bbox_inches='tight',facecolor="#FFFFFF")
plt.show()
```
| github_jupyter |
# WELCOME TO MATPLOT VISUALIZATION
# Data Visualization with Matplotlib
This project is all about Matplotlib, the basic data visualization tool of Python programming language. I have discussed Matplotlib object hierarchy, various plot types with Matplotlib and customization techniques associated with Matplotlib.
# Contents
1] Introduction
2] Overview of Python Data Visualization Tools
3] Introduction to Matplotlib
4] Import Matplotlib
5] Displaying Plots in Matplotlib
6] Matplotlib Object Hierarchy
7] Matplotlib interfaces
8] Pyplot API
9] Object-Oriented API
10] Figure and Subplots
11] First plot with Matplotlib
12] Multiline Plots
13] Parts of a Plot
14] Saving the Plot
15] Line Plot
16] Scatter Plot
17] Histogram
18] Bar Chart
19] Horizontal Bar Chart
20] Error Bar Chart
21] Stacked Bar Chart
22] Pie Chart
23] Box Plot
24] Area Chart
25] Contour Plot
26] Styles with Matplotlib Plots
27] Adding a grid
28] Handling axes
29] Handling X and Y ticks
30] Adding labels
31] Adding a title
32] Adding a legend
33] Control colours
34] Control line styles
# Introduction
When we want to convey some information to others, there are several ways to do so. The process of conveying the information with the help of plots and graphics is called Data Visualization. The plots and graphics take numerical data as input and display output in the form of charts, figures and tables. It helps to analyze and visualize the data clearly and make concrete decisions. It makes complex data more accessible and understandable. The goal of data visualization is to communicate information in a clear and efficient manner.
In this project, I shed some light on Matplotlib, which is the basic data visualization tool of Python programming language. Python has different data visualization tools available which are suitable for different purposes. First of all, I will list these data visualization tools and then I will discuss Matplotlib.
# Overview of Python Visualization Tools
Python is the preferred language of choice for data scientists. Python have multiple options for data visualization. It has several tools which can help us to visualize the data more effectively. These Python data visualization tools are as follows:-
• Matplotlib
• Seaborn
• pandas
• Bokeh
• Plotly
• ggplot
• pygal
In the following sections, I discuss Matplotlib as the data visualization tool.
# Introduction to Matplotlib
Matplotlib is the basic plotting library of Python programming language. It is the most prominent tool among Python visualization packages. Matplotlib is highly efficient in performing wide range of tasks. It can produce publication quality figures in a variety of formats. It can export visualizations to all of the common formats like PDF, SVG, JPG, PNG, BMP and GIF. It can create popular visualization types – line plot, scatter plot, histogram, bar chart, error charts, pie chart, box plot, and many more types of plot. Matplotlib also supports 3D plotting. Many Python libraries are built on top of Matplotlib. For example, pandas and Seaborn are built on Matplotlib. They allow to access Matplotlib’s methods with less code.
The project Matplotlib was started by John Hunter in 2002. Matplotlib was originally started to visualize Electrocorticography (ECoG) data of epilepsy patients during post-doctoral research in Neurobiology. The open-source tool Matplotlib emerged as the most widely used plotting library for the Python programming language. It was used for data visualization during landing of the Phoenix spacecraft in 2008.
# Import Matplotlib
Before, we need to actually start using Matplotlib, we need to import it. We can import Matplotlib as follows:-
import matplotlib
Most of the time, we have to work with pyplot interface of Matplotlib. So, I will import pyplot interface of Matplotlib as follows:-
import matplotlib.pyplot
To make things even simpler, we will use standard shorthand for Matplotlib imports as follows:-
import matplotlib.pyplot as plt
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
# 5. Displaying Plots in Matplotlib
Viewing the Matplotlib plot is context based. The best usage of Matplotlib differs depending on how we are using it. There are three applicable contexts for viewing the plots. The three applicable contexts are using plotting from a script, plotting from an IPython shell or plotting from a Jupyter notebook.
## Plotting from a script
If we are using Matplotlib from within a script, then the plt.show() command is of great use. It starts an event loop, looks for all currently active figure objects, and opens one or more interactive windows that display the figure or figures.
The plt.show() command should be used only once per Python session. It should be used only at the end of the script. Multiple plt.show() commands can lead to unpredictable results and should mostly be avoided.
## Plotting from an IPython shell
We can use Matplotlib interactively within an IPython shell. IPython works well with Matplotlib if we specify Matplotlib mode. To enable this mode, we can use the %matplotlib magic command after starting ipython. Any plt plot command will cause a figure window to open and further commands can be run to update the plot.
## Plotting from a Jupyter notebook
The Jupyter Notebook (formerly known as the IPython Notebook) is a data analysis and visualization tool that provides multiple tools under one roof. It provides code execution, graphical plots, rich text and media display, mathematics formula and much more facilities into a single executable document.
Interactive plotting within a Jupyter Notebook can be done with the %matplotlib command. There are two possible options to work with graphics in Jupyter Notebook. These are as follows:-
• %matplotlib notebook – This command will produce interactive plots embedded within the notebook.
• %matplotlib inline – It will output static images of the plot embedded in the notebook.
After this command (it needs to be done only once per kernel per session), any cell within the notebook that creates a plot will embed a PNG image of the graphic.
```
%matplotlib inline
x1 = np.linspace(0, 10, 100)
# create a plot figure
fig = plt.figure()
plt.plot(x1, np.sin(x1), '-')
plt.plot(x1, np.cos(x1), '--');
```
# Matplotlib Object Hierarchy
There is an Object Hierarchy within Matplotlib. In Matplotlib, a plot is a hierarchy of nested Python objects. Ahierarch means that there is a tree-like structure of Matplotlib objects underlying each plot.
A Figure object is the outermost container for a Matplotlib plot. The Figure object contain multiple Axes objects. So, the Figure is the final graphic that may contain one or more Axes. The Axes represent an individual plot.
So, we can think of the Figure object as a box-like container containing one or more Axes. The Axes object contain smaller objects such as tick marks, lines, legends, title and text-boxes.
# Matplotlib API Overview
Matplotlib has two APIs to work with. A MATLAB-style state-based interface and a more powerful object-oriented (OO) interface. The former MATLAB-style state-based interface is called pyplot interface and the latter is called Object-Oriented interface.
There is a third interface also called pylab interface. It merges pyplot (for plotting) and NumPy (for mathematical functions) together in an environment closer to MATLAB. This is considered bad practice nowadays. So, the use of pylab is strongly discouraged and hence, I will not discuss it any further.
# Pyplot API
Matplotlib.pyplot provides a MATLAB-style, procedural, state-machine interface to the underlying object-oriented library in Matplotlib. Pyplot is a collection of command style functions that make Matplotlib work like MATLAB. Each pyplot function makes some change to a figure - e.g., creates a figure, creates a plotting area in a figure etc.
Matplotlib.pyplot is stateful because the underlying engine keeps track of the current figure and plotting area information and plotting functions change that information. To make it clearer, we did not use any object references during our plotting we just issued a pyplot command, and the changes appeared in the figure.
We can get a reference to the current figure and axes using the following commands-
plt.gcf ( ) # get current figure
plt.gca ( ) # get current axes
Matplotlib.pyplot is a collection of commands and functions that make Matplotlib behave like MATLAB (for plotting). The MATLAB-style tools are contained in the pyplot (plt) interface.
This is really helpful for interactive plotting, because we can issue a command and see the result immediately. But, it is not suitable for more complicated cases. For these cases, we have another interface called Object-Oriented interface, described later.
The following code produces sine and cosine curves using Pyplot API.
```
# create a plot figure
plt.figure()
# create the first of two panels and set current axis
plt.subplot(2, 1, 1) # (rows, columns, panel number)
plt.plot(x1, np.sin(x1))
# create the second of two panels and set current axis
plt.subplot(2, 1, 2) # (rows, columns, panel number)
plt.plot(x1, np.cos(x1));
print(plt.gcf())
print(plt.gca())
```
# Visualization with Pyplot
Generating visualization with Pyplot is very easy. The x-axis values ranges from 0-3 and the y-axis from 1-4. If we provide a single list or array to the plot() command, matplotlib assumes it is a sequence of y values, and automatically generates the x values. Since python ranges start with 0, the default x vector has the same length as y but starts with 0. Hence the x data are [0,1,2,3] and y data are [1,2,3,4].
```
plt.plot([1, 2, 3, 4])
plt.ylabel('Numbers')
plt.show()
```
# plot() - A versatile command
plot() is a versatile command. It will take an arbitrary number of arguments. For example, to plot x versus y, we can issue the following command:-
```
plt.plot([1, 2, 3, 4], [1, 4, 9, 16])
plt.show()
```
# State-machine interface
Pyplot provides the state-machine interface to the underlying object-oriented plotting library. The state-machine implicitly and automatically creates figures and axes to achieve the desired plot. For example:
```
x = np.linspace(0, 2, 100)
plt.plot(x, x, label='linear')
plt.plot(x, x**2, label='quadratic')
plt.plot(x, x**3, label='cubic')
plt.xlabel('x label')
plt.ylabel('y label')
plt.title("Simple Plot")
plt.legend()
plt.show()
```
# Formatting the style of plot
For every x, y pair of arguments, there is an optional third argument which is the format string that indicates the color and line type of the plot. The letters and symbols of the format string are from MATLAB. We can concatenate a color string with a line style string. The default format string is 'b-', which is a solid blue line. For example, to plot the above line with red circles, we would issue the following command:-
```
plt.plot([1, 2, 3, 4], [1, 4, 9, 16], 'ro')
plt.axis([0, 6, 0, 20])
plt.show()
```
# Working with NumPy arrays
Generally, we have to work with NumPy arrays. All sequences are converted to numpy arrays internally. The below example illustrates plotting several lines with different format styles in one command using arrays.
```
t = np.arange(0., 5., 0.2)
# red dashes, blue squares and green triangles
plt.plot(t, t, 'r--', t, t**2, 'bs', t, t**3, 'g^')
plt.show()
```
# 9. Object-Oriented API
The Object-Oriented API is available for more complex plotting situations. It allows us to exercise more control over the figure. In Pyplot API, we depend on some notion of an "active" figure or axes. But, in the Object-Oriented API the plotting functions are methods of explicit Figure and Axes objects.
Figure is the top level container for all the plot elements. We can think of the Figure object as a box-like container containing one or more Axes.
The Axes represent an individual plot. The Axes object contain smaller objects such as axis, tick marks, lines, legends, title and text-boxes.
The following code produces sine and cosine curves using Object-Oriented API.
```
# First create a grid of plots
# ax will be an array of two Axes objects
fig, ax = plt.subplots(2)
# Call plot() method on the appropriate object
ax[0].plot(x1, np.sin(x1), 'b-')
ax[1].plot(x1, np.cos(x1), 'b-');
```
# Objects and Reference
The main idea with the Object Oriented API is to have objects that one can apply functions and actions on. The real advantage of this approach becomes apparent when more than one figure is created or when a figure contains more than one subplot.
We create a reference to the figure instance in the fig variable. Then, we ceate a new axis instance axes using the add_axes method in the Figure class instance fig as follows:-
```
fig = plt.figure()
x2 = np.linspace(0, 5, 10)
y2 = x2 ** 2
axes = fig.add_axes([0.1, 0.1, 0.8, 0.8])
axes.plot(x2, y2, 'r')
axes.set_xlabel('x2')
axes.set_ylabel('y2')
axes.set_title('title');
```
# Figure and Axes
I start by creating a figure and an axes. A figure and axes can be created as follows:
fig = plt.figure()
ax = plt.axes()
In Matplotlib, the figure (an instance of the class plt.Figure) is a single container that contains all the objects representing axes, graphics, text and labels. The axes (an instance of the class plt.Axes) is a bounding box with ticks and labels. It will contain the plot elements that make up the visualization. I have used the variable name fig to refer to a figure instance, and ax to refer to an axes instance or group of axes instances.
```
fig = plt.figure()
ax = plt.axes()
```
# Figure and Subplots
Plots in Matplotlib reside within a Figure object. As described earlier, we can create a new figure with plt.figure() as follows:-
fig = plt.figure()
Now, I create one or more subplots using fig.add_subplot() as follows:-
ax1 = fig.add_subplot(2, 2, 1)
The above command means that there are four plots in total (2 * 2 = 4). I select the first of four subplots (numbered from 1).
I create the next three subplots using the fig.add_subplot() commands as follows:-
ax2 = fig.add_subplot(2, 2, 2)
ax3 = fig.add_subplot(2, 2, 3)
ax4 = fig.add_subplot(2, 2, 4)
The above command result in creation of subplots. The diagrammatic representation of subplots are as follows:-
Subplots.png
# First plot with Matplotlib
Now, I will start producing plots. Here is the first example:-
```
plt.plot([1, 3, 2, 4], 'b-')
plt.show( )
```
plt.plot([1, 3, 2, 4], 'b-')
This code line is the actual plotting command. Only a list of values has been plotted that represent the vertical coordinates of the points to be plotted. Matplotlib will use an implicit horizontal values list, from 0 (the first value) to N-1 (where N is the number of items in the list).
# Specify both Lists
Also, we can explicitly specify both the lists as follows:-
x3 = range(6)
plt.plot(x3, [xi**2 for xi in x3])
plt.show()
```
x3 = np.arange(0.0, 6.0, 0.01)
plt.plot(x3, [xi**2 for xi in x3], 'b-')
plt.show()
```
# 12. Multiline Plots
Multiline Plots mean plotting more than one plot on the same figure. We can plot more than one plot on the same figure.
It can be achieved by plotting all the lines before calling show(). It can be done as follows:-
```
x4 = range(1, 5)
plt.plot(x4, [xi*1.5 for xi in x4])
plt.plot(x4, [xi*3 for xi in x4])
plt.plot(x4, [xi/3.0 for xi in x4])
plt.show()
```
# 13. Parts of a Plot
There are different parts of a plot. These are title, legend, grid, axis and labels etc. These are denoted in the following figure:-
Parts%20of%20a%20plot.png
# Saving the Plot
We can save the figures in a wide variety of formats. We can save them using the savefig() command as follows:-
fig.savefig(‘fig1.png’)
We can explore the contents of the file using the IPython Image object.
from IPython.display import Image
Image(‘fig1.png’)
In savefig() command, the file format is inferred from the extension of the given filename. Depending on the backend, many different file formats are available. The list of supported file types can be found by using the get_supported_filetypes() method of the figure canvas object as follows:-
fig.canvas.get_supported_filetypes()
```
fig.savefig('plot1.png')
from IPython.display import Image
Image('plot1.png')
fig.canvas.get_supported_filetypes()
```
# 15. Line Plot
We can use the following commands to draw the simple sinusoid line plot:-
```
# Create figure and axes first
fig = plt.figure()
ax = plt.axes()
# Declare a variable x5
x5 = np.linspace(0, 10, 1000)
# Plot the sinusoid function
ax.plot(x5, np.sin(x5), 'b-');
```
# 16. Scatter Plot
Another commonly used plot type is the scatter plot. Here the points are represented individually with a dot or a circle.
# Scatter Plot with plt.plot()
We have used plt.plot/ax.plot to produce line plots. We can use the same functions to produce the scatter plots as follows:-
```
x7 = np.linspace(0, 10, 30)
y7 = np.sin(x7)
plt.plot(x7, y7, 'o', color = 'black');
```
# 17. Histogram
Histogram charts are a graphical display of frequencies. They are represented as bars. They show what portion of the dataset falls into each category, usually specified as non-overlapping intervals. These categories are called bins.
The plt.hist() function can be used to plot a simple histogram as follows:-
```
data1 = np.random.randn(1000)
plt.hist(data1);
```
# 18. Bar Chart
Bar charts display rectangular bars either in vertical or horizontal form. Their length is proportional to the values they represent. They are used to compare two or more values.
We can plot a bar chart using plt.bar() function. We can plot a bar chart as follows:-
```
data2 = [5. , 25. , 50. , 20.]
plt.bar(range(len(data2)), data2)
plt.show()
```
# 19. Horizontal Bar Chart
We can produce Horizontal Bar Chart using the plt.barh() function. It is the strict equivalent of plt.bar() function.
```
data2 = [5. , 25. , 50. , 20.]
plt.barh(range(len(data2)), data2)
plt.show()
```
# 20. Error Bar Chart
In experimental design, the measurements lack perfect precision. So, we have to repeat the measurements. It results in obtaining a set of values. The representation of the distribution of data values is done by plotting a single data point (known as mean value of dataset) and an error bar to represent the overall distribution of data.
We can use Matplotlib's errorbar() function to represent the distribution of data values. It can be done as follows:-
```
x9 = np.arange(0, 4, 0.2)
y9 = np.exp(-x9)
e1 = 0.1 * np.abs(np.random.randn(len(y9)))
plt.errorbar(x9, y9, yerr = e1, fmt = '.-')
plt.show();
```
# 21. Stacked Bar Chart
We can draw stacked bar chart by using a special parameter called bottom from the plt.bar() function. It can be done as follows:-
```
A = [15., 30., 45., 22.]
B = [15., 25., 50., 20.]
z2 = range(4)
plt.bar(z2, A, color = 'b')
plt.bar(z2, B, color = 'r', bottom = A)
plt.show()
```
The optional bottom parameter of the plt.bar() function allows us to specify a starting position for a bar. Instead of running from zero to a value, it will go from the bottom to value. The first call to plt.bar() plots the blue bars. The second call to plt.bar() plots the red bars, with the bottom of the red bars being at the top of the blue bars.
# Pie Chart
Pie charts are circular representations, divided into sectors. The sectors are also called wedges. The arc length of each sector is proportional to the quantity we are describing. It is an effective way to represent information when we are interested mainly in comparing the wedge against the whole pie, instead of wedges against each other.
Matplotlib provides the pie() function to plot pie charts from an array X. Wedges are created proportionally, so that each value x of array X generates a wedge proportional to x/sum(X).
```
plt.figure(figsize=(7,7))
x10 = [35, 25, 20, 20]
labels = ['Computer', 'Electronics', 'Mechanical', 'Chemical']
plt.pie(x10, labels=labels);
plt.show()
```
# 23. Boxplot
Boxplot allows us to compare distributions of values by showing the median, quartiles, maximum and minimum of a set of values.
We can plot a boxplot with the boxplot() function as follows:-
```
data3 = np.random.randn(100)
plt.boxplot(data3)
plt.show();
```
The boxplot() function takes a set of values and computes the mean, median and other statistical quantities. The following points describe the preceeding boxplot:
• The red bar is the median of the distribution.
• The blue box includes 50 percent of the data from the lower quartile to the upper quartile. Thus, the box is centered on the median of the data.
• The lower whisker extends to the lowest value within 1.5 IQR from the lower quartile.
• The upper whisker extends to the highest value within 1.5 IQR from the upper quartile.
• Values further from the whiskers are shown with a cross marker.
# Area Chart
An Area Chart is very similar to a Line Chart. The area between the x-axis and the line is filled in with color or shading. It represents the evolution of a numerical variable following another numerical variable.
We can create an Area Chart as follows:-
```
# Create some data
x12 = range(1, 6)
y12 = [1, 4, 6, 8, 4]
# Area plot
plt.fill_between(x12, y12)
plt.show()
```
# 25. Contour Plot
Contour plots are useful to display three-dimensional data in two dimensions using contours or color-coded regions. Contour lines are also known as level lines or isolines. Contour lines for a function of two variables are curves where the function has constant values. They have specific names beginning with iso- according to the nature of the variables being mapped.
There are lot of applications of Contour lines in several fields such as meteorology(for temperature, pressure, rain, wind speed), geography, magnetism, engineering, social sciences and so on.
The density of the lines indicates the slope of the function. The gradient of the function is always perpendicular to the contour lines. When the lines are close together, the length of the gradient is large and the variation is steep.
A Contour plot can be created with the plt.contour() function as follows:-
```
matrix1 = np.random.rand(10, 20)
cp = plt.contour(matrix1)
plt.show()
```
The contour() function draws contour lines. It takes a 2D array as input.Here, it is a matrix of 10 x 20 random elements.
The number of level lines to draw is chosen automatically, but we can also specify it as an additional parameter, N.
plt.contour(matrix, N)
# Styles with Matplotlib Plots
The Matplotlib version 1.4 which was released in August 2014 added a very convenient style module. It includes a number of new default stylesheets, as well as the ability to create and package own styles.
We can view the list of all available styles by the following command.
print(plt.style.availabe)
```
print(plt.style.available)
```
# 27. Adding a grid
In some cases, the background of a plot was completely blank. We can get more information, if there is a reference system in the plot. The reference system would improve the comprehension of the plot. An example of the reference system is adding a grid. We can add a grid to the plot by calling the grid() function. It takes one parameter, a Boolean value, to enable(if True) or disable(if False) the grid.
```
x15 = np.arange(1, 5)
plt.plot(x15, x15*1.5, x15, x15*3.0, x15, x15/3.0)
plt.grid(True)
plt.show()
```
# 28. Handling axes
Matplotlib automatically sets the limits of the plot to precisely contain the plotted datasets. Sometimes, we want to set the axes limits ourself. We can set the axes limits with the axis() function as follows:-
```
x15 = np.arange(1, 5)
plt.plot(x15, x15*1.5, x15, x15*3.0, x15, x15/3.0)
plt.axis() # shows the current axis limits values
plt.axis([0, 5, -1, 13])
plt.show()
```
We can see that we now have more space around the lines.
If we execute axis() without parameters, it returns the actual axis limits.
We can set parameters to axis() by a list of four values.
The list of four values are the keyword arguments [xmin, xmax, ymin, ymax] allows the minimum and maximum limits for X and Y axis respectively.
We can control the limits for each axis separately using the xlim() and ylim() functions. This can be done as follows:-
```
x15 = np.arange(1, 5)
plt.plot(x15, x15*1.5, x15, x15*3.0, x15, x15/3.0)
plt.xlim([1.0, 4.0])
plt.ylim([0.0, 12.0])
```
# 29. Handling X and Y ticks
Vertical and horizontal ticks are those little segments on the axes, coupled with axes labels, used to give a reference system on the graph.So, they form the origin and the grid lines.
Matplotlib provides two basic functions to manage them - xticks() and yticks().
Executing with no arguments, the tick function returns the current ticks' locations and the labels corresponding to each of them.
We can pass arguments(in the form of lists) to the ticks functions. The arguments are:-
Locations of the ticks
Labels to draw at these locations.
We can demonstrate the usage of the ticks functions in the code snippet below:-
```
u = [5, 4, 9, 7, 8, 9, 6, 5, 7, 8]
plt.plot(u)
plt.xticks([2, 4, 6, 8, 10])
plt.yticks([2, 4, 6, 8, 10])
plt.show()
```
# 30. Adding labels
Another important piece of information to add to a plot is the axes labels, since they specify the type of data we are plotting.
```
plt.plot([1, 3, 2, 4])
plt.xlabel('This is the X axis')
plt.ylabel('This is the Y axis')
plt.show()
```
# 31. Adding a title
The title of a plot describes about the plot. Matplotlib provides a simple function title() to add a title to an image.
```
plt.plot([1, 3, 2, 4])
plt.title('First Plot')
plt.show()
```
# 32. Adding a legend
Legends are used to describe what each line or curve means in the plot.
Legends for curves in a figure can be added in two ways. One method is to use the legend method of the axis object and pass a list/tuple of legend texts as follows:-
```
x15 = np.arange(1, 5)
fig, ax = plt.subplots()
ax.plot(x15, x15*1.5)
ax.plot(x15, x15*3.0)
ax.plot(x15, x15/3.0)
ax.legend(['Normal','Fast','Slow']);
```
The above method follows the MATLAB API. It is prone to errors and unflexible if curves are added to or removed from the plot. It resulted in a wrongly labelled curve.
A better method is to use the label keyword argument when plots are added to the figure. Then we use the legend method without arguments to add the legend to the figure.
The advantage of this method is that if curves are added or removed from the figure, the legend is automatically updated accordingly. It can be achieved by executing the code below:-
```
x15 = np.arange(1, 5)
fig, ax = plt.subplots()
ax.plot(x15, x15*1.5, label='Normal')
ax.plot(x15, x15*3.0, label='Fast')
ax.plot(x15, x15/3.0, label='Slow')
ax.legend();
```
The legend function takes an optional keyword argument loc. It specifies the location of the legend to be drawn. The loc takes numerical codes for the various places the legend can be drawn. The most common loc values are as follows:-
ax.legend(loc=0) # let Matplotlib decide the optimal location
ax.legend(loc=1) # upper right corner
ax.legend(loc=2) # upper left corner
ax.legend(loc=3) # lower left corner
ax.legend(loc=4) # lower right corner
ax.legend(loc=5) # right
ax.legend(loc=6) # center left
ax.legend(loc=7) # center right
ax.legend(loc=8) # lower center
ax.legend(loc=9) # upper center
ax.legend(loc=10) # center
# 33. Control colours
We can draw different lines or curves in a plot with different colours. In the code below, we specify colour as the last argument to draw red, blue and green lines.
```
x16 = np.arange(1, 5)
plt.plot(x16, 'r')
plt.plot(x16+1, 'g')
plt.plot(x16+2, 'b')
plt.show()
```
The colour names and colour abbreviations is given in the following table:-
Colour abbreviation Colour name
b blue
c cyan
g green
k black
m magenta
r red
w white
y yellow
There are several ways to specify colours, other than by colour abbreviations:
• The full colour name, such as yellow
• Hexadecimal string such as ##FF00FF
• RGB tuples, for example (1, 0, 1)
• Grayscale intensity, in string format such as ‘0.7’.
# 34.Control line styles
Matplotlib provides us different line style options to draw curves or plots. In the code below, I use different line styles to draw different plots.
```
x16 = np.arange(1, 5)
plt.plot(x16, '--', x16+1, '-.', x16+2, ':')
plt.show()
```
# That is all for this notebook, I hope this will be helpfull for some Data Science Aspirants
| github_jupyter |
# constellation_line_csv
星座線の始点、終点の位置をCSVファイルに保存する。
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
import struct
hip = pd.read_table("BrowseTargets.13041.1545121565", sep="|", header=2, usecols=range(1, 7), skipinitialspace=True)
hip.head()
```
# 星座線データの読み込み
星座線の始点・終点となる星のHIP IDが入っているCSVファイルを読み込む。
ソースはこちら↓
[星座線データ - Astro Commons](http://astronomy.webcrow.jp/hip/#line)
```
line = pd.read_csv("hip_constellation_line.csv", names=("name", "HIP_S", "HIP_E"))
line.head()
```
# 欠損値処理
各項目の欠損値の数
```
hip.isnull().sum()
```
Parallaxが0以下の星は除外する。
```
hip = hip[(hip["parallax"] > 0)]
```
bv_colorはなんか適当に埋めてみる。
とりあえず中央値で埋める。
```
hip["bv_color"] = hip["bv_color"].fillna(hip["bv_color"].median())
```
欠けたデータを除外した後の星の数
```
len(hip)
```
# 座標変換
年周視差([parallax](https://heasarc.gsfc.nasa.gov/W3Browse/star-catalog/hipparcos.html#parallax)) $pi$と距離(parsec) $D$の関係は$D = 1000 / pi$。1 parsecは3.26光年。
球座標系を直交座標系に変換する式はこんな感じになるはず。
$$
\begin{eqnarray}
x &=& \dfrac {1000}{P_{parallax}}\cos \theta_{dec} \cos \phi_{ra} \\
y &=& \dfrac {1000}{P_{parallax}}\cos \theta_{dec} \sin \phi_{ra} \\
z &=& \dfrac {1000}{P_{parallax}}\sin \theta_{dec}
\end{eqnarray}
$$
```
def radec2xyz(star):
x = 1000 / star["parallax"] * np.cos(np.deg2rad(star["dec_deg"])) * np.cos(np.deg2rad(star["ra_deg"]))
y = 1000 / star["parallax"] * np.cos(np.deg2rad(star["dec_deg"])) * np.sin(np.deg2rad(star["ra_deg"]))
z = 1000 / star["parallax"] * np.sin(np.deg2rad(star["dec_deg"]))
# Unityはyが上、zが前なので順序を変更
return [x, z, y]
```
# CSVファイルの作成
星座線の始点、終点となる星のIDを順に読み出し、星の位置を計算してリストを作成する。
```
constellation_line = pd.DataFrame({"xs": [],
"ys": [],
"zs": [],
"xe": [],
"ye": [],
"ze": [],
"name": []})
for index, i in line.iterrows():
try:
start_pos = radec2xyz(hip[hip["name"].str.startswith("HIP "+str(i["HIP_S"])+" ")].iloc[0])
except IndexError:
start_pos = radec2xyz(hip[hip["name"].str.startswith("HIP "+str(i["HIP_S"]))].iloc[0])
try:
end_pos = radec2xyz(hip[hip["name"].str.startswith("HIP "+str(i["HIP_E"])+" ")].iloc[0])
except IndexError:
end_pos = radec2xyz(hip[hip["name"].str.startswith("HIP "+str(i["HIP_E"]))].iloc[0])
name = [i["name"]]
constellation_line = constellation_line.append(pd.Series(start_pos+end_pos+name, index=constellation_line.columns), ignore_index=True)
constellation_line.to_csv("constellation_line.csv",header=False, index=False)
```
| github_jupyter |
```
from __future__ import print_function, division
import scipy
import pandas as pd
import numpy as np
import os
from keras.datasets import mnist
from keras_contrib.layers.normalization.instancenormalization import InstanceNormalization
from keras.layers import Input, Dense, Reshape, Flatten, Dropout, Concatenate
from keras.layers import BatchNormalization, Activation, ZeroPadding2D, Add
from keras.layers.advanced_activations import LeakyReLU
from keras.layers.convolutional import UpSampling2D, Conv2D
from keras.models import Sequential, Model
from keras.optimizers import Adam
from keras.utils import to_categorical
import datetime
import matplotlib.pyplot as plt
import sys
import scipy
from glob import glob
from keras.datasets import mnist
from skimage.transform import resize as imresize
import pickle
import os
import urllib
import gzip
class DataLoader():
"""Loads images from MNIST (domain A) and MNIST-M (domain B)"""
def __init__(self, img_res=(28, 28)):
self.img_res = img_res
self.mnistm_url = 'https://github.com/VanushVaswani/keras_mnistm/releases/download/1.0/keras_mnistm.pkl.gz'
self.setup_mnist(img_res)
self.setup_mnistm(img_res)
def normalize(self, images):
return images.astype(np.float32) / 127.5 - 1.
def setup_mnist(self, img_res):
print ("Setting up MNIST...")
if not os.path.exists('datasets/mnist_x.npy'):
# Load the dataset
(mnist_X, mnist_y), (_, _) = mnist.load_data()
# Normalize and rescale images
mnist_X = self.normalize(mnist_X)
mnist_X = np.array([imresize(x, img_res) for x in mnist_X])
mnist_X = np.expand_dims(mnist_X, axis=-1)
mnist_X = np.repeat(mnist_X, 3, axis=-1)
self.mnist_X, self.mnist_y = mnist_X, mnist_y
# Save formatted images
np.save('datasets/mnist_x.npy', self.mnist_X)
np.save('datasets/mnist_y.npy', self.mnist_y)
else:
self.mnist_X = np.load('datasets/mnist_x.npy')
self.mnist_y = np.load('datasets/mnist_y.npy')
print ("+ Done.")
def setup_mnistm(self, img_res):
print ("Setting up MNIST-M...")
if not os.path.exists('datasets/mnistm_x.npy'):
# Download the MNIST-M pkl file
filepath = 'datasets/keras_mnistm.pkl.gz'
if not os.path.exists(filepath.replace('.gz', '')):
print('+ Downloading ' + self.mnistm_url)
data = urllib.request.urlopen(self.mnistm_url)
with open(filepath, 'wb') as f:
f.write(data.read())
with open(filepath.replace('.gz', ''), 'wb') as out_f, \
gzip.GzipFile(filepath) as zip_f:
out_f.write(zip_f.read())
os.unlink(filepath)
# load MNIST-M images from pkl file
with open('datasets/keras_mnistm.pkl', "rb") as f:
data = pickle.load(f, encoding='bytes')
# Normalize and rescale images
mnistm_X = np.array(data[b'train'])
mnistm_X = self.normalize(mnistm_X)
mnistm_X = np.array([imresize(x, img_res) for x in mnistm_X])
self.mnistm_X, self.mnistm_y = mnistm_X, self.mnist_y.copy()
# Save formatted images
np.save('datasets/mnistm_x.npy', self.mnistm_X)
np.save('datasets/mnistm_y.npy', self.mnistm_y)
else:
self.mnistm_X = np.load('datasets/mnistm_x.npy')
self.mnistm_y = np.load('datasets/mnistm_y.npy')
print ("+ Done.")
def load_data(self, domain, batch_size=1):
X = self.mnist_X if domain == 'A' else self.mnistm_X
y = self.mnist_y if domain == 'A' else self.mnistm_y
idx = np.random.choice(list(range(len(X))), size=batch_size)
return X[idx], y[idx]
class PixelDA():
def __init__(self):
# Input shape
self.img_rows = 28
self.img_cols = 28
self.channels = 3
self.img_shape = (self.img_rows, self.img_cols, self.channels)
self.num_classes = 10
# Configure MNIST and MNIST-M data loader
self.data_loader = DataLoader(img_res=(self.img_rows, self.img_cols))
# Loss weights
lambda_adv = 10
lambda_clf = 1
# Calculate output shape of D (PatchGAN)
#patch = int(self.img_rows / 2**4)
patch = 2
self.disc_patch = (patch, patch, 1)
# Number of residual blocks in the generator
self.residual_blocks = 6
optimizer = Adam(0.0002, 0.5)
# Number of filters in first layer of discriminator and classifier
self.df = 64
self.cf = 64
# Build and compile the discriminators
self.discriminator = self.build_discriminator()
self.discriminator.compile(loss='mse',
optimizer=optimizer,
metrics=['accuracy'])
# Build the generator
self.generator = self.build_generator()
# Build the task (classification) network
self.clf = self.build_classifier()
# Input images from both domains
img_A = Input(shape=self.img_shape)
img_B = Input(shape=self.img_shape)
# Translate images from domain A to domain B
fake_B = self.generator(img_A)
# Classify the translated image
class_pred = self.clf(fake_B)
# For the combined model we will only train the generator and classifier
self.discriminator.trainable = False
# Discriminator determines validity of translated images
valid = self.discriminator(fake_B)
self.combined = Model(img_A, [valid, class_pred])
self.combined.compile(loss=['mse', 'categorical_crossentropy'],
loss_weights=[lambda_adv, lambda_clf],
optimizer=optimizer,
metrics=['accuracy'])
def build_generator(self):
"""Resnet Generator"""
def residual_block(layer_input):
"""Residual block described in paper"""
d = Conv2D(64, kernel_size=3, strides=1, padding='same')(layer_input)
d = BatchNormalization(momentum=0.8)(d)
d = Activation('relu')(d)
d = Conv2D(64, kernel_size=3, strides=1, padding='same')(d)
d = BatchNormalization(momentum=0.8)(d)
d = Add()([d, layer_input])
return d
# Image input
img = Input(shape=self.img_shape)
l1 = Conv2D(64, kernel_size=3, padding='same', activation='relu')(img)
# Propogate signal through residual blocks
r = residual_block(l1)
for _ in range(self.residual_blocks - 1):
r = residual_block(r)
output_img = Conv2D(self.channels, kernel_size=3, padding='same', activation='tanh')(r)
return Model(img, output_img)
def build_discriminator(self):
def d_layer(layer_input, filters, f_size=4, normalization=True):
"""Discriminator layer"""
d = Conv2D(filters, kernel_size=f_size, strides=2, padding='same')(layer_input)
d = LeakyReLU(alpha=0.2)(d)
if normalization:
d = InstanceNormalization()(d)
return d
img = Input(shape=self.img_shape)
d1 = d_layer(img, self.df, normalization=False)
d2 = d_layer(d1, self.df*2)
d3 = d_layer(d2, self.df*4)
d4 = d_layer(d3, self.df*8)
validity = Conv2D(1, kernel_size=4, strides=1, padding='same')(d4)
return Model(img, validity)
def build_classifier(self):
def clf_layer(layer_input, filters, f_size=4, normalization=True):
"""Classifier layer"""
d = Conv2D(filters, kernel_size=f_size, strides=2, padding='same')(layer_input)
d = LeakyReLU(alpha=0.2)(d)
if normalization:
d = InstanceNormalization()(d)
return d
img = Input(shape=self.img_shape)
c1 = clf_layer(img, self.cf, normalization=False)
c2 = clf_layer(c1, self.cf*2)
c3 = clf_layer(c2, self.cf*4)
c4 = clf_layer(c3, self.cf*8)
c5 = clf_layer(c4, self.cf*8)
class_pred = Dense(self.num_classes, activation='softmax')(Flatten()(c5))
return Model(img, class_pred)
def train(self, epochs, batch_size=128, sample_interval=50):
half_batch = int(batch_size / 2)
# Classification accuracy on 100 last batches of domain B
test_accs = []
# Adversarial ground truths
valid = np.ones((batch_size, *self.disc_patch))
fake = np.zeros((batch_size, *self.disc_patch))
for epoch in range(epochs):
# ---------------------
# Train Discriminator
# ---------------------
imgs_A, labels_A = self.data_loader.load_data(domain="A", batch_size=batch_size)
imgs_B, labels_B = self.data_loader.load_data(domain="B", batch_size=batch_size)
# Translate images from domain A to domain B
fake_B = self.generator.predict(imgs_A)
# Train the discriminators (original images = real / translated = Fake)
d_loss_real = self.discriminator.train_on_batch(imgs_B, valid)
d_loss_fake = self.discriminator.train_on_batch(fake_B, fake)
d_loss = 0.5 * np.add(d_loss_real, d_loss_fake)
# --------------------------------
# Train Generator and Classifier
# --------------------------------
# One-hot encoding of labels
labels_A = to_categorical(labels_A, num_classes=self.num_classes)
# Train the generator and classifier
g_loss = self.combined.train_on_batch(imgs_A, [valid, labels_A])
#-----------------------
# Evaluation (domain B)
#-----------------------
pred_B = self.clf.predict(imgs_B)
test_acc = np.mean(np.argmax(pred_B, axis=1) == labels_B)
# Add accuracy to list of last 100 accuracy measurements
test_accs.append(test_acc)
if len(test_accs) > 100:
test_accs.pop(0)
# Plot the progress
# print ( "%d : [D - loss: %.5f, acc: %3d%%], [G - loss: %.5f], [clf - loss: %.5f, acc: %3d%%, test_acc: %3d%% (%3d%%)]" % \
# (epoch, d_loss[0], 100*float(d_loss[1]),
# g_loss[1], g_loss[2], 100*float(g_loss[-1]),
# 100*float(test_acc), 100*float(np.mean(test_accs))))
# If at save interval => save generated image samples
if epoch % sample_interval == 0:
self.sample_images(epoch)
print ( "%d : [D - loss: %.5f, acc: %3d%%], [G - loss: %.5f], [clf - loss: %.5f, acc: %3d%%, test_acc: %3d%% (%3d%%)]" % \
(epoch, d_loss[0], 100*float(d_loss[1]),
g_loss[1], g_loss[2], 100*float(g_loss[-1]),
100*float(test_acc), 100*float(np.mean(test_accs))))
def sample_images(self, epoch):
r, c = 2, 5
imgs_A, _ = self.data_loader.load_data(domain="A", batch_size=5)
# Translate images to the other domain
fake_B = self.generator.predict(imgs_A)
gen_imgs = np.concatenate([imgs_A, fake_B])
# Rescale images 0 - 1
gen_imgs = 0.5 * gen_imgs + 0.5
#titles = ['Original', 'Translated']
fig, axs = plt.subplots(r, c)
cnt = 0
for i in range(r):
for j in range(c):
axs[i,j].imshow(gen_imgs[cnt])
#axs[i, j].set_title(titles[i])
axs[i,j].axis('off')
cnt += 1
fig.savefig("images/%d.png" % (epoch))
plt.imread("images/%d.png" % (epoch))
plt.show()
plt.close()
gan = PixelDA()
gan.train(epochs=10000, batch_size=8, sample_interval=1000)
```
| github_jupyter |
```
import os
path = os.path.join('/home/santiago/Documents/dev/reservoirpy')
import sys
sys.path.insert(0,path)
import matplotlib.pyplot as plt
import numpy as np
from zmapio import ZMAPGrid
from scipy.interpolate import griddata
from scipy.integrate import simps, trapz
from skimage import measure
import reservoirpy.volumetricspy as vc
import pyvista as pv
import seaborn as sns
z_file = ZMAPGrid('mir_sup_map')
df = z_file.to_dataframe()
df.dropna(inplace=True)
```
# Interpolate for a gridding
```
x = np.arange(df['X'].min(), df['X'].max(), 100)
y = np.arange(df['Y'].min(), df['Y'].max(), 100)
xx,yy = np.meshgrid(x,y)
z2 = griddata(df[['X','Y']].values, df[['Z']].values, (xx,yy), method='cubic')
z2 = np.squeeze(z2)
z2.shape
fig, ax = plt.subplots()
ax.contour(xx,yy,z2, levels=np.linspace(df['Z'].min(),df['Z'].max(),30))
for contour in measure.find_contours(z2,7790):
ax.plot(contour[:, 1]+df['X'].min(), contour[:, 0]+df['Y'].min(), linewidth=5, color='r')
c = measure.find_contours(z2,7760)
arr = np.empty((0,2))
for contour in c:
arr=np.concatenate((arr,contour),axis=0)
arr[:,1] = (arr[:,1]/z2.shape[1]) * (df['X'].max() - df['X'].min()) + df['X'].min()
arr[:,0] = (arr[:,0]/z2.shape[0]) * (df['Y'].max() - df['Y'].min()) + df['Y'].min()
measure.find_contours(z2,2)
fig, ax = plt.subplots()
ax.contour(xx,yy,z2, levels=np.linspace(df['Z'].min(),df['Z'].max(),30))
ax.plot(arr[:, 1], arr[:, 0], linewidth=5, color='r')
```
# Make a pivoting to make the mesh original
```
df['Z'] *= -1
p = df.pivot(index='Y',columns='X',values='Z')
p.sort_index(axis=0, inplace=True)
p.sort_index(axis=1, inplace=True)
xx,yy = np.meshgrid(p.columns,p.index)
fig, ax = plt.subplots()
ax.contour(xx,yy,p.values)
```
# Make the surface object
```
top = vc.surface(x=xx,y=yy,z=p.values)
bottom = vc.surface(x=xx,y=yy,z=p.values-10)
top.contourf(levels=5)
ss = top.structured_surface_vtk()
ss.plot(notebook=True)
levels = np.linspace(-7790,-7760,25)
top_area = top.get_contours_area(levels=levels,group=True)
bottom_area = bottom.get_contours_area(levels=levels,group=True)
contours = top.get_contours(levels=levels)
print(contours)
fig, ax = plt.subplots(figsize=(10,10))
top.contourf(ax=ax)
#sns.lineplot(x='x', y='y', data=contours, hue='level')
ax.plot(contours['x'], contours['y'], color='r')
vol=top_area.merge(bottom_area,how='outer',left_index=True,right_index=True,suffixes=['_top','_bottom']).fillna(0)
vol['dif']= vol['area_top'] - vol['area_bottom']
vol
fig, ax = plt.subplots()
ax.plot(vol.index,vol['area_top'], color='r')
ax.plot(vol.index,vol['area_bottom'], color='g')
fig, ax = plt.subplots()
ax.plot(vol.index,vol['dif'], color='r')
from scipy.integrate import simps
rv=simps(vol.index,vol['dif'])
rv
vol
```
# surface group
```
sg = vc.surface_group()
sg.add_surface({'top':top,'bottom':bottom})
rv, ar= sg.get_volume(top_surface='top',bottom_surface='bottom',levels = np.linspace(-7790,-7760,25))
print('rv ',rv)
print(ar)
x = np.linspace(10,20,100)
y = np.linspace(10,20,100)
xx, yy = np.meshgrid(x,y)
#zz = np.full(xx.shape,5)
zz = np.power(np.power(5,2) - np.power(xx-15,2) - np.power(yy-15,2),0.5)
test=vc.surface(x=xx,y=yy,z=zz)
test.structured_surface_vtk().plot(notebook=True, show_grid=True)
zmin=np.nanmin(zz)
zmax=np.nanmax(zz)
levels = np.linspace(0,10,1000)
#test.get_contours_area(c=1, levels=levels)
#test.contour(levels=levels)
rv,ar = test.get_volume(c=1,levels=levels)
rv
ar
plt.plot(ar.index,ar['area'])
yc= np.power(np.power(10,2) - np.power(x-20,2),0.5)
vc.poly_area(x,yc)
```
| github_jupyter |
## Assemble website data
This notebook saves collections in .json files in the website_data/ directory
StudySets.json
Studies.json
Recordings.json
TrueUnits.json
UnitResults.json
Sorters.json
## Schema
StudySet
* name (str)
* [type (str) -- synthetic, real, hybrid, etc.]
* [description (str)]
Study
* name (str)
* studySet (str)
* description (str)
* sorterNames (array of str)
Note: study name is unique, even across study sets
Recording
* name (str)
* study (str)
* directory (str) -- i.e., kbucket address
* description (str)
* sampleRateHz (float)
* numChannels (int)
* durationSec (float)
* numTrueUnits (int)
* [fileSizeBytes (int)]
* spikeSign (int) [Hard-coded for now. In future, grab from params.json]
TrueUnit
* unitId (int)
* recording (str)
* study (str)
* meanFiringRateHz (float)
* numEvents (int)
* peakChannel (int)
* snr (float)
SortingResult
* recording (str)
* recordingExt (str)
* study (str)
* sorter (str)
* cpuTimeSec (float)
* [runtime_info (object): timestamps, wall time, CPU time, RAM usage, error status]
* [firingsOutputUrl (str)] TODO: jfm (two weeks)
UnitResult
* unitId (int)
* recording (str)
* recordingExt (str)
* study (str)
* sorter (str)
* numMatches (int)
* numFalsePositives (int)
* numFalseNegatives (int)
* checkAccuracy (float)
* checkRecall (float)
* checkPrecision (float)
* bestSortedUnitId (int)
* spikeSprayUrl (str) TODO: jfm to make this (next week)
Sorter
* name (str)
* algorithm (str)
* [algorithmVersion (str)] - future
* processorName (str)
* processorVersion (str)
* sortingParameters (object)
```
%load_ext autoreload
%autoreload 2
from mountaintools import client as mt
import os
mt.configRemoteReadonly(collection='spikeforest', share_id='spikeforest.spikeforest2')
output_ids=['mearec_neuronexus', 'visapy_mea', 'magland_synth', 'paired', 'mearec_tetrode', 'manual_tetrode', 'bionet']
result_objects=[
mt.loadObject(
key=dict(
name='spikeforest_results'
),
subkey=output_id
)
for output_id in output_ids
]
studies=[study for X in result_objects for study in X['studies']]
recordings=[recording for X in result_objects for recording in X['recordings']]
sorting_results=[sorting_result for X in result_objects for sorting_result in X['sorting_results']]
if not os.path.exists('website_data'):
os.mkdir('website_data')
### STUDY SETS
study_sets_by_name=dict()
for study in studies:
study_sets_by_name[study['study_set']]=dict(name=study['study_set'])
StudySets=[]
for study_set in study_sets_by_name.values():
StudySets.append(dict(
name=study_set['name']
))
mt.saveObject(object=StudySets, dest_path=os.path.abspath(os.path.join('website_data', 'StudySets.json')))
print(StudySets)
### RECORDINGS and TRUE UNITS
Recordings=[]
TrueUnits=[]
for recording in recordings:
true_units_info=mt.loadObject(path=recording['summary']['true_units_info'])
for unit_info in true_units_info:
TrueUnits.append(dict(
unitId=unit_info['unit_id'],
recording=recording['name'],
recordingExt=recording['study']+':'+recording['name'],
study=recording['study'],
meanFiringRateHz=unit_info['firing_rate'],
numEvents=unit_info['num_events'],
peakChannel=unit_info['peak_channel'],
snr=unit_info['snr'],
))
Recordings.append(dict(
name=recording['name'],
study=recording['study'],
directory=recording['directory'],
description=recording['description'],
sampleRateHz=recording['summary']['computed_info']['samplerate'],
numChannels=recording['summary']['computed_info']['num_channels'],
durationSec=recording['summary']['computed_info']['duration_sec'],
numTrueUnits=len(true_units_info),
spikeSign=-1
))
mt.saveObject(object=Recordings, dest_path=os.path.abspath(os.path.join('website_data', 'Recordings.json')))
mt.saveObject(object=TrueUnits, dest_path=os.path.abspath(os.path.join('website_data', 'TrueUnits.json')))
print('Num recordings:',len(Recordings))
print('Num true units:',len(TrueUnits))
print('studies for recordings:',set([recording['study'] for recording in Recordings]))
### UNIT RESULTS and SORTING RESULTS
UnitResults=[]
SortingResults=[]
sorter_names_by_study=dict()
for sr in sorting_results:
if ('comparison_with_truth' in sr) and (sr['comparison_with_truth']):
SortingResults.append(dict(
recording=sr['recording']['name'],
study=sr['recording']['study'],
sorter=sr['sorter']['name'],
cpuTimeSec=sr['execution_stats'].get('elapsed_sec',None)
))
comparison_with_truth=mt.loadObject(path=sr['comparison_with_truth']['json'])
for unit_result in comparison_with_truth.values():
study_name=sr['recording']['study']
sorter_name=sr['sorter']['name']
if study_name not in sorter_names_by_study:
sorter_names_by_study[study_name]=set()
sorter_names_by_study[study_name].add(sorter_name)
n_match=unit_result['num_matches']
n_fp=unit_result['num_false_positives']
n_fn=unit_result['num_false_negatives']
UnitResults.append(dict(
unitId=unit_result['unit_id'],
recording=sr['recording']['name'],
recordingExt=sr['recording']['study']+':'+sr['recording']['name'],
study=study_name,
sorter=sorter_name,
numMatches=n_match,
numFalsePositives=n_fp,
numFalseNegatives=n_fn,
checkAccuracy=n_match/(n_match+n_fp+n_fn),
#checkPrecision=n_match/(n_match+n_fp),
checkRecall=n_match/(n_match+n_fn),
bestSortedUnitId=unit_result['best_unit']
))
else:
print('Warning: comparison with truth not found for sorting result: {} {}/{}', sr['sorter']['name'], sr['recording']['study'], sr['recording']['name'])
for study in sorter_names_by_study.keys():
sorter_names_by_study[study]=list(sorter_names_by_study[study])
sorter_names_by_study[study].sort()
mt.saveObject(object=UnitResults, dest_path=os.path.abspath(os.path.join('website_data', 'UnitResults.json')))
mt.saveObject(object=SortingResults, dest_path=os.path.abspath(os.path.join('website_data', 'SortingResults.json')))
print('Num unit results:',len(UnitResults))
### SORTERS
sorters_by_name=dict()
for sr in sorting_results:
sorters_by_name[sr['sorter']['name']]=sr['sorter']
Sorters=[]
for name,sorter in sorters_by_name.items():
Sorters.append(dict(
name=sorter['name'],
algorithm=sorter['processor_name'], # right now the algorithm is the same as the processor name
processorName=sorter['processor_name'],
processorVersion='0', # jfm needs to provide this
sorting_parameters=sorter['params'] # Liz, even though most sorters have similar parameter names, it won't always be like that. The params is an arbitrary json object.
))
mt.saveObject(object=Sorters, dest_path=os.path.abspath(os.path.join('website_data', 'Sorters.json')))
print([S['name'] for S in Sorters])
### STUDIES
Studies=[]
for study in studies:
Studies.append(dict(
name=study['name'],
studySet=study['study_set'],
description=study['description'],
sorterNames=sorter_names_by_study[study['name']]
# the following can be obtained from the other collections
# numRecordings, sorters, etc...
))
mt.saveObject(object=Studies, dest_path=os.path.abspath(os.path.join('website_data', 'Studies.json')))
print([S['name'] for S in Studies])
int(1.3)
def _adjust_srun_opts_for_num_jobs(srun_opts, num_workers, num_jobs):
vals = srun_opts.split()
for i in range(len(vals)):
if vals[i] == '-n' and (i+1<len(vals)):
nval = int(vals[i+1])
if num_jobs <= nval:
nval = num_jobs
num_workers = 1
elif num_jobs <= nval * (num_workers-1):
num_workers = int((num_jobs-1)/nval) + 1
vals[i+1] = str(nval)
return ' '.join(vals), num_workers
a, b = _adjust_srun_opts_for_num_jobs(srun_opts='-n 12', num_workers=4, num_jobs=37)
print(a, b)
```
| github_jupyter |
# Ansatz Sequencing: tket example
When performing variational algorithms like VQE, one common approach to generating circuit ansätze is to take an operator $U$ representing excitations and use this to act on a reference state $\lvert \phi_0 \rangle$. One such ansatz is the Unitary Coupled Cluster ansatz. Each excitation, indexed by $j$, within $U$ is given a real coefficient $a_j$ and a parameter $t_j$, such that $U = e^{i \sum_j \sum_k a_j t_j P_{jk}}$, where $P_{jk} \in \{I, X, Y, Z \}^{\otimes n}$. The exact form is dependent on the chosen qubit encoding. This excitation gives us a variational state $\lvert \psi (t) \rangle = U(t) \lvert \phi_0 \rangle$. The operator $U$ must be Trotterised, to give a product of Pauli exponentials, and converted into native quantum gates to create the ansatz circuit.<br>
<br>
This notebook will describe how to use an advanced feature of `pytket` to enable automated circuit synthesis for $U$ and reduce circuit depth dramatically.<br>
<br>
We must create a `pytket` `QubitPauliOperator`, which represents such an operator $U$, and contains a dictionary from Pauli string $P_{jk}$ to symbolic expression. Here, we make a mock operator ourselves, which resembles the UCCSD excitation operator for the $\mathrm{H}_2$ molecule using the Jordan-Wigner qubit encoding. In the future, operator generation will be handled automatically using CQC's upcoming software for enterprise quantum chemistry, EUMEN. We also offer conversion to and from the `OpenFermion` `QubitOperator` class, although at the time of writing a `QubitOperator` cannot handle arbitrary symbols.<br>
<br>
First, we create a series of `QubitPauliString` objects, which represent each $P_{jk}$.
```
from pytket.pauli import Pauli, QubitPauliString
from pytket.circuit import Qubit
q = [Qubit(i) for i in range(4)]
qps0 = QubitPauliString([q[0], q[1], q[2]], [Pauli.Y, Pauli.Z, Pauli.X])
qps1 = QubitPauliString([q[0], q[1], q[2]], [Pauli.X, Pauli.Z, Pauli.Y])
qps2 = QubitPauliString(q, [Pauli.X, Pauli.X, Pauli.X, Pauli.Y])
qps3 = QubitPauliString(q, [Pauli.X, Pauli.X, Pauli.Y, Pauli.X])
qps4 = QubitPauliString(q, [Pauli.X, Pauli.Y, Pauli.X, Pauli.X])
qps5 = QubitPauliString(q, [Pauli.Y, Pauli.X, Pauli.X, Pauli.X])
```
Now, create some symbolic expressions for the $a_j t_j$ terms.
```
from pytket.circuit import fresh_symbol
symbol1 = fresh_symbol("s0")
expr1 = 1.2 * symbol1
symbol2 = fresh_symbol("s1")
expr2 = -0.3 * symbol2
```
We can now create our `QubitPauliOperator`.
```
from pytket.utils import QubitPauliOperator
dict1 = dict((string, expr1) for string in (qps0, qps1))
dict2 = dict((string, expr2) for string in (qps2, qps3, qps4, qps5))
operator = QubitPauliOperator({**dict1, **dict2})
print(operator)
```
Now we can let `pytket` sequence the terms in this operator for us, using a selection of strategies. First, we will create a `Circuit` to generate an example reference state, and then use the `gen_term_sequence_circuit` method to append the Pauli exponentials.
```
from pytket.circuit import Circuit
from pytket.utils import gen_term_sequence_circuit
from pytket.partition import PauliPartitionStrat, GraphColourMethod
reference_circ = Circuit(4).X(1).X(3)
ansatz_circuit = gen_term_sequence_circuit(
operator, reference_circ, PauliPartitionStrat.CommutingSets, GraphColourMethod.Lazy
)
```
This method works by generating a graph of Pauli exponentials and performing graph colouring. Here we have chosen to partition the terms so that exponentials which commute are gathered together, and we have done so using a lazy, greedy graph colouring method.<br>
<br>
Alternatively, we could have used the `PauliPartitionStrat.NonConflictingSets`, which puts Pauli exponentials together so that they only require single-qubit gates to be converted into the form $e^{i \alpha Z \otimes Z \otimes ... \otimes Z}$. This strategy is primarily useful for measurement reduction, a different problem.<br>
<br>
We could also have used the `GraphColourMethod.LargestFirst`, which still uses a greedy method, but builds the full graph and iterates through the vertices in descending order of arity. We recommend playing around with the options, but we typically find that the combination of `CommutingSets` and `Lazy` allows the best optimisation.<br>
<br>
In general, not all of our exponentials will commute, so the semantics of our circuit depend on the order of our sequencing. As a result, it is important for us to be able to inspect the order we have produced. `pytket` provides functionality to enable this. Each set of commuting exponentials is put into a `CircBox`, which lets us inspect the partitoning.
```
from pytket.circuit import OpType
for command in ansatz_circuit:
if command.op.type == OpType.CircBox:
print("New CircBox:")
for pauli_exp in command.op.get_circuit():
print(
" {} {} {}".format(
pauli_exp, pauli_exp.op.get_paulis(), pauli_exp.op.get_phase()
)
)
else:
print("Native gate: {}".format(command))
```
We can convert this circuit into basic gates using a `pytket` `Transform`. This acts in place on the circuit to do rewriting, for gate translation and optimisation. We will start off with a naive decomposition.
```
from pytket.transform import Transform
naive_circuit = ansatz_circuit.copy()
Transform.DecomposeBoxes().apply(naive_circuit)
print(naive_circuit.get_commands())
```
This is a jumble of one- and two-qubit gates. We can get some relevant circuit metrics out:
```
print("Naive CX Depth: {}".format(naive_circuit.depth_by_type(OpType.CX)))
print("Naive CX Count: {}".format(naive_circuit.n_gates_of_type(OpType.CX)))
```
These metrics can be improved upon significantly by smart compilation. A `Transform` exists precisely for this purpose:
```
from pytket.transform import PauliSynthStrat, CXConfigType
smart_circuit = ansatz_circuit.copy()
Transform.UCCSynthesis(PauliSynthStrat.Sets, CXConfigType.Tree).apply(smart_circuit)
print("Smart CX Depth: {}".format(smart_circuit.depth_by_type(OpType.CX)))
print("Smart CX Count: {}".format(smart_circuit.n_gates_of_type(OpType.CX)))
```
This `Transform` takes in a `Circuit` with the structure specified above: some arbitrary gates for the reference state, along with several `CircBox` gates containing `PauliExpBox` gates.<br>
<br>
We have chosen `PauliSynthStrat.Sets` and `CXConfigType.Tree`. The `PauliSynthStrat` dictates the method for decomposing multiple adjacent Pauli exponentials into basic gates, while the `CXConfigType` dictates the structure of adjacent CX gates.<br>
<br>
If we choose a different combination of strategies, we can produce a different output circuit:
```
last_circuit = ansatz_circuit.copy()
Transform.UCCSynthesis(PauliSynthStrat.Individual, CXConfigType.Snake).apply(
last_circuit
)
print(last_circuit.get_commands())
print("Last CX Depth: {}".format(last_circuit.depth_by_type(OpType.CX)))
print("Last CX Count: {}".format(last_circuit.n_gates_of_type(OpType.CX)))
```
Other than some single-qubit Cliffords we acquired via synthesis, you can check that this gives us the same circuit structure as our `Transform.DecomposeBoxes` method! It is a suboptimal synthesis method.<br>
<br>
As with the `gen_term_sequence` method, we recommend playing around with the arguments and seeing what circuits come out. Typically we find that `PauliSynthStrat.Sets` and `CXConfigType.Tree` work the best, although routing can affect this somewhat.
| github_jupyter |
# Tabular Data Classification Example
In this example, we demonstrate how to use dlpy to build a simple neural network for tabular data classification problems.
The breast cancer dataset we used here is contained in the sklearn package. More details about the data can be found here:https://scikit-learn.org/stable/datasets/index.html#breast-cancer-dataset
Key steps involved:
1. Upload data to the server
2. Build and train a fully connected feedforward neural network model with dlpy.
3. Predict on the testing dataset.
4. Evaluate confusion matrix
5. Assess and visualize ROC curve, calculate Area under the curve(AUC).
6. Evaluate precision recall curve, f1 score and average precision score.
### 1. Import python modules
```
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
from sklearn.datasets import load_breast_cancer #The dataset is from sklearn package, with version >= 0.20.1
from swat.cas import datamsghandlers
from swat import CAS, CASTable
from dlpy import Model, Sequential
from dlpy.model import Optimizer, AdamSolver
from dlpy.layers import *
from dlpy.splitting import two_way_split
from dlpy.metrics import (accuracy_score, confusion_matrix, plot_roc,
plot_precision_recall, roc_auc_score, f1_score, average_precision_score)
%matplotlib inline
```
### 2. Connect to CAS server
```
conn = CAS('host_name', port_number)
```
### 3. Load examples dataset from scikit-learn
```
# load the breast cancer data
bc_data = load_breast_cancer()
# Get data into two pandas dataframes for the predictors(bc_x_df) and target(bc_y_df).
bc_x_df = pd.DataFrame(bc_data['data'], columns=bc_data['feature_names'])
bc_y_df = pd.DataFrame(bc_data['target'], columns=['target_class'])
# Concatenate the data into one dataframe
bc_df = pd.concat([bc_x_df, bc_y_df], axis=1)
bc_df.loc[:10]
```
There are 212 malignant tumors ('0' in the data), 357 benign tumors ('1' in the data)
```
bc_df.target_class.value_counts()
```
### 4. Upload the pandas dataframe to server
```
tbl = conn.upload_frame(bc_df, casout=dict(name='bc_df', replace=True))
```
### 5. Split the data
Here we show the train-test spliting using the `two_way_split` function in `dlpy.splitting`. It performs random sampling and split the data into `train_tbl` and `test_tbl` (on the server side). It is perferred over splitting on the local client side when the data is large, or already hosted on the server.
```
train_tbl, test_tbl = two_way_split(tbl, test_rate=30, seed=1234, stratify=False, im_table=False)
```
### 6. Build one layer fully connected neural network
Network specifications:
1. One dense layer
2. Activation function: relu
3. Number of neurons: 20
4. Output layer loss function: entropy.
5. Output layer activation function: softmax (to match with entropy).
6. number of neuron in output layer: n=2 (two classes).
```
model1 = Sequential(conn, model_table=CASTable('simple_dnn_classifier', replace=True))
model1.add(InputLayer(std='STD'))
model1.add(Dense(20, act='relu'))
model1.add(OutputLayer(act='softmax', n=2, error='entropy'))
```
### 7. Train fully connected neural network
Here we specify the optimizer to show a **scheduled training approach**:
We use `learning_rate_policy='step'` to specify stepwise learning rate decreasing policy. The step size is `step_size=5`, which means every 5 epochs, the `learning_rate` will be multiplied by factor `gamma=0.9`. In addition, we set `log_level=2` to visualize the training log.
```
optimizer = Optimizer(algorithm=AdamSolver(learning_rate=0.005, learning_rate_policy='step',
gamma=0.9, step_size=5),
mini_batch_size=4, seed=1234, max_epochs=50, log_level=2)
result = model1.fit(train_tbl, inputs=bc_x_df.columns.tolist(), nominals=['target_class'],
target='target_class', optimizer=optimizer)
# Plot the training history.
model1.plot_training_history()
```
### 8. Evaluate testing results with various classification metrics
Note: for example simplicity, we just evaluate the testing results. Feel free to check the training result by changing `test_tbl` to `train_tbl`.
```
test_result = model1.predict(test_tbl)
test_result_table = model1.valid_res_tbl
test_result_table.head()
# Show the confusion matrix
display(confusion_matrix(test_result_table['target_class'], test_result_table['I_target_class']))
# Calculate the accuracy score
acc_score = accuracy_score(test_result_table['target_class'], test_result_table['I_target_class'])
print('the accuracy score is {:.6f}'.format(acc_score))
# Plot the ROC curve for target_class = 1.
plot_roc(test_result_table['target_class'], test_result_table['P_target_class1'], pos_label=1, figsize=(6,6), linewidth=2)
# Calculate the area under the ROC curve.
auc_of_roc = roc_auc_score(test_result_table['target_class'], test_result_table['P_target_class1'], pos_label=1)
print('the area under the ROC curve is {:.6f}'.format(auc_of_roc))
# Plot the precision recall curve for target_class = 1.
plot_precision_recall(test_result_table['target_class'], test_result_table['P_target_class1'], pos_label=1, figsize=(6,6),
linewidth=2)
# Calculate the average precision score
ap = average_precision_score(test_result_table['target_class'], test_result_table['P_target_class1'], pos_label=1)
print('the average precision score is {:.6f}'.format(ap))
# Calculate the f1 score
f1sc = f1_score(test_result_table['target_class'], test_result_table['I_target_class'], pos_label=1)
print('the f1 score is {:.6f}'.format(f1sc))
```
**Note: For more classification related metrics, please see the dlpy.metrics**
| github_jupyter |
```
# Setting up the Colab page to use neqsim
%%capture
!pip install neqsim
from neqsim.process import openprocess
import urllib
import time
# Importing the gas-oil process from file
!wget https://github.com/equinor/neqsimprocess/raw/master/lib/offshorePro.neqsim
```
#Documentation
1. https://github.com/equinor/neqsimprocess/blob/master/doc/oilstabilizationprocess.pdf
2. [Exergy analysis of offshore oil and gas processing](https://ntnuopen.ntnu.no/ntnu-xmlui/handle/11250/248017)
```
# importing the process from the stored file
# See: https://github.com/equinor/neqsimprocess/blob/master/src/neqsimprocess/oilgasprocess/sepproces.java
# See Figure 1 for process flow diagram
GasOilProcess = openprocess('offshorePro.neqsim')
```
## figure oil process

```
from neqsim.process import openprocess
import time
import pandas as pd
from neqsim.thermo import createfluid,fluid_df,TPflash
# Read gas-oil process from file
# Uncomment to create alternative well stream
#wellStreamComposition = {'ComponentName': ["nitrogen", "CO2", "methane", "ethane", "propane", "i-butane", "n-butane", "i-pentane", "n-pentane", "n-hexane", "C7", "C8", "C9", "C10", "C11", "C12", "C13", "C14", "C15", "C16", "C17", "C18", "C19", "C20", "water"],
# 'MolarComposition[-]': [0.53, 3.3, 72.98, 7.68, 4.1, 0.7, 1.42, 0.54, 0.67, 0.85, 1.33, 1.33, 0.78, 0.61, 0.42, 0.33, 0.42, 0.24, 0.3, 0.17, 0.21, 0.15, 0.15, 0.8, 10.0],
# 'MolarMass[kg/mol]': [None,None, None,None,None,None,None,None,None,None,0.0913, 0.1041, 0.1188, 0.136, 0.150, 0.164, 0.179, 0.188, 0.204, 0.216, 0.236, 0.253, 0.27, 0.391, None],
# 'RelativeDensity[-]': [None,None, None,None,None,None,None,None,None,None, 0.746, 0.768, 0.79, 0.787, 0.793, 0.804, 0.817, 0.83, 0.835, 0.843, 0.837, 0.84, 0.85, 0.877, None]
# }
#
#wellStreamCompositiondf = pd.DataFrame(wellStreamComposition)
#wellStream = fluid_df(wellStreamCompositiondf)
#GasOilProcess.getUnit("well stream").setThermoSystem(wellStream)
GasOilProcess.getUnit("well stream").setFlowRate(12.23, 'MSm3/day')
GasOilProcess.getUnit("well stream").setPressure(120.0, "bara")
GasOilProcess.getUnit("well stream").setTemperature(65.0, "C")
GasOilProcess.getUnit("well stream cooler/heater").setOutTemperature(66.0, "C")
GasOilProcess.getUnit("well stream cooler/heater").setOutPressure(52.21, "bara")
GasOilProcess.getUnit("inlet choke valve").setOutletPressure(51.21, "bara")
GasOilProcess.getUnit("1st stage separator").setEntrainment(0.1, "", "oil", "aqueous")
GasOilProcess.getUnit("oil HP to MP valve").setOutletPressure(15.0, "bara")
GasOilProcess.getUnit("oil cooler/heater to 2nd stage").setOutTemperature(85.0, "C")
GasOilProcess.getUnit("oil MP to LP valve").setOutletPressure(1.8, "bara")
GasOilProcess.getUnit("Water HP to LP valve").setOutletPressure(1.01325, "bara")
GasOilProcess.getUnit("dew point scrubber cooler2").setOutTemperature(33.0, "C")
GasOilProcess.getUnit("2nd stage cooler").setOutTemperature(33.0, "C")
GasOilProcess.getUnit("1st stage gas heat exchanger").setOutTemperature(33.0, "C")
GasOilProcess.getUnit("1st stage recompressor").setIsentropicEfficiency(0.75)
GasOilProcess.getUnit("2nd stage recompressor").setIsentropicEfficiency(0.75)
#Run the process calculations
tic = time.perf_counter()
GasOilProcess.run()
toc = time.perf_counter()
print(f"Simulation run in {toc - tic:0.4f} seconds")
```
#Exergy analysis of the process
In the following simulation we will evaluate the entropy production and lost work of the total process and the individual unit operations.
```
refereceTemperature = 298.15
totalEntropyProduction = GasOilProcess.getEntropyProduction("J/K")
totalLostWork = refereceTemperature*totalEntropyProduction
totalCoolingDuty = GasOilProcess.getCoolerDuty("J/sec")/1.0e6 #MW
totalHeaterDuty = GasOilProcess.getHeaterDuty("J/sec")/1.0e6 #MW
totalPower = GasOilProcess.getPower("W")/1.0e6
print('total entropy production ', totalEntropyProduction, ' [J/K] total lost work ', totalLostWork/1.0e6, 'MW \n\n')
for unitoperation in GasOilProcess.getUnitOperations():
print('name: ', unitoperation.getName(), ' entropy production ', unitoperation.getEntropyProduction("J/K"), ' [J/K] lost work ', unitoperation.getEntropyProduction("J/K")*refereceTemperature/1.0e6, ' [MW]')
```
| github_jupyter |
## Analysis
In this notebook, we generate some of the summary statistics for the THOR runs on both simulations and ZTF alerts.
Data and results files for this notebook may be downloaded [here](https://dirac.astro.washington.edu/~moeyensj/projects/thor/paper1/).
```
%load_ext autoreload
%autoreload 2
import glob
import os
import numpy as np
import pandas as pd
import sqlite3 as sql
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.colors as colors
import seaborn as sns
sns.set(font_scale=1.0, context="paper", style="ticks")
sns.set_palette("viridis")
from astropy.time import Time
%matplotlib inline
import thor
from thor import __version__
print("THOR version: {}".format(__version__))
import difi
from difi import __version__
print("difi version: {}".format(__version__))
def createComponentSummary(
all_linkages,
all_truths,
components=["clustering", "iod", "od", "od+a"]
):
linkage_types = ["mixed", "partial", "pure", "pure_complete"]
summary_quantities = [
"component",
"num_linkages", "num_mixed", "num_partial", "num_pure", "num_pure_complete",
"num_findable", "num_found", "num_found_pure", "num_found_partial"
]
component_summary = {q : [] for q in summary_quantities}
for component in components:
component_mask_all_linkages = (all_linkages["component"] == component)
component_mask_all_truths = (all_truths["component"] == component)
component_summary["component"].append(component)
# Calculate the total number of different linkage types
num_total_linkages = len(all_linkages[component_mask_all_linkages])
component_summary["num_linkages"].append(num_total_linkages)
for linkage_type in linkage_types:
num_linkages = all_linkages[component_mask_all_linkages][linkage_type].sum()
component_summary[f"num_{linkage_type}"].append(num_linkages)
# Calculate the total numbers of objects found, findable, etc..
num_findable = all_truths[component_mask_all_truths & (all_truths["findable"] == 1)]["obj_id"].nunique()
component_summary["num_findable"].append(num_findable)
num_found_pure = all_truths[component_mask_all_truths & (all_truths["found_pure"] >= 1)]["obj_id"].nunique()
component_summary["num_found_pure"].append(num_found_pure)
num_found_partial = all_truths[component_mask_all_truths & (all_truths["found_partial"] >= 1)]["obj_id"].nunique()
component_summary["num_found_partial"].append(num_found_partial)
num_found = all_truths[component_mask_all_truths & (all_truths["found"] >= 1)]["obj_id"].nunique()
component_summary["num_found"].append(num_found)
component_summary = pd.DataFrame(component_summary)
component_summary["completeness"] = 100 * component_summary["num_found"] / component_summary["num_findable"]
component_summary["completeness_pure"] = 100 * component_summary["num_found_pure"] / component_summary["num_findable"]
component_summary["completeness_partial"] = 100 * component_summary["num_found_partial"] / component_summary["num_findable"]
component_summary["purity"] = 100 * component_summary["num_pure"] / component_summary["num_linkages"]
component_summary["linkage_efficiency"] = 100 * component_summary["num_found_pure"] / component_summary["num_linkages"]
return component_summary
```
### Simulations
```
DATA_DIR = "/mnt/data/projects/thor/thor_data/msst_4x4/"
preprocessed_observations = pd.read_csv(
os.path.join(DATA_DIR, "preprocessed_observations.csv"),
index_col=False,
dtype={"obs_id" : str}
)
preprocessed_associations = pd.read_csv(
os.path.join(DATA_DIR, "preprocessed_associations.csv"),
index_col=False,
dtype={"obs_id" : str}
)
RUN_DIR = "/mnt/data/projects/thor/thor_results/msst_4x4/v1.1/run_4/"
DATABASE = "/home/moeyensj/projects/thor/thor_data/msst_4x4/msst_survey.db"
con = sql.connect(DATABASE)
known_orbits = pd.read_sql("SELECT * FROM mpcOrbitCat", con)
known_orbits = known_orbits[known_orbits["designation"].isin(preprocessed_associations["obj_id"].unique())]
con.close()
sma_bins = [0, 1.7, 2.06, 2.5, 2.82, 2.95, 3.27, 5.0, 50, 1000.0]
classes = {}
for i, (bin_start, bin_end) in enumerate(zip(sma_bins[:-1], sma_bins[1:])):
bin_mask = (known_orbits["a_au"] >= bin_start) & (known_orbits["a_au"] < bin_end)
classes["{}<=a<{}".format(bin_start, bin_end)] = known_orbits[bin_mask]["designation"].unique()
classes["Noise"] = preprocessed_associations[preprocessed_associations["obj_id"].str.contains("^u[0-9]{8}$", regex=True)]["obj_id"].unique()
from thor.orbits import Orbits
from thor import analyzeTHOR
ANALYSIS_DIR = os.path.join(RUN_DIR, "analysis")
os.makedirs(ANALYSIS_DIR, exist_ok=True)
# Read recovered orbits and orbit members
recovered_orbits = Orbits.from_csv(
os.path.join(RUN_DIR, "recovered_orbits.csv"),
).to_df(include_units=False)
recovered_orbit_members = pd.read_csv(
os.path.join(RUN_DIR, "recovered_orbit_members.csv"),
index_col=False,
dtype={"obs_id" : str}
)
if not os.path.exists(os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv")):
# Analyze THOR run
run_analysis, test_orbit_analysis = analyzeTHOR(
preprocessed_associations,
RUN_DIR,
classes=classes
)
all_orbits_recovered, all_truths_recovered, summary_recovered = run_analysis
all_linkages, all_truths, summary = test_orbit_analysis
# Compute component summary
summary_components = createComponentSummary(all_linkages, all_truths)
all_orbits_recovered.to_csv(
os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv"),
index=False
)
all_truths_recovered.to_csv(
os.path.join(ANALYSIS_DIR, "all_truths_recovered.csv"),
index=False
)
summary_recovered.to_csv(
os.path.join(ANALYSIS_DIR, "summary_recovered.csv"),
index=False
)
all_linkages.to_csv(
os.path.join(ANALYSIS_DIR, "all_linkages.csv"),
index=False
)
all_truths.to_csv(
os.path.join(ANALYSIS_DIR, "all_truths.csv"),
index=False
)
summary.to_csv(
os.path.join(ANALYSIS_DIR, "summary.csv"),
index=False
)
summary_components.to_csv(
os.path.join(ANALYSIS_DIR, "summary_components.csv"),
index=False
)
else:
all_orbits_recovered = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv"),
index_col=False
)
all_truths_recovered = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_truths_recovered.csv"),
index_col=False
)
summary_recovered = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary_recovered.csv"),
index_col=False
)
all_linkages = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_linkages.csv"),
index_col=False
)
all_truths = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_truths.csv"),
index_col=False
)
summary = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary.csv"),
index_col=False
)
summary_components = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary_components.csv"),
index_col=False
)
# Number of noise detections
num_noise_obs = len(preprocessed_associations[preprocessed_associations["obj_id"].str.contains("^u[0-9]{8}$", regex=True)])
num_detections = len(preprocessed_associations)
percent_noise = 100 * num_noise_obs / num_detections
print(f"{num_noise_obs} [{percent_noise:.3f}%]")
# Number of real detections
num_object_obs = len(preprocessed_associations[~preprocessed_associations["obj_id"].str.contains("^u[0-9]{8}$", regex=True)])
percent_object = 100 * num_object_obs / num_detections
print(f"{num_object_obs} [{percent_object:.3f}%]")
summary_components
summary_recovered
findable = all_truths_recovered[all_truths_recovered["findable"] == 1]["obj_id"].values
found = all_truths_recovered[all_truths_recovered["found"] >= 1]["obj_id"].values
sma_bins = [0.0, 1.7, 50.0]
for a_min, a_max in zip(sma_bins[:-1], sma_bins[1:]):
found_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(found)])
findable_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(findable)])
print("Completeness between {} and {} AU: {:.2f} %, {}".format(a_min, a_max, found_in_bin / findable_in_bin * 100.0, found_in_bin))
sma_bins = [0.0, 2.5, 50.0]
for a_min, a_max in zip(sma_bins[:-1], sma_bins[1:]):
found_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(found)])
findable_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(findable)])
print("Completeness between {} and {} AU: {:.2f} %, {}".format(a_min, a_max, found_in_bin / findable_in_bin * 100.0, found_in_bin))
```
### ZTF Analysis with 2018 Catalog
```
DATA_DIR = "/mnt/data/projects/thor/thor_data/ztf/"
RUN_DIR = "/mnt/data/projects/thor/thor_results/ztf/v1.1/run4"
preprocessed_observations = pd.read_csv(
os.path.join(DATA_DIR, "preprocessed_observations.csv"),
index_col=False,
dtype={"obs_id" : str}
)
preprocessed_associations = pd.read_csv(
os.path.join(DATA_DIR, "preprocessed_associations.csv"),
index_col=False,
dtype={"obs_id" : str}
)
# Read original observations
observations = pd.read_csv(
os.path.join(DATA_DIR, "ztf_observations_610_624.csv"),
index_col=False,
sep=" ",
dtype={"candid" : str},
low_memory=False
)
# Add magnitudes, magnitude errors, filter IDs, night IDs to the preprocessed observations
preprocessed_observations = preprocessed_observations.merge(
observations[["candid", "nid", "magpsf", "sigmapsf", "fid"]],
left_on="obs_id",
right_on="candid"
)
preprocessed_observations.drop(
columns=["candid"],
inplace=True
)
preprocessed_observations.rename(
columns={
"nid" : "night_id",
"magpsf" : "mag",
"sigmapsf" : "mag_sigma",
"fid" : "filter"
},
inplace=True
)
# Filter ID (1=g; 2=r; 3=i) from https://zwickytransientfacility.github.io/ztf-avro-alert/schema.html
for i, f in enumerate(["g", "r", "i"]):
preprocessed_observations.loc[preprocessed_observations["filter"].isin([i + 1]), "filter"] = f
preprocessed_observations.head()
from thor.utils import unpackMPCDesignation
# Read orbits file (MPCORB in OORB format from 2018)
known_orbits = pd.read_csv(
"/mnt/data/projects/thor/thor_data/ztf/MPCORB_20181106_ZTF_keplerian.orb",
delim_whitespace=True,
skiprows=4,
names=["designation", "a_au", "e", "i_deg", "ascNode_deg", "argPeri_deg", "meanAnom_deg", "epoch_mjd_tt", "H", "G"],
low_memory=False
)
known_orbits.loc[:, "designation"] = known_orbits["designation"].apply(unpackMPCDesignation)
known_orbits = known_orbits[known_orbits["designation"].isin(preprocessed_associations["obj_id"].unique())]
known_orbits
sma_bins = [0, 1.7, 2.06, 2.5, 2.82, 2.95, 3.27, 5.0, 50, 1000.0]
classes = {}
for i, (bin_start, bin_end) in enumerate(zip(sma_bins[:-1], sma_bins[1:])):
bin_mask = (known_orbits["a_au"] >= bin_start) & (known_orbits["a_au"] < bin_end)
classes["{}<=a<{}".format(bin_start, bin_end)] = known_orbits[bin_mask]["designation"].unique()
# Observations unattributed by ZTF
classes["Unknown"] = preprocessed_associations[preprocessed_associations["obj_id"].str.contains("^u[0-9]{8}$", regex=True)]["obj_id"].unique()
# Observations attributed by ZTF that could not be matched the known catalog (probably designation changes or comets)
unclassified_mask = ~preprocessed_associations["obj_id"].isin(known_orbits["designation"].unique()) & (~preprocessed_associations["obj_id"].str.contains("^u[0-9]{8}$", regex=True))
classes["Unmatched"] = preprocessed_associations[unclassified_mask]["obj_id"].unique()
from thor.orbits import Orbits
from difi import analyzeLinkages
from difi import analyzeObservations
ANALYSIS_DIR = os.path.join(RUN_DIR, "analysis_2018")
os.makedirs(ANALYSIS_DIR, exist_ok=True)
# Read the recovered orbits and recovered_orbit_members which where combined from
# the patches to see how the overall run performed
recovered_orbits = Orbits.from_csv(
os.path.join(RUN_DIR, "recovered_orbits.csv"),
).to_df(include_units=False)
recovered_orbit_members = pd.read_csv(
os.path.join(RUN_DIR, "recovered_orbit_members.csv"),
index_col=False,
dtype={"obs_id" : str}
)
column_mapping = {
'linkage_id': 'orbit_id',
'obs_id': 'obs_id',
'truth': 'obj_id'
}
analysis_observations = preprocessed_observations.merge(preprocessed_associations, on="obs_id")
all_truths_survey, findable_observations, summary_survey = analyzeObservations(
analysis_observations,
classes=classes,
metric='min_obs',
column_mapping=column_mapping,
min_obs=5,
)
if not os.path.exists(os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv")):
# Go through each individual patch directory (which themselves
# are individual THOR runs) and analyze the performance
all_linkages_patches = []
all_truths_patches = []
summary_patches = []
contents = sorted(glob.glob(os.path.join(RUN_DIR, "patch_*")))
for c in contents:
if os.path.isdir(c):
print(f"Analyzing Patch {os.path.basename(c)}")
run_analysis, test_orbit_analysis = analyzeTHOR(
preprocessed_associations,
c,
classes=classes,
)
all_linkages_patches_i, all_truths_patches_i, summary_patches_i = test_orbit_analysis
all_linkages_patches.append(all_linkages_patches_i)
all_truths_patches.append(all_truths_patches_i)
summary_patches.append(summary_patches_i)
all_linkages_patches = pd.concat(
all_linkages_patches,
ignore_index=True
)
all_truths_patches = pd.concat(
all_truths_patches,
ignore_index=True
)
summary_patches = pd.concat(
summary_patches,
ignore_index=True
)
all_orbits_recovered, all_truths_recovered, summary_recovered = analyzeLinkages(
analysis_observations,
recovered_orbit_members,
all_truths=all_truths_survey,
min_obs=5,
contamination_percentage=0.0,
classes=classes,
column_mapping=column_mapping
)
for df in [all_orbits_recovered, all_truths_recovered, summary_recovered]:
df.insert(0, "component", "combined")
all_linkages = pd.concat([all_linkages_patches, all_orbits_recovered], ignore_index=True)
all_truths = pd.concat([all_truths_patches, all_truths_recovered], ignore_index=True)
summary = pd.concat([summary_patches, summary_recovered], ignore_index=True)
for df in [all_orbits_recovered, all_truths_recovered, summary_recovered]:
df.drop(columns=["component"], inplace=True)
summary_components = createComponentSummary(
all_linkages,
all_truths,
components=["clustering", "iod", "od", "od+a", "combined"]
)
all_orbits_recovered.to_csv(
os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv"),
index=False
)
all_truths_recovered.to_csv(
os.path.join(ANALYSIS_DIR, "all_truths_recovered.csv"),
index=False
)
summary_recovered.to_csv(
os.path.join(ANALYSIS_DIR, "summary_recovered.csv"),
index=False
)
all_linkages.to_csv(
os.path.join(ANALYSIS_DIR, "all_linkages.csv"),
index=False
)
all_truths.to_csv(
os.path.join(ANALYSIS_DIR, "all_truths.csv"),
index=False
)
summary.to_csv(
os.path.join(ANALYSIS_DIR, "summary.csv"),
index=False
)
summary_components.to_csv(
os.path.join(ANALYSIS_DIR, "summary_components.csv"),
index=False
)
else:
all_orbits_recovered = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv"),
index_col=False
)
all_truths_recovered = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_truths_recovered.csv"),
index_col=False
)
summary_recovered = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary_recovered.csv"),
index_col=False
)
all_linkages = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_linkages.csv"),
index_col=False
)
all_truths = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_truths.csv"),
index_col=False
)
summary = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary.csv"),
index_col=False
)
summary_components = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary_components.csv"),
index_col=False
)
# Number of noise detections
num_noise_obs = len(preprocessed_associations[preprocessed_associations["obj_id"].str.contains("^u[0-9]{8}$", regex=True)])
num_detections = len(preprocessed_associations)
percent_noise = 100 * num_noise_obs / num_detections
print(f"{num_noise_obs} [{percent_noise:.3f}%]")
# Number of real detections
num_object_obs = len(preprocessed_associations[~preprocessed_associations["obj_id"].str.contains("^u[0-9]{8}$", regex=True)])
percent_object = 100 * num_object_obs / num_detections
print(f"{num_object_obs} [{percent_object:.3f}%]")
summary_recovered
mask = ((~summary["test_orbit_id"].isna()) & (summary["class"] == "All") & (summary["component"] == "clustering"))
len(summary[mask & (summary["findable"] == 0)])
mask = ((~summary["test_orbit_id"].isna()) & (summary["class"] == "All") & (summary["component"] == "od+a"))
len(summary[mask & (summary["found"] == 0)])
summary_components
duplicate_orbits = all_orbits_recovered["linked_truth"].value_counts()
len(duplicate_orbits.index.values[duplicate_orbits.values > 1]), duplicate_orbits.values[duplicate_orbits.values > 1].sum()
def addNumNights(
linkages,
linkage_members,
preprocessed_observations,
linkage_id_col="orbit_id"
):
linkages_ = linkages.copy()
linkage_members_ = linkage_members.copy()
linkage_members_ = linkage_members_.merge(
preprocessed_observations,
on="obs_id"
)
linkages_ = linkages_.merge(
linkage_members_.groupby(by=[linkage_id_col])["night_id"].nunique().to_frame("num_nights").reset_index(),
on="orbit_id"
)
return linkages_, linkage_members_
def calculateDeltas(
linkage_members,
preprocessed_observations
):
linkage_members_ = linkage_members.copy()
deltas = linkage_members_.groupby(by=["orbit_id", "night_id"])[["mjd_utc", "RA_deg", "Dec_deg", "mag"]].diff()
deltas["mjd_utc"] = deltas["mjd_utc"].values
deltas.rename(
columns={
"mjd_utc" : "dt",
"RA_deg" : "dRA_deg",
"Dec_deg" : "dDec_deg",
"mag" : "dmag",
},
inplace=True
)
linkage_members_ = linkage_members_.join(deltas)
linkage_members_["dt_sec"] = linkage_members_["dt"] * 86400
return linkage_members_
recovered_orbits, recovered_orbit_members = addNumNights(recovered_orbits, recovered_orbit_members, preprocessed_observations)
recovered_orbit_members = calculateDeltas(recovered_orbit_members, preprocessed_observations)
analysis_orbit_members = recovered_orbit_members.merge(preprocessed_associations, on="obs_id")
analysis_orbit_members.head()
recovered_orbits
len(recovered_orbits[recovered_orbits["num_nights"] == recovered_orbits["num_obs"]])
pure_orbits_2018 = all_orbits_recovered[all_orbits_recovered["pure"] == 1]["orbit_id"].values
mixed_orbits_2018 = all_orbits_recovered[all_orbits_recovered["mixed"] == 1]["orbit_id"].values
# Total number of observations recovered
analysis_orbit_members["obs_id"].nunique()
# Total number of observations recovered in pure orbits
analysis_orbit_members[analysis_orbit_members["orbit_id"].isin(pure_orbits_2018)]["obs_id"].nunique()
mixed_orbit_members = analysis_orbit_members[analysis_orbit_members["orbit_id"].isin(mixed_orbits_2018)]
unknown_observations_in_mixed = mixed_orbit_members[mixed_orbit_members["obj_id"].str.contains("^u[0-9]{8}$", regex=True)]
known_observations_in_mixed = mixed_orbit_members[~mixed_orbit_members["obj_id"].str.contains("^u[0-9]{8}$", regex=True)]
len(unknown_observations_in_mixed)
len(known_observations_in_mixed)
unknown_observations_in_mixed["orbit_id"].nunique(), unknown_observations_in_mixed["obj_id"].nunique()
known_observations_in_mixed["orbit_id"].nunique(), known_observations_in_mixed["obj_id"].nunique()
```
How would MOPS and ZMODE perform?
```
from difi import analyzeObservations
column_mapping = {
'obs_id': 'obs_id',
'truth': 'obj_id',
'night' : 'night_id',
'time' : 'mjd_utc'
}
all_truths_survey_MOPS, findable_observations_MOPS, summary_survey_MOPS = analyzeObservations(
analysis_observations,
classes=None,
metric='nightly_linkages',
linkage_min_obs=2,
max_obs_separation=3.0/24,
min_linkage_nights=3,
column_mapping=column_mapping
)
summary_survey_MOPS["findable"].values[0]
summary_survey_MOPS["findable"].values[0] / summary_recovered[summary_recovered["class"] == "All"]["found"].values[0]
percent_mops = 100 * summary_recovered[summary_recovered["class"] == "All"]["found"].values[0] / summary_survey_MOPS["findable"].values[0]
print(f"THOR discovery potential over ideal MOPS: {percent_mops:.3f}%")
def calcDiscoverableZMODE(
observations,
min_tracklets=2,
tracklet_min_obs=2,
min_obs_per_track=4,
max_track_night_span=4,
column_mapping=column_mapping
):
# Count number of observations per object
obs_per_obj = observations[column_mapping["truth"]].value_counts()
possibly_findable = obs_per_obj.index.values[obs_per_obj >= min_obs_per_track]
night_designation_count = observations.groupby(by=[column_mapping["night"]])[column_mapping["truth"]].value_counts()
night_designation_count = pd.DataFrame(night_designation_count)
night_designation_count.rename(columns={"obj_id": "num_obs"}, inplace=True)
night_designation_count.reset_index(inplace=True)
night_designation_count["delta_night"] = night_designation_count.groupby([column_mapping["truth"]])[column_mapping["night"]].diff()
night_designation_count.loc[night_designation_count["delta_night"].isna(), "delta_night"] = 0
night_designation_count["possible_tracklet"] = np.where(night_designation_count["num_obs"] >= tracklet_min_obs, 1, 0)
night_designation_count = night_designation_count[night_designation_count["delta_night"] < max_track_night_span]
tracklets_per_designation = night_designation_count.groupby(by=[column_mapping["truth"]])["possible_tracklet"].sum()
possibly_findable = tracklets_per_designation.index.values[tracklets_per_designation >= min_tracklets]
obs_per_designation = night_designation_count[night_designation_count[column_mapping["truth"]].isin(possibly_findable)].groupby([column_mapping["truth"]])["num_obs"].sum()
return obs_per_designation.index.values[obs_per_designation >= min_obs_per_track]
findableZMODE = calcDiscoverableZMODE(analysis_observations)
len(findableZMODE)
len(findableZMODE) / summary_recovered[summary_recovered["class"] == "All"]["found"].values[0]
percent_zmode = 100 * summary_recovered[summary_recovered["class"] == "All"]["found"].values[0] / len(findableZMODE)
print(f"THOR discovery potential over ideal ZMODE: {percent_zmode:.3f}%")
findable = all_truths_recovered[all_truths_recovered["findable"] == 1]["obj_id"].values
found = all_truths_recovered[all_truths_recovered["found"] >= 1]["obj_id"].values
sma_bins = [0.0, 1.7, 50.0]
for a_min, a_max in zip(sma_bins[:-1], sma_bins[1:]):
found_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(found)])
findable_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(findable)])
print("Completeness between {} and {} AU: {:.2f} %, {}".format(a_min, a_max, found_in_bin / findable_in_bin * 100.0, found_in_bin))
sma_bins = [0.0, 2.5, 50.0]
for a_min, a_max in zip(sma_bins[:-1], sma_bins[1:]):
found_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(found)])
findable_in_bin = len(known_orbits[(known_orbits["a_au"] >= a_min) & (known_orbits["a_au"] < a_max) & known_orbits["designation"].isin(findable)])
print("Completeness between {} and {} AU: {:.2f} %, {}".format(a_min, a_max, found_in_bin / findable_in_bin * 100.0, found_in_bin))
```
### ZTF Analysis with 2021 Catalog
```
DATA_DIR = "/mnt/data/projects/thor/thor_data/ztf/"
preprocessed_associations_2021 = pd.read_csv(
os.path.join(DATA_DIR, "preprocessed_associations_20210420_3arcsec.csv"),
index_col=False,
dtype={"obs_id" : str}
)
RUN_DIR = "/mnt/data/projects/thor/thor_results/ztf/v1.1/run4"
# Read orbits file from 2021
known_orbits = pd.read_csv(
"/mnt/data/projects/thor/thor_data/ztf/MPCORB_20210420.csv",
index_col=False,
low_memory=False
)
known_orbits = known_orbits[known_orbits["designation"].isin(preprocessed_associations_2021["obj_id"].unique())]
known_orbits
sma_bins = [0, 1.7, 2.06, 2.5, 2.82, 2.95, 3.27, 5.0, 50, 1000.0]
classes_2021 = {}
for i, (bin_start, bin_end) in enumerate(zip(sma_bins[:-1], sma_bins[1:])):
bin_mask = (known_orbits["a_au"] >= bin_start) & (known_orbits["a_au"] < bin_end)
classes_2021["{}<=a<{}".format(bin_start, bin_end)] = known_orbits[bin_mask]["designation"].unique()
# Observations unattributed by ZTF
classes_2021["Unknown"] = preprocessed_associations_2021[preprocessed_associations_2021["obj_id"].str.contains("^u[0-9]{8}$", regex=True)]["obj_id"].unique()
# Observations attributed by ZTF that could not be matched the known catalog (probably designation changes or comets)
unclassified_mask = ~preprocessed_associations_2021["obj_id"].isin(known_orbits["designation"].unique()) & (~preprocessed_associations_2021["obj_id"].str.contains("^u[0-9]{8}$", regex=True))
classes_2021["Unmatched"] = preprocessed_associations_2021[unclassified_mask]["obj_id"].unique()
from thor.orbits import Orbits
from difi import analyzeLinkages
from difi import analyzeObservations
ANALYSIS_DIR = os.path.join(RUN_DIR, "analysis_2021")
os.makedirs(ANALYSIS_DIR, exist_ok=True)
column_mapping = {
'linkage_id': 'orbit_id',
'obs_id': 'obs_id',
'truth': 'obj_id'
}
analysis_observations_2021 = preprocessed_observations.merge(preprocessed_associations_2021, on="obs_id")
all_truths_survey_2021, findable_observations_2021, summary_survey_2021 = analyzeObservations(
analysis_observations_2021,
classes=classes_2021,
metric='min_obs',
column_mapping=column_mapping,
min_obs=5,
)
if not os.path.exists(os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv")):
# Go through each individual patch directory (which themselves
# are individual THOR runs) and analyze the performance
all_linkages_patches_2021 = []
all_truths_patches_2021 = []
summary_patches_2021 = []
contents = sorted(glob.glob(os.path.join(RUN_DIR, "patch_*")))
for c in contents:
if os.path.isdir(c):
print(f"Analyzing Patch {os.path.basename(c)}")
run_analysis, test_orbit_analysis = analyzeTHOR(
preprocessed_associations_2021,
c,
classes=classes_2021,
)
all_linkages_patches_i, all_truths_patches_i, summary_patches_i = test_orbit_analysis
all_linkages_patches_2021.append(all_linkages_patches_i)
all_truths_patches_2021.append(all_truths_patches_i)
summary_patches_2021.append(summary_patches_i)
all_linkages_patches_2021 = pd.concat(
all_linkages_patches_2021,
ignore_index=True
)
all_truths_patches_2021 = pd.concat(
all_truths_patches_2021,
ignore_index=True
)
summary_patches_2021 = pd.concat(
summary_patches_2021,
ignore_index=True
)
all_orbits_recovered_2021, all_truths_recovered_2021, summary_recovered_2021 = analyzeLinkages(
analysis_observations_2021,
recovered_orbit_members,
all_truths=all_truths_survey_2021,
min_obs=5,
contamination_percentage=0.0,
classes=classes_2021,
column_mapping=column_mapping
)
for df in [all_orbits_recovered_2021, all_truths_recovered_2021, summary_recovered_2021]:
df.insert(0, "component", "combined")
all_linkages_2021 = pd.concat([all_linkages_patches_2021, all_orbits_recovered_2021], ignore_index=True)
all_truths_2021 = pd.concat([all_truths_patches_2021, all_truths_recovered_2021], ignore_index=True)
summary_2021 = pd.concat([summary_patches_2021, summary_recovered_2021], ignore_index=True)
for df in [all_orbits_recovered_2021, all_truths_recovered_2021, summary_recovered_2021]:
df.drop(columns=["component"], inplace=True)
summary_components_2021 = createComponentSummary(
all_linkages_2021,
all_truths_2021,
components=["clustering", "iod", "od", "od+a", "combined"]
)
all_orbits_recovered_2021.to_csv(
os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv"),
index=False
)
all_truths_recovered_2021.to_csv(
os.path.join(ANALYSIS_DIR, "all_truths_recovered.csv"),
index=False
)
summary_recovered_2021.to_csv(
os.path.join(ANALYSIS_DIR, "summary_recovered.csv"),
index=False
)
all_linkages_2021.to_csv(
os.path.join(ANALYSIS_DIR, "all_linkages.csv"),
index=False
)
all_truths_2021.to_csv(
os.path.join(ANALYSIS_DIR, "all_truths.csv"),
index=False
)
summary_2021.to_csv(
os.path.join(ANALYSIS_DIR, "summary.csv"),
index=False
)
summary_components_2021.to_csv(
os.path.join(ANALYSIS_DIR, "summary_components.csv"),
index=False
)
else:
all_orbits_recovered_2021 = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_orbits_recovered.csv"),
index_col=False
)
all_truths_recovered_2021 = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_truths_recovered.csv"),
index_col=False
)
summary_recovered_2021 = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary_recovered.csv"),
index_col=False
)
all_linkages_2021 = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_linkages.csv"),
index_col=False
)
all_truths_2021 = pd.read_csv(
os.path.join(ANALYSIS_DIR, "all_truths.csv"),
index_col=False
)
summary_2021 = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary.csv"),
index_col=False
)
summary_components_2021 = pd.read_csv(
os.path.join(ANALYSIS_DIR, "summary_components.csv"),
index_col=False
)
summary_components
summary_components_2021
summary_recovered
summary_recovered_2021
```
Filtering out suspect orbits for 2018 and 2021
```
mixed_orbits_2021 = all_orbits_recovered_2021[all_orbits_recovered_2021["mixed"] == 1]["orbit_id"].unique()
pure_orbits_2021 = all_orbits_recovered_2021[all_orbits_recovered_2021["pure"] == 1]["orbit_id"].unique()
analysis_orbit_members_2021 = recovered_orbit_members.merge(preprocessed_associations_2021, on="obs_id")
# Remove observations that are made within 5 minutes (using 2018)
high_quality_mask = (
analysis_orbit_members["orbit_id"].isin(mixed_orbits_2018)
& ((analysis_orbit_members["dt_sec"].isna()) | (analysis_orbit_members["dt_sec"] > 1800))
)
occurences = analysis_orbit_members[high_quality_mask]["orbit_id"].value_counts()
orbit_ids_keep = occurences.index.values[occurences.values >= 5]
print(len(orbit_ids_keep))
mixed_orbits_high_quality_2018 = orbit_ids_keep
# Remove observations of known objects (using 2018)
high_quality_mask_2 = (
high_quality_mask & analysis_orbit_members["obj_id"].str.contains("^u[0-9]{8}$", regex=True)
)
occurences = analysis_orbit_members[high_quality_mask_2]["orbit_id"].value_counts()
orbit_ids_keep = occurences.index.values[occurences.values >= 5]
print(len(orbit_ids_keep))
mixed_orbits_high_quality_2018 = orbit_ids_keep
print("Known in 2021")
known_2021 = recovered_orbits[
recovered_orbits["orbit_id"].isin(pure_orbits_2021)
]
print(len(known_2021), all_orbits_recovered_2021[all_orbits_recovered_2021["orbit_id"].isin(known_2021["orbit_id"].values)]["linked_truth"].nunique())
print("Unknown in 2021")
unknown_2021 = recovered_orbits[
recovered_orbits["orbit_id"].isin(mixed_orbits_2021)
]
print(len(unknown_2021))
print("Unknown in 2018, Found in 2021")
found_since_2018 = recovered_orbits[
(recovered_orbits["orbit_id"].isin(mixed_orbits_2018)
& recovered_orbits["orbit_id"].isin(pure_orbits_2021))
]
print(len(found_since_2018))
print("Hiqh Quality Unknown in 2018, Found in 2021")
found_since_2018_high_quality = recovered_orbits[
(recovered_orbits["orbit_id"].isin(mixed_orbits_high_quality_2018)
& recovered_orbits["orbit_id"].isin(pure_orbits_2021))
]
print(len(found_since_2018_high_quality))
print("Hiqh Quality Unknown in 2018, Unknown in 2021")
unknown_2021_high_quality = recovered_orbits[
(recovered_orbits["orbit_id"].isin(mixed_orbits_high_quality_2018)
& (~recovered_orbits["orbit_id"].isin(pure_orbits_2021)))
]
print(len(unknown_2021_high_quality))
Orbits.from_df(unknown_2021_high_quality).to_csv(
os.path.join(RUN_DIR, "discovery_candidates.csv")
)
unknown_2021_high_quality
import astropy.units as u
from astroquery.imcce import Skybot
from astropy.coordinates import SkyCoord
from astropy.time import Time
def createMPCCheckerQuery(observation):
assert len(observation) == 1
# Configure RA, Dec
coords = SkyCoord(
ra=observation["RA_deg"].values[:1]*u.deg,
dec=observation["Dec_deg"].values[:1]*u.deg
)
ra_hms = "{:02.0f} {:02.0f} {:05.2f}".format(*coords.ra[0].hms)
dec_dms = "{:+03.0f} {:02.0f} {:05.2f}".format(*coords.dec[0].dms)
# Configure observation time
observation_time = Time(
observation["mjd_utc"].values[0],
scale="utc",
format="mjd"
)
decimal_day = np.modf(observation["mjd_utc"].values[0])[0]
time = "{} {} {}{}".format(
*observation_time.utc.isot.split("T")[0].split("-"),
"{:.2f}".format(decimal_day).lstrip("0")
)
observatory_code = observation["observatory_code"].values[0]
return (ra_hms, dec_dms, time, observatory_code)
def querySkyBot(observations, radius=50*u.arcsecond):
# Configure RA, Dec
coords = SkyCoord(
ra=observations["RA_deg"].values*u.deg,
dec=observations["Dec_deg"].values*u.deg
)
# Configure observation time
observation_times = Time(
observations["mjd_utc"].values,
scale="utc",
format="mjd"
)
observatory_codes = observations["observatory_code"].values
obs_ids = observations["obs_id"].values
results = []
for i, obs_id in enumerate(obs_ids):
try:
result = Skybot.cone_search(
coords[i],
radius,
observation_times[i],
location=observatory_codes[i]
)
result = result.to_pandas()
result.insert(0, "orbit_id", orbit_id)
result.insert(1, "obs_id", obs_id)
except RuntimeError as e:
result = pd.DataFrame({
"orbit_id" : [orbit_id],
"obs_id" : [obs_id]
})
results.append(result)
results = pd.concat(
results,
ignore_index=True
)
return results
result_dfs = []
for orbit_id in unknown_2021_high_quality["orbit_id"].unique():
print(orbit_id)
obs_ids = recovered_orbit_members[recovered_orbit_members["orbit_id"].isin([orbit_id])]["obs_id"].values
selected_obs = preprocessed_observations[preprocessed_observations["obs_id"].isin(obs_ids)]
results = querySkyBot(selected_obs)
result_dfs.append(results)
results = pd.concat(
result_dfs,
ignore_index=True
)
results
recovered_orbits[recovered_orbits["orbit_id"] == "5f5f205516d5445b9bbae9e80d6ab0ca"]
q = (-47.494 * (1 - 1.101))
print(q)
```
| github_jupyter |
```
import pandas as pd
import sqlalchemy
from sqlalchemy import create_engine
from sqlalchemy import Column, Integer, String, create_engine
from sqlalchemy.ext.declarative import declarative_base
from sqlalchemy.orm import sessionmaker
engine = engine = create_engine('postgresql://live_crimehtx:%7C_IVE_C%7C3IME_hTX001@crimehtxdb.c0fjj9b9p4wl.us-east-1.rds.amazonaws.com/crimehtxdb')
Base = declarative_base(bind=engine)
Session = sessionmaker(bind=engine)
session = Session()
conn = session.bind
# with engine.connect() as con:
# rs = con.execute('SELECT * FROM book')
# for row in rs:
# print row
# test x : -95.322341
# test y : 29.800429
df = pd.read_sql_query('Select * from crime09_15', engine)
df
df = pd.read_sql_query('Select * from crime09_15', engine)
list_snb = df['snbname'].unique().tolist()
list_snb
df.offensetype.value_counts()
list = df.snbname.value_counts()
list
df_murder = df[df['offensetype'] == 'Murder']
df_murder
len(list)
l = pd.DataFrame({"Area":list})
l = l.reset_index()
l = l.rename(columns={"Index": "name", "Area": "area"})
l.to_csv('area.csv',index=None)
df1 = pd.read_sql_query("""select hour,offensetype, snbname from crime09_15 where snbname = 'DOWNTOWN' and offensetype in (Select offensetype from crime09_15 where snbname = 'DOWNTOWN' group by offensetype
order by count(*) desc limit 3) order by hour asc""", engine)
df1
m = df1.groupby(['hour','offensetype']).count()
m#m.sort_values('hour')[]
df1['hour']
```
select
case hour
when '00' THEN 0
when '0' THEN 0
when '0.0' THEN 0
when '''00' THEN 0
when '1' THEN 1
when '01' THEN 1
when '1.0' THEN 1
when '''01' then 1
when '2' THEN 2
when '02' THEN 2
when '2.0' THEN 2
when '''02' then 2
when '3' THEN 3
when '03' THEN 3
when '3.0' THEN 3
when '''03' then 3
when '4' THEN 4
when '04' THEN 4
when '4.0' THEN 4
when '''04' then 4
when '5' THEN 5
when '05' THEN 5
when '5.0' THEN 5
when '''05' then 5
when '6' THEN 6
when '06' THEN 6
when '6.0' THEN 6
when '''06' then 6
when '7' THEN 7
when '07' THEN 7
when '7.0' THEN 7
when '''07' then 7
when '8' THEN 8
when '08' THEN 8
when '8.0' THEN 8
when '''08' then 8
when '9' THEN 9
when '09' THEN 9
when '9.0' THEN 9
when '''09' then 9
when '10' THEN 10
when '010' THEN 10
when '10.0' THEN 10
when '''10' then 10
when '11' THEN 11
when '011' THEN 11
when '11.0' THEN 11
when '''11' then 11
when '12' THEN 12
when '012' THEN 12
when '12.0' THEN 12
when '''12' then 12
when '13' THEN 13
when '013' THEN 13
when '13.0' THEN 13
when '''13' then 13
when '14' THEN 14
when '014' THEN 14
when '14.0' THEN 14
when '''14' then 14
when '15' THEN 15
when '015' THEN 15
when '15.0' THEN 15
when '''15' then 15
when '16' THEN 16
when '016' THEN 16
when '16.0' THEN 16
when '''16' then 16
when '17' THEN 17
when '017' THEN 17
when '17.0' THEN 17
when '''17' then 17
when '18' THEN 18
when '018' THEN 18
when '18.0' THEN 18
when '''18' then 18
when '19' THEN 19
when '019' THEN 19
when '19.0' THEN 19
when '''19' then 19
when '20' THEN 20
when '020' THEN 20
when '20.0' THEN 20
when '''20' then 20
when '21' THEN 21
when '021' THEN 21
when '21.0' THEN 21
when '''21' then 21
when '22' THEN 22
when '022' THEN 22
when '22.0' THEN 22
when '''22' then 22
when '23' THEN 23
when '023' THEN 23
when '23.0' THEN 23
when '''23' then 23
when '24' THEN 24
when '024' THEN 24
when '24.0' THEN 24
when '''24' then 24
else 999
end as hour,
--CAST(hour AS INT) as hour,
offensetype,
snbname from crime09_15 where snbname = 'GREATER GREENSPOINT' and
offensetype in (Select offensetype from crime09_15 where snbname = 'GREATER GREENSPOINT' group by offensetype order by count(*) desc ) order by hour desc
```
incident_df = pd.read_sql_query("""select
case hour
when '00' THEN 0
when '0' THEN 0
when '0.0' THEN 0
when '''00' THEN 0
when '1' THEN 1
when '01' THEN 1
when '1.0' THEN 1
when '''01' then 1
when '2' THEN 2
when '02' THEN 2
when '2.0' THEN 2
when '''02' then 2
when '3' THEN 3
when '03' THEN 3
when '3.0' THEN 3
when '''03' then 3
when '4' THEN 4
when '04' THEN 4
when '4.0' THEN 4
when '''04' then 4
when '5' THEN 5
when '05' THEN 5
when '5.0' THEN 5
when '''05' then 5
when '6' THEN 6
when '06' THEN 6
when '6.0' THEN 6
when '''06' then 6
when '7' THEN 7
when '07' THEN 7
when '7.0' THEN 7
when '''07' then 7
when '8' THEN 8
when '08' THEN 8
when '8.0' THEN 8
when '''08' then 8
when '9' THEN 9
when '09' THEN 9
when '9.0' THEN 9
when '''09' then 9
when '10' THEN 10
when '010' THEN 10
when '10.0' THEN 10
when '''10' then 10
when '11' THEN 11
when '011' THEN 11
when '11.0' THEN 11
when '''11' then 11
when '12' THEN 12
when '012' THEN 12
when '12.0' THEN 12
when '''12' then 12
when '13' THEN 13
when '013' THEN 13
when '13.0' THEN 13
when '''13' then 13
when '14' THEN 14
when '014' THEN 14
when '14.0' THEN 14
when '''14' then 14
when '15' THEN 15
when '015' THEN 15
when '15.0' THEN 15
when '''15' then 15
when '16' THEN 16
when '016' THEN 16
when '16.0' THEN 16
when '''16' then 16
when '17' THEN 17
when '017' THEN 17
when '17.0' THEN 17
when '''17' then 17
when '18' THEN 18
when '018' THEN 18
when '18.0' THEN 18
when '''18' then 18
when '19' THEN 19
when '019' THEN 19
when '19.0' THEN 19
when '''19' then 19
when '20' THEN 20
when '020' THEN 20
when '20.0' THEN 20
when '''20' then 20
when '21' THEN 21
when '021' THEN 21
when '21.0' THEN 21
when '''21' then 21
when '22' THEN 22
when '022' THEN 22
when '22.0' THEN 22
when '''22' then 22
when '23' THEN 23
when '023' THEN 23
when '23.0' THEN 23
when '''23' then 23
when '24' THEN 24
when '024' THEN 24
when '24.0' THEN 24
when '''24' then 24
else 999
end as hour,
--CAST(hour AS INT) as hour,
offensetype,
snbname from crime09_15 where snbname = 'GREATER GREENSPOINT' and
offensetype in (Select offensetype from crime09_15 where snbname = 'GREATER GREENSPOINT' group by offensetype order by count(*) desc ) order by hour desc
""", engine)
def insert_hour(sbn):
s = "\'"+sbn+"\'"
print(s)
query = f"""select
case hour
when '00' THEN 0
when '0' THEN 0
when '0.0' THEN 0
when '''00' THEN 0
when '1' THEN 1
when '01' THEN 1
when '1.0' THEN 1
when '''01' then 1
when '2' THEN 2
when '02' THEN 2
when '2.0' THEN 2
when '''02' then 2
when '3' THEN 3
when '03' THEN 3
when '3.0' THEN 3
when '''03' then 3
when '4' THEN 4
when '04' THEN 4
when '4.0' THEN 4
when '''04' then 4
when '5' THEN 5
when '05' THEN 5
when '5.0' THEN 5
when '''05' then 5
when '6' THEN 6
when '06' THEN 6
when '6.0' THEN 6
when '''06' then 6
when '7' THEN 7
when '07' THEN 7
when '7.0' THEN 7
when '''07' then 7
when '8' THEN 8
when '08' THEN 8
when '8.0' THEN 8
when '''08' then 8
when '9' THEN 9
when '09' THEN 9
when '9.0' THEN 9
when '''09' then 9
when '10' THEN 10
when '010' THEN 10
when '10.0' THEN 10
when '''10' then 10
when '11' THEN 11
when '011' THEN 11
when '11.0' THEN 11
when '''11' then 11
when '12' THEN 12
when '012' THEN 12
when '12.0' THEN 12
when '''12' then 12
when '13' THEN 13
when '013' THEN 13
when '13.0' THEN 13
when '''13' then 13
when '14' THEN 14
when '014' THEN 14
when '14.0' THEN 14
when '''14' then 14
when '15' THEN 15
when '015' THEN 15
when '15.0' THEN 15
when '''15' then 15
when '16' THEN 16
when '016' THEN 16
when '16.0' THEN 16
when '''16' then 16
when '17' THEN 17
when '017' THEN 17
when '17.0' THEN 17
when '''17' then 17
when '18' THEN 18
when '018' THEN 18
when '18.0' THEN 18
when '''18' then 18
when '19' THEN 19
when '019' THEN 19
when '19.0' THEN 19
when '''19' then 19
when '20' THEN 20
when '020' THEN 20
when '20.0' THEN 20
when '''20' then 20
when '21' THEN 21
when '021' THEN 21
when '21.0' THEN 21
when '''21' then 21
when '22' THEN 22
when '022' THEN 22
when '22.0' THEN 22
when '''22' then 22
when '23' THEN 23
when '023' THEN 23
when '23.0' THEN 23
when '''23' then 23
when '24' THEN 24
when '024' THEN 24
when '24.0' THEN 24
when '''24' then 24
else 999
end as hour,
--CAST(hour AS INT) as hour,
offensetype,
snbname from crime09_15 where snbname = {s} and
offensetype in (Select offensetype from crime09_15 where snbname = {s} group by offensetype order by count(*) desc ) order by hour desc
"""
incident_df = pd.read_sql_query(query, engine)
print('length')
print(len(incident_df))
print("query successful")
incident_df.to_sql('incident_hour_count', con=engine, if_exists='append',method = 'multi',chunksize=10000,index=False)
for s in list_snb:
print(s)
insert_hour(s)
print(s)
insert_hour('KASHMERE GARDENS')
list_snb
test_df = pd.read_sql_query("SELECT * FROM incident_hour_count where snbname = 'MEYERLAND AREA'", engine)
len(test_df)
sbn = 'DOWNTOWN'
s = "\'"+sbn+"\'"
query = f"""select
case hour
when '00' THEN 0
when '0' THEN 0
when '0.0' THEN 0
when '''00' THEN 0
when '1' THEN 1
when '01' THEN 1
when '1.0' THEN 1
when '''01' then 1
when '2' THEN 2
when '02' THEN 2
when '2.0' THEN 2
when '''02' then 2
when '3' THEN 3
when '03' THEN 3
when '3.0' THEN 3
when '''03' then 3
when '4' THEN 4
when '04' THEN 4
when '4.0' THEN 4
when '''04' then 4
when '5' THEN 5
when '05' THEN 5
when '5.0' THEN 5
when '''05' then 5
when '6' THEN 6
when '06' THEN 6
when '6.0' THEN 6
when '''06' then 6
when '7' THEN 7
when '07' THEN 7
when '7.0' THEN 7
when '''07' then 7
when '8' THEN 8
when '08' THEN 8
when '8.0' THEN 8
when '''08' then 8
when '9' THEN 9
when '09' THEN 9
when '9.0' THEN 9
when '''09' then 9
when '10' THEN 10
when '010' THEN 10
when '10.0' THEN 10
when '''10' then 10
when '11' THEN 11
when '011' THEN 11
when '11.0' THEN 11
when '''11' then 11
when '12' THEN 12
when '012' THEN 12
when '12.0' THEN 12
when '''12' then 12
when '13' THEN 13
when '013' THEN 13
when '13.0' THEN 13
when '''13' then 13
when '14' THEN 14
when '014' THEN 14
when '14.0' THEN 14
when '''14' then 14
when '15' THEN 15
when '015' THEN 15
when '15.0' THEN 15
when '''15' then 15
when '16' THEN 16
when '016' THEN 16
when '16.0' THEN 16
when '''16' then 16
when '17' THEN 17
when '017' THEN 17
when '17.0' THEN 17
when '''17' then 17
when '18' THEN 18
when '018' THEN 18
when '18.0' THEN 18
when '''18' then 18
when '19' THEN 19
when '019' THEN 19
when '19.0' THEN 19
when '''19' then 19
when '20' THEN 20
when '020' THEN 20
when '20.0' THEN 20
when '''20' then 20
when '21' THEN 21
when '021' THEN 21
when '21.0' THEN 21
when '''21' then 21
when '22' THEN 22
when '022' THEN 22
when '22.0' THEN 22
when '''22' then 22
when '23' THEN 23
when '023' THEN 23
when '23.0' THEN 23
when '''23' then 23
when '24' THEN 24
when '024' THEN 24
when '24.0' THEN 24
when '''24' then 24
else 999
end as hour,
--CAST(hour AS INT) as hour,
offensetype,
snbname from crime09_15 where snbname = {s} and
offensetype in (Select offensetype from crime09_15 where snbname = {s} group by offensetype order by count(*) desc ) order by hour desc
"""
incident_df = pd.read_sql_query(query, engine)
print('length')
print(len(incident_df))
print("query successful")
incident_df.head()
incident_df.to_sql('incident_hour_count', con=engine, if_exists='append',method = 'multi',chunksize=10000,index=False)
test_df = pd.read_sql_query("SELECT * FROM incident_hour_count where snbname = 'DOWNTOWN'", engine)
len(test_df)
# FUnction to convert lat, lng to neighbourhood
agora = {'lat':29.801320, 'lon':-95.322341}
# closest_coord= dfs_09_15.ix[(dfs_09_15[['X','Y']-agora).abs().argsort()[:1]]
from math import cos, asin, sqrt
def distance(lat1, lon1, lat2, lon2):
p = 0.017453292519943295
a = 0.5 - cos((lat2-lat1)*p)/2 + cos(lat1*p)*cos(lat2*p) * (1-cos((lon2-lon1)*p)) / 2
return 12742 * asin(sqrt(a))
def closest(data, agora):
return min(data, key=lambda p: distance(agora['lat'],agora['lon'],p['y'],p['x']))
tempDataList = [{'lat': 29.743340, 'lon': -95.391237, 'SNBNAME': 'NEARTOWN - MONTROSE'},
{'lat': 29.823556, 'lon': -95.502198, 'SNBNAME': 'LANGWOOD' },
{'lat': 29.755974, 'lon': -95.366992, 'SNBNAME': 'DOWNTOWN'}]
#print(closest(tempDataList, agora))
df_t = pd.read_sql_query('Select y,x,snbname from crime09_15', engine)
df_t = pd.read_sql_query('Select y,x,snbname from crime09_15', engine)
dict_incident = df_t.head().to_dict('records')
agora = {'lat':29.801320, 'lon':-95.322341}
print(closest(dict_incident, agora))
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#1.-About-Open-Power-System-Data" data-toc-modified-id="1.-About-Open-Power-System-Data-1">1. About Open Power System Data</a></span></li><li><span><a href="#2.-About-Jupyter-Notebooks-and-GitHub" data-toc-modified-id="2.-About-Jupyter-Notebooks-and-GitHub-2">2. About Jupyter Notebooks and GitHub</a></span></li><li><span><a href="#3.-About-this-datapackage" data-toc-modified-id="3.-About-this-datapackage-3">3. About this datapackage</a></span></li><li><span><a href="#4.-Data-sources" data-toc-modified-id="4.-Data-sources-4">4. Data sources</a></span></li><li><span><a href="#5.-Naming-conventions" data-toc-modified-id="5.-Naming-conventions-5">5. Naming conventions</a></span></li><li><span><a href="#6.-License" data-toc-modified-id="6.-License-6">6. License</a></span></li></ul></div>
<div style="width:100%; background-color: #D9EDF7; border: 1px solid #CFCFCF; text-align: left; padding: 10px;">
<b>Time series: Processing Notebook</b>
<ul>
<li>Main Notebook</li>
<li><a href="processing.ipynb">Processing Notebook</a></li>
</ul>
<br>This Notebook is part of the <a href="http://data.open-power-system-data.org/time_series">Time series Data Package</a> of <a href="http://open-power-system-data.org">Open Power System Data</a>.
</div>
# 1. About Open Power System Data
This notebook is part of the project [Open Power System Data](http://open-power-system-data.org). Open Power System Data develops a platform for free and open data for electricity system modeling. We collect, check, process, document, and provide data that are publicly available but currently inconvenient to use.
More info on Open Power System Data:
- [Information on the project on our website](http://open-power-system-data.org)
- [Data and metadata on our data platform](http://data.open-power-system-data.org)
- [Data processing scripts on our GitHub page](https://github.com/Open-Power-System-Data)
# 2. About Jupyter Notebooks and GitHub
This file is a [Jupyter Notebook](http://jupyter.org/). A Jupyter Notebook is a file that combines executable programming code with visualizations and comments in markdown format, allowing for an intuitive documentation of the code. We use Jupyter Notebooks for combined coding and documentation. We use Python 3 as programming language. All Notebooks are stored on [GitHub](https://github.com/), a platform for software development, and are publicly available. More information on our IT-concept can be found [here](http://open-power-system-data.org/it). See also our [step-by-step manual](http://open-power-system-data.org/step-by-step) how to use the dataplatform.
# 3. About this datapackage
We provide data in different chunks, or [data packages](http://frictionlessdata.io/data-packages/). The one you are looking at right now, [Time series](http://data.open-power-system-data.org/time_series/), contains various kinds of time series data in 15min, 30min or 60min resolution, namely:
- electricity consumption (load)
- wind and solar power: capacity, generation forecast, actual generation
- day-ahead spot prices
The main focus of this datapackage is German data, but we include data from other countries wherever possible.
The timeseries become available at different points in time depending on the sources. The full dataset is only available from 2015 onwards.
The data has been downloaded from the sources, resampled and merged in a large CSV file with hourly resolution. Additionally, the data available at a higher resolution (some renewables in-feed, 15 minutes) is provided in a separate file.
# 4. Data sources
The main data sources are the various European Transmission System Operators (TSOs) and the [ENTSO-E Data Portal](https://www.entsoe.eu/data/data-portal/Pages/default.aspx). Where no data is available from hte TSOs directly, data are taken from the [ENTSO-E Transparency Plstform](https://transparency.entsoe.eu). A complete list of data sources is provided on the [datapackage information website](http://data.open-power-system-data.org/time_series/). They are also contained in the JSON file that contains all metadata.
# 5. Naming conventions
```
import pandas as pd; pd.read_csv('input/notation.csv', index_col=list(range(4)))
```
# 6. License
This notebook as well as all other documents in this repository is published under the [MIT License](LICENSE.md).
| github_jupyter |
# The Alien Blaster problem
This notebook presents solutions to exercises in Think Bayes.
Copyright 2016 Allen B. Downey
MIT License: https://opensource.org/licenses/MIT
```
from __future__ import print_function, division
% matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import numpy as np
from thinkbayes2 import Hist, Pmf, Cdf, Suite, Beta
import thinkplot
```
## Part One
In preparation for an alien invasion, the Earth Defense League has been working on new missiles to shoot down space invaders. Of course, some missile designs are better than others; let's assume that each design has some probability of hitting an alien ship, $x$.
Based on previous tests, the distribution of $x$ in the population of designs is well-modeled by a beta distribution with parameters $\alpha=2$ and $\beta=3$. What is the average missile's probability of shooting down an alien?
```
prior = Beta(2, 3)
thinkplot.Pdf(prior.MakePmf())
prior.Mean()
```
In its first test, the new Alien Blaster 9000 takes 10 shots and hits 2 targets. Taking into account this data, what is the posterior distribution of $x$ for this missile? What is the value in the posterior with the highest probability, also known as the MAP?
```
posterior = Beta(3, 2)
posterior.Update((2, 8))
posterior.MAP()
```
Now suppose the new ultra-secret Alien Blaster 10K is being tested. In a press conference, an EDF general reports that the new design has been tested twice, taking two shots during each test. The results of the test are confidential, so the general won't say how many targets were hit, but they report: "The same number of targets were hit in the two tests, so we have reason to think this new design is consistent."
Write a class called `AlienBlaster` that inherits from `Suite` and provides a likelihood function that takes this data -- two shots and a tie -- and computes the likelihood of the data for each hypothetical value of $x$. If you would like a challenge, write a version that works for any number of shots.
```
from scipy import stats
class AlienBlaster(Suite):
def Likelihood(self, data, hypo):
"""Computes the likeliood of data under hypo.
data: number of shots they took
hypo: probability of a hit, p
"""
n = data
x = hypo
# specific version for n=2 shots
likes = [x**4, (1-x)**4, (2*x*(1-x))**2]
# general version for any n shots
likes = [stats.binom.pmf(k, n, x)**2 for k in range(n+1)]
return np.sum(likes)
```
If we start with a uniform prior, we can see what the likelihood function looks like:
```
pmf = Beta(1, 1).MakePmf()
blaster = AlienBlaster(pmf)
blaster.Update(2)
thinkplot.Pdf(blaster)
```
A tie is most likely if they are both terrible shots or both very good.
Is this data good or bad; that is, does it increase or decrease your estimate of $x$ for the Alien Blaster 10K?
Now let's run it with the specified prior and see what happens when we multiply the convex prior and the concave posterior:
```
pmf = Beta(2, 3).MakePmf()
blaster = AlienBlaster(pmf)
blaster.Update(2)
thinkplot.Pdf(blaster)
```
The posterior mean and MAP are lower than in the prior.
```
prior.Mean(), blaster.Mean()
prior.MAP(), blaster.MAP()
```
So if we learn that the new design is "consistent", it is more likely to be consistently bad (in this case).
## Part Two
Suppose we
have we have a stockpile of 3 Alien Blaster 9000s and 7 Alien
Blaster 10Ks. After extensive testing, we have concluded that
the AB9000 hits the target 30% of the time, precisely, and the
AB10K hits the target 40% of the time.
If I grab a random weapon from the stockpile and shoot at 10 targets,
what is the probability of hitting exactly 3? Again, you can write a
number, mathematical expression, or Python code.
```
k = 3
n = 10
x1 = 0.3
x2 = 0.4
0.3 * stats.binom.pmf(k, n, x1) + 0.7 * stats.binom.pmf(k, n, x2)
```
The answer is a value drawn from the mixture of the two distributions.
Continuing the previous problem, let's estimate the distribution
of `k`, the number of successful shots out of 10.
1. Write a few lines of Python code to simulate choosing a random weapon and firing it.
2. Write a loop that simulates the scenario and generates random values of `k` 1000 times.
3. Store the values of `k` you generate and plot their distribution.
```
def flip(p):
return np.random.random() < p
def simulate_shots(n, p):
return np.random.binomial(n, p)
ks = []
for i in range(1000):
if flip(0.3):
k = simulate_shots(n, x1)
else:
k = simulate_shots(n, x2)
ks.append(k)
```
Here's what the distribution looks like.
```
pmf = Pmf(ks)
thinkplot.Hist(pmf)
len(ks), np.mean(ks)
```
The mean should be near 3.7. We can run this simulation more efficiently using NumPy. First we generate a sample of `xs`:
```
xs = np.random.choice(a=[x1, x2], p=[0.3, 0.7], size=1000)
Hist(xs)
```
Then for each `x` we generate a `k`:
```
ks = np.random.binomial(n, xs)
```
And the results look similar.
```
pmf = Pmf(ks)
thinkplot.Hist(pmf)
np.mean(ks)
```
One more way to do the same thing is to make a meta-Pmf, which contains the two binomial `Pmf` objects:
```
from thinkbayes2 import MakeBinomialPmf
pmf1 = MakeBinomialPmf(n, x1)
pmf2 = MakeBinomialPmf(n, x2)
metapmf = Pmf({pmf1:0.3, pmf2:0.7})
metapmf.Print()
```
Here's how we can draw samples from the meta-Pmf:
```
ks = [metapmf.Random().Random() for _ in range(1000)]
```
And here are the results, one more time:
```
pmf = Pmf(ks)
thinkplot.Hist(pmf)
np.mean(ks)
```
This result, which we have estimated three ways, is a predictive distribution, based on our uncertainty about `x`.
We can compute the mixture analtically using `thinkbayes2.MakeMixture`:
def MakeMixture(metapmf, label='mix'):
"""Make a mixture distribution.
Args:
metapmf: Pmf that maps from Pmfs to probs.
label: string label for the new Pmf.
Returns: Pmf object.
"""
mix = Pmf(label=label)
for pmf, p1 in metapmf.Items():
for k, p2 in pmf.Items():
mix[k] += p1 * p2
return mix
The outer loop iterates through the Pmfs; the inner loop iterates through the items.
So `p1` is the probability of choosing a particular Pmf; `p2` is the probability of choosing a value from the Pmf.
In the example, each Pmf is associated with a value of `x` (probability of hitting a target). The inner loop enumerates the values of `k` (number of targets hit after 10 shots).
```
from thinkbayes2 import MakeMixture
mix = MakeMixture(metapmf)
thinkplot.Hist(mix)
mix.Mean()
```
**Exercise**: Assuming again that the distribution of `x` in the population of designs is well-modeled by a beta distribution with parameters α=2 and β=3, what the distribution if `k` if I choose a random Alien Blaster and fire 10 shots?
| github_jupyter |
# Always run these first three cells.
```
reset
# IMPORT PACKAGES
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker as mticker
from netCDF4 import Dataset
import cartopy.crs as ccrs
import cartopy.feature as feature
import cmocean.cm
import pandas as pd
import xarray as xr
import collections
from scipy.stats import ttest_ind, ttest_rel
# fix to cartopy issue right now
from matplotlib.axes import Axes
from cartopy.mpl.geoaxes import GeoAxes
GeoAxes._pcolormesh_patched = Axes.pcolormesh
# ADDITIONAL FUNCTIONS
# CODE TO CALCULATE SEASONAL AVERAGE
# Taken from: https://xarray.pydata.org/en/v0.14.0/examples/monthly-means.html
# However test plot in "Seasonal Mean Example" produces somewhat different results-- not sure why.
# usage example: ds_weighted = season_mean(ds,calendar='noleap')
dpm = {'noleap': [0, 31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31],
'365_day': [0, 31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31],
'standard': [0, 31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31],
'gregorian': [0, 31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31],
'proleptic_gregorian': [0, 31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31],
'all_leap': [0, 31, 29, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31],
'366_day': [0, 31, 29, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31],
'360_day': [0, 30, 30, 30, 30, 30, 30, 30, 30, 30, 30, 30, 30],
'julian': [0, 31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31]}
def leap_year(year, calendar='standard'):
"""Determine if year is a leap year"""
leap = False
if ((calendar in ['standard', 'gregorian',
'proleptic_gregorian', 'julian']) and
(year % 4 == 0)):
leap = True
if ((calendar == 'proleptic_gregorian') and
(year % 100 == 0) and
(year % 400 != 0)):
leap = False
elif ((calendar in ['standard', 'gregorian']) and
(year % 100 == 0) and (year % 400 != 0) and
(year < 1583)):
leap = False
return leap
def get_dpm(time, calendar='standard'):
"""
return a array of days per month corresponding to the months provided in `months`
"""
month_length = np.zeros(len(time), dtype=np.int)
cal_days = dpm[calendar]
for i, (month, year) in enumerate(zip(time.month, time.year)):
month_length[i] = cal_days[month]
if leap_year(year, calendar=calendar):
month_length[i] += 1
return month_length
def season_mean(ds, calendar='standard'):
# Make a DataArray with the number of days in each month, size = len(time)
month_length = xr.DataArray(get_dpm(ds.time.to_index(), calendar=calendar),
coords=[ds.time], name='month_length')
# Calculate the weights by grouping by 'time.season'
weights = month_length.groupby('time.season') / month_length.groupby('time.season').sum()
# Test that the sum of the weights for each season is 1.0
np.testing.assert_allclose(weights.groupby('time.season').sum().values, np.ones(4))
# Calculate the weighted average
return (ds * weights).groupby('time.season').sum(dim='time')
def annual_season_mean(ds, calendar='standard'): # Added 7-9-20
# TRUNCATE TIME SERIES IN ADVANCE SO THERE ARE ONLY COMPLETE YEARS, OTHERWISE GET INF/0 AT END FOR SOME SEASONS
ml = xr.DataArray(get_dpm(ds.time.to_index(), calendar=calendar),
coords=[ds.time], name='month_length') # ml stands for month_length
ml_seas = {}
ml_seas_sums = {}
weights = {}
ds_seas = {}
ann_seas_mean_v1 = {}
for seas in ['DJF','MAM','JJA','SON']:
ml_seas[seas] = ml.where(ds['time.season'] == seas)
ml_seas_sums[seas] = ml_seas[seas].rolling(min_periods=3, center=True, time=3).sum()
if seas == 'DJF':
ml_seas[seas] = ml_seas[seas].shift(time=1) # done to ensure weights for consecutive D-J-F add up to 1 despite crossing year
weights[seas] = ml_seas[seas].groupby('time.year') / ml_seas_sums[seas].groupby('time.year').sum()
ds_seas[seas] = ds.where(ds['time.season'] == seas)
if seas == 'DJF':
ds_seas[seas] = ds_seas[seas].shift(time=1)
ann_seas_mean_v1[seas] = (ds_seas[seas] * weights[seas]).groupby('time.year').sum(dim='time') # 0 AT END IF NOT COMPLETE YEAR
ann_seas_mean_v1['DJF'] = ann_seas_mean_v1['DJF'].isel(year=slice(1,len(ann_seas_mean_v1['DJF'].year)))
return ann_seas_mean_v1
# CODE TO SHIFT CCSM DATA BY 1 MONTH (given weird NCAR conventions that calendar date is end of time bound for each month)
def shift_dates_1M(dat):
y_first = str(dat.time.values[0].year).zfill(4)
y_last = str(dat.time.values[-2].year).zfill(4)
time2 = xr.cftime_range(start=str(y_first)+'-01-01', end=str(y_last)+'-12-01', freq="1MS", calendar="noleap")
dat = dat.assign_coords(time=time2)
return dat
```
# Can skip these cells if ran already.
```
# PATHS TO DATA FILES
# direc = '/tigress/GEOCLIM/janewb/MODEL_OUT'
# files = {}
# files['ctrl'] = direc+'/CTL1860_newdiag_tigercpu_intelmpi_18_576PE_coldstart/TIMESERIES/precip.00010101-03000101.atmos_month.nc'
# files['cam'] = '/tigress/janewb/MODEL_OUT_HERE/CTL1860_newdiag_tigercpu_intelmpi_18_576PE_coldstart_HiTopo1_CAM/TIMESERIES/precip.00010101-02050101.atmos_month.nc'
# files['hitopo'] = direc+'/CTL1860_newdiag_tigercpu_intelmpi_18_576PE_coldstart_HiTopo1/TIMESERIES/precip.00010101-06000101.atmos_month.nc'
# files['cesm_ctrl'] = '/tigress/janewb/HiTopo/FROM_ALYSSA/CESM_data/b40.1850.track1.1deg.006.cam2.h0.PRECT.120001-130012.nc'
# files['cesm_cam'] = '/tigress/janewb/HiTopo/FROM_ALYSSA/CESM_data/ccsm4pi_topo2.cam2.h0.PRECT.000101-029912.nc'
# files['gpcp'] = '/tigress/wenchang/data/gpcp/v2p3/precip.mon.mean.197901_201808.nc'
# files['cmap'] = '/tigress/janewb/OBS/CMAP/precip.mon.mean.nc'
# files['imerg'] = '/tigress/janewb/OBS/GPM/precip.mon.mean.nc'
# land_files = {}
# land_files['ctrl'] = direc+'/CTL1860_newdiag_tigercpu_intelmpi_18_576PE_coldstart/TIMESERIES/land_mask.nc'
# land_files['imerg'] = '/tigress/janewb/OBS/GPM/IMERG_land_sea_mask.nc'
# land_files['cesm_ctrl'] = '/tigress/janewb/HiTopo/FROM_ALYSSA/CESM_topo_mods/hose1Sv_ccsm4_pi01.cam2.h0.0001-01.nc'
# land_files['cesm_cam'] = '/tigress/janewb/HiTopo/FROM_ALYSSA/CESM_topo_mods/shf_heating_topo6.cam2.h0.0001-01.nc'
direc = '/tigress/janewb/public_html/HITOPO/'
files = {}
files['ctrl'] = direc+'flor.ctrl.precip.nc'
files['cam'] = direc+'flor.cam.precip.nc'
files['hitopo'] = direc+'flor.hitopo.precip.nc'
files['cesm_ctrl'] = direc+'ccsm4.ctrl.precip.nc'
files['cesm_cam'] = direc+'ccsm4.ideal_cam.precip.nc'
files['gpcp'] = direc+'obs.gpcp.precip.nc'
files['cmap'] = direc+'obs.cmap.precip.nc'
files['imerg'] = direc+'obs.imerg.precip*.nc'
land_files = {}
land_files['ctrl'] = direc+'flor.land_mask.nc'
land_files['imerg'] = direc+'obs.imerg.land.nc'
land_files['cesm_ctrl'] = direc+'ccsm4.ctrl.land.nc'
land_files['cesm_cam'] = direc+'ccsm4.ideal_cam.land.nc'
%%time
# DATA CLEANING
dat = {}
pr = {}
tsel = {}
x = 'lon'
y = 'lat'
precip_var = 'precip'
units = 'mm/day'
model_tmin = '0031'
model_tmax = '0200'
calendars = {'ctrl':'noleap',
'cam':'noleap',
'hitopo':'noleap',
'cmap':'gregorian',
'gpcp':'gregorian',
'imerg':'julian'
}
# FLOR Runs
for key in ['ctrl','cam','hitopo']:
dat[key] = xr.open_dataset(files[key])
pr[key] = dat[key].precip*86400
pr[key].attrs = dat[key].attrs
pr[key].attrs['units'] = units
pr[key] = pr[key].rename({'grid_xt': 'lon','grid_yt': 'lat'})
tsel[key] = pr[key].sel(time = slice(model_tmin,model_tmax))
# CESM RUNS
for key in ['cesm_ctrl','cesm_cam']:
dat[key] = xr.open_dataset(files[key])
dat[key] = shift_dates_1M(dat[key])
dat[key] = dat[key].rename({'PRECT': 'precip'})
pr[key] = dat[key].precip*86400*1000
pr[key].attrs = dat[key].attrs
pr[key].attrs['units'] = units
tsel[key] = pr[key]
# OBSERVATIONS
# # CMAP
dat['cmap'] = xr.open_dataset(files['cmap'])
pr['cmap'] = dat['cmap'].precip
pr['cmap'].attrs = dat['cmap'].attrs
tsel['cmap'] = pr['cmap'].sel(time = slice('1979','2018'))
# # GPCP
dat['gpcp'] = xr.open_dataset(files['gpcp'])
pr['gpcp'] = dat['gpcp'].precip
pr['gpcp'].attrs = dat['gpcp'].attrs
tsel['gpcp'] = pr['gpcp'].sel(time = slice('1979','2017'))
# IMERG
dat['imerg'] = xr.open_mfdataset(files['imerg'])
dat['imerg'] = dat['imerg'].transpose('time','lat','lon')
dat['imerg'] = dat['imerg'].assign_coords(lon=(dat['imerg'].lon % 360)).roll(lon=1800, roll_coords=True)
dat['imerg'] = dat['imerg'].rename({'precipitation':'precip'})
pr['imerg'] = dat['imerg'].precip*24
pr['imerg'].attrs = dat['imerg'].attrs
pr['imerg'].attrs['units'] = units
tsel['imerg'] = pr['imerg'].sel(time = slice('2001','2018'))
land_dat = {}
land = {}
land_units = 'fractional amount of ocean'
# LAND MASKS
land_dat['imerg'] = xr.open_dataset(land_files['imerg'])
land_dat['imerg'] = land_dat['imerg'].rename({'landseamask':'land_mask'})
land['imerg'] = land_dat['imerg'].land_mask/100
land['imerg'].attrs['units'] = land_units
land['cmap'] = land['imerg'].interp_like(tsel['cmap'],method='linear')
land['gpcp'] = land['imerg'].interp_like(tsel['gpcp'],method='linear')
land['imerg'] = land['imerg'].sel(lon=slice(0.05,360.0)) # land mask has two extra points compared to precip. data
land_dat['ctrl'] = xr.open_dataset(land_files['ctrl'])
land_dat['ctrl'] = land_dat['ctrl'].rename({'LAND_MASK':'land_mask'})
land['ctrl'] = 1-land_dat['ctrl'].land_mask
land['ctrl'].attrs['units'] = land_units
land['hitopo'] = land['ctrl']
land['cam'] = land['ctrl']
land_dat['cesm_ctrl'] = xr.open_dataset(land_files['cesm_ctrl'])
land_dat['cesm_ctrl'] = land_dat['cesm_ctrl'].rename({'LANDFRAC':'land_mask'})
land['cesm_ctrl'] = 1-land_dat['cesm_ctrl'].land_mask
land['cesm_ctrl'].attrs['units'] = land_units
land['cesm_cam'] = land['cesm_ctrl']
# PRECIP SAVE DATA TO NETCDF SO DON'T HAVE TO RECALCULATE EACH TIME
tsel['ctrl'].to_netcdf('PRECIP/precip_ctrl.nc')
tsel['hitopo'].to_netcdf('PRECIP/precip_hitopo.nc')
tsel['cam'].to_netcdf('PRECIP/precip_cam.nc')
tsel['imerg'].to_netcdf('PRECIP/precip_imerg.nc')
tsel['cmap'].to_netcdf('PRECIP/precip_cmap.nc')
tsel['gpcp'].to_netcdf('PRECIP/precip_gpcp.nc')
tsel['cesm_ctrl'].to_netcdf('PRECIP/precip_cesm_ctrl.nc')
tsel['cesm_cam'].to_netcdf('PRECIP/precip_cesm_cam.nc')
# LAND SAVE DATA
land['ctrl'].to_netcdf('PRECIP/land_model.nc')
land['imerg'].to_netcdf('PRECIP/land_imerg.nc')
land['cmap'].to_netcdf('PRECIP/land_cmap.nc')
land['gpcp'].to_netcdf('PRECIP/land_gpcp.nc')
land['cesm_ctrl'].to_netcdf('PRECIP/land_cesm_ctrl.nc')
land['cesm_cam'].to_netcdf('PRECIP/land_cesm_cam.nc')
```
# Start from here if saved out analyzed data.
```
# OPEN SAVED OUT DATA
land = {}
land['ctrl'] = xr.open_dataset('PRECIP/land_model.nc').land_mask
land['imerg'] = xr.open_dataset('PRECIP/land_imerg.nc').land_mask
land['cmap'] = xr.open_dataset('PRECIP/land_cmap.nc').land_mask
land['gpcp'] = xr.open_dataset('PRECIP/land_gpcp.nc').land_mask
land['cesm_ctrl'] = xr.open_dataset('PRECIP/land_cesm_ctrl.nc').land_mask
land['cesm_cam'] = xr.open_dataset('PRECIP/land_cesm_cam.nc').land_mask
tsel = {}
tsel['ctrl'] = xr.open_dataset('PRECIP/precip_ctrl.nc').precip
tsel['hitopo'] = xr.open_dataset('PRECIP/precip_hitopo.nc').precip
tsel['cam'] = xr.open_dataset('PRECIP/precip_cam.nc').precip
tsel['imerg'] = xr.open_dataset('PRECIP/precip_imerg.nc').precip
tsel['cmap'] = xr.open_dataset('PRECIP/precip_cmap.nc').precip
tsel['gpcp'] = xr.open_dataset('PRECIP/precip_gpcp.nc').precip
tsel['cesm_ctrl'] = xr.open_dataset('PRECIP/precip_cesm_ctrl.nc').precip
tsel['cesm_cam'] = xr.open_dataset('PRECIP/precip_cesm_cam.nc').precip
# REGIONS FOR AVERAGING
# Pacific Zonal Limits
xmin = 230
xmax = 285
# Tropical meridional limits
tymin = -23.5
tymax = 23.5
# CREATE OCEAN MASK
mask = {}
mask['ctrl'] = np.array(land['ctrl'].where(land['ctrl'] == 1))
mask['cam'] = mask['ctrl']
mask['hitopo'] = mask['ctrl']
mask['imerg'] = np.array(land['imerg'].where(land['imerg'] == 1))
mask['gpcp'] = np.array(land['gpcp'].where(land['gpcp'] == 1))
mask['cmap'] = np.array(land['cmap'].where(land['cmap'] == 1))
mask['cesm_ctrl'] = np.array(land['cesm_ctrl'].where(land['cesm_ctrl'] == 1))
mask['cesm_cam'] = mask['cesm_ctrl']
# MASK DATA
tsel_seamask = {}
for key in mask.keys():
tsel_seamask[key] = tsel[key]*mask[key]
# DATA ANALYSIS
calendars = {'ctrl':'noleap',
'cam':'noleap',
'hitopo':'noleap',
'cmap':'gregorian',
'gpcp':'gregorian',
'imerg':'julian',
'cesm_ctrl':'noleap',
'cesm_cam':'noleap'
}
tmean = {}
seasmean = {}
ann_seasmean = {}
xtmean = {}
pac_xtmean = {}
seas_xtmean = {}
seas_pac_xtmean = {}
tmean_seamask = {}
seasmean_seamask = {}
pac_xtmean_seamask = {}
seas_pac_xtmean_seamask = {}
for key in tsel_seamask.keys():
tmean[key] = tsel[key].mean(dim='time')
seasmean[key] = season_mean(tsel[key],calendar=calendars[key])
ann_seasmean[key] = annual_season_mean(tsel[key],calendar=calendars[key])
xtmean[key] = tmean[key].mean(dim='lon')
seas_xtmean[key] = seasmean[key].mean(dim='lon')
pac_xtmean[key] = tmean[key].sel(lon = slice(xmin,xmax)).mean(dim='lon')
seas_pac_xtmean[key] = seasmean[key].sel(lon = slice(xmin,xmax)).mean(dim='lon')
tmean_seamask[key] = tsel_seamask[key].mean(dim='time')
seasmean_seamask[key] = season_mean(tsel_seamask[key],calendar=calendars[key])
pac_xtmean_seamask[key] = tmean_seamask[key].sel(lon = slice(xmin,xmax)).mean(dim='lon')
seas_pac_xtmean_seamask[key] = seasmean_seamask[key].sel(lon = slice(xmin,xmax)).mean(dim='lon')
#STATISTICAL SIGNIFICANCE TEST
#test with identical sample sizes
def sigtest(yearmean1,yearmean2,timemean1,timemean2):
ptvals = ttest_rel(yearmean1,yearmean2, axis=0)
diff = timemean1-timemean2
diff_mask = np.ma.masked_where(ptvals[1] > 0.1,diff)
return diff,diff_mask,ptvals
def sigtest2n(yearmean1,yearmean2,timemean1,timemean2):
ptvals = ttest_ind(yearmean1,yearmean2, axis=0, equal_var = False)
diff = timemean1-timemean2
diff_mask = np.ma.masked_where(ptvals[1] > 0.1,diff)
return diff,diff_mask,ptvals
# Calculate difference and statistical significance mask
diff_mask = collections.defaultdict(dict)
diff_mask_cesm = collections.defaultdict(dict)
diff = collections.defaultdict(dict)
diff_cesm = collections.defaultdict(dict)
ptvals = collections.defaultdict(dict)
ptvals_cesm = collections.defaultdict(dict)
seasons = ['DJF','MAM','JJA','SON']
# COMPARING CONTROL AND TOPO. MODEL RUNS
for key in ['hitopo','cam']:
for seas in seasons:
diff[key][seas],diff_mask[key][seas],ptvals[key][seas] = sigtest(ann_seasmean[key][seas],ann_seasmean['ctrl'][seas],seasmean[key].sel(season=seas),seasmean['ctrl'].sel(season=seas))
for seas in seasons:
diff['cesm_cam'][seas],diff_mask['cesm_cam'][seas],ptvals['cesm_cam'][seas] = sigtest2n(ann_seasmean['cesm_cam'][seas].values,ann_seasmean['cesm_ctrl'][seas].values,seasmean['cesm_cam'].sel(season=seas).values,seasmean['cesm_ctrl'].sel(season=seas).values)
# COMPARING CONTROL MODEL RUNS WITH OBS.
seasmean_regrid = {}
ann_seasmean_regrid = collections.defaultdict(dict)
for key in ['imerg','cmap','gpcp']:
seasmean_regrid[key] = seasmean[key].interp_like(tmean['ctrl'],method='linear')
ann_seasmean_regrid[key][seas] = ann_seasmean[key][seas].interp_like(tmean['ctrl'],method='linear')
for seas in seasons:
ann_seasmean_regrid[key][seas] = ann_seasmean[key][seas].interp_like(tmean['ctrl'],method='linear')
diff[key][seas],diff_mask[key][seas],ptvals[key][seas] = sigtest2n(ann_seasmean_regrid[key][seas].values,ann_seasmean['ctrl'][seas].values,seasmean_regrid[key].sel(season=seas).values,seasmean['ctrl'].sel(season=seas).values)
# Note: mask may not be working for DJF
for key in ['imerg','cmap','gpcp']:
seasmean_regrid[key] = seasmean[key].interp_like(tmean['cesm_ctrl'],method='linear')
ann_seasmean_regrid[key][seas] = ann_seasmean[key][seas].interp_like(tmean['ctrl'],method='linear')
for seas in seasons:
ann_seasmean_regrid[key][seas] = ann_seasmean[key][seas].interp_like(tmean['cesm_ctrl'],method='linear')
diff_cesm[key][seas],diff_mask_cesm[key][seas],ptvals_cesm[key][seas] = sigtest2n(ann_seasmean_regrid[key][seas].values,ann_seasmean['cesm_ctrl'][seas].values,seasmean_regrid[key].sel(season=seas).values,seasmean['cesm_ctrl'].sel(season=seas).values)
# Calculate annual tmean for statistical significance testing
ann_tmean = {}
for key in tsel_seamask.keys():
ann_tmean[key] = tsel[key].groupby('time.year').mean(dim='time')
for key in ['hitopo','cam']:
diff[key]['ANN'],diff_mask[key]['ANN'],ptvals[key]['ANN'] = sigtest(ann_tmean[key],ann_tmean['ctrl'],tmean[key],tmean['ctrl'])
# REGRID AND CALCULATE DIFFERENCE FROM OBS
regrid = collections.defaultdict(dict)
seasmean_regrid = collections.defaultdict(dict)
diff = collections.defaultdict(dict)
seas_diff = collections.defaultdict(dict)
pdiff = collections.defaultdict(dict)
seas_pdiff = collections.defaultdict(dict)
for key in ['imerg','cmap','gpcp']:
regrid[key] = tmean[key].interp_like(tmean['ctrl'],method='linear')
seasmean_regrid[key] = seasmean[key].interp_like(tmean['ctrl'],method='linear')
for run in ['ctrl','hitopo','cam']:
diff[key][run] = regrid[key] - tmean[run]
seas_diff[key][run] = seasmean_regrid[key] - seasmean[run]
pdiff[key][run] = (regrid[key] - tmean[run])/regrid[key]
seas_pdiff[key][run] = (seasmean_regrid[key] - seasmean[run])/seasmean_regrid[key]
for key in ['imerg','cmap','gpcp']:
regrid[key] = tmean[key].interp_like(tmean['cesm_ctrl'],method='linear')
seasmean_regrid[key] = seasmean[key].interp_like(tmean['cesm_ctrl'],method='linear')
for run in ['cesm_ctrl','cesm_cam']:
diff[key][run] = regrid[key] - tmean[run]
seas_diff[key][run] = seasmean_regrid[key] - seasmean[run]
pdiff[key][run] = (regrid[key] - tmean[run])/regrid[key]
seas_pdiff[key][run] = (seasmean_regrid[key] - seasmean[run])/seasmean_regrid[key]
# CALCULATE DIFFERENCE BETWEEN DIFFERENT MODEL RUNS
mdiff = {}
seas_mdiff = {}
for key in ['hitopo','cam']:
mdiff[key] = tmean['ctrl'] - tmean[key]
seas_mdiff[key] = seasmean['ctrl'] - seasmean[key]
# 6-5-20: Having problem with grids-- cutting off 50% of latitudes for CESM data when subtracting dataarrays, so directly subtracting arrays
mdiff0 = {}
seas_mdiff0 = {}
lat = tmean['cesm_ctrl']['lat'].lat.values
lon = tmean['cesm_ctrl']['lon'].lon.values
season = seasmean['cesm_ctrl'].season.values
for key in ['cesm_cam']:
mdiff0[key] = tmean['cesm_ctrl'].values - tmean[key].values
mdiff[key] = xr.DataArray(mdiff0[key], coords={'lat': lat, 'lon': lon}, dims=['lat', 'lon'])
seas_mdiff0[key] = seasmean['cesm_ctrl'].values - seasmean[key].values
seas_mdiff[key] = xr.DataArray(seas_mdiff0[key], coords={'season': season, 'lat': lat, 'lon': lon}, dims=['season','lat', 'lon'])
# Lon and lats
lonf = seasmean['ctrl'].lon
latf = seasmean['ctrl'].lat
lonc = seasmean['cesm_ctrl'].lon
latc = seasmean['cesm_ctrl'].lat
# COMBINED FLOR CCSM4 ZONAL MEAN PRECIPITATION PLOT
fig = plt.figure(figsize=(12,12))
plt.rcParams.update({'font.size': 14})
fs_legend = 12
fs_label = 18
xlim1 = -23.5
xlim2 = 23.5
ylim1 = 0
ylim2 = 13.5
ax1 = plt.subplot(321)
pac_xtmean_seamask['imerg'].plot(color='black', label = 'Obs. 1: IMERG')
pac_xtmean['cmap'].plot(color='black',dashes = [4,4,4,4], label='Obs. 2: CMAP')
pac_xtmean['gpcp'].plot(color='black',dashes=[2,1,2,1], label='Obs. 3: GPCP')
pac_xtmean_seamask['ctrl'].plot(color='blue', label = 'FLOR Control')
pac_xtmean_seamask['hitopo'].plot(color='red', label = 'FLOR HiTopo')
pac_xtmean_seamask['cam'].plot(color='red',dashes=[1,1,1,1], label = 'FLOR CAm')
ax1.set_title('a)')
ax1.set_ylabel('Zonal Mean Precipitation [mm/day]')
ax1.set_xlabel('')
ax1.set_xlim([xlim1,xlim2])
ax1.set_ylim([ylim1,ylim2])
ax1.legend(fontsize=fs_legend)
ax2 = plt.subplot(322)
pac_xtmean_seamask['imerg'].plot(color='black', label = 'Obs. 1: IMERG')
pac_xtmean['cmap'].plot(color='black',dashes = [4,4,4,4], label='Obs. 2: CMAP')
pac_xtmean['gpcp'].plot(color='black',dashes=[2,1,2,1], label='Obs. 3: GPCP')
pac_xtmean_seamask['cesm_ctrl'].plot(color='blue', linewidth = 1, label = 'CCSM4 Control')
pac_xtmean_seamask['cesm_cam'].plot(color='red',dashes=[1,1,1,1], label = 'CCSM4 Ideal CAm')
ax2.set_title('b)')
ax2.set_xlabel('')
ax2.set_ylabel('')
ax2.set_xlim([xlim1,xlim2])
ax2.set_ylim([ylim1,ylim2])
ax2.legend(fontsize=fs_legend)
ax3 = plt.subplot(323)
seas_pac_xtmean_seamask['imerg'].sel(season='MAM').plot(color='black', label = 'Obs. 1: IMERG')
seas_pac_xtmean['cmap'].sel(season='MAM').plot(color='black',dashes = [4,4,4,4], label='Obs. 2: CMAP')
seas_pac_xtmean['gpcp'].sel(season='MAM').plot(color='black',dashes=[2,1,2,1], label='Obs. 3: GPCP')
seas_pac_xtmean_seamask['ctrl'].sel(season='MAM').plot(color='blue', label = 'FLOR Control')
seas_pac_xtmean_seamask['hitopo'].sel(season='MAM').plot(color='red', label = 'FLOR HiTopo')
seas_pac_xtmean_seamask['cam'].sel(season='MAM').plot(color='red',dashes=[1,1,1,1], label = 'FLOR CAm')
ax3.set_title('c)')
ax3.set_ylabel('Zonal Mean Precipitation [mm/day]')
ax3.set_xlabel('')
ax3.set_xlim([xlim1,xlim2])
ax3.set_ylim([ylim1,ylim2])
ax3.legend(fontsize=fs_legend)
ax4 = plt.subplot(324)
seas_pac_xtmean_seamask['imerg'].sel(season='MAM').plot(color='black', label = 'Obs. 1: IMERG')
seas_pac_xtmean['cmap'].sel(season='MAM').plot(color='black',dashes = [4,4,4,4], label='Obs. 2: CMAP')
seas_pac_xtmean['gpcp'].sel(season='MAM').plot(color='black',dashes=[2,1,2,1], label='Obs. 3: GPCP')
seas_pac_xtmean_seamask['cesm_ctrl'].sel(season='MAM').plot(color='blue', label = 'CCSM4 Control')
seas_pac_xtmean_seamask['cesm_cam'].sel(season='MAM').plot(color='red',dashes=[1,1,1,1], label = 'CCSM4 Ideal CAm')
ax4.set_title('d)')
ax4.set_xlabel('')
ax4.set_xlim([xlim1,xlim2])
ax4.set_ylim([ylim1,ylim2])
ax4.legend(fontsize=fs_legend)
ax5 = plt.subplot(325)
seas_pac_xtmean_seamask['imerg'].sel(season='SON').plot(color='black', label = 'Obs. 1: IMERG')
seas_pac_xtmean['cmap'].sel(season='SON').plot(color='black',dashes = [4,4,4,4], label='Obs. 2: CMAP')
seas_pac_xtmean['gpcp'].sel(season='SON').plot(color='black',dashes=[2,1,2,1], label='Obs. 3: GPCP')
seas_pac_xtmean_seamask['ctrl'].sel(season='SON').plot(color='blue', label = 'FLOR Control')
seas_pac_xtmean_seamask['hitopo'].sel(season='SON').plot(color='red', label = 'FLOR HiTopo')
seas_pac_xtmean_seamask['cam'].sel(season='SON').plot(color='red',dashes=[1,1,1,1], label = 'FLOR CAm')
ax5.set_title('e)')
ax5.set_ylabel('Zonal Mean Precipitation [mm/day]')
ax5.set_xlabel('Latitude [$^{\circ}$N]')
ax5.set_xlim([xlim1,xlim2])
ax5.set_ylim([ylim1,ylim2])
ax5.legend(fontsize=fs_legend)
ax6 = plt.subplot(326)
seas_pac_xtmean_seamask['imerg'].sel(season='SON').plot(color='black', label = 'Obs. 1: IMERG')
seas_pac_xtmean['cmap'].sel(season='SON').plot(color='black',dashes = [4,4,4,4], label='Obs. 2: CMAP')
seas_pac_xtmean['gpcp'].sel(season='SON').plot(color='black',dashes=[2,1,2,1], label='Obs. 3: GPCP')
seas_pac_xtmean_seamask['cesm_ctrl'].sel(season='SON').plot(color='blue', label = 'CCSM4 Control')
seas_pac_xtmean_seamask['cesm_cam'].sel(season='SON').plot(color='red',dashes=[1,1,1,1], label = 'CCSM4 Ideal CAm')
ax6.set_title('f)')
ax6.set_xlabel('Latitude [$^{\circ}$N]')
ax6.set_xlim([xlim1,xlim2])
ax6.set_ylim([ylim1,ylim2])
ax6.legend(fontsize=fs_legend)
t1 = ax1.text(0.5,1.15,'FLOR',fontsize=fs_label, fontweight='bold', style = 'italic', horizontalalignment='center', verticalalignment='center', transform=ax1.transAxes)
t1b = ax1.text(-0.15,0.5,'Annual Mean',fontsize=fs_label, fontweight='bold', style = 'italic', horizontalalignment='center', verticalalignment='center', rotation = 90, transform=ax1.transAxes)
t2 = ax2.text(0.5,1.15,'CCSM4',fontsize=fs_label, fontweight='bold', style = 'italic', horizontalalignment='center', verticalalignment='center', transform=ax2.transAxes)
t3 = ax3.text(-0.15,0.5,'MAM',fontsize=fs_label, fontweight='bold', style = 'italic', horizontalalignment='center', verticalalignment='center', rotation = 90, transform=ax3.transAxes)
t5 = ax5.text(-0.15,0.5,'SON',fontsize=fs_label, fontweight='bold', style = 'italic', horizontalalignment='center', verticalalignment='center', rotation = 90, transform=ax5.transAxes)
#plt.suptitle('East Pacific (130W to 75W)')
plt.tight_layout(pad=0.5)
#plt.savefig('precip_zonalmean_epac_bothmodels.png',dpi=600)
#plt.savefig('precip_zonalmean_epac_bothmodels.pdf')
# w/ stat sig: FLOR MAPS COMPARING DIFFERENT MODEL RUNS SEASON ZOOMED IN
fig = plt.figure(figsize=(10,9))
plt.rcParams.update({'font.size': 12})
lev = np.arange(0,13,1)
levb = np.arange(0,12,4)
lev2 = np.arange(-4.5,5,0.5)
cmap = cmocean.cm.rain
cmap2 =cmocean.cm.balance_r
cmap2r = cmocean.cm.balance
contour_color = 'darkslategrey'
lw = 0.5
proj = ccrs.PlateCarree(central_longitude=-180)
ymax = 23.5
ymin = -23.5
xmin = 120
xmax = 300
season = 'MAM'
ax1 = plt.subplot(5,2,1,projection=proj)
ax1.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im1 = plt.contourf(lonf,latf,seasmean['ctrl'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max')
ax1.coastlines()
ax1.set_title('a) FLOR Control')
gl = ax1.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax2 = plt.subplot(5,2,2,projection=proj)
ax2.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im2 = plt.contourf(lonf,latf,diff_mask['imerg'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
CS = plt.contour(lonf,latf,seasmean['ctrl'].sel(season=season),levels=levb,transform=ccrs.PlateCarree(),extend='max',colors=contour_color,linewidths=1)
ax2.clabel(CS, fontsize=9, inline=1, fmt = '%1.f')
ax2.coastlines()
ax2.set_title('b) Obs. - FLOR Control')
gl = ax2.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.ylabels_left = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax3 = plt.subplot(5,2,3,projection=proj)
ax3.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im3 = plt.contourf(lonc,latc,seasmean['cesm_ctrl'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max',color='k')
ax3.coastlines()
ax3.set_title('c) CCSM4 Control')
gl = ax3.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax4 = plt.subplot(5,2,4,projection=proj)
ax4.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im4 = plt.contourf(lonc,latc,diff_mask_cesm['imerg'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
CS = plt.contour(lonc,latc,seasmean['cesm_ctrl'].sel(season=season),levels=levb,transform=ccrs.PlateCarree(),extend='max',colors=contour_color,linewidths=1)
ax4.clabel(CS, fontsize=9, inline=1, fmt = '%1.f')
ax4.coastlines()
ax4.set_title('d) Obs. - CCSM4 Control')
gl = ax4.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.ylabels_left = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax5 = plt.subplot(5,2,5,projection=proj)
ax5.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im5 = plt.contourf(lonf,latf,seasmean['hitopo'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max')
ax5.coastlines()
ax5.set_title('e) FLOR HiTopo')
gl = ax5.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax6 = plt.subplot(5,2,6,projection=proj)
ax6.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im6 = plt.contourf(lonf,latf,diff_mask['hitopo'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
CS = plt.contour(lonf,latf,seasmean['ctrl'].sel(season=season),levels=levb,transform=ccrs.PlateCarree(),extend='max',colors=contour_color,linewidths=1)
ax6.clabel(CS, fontsize=9, inline=1, fmt = '%1.f')
ax6.coastlines()
ax6.set_title('f) FLOR HiTopo - FLOR Control')
gl = ax6.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.ylabels_left = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax7 = plt.subplot(5,2,7,projection=proj)
ax7.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im7 = plt.contourf(lonf,latf,seasmean['cam'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max')
ax7.coastlines()
ax7.set_title('g) FLOR CAm')
gl = ax7.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax8 = plt.subplot(5,2,8,projection=proj)
ax8.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im8 = plt.contourf(lonf,latf,diff_mask['cam'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
CS = plt.contour(lonf,latf,seasmean['ctrl'].sel(season=season),levels=levb,transform=ccrs.PlateCarree(),extend='max',colors=contour_color,linewidths=1)
ax8.clabel(CS, fontsize=9, inline=1, fmt = '%1.f')
ax8.coastlines()
ax8.set_title('h) FLOR CAm - FLOR Control')
gl = ax8.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = False
gl.ylabels_right = False
gl.ylabels_left = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax9 = plt.subplot(5,2,9,projection=proj)
ax9.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im9 = plt.contourf(lonc,latc,seasmean['cesm_cam'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max')
ax9.coastlines()
ax9.set_title('i) CCSM4 Ideal CAm')
gl = ax9.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = True
gl.ylabels_right = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
ax10 = plt.subplot(5,2,10,projection=proj)
ax10.set_extent([xmin, xmax, ymin, ymax], ccrs.PlateCarree())
im10 = plt.contourf(lonc,latc,diff_mask['cesm_cam'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
CS = plt.contour(lonc,latc,seasmean['cesm_ctrl'].sel(season=season),levels=levb,transform=ccrs.PlateCarree(),extend='max',colors=contour_color,linewidths=1)
ax10.clabel(CS, fontsize=9, inline=1, fmt = '%1.f')
ax10.coastlines()
ax10.set_title('j) CCSM4 Ideal CAm - CCSM4 Control')
gl = ax10.gridlines(crs=ccrs.PlateCarree(), draw_labels=True,
linewidth=lw, color='gray', alpha=0.5, linestyle='--')
gl.xlabels_top = False
gl.xlabels_bottom = True
gl.ylabels_right = False
gl.ylabels_left = False
gl.xlocator = mticker.FixedLocator([140,190,-170,-120,-70,0])
gl.ylocator = mticker.FixedLocator([-30,-20,-10,0,10,20,30])
#fig.subplots_adjust(wspace=0.7)
cb1_ax = fig.add_axes([0.15, 0.07, 0.3, 0.02]) # rect = l, b, w, h
cb1 = fig.colorbar(im1, cax=cb1_ax, orientation = 'horizontal')
cb1.ax.set_xlabel('precipitation [mm/day]', rotation=0)
cb2_ax = fig.add_axes([0.57, 0.07, 0.3, 0.02])
cb2 = fig.colorbar(im2, cax=cb2_ax, orientation = 'horizontal')
cb2.ax.set_xlabel('precipitation [mm/day]', rotation=0)
#plt.tight_layout()
#plt.savefig('precip_map_'+season+'_zoom.pdf')
# w/ stat sig: North America DJF
fig = plt.figure(figsize=(9,10))
plt.rcParams.update({'font.size': 14})
lev = np.arange(0,7.5,0.5)
lev2 = np.arange(-3,3.5,0.5)
cmap = cmocean.cm.rain
cmap2 =cmocean.cm.balance_r
cmap2r = cmocean.cm.balance
proj = ccrs.PlateCarree(central_longitude=-180)
xmin = -130
xmax = -65
ymin = 22
ymax = 60
season = 'DJF'
ax1 = plt.subplot(3,2,1,projection=proj)
im1 = plt.contourf(lonf,latf,seasmean['ctrl'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max')
ax1.coastlines()
ax1.add_feature(feature.BORDERS, linestyle='-', alpha=.5)
ax1.set_title('a) FLOR Control')
ax1.set_extent([xmin, xmax, ymin, ymax])
ax2 = plt.subplot(3,2,2,projection=proj)
im2 = plt.contourf(lonf,latf,diff_mask['imerg'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
ax2.coastlines()
ax2.add_feature(feature.BORDERS, linestyle='-', alpha=.5)
ax2.set_title('b) Obs. - FLOR Control')
ax2.set_extent([xmin, xmax, ymin, ymax])
ax3 = plt.subplot(3,2,3,projection=proj)
im1 = plt.contourf(lonf,latf,seasmean['hitopo'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max')
ax3.coastlines()
ax3.add_feature(feature.BORDERS, linestyle='-', alpha=.5)
ax3.add_feature(feature.BORDERS, linestyle='-', alpha=.5)
ax3.set_title('c) FLOR HiTopo')
ax3.set_extent([xmin, xmax, ymin, ymax])
ax4 = plt.subplot(3,2,4,projection=proj)
im4 = plt.contourf(lonf,latf,diff_mask['hitopo'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
ax4.coastlines()
ax4.add_feature(feature.BORDERS, linestyle='-', alpha=.5)
ax4.set_title('d) FLOR HiTopo - FLOR Control')
ax4.set_extent([xmin, xmax, ymin, ymax])
ax5 = plt.subplot(3,2,5,projection=proj)
im1 = plt.contourf(lonf,latf,seasmean['cam'].sel(season=season),levels=lev,cmap=cmap,transform=ccrs.PlateCarree(),extend='max')
ax5.coastlines()
ax5.add_feature(feature.BORDERS, linestyle='-', alpha=.5)
ax5.set_title('e) FLOR CAm')
ax5.set_extent([xmin, xmax, ymin, ymax])
ax6 = plt.subplot(3,2,6,projection=proj)
im6 = plt.contourf(lonf,latf,diff_mask['cam'][season],levels=lev2,cmap=cmap2,transform=ccrs.PlateCarree(),extend='both')
ax6.coastlines()
ax6.add_feature(feature.BORDERS, linestyle='-', alpha=.5)
ax6.set_title('f) FLOR CAm - FLOR Control')
ax6.set_extent([xmin, xmax, ymin, ymax])
#fig.subplots_adjust(wspace=0.7)
cb1_ax = fig.add_axes([0.15, 0.07, 0.3, 0.02]) # rect = l, b, w, h
cb1 = fig.colorbar(im1, cax=cb1_ax, orientation = 'horizontal')
cb1.ax.set_xlabel('precipitation [mm/day]', rotation=0)
cb2_ax = fig.add_axes([0.57, 0.07, 0.3, 0.02])
cb2 = fig.colorbar(im2, cax=cb2_ax, orientation = 'horizontal')
cb2.ax.set_xlabel('precipitation [mm/day]', rotation=0)
#plt.tight_layout(w_pad = 7)
#plt.savefig('us_precip_map_djf.png',dpi = 600)
#plt.savefig('us_precip_map_djf.pdf')
```
| github_jupyter |
# First steps with xmovie
```
# load modules
import xarray as xr
from xmovie import Movie
%matplotlib inline
# load test dataset
ds = xr.tutorial.open_dataset('air_temperature').isel(time=slice(0,150))
# create movie object
mov = Movie(ds.air)
# preview 10th frame
mov.preview(10)
! rm frame*.png rm *.mp4 *.mp4
mov.save('movie.mp4')# Use to save a high quality mp4 movie
mov.save('movie_gif.gif') #Use to save a gif
```
In many cases it is useful to have both a high quality movie and a lower resolution gif of the same animation. If that is desired, just deactivate the `remove_movie` option and give a filename with `.gif`. xmovie will first render a high quality movie and then convert it to a gif, without removing the movie afterwards.
```
# Display a progressbar with `progress`=True, (requires tqdm). This can be helpful for long running animations.
mov.save('movie_combo.gif', remove_movie=False, progress=True)
# Modify the framerate of the output with the keyword arguments `framerate` (for movies) and `gif_framerate` (for gifs)
mov.save('movie_fast.gif', remove_movie=False, progress=True, framerate=20, gif_framerate=20)
mov.save('movie_slow.gif', remove_movie=False, progress=True, framerate=5, gif_framerate=5)
```



```
from xmovie.presets import rotating_globe
# specify custom plotfunctions (here a preset from xmovie)
mov = Movie(ds.air, plotfunc=rotating_globe)
mov.save('movie_rotating.gif', progress=True)
```

```
mov = Movie(ds.air, plotfunc=rotating_globe, style='dark')
mov.save('movie_rotating_dark.gif', progress=True)
```

```
# Change the plotting function with the parameter `plotmethod`
mov = Movie(ds.air, rotating_globe, plotmethod='contour')
mov.save('movie_cont.gif')
mov = Movie(ds.air, rotating_globe, plotmethod='contourf')
mov.save('movie_contf.gif')
```


```
import numpy as np
ds = xr.tutorial.open_dataset('rasm', decode_times=False).Tair
# Interpolate time for smoother animation
ds['time'].data = np.arange(len(ds['time']))
ds = ds.interp(time=np.linspace(0,10, 60))
# `Movie` accepts keywords for the xarray plotting interface and provides a set of 'own' keywords like
# `coast`, `land` and `style` to facilitate the styling of plots
mov = Movie(ds, rotating_globe,
cmap='RdYlBu_r',
x='xc',
y='yc', #accepts keyword arguments from the xarray plotting interface
lat_start=45, # Custom keywords from `rotating_globe_dark
lat_rotations=0.05,
lon_rotations=0.2,
land=False,
coastline=True,
style='dark')
mov.save('movie_rasm.gif', progress=True)
# mov.preview(10)
```

Besides the presets xmovie is designed to animate any custom plot which can be wrapped in a function acting on a matplotlib figure. This can contain xarray plotting commands, 'pure' matplotlib or a combination of both. This can come in handy when you want to animate a complex static plot.
```
# some awesome static plot
import matplotlib.pyplot as plt
ds = xr.tutorial.open_dataset('rasm', decode_times=False).Tair
fig = plt.figure(figsize=[10,5])
tt = 30
station = dict(x=100, y=150)
ds_station = ds.sel(**station)
(ax1, ax2) = fig.subplots(ncols=2)
ds.isel(time=tt).plot(ax=ax1)
ax1.plot(station['x'], station['y'], marker='*', color='k' ,markersize=15)
ax1.text(station['x']+4, station['y']+4, 'Station', color='k' )
ax1.set_aspect(1)
ax1.set_facecolor('0.5')
ax1.set_title('');
# Time series
ds_station.isel(time=slice(0,tt+1)).plot.line(ax=ax2, x='time')
ax2.set_xlim(ds.time.min().data, ds.time.max().data)
ax2.set_ylim(ds_station.min(), ds_station.max())
ax2.set_title('Data at station');
fig.subplots_adjust(wspace=0.6)
```
All you need to do is wrap your plotting calls into a functions `func(ds, fig, frame)`, where ds is an xarray dataset you pass to `Movie`, fig is a matplotlib.figure handle and tt is the movie frame.
```
def custom_plotfunc(ds, fig, tt):
# Define station location for timeseries
station = dict(x=100, y=150)
ds_station = ds.sel(**station)
(ax1, ax2) = fig.subplots(ncols=2)
# Map axis
# Colorlimits need to be fixed or your video is going to cause seizures.
# This is the only modification from the code above!
ds.isel(time=tt).plot(ax=ax1, vmin=ds.min(), vmax=ds.max(), cmap='RdBu_r')
ax1.plot(station['x'], station['y'], marker='*', color='k' ,markersize=15)
ax1.text(station['x']+4, station['y']+4, 'Station', color='k' )
ax1.set_aspect(1)
ax1.set_facecolor('0.5')
ax1.set_title('');
# Time series
ds_station.isel(time=slice(0,tt+1)).plot.line(ax=ax2, x='time')
ax2.set_xlim(ds.time.min().data, ds.time.max().data)
ax2.set_ylim(ds_station.min(), ds_station.max())
ax2.set_title('Data at station');
fig.subplots_adjust(wspace=0.6)
return None, None #This is not strictly necessary, but otherwise a warning will be raised.
mov_custom = Movie(ds, custom_plotfunc)
mov_custom.preview(30)
mov_custom.save('movie_custom.gif', progress=True)
```

| github_jupyter |
# Bayesian Temporal Tensor Factorization
**Published**: December 27, 2020
**Author**: Xinyu Chen [[**GitHub homepage**](https://github.com/xinychen)]
**Download**: This Jupyter notebook is at our GitHub repository. If you want to evaluate the code, please download the notebook from the [**transdim**](https://github.com/xinychen/transdim/blob/master/predictor/BTTF.ipynb) repository.
This notebook shows how to implement the Bayesian Temporal Tensor Factorization (BTTF), a fully Bayesian matrix factorization model, on some real-world data sets. To overcome the missing data problem in multivariate time series, BTTF takes into account both low-rank matrix structure and time series autoregression. For an in-depth discussion of BTTF, please see [1].
<div class="alert alert-block alert-info">
<font color="black">
<b>[1]</b> Xinyu Chen, Lijun Sun (2019). <b>Bayesian temporal factorization for multidimensional time series prediction</b>. arXiv:1910.06366. <a href="https://arxiv.org/pdf/1910.06366.pdf" title="PDF"><b>[PDF]</b></a>
</font>
</div>
```
import numpy as np
from numpy.linalg import inv as inv
from numpy.random import normal as normrnd
from numpy.random import multivariate_normal as mvnrnd
from scipy.linalg import khatri_rao as kr_prod
from scipy.stats import wishart
from scipy.stats import invwishart
from numpy.linalg import solve as solve
from numpy.linalg import cholesky as cholesky_lower
from scipy.linalg import cholesky as cholesky_upper
from scipy.linalg import solve_triangular as solve_ut
import matplotlib.pyplot as plt
%matplotlib inline
def mvnrnd_pre(mu, Lambda):
src = normrnd(size = (mu.shape[0],))
return solve_ut(cholesky_upper(Lambda, overwrite_a = True, check_finite = False),
src, lower = False, check_finite = False, overwrite_b = True) + mu
def cov_mat(mat, mat_bar):
mat = mat - mat_bar
return mat.T @ mat
def ten2mat(tensor, mode):
return np.reshape(np.moveaxis(tensor, mode, 0), (tensor.shape[mode], -1), order = 'F')
def sample_factor_u(tau_sparse_tensor, tau_ind, U, V, X, beta0 = 1):
"""Sampling M-by-R factor matrix U and its hyperparameters (mu_u, Lambda_u)."""
dim1, rank = U.shape
U_bar = np.mean(U, axis = 0)
temp = dim1 / (dim1 + beta0)
var_mu_hyper = temp * U_bar
var_U_hyper = inv(np.eye(rank) + cov_mat(U, U_bar) + temp * beta0 * np.outer(U_bar, U_bar))
var_Lambda_hyper = wishart.rvs(df = dim1 + rank, scale = var_U_hyper)
var_mu_hyper = mvnrnd_pre(var_mu_hyper, (dim1 + beta0) * var_Lambda_hyper)
var1 = kr_prod(X, V).T
var2 = kr_prod(var1, var1)
var3 = (var2 @ ten2mat(tau_ind, 0).T).reshape([rank, rank, dim1]) + var_Lambda_hyper[:, :, None]
var4 = var1 @ ten2mat(tau_sparse_tensor, 0).T + (var_Lambda_hyper @ var_mu_hyper)[:, None]
for i in range(dim1):
U[i, :] = mvnrnd_pre(solve(var3[:, :, i], var4[:, i]), var3[:, :, i])
return U
def sample_factor_v(tau_sparse_tensor, tau_ind, U, V, X, beta0 = 1):
"""Sampling N-by-R factor matrix V and its hyperparameters (mu_v, Lambda_v)."""
dim2, rank = V.shape
V_bar = np.mean(V, axis = 0)
temp = dim2 / (dim2 + beta0)
var_mu_hyper = temp * V_bar
var_V_hyper = inv(np.eye(rank) + cov_mat(V, V_bar) + temp * beta0 * np.outer(V_bar, V_bar))
var_Lambda_hyper = wishart.rvs(df = dim2 + rank, scale = var_V_hyper)
var_mu_hyper = mvnrnd_pre(var_mu_hyper, (dim2 + beta0) * var_Lambda_hyper)
var1 = kr_prod(X, U).T
var2 = kr_prod(var1, var1)
var3 = (var2 @ ten2mat(tau_ind, 1).T).reshape([rank, rank, dim2]) + var_Lambda_hyper[:, :, None]
var4 = var1 @ ten2mat(tau_sparse_tensor, 1).T + (var_Lambda_hyper @ var_mu_hyper)[:, None]
for j in range(dim2):
V[j, :] = mvnrnd_pre(solve(var3[:, :, j], var4[:, j]), var3[:, :, j])
return V
def mnrnd(M, U, V):
"""
Generate matrix normal distributed random matrix.
M is a m-by-n matrix, U is a m-by-m matrix, and V is a n-by-n matrix.
"""
dim1, dim2 = M.shape
X0 = np.random.randn(dim1, dim2)
P = cholesky_lower(U)
Q = cholesky_lower(V)
return M + P @ X0 @ Q.T
def sample_var_coefficient(X, time_lags):
dim, rank = X.shape
d = time_lags.shape[0]
tmax = np.max(time_lags)
Z_mat = X[tmax : dim, :]
Q_mat = np.zeros((dim - tmax, rank * d))
for k in range(d):
Q_mat[:, k * rank : (k + 1) * rank] = X[tmax - time_lags[k] : dim - time_lags[k], :]
var_Psi0 = np.eye(rank * d) + Q_mat.T @ Q_mat
var_Psi = inv(var_Psi0)
var_M = var_Psi @ Q_mat.T @ Z_mat
var_S = np.eye(rank) + Z_mat.T @ Z_mat - var_M.T @ var_Psi0 @ var_M
Sigma = invwishart.rvs(df = rank + dim - tmax, scale = var_S)
return mnrnd(var_M, var_Psi, Sigma), Sigma
def sample_factor_x(tau_sparse_tensor, tau_ind, time_lags, U, V, X, A, Lambda_x):
"""Sampling T-by-R factor matrix X."""
dim3, rank = X.shape
tmax = np.max(time_lags)
tmin = np.min(time_lags)
d = time_lags.shape[0]
A0 = np.dstack([A] * d)
for k in range(d):
A0[k * rank : (k + 1) * rank, :, k] = 0
mat0 = Lambda_x @ A.T
mat1 = np.einsum('kij, jt -> kit', A.reshape([d, rank, rank]), Lambda_x)
mat2 = np.einsum('kit, kjt -> ij', mat1, A.reshape([d, rank, rank]))
var1 = kr_prod(V, U).T
var2 = kr_prod(var1, var1)
var3 = (var2 @ ten2mat(tau_ind, 2).T).reshape([rank, rank, dim3]) + Lambda_x[:, :, None]
var4 = var1 @ ten2mat(tau_sparse_tensor, 2).T
for t in range(dim3):
Mt = np.zeros((rank, rank))
Nt = np.zeros(rank)
Qt = mat0 @ X[t - time_lags, :].reshape(rank * d)
index = list(range(0, d))
if t >= dim3 - tmax and t < dim3 - tmin:
index = list(np.where(t + time_lags < dim3))[0]
elif t < tmax:
Qt = np.zeros(rank)
index = list(np.where(t + time_lags >= tmax))[0]
if t < dim3 - tmin:
Mt = mat2.copy()
temp = np.zeros((rank * d, len(index)))
n = 0
for k in index:
temp[:, n] = X[t + time_lags[k] - time_lags, :].reshape(rank * d)
n += 1
temp0 = X[t + time_lags[index], :].T - np.einsum('ijk, ik -> jk', A0[:, :, index], temp)
Nt = np.einsum('kij, jk -> i', mat1[index, :, :], temp0)
var3[:, :, t] = var3[:, :, t] + Mt
if t < tmax:
var3[:, :, t] = var3[:, :, t] - Lambda_x + np.eye(rank)
X[t, :] = mvnrnd_pre(solve(var3[:, :, t], var4[:, t] + Nt + Qt), var3[:, :, t])
return X
def compute_mape(var, var_hat):
return np.sum(np.abs(var - var_hat) / var) / var.shape[0]
def compute_rmse(var, var_hat):
return np.sqrt(np.sum((var - var_hat) ** 2) / var.shape[0])
def ar4cast(A, X, Sigma, time_lags, multi_step):
dim, rank = X.shape
d = time_lags.shape[0]
X_new = np.append(X, np.zeros((multi_step, rank)), axis = 0)
for t in range(multi_step):
var = A.T @ X_new[dim + t - time_lags, :].reshape(rank * d)
X_new[dim + t, :] = mvnrnd(var, Sigma)
return X_new
```
#### BTTF Implementation
```
def BTTF(dense_tensor, sparse_tensor, init, rank, time_lags, burn_iter, gibbs_iter, multi_step = 1):
"""Bayesian Temporal Tensor Factorization, BTTF."""
dim1, dim2, dim3 = sparse_tensor.shape
d = time_lags.shape[0]
U = init["U"]
V = init["V"]
X = init["X"]
if np.isnan(sparse_tensor).any() == False:
ind = sparse_tensor != 0
pos_obs = np.where(ind)
pos_test = np.where((dense_tensor != 0) & (sparse_tensor == 0))
elif np.isnan(sparse_tensor).any() == True:
pos_test = np.where((dense_tensor != 0) & (np.isnan(sparse_tensor)))
ind = ~np.isnan(sparse_tensor)
pos_obs = np.where(ind)
sparse_tensor[np.isnan(sparse_tensor)] = 0
dense_test = dense_tensor[pos_test]
del dense_tensor
U_plus = np.zeros((dim1, rank, gibbs_iter))
V_plus = np.zeros((dim2, rank, gibbs_iter))
X_plus = np.zeros((dim3 + multi_step, rank, gibbs_iter))
A_plus = np.zeros((rank * d, rank, gibbs_iter))
tau_plus = np.zeros(gibbs_iter)
Sigma_plus = np.zeros((rank, rank, gibbs_iter))
temp_hat = np.zeros(len(pos_test[0]))
show_iter = 500
tau = 1
tensor_hat_plus = np.zeros(sparse_tensor.shape)
tensor_new_plus = np.zeros((dim1, dim2, multi_step))
for it in range(burn_iter + gibbs_iter):
tau_ind = tau * ind
tau_sparse_tensor = tau * sparse_tensor
U = sample_factor_u(tau_sparse_tensor, tau_ind, U, V, X)
V = sample_factor_v(tau_sparse_tensor, tau_ind, U, V, X)
A, Sigma = sample_var_coefficient(X, time_lags)
X = sample_factor_x(tau_sparse_tensor, tau_ind, time_lags, U, V, X, A, inv(Sigma))
tensor_hat = np.einsum('is, js, ts -> ijt', U, V, X)
tau = np.random.gamma(1e-6 + 0.5 * np.sum(ind),
1 / (1e-6 + 0.5 * np.sum(((sparse_tensor - tensor_hat) ** 2) * ind)))
temp_hat += tensor_hat[pos_test]
if (it + 1) % show_iter == 0 and it < burn_iter:
temp_hat = temp_hat / show_iter
print('Iter: {}'.format(it + 1))
print('MAPE: {:.6}'.format(compute_mape(dense_test, temp_hat)))
print('RMSE: {:.6}'.format(compute_rmse(dense_test, temp_hat)))
temp_hat = np.zeros(len(pos_test[0]))
print()
if it + 1 > burn_iter:
U_plus[:, :, it - burn_iter] = U
V_plus[:, :, it - burn_iter] = V
A_plus[:, :, it - burn_iter] = A
Sigma_plus[:, :, it - burn_iter] = Sigma
tau_plus[it - burn_iter] = tau
tensor_hat_plus += tensor_hat
X0 = ar4cast(A, X, Sigma, time_lags, multi_step)
X_plus[:, :, it - burn_iter] = X0
tensor_new_plus += np.einsum('is, js, ts -> ijt', U, V, X0[- multi_step :, :])
tensor_hat = tensor_hat_plus / gibbs_iter
print('Imputation MAPE: {:.6}'.format(compute_mape(dense_test, tensor_hat[:, :, : dim3][pos_test])))
print('Imputation RMSE: {:.6}'.format(compute_rmse(dense_test, tensor_hat[:, :, : dim3][pos_test])))
print()
tensor_hat = np.append(tensor_hat, tensor_new_plus / gibbs_iter, axis = 2)
tensor_hat[tensor_hat < 0] = 0
return tensor_hat, U_plus, V_plus, X_plus, A_plus, Sigma_plus, tau_plus
def sample_factor_x_partial(tau_sparse_tensor, tau_ind, time_lags, U, V, X, A, Lambda_x, back_step):
"""Sampling T-by-R factor matrix X."""
dim3, rank = X.shape
tmax = np.max(time_lags)
tmin = np.min(time_lags)
d = time_lags.shape[0]
A0 = np.dstack([A] * d)
for k in range(d):
A0[k * rank : (k + 1) * rank, :, k] = 0
mat0 = Lambda_x @ A.T
mat1 = np.einsum('kij, jt -> kit', A.reshape([d, rank, rank]), Lambda_x)
mat2 = np.einsum('kit, kjt -> ij', mat1, A.reshape([d, rank, rank]))
var1 = kr_prod(V, U).T
var2 = kr_prod(var1, var1)
var3 = (var2 @ ten2mat(tau_ind[:, :, - back_step :], 2).T).reshape([rank, rank, back_step]) + Lambda_x[:, :, None]
var4 = var1 @ ten2mat(tau_sparse_tensor[:, :, - back_step :], 2).T
for t in range(dim3 - back_step, dim3):
Mt = np.zeros((rank, rank))
Nt = np.zeros(rank)
Qt = mat0 @ X[t - time_lags, :].reshape(rank * d)
index = list(range(0, d))
if t >= dim3 - tmax and t < dim3 - tmin:
index = list(np.where(t + time_lags < dim3))[0]
if t < dim3 - tmin:
Mt = mat2.copy()
temp = np.zeros((rank * d, len(index)))
n = 0
for k in index:
temp[:, n] = X[t + time_lags[k] - time_lags, :].reshape(rank * d)
n += 1
temp0 = X[t + time_lags[index], :].T - np.einsum('ijk, ik -> jk', A0[:, :, index], temp)
Nt = np.einsum('kij, jk -> i', mat1[index, :, :], temp0)
var3[:, :, t + back_step - dim3] = var3[:, :, t + back_step - dim3] + Mt
X[t, :] = mvnrnd_pre(solve(var3[:, :, t + back_step - dim3],
var4[:, t + back_step - dim3] + Nt + Qt), var3[:, :, t + back_step - dim3])
return X
def BTTF_partial(dense_tensor, sparse_tensor, init, rank, time_lags, burn_iter, gibbs_iter, multi_step = 1):
"""Bayesian Temporal Tensor Factorization, BTTF."""
dim1, dim2, dim3 = sparse_tensor.shape
U_plus = init["U_plus"]
V_plus = init["V_plus"]
X_plus = init["X_plus"]
A_plus = init["A_plus"]
Sigma_plus = init["Sigma_plus"]
tau_plus = init["tau_plus"]
if np.isnan(sparse_tensor).any() == False:
ind = sparse_tensor != 0
pos_obs = np.where(ind)
elif np.isnan(sparse_tensor).any() == True:
ind = ~np.isnan(sparse_tensor)
pos_obs = np.where(ind)
sparse_tensor[np.isnan(sparse_tensor)] = 0
X_new_plus = np.zeros((dim3 + multi_step, rank, gibbs_iter))
tensor_new_plus = np.zeros((dim1, dim2, multi_step))
back_step = 10 * multi_step
for it in range(gibbs_iter):
tau_ind = tau_plus[it] * ind
tau_sparse_tensor = tau_plus[it] * sparse_tensor
X = sample_factor_x_partial(tau_sparse_tensor, tau_ind, time_lags, U_plus[:, :, it], V_plus[:, :, it],
X_plus[:, :, it], A_plus[:, :, it], inv(Sigma_plus[:, :, it]), back_step)
X0 = ar4cast(A_plus[:, :, it], X, Sigma_plus[:, :, it], time_lags, multi_step)
X_new_plus[:, :, it] = X0
tensor_new_plus += np.einsum('is, js, ts -> ijt', U_plus[:, :, it], V_plus[:, :, it], X0[- multi_step :, :])
tensor_hat = tensor_new_plus / gibbs_iter
tensor_hat[tensor_hat < 0] = 0
return tensor_hat, U_plus, V_plus, X_new_plus, A_plus, Sigma_plus, tau_plus
from ipywidgets import IntProgress
from IPython.display import display
def BTTF_forecast(dense_tensor, sparse_tensor, pred_step, multi_step, rank, time_lags, burn_iter, gibbs_iter):
dim1, dim2, T = dense_tensor.shape
start_time = T - pred_step
max_count = int(np.ceil(pred_step / multi_step))
tensor_hat = np.zeros((dim1, dim2, max_count * multi_step))
f = IntProgress(min = 0, max = max_count) # instantiate the bar
display(f) # display the bar
for t in range(max_count):
if t == 0:
init = {"U": 0.1 * np.random.randn(dim1, rank),
"V": 0.1 * np.random.randn(dim2, rank),
"X": 0.1 * np.random.randn(start_time, rank)}
tensor, U, V, X_new, A, Sigma, tau = BTTF(dense_tensor[:, :, : start_time],
sparse_tensor[:, :, : start_time], init, rank, time_lags, burn_iter, gibbs_iter, multi_step)
else:
init = {"U_plus": U, "V_plus": V, "X_plus": X_new, "A_plus": A, "Sigma_plus": Sigma, "tau_plus": tau}
tensor, U, V, X_new, A, Sigma, tau = BTTF_partial(dense_tensor[:, :, : start_time + t * multi_step],
sparse_tensor[:, :, : start_time + t * multi_step], init, rank, time_lags, burn_iter, gibbs_iter, multi_step)
tensor_hat[:, :, t * multi_step : (t + 1) * multi_step] = tensor[:, :, - multi_step :]
f.value = t
small_dense_tensor = dense_tensor[:, :, start_time : T]
pos = np.where(small_dense_tensor != 0)
print('Prediction MAPE: {:.6}'.format(compute_mape(small_dense_tensor[pos], tensor_hat[pos])))
print('Prediction RMSE: {:.6}'.format(compute_rmse(small_dense_tensor[pos], tensor_hat[pos])))
print()
return tensor_hat
```
## Eavluation on NYC Taxi Flow Data
**Scenario setting**:
- Tensor size: $30\times 30\times 1461$ (origin, destination, time)
- Test on original data
```
import scipy.io
import warnings
warnings.simplefilter('ignore')
dense_tensor = scipy.io.loadmat('../datasets/NYC-data-set/tensor.mat')['tensor'].astype(np.float32)
rm_tensor = scipy.io.loadmat('../datasets/NYC-data-set/rm_tensor.mat')['rm_tensor']
sparse_tensor = dense_tensor.copy()
```
**Model setting**:
- Low rank: 30
- Total (rolling) prediction horizons: 7 * 24
- Time lags: {1, 2, 24, 24 + 1, 24 + 2, 7 * 24, 7 * 24 + 1, 7 * 24 + 2}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
```
import time
rank = 30
pred_step = 7 * 24
time_lags = np.array([1, 2, 3, 24, 25, 26, 7 * 24, 7 * 24 + 1, 7 * 24 + 2])
burn_iter = 1000
gibbs_iter = 200
multi_step = 2
start = time.time()
print('Prediction time horizon (delta) = {}.'.format(multi_step))
tensor_hat = BTTF_forecast(dense_tensor, sparse_tensor, pred_step, multi_step, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
print()
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('classic')
plt.style.use('ggplot')
plt.style.use('bmh')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize = (4, 3.1))
ax = fig.add_axes([0.12, 0.14, 0.80, 0.83])
ax = sns.heatmap(tensor_hat[:, :, 24 * 3 + 8], cmap = 'OrRd', vmin = 0, vmax = 200, linewidth = 0.01,
cbar_kws={'label': 'Volume'})
plt.xticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
plt.yticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
ax.set_xlabel("Zone")
ax.set_ylabel("Zone")
plt.show()
fig.savefig("../images/Ndata_heatmap_predicted_values_32_0.pdf")
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('classic')
plt.style.use('ggplot')
plt.style.use('bmh')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize = (4, 3.1))
ax = fig.add_axes([0.12, 0.14, 0.80, 0.83])
ax = sns.heatmap(tensor_hat[:, :, 24 * 3 + 9], cmap = 'OrRd', vmin = 0, vmax = 150, linewidth = 0.01,
cbar_kws={'label': 'Volume'})
plt.xticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
plt.yticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
ax.set_xlabel("Zone")
ax.set_ylabel("Zone")
plt.show()
fig.savefig("../images/Ndata_heatmap_predicted_values_33_0.pdf")
```
**Scenario setting**:
- Tensor size: $30\times 30\times 1461$ (origin, destination, time)
- Non-random missing (NM)
- 40% missing rate
```
import scipy.io
dense_tensor = scipy.io.loadmat('../datasets/NYC-data-set/tensor.mat')['tensor'].astype(np.float32)
nm_tensor = scipy.io.loadmat('../datasets/NYC-data-set/nm_tensor.mat')['nm_tensor']
missing_rate = 0.4
## Non-random missing (NM)
binary_tensor = np.zeros(dense_tensor.shape)
for i1 in range(dense_tensor.shape[0]):
for i2 in range(dense_tensor.shape[1]):
for i3 in range(61):
binary_tensor[i1, i2, i3 * 24 : (i3 + 1) * 24] = np.round(nm_tensor[i1, i2, i3] + 0.5 - missing_rate)
sparse_tensor = dense_tensor.copy()
sparse_tensor[binary_tensor == 0] = np.nan
```
**Model setting**:
- Low rank: 30
- Total (rolling) prediction horizons: 7 * 24
- Time lags: {1, 2, 24, 24 + 1, 24 + 2, 7 * 24, 7 * 24 + 1, 7 * 24 + 2}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
```
import time
rank = 30
pred_step = 7 * 24
time_lags = np.array([1, 2, 3, 24, 25, 26, 7 * 24, 7 * 24 + 1, 7 * 24 + 2])
burn_iter = 1000
gibbs_iter = 200
multi_step = 2
start = time.time()
print('Prediction time horizon (delta) = {}.'.format(multi_step))
tensor_hat = BTTF_forecast(dense_tensor, sparse_tensor, pred_step, multi_step, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
print()
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('classic')
plt.style.use('ggplot')
plt.style.use('bmh')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize = (4, 3.1))
ax = fig.add_axes([0.12, 0.14, 0.80, 0.83])
ax = sns.heatmap(tensor_hat[:, :, 24 * 3 + 8], cmap = 'OrRd', vmin = 0, vmax = 200, linewidth = 0.01,
cbar_kws={'label': 'Volume'})
plt.xticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
plt.yticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
ax.set_xlabel("Zone")
ax.set_ylabel("Zone")
plt.show()
fig.savefig("../images/Ndata_heatmap_predicted_values_32.pdf")
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('classic')
plt.style.use('ggplot')
plt.style.use('bmh')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize = (4, 3.1))
ax = fig.add_axes([0.12, 0.14, 0.80, 0.83])
ax = sns.heatmap(tensor_hat[:, :, 24 * 3 + 9], cmap = 'OrRd', vmin = 0, vmax = 150, linewidth = 0.01,
cbar_kws={'label': 'Volume'})
plt.xticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
plt.yticks(np.arange(0.5, 30, 2), ["1", "3", "5", "7", "9",
"11", "13", "15", "17", "19",
"21", "23", "25", "27", "29"], rotation = 0)
ax.set_xlabel("Zone")
ax.set_ylabel("Zone")
plt.show()
fig.savefig("../images/Ndata_heatmap_predicted_values_33.pdf")
import matplotlib.pyplot as plt
import matplotlib.patches as patches
small_dense_tensor = dense_tensor[:, :, - pred_step :]
small_sparse_tensor = sparse_tensor[:, :, - pred_step :]
plt.style.use('classic')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize=(4.25, 1.55))
ax = fig.add_axes([0.14, 0.20, 0.84, 0.75])
plt.plot(small_dense_tensor[16, 12, :], color = "#006ea3",linewidth = 1.0, label = "Actual value")
plt.plot(tensor_hat[16, 12, :], color = "#e3120b", linewidth = 1.5, label = "Predicted value")
ax.set_xlim([0, pred_step])
ax.set_ylim([0, 150])
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.1, alpha = 0.2)
for j in range(7):
if small_sparse_tensor[16, 12, 24 * j] > 0:
someX, someY = j * 24, 0
currentAxis = plt.gca()
ax.add_patch(patches.Rectangle((someX, someY), 24, 300, alpha = 0.1, facecolor = 'green'))
plt.xticks(np.arange(0, 7 * 24 + 1, 24))
plt.yticks(np.arange(0, 150, 40))
ax.set_ylabel("Volume")
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.4, alpha = 0.5, axis = 'x')
plt.show()
fig.savefig("../images/NYC_time_series_volume_1713.pdf")
import matplotlib.pyplot as plt
import matplotlib.patches as patches
small_dense_tensor = dense_tensor[:, :, - pred_step :]
small_sparse_tensor = sparse_tensor[:, :, - pred_step :]
plt.style.use('classic')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize=(4.25, 1.55))
ax = fig.add_axes([0.14, 0.20, 0.84, 0.75])
plt.plot(small_dense_tensor[26, 26, :], color = "#006ea3",linewidth = 1.0, label = "Actual value")
plt.plot(tensor_hat[26, 26, :], color = "#e3120b", linewidth = 1.5, label = "Predicted value")
ax.set_xlim([0, pred_step])
ax.set_ylim([0, 210])
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.1, alpha = 0.2)
for j in range(7):
if small_sparse_tensor[26, 26, 24 * j] > 0:
someX, someY = j * 24, 0
currentAxis = plt.gca()
ax.add_patch(patches.Rectangle((someX, someY), 24, 300, alpha = 0.1, facecolor = 'green'))
plt.xticks(np.arange(0, 7 * 24 + 1, 24))
plt.yticks(np.arange(0, 210, 50))
ax.set_ylabel("Volume")
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.4, alpha = 0.5, axis = 'x')
plt.show()
fig.savefig("../images/NYC_time_series_volume_2727.pdf")
import matplotlib.pyplot as plt
import matplotlib.patches as patches
small_dense_tensor = dense_tensor[:, :, - pred_step :]
small_sparse_tensor = sparse_tensor[:, :, - pred_step :]
plt.style.use('classic')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize=(4.25, 1.55))
ax = fig.add_axes([0.14, 0.20, 0.84, 0.75])
plt.plot(small_dense_tensor[16, 26, :], color = "#006ea3",linewidth = 1.0, label = "Actual value")
plt.plot(tensor_hat[16, 26, :], color = "#e3120b", linewidth = 1.5, label = "Predicted value")
ax.set_xlim([0, pred_step])
ax.set_ylim([0, 210])
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.1, alpha = 0.2)
for j in range(7):
if small_sparse_tensor[16, 26, 24 * j] > 0:
someX, someY = j * 24, 0
currentAxis = plt.gca()
ax.add_patch(patches.Rectangle((someX, someY), 24, 300, alpha = 0.1, facecolor = 'green'))
plt.xticks(np.arange(0, 7 * 24 + 1, 24))
plt.yticks(np.arange(0, 210, 50))
ax.set_ylabel("Volume")
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.4, alpha = 0.5, axis = 'x')
plt.show()
fig.savefig("../images/NYC_time_series_volume_1727.pdf")
import matplotlib.pyplot as plt
import matplotlib.patches as patches
small_dense_tensor = dense_tensor[:, :, - pred_step :]
small_sparse_tensor = sparse_tensor[:, :, - pred_step :]
plt.style.use('classic')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize=(4.25, 1.55))
ax = fig.add_axes([0.14, 0.20, 0.84, 0.75])
plt.plot(small_dense_tensor[16, 20, :], color = "#006ea3",linewidth = 1.0, label = "Actual value")
plt.plot(tensor_hat[16, 20, :], color = "#e3120b", linewidth = 1.5, label = "Predicted value")
ax.set_xlim([0, pred_step])
ax.set_ylim([0, 170])
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.1, alpha = 0.2)
for j in range(7):
if small_sparse_tensor[16, 20, 24 * j] > 0:
someX, someY = j * 24, 0
currentAxis = plt.gca()
ax.add_patch(patches.Rectangle((someX, someY), 24, 300, alpha = 0.1, facecolor = 'green'))
plt.xticks(np.arange(0, 7 * 24 + 1, 24))
plt.yticks(np.arange(0, 170, 40))
ax.set_ylabel("Volume")
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.4, alpha = 0.5, axis = 'x')
plt.show()
fig.savefig("../images/NYC_time_series_volume_1721.pdf")
import matplotlib.pyplot as plt
import matplotlib.patches as patches
small_dense_tensor = dense_tensor[:, :, - pred_step :]
small_sparse_tensor = sparse_tensor[:, :, - pred_step :]
plt.style.use('classic')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize=(4.25, 1.55))
ax = fig.add_axes([0.14, 0.20, 0.84, 0.75])
plt.plot(small_dense_tensor[12, 20, :], color = "#006ea3",linewidth = 1.0, label = "Actual value")
plt.plot(tensor_hat[12, 20, :], color = "#e3120b", linewidth = 1.5, label = "Predicted value")
ax.set_xlim([0, pred_step])
ax.set_ylim([0, 60])
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.1, alpha = 0.2)
for j in range(7):
if small_sparse_tensor[12, 20, 24 * j] > 0:
someX, someY = j * 24, 0
currentAxis = plt.gca()
ax.add_patch(patches.Rectangle((someX, someY), 24, 300, alpha = 0.1, facecolor = 'green'))
plt.xticks(np.arange(0, 7 * 24 + 1, 24))
plt.yticks(np.arange(0, 61, 15))
ax.set_ylabel("Volume")
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.4, alpha = 0.5, axis = 'x')
plt.show()
fig.savefig("../images/NYC_time_series_volume_1321.pdf")
import matplotlib.pyplot as plt
import matplotlib.patches as patches
small_dense_tensor = dense_tensor[:, :, - pred_step :]
small_sparse_tensor = sparse_tensor[:, :, - pred_step :]
plt.style.use('classic')
plt.rcParams['font.family'] = 'Arial'
fig = plt.figure(figsize=(4.25, 1.55))
ax = fig.add_axes([0.14, 0.20, 0.84, 0.75])
plt.plot(small_dense_tensor[25, 20, :], color = "#006ea3",linewidth = 1.0, label = "Actual value")
plt.plot(tensor_hat[25, 20, :], color = "#e3120b", linewidth = 1.5, label = "Predicted value")
ax.set_xlim([0, pred_step])
ax.set_ylim([0, 120])
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.1, alpha = 0.2)
for j in range(7):
if small_sparse_tensor[25, 20, 24 * j] > 0:
someX, someY = j * 24, 0
currentAxis = plt.gca()
ax.add_patch(patches.Rectangle((someX, someY), 24, 300, alpha = 0.1, facecolor = 'green'))
plt.xticks(np.arange(0, 7 * 24 + 1, 24))
plt.yticks(np.arange(0, 121, 30))
ax.set_ylabel("Volume")
ax.grid(color = 'gray', linestyle = '-', linewidth = 0.4, alpha = 0.5, axis = 'x')
plt.show()
fig.savefig("../images/NYC_time_series_volume_2621.pdf")
```
## Evaluation on Pacific Surface Temperature Data
**Scenario setting**:
- Tensor size: $30\times 84\times 396$ (location x, location y, month)
- Test on original data
```
import numpy as np
import warnings
warnings.simplefilter('ignore')
dense_tensor = np.load('../datasets/Temperature-data-set/tensor.npy').astype(np.float32)
pos = np.where(dense_tensor[:, 0, :] > 50)
dense_tensor[pos[0], :, pos[1]] = 0
sparse_tensor = dense_tensor.copy()
import time
rank = 30
pred_step = 10 * 12
time_lags = np.array([1, 2, 3, 12, 13, 14, 2 * 12, 2 * 12 + 1, 2 * 12 + 2])
burn_iter = 1000
gibbs_iter = 200
for multi_step in [2, 4, 6]:
start = time.time()
print('Prediction time horizon (delta) = {}.'.format(multi_step))
tensor_hat = BTTF_forecast(dense_tensor, sparse_tensor, pred_step, multi_step, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
print()
```
**Scenario setting**:
- Tensor size: $30\times 84\times 396$ (location x, location y, month)
- Random missing (RM)
- 40% missing rate
```
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Temperature-data-set/tensor.npy').astype(np.float32)
pos = np.where(dense_tensor[:, 0, :] > 50)
dense_tensor[pos[0], :, pos[1]] = 0
random_tensor = np.random.rand(dense_tensor.shape[0], dense_tensor.shape[1], dense_tensor.shape[2])
missing_rate = 0.4
## Random missing (RM)
binary_tensor = np.round(random_tensor + 0.5 - missing_rate)
sparse_tensor = dense_tensor.copy()
sparse_tensor[binary_tensor == 0] = np.nan
sparse_tensor[sparse_tensor == 0] = np.nan
import time
rank = 30
pred_step = 10 * 12
time_lags = np.array([1, 2, 3, 12, 13, 14, 2 * 12, 2 * 12 + 1, 2 * 12 + 2])
burn_iter = 1000
gibbs_iter = 200
for multi_step in [2, 4, 6]:
start = time.time()
print('Prediction time horizon (delta) = {}.'.format(multi_step))
tensor_hat = BTTF_forecast(dense_tensor, sparse_tensor, pred_step, multi_step, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
print()
```
**Scenario setting**:
- Tensor size: $30\times 84\times 396$ (location x, location y, month)
- Random missing (RM)
- 60% missing rate
```
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Temperature-data-set/tensor.npy').astype(np.float32)
pos = np.where(dense_tensor[:, 0, :] > 50)
dense_tensor[pos[0], :, pos[1]] = 0
random_tensor = np.random.rand(dense_tensor.shape[0], dense_tensor.shape[1], dense_tensor.shape[2])
missing_rate = 0.6
## Random missing (RM)
binary_tensor = np.round(random_tensor + 0.5 - missing_rate)
sparse_tensor = dense_tensor.copy()
sparse_tensor[binary_tensor == 0] = np.nan
sparse_tensor[sparse_tensor == 0] = np.nan
import time
rank = 30
pred_step = 10 * 12
time_lags = np.array([1, 2, 3, 12, 13, 14, 2 * 12, 2 * 12 + 1, 2 * 12 + 2])
burn_iter = 1000
gibbs_iter = 200
for multi_step in [2, 4, 6]:
start = time.time()
print('Prediction time horizon (delta) = {}.'.format(multi_step))
tensor_hat = BTTF_forecast(dense_tensor, sparse_tensor, pred_step, multi_step, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
print()
```
**Scenario setting**:
- Tensor size: $30\times 84\times 396$ (location x, location y, month)
- Non-random missing (NM)
- 40% missing rate
```
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Temperature-data-set/tensor.npy').astype(np.float32)
pos = np.where(dense_tensor[:, 0, :] > 50)
dense_tensor[pos[0], :, pos[1]] = 0
random_tensor = np.random.rand(dense_tensor.shape[0], dense_tensor.shape[1], int(dense_tensor.shape[2] / 3))
missing_rate = 0.4
## Non-random missing (NM)
binary_tensor = np.zeros(dense_tensor.shape)
for i1 in range(dense_tensor.shape[0]):
for i2 in range(dense_tensor.shape[1]):
for i3 in range(int(dense_tensor.shape[2] / 3)):
binary_tensor[i1, i2, i3 * 3 : (i3 + 1) * 3] = np.round(random_tensor[i1, i2, i3] + 0.5 - missing_rate)
sparse_tensor = dense_tensor.copy()
sparse_tensor[binary_tensor == 0] = np.nan
sparse_tensor[sparse_tensor == 0] = np.nan
import time
rank = 30
pred_step = 10 * 12
time_lags = np.array([1, 2, 3, 12, 13, 14, 2 * 12, 2 * 12 + 1, 2 * 12 + 2])
burn_iter = 1000
gibbs_iter = 200
for multi_step in [2, 4, 6]:
start = time.time()
print('Prediction time horizon (delta) = {}.'.format(multi_step))
tensor_hat = BTTF_forecast(dense_tensor, sparse_tensor, pred_step, multi_step, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
print()
```
### License
<div class="alert alert-block alert-danger">
<b>This work is released under the MIT license.</b>
</div>
| github_jupyter |
download datasetnya https://drive.google.com/file/d/1IX9cWMwzc4v8lLivk19k2LV2JrCj0KD1/view?usp=sharing
```
import pandas as pd
import numpy as np
df = pd.read_csv('Amazon_Unlocked_Mobile.csv')
# df = df.sample(frac=0.1, random_state=10)
df.head()
df.dropna(inplace=True)
df = df[df['Rating'] != 3]
df['Positively Rated'] = np.where(df['Rating'] > 3, 1, 0)
df.head(10)
df['Positively Rated'].mean()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df['Reviews'],
df['Positively Rated'],
random_state=0)
print('X_train first entry:\n\n', X_train.iloc[0])
print('\n\nX_train shape: ', X_train.shape)
```
# CountVectorizer
```
from sklearn.feature_extraction.text import CountVectorizer
vect = CountVectorizer().fit(X_train)
vect.get_feature_names()[::2000]
len(vect.get_feature_names())
X_train_vectorized = vect.transform(X_train)
X_train_vectorized
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X_train_vectorized, y_train)
from sklearn.metrics import roc_auc_score
predictions = model.predict(vect.transform(X_test))
print('AUC: ', roc_auc_score(y_test, predictions))
feature_names = np.array(vect.get_feature_names())
sorted_coef_index = model.coef_[0].argsort()
print('Smallest Coefs:\n{}\n'.format(feature_names[sorted_coef_index[:10]]))
print('Largest Coefs: \n{}'.format(feature_names[sorted_coef_index[:-11:-1]]))
```
# TfIdf
```
from sklearn.feature_extraction.text import TfidfVectorizer
# Fit the TfidfVectorizer to the training data specifiying a minimum document frequency of 5
vect = TfidfVectorizer(min_df=5).fit(X_train)
len(vect.get_feature_names())
X_train_vectorized = vect.transform(X_train)
model = LogisticRegression()
model.fit(X_train_vectorized, y_train)
predictions = model.predict(vect.transform(X_test))
print('AUC: ', roc_auc_score(y_test, predictions))
feature_names = np.array(vect.get_feature_names())
sorted_tfidf_index = X_train_vectorized.max(0).toarray()[0].argsort()
print('Smallest tfidf:\n{}\n'.format(feature_names[sorted_tfidf_index[:10]]))
print('Largest tfidf: \n{}'.format(feature_names[sorted_tfidf_index[:-11:-1]]))
sorted_coef_index = model.coef_[0].argsort()
print('Smallest Coefs:\n{}\n'.format(feature_names[sorted_coef_index[:10]]))
print('Largest Coefs: \n{}'.format(feature_names[sorted_coef_index[:-11:-1]]))
print(model.predict(vect.transform(['not an issue, phone is working',
'an issue, phone is not working'])))
vect = CountVectorizer(min_df=5, ngram_range=(1,2)).fit(X_train)
X_train_vectorized = vect.transform(X_train)
len(vect.get_feature_names())
model = LogisticRegression()
model.fit(X_train_vectorized, y_train)
predictions = model.predict(vect.transform(X_test))
print('AUC: ', roc_auc_score(y_test, predictions))
feature_names = np.array(vect.get_feature_names())
sorted_coef_index = model.coef_[0].argsort()
print('Smallest Coefs:\n{}\n'.format(feature_names[sorted_coef_index[:10]]))
print('Largest Coefs: \n{}'.format(feature_names[sorted_coef_index[:-11:-1]]))
print(model.predict(vect.transform(['not an issue, phone is working',
'an issue, phone is not working'])))
```
| github_jupyter |
```
%load_ext nb_black
%load_ext autoreload
%autoreload 2
import os
from pathlib import Path
from requests import get
import pandas as pd
import numpy as np
from sklearn.metrics import roc_auc_score, log_loss
from sklearn.preprocessing import LabelEncoder
from tensorflow.keras.utils import plot_model
from tensorflow.keras.callbacks import EarlyStopping
import logging
logging.basicConfig(level=logging.WARN)
from xplainet.input_utils import preproc_dataset
from xplainet.model import build_model
from xplainet.random_utils import setup_seed, SEED
from sklearn.model_selection import StratifiedShuffleSplit
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
%matplotlib inline
from zipfile import ZipFile
setup_seed()
def download(url, out, force=False, verify=True):
out.parent.mkdir(parents=True, exist_ok=True)
if force and out.exists():
print(f"Removing file at {str(out)}")
out.unlink()
if out.exists():
print("File already exists.")
return
print(f"Downloading {url} at {str(out)} ...")
# open in binary mode
with out.open(mode="wb") as file:
# get request
response = get(url, verify=verify)
for chunk in response.iter_content(100000):
# write to file
file.write(chunk)
def plot_history(history):
loss_list = [s for s in history.history.keys() if "loss" in s and "val" not in s]
val_loss_list = [s for s in history.history.keys() if "loss" in s and "val" in s]
acc_list = [s for s in history.history.keys() if "AUC" in s and "val" not in s]
val_acc_list = [s for s in history.history.keys() if "AUC" in s and "val" in s]
if len(loss_list) == 0:
print("Loss is missing in history")
return
## As loss always exists
epochs = range(1, len(history.history[loss_list[0]]) + 1)
## Loss
plt.figure(1)
for l in loss_list:
plt.plot(
epochs,
history.history[l],
"b",
label="Training loss ("
+ str(str(format(history.history[l][-1], ".5f")) + ")"),
)
for l in val_loss_list:
plt.plot(
epochs,
history.history[l],
"g",
label="Validation loss ("
+ str(str(format(history.history[l][-1], ".5f")) + ")"),
)
plt.title("Loss")
plt.xlabel("Epochs")
plt.ylabel("Loss")
plt.legend()
plt.show()
```
## Census income : loading data
```
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data"
url_test = "https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.test"
dataset_name = "census-income"
out = Path(os.getcwd() + "/data/" + dataset_name + ".csv")
out_test = Path(os.getcwd() + "/data/" + dataset_name + "_test.csv")
download(url, out, force=False)
download(url_test, out_test, force=False)
cols = [
"age",
"workclass",
"fnlwgt",
"education",
"education-num",
"marital-status",
"occupation",
"relationship",
"race",
"sex",
"capital-gain",
"capital-loss",
"hours-per-week",
"native-country",
"target",
]
train = pd.read_csv(out, names=cols)
test = pd.read_csv(out_test, names=cols, skiprows=2)
target = "target"
train[target] = train[target].str.strip()
# Test has . in label, let's clean it
test[target] = test[target].str.strip().str.strip(".")
if "Set" not in train.columns:
print("Building tailored column")
train_index, valid_index = next(
StratifiedShuffleSplit(n_splits=1, test_size=0.1, random_state=SEED).split(
range(train[target].shape[0]), train[target].values
)
)
train["Set"] = "train"
train["Set"][valid_index] = "valid"
train_indices = train[train.Set == "train"].index
valid_indices = train[train.Set == "valid"].index
# test_indices = train[train.Set == "test"].index
input_train, params = preproc_dataset(train.loc[train_indices], target, ["Set"])
params
len(train_indices)
input_valid, _ = preproc_dataset(train.loc[valid_indices], target, ["Set"], params)
input_test, _ = preproc_dataset(test, target, ["Set"], params)
target_encoder = LabelEncoder()
train[target] = target_encoder.fit_transform(train[target].values.reshape(-1))
y_train = train[target].values[train_indices]
y_valid = train[target].values[valid_indices]
y_test = target_encoder.fit_transform(test[target].values)
params
model = build_model(
params,
lconv_dim=[4],
lconv_num_dim=[8],
emb_size=16,
activation_num_first_layer=None, # "tanh",
)
model.get_layer("output")._build_input_shape
model.summary()
#!pip install pydot graphviz
plot_model(
model,
# to_file="model.png",
show_shapes=True,
show_layer_names=True,
rankdir="TB",
expand_nested=False,
dpi=96,
)
y_train.shape
counts = np.unique(y_train, return_counts=True)[1]
counts = counts.sum() / counts
class_weight = {
0: counts[0],
1: counts[1],
}
class_weight
```
```
%%time
history = model.fit(
input_train,
y_train.reshape(-1, 1),
epochs=2000,
batch_size=1024,
validation_data=(input_valid, y_valid.reshape(-1, 1),),
verbose=2,
callbacks=[EarlyStopping(monitor="val_loss", patience=20, verbose=1)],
class_weight=class_weight
)
plot_history(history)
model_auc = roc_auc_score(
y_true=y_valid,
y_score=model.predict(input_valid).reshape(-1),
)
model_auc
model_auc = roc_auc_score(
y_true=y_test,
y_score=model.predict(input_test).reshape(-1),
)
model_auc
from xplainet.model import predict, encode
probs, explanations = predict(model, input_test)
probs, encoded_output = encode(model, input_test)
y_test.shape
encoded_output.shape
explanations.shape
import matplotlib.pyplot as plt
# plt.rcdefaults()
import numpy as np
import matplotlib.pyplot as plt
def explain_plot(importances, columns):
selection = np.argsort(-np.absolute(importances))[:10]
# indexes = np.argsort(importances)
performance = importances[selection]
# print(performance.shape)512, 256, 128, 64, 32, 1
y_pos = np.arange(performance.shape[0])
plt.barh(y_pos, performance, align="center", alpha=0.5)
plt.yticks(y_pos, columns[selection])
# plt.xlabel('Usage')
plt.title("Feature importance")
plt.show()
all_cols = np.array(params["bool_cols"] + params["num_cols"] + params["cat_cols"])
all_cols
```
## Explain global
```
probs_train, explanations_train = predict(model, input_train)
global_explain = np.sum(explanations_train, axis=0)
global_explain = global_explain / np.abs(global_explain).sum()
explain_plot(global_explain, all_cols)
global_explain = np.abs(explanations_train).sum(axis=0)
global_explain = global_explain / global_explain.sum()
explain_plot(global_explain, all_cols)
global_explain = np.where(explanations_train >= 0, explanations_train, 0).sum(axis=0)
global_explain = global_explain / np.abs(global_explain).sum()
explain_plot(global_explain, all_cols)
global_explain = np.where(explanations_train < 0, explanations_train, 0).sum(axis=0)
global_explain = global_explain / np.abs(global_explain).sum()
explain_plot(global_explain, all_cols)
```
## Explain local
```
for i in range(20):
explain_plot(explanations[i], all_cols)
print(probs[i].item())
from sklearn.metrics import confusion_matrix
np.unique(y_test, return_counts=True)
y_test
target_encoder.classes_
confusion_matrix(
y_true=y_test,
y_pred=model.predict(input_test).reshape(-1) >= 0.5,
# labels=target_encoder.classes_,
)
```
| github_jupyter |
# Shifting moons with dynamic rotation
```
%load_ext autoreload
%autoreload 2
import jax
import optax
import dojax
import jax.numpy as jnp
import flax.linen as nn
import pandas as pd
import matplotlib.pyplot as plt
from celluloid import Camera
from sklearn.datasets import make_moons
from sklearn.decomposition import PCA, KernelPCA
from sklearn.manifold import TSNE
%config InlineBackend.figure_format = "retina"
plt.rcParams["axes.spines.right"] = False
plt.rcParams["axes.spines.top"] = False
X, y = make_moons(n_samples=100, noise=0.12, random_state=314)
class MLP(nn.Module):
@nn.compact
def __call__(self, x):
x = nn.relu(nn.Dense(30)(x))
x = nn.relu(nn.Dense(30, name="last_layer")(x))
x = nn.Dense(1)(x)
x = nn.sigmoid(x)
return x
model = MLP()
loss = dojax.make_mse_func(model, X, y)
batch = jnp.ones((1, 2))
key = jax.random.PRNGKey(314)
params = model.init(key, batch)
alpha = 0.1
tx = optax.adam(learning_rate=alpha)
opt_state = tx.init(params)
loss_grad_fn = jax.value_and_grad(loss)
for i in range(201):
loss_val, grads = loss_grad_fn(params)
updates, opt_state = tx.update(grads, opt_state)
params = optax.apply_updates(params, updates)
if i % 50 == 0:
print('Loss step {}: '.format(i), loss_val)
```
## Multiple
```
def estimate_weights(model, X, y, key, optimizer, n_epochs=200, seed=None, output_progress=False, print_final_loss=False):
loss = dojax.make_mse_func(model, X, y)
batch = jnp.ones((1, 2))
params = model.init(key, batch)
opt_state = optimizer.init(params)
loss_grad_fn = jax.value_and_grad(loss)
for i in range(n_epochs):
loss_val, grads = loss_grad_fn(params)
updates, opt_state = tx.update(grads, opt_state)
params = optax.apply_updates(params, updates)
if i % 50 == 0 and output_progress:
print('Loss step {}: '.format(i), loss_val)
if print_final_loss:
print(f"Final loss: {loss_val}")
output = {
"params": params,
"final_loss": loss_val,
"train_accuracy": (model.apply(params, X).round().ravel() == y).mean()
}
return output
model = MLP()
alpha = 0.1
tx = optax.adam(learning_rate=alpha)
X, y = make_moons(n_samples=100, noise=0.12, random_state=314)
X = jnp.einsum("nm,mk->nk", X, dojax.rotation_matrix(0))
res = estimate_weights(model, X, y, key, tx)
params = res["params"]
params_flat, _ = jax.flatten_util.ravel_pytree(params)
pred_map = jax.vmap(model.apply, (None, 1))
pred_map = jax.vmap(pred_map, (None, 2))
X_grid = jnp.mgrid[-2:2.5:0.1, -2:2.5:0.1]
Z = jnp.einsum("ijn->ji", pred_map(params, X_grid))
plt.contourf(*X_grid, Z, cmap="bone")
plt.scatter(*X.T, c=y, cmap="Dark2")
```
## Weight's dynamics
```
def train_mlp_model(key, data_generator, model, optimiser, eval_elements,
n_epochs=200, centre=False, **kwargs):
"""
Train an MLP model iterating over eval elements and applying each element to the
data-gerating process.
Parameters
----------
data_generator: function
Data generation function. It returns a tuple of
X, y elements.
model: Flax model
The model to train the X and y elements.
optimiser: Optax element
Optimiser to train falx model
eval_elements: array
The range of values to iterate the model configuration
n_epoch: int
Number of epochs to train the model
centre: Bool
Whether to centre the data
Returns
-------
dictionary
"""
data_hist = []
params_hist = []
train_acc_hist = []
n_elements = len(eval_elements)
for it, val in enumerate(eval_elements):
X, y = data_generator(val)
X_train = X.copy()
if centre:
X_train = X_train - X_train.mean(axis=0, keepdims=True)
res = estimate_weights(model, X_train, y, key, optimiser, n_epochs=n_epochs)
params = res["params"]
loss = res["final_loss"]
train_acc = res["train_accuracy"].item()
data_hist.append([X, y])
params_hist.append(params)
train_acc_hist.append(train_acc)
print(f"@it: {it+1:03}/{n_elements:03} || {loss=:0.4e}", end="\r")
hist = {
"data": data_hist,
"params": params_hist,
"train_accuracy": jnp.array(train_acc_hist)
}
return hist
def flat_and_concat_params(params_hist):
"""
Flat and concat a list of parameters trained using
a Flax model
Parameters
----------
params_hist: list of flax FrozenDicts
List of flax FrozenDicts containing trained model
weights.
Returns
-------
jnp.array: flattened and concatenated weights
"""
flat_params = [jax.flatten_util.ravel_pytree(params)[0] for params in params_hist]
flat_params = jnp.r_[flat_params]
return flat_params
def make_rotating_moons(radians, n_samples=100, **kwargs):
"""
Make two interleaving half circles rotated by 'radians' radians
Parameters
----------
radians: float
Angle of rotation
n_samples: int
Number of samples
**kwargs:
Extra arguments passed to the `make_moons` function
"""
X, y = make_moons(n_samples=n_samples, **kwargs)
X = jnp.einsum("nm,mk->nk", X, dojax.rotation_matrix(radians))
return X, y
def make_rotating_translating_moons(radians, n_samples=100, **kwargs):
"""
Make two interleaving half circles rotated by 'radians' radians
Parameters
----------
radians: float
Angle of rotation
n_samples: int
Number of samples
**kwargs:
Extra arguments passed to the `make_moons` function
"""
X, y = make_moons(n_samples=n_samples, **kwargs)
# 1. rotate
X = jnp.einsum("nm,mk->nk", X, dojax.rotation_matrix(radians))
# 2. translate
X = X + 4 * jnp.c_[jnp.cos(radians), jnp.sin(2 * radians)]
return X, y
```
## Rotate
```
alpha = 0.05
n_steps = 200
model = MLP()
key = jax.random.PRNGKey(314)
tx = optax.adam(learning_rate=alpha)
radii = jnp.linspace(0, 2 * 2 * jnp.pi, n_steps)
hist = train_mlp_model(key, lambda rad: make_rotating_moons(rad, noise=0.2), model, tx, radii)
data_hist, params_hist = hist["data"], hist["params"]
import os
import pickle
from datetime import datetime
date_fmt = "%y%m%d%H%m"
date_str = datetime.now().strftime(date_fmt)
file_params_name = f"moons-rotating-params-{date_str}.pkl"
file_params_name = os.path.join("outputs", file_params_name)
file_dataset_name = f"moons-rotating-dataset-{date_str}.pkl"
file_dataset_name = os.path.join("outputs", file_dataset_name)
with open(file_params_name, "wb") as f:
pickle.dump(params_hist, f)
with open(file_dataset_name, "wb") as f:
pickle.dump(data_hist, f)
import pickle
file_name = "./outputs/moons-rotating-params-2203010703.pkl"
with open(file_name, "rb") as f:
params_hist = pickle.load(f)
params_flat_hist = flat_and_concat_params(params_hist)
proj = PCA(n_components=2)
# proj = KernelPCA(n_components=2, kernel="cosine")
# proj = TSNE(n_components=2, init="pca", random_state=314, perplexity=10)
w_transformed = proj.fit_transform(params_flat_hist)
plt.title("Projected weights")
domain = radii % (2 * jnp.pi)
plt.scatter(*w_transformed.T, c=domain, cmap="twilight")
plt.axis("equal");
def plot_history(params_hist, data_hist, ranges):
"""
Animate projected weight dynamics and observations
Parameters
----------
params_hist: list of FrozenDict
List of trained weights
data_hist: list of (array(N).
"""
...
fig, ax = plt.subplots(1, 2, figsize=(12, 4))
camera = Camera(fig)
xmin, xmax = -2, 2
ymin, ymax = -2, 2
X_grid = jnp.mgrid[xmin:xmax:0.1, ymin:ymax:0.1]
ax[0].set_title("Observation space")
ax[1].set_title("(projected) weight space")
for it in range(n_steps):
# plt.cla()
params = params_hist[it]
X_step, y_step = data_hist[it]
Z = jnp.einsum("ijn->ji", pred_map(params, X_grid))
ax[1].scatter(*w_transformed[:it].T, c=radii[:it] % (2 * jnp.pi), cmap="twilight")
ax[1].axis("equal")
ax[0].contourf(*X_grid, Z, cmap="bone")
ax[0].scatter(*X_step.T, c=y_step, cmap="Dark2")
ax[0].set_xlim(xmin, xmax)
ax[0].set_ylim(ymin, ymax)
camera.snap()
animation = camera.animate()
# animation.save('half-moons.gif', writer = 'imagemagick')
animation.save('half-moons.mp4', fps=20, dpi=150)
```
## Approximate projection
In this section, we investigate the effect of projecting the weights to a linear subspace and projecting them back to the full space using an approximate mapping.
In this test, we take $d$ number of components and
```
def create_comparisson_df(model, projected_weights, full_weights, configurations, projection,
test_samples=100, seed=None):
hist_values = []
n_configurations = len(configurations)
iterables = zip(configurations, projected_weights, full_weights)
for ix, (config, w_proj, w_full) in enumerate(iterables):
seed_ix = None if seed is None else seed + ix
print(f"@it{ix+1:03}/{n_configurations}", end="\r")
w_full_approx = projection.inverse_transform(w_proj)
w_full_approx = rebuild_params(w_full_approx)
X_test, y_test = make_rotating_moons(config, n_samples=test_samples, random_state=seed_ix)
accuracy_full = (model.apply(w_full, X_test).round().ravel() == y_test).mean().item()
accuracy_proj = (model.apply(w_full_approx, X_test).round().ravel() == y_test).mean().item()
entry = {
"radius": config.item(),
"acc_full": accuracy_full,
"acc_proj": accuracy_proj
}
hist_values.append(entry)
hist_values = pd.DataFrame(hist_values).set_index("radius")
return hist_values
n_components = 100
proj = PCA(n_components=n_components)
w_transformed = proj.fit_transform(params_flat_hist)
_, rebuild_params = jax.flatten_util.ravel_pytree(params_hist[0])
components = range(0, 210, 10)
errors = []
for n_components in components:
n_components = 1 if n_components == 0 else n_components
print(f"Evaluating component {n_components}")
proj = PCA(n_components=n_components)
w_transformed = proj.fit_transform(params_flat_hist)
_, rebuild_params = jax.flatten_util.ravel_pytree(params_hist[0])
hist_values = create_comparisson_df(model, w_transformed, params_hist, radii, proj)
error = hist_values.diff(axis=1).dropna(axis=1)
mean_abs_error = error.abs().mean().item()
mean_error = error.mean().item()
errors_entry = {
"n_components": n_components,
"mean_abs_error": mean_abs_error,
"mean_error": mean_error
}
errors.append(errors_entry)
print(f"{mean_abs_error=:0.4f}")
print(f"{mean_error=:0.4f}", end="\n"*2)
errors_df = pd.DataFrame(errors, index=components)
fig, ax = plt.subplots(1, 2, figsize=(12, 4))
ax[0].set_title("Mean error")
ax[1].set_title("Mean absolute error")
errors_df["mean_error"].plot(marker="o", ax=ax[0])
errors_df["mean_abs_error"].plot(marker="o", ax=ax[1])
ax[0].grid(alpha=0.5)
ax[1].grid(alpha=0.5)
plt.tight_layout()
hist_values = []
for ix, radius in enumerate(radii):
print(f"@it{ix+1:03}/{len(radii)}", end="\r")
deg = (180 * (radii[ix] % (2 * jnp.pi))).item()
w_approx = proj.inverse_transform(w_transformed[ix])
w_approx = rebuild_params(w_approx)
w_full = params_hist[ix]
X_test, y_test = make_rotating_moons(radii[ix], n_samples=100)
accuracy_proj = (model.apply(w_approx, X_test).round().ravel() == y_test).mean().item()
accuracy_full = (model.apply(w_full, X_test).round().ravel() == y_test).mean().item()
entry = {
"radius": radius.item(),
"accuracy_full": accuracy_full,
"accuracy_approx": accuracy_proj,
}
hist_values.append(entry)
df_values = pd.DataFrame(hist_values).set_index("radius")
df_values = pd.DataFrame(hist_values).set_index("radius")
df_values.diff(axis=1).iloc[:, 1].plot()
plt.axhline(y=0, c="tab:gray", linestyle="--")
plt.xticks(rotation=45);
fig, ax = plt.subplots()
pd.DataFrame(hist_values).plot(x="radius", y="accuracy_approx", ax=ax)
pd.DataFrame(hist_values).plot(x="radius", y="accuracy_full", ax=ax)
plt.xticks(rotation=45);
```
## Rotate and shift
```
for r in jnp.linspace(0, 2 * jnp.pi, 100):
x = 5 * jnp.cos(r)
y = 5 * jnp.sin(2 * r)
plt.scatter(x, y, c="tab:blue")
plt.axis("equal");
alpha = 0.01
n_steps = 200
model = MLP()
key = jax.random.PRNGKey(314)
tx = optax.adam(learning_rate=alpha)
radii = jnp.linspace(0, 2 * 2 * jnp.pi, n_steps)
def rotate_shift_centre():
"""
"""
...
hist = train_mlp_model(key, lambda rad: make_rotating_translating_moons(rad, noise=0.1),
model, tx, radii, centre=True)
data_hist, params_hist = hist["data"], hist["params"]
params_flat_hist = flat_and_concat_params(params_hist)
radii_mod = radii % (2 * jnp.pi)
plt.scatter(radii_mod, hist["train_accuracy"])
proj = PCA(n_components=2)
# proj = KernelPCA(n_components=2, kernel="cosine")
proj = TSNE(n_components=2, init="pca", random_state=314,
perplexity=20, learning_rate="auto")
w_transformed = proj.fit_transform(params_flat_hist)
plt.title("Projected weights")
plt.scatter(*w_transformed.T, c=radii_mod, cmap="twilight")
plt.axis("equal")
plt.axis("off")
```
## Higher dimensions
Quickly adapting model. When corrupting, thing always on a way to uncorrupt.
* We can go from model A to model B in different ways (or paths). Think of driving in a car during the day and eventually going into night v.s.
* How do we move to find a new datapoint? What triggers the switch to go find another model? Kevin suggested doing marginal likelihood test.
* We seek a highly adaptive model that is able to revisit a state if it needs to (remember)
* Distance in weight space.
* We can track distribution shift through the weights of NNet if the transformation is smooth.
* Neural generative models. Take VAE => Apply weights to the VAE.
* Conjectures: Neural architectures preserve the topology of the weight-space dynamics.
* Q: how to make the predictions? / how to choose the learned weights?
## ToDo
1. Dimensionality
2. Stochasticity
3. Abrupt changes
# * Increase dimensionality of points (20D/40D)
* Nonlinear transformation of points
1. Try transformation over MNIST dataset
2. Try _topological_ transformation of the moons to two circles. (Try transformation with discretised ODEs)
3. Abrupt change of relabeling of points.
| github_jupyter |
# Practice: Question Answering with a Fine-Tuned BERT (and TTS example)
This notebook is based on great [post and corresponding notebook](https://mccormickml.com/2020/03/10/question-answering-with-a-fine-tuned-BERT/) *by Chris McCormick*. It contains some minor changes and additions (especially parts 3 and 4).
What does it mean for BERT to achieve "human-level performance on Question Answering"? Is BERT the greatest search engine ever, able to find the answer to any question we pose it?
In **Part 1** of this notebook, we will discuss what it really means to apply BERT to QA, and illustrate the details.
**Part 2** contains example code--we'll be downloading a model that's *already been fine-tuned* for question answering, and try it out on our own text!
In **Part 3** we will apply the same approach to Russian language using the model pre-trained on SberQuAD dataset.
And in **Part 4** and **Part 5** we will generate question and answer as audio in english and russian languages.
**Links**
* The [video walkthrough](https://youtu.be/l8ZYCvgGu0o) on this topic.
* The [original blog post](https://mccormickml.com/2020/03/10/question-answering-with-a-fine-tuned-BERT/) version.
* The [original Colab Notebook](https://colab.research.google.com/drive/1uSlWtJdZmLrI3FCNIlUHFxwAJiSu2J0-).
```
!pip install -U transformers deeppavlov unidecode omegaconf
# This cell is optional and needed only for Russian language inference
# !python -m deeppavlov install squad_ru_rubert
# # Pre-downloading the BERT for Russian language. Same result can be achieved with
# # `!python -m deeppavlov download squad_ru_rubert`
# # But it works significantly slower.
# !wget -nc https://www.dropbox.com/s/7za1o6vaffbdlcg/rubert_cased_L-12_H-768_A-12_v1.tar.gz
# !mkdir -p /root/.deeppavlov/downloads/bert_models/
# !tar -xzvf rubert_cased_L-12_H-768_A-12_v1.tar.gz -C /root/.deeppavlov/downloads/bert_models
# !wget -nc https://www.dropbox.com/s/ns8280pd9t9n9dc/squad_model_ru_rubert.tar.gz
# !mkdir -p /root/.deeppavlov/models/
# !tar -xzvf squad_model_ru_rubert.tar.gz -C /root/.deeppavlov/models
import torch
assert torch.cuda.is_available(), 'Tacotron2 by NVIDIA infers only on GPU, so the Part 4 will not work on CPU-only machine'
device = torch.device('cuda:0')
tacotron2 = torch.hub.load('nvidia/DeepLearningExamples:torchhub', 'nvidia_tacotron2', **{'map_location': device})
tacotron2.to(device)
tacotron2.eval()
waveglow = torch.hub.load('nvidia/DeepLearningExamples:torchhub', 'nvidia_waveglow')
waveglow = waveglow.remove_weightnorm(waveglow)
waveglow.to(device)
waveglow.eval();
```
## Part 1: Applying BERT to Question Answering
### The SQuAD v1.1 Benchmark
When someone mentions "Question Answering" as an application of BERT, what they are really referring to is applying BERT to the Stanford Question Answering Dataset (SQuAD).
The task posed by the SQuAD benchmark is a little different than you might think. Given a question, and *a passage of text containing the answer* (often refered to as context), BERT needs to highlight the "span" of text corresponding to the correct answer.
The SQuAD homepage has a fantastic tool for exploring the questions and reference text for this dataset, and even shows the predictions made by top-performing models.
For example, here are some [interesting examples](https://rajpurkar.github.io/SQuAD-explorer/explore/1.1/dev/Super_Bowl_50.html?model=r-net+%20(ensemble)%20(Microsoft%20Research%20Asia)&version=1.1) on the topic of Super Bowl 50.
### BERT Input Format
To feed a QA task into BERT, we pack both the question and the reference text into the input.
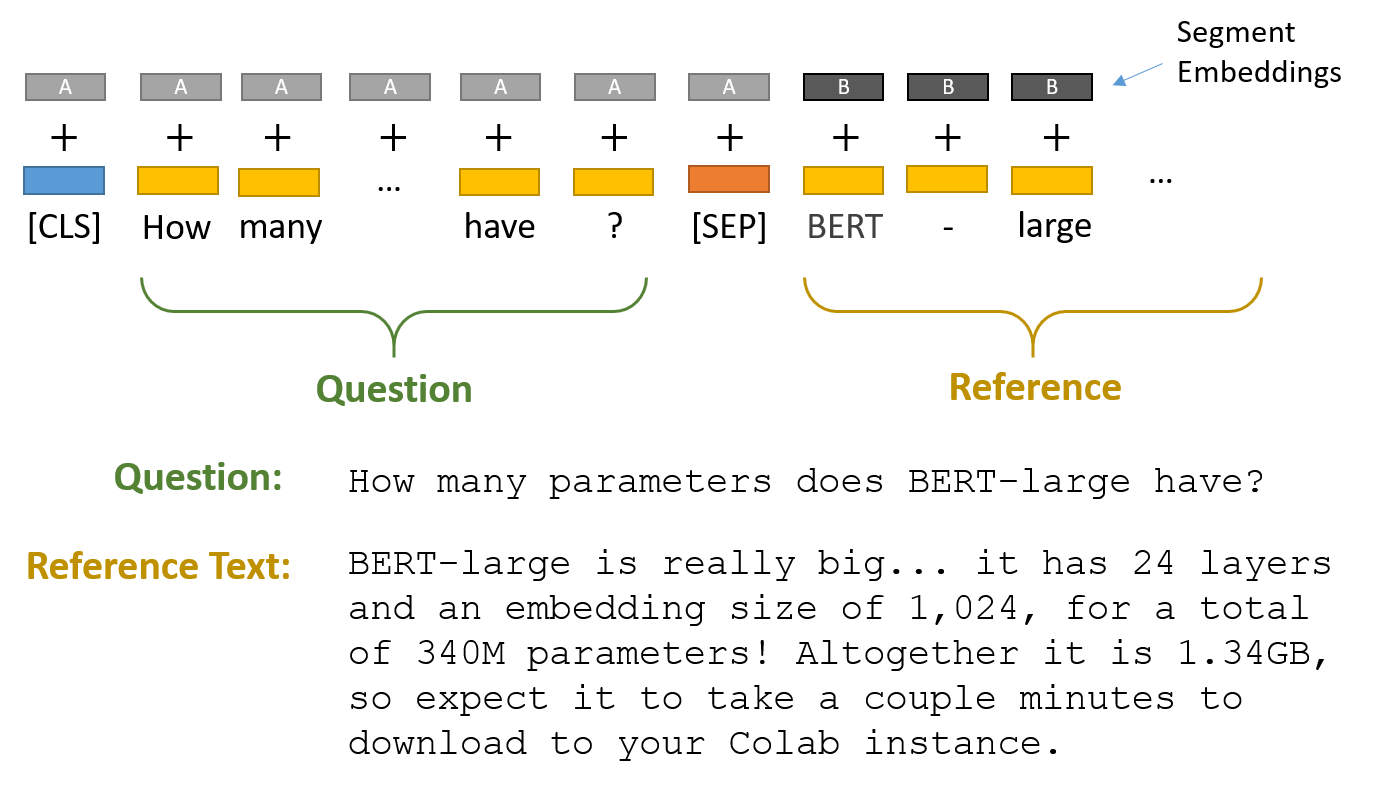
*Image credits: [Chris McCormick](https://mccormickml.com/2020/03/10/question-answering-with-a-fine-tuned-BERT/)*
The two pieces of text are separated by the special `[SEP]` token.
> _Side note:_ Original BERT also uses "Segment Embeddings" to differentiate the question from the reference text. These are simply two embeddings (for segments "A" and "B") that BERT learned, and which it adds to the token embeddings before feeding them into the input layer. However today we will be using DistilBERT model, which relies solely on the special tokens.
### Start & End Token Classifiers
BERT needs to highlight a "span" of text containing the answer--this is represented as simply predicting which token marks the start of the answer, and which token marks the end.
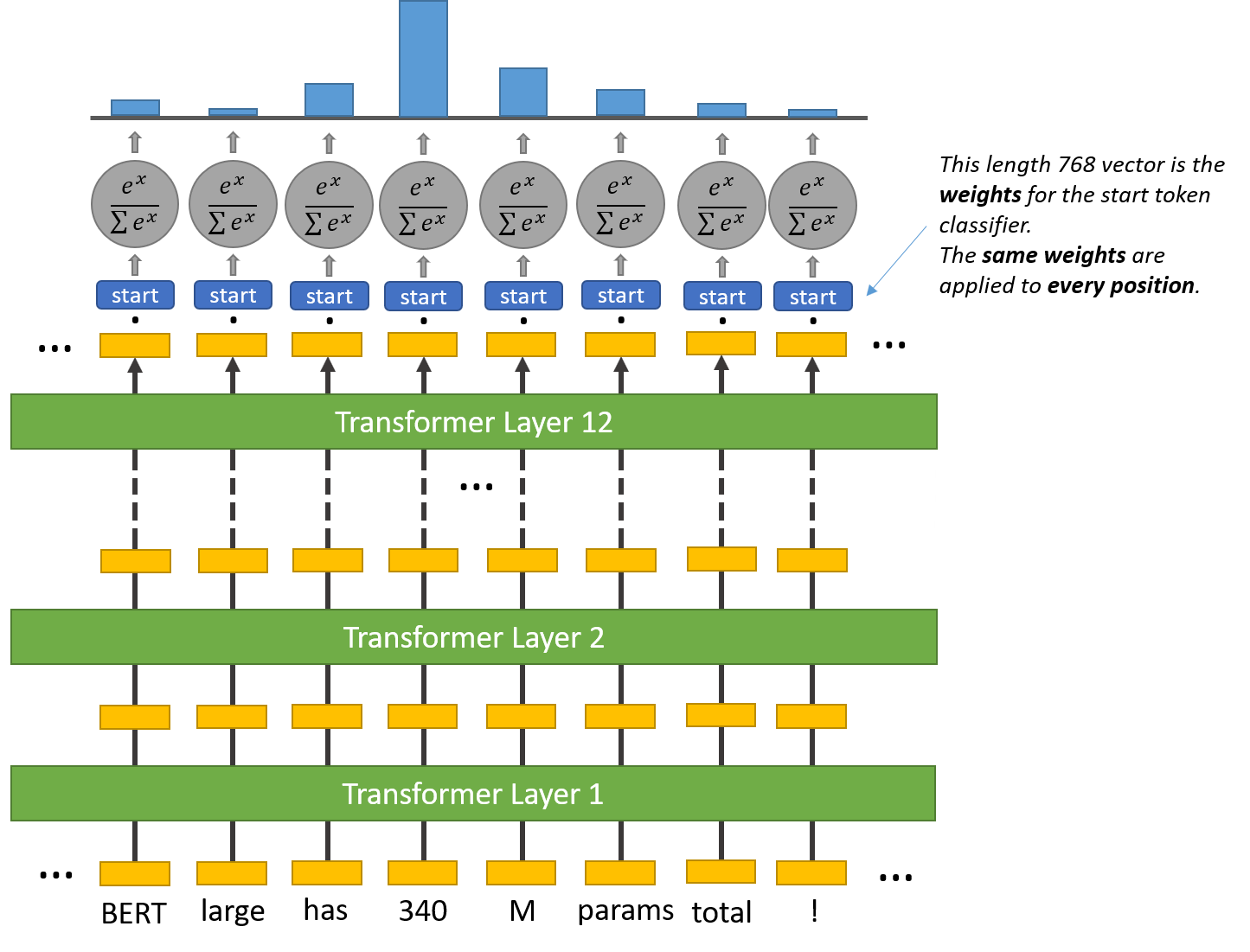
*Image credits: [Chris McCormick](https://mccormickml.com/2020/03/10/question-answering-with-a-fine-tuned-BERT/)*
For every token in the text, we feed its final embedding into the start token classifier. The start token classifier only has a single set of weights (represented by the blue "start" rectangle in the above illustration) which it applies to every word.
After taking the dot product between the output embeddings and the 'start' weights, we apply the softmax activation to produce a probability distribution over all of the words. Whichever word has the highest probability of being the start token is the one that we pick.
We repeat this process for the end token--we have a separate weight vector this.
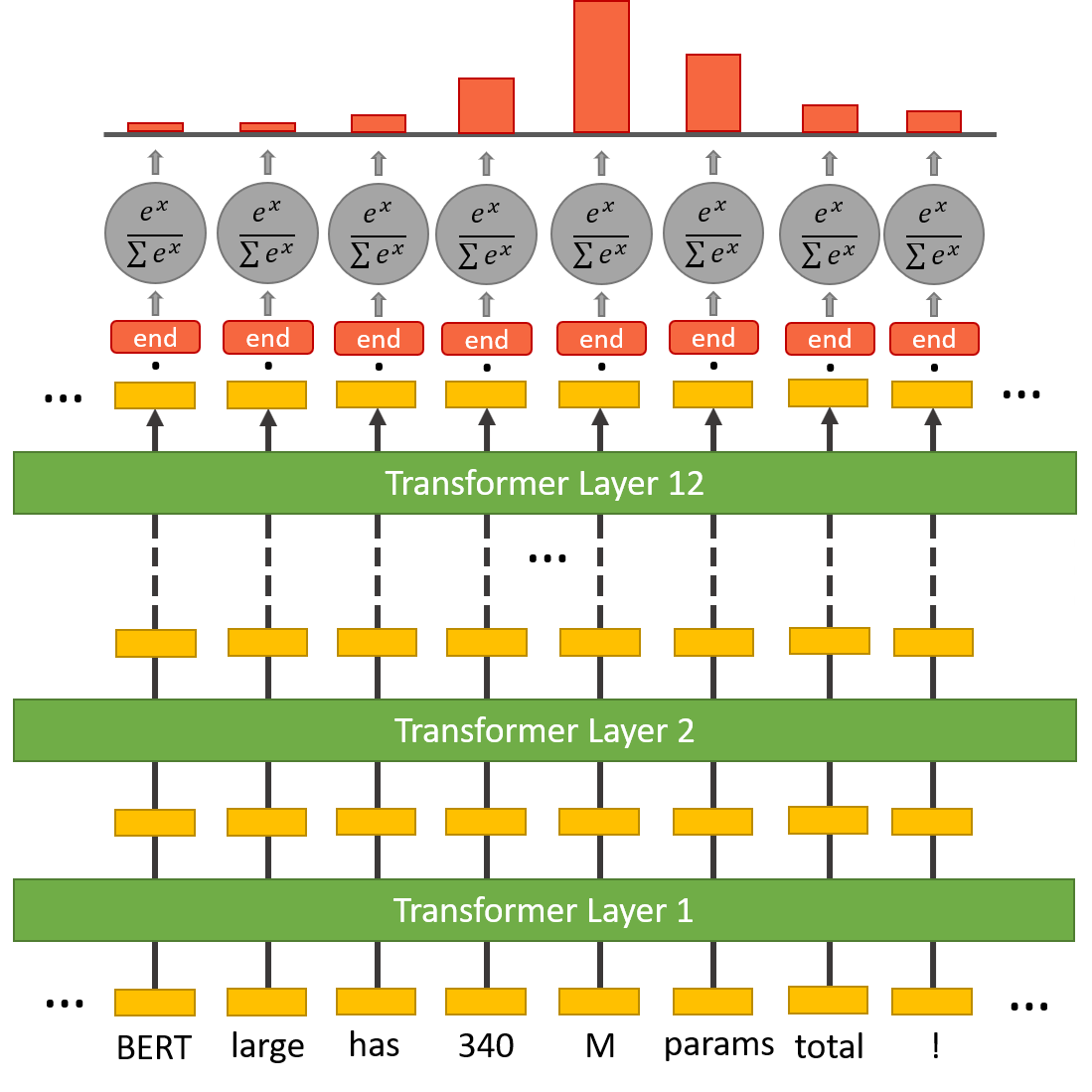
*Image credits: [Chris McCormick](https://mccormickml.com/2020/03/10/question-answering-with-a-fine-tuned-BERT/)*
## Part 2: Example Code
In the example code below, we'll be downloading a model that's *already been fine-tuned* for question answering, and try it out on our own text.
If you do want to fine-tune on your own dataset, it is possible to fine-tune BERT for question answering yourself. See [run_squad.py](https://github.com/huggingface/transformers/blob/master/examples/run_squad.py) in the `transformers` library. However, you may find that the "fine-tuned-on-squad" model already does a good job, even if your text is from a different domain.
### 1. Load Fine-Tuned BERT
This example uses the `transformers` [library](https://github.com/huggingface/transformers/) by huggingface. We've already installed it in the top of this notebook.
For Question Answering we use the `DistilBertForQuestionAnswering` class from the `transformers` library.
This class supports fine-tuning, but for this example we will keep things simpler and load a BERT model that has already been fine-tuned for the SQuAD benchmark.
The `transformers` library has a large collection of pre-trained models which you can reference by name and load easily. The full list is in their documentation [here](https://huggingface.co/transformers/pretrained_models.html).
```
from transformers import DistilBertTokenizer, DistilBertForQuestionAnswering
tokenizer = DistilBertTokenizer.from_pretrained('distilbert-base-uncased-distilled-squad')
model = DistilBertForQuestionAnswering.from_pretrained('distilbert-base-uncased-distilled-squad')
```
> _Side note:_ Apparently the vocabulary of this model is identicaly to the one in bert-base-uncased. You can load the tokenizer from `bert-base-uncased` and that works just as well.
### 2. Ask a Question
Now we're ready to feed in an example!
A QA example consists of a question and a passage of text containing the answer to that question.
```
question = "How many parameters does BERT-large have?"
context = (
"BERT-large is really big... it has 24-layers and an embedding size of 1,024, "
"for a total of 340M parameters! Altogether it is 1.34GB, so expect it to "
"take a couple minutes to download to your Colab instance."
)
```
We'll need to run the BERT tokenizer against both the `question` and the `context`. To feed these into BERT, we actually concatenate them together and place the special `[SEP]` token in between.
```
# Apply the tokenizer to the input text, treating them as a text-pair.
input_ids = tokenizer.encode(question, context)
print(f'The input has a total of {len(input_ids)} tokens.')
```
Just to see exactly what the tokenizer is doing, let's print out the tokens with their IDs.
```
# BERT only needs the token IDs, but for the purpose of inspecting the
# tokenizer's behavior, let's also get the token strings and display them.
tokens = tokenizer.convert_ids_to_tokens(input_ids)
# Display tokens and ids as table.
# For each token and its id...
for token, token_id in zip(tokens, input_ids):
# If this is the [SEP] token, add some space around it to make it stand out.
if token_id == tokenizer.sep_token_id:
print()
# Print the token string and its ID in two columns.
print('{:<12} {:>6,}'.format(token, token_id))
if token_id == tokenizer.sep_token_id:
print()
```
We're ready to feed our example into the model!
```
import torch
inputs = tokenizer(question, context, return_tensors='pt')
with torch.no_grad():
outputs = model(**inputs)
start_scores = outputs.start_logits
end_scores = outputs.end_logits
start_scores
```
>*Side Note: Where's the padding?*
>
> The original [example code](https://huggingface.co/transformers/model_doc/bert.html?highlight=bertforquestionanswering#transformers.BertForQuestionAnswering) does not perform any padding. I suspect that this is because we are only feeding in a *single example*. If we instead fed in a batch of examples, then we would need to pad or truncate all of the samples in the batch to a single length, and supply an attention mask to tell BERT to ignore the padding tokens.
Now we can highlight the answer just by looking at the most probable start and end words.
```
# Find the tokens with the highest `start` and `end` scores.
answer_start = torch.argmax(start_scores)
answer_end = torch.argmax(end_scores)
# Combine the tokens in the answer and print it out.
answer = ' '.join(tokens[answer_start : answer_end + 1])
print(f'Answer: "{answer}"')
```
It got it right! Awesome :)
> *Side Note: It's a little naive to pick the highest scores for start and end--what if it predicts an end word that's before the start word?! The correct implementation is to pick the highest total score for which end >= start.*
With a little more effort, we can reconstruct any words that got broken down into subwords.
```
answer = tokenizer.convert_tokens_to_string(tokens[answer_start : answer_end + 1])
print(f'Answer: "{answer}"')
```
### 3. Visualizing Scores
Let's see what the scores were for all of the words. The following cells generate bar plots showing the start and end scores for every word in the input.
```
import matplotlib.pyplot as plt
import seaborn as sns
# Use plot styling from seaborn.
sns.set(style='darkgrid')
# Increase the plot size and font size.
plt.rcParams['figure.figsize'] = (16, 8)
plt.rcParams['font.size'] = 16
```
Retrieve all of the start and end scores, and use all of the tokens as x-axis labels.
```
# Pull the scores out of PyTorch Tensors and convert them to 1D numpy arrays.
start_scores = start_scores.numpy().flatten()
end_scores = end_scores.numpy().flatten()
# We'll use the tokens as the x-axis labels. In order to do that, they all need
# to be unique, so we'll add the token index to the end of each one.
token_labels = []
for (i, token) in enumerate(tokens):
token_labels.append('{:} - {:>2}'.format(token, i))
```
Create a bar plot showing the score for every input word being the "start" word.
```
# Create a barplot showing the start word score for all of the tokens.
ax = sns.barplot(x=token_labels, y=start_scores, ci=None)
# Turn the xlabels vertical.
ax.set_xticklabels(ax.get_xticklabels(), rotation=90, ha="center")
# Turn on the vertical grid to help align words to scores.
ax.grid(True)
plt.title('Start Word Scores');
```
Create a second bar plot showing the score for every input word being the "end" word.
```
# Create a barplot showing the end word score for all of the tokens.
ax = sns.barplot(x=token_labels, y=end_scores, ci=None)
# Turn the xlabels vertical.
ax.set_xticklabels(ax.get_xticklabels(), rotation=90, ha="center")
# Turn on the vertical grid to help align words to scores.
ax.grid(True)
plt.title('End Word Scores');
```
**Alternate View**
I also tried visualizing both the start and end scores on a single bar plot, but I think it may actually be more confusing then seeing them separately.
```
import pandas as pd
# Store the tokens and scores in a DataFrame.
# Each token will have two rows, one for its start score and one for its end
# score. The "marker" column will differentiate them. A little wacky, I know.
scores = []
for (i, token_label) in enumerate(token_labels):
# Add the token's start score as one row.
scores.append({'token_label': token_label,
'score': start_scores[i],
'marker': 'start'})
# Add the token's end score as another row.
scores.append({'token_label': token_label,
'score': end_scores[i],
'marker': 'end'})
df = pd.DataFrame(scores)
# Draw a grouped barplot to show start and end scores for each word.
# The "hue" parameter is where we tell it which datapoints belong to which
# of the two series.
plot = sns.catplot(
x="token_label", y="score", hue="marker",
data=df, kind="bar", height=6, aspect=4
)
# Turn the xlabels vertical.
plot.set_xticklabels(plot.ax.get_xticklabels(), rotation=90, ha="center")
# Turn on the vertical grid to help align words to scores.
plot.ax.grid(True);
```
### 4. More Examples
Turn the QA process into a function so we can easily try out other examples.
```
def answer_question(question, context):
# ======== Tokenize ========
# Apply the tokenizer to the input text, treating them as a text-pair.
inputs = tokenizer(question, context, return_tensors='pt')
input_ids = inputs.input_ids.numpy().flatten()
# ======== Evaluate ========
# Run our example question through the model.
outputs = model(**inputs)
start_scores = outputs.start_logits
end_scores = outputs.end_logits
# ======== Reconstruct Answer ========
# Find the tokens with the highest `start` and `end` scores.
answer_start = torch.argmax(start_scores)
answer_end = torch.argmax(end_scores)
# Get the string versions of the input tokens.
token_ids = input_ids[answer_start : answer_end + 1]
tokens = tokenizer.convert_ids_to_tokens(token_ids)
answer = tokenizer.convert_tokens_to_string(tokens)
return answer
```
As our reference text, we've taken the Abstract of the [BERT paper](https://arxiv.org/pdf/1810.04805.pdf).
```
bert_abstract = (
'We introduce a new language representation model called BERT, which stands for '
'Bidirectional Encoder Representations from Transformers. Unlike recent language '
'representation models (Peters et al., 2018a; Radford et al., 2018), BERT is '
'designed to pretrain deep bidirectional representations from unlabeled text by '
'jointly conditioning on both left and right context in all layers. As a result, '
'the pre-trained BERT model can be finetuned with just one additional output '
'layer to create state-of-the-art models for a wide range of tasks, such as '
'question answering and language inference, without substantial taskspecific '
'architecture modifications. BERT is conceptually simple and empirically '
'powerful. It obtains new state-of-the-art results on eleven natural language '
'processing tasks, including pushing the GLUE score to 80.5% (7.7% point absolute '
'improvement), MultiNLI accuracy to 86.7% (4.6% absolute improvement), SQuAD v1.1 '
'question answering Test F1 to 93.2 (1.5 point absolute improvement) and SQuAD '
'v2.0 Test F1 to 83.1 (5.1 point absolute improvement).'
)
```
Let's ask BERT what its name stands for (the answer is in the first sentence of the abstract).
```
question = "What does the 'B' in BERT stand for?"
answer = answer_question(question, bert_abstract)
print(f'Answer: "{answer}"')
```
Let's ask BERT about example applications of itself :)
The answer to the question comes from this passage from the abstract:
> "...BERT model can be finetuned with just one additional output
layer to create state-of-the-art models for **a wide range of tasks, such as
question answering and language inference,** without substantial taskspecific
architecture modifications."
```
question = "What are some example applications of BERT?"
answer = answer_question(question, bert_abstract)
print(f'Answer: "{answer}"')
```
## [Optional] Part 3. RuBERT for question answering.
Here we will use the model pre-trained on the SberQuAD dataset from the [SDSJ-2017 challenge problem B](https://github.com/sberbank-ai/data-science-journey-2017/tree/master/problem_B).
```
from deeppavlov import build_model, configs
model_ru = build_model(configs.squad.squad_ru_rubert, download=False)
```
The following text is copied from [habr post on Crew Dragon flight](https://habr.com/ru/news/t/504642/).
```
context = (
'Первая многоразовая ступень ракеты-носителя Falcon 9 успешно отделилась через две с половиной '
'минуты после старта и автоматически приземлилась на плавучую платформу Of Course I Still '
'Love You у берегов Флориды. Через 12 минут после запуска космический корабль Crew Dragon '
'вышел на расчетную орбиту и отделился от второй ступени ракеты.'
'\n\n'
'Сближение корабля Crew Dragon с Международной космической станцией запланировано на 31 мая. '
'К стыковочному адаптеру на узловом модуле «Гармония» американского сегмента МКС Crew Dragon '
'должен причалить в ручном или, при необходимости, в автоматическом режиме. Эта процедура '
'запланирована на 10:29 по времени Восточного побережья США (17:29 по московскому времени).'
'\n\n'
'В испытательном полете DM2 астронавт Херли является командиром космического корабля (spacecraft '
'commander), а его напарник Бенкен — командир по операциям стыковки и расстыковки (joint '
'operations commander). Фактически это означает, что именно Херли управляет Crew Dragon в '
'полете к МКС, к которой они должны пристыковаться в течение суток после старта. Херли и Бенкен '
'также будут выполнять необходимые для сертификации НАСА проверки систем корабля в полете.'
'\n\n'
'Во время полета Херли и Бенкен провели небольшую экскурсию по Crew Dragon.'
)
```
And here is how to use deeppavlov's model:
```
question = 'Когда отделилась первая ступень?'
model_ru([context], [question])
```
The model returns list with answer, answer starting position in context and the answer logit.
This yields the following `answer_question` function.
```
def answer_question_ru(question, context):
output = model_ru([context], [question])
return output[0][0]
```
Let's ask a bunch of other questions to the model.
```
question = 'На какую дату запланирована стыковка?'
answer = answer_question_ru(question, context)
print(f'Ответ: "{answer}"')
question = 'Кто участвует в полете?'
answer = answer_question_ru(question, context)
print(f'Ответ: "{answer}"')
question = 'Кто участвует в полете кроме астронавта Херли?'
answer = answer_question_ru(question, context)
print(f'Ответ: "{answer}"')
question = 'Какие астронавты участвовали в полете?'
answer = answer_question_ru(question, context)
# Notice how model finds the appropriate answer dispite slightly different context.
print(f'Ответ: "{answer}"')
question = 'Какая ступень приземлилась на плавучую платформу Of Course I Still Love You?'
answer = answer_question_ru(question, context)
print(f'Ответ: "{answer}"')
```
## Part 4. Question answering with speech using Tacotron 2.
### Text to speech using Tacotron 2.
Tacotron 2 is a network proposed in 2017 in [Natural TTS Synthesis By Conditioning
Wavenet On Mel Spectrogram Predictions](https://arxiv.org/pdf/1712.05884.pdf) paper. This network takes an input text and maps it into the mel-frequency spectrogram. This spectrogram is then passed through a modified WaveNet (generative model for audio, original paper can be found [here](https://arxiv.org/pdf/1609.03499.pdf)) to generate the actual speech.
Let's look more closely at a mel spectrogram (for more info on its nature please refer to the [Tacotron 2 paper](https://arxiv.org/pdf/1712.05884.pdf)).
```
assert tacotron2 is not None and waveglow is not None, 'Tacotron2 by NVIDIA infers only on GPU, so the Part 4 will not work on CPU-only machine'
utils = torch.hub.load('NVIDIA/DeepLearningExamples:torchhub', 'nvidia_tts_utils')
text = 'Some test text.'
sequences, lengths = utils.prepare_input_sequence([text])
with torch.no_grad():
mel, _, _ = tacotron2.infer(sequences, lengths)
sns.reset_orig()
plt.imshow(mel[0].cpu().numpy())
plt.title('mel-frequency spectrogram');
```
After obtaining this spectrogram, we can generate the audio with `waveglow` model.
```
from IPython.display import Audio
sampling_rate = 22050
with torch.no_grad():
audio = waveglow.infer(mel)
audio_numpy = audio[0].cpu().numpy()
Audio(audio_numpy, rate=sampling_rate)
```
We've generated a `.wav` format audio. We can save it using the `scipy.io.wavfile.write`.
```
from scipy.io.wavfile import write
write('audio.wav', sampling_rate, audio_numpy)
```
This yields the following `text_to_speech` function.
```
def text_to_speech(text):
# preprocessing
sequences, lengths = utils.prepare_input_sequence([text])
# run the models
with torch.no_grad():
mel, _, _ = tacotron2.infer(sequences, lengths)
audio = waveglow.infer(mel)
audio_numpy = audio[0].cpu().numpy()
return audio_numpy
text = 'Another test text.'
audio_numpy = text_to_speech(text)
Audio(audio_numpy, rate=sampling_rate)
```
### Tying text to speech with question answering.
Let's take a look at [Mail.ru group blog post on Computer Vision on habr.com](https://habr.com/ru/company/mailru/blog/467905/)
```
context = (
'One of Mail.ru Cloud’s objectives is to provide the handiest means for accessing '
'and searching your own photo and video archives. For this purpose, we at Mail.ru '
'Computer Vision Team have created and implemented systems for smart image '
'processing: search by object, by scene, by face, etc. Another spectacular '
'technology is landmark recognition. Today, I am going to tell you how we made '
'this a reality using Deep Learning.'
'\n\n'
'Imagine the situation: you return from your vacation with a load of photos. Talking '
'to your friends, you are asked to show a picture of a place worth seeing, like '
'palace, castle, pyramid, temple, lake, waterfall, mountain, and so on. You rush to '
'scroll your gallery folder trying to find one that is really good. Most likely, it '
'is lost amongst hundreds of images, and you say you will show it later.'
'\n\n'
'We solve this problem by grouping user photos in albums. This will let you find '
'pictures you need just in few clicks. Now we have albums compiled by face, by '
'object and by scene, and also by landmark.'
'\n\n'
'Photos with landmarks are essential because they often capture highlights of our '
'lives (journeys, for example). These can be pictures with some architecture or '
'wilderness in the background. This is why we seek to locate such images and make '
'them readily available to users.'
)
question = 'Why photos with landmarks are essential?'
answer = answer_question(question, context)
print(f'Answer: "{answer}"')
```
Let's cat question and answer into one phrase and convert it to audio!
```
text = f'{question}\n{answer}'
audio_numpy = text_to_speech(text)
Audio(audio_numpy, rate=sampling_rate)
```
And another one.
```
question = "Which places except mountain are worth seeing?"
answer = answer_question(question, context)
print(f'Answer: "{answer}"')
text = f'{question}\n{answer}'
audio_numpy = text_to_speech(text)
Audio(audio_numpy, rate=sampling_rate)
# Take your time, experiment with questions and the generated audio
```
## [Optional] 5. Russian langugage speech generation
Of course, text to speech is not specific to english language. Here is how you can do it with russian.
```
from omegaconf import OmegaConf
torch.hub.download_url_to_file(
'https://raw.githubusercontent.com/snakers4/silero-models/master/models.yml',
'latest_silero_models.yml',
progress=False
)
models = OmegaConf.load('latest_silero_models.yml')
# see latest avaiable models
available_languages = list(models['tts_models'].keys())
print(f'Available languages {available_languages}')
for lang in available_languages:
speakers = list(models['tts_models'][lang].keys())
print(f'Available speakers for {lang}: {speakers}')
```
Let's choose our language and speaker and try using them!
```
language = 'ru'
speaker = 'kseniya_16khz'
device = torch.device('cpu')
model, symbols, sample_rate, example_text, apply_tts = torch.hub.load(
'snakers4/silero-models', 'silero_tts',
language=language, speaker=speaker
)
model = model.to(device)
audio = apply_tts(
texts=[example_text],
model=model,
sample_rate=sample_rate,
symbols=symbols,
device=device
)
print(example_text)
Audio(audio[0], rate=sample_rate)
audio = apply_tts(
texts=["Дерзайте знать! Спасибо за внимание!"],
model=model,
sample_rate=sample_rate,
symbols=symbols,
device=device
)
Audio(audio[0], rate=sample_rate)
```
| github_jupyter |
# Notable Telescope Apertures
This notebook will show how to use prysm to paint the apertures of notable telescopes. Further modeling of these observatories will not be given here, and requries additional data (e.g., OPD maps or coefficients, masks) not widely available. It is assumed the user sufficiently understands the components used to not require explanation of details. All parameters are based on publically shown values and may be imprecise. If you are a member of the science or engineering team for these systems, you should check all parameters against internal values.
Most apertures include the steps to repeat this synthesis for any similar aperture, and do not jump directly to the solution. They all conclude with a mask and a figure showing the fully composited aperture.
Links jump to telescopes:
- [HST](#HST)
- [JWST](#JWST)
- [TMT](#TMT)
- [LUVOIR-A](#LUVOIR-A)
- [LUVOIR-B](#LUVOIR-B)
- [HabEx-A](#HabEx-A)
- [HabEx-B](#HabEx-B)
```
import numpy as np
from prysm.coordinates import make_xy_grid, cart_to_polar
from prysm.geometry import spider, circle, offset_circle
from prysm.segmented import CompositeHexagonalAperture
from matplotlib import pyplot as plt
```
## HST
HST has a primary mirror of diameter 2.4 m with 32% linear obscuration, and four spiders of 38 mm diameter rotated 45$^\circ$ from the cardinal axes. There are an additional three small circular obscurations from pads used to secure the primary mirror. The pads are 90% of the way to the edge of the mirror at ccw angles w.r.t. the x axis of -45, -165, and +75 degrees and have each a diameter of 150 mm.
```
x, y = make_xy_grid(512, diameter=2.4)
r, t = cart_to_polar(x, y)
pm_od = circle(2.4/2, r)
pm_id = circle(2.4/2*.32, r)
mask = pm_od ^ pm_id # or pm_od & ~pm_id
plt.imshow(mask, cmap='gray')
```
After shading the primary, we now compute the spider and pad obscurations:
```
spider_ = spider(4, 0.038, x, y, 45)
pads_r = 0.90*2.4/2
pad_angles = [np.radians(a) for a in [-45, -165, 75]]
pad_centers = [(pads_r*np.cos(a), pads_r*np.sin(a)) for a in pad_angles]
pads = [offset_circle(.075, x, y, c) for c in pad_centers]
# pads before this point is a list of the points INSIDE each circle.
# logical or, |, below produces a mask of "pixels inside ANY circle"
# these are an obscuration, so we invert it with ~
pads = (pads[0]|pads[1]|pads[2])
hst_pupil = mask & spider_ & ~pads
plt.imshow(hst_pupil, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
plt.title('Fully composited HST aperture')
```
## JWST
<!-- This notebook will show how to use prysm to model the JWST. The principle features of the JWST shown here are the shape of its pupil as well as both per-segment and aperture-wide wavefront errors. Obviously JWST is a high complexity observatory, and each of its instruments are also complicated. We will begin from the front of the observatory and not cover any radiometric or polychromatic topics in detail, users can compose these topics with what is presented here. -->
<!-- To avoid running afoul of any export restrictions, we will also use only publicly available, unlimited release values in constructing this model. This means that if you actually want to JWST with a program inspired by what is here, you need to check and likely replace all values with the unpublicized versions. -->
JWST is a 2-ring segmented hexagonal design. The central segment is missing, and there is a upside-down "Y" strut system to hold the secondary. The segments are 1.32 m flat-to-flat, with 7 mm airgaps between. We first paint the hexagons:
```
x, y = make_xy_grid(512, diameter=6.6)
cha = CompositeHexagonalAperture(x,y,2,1.32,0.007,exclude=(0,))
plt.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
```
And create the secondary struts, adding them to the mask:
```
m1 = spider(1, .1, x, y, rotation=-120)
m2 = spider(1, .1, x, y, rotation=-60)
m3 = spider(1, .1, x, y, rotation=90)
spider_ = m1&m2&m3
plt.imshow(cha.amp&spider_, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
plt.title('Fully composited JWST aperture')
```
## TMT
TMT is a hexagonally tiled aperture with 1.44 m segments (diameter, not flat-to-flat) and only 2.5 mm gaps. The gaps cannot be drawn properly except on a very fine grid (30M/2.5mm ~= 12K array to get 1 sample per gap). 13 rings are required to shade the entire aperture. The first step in defining the aperture is to indicate which segment has which ID from prysm (which are deterministic) and mark the ones missing from the observatory for exclusion:
```
x, y = make_xy_grid(1024, diameter=30)
r, t = cart_to_polar(x, y)
flat_to_flat_to_vertex_vertex = 2 / np.sqrt(3)
vtov_to_flat_to_flat = 1 / flat_to_flat_to_vertex_vertex
segdiam = vtov_to_flat_to_flat * 1.44
cha = CompositeHexagonalAperture(x,y,13,segdiam,0.0025)
fig, ax = plt.subplots(figsize=(15,15))
ax.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
for center, id_ in zip(cha.all_centers, cha.segment_ids):
plt.text(*center, id_, ha='center', va='center')
```
The inner ring and center segment should be excluded, and only 6 segments exist per horizontal side, nor should the most extreme "columns" be present. The topmost segments are also not present. Let's start with this as an exclusion list:
```
exclude = [
0, 1, 2, 3, 4, 5, 6, # center
469, 470, 508, 509, 507, 510, 506, 545,
471, 511, 505, 544, 472, 397, 433, 546, # top, bottom
534, 533, 532, 531, 521, 522, 523, 524, # left edge
482, 483, 484, 485, 495, 494, 493, 492, # right edge
]
cha = CompositeHexagonalAperture(x,y,13,segdiam,0.0025, exclude=exclude)
fig, ax = plt.subplots(figsize=(15,15))
ax.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
for center, id_ in zip(cha.all_centers, cha.segment_ids):
plt.text(*center, id_, ha='center', va='center')
```
Next we can see that the diagonal "corners" are too large. With the exclusion list below, we can create a TMT pupil, excepting struts and SM obscuration, in only two lines of code.
```
exclude = [
0, 1, 2, 3, 4, 5, 6, # center
469, 470, 508, 509, 507, 510, 506, 545,
471, 511, 505, 544, 472, 397, 433, 546, # top, bottom
534, 533, 532, 531, 521, 522, 523, 524, # left edge
482, 483, 484, 485, 495, 494, 493, 492, # right edge
457, 535, 445, 520, 481, 409, 421, 496, # corners
536, 537, 479, 480, 497, 498, 519, 518, # next 'diagonal' from corners
]
cha = CompositeHexagonalAperture(x,y,13,segdiam,0.0025, exclude=exclude)
fig, ax = plt.subplots(figsize=(15,15))
ax.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
```
The TMT secondary obscuration is of 3.65 m diameter, we add it and struts of 50 cm diameter that are equiangular:
```
spider_ = spider(3, .5, x, y, rotation=90)
sm_obs = ~circle(3.65/2, r)
plt.imshow(cha.amp&spider_&sm_obs, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
```
Last of all are the six cables, of 20 mm diameter. These are a bit tricky, but they have a meeting point at 90% the radius of the SM obscuration. We will form them similar to the JWST and LUVOIR-A spiders, by shifting the coordinate grid and specifying the angle. The angles are about 10$^\circ$ from the radial normal.
```
# first cable bundle
r_offset = 3.65/2*.8
center_angle = np.radians(90)
center_c1 = (np.cos(center_angle) * r_offset, np.sin(center_angle) * r_offset)
cable1 = spider(1, 0.02, x, y, rotation=25.5, center=center_c1)
cable2 = spider(1, 0.02, x, y, rotation=180-25.5, center=center_c1)
center_angle = np.radians(-30)
center_c1 = (np.cos(center_angle) * r_offset, np.sin(center_angle) * r_offset)
cable3 = spider(1, 0.02, x, y, rotation=34.5, center=center_c1)
cable4 = spider(1, 0.02, x, y, rotation=-90-4.5, center=center_c1)
center_angle = np.radians(210)
center_c1 = (np.cos(center_angle) * r_offset, np.sin(center_angle) * r_offset)
cable5 = spider(1, 0.02, x, y, rotation=180-34.5, center=center_c1)
cable6 = spider(1, 0.02, x, y, rotation=-90+4.5, center=center_c1)
cables = cable1&cable2&cable3&cable4&cable5&cable6
fig, ax = plt.subplots(figsize=(15,15))
ax.imshow(cha.amp&spider_&sm_obs&cables, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
ax.set_title('Fully composited TMT aperture')
```
## LUVOIR-A
LUVOIR-A (as of the 2018 new design) contains 120 hexagonal segments of flat-to-flat dimension 1.223 m. Only the central segment is missing. The strut design is essentially the same as JWST. The first step in defining the aperture is to indicate which segment has which ID from prysm (which are deterministic) and mark the ones missing from the observatory for exclusion:
```
x, y = make_xy_grid(512, diameter=15)
cha = CompositeHexagonalAperture(x,y,6,1.223,0.007)
fig, ax = plt.subplots(figsize=(10,10))
ax.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
for center, id_ in zip(cha.all_centers, cha.segment_ids):
plt.text(*center, id_)
```
Note that we have discarded all of the other information from the composition process, which will be identical to the previous invocation. We now add the spider, pretty much the same as JWST:
```
exclude = [
0,
91,
109,
97,
103,
115,
121
]
cha = CompositeHexagonalAperture(x,y,6,1.223,0.007, exclude=exclude)
m1 = spider(1, .2, x, y, rotation=-105)
m2 = spider(1, .2, x, y, rotation=-75)
m3 = spider(1, .2, x, y, rotation=90)
spider_ = m1&m2&m3
plt.imshow(cha.amp&spider_, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
plt.title('Fully composited LUVOIR-A aperture')
```
## LUVOIR-B
LUVOIR-B is a smaller, unobscured co-design to LUVOIR-A using the same segment architecture. We follow a similar two-step shading process to find which segment IDs must be excluded:
```
x, y = make_xy_grid(512, diameter=8)
cha = CompositeHexagonalAperture(x,y,4,.955,.007)
fig, ax = plt.subplots(figsize=(10,10))
ax.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
for center, id_ in zip(cha.all_centers, cha.segment_ids):
plt.text(*center, id_)
exclude = [
37,
41,
45,
49,
53,
57
]
cha = CompositeHexagonalAperture(x,y,4,.955,.007, exclude=exclude)
plt.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
plt.title('Fully composited LUVOIR-B aperture')
```
## HabEx-A
Habex architecture A is a 4m unobscured system, which is extremely simple to model:
```
x, y = make_xy_grid(512, diameter=4)
r, t = cart_to_polar(x, y)
mask = circle(2, r)
plt.imshow(mask, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
plt.title('Fully composited HabEx A pupil')
```
## HabEx-B
Habex architecture B is an unobscured pupil of 6.5 m diameter based on a 3-ring fully populated hexagonal composition
```
x, y = make_xy_grid(512, diameter=6.5)
# vtov, centers, windows, local_coords, local_masks, segment_ids, mask = composite_hexagonal_aperture(3, 0.825, 0.007, x, y, exclude=[])
cha = CompositeHexagonalAperture(x,y,3,.825,0.007)
plt.imshow(cha.amp, origin='lower', cmap='gray', extent=[x.min(), x.max(), y.min(), y.max()])
plt.title('Fully composited HabEx B pupil')
```
| github_jupyter |
```
!pip install tensorflow==1.0
import tensorflow
tensorflow.__version__
!pip install gym
```
# Reinforcement Learning
Reinforcement learning is an area of machine learning inspired by behaviorist psychology, concerned with how software agents ought to take actions in an environment so as to maximize some notion of cumulative reward. (Source: Wikipedia)
Reinforcement learning provides the capacity for us not only to teach an artificial agent how to act, but to allow it to learn through it’s own interactions with an environment. By combining the complex representations that deep neural networks can learn with the goal-driven learning of an RL agent, computers have accomplished some amazing feats, like beating humans at over a dozen Atari games, and defeating the Go world champion.
RL algorithms must enable the agent to learn the correct pairings itself through the use of *observations*, *rewards*, and *actions*.
Typical aspects of a task that make it an RL problem are the following:
1. Different actions yield different rewards. For example, when looking for treasure in a maze, going left may lead to the treasure, whereas going right may lead to a pit of snakes.
2. Rewards are delayed over time. This just means that even if going left in the above example is the right things to do, we may not know it till later in the maze.
3. Reward for an action is conditional on the state of the environment. Continuing the maze example, going left may be ideal at a certain fork in the path, but not at others.
## Multi-armed bandit
The simplest reinforcement learning problem is the multi-armed bandit. Essentially, there are $n$-many slot machines, each with a different fixed payout probability.
The goal is to discover the machine with the best payout, and maximize the returned reward by always choosing it.
<img src="images/slot.jpg" alt="" style="width: 400px;"/>
This question has been the subject of active research since the 1950s, and many variations have been studied.
#### A-B Testing
Traditional A-B testing can be thought of as a special case of the multi-armed bandit problem, in which we choose to pursue a strategy of pure exploration in the initial testing phase, followed by a period of pure exploitation in which we choose the most valuable “arm” 100% of the time.
If the exploitation phase can be assumed to be much longer than the exploration phase, this approach is usually reasonable, as the wasted resources during the exploration are insignificant relative to the total rewards. However, in cases where the cost of the exploration phase is non-negligible, or in cases in which arm values are changing dynamically on short enough timescales that it becomes impractical to repeatedly perform new A-B tests, alternative approaches are needed.
The n-armed bandit is a nice starting place because we don’t have to worry about aspects #2 and #3. All we need to focus on is learning which rewards we get for each of the possible actions, and ensuring we chose the optimal ones.
### Epsilon-greedy
The most straightforward algorithm for continuously balancing exploration with exploitation is called “epsilon-greedy”.
Here, we pull a randomly chosen arm a fraction $\epsilon$ of the time. The other $1-\epsilon$ of the time, we pull the arm which we estimate to be the most profitable. As each arm is pulled and rewards are received, our estimates of arm values are updated. This method can be thought of a a continuous testing setup, where we devote a fraction $\epsilon$ of our resources to testing.
The following python code implements a simple 10-Armed Bandit using the epsilon-greedy algorithm.
The payout rate of the arms are normally distributed with $\mu = 0$ and $\sigma = 1$. Gaussian noise is also added to the rewards, also with $\mu = 0$ and $\sigma = 1$. (See [Sutton and Barto](http://people.inf.elte.hu/lorincz/Files/RL_2006/SuttonBook.pdf) book, section 2.1)
```
import numpy as np
import seaborn
class Bandit:
def __init__(self):
self.arm_values = np.random.normal(0,1,10)
self.K = np.zeros(10)
self.est_values = np.zeros(10)
def get_reward(self,action):
noise = np.random.normal(0,1)
reward = self.arm_values[action] + noise
return reward
def choose_eps_greedy(self,epsilon):
rand_num = np.random.random()
if epsilon>rand_num:
return np.random.randint(10)
else:
return np.argmax(self.est_values)
def update_est(self,action,reward):
self.K[action] += 1
alpha = 1./self.K[action]
self.est_values[action] += alpha * (reward - self.est_values[action])
```
How are we estimating the value of an action?
If by the $t$-th time step action $a$ has been chosen $K_a$ times prior to $t$, yielding rewards
$R_1, R_2, . . . , R_{Ka}$, then its value is estimated to be:
$$
Q_t(a) = \frac{R_1 + R_2 + . . .+ R_{Ka}}{K_a}
$$
A problem with this straightforward implementation is that its memory and computational requirements grow over time without
bound (we have to maintain, for each action $a$, a record of all the rewards that have followed the
selection of that action), but we can derive an incremental formula for computing averages with small, constant computation
required to process each new reward.
\begin{eqnarray*}
Q_{k+1} & = & \frac{1}{k+1} \sum_{i=1}^{k+1} R_i \\
& = & \frac{1}{k+1} \left( R_{k+1} + \sum_{i=1}^k R_i
\right) \\
& = & \frac{1}{k+1} \left( R_{k+1} + kQ_k + Q_k - Q_k
\right) \\
& = & \frac{1}{k+1} \left(R_{k+1} + (k+1) Q_k - Q_k
\right) \\
& = & Q_k + \frac{1}{k+1} \left( R_{k+1} - Q_k \right)
\end{eqnarray*}
```
def experiment(bandit,Npulls,epsilon):
history = []
for i in range(Npulls):
action = bandit.choose_eps_greedy(epsilon)
R = bandit.get_reward(action)
bandit.update_est(action,R)
history.append(R)
return np.array(history)
```
Let's make three different experiments: $\epsilon = 0$, $\epsilon = 0.1$ and $\epsilon = 0.01$. Data will be averages over 2000 tasks.
```
Nexp = 2000
Npulls = 5000
avg_outcome_eps0p0 = np.zeros(Npulls)
avg_outcome_eps0p01 = np.zeros(Npulls)
avg_outcome_eps0p1 = np.zeros(Npulls)
for i in range(Nexp):
bandit = Bandit()
avg_outcome_eps0p0 += experiment(bandit,Npulls,0.0)
bandit = Bandit()
avg_outcome_eps0p01 += experiment(bandit,Npulls,0.01)
bandit = Bandit()
avg_outcome_eps0p1 += experiment(bandit,Npulls,0.1)
avg_outcome_eps0p0 /= np.float(Nexp)
avg_outcome_eps0p01 /= np.float(Nexp)
avg_outcome_eps0p1 /= np.float(Nexp)
# plot results
import matplotlib.pyplot as plt
plt.plot(avg_outcome_eps0p0,label="eps = 0.0", alpha=0.5)
plt.plot(avg_outcome_eps0p01,label="eps = 0.01", alpha=0.5)
plt.plot(avg_outcome_eps0p1,label="eps = 0.1", alpha=0.5)
plt.ylim(0,2.2)
plt.legend()
plt.gcf().set_size_inches((8,3))
plt.show()
```
Although ε-greedy action selection is an effective and popular means of balancing exploration and exploitation in reinforcement learning, one drawback is that when it explores it chooses equally among all actions. This means that it is as likely to choose the worst-appearing action as it is to choose the next-to-best action. In tasks where the worst actions are very bad, this may
be unsatisfactory. The obvious solution is to vary the action probabilities as a graded function of estimated value.
The greedy action will be given the highest selection probability, but all the others will be ranked and weighted according to
their value estimates. To this end we can use a softmax action selection rule:
$$
\frac{ e^{Q_t(a)/ \tau}}{\sum_{i=1}^n e^{Q_t(i)/ \tau}}
$$
### Contextual Bandit
In the bandit problem described above, it is assumed that nothing is known about each arm other than what we have learned from prior pulls. We can relax this assumption and assume that for each arm there is a d-dimensional “context” vector. For example, if each arm represents a digital ad, the features in these vectors may correspond to things like banner size, web browser type, font color, etc. We can now model the value of each arm using these context vectors as well as past rewards in order to inform our choice of which arm to pull. This scenario is known as the contextual bandit problem.
## Q-learning
Unlike other methods, which attempt to learn functions which directly map an observation to an action, Q-Learning attempts **to learn the value of being in a given state, and taking a specific action there**.
The agent is in a state $s$ and has to choose one action $a$, upon which it receives a reward $r$ and come to a new state $s’$. The way the agent chooses actions is called **policy**.
Let’s define a function $Q(s, a)$ such that for given state $s$ and action $a$ it returns an estimate of a total reward we would achieve starting at this state, taking the action and then following some policy. Under certain conditions, there certainly exist policies that are optimal, meaning that they always select an action which is the best in the context. Let’s call the $Q$ function for these optimal policies $Q^*$.
If we knew the true $Q^*$ function, the solution would be straightforward. We would just apply a greedy policy to it. That means that in each state $s$, we would just choose an action a that maximizes the function $Q^*$, $argmax_a Q^*(s, a)$. Knowing this, our problem reduces to find a good estimate of the $Q^*$ function and apply the greedy policy to it.
Let’s write a formula for this function in a symbolic way. It is a sum of rewards we achieve after each action, but we will discount every member with γ:
$$ Q^*(s, a) = r_0 + \gamma r_1 + \gamma^2 r_2 + \gamma^3 r_3 + ... $$
$\gamma$ is called a discount factor and when set it to $\gamma < 1$ , it makes sure that the sum in the formula is finite. Value of each member exponentially diminish as they are more and more in the future and become zero in the limit. The $\gamma$ therefore controls how much the function Q in state s depends on the future and so it can be thought of as how much ahead the agent sees.
Typically we set it to a value close, but lesser to one. The actions are chosen according to the greedy policy, maximizing the $Q^*$ function.
When we look again at the formula, we see that we can write it in a recursive form:
$$Q^*(s, a) = r_0 + \gamma (r_1 + \gamma r_2 + \gamma^2 r_3 + ...) = r_0 + \gamma max_a Q^*(s', a)$$
We just derived a so called **Bellman equation**.
One of the possible strategies to solve the Bellman equation is by applying the **Q-learning** algorithm:
```
For each state-action pair (s, a), initialize the table entry Q(s,a) to zero
Observe the current state s
Do forever:
- Select an action a from s and execute it
- Receive immediate reward r
- Observe the new state s'
- Update the table entry for Q
- s=s'
```
#### Action selection
We could apply different strategies for action selection:
+ **Random approach**. Only in circumstances where a random policy is optimal would this approach be ideal.
+ **$\epsilon$- greedy approach**: A simple combination of the greedy and random approaches yields one of the most used exploration strategies. At the start of the training process the $\epsilon$ value is often initialized to a large probability, to encourage exploration in the face of knowing little about the environment. The value is then annealed down to a small constant (often 0.1), as the agent is assumed to learn most of what it needs about the environment. Despite the prevalence of usage that it enjoys, this method is far from optimal, since it takes into account only whether actions are most rewarding or not.
+ **Boltzmann Approach**. Instead of always taking the optimal action, or taking a random action, this approach involves choosing an action with weighted probabilities. To accomplish this we use a softmax over the networks estimates of value for each action. In practice we utilize an additional temperature parameter ($\tau$) which is annealed over time. While this measure can be a useful proxy, it is not exactly what would best aid exploration. What we really want to understand is the agent’s uncertainty about the value of different actions.
+ **Bayesian Approaches**. What if an agent could exploit its own uncertainty about its actions? This is exactly the ability that a class of neural network models referred to as Bayesian Neural Networks (BNNs) provide. Unlike traditional neural network which act deterministically, BNNs act probabilistically. This means that instead of having a single set of fixed weights, a BNN maintains a probability distribution over possible weights. In a reinforcement learning setting, the distribution over weight values allows us to obtain distributions over actions as well. The variance of this distribution provides us an estimate of the agent’s uncertainty about each action. In order to get true uncertainty estimates, multiple samples are required, thus increasing computational complexity.
#### Table updating
The table entry for $Q$ is updated by using this formula:
$$
Q(s,a) = Q(s,a) + \alpha [r + \gamma max_{a'} Q(s',a') - Q(s,a) ]
$$
where
+ $0<\alpha<1$ is the learning rate. Setting it to 0 means that the Q-values are never updated, hence nothing is learned. Setting a high value such as 0.9 means that learning can occur quickly.
+ $0<\gamma<1$ is the discount factor. This models the fact that future rewards are worth less than immediate rewards.
See http://www.scholarpedia.org/article/Temporal_difference_learning for a short description.
### Solving the FrozenLake problem
We are going to to solve the FrozenLake environment from the OpenAI gym.
For those unfamiliar, the OpenAI gym provides an easy way for people to experiment with their learning agents in an array of provided toy games. The FrozenLake environment consists of a 4x4 grid of blocks, each one either being the start block, the goal block, a safe frozen block, or a dangerous hole.
> **FrozenLake-v0**
> The agent controls the movement of a character in a grid world. Some tiles of the grid are walkable, and others lead to the agent falling into the water. Additionally, the movement direction of the agent is uncertain and only partially depends on the chosen direction. The agent is rewarded for finding a walkable path to a goal tile.
> The surface is described using a grid like the following:
>``SFFF (S: starting point, safe)``
>``FHFH (F: frozen surface, safe)``
>``FFFH (H: hole, fall to your doom)``
>``HFFG (G: goal, where the frisbee is located)``
> The episode ends when you reach the goal or fall in a hole. You receive a reward of 1 if you reach the goal, and zero otherwise.
The objective is to have an agent learn to navigate from the start to the goal without moving onto a hole. At any given time the agent can choose to move either ``up, down, left``, or ``right``.
The catch is that there is a wind which occasionally blows the agent onto a space they didn’t choose. As such, perfect performance every time is impossible, but learning to avoid the holes and reach the goal are certainly still doable. The reward at every step is 0, except for entering the goal, which provides a reward of 1. Thus, we will need an algorithm that learns long-term expected rewards. This is exactly what Q-Learning is designed to provide.
In it’s simplest implementation, Q-Learning is a table of values for every state (row) and action (column) possible in the environment. Within each cell of the table, we learn a value for how good it is to take a given action within a given state.
In the case of the FrozenLake environment, we have 16 possible states (one for each block), and 4 possible actions (the four directions of movement), giving us a $16 \times 4$ table of Q-values. We start by initializing the table to be uniform (all zeros), and then as we observe the rewards we obtain for various actions, we update the table accordingly.
```
import gym
import numpy as np
env = gym.make('FrozenLake-v0')
#Initialize table with all zeros
Q = np.zeros([env.observation_space.n,env.action_space.n])
# Set learning parameters
lr = .9
gamma = 0.95
num_episodes = 10000
#create lists to contain total rewards and steps per episode
rList = []
for i in range(num_episodes):
#Reset environment and get first new observation
s = env.reset()
rAll = 0
d = False
j = 0
#The Q-Table learning algorithm
while j < 999999:
j+=1
#Choose an action by greedily (with noise) picking from Q table
a = np.argmax(Q[s,:] + np.random.randn(1,env.action_space.n)*(1./(i+1)))
#Get new state and reward from environment
s1,r,d,_ = env.step(a)
#Update Q-Table with new knowledge
Q[s,a] = Q[s,a] + lr*(r + gamma*np.max(Q[s1,:]) - Q[s,a])
rAll += r
s = s1
if d == True:
break
rList.append(rAll)
```
FrozenLake-v0 is considered "solved" when the agent obtains an average reward of at least 0.78 over 100 consecutive episodes.
```
print "Score over time: " + str(sum(rList[-100:])/100)
print "Final Q-Table Values"
print Q
```
## Q-Learning with Neural Networks
Now, you may be thinking: tables are great, but they don’t really scale, do they? While it is easy to have a 16x4 table for a simple grid world, the number of possible states in any modern game or real-world environment is nearly infinitely larger. For most interesting problems, tables simply don’t work.
We instead need some way to take a description of our state, and produce $Q$-values for actions without a table: that is where neural networks come in. By acting as a function approximator, we can take any number of possible states that can be represented as a vector and learn to map them to $Q$-values.
In the case of the FrozenLake example, we will be using a one-layer network which takes the state encoded in a one-hot vector (1x16), and produces a vector of 4 $Q$-values, one for each action.
Such a simple network acts kind of like a glorified table, with the network weights serving as the old cells. The key difference is that we can easily expand the Tensorflow network with added layers, activation functions, and different input types, whereas all that is impossible with a regular table.
<img src="images/pong.jpg" alt="" style="width: 300px;"/>
The method of updating is a little different as well. Instead of directly updating our table, with a network we will be using backpropagation and a loss function. Our loss function will be sum-of-squares loss, where the difference between the current predicted $Q$-values, and the “target” value is computed and the gradients passed through the network. In this case, our $Q_{target}$ for the chosen action is the equivalent to the $Q$-value computed in equation above ($
Q(s,a) + \alpha [r + \gamma max_{a'} Q(s',a') - Q(s,a) ]
$).
$$
Loss = \sum (Q_{target} - Q_{predicted})^2
$$
```
import gym
import numpy as np
import random
import tensorflow as tf
import matplotlib.pyplot as plt
%matplotlib inline
env = gym.make('FrozenLake-v0')
tf.reset_default_graph()
#These lines establish the feed-forward part of the network used to choose actions
inputs1 = tf.placeholder(shape=[1,16],dtype=tf.float32)
W = tf.Variable(tf.random_uniform([16,4],0,0.01))
Qout = tf.matmul(inputs1,W)
predict = tf.argmax(Qout,1)
#Below we obtain the loss by taking the sum of squares difference between the target and prediction Q values.
nextQ = tf.placeholder(shape=[1,4],dtype=tf.float32)
loss = tf.reduce_sum(tf.square(nextQ - Qout))
trainer = tf.train.GradientDescentOptimizer(learning_rate=0.1)
updateModel = trainer.minimize(loss)
init = tf.global_variables_initializer()
# Set learning parameters
y = .99
e = 0.1
num_episodes = 2000
#create lists to contain total rewards and steps per episode
jList = []
rList = []
with tf.Session() as sess:
sess.run(init)
for i in range(num_episodes):
#Reset environment and get first new observation
s = env.reset()
rAll = 0
d = False
j = 0
#The Q-Network
while j < 99:
j+=1
#Choose an action by greedily (with e chance of random action) from the Q-network
a,allQ = sess.run([predict,Qout],
feed_dict={inputs1:np.identity(16)[s:s+1]})
if np.random.rand(1) < e:
a[0] = env.action_space.sample()
#Get new state and reward from environment
s1,r,d,_ = env.step(a[0])
#Obtain the Q' values by feeding the new state through our network
Q1 = sess.run(Qout,
feed_dict={inputs1:np.identity(16)[s1:s1+1]})
#Obtain maxQ' and set our target value for chosen action.
maxQ1 = np.max(Q1)
targetQ = allQ
targetQ[0,a[0]] = r + y*maxQ1
#Train our network using target and predicted Q values
_,W1 = sess.run([updateModel,W],
feed_dict={inputs1:np.identity(16)[s:s+1],
nextQ:targetQ})
rAll += r
s = s1
if d == True:
#Reduce chance of random action as we train the model.
e = 1./((i/50) + 10)
break
jList.append(j)
rList.append(rAll)
print "Percent of succesful episodes: " + str(sum(rList[-100:])/100) + "%"
```
We can see that the network beings to consistly reach the goal around the 1000 episode mark.
```
plt.plot(rList)
```
It also begins to progress through the environment for longer than chance aroudn the 1000 mark as well.
```
plt.plot(jList)
```
## Deep Q-networks
While our ordinary Q-network was able to barely perform as well as the Q-Table in a simple game environment, Deep $Q$-Networks are much more capable. In order to transform an ordinary Q-Network into a DQN we will be making the following improvements:
+ Going from a single-layer network to a multi-layer convolutional network.
+ Implementing Experience Replay, which will allow our network to train itself using stored memories from it’s experience.
+ Utilizing a second “target” network, which we will use to compute target $Q$-values during our updates.
<img src="images/deepq1.png" alt="" style="width: 800px;"/>
See https://jaromiru.com/2016/09/27/lets-make-a-dqn-theory/
### Convolutional Layers
Since our agent is going to be learning to play video games, it has to be able to make sense of the game’s screen output in a way that is at least similar to how humans or other intelligent animals are able to. Instead of considering each pixel independently, convolutional layers allow us to consider regions of an image, and maintain spatial relationships between the objects on the screen as we send information up to higher levels of the network.
### Experience Replay
The second major addition to make DQNs work is Experience Replay.
The problem with online learning is that the *samples arrive in order* they are experienced and as such are highly correlated. Because of this, our network will most likely overfit and fail to generalize properly.
The key idea of **experience replay** is that we store these transitions in our memory and during each learning step, sample a random batch and perform a gradient descend on it.
The Experience Replay buffer stores a fixed number of recent memories, and as new ones come in, old ones are removed. When the time comes to train, we simply draw a uniform batch of random memories from the buffer, and train our network with them.
### Separate Target Network
This second network is used to generate the $Q$-target values that will be used to compute the loss for every action during training.
The issue is that at every step of training, the $Q$-network’s values shift, and if we are using a constantly shifting set of values to adjust our network values, then the value estimations can easily spiral out of control. The network can become destabilized by falling into feedback loops between the target and estimated $Q$-values. In order to mitigate that risk, the target network’s weights are fixed, and only periodically or slowly updated to the primary $Q$-networks values. In this way training can proceed in a more stable manner.
Instead of updating the target network periodically and all at once, we will be updating it frequently, but slowly.
While the DQN we have described above could learn ATARI games with enough training, getting the network to perform well on those games takes at least a day of training on a powerful machine.
```
from __future__ import division
import gym
import numpy as np
import random
import tensorflow as tf
import tensorflow.contrib.slim as slim
import matplotlib.pyplot as plt
import scipy.misc
import os
%matplotlib inline
import numpy as np
import random
import itertools
import scipy.misc
import matplotlib.pyplot as plt
class gameOb():
def __init__(self,coordinates,size,intensity,channel,reward,name):
self.x = coordinates[0]
self.y = coordinates[1]
self.size = size
self.intensity = intensity
self.channel = channel
self.reward = reward
self.name = name
class gameEnv():
def __init__(self,partial,size):
self.sizeX = size
self.sizeY = size
self.actions = 4
self.objects = []
self.partial = partial
a = self.reset()
plt.imshow(a,interpolation="nearest")
def reset(self):
self.objects = []
hero = gameOb(self.newPosition(),1,1,2,None,'hero')
self.objects.append(hero)
bug = gameOb(self.newPosition(),1,1,1,1,'goal')
self.objects.append(bug)
hole = gameOb(self.newPosition(),1,1,0,-1,'fire')
self.objects.append(hole)
bug2 = gameOb(self.newPosition(),1,1,1,1,'goal')
self.objects.append(bug2)
hole2 = gameOb(self.newPosition(),1,1,0,-1,'fire')
self.objects.append(hole2)
bug3 = gameOb(self.newPosition(),1,1,1,1,'goal')
self.objects.append(bug3)
bug4 = gameOb(self.newPosition(),1,1,1,1,'goal')
self.objects.append(bug4)
state = self.renderEnv()
self.state = state
return state
def moveChar(self,direction):
# 0 - up, 1 - down, 2 - left, 3 - right
hero = self.objects[0]
heroX = hero.x
heroY = hero.y
penalize = 0.
if direction == 0 and hero.y >= 1:
hero.y -= 1
if direction == 1 and hero.y <= self.sizeY-2:
hero.y += 1
if direction == 2 and hero.x >= 1:
hero.x -= 1
if direction == 3 and hero.x <= self.sizeX-2:
hero.x += 1
if hero.x == heroX and hero.y == heroY:
penalize = 0.0
self.objects[0] = hero
return penalize
def newPosition(self):
iterables = [ range(self.sizeX), range(self.sizeY)]
points = []
for t in itertools.product(*iterables):
points.append(t)
currentPositions = []
for objectA in self.objects:
if (objectA.x,objectA.y) not in currentPositions:
currentPositions.append((objectA.x,objectA.y))
for pos in currentPositions:
points.remove(pos)
location = np.random.choice(range(len(points)),replace=False)
return points[location]
def checkGoal(self):
others = []
for obj in self.objects:
if obj.name == 'hero':
hero = obj
else:
others.append(obj)
ended = False
for other in others:
if hero.x == other.x and hero.y == other.y:
self.objects.remove(other)
if other.reward == 1:
self.objects.append(gameOb(self.newPosition(),1,1,1,1,'goal'))
else:
self.objects.append(gameOb(self.newPosition(),1,1,0,-1,'fire'))
return other.reward,False
if ended == False:
return 0.0,False
def renderEnv(self):
#a = np.zeros([self.sizeY,self.sizeX,3])
a = np.ones([self.sizeY+2,self.sizeX+2,3])
a[1:-1,1:-1,:] = 0
hero = None
for item in self.objects:
a[item.y+1:item.y+item.size+1,item.x+1:item.x+item.size+1,item.channel] = item.intensity
if item.name == 'hero':
hero = item
if self.partial == True:
a = a[hero.y:hero.y+3,hero.x:hero.x+3,:]
b = scipy.misc.imresize(a[:,:,0],[84,84,1],interp='nearest')
c = scipy.misc.imresize(a[:,:,1],[84,84,1],interp='nearest')
d = scipy.misc.imresize(a[:,:,2],[84,84,1],interp='nearest')
a = np.stack([b,c,d],axis=2)
return a
def step(self,action):
penalty = self.moveChar(action)
reward,done = self.checkGoal()
state = self.renderEnv()
if reward == None:
print(done)
print(reward)
print(penalty)
return state,(reward+penalty),done
else:
return state,(reward+penalty),done
env = gameEnv(partial=False,size=5)
```
Above is an example of a starting environment in our simple game. The agent controls the blue square, and can move up, down, left, or right. The goal is to move to the green square (for +1 reward) and avoid the red square (for -1 reward). The position of the three blocks is randomized every episode.
```
class Qnetwork():
def __init__(self,h_size):
#The network recieves a frame from the game, flattened into an array.
#It then resizes it and processes it through four convolutional layers.
self.scalarInput = tf.placeholder(shape=[None,21168],dtype=tf.float32)
self.imageIn = tf.reshape(self.scalarInput,shape=[-1,84,84,3])
self.conv1 = slim.conv2d( \
inputs=self.imageIn,num_outputs=32,kernel_size=[8,8],stride=[4,4],padding='VALID', biases_initializer=None)
self.conv2 = slim.conv2d( \
inputs=self.conv1,num_outputs=64,kernel_size=[4,4],stride=[2,2],padding='VALID', biases_initializer=None)
self.conv3 = slim.conv2d( \
inputs=self.conv2,num_outputs=64,kernel_size=[3,3],stride=[1,1],padding='VALID', biases_initializer=None)
self.conv4 = slim.conv2d( \
inputs=self.conv3,num_outputs=h_size,kernel_size=[7,7],stride=[1,1],padding='VALID', biases_initializer=None)
#We take the output from the final convolutional layer and split it into separate advantage and value streams.
self.streamAC,self.streamVC = tf.split(self.conv4,2,3)
self.streamA = slim.flatten(self.streamAC)
self.streamV = slim.flatten(self.streamVC)
xavier_init = tf.contrib.layers.xavier_initializer()
self.AW = tf.Variable(xavier_init([h_size//2,env.actions]))
self.VW = tf.Variable(xavier_init([h_size//2,1]))
self.Advantage = tf.matmul(self.streamA,self.AW)
self.Value = tf.matmul(self.streamV,self.VW)
#Then combine them together to get our final Q-values.
self.Qout = self.Value + tf.subtract(self.Advantage,tf.reduce_mean(self.Advantage,axis=1,keep_dims=True))
self.predict = tf.argmax(self.Qout,1)
#Below we obtain the loss by taking the sum of squares difference between the target and prediction Q values.
self.targetQ = tf.placeholder(shape=[None],dtype=tf.float32)
self.actions = tf.placeholder(shape=[None],dtype=tf.int32)
self.actions_onehot = tf.one_hot(self.actions,env.actions,dtype=tf.float32)
self.Q = tf.reduce_sum(tf.multiply(self.Qout, self.actions_onehot), axis=1)
self.td_error = tf.square(self.targetQ - self.Q)
self.loss = tf.reduce_mean(self.td_error)
self.trainer = tf.train.AdamOptimizer(learning_rate=0.0001)
self.updateModel = self.trainer.minimize(self.loss)
class experience_buffer():
def __init__(self, buffer_size = 50000):
self.buffer = []
self.buffer_size = buffer_size
def add(self,experience):
if len(self.buffer) + len(experience) >= self.buffer_size:
self.buffer[0:(len(experience)+len(self.buffer))-self.buffer_size] = []
self.buffer.extend(experience)
def sample(self,size):
return np.reshape(np.array(random.sample(self.buffer,size)),[size,5])
def processState(states):
return np.reshape(states,[21168])
def updateTargetGraph(tfVars,tau):
total_vars = len(tfVars)
op_holder = []
for idx,var in enumerate(tfVars[0:total_vars//2]):
op_holder.append(tfVars[idx+total_vars//2].assign((var.value()*tau) + ((1-tau)*tfVars[idx+total_vars//2].value())))
return op_holder
def updateTarget(op_holder,sess):
for op in op_holder:
sess.run(op)
batch_size = 32 #How many experiences to use for each training step.
update_freq = 4 #How often to perform a training step.
y = .99 #Discount factor on the target Q-values
startE = 1 #Starting chance of random action
endE = 0.1 #Final chance of random action
anneling_steps = 10000. #How many steps of training to reduce startE to endE.
num_episodes = 10000 #How many episodes of game environment to train network with.
pre_train_steps = 10000 #How many steps of random actions before training begins.
max_epLength = 50 #The max allowed length of our episode.
load_model = False #Whether to load a saved model.
path = "./dqn" #The path to save our model to.
h_size = 512 #The size of the final convolutional layer before splitting it into Advantage and Value streams.
tau = 0.001 #Rate to update target network toward primary network
tf.reset_default_graph()
mainQN = Qnetwork(h_size)
targetQN = Qnetwork(h_size)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
trainables = tf.trainable_variables()
targetOps = updateTargetGraph(trainables,tau)
myBuffer = experience_buffer()
#Set the rate of random action decrease.
e = startE
stepDrop = (startE - endE)/anneling_steps
#create lists to contain total rewards and steps per episode
jList = []
rList = []
total_steps = 0
#Make a path for our model to be saved in.
if not os.path.exists(path):
os.makedirs(path)
with tf.Session() as sess:
sess.run(init)
if load_model == True:
print('Loading Model...')
ckpt = tf.train.get_checkpoint_state(path)
saver.restore(sess,ckpt.model_checkpoint_path)
updateTarget(targetOps,sess) #Set the target network to be equal to the primary network.
for i in range(num_episodes):
episodeBuffer = experience_buffer()
#Reset environment and get first new observation
s = env.reset()
s = processState(s)
d = False
rAll = 0
j = 0
#The Q-Network
while j < max_epLength: #If the agent takes longer than 200 moves to reach either of the blocks, end the trial.
j+=1
#Choose an action by greedily (with e chance of random action) from the Q-network
if np.random.rand(1) < e or total_steps < pre_train_steps:
a = np.random.randint(0,4)
else:
a = sess.run(mainQN.predict,feed_dict={mainQN.scalarInput:[s]})[0]
s1,r,d = env.step(a)
s1 = processState(s1)
total_steps += 1
episodeBuffer.add(np.reshape(np.array([s,a,r,s1,d]),[1,5])) #Save the experience to our episode buffer.
if total_steps > pre_train_steps:
if e > endE:
e -= stepDrop
if total_steps % (update_freq) == 0:
trainBatch = myBuffer.sample(batch_size) #Get a random batch of experiences.
#Below we perform the Double-DQN update to the target Q-values
Q1 = sess.run(mainQN.predict,feed_dict={mainQN.scalarInput:np.vstack(trainBatch[:,3])})
Q2 = sess.run(targetQN.Qout,feed_dict={targetQN.scalarInput:np.vstack(trainBatch[:,3])})
end_multiplier = -(trainBatch[:,4] - 1)
doubleQ = Q2[range(batch_size),Q1]
targetQ = trainBatch[:,2] + (y*doubleQ * end_multiplier)
#Update the network with our target values.
_ = sess.run(mainQN.updateModel, \
feed_dict={mainQN.scalarInput:np.vstack(trainBatch[:,0]),mainQN.targetQ:targetQ, mainQN.actions:trainBatch[:,1]})
updateTarget(targetOps,sess) #Set the target network to be equal to the primary network.
rAll += r
s = s1
if d == True:
break
myBuffer.add(episodeBuffer.buffer)
jList.append(j)
rList.append(rAll)
#Periodically save the model.
if i % 1000 == 0:
saver.save(sess,path+'/model-'+str(i)+'.cptk')
print("Saved Model")
if len(rList) % 10 == 0:
print(total_steps,np.mean(rList[-10:]), e)
saver.save(sess,path+'/model-'+str(i)+'.cptk')
print("Percent of succesful episodes: " + str(sum(rList)/num_episodes) + "%")
```
| github_jupyter |
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
#print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
```
## Ground-Truthing Local Weather Data
As a preliminary step toward participation in [Project GROW](https://growobservatory.org/), this is to test two elements of the mix that will be critical to our success, namely:
- Data logs from the FlowerPower soil sensor; and
- Weather reports from our nearest weather station.
First, let's take a look at representative data from these two sources, heads and tails, to see if we've caught a good month of overlapping records:
```
#df_s=pd.read_csv('../input/soil-sensor-test/Flower power 6A22.csv')
df_s=pd.read_csv('Flower power 6A22.csv',parse_dates=["capture_datetime_utc"], index_col="capture_datetime_utc")
df_w=pd.read_csv('portimao_hist_meteoclean.csv')
df_s.head()
df_s.tail()
df_w.head()
df_w.tail()
```
Great: the soil sensor data covers period from June 18 to July 18, and the weather data overlaps that period at both ends, so it looks like we've got a reasonable basis for comparison across these two data sets.
Thing is: the weather data has been recorded ever hour on the hour, while soil sensor data is recorded at 15 minute intervals that don't appear to fall exactly on the hourly moment... So to achieve alignment of this time-series data, we must munge the soil sensor records a bit to get a (reduced) set of records that aligns in terms of time with these hourly weather records.
```
# To match df_w, We downsample df_s from 15 minute intervals to hourly frequency, aggregated using mean
df_s_w = df_s.resample('H').mean()
print(df_s.shape)
print(df_s_w.shape)
df_s_w.head()
```
Great: we've got the soil sensor data at hourly intervals, so we can match it to the weather data... Once we concatenate the related columns in weather df into a single datetime index.
```
# Now we create a datetime index composed of the 1st 5 columns (integers), concatenated
df_w.index = pd.to_datetime(df_w[['Year', 'Month', 'Day', 'Hour', 'Minute']])
df_w.head()
```
Now, we can simplify the weather dataframe in both dimensions, i.e.:
- drop all extraneous columns; and
- filter rows to match the 1-month range of dates in df_s (df_w both starts earlier and ends later).
```
df_w.drop(['Year','Month','Day','Hour','Minute','Snowfall amount raw [sfc]','High Cloud Cover [high cld lay]','Medium Cloud Cover [mid cld lay]','Low Cloud Cover [low cld lay]','Shortwave Radiation [sfc]','Wind Speed [80 m above gnd]','Wind Direction [80 m above gnd]','Wind Speed [900 mb]','Wind Direction [900 mb]','Wind Gust [sfc]'], axis = 1, inplace=True)
df_w.info()
df_w_s = df_w.loc['2018-06-18 19:00:00':'2018-07-18 15:18:29']
df_w_s.shape
print("soil sensor data:")
print(df_s_w.info())
print("")
print("weather data:")
print(df_w_s.info())
#test_merge = pd.merge(df_s_w, df_w_s, on=index)
test_merge = pd.merge(df_s_w, df_w_s, on=index)
test_merge.head()
```
| github_jupyter |
## Welcome to Week 4, Single Cell RNA (cont.)!
### This week, we're going to go a bit deeper into scRNA analysis, such as how to interact with Seurat objects, add additional datatypes including CITE-seq and TCR/BCR-seq data, and create custom, publication-ready plots.
We'll continue to use Scanpy, which has some nice capabilities for multi-modal data analysis. The two datatypes we will be working with today at **CITE-seq** and **TCR/BCR-seq** data. The main idea of both is that additional information about the cell is captured using the same cell barcode from reverse transcription so that multiple types of data can be assigned to the same cell. CITE-seq is a method for capturing surface protein information using oligo-conjugated antibodies developed at the New York Genome Center. Here antibodies are conjugated to oligos which contain two important sequences: an antibody specific barcode which is used to quantify surface protein levels in individual cells and a capture sequence (either poly-A sequence or a 10X specific capture sequence) which enables the antibody oligo to be tagged with the cell barcode during reverse transcription. You can look at more details in the publication here:
* https://www.ncbi.nlm.nih.gov/pubmed/28759029
Oligo-conjugated anitbodies compatible with 10X scRNA (both 5' and 3') are commercially available from BioLegend (https://www.biolegend.com/en-us/totalseq) and can also be used to multiplex different samples in the same 10X capture. This works by using an antibody which recognizes a common surface antigen and using the antibody barcode to distinguish between samples, a process known as **cell hashing**:
* https://www.ncbi.nlm.nih.gov/pubmed/30567574
We won't be using hashtag data today, but many of the same strategies apply and feel free to reach out if you are interested in learning more!
The second data type we will be working with is TCR/BCR sequencing data. T and B cells express a highly diverse repertoire of transcripts resulting from V(D)J recombination - the T cell receptor (TCR) in T cells and immunoglobulin (Ig) or BCR in B cells. Daughter cells will share the same TCR/BCR sequence, allowing this sequence to be used to track clonal cell populations over time and space, as well as infer lineage relationships. TCR/BCR sequences are amplified from the cDNA library in the 5' immune profiling 10X kit, allowing these sequences to be matched to the gene expression library from the same cell. For more details, see the 10X website:
* https://www.10xgenomics.com/products/vdj/
For both of these applications, we'll be following this tutorial:
* https://scanpy-tutorials.readthedocs.io/en/multiomics/cite-seq/pbmc5k.html
### Import Statements
```
import scanpy as sc
import pandas as pd
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
from collections import Counter, defaultdict
from scipy import stats as scistats
import scrublet as scr
import scipy.io
%matplotlib inline
# you'll need to change these for yourself
path = '/Users/kevin/changlab/covid19/3_scRNA/data/filtered_feature_bc_matrix/'
figpath = '/Users/kevin/changlab/covid19/4_scRNA-part-2/figures/'
# lets set the default figure settings
sc.settings.set_figure_params(dpi_save=300)
sc.settings.figdir = figpath
# helpful plotting functions, "sax" or "simple ax" and "prettify ax" or "pax"
def pax(ax):
mpl.rcParams['font.sans-serif'] = 'Helvetica'
for spine in ax.spines.values():
spine.set_color('k')
ax.set_frameon=True
ax.patch.set_facecolor('w')
ax.tick_params(direction='out', color = 'k', length=5, width=.75, pad=8)
ax.set_axisbelow(True)
ax.grid(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
mpl.rcParams['font.sans-serif'] = 'Helvetica'
def sax(figsize=(6,6)):
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(111)
pax(ax)
return fig, ax
def sf(fig, fn, formats=['png'], dpi=300, figpath=figpath):
for f in formats:
fig.savefig(figpath + fn + '.' + f, dpi=dpi, bbox_inches='tight')
```
### First, go back to the week three notebook, re-run everything, and save the output so you can just re-import the procssed dataset here.
Or, you can use the file that I outputted to have the same input. I've included the code that I ran to generate it below.
```
# # process with scrublet
# print('processing with scrublet')
# counts_matrix = scipy.io.mmread(path + '/matrix.mtx.gz').T.tocsc()
# cells = pd.read_csv(path + '/barcodes.tsv.gz', sep='\t', header=None, names=['barcode'])
# cells = cells.set_index('barcode', drop=False)
# scrub = scr.Scrublet(counts_matrix, expected_doublet_rate=0.08)
# doublet_scores, predicted_doublets = scrub.scrub_doublets(min_counts=2,
# min_cells=3,
# min_gene_variability_pctl=85,
# n_prin_comps=30)
# predicted_doublets = scrub.call_doublets(threshold=0.25)
# cells['doublet_score'] = doublet_scores
# cells['predicted_doublet'] = predicted_doublets
# # import data
# print('importing data')
# gex = sc.read_10x_mtx(path, gex_only=True)
# gex.obs['doublet_score'] = cells.loc[gex.obs.index, 'doublet_score']
# gex.obs['predicted_doublet'] = cells.loc[gex.obs.index, 'predicted_doublet']
# # preliminary processing
# print('preliminary processing')
# sc.pp.filter_cells(gex, min_genes=200)
# sc.pp.filter_genes(gex, min_cells=3)
# mito_genes = gex.var_names.str.startswith('MT-')
# gex.obs['percent_mito'] = np.sum(
# gex[:, mito_genes].X, axis=1).A1 / np.sum(gex.X, axis=1).A1
# gex.obs['n_counts'] = gex.X.sum(axis=1).A1
# gex = gex[gex.obs.n_genes >= 500, :]
# gex = gex[gex.obs.percent_mito < 0.1, :]
# sc.pp.normalize_total(gex, target_sum=1e4)
# sc.pp.log1p(gex)
# gex.raw = gex
# # dimensionality reduction
# print('secondary processing')
# sc.pp.highly_variable_genes(gex, n_top_genes=2000)
# gex = gex[:, gex.var.highly_variable]
# sc.pp.regress_out(gex, ['n_genes'])
# sc.pp.scale(gex, max_value=10)
# sc.tl.pca(gex, svd_solver='arpack', n_comps=50)
# sc.pp.neighbors(gex, n_neighbors=10, n_pcs=50, random_state=1)
# sc.tl.leiden(gex, random_state=1, resolution=.4)
# sc.tl.umap(gex)
# new_cluster_names = ['Mono_CD14', #0
# 'CD4 T', #1
# 'B', #2
# 'CD8 T', #3
# 'NK', #4
# 'CD8 Tem', #5
# 'Mono_FCGR3A', #6
# 'Und1_Doublets', #7
# 'cDC', #8
# 'gd T', #9 gamma delta t cells
# 'pDCs', #10
# 'Platelets', #11
# 'Plasma B', #12
# 'Und2', #13
# ]
# gex.rename_categories('leiden', new_cluster_names)
# # plot things
# print('plotting')
# # plot1
# fig = sc.pl.umap(gex, color=['leiden'],
# legend_fontsize = 8,
# legend_loc = 'on data', return_fig=True)
# fig.savefig(figpath + '0_leiden-clustering-renamed.png', dpi=300, bbox_inches='tight')
# # plot2
# genes_to_plot = ['predicted_doublet','n_genes',
# 'n_counts','percent_mito']
# fig = sc.pl.umap(gex, color=genes_to_plot, use_raw=True,
# sort_order=True, ncols=2, return_fig=True)
# fig.savefig(figpath + '0_umap-metadata.png', dpi=300, bbox_inches='tight')
# # plot3
# genes_to_plot = ['CD3G','CD4','CD8A',
# 'TRDV2','KLRB1','NKG7',
# 'CD14','FCGR3A','FCER1A',
# 'MS4A1','JCHAIN','PPBP',
# ]
# fig = sc.pl.umap(gex, color=genes_to_plot, use_raw=True,
# sort_order=True, ncols=3,return_fig=True, color_map='Reds')
# fig.savefig(figpath + '0_umap-gene-expression.png', dpi=300, bbox_inches='tight')
# # save the results
# gex.write(figpath + 'scrna_wk3_processed.h5ad', compression='gzip')
# import the data
# reminder that you'll need to change the path to this
gex = sc.read_h5ad(figpath + 'scrna_wk3_processed.h5ad')
gex
# make sure that everything looks good
genes_to_plot = ['CD3G','CD4','CD8A',
'TRDV2','KLRB1','NKG7',
'CD14','FCGR3A','FCER1A',
'MS4A1','JCHAIN','PPBP',
]
fig = sc.pl.umap(gex, color=genes_to_plot, use_raw=True,
sort_order=True, ncols=3,return_fig=True, color_map='Reds')
plt.show()
fig = sc.pl.umap(gex, color=['leiden'],
legend_fontsize = 8,
legend_loc = 'on data', return_fig=True)
plt.show()
```
### CITE-seq Analysis
```
# first, read in the cite seq information
# remember that gex_only=False will let you read them both in
data = sc.read_10x_mtx(path, gex_only=False)
data
# what cite seq features do we have?
# how many genes?
# how many cite-seq?
# rename the antibody capture genes
# get rid of the "_TotalSeqC" part of the name just to make our lives easier
# e.g. CD3_TotalSeqC to CD3
# filter this to just include cells that we analyzed previously, so the datasets will align
# you can do this with
data = data[data.obs.index.isin(gex.obs.index), :]
# now lets get just the protein information, and make that its own anndata object
protein = data[:, data.var['feature_types'] == 'Antibody Capture'].copy()
protein
```
### Now let's break out of scanpy for a minute to inspect, normalize, and scale this data on our own
Scanpy seems to be developing some functions specifically for protein data, but hasn't yet implemented them. But this isn't a problem! We can do things on our own, and transform the data into a format that scanpy wants.
**We're going to break this down in a few steps:**
1. get the raw antibody count data from the protein anndata object.
2. compute the centered log ratio (CLR) of antibody counts (this is different than for RNA!) - more notes on this below.
3. scale the data to be mean centered and have unit variance (i.e., z-normalization). This is the same as for RNA.
4. save the CLR normalized antibody counts as the raw data of the protein object, and the scaled data as the (normal) data of the protein object, which will be used for dimensionality reduction.
Now, in terms of what the actual normalizations are: we're going to do this with the .apply() function with dataframes. I'm providing an example for how to you would do the depth normalization that you'd normally do for RNA-seq below, but you should play around on your own with implementing the normalizations in 2 and 3.
**Normalization methods:**
* depth normalization (as a comparison). For a cell, divide the counts for each gene/antibody by the sum of all gene/antibody counts for that cell, then multiply by some scaling factor (e.g. 10,000). Commonly, you would also log transform this, and add a pseudocount (say 1). This is sometimes referred to as **log1p**.
* CLR. For an antibody, divide the counts for each antibody by the geometric mean antibody counts across all cells, then take the natural log of this. Similarly, you'll add a pseudocount of 1.
* z-normalization (scaling to zero mean and unit variance). Basically, you're making all datapoints have similar distributions. For a gene, return the count for a cell minus the mean count across all cells, divided by the standard deviation across all cells.
* clipping extremes. You can use the np.clip() function to do this. Basically, this will take any value lower than the lower bound in np.clip and make it equal to the lower bound, and do the same for the upper bound. You might combine this with computing the mean and standard deviation, to clip values > 3 stds away from the mean; or np.percentile() to clip values that are less or greater than a given percentile in the data.
It's worth taking the time to look at why the CLR transformation is better than a simple log transformation. Why? Because antibodies aren't genes - when a gene is negative, the count is 0; when a gene is positive, the count is greater than 0. But does this hold true with antibodies? When an antibody is negative, the count isn't necessarily 0 - the antibody might have background! The CLR transformation does a better job of dealing with this, by looking at the relative abundance of the antibody.
```
# get the raw data
protein_orig = pd.DataFrame(protein.X.todense(), index=protein.obs.index, columns=protein.var.index).T
# what does your data look like?
# I'd recommend first plotting the distribution of total antibody counts across all cells
# sf(fig, '1_preprocess_histogram_antibody-counts')
# what if we just take a 'naive' approach to normalization?
protein_norm_depth = protein_orig.apply(lambda x: 10000 * x / x.sum(), axis=0)
protein_norm_depth = np.log(protein_norm_depth + 1)
# plot the distribution of counts for all of these
fig = plt.figure(figsize=(10,10))
axes = [fig.add_subplot(5,4,i+1) for i in range(len(protein_norm_depth.index))]
xlim = [protein_norm_depth.min().min(), protein_norm_depth.max().max()]
bins = np.linspace(xlim[0], xlim[1], 100)
for ix, p in enumerate(protein_orig.index):
ax = axes[ix]
pax(ax)
vals = protein_norm_depth.loc[p]
ax.hist(vals, bins=bins)
ax.set_title(p, size=16)
ax.set_xlim(xlim)
fig.tight_layout()
plt.show()
sf(fig, '1_preprocess_log1p-distributions')
# now lets compare this with the CLR approach
def clr(x, pseudo=1):
x = x + pseudo
geo_mean = scistats.gmean(x)
return np.log(x / geo_mean)
protein_norm_clr = protein_orig.apply(clr, axis=1)
protein_norm_clr.head()
# plot the distribution of counts for all of these
# sf(fig, '1_preprocess_clr-distributions')
# now lets compare the two with a scatter plot
# sf(fig, '1_preprocess_scatter-norm-methods')
# now scale this to unit variance
# see https://en.wikipedia.org/wiki/Feature_scaling under z-normalization
# also, clip extremes - clip anything less than -10 and above 10
# plot the distribution of counts for all of the scaled data
# note how the distributions are relatively similar
# sf(fig, '1_preprocess_scaled_clr-distributions')
# what if we want to make a scatter plot of one CD4 vs CD8a?
# compare the depth-normalized vs CLR normalized counts
# make it once with depth-normalized counts and once with CLR normalized
# sf(fig, '1_preprocess_scatter_log1p_cd4-8')
# what if we want to make a scatter plot of one antibody?
# sf(fig, '1_preprocess_scatter_clr_cd4-8')
```
### Now go back to scanpy
Let's save the protein_norm_clr values as the raw data in protein, and the protein_scaled values in the data slot of protein. Let's also exclude the control proteins from the main data slot.
```
protein = data[:, data.var['feature_types'] == 'Antibody Capture'].copy()
protein.var['control'] = ['control' in i for i in protein.var.index]
protein.X = protein_norm_clr.T
protein.raw = protein
protein.X = protein_scaled.T
protein = protein[:, ~protein.var['control']]
protein
protein.var
protein_genes = ['CD3D','CD19','PTPRC',
'CD4','CD8A','CD14','FCGR3A',
'NCAM1','IL2RA','PTPRC',
'PDCD1','TIGIT','IL7R','FUT4']
protein.var['rna_name'] = protein_genes
name_dict = dict(zip(protein.var.index, protein.var['rna_name']))
protein.var.head()
sc.pp.pca(protein, n_comps=len(protein.var)-1)
sc.pp.neighbors(protein, n_neighbors=30, n_pcs=len(protein.var)-1)
sc.tl.leiden(protein, key_added="protein_leiden", resolution=.33)
sc.tl.umap(protein)
genes_to_plot = protein.var.index.tolist() + ['protein_leiden']
fig = sc.pl.umap(protein, color=genes_to_plot,
sort_order=True, ncols=4,return_fig=True, color_map='Blues', use_raw=True,
vmin='p5', vmax='p99.9')
fig.set_size_inches(12,12)
sf(fig,'2_umap_with_cite-clustering')
plt.show()
```
### Now let's integrate this with the RNA data
I'm going to do this a little fast and loose because I think that scanpy hasn't yet fully implemented the CITE-seq stuff too well. Basically, we're going to add the umap coordinates and clustering information from the RNA processed data to the protein-processed data, and vice versa.
```
# add gex to protein
protein.obsm['RNA_umap'] = gex[protein.obs.index].obsm['X_umap']
protein.obs['rna_leiden'] = gex.obs.loc[protein.obs.index, 'leiden']
# add protein to gex
# I'll leave you do to this
# now, let's plot the cite-seq information on top of the rna clusters
genes_to_plot = protein.var.index.tolist() + ['rna_leiden']
fig = sc.pl.embedding(protein, 'RNA_umap', color=genes_to_plot,
sort_order=True, ncols=4,return_fig=True, color_map='Blues', use_raw=True,
vmin='p5', vmax='p99.9', legend_fontsize=8)
fig.set_size_inches(12,12)
sf(fig,'3_RNA-umap_with_CITE-counts')
plt.show()
# and, let's plot some rna-seq information on top of the cite clusters
# I'll leave you to do this one
# sf(fig,'3_CITE-umap_with_RNA-counts')
```
### Now, let's plot RNA information against CITE information to see how they compare.
```
# first, get the metadata from the scanpy .obs dataframe
meta = gex.obs
meta.head()
# and add in the umap coordinates from the RNA
meta['umap_1'] = gex.obsm['X_umap'][:, 0]
meta['umap_2'] = gex.obsm['X_umap'][:, 1]
# now add in the umap coordinates from the CITE-seq
meta['umap-cite_1'] = protein[meta.index].obsm['X_umap'][:, 0]
meta['umap-cite_2'] = protein[meta.index].obsm['X_umap'][:, 1]
meta.head()
# here's two helper functions to get gene/protein expression information
def get_gene_expression(gene, adata=gex, undo_log=False, cells=''):
gene_ix = adata.raw.var.index.get_loc(gene)
vals = adata.raw.X[:, gene_ix].toarray().ravel()
if undo_log:
vals = np.exp(vals) - 1
vals = pd.Series(vals, index=adata.obs.index)
return vals
def get_protein_expression(gene, data=protein_norm_clr):
vals = protein_norm_clr.loc[gene]
return vals
# make a scatter plot of RNA expression vs CITE-seq counts
for gene in protein.var.index:
rna_vals = get_gene_expression(name_dict[gene])
protein_vals = get_protein_expression(gene)
sf(fig, '4_scatter_rna-cite_' + gene)
# plot the RNA and CITE counts on top of the UMAP from the RNA data
for gene in protein.var.index:
rna_vals = get_gene_expression(name_dict[gene])
protein_vals = get_protein_expression(gene)
fig = plt.figure(figsize=(10,5))
# plot RNA
# plot PROTEIN
sf(fig, '4_umap_rna-cite_' + gene)
```
| github_jupyter |
## Coding Basics for Researchers - Day 4
*Notebook by [Pedro V Hernandez Serrano](https://github.com/pedrohserrano)*
There are many ways and technologies to harness and manipulate data, from open source solutions to licensed software, from scientific software to GUI programs like Excel. Usually, there is no one-size-fits-all solution for data manipulation. The main advantage of GUI software is the short learning curve. However, the more operations you perform to the data, the more difficult it is to track the changes in the data, resulting in difficulties reproducing the research, either for yourself or for future researchers.

Part of the research process is to conduct data analysis to answer research questions. However, data can come in different shapes and forms. It can be observations, experimental or simulated. Moreover, most of the time, there is a need to transform the data to make sense of it and bring it to the level of analysis.
In the following notebook, we will explore why performing data manipulation, transformation and analysis with an open-source programming language, Python, can automate your research, making it reproducible and transparent according to the open science efforts. Furthermore, using jupyter notebooks, one can share and publish data analysis processes.

---
# 4. Data Analysis with Python
* [4.1. Data Simulation](#4.1)
* [4.2. Data Filtering and Aggregation](#4.2)
When we are doing data analysis, we normally will encounter the following type of data problems:
+ **Oh, I have to simulate some data (Data Simulation)**
+ **I'm not interested in the whole dataset, only one category (Data Filtering)**
+ **Well, this is observation level I need analysis level (Data Aggregation)**
+ **Oh, they gave me the dataset in panel format, and I can't do descriptive statistics (Data Melting)**
+ **Hmm, my observations are identifiers, and the patients are in another catalogue (Data Merging)**
+ **Survey data has more than one response per entry! (Data Expand)**
+ **My data comes in a format I've never seen! (e.g. JSON) (Data Mapping)**
---
## 4.1. Data Simulation
<a id="4.1">
[NumPy](https://numpy.org/) Is a fundamental package for scientific computing with Python including statistical methds for quantitative analysis.
### List
- A collection of elements in a vector, not nescesarrily of the same type.
- The elements in a list are indexed with 0 being the first index.
- A **List** is the standard ordered-sequence data structure in Python.
- No external package is needed to create and read **Lists**
A list is created simply by adding elements inside squared brakets`[ ]`
```
my_list = [11, 12, 33, 40.5]
print(my_list)
```
And it can contain numbers and strings (words)
```
weird_list = ['a','b','c','d',1,2,3,2,4]
print(weird_list)
```
Once a list is defined, we can access its elements using the indexing method `[ ]` (no space)
```
weird_list[2]
```
We can check that indeed some elements in the same list are not of the same type
```
print(type(weird_list[2]), type(weird_list[5]))
```
### Array
- A more efficient way to store an ordered set (vector) of elements of the same type.
- For efficient computation, NumPy arrays are far more suitable
- To use arrays is necessary to import the Numpy library, e.g., `import numpy as np`
An array is created using the `np.array()` function from NumPy libray (using the same vector)
**Note:** We need to use both parentheses and square brackets
```
import numpy as np
my_array = np.array([11, 12, 33])
print(my_array)
```
But introducing different types of data, things may change
```
np.array([11, 12, 33, "Billie Eilish"])
```
Do you see what happened? Python automatically typecast all of the elements in the array to be of the same type.
The main advantage is that one is able to make math operations easily
```
# 3 times each element of the array
3 * my_array
# substracting 10
my_array - 10
```
An array can be converted to a List by simply using the `list()` function
```
new_list = list(my_array)
print(new_list)
```
### Using SciPy to Generate Data
- The main library to perform Statistical analysis in Python is [SciPy (Scientific Python)](https://www.scipy.org/)
- SciPy builds on NumPy, and for all basic array handling needs you can use NumPy functions.
**Note:** SciPy library is huge, therefore, we might not want to import the whole library but the functions that we need in that moment, for example, to import the Normal distribution `from scipy.stats import norm`
#### Generating from normal distribution
- The location (`loc`) parameter specifies the `mean`. The scale (`scale`) parameter specifies the `standard deviation`.
- See [scipy.stats.norm](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.norm.html#scipy.stats.norm) for more details
The case can be *The mean daily temperature of Maastricht in April*". Let's suppose that we know the daily temperature in April is distributed Normal with `mean = 11.5` and `sd = 5.5` (i.e. between 6 and 17 Celsius on most of days).
For a `size` of 30 days:
```
from scipy.stats import norm
X = norm.rvs(loc=11.5, scale=5.5, size=30)
print(X)
```
To get an idea of whether or not the data we generated has a Normal dstribution, we can quickly see if it has the expected bell curve by plotting it.
```
import matplotlib.pyplot as plt
plt.hist(X)
```
**Note:**
By adding the parameter `random_state` we can reproduce the same generated values if the code is run again, random state is also called **seed**. This number can be anything you want (e.g the number of the beast)
```
norm.rvs(size=50, random_state=666)
```
#### Generating from a discrete distribution
- Poisson distribution takes as `shape` parameter `mu`. When $\mu = 0$ then method returns 1.0.
- See [scipy.stats.poisson](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.poisson.html#scipy.stats.poisson) for more details
The Poisson distribution is popular for modelling the number of times an event occurs in an interval of time. An example can be the "Number of goals scored in a Football match" *. Let's supose that we know `mu = 3` (i.e., 3 goals expected in total).
- We can generate random values that are representative of the real data, we just have to specify the size of the sample that we want.
With a `size` of 60 matches:
```
from scipy.stats import poisson
X = poisson.rvs(mu=3, size=30)
print(X)
```
#### Sampling Elements
- The main goal of simple random sampling is to have an unbiased representation of the total population.
**Note:** There are many other techniques for sampling depending on the experiment, however, we are focusing on simple random sampling using `random.sample()` from the Random library
Given a set Y of 10 consecutive numbers, we can sample 4 of them making sure that all the elements have the same chances
```
#import the library
import random
#this is just a vector represented as a list
Y = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
# use the function sample on the vector Y to select 3 elements
random.sample(Y, 4)
```
It is also possible to sample non numerical elements in a vector
```
# yet another vector
teams_spain = ["Real Madrid", "Villareal", "Barcelona", "Sevilla", "Atletico"]
random.sample(teams_spain, 2)
```
Let's create another vector and try to come up with a table that can simulate calendar matches
```
teams_england = ["M. City", "M. United", "Arsenal", "Liverpool", "Chelsea"]
```
Let's use a basic for loop to generate the match number
```
# getting 1 sample from teams_spain repeated 30 times
samples_spain = [random.sample(teams_spain, 1)[0] for i in range(30)]
# getting 1 sample from teams_england repeated 30 times
samples_england = [random.sample(teams_england, 1)[0] for i in range(30)]
```
Let's convert it into a table that we can export
```
import pandas as pd
table = {
'spanish_team': samples_spain,
'english_team': samples_england,
'match_score': X
}
df_matches = pd.DataFrame(table)
df_matches.head()
```
``` python
# In quotations is the name we WANT to give to the file
df_matches.to_excel('simulated_matches.xls')
```
---
## 4.2. Data Filtering and Aggregation
<a id="4.2">
* Load the library with `import pandas as pd`. The alias pd is commonly used for Pandas. (recommended)
* Read a Comma Separated Values (CSV) data file with `pd.read_csv`.
* We will use the Gapminder dataset
#### Data
We will be using the **[Gapminder example](https://www.gapminder.org/fw/world-health-chart/)** a dataset that is already contained in the Plotly library (how convenient!). The dataset contains information on countries' life expectancy, population and GDP per capita per year.
This dataset became famous since it has been constantly used for illustrating the power of data visualization, in different conferences, presentations, dashboards and a number of infographics.
Likewise, we will walk through different tasks using it.
Down here there is a video explaining the **Gapminder** data in an amazing real-life data visualization exercise (click play ►)
Here is a [link to the video](youtube.com/watch?v=jbkSRLYSojo) as well
#### Descriptions
- **country:** Name of the country **[Zimbabwe = ZWE]**
- **continent:** Name of the comntinent **[Europe]**
- **year:** year**
- **lifeExp:**
- **pop:**
- **gdpPercap:**
- **iso_alpha:**
- **iso_num:** Country three letters ISO official code **[ZWE = Zimbabwe]**
**Note:** Since Plotly Express is an external library, it is required to be imported (sometimes also installed) before executing. But no worries!! Installing and importing things in a programming environment is quite simple
```
import plotly.express as px
df = px.data.gapminder()
```
- Normally, as a convention, people use `df` as the standard name to call the data, df stands for DataFrame, indicating that one should not confuse this data structure with any other
- The columns in a dataframe are the observed variables, and the rows are the observations.
- You can simply display the data by executing the `df` object in a new cell
Use `df.head()` to get the first rows of the dataframe
```
df.head(3)
df.columns
```
Use `df.describe()` to get summary statistics about the data
- This method returns a new samaller dataframe with only the summary statistics of the columns that have numerical data. All other columns are ignored.
```
df.describe()
```
### Filtering by creating subsets with Pandas
- Using Pandas it is possible to select entire rows or entire columns from a dataframe.
- Also, select a subset of a dataframe by defined criterion.
- The DataFrame is the way Pandas represents a table
- Pandas is built on top of the NumPy library, which in practice means that most of the methods defined for Numpy Arrays apply to DataFrames.
We can get a slice of the data by using the `df.query()` function to select all the records relating to a specific year. For example, by specifying the corresponding column to filter (`year`) and the desired value (`2007`).
```
df.query('year == 2007')
```
**Note:** This was just a slice of the data, this new subset is not saved anywhere, in order to preserve it, we need to save it in an object. Let's define the `df_2007` object then.
```
df_2007 = df.query('year == 2007')
df_2007.head(2)
```
We can get yet another subset of the `df_2007` subset by using the `df.query()`. Let's filter now by a column that is not numerical, for example `continent` selecting `Europe`. Please note we are using double quotations now!
```
df_2007_europe = df_2007.query('continent == "Europe" ')
df_2007_europe.head(2)
```
Use `DataFrame.sort_values()` to order the values
- This function expects a parameter to indicate which column you want to order by
- The second parameter indicates ascending or descending order (when ascending is `False` it's the same as descending)
- **Note:** This time we are not using `df` anymore since now we are interested only in the European subset
```
df_2007_europe.sort_values(by='pop', ascending=False)
```
This last table looks quite neat, the following command will make it even better, by assigning the country as the index (the row names) of the dataframe
```
# european 2007 dataset sets the country as index and applies it to the current dataframe with inplace=True (with inplace=False a new dataframe is returned)
df_2007_europe.set_index('country', inplace=True )
```
Use `DataFrame.loc[ , ]` to select values by their (entry) label. e.g. What is the `Netherlands` population `pop`?. First the index of the row and then the column name
```
df_2007_europe.loc['Netherlands','pop']
```
Finally, it is possible to create a subset based on a condition, e.g. "Selecting all the countries with a life expectancy above 80 years" We save it as the variable `final_subset`
```
# Subset based on logical condition (>)
final_subset = df_2007_europe.query('lifeExp > 80')
# display it
final_subset
```
**IMPORTANT**
- Having this last subset we have created: "Pop and GDP of European countries in 2007 with Life expectancy above 80"
- We can now save it in a real file, this can be CSV or Excel
- To handle Excel files an aditional library needs to be installed `xlrd`
Use `DataFrame.to_csv()` to save the file in the same directory your notebook is located
### Aggregate the data for analysis
```
# Filter the Africa region
df_Africa = df.query('continent == "Africa"')
df_Africa.head(3)
df_Africa_table = df_Africa\
.groupby('country')\
.mean()['lifeExp']\
.reset_index()
df_Africa_table.head(3)
df_Africa_years = df_Africa\
.groupby('year')\
.mean()['lifeExp']\
.reset_index()
df_Africa_years.head(3)
summary_Africa = pd.pivot_table(df_Africa, values='lifeExp', columns='year', index='country', aggfunc='mean')
summary_Africa.head(3)
```
``` python
# In quotations is the name we WANT to give to the file
summary_Africa.to_excel('africa_summary_lifeExp.xls')
```
---
## [Seaborn](https://seaborn.pydata.org/)
is a Python data visualization library based on matplotlib. It provides a high-level interface for drawing attractive and informative statistical graphics.
```
# import seaborn package
import seaborn as sns
```
#### Scatterplot:
- A scatterplot displays the relationship between 2 numeric variables.
- For each data point, the value of its first variable is represented on the X axis, the second on the Y axis.
#### What for?
A scatterplot is made to visually look at the relationship between 2 variables. It is often accompanied by a regression line that tries to capture the *linear relationship*.
```
# Create a visualization
sns.regplot(data=df_Africa, x='lifeExp', y='gdpPercap')
```
#### Lineplot:
- Seaborn allows you to easily put a dataframe as input and it will recognize the categoriez you want to plot.
#### What for?
A lineplot intends to show a linera pattern over the axis X, time is often commin.
```
# Making a list of the countries I want to filter
north_africa = ['Algeria','Morocco','Tunisia','Libya','Egypt']
# Create a filtered subset including only the countries of my list
subset_north_africa = summary_Africa[summary_Africa.index.isin(north_africa)].T
# input my subset in the lineplot function of Seaborn
sns.lineplot(data = subset_north_africa)
```
---
## EXERCISES
+ _1. When we print `my_list` and `my_array` it appears to produce the same result. How would you check the type of data structure for each one?_
```python
my_list = [11, 12, 33]
print(my_list)
my_array = np.array([11, 12, 33])
print(my_array)
```
___
+ _2. Let's say that I want to import the whole stats library and then use the Uniform distribution (which is part of the stats library)_
```python
import scipy.stats as stats
```
Why is the following function not working? Make the appropiate changes to fix the problem.
```python
uniform.rvs(size=100)
```
___
+ _3. Given the following vector in an array form. Explore why it cannot be sampled. How do you fix this?_
```python
V = np.array([0, 1, 2, 3, 4, 5])
random.sample(V, 2)
```
___
+ _6. Create a list with 5 elements of your wish. Then write a code that will take a sample corresponding to the 20% total amount of elements
Hint: you can use `len()` function
___
+ _7. Read the gapminder data again. Which one of the following query functions is going to work and why?_
```python
df.query('year == "1992" ')
df.query('year == 1992 ')
df.query('year = "1992" ')
```
___
+ _8. Select the column `continent` from the `df` dataframe with the command `df['continent']`, and call it `my_column`_
_What would be the purpose of the following function?:_
```python
my_column.unique()
```
___
+ _9. We did a Filter of gapminder by creating a subset `Europe 2007`, create another data Filter for `Europe` but this time for the year `1952`._
_What is the average life expectancy for 1952 and 2007? How much has changed?_
___
+ _10. We did an Aggregation of gapminder by pivoting `african countries` and `years`. Create another data Aggregation but now for `Asia`._
___
+ _11. Reproduce the Lineplot example, this time you will select Japan, China, North Korea and South Korea
Comment the results
._
___
| github_jupyter |
```
import ctypes
import locale
import os
import platform
from ctypes.util import find_library
import cffi
from PIL import Image, ImageDraw, ImageFont
# matplotlib&numpy are needed only for displying in jupyter.
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
def get_abs_path_of_library(library):
"""Get absolute path of library."""
abs_path = None
lib_name = find_library(library)
if os.path.exists(lib_name):
abs_path = os.path.abspath(lib_name)
return abs_path
libdl = ctypes.CDLL(lib_name)
if not libdl:
return abs_path # None
try:
dlinfo = libdl.dlinfos
except AttributeError as err:
# Workaroung for linux
abs_path = str(err).split(":")[0]
return abs_path
ffi = cffi.FFI()
ffi.cdef("""
typedef signed char l_int8;
typedef unsigned char l_uint8;
typedef short l_int16;
typedef unsigned short l_uint16;
typedef int l_int32;
typedef unsigned int l_uint32;
typedef float l_float32;
typedef double l_float64;
typedef long long l_int64;
typedef unsigned long long l_uint64;
typedef int l_ok; /*!< return type 0 if OK, 1 on error */
struct Pix;
typedef struct Pix PIX;
typedef enum lept_img_format {
IFF_UNKNOWN = 0,
IFF_BMP = 1,
IFF_JFIF_JPEG = 2,
IFF_PNG = 3,
IFF_TIFF = 4,
IFF_TIFF_PACKBITS = 5,
IFF_TIFF_RLE = 6,
IFF_TIFF_G3 = 7,
IFF_TIFF_G4 = 8,
IFF_TIFF_LZW = 9,
IFF_TIFF_ZIP = 10,
IFF_PNM = 11,
IFF_PS = 12,
IFF_GIF = 13,
IFF_JP2 = 14,
IFF_WEBP = 15,
IFF_LPDF = 16,
IFF_TIFF_JPEG = 17,
IFF_DEFAULT = 18,
IFF_SPIX = 19
};
char * getLeptonicaVersion ( );
PIX * pixRead ( const char *filename );
PIX * pixCreate ( int width, int height, int depth );
PIX * pixEndianByteSwapNew(PIX *pixs);
l_int32 pixSetData ( PIX *pix, l_uint32 *data );
l_ok pixSetPixel ( PIX *pix, l_int32 x, l_int32 y, l_uint32 val );
l_ok pixWrite ( const char *fname, PIX *pix, l_int32 format );
l_int32 pixFindSkew ( PIX *pixs, l_float32 *pangle, l_float32 *pconf );
PIX * pixDeskew ( PIX *pixs, l_int32 redsearch );
void pixDestroy ( PIX **ppix );
l_ok pixGetResolution ( const PIX *pix, l_int32 *pxres, l_int32 *pyres );
l_ok pixSetResolution ( PIX *pix, l_int32 xres, l_int32 yres );
l_int32 pixGetWidth ( const PIX *pix );
typedef struct TessBaseAPI TessBaseAPI;
typedef struct ETEXT_DESC ETEXT_DESC;
typedef struct TessPageIterator TessPageIterator;
typedef struct TessResultIterator TessResultIterator;
typedef int BOOL;
typedef enum TessOcrEngineMode {
OEM_TESSERACT_ONLY = 0,
OEM_LSTM_ONLY = 1,
OEM_TESSERACT_LSTM_COMBINED = 2,
OEM_DEFAULT = 3} TessOcrEngineMode;
typedef enum TessPageSegMode {
PSM_OSD_ONLY = 0,
PSM_AUTO_OSD = 1,
PSM_AUTO_ONLY = 2,
PSM_AUTO = 3,
PSM_SINGLE_COLUMN = 4,
PSM_SINGLE_BLOCK_VERT_TEXT = 5,
PSM_SINGLE_BLOCK = 6,
PSM_SINGLE_LINE = 7,
PSM_SINGLE_WORD = 8,
PSM_CIRCLE_WORD = 9,
PSM_SINGLE_CHAR = 10,
PSM_SPARSE_TEXT = 11,
PSM_SPARSE_TEXT_OSD = 12,
PSM_COUNT = 13} TessPageSegMode;
typedef enum TessPageIteratorLevel {
RIL_BLOCK = 0,
RIL_PARA = 1,
RIL_TEXTLINE = 2,
RIL_WORD = 3,
RIL_SYMBOL = 4} TessPageIteratorLevel;
TessPageIterator* TessBaseAPIAnalyseLayout(TessBaseAPI* handle);
TessPageIterator* TessResultIteratorGetPageIterator(TessResultIterator* handle);
BOOL TessPageIteratorNext(TessPageIterator* handle, TessPageIteratorLevel level);
BOOL TessPageIteratorBoundingBox(const TessPageIterator* handle, TessPageIteratorLevel level,
int* left, int* top, int* right, int* bottom);
const char* TessVersion();
TessBaseAPI* TessBaseAPICreate();
int TessBaseAPIInit3(TessBaseAPI* handle, const char* datapath, const char* language);
int TessBaseAPIInit2(TessBaseAPI* handle, const char* datapath, const char* language, TessOcrEngineMode oem);
void TessBaseAPISetPageSegMode(TessBaseAPI* handle, TessPageSegMode mode);
void TessBaseAPISetImage(TessBaseAPI* handle,
const unsigned char* imagedata, int width, int height,
int bytes_per_pixel, int bytes_per_line);
void TessBaseAPISetImage2(TessBaseAPI* handle, struct Pix* pix);
BOOL TessBaseAPISetVariable(TessBaseAPI* handle, const char* name, const char* value);
BOOL TessBaseAPIDetectOrientationScript(TessBaseAPI* handle, char** best_script_name,
int* best_orientation_deg, float* script_confidence,
float* orientation_confidence);
int TessBaseAPIRecognize(TessBaseAPI* handle, ETEXT_DESC* monitor);
TessResultIterator* TessBaseAPIGetIterator(TessBaseAPI* handle);
BOOL TessResultIteratorNext(TessResultIterator* handle, TessPageIteratorLevel level);
char* TessResultIteratorGetUTF8Text(const TessResultIterator* handle, TessPageIteratorLevel level);
float TessResultIteratorConfidence(const TessResultIterator* handle, TessPageIteratorLevel level);
char* TessBaseAPIGetUTF8Text(TessBaseAPI* handle);
const char* TessResultIteratorWordFontAttributes(const TessResultIterator* handle, BOOL* is_bold, BOOL* is_italic,
BOOL* is_underlined, BOOL* is_monospace, BOOL* is_serif,
BOOL* is_smallcaps, int* pointsize, int* font_id);
void TessBaseAPIEnd(TessBaseAPI* handle);
void TessBaseAPIDelete(TessBaseAPI* handle);
""")
def pil2PIX32(im, leptonica):
"""Convert PIL to leptonica PIX."""
# At the moment we handle everything as RGBA image
if im.mode != "RGBA":
im = im.convert("RGBA")
depth = 32
width, height = im.size
data = im.tobytes("raw", "RGBA")
pixs = leptonica.pixCreate(width, height, depth)
leptonica.pixSetData(pixs, ffi.from_buffer("l_uint32[]", data))
try:
resolutionX = im.info['resolution'][0]
resolutionY = im.info['resolution'][1]
leptonica.pixSetResolution(pixs, resolutionX, resolutionY)
except KeyError:
pass
try:
resolutionX = im.info['dpi'][0]
resolutionY = im.info['dpi'][1]
leptonica.pixSetResolution(pixs, resolutionX, resolutionY)
except KeyError:
pass
return leptonica.pixEndianByteSwapNew(pixs)
# Setup path and library names
architecture = platform.architecture()
dll_dir = ""
if platform.architecture()[1].lower().startswith('windows'):
dll_dir = "win"
if platform.architecture()[0] == '64bit':
dll_dir += "64"
elif platform.architecture()[0] == '32bit':
dll_dir += "32"
abs_path = os.path.join(os.getcwd(), dll_dir)
env_path = os.environ['PATH']
if abs_path not in env_path:
os.environ['PATH'] = abs_path + ";" + env_path
tess_libname = os.path.join(abs_path, "tesseract41.dll")
lept_libname = os.path.join(abs_path, "leptonica-1.78.0.dll")
else:
tess_libname = get_abs_path_of_library('tesseract')
lept_libname = get_abs_path_of_library('lept')
# Use project tessdata
tessdata = os.path.join(os.getcwd(), "tessdata")
os.environ['TESSDATA_PREFIX'] = tessdata
# Load libraries in ABI mode
if os.path.exists(tess_libname):
tesseract = ffi.dlopen(tess_libname)
else:
print(f"'{tess_libname}' does not exists!")
tesseract_version = ffi.string(tesseract.TessVersion())
print('Tesseract-ocr version', tesseract_version.decode('utf-8'))
if os.path.exists(lept_libname):
leptonica = ffi.dlopen(lept_libname)
else:
print(f"'{lept_libname}' does not exists!")
leptonica_version = ffi.string(leptonica.getLeptonicaVersion())
print(leptonica_version.decode('utf-8'))
api = None
# Load image with PIL and convert it to leptonica PIX
filename = r"test.jpg"
im = Image.open(filename)
pix = pil2PIX32(im, leptonica)
# Get information about DPI
x_dpi = ffi.new('int *')
y_dpi = ffi.new('int *')
leptonica.pixGetResolution(pix, x_dpi, y_dpi)
print(f"Image {filename} has {x_dpi[0]}x{y_dpi[0]} DPI.")
# Create tesseract API
if api:
tesseract.TessBaseAPIEnd(api)
tesseract.TessBaseAPIDelete(api)
api = tesseract.TessBaseAPICreate()
# Parameters fro API initialization
# use xz compressed traineddata file - feature is available in recent github code
lang = "eng_xz"
# OEM_DEFAULT OEM_LSTM_ONLY OEM_TESSERACT_ONLY OEM_TESSERACT_LSTM_COMBINED
oem = tesseract.OEM_DEFAULT
# Initialize API, set image and regonize it
tesseract.TessBaseAPIInit2(api, tessdata.encode(), lang.encode(), oem)
# PSM (Page segmentation mode):
# PSM_OSD_ONLY, PSM_AUTO_OSD, PSM_AUTO_ONLY, PSM_AUTO, PSM_SINGLE_COLUMN,
# PSM_SINGLE_BLOCK_VERT_TEXT, PSM_SINGLE_BLOCK, PSM_SINGLE_LINE,
# PSM_SINGLE_WORD, PSM_CIRCLE_WORD, PSM_SINGLE_CHAR, PSM_SPARSE_TEXT,
# PSM_SPARSE_TEXT_OSD
tesseract.TessBaseAPISetPageSegMode(api, tesseract.PSM_AUTO)
tesseract.TessBaseAPISetImage2(api, pix)
tesseract.TessBaseAPIRecognize(
api, ffi.NULL) # recognize is needed to get result iterator
# Print whole recognized text
utf8_text = ffi.string(tesseract.TessBaseAPIGetUTF8Text(api)).decode('utf-8')
print(utf8_text)
# Use result iterator to get confidence and bouding box for words
idx = 0
results = []
left = ffi.new('int *')
top = ffi.new('int *')
right = ffi.new('int *')
bottom = ffi.new('int *')
level = tesseract.RIL_WORD # RIL_SYMBOL RIL_WORD RIL_TEXTLINE RIL_PARA RIL_BLOCK
result_iterator = tesseract.TessBaseAPIGetIterator(api)
while 1:
page_iterator = tesseract.TessResultIteratorGetPageIterator(
result_iterator)
tesseract.TessPageIteratorBoundingBox(page_iterator, level, left, top,
right, bottom)
text = ffi.string(
tesseract.TessResultIteratorGetUTF8Text(result_iterator,
level)).decode('utf-8')
conf = tesseract.TessResultIteratorConfidence(result_iterator, level)
print(
f"""{idx} confidence: {conf} - [{left[0]}, {top[0]}, {right[0]}, {bottom[0]}]; {text}"""
)
idx = idx + 1
results.append((left[0], top[0], right[0], bottom[0], text, conf))
if not tesseract.TessResultIteratorNext(result_iterator, level):
break
# Display Bound boxes with result text
result_img = None
tmp = Image.new('RGBA', im.size, (0,0,0,0))
font_size = 26
conf_size = int(font_size*.90)
text_font = ImageFont.truetype('Roboto-Black.ttf', font_size)
conf_font = ImageFont.truetype('Roboto-Black.ttf', conf_size)
padding = 0
box_color = (255, 255, 255, 150)
text_color = (255, 0, 0, 255)
conf_color = (255, 0, 255, 255)
draw = ImageDraw.Draw(tmp)
for (startX, startY, endX, endY, text, conf) in results:
draw.rectangle(((startX, startY), (endX, endY)), fill=box_color)
draw.text((startX + padding, startY + padding),
text,
font=text_font,
fill=text_color)
cont_text = f"{conf:.3}"
y_shifted = startY + padding + conf_font.getsize(cont_text)[1]
if y_shifted > tmp.size[1]*0.9:
y_shifted = tmp.size[1] - text_font.getsize(text)[1] - padding - conf_font.getsize(cont_text)[1]
print(y_shifted, tmp.size[1])
draw.text((startX + padding, y_shifted),
cont_text,
font=conf_font,
fill=conf_color)
result_img = Image.alpha_composite(im.convert("RGBA"), tmp)
# If you do not have/want to use matplotlib&numpy uncomment folowing line
# result_img.show()
# To save this image you can use:
result_img.save("BoundingBox.png", "PNG")
# display bounding boxes and recognized text with matplot&numpy
plt.rcParams['figure.dpi'] = 300 # dpi of displayed plot
f = plt.figure()
f.add_subplot(1, 2, 1)
plt.imshow(np.asarray(im))
f.add_subplot(1, 2, 2)
plt.imshow(np.asarray(result_img))
plt.show(block=True)
# Delete api and pix
if api:
tesseract.TessBaseAPIEnd(api)
tesseract.TessBaseAPIDelete(api)
result = ffi.new('PIX**')
result[0] = pix
leptonica.pixDestroy(result)
del pix
del result
api = None
# Store results to csv file
import csv
with open('result.csv', 'wt', encoding='utf-8', newline='\n') as csvFile:
writer = csv.writer(csvFile)
writer.writerows(results)
csvFile.close()
```
| github_jupyter |
# Selection Algorithm
```
import numpy as np
from mercs.algo.selection import _set_missing, _ensure_desc_atts, _set_nb_targets
TARG_ENCODING = 1
def _nb_models_and_deficit(nb_targets, potential_targets):
nb_potential_targets = potential_targets.shape[0]
nb_models_with_regular_nb_targets = nb_potential_targets // nb_targets
nb_leftover_targets = nb_potential_targets % nb_targets
if nb_leftover_targets:
nb_models = nb_models_with_regular_nb_targets + 1
deficit = nb_targets - nb_leftover_targets
else:
nb_models = nb_models_with_regular_nb_targets
deficit = 0
return nb_models, deficit
def _init(nb_models, nb_attributes):
return np.zeros((nb_models, nb_attributes), dtype=int)
def _target_sets(potential_targets, nb_targets, nb_models, deficit):
np.random.shuffle(potential_targets)
choices = np.r_[potential_targets, potential_targets[:deficit]]
return np.random.choice(choices, replace=False, size=(nb_models, nb_targets))
def _set_targets(m_codes, target_sets):
row_idx = np.arange(m_codes.shape[0]).reshape(-1,1)
col_idx = target_sets
m_codes[row_idx, col_idx] = TARG_ENCODING
return m_codes
def _single_iteration_random_selection(nb_attributes, nb_targets, fraction_missing, potential_targets):
nb_models, deficit = _nb_models_and_deficit(nb_targets, potential_targets)
# Init
m_codes = _init(nb_models, nb_attributes)
target_sets = _target_sets(potential_targets, nb_targets, nb_models, deficit)
m_codes = _set_targets(m_codes, target_sets)
m_codes = _set_missing(m_codes, fraction_missing)
m_codes.astype(int)
return m_codes
def base_selection_algorithm(metadata, nb_targets=1, nb_iterations=1, random_state=997):
m_codes = random_selection_algorithm(metadata, nb_targets=nb_targets, nb_iterations=nb_iterations, fraction_missing=0., random_state=random_state)
return m_codes
def random_selection_algorithm(metadata, nb_targets=1, nb_iterations=1, fraction_missing=0.2, random_state=997):
# Init
np.random.seed(random_state)
nb_attributes = metadata["n_attributes"]
nb_targets = _set_nb_targets(nb_targets, nb_attributes)
codes = []
for attribute_kind in {'nominal_attributes', 'numeric_attributes'}:
potential_targets = np.array(list(metadata[attribute_kind]))
for iterations in range(nb_iterations):
m_codes = _single_iteration_random_selection(nb_attributes, nb_targets, fraction_missing, potential_targets)
codes.append(m_codes)
m_codes = np.vstack(codes)
m_codes = ensure_desc_atts(m_codes)
return m_codes
metadata
metadata={'n_attributes': 10,
'nominal_attributes': {0,1,2,3,4,5,6},
'numeric_attributes': {7, 8, 9}}
m_codes = base_selection_algorithm(metadata, nb_targets=2, nb_iterations=1, random_state=3)
m_codes
np.where(m_codes==1)[1]
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import time
from scipy.stats import multivariate_normal
class VAE(tf.keras.Model):
"""Class of basic Variational Autoencoder
This class contains basic components and main functions of VAE model.
Args:
latent_dim: size of latent variables. 2,4,6 ... 2n
input_shape: shape of input data. [28,28]...
"""
def __init__(self, latent_dim,input_shape):
super(VAE, self).__init__()
self.latent_dim = latent_dim
# Encoder NN
self.encoder = tf.keras.Sequential(
[
tf.keras.layers.InputLayer(input_shape=input_shape),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(latent_dim + latent_dim),
]
)
# Decoder NN
self.decoder = tf.keras.Sequential(
[
tf.keras.layers.InputLayer(input_shape=(latent_dim)),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(784),
tf.keras.layers.Reshape(target_shape=input_shape),
]
)
@tf.function
def sample(self, eps=None):
""" sample data from latent variables
Args:
eps: it will be used for decode and
eps will be set by a random data when None
Return:
log x'(the reconstruction data of x)
"""
if eps is None:
eps = tf.random.normal(shape=(100, self.latent_dim))
return self.decode(eps, apply_sigmoid=True)
def encode(self, x):
"""do encode jobs
for each input data x, will construct multiple Gausian Distribution
Args:
x: data that will be encoded.
Return:
mean: array like. list of mu of that distribution.
logvar: array like. list of logvar of that distribution.
"""
mean, logvar = tf.split(self.encoder(x), num_or_size_splits=2, axis=1)
return mean, logvar
def reparameterize(self, mean, logvar):
""" reparameterize trick """
eps = tf.random.normal(shape=mean.shape)
return eps * tf.exp(logvar * .5) + mean
# decode
def decode(self, z, apply_sigmoid=False):
""" do decode job
Args:
z: latent variables.
apply_sigmoid: use sigmoid or not.
Return:
log x'(the reconstruction data of x)
"""
logits = self.decoder(z)
if apply_sigmoid:
probs = tf.sigmoid(logits)
return probs
return logits
def log_normal_pdf(sample, mean, logvar, raxis=1):
"""Log of the normal probability density function.
Args:
sample: sample that need compute pdf
mean,logvar: parametes of distribution
Return:
log pdf
"""
log2pi = tf.math.log(2. * np.pi)
return tf.reduce_sum(
-.5 * ((sample - mean) ** 2. * tf.exp(-logvar) + logvar + log2pi),
axis=raxis)
def compute_loss(model, x):
"""calc ELBO
Args:
x: input data
Return:
loss
"""
# 1. encode process
mean, logvar = model.encode(x)
# 2. reparameterize
z = model.reparameterize(mean, logvar)
# 3. get log x'
x_logit = model.decode(z)
# 4. Compute sigmoid cross entropy given `logits`.
cross_ent = tf.nn.sigmoid_cross_entropy_with_logits(logits=x_logit, labels=x)
# 5. Compute log p(x|z),log p(z),log q(z|x)
logpx_z = -tf.reduce_sum(cross_ent, axis=[1, 2, 3])
logpz = log_normal_pdf(z, 0., 0.)
logqz_x = log_normal_pdf(z, mean, logvar)
# 5. ELBO
return -tf.reduce_mean(logpx_z + logpz - logqz_x)
# optimizers
optimizer = tf.keras.optimizers.Adam(1e-4)
@tf.function
def train_step(model, x, optimizer):
"""Executes one training step and returns the loss.
This function computes the loss and gradients, and uses the latter to
update the model's parameters.
"""
with tf.GradientTape() as tape:
loss = compute_loss(model, x)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
def reconstructed_probability(X):
"""Get the reconstruction probability for X
Args:
X: test data. X.shape = (32,28,28,1)
Return:
log of probability
"""
reconstructed_prob = np.zeros(X.shape[0], dtype='float32')
# 1. encode,
mu_hat, log_sigma_hat = model.encode(X)
sigma_hat = tf.exp(log_sigma_hat)+0.0001
# for each piece of data
for j in range(X.shape[0]):
p_l = multivariate_normal.logpdf(X[j], mu_hat[j,:], np.diag(sigma_hat[j,:]))
reconstructed_prob[j] += tf.reduce_sum(p_l)
return reconstructed_prob
def judege_anomaly(scores, threshold):
""" judge anomaly
if scores[i] > threshold:
label for scores[i]: 1
Args:
scores: Array like.
threshold: float
Return:
labels: Array like.
"""
labels = np.zeros(len(scores),dtype=int)
for i in range(len(scores)):
if (scores[i]>threshold):
labels[i] = 1
return labels
model = VAE(latent_dim=4,input_shape=(28,28,1))
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.mnist.load_data()
# preprocess image data
def preprocess_images(images):
images = images.reshape((images.shape[0], 28, 28, 1)) / 255.
return np.where(images > .5, 1.0, 0.0).astype('float32')
# exclude error num
def exclude_num(x, y, num):
keep = (y != num)
x = x[keep]
return x
# label anomaly
def label_anomaly(anomaly_num,labels):
true_labels = np.zeros(len(labels),dtype=int)
for i in range(len(labels)):
if (labels[i] == anomaly_num):
true_labels[i] = 1
return true_labels
train_images = preprocess_images(train_images)
test_images = preprocess_images(test_images)
print("Before exclusion, Train_images.shape:",train_images.shape)
#choose error num
anomaly_num = 8
train_images = exclude_num(train_images,train_labels,anomaly_num)
# get true labels
true_labels = label_anomaly(anomaly_num, test_labels)
print("Take num {} as error data.".format(anomaly_num))
print("After exclusion, Train_images.shape:",train_images.shape)
# get validataion data from train_images
validation_size = int(train_images.shape[0]*0.3)
train_data = train_images[:-validation_size]
validation_data = train_images[-validation_size:]
# test_data
# test_dataset = test_images
train_size = 60000
batch_size = 32
test_size = 10000
train_dataset = (tf.data.Dataset.from_tensor_slices(train_data)
.shuffle(train_size).batch(batch_size))
validation_dataset = (tf.data.Dataset.from_tensor_slices(validation_data)
.shuffle(train_size).batch(batch_size))
# for test data, do not shuffle
test_dataset = (tf.data.Dataset.from_tensor_slices(test_images)
.batch(batch_size))
epochs = 20
# training
for epoch in range(1, epochs + 1):
# training
start_time = time.time()
for train_x in train_dataset:
train_step(model, train_x, optimizer)
end_time = time.time()
# compute loss
loss = tf.keras.metrics.Mean()
for validation_x in validation_dataset:
loss(compute_loss(model, validation_x))
elbo = -loss.result()
print('Epoch: {}, Test set ELBO: {}, time elapse for current epoch: {}'
.format(epoch, elbo, end_time - start_time))
# def judege_anomaly(scores,threshold):
# labels = np.zeros(len(scores),dtype=int)
# for i in range(len(scores)):
# if (scores[i]>threshold):
# labels[i] = 1
# return labels
# true_labels = np.zeros(len(test_labels),dtype=int)
# for i in range(len(test_labels)):
# if (test_labels[i] == error_num):
# true_labels[i] = 1
# true_labels
from tqdm import tqdm
from time import sleep
predict = []
with tqdm(total=len(test_dataset)) as pbar:
for test_data in test_dataset:
results = reconstructed_probability(X=test_data)
pre_labels = judege_anomaly(results,-31*1000)
predict.extend(pre_labels)
pbar.update(1)
from sklearn.metrics import accuracy_score
accuracy_score(predict,true_labels)
count/len(true_labels)
```
| github_jupyter |
# <center>Python Basics<center/>
<img height="60" width="120" src="https://www.python.org/static/img/python-logo-large.png?1414305901"></img>
# Table of contents
<br/>
<a href = "#6.-Variables">6. Variables</a><br/>
<a href = "#7.-Data-Types">7. Data Types</a><br/>
<a href = "#8.-Conversion-between-Datatypes">8. Conversion between Datatypes</a>
# 6. Variables
A variable is an alias given to a location in memory that is used to store some data (value).
Each data point stored in the memory is given unique name to differentiate between different memory locations (and thus data points). <br/><b>The rules for writing a variable name is same as the rules for writing identifiers in Python.</b>
We don't need to declare a variable before using it. In Python, we simply assign a value to a variable and it will exist. We don't even have to declare the <b>type</b> of the variable. This is handled internally according to the type of value we assign to the variable.
<br/>
<br/>
### Variable assignements
We use the assignment operator (=) to assign values to a variable
```
intergerNumber = 4
decimalNumber = 22.2
stringVariable = "StringValue"
print(intergerNumber, decimalNumber, stringVariable)
```
### Multiple assignements
Use comma to seperate the variables and their corresponding values
```
intergerNumber, decimalNumber, stringVariable = 4, 22.2, "StringValue"
print(intergerNumber, decimalNumber, stringVariable)
```
To initialize multiple variables with same value use 1 assignment
```
variable1=variable2=variable3 = " All variable assigned same value "
print(variable1,variable2,variable3)
```
### Storage Locations
```
print(id(intergerNumber)) #print address of variable: 'intergerNumber'
print(id(decimalNumber)) #print address of variable: 'decimalNumber'
```
<b>interger1</b> and <b>interger2</b> points to same memory location in the below example
```
interger1 = 10
interger2 = 10
print(id(interger1))
print(id(interger2))
```
# 7. Data Types
Every value in Python has a datatype. Since everything is an object in Python programming, data types are actually classes and variables are instance (object) of these classes.
<br/><br/>
Below are few of the commonly used data types<br/>
### Python Numbers
<b>Integers, floating point numbers and complex numbers</b>
falls under Python numbers category. They are defined as int, float and complex class in Python.
We can use the <b>type()</b> function to know which class a variable or a value belongs to <br/> and the <b>isinstance()</b> function to check if an object belongs to a particular class.
```
data1 = 12 # integer data type
print(data1, " is of type", type(data1))
data2 = 12.5 # float data type
print(data2, " is of type", type(data2))
data3 = 100 + 20j #data type is changed to complex number
print(data3, " is complex number?", isinstance(data3, complex))
```
### Boolean
Boolean represents True or False values
```
booleanVariable = True # 'booleanVariable' is a boolean type & True is a keyword
print(type(booleanVariable))
```
### Python Strings
String is sequence of Unicode characters. We can use single quotes or double quotes to represent strings. <br/>
Multi-line strings can be denoted using triple quotes(single/double), <b>''' or """. </b><br/>
A string in Python consists of a series or <u>sequence of characters</u> - <b>letters, numbers, and special characters.</b>
Strings can be indexed - often synonymously called subscripted as well.
The first character of a string has the index 0.
```
stringVariable = "This is a string variable"
print('stringVariable is : ',stringVariable)
print('First character in stringVariable is : ',stringVariable[0])
# Slicing of String variable
print(stringVariable[0:7])
print(stringVariable[:7])
print(stringVariable[5:])
```
### Python List
List is an <u>ordered sequence</u> of items. It is one of the most used datatype in Python and is very flexible.
All the items in a list do not need to be of the same type.
Declaring a list is pretty straight forward. Items separated by commas are enclosed within square brackets <b>'[ ]'</b>.
```
myList = [1, 2.2, "ThirdElement",12,24,"More values"]
print(myList)
print(myList[2]) # Print an element based on its index. Index starts from 0
```
Lists are mutable, which means, value of elements of a list can be changed.
```
myList[2] = "ElementThree"
print(myList)
myList.remove(2.2) # Remove item from a particular index
print(myList)
myList.pop(2) # Remove item from a particular index
print(myList)
myList.append(2.2) # Add item at the last index
print(myList)
```
### Python Tuple
Tuple is an <u>ordered sequence</u> of items same as list. <br/>
The only difference is that tuples are <u>immutable</u>. <b>Tuples once created cannot be modified</b>.
Tuples are used to write-protect data and are usually faster than list as it cannot change dynamically.
It is defined within parentheses <b>()</b> where items are separated by commas.
```
myTuple = (10,20,30,"Text")
print(myTuple[2]) # Read elements by their corresponding index values
myTuple[2] = 120
```
### Python Set
Set is an <u>unordered collection</u> of unique items. <br/>
Set is defined by values separated by comma inside curly braces <b>{ }</b>. <br/>
```
mySet = {10, 20, 30, 40, 50}
print(mySet)
mySet = {10, 20, 20, 30, 30} # Only unique values considered, duplicates removed automatically
print(mySet)
print(type(mySet)) # Check the type of the variable
print(mySet[0]) # Cannot print any particular element in a set because
# it's an unorder collections of items
```
### Python Dictionary
<b>Dictionary</b> is an <u>unordered collection</u> of <b>key-value</b> pairs.
It is generally used when we have a huge amount of data.<br/>
Dictionaries are optimized for retrieving data. <br/>
We must know the <b>key</b> to retrieve the <b>value</b>.
In Python, dictionaries are defined within curly braces <b>{}</b> <br/>
with each item being a pair in the form <b>key:value</b>. <br/>
<b>Key and value</b> can be of <u>any type</u>.
```
myDictionary = {'key1': "value1", 'key2': "value2", 'key3': "value3",}
print(myDictionary['key2'])
```
<b>Update</b> a value in the dictionary
```
myDictionary['key2'] = "value2.2"
print(myDictionary)
```
<b>Add</b> a new key & value
```
myDictionary["key4"] = "value4"
print(myDictionary)
```
<b>Delete</b> Dictionary Elements
```
del myDictionary["key2"]
print(myDictionary)
```
# 8. Conversion between Datatypes
To convert between different data types use different type conversion functions like <b>int(), float(), str()</b> etc.
```
int(5.5)
float(5)
```
Conversion to and from string must contain compatible values.
```
str(5)
str(5.5)
int('5')
int('5.5')
float('5.5')
float('5')
int('CharIn10')
myNewString = 'text1 '+ 20 + ' text2'
myNewString = 'text1 '+ str(20) + ' text2'
print(myNewString)
```
Convert,<br/>
1> list to set<br/>
2> String to list
```
# 1> list to set
myList1 = ['a','b','c','c']
print(myList1)
print(type(myList1))
mySet1 = set(myList1)
print(type(mySet1))
print(mySet1)
# 2> String to list
myString1 = 'This is string to list'
print(type(myString1))
strToList = list(myString1)
print(type(strToList))
print(strToList)
```
| github_jupyter |
# Logistic Regression, Gradient Descent
This notebook will show a few things.
* HOWTO simulate data for testing logistic regression
* HOWTO solve a logistic regression problem using [Autograd](https://github.com/HIPS/autograd).
## Simulate data
Here, we simulate data according to the following equation.
$\log \frac{p}{1-p} = 1.0 + 2.0 * x_1 + 3.0 * x_2$
It's easier to understand the simulation if we rewrite the equation to look like the following.
$p = \frac{ 1 }{ 1 + \mathrm{e}^{1.0 + 2.0 * x_1 + 3.0 * x_2}}$
Note that the values $x_1$ and $x_2$ are sampled from a Gaussian distribution.
* $X_1 \sim \mathcal{N}(0, 1)$
* $X_2 \sim \mathcal{N}(0, 1)$
The value $p$ that we get is the plugged into the [Binomial distribution](https://en.wikipedia.org/wiki/Binomial_distribution) to sample our output labels of 1's and 0's.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from numpy.random import binomial, normal
from scipy.stats import bernoulli, binom
np.random.seed(37)
sns.set(color_codes=True)
n = 10000
X = np.hstack([
np.array([1 for _ in range(n)]).reshape(n, 1),
normal(0.0, 1.0, n).reshape(n, 1),
normal(0.0, 1.0, n).reshape(n, 1)
])
z = np.dot(X, np.array([1.0, 2.0, 3.0])) + normal(0.0, 1.0, n)
p = 1.0 / (1.0 + np.exp(-z))
y = binom.rvs(1, p)
```
## Visualize data
Here's a plot of the positive (the one's or red dots) and negative (the zero's or green dots) examples.
```
fig, ax = plt.subplots(1, 1, figsize=(10, 5), sharex=False, sharey=False)
colors = ['r' if 1 == v else 'g' for v in y]
ax.scatter(X[:, 1], X[:, 2], c=colors, alpha=0.4)
ax.set_title('Scatter plot of classes')
ax.set_xlabel(r'$x_0$')
ax.set_ylabel(r'$x_1$')
```
This plot is the density over the `score` $Xw$ (the data $X$ dot the weights $w$).
```
fig, ax = plt.subplots(1, 1, figsize=(10, 5), sharex=False, sharey=False)
sns.distplot(z, ax=ax)
ax.set_title('Density plot of scores')
ax.set_xlabel(r'score')
ax.set_ylabel(r'probability')
```
These next two plots are of the density of the probabilities (note how they center around 0 and 1) and a bar chart of the frequency of the classes.
```
fig, ax = plt.subplots(1, 2, figsize=(20, 5), sharex=False, sharey=False)
sns.distplot(p, bins=50, ax=ax[0])
ax[0].set_title('Density plot of probabilities')
ax[0].set_xlabel(r'probability')
ax[0].set_xlim([0, 1])
sns.countplot(y, ax=ax[1])
ax[1].set_title('Counts of classes')
ax[1].set_xlabel(r'classes')
ax[1].set_ylabel(r'count')
```
## Use gradient descent through Autograd to learn the weights
Now we use Autograd to help learn the coefficients of the logistic model through gradient descent. Note that the loss function is defined as follows.
$\frac{1}{n} \sum -(\hat{y}_i - y_i) + \log (1 + \mathrm{e}^{\hat{y}_i})$
* $\hat{y}_i$ is the i-th predicted probability
* $y_i$ is the i-th class label (0 or 1)
Using Autograd, we do not have to specify (or code up) the gradients of this loss function. We will run the gradient descent algorithm to learn the weights of the logistic regression model using different learning rates, $\alpha$ e.g. 0.01, 0.05, 0.1, 1.0.
```
import autograd.numpy as np
from autograd import grad
from autograd.numpy import exp, log, sqrt
# define the loss function
def loss(w, X, y):
n = float(len(X))
y_pred = np.dot(X, w)
return np.sum(-(y_pred * y) + log(1.0 + exp(y_pred))) / n
#the magic line that gives you the gradient of the loss function
loss_grad = grad(loss)
def learn_weights(X, y, alpha=0.05, max_iter=30000, debug=False):
w = np.array([0.0 for _ in range(X.shape[1])])
if debug is True:
print('initial weights = {}'.format(w))
loss_trace = []
weight_trace = []
for i in range(max_iter):
loss = loss_grad(w, X, y)
w = w - (loss * alpha)
if i % 2000 == 0 and debug is True:
print('{}: loss = {}, weights = {}'.format(i, loss, w))
loss_trace.append(loss)
weight_trace.append(w)
if debug is True:
print('intercept + weights: {}'.format(w))
loss_trace = np.array(loss_trace)
weight_trace = np.array(weight_trace)
return w, loss_trace, weight_trace
def plot_traces(loss_trace, weight_trace, alpha):
fig, ax = plt.subplots(1, 2, figsize=(20, 5))
ax[0].set_title(r'Log-loss of the weights over iterations, $\alpha=${}'.format(alpha))
ax[0].set_xlabel('iteration')
ax[0].set_ylabel('log-loss')
ax[0].plot(loss_trace[:, 0], label=r'$\beta$')
ax[0].plot(loss_trace[:, 1], label=r'$x_0$')
ax[0].plot(loss_trace[:, 2], label=r'$x_1$')
ax[0].legend()
ax[1].set_title(r'Weight learning over iterations, $\alpha=${}'.format(alpha))
ax[1].set_xlabel('iteration')
ax[1].set_ylabel('weight')
ax[1].plot(weight_trace[:, 0], label=r'$\beta$')
ax[1].plot(weight_trace[:, 1], label=r'$x_0$')
ax[1].plot(weight_trace[:, 2], label=r'$x_1$')
ax[1].legend()
```
The important thing to note here is that the learning rate impacts how quickly we converge to the estimated coefficients. For smaller learning rates, it takes longer to find the optimal solution (weights). For the running example, $\alpha=0.01$ takes over 8,000 iterations to reach the optimal solution. Be careful, however, as $\alpha$ will need to be fined tune so that the solution can converge.
We plot the traces of the loss over the number of iterations as well as the change in the coefficients over the number of iterations.
```
w, loss_trace, weight_trace = learn_weights(X, y, alpha=0.01, max_iter=9000)
plot_traces(loss_trace, weight_trace, alpha=0.01)
w, loss_trace, weight_trace = learn_weights(X, y, alpha=0.05, max_iter=6000)
plot_traces(loss_trace, weight_trace, alpha=0.05)
w, loss_trace, weight_trace = learn_weights(X, y, alpha=0.1, max_iter=6000)
plot_traces(loss_trace, weight_trace, alpha=0.1)
w, loss_trace, weight_trace = learn_weights(X, y, alpha=1.0, max_iter=6000)
plot_traces(loss_trace, weight_trace, alpha=1.0)
```
## Verify the results with scikit-learn
```
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression(fit_intercept=False, solver='lbfgs')
lr.fit(X, y)
print(lr.coef_)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# How to Publish a Pipeline and Invoke the REST endpoint
In this notebook, we will see how we can publish a pipeline and then invoke the REST endpoint.
## Prerequisites and Azure Machine Learning Basics
If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the [configuration Notebook](https://aka.ms/pl-config) first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc.
### Initialization Steps
```
import azureml.core
from azureml.core import Workspace, Datastore, Experiment, Dataset
from azureml.data import OutputFileDatasetConfig
from azureml.core.compute import AmlCompute
from azureml.core.compute import ComputeTarget
# Check core SDK version number
print("SDK version:", azureml.core.VERSION)
from azureml.pipeline.core import Pipeline
from azureml.pipeline.steps import PythonScriptStep
from azureml.pipeline.core.graph import PipelineParameter
print("Pipeline SDK-specific imports completed")
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# Default datastore (Azure blob storage)
# def_blob_store = ws.get_default_datastore()
def_blob_store = Datastore(ws, "workspaceblobstore")
print("Blobstore's name: {}".format(def_blob_store.name))
```
### Compute Targets
#### Retrieve an already attached Azure Machine Learning Compute
```
from azureml.core.compute_target import ComputeTargetException
aml_compute_target = "cpu-cluster"
try:
aml_compute = AmlCompute(ws, aml_compute_target)
print("found existing compute target.")
except ComputeTargetException:
print("creating new compute target")
provisioning_config = AmlCompute.provisioning_configuration(vm_size = "STANDARD_D2_V2",
min_nodes = 1,
max_nodes = 4)
aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)
aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current Azure Machine Learning Compute status, use get_status()
# example: un-comment the following line.
# print(aml_compute.get_status().serialize())
```
## Building Pipeline Steps with Inputs and Outputs
A step in the pipeline can take [dataset](https://docs.microsoft.com/python/api/azureml-core/azureml.data.filedataset?view=azure-ml-py) as input. This dataset can be a data source that lives in one of the accessible data locations, or intermediate data produced by a previous step in the pipeline.
```
# Uploading data to the datastore
data_path = def_blob_store.upload_files(["./20news.pkl"], target_path="20newsgroups", overwrite=True)
# Reference the data uploaded to blob storage using file dataset
# Assign the datasource to blob_input_data variable
blob_input_data = Dataset.File.from_files(data_path).as_named_input("test_data")
print("Dataset created")
# Define intermediate data using OutputFileDatasetConfig
processed_data1 = OutputFileDatasetConfig(name="processed_data1")
print("Output dataset object created")
```
#### Define a Step that consumes a dataset and produces intermediate data.
In this step, we define a step that consumes a dataset and produces intermediate data.
**Open `train.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**
The best practice is to use separate folders for scripts and its dependent files for each step and specify that folder as the `source_directory` for the step. This helps reduce the size of the snapshot created for the step (only the specific folder is snapshotted). Since changes in any files in the `source_directory` would trigger a re-upload of the snapshot, this helps keep the reuse of the step when there are no changes in the `source_directory` of the step.
```
# trainStep consumes the datasource (Datareference) in the previous step
# and produces processed_data1
source_directory = "publish_run_train"
trainStep = PythonScriptStep(
script_name="train.py",
arguments=["--input_data", blob_input_data.as_mount(), "--output_train", processed_data1],
compute_target=aml_compute,
source_directory=source_directory
)
print("trainStep created")
```
#### Define a Step that consumes intermediate data and produces intermediate data
In this step, we define a step that consumes an intermediate data and produces intermediate data.
**Open `extract.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**
```
# extractStep to use the intermediate data produced by trainStep
# This step also produces an output processed_data2
processed_data2 = OutputFileDatasetConfig(name="processed_data2")
source_directory = "publish_run_extract"
extractStep = PythonScriptStep(
script_name="extract.py",
arguments=["--input_extract", processed_data1.as_input(), "--output_extract", processed_data2],
compute_target=aml_compute,
source_directory=source_directory)
print("extractStep created")
```
#### Define a Step that consumes multiple intermediate data and produces intermediate data
In this step, we define a step that consumes multiple intermediate data and produces intermediate data.
### PipelineParameter
This step also has a [PipelineParameter](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.graph.pipelineparameter?view=azure-ml-py) argument that help with calling the REST endpoint of the published pipeline.
```
# We will use this later in publishing pipeline
pipeline_param = PipelineParameter(name="pipeline_arg", default_value=10)
print("pipeline parameter created")
```
**Open `compare.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**
```
# Now define compareStep that takes two inputs (both intermediate data), and produce an output
processed_data3 = OutputFileDatasetConfig(name="processed_data3")
# You can register the output as dataset after job completion
processed_data3 = processed_data3.register_on_complete("compare_result")
source_directory = "publish_run_compare"
compareStep = PythonScriptStep(
script_name="compare.py",
arguments=["--compare_data1", processed_data1.as_input(), "--compare_data2", processed_data2.as_input(), "--output_compare", processed_data3, "--pipeline_param", pipeline_param],
compute_target=aml_compute,
source_directory=source_directory)
print("compareStep created")
```
#### Build the pipeline
```
pipeline1 = Pipeline(workspace=ws, steps=[compareStep])
print ("Pipeline is built")
```
## Run published pipeline
### Publish the pipeline
```
published_pipeline1 = pipeline1.publish(name="My_New_Pipeline", description="My Published Pipeline Description", continue_on_step_failure=True)
published_pipeline1
```
Note: the continue_on_step_failure parameter specifies whether the execution of steps in the Pipeline will continue if one step fails. The default value is False, meaning when one step fails, the Pipeline execution will stop, canceling any running steps.
### Publish the pipeline from a submitted PipelineRun
It is also possible to publish a pipeline from a submitted PipelineRun
```
# submit a pipeline run
pipeline_run1 = Experiment(ws, 'Pipeline_experiment').submit(pipeline1)
# publish a pipeline from the submitted pipeline run
published_pipeline2 = pipeline_run1.publish_pipeline(name="My_New_Pipeline2", description="My Published Pipeline Description", version="0.1", continue_on_step_failure=True)
published_pipeline2
```
### Get published pipeline
You can get the published pipeline using **pipeline id**.
To get all the published pipelines for a given workspace(ws):
```css
all_pub_pipelines = PublishedPipeline.get_all(ws)
```
```
from azureml.pipeline.core import PublishedPipeline
pipeline_id = published_pipeline1.id # use your published pipeline id
published_pipeline = PublishedPipeline.get(ws, pipeline_id)
published_pipeline
```
### Run published pipeline using its REST endpoint
[This notebook](https://aka.ms/pl-restep-auth) shows how to authenticate to AML workspace.
```
from azureml.core.authentication import InteractiveLoginAuthentication
import requests
auth = InteractiveLoginAuthentication()
aad_token = auth.get_authentication_header()
rest_endpoint1 = published_pipeline.endpoint
print("You can perform HTTP POST on URL {} to trigger this pipeline".format(rest_endpoint1))
# specify the param when running the pipeline
response = requests.post(rest_endpoint1,
headers=aad_token,
json={"ExperimentName": "My_Pipeline1",
"RunSource": "SDK",
"ParameterAssignments": {"pipeline_arg": 45}})
try:
response.raise_for_status()
except Exception:
raise Exception('Received bad response from the endpoint: {}\n'
'Response Code: {}\n'
'Headers: {}\n'
'Content: {}'.format(rest_endpoint, response.status_code, response.headers, response.content))
run_id = response.json().get('Id')
print('Submitted pipeline run: ', run_id)
```
# Next: Data Transfer
The next [notebook](https://aka.ms/pl-data-trans) will showcase data transfer steps between different types of data stores.
| github_jupyter |
# Modelling poisson using PINN
__Author: Manu Jayadharan__
Written as part of FlowNet package, a TensorFlow based neural network package to solve fluid flow PDEs.
Solving the poisson equation $-\Delta u = f$ using a physics informed neural network
## 1D problem poisson problem
### Manufactured solution
We use $u = 3sin(4x)$ for $x\in [-1,1]$
### Importing packages
```
import numpy as np
import tensorflow as tf
from tensorflow import keras
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
```
## Manufacturing data for trainig
```
np.random.seed(123)
X_tr_pde = np.random.uniform(-1,1,500).reshape(500,1)
```
#### Plotting histogram of randomly selected points to make sure they are uniformly distributed
```
plt.hist(X_tr_pde)
plt.xlabel("training points")
plt.ylabel("frequency ")
Y_tr_pde = np.zeros((X_tr_pde.shape[0],1))
Y_tr_pde = np.concatenate([Y_tr_pde,np.zeros((Y_tr_pde.shape[0],1))],axis=1)
Y_tr_pde.shape
X_tr_Dr_bc_left = -1*np.ones(200).reshape(200,1)
X_tr_Dr_bc_right = 1*np.ones(200).reshape(200,1)
X_bc = np.concatenate([X_tr_Dr_bc_left,X_tr_Dr_bc_right],axis=0)
Y_tr_bc = 3*np.sin(4*X_bc)
Y_tr_bc = np.concatenate([Y_tr_bc,np.ones((Y_tr_bc.shape[0],1))],axis=1)
```
### Scaling the inputs(optional)
```
# from sklearn.preprocessing import StandardScaler
# scaler = StandardScaler()
# X_tr_pde = scaler.fit_transform(X_tr_pde)
X_tr = np.concatenate((X_tr_pde, X_bc), axis=0)
Y_tr = np.concatenate((Y_tr_pde, Y_tr_bc), axis=0)
# from sklearn.preprocessing import StandardScaler
# scaler = StandardScaler()
# X_tr = scaler.fit_transform(X_tr)
# X_tr.std()
(X_tr[0:3] +0.0056276)/4.520744138916567
```
## Defining the NN model (custom Keras model)
- __Model specifications__: 7 layers: 1 input layer, 3 hidden layers with 20 neurons each, 1 dense intermediate layer, 1 gradient layer, 1 laplacian layer, 1 pde layer.
- Output is a list of two elements: value of function and value of the pde operator.
- mean squared error is used for finding the cost function.
- Specialized, sgd( stochastic gradient descenet) type optimizer is used: either nadam or adam.
- tanh activation functions are used.
```
from tensorflow.keras import backend as K
class Poisson1d(tf.keras.Model):
def __init__(self):
super(Poisson1d, self).__init__()
# self.batch_norm_ = keras.layers.BatchNormalization()
self.flatten_input = keras.layers.Flatten()
he_kernel_init = keras.initializers.he_uniform()
self.dense_1 = keras.layers.Dense(20, activation="tanh",
kernel_initializer=he_kernel_init,
name="dense_1")
self.dense_2 = keras.layers.Dense(20, activation="tanh",
kernel_initializer=he_kernel_init,
name="dense_2")
self.dense_3 = keras.layers.Dense(20, activation="tanh",
kernel_initializer=he_kernel_init,
name="dense_3")
self.dense_4 = keras.layers.Dense(1,
name="dense_4")
def findGrad(self,func,argm):
return keras.layers.Lambda(lambda x: K.gradients(x[0],x[1])[0]) ([func,argm])
def findPdeLayer(self, pde_lhs, input_arg):
return keras.layers.Lambda(lambda z: z[0] + 48*tf.sin(4*z[1])) ([pde_lhs, input_arg])
def call(self, inputs):
# layer_0 = self.batch_norm_input(inputs)
layer_0 = self.flatten_input(inputs)
# layer_0_1 = self.batch_norm_input(layer_0)
layer_1 = self.dense_1(layer_0)
# layer_2_0 = self.batch_norm_input(layer_1)
layer_2 = self.dense_2(layer_1)
# layer_3_0 = self.batch_norm_(layer_2)
layer_3 = self.dense_3(layer_2)
layer_4 = self.dense_4(layer_3)
grad_layer = self.findGrad(layer_4, inputs)
laplace_layer = self.findGrad(grad_layer, inputs)
pde_layer = self.findPdeLayer(laplace_layer, inputs)
return layer_4, pde_layer
```
### Defining the loss functions
```
#Loss coming from the boundary terms
def u_loss(y_true, y_pred):
y_true_act = y_true[:,:-1]
at_boundary = tf.cast(y_true[:,-1:,],bool)
u_sq_error = (1/2)*tf.square(y_true_act-y_pred)
return tf.where(at_boundary, u_sq_error, 0.)
#Loss coming from the PDE constrain
def pde_loss(y_true, y_pred):
y_true_act = y_true[:,:-1]
at_boundary = tf.cast(y_true[:,-1:,],bool)
#need to change this to just tf.square(y_pred) after pde constrain is added to grad_layer
# pde_sq_error = (1/2)*tf.square(y_true_act-y_pred)
pde_sq_error = (1/2)*tf.square(y_pred)
return tf.where(at_boundary,0.,pde_sq_error)
```
### Instantiating and compiling the poisson_model
```
poisson_NN = Poisson1d()
poisson_NN.compile(loss=[u_loss,pde_loss],optimizer="adam")
poisson_NN.fit(x=X_tr, y=Y_tr,epochs=100)
```
## Testing the trained network
```
X_test_st = np.random.uniform(-1,1,100).reshape(100,1)
```
### Scaling the test set (only if the trainng data was scaled)
```
# #Scaling test set
# X_test_st_2 = scaler.transform(X_test_st)
#xtrain: mean, std: -0.005627660222786496 4.520744138916567
Y_test = poisson_NN.predict(X_test_st)
```
### Plotting the true and predicted solutions
```
# fig, ax = plt.subplots(nrows=2,ncols=2, figsize=(10,10))
#plotting predicted solution
plt.figure(figsize=(15,7))
plt.subplot(1,2,1)
plt.scatter(X_test_st, Y_test[0][:,0])
plt.title("Predicted solution")
plt.subplot(1,2,2)
plt.scatter(X_test_st, 3*np.sin(4*X_test_st), c="r")
plt.title("True solution")
#True vs predicted solution
plt.figure(figsize=(10,5))
plt.scatter(np.sin(4*X_test_st), Y_test[0][:,0], c="g")
plt.title("True solution vs predicted solution")
plt.xlabel("True solution")
plt.ylabel("Predicted solution")
plt.show()
```
### Notes to be made
- For second order pde, some form of normalization is needed for convergence.
- If the the input data already comes normalized, there is no problem.
- If the data is not normalized, then we would want to some kind of normalization technique like batch normalization or normalization of incoming data.
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Image captioning with visual attention
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/text/image_captioning">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/text/image_captioning.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/image_captioning.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/text/image_captioning.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
Given an image like the example below, our goal is to generate a caption such as "a surfer riding on a wave".

*[Image Source](https://commons.wikimedia.org/wiki/Surfing#/media/File:Surfing_in_Hawaii.jpg); License: Public Domain*
To accomplish this, you'll use an attention-based model, which enables us to see what parts of the image the model focuses on as it generates a caption.
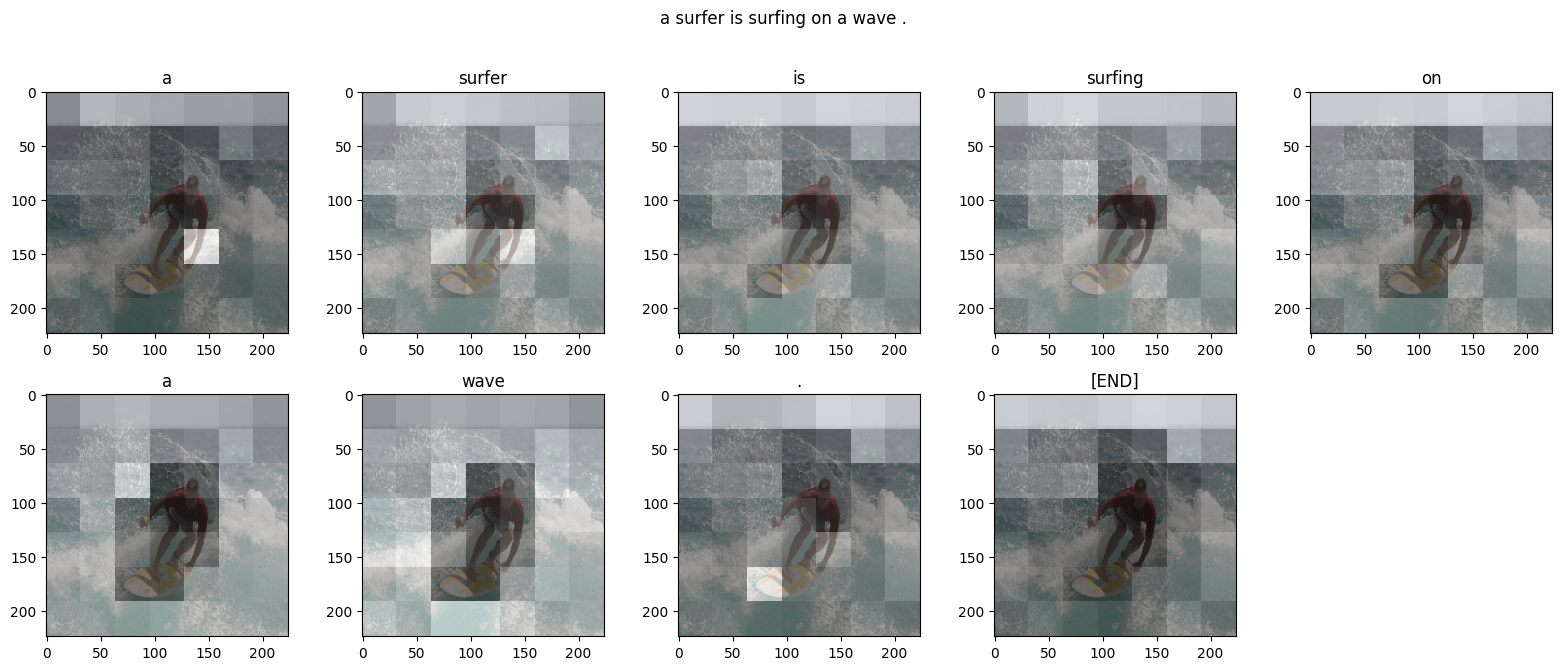
The model architecture is similar to [Show, Attend and Tell: Neural Image Caption Generation with Visual Attention](https://arxiv.org/abs/1502.03044).
This notebook is an end-to-end example. When you run the notebook, it downloads the [MS-COCO](http://cocodataset.org/#home) dataset, preprocesses and caches a subset of images using Inception V3, trains an encoder-decoder model, and generates captions on new images using the trained model.
In this example, you will train a model on a relatively small amount of data—the first 30,000 captions for about 20,000 images (because there are multiple captions per image in the dataset).
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# You'll generate plots of attention in order to see which parts of an image
# our model focuses on during captioning
import matplotlib.pyplot as plt
# Scikit-learn includes many helpful utilities
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
import re
import numpy as np
import os
import time
import json
from glob import glob
from PIL import Image
import pickle
```
## Download and prepare the MS-COCO dataset
You will use the [MS-COCO dataset](http://cocodataset.org/#home) to train our model. The dataset contains over 82,000 images, each of which has at least 5 different caption annotations. The code below downloads and extracts the dataset automatically.
**Caution: large download ahead**. You'll use the training set, which is a 13GB file.
```
# Download caption annotation files
annotation_folder = '/annotations/'
if not os.path.exists(os.path.abspath('.') + annotation_folder):
annotation_zip = tf.keras.utils.get_file('captions.zip',
cache_subdir=os.path.abspath('.'),
origin = 'http://images.cocodataset.org/annotations/annotations_trainval2014.zip',
extract = True)
annotation_file = os.path.dirname(annotation_zip)+'/annotations/captions_train2014.json'
os.remove(annotation_zip)
# Download image files
image_folder = '/train2014/'
if not os.path.exists(os.path.abspath('.') + image_folder):
image_zip = tf.keras.utils.get_file('train2014.zip',
cache_subdir=os.path.abspath('.'),
origin = 'http://images.cocodataset.org/zips/train2014.zip',
extract = True)
PATH = os.path.dirname(image_zip) + image_folder
os.remove(image_zip)
else:
PATH = os.path.abspath('.') + image_folder
```
## Optional: limit the size of the training set
To speed up training for this tutorial, you'll use a subset of 30,000 captions and their corresponding images to train our model. Choosing to use more data would result in improved captioning quality.
```
# Read the json file
with open(annotation_file, 'r') as f:
annotations = json.load(f)
# Store captions and image names in vectors
all_captions = []
all_img_name_vector = []
for annot in annotations['annotations']:
caption = '<start> ' + annot['caption'] + ' <end>'
image_id = annot['image_id']
full_coco_image_path = PATH + 'COCO_train2014_' + '%012d.jpg' % (image_id)
all_img_name_vector.append(full_coco_image_path)
all_captions.append(caption)
# Shuffle captions and image_names together
# Set a random state
train_captions, img_name_vector = shuffle(all_captions,
all_img_name_vector,
random_state=1)
# Select the first 30000 captions from the shuffled set
num_examples = 30000
train_captions = train_captions[:num_examples]
img_name_vector = img_name_vector[:num_examples]
len(train_captions), len(all_captions)
```
## Preprocess the images using InceptionV3
Next, you will use InceptionV3 (which is pretrained on Imagenet) to classify each image. You will extract features from the last convolutional layer.
First, you will convert the images into InceptionV3's expected format by:
* Resizing the image to 299px by 299px
* [Preprocess the images](https://cloud.google.com/tpu/docs/inception-v3-advanced#preprocessing_stage) using the [preprocess_input](https://www.tensorflow.org/api_docs/python/tf/keras/applications/inception_v3/preprocess_input) method to normalize the image so that it contains pixels in the range of -1 to 1, which matches the format of the images used to train InceptionV3.
```
def load_image(image_path):
img = tf.io.read_file(image_path)
img = tf.image.decode_jpeg(img, channels=3)
img = tf.image.resize(img, (299, 299))
img = tf.keras.applications.inception_v3.preprocess_input(img)
return img, image_path
```
## Initialize InceptionV3 and load the pretrained Imagenet weights
Now you'll create a tf.keras model where the output layer is the last convolutional layer in the InceptionV3 architecture. The shape of the output of this layer is ```8x8x2048```. You use the last convolutional layer because you are using attention in this example. You don't perform this initialization during training because it could become a bottleneck.
* You forward each image through the network and store the resulting vector in a dictionary (image_name --> feature_vector).
* After all the images are passed through the network, you pickle the dictionary and save it to disk.
```
image_model = tf.keras.applications.InceptionV3(include_top=False,
weights='imagenet')
new_input = image_model.input
hidden_layer = image_model.layers[-1].output
image_features_extract_model = tf.keras.Model(new_input, hidden_layer)
```
## Caching the features extracted from InceptionV3
You will pre-process each image with InceptionV3 and cache the output to disk. Caching the output in RAM would be faster but also memory intensive, requiring 8 \* 8 \* 2048 floats per image. At the time of writing, this exceeds the memory limitations of Colab (currently 12GB of memory).
Performance could be improved with a more sophisticated caching strategy (for example, by sharding the images to reduce random access disk I/O), but that would require more code.
The caching will take about 10 minutes to run in Colab with a GPU. If you'd like to see a progress bar, you can:
1. install [tqdm](https://github.com/tqdm/tqdm):
`!pip install tqdm`
2. Import tqdm:
`from tqdm import tqdm`
3. Change the following line:
`for img, path in image_dataset:`
to:
`for img, path in tqdm(image_dataset):`
```
# Get unique images
encode_train = sorted(set(img_name_vector))
# Feel free to change batch_size according to your system configuration
image_dataset = tf.data.Dataset.from_tensor_slices(encode_train)
image_dataset = image_dataset.map(
load_image, num_parallel_calls=tf.data.experimental.AUTOTUNE).batch(16)
for img, path in image_dataset:
batch_features = image_features_extract_model(img)
batch_features = tf.reshape(batch_features,
(batch_features.shape[0], -1, batch_features.shape[3]))
for bf, p in zip(batch_features, path):
path_of_feature = p.numpy().decode("utf-8")
np.save(path_of_feature, bf.numpy())
```
## Preprocess and tokenize the captions
* First, you'll tokenize the captions (for example, by splitting on spaces). This gives us a vocabulary of all of the unique words in the data (for example, "surfing", "football", and so on).
* Next, you'll limit the vocabulary size to the top 5,000 words (to save memory). You'll replace all other words with the token "UNK" (unknown).
* You then create word-to-index and index-to-word mappings.
* Finally, you pad all sequences to be the same length as the longest one.
```
# Find the maximum length of any caption in our dataset
def calc_max_length(tensor):
return max(len(t) for t in tensor)
# Choose the top 5000 words from the vocabulary
top_k = 5000
tokenizer = tf.keras.preprocessing.text.Tokenizer(num_words=top_k,
oov_token="<unk>",
filters='!"#$%&()*+.,-/:;=?@[\]^_`{|}~ ')
tokenizer.fit_on_texts(train_captions)
train_seqs = tokenizer.texts_to_sequences(train_captions)
tokenizer.word_index['<pad>'] = 0
tokenizer.index_word[0] = '<pad>'
# Create the tokenized vectors
train_seqs = tokenizer.texts_to_sequences(train_captions)
# Pad each vector to the max_length of the captions
# If you do not provide a max_length value, pad_sequences calculates it automatically
cap_vector = tf.keras.preprocessing.sequence.pad_sequences(train_seqs, padding='post')
# Calculates the max_length, which is used to store the attention weights
max_length = calc_max_length(train_seqs)
```
## Split the data into training and testing
```
# Create training and validation sets using an 80-20 split
img_name_train, img_name_val, cap_train, cap_val = train_test_split(img_name_vector,
cap_vector,
test_size=0.2,
random_state=0)
len(img_name_train), len(cap_train), len(img_name_val), len(cap_val)
```
## Create a tf.data dataset for training
Our images and captions are ready! Next, let's create a tf.data dataset to use for training our model.
```
# Feel free to change these parameters according to your system's configuration
BATCH_SIZE = 64
BUFFER_SIZE = 1000
embedding_dim = 256
units = 512
vocab_size = top_k + 1
num_steps = len(img_name_train) // BATCH_SIZE
# Shape of the vector extracted from InceptionV3 is (64, 2048)
# These two variables represent that vector shape
features_shape = 2048
attention_features_shape = 64
# Load the numpy files
def map_func(img_name, cap):
img_tensor = np.load(img_name.decode('utf-8')+'.npy')
return img_tensor, cap
dataset = tf.data.Dataset.from_tensor_slices((img_name_train, cap_train))
# Use map to load the numpy files in parallel
dataset = dataset.map(lambda item1, item2: tf.numpy_function(
map_func, [item1, item2], [tf.float32, tf.int32]),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# Shuffle and batch
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
dataset = dataset.prefetch(buffer_size=tf.data.experimental.AUTOTUNE)
```
## Model
Fun fact: the decoder below is identical to the one in the example for [Neural Machine Translation with Attention](../sequences/nmt_with_attention.ipynb).
The model architecture is inspired by the [Show, Attend and Tell](https://arxiv.org/pdf/1502.03044.pdf) paper.
* In this example, you extract the features from the lower convolutional layer of InceptionV3 giving us a vector of shape (8, 8, 2048).
* You squash that to a shape of (64, 2048).
* This vector is then passed through the CNN Encoder (which consists of a single Fully connected layer).
* The RNN (here GRU) attends over the image to predict the next word.
```
class BahdanauAttention(tf.keras.Model):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, features, hidden):
# features(CNN_encoder output) shape == (batch_size, 64, embedding_dim)
# hidden shape == (batch_size, hidden_size)
# hidden_with_time_axis shape == (batch_size, 1, hidden_size)
hidden_with_time_axis = tf.expand_dims(hidden, 1)
# score shape == (batch_size, 64, hidden_size)
score = tf.nn.tanh(self.W1(features) + self.W2(hidden_with_time_axis))
# attention_weights shape == (batch_size, 64, 1)
# you get 1 at the last axis because you are applying score to self.V
attention_weights = tf.nn.softmax(self.V(score), axis=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * features
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
class CNN_Encoder(tf.keras.Model):
# Since you have already extracted the features and dumped it using pickle
# This encoder passes those features through a Fully connected layer
def __init__(self, embedding_dim):
super(CNN_Encoder, self).__init__()
# shape after fc == (batch_size, 64, embedding_dim)
self.fc = tf.keras.layers.Dense(embedding_dim)
def call(self, x):
x = self.fc(x)
x = tf.nn.relu(x)
return x
class RNN_Decoder(tf.keras.Model):
def __init__(self, embedding_dim, units, vocab_size):
super(RNN_Decoder, self).__init__()
self.units = units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc1 = tf.keras.layers.Dense(self.units)
self.fc2 = tf.keras.layers.Dense(vocab_size)
self.attention = BahdanauAttention(self.units)
def call(self, x, features, hidden):
# defining attention as a separate model
context_vector, attention_weights = self.attention(features, hidden)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# passing the concatenated vector to the GRU
output, state = self.gru(x)
# shape == (batch_size, max_length, hidden_size)
x = self.fc1(output)
# x shape == (batch_size * max_length, hidden_size)
x = tf.reshape(x, (-1, x.shape[2]))
# output shape == (batch_size * max_length, vocab)
x = self.fc2(x)
return x, state, attention_weights
def reset_state(self, batch_size):
return tf.zeros((batch_size, self.units))
encoder = CNN_Encoder(embedding_dim)
decoder = RNN_Decoder(embedding_dim, units, vocab_size)
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction='none')
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
```
## Checkpoint
```
checkpoint_path = "./checkpoints/train"
ckpt = tf.train.Checkpoint(encoder=encoder,
decoder=decoder,
optimizer = optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
start_epoch = 0
if ckpt_manager.latest_checkpoint:
start_epoch = int(ckpt_manager.latest_checkpoint.split('-')[-1])
# restoring the latest checkpoint in checkpoint_path
ckpt.restore(ckpt_manager.latest_checkpoint)
```
## Training
* You extract the features stored in the respective `.npy` files and then pass those features through the encoder.
* The encoder output, hidden state(initialized to 0) and the decoder input (which is the start token) is passed to the decoder.
* The decoder returns the predictions and the decoder hidden state.
* The decoder hidden state is then passed back into the model and the predictions are used to calculate the loss.
* Use teacher forcing to decide the next input to the decoder.
* Teacher forcing is the technique where the target word is passed as the next input to the decoder.
* The final step is to calculate the gradients and apply it to the optimizer and backpropagate.
```
# adding this in a separate cell because if you run the training cell
# many times, the loss_plot array will be reset
loss_plot = []
@tf.function
def train_step(img_tensor, target):
loss = 0
# initializing the hidden state for each batch
# because the captions are not related from image to image
hidden = decoder.reset_state(batch_size=target.shape[0])
dec_input = tf.expand_dims([tokenizer.word_index['<start>']] * target.shape[0], 1)
with tf.GradientTape() as tape:
features = encoder(img_tensor)
for i in range(1, target.shape[1]):
# passing the features through the decoder
predictions, hidden, _ = decoder(dec_input, features, hidden)
loss += loss_function(target[:, i], predictions)
# using teacher forcing
dec_input = tf.expand_dims(target[:, i], 1)
total_loss = (loss / int(target.shape[1]))
trainable_variables = encoder.trainable_variables + decoder.trainable_variables
gradients = tape.gradient(loss, trainable_variables)
optimizer.apply_gradients(zip(gradients, trainable_variables))
return loss, total_loss
EPOCHS = 20
for epoch in range(start_epoch, EPOCHS):
start = time.time()
total_loss = 0
for (batch, (img_tensor, target)) in enumerate(dataset):
batch_loss, t_loss = train_step(img_tensor, target)
total_loss += t_loss
if batch % 100 == 0:
print ('Epoch {} Batch {} Loss {:.4f}'.format(
epoch + 1, batch, batch_loss.numpy() / int(target.shape[1])))
# storing the epoch end loss value to plot later
loss_plot.append(total_loss / num_steps)
if epoch % 5 == 0:
ckpt_manager.save()
print ('Epoch {} Loss {:.6f}'.format(epoch + 1,
total_loss/num_steps))
print ('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
plt.plot(loss_plot)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Loss Plot')
plt.show()
```
## Caption!
* The evaluate function is similar to the training loop, except you don't use teacher forcing here. The input to the decoder at each time step is its previous predictions along with the hidden state and the encoder output.
* Stop predicting when the model predicts the end token.
* And store the attention weights for every time step.
```
def evaluate(image):
attention_plot = np.zeros((max_length, attention_features_shape))
hidden = decoder.reset_state(batch_size=1)
temp_input = tf.expand_dims(load_image(image)[0], 0)
img_tensor_val = image_features_extract_model(temp_input)
img_tensor_val = tf.reshape(img_tensor_val, (img_tensor_val.shape[0], -1, img_tensor_val.shape[3]))
features = encoder(img_tensor_val)
dec_input = tf.expand_dims([tokenizer.word_index['<start>']], 0)
result = []
for i in range(max_length):
predictions, hidden, attention_weights = decoder(dec_input, features, hidden)
attention_plot[i] = tf.reshape(attention_weights, (-1, )).numpy()
predicted_id = tf.random.categorical(predictions, 1)[0][0].numpy()
result.append(tokenizer.index_word[predicted_id])
if tokenizer.index_word[predicted_id] == '<end>':
return result, attention_plot
dec_input = tf.expand_dims([predicted_id], 0)
attention_plot = attention_plot[:len(result), :]
return result, attention_plot
def plot_attention(image, result, attention_plot):
temp_image = np.array(Image.open(image))
fig = plt.figure(figsize=(10, 10))
len_result = len(result)
for l in range(len_result):
temp_att = np.resize(attention_plot[l], (8, 8))
ax = fig.add_subplot(len_result//2, len_result//2, l+1)
ax.set_title(result[l])
img = ax.imshow(temp_image)
ax.imshow(temp_att, cmap='gray', alpha=0.6, extent=img.get_extent())
plt.tight_layout()
plt.show()
# captions on the validation set
rid = np.random.randint(0, len(img_name_val))
image = img_name_val[rid]
real_caption = ' '.join([tokenizer.index_word[i] for i in cap_val[rid] if i not in [0]])
result, attention_plot = evaluate(image)
print ('Real Caption:', real_caption)
print ('Prediction Caption:', ' '.join(result))
plot_attention(image, result, attention_plot)
```
## Try it on your own images
For fun, below we've provided a method you can use to caption your own images with the model we've just trained. Keep in mind, it was trained on a relatively small amount of data, and your images may be different from the training data (so be prepared for weird results!)
```
image_url = 'https://tensorflow.org/images/surf.jpg'
image_extension = image_url[-4:]
image_path = tf.keras.utils.get_file('image'+image_extension,
origin=image_url)
result, attention_plot = evaluate(image_path)
print ('Prediction Caption:', ' '.join(result))
plot_attention(image_path, result, attention_plot)
# opening the image
Image.open(image_path)
```
# Next steps
Congrats! You've just trained an image captioning model with attention. Next, take a look at this example [Neural Machine Translation with Attention](../sequences/nmt_with_attention.ipynb). It uses a similar architecture to translate between Spanish and English sentences. You can also experiment with training the code in this notebook on a different dataset.
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import math
import warnings
warnings.filterwarnings("ignore")
# fix_yahoo_finance is used to fetch data
import fix_yahoo_finance as yf
yf.pdr_override()
# input
symbol = 'AMD'
start = '2007-01-01'
end = '2019-01-01'
# Read data
dataset = yf.download(symbol,start,end)
# View Columns
dataset.head()
dataset['Open_Close'] = (dataset['Open'] - dataset['Adj Close'])/dataset['Open']
dataset['High_Low'] = (dataset['High'] - dataset['Low'])/dataset['Low']
dataset['Increase_Decrease'] = np.where(dataset['Volume'].shift(-1) > dataset['Volume'],1,0)
dataset['Buy_Sell_on_Open'] = np.where(dataset['Open'].shift(-1) > dataset['Open'],1,0)
dataset['Buy_Sell'] = np.where(dataset['Adj Close'].shift(-1) > dataset['Adj Close'],1,0)
dataset['Returns'] = dataset['Adj Close'].pct_change()
dataset = dataset.dropna()
dataset.head()
dataset['High'].plot(figsize=(16,8))
from pylab import rcParams
import statsmodels.api as sm
rcParams['figure.figsize'] = 11, 9
decomposed_volume = sm.tsa.seasonal_decompose(dataset["High"],freq=360) # The frequncy is annual
figure = decomposed_volume.plot()
plt.show()
from statsmodels.tsa.stattools import adfuller
# Augmented Dickey-Fuller test on volume of stock
adf = adfuller(dataset["Volume"])
print("p-value of stock: {}".format(float(adf[1])))
# The original non-stationary plot
decomposed_volume.trend.plot()
# The new stationary plot
decomposed_volume.trend.diff().plot()
```
## ARMA models
```
from statsmodels.tsa.arima_model import ARMA
# Predicting stock closing prices
humid = ARMA(dataset["Adj Close"].diff().iloc[1:].values, order=(1,0))
res = humid.fit()
res.plot_predict(start=900, end=1010)
plt.show()
# Forecasting and predicting stocks volume
model = ARMA(dataset["Volume"].diff().iloc[1:].values, order=(3,3))
result = model.fit()
print(result.summary())
print("μ={}, ϕ={}, θ={}".format(result.params[0],result.params[1],result.params[2]))
result.plot_predict(start=1000, end=1100)
plt.show()
from sklearn.metrics import mean_squared_error
rmse = math.sqrt(mean_squared_error(dataset["Volume"].diff().iloc[1000:1101].values, result.predict(start=1000,end=1100)))
print("The root mean squared error is {}.".format(rmse))
```
## ARIMA models
```
from statsmodels.tsa.arima_model import ARIMA
# Predicting the stocks volume
rcParams['figure.figsize'] = 16, 6
model = ARIMA(dataset["Volume"].diff().iloc[1:].values, order=(2,1,0))
result = model.fit()
print(result.summary())
result.plot_predict(start=700, end=1000)
plt.show()
rmse = math.sqrt(mean_squared_error(dataset["Volume"].diff().iloc[700:1001].values, result.predict(start=700,end=1000)))
print("The root mean squared error is {}.".format(rmse))
```
| github_jupyter |
# Implementing a Neural Network
In this exercise we will develop a neural network with fully-connected layers to perform classification, and test it out on the CIFAR-10 dataset.
```
# A bit of setup
import numpy as np
import matplotlib.pyplot as plt
from cs231n.classifiers.neural_net import TwoLayerNet
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
```
We will use the class `TwoLayerNet` in the file `cs231n/classifiers/neural_net.py` to represent instances of our network. The network parameters are stored in the instance variable `self.params` where keys are string parameter names and values are numpy arrays. Below, we initialize toy data and a toy model that we will use to develop your implementation.
```
# Create a small net and some toy data to check your implementations.
# Note that we set the random seed for repeatable experiments.
input_size = 4
hidden_size = 10
num_classes = 3
num_inputs = 5
def init_toy_model():
np.random.seed(0)
return TwoLayerNet(input_size, hidden_size, num_classes, std=1e-1)
def init_toy_data():
np.random.seed(1)
X = 10 * np.random.randn(num_inputs, input_size)
y = np.array([0, 1, 2, 2, 1])
return X, y
net = init_toy_model()
X, y = init_toy_data()
```
# Forward pass: compute scores
Open the file `cs231n/classifiers/neural_net.py` and look at the method `TwoLayerNet.loss`. This function is very similar to the loss functions you have written for the SVM and Softmax exercises: It takes the data and weights and computes the class scores, the loss, and the gradients on the parameters.
Implement the first part of the forward pass which uses the weights and biases to compute the scores for all inputs.
```
scores = net.loss(X)
print 'Your scores:'
print scores
print
print 'correct scores:'
correct_scores = np.asarray([
[-0.81233741, -1.27654624, -0.70335995],
[-0.17129677, -1.18803311, -0.47310444],
[-0.51590475, -1.01354314, -0.8504215 ],
[-0.15419291, -0.48629638, -0.52901952],
[-0.00618733, -0.12435261, -0.15226949]])
print correct_scores
print
# The difference should be very small. We get < 1e-7
print 'Difference between your scores and correct scores:'
print np.sum(np.abs(scores - correct_scores))
```
# Forward pass: compute loss
In the same function, implement the second part that computes the data and regularizaion loss.
```
loss, _ = net.loss(X, y, reg=0.1)
correct_loss = 1.30378789133
# should be very small, we get < 1e-12
print 'Difference between your loss and correct loss:'
print np.sum(np.abs(loss - correct_loss))
```
# Backward pass
Implement the rest of the function. This will compute the gradient of the loss with respect to the variables `W1`, `b1`, `W2`, and `b2`. Now that you (hopefully!) have a correctly implemented forward pass, you can debug your backward pass using a numeric gradient check:
```
from cs231n.gradient_check import eval_numerical_gradient
# Use numeric gradient checking to check your implementation of the backward pass.
# If your implementation is correct, the difference between the numeric and
# analytic gradients should be less than 1e-8 for each of W1, W2, b1, and b2.
loss, grads = net.loss(X, y, reg=0.1)
# these should all be less than 1e-8 or so
for param_name in grads:
f = lambda W: net.loss(X, y, reg=0.1)[0]
param_grad_num = eval_numerical_gradient(f, net.params[param_name], verbose=False)
print '%s max relative error: %e' % (param_name, rel_error(param_grad_num, grads[param_name]))
```
# Train the network
To train the network we will use stochastic gradient descent (SGD), similar to the SVM and Softmax classifiers. Look at the function `TwoLayerNet.train` and fill in the missing sections to implement the training procedure. This should be very similar to the training procedure you used for the SVM and Softmax classifiers. You will also have to implement `TwoLayerNet.predict`, as the training process periodically performs prediction to keep track of accuracy over time while the network trains.
Once you have implemented the method, run the code below to train a two-layer network on toy data. You should achieve a training loss less than 0.2.
```
net = init_toy_model()
stats = net.train(X, y, X, y,
learning_rate=1e-1, reg=1e-5,
num_iters=100, verbose=False)
print 'Final training loss: ', stats['loss_history'][-1]
# plot the loss history
plt.plot(stats['loss_history'])
plt.xlabel('iteration')
plt.ylabel('training loss')
plt.title('Training Loss history')
plt.show()
```
# Load the data
Now that you have implemented a two-layer network that passes gradient checks and works on toy data, it's time to load up our favorite CIFAR-10 data so we can use it to train a classifier on a real dataset.
```
from cs231n.data_utils import load_CIFAR10
def get_CIFAR10_data(num_training=49000, num_validation=1000, num_test=1000):
"""
Load the CIFAR-10 dataset from disk and perform preprocessing to prepare
it for the two-layer neural net classifier. These are the same steps as
we used for the SVM, but condensed to a single function.
"""
# Load the raw CIFAR-10 data
cifar10_dir = 'cs231n/datasets/cifar-10-batches-py'
X_train, y_train, X_test, y_test = load_CIFAR10(cifar10_dir)
# Subsample the data
mask = range(num_training, num_training + num_validation)
X_val = X_train[mask]
y_val = y_train[mask]
mask = range(num_training)
X_train = X_train[mask]
y_train = y_train[mask]
mask = range(num_test)
X_test = X_test[mask]
y_test = y_test[mask]
# Normalize the data: subtract the mean image
mean_image = np.mean(X_train, axis=0)
X_train -= mean_image
X_val -= mean_image
X_test -= mean_image
# Reshape data to rows
X_train = X_train.reshape(num_training, -1)
X_val = X_val.reshape(num_validation, -1)
X_test = X_test.reshape(num_test, -1)
return X_train, y_train, X_val, y_val, X_test, y_test
# Invoke the above function to get our data.
X_train, y_train, X_val, y_val, X_test, y_test = get_CIFAR10_data()
print 'Train data shape: ', X_train.shape
print 'Train labels shape: ', y_train.shape
print 'Validation data shape: ', X_val.shape
print 'Validation labels shape: ', y_val.shape
print 'Test data shape: ', X_test.shape
print 'Test labels shape: ', y_test.shape
```
# Train a network
To train our network we will use SGD with momentum. In addition, we will adjust the learning rate with an exponential learning rate schedule as optimization proceeds; after each epoch, we will reduce the learning rate by multiplying it by a decay rate.
```
input_size = 32 * 32 * 3
hidden_size = 100
num_classes = 10
net = TwoLayerNet(input_size, hidden_size, num_classes)
# Train the network
stats = net.train(X_train, y_train, X_val, y_val,
num_iters=10000, batch_size=200,
learning_rate=1e-4, learning_rate_decay=0.95,
reg=0.4, verbose=True)
# Predict on the validation set
val_acc = (net.predict(X_val) == y_val).mean()
print 'Validation accuracy: ', val_acc
```
# Debug the training
With the default parameters we provided above, you should get a validation accuracy of about 0.29 on the validation set. This isn't very good.
One strategy for getting insight into what's wrong is to plot the loss function and the accuracies on the training and validation sets during optimization.
Another strategy is to visualize the weights that were learned in the first layer of the network. In most neural networks trained on visual data, the first layer weights typically show some visible structure when visualized.
```
# Plot the loss function and train / validation accuracies
plt.subplot(2, 1, 1)
plt.plot(stats['loss_history'])
plt.title('Loss history')
plt.xlabel('Iteration')
plt.ylabel('Loss')
plt.subplot(2, 1, 2)
plt.plot(stats['train_acc_history'], label='train')
plt.plot(stats['val_acc_history'], label='val')
plt.title('Classification accuracy history')
plt.xlabel('Epoch')
plt.ylabel('Clasification accuracy')
plt.show()
from cs231n.vis_utils import visualize_grid
# Visualize the weights of the network
def show_net_weights(net):
W1 = net.params['W1']
W1 = W1.reshape(32, 32, 3, -1).transpose(3, 0, 1, 2)
plt.imshow(visualize_grid(W1, padding=3).astype('uint8'))
plt.gca().axis('off')
plt.show()
show_net_weights(net)
```
# Tune your hyperparameters
**What's wrong?**. Looking at the visualizations above, we see that the loss is decreasing more or less linearly, which seems to suggest that the learning rate may be too low. Moreover, there is no gap between the training and validation accuracy, suggesting that the model we used has low capacity, and that we should increase its size. On the other hand, with a very large model we would expect to see more overfitting, which would manifest itself as a very large gap between the training and validation accuracy.
**Tuning**. Tuning the hyperparameters and developing intuition for how they affect the final performance is a large part of using Neural Networks, so we want you to get a lot of practice. Below, you should experiment with different values of the various hyperparameters, including hidden layer size, learning rate, numer of training epochs, and regularization strength. You might also consider tuning the learning rate decay, but you should be able to get good performance using the default value.
**Approximate results**. You should be aim to achieve a classification accuracy of greater than 48% on the validation set. Our best network gets over 52% on the validation set.
**Experiment**: You goal in this exercise is to get as good of a result on CIFAR-10 as you can, with a fully-connected Neural Network. For every 1% above 52% on the Test set we will award you with one extra bonus point. Feel free implement your own techniques (e.g. PCA to reduce dimensionality, or adding dropout, or adding features to the solver, etc.).
```
best_net = None # store the best model into this
#################################################################################
# TODO: Tune hyperparameters using the validation set. Store your best trained #
# model in best_net. #
# #
# To help debug your network, it may help to use visualizations similar to the #
# ones we used above; these visualizations will have significant qualitative #
# differences from the ones we saw above for the poorly tuned network. #
# #
# Tweaking hyperparameters by hand can be fun, but you might find it useful to #
# write code to sweep through possible combinations of hyperparameters #
# automatically like we did on the previous exercises. #
#################################################################################
best_net=net
#################################################################################
# END OF YOUR CODE #
#################################################################################
# visualize the weights of the best network
show_net_weights(best_net)
```
# Run on the test set
When you are done experimenting, you should evaluate your final trained network on the test set; you should get above 48%.
**We will give you extra bonus point for every 1% of accuracy above 52%.**
```
test_acc = (best_net.predict(X_test) == y_test).mean()
print 'Test accuracy: ', test_acc
```
| github_jupyter |
```
################### BLINKAH Lane Detection ###################
# shoutout Matt Hardwick from https://medium.com/@mrhwick/simple-lane-detection-with-opencv-bfeb6ae54ec0
# Loading an image into memory and cropping to a region of interest
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# reading in an image
image = mpimg.imread('sample_lane.jpg')
# printing out some stats and plotting the image
print('This image is:', type(image), 'with dimensions:', image.shape)
plt.imshow(image)
plt.show()
height = image.shape[0]
width = image.shape[1]
color_channels = image.shape[2]
# region_of_interest_vertices = [
# (0, height),
# (width / 2, height / 2),
# (width, height),]
# actual cropping of image, defining region_of_interest()
import numpy as np
import cv2
def region_of_interest(img, vertices):
# Define a blank matrix that matches the image height/width.
mask = np.zeros_like(img)
# Retrieve the number of color channels of the image.
channel_count = img.shape[2]
# Create a match color with the same color channel counts.
match_mask_color = (255,) * channel_count
# Fill inside the polygon
cv2.fillPoly(mask, vertices, match_mask_color)
# Returning the image only where mask pixels match
masked_image = cv2.bitwise_and(img, mask)
return masked_image
# runnning cropping function on our image
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
region_of_interest_vertices = [
(0, height),
(width / 2, height / 2),
(width, height),
]
image = mpimg.imread('sample_lane.jpg')
cropped_image = region_of_interest(
image,
np.array([region_of_interest_vertices], np.int32),
)
plt.figure()
plt.imshow(cropped_image)
plt.show()
# detecting edges in the cropped image
# grayscale conversion and canny edge detection using single intensity values as each pixel in the image
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import cv2
import math
def region_of_interest(img, vertices):
mask = np.zeros_like(img)
channel_count = img.shape[2]
match_mask_color = (255,) * channel_count
cv2.fillPoly(mask, vertices, match_mask_color)
masked_image = cv2.bitwise_and(img, mask)
return masked_image
region_of_interest_vertices = [
(0, height),
(width / 2, height / 2),
(width, height),
]
image = mpimg.imread('sample_lane.jpg')
cropped_image = region_of_interest(
image,
np.array([region_of_interest_vertices], np.int32),
)
plt.figure()
plt.imshow(cropped_image)
# Convert to grayscale here.
gray_image = cv2.cvtColor(cropped_image, cv2.COLOR_RGB2GRAY)
# Call Canny Edge Detection here.
cannyed_image = cv2.Canny(gray_image, 100, 200)
plt.figure()
plt.imshow(cannyed_image)
plt.show()
# image now contains single pixels indicative of edges as well as edge of region of interest UH OH!!!
# SOLUTION: Place region of interest cropping after Canny edge detection
# as shown below #
def region_of_interest(img, vertices):
mask = np.zeros_like(img)
match_mask_color = 255 # <-- This line altered for grayscale.
cv2.fillPoly(mask, vertices, match_mask_color)
masked_image = cv2.bitwise_and(img, mask)
return masked_image
region_of_interest_vertices = [
(0, height),
(width / 2, height / 2),
(width, height),
]
image = mpimg.imread('sample_lane.jpg')
plt.figure()
plt.imshow(image)
plt.show()
gray_image = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
cannyed_image = cv2.Canny(gray_image, 100, 200)
# Moved the cropping operation to the end of the pipeline.
cropped_image = region_of_interest(
cannyed_image,
np.array([region_of_interest_vertices], np.int32)
)
plt.figure()
plt.imshow(cropped_image)
plt.show()
# Generate lines from edge pixels using Hough transforms
image = mpimg.imread('sample_lane.jpg')
gray_image = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
cannyed_image = cv2.Canny(gray_image, 200, 300)
cropped_image = region_of_interest(
cannyed_image,
np.array(
[region_of_interest_vertices],
np.int32
),
)
lines = cv2.HoughLinesP(
cropped_image,
rho=6,
theta=np.pi / 60,
threshold=160,
lines=np.array([]),
minLineLength=40,
maxLineGap=25
)
print(lines)
# rendering detected Hough lines as an overlay
def draw_lines(img, lines, color=[255, 0, 0], thickness=3):
# If there are no lines to draw, exit.
if lines is None:
return
# Make a copy of the original image.
img = np.copy(img)
# Create a blank image that matches the original in size.
line_img = np.zeros(
(
height,
width,
color_channels
),
dtype=np.uint8,
)
# Loop over all lines and draw them on the blank image.
for line in lines:
for x1, y1, x2, y2 in line:
#cv2.line(line_img, (x1, y1), (x2, y2), color, thickness)
cv2.line(line_img, (int(x1), int(y1)), (int(x2), int(y2)), color, thickness)
# Merge the image with the lines onto the original.
img = cv2.addWeighted(image, 0.8, line_img, 1.0, 0.0)
# Return the modified image.
return img
image = mpimg.imread('sample_lane.jpg')
plt.figure()
plt.imshow(image)
plt.show()
gray_image = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
#gray_image = cv2.cvtColor(np.float32(imgUMat), cv2.COLOR_RGB2GRAY)
cannyed_image = cv2.Canny(gray_image, 100, 200)
cropped_image = region_of_interest(
cannyed_image,
np.array(
[region_of_interest_vertices],
np.int32
),
)
lines = cv2.HoughLinesP(
cropped_image,
rho=6,
theta=np.pi / 60,
threshold=160,
lines=np.array([]),
minLineLength=40,
maxLineGap=25
)
line_image = draw_lines(image, lines) # <---- Add this call.
plt.figure()
plt.imshow(line_image)
plt.show()
# create single linear represenatation of each line group (left & right)
left_line_x = []
left_line_y = []
right_line_x = []
right_line_y = []
for line in lines:
for x1, y1, x2, y2 in line:
slope = (y2 - y1) / (x2 - x1) # <-- Calculating the slope.
if math.fabs(slope) < 0.5: # <-- Only consider extreme slope
continue
if slope <= 0: # <-- If the slope is negative, left group.
left_line_x.extend([x1, x2])
left_line_y.extend([y1, y2])
else: # <-- Otherwise, right group.
right_line_x.extend([x1, x2])
right_line_y.extend([y1, y2])
min_y = image.shape[0] * (3 / 5) # <-- Just below the horizon
max_y = image.shape[0] # <-- The bottom of the image
poly_left = np.poly1d(np.polyfit(
left_line_y,
left_line_x,
deg=1
))
left_x_start = int(poly_left(max_y))
left_x_end = int(poly_left(min_y))
poly_right = np.poly1d(np.polyfit(
right_line_y,
right_line_x,
deg=1
))
right_x_start = int(poly_right(max_y))
right_x_end = int(poly_right(min_y))
image = mpimg.imread('sample_lane.jpg')
plt.figure()
plt.imshow(image)
gray_image = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
cannyed_image = cv2.Canny(gray_image, 100, 200)
cropped_image = region_of_interest(
cannyed_image,
np.array(
[region_of_interest_vertices],
np.int32
),
)
lines = cv2.HoughLinesP(
cropped_image,
rho=6,
theta=np.pi / 60,
threshold=160,
lines=np.array([]),
minLineLength=40,
maxLineGap=25
)
left_line_x = []
left_line_y = []
right_line_x = []
right_line_y = []
for line in lines:
for x1, y1, x2, y2 in line:
slope = (y2 - y1) / (x2 - x1) # <-- Calculating the slope.
if math.fabs(slope) < .5: # <-- Only consider extreme slope
continue
if slope <= 0: # <-- If the slope is negative, left group.
left_line_x.extend([x1, x2])
left_line_y.extend([y1, y2])
else: # <-- Otherwise, right group.
right_line_x.extend([x1, x2])
right_line_y.extend([y1, y2])
min_y = image.shape[0] * (3 / 5) # <-- Just below the horizon
max_y = image.shape[0] # <-- The bottom of the image
poly_left = np.poly1d(np.polyfit(
left_line_y,
left_line_x,
deg=1
))
left_x_start = int(poly_left(max_y))
left_x_end = int(poly_left(min_y))
poly_right = np.poly1d(np.polyfit(
right_line_y,
right_line_x,
deg=1
))
right_x_start = int(poly_right(max_y))
right_x_end = int(poly_right(min_y))
line_image = draw_lines(
image,
[[
[left_x_start, max_y, left_x_end, min_y],
[right_x_start, max_y, right_x_end, min_y],
]],
thickness=5,
)
plt.figure()
plt.imshow(line_image)
plt.show()
######################### Applying to video #########################
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import cv2
import math
def region_of_interest(img, vertices):
mask = np.zeros_like(img)
match_mask_color = 255
cv2.fillPoly(mask, vertices, match_mask_color)
masked_image = cv2.bitwise_and(img, mask)
return masked_image
def draw_lines(img, lines, color=[255, 0, 0], thickness=3):
line_img = np.zeros(
(
img.shape[0],
img.shape[1],
3
),
dtype=np.uint8
)
img = np.copy(img)
if lines is None:
return
for line in lines:
for x1, y1, x2, y2 in line:
cv2.line(line_img, (x1, y1), (x2, y2), color, thickness)
img = cv2.addWeighted(img, 0.8, line_img, 1.0, 0.0)
return img
def pipeline(image):
"""
An image processing pipeline which will output
an image with the lane lines annotated.
"""
height = image.shape[0]
width = image.shape[1]
region_of_interest_vertices = [
(0, height),
(width / 2, height / 2),
(width, height),
]
gray_image = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
cannyed_image = cv2.Canny(gray_image, 100, 200)
cropped_image = region_of_interest(
cannyed_image,
np.array(
[region_of_interest_vertices],
np.int32
),
)
lines = cv2.HoughLinesP(
cropped_image,
rho=6,
theta=np.pi / 60,
threshold=160,
lines=np.array([]),
minLineLength=40,
maxLineGap=25
)
left_line_x = []
left_line_y = []
right_line_x = []
right_line_y = []
for line in lines:
for x1, y1, x2, y2 in line:
slope = (y2 - y1) / (x2 - x1)
#if math.fabs(slope) < 0.5:
# continue
if slope <= 0:
left_line_x.extend([x1, x2])
left_line_y.extend([y1, y2])
else:
right_line_x.extend([x1, x2])
right_line_y.extend([y1, y2])
min_y = int(image.shape[0] * (3 / 5))
max_y = int(image.shape[0])
poly_left = np.poly1d(np.polyfit(
left_line_y,
left_line_x,
deg=1
))
left_x_start = int(poly_left(max_y))
left_x_end = int(poly_left(min_y))
poly_right = np.poly1d(np.polyfit(
right_line_y,
right_line_x,
deg=1
))
right_x_start = int(poly_right(max_y))
right_x_end = int(poly_right(min_y))
line_image = draw_lines(
image,
[[
[left_x_start, max_y, left_x_end, min_y],
[right_x_start, max_y, right_x_end, min_y],
]],
thickness=5,
)
return line_image
from moviepy.editor import VideoFileClip
from IPython.display import HTML
white_output = 'solidWhiteRight_output.mp4'
clip1 = VideoFileClip("solidWhiteRight.mp4")
white_clip = clip1.fl_image(pipeline)
white_clip.write_videofile(white_output, audio=False)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as pl
import xavierUtils as xu
import datetime as dt
%autosave 1000000
pd.to_datetime('2019-02-01') + pd.to_timedelta(99, unit='d')
today = dt.datetime.now().strftime("%Y-%m-%d")
print today
```
### Carregando dados
```
# Dados brutos sobre órgãos (comissões, etc.):
query = "SELECT * FROM `gabinete-compartilhado.camara_v2.deputados_orgaos`"
rorgaos = pd.read_gbq(query, project_id='gabinete-compartilhado',
dialect='standard',
private_key='../keys-configs/gabinete-compartilhado.json')
rorgaos['id_deputado'] = rorgaos.api_url.map(lambda s: int(s.split('/')[-2]))
# Ocupação de tipos de cargos (nossa classificação) e score de poder:
query = "SELECT * FROM `gabinete-compartilhado.analise_congresso_poder.camara_cargos_score`"
rcargos = pd.read_gbq(query, project_id='gabinete-compartilhado',
dialect='standard',
private_key='../keys-configs/gabinete-compartilhado.json')
# Ocupação de lideranças partidárias e de bloco:
query = "SELECT * FROM `gabinete-compartilhado.analise_congresso_poder.camara_liderancas_socre_`"
rlider = pd.read_gbq(query, project_id='gabinete-compartilhado',
dialect='standard',
private_key='../keys-configs/gabinete-compartilhado.json')
# Informações sobre os deputados:
query = "SELECT * FROM `gabinete-compartilhado.camara_v2.deputados_detalhes`"
rdep = pd.read_gbq(query, project_id='gabinete-compartilhado',
dialect='standard',
private_key='../keys-configs/gabinete-compartilhado.json')
# Informações sobre os deputados:
query = "SELECT * FROM `gabinete-compartilhado.camara_v2.deputados`"
rdep0 = pd.read_gbq(query, project_id='gabinete-compartilhado',
dialect='standard',
private_key='../keys-configs/gabinete-compartilhado.json')
# Informações sobre os deputados:
#query = "SELECT * FROM `gabinete-compartilhado.congresso.camara_deputado_`"
#rdep0 = pd.read_gbq(query, project_id='gabinete-compartilhado',
# dialect='standard',
# private_key='../keys-configs/gabinete-compartilhado.json')
columns(rdep0)
```
### Exploração
#### Base Cargos
```
print xu.Bold('-- Cargos --')
xu.unique(rcargos.columns)
rcargos.dtypes
xu.checkMissing(rcargos)
xu.mapUnique(rcargos)
```
#### Base Lideranças
```
print xu.Bold('-- Lideranças --')
xu.unique(rlider.columns)
rlider.dtypes
xu.checkMissing(rlider)
xu.mapUnique(rlider)
```
#### Base orgaos (bruta)
```
xu.unique(rorgaos.columns)
rdep.loc[rdep.id==204534]
```
#### Base deputados
```
xu.unique(rdep.columns)
xu.checkMissing(rdep)
# ATENÇÃO: Data de falecimento faltando é uma string vazia:
#rdep.sort_values('dataNascimento').dataFalecimento[1009]
```
### Pesos dos cargos e lideranças
```
pd.concat([rcargos[['cargo','score']].drop_duplicates(),rlider[['cargo','score']].drop_duplicates()])\
.sort_values('score', ascending=False).set_index('cargo', drop=True)
rlider[['cargo','score']].drop_duplicates().sort_values('score',ascending=False)
```
### Evolução temporal do número de cargos por deputado
```
# Função para selecionar cargos ativos num certo período:
def activeInPeriod(df, start, end):
if np.any(df.columns.values=='data_inicio') and np.any(df.columns.values=='data_fim'):
return df.loc[(df.data_fim>=start) & (df.data_inicio<=end)]
if np.any(df.columns.values=='dataInicio') and np.any(df.columns.values=='dataFim'):
return df.loc[(df.dataFim>=start) & (df.dataInicio<=end)]
if np.any(df.columns.values=='timestamp'):
return df.loc[(df.timestamp>=start) & (df.timestamp<=end)]
orgaosActive = activeInPeriod(rorgaos,'2019-02-01','2019-04-23').sort_values('id_deputado')
cargosActive = activeInPeriod(rcargos,'2019-02-01','2019-04-23').sort_values('id_deputado')
# Teste para verificar quais cargos e deputados aparecem nas bases:
#idList = orgaosActive.loc[orgaosActive.titulo!='Suplente'].id_deputado.unique()
#t=6
#cargosActive.loc[cargosActive.id_deputado==idList[t]][['id_deputado','cargo','score','data_inicio','data_fim']]
#orgaosActive.loc[(orgaosActive.titulo!='Suplente')&(orgaosActive.id_deputado==idList[t])]\
#[['id_deputado','siglaOrgao','titulo','dataInicio','dataFim']]
# Vimos que a CESPO - Comissão do Esporte - não é contabilizada na base cargos.
#rorgaos.loc[rorgaos.siglaOrgao=='CESPO']
def nCargosPreenchidos(ano):
return len(activeInPeriod(rcargos, str(ano)+'-02-01', str(ano)+'-04-16')[['id_deputado','cargo']])
anos = np.arange(1999,2020)
nCargosByAno = np.array([nCargosPreenchidos(a) for a in anos])
pl.plot(anos, nCargosByAno,'r-')
pl.plot(anos, nCargosByAno,'r.')
pl.xlim([1998.5,2019.5])
pl.xticks(np.arange(1999,2020,4))
pl.grid(axis='x')
pl.show()
y = nCargosByAno
x = anos
yIdx = pd.date_range(start=str(x[0])+'-'+str(1).zfill(2)+'-01', periods=len(x),freq='A-JAN')
yFull = pd.Series(y, index=yIdx)
from statsmodels.tsa.seasonal import seasonal_decompose
result = seasonal_decompose(yFull.iloc[:-1], model='additive', freq=4)
#xPeriod = np.arange(1999,2020,4) # Para grid no plot.
xPeriod = yIdx[::4]
fullTS = pd.Series(y,x, name='fullTimeSeries')
scale = 1
ymin = y[y>0].min()/scale
ymax = y[y>0].max()/scale
deltay = ymax-ymin
sMean = np.mean(y)/scale
fig = pl.figure(figsize=(10,7))
# Observed:
ax1 = pl.subplot(4,1,1)
pl.text(0.93,0.9,'Observado', horizontalalignment='right', verticalalignment='top', transform=pl.gca().transAxes,
fontsize=16)
pl.plot(yFull/scale)
pl.plot(yFull/scale, 'b.')
pl.ylim([ymin,ymax])
pl.gca().tick_params(labelsize=14)
pl.axhline(sMean,color='firebrick')
# Format x-axis:
pl.xticks(xPeriod)
pl.grid(axis='x', linestyle='--')
pl.gca().tick_params(labelbottom=False)
# Novas legislaturas:
#[pl.axvline(a, color='k') for a in anoLegislatura]
# Trend:
pl.subplot(4,1,2, sharex=ax1)
pl.text(0.93,0.9,u'Tendência', horizontalalignment='right', verticalalignment='top', transform=pl.gca().transAxes,
fontsize=16)
pl.plot(result.trend/scale)
pl.ylim([ymin,ymax])
pl.gca().tick_params(labelsize=14)
pl.axhline(sMean,color='firebrick')
# Format x-axis:
pl.xticks(xPeriod)
pl.grid(axis='x', linestyle='--')
pl.gca().tick_params(labelbottom=False)
# Novas legislaturas:
#[pl.axvline(a, color='k') for a in anoLegislatura]
# Seasonal:
pl.subplot(4,1,3, sharex=ax1)
pl.text(0.93,0.9,'Sazonalidade', horizontalalignment='right', verticalalignment='top', transform=pl.gca().transAxes,
fontsize=16)
pl.plot(result.seasonal/scale)
pl.ylim([-deltay/2, deltay/2])
pl.gca().tick_params(labelsize=14)
# Format x-axis:
pl.xticks(xPeriod)
pl.grid(axis='x', linestyle='--')
pl.gca().tick_params(labelbottom=False)
pl.axhline(0,color='firebrick')
# Novas legislaturas:
#[pl.axvline(a, color='k') for a in anoLegislatura]
# Residual:
pl.subplot(4,1,4, sharex=ax1)
pl.text(0.93,0.9,u'Resíduo', horizontalalignment='right', verticalalignment='top', transform=pl.gca().transAxes,
fontsize=16)
pl.plot(result.resid/scale)
pl.ylim([-deltay/2, deltay/2])
pl.gca().tick_params(labelsize=14)
# Format x-axis:
pl.xticks(xPeriod)
pl.grid(axis='x', linestyle='--')
pl.axhline(0,color='firebrick')
pl.xlabel('Ano', fontsize=16)
# Novas legislaturas:
#[pl.axvline(a, color='k') for a in anoLegislatura]
# Label x comum:
axComum = fig.add_subplot(111, frameon=False)
pl.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
axComum.set_ylabel(u'# de deputados com cargos nos 100 primeiros dias', fontsize=16, labelpad=15)
# Ajustes finais:
pl.gca().tick_params(labelsize=14)
pl.subplots_adjust(hspace=0.1)
#xu.saveFigWdate('graficos/cargos_sazonalidade.pdf')
pl.show()
def nOrgaosPreenchidos(ano):
return len(activeInPeriod(rorgaos.loc[rorgaos.titulo!='Suplente'],
str(ano)+'-02-01', str(ano)+'-04-18')[['id_deputado','siglaOrgao']])
xu.unique(rorgaos.columns)
xu.unique(rorgaos.nomeOrgao)
anos = np.arange(1990,2020)
nOrgaosByAno = np.array([nOrgaosPreenchidos(a) for a in anos])
pl.figure(figsize=(7,5))
pl.plot(anos, nOrgaosByAno,'r-')
pl.plot(anos, nOrgaosByAno,'r.')
pl.xticks(np.arange(2019,1990,-4))
pl.grid(axis='x', linestyle='--')
pl.gca().tick_params(labelsize=14)
pl.xlabel('Ano', fontsize=16)
pl.ylabel(u'# vagas ocupadas em órgãos', fontsize=16)
#xu.saveFigWdate('graficos/orgaos-ocupados-por-ano.pdf')
pl.show()
# CONCLUSÃO: O ano de 2019 está com problemas na base de dados. Vamos realizar a análise ignorando esse ano.
```
# Análise histórica
### Quantos dias dura o cargo
```
# Constrói base de dados histórica (antes de 2019) com número de dias no cargo:
rcargos['n_dias'] = (rcargos.data_fim - rcargos.data_inicio).dt.days
cargos_hist = rcargos.loc[(rcargos.data_fim<='2018-12-31') & (rcargos.data_inicio>='1999-02-01')]
# Existem datas erradas na base de dados:
rcargos.loc[rcargos.n_dias<0]
leg_ano_inicial = dict(zip(np.arange(56, 1, -1), np.arange(2019, 2019 - 4*(56-1), -4)))
nRows = 2
nCols = 2
pl.figure(figsize=(13,7))
for i in range(1,1+4):
pl.subplot(nRows,nCols,i)
# Seleciona legislatura:
leg = 51+i
ano = leg_ano_inicial[leg]
cargos_leg = activeInPeriod(cargos_hist, str(ano)+'-02-01', str(ano+3)+'-12-31')
# Texto:
pl.text(0.1,0.9,'Leg: '+str(leg),transform = pl.gca().transAxes, fontsize=14)
# Histograma:
ncargos_edges = np.arange(-36.5,1600,36.5)
pl.hist(cargos_leg.n_dias - 1, bins=ncargos_edges)
# Formatação:
pl.gca().tick_params(labelsize=14)
# Formatação do eixo x:
if (i-1) < nCols*(nRows-1):
pl.gca().tick_params(labelbottom=False)
else:
xlabel = '# dias na vaga'
if (nCols%2 == 0 and (i-1)%nCols == nCols/2 - 1):
pl.xlabel(xlabel, fontsize=16, position=(1,0), labelpad=14)
elif (nCols%2 == 1 and (i-1)%nCols == nCols/2):
pl.xlabel(xlabel, fontsize=16, labelpad=14)
# Formatação do eixo y:
pl.ylim([0,570])
if (i-1)%nCols!=0:
pl.gca().tick_params(labelleft=False)
else:
ylabel = u'# vagas ocupadas em órgãos'
if (nRows%2==0 and (i-1)/nCols == nRows/2):
pl.ylabel(ylabel, fontsize=16, position=(0,1))
elif (nRows%2==1 and (i-1)/nCols == nRows/2):
pl.ylabel(ylabel, fontsize=16)
pl.axvline(0,color='k')
pl.axvline(365,color='gray')
pl.axvline(2*365,color='gray')
pl.axvline(3*365,color='gray')
pl.axvline(4*365,color='k')
pl.subplots_adjust(wspace=0.02, hspace=0.05)
#xu.saveFigWdate('graficos/vagas_ocupadas_por_tempo_na_vaga.pdf')
pl.show()
# Cargos que ficam mais de 3 anos:
xu.unique(cargos_hist.loc[cargos_hist.n_dias>3*365]['cargo'])
```
## Distribuição por deputado
```
def columns(df):
xu.unique(df.columns)
columns(rdep)
rdep['idLegislatura'] = rdep.ultimoStatus.apply(lambda x: x[u'idLegislatura'])
len(rdep.groupby('id')['idLegislatura'].nunique())
np.sum(rdep.groupby('id')['idLegislatura'].nunique())
ano = 2015
activeInPeriod(cargos_hist, str(ano)+'-02-01', str(ano+3)+'-12-31').id_deputado
columns(rcargos)
leg = 55
ano = leg_ano_inicial[leg]
Ndeps_leg = len(rdep0.loc[(rdep0.idLegislaturaInicial<=leg) & (rdep0.idLegislaturaFinal>=leg)])
Ncargos_by_dep = activeInPeriod(cargos_hist, str(ano)+'-02-01', str(ano+3)+'-12-31').groupby('id_deputado').size()
Ndeps_zero_cargos = Ndeps_leg - len(Ncargos_by_dep)
rdep0.loc[(rdep0.idLegislaturaInicial<=leg) & (rdep0.idLegislaturaFinal>=leg)]
set(rset(Ncargos_by_dep.index)
```
# Lixo
### Distribuição de poder por deputados
#### Cargos em comissões
```
rdep0.loc[(rdep0.id<=165429.5)&(rdep0.id>=165428.5)]
rdep.merge(rcargos,how='outer',left_on='id',right_on='id_deputado')
activeInPeriod(rcargos,'2018-02-01','2018-04-25').groupby('id_deputado')['score'].size()
rdep.loc[rdep.id_deputado==178849]
rcargos.loc[rcargos.id_deputado==178849]
activeInPeriod(rcargos,'2018-01-01','2019-01-01')
```
#### Lideranças de partidos/blocos (só vale para 2019)
```
def scoreFinalLider(df, score0, gain):
bancadas = df.loc[df.total_membros.isnull()==False]
# Verifica se bancada mudou de tamanho:
Ncte = np.all(bancadas.groupby('sigla_bloco')['total_membros'].nunique()==1)
if Ncte==False:
raise Exception('Tamanho da bancada variou no período.')
# Calcula tamanho do bloco de referência:
ref = np.mean(bancadas[['sigla_bloco','total_membros']].drop_duplicates()).values[0]
# Calcula o score levando em conta o tamanho da bancada:
liderPartido = (df.id_cargo==3 * ~df.sigla_bloco.isin(['Minoria','Maioria','Governo',u'Oposição'])).astype(int)
scoreFinal = liderPartido*(score0 + gain * df.total_membros/ref * df.score).fillna(0) + (1-liderPartido)*df.score
# Retorna o resultado:
scoreFinal.name = 'scoreFinal'
return scoreFinal
rlider['scoreFinal'] = scoreFinalLider(rlider,0.3,1)
```
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import pycountry
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe():
return pd.read_csv("https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/owid-covid-data.csv")
confirmed_df_ = download_cases_dataframe()
confirmed_df_.iloc[0]
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["date", "new_cases", "iso_code"]]
confirmed_df.rename(
columns={
"date": "sample_date",
"iso_code": "country_code",
},
inplace=True)
def convert_iso_alpha_3_to_alpha_2(x):
try:
return pycountry.countries.get(alpha_3=x).alpha_2
except Exception as e:
logging.info(f"Error converting country ISO Alpha 3 code '{x}': {repr(e)}")
return None
confirmed_df["country_code"] = confirmed_df.country_code.apply(convert_iso_alpha_3_to_alpha_2)
confirmed_df.dropna(inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
confirmed_days_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
confirmed_days_df["sample_date_string"] = \
confirmed_days_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_days_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = report_backend_client.source_regions_for_date(
date=extraction_datetime.date())
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
report_source_regions
def get_cases_dataframe(source_regions_for_date_function, columns_suffix=None):
source_regions_at_date_df = confirmed_days_df.copy()
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: source_regions_for_date_function(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df["_source_regions_group"] = source_regions_at_date_df. \
source_regions_at_date.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_at_date_df.tail()
#%%
source_regions_for_summary_df_ = \
source_regions_at_date_df[["sample_date", "_source_regions_group"]].copy()
source_regions_for_summary_df_.rename(columns={"_source_regions_group": "source_regions"}, inplace=True)
source_regions_for_summary_df_.tail()
#%%
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group, source_regions_group_series in \
source_regions_at_date_df.groupby("_source_regions_group"):
source_regions_set = set(source_regions_group.split(","))
confirmed_source_regions_set_df = \
confirmed_df[confirmed_df.country_code.isin(source_regions_set)].copy()
confirmed_source_regions_group_df = \
confirmed_source_regions_set_df.groupby("sample_date").new_cases.sum() \
.reset_index().sort_values("sample_date")
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df.merge(
confirmed_days_df[["sample_date_string"]].rename(
columns={"sample_date_string": "sample_date"}),
how="right")
confirmed_source_regions_group_df["new_cases"] = \
confirmed_source_regions_group_df["new_cases"].clip(lower=0)
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df = confirmed_source_regions_group_df.replace(0, np.nan)
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[
confirmed_source_regions_group_df.sample_date.isin(
source_regions_group_series.sample_date_string)]
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
result_df = confirmed_output_df.copy()
result_df.tail()
#%%
result_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
result_df = confirmed_days_df[["sample_date_string"]].merge(result_df, how="left")
result_df.sort_values("sample_date_string", inplace=True)
result_df.fillna(method="ffill", inplace=True)
result_df.tail()
#%%
result_df[["new_cases", "covid_cases"]].plot()
if columns_suffix:
result_df.rename(
columns={
"new_cases": "new_cases_" + columns_suffix,
"covid_cases": "covid_cases_" + columns_suffix},
inplace=True)
return result_df, source_regions_for_summary_df_
confirmed_eu_df, source_regions_for_summary_df = get_cases_dataframe(
report_backend_client.source_regions_for_date)
confirmed_es_df, _ = get_cases_dataframe(
lambda date: [spain_region_country_code],
columns_suffix=spain_region_country_code.lower())
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
base_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=base_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_base_df = tek_list_df[tek_list_df.region == report_backend_identifier]
tek_list_base_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_base_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_base_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_base_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Official Statistics
```
import requests
import pandas.io.json
official_stats_response = requests.get("https://radarcovid.covid19.gob.es/kpi/statistics/basics")
official_stats_response.raise_for_status()
official_stats_df_ = pandas.io.json.json_normalize(official_stats_response.json())
official_stats_df = official_stats_df_.copy()
official_stats_df["date"] = pd.to_datetime(official_stats_df["date"], dayfirst=True)
official_stats_df.head()
official_stats_column_map = {
"date": "sample_date",
"applicationsDownloads.totalAcummulated": "app_downloads_es_accumulated",
"communicatedContagions.totalAcummulated": "shared_diagnoses_es_accumulated",
}
accumulated_suffix = "_accumulated"
accumulated_values_columns = \
list(filter(lambda x: x.endswith(accumulated_suffix), official_stats_column_map.values()))
interpolated_values_columns = \
list(map(lambda x: x[:-len(accumulated_suffix)], accumulated_values_columns))
official_stats_df = \
official_stats_df[official_stats_column_map.keys()] \
.rename(columns=official_stats_column_map)
official_stats_df["extraction_date"] = extraction_date
official_stats_df.head()
official_stats_path = "Data/Statistics/Current/RadarCOVID-Statistics.json"
previous_official_stats_df = pd.read_json(official_stats_path, orient="records", lines=True)
previous_official_stats_df["sample_date"] = pd.to_datetime(previous_official_stats_df["sample_date"], dayfirst=True)
official_stats_df = official_stats_df.append(previous_official_stats_df)
official_stats_df.head()
official_stats_df = official_stats_df[~(official_stats_df.shared_diagnoses_es_accumulated == 0)]
official_stats_df.sort_values("extraction_date", ascending=False, inplace=True)
official_stats_df.drop_duplicates(subset=["sample_date"], keep="first", inplace=True)
official_stats_df.head()
official_stats_stored_df = official_stats_df.copy()
official_stats_stored_df["sample_date"] = official_stats_stored_df.sample_date.dt.strftime("%Y-%m-%d")
official_stats_stored_df.to_json(official_stats_path, orient="records", lines=True)
official_stats_df.drop(columns=["extraction_date"], inplace=True)
official_stats_df = confirmed_days_df.merge(official_stats_df, how="left")
official_stats_df.sort_values("sample_date", ascending=False, inplace=True)
official_stats_df.head()
official_stats_df[accumulated_values_columns] = \
official_stats_df[accumulated_values_columns] \
.astype(float).interpolate(limit_area="inside")
official_stats_df[interpolated_values_columns] = \
official_stats_df[accumulated_values_columns].diff(periods=-1)
official_stats_df.drop(columns="sample_date", inplace=True)
official_stats_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
official_stats_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_eu_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df = confirmed_es_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(result_summary_df.shared_diagnoses_es / result_summary_df.covid_cases_es).fillna(0)
result_summary_df.head(daily_plot_days)
def compute_aggregated_results_summary(days) -> pd.DataFrame:
aggregated_result_summary_df = result_summary_df.copy()
aggregated_result_summary_df["covid_cases_for_ratio"] = \
aggregated_result_summary_df.covid_cases.mask(
aggregated_result_summary_df.shared_diagnoses == 0, 0)
aggregated_result_summary_df["covid_cases_for_ratio_es"] = \
aggregated_result_summary_df.covid_cases_es.mask(
aggregated_result_summary_df.shared_diagnoses_es == 0, 0)
aggregated_result_summary_df = aggregated_result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(days).agg({
"covid_cases": "sum",
"covid_cases_es": "sum",
"covid_cases_for_ratio": "sum",
"covid_cases_for_ratio_es": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum",
"shared_diagnoses_es": "sum",
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
aggregated_result_summary_df = aggregated_result_summary_df.fillna(0).astype(int)
aggregated_result_summary_df["teks_per_shared_diagnosis"] = \
(aggregated_result_summary_df.shared_teks_by_upload_date /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case"] = \
(aggregated_result_summary_df.shared_diagnoses /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(aggregated_result_summary_df.shared_diagnoses_es /
aggregated_result_summary_df.covid_cases_for_ratio_es).fillna(0)
return aggregated_result_summary_df
aggregated_result_with_7_days_window_summary_df = compute_aggregated_results_summary(days=7)
aggregated_result_with_7_days_window_summary_df.head()
last_7_days_summary = aggregated_result_with_7_days_window_summary_df.to_dict(orient="records")[1]
last_7_days_summary
aggregated_result_with_14_days_window_summary_df = compute_aggregated_results_summary(days=13)
last_14_days_summary = aggregated_result_with_14_days_window_summary_df.to_dict(orient="records")[1]
last_14_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases (Source Countries)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date (Source Countries)",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date (Source Countries)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date (Source Countries)",
"shared_diagnoses": "Shared Diagnoses (Source Countries – Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis (Source Countries)",
"shared_diagnoses_per_covid_case": "Usage Ratio (Source Countries)",
"covid_cases_es": "COVID-19 Cases (Spain)",
"app_downloads_es": "App Downloads (Spain – Official)",
"shared_diagnoses_es": "Shared Diagnoses (Spain – Official)",
"shared_diagnoses_per_covid_case_es": "Usage Ratio (Spain)",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
"covid_cases_es",
"app_downloads_es",
"shared_diagnoses_es",
"shared_diagnoses_per_covid_case_es",
]
summary_percentage_columns= [
"shared_diagnoses_per_covid_case_es",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 30), legend=False)
ax_ = summary_ax_list[0]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
for percentage_column in summary_percentage_columns:
percentage_column_index = summary_columns.index(percentage_column)
summary_ax_list[percentage_column_index].yaxis \
.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case_es"]: lambda x: f"{x:.2%}" if x != 0 else "",
}
general_columns = \
list(filter(lambda x: x not in display_formatters, display_column_name_mapping.values()))
general_formatter = lambda x: f"{x}" if x != 0 else ""
display_formatters.update(dict(map(lambda x: (x, general_formatter), general_columns)))
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.item()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.item()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.item()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.item()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.item()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.item()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
df = df.copy()
df_styler = df.style.format(display_formatters)
media_path = get_temporary_image_path()
dfi.export(df_styler, media_path)
return media_path
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
def generate_summary_api_results(df: pd.DataFrame) -> list:
api_df = df.reset_index().copy()
api_df["sample_date_string"] = \
api_df["sample_date"].dt.strftime("%Y-%m-%d")
api_df["source_regions"] = \
api_df["source_regions"].apply(lambda x: x.split(","))
return api_df.to_dict(orient="records")
summary_api_results = \
generate_summary_api_results(df=result_summary_df)
today_summary_api_results = \
generate_summary_api_results(df=extraction_date_result_summary_df)[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_api_results,
last_7_days=last_7_days_summary,
last_14_days=last_14_days_summary,
daily_results=summary_api_results)
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
def format_shared_diagnoses_per_covid_case(value) -> str:
if value == 0:
return "–"
return f"≤{value:.2%}"
display_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=shared_diagnoses_per_covid_case)
display_last_14_days_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case"])
display_last_14_days_shared_diagnoses_per_covid_case_es = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case_es"])
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: {display_shared_diagnoses_per_covid_case}
Last 14 Days:
- Usage Ratio (Estimation): {display_last_14_days_shared_diagnoses_per_covid_case}
- Usage Ratio (Official): {display_last_14_days_shared_diagnoses_per_covid_case_es}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
```
# dependencies
import tensorflow as tf
import numpy as np
from sklearn.cross_validation import train_test_split
import time
import matplotlib.pyplot as plt
import pickle
import codecs
def read_dataset(filepath):
with open(filepath, 'rb') as fp:
return pickle.load(fp)
# read dataset
dataset_location = "./data.p"
X, Y, l1_word2idx, l1_idx2word, l1_vocab, l2_word2idx, l2_idx2word, l2_vocab = read_dataset(dataset_location)
input_seq_len = 20
output_seq_len = 22
l1_vocab_size = len(l1_vocab) + 2 # + <pad>, <ukn>
l2_vocab_size = len(l2_vocab) + 4 # + <pad>, <ukn>, <eos>, <go>
# let's define some helper functions
# simple softmax function
def softmax(x):
n = np.max(x)
e_x = np.exp(x - n)
return e_x / e_x.sum()
# feed data into placeholders
def feed_dict(x, y, batch_size = 64):
feed = {}
idxes = np.random.choice(len(x), size = batch_size, replace = False)
for i in range(input_seq_len):
feed[encoder_inputs[i].name] = np.array([x[j][i] for j in idxes])
for i in range(output_seq_len):
feed[decoder_inputs[i].name] = np.array([y[j][i] for j in idxes])
feed[targets[len(targets)-1].name] = np.full(shape = [batch_size], fill_value = l2_word2idx['<pad>'])
for i in range(output_seq_len-1):
batch_weights = np.ones(batch_size, dtype = np.float32)
target = feed[decoder_inputs[i+1].name]
for j in range(batch_size):
if target[j] == l2_word2idx['<pad>']:
batch_weights[j] = 0.0
feed[target_weights[i].name] = batch_weights
feed[target_weights[output_seq_len-1].name] = np.zeros(batch_size, dtype = np.float32)
return feed
# decode output sequence
def decode_output(output_seq):
words = []
for i in range(output_seq_len):
smax = softmax(output_seq[i])
idx = np.argmax(smax)
words.append(l2_idx2word[idx])
return words
def data_padding(x, l1_word2idx, length = 20):
for i in range(len(x)):
x[i] = x[i] + (length - len(x[i])) * [l1_word2idx['<pad>']]
return x
def translate_model(sentences):
result = []
# read dataset
dataset_location = "./data.p"
X, Y, l1_word2idx, l1_idx2word, l1_vocab, l2_word2idx, l2_idx2word, l2_vocab = read_dataset(dataset_location)
with tf.Graph().as_default():
# placeholders
encoder_inputs = [tf.placeholder(dtype = tf.int32, shape = [None], name = 'encoder{}'.format(i)) for i in range(input_seq_len)]
decoder_inputs = [tf.placeholder(dtype = tf.int32, shape = [None], name = 'decoder{}'.format(i)) for i in range(output_seq_len)]
# output projection
size = 512
w_t = tf.get_variable('proj_w', [l2_vocab_size, size], tf.float32)
b = tf.get_variable('proj_b', [l2_vocab_size], tf.float32)
w = tf.transpose(w_t)
output_projection = (w, b)
# change the model so that output at time t can be fed as input at time t+1
outputs, states = tf.nn.seq2seq.embedding_attention_seq2seq(
encoder_inputs,
decoder_inputs,
tf.nn.rnn_cell.BasicLSTMCell(size),
num_encoder_symbols = l1_vocab_size,
num_decoder_symbols = l2_vocab_size,
embedding_size = 80,
feed_previous = True, # <-----this is changed----->
output_projection = output_projection,
dtype = tf.float32)
# ops for projecting outputs
outputs_proj = [tf.matmul(outputs[i], output_projection[0]) + output_projection[1] for i in range(output_seq_len)]
sentences = [[l1_word2idx.get(word.strip(',." ;:)(|][?!<>'), 0) for word in sentence.split(' ')] for sentence in sentences]
encoded_sentences = data_padding(sentences, l1_word2idx)
# restore all variables - use the last checkpoint saved
saver = tf.train.Saver()
path = tf.train.latest_checkpoint('./checkpoints/')
with tf.Session() as sess:
# restore
saver.restore(sess, path)
# feed data into placeholders
feed = {}
for i in range(input_seq_len):
feed[encoder_inputs[i].name] = np.array([encoded_sentences[j][i] for j in range(len(encoded_sentences))])
feed[decoder_inputs[0].name] = np.array([l2_word2idx['<go>']] * len(encoded_sentences))
# translate
output_sequences = sess.run(outputs_proj, feed_dict = feed)
for i in range(len(encoded_sentences)):
ouput_seq = [output_sequences[j][i] for j in range(output_seq_len)]
#decode output sequence
words = decode_output(ouput_seq)
temp = """"""
for i in range(len(words)):
if words[i] not in ['<eos>', '<pad>', '<go>']:
temp += words[i] + " "
result.append(temp.strip())
return result
test = [ "आपका नाम क्या है", "यह कौन है", "कल शनिवार है", "बच्चे पानी प्यार करते हैं", "पानी पिएं", "अधिक नृत्य करें"]
result = translate_model(test)
print(result)
```
| github_jupyter |
# Paso 4.- Datos Holiday
## Determinar la variable de festividad de un día
Determinar la variable de festividad de un día:
- laborables = 0
- Sábados = 0.75
- Domingos y festivos = 1
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
from esiosdata import PVPC
from esiosdata.prettyprinting import *
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from datetime import datetime
import time
data_total = pd.read_csv("data_total.csv")
data_total.head()
#Some cleaning
data_total = data_total.loc[:, ~data_total.columns.str.contains('^Unnamed')]
data_total = data_total.drop(columns=['date_x', 'date_y', 'date.1'])
data_total['date_datetime'] = pd.to_datetime(data_total['fecha'])
data_total.head()
```
## Creación del calendario festivo de españa
```
from pandas.tseries.holiday import Holiday,AbstractHolidayCalendar
from pandas.tseries.offsets import CustomBusinessDay
class EsBusinessCalendar(AbstractHolidayCalendar):
rules = [
Holiday('Año Nuevo', month=1, day=1, observance=nearest_workday),
Holiday('Epifanía del Señor', month=1, day=6, observance=nearest_workday),
Holiday('Viernes Santo', month=1, day=1, offset=[Easter(), Day(-2)]),
Holiday('Día del Trabajador', month=5, day=1, observance=nearest_workday),
Holiday('Asunción de la Virgen', month=8, day=15, observance=nearest_workday),
Holiday('Día de la Hispanidad', month=10, day=12, observance=nearest_workday),
Holiday('Todos los Santos', month=11, day=1, observance=nearest_workday),
Holiday('Día Constitución', month=12, day=6, observance=nearest_workday),
Holiday('Inmaculada Concepción', month=12, day=8, observance=nearest_workday),
Holiday('Navidad', month=12, day=25, observance=nearest_workday)
]
cal = EsBusinessCalendar()
holidays = cal.holidays(start=data_total['fecha'].min(),
end=data_total['fecha'].max()).to_pydatetime()
holidays_array_by_date = list()
for holiday in holidays:
holidays_array_by_date.append(holiday.strftime('%Y-%m-%d'))
print(holidays_array_by_date)
from datetime import date
def is_holiday(df):
for i, row in df.iterrows():
holiday_value = 0
if row['date_datetime'].weekday() == 5:
holiday_value = 0.75
elif row['date_datetime'].weekday() == 6:
holiday_value = 1
else:
if row['date_datetime'] in holidays_array_by_date:
holiday_value = 1
df.set_value(i,'Holiday',holiday_value)
is_holiday(data_total)
data_total = data_total.drop(columns=['date_datetime'])
data_total.tail()
data_total[data_total['Holiday'] == 1]
data_total.to_csv('data_total.csv')
```
| github_jupyter |
# Configuration
NOTES: The warnings after the import are referred to the fact that Tensorflow 2.x versions are built to directly look for a GPU in the system. The warning can be forgot if you are not going to use the GPU.
```
!source myenv/bin/activate
import os
import librosa
import numpy as np
from tqdm.notebook import tqdm
import pandas as pd
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
import seaborn as sns
sns.set_style('whitegrid')
import IPython.display as ipd
import librosa.display
import numpy as np
import pickle
import scipy
import ipywidgets
import math
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import StandardScaler, LabelEncoder
from sklearn.pipeline import make_pipeline
from sklearn.cluster import KMeans
from sklearn.metrics import accuracy_score, confusion_matrix
from scipy.cluster.hierarchy import dendrogram
from sklearn.cluster import AgglomerativeClustering
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import KFold, StratifiedKFold
from tqdm import tqdm
import tensorflow as tf
from tensorflow.keras.layers import Dense, Dropout, Conv2D, AveragePooling1D, MaxPooling2D, Flatten
from tensorflow.keras.optimizers import SGD, Adam
from tensorflow.keras.models import Sequential
from tensorflow.keras import layers
from tensorflow.keras import regularizers
# from livelossplot import PlotLossesKeras
tf.config.list_physical_devices('GPU')
```
# Utils
# Compute dataframes for datasets and split in Train, Val, Test
```
main_path = '/media/helemanc/OS/Users/i2CAT/Desktop/Datasets SER/'
TESS = os.path.join(main_path, "tess/TESS Toronto emotional speech set data/")
RAV = os.path.join(main_path, "ravdess-emotional-speech-audio/audio_speech_actors_01-24")
SAVEE = os.path.join(main_path, "savee/ALL/")
CREMA = os.path.join(main_path, "creamd/AudioWAV/")
lst = []
emotion = []
voc_channel = []
full_path = []
modality = []
intensity = []
actors = []
phrase =[]
for root, dirs, files in tqdm(os.walk(RAV)):
for file in files:
try:
#Load librosa array, obtain mfcss, store the file and the mfcss information in a new array
# X, sample_rate = librosa.load(os.path.join(root,file), res_type='kaiser_fast')
# mfccs = np.mean(librosa.feature.mfcc(y=X, sr=sample_rate, n_mfcc=40).T,axis=0)
# The instruction below converts the labels (from 1 to 8) to a series from 0 to 7
# This is because our predictor needs to start from 0 otherwise it will try to predict also 0.
modal = int(file[1:2])
vchan = int(file[4:5])
lab = int(file[7:8])
ints = int(file[10:11])
phr = int(file[13:14])
act = int(file[18:20])
# arr = mfccs, lab
# lst.append(arr)
modality.append(modal)
voc_channel.append(vchan)
emotion.append(lab) #only labels
intensity.append(ints)
phrase.append(phr)
actors.append(act)
full_path.append((root, file)) # only files
# If the file is not valid, skip it
except ValueError:
continue
# 01 = neutral, 02 = calm, 03 = happy, 04 = sad, 05 = angry, 06 = fearful, 07 = disgust, 08 = surprised
# merge neutral and calm
emotions_list = ['neutral', 'neutral', 'happy', 'sadness', 'angry', 'fear', 'disgust', 'surprise']
emotion_dict = {em[0]+1:em[1] for em in enumerate(emotions_list)}
df = pd.DataFrame([emotion, voc_channel, modality, intensity, actors, actors,phrase, full_path]).T
df.columns = ['emotion', 'voc_channel', 'modality', 'intensity', 'actors', 'gender', 'phrase', 'path']
df['emotion'] = df['emotion'].map(emotion_dict)
df['voc_channel'] = df['voc_channel'].map({1: 'speech', 2:'song'})
df['modality'] = df['modality'].map({1: 'full AV', 2:'video only', 3:'audio only'})
df['intensity'] = df['intensity'].map({1: 'normal', 2:'strong'})
df['actors'] = df['actors']
df['gender'] = df['actors'].apply(lambda x: 'female' if x%2 == 0 else 'male')
df['phrase'] = df['phrase'].map({1: 'Kids are talking by the door', 2:'Dogs are sitting by the door'})
df['path'] = df['path'].apply(lambda x: x[0] + '/' + x[1])
# remove files with noise to apply the same noise to all files for data augmentation
df = df[~df.path.str.contains('noise')]
# only speech
RAV_df = df
RAV_df = RAV_df.loc[RAV_df.voc_channel == 'speech']
RAV_df.insert(0, "emotion_label", RAV_df.emotion, True)
RAV_df = RAV_df.drop(['emotion', 'voc_channel', 'modality', 'intensity', 'phrase'], 1)
RAV_train = []
RAV_val = []
RAV_test = []
for index, row in RAV_df.iterrows():
if row['actors'] in range(1,21):
RAV_train.append(row)
elif row['actors'] in range(21,23):
RAV_val.append(row)
elif row['actors'] in range(23,25):
RAV_test.append(row)
len(RAV_train), len(RAV_val), len(RAV_test)
RAV_train = pd.DataFrame(RAV_train)
RAV_val = pd.DataFrame(RAV_val)
RAV_test = pd.DataFrame(RAV_test)
RAV_train = RAV_train.drop(['actors'], 1)
RAV_val = RAV_val.drop(['actors'], 1)
RAV_test = RAV_test.drop(['actors'], 1)
df_train = RAV_train.reset_index(drop=True)
df_val = RAV_val.reset_index(drop=True)
df_test = RAV_test.reset_index(drop=True)
df_train.head()
```
# Create Noise Files
```
from pydub import AudioSegment
import random
from pydub.utils import make_chunks
def create_noise_files(df_train, df_val, df_test):
'''
Apply noise only on training files, so double the number of training files and keep
validation and test the same
'''
path_noise_sound_1 = '/home/helemanc/Desktop/Binary_Model/noise_sounds/freight_train.wav'
path_noise_sound_2 = '/home/helemanc/Desktop/Binary_Model/noise_sounds/inside_train.wav'
path_noise_sound_3 = '/home/helemanc/Desktop/Binary_Model/noise_sounds/small_crowd.wav'
path_noise_dataset_train = '/home/helemanc/Desktop/Binary_Model/noise_datasets/ravdess/train'
#path_noise_dataset_val = '/home/helemanc/Desktop/Binary_Model/noise_datasets/ravdess/val'
#path_noise_dataset_test = '/home/helemanc/Desktop/Binary_Model/noise_datasets/ravdess/test'
#df_list = [df_train, df_val, df_test]
#count_df = 0
train_emotions = []
train_genders = []
train_paths = []
#val_emotions = []
#val_genders = []
#val_paths = []
#test_emotions = []
#test_genders = []
#test_paths = []
#for df in df_list:
for index, row in tqdm(df_train.iterrows()):
path = row['path']
sound1 = AudioSegment.from_file(path)
samples, sr = librosa.load(path, res_type='kaiser_fast', sr=16000)
duration = librosa.get_duration(y = samples, sr = sr)
# pick a noise sound file randomly
noise_list = [path_noise_sound_1, path_noise_sound_2, path_noise_sound_3]
random_noise = random.choice(noise_list)
lower_volume = 0
# adjust volume to not cover the voice of the audio file
# warning: different levels of dB need to be calibrate for each dataset
'''
if random_noise == path_noise_sound_1:
lower_volume = 40
elif random_noise == path_noise_sound_2:
lower_volume = 25
else:
lower_volume = 40
'''
# other strategy:
# compute db of both files, compute the difference, and lower the volume of the file to make it
# a bit lower than the original file -almost equal-
sound2 = AudioSegment.from_file(random_noise)
# make chunks of duration equal to the audio file
chunk_length_ms = duration*1000 #ms
chunks = make_chunks(sound2, chunk_length_ms)
# pick a random chunk
random_chunk = random.choice(chunks)
difference = random_chunk.dBFS - sound1.dBFS
abs_difference = abs(difference)
lower = random_chunk - abs_difference - 2
# lower the volume of the noise file to be overlayed with the voice_sound
#lower = random_chunk - lower_volume
combined = sound1.overlay(lower)
parts = path.split('/')
fname = parts[-1]
new_path = path_noise_dataset_train + '/' + fname
train_emotions.append(row['emotion_label'])
train_genders.append(row['gender'])
train_paths.append(new_path)
'''
if count_df == 0:
new_path = path_noise_dataset_train + '/' + fname
train_emotions.append(row['emotion_label'])
train_genders.append(row['gender'])
train_paths.append(new_path)
elif count_df == 1:
new_path = path_noise_dataset_val + '/' + fname
val_emotions.append(row['emotion_label'])
val_genders.append(row['gender'])
val_paths.append(new_path)
elif count_df == 2:
new_path = path_noise_dataset_test + '/' + fname
test_emotions.append(row['emotion_label'])
test_genders.append(row['gender'])
test_paths.append(new_path)
'''
combined.export(new_path, format= 'wav')
#count_df +=1
df_train_noise = pd.DataFrame([train_emotions, train_genders, train_paths]).T
df_train_noise.columns = ['emotion_label', 'gender', 'path']
#df_val_noise = pd.DataFrame([val_emotions, val_genders, val_paths]).T
#df_val_noise.columns = ['emotion_label', 'gender', 'path']
#df_test_noise = pd.DataFrame([test_emotions, test_genders, test_paths]).T
#df_test_noise.columns = ['emotion_label', 'gender', 'path']
df_train_combined = pd.concat([df_train, df_train_noise])
df_train_combined.reset_index(drop=True, inplace=True)
#df_val_combined = pd.concat([df_val, df_val_noise])
#df_val_combined.reset_index(drop=True, inplace=True)
#df_test_combined = pd.concat([df_test, df_test_noise])
#df_test_combined.reset_index(drop=True, inplace=True)
return df_train_combined, df_val, df_test
# have to save df
new_df_train, new_df_val, new_df_test = create_noise_files(df_train, df_val, df_test)
new_df_train.shape, new_df_val.shape, new_df_test.shape
```
## Save dataframes
```
preprocess_path = "/home/helemanc/Desktop/Binary_Model/df_csv_noise/ravdess"
new_df_train.to_csv(os.path.join(preprocess_path,"df_train.csv"), index=False)
new_df_val.to_csv(os.path.join(preprocess_path,"df_val.csv"), index=False)
new_df_test.to_csv(os.path.join(preprocess_path,"df_test.csv"), index=False)
```
## Trial Code
```
path_noise_sound_1 = '/home/helemanc/Desktop/Binary_Model/noise_sounds/freight_train.wav'
path_noise_sound_2 = '/home/helemanc/Desktop/Binary_Model/noise_sounds/inside_train.wav'
path_noise_sound_3 = '/home/helemanc/Desktop/Binary_Model/noise_sounds/small_crowd.wav'
path_noise_dataset = '/home/helemanc/Desktop/Binary_Model/noise_datasets/ravdess/train'
# load a file
from pydub import AudioSegment
import random
from pydub.utils import make_chunks
sound1 = AudioSegment.from_file(RAV_df.path[0])
samples, sr = librosa.load(RAV_df.path[0], res_type='kaiser_fast', sr=16000)
duration = librosa.get_duration(y = samples, sr = sr)
noise_list = [path_noise_sound_1, path_noise_sound_2, path_noise_sound_3]
lower_volume = 0
random_noise = random.choice(noise_list)
'''
if random_noise == path_noise_sound_1:
lower_volume = 30
elif random_noise == path_noise_sound_2:
lower_volume = 25
else:
lower_volume = 40
'''
sound2 = AudioSegment.from_file(random_noise)
chunk_length_ms = duration*1000 #ms
chunks = make_chunks(sound2, chunk_length_ms) # divide the audio file to the original length
random_chunk = random.choice(chunks)
#lower = random_chunk - lower_volume
print(random_chunk.dBFS)
difference = random_chunk.dBFS - sound1.dBFS
abs_difference = abs(difference)
print(abs_difference)
lower = random_chunk - abs_difference -2
print(lower.dBFS)
combined = sound1.overlay(lower)
parts = RAV_df.path[0].split('/')
fname = parts[-1]
new_path = path_noise_dataset + '/' + fname
combined.export(new_path, format= 'wav')
print(sound1.dBFS)
```
| github_jupyter |
# Lesson 2 - Image Classification Models from Scratch
## Lesson Video:
```
#hide_input
from IPython.lib.display import YouTubeVideo
YouTubeVideo('_SKqrTlXNt8')
#hide
#Run once per session
!pip install fastai wwf -q --upgrade
#hide_input
from wwf.utils import state_versions
state_versions(['fastai', 'fastcore', 'wwf'])
```
Grab our vision related libraries
```
from fastai.vision.all import *
```
Below you will find the exact imports for everything we use today
```
from torch import nn
from fastai.callback.hook import summary
from fastai.callback.schedule import fit_one_cycle, lr_find
from fastai.callback.progress import ProgressCallback
from fastai.data.core import Datasets, DataLoaders, show_at
from fastai.data.external import untar_data, URLs
from fastai.data.transforms import Categorize, GrandparentSplitter, parent_label, ToTensor, IntToFloatTensor, Normalize
from fastai.layers import Flatten
from fastai.learner import Learner
from fastai.metrics import accuracy, CrossEntropyLossFlat
from fastai.vision.augment import CropPad, RandomCrop, PadMode
from fastai.vision.core import PILImageBW
from fastai.vision.utils import get_image_files
```
And our data
```
path = untar_data(URLs.MNIST)
```
## Working with the data
```
items = get_image_files(path)
items[0]
```
Create an image object. Done automatically with `ImageBlock`.
```
im = PILImageBW.create(items[0])
im.show()
```
Split our data with `GrandparentSplitter`, which will make use of a `train` and `valid` folder.
```
splits = GrandparentSplitter(train_name='training', valid_name='testing')
items[:3]
```
Splits need to be applied to some items
```
splits = splits(items)
splits[0][:5], splits[1][:5]
```
* Make a `Datasets`
* Expects items, transforms for describing our problem, and a splitting method
```
dsrc = Datasets(items, tfms=[[PILImageBW.create], [parent_label, Categorize]],
splits=splits)
```
We can look at an item in our `Datasets` with `show_at`
```
show_at(dsrc.train, 3)
```
We can see that it's a `PILImage` of a three, along with a label of `3`
Next we need to give ourselves some transforms on the data! These will need to:
1. Ensure our images are all the same size
2. Make sure our output are the `tensor` our models are wanting
3. Give some image augmentation
```
tfms = [ToTensor(), CropPad(size=34, pad_mode=PadMode.Zeros), RandomCrop(size=28)]
```
* `ToTensor`: Converts to tensor
* `CropPad` and `RandomCrop`: Resizing transforms
* Applied on the `CPU` via `after_item`
```
gpu_tfms = [IntToFloatTensor(), Normalize()]
```
* `IntToFloatTensor`: Converts to a float
* `Normalize`: Normalizes data
```
dls = dsrc.dataloaders(bs=128, after_item=tfms, after_batch=gpu_tfms)
```
And show a batch
```
dls.show_batch()
```
From here we need to see what our model will expect
```
xb, yb = dls.one_batch()
```
And now the shapes:
```
xb.shape, yb.shape
dls.c
```
So our input shape will be a [128 x 1 x 28 x 28] and our output shape will be a [128] tensor that we need to condense into 10 classes
## The Model
Our models are made up of **layers**, and each layer represents a matrix multiplication to end up with our final `y`. For this image problem, we will use a **Convolutional layer**, a **Batch Normalization layer**, an **Activation Function**, and a **Flattening layer**
### Convolutional Layer
These are always the first layer in our network. I will be borrowing an analogy from [here](https://adeshpande3.github.io/A-Beginner%27s-Guide-To-Understanding-Convolutional-Neural-Networks/) by Adit Deshpande.
Our example Convolutional layer will be 5x5x1
Imagine a flashlight that is shining over the top left of an image, which covers a 5x5 section of pixels at one given moment. This flashlight then slides crosses our pixels at all areas in the picture. This flashlight is called a **filter**, which can also be called a **neuron** or **kernel**. The region it is currently looking over is called a **receptive field**. This filter is also an array of numbers called **weights** (or **parameters**). The depth of this filter **must** be the same as the depth of our input. In our case it is 1 (in a color image this is 3). Now once this filter begins moving (or **convolving**) around the image, it is multiplying the values inside this filter with the original pixel value of our image (also called **element wise multiplications**). These are then summed up (in our case this is just one multiplication of 28x28) to an individual value, which is a representation of **just** the top left of our image. Now repeat this until every unique location has a number and we will get what is called an **activation** or **feature map**. This feature map will be 784 different locations, which turns into a 28x28 array
```
def conv(ni, nf): return nn.Conv2d(ni, nf, kernel_size=3, stride=2, padding=1)
```
Here we can see our `ni` is equivalent to the depth of the filter, and `nf` is equivalent to how many filters we will be using. (Fun fact this always has to be divisible by the size of our image).
### Batch Normalization
As we send our tensors through our model, it is important to normalize our data throughout the network. Doing so can allow for a much larger improvement in training speed, along with allowing each layer to learn independantly (as each layer is then re-normalized according to it's outputs)
```
def bn(nf): return nn.BatchNorm2d(nf)
```
`nf` will be the same as the filter output from our previous convolutional layer
### Activation functions
They give our models non-linearity and work with the `weights` we mentioned earlier along with a `bias` through a process called **back-propagation**. These allow our models to learn and perform more complex tasks because they can choose to fire or activate one of those neurons mentioned earlier. On a simple sense, let's look at the `ReLU` activation function. It operates by turning any negative values to zero, as visualized below:
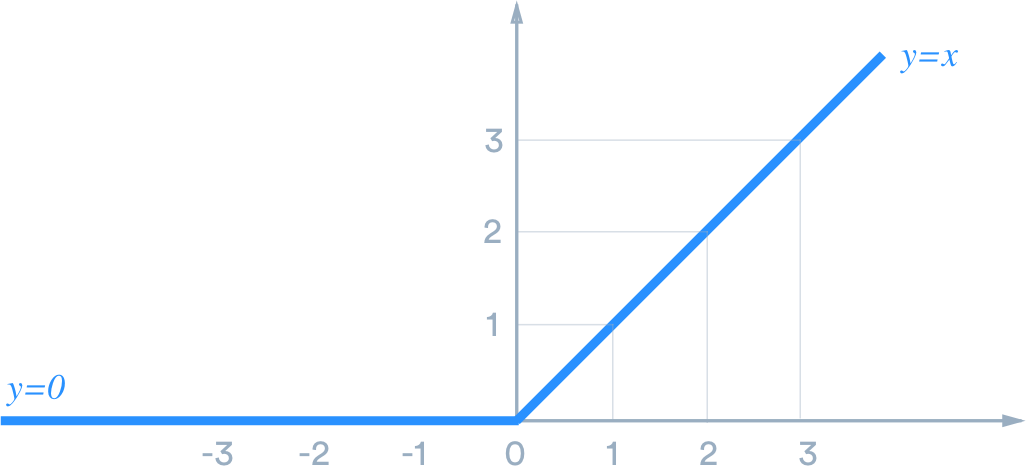
From "A Practical Guide to ReLU by Danqing Liu [URL](https://medium.com/@danqing/a-practical-guide-to-relu-b83ca804f1f7).
```
def ReLU(): return nn.ReLU(inplace=False)
```
### Flattening
The last bit we need to do is take all these activations and this outcoming matrix and flatten it into a single dimention of predictions. We do this with a `Flatten()` module
```
Flatten??
```
## Making a Model
* Five convolutional layers
* `nn.Sequential`
* 1 -> 32 -> 10
```
model = nn.Sequential(
conv(1, 8),
bn(8),
ReLU(),
conv(8, 16),
bn(16),
ReLU(),
conv(16,32),
bn(32),
ReLU(),
conv(32, 16),
bn(16),
ReLU(),
conv(16, 10),
bn(10),
Flatten()
)
```
Now let's make our `Learner`
```
learn = Learner(dls, model, loss_func=CrossEntropyLossFlat(), metrics=accuracy)
```
We can then also call `learn.summary` to take a look at all the sizes with thier **exact** output shapes
```
learn.summary()
```
`learn.summary` also tells us:
* Total parameters
* Trainable parameters
* Optimizer
* Loss function
* Applied `Callbacks`
```
learn.lr_find()
```
Let's use a learning rate around 1e-1 (0.1)
```
learn.fit_one_cycle(3, lr_max=1e-1)
```
## Simplify it
* Try to make it more like `ResNet`.
* `ConvLayer` contains a `Conv2d`, `BatchNorm2d`, and an activation function
```
def conv2(ni, nf): return ConvLayer(ni, nf, stride=2)
```
And make a new model
```
net = nn.Sequential(
conv2(1,8),
conv2(8,16),
conv2(16,32),
conv2(32,16),
conv2(16,10),
Flatten()
)
```
Great! That looks much better to read! Let's make sure we get (roughly) the same results with it.
```
learn = Learner(dls, net, loss_func=CrossEntropyLossFlat(), metrics=accuracy)
learn.fit_one_cycle(3, lr_max=1e-1)
```
Almost the exact same! Perfect! Now let's get a bit more advanced
## ResNet (kinda)
The ResNet architecture is built with what are known as ResBlocks. Each of these blocks consist of two `ConvLayers` that we made before, where the number of filters do not change. Let's generate these layers.
```
class ResBlock(Module):
def __init__(self, nf):
self.conv1 = ConvLayer(nf, nf)
self.conv2 = ConvLayer(nf, nf)
def forward(self, x): return x + self.conv2(self.conv1(x))
```
* Class notation
* `__init__`
* `foward`
Let's add these in between each of our `conv2` layers of that last model.
```
net = nn.Sequential(
conv2(1,8),
ResBlock(8),
conv2(8,16),
ResBlock(16),
conv2(16,32),
ResBlock(32),
conv2(32,16),
ResBlock(16),
conv2(16,10),
Flatten()
)
net
```
Awesome! We're building a pretty substantial model here. Let's try to make it **even simpler**. We know we call a convolutional layer before each `ResBlock` and they all have the same filters, so let's make that layer!
```
def conv_and_res(ni, nf): return nn.Sequential(conv2(ni, nf), ResBlock(nf))
net = nn.Sequential(
conv_and_res(1,8),
conv_and_res(8,16),
conv_and_res(16,32),
conv_and_res(32,16),
conv2(16,10),
Flatten()
)
```
And now we have something that resembles a ResNet! Let's see how it performs
```
learn = Learner(dls, net, loss_func=CrossEntropyLossFlat(), metrics=accuracy)
learn.lr_find()
```
Let's do 1e-1 again
```
learn.fit_one_cycle(3, lr_max=1e-1)
```
| github_jupyter |
[Table of Contents](./table_of_contents.ipynb)
# Least Squares Filters
```
#format the book
%matplotlib inline
from __future__ import division, print_function
from book_format import load_style
load_style()
```
## Introduction
**author's note**: This was snipped from the g-h chapter, where it didn't belong. This chapter is not meant to be read yet! I haven't written it yet.
Near the beginning of the chapter I used `numpy.polyfit()` to fit a straight line to the weight measurements. It fits a n-th degree polynomial to the data using a 'least squared fit'. How does this differ from the g-h filter?
Well, it depends. We will eventually learn that the Kalman filter is optimal from a least squared fit perspective under certain conditions. However, `polyfit()` fits a polynomial to the data, not an arbitrary curve, by minimizing the value of this formula:
$$E = \sum_{j=0}^k |p(x_j) - y_j|^2$$
I assumed that my weight gain was constant at 1 lb/day, and so when I tried to fit a polynomial of $n=1$, which is a line, the result very closely matched the actual weight gain. But, of course, no one consistently only gains or loses weight. We fluctuate. Using 'polyfit()' for a longer series of data would yield poor results. In contrast, the g-h filter reacts to changes in the rate - the $h$ term controls how quickly the filter reacts to these changes. If we gain weight, hold steady for awhile, then lose weight, the filter will track that change automatically. 'polyfit()' would not be able to do that unless the gain and loss could be well represented by a polynomial.
Another advantage of this form of filter, even if the data fits a *n*-degree polynomial, is that it is *recursive*. That is, we can compute the estimate for this time period knowing nothing more than the estimate and rate from the last time period. In contrast, if you dig into the implementation for `polyfit()` you will see that it needs all of the data before it can produce an answer. Therefore algorithms like `polyfit()` are not well suited for real-time data filtering. In the 60's when the Kalman filter was developed computers were very slow and had extremely limited memory. They were utterly unable to store, for example, thousands of readings from an aircraft's inertial navigation system, nor could they process all of that data in the short period of time needed to provide accurate and up-to-date navigation information.
Up until the mid 20th century various forms of Least Squares Estimation was used for this type of filtering. For example, for NASA's Apollo program had a ground network for tracking the Command and Service Model (CSM) and the Lunar Module (LM). They took measurements over many minutes, batched the data together, and slowly computed an answer. In 1960 Stanley Schmidt at NASA Ames recognized the utility of Rudolf Kalman's seminal paper and invited him to Ames. Schmidt applied Kalman's work to the on board navigation systems on the CSM and LM, and called it the "Kalman filter".[1] Soon after, the world moved to this faster, recursive filter.
The Kalman filter only needs to store the last estimate and a few related parameters, and requires only a relatively small number of computations to generate the next estimate. Today we have so much memory and processing power that this advantage is somewhat less important, but at the time the Kalman filter was a major breakthrough not just because of the mathematical properties, but because it could (barely) run on the hardware of the day.
This subject is much deeper than this short discussion suggests. We will consider these topics many more times throughout the book.
| github_jupyter |
```
import numpy as np
import pandas as pd
import linearsolve as ls
import matplotlib.pyplot as plt
plt.style.use('classic')
%matplotlib inline
```
# Class 15: Prescott's Real Business Cycle Model II
In this notebook, we continue to examine the centralized version of the model from pages 11-17 in Edward Prescott's article "Theory Ahead of Business Cycle Measurement in the Fall 1986 of the Federal Reserve Bank of Minneapolis' *Quarterly Review* (link to article: [https://www.minneapolisfed.org/research/qr/qr1042.pdf](https://www.minneapolisfed.org/research/qr/qr1042.pdf)). In this notebook, we:
1. Look at the effect of changing $\rho$ (the autoregressive coefficient on log TFP) on the simulated impulse responses of model variables to a TFP shock
2. Compute a stochastic simulation of the model and compute summary statistics.
## Example: Effect of Changing $\rho$ on Impulse Responses
Recall that the equilibrium conditions for Prescott's RBC model are:
\begin{align}
\frac{1}{C_t} & = \beta E_t \left[\frac{\alpha A_{t+1}K_{t+1}^{\alpha-1}L_{t+1}^{1-\alpha} +1-\delta }{C_{t+1}}\right]\\
\frac{\varphi}{1-L_t} & = \frac{(1-\alpha)A_tK_t^{\alpha}L_t^{-\alpha}}{C_t} \\
Y_t & = A_t K_t^{\alpha}L_t^{1-\alpha}\\
K_{t+1} & = I_t + (1-\delta) K_t\\
Y_t & = C_t + I_t\\
\log A_{t+1} & = \rho \log A_t + \epsilon_{t+1}
\end{align}
where $\epsilon_{t+1} \sim \mathcal{N}(0,\sigma^2)$.
The objective is use `linearsolve` to simulate impulse responses to a TFP shock using the following parameter values for the simulation for $\rho = 0.5,0.75,0.9,0.99$. Other parameter values are given in the table below:
| $\sigma$ | $\beta$ | $\varphi$ | $\alpha$ | $\delta $ |
|----------|---------|-----------|----------|-----------|
| 0.006 | 0.99 | 1.7317 | 0.35 | 0.025 |
## Model Preparation
As usual, we recast the model in the form required for `linearsolve`. Write the model with all variables moved to the left-hand side of the equations and dropping the expecations operator $E_t$ and the exogenous shock $\epsilon_{t+1}$:
\begin{align}
0 & = \beta\left[\frac{\alpha A_{t+1}K_{t+1}^{\alpha-1}L_{t+1}^{1-\alpha} +1-\delta }{C_{t+1}}\right] - \frac{1}{C_t}\\
0 & = \frac{(1-\alpha)A_tK_t^{\alpha}L_t^{-\alpha}}{C_t} - \frac{\varphi}{1-L_t}\\
0 & = A_t K_t^{\alpha}L_t^{1-\alpha} - Y_t\\
0 & = I_t + (1-\delta) K_t - K_{t+1}\\
0 & = C_t + I_t - Y_t\\
0 & = \rho \log A_t - \log A_{t+1}
\end{align}
Remember, capital and TFP are called *state variables* because they're $t+1$ values are predetermined. Output, consumption, and investment are called a *costate* or *control* variables. Note that the model as 5 equations in 5 endogenous variables.
## Initialization, Approximation, and Solution
The next several cells initialize the model in `linearsolve` and then approximate and solve it.
```
# Create a variable called 'parameters' that stores the model parameter values in a Pandas Series. CELL PROVIDED
# Note that a value for rho is absent
parameters = pd.Series()
parameters['sigma_squared'] = 0.006**2
parameters['beta'] = 0.99
parameters['phi'] = 1.7317
parameters['alpha'] = 0.35
parameters['delta'] = 0.025
# Print the model's parameters
print(parameters)
# Create variable called 'varNames' that stores the variable names in a list with state variables ordered first. CELL PROVIDED
varNames = ['a','k','y','c','i','l']
# Create variable called 'shockNames' that stores an exogenous shock name for each state variable. CELL PROVIDED
shockNames = ['e_a','e_k']
# Define a function that evaluates the equilibrium conditions of the model solved for zero. CELL PROVIDED
def equilibrium_equations(variables_forward,variables_current,parameters):
# Parameters. PROVIDED
p = parameters
# Current variables. PROVIDED
cur = variables_current
# Forward variables. PROVIDED
fwd = variables_forward
# Euler equation
euler_equation = p.beta*(p.alpha*fwd.a*fwd.k**(p.alpha-1)*fwd.l**(1-p.alpha)+1-p.delta)/fwd.c - 1/cur.c
# Labor-labor choise
labor_leisure = (1-p.alpha)*cur.a*cur.k**p.alpha*cur.l**(-p.alpha)/cur.c - p.phi/(1-cur.l)
# Production function
production_function = cur.a*cur.k**p.alpha*cur.l**(1-p.alpha) - cur.y
# Capital evolution
capital_evolution = cur.i + (1 - p.delta)*cur.k - fwd.k
# Market clearing
market_clearing = cur.c+cur.i - cur.y
# Exogenous tfp
tfp_process = p.rho*np.log(cur.a) - np.log(fwd.a)
# Stack equilibrium conditions into a numpy array
return np.array([
euler_equation,
labor_leisure,
production_function,
capital_evolution,
market_clearing,
tfp_process
])
```
Next, initialize the model using `ls.model` which takes the following required arguments:
* `equations`
* `nstates`
* `varNames`
* `shockNames`
* `parameters`
```
# Initialize the model into a variable named 'rbc_model'. CELL PROVIDED.
rbc_model = ls.model(equations = equilibrium_equations,
nstates=2,
varNames=varNames,
shockNames=shockNames,
parameters=parameters)
```
### SImulation and Plotting
The objective is to create a $2\times 2$ grid of plots containing the impulse responses of TFP, output, labor, and consumption to a one percent shock to TFP for each of the values for $\rho$: 0.5,0.75,0.9,0.99. Here are the steps that we'll take:
1. Initilize figure and axes for plotting.
2. Iterate over each desired value for $\rho$.
1. Set `rbc_model.parameters['rho']` equal to current value of $\rho$.
2. Use `rbc_model.compute_ss()` to compute the steady state with `guess` equal to `[1,4,1,1,1,0.5]`.
3. Use `rbc_model.approximate_and_solve()` to approximate and solve the model with the current value of $\rho$.
4. Use `rbc_model.impulse()` to compute the 51 period impulse response to a 0.01 unit shock to TFP in period 5.
5. Add the computed impulse responses to the axes.
```
# Create a 12x8 figure
# Create four axis variables: 'ax1', 'ax2', 'ax3', 'ax4'
# Create an axis equal to the size of the figure. PROVIDED
ax0 = fig.add_subplot(1,1,1)
# Turn off the axis so that the underlying axes are visible. PROVIDED
ax0.set_frame_on(False)
# Hide the x-axis. PROVIDED
ax0.get_xaxis().set_visible(False)
# Hide the y-axis. PROVIDED
ax0.get_yaxis().set_visible(False)
# Create variable called 'rho_values' that stores the desired values of rho
# Iterate over the elements of rho_values
# Update the value of rho in rbc_model.parameters
# Compute the steady state with initial guess equal to [1,4,1,1,1,0.5]
# Approximate the model and solve
# Compute the impulse responses to a 0.01 unit shock to TFP
# Add plots of TFP, output, labor, and consumption to ax1, ax2, ax3, and ax4
# Plot the point 0,0 on ax0 with the same line properties used for the other plotted lines and provide a label
# Set axis titles to the axes
# Add grids to the axes
# Set ax1 y-axis limits to [0,2]
# Set ax2 y-axis limits to [0,2]
# Set ax3 y-axis limits to [-0.5,1.25]
# Set ax4 y-axis limits to [-0.5,1.5]
# Add legend below the figure. PROVIDED
legend = ax0.legend(loc='upper center',bbox_to_anchor=(0.5,-0.075), ncol=4,fontsize=15)
```
## Example: Stochastic Simulation
Compute a 401 period stochastic simulation of the model. Set $\rho=0.75$ to match US business cycle data. And set the seed for the simulation to 126.
```
# Set the value of rho in rbc_model.parameters to 0.75
# Compute the steady state with initial guess equal to [1,4,1,1,1,0.5]
# Approximate the model and solve
# Compute the stochastic simulation using the .stoch_sim() method of rbc_model
```
On a single axis, plot the simulated values for output, consumption, investment, and labor.
```
# Print the standard deviations (times 100) of output, consumption, investment, and labor in the simulated data
# Print the correlations of output, consumption, investment, and labor in the simulated data
```
| github_jupyter |
```
from bokeh.io import output_notebook, show
from bokeh.plotting import figure
from bokeh.models import Range1d, ColumnDataSource, HoverTool, CrosshairTool
from bokeh.layouts import gridplot
from bokeh.palettes import Category20
import pandas as pd
import numpy as np
import requests
import io
output_notebook()
import rasterio
from bokeh.models.mappers import LinearColorMapper
print('start')
ds = rasterio.open('/out/luh2/ssp1_rcp2.6_image-BIIAb-2100.tif')
data = ds.read(1, masked=True)[::-1]
print(data.shape)
p = figure(title='Test of image()', plot_width=ds.width>>1, plot_height=ds.height>>1, x_range=(0, 10), y_range=(0, 10))
cm = LinearColorMapper(palette='Viridis256', nan_color='black')
p.image(image=[data], x=0, y=0, dw=10, dh=10, color_mapper=cm)
show(p)
print('done')
scenarios = ('historical', 'ssp1_rcp2.6_image', 'ssp3_rcp7.0_aim', 'ssp4_rcp3.4_gcam', 'ssp4_rcp6.0_gcam', 'ssp5_rcp8.5_remind-magpie')
plots = []
for scenario in scenarios:
print(scenario)
row = []
for indicator in ('BIIAb', 'BIISR'):
title = 'historical'
base_url = "http://ipbes.s3.amazonaws.com/summary/%s-%s-%s- 900-2014.csv"
if scenario != 'historical':
ssp, rcp, model = scenario.upper().split('_')
title = '%s -- %s / %s' % (indicator, ssp, rcp)
base_url = "http://ipbes.s3.amazonaws.com/summary/%s-%s-%s-2015-2100.csv"
p = figure(title=title)
p.y_range = Range1d(0.45, 1)
print(base_url % (scenario, indicator, 'subreg'))
s = requests.get(base_url % (scenario, indicator, 'subreg')).content
df = pd.read_csv(io.StringIO(s.decode('utf-8')))
#df = pd.read_csv('ipbes-upload/%s-%s-subreg-2015-2100.csv' % (scenario, indicator))
subset = df.loc[:, '2015':'2100'].T
subset.columns = df['Name']
subset.reset_index(inplace=True)
subset = subset.rename(columns={'index': 'Year'})
mypalette=Category20[len(subset.columns)]
for idx, col in enumerate(subset.columns):
if col in ('Year', 'Excluded'):
continue
src = ColumnDataSource(data={
'year': subset.Year,
'data': subset[col],
'name': [col for n in range(len(subset))]
})
p.line('year', 'data', source=src, line_width=4, color=mypalette[idx])
base_url = "http://ipbes.s3.amazonaws.com/summary/%s-%s-%s-2015-2100.csv"
s = requests.get(base_url % (scenario, indicator, 'global')).content
df = pd.read_csv(io.StringIO(s.decode('utf-8')))
glob = df.loc[:, '2015':'2100'].T.reset_index()
src = ColumnDataSource(data={
'year': glob['index'],
'data': glob[0],
'name': ['Global' for n in range(len(glob))]
})
p.line('year', 'data', source=src, line_width=8, color='black')
p.add_tools(HoverTool(tooltips=[('Year', '@year'), (indicator, '@data'), ('Region', '@name')]))
row.append(p)
plots.append(row)
grid = gridplot(plots, sizing_mode='scale_width')
show(grid)
```
| github_jupyter |
**CogDL Notebook**
created by CogDL Team
[cogdlteam@gmail.com]
This notebook shows how to write your first GCN model.
CogDL Link: https://github.com/THUDM/CogDL
Colab Link: https://colab.research.google.com/drive/1V47IIanXxDxi0Qsd6feOvvyYuqXcFP6P?usp=sharing
```
import torch
import torch.nn as nn
import torch.nn.functional as F
```
**第一部分:手动模拟GCN的计算和训练过程。**
---
1. 根据初始的邻接矩阵A得到正则化后的邻接矩阵normA。
```
A = torch.tensor([[0, 1, 1, 1], [1, 0, 1, 0], [1, 1, 0, 1], [1, 0, 1, 0]])
A = A + torch.eye(4)
print("A=", A)
# 计算度数矩阵D,并对A进行正则化得到normA
D = torch.diag(A.sum(1))
D_hat = torch.diag(1.0 / torch.sqrt(A.sum(1)))
normA = torch.mm(torch.mm(D_hat, A), D_hat)
print("normA=", normA)
```
2. 根据初始特征X,模型参数W1,邻接矩阵normA来计算第一层的输出H1。
```
H0 = X = torch.FloatTensor([[1,0], [0,1], [1,0], [1,1]])
W1 = torch.tensor([[1, -0.5], [0.5, 1]], requires_grad=True)
# 通过normA/H0/W1计算得到H1
H1 = F.relu(torch.mm(normA, torch.mm(H0, W1)))
print(H1)
```
3. 计算第二层的输出H2和最后的输出Z。
```
W2 = torch.tensor([[0.5, -0.5], [1, 0.5]], requires_grad=True)
# 通过normA/H1/W2计算得到H2和Z
H2 = torch.mm(normA, torch.mm(H1, W2))
print("H2=", H2)
Z = F.softmax(H2, dim=-1)
print("Z=", Z)
```
4. 计算损失函数loss。
```
Y = torch.LongTensor([0, 1, 0, 0])
# 根据输出Z和标签Y来计算最后的loss
loss = F.nll_loss(Z.log(), Y)
print(loss.item())
```
5. 通过loss进行反向传播。可以看到模型参数W1/W2的梯度值。
```
loss.backward(retain_graph=True)
print(W1)
print(W1.grad)
print(W2)
print(W2.grad)
```
**第二部分:使用你实现的GCN模型来运行cora数据集**
---
1. 通过pip install来安装cogdl。
```
!pip install cogdl
```
2. 从cogdl中加载cora数据集(x表示特征,y表示标签,mask表示训练/验证/测试集的划分)
```
from cogdl.datasets import build_dataset_from_name
dataset = build_dataset_from_name("cora")
data = dataset[0]
print(data)
n = data.x.shape[0]
edge_index = torch.stack(data.edge_index)
A = torch.sparse_coo_tensor(edge_index, torch.ones(edge_index.shape[1]), (n, n)).to_dense()
```
3. 使用你实现的GCN模型进行训练(在GCN模型的forward中填入你在第一部分中写的代码)
```
import math
import copy
from tqdm import tqdm
def accuracy(y_pred, y_true):
y_true = y_true.squeeze().long()
preds = y_pred.max(1)[1].type_as(y_true)
correct = preds.eq(y_true).double()
correct = correct.sum().item()
return correct / len(y_true)
class GCN(nn.Module):
def __init__(
self,
in_feats,
hidden_size,
out_feats,
):
super(GCN, self).__init__()
self.out_feats = out_feats
self.W1 = nn.Parameter(torch.FloatTensor(in_feats, hidden_size))
self.W2 = nn.Parameter(torch.FloatTensor(hidden_size, out_feats))
self.reset_parameters()
def reset_parameters(self):
stdv = 1.0 / math.sqrt(self.out_feats)
torch.nn.init.uniform_(self.W1, -stdv, stdv)
torch.nn.init.uniform_(self.W2, -stdv, stdv)
def forward(self, A, X):
n = X.shape[0]
A = A + torch.eye(n, device=X.device)
# 依次计算normA/H1/H2,然后返回H2。注意:此处不需要计算Z,因为通常直接根据H2和Y来计算loss。
# 注意使用self.W1/W2来调用模型参数。
D_hat = torch.diag(1.0 / torch.sqrt(A.sum(1)))
normA = torch.mm(torch.mm(D_hat, A), D_hat)
H1 = F.relu(torch.mm(normA, torch.mm(X, self.W1)))
H2 = torch.mm(normA, torch.mm(H1, self.W2))
return H2
hidden_size = 64
model = GCN(data.x.shape[1], hidden_size, data.y.max() + 1)
if torch.cuda.is_available():
device = torch.device("cuda")
model = model.to(device)
A = A.to(device)
data.apply(lambda x: x.to(device))
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
epoch_iter = tqdm(range(100), position=0, leave=True)
best_model = None
best_loss = 1e8
for epoch in epoch_iter:
model.train()
optimizer.zero_grad()
logits = model(A, data.x)
loss = F.cross_entropy(logits[data.train_mask], data.y[data.train_mask])
loss.backward()
optimizer.step()
train_loss = loss.item()
model.eval()
with torch.no_grad():
logits = model(A, data.x)
val_loss = F.cross_entropy(logits[data.val_mask], data.y[data.val_mask]).item()
val_acc = accuracy(logits[data.val_mask], data.y[data.val_mask])
if val_loss < best_loss:
best_loss = val_loss
best_model = copy.deepcopy(model)
epoch_iter.set_description(f"Epoch: {epoch:03d}, Train Loss: {train_loss:.4f}, Val Loss: {val_loss:.4f}, Val Acc: {val_acc:.4f}")
with torch.no_grad():
logits = best_model(A, data.x)
val_acc = accuracy(logits[data.val_mask], data.y[data.val_mask])
test_acc = accuracy(logits[data.test_mask], data.y[data.test_mask])
print("Val Acc", val_acc)
print("Test Acc", test_acc)
```
4. 调用cogdl的GCN模型来运行cora数据集,观察两者的区别(包括Acc和训练时间)
```
from cogdl import experiment
experiment(dataset="cora", model="gcn", epochs=100)
```
| github_jupyter |
# Diffusion Maps
Author: Ketson R. M. dos Santos,
Date: June 3rd, 2020
This example shows how to use the UQpy DiffusionMaps class to
* reveal the embedded structure of noisy data;
Import the necessary libraries. Here we import standard libraries such as numpy and matplotlib, but also need to import the DiffusionMaps class from UQpy implemented in the DimensionReduction module.
```
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from UQpy.DimensionReduction import DiffusionMaps
```
Sample points randomly following a parametric curve and plot the 3D graphic.
```
a=6
b=1
k=10
u = np.linspace(0, 2*np.pi, 1000)
v = k*u
x0 = (a+b*np.cos(0.8*v))*(np.cos(u))
y0 = (a+b*np.cos(0.8*v))*(np.sin(u))
z0 = b*np.sin(0.8*v)
rox = 0.2
roy = 0.2
roz = 0.2
x = x0 + rox*np.random.normal(0,1,len(x0))
y = y0 + roy*np.random.normal(0,1,len(y0))
z = z0 + roz*np.random.normal(0,1,len(z0))
X = np.array([x, y, z]).transpose()
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.scatter(x, y, z, c='b', cmap=plt.cm.Spectral, s=8)
ax.plot(x0, y0, z0,'r',label='parametric curve')
plt.show()
```
Instantiate the class `DiffusionMaps` using `alpha=1`; `n_evecs=3`, because the first eigenvector is non-informative. Moreover, a Gaussian is used in the kernel construction.
```
dfm = DiffusionMaps(alpha=1, n_evecs=3, kernel_object=DiffusionMaps.gaussian_kernel)
```
Use the method `mapping` to compute the diffusion coordinates assuming `epsilon=0.3`.
```
diff_coords, evals, evecs = dfm.mapping(data=X, epsilon=0.3)
```
Plot the second and third diffusion coordinates to reveal the embedded structure of the data.
```
color = evecs[:,1]
plt.scatter(diff_coords[:,1], diff_coords[:,2], c=color, cmap=plt.cm.Spectral,s=8)
plt.axis('equal')
plt.show()
```
Use the colormap to observe how the embedded structure is distributed in the original set.
```
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.scatter(x, y, z, c=color, cmap=plt.cm.Spectral, s=8)
plt.show()
```
Now, use an user defined kernel capable to handle this kind of data.
```
from user_kernel_diffusion import my_kernel_diffusion
dfm = DiffusionMaps(alpha=1, n_evecs=3, kernel_object=my_kernel_diffusion)
```
Use the method `mapping` to compute the diffusion coordinates assuming `epsilon=0.3`.
```
diff_coord_new, evals, evecs = dfm.mapping(data=X)
color = evecs[:,1]
plt.scatter(diff_coords[:,1], diff_coords[:,2], c=color, cmap=plt.cm.Spectral,s=8)
plt.axis('equal')
plt.show()
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.scatter(x, y, z, c=color, cmap=plt.cm.Spectral, s=8)
plt.show()
```
| github_jupyter |
# Lesson 2.2:
# Using the PowerGrid Models API with SPARQL Queries
This tutorial focuses on how to make generic queries of the PowerGrid Models API to obtain names, mRIDs, measurements, and control setting objects modeled in CIM XML for common power system equipment used in GridAPPS-D.
The lesson reviews the format used for making generic SPARQL queries and then presents an extensive catalog of cut-and-paste code blocks for the most common queries.
__Learning Objectives:__
At the end of the tutorial, the user should be able to use the PowerGrid Models API to
*
*
*
## Getting Started
Before running any of the sample routines in this tutorial, it is first necessary to start the GridAPPS-D Platform and establish a connection to this notebook so that we can start passing calls to the API.
_Open the Ubuntu terminal and start the GridAPPS-D Platform if it is not running already:_
`cd gridappsd-docker`
~/gridappsd-docker$ `./run.sh -t develop`
_Once containers are running,_
gridappsd@[container]:/gridappsd$ `./run-gridappsd.sh`
```
# Establish connection to GridAPPS-D Platform:
from gridappsd import GridAPPSD
gapps = GridAPPSD("('localhost', 61613)", username='system', password='manager')
model_mrid = "_49AD8E07-3BF9-A4E2-CB8F-C3722F837B62" # IEEE 13 Node used for all example queries
```
---
# Table of Contents
* [1. Structure of Generic SPARQL Queries](#1.-Structure-of-Generic-SPARQL-Queries)
* [2. Making SPARQL Queries using the GridAPPSD-Python API](#2.-Making-SPARQL-Queries-using-the-GridAPPSD-Python-API)
* [3. Making SPARQL Queries using the STOMP Client](#3.-Making-SPARQL-Queries-using-the-STOMP-Client)
* [4. Making SPARQL Queries using the Blazegraph Workbench](#4.-Making-SPARQL-Queries-using-the-Blazegraph-Workbench)
* [5. Catalog of Common SPARQL Queries for CIM Objects](#5.-Catalog-of-Common-SPARQL-Queries-for-CIM-Objects)
# 1. Structure of Generic SPARQL Queries
The format of SPARQL queries was discussed in detail previously in [Lesson 1.7]().
# 2. Making SPARQL Queries using the GridAPPSD-Python API
---
# 3. Making SPARQL Queries using the Blazegraph Workbench
Open the [Blazegraph Workbench](http://localhost:8889/bigdata/#query) hosted on [localhost:8889/bigdata](http://localhost:8889/bigdata/#query)

In the query input window, it is possible to directly copy and paste queries to the Blazegraph database.
---
# 5. Catalog of Common SPARQL Queries for CIM Objects
The sections below present common power system equipment and measurement objects. The
## 5.1. Queries for all feeder models and core objects
### List all the feeders, with substations and regions
```
query = """
# list all the feeders, with substations and regions - DistFeeder
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?feeder ?fid ?station ?sid ?subregion ?sgrid ?region ?rgnid WHERE {
?s r:type c:Feeder.
?s c:IdentifiedObject.name ?feeder.
?s c:IdentifiedObject.mRID ?fid.
?s c:Feeder.NormalEnergizingSubstation ?sub.
?sub c:IdentifiedObject.name ?station.
?sub c:IdentifiedObject.mRID ?sid.
?sub c:Substation.Region ?sgr.
?sgr c:IdentifiedObject.name ?subregion.
?sgr c:IdentifiedObject.mRID ?sgrid.
?sgr c:SubGeographicalRegion.Region ?rgn.
?rgn c:IdentifiedObject.name ?region.
?rgn c:IdentifiedObject.mRID ?rgnid.
}
ORDER by ?station ?feeder
"""
# Preview API call output for the query
gapps.query_data(query)
```
## 2. Get all mRID values by class and name
```
query = """
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?type ?name ?id WHERE {
?s c:IdentifiedObject.name ?name.
?s c:IdentifiedObject.mRID ?id.
?s r:type ?rawtype.
bind(strafter(str(?rawtype),"#") as ?type)
}
ORDER by ?type ?name
"""
# Preview API call output
gapps.query_data(query)
```
# 1. Querying for Buses and Nodes
## 1.1. List the bus name and xy coordinates
Note: this query is the basis of qbus_template in InsertDER.py
```
query = """
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?bus ?seq ?locid ?x ?y WHERE { # ?cnid ?tid ?eqid
VALUES ?fdrid {"%s"} # inserts model_mrid
?fdr c:IdentifiedObject.mRID ?fdrid.
?cn c:ConnectivityNode.ConnectivityNodeContainer ?fdr.
?trm c:Terminal.ConnectivityNode ?cn.
?trm c:ACDCTerminal.sequenceNumber ?seq.
?trm c:Terminal.ConductingEquipment ?eq.
?eq c:PowerSystemResource.Location ?loc.
?trm c:IdentifiedObject.mRID ?tid.
?cn c:IdentifiedObject.mRID ?cnid.
?cn c:IdentifiedObject.name ?bus.
?eq c:IdentifiedObject.mRID ?eqid.
?loc c:IdentifiedObject.mRID ?locid.
?pt c:PositionPoint.Location ?loc.
# caution - these next three triples make the query very slow, uncomment only if needed
# ?pt c:PositionPoint.sequenceNumber ?seq.
# ?pt c:PositionPoint.xPosition ?x.
# ?pt c:PositionPoint.yPosition ?y
}
ORDER BY ?bus ?locid
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## List all the connectivity nodes by feeder
```
query = """
# list all the connectivity nodes by feeder
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?feeder ?name WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?fdr c:IdentifiedObject.mRID ?fdrid.
?s c:ConnectivityNode.ConnectivityNodeContainer ?fdr.
?s r:type c:ConnectivityNode.
?s c:IdentifiedObject.name ?name.
?fdr c:IdentifiedObject.name ?feeder.
}
ORDER by ?feeder ?name
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## List all the connectivity nodes by feeder, with voltage limits
```
query = '''
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?feeder ?bus ?cnid ?val ?dur ?dir WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?fdr c:IdentifiedObject.mRID ?fdrid.
?s c:ConnectivityNode.ConnectivityNodeContainer ?fdr.
?s r:type c:ConnectivityNode.
?s c:IdentifiedObject.name ?bus.
?s c:IdentifiedObject.mRID ?cnid.
?fdr c:IdentifiedObject.name ?feeder.
?s c:ConnectivityNode.OperationalLimitSet ?ols.
?vlim c:OperationalLimit.OperationalLimitSet ?ols.
?vlim r:type c:VoltageLimit.
?vlim c:OperationalLimit.OperationalLimitType ?olt.
?olt c:OperationalLimitType.acceptableDuration ?dur.
?olt c:OperationalLimitType.direction ?rawdir.
bind(strafter(str(?rawdir),"OperationalLimitDirectionKind.") as ?dir)
?vlim c:VoltageLimit.value ?val.
}
ORDER by ?feeder ?bus ?val
''' % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## 1.2. List all the connectivity node base voltages by feeder, for sensor service
```
query = """
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT DISTINCT ?feeder ?busname ?cnid ?nomv WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?fdr c:IdentifiedObject.mRID ?fdrid.
?bus c:ConnectivityNode.ConnectivityNodeContainer ?fdr.
?bus r:type c:ConnectivityNode.
?bus c:IdentifiedObject.name ?busname.
?bus c:IdentifiedObject.mRID ?cnid.
?fdr c:IdentifiedObject.name ?feeder.
?trm c:Terminal.ConnectivityNode ?bus.
?trm c:Terminal.ConductingEquipment ?ce.
?ce c:ConductingEquipment.BaseVoltage ?bv.
?bv c:BaseVoltage.nominalVoltage ?nomv.
}
ORDER by ?feeder ?busname ?nomv
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## List all the connectivity node base voltages by feeder, for visualization
```
query = """
# list all the connectivity node base voltages by feeder, for visualization
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT DISTINCT ?feeder ?busname ?nomv WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?fdr c:IdentifiedObject.mRID ?fdrid.
?bus c:ConnectivityNode.ConnectivityNodeContainer ?fdr.
?bus r:type c:ConnectivityNode.
?bus c:IdentifiedObject.name ?busname.
?fdr c:IdentifiedObject.name ?feeder.
?trm c:Terminal.ConnectivityNode ?bus.
?trm c:Terminal.ConductingEquipment ?ce.
?ce c:ConductingEquipment.BaseVoltage ?bv.
?bv c:BaseVoltage.nominalVoltage ?nomv.
}
ORDER by ?feeder ?busname ?nomv
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
# Querying for Switching Equipment
The sample queries below present the most commonly needed queries for power system model information related to switching objects.
The models currently stored in GridAPPS-D only use the _LoadBreakSwitch_ object for switching objects, so queries for other classes of switching objects are commented out in the sample code blocks below.
## Fuse, Breaker, Recloser, LoadBreakSwitch, Sectionaliser
This query features several different types of distribution switches available within CIM XML. The models currently stored in GridAPPS-D only use the _LoadBreakSwitch_ object for switching objects, so the other queries are commented out in this sample code block.
Note: the sectionalizer object will be supported in a future release.
```
query = """
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?name ?basev ?open ?continuous ?breaking ?fdrid (group_concat(distinct ?bus;separator="\n") as ?buses) (group_concat(distinct ?phs;separator="\n") as ?phases) WHERE {
# ?s r:type c:Sectionaliser.
# ?s r:type c:Disconnector.
# ?s r:type c:Fuse.
# ?s r:type c:Recloser.
# ?s r:type c:Breaker.
?s r:type c:LoadBreakSwitch.
?s c:Equipment.EquipmentContainer ?fdr.
?fdr c:IdentifiedObject.mRID ?fdrid.
?s c:IdentifiedObject.name ?name.
?s c:ConductingEquipment.BaseVoltage ?bv.
?bv c:BaseVoltage.nominalVoltage ?basev.
?s c:Switch.normalOpen ?open.
?s c:Switch.ratedCurrent ?continuous.
OPTIONAL {?s c:ProtectedSwitch.breakingCapacity ?breaking.}
?t c:Terminal.ConductingEquipment ?s.
?t c:Terminal.ConnectivityNode ?cn.
?cn c:IdentifiedObject.name ?bus
OPTIONAL {?swp c:SwitchPhase.Switch ?s.
?swp c:SwitchPhase.phaseSide1 ?phsraw.
bind(strafter(str(?phsraw),"SinglePhaseKind.") as ?phs) }
}
GROUP BY ?name ?basev ?open ?continuous ?breaking ?fdrid
ORDER BY ?name
"""
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
query = '''
# Storage - DistStorage
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?name ?bus ?ratedS ?ratedU ?ipu ?ratedE ?storedE ?state ?p ?q ?id ?fdrid (group_concat(distinct ?phs;separator="\\n") as ?phases) WHERE {
?s r:type c:BatteryUnit.
?s c:IdentifiedObject.name ?name.
?pec c:PowerElectronicsConnection.PowerElectronicsUnit ?s.
# feeder selection options - if all commented out, query matches all feeders
#VALUES ?fdrid {"_C1C3E687-6FFD-C753-582B-632A27E28507"} # 123 bus
#VALUES ?fdrid {"_49AD8E07-3BF9-A4E2-CB8F-C3722F837B62"} # 13 bus
#VALUES ?fdrid {"_5B816B93-7A5F-B64C-8460-47C17D6E4B0F"} # 13 bus assets
#VALUES ?fdrid {"_4F76A5F9-271D-9EB8-5E31-AA362D86F2C3"} # 8500 node
#VALUES ?fdrid {"_67AB291F-DCCD-31B7-B499-338206B9828F"} # J1
#VALUES ?fdrid {"_9CE150A8-8CC5-A0F9-B67E-BBD8C79D3095"} # R2 12.47 3
?pec c:Equipment.EquipmentContainer ?fdr.
?fdr c:IdentifiedObject.mRID ?fdrid.
?pec c:PowerElectronicsConnection.ratedS ?ratedS.
?pec c:PowerElectronicsConnection.ratedU ?ratedU.
?pec c:PowerElectronicsConnection.maxIFault ?ipu.
?s c:BatteryUnit.ratedE ?ratedE.
?s c:BatteryUnit.storedE ?storedE.
?s c:BatteryUnit.batteryState ?stateraw.
bind(strafter(str(?stateraw),"BatteryState.") as ?state)
?pec c:PowerElectronicsConnection.p ?p.
?pec c:PowerElectronicsConnection.q ?q.
OPTIONAL {?pecp c:PowerElectronicsConnectionPhase.PowerElectronicsConnection ?pec.
?pecp c:PowerElectronicsConnectionPhase.phase ?phsraw.
bind(strafter(str(?phsraw),"SinglePhaseKind.") as ?phs) }
bind(strafter(str(?s),"#_") as ?id).
?t c:Terminal.ConductingEquipment ?pec.
?t c:Terminal.ConnectivityNode ?cn.
?cn c:IdentifiedObject.name ?bus
}
GROUP by ?name ?bus ?ratedS ?ratedU ?ipu ?ratedE ?storedE ?state ?p ?q ?id ?fdrid
ORDER by ?name
'''
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## SynchronousMachine - DistSyncMachine
```
query = """
# SynchronousMachine - DistSyncMachine
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?name ?bus (group_concat(distinct ?phs;separator="\\n") as ?phases) ?ratedS ?ratedU ?p ?q ?id ?fdrid WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?s r:type c:SynchronousMachine.
?s c:IdentifiedObject.name ?name.
?s c:Equipment.EquipmentContainer ?fdr.
?fdr c:IdentifiedObject.mRID ?fdrid.
?s c:SynchronousMachine.ratedS ?ratedS.
?s c:SynchronousMachine.ratedU ?ratedU.
?s c:SynchronousMachine.p ?p.
?s c:SynchronousMachine.q ?q.
bind(strafter(str(?s),"#_") as ?id).
OPTIONAL {?smp c:SynchronousMachinePhase.SynchronousMachine ?s.
?smp c:SynchronousMachinePhase.phase ?phsraw.
bind(strafter(str(?phsraw),"SinglePhaseKind.") as ?phs) }
?t c:Terminal.ConductingEquipment ?s.
?t c:Terminal.ConnectivityNode ?cn.
?cn c:IdentifiedObject.name ?bus
}
GROUP by ?name ?bus ?ratedS ?ratedU ?p ?q ?id ?fdrid
ORDER by ?name
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
query = """
# Solar - DistSolar
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?name ?bus ?ratedS ?ratedU ?ipu ?p ?q ?fdrid (group_concat(distinct ?phs;separator="\\n") as ?phases) WHERE {
?s r:type c:PhotovoltaicUnit.
?s c:IdentifiedObject.name ?name.
?pec c:PowerElectronicsConnection.PowerElectronicsUnit ?s.
# feeder selection options - if all commented out, query matches all feeders
#VALUES ?fdrid {"_C1C3E687-6FFD-C753-582B-632A27E28507"} # 123 bus
#VALUES ?fdrid {"_49AD8E07-3BF9-A4E2-CB8F-C3722F837B62"} # 13 bus
#VALUES ?fdrid {"_5B816B93-7A5F-B64C-8460-47C17D6E4B0F"} # 13 bus assets
#VALUES ?fdrid {"_4F76A5F9-271D-9EB8-5E31-AA362D86F2C3"} # 8500 node
#VALUES ?fdrid {"_67AB291F-DCCD-31B7-B499-338206B9828F"} # J1
#VALUES ?fdrid {"_9CE150A8-8CC5-A0F9-B67E-BBD8C79D3095"} # R2 12.47 3
?pec c:Equipment.EquipmentContainer ?fdr.
?fdr c:IdentifiedObject.mRID ?fdrid.
?pec c:PowerElectronicsConnection.ratedS ?ratedS.
?pec c:PowerElectronicsConnection.ratedU ?ratedU.
?pec c:PowerElectronicsConnection.maxIFault ?ipu.
?pec c:PowerElectronicsConnection.p ?p.
?pec c:PowerElectronicsConnection.q ?q.
OPTIONAL {?pecp c:PowerElectronicsConnectionPhase.PowerElectronicsConnection ?pec.
?pecp c:PowerElectronicsConnectionPhase.phase ?phsraw.
bind(strafter(str(?phsraw),"SinglePhaseKind.") as ?phs) }
?t c:Terminal.ConductingEquipment ?pec.
?t c:Terminal.ConnectivityNode ?cn.
?cn c:IdentifiedObject.name ?bus
}
GROUP by ?name ?bus ?ratedS ?ratedU ?ipu ?p ?q ?fdrid
ORDER by ?name
"""
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
query = """
# list houses - DistHouse
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?fdrname ?name ?parent ?coolingSetpoint ?coolingSystem ?floorArea ?heatingSetpoint ?heatingSystem ?hvacPowerFactor ?numberOfStories ?thermalIntegrity ?id ?fdrid
WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?h r:type c:House.
?h c:IdentifiedObject.name ?name.
?h c:IdentifiedObject.mRID ?id.
?h c:House.floorArea ?floorArea.
?h c:House.numberOfStories ?numberOfStories.
OPTIONAL{?h c:House.coolingSetpoint ?coolingSetpoint.}
OPTIONAL{?h c:House.heatingSetpoint ?heatingSetpoint.}
OPTIONAL{?h c:House.hvacPowerFactor ?hvacPowerFactor.}
?h c:House.coolingSystem ?coolingSystemRaw.
bind(strafter(str(?coolingSystemRaw),"HouseCooling.") as ?coolingSystem)
?h c:House.heatingSystem ?heatingSystemRaw.
bind(strafter(str(?heatingSystemRaw),"HouseHeating.") as ?heatingSystem)
?h c:House.thermalIntegrity ?thermalIntegrityRaw.
bind(strafter(str(?thermalIntegrityRaw),"HouseThermalIntegrity.") as ?thermalIntegrity)
?h c:House.EnergyConsumer ?econ.
?econ c:IdentifiedObject.name ?parent.
?fdr c:IdentifiedObject.mRID ?fdrid.
?fdr c:IdentifiedObject.name ?fdrname.
?econ c:Equipment.EquipmentContainer ?fdr.
}
ORDER BY ?fdrname ?name
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
# Querying for measurements
## List all measurements, with buses and equipments
```
query = """
# list all measurements, with buses and equipments
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?class ?type ?name ?bus ?phases ?eqtype ?eqname ?eqid ?trmid ?id WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?eq c:Equipment.EquipmentContainer ?fdr.
?fdr c:IdentifiedObject.mRID ?fdrid.
{ ?s r:type c:Discrete. bind ("Discrete" as ?class)}
UNION
{ ?s r:type c:Analog. bind ("Analog" as ?class)}
?s c:IdentifiedObject.name ?name .
?s c:IdentifiedObject.mRID ?id .
?s c:Measurement.PowerSystemResource ?eq .
?s c:Measurement.Terminal ?trm .
?s c:Measurement.measurementType ?type .
?trm c:IdentifiedObject.mRID ?trmid.
?eq c:IdentifiedObject.mRID ?eqid.
?eq c:IdentifiedObject.name ?eqname.
?eq r:type ?typeraw.
bind(strafter(str(?typeraw),"#") as ?eqtype)
?trm c:Terminal.ConnectivityNode ?cn.
?cn c:IdentifiedObject.name ?bus.
?s c:Measurement.phases ?phsraw .
{bind(strafter(str(?phsraw),"PhaseCode.") as ?phases)}
}
ORDER BY ?class ?type ?name
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## List measurement points for PowerTransformer with no tanks
```
query = """
# list measurement points for PowerTransformer with no tanks
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
PREFIX xsd: <http://www.w3.org/2001/XMLSchema#>
SELECT ?name ?wnum ?bus ?eqid ?trmid WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?s c:Equipment.EquipmentContainer ?fdr.
?fdr c:IdentifiedObject.mRID ?fdrid.
?s r:type c:PowerTransformer.
?s c:IdentifiedObject.name ?name.
?s c:IdentifiedObject.mRID ?eqid.
?end c:PowerTransformerEnd.PowerTransformer ?s.
?end c:TransformerEnd.Terminal ?trm.
?end c:TransformerEnd.endNumber ?wnum.
?trm c:IdentifiedObject.mRID ?trmid.
?trm c:Terminal.ConnectivityNode ?cn.
?cn c:IdentifiedObject.name ?bus.
}
ORDER BY ?name ?wnum
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## List measurement points for Breakers, Reclosers, LoadBreakSwitches in a selected feeder
This query obtains
```
query = """
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?cimtype ?name ?bus1 ?bus2 ?id (group_concat(distinct ?phs;separator="") as ?phases) WHERE {
SELECT ?cimtype ?name ?bus1 ?bus2 ?phs ?id WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
VALUES ?cimraw {c:LoadBreakSwitch c:Recloser c:Breaker}
?fdr c:IdentifiedObject.mRID ?fdrid.
?s r:type ?cimraw.
bind(strafter(str(?cimraw),"#") as ?cimtype)
?s c:Equipment.EquipmentContainer ?fdr.
?s c:IdentifiedObject.name ?name.
?s c:IdentifiedObject.mRID ?id.
?t1 c:Terminal.ConductingEquipment ?s.
?t1 c:ACDCTerminal.sequenceNumber "1".
?t1 c:Terminal.ConnectivityNode ?cn1.
?cn1 c:IdentifiedObject.name ?bus1.
?t2 c:Terminal.ConductingEquipment ?s.
?t2 c:ACDCTerminal.sequenceNumber "2".
?t2 c:Terminal.ConnectivityNode ?cn2.
?cn2 c:IdentifiedObject.name ?bus2
OPTIONAL {?swp c:SwitchPhase.Switch ?s.
?swp c:SwitchPhase.phaseSide1 ?phsraw.
bind(strafter(str(?phsraw),"SinglePhaseKind.") as ?phs) }
} ORDER BY ?name ?phs
}
GROUP BY ?cimtype ?name ?bus1 ?bus2 ?id
ORDER BY ?cimtype ?name
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## List measurement points for PowerElectronicsConnection with BatteryUnit in a selected feeder
```
query = """
# list measurement points for PowerElectronicsConnection with BatteryUnit in a selected feeder
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?name ?uname ?bus ?id (group_concat(distinct ?phs;separator="") as ?phases) WHERE {
SELECT ?name ?uname ?bus ?phs ?id WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?fdr c:IdentifiedObject.mRID ?fdrid.
?s r:type c:PowerElectronicsConnection.
?s c:Equipment.EquipmentContainer ?fdr.
?s c:IdentifiedObject.name ?name.
?s c:IdentifiedObject.mRID ?id.
?peu r:type c:BatteryUnit.
?peu c:IdentifiedObject.name ?uname.
?s c:PowerElectronicsConnection.PowerElectronicsUnit ?peu.
?t1 c:Terminal.ConductingEquipment ?s.
?t1 c:ACDCTerminal.sequenceNumber "1".
?t1 c:Terminal.ConnectivityNode ?cn1.
?cn1 c:IdentifiedObject.name ?bus.
OPTIONAL {?pep c:PowerElectronicsConnectionPhase.PowerElectronicsConnection ?s.
?pep c:PowerElectronicsConnectionPhase.phase ?phsraw.
bind(strafter(str(?phsraw),"SinglePhaseKind.") as ?phs) }
} ORDER BY ?name ?phs
}
GROUP BY ?name ?uname ?bus ?id
ORDER BY ?name
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
## List measurement points for ACLineSegments in a selected feeder
```
query = """
# list measurement points for ACLineSegments in a selected feeder
PREFIX r: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX c: <http://iec.ch/TC57/CIM100#>
SELECT ?name ?bus1 ?bus2 ?id (group_concat(distinct ?phs;separator="") as ?phases) WHERE {
SELECT ?name ?bus1 ?bus2 ?phs ?id WHERE {
VALUES ?fdrid {"%s"} # inserts model_mrid
?fdr c:IdentifiedObject.mRID ?fdrid.
?s r:type c:ACLineSegment.
?s c:Equipment.EquipmentContainer ?fdr.
?s c:IdentifiedObject.name ?name.
?s c:IdentifiedObject.mRID ?id.
?t1 c:Terminal.ConductingEquipment ?s.
?t1 c:ACDCTerminal.sequenceNumber "1".
?t1 c:Terminal.ConnectivityNode ?cn1.
?cn1 c:IdentifiedObject.name ?bus1.
?t2 c:Terminal.ConductingEquipment ?s.
?t2 c:ACDCTerminal.sequenceNumber "2".
?t2 c:Terminal.ConnectivityNode ?cn2.
?cn2 c:IdentifiedObject.name ?bus2
OPTIONAL {?acp c:ACLineSegmentPhase.ACLineSegment ?s.
?acp c:ACLineSegmentPhase.phase ?phsraw.
bind(strafter(str(?phsraw),"SinglePhaseKind.") as ?phs) }
} ORDER BY ?name ?phs
}
GROUP BY ?name ?bus1 ?bus2 ?id
ORDER BY ?name
""" % model_mrid
# Preview API call output for the query on the IEEE 13 node model
gapps.query_data(query)
```
---
# 4. Making SPARQL Queries using the STOMP Client
Open the [GridAPPS-D Viz on localhost:8080](http://localhost:8080/) and log in.
Open the menu in the top left corner and select `Stomp Client`

Change the topic to the Powergrid Model API topic: `goss.gridappsd.process.request.data.powergridmodel`
Enter the SPARQL query wrapped as a JSON string, like this, and click `Send request`
```
{
"requestType": "QUERY",
"resultFormat": "JSON",
"queryString": "select ?feeder_name ?subregion_name ?region_name WHERE {?line r:type c:Feeder.?line c:IdentifiedObject.name ?feeder_name.?line c:Feeder.NormalEnergizingSubstation ?substation.?substation r:type c:Substation.?substation c:Substation.Region ?subregion.?subregion c:IdentifiedObject.name ?subregion_name .?subregion c:SubGeographicalRegion.Region ?region . ?region c:IdentifiedObject.name ?region_name}"
}
```

The PowerGrid Model API response will be displayed in the dialogue box below.
| github_jupyter |
# Evaluate the enrichment of Decipher shared phenotype patients on our ASD-CHD interactome
Update 19-12-12: found all damaging SNVs/indels in DECIPHER, by looking up karyotype for abnormal nervous and cardiovascular systems. Use this to make an ROC curve
Update 20-04-28: make sure results can be replicated from supplemental tables, add more description
Update 21-04-19: update for consistency with revision
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import networkx as nx
import pandas as pd
import random
# latex rendering of text in graphs
import matplotlib as mpl
from matplotlib import rcParams
rcParams['font.family'] = 'sans-serif'
rcParams['font.sans-serif'] = ['Arial']
sns.set_style('white')
import sys
% matplotlib inline
sns.set_style("ticks", {"xtick.major.size": 15, "ytick.major.size": 15})
plt.rcParams['svg.fonttype'] = 'none'
```
# Load the ASD-CHD network genes, (z_ASD-CHD>=3; 844 genes)
```
ASD_CHD_df = pd.read_excel('data/supplemental_tables_cell_systems_210416.xlsx',
sheet_name='Table S5',skiprows=1)
ASD_CHD_df.index=ASD_CHD_df['gene']
print(len(ASD_CHD_df))
display(ASD_CHD_df.head())
# number of dual-phenotype genes harboring at least one damaging variant (from DECIPHER v9.25, SNVs/INDELS/smallCNVs, + PCGC/PHN dual-phenotype dDNVs)
# but MLL2 is included here... set to nan because it is a duplicate of KMT2A in PCnet
sum(ASD_CHD_df['sum_dual_pheno_damaging_variants (DECIPHER 11.1, PCGC/PHN)']>0)
```
# Load the DECIPHER all dual-condition results
DECIPHER all dual-condition results come from DECIPHER v11.1 (manually aqcuired from the webtool), and include only SNVs/indels which were classified as likely LOF or protein changing. We were not able to collect comprehensive CNV information from the DECIPHER webtool.
```
# dec_kary_shared = pd.read_excel('data/DECIPHER_karyotypes_9.31.xlsx',sheet_name='ASD-CHD')
# dec_kary_shared = pd.read_excel('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/tables_19_01/DECIPHER_karyotypes_210129.xlsx',
# sheet_name = 'ASD-CHD_210129')
# dec_kary_shared = pd.read_excel('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/CELL_SYSTEMS/supplemental_tables_cell_systems_210212.xlsx',
# sheet_name='DECIPHER_11.1_DETAILED')
dec_kary_shared = pd.read_excel('data/supplemental_tables_cell_systems_210416.xlsx',
sheet_name='Table S6',skiprows=1)
dec_kary_shared.index=dec_kary_shared['gene']
dec_kary_shared = dec_kary_shared.fillna(0)
dec_kary_shared['num_LOF_PC']=dec_kary_shared['num_likely_LOF']+dec_kary_shared['num_PC']
print(dec_kary_shared['num_LOF_PC'].sum())
print(len(dec_kary_shared))
dec_kary_shared.head()
print(len(dec_kary_shared))
print(sum(dec_kary_shared['num_LOF_PC']))
print(sum(dec_kary_shared['num_LOF_PC']>1))
# bar-plot genes harboring most variants
# sns.set_style('whitegrid')
sns.set_style("ticks", {"xtick.major.size": 15, "ytick.major.size": 15})
# dual_plot =ASD_CHD_df[ASD_CHD_df['sum_damaging_variants (DECIPHER 9.25, PCGC/PHN)']>3]['sum_damaging_variants (DECIPHER 9.25, PCGC/PHN)']
dual_plot =ASD_CHD_df[ASD_CHD_df['sum_damaging_variants (DECIPHER 11.1, PCGC/PHN)']>3]['sum_damaging_variants (DECIPHER 11.1, PCGC/PHN)']
dual_plot=dual_plot.sort_values(ascending=False)
plt.figure(figsize=(4,1.41))
plt.bar(np.arange(len(dual_plot)),dual_plot,color='black')
tmp=plt.xticks(np.arange(len(dual_plot)),dual_plot.index.tolist(),fontsize=6,rotation='vertical',va='top')
plt.xlim([-1,len(dual_plot)])
plt.yticks([0,5,10,15],fontsize=9)
plt.ylabel('# dual-phenotype variants',fontsize=8)
# plt.savefig('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/figures_revisions_2102/Figure3/barchart_dualpheno_g2_11.1.png',dpi=300,bbox_inches='tight')
# plt.savefig('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/figures_revisions_2102/Figure3/barchart_dualpheno_g2_11.1.svg',dpi=300,bbox_inches='tight')
# plt.savefig('../../manuscript/figures_1911/Figure3/Figure3_final assets/barchart_dualpheno_g2.png',dpi=300,bbox_inches='tight')
# plt.savefig('../../manuscript/figures_1911/Figure3/Figure3_final assets/barchart_dualpheno_g2.svg',dpi=300,bbox_inches='tight',)
# plt.savefig('../../manuscript/figures_1911/Figure3/barchart_dualpheno_g2.svg',dpi=300,bbox_inches='tight')
dual_plot
```
# DECIPHER ROC curves
```
# load the full z-score results for ASD and CHD
num_reps = 5000
focal_interactome='PCnet'
rand_method = 'degree_binning' # can be 'degree_binning' or 'degree_ks_test' (deprecated)
z_ASD = pd.read_csv('data/z_score_results/z_ASD_'+str(num_reps)+'_reps'+focal_interactome+'_'+rand_method+'.tsv',
sep='\t',names=['gene','zscore'],index_col='gene')
z_CHD = pd.read_csv('data/z_score_results/z_CHD_'+str(num_reps)+'_reps'+focal_interactome+'_'+rand_method+'.tsv',
sep='\t',names=['gene','zscore'],index_col='gene')
# z_ASD = z_ASD['zscore'].copy()
# z_CHD = z_CHD['zscore'].copy()
z_ASD.head()
# multiply by the sign (+1 if both +, -1 if either -).
z_ASDCHD = z_ASD['zscore'].abs()*z_CHD['zscore'].abs()*(((np.sign(z_ASD['zscore'])>0)&(np.sign(z_CHD['zscore'])>0))*1.0-.5)*2
z_ASDCHD.head()
seed_genes = ASD_CHD_df[(ASD_CHD_df['ASD_seed']==1)|(ASD_CHD_df['CHD_seed']==1)].index.tolist()
print(len(seed_genes))
non_seed_genes = list(np.setdiff1d(z_ASDCHD.index.tolist(),seed_genes))
z_ASDCHD_noseeds = z_ASDCHD.loc[non_seed_genes]
print(len(z_ASDCHD_noseeds))
z_ASDCHD_noseeds.max()
z_comb = z_ASD.join(z_CHD,lsuffix='_ASD',rsuffix='_CHD')
z_comb['zprod_both']=z_ASDCHD
z_comb.head()
z_comb_no_seeds = z_comb.loc[non_seed_genes]
z_comb_no_seeds.head()
focal_genes_temp = list(np.intersect1d(z_ASDCHD[z_ASDCHD>3].index.tolist(),dec_kary_shared.index.tolist()))
print(len(focal_genes_temp))
print(dec_kary_shared['num_LOF_PC'].loc[focal_genes_temp].dropna().sum())
def calc_FPR_TPR(z_ASDCHD, both_DECIPHER,thresh_list):
TP_both_list,FP_both_list,TN_both_list,FN_both_list = [],[],[],[]
# thresh_list = np.arange(0,30)
# thresh_list = np.linspace(-17,400,1000)
# thresh_list = np.linspace(0,2,1000)
for thresh_temp in thresh_list:
both_predicted = z_ASDCHD[z_ASDCHD>=thresh_temp].index.tolist()
# ------ both -------
num_TP_both = len(np.intersect1d(both_DECIPHER,both_predicted))
num_FP_both = len(both_predicted)-num_TP_both
num_FN_both = len(both_DECIPHER)-num_TP_both
# number true negatives is everything that wasn't TP, FN, or FP
num_TN_both = len(z_ASDCHD)-num_TP_both-num_FP_both-num_FN_both
TP_both_list.append(num_TP_both)
FP_both_list.append(num_FP_both)
TN_both_list.append(num_TN_both)
FN_both_list.append(num_FN_both)
# P is number of true positives in the data
P_both = len(both_DECIPHER)
# N is number of true negatives in the data (# number of interactome genes- P..?)
N_both = len(z_ASDCHD)-P_both
# (TPR = sensitivity/ recall)
TPR_both = np.divide(TP_both_list,float(P_both))
FPR_both = np.divide(FP_both_list,float(N_both))
return(TPR_both,FPR_both)
# make an ROC curve sweeping z score
from sklearn.metrics import roc_curve,auc
thresh_list = np.linspace(-17,400,2000)
# -------- set threshold for number of variants observed per gene (1 used in main text, 0 used in supplement ) --------
dec_thresh=1
# ------------------------------------------------
both_DECIPHER = dec_kary_shared[dec_kary_shared['num_LOF_PC']>dec_thresh].index.tolist()
print(len(both_DECIPHER))
# make sure the genes are able to be recovered
both_DECIPHER = list(np.intersect1d(z_ASDCHD.index.tolist(),both_DECIPHER))
print(len(both_DECIPHER))
TPR_both,FPR_both=calc_FPR_TPR(z_comb['zprod_both'], both_DECIPHER,thresh_list)
both_DECIPHER_noseeds = list(np.setdiff1d(both_DECIPHER,seed_genes))
print(len(both_DECIPHER_noseeds))
TPR_both_noseeds,FPR_both_noseeds=calc_FPR_TPR(z_comb_no_seeds['zprod_both'], both_DECIPHER_noseeds,thresh_list)
# TPR_both
auc_both = auc(FPR_both,TPR_both)
print('\nAUC all:')
print(auc_both)
auc_both_noseeds = auc(FPR_both_noseeds,TPR_both_noseeds)
print('\nAUC seeds excluded:')
print(auc_both_noseeds)
auc_both_rand=[]
# randomize xeno_df
for r in np.arange(100):
# print(r)
if (r%10)==0:
print(r)
z_ASDrand = z_ASD.copy(deep=True)
rand_index=z_ASDrand.index.tolist()
np.random.shuffle(rand_index)
z_ASDrand.index=rand_index
z_CHDrand = z_CHD.copy(deep=True)
rand_index=z_CHDrand.index.tolist()
np.random.shuffle(rand_index)
z_CHDrand.index=rand_index
z_ASDrand.head()
z_ASDCHDrand = z_ASDrand['zscore'].abs()*z_CHDrand['zscore'].abs()*(((np.sign(z_ASDrand['zscore'])>0)&(np.sign(z_CHDrand['zscore'])>0))*1.0-.5)*2
TPR_both_rand,FPR_both_rand=calc_FPR_TPR(z_ASDCHDrand, both_DECIPHER,thresh_list)
auc_both_rand.append(auc(TPR_both_rand,FPR_both_rand))
print(np.mean(auc_both_rand))
from scipy.stats import norm
sns.set_style('white')
sns.set_style("ticks", {"xtick.major.size": 15, "ytick.major.size": 15})
# from scipy.special import ndtr
plt.figure(figsize=(1.8,1.8))
sns.distplot(auc_both_rand,label='permuted labels',color='k')
z_both = (auc_both-np.mean(auc_both_rand))/np.std(auc_both_rand)
print(z_both)
print(norm.sf(z_both))
z_both_noseeds = (auc_both_noseeds-np.mean(auc_both_rand))/np.std(auc_both_rand)
print(z_both_noseeds)
print(norm.sf(z_both_noseeds))
plt.vlines(auc_both,0,4,color='#A81012',label='z_ASD-CHD')
plt.vlines(auc_both_noseeds,0,4,color='#FF4F4F',label='z_ASD-CHD\nno seeds')
plt.legend(loc='upper right',fontsize=6)
plt.xlabel('AUC',fontsize=8)
plt.ylabel('count',fontsize=8)
# plt.savefig('../../manuscript/figures_1911/Supplement/DECIPHER_SNV_INDEL_AUC_'+str(dec_thresh)+'.png',dpi=300,bbox_inches='tight')
# plt.savefig('../../manuscript/figures_1911/Supplement/DECIPHER_SNV_INDEL_AUC_'+str(dec_thresh)+'.svg',dpi=300,bbox_inches='tight')
# plt.savefig('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/figures_revisions_2102/Figure3/DECIPHER_11.1_SNV_INDEL_AUC_'+str(dec_thresh)+'.png',dpi=300,bbox_inches='tight')
# plt.savefig('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/figures_revisions_2102/Figure3/DECIPHER_11.1_SNV_INDEL_AUC_'+str(dec_thresh)+'.svg',dpi=300,bbox_inches='tight')
from sklearn.metrics import roc_curve,auc
sns.set_style('white')
sns.set_style("ticks", {"xtick.major.size": 15, "ytick.major.size": 15})
threshlist = np.linspace(-17,400,2000)
ROC_df = pd.DataFrame({'thresh':threshlist,
'FPR_both':FPR_both,'TPR_both':TPR_both,
'FPR_both_noseeds':FPR_both_noseeds,'TPR_both_noseeds':TPR_both_noseeds,
'FPR_both_rand':FPR_both_rand,'TPR_both_rand':TPR_both_rand})
ROC_df.head()
plt.figure(figsize=(1.8,1.8))
plt.plot(FPR_both_noseeds,TPR_both_noseeds,'.-',color='#FF4F4F',label='$z_{ASD-CHD}$, no seeds',lw=.5,ms=2)
plt.plot(FPR_both,TPR_both,'.-',color='#A81012',label='$z_{ASD-CHD}$, all',lw=.5,ms=2)
# circle z=3
ROC_z3 = ROC_df[(ROC_df['thresh']>2.9)&(ROC_df['thresh']<3.1)]
display(ROC_z3)
plt.plot(ROC_z3['FPR_both'],ROC_z3['TPR_both'],'o',ms=4,color='k',fillstyle='none',label='$z=3$')
plt.plot(ROC_z3['FPR_both_noseeds'],ROC_z3['TPR_both_noseeds'],'o',ms=4,color='k',fillstyle='none',label='')
# plt.plot(FPR_both_rand,TPR_both_rand,'k.-')
plt.plot([0,1],[0,1],'k--',alpha=.5)
plt.legend(loc='upper left', fontsize=6,frameon=False)
plt.xlabel('DECIPHER false positive rate',fontsize=8)
plt.ylabel('DECIPHER true positive rate',fontsize=8)
plt.xticks(fontsize=8)
plt.yticks(fontsize=8)
plt.xlim([-.03,1.1])
plt.ylim([-.03,1.1])
# plt.savefig('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/figures_revisions_2102/Figure3/DECIPHER_11.1_SNV_INDEL_ROC_'+str(dec_thresh)+'.png',dpi=300,bbox_inches='tight')
# plt.savefig('/Users/brinrosenthal/Google Drive/UCSD/CCBB_tickets/Ideker_17_05_neuro/manuscript/figures_revisions_2102/Figure3/DECIPHER_11.1_SNV_INDEL_ROC_'+str(dec_thresh)+'.svg',dpi=300,bbox_inches='tight')
```
| github_jupyter |
# 1. Import libraries to be used in this notebook
```
import numpy as np # for array manipulation and basic scientific calculation
import xarray as xr # To read NetCDF files
import matplotlib.pyplot as plt # Core library for plotting
from functions.Calc_Emis import Calc_Emis_T # Emission calculation
from scipy.interpolate import griddata # Simple regridding
from functions.ZonalMeridional import SE_ZonalMeridional # To calculate zonal/meridional mean
```
# 2. Read files
## 2.1. Model output
```
Filebase_CONUS = '/glade/campaign/acom/acom-climate/tilmes/CO_CONUS/f.e22.FCcotagsNudged.ne0CONUSne30x8.cesm220.2012-01/atm/hist/f.e22.FCcotagsNudged.ne0CONUSne30x8.cesm220.2012-01.cam.h1.2013-MM.nc'
# Change "MM" to 01~12 for a year calculation
Files_CONUS = [] # empty list for a start
# file list with 12 files
for mm in np.arange(12):
Files_CONUS.append( Filebase_CONUS.replace('MM',str(mm+1).zfill(2)) )
# read 12 files at once using "mf"dataset
ds_CONUS = xr.open_mfdataset( Files_CONUS )
```
## 2.2. SCRIP file
### SCRIP file has grid information that is needed to calculate area of each grid
### Don't have to read, just locate where the files are
```
SCRIP_CONUS = '/glade/p/acom/MUSICA/grids/ne0CONUSne30x8/ne0CONUS_ne30x8_np4_SCRIP.nc'
```
# 3. Emission total
## 3.1. Calculate emissions for 12 months
```
# Calculate CO emissions - global
CO_Emis_global = Calc_Emis_T( ds_CONUS['SFCO'], print_results=False,
mw=28., scrip_file=SCRIP_CONUS )
# Calculate CO emissions - specify longitudes and latitudes
CO_Emis_CONUS = Calc_Emis_T( ds_CONUS['SFCO'], print_results=False,
mw=28., scrip_file=SCRIP_CONUS,
lon_range=[-130,-60], lat_range=[20,55] )
```
## 3.2 Draw a simple plot
```
fig = plt.figure( figsize=(8,5) )
ax = fig.add_subplot(1,1,1)
# Plot global monthly CO emissions
ax.plot( np.arange(12)+1, CO_Emis_global.emissions_total/1E9, 'ko-', lw=3, ms=10 )
# === Polish the plot ===
# Font size
ax.set_xticks( np.arange(12)+1 )
plt.setp( ax.get_xticklabels(), fontsize=15 )
plt.setp( ax.get_yticklabels(), fontsize=15 )
# Add x & y axis titles
ax.set_xlabel( 'Month', fontsize=15 )
ax.set_ylabel( 'CO emission [Gg/month]', fontsize=15 )
# Add a plot title
ax.set_title( 'Global CO emissions in 2013', fontsize=17, weight='semibold' )
```
# 4. Vertical profile
## 4.1. Calculate vertical profile of ozone over the region of interest
```
# Get the first month using the isel function of xarray
# You can use it for any dimensions such as altitude
ds_CONUS_Jan = ds_CONUS.isel(time=0)
# Colorado region
# Longitude in MUSICAv0 (CESM) is from 0 to 360
lon_range = [251, 258]; lat_range=[37,41]
# Get longitude & latitude
lon = ds_CONUS_Jan['lon'].values
lat = ds_CONUS_Jan['lat'].values
# Get array indices for Colorado, based on center longitude and latitude
Indices = np.where( ( lon >= lon_range[0] ) & ( lon <= lon_range[1] ) \
& ( lat >= lat_range[0] ) & ( lat <= lat_range[1] ) )[0]
# Calculate mean ozone profile
# The first dimension is altitude, the second is ncol
# Use np.mean to average it over the whole Colorado region
O3_profile = np.mean( ds_CONUS_Jan['O3'].values[:,Indices], 1 ) * 1e9
# O3S is ozone from stratosphere
O3S_profile = np.mean( ds_CONUS_Jan['O3S'].values[:,Indices], 1 ) * 1e9
# Also get geopotential height for y-axis
Z3_profile = np.mean( ds_CONUS_Jan['Z3'].values[:,Indices], 1 ) / 1e3
```
## 4.2. Plot vertical profile
```
fig = plt.figure( figsize=(8,5) )
ax = fig.add_subplot(1,1,1)
# Plot O3 profile
ax.plot( O3_profile[15:], Z3_profile[15:], 'ko-', lw=3, label='O3' )
ax.plot( O3S_profile[15:], Z3_profile[15:], 'bo-', lw=3, label='O3S' )
# === Polish the plot ===
# Font size
plt.setp( ax.get_xticklabels(), fontsize=15 )
plt.setp( ax.get_yticklabels(), fontsize=15 )
# Add x & y axis titles
ax.set_xlabel( 'O3 [ppbv]', fontsize=15 )
ax.set_ylabel( 'Geopotential Height [km]', fontsize=15 )
# Add a plot title
ax.set_title( 'O3 profile over Colorado', fontsize=17, weight='semibold' )
# Add a legend
ax.legend( fontsize=15, loc=0 )
```
# 5. Cross section
## 5.1. Regrid data to a traditional lat/lon grid
### This example uses griddata function by Scipy library.
### For advanced regridding methods, use ESMPy (https://earthsystemmodeling.org/esmpy_doc/release/latest/html/intro.html)
```
# This will define the lat/lon range we are using to correspond to global 1x1 grid
x = np.linspace(0.5,359.5,360)
y = np.linspace(-89.5,89.5,180)
# This will put lat and lon into arrays, something that is needed for plotting
X, Y = np.meshgrid(x,y)
# Here we are putting the unstructured data onto our defined 1x1 grid using linear interpolation
Regridded_PRECC = griddata( (lon,lat), ds_CONUS_Jan['PRECC'].values, (X, Y), method='linear')
Regridded_PRECL = griddata( (lon,lat), ds_CONUS_Jan['PRECL'].values, (X, Y), method='linear')
# Make zonal-mean
# use nanmean instead of mean to deal with nan values around the pole
# PRECL and PRECC are in m/s. 8.64e7 is used to convert m/s to mm/day
Zmean_PRECC = np.nanmean( Regridded_PRECC, 1 ) * 8.64e7
Zmean_PRECL = np.nanmean( Regridded_PRECL, 1 ) * 8.64e7
```
## 5.2. Plot meridional cross section
```
fig = plt.figure( figsize=(8,5) )
ax = fig.add_subplot(1,1,1)
# Plot global monthly CO emissions
ax.plot( y, Zmean_PRECC, 'k-', label='Convective' )
ax.plot( y, Zmean_PRECL, 'b-', label='Large-scale' )
# === Polish the plot ===
# Set x-axis ticks
ax.set_xticks( [-90,-60,-30,0,30,60,90] )
# Font size
plt.setp( ax.get_xticklabels(), fontsize=15 )
plt.setp( ax.get_yticklabels(), fontsize=15 )
# Add x & y axis titles
ax.set_xlabel( 'Latitude [degrees]', fontsize=15 )
ax.set_ylabel( 'Precipitation rate [mm/day]', fontsize=15 )
# Add a plot title
ax.set_title( 'Zonal-mean precipitation rate', fontsize=17 )
# Add a legend
ax.legend( fontsize=15, loc=0 )
```
## 5.3. Another way to calculate cross section
### This example uses a function that calculates area-weighted zonal/meridional cross section
```
# Regrid the data for zonal plot to a regular grid with the following resolution
Regridding_res=1. #in degree
# Get area information
area= ds_CONUS_Jan['area'].values
# PRECL and PRECC are in m/s. 8.64e7 is used to convert m/s to mm/day
PRECC = ds_CONUS_Jan['PRECC'].values* 8.64e7
PRECL = ds_CONUS_Jan['PRECL'].values* 8.64e7
Zmean_PRECC,Z_coord=SE_ZonalMeridional(PRECC,area,lat,Regridding_res)
Zmean_PRECL,Z_coord=SE_ZonalMeridional(PRECL,area,lat,Regridding_res)
```
## 5.4. Same as 5.2 but with different regridding method used in 5.3
```
fig = plt.figure( figsize=(8,5) )
ax = fig.add_subplot(1,1,1)
# Plot global monthly CO emissions
ax.plot( Z_coord, Zmean_PRECC, 'k-', label='Convective' )
ax.plot( Z_coord, Zmean_PRECL, 'b-', label='Large-scale' )
# === Polish the plot ===
# Set x-axis ticks
ax.set_xticks( [-90,-60,-30,0,30,60,90] )
# Font size
plt.setp( ax.get_xticklabels(), fontsize=15 )
plt.setp( ax.get_yticklabels(), fontsize=15 )
# Add x & y axis titles
ax.set_xlabel( 'Latitude [degrees]', fontsize=15 )
ax.set_ylabel( 'Precipitation rate [mm/day]', fontsize=15 )
# Add a plot title
ax.set_title( 'Zonal-mean precipitation rate', fontsize=17 )
# Add a legend
ax.legend( fontsize=15, loc=0 )
```
| github_jupyter |
## Configuration
_Initial steps to get the notebook ready to play nice with our repository. Do not delete this section._
Code formatting with [black](https://pypi.org/project/nb-black/).
```
%load_ext lab_black
import os
import pathlib
this_dir = pathlib.Path(os.path.abspath(""))
data_dir = this_dir / "data"
import pytz
import glob
import json
import requests
import pandas as pd
from datetime import datetime
```
## Download
Retrieve the page
```
url = "https://services3.arcgis.com/ibgDyuD2DLBge82s/arcgis/rest/services/Fresno_County_Zip_Data_Summary/FeatureServer/0/query?f=json&where=FZIP%3C%3E%2799999%27%20AND%20COUNT_Cases%3E%3D75%20AND%20FZIP%20IS%20NOT%20NULL&returnGeometry=false&spatialRel=esriSpatialRelIntersects&outFields=*&groupByFieldsForStatistics=FCITY&orderByFields=value%20desc&outStatistics=%5B%7B%22statisticType%22%3A%22sum%22%2C%22onStatisticField%22%3A%22COUNT_Cases%22%2C%22outStatisticFieldName%22%3A%22value%22%7D%5D&resultType=standard&cacheHint=true"
r = requests.get(url)
data = r.json()
```
## Parse
```
dict_list = []
for item in data["features"]:
d = dict(
county="Fresno",
area=item["attributes"]["FCITY"],
confirmed_cases=item["attributes"]["value"],
)
dict_list.append(d)
df = pd.DataFrame(dict_list)
df["area"] = df["area"].str.title()
```
Get timestamp
```
date_url = "https://services3.arcgis.com/ibgDyuD2DLBge82s/arcgis/rest/services/Fresno_County_Zip_Data_Summary/FeatureServer/0/?f=json"
date_r = requests.get(date_url)
date_data = date_r.json()
timestamp = date_data["editingInfo"]["lastEditDate"]
tz = pytz.timezone("America/Los_Angeles")
latest_date = datetime.fromtimestamp(timestamp / 1000, tz).date()
df["county_date"] = latest_date
```
## Trim
```
df["county"] = "Fresno"
df["area"] = df["area"].str.title()
c = df[["county", "area", "county_date", "confirmed_cases"]]
```
Remove county row
```
df = df[~(df.area == "Fresno")]
```
## Vet
```
try:
assert not len(df) > 27
except AssertionError:
raise AssertionError("Fresno County's place scraper has additional rows")
try:
assert not len(df) < 27
except AssertionError:
raise AssertionError("Fresno County's place scraper is missing rows")
```
## Export
Mark the current date
```
tz = pytz.timezone("America/Los_Angeles")
today = datetime.now(tz).date()
slug = "fresno"
df.to_csv(data_dir / slug / f"{today}.csv", index=False)
```
## Combine
```
csv_list = [
i
for i in glob.glob(str(data_dir / slug / "*.csv"))
if not str(i).endswith("timeseries.csv")
]
df_list = []
for csv in csv_list:
if "manual" in csv:
df = pd.read_csv(csv, parse_dates=["date"])
else:
file_date = csv.split("/")[-1].replace(".csv", "")
df = pd.read_csv(csv, parse_dates=["county_date"])
df["date"] = file_date
df_list.append(df)
df = pd.concat(df_list).sort_values(["date", "area"])
df.to_csv(data_dir / slug / "timeseries.csv", index=False)
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.optimize import fmin
import seaborn as sns
sns.set_context('talk')
sns.set_style('white')
RANDOM_SEED = 20090425
```
---
# Regression modeling
A general, primary goal of many statistical data analysis tasks is to relate the influence of one variable on another.
For example:
- how different medical interventions influence the incidence or duration of disease
- how baseball player's performance varies as a function of age.
- [how test scores are correlated with tissue LSD concentration](http://onlinelibrary.wiley.com/doi/10.1002/cpt196895635/abstract)
```
from io import StringIO
data_string = """
Drugs Score
0 1.17 78.93
1 2.97 58.20
2 3.26 67.47
3 4.69 37.47
4 5.83 45.65
5 6.00 32.92
6 6.41 29.97
"""
lsd_and_math = pd.read_table(StringIO(data_string), sep='\t', index_col=0)
lsd_and_math
```

> Taking LSD was a profound experience, one of the most important things in my life --Steve Jobs
```
lsd_and_math.plot(x='Drugs', y='Score', style='ro', legend=False, xlim=(0,8));
```
We can build a model to characterize the relationship between $X$ and $Y$, recognizing that additional factors other than $X$ (the ones we have measured or are interested in) may influence the response variable $Y$.
- $M(Y|X) = E(Y|X)$
- $M(Y|X) = Pr(Y=1|X)$
In general,
$$M(Y|X) = f(X)$$
for a regression model,
$$M(Y|X) = f(X\beta)$$
where $f$ is some function, for example a linear function:
<div style="font-size: 150%;">
$y_i = \beta_0 + \beta_1 x_{1i} + \ldots + \beta_k x_{ki} + \epsilon_i$
</div>
Regression is a **weighted sum** of independent predictors
and $\epsilon_i$ accounts for the difference between the observed response $y_i$ and its prediction from the model $\hat{y_i} = \beta_0 + \beta_1 x_i$. This is sometimes referred to as **process uncertainty**.
Interpretation: coefficients represent the change in Y for a unit increment of the predictor X.
Two important linear regression **assumptions**:
1. normal errors
2. homoscedasticity
## Parameter estimation
We would like to select $\beta_0, \beta_1$ so that the difference between the predictions and the observations is zero, but this is not usually possible. Instead, we choose a reasonable criterion: ***the smallest sum of the squared differences between $\hat{y}$ and $y$***.
<div style="font-size: 120%;">
$$R^2 = \sum_i (y_i - [\beta_0 + \beta_1 x_i])^2 = \sum_i \epsilon_i^2 $$
</div>
Squaring serves two purposes:
1. to prevent positive and negative values from cancelling each other out
2. to strongly penalize large deviations.
Whether or not the latter is a desired depends on the goals of the analysis.
In other words, we will select the parameters that minimize the squared error of the model.
```
sum_of_squares = lambda θ, x, y: np.sum((y - θ[0] - θ[1]*x) ** 2)
```
Here is a sample calculation, using aribitrary parameter values:
```
sum_of_squares([0,0.7], lsd_and_math.Drugs, lsd_and_math.Score)
```
However, we have the stated objective of minimizing the sum of squares, so we can pass this function to one of several optimizers in SciPy:
```
x, y = lsd_and_math.T.values
b0, b1 = fmin(sum_of_squares, [0,1], args=(x,y))
b0, b1
ax = lsd_and_math.plot(x='Drugs', y='Score', style='ro', legend=False, xlim=(0,8), ylim=(20, 90))
ax.plot([0,10], [b0, b0+b1*10])
for xi, yi in zip(x,y):
ax.plot([xi]*2, [yi, b0+b1*xi], 'k:')
```
## Alternative loss functions
Minimizing the sum of squares is not the only criterion we can use; it is just a very popular (and successful) one. For example, we can try to minimize the sum of absolute differences:
```
sum_of_absval = lambda θ, x, y: np.sum(np.abs(y - θ[0] - θ[1]*x))
b0, b1 = fmin(sum_of_absval, [0,0], args=(x,y))
print('\nintercept: {0:.2}, slope: {1:.2}'.format(b0,b1))
ax = lsd_and_math.plot(x='Drugs', y='Score', style='ro', legend=False, xlim=(0,8))
ax.plot([0,10], [b0, b0+b1*10])
```
### Exercise
1. Append an additional, extreme observation: `{'Drugs': 6, 'Score': 70}`.
2. Re-fit the model using both the sum of squares error and sum of absolute values error, and compare the resulting estimates.
```
# %load ../exercises/extreme_fit.py
```
## Polynomial regession
We are not restricted to a straight-line regression model; we can represent a curved relationship between our variables by introducing **polynomial** terms. For example, a cubic model:
<div style="font-size: 150%;">
$y_i = \beta_0 + \beta_1 x_i + \beta_2 x_i^2 + \epsilon_i$
</div>
```
sum_squares_quad = lambda θ, x, y: np.sum((y - θ[0] - θ[1]*x - θ[2]*(x**2)) ** 2)
b0,b1,b2 = fmin(sum_squares_quad, [1,1,-1], args=(x,y))
print('\nintercept: {0:.2}, x: {1:.2}, x2: {2:.2}'.format(b0,b1,b2))
ax = lsd_and_math.plot(x='Drugs', y='Score', style='ro', legend=False, xlim=(0,8))
xvals = np.linspace(0, 8, 100)
ax.plot(xvals, b0 + b1*xvals + b2*(xvals**2))
```
Although a polynomial model characterizes a nonlinear relationship, it is a linear problem in terms of estimation. That is, the regression model $f(y | x)$ is **linear in the parameters**.
For some data, it may be reasonable to consider polynomials of order>2. For example, consider the relationship between the number of spawning salmon and the number of juveniles recruited into the population the following year; one would expect the relationship to be positive, but not necessarily linear.
```
salmon = pd.read_table("../data/salmon.dat", delim_whitespace=True, index_col=0)
salmon.plot.scatter(x='spawners', y='recruits');
```
### Exercise
Create a sum of squares function for a cubic regression (order=3), and fit this function to the salmon spawning data.
```
# %load ../exercises/salmon_cubic.py
```
## Bayesian Linear Regression with PyMC3
In practice, we need not fit least squares models by hand because they are implemented generally in packages such as [`scikit-learn`](http://scikit-learn.org/) and [`statsmodels`](https://github.com/statsmodels/statsmodels/). Moreover, we are interested not only in obtaining a line of best fit, but also **estimates of uncertainty** in the line and the parameters used to calculate the line.
Here, we will return to a Bayesian approach and build a regression model in PyMC3.
### Priors
The first step in specifying our model is to specify priors for our model. Since regression parameters are continuous, and potentially positive or negative, we can use a Normal distribution with a variance set to an appropriate value that reflects our prior knowledsge of the parameter.
$$\beta_i \sim \text{Normal}(0, 100)$$
The other latent variable is the residual variance of the observations after applying our model. This is the **process uncertainty** that we identified earlier.
$$\sigma \sim \text{HalfCauchy}(1)$$
The **half-Cauchy** distribution used here provides support over positive continuous values, and has relatively large tail probabilities, allowing for the possibility of extreme values.
```
from pymc3 import HalfCauchy
ax = sns.kdeplot(HalfCauchy.dist(1).random(size=10000), gridsize=2000)
ax.set_xlim(0, 100);
from pymc3 import Normal, Model
with Model() as drugs_model:
intercept = Normal('intercept', 0, sd=100)
slope = Normal('slope', 0, sd=100)
σ = HalfCauchy('σ', 1)
```
### Likelihood
The sampling distribution of the data for a regression model is a normal distribution, and we specified the standard deviation for this sampling distribution in $\sigma$ above.
$$y_i \sim \text{Normal}(\mu_i, \sigma)$$
Here, $\mu_i$ is the expected value of the *i*th observation, which is generated by the regression model at the corresponding value of $x$. We can calculate this expected value as a function of the regression parameters and the data, and pass it to the normal likelihood:
```
with drugs_model:
μ = intercept + slope*x
score = Normal('score', μ, sd=σ, observed=y)
```
That's it!
The regression model is fully specified with these 5 lines of Python code. We can now use the fitting method of our choice to estimate a posterior distribution. In the previous module, we used variational inference, here we will use a **Markov chain Monte Carlo** algorithm, called **NUTS** (the No U-Turn Sampler).
```
from pymc3 import sample
with drugs_model:
drugs_sample = sample(1000, random_seed=RANDOM_SEED)
from pymc3 import plot_posterior
plot_posterior(drugs_sample[500:], varnames=['slope']);
```
Because we have a vector of samples from the estimated posterior, it is easy to calculate means, medians, standard deviations, and probability intervals. Above, we see the mean and the 95% **posterior credible interval** reported.
### Exercise
Fit a quadratic model for the salmon spawning data, using PyMC3.
```
# %load ../exercises/salmon_pymc3.py
```
Is this a good model? Why or why not?
## Checking model fit
One intuitive way of evaluating model fit is to compare model predictions with the observations used to fit the model. In other words, the fitted model can be used to **simulate
data**, and the distribution of the simulated data should resemble the distribution of the actual data.
Fortunately, simulating data from the model is a natural component of the Bayesian modelling framework. Recall, from our introduction to Bayesian inference, the **posterior predictive distribution**:
$$p(\tilde{y}|y) = \int p(\tilde{y}|\theta) f(\theta|y) d\theta$$
Here, $\tilde{y}$ represents some hypothetical new data that would be expected, taking into account the posterior uncertainty in the model parameters.
Sampling from the posterior predictive distribution is straighforward in PyMC; the `sample_ppc` function draws posterior predictive checks from all of the data likelihoods.
```
from pymc3 import sample_ppc
with drugs_model:
drugs_ppc = sample_ppc(drugs_sample, 1000)
```
This yields 1000 samples corresponding to each of the seven data points in our observation vector.
```
drugs_ppc['score'].shape
```
We can then compare these simulated data to the data we used to fit the model. Our claim is that the model might have plausibly been used to generate the data that we observed.
```
fig, axes = plt.subplots(4, 2)
axes_flat = axes.flatten()
for ax, real_data, sim_data in zip(axes_flat[:-1], y, drugs_ppc['score'].T):
ax.hist(sim_data, bins=20)
ax.vlines(real_data, *ax.get_ylim(), colors='red')
ax.set_yticklabels([])
sns.despine(left=True)
axes_flat[-1].axis('off')
plt.tight_layout()
```
## Generalized linear models
Often our data violates one or more of the linear regression assumptions:
- non-linear
- non-normal error distribution
- heteroskedasticity
this forces us to **generalize** the regression model in order to account for these characteristics.
As a motivating example, consider the Olympic medals data that we compiled earlier in the tutorial.

```
medals = pd.read_csv('../data/medals.csv')
medals.head()
```
We expect a positive relationship between population and awarded medals, but the data in their raw form are clearly not amenable to linear regression.
```
medals.plot(x='population', y='medals', kind='scatter')
```
Part of the issue is the scale of the variables. For example, countries' populations span several orders of magnitude. We can correct this by using the logarithm of population, which we have already calculated.
```
medals.plot(x='log_population', y='medals', kind='scatter')
```
This is an improvement, but the relationship is still not adequately modeled by least-squares regression.
```
from pymc3 import fit
x = medals.log_population
y = medals.medals
with Model() as medals_linear:
intercept = Normal('intercept', 0, sd=100)
slope = Normal('slope', 0, sd=100)
σ = HalfCauchy('σ', 1)
μ = intercept + slope*x
score = Normal('score', μ, sd=σ, observed=y)
samples = sample(500, random_seed=RANDOM_SEED)
yhat = samples['intercept'].mean() + samples['slope'].mean() * np.linspace(10, 22)
ax = medals.plot(x='log_population', y='medals', kind='scatter')
ax.plot(np.linspace(10, 22), yhat, color='red',
linewidth=2)
```
This is due to the fact that the response data are **counts**. As a result, they tend to have characteristic properties.
- discrete
- positive
- variance grows with mean
to account for this, we can do two things: (1) model the medal count on the **log scale** and (2) assume **Poisson errors**, rather than normal.
Recall the Poisson distribution from the previous section:
$$p(y)=\frac{e^{-\lambda}\lambda^y}{y!}$$
* $Y=\{0,1,2,\ldots\}$
* $\lambda > 0$
$$E(Y) = \text{Var}(Y) = \lambda$$
So, we will model the logarithm of the expected value as a linear function of our predictors:
$$\log(\lambda) = X\beta$$
In this context, the log function is called a **link function**. This transformation implies the mean of the Poisson is:
$$\lambda = \exp(X\beta)$$
We can plug this into the Poisson likelihood and use maximum likelihood to estimate the regression covariates $\beta$.
$$\log L = \sum_{i=1}^n -\exp(X_i\beta) + Y_i (X_i \beta)- \log(Y_i!)$$
As we have already done, we just need to code the kernel of this likelihood, and optimize!
```
# Poisson negative log-likelhood
poisson_loglike = lambda β, X, y: -(-np.exp(X.dot(β)) + y*X.dot(β)).sum()
```
Let's use the `assign` method to add a column of ones to the design matrix.
```
poisson_loglike([0,1], medals[['log_population']].assign(intercept=1), medals.medals)
```
We will use Nelder-Mead to minimize the negtive log-likelhood.
```
b1, b0 = fmin(poisson_loglike, [0,1], args=(medals[['log_population']].assign(intercept=1).values,
medals.medals.values))
b0, b1
```
The resulting fit looks reasonable.
```
ax = medals.plot(x='log_population', y='medals', kind='scatter')
xvals = np.arange(12, 22)
ax.plot(xvals, np.exp(b0 + b1*xvals), 'r--')
```
### Exercise: Poisson regression
Code the Poisson regression model above in PyMC3 (*Hint*: you will need to specify a Poisson distribution somewhere!).
```
from pymc3 import Poisson
from pymc3.math import exp
# %load ../exercises/medals.py
```
### Exercise: Multivariate GLM
Add the OECD indicator variable to the model, and estimate the model coefficients.
```
# %load ../exercises/medals_oecd.py
```
## Logistic Regression
Fitting a line to the relationship between two variables using the least squares approach is sensible when the variable we are trying to predict is continuous, but what about when the data are dichotomous?
- male/female
- pass/fail
- died/survived
Let's revisit the *very low birthweight infants* dataset from the last module.
```
vlbw = pd.read_csv('../data/vlbw.csv', index_col=0).dropna(axis=0, subset=['ivh', 'pneumo'])
ivh = vlbw.ivh.isin(['definite', 'possible']).astype(int).values
pneumo = vlbw.pneumo.values
```
Previously, we were comparing the number of **intra-ventricular hemorrhage** events between two groups, those with and without a pneumothorax, where we found a higher probability of IVH associated with pneumothorax. However, its possible that lower birthweight accounts for the difference, since birthweight is nominally lower in the pneumothorax group:
```
vlbw.groupby('pneumo').bwt.mean()
```
Let's consider the relative number of events as a function of birthweight, then.
```
bwt_kg = vlbw.bwt.values/1000
jitter = np.random.normal(scale=0.02, size=len(vlbw))
plt.scatter(bwt_kg, ivh + jitter, alpha=0.3)
plt.yticks([0,1])
plt.ylabel("IVH")
plt.xlabel("Birth weight")
```
I have added random jitter on the y-axis to help visualize the density of the points, and have plotted fare x-axis.
Clearly, fitting a line through this data makes little sense, for several reasons. First, for most values of the predictor variable, the line would predict values that are not zero or one. Second, it would seem odd to choose least squares (or similar) as a criterion for selecting the best line.
```
betas_vlbw = fmin(sum_of_squares, [1,1], args=(bwt_kg,ivh))
plt.scatter(bwt_kg, ivh + jitter, alpha=0.3)
plt.yticks([0,1])
plt.ylabel("IVH")
plt.xlabel("Birth weight")
plt.plot([0,2.5], [betas_vlbw[0] + betas_vlbw[1]*0, betas_vlbw[0] + betas_vlbw[1]*2.5], 'r-')
```
### Stochastic model
Rather than model the binary outcome explicitly, it makes sense instead to model the *probability* of death or survival in a **stochastic** model. Probabilities are measured on a continuous [0,1] scale, which may be more amenable for prediction using a regression line. We need to consider a different probability model for this exerciese however; let's consider the **Bernoulli** distribution as a generative model for our data:
<div style="font-size: 120%;">
$$f(y|p) = p^{y} (1-p)^{1-y}$$
</div>
where $y = \{0,1\}$ and $p \in [0,1]$. So, this model predicts whether $y$ is zero or one as a function of the probability $p$. Notice that when $y=1$, the $1-p$ term disappears, and when $y=0$, the $p$ term disappears.
So, the model we want to fit should look something like this:
<div style="font-size: 120%;">
$$p_i = \beta_0 + \beta_1 x_i + \epsilon_i$$
</div>
However, since $p$ is constrained to be between zero and one, it is easy to see where a linear (or polynomial) model might predict values outside of this range. As with the Poisson regression, we can modify this model slightly by using a link function to transform the probability to have an unbounded range on a new scale. Specifically, we can use a **logit transformation** as our link function:
<div style="font-size: 120%;">
$$\text{logit}(p) = \log\left[\frac{p}{1-p}\right] = x$$
</div>
Here's a plot of $p/(1-p)$
```
logit = lambda p: np.log(p/(1.-p))
unit_interval = np.linspace(0,1)
plt.plot(unit_interval/(1-unit_interval), unit_interval)
plt.xlabel(r'$p/(1-p)$')
plt.ylabel('p');
```
And here's the logit function:
```
plt.plot(logit(unit_interval), unit_interval)
plt.xlabel('logit(p)')
plt.ylabel('p');
```
The inverse of the logit transformation is:
<div style="font-size: 150%;">
$$p = \frac{1}{1 + \exp(-x)}$$
</div>
```
invlogit = lambda x: 1. / (1 + np.exp(-x))
```
So, now our model is:
<div style="font-size: 120%;">
$$\text{logit}(p_i) = \beta_0 + \beta_1 x_i + \epsilon_i$$
</div>
We can fit this model using maximum likelihood. Our likelihood, again based on the Bernoulli model is:
<div style="font-size: 120%;">
$$L(y|p) = \prod_{i=1}^n p_i^{y_i} (1-p_i)^{1-y_i}$$
</div>
which, on the log scale is:
<div style="font-size: 120%;">
$$l(y|p) = \sum_{i=1}^n y_i \log(p_i) + (1-y_i)\log(1-p_i)$$
</div>
We can easily implement this in Python, keeping in mind that `fmin` minimizes, rather than maximizes functions:
```
def logistic_like(θ, x, y):
p = invlogit(θ[0] + θ[1] * x)
# Return negative of log-likelihood
return -np.sum(y * np.log(p) + (1-y) * np.log(1 - p))
b0, b1 = fmin(logistic_like, [0.5,0], args=(bwt_kg, ivh))
b0, b1
jitter = np.random.normal(scale=0.01, size=len(bwt_kg))
plt.plot(bwt_kg, ivh+jitter, 'r.', alpha=0.3)
plt.yticks([0,.25,.5,.75,1])
xvals = np.linspace(0.4, 1.6)
plt.plot(xvals, invlogit(b0+b1*xvals))
```
The logistic regression is just another type of GLM, this time with a Bernoulli distribution representing the sampling distribution of the data.
## Bayesian multivariate logistic regression
Let's once again drop our hand-fit model into a Bayesian context. This time we will model two variables, one discrete (pneumothorax) and one continuous (birth weight). First, we will center birth weight to aid interpretation:
```
bwt_centered = bwt_kg - bwt_kg.mean()
```
Our regression model for predicting the latent probability of IVH for each individual will be:
$$p_i = \mu + \alpha * \text{bwt}_i + \beta * \text{pneumo}_i$$
here, $\mu$ is a baseline probability, $\alpha$ is a coefficient for centered birthweight, and $\beta$ is a coefficient for pneumothorax.
We then step through the procedure for coding a Bayesian model, first by specifing priors for the linear model coefficients:
```
with Model() as ivh_glm:
μ = Normal('μ', 0, sd=10)
α = Normal('α', 0, sd=10)
β = Normal('β', 0, sd=10)
```
Then, apply the `invlogit` transformation to the linear combination of predictors:
```
from pymc3.math import invlogit
with ivh_glm:
p = invlogit(μ + α*bwt_kg + β*pneumo)
```
Finally, the sampling distribution:
```
from pymc3 import Bernoulli
with ivh_glm:
bb_like = Bernoulli('bb_like', p=p, observed=ivh)
with ivh_glm:
trace_ivh = sample(1000, random_seed=RANDOM_SEED)
plot_posterior(trace_ivh);
```
## Interactions among variables
Interactions imply that the effect of one covariate $X_1$ on $Y$ depends on the value of another covariate $X_2$.
$$M(Y|X) = \beta_0 + \beta_1 X_1 + \beta_2 X_2 +\beta_3 X_1 X_2$$
the effect of a unit increase in $X_1$:
$$M(Y|X_1+1, X_2) - M(Y|X_1, X_2)$$
$$\begin{align}
&= \beta_0 + \beta_1 (X_1 + 1) + \beta_2 X_2 +\beta_3 (X_1 + 1) X_2
- [\beta_0 + \beta_1 X_1 + \beta_2 X_2 +\beta_3 X_1 X_2] \\
&= \beta_1 + \beta_3 X_2
\end{align}$$
```
ax = medals[medals.oecd==1].plot(x='log_population', y='medals', kind='scatter', alpha=0.8)
medals[medals.oecd==0].plot(x='log_population', y='medals', kind='scatter', color='red', alpha=0.5, ax=ax)
```
Interaction can be interpreted as:
- $X_1$ interacts with $X_2$
- $X_1$ modifies the effect of $X_2$
- $X_2$ modifies the effect of $X_1$
- $X_1$ and $X_2$ are non-additive or synergistic
Let's construct a model that predicts medal count based on population size and OECD status, as well as the interaction.
We can use the `dmatrix` function from the `patsy` library to set up a **design matrix**. This is a matrix of independent observations (rows) and predictors (columns) that defines the input to the model.
```
from patsy import dmatrix
y = medals.medals
X = dmatrix('log_population * oecd', data=medals)
X
```
This matrix is multiplied by the vector of model coefficients to generate an estimate of the response corresponding to each row.
Now, fit the model.
```
interaction_params = fmin(poisson_loglike, [0,1,1,0], args=(X, y), maxiter=500)
interaction_params
```
Notice anything odd about these estimates?
The main effect of the OECD effect is negative, which seems counter-intuitive. This is because the variable is interpreted as the OECD effect when the **log-population is zero**. This is not particularly meaningful.
Asw we did with birth weight in the previous example, we can improve the interpretability of this parameter by centering the log-population variable prior to entering it into the model. This will result in the OECD main effect being interpreted as the marginal effect of being an OECD country for an **average-sized** country.
```
y = medals.medals
X = dmatrix('center(log_population) * oecd', data=medals)
X
fmin(poisson_loglike, [0,1,1,0], args=(X, y))
```
## Model Selection
How do we choose among competing models for a given dataset? More parameters are not necessarily better, from the standpoint of model fit. For example, fitting a 6th order polynomial to the LSD example certainly results in an overfit.
```
def calc_poly(params, data):
x = np.c_[[data**i for i in range(len(params))]]
return np.dot(params, x)
x, y = lsd_and_math.T.values
sum_squares_poly = lambda θ, x, y: np.sum((y - calc_poly(θ, x)) ** 2)
betas = fmin(sum_squares_poly, np.zeros(7), args=(x,y), maxiter=1e6)
plt.plot(x, y, 'ro')
xvals = np.linspace(0, max(x), 100)
plt.plot(xvals, calc_poly(betas, xvals))
```
One approach is to use an information-theoretic criterion to select the most appropriate model. For example **Akaike's Information Criterion (AIC)** balances the fit of the model (in terms of the likelihood) with the number of parameters required to achieve that fit. We can easily calculate AIC as:
$$AIC = n \log(\hat{\sigma}^2) + 2p$$
where $p$ is the number of parameters in the model and $\hat{\sigma}^2 = RSS/(n-p-1)$.
Notice that as the number of parameters increase, the residual sum of squares goes down, but the second term (a penalty) increases.
AIC is a metric of **information distance** between a given model and a notional "true" model. Since we don't know the true model, the AIC value itself is not meaningful in an absolute sense, but is useful as a relative measure of model quality.
To apply AIC to model selection, we choose the model that has the **lowest** AIC value.
```
n = len(x)
aic = lambda rss, p, n: n * np.log(rss/(n-p-1)) + 2*p
RSS1 = sum_of_squares(fmin(sum_of_squares, [0,1], args=(x,y)), x, y)
RSS2 = sum_squares_quad(fmin(sum_squares_quad, [1,1,-1], args=(x,y)), x, y)
print('\nModel 1: {0}\nModel 2: {1}'.format(aic(RSS1, 2, n), aic(RSS2, 3, n)))
```
Hence, on the basis of "information distance", we would select the 2-parameter (linear) model.
### Exercise: Olympic medals model selection
Use AIC to select the best model from the following set of Olympic medal prediction models:
- population only
- population and OECD
- interaction model
For these models, use the alternative form of AIC, which uses the log-likelhood rather than the residual sums-of-squares:
$$AIC = -2 \log(L) + 2p$$
```
# Write your answer here
```
## References
- Harrell, F. E. (2015). Regression Modeling Strategies (pp. 1–397).
- Gelman, A., & Hill, J. (2006). Data Analysis Using Regression and Multilevel/Hierarchical Models (1st ed.). Cambridge University Press.
| github_jupyter |
```
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
<a target="_blank" href="https://colab.research.google.com/github/GoogleCloudPlatform/keras-idiomatic-programmer/blob/master/workshops/Training/Idiomatic%20Programmer%20-%20handbook%203%20-%20Codelab%202.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# Idiomatic Programmer Code Labs
## Code Labs #2 - Get Familiar with Training
## Prerequistes:
1. Familiar with Python
2. Completed Handbook 3/Part 11: Training & Deployment
## Objectives:
1. Pretraining for Weight Initialization
2. Early Stopping during Training
3. Model Saving and Restoring
## Pretraining
We are going to do some pre-training runs to find a good initial weight initialization. Each time the weights are initialized, they are randomly choosen from the selected distribution (i.e., kernel_initializer).
We will do the following:
1. Make three instances of the same model, each with their own weight initialization.
2. Take a subset of the training data (20%)
3. Train each model instance for a few epochs.
4. Pick the instance with the highest valuation accuracy.
5. Use this instance to train the model with the entire training data.
You fill in the blanks (replace the ??), make sure it passes the Python interpreter.
```
from keras import Sequential, optimizers
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout
from keras.utils import to_categorical
from keras.datasets import cifar10
import numpy as np
# Let's use the CIFAR-10 dataset
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# Normalize the pixel data
x_train = (x_train / 255.0).astype(np.float32)
x_test = (x_test / 255.0).astype(np.float32)
# One-hot encode the labels
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# Let's take a fraction of the training data to test the weight initialization (20%)
# Generally, we like to use all the training data for this purpose, but for brevity we will use 20%
x_tmp = x_train[0:10000]
y_tmp = y_train[0:10000]
# We will use this function to build a simple CNN, using He-Normal initialization for the weights.
def convNet(input_shape, nclasses):
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', kernel_initializer='he_normal',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal'))
model.add(Dropout(0.25))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu', kernel_initializer='he_normal'))
model.add(Dense(nclasses, activation='softmax'))
return model
# Let's make 3 versions of the model, each with their own weight initialization.
models = []
for _ in range(3):
model = convNet((32, 32, 3), 10)
# We will use (assume best) learning rate of 0.001
model.compile(loss='categorical_crossentropy', optimizer=optimizers.Adam(lr=0.001), metrics=['accuracy'])
# Let's do the short training of 20% of training data for 5 epochs.
model.fit(x_tmp, y_tmp, epochs=5, batch_size=32, validation_split=0.1, verbose=1)
# Save a copy of the model
# HINT: We are saving the in-memory partially trained model
models.append(??)
# Now let's pick the model instance with the highest val_acc and train it with the full training data
# HINT: Index will be 0, 1 or 2
models[1].fit(x_train, y_train, epochs=10, batch_size=32, validation_split=0.1, verbose=1)
score = model.evaluate(x_test, y_test)
print(score)
```
## EarlyStopping
Note that the training accuracy in the above example keeps going up, but at some point the validation loss swings back up and validation goes down. That means you are overfitting -- even with the dropout.
Let's now look on how to decide how many epochs we should run. We can use early stopping technique. In this case, we set the number of epochs larger than we anticipate, and then set an objective to reach. When the objective is reached, we stop training.
```
from keras.callbacks import EarlyStopping
# Let's try this with a fresh model, and not care about the weight initialization this time.
model = convNet((32, 32, 3), 10)
model.compile(loss='categorical_crossentropy', optimizer=optimizers.Adam(lr=0.001), metrics=['accuracy'])
# Set an early stop (termination of training) when the valuation loss has stopped
# reducing (default setting).
earlystop = EarlyStopping(monitor='val_loss')
# Train the model and use early stop to stop training early if the valuation loss
# stops decreasing.
# HINT: what goes in the callbacks list is the instance (variable) of the EarlyStopping object
model.fit(x_train, y_train, epochs=10, batch_size=32, validation_split=0.1, verbose=1, callbacks=[??])
```
## Model Saving and Restoring
Let's do a basic store of the model and weights to disk, and then mimic restoring the model from disk to in memory.
```
from keras.models import load_model
# Save the model and trained weights and biases.
model.save('mymodel.h5')
# load a pre-trained model as a different model instance (mymodel instead of model)
mymodel = load_model('mymodel.h5')
# Let's verify we really do that.
score = mymodel.evaluate(x_test, y_test)
print(score)
```
## End of Code Lab
| github_jupyter |
# Chapter 1 - Importing Data in Python (Part 2)
### The web is a rich source of data from which you can extract various types of insights and findings. In this chapter, you will learn how to get data from the web, whether it be stored in files or in HTML. You'll also learn the basics of scraping and parsing web data.
### Importing flat files from the web: your turn!
You are about to import your first file from the web! The flat file you will import will be `'winequality-red.csv'` from the University of California, Irvine's [Machine Learning repository](http://archive.ics.uci.edu/ml/index.php). The flat file contains tabular data of physiochemical properties of red wine, such as pH, alcohol content and citric acid content, along with wine quality rating.
The URL of the file is
'https://s3.amazonaws.com/assets.datacamp.com/production/course_1606/datasets/winequality-red.csv'
After you import it, you'll check your working directory to confirm that it is there and then you'll load it into a pandas DataFrame.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
!ls
# Import package
from urllib.request import urlretrieve
# Assign url of file: url
url = 'https://s3.amazonaws.com/assets.datacamp.com/production/course_1606/datasets/winequality-red.csv'
# Save file locally
urlretrieve(url, 'winequality-red.csv')
# Read file into a DataFrame and print its head
df = pd.read_csv('winequality-red.csv', sep=';')
df.head()
!ls
```
### Opening and reading flat files from the web
You have just imported a file from the web, saved it locally and loaded it into a DataFrame. If you just wanted to load a file from the web into a DataFrame without first saving it locally, you can do that easily using pandas. In particular, you can use the function pd.read_csv() with the URL as the first argument and the separator sep as the second argument.
The URL of the file, once again, is
'https://s3.amazonaws.com/assets.datacamp.com/production/course_1606/datasets/winequality-red.csv'
```
import warnings
warnings.filterwarnings('ignore')
# Assign url of file: url
url = 'https://s3.amazonaws.com/assets.datacamp.com/production/course_1606/datasets/winequality-red.csv'
# Read file into a DataFrame: df
df = pd.read_csv(url, sep = ';')
# Print the head of the DataFrame
print(df.head())
# Plot first column of df
pd.DataFrame.hist(df.ix[:, 0:1])
plt.xlabel('fixed acidity (g(tartaric acid)/dm$^3$)')
plt.ylabel('count')
```
### Importing non-flat files from the web
Congrats! You've just loaded a flat file from the web into a DataFrame without first saving it locally using the pandas function pd.read_csv(). This function is super cool because it has close relatives that allow you to load all types of files, not only flat ones. In this interactive exercise, you'll use pd.read_excel() to import an Excel spreadsheet.
The URL of the spreadsheet is
'http://s3.amazonaws.com/assets.datacamp.com/course/importing_data_into_r/latitude.xls'
Your job is to use pd.read_excel() to read in all of its sheets, print the sheet names and then print the head of the first sheet using its name, not its index.
Note that the output of pd.read_excel() is a Python dictionary with sheet names as keys and corresponding DataFrames as corresponding values.
__Instructions__
- Assign the URL of the file to the variable url.
- Read the file in url into a dictionary xl using pd.read_excel() recalling that, in order to import all sheets you need to pass None to the argument sheetname.
- Print the names of the sheets in the Excel spreadsheet; these will be the keys of the dictionary xl.
- Print the head of the first sheet using the sheet name, not the index of the sheet! The sheet name is '1700'
```
# Assign url of file: url
url = 'http://s3.amazonaws.com/assets.datacamp.com/course/importing_data_into_r/latitude.xls'
# Read in all sheets of Excel file: xl
x1 = pd.read_excel(url, sheetname = None)
# Print the sheetnames to the shell
print(x1.keys())
# Print the head of the first sheet (using its name, NOT its index)
x1['1700'].head()
```
### Performing HTTP requests in Python using urllib
Now that you know the basics behind HTTP GET requests, it's time to perform some of your own. In this interactive exercise, you will ping our very own DataCamp servers to perform a GET request to extract information from our teach page, "http://www.datacamp.com/teach/documentation".
In the next exercise, you'll extract the HTML itself. Right now, however, you are going to package and send the request and then catch the response.
```
# Import packages
from urllib.request import urlopen, Request
# Specify the url
url = "http://www.datacamp.com/teach/documentation"
# This packages the request: request
request = Request(url)
# Sends the request and catches the response: response
response = urlopen(request)
# Print the datatype of response
print(type(response))
# Be polite and close the response!
response.close()
```
### Printing HTTP request results in Python using urllib
You have just packaged and sent a GET request to "http://www.datacamp.com/teach/documentation" and then caught the response. You saw that such a response is a http.client.HTTPResponse object. The question remains: what can you do with this response?
Well, as it came from an HTML page, you could read it to extract the HTML and, in fact, such a http.client.HTTPResponse object has an associated read() method. In this exercise, you'll build on your previous great work to extract the response and print the HTML.
```
# Import packages
from urllib.request import urlopen, Request
# Specify the url
url = "http://www.datacamp.com/teach/documentation"
# This packages the request
request = Request(url)
# Sends the request and catches the response: response
response = urlopen(request)
# Extract the response: html
html = response.read()
# Print the html
print(html)
# Be polite and close the response!
response.close()
```
### Performing HTTP requests in Python using requests
Now that you've got your head and hands around making HTTP requests using the urllib package, you're going to figure out how to do the same using the higher-level requests library. You'll once again be pinging DataCamp servers for their "http://www.datacamp.com/teach/documentation" page.
Note that unlike in the previous exercises using urllib, you don't have to close the connection when using requests!
```
# Import package
import requests
# Specify the url: url
url = "http://www.datacamp.com/teach/documentation"
# Packages the request, send the request and catch the response: r
r = requests.get(url)
# Extract the response: text
text = r.text
# Print the html
print(text)
```
### Parsing HTML with BeautifulSoup
In this interactive exercise, you'll learn how to use the BeautifulSoup package to parse, prettify and extract information from HTML. You'll scrape the data from the webpage of Guido van Rossum, Python's very own [Benevolent Dictator for Life](https://en.wikipedia.org/wiki/Benevolent_dictator_for_life). In the following exercises, you'll prettify the HTML and then extract the text and the hyperlinks.
The URL of interest is url = 'https://www.python.org/~guido/'.
```
#! pip install bs4
# Import packages
import requests
from bs4 import BeautifulSoup
# Specify url: url
url = 'https://www.python.org/~guido/'
# Package the request, send the request and catch the response: r
r = requests.get(url)
# Extracts the response as html: html_doc
html_doc = r.text
# Create a BeautifulSoup object from the HTML: soup
soup = BeautifulSoup(html_doc)
# Prettify the BeautifulSoup object: pretty_soup
pretty_soup = soup.prettify()
# Print the response
print(pretty_soup)
```
### Turning a webpage into data using BeautifulSoup: getting the text
As promised, in the following exercises, you'll learn the basics of extracting information from HTML soup. In this exercise, you'll figure out how to extract the text from the BDFL's webpage, along with printing the webpage's title.
```
# Get the title of Guido's webpage: guido_title
guido_title = soup.title
# Print the title of Guido's webpage to the shell
print(guido_title)
# Get Guido's text: guido_text
guido_text = soup.get_text()
# Print Guido's text to the shell
print(guido_text)
```
### Turning a webpage into data using BeautifulSoup: getting the hyperlinks
In this exercise, you'll figure out how to extract the URLs of the hyperlinks from the BDFL's webpage. In the process, you'll become close friends with the soup method find_all().
```
# Print the title of Guido's webpage
print(soup.title)
# Find all 'a' tags (which define hyperlinks): a_tags
a_tags = soup.find_all('a')
# Print the URLs to the shell
for link in a_tags:
print(link.get('href'))
```
---
# Chapter 2 - Interacting with APIs to import data from the web
### In this chapter, you will push further on your knowledge of importing data from the web. You will learn the basics of extracting data from APIs, gain insight on the importance of APIs and practice getting data from them with dives into the OMDB and Library of Congress APIs.
### JSONs
- JSONs consist of key-value pairs.
- JSONs are human-readable.
- The JSON file format arose out of a growing need for real-time server-to-browser communication.
- The function json.load() will load the JSON into Python as a dictionary.
### Loading and exploring a JSON
Now that you know what a JSON is, you'll load one into your Python environment and explore it yourself. Here, you'll load the JSON 'a_movie.json' into the variable json_data, which will be a dictionary. You'll then explore the JSON contents by printing the key-value pairs of json_data to the shell.
```
# Load JSON: json_data
with open("a_movie.json") as json_file:
json_data = json.load(json_file)
# Print each key-value pair in json_data
for k in json_data.keys():
print(k + ': ',json_data[k])
```
### What's an API
- An API is a set of protocols and routines for building and interacting with software applications.
- API is an acronym and is short for Application Program interface.
- It is common to pull data from APIs in the JSON file format.
- An API is a bunch of code that allows two software programs to communicate with each other.
### API requests
Now it's your turn to pull some movie data down from the Open Movie Database (OMDB) using their API. The movie you'll query the API about is The Social Network.
Note: recently, OMDB has changed their API: you now also have to specify an API key. This means you'll have to add another argument to the URL: `apikey=72bc447a`
```
# Import requests package
import requests
# Assign URL to variable: url
url = 'http://www.omdbapi.com/?apikey=72bc447a&t=the+social+network'
# Package the request, send the request and catch the response: r
r = requests.get(url)
# Print the text of the response
print(r.text)
```
### JSON–from the web to Python
Wow, congrats! You've just queried your first API programmatically in Python and printed the text of the response to the shell. However, as you know, your response is actually a JSON, so you can do one step better and decode the JSON. You can then print the key-value pairs of the resulting dictionary. That's what you're going to do now!
```
# Import package
import requests
# Assign URL to variable: url
url = 'http://www.omdbapi.com/?apikey=72bc447a&t=social+network'
# Package the request, send the request and catch the response: r
r = requests.get(url)
# Decode the JSON data into a dictionary: json_data
json_data = r.json()
# Print each key-value pair in json_data
for k in json_data.keys():
print(k + ': ', json_data[k])
```
### Checking out the Wikipedia API
You're doing so well and having so much fun that we're going to throw one more API at you: the Wikipedia API (documented [here](https://www.mediawiki.org/wiki/API:Main_page). You'll figure out how to find and extract information from the Wikipedia page for Pizza. What gets a bit wild here is that your query will return nested JSONs, that is, JSONs with JSONs, but Python can handle that because it will translate them into dictionaries within dictionaries.
The URL that requests the relevant query from the Wikipedia API is
https://en.wikipedia.org/w/api.php?action=query&prop=extracts&format=json&exintro=&titles=pizza
```
# Assign URL to variable: url
url = 'https://en.wikipedia.org/w/api.php?action=query&prop=extracts&format=json&exintro=&titles=pizza'
# Package the request, send the request and catch the response: r
r = requests.get(url)
# Decode the JSON data into a dictionary: json_data
json_data = r.json()
# Print the Wikipedia page extract
pizza_extract = json_data['query']['pages']['24768']['extract']
print(pizza_extract)
```
# Chapter 3 - Diving deep into the Twitter API
### In this chapter, you will consolidate your knowledge of interacting with APIs in a deep dive into the Twitter streaming API. You'll learn how to stream real-time Twitter data and to analyze and visualize it!
### API Authentication
The package tweepy is great at handling all the Twitter API OAuth Authentication details for you. All you need to do is pass it your authentication credentials. In this interactive exercise, we have created some mock authentication credentials. Your task is to pass these credentials to tweepy's OAuth handler.
__Instructions__
- Import the package tweepy.
- Pass the parameters consumer_key and consumer_secret to the function tweepy.OAuthHandler().
- Complete the passing of OAuth credentials to the OAuth handler auth by applying to it the method set_access_token(), along with arguments access_token and access_token_secret.
```
#! pip install tweepy
# Import package
import tweepy
# Store OAuth authentication credentials in relevant variables
access_token = "1092294848-aHN7DcRP9B4VMTQIhwqOYiB14YkW92fFO8k8EPy"
access_token_secret = "X4dHmhPfaksHcQ7SCbmZa2oYBBVSD2g8uIHXsp5CTaksx"
consumer_key = "nZ6EA0FxZ293SxGNg8g8aP0HM"
consumer_secret = "fJGEodwe3KiKUnsYJC3VRndj7jevVvXbK2D5EiJ2nehafRgA6i"
# Pass OAuth details to tweepy's OAuth handler
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
```
### Streaming tweets
Now that you have set up your authentication credentials, it is time to stream some tweets! We have already defined the tweet stream listener class, MyStreamListener. You can find the code for the tweet stream listener class [here](https://gist.github.com/hugobowne/18f1c0c0709ed1a52dc5bcd462ac69f4).
Your task is to create the Streamobject and to filter tweets according to particular keywords.
```
class MyStreamListener(tweepy.StreamListener):
def __init__(self, api=None):
super(MyStreamListener, self).__init__()
self.num_tweets = 0
self.file = open("tweets.txt", "w")
def on_status(self, status):
tweet = status._json
self.file.write( json.dumps(tweet) + '\n' )
self.num_tweets += 1
if self.num_tweets < 100:
return True
else:
return False
self.file.close()
def on_error(self, status):
print(status)
# Initialize Stream listener
l = MyStreamListener()
# Create your Stream object with authentication
stream = tweepy.Stream(auth, l)
# Filter Twitter Streams to capture data by the keywords:
stream.filter(track = ['clinton', 'trump', 'sanders', 'cruz']);
# Import package
import json
# String of path to file: tweets_data_path
tweets_data_path = 'tweets3.txt'
# Initialize empty list to store tweets: tweets_data
tweets_data = []
# Open connection to file
tweets_file = open(tweets_data_path, "r")
# Read in tweets and store in list: tweets_data
for line in tweets_file:
tweet = json.loads(line)
tweets_data.append(tweet)
# Close connection to file
tweets_file.close()
# Print the keys of the first tweet dict
print(tweets_data[0].keys())
```
### Twitter data to DataFrame
Now you have the Twitter data in a list of dictionaries, tweets_data, where each dictionary corresponds to a single tweet. Next, you're going to extract the text and language of each tweet. The text in a tweet, t1, is stored as the value t1['text']; similarly, the language is stored in t1['lang']. Your task is to build a DataFrame in which each row is a tweet and the columns are 'text' and 'lang'.
```
# Build DataFrame of tweet texts and languages
df = pd.DataFrame(tweets_data, columns=['text', 'lang'])
# Print head of DataFrame
df.head()
```
### A little bit of Twitter text analysis
Now that you have your DataFrame of tweets set up, you're going to do a bit of text analysis to count how many tweets contain the words 'clinton', 'trump', 'sanders' and 'cruz'. In the pre-exercise code, we have defined the following function word_in_text(), which will tell you whether the first argument (a word) occurs within the 2nd argument (a tweet).
```python
import re
def word_in_text(word, text):
word = word.lower()
text = tweet.lower()
match = re.search(word, text)
if match:
return True
return False
```
You're going to iterate over the rows of the DataFrame and calculate how many tweets contain each of our keywords! The list of objects for each candidate has been initialized to 0.
```
import re
def word_in_text(word, text):
word = word.lower()
text = text.lower()
match = re.search(word, text)
if match:
return True
return False
# Initialize list to store tweet counts
[clinton, trump, sanders, cruz] = [0, 0, 0, 0]
# Iterate through df, counting the number of tweets in which
# each candidate is mentioned
for index, row in df.iterrows():
clinton += word_in_text('clinton', row['text'])
trump += word_in_text('trump', row['text'])
sanders += word_in_text('sanders', row['text'])
cruz += word_in_text('cruz', row['text'])
```
### Plotting your Twitter data
Now that you have the number of tweets that each candidate was mentioned in, you can plot a bar chart of this data. You'll use the statistical data visualization library seaborn, which you may not have seen before, but we'll guide you through. You'll first import seaborn as sns. You'll then construct a barplot of the data using sns.barplot, passing it two arguments:
1. a list of labels and
2. a list containing the variables you wish to plot (clinton, trump and so on.)
Hopefully, you'll see that Trump was unreasonably represented!
```
clinton, trump, sanders, cruz
# Import packages
import matplotlib.pyplot as plt
import seaborn as sns
# Set seaborn style
sns.set(color_codes=True)
# Create a list of labels:cd
cd = ['clinton', 'trump', 'sanders', 'cruz']
# Plot histogram
ax = sns.barplot(cd, [clinton, trump, sanders, cruz])
ax.set(ylabel="count")
plt.show()
```
| github_jupyter |
# LSTM Experiment
This notebook is an entry point to reproduce the experiments describe in the LSTM section of the paper.
It consists of three sections:
1) Data Loading
2) Example on how to train and test a LSTM on our data
3) The actual experiment code from the paper
```
%load_ext autoreload
%autoreload 2
# For use in google colab
!pip install python-Levenshtein
import DataSet
import Evaluation
import datetime
import numpy as np
import matplotlib.pyplot as plt
import random
from matplotlib.backends.backend_pdf import PdfPages
from Utils import *
from DataSet import UniHHIMUGestures
from torch.utils.data import Dataset, DataLoader
import torch
import torch.nn as nn
import torch.optim as optim
def getProjectPath():
return './'
```
We need to specify a bunch of parameters the expriment:
```
#===========================================================================
# Give this run a name.
# If name equals 'test', no log will be generated
#===========================================================================
name = 'test'
#===========================================================================
# Decide which gesture data shall be used for training
#===========================================================================
inputGestures = [0,1,2,3,4,5,6,7,8,9]
#===========================================================================
# Decide which target signals shall be used for training
#===========================================================================
usedGestures = [0,1,2,3,4,5,6,7,8,9]
#===========================================================================
# Concatenate data to create "more" training samples, 1 corresponds to no concatenations
#===========================================================================
concFactor = 1
#===========================================================================
# Add noise to the data, 0 corresponds to no noise. Noise above 2 has shown to weaken recognition
#===========================================================================
noiseFactor = 1
#===========================================================================
# Decide wether gestures shall be shuffled before training. If true, nFolds many
# pieces will be generated. Not every piece is garanteed to contain every gesture, so do not use too many.
#===========================================================================
shuffle = True
nFolds = 4
#===========================================================================
# Function used to evaluate during cross validation. Possible functions are:
# Evaluation.calc1MinusF1FromMaxApp (best working, used in thesis)
# Oger.utils.nmse (normalised mean square error, tells nothing about classifier perfomance but works okay)
# Evaluation.calcLevenshteinError (use the Levenshtein error, disadvantages are highlighted in thesis)
# Evaluation.calc1MinusF1FromInputSegment (use segmentation by supervised signal)
#===========================================================================
evaluationFunction = Evaluation.calc1MinusF1FromMaxApp
#===========================================================================
# Set this to true if another output neuron shall be added to represent "no gesture"
#===========================================================================
learnTreshold = False
#===========================================================================
# Use on of the optimisation dictionaries from the optDicts file
#===========================================================================
optDict = 'bestParas'
#===========================================================================
# Use normalizer
#===========================================================================
useNormalized = 2
#===========================================================================
# Pick datasets to train on, and datasets to test on
#===========================================================================
inputFiles = ['ni','j','na','l']
testFiles = ['s']
# If desired add a specific file to test on, e.g. randTestFiles = ['lana_0_0.npz']
randTestFiles = []
#===========================================================================
# Setup project directory
#===========================================================================
now = datetime.datetime.now()
resultsPath = getProjectPath()+'results/'
pdfFileName = now.strftime("%Y-%m-%d-%H-%M")+'_'+name+'.pdf'
pdfFilePath = resultsPath+'pdf/'+pdfFileName
npzFileName = now.strftime("%Y-%m-%d-%H-%M")+'_'+name+'.npz'
npzFilePath = resultsPath+'npz/'+npzFileName
bestFlowPath = resultsPath+'nodes/'+now.strftime("%Y-%m-%d-%H-%M")+'_'+name+'.p'
pp = PdfPages(pdfFilePath)
#===========================================================================
# Add labels for gestures
#===========================================================================
totalGestureNames = ['left','right','forward','backward','bounce up','bounce down',
'turn left','turn right','shake lr','shake ud',
'tap 1','tap 2','tap 3','tap 4','tap 5','tap 6','no gesture']
gestureNames = []
for i in usedGestures:
gestureNames.append(totalGestureNames[i])
gestureNames.append('no gesture')
```
Create dataset and dataloaders
```
def createData(inputFiles, testFiles, validation_size=10):
trainset_ = UniHHIMUGestures(dataDir=getProjectPath() + 'dataSets/',
train=True,
inputFiles=inputFiles,
testFiles=testFiles,
useNormalized=useNormalized,
learnTreshold=learnTreshold,
nFolds=100,
shuffle=True,
)
testset = UniHHIMUGestures(dataDir=getProjectPath() + 'dataSets/',
train=False,
inputFiles=inputFiles,
testFiles=testFiles,
useNormalized=useNormalized,
learnTreshold=learnTreshold,
shuffle=True
)
trainset, validationset = torch.utils.data.random_split(trainset_, [len(trainset_)-validation_size,validation_size])
trainloader = DataLoader(trainset, batch_size=1,
shuffle=True, num_workers=1)
validationloader = DataLoader(validationset, batch_size=1,
shuffle=True, num_workers=1)
testloader = DataLoader(testset, batch_size=1,
shuffle=True, num_workers=1)
return trainset, validationset, testset, trainloader, validationloader, testloader
trainset, validationset, testset, trainloader, validationloader, testloader = createData(inputFiles, testFiles)
```
Let's take a look at the scaled input data:
```
fig, ax = plt.subplots(3,1, figsize=(15,9))
ax[0].plot(trainset[0][0][:800,0:3])
ax[1].plot(trainset[0][0][:800,3:6])
ax[2].plot(trainset[0][0][:800,6:9])
plt.tight_layout()
```
Looks all good, we can clearly see the gesture sequences intercepted by non gesture seqeuences in between.
Now let's create a Leaky Integrator ESN with parameters as defined in the paper.
## Section 2: Train and test LSTM
This section shows how to create and train LSTM on the give data.
```
class LSTMClassifier(nn.Module):
def __init__(self, hidden_dim=25):
super(LSTMClassifier, self).__init__()
self.lstm = torch.nn.LSTM(input_size=9, hidden_size=hidden_dim, batch_first=True)
self.classifier = torch.nn.Sequential(
torch.nn.Linear(hidden_dim,10, bias=False),
#torch.nn.Sigmoid()
)
def forward(self, inputs):
lstm_outputs, (hidden_acts, cell_states) = self.lstm(inputs)
predictions = self.classifier(lstm_outputs)
return predictions
lstm = LSTMClassifier(20)
loss_function = torch.nn.MSELoss()
optimizer = optim.Adam(lstm.parameters(), lr=1e-3)
lstm.cuda()
import sklearn
def testLSTM(testloader, lstm, plot=True, plotConf=False):
testF1MaxApps = []
testAccuracies = []
testCms = []
losses = []
for inputs, targets in testloader:
test_inputs, test_targets = inputs.float(), targets.float()
#inputs[0,:,:9] = targets[0,:,:9]
test_inputs = (test_inputs/test_inputs.std(1))
test_inputs = test_inputs.cuda()
test_targets = test_targets.cuda()
lstm.eval()
outputs = lstm(test_inputs)
loss = loss_function(outputs, test_targets)
losses.append(loss.item())
test_preds = outputs.cpu().detach().numpy()[0]
if plot:
plt.figure(figsize=(15,5))
plt.plot(test_preds[:1600,:])
plt.plot(test_targets[0,:1600,:].cpu())
plt.pause(1)
fixed_threshold = 0.4
t_target = test_targets[0].cpu().numpy()
prediction = test_preds[:,:10]
if learnTreshold: # if threshold is learned, then it's the las collumn of the prediction
threshold = outputs[0].numpy()[:,10]
else: #else add a constant threshold
threshold = np.ones((prediction.shape[0],1))*fixed_threshold
t_maxApp_prediction = Evaluation.calcMaxActivityPrediction(prediction,t_target,threshold, 10)
pred_MaxApp, targ_MaxApp = Evaluation.calcInputSegmentSeries(t_maxApp_prediction, t_target, 0.5)
testF1MaxApps.append(np.mean(sklearn.metrics.f1_score(targ_MaxApp,pred_MaxApp,average=None)))
testAccuracies.append(np.mean(sklearn.metrics.accuracy_score(targ_MaxApp,pred_MaxApp)))
if False:
#print(t_maxApp_prediction.shape, prediction.shape, pred_MaxApp, targ_MaxApp)
plt.figure(figsize=(20,3))
plt.plot(t_maxApp_prediction[:800])
plt.plot(t_target[:800])
conf = sklearn.metrics.confusion_matrix(targ_MaxApp, pred_MaxApp)
testCms.append(conf)
if plotConf:
Evaluation.plot_confusion_matrix(testCms[0], gestureNames, 'test set')
plt.tight_layout()
plt.ylim(10.5,-0.5)
#print("Test f1 score for maxactivity: {:.4f}, MSE: {:.4f}".format(np.mean(testF1MaxApps),loss.item()))
return np.mean(testF1MaxApps), np.mean(losses), np.mean(testAccuracies), testCms
testLSTM(testloader, lstm)
import copy
def trainLSTM(inputFiles, testFiles, plot=True):
trainset, validationset, testset, trainloader, validationloader, testloader = createData(inputFiles, testFiles)
lstm = LSTMClassifier(80)
loss_function = torch.nn.MSELoss()
optimizer = optim.Adam(lstm.parameters(), lr=1e-3)
lstm.cuda()
best_score = 0.
best_model = lstm.state_dict()
train_losses = []
test_losses = []
test_scores = []
for epoch in range(100):
epoch_losses = []
for inputs, targets in trainloader:
lstm.train()
inputs, targets = inputs.float(), targets.float()
#inputs[0,:,:9] = targets[0,:,:9]
inputs = (inputs/inputs.std(1))
inputs = inputs[:,:,:].cuda()
targets = targets[:,:,:].cuda()
lstm.zero_grad()
outputs = lstm(inputs)
loss = loss_function(outputs, targets)
loss.backward()
epoch_losses.append(loss.item())
optimizer.step()
train_losses.append(epoch_losses)
score, mse, accuracies, cm = testLSTM(validationloader, lstm, plot=False)
test_losses.append(mse.item())
test_scores.append(score.item())
if score > best_score:
best_model = copy.deepcopy(lstm.state_dict())
torch.save(lstm.state_dict(), 'weights_only.pth')
best_score = score
if score + 0.1 < best_score:
print('Overfitted, breaking the loop')
break
if epoch % 25 == 0:
print(loss.item())
lstm.eval()
plt.figure(figsize=(15,5))
plt.title('Epoch: {:}, loss: {:.4f}'.format(epoch, loss))
#test_preds = lstm(test_inputs.float()).detach().numpy()[0]
#plt.plot(test_preds[:800,:])
#plt.plot(test_targets[0,:800,:])
preds = lstm(inputs)
if plot:
plt.plot(preds[0,:800].cpu().detach())
plt.plot(targets[0,:800].cpu().detach())
plt.ylim(-0.1,1.1)
plt.pause(1)
score, mse, accuracies, cm = testLSTM(validationloader,lstm, plot=False)
print("Best score: {:.2f}, current score: {:.2f}".format(best_score, score, mse))
plt.plot([np.mean(epoch_losses) for epoch_losses in train_losses])
plt.plot([x for x in test_losses])
lstm.load_state_dict(best_model)
train_score, train_mse, train_accuracy, train_cm = testLSTM(trainloader, lstm, plot=False, plotConf=False)
score, mse, accuracy, cm = testLSTM(testloader, lstm, plot=plot, plotConf=plot)
print("Best score: {:.2f}, current score: {:.2f}".format(best_score, score, mse))
print('###################################################################')
return score, mse, accuracy, cm, train_score, train_mse, train_accuracy, lstm
inputFiles = ['s','ni','na','l']
testFiles = ['j']
scores = []
for trial in range(1):
print('################# TRIAL: {} #########################'.format(trial))
score, mse, accuracy, cm, train_score, train_mse, train_accuracy, lstm = trainLSTM(inputFiles=inputFiles, testFiles=testFiles, plot=True)
scores.append(score)
best_lstm = LSTMClassifier(80)
best_lstm.load_state_dict(torch.load('weights_only.pth'))
best_lstm.cuda().eval()
testLSTM(testloader, best_lstm, plot=True, plotConf=True)
score, mse, accuracies, cms = testLSTM(testloader, best_lstm, plot=True)
```
## Section 3: Run experiment from paper
In this experiment we train an LSTM on four of the testsets and evaluate on the fith one. Training and evaluation is repeated several times to average the results.
```
# Prepare arrays to store evaualtion results
files = ['s','j','na','l','ni']
trials = 10
if False:
all_scores = np.zeros((len(files),trials))
all_train_scores = np.zeros((len(files),trials))
all_cms = np.zeros((len(files),trials, 11, 11))
all_accuracies = np.zeros((len(files),trials))
all_train_accuracies = np.zeros((len(files),trials))
np.save(file=getProjectPath()+"lstm_scores", arr=np.array(all_scores))
np.save(file=getProjectPath()+"lstm_train_scores", arr=np.array(all_train_scores))
np.save(file=getProjectPath()+"lstm_cms", arr=np.array(all_cms))
np.save(file=getProjectPath()+"lstm_accuracies", arr=np.array(all_accuracies))
np.save(file=getProjectPath()+"lstm_train_accuracies", arr=np.array(all_train_accuracies))
if False:
all_scores = np.load("lstm_scores.npy")
all_train_scores = np.load("lstm_train_scores.npy")
all_cms = np.load("lstm_cms.npy")
all_accuracies = np.load("lstm_accuracies.npy")
all_train_accuracies = np.load("lstm_train_accuracies.npy")
from tqdm import tqdm_notebook
def fix_seed(manualSeed):
np.random.seed(manualSeed)
random.seed(manualSeed)
torch.manual_seed(manualSeed)
# if you are using GPU
torch.cuda.manual_seed(manualSeed)
torch.cuda.manual_seed_all(manualSeed)
# run evaluation for each testset
fix_seed(1)
start_ind=0
for idx in tqdm_notebook(range(int(start_ind/trials) ,5)):
scores = []
train_scores = []
accuracies = []
networks = []
# Shuffle testsets
inputFiles = files[:idx] + files[idx+1:]
testFiles = files[idx:idx+1]
for t in tqdm_notebook(range(int(start_ind%trials),trials)):
print('Exp id:', idx, t)
score, mse, accuracy, cm, train_score, train_mse, train_accuracy, lstm = \
trainLSTM(inputFiles=inputFiles, testFiles=testFiles, plot=False)
all_scores[idx,t] = score
all_train_scores[idx,t] = train_score
all_accuracies[idx,t] = accuracy
all_train_accuracies[idx,t] = train_accuracy
all_cms[idx,t] = cm[0]
#np.save(file=getProjectPath()+"lstm_scores", arr=np.array(all_scores))
#np.save(file=getProjectPath()+"lstm_train_scores", arr=np.array(all_train_scores))
#np.save(file=getProjectPath()+"lstm_cms", arr=np.array(all_cms))
#np.save(file=getProjectPath()+"lstm_accuracies", arr=np.array(all_accuracies))
#np.save(file=getProjectPath()+"lstm_train_accuracies", arr=np.array(all_train_accuracies))
# print results
import pandas as pd
pd.DataFrame(np.vstack([
all_accuracies.mean(1),
all_accuracies.std(1),
all_train_accuracies.mean(1),
all_train_accuracies.std(1),
all_scores.mean(1),
all_scores.std(1),
all_train_scores.mean(1),
all_train_scores.std(1),
]),
columns=files,
index=['accuracies','accuracies_std','accuracies_train','accuracies_train_std','f1','f1_std','f1_train','f1_train_std'])
all_scores.mean(), all_scores.std(), all_accuracies.mean(), all_accuracies.std(), all_train_scores.mean(), all_train_scores.std(),
```
## Additional methods and figures
```
score, mse, accuracy, cm, train_score, train_mse, train_accuracy, lstm = \
trainLSTM(inputFiles=inputFiles, testFiles=testFiles, plot=True)
#testLSTM(testloader,lstm)
trainset, validationset, testset, trainloader, validationloader, testloader = createData(inputFiles, testFiles)
score, mse, accuracy, cm = testLSTM(testloader, lstm, plot=True, plotConf=True)
# plot median confustion matrices
from Evaluation import plot_confusion_matrix
median_cms = []
for i, (testfile, scores, cms) in enumerate(zip(files, all_scores, all_cms)):
print(i, testfile)
# get median model
median_idx = np.argsort(scores)[round(len(scores)/2)]
median_idx.argsort
median_score = scores[median_idx]
median_cm = cms[median_idx]
median_cms.append(median_cm)
fig = plot_confusion_matrix(median_cm.astype('int'), gestures=totalGestureNames[:10] + totalGestureNames[-1:])
fig.tight_layout(pad=3.0)
pp = PdfPages('figures/lstm_experiment_{}_f1_score_{:.2f}.pdf'.format(testfile, median_score))
pp.savefig()
pp.close()
plt.savefig('figures/lstm_experiment_{}_f1_score_{:.2f}.eps'.format(testfile, median_score), format='eps')
# plot average confusion matrices
from Evaluation import plot_confusion_matrix
median_cms = []
for i, (testfile, scores, cms) in enumerate(zip(files, all_scores, all_cms)):
print(i, testfile)
# get median model
median_idx = np.argsort(scores)[round(len(scores)/2)]
median_idx.argsort
median_score = scores[median_idx]
median_cm = cms[median_idx]
median_cms.append(median_cm)
fig = plot_confusion_matrix(cms.mean(0), gestures=totalGestureNames[:10] + totalGestureNames[-1:])
fig.tight_layout(pad=3.0)
pp = PdfPages('figures/lstm_experiment_{}_avg__f1_score_{:.2f}.pdf'.format(testfile, scores.mean()))
pp.savefig()
pp.close()
plt.savefig('figures/lstm_experiment_{}_avg_f1_score_{:.2f}.eps'.format(testfile, scores.mean()), format='eps')
inputFiles = [ 'j', 'na', 'l', 'ni']
testFiles = ['s']
trainset, validationset, testset, trainloader, validationloader, testloader = createData(inputFiles, testFiles)
plt.figure(figsize=(15,5))
plt.plot(testset[0][1][:,9])
# create activations plot
import matplotlib
old_font = matplotlib.rcParams.get('font.size')
matplotlib.rcParams.update({'font.size': 20})
for inputs, targets in testloader:
test_inputs, test_targets = inputs.float(), targets.float()
#inputs[0,:,:9] = targets[0,:,:9]
test_inputs = (test_inputs/test_inputs.std(1))
test_inputs = test_inputs.cuda()
test_targets = test_targets.cuda()
lstm.eval()
outputs = lstm(test_inputs)
loss = loss_function(outputs, test_targets)
#losses.append(loss.item())
test_preds = outputs.cpu().detach().numpy()[0]
if True:
plt.figure(figsize=(15,6))
for signal, target, name in zip(test_preds.T,test_targets[0].T, gestureNames):
plt.plot(signal[0:480], label=name)
plt.fill_between(np.arange(480), np.zeros(480), target[0:480,].cpu(), alpha=0.05)
plt.plot(test_targets[0,0:480,:].cpu())
plt.xlim(0,700)
#plt.pause(1)
plt.legend()
plt.xlabel('Timesteps')
plt.ylabel('Activation')
fixed_threshold = 0.4
t_target = test_targets[0].cpu().numpy()
prediction = test_preds[:,:10]
if learnTreshold: # if threshold is learned, then it's the las collumn of the prediction
threshold = outputs[0].numpy()[:,10]
else: #else add a constant threshold
threshold = np.ones((prediction.shape[0],1))*fixed_threshold
t_maxApp_prediction = Evaluation.calcMaxActivityPrediction(prediction,t_target,threshold, 10)
pred_MaxApp, targ_MaxApp = Evaluation.calcInputSegmentSeries(t_maxApp_prediction, t_target, 0.5)
#testF1MaxApps.append(np.mean(sklearn.metrics.f1_score(targ_MaxApp,pred_MaxApp,average=None)))
#testAccuracies.append(np.mean(sklearn.metrics.accuracy_score(targ_MaxApp,pred_MaxApp)))
if False:
#print(t_maxApp_prediction.shape, prediction.shape, pred_MaxApp, targ_MaxApp)
plt.figure(figsize=(20,3))
plt.plot(t_maxApp_prediction[:800])
plt.plot(t_target[:800])
conf = sklearn.metrics.confusion_matrix(targ_MaxApp, pred_MaxApp)
#testCms.append(conf)
matplotlib.rcParams.update({'font.size': old_font})
pp = PdfPages('figures/subgesture.pdf'.format(testfile, scores.mean()))
pp.savefig()
pp.close()
plt.savefig('figures/subgesture.eps'.format(testfile, scores.mean()), format='eps')
plt.plot(targets[0])
(targets[0,:-1] < targets[0,1:]).sum(0)
```
| github_jupyter |
```
"""Collects data for file_properties_df.
file_properties_df holds metadata about data in our dataset,
details of metadata can be seen at src/nna/tests/mock_data.py::mock_file_properties_df
"""
from pathlib import Path
import pandas as pd
import datetime
from collections import Counter
from nna.fileUtils import list_files,getLength,read_file_properties_v2
# PARAMETERS
# increase version number accordinly
new_database_ver_str = 'V4'
# where to save txt file storing length info
old_data_folder="/home/enis/projects/nna/data/"
data_folder = '/scratch/enis/data/nna/database/'
#/scratch/enis/data/nna/database
# path to search for audio files
# where
# ignore_folders=['/tank/data/nna/real/stinchcomb/']
search_path="/tank/data/nna/real/"
ignore_folders=["/tank/data/nna/real/stinchcomb/dups/","/tank/data/nna/real/stinchcomb/excerpts/"]
# search_path="/tank/data/nna/real/stinchcomb/"
# if we already have a list of files we can load them
# files_list_path=data_folder+"stinchcomb_files_pathV1.txt"
files_list_path=data_folder+ f"allFields_path{new_database_ver_str}.txt"
# if we calculated audio lengths and saved them into text file,
# we can load them
fileswlen_path = data_folder+ f"allFields_wlen_f{new_database_ver_str}.txt"
filesWError = data_folder+f"allFields_wERROR_f{new_database_ver_str}.txt"
# do NOT add pkl at the end
pkl_file_name=f"allFields_dataf{new_database_ver_str}"
# this is the current info we have so we can check if we already processed a file before
current_pkl_file = old_data_folder + "allFields_dataV3.pkl"
%%time
# Find files
# in given search path ignoring given directories
if not Path(fileswlen_path).exists():
files_path_list=list_files(search_path,ignore_folders)
else:
with open(files_list_path,"r") as f:
lines=f.readlines()
files_path_list=[line.strip() for line in lines]
print('example file',files_path_list[0])
# count file extension and filter if required
files_suffixes=[]
files_path_list_filtered=[]
for m in files_path_list:
m=Path(m)
mSuffix = m.suffix.lower()
files_suffixes.append(mSuffix)
if "~" in str(m):
continue
# if mSuffix!=".flac" and mSuffix!=".aac" and mSuffix!=".mp3":
# print(m)
# break
files_path_list_filtered.append((m))
print(Counter(files_suffixes))
files_path_list = files_path_list_filtered[:]
#
# Load previous data
current_file_properties_df=pd.read_pickle(str(current_pkl_file))
# remove files we already know about
currentFileSet = set(current_file_properties_df.index)
foundFileSet = set(files_path_list)
foundFileSet = foundFileSet.difference(currentFileSet)
New_files_path_list = list(foundFileSet)
"new",len(New_files_path_list),"previously",len(currentFileSet),"total",len(files_path_list),
# New_files_path_list
# Load or calculate Audio length
import subprocess
filesWError = []
# learn length of each audio and store in a text file,
# if file already exists, it tries to get data from there
if not Path(fileswlen_path).exists():
length_dict={}
for f in New_files_path_list:
# length=float(getLength(f))
##################
input_video = f
ffprobe_path = '/scratch/enis/conda/envs/speechEnv/bin/ffprobe'
cmd=[]
cmd.extend( [ffprobe_path, '-i', '{}'.format(input_video), '-show_entries' ,'format=duration', '-v', 'quiet' ])
result = subprocess.Popen(cmd, stdout=subprocess.PIPE,stderr=subprocess.PIPE,)
output = result.communicate(b'\n')
output = [i.decode('ascii') for i in output]
if output[0]=="":
length = -1
print("ERROR file is too short {}".format(input_video))
print("command run with ERROR: {}".format(cmd))
filesWError.append(input_video)
else:
length = output[0].split("\n")[1].split("=")[1]
###############
length_dict[f]=length
length_list=list(length_dict.items())
with open(fileswlen_path,"w") as f:
for line in length_list:
f.write(",".join([line[0],str(line[1])])+"\n")
with open(fileswlen_path,"r") as f:
lines=f.readlines()
fileswlen=[line.strip().split(",") for line in lines]
# print and save files with errors
print(len(filesWError))
with open(filesWError_path,"w") as f:
for line in length_list:
f.write(",".join([line[0],str(line[1])])+"\n")
# turn results into a dict
fileswlen=dict([(i[0],float(i[1])) for i in fileswlen])
# file_properties
# file_properties,exceptions = read_file_properties_v2(New_files_path_list,debug=0)
for f,lengthSeconds in fileswlen.items():
if file_properties.get(Path(f)) is not None:
file_properties[Path(f)]["durationSec"] = lengthSeconds
file_properties[Path(f)]["timestampEnd"] = file_properties[Path(f)]["timestamp"] + datetime.timedelta(seconds=lengthSeconds)
file_properties_df=pd.DataFrame(file_properties).T
# exceptions
def str2timestamp(fileinfo_dict):
# x=file_properties[file]
# print(x)
hour_min_sec=fileinfo_dict["hour_min_sec"]
hour=int(hour_min_sec[:2])
minute=int(hour_min_sec[2:4])
second=int(hour_min_sec[4:6])
year = int(fileinfo_dict["year"])
timestamp=datetime.datetime(year, int(fileinfo_dict["month"]), int(fileinfo_dict["day"]),
hour=hour, minute=minute, second=second, microsecond=0)
fileinfo_dict["timestamp"]=timestamp
return fileinfo_dict
# merge with previous file properties
merged_file_properties_df = pd.concat([file_properties_df,current_file_properties_df])
merged_file_properties_df.to_pickle(data_folder+pkl_file_name+".pkl")
```
### Keep only prudhoe and anwr filter others
##### since they are only ones with images
```
# file_properties_df = pd.read_pickle(str(file_properties_df_FilePath))
len(merged_file_properties_df)
def mask2(df, key, value):
return df[df[key] == value]
pd.DataFrame.mask2 = mask2
prudhoe = merged_file_properties_df.mask2("region",'prudhoe')
anwr = merged_file_properties_df.mask2("region",'anwr')
len(prudhoe),len(anwr)
prudhoeAndAnwr4photoExp = pd.concat([prudhoe,anwr])
len(prudhoeAndAnwr4photoExp)
prudhoeAndAnwr4photoExp.to_pickle(data_folder+"prudhoeAndAnwr4photoExp_dataV1"+".pkl")
(data_folder+"prudhoeAndAnwr4photoExp_dataV1"+".pkl")
```
| github_jupyter |
# Generate parameterized datapackage
```
import bw2data as bd
import bw2io as bi
import bw2calc as bc
from pprint import pprint
from tqdm import tqdm
import bw2parameters as bwp
import numpy as np
import traceback
import sys
import re
from gsa_framework.utils import write_pickle, read_pickle
sys.path.append('/Users/akim/PycharmProjects/akula')
from akula.markets import DATA_DIR
assert bi.__version__ >= (0, 9, "DEV7")
from asteval import Interpreter
from numbers import Number
from bw2parameters.errors import BroadcastingError
from stats_arrays import uncertainty_choices
MC_ERROR_TEXT = """Formula returned array of wrong shape:
Name: {}
Formula: {}
Expected shape: {}
Returned shape: {}"""
class PatchedParameterSet(bwp.ParameterSet):
def evaluate_monte_carlo(self, iterations=1000):
"""Evaluate each formula using Monte Carlo and variable uncertainty data, if present.
Formulas **must** return a one-dimensional array, or ``BroadcastingError`` is raised.
Returns dictionary of ``{parameter name: numpy array}``."""
interpreter = Interpreter()
result = {}
def get_rng_sample(obj):
if isinstance(obj, np.ndarray):
# Already a Monte Carlo sample
return obj
if 'uncertainty_type' not in obj:
if 'uncertainty type' not in obj:
obj = obj.copy()
obj['uncertainty_type'] = 0
obj['loc'] = obj['amount']
else:
obj['uncertainty_type'] = obj['uncertainty type']
kls = uncertainty_choices[obj['uncertainty_type']]
return kls.bounded_random_variables(kls.from_dicts(obj), iterations).ravel()
def fix_shape(array):
# This is new
if array is None:
return np.zeros((iterations,))
elif isinstance(array, Number):
return np.ones((iterations,)) * array
elif not isinstance(array, np.ndarray):
return np.zeros((iterations,))
# End new section
elif array.shape in {(1, iterations), (iterations, 1)}:
return array.reshape((iterations,))
else:
return array
for key in self.order:
if key in self.global_params:
interpreter.symtable[key] = result[key] = get_rng_sample(self.global_params[key])
elif self.params[key].get('formula'):
sample = fix_shape(interpreter(self.params[key]['formula']))
if sample.shape != (iterations,):
raise BroadcastingError(MC_ERROR_TEXT.format(
key, self.params[key]['formula'], (iterations,), sample.shape)
)
interpreter.symtable[key] = result[key] = sample
else:
interpreter.symtable[key] = result[key] = get_rng_sample(self.params[key])
return result
bd.projects.set_current('GSA for archetypes')
bd.databases
ei = bd.Database("ecoinvent 3.8 cutoff")
# Takes forever and not necessary, skip it...
if not ei.metadata.get('fixed chemical formula name') and False:
from bw2data.backends.schema import ExchangeDataset as ED
qs = ED.select().where(ED.output_database == "ecoinvent 3.8 cutoff")
print("this will take a while, maybe 30 minutes")
for exc in tqdm(qs, total=629959):
if 'formula' in exc.data:
exc.data['chemical formula'] = exc.data.pop('formula')
exc.save()
ei.metadata['fixed chemical formula name'] = True
bd.databases.flush()
# fp_ecoinvent_38 = "/Users/cmutel/Documents/lca/Ecoinvent/3.8/cutoff/datasets"
fp_ecoinvent_38 = "/Users/akim/Documents/LCA_files/ecoinvent_38_cutoff/datasets"
fp_ei = DATA_DIR / "ecoinvent.pickle"
if fp_ei.exists():
eii = read_pickle(fp_ei)
else:
eii = bi.SingleOutputEcospold2Importer(fp_ecoinvent_38, "ecoinvent 3.8 cutoff")
eii.apply_strategies()
write_pickle(eii, fp_ei)
found = set()
for act in eii.data:
if any(exc.get('formula') for exc in act['exchanges']):
found.add(
(sum(1 for exc in act['exchanges'] if exc.get('formula')),
act['name'],
act['reference product'],
act['location'],
act['unit'])
)
len(found)
# list(sorted(found, reverse=True))[:25]
```
Don't trust pedigree uncertainty increases for variables
```
def drop_pedigree_uncertainty(dct):
if 'scale' in dct and 'scale with pedigree' in dct:
dct['scale with pedigree'] = dct.pop('scale')
dct['scale'] = dct.pop('scale without pedigree')
return dct
```
Change `10,43` to `10.42`.
```
test = "0,034 * 10,42"
result = re.sub(r'(\d)\,(\d)', r'\1.\2', test)
assert result == '0.034 * 10.42'
```
Fix Python reserved words used as variable names
```
substitutions = {
'yield': 'yield_',
'import': 'import_',
'load': 'load_',
}
```
Apply above fixes and a few others
```
def clean_formula(string):
string = string.strip().replace("%", " / 100").replace("^", " ** ").replace("\r\n", " ").replace("\n", "")
for k, v in substitutions.items():
string = string.replace(k, v)
string = re.sub(r'(\d)\,(\d)', r'\1.\2', string)
return string
def clean_dct(dct):
if dct.get('formula'):
dct['formula'] = clean_formula(dct['formula'])
if dct.get('name') in substitutions:
dct['name'] = substitutions[dct['name']]
return dct
def reformat_parameters(act):
parameters = {
substitutions.get(dct['name'], dct['name']): clean_dct(drop_pedigree_uncertainty(dct))
for dct in act['parameters'] if 'name' in dct
}
for index, exc in enumerate(act['exchanges']):
if exc.get('formula'):
pn = f'__exchange_{index}'
exc['parameter_name'] = pn
parameters[pn] = {'formula': clean_formula(exc['formula'])}
return parameters
def stochastic_parameter_set_for_activity(act, iterations=250):
ps = PatchedParameterSet(reformat_parameters(act))
return ps.evaluate_monte_carlo(iterations=iterations)
def check_that_parameters_are_reasonable(act, results, rtol=0.1):
for exc in act['exchanges']:
if exc.get('formula'):
arr = results[exc['parameter_name']]
if not np.isclose(exc['amount'], np.median(arr), rtol=rtol):
print(
act['name'],
exc['name'],
act['location'],
act['unit'],
)
print("\t", exc['amount'], np.median(arr), exc['formula'])
return False
return True
from bw2data.backends.schema import ActivityDataset as AD
lookup_cache = {(x, y): z
for x, y, z in AD.select(AD.database, AD.code, AD.id)
.where(AD.database << ("biosphere3", "ecoinvent 3.8 cutoff"))
.tuples()
}
tech_data, bio_data = [], []
found, errors, unreasonable, missing = 0, 0, 0, 0
error_log = open("error.log", "w")
missing_reference_log = open("undefined_reference.log", "w")
for act in tqdm(eii.data):
if any(exc.get('formula') for exc in act['exchanges']):
try:
params = stochastic_parameter_set_for_activity(act, iterations=25000)
break
if check_that_parameters_are_reasonable(act, params):
found += 1
for exc in act['exchanges']:
if not exc.get('formula'):
continue
if exc['input'][0] == "ecoinvent 3.8 cutoff":
tech_data.append((
(lookup_cache[exc['input']], lookup_cache[(act['database'], act['code'])]),
params[exc['parameter_name']],
exc['type'] != 'production' # TODO Chris please check, changed this from == to !=
))
else:
bio_data.append((
(lookup_cache[exc['input']], lookup_cache[(act['database'], act['code'])]),
params[exc['parameter_name']],
False
))
else:
unreasonable += 1
except (ValueError, SyntaxError, bwp.errors.DuplicateName):
error_log.write(act['filename'] + "\n")
traceback.print_exc(file=error_log)
errors += 1
except bwp.errors.ParameterError:
missing_reference_log.write(act['filename'] + "\n")
traceback.print_exc(file=missing_reference_log)
missing += 1
error_log.close()
missing_reference_log.close()
found, errors, unreasonable, missing
len(tech_data), len(bio_data)
```
Uncertain production exchanges. Could be bad data.
```
np.hstack([z for x, y, z in tech_data]).sum()
import bw_processing as bp
from fs.zipfs import ZipFS
dp = bp.create_datapackage(
fs=ZipFS("ecoinvent-parameterization.zip", write=True),
name="ecoinvent-parameterization",
seed=42,
)
indices = np.empty(len(tech_data), dtype=bp.INDICES_DTYPE)
indices[:] = [x for x, y, z in tech_data]
dp.add_persistent_array(
matrix="technosphere_matrix",
data_array=np.vstack([y for x, y, z in tech_data]),
name="ecoinvent-parameterization-tech",
indices_array=indices,
flip_array=np.hstack([z for x, y, z in tech_data]),
)
indices = np.empty(len(bio_data), dtype=bp.INDICES_DTYPE)
indices[:] = [x for x, y, z in bio_data]
dp.add_persistent_array(
matrix="biosphere_matrix",
data_array=np.vstack([y for x, y, z in bio_data]),
name="ecoinvent-parameterization-bio",
indices_array=indices,
flip_array=np.hstack([z for x, y, z in bio_data]),
)
dp.finalize_serialization()
```
# [archived] Check values in the parameterized datapackage
```
from pathlib import Path
import numpy as np
from fs.zipfs import ZipFS
import bw2calc as bc
import bw2data as bd
import bw_processing as bwp
import sys
sys.path.append('/Users/akim/PycharmProjects/akula')
from akula.virtual_markets import DATA_DIR
fp_ei_parameterization = DATA_DIR / "ecoinvent-parameterization.zip"
dp_params = bwp.load_datapackage(ZipFS(fp_ei_parameterization))
project = "GSA for archetypes"
bd.projects.set_current(project)
method = ("IPCC 2013", "climate change", "GWP 100a", "uncertain")
me = bd.Method(method)
bs = bd.Database("biosphere3")
ei = bd.Database("ecoinvent 3.8 cutoff")
co_name = "swiss consumption 1.0"
co = bd.Database(co_name)
list_ = [me, bs, ei, co]
dps = [
bwp.load_datapackage(ZipFS(db.filepath_processed()))
for db in list_
]
hh_average = [act for act in co if "ch hh average consumption aggregated" == act['name']]
assert len(hh_average) == 1
demand_act = hh_average[0]
demand = {demand_act: 1}
demand_id = {demand_act.id: 1}
iterations = 5
lca = bc.LCA(
demand_id,
data_objs=dps,
use_distributions=True,
use_arrays=True,
seed_override=11111000
)
lca.lci()
lca.lcia()
scores = [lca.score for _, _ in zip(lca, range(iterations))]
scores
lca_params = bc.LCA(
demand_id,
data_objs=dps + [dp_params],
use_distributions=True,
use_arrays=True,
seed_override=11111000,
)
lca_params.lci()
lca_params.lcia()
scores_params = [lca_params.score for _, _ in zip(lca_params, range(iterations))]
scores_params
dp_params_bio = dp_params.filter_by_attribute("group", "ecoinvent-parameterization-bio")
lca_params_bio = bc.LCA(
demand_id,
data_objs=dps + [dp_params_bio],
use_distributions=True,
use_arrays=True,
seed_override=11111000,
)
lca_params_bio.lci()
lca_params_bio.lcia()
scores_params_bio = [lca_params_bio.score for _, _ in zip(lca_params_bio, range(iterations))]
scores_params_bio
dp_params_tech = dp_params.filter_by_attribute("group", "ecoinvent-parameterization-tech")
lca_params_tech = bc.LCA(
demand_id,
data_objs=dps + [dp_params_tech],
use_distributions=True,
use_arrays=True,
seed_override=11111000,
)
lca_params_tech.lci()
lca_params_tech.lcia()
scores_params_tech = [lca_params_tech.score for _, _ in zip(lca_params_tech, range(iterations))]
scores_params_tech
dp_ei = bd.Database("ecoinvent 3.8 cutoff").datapackage()
ei_indices = dp_ei.get_resource("ecoinvent_3.8_cutoff_technosphere_matrix.indices")[0]
ei_data = dp_ei.get_resource("ecoinvent_3.8_cutoff_technosphere_matrix.data")[0]
ei_flip_raw = dp_ei.get_resource("ecoinvent_3.8_cutoff_technosphere_matrix.flip")[0]
ei_selected = []
ei_flip = []
for i, inds in enumerate(dp_params_tech.data[0]):
ei_where = np.where(ei_indices==inds)[0][0]
ei_selected.append(ei_data[ei_where])
ei_flip.append(ei_flip_raw[ei_where])
params_selected = dp_params_tech.data[1][:,0]
ei_selected = np.array(ei_selected)
ei_flip = np.array(ei_flip)
indices_selected = dp_params_tech.data[0]
wdiff = abs(params_selected - ei_selected)
# np.where(wdiff==min(wdiff))
%%time
res = bc.GraphTraversal().calculate(
lca, cutoff=1e-3, max_calc=1e3
)
%%time
res_params_tech = bc.GraphTraversal().calculate(
lca_params_tech, cutoff=1e-3, max_calc=1e3
)
import pandas as pd
df = pd.DataFrame.from_dict(res['edges'])
df_params = pd.DataFrame.from_dict(res_params_tech['edges'])
df_both = df.merge(df_params, on=['to', 'from'], how='outer')
df_both.to_excel("sct.xlsx")
lca_params_tech1 = bc.LCA(
{4916: 1212.188043},
data_objs=dps + [dp_params_tech],
use_distributions=False,
use_arrays=True,
seed_override=11111000,
)
lca_params_tech1.lci()
lca_params_tech1.lcia()
lca_params_tech1.score
lca_params_tech1 = bc.LCA(
{4916: 1212.188043},
data_objs=dps,
use_distributions=False,
use_arrays=True,
seed_override=11111000,
)
lca_params_tech1.lci()
lca_params_tech1.lcia()
lca_params_tech1.score
params_flip = dp_params_tech.get_resource('ecoinvent-parameterization-tech.flip')[0]
sum(ei_flip), sum(params_flip)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/cltl/python-for-text-analysis/blob/colab/Assignments-colab/ASSIGNMENT_RESIT_B.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
%%capture
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Data.zip
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/images.zip
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Extra_Material.zip
!unzip Data.zip -d ../
!unzip images.zip -d ./
!unzip Extra_Material.zip -d ../
!rm Data.zip
!rm Extra_Material.zip
!rm images.zip
```
# Assignment Resit - Part B
Deadline: Friday, November 13, 2020 before 17:00
This part of the assignment should be submitted as a zip file containing two python modules:
* utils.py
* texts_to_coll.py
* ASSIGNMENT-RESIT-A.ipynb (notebook containing part A)
Please name your zip file as follows: RESIT-ASSIGNMENT.zip and upload it via Canvas (Resit Assignment).
Please submit your assignment on Canvas: Resit Assignment
If you have questions about this topic
* If you have **questions** about this topic, please contact **cltl.python.course@gmail.com**.
Questions and answers will be collected in this [Q&A document](https://docs.google.com/document/d/1Yf2lE6HdApz4wSgNpxWL_nnVcXED1YNW8Rg__wCKcvs/edit?usp=sharing), so please check if your question has already been answered.
All of the covered chapters are important to this assignment. However, please pay special attention to:
* Chapter 14 - Reading and writing text files
* Chapter 15 - Off to analyzing text
* Chapter 16 - Data formats I (CSV and TSV)
* Chapter 19 - More about Natural Language Processing Tools (spaCy)
In this assignment, we are going to write code which conversts raw text to a structured format frequently used in Natural Lanugage Processing. No matter what field you will end up working in, you will always have to be able to convert data from format A to format B. You have already gained some experience with such conversions in Block 4.
**The CoNLL format**
Before you use the output of a text analysis system, you usually want to store the output in a structured format. One way of doing this is to use naf - a format using xml. In this assignment, we are going to look at CoNLL, which is a table-based format (i.e. it is similar to csv/tsv).
The format we are converting to is called CoNLL. CoNLL is the name of a conference (Conference on Natural Language Learning). Every year, the conference hosts a 'competion'. In this competition, participants have to build systems for a certain Natural Language Processing problem (usually referred to as 'task'). To compare results, participants have to stick to the CoNLL format. The format has become a popular format for storing the output of NLP systems.
The goal of this assignment is to write a python module which processes all texts in ../Data/Dreams/. The output should be written to a new directory, in which each text is stored as a csv/tsv file following CoNLL conventions.
**Text analysis with SpaCy**
In part A of this assignment, you have already used SpaCy to process text. In this part of the assignment, you can make use of the code you have already written. The output files will contain the following information:
* The tokens in each text
* Information about the sentences in each text
* Part-of-speech tags for each token
* The lemma of each token
* information about entities in a text (i.e. people, places, organizations, etc that are mentioned)
**The assignment**
We will guide you towards the final file-conversion step-by-step. The assignment is divided in 3 parts. We provide small toy exampls you can use to develop your code. As a final step, you will be asked to transfer all your code to python modules and process a directory of text files with it.
Exercise 1: A guided tour of the CoNLL format
Exercise 2: Writing a conversion function (text_to_conll)
Exercise 3: Processing multiple files using python modules
**Attention: This notebook should be placed in the same folder as the other Assignments!**
## 1. Understanding the CoNLL format
The CoNLL format represents information about a text in table format. Each token is represented on a line. Each column contains a piece of information. Sentence-boundaries are marked by empty lines. In addition, each token has an index. This index starts with 1 and indentifies the positoion of the token in the sentence. Punctuation marks are also included.
Consider the following example text:
*This is an example text. The text mentions a former president of the United States, Barack Obama.*
The representation of this sentence in CoNLL format looks like this:
| | | | | | |
|----|-----------|-----|-----------|--------|---|
| 1 | This | DT | this | | O |
| 2 | is | VBZ | be | | O |
| 3 | an | DT | an | | O |
| 4 | example | NN | example | | O |
| 5 | text | NN | text | | O |
| 6 | . | . | . | | O |
| | | | | | |
| 1 | The | DT | the | | O |
| 2 | text | NN | text | | O |
| 3 | mentions | VBZ | mention | | O |
| 4 | a | DT | a | | O |
| 5 | former | JJ | former | | O |
| 6 | president | NN | president | | O |
| 7 | of | IN | of | | O |
| 8 | the | DT | the | GPE | B |
| 9 | United | NNP | United | GPE | I |
| 10 | States | NNP | States | GPE | I |
| 11 | , | , | , | | O |
| 12 | Barack | NNP | Barack | PERSON | B |
| 13 | Obama | NNP | Obama | PERSON | I |
| 14 | . | . | . | | O |
**The columns represent the following information:**
* Column 1: Token index in sentence
* Column 2: The token as it appears in the text (including punctuation)
* Column 3: The part-of-speech tag
* Column 4: The lemma of the token
Column 5: Information about the type of entity (if the token is part of an expression referring to an entity). For example, Barack Obama is recognized as a person
Column 6: Information about the position of the token in the entiy-mention. B stands for 'beginning', I stands for 'inside' and O stands for 'outside'. Anything that is not part of an entity mention is marked as 'outside'. (This is important information for dealing with entity mentions. Don't worry, you do not have to make use of this information here.)
## 2. Writing the conversion function
In this section of the assignment, we will guide you through writing your function. You can accomplish the entire conversion in a single function (i.e. there will be no helper functions at this point). We will first describe what your function should do and then provide small toy examples to help you with some of the steps.
**The conversion function: text_to_conll**
(1) Define a function called text_to_conll
(2) The function should have the following parameters:
* text: The input text (str) that should be processed and written to a conll file
* nlp: the SpaCy model
* output_dir: the directory the file should be written to
* basename: the name of the output file without the path (i.e the file will be written to output_dir/basename
* delimiter: the field delimiter (by default, it should be a tab)
* start_with_index: By default, this should be True.
* overwrite_existing_conll_file: By default, this should be set to True.
(3) The function should do the following:
* Convert text to CoNll format as shown in the example in exercise 1.
* The file should have the following columns:
* Token index in sentence (as shown in example) If start_with_index is set to False, the first column should be the token.
* token
* part of speech tag (see tips below)
* lemma
* entity type (see tips below)
* entity iob label (indicates the position of a token in an entity-expression (see tips below)
* If the parameter overwrite_existing_conll_file is set to True, the file should be written to output_dir/basename.
* If the parameter overwrite_existing_conll_file is set to False, the function should check whether the file (path: output_dir/basename) exists. If it does, it should print 'File exists. Set param overwrite_exisiting_conll_file to True if you want to overwrite it.' If it does not exist, it should write it to the specified file. (See tips below)
* The delimiter between fields should be the delimiter specified by the parameter delimiter.
You can define the function in the notebook. Please test it using the following test text. Make sure to test the different paprameters. Your test file should be written to `test_dir/test_text.tsv`.
```
# your function
# test your function
text = 'This is an example text. The text mentions a former president of the United States, Barack Obama.'
basename = 'test_text.tsv'
output_dir = 'test_dir'
text_to_conll_simple(text,
nlp,
output_dir,
basename,
start_with_index = False,
overwrite_existing_conll_file = True)
```
## Tip 0: Import spacy and load your model
(See part A and chapter on SpaCy for more information)
```
import spacy
nlp = spacy.load('en_core_web_sm')
```
### Tip 1: Tokens, POS tags, and lemmas
Experiment with a small example to get the tokens and pos tags. Please refer to the chapter on SpaCy for an example on how to process text with spacy.
Spacy has different pos tags. For this exercise, it does not matter which one you use. Hint: To get a string (rather than a number, use the SpaCy attributes ending with '_').
You can use the code below to experiment:
```
test = 'This is a test.'
doc = nlp(test)
tok = doc[0]
tok.text
```
### Tip 2: Entities
**Entity types**
Entities are things (usually people/places/organizations/etc) that exist in the real world. SpaCy can tag texts with entity types. If an expression refers to an entity in the world, it will receive a lable indicating the type (for example, Barack Obama will be tagged as 'PERSON'. Since the expression 'Barack Obama' consists of two tokens, each token will receive such a label. Use dir() on a token object to find out how to get this information. Hint: **Everything about entities starts with ('ent_')**
**Position of the entity token**
An expression referring to an entity can consist of multiple tokens. To indicate that multiple tokens are part of the same/of different expressions, we often use the IOB system. In this system, we indicate whether a token is outside an entity mention, inside an entity mention or at the beginning of an entity mention. In practice, most tokens of a text will thus be tagged as 'O'. 'Barack' will be tagged as 'B' and 'Obama' as 'I' (see example above). SpaCy can do this type of labeling. Use dir() on a token object to find out how to get this information.
```
test = 'This is a test.'
doc = nlp(test)
tok = doc[0]
tok.text
dir(tok)
```
### Tip 3: Dealing with directories and files
Use os to check if files or directories exist. You can also use os to make a directory if it does not exist yet.
* os.path.isdir(path_to_dir) returns a boolean value. If the directory exists, it returns True. Else it returns False. You can use this to check if a directory exists. If it does not, you can make it.
* os.path.isfile(path_to_file) returns a boolean value. If the file exists, it returns True. Else it returns False.
* os.mkdir(path_to_dir) makes a new directory. Try it out and create a directory called 'test_dir' in the current directory.
```
# Check if file exists
import os
a_path_to_a_file = '../Data/books/Macbeth.txt'
if os.path.isfile(a_path_to_a_file):
print('File exists:', a_path_to_a_file)
else:
print('File not found:', a_path_to_a_file)
another_path_to_a_file = '../Data/books/KingLear.txt'
if os.path.isfile(another_path_to_a_file):
print('File exists:', another_path_to_a_file)
else:
print('File not found:', another_path_to_a_file)
# check if directory exists
a_path_to_a_dir = '../Data/books/'
if os.path.isdir(a_path_to_a_dir):
print('Directory exists:', a_path_to_a_dir)
else:
print('Directory not found:', a_path_to_a_dir)
another_path_to_a_dir = '../Data/films/'
if os.path.isdir(another_path_to_a_dir):
print('Directory exists:', another_path_to_a_dir)
else:
print('Directory not found:', another_path_to_a_dir)
```
## 3. Building python modules to process files in a directory
In this exercise, you will write two python modules:
* utils.py
* texts_to_conll.py
The module texts_to_conll.py should do the following:
* process all text files in a specified directory (we will use '../Data/Dreams')
* write conll files representing these texts to another directory
**Step 1: Preparation**:
* Create the two python modules in the same directory as this notebook
* copy your function `text_to_conll` to the python module `texts_to_conll.py`
* Move the function `load_text` you have defined in part A to `utils.py` and import it in `text_to_conll.py`
* Move the function `get_paths` you have defined in part A to `utils.py` and import in it `text_to_conll.py`
**Step 2: convert all text files in ../Data/Dreams**:
Use your functions to convert all files in `../Data/Dreams/`. Please fulfill the following criteria:
* The new files should be placed in a directory placed in the current directory called dreams_conll/
* Each file should be named as follows: [original name without extension].tsv (e.g.vicky1.tsv)
* The files should contain an index column
Tips:
* Use a loop to iterate over the files in ../Data/Dreams.
* Use string methods and slicing to create the new filename from the original filename (e.g. split on '/' and/or '.', use indices to extract certain substrings, etc.)
* Look at the resulting files to check if your code works.
**Step 3: Test and submit**
Please test your code carefully. Them submitt all your files in a .zip file via Canvas.
**Congratulations! You have completed your first file conversion exercise!**
```
# Files in '../Data/Dreams':
%ls ../Data/Dreams/
```
| github_jupyter |
<a href="https://colab.research.google.com/github/bkkaggle/pytorch-CycleGAN-and-pix2pix/blob/master/pix2pix.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Install
```
# this mounts your Google Drive to the Colab VM.
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# enter the foldername in your Drive where you have saved the unzipped
# assignment folder, e.g. 'cs231n/assignments/assignment3/'
FOLDERNAME = 'CS231N/'
assert FOLDERNAME is not None, "[!] Enter the foldername."
# now that we've mounted your Drive, this ensures that
# the Python interpreter of the Colab VM can load
# python files from within it.
import sys
sys.path.append('/content/drive/My Drive/{}'.format(FOLDERNAME))
#Imports
import shutil
import os
import matplotlib.pyplot as plt
#Do all of the required pix2pix stuff
!git clone https://github.com/michaele77/CS231N-pytorch
import os
# os.chdir('pytorch-CycleGAN-and-pix2pix/')
os.chdir('CS231N-pytorch/')
!pip install -r requirements.txt
#Run the combination script
!python datasets/combine_A_and_B.py --fold_A path/to/data/A --fold_B path/to/data/B --fold_AB path/to/data
#Test directories
!ls
####THIS IS THE MAIN TRAINING LOOP####
#First, let's train for 70 epochs, 35 main and 35 decay
#Everything default, print every 100
#Use the WGAN loss model
import shutil
import os
# !python train.py --dataroot ./path/to/data/ --name test0_WGAN_1 --model pix2pix --batch_size=8 --direction AtoB --n_epochs=25 --n_epochs_decay=25 --print_freq=100 --gan_mode=wgangp
# dirNum = 2
# lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
# shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + lossFileSrc)
# fileSrc = 'test0_WGAN_1'
# shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc)
#Now, let's train for 70 epochs, 35 main and 35 decay
#Everything default, print every 100
#ITER 0
# !python train.py --dataroot ./path/to/data/ --name test0_nom --model pix2pix --batch_size=8 --direction AtoB --n_epochs=25 --n_epochs_decay=25 --print_freq=100 --gan_mode=lsgan --continue_train --epoch_count=28
# fileSrc = 'test0_nom'
# modifier = '_iter1'
# flag = True
# initDirNum = 10
# dirCntr = 0
# dirNum = initDirNum - dirCntr
# while flag and dirNum >=0:
# try:
# c = dirNum
# lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
# shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + lossFileSrc)
# flag = False
# except:
# print('Directory ' + str(dirNum) + ' not occupied, moving down')
# dirCntr += 1
# dirNum = initDirNum - dirCntr
# shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc+modifier)
# #ITER 0
# !python train.py --dataroot ./path/to/data/ --name test0_nom --model pix2pix --batch_size=8 --direction AtoB --n_epochs=25 --n_epochs_decay=25 --print_freq=100 --gan_mode=lsgan --continue_train --epoch_count=28
# fileSrc = 'test0_nom_iter0'
# flag = True
# initDirNum = 10
# dirCntr = 0
# dirNum = initDirNum - dirCntr
# while flag and dirNum >=0:
# try:
# c = dirNum
# lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
# shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + lossFileSrc)
# flag = False
# except:
# print('Directory ' + str(dirNum) + ' not occupied, moving down')
# dirCntr += 1
# dirNum = initDirNum - dirCntr
# shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc)
# #ITER 1, theoretically start from 38
# !python train.py --dataroot ./path/to/data/ --name test0_nom --model pix2pix --batch_size=8 --direction AtoB --n_epochs=25 --n_epochs_decay=25 --print_freq=100 --gan_mode=lsgan --continue_train --epoch_count=28
# fileSrc = 'test0_nom_iter1'
# flag = True
# initDirNum = 10
# dirCntr = 0
# dirNum = initDirNum - dirCntr
# while flag and dirNum >=0:
# try:
# c = dirNum
# lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
# shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + lossFileSrc)
# flag = False
# except:
# print('Directory ' + str(dirNum) + ' not occupied, moving down')
# dirCntr += 1
# dirNum = initDirNum - dirCntr
# shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc)
# #ITER 0
# !python train.py --dataroot ./path/to/data/ --name test0_nom --model pix2pix --batch_size=8 --direction AtoB --n_epochs=25 --n_epochs_decay=25 --print_freq=100 --gan_mode=lsgan --continue_train --epoch_count=28
# fileSrc = 'test0_nom_iter0'
# flag = True
# initDirNum = 10
# dirCntr = 0
# dirNum = initDirNum - dirCntr
# while flag and dirNum >=0:
# try:
# c = dirNum
# lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
# shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + lossFileSrc)
# flag = False
# except:
# print('Directory ' + str(dirNum) + ' not occupied, moving down')
# dirCntr += 1
# dirNum = initDirNum - dirCntr
# shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc)
# #ITER 0
# !python train.py --dataroot ./path/to/data/ --name test0_nom --model pix2pix --batch_size=8 --direction AtoB --n_epochs=25 --n_epochs_decay=25 --print_freq=100 --gan_mode=lsgan --continue_train --epoch_count=28
# fileSrc = 'test0_nom_iter0'
# flag = True
# initDirNum = 10
# dirCntr = 0
# dirNum = initDirNum - dirCntr
# while flag and dirNum >=0:
# try:
# c = dirNum
# lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
# shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + lossFileSrc)
# flag = False
# except:
# print('Directory ' + str(dirNum) + ' not occupied, moving down')
# dirCntr += 1
# dirNum = initDirNum - dirCntr
# shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc)
#First, let's train for 70 epochs, 35 main and 35 decay
#Everything default, print every 100
#Use the WGAN loss model
#Train at a higher learning rate
!python train.py --dataroot ./path/to/data/ --name test0_WGAN_lorate --model pix2pix --batch_size=8 --direction AtoB --n_epochs=25 --n_epochs_decay=25 --print_freq=100 --gan_mode=wgangp --lr=0.001
fileSrc = 'test0_WGAN_lorate'
modifier = '_iter0'
flag = True
initDirNum = 10
dirCntr = 0
dirNum = initDirNum - dirCntr
shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc + modifier)
while flag and dirNum >=0:
try:
c = dirNum
lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + fileSrc + modifier + '/' + lossFileSrc)
flag = False
except:
print('Directory ' + str(dirNum) + ' not occupied, moving down')
dirCntr += 1
dirNum = initDirNum - dirCntr
# !python train.py --dataroot ./path/to/data/ --name test0_WGAN_hirate --model pix2pix --batch_size=8 --direction AtoB --n_epochs=12 --n_epochs_decay=12 --print_freq=100 --gan_mode=wgangp --lr=0.007 --continue_train --epoch_count=24
# fileSrc = 'test0_WGAN_hirate'
# modifier = '_iter1'
# flag = True
# initDirNum = 10
# dirCntr = 0
# dirNum = initDirNum - dirCntr
# while flag and dirNum >=0:
# try:
# c = dirNum
# lossFileSrc = 'ershov_lossFolder_' + str(dirNum)
# shutil.copytree(lossFileSrc, '/content/drive/My Drive/CS231N/' + lossFileSrc)
# flag = False
# except:
# print('Directory ' + str(dirNum) + ' not occupied, moving down')
# dirCntr += 1
# dirNum = initDirNum - dirCntr
# shutil.copytree('checkpoints/' + fileSrc, '/content/drive/My Drive/CS231N/' + fileSrc)
####THIS IS THE MAIN TRAINING LOOP####
#Define loading and plotting functions
def lossDataLoad(inStr):
G_GAN = []
G_loss = []
D_real = []
D_fake = []
with open(inStr + '/loss.txt', 'r') as filehandle:
#It's a list of lists of floats
fullFile = filehandle.readlines()
cntr = 0
for i in fullFile:
cntr += 1
if i == '-\n':
cntr = 0
continue
elif cntr == 1:
G_GAN.append(float(i[:-2]))
elif cntr == 2:
G_loss.append(float(i[:-2]))
elif cntr == 3:
D_real.append(float(i[:-2]))
elif cntr == 4:
D_fake.append(float(i[:-2]))
epoch = []
epoch_iter = []
with open(inStr + '/epoch.txt', 'r') as filehandle:
#It's a list of 2 numbers for epoch
fullFile = filehandle.readlines()
cntr = 0
for i in fullFile:
if i == '-\n':
cntr = 0
continue
else:
cntr += 1
if cntr == 1:
epoch.append(float(i[:-2]))
else:
epoch_iter.append(float(i[:-2]))
combEpoch = []
for i in range(len(epoch)):
bigE = epoch[i]
endE = epoch_iter[-1]
temp = (bigE - 1)*endE + epoch_iter[i]
combEpoch.append(temp)
return G_GAN, G_loss, D_real, D_fake, combEpoch
#plotting function
def lossPlotFunc(G_GAN, G_loss, D_real, D_fake, combEpoch):
fig, axs = plt.subplots(1, 2, figsize=(15,5))
axs[0].plot(combEpoch, G_GAN)
axs[0].set_title('Generator Error', fontsize = 20)
axs[0].set_xlabel('Batch Number', fontsize = 20)
axs[0].set_ylabel('Error', fontsize = 20)
axs[0].grid(True)
axs[1].plot(combEpoch, G_loss)
axs[1].set_title('Generator Loss', fontsize = 20)
axs[1].set_xlabel('Batch Number', fontsize = 20)
axs[1].set_ylabel('Loss', fontsize = 20)
axs[1].grid(True)
plt.savefig('/content/drive/My Drive/CS231N/tempPlot1.jpg')
fig, axs = plt.subplots(1, 2, figsize=(15,5))
axs[0].plot(combEpoch, D_real)
axs[0].set_title('Discr Real Confidence', fontsize = 20)
axs[0].set_xlabel('Batch Number', fontsize = 20)
axs[0].set_ylabel('Confidence', fontsize = 20)
axs[0].grid(True)
axs[1].plot(combEpoch, D_fake)
axs[1].set_title('Discr Fake Confidence', fontsize = 20)
axs[1].set_xlabel('Batch Number', fontsize = 20)
axs[1].set_ylabel('Confidence', fontsize = 20)
axs[1].grid(True)
plt.savefig('/content/drive/My Drive/CS231N/tempPlot2.jpg')
#Now, let's plot this shit!
#First, create an epoch and log list
# ershStr = 'ershov_lossFolder_0'
# ershStr = 'test0_WGAN_lorate_iter0/ershov_lossFolder_5'
ershStr = 'test0_nom_iter1/ershov_lossFolder_2'
fullString = '/content/drive/My Drive/CS231N/' + ershStr
G_GAN, G_loss, D_real, D_fake, combEpoch = lossDataLoad(fullString)
print(G_loss)
lossPlotFunc(G_GAN, G_loss, D_real, D_fake, combEpoch)
!ls
#Now test
# ershStr = 'ershov_lossFolder_0'
# ershStr = 'test0_WGAN_lorate_iter0'
ershStr = 'test0_nom_iter1'
#First need to copy the file to the mounted checkpoints folder
shutil.copytree('/content/drive/My Drive/CS231N/' + ershStr, 'checkpoints/' + ershStr)
!ls checkpoints/test0_nom_iter1
!python test.py --dataroot ./path/to/data/ --name test0_nom_iter1 --model pix2pix --batch_size=8 --direction AtoB
!ls results/test0_nom_iter1/test_latest/
shutil.copytree('results/' + ershStr + '/test_latest/images', '/content/drive/My Drive/CS231N/' + ershStr + '/results/')
```
# Testing
- `python test.py --dataroot ./datasets/facades --direction BtoA --model pix2pix --name facades_pix2pix`
Change the `--dataroot`, `--name`, and `--direction` to be consistent with your trained model's configuration and how you want to transform images.
> from https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix:
> Note that we specified --direction BtoA as Facades dataset's A to B direction is photos to labels.
> If you would like to apply a pre-trained model to a collection of input images (rather than image pairs), please use --model test option. See ./scripts/test_single.sh for how to apply a model to Facade label maps (stored in the directory facades/testB).
> See a list of currently available models at ./scripts/download_pix2pix_model.sh
```
!ls checkpoints/greyGoal_1
!python test.py --dataroot ./path/to/data/ --name greyGoal_1 --model pix2pix --batch_size=8 --direction AtoB
!ls results/greyGoal_1/test_latest/images/
#Try and save some of the created images to the directory:
testDir = 'results/greyGoal_1/test_latest/images/25986_fake_B.png'
!ls
!ls ..
import cv2
a = cv2.imread(testDir)
print(a.shape)
cv2.imwrite('/content/drive/My Drive/CS231N/testIm2.jpg', a)
```
# Visualize
```
import matplotlib.pyplot as plt
img = plt.imread('results/greyGoal_1/test_latest/images/25986_fake_B.png')
plt.imshow(img)
img = plt.imread('results/greyGoal_1/test_latest/images/25986_real_A.png')
plt.imshow(img)
img = plt.imread('results/greyGoal_1/test_latest/images/25986_real_B.png')
plt.imshow(img)
```
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!# Network Graphs with Plotly.
Install the Python library `networkx` with `sudo pip install networkx`.
#### Get Node Positions
Store position as node attribute data for random_geometric_graph and find node near center (0.5, 0.5)
```
import plotly.plotly as py
import plotly.graph_objs as go
import networkx as nx
G=nx.random_geometric_graph(200,0.125)
pos=nx.get_node_attributes(G,'pos')
dmin=1
ncenter=0
for n in pos:
x,y=pos[n]
d=(x-0.5)**2+(y-0.5)**2
if d<dmin:
ncenter=n
dmin=d
p=nx.single_source_shortest_path_length(G,ncenter)
```
#### Create Edges
Add edges as disconnected lines in a single trace and nodes as a scatter trace
```
edge_trace = go.Scatter(
x=[],
y=[],
line=dict(width=0.5,color='#888'),
hoverinfo='none',
mode='lines')
for edge in G.edges():
x0, y0 = G.node[edge[0]]['pos']
x1, y1 = G.node[edge[1]]['pos']
edge_trace['x'] += tuple([x0, x1, None])
edge_trace['y'] += tuple([y0, y1, None])
node_trace = go.Scatter(
x=[],
y=[],
text=[],
mode='markers',
hoverinfo='text',
marker=dict(
showscale=True,
# colorscale options
#'Greys' | 'YlGnBu' | 'Greens' | 'YlOrRd' | 'Bluered' | 'RdBu' |
#'Reds' | 'Blues' | 'Picnic' | 'Rainbow' | 'Portland' | 'Jet' |
#'Hot' | 'Blackbody' | 'Earth' | 'Electric' | 'Viridis' |
colorscale='YlGnBu',
reversescale=True,
color=[],
size=10,
colorbar=dict(
thickness=15,
title='Node Connections',
xanchor='left',
titleside='right'
),
line=dict(width=2)))
for node in G.nodes():
x, y = G.node[node]['pos']
node_trace['x'] += tuple([x])
node_trace['y'] += tuple([y])
```
#### Color Node Points
Color node points by the number of connections.
Another option would be to size points by the number of connections
i.e. ```node_trace['marker']['size'].append(len(adjacencies))```
```
for node, adjacencies in enumerate(G.adjacency()):
node_trace['marker']['color']+=tuple([len(adjacencies[1])])
node_info = '# of connections: '+str(len(adjacencies[1]))
node_trace['text']+=tuple([node_info])
```
#### Create Network Graph
```
fig = go.Figure(data=[edge_trace, node_trace],
layout=go.Layout(
title='<br>Network graph made with Python',
titlefont=dict(size=16),
showlegend=False,
hovermode='closest',
margin=dict(b=20,l=5,r=5,t=40),
annotations=[ dict(
text="Python code: <a href='https://plot.ly/ipython-notebooks/network-graphs/'> https://plot.ly/ipython-notebooks/network-graphs/</a>",
showarrow=False,
xref="paper", yref="paper",
x=0.005, y=-0.002 ) ],
xaxis=dict(showgrid=False, zeroline=False, showticklabels=False),
yaxis=dict(showgrid=False, zeroline=False, showticklabels=False)))
py.iplot(fig, filename='networkx')
```
### Dash Example
[Dash](https://plot.ly/products/dash/) is an Open Source Python library which can help you convert plotly figures into a reactive, web-based application. Below is a simple example of a dashboard created using Dash. Its [source code](https://github.com/plotly/simple-example-chart-apps/tree/master/dash-networkplot) can easily be deployed to a PaaS.
```
from IPython.display import IFrame
IFrame(src= "https://dash-simple-apps.plotly.host/dash-networkplot/", width="100%", height="650px", frameBorder="0")
from IPython.display import IFrame
IFrame(src= "https://dash-simple-apps.plotly.host/dash-networkplot/code", width="100%", height=500, frameBorder="0")
```
#### Reference
See https://plot.ly/python/reference/#scatter for more information and chart attribute options!
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'networkx.ipynb', 'python/network-graphs/', 'Python Network Graphs',
'How to make Network Graphs in Python with Plotly. '
'One examples of a network graph with NetworkX',
name = 'Network Graphs',
thumbnail='thumbnail/net.jpg', language='python',
has_thumbnail='true', display_as='scientific', order=14, redirect_from='ipython-notebooks/networks/',
ipynb= '~notebook_demo/223')
```
| github_jupyter |
### Forest structure using PDAL + Python
Dr Adam Steer, November 2019.
This work is a set of python modules to replace MATLAB code for generating TERN forest metrics from airborne LIDAR.
## Fundamental ideas
Existing code uses a series of nested loops, meaning we can't take advantage of array operations or easily reformat or paralellise functionality
The approach used here defines a transportable function for each TERN product. These are applied to the data using a single loop (which could be chunked and parallelised).
A simple process step-through looks like:
1. Read LAS tile using PDAL. This removes an uncompression step. It also removes low outliers and computes normalised height for each point on the fly
2. Read numpy labelled arrays from PDAL output into a GeoPandas dataframe, and apply a 2D spatial index
3. From LAS file metadata, produce a fishnet grid with cells of size 'output resolution X output resolution'
4. Iterate over grid cells, select valid points and generate TERN products for each grid cell
5. Assemble an output array for each TERN product and write to GeoTIFF
This set of functions operates per-las-tile. An additional layer may be added to merge mutliple raster outputs into larger datasets
## to do:
- snake_casify variable names
```
NODATA_VALUE = -9999
#imports
import pdal
import numpy as np
import json
from shapely.geometry import Point
from shapely.geometry import MultiPolygon
from shapely.geometry import box
#from shapely.strtree import STRtree
import geopandas as gpd
import pandas as pd
import osmnx as ox
import os
# not using this, using geopandas instead
from rtree import index
# this is needed to create a raster from the output array
from osgeo import gdal
import osgeo.osr as osr
def writegeotiff(griddedpoints, outfile, parameters):
"""
writes out a geotiff from a numpy array of forest metric
results.
inputs:
- a numpy array of metrics [griddedpoints]
- an outfile name [outfile]
- a dictionary of parameters for the raster
outputs:
- a gdal dataset object
- [outfile] written to disk
"""
width = parameters["width"]
height = parameters["height"]
drv = gdal.GetDriverByName("GTiff")
ds = drv.Create(outfile, width, height, 6, gdal.GDT_Float32)
ds.SetGeoTransform(parameters["upperleft_y"],
parameters["resolution"],
0,
parameters["upperleft_y"],
0,
parameters["resolution"])
ds.setProjection = parameters["projection"]
ds.GetRasterBand(1).WriteArray(arr)
return(ds)
def pdal2df(points):
"""
Feed me a PDAL pipeline return array, get back a
GeoPandas dataframe
"""
arr = points[0]
description = arr.dtype.descr
cols = [col for col, __ in description]
gdf = gpd.GeoDataFrame({col: arr[col] for col in cols})
gdf.name = 'nodes'
gdf['geometry'] = gdf.apply(lambda row: Point((row['X'], row['Y'])), axis=1)
return(gdf_nodes)
def spatialindex(dataframe):
sindex = dataframe.sindex
return(sindex)
#get a pointview from PDAL
def readlasfile(lasfile):
"""
Run a PDAL pipeline. Input is a JSON declaration to
deliver to PDAL. Output is a labelled numpy array.
Data are filtered to:
- label local minima as noise
- compute height above ground using nearest ground point
neighbours (TIN method arriving soon)
- sort using a morton order (space filling curve) to
speed indexing later.
"""
pipeline = {
"pipeline": [
{
"type": "readers.las",
"filename": lasfile
},
{
"type": "filters.hag"
}
]
}
pipeline = pdal.Pipeline(json.dumps(pipeline))
count = pipeline.execute()
#read points into labelled arrays
arrays = pipeline.arrays
#return a numpy array to operate on
return(arrays)
def readlasmetadata(lasfile):
pipeline = {
"pipeline": [
{
"type": "readers.las",
"filename": lasfile,
"count": 1
},
{
"type": "filters.info"
}
]
}
pipeline = pdal.Pipeline(json.dumps(pipeline))
pipeline.validate()
pipeline.loglevel = 2 # stay quiet
count = pipeline.execute()
#extract metadata into a JSON blob
metadata = json.loads(pipeline.metadata)
return(metadata)
def readlaschunk(lasfile, poly):
"""
Run a PDAL pipeline to collect a chunk of points from a LASfile.
Inputs
- a PDAL readable file name
- a geometry bounding the chunk of data to be clipped
Output
- a labelled numpy array.
"""
xmin = str(np.int(np.floor(poly.bounds[0])))
ymin = str(np.int(np.floor(poly.bounds[1])))
xmax = str(np.int(np.ceil(poly.bounds[2])))
ymax = str(np.int(np.ceil(poly.bounds[3])))
cropbox = "([" + xmin + "," + ymin + "],[" + xmax + "," + ymax + "])"
print(cropbox)
pipeline = {
"pipeline": [
{
"type": "readers.las",
"filename": lasfile
},
{
"type": "filters.crop",
"bounds": cropbox
}
]
}
pipeline = pdal.Pipeline(json.dumps(pipeline))
count = pipeline.execute()
#read points into labelled arrays
array = pipeline.arrays
#return a numpy array to operate on
return(array)
def extract_vars(df):
"""
extract relevant variables
do we need to do this now? or wait till we've grabbed the indexed chunk?
lets write it anyway, then the index chunkifier can call it...
inputs:
- a numpy labelled array resulting from a PDAL LAS/LAZ file read
outputs:
- 1D arrays containing relevant variables
"""
classification = df["Classification"].values
intensity = df["Intensity"].values
returnnumber = df["ReturnNumber"].values
numberofreturns = df["NumberOfReturns"].values
elevation = df["Z"].values
hag = df["HeightAboveGround"].values
return(intensity, returnnumber, numberofreturns, elevation, hag)
def gen_raster_cells(metadata, resolution):
"""
Generate cells of 'resolution x resolution' for point querying
input:
- PDAL metadata
output:
- shapely geometry containing polygons defining 'resolution x resolution'
boxes covering the LAS tile extent
"""
bbox = box(metadata["metadata"]["readers.las"][0]["minx"],
metadata["metadata"]["readers.las"][0]["miny"],
metadata["metadata"]["readers.las"][0]["maxx"],
metadata["metadata"]["readers.las"][0]["maxy"])
tiledBBox = ox.quadrat_cut_geometry(bbox, quadrat_width=resolution)
return(tiledBBox)
def get_cell_points(poly, df, sindex):
poly = poly.buffer(1e-14).buffer(0)
possible_matches_index = list(sindex.intersection(poly.bounds))
possible_matches = df.iloc[possible_matches_index]
precise_matches = possible_matches[possible_matches.intersects(poly)]
return(precise_matches)
# Vegetation cover fraction: (Nfirst - Nsingle) / Nfirst
def comp_vcf(points):
"""
Computes vegetation cover fraction according to the TERN product manual.
inputs:
- a labelled array of points from an input LAS tile
outputs:
- a numpy array of grid cells containing the result of:
(Nfirst - Nsingle) / Nfirst
...where:
Nfirst = count of first returns
Nsingle = count of single returns
...per grid cell.
"""
# collect all the first and single return indices
nSingle = np.size(np.where(points["NumberOfReturns"].values == 1))
nFirst = np.size(np.where(points["ReturnNumber"].values == 1))
if (nFirst > 0):
vcf = (nFirst - nSingle) / nFirst
else:
print('no first returns, set vcf to {}'.format(NODATA_VALUE))
vcf = -9999
return(vcf)
# Canopy layering index:
# R = total returns
#
# vegetation layer cover fraction: LCF
def comp_lcf(points, heights, vcf):
"""
Compute LCF as per the TERN product manual:
LCF = VCF * (((veg returns below H2) - (veg returns below H1)) / (veg returns below H2))
Inputs:
- a set of points to compute LCF over
- a height threshold pair, containing H1 and H2 as an array [h1, h2]
- a precomputed VCF
Outputs:
- a floating point number denoting LCF
Conditions:
The LCF *must* be computed over the same set of points as the VCF used as input.
"""
h1 = heights[0]
h2 = heights[1]
#find veg returns - ASPRS classes 3,4,5
veg_returns = np.where(np.logical_or(points["Classification"].values == 3,
points["Classification"].values == 4,
points["Classification"].values == 5))
# how many veg returns have height below the first threshold?
vegbelowh1 = np.size(np.where(points["HeightAboveGround"][vegreturns] < h1))
# how many veg returns have height below the second threshold?
vegbelowh2 = np.size(np.where(points["HeightAboveGround"][vegreturns] < h2))
# compute the LCF
lcf = vcf * ( (vegbelowh2 - vegbelowh1) / vegbelowh2)
return(lcf)
#CTH
def comp_cth(points):
# compute the highest vegetation point in each grid cell
veg_returns = np.where(np.logical_or(points["Classification"].values == 3,
points["Classification"].values == 4,
points["Classification"].values == 5))
vegpoints = points["HeightAboveGround"].values[veg_returns]
cth = np.max(vegpoints)
return(cth)
def comp_dem(points):
# interpolate ground returns in a grid and output a raster
return()
def comp_fbf(points):
# if building classes exist, compute a fractional conver per grid cell...
return()
def read_data(lasfile):
"""
wrapper to read in LAS data and produce a dataframe + spatial index
"""
metadata, points = readlasfile(lasfile)
dataframe, spatial_index = pdal2df(points)
return(metadata, dataframe, spatial_index)
def compute_tern_products(metadata, points, sindex, resolution):
"""
Wrapper to iterate over the input data and generate rasters for each product.
*note this part could be paralellised - maybe per-product, or per-cell
Each grid square processed in this loop corresponds to one pixel in an output raster.
"""
#set up an 'output resolution' sized grid - like a fishnet grid.
# each polygon in the resulting set covers an area of 'resolution X resolution'
pixel_grid = gen_raster_cells(metadata, resolution)
#set up output rasters
# get tile width and height
tile_width = metadata["metadata"]["readers.las"][0]["maxx"] - metadata["metadata"]["readers.las"][0]["minx"]
tile_height = metadata["metadata"]["readers.las"][0]["maxy"] - metadata["metadata"]["readers.las"][0]["miny"]
raster_xsize = int(np.ceil(tile_width) / resolution)
raster_ysize = int(np.ceil(tile_height) / resolution)
print(tile_width)
print(raster_xsize)
vcf_raster = np.zeros((raster_xsize, raster_ysize))
print(np.shape(vcf_raster))
lcf_raster = np.zeros((raster_xsize, raster_ysize))
cth_raster = np.zeros((raster_xsize, raster_ysize))
for pixel in pixel_grid:
#compute output array index for this cell:
poly_x, poly_y = pixel.centroid.xy
poly_base_x = poly_x[0] - metadata["metadata"]["readers.las"][0]["minx"]
poly_base_y = poly_y[0] - metadata["metadata"]["readers.las"][0]["miny"]
print(poly_base_x)
print(poly_base_y)
array_x = int(np.floor((poly_base_x / (resolution)) ))
array_y = int(np.floor((poly_base_y / (resolution)) ))
#print('array X: {}; array Y: {}'.format(array_x, array_y))
#get points for this cell
matches = get_cell_points(pixel, points, sindex)
#compute in order
#VCF
vcf_raster[array_x, array_y] = comp_vcf(matches)
#LCF - need stuff about levels here...
#lcf_raster[array_x, array_y] = comp_lcf(points)
#CTH
try:
cth_raster[array_x, array_y] = comp_cth(matches)
except ValueError:
print('no vegetation returns were present, CTH set to {} for array index {} {}'.format(NODATA_VALUE, array_x, array_y))
cth_raster[array_x, array_y] = NODATA_VALUE
#end of computing stuff
#extract EPSG code from LAS:
return(vcf_raster, cth_raster)
```
## Testing functionality using a local file
The following section generates metrics from a local LAZ file. Plugging in download mechanics from ELVIS will be added later
```
#lidar test file - Mt Ainslie, chosen for varied vegetation cover and topography
# this is pretty big, may need splitting up!
#lasfile = "/Volumes/Antares/ACT-lidar/8ppm/callingelvis-testdata/ACT2015_8ppm-C3-AHD_6966094_55.las"
lasfile = "/Volumes/Antares/fire-test/NSW Government - Spatial Services-2/Point Clouds/AHD/StAlbans201709-LID2-C3-AHD_2866308_56_0002_0002/StAlbans201709-LID2-C3-AHD_2866308_56_0002_0002.las"
lasfile = "./Berridale201802-LID2-C3-AHD_6585974_55_0002_0002.las"
%%time
metadata = readlasmetadata(lasfile)
metadata
resolution = 25
pixel_grid = gen_raster_cells(metadata, resolution)
pixel_grid[0]
boxbounds = pixel_grid[0].bounds[0]
boxbounds
#get a pointview from PDAL
def readlaschunk(lasfile, poly):
"""
Run a PDAL pipeline to collect a chunk of points from a LASfile.
Inputs
- a PDAL readable file name
- a geometry bounding the chunk of data to be clipped
Output
- a labelled numpy array.
"""
xmin = str(np.int(np.floor(poly.bounds[0])))
ymin = str(np.int(np.floor(poly.bounds[1])))
xmax = str(np.int(np.ceil(poly.bounds[2])))
ymax = str(np.int(np.ceil(poly.bounds[3])))
cropbox = "([" + xmin + "," + ymin + "],[" + xmax + "," + ymax + "])"
print(cropbox)
pipeline = {
"pipeline": [
{
"type": "readers.las",
"filename": lasfile
},
{
"type": "filters.crop",
"bounds": cropbox
}
]
}
pipeline = pdal.Pipeline(json.dumps(pipeline))
count = pipeline.execute()
#read points into labelled arrays
array = pipeline.arrays
#return a numpy array to operate on
return(array)
%%time
laschunk = readlaschunk(lasfile, pixel_grid[0])
laschunk
%%time
laspoints = readlasfile(lasfile)
%%time
#here we read points into a GeoDataFrame and dump the labelled array.
# this is a pretty expensive step RAM wise, we're duplicating all the points...
df = pdal2df(laspoints)
# set points to None, we don't use them anymore
laspoints = None
%%time
sindex = spatialindex(df)
# dump everything from memory
points = None
df = None
vcf_raster = None
cth_raster = None
%%time
# this part of the process is simply reading from the source file. No analysis yet.
metadata, points = readlasfile(lasfile)
%%time
#here we read points into a GeoDataFrame and dump the labelled array.
# this is a pretty expensive step RAM wise, we're duplicating all the points...
df = pdal2df(points)
# set points to None, we don't use them anymore
points = None
%%time
# here we generate an RTree index on the dataframe using GeoPandas.
# also pretty expensive...
sindex = spatialindex(df)
%%time
## rtree index building straight from the point dataset...
idx = index.Index()
for pid, point in enumerate(points[0]):
idx.insert(pid, (point[0], point[1],point[0], point[1]), point)
# set an output resolution
resolution = 100
%%time
vcf_raster, cth_raster = compute_tern_products(metadata, df, sindex, resolution)
from matplotlib import pyplot as plt
%matplotlib inline
plt.imshow(vcf_rasters)
plt.colorbar()
plt.imshow(cth_rasters)
plt.colorbar()
wktcrs = metadata["metadata"]["readers.las"][0]["comp_spatialreference"]
type(wktcrs)
srs = osr.SpatialReference()
srs.SetFromUserInput("EPSG:28356")
srs.ImportFromWkt(wktcrs)
srs.GetAuthorityCode(None)
srs.GetAuthorityName(None)
tiledBBox = gen_raster_cells(metadata,resolution)
tiledBBox[50]
%%time
idxpoints, tree = create_spatial_index(points)
%%time
thepoints = tree.query(geometry_cut[50])
query
```
## Run a sample workflow on one square
```
pixel_grid[0]
matches = get_cell_points(pixel_grid[10],df, sindex)
def comp_cth1(points):
# compute the highest vegetation point in each grid cell
veg_returns = np.where(np.logical_or(points["Classification"].values == 3,
points["Classification"].values == 4,
points["Classification"].values == 5))
vegpoints = points["HeightAboveGround"].values[veg_returns]
cth = np.max(vegpoints)
return(cth)
comp_cth1(matches)
```
## create a dataframe for pretty querying purposes..
```
points
#import pandas as pd
#points = None
# find the points that intersect with each subpolygon and add them to points_within_geometry
points_within_geometry = pd.DataFrame()
def get_points(poly, df, sindex):
poly = poly.buffer(1e-14).buffer(0)
possible_matches_index = list(sindex.intersection(poly.bounds))
possible_matches = df.iloc[possible_matches_index]
precise_matches = possible_matches[possible_matches.intersects(poly)]
return(precise_matches)
polyX, polyY = poly.centroid.xy
polyX
polyY
polyBaseXCoord = polyX[0] - metadata["metadata"]["readers.las"][0]["minx"]
arrayXindex = (polyBaseXCoord / (resolution/2 )) -1
arrayXindex
polyBaseYCoord = polyY[0] - metadata["metadata"]["readers.las"][0]["miny"]
arrayYindex = (polyBaseYCoord / (resolution/2 )) - 1
arrayYindex
arrayX = PolyBaseCoord - numberofcells
polyY[0] - metadata["metadata"]["readers.las"][0]["miny"]
tilewidth = metadata["metadata"]["readers.las"][0]["maxx"] - metadata["metadata"]["readers.las"][0]["minx"]
tilewidth
tileheight = metadata["metadata"]["readers.las"][0]["maxy"] - metadata["metadata"]["readers.las"][0]["miny"]
tileheight
vcfRaster = np.zeros((tilewidth, tileheight))
vcfRaster
%%time
matches = get_points(poly, df)
matches
intensity, returnnumber, numberofreturns, elevation, hag = extract_vars(matches)
np.size(np.where(matches["NumberOfReturns"].values == 1))
## OK now we can make magic - extracting each grid cell, we can rasterify it...
vcf = comp_vcf(matches)
vcf
vcfRaster[0,138] = vcf
vcfRaster
this = matches["Classification"].values
veg = np.where(np.logical_or(matches["Classification"].values == 3,matches["Classification"].values == 4, matches["Classification"].values == 5))
this = matches["Classification"].values
np.where(matches["HeightAboveGround"].values[veg] < 1)
lcf0105 = comp_lcf(matches, [1, 5], vcf)
```
## code purgatory
stuff here might be useful, or not
```
# this will likely evolve to take a 'what to grid' input
# actually likely not needed...
def grid_setup(pointmetadata, resolution):
"""
Sets up array indexes for an incoming las file, using an input resolution
input:
- las file metadata
- a scalar resolution
output:
- a numpy array of indexing values to divide the input points into
'resolution' x 'resolution' bins.
implicit assumptions:
- 'resolution' is always set in native LAS file units
"""
return()
# not used yet - geopandas is doing this part!
def create_spatial_index(arrays):
"""
task here is to map geospatial space to numpy array space.
- using the point metadata set up a grid of [resolution] x [resolution]
- scan the point coordinates array to see which points live in which cell
- create an index which maps point array indexes to grid indexes
idly wondering if a fast point-in-polygon does this job. Shapely / OGR to the rescue?
...this would be parallelisable...
"""
points = []
for thepoint in arrays[0]:
#print(thepoint[0])
points.append(Point(thepoint[0], thepoint[1]))
tree = STRtree(points)
return(points, tree)
```
| github_jupyter |
## 1. Where are the old left-handed people?
<p><img src="https://s3.amazonaws.com/assets.datacamp.com/production/project_479/img/Obama_signs_health_care-20100323.jpg" alt="Barack Obama signs the Patient Protection and Affordable Care Act at the White House, March 23, 2010"></p>
<p>Barack Obama is left-handed. So are Bill Gates and Oprah Winfrey; so were Babe Ruth and Marie Curie. A <a href="https://www.nejm.org/doi/full/10.1056/NEJM199104043241418">1991 study</a> reported that left-handed people die on average nine years earlier than right-handed people. Nine years! Could this really be true? </p>
<p>In this notebook, we will explore this phenomenon using age distribution data to see if we can reproduce a difference in average age at death purely from the changing rates of left-handedness over time, refuting the claim of early death for left-handers. This notebook uses <code>pandas</code> and Bayesian statistics to analyze the probability of being a certain age at death given that you are reported as left-handed or right-handed.</p>
<p>A National Geographic survey in 1986 resulted in over a million responses that included age, sex, and hand preference for throwing and writing. Researchers Avery Gilbert and Charles Wysocki analyzed this data and noticed that rates of left-handedness were around 13% for people younger than 40 but decreased with age to about 5% by the age of 80. They concluded based on analysis of a subgroup of people who throw left-handed but write right-handed that this age-dependence was primarily due to changing social acceptability of left-handedness. This means that the rates aren't a factor of <em>age</em> specifically but rather of the <em>year you were born</em>, and if the same study was done today, we should expect a shifted version of the same distribution as a function of age. Ultimately, we'll see what effect this changing rate has on the apparent mean age of death of left-handed people, but let's start by plotting the rates of left-handedness as a function of age.</p>
<p>This notebook uses two datasets: <a href="https://www.cdc.gov/nchs/data/statab/vs00199_table310.pdf">death distribution data</a> for the United States from the year 1999 (source website <a href="https://www.cdc.gov/nchs/nvss/mortality_tables.htm">here</a>) and rates of left-handedness digitized from a figure in this <a href="https://www.ncbi.nlm.nih.gov/pubmed/1528408">1992 paper by Gilbert and Wysocki</a>. </p>
```
# import libraries
# ... YOUR CODE FOR TASK 1 ...
import pandas as pd
import matplotlib.pyplot as plt
# load the data
data_url_1 = "https://gist.githubusercontent.com/mbonsma/8da0990b71ba9a09f7de395574e54df1/raw/aec88b30af87fad8d45da7e774223f91dad09e88/lh_data.csv"
lefthanded_data = pd.read_csv(data_url_1)
# plot male and female left-handedness rates vs. age
%matplotlib inline
fig, ax = plt.subplots() # create figure and axis objects
ax.plot("Age", "Female", data=lefthanded_data, marker = 'o') # plot "Female" vs. "Age"
ax.plot("Age", "Male", data=lefthanded_data, marker = 'x') # plot "Male" vs. "Age"
ax.legend() # add a legend
ax.set_xlabel("Sex")
ax.set_ylabel("Age")
```
## 2. Rates of left-handedness over time
<p>Let's convert this data into a plot of the rates of left-handedness as a function of the year of birth, and average over male and female to get a single rate for both sexes. </p>
<p>Since the study was done in 1986, the data after this conversion will be the percentage of people alive in 1986 who are left-handed as a function of the year they were born. </p>
```
lefthanded_data["Birth_year"] = 1986 - lefthanded_data["Age"]
# create a new column for the average of male and female
# ... YOUR CODE FOR TASK 2 ...
lefthanded_data["Mean_lh"] = lefthanded_data[["Female","Male"]].mean(axis=1)
# create a plot of the 'Mean_lh' column vs. 'Birth_year'
fig, ax = plt.subplots()
ax.plot("Birth_year", "Mean_lh", data=lefthanded_data) # plot 'Mean_lh' vs. 'Birth_year'
ax.set_xlabel("Mean_lh") # set the x label for the plot
ax.set_ylabel("Birth_year") # set the y label for the plot
```
## 3. Applying Bayes' rule
<p>The probability of dying at a certain age given that you're left-handed is <strong>not</strong> equal to the probability of being left-handed given that you died at a certain age. This inequality is why we need <strong>Bayes' theorem</strong>, a statement about conditional probability which allows us to update our beliefs after seeing evidence. </p>
<p>We want to calculate the probability of dying at age A given that you're left-handed. Let's write this in shorthand as P(A | LH). We also want the same quantity for right-handers: P(A | RH). </p>
<p>Here's Bayes' theorem for the two events we care about: left-handedness (LH) and dying at age A.</p>
<p>$$P(A | LH) = \frac{P(LH|A) P(A)}{P(LH)}$$</p>
<p>P(LH | A) is the probability that you are left-handed <em>given that</em> you died at age A. P(A) is the overall probability of dying at age A, and P(LH) is the overall probability of being left-handed. We will now calculate each of these three quantities, beginning with P(LH | A).</p>
<p>To calculate P(LH | A) for ages that might fall outside the original data, we will need to extrapolate the data to earlier and later years. Since the rates flatten out in the early 1900s and late 1900s, we'll use a few points at each end and take the mean to extrapolate the rates on each end. The number of points used for this is arbitrary, but we'll pick 10 since the data looks flat-ish until about 1910. </p>
```
import numpy as np
# create a function for P(LH | A)
def P_lh_given_A(ages_of_death, study_year = 1990):
""" P(Left-handed | ages of death), calculated based on the reported rates of left-handedness.
Inputs: numpy array of ages of death, study_year
Returns: probability of left-handedness given that subjects died in `study_year` at ages `ages_of_death` """
# Use the mean of the 10 last and 10 first points for left-handedness rates before and after the start
early_1900s_rate = lefthanded_data["Mean_lh"][-10:].mean()
late_1900s_rate = lefthanded_data["Mean_lh"][:10].mean()
middle_rates = lefthanded_data.loc[lefthanded_data['Birth_year'].isin(study_year - ages_of_death)]['Mean_lh']
youngest_age = study_year - 1986 + 10 # the youngest age is 10
oldest_age = study_year - 1986 + 86 # the oldest age is 86
P_return = np.zeros(ages_of_death.shape) # create an empty array to store the results
# extract rate of left-handedness for people of ages 'ages_of_death'
P_return[ages_of_death > oldest_age] = early_1900s_rate/100
P_return[ages_of_death < youngest_age] = late_1900s_rate/100
P_return[np.logical_and((ages_of_death <= oldest_age), (ages_of_death >= youngest_age))] = middle_rates/100
return P_return
```
## 4. When do people normally die?
<p>To estimate the probability of living to an age A, we can use data that gives the number of people who died in a given year and how old they were to create a distribution of ages of death. If we normalize the numbers to the total number of people who died, we can think of this data as a probability distribution that gives the probability of dying at age A. The data we'll use for this is from the entire US for the year 1999 - the closest I could find for the time range we're interested in. </p>
<p>In this block, we'll load in the death distribution data and plot it. The first column is the age, and the other columns are the number of people who died at that age. </p>
```
# Death distribution data for the United States in 1999
data_url_2 = "https://gist.githubusercontent.com/mbonsma/2f4076aab6820ca1807f4e29f75f18ec/raw/62f3ec07514c7e31f5979beeca86f19991540796/cdc_vs00199_table310.tsv"
death_distribution_data = pd.read_csv(data_url_2, sep = "\t", skiprows=[1])
# drop NaN values from the `Both Sexes` column
# ... YOUR CODE FOR TASK 4 ...
death_distribution_data = death_distribution_data.dropna(subset = ["Both Sexes"])
# plot number of people who died as a function of age
fig, ax = plt.subplots()
ax.plot("Age", "Both Sexes", data = death_distribution_data, marker='o') # plot 'Both Sexes' vs. 'Age'
ax.set_xlabel("Both Sexes")
ax.set_ylabel("Age")
```
## 5. The overall probability of left-handedness
<p>In the previous code block we loaded data to give us P(A), and now we need P(LH). P(LH) is the probability that a person who died in our particular study year is left-handed, assuming we know nothing else about them. This is the average left-handedness in the population of deceased people, and we can calculate it by summing up all of the left-handedness probabilities for each age, weighted with the number of deceased people at each age, then divided by the total number of deceased people to get a probability. In equation form, this is what we're calculating, where N(A) is the number of people who died at age A (given by the dataframe <code>death_distribution_data</code>):</p>
<p><img src="https://i.imgur.com/gBIWykY.png" alt="equation" width="220"></p>
<!--- $$P(LH) = \frac{\sum_{\text{A}} P(LH | A) N(A)}{\sum_{\text{A}} N(A)}$$ -->
```
def P_lh(death_distribution_data, study_year = 1990): # sum over P_lh for each age group
p_list = death_distribution_data["Both Sexes"]*P_lh_given_A(death_distribution_data["Age"], study_year) # multiply number of dead people by P_lh_given_A
p = np.sum(p_list) # calculate the sum of p_list
return p/np.sum(death_distribution_data["Both Sexes"]) # normalize to total number of people (sum of death_distribution_data['Both Sexes'])
print(P_lh(death_distribution_data, 1990))
```
## 6. Putting it all together: dying while left-handed (i)
<p>Now we have the means of calculating all three quantities we need: P(A), P(LH), and P(LH | A). We can combine all three using Bayes' rule to get P(A | LH), the probability of being age A at death (in the study year) given that you're left-handed. To make this answer meaningful, though, we also want to compare it to P(A | RH), the probability of being age A at death given that you're right-handed. </p>
<p>We're calculating the following quantity twice, once for left-handers and once for right-handers.</p>
<p>$$P(A | LH) = \frac{P(LH|A) P(A)}{P(LH)}$$</p>
<p>First, for left-handers.</p>
<!--Notice that I was careful not to call these "probability of dying at age A", since that's not actually what we're calculating: we use the exact same death distribution data for each. -->
```
def P_A_given_lh(ages_of_death, death_distribution_data, study_year = 1990):
P_A = death_distribution_data["Both Sexes"][ages_of_death]/np.sum(death_distribution_data["Both Sexes"])
P_left = P_lh(death_distribution_data, study_year) # use P_lh function to get probability of left-handedness overall
P_lh_A = P_lh_given_A(ages_of_death, study_year) # use P_lh_given_A to get probability of left-handedness for a certain age
return P_lh_A*P_A/P_left
```
## 7. Putting it all together: dying while left-handed (ii)
<p>And now for right-handers.</p>
```
def P_A_given_rh(ages_of_death, death_distribution_data, study_year = 1990):
P_A = death_distribution_data["Both Sexes"][ages_of_death]/np.sum(death_distribution_data["Both Sexes"])
P_right = 1 - P_lh(death_distribution_data, study_year)# either you're left-handed or right-handed, so P_right = 1 - P_left
P_rh_A = 1 - P_lh_given_A(ages_of_death, study_year) # P_rh_A = 1 - P_lh_A
return P_rh_A*P_A/P_right
```
## 8. Plotting the distributions of conditional probabilities
<p>Now that we have functions to calculate the probability of being age A at death given that you're left-handed or right-handed, let's plot these probabilities for a range of ages of death from 6 to 120. </p>
<p>Notice that the left-handed distribution has a bump below age 70: of the pool of deceased people, left-handed people are more likely to be younger. </p>
```
ages = np.arange(6, 120) # make a list of ages of death to plot
# calculate the probability of being left- or right-handed for each
left_handed_probability = P_A_given_lh(ages, death_distribution_data)
right_handed_probability = P_A_given_rh(ages, death_distribution_data)
# create a plot of the two probabilities vs. age
fig, ax = plt.subplots() # create figure and axis objects
ax.plot(ages, left_handed_probability, label = "Left-handed")
ax.plot(ages, right_handed_probability, label = "Right-handed")
ax.legend() # add a legend
ax.set_xlabel("Age at death")
ax.set_ylabel(r"Probability of being age A at death")
```
## 9. Moment of truth: age of left and right-handers at death
<p>Finally, let's compare our results with the original study that found that left-handed people were nine years younger at death on average. We can do this by calculating the mean of these probability distributions in the same way we calculated P(LH) earlier, weighting the probability distribution by age and summing over the result.</p>
<p>$$\text{Average age of left-handed people at death} = \sum_A A P(A | LH)$$</p>
<p>$$\text{Average age of right-handed people at death} = \sum_A A P(A | RH)$$</p>
```
# calculate average ages for left-handed and right-handed groups
# use np.array so that two arrays can be multiplied
average_lh_age = np.nansum(ages*np.array(left_handed_probability))
average_rh_age = np.nansum(ages*np.array(right_handed_probability))
# print the average ages for each group
# ... YOUR CODE FOR TASK 9 ...
print("Average age of lefthanded" + str(average_lh_age))
print("Average age of righthanded" + str(average_rh_age))
# print the difference between the average ages
print("The difference in average ages is " + str(round(average_lh_age - average_rh_age, 1)) + " years.")
```
## 10. Final comments
<p>We got a pretty big age gap between left-handed and right-handed people purely as a result of the changing rates of left-handedness in the population, which is good news for left-handers: you probably won't die young because of your sinisterness. The reported rates of left-handedness have increased from just 3% in the early 1900s to about 11% today, which means that older people are much more likely to be reported as right-handed than left-handed, and so looking at a sample of recently deceased people will have more old right-handers.</p>
<p>Our number is still less than the 9-year gap measured in the study. It's possible that some of the approximations we made are the cause: </p>
<ol>
<li>We used death distribution data from almost ten years after the study (1999 instead of 1991), and we used death data from the entire United States instead of California alone (which was the original study). </li>
<li>We extrapolated the left-handedness survey results to older and younger age groups, but it's possible our extrapolation wasn't close enough to the true rates for those ages. </li>
</ol>
<p>One thing we could do next is figure out how much variability we would expect to encounter in the age difference purely because of random sampling: if you take a smaller sample of recently deceased people and assign handedness with the probabilities of the survey, what does that distribution look like? How often would we encounter an age gap of nine years using the same data and assumptions? We won't do that here, but it's possible with this data and the tools of random sampling. </p>
<!-- I did do this if we want to add more tasks - it would probably take three more blocks.-->
<p>To finish off, let's calculate the age gap we'd expect if we did the study in 2018 instead of in 1990. The gap turns out to be much smaller since rates of left-handedness haven't increased for people born after about 1960. Both the National Geographic study and the 1990 study happened at a unique time - the rates of left-handedness had been changing across the lifetimes of most people alive, and the difference in handedness between old and young was at its most striking. </p>
```
# Calculate the probability of being left- or right-handed for all ages
left_handed_probability_2018 = ...
right_handed_probability_2018 = ...
# calculate average ages for left-handed and right-handed groups
average_lh_age_2018 = np.nansum(ages*np.array(left_handed_probability_2018))
average_rh_age_2018 = np.nansum(ages*np.array(right_handed_probability_2018))
print("The difference in average ages is " +
str(round(average_rh_age_2018 - average_lh_age_2018, 1)) + " years.")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/AI4Finance-LLC/FinRL-Library/blob/master/FinRL_stock_trading_NeurIPS_2018.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Deep Reinforcement Learning for Stock Trading from Scratch: Multiple Stock Trading
Tutorials to use OpenAI DRL to trade multiple stocks in one Jupyter Notebook | Presented at NeurIPS 2020: Deep RL Workshop
* This blog uses FinRL to reproduce the paper: Practical Deep Reinforcement Learning Approach for Stock Trading, Workshop on Challenges and Opportunities for AI in Financial Services, NeurIPS 2018.
* Check out medium blog for detailed explanations: https://towardsdatascience.com/finrl-for-quantitative-finance-tutorial-for-multiple-stock-trading-7b00763b7530
* Please report any issues to our Github: https://github.com/AI4Finance-LLC/FinRL-Library/issues
* **Pytorch Version**
# Content
* [1. Problem Definition](#0)
* [2. Getting Started - Load Python packages](#1)
* [2.1. Install Packages](#1.1)
* [2.2. Check Additional Packages](#1.2)
* [2.3. Import Packages](#1.3)
* [2.4. Create Folders](#1.4)
* [3. Download Data](#2)
* [4. Preprocess Data](#3)
* [4.1. Technical Indicators](#3.1)
* [4.2. Perform Feature Engineering](#3.2)
* [5.Build Environment](#4)
* [5.1. Training & Trade Data Split](#4.1)
* [5.2. User-defined Environment](#4.2)
* [5.3. Initialize Environment](#4.3)
* [6.Implement DRL Algorithms](#5)
* [7.Backtesting Performance](#6)
* [7.1. BackTestStats](#6.1)
* [7.2. BackTestPlot](#6.2)
* [7.3. Baseline Stats](#6.3)
* [7.3. Compare to Stock Market Index](#6.4)
<a id='0'></a>
# Part 1. Problem Definition
This problem is to design an automated trading solution for single stock trading. We model the stock trading process as a Markov Decision Process (MDP). We then formulate our trading goal as a maximization problem.
The algorithm is trained using Deep Reinforcement Learning (DRL) algorithms and the components of the reinforcement learning environment are:
* Action: The action space describes the allowed actions that the agent interacts with the
environment. Normally, a ∈ A includes three actions: a ∈ {−1, 0, 1}, where −1, 0, 1 represent
selling, holding, and buying one stock. Also, an action can be carried upon multiple shares. We use
an action space {−k, ..., −1, 0, 1, ..., k}, where k denotes the number of shares. For example, "Buy
10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or −10, respectively
* Reward function: r(s, a, s′) is the incentive mechanism for an agent to learn a better action. The change of the portfolio value when action a is taken at state s and arriving at new state s', i.e., r(s, a, s′) = v′ − v, where v′ and v represent the portfolio
values at state s′ and s, respectively
* State: The state space describes the observations that the agent receives from the environment. Just as a human trader needs to analyze various information before executing a trade, so
our trading agent observes many different features to better learn in an interactive environment.
* Environment: Dow 30 consituents
The data of the single stock that we will be using for this case study is obtained from Yahoo Finance API. The data contains Open-High-Low-Close price and volume.
<a id='1'></a>
# Part 2. Getting Started- Load Python Packages
<a id='1.1'></a>
## 2.1. Install all the packages through FinRL library
```
## install finrl library
!pip install git+https://github.com/AI4Finance-LLC/FinRL-Library.git
```
<a id='1.2'></a>
## 2.2. Check if the additional packages needed are present, if not install them.
* Yahoo Finance API
* pandas
* numpy
* matplotlib
* stockstats
* OpenAI gym
* stable-baselines
* tensorflow
* pyfolio
<a id='1.3'></a>
## 2.3. Import Packages
```
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# matplotlib.use('Agg')
import datetime
%matplotlib inline
from finrl.config import config
from finrl.marketdata.yahoodownloader import YahooDownloader
from finrl.preprocessing.preprocessors import FeatureEngineer
from finrl.preprocessing.data import data_split
from finrl.env.env_stocktrading import StockTradingEnv
from finrl.model.models import DRLAgent
from finrl.trade.backtest import backtest_stats, backtest_plot, get_daily_return, get_baseline
from pprint import pprint
import sys
sys.path.append("../FinRL-Library")
import itertools
```
<a id='1.4'></a>
## 2.4. Create Folders
```
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR):
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
```
<a id='2'></a>
# Part 3. Download Data
Yahoo Finance is a website that provides stock data, financial news, financial reports, etc. All the data provided by Yahoo Finance is free.
* FinRL uses a class **YahooDownloader** to fetch data from Yahoo Finance API
* Call Limit: Using the Public API (without authentication), you are limited to 2,000 requests per hour per IP (or up to a total of 48,000 requests a day).
-----
class YahooDownloader:
Provides methods for retrieving daily stock data from
Yahoo Finance API
Attributes
----------
start_date : str
start date of the data (modified from config.py)
end_date : str
end date of the data (modified from config.py)
ticker_list : list
a list of stock tickers (modified from config.py)
Methods
-------
fetch_data()
Fetches data from yahoo API
```
# from config.py start_date is a string
config.START_DATE
# from config.py end_date is a string
config.END_DATE
print(config.DOW_30_TICKER)
df = YahooDownloader(start_date = '2009-01-01',
end_date = '2021-01-01',
ticker_list = config.DOW_30_TICKER).fetch_data()
df.shape
df.sort_values(['date','tic'],ignore_index=True).head()
```
# Part 4: Preprocess Data
Data preprocessing is a crucial step for training a high quality machine learning model. We need to check for missing data and do feature engineering in order to convert the data into a model-ready state.
* Add technical indicators. In practical trading, various information needs to be taken into account, for example the historical stock prices, current holding shares, technical indicators, etc. In this article, we demonstrate two trend-following technical indicators: MACD and RSI.
* Add turbulence index. Risk-aversion reflects whether an investor will choose to preserve the capital. It also influences one's trading strategy when facing different market volatility level. To control the risk in a worst-case scenario, such as financial crisis of 2007–2008, FinRL employs the financial turbulence index that measures extreme asset price fluctuation.
```
fe = FeatureEngineer(
use_technical_indicator=True,
tech_indicator_list = config.TECHNICAL_INDICATORS_LIST,
use_turbulence=True,
user_defined_feature = False)
processed = fe.preprocess_data(df)
list_ticker = processed["tic"].unique().tolist()
list_date = list(pd.date_range(processed['date'].min(),processed['date'].max()).astype(str))
combination = list(itertools.product(list_date,list_ticker))
processed_full = pd.DataFrame(combination,columns=["date","tic"]).merge(processed,on=["date","tic"],how="left")
processed_full = processed_full[processed_full['date'].isin(processed['date'])]
processed_full = processed_full.sort_values(['date','tic'])
processed_full = processed_full.fillna(0)
processed_full.sort_values(['date','tic'],ignore_index=True).head(10)
```
<a id='4'></a>
# Part 5. Design Environment
Considering the stochastic and interactive nature of the automated stock trading tasks, a financial task is modeled as a **Markov Decision Process (MDP)** problem. The training process involves observing stock price change, taking an action and reward's calculation to have the agent adjusting its strategy accordingly. By interacting with the environment, the trading agent will derive a trading strategy with the maximized rewards as time proceeds.
Our trading environments, based on OpenAI Gym framework, simulate live stock markets with real market data according to the principle of time-driven simulation.
The action space describes the allowed actions that the agent interacts with the environment. Normally, action a includes three actions: {-1, 0, 1}, where -1, 0, 1 represent selling, holding, and buying one share. Also, an action can be carried upon multiple shares. We use an action space {-k,…,-1, 0, 1, …, k}, where k denotes the number of shares to buy and -k denotes the number of shares to sell. For example, "Buy 10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or -10, respectively. The continuous action space needs to be normalized to [-1, 1], since the policy is defined on a Gaussian distribution, which needs to be normalized and symmetric.
## Training data split: 2009-01-01 to 2018-12-31
## Trade data split: 2019-01-01 to 2020-09-30
```
train = data_split(processed_full, '2009-01-01','2019-01-01')
trade = data_split(processed_full, '2019-01-01','2021-01-01')
print(len(train))
print(len(trade))
train.head()
trade.head()
config.TECHNICAL_INDICATORS_LIST
stock_dimension = len(train.tic.unique())
state_space = 1 + 2*stock_dimension + len(config.TECHNICAL_INDICATORS_LIST)*stock_dimension
print(f"Stock Dimension: {stock_dimension}, State Space: {state_space}")
env_kwargs = {
"hmax": 100,
"initial_amount": 1000000,
"buy_cost_pct": 0.001,
"sell_cost_pct": 0.001,
"state_space": state_space,
"stock_dim": stock_dimension,
"tech_indicator_list": config.TECHNICAL_INDICATORS_LIST,
"action_space": stock_dimension,
"reward_scaling": 1e-4
}
e_train_gym = StockTradingEnv(df = train, **env_kwargs)
```
## Environment for Training
```
env_train, _ = e_train_gym.get_sb_env()
print(type(env_train))
```
<a id='5'></a>
# Part 6: Implement DRL Algorithms
* The implementation of the DRL algorithms are based on **OpenAI Baselines** and **Stable Baselines**. Stable Baselines is a fork of OpenAI Baselines, with a major structural refactoring, and code cleanups.
* FinRL library includes fine-tuned standard DRL algorithms, such as DQN, DDPG,
Multi-Agent DDPG, PPO, SAC, A2C and TD3. We also allow users to
design their own DRL algorithms by adapting these DRL algorithms.
```
agent = DRLAgent(env = env_train)
```
### Model Training: 5 models, A2C DDPG, PPO, TD3, SAC
### Model 1: A2C
```
agent = DRLAgent(env = env_train)
model_a2c = agent.get_model("a2c")
trained_a2c = agent.train_model(model=model_a2c,
tb_log_name='a2c',
total_timesteps=100000)
```
### Model 2: DDPG
```
agent = DRLAgent(env = env_train)
model_ddpg = agent.get_model("ddpg")
trained_ddpg = agent.train_model(model=model_ddpg,
tb_log_name='ddpg',
total_timesteps=50000)
```
### Model 3: PPO
```
agent = DRLAgent(env = env_train)
PPO_PARAMS = {
"n_steps": 2048,
"ent_coef": 0.01,
"learning_rate": 0.00025,
"batch_size": 128,
}
model_ppo = agent.get_model("ppo",model_kwargs = PPO_PARAMS)
trained_ppo = agent.train_model(model=model_ppo,
tb_log_name='ppo',
total_timesteps=50000)
```
### Model 4: TD3
```
agent = DRLAgent(env = env_train)
TD3_PARAMS = {"batch_size": 100,
"buffer_size": 1000000,
"learning_rate": 0.001}
model_td3 = agent.get_model("td3",model_kwargs = TD3_PARAMS)
trained_td3 = agent.train_model(model=model_td3,
tb_log_name='td3',
total_timesteps=30000)
```
### Model 5: SAC
```
agent = DRLAgent(env = env_train)
SAC_PARAMS = {
"batch_size": 128,
"buffer_size": 1000000,
"learning_rate": 0.0001,
"learning_starts": 100,
"ent_coef": "auto_0.1",
}
model_sac = agent.get_model("sac",model_kwargs = SAC_PARAMS)
trained_sac = agent.train_model(model=model_sac,
tb_log_name='sac',
total_timesteps=80000)
```
## Trading
Assume that we have $1,000,000 initial capital at 2019-01-01. We use the DDPG model to trade Dow jones 30 stocks.
### Set turbulence threshold
Set the turbulence threshold to be greater than the maximum of insample turbulence data, if current turbulence index is greater than the threshold, then we assume that the current market is volatile
```
data_turbulence = processed_full[(processed_full.date<'2019-01-01') & (processed_full.date>='2009-01-01')]
insample_turbulence = data_turbulence.drop_duplicates(subset=['date'])
insample_turbulence.turbulence.describe()
turbulence_threshold = np.quantile(insample_turbulence.turbulence.values,1)
turbulence_threshold
```
### Trade
DRL model needs to update periodically in order to take full advantage of the data, ideally we need to retrain our model yearly, quarterly, or monthly. We also need to tune the parameters along the way, in this notebook I only use the in-sample data from 2009-01 to 2018-12 to tune the parameters once, so there is some alpha decay here as the length of trade date extends.
Numerous hyperparameters – e.g. the learning rate, the total number of samples to train on – influence the learning process and are usually determined by testing some variations.
```
trade = data_split(processed_full, '2019-01-01','2021-01-01')
e_trade_gym = StockTradingEnv(df = trade, turbulence_threshold = 380, **env_kwargs)
# env_trade, obs_trade = e_trade_gym.get_sb_env()
trade.head()
df_account_value, df_actions = DRLAgent.DRL_prediction(
model=trained_sac,
environment = e_trade_gym)
df_account_value.shape
df_account_value.tail()
df_actions.head()
```
<a id='6'></a>
# Part 7: Backtest Our Strategy
Backtesting plays a key role in evaluating the performance of a trading strategy. Automated backtesting tool is preferred because it reduces the human error. We usually use the Quantopian pyfolio package to backtest our trading strategies. It is easy to use and consists of various individual plots that provide a comprehensive image of the performance of a trading strategy.
<a id='6.1'></a>
## 7.1 BackTestStats
pass in df_account_value, this information is stored in env class
```
print("==============Get Backtest Results===========")
now = datetime.datetime.now().strftime('%Y%m%d-%Hh%M')
perf_stats_all = backtest_stats(account_value=df_account_value)
perf_stats_all = pd.DataFrame(perf_stats_all)
perf_stats_all.to_csv("./"+config.RESULTS_DIR+"/perf_stats_all_"+now+'.csv')
#baseline stats
print("==============Get Baseline Stats===========")
baseline_df = get_baseline(
ticker="^DJI",
start = '2019-01-01',
end = '2021-01-01')
stats = backtest_stats(baseline_df, value_col_name = 'close')
```
<a id='6.2'></a>
## 7.2 BackTestPlot
```
print("==============Compare to DJIA===========")
%matplotlib inline
# S&P 500: ^GSPC
# Dow Jones Index: ^DJI
# NASDAQ 100: ^NDX
backtest_plot(df_account_value,
baseline_ticker = '^DJI',
baseline_start = '2019-01-01',
baseline_end = '2021-01-01')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/LuxusTrek/Notebooks/blob/main/1%20-%20Missing%20Data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

<hr style="margin-bottom: 40px;">
<img src="https://user-images.githubusercontent.com/7065401/39117173-a433bf6a-46e6-11e8-8a40-b4d4d6422493.jpg"
style="width:300px; float: right; margin: 0 40px 40px 40px;"></img>
# Missing Data

## Hands on!
```
import numpy as np
import pandas as pd
```
What does "missing data" mean? What is a missing value? It depends on the origin of the data and the context it was generated. For example, for a survey, a _`Salary`_ field with an empty value, or a number 0, or an invalid value (a string for example) can be considered "missing data". These concepts are related to the values that Python will consider "Falsy":
```
falsy_values = (0, False, None, '', [], {})
```
For Python, all the values above are considered "falsy":
```
any(falsy_values)
```
Numpy has a special "nullable" value for numbers which is `np.nan`. It's _NaN_: "Not a number"
```
np.nan
```
The `np.nan` value is kind of a virus. Everything that it touches becomes `np.nan`:
```
3 + np.nan
a = np.array([1, 2, 3, np.nan, np.nan, 4])
a.sum()
a.mean()
```
This is better than regular `None` values, which in the previous examples would have raised an exception:
```
3 + None
```
For a numeric array, the `None` value is replaced by `np.nan`:
```
a = np.array([1, 2, 3, np.nan, None, 4], dtype='float')
a
```
As we said, `np.nan` is like a virus. If you have any `nan` value in an array and you try to perform an operation on it, you'll get unexpected results:
```
a = np.array([1, 2, 3, np.nan, np.nan, 4])
a.mean()
a.sum()
```
Numpy also supports an "Infinite" type:
```
np.inf
```
Which also behaves as a virus:
```
3 + np.inf
np.inf / 3
np.inf / np.inf
b = np.array([1, 2, 3, np.inf, np.nan, 4], dtype=np.float)
b.sum()
```

### Checking for `nan` or `inf`
There are two functions: `np.isnan` and `np.isinf` that will perform the desired checks:
```
np.isnan(np.nan)
np.isinf(np.inf)
```
And the joint operation can be performed with `np.isfinite`.
```
np.isfinite(np.nan), np.isfinite(np.inf)
```
`np.isnan` and `np.isinf` also take arrays as inputs, and return boolean arrays as results:
```
np.isnan(np.array([1, 2, 3, np.nan, np.inf, 4]))
np.isinf(np.array([1, 2, 3, np.nan, np.inf, 4]))
np.isfinite(np.array([1, 2, 3, np.nan, np.inf, 4]))
```
_Note: It's not so common to find infinite values. From now on, we'll keep working with only `np.nan`_

### Filtering them out
Whenever you're trying to perform an operation with a Numpy array and you know there might be missing values, you'll need to filter them out before proceeding, to avoid `nan` propagation. We'll use a combination of the previous `np.isnan` + boolean arrays for this purpose:
```
a = np.array([1, 2, 3, np.nan, np.nan, 4])
a[~np.isnan(a)]
```
Which is equivalent to:
```
a[np.isfinite(a)]
```
And with that result, all the operation can be now performed:
```
a[np.isfinite(a)].sum()
a[np.isfinite(a)].mean()
```

| github_jupyter |
# Demo notebook for Kamodo Flythrough "RealFlight" function
The RealFlight function flies a real satellite trajectory through the chosen model data.
You may run the notebook as is if you have the sample data file, but you must
change the 'file_dir', 'output_name', and 'plot_output' variables in block 6 to have the correct file path.
```
#import satellite flythrough code
from kamodo_ccmc.flythrough import SatelliteFlythrough as SF
import kamodo_ccmc.flythrough.model_wrapper as MW
#The testing data file is available at https://drive.google.com/file/d/1pHx9Q8v4vO59_RUMX-SJqYv_-dE3h-st/view?usp=sharing
#What models are possible?
MW.Choose_Model('')
#Choose which model to view the example for
model = 'TIEGCM'
#What are the variable names available from that model?
MW.Model_Variables(model)
#variable name, description, variable number, coordinate type, coordinate grid, list of coordinate names, units of data
#What are the time ranges available in my data?
file_dir = 'C:/Users/rringuet/Kamodo_WinDev1/TIEGCM/Data/' #full file path to where the model output data is stored
#Change file_dir to match the file path for your data.
MW.File_Times(model, file_dir)
#This function also automatically performs any data preparation needed.
#What are the variable names available in my data?
MW.File_Variables(model, file_dir)
#variable name, description, variable number, coordinate type, coordinate grid, list of coordinate names, units of data
help(SF.RealFlight)
#Choosing input values for RealFlight function call
#----------------------------
dataset = 'grace1'
start_utcts, end_utcts = 1506039600, 1506124800
#Use https://sscweb.gsfc.nasa.gov/ to find the satellite name and time range desired
#The chosen time range should match the length of time in the model data files.
#See the times.csv file in the directory where the model data is stored for the available time ranges
#The file will appear after attempting to execute a flythrough function.
#Time values found not to be contained in the model data are automatically discarded (see output of next block).
variable_list = ['rho','u_n','T_e'] #list of desired variable names from above list.
#not all variables in the list will be available in the file(s) found.
coord_type = 'GEO' #GEO cartesian coordinates as the sample coordinate system for trajectory.
#Choose from any option available in SpacePy.
#See https://sscweb.gsfc.nasa.gov/users_guide/Appendix_C.shtml for a description of coordinate types
#choose naming convention for output files
output_type = 'csv' #chosen file format for data output
output_name = 'C:/Users/rringuet/Kamodo_NasaTest/RealFlightExample_TIEGCM' #filename for DATA output without extension
plot_output = 'C:/Users/rringuet/Kamodo_NasaTest/RealFlightExample_TIEGCM' #filename for PLOT outputs without extension
plot_coord = 'GSE' #coordinate system chosen for output plots
#run RealFlight with champ satellite trajectory
results = SF.RealFlight(dataset, start_utcts, end_utcts, model, file_dir, variable_list, coord_type,
output_type=output_type, output_name=output_name, plot_output=plot_output, plot_coord=plot_coord)
#open plots in separate internet browser window for interactivity. Nothing will open here.
```
| github_jupyter |
# 05 - Logistic Regression
by [Alejandro Correa Bahnsen](albahnsen.com/)
version 0.2, May 2016
## Part of the class [Machine Learning for Security Informatics](https://github.com/albahnsen/ML_SecurityInformatics)
This notebook is licensed under a [Creative Commons Attribution-ShareAlike 3.0 Unported License](http://creativecommons.org/licenses/by-sa/3.0/deed.en_US). Special thanks goes to [Kevin Markham](https://github.com/justmarkham)
# Review: Predicting a Continuous Response
```
import pandas as pd
import zipfile
with zipfile.ZipFile('../datasets/glass.csv.zip', 'r') as z:
f = z.open('glass.csv')
glass = pd.read_csv(f, sep=',', index_col=0)
glass.head()
```
**Question:** Pretend that we want to predict **ri**, and our only feature is **al**. How could we do it using machine learning?
**Answer:** We could frame it as a regression problem, and use a linear regression model with **al** as the only feature and **ri** as the response.
**Question:** How would we **visualize** this model?
**Answer:** Create a scatter plot with **al** on the x-axis and **ri** on the y-axis, and draw the line of best fit.
```
%matplotlib inline
import matplotlib.pyplot as plt
# scatter plot using Pandas
glass.plot(kind='scatter', x='al', y='ri')
# equivalent scatter plot using Matplotlib
plt.scatter(glass.al, glass.ri)
plt.xlabel('al')
plt.ylabel('ri')
# fit a linear regression model
from sklearn.linear_model import LinearRegression
linreg = LinearRegression()
feature_cols = ['al']
X = glass[feature_cols]
y = glass.ri
linreg.fit(X, y)
# make predictions for all values of X
glass['ri_pred'] = linreg.predict(X)
glass.head()
# put the plots together
plt.scatter(glass.al, glass.ri)
plt.plot(glass.al, glass.ri_pred, color='red')
plt.xlabel('al')
plt.ylabel('ri')
```
### Refresher: interpreting linear regression coefficients
Linear regression equation: $y = \beta_0 + \beta_1x$
```
# compute prediction for al=2 using the equation
linreg.intercept_ + linreg.coef_ * 2
# compute prediction for al=2 using the predict method
linreg.predict(2)
# examine coefficient for al
print(feature_cols, linreg.coef_)
```
**Interpretation:** A 1 unit increase in 'al' is associated with a 0.0025 unit decrease in 'ri'.
```
# increasing al by 1 (so that al=3) decreases ri by 0.0025
1.51699012 - 0.0024776063874696243
# compute prediction for al=3 using the predict method
linreg.predict(3)
```
# Predicting a Categorical Response
```
# examine glass_type
glass.glass_type.value_counts().sort_index()
# types 1, 2, 3 are window glass
# types 5, 6, 7 are household glass
glass['household'] = glass.glass_type.map({1:0, 2:0, 3:0, 5:1, 6:1, 7:1})
glass.head()
```
Let's change our task, so that we're predicting **household** using **al**. Let's visualize the relationship to figure out how to do this:
```
plt.scatter(glass.al, glass.household)
plt.xlabel('al')
plt.ylabel('household')
```
Let's draw a **regression line**, like we did before:
```
# fit a linear regression model and store the predictions
feature_cols = ['al']
X = glass[feature_cols]
y = glass.household
linreg.fit(X, y)
glass['household_pred'] = linreg.predict(X)
# scatter plot that includes the regression line
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred, color='red')
plt.xlabel('al')
plt.ylabel('household')
```
If **al=3**, what class do we predict for household? **1**
If **al=1.5**, what class do we predict for household? **0**
We predict the 0 class for **lower** values of al, and the 1 class for **higher** values of al. What's our cutoff value? Around **al=2**, because that's where the linear regression line crosses the midpoint between predicting class 0 and class 1.
Therefore, we'll say that if **household_pred >= 0.5**, we predict a class of **1**, else we predict a class of **0**.
## $$h_\beta(x) = \beta_0 + \beta_1x_1 + \beta_2x_2 + ... + \beta_nx_n$$
- $h_\beta(x)$ is the response
- $\beta_0$ is the intercept
- $\beta_1$ is the coefficient for $x_1$ (the first feature)
- $\beta_n$ is the coefficient for $x_n$ (the nth feature)
### if $h_\beta(x)\le 0.5$ then $\hat y = 0$
### if $h_\beta(x)> 0.5$ then $\hat y = 1$
```
# understanding np.where
import numpy as np
nums = np.array([5, 15, 8])
# np.where returns the first value if the condition is True, and the second value if the condition is False
np.where(nums > 10, 'big', 'small')
# transform household_pred to 1 or 0
glass['household_pred_class'] = np.where(glass.household_pred >= 0.5, 1, 0)
glass.head()
# plot the class predictions
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_class, color='red')
plt.xlabel('al')
plt.ylabel('household')
```
$h_\beta(x)$ can be lower 0 or higher than 1, which is countra intuitive
## Using Logistic Regression Instead
Logistic regression can do what we just did:
```
# fit a logistic regression model and store the class predictions
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(C=1e9)
feature_cols = ['al']
X = glass[feature_cols]
y = glass.household
logreg.fit(X, y)
glass['household_pred_class'] = logreg.predict(X)
# plot the class predictions
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_class, color='red')
plt.xlabel('al')
plt.ylabel('household')
```
What if we wanted the **predicted probabilities** instead of just the **class predictions**, to understand how confident we are in a given prediction?
```
# store the predicted probabilites of class 1
glass['household_pred_prob'] = logreg.predict_proba(X)[:, 1]
# plot the predicted probabilities
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_prob, color='red')
plt.xlabel('al')
plt.ylabel('household')
# examine some example predictions
print(logreg.predict_proba(1))
print(logreg.predict_proba(2))
print(logreg.predict_proba(3))
```
The first column indicates the predicted probability of **class 0**, and the second column indicates the predicted probability of **class 1**.
## Probability, odds, e, log, log-odds
$$probability = \frac {one\ outcome} {all\ outcomes}$$
$$odds = \frac {one\ outcome} {all\ other\ outcomes}$$
Examples:
- Dice roll of 1: probability = 1/6, odds = 1/5
- Even dice roll: probability = 3/6, odds = 3/3 = 1
- Dice roll less than 5: probability = 4/6, odds = 4/2 = 2
$$odds = \frac {probability} {1 - probability}$$
$$probability = \frac {odds} {1 + odds}$$
```
# create a table of probability versus odds
table = pd.DataFrame({'probability':[0.1, 0.2, 0.25, 0.5, 0.6, 0.8, 0.9]})
table['odds'] = table.probability/(1 - table.probability)
table
```
What is **e**? It is the base rate of growth shared by all continually growing processes:
```
# exponential function: e^1
np.exp(1)
```
What is a **(natural) log**? It gives you the time needed to reach a certain level of growth:
```
# time needed to grow 1 unit to 2.718 units
np.log(2.718)
```
It is also the **inverse** of the exponential function:
```
np.log(np.exp(5))
# add log-odds to the table
table['logodds'] = np.log(table.odds)
table
```
## What is Logistic Regression?
**Linear regression:** continuous response is modeled as a linear combination of the features:
$$y = \beta_0 + \beta_1x$$
**Logistic regression:** log-odds of a categorical response being "true" (1) is modeled as a linear combination of the features:
$$\log \left({p\over 1-p}\right) = \beta_0 + \beta_1x$$
This is called the **logit function**.
Probability is sometimes written as pi:
$$\log \left({\pi\over 1-\pi}\right) = \beta_0 + \beta_1x$$
The equation can be rearranged into the **logistic function**:
$$\pi = \frac{e^{\beta_0 + \beta_1x}} {1 + e^{\beta_0 + \beta_1x}}$$
In other words:
- Logistic regression outputs the **probabilities of a specific class**
- Those probabilities can be converted into **class predictions**
The **logistic function** has some nice properties:
- Takes on an "s" shape
- Output is bounded by 0 and 1
We have covered how this works for **binary classification problems** (two response classes). But what about **multi-class classification problems** (more than two response classes)?
- Most common solution for classification models is **"one-vs-all"** (also known as **"one-vs-rest"**): decompose the problem into multiple binary classification problems
- **Multinomial logistic regression** can solve this as a single problem
## Part 6: Interpreting Logistic Regression Coefficients
```
# plot the predicted probabilities again
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_prob, color='red')
plt.xlabel('al')
plt.ylabel('household')
# compute predicted log-odds for al=2 using the equation
logodds = logreg.intercept_ + logreg.coef_[0] * 2
logodds
# convert log-odds to odds
odds = np.exp(logodds)
odds
# convert odds to probability
prob = odds/(1 + odds)
prob
# compute predicted probability for al=2 using the predict_proba method
logreg.predict_proba(2)[:, 1]
# examine the coefficient for al
feature_cols, logreg.coef_[0]
```
**Interpretation:** A 1 unit increase in 'al' is associated with a 4.18 unit increase in the log-odds of 'household'.
```
# increasing al by 1 (so that al=3) increases the log-odds by 4.18
logodds = 0.64722323 + 4.1804038614510901
odds = np.exp(logodds)
prob = odds/(1 + odds)
prob
# compute predicted probability for al=3 using the predict_proba method
logreg.predict_proba(3)[:, 1]
```
**Bottom line:** Positive coefficients increase the log-odds of the response (and thus increase the probability), and negative coefficients decrease the log-odds of the response (and thus decrease the probability).
```
# examine the intercept
logreg.intercept_
```
**Interpretation:** For an 'al' value of 0, the log-odds of 'household' is -7.71.
```
# convert log-odds to probability
logodds = logreg.intercept_
odds = np.exp(logodds)
prob = odds/(1 + odds)
prob
```
That makes sense from the plot above, because the probability of household=1 should be very low for such a low 'al' value.

Changing the $\beta_0$ value shifts the curve **horizontally**, whereas changing the $\beta_1$ value changes the **slope** of the curve.
## Comparing Logistic Regression with Other Models
Advantages of logistic regression:
- Highly interpretable (if you remember how)
- Model training and prediction are fast
- No tuning is required (excluding regularization)
- Features don't need scaling
- Can perform well with a small number of observations
- Outputs well-calibrated predicted probabilities
Disadvantages of logistic regression:
- Presumes a linear relationship between the features and the log-odds of the response
- Performance is (generally) not competitive with the best supervised learning methods
- Can't automatically learn feature interactions
| github_jupyter |
# <img src="https://raw.githubusercontent.com/behavioral-ds/BirdSpotter/master/birdspotter_logo.png" width="200"> Birdspotter: A Tool for Analyzing and Labeling Twitter Users
### Rohit Ram [@rohitram96](https://twitter.com/rohitram96), Quyu Kong [@kaserty](https://twitter.com/kaserty), Marian-Andrei Rizoiu [@andrei_rizoiu](https://twitter.com/andrei_rizoiu)
***
<div style="display:flex; justify-content: flex-end; text-align: right;">
<img src='uts.png' width="100" alt='' style="flex:1">
<img src='https://style.anu.edu.au/_anu/4/images/logos/anu_logo_fb_350.png' width="100" alt='' style="flex:1">
<img src='https://chart.googleapis.com/chart?cht=qr&chl=https%3A%2F%2Fgithub.com%2Fbehavioral-ds%2FBirdSpotter&chs=180x180&choe=UTF-8&chld=L|2' width="100" alt='' style="flex:1">
</div>
- the primary developer of `birdspotter`.
- collaboration
- QR to the github repository.
## Problems
***
***
<div class="mytricontainer">
<div class="mydiv">
<h3> Bots <br/></h3>
<img class="myimg" src="https://static.thenounproject.com/png/1812216-200.png" width="200"/>
</div>
<div class="mydiv">
<h3> Political Leaning</h3>
<img class="myimg" src="https://static.thenounproject.com/png/2335026-200.png" width="200"/>
</div>
<div class="mydiv">
<h3> Disinformation <br/></h3>
<img class="myimg" src="https://static.thenounproject.com/png/881233-200.png" width="200"/>
</div>
</div>
- Social Media usage taking off, implications for society significant
- Advent: OSB, divergence of ideology, spread disinformation, concerning impacts on instituions
- Increasigly researchers are interested in understanding the implications of these online phenomena.
- [stakeholders] curate expansive datasets, to understand phenomena
- [problem] stakeholders face a lack of tooling to analyze such datasets
- context
- problems:
- solution:
- context
- porblem
# Birdspotter
***
<div class="mydicontainer">
<div>
<ul>
<li>
An <b>easy-to-use</b> Twitter user analysis tool
<ul>
<li>Designed for political scientists, sociologists and, data-practitioners alike</li>
<li>Models the attributes of Twitter users, and labels them</li>
<li>Prepackaged with a state-of-the-art offline bot detector</li>
<li>Quantifies influence via a Tweet Dynamics System</li>
</ul>
<li>The <b>intuitive visualization birdspotter.ml</b> for dataset exploration and narrative construction.</li>
<li>A <b>versatile label classifier</b> which can be applied to diverse aforementioned use-cases.</li>
</ul>
</div>
<div class="myimg">
<img src="auspol_teaser.png" width=900>
</div>
</div>
Birdspotter provides several contributions to the research community.
Firstly, its basic workflow is simple to operate. It comes with an offline bot detector comparable to the state of the art, and a influence quantification system.
Secondly, it comes with an accompanying visualization, birdspotter.ml, allowing dataset exploration and narrative construction.
Lastly, birdspotter is versitile and can be used for diverse use-cases which practitioners are likely interested in.
## Outline
***
* A peak under-the-hood
* A basic walkthrough for `birdspotter` workflow
* A customisation `birdspotter` to detect political leaning
In this video, we'll take you through three aspects of birdspotter.
We'll start by highlighting the features of users birdspotter focusses on.
We'll then walk-through the basic workflow for birdspotter, and show some simple visualisations
Finally, we'll show how birdspotter can be customised for another use case.
## Under the hood
<div class="mytrirowcontainer">
<div>
<h5>Twitter User Features</h5>
<div class="mydicontainer">
<div class="myimg">
<img class="myimg" src="https://cdn.iconscout.com/icon/free/png-512/twitter-241-721979.png" width="100"/>
</div>
<div style="width: 1000px">
<small>
These features are engineered directly from twitter user attributes, providing insights into the nature of the users. Features include <code>follower_count</code>, <code>years_on_twitter</code>, <code>statuses_rate</code>, ...
</small>
</div>
</div>
</div>
<div>
<h5>Semantic Features</h5>
<div class="mydicontainer">
<div>
<img class="myimg" src="https://static.thenounproject.com/png/1714867-200.png" width="100"/>
</div>
<div style="width: 1000px">
<small>
These features focus on the language of users by embedding their tweets and description with Fasttext wiki-news-300d-1M word embeddings.
</small>
</div>
</div>
</div>
<div>
<h5>Topic Features</h5>
<div class="mydicontainer">
<div>
<img class="myimg" src="https://static.thenounproject.com/png/1308644-200.png" width="100"/>
</div>
<div style="width: 1000px">
<small>
These features focus on the topics users talk about, using the TF-IDF of the 1,000 most frequent hashtags.
</small>
</div>
</div>
</div>
<div>
<h5>Retweet Dynamics</h5>
<div class="mydicontainer">
<div>
<img class="myimg" src="https://static.thenounproject.com/png/635941-200.png" width="100"/>
</div>
<div style="width: 1000px">
<small>
These features use the timing and follower counts of tweets/users, to reconstruct an expected retweet diffusion of a cascade.
</small>
</div>
</div>
</div>
</div>
Birdspotter has two main systems; it's labeller and it influence quantification.
These first three feature sets are engineered to provide informative representations of users for the labeller.
The Twitter User Features, are engineered directly from twitter user attributes to provide information about a users status and behaviour on the platform. Constructed features include the `statuses_rate`, `years_on_twitter`, etc. If `statuses_rate` is high this might indicate a bot. If `year_on_twitter` are low, this might indicate a sock puppet account.
The Semantic features are engineered to focus on the language users use. We embed a users tweets and descriptions using the Fasttext word2vec embeddings, to provide a indication of a users language profile.
The Topic features relate to the hashtags that users use, and conseuqnetly the topics they talk about. We use the TF-IDF of the 1,000 most frequent hashtags in a dataset, to represent this.
Finally, the influence quantification system, uses the timing and follower_counts of users to infer if a retweet is a direct retweet of another. This is used to quantify the influence of users.
### Bot Detector Performance
***
<img class="myimg" src="ablation_1.svg"/>
### Bot Detector Performance
***
<img class="myimg" src="ablation_2.svg"/>
### Bot Detector Performance
***
<img class="myimg" src="ablation_3.svg"/>
* Ablation shows higher scores, but we keep all features for feature diversity for broader use-cases
## Influence Quantification
***
<img class="myimg" src="diffusion.PNG"/>
* Compute influence as number of descendants
* {Point to two examples}
* Problem: This structure of tweets and retweets isn't provide
* Solution: We reconstruct it
```
import seaborn as sns
import matplotlib.pyplot as plt
from birdspotter import BirdSpotter
import pandas as pd
from tqdm import tqdm
import json
import pickle as pk
import numpy as np
import os
sns.set(rc={'figure.figsize':(11.7,8.27)})
```
## Basic Usage
#### Installation
We can now walk-through a simple workflow for birdspotter.
Firstly, birdspotter is provided through the pypi package manager, and can be installed with `pip install birdspotter`
#### Basic Workflow
```
# We got the tweets from the following the github repository https://github.com/echen102/COVID-19-TweetIDs
# We used their instructions to rehydrate the tweets from 31/01/2020 and placed it into covid19.jsonl
from birdspotter import BirdSpotter ## Import birdspotter
bs = BirdSpotter('covid19.jsonl') ## Extracts the tweets from the raw jsonl [https://github.com/echen102/COVID-19-TweetIDs]
bs.getLabeledUsers() ## Uses the default bot labeller and influence quantification systems
bs.getCascadesDataFrame() ## Formats the retweet cascades, such that expected retweet structures can extracted
bs.featureDataframe[['botness', 'influence']]
fig, ax =plt.subplots(1,2)
botness_dist = sns.histplot(data=bs.featureDataframe, x="botness", ax=ax[0])
influence_eccdf = sns.ecdfplot(data=bs.featureDataframe, x="influence", complementary=True, ax=ax[1]).set(xscale="log", yscale="log")
fig.show()
```
* botness shows several humans and bots
* influence shows long-tail signature, a.k.a. rich-gets-richer, where a minority hold a significant influence
[birdspotter.ml](http://birdspotter.ml/?uid=20986375)
- explain the plot
- explain what the plot shows
- explore the data
- show how the explored data fits a narrative
bot: http://localhost:8050/?uid=1092530860618907653
## Customising `birdspotter` for broader use-cases
- the complex nature of online phenomena, demand diverse analyses
- researchers desirce view data different lenses
- show blah
- show use public dataset, detect political leaning
1. We hydrate some tweets from the Twitter Parlimentarian Database
2. We filter the tweets to include only **Australian Politicians**.
3. We **label right-wing partied politicians positively**, and others negatively (with `bs_pol.getBotAnnotationTemplate` for example)
4. We **retrain `birdspotter`** with these new labels and label all users (i.e. including users the politicians retweeted) using the new model
## Customising `birdspotter` for broader use-cases
```
# This is the guts of the code; it does what is described above
politicians = pd.read_csv('./full_member_info.csv', encoding='utf16')
politicians_aus = politicians[politicians['country'] == 'Australia']
politicians_aus_available = politicians_aus[~politicians_aus['uid'].isnull()]
def classify_party(party_id):
mapping = {
464 : 1, # Liberal Party of Australia
465 : -1, # Australian Labor Party
467 : 1, # The Nationals
468 : 0, # Nick Xenophon Team
469 : -1, # Australian Greens
471 : np.nan,
475 : 1, # Katter's Australian Party
}
return mapping[party_id]
politicians_aus_available['isright'] = politicians_aus_available['party_id'].apply(classify_party)
politicians_aus_available['user_id'] = politicians_aus_available['uid'].astype(int).astype(str)
politicians_aus_available = politicians_aus_available.set_index('user_id')
with open('./tweets.jsonl', 'r') as rf, open('./aus_tweets.jsonl', 'w') as wf:
for line in tqdm(rf):
try:
j = json.loads(line)
if j['user']['id_str'] in politicians_aus_available['uid'].astype(int).astype(str).values:
wf.write(json.dumps(j) + '\n')
except Exception as e:
print(j)
print(e)
break
bs = BirdSpotter('aus_tweets.jsonl')
bs.getLabeledUsers()
bs.getCascadesDataFrame()
with open('bs_aus_module.pk', 'wb') as wf:
pk.dump(bs,wf, protocol=4)
bs.featureDataframe['isright'] = politicians_aus_available['isright']
ground_truth = bs.featureDataframe[~bs.featureDataframe['isright'].isnull()][['isright']]
ground_truth['isbot'] = ground_truth['isright'] == 1
ground_truth = ground_truth[~ground_truth.index.duplicated()]
data = bs.featureDataframe.copy()[bs.featureDataframe.index.isin(ground_truth.index)]
data = data[~data.index.duplicated()]
del data['isright']
del data['botness']
del data['influence']
del data['cascade_membership']
data = data[list(data.columns[data.dtypes != 'object'])]
data['isbot'] = ground_truth['isbot'].loc[data.index]
with open('pol_training_data.pickle', 'wb') as wf:
pk.dump(data,wf, protocol=4)
from birdspotter import BirdSpotter
import pickle as pk
# bs_pol = BirdSpotter('aus_tweets.jsonl')
with open('bs_aus_module.pk', 'rb') as rf:
bs_pol = pk.load(rf)
print("Loaded module")
bs_pol.trainClassifierModel('pol_training_data.pickle')
print("finished training")
del bs_pol.featureDataframe['botness']
print("removed botness column")
bs_pol.getBotness()
bs_pol.getLabeledUsers()
print("got labels")
with open('pol_booster.pickle', 'wb') as wf:
pk.dump(bs_pol.booster, wf, protocol=4)
print("pickled booster")
with open('aus_pol_bs_module.pickle', 'wb') as wf:
pk.dump(bs_pol, wf, protocol=4)
with open('pol_booster.pickle', 'wb') as wf:
pk.dump(bs.booster, wf, protocol=4)
import pickle as pk
with open('aus_pol_bs_module.pickle', 'rb') as rf:
bs_pol = pk.load(rf)
bs_pol = BirdSpotter('aus_tweets.jsonl')
# bs_pol.trainClassifierModel('pol_training_data.pickle')
bs_pol.loadPickledBooster('pol_booster.pickle')
bs_pol.getLabeledUsers()
```
## Results
***
On this limited dataset, a 10-fold CV of <code>birdspotter</code> garners an average AUC (Area under ROC) of 0.986.
##### Lefties
<div class="mytricontainer">
<div class="mydiv">
<h5>Katharine Murphy</h5>
<span> Score = 0.231608</span>
<div class="myimg">
<img class="myimg" src="https://pbs.twimg.com/profile_images/687958069733494784/1SUBcHBJ_400x400.jpg" width="200"/>
</div>
<span>Guardian Journalist (a left-leaning news organisation).</span>
</div>
<div class="mydiv">
<h5>Josh Taylor</h5>
<span> Score = 0.232707</span>
<img class="myimg" src="https://pbs.twimg.com/profile_images/1255254018147987457/vuvJbsoJ_400x400.jpg" width="200"/>
<span>Guardian Reporter (a left-leaning news organisation).</span>
</div>
<div class="mydiv">
<h5>Shorten Suite</h5>
<span> Score = 0.237657</span>
<img class="myimg" src="https://pbs.twimg.com/profile_images/882886962754895873/Ya_ZZwIV_400x400.jpg" width="200"/>
<span>Tweets memes supporting Bill Shorten (Former ALP Leader)</span>
</div>
</div>
## Results
***
On this limited dataset, a 10-fold CV of <code>birdspotter</code> garners an average AUC (Area under ROC) of 0.986.
##### Righties
<div class="mytricontainer">
<div class="mydiv">
<h5>Gavan Macrides</h5>
<span> Score = 0.807822</span>
<div class="myimg">
<img class="myimg" src="https://pbs.twimg.com/profile_images/851294256547352576/_bYWJQwa_400x400.jpg" width="200"/>
</div>
<span>Former LNP Campaign Director</span>
</div>
<div class="mydiv">
<h5>David O'Brien</h5>
<span> Score = 0.801732</span>
<img class="myimg" src="https://pbs.twimg.com/profile_images/1134760668911902721/FSNPNM7s_400x400.png" width="200"/>
<span>Tweets about and supports conservative politicians, particularly the Nationals.</span>
</div>
<div class="mydiv">
<h5>Minerals Council Australia</h5>
<span> Score = 0.797240</span>
<img class="myimg" src="https://pbs.twimg.com/profile_images/595401078796980224/M1ZCxkuo_400x400.jpg" width="200"/>
<span>A group presenting the interests of the mining industry, who're typically right leaning.</span>
</div>
</div>
```
bs_pol.featureDataframe.sort_values('botness').iloc[[42, 45, 50, 7693 - 7 - 1,7693 - 28 - 1, 7693 - 67 - 1]]
```
## Conclusion
***
* We present a simple, versatile, and powerful tool, `birdspotter`, for analyzing Twitter users
* It has an extremely simple basic workflow, utilising many of the curated datasets practioners likely have.
* It can be customized for the broad-range of use cases practioners are likely interested in.
<small>All code for this presentation can be found here: <img src='https://chart.googleapis.com/chart?cht=qr&chl=https%3A%2F%2Fgithub.com%2Fbehavioral-ds%2FBirdSpotter&chs=180x180&choe=UTF-8&chld=L|2' width="300" alt='' style="flex:1"></small>
<h1 style="color:#c9245d; text-align:center">Thank You!</h1>
#### References
* van Vliet, L., Törnberg, P., & Uitermark, J. (2020). The Twitter parliamentarian database: Analyzing Twitter politics across 26 countries. PloS one, 15(9), e0237073.
* Rizoiu, M. A., Graham, T., Zhang, R., Zhang, Y., Ackland, R., & Xie, L. (2018, June). # debatenight: The role and influence of socialbots on twitter during the 1st 2016 us presidential debate. In Proceedings of the International AAAI Conference on Web and Social Media (Vol. 12, No. 1).
#### Acknowledgements
* SIGIR Travel Grant
<div style="display:flex; justify-content: flex-end; text-align: right;">
<img src='uts.png' width="100" alt='' style="flex:1">
<img src='https://style.anu.edu.au/_anu/4/images/logos/anu_logo_fb_350.png' width="100" alt='' style="flex:1">
<img src='https://chart.googleapis.com/chart?cht=qr&chl=https%3A%2F%2Fgithub.com%2Fbehavioral-ds%2FBirdSpotter&chs=180x180&choe=UTF-8&chld=L|2' width="100" alt='' style="flex:1">
</div>
| github_jupyter |
```
import ipyvuetify as v
```
First histogram plot
```
import ipywidgets as widgets
import numpy as np
from bqplot import pyplot as plt
import bqplot
n = 200
x = np.linspace(0.0, 10.0, n)
y = np.cumsum(np.random.randn(n)*10).astype(int)
fig = plt.figure( title='Histogram')
np.random.seed(0)
hist = plt.hist(y, bins=25)
hist.scales['sample'].min = float(y.min())
hist.scales['sample'].max = float(y.max())
fig.layout.width = 'auto'
fig.layout.height = 'auto'
fig.layout.min_height = '300px' # so it shows nicely in the notebook
fig
slider = v.Slider(thumb_label='always', class_="px-4", v_model=30)
widgets.link((slider, 'v_model'), (hist, 'bins'))
slider
```
Line chart
```
fig2 = plt.figure( title='Line Chart')
np.random.seed(0)
p = plt.plot(x, y)
fig2.layout.width = 'auto'
fig2.layout.height = 'auto'
fig2.layout.min_height = '300px' # so it shows nicely in the notebook
fig2
brushintsel = bqplot.interacts.BrushIntervalSelector(scale=p.scales['x'])
def update_range(*args):
if brushintsel.selected is not None and brushintsel.selected.shape == (2,):
mask = (x > brushintsel.selected[0]) & (x < brushintsel.selected[1])
hist.sample = y[mask]
brushintsel.observe(update_range, 'selected')
fig2.interaction = brushintsel
```
Second histogram plot
```
n2 = 200
x2 = np.linspace(0.0, 10.0, n)
y2 = np.cumsum(np.random.randn(n)*10).astype(int)
figHist2 = plt.figure( title='Histogram 2')
np.random.seed(0)
hist2 = plt.hist(y2, bins=25)
hist2.scales['sample'].min = float(y2.min())
hist2.scales['sample'].max = float(y2.max())
figHist2.layout.width = 'auto'
figHist2.layout.height = 'auto'
figHist2.layout.min_height = '300px' # so it shows nicely in the notebook
sliderHist2 = v.Slider(_metadata={'mount_id': 'histogram_bins2'}, thumb_label='always', class_='px-4', v_model=5)
from traitlets import link
link((sliderHist2, 'v_model'), (hist2, 'bins'))
display(figHist2)
display(sliderHist2)
```
Set up voila vuetify layout
The voila vuetify template does not render output from the notebook, it only shows widget with the mount_id metadata.
```
v.Tabs(_metadata={'mount_id': 'content-main'}, children=[
v.Tab(children=['Tab1']),
v.Tab(children=['Tab2']),
v.TabItem(children=[
v.Layout(row=True, wrap=True, align_center=True, children=[
v.Flex(xs12=True, lg6=True, xl4=True, children=[
fig, slider
]),
v.Flex(xs12=True, lg6=True, xl4=True, children=[
figHist2, sliderHist2
]),
v.Flex(xs12=True, xl4=True, children=[
fig2
]),
])
]),
v.TabItem(children=[
v.Container(children=['Lorum ipsum'])
])
])
import ipywidgets as widgets
import ipyvuetify as v
buttons = [widgets.Button(description=f'button {i}') for i in range(4)]
v.Row(_metadata={'mount_id': 'content'}, children=buttons)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_style("whitegrid")
```
# Model: random forrest
```
from sklearn.metrics import classification_report, confusion_matrix, plot_confusion_matrix, roc_auc_score, roc_curve, precision_recall_curve
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
```
## Data
Load the dataset, applying no major transformations to it.
```
data = pd.read_csv('../dataset/creditcard.csv')
data.head()
X = data.drop(columns=['Class'])
y = data['Class']
```
Since the data is largely unbalanced we must use a stratified sampling to make sure we get both negative and positive samples to train with.
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=0, stratify=y)
```
## Pipeline (build)
```
numeric_feature_indexes = slice(0, 30)
preprocessor = ColumnTransformer(
transformers=[
('num', StandardScaler(), numeric_feature_indexes),
])
pipeline = Pipeline([
('preprocessor', preprocessor),
('classifier', RandomForestClassifier())
])
num_features_type_map = {feature: 'float64' for feature in X_train.columns[numeric_feature_indexes]}
X_train = X_train.astype(num_features_type_map)
X_test = X_test.astype(num_features_type_map)
```
## Pipeline (train)
```
model = pipeline.fit(X_train, y_train)
model
```
## Pipeline (evaluate)
```
y_pred = model.predict(X_test)
print(classification_report(y_test, y_pred))
disp = plot_confusion_matrix(model, X_test, y_test, display_labels=['normal', 'fraudulent'], cmap=plt.cm.Blues)
disp.ax_.grid(False)
```
Some great material is available here: https://machinelearningmastery.com/roc-curves-and-precision-recall-curves-for-classification-in-python/
```
y_pred_proba = pipeline.predict_proba(X_test)[::,1]
fpr, tpr, _ = roc_curve(y_test, y_pred_proba)
auc = roc_auc_score(y_test, y_pred_proba)
fig, ax = plt.subplots(figsize=(5,5))
ax.plot(fpr,tpr,label=f"auc {auc:2.2f}")
ax.legend(loc=4)
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate');
precision, recall, _ = precision_recall_curve(y_test, y_pred_proba)
fig, ax = plt.subplots(figsize=(5,5))
no_skill = len(y_test[y_test==1]) / len(y_test)
ax.plot([0, 1], [no_skill, no_skill], linestyle='--', label='No Skill')
ax.plot(recall, precision)
ax.set_xlabel('Precision')
ax.set_ylabel('Recall');
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
#@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Bayesian Gaussian Mixture Model and Hamiltonian MCMC
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/probability/blob/master/tensorflow_probability/examples/jupyter_notebooks/Bayesian_Gaussian_Mixture_Model.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/probability/blob/master/tensorflow_probability/examples/jupyter_notebooks/Bayesian_Gaussian_Mixture_Model.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
In this colab we'll explore sampling from the posterior of a Bayesian Gaussian Mixture Model (BGMM) using only TensorFlow Probability primitives.
## Model
For $k\in\{1,\ldots, K\}$ mixture components each of dimension $D$, we'd like to model $i\in\{1,\ldots,N\}$ iid samples using the following Bayesian Gaussian Mixture Model:
$$\begin{align*}
\theta &\sim \text{Dirichlet}(\text{concentration}=\alpha_0)\\
\mu_k &\sim \text{Normal}(\text{loc}=\mu_{0k}, \text{scale}=I_D)\\
T_k &\sim \text{Wishart}(\text{df}=5, \text{scale}=I_D)\\
Z_i &\sim \text{Categorical}(\text{probs}=\theta)\\
Y_i &\sim \text{Normal}(\text{loc}=\mu_{z_i}, \text{scale}=T_{z_i}^{-1/2})\\
\end{align*}$$
Note, the `scale` arguments all have `cholesky` semantics. We use this convention because it is that of TF Distributions (which itself uses this convention in part because it is computationally advantageous).
Our goal is to generate samples from the posterior:
$$p\left(\theta, \{\mu_k, T_k\}_{k=1}^K \Big| \{y_i\}_{i=1}^N, \alpha_0, \{\mu_{ok}\}_{k=1}^K\right)$$
Notice that $\{Z_i\}_{i=1}^N$ is not present--we're interested in only those random variables which don't scale with $N$. (And luckily there's a TF distribution which handles marginalizing out $Z_i$.)
It is not possible to directly sample from this distribution owing to a computationally intractable normalization term.
[Metropolis-Hastings algorithms](https://en.wikipedia.org/wiki/Metropolis%E2%80%93Hastings_algorithm) are technique for for sampling from intractable-to-normalize distributions.
TensorFlow Probability offers a number of MCMC options, including several based on Metropolis-Hastings. In this notebook, we'll use [Hamiltonian Monte Carlo](https://en.wikipedia.org/wiki/Hamiltonian_Monte_Carlo) (`tfp.mcmc.HamiltonianMonteCarlo`). HMC is often a good choice because it can converge rapidly, samples the state space jointly (as opposed to coordinatewise), and leverages one of TF's virtues: automatic differentiation. That said, sampling from a BGMM posterior might actually be better done by other approaches, e.g., [Gibb's sampling](https://en.wikipedia.org/wiki/Gibbs_sampling).
```
%matplotlib inline
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import functools
import matplotlib.pyplot as plt; plt.style.use('ggplot')
import numpy as np
import seaborn as sns; sns.set_context('notebook')
import tensorflow as tf
import tensorflow_probability as tfp
tfd = tfp.distributions
tfb = tfp.bijectors
def session_options(enable_gpu_ram_resizing=True):
"""Convenience function which sets common `tf.Session` options."""
config = tf.ConfigProto()
config.log_device_placement = True
if enable_gpu_ram_resizing:
# `allow_growth=True` makes it possible to connect multiple colabs to your
# GPU. Otherwise the colab malloc's all GPU ram.
config.gpu_options.allow_growth = True
return config
def reset_sess(config=None):
"""Convenience function to create the TF graph and session, or reset them."""
if config is None:
config = session_options()
tf.reset_default_graph()
global sess
try:
sess.close()
except:
pass
sess = tf.InteractiveSession(config=config)
reset_sess()
```
Before actually building the model, we'll need to define a new type of distribution. From the model specification above, its clear we're parameterizing the MVN with an inverse covariance matrix, i.e., [precision matrix](https://en.wikipedia.org/wiki/Precision_(statistics%29). To accomplish this in TF, we'll need to roll out our `Bijector`. This `Bijector` will use the forward transformation:
- `Y =` [`tf.matrix_triangular_solve`](https://www.tensorflow.org/api_docs/python/tf/matrix_triangular_solve)`(tf.matrix_transpose(chol_precision_tril), X, adjoint=True) + loc`.
And the `log_prob` calculation is just the inverse, i.e.:
- `X =` [`tf.matmul`](https://www.tensorflow.org/api_docs/python/tf/matmul)`(chol_precision_tril, X - loc, adjoint_a=True)`.
Since all we need for HMC is `log_prob`, this means we avoid ever calling `tf.matrix_triangular_solve` (as would be the case for `tfd.MultivariateNormalTriL`). This is advantageous since `tf.matmul` is usually faster owing to better cache locality.
```
class MVNCholPrecisionTriL(tfd.TransformedDistribution):
"""MVN from loc and (Cholesky) precision matrix."""
def __init__(self, loc, chol_precision_tril, name=None):
super(MVNCholPrecisionTriL, self).__init__(
distribution=tfd.Independent(tfd.Normal(tf.zeros_like(loc),
scale=tf.ones_like(loc)),
reinterpreted_batch_ndims=1),
bijector=tfb.Chain([
tfb.Affine(shift=loc),
tfb.Invert(tfb.Affine(scale_tril=chol_precision_tril,
adjoint=True)),
]),
name=name)
```
The `tfd.Independent` distribution turns independent draws of one distribution, into a multivariate distribution with statistically independent coordinates. In terms of computing `log_prob`, this "meta-distribution" manifests as a simple sum over the event dimension(s).
Also notice that we took the `adjoint` ("transpose") of the scale matrix. This is because if precision is inverse covariance, i.e., $P=C^{-1}$ and if $C=AA^\top$, then $P=BB^{\top}$ where $B=A^{-\top}$.
Since this distribution is kind of tricky, let's quickly verify that our `MVNCholPrecisionTriL` works as we think it should.
```
def compute_sample_stats(d, seed=42, n=int(1e6)):
x = d.sample(n, seed=seed)
sample_mean = tf.reduce_mean(x, axis=0, keepdims=True)
s = x - sample_mean
sample_cov = tf.matmul(s, s, adjoint_a=True) / tf.cast(n, s.dtype)
sample_scale = tf.cholesky(sample_cov)
sample_mean = sample_mean[0]
return [
sample_mean,
sample_cov,
sample_scale,
]
dtype = np.float32
true_loc = np.array([1., -1.], dtype=dtype)
true_chol_precision = np.array([[1., 0.],
[2., 8.]],
dtype=dtype)
true_precision = np.matmul(true_chol_precision, true_chol_precision.T)
true_cov = np.linalg.inv(true_precision)
d = MVNCholPrecisionTriL(
loc=true_loc,
chol_precision_tril=true_chol_precision)
[
sample_mean_,
sample_cov_,
sample_scale_,
] = sess.run(compute_sample_stats(d))
print('true mean:', true_loc)
print('sample mean:', sample_mean_)
print('true cov:\n', true_cov)
print('sample cov:\n', sample_cov_)
```
Since the sample mean and covariance are close to the true mean and covariance, it seems like the distribution is correctly implemented. Now, we'll use `MVNCholPrecisionTriL` and stock`tfp.distributions` to specify the BGMM prior random variables:
```
dtype = np.float32
dims = 2
components = 3
rv_mix_probs = tfd.Dirichlet(
concentration=np.ones(components, dtype) / 10.,
name='rv_mix_probs')
rv_loc = tfd.Independent(
tfd.Normal(
loc=np.stack([
-np.ones(dims, dtype),
np.zeros(dims, dtype),
np.ones(dims, dtype),
]),
scale=tf.ones([components, dims], dtype)),
reinterpreted_batch_ndims=1,
name='rv_loc')
rv_precision = tfd.WishartTriL(
df=5,
scale_tril=np.stack([np.eye(dims, dtype=dtype)]*components),
input_output_cholesky=True,
name='rv_precision')
print(rv_mix_probs)
print(rv_loc)
print(rv_precision)
```
Using the three random variables defined above, we can now specify the joint log probability function. To do this we'll use `tfd.MixtureSameFamily` to automatically integrate out the categorical $\{Z_i\}_{i=1}^N$ draws.
```
def joint_log_prob(observations, mix_probs, loc, chol_precision):
"""BGMM with priors: loc=Normal, precision=Inverse-Wishart, mix=Dirichlet.
Args:
observations: `[n, d]`-shaped `Tensor` representing Bayesian Gaussian
Mixture model draws. Each sample is a length-`d` vector.
mix_probs: `[K]`-shaped `Tensor` representing random draw from
`SoftmaxInverse(Dirichlet)` prior.
loc: `[K, d]`-shaped `Tensor` representing the location parameter of the
`K` components.
chol_precision: `[K, d, d]`-shaped `Tensor` representing `K` lower
triangular `cholesky(Precision)` matrices, each being sampled from
a Wishart distribution.
Returns:
log_prob: `Tensor` representing joint log-density over all inputs.
"""
rv_observations = tfd.MixtureSameFamily(
mixture_distribution=tfd.Categorical(probs=mix_probs),
components_distribution=MVNCholPrecisionTriL(
loc=loc,
chol_precision_tril=chol_precision))
log_prob_parts = [
rv_observations.log_prob(observations), # Sum over samples.
rv_mix_probs.log_prob(mix_probs)[..., tf.newaxis],
rv_loc.log_prob(loc), # Sum over components.
rv_precision.log_prob(chol_precision), # Sum over components.
]
sum_log_prob = tf.reduce_sum(tf.concat(log_prob_parts, axis=-1), axis=-1)
# Note: for easy debugging, uncomment the following:
# sum_log_prob = tf.Print(sum_log_prob, log_prob_parts)
return sum_log_prob
```
Notice that this function internally defines a new random variable. This is necessary since the `observations` RV depends on samples from the RVs defined further above.
## Generate "Training" Data
For this demo, we'll sample some random data.
```
num_samples = 1000
true_loc = np.array([[-2, -2],
[0, 0],
[2, 2]], dtype)
random = np.random.RandomState(seed=42)
true_hidden_component = random.randint(0, components, num_samples)
observations = (true_loc[true_hidden_component] +
random.randn(num_samples, dims).astype(dtype))
```
## Bayesian Inference using HMC
Now that we've used TFD to specify our model and obtained some observed data, we have all the necessary pieces to run HMC.
To do this, we'll use a [partial application](https://en.wikipedia.org/wiki/Partial_application) to "pin down" the things we don't want to sample. In this case that means we need only pin down `observations`. (The hyper-parameters are already baked in to the prior distributions and not part of the `joint_log_prob` function signature.)
```
unnormalized_posterior_log_prob = functools.partial(joint_log_prob, observations)
initial_state = [
tf.fill([components],
value=np.array(1. / components, dtype),
name='mix_probs'),
tf.constant(np.array([[-2, -2],
[0, 0],
[2, 2]], dtype),
name='loc'),
tf.eye(dims, batch_shape=[components], dtype=dtype, name='chol_precision'),
]
```
### Unconstrained Representation
Hamiltonian Monte Carlo (HMC) requires the target log-probability function be differentiable with respect to its arguments. Furthermore, HMC can exhibit dramatically higher statistical efficiency if the state-space is unconstrained.
This means we'll have to work out two main issues when sampling from the BGMM posterior:
1. $\theta$ represents a discrete probability vector, i.e., must be such that $\sum_{k=1}^K \theta_k = 1$ and $\theta_k>0$.
2. $T_k$ represents an inverse covariance matrix, i.e., must be such that $T_k \succ 0$, i.e., is [positive definite](https://en.wikipedia.org/wiki/Positive-definite_matrix).
To address this requirement we'll need to:
1. transform the constrained variables to an unconstrained space
2. run the MCMC in unconstrained space
3. transform the unconstrained variables back to the constrained space.
As with `MVNCholPrecisionTriL`, we'll use [`Bijector`s](https://www.tensorflow.org/api_docs/python/tf/distributions/bijectors/Bijector) to transform random variables to unconstrained space.
- The [`Dirichlet`](https://en.wikipedia.org/wiki/Dirichlet_distribution) is transformed to unconstrained space via the [softmax function](https://en.wikipedia.org/wiki/Softmax_function).
- Our precision random variable is a distribution over postive semidefinite matrices. To unconstrain these we'll use the `FillTriangular` and `TransformDiagonal` bijectors. These convert vectors to lower-triangular matrices and ensure the diagonal is positive. The former is useful because it enables sampling only $d(d+1)/2$ floats rather than $d^2$.
```
unconstraining_bijectors = [
tfb.SoftmaxCentered(),
tfb.Identity(),
tfb.Chain([
tfb.TransformDiagonal(tfb.Softplus()),
tfb.FillTriangular(),
])]
[mix_probs, loc, chol_precision], kernel_results = tfp.mcmc.sample_chain(
num_results=2000,
num_burnin_steps=500,
current_state=initial_state,
kernel=tfp.mcmc.TransformedTransitionKernel(
inner_kernel=tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=unnormalized_posterior_log_prob,
step_size=0.065,
num_leapfrog_steps=5),
bijector=unconstraining_bijectors))
acceptance_rate = tf.reduce_mean(tf.to_float(kernel_results.inner_results.is_accepted))
mean_mix_probs = tf.reduce_mean(mix_probs, axis=0)
mean_loc = tf.reduce_mean(loc, axis=0)
mean_chol_precision = tf.reduce_mean(chol_precision, axis=0)
```
Note: through trial-and-error we've predetermined the `step_size` and `num_leapfrog_steps` to approximately achieve an [asymptotically optimal rate of 0.651](https://arxiv.org/abs/1001.4460). For a technique to do this automatically, see the examples section in `help(tfp.mcmc.HamiltonianMonteCarlo)`.
We'll now execute the chain and print the posterior means.
```
[
acceptance_rate_,
mean_mix_probs_,
mean_loc_,
mean_chol_precision_,
mix_probs_,
loc_,
chol_precision_,
] = sess.run([
acceptance_rate,
mean_mix_probs,
mean_loc,
mean_chol_precision,
mix_probs,
loc,
chol_precision,
])
print(' acceptance_rate:', acceptance_rate_)
print(' avg mix probs:', mean_mix_probs_)
print('\n avg loc:\n', mean_loc_)
print('\navg chol(precision):\n', mean_chol_precision_)
ax = sns.kdeplot(loc_[:,0,0], loc_[:,0,1], shade=True)
ax = sns.kdeplot(loc_[:,1,0], loc_[:,1,1], shade=True)
ax = sns.kdeplot(loc_[:,2,0], loc_[:,2,1], shade=True)
plt.title('KDE of loc draws');
```
## Conclusion
This simple colab demonstrated how TensorFlow Probability primitives can be used to build hierarchical Bayesian mixture models.
| github_jupyter |
# Revision of Functions in Scripts
## Overview:
- **Teaching:** 15 min
- **Exercises:** 15 min
**Questions**
- How can we put our new assertions into a scipt that we can run or import?
**Objectives**
- Revise how to define a function in a script.
- Recall how to use docstrings to auto-document your functions.
- Know how to call the function within a script.
- Know how to make a script importable.
In the previous lesson we wrote our function with its assertion in `notebooks`. As we have discussed previously, it is much more useful to write these as a script which we can then call, or in the case of libraries import. In this lesson we will write our mean function as a script and revise how to display documentation and call the function. We will then explore how to include our own libraries within other scripts revise why it is good practice to include the main part of the script in its own function.
If you have not already created a directory `my_testing` in your `intro-testing` directory, do so and `cd` into it. Then create a new file called `mean.py`. You are free to use the editor of your choice, e.g. `nano` or others on `linux`, or use the notebooks text editor.
Note that this will not be available on remote linux machines so you should be confident to use editors like `nano` if required.
## Writing Scripts
Edit your new file `mean.py` so that it reads:
```python
def mean(sample):
'''
Takes a list of numbers, sample
and returns the mean.
'''
assert len(sample) != 0, "Unable to take the mean of an empty list"
for value in sample:
assert isinstance(value,int) or isinstance(value,float), "Value in list is not a number."
sample_mean = sum(sample) / len(sample)
return sample_mean
numbers = [1, 2, 3, 4, 5]
print( mean(numbers) )
no_numbers = []
print( mean(no_numbers) )
word_and_numbers = [1, 2, 3, 4, "apple"]
print( mean(word_and_numbers) )
```
Finally we can execute our script, you can do this form the terminal but we will run from within notebooks as we did before. in your library, navigate to the folder containing your `mean.py` file and luanch a new notebook:
```python
%run mean.py
```
You should see an output that looks like:
```brainfuck
Traceback (most recent call last):
File "./mean.py", line 22, in <module>
mean(no_numbers)
File "./mean.py", line 10, in mean
assert len(sample) != 0, "Unable to take the mean of an empty list"
AssertionError: Unable to take the mean of an empty list
```
Our scripts executes each of line of code in turn and stops when it reaches the first `AssertionError`, even though this is how we intend the program to execture, Python doesn't know this and it means that it does not yet run the third of our tests. We will learn how we can run multiple tests, and encapsulate them in successful tests even if they intentionally raise errors.
Now we want to explore our script in a little more detail. Remember in our previous lesson we `import`ed libraries into our interactive session. Let's see what happens when we do this with our new script. Let's try to import `mean.py` as a library:
```python
import mean
3.0
---------------------------------------------------------------------------
AssertionError Traceback (most recent call last)
<ipython-input-1-330bb4995352> in <module>
----> 1 import mean
/home/rjg20/training/arc-training/intro-testing/mean.py in <module>
20 no_numbers = []
21
---> 22 print( mean(no_numbers) )
23
24 word_and_numbers = [1, 2, 3, 4, "apple"]
/home/rjg20/training/arc-training/intro-testing/mean.py in mean(sample)
8 '''
9
---> 10 assert len(sample) != 0, "Unable to take the mean of an empty list"
11 for value in sample:
12 assert isinstance(value,int) or isinstance(value,float), "Value in list is not a number."
AssertionError: Unable to take the mean of an empty list
```
## The Scope of a Script
What we would like to do is change the behaviour of our program so that the functions are available when it is imported, but the remainder of the scripts only executes when we run it from the command line. This is called limiting the scope of the script, open your script and edit it so that it reads:
```python
def mean(sample):
'''
Takes a list of numbers, sample
and returns the mean.
'''
assert len(sample) != 0, "Unable to take the mean of an empty list"
for value in sample:
assert isinstance(value,int) or isinstance(value,float), "Value in list is not a number."
sample_mean = sum(sample) / len(sample)
return sample_mean
def main():
numbers = [1, 2, 3, 4, 5]
print( mean(numbers) )
no_numbers = []
print( mean(no_numbers) )
word_and_numbers = [1, 2, 3, 4, "apple"]
print( mean(word_and_numbers) )
if __name__ == '__main__':
main()
```
Now run this as a script and also try importing it in ipython. `__name__` determines the scope of a script, if it is run as a script it is set to `__main__`, but not when it is imported, allowing the behaviour to be different in the two cases. You can find out more [here](https://docs.python.org/3/library/__main__.html#module-__main__).
## Exercise: Update the documentation
We have some documentation in the `mean` function but this could be improved to give more details on the functions behaviour, and there is no documentation for our main function.
Add appropriate documentation and verify that it works correctly.
[Solution]()
## Solution: Update the documentation
```python
def mean(num_list):
'''
Function to calculate the mean of a list of numbers
Usage: mean(list_of_numbers)
Checks that length of list is not 0, else raises assertion error.
Checks that all items are ints of floats, else raises assertion error.
returns sum(list_of_numbers)/len(list_of_numbers)
'''
assert len(sample) != 0, "Unable to take the mean of an empty list"
for value in sample:
assert isinstance(value,int) or isinstance(value,float), "Value in list is not a number."
sample_mean = sum(sample) / len(sample)
return sample_mean
def main():
'''
Simple check of mean(num_list):
calls mean on:
numbers = [1, 2, 3, 4, 5],
returning the mean, 3.0
nonumbers = [], empty list
which causes an assertion error
word_and_numbers = [1, 2, 3, 4, "apple"], string in list
which would raise assertion error is executed
'''
numbers = [1, 2, 3, 4, 5]
print(mean(numbers))
nonumbers = []
print(mean(nonumbers))
word_and_numbers = [1, 2, 3, 4, "apple"]
print( mean(word_and_numbers) )
if __name__ == '__main__':
main()
```
Check that your documentation is working correctly by importing the script in ipython3 and running `help(my_script)`.
## Exercise: What is missing?
You are putting your code into scripts, preserving the code and enabling it’s re-use. You are developing tests allowing you to help demonstrate that the code is working as in tended. And you have documented your code so that you and other users can make use of the code in the future, But what have we forgotten?
[Solution]()
## Solution: What is missing?
Version Control!
In the directory containing your new script initialise a new git repository and add the script to it.
```bash
% git init
Initialised empty Git repository in /u/q/rjg20/intro-testing/my_testing/.git/
% git add mean.py
% git commit -m "Intiailised repository, a little late but with code, test and documentation"
[master (root-commit) c01f726] Intiailised repository, a little late but with code, test and documentation
1 files changed, 29 insertions(+)
create mode 100644 mean.py
```
Don’t forget to keep the repository upto date as we progress this lesson!
## Key Points:
- Writing functions in scripts makes reusable
- Documenting your functions is essential to make them re-usable
- All code in a script is executed by default.
- Specify a ‘main’ function which is only executed when run as a script.
- The ‘script’ can then be imported as any other Python library.
| github_jupyter |
# StructN2V - 2D Example for Convallaria data
```
# We import all our dependencies.
from n2v.models import N2VConfig, N2V
import numpy as np
from csbdeep.utils import plot_history
from n2v.utils.n2v_utils import manipulate_val_data
from n2v.internals.N2V_DataGenerator import N2V_DataGenerator
from matplotlib import pyplot as plt
import urllib
import os
import zipfile
from tifffile import imread
import ssl
ssl._create_default_https_context = ssl._create_unverified_context
```
# Download Example Data
*C. majalis* data acquired by Britta Schroth-Diez of the MPI-CBG Light Microscopy Facility.<br>
Thank you Britta!
```
# create a folder for our data.
if not os.path.isdir('./data'):
os.mkdir('./data')
# check if data has been downloaded already
zipPath="data/flower.tif"
if not os.path.exists(zipPath):
urllib.request.urlretrieve('https://download.fht.org/jug/n2v/flower.tif', zipPath)
data = imread("data/flower.tif")
```
# Training Data Preparation
For training we use the <code>N2V_DataGenerator</code> to extract training <code>X</code> and validation <code>X_val</code> patches.
```
datagen = N2V_DataGenerator()
imgs = datagen.load_imgs_from_directory(directory = "data/", dims="TYX")
print(imgs[0].shape)
# The function automatically added an extra "channels" dimensions to the images at the end
# Lets' look at the images.
# Select channel=0 in the last dimension, as `imshow()` doesn't really understand channels
plt.imshow(imgs[0][0,...,0], cmap='magma')
plt.show()
# split up image into little non-overlapping patches for training.
# y<832 (top of image) is training, y>=832 (bottom of image) is validation
imgs_train = [imgs[0][:,:832]]
X = datagen.generate_patches_from_list(imgs_train,shape=(96,96))
imgs_vali = [imgs[0][:,832:]]
X_val = datagen.generate_patches_from_list(imgs_vali,shape=(96,96))
# Patches are created so they do not overlap.
# (Note: this is not the case if you specify a number of patches. See the docstring for details!)
# Just in case you don't know how to access the docstring of a method:
datagen.generate_patches_from_list?
# Let's look at one of our training and validation patches.
plt.figure(figsize=(14,7))
plt.subplot(1,2,1)
plt.imshow(X[0,...,0], cmap='magma')
plt.title('Training Patch');
plt.subplot(1,2,2)
plt.imshow(X_val[0,...,0], cmap='magma')
plt.title('Validation Patch');
```
# Configure
Noise2Void comes with a special config-object, where we store network-architecture and training specific parameters. See the docstring of the <code>N2VConfig</code> constructor for a description of all parameters.
When creating the config-object, we provide the training data <code>X</code>. From <code>X</code> we extract <code>mean</code> and <code>std</code> that will be used to normalize all data before it is processed by the network. We also extract the dimensionality and number of channels from <code>X</code>.
Compared to supervised training (i.e. traditional CARE), we recommend to use N2V with an increased <code>train_batch_size</code> and <code>batch_norm</code>.
To keep the network from learning the identity we have to manipulate the input pixels during training. For this we have the parameter <code>n2v_manipulator</code> with default value <code>'uniform_withCP'</code>. Most pixel manipulators will compute the replacement value based on a neighborhood. With <code>n2v_neighborhood_radius</code> we can control its size.
Other pixel manipulators:
* normal_withoutCP: samples the neighborhood according to a normal gaussian distribution, but without the center pixel
* normal_additive: adds a random number to the original pixel value. The random number is sampled from a gaussian distribution with zero-mean and sigma = <code>n2v_neighborhood_radius</code>
* normal_fitted: uses a random value from a gaussian normal distribution with mean equal to the mean of the neighborhood and standard deviation equal to the standard deviation of the neighborhood.
* identity: performs no pixel manipulation
For faster training multiple pixels per input patch can be manipulated. In our experiments we manipulated about 0.198% of the input pixels per patch. For a patch size of 64 by 64 pixels this corresponds to about 8 pixels. This fraction can be tuned via <code>n2v_perc_pix</code>.
For Noise2Void training it is possible to pass arbitrarily large patches to the training method. From these patches random subpatches of size <code>n2v_patch_shape</code> are extracted during training. Default patch shape is set to (64, 64).
In the past we experienced bleedthrough artifacts between channels if training was terminated to early. To counter bleedthrough we added the `single_net_per_channel` option, which is turned on by default. In the back a single U-Net for each channel is created and trained independently, thereby removing the possiblity of bleedthrough. <br/>
__Note:__ Essentially the network gets multiplied by the number of channels, which increases the memory requirements. If your GPU gets too small, you can always split the channels manually and train a network for each channel one after another.
<font color='red'>Warning:</font> to make this example notebook execute faster, we have set <code>train_epochs</code> to only 10. <br>For better results we suggest 100 to 200 <code>train_epochs</code>.
```
# train_steps_per_epoch is set to (number of training patches)/(batch size), like this each training patch
# is shown once per epoch.
config = N2VConfig(X, unet_kern_size=3,
train_steps_per_epoch=int(X.shape[0]/128), train_epochs=10, train_loss='mse', batch_norm=True,
train_batch_size=128, n2v_perc_pix=0.198, n2v_patch_shape=(64, 64),
n2v_manipulator='uniform_withCP', n2v_neighborhood_radius=5, structN2Vmask = [[0,1,1,1,1,1,1,1,1,1,0]])
# Let's look at the parameters stored in the config-object.
vars(config)
# a name used to identify the model
model_name = 'n2v_2D'
# the base directory in which our model will live
basedir = 'models'
# We are now creating our network model.
model = N2V(config, model_name, basedir=basedir)
```
# Training
Training the model will likely take some time. We recommend to monitor the progress with TensorBoard, which allows you to inspect the losses during training. Furthermore, you can look at the predictions for some of the validation images, which can be helpful to recognize problems early on.
You can start TensorBoard in a terminal from the current working directory with tensorboard --logdir=. Then connect to http://localhost:6006/ with your browser.
```
# We are ready to start training now.
history = model.train(X, X_val)
```
### After training, lets plot training and validation loss.
```
print(sorted(list(history.history.keys())))
plt.figure(figsize=(16,5))
plot_history(history,['loss','val_loss']);
```
## Export Model in BioImage ModelZoo Format
See https://imagej.net/N2V#Prediction for details.
```
model.export_TF(name='Struct Noise2Void - Convallaria Example',
description='This is the Struct Noise2Void example trained on the Convallaria data in python.',
authors=["Coleman Broaddus"],
test_img=X_val[0], axes='YXC',
patch_shape=(96,96))
```
| github_jupyter |
# The Laplace Transform
*This Jupyter notebook is part of a [collection of notebooks](../index.ipynb) in the bachelors module Signals and Systems, Communications Engineering, Universität Rostock. Please direct questions and suggestions to [Sascha.Spors@uni-rostock.de](mailto:Sascha.Spors@uni-rostock.de).*
## Theorems
The theorems of the Laplace transformation relate basic time-domain operations to their equivalents in the Laplace domain. They are of use for the computation of Laplace transforms of signals composed from modified [standard signals](../continuous_signals/standard_signals.ipynb) and for the computation of the response of systems to an input signal. The theorems allow further to predict the consequences of modifying a signal or system by certain operations.
### Temporal Scaling Theorem
A signal $x(t)$ is given for which the Laplace transform $X(s) = \mathcal{L} \{ x(t) \}$ exists. The Laplace transform of the [temporally scaled signal](../continuous_signals/operations.ipynb#Temporal-Scaling) $x(a t)$ with $a \in \mathbb{R} \setminus \{0\}$ reads
\begin{equation}
\mathcal{L} \{ x(a t) \} = \frac{1}{|a|} \cdot X \left( \frac{s}{a} \right)
\end{equation}
The Laplace transformation of a temporally scaled signal is given by weighting the inversely scaled Laplace transform of the unscaled signal with $\frac{1}{|a|}$. The scaling of the Laplace transform can be interpreted as a scaling of the complex $s$-plane. The region of convergence (ROC) of the temporally scaled signal $x(a t)$ is consequently the inversely scaled ROC of the unscaled signal $x(t)$
\begin{equation}
\text{ROC} \{ x(a t) \} = \left\{ s: \frac{s}{a} \in \text{ROC} \{ x(t) \} \right\}
\end{equation}
Above relation is known as scaling theorem of the Laplace transform. The scaling theorem can be proven by introducing the scaled signal $x(a t)$ into the definition of the Laplace transformation
\begin{equation}
\mathcal{L} \{ x(a t) \} = \int_{-\infty}^{\infty} x(a t) \, e^{- s t} \; dt = \frac{1}{|a|} \int_{-\infty}^{\infty} x(t') \, e^{-\frac{s}{a} t'} \; dt' = \frac{1}{|a|} \cdot X \left( \frac{s}{a} \right)
\end{equation}
where the substitution $t' = a t$ was used. Note that a negative value of $a$ would result in a reversal of the integration limits. In this case a second reversal of the integration limits together with the sign of the integration element $d t'= a \, dt$ was consolidated into the absolute value of $a$.
### Convolution Theorem
The convolution theorem states that the Laplace transform of the convolution of two signals $x(t)$ and $y(t)$ is equal to the scalar multiplication of their Laplace transforms $X(s)$ and $Y(s)$
\begin{equation}
\mathcal{L} \{ x(t) * y(t) \} = X(s) \cdot Y(s)
\end{equation}
under the assumption that both Laplace transforms $X(s) = \mathcal{L} \{ x(t) \}$ and $Y(s) = \mathcal{L} \{ y(t) \}$ exist, respectively. The ROC of the convolution $x(t) * y(t)$ includes at least the intersection of the ROCs of $x(t)$ and $y(t)$
\begin{equation}
\text{ROC} \{ x(t) * y(t) \} \supseteq \text{ROC} \{ x(t) \} \cap \text{ROC} \{ y(t) \}
\end{equation}
The theorem can be proven by introducing the [definition of the convolution](../systems_time_domain/convolution.ipynb) into the [definition of the Laplace transform](definition.ipynb) and changing the order of integration
\begin{align}
\mathcal{L} \{ x(t) * y(t) \} &= \int_{-\infty}^{\infty} \left( \int_{-\infty}^{\infty} x(\tau) \cdot y(t-\tau) \; d \tau \right) e^{-s t} \; dt \\
&= \int_{-\infty}^{\infty} \left( \int_{-\infty}^{\infty} y(t-\tau) \, e^{-s t} \; dt \right) x(\tau) \; d\tau \\
&= Y(s) \cdot \int_{-\infty}^{\infty} x(\tau) \, e^{-s \tau} \; d \tau \\
&= Y(s) \cdot X(s)
\end{align}
The convolution theorem is very useful in the context of linear time-invariant (LTI) systems. The output signal $y(t)$ of an LTI system is given as the convolution of the input signal $x(t)$ with the impulse response $h(t)$. The signals can be represented either in the time or Laplace domain. This leads to the following equivalent representations of an LTI system in the time and Laplace domain, respectively

Calculation of the system response by transforming the problem into the Laplace domain can be beneficial since this replaces the evaluation of the convolution integral by a scalar multiplication. In many cases this procedure simplifies the calculation of the system response significantly. A prominent example is the [analysis of a passive electrical network](network_analysis.ipynb). The convolution theorem can also be useful to derive an unknown Laplace transform. The key is here to express the signal as convolution of two other signals for which the Laplace transforms are known. This is illustrated by the following example.
**Example**
The Laplace transform of the convolution of a causal cosine signal $\epsilon(t) \cdot \cos(\omega_0 t)$ with a causal sine signal $\epsilon(t) \cdot \sin(\omega_0 t)$ is derived by the convolution theorem
\begin{equation}
\mathcal{L} \{ \epsilon(t) \cdot ( \cos(\omega_0 t) * \sin(\omega_0 t) \}
= \frac{s}{s^2 + \omega_0^2} \cdot \frac{\omega_0}{s^2 + \omega_0^2}
= \frac{\omega_0 s}{(s^2 + \omega_0^2)^2}
\end{equation}
where the [Laplace transforms of the causal cosine and sine signals](properties.ipynb#Transformation-of-the-cosine-and-sine-signal) were used. The ROC of the causal cosine and sine signal is $\Re \{ s \} > 0$. The ROC for their convolution is also $\Re \{ s \} > 0$, since no poles and zeros cancel out. Above Laplace transform has one zero $s_{00} = 0$, and two poles of second degree $s_{\infty 0} = s_{\infty 1} = j \omega_0$ and $s_{\infty 2} = s_{\infty 3} = - j \omega_0$.
This example is evaluated numerically in the following. First the convolution of the causal cosine and sine signal is computed
```
%matplotlib inline
import sympy as sym
sym.init_printing()
t, tau = sym.symbols('t tau', real=True)
s = sym.symbols('s', complex=True)
w0 = sym.symbols('omega0', positive=True)
x = sym.integrate(sym.cos(w0*tau) * sym.sin(w0*(t-tau)), (tau, 0, t))
x = x.doit()
x
```
For the sake of illustration let's plot the signal for $\omega_0 = 1$
```
sym.plot(x.subs(w0, 1), (t, 0, 50), xlabel=r'$t$', ylabel=r'$x(t)$');
```
The Laplace transform is computed
```
X, a, cond = sym.laplace_transform(x, t, s)
X, a
```
which exists for $\Re \{ s \} > 0$. Its zeros are given as
```
sym.roots(sym.numer(X), s)
```
and its poles as
```
sym.roots(sym.denom(X), s)
```
### Temporal Shift Theorem
The [temporal shift of a signal](../continuous_signals/operations.ipynb#Temporal-Shift) $x(t - \tau)$ for $\tau \in \mathbb{R}$ can be expressed by the convolution of the signal $x(t)$ with a shifted Dirac impulse
\begin{equation}
x(t - \tau) = x(t) * \delta(t - \tau)
\end{equation}
This follows from the sifting property of the Dirac impulse. Applying a two-sided Laplace transform to the left- and right-hand side and exploiting the convolution theorem yields
\begin{equation}
\mathcal{L} \{ x(t - \tau) \} = X(s) \cdot e^{- s \tau}
\end{equation}
where $X(s) = \mathcal{L} \{ x(t) \}$ is assumed to exist. Note that $\mathcal{L} \{ \delta(t - \tau) \} = e^{- s \tau}$ can be derived from the definition of the two-sided Laplace transform together with the sifting property of the Dirac impulse. The Laplace transform of a shifted signal is given by multiplying the Laplace transform of the original signal with $e^{- s \tau}$. The ROC does not change
\begin{equation}
\text{ROC} \{ x(t-\tau) \} = \text{ROC} \{ x(t) \}
\end{equation}
This result is known as shift theorem of the Laplace transform. For a causal signal $x(t)$ and $\tau > 0$ the shift theorem of the one-sided Laplace transform is equal to the shift theorem of the two-sided transform.
#### Transformation of the rectangular signal
The Laplace transform of the [rectangular signal](../continuous_signals/standard_signals.ipynb#Rectangular-Signal) $x(t) = \text{rect}(t)$ is derived by expressing it by the Heaviside signal
\begin{equation}
\text{rect}(t) = \epsilon \left(t + \frac{1}{2} \right) - \epsilon \left(t - \frac{1}{2} \right)
\end{equation}
Applying the shift theorem to the [transform of the Heaviside signal](definition.ipynb#Transformation-of-the-Heaviside-Signal) and the linearity of the Laplace transform yields
\begin{equation}
\mathcal{L} \{ \text{rect}(t) \} = \frac{1}{s} e^{s \frac{1}{2}} - \frac{1}{s} e^{- s \frac{1}{2}} = \frac{\sinh \left( \frac{s}{2} \right) }{\frac{s}{2}}
\end{equation}
where $\sinh(\cdot)$ denotes the [hyperbolic sine function](https://en.wikipedia.org/wiki/Hyperbolic_function#Sinh). The ROC of the Heaviside signal is given as $\Re \{ s \} > 0$. Applying [l'Hopitals rule](https://en.wikipedia.org/wiki/L'H%C3%B4pital's_rule) the pole at $s=0$ can be disregarded leading to
\begin{equation}
\text{ROC} \{ \text{rect}(t) \} = \mathbb{C}
\end{equation}
For illustration, the magnitude of the Laplace transform $|X(s)|$ is plotted in the $s$-plane, as well as $X(\sigma)$ and $X(j \omega)$ for the real and imaginary part of the complex frequency $s = \sigma + j \omega$.
```
sigma, omega = sym.symbols('sigma omega')
X = sym.sinh(s/2)*2/s
sym.plotting.plot3d(abs(X.subs(s, sigma+sym.I*omega)), (sigma, -5, 5), (omega, -20, 20),
xlabel=r'$\Re\{s\}$', ylabel=r'$\Im\{s\}$', title=r'$|X(s)|$')
sym.plot(X.subs(s, sigma) , (sigma, -5, 5), xlabel=r'$\Re\{s\}$', ylabel=r'$X(s)$', ylim=(0, 3))
sym.plot(X.subs(s, sym.I*omega) , (omega, -20, 20), xlabel=r'$\Im\{s\}$', ylabel=r'$X(s)$');
```
**Exercise**
* Derive the Laplace transform $X(s) = \mathcal{L} \{ x(t) \}$ of the causal rectangular signal $x(t) = \text{rect} (a t - \frac{1}{2 a})$
* Derive the Laplace transform of the [triangular signal](../fourier_transform/theorems.ipynb#Transformation-of-the-triangular-signal) $x(t) = \Lambda(a t)$ with $a \in \mathbb{R} \setminus \{0\}$
### Differentiation Theorem
Derivatives of signals are the fundamental operations of differential equations. Ordinary differential equations (ODEs) with constant coefficients play an important role in the theory of time-invariant (LTI) systems. Consequently, the representation of the derivative of a signal in the Laplace domain is of special interest.
#### Two-sided transform
A differentiable signal $x(t)$ whose temporal derivative $\frac{d x(t)}{dt}$ exists is given. Using the [derivation property of the Dirac impulse](../continuous_signals/standard_signals.ipynb#Dirac-Impulse), the derivative of the signal can be expressed by the convolution
\begin{equation}
\frac{d x(t)}{dt} = \frac{d \delta(t)}{dt} * x(t)
\end{equation}
Applying a two-sided Laplace transformation to the left- and right-hand side together with the [convolution theorem](#Convolution-Theorem) yields the Laplace transform of the derivative of $x(t)$
\begin{equation}
\mathcal{L} \left\{ \frac{d x(t)}{dt} \right\} = s \cdot X(s)
\end{equation}
where $X(s) = \mathcal{L} \{ x(t) \}$. The two-sided Laplace transform $\mathcal{L} \{ \frac{d \delta(t)}{dt} \} = s$ can be derived by applying the definition of the Laplace transform together with the derivation property of the Dirac impulse. The ROC is given as a superset of the ROC for $x(t)$
\begin{equation}
\text{ROC} \left\{ \frac{d x(t)}{dt} \right\} \supseteq \text{ROC} \{ x(t) \}
\end{equation}
due to the zero at $s=0$ which may cancel out a pole.
Above result is known as differentiation theorem of the two-sided Laplace transform. It states that the differentiation of a signal in the time domain is equivalent to a multiplication of its spectrum by $s$.
#### One-sided transform
Many practical signals and systems are causal, hence $x(t) = 0$ for $t < 0$. A causal signal is potentially discontinuous for $t=0$. The direct application of above result for the two-sided Laplace transform is not possible since it assumes that the signal is differentiable for every time $t$. The potential discontinuity at $t=0$ has to be considered explicitly for the derivation of the differentiation theorem for the one-sided Laplace transform [[Girod et al.](index.ipynb#Literature)]
\begin{equation}
\mathcal{L} \left\{ \frac{d x(t)}{dt} \right\} = s \cdot X(s) - x(0+)
\end{equation}
where $x(0+) := \lim_{\epsilon \to 0} x(0+\epsilon)$ denotes the right sided limit value of $x(t)$ for $t=0$. The ROC is given as a superset of the ROC of $x(t)$
\begin{equation}
\text{ROC} \left\{ \frac{d x(t)}{dt} \right\} \supseteq \text{ROC} \{ x(t) \}
\end{equation}
due to the zero at $s=0$ which may cancel out a pole. The one-sided Laplace transform of a causal signal is equal to its two-sided transform. Above result holds therefore also for the two-sided transform of a causal signal.
The main application of the differentiation theorem is the transformation and solution of differential equations under consideration of initial values. Another application area is the derivation of transforms of signals which can be expressed as derivatives of other signals.
### Integration Theorem
An integrable signal $x(t)$ for which the integral $\int_{-\infty}^{t} x(\tau) \; d\tau$ exists is given. The integration can be represented as convolution with the rectangular signal $\epsilon(t)$
\begin{equation}
\int_{-\infty}^{t} x(\tau) \; d\tau = \int_{-\infty}^{\infty} x(\tau) \cdot \epsilon(t - \tau) \; d\tau = \epsilon(t) * x(t)
\end{equation}
as illustrated below

Two-sided Laplace transformation of the left- and right-hand side of above equation, application of the convolution theorem and using the Laplace transform of the Heaviside signal $\epsilon(t)$ yields
\begin{equation}
\mathcal{L} \left\{ \int_{-\infty}^{t} x(\tau) \; d\tau \right\}
= \frac{1}{s} \cdot X(s)
\end{equation}
The ROC is given as a superset of the intersection of the ROC of $x(t)$ and the right $s$-half-plane
\begin{equation}
\text{ROC} \left\{ \int_{-\infty}^{t} x(\tau) \; d\tau \right\} \supseteq \text{ROC} \{ x(t) \} \cap \{s : \Re \{ s \} > 0\}
\end{equation}
due to the pole at $s=0$. This integration theorem holds also for the one-sided Laplace transform.
#### Transformation of the ramp signal
The Laplace transform of the causal [ramp signal](https://en.wikipedia.org/wiki/Ramp_function) $t \cdot \epsilon(t)$ is derived by the integration theorem. The ramp signal can be expressed as integration over the Heaviside signal
\begin{equation}
t \cdot \epsilon(t) = \int_{-\infty}^{t} \tau \cdot \epsilon(\tau) \; d \tau
\end{equation}
Laplace transformation of the left- and right-hand side and application of the integration theorem together with the Laplace transform of the Heaviside signal yields
\begin{equation}
\mathcal{L} \{ t \cdot \epsilon(t) \} = \frac{1}{s^2}
\end{equation}
with
\begin{equation}
\text{ROC} \{ t \cdot \epsilon(t) \} = \{s : \Re \{ s \} > 0\}
\end{equation}
**Exercise**
* Derive the Laplace transform $X(s) = \mathcal{L} \{ x(t) \}$ of the signal $x(t) = t^n \cdot \epsilon(t)$ with $n \geq 0$ by repeated application of the integration theorem.
* Compare your result to the numerical result below. Note that $\Gamma(n+1) = n!$ for $n \in \mathbb{N}$.
```
n = sym.symbols('n', integer=True)
X, a, cond = sym.laplace_transform(t**n, t, s)
X, a, cond
```
### Modulation Theorem
The complex modulation of a signal $x(t)$ is defined as $e^{s_0 t} \cdot x(t)$ with $s_0 \in \mathbb{C}$. The Laplace transform of a modulated signal is derived by introducing it into the definition of the two-sided Laplace transform
\begin{align}
\mathcal{L} \left\{ e^{s_0 t} \cdot x(t) \right\} &=
\int_{-\infty}^{\infty} e^{s_0 t} x(t) \, e^{-s t} \; dt =
\int_{-\infty}^{\infty} x(t) \, e^{- (s - s_0) t} \; dt \\
&= X(s-s_0)
\end{align}
where $X(s) = \mathcal{L} \{ x(t) \}$. Modulation of the signal $x(t)$ leads to a translation of the $s$-plane into the direction given by the complex value $s_0$. Consequently, the ROC is also shifted
\begin{equation}
\text{ROC} \{ e^{s_0 t} \cdot x(t) \} = \{s: s - \Re \{ s_0 \} \in \text{ROC} \{ x(t) \} \}
\end{equation}
This relation is known as modulation theorem.
**Example**
The Laplace transform of the signal $t^n \cdot \epsilon(t)$
\begin{equation}
\mathcal{L} \{ t^n \cdot \epsilon(t) \} = \frac{n!}{s^{n+1}}
\end{equation}
for $\Re \{ s \} > 0$ was derived in the previous example. This result can be extended to the class of signals $t^n e^{-s_0 t} \epsilon(t)$ with $s_0 \in \mathbb{C}$ using the modulation theorem
\begin{equation}
\mathcal{L} \{ t^n e^{-s_0 t} \epsilon(t) \} = \frac{n!}{(s + s_0)^{n+1}} \qquad \text{for } \Re \{ s \} > \Re \{ - s_0 \}.
\end{equation}
**Copyright**
The notebooks are provided as [Open Educational Resource](https://de.wikipedia.org/wiki/Open_Educational_Resources). Feel free to use the notebooks for your own educational purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Lecture Notes on Signals and Systems* by Sascha Spors.
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D3_ModelFitting/W1D3_Tutorial4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy: Week 1, Day 3, Tutorial 4
# Model Fitting: Multiple linear regression and polynomial regression
**Content creators**: Pierre-Étienne Fiquet, Anqi Wu, Alex Hyafil with help from Byron Galbraith, Ella Batty
**Content reviewers**: Lina Teichmann, Saeed Salehi, Patrick Mineault, Michael Waskom
---
#Tutorial Objectives
This is Tutorial 4 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of regression models by generalizing to multiple linear regression and polynomial regression (Tutorial 4). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 5) and Cross Validation for model selection (Tutorial 6).
In this tutorial, we will generalize the regression model to incorporate multiple features.
- Learn how to structure inputs for regression using the 'Design Matrix'
- Generalize the MSE for multiple features using the ordinary least squares estimator
- Visualize data and model fit in multiple dimensions
- Fit polynomial regression models of different complexity
- Plot and evaluate the polynomial regression fits
```
#@title Video 1: Multiple Linear Regression and Polynomial Regression
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="d4nfTki6Ejc", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
---
# Setup
```
import numpy as np
import matplotlib.pyplot as plt
#@title Figure Settings
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
#@title Helper Functions
def plot_fitted_polynomials(x, y, theta_hat):
""" Plot polynomials of different orders
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
theta_hat (dict): polynomial regression weights for different orders
"""
x_grid = np.linspace(x.min() - .5, x.max() + .5)
plt.figure()
for order in range(0, max_order + 1):
X_design = make_design_matrix(x_grid, order)
plt.plot(x_grid, X_design @ theta_hat[order]);
plt.ylabel('y')
plt.xlabel('x')
plt.plot(x, y, 'C0.');
plt.legend([f'order {o}' for o in range(max_order + 1)], loc=1)
plt.title('polynomial fits')
plt.show()
```
---
# Section 1: Multiple Linear Regression
Now that we have considered the univariate case and how to produce confidence intervals for our estimator, we turn to the general linear regression case, where we can have more than one regressor, or feature, in our input.
Recall that our original univariate linear model was given as
\begin{align}
y = \theta x + \epsilon
\end{align}
where $\theta$ is the slope and $\epsilon$ some noise. We can easily extend this to the multivariate scenario by adding another parameter for each additional feature
\begin{align}
y = \theta_0 + \theta_1 x_1 + \theta_1 x_2 + ... +\theta_d x_d + \epsilon
\end{align}
where $\theta_0$ is the intercept and $d$ is the number of features (it is also the dimensionality of our input).
We can condense this succinctly using vector notation for a single data point
\begin{align}
y_i = \boldsymbol{\theta}^{\top}\mathbf{x}_i + \epsilon
\end{align}
and fully in matrix form
\begin{align}
\mathbf{y} = \mathbf{X}\boldsymbol{\theta} + \mathbf{\epsilon}
\end{align}
where $\mathbf{y}$ is a vector of measurements, $\mathbf{X}$ is a matrix containing the feature values (columns) for each input sample (rows), and $\boldsymbol{\theta}$ is our parameter vector.
This matrix $\mathbf{X}$ is often referred to as the "[design matrix](https://en.wikipedia.org/wiki/Design_matrix)".
For this tutorial we will focus on the two-dimensional case ($d=2$), which allows us to easily visualize our results. As an example, think of a situation where a scientist records the spiking response of a retinal ganglion cell to patterns of light signals that vary in contrast and in orientation. Then contrast and orientation values can be used as features / regressors to predict the cells response.
In this case our model can be writen as:
\begin{align}
y = \theta_0 + \theta_1 x_1 + \theta_2 x_2 + \epsilon
\end{align}
or in matrix form where
\begin{align}
\mathbf{X} =
\begin{bmatrix}
1 & x_{1,1} & x_{1,2} \\
1 & x_{2,1} & x_{2,2} \\
\vdots & \vdots & \vdots \\
1 & x_{n,1} & x_{n,2}
\end{bmatrix},
\boldsymbol{\theta} =
\begin{bmatrix}
\theta_0 \\
\theta_1 \\
\theta_2 \\
\end{bmatrix}
\end{align}
For our actual exploration dataset we shall set $\boldsymbol{\theta}=[0, -2, -3]$ and draw $N=40$ noisy samples from $x \in [-2,2)$. Note that setting the value of $\theta_0 = 0$ effectively ignores the offset term.
```
#@title
#@markdown Execute this cell to simulate some data
# Set random seed for reproducibility
np.random.seed(1234)
# Set parameters
theta = [0, -2, -3]
n_samples = 40
# Draw x and calculate y
n_regressors = len(theta)
x0 = np.ones((n_samples, 1))
x1 = np.random.uniform(-2, 2, (n_samples, 1))
x2 = np.random.uniform(-2, 2, (n_samples, 1))
X = np.hstack((x0, x1, x2))
noise = np.random.randn(n_samples)
y = X @ theta + noise
ax = plt.subplot(projection='3d')
ax.plot(X[:,1], X[:,2], y, '.')
ax.set(
xlabel='$x_1$',
ylabel='$x_2$',
zlabel='y'
)
plt.tight_layout()
```
Now that we have our dataset, we want to find an optimal vector of paramters $\boldsymbol{\hat\theta}$. Recall our analytic solution to minimizing MSE for a single regressor:
\begin{align}
\hat\theta = \frac{\sum_{i=1}^N x_i y_i}{\sum_{i=1}^N x_i^2}.
\end{align}
The same holds true for the multiple regressor case, only now expressed in matrix form
\begin{align}
\boldsymbol{\hat\theta} = (\mathbf{X}^\top\mathbf{X})^{-1}\mathbf{X}^\top\mathbf{y}.
\end{align}
This is called the [ordinary least squares](https://en.wikipedia.org/wiki/Ordinary_least_squares) (OLS) estimator.
### Exercise 1: Ordinary Least Squares Estimator
In this exercise you will implement the OLS approach to estimating $\boldsymbol{\hat\theta}$ from the design matrix $\mathbf{X}$ and measurement vector $\mathbf{y}$. You can use the `@` symbol for matrix multiplication, `.T` for transpose, and `np.linalg.inv` for matrix inversion.
```
def ordinary_least_squares(X, y):
"""Ordinary least squares estimator for linear regression.
Args:
x (ndarray): design matrix of shape (n_samples, n_regressors)
y (ndarray): vector of measurements of shape (n_samples)
Returns:
ndarray: estimated parameter values of shape (n_regressors)
"""
######################################################################
## TODO for students: solve for the optimal parameter vector using OLS
# Fill out function and remove
raise NotImplementedError("Student exercise: solve for theta_hat vector using OLS")
######################################################################
# Compute theta_hat using OLS
theta_hat = ...
return theta_hat
# Uncomment below to test your function
# theta_hat = ordinary_least_squares(X, y)
# print(theta_hat)
# to_remove solution
def ordinary_least_squares(X, y):
"""Ordinary least squares estimator for linear regression.
Args:
X (ndarray): design matrix of shape (n_samples, n_regressors)
y (ndarray): vector of measurements of shape (n_samples)
Returns:
ndarray: estimated parameter values of shape (n_regressors)
"""
# Compute theta_hat using OLS
theta_hat = np.linalg.inv(X.T @ X) @ X.T @ y
return theta_hat
theta_hat = ordinary_least_squares(X, y)
print(theta_hat)
```
After filling in this function, you should see that $\hat{\theta}$ = [ 0.13861386, -2.09395731, -3.16370742]
Now that we have our $\mathbf{\hat\theta}$, we can obtain $\mathbf{\hat y}$ and thus our mean squared error.
```
# Compute predicted data
theta_hat = ordinary_least_squares(X, y)
y_hat = X @ theta_hat
# Compute MSE
print(f"MSE = {np.mean((y - y_hat)**2):.2f}")
```
Finally, the following code will plot a geometric visualization of the data points (blue) and fitted plane. You can see that the residuals (green bars) are orthogonal to the plane.
Extra: how would you check that the residuals are orthogonal to the fitted plane? (this is sometimes called the orthogonality principle, see [least squares notes](https://www.cns.nyu.edu/~eero/NOTES/leastSquares.pdf))
```
#@title
#@markdown Execute this cell to visualize data and predicted plane
theta_hat = ordinary_least_squares(X, y)
xx, yy = np.mgrid[-2:2:50j, -2:2:50j]
y_hat_grid = np.array([xx.flatten(), yy.flatten()]).T @ theta_hat[1:]
y_hat_grid = y_hat_grid.reshape((50, 50))
ax = plt.subplot(projection='3d')
ax.plot(X[:, 1], X[:, 2], y, '.')
ax.plot_surface(xx, yy, y_hat_grid, linewidth=0, alpha=0.5, color='C1',
cmap=plt.get_cmap('coolwarm'))
for i in range(len(X)):
ax.plot((X[i, 1], X[i, 1]),
(X[i, 2], X[i, 2]),
(y[i], y_hat[i]),
'g-', alpha=.5)
ax.set(
xlabel='$x_1$',
ylabel='$x_2$',
zlabel='y'
)
plt.tight_layout()
```
---
# Section 2: Polynomial Regression
So far today, you learned how to predict outputs from inputs by fitting a linear regression model. We can now model all sort of relationships, including in neuroscience!
One potential problem with this approach is the simplicity of the model. Linear regression, as the name implies, can only capture a linear relationship between the inputs and outputs. Put another way, the predicted outputs are only a weighted sum of the inputs. What if there are more complicated computations happening? Luckily, many more complex models exist (and you will encounter many more over the next 3 weeks). One model that is still very simple to fit and understand, but captures more complex relationships, is **polynomial regression**, an extension of linear regression.
Since polynomial regression is an extension of linear regression, everything you learned so far will come in handy now! The goal is the same: we want to predict the dependent variable $y_{n}$ given the input values $x_{n}$. The key change is the type of relationship between inputs and outputs that the model can capture.
Linear regression models predict the outputs as a weighted sum of the inputs:
$$y_{n}= \theta_0 + \theta x_{n} + \epsilon_{n}$$
With polynomial regression, we model the outputs as a polynomial equation based on the inputs. For example, we can model the outputs as:
$$y_{n}= \theta_0 + \theta_1 x_{n} + \theta_2 x_{n}^2 + \theta_3 x_{n}^3 + \epsilon_{n}$$
We can change how complex a polynomial is fit by changing the order of the polynomial. The order of a polynomial refers to the highest power in the polynomial. The equation above is a third order polynomial because the highest value x is raised to is 3. We could add another term ($+ \theta_4 x_{n}^4$) to model an order 4 polynomial and so on.
First, we will simulate some data to practice fitting polynomial regression models. We will generate random inputs $x$ and then compute y according to $y = x^2 - x - 2 $, with some extra noise both in the input and the output to make the model fitting exercise closer to a real life situation.
```
#@title
#@markdown Execute this cell to simulate some data
# setting a fixed seed to our random number generator ensures we will always
# get the same psuedorandom number sequence
np.random.seed(121)
n_samples = 30
x = np.random.uniform(-2, 2.5, n_samples) # inputs uniformly sampled from [-2, 2.5)
y = x**2 - x - 2 # computing the outputs
output_noise = 1/8 * np.random.randn(n_samples)
y += output_noise # adding some output noise
input_noise = 1/2 * np.random.randn(n_samples)
x += input_noise # adding some input noise
fig, ax = plt.subplots()
ax.scatter(x, y) # produces a scatter plot
ax.set(xlabel='x', ylabel='y');
```
## Section 2.1: Design matrix for polynomial regression
Now we have the basic idea of polynomial regression and some noisy data, let's begin! The key difference between fitting a linear regression model and a polynomial regression model lies in how we structure the input variables.
For linear regression, we used $X = x$ as the input data. To add a constant bias (a y-intercept in a 2-D plot), we use $X = \big[ \boldsymbol 1, x \big]$, where $\boldsymbol 1$ is a column of ones. When fitting, we learn a weight for each column of this matrix. So we learn a weight that multiples with column 1 - in this case that column is all ones so we gain the bias parameter ($+ \theta_0$). We also learn a weight for every column, or every feature of x, as we learned in Section 1.
This matrix $X$ that we use for our inputs is known as a **design matrix**. We want to create our design matrix so we learn weights for $x^2, x^3,$ etc. Thus, we want to build our design matrix $X$ for polynomial regression of order $k$ as:
$$X = \big[ \boldsymbol 1 , x^1, x^2 , \ldots , x^k \big],$$
where $\boldsymbol{1}$ is the vector the same length as $x$ consisting of of all ones, and $x^p$ is the vector or matrix $x$ with all elements raised to the power $p$. Note that $\boldsymbol{1} = x^0$ and $x^1 = x$
### Exercise 2: Structure design matrix
Create a function (`make_design_matrix`) that structures the design matrix given the input data and the order of the polynomial you wish to fit. We will print part of this design matrix for our data and order 5.
```
def make_design_matrix(x, order):
"""Create the design matrix of inputs for use in polynomial regression
Args:
x (ndarray): input vector of shape (n_samples)
order (scalar): polynomial regression order
Returns:
ndarray: design matrix for polynomial regression of shape (samples, order+1)
"""
########################################################################
## TODO for students: create the design matrix ##
# Fill out function and remove
raise NotImplementedError("Student exercise: create the design matrix")
########################################################################
# Broadcast to shape (n x 1) so dimensions work
if x.ndim == 1:
x = x[:, None]
#if x has more than one feature, we don't want multiple columns of ones so we assign
# x^0 here
design_matrix = np.ones((x.shape[0], 1))
# Loop through rest of degrees and stack columns (hint: np.hstack)
for degree in range(1, order + 1):
design_matrix = ...
return design_matrix
# Uncomment to test your function
order = 5
# X_design = make_design_matrix(x, order)
# print(X_design[0:2, 0:2])
# to_remove solution
def make_design_matrix(x, order):
"""Create the design matrix of inputs for use in polynomial regression
Args:
x (ndarray): input vector of shape (samples,)
order (scalar): polynomial regression order
Returns:
ndarray: design matrix for polynomial regression of shape (samples, order+1)
"""
# Broadcast to shape (n x 1) so dimensions work
if x.ndim == 1:
x = x[:, None]
#if x has more than one feature, we don't want multiple columns of ones so we assign
# x^0 here
design_matrix = np.ones((x.shape[0], 1))
# Loop through rest of degrees and stack columns
for degree in range(1, order + 1):
design_matrix = np.hstack((design_matrix, x**degree))
return design_matrix
order = 5
X_design = make_design_matrix(x, order)
print(X_design[0:2, 0:2])
```
You should see that the printed section of this design matrix is `[[ 1. -1.51194917]
[ 1. -0.35259945]]`
## Section 2.2: Fitting polynomial regression models
Now that we have the inputs structured correctly in our design matrix, fitting a polynomial regression is the same as fitting a linear regression model! All of the polynomial structure we need to learn is contained in how the inputs are structured in the design matrix. We can use the same least squares solution we computed in previous exercises.
### Exercise 3: Fitting polynomial regression models with different orders
Here, we will fit polynomial regression models to find the regression coefficients ($\theta_0, \theta_1, \theta_2,$ ...) by solving the least squares problem. Create a function `solve_poly_reg` that loops over different order polynomials (up to `max_order`), fits that model, and saves out the weights for each. You may invoke the `ordinary_least_squares` function.
We will then qualitatively inspect the quality of our fits for each order by plotting the fitted polynomials on top of the data. In order to see smooth curves, we evaluate the fitted polynomials on a grid of $x$ values (ranging between the largest and smallest of the inputs present in the dataset).
```
def solve_poly_reg(x, y, max_order):
"""Fit a polynomial regression model for each order 0 through max_order.
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
max_order (scalar): max order for polynomial fits
Returns:
dict: fitted weights for each polynomial model (dict key is order)
"""
# Create a dictionary with polynomial order as keys, and np array of theta_hat
# (weights) as the values
theta_hats = {}
# Loop over polynomial orders from 0 through max_order
for order in range(max_order + 1):
##################################################################################
## TODO for students: Create design matrix and fit polynomial model for this order
# Fill out function and remove
raise NotImplementedError("Student exercise: fit a polynomial model")
##################################################################################
# Create design matrix
X_design = ...
# Fit polynomial model
this_theta = ...
theta_hats[order] = this_theta
return theta_hats
# Uncomment to test your function
max_order = 5
#theta_hats = solve_poly_reg(x, y, max_order)
#plot_fitted_polynomials(x, y, theta_hats)
# to_remove solution
def solve_poly_reg(x, y, max_order):
"""Fit a polynomial regression model for each order 0 through max_order.
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
max_order (scalar): max order for polynomial fits
Returns:
dict: fitted weights for each polynomial model (dict key is order)
"""
# Create a dictionary with polynomial order as keys, and np array of theta
# (weights) as the values
theta_hats = {}
# Loop over polynomial orders from 0 through max_order
for order in range(max_order + 1):
X_design = make_design_matrix(x, order)
this_theta = ordinary_least_squares(X_design, y)
theta_hats[order] = this_theta
return theta_hats
max_order = 5
theta_hats = solve_poly_reg(x, y, max_order)
with plt.xkcd():
plot_fitted_polynomials(x, y, theta_hats)
```
## Section 2.4: Evaluating fit quality
As with linear regression, we can compute mean squared error (MSE) to get a sense of how well the model fits the data.
We compute MSE as:
$$ MSE = \frac 1 N ||y - \hat y||^2 = \sum_{i=1}^N (y_i - \hat y_i)^2 $$
where the predicted values for each model are given by $ \hat y = X \hat \theta$.
*Which model (i.e. which polynomial order) do you think will have the best MSE?*
### Exercise 4: Compute MSE and compare models
We will compare the MSE for different polynomial orders with a bar plot.
```
mse = np.zeros((max_order + 1))
for order in range(0, max_order + 1):
X_design = make_design_matrix(x, order)
############################################################################
## TODO for students: Compute MSE
#############################################################################
## Uncomment below and fill in with your code
# Get prediction for the polynomial regression model of this order
#y_hat = ...
# Compute the residuals
#residuals = ...
# Compute the MSE
#mse[order] = ...
pass
fig, ax = plt.subplots()
# Uncomment once above exercise is complete
# ax.bar(range(max_order + 1), mse);
ax.set(title='Comparing Polynomial Fits', xlabel='Polynomial order', ylabel='MSE')
# to_remove solution
mse = np.zeros((max_order + 1))
for order in range(0, max_order + 1):
X_design = make_design_matrix(x, order)
# Get prediction for the polynomial regression model of this order
y_hat = X_design @ theta_hats[order]
# Compute the residuals
residuals = y - y_hat
# Compute the MSE
mse[order] = np.mean(residuals ** 2)
with plt.xkcd():
fig, ax = plt.subplots()
ax.bar(range(max_order + 1), mse);
ax.set(title='Comparing Polynomial Fits', xlabel='Polynomial order', ylabel='MSE')
```
---
# Summary
* Linear regression generalizes naturally to multiple dimensions
* Linear algebra affords us the mathematical tools to reason and solve such problems beyond the two dimensional case
* To change from a linear regression model to a polynomial regression model, we only have to change how the input data is structured
* We can choose the complexity of the model by changing the order of the polynomial model fit
* Higher order polynomial models tend to have lower MSE on the data they're fit with
**Note**: In practice, multidimensional least squares problems can be solved very efficiently (thanks to numerical routines such as LAPACK).
**Suggested readings**
[Introduction to Applied Linear Algebra – Vectors, Matrices, and Least Squares](http://vmls-book.stanford.edu/)
Stephen Boyd and Lieven Vandenberghe
| github_jupyter |
# Working on weather data for a project
[Citrics](https://b.citrics.dev/) is a project that helps people decide before moving to a new city by providing them valuable informations on different cities. One of the core features of the project is being able to get information about top job industries of different cities and compare them. This notebook shows how the data was cleaned, wrangled and new features were created so that they can be used for getting job industry insights.
The data were collected from [Bureau of Labor Statistics (BLS)](https://www.bls.gov/oes/)
```
import pandas as pd
import numpy as np
# The first dataset is from BLS
# The second dataset contains unique identifiers
# for the 100 cities we worked on
df = pd.read_csv("/content/us_job_industry_data_2019.csv")
df_cities = pd.read_csv("/content/100city_state_data.csv")
df.head()
# Cleaning up some area titles
df = df.replace(['Phoenix-Mesa-Scottsdale, AZ', 'Los Angeles-Long Beach-Anaheim, CA', 'Killeen-Temple, TX', 'Virginia Beach-Norfolk-Newport News, VA-NC'], ['Phoenix-Mesa-Scottsdale-Chandler-Gilbert, AZ', 'Los Angeles-Long Beach-Anaheim-Glendale-Santa Ana, CA', 'Killeen-Temple-Irving-Plano, TX', 'Virginia Beach-Norfolk-Newport News-Chesapeake, VA-NC'])
df.shape, df_cities.shape
# Restricting area type to city
df = df[ df["area_type"] == 4]
df.shape, df_cities.shape
# A function to create city names from area title
def create_cities(areaname):
#print(areaname)
areaname = areaname.replace("--","-")
cities, garbage1 = areaname.split(",")
areaname = cities.split("-")
#print(f"{cities} ---- {list_cities}")
return areaname
# Applying the function
df['city'] = df.area_title.apply(create_cities)
df.head()
df_explode = df.explode('city')
df_explode.shape
df_explode.columns
df_explode[['area_title', 'city']].head()
missing = ['Chandler', 'Gilbert', 'Glendale', 'Santa Ana', 'Honolulu', 'Boise', 'Louisville', 'Winston Salem', 'Irving', 'Plano', 'Chesapeake']
sorted(missing)
len(df_explode.city.unique())
len(df_explode.area_title.unique())
# Cleaning up some more city names
df_explode = df_explode.replace(['Boise City', 'Louisville/Jefferson County', 'Urban Honolulu', 'Winston'], ['Boise', 'Louisville', 'Honolulu', 'Winston Salem'])
# merging with the dataframe with unique identifier
merged = df_cities.merge(df_explode, on="city", how="left")
merged.head()
merged.shape
merged.isnull().sum()
len(merged.city.unique())
# Downloading the data
merged.to_csv('test1.csv')
```
To see how the endpoints were created using this data click [here](https://colab.research.google.com/drive/1IJD9aRTVFZiIvYKZztM2xzUEbmkM3Y6E?usp=sharing)
| github_jupyter |
```
# NOTE: PLEASE MAKE SURE YOU ARE RUNNING THIS IN A PYTHON3 ENVIRONMENT
import tensorflow as tf
print(tf.__version__)
# This is needed for the iterator over the data
# But not necessary if you have TF 2.0 installed
#!pip install tensorflow==2.0.0-beta0
tf.enable_eager_execution()
# !pip install -q tensorflow-datasets
import tensorflow_datasets as tfds
imdb, info = tfds.load("imdb_reviews", with_info=True, as_supervised=True)
import numpy as np
train_data, test_data = imdb['train'], imdb['test']
training_sentences = []
training_labels = []
testing_sentences = []
testing_labels = []
# str(s.tonumpy()) is needed in Python3 instead of just s.numpy()
for s,l in train_data:
training_sentences.append(str(s.numpy()))
training_labels.append(l.numpy())
for s,l in test_data:
testing_sentences.append(str(s.numpy()))
testing_labels.append(l.numpy())
training_labels_final = np.array(training_labels)
testing_labels_final = np.array(testing_labels)
vocab_size = 10000
embedding_dim = 16
max_length = 120
trunc_type='post'
oov_tok = "<OOV>"
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tokenizer = Tokenizer(num_words = vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
sequences = tokenizer.texts_to_sequences(training_sentences)
padded = pad_sequences(sequences,maxlen=max_length, truncating=trunc_type)
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences,maxlen=max_length)
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
print(decode_review(padded[1]))
print(training_sentences[1])
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(6, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
num_epochs = 10
model.fit(padded, training_labels_final, epochs=num_epochs, validation_data=(testing_padded, testing_labels_final))
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
import io
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for word_num in range(1, vocab_size):
word = reverse_word_index[word_num]
embeddings = weights[word_num]
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in embeddings]) + "\n")
out_v.close()
out_m.close()
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
sentence = "I really think this is amazing. honest."
sequence = tokenizer.texts_to_sequences(sentence)
print(sequence)
```
| github_jupyter |
# EECS 445: Machine Learning
## Hands On 11: Info Theory and Decision Trees
Prove that the KL divergence $KL(p,q)$ for any distributions $p,q$ is non negative.
*Hint*: Show that $$\min_{q} KL(p,q) \geq 0$$
Prove that two random variables $X$ and $Y$ are independent if and only if $I(X,Y) = 0$ where $I$ is the mutual information.
*Hint*: In the last problem, we almost proved
$$KL(p,q) = 0 \iff p = q$$
but we swear it's true
Prove that
* $I(X, Y) = H(X) + H(Y) - H(X,Y)$
* $I(X, Y) = H(X) - H(X | Y)$
Prove that
$$I(X,Y) = H(X)$$ if $Y$ is a determinisitc, one-to-one function of $X$
## Decision trees vs linear models
Let's explore cases where decision trees perform better and worse than linear models.
Note: we're using the function 'plot_decision_regions' from the [mlextend library](https://github.com/rasbt/mlxtend); you can install with pip, conda or just copy and paste the function into a cell from [here](https://raw.githubusercontent.com/rasbt/mlxtend/master/mlxtend/plotting/decision_regions.py).
```
$ conda install -c rasbt mlxtend
```
```
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression, Perceptron
import numpy as np
import matplotlib.pyplot as plt
from mlxtend.evaluate import plot_decision_regions
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
```
## Trees can fit XOR
The classic minimal function that a linear model can't fit is XOR. Let's visualize how linear and tree models manage to fit AND and XOR.
```
X = np.array([
[-1, -1],
[-1, 1],
[1, -1],
[1, 1]
])
y_and = np.array([0, 1, 1, 1])
y_xor = np.array([0, 1, 1, 0])
lr = LogisticRegression(C=100000)
for label, y in [('and', y_and), ('xor', y_xor)]:
lr.fit(X, y)
plot_decision_regions(X, y, lr)
plt.xlabel('x')
plt.ylabel('y')
plt.title('linear model fitting "{}"'.format(label))
plt.show()
tree = DecisionTreeClassifier(criterion='entropy', max_depth=2, random_state=0)
tree.fit(X, y)
for label, y in [('and', y_and), ('xor', y_xor)]:
tree.fit(X, y)
plot_decision_regions(X, y, tree)
plt.xlabel('x')
plt.ylabel('y')
plt.title('decision tree model fitting "{}"'.format(label))
plt.show()
```
## When a linear models beat a decision tree
For this exercise, construct a dataset that a linear model can fit, but that a decision tree of depth 2 cannot.
```
from sklearn.metrics import accuracy_score
X_linear_wins = np.array([
# place 2d samples here, each value between -1 and 1
])
y_linear_wins = np.array([
# place class label 0, 1 for each 2d point here
])
# uncommment code below to test out whether your dataset is more accurately predicted by a linear model
# than a tree of depth 3.
# for label, model in [('linear', lr), ('tree', tree)]:
# model.fit(X_linear_wins, y_linear_wins)
# plot_decision_regions(X_linear_wins, y_linear_wins, model)
# plt.xlabel('x')
# plt.ylabel('y')
# title = "{}: accuracy {:.2f}".format(label, accuracy_score(y_linear_wins, model.predict(X_linear_wins)))
# plt.title(title)
# plt.show()
```
## Depth matters
For this exercise, construct a dataset that cannot be 100% accurately classified with a tree of depth 2 but *can* be by a tree of depth 3.
```
X_needs_depth = np.array([
# place 2d samples here, each value between -1 and 1
])
y_needs_depth = np.array([
# place class label 0, 1 for each 2d point here
])
# uncomment the code below to compare
# tree_d2 = DecisionTreeClassifier(criterion='entropy', max_depth=2, random_state=0)
# tree_d4 = DecisionTreeClassifier(criterion='entropy', max_depth=4, random_state=0)
# tree_d2.fit(X_needs_depth, y_needs_depth)
# plot_decision_regions(X_needs_depth, y_needs_depth, tree_d2)
# plt.title("depth 2 fit: {:.2f}".format(accuracy_score(y_needs_depth, tree_d2.predict(X_needs_depth))))
# plt.show()
# tree_d4.fit(X_needs_depth, y_needs_depth)
# plot_decision_regions(X_needs_depth, y_needs_depth, tree_d4)
# plt.title("depth 4 fit: {:.2f}".format(accuracy_score(y_needs_depth, tree_d4.predict(X_needs_depth))))
# plt.show()
```
| github_jupyter |
```
# http://pytorch.org/
from os.path import exists
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
platform = '{}{}-{}'.format(get_abbr_impl(), get_impl_ver(), get_abi_tag())
cuda_output = !ldconfig -p|grep cudart.so|sed -e 's/.*\.\([0-9]*\)\.\([0-9]*\)$/cu\1\2/'
accelerator = cuda_output[0] if exists('/dev/nvidia0') else 'cpu'
'''
code by Tae Hwan Jung(Jeff Jung) @graykode
Reference : https://github.com/jadore801120/attention-is-all-you-need-pytorch
https://github.com/JayParks/transformer
'''
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
import torch.nn.functional as F
import matplotlib.pyplot as plt
dtype = torch.FloatTensor
# S: Symbol that shows starting of decoding input
# E: Symbol that shows starting of decoding output
# P: Symbol that will fill in blank sequence if current batch data size is short than time steps
sentences = ['ich mochte ein bier P', 'S i want a beer', 'i want a beer E']
# Transformer Parameters
src_vocab = {'PAD' : 0}
for i, w in enumerate(sentences[0].split()):
src_vocab[w] = i+1
src_vocab_size = len(src_vocab)
tgt_vocab = {'PAD' : 0}
number_dict = {0 : 'PAD'}
for i, w in enumerate(set((sentences[1]+' '+sentences[2]).split())):
tgt_vocab[w] = i+1
number_dict[i+1] = w
tgt_vocab_size = len(tgt_vocab)
src_len = 5
tgt_len = 5
d_model = 512 # Embedding Size
d_ff = 2048 # FeedForward dimension
d_k = d_v = 64 # dimension of K(=Q), V
n_layers = 6 # number of Encoder of Decoder Layer
n_heads = 8 # number of heads in Multi-Head Attention
def make_batch(sentences):
input_batch = [[src_vocab[n] for n in sentences[0].split()]]
output_batch = [[tgt_vocab[n] for n in sentences[1].split()]]
target_batch = [[tgt_vocab[n] for n in sentences[2].split()]]
return Variable(torch.LongTensor(input_batch)), Variable(torch.LongTensor(output_batch)), Variable(torch.LongTensor(target_batch))
def get_sinusoid_encoding_table(n_position, d_model):
def cal_angle(position, hid_idx):
return position / np.power(10000, 2 * (hid_idx // 2) / d_model)
def get_posi_angle_vec(position):
return [cal_angle(position, hid_j) for hid_j in range(d_model)]
sinusoid_table = np.array([get_posi_angle_vec(pos_i) for pos_i in range(n_position)])
sinusoid_table[:, 0::2] = np.sin(sinusoid_table[:, 0::2]) # dim 2i
sinusoid_table[:, 1::2] = np.cos(sinusoid_table[:, 1::2]) # dim 2i+1
return torch.FloatTensor(sinusoid_table)
def get_attn_pad_mask(seq_q, seq_k):
batch_size, len_q = seq_q.size()
batch_size, len_k = seq_k.size()
# eq(zero) is PAD token
pad_attn_mask = seq_k.data.eq(0).unsqueeze(1) # batch_size x 1 x len_k(=len_q), one is masking
return pad_attn_mask.expand(batch_size, len_q, len_k) # batch_size x len_q x len_k
def get_attn_subsequent_mask(seq):
attn_shape = [seq.size(0), seq.size(1), seq.size(1)]
subsequent_mask = np.triu(np.ones(attn_shape), k=1)
subsequent_mask = torch.from_numpy(subsequent_mask).byte()
return subsequent_mask
class ScaledDotProductAttention(nn.Module):
def __init__(self):
super(ScaledDotProductAttention, self).__init__()
def forward(self, Q, K, V, attn_mask):
scores = torch.matmul(Q, K.transpose(-1, -2)) / np.sqrt(d_k) # scores : [batch_size x n_heads x len_q(=len_k) x len_k(=len_q)]
scores.masked_fill_(attn_mask, -1e9) # Fills elements of self tensor with value where mask is one.
attn = nn.Softmax(dim=-1)(scores)
context = torch.matmul(attn, V)
return context, attn
class MultiHeadAttention(nn.Module):
def __init__(self):
super(MultiHeadAttention, self).__init__()
self.W_Q = nn.Linear(d_model, d_k * n_heads)
self.W_K = nn.Linear(d_model, d_k * n_heads)
self.W_V = nn.Linear(d_model, d_v * n_heads)
def forward(self, Q, K, V, attn_mask):
# q: [batch_size x len_q x d_model], k: [batch_size x len_k x d_model], v: [batch_size x len_k x d_model]
residual, batch_size = Q, Q.size(0)
# (B, S, D) -proj-> (B, S, D) -split-> (B, S, H, W) -trans-> (B, H, S, W)
q_s = self.W_Q(Q).view(batch_size, -1, n_heads, d_k).transpose(1,2) # q_s: [batch_size x n_heads x len_q x d_k]
k_s = self.W_K(K).view(batch_size, -1, n_heads, d_k).transpose(1,2) # k_s: [batch_size x n_heads x len_k x d_k]
v_s = self.W_V(V).view(batch_size, -1, n_heads, d_v).transpose(1,2) # v_s: [batch_size x n_heads x len_k x d_v]
attn_mask = attn_mask.unsqueeze(1).repeat(1, n_heads, 1, 1) # attn_mask : [batch_size x n_heads x len_q x len_k]
# context: [batch_size x n_heads x len_q x d_v], attn: [batch_size x n_heads x len_q(=len_k) x len_k(=len_q)]
context, attn = ScaledDotProductAttention()(q_s, k_s, v_s, attn_mask)
context = context.transpose(1, 2).contiguous().view(batch_size, -1, n_heads * d_v) # context: [batch_size x len_q x n_heads * d_v]
output = nn.Linear(n_heads * d_v, d_model)(context)
return nn.LayerNorm(d_model)(output + residual), attn # output: [batch_size x len_q x d_model]
class PoswiseFeedForwardNet(nn.Module):
def __init__(self):
super(PoswiseFeedForwardNet, self).__init__()
self.conv1 = nn.Conv1d(in_channels=d_model, out_channels=d_ff, kernel_size=1)
self.conv2 = nn.Conv1d(in_channels=d_ff, out_channels=d_model, kernel_size=1)
def forward(self, inputs):
residual = inputs # inputs : [batch_size, len_q, d_model]
output = nn.ReLU()(self.conv1(inputs.transpose(1, 2)))
output = self.conv2(output).transpose(1, 2)
return nn.LayerNorm(d_model)(output + residual)
class EncoderLayer(nn.Module):
def __init__(self):
super(EncoderLayer, self).__init__()
self.enc_self_attn = MultiHeadAttention()
self.pos_ffn = PoswiseFeedForwardNet()
def forward(self, enc_inputs, enc_self_attn_mask):
enc_outputs, attn = self.enc_self_attn(enc_inputs, enc_inputs, enc_inputs, enc_self_attn_mask) # enc_inputs to same Q,K,V
enc_outputs = self.pos_ffn(enc_outputs) # enc_outputs: [batch_size x len_q x d_model]
return enc_outputs, attn
class DecoderLayer(nn.Module):
def __init__(self):
super(DecoderLayer, self).__init__()
self.dec_self_attn = MultiHeadAttention()
self.dec_enc_attn = MultiHeadAttention()
self.pos_ffn = PoswiseFeedForwardNet()
def forward(self, dec_inputs, enc_outputs, dec_self_attn_mask, dec_enc_attn_mask):
dec_outputs, dec_self_attn = self.dec_self_attn(dec_inputs, dec_inputs, dec_inputs, dec_self_attn_mask)
dec_outputs, dec_enc_attn = self.dec_enc_attn(dec_outputs, enc_outputs, enc_outputs, dec_enc_attn_mask)
dec_outputs = self.pos_ffn(dec_outputs)
return dec_outputs, dec_self_attn, dec_enc_attn
class Encoder(nn.Module):
def __init__(self):
super(Encoder, self).__init__()
self.src_emb = nn.Embedding(src_vocab_size, d_model)
self.pos_emb = nn.Embedding.from_pretrained(get_sinusoid_encoding_table(src_len+1 , d_model),freeze=True)
self.layers = nn.ModuleList([EncoderLayer() for _ in range(n_layers)])
def forward(self, enc_inputs): # enc_inputs : [batch_size x source_len]
enc_outputs = self.src_emb(enc_inputs) + self.pos_emb(torch.LongTensor([[1,2,3,4,5]]))
enc_self_attn_mask = get_attn_pad_mask(enc_inputs, enc_inputs)
enc_self_attns = []
for layer in self.layers:
enc_outputs, enc_self_attn = layer(enc_outputs, enc_self_attn_mask)
enc_self_attns.append(enc_self_attn)
return enc_outputs, enc_self_attns
class Decoder(nn.Module):
def __init__(self):
super(Decoder, self).__init__()
self.tgt_emb = nn.Embedding(tgt_vocab_size, d_model)
self.pos_emb = nn.Embedding.from_pretrained(get_sinusoid_encoding_table(tgt_len+1 , d_model),freeze=True)
self.layers = nn.ModuleList([DecoderLayer() for _ in range(n_layers)])
def forward(self, dec_inputs, enc_inputs, enc_outputs): # dec_inputs : [batch_size x target_len]
dec_outputs = self.tgt_emb(dec_inputs) + self.pos_emb(torch.LongTensor([[1,2,3,4,5]]))
dec_self_attn_pad_mask = get_attn_pad_mask(dec_inputs, dec_inputs)
dec_self_attn_subsequent_mask = get_attn_subsequent_mask(dec_inputs)
dec_self_attn_mask = torch.gt((dec_self_attn_pad_mask + dec_self_attn_subsequent_mask), 0)
dec_enc_attn_mask = get_attn_pad_mask(dec_inputs, enc_inputs)
dec_self_attns, dec_enc_attns = [], []
for layer in self.layers:
dec_outputs, dec_self_attn, dec_enc_attn = layer(dec_outputs, enc_outputs, dec_self_attn_mask, dec_enc_attn_mask)
dec_self_attns.append(dec_self_attn)
dec_enc_attns.append(dec_enc_attn)
return dec_outputs, dec_self_attns, dec_enc_attns
class Transformer(nn.Module):
def __init__(self):
super(Transformer, self).__init__()
self.encoder = Encoder()
self.decoder = Decoder()
self.projection = nn.Linear(d_model, tgt_vocab_size, bias=False)
def forward(self, enc_inputs, dec_inputs):
enc_outputs, enc_self_attns = self.encoder(enc_inputs)
dec_outputs, dec_self_attns, dec_enc_attns = self.decoder(dec_inputs, enc_inputs, enc_outputs)
dec_logits = self.projection(dec_outputs) # dec_logits : [batch_size x src_vocab_size x tgt_vocab_size]
return dec_logits.view(-1, dec_logits.size(-1)), enc_self_attns, dec_self_attns, dec_enc_attns
model = Transformer()
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
for epoch in range(100):
optimizer.zero_grad()
enc_inputs, dec_inputs, target_batch = make_batch(sentences)
outputs, enc_self_attns, dec_self_attns, dec_enc_attns = model(enc_inputs, dec_inputs)
loss = criterion(outputs, target_batch.contiguous().view(-1))
if (epoch + 1) % 20 == 0:
print('Epoch:', '%04d' % (epoch + 1), 'cost =', '{:.6f}'.format(loss))
loss.backward()
optimizer.step()
def showgraph(attn):
attn = attn[-1].squeeze(0)[0]
attn = attn.squeeze(0).data.numpy()
fig = plt.figure(figsize=(n_heads, n_heads)) # [n_heads, n_heads]
ax = fig.add_subplot(1, 1, 1)
ax.matshow(attn, cmap='viridis')
ax.set_xticklabels(['']+sentences[0].split(), fontdict={'fontsize': 14}, rotation=90)
ax.set_yticklabels(['']+sentences[2].split(), fontdict={'fontsize': 14})
plt.show()
# Test
predict, _, _, _ = model(enc_inputs, dec_inputs)
predict = predict.data.max(1, keepdim=True)[1]
print(sentences[0], '->', [number_dict[n.item()] for n in predict.squeeze()])
print('first head of last state enc_self_attns')
showgraph(enc_self_attns)
print('first head of last state dec_self_attns')
showgraph(dec_self_attns)
print('first head of last state dec_enc_attns')
showgraph(dec_enc_attns)
```
| github_jupyter |
# Basic Regression Analysis with Time Series Data
## The Nature of Time Series Data
One key feature that distinguishs time series data from cross-sectional data is, temporal ordering. And another one, that the OLS estimators are considered to be $r.v.$s while the time series data set, is considered as a ***realization*** of a **stochastic process**. And since time is irreversible we can only obtain one **realization**. The **sample size** for a time series data set is the *number of time periods* over which we observe the variables of interest.
## Examples of Time Series Regression Models
### Static Models
Suppose that we have time series data available on two variables, $y$ and $z$, where $y_t$ and $z_t$ are dated contemporaneously (marked with same date). Then, we have a typical ***static model*** relating $y$ to $z$:
$$\DeclareMathOperator*{\argmin}{argmin}
\DeclareMathOperator*{\argmax}{argmax}
\DeclareMathOperator*{\plim}{plim}
\newcommand{\ffrac}{\displaystyle \frac}
\newcommand{\d}[1]{\displaystyle{#1}}
\newcommand{\space}{\text{ }}
\newcommand{\bspace}{\;\;\;\;}
\newcommand{\bbspace}{\;\;\;\;\;\;\;\;}
\newcommand{\QQQ}{\boxed{?\:}}
\newcommand{\void}{\left.\right.}
\newcommand{\Tran}[1]{{#1}^{\mathrm{T}}}
\newcommand{\CB}[1]{\left\{ #1 \right\}}
\newcommand{\SB}[1]{\left[ #1 \right]}
\newcommand{\P}[1]{\left( #1 \right)}
\newcommand{\abs}[1]{\left| #1 \right|}
\newcommand{\norm}[1]{\left\| #1 \right\|}
\newcommand{\given}[1]{\left. #1 \right|}
\newcommand{\using}[1]{\stackrel{\mathrm{#1}}{=}}
\newcommand{\asim}{\overset{\text{a}}{\sim}}
\newcommand{\RR}{\mathbb{R}}
\newcommand{\EE}{\mathbb{E}}
\newcommand{\II}{\mathbb{I}}
\newcommand{\NN}{\mathbb{N}}
\newcommand{\ZZ}{\mathbb{Z}}
\newcommand{\QQ}{\mathbb{Q}}
\newcommand{\PP}{\mathbb{P}}
\newcommand{\AcA}{\mathcal{A}}
\newcommand{\FcF}{\mathcal{F}}
\newcommand{\AsA}{\mathscr{A}}
\newcommand{\FsF}{\mathscr{F}}
\newcommand{\dd}{\mathrm{d}}
\newcommand{\I}[1]{\mathrm{I}\left( #1 \right)}
\newcommand{\N}[1]{\mathcal{N}\left( #1 \right)}
\newcommand{\Exp}[1]{\mathrm{E}\left[ #1 \right]}
\newcommand{\Var}[1]{\mathrm{Var}\left[ #1 \right]}
\newcommand{\Avar}[1]{\mathrm{Avar}\left[ #1 \right]}
\newcommand{\Cov}[1]{\mathrm{Cov}\left( #1 \right)}
\newcommand{\Corr}[1]{\mathrm{Corr}\left( #1 \right)}
\newcommand{\ExpH}{\mathrm{E}}
\newcommand{\VarH}{\mathrm{Var}}
\newcommand{\AVarH}{\mathrm{Avar}}
\newcommand{\CovH}{\mathrm{Cov}}
\newcommand{\CorrH}{\mathrm{Corr}}
\newcommand{\ow}{\text{otherwise}}y_t = \beta_0 + \beta_1 z_t + u_t,\bspace t = 1,2,\dots, n$$
Usually, the static model is postulated when a change in $z$ at time $t$ is believed to have an immediate effect on $y$: $\Delta y_t = \beta_1 \Delta z_t$, when $\Delta z_t = 0$. Another possible reason is we want to inspect the tradeoff between $y$ and $z$.
### Finite Distributed Lag Models
In a finite distributed lag (***FDL***) model, one or more variables with a lag can affect $y$. An **FDL** of order $k$ can be written as
$$y_t = \alpha_0 + \delta_0 z_t + \delta_1 z_{t-1} + \cdots + \delta_k z_{t-k} + u_t$$
Here, $\delta_0$ is called the ***impact propensity*** or ***impact multiplier***. And the ***lag distribution*** can be obtained by graphing the $\delta_j$ as a function of $j$, and we plot the graph uses estimated values.
Given a permanent increase in $z$, we find that $y$ also increases permanently by $\sum_{i=0}^k \delta_i$ multiply that increase in $z$, after at most $k$ periods. We call this permanent increase in $y$, the ***long-run propensity*** (***LRP***), or ***long-run multiplier***. The **LRP** is often of interest in distributed lag models.
We can transfer the **FDL** model to **static** model by setting the lagged variables to $0$. Sometimes, the primary purpose assuming this model is to test whether $z$ has a lagged effect on $y$. We also omit $z_t$, the **impact propensity** occasionally.
And for any time horizon $h$, we can define the ***cumulative effect*** as $\delta_0 + \delta_1 + \cdots + \delta_h$, as the change in the expected outcome $h$ periods after a permanent, one unit increase in $z$. And the estimated **cumulative effect** can be plotted as a function of $h$. Then **LRP** can be written as
$$\text{LRP} = \delta_0 + \delta_1 + \cdots + \delta_k$$
Multicollinearity in **FDL** of order $k$ happens a lot. Estimating individual $\delta_j$ precisely would be hard. $\text{LRP}$, however, can often be estimated nicely.
**e.f.1**
Suppose that $\text{int}_t = 1.6 + 0.48 \text{inf}_t - 0.15\text{inf}_{t-1} + 0.32\text{inf}_{t-2} + u_t$ where $\text{int}$ is an interest rate and $\text{inf}$ is the inflation rate. What are the impact and long-run propensities?
>$0.48$ and $0.48-0.15+0.32 = 0.65$
***
### A Convention about the Time Index
We can always start the time index at $t=1$. As for the initial observation, though the dependent variable at $t=1$ should be related to $z_{1}$, $z_0$ and $z_{-1}$. These three are the initial values in our sample.
## Finite Sample Properties of OLS under Classical Assumptions
### Unbiasedness of OLS
$Assumption$ $\text{TS}.1$
$\bspace$Linear in Parameters
>The stochastic process $\CB{\P{x_{t1},x_{t2}, \dots, x_{tk} ,y_t}: t = 1,2,\dots,n}$ follows the linear model:
>
>$$y_t = \beta_0 + \beta_1 x_{t1} + \cdots + \beta_k x_{tk} + u_t$$
>
>where $\CB{u_t:t=1,2,\dots,n}$ is the sequence of errors or disturbances. Here, $n$ is the number of observations (time periods).
Here, $x_{tj}$ denotes the observation at time period $t$, of $j^{\text{th}}$ **explanatory variables**, or **regressors**, or **independent variables**. And just like the terminology in cross-sectional regression, we call $y_t$ the **dependent variable**, **explained variable**, or **regressand**.
And we should think of $\text{TS}.1$ essentially, as same as $\text{MLR}.1$. Let $\mathbf{x}_t = \P{x_{t1}, x_{t2}, \dots,x_{tk}}$ denote the set of all independent variables in the equation at time $t$. Further let $\mathbf{X}$ denotes the collection of all independent variables for all time periods. $X$ can be thought as a matrix with $n$ rows and $k$ columns.
The $t^{\text{th}}$ row of $\mathbf{X}$ is $\mathbf{x}_t$, consisting of all independent variables for time period $t$.
$Assumption$ $\text{TS}.2$
$\bspace$No Perfect Collinearity
>In the sample (and therefore in the underlying time series process), no independent variable is constant nor a *perfect linear* combination of the others.
The issues are essentially same with cross-sectional data.
$Assumption$ $\text{TS}.3$
$\bspace$Zero Conditional Mean
>For each $t$, the expected value of the error $u_t$, given the explanatory variables for *all* time periods, is $0$. Mathematically,
>
>$$\Exp{u_t\mid \mathbf{X}} = 0,\bspace t = 1,2,\dots,n$$
And if $u_t$ is independent of $\mathbf{X}$ and $\Exp{u_t} = 0$, then this assumption automatically holds. One special condition of this assumption, are the ***contemporaneously exogenous*** explanatory variables. In conditional mean terms, we write
$$\Exp{u_t\mid x_{t1},x_{t2},\dots x_{tk}} = \Exp{u_t\mid \mathbf{x}_t} = 0 \Rightarrow \Corr{x_{tj},u_t} = 0,\forall\; j$$
The complete assumption $\text{TS}.3$ is called the ***strict exogeneity***, where $u_t$ must be uncorrelated with $x_{sj}$ where $s\neq t$.
$Remark$
>**Exogeneity** can gurantee the consistency while unbiasedness requires **strict exogeneity**.
>
>And another way to interpret $\text{TS}.3$ is that the average value of $u_t$ is unrelated to the independent variables in all time periods. It really doesn't matter whether there's correlation in the independent variables or in the $u_t$ across time.
***
How would $\text{TS}.3$ fails? Two leading candidates for failure are *omitted variables* and *measurement error* in some of the regressors. And to keep the **strict exogeneity** assumption in **static** regression model, we need to make sure that $u_t$ is uncorrelated with not just $z_t$, but also the past and future value of $z$. This implies that
- $z$ has no lagged effect on $y$, otherwise the model is wrong.
- Changes in error term $u_t$ today, will have NO chance to cause future changes in $z$. There's no feedback from $y$ to future values of $z$.
And in **finite distributed lag** model, the second implication keeps. Feedback from $u$ to future $z$ should be ruled out.
**e.f.2**
In the **FDL** model $y_t = \alpha_0 + \beta_0 z_t + \beta_1 z_{t-1} +u_t$, what to assume about the sequence $\CB{z_0,z_1,\dots,z_n}$ in order for $\text{TS}.3$ to hold?
>The explanatory variables are $x_{t1} = z_t$ and $x_{t2} = z_{t-1}$. The absence of perfect collinearity means that these CANNOT be constant, and there CANNOT be an exact linear relationship between them in the sample.
***
$Theorem.1$ Unbiasedness of OLS
>Under assumptions $\text{TS}.1$ through $\text{TS}.3$, the OLS estimators are unbiased, conditional on $\mathbf X$, and therefore unconditionally as well: $\Exp{\hat\beta_j} = \beta_j$, $j = 0,1,\dots,k$
### The Variances of the OLS Estimators and the Gauss-Markov Theorem
$Assumption$ $\text{TS}.4$
$\bspace$Homoskedasticity
>Conditional on $\mathbf X$, the variance of $u_t$ is the same for all $t$: $\Var{u_t\mid \mathbf X} = \Var{u_t} = \sigma^2$, for $t=1,2,\dots,n$
And if not, we say that the errors are ***heteroskedastic***. Homoskedasticity says two things
- $u_t$ and $\mathbf{X}$ are independent
- $\Var{u_t}$ is a constant over time
$Assumption$ $\text{TS}.5$
$\bspace$No serial correlation
>Conditional on $\mathbf{X}$, the errors in two different time periods are uncorrelated: $\Corr{u_t,u_s\mid \mathbf{X}} = 0$, $\forall t\neq s$
And currently we'll only consider this assumption as $\Corr{u_t,u_s} = 0$, $\forall t\neq s$. When it's failed, we say the errors suffer from ***serial correlation***, or ***autocorrelation***. And also note that this assumption rules out the temporal correlation only in the error term, not the independent variables.
You may ask why this is not assumed in cross-sectional observations. That's because the random sampling assumption, which guarantee that $u_i$ and $u_h$ are independent for any two observations $i$ and $h$. And actually if random sampling assumption is violated, this could be an ideal substitute.
$\text{TS}.1$ through $\text{TS}.5$ are the **Gauss-Markov assumptions** for time series applications.
$Theorem.2$ OLS Sampling Variances
> Under the time series Gauss-Markov Assumptions $\text{TS}.1$ through $\text{TS}.5$, the variance of $\hat\beta_j$, conditional on $\mathbf{X}$, is
>
>$$\Var{\hat\beta_j\mid \mathbf X} = \ffrac{\sigma^2}{\text{SST}_j \P{1-R_j^2}},\bspace j=1,2,\dots,k$$
>
>where $\text{SST}_j$ is the total sum of squares of $x_{tj}$ and $R_j^2$ is the $R$-squared from the regression of $x_j$ on other independent variables.
This theorem and its proof are just like thoes in cross-sectional case.
$Theorem.3$ Unbiased estimation of $\sigma^2$
> Under $\text{TS}.1$ through $\text{TS}.5$, the estimator $\hat\sigma^2 = \ffrac{\text{SSR}}{df}$ is an unbiased estimator of $\sigma^2$, where $df = n-k-1$
$Theorem.4$ Gauss-Markov Theorem
>Under $\text{TS}.1$ through $\text{TS}.5$, the OLS estimators are the best linear unbiased estimators conditional on $\mathbf X$.
**e.f.3**
In the **FDL** model $y_t = \alpha_0 + \delta_0 z_t + \delta_1 x_{t-1} + u_t$, explain the multicollinearity in the explanatory variables.
>Suppose that $\CB{z_t}$ moves slowly over time, then $z_t$ and $z_{t-1}$ can be highly correlated, as can be seen in logs of many economic time series.
### Inference under the Classical Linear Model Assumptions
$Assumption$ $\text{TS}.6$
$\bspace$Normality
> The errors $u_t$ are independent of $X$ and are $i.i.d.$ of $\N{0,\sigma^2}$
Note that this assumption implies $\text{TS}.3$ through $\text{TS}.5$. And taking all assumptions together, we have
$Theorem.5$ Normal sampling distributions
>Under $\text{TS}.1$ through $\text{TS}.6$, the $\text{CLM}$ assumptions for time series, the OLS estimators are **normally distributed**, *conditional* on $\mathbf X$. Further, under the null hypothesis, each $t$ statistic has a $t$ distribution, and each $F$ statistic has an $F$ distribution. The usual construction of confidence intervals is also valid.
## Functional Form, Dummy Variables, and Index Numbers
We use natural logarithm often, that shows time series regression with constant percentage effects, in **static** or **lag** models.
And in **FDL**, **impact multiplier** $\delta_0$ is also called the ***short-run elasticity***, which is the immediate percentage change in $y_t$ given a $1\%$ increase in $z_t$. And similarly, **long-run multiplier** changes into ***long-run elasticity***.
Dummy variables is also very useful since it can indicate whether certain event happened at time $t$. They are key to ***event study***. And one more thing to notice is the ***index number***, which is usually called index. Like S&P index, they aggregate a vast amount of information into a single quantity, comparing a predetermined ***base period*** and the ***base value*** at that time.
## Trends and Seasonality
### Characterizing Trending Time Series
One popular formulation with trending behavior is to write the series $\CB{y_i}$ as
$$y_t = \alpha_0 + \alpha_1 t + e_t,\bspace t=1,2,\dots$$
where, in the simplest case, $\CB{e_t}$ is an $i.i.d.$ sequence with $\Exp{e_t} = 0$ and $\Var{e_t} = \sigma_t^2$ and we call this ***linear time trend***. We'll interpret $\alpha_1$ as, *holding all other factors fixed, the change in $y_i$ from one period to the next due to the passage of time*.
This **linear time trend** implies that if $\Delta e_t = 0$, $\Delta y_t = y_t - y_{t-1} = \alpha1 + \Delta e_t = \alpha_1$. And another way to think this is that *the average value of this sequence is a linear function of time*: $\Exp{y_t} = \alpha_0 + \alpha_1 t$; and as for the variance we have $\Var{y_t} = \Var{e_t} = \sigma_e^2$.
**e.f.4**
Can a **linear trend** with $\alpha_1<0$ be realistic for all future time periods?
>It depends. It'll be more and more negative as $t$ gets larger. For some $y_t$ which can never be negative, a linear time trend with a negative trend coefficient cannot represent it in all future time periods.
***
Another typical trend is the ***exponential trend*** where we would write
$$\log\P{y_t} = \beta_0 + \beta_1 t + e_t,\bspace t =1,2,\dots$$
To interpret first we notive the approximation for small changes
$$\Delta \log\P{y_t} = \log\P{y_t} - \log\P{y_{t-1}} \approx\ffrac{y_t - y_{t-1}}{y_{t-1}}$$
The right hand side is called the ***growth rate*** in $y$ from period $t-1$ to period $t$. And if further assumed that $\Delta e_t = 0$, we have $\forall\,t$, $\Delta \log\P{y_t} = \beta_1$.
### Using Trending Variables in Regression Analysis
***Spurious regression problem*** would happen if there's a relationship between two or more trending variables. And to eliminate this problem we can simply add the time trend. And commonly, adding trends $t$ and $t^2$ would be enough.
$Remark$
>We'll include a trend when
>
>- if the dependent variable displays an obvious trending behaviour
>- if both the dependent and some independent variables have trends
>- if only some of the independent variables have trends, and *only when a trend is added* their effect on the dependent variable will be visible
### A Detrending Interpretation of Regressions with a Time Trend
Including a time trend is like detrending the original data series before regression. For instance we first obtain the fitted equation
$$\hat y_t = \hat \beta_0 + \hat \beta_1 x_{t1}+ \hat\beta_2 x_{t2} + \hat\beta_3 t$$
Similar to what we did in Chap_03, we can apply three-step OLS regression. $x_{t2}$ on $t$, then $x_{t2}$ on $x_{t1}$ then $y$ on $r_{t2}$.
$\P{1}$ Regress each of $y_t$, $x_{t1}$ and $x_{t2}$ on a constant and the time trend $t$ and save the residuals as $\ddot y_t$, $\ddot x_{t1}$ and $\ddot x_{t2}$.
$$
\begin{align}
y_t &= \hat\alpha_0 + \hat\alpha_1 t + \ddot y_t \\
x_{t1} &= \hat\eta_0 + \hat\eta_1 t +\ddot x_{t1}\\
x_{t2} &= \hat\xi_0 + \hat\xi_1 t + \ddot x_{t2}
\end{align}$$
Then $\ddot y_t$, $\ddot x_{t1}$ and $\ddot x_{t2}$ can be regarded as the detrended variables.
$\P 2$ Run the regression of $\ddot y$ on $\ddot x_{t1}$ and $\ddot x_{t2}$. This regression exactly yields $\hat\beta_1$ and $\hat\beta_2$. Here's a proof for one-variable case.
$Proof$ case: $k=1$
>Write the model as $y_t = \beta_0 + \beta_1 x_{t1} + \beta_2 t + u_t$. To detrend, first we have $y_t = \hat\alpha_0 + \hat\alpha_1 t + \ddot y_t$ and $x_{t1} = \hat\eta_0 + \hat\eta_1 t + \ddot x_{t1}$. Then regress the equation
>
>$$\ddot y_t = \hat\theta_1 \ddot x_{t1} + v_t \Rightarrow \hat\theta_1 = \ffrac{\sum\limits_{t=1}^{n} \ddot x_{t1} \ddot y_t}{\sum\limits_{t=1}^{n} \ddot x_{t1}^2}$$
>
>Then using the two-step OLS estimation, we have $x_{t1} = \hat\delta_0 + \hat\delta_1 t + \hat r_{t1}$ and $y_t = \hat\beta_0 + \hat\beta_1 \hat r_{t1} + w_t$. Then,
>
>$$\hat\beta_1 = \ffrac{\sum\limits_{t=1}^n \hat r_{t1} y_t}{\sum\limits_{t=1}^n \hat r_{t1}^2}$$
>
>Remember that $\sum\limits_{t=1}^n t\cdot \hat r_{t1} = 0$ and notice that $\hat r_{t1} = \ddot x_{t1}$, then
>
>$$\hat\beta_1 = \ffrac{\sum\limits_{t=1}^n \hat r_{t1} \P{\hat\alpha_0 + \hat\alpha_1 t + \ddot y_t}}{\sum\limits_{t=1}^n \hat r_{t1}^2} = \ffrac{\sum\limits_{t=1}^n \ddot x_{t1}\ddot y_t}{\sum\limits_{t=1}^n \ddot x_{t1}^2} + \alpha_0 \ffrac{\sum\limits_{t=1}^n \hat r_{t1}}{\sum\limits_{t=1}^n \hat r_{t1}^2} + \alpha_1 \ffrac{\sum\limits_{t=1}^n \hat r_{t1} \cdot t }{\sum\limits_{t=1}^n \hat r_{t1}^2} = \hat\theta_1$$
So we may draw the conclusion that *the OLS coefficients in a regression **including a trend** are the **same** as the coefficients in a regression **without a trend** but where **all** the variables have been **detrended before the regression***.
### Computing $R$-Squared when the Dependent Variable is Trending
Usually it'll be higher, comparing with typical $R$-squareds for cross-sectional data, expecially when the dependent variable is trending.
The adjusted $R$-squared is written as $\bar R^2 = 1 - \ffrac{\hat\sigma_u^2}{\hat\sigma_y^2}$, where $\hat \sigma_u^2$ is the **unbiased estimator of the error variance**, $\hat\sigma_y^2 = \ffrac{\text{SST}}{n-1} = \ffrac{\sum_{t=1}^n \P{y_t - \bar y}^2}{n-1}$. With these formulas we can easily calculate the **error variance** *given that a time trend is included in the regression.* However, $\ffrac{\text{SST}}{n-}$ may overestimate the variance in $y_t$, because it does NOT account for the trend in $y_t$.
And if there's any polynomial trend, we can calculate the $R$-squared of $y = \beta_0 + \beta_1 x_{t1} + \beta_2 x_{t2} + \beta_3 t + u_t$ by
1. detrend the dependent variable: regress $y_t$ on $t$ to obtain the residuals $\ddot y_t$
2. regress the residual on the independent variables: $\ddot y_t$ on $x_{t1}$, $x_{t2}$ and $t$
3. $R^2 = 1 - \ffrac{\text{SSR}}{\sum_{t=1}^n \ddot y_t^2}$ where $\text{SSR}$ is identical to the sum of squared residuals from the original model
We prefer using this because it's smaller and has net out the effect of the time trend, based on the fact that $\sum_{t=1}^n \ddot y_t^2 \leq \sum_{t=1}^n \P{y_t - \bar y}^2$. And the corresponding adjusted $R$-squared is $\bar R^2 = 1 - \ffrac{\frac{1}{n-4}\text{SSR}}{\frac{1}{n-2}\sum_{t=1}^n \ddot y_t^2}$ where $n-4$ is the $df$ of the original model and $n-2$ is the $df$ of the restricted model.
***
One more thing, in computing the $R$-squared form of an $F$ statistic for testing multiple hypotheses, usual $R$-squared without any detrending would be enough.
### Seasonality
Some data series display seasonal patterns, and are often ***seasonally adjusted*** before they are reported for public use, meaning the seasonal factors are removed. However there's time when we work with seasonally unadjusted data and that's the time we'll add a set of ***seasonal dummy variables*** to account for seasonality in the dependent variable, the independent variables, or both. We call this ***deseasonalizing*** the data and we'll write
$$y_t = \beta_0 + \beta_1 x_{t1} + \beta_2 x_{t2} + \cdots + \beta_k x_{tk} + u_t\\
\Downarrow\\
y_t = \beta_0 + \beta_1 x_{t1} + \beta_2 x_{t2} + \cdots + \beta_k x_{tk} + u_t + \delta_1 \text{Jan}_t + \delta_2 \text{Feb}_t + \cdots + \delta_{11} \text{Nov}_t$$
to eliminate the monthly effect to the model. Note that here we would put only $11$ months rather than $12$ to avoid the... some kind of trap, with $\text{Dec}$ as the base month, $\beta_0$ is the intercept for December. Besides, thougth $F$ test, $\delta_1 = \cdots = \delta_{11} = 0$, we can determine whether there's seasonality in $y_t$ or not.
Further discussions are skipped.
***
**e.f.5**
What's the intercept for March? And why the seasonal dummy variables satisfy the strict exogeneity assumption?
>For March, its intercept is $\mathbf \beta_0 + \delta_3$. Seasonal dummy variables are strictly exogenous because they follow a deterministic pattern. For example, the months do not change based upon whether either the explanatory variables or the dependent variable change.
***
| github_jupyter |
Notebook prepared by Mathieu Blondel (lecture 1) and Pierre Ablin (lecture 2).
# Lecture 1
## Ridge regression
```
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
# Load the Boston housing dataset.
X_boston, y_boston = load_boston(return_X_y=True)
# Split into 60% training, 20% validation and 20% test.
X_boston_tr, X_rest, y_boston_tr, y_rest = \
train_test_split(X_boston, y_boston, test_size=0.4, random_state=0)
X_boston_val, X_boston_te, y_boston_val, y_boston_te = \
train_test_split(X_rest, y_rest, test_size=0.5, random_state=0)
```
**Exercise 1.** Implement the analytical solution of ridge regression $(X^\top X + \alpha I) w = X^\top y$ (see [slides](https://data-psl.github.io/lectures2021/slides/05_optimization_linear_models)) using [scipy.linalg.solve](https://docs.scipy.org/doc/scipy/reference/generated/scipy.linalg.solve.html). Compute the solution on the training data. Make sure that the gradient at the solution is zero (up to machine precision).
```
from scipy.linalg import solve
def ridge_regression_solution(X, y, alpha):
# Write your function here
return w
def ridge_regression_gradient(w, X, y, alpha):
# Write your function here
return gradient
w_star = ridge_regression_solution(X_boston_tr, y_boston_tr, alpha=0.1)
gradient = ridge_regression_gradient(w_star, X_boston_tr, y_boston_tr, alpha=0.1)
np.sqrt(np.sum(gradient ** 2))
```
**Exercise 2.** Train the models for several possible values of alpha (see below). Plot the mean squared error on the test set as a function of alpha. Use the validation data to find the best alpha and display it on the graph using a circle.
```
alphas = np.logspace(-3, 3, 20)
```
**Bonus exercise.** Implement a scikit-learn compatible estimator class (with fit and predict methods). Compare that you obtain the same results as `sklearn.linear_model.Ridge(fit_intercept=False)`.
```
from sklearn.base import BaseEstimator, RegressorMixin
class MyRidge(BaseEstimator, RegressorMixin):
def __init__(self, alpha=1.0):
self.alpha = alpha
def fit(self, X, y):
# Write your code here
return self
def predict(self, X):
# Write your code here
return
from sklearn.linear_model import Ridge
print(MyRidge().fit(X_boston_tr, y_boston_tr).predict(X_boston_te)[:10])
print(Ridge(fit_intercept=False).fit(X_boston_tr, y_boston_tr).predict(X_boston_te)[:10])
```
## Logistic regression
```
from sklearn.datasets import load_iris
X_iris, y_iris = load_iris(return_X_y=True)
# Keep only two classes for this exercise.
X_iris = X_iris[y_iris <= 1]
y_iris = y_iris[y_iris <= 1]
```
**Exercise 3.** Make a function that computes
$$
\text{softplus}(u) = \log(1 + e^u)
$$
and notice that its derivative is
$$
(\text{softplus}(u))' = \frac{e^u}{1 + e^u} = \frac{1}{1 + e^{-u}} = \text{sigmoid}(u).
$$
Using the finite difference formula $f'(u) \approx \frac{f(u + \epsilon) - f(u)}{\epsilon}$ where epsilon is small value (e.g. 10^-6), check that the derivative of softplus is indeed the sigmoid.
```
import numpy as np
from scipy.special import expit as sigmoid
print(softplus(3))
print(finite_difference(softplus, 3))
print(sigmoid(3))
```
**Exercise 4.**
Make a function that computes the likelihood
$$
\text{likelihood}(u_i, y_i) = y_i \log \text{sigmoid}(u_i) + (1-y_i) \log (1-\text{sigmoid}(u_i))
$$
where $u_i = \mathbf{w}^\top \mathbf{x}_i$.
Using
$$
\log \text{sigmoid}(u) = -\text{softplus}(-u)
$$
and
$$
\log(1 - \text{sigmoid}(u)) = -\text{softplus}(u)
$$
make a function that computes the derivative of $\text{likelihood}(u_i, y_i)$ with respect to $u_i$. Check the result by finite difference. Be careful of signs!
```
def likelihood(u_i, y_i):
# Write function here
return
def likelihood_derivative(u_i, y_i):
# Write function here
return
print(likelihood_derivative(3, 1))
print(finite_difference(likelihood, 3, 1))
```
**Exercise 5.** Write a function that implements the penalized objective function
$$
L(\mathbf{w})
= -\sum_{i=1}^n y_i \log \text{sigmoid}(\mathbf{w}^\top \mathbf{x}_i) + (1-y_i) \log (1-\text{sigmoid}(\mathbf{w}^\top \mathbf{x}_i)) + \frac{\alpha}{2} \|\mathbf{w}\|^2
$$
and another function that computes its gradient. Reuse `likelihood(u_i, y_i)` and `likelihood_derivative(u_i, y_i)` (you can use a for loop). Check that the gradient is correct using finite differences.
```
def objective_value(w, X, y, alpha):
# Write function here
return
def objective_gradient(w, X, y, alpha):
# Write function here
return
def finite_difference_gradient(func, w, *args, eps=1e-6):
gradient = np.zeros_like(w)
for j in range(len(w)):
e_j = np.zeros(len(w))
e_j[j] = 1
gradient[j] = (func(w + eps * e_j, *args) - func(w, *args)) / eps
return gradient
n_samples, n_features = X_iris.shape
w = np.random.randn(n_features)
alpha = 0.1
print(objective_gradient(w, X_iris, y_iris, alpha))
print(finite_difference_gradient(objective_value, w, X_iris, y_iris, alpha))
```
**Exercise 6.** Implement gradient descent. Check that the objective value is decreasing. Plot the objective value as a function of the number of iterations.
```
def gradient_descent(value_function, gradient_function, w_init, *args,
step_size=1e-4, num_iterations=1000):
values = []
w = w_init
# Write gradient descent iteration here.
return values
n_samples, n_features = X_iris.shape
w_init = np.random.randn(n_features)
values = gradient_descent(objective_value, objective_gradient, w, X_iris, y_iris, alpha)
import matplotlib.pyplot as plt
plt.figure()
plt.plot(np.arange(len(values)), values)
plt.show()
```
**Exercise 7.** Rewrite `objective_value` and `objective_gradient` without for loop. Check the correctness of your implementation against the version with a for loop.
```
def objective_value_no_loop(w, X, y, alpha):
# Write your code here
return
def objective_gradient_no_loop(w, X, y, alpha):
# Write your code here
return
n_samples, n_features = X_iris.shape
w = np.random.randn(n_features)
alpha = 0.1
print(objective_value(w, X_iris, y_iris, alpha))
print(objective_value_no_loop(w, X_iris, y_iris, alpha))
print(objective_gradient(w, X_iris, y_iris, alpha))
print(objective_gradient_no_loop(w, X_iris, y_iris, alpha))
```
Time the two implementations.
```
%time objective_value(w, X_iris, y_iris, alpha)
%time objective_value_no_loop(w, X_iris, y_iris, alpha)
```
# Lecture 2
# Stochastic gradient descent
We will focus on the Boston dataset and ridge regression. We will start by scaling the dataset
```
from sklearn.preprocessing import StandardScaler
X = StandardScaler().fit_transform(X_boston)
y = y_boston - y_boston.mean()
y /= np.std(y_boston)
```
**Exercise 8.** Write a function that computes the stochastic gradient of ridge regression
$$
L(\mathbf{w}) = \frac{1}{n} \|\mathbf{y} - \mathbf{X} \mathbf{w}\|^2
+ \frac{\alpha}{2} \|\mathbf{w}\|^2
$$
(notice the 1/n factor).
Check that the mean of the stochastic gradients gives the gradient.
```
def ridge_objective(w, X, y, alpha):
# Write your code here
return
def ridge_gradient(w, X, y, alpha):
# Write your code here
return
def stochastic_gradient(w, i, X, y, alpha):
# Write your code here
return
```
**Exercise 9.** Write a function that implements stochastic gradient descent. Implement two rules for sampling the index: cyclic, and at random. Compare the convergence of both algorithms. What is the role of the step size?
You should especially look at the convergence speed and the value at which the algorithm plateaus.
```
def stochastic_gradient_descent(value_function, gradient_function, w_init, idx_list, *args,
step_size=1e-4, num_iterations=1000):
values = []
w = w_init
# Write SGD code here
return values
```
# L-BFGS
L-BFGS is the go-to second order method. It is already implemented in `scipy`:
```
from scipy.optimize import fmin_l_bfgs_b
```
**Exercise 10.** Use the L-BFGS code to optimize the logistic regression on the Iris dataset. Compare it with your gradient descent.
Hint: in order to store the function values, you can use the callback function in `fmin_l_bfgs_b`. Don't forget to read the documentation !
```
class callback(object):
def __init__(self):
self.values = []
def __call__(self, w):
self.values.append(objective_value_no_loop(w, X_iris, y_iris, alpha))
```
| github_jupyter |
# Deep Q-Network (DQN)
---
In this notebook, you will implement a DQN agent with OpenAI Gym's LunarLander-v2 environment.
### 1. Import the Necessary Packages
```
import gym
import random
import torch
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
%matplotlib inline
EXPERIMENT_NAME = "rainbow"
EXPERIMENT_DETAIL = "NoNoise"
```
### 2. Instantiate the Environment and Agent
Initialize the environment in the code cell below.
```
env = gym.make('LunarLander-v2')
env.seed(0)
print('State shape: ', env.observation_space.shape)
print('Number of actions: ', env.action_space.n)
```
Before running the next code cell, familiarize yourself with the code in **Step 2** and **Step 3** of this notebook, along with the code in `dqn_agent.py` and `model.py`. Once you have an understanding of how the different files work together,
- Define a neural network architecture in `model.py` that maps states to action values. This file is mostly empty - it's up to you to define your own deep Q-network!
- Finish the `learn` method in the `Agent` class in `dqn_agent.py`. The sampled batch of experience tuples is already provided for you; you need only use the local and target Q-networks to compute the loss, before taking a step towards minimizing the loss.
Once you have completed the code in `dqn_agent.py` and `model.py`, run the code cell below. (_If you end up needing to make multiple changes and get unexpected behavior, please restart the kernel and run the cells from the beginning of the notebook!_)
You can find the solution files, along with saved model weights for a trained agent, in the `solution/` folder. (_Note that there are many ways to solve this exercise, and the "solution" is just one way of approaching the problem, to yield a trained agent._)
```
# Hyperparameters
hyperparams = {
'seed': 101,
'buffer_size': int(1e5),
'batch_size': 32,
'start_since': int(8e3),
'gamma': 0.99,
'target_update_every': int(32e2),
'tau': 1,
'lr': 1e-4,
'weight_decay': 0,
'update_every': 4,
'priority_eps': 1e-5,
'a': 0.5,
'n_multisteps': 3,
'v_min': -100,
'v_max': 300,
'clip': None,
'n_atoms': 51,
'initial_sigma': 0.1,
'linear_type': 'linear'
}
# Training Parameters
train_params = {
'n_episodes': 2000, 'max_t': 1000,
'eps_start': 1., 'eps_end': 0.01, 'eps_decay': 0.9975,
'beta_start': 0.4, 'beta_end': 1.0
}
from dqn_agent import Agent
agent = Agent(state_size=8, action_size=4, **hyperparams)
```
### 3. Train the Agent with DQN
Run the code cell below to train the agent from scratch. You are welcome to amend the supplied values of the parameters in the function, to try to see if you can get better performance!
```
def dqn(n_episodes=2000, max_t=1000,
eps_start=1.0, eps_end=0.01, eps_decay=0.995,
beta_start=0., beta_end=1.0,
continue_after_solved=True,
save_name="checkpoint_dueling_solved.pth"):
"""Deep Q-Learning.
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of timesteps per episode
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float): minimum value of epsilon
eps_decay (float): multiplicative factor (per episode) for decreasing epsilon
"""
scores = [] # list containing scores from each episode
scores_window = deque(maxlen=100) # last 100 scores
eps = eps_start # initialize epsilon
prioritized = hasattr(agent, 'beta') # if using prioritized experience replay, initialize beta
if prioritized:
print("Priority Used")
agent.beta = beta_start
beta_increment = (beta_end - beta_start) / n_episodes
else:
print("Priority Not Used")
solved = False
epi_str_max_len = len(str(n_episodes))
for i_episode in range(1, n_episodes+1):
state = env.reset()
score = 0
for t in range(max_t):
action = agent.act(state, eps)
next_state, reward, done, _ = env.step(action)
agent.step(state, action, reward, next_state, done)
state = next_state
score += reward
if done:
break
else: # if not done (reached max_t)
agent.memory.reset_multisteps()
scores_window.append(score) # save most recent score
scores.append(score) # save most recent score
eps = max(eps_end, eps_decay*eps) # decrease epsilon
if prioritized:
agent.beta = min(beta_end, agent.beta + beta_increment)
print('\rEpisode {:>{epi_max_len}d} | Current Score: {:>7.2f} | Average Score: {:>7.2f} | Epsilon: {:>6.4f}'\
.format(i_episode, score, np.mean(scores_window), eps, epi_max_len=epi_str_max_len), end="")
if prioritized:
print(' | A: {:>6.4f} | Beta: {:>6.4f}'.format(agent.a, agent.beta), end='')
print(' ', end='')
if i_episode % 100 == 0:
print('\rEpisode {:>{epi_max_len}} | Current Score: {:>7.2f} | Average Score: {:>7.2f} | Epsilon: {:>6.4f}'\
.format(i_episode, score, np.mean(scores_window), eps, epi_max_len=epi_str_max_len), end='')
if prioritized:
print(' | A: {:>6.4f} | Beta: {:>6.4f}'.format(agent.a, agent.beta), end='')
print(' ')
if not solved and np.mean(scores_window)>=200.0:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_window)))
torch.save(agent.qnetwork_local.state_dict(), save_name)
solved = True
if not continue_after_solved:
break
return scores
scores = dqn(**train_params,
continue_after_solved=True,
save_name="experiment_{}_{}_solved.pth".format(EXPERIMENT_NAME, EXPERIMENT_DETAIL))
# plot the scores
plt.rcParams['figure.facecolor'] = 'w'
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
torch.save(agent.qnetwork_local.state_dict(), 'experiment_{}_{}_final.pth'.format(EXPERIMENT_NAME, EXPERIMENT_DETAIL))
agent.qnetwork_local.load_state_dict(torch.load('experiment_{}_{}_final.pth'.format(EXPERIMENT_NAME, EXPERIMENT_DETAIL)))
```
### 4. Watch a Smart Agent!
In the next code cell, you will load the trained weights from file to watch a smart agent!
```
agent.qnetwork_local.noise(False)
for i in range(10):
state = env.reset()
score = 0
for j in range(1000):
action = agent.act(state)
env.render()
state, reward, done, _ = env.step(action)
score += reward
if done:
break
print("Game {} Score: {} in {} steps".format(i, score, j + 1))
agent.qnetwork_local.noise(True)
env.close()
```
### 5. Explore
In this exercise, you have implemented a DQN agent and demonstrated how to use it to solve an OpenAI Gym environment. To continue your learning, you are encouraged to complete any (or all!) of the following tasks:
- Amend the various hyperparameters and network architecture to see if you can get your agent to solve the environment faster. Once you build intuition for the hyperparameters that work well with this environment, try solving a different OpenAI Gym task with discrete actions!
- You may like to implement some improvements such as prioritized experience replay, Double DQN, or Dueling DQN!
- Write a blog post explaining the intuition behind the DQN algorithm and demonstrating how to use it to solve an RL environment of your choosing.
```
def reset_env():
state = torch.from_numpy(env.reset()).unsqueeze(0).cuda()
with torch.no_grad():
p = agent.qnetwork_local(state).softmax(dim=-1)
action = np.argmax(agent.supports.mul(p).sum(dim=-1, keepdim=False).cpu().numpy())
env.render()
p = p.cpu().squeeze().numpy()
supports = agent.supports.cpu().numpy()
plt.rcParams['figure.facecolor'] = 'w'
fig, axes = plt.subplots(2, 2, figsize=(8, 8))
for ax in axes.reshape(-1):
ax.grid(True)
ax.set_ylabel("estimated probability")
ax.set_xlabel("supports")
axes[0, 0].set_title("do nothing")
axes[0, 1].set_title("left engine")
axes[1, 0].set_title("main engine")
axes[1, 1].set_title("right engine")
axes[0, 0].bar(x=supports, height=p[0], width=5)
axes[0, 1].bar(x=supports, height=p[1], width=5)
axes[1, 0].bar(x=supports, height=p[2], width=5)
axes[1, 1].bar(x=supports, height=p[3], width=5)
plt.tight_layout()
return action
def step(action, n_steps):
print(['nothing', 'left', 'main', 'right'][action])
score_gained = 0
for _ in range(n_steps):
state, reward, done, _ = env.step(action)
score_gained += reward
with torch.no_grad():
state = torch.from_numpy(state).unsqueeze(0).cuda()
p = agent.qnetwork_local(state).softmax(dim=-1)
action = np.argmax(agent.supports.mul(p).sum(dim=-1, keepdim=False).cpu().numpy())
env.render()
if done:
print(done)
break
print(score_gained)
p = p.cpu().squeeze().numpy()
supports = agent.supports.cpu().numpy()
plt.rcParams['figure.facecolor'] = 'w'
fig, axes = plt.subplots(2, 2, figsize=(8, 8))
for ax in axes.reshape(-1):
ax.grid(True)
ax.set_ylabel("estimated probability")
ax.set_xlabel("supports")
axes[0, 0].set_title("do nothing")
axes[0, 1].set_title("left engine")
axes[1, 0].set_title("main engine")
axes[1, 1].set_title("right engine")
axes[0, 0].bar(x=supports, height=p[0], width=5)
axes[0, 1].bar(x=supports, height=p[1], width=5)
axes[1, 0].bar(x=supports, height=p[2], width=5)
axes[1, 1].bar(x=supports, height=p[3], width=5)
plt.tight_layout()
return action
action = reset_env()
action = step(action, 50)
env.close()
```
---
| github_jupyter |
# Principle Componet Analysis
`Principal component analysis (PCA)` is a statistical procedure that uses an orthogonal transformation to convert a set of observations of possibly correlated variables into a set of values of linearly uncorrelated variables called principal components. If there are `n` observations with `p` variables, then the number of distinct principal components is `min(n-1,p)`. This transformation is defined in such a way that the first principal component has the largest possible variance (that is, accounts for as much of the variability in the data as possible), and each succeeding component in turn has the highest variance possible under the constraint that it is orthogonal to the preceding components. The resulting vectors are an uncorrelated orthogonal basis set. PCA is sensitive to the relative scaling of the original variables.
### PCA降维
* 目标:提取最有价值的信息(基于方差)<br/>
* 问题:降维后数据失去原来意义(可以保密)
<br/><br/>
### 原理:
* 向量、基:基是正交的(相互垂直、线性无关)<br/>
* 基变换:空间变换
<br/><br/>
### 基变换依据:
* 协方差矩阵:寻找一个一维基,使得所有数据变换为这个基上的坐标表示后,方差值最大. 直观:投影后的投影值尽可能的分散<br/>
* 方差: $Var(a)=\frac{1}{m}\sum_{i=1}^m (a_{i}-{\eta})^2$<br/>
* 协方差(假设均值为0): $Cov(a,b)=\frac{1}{m}\sum_{i=1}^m (a_{i}b_{i})^2$<br/>
* 说明:协方差为0表示两个字段完全独立,表现为两个基正交
<br/><br/>
### 优化目标:
* 将一组N维向量降为K维,目标是选择K个单位正交基,使原始数据变换到这组基上后,各字段两两间协方差为0,字段的方差尽可能大
* `协方差矩阵`:对角线元素为两个字段的方差,其他元素为协方差
* 数学原理:`特征值、特征向量`
* 采取措施:协方差矩阵`对角化`:除对角线外,其他元素化为0,并且对角线上元素按大小从上到下排序
* 数学定理:实对称矩阵(一个n行n列的实对称矩阵一定可以找到n个单位正交特征向量)——实对称阵对角化
```
import numpy as np
import pandas as pd
df = pd.read_csv('./resources/iris.data')
df.head()
df.columns=['sepal_len', 'sepal_wid', 'petal_len', 'petal_wid', 'class']
df.head()
# 查看类别值
df["class"].unique()
# 将特征与类别分开
X = df.iloc[:,0:4].values
y = df.iloc[:,4].values
from matplotlib import pyplot as plt
%matplotlib inline
import math
label_dict = {1: 'Iris-Setosa',
2: 'Iris-Versicolor',
3: 'Iris-Virgnica'}
feature_dict = {0: 'sepal length [cm]',
1: 'sepal width [cm]',
2: 'petal length [cm]',
3: 'petal width [cm]'}
plt.figure(figsize=(15, 12))
for cnt in range(4):
plt.subplot(2, 2, cnt+1)
for lab in ('Iris-setosa', 'Iris-versicolor', 'Iris-virginica'):
plt.hist(X[y==lab, cnt],
label=lab,
bins=10,
alpha=0.3,)
plt.xlabel(feature_dict[cnt])
plt.legend(loc='upper right', fancybox=True, fontsize=8)
plt.tight_layout()
plt.show()
```
### Normalisation
```
from sklearn.preprocessing import StandardScaler
X_std = StandardScaler().fit_transform(X)
print (X_std[:5])
```
### np.mean求均值:
axis = 0 :压缩行,对各列求均值
axis = 1 :压缩列,对各行求均值
```
mean_vec = np.mean(X_std, axis=0)
mean_vec
```
### 协方差矩阵
```
cov_mat = (X_std - mean_vec).T.dot((X_std - mean_vec)) / (X_std.shape[0]-1)
print('Covariance matrix \n%s' %cov_mat)
# using Numpy
cov_mat = np.cov(X_std.T)
print('NumPy covariance matrix: \n%s' %cov_mat)
```
### numpy.linalg.eig计算特征值与特征向量
```
eig_vals, eig_vecs = np.linalg.eig(cov_mat)
print('\nEigenvalues \n%s' %eig_vals)
print('Eigenvectors \n%s' %eig_vecs)
```
### 特征值由大到小排,选择其中最大的2个,然后将对应的2个特征向量分别作为列向量组成` 特征向量矩阵`
```
# Make a list of (eigenvalue, eigenvector) tuples
eig_pairs = [(np.abs(eig_vals[i]), eig_vecs[:,i]) for i in range(len(eig_vals))]
print (eig_pairs)
print ('----------')
# Sort the (eigenvalue, eigenvector) tuples from high to low
eig_pairs.sort(key=lambda x: x[0], reverse=True)
# Visually confirm that the list is correctly sorted by decreasing eigenvalues
print('Eigenvalues in descending order:')
for i in eig_pairs:
print(i[0])
tot = sum(eig_vals)
var_exp = [(i / tot)*100 for i in sorted(eig_vals, reverse=True)]
print (var_exp)
cum_var_exp = np.cumsum(var_exp)
cum_var_exp
plt.figure(figsize=(6, 4))
plt.bar(range(4), var_exp, alpha=0.5, align='center',
label='individual explained variance')
plt.step(range(4), cum_var_exp, where='mid',
label='cumulative explained variance')
plt.ylabel('Explained variance ratio')
plt.xlabel('Principal components')
plt.legend(loc='best')
plt.tight_layout()
plt.show()
matrix_w = np.hstack((eig_pairs[0][1].reshape(4,1),
eig_pairs[1][1].reshape(4,1)))
print('Matrix W:\n', matrix_w)
Y = X_std.dot(matrix_w)
Y.shape
plt.figure(figsize=(6, 4))
for lab, col in zip(('Iris-setosa', 'Iris-versicolor', 'Iris-virginica'),
('blue', 'red', 'green')):
plt.scatter(X[y==lab, 0],
X[y==lab, 1],
label=lab,
c=col)
plt.xlabel('sepal_len')
plt.ylabel('sepal_wid')
plt.legend(loc='best')
plt.tight_layout()
plt.show()
plt.figure(figsize=(6, 4))
for lab, col in zip(('Iris-setosa', 'Iris-versicolor', 'Iris-virginica'),
('blue', 'red', 'green')):
plt.scatter(Y[y==lab, 0],
Y[y==lab, 1],
label=lab,
c=col)
plt.xlabel('Principal Component 1')
plt.ylabel('Principal Component 2')
plt.legend(loc='lower center')
plt.tight_layout()
plt.show()
```
| github_jupyter |
# 📃 Solution for Exercise M3.02
The goal is to find the best set of hyperparameters which maximize the
statistical performance on a training set.
Here again with limit the size of the training set to make computation
run faster. Feel free to increase the `train_size` value if your computer
is powerful enough.
```
import numpy as np
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census.csv")
target_name = "class"
target = adult_census[target_name]
data = adult_census.drop(columns=[target_name, "education-num"])
from sklearn.model_selection import train_test_split
data_train, data_test, target_train, target_test = train_test_split(
data, target, train_size=0.2, random_state=42)
```
You should:
* preprocess the categorical columns using a `OneHotEncoder` and use a
`StandardScaler` to normalize the numerical data.
* use a `LogisticRegression` as a predictive model.
Start by defining the columns and the preprocessing pipelines to be applied
on each columns.
```
from sklearn.compose import make_column_selector as selector
categorical_columns_selector = selector(dtype_include=object)
categorical_columns = categorical_columns_selector(data)
numerical_columns_selector = selector(dtype_exclude=object)
numerical_columns = numerical_columns_selector(data)
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import StandardScaler
categorical_processor = OneHotEncoder(handle_unknown="ignore")
numerical_processor = StandardScaler()
```
Subsequently, create a `ColumnTransformer` to redirect the specific columns
a preprocessing pipeline.
```
from sklearn.compose import ColumnTransformer
preprocessor = ColumnTransformer(
[('cat-preprocessor', categorical_processor, categorical_columns),
('num-preprocessor', numerical_processor, numerical_columns)]
)
```
Finally, concatenate the preprocessing pipeline with a logistic regression.
```
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LogisticRegression
model = make_pipeline(preprocessor, LogisticRegression())
```
Use `RandomizedSearchCV` with `n_iter=20` to find the best set of
hyperparameters by tuning the following parameters of the `model`:
- the parameter `C` of the `LogisticRegression` with values ranging from
0.001 to 10. You can use a log-uniform distribution
(i.e. `scipy.stats.loguniform`);
- the parameter `with_mean` of the `StandardScaler` with possible values
`True` or `False`;
- the parameter `with_std` of the `StandardScaler` with possible values
`True` or `False`.
Once the computation has completed, print the best combination of parameters
stored in the `best_params_` attribute.
```
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import loguniform
param_distributions = {
"logisticregression__C": loguniform(0.001, 10),
"columntransformer__num-preprocessor__with_mean": [True, False],
"columntransformer__num-preprocessor__with_std": [True, False],
}
model_random_search = RandomizedSearchCV(
model, param_distributions=param_distributions,
n_iter=20, error_score=np.nan, n_jobs=-1, verbose=1)
model_random_search.fit(data_train, target_train)
model_random_search.best_params_
```
So the best hyperparameters give a model where the features are scaled but
not centered and the final model is regularized.
Getting the best parameter combinations is the main outcome of the
hyper-parameter optimization procedure. However it is also interesting
to assess the sensitivity of the best models to the choice of those
parameters. The following code, not required to answer the quiz question
shows how to conduct such an analysis for this this pipeline using a
parallel coordinate plot.
We could use `cv_results = model_random_search.cv_results_` to make a
parallel coordinate plot as we did in the previous notebook (you are more
than welcome to try!). Instead we are going to load the results obtained from
a similar search with many more iterations (1,000 instead of 20).
```
cv_results = pd.read_csv(
"../figures/randomized_search_results_logistic_regression.csv")
```
To simplify the axis of the plot, we will rename the column of the dataframe
and only select the mean test score and the value of the hyperparameters.
```
column_name_mapping = {
"param_logisticregression__C": "C",
"param_columntransformer__num-preprocessor__with_mean": "centering",
"param_columntransformer__num-preprocessor__with_std": "scaling",
"mean_test_score": "mean test accuracy",
}
cv_results = cv_results.rename(columns=column_name_mapping)
cv_results = cv_results[column_name_mapping.values()].sort_values(
"mean test accuracy", ascending=False)
```
In addition, the parallel coordinate plot from `plotly` expects all data to
be numeric. Thus, we convert the boolean indicator informing whether or not
the data were centered or scaled into an integer, where True is mapped to 1
and False is mapped to 0.
We also take the logarithm of the `C` values to span the data on a broader
range for a better visualization.
```
column_scaler = ["centering", "scaling"]
cv_results[column_scaler] = cv_results[column_scaler].astype(np.int64)
cv_results['log C'] = np.log10(cv_results['C'])
import plotly.express as px
fig = px.parallel_coordinates(
cv_results,
color="mean test accuracy",
dimensions=["log C", "centering", "scaling", "mean test accuracy"],
color_continuous_scale=px.colors.diverging.Tealrose,
)
fig.show()
```
We recall that it is possible to select a range of results by clicking and
holding on any axis of the parallel coordinate plot. You can then slide
(move) the range selection and cross two selections to see the intersections.
Selecting the best performing models (i.e. above an accuracy of ~0.845), we
observe the following pattern:
- scaling the data is important. All the best performing models are scaling
the data;
- centering the data does not have a strong impact. Both approaches,
centering and not centering, can lead to good models;
- using some regularization is fine but using too much is a problem. Recall
that a smaller value of C means a stronger regularization. In particular
no pipeline with C lower than 0.001 can be found among the best
models.
| github_jupyter |
```
from IPython.core.display import display,HTML
display(HTML('<style>.prompt{width: 0px; min-width: 0px; visibility: collapse}</style>'))
%matplotlib inline
%load_ext autoreload
%autoreload 2
%config Completer.use_jedi=False
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from ipywidgets import interact, Dropdown, IntSlider
from tqdm import tqdm_notebook
train_acl = pd.read_csv('/home/medmantra/Downloads/KneeMR_AI/DB/MRNet-v1.0/MRNet-v1.0/train-acl.csv', header=None,
names=['Case', 'Abnormal'],
dtype={'Case': str, 'Abnormal': np.int64})
train_acl.head()
print(train_acl.shape)
# (1130, 2)
train_acl.Abnormal.value_counts(normalize=True)
# 0 0.815929
# 1 0.184071
# Name: Abnormal, dtype: float64
case = '0000'
mri_coronal = np.load('/home/medmantra/Downloads/KneeMR_AI/DB/MRNet-v1.0/MRNet-v1.0/train/coronal/0000.npy')
mri_axial = np.load('/home/medmantra/Downloads/KneeMR_AI/DB/MRNet-v1.0/MRNet-v1.0/train/axial/0000.npy')
mri_sagittal = np.load('/home/medmantra/Downloads/KneeMR_AI/DB/MRNet-v1.0/MRNet-v1.0/train/sagittal/0000.npy')
print(f'MRI scan on coronal plane: {mri_coronal.shape}')
print(f'MRI scan on axial plane: {mri_axial.shape}')
print(f'MRI scan on sagittal plane: {mri_sagittal.shape}')
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(15, 5))
ax1.imshow(mri_coronal[0, :, :], 'gray');
ax1.set_title('Case 0 | Slice 1 | Sagittal');
ax2.imshow(mri_axial[0, :, :], 'gray');
ax2.set_title('Case 0 | Slice 1 | Axial');
ax3.imshow(mri_sagittal[0, :, :], 'gray');
ax3.set_title('Case 0 | Slice 1 | Coronal');
train_path = '/home/medmantra/Downloads/KneeMR_AI/DB/MRNet-v1.0/MRNet-v1.0/train/'
def load_one_stack(case, data_path=train_path, plane='coronal'):
fpath = '{}/{}/{}.npy'.format(data_path, plane, case)
return np.load(fpath)
def load_stacks(case, data_path=train_path):
x = {}
planes = ['coronal', 'sagittal', 'axial']
for i, plane in enumerate(planes):
x[plane] = load_one_stack(case, plane=plane)
return x
def load_cases(train=True, n=None):
assert (type(n) == int) and (n < 1250)
if train:
case_list = pd.read_csv('/home/medmantra/Downloads/KneeMR_AI/DB/MRNet-v1.0/MRNet-v1.0/train-acl.csv', names=['case', 'label'], header=None,
dtype={'case': str, 'label': np.int64})['case'].tolist()
else:
case_list = pd.read_csv('/home/medmantra/Downloads/KneeMR_AI/DB/MRNet-v1.0/MRNet-v1.0/valid-acl.csv', names=['case', 'label'], header=None,
dtype={'case': str, 'label': np.int64})['case'].tolist()
cases = {}
if n is not None:
case_list = case_list[:n]
for case in tqdm_notebook(case_list, leave=False):
x = load_stacks(case)
cases[case] = x
return cases
cases = load_cases(n=100)
print(cases['0000'].keys())
#dict_keys(['coronal', 'sagittal', 'axial'])
print(cases['0000']['axial'].shape)
print(cases['0000']['coronal'].shape)
print(cases['0000']['sagittal'].shape)
# (44, 256, 256)
# (36, 256, 256)
# (36, 256, 256)
class KneePlot():
def __init__(self, cases, figsize=(15, 5)):
self.cases = cases
self.planes = {case: ['coronal', 'sagittal', 'axial'] for case in self.cases}
self.slice_nums = {}
for case in self.cases:
self.slice_nums[case] = {}
for plane in ['coronal', 'sagittal', 'axial']:
self.slice_nums[case][plane] = self.cases[case][plane].shape[0]
self.figsize = figsize
def _plot_slices(self, case, im_slice_coronal, im_slice_sagittal, im_slice_axial):
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=self.figsize)
ax1.imshow(self.cases[case]['coronal'][im_slice_coronal, :, :], 'gray')
ax1.set_title(f'MRI slice {im_slice_coronal} on coronal plane')
ax2.imshow(self.cases[case]['sagittal'][im_slice_sagittal, :, :], 'gray')
ax2.set_title(f'MRI slice {im_slice_sagittal} on sagittal plane')
ax3.imshow(self.cases[case]['axial'][im_slice_axial, :, :], 'gray')
ax3.set_title(f'MRI slice {im_slice_axial} on axial plane')
plt.show()
def draw(self):
case_widget = Dropdown(options=list(self.cases.keys()),
description='Case'
)
case_init = list(self.cases.keys())[0]
slice_init_coronal = self.slice_nums[case_init]['coronal'] - 1
slices_widget_coronal = IntSlider(min=0,
max=slice_init_coronal,
value=slice_init_coronal // 2,
description='Coronal')
slice_init_sagittal = self.slice_nums[case_init]['sagittal'] - 1
slices_widget_sagittal = IntSlider(min=0,
max=slice_init_sagittal,
value=slice_init_sagittal // 2,
description='Sagittal'
)
slice_init_axial = self.slice_nums[case_init]['axial'] - 1
slices_widget_axial = IntSlider(min=0,
max=slice_init_axial,
value=slice_init_axial // 2,
description='Axial'
)
def update_slices_widget(*args):
slices_widget_coronal.max = self.slice_nums[case_widget.value]['coronal'] - 1
slices_widget_coronal.value = slices_widget_coronal.max // 2
slices_widget_sagittal.max = self.slice_nums[case_widget.value]['sagittal'] - 1
slices_widget_sagittal.value = slices_widget_sagittal.max // 2
slices_widget_axial.max = self.slice_nums[case_widget.value]['axial'] - 1
slices_widget_axial.value = slices_widget_axial.max // 2
case_widget.observe(update_slices_widget, 'value')
interact(self._plot_slices,
case=case_widget,
im_slice_coronal=slices_widget_coronal,
im_slice_sagittal=slices_widget_sagittal,
im_slice_axial=slices_widget_axial
)
def resize(self, figsize):
self.figsize = figsize
plot = KneePlot(cases)
plot.draw()
```
| github_jupyter |
# Table of Contents
<p><div class="lev1 toc-item"><a href="#Neural-Networks-Overview" data-toc-modified-id="Neural-Networks-Overview-1"><span class="toc-item-num">1 </span>Neural Networks Overview</a></div><div class="lev1 toc-item"><a href="#Neural-Network-Representation" data-toc-modified-id="Neural-Network-Representation-2"><span class="toc-item-num">2 </span>Neural Network Representation</a></div><div class="lev1 toc-item"><a href="#Computing-a-Neural-Network's-Output" data-toc-modified-id="Computing-a-Neural-Network's-Output-3"><span class="toc-item-num">3 </span>Computing a Neural Network's Output</a></div><div class="lev1 toc-item"><a href="#Vectorizing-across-multiple-examples" data-toc-modified-id="Vectorizing-across-multiple-examples-4"><span class="toc-item-num">4 </span>Vectorizing across multiple examples</a></div><div class="lev1 toc-item"><a href="#Explanation-for-Vectorized-Implementation" data-toc-modified-id="Explanation-for-Vectorized-Implementation-5"><span class="toc-item-num">5 </span>Explanation for Vectorized Implementation</a></div><div class="lev1 toc-item"><a href="#Activation-functions" data-toc-modified-id="Activation-functions-6"><span class="toc-item-num">6 </span>Activation functions</a></div><div class="lev1 toc-item"><a href="#Why-do-you-need-non-linear-activation-functions?" data-toc-modified-id="Why-do-you-need-non-linear-activation-functions?-7"><span class="toc-item-num">7 </span>Why do you need non-linear activation functions?</a></div><div class="lev1 toc-item"><a href="#Derivatives-of-activation-functions" data-toc-modified-id="Derivatives-of-activation-functions-8"><span class="toc-item-num">8 </span>Derivatives of activation functions</a></div><div class="lev2 toc-item"><a href="#Sigmoid-activation-Function" data-toc-modified-id="Sigmoid-activation-Function-81"><span class="toc-item-num">8.1 </span>Sigmoid activation Function</a></div><div class="lev2 toc-item"><a href="#Tanh-activation-Function" data-toc-modified-id="Tanh-activation-Function-82"><span class="toc-item-num">8.2 </span>Tanh activation Function</a></div><div class="lev2 toc-item"><a href="#Relu-and-Leaky-Relu-activation-Functions" data-toc-modified-id="Relu-and-Leaky-Relu-activation-Functions-83"><span class="toc-item-num">8.3 </span>Relu and Leaky Relu activation Functions</a></div><div class="lev1 toc-item"><a href="#Gradient-descent-for-Neural-Networks" data-toc-modified-id="Gradient-descent-for-Neural-Networks-9"><span class="toc-item-num">9 </span>Gradient descent for Neural Networks</a></div><div class="lev1 toc-item"><a href="#Backpropagation-intuition" data-toc-modified-id="Backpropagation-intuition-10"><span class="toc-item-num">10 </span>Backpropagation intuition</a></div><div class="lev1 toc-item"><a href="#Random-Initialization" data-toc-modified-id="Random-Initialization-11"><span class="toc-item-num">11 </span>Random Initialization</a></div>
# Neural Networks Overview

# Neural Network Representation

# Computing a Neural Network's Output



# Vectorizing across multiple examples


# Explanation for Vectorized Implementation


# Activation functions

**some rules of thumb for choosing activation functions:**
- if your output is 0 1 value if you're I'm using binary classification then the sigmoid activation function is very natural for the upper layer and then for all other units on varalu or the rectified linear unit is increasingly the default choice of activation function so if you're not sure what to use, I would just use the relu activation function that's what you see most people using these days although sometimes people also use the tannish activation function once this advantage of the value is that the derivative is equal to zero when V is negative in practice this works just fine but there is another version of the value called the least G value will give you.

# Why do you need non-linear activation functions?

# Derivatives of activation functions
- when you implement back-propagation for your neural network you need to really compute the slope or the derivative of the activation functions. so
- let's take a look at our choices of activation functions and
- how you can compute the slope of these functions
## Sigmoid activation Function

## Tanh activation Function

## Relu and Leaky Relu activation Functions

# Gradient descent for Neural Networks


# Backpropagation intuition




# Random Initialization


| github_jupyter |
# Histogram based Gradient Boosting Regression with RobustScaler
This Code template is for the regression analysis using a simple Histogram based Gradient Boosting Trees Regressor and the feature rescaling technique RobustScaler .
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import RobustScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.experimental import enable_hist_gradient_boosting
from sklearn.ensemble import HistGradientBoostingRegressor
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Data Rescaling
#### Robust Scaler:
It Scales features using statistics that are robust to outliers.
For more information [click here](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html)
```
x_fit = RobustScaler()
x_train = x_fit.fit_transform(x_train)
x_test = x_fit.transform(x_test)
```
### Model
Histogram-based Gradient Boosting Regression Tree.This estimator is much faster than GradientBoostingRegressor for big datasets (n_samples >= 10 000).This estimator has native support for missing values (NaNs).
#### Tuning Parameters
[Reference](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.HistGradientBoostingRegressor.html)
> **loss**: The loss function to use in the boosting process. Note that the “least squares” and “poisson” losses actually implement “half least squares loss” and “half poisson deviance” to simplify the computation of the gradient. Furthermore, “poisson” loss internally uses a log-link and requires y >= 0
> **learning_rate**: The learning rate, also known as shrinkage. This is used as a multiplicative factor for the leaves values. Use 1 for no shrinkage.
> **max_iter**: The maximum number of iterations of the boosting process, i.e. the maximum number of trees.
> **max_depth**: The maximum depth of each tree. The depth of a tree is the number of edges to go from the root to the deepest leaf. Depth isn’t constrained by default.
> **l2_regularization**: The L2 regularization parameter. Use 0 for no regularization (default).
> **early_stopping**: If ‘auto’, early stopping is enabled if the sample size is larger than 10000. If True, early stopping is enabled, otherwise early stopping is disabled.
> **n_iter_no_change**: Used to determine when to “early stop”. The fitting process is stopped when none of the last n_iter_no_change scores are better than the n_iter_no_change - 1 -th-to-last one, up to some tolerance. Only used if early stopping is performed.
> **tol**: The absolute tolerance to use when comparing scores during early stopping. The higher the tolerance, the more likely we are to early stop: higher tolerance means that it will be harder for subsequent iterations to be considered an improvement upon the reference score.
> **scoring**: Scoring parameter to use for early stopping.
```
# model initialization and fitting
model = HistGradientBoostingRegressor(random_state=123)
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
score: The score function returns the coefficient of determination R2 of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),model.predict(x_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Neel Pawar, Github: [Profile]( https://github.com/neel-ntp)
| github_jupyter |
```
import tensorflow as tf
import os
from utils import *
from tqdm import tqdm
def encoder_block(inp, n_hidden, filter_size):
inp = tf.expand_dims(inp, 2)
inp = tf.pad(inp, [[0, 0], [(filter_size[0]-1)//2, (filter_size[0]-1)//2], [0, 0], [0, 0]])
conv = tf.layers.conv2d(inp, n_hidden, filter_size, padding="VALID", activation=None)
conv = tf.squeeze(conv, 2)
return conv
def decoder_block(inp, n_hidden, filter_size):
inp = tf.expand_dims(inp, 2)
inp = tf.pad(inp, [[0, 0], [filter_size[0]-1, 0], [0, 0], [0, 0]])
conv = tf.layers.conv2d(inp, n_hidden, filter_size, padding="VALID", activation=None)
conv = tf.squeeze(conv, 2)
return conv
def glu(x):
return tf.multiply(x[:, :, :tf.shape(x)[2]//2], tf.sigmoid(x[:, :, tf.shape(x)[2]//2:]))
def layer(inp, conv_block, kernel_width, n_hidden, residual=None):
z = conv_block(inp, n_hidden, (kernel_width, 1))
return glu(z) + (residual if residual is not None else 0)
def layer_norm(inputs, epsilon=1e-8):
mean, variance = tf.nn.moments(inputs, [-1], keep_dims=True)
normalized = (inputs - mean) / (tf.sqrt(variance + epsilon))
params_shape = inputs.get_shape()[-1:]
gamma = tf.get_variable('gamma', params_shape, tf.float32, tf.ones_initializer())
beta = tf.get_variable('beta', params_shape, tf.float32, tf.zeros_initializer())
return gamma * normalized + beta
def cnn_block(x, dilation_rate, pad_sz, hidden_dim, kernel_size):
x = layer_norm(x)
pad = tf.zeros([tf.shape(x)[0], pad_sz, hidden_dim])
x = tf.layers.conv1d(inputs = tf.concat([pad, x, pad], 1),
filters = hidden_dim,
kernel_size = kernel_size,
dilation_rate = dilation_rate)
x = x[:, :-pad_sz, :]
x = tf.nn.relu(x)
return x
class Model:
def __init__(
self,
num_layers,
size_layers,
learning_rate = 1e-4,
n_attn_heads = 16,
kernel_size = 3
):
self.X = tf.placeholder(tf.int32, (None, None))
self.training = tf.placeholder(tf.bool, None)
lookup_table = tf.get_variable(
'lookup_table',
dtype = tf.float32,
shape = [len(vocab), size_layers],
initializer = tf.truncated_normal_initializer(
mean = 0.0, stddev = 0.01
),
)
lookup_table = tf.concat(
(tf.zeros(shape = [1, size_layers]), lookup_table[1:, :]), 0
)
self.X_seq_len = tf.count_nonzero(self.X, 1, dtype=tf.int32)
self.gts = tf.convert_to_tensor(guided_attention())
forward = tf.nn.embedding_lookup(lookup_table, self.X)
self.Y = tf.placeholder(tf.float32, (None, None, n_mels * resampled))
batch_size = tf.shape(self.Y)[0]
self.decoder_inputs = tf.concat(
(tf.ones_like(self.Y[:, :1, :]), self.Y[:, :-1, :]), 1
)
self.decoder_inputs = self.decoder_inputs[:, :, -n_mels:]
self.Z = tf.placeholder(
tf.float32, (None, None, fourier_window_size // 2 + 1)
)
seq_lens = tf.count_nonzero(
tf.reduce_sum(self.decoder_inputs, -1), 1, dtype = tf.int32
)
e = tf.identity(forward)
for i in range(num_layers):
z = layer(forward, encoder_block, kernel_size, size_layers * 2, forward)
forward = z
encoder_output, output_memory = z, z + e
decoder_inputs = tf.layers.dense(self.decoder_inputs, size_layers)
g = tf.identity(decoder_inputs)
for i in range(num_layers):
attn_res = h = layer(decoder_inputs, decoder_block, kernel_size, size_layers * 2,
residual=tf.zeros_like(decoder_inputs))
C = []
for j in range(n_attn_heads):
h_ = tf.layers.dense(h, size_layers//n_attn_heads)
g_ = tf.layers.dense(g, size_layers//n_attn_heads)
zu_ = tf.layers.dense(encoder_output, size_layers//n_attn_heads)
ze_ = tf.layers.dense(output_memory, size_layers//n_attn_heads)
d = tf.layers.dense(h_, size_layers//n_attn_heads) + g_
dz = tf.matmul(d, tf.transpose(zu_, [0, 2, 1]))
a = tf.nn.softmax(dz)
c_ = tf.matmul(a, ze_)
C.append(c_)
c = tf.concat(C, 2)
h = tf.layers.dense(attn_res + c, size_layers)
decoder_inputs = h
decoder_output = tf.sigmoid(h)
self.Y_hat = tf.layers.dense(decoder_output, n_mels * resampled)
self.loss1 = tf.reduce_mean(tf.abs(self.Y_hat - self.Y))
self.loss_bd1 = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=self.Y_hat,
labels=self.Y))
out_decoder = tf.reshape(
self.Y, [tf.shape(self.Y)[0], -1, n_mels]
)
out_decoder = tf.layers.dense(out_decoder, size_layers)
for i in range(num_layers):
dilation_rate = 2 ** i
pad_sz = (kernel_size - 1) * dilation_rate
with tf.variable_scope('block_%d'%i):
out_decoder += cnn_block(out_decoder, dilation_rate, pad_sz, size_layers, kernel_size)
self.Z_hat = tf.layers.dense(out_decoder, 1 + fourier_window_size // 2)
self.loss2 = tf.reduce_mean(tf.abs(self.Z_hat - self.Z))
self.loss_bd2 = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=self.Z_hat,
labels=self.Z))
self.loss = self.loss1 + self.loss2 + self.loss_bd1 + self.loss_bd2
self.optimizer = tf.train.AdamOptimizer(
learning_rate = learning_rate
).minimize(self.loss)
tf.reset_default_graph()
sess = tf.InteractiveSession()
size_layers = 256
learning_rate = 1e-4
num_layers = 4
model = Model(num_layers, size_layers, learning_rate)
sess.run(tf.global_variables_initializer())
paths, lengths, texts, raw_texts = [], [], [], []
text_files = [f for f in os.listdir('mel') if f.endswith('.npy')]
for fpath in text_files:
with open('%s/%s' % (path, fpath.replace('npy', 'txt'))) as fopen:
text = fopen.read()
paths.append(fpath.replace('.npy', ''))
text = text_normalize(text)
raw_texts.append(text)
text = text + 'E'
texts.append(np.array([char2idx[char] for char in text], np.int32))
lengths.append(len(text))
def dynamic_batching(paths):
files, max_y, max_z = [], 0, 0
for n in range(len(paths)):
files.append(get_cached(paths[n]))
if files[-1][0].shape[0] > max_y:
max_y = files[-1][0].shape[0]
if files[-1][1].shape[0] > max_z:
max_z = files[-1][1].shape[0]
return files, max_y, max_z
EPOCH = 50
for i in range(EPOCH):
pbar = tqdm(range(0, len(paths), batch_size), desc = 'minibatch loop')
for k in pbar:
index = min(k + batch_size, len(paths))
files, max_y, max_z = dynamic_batching(paths[k:index])
max_x = max(lengths[k:index])
batch_x = np.zeros((batch_size, max_x))
batch_y = np.zeros((batch_size, max_y, n_mels * resampled))
batch_z = np.zeros((batch_size, max_z, fourier_window_size // 2 + 1))
for n in range(len(files)):
batch_x[n, :] = np.pad(
texts[k + n],
((0, max_x - texts[k + n].shape[0])),
mode = 'constant',
)
batch_y[n, :, :] = np.pad(
files[n][0],
((0, max_y - files[n][0].shape[0]), (0, 0)),
mode = 'constant',
)
batch_z[n, :, :] = np.pad(
files[n][1],
((0, max_z - files[n][1].shape[0]), (0, 0)),
mode = 'constant',
)
_, cost, loss1, loss2, loss_bd1, loss_bd2 = sess.run(
[model.optimizer, model.loss,
model.loss1, model.loss2, model.loss_bd1,
model.loss_bd2],
feed_dict = {model.X: batch_x, model.Y: batch_y, model.Z: batch_z},
)
pbar.set_postfix(cost = cost, loss1 = loss1, loss2 = loss2, loss_bd1 = loss_bd1,
loss_bd2 = loss_bd2)
y_hat = np.ones((1, batch_y.shape[1], n_mels * resampled), np.float32)
for j in tqdm(range(batch_y.shape[1])):
_y_hat = sess.run(model.Y_hat, {model.X: [texts[0]], model.Y: y_hat})
y_hat[:, j, :] = _y_hat[:, j, :]
mags = sess.run(model.Z_hat, {model.Y: y_hat})
audio = spectrogram2wav(mags[0])
import IPython.display as ipd
ipd.Audio(audio, rate = sample_rate)
from scipy.io.wavfile import write
print('saving: %s'%(raw_texts[0]))
write(os.path.join('test.wav'), sample_rate, audio)
mags = sess.run(model.Z_hat, {model.Y: [batch_y[0]]})
audio = spectrogram2wav(mags[0])
ipd.Audio(audio, rate = sample_rate)
real_audio = spectrogram2wav(batch_z[0])
ipd.Audio(real_audio, rate = sample_rate)
_y_hat = sess.run(model.Y_hat, {model.X: [texts[0]], model.Y: [batch_y[0]]})
mags = sess.run(model.Z_hat, {model.Y: y_hat})
audio = spectrogram2wav(mags[0])
ipd.Audio(audio, rate = sample_rate)
```
| github_jupyter |
# ResNet in `gluon`
In this notebook we are going to:
1. Implement the popular ResNet18 convolutional neural network in just a few lines of code.
2. Use a pretrained model to make predictions on a new image.
ResNet is a powerful model that achieves high accuracy on a number computer vision tasks. The main innovation in the development of ResNets was the introduction of *residual connections*. In short, the residual connections change the mappings applied by neural network layers from $h_l = f(h_{1-1})$ (where $f$ is some linear mapping followed by an activation function to $h_l = f(h_{1-1}) + h_{l-1}$. Intuitively, while the easiest function for a normal neural network to learn is to map every vector to the zero vector (by setting all weights to $0$), the easiest mapping for a residual network to learn is the identity function. Another way to think about the benefit of residual networks is that they help with the vanishing gradient problem because the skip connections create shorter paths from the loss to the parameters.
## ResNet Architecture
<img src="https://github.com/zhreshold/ICCV19-GluonCV/blob/master/01_basics/resnet34-landscape.png?raw=true">
The ResNet we're going to work with in this tutorial consists of several parts.
- Conv layer with 7x7 kernel
- Max pooling
- Stage 1
- Stage 2
- Stage 3
- Stage 4
- Global average pooling
- Fully-connected output layer
Where each stage is a block following the architecture described below.
###### Each stage is built with repeated basic building blocks with residual connection.
<img src="https://github.com/zhreshold/ICCV19-GluonCV/blob/master/01_basics/resnet-basicblock.png?raw=true" width=512>
A basic block has the following components:
- Conv
- Conv
- Add result with input
- RELU Activation
Note that starting from stage 2, we need to double the number of channels and downsample the feature map in the first block. Therefore, the first convolutional layer in the main body needs strides, and the residual path also needs a convolutional layer with strides to downsample the feature map.
This pattern is easy to implement in Gluon.
## Gluon ResNet Implementation
First, let's implement the basic building block in ResNet.
```
%pylab inline
import matplotlib.pyplot as plt
from mxnet import nd, image
from mxnet.gluon import nn
from mxnet.gluon.block import HybridBlock
from mxnet.gluon.data.vision import transforms
from mxnet.gluon.model_zoo.vision import get_model
class BasicBlock(HybridBlock):
def __init__(self, channels, strides, downsample=False, **kwargs):
super(BasicBlock, self).__init__(**kwargs)
self.body = nn.HybridSequential()
self.body.add(nn.Conv2D(channels, kernel_size=3, strides=strides, padding=1, use_bias=False),
nn.BatchNorm(),
nn.Activation('relu'),
nn.Conv2D(channels, kernel_size=3, strides=1, padding=1, use_bias=False),
nn.BatchNorm())
self.relu = nn.Activation('relu')
self.downsample = None
if downsample:
self.downsample = nn.HybridSequential()
self.downsample.add(nn.Conv2D(channels, kernel_size=1, strides=strides, padding=1, use_bias=False),
nn.BatchNorm())
def hybrid_forward(self, F, x):
residual = x
x = self.body(x)
if self.downsample:
residual = self.downsample(residual)
return self.relu(residual+x)
```
Next - the function to construct one stage.
```
def make_stage(num_blocks, channels, strides, downsample):
stage = nn.HybridSequential()
stage.add(BasicBlock(channels=channels, strides=strides, downsample=downsample))
for i in range(num_blocks-1):
stage.add(BasicBlock(channels=channels, strides=1, downsample=False))
return stage
```
Finally, we can define a ResNet18 structure in gluon easily:
```
def resnet18(classes):
net = nn.HybridSequential()
net.add(nn.Conv2D(channels=64, kernel_size=7, strides=2,
padding=3, use_bias=False),
nn.BatchNorm(),
nn.Activation('relu'),
nn.MaxPool2D(pool_size=3, strides=2, padding=1),
# Stages
make_stage(2, 64, 1, False),
make_stage(2, 128, 2, True),
make_stage(2, 256, 2, True),
make_stage(2, 512, 2, True),
# Output
nn.GlobalAvgPool2D(),
nn.Dense(classes))
return net
num_classes = 10
net = resnet18(num_classes)
# print(net)
```
## Make predictions with a pre-trained ResNet
ImageNet is a large database with over one million natural images. Models trained on this dataset are capable of making accurate predictions on a wide variety of natural images. `Gluon` offers a model zoo with many standard architectures and easily accessible pretrinaed paramameters.
We can demonstrate the power of pretrained models by making prediction on an image.
```
net = get_model('resnet18_v1', pretrained=True, classes=1000)
img = image.imread('mt_baker.jpg')
plt.figure(figsize=(20, 40))
plt.imshow(img.asnumpy())
plt.show()
```
We need to first process the image with:
- Resize the shorter edge to 256 px, keeping the aspect ratio
- Crop the center 224x224 square
- Transpose the image to tensor format
- Normalize the image with ImageNet parameters
```
transform_fn = transforms.Compose([
transforms.Resize(256, keep_ratio=True),
transforms.CenterCrop(256),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
img = transform_fn(img).expand_dims(0)
img.shape
```
Now we can simply predict with:
```
pred = net(img)
```
Let's see how the result looks like:
```
with open('imagenet_labels.txt', 'r') as f:
class_names = [l.strip('\r\n') for l in f.readlines()]
topK = 5
ind = nd.topk(pred, k=topK)[0].astype('int')
print('The input picture is classified to be')
for i in range(topK):
name = class_names[ind[i].asscalar()]
prob = nd.softmax(pred[0])[ind[i]].asscalar()
print('\t[%s], with probability %.3f.'% (name, prob))
```
## Practice
Find a PNG image online. Can the pre-trained model classify it correctly?
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.