code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Блокнот к вопросу https://ru.stackoverflow.com/questions/1294279/
#
# +
import scipy.io as io
import scipy as sp
import scipy.fft as fft
import scipy.signal as signal
import matplotlib.pyplot as plt
import numpy as np
# -
ecg_list = []
# Электрокардиограмма из набора https://data.mendeley.com/datasets/7dybx7wyfn/3
#
# Кардиограмма состоит из 10 последовательных неперекрывающихся наборов, снятых с частотой 360 Гц. В каждом файле 3600 замеров, всего 36000 замеров.
for i in range(10):
mat = io.loadmat('./MLII/1 NSR/100m (0).mat')
ecg_list.append(mat["val"].flatten())
ecg = np.concatenate(ecg_list)
ecg.shape
plt.plot(ecg[2000:4000])
# Построение спектра средствами периодограммы
#
# Периодограмма в простейшем виде - модуль амплитуды преобразования Фурье
freqs, psd = signal.periodogram(ecg, fs=360)
plt.plot(freqs, psd)
# Диапазон частот менее 4 Гц (т.е. процессы с частотой не более 240 сокращений в минуту)
low_freqs = freqs[freqs < 4.0]
plt.plot(low_freqs, psd[:len(low_freqs)])
# **Построение спектра средствами быстрого преобразования Фурье**
fft_data = np.abs(np.fft.fft(ecg))
# Удаление нулевой частоты - постоянной части сигнала.
fft_data[0] = np.nan
# Функция `numpy.fft.fft` возвращает значения для дискретного набора частот. Для заданного числа замеров частоты возвращает функция `fftfreq`. Второй параметр задаёт время дискретизации, интервал в секундах между смежными замерами.
time_step = 1 / 360
freqs = np.fft.fftfreq(len(ecg), time_step)
# Удаление частот менее 4 Гц.
# Так как сигнал получился симметричным, отрицательные частоты отброшены.
idx = np.argsort(freqs)
idx = idx[np.abs(freqs[idx]) < 4.0]
idx = idx[freqs[idx] > 0]
plt.plot(freqs[idx], spectre[idx])
plt.xlabel("Частота, Гц")
plt.ylabel("Амплитуда")
|
python/1294279/.ipynb_checkpoints/ecg-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
python argparse_type_int.py 100
python argparse_type_int.py foo
python argparse_type_int.py 1.23
python argparse_type_bool.py True
python argparse_type_bool.py False
python argparse_type_bool.py bar
python argparse_option_bool.py --en
python argparse_option_bool.py
python argparse_type_strtobool.py true
python argparse_type_strtobool.py false
|
notebook/argparse_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KinsleyDavis/Novo/blob/main/Colab_ArteMaisComp.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="tCkT6_ZjYe8o"
def escolher_arquivo():
import ipywidgets as widgets
from IPython.display import clear_output
import os
import matplotlib.pyplot as plt
# !pip install chainer &> /dev/null
# !pip install cupy-cuda101==7.7.0
# !git clone https://github.com/artemaiscomp/chainer-fast-neuralstyle &> /dev/null
from google.colab import files
content_img = files.upload()
CONTENT_IMAGE_FN = list(content_img)[0]
CONTENT_IMAGE_FN_temp = CONTENT_IMAGE_FN.strip().replace(" ", "_")
if CONTENT_IMAGE_FN != CONTENT_IMAGE_FN_temp:
os.rename(CONTENT_IMAGE_FN, CONTENT_IMAGE_FN_temp)
CONTENT_IMAGE_FN = CONTENT_IMAGE_FN_temp
#print("Nome do arquivo da imagem :", CONTENT_IMAGE_FN)
# %matplotlib inline
fig = plt.figure(figsize=(10, 10))
img = plt.imread(CONTENT_IMAGE_FN)
plt.axis('off')
#plt.title('Content image')
plt.imshow(img)
# + id="qAOd5t2qP1eZ" cellView="form"
#@title Clique no Play e após em "Escolher Arquivo" para escolher sua imagem a ser estilizada
import ipywidgets as widgets
from IPython.display import clear_output
import os
import matplotlib.pyplot as plt
# !pip install chainer &> /dev/null
# !pip install cupy-cuda101 &> /dev/null
# !git clone https://github.com/artemaiscomp/chainer-fast-neuralstyle &> /dev/null
from google.colab import files
content_img = files.upload()
CONTENT_IMAGE_FN = list(content_img)[0]
CONTENT_IMAGE_FN_temp = CONTENT_IMAGE_FN.strip().replace(" ", "_")
if CONTENT_IMAGE_FN != CONTENT_IMAGE_FN_temp:
os.rename(CONTENT_IMAGE_FN, CONTENT_IMAGE_FN_temp)
CONTENT_IMAGE_FN = CONTENT_IMAGE_FN_temp
#print("Nome do arquivo da imagem :", CONTENT_IMAGE_FN)
# %matplotlib inline
fig = plt.figure(figsize=(10, 10))
img = plt.imread(CONTENT_IMAGE_FN)
plt.axis('off')
#plt.title('Content image')
plt.imshow(img)
# + id="TXLFpBasYbmL"
# + id="p9p-LAAqVsFw" cellView="form"
#@title Selecione a arte a ser aplicada.
import os, ipywidgets as widgets
from IPython.display import clear_output
model_files = [f for f in os.listdir('/content/chainer-fast-neuralstyle/models') if f.endswith('.model')]
model=widgets.Dropdown(
options=model_files,
value='hokusai.model',
description='Modelo:',
disabled=False,
)
model
# + id="hFRk-GJhDoS6" cellView="form"
#@title Clique no botão Play e abaixo no botao OK para converter a imagem com o estilo escolhido.
clear_output()
#@title Clique no botão Play e abaixo no botao OK para converter a imagem com o estilo escolhido.
from IPython.display import clear_output
button = widgets.Button(description='OK')
# !pip install chainer &> /dev/null
# !pip install cupy-cuda101==7.7.0 &> /dev/null
clear_output()
out = widgets.Output()
def on_button_clicked(_):
# "linkar funcão com saída"
with out:
# !python chainer-fast-neuralstyle/generate.py $CONTENT_IMAGE_FN unique -m chainer-fast-neuralstyle/models/$model.value -o output.jpg --gpu 0 &> /dev/null
fig = plt.figure(figsize=(10, 10))
img = plt.imread('output.jpg')
plt.axis('off')
plt.title('imagem estilizada')
plt.imshow(img)
# unir butão e funcão juntos usando um métodos no butão
button.on_click(on_button_clicked)
# mostrar butão e sua saída juntos
widgets.VBox([button,out])
|
Colab_ArteMaisComp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# ## FLAG Example
# +
import argparse
from ogb.nodeproppred import DglNodePropPredDataset, Evaluator
import torch
from torch import nn
import torch.nn.functional as F
from utils import Logger, EarlyStopping
from dgl.nn import GraphConv, SAGEConv
# -
# import flag
from gtrick import FLAG
# ### Define Models
class GCN(nn.Module):
def __init__(self, in_channels, hidden_channels, out_channels, num_layers,
dropout):
super(GCN, self).__init__()
self.convs = nn.ModuleList()
self.convs.append(GraphConv(in_channels, hidden_channels))
self.bns = nn.ModuleList()
self.bns.append(nn.BatchNorm1d(hidden_channels))
for _ in range(num_layers - 2):
self.convs.append(
GraphConv(hidden_channels, hidden_channels))
self.bns.append(nn.BatchNorm1d(hidden_channels))
self.convs.append(GraphConv(hidden_channels, out_channels))
self.dropout = dropout
def reset_parameters(self):
for conv in self.convs:
conv.reset_parameters()
for bn in self.bns:
bn.reset_parameters()
# add a param `perturb` to pass perturb
def forward(self, g, x, perturb=None):
# add perturb to x, note that do not use x += perturb
if perturb is not None:
x = x + perturb
for i, conv in enumerate(self.convs[:-1]):
x = conv(g, x)
x = self.bns[i](x)
x = F.relu(x)
x = F.dropout(x, p=self.dropout, training=self.training)
x = self.convs[-1](g, x)
return x
class SAGE(nn.Module):
def __init__(self, in_channels, hidden_channels, out_channels, num_layers,
dropout):
super(SAGE, self).__init__()
self.convs = nn.ModuleList()
self.convs.append(SAGEConv(in_channels, hidden_channels, 'mean'))
self.bns = nn.ModuleList()
self.bns.append(nn.BatchNorm1d(hidden_channels))
for _ in range(num_layers - 2):
self.convs.append(SAGEConv(hidden_channels, hidden_channels, 'mean'))
self.bns.append(nn.BatchNorm1d(hidden_channels))
self.convs.append(SAGEConv(hidden_channels, out_channels, 'mean'))
self.dropout = dropout
def reset_parameters(self):
for conv in self.convs:
conv.reset_parameters()
for bn in self.bns:
bn.reset_parameters()
# add a param perturb `perturb` to pass perturb
def forward(self, g, x, perturb=None):
# add perturb to x, note that do not use x += perturb
if perturb is not None:
x = x + perturb
for i, conv in enumerate(self.convs[:-1]):
x = conv(g, x)
x = self.bns[i](x)
x = F.relu(x)
x = F.dropout(x, p=self.dropout, training=self.training)
x = self.convs[-1](g, x)
return x
# ### Define Train Process
# pass flag func to train
def train(model, g, x, y, train_idx, flag):
y = y[train_idx].squeeze(1)
forward = lambda perturb: model(g, x, perturb)[train_idx]
loss, out = flag(model, forward, g.num_nodes(), y)
return loss.item()
@torch.no_grad()
def test(model, g, x, y, split_idx, evaluator, eval_metric):
model.eval()
out = model(g, x)
y_pred = out.argmax(dim=-1, keepdim=True)
train_metric = evaluator.eval({
'y_true': y[split_idx['train']],
'y_pred': y_pred[split_idx['train']],
})[eval_metric]
valid_metric = evaluator.eval({
'y_true': y[split_idx['valid']],
'y_pred': y_pred[split_idx['valid']],
})[eval_metric]
test_metric = evaluator.eval({
'y_true': y[split_idx['test']],
'y_pred': y_pred[split_idx['test']],
})[eval_metric]
return train_metric, valid_metric, test_metric
def run_node_pred(args, model, dataset):
device = f'cuda:{args.device}' if torch.cuda.is_available() else 'cpu'
device = torch.device(device)
model.to(device)
evaluator = Evaluator(name=args.dataset)
g, y = dataset[0]
# add reverse edges
srcs, dsts = g.all_edges()
g.add_edges(dsts, srcs)
# add self-loop
print(f"Total edges before adding self-loop {g.number_of_edges()}")
g = g.remove_self_loop().add_self_loop()
print(f"Total edges after adding self-loop {g.number_of_edges()}")
g, y = g.to(device), y.to(device)
if args.dataset == 'ogbn-proteins':
x = g.ndata['species']
else:
x = g.ndata['feat']
split_idx = dataset.get_idx_split()
train_idx = split_idx['train']
logger = Logger(args.runs, mode='max')
for run in range(args.runs):
model.reset_parameters()
optimizer = torch.optim.Adam(model.parameters(), lr=args.lr)
early_stopping = EarlyStopping(
patience=args.patience, verbose=True, mode='max')
if dataset.task_type == 'binary classification':
loss_func = nn.BCEWithLogitsLoss()
elif dataset.task_type == 'multiclass classification':
loss_func = nn.CrossEntropyLoss()
# define flag, params: in_feats, loss_func, optimizer
flag = FLAG(g.ndata['feat'].shape[1], loss_func, optimizer)
for epoch in range(1, 1 + args.epochs):
loss = train(model, g, x, y, train_idx,
flag)
result = test(model, g, x, y, split_idx,
evaluator, dataset.eval_metric)
logger.add_result(run, result)
train_acc, valid_acc, test_acc = result
if epoch % args.log_steps == 0:
print(f'Run: {run + 1:02d}, '
f'Epoch: {epoch:02d}, '
f'Loss: {loss:.4f}, '
f'Train: {100 * train_acc:.2f}%, '
f'Valid: {100 * valid_acc:.2f}% '
f'Test: {100 * test_acc:.2f}%')
if early_stopping(valid_acc, model):
break
logger.print_statistics(run)
logger.print_statistics()
# ### Run Node Property Prediction Experiment
parser = argparse.ArgumentParser(
description='train node property prediction')
parser.add_argument("--dataset", type=str, default="ogbn-arxiv",
choices=["ogbn-arxiv"])
parser.add_argument("--dataset_path", type=str, default="/home/ubuntu/.dgl_dataset",
help="path to dataset")
parser.add_argument('--device', type=int, default=0)
parser.add_argument('--log_steps', type=int, default=1)
parser.add_argument('--model', type=str, default='gcn')
parser.add_argument('--num_layers', type=int, default=3)
parser.add_argument('--hidden_channels', type=int, default=256)
parser.add_argument('--dropout', type=float, default=0.5)
parser.add_argument('--lr', type=float, default=0.01)
parser.add_argument('--epochs', type=int, default=500)
parser.add_argument('--runs', type=int, default=5)
parser.add_argument('--patience', type=int, default=30)
args = parser.parse_args(args=[])
print(args)
# +
dataset = DglNodePropPredDataset(
name=args.dataset,
root=args.dataset_path
)
g, _ = dataset[0]
num_features = g.ndata['feat'].shape[1]
if args.model == 'gcn':
model = GCN(num_features, args.hidden_channels,
dataset.num_classes, args.num_layers,
args.dropout)
elif args.model == 'sage':
model = SAGE(num_features, args.hidden_channels,
dataset.num_classes, args.num_layers,
args.dropout)
# -
run_node_pred(args, model, dataset)
|
benchmark/dgl/FLAG.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **2 September 2020**
#
# # Introduction: INFO 3350/6350
#
# **Welcome to Information Science 3350 and 6350: Text Mining for History and Literature.**
#
# Here's today's agenda:
#
# ## Review the syllabus
#
# * What is this class about?
# * Who is it for?
# * Who might *not* want to take this class?
# * What will we do and how will it run?
# * Note that you will often have a problem set due on Tuesday evening.
# * Four times during the semester, on dates of your choice, you will *also* have a reading response due on Tuesday.
#
# ## Introduce staff
#
# * Prof. <NAME>
# * <NAME> (grad TA)
# * <NAME> (undergrad TA)
# * <NAME> (undergrad TA)
# * <NAME> (undergrad TA)
#
# ## Whom to contact?
#
# * Homework problem, due date question, GitHub issue, lecture clarification, etc: post to Campuswire.
# * Course admin issue (switch sections, access CMS, etc.): email Maria.
# * Everything else: email Matt or come to Matt or Maria's office hours.
#
# When in doubt, post to Campuswire. Questions there will be flagged for Matt and Maria as needed. Note that you can get extra credit for answering (correctly!) other students' questions on Campuswire.
#
# **Links to all course resources are on [Canvas](https://canvas.cornell.edu/courses/20174).**
#
# ## Friday sections
#
# * Sections 201 and 202 are in-person in Upson Hall. If you are enrolled in one of these sections, **do not come to class unless you have a seat assignment**. Seat assignments will be sent to you via Canvas no later than Thursday night. If you do not have a seat assignment by Friday morning, stay home and join one of the later online sections.
# * Sections 203, 204, and 205 are online at the same Zoom link as the lectures.
#
# ## Graduate section
#
# Grad students enrolled in 6350 will have an additional meeting each week to discuss research and advanced topics. If you are enrolled in 6350, please fill out the poll launching ... now.
|
lectures/lec-09-02-intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import h5py
import condor
import copy
import numpy as np
import condor.utils.linalg as linalg
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.cm as cm
fullp_res_edge = lambda det,w: 1./linalg.length(det.get_q_max(w, pos='edge')/(2*np.pi))
fullp_res_corner = lambda det,w: 1./linalg.length(det.get_q_max(w, pos='corner')/(2*np.pi))
def plot_diffraction(data, mask=None):
if mask is None:
mask = np.ones_like(data).astype(np.bool)
image = np.ma.array(np.copy(data).astype(np.float), mask=~mask)
image.data[image.data<0.5] = 1e-5
palette = copy.copy(cm.magma)
palette.set_bad('w', 1.)
palette.set_under('0.9',1.)
plt.figure()
plt.imshow(image, norm=colors.LogNorm(vmin=1), interpolation='none', cmap=palette)
plt.axis('off')
cb = plt.colorbar(pad=0)
cb.ax.set_ylabel('Intensity [photons / px]')
cb.outline.set_visible(False)
plt.tight_layout()
plt.show()
# # Parameters
# Detector (pnCCD)
pixelsize = 8*75e-6
nx,ny = (1024//8,1024//8)
detector_distance = 300e-3
# Source
N = 10
photon_energy = np.random.normal(loc=3500, scale=10, size=(N,)) # [eV]
ph = condor.utils.photon.Photon(energy_eV=photon_energy)
wavelength = ph.get_wavelength()
fluence = 1e15 #[ph/um2]
focus_diameter = 0.2e-6
pulse_energy = fluence * ph.get_energy() * (np.pi*((1e6*focus_diameter/2.)**2)) # [J]
# Sample
pdb_id = '1FFK'
sample_size = 18e-9
# # Fix particle orientation
angle_degrees = 72.5
angle = angle_degrees/360.*2*np.pi
rotation_axis = np.array([1.,1.,0.])/np.sqrt(2.)
quaternion = condor.utils.rotation.quat(angle,rotation_axis[0],rotation_axis[1], rotation_axis[2])
rotation_values = np.array([quaternion])
rotation_formalism = "quaternion"
rotation_mode = "extrinsic"
# # Run simulation
incoherent_average = 0
det = condor.Detector(distance=detector_distance, pixel_size=pixelsize, nx=nx, ny=ny, noise="poisson")
par = condor.ParticleAtoms(pdb_id=pdb_id,
rotation_formalism=rotation_formalism,
rotation_values=rotation_values,
rotation_mode=rotation_mode)
for i in range(N):
src = condor.Source(wavelength=wavelength[i], pulse_energy=pulse_energy[i], focus_diameter=focus_diameter)
E = condor.Experiment(source=src, particles={"particle_atoms":par}, detector=det)
o = E.propagate()
incoherent_average += o["entry_1"]["data_1"]["data"][:]
incoherent_average /= N
o["entry_1"]["data_1"]["data"] = incoherent_average
# # Output
print "Photon energy: %d eV" %photon_energy.mean()
print "Fluence: %g ph/um2" %fluence
print "Pulse energy: %.4f mJ" %(1e3*pulse_energy.mean())
print "Sample size: %d nm" %(1e9*sample_size)
print "Detector distance: %d mm" %(1e3*detector_distance)
print "Full period resolution (corner): %.2f nm" %(1e9*fullp_res_corner(det,wavelength.mean()))
print "Full period resolution (edge): %.2f nm" %(1e9*fullp_res_edge(det,wavelength.mean()))
print "Nr. of resolution elements: %.2f" %(sample_size / fullp_res_corner(det,wavelength.mean())*2)
plot_diffraction(o['entry_1']['data_1']['data'])
print "\n"
# # Write to CXI file
W = condor.utils.cxiwriter.CXIWriter("../data/single_protein_polychromatic.h5")
W.write(o)
W.close()
|
ipynb/simulate_protein_polychromatic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''thoughts-dev'': conda)'
# language: python
# name: python3
# ---
# ## Identifying Poker Hands
#
# As it turns out, identifying poker hands is a great way to learn how sequence and set rules work. Begin by importing the standard libraries and creating a new rules engine.
#
# ## ♥ ♦ ♠ ♣
# +
import os, sys
sys.path.insert(1, os.path.abspath('..\\..'))
from thoughts.rules_engine import RulesEngine
import pprint
engine = RulesEngine()
# -
# ## High Card
#
# Detecting high cards is relatively straight forward. Since any card could be a high card* we'll simply forward the card's rank information on as a high card.
#
# Don't worry for now about picking the highest scoring one of these results. These initial rules are simply to generate candidate card rankings for all possibilities. Later we'll pick the highest scoring pattern out of the candidates.
#
# *Except for 2's, since there would always be guaranteed at least at 3 in 10 cards shared among 2 people using 2 decks, but will ignore that anomaly.
# +
# define the rule
high_card_rule = {"#when": [{"suit": "?suit", "rank": "?rank1"}],
"#then": {"high-card": "?rank1", "score": 1}}
engine.add_rule(high_card_rule)
# assert a hand to test the rule
hand = [
{"rank": "3", "suit": "spades"},
{"rank": "5", "suit": "clubs"},
{"rank": "7", "suit": "hearts"},
{"rank": "9", "suit": "diamonds"},
{"rank": "K", "suit": "clubs"}]
result = engine.process(hand)
pprint.pprint(result)
# -
# ## One Pair
#
# To detect pairs, we are interested whenever there are two cards that follow in sequence that have the same rank.
#
# In the rule below, note that the ?rank1 variable is the same between both constituents in the #when sequence. The engine will use this informtion to make sure the value of the rank in the second card matches the rank in the first card.
#
# Also note the #seq-type of "allow-junk" as part of the rule. This allows for extra constituents to be in-between the constituents we are looking for. Without this, the second card in the pair would have to come immediately after first card in the sequence. For example, it would detect 2♥ followed by 2♦, but not 2♥ followed by 3♠ followed by 2♦. The 3♠ is the "junk" consituent which is ignored whenever #seq-type is "allow-junk".
# +
# define the rule
one_pair = {"#when": [{"rank": "?rank1"}, {"rank": "?rank1"}],
"#seq-type": "allow-junk",
"#then": {"one-pair": "?rank1", "score": 2}}
engine.add_rule(one_pair)
# assert a hand to test the rule
hand = [
{"rank": "J", "suit": "spades"},
{"rank": "3", "suit": "clubs"},
{"rank": "J", "suit": "hearts"},
{"rank": "7", "suit": "diamonds"},
{"rank": "5", "suit": "clubs"}]
result = engine.process(hand)
pprint.pprint(result, sort_dicts=False)
# -
# ## Two Pair
#
# Detecting two-pair works by matching whenever you find a single (one) pair, followed by another single (one) pair, with junk cards allowed in between.
#
# Here the ranks can be the same, but do not have to be the same. This means this rule will detect four 4's as two pairs, though we know it also be better known as four of a kind. That's OK - we'll let the engine detect the two pairs and ALSO detect the four of a kind. Technically, that's an accurate identification the allowed patterns. In general it's better for the engine to over-generate matches as possible "ideas", and then let another set of rules filter these down into the correct conclusion based on some other criteria.
# +
# add the rule
rule = {"#when": [{"one-pair": "?rank1"}, {"one-pair": "?rank2"}],
"#seq-type": "allow-junk",
"#then": {"two-pair": {"pair1": "?rank1", "pair2": "?rank2"}, "score": 3}}
engine.add_rule(rule)
# test it
hand = [
{"rank": "J", "suit": "spades"},
{"rank": "J", "suit": "spades"},
{"rank": "7", "suit": "hearts"},
{"rank": "3", "suit": "clubs"},
{"rank": "7", "suit": "hearts"}
]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Three of a Kind
#
# Three of a Kind works similiar to the one-pair and two-pair rules. Look for a sequence of three cards with the same rank, and allow junk cards in between.
# +
# add the rule
rule = {"#when": [{"rank": "?rank1"}, {"rank": "?rank1"}, {"rank": "?rank1"}],
"#seq-type": "allow-junk",
"#name": "Three of a Kind", "#then": {"three-of-a-kind": "?rank1", "score": 4}}
engine.add_rule(rule)
# test it
hand = [
{"rank": "J", "suit": "spades"},
{"rank": "J", "suit": "hearts"},
{"rank": "3", "suit": "clubs"},
{"rank": "J", "suit": "diamonds"},
{"rank": "7", "suit": "clubs"}]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Straight
#
# With a straight, things get more interesting. We need to detect whenever we have 5 cards in a sequence, where each card is one rank higher than the card before it.
#
# We can do this by first detecting the "mini-runs", where there is one card which is one higher than the card before it. Then we'll look for a number of mini-runs with are connected by the ending and beginning card.
#
# To allow for this type of overlap in sequence, where the ending of one constituent can be the beginning of the next constituent, we set the #seq-type to "overlap-connected".
# +
rules = [
{"#when": [{"rank": "2"}, {"rank": "3"}],
"#then": {"mini-run": "3"}},
{"#when": [{"rank": "3"}, {"rank": "4"}],
"#then": {"mini-run": "4"}},
{"#when": [{"rank": "4"}, {"rank": "5"}],
"#then": {"mini-run": "5"}},
{"#when": [{"rank": "5"}, {"rank": "6"}],
"#then": {"mini-run": "6"}},
{"#when": [{"rank": "6"}, {"rank": "7"}],
"#then": {"mini-run": "7"}},
{"#when": [{"rank": "7"}, {"rank": "8"}],
"#then": {"mini-run": "8"}},
{"#when": [{"rank": "8"}, {"rank": "9"}],
"#then": {"mini-run": "9"}},
{"#when": [{"rank": "9"}, {"rank": "10"}],
"#then": {"mini-run": "10"}},
{"#when": [{"rank": "10"}, {"rank": "J"}],
"#then": {"mini-run": "J"}},
{"#when": [{"rank": "J"}, {"rank": "Q"}],
"#then": {"mini-run": "Q"}},
{"#when": [{"rank": "Q"}, {"rank": "K"}],
"#then": {"mini-run": "K"}},
{"#when": [{"rank": "K"}, {"rank": "A"}],
"#then": {"mini-run": "A"}},
{"#when": [{"mini-run": "?run1"}, {"mini-run": "?run2"}, {"mini-run": "?run3"}, {"mini-run": "?run4"}],
"#seq-type": "overlap-connected",
"#then": {"straight": "?run4", "score": 5}}
]
engine.add_rules(rules)
hand = [
{"rank": "3", "suit": "spades"},
{"rank": "4", "suit": "hearts"},
{"rank": "5", "suit": "diamonds"},
{"rank": "6", "suit": "clubs"},
{"rank": "7", "suit": "spades"}]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Flush
#
# A flush is quite a bit easier. Similiar to the pair rule, we need to detect 5 cards in sequence which are share the same suit.
#
# We do this by using the same ?suit1 variable in all five consituents in the pattern.
# +
rule = {"#when": [{"suit": "?suit1"}, {"suit": "?suit1"}, {"suit": "?suit1"}, {"suit": "?suit1"}, {"suit": "?suit1"}],
"#then": {"flush": "?suit1", "score": 6}}
engine.add_rule(rule)
hand = [
{"rank": "3", "suit": "spades"},
{"rank": "5", "suit": "spades"},
{"rank": "7", "suit": "spades"},
{"rank": "9", "suit": "spades"},
{"rank": "J", "suit": "spades"}]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Full House
#
# A full house sounds trivial at first, but there's a catch. The cards can be arranged either with the three of a kind going first and the pair going second; the pair going first and the three of a kind going second; or the cards interlaced so that the pair is hiding in the middle of the three of a kind!
#
# For this scenario, rather than look for a sequence of cards in order, we'll look for a "set", which means the cards can be arranged in any order, as long as the constituents do not contain any sub-constituents of each other. Fortunately the engine has an easy way to handle this, by setting the #seq-type to "set".
# +
rule = {"#when": [{"three-of-a-kind": "?rank1"}, {"one-pair": "?rank2"}],
"#seq-type": "set",
"#then": {"full-house": {"three-of-a-kind": "?rank1", "one-pair": "?rank2"}, "score": 7}}
engine.add_rule(rule)
hand = [
{"rank": "3", "suit": "spades"},
{"rank": "7", "suit": "spades"},
{"rank": "3", "suit": "hearts"},
{"rank": "7", "suit": "hearts"},
{"rank": "3", "suit": "diamonds"}]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Four of a Kind
#
# A four of a kind returns to normalcy - just detect four cards that have the same rank, and allow for junk cards in between constituents.
#
# Notice that the engine detects quite a bit of other patterns here too! That's OK, it is detecting legal sequences based on the previous rules. Just like in real poker, you can look at the cards in your hand in different ways, with the goal of picking the *highest* ranking arrangement in your hand.
# +
rule = {"#when": [{"rank": "?rank1"}, {"rank": "?rank1"}, {"rank": "?rank1"}, {"rank": "?rank1"}],
"#seq-type": "allow-junk",
"#then": {"four-of-a-kind": "?rank1", "score": 8}}
engine.add_rule(rule)
hand = [
{"rank": "3", "suit": "spades"},
{"rank": "3", "suit": "clubs"},
{"rank": "3", "suit": "hearts"},
{"rank": "7", "suit": "hearts"},
{"rank": "3", "suit": "diamonds"}]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Straight Flush
#
# Detecting straight flushes presents a challenge. We want to detect whenever the arrangement of cards matches both a straight *and* and flush, and the members in both sets can overlap.
#
# At first it's tempting to use the #seq-type = "set" option above, which gets us part of the way there. However, by default this option does not allow the constituents to contain members that are already in other constituents in the pattern, and so would fail by itself.
#
# We can relax this option, so that the matching *will* allow constituents to share members, by using the #seq-allow-multi option. In this way the straight can contain members from the flush, and vice-versa.
# +
rule = {"#when": [{"straight": "?rank"}, {"flush": "?suit"}],
"#seq-type": "set",
"#seq-allow-multi": True,
"#then": {"straight-flush": "?suit", "rank": "?rank", "score": 9}}
engine.add_rule(rule)
hand = [
{"rank": "3", "suit": "spades"},
{"rank": "4", "suit": "spades"},
{"rank": "5", "suit": "spades"},
{"rank": "6", "suit": "spades"},
{"rank": "7", "suit": "spades"}]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Royal Flush
#
# Ah, the infamous royal flush. This turns out to be a special case of the straight flush rule, where the rank of the straight is an ace.
#
# Easy enough:
# +
rule = {"#when": [{"straight-flush": "?suit", "rank": "A"}],
"#then": {"royal-flush": "?suit", "score": 10}}
engine.add_rule(rule)
hand = [
{"rank": "10", "suit": "hearts"},
{"rank": "J", "suit": "hearts"},
{"rank": "Q", "suit": "hearts"},
{"rank": "K", "suit": "hearts"},
{"rank": "A", "suit": "hearts"}]
result = engine.process(hand, extract_conclusions=True)
pprint.pprint(result, sort_dicts=False)
# -
# ## Choosing the Highest Scoring Arrangement
#
# Choosing the highest scoring arrangment is a matter of picking out the conclusion with the highest score.
#
# Let's try it on a sample hand we used previously.
# +
hand = [
{"rank": "3", "suit": "spades"},
{"rank": "4", "suit": "spades"},
{"rank": "5", "suit": "spades"},
{"rank": "6", "suit": "spades"},
{"rank": "7", "suit": "spades"}]
result = engine.process(hand, extract_conclusions=True)
# pprint.pprint(result, sort_dicts=False)
max_score = max(filter(lambda x: "score" in x, result), key=lambda x: x['score'])
print("BEST RANKING:", max_score)
# -
# ## Conclusion
#
# Detecting poker hands illustrates many of the concepts in sequence and set detection. Hopefully you found it as a fun way to learn how the engine can help with this kind of scenario.
|
notebooks/fun/poker.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false id="NYJWmpEfR9lJ" nbgrader={"cell_type": "markdown", "checksum": "65581aa445e47361f83d64ec69310261", "grade": false, "grade_id": "cell-92665c32235efd5a", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # Linear Support Vector Machine (SVM)
#
# We've now seen how to optimise analytic functions using PyTorch's optimisers, and in the previous labs and exercises we played with training simple machine learning models with hand-coded gradient descent. Let's put everything together and implement a Soft-Margin Linear Support Vector Machine, which we'll train on some artifically generated data using a range of optimisers.
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "ba82988994ec42e9cc63d42b118605ce", "grade": false, "grade_id": "cell-51814571a361e8a4", "locked": true, "schema_version": 3, "solution": false, "task": false} id="o2LKot32e8NK"
# We're going to use a library called celluloid to make animations that work on colab
try:
from celluloid import Camera
except:
# !pip install celluloid
from IPython.display import HTML
import torch
import torch.optim as optim
# + [markdown] deletable=false editable=false id="a31Upx80S0Wf" nbgrader={"cell_type": "markdown", "checksum": "ef09680740eca5e908063bd1cbc2f8b5", "grade": false, "grade_id": "cell-ad34b4924532e881", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## SVM Recap
#
# Recall that an SVM tries to find the maximum margin hyperplane which separates the data classes. For a soft margin SVM
# where $\textbf{x}$ is our data, we minimize:
#
# \begin{equation}
# \left[\frac 1 n \sum_{i=1}^n \max\left(0, 1 - y_i(\textbf{w}\cdot \textbf{x}_i - b)\right) \right] + \lambda\lVert \textbf{w} \rVert^2
# \end{equation}
#
# We can formulate this as an optimization over our weights $\textbf{w}$ and bias $b$, where we minimize the
# hinge loss subject to a level 2 weight decay term. The hinge loss for some model outputs
# $z = \textbf{w}\textbf{x} + b$ with targets $y$ is given by:
#
# \begin{equation}
# \ell(y,z) = \max\left(0, 1 - yz \right)
# \end{equation}
#
# First, complete the following function to implement the hinge loss for batches of predictions `y_pred` and targets `y_true`. You should return the mean of the hinge loss across the batch. Note that this is a binary problem with labels are chosen to be $\{-1,1\}$.
# + deletable=false id="a-0v2QecS6YP" nbgrader={"cell_type": "code", "checksum": "4a913c1bb199d596ad24ad4408c9eeb0", "grade": false, "grade_id": "cell-420f491f3b45382b", "locked": false, "schema_version": 3, "solution": true, "task": false}
def hinge_loss(y_pred, y_true):
return torch.mean(torch.clamp(1 - y_pred * y_true, min=0))
raise NotImplementedError()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "75107b63da640aa87fdbbd284dacb5e2", "grade": false, "grade_id": "cell-5057286d33fba508", "locked": true, "schema_version": 3, "solution": false, "task": false} id="VLBNidVze8NM"
# ## Defining the SVM
#
# Defining the SVM is pretty simple - it's just a basic linear classifier like a Perceptron; what distinguishes it is the loss. We'll wrap it up in a function:
# + deletable=false editable=false id="27hRy0i8Sze4" nbgrader={"cell_type": "code", "checksum": "8fecc13d5366fa71717a03cf02f1c883", "grade": false, "grade_id": "cell-41c9a1a8a2140213", "locked": true, "schema_version": 3, "solution": false, "task": false}
def svm(x, w, b):
h = (w*x).sum(1) + b
return h
# + [markdown] deletable=false editable=false id="diJonMlwS7z4" nbgrader={"cell_type": "markdown", "checksum": "8477a6a51fa5c79348734e5f43c648f1", "grade": false, "grade_id": "cell-92a878d686b53c98", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Creating Synthetic Data
# -----------------------------------------------
#
# Now for some data, 1024 samples should do the trick. We normalise here so that our random init is in the same space as
# the data:
# + deletable=false editable=false id="U4U7FpoiS946" nbgrader={"cell_type": "code", "checksum": "d2e3a87b86eb49d5a5420123397df03d", "grade": false, "grade_id": "cell-210ee9436a431b1d", "locked": true, "schema_version": 3, "solution": false, "task": false} colab={"base_uri": "https://localhost:8080/"} outputId="8ad3b72d-0e96-424c-d6f7-833a16aa0682"
import numpy as np
from sklearn.datasets.samples_generator import make_blobs
X, Y = make_blobs(n_samples=1024, centers=2, cluster_std=1.2, random_state=1)
X = (X - X.mean()) / X.std()
Y[np.where(Y == 0)] = -1
X, Y = torch.FloatTensor(X), torch.FloatTensor(Y)
print(X.shape)
print(Y.shape)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "bb5c51de1529fce62d95430e304b787f", "grade": false, "grade_id": "cell-a5d0c7fc6368409c", "locked": true, "schema_version": 3, "solution": false, "task": false} id="PrHKpXGUe8NQ"
# For the first time, we're going to do proper mini-batch gradient descent. As such, we actually need to be able to produce batches of data. PyTorch has the concept of datasets (which represent entire collections of data) and data loaders (which allow us to iterate batches of data from a dataset). This allows the framework to do all the hard work for us:
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "3a3dc4137d085f28bb284143c91736f8", "grade": false, "grade_id": "cell-e65bbcc750fada1a", "locked": true, "schema_version": 3, "solution": false, "task": false} id="VQPVZI1ye8NS"
from torch.utils import data
dataset = data.TensorDataset(X,Y) # create your datset
dataloader = data.DataLoader(dataset, batch_size=32, shuffle=True) # create your dataloader
# + [markdown] deletable=false editable=false id="OR_iJX_6TJRF" nbgrader={"cell_type": "markdown", "checksum": "2653001f82321a71bde17f19a62975c6", "grade": false, "grade_id": "cell-ba76c22eaa870aa9", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Visualizing the Training
# ----------------------------------------
#
# We now aim to create a nice visualisation, such as the one below, that shows what happens as our SVM learns.
#
# 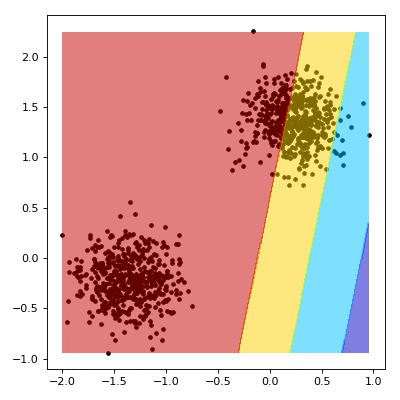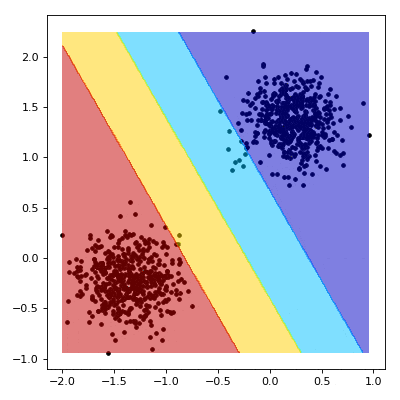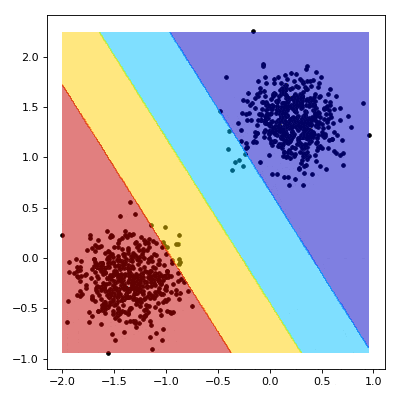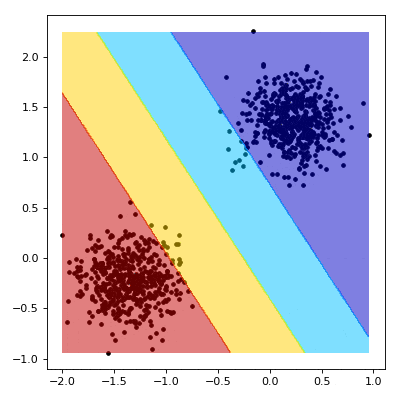
#
# The code for the visualisation (using [pyplot](https://matplotlib.org/api/pyplot_api.html)) is a bit ugly but we'll
# try to explain it to some degree. First, we need a mesh grid `xy` over the range of our data:
# + deletable=false editable=false id="9WWuOIt5TeAA" nbgrader={"cell_type": "code", "checksum": "2b0a1274964e386c9899b503c6db3e88", "grade": false, "grade_id": "cell-527fa37ff55bde6c", "locked": true, "schema_version": 3, "solution": false, "task": false}
delta = 0.01
x = np.arange(X[:, 0].min(), X[:, 0].max(), delta)
y = np.arange(X[:, 1].min(), X[:, 1].max(), delta)
x, y = np.meshgrid(x, y)
xy = list(map(np.ravel, [x, y]))
# + [markdown] deletable=false editable=false id="Wm9gBsuzTy7t" nbgrader={"cell_type": "markdown", "checksum": "fde84120db02f77f649b1ac393a38591", "grade": false, "grade_id": "cell-8e0141bbdb2b155e", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Now things get a little strange. We start by evaluating our model over the mesh grid from earlier.
#
# For our outputs $z \in \textbf{Z}$, we can make some observations about the decision boundary. First, that we are
# outside the margin if $z \lt -1$ or $z \gt 1$. Conversely, we are inside the margine where $z \gt -1$
# or $z \lt 1$.
#
# This whole process is shown in the function below, which we can call at the end of every epoch. The `camera` takes snapshots of the current plot and is used later to render a video.
# + deletable=false editable=false id="QEcC8BsoTzQ9" nbgrader={"cell_type": "code", "checksum": "785e22b2bb4b7f867908edc9792e8536", "grade": false, "grade_id": "cell-6efabcf77a2e0515", "locked": true, "schema_version": 3, "solution": false, "task": false}
import matplotlib
import matplotlib.pyplot as plt
def draw_margin(w, b, camera):
w = w.data.numpy()
b = b.data.numpy()
z = (w.dot(xy) + b).reshape(x.shape)
z[np.where(z > 1.)] = 4
z[np.where((z > 0.) & (z <= 1.))] = 3
z[np.where((z > -1.) & (z <= 0.))] = 2
z[np.where(z <= -1.)] = 1
plt.scatter(x=X[:, 0], y=X[:, 1], c="black", s=10)
plt.contourf(x, y, z, cmap=plt.cm.jet, alpha=0.5)
camera.snap()
# + [markdown] deletable=false editable=false id="GAdaOug0S_Nf" nbgrader={"cell_type": "markdown", "checksum": "3a0668d3f64bd7f21a64312127013411", "grade": false, "grade_id": "cell-f15891d25a807ea4", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Since we don't know that our data is linearly separable, we would like to use a soft-margin SVM. That is, an SVM for
# which the data does not all have to be outside of the margin. This takes the form of a weight decay term,
# $\lambda\lVert \textbf{w} \rVert^2$ in the above equation. This term is called weight decay because the gradient
# corresponds to subtracting some amount ($2\lambda\textbf{w}$) from our weights at each step.
#
# Most PyTorch optimisers actually have weight decay built in to them as an option (`weight_decay=...`), so its trivial to incorporate this.
#
# At this point we are ready to create and train our model. We've written most of the code, but you'll need to implement the forward and backward pass:
# + deletable=false id="gpKBohTtTHdr" nbgrader={"cell_type": "code", "checksum": "531b2437be629976814f210aeaf54246", "grade": false, "grade_id": "cell-1631d2d34dd5f1d3", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/", "height": 381} outputId="f0c30dc7-c6d7-4c43-c051-2b2323153ac6"
# Set up drawing
fig = plt.figure(figsize=(5, 5))
camera = Camera(fig)
w = torch.randn(1, 2, requires_grad=True)
b = torch.randn(1, requires_grad=True)
opt = optim.SGD([w,b], lr=0.1, weight_decay=0.01)
for epoch in range(50):
for batch in dataloader:
opt.zero_grad()
output = svm(batch[0],w,b)
loss = hinge_loss(output, batch[1])
loss.backward()
# raise NotImplementedError()
opt.step()
draw_margin(w, b, camera)
# create the animation and display it
anim = camera.animate()
plt.close()
HTML(anim.to_html5_video())
# + [markdown] deletable=false editable=false id="tZwqaO7pT0le" nbgrader={"cell_type": "markdown", "checksum": "f8be0f68a23e13fddd5f2ca815055f66", "grade": false, "grade_id": "cell-5b71776c13df59d7", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Now do some further experiments. What optimiser and parameters gets you to a good solution the quickest? Do you notice that when the model is near a solution it jitters around upon each step? Can you add some kind of learning rate decay or schedule from the `torch.optim.lr_scheduler` package to reduce the learning rate over time?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "9078d7fbc64e6cd09b31c42ce52e25b8", "grade": false, "grade_id": "cell-e3497d120884f361", "locked": false, "schema_version": 3, "solution": true, "task": false} id="SgaIRns-e8NZ"
# YOUR CODE HERE
raise NotImplementedError()
|
Pytorch Practical Tasks/3_2_SVM.ipynb
|
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// # Tastaturabfragen
//
// In dieser Lesson beschäftigen wir uns mit der Möglichkeit AlgoViz-Programme über die Tastatur zu steuern und werden ein paar Beipiele vorstellen. Dabei kommen alle algorithmischen Konstrukte in Kombination zum Einsatz.
// <div class="prereq">
// <h3>Was man wissen sollte</h3>
// <div>
// MAn sollte bereits <a class="prereq" href="/user-redirect/algoviz/lessons/02_Grundlagen/14_ErsterKontaktMitObjekten.ipynb">einen ersten Kontakt mot Objekten </a> gehabt haben und <a class="prereq" href="/user-redirect/algoviz/lessons/02_Grundlagen/12_Schleifen.ipynb"> kennen.
// </div>
// </div>
// ## Erste Schritte
//
// Wir benötigen eine AlgoViz-Fenster. Das kann entweder SVG oder Turtle sein. Fangen wir mal mit einem kleinen SVG an.
// +
#include <algoviz/SVG.hpp>
#include <algoviz/Turtle.hpp>
#include <iostream>
using namespace std; // Hierdurch müssen wir nicht mehr std:: vor cout etc. schreiben
// -
AlgoViz::clear();
SVG zeichnung = SVG(100,100,"Testfenster");
// Als nächstes wollen wir darauf warten, dass in dem Fenster eine Taste gedrückt wird. Das können wir über den Befehl `zeichnung.waitForKey()` erreichen.
cout << zeichnung.waitForKey() << endl;
// Der Befehl wartet bis in das Fenster klickt und anschließend eine Taste drückt. Er gibt einen String zurück, der den Namen der Taste enthält.
// <div class="task">
// <h3>Aufgabe</h3>
// <div>
// Ermitteln Sie die Tastennamen für die Pfeiltasten heraus.
// </div>
// </div>
// Da `waitForKey()` tatsächlich wartet bis eine Taste gedrückt wurde, ist es nicht für alle Situationen geeignet. Man braucht häufig eine Möglichkeit festzustellen, ob eine Taste gedrückt wurde, ohne dass das Programm wartet. In AlgoViz geht das mit `lastKey()`. Dabei wir immer die seit dem letzten Aufruf zuletzt gedrückte Taste zurückgegeben.
//
// Am Besten kann mand das verstehen, wenn man sich das folgende Beispielprogramm ansieht. Es fragt in do-while-Schleife immer wieder nach der letzten gedrückten Taste. Der Name wird in `key`gespeichert. Wurde seit der letzten Abgrage keine Taste gedrückt ist das Ergebnis der leere String `""`. D.h. die Entscheidungsanweisung prüft, ob eine Taste gedrückt wurde und gibt sie aus.
//
// Die Schleife wiederrum wird beendet, sobald x gedückt wurde.
// +
string key;
do {
key = zeichnung.lastKey();
if ( key != "" ) {
cout << key << endl;
}
} while ( key != "x" );
cout << "Ende der Schleife" << endl;
// -
// Mit diesem Mechanismus lassen sich hervorragend Tastatursteuerungen implementieren. Hier ein ganz einfaches Beispiel mit dem die Turtle sich vorwärts bewegen lässt.
// +
AlgoViz::clear();
Turtle bowser = Turtle(400,400);
string key;
do {
key = bowser.lastKey();
if ( key == "ArrowUp" ) {
bowser.forward(1);
}
} while ( key != "x" );
cout << "Ende der Schleife" << endl;
// -
// <div class="task">
// <h3>Aufgabe</h3>
// <div>
// Ergänzen Sie das Programm um die Tasten <em>Links</em> und <em>Rechts</em>, die die Turtle
// entsprechend drehen. Mit <em>Unten</em> soll sie außerdem rückwärts gehen. Andere Tasten
// könnten für die Einstellung der Farbe genutzt werden.
// </div>
// </div>
// ## Der hüpfende Ball
//
// Mit der Tastatursteuerung lääst sich auch das Problem der endlichen Schleife für den Ball lösen. Statt einer For-Schleife, die eine feste Anzahl von Durchläufen hat, verwenden wir eine do-while-Schleife, die auf das Drücken einer bestimmten Taste wartet.
// +
AlgoViz::clear();
SVG zeichnung = SVG(400,400);
string key;
Circle ball = Circle(200,50,10,&zeichnung);
// Image tardis = Image("/user-redirect/algoviz/img/tardis.png",200,50,20,20,&zeichnung);
int x = 0;
int y = 0;
int vx = 2;
int vy = 1;
do {
key = zeichnung.lastKey();
vy = vy + 1;
x = x + vx;
y = y + vy;
if ((x < 0) || (x > 399)) {
vx = -vx;
}
if ((y < 0) || (y > 399)) {
vy = -vy;
}
ball.moveTo(x,y);
AlgoViz::sleep(10);
} while ( key != "x" );
// -
string
// +
int x = 200;
int y = 20;
int vx = 0;
int vy = 0;
int ax = 0;
int ay = 1;
int winkel = 0;
std::string key;
do {
vx = vx + ax;
vy = vy + ay;
x = x + vx;
y = y + vy;
if ( y >= 400 ) {
y = (800-y);
vy = -vy;
}
if ( x < 0 ) {
x = -x;
vx = -vx;
}
if ( x >= 400 ) {
x = (800-x);
vx = -vx;
}
winkel = (winkel+10) % 360;
tardis.rotateTo(winkel);
tardis.moveTo(x,y);
AlgoViz::sleep(20);
key = zeichnung.lastKey();
if ( key == "ArrowLeft" ) vx = vx - 1;
if ( key == "ArrowRight" ) vx = vx + 1;
if ( key == "0" ) vx = 0;
} while ( key != "x" );
// -
// <div class="followup">
// <h3>Wo es weiter geht</h3>
// <div>
// <a class="followup" href="/user-redirect/algoviz/lessons/02_Grundlagen/19_BouncingBall.ipynb">Das Beispiel des hüpfenden Balls</a>.
// </div>
// </div>
|
lessons/02_Grundlagen/18_Tastatur.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # wagtail install
# +
$ pip install wagtail
$ wagtail start mysite
$ cd mysite
$ pip install -r requirements.txt
$ ./manage.py migrate
$ ./manage.py createsuperuser
$ ./manage.py runserver
# -
# wagtail start mysite创建基本的DJANGO CMS(wagtail)项目模板,只包含了基本的配置项和home APP.
# pip install -r requirements.txt 安装其它所需要的基本组件。
# # settings
|
netplan/docs/01.CreateDjangoCmsProject_wagtail.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A couple of practical things
#
# ## Overview
#
# As in any class, we're going to use a couple of 'technologies' in order to learn the material. Below a couple of videos to explain.
#
# * The first thing is Github. You've already worked with this (in fact, you're probably looking at this notebook file on the web), but in the video below, I provide some more examples.
#
# * The second tool is Python and Jupyter Notebooks. While you should be by now very familiar with it, please read carefully the instructions to make sure we all use the same Python version, etc...
# ## Github to organize things
#
# The video is taken from another course run by <NAME>.
# We will use Github in exactly the same way, so this video explains most of how the course work.
#
# If you are not familiar with Git/Github, I suggest going through these two tutorials:
# * [Intro Git Tutorial](http://jmausolf.github.io/code/intro_git/)
# * [Intermediate Git Tutorial](http://jmausolf.github.io/code/intermediate_git/)
from IPython.display import YouTubeVideo
YouTubeVideo("42KjmxgYYNI",width=800, height=450)
# # Python
# The exercise you will solve during the class require using Python and Jupyter Notebooks.
# I suggest to use the Anaconda distribution for Python 3.8. Note that if you want to use another setup, that's fine, but I cannot promise to help you with anything other than Anaconda.
#
# I suggest to follow these instructions for set up:
#
# * Download the _Anaconda distribution_ of Python [**here**](https://www.anaconda.com/download/).
# * Create an Anaconda [environment](https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html) for this class: `conda create -n comp_soc_sci_2021 python=3.8`
# * Activate the environment "`conda activate comp_soc_sci_2021`", start up the notebook by typing "`jupyter notebook`" and your terminal, and everything should be ready to use in your favorite browser.
# * Be sure to check the keyboards shortcuts under the heading of "Help" where you will find for instance shortcut to code-completion (Tab) and tooltip with documentation (Shift-Tab) which will save you a ton of time.
#
#
# ### Super important notice
#
# Everything we do going forward in this class will depend on you being comfortable with Python. There is simply **no way** that you will be able to do well, if you're also struggling with Python on top of everything else you'll be learning.
#
# **So if you're not 100% comfortable with Python, I recommend you follow a tutorial to teach you Python, for example [this one](https://www.learnpython.org), before proceeding**.
|
lectures/AdminStuff.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: feml
# language: python
# name: feml
# ---
import pandas as pd
# +
# first, let's create a toy dataframe with some timestamps in different time zones
# variable 1
df = pd.DataFrame()
df['time1'] = pd.concat([
pd.Series(
pd.date_range(
start='2015-06-10 09:00', freq='H', periods=3,
tz='Europe/Berlin')),
pd.Series(
pd.date_range(
start='2015-09-10 09:00', freq='H', periods=3, tz='US/Central'))
], axis=0)
df
# +
# first, let's create a toy dataframe with some timestamps in different time zones
# variable 2
df['time2'] = pd.concat([
pd.Series(
pd.date_range(
start='2015-07-01 09:00', freq='H', periods=3,
tz='Europe/Berlin')),
pd.Series(
pd.date_range(
start='2015-08-01 09:00', freq='H', periods=3, tz='US/Central'))
], axis=0)
df
# +
# to work with different time zones, first we unify the timezone to the central one
# setting utc = True
df['time1_utc'] = pd.to_datetime(df['time1'], utc=True)
df['time2_utc'] = pd.to_datetime(df['time2'], utc=True)
df
# +
# let's explore the variable type
df['elapsed_days'] = (df['time2_utc'] - df['time1_utc']).dt.days
df['elapsed_days'].head()
# +
# next we change all timestamps to the desired timezone, eg Europe/London
# in this example
df['time1_london'] = df['time1_utc'].dt.tz_convert('Europe/London')
df['time2_berlin'] = df['time1_utc'].dt.tz_convert('Europe/Berlin')
df[['time1_london', 'time2_berlin']]
|
Chapter07/Recipe6--different-time-zones.ipynb
|
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [Intro to Deep Learning](https://www.kaggle.com/learn/intro-to-deep-learning) course. You can reference the tutorial at [this link](https://www.kaggle.com/ryanholbrook/deep-neural-networks).**
#
# ---
#
# # Introduction #
#
# In the tutorial, we saw how to build deep neural networks by stacking layers inside a `Sequential` model. By adding an *activation function* after the hidden layers, we gave the network the ability to learn more complex (non-linear) relationships in the data.
#
# In these exercises, you'll build a neural network with several hidden layers and then explore some activation functions beyond ReLU. Run this next cell to set everything up!
# +
import tensorflow as tf
# Setup plotting
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
# Set Matplotlib defaults
plt.rc('figure', autolayout=True)
plt.rc('axes', labelweight='bold', labelsize='large',
titleweight='bold', titlesize=18, titlepad=10)
# Setup feedback system
from learntools.core import binder
binder.bind(globals())
from learntools.deep_learning_intro.ex2 import *
# -
# In the *Concrete* dataset, your task is to predict the compressive strength of concrete manufactured according to various recipes.
#
# Run the next code cell without changes to load the dataset.
# +
import pandas as pd
concrete = pd.read_csv('../input/dl-course-data/concrete.csv')
concrete.head()
# -
# # 1) Input Shape #
#
# The target for this task is the column `'CompressiveStrength'`. The remaining columns are the features we'll use as inputs.
#
# What would be the input shape for this dataset?
# +
# YOUR CODE HERE
input_shape = ____
# Check your answer
q_1.check()
# +
# Lines below will give you a hint or solution code
#q_1.hint()
#q_1.solution()
# -
# # 2) Define a Model with Hidden Layers #
#
# Now create a model with three hidden layers, each having 512 units and the ReLU activation. Be sure to include an output layer of one unit and no activation, and also `input_shape` as an argument to the first layer.
# +
from tensorflow import keras
from tensorflow.keras import layers
# YOUR CODE HERE
model = ____
# Check your answer
q_2.check()
# +
# Lines below will give you a hint or solution code
#q_2.hint()
#q_2.solution()
# -
# # 3) Activation Layers #
#
# Let's explore activations functions some.
#
# The usual way of attaching an activation function to a `Dense` layer is to include it as part of the definition with the `activation` argument. Sometimes though you'll want to put some other layer between the `Dense` layer and its activation function. (We'll see an example of this in Lesson 5 with *batch normalization*.) In this case, we can define the activation in its own `Activation` layer, like so:
#
# ```
# layers.Dense(units=8),
# layers.Activation('relu')
# ```
#
# This is completely equivalent to the ordinary way: `layers.Dense(units=8, activation='relu')`.
#
# Rewrite the following model so that each activation is in its own `Activation` layer.
# +
### YOUR CODE HERE: rewrite this to use activation layers
model = keras.Sequential([
layers.Dense(32, activation='relu', input_shape=[8]),
layers.Dense(32, activation='relu'),
layers.Dense(1),
])
# Check your answer
q_3.check()
# +
# Lines below will give you a hint or solution code
#q_3.hint()
#q_3.solution()
# -
# # Optional: Alternatives to ReLU #
#
# There is a whole family of variants of the `'relu'` activation -- `'elu'`, `'selu'`, and `'swish'`, among others -- all of which you can use in Keras. Sometimes one activation will perform better than another on a given task, so you could consider experimenting with activations as you develop a model. The ReLU activation tends to do well on most problems, so it's a good one to start with.
#
# Let's look at the graphs of some of these. Change the activation from `'relu'` to one of the others named above. Then run the cell to see the graph. (Check out the [documentation](https://www.tensorflow.org/api_docs/python/tf/keras/activations) for more ideas.)
# +
# YOUR CODE HERE: Change 'relu' to 'elu', 'selu', 'swish'... or something else
activation_layer = layers.Activation('relu')
x = tf.linspace(-3.0, 3.0, 100)
y = activation_layer(x) # once created, a layer is callable just like a function
plt.figure(dpi=100)
plt.plot(x, y)
plt.xlim(-3, 3)
plt.xlabel("Input")
plt.ylabel("Output")
plt.show()
# -
# # Keep Going #
#
# Now move on to Lesson 3 and [**learn how to train neural networks**](https://www.kaggle.com/ryanholbrook/stochastic-gradient-descent) with stochastic gradient descent.
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/191966) to chat with other Learners.*
|
corso-data-science-2021/hands-on/05-geovis-and-dnn/exercises/exercise-deep-neural-networks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_pytorch_p36)
# language: python
# name: conda_pytorch_p36
# ---
# +
% load_ext autoreload
% autoreload 2
% matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import torch
device = 'cuda' if torch.cuda.is_available() else 'cpu'
from copy import deepcopy
from sklearn.linear_model import LinearRegression
from sklearn import metrics
from sklearn.model_selection import cross_val_score
import time
# adaptive-wavelets modules
from awd.data.biology import get_dataloader, load_pretrained_model
from awd.utils.misc import tuple_to_tensor
from awd.trim import TrimModel
# evaluation
from eval_biology import load_results, max_transformer
from ex_biology import p
# -
# # load results
dirs = [
"db5_saliency_warmstart_seed=1",
"db5_saliency_warmstart_seed=10",
"db5_saliency_warmstart_seed=100",
"db5_saliency_warmstart_seed=1000",
"db5_saliency_warmstart_seed=10000"
]
dics, results, models = load_results(dirs)
# # prediction accuracy, compression, and computation time
def r2_bootstrap(y, y_pred, m=10000):
"""Return bootstrap mean and std error."""
np.random.seed(p.seed)
e = []
for j in range(m):
idx = np.arange(len(y))
sel = np.random.choice(idx, len(idx), replace=True)
e.append(metrics.r2_score(y[sel], y_pred[sel]))
return metrics.r2_score(y_test, preds), np.std(e)
# +
r = {
'Standard Wavelet (DB5)': [],
'AWD (Ours)': [],
'LSTM': [],
}
s = {
'Standard Wavelet (DB5)': [],
'AWD (Ours)': [],
'LSTM': [],
}
c = {
'Standard Wavelet (DB5)': [],
'AWD (Ours)': []
}
t = {
'Standard Wavelet (DB5)': [],
'AWD (Ours)': [],
'LSTM': [],
}
best_lam = []
best_wt = []
# params for feature transformer
sgn = "pos"
m = 6
for index in range(len(dirs)):
res = results[index]
mos = models[index]
# load data
(train_loader, test_loader) = get_dataloader(p.data_path,
batch_size=p.batch_size,
is_continuous=True)
# cross validation
reg_score = []
for i, wt in enumerate(list(dics[index]['wt'].values())):
wt = wt.to('cpu')
(X, y), (X_test, y_test) = max_transformer(wt,
train_loader,
test_loader,
sgn=sgn,
m=m)
clf = LinearRegression()
scores = cross_val_score(clf, X, y, cv=5, scoring='r2')
reg_score.append(scores.mean())
reg_score = np.array(reg_score)
# select best wavelet
idx1, idx2 = list(dics[index]['wt'].keys())[np.argmax(reg_score).flatten()[0]]
idx = dics[index]['index'][(idx1, idx2)]
lamL1wave = dics[index]['lamL1wave'][(idx1, idx2)]
lamL1attr = dics[index]['lamL1attr'][(idx1, idx2)]
best_lam.append((lamL1wave, lamL1attr))
wt = dics[index]['wt'][(idx1, idx2)]
best_wt.append(wt)
# load pre-trained model
model = load_pretrained_model(p.model_path, device=device)
p.batch_size = 3000
(train_loader, test_loader) = get_dataloader(p.data_path,
batch_size=p.batch_size,
is_continuous=True)
(X, y), (X_test, y_test) = max_transformer(wt,
train_loader,
test_loader,
sgn=sgn,
m=m)
clf = LinearRegression().fit(X, y)
start_time = time.time()
preds = clf.predict(X_test)
t['AWD (Ours)'].append(time.time() - start_time)
acc, std = r2_bootstrap(y_test, preds)
r['AWD (Ours)'].append(acc)
s['AWD (Ours)'].append(std)
wt_o = adaptive_wavelets.DWT1d(wave='db5', mode='zero', J=4, init_factor=1, noise_factor=0.0)
(X, y), (X_test, y_test) = max_transformer(wt_o,
train_loader,
test_loader,
sgn=sgn,
m=m)
clf_o = LinearRegression().fit(X, y)
start_time = time.time()
preds = clf_o.predict(X_test)
t['Standard Wavelet (DB5)'].append(time.time() - start_time)
acc, std = r2_bootstrap(y_test, preds)
r['Standard Wavelet (DB5)'].append(acc)
s['Standard Wavelet (DB5)'].append(std)
preds = []
y_test = []
start_time = time.time()
for data, labels in test_loader:
preds.append(model(data).detach().numpy())
y_test.append(labels.detach().numpy())
t['LSTM'].append(time.time() - start_time)
preds = np.vstack(preds)
y_test = np.vstack(y_test)
acc, std = r2_bootstrap(y_test, preds)
r['LSTM'].append(acc)
s['LSTM'].append(std)
# define trim model
mt = TrimModel(model, wt.inverse, use_residuals=True)
mt_o = TrimModel(model, wt_o.inverse, use_residuals=True)
attributer = adaptive_wavelets.Attributer(mt, attr_methods='Saliency', device='cuda')
attributer_o = adaptive_wavelets.Attributer(mt_o, attr_methods='Saliency', device='cuda')
# compute compression rate and representations
attrs = {'AWD': torch.tensor([]).to(device),
'DB5': torch.tensor([]).to(device)}
reps = {'AWD': torch.tensor([]).to(device),
'DB5': torch.tensor([]).to(device)}
wt, wt_o = wt.to(device), wt_o.to(device)
for data, _ in test_loader:
data = data.to(device)
i = 0
for w in [wt, wt_o]:
if i == 0:
data_t = w(data)
with torch.backends.cudnn.flags(enabled=False):
attributions = attributer(data_t, target=0, additional_forward_args=deepcopy(data))
y, _ = tuple_to_tensor(data_t)
reps['AWD'] = torch.cat((reps['AWD'], y), dim=0)
z, _ = tuple_to_tensor(attributions)
attrs['AWD'] = torch.cat((attrs['AWD'], z), dim=0)
else:
data_t = w(data)
with torch.backends.cudnn.flags(enabled=False):
attributions = attributer_o(data_t, target=0, additional_forward_args=deepcopy(data))
y, _ = tuple_to_tensor(data_t)
reps['DB5'] = torch.cat((reps['DB5'], y), dim=0)
z, _ = tuple_to_tensor(attributions)
attrs['DB5'] = torch.cat((attrs['DB5'], z), dim=0)
i += 1
reps['AWD'] = reps['AWD'].reshape(-1)
reps['DB5'] = reps['DB5'].reshape(-1)
attrs['AWD'] = attrs['AWD'].reshape(-1)
attrs['DB5'] = attrs['DB5'].reshape(-1)
thresh1 = 1e-3
thresh2 = 1e-3
c_rate_AWD = 1.0 * ((abs(reps['AWD']) > thresh1) & (abs(attrs['AWD']) > thresh2)).sum() / reps['AWD'].shape[0]
c_rate_DB5 = 1.0 * ((abs(reps['DB5']) > thresh1) & (abs(attrs['DB5']) > thresh2)).sum() / reps['DB5'].shape[0]
c['AWD (Ours)'].append(c_rate_AWD.item())
c['Standard Wavelet (DB5)'].append(c_rate_DB5.item())
# -
print("Acc :", np.array(r['AWD (Ours)']).mean().round(3), np.array(r['Standard Wavelet (DB5)']).mean().round(3),
np.array(r['LSTM']).mean().round(3))
print("Std :", np.array(r['AWD (Ours)']).std().round(3))
print("Comp :", np.array(c['AWD (Ours)']).mean().round(3), np.array(c['Standard Wavelet (DB5)']).mean().round(3))
print("Std :", np.array(c['AWD (Ours)']).std().round(3))
print("Time :", np.array(t['AWD (Ours)']).mean().round(4), np.array(t['Standard Wavelet (DB5)']).mean().round(4),
np.array(t['LSTM']).mean().round(4))
print("Std :", np.array(t['AWD (Ours)']).std().round(4))
# # plot wavelets
# +
# plot
num_rows = 5
num_cols = 2
titsize = 6
fig = plt.figure(dpi=300, figsize=(2, 4))
for j in range(len(dirs)):
wt = best_wt[j]
phi, psi, x = adaptive_wavelets.get_wavefun(wt)
plt.subplot(num_rows, num_cols, j * num_cols + 1)
plt.plot(x, phi)
plt.axis('off')
if j == 0:
plt.title('scaling function', fontsize=titsize)
plt.subplot(num_rows, num_cols, j * num_cols + 2)
plt.plot(x, psi)
plt.axis('off')
if j == 0:
plt.title('wavelet function', fontsize=titsize)
plt.tight_layout()
plt.subplots_adjust(wspace=0.0, hspace=0.05)
# plt.savefig('figures/bio_wave.pdf', bbox_inches='tight')
# -
|
notebooks/biology/03_analyze.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pulling data from (open) REST APIs
#
# [Big source of public APIs](https://rapidapi.com/collection/list-of-free-apis)
#
# We have already seen how to use `requests` to fetch a webpage:
import requests
r = requests.get('http://www.cnn.com')
r.text[0:300]
# If the URL is to a page that gives you HTML, we would say that we are fetching a webpage. On the other hand, if the URL is returning data in some form, we would say that we are accessing a *REST* api.
#
# **REST** is an acronym for *REpresentational State Transfer* and is a very handy way to make something trivial sound very complicated. Anytime you see the word REST, just think "webpage that gives me data not HTML." There is a massive industry and giant following behind this term but I don't see anything beyond "fetch data from webpage".
#
# Anyway, we are going to pull data from web servers that intentionally provide nice data spigot URLs. Information you need in order to get data is typically:
#
# * Base URL, including machine name, port number, and "file" path
# * The names and values of parameters
# * What data comes back and in what format (XML, JSON, CSV, ...)
# ## Looking up word definitions
#
# The [dictionaryapi.dev](https://dictionaryapi.dev/) API lets us look up words in various languages and get the definitions. The format of the URL to access the API is just:
#
# ```
# https://api.dictionaryapi.dev/api/v2/entries/<language_code>/<word>
# ```
#
# So, we can get the English definition for *science* like this (and parse the json result):
# +
import requests
import json
r = requests.get('https://api.dictionaryapi.dev/api/v2/entries/en_US/science')
data = json.loads(r.text)
data
# -
# The JSON looks like the following when formatted in the browser (I think I have a JSON viewer plug-in).
#
# <img src="figures/dictionary-science-json.png" width="400">
#
# That looks like there is a list with one element, which is the actual dictionary of stuff we want so `data[0]` is the dictionary of stuff. This lets us get access to the definition and phonetics if we dig down.
data = data[0]
phonetic = data['phonetic']
sciencedef = data['meanings'][0]['definitions'][0]['definition']
print(phonetic)
print(sciencedef)
# **Exercise**: Print out the origin of the word science from that JSON.
# **Exercise**: Use the API to fetch and print out the definition of *Merhaba* (a greeting) in the Turkish language. The result should be *karşılaşıldığında söylenilen bir selamlaşma sözü.* (*a word of greeting when encountered.*)
# ## JSON from openpayments.us
#
# (This site seems to go down a lot when they reboot our computer science machine so forgive me if it's not up...)
#
# Now, let's look at a website that will give us JSON data: [www.openpayments.us](http://www.openpayments.us).
#
# There is a REST data API available at URL template:
#
# ```
# URL = f"http://openpayments.us/data?query={q}" # for some q
# ```
# **Exercise**: Use `curl` to fetch data about a doctor.
#
# Here's how to fetch the data for a doctor's name, such as `<NAME>`:
# + tags=[]
import requests
import json
import sys
name = "<NAME>"
URL = f"http://openpayments.us/data?query={name}"
r = requests.get(URL)
data = json.loads(r.text)
print(json.dumps(data)[0:1000])
# -
# This website gives you JSON, which is very easy to load and dump using the default `json` package as you can see from that code snippet. As before, you can grab one of the elements using dictionary like indexing:
results = data['results']
results[0:2]
# It is convenient to look at the records in a data frame:
import pandas as pd
pd.DataFrame.from_dict(results).head(3)
# A **technical detail** related to valid strings you can include as part of a URL. Spaces are not allowed so `<NAME>` has to be encoded or "quoted". Fortunately, `requests` does this automatically for us. If you ever need to quote parameter values in URLs, you can do this:
#
# ```python
# from urllib.parse import quote
# value = quote(value)
# ```
#
# Because `&` is the separator between parameters, it is also invalid in a parameter name or value. Here are some example conversions:
#
# ```python
# >>> quote("<NAME>")
# 'john%20chan'
# >>> quote("john&chan")
# 'john%26chan'
# ```
#
# The conversion uses the ASCII character code (in 2-digit hexadecimal) for space and ampersand. Sometimes you will see the space converted to a `+`, which also works: `John+Chan`.
|
notes/openapi.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import defaultdict
class Solution(object):
def nextGreaterElement(self, nums1, nums2):
"""
:type nums1: List[int]
:type nums2: List[int]
:rtype: List[int]
"""
value_to_greater = defaultdict(lambda: -1)
stack = []
for num in nums2:
while stack and num > stack[-1]:
value_to_greater[stack.pop()] = num
stack.append((num))
return [value_to_greater[num] for num in nums1]
# -
s = Solution()
s.nextGreaterElement([4,1,2], [1,3,4,2])
|
algorithms/496-next-greater-element-i.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import functools
# ## Part 2: Plotting
# Part 2: Plotting
data = np.genfromtxt ('ex1data1.txt', delimiter=",")
X = np.matrix(data[:, 0]).T
y = np.matrix(data[:, 1]).T
m = len(y)
plt.scatter(X, y, alpha=0.7)
ones = np.ones((m, 1))
X = np.hstack((ones, X)) # Add a column of ones to x
# ## Part 3: Cost and Gradient descent
# +
def derive(f):
def dfdx(x, step):
dy = f(x + step) - f(x)
dx = step
#print("dx, dy\n", dx,"\n", dy, "\n")
return dy/dx
return dfdx
def computeCost(X, y, theta):
m = len(y)
costs = np.power((X*theta - y), 2)
J = (sum(costs)) / (2*m)
return J
partialComputeCost = functools.partial(computeCost, X, y)
dcostdtheta = derive(partialComputeCost)
def gradientDescent(X, y, theta, alpha, num_iters):
"""
GRADIENTDESCENT Performs gradient descent to learn theta
theta = GRADIENTDESCENT(X, y, theta, alpha, num_iters) updates theta by
taking num_iters gradient steps with learning rate alpha
"""
# Initialize some useful values
m = len(y) # number of training examples
J_history = np.zeros((num_iters, 1))
#print("theta0", theta)
for i in range(num_iters):
J_history[i] = computeCost(X, y, theta)
#print("derviation", derive(partialCost, i).T)
for j in range(len(theta)):
temp = np.matrix(np.zeros(len(theta))).T
temp[j] = alpha
#print(temp)
theta[j] = theta[j] - alpha * dcostdtheta(theta, temp)[j]
#print(J_history[i])
print(theta)
print("Result, theta = ", theta, "with cost = ", J_history[-1])
return theta, J_history
# -
# +
theta = np.matrix('10 ; 10')
iterations, alpha = 100, 0.001 # Some gradient descent settings
theta, cost = gradientDescent(X, y, theta, alpha, iterations);
# -
cost
# +
iterations, alpha = 1500, 0.01 # Some gradient descent settings
print('\nTesting the cost function ...\n')
# compute and display initial cost
theta = np.zeros((2, 1))
J = computeCost(X, y, theta);
print('With theta = [0 ; 0]\nCost computed = ', J);
print('Expected cost value (approx) 32.07\n');
# further testing of the cost function
theta = np.matrix('-1 ; 2')
J = computeCost(X, y, theta);
print('\nWith theta = [-1 ; 2]\nCost computed = ', J);
print('Expected cost value (approx) 54.24\n');
print('\nRunning Gradient Descent ...\n')
# run gradient descent
theta = gradientDescent(X, y, theta, alpha, iterations);
# print theta to screen
print('Theta found by gradient descent:\n');
fprintf('%f\n', theta);
fprintf('Expected theta values (approx)\n');
fprintf(' -3.6303\n 1.1664\n\n');
% Plot the linear fit
hold on; % keep previous plot visible
plot(X(:,2), X*theta, '-')
legend('Training data', 'Linear regression')
hold off % don't overlay any more plots on this figure
% Predict values for population sizes of 35,000 and 70,000
predict1 = [1, 3.5] *theta;
fprintf('For population = 35,000, we predict a profit of %f\n',...
predict1*10000);
predict2 = [1, 7] * theta;
fprintf('For population = 70,000, we predict a profit of %f\n',...
predict2*10000);
fprintf('Program paused. Press enter to continue.\n');
pause;
# -
# ## Part 4: Visualizing J(theta_0, theta_1)
# +
fprintf('Visualizing J(theta_0, theta_1) ...\n')
% Grid over which we will calculate J
theta0_vals = linspace(-10, 10, 100);
theta1_vals = linspace(-1, 4, 100);
% initialize J_vals to a matrix of 0's
J_vals = zeros(length(theta0_vals), length(theta1_vals));
% Fill out J_vals
for i = 1:length(theta0_vals)
for j = 1:length(theta1_vals)
t = [theta0_vals(i); theta1_vals(j)];
J_vals(i,j) = computeCost(X, y, t);
end
end
% Because of the way meshgrids work in the surf command, we need to
% transpose J_vals before calling surf, or else the axes will be flipped
J_vals = J_vals';
% Surface plot
figure;
surf(theta0_vals, theta1_vals, J_vals)
xlabel('\theta_0'); ylabel('\theta_1');
% Contour plot
figure;
% Plot J_vals as 15 contours spaced logarithmically between 0.01 and 100
contour(theta0_vals, theta1_vals, J_vals, logspace(-2, 3, 20))
xlabel('\theta_0'); ylabel('\theta_1');
hold on;
plot(theta(1), theta(2), 'rx', 'MarkerSize', 10, 'LineWidth', 2);
# -
# Testing derivative function
STEP = 0.01
def cube(x): return x*x*x
d = derive(cube)
dd = derive(d)
xs = np.arange(-10, 10, .1)
plt.plot(xs, cube(xs))
plt.plot(xs, d(xs, STEP))
plt.plot(xs, dd(xs, STEP))
print(dd)
|
Machine Learning - Coursera/machine-learning-ex1/ex1/_ex1 - Copy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%
# %%
from sect_sens.constants import get_plotpath
from sect_sens.util.practical_functions import make_folders
from sect_sens.util.slice_average import one_val_tab
# load and autoreload
from IPython import get_ipython
from sect_sens.util.naming_conventions.var_info import get_fancy_var_name, get_fancy_unit_xr
# noinspection PyBroadException
from sect_sens.util.slice_average.one_val_tab import plt_var, get_diff_by_type, get_mean_std_by_type
try:
_ipython = get_ipython()
_magic = _ipython.magic
_magic('load_ext autoreload')
_magic('autoreload 2')
except:
pass
# %%
from sect_sens.util.plot.colors import get_case_col
version='review'
plt_path = get_plotpath('one_value')
def create_filename(name):
fn= f'{plt_path}_2d_{version}{name}.'
make_folders(fn)
return fn
# %%
startyear = '0004-01'
endyear = '0008-12'
pmin = 850. # minimum pressure level
avg_over_lev = True # True#True#False#True
pressure_adjust = True # Can only be false if avg_over_lev false. Plots particular hybrid sigma lev
# %%
varl = ['N_AER', 'NCONC01', 'TGCLDCWP', 'CDNUMC', 'NCFT_Ghan', 'DIR_Ghan', 'LWDIR_Ghan', 'SWDIR_Ghan', 'SWCF_Ghan',
'LWCF_Ghan' ,'cb_SO4_NA','cb_SOA_NA','cb_NA', 'SOA_NA','SO4_NA',
'ACTNL_incld','ACTREL_incld', 'SFisoprene','SFmonoterp']
cases = ['SECTv21_ctrl_koagD', 'SECTv21_incY', 'SECTv21_decY', 'noSECTv21_ox_ricc_dd', 'noSECTv21_ox_ricc_decY',
'noSECTv21_ox_ricc_incY']
cases_sec = [
'NF1850_aeroxid2014_SECT_ctrl',
'NF1850_SECT_ctrl',
#'NF1850_aeroxid2014_SECT_ctrl_smax',
'NF1850_SECT_paas',
'NF1850_SECT_depT',
'NF1850_aeroxid2014_SECT_paas',
'NF1850_aeroxid2014_SECT_depT',
]
cases_nsec = [
#'NF1850_noSECT_def_smax',
#'NF1850_aeroxid2014_noSECT_def_smax',
'NF1850_noSECT_def',
'NF1850_aeroxid2014_noSECT_def',
'NF1850_noSECT_ox_ricc',
'NF1850_aeroxid2014_noSECT_ox_ricc',
'NF1850_noSECT_ox_ricc_depT',
'NF1850_aeroxid2014_noSECT_ox_ricc_depT',
]
# %%
varl_ex = ['FSNT','FSNT_DRF','FLNT','FLNT_DRF','FSNTCDRF']
varl =varl+ varl_ex
# %%
case_types = ['PI', 'PD']
model_types = ['OsloAeroSec','OsloAeroSec$_{paas}$','OsloAeroSec$_{depT}$', 'OsloAero$_{depT}$', 'OsloAero$_{imp}$', 'OsloAero$_{def}$'][::-1]#'OsloAeroSec$_{gord}$','OsloAero$_{gord}$'
mod_types = ['OsloAeroSec','OsloAeroSec$_{paas}$','OsloAeroSec$_{depT}$', 'OsloAero$_{depT}$', 'OsloAero$_{imp}$', 'OsloAero$_{def}$'][::-1]#'OsloAeroSec$_{gord}$','OsloAero$_{gord}$'
cdic = {key: get_case_col(key) for key in mod_types} # , ['r','g','b'])}
# %%
# varl = ['N_AER']
df2, dic_vals = one_val_tab.get_tab_yearly_mean(varl,
cases_sec + cases_nsec,
startyear,
endyear,
pmin=pmin,
pressure_adjust=pressure_adjust,
average_over_lev=avg_over_lev,
groupby='time.year', # 'time',
dims=None,
area='Global',
invert_dic=True
)
# %%
from useful_scit.imps import ( sns,pd, plt)
# %%
from sect_sens.data_info.simulation_types import get_abs_by_type
# %%
di = get_abs_by_type(dic_vals, case_types=['PI','PD'], mod_types=model_types)
di.keys()
# %%
ls =[]
for ct in di.keys():
_di = di[ct]
for cn in _di.keys():
print(cn)
_df = _di[cn]
_df['case']=cn
_df['case_type']=ct
ls.append(_df.reset_index())
df_tot = pd.concat(ls)
df_tot
# %%
svarl = ['ACTNL_incld','cb_NA']#,'FSNT_DRF','FLNT','FLNT_DRF','FSNTCDRF']#'', 'SWCF_Ghan', 'LWCF_Ghan','DIR_Ghan']
v1 = 'ACTNL_incld'
v2 = 'ACTREL_incld'
di = get_abs_by_type(dic_vals, case_types=['PI','PD'], mod_types=model_types)
_df = pd.DataFrame(columns=['val', 'type','var','model'])
di_var = {}
for t in di.keys():
for m in di[t].keys():
di[t][m]=di[t][m].mean()
for v in varl:
_df_v = pd.DataFrame(columns=['val', 'type','model'])
for t in di.keys():
for m in di[t].keys():
_df = _df.append(pd.DataFrame([di[t][m][v], t, v, m],index=['val', 'type','var','model']).transpose(), ignore_index=True)
_df_v=_df_v.append(pd.DataFrame([di[t][m][v], t, m],index=['val', 'type','model']).transpose(), ignore_index=True)
_df_v['val'] = pd.to_numeric(_df_v['val'])
di_var[v]=_df_v.copy()
# %%
di_2 = di_var[v].set_index(['type','model']).rename({'val':v}, axis=1)#.plot.bar()
for v in varl:
di_2[v] = di_var[v].set_index(['type','model']).rename({'val':v}, axis=1)
# %%
di_2.reset_index()
# %%
# %%
df_tot.head()
# %%
mod_types
# %%
y='NCFT_Ghan'
x='cb_NA'
_df = di_2.reset_index()
fig, ax = plt.subplots(1, figsize=[6,3], dpi=150)
for mod in model_types:
print(mod)
sdf = _df[_df['model']==mod]
c=get_case_col(mod)
plt.plot(sdf[x],sdf[y], label=mod, marker='o', c=c)
plt.text(sdf[x].iloc[0],sdf[y].iloc[0]+0.05, sdf['type'].iloc[0], color=c, size=12)
plt.text(sdf[x].iloc[1],sdf[y].iloc[1]+0.05, sdf['type'].iloc[1], color=c, size=12)
for mod in model_types:
c=get_case_col(mod)
_df = df_tot[df_tot['case']==mod][[x,y]]
plt.scatter(_df[x],_df[y], alpha=.4, edgecolor=c,facecolors='none',)
def label(v):
_n = get_fancy_var_name(v)
_u = df2[v]['unit']
return f'{_n} [{_u}]'
plt.xlabel(label(x))#'Col. burden N$_{NPF}$ [kg m$^{-2}$]')
plt.ylabel(label(y))
plt.legend(frameon=False, bbox_to_anchor=(1.05, 1), loc='upper left',)
sns.despine(fig)
fn= create_filename(f'{x}_{y}')
plt.tight_layout()
plt.savefig(fn+'pdf', dpi=300)
# %%
_df = di_2.reset_index()
_df[_df['model']=='OsloAero$_{depT}$']#"=='OsloAeroSec']
# %%
df_tot[df_tot['case']=='OsloAeroSec$_{depT}$'][['SWCF_Ghan','LWCF_Ghan','NCFT_Ghan']]#[df_tot['case']=='OsloAero$_{depT}$']
# %%
df_tot[df_tot['case']=='OsloAero$_{depT}$'][['SWCF_Ghan','LWCF_Ghan','NCFT_Ghan']]#[df_tot['case']=='OsloAero$_{depT}$']
# %%
_df['model']
# %%
trans_dic = {v:get_fancy_var_name(v) for v in varl}
# %%
rn_dic = {
'NCRE$_{Ghan}$':'ERF$_{aci}$',
'SWCRE$_{Ghan}$':'ERF$_{aci,SW}$',
'LWCRE$_{Ghan}$':'ERF$_{aci,LW}$',
'DRE$_{Ghan}$': 'ERF$_{ari}$'
}
# %%
svarl = ['NCFT_Ghan', 'SWCF_Ghan', 'LWCF_Ghan','DIR_Ghan']
df1 = df_tot[[*svarl,'case']]
df1 = df1.rename(trans_dic, axis=1)
df1 = df1.rename(rn_dic, axis=1)
df2 = pd.melt(df1,id_vars='case')
# %%
cols=[cdic[c] for c in df_tot['case'].unique()]
cols
# %%
mean_nn.rename(rn_dic, axis=0)
# %%
figsize=[5,3]
svarl = ['NCFT_Ghan', 'SWCF_Ghan', 'LWCF_Ghan','DIR_Ghan']
mean, std, mean_nn, std_nn = get_mean_std_by_type(dic_vals, svarl, case_types=['PI', 'PIaerPD'], ctrl='PI',model_types=model_types)
mean_nn = mean_nn.rename(rn_dic, axis=0)
std_nn = std_nn.rename(rn_dic, axis=0)
fig, ax = plt.subplots(1, figsize=figsize, dpi=200)
mean_nn.plot.bar(alpha=0.5, ax=ax, color=[cdic[c] for c in pd.DataFrame(mean).columns],
yerr=std_nn) # , colors={'OsloAeroSec':'b'})
ax.axhline(0, linewidth=0.4, c='k')
sns.despine(fig, bottom=True, trim=True, offset=10)
ax.legend(frameon=False)
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
labeltop=True,
labelbottom=False
) # labels along the bottom edge are off
ax.set_ylabel('PD-PI [Wm$^{-2}$]')
fn = create_filename('forcing')
plt.tight_layout()
fig.savefig(fn+'pdf', dpi=300)
plt.show()
# %%
fn
# %%
# %%
# %%
# %%
# varl = ['N_AER']
df2, dic_vals = one_val_tab.get_tab_yearly_mean(varl,
cases_sec + cases_nsec,
startyear,
endyear,
pmin=pmin,
pressure_adjust=pressure_adjust,
average_over_lev=avg_over_lev,
groupby='time.year', # 'time',
dims=None,
area='Global',
invert_dic=True
)
# %%
from useful_scit.imps import (plt, pd, sns)
# %%
from sect_sens.data_info.simulation_types import get_abs_by_type
# %%
from sect_sens.data_info import get_nice_name_case, simulation_types
# %%
relative=False
dic_diff = simulation_types.get_diff_by_type(dic_vals, varl, case_types=['PI','PD'],
relative=relative,
mod_types=model_types,
ctrl='PI'
)['PD-PI']
ls =[]
for key in dic_diff.keys():
print(key)
_df = dic_diff[key]
_df['case'] = key
print(_df.keys())
ls.append(_df.reset_index())
df_tot = pd.concat(ls)
df_tot
# %%
trans_dic = {v:get_fancy_var_name(v) for v in varl}
# %%
rn_dic = {
'NCRE$_{Ghan}$':'ERF$_{aci}$',
'SWCRE$_{Ghan}$':'ERF$_{aci,SW}$',
'LWCRE$_{Ghan}$':'ERF$_{aci,LW}$',
'DRE$_{Ghan}$': 'ERF$_{ari}$'
}
# %%
svarl = ['NCFT_Ghan', 'SWCF_Ghan', 'LWCF_Ghan','DIR_Ghan']
df1 = df_tot[[*svarl,'case']]
df1 = df1.rename(trans_dic, axis=1)
df1 = df1.rename(rn_dic, axis=1)
df2 = pd.melt(df1,id_vars='case')
# %%
cols=[cdic[c] for c in df_tot['case'].unique()]
cols
# %%
df2.groupby(['variable','case']).mean()
# %%
import matplotlib as mpl
import numpy as np
figsize=[5,3]
figsize=[4.,4.7]
f,ax = plt.subplots(figsize=figsize,dpi=150)
pts = np.linspace(0, np.pi * 2, 24)
circ = np.c_[np.sin(pts) / 2, -np.cos(pts) / 2]
vert = np.r_[circ, circ[::-1] * .7]
open_circle = mpl.path.Path(vert)
sns.barplot(y='value',x='variable', hue='case', data=df2,errcolor='.55',ci=90
,palette=cols, alpha=.6)
g = sns.stripplot(y='value',x='variable', hue='case', data=df2,dodge=True,marker=open_circle,palette=['none']*3,jitter=.2,label='_nolegend_')#, add_=False)#label='_nolabel_')
handles, labels = ax.get_legend_handles_labels()
# When creating the legend, only use the first two elements
# to effectively remove the last two.
l = plt.legend(handles[3:], labels[3:], frameon=False, loc='center right')
ax.axhline(0, linewidth=0.4, c='k')
sns.despine(f, bottom=True, trim=True, offset=10)
#ax.legend(frameon=False)
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
labeltop=True,
labelbottom=False
) # labels along the bottom edge are off
ax.set_ylabel('[Wm$^{-2}$]')
ax.set_xlabel('')
fn = create_filename('forcing')
plt.tight_layout()
f.savefig(fn+'pdf', dpi=300)
print(fn)
plt.show()
print(fn)
|
oas_erf/notebooks/06_review/01_one_val.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Natural Langugage Processing
# ## Description
#
# NLP or Natural Language Processing as is normally referred to, refers to working (or processing) **text data**, either for machine learning, or any of the host of use cases textual data comprises of. Working with text, is very different from working with numerical or categorical data. We have worked extensively with data, numerical, categorical and boolean, however text data is a different paradigm altogether and this tutorial aims to get you acquanited with the basics of working with text and understanding the underlying implications in Machine learning.
#
# ## Overview
#
# - Introduction to the problem statement **Consumer Complaints Database**
# - What is NLP (Introduction and usecases)
# - Tokenization and Introduction to NLTK
# - Vectorization and vector space models **Count Vectorizer**
# - Applying our first classification algorithm **Logistic Regression**
# - Stopwords
# - Basic Stemming
# - TFIDF
# - Naive Bayes Classifier
# - Linear kernel SVM
# - Text Classification (Build a text classifier using NLTK)
#
#
# ## Pre-requisite
#
# - Python (along with NumPy and pandas libraries)
# - Basic statistics (knowledge of central tendancy)
#
#
# ## Learning Outcomes
#
# - Understanding why working with text data isn't like numerical or categorical data
# - What is NLP
# - The basic building blocks of text
# - Tokenization, Stemming and what constitutes as a stopword
# - Preliminary cleaning of text data
# ## Chapter 1: Introduction to text data
#
# ### Description:
# Uptill here, all of our problem statements had data in either a numerical format, a categorical format, or a Boolean format. In the real-world we usually do, and might very well encounter text data. We will now try to understand how we can use text analytics to solve data with text.
#
# ### 1.1 Introduction to the problem statement: <font color='green'> Categorize complaints into categories</font>
#
# **What is the problem?**
# #### The Dataset is a consumer complaints database where every complaint needs to be categorized into one of the pre-defined 12 categories. A multi- class classification problem.
#
# Each row in the dataset describes a single compliant. Including the complaint narrative, the issue, the category of the complaint, the date it was received on, the zip code and details of the customer placing the complaint and the current status of the complaint. The final idea is to build a model that will categorize each customer's complaint into a product (12 categories in all). You can download the dataset below.
#
# We will work with the csv file.
#
# https://catalog.data.gov/dataset/consumer-complaint-database
#
# However, for the purpose of understanding how text processing works, we will specifically, work on only 2 columns of this dataset. It is evident that if we add more features, the model accuracy will rise and be more robust, however initially, we will just have 2 columns - the consumer complaint narrative and the product the complaint has to be categorized into.
#
# - Consumer Complaint Narrative
# - Product
#
# The 2 categories the compliants need to be categorized into are:
#
# ['Mortgage', 'Student loan', 'Credit card or prepaid card', 'Credit card', 'Debt collection', 'Credit reporting', 'Credit reporting, credit repair services, or other personal consumer reports', 'Bank account or service', 'Consumer Loan', 'Money transfers', 'Vehicle loan or lease', 'Money transfer, virtual currency, or money service', 'Checking or savings account', 'Payday loan', 'Payday loan, title loan, or personal loan', 'Other financial service', 'Prepaid card']
#
# **Brief explanation of the dataset & features**
#
# * `Consumer Complaint Narrative`: Is a paragraph (or text) written by the customer explianing his complaint in detail. It is not a numerical or categorical type, the data is a string type consisting of text in the form of paragraphs
#
# * `Product`: Is the category we are to classify each complaint to.
#
# **What we want as the outcome?**
#
# Using a classification algorithm, classify each complaint to it's respective category
#
# #### Why work with text
#
# ***
#
# **Intuition for text**
#
# Let's start with what information we have: The main goal is to build a machine learning model which can predict the category of the complaint based on the customer's written data. The written data here is in the form of a paragraph comprising of sentences (or natural language). How do we convert this text data to a form fit for machine learning? The usual ways of working with numerical or categorical data will not work here, as the data type is completely different, and the algorithm has to make sense of the written data, not a single variable unlike categorical or numerical.
#
#
# If you have a look at the Consumer Complaint Narrative columns, the values are paragraphs! Not numbers or categories. This needs to be pre-processed before running an algorithm onto this.
#
# **Why NLP for this data**
#
# These complaints in the Narratives columns are typical examples of text data. Normal paragraphs, sentences in the form of text. The column Consumer complaint narrative has all rows in the form of either NaNs or text data. How do we make sense of this data?
#
# Do we convert this to categorical by one-hot encoding the text? If yes, how do we do it?
#
# How are we supposed to convert this text data to a numerical format to make sure the Machine Learning Algorithm can be applied to this?
#
# Can this column be used in a multi category classfication model to predict the class of the complaint?
# All these questions (and others) can be answered through a particular branch of ML. Enter Natural Language Processing.
#
# ### Have a look at the data set
#
# In this task you will load Consumer_complaints.csv into a dataframe using pandas and explore the column Consumer Complaint Narrative.
#
# We will see at the end of this exercise that the **The cell values of the consumer complaint narrative column is a paragraph!**
#
# This is a typical example of text data. Normal paragraphs, sentences in the form of text. The column Consumer complaint narrative has all rows in the form of either NaNs or text data. How do we make sense of this data?
# Do we convert this to categorical by one-hot encoding the text? If yes, how do we do it?
# How are we supposed to convert this text data to a numerical format to make sure the Machine Learning Algorithm can be applied to this?
# Can this column be used in a multi-nominal classfication model to predict the class of the complaint?
#
# All these questions (and others) can be answered through a particular branch of ML. **Enter Natural Language Processing.**
#
#
# ### Instructions
# - Load the csv into a dataframe
# - Drop all columns except the Product and the Consumer Complaints Narratives. Make sure to keep a copy of the original dataframe in a different instance.
# - Print out the first 5 instances of our 2 column dataframe. Name it df.
# - Rename column Consumer compliant narrative to X and Product to y (The reason we have done this is obvious. We intend to classify the complaints in the narrative (X) to each of the categories listed in the Product (or the Y column))
# - print out the first value of the X column
# - Have a look at the various categories that exist in the Product (the renamed y column, these are the categories that exist).
import os
os.chdir('C:\Learning')
import pandas as pd
df = pd.read_pickle("Consumer_complaints.pkl")
df_copy = df.head(1000).copy()
df = df_copy[["Consumer complaint narrative", "Product"]] #keeping the relevant columns
df.columns = ["X","y"]
df.head()
#Printing out the first non-empty value of the X column. Hence the second value, index is 1
print(df["X"][1])
print ("\n")
print (list(df["y"].unique()))
# ## Chapter 2: What is NLP? (Introduction and Usecases)
#
# ### 2.1 Introduction
#
# ***
#
# NLP or the Natural Language part in NLP, is called that, because it is the language that exists all around us.
# NLP can broadly be defined as the "cleaning" and "getting the text" to a form fit for machine learning. That's all it really is. Of course you can derive insights from the text as well just like the EDA operation, which aims to bring the data to a more ML application frinedly approach. There are other off shoots of Natural Langugage such as NLG - Natural language Generation which aims to generate new text data based on prior data and Natural Language Understanding - NLU which is the backbone of all intelligent chatbots out there currently, which focuses on recognizing the intent of a conversation. For the sake of this tutorial and brevity we will stick to NLP.
#
# #### Why is it difficult to work with text?
#
# ***
#
# Comprehending Language is hard for computers. Several reasons exist.
#
# - Different ways of saying the same thing, is one of the prime reasons, why computers have a hard time deciphering the meaning or intent of those statements. "I like the rains of Mumbai" and "Mumbai is beautiful during the monsoons and that's why I like it" are basically advocating the same sentiment, however since they are 2 completely different sentences syntactically, computers have a hard time figuring out the intent of the user and get stuck.
#
# - Ambiguity - "The shop is by the road" and "The shop was found to be closed by him". Are 2 completely different statements. The word "by" used in completely different meanings here. In the first case it represents proximity, in the second it refers to the person.
#
# - Context. "Anil is my friend. He likes football". In the second statement "he" refers to Anil. Computers are not inherently able to store the context of the first statement and use it to the decipher the second statement.
#
# - Understanding language. All code to a machine is just numbers. A statement in human language is just a sequence of numbers to a computer. Rudimentary Chatbots work because they detect key words in your statement. As long as the keywords remain the same, you could use any words in your statements and the end result will remain the same.
#
# - Every language has its own uniqueness. Like in the case of English we have words, sentences, paragraphs and so on to limit our language. But in Thai, there is no concept of sentences. That’s why Google Translator or any other translator struggles to perfectly convert a piece of text from one language to another.
#
# - Machines have a hard time adapting to any new constructs that humans come up with. Suppose a teenager is looking at his twitter feed and comes across a word he has never seen before, he might not understand it’s meaning instantly. But this does not mean he cannot adapt. After looking at the word in several different tweets he might be able to understand why and in which context that word is to be used. This is not possible with machines. Machines can only handle data that they have seen before. If something new comes up, they get confused and are unable to respond.
#
# #### Usecases of NLP
#
# ***
#
# The usecases of NLP encompass almost anything you can do with Language in relation to a problem.
#
# 1) Sentiment Analysis - Finding if the text opinionated a positive or negative sentiment.
#
# (Sentiment analysis is immensely useful in figuring the overall sentiment of products (Amazon), movies (Netflix), food (Zomato) by parsing the reviews and doing a sentiment analysis on them)
#
# 2) Text Classfication - categorizing text to various categories
# (Some examples of text classification are:
#
# - Understanding audience sentiment from social media,
# - Detection of spam and non-spam emails,
# - Auto tagging of customer queries, and
# - Categorization of news articles into defined topics.
# )
#
# 3) Summarizing - Summarzing a paragraph into "n" words or sentences
#
# (Example: Inshorts, news in 60 words or less)
#
# 4) Parts of Speech - Tagging - Figuring out the various nouns, adverbs, verbs etc; in your text
#
# (Chatbots)
#
# 5) Language translation
#
# (Google translate)
#
# 6) Grammar correction
#
# (Autocorrect in messaging services)
#
# 7) Entity recognition - Finding places, animals, people from the text in question
#
# (Chatbots
#
# 8) Intent recognition - Chatbots usually use this extensively. To figure what exactly you, or the customer in question needs information or services about.
#
# We will deal with each of them in detail in the subsequent tutorials.
# ## Chapter 3: Tokenization and Introduction to NLTK
#
#
# ### 3.1 Building blocks of text:Motivation for tokenization
#
# ***
# Now we can see that unlike all the machine learning datasets we have worked with previously, the data isn't boolean, numeric, cateorigical etc; How do we apply this text data to a ML algorithm?
# The first step is understanding what text data consists of..
#
# Usually a text is composed of paragraphs, paragraphs are composed of sentences, and sentences are composed of words.
#
# #### Words are the basic building blocks of any text.
#
# Sure you could do deeper into letters, but the letters as themselves have no meaning, it's only when they are combined into words, that the text starts to make sense. Hence NLP considers words as the absolute unit of text.
#
# Tokenization is exactly what it sounds like. Breaking down anything to "tokens". Tokens are the basic units of a particular dataset. In this case, our data is text and tokenization implies breaking it down into it's basic tokens. Which are words.
#
# We could also tokenize a paragrah into sentences. Since a paragraph is composed of sentences.
#
# #### Introduction to NLTK
#
# ***
# Working with text data as we have seen is not as straightforward as working with numeric or other data types, hence it is no surprise, that this processing is achieved through special libraries. NLTK or the Natural Langugage Tool Kit is the de-facto standard library in python which specifically deals with text. Tasks, such as tokenization, Lemmatization, text_classification, vectorizing are methods built in and help working with text much simpler. Do not worry if the above terms do not make sense to you, we will get to them and cover them in detail eventually.
#
# NLTK is not part of the standard Anaconda 3.6 installation and will require an independent set-up.
#
# #### Installing NLTK.
#
# ***
# To install NLTK and all it's dependencies, go to your terminal(mac) or command window(windows) and type pip install nltk. This could take a while, depending on your internet speed. It is to be noted that NLTK is not the only library that can be used for text processing, there are a ton of others, however NLTK is one of the first ones that came up and is generally astarting point from a beginner's perspective. There are obvious tasks the NLTK library cannot do, which have to be compensated via other libraries, howeer for thr purpose of this particukar tutorial, NLTK should more than suffice presently.
#
# #### Tokenizing with NLTK - The problem intuition
#
# ***
#
# Now that we have already defined our present dataframe df as below, we will need to find a way to convert the text in the X column to numbers to get them to a form where I would be able to apply an alogorithm to this. Think of this like sklearn, which require all non-numeric data to be encoded (label or one-hot) prior to the sklearn pipeline.
#
# Intuitively, it would make sense to divide each paragraph of text to it's basic form (words) and then convert each of those words to numbers. We could assign a particular number to each word, in which case a sentence could look like a set of numbers to us, each number representing a particular word.
#
# The first step to acheiving that would be to break the text down to words. That's what tokenization aims to do. NLTK has a built in for tokenization. Assuming NLTK is installed on your machines now, let us quickly run through a few sample tokenization exercises, before your assignment 3 where you will break doen every cel in the Y column to its words and create a new colunm for the same.
# ### Tokenize the first complaint into words
#
# In this task you will assign a variable to the first row of the consumer complaints narrative column (X column) and break down the text into it's constituent words.
# ### Instructions
# - Load the dataframe defined earlier
# - Assign a variable **first_complaint** to the paragraph listed in the first non-empty row of the consumer complaint narrative column (or X)
# - Break it down into words using the split command initially and then using the nltk.word_tokenize function.
# - Assign this list of words to another list called bag_of_words
# - We can see from both lists (the one using split and the one using word_tokenize) that the word_tokenize function is more robust as it splits the paragraph into purely words and seperates the punctuation into seperate tokens. In the case of split, full-stops have appeared along with certain words.
# +
import nltk
from nltk.tokenize import word_tokenize
nltk.download('punkt')
df["X"].head(3)
df = df.dropna() #dropping nans
first_complaint = df["X"].iloc[0]
print (first_complaint)
print ("\n")
#Using the split command
print ("Using the Split Command")
print ("\n")
bag_of_words = first_complaint.split(" ")
print (bag_of_words)
bag_of_words = word_tokenize(first_complaint)
print ("\n")
print (bag_of_words)
# -
# ### 3.2 Sent Tokenize
#
# ### Tokenize the first complaint into sentences
#
# One could also tokenize a paragraph into constituent sentences.
#
# ### Assignment 2
#
# #### Tokenize the Second non-empty complaint into words and convert all words to lower case and assign the list of words to a list
#
# #### Description:
# The importance of converting words to lower case - All words should be converted to lowercase while doing NLP. The reason behind being, that "Mumbai" and "mumbai" even though are the same word, will be considered 2 seperate words whilst converting the words into numbers, and that would be a biased conversion. To avoid such problems, it is standard practice to convert all words or text to lower case, before beginning NLP.
#
# ### Instructions
# - Load the dataframe defined earlier
# - Assign a variable **first_complaint** to the paragraph listed in the first non-empty row of the consumer complaint narrative column (or X)
# - Break it down into sentences using the sent_tokenize function from nltk
# - Assign this list of words to another list called list_of_sentences
# +
first_complaint #Is already loaded onto the workspace
from nltk.tokenize import sent_tokenize
list_of_sentences = sent_tokenize(first_complaint)
(list_of_sentences)
print ("\n")
print (df["X"].iloc[1])
bag_of_words_lower = word_tokenize(df["X"].iloc[1].lower())
print ("\n")
print (bag_of_words_lower)
# -
# ## Chapter 4: Vectorization
#
#
# ### 4.1 Converting your text to numbers: The crux of NLP
#
# ***
# We managed to convert our complaints into a bag of words or a list of words. But that is no good until we figure a way out to convert these words to a numeric format. And that is necessary to apply any sort of algorithm (machine learning or otherwise).
#
# **This process of converting text data to numbers is called vectorization**
#
# There are multiple methods to convert words to numbers. We will be initially dealing with 1 of them: A count Vectorizer.
#
# ### 4.2 The intuition behind vectorization: The count vectorizer
#
# ***
# Every list of words corresponds to a row in our dataframe, something like this. Where the X column would now be the list of lowercased words and our y column would be the product category.
#
# <img src="img.png">
# The idea now is to convert the X column to numbers.
#
# **One way to do that would be to represent every word as a key value pair in the form of a dictionary, where the key would be the word and the vale would be the number of times that word has appeared in the list.**
#
# This method of converting the counts of words in the list to convert them to a numeric format is called Count vectorization. We will initially do this manually, and then explot sklearn to do this automatically to understand the intuition behind this.
#
#
# ### Convert the first complaint to numbers using the counts of words in the form of a dictionary
#
# In this task you will implement your own code for count vectorization
# ### Instructions
# - Load the df.
# - Take the first complaint in the X column and assign that to a list called **first_complaint**
# - Tokenize the list to its words and convert them to lower case.
# - Create a dictionary (any method you prefer) so that the keys are the words themselves and the values are the number of times the word has appeared in the list **first_complaint**
# - Name this dictionary as Count_Vectorizer
#
first_complaint = word_tokenize(df["X"].iloc[0].lower())
from collections import Counter
Count_Vectorizer = {}
Count_Vectorizer = Counter(first_complaint)
print (Count_Vectorizer)
# ### 4.3 Introduction to the sklearn's Count Vectorizer
#
# ***
# We did manage to convert our list of words to numbers. However the problem still remain unresolved.
#
# **How do we apply our algorithm to this?**
#
# **Could we convert every word to a feature (or column) and the count associated with it to it's value and then apply a Classification algorith to it? Something like below?
#
# <img src="img2.png">
#
# ** This looks very similar to one-hot encoding and is a typical method of applying ML to text data**.
#
# This is simiar to one-hot in the way that when we add the second row, and the the third row and subsequent rows, the features or the columns will increase as more and more words come in and there will be words which do not appear in say the first_complaint, the vectorizer will automatically assign 0 to those words. **Hence the number of features will be equal to the total number of unique words in all the complaints combined and the values for those features will be the count of those words in that particular complaint.**
#
# A normal classifcation algorithm can now be applied where X is all the features except the Product column and y is the Product column.
#
# We could just use the sklearn's Count Vectorizer and convert all the text into numbers in a single step instead of breaking them down into individual words, lowercasing them, and then making a dictionary assigning the counts.
# This is how we would do it for the first row.
df.head(5)
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer()
txt = [df["X"].iloc[0]]
print (txt)
print ("\n")
print ("Applying the count vectorizer")
cv.fit(txt)
vector = cv.transform(txt)
print ("Vector Shape")
print ("\n")
print (vector.shape)#Has 69 unique words
vector_values = vector.toarray()
print ("\n")
print ("vector Values")
print (vector_values)
print ('These are the counts of the 69 unique words in our first complaint. To find which words have these counts, we can execute the command below:cv.vocabulary_')
print ("\n")
print ("Count Vectorizer Vocabulary")
print (cv.vocabulary_)
# #### This does not specify the counts of the word. This specifies the index of the word in the vector_values list. So for the first word expalin, we see it's index is 22, if we see the index 22 in the vector_values list, we will see that it's value is 1,as specified in the figure above.
vector_values = vector_values.tolist()
vector_values = vector_values[0]
print (vector_values)
print ("\n")
print ("count value of the word at index 22")
print (vector_values[22]) #Value of 1
print ("\n")
print ("count value of the word at index 34, the word is 'loan'")
print (vector_values[34]) #Value of 2
print ("\n")
print ("Seeing the cv.vocabulary_ dictionary we see that the word is 'loan' and it's value in the vector values list is 2. ")
# <img src="img2.png">
# ### Coding the count vectorizer and getting the data to a form for algorithm application.
#
# #### Use the count vectorizer to numerize the X column of the dataframe and make a new dataframe with these features and the product column. We will consider just the top 3 rows of the entire dataframe to aid better understanding and then run it over the entire dataframe.
#
#Importing count vectorizer from sklearn
from sklearn.feature_extraction.text import CountVectorizer
#Initializing the Count vectorizer
cv = CountVectorizer()
#Initializing a dataframe "all text" with the first 3 rows of df
all_text = df["X"][:3]
all_text = pd.DataFrame(all_text)
#Renaming the column for that dataframe (has only one column) to "text"
all_text.columns = ["Text"]
#Converting to lower case
all_text["Text"] = all_text['Text'].str.lower()
all_text
#Fitting the Count vectorizer all text
cv.fit(all_text["Text"])
vector = cv.transform(all_text["Text"])
vector_values_array = vector.toarray()
#Converting the text to numbers - The transform function does this.
vector_values_list = vector_values_array.tolist()
print (len(vector_values_list))
print ("\n")
#Because there are 3 rows in the entire dataframe.
print (len(vector_values_list[0]))
# 214 is the number of unique words in all 3 rows combined. This value will be constant for every list element in the vector_values_list because all the unique words in the entire dataframe have been converted to features and the values for these features per row depends on the count of those words in that row, 0 in case the word does not exist in the row.
len(vector_values_list[1])
# +
len(vector_values_list[2])
#As you can see that every document has been converted to a fixed length vector of 214 words, and
#have values coresponding to the occurences of those words in the particular document:0 in case those words
# words aren't present in the document
# -
# The y values for these 3 rows are:
# We will have to label encode these categories to numerize them.
from sklearn.preprocessing import LabelEncoder
labels = pd.DataFrame(df["y"][:3])
#label encoding the y values
labels.columns = ["labels"]
le = LabelEncoder()
labels["labels"] = le.fit_transform(labels["labels"])
print (labels)
# Now our final dataframe for the X is the vector_values_array, which is the vectorized form of the text, and the Y is the labels dataframe.
vector_values_array[:1][:5] #This is the row 1 of the original dataframe df now numerized
# If we now, include all the 335 rows and vectorize them and label it as the X, and the corresponding y labels, we can train a classification algorithm on the same, and figure out the accuracy.
#Running the exact same code as the earlier one on 335 rows
import numpy as np
all_text = df["X"]
all_text = pd.DataFrame(all_text)
all_text.columns = ["Text"]
all_text["Text"] = all_text['Text'].str.lower()
cv = CountVectorizer()
cv.fit(all_text["Text"])
vector = cv.transform(all_text["Text"])
vector_values_array = vector.toarray()
labels = pd.DataFrame(df["y"])
labels.columns = ["labels"]
labels["labels"] = le.fit_transform(labels["labels"])
len(vector_values_array) #Because 335 documents in all
len(labels) #Because each document or complaint has a label, hence 335 labels in all
# Applying a normal Logistic Regression function to this X and y post breaking it down to a train and a test set, we can calculate the accuracy of the same.
from sklearn.metrics import accuracy_score,roc_auc_score
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split as tts
X = vector_values_array
y = labels["labels"]
X_train,X_test,y_train,y_test = tts(X,y,test_size=0.4,random_state=42)
log_reg = LogisticRegression(random_state=42)
log_reg.fit(X_train,y_train)
y_pred = log_reg.predict(X_test)
print (accuracy_score(y_test,y_pred))
# That's a 42% accuracy of predicting the product category. Which isn't really a good number. A look at the precision recall shows that the product category was heavily imblanaced, hence a few categories have not been detected by the algorithm. Before we rectify the same, let us first look at the distribtion of the y column.
labels["labels"].value_counts()
# We can see very clearly that the data set clearly has heavily imbalanced labels. Labels 10,8,12 are under represented and hence the probability of the model catching those labels is less, hence the accuracy is going to suffer. A look at the classification report should reinfornce this.
from sklearn.metrics import classification_report
import warnings
warnings.filterwarnings("ignore")
print (classification_report(y_test,y_pred))
# As predicted, the recall for Labels, 8,10,11,12,2,0 and 3 are 0. A typical sampling issue here. This can be rectified by an oversampling technique such as SMOTE or ROS, however before we get to that, it's time to understand what the other reasons for a lower accuracy could be.
# It is important to realize that count vectorizer is essentially a Ranking algorithm, in the way that it gives a higher weight to words which have appeared more number of times, in other words, the value of the key_wvalue pair is plain the count of the word in the dataset. It does not assign any importance to the order in whoch the words are sentences have appeared.
#
# Another point to be considered is the fact that words like "a","an","the"... will appear more number of times than the rest of the words as they are common articles. Using a Count vectorizer out of the box on a paragraph or a body of text will invariably give the highest count to these common words. Hence the words we are actually interested in will be underneath these words. one way to rectify this, is to remove these commonly occurng words. NLTK offers this functionality and has rightly defined these particluar words as "stopwords" or words we wouldn't include in out bag of words. We will have to remove the punctutaion as well, as you can see our initial bag of words had the commas, full-stops and all of the other symbols as well. We will want to remove them as well, to avoid being vectorized.
# To see the list of stopwords, NLTK currently includes, we could check that by just running
import nltk
nltk.download("stopwords")
from nltk.corpus import stopwords
print (set(stopwords.words('english')))
# The punctutaion list can be derived as follows.
from string import punctuation
print (list(punctuation))
# +
#The good thing is we can add our own list of words we want to remove our body of text over and above #this list.
custom_set_of_stopwords = set(stopwords.words('english')+list(punctuation)+["Bangalore"])
#This will include the word Bangalore as a topword and remoe the same from our body of text before vectorizing it.
print ("Bangalore" in custom_set_of_stopwords)
# -
# If we re-do the exercise for prediction removing the stopwords, the accuracy should increase, as now we are not giving any weight to meaningless words but to words which actually matter.
all_text = df["X"]
all_text = pd.DataFrame(all_text)
all_text.columns = ["Text"]
all_text["Text"] = all_text['Text'].str.lower()
# Taking just the first row
first_complaint = all_text['Text'].iloc[0]
first_complaint
first_complaint_bow = word_tokenize(first_complaint)
print (first_complaint_bow)
len(first_complaint_bow)
first_complaint_bow_stopwords_removed = [x for x in first_complaint_bow if x not in custom_set_of_stopwords]
print (first_complaint_bow_stopwords_removed)
len(first_complaint_bow_stopwords_removed)
# We see that the less important words such as "a","an","the","where" have been removed including the punctutaion.
# Redoing this for all the rows of the dataframe.
all_text = df["X"]
all_text = pd.DataFrame(all_text)
all_text.columns = ["Text"]
all_text["Text"] = all_text['Text'].str.lower()
cv = CountVectorizer(stop_words="english")
cv.fit(all_text["Text"])
vector = cv.transform(all_text["Text"])
vector_values_array = vector.toarray()
labels = pd.DataFrame(df["y"])
labels.columns = ["labels"]
labels["labels"] = le.fit_transform(labels["labels"])
from sklearn.metrics import accuracy_score,roc_auc_score
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split as tts
X = vector_values_array
y = labels["labels"]
X_train,X_test,y_train,y_test = tts(X,y,test_size=0.4,random_state=42)
log_reg = LogisticRegression(random_state=42)
log_reg.fit(X_train,y_train)
y_pred = log_reg.predict(X_test)
print (accuracy_score(y_test,y_pred))
# We see a massive 5% increase in accuracy by just removing the stopwords. The model is now being trained on the words which actually matter. Not the more commonly occuring stopwords.
# ### 4.4 Introduction to the TF-IDF Vectorizer
#
# In the last tutorial we saw how text was converted to numerics using a count vectorizer.
#
# In other words, a count vectorizer, counts the occurences of the words in a document and all the documents are considered independent of each other. Very similar to a one hot encoding or pandas getdummies function. However in cases where multiple documents are involved, count vectorizer still does not assume any interdependence between the documents and considers each of the documents as a seperate entity.
#
# It does not rank the words based on their importance in the document, but just based on whether they exist or not. This is not a wrong approach, but it intuitively makes more sense to rank words based on their importance in the document right? In fact, the process of converting, text to numbers should essentially be a ranking system of the words so that the documents can each get a score based on what words they contain. All words cannot have the same imprtance or relevance in the document right?
#
# #### Enter TF-IDF!!
#
# TF-IDF or Term Frequency and Inverse Document Frequency is kind of the holy grail of ranking metrics to convert text to numbers. Consider the count vectorizer as a metric which just counts the occurences of words in a document.
#
# ** The ranking system in a count vectorizer is purely occurence based on a single document only!**
#
# TF-IDF takes it a step further and ranks the words based not just on their occurences in one document but across all the documents. Hence if CV or Count vectorizer was giving more importance to words because they have appeared multiple times in the document, TF-IDF will rank them high if they have appeared only in that document, meaning that they are rare, hence higher importance and lower if they have appeared in all or most documents, because they are more common, hence lower ranking.
#
# Consider a scenario where there are 5 documents and all are talking aout football. The word football would have appeared multiple times in each document. CV is going to rank football consistently high and infact give the word football a different value across all 5 documents based on how many times that word has appeared in that document. In other words, it is assuming, that the more number of times a word appears, the more important it is. That is exactly what the TF or the Term Frequency component in TF-IDF does.
#
# IDF on the other hand now is the dominating factor in TFIDF which is going to find out the number of times football has also appeared in the other 4 documents except for the one it is currently seeing. If football has also appeared in rest of the documents, it means that though football is important to that one document based on the number of occurences, considering it has appeared in the rest as well, it is not that rare or more common, hence the importance now is going to reduce instead of going high!
#
# **The ranking system is across the entire corpus or all documents. It is not a single document based metric!**
#
# We have seen how CV is calculated for a word in a document. Let us now see how TF IDF is...
#
# The tf-idf weight is composed by two terms: the first computes the normalized Term Frequency (TF), aka. the number of times a word appears in a document, divided by the total number of words in that document; the second term is the Inverse Document Frequency (IDF), computed as the logarithm of the number of the documents in the corpus divided by the number of documents where the specific term appears.
#
# TF: Term Frequency, which measures how frequently a term occurs in a document. Since every document is different in length, it is possible that a term would appear much more times in long documents than shorter ones. Thus, the term frequency is often divided by the document length (aka. the total number of terms in the document) as a way of normalization:
#
# TF(t) = (Number of times term t appears in a document) / (Total number of terms in the document).
#
# IDF: Inverse Document Frequency, which measures how important a term is. While computing TF, all terms are considered equally important. However it is known that certain terms, such as "is", "of", and "that", may appear a lot of times but have little importance. Thus we need to weigh down the frequent terms while scale up the rare ones, by computing the following:
#
# IDF(t) = log_e(Total number of documents / Number of documents with term t in it).
#
# #### Example
#
# Consider a document containing 100 words wherein the word cat appears 3 times. The term frequency (i.e., tf) for cat is then (3 / 100) = 0.03. Now, assume we have 10 million documents and the word cat appears in one thousand of these. Then, the inverse document frequency (i.e., idf) is calculated as log(10,000,000 / 1,000) = 4. Thus, the Tf-idf weight is the product of these quantities: 0.03 * 4 = 0.12.
#
# Let us now take the first 2 complaints and run a TF-IDF vectorizer on the same. So, in this case, the 2 complaints are our 2 documents. and instead of a CV which considers each document independent of each other and just calculates the count of every word in the document. now the corpus will be the sum total of both documents.
complaint_1 = df["X"].iloc[0]
complaint_2 = df["X"].iloc[1]
complaint_3 = df["X"].iloc[2]
print ("Complaint 1: ", complaint_1)
print ("\n")
print ("Complaint 2: ", complaint_2)
print ("\n")
print ("Complaint 3: ", complaint_3)
from sklearn.feature_extraction.text import TfidfVectorizer
# list of text documents called sents
sents = [complaint_1, complaint_2, complaint_3]
# create the transform
vectorizer = TfidfVectorizer()
# tokenize and build vocab
sents
vectorizer.fit(sents)
vector = vectorizer.transform(sents)
vector
vector.shape #214 Unique words in both sentences combined. 3 documents in total.
vector_values = vector.toarray().tolist()[0]
# vector_values would be the tf-idf score for each of the 214 words. Printing the first 5 elements of this list.
vector_values[:5]
# To figure out which these words are, just like the count vectorizer, we have the vectorizer.vocabulary_
print (vectorizer.vocabulary_)
import operator
sorted_x = sorted(vectorizer.vocabulary_.items(), key=operator.itemgetter(1))
words = [x[0] for x in sorted_x]
d = dict(zip(words,vector_values))
print (d)
# Sorting this dictionary by value in the descending order to see the ranking.
print (sorted(d.items(), key=operator.itemgetter(1), reverse = True))
# We can see that the model learns to give lesser importance to words like is,it,in etc;. Unfortunately, it also gives a low importance to important words like financial, mortgage and a fairly high importance to unwanted words like the, was. It does give higher importance to words such as credit. And that is because TF-DF works better with larger corpuses. Just like a machine learning model, the larger the data, the better the model. With a larger corpus, these issues would be resolved when a lot more documents would have words like financial but not the.
#
# Rerunning this fo about 100 documents, we see that the ranking is completely different.
sents=[]
for x in range(100):
sents.append(df["X"].iloc[x])
len(sents)
from sklearn.feature_extraction.text import TfidfVectorizer
# list of text documents called sents
# create the transform
vectorizer = TfidfVectorizer()
# tokenize and build vocab
vectorizer.fit(sents)
vector = vectorizer.transform(sents)
vector.shape
vector_values = vector.toarray().tolist()[0]
import operator
sorted_x = sorted(vectorizer.vocabulary_.items(), key=operator.itemgetter(1))
words = [x[0] for x in sorted_x]
d = dict(zip(words,vector_values))
print ((sorted(d.items(), key=operator.itemgetter(1), reverse = True))[:20])
# "the" has moved down from 0.42 to 0.24. 'Navient' has increased from 0.20 to 0.35. So has 'bureaus' from 0.13 to 0.19. As we include more and more sentences, the words whch have appeared more and more frequently across all the documents, such as "the" are moving down in value, and words like bureau and navient, which have appeared far lesser number of times have started increasing. Which reiterates the point we had. TF-DF works better with larger corpuses. Just like a machine learning model, the larger the data, the better the model.
#
# Let's run our initial Logistic Regression model using a tf-idf and see if there is a difference in the accuracies.
all_text = df["X"]
all_text = pd.DataFrame(all_text)
all_text.columns = ["Text"]
all_text["Text"] = all_text['Text'].str.lower()
tfidf = TfidfVectorizer(stop_words="english")
tfidf.fit(all_text["Text"])
vector = tfidf.transform(all_text["Text"])
vector_values_array = vector.toarray()
labels = pd.DataFrame(df["y"])
labels.columns = ["labels"]
labels["labels"] = le.fit_transform(labels["labels"])
from sklearn.metrics import accuracy_score,roc_auc_score
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split as tts
X = vector_values_array
y = labels["labels"]
X_train,X_test,y_train,y_test = tts(X,y,test_size=0.4,random_state=42)
log_reg = LogisticRegression(random_state=42)
log_reg.fit(X_train,y_train)
y_pred = log_reg.predict(X_test)
print (accuracy_score(y_test,y_pred))
# And we can see that the overall accuracy of the model is **low** compared to the initial CV model. A 10% reduction. As far we have seen, do you think this will increase if we add more data to it.
import pandas as pd
df = pd.read_pickle("Consumer_complaints.pkl")
print ("reading")
df_copy = df.tail(2000).copy()
df = df_copy[["Consumer complaint narrative", "Product"]] #keeping the relevant columns
df.shape
df.columns = ["X","y"]
df = df.dropna()
df = df.iloc[:2000]
print (df.shape)
print ("Building model")
all_text = df["X"]
all_text = pd.DataFrame(all_text)
all_text.columns = ["Text"]
all_text["Text"] = all_text['Text'].str.lower()
tfidf = TfidfVectorizer(stop_words="english")
tfidf.fit(all_text["Text"])
vector = tfidf.transform(all_text["Text"])
vector_values_array = vector.toarray()
labels = pd.DataFrame(df["y"])
labels.columns = ["labels"]
labels["labels"] = le.fit_transform(labels["labels"])
from sklearn.metrics import accuracy_score,roc_auc_score
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split as tts
X = vector_values_array
y = labels["labels"]
X_train,X_test,y_train,y_test = tts(X,y,test_size=0.3,random_state=42)
log_reg = LogisticRegression(random_state=42)
log_reg.fit(X_train,y_train)
y_pred = log_reg.predict(X_test)
print (accuracy_score(y_test,y_pred))
# From 43% at 359 rows, to 59% at 2000 rows, we are starting to see how TF-IDF works better with larger
# data sets. The last value for the word "the" was around 0.27. Just out of curiosity, let's see the value of the word "the" now at 2000 docs
### from sklearn.feature_extraction.text import TfidfVectorizer
# list of text documents called sents
# create the transform
sents=[]
for x in range(df.shape[0]):
sents.append(df["X"].iloc[x])
vectorizer = TfidfVectorizer()
# tokenize and build vocab
vectorizer.fit(sents)
vector = vectorizer.transform(sents)
vector.shape
vector_values = vector.toarray().tolist()[0]
import operator
sorted_x = sorted(vectorizer.vocabulary_.items(), key=operator.itemgetter(1))
words = [x[0] for x in sorted_x]
d = dict(zip(words,vector_values))
print ((sorted(d.items(), key=operator.itemgetter(1), reverse = True))[:20])
# "the" is at 0.21 at 2000 documents. "And" has been pushed down, expalin has come up, deliquent has increased etc; a few words have been pushed down as well, based on their importance **across all documents** not just each document alone.
# ## Chapter 5: The Naive Bayes Classifier
# Naive Bayes classifers are based on the Bayes' theorem. A pure classification algorithm, it predicts the various categories or classes of the target based on the fundamental premise that the features responsible are independent of each other. These features are assumed to independently contribute to the the probability to the target variable belonging to a certain class.
#
# Let us try running a Naive Bayes classifier using the TF-IDF method, exactly the same way we ran a log-reg model in the earlier section.
import pandas as pd
df = pd.read_pickle("Consumer_complaints.pkl")
print ("reading")
df_copy = df.copy()
df = df[["Consumer complaint narrative", "Product"]] #keeping the relevant columns
df.shape
df.columns = ["X","y"]
df = df.dropna()
df = df.iloc[:2000]
print (df.shape)
print ("Building model")
all_text = df["X"]
all_text = pd.DataFrame(all_text)
all_text.columns = ["Text"]
all_text["Text"] = all_text['Text'].str.lower()
tfidf = TfidfVectorizer(stop_words="english")
tfidf.fit(all_text["Text"])
vector = tfidf.transform(all_text["Text"])
vector_values_array = vector.toarray()
labels = pd.DataFrame(df["y"])
labels.columns = ["labels"]
labels["labels"] = le.fit_transform(labels["labels"])
from sklearn.metrics import accuracy_score,roc_auc_score
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split as tts
X = vector_values_array
y = labels["labels"]
X_train,X_test,y_train,y_test = tts(X,y,test_size=0.3,random_state=42)
# log_reg = LogisticRegression(random_state=42)
nb = MultinomialNB()
nb.fit(X_train,y_train)
y_pred = nb.predict(X_test)
print (accuracy_score(y_test,y_pred))
# One of the fundamental reasons the model isn't giving an accuracy can be deduced from the classification report, where we see that the recall of multiple categories is 0 and that means that the data isn't balanced well enough to reflect an equal weightage. Since a classfication algorithm, tends to predict the majority class unless, the output categories are more or less equally balanced, the error in preeicting the majority class will increase as it classfies more and more data, and classifies tg=hem worng, hence reducing the overall accuracy.
#
# A recap on the Classification report of the NB classifier.
import pandas as pd
df = pd.read_pickle("Consumer_complaints.pkl")
print ("reading")
df_copy = df.copy()
df = df[["Consumer complaint narrative", "Product"]] #keeping the relevant columns
df.shape
df.columns = ["X","y"]
df = df.dropna()
df = df.iloc[:2000]
print (df.shape)
print ("Building model")
all_text = df["X"]
all_text = pd.DataFrame(all_text)
all_text.columns = ["Text"]
all_text["Text"] = all_text['Text'].str.lower()
tfidf = TfidfVectorizer(stop_words="english")
tfidf.fit(all_text["Text"])
vector = tfidf.transform(all_text["Text"])
vector_values_array = vector.toarray()
labels = pd.DataFrame(df["y"])
labels.columns = ["labels"]
labels["labels"] = le.fit_transform(labels["labels"])
from sklearn.metrics import accuracy_score,roc_auc_score
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split as tts
X = vector_values_array
y = labels["labels"]
X_train,X_test,y_train,y_test = tts(X,y,test_size=0.3,random_state=42)
# log_reg = LogisticRegression(random_state=42)
nb = MultinomialNB()
nb.fit(X_train,y_train)
y_pred = nb.predict(X_test)
print (accuracy_score(y_test,y_pred))
print (classification_report(y_test,y_pred))
y.value_counts()
# ### As predicted, the dataset is heavily imbalanced. With Category 6 and 7 being over represented , while all else have a less than 10% weightage.
# We might have to oversample the under-represented categories. We will use the Random Over sampler to do this from the package IMBLEARN
df.columns = ["X","y"]
df = df.dropna()
df = df.iloc[:2000]
# !pip install imblearn
X = df["X"]
y = df["y"]
X = pd.DataFrame(X)
tfidf.fit(X["X"])
vector = tfidf.transform(X["X"])
vector_values = vector.toarray()
vector_values = pd.DataFrame(vector_values)
labels = pd.DataFrame(y)
labels.columns = ["labels"]
labels["labels"] = le.fit_transform(labels["labels"])
y = labels["labels"]
from imblearn.over_sampling import RandomOverSampler
ros = RandomOverSampler()
X_ros, y_ros = ros.fit_sample(vector_values, y)
y_ros = pd.Series(y_ros)
X_train,X_test,y_train,y_test = tts(X_ros,y_ros,test_size = 0.3, random_state = 0)
nb = MultinomialNB()
nb.fit(X_train,y_train)
y_pred = nb.predict(X_test)
accuracy_score(y_test,y_pred)
# #### We managed a 94% accuracy with multinominal Naive Bayes, TF-IDF and oversampling.
# Using a Linear SVC as our last algorithm, to test the results
from sklearn.svm import SVC
X_train,X_test,y_train,y_test = tts(X_ros,y_ros,test_size = 0.3, random_state = 0)
svc = SVC(kernel="linear")
svc.fit(X_train,y_train)
y_pred = svc.predict(X_test)
accuracy_score(y_test,y_pred)
|
Introduction to NLP - Block - 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="4bkNVc42_H9r"
# # 3D segmentation with [MONAI](https://github.com/Project-MONAI/MONAI) and [Catalyst](https://github.com/catalyst-team/catalyst)
#
# This tutorial demonstrates how [MONAI](https://github.com/Project-MONAI/MONAI) can be used with the [Catalyst](https://github.com/catalyst-team/catalyst) framework for 3D segmentation task.
# And easily use below features:
#
# * Prepare synthetic data.
# * Load Nifti image with metadata.
# * Transforms for dictionary format data.
# * Add channel dim to the data if no channel dimension.
# * Scale medical image intensity with expected range.
# * Crop out a batch of balanced images based on positive / negative label ratio.
# * 3D UNet model, Dice loss function, Mean Dice metric for 3D segmentation task.
# * Sliding window inference method.
# * Deterministic training for reproducibility.
#
# This tutorial is based on [unet_training_dict.py](https://github.com/Project-MONAI/tutorials/blob/master/3d_segmentation/torch/unet_training_dict.py) and [spleen_segmentation_3d.ipynb](https://github.com/Project-MONAI/tutorials/blob/master/3d_segmentation/spleen_segmentation_3d.ipynb).
#
# [](https://colab.research.google.com/github/Project-MONAI/tutorials/blob/master/3d_segmentation/unet_segmentation_3d_catalyst.ipynb)
# + [markdown] colab_type="text" id="hGYGdy1yBLr5"
# ## Setup environment
# + tags=[]
# %pip install -q "monai[nibabel, tensorboard]"
# + tags=[]
# %pip install -q matplotlib
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="el520dTNfmZm" outputId="6668fb1a-3c73-4cc3-97b6-8d8260b33433" tags=[]
# %pip install -q catalyst==20.07
# -
# ## Setup imports
# + tags=[]
# Copyright 2020 MONAI Consortium
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import glob
import logging
import os
import shutil
import sys
import tempfile
import catalyst.dl
import matplotlib.pyplot as plt
import nibabel as nib
import numpy as np
import torch
from monai.config import print_config
from monai.data import Dataset, create_test_image_3d, list_data_collate
from monai.inferers import sliding_window_inference
from monai.losses import DiceLoss
from monai.metrics import DiceMetric
from monai.networks.nets import UNet
from monai.transforms import (
AsChannelFirstd,
Compose,
LoadNiftid,
RandCropByPosNegLabeld,
RandRotate90d,
ScaleIntensityd,
ToTensord,
)
from monai.utils import first
print_config()
# -
# ## Setup data directory
#
# You can specify a directory with the `MONAI_DATA_DIRECTORY` environment variable.
# This allows you to save results and reuse downloads.
# If not specified a temporary directory will be used.
# + tags=[]
directory = os.environ.get("MONAI_DATA_DIRECTORY")
root_dir = tempfile.mkdtemp() if directory is None else directory
print(root_dir)
# -
# ## Setup logging
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="r1P0mVvymvsY" outputId="930a36c6-bbb7-49a6-9520-8335df0b9164"
logging.basicConfig(stream=sys.stdout, level=logging.INFO)
# + [markdown] colab_type="text" id="XJpDdAo1-ZF4"
# # [MONAI](https://github.com/Project-MONAI/MONAI) components
# + [markdown] colab_type="text" id="G2Wo7P7EBRdQ"
# ## Prepare synthetic data
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="eq3RgOT2BclX" outputId="3a5039ac-bb44-443a-bdc3-38fca80cc6eb"
for i in range(40):
im, seg = create_test_image_3d(128, 128, 128, num_seg_classes=1, channel_dim=-1)
n = nib.Nifti1Image(im, np.eye(4))
nib.save(n, os.path.join(root_dir, f"img{i}.nii.gz"))
n = nib.Nifti1Image(seg, np.eye(4))
nib.save(n, os.path.join(root_dir, f"seg{i}.nii.gz"))
images = sorted(glob.glob(os.path.join(root_dir, "img*.nii.gz")))
segs = sorted(glob.glob(os.path.join(root_dir, "seg*.nii.gz")))
# + [markdown] colab_type="text" id="WX6w-86XBjeh"
# ## Prepare transforms and datasets
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="CjeN90W-feaM" outputId="2493d49b-8457-4139-ae7a-3c5fa5bd9087" tags=[]
train_files = [{"img": img, "seg": seg} for img, seg in zip(images[:20], segs[:20])]
val_files = [{"img": img, "seg": seg} for img, seg in zip(images[-20:], segs[-20:])]
# define transforms for image and segmentation
train_transforms = Compose(
[
LoadNiftid(keys=["img", "seg"]),
AsChannelFirstd(keys=["img", "seg"], channel_dim=-1),
ScaleIntensityd(keys=["img", "seg"]),
RandCropByPosNegLabeld(
keys=["img", "seg"],
label_key="seg",
spatial_size=[96, 96, 96],
pos=1,
neg=1,
num_samples=4,
),
RandRotate90d(keys=["img", "seg"], prob=0.5, spatial_axes=[0, 2]),
ToTensord(keys=["img", "seg"]),
]
)
val_transforms = Compose(
[
LoadNiftid(keys=["img", "seg"]),
AsChannelFirstd(keys=["img", "seg"], channel_dim=-1),
ScaleIntensityd(keys=["img", "seg"]),
ToTensord(keys=["img", "seg"]),
]
)
# define dataset, data loader
check_ds = Dataset(data=train_files, transform=train_transforms)
# use batch_size=2 to load images and use RandCropByPosNegLabeld to generate 2 x 4 images for network training
check_loader = torch.utils.data.DataLoader(
check_ds, batch_size=2, num_workers=4, collate_fn=list_data_collate
)
check_data = first(check_loader)
print(check_data["img"].shape, check_data["seg"].shape)
# create a training data loader
train_ds = Dataset(data=train_files, transform=train_transforms)
# use batch_size=2 to load images and use RandCropByPosNegLabeld to generate 2 x 4 images for network training
train_loader = torch.utils.data.DataLoader(
train_ds,
batch_size=2,
shuffle=True,
num_workers=4,
collate_fn=list_data_collate,
pin_memory=torch.cuda.is_available(),
)
# create a validation data loader
val_ds = Dataset(data=val_files, transform=val_transforms)
val_loader = torch.utils.data.DataLoader(
val_ds, batch_size=1, num_workers=4, collate_fn=list_data_collate
)
# + [markdown] colab_type="text" id="BB8EGm5OBuIR"
# ## Prepare model, optimizer and metrics
# + colab={} colab_type="code" id="gIh_W821Bvdd"
# create UNet, DiceLoss and Adam optimizer
# device = torch.device("cuda:0") # you don't need device, because Catalyst uses autoscaling
model = UNet(
dimensions=3,
in_channels=1,
out_channels=1,
channels=(16, 32, 64, 128, 256),
strides=(2, 2, 2, 2),
num_res_units=2,
)
loss_function = DiceLoss(sigmoid=True)
optimizer = torch.optim.Adam(model.parameters(), 1e-3)
dice_metric = DiceMetric(include_background=True, to_onehot_y=False, sigmoid=True, reduction="mean")
# + [markdown] colab_type="text" id="BSHNgJ2e8908"
# # [Catalyst](https://github.com/catalyst-team/catalyst) experiment
# + [markdown] colab_type="text" id="oO8ijD62CCvm"
# ## Setup Runner
# + colab={} colab_type="code" id="wtiY6gC1CCb6"
class MonaiSupervisedRunner(catalyst.dl.SupervisedRunner):
def forward(self, batch):
if self.is_train_loader:
output = {self.output_key: self.model(batch[self.input_key])}
elif self.is_valid_loader:
roi_size = (96, 96, 96)
sw_batch_size = 4
output = {
self.output_key: sliding_window_inference(
batch[self.input_key], roi_size, sw_batch_size, self.model
)
}
elif self.is_infer_loader:
roi_size = (96, 96, 96)
sw_batch_size = 4
batch = self._batch2device(batch, self.device)
output = {
self.output_key: sliding_window_inference(
batch[self.input_key], roi_size, sw_batch_size, self.model
)
}
output = {**output, **batch}
return output
# + [markdown] colab_type="text" id="OVBQ44E3CJst"
# ## Run experiment
# + colab={"base_uri": "https://localhost:8080/", "height": 819} colab_type="code" id="NogjkLJaf-1U" outputId="d96f2f00-42f1-4863-ccfb-1a1facbbe652"
log_dir = os.path.join(root_dir, "logs")
runner = MonaiSupervisedRunner(
input_key="img", input_target_key="seg", output_key="logits"
) # you can also specify `device` here
runner.train(
loaders={"train": train_loader, "valid": val_loader},
model=model,
criterion=loss_function,
optimizer=optimizer,
num_epochs=6,
logdir=log_dir,
main_metric="dice_metric",
minimize_metric=False,
verbose=False,
timeit=True, # let's use minimal logs, but with time checkers
callbacks={
"loss": catalyst.dl.CriterionCallback(input_key="seg", output_key="logits"),
"periodic_valid": catalyst.dl.PeriodicLoaderCallback(valid=2),
"dice_metric": catalyst.dl.MetricCallback(
prefix="dice_metric", metric_fn=dice_metric, input_key="seg", output_key="logits"
),
},
load_best_on_end=True, # user-friendly API :)
)
# + [markdown] colab_type="text" id="ugpGCkyS83e0"
# # Tensorboard logs
# -
# %load_ext tensorboard
# %tensorboard --logdir=log_dir
# + [markdown] colab_type="text" id="Z1RDmcUa8tQy"
# # Best model performance visualisation
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="nHYP-ltLqzoC" outputId="5a75dac9-8f25-459f-cddc-cc8b9dfc4767"
for i, valid_output in enumerate(runner.predict_loader(loader=val_loader)):
if i > 4:
break
plt.figure("check", (9, 3))
plt.subplot(1, 3, 1)
plt.title("image " + str(i))
plt.imshow(valid_output["img"].detach().cpu()[0, 0, :, :, 48], cmap="gray")
plt.subplot(1, 3, 2)
plt.title("label " + str(i))
plt.imshow(valid_output["seg"].detach().cpu()[0, 0, :, :, 48])
plt.subplot(1, 3, 3)
plt.title("output " + str(i))
logits = valid_output["logits"]
plt.imshow((logits[0] > 0.5).float().detach().cpu()[0, :, :, 48])
plt.show()
# -
# ## Cleanup data directory
#
# Remove directory if a temporary was used.
# + colab={} colab_type="code" id="8iQXpWNwlUjG"
if directory is None:
shutil.rmtree(root_dir)
|
3d_segmentation/unet_segmentation_3d_catalyst.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
data = pd.DataFrame({'Country': ['Russia','Colombia','Chile','Equador','Nigeria'],
'Rank':[121,40,100,130,11]})
data
data.describe()
#describe() method computes summary statistics of integer / double variables
data.info()
#get the complete information about the data set, we can use info() function
data = pd.DataFrame({'group':['a', 'a', 'a', 'b','b', 'b', 'c', 'c','c'],'ounces':[4, 3, 12, 6, 7.5, 8, 3, 5, 6]})
data
#sort the data frame by ounces - inplace = True will make changes to the data
data.sort_values(by=['ounces'],ascending=True,inplace=False)
#sort the data by not just one column but multiple columns.
data.sort_values(by=['group','ounces'],ascending=[True,False],inplace=False)
#remove duplicate rows.
data = pd.DataFrame({'k1':['one']*3 + ['two']*4, 'k2':[3,2,1,3,3,4,4]})
data
#sort values
data.sort_values(by='k2')
#remove duplicates
data.drop_duplicates()
#remove duplicates based on a particular column
data.drop_duplicates(subset='k1')
data = pd.DataFrame({'food': ['bacon', 'pulled pork', 'bacon', 'Pastrami','corned beef', 'Bacon', 'pastrami', 'honey ham','nova lox'],
'ounces': [4, 3, 12, 6, 7.5, 8, 3, 5, 6]})
data
# +
meat_to_animal = {
'bacon': 'pig',
'pulled pork': 'pig',
'pastrami': 'cow',
'corned beef': 'cow',
'honey ham': 'pig',
'nova lox': 'salmon'
}
def meat_2_animal(series):
if series['food'] == 'bacon':
return 'pig'
elif series['food'] == 'pulled pork':
return 'pig'
elif series['food'] == 'pastrami':
return 'cow'
elif series['food'] == 'corned beef':
return 'cow'
elif series['food'] == 'honey ham':
return 'pig'
else:
return 'salmon'
#create a new variable
data['animal'] = data['food'].map(str.lower).map(meat_to_animal)
data
# -
#another way of doing it is: convert the food values to the lower case and apply the function
lower = lambda x: x.lower()
data['food'] = data['food'].apply(lower)
data['animal2'] = data.apply(meat_2_animal, axis='columns')
data
data.assign(new_variable = data['ounces']*10)
#Another way to create a new variable is by using the assign function
#remove columns
data.drop('animal2',axis='columns',inplace=True)
data
#A quick method for imputing missing values is by filling the missing value with any random number
#also to replace outliers
#Series function from pandas are used to create arrays
data = pd.Series([1., -999., 2., -999., -1000., 3.])
data
data.replace(-999, np.nan,inplace=True)
data
#We can also replace multiple values at once.
data = pd.Series([1., -999., 2., -999., -1000., 3.])
data.replace([-999,-1000],np.nan,inplace=True)
data
data = pd.DataFrame(np.arange(12).reshape((3, 4)),index=['Ohio', 'Colorado', 'New York'],columns=['one', 'two', 'three', 'four'])
data
#Using rename function
data.rename(index = {'Ohio':'SanF'}, columns={'one':'one_p','two':'two_p'},inplace=True)
data
#You can also use string functions
data.rename(index = str.upper, columns=str.title,inplace=True)
data
ages = [20, 22, 25, 27, 21, 23, 37, 31, 61, 45, 41, 32]
#categorize(bin) continuous variables
#Understand the output - '(' means the value is included in the bin, '[' means the value is excluded
bins = [18, 25, 35, 60, 100]
cats = pd.cut(ages, bins)
cats
#To include the right bin value, we can do:
pd.cut(ages,bins,right=False)
#pandas library intrinsically assigns an encoding to categorical variables.
cats.labels
#Let's check how many observations fall under each bin
pd.value_counts(cats)
# +
bin_names = ['Youth', 'YoungAdult', 'MiddleAge', 'Senior']
new_cats = pd.cut(ages, bins,labels=bin_names)
pd.value_counts(new_cats)
# -
pd.value_counts(new_cats).cumsum()
#cumulative sum
df = pd.DataFrame({'key1' : ['a', 'a', 'b', 'b', 'a'],
'key2' : ['one', 'two', 'one', 'two', 'one'],
'data1' : np.random.randn(5),
'data2' : np.random.randn(5)})
df
#calculate the mean of data1 column by key1
grouped = df['data1'].groupby(df['key1'])
grouped.mean()
#slice data
dates = pd.date_range('20130101',periods=6)
df = pd.DataFrame(np.random.randn(6,4),index=dates,columns=list('ABCD'))
df
#get first n rows from the data frame
df[:3]
#slice based on date range
df['20130101':'20130104']
#slicing based on column names
df.loc[:,['A','B']]
#slicing based on both row index labels and column names
df.loc['20130102':'20130103',['A','B']]
#slicing based on index of columns
df.iloc[3] #returns 4th row (index is 3rd)
#returns a specific range of rows
df.iloc[2:4, 0:2]
#returns specific rows and columns using lists containing columns or row indexes
df.iloc[[1,5],[0,2]]
#Boolean indexing based on column values
df[df.A > 0]
#we can copy the data set
df2 = df.copy()
df2['E']=['one', 'one','two','three','four','three']
df2
#select rows based on column values
df2[df2['E'].isin(['two','four'])]
#select all rows except those with two and four
df2[~df2['E'].isin(['two','four'])]
#list all columns where A is greater than C
df.query('A > C')
#using OR condition
df.query('A < B | C > A')
#create a data frame
data = pd.DataFrame({'group': ['a', 'a', 'a', 'b','b', 'b', 'c', 'c','c'],
'ounces': [4, 3, 12, 6, 7.5, 8, 3, 5, 6]})
data
data.pivot_table(values='ounces',index='group',aggfunc=np.mean)
#calculate count by each group
data.pivot_table(values='ounces',index='group',aggfunc='count')
|
NumpyAndPandasTutorial/PandasTutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: rapids-0.20_sc
# language: python
# name: python3
# ---
# # Setup of the AnnData object
# **Author:** [<NAME>](https://github.com/Intron7) (IBSM Freiburg)
# This notebook is just downloader and sets up the AnnData object (https://anndata.readthedocs.io/en/latest/index.html) we will be working with. In this example workflow we'll be looking at a dataset of ca. 90000 cells from lungcancer patients published by [<NAME> al., Cell Research 2020](https://www.nature.com/articles/s41422-020-0355-0).
import wget
import scanpy as sc
import os
import tarfile
import pandas as pd
# First we download the countmartix and metadata file from the Lambrechts lab website.
count_file = './data/LC_counts.tar.gz'
if not os.path.exists(count_file):
os.makedirs("./data",exist_ok=True)
wget.download("http://blueprint.lambrechtslab.org/download/LC_counts.tar.gz", out="./data")
wget.download("http://blueprint.lambrechtslab.org/download/LC_metadata.csv.gz", out="./data")
# We than decompress the data.
tar = tarfile.open(count_file, "r:gz")
tar.extractall("./data")
tar.close()
# Now we can start creating our AnnData object with scanpy (https://scanpy.readthedocs.io/en/stable/index.html).
adata = sc.read_10x_mtx("./data/export/LC_counts/")
# Next we have to append the metadata to `adata.obs`.
obs_df = pd.read_csv("./data/LC_metadata.csv.gz",compression="gzip", index_col=0)
obs_df
# In this case `adata.obs` and the meta_data in `obs_df` have the identical number of cells and the cell barcodes are in the same order. We can therefore just replace `.obs` with `obs_df`
adata.obs = obs_df
# Since `PatientNumber` is a category and not a numerical value we have to change its type. In some cases scanpy doesn't like integers as categories. So we convert it to `str`
adata.obs.PatientNumber = adata.obs.PatientNumber.astype(str)
# During the saving of the adata object string based columns in `.obs` are transformed are changed into categorical data.
os.makedirs("./h5",exist_ok=True)
adata.write("./h5/adata.raw.h5ad")
# If you want to you can now delete the "./data" folder since we won't need it anymore
|
notebooks/data_downloader.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (thesisp375_clone)
# language: python
# name: thesisp375_clone
# ---
# +
import sys
sys.path.append('..')
from msidata.dataset_msi_features_with_patients import PreProcessedMSIFeatureDataset
from testing.logistic_regression import get_precomputed_dataloader
import matplotlib.pyplot as plt
from modules.deepmil import Attention
import torch
import pandas as pd
import os
# -
# +
# We need a model with saved weights
model = Attention(hidden_dim=512)
model.load_state_dict(torch.load('./logs/pretrain/97/classifier_checkpoint_10.tar', map_location='cpu'))
print(model)
# +
# We need a dataset to visualize
test_root_dir="/home/yonis/histogenomics-msc-2019/yoni-code/MsiPrediction/data/msidata/crc_dx/test/"
train_root_dir="/home/yonis/histogenomics-msc-2019/yoni-code/MsiPrediction/data/msidata/crc_dx/train/"
train_dataset = PreProcessedMSIFeatureDataset(
root_dir=train_root_dir,
transform=None,
data_fraction=1,
sampling_strategy='patient',
append_img_path_with=f'_61',
tensor_per_patient=True
)
test_dataset = PreProcessedMSIFeatureDataset(
root_dir=test_root_dir,
transform=None,
data_fraction=1,
sampling_strategy='patient',
append_img_path_with=f'_61',
tensor_per_patient=True
)
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=1,
shuffle=True,
drop_last=False,
num_workers=16,
)
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=1,
shuffle=False,
drop_last=False,
num_workers=16,
)
# +
# We need to visualize
from PIL import Image
import os
test_data=pd.read_csv(os.path.join(test_root_dir, 'data.csv'))
mss=0
msi=0
model.train()
for step, data in enumerate(test_loader):
x=data[0]
x.requires_grad = True
y=data[1]
patient=data[2][0]
img_path=data[3]
if mss == 5 and msi == 5:
break
else:
if y.item()==0:
if mss == 5:
continue
mss+=1
else:
if msi == 5:
continue
msi+=1
Y_out, Y_hat, A = model.forward(x)
# binary_Y_prob = Y_out.softmax(dim=1)[0][1]
binary_Y_prob = Y_out.softmax(dim=1)[0][1]
# --- Get MSI gradient labels
frac = Y_out[0][1] / (Y_out[0].sum())
frac.backward()
dMSIdA = model.A_grad
# --- Done with MSI Grad labels
test_data.loc[test_data['patient_id']==patient, 'attention'] = A.flatten().detach().numpy()
test_data.loc[test_data['patient_id']==patient, 'dMSIdA'] = dMSIdA.flatten().detach().numpy()
test_data.loc[test_data['patient_id']==patient, 'a_dMSIdA'] = (A.flatten()*dMSIdA.flatten()).detach().numpy()
att_data = test_data[test_data['patient_id']==patient].copy()
grad_data = test_data[test_data['patient_id']==patient].copy()
# assert (patient_image_paths == patient_image_paths.sort_values()).all(), "Sorting gone wrong. Since we have no mapping between tensor index and image, we can't do anything now."
att_sorted = att_data.sort_values(by=['attention'],ascending=False).copy()
grad_sorted = grad_data.sort_values(by=['a_dMSIdA'], ascending=False).copy()
top_attention = att_sorted['attention'].head(5)
bottom_attention = att_sorted['attention'].tail(5)
top_att_images = att_sorted['img'].head(5)
bottom_att_images = att_sorted['img'].tail(5)
top_grad = grad_sorted['a_dMSIdA'].head(5)
bottom_grad = grad_sorted['a_dMSIdA'].tail(5)
top_grad_images = grad_sorted['img'].head(5)
bottom_grad_images = grad_sorted['img'].tail(5)
plot_attention=False
if plot_attention:
f, axarr = plt.subplots(2,5, sharex=True, sharey=True, figsize=[12,6])
plt.suptitle(f'Patient #{patient}. Prediction: {binary_Y_prob:.4f}. Label: {y.item()}')
for idx, (atts, ims, tb) in enumerate(zip([top_attention, bottom_attention], [top_att_images, bottom_att_images], ['top', 'bottom'])):
for ix, (att, im_path) in enumerate(zip(list(atts), list(ims))):
im = Image.open(os.path.join(test_root_dir, im_path))
axarr[idx, ix].imshow(im)
axarr[idx, ix].set_title(f'A: {att:.4f}')
plt.show()
plot_gradient=True
if plot_gradient:
f, axarr = plt.subplots(2,5, sharex=True, sharey=True, figsize=[12,6])
plt.suptitle(f'Patient #{patient}. Prediction: {binary_Y_prob:.4f}. Label: {y.item()}')
for idx, (atts, ims, tb) in enumerate(zip([top_grad, bottom_grad], [top_grad_images, bottom_grad_images], ['top', 'bottom'])):
for ix, (att, im_path) in enumerate(zip(list(atts), list(ims))):
im = Image.open(os.path.join(test_root_dir, im_path))
axarr[idx, ix].imshow(im)
axarr[idx, ix].set_title(f'dMSI/da: {att:.4f}')
plt.show()
# -
|
viz/attention_visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/YeonKang/Tensorflow-with-Colab/blob/master/Lab7_2_normalization_%26_decay_%26_L2_loss.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="NjcyTgO4kbnC" outputId="08e390c9-7e9a-4abe-c080-eb197639aa4f"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
from mpl_toolkits.mplot3d import Axes3D
tf.random.set_seed(777) #for reproducibility
print(tf.__version__)
# + [markdown] id="SpSmqC-vl253"
# **Data**
# + colab={"base_uri": "https://localhost:8080/", "height": 276} id="wAyGHl3Tl4Fx" outputId="bd276b71-ca0f-4f2a-9467-9cde3dfcbecd"
xy = np.array([[828.659973, 833.450012, 908100, 828.349976, 831.659973],
[823.02002, 828.070007, 1828100, 821.655029, 828.070007],
[819.929993, 824.400024, 1438100, 818.97998, 824.159973],
[816, 820.958984, 1008100, 815.48999, 819.23999],
[819.359985, 823, 1188100, 818.469971, 818.97998],
[819, 823, 1198100, 816, 820.450012],
[811.700012, 815.25, 1098100, 809.780029, 813.669983],
[809.51001, 816.659973, 1398100, 804.539978, 809.559998]])
x_train = xy[:, 0:-1]
y_train = xy[:, [-1]]
plt.plot(x_train, 'ro')
plt.plot(y_train)
plt.show()
# + [markdown] id="Mq7GeZUllwbk"
# **normalization**
# + id="62IT_42Blzxd"
def normalization(data):
numerator = data - np.min(data, 0)
denominator = np.max(data, 0) - np.min(data, 0)
return numerator / denominator
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="YvSD4PbWl7WR" outputId="3be5b41f-3f5c-436d-fb20-405090a7508c"
xy = normalization(xy)
print(xy)
x_train = xy[:, 0:-1]
y_train = xy[:, [-1]]
plt.plot(x_train, 'ro')
plt.plot(y_train)
plt.show()
# + [markdown] id="LakUVkvdl1Dn"
# **Linear Regression model**
# + id="oCiLPxaxmBlO"
dataset = tf.data.Dataset.from_tensor_slices((x_train, y_train)).batch(len(x_train))
W = tf.Variable(tf.random.normal((4, 1)), dtype=tf.float32)
b = tf.Variable(tf.random.normal((1,)), dtype=tf.float32)
# + id="VTp7cr4ymFSg"
def linearReg_fn(features):
hypothesis = tf.matmul(features, W) + b
return hypothesis
# + [markdown] id="Awhv8JFtmH6j"
# **L2 loss**
# + id="_NiSVuBomL45"
def l2_loss(loss, beta = 0.01):
W_reg = tf.nn.l2_loss(W) #output = sum(t ** 2) / 2
loss = tf.reduce_mean(loss + W_reg * beta)
return loss
# + id="VURBPa9amOg_"
def loss_fn(hypothesis, features, labels, flag = False):
cost = tf.reduce_mean(tf.square(hypothesis - labels))
if(flag):
cost = l2_loss(cost)
return cost
# + [markdown] id="zjAVGTF8mkkO"
# **Set Learning Decay (5 parameters) to control Lerning Rate**
# + [markdown] id="TRq3ban0mmFe"
#
#
# * starter_learning_rate : initial learning rate
# * global_step : current number of learning
# * 1000 : decay_steps
# * 0.96 : decay_rate
# * decayed_learning_rate = learning_rate * decay_rate ^ (global_step/decay_steps)
#
#
#
#
# + id="Gub7KYAjmnDc"
is_decay = True
starter_learning_rate = 0.1
if(is_decay):
learning_rate = tf.keras.optimizers.schedules.ExponentialDecay(initial_learning_rate=starter_learning_rate,
decay_steps=50,
decay_rate=0.96,
staircase=True)
optimizer = tf.keras.optimizers.SGD(learning_rate)
else:
optimizer = tf.keras.optimizers.SGD(learning_rate=starter_learning_rate)
def grad(hypothesis, features, labels, l2_flag):
with tf.GradientTape() as tape:
loss_value = loss_fn(linearReg_fn(features),features,labels, l2_flag)
return tape.gradient(loss_value, [W,b]), loss_value
# + [markdown] id="du40lgwXmr9T"
# **train model**
# + colab={"base_uri": "https://localhost:8080/"} id="JFRdpaOWmtoq" outputId="e8fccfc7-b9c2-40b1-b248-63a5af1d0171"
EPOCHS = 101
for step in range(EPOCHS):
for features, labels in dataset:
features = tf.cast(features, tf.float32)
labels = tf.cast(labels, tf.float32)
grads, loss_value = grad(linearReg_fn(features), features, labels, False)
optimizer.apply_gradients(grads_and_vars=zip(grads,[W,b]))
if step % 10 == 0:
print("Iter: {}, Loss: {:.4f}".format(step, loss_value))
|
Lab7_2_normalization_&_decay_&_L2_loss.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from notebook_utils import synthesize
from utils import hparams as hp
import utils.text.cleaners as cleaners
import numpy as np
import string
SAMPLE_RATE = 22050
# +
forward_model_path = 'pretrained/forward_300K.pyt'
voc_model_path = 'pretrained/model_loss0.028723_step860000_weights.pyt'
print('*** Configure hparams...')
synthesize.init_hparams('hparams.py')
print('*** Loading forward model')
forward_model = synthesize.get_forward_model(forward_model_path, gpu=False)
print('*** Loading VOC model')
voc_model = synthesize.get_wavernn_model(voc_model_path, gpu=False)
# +
text = 'Đó là lần tôi ở <NAME> non một tháng làm nhiệm vụ nghiên cứu chính sách đất đai cho đồng bào dân tộc thiểu số Tây Nguyên cuối năm 2004'
# Using WaveRNN vocoder
wav = synthesize.synthesize(text, forward_model, voc_model)
# Using Griffin-Lim
gl_wav = synthesize.synthesize(text, forward_model, 'griffinlim')
# +
from IPython.display import Audio
Audio(wav, rate=SAMPLE_RATE)
# +
from IPython.display import Audio
Audio(gl_wav, rate=SAMPLE_RATE)
# -
|
demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# # Bernstein-Vazirani Algorithm
# -
# In this section, we first introduce the Bernstein-Vazirani problem, its classical solution, and the quantum algorithm to solve it. We then implement the quantum algorithm using Qiskit and run it on both a simulator and a device.
#
# ## Contents
#
# 1. [The Bernstein-Vazirani Algorithm](#algorithm)
# 1.1 [Bernstein-Vazirani Problem](#bvproblem)
# 1.2 [The Classical Solution](#bclassical-solution)
# 1.3 [The Quantum Solution](#quantum-solution)
# 2. [Example](#example)
# 3. [Qiskit Implementation](#implementation)
# 3.1 [Simulation](#simulation)
# 3.2 [Device](#device)
# 4. [Problems](#problems)
# 5. [References](#references)
# ## 1. The Bernstein-Vazirani Algorithm<a id='algorithm'></a>
#
# The Bernstein-Vazirani algorithm, first introduced in Reference [1], can be seen as an extension of the Deutsch-Josza algorithm we covered in the last section. It showed that there can be advantages in using a quantum computer as a computational tool for more complex problems than the Deutsch-Josza problem.
#
# ### 1.1 The Bernstein-Vazirani Problem <a id='bvproblem'> </a>
#
# We are again given a black-box function $f$, which takes as input a string of bits ($x$), and returns either $0$ or $1$, that is:
# $$f(\{x_0,x_1,x_2,...\}) \rightarrow 0 \textrm{ or } 1 \textrm{ where } x_n \textrm{ is }0 \textrm{ or } 1 $$
#
# Instead of the function being balanced or constant as in the Deutsch-Josza problem, now the function is guaranteed to return the bitwise product of the input with some string, $s$. In other words, given an input $x$, $f(x) = s \cdot x \, \text{(mod 2)}$. We are expected to find $s$. As a classical reversible circuit, the Bernstein-Vazirani oracle looks like this:
#
# 
#
#
#
# ### 1.2 The Classical Solution <a id='classical-solution'> </a>
#
# Classically, the oracle returns:
# $$f_s(x) = s \cdot x \mod 2$$
# given an input $x$. Thus, the hidden bit string $s$ can be revealed by querying the oracle with the sequence of inputs:
#
# |Input(x)|
# |:-----:|
# |100...0|
# |010...0|
# |001...0|
# |000...1|
#
# Where each query reveals a different bit of $s$ (the bit $s_i$). For example, with `x = 1000...0` one can obtain the least significant bit of $s$, with `x = 0100...0` we can find the next least significant, and so on. This means we would need to call the function $f_s(x)$, $n$ times.
# ### 1.3 The Quantum Solution <a id='quantum-solution'> </a>
#
# Using a quantum computer, we can solve this problem with 100% confidence after only one call to the function $f(x)$. The quantum Bernstein-Vazirani algorithm to find the hidden bit string is very simple:
#
# 1. Initialize the inputs qubits to the $|0\rangle^{\otimes n}$ state, and output qubit to $|{-}\rangle$.
# 2. Apply Hadamard gates to the input register
# 3. Query the oracle
# 4. Apply Hadamard gates to the input register
# 5. Measure
#
# 
#
# To explain the algorithm, let’s look more closely at what happens when we apply a H-gate to each qubit. If we have an $n$-qubit state, $|a\rangle$, and apply the H-gates, we will see the transformation:
#
# $$
# |a\rangle \xrightarrow{H^{\otimes n}} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} (-1)^{a\cdot x}|x\rangle.
# $$
#
#
#
# <details>
# <summary>Explain Equation (Click to Expand)</summary>
# We remember the Hadamard performs the following transformations on one qubit:
#
# $$
# H|0\rangle = \tfrac{1}{\sqrt{2}}(|0\rangle + |1\rangle)
# $$ $$
# H|1\rangle = \tfrac{1}{\sqrt{2}}(|0\rangle - |1\rangle)
# $$
#
# Using summation notation, we could rewrite it like this:
#
# $$
# H|a\rangle = \frac{1}{\sqrt{2}}\sum_{x\in \{0,1\}} (-1)^{a\cdot x}|x\rangle.
# $$
#
# For two qubits, applying a Hadamard to each performs the following transformations:
#
# $$
# H^{\otimes 2}|00\rangle = \tfrac{1}{2}(|00\rangle + |01\rangle + |10\rangle + |11\rangle)
# $$ $$
# H^{\otimes 2}|01\rangle = \tfrac{1}{2}(|00\rangle - |01\rangle + |10\rangle - |11\rangle)
# $$ $$
# H^{\otimes 2}|10\rangle = \tfrac{1}{2}(|00\rangle + |01\rangle - |10\rangle - |11\rangle)
# $$ $$
# H^{\otimes 2}|11\rangle = \tfrac{1}{2}(|00\rangle - |01\rangle - |10\rangle + |11\rangle)
# $$
#
# We can express this using the summation below:
#
# $$
# H^{\otimes 2}|a\rangle = \frac{1}{2}\sum_{x\in \{0,1\}^2} (-1)^{a\cdot x}|x\rangle
# $$
#
# You will hopefully now see how we arrive at the equation above.
# </details>
#
# In particular, when we start with a quantum register $|00\dots 0\rangle$ and apply $n$ Hadamard gates to it, we have the familiar quantum superposition:
#
# $$
# |00\dots 0\rangle \xrightarrow{H^{\otimes n}} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} |x\rangle
# $$
#
# In this case, the phase term $(-1)^{a\cdot x}$ disappears, since $a=0$, and thus $(-1)^{a\cdot x} = 1$.
#
# The classical oracle $f_s$ returns $1$ for any input $x$ such that $s \cdot x\mod 2 = 1$, and returns $0$ otherwise. If we use the same phase kickback trick from the Deutsch-Joza algorithm and act on a qubit in the state $|{-}\rangle$, we get the following transformation:
#
# $$
# |x \rangle \xrightarrow{f_s} (-1)^{s\cdot x} |x \rangle
# $$
#
# The algorithm to reveal the hidden bit string follows naturally by querying the quantum oracle $f_s$ with the quantum superposition obtained from the Hadamard transformation of $|00\dots 0\rangle$. Namely,
#
# $$
# |00\dots 0\rangle \xrightarrow{H^{\otimes n}} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} |x\rangle \xrightarrow{f_a} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} (-1)^{a\cdot x}|x\rangle
# $$
#
# Because the inverse of the $n$ Hadamard gates is again the $n$ Hadamard gates, we can obtain $a$ by
#
# $$
# \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} (-1)^{a\cdot x}|x\rangle \xrightarrow{H^{\otimes n}} |a\rangle
# $$
# ## 2. Example <a id='example'></a>
#
# Let's go through a specific example for $n=2$ qubits and a secret string $s=11$. Note that we are following the formulation in Reference [2] that generates a circuit for the Bernstein-Vazirani quantum oracle using only one register.
#
# <ol>
# <li> The register of two qubits is initialized to zero:
#
#
# $$\lvert \psi_0 \rangle = \lvert 0 0 \rangle$$
#
#
# </li>
#
# <li> Apply a Hadamard gate to both qubits:
#
#
# $$\lvert \psi_1 \rangle = \frac{1}{2} \left( \lvert 0 0 \rangle + \lvert 0 1 \rangle + \lvert 1 0 \rangle + \lvert 1 1 \rangle \right) $$
#
#
# </li>
#
# <li> For the string $s=11$, the quantum oracle performs the operation:
# $$
# |x \rangle \xrightarrow{f_s} (-1)^{x\cdot 11} |x \rangle.
# $$
#
# $$\lvert \psi_2 \rangle = \frac{1}{2} \left( (-1)^{00\cdot 11}|00\rangle + (-1)^{01\cdot 11}|01\rangle + (-1)^{10\cdot 11}|10\rangle + (-1)^{11\cdot 11}|11\rangle \right)$$
#
# $$\lvert \psi_2 \rangle = \frac{1}{2} \left( \lvert 0 0 \rangle - \lvert 0 1 \rangle - \lvert 1 0 \rangle + \lvert 1 1 \rangle \right)$$
#
#
# </li>
#
# <li> Apply a Hadamard gate to both qubits:
#
#
# $$\lvert \psi_3 \rangle = \lvert 1 1 \rangle$$
#
#
# </li>
#
# <li> Measure to find the secret string $s=11$
# </li>
#
#
# </ol>
#
# Use the widget `bv_widget` below. Press the buttons to apply the different steps, and try to follow the algorithm through. You can change the number of input qubits and the value of the secret string through the first two positional arguments.
from qiskit_textbook.widgets import bv_widget
bv_widget(2, "11")
# ## 3. Qiskit Implementation <a id='implementation'></a>
#
# We'll now walk through the Bernstein-Vazirani algorithm implementation in Qiskit for a three bit function with $s=011$.
# +
# initialization
import matplotlib.pyplot as plt
import numpy as np
# importing Qiskit
from qiskit import IBMQ, Aer
from qiskit.providers.ibmq import least_busy
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, transpile, assemble
# import basic plot tools
from qiskit.visualization import plot_histogram
# -
# We first set the number of qubits used in the experiment, and the hidden bit string $s$ to be found by the algorithm. The hidden bit string $s$ determines the circuit for the quantum oracle.
n = 3 # number of qubits used to represent s
s = '011' # the hidden binary string
# We then use Qiskit to program the Bernstein-Vazirani algorithm.
# +
# We need a circuit with n qubits, plus one auxiliary qubit
# Also need n classical bits to write the output to
bv_circuit = QuantumCircuit(n+1, n)
# put auxiliary in state |->
bv_circuit.h(n)
bv_circuit.z(n)
# Apply Hadamard gates before querying the oracle
for i in range(n):
bv_circuit.h(i)
# Apply barrier
bv_circuit.barrier()
# Apply the inner-product oracle
s = s[::-1] # reverse s to fit qiskit's qubit ordering
for q in range(n):
if s[q] == '0':
bv_circuit.i(q)
else:
bv_circuit.cx(q, n)
# Apply barrier
bv_circuit.barrier()
#Apply Hadamard gates after querying the oracle
for i in range(n):
bv_circuit.h(i)
# Measurement
for i in range(n):
bv_circuit.measure(i, i)
bv_circuit.draw()
# -
# ### 3a. Experiment with Simulators <a id='simulation'></a>
#
# We can run the above circuit on the simulator.
# +
# use local simulator
qasm_sim = Aer.get_backend('qasm_simulator')
shots = 1024
qobj = assemble(bv_circuit)
results = qasm_sim.run(qobj).result()
answer = results.get_counts()
plot_histogram(answer)
# -
# We can see that the result of the measurement is the hidden string `011`.
# ### 3b. Experiment with Real Devices <a id='device'></a>
#
# We can run the circuit on the real device as below.
# + tags=["uses-hardware"]
# Load our saved IBMQ accounts and get the least busy backend device with less than or equal to 5 qubits
IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q')
provider.backends()
backend = least_busy(provider.backends(filters=lambda x: x.configuration().n_qubits <= 5 and
x.configuration().n_qubits >= 2 and
not x.configuration().simulator and x.status().operational==True))
print("least busy backend: ", backend)
# + tags=["uses-hardware"]
# Run our circuit on the least busy backend. Monitor the execution of the job in the queue
from qiskit.tools.monitor import job_monitor
shots = 1024
transpiled_bv_circuit = transpile(bv_circuit, backend)
qobj = assemble(transpiled_bv_circuit, shots=shots)
job = backend.run(qobj)
job_monitor(job, interval=2)
# + tags=["uses-hardware"]
# Get the results from the computation
results = job.result()
answer = results.get_counts()
plot_histogram(answer)
# -
# As we can see, most of the results are `011`. The other results are due to errors in the quantum computation.
# ## 4. Exercises <a id='problems'></a>
#
# 1. Use the widget below to see the Bernstein-Vazirani algorithm in action on different oracles:
from qiskit_textbook.widgets import bv_widget
bv_widget(3, "011", hide_oracle=False)
# 2. The above [implementation](#implementation) of Bernstein-Vazirani is for a secret bit string $s = 011$. Modify the implementation for a secret string $s = 1011$. Are the results what you expect? Explain.
# 3. The above [implementation](#implementation) of Bernstein-Vazirani is for a secret bit string $s = 011$. Modify the implementation for a secret string $s = 11101101$. Are the results what you expect? Explain.
# ## 5. References <a id='references'></a>
# 1. <NAME> and <NAME> (1997) "Quantum Complexity Theory" SIAM Journal on Computing, Vol. 26, No. 5: 1411-1473, [doi:10.1137/S0097539796300921](https://doi.org/10.1137/S0097539796300921).
# 2. <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> (2001) "Implementation of a quantum algorithm to solve the Bernstein-Vazirani parity problem without entanglement on an ensemble quantum computer", Phys. Rev. A 64, 042306, [10.1103/PhysRevA.64.042306](https://doi.org/10.1103/PhysRevA.64.042306), [arXiv:quant-ph/0012114](https://arxiv.org/abs/quant-ph/0012114).
import qiskit
qiskit.__qiskit_version__
|
content/ch-algorithms/bernstein-vazirani.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # t-SNE 範例
# (Optional) 若尚未安裝相關套件,執行下一行,然後 restart kernel
# !pip3 install --user sklearn
# !pip3 install --user --upgrade matplotlib
# 載入套件
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import offsetbox
from sklearn import manifold, datasets
# %matplotlib inline
# -
# 載入 digits (4個數字) 資料集
digits = datasets.load_digits(n_class=4)
X = digits.data
y = digits.target
# 設定 模型 估計參數
n_samples, n_features = X.shape
n_neighbors = 30
tsne = manifold.TSNE(n_components=2, random_state=0, init='pca', learning_rate=200., early_exaggeration=12.)
# 資料建模 並 視覺化 結果
# +
# Scale and visualize the embedding vectors
def plot_embedding(X, title=None):
x_min, x_max = np.min(X, 0), np.max(X, 0)
X = (X - x_min) / (x_max - x_min)
plt.figure()
ax = plt.subplot(111)
for i in range(X.shape[0]):
plt.text(X[i, 0], X[i, 1], str(y[i]),
color=plt.cm.Set1(y[i] / 10.),
fontdict={'weight': 'bold', 'size': 9})
if hasattr(offsetbox, 'AnnotationBbox'):
# only print thumbnails with matplotlib > 1.0
shown_images = np.array([[1., 1.]]) # just something big
for i in range(X.shape[0]):
dist = np.sum((X[i] - shown_images) ** 2, 1)
if np.min(dist) < 4e-3:
# don't show points that are too close
continue
shown_images = np.r_[shown_images, [X[i]]]
imagebox = offsetbox.AnnotationBbox(
offsetbox.OffsetImage(digits.images[i], cmap=plt.cm.gray_r),
X[i])
ax.add_artist(imagebox)
plt.xticks([]), plt.yticks([])
if title is not None:
plt.title(title)
# t-SNE embedding of the digits dataset
print("Computing t-SNE embedding")
X_tsne = tsne.fit_transform(X)
plot_embedding(X_tsne,
"t-SNE embedding of the digits")
plt.show()
# -
|
homeworks/D061/Day_061_tsne_sample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="left" src="https://ithaka-labs.s3.amazonaws.com/static-files/images/tdm/tdmdocs/CC_BY.png"><br />
#
# Created by [<NAME>](http://nkelber.com) and Ted Lawless for [JSTOR Labs](https://labs.jstor.org/) under [Creative Commons CC BY License](https://creativecommons.org/licenses/by/4.0/)<br />
# For questions/comments/improvements, email <EMAIL>.<br />
# ____
# # Exploring Word Frequencies
#
# **Description:**
# This [notebook](https://docs.constellate.org/key-terms/#jupyter-notebook) shows how to find the most common words in a
# [dataset](https://docs.constellate.org/key-terms/#dataset). The following processes are described:
#
# * Using the `constellate` client to create a Pandas DataFrame
# * Filtering based on a pre-processed ID list
# * Filtering based on a [stop words list](https://docs.constellate.org/key-terms/#stop-words)
# * Using a `Counter()` object to get the most common words
#
# **Use Case:** For Learners (Detailed explanation, not ideal for researchers)
#
# [Take me to the **Research Version** of this notebook ->](./exploring-word-frequencies-for-research.ipynb)
#
# **Difficulty:** Intermediate
#
# **Completion time:** 60 minutes
#
# **Knowledge Required:**
# * Python Basics ([Start Python Basics I](./python-basics-1.ipynb))
#
# **Knowledge Recommended:**
#
# * [Working with Dataset Files](./working-with-dataset-files.ipynb)
# * [Pandas I](./pandas-1.ipynb)
# * [Counter Objects](./counter-objects.ipynb)
# * [Creating a Stopwords List](./creating-stopwords-list.ipynb)
#
# **Data Format:** [JSON Lines (.jsonl)](https://docs.constellate.org/key-terms/#jsonl)
#
# **Libraries Used:**
# * [constellate](https://docs.constellate.org/key-terms/#tdm-client) client to collect, unzip, and read our dataset
# * [NLTK](https://docs.constellate.org/key-terms/#nltk) to help [clean](https://docs.constellate.org/key-terms/#clean-data) up our dataset
# * [Counter](https://docs.constellate.org/key-terms/#python-counter) from **Collections** to help sum up our word frequencies
#
# **Research Pipeline:**
#
# 1. Build a dataset
# 2. Create a "Pre-Processing CSV" with [Exploring Metadata](./exploring-metadata.ipynb) (Optional)
# 3. Create a "Custom Stopwords List" with [Creating a Stopwords List](./creating-stopwords-list.ipynb) (Optional)
# 4. Complete the word frequencies analysis with this notebook
# ___
# ## Import your dataset
#
# We'll use the `constellate` client to automatically retrieve the dataset in the JSON file format.
#
# Enter a [dataset ID](https://docs.constellate.org/key-terms/#dataset-ID) in the next code cell.
#
# If you don't have a dataset ID, you can:
# * Use the sample dataset ID already in the code cell
# * [Create a new dataset](https://constellate.org/builder)
# * [Use a dataset ID from other pre-built sample datasets](https://constellate.org/dataset/dashboard)
# Creating a variable `dataset_id` to hold our dataset ID
# The default dataset is Shakespeare Quarterly, 1950-present
dataset_id = "7e41317e-740f-e86a-4729-20dab492e925"
# Next, import the `constellate` client, passing the `dataset_id` as an argument using the `get_dataset` method.
# +
# Importing your dataset with a dataset ID
import constellate
# Pull in the sampled dataset (1500 documents) that matches `dataset_id`
# in the form of a gzipped JSON lines file.
# The .get_dataset() method downloads the gzipped JSONL file
# to the /data folder and returns a string for the file name and location
dataset_file = constellate.get_dataset(dataset_id)
# To download the full dataset (up to a limit of 25,000 documents),
# request it first in the builder environment. See the Constellate Client
# documentation at: https://constellate.org/docs/constellate-client
# Then use the `constellate.download` method show below.
#dataset_file = constellate.download(dataset_id, 'jsonl')
# -
# ## Apply Pre-Processing Filters (if available)
# If you completed pre-processing with the "Exploring Metadata and Pre-processing" notebook, you can use your CSV file of dataset IDs to automatically filter the dataset. Your pre-processed CSV file should be in the /data directory.
# +
# Import a pre-processed CSV file of filtered dataset IDs.
# If you do not have a pre-processed CSV file, the analysis
# will run on the full dataset and may take longer to complete.
import pandas as pd
import os
# Define a string that describes the path to the CSV
pre_processed_file_name = f'data/pre-processed_{dataset_id}.csv'
# Test if the path to the CSV exists
# If true, then read the IDs into filtered_id_list
if os.path.exists(pre_processed_file_name):
df = pd.read_csv(pre_processed_file_name)
filtered_id_list = df["id"].tolist()
use_filtered_list = True
print(f'Pre-Processed CSV found. Filtered dataset is ' + str(len(df)) + ' documents.')
else:
use_filtered_list = False
print('No pre-processed CSV file found. Full dataset will be used.')
# -
# ## Extract Unigram Counts from the JSON file (No cleaning)
#
# We pulled in our dataset using a `dataset_id`. The file, which resides in the datasets/ folder, is a compressed JSON Lines file (jsonl.gz) that contains all the metadata information found in the metadata CSV *plus* the textual data necessary for analysis including:
#
# * Unigram Counts
# * Bigram Counts
# * Trigram Counts
# * Full-text (if available)
#
# To complete our analysis, we are going to pull out the unigram counts for each document and store them in a Counter() object. We will import `Counter` which will allow us to use Counter() objects for counting unigrams. Then we will initialize an empty Counter() object `word_frequency` to hold all of our unigram counts.
# +
# Import Counter()
from collections import Counter
# Create an empty Counter object called `word_frequency`
word_frequency = Counter()
# -
# We can read in each document using the tdm_client.dataset_reader.
# +
# Gather unigramCounts from documents in `filtered_id_list` if it is available
for document in constellate.dataset_reader(dataset_file):
if use_filtered_list is True:
document_id = document['id']
# Skip documents not in our filtered_id_list
if document_id not in filtered_id_list:
continue
unigrams = document.get("unigramCount", [])
for gram, count in unigrams.items():
word_frequency[gram] += count
# Print success message
if use_filtered_list is True:
print('Unigrams have been collected only for the ' + str(len(df)) + ' documents listed in your CSV file.')
else:
print('Unigrams have been collected for all documents without filtering from a CSV file.')
# -
# ### Find Most Common Unigrams
# Now that we have a list of the frequency of all the unigrams in our corpus, we need to sort them to find which are most common
for gram, count in word_frequency.most_common(25):
print(gram.ljust(20), count)
# ### Some issues to consider
#
# We have successfully created a word frequency list. There are a couple small issues, however, that we still need to address:
# 1. There are many [function words](https://docs.constellate.org/key-terms/#function-words), words like "the", "in", and "of" that are grammatically important but do not carry as much semantic meaning like [content words](https://docs.constellate.org/key-terms/#content-words), such as nouns and verbs.
# 2. The words represented here are actually case-sensitive [strings](https://docs.constellate.org/key-terms/#string). That means that the string "the" is a different from the string "The". You may notice this in your results above.
#
#
# ## Extract Unigram Counts from the JSON File (with cleaning)
# To address these issues, we need to find a way to remove common [function words](https://docs.constellate.org/key-terms/#function-words) and combine [strings](https://docs.constellate.org/key-terms/#string) that may have capital letters in them. We can address these issues by:
#
# 1. Using a [stopwords](https://docs.constellate.org/key-terms/#stop-words) list to remove common [function words](https://docs.constellate.org/key-terms/#function-words)
# 2. Lowercasing all the characters in each string to combine our counts
# ### Load Stopwords List
#
# If you have created a stopword list in the stopwords notebook, we will import it here. (You can always modify the CSV file to add or subtract words then reload the list.) Otherwise, we'll load the NLTK [stopwords](https://docs.constellate.org/key-terms/#stop-words) list automatically.
# +
# Load a custom data/stop_words.csv if available
# Otherwise, load the nltk stopwords list in English
# Create an empty Python list to hold the stopwords
stop_words = []
# The filename of the custom data/stop_words.csv file
stopwords_list_filename = 'data/stop_words.csv'
if os.path.exists(stopwords_list_filename):
import csv
with open(stopwords_list_filename, 'r') as f:
stop_words = list(csv.reader(f))[0]
print('Custom stopwords list loaded from CSV')
else:
# Load the NLTK stopwords list
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
print('NLTK stopwords list loaded')
# -
# Preview stop words
list(stop_words)
# ### Modify Stopwords List
# The following code examples can be used to modify a stopwords list. We recommend storing your stopwords in a CSV file as shown in the [Creating Stopwords List](./creating-stopwords-list.ipynb) notebook.
#
# |code|change|
# |---|---|
# |stop_words.append('word_to_add')| Append a single word to the list|
# |stop_words = stop_words + ['word_one', 'word_two', 'word_three']| Concatenate multiple words to the list|
# |stop_words.remove('word_to_remove')| Delete a word from the list|
# ### Gather unigrams again with extra cleaning steps
# In addition to using a stopwords list, we will clean up the tokens by lowercasing all tokens and combining them. This will combine tokens with different capitalization such as "quarterly" and "Quarterly." We will also remove any tokens that are not alphanumeric.
# +
# Gather unigramCounts from documents in `filtered_id_list` if available
# and apply the processing.
word_frequency = Counter()
for document in constellate.dataset_reader(dataset_file):
if use_filtered_list is True:
document_id = document['id']
# Skip documents not in our filtered_id_list
if document_id not in filtered_id_list:
continue
unigrams = document.get("unigramCount", [])
for gram, count in unigrams.items():
clean_gram = gram.lower()
if clean_gram in stop_words:
continue
if not clean_gram.isalpha():
continue
if len(clean_gram) < 4:
continue
word_frequency[clean_gram] += count
# -
# ## Display Results
# Finally, we will display the 20 most common words by using the `.most_common()` method on the `Counter()` object.
# Print the most common processed unigrams and their counts
for gram, count in word_frequency.most_common(25):
print(gram.ljust(20), count)
# ## Export Results to a CSV File
# The word frequency data can be exported to a CSV file.
# +
# Add output method to csv
import csv
with open(f'./data/word_frequencies_{dataset_id}.csv', 'w') as f:
writer = csv.writer(f)
writer.writerow(['unigram', 'count'])
for gram, count in word_frequency.most_common():
writer.writerow([gram, count])
# -
# ## Create a Word Cloud to Visualize the Data
# A visualization using the WordCloud library in Python. To learn more about customizing a wordcloud, [see the documentation](http://amueller.github.io/word_cloud/generated/wordcloud.WordCloud.html).
# Add wordcloud
from wordcloud import WordCloud
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from PIL import Image
### Download cloud image for our word cloud shape
import urllib.request
download_url = 'https://ithaka-labs.s3.amazonaws.com/static-files/images/tdm/tdmdocs/sample_cloud.png'
urllib.request.urlretrieve(download_url, './data/sample_cloud.png')
# +
# Create a wordcloud from our data
# Adding a mask shape of a cloud to your word cloud
# By default, the shape will be a rectangle
# You can specify any shape you like based on an image file
cloud_mask = np.array(Image.open('./data/sample_cloud.png')) # Specifies the location of the mask shape
cloud_mask = np.where(cloud_mask > 3, 255, cloud_mask) # this line will take all values greater than 3 and make them 255 (white)
### Specify word cloud details
wordcloud = WordCloud(
width = 800, # Change the pixel width of the image if blurry
height = 600, # Change the pixel height of the image if blurry
background_color = "white", # Change the background color
colormap = 'viridis', # The colors of the words, see https://matplotlib.org/stable/tutorials/colors/colormaps.html
max_words = 150, # Change the max number of words shown
min_font_size = 4, # Do not show small text
# Add a shape and outline (known as a mask) to your wordcloud
contour_color = 'blue', # The outline color of your mask shape
mask = cloud_mask, #
contour_width = 1
).generate_from_frequencies(word_frequency)
mpl.rcParams['figure.figsize'] = (20,20) # Change the image size displayed
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# -
|
exploring-word-frequencies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Creating Dataframes of Different Statistics
# ##### Creating timeframe
# +
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import dates as md
from matplotlib import ticker
import scipy as scp
import scipy.optimize as opt
import csv
import math
import random
import pandas as pd
import copy
from datetime import datetime, timedelta
import pytz
import matplotlib.colors as mcolors
from matplotlib.patches import Polygon
# +
start_date = datetime(2021, 12, 26, 0, 0, 0) #1th of October, 2021, 00:00
start_date = pd.to_datetime(start_date, utc=True)
end_date = datetime(2022, 5, 10, 0, 0, 0) #1st of January, 2022, 00:00
end_date = pd.to_datetime(end_date, utc=True)
dateColumn = pd.date_range(start_date, end_date, freq='D')
dateColumn = pd.DataFrame(dateColumn, columns=['date'])
dateColumn.head()
# -
# ##### Creating LUSD dataframe
# +
LUSD_Utilization = pd.read_csv('bb-USD_data/liquity/LUSD Utilization.csv')
LUSD = pd.DataFrame()
LUSD['date'] = pd.to_datetime(LUSD_Utilization['hour'], utc=True)
LUSD['LUSD in SP'] = LUSD_Utilization['stabilityPool']
#LUSD['LUSD other'] = LUSD_Utilization.iloc[:, 1:].sum(1, numeric_only=True) - LUSD_Utilization['stabilityPool']
LUSD = LUSD.merge(right=dateColumn, on='date', how='right')
LUSD = LUSD.groupby(LUSD['date'].dt.date).mean().reset_index()
LUSD['date'] = pd.to_datetime(LUSD['date'], utc=True)
LUSD.head()
# -
# ##### Creating LQTY dataframe
# +
Total_LQTY_Staked = pd.read_csv('bb-USD_data/liquity/LQTY Supply.csv')
LQTY = pd.DataFrame()
LQTY['date'] = pd.to_datetime(Total_LQTY_Staked['day'], utc=True)
LQTY['LQTY total'] = Total_LQTY_Staked['circulatingSupply']
LQTY = LQTY.merge(right=dateColumn, on='date', how='right')
LQTY = LQTY.fillna(method='bfill')
LQTY = LQTY.groupby(LQTY['date'].dt.date).mean().reset_index()
LQTY['date'] = pd.to_datetime(LQTY['date'], utc=True)
plt.plot(LQTY['LQTY total'])
# +
# Total_LQTY_Staked = pd.read_csv('bb-USD_data/liquity/Total LQTY Staked.csv')
# LQTY = pd.DataFrame()
# LQTY['date'] = pd.to_datetime(Total_LQTY_Staked['hour'], utc=True)
# LQTY['LQTY total'] = Total_LQTY_Staked['totalLQTYClaimed']
# LQTY = LQTY.merge(right=dateColumn, on='date', how='right')
# LQTY = LQTY.fillna(method='bfill')
# LQTY = LQTY.groupby(LQTY['date'].dt.date).mean().reset_index()
# LQTY['date'] = pd.to_datetime(LQTY['date'], utc=True)
# LQTY.head()
# -
# ##### Creating Liquidations dataframe
# +
Liquidations_ = pd.read_csv('bb-USD_data/liquity/Recent Liquidations.csv')
Liquidations = pd.DataFrame()
Liquidations['date'] = pd.to_datetime(Liquidations_['timestamp'], utc=True)
Liquidations['LIQ col'] = Liquidations_['collateral']
Liquidations['LIQ debt'] = Liquidations_['debt']
Liquidations['LIQ price'] = Liquidations_['price']
Liquidations['LIQ CR'] = Liquidations_['collateralRatio']
Liquidations['LIQ mode'] = Liquidations_['mode']
Liquidations = Liquidations[(Liquidations['date'] >= start_date) & (Liquidations['date'] <= end_date)].merge(right=dateColumn, on='date', how='outer')
#Liquidations = Liquidations.merge(right=dateColumn, on='date', how='outer')
Liquidations.sort_values(by='date', ignore_index=True, inplace=True)
Liquidations.loc[:, 'LIQ col':'LIQ CR'] = Liquidations.loc[:, 'LIQ col':'LIQ CR'].fillna(value=0)
Liquidations.loc[:, 'LIQ mode'] = Liquidations.loc[:, 'LIQ mode'].fillna(value = 'none')
#Liquidations = Liquidations[Liquidations['LIQ col'] != 0]
Liquidations.head()
# -
# ##### Loading aUSD Data
# +
aUSD_TVL = pd.read_csv('bb-USD_data/balancer/aUSD-TVL.csv')
aUSD_Rewards = pd.read_csv('bb-USD_data/balancer/BAL-Rewards.csv')
aUSD_TVL = aUSD_TVL.rename(columns={'day':'date', 'tvl':'aUSD TVL'})
aUSD_TVL = aUSD_TVL[aUSD_TVL['pool'] == 'BB-A-USD (7b507753)'].drop(columns=['pool'])
aUSD_TVL['date'] = pd.to_datetime(aUSD_TVL['date'], utc=True)
aUSD_Rewards = aUSD_Rewards.rename(columns={'week':'date', 'amount':'BAL Reward'}).reset_index(drop=True)
aUSD_Rewards['date'] = pd.to_datetime(aUSD_Rewards['date'])
aUSD = aUSD_TVL.merge(right=dateColumn, on='date', how='outer')
aUSD = aUSD.merge(right=aUSD_Rewards, on='date', how='outer').sort_values(by='date').reset_index(drop=True)
aUSD = aUSD.fillna(0)
aUSD.head(50)
# -
# ## Loading Prices
# ##### Loading ETH Price
ETHprice = pd.read_csv('bb-USD_data/prices/ETH-Price.csv')
ETHprice.drop(ETHprice.loc[:, ['market_cap', 'total_volume']], axis=1, inplace=True)
ETHprice['snapped_at'] = pd.to_datetime(ETHprice['snapped_at'], utc=True )
ETHprice = ETHprice.sort_values(by='snapped_at', ascending=True, ignore_index=True)
ETHprice.rename(columns = {'snapped_at':'date', 'price':'ETH Price'}, inplace=True)
ETHprice = ETHprice.merge(right=dateColumn, how='outer')
ETHprice = ETHprice[(ETHprice['date'] >= start_date) & (ETHprice['date'] <= end_date)].sort_values(by='date').reset_index(drop=True).fillna(method='ffill')
ETHprice = ETHprice.groupby(ETHprice['date'].dt.date).mean().reset_index()
ETHprice['date'] = pd.to_datetime(ETHprice['date'], utc=True)
ETHprice.head()
# ##### Loading BAL Price
BALprice = pd.read_csv('bb-USD_data/prices/BAL-Price.csv')
BALprice = BALprice.loc[:, ('snapped_at', 'price')].rename(columns={'snapped_at': 'date', 'price':'BAL Price'})
BALprice.loc[:, 'date'] = pd.to_datetime(BALprice['date'])
BALprice = BALprice.merge(right=dateColumn, how='outer', on='date')
BALprice = BALprice.sort_values(by='date', ignore_index=True)
BALprice = BALprice[(BALprice['date'] >= start_date) & (BALprice['date'] <= end_date)].fillna(method='ffill')
BALprice = BALprice.reset_index(drop=True)
BALprice = BALprice.groupby(BALprice['date'].dt.date).mean().reset_index()
BALprice['date'] = pd.to_datetime(BALprice['date'], utc=True)
BALprice.head()
# ##### Loading LQTY Price
LQTYprice = pd.read_csv('bb-USD_data/prices/LQTY-Price.csv')
LQTYprice = LQTYprice.loc[:, ('snapped_at', 'price')].rename(columns={'snapped_at':'date', 'price':'LQTY Price'})
LQTYprice['date'] = pd.to_datetime(LQTYprice['date'])
LQTYprice = LQTYprice.merge(right=dateColumn, how='outer', on='date')
LQTYprice = LQTYprice.sort_values(by='date', ignore_index=True)
LQTYprice = LQTYprice[(LQTYprice['date'] >= start_date) & (LQTYprice['date'] <= end_date)].fillna(method='ffill')
LQTYprice = LQTYprice.reset_index(drop=True)
LQTYprice = LQTYprice.groupby(LQTYprice['date'].dt.date).mean().reset_index()
LQTYprice['date'] = pd.to_datetime(LQTYprice['date'], utc=True)
LQTYprice.head()
# ##### Loading Gas Prices
#
# Gas Prices are loaded in ETH
# +
Gas = pd.read_csv('bb-USD_data/prices/GAS-Price.csv')
Gas['Date(UTC)'] = pd.to_datetime(Gas['Date(UTC)'], utc=True)
Gas = Gas.drop(columns=['UnixTimeStamp']).rename(columns={'Date(UTC)':'date', 'Value (Wei)':'GAS Price'})
Gas = Gas[(Gas['date'] >= start_date) & (Gas['date'] <= end_date)]
Gas['GAS Price'] = Gas['GAS Price'] / 10**18
Gas.head()
# -
# ## Loading Swaps on Curve and Uniswap
# +
Curve_TVL = pd.read_csv('bb-USD_data/dex/Curve-LUSD-TVL.csv')
Curve_Volume = pd.read_csv('bb-USD_data/dex/Curve-LUSD-Volume.csv')
Curve_TVL['Curve TVL'] = Curve_TVL['3crvBalance'] + Curve_TVL['lusdBalance']
Curve_TVL = Curve_TVL.rename(columns={'hour':'date'})[['date', 'Curve TVL']]
Curve_TVL['date'] = pd.to_datetime(Curve_TVL['date'], utc=True)
Curve_TVL = Curve_TVL.groupby(Curve_TVL['date'].dt.date).mean().reset_index()
Curve_TVL['date'] = pd.to_datetime(Curve_TVL['date'], utc=True)
Curve_Volume = Curve_Volume.rename(columns={'day':'date', 'lusd':'Curve Volume'})
Curve_Volume['date'] = pd.to_datetime(Curve_Volume['date'], utc=True)
Curve_Volume = Curve_Volume.groupby(Curve_Volume['date'].dt.date).sum().reset_index()
Curve_Volume['date'] = pd.to_datetime(Curve_Volume['date'], utc=True)
Curve = Curve_TVL.merge(right=Curve_Volume, on='date', how='outer').sort_values(by='date').reset_index(drop=True)
Curve['Curve TVL'] = Curve['Curve TVL'].fillna(method='ffill')
Curve['Curve Volume'] = Curve['Curve Volume'].fillna(0)
Curve['date'] = pd.to_datetime(Curve['date'], utc=True)
Curve = Curve.sort_values(by='date').reset_index(drop=True)
Curve.tail()
# -
# ### Merging All Data into Single Dataframe
#
# ##### Creating main dataframe and merging info
# +
Data = pd.DataFrame()
Data = copy.deepcopy(dateColumn)
Data = Data.merge(LUSD, how='outer', on='date')
Data = Data.merge(LQTY, how='outer', on='date')
Data = Data.merge(Liquidations, how='outer', on='date')
Data = Data.merge(ETHprice, on='date', how='outer')
Data = Data.merge(BALprice, on='date', how='outer')
Data = Data.merge(LQTYprice, on='date', how='outer')
Data = Data.merge(aUSD, on='date', how='outer')
Data = Data.merge(Gas, on='date', how='outer')
Data = Data.merge(Curve, on='date', how='outer')
Data = Data[(Data['date'] >= start_date) & (Data['date'] <= end_date)]
Data = Data.sort_values(by=['date'], ignore_index=True).reset_index(drop=True)#.fillna(method='ffill')
#Data[['ETH Price', 'BAL Price', 'LQTY Price']] = Data[['ETH Price', 'BAL Price', 'LQTY Price']].fillna(method='ffill')
#Data[['ETH Price', 'BAL Price', 'LQTY Price']] = Data[['ETH Price', 'BAL Price', 'LQTY Price']].fillna(method='bfill')
#Data[Data.columns.drop(['ETH Price', 'LIQ mode'])] = Data[Data.columns.drop(['ETH Price', 'LIQ mode'])].fillna(method='ffill')
for c in ['LUSD in SP', 'LQTY total', 'ETH Price', 'BAL Price', 'LQTY Price', 'aUSD TVL', 'GAS Price', 'Curve TVL']:
Data[c].fillna(method='ffill', inplace=True)
for c in ['BAL Reward', 'Curve Volume']:
Data[c].fillna(value=0, inplace=True)
Data.head()
# -
# ## Initializing Pool with 50/50 $ 60M Total TVL
# ### Creating a Pool Dataframe
# ### Operations Gas Requirement
# We take into account the gas price for such operations like ETH, BAL and LQTY withdrawal and Stability Pool liquidity provision.
# The respective gas estimations for these operations are:
#
# * SP add liquidity: 300,000
# * ETH withdrawal: 500,000
# * LQTY withdrawal: 350,000
# * BAL withdrawal: 100,000
# * Trading operation: 1% fee + gas
# ### B.Protocol comissions
#
# In case we route the liquidity of LUSD through B.Protocol, we use the following rules (according to B.Protocol policy):
#
# * Comission - 1% (This means 1% of ETH income from liquidations will be paid to B.Protocol)
# * ETH price sale - 0-4% (When B.Protocol puts ETH for sale, it allows maximum decrease in price of 4% relative to the market price)
def create_pool(Data):
Pool_ = pd.DataFrame()
#columns = ['date', 'LUSD', 'aUSD', 'LQTY', 'ETH', 'SP share', 'ETH received', 'BAL received']
#)
Pool_['date'] = Data['date']
# ---Main balances---
Pool_.loc[:, 'LUSD'] = 30e6 #LUSD in Pool_
Pool_.loc[:, 'aUSD'] = 30e6 #aUSD in Pool_
# ---Pool_ shares---
# Calculated as (Amount in Pool_) / (Amount in common Pool_)
# For example,
# for LUSD share it will be (LUSD in Pool_ (staked)) / (LUSD in Liquity Stability Pool_)
# for aUSD share it will be (aUSD in Pool_) / (TVL in bb-a-USD Balancer Pool_)
Pool_.loc[:, 'SP share'] = 0 #Stability Pool_ share
Pool_.loc[:, 'aUSD share'] = 0 #Balancer aUSD pool_ share
#Reward tokens balances
Pool_.loc[:, 'LQTY'] = 0 #LQTY Rewards collected, but not withdrawn
Pool_.loc[:, 'BAL'] = 0 #BAL Rewards collected, but not withdrawn
Pool_.loc[:, 'ETH'] = 0 #ETH Rewards collected, but not withdrawn
#Reward tokens income
Pool_.loc[:, 'ETH received'] = 0 #ETH received at the timestamp
Pool_.loc[:, 'BAL received'] = 0 #BAL received at the timestamp
Pool_.loc[:, 'LQTY received'] = 0 #LQTY received at the timestamp
#Reward tokens sold
Pool_.loc[:, 'ETH sold'] = 0 #ETH sold at the timestamp
Pool_.loc[:, 'BAL sold'] = 0 #BAL sold at the timestamp
Pool_.loc[:, 'LQTY sold'] = 0 #LQTY sold at the timestamp
#Trading fees
#Trading income is calculated as follows: we assume that the trading volume on Curve is split amongst our pool_ and Curve LUSD-3crv pool_
# Income = Fee% * Curve Trading Volume * Pool_ TVL / (Pool_ TVL + Curve TVL)
Pool_.loc[:, 'Trading income'] = 0 # Total income (in USD) from trading operations
Pool_.loc[:, 'Trading fees'] = 0 # Total fees paid for trading operations (1% of the amount traded (in USD))
#Gas spendings
Pool_.loc[:, 'BAL gas'] = 0
Pool_.loc[:, 'ETH gas'] = 0
Pool_.loc[:, 'LQTY gas'] = 0
Pool_.loc[:, 'SP gas'] = 0
#B.Protocol spendings
Pool_.loc[:, 'ETH sale'] = 0 # Total losses from ETH being sold with sale (in USD)
Pool_.loc[:, 'ETH comission'] = 0 # Comission paid to B.Protocol (in USD)
#DAO fees reserve
Pool_.loc[:, 'LQTY reserve'] = 0 # LQTY accumulated in PowerPool_ DAO Treasury
Pool_.loc[:, 'BAL reserve'] = 0
#LUSD burned
Pool_.loc[:, 'LUSD burn'] = 0 # LUSD in our pool_ that was burned during liquidations
Pool_ = Pool_.sort_values('date', ignore_index=True).reset_index(drop=True)
return Pool_
def run(Pool_, Data_, BProt = False, sale_ = 0, reserve_ = 0, DAO_fees_ = 0, swaps = False):
# Bring all parameters from percentage to decimal
sale = sale_/100
reserve = reserve_/100
DAO_fees = DAO_fees_/100
for i in range(len(Pool_)):
# Bring the previous balances to the current timestamp
if (i > 0):
Pool_.loc[i, 'LUSD'] = Pool_['LUSD'][i-1]
Pool_.loc[i, 'ETH'] = Pool_['ETH'][i-1]
Pool_.loc[i, 'aUSD'] = Pool_['aUSD'][i-1]
Pool_.loc[i, 'LQTY'] = Pool_['LQTY'][i-1]
Pool_.loc[i, 'BAL'] = Pool_['BAL'][i-1]
Pool_.loc[i, 'LQTY reserve'] = Pool_['LQTY reserve'][i-1]
# Calculate shares of the pool_ parts at the timestep
Pool_.loc[i, 'SP share'] = Pool_['LUSD'][i] / Data_['LUSD in SP'][i] * (1 - reserve)
Pool_.loc[i, 'aUSD share'] = Pool_['aUSD'][i] / Data_['aUSD TVL'][i]
# If liquidation occurs, process liquidation gains and LUSD burn
ETH_received = Data_['LIQ col'][i]*(1-0.005)*Pool_['SP share'][i]
LUSD_burned = Data_['LIQ debt'][i]*Pool_['SP share'][i]
Pool_.loc[i, 'ETH received'] = ETH_received
Pool_.loc[i, 'ETH'] += ETH_received
Pool_.loc[i, 'LUSD'] -= LUSD_burned
Pool_.loc[i, 'LUSD burn'] += LUSD_burned
gas_price = Data_['GAS Price'][i] * Data_['ETH Price'][i]
if (BProt):
#Using B.Protocol:
#No gas spendings
#1% B.Protocol comission
#0-4% ETH sale
ETH_in_pool_ = Pool_['ETH'][i]
ETH_sold = ETH_in_pool_ * (1 - 0.01) #1% comission
ETH_comission = ETH_in_pool_ * 0.01
LUSD_bought = ETH_sold * Data_['ETH Price'][i] * (1 - sale)
ETH_sale = ETH_sold * Data_['ETH Price'][i] * sale
Pool_.loc[i, 'ETH'] = 0
Pool_.loc[i, 'LUSD'] += LUSD_bought
Pool_.loc[i, 'ETH sold'] += ETH_sold
Pool_.loc[i, 'ETH sale'] += ETH_sale
Pool_.loc[i, 'ETH comission'] += ETH_comission
#gas price on the current timestamp (USD per 1 unit of Gas)
# ---Check Pool_ "account" and sell reward tokens if conditions met---
else:
#checking if any ETH is on the account
ETH_in_pool_ = Pool_['ETH'][i]
#LUSD deposit gas: 300,000 gwei
#ETH withdraw gas: 500,000 gwei
# We allow gas spendings of up to 10% of the trade volume
if (ETH_in_pool_ * Data_['ETH Price'][i] >= 10 * 500000 * gas_price):
ETH_sold = ETH_in_pool_
LUSD_bought = ETH_sold * Data_['ETH Price'][i] * (1 - 0.01) - 500000 * gas_price #we spend gas to withdraw ETH from Liquity
Pool_.loc[i, 'ETH'] = 0
Pool_.loc[i, 'LUSD'] += LUSD_bought - 300000 * gas_price #we spend gas to restake LUSD into Stability Pool_
Pool_.loc[i, 'ETH sold'] += ETH_sold
Pool_.loc[i, 'SP gas'] += 300000 * gas_price
Pool_.loc[i, 'ETH gas'] += 500000 * gas_price
Pool_.loc[i, 'Trading fees'] += ETH_sold * Data_['ETH Price'][i] * 0.01
#checking if we have enough BAL tokens to sell and add liquidity to the pool_
# We allow gas spendings of up to 10% of the trade volume
if (Pool_['BAL'][i] >= 10 * 100000 * gas_price):
BAL_sold = Pool_['BAL'][i]
aUSD_bought = BAL_sold * Data_['BAL Price'][i] * (1 - 0.01) - 100000 * gas_price #we spend gas to withdraw BAL rewards
Pool_.loc[i, 'BAL'] = 0
Pool_.loc[i, 'aUSD'] += aUSD_bought
Pool_.loc[i, 'BAL sold'] += BAL_sold
Pool_.loc[i, 'BAL gas'] += 100000 * gas_price
Pool_.loc[i, 'Trading fees'] += BAL_sold * Data_['BAL Price'][i] * 0.01
#checking if any LQTY is on the account
if (Pool_['LQTY'][i] > 10 * 350000 * gas_price):
LQTY_sold = Pool_['LQTY'][i]
LUSD_bought = LQTY_sold * Data_['LQTY Price'][i] * (1 - 0.01) - 350000 * gas_price #we spend gas to withdraw LQTY rewards
Pool_.loc[i, 'LQTY'] = 0
Pool_.loc[i, 'LUSD'] += LUSD_bought - 300000 * gas_price #we spend gas to restake LUSD into Stability Pool_
Pool_.loc[i, 'LQTY sold'] += LQTY_sold
Pool_.loc[i, 'LQTY gas'] += 350000 * gas_price
Pool_.loc[i, 'SP gas'] += 300000 * gas_price
Pool_.loc[i, 'Trading fees'] += LQTY_sold * Data_['LQTY Price'][i] * 0.01
#calculating hypothetical trading volume fees
# 0.04% fee
if swaps:
Trading_volume_income = (Pool_['LUSD'][i] + Pool_['aUSD'][i])/Data_['Curve TVL'][i] * Data_['Curve Volume'][i] * 0.04 / 100
# We assume the trading income is split equally between the parts of our pool_
Pool_.loc[i, 'aUSD'] += Trading_volume_income / 2
Pool_.loc[i, 'LUSD'] += Trading_volume_income / 2
Pool_.loc[i, 'Trading income'] += Trading_volume_income
# Calculating reward incomes
if (i > 0):
#calculating LQTY reward
# LQTY rewards = SP share * (LQTY Total Supply (t) - LQTY Total Supply (t-1))
LQTY_minted = Data_['LQTY total'][i] - Data_['LQTY total'][i-1]
Pool_.loc[i, 'LQTY'] += LQTY_minted * Pool_['SP share'][i] * (1 - DAO_fees)
Pool_.loc[i, 'LQTY received'] += LQTY_minted * Pool_['SP share'][i]
Pool_.loc[i, 'LQTY reserve'] += LQTY_minted * Pool_['SP share'][i] * DAO_fees
#calculating aUSD revenue (this part is not accounted for)
'''fees = Data_['aUSD revenues'][i]*Pool_['aUSD share'][i]
Pool_['aUSD'] += fees'''
#calculating BAL rewards
# BAL rewards = aUSD share * aUSD BAL rewards (t)
BAL_received = Data['BAL Reward'][i] * Pool_['aUSD share'][i]
Pool_.loc[i, 'BAL received'] = BAL_received * (1 - DAO_fees)
Pool_.loc[i, 'BAL'] += BAL_received * (1 - DAO_fees)
Pool_.loc[i, 'BAL reserve'] += BAL_received * DAO_fees
return Pool_
# +
BProt = False
Sale = 0
Reserve = 0
DAO_fees = 7.5
Swaps = True
Pool = create_pool(Data)
Pool_0 = create_pool(Data)
Pool_4 = create_pool(Data)
Pool_2 = create_pool(Data)
Pool = run(Pool, Data, False, Sale, Reserve, DAO_fees, Swaps)
Pool_0 = run(Pool_0, Data, True, 0, Reserve, DAO_fees, Swaps)
Pool_4 = run(Pool_4, Data, True, 4, Reserve, DAO_fees, Swaps)
Pool_2 = run(Pool_2, Data, True, 2, Reserve, DAO_fees, Swaps)
# -
(Pool_4['SP share'] == Pool['SP share']).all()
# ### Data peculiarities
# +
# %config InlineBackend.figure_format = 'svg'
plt.plot(Data['date'], Data['Curve Volume'], label='Trading Volume', lw=0, marker='.')
plt.plot(Data['date'], Data['Curve TVL'], label='TVL', lw=0, marker='.')
plt.title('Curve LUSD-3CRV Key Metrics')
plt.xlabel('Date')
plt.ylabel('Millions of $')
plt.legend()
scale_y = 1e6
ticks_y = ticker.FuncFormatter(lambda x, pos: '{0:g}'.format(x/scale_y))
plt.gca().yaxis.set_major_formatter(ticks_y)
plt.grid()
date_form = md.DateFormatter("%b-%d")
plt.gca().xaxis.set_major_formatter(date_form)
# -
# ### Results
def gradient_fill(x, y, fill_color=None, ax=None, **kwargs):
"""
Plot a line with a linear alpha gradient filled beneath it.
Parameters
----------
x, y : array-like
The data values of the line.
fill_color : a matplotlib color specifier (string, tuple) or None
The color for the fill. If None, the color of the line will be used.
ax : a matplotlib Axes instance
The axes to plot on. If None, the current pyplot axes will be used.
Additional arguments are passed on to matplotlib's ``plot`` function.
Returns
-------
line : a Line2D instance
The line plotted.
im : an AxesImage instance
The transparent gradient clipped to just the area beneath the curve.
"""
if ax is None:
ax = plt.gca()
line, = ax.plot(x, y, **kwargs)
if fill_color is None:
fill_color = line.get_color()
zorder = line.get_zorder()
alpha = line.get_alpha()
alpha = 1.0 if alpha is None else alpha
z = np.empty((100, 1, 4), dtype=float)
rgb = mcolors.colorConverter.to_rgb(fill_color)
z[:,:,:3] = rgb
z[:,:,-1] = np.linspace(0, alpha, 100)[:,None]
xmin, xmax, ymin, ymax = x.min(), x.max(), y.min(), y.max()
im = ax.imshow(z, aspect='auto', extent=[xmin, xmax, ymin, ymax],
origin='lower', zorder=zorder)
xy = np.column_stack([x, y])
xy = np.vstack([[xmin, ymin], xy, [xmax, ymin], [xmin, ymin]])
clip_path = Polygon(xy, facecolor='none', edgecolor='none', closed=True)
ax.add_patch(clip_path)
im.set_clip_path(clip_path)
ax.autoscale(True)
return line, im
# ##### Pool State vs Time:
# +
# %config InlineBackend.figure_format='svg'
# %matplotlib inline
#cut = pd.to_datetime(datetime(2022, 1, 21, 0, 0, 0), utc=True)
#Pool_0 = Pool_0[Pool_0['date'] <= cut]
fig, (ax1, ax5, ax2, ax3, ax4) = plt.subplots(5, 1, gridspec_kw={'height_ratios': [5,5,3,2,2]})
fig = plt.gcf()
dates = np.arange(0, len(Pool_0), 48)
date_ticks = [Pool_0['date'][i] for i in dates]
plot = ax1.plot(Pool_0['date'], Pool_0['LUSD'], color='blue')
plot = ax1.plot(Pool_0['date'], Pool_0['aUSD'], color='brown')
ax1.legend(('LUSD', 'aUSD'))
ax1.grid()
ax1.set(
title = 'Pool_0 Composition',
ylabel = 'Millions of $'
)
date_form = md.DateFormatter("%b-%d")
ax1.xaxis.set_major_formatter(date_form)
scale_y = 1e6
ticks_y = ticker.FuncFormatter(lambda x, pos: '{0:g}'.format(x/scale_y))
ax1.yaxis.set_major_formatter(ticks_y)
ax51 = ax5.twinx()
ax5.plot(Data['date'], Pool_0['Trading income'], color='brown')
#gradient_fill(Data.index.to_numpy(), y=Data['Curve Volume'].to_numpy(), ax=ax51)
ax51.bar(Data['date'], Data['LIQ col'], width=0.1)
ax5.set(
title='Liquidations and Trading Volume',
xlabel='Date',
ylabel='Trading Volume, USD'
)
ax51.set(
ylabel='Liquidated Collateral, ETH'
)
ax5.grid()
date_form = md.DateFormatter("%b-%d")
ax5.xaxis.set_major_formatter(date_form)
ax5.legend(('Trading Volume on Curve',), loc='upper left')
ax51.legend(('Liquidated Collateral on LQTY',), loc='center left')
ax2.bar(Pool_0['date'], Pool_0['ETH received'], color='red', label='ETH received from liquidations', width=0.1)
ax2.bar(Pool_0['date'], -Pool_0['ETH sold'], color='blue', label='ETH sold', width=0.1)
ax2.set_title('Ether Events')
ax2.set_ylabel('ETH')
#ax2.set_xlabel('Date')
ax2.legend()
ax2.grid()
ax3.bar(Data['date'], Data['BAL Reward']*Pool_0['aUSD share'], color='brown')
ax3.set_title('BAL Rewards')
ax3.set_ylabel('Rewards, BAL')
#ax3.set_xlabel('Date')
ax3.grid()
ax4.bar(Pool_0['date'], Pool_0['LQTY received'])
ax4.set_title('LQTY Rewards')
ax4.set_ylabel('LQTY')
ax4.set_xlabel('Date')
ax4.grid()
fig.set_size_inches(8, 10)
plt.tight_layout()
# -
Pool.iloc[-2, :]['LUSD'] / Data.iloc[-2, :]['LUSD in SP']
Pool_4['SP share']
# ##### Gains:
plt.plot(Pool['date'], Pool['LUSD'])
plt.plot(Pool_4['date'], Pool_4['LUSD'])
plt.twinx()
plt.plot(Data['date'], Data['LIQ col'])
plt.twinx()
plt.plot(Pool['date'], Pool['ETH sold'], color='red')
# +
##### Gains:
# +
def calculate_TVL(pool):
timeDelta = pool.iloc[-1]['date'] - pool.iloc[0]['date']
year = pd.to_timedelta(timedelta(days=365))
Init_TVL = pool.iloc[0]['LUSD'] + pool.iloc[0]['aUSD']
Final_TVL = pool.iloc[-1]['LUSD'] + pool.iloc[-1]['aUSD']
Total_Gain = Final_TVL - Init_TVL
LUSD_gain = pool.iloc[-1]['LUSD'] - pool['LUSD'][0]
aUSD_gain = (pool.iloc[-1]['aUSD'] - pool['aUSD'][0])
Total_percentage = Total_Gain/Init_TVL
LUSD_percentage = LUSD_gain/pool.iloc[0]['LUSD']
aUSD_percentage = aUSD_gain/pool.iloc[0]['aUSD']
Total_APY = Total_percentage * (year / timeDelta)
LUSD_APY = LUSD_percentage * (year / timeDelta)
aUSD_APY = aUSD_percentage * (year / timeDelta)
#print('Initial TVL: {}'.format(Init_TVL), 'Final_TVL: {}'.format(Final_TVL))
#print('Gain: {}'.format(Total_Gain))
#print('B.Protocol: {}, Sale: {}%, Reserve: {}%, DAO fees: {}%, Swaps: {}'.format(BProt, Sale, Reserve, DAO_fees, Swaps))
print('LUSD APR: {:0,.1%}'.format(LUSD_APY))
print('aUSD APR: {:0,.1%}'.format(aUSD_APY))
print('Total APR: {:0,.1%}'.format(Total_APY))
print('--- Full timeline ---')
print('No B.Protocol')
calculate_TVL(Pool)
print()
print('B.Protocol with 2% sale')
calculate_TVL(Pool_2)
print()
print('B.Protocol with 4% sale')
calculate_TVL(Pool_4)
print()
print('--- Cut timeline ---')
calculate_TVL(Pool_cut)
print()
print('B.Protocol with 0% sale')
calculate_TVL(Pool_0_cut)
print()
print('B.Protocol with 4% sale')
calculate_TVL(Pool_4)
# -
# ##### Details:
# ##### Gains
# +
# DAO gains
print('DAO gains:')
print(Pool['BAL reserve'].sum())
print(
(Pool['BAL reserve'].sum() * Data.iloc[-1, :]['BAL Price']) + \
(Pool.iloc[-1, :]['LQTY reserve'] * Data.iloc[-1, :]['LQTY Price'])
)
# +
# Trading fees:
print('--- Trading fees: ---')
print('Standalone: {}'.format(Pool['Trading income'].sum()))
print('B.Protocol: {}'.format(Pool_0['Trading income'].sum()))
print('B.Protocol (discount): {}'.format(Pool_4['Trading income'].sum()))
print('--- Token gains: ---')
def token_gains(pool, label):
print('{}:'.format(label))
print('ETH: {} (${}) \t LQTY: {} (${}) \t BAL: {} (${})'.format(
pool['ETH sold'].sum(), (pool['ETH sold'] * Data['ETH Price']).sum(),
pool['LQTY sold'].sum(), (pool['LQTY sold'] * Data['LQTY Price']).sum(),
pool['BAL sold'].sum(), (pool['BAL sold'] * Data['BAL Price']).sum()
))
token_gains(Pool, 'Standalone')
token_gains(Pool_0, 'B.Protocol')
token_gains(Pool_4, 'B.Protocol (discount)')
# +
# total gains
def total_gains(pool, label):
print('{}:'.format(label))
print('${}'.format(
(pool['ETH sold'] * Data['ETH Price']).sum() +
(pool['LQTY sold'] * Data['LQTY Price']).sum() +
(pool['BAL sold'] * Data['BAL Price']).sum() +
pool['Trading income'].sum()
))
print('--- Total gains: ---')
total_gains(Pool, 'Standalone')
total_gains(Pool_0, 'B.Protocol')
total_gains(Pool_4, 'B.Protocol (discount)')
# -
# ##### Comissions
# +
def spendings(pool, label):
print('{}:'.format(label))
print('LUSD burn: ${}'.format(pool['LUSD burn'].sum()))
print('Trading fees: ${}'.format(pool['Trading fees'].sum()))
print('--Gas--:')
print('ETH: ${} \t LQTY: ${} \t BAL: ${} \t Stability Pool: ${} \t '.format(
pool['ETH gas'].sum(),
pool['LQTY gas'].sum(),
pool['BAL gas'].sum(),
pool['SP gas'].sum()
))
print('--B.Protocol:--')
print('ETH commission: {} (${}) \t ETH discount: ${}'.format(
pool['ETH comission'].sum(), (pool['ETH comission'] * Data['ETH Price']).sum(),
pool['ETH sale'].sum()
))
spendings(Pool, 'Standalone')
spendings(Pool_0, 'B.Protocol')
spendings(Pool_4, 'B.Protocol (discount)')
# +
# total spendings
def total_spendings(pool, label):
print('{}'.format(label))
print('${}'.format(
pool['ETH gas'].sum() +
pool['LQTY gas'].sum() +
pool['BAL gas'].sum() +
pool['SP gas'].sum() +
#pool['LUSD burn'].sum() +
(pool['ETH comission'] * Data['ETH Price']).sum() +
pool['ETH sale'].sum()
))
total_spendings(Pool, 'Standalone')
total_spendings(Pool_0, 'B.Protocol')
total_spendings(Pool_4, 'B.Protocol (discount)')
# -
# # Monthly stats
# +
Pool = create_pool(Data)
Pool2 = create_pool(Data)
Res = run(Pool, Data, True, 2, 0, 7.5, True)
Res2 = run(Pool2, Data, True, 2, 0, 7.5, False)
# -
plt.plot(Res['date'], Res['LUSD'])
plt.plot(Res2['date'], Res2['LUSD'])
Data.columns
# +
d26 = pd.to_datetime(pd.Timestamp(2021, 12, 26), utc=True)
j1 = pd.to_datetime(pd.Timestamp(2022, 1, 1), utc=True)
j15 = pd.to_datetime(pd.Timestamp(2022, 1, 15), utc=True)
f1 = pd.to_datetime(pd.Timestamp(2022, 2, 1), utc=True)
f15 = pd.to_datetime(pd.Timestamp(2022, 2, 15), utc=True)
m1 = pd.to_datetime(pd.Timestamp(2022, 3, 1), utc=True)
m15 = pd.to_datetime(pd.Timestamp(2022, 3, 15), utc=True)
a1 = pd.to_datetime(pd.Timestamp(2022, 4, 1), utc=True)
a15 = pd.to_datetime(pd.Timestamp(2022, 4, 15), utc=True)
ma1 = pd.to_datetime(pd.Timestamp(2022, 5, 1), utc=True)
ma15 = pd.to_datetime(pd.Timestamp(2022, 5, 15), utc=True)
end = Data['date'].iloc[-1]
dates = [d26, j1, j15, f1, f15, m1, m15, a1, a15, ma1, ma15]
dates
d = pd.Timedelta('14d')
# +
# dates2 = [d26, j1, f1, m1, m27]
# dates2
# +
# d = pd.Timedelta('30d')
# dates3 = []
# dates3.append(d26)
# dates3 += ([j1 + i*d for i in range(200) if (j1 + i*d <= m27)])
# dates3.append(m27)
# dates3
# +
dates_ = dates
LUSD_APRs = np.zeros(len(dates_))
aUSD_APRs = np.zeros(len(dates_))
Total_APRs = np.zeros(len(dates_))
DAO = np.zeros(len(dates_))
Liqs = np.zeros(len(dates_))
BAL_rewards = np.zeros(len(dates_))
LQTY_rewards = np.zeros(len(dates_))
Trading_income = np.zeros(len(dates_))
LIQ_income = np.zeros(len(dates_))
for i, _ in enumerate(dates_):
if (i == len(dates_)-1):
break
s = dates_[i]
e = dates_[i+1]
delta = e - s
year = pd.Timedelta('365d')
Data_ = Data[(Data['date'] >= s) & (Data['date'] <= e)].reset_index(drop=True)
Pool_ = create_pool(Data_)
Res = run(Pool_, Data_, True, 2, 0, 7.5, True)
LUSD_gain = Res['LUSD'].iloc[-1] - Res['LUSD'].iloc[0]; LUSD_apr = LUSD_gain / Res['LUSD'].iloc[-1] * (year / delta)
aUSD_gain = Res['aUSD'].iloc[-1] - Res['aUSD'].iloc[0]; aUSD_apr = aUSD_gain / Res['aUSD'].iloc[-1] * (year / delta)
Total_apr = (LUSD_gain + aUSD_gain) / (Res['LUSD'].iloc[-1] + Res['aUSD'].iloc[-1]) * (year / delta)
DAO_LQTY_gain = Res['LQTY reserve'].iloc[-1] - Res['LQTY reserve'].iloc[0]; DAO_LQTY_gain = DAO_LQTY_gain * Data['LQTY Price'].iloc[-1]
DAO_BAL_gain = Res['BAL reserve'].iloc[-1] - Res['BAL reserve'].iloc[0]; DAO_BAL_gain = DAO_BAL_gain * Data['BAL Price'].iloc[-1]
LUSD_APRs[i+1] = LUSD_apr
aUSD_APRs[i+1] = aUSD_apr
Total_APRs[i+1] = Total_apr
DAO[i+1] = DAO_LQTY_gain + DAO_BAL_gain
Liqs[i+1] = Data_['LIQ col'].sum()
BAL_rewards[i+1] = (Res['BAL received'] * Data_['BAL Price']).sum()
LQTY_rewards[i+1] = (Res['LQTY received'] * Data_['LQTY Price']).sum()
Trading_income[i+1] = Res['Trading income'].sum()
LIQ_income[i+1] = (Res['ETH sold'] * Data_['ETH Price'] - Res['LUSD burn']).sum()
# print('Start date: {}; End date: {}'.format(s, e))
# print('LUSD Gain: {:,.0f}$; \t LUSD APR: {:.1f}%'.format(LUSD_gain, LUSD_apr*100))
# print('aUSD gain: {:,.0f}$; \t aUSD APR: {:.1f}%'.format(aUSD_gain, aUSD_apr*100))
# +
import matplotlib.ticker as mtick
date_form = md.DateFormatter("%b-%d")
dateticks1 = np.array(dates_) - pd.Timedelta('7d')
#date_shift = pd.Timedelta('36h')
#dateticks1[-1] += date_shift
dateticks2 = np.array(dates_[:2]) - pd.Timedelta('3d')
yticks = np.array([0, 5, 7.5, 10, 12.5, 15, 20, 25, 30, 35, 40, 45])
fig, (ax1, ax2, ax3) = plt.subplots(3, 1, gridspec_kw={'height_ratios': [4,3,2]})
ax1.bar(dateticks1[2:], Total_APRs[2:]*100, width=10, color='green', alpha=0.5)
#ax1.bar(dateticks2[:2], Total_APRs[:2]*100, width=3, color='green', alpha=0.5)
# ax11 = ax1.twinx()
# ax11.plot(Data['date'], Data['LQTY Price'], lw=2, label='LQTY Price', color='blue')
# ax11.plot(Data['date'], Data['BAL Price'], lw=2, label='BAL Price', color='brown')
# ax11.legend()
# for i, _ in enumerate(Liqs):
# s = 'Liquidated ETH:\n{:,.1f}\nBAL Rewards: {:,.0f}\nLQTY Rewards: {:,.0f}\nTrading: {:,.0f}'.format(
# Liqs[i],
# BAL_rewards[i],
# LQTY_rewards[i],
# Trading_income[i])
# t = None
# # if (i == 1):
# # t = ax1.text(dateticks2[i], Total_APRs[i]*100/2, s, ha='center', backgroundcolor='white')
# if (i > 1):
# t = ax1.text(dateticks1[i], Total_APRs[i]*100/2, s, ha='center', backgroundcolor='white')
# if (t):
# t.set_bbox(dict(facecolor='white', alpha=0.5, edgecolor='white'))
# #ax12 = ax1.twinx()
#ax12.bar(dateticks1[2:], Liqs[2:], width=5, color='red', alpha=0.25)
ax11 = ax1.twinx()
ax11.plot(Data['date'], Data['ETH Price'], lw=2, label='ETH Price', color='red', alpha=0.75)
ax11.set_ylabel('Price, $')
ax11.legend()
ax1.set_title('Pool APR ({} days period)'.format(d.days))
ax1.set_xlabel('Period')
ax1.set_ylabel('APR')
# ax11.set_ylabel('Price, $')
ax1.xaxis.set_major_formatter(date_form)
ax1.set_xticks(dates_[1:])
ax1.set_yticks(yticks)
fmt = '%.1f%%' # Format you want the ticks, e.g. '40%'
ylabels = mtick.FormatStrFormatter(fmt)
ax1.yaxis.set_major_formatter(ylabels)
ax1.grid(alpha=0.3)
shift = pd.Timedelta('3d')
#ax2.bar(dateticks1[2:]-shift, DAO[2:], width=4, color='red', label='', alpha=0.5)
#
# ax2.bar(dateticks1[2:]-shift, LQTY_rewards[2:], width=3, color='blue', label='LQTY rewards', alpha=0.5)
# ax2.bar(dateticks1[2:], BAL_rewards[2:], width=3, color='brown', label='BAL rewards', alpha=0.5)
# ax2.bar(dateticks1[2:]+shift, Trading_income[2:], width=3, color='green', label='Trading income', alpha=0.5)
ax2.bar(dateticks1[2:], LQTY_rewards[2:], width=9, color='blue', label='LQTY rewards', alpha=0.5)
ax2.bar(dateticks1[2:], BAL_rewards[2:], width=9, bottom=LQTY_rewards[2:], color='brown', label='BAL rewards', alpha=0.5)
ax2.bar(dateticks1[2:], Trading_income[2:], bottom=LQTY_rewards[2:] + BAL_rewards[2:], width=9, color='green', label='Trading income', alpha=0.5)
ax2.bar(dateticks1[2:], LIQ_income[2:], width=9, color='red', label='Liquidation income', alpha=0.5, bottom=LQTY_rewards[2:] + BAL_rewards[2:] + Trading_income[2:])
# for i, _ in enumerate(Liqs):
# bot = LIQ_income[i] + BAL_rewards[i] + LQTY_rewards[i] + Trading_income[i]
# percent = LIQ_income[i] / bot *100 if bot != 0 else 0
# s = '{:,.1f}%'.format(percent)
# t = None
# if (i == 3):
# h = bot - 350000
# else:
# h = bot + 100000
# if (i > 1):
# t = ax2.text(dateticks1[i], h, s, ha='center', backgroundcolor='white')
# if (t):
# t.set_bbox(dict(facecolor='white', alpha=0.5, edgecolor='white'))
# ax21 = ax2.twinx()
# #bottom=LQTY_rewards[2:] + BAL_rewards[2:] + Trading_income[2:]
# ax21.bar(dateticks1[2:]+shift, LIQ_income[2:], width=3, color='red', label='Liquidation income', alpha=0.5)
# ax21.legend(loc=(0.6, 0.8))
# ax21.set_ylabel('Liquidation gain, Millions of $')
ax2.legend(loc=(0.6, 0.5))
#ax2.bar(dateticks2[:2], DAO[:2], width=2, color='brown')
ax2.set_title('Pool Gains')
ax2.set_xlabel('Period')
ax2.set_ylabel('Gain, Millions of $')
ax2.xaxis.set_major_formatter(date_form)
ax2.set_xticks(dates_[1:])
ax2.grid(alpha=0.3)
ax2.set_xlim(ax1.get_xlim())
ax3.set_xlim(ax1.get_xlim())
ax3.bar(dateticks1[2:], DAO[2:], width=5, color='brown', alpha=0.75)
ax3.set_xlabel('Period')
ax3.set_ylabel('Gain, $')
ax3.set_title('DAO Gains')
ax3.grid(alpha=0.3)
ax3.xaxis.set_major_formatter(date_form)
ax3.set_xticks(dates_[1:])
# ax31 = ax3.twinx()
# ax31.set_ylabel('Price, $')
# ax31.plot(Data['date'], Data['LQTY Price'], lw=2, label='LQTY Price', color='blue')
# ax31.plot(Data['date'], Data['BAL Price'], lw=2, label='BAL Price', color='brown')
# ax31.legend()
fig.set_size_inches(10, 15)
# -
plt.plot(Data['date'], Data['LUSD in SP'])
plt.gca().xaxis.set_major_formatter(date_form)
plt.gcf().set_size_inches(8, 5)
plt.title('LUSD in Stability Pool')
plt.ylim((0, 5.5e8))
plt.xlabel('Date')
plt.ylabel('LUSD')
plt.grid()
plt.bar(Data['date'], Data['LIQ col'])
plt.gca().xaxis.set_major_formatter(date_form)
plt.gcf().set_size_inches(8, 5)
plt.title('Liquidated ETH')
plt.xlabel('Date')
plt.ylabel('ETH')
plt.grid()
|
LUSDaUsdStablePool/LUSD-aUSD Stable Swap Pool.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
browser = webdriver.Chrome('./chromedriver.exe')
browser.get('https://www.genie.co.kr/chart/top200')
html = browser.page_source
from bs4 import BeautifulSoup
soup = BeautifulSoup(html, 'html.parser')
len(soup), type(soup)
tags = soup.select('td.info')
len(tags), type(tags)
tag = tags[0]
# 곡명 : a.title, 가수이름 : a.artist
title = tag.select('a.title')
len(title), type(title)
title[0]
title[0].text.strip()
artist = tag.select('a.artist')
len(artist), type(artist)
artist[0]
artist[0].text.strip()
# [
# [title01, artist01],
# [title02, artist02],
# [title03, artist03],
# ...
# ]
# +
contents = list()
for tag in tags:
title = tag.select('a.title')
artist = tag.select('a.artist')
# print(title[0].text.strip(),'-', artist[0].text.strip())
contents.append([title[0].text.strip(), artist[0].text.strip()])
contents
# -
import pandas as pd
pd_data = pd.DataFrame(contents, columns=['title', 'artist'])
pd_data
pd_data.to_excel('./saves/genie_scraping.xls', index=False)
|
genie_scraping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="e5O1UdsY202_"
# ##### Copyright 2020 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + [markdown] id="jUW1g2_jWmBk"
# ## Measuring Signal Properties of Various Initializations
# For a random signal x ~ normal(0, 1), and a neural network denoted with f(x)=y; ensuring std(y)=1 at initialization is a common goal for popular NN initialization schemes. Here we measure signal propagation for different sparse initializations.
# + cellView="form" id="4rvDSX8FFYTI"
#@title Imports and Definitions
import numpy as np
import os
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
import gin
from rigl import sparse_utils
from rigl.rigl_tf2 import init_utils
from rigl.rigl_tf2 import utils
from rigl.rigl_tf2 import train
from rigl.rigl_tf2 import networks
from rigl.rigl_tf2 import mask_updaters
import functools
pruning_params = utils.get_pruning_params(mode='constant', final_sparsity = 0., begin_step=int(1e10))
INPUT_SHAPE = (28, 28, 3)
class Lenet5(tf.keras.Model):
def __init__(self,
input_shape,
num_classes,
activation: str,
hidden_sizes = (6, 16, 120, 84)):
super(Lenet5, self).__init__()
l = tf.keras.layers
kwargs = {'activation': activation}
filter_fn = lambda _: True
wrap_fn = functools.partial(utils.maybe_prune_layer, params=pruning_params, filter_fn=filter_fn)
self.conv1 = wrap_fn(l.Conv2D(hidden_sizes[0], 5, input_shape=input_shape, **kwargs))
self.pool1 = l.MaxPool2D(pool_size=(2, 2))
self.conv2 = wrap_fn(l.Conv2D(hidden_sizes[1], 5, input_shape=input_shape, **kwargs))
self.pool2 = l.MaxPool2D(pool_size=(2, 2))
self.flatten = l.Flatten()
self.dense1 = wrap_fn(l.Dense(hidden_sizes[2], **kwargs))
self.dense2 = wrap_fn(l.Dense(hidden_sizes[3], **kwargs))
self.dense3 = wrap_fn(l.Dense(num_classes, **kwargs))
self.build((1,)+input_shape)
def call(self, inputs):
x = inputs
results = {}
for l_name in ['conv1', 'pool1', 'conv2', 'pool2', 'flatten', 'dense1', 'dense2', 'dense3']:
x = getattr(self, l_name)(x)
results[l_name] = x
return results
def get_mask_random_numpy(mask_shape, sparsity):
"""Creates a random sparse mask with deterministic sparsity.
Args:
mask_shape: list, used to obtain shape of the random mask.
sparsity: float, between 0 and 1.
Returns:
numpy.ndarray
"""
all_ones = np.abs(np.ones(mask_shape))
n_zeros = int(np.floor(sparsity * all_ones.size))
rand_vals = np.random.uniform(size=mask_shape, high=range(1,mask_shape[-1]+1))
randflat=rand_vals.flatten()
randflat.sort()
t = randflat[n_zeros]
all_ones[rand_vals<=t] = 0
return all_ones
def create_convnet(sparsity=0, weight_init_method = None, scale=2, method='fanin_normal'):
model = Lenet5(INPUT_SHAPE, num_classes, 'relu')
if sparsity > 0:
all_masks = [layer.pruning_vars[0][1] for layer in model.layers if isinstance(layer, utils.PRUNING_WRAPPER)]
for mask in all_masks:
new_mask = tf.cast(get_mask_random_numpy(mask.shape, sparsity), dtype=mask.dtype)
mask.assign(new_mask)
if weight_init_method:
all_weights = [layer.pruning_vars[0][0] for layer in model.layers if isinstance(layer, utils.PRUNING_WRAPPER)]
for mask, param in zip(all_masks, all_weights):
if weight_init_method == 'unit':
new_init = init_utils.unit_scaled_init(mask, method=method, scale=scale)
elif weight_init_method == 'layer':
new_init = init_utils.layer_scaled_init(mask, method=method, scale=scale)
else:
raise ValueError
param.assign(new_init)
return model
# + [markdown] id="fkZ_GNjyYYqZ"
# Here we demonstrate how we can calculate the standard deviation of random noise at initialization for `layer-wise` scaled initialization of Liu et. al.
# + id="NsmPRCuZnxDA"
# Let's create a 95% sparse Lenet-5.
model = create_convnet(sparsity=0.95, weight_init_method='layer', scale=2, method='fanin_normal')
# Random input signal
random_input = tf.random.normal((1000,) + INPUT_SHAPE)
output_dict = model(random_input)
all_stds = []
for k in ['dense1', 'dense2', 'dense3']:
out_dim = output_dict[k].shape[-1]
stds = np.std(np.reshape(output_dict[k], (-1,out_dim)),axis=0)
all_stds.append(stds)
print('Mean deviation per neuron', np.mean(np.concatenate(all_stds, axis=0)))
print('Mean deviation per output neuron', np.mean(all_stds[-1]))
print('Deviation at output', np.std(random_input))
# + [markdown] id="l3ttY88rYovo"
# Now we define the code above as a function and use it on a grid to plot signal propagation at different sparsities.
# + executionInfo={"elapsed": 320, "status": "ok", "timestamp": 1613388807790, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": -180} id="4rfMGKciOOHf"
def propagate_signal(sparsity, init_method, batch_size=500):
model = create_convnet(sparsity=sparsity, weight_init_method=init_method)
random_input = tf.random.normal((batch_size,) + INPUT_SHAPE)
# print(np.mean(random_input), np.std(random_input))
output_dict = model(random_input)
out_std = np.std(output_dict['dense3'])
all_stds = []
for k in ['dense1', 'dense2', 'dense3']:
out_dim = output_dict[k].shape[-1]
stds = np.std(np.reshape(output_dict[k], (-1,out_dim)),axis=0)
all_stds.append(stds)
meanstd = np.mean(np.concatenate(all_stds, axis=0))
return meanstd, out_std
# + id="F1rNPLXk7Ins"
import itertools, collections
import numpy as np
all_results = collections.defaultdict(dict)
N_EXP = 3
for s in np.linspace(0.8,0.98,5):
print(s)
for method, name in zip((None, 'unit', 'layer'), ('Masked Dense', 'Ours', 'Scaled-Init')):
all_results[name][s] = [propagate_signal(s, method) for _ in range(N_EXP)]
# + id="Sbjc7LxpVGl0"
import matplotlib.pyplot as plt
for k, v in all_results.items():
# if k == 'Masked Dense':
# continue
x = sorted(v.keys())
y = [np.mean([vv[1] for vv in v[kk]])+1e-5 for kk in x]
plt.plot(x, y, label=k)
plt.hlines(y=1, color='r', xmin=0, xmax=1)
plt.yscale('log')
plt.title('std(output)')
plt.legend()
plt.show()
for k, v in all_results.items():
# if k == 'Masked Dense':
# continue
x = sorted(v.keys())
y = [np.mean([vv[0] for vv in v[kk]])+1e-5 for kk in x]
plt.plot(x, y, label=k)
plt.yscale('log')
plt.hlines(y=1, color='r', xmin=0, xmax=1)
plt.title('mean(std_per_neuron)')
plt.legend()
plt.show()
|
rigl/rigl_tf2/colabs/MnistProp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Neste tutorial, vamos tentar entender a lógica subjacente à inferência por reamostragem *bootstrapping*. Trata-se de uma apresentação muito introdutória.
import numpy as np
import matplotlib.pyplot as plt
# Cria parâmetros de uma distribuição de escores em um teste de inteligência
#
# **escores ~ *N*(100, 15)**
# +
# parâmetros da distribuição
media = 100
dp = 15
tamanho_amostral = 15
# definindo uma semente para replicabilidade
np.random.seed(1)
amostra = np.random.normal(100, 15, size = tamanho_amostral)
# imprime escores da amostra, média e desvio padrão
print(amostra)
print(f"Média: {np.mean(amostra):.3f}")
print(f"DP: {np.std(amostra):.3f}")
# -
# Vamos ver a distribuição dessa amostra
# +
plt.hist(amostra, color = "pink")
plt.title("Distribuição Amostral", fontsize = 20)
plt.xlabel("QI", fontsize = 16)
plt.ylabel("Frequência", fontsize = 16)
plt.savefig('006 - Bootstrapping.jpg', dpi = 600, bbox_inches = "tight")
plt.show()
# -
# a distribuição acima é toda estranha. Agora, vamos tratar a nossa amostra de 15 participantes como uma **minipopulação**. A partir dela, extraíremos uma amostra de 15 casos com reposição. Em outras palavras, sortearemos aleatoriamente um participante, anotaremos seu escore, devolveremos ele para a amostra, sortearemos outro, até completarmos um tamanho amostral igual ao de nossa amostra original.
#
# Por conta desse processo probabilístico, é possível que alguns participantes não sejam representados nessa nova amostra, enquanto outros podem ser representados 2 ou mais vezes. Nessa nova amostra, podemos calcular a média e guardar esse valor.
#
# Mais importante, podemos repetir esse procedimento milhares de vezes (digamos, 5 mil). Ao final, teremos 5 mil médias de amostras *bootstrap*. Vamos lá?
# +
# define o número de simulações
n_simulacoes = 5_000 # modifique este valor, caso queira
# lista armazenará as médias de cada uma das 5 mil simulações
medias_das_amostras_bootstrap = list()
# realiza as n_simulacoes simulacoes
for simulacao in range(n_simulacoes):
# amostragem com reposição: participantes são sorteados da amostra e depois devolvidos para novo sorteio
amostra_bootstrap = np.random.choice(amostra, size = tamanho_amostral, replace = True)
# salva a média da i-ésima amostra bootstrap em uma lista
medias_das_amostras_bootstrap.append(np.mean(amostra_bootstrap))
print("Terminamos a simulação!")
# -
# Após terminarmos a simulação, vamos fazer isso e ver como é a distribuição dessas 5 mil médias?
plt.hist(medias_das_amostras_bootstrap, color = "pink")
plt.title("Distribuição de Médias Amostrais\ndas Amostras Bootstrap", fontsize = 20)
plt.xlabel("QI", fontsize = 16)
plt.ylabel("Frequência", fontsize = 16)
plt.show()
# Note que a distribuição amostral de médias das amostras *bootstrap* tem distribuição normal, mesmo que os dados originalmente fossem todos zoados. Vamos ver a média e o DP dessa distribuição amostral de médias das amostras *bootstrap*...
# +
print(f"Média: {np.mean(medias_das_amostras_bootstrap):.3f}")
print(f"DP: {np.std(medias_das_amostras_bootstrap):.3f}")
# print(f"EPM: {np.std(amostra / np.sqrt(tamanho_amostral)):.3f}")
# -
# Podemos usar os valores acima para calcular intervalos de confiança. Você pode se perguntar: "mas por que não calcular ICs do jeito tradicional?"
#
# O problema dos ICs tradicionais em nosso exemplo é que, pelo tamanho amostral ser pequeno, isso significa que a média amostral tende a variar mais de amostragem para amostragem. Isso quer dizer que há mais chances de que a média da nossa amostra, por azar, esteja mais distante da verdadeira média populacional. Os intervalos de confiança *bootstrapping*, que não se baseiam na teoria normal clássica, tendem a levar a inferências mais precisas nessas situações.
#
# Primeiro, vamos calcular o intervalo de confiança para a média populacional com base na nossa amostra.
# Intervalos de confiança da amostra
print("Intervalos de confiança da amostra")
limite_inferior = np.mean(amostra) - 1.96 * np.std(amostra) / np.sqrt(10)
limite_superior = np.mean(amostra) + 1.96 * np.std(amostra) / np.sqrt(10)
print(f"IC 95% = [{limite_inferior:.3f}, {limite_superior:.3f}]")
# Agora, vamos repetir o procedimento, mas calculando o IC *bootstrap* percentílico.
# Intervalos de confiança percentílicos das amostras bootstrap
# produz ICs mais estreitos (i.e., mais precisos) que aqueles baseados na teoria normal
print("Intervalos de confiança percentílicos das amostras bootstrap")
limite_inferior_boot = np.percentile(medias_das_amostras_bootstrap, q = 2.5)
limite_superior_boot = np.percentile(medias_das_amostras_bootstrap, q = 97.5)
print(f"IC 95% percentílico = [{limite_inferior_boot:.3f}, {limite_superior_boot:.3f}]")
|
006 - Bootstrapping/006 - Bootstrapping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parse copyright data
import pandas as pd
import xml.etree.ElementTree as ET
import glob
# +
entries = []
for folderyear in range(1923, 1970):
ctr = 0
xmlfiles = glob.glob('/Users/tunder/work/copyright/xml/' + str(folderyear) + '/*.xml')
for afilepath in xmlfiles:
tree = ET.parse(afilepath)
root = tree.getroot()
for child in root.findall('copyrightEntry'):
authors = []
for author in child.findall('author'):
for authorname in author.findall('authorName'):
if authorname.text is not None:
authors.append(authorname.text)
for title in child.findall('title'):
if title.text is not None:
title = title.text
else:
title = ''
for regdate in child.findall('regDate'):
try:
date = int(regdate.attrib['date'][0:4])
except:
date = 0
print('exception')
if len(authors) > 0 and len(title) > 0 and date > 0:
row = dict()
row['authors'] = '|'.join(authors)
row['title'] = title
row['copydate'] = date
entries.append(row)
ctr += 1
print(folderyear, ctr, len(entries))
# -
df = pd.DataFrame(entries)
df.head()
df.to_csv('/Users/tunder/work/copyright/tabularcopyright.tsv', sep = '\t', index = False)
df = pd.read_csv('/Users/tunder/work/copyright/tabularcopyright.tsv', sep = '\t', low_memory = False)
import pandas as pd
from difflib import SequenceMatcher
import re
titles = pd.read_csv('../noveltmmeta/metadata/titlemeta.tsv', sep = '\t', low_memory = False)
titles = titles.loc[titles.latestcomp > 1920, : ]
titles.shape
# +
def getinitial(astring):
if pd.isnull(astring):
return('xx')
astring = re.sub(r'[^\w\s]', '', astring.lower())
if len(astring) < 1:
return 'xx'
elif len(astring) == 1:
return astring.lower()
else:
return astring.lower()[0:2]
titles['initial'] = titles['author'].map(getinitial)
titles.head()
# +
def get_ratio(stringA, stringB):
'''
A generic function to get fuzzy similarity between two strings.
'''
m = SequenceMatcher(None, stringA, stringB)
thefilter = m.real_quick_ratio()
if thefilter < 0.75:
return thefilter
else:
return m.ratio()
def title_compare(stringA, stringB):
'''
When searching for the similarity between two titles, we use this modified comparison,
which gives a boost to the similarity when it applies to two long strings, but also
permits relatively high similarity in cases where one string is much longer than
the other and only the overlapping starts are similar. This is useful because there
are often variant fiction titles like
Shane
and
Shane: A Story of the American West.
'''
stringA = stringA.lower().replace('the ', 'x') # we replace 'the' because it's a word that
stringB = stringB.lower().replace('the ', 'x') # is relatively long for how common it is,
# and also quite likely to appear at the start
# of a title, producing false matches
minlen = min(len(stringA), len(stringB))
maxlen = max(len(stringA), len(stringB))
diffpenalty = (maxlen - minlen) / 300
if minlen > 3:
stringA = stringA[0: minlen]
stringB = stringB[0: minlen]
if minlen < 25:
lendiscount = 1.2 - (((25 - minlen) ** 1.1) / 100)
else:
lendiscount = 1.2
m = SequenceMatcher(None, stringA, stringB)
thefilter = m.quick_ratio()
if thefilter < 0.5:
return round((thefilter * lendiscount) - diffpenalty, 4)
else:
return round((m.ratio() * lendiscount) - diffpenalty, 4)
def given_name_similarity(namesA, namesB):
initialsA = set([x[0] for x in namesA])
initialsB = set([x[0] for x in namesB])
overlap = len(initialsA.intersection(initialsB))
difference = len(initialsA.symmetric_difference(initialsB))
surplus = overlap - difference
if surplus < 1:
return surplus * .04
else:
for name in namesA:
if len(name) > 2 and name in namesB:
surplus += 1
return surplus * .04
# +
matches = []
matchedalready = set()
ctr = 0
for year in range(1922, 1941):
print('year', year)
dfslice = df.loc[df.copydate == year, : ]
titlesneardate = titles.loc[(titles.latestcomp > (year -3)) & (titles.latestcomp < (year + 30)), : ]
blocks = dict()
for idx, row in dfslice.iterrows():
ctr += 1
if ctr % 1000 == 1:
print(ctr)
title = row['title']
if not pd.isnull(title):
title = title.lower()
else:
continue
title = re.sub(r'[^\w\s]', '', title) # remove punctuation
author = row['authors']
if not pd.isnull(author):
author = author.lower().split('|')[0]
else:
continue
author = re.sub(r'[^\w\s]', '', author)
names = author.split()
if len(names) > 0:
surname = names[0]
last_initial = getinitial(surname)
else:
continue
if len(names) > 1:
given_names = names[1: ]
else:
given_names = []
if last_initial in blocks:
block = blocks[last_initial]
else:
block = titlesneardate.loc[titlesneardate['initial'] == last_initial, : ]
blocks[last_initial] = block
for idx, matchrow in block.iterrows():
m_author = matchrow['author']
if not pd.isnull(m_author):
m_author = m_author.lower()
else:
continue
m_author = re.sub(r'[^\w\s]', '', m_author)
m_names = m_author.split()
if len(m_names) > 0:
m_surname = m_names[0]
else:
continue
surname_match = get_ratio(surname, m_surname)
if surname_match < .85:
continue
if len(m_names) > 1:
m_given_names = m_names[1: ]
else:
m_given_names = []
if len(m_given_names) > 0:
given_supp = given_name_similarity(given_names, m_given_names)
else:
given_supp = 0
m_title = matchrow['shorttitle']
if pd.isnull(m_title):
continue
m_title = m_title.lower()
m_title = re.sub(r'[^\w\s]', '', m_title)
title_match = title_compare(title, m_title)
if (surname_match + given_supp) > .98 and title_match > .95:
matchrec = dict()
matchrec['copy_author'] = author
matchrec['copy_title'] = title
matchrec['hathi_author'] = m_author
matchrec['hathi_title'] = m_title
matchrec['surname_match'] = surname_match
matchrec['given_supp'] = given_supp
matchrec['title_match'] = title_match
matchrec['docid'] = matchrow['docid']
matchrec['copy_date'] = row['copydate']
matchrec['hathi_date'] = matchrow['latestcomp']
matches.append(matchrec)
matchlen = len(matches)
if matchlen % 100 == 1:
print(len(matches), 'matches')
matchedalready.add(matchrec['docid'])
# titles = titles.loc[~titles.docid.isin(matchedalready)]
print('Matched so far: ', len(matchedalready))
matched_df = pd.DataFrame(matches)
outdf = matched_df[['copy_author', 'copy_title', 'hathi_author', 'hathi_title', 'title_match', 'surname_match', 'given_supp', 'docid', 'copy_date', 'hathi_date']]
outdf.to_csv('copymatches_1922-40.tsv', sep= '\t', index = False)
# -
outdf = matched_df[['author', 'title', 'm_author', 'm_title', 'title_match', 'surname_match', 'given_supp', 'docid', 'copydate', 'hathidate']]
outdf.to_csv('copymatches_27-40.tsv', sep= '\t', index = False)
matched_df.head()
# +
from IPython.display import display
ctr = 0
for idx, df in matched_df.groupby('docid'):
if df.shape[0] > 1:
display(df)
ctr += 1
if ctr > 8:
break
# -
|
dataconstruction/ParseCopyright.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Steps for training detectron2 on custom data
# ### Loading necessary libraries
# +
import h5py
import os
from matplotlib import pyplot as plt
import cv2
import numpy as np
import pandas as pd
import random
import time
# %config Completer.use_jedi = False
os.chdir("C:/Users/admin/Documents/Python Scripts/Soma-Segmentation/train")
from utils import ImageJ2COCO
from config import configuration
from segmentation_predictor import predict_img
#from train import run_train
from detectron2.data.datasets import register_coco_instances
from detectron2.engine import DefaultTrainer
from detectron2.config import get_cfg
from detectron2 import model_zoo
from detectron2.data import MetadataCatalog, DatasetCatalog
import detectron2.data.transforms as T
from detectron2.data import DatasetMapper # the default mapper
from detectron2.data import build_detection_train_loader, build_detection_test_loader
from detectron2.evaluation import COCOEvaluator, inference_on_dataset
from detectron2.utils.visualizer import ColorMode
from detectron2.utils.visualizer import Visualizer
from detectron2.engine import DefaultPredictor
# -
# ### Registering data
# +
# if data already is in COCO style
register_coco_instances("train", {}, "I:/Sina/Medical report segmentation/publaynet/train.json",
"I:/Sina/Medical report segmentation/publaynet/train")
register_coco_instances("val", {}, "I:/Sina/Medical report segmentation/publaynet/val.json",
"I:/Sina/Medical report segmentation/publaynet/val")
#MetadataCatalog.get("val").set(thing_classes=["title", "text", "figure", "table", "list"])
#MetadataCatalog.get("train").set(thing_classes=["title", "text", "figure", "table", "list"])
# +
# for custom data with custom written dataloader
# dataloader for train dataset (this custom written data loader is able to load multple datasets from different location)
img2coco_train = ImageJ2COCO(image_path=["G:/Data & Analysis/150802_p3.5_gcamp6/Data/150802_p3.5_gcamp6 H5/150802_a3_1h40min.h5",
"G:/Data & Analysis/150802_p3.5_gcamp6/Data/150802_p3.5_gcamp6 H5/150802_a3_1h40min.h5"],
label_path=["G:/Data & Analysis/150802_p3.5_gcamp6/Analysis/ROIS and Inside Activities/RoiSetFull.zip",
"G:/Data & Analysis/150802_p3.5_gcamp6/Analysis/ROIS and Inside Activities/RoiSetFull.zip"],
output_path="C:/Users/admin/Desktop/test",
start_index=[10000, 12000],
end_index=[12000, 14000],
image_nr=[1400, 1500],
id_starter=[1, 2000],
min_intensity = [100, 100],
max_intensity = [4000, 3000],
image_scale = [(128,128), (128,128)],
image_rotation = [-90, -90],
key = ["GroupHierarchy.Groups.Datasets",
"GroupHierarchy.Groups.Datasets"])
# dataloader for validation dataset (this custom written data loader is able to load multple datasets from different location)
img2coco_val = ImageJ2COCO(image_path=["G:/Data & Analysis/150802_p3.5_gcamp6/Data/150802_p3.5_gcamp6 H5/150802_a3_1h40min.h5",
"G:/Data & Analysis/150802_p3.5_gcamp6/Data/150802_p3.5_gcamp6 H5/150802_a3_1h40min.h5"],
label_path=["G:/Data & Analysis/150802_p3.5_gcamp6/Analysis/ROIS and Inside Activities/RoiSetFull.zip",
"G:/Data & Analysis/150802_p3.5_gcamp6/Analysis/ROIS and Inside Activities/RoiSetFull.zip"],
output_path="C:/Users/admin/Desktop/test",
start_index=[1000, 1200],
end_index=[20000, 40000],
image_nr=[800, 800],
id_starter=[3000, 4000],
min_intensity = [100, 100],
max_intensity = [4000, 3000],
image_scale = [(128,128), (128,128)],
image_rotation = [-90, -90],
key = ["GroupHierarchy.Groups.Datasets",
"GroupHierarchy.Groups.Datasets"])
# register train data set
DatasetCatalog.register("train", img2coco_train.transform)
MetadataCatalog.get("train").set(thing_classes=["soma"]) # define classes as well
# register validation data set
DatasetCatalog.register("val", img2coco_val.transform)
MetadataCatalog.get("val").set(thing_classes=["soma"]) # define classes as well
metadata = MetadataCatalog.get("train")
# -
# ### Check visually the data load step
# +
# prepare data
mydata = DatasetCatalog.get("train")
mydata_metdata = MetadataCatalog.get("train")
# +
# plot
# %matplotlib inline
for d in random.sample(mydata, 1):
im = cv2.imread(d["file_name"])
v = Visualizer(im[:, :, ::-1],
metadata=mydata_metdata,
scale=1,
instance_mode=ColorMode.IMAGE_BW # remove the colors of unsegmented pixels. This option is only available for segmentation models
)
out = v.draw_dataset_dict(d)
plt.imshow(out.get_image()[:, :, ::-1])
plt.show()
# -
# ### Model configuration
# configuration file
cfg = configuration(num_classes=5,
train_output_path="C:/Users/admin/Desktop/test/out2",
min_image_size=125,
image_per_batch=1,
max_iter=50000,
base_lr = 0.001,
model_weights=False, #"C:/Users/admin/Desktop/test/out/model_final.pth", # if you have another weights give that one
validation=True) # if you have validation turn it to True
# ### Model training
# #### Default params
# +
# start training
trainer = DefaultTrainer(cfg)
trainer.resume_or_load(resume=False)
trainer.train()
# -
# #### Custom data loader (with data augmentation)
class CustomTrainer(DefaultTrainer):
# trainer
@classmethod
def build_train_loader(cls, cfg):
dataloader = build_detection_train_loader(cfg,
mapper=DatasetMapper(cfg, is_train=True, augmentations=[
T.Resize((800, 800)),
T.Resize((100, 100)),
T.RandomBrightness(intensity_min=0.1, intensity_max=4),
T.Resize((200, 200)),
T.RandomBrightness(intensity_min=0.5, intensity_max=2),
T.RandomContrast(intensity_min=0.5, intensity_max=2),
T.RandomCrop(crop_type="relative", crop_size=(0.8, 0.8)),
T.RandomFlip(),
T.RandomFlip(vertical=True, horizontal=False)
]))
return dataloader
#evaluator
#@classmethod
#def build_evaluator(cls, cfg):
# return COCOEvaluator("val", cfg, True, os.path.join(cfg.OUTPUT_DIR,"inference"))
# +
# https://www.kaggle.com/dhiiyaur/detectron-2-compare-models-augmentation
#https://ortegatron.medium.com/training-on-detectron2-with-a-validation-set-and-plot-loss-on-it-to-avoid-overfitting-6449418fbf4e
# first time install shapely
trainer = CustomTrainer(cfg)
trainer.resume_or_load(resume=False)
trainer.train()
# -
# ### Model evaluator
evaluator = COCOEvaluator("val", cfg, False, output_dir="C:/Users/admin/Desktop/test/out2")
val_loader = build_detection_test_loader(cfg, "val")
inference_on_dataset(trainer.model, val_loader, evaluator)
# ### Model prediction
# +
# on single image
# save result's image
save_img = True
# get dir path to image
img_path = "C:/Users/admin/Desktop/test/50.png"
# model weights (after training)
model_weights_path = "C:/Users/admin/Desktop/test/out2/model_final.pth"
# path for saving final results
save_path = "C:/Users/admin/Desktop/test/test"
# get configuration file same as used during training
cfg = configuration(num_classes=1,
train_output_path="C:/Users/admin/Desktop/test/out2",
min_image_size=125,
image_per_batch=1,
max_iter=150,
model_weights=model_weights_path,
validation=True)
# initialize main dataframe
df = pd.DataFrame()
df = predict_img(cfg=cfg, img_path=img_path, save_path=save_path,
img_save=save_img, df_save=False, score_thresh=0.8)
# save main dataframe
print("saving data frame!")
df.to_csv(os.path.join(save_path, "main_results") + ".csv", index=False)
# +
# on multiple images
# save result's image
save_img = False
# get dir path to images
imgs_path = "C:/Users/admin/Desktop/test/test_images"
# get images info
imgs = os.listdir(imgs_path)
# model weights (after training)
model_weights_path = "C:/Users/admin/Desktop/test/out2/model_final.pth"
# path for saving final results
save_path = "C:/Users/admin/Desktop/test/test"
# get configuration file same as used during training
cfg = configuration(num_classes=1,
train_output_path="C:/Users/admin/Desktop/test/out2",
min_image_size=125,
image_per_batch=1,
max_iter=150,
model_weights=model_weights_path,
validation=True)
# initialize main dataframe
df = pd.DataFrame()
# claculate time
start_time = time.clock()
# starting looping results
for img in imgs:
df = df.append(predict_img(cfg=cfg, img_path=os.path.join(imgs_path, img), save_path=save_path,
img_save=save_img, df_save=False, score_thresh=0.7), ignore_index=True)
print(f'execution time: {np.rint(time.clock() - start_time)} seconds')
# save main dataframe
print("saving data frame!")
df.to_csv(os.path.join(save_path, "main_results") + ".csv", index=False)
# -
|
train/soma_segmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Now that we've created a predictive model we can apply this model to a new set of molecules. In many cases, we will build a predictive model based on literature data, then apply that model to a set of molecules that we want to screen. The molecules we want to screen may come from an internal database or from a commercially available screening collection. As an example, we will use the predictive model we created to screen a small sample of 100,000 compounds from the ZINC database.
#
# One potential source of difficulty when carrying out a virtual screen is the presence of molecules which have the potential to interfere with biological assays. Over the last 25 years, many groups have developed sets of computational filters to identify potentially reactive or problematic molecules. Several of these rule sets, which are encoded as SMARTS strings have been collected by the group that curates the ChEMBL database. These rule sets have been made avaiable through a Python script called rd_filters.py. In this example, we will use rd_filters.py to identify potentially problematic molecules in our set of 100,000 molecules from the ZINC database.
#
# **IMPORTANT** In order to run this notebook you must install the rdfiters script which can be downloaded from https://github.com/PatWalters/rd_filters the script rd_filters must be in your path.
#
# The rd_filters script can be called as follows.
# !rd_filters -h
# To call the script on our input file, which is called zinc_100k.smi we can specify the input file and a prefix for output file names.
# !rd_filters filter --in zinc_100k.smi --prefix zinc
# The output above indicates the following.
# * The script runs in parallel across multiple cores, the number of cores can be selected with the "-np" flag
# * The script is using the "Inpharmatica" set of alerts. It has 7 other alert sets available. Please see the rd_filters.py documentaiton for more information
# * SMILES for the molecules passing the filters was written to a file called zinc.smi. We will use this as the input when we use the predictive model.
# * Detailed information on which compounds triggered particular structural alerts was written to a file called zinc.csv.
# * 68% of the structures passed the filters
#
# It is informative to take a look at the reasons molecules were rejected. This can let us know whether we need to adjust any of the filters.
import pandas as pd
df = pd.read_csv("zinc.csv")
df.head()
# We can use the Counter class from the Python "collections" library to identify which filters were responsible for removing the largest number of molecules.
from collections import Counter
count_list = list(Counter(df.FILTER).items())
count_df = pd.DataFrame(count_list,columns=["Rule","Count"])
count_df.sort_values("Count",inplace=True,ascending=False)
count_df.head()
# The largest number of molecules (19,330) were rejected because they contained a 1,2 dicarbonyl group. Molecules of this type have a tendency to act a Michael Acceptors and may reactive with nucleophilic protein residues such as serine and cysteine. Let's take a look at a few of these molecules.
smiles_list = df[df.FILTER == "Filter41_12_dicarbonyl > 0"].SMILES[:10]
# +
from rdkit import Chem
from rdkit.Chem import Draw
mol_list = [Chem.MolFromSmiles(x) for x in smiles_list]
dicarbonyl = Chem.MolFromSmarts('*C(=O)C(=O)*')
match_list = [mol.GetSubstructMatch(dicarbonyl) for mol in mol_list]
Draw.MolsToGridImage(mol_list,highlightAtomLists=match_list,molsPerRow=5)
# -
# As we can see above, the molecules do indeed have dicarbonyl groups. If we wanted to we could similarly evaluate other filters.
|
Chapter11/chapter_11_03_filtering_chemical_libraries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## EDA & Cleaning: Exploring continuous features
#
# Using the Titanic dataset from [this](https://www.kaggle.com/c/titanic/overview) Kaggle competition.
#
# This dataset contains information about 891 people who were on board the ship when departed on April 15th, 1912. As noted in the description on Kaggle's website, some people aboard the ship were more likely to survive the wreck than others. There were not enough lifeboats for everybody so women, children, and the upper-class were prioritized. Using the information about these 891 passengers, the challenge is to build a model to predict which people would survive based on the following fields:
#
# - **Name** (str) - Name of the passenger
# - **Pclass** (int) - Ticket class
# - **Sex** (str) - Sex of the passenger
# - **Age** (float) - Age in years
# - **SibSp** (int) - Number of siblings and spouses aboard
# - **Parch** (int) - Number of parents and children aboard
# - **Ticket** (str) - Ticket number
# - **Fare** (float) - Passenger fare
# - **Cabin** (str) - Cabin number
# - **Embarked** (str) - Port of embarkation (C = Cherbourg, Q = Queenstown, S = Southampton)
#
# **This section focuses on exploring the `Pclass`, `Age`, `SibSp`, `Parch`, and `Fare` features.**
# ### Read in data
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
titanic_df = pd.read_csv("../Data/titanic.csv")
titanic_df.head()
# +
# drop all categorial features
cat_features = ['PassengerId', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked']
titanic_df.drop(cat_features, axis=1, inplace=True)
# -
titanic_df.head()
# ### Explore continuous features
titanic_df.describe()
# to see the overview glance of survived / non survied group
titanic_df.groupby('Survived').mean()
titanic_df['Age'].isnull()
# see the overview glance of missing age group and non missing age group
titanic_df.groupby(titanic_df['Age'].isnull()).mean()
# ### Plot continuous features
titanic_df[titanic_df['Survived'] == 0]['Age']
# #### seaborn distribution plot
for i in ['Age', 'Fare']:
died = list(titanic_df[titanic_df['Survived'] == 0][i].dropna())
survived = list(titanic_df[titanic_df['Survived'] == 1][i].dropna())
xmin = min(min(died), min(survived))
xmax = max(max(died), max(survived))
width = (xmax - xmin) / 40
sns.distplot(died, color='r', kde=False, bins=np.arange(xmin, xmax, width))
sns.distplot(survived, color= 'g', kde=False, bins=np.arange(xmin, xmax, width))
plt.legend(['Did not survived', 'Survived'])
plt.title('Overlaid Histogram for {}'.format(i))
plt.show()
# #### using seaborn categorial plot to explore more
# +
for i, col in enumerate(['Pclass', 'SibSp', 'Parch']):
plt.figure(i);
sns.catplot(x=col, y='Survived', data=titanic_df , kind='point', aspect=2); # kind is what kind of graph, aspect is for how big the point is
# in the below charts, the shorter the line is the more confidence we are.
# some lines are longer because we may have less or missing data and don't have much confidence in it.
# -
# ### as both Parch and SpSib have similarity, we will merge both data and explore a whole
titanic_df['family_cnt'] = titanic_df['SibSp'] + titanic_df['Parch']
sns.catplot(x='family_cnt', y='Survived', data=titanic_df, kind='point', aspect=2)
# #### we can see that the more family members, the less likely that person will survive.
|
ML - Applied Machine Learning Foundation/02.Exploratory Data Analysis and Data Cleaning/01.EDA & Cleaning - Exploring continuous features.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ATACseq_BROCKMAN3]
# language: python
# name: conda-env-ATACseq_BROCKMAN3-py
# ---
# `ls ./bed/ | grep bed | cut -d. -f1 > cells_input.txt`
# `ls ./kmers/ | grep freq.gz | cut -d. -f1 > cells_output.txt`
import pandas as pd
import numpy as np
import os
import shutil
df_input = pd.read_csv('./cells_input.txt',sep='\t',header=None,names=['cells'])
df_input.index.name = None
df_output = pd.read_csv('./cells_output.txt',sep='\t',header=None,names=['cells'])
df_output.index.name = None
df_input.head()
df_output.head()
missing_cells = list(set(df_input['cells'])-set(df_output['cells']))
if not os.path.exists('bed_missing'):
os.mkdir('bed_missing')
for x in missing_cells:
print(x+'.bed')
shutil.copy2('./bed/'+x+'.bed', './bed_missing/')
# `bsub < get_missing_gapped_kmers.sh`
|
Synthetic_Data/BoneMarrow_cov1000/run_methods/BROCKMAN_preprocess/find_missing_files.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# [](https://github.com/awslabs/aws-data-wrangler)
#
# # 7 - Databases (Redshift, MySQL and PostgreSQL)
#
# [Wrangler](https://github.com/awslabs/aws-data-wrangler)'s Database module (`wr.db.*`) has two mainly functions that tries to follow the Pandas conventions, but add more data type consistency.
#
# - [wr.db.to_sql()](https://aws-data-wrangler.readthedocs.io/en/latest/stubs/awswrangler.db.to_sql.html#awswrangler.db.to_sql)
#
# - [wr.db.read_sql_query()](https://aws-data-wrangler.readthedocs.io/en/latest/stubs/awswrangler.db.read_sql_query.html#awswrangler.db.read_sql_query)
# +
import awswrangler as wr
import pandas as pd
df = pd.DataFrame({
"id": [1, 2],
"name": ["foo", "boo"]
})
# -
# ### Creating an engine (SQLAlchemy Engine)
#
# The Wrangler offers basically three diffent ways to create a SQLAlchemy engine.
#
# 1 - [wr.catalog.get_engine()](https://aws-data-wrangler.readthedocs.io/en/latest/stubs/awswrangler.catalog.get_engine.html#awswrangler.catalog.get_engine): Get the engine from a Glue Catalog Connection.
#
# 2 - [wr.db.get_engine()](https://aws-data-wrangler.readthedocs.io/en/latest/stubs/awswrangler.db.get_engine.html#awswrangler.db.get_engine): Get the engine from primitives values (host, user, password, etc).
#
# 3 - [wr.db.get_redshift_temp_engine()](https://aws-data-wrangler.readthedocs.io/en/latest/stubs/awswrangler.db.get_redshift_temp_engine.html#awswrangler.db.get_redshift_temp_engine): Get redshift engine with temporary credentials.
eng_postgresql = wr.catalog.get_engine("aws-data-wrangler-postgresql")
eng_mysql = wr.catalog.get_engine("aws-data-wrangler-mysql")
eng_redshift = wr.catalog.get_engine("aws-data-wrangler-redshift")
# ## Raw SQL queries (No Pandas)
with eng_postgresql.connect() as con:
for row in con.execute("SELECT 1"):
print(row)
# ## Loading data to Database
wr.db.to_sql(df, eng_postgresql, schema="public", name="tutorial", if_exists="replace", index=False) # PostgreSQL
wr.db.to_sql(df, eng_mysql, schema="test", name="tutorial", if_exists="replace", index=False) # MySQL
wr.db.to_sql(df, eng_redshift, schema="public", name="tutorial", if_exists="replace", index=False) # Redshift
# ## Unloading data from Database
wr.db.read_sql_query("SELECT * FROM public.tutorial", con=eng_postgresql) # PostgreSQL
wr.db.read_sql_query("SELECT * FROM test.tutorial", con=eng_mysql) # MySQL
wr.db.read_sql_query("SELECT * FROM public.tutorial", con=eng_redshift) # Redshift
|
tutorials/007 - Redshift, MySQL, PostgreSQL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.0 64-bit (''devlop'': venv)'
# name: python370jvsc74a57bd0ad645be9652fb026c3dd671def0ef771d9b9fe02c8c0a7fb4ce068b245535155
# ---
# +
import os
import yfinance as yf
import pandas as pd
import plotly.graph_objects as go
import plotly.express as px
from datetime import datetime
from pyquery import PyQuery
from dateutil.relativedelta import relativedelta
from FFI import rust_lib
# -
symbols = [
# {"name": "^TWII", "remark": "台灣加權指數"},
# {
# "name": "^TAIEX",
# "remark": "台灣加權報酬指數",
# "fromPath": os.path.join("./extraData", "臺灣加權股價指數"),
# },
# {"name": "^0050", "remark": "0050報酬指數", "fromPath": os.path.join("./extraData", "臺灣50指數")},
# {"name": "0050.TW", "remark": "元大台灣50", "replaceDiv": True},
# {"name": "006208.TW", "remark": "富邦台50", "replaceDiv": True},
{"name": "0051.TW", "remark": "元大中型100", "replaceDiv": True},
# {"name": "0056.TW", "remark": "元大高股息", "replaceDiv": True},
# {"name": "2412.TW", "remark": "中華電信", "replaceDiv": True},
# {"name": "2002.TW", "remark": "中鋼", "replaceDiv": True},
# {"name": "2330.TW", "remark": "台積電", "replaceDiv": True},
# {"name": "2317.TW", "remark": "鴻海", "replaceDiv": True},
# {"name": "6505.TW", "remark": "台塑石化", "replaceDiv": True},
# {"name": "3481.TW", "remark": "群創", "replaceDiv": True},
# {"name": "2303.TW", "remark": "聯電", "replaceDiv": True},
# {"name": "2308.TW", "remark": "台達電", "replaceDiv": True},
]
from stock import *
start="1911-1-1"
end=datetime.now().strftime("%Y-%m-%d")
prefix="TW"
iYear=5
stocks = []
for symbol in symbols:
stocks.append(
Stock(
symbol["name"],
remark=symbol["remark"],
start=start,
end=end,
extraDiv=symbol.get("extraDiv", {}),
replaceDiv=symbol.get("replaceDiv", False),
fromPath=symbol.get("fromPath", False),
)
)
# # ===============================================================
# +
data = {}
for st in stocks:
df = st.history.set_index("Date")
if "Open" not in df.columns:
df["Open"] = 0
if "High" not in df.columns:
df["High"] = 0
if "Low" not in df.columns:
df["Low"] = 0
if "Volume" not in df.columns:
df["Volume"] = 0
df = df[["Open", "High", "Low", "Close", "Adj Close Cal", "Volume"]]
df = df.rename({"Adj Close Cal": "CloseAdj"}, axis="columns")
data[st.name] = df
df = pd.concat(data, axis=1)
data_stat_year = {}
for st in stocks:
df1 = df[st.name]
df1 = df1.dropna()
df1.loc[:, "Volume"] = df1["Volume"].astype(int)
with rust_lib.Stock(df1) as stock:
activeYear = stock.stat_active_year()
holdYear = stock.stat_hold_year()
data_stat_year[f"{st.symbol:7s} A {st.remark}"] = activeYear * 100
data_stat_year[f"{st.symbol:7s} P {st.remark}"] = holdYear * 100
# -
data_stat_year
df=pd.concat(data_stat_year)
df
dataList = []
for symbol, data in df.groupby(level=0):
data = data.dropna(axis=1)
dd = {
"type": "box",
"name": symbol,
"x": data.columns,
"q1": data.loc[:, "25%", :].values[0],
"median": data.loc[:, "50%", :].values[0],
"q3": data.loc[:, "75%", :].values[0],
"lowerfence": data.loc[:, "min", :].values[0],
"upperfence": data.loc[:, "max", :].values[0],
"mean": data.loc[:, "mean", :].values[0],
"sd": data.loc[:, "std", :].values[0],
}
dataList.append(dd)
dataList
data.dropna(axis=1)
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Limpieza de datos y preparacion df
# Basado en Hair et AI. (2013), capítulo 'Examining your data', vamos a proceder con los siguientes pasos
#
# * **1. Entender el problema:** Mirar cada variable y su relevancia para resolver el problema
# * **2. Análisis univariante:** realizado sobre la variable target (SalesPrice)
# * **3. Análisis multiunivariante:** para obtener variables dependientes e independientes
# * **4. Limpieza de datos:** detectar NAs, outliers y variables categóricas
# * **5. Transformación de datos:** aplicando análisis multivariante, vamos a tener que
# - Normalizar datos: hacer que sigan una distribución normal (dado que luego cuando usemos algunos análisis estadísticos, si no lo siguen nos saldrán mal). Lo haremos solo respecto a la variable target y si tenemos pocos datos, dado que este punto para grandes volumenes de datos no suele ser un problema
# - Heterocedasticidad: para asegurar que un error no es constante para todas las variables independientes
# - Linealidad
# - Asegurar que no hay errores correlacionados
#
# * **6. Conclusiones**
#
# ---
#
# En este apartado, el del limpieza y preparación de los datos, abordaremos los puntos 4 a 6
#
# ---
#
# ## ( Obtención de datos)
#
# Realizamos Importación de librerías y ficheros
# +
# Importación de librerías
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from scipy.stats import norm
from sklearn.preprocessing import StandardScaler
from scipy import stats
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# Importación de ficheros
df_train = pd.read_csv('data/PreciosCasas/train.csv')
df_train.describe();
df_train.columns
# -
df_train.describe()
# Como se puede observar, hemos importado 1460 registros distintos, y disponemos de varias variables para determinar el precio de las casas
# Tenemos de variables numéricas y categóricas, además, para empezar el análisis vamos a:
# * seleccionar aquellas que por lógica, nos parece que guardarán más relación con la variable target (SalesPrice). Por ejemplo, si el tener o no garaje puede encarecer un piso (que parece indicar que si), o
# * y que variables a priori, parecen relacionadas entre sí. Por ejemplo, hasta que punto necesito LandScope, que me indica ya la inclinación de la propiedad, si tengo ya LandContour
#
# A priori, suponemos que las siguientes variables son importantes: OverallQual, YearBuilt, TotalBsmtSF, GrLivArea, Neigborhood
#
# y analicemos entonces la **variable target**. Se ve el los resultados anteriores ( o por ``df_train['SalePrice'].describe()`` ) que sí tenemos datos para las 1460 variables, que el mínimo es mayor que cero (y por lo tanto, sí hay información) y que además se distribuye siguiendo una distribución normal con asimetría positiva (possitive skewness), es decir, son sesgo en valores superiores a la media (tenemos propiedades cuyo precio es muy superior al de la mayoría), y con los valores concentrados en la región central de la distribución (curtosis > D. normal).
# **Podemos por tanto continuar el análisis para predicción de esa variable **
# +
#histograma
sns.distplot(df_train['SalePrice']);
# Valor de la asimetría y curtosis
print("Skewness: %f" % df_train['SalePrice'].skew())
print("Kurtosis: %f" % df_train['SalePrice'].kurt())
# -
# ## 4. Limpieza de datos
#
# - **Missing data**
#
# Este tema es muy relevante, pues ¿hasta que punto los NAs siguen un patrón aleatorio o son constantes en nuestros datos? Puede llevarnos a reducir tanto el tamaño de la muestra que hasta nos impida hacer el análisis. Pasemos pues a analizarlo y ver hasta que punto las variables que hemos visto más significativas, están completas o no
#
#missing data ordenados por %
total = df_train.isnull().sum().sort_values(ascending=False)
percent = (df_train.isnull().sum()/df_train.isnull().count()).sort_values(ascending=False)
missing_data = pd.concat([total, percent], axis=1, keys=['Total', 'Percent'])
missing_data.head(20)
# *Hipótesis: un dato no es correcto si tiene más de un 15 % de NAs*
#
# Según esto
# - 'PoolQC', 'MiscFeature', 'Alley', etc no podremos considerarlas (pero no importa, no parecían relevantes tampoco, e incluso ser outliers)
# - 'Garage..' lo mismo, pero como 'GarageCars' no está en la lista, sí (tendrá como máximo un 5% de los valores NAs, pero no más)
# - 'MasVnrArea' y 'MasVnrType' tampoco parecían relevantes, así que bien (y mirando el mapa de calor, estabas correlacionadas con 'YearBuilt' y 'OverallQual' así que no perdemos nada)
# - 'Electrical' tiene solo una observación, así que la borraremos pero sí mantendremos la variable
#
# +
#Creación de los nuevo data frames
df_train = df_train.drop((missing_data[missing_data['Total'] > 1]).index,1)
df_train = df_train.drop(df_train.loc[df_train['Electrical'].isnull()].index)
# Veamos que está bien (debe dar 0
print ("El valor seberia salir 0 y sale: ");
df_train.isnull().sum().max()
# -
# - **Imputación**
#
# La opción para quitar los NAs cuando no queremos simplemente quitar las columnas que los contengan, es la imputación: rellenar esos "huecos" el valor medio. Una extensión a esto sería ademas indicar en que casos se ha hecho eso (en otra columna) para que le modelo lo tenga en cuenta, pero normalmente no aporta tanto como el tiempo que tarda.
# Vamos a hacer una imputación simple, para los NAs de los valores numéricos (obvio)
#
Col_numeros = ['LotArea', 'YearBuilt', '1stFlrSF', '2ndFlrSF', 'FullBath', 'BedroomAbvGr', 'TotRmsAbvGrd', 'SalePrice']
df_train_numeros = df_train[Col_numeros]
# +
from sklearn.preprocessing import Imputer
# Imputation
my_imputer = Imputer()
df_train_numeros_imp = my_imputer.fit_transform(df_train_numeros)
# -
# Pero **¿Cómo sabemos que esto es mejor?** hagamos una prueba simple, una regresión de los dos df a ver que error se produce
# +
#Cojamos por ejemplo un randomforest (función)
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
def score_dataset(X_train, X_test, y_train, y_test):
model = RandomForestRegressor()
model.fit(X_train, y_train)
preds = model.predict(X_test)
return mean_absolute_error(y_test, preds)
# +
#Hagamos el grupo train y test
from sklearn.model_selection import train_test_split
X_train_N, X_test_N, y_train_N, y_test_N = train_test_split(df_train_numeros,
df_train_numeros['SalePrice'],
train_size=0.7,
test_size=0.3,
random_state=0)
# +
df_train_numeros
##1) Quitando todos los nulos
cols_with_missing = [col for col in X_train_N.columns
if X_train_N[col].isnull().any()]
reduced_X_train = X_train_N.drop(cols_with_missing, axis=1)
reduced_X_test = X_test_N.drop(cols_with_missing, axis=1)
print("Mean Absolute Error para los datos numericos:")
print(score_dataset(reduced_X_train, reduced_X_test, y_train_N, y_test_N));
##2) imputacióon
my_imputer = Imputer()
imputed_X_train = my_imputer.fit_transform(X_train_N)
imputed_X_test = my_imputer.transform(X_test_N)
print("Mean Absolute Error para los datos con imputación:")
print(score_dataset(imputed_X_train, imputed_X_test, y_train_N, y_test_N))
# -
# # No deberia salir esto
imputed_X_train.shape
# - **Outliars**
#
# Hay que definir un umbral a partir del cual definimos un dato como outliar, para lo cual necesitaremos estandarizar los datos, y analizar cuanto se desvian de 0 por encima y por debajo
#
#Estandarizar datos del target
saleprice_scaled = StandardScaler().fit_transform(df_train['SalePrice'][:,np.newaxis]);
low_range = saleprice_scaled[saleprice_scaled[:,0].argsort()][:10]
high_range= saleprice_scaled[saleprice_scaled[:,0].argsort()][-10:]
print('Rango bajo (low) de la distribución:')
print(low_range)
print('\nRango alto (high) de la distribución')
print(high_range)
# los valores por debajo no preocupan (en torno a -1), pero los por encima si, especiamente esos de más de 7 (seguramente son outliars). Veamos estos datos en perspectiva con las dos variables más significativas o relacionadas con la target
# saleprice/grlivarea
var = 'GrLivArea'
data = pd.concat([df_train['SalePrice'], df_train[var]], axis=1)
data.plot.scatter(x=var, y='SalePrice', ylim=(0,800000));
# en efecto, tenemos dos valores que para ese tamaño de propiedad, no pega que sean tan bajos y se desvian de la tendencia, así que los quitaremos. Mantendremos sin embargo los dos valores de precio más alto, dado que SI parecen mantener esa tendencia
#Eliminar outliers
df_train.sort_values(by = 'GrLivArea', ascending = False)[:2]
df_train = df_train.drop(df_train[df_train['Id'] == 1299].index)
df_train = df_train.drop(df_train[df_train['Id'] == 524].index)
# saleprice/TotalBsmtSF
var = 'TotalBsmtSF'
data = pd.concat([df_train['SalePrice'], df_train[var]], axis=1)
data.plot.scatter(x=var, y='SalePrice', ylim=(0,800000));
# Para este caso, no merece la pena quitar nada
# saleprice/OverallQual
var = 'OverallQual'
data = pd.concat([df_train['SalePrice'], df_train[var]], axis=1)
data.plot.scatter(x=var, y='SalePrice', ylim=(0,800000));
# ## 5. Transformación de datos
#
# Veamos como han quedado los datos y que distribución siguen para saber si debemos ajustarlos de alguna manera
#
# ### Normalización
#histograma y normal probability plot
sns.distplot(df_train['SalePrice'], fit=norm);
fig = plt.figure()
res = stats.probplot(df_train['SalePrice'], plot=plt)
# Para casos de asimetría positiva, se puede lograr que los datos sigan una distribución normal mediante una transformación logartimica, esto es:
# +
# transformación logarítmica
df_train_log = pd.DataFrame(df_train)
df_train_log ['SalePrice'] = np.log(df_train['SalePrice'])
# nuevo histograma y normal probability plot
sns.distplot(df_train_log['SalePrice'], fit=norm);
fig = plt.figure()
res = stats.probplot(df_train_log['SalePrice'], plot=plt)
# -
# Y claro, pasará lo mismo con
#
# 1) 'GrLivArea'
# 2) 'TotalBsmtSF'
# 1) GrLivArea
#histograma y normal probability plot
sns.distplot(df_train_log['GrLivArea'], fit=norm);
fig = plt.figure()
res = stats.probplot(df_train_log['GrLivArea'], plot=plt)
# +
# transformación logaritmica
df_train_log ['GrLivArea'] = np.log(df_train_log['GrLivArea'])
#transformed histogram and normal probability plot
sns.distplot(df_train_log['GrLivArea'], fit=norm);
fig = plt.figure()
res = stats.probplot(df_train_log['GrLivArea'], plot=plt)
# -
# 2) TotalBsmtSF
#histogram and normal probability plot
sns.distplot(df_train_log['TotalBsmtSF'], fit=norm);
fig = plt.figure()
res = stats.probplot(df_train_log['TotalBsmtSF'], plot=plt)
# En este caso hay valores = 0, por lo tanto NO se puede aplicar logaritmos. De nuevo, hay que hacer una suposición, cuando esos ceros seguramente se refieren a que no hay sotano, así que en este caso, la teoría dice que hay que hacer una nueva variable binaria (siene sotano si/no) y luego a los que sí, aplicar ya la transformación de los datos
#
# +
df_train_log['ConBasement'] = 1
df_train_log['ConBasement'][df_train_log['TotalBsmtSF'] ==0] = int (0)
# Y ahora quitamos los 0 de TotalBsmtSF y hacemos el logaritmo
df_train_log2 = pd.DataFrame(df_train_log)
# transformación logaritmica
df_train_log2 ['TotalBsmtSF'][df_train_log2 ['TotalBsmtSF']!= 0] = np.log1p(df_train_log2['TotalBsmtSF'])
#transformed histogram and normal probability plot
sns.distplot(df_train_log2['TotalBsmtSF'], fit=norm);
fig = plt.figure()
res = stats.probplot(df_train_log2['TotalBsmtSF'], plot=plt)
# -
#
# ### Heterocedasticidad
# Lo haremos de forma gráfica: si tiene forma cónica o de diamante
#scatter plot
plt.scatter(df_train['GrLivArea'], df_train['SalePrice']);
plt.scatter(df_train_log['GrLivArea'], df_train_log['SalePrice']);
# Antes los datos tenian forma de diamante, pero tras la normalización ya no, por lo que no tenemos problemas en este punto para 'GrLivArea'
#scatter plot
plt.scatter(df_train[df_train['TotalBsmtSF']>0]['TotalBsmtSF'], df_train[df_train['TotalBsmtSF']>0]['SalePrice']);
#scatter plot
plt.scatter(df_train_log[df_train_log['TotalBsmtSF']>0]['TotalBsmtSF'], df_train_log[df_train_log['TotalBsmtSF']>0]['SalePrice']);
# en este caso igual, parece que la varianza del target respecto a TotalBsmtSF es similar en todo el rango
# ### Variables Dummy
# Hay una función que lo hace
#convert categorical variable into dummy
df_train_log = pd.get_dummies(df_train_log)
# ## Conclusiones
# Disponemos de un data frame limpio y listo para ser usado en los siguientes apartados de modelización. Subámoslo pues a nuestro repositorio para poder usarlo
#
df_train_log.to_csv('data/PreciosCasas/train_final.csv', sep='\t', encoding='utf-8')
df_train_log.describe()
|
0B. Limpieza de datos y preparacion df.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="suspected-lambda"
# # Auxiliary Lines in Planar Geometry
#
# ## Preface
#
# Proving a proposition in planar geometry is like an outdoor exploration -- to find a path from the starting point (the problem) to the destination (the conclusion). Yet the path can be a broad highway, or a meandering trail, or -- you may even find yourself in front of a river.
#
# The auxiliary lines are the bridges to get you across. Such lines are indispensible in many problems, or can drastically simplify the proof in others. Just like there is no universal rule on where or how to build the bridge for all kind of terrains, the auxiliary lines have to be designed for each individual problem. Difficult as it can be for beginners, the process of analyzing the problem and finding the solution is rigorous, creative, fascinating and extremely rewarding. This booklet is intended to give you a helping hand.
# + [markdown] id="massive-introduction"
# ## Basic problems
#
# Let's look at a simple example. In $\triangle ABC$, $AD$ is a median. $DE$ extends $AD$ and $DE = AD$ (Figure 1). Show that $BE \| AC$ and $BE = AC$.
#
# 
#
# **Figure 1**
#
# It's obvious that to prove $BE \| AC$, we can start from proving $\angle EBC = \angle ACB$, or $\angle BEA = \angle CAE$. To prove $BE = AC$, we can try to prove $\triangle BED \cong \triangle CAD$. Note that as the corresponding angles are equal in congruent triangles, $\angle EBC = \angle ACB$ or $\angle BEA = \angle CAE$ will already be implied when the congruency is established, which is all we need. The proof can be written as follows:
#
# $\because$ $AD$ is a median of $\triangle ABC$
#
# $\therefore BD = DC$
#
# Also $\because$ $\angle ADC = \angle BDE$ and $DE = AD$
#
# $\therefore \triangle BED \cong \triangle CAD$ (SAS)
#
# $\therefore BE = AC, \angle EBD = \angle ACD$
#
# $\therefore BE \| AC$.
#
# Given the drawing, we came to the solution fairly smoothly in this example. But in many cases, the drawing itself may not be enough, and auxiliary lines are required. Even in this example, if we connect $CE$, we get $\text{▱}ABEC$, from which we can easily prove $BE = AC$ and $BE \| AC$.
#
# Let's see some more examples.
# + [markdown] id="introductory-robin"
# ### Medians
#
# 
#
# **Figure 2**
#
# 
#
# **Figure 3**
#
# 
#
# **Figure 4**
#
# 
#
# **Figure 5**
#
# 
#
# **Figure 6**
# + [markdown] id="related-republic"
# ### Midpoints
# + [markdown] id="sustained-crawford"
# ### Angle bisects
# + [markdown] id="spare-thing"
# ### Trapezoids, squares and triangles
#
# 
#
# **Figure 20**
#
# 
#
# **Figure 21**
#
# 
#
# **Figure 22**
#
# 
#
# **Figure 23**
#
# 
#
# **Figure 24**
#
# 
#
# **Figure 25**
#
# 
#
# **Figure 26**
#
# 
#
# **Figure 27**
#
# 
#
# **Figure 28**
#
# 
#
# **Figure 29**
#
# + [markdown] id="packed-product"
# ### Special points in triangles
#
# + [markdown] id="injured-concern"
# ### Double angles
# + [markdown] id="single-hammer"
# ### Right triangles
# + [markdown] id="representative-transcription"
# ### Proportional segments
#
# 
#
# **Figure 47**
#
# 
#
# **Figure 48**
#
# 
#
# **Figure 49**
#
# 
#
# **Figure 50**
#
# 
#
# **Figure 51**
#
# 
#
# **Figure 52**
#
# 
#
# **Figure 53**
#
# 
#
# **Figure 54**
#
# 
#
# **Figure 55**
#
# 
#
# **Figure 56**
#
# 
#
# **Figure 57**
#
# + [markdown] id="zeFAuOr8l54Z"
# Problem 24
#
# As show in Figure 58, A line segment $XY$ intersects $\triangle$$ABC$ at $X$,$Y$,$Z$. Prove $\frac {AX}{XB} \frac{BY}{YC} \frac{CZ}{ZA}$=1.
#
# This claim is called Monelaus Theorem
#
#
# 
#
# **Figure 58**
#
# Proof 1
#
#
# Analysis To prove that $\frac {AX}{XB} \frac{BY}{YC} \frac{CZ}{ZA}$=1, first change it to $ \frac{BY}{YC} \frac{CZ}{ZA} = \frac {XB}{AX}$. To prove $ \frac{BY}{YC} \frac{CZ}{ZA} = \frac {XB}{AX}$, need to find a line segment a, such that $ \frac{BY}{YC} \frac{CZ}{ZA} = \frac {XB}{a} \frac{a}{AX}$. This equation is a multiplication of the proportions $ \frac{BY}{YC}= \frac {XB}{a}$, and $ \frac{CZ}{ZA} = \frac {a}{AX}$. Considering $ \frac{BY}{YC} = \frac {XB}{a}$, it is possible to create a line $CD$ passing C with $CD\||XY$, therefore, $ \frac{BY}{YC} = \frac {BX}{XD}$ (Refer to Figure 59). At $\triangle$ACD, we get $\frac{CZ}{ZA} = \frac {DXB}{AX}$.(Here DX is the desired auxiliary line segment a).With this analysis, it is not difficult to prove the claim.
#
#
# 
#
# **Figure 59**
#
# Proof Omitted.
#
# Proof 2
#
# Analysis Similar to the analysis of Proof 1, to prove $ \frac{BY}{YC}=\frac {XB}{a}$ and $\frac{CZ}{ZA} = \frac {a}{AX}$, an alternative method is to create a line $CD$ passing C with $CD\||AB$ (refer to Figure 60). The proof can be made.
#
#
# 
#
# **Figure 60**
#
# Proof Reader please prove it yourself.
#
# Proof 3
#
# Analysis If change $\frac {AX}{XB} \frac{BY}{YC} \frac{CZ}{ZA}$=1, to $ \frac{AX}{XB}\frac{CZ}{ZA} = \frac {YC}{BY}$, the next step is to find line segment b, such that $ \frac{AX}{XB} \frac{CZ}{ZA} = \frac {b}{BY} \frac{YC}{b}$. This equation is a multiplication of two proportionals $ \frac{AX}{XB} = \frac{b}{BY}$ and $\frac{CZ}{ZA} = \frac {YC}{b}$. Refer to Figure 61, it is possible to create a line segment AD passing A to get $AD\||XY$, which intersects the extension of $BC$ at D. At $\triangle$ABD, $\frac {AX}{XB} =\frac{YD}{BY}$, at $\triangle$ACD, $\frac {CZ}{ZA} =\frac{YC}{YD}$. (Here $YD$ is the desired auxiliary line segment b). It is not difficult to prove the claim.
#
#
# 
#
# **Figure 61**
#
# Proof oimitted.
#
# Passing B to create parallel line to $XY$ or to $AC$ can do the proof the same way. We do not go details for those approaches.
#
# Proof 4
#
# Analysis To prove $\frac {AX}{XB} \frac{BY}{YC} \frac{CZ}{ZA}$=1, if change the three ratios $\frac {AX}{XB}$, $\frac{BY}{YC}$, $\frac{CZ}{ZA}$ to $\frac{a}{b}$,$\frac{b}{c}$, $\frac{c}{a}$ respectively, then the product of the three must equal 1. To get $\frac{a}{b}$,$\frac{b}{c}$, $\frac{c}{a}$, need to find a line segment that is relavent to $\frac {AX}{XB}$, $\frac{BY}{YC}$, $\frac{CZ}{ZA}$. As shown in Figure 62, if connect $BZ$, create parallel lines of $BZ$ passing A and passing C, intersecting $XY$ and its extension line at $M,N$, then $MY$ is the desired line segment. Therefore, we have $\frac {AX}{XB} = \frac{AM}{BZ}$, $\frac{BY}{YC} = \frac{BZ}{CN}$, $\frac{CZ}{ZA} = \frac{CN}{AM}$, a multiplication of the three equations get $\frac {AX}{XB} \frac{BY}{YC} \frac{CZ}{ZA}$ = $\frac {AM}{BZ} \frac{BZ}{CN} \frac{CN}{AM}$ = 1. Similarly, connect $CX$(or $AY$), create $AM$,$BN$ parallel to $CX$, intersecting $XY$ and its extension at $M$,$N$ could get the same result.
#
#
# 
#
# **Figure 62**
#
#
# Proof Omitted
#
# + [markdown] id="jNFlUmGB6-P6"
# Problem 25
#
# As shown in Figure 63, inside $\odot o$ from the end of the diameter AB create perpendicular line $AE$, $BF$ ($E,F$ are the feet of perpendicular). Prove $OE=OF$.
#
# 
#
# **Figure 63**
#
# Analysis To prove $OE=OF$, the most straightforward approach is to prove $\triangle OEF$ is isoceles triangle, i.e., start with proving $\angle OEF$=$\angle OFE$. But the vortexes of the two angles are not on the cicle, need more steps to prove. As illustrated in the Firgure, create $OM \bot CD$, if we could prove $OM$ is the perpendicular bisector of $EF$, then we get $OE=OF$. So we need to prove $EM=MF$. Since $AE \bot CD$, $BF \bot CD$, therefore $AE \|| OM \|| BF$. Apply parallel intercept theorem, we get $EM=MF$.
#
# Proof Omitted
#
# Let's lok at the application of the intercept theorem. If we change the proof to a difference approach as shown in Figure 64, using intercept theorem to create the auxilliary line $ OM \bot CD$, it is easier for beginners. It is more difficult to figure out the relationship using Figure 63. There are two cases of the intercept line in the theorem, one is that the two intercept lines intersect outside of the parallel lines, the other is that the two intercept lines intersect inside of the parallel lines. The latter needs particular attention.
#
#
# 
#
# **Figure 64**
#
# + [markdown] id="GRj1c6GqPFD8"
# Problem 26
#
# As shown in Figure 65, $AD$ is the altitude of $\triangle ABC$, extend $AD$ and intercept circumcircle at $H$, then use $AD$ as diameter to create a circle, intersecting $AB$,$AC$ at $E,F$. $EF$ and $AD$ intersect at $G$. Prove $AD^2 = AD^.AH$
#
#
# 
#
# **Figure 65**
#
# Proof 1
#
# Analysis To prove $AD^2 = AD^.AH$ is equivalent to prove $\frac{AD}{AG}=\frac{AH}{AD}$. Since $AG, AH, AD$ are three coline segments, no similar triangle is composed of the three line segments. Therefore, we must find a transitional proportion to connect the original proportion and the conclusion. Since $AD$ is the diameter of $\odot$$AEF$, connect $DE$, $\triangle$DAB is right triangle.Therefore, $AD^2 = AE^.AB$. To prove $AD^2 = AD^.AH$, we only need to get $AD^. AB=AG^.AH$, i.e. to get $\frac{AE}{AG}=\frac{AH}{AB}$, which is straightforward if we could prove $\triangle AEG \sim \triangle AHB$. Since both triangles shared an angle, only need to prove $\angle AEG = \angle H$. Since $\angle H=\angle C$, only need to prove $\angle AEG=\angle C$. And since $\angle C$ is supplementary of $\angle CAD$, $\angle AEG$ is supplementary of $\angle DEF$, and $\angle CAD = \angle DEF$, therefore, the proof is done.
#
# Proof omitted.
#
# Proof 2
#
# Analysis To prove $AE^.AB=AG^.AH$, only need to prove $E,G,H,B$ are concyclic. To prove $E,G,H,B$ are concyclic, we may start with proving $\angle ABH = \angle AGE$. Since $\angle ABH = \angle ABD + \angle DBH$, $\angle AGE = \angle ADE + \angle DEG$, thereforre, we only need to prove $\angle ABD = \angle ADE$, and $\angle DBH= \angle DEG$", which are straightforward.
#
# Proof omitted.
# + [markdown] id="smgM0C58zXLA"
# The claims from problem 22 to problem 26 are all related to proportional line segments. To prove the problem related to proportional line segments, first need to convert the conclusion into proportions. For example, to prove $ab=cd$, first convert $ab=cd$ to $\frac{a}{c}=\frac{d}{b}$ or$\frac{a}{d}=\frac{c}{b}$; to prove $a^2=bc$, first convert to $\frac{a}{b}=\frac{c}{a}$ or $\frac{a}{c}=\frac{b}{a}$. In euclidian geometry, to prove a theorem related to proportional line segments, we may start from a theorem related to similar triangles, to find whether the link segments in the proportional are mapped to the corresponding pair of sides in the similar triangles. If such triangles exist in the figure, we may prove they are smiliar triangles If such triangles does not exist, we need to add auxiliary line to connect some end points of the line segments to construct triangles (refer to problem 22), then prove they are similar. If the line segments in the proportional cannot construct corresponding sides of similar triangles, usually we need to select intersect pints to construct auxiliary lines parallel to other line segments, then apply parallem theory to get proportional relationship. (refer to problem 23 and problem 24 proof 1,2,3).
#
# If the line segment in the proportinals are colinear, they cannot construct two similar triangles, and cannot apply parallel theorem, then we need to find two transitional line segment based on the characteristics of the figure then approve. (refer to problem 26 proof 1 and 2).
#
# In addition, the right triangle altitude theorem,Thales' theorem and angle bisector theorem may be used to prove the problems related to proportional line segments.
# + [markdown] id="Ck876R0dSuAF"
# Practice Questions
#
# 1. At $\triangle ABC$, a line intersects $AB$, $AC$ and the extension of $BC$ at $D,E,F$, and $AD=AE$. Prove $\frac{CF}{BF}=\frac{CE}{BD}$.
#
# 2. At $\triangle ABC$, a line intersects $AB$, $AC$ and the extension of $BC$ at $D,E,F$, and $\frac{AE}{CE}=\frac{BF}{CF}$.
#
# 3. At $\triangle ABC$, $AT$ is the bisector of exterior angle of $\angle A$. Prove $\frac{BT}{CT} = \frac{AB}{AC}$.
#
# 4. $AB$ is the diameter of a semicircle. $PQ\bot AB$ at $P$, and intersects the semicircle at Q. R is a random point at the semicircle. $AR, BR$ intersect the line $PQ$ at $X,Y$ respectively. Prove $PQ^2=PX^.PY$.
# + [markdown] id="powered-progressive"
# ### Concyclic points
#
# 
#
# **Figure 66**
#
# 
#
# **Figure 66**
#
# 
#
# **Figure 66**
#
#    
#
# **Figure 69**
# + [markdown] id="major-impossible"
# ### Tangent and intersecting circles
#
# #### Exercises
#
# 1. Two circles intersect at $A$ and $B$. $AC$ and $AD$ are the diameters of two circles respectively. Show that $C, B$, and $D$ are collinear.
#
# 2. Two circles intersect at $A$ and $B$. $AD$ and $BF$ are the chords of the two circles, and intersect with the other circle at $C$ and $E$ respectively. Show that $CF\|DE$.
#
# 3. Two circles are tangent at $P$. Chord $AB$ of the first circle is tangent to the second circle at $C$. The extension of $AP$ intersects the second circle at $D$. Show that $\angle BPC=\angle CPD$.
#
# 4. Given semicircle $O$ with $AB$ as a diameter, $C$ is a point on the semicircle, and $CD\bot AB$ at $D$. $\odot P$ is tangent to $\odot O$ externally at $E$, and to line $CD$ at $F$. $A$ and $E$ are on the same side of $CD$. Show that $A, E, F$ are collinear.
# + [markdown] id="infrared-section"
# ## Problem Set 1
#
# 1. $AD$ is a median of $\triangle ABC$, and $AE$ is a median of $\triangle ABD$. $BA=BD$. Show that $AC=2AE$.
#
# 
#
# **Problem 1**
#
# 2. Prove the perimeter of a triangle is greater than the sum of the three medians.
#
# 3. Given triangle $ABC$, $P$ is a point on the exterior bisector of angle $A$. Show that $PB+PC > AB+AC$.
#
# 
#
# **Problem 3**
#
# 4. For right triangle $ABC$ with $AB$ as the hypotenuse, the perpendicular bisector $ME$ of $AB$ intersects the angle bisector of $C$ at $E$. Show that $MC=ME$.
#
# 
#
# **Problem 4**
#
# 5. For isosceles triangle $ABC$ with $AB=AC$, $CX$ is the altitude on $AB$. $XP\bot BC$ at $P$. Show that $AB^2=PA^2+PX^2$.
#
# 
#
# **Problem 5**
#
# 6. Show that the diagonal of a rectangle is longer than any line segment between opposite sides.
#
# 7. For square $ABCD$, $E$ is the midpoint of $CD$. $BF\bot AE$ at $F$. Show that $CF=CB$.
#
# 
#
# **Problem 7**
#
# 8. In isosceles triangle $ABC$ with $AB=AC$, the circle with $AB$ as a diameter intersects $AC$ and $BC$ at $E$ and $D$ respectively. Make $DF\bot AC$ at $F$. Show that $DF^2=EF\cdot FA$.
#
# 
#
# **Problem 8**
#
# 9. Show that for a triangle, the reflection points of the orthocenter along three sides are on the circumcircle.
#
# 10. As shown in the figure, $AB$ is a diameter of $\odot O$, and $AT$ is a tangent line of $\odot O$. $P$ is on the extension of $BM$ such that $PT\bot AT$ and $PT=PM$. Show that $PB=AB$.
#
# 
#
# **Problem 10**
#
# 11. As shown in the figure, $AB$ is a diameter of $\odot O$, and $P$ is a point on the circle. $Q$ is the midpoint of arc $\widearc{BP}$ and tangent $QH$ intersects $AP$ at $H$. Show that $QH\bot AP$.
#
# 
#
# **Problem 11**
#
# 12. As shown in the figure, two circles are tangent internally at $P$. A secant intersects the two circles at $A, B, C, D$. Show that $\angle APB=\angle CPD$.
#
# 
#
# **Problem 12**
#
# 13. As shown in the figure, two circles are tangent externally at $P$. A secant intersects the two circles at $A, B, C, D$. Show that $\angle APD+\angle BPC=180\degree$.
#
# 
#
# **Problem 13**
#
# 14. Two circles intersect at $A,B$. A line through $A$ intersects the two circles at $C$ and $D$. The tangent lines at $C$ and $D$ intersect at $P$. Show that $B, C, P, D$ are concyclic.
#
# 
#
# **Problem 14**
#
# 15. In $\triangle ABC$, $\angle C=90\degree$, and $CD$ is an altitude. The circle with $CD$ as a diameter intersects $AC$ and $BC$ at $E$ and $F$ respectively. Show that $\frac{BF}{AE}=\frac{BC^3}{AC^3}$.
#
# 
#
# **Problem 15**
#
# 16. In $\triangle ABC$, $\angle B=3\angle C$. $AD$ is the angle bisector of $\angle A$. $BD\bot AD$. Show that $BD=\frac{1}{2}(AC-AB)$.
#
# 
#
# **Problem 16**
#
# 17. In right triangle $ABC$, $\angle A=90\degree$, and $AD$ is the altitude on $BC$. $BF$ is the angle bisector of $\angle B$, and $AD$ and $BF$ intersect at $E$. $EG\|BC$. Show that $CG=AF$.
#
# 
#
# **Problem 17**
#
# 18. As shown in the figure, $D$ and $E$ are the midpoints of $AB$ and $AC$ respectively. $AB>AC$. $F$ is a point between $B$ and $D$ such that $DF=AE$. $AH$ is the angle bisector of $\angle BAC$. $FH\bot AH$, and $FH$ intersects $BC$ at $M$. Show that $BM=MC$.
#
# 
#
# **Problem 18**
#
# 19. In trapezoid $ABCD$, $AD\|BC$ and $AD+BC=AB$. $F$ is the midpoint of $CD$. Show that the angle bisectors of $\angle A$ and $\angle B$ intersect at $F$.
#
# 
#
# **Problem 19**
#
# 20. In $\triangle ABC$, $AC=BC$, and $\angle B=2\angle C$. Show that $AC^2=AB^2+AC\cdot AB$.
#
# 21.
#
# 
#
# **Problem 21**
#
# 22.
#
# 
#
# **Problem 22**
#
# 23.
#
# 
#
# **Problem 23**
#
# 24.
#
# 
#
# **Problem 24**
#
# 25.
#
# 
#
# **Problem 25**
#
# 26.
#
# 
#
# **Problem 26**
#
# 27.
#
#
# 28.
#
# 
#
# **Problem 28**
#
# 29.
#
# 
#
# **Problem 29**
#
# 30.
#
#
# 31.
#
# 
#
# **Problem 31**
#
# 32.
#
# 
#
# **Problem 32**
#
# 33.
#
# 
#
# **Problem 33**
#
# 34.
#
# 
#
# **Problem 34**
#
# 35.
#
# 
#
# **Problem 35**
#
# 36.
#
# 
#
# **Problem 36**
#
# 37.
#
# 
#
# **Problem 37**
#
# 38.
#
# 
#
# **Problem 38**
#
# 39.
#
# 
#
# **Problem 39**
# + [markdown] id="appropriate-seven"
# ## Advanced problems
# + [markdown] id="worthy-official"
# ### Problem Set 2
#
# 
#
# **Problem 2**
#
# 
#
# **Problem 3**
#
# 
#
# **Problem 4**
#
# 
#
# **Problem 5**
#
# 
#
# **Problem 6**
#
# 
#
# **Problem 7**
#
# 
#
# **Problem 8**
#
# 
#
# **Problem 9**
#
# 
#
# **Problem 10**
#
# 
#
# **Problem 11**
#
# 
#
# **Problem 12**
#
# 
#
# **Problem 13**
#
# 
#
# **Problem 14**
#
# 
#
# **Problem 15**
#
# 
#
# **Problem 19**
#
# 
#
# **Problem 20**
#
# + [markdown] id="medium-basement"
# ## Hints and answer keys
|
Auxiliary Lines.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (enm)
# language: python
# name: enm
# ---
import pandas as pd
import numpy as np
from enm.utils import *
# +
gaf = snakemake.input['gaf']
obo = snakemake.input['obo']
background_file = snakemake.input['background_file']
sgd_info = snakemake.input['sgd_info']
goea , geneid2name , obo = create_goea(gaf = gaf, obo_fname=obo,
background=background_file, sgd_info_tab = sgd_info,
goset=['BP'], methods=['fdr','fdr_bh','bonferroni'])
# -
signaling_related_gos = ['GO:0007165', 'GO:0023052','GO:0007154' ]
sensors_df = pd.read_csv(snakemake.input.sensors_pcc)
# + tags=[]
# %%capture
query_gene_ids = [key for key,value in geneid2name.items() if value in sensors_df.loc[:,'Systematic gene name'].unique()]
goea_res_all = goea.run_study(query_gene_ids)
goea_res_sig = [r for r in goea_res_all if r.p_fdr<0.1]
go_df_sensor = goea_to_pandas(goea_res_sig, geneid2name)
# -
go_df_sensor
sensor_signaling_go = [i for i in goea_res_all if i.GO in signaling_related_gos]
sgd_desc = pd.read_csv(snakemake.input.sgd_info,'\t',header=None)
(sensors_df
.loc[sensors_df['Systematic gene name'].isin([*[geneid2name[i] for i in sensor_signaling_go[0].study_items],*['YCL025C','YDR463W','YHR006W','YBR068C']])]
.loc[:,['orf_name','Systematic gene name','go_group','label']].reset_index(drop=True)
.merge(sgd_desc.loc[:,[3,15]],left_on='Systematic gene name',right_on = 3).rename(columns={15:'Description'})
.drop(labels=3,axis=1)
.to_csv(snakemake.output.sensors_signaling_df))
effectors_df =pd.read_csv(snakemake.input.effector_pcc)
# + tags=[]
# %%capture
query_gene_ids = [key for key,value in geneid2name.items() if value in effectors_df.loc[:,'Systematic gene name'].unique()]
goea_res_all_eff = goea.run_study(query_gene_ids)
goea_res_sig_eff = [r for r in goea_res_all_eff if r.p_fdr<0.1]
go_df_effector = goea_to_pandas(goea_res_sig_eff, geneid2name)
# -
eff_go_sig = [i for i in goea_res_all_eff if i.GO in signaling_related_gos]#[0]
(effectors_df
.loc[effectors_df['Systematic gene name'].isin([geneid2name[i] for i in eff_go_sig[0].study_items])]
.loc[:,['orf_name','Systematic gene name','go_group','effector_cluster']].reset_index(drop=True)
.merge(sgd_desc.loc[:,[3,15]],left_on='Systematic gene name',right_on = 3).rename(columns={15:'Description'})
.drop(labels=3,axis=1))
|
notebooks/04-Signaling-related-effector-sensors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seaborn as sns;
import pandas as pd;
sales = pd.read_excel('Datasets/sales.xls',sheet_name=1)
sales.columns
sns.relplot('Sales','Profit',data=sales, col='Customer Segment', col_wrap=2)
sns.relplot('Sales','Profit',data=sales, hue="Ship Mode", col="Customer Segment", row='Ship Mode')
sns.relplot('Sales','Profit',data=sales,col='Customer Segment', row='Product Category', hue='Ship Mode')
|
Seaborn - Crash Course/RowColumn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Tutorial: Compressing Natural Language With an Autoregressive Machine Learning Model and `constriction`
#
# - **Author:** <NAME>, University of Tuebingen
# - **Initial Publication Date:** Jan 7, 2022
#
# This is an interactive jupyter notebook.
# You can read this notebook [online](https://github.com/bamler-lab/constriction/blob/main/examples/python/03-tuorial-autoregressive-nlp-compression.ipynb) but if you want to execute any code, we recommend to [download](https://raw.githubusercontent.com/bamler-lab/constriction/main/examples/python/03-tuorial-autoregressive-nlp-compression.ipynb) it.
#
# More examples, tutorials, and reference materials are available at <https://bamler-lab.github.io/constriction/>.
#
# ## Introduction
#
# This notebook teaches you how to losslessly compress and decompress data with `constriction` using an autoregressive machine learning model.
# We will use the simple character based recurrent neural network toy-model for natural language from the [Practical PyTorch Series](https://github.com/spro/practical-pytorch/blob/master/char-rnn-generation/char-rnn-generation.ipynb).
# You won't need any fancy hardware (like a GPU) to go along with this tutorial since the model is so simplistic that it can be trained on a normal consumer PC in about 30 minutes.
#
# The compression method we develop in this tutorial is for demonstration purpose only and not meant as a proposal for practical text compression.
# While you will find that our method can compress text that is similar to the training data very well (with a 40% reduction in bitrate compared to `bzip2`), you'll also find that it will generalize poorly to other text forms and that compression and decompression are excruciatingly slow.
# The poor generalization performance is not a fundamental issue of the presented approach; it is just a result of using a very simplistic model combined with a very small training set.
# The runtime performance, too, would improve to *some* degree if we used a better suited model and if we ported the implementation to a compiled language and avoid permanent data-copying between Python, PyTorch, and `constriction`.
# However, even so, autoregressive models tend to be quite runtime-inefficient in general due to poor parallelizability.
# An alternative to autoregressive models for exploiting correlations in data compression is the so-called bits-back technique.
# An example of bits-back coding with `constriction` is provided in [this problem set](https://robamler.github.io/teaching/compress21/problem-set-05.zip) (with [solutions](https://robamler.github.io/teaching/compress21/problem-set-05-solutions.zip)).
# ## Step 1: Prepare Your Environment
#
# We'll use the PyTorch deep learning framework to train and evaluate our entropy model.
# Follow the [installation instructions](https://pytorch.org/get-started/locally/) (you'll save a *lot* of download time and disk space if you install the CPU-only version).
# Then restart your jupyter kernel and test if you can import PyTorch:
import torch
# Next, make sure you have a recent enough version of `constriction` and a few other small Python packages:
# !python -m pip install constriction~=0.2.2 tqdm~=4.62.3 unidecode~=1.3.2
# **Restart your jupyter kernel again.**
# Then let's get started.
# ## Step 2: Get Some Training Data
#
# We'll train our text model on the same [100,000 character Shakespeare sample](https://raw.githubusercontent.com/karpathy/char-rnn/master/data/tinyshakespeare/input.txt) that was used in the [Practical PyTorch Series](https://github.com/spro/practical-pytorch/blob/master/char-rnn-generation/char-rnn-generation.ipynb).
# Different from the Practical PyTorch Series, however, we'll split the data set into a training set and a test set so that we can test our compression method on text on which the model wasn't explicitly trained.
#
# Start with downloading the full data set:
# !wget -O shakespeare_all.txt https://raw.githubusercontent.com/karpathy/char-rnn/master/data/tinyshakespeare/input.txt
# Now, let's split the data set into, say, 90% training data and 10% test data by randomly assigning each line of the full data set to either one of those two subsets.
# In a real scientific project, you should usually split into more than just two sets (e.g., add an additional "validation" set that you then use to tune model hyperparameters).
# But we'll keep it simple for this tutorial.
# +
import numpy as np
target_test_set_ratio = 0.1 # means that about 10% of the lines will end up in the test set.
train_count, test_count = 0, 0
rng = np.random.RandomState(830472) # (always set a random seed to make results reproducible)
with open("shakespeare_all.txt", "r") as in_file, \
open("shakespeare_train.txt", "w") as train_file, \
open("shakespeare_test.txt", "w") as test_file:
for line in in_file.readlines():
if line == "\n" or line == "":
continue # Let's leave out empty lines
if rng.uniform() < target_test_set_ratio:
test_file.write(line)
test_count += 1
else:
train_file.write(line)
train_count += 1
total_count = train_count + test_count
print(f"Total number of non-empty lines in the data set: {total_count}")
print(f"File `shakespeare_train.txt` has {train_count} lines ({100 * train_count / total_count:.1f}%).")
print(f"File `shakespeare_test.txt` has {test_count} lines ({100 * test_count / total_count:.1f}%).")
# -
# ## Step 3: Train The Model
#
# We'll use the toy model described in [this tutorial](https://github.com/spro/practical-pytorch/blob/master/char-rnn-generation/char-rnn-generation.ipynb).
# Luckily, someone has already extracted the relevant code blocks from the tutorial and built a command line application around it, so we'll just use that.
#
# Clone the repository into a subdirectory `char-rnn.pytorch` right next to this notebook:
# !git clone https://github.com/spro/char-rnn.pytorch
# At the time of writing, the `char-rnn.pytorch` repository seems to have a small bug that will result in an exception being raised when training the model (a pull request that fixes it [exists](https://github.com/spro/char-rnn.pytorch/pull/10)).
# But we can fix it easily.
# Open the file `char-rnn.pytorch/train.py` in a text editor, look for the line in the `train` function that reads
#
# ```python
# return loss.data[0] / args.chunk_len
# ```
#
# and replace "`[0]`" with "`.item()`" so that the line now reads:
#
# ```python
# return loss.data.item() / args.chunk_len
# ```
#
# If you can't find the original line then the bug has probably been fixed in the meantime.
#
# Now, start the training procedure.
# **This will take about 30 minutes** but you'll only have to do it once since the trained model will be saved to a file.
# !python char-rnn.pytorch/train.py shakespeare_train.txt
# The above training script should print a few lines of text after each completed 5% of training progress.
# The generated text snippets are random samples from the trained model.
# You should be able to observe that the quality of the sampled text should improve as training proceeds.
#
# At the end of the training cycle, the generated text will not be perfect, but that's OK for the purpose of compression.
# When we'll use the trained model to compress some new text below, we won't be blindly following the model's predictions as in the randomly generated text here.
# Instead, we'll compare the model's predictions with the actual text that we want to compress, and we'll then essentially encode the difference.
# Thus, model predictions don't need to be perfect; as long as they're better than a completely random (uniform) guess, they will allow us to reduce the bitrate.
# ## Step 4: While the Model is Being Trained ...
#
# While you wait for the model to be trained, let's start thinking about how we'll use the trained model for our compression method.
#
# ### Overall Encoder/Decoder Design
#
# The model is an autoregressive model that processes one character after the other.
# Each step takes the previous character as input, updates some internal hidden state, and outputs a probability distribution over all possible values for the next character.
#
# This autoregressive model architecture pretty much dictates how our compression method must operate:
#
# - The *encoder* reads the message that we want to compress character by character and updates the autoregressive model with each step.
# It uses the output of the model at each step to define an entropy model for encoding the next character.
# - The *decoder* decodes one character at a time and uses it to perform the exact same model updates as the encoder did.
# The entropy model used for each decoding step is defined by the model output from the previous step.
#
# The very first model update on both encoder and decoder side is a bit subtle because we don't yet have a "previous" character to provide as input for the model update here.
# We'll just make up a fake character that we pretend exists before the start of the actual message, and we'll inject this fake character at the very beginning to both the encoder and the decoder.
# Let's use the newline character `"\n"` for this purpose since this seems like a character that could indeed naturally precede the beginning of any message.
#
# Since encoding and decoding iterate over the characters in the same order, we'll need to use an entropy coder with *queue* semantics (i.e., "first in first out").
# The `constriction` library provides two entropy coders with queue semantics: [Range Coding](https://bamler-lab.github.io/constriction/apidoc/python/stream/queue.html) and [Huffman Coding](https://bamler-lab.github.io/constriction/apidoc/python/symbol.html).
# We'll use Range Coding here since it has better compression performance.
# At the end of this notebook you'll find an empirical comparison to Huffman Coding (which will turn out to perform worse).
#
# ### How to Implement Iterative Model Updates
#
# So how are we going to perform these iterative model updates described above in practice?
# Let's just start from how it's done in the the implementation of the model (which you cloned into the subdirectory `char-rnn.pytorch` in Step 3 above) and adapt it to our needs.
# The file `generate.py` seems like a good place to look for inspiration since generating random text from the model is not all that different from what we're trying to do: both have to update the model character by character, continuously obtaining a probability distribution over the next character.
# The only difference is that `generate.py` uses the obtained probability distribution to sample from it while we'll use it to define an entropy model.
#
# The function `generate` in `char-rnn.pytorch/generate.py` shows us how to do all the steps we need.
# The function takes a model as argument, which is—confusingly—called `decoder`.
# The function body starts by initializing a hidden state as follows,
#
# ```python
# hidden = decoder.init_hidden(1)
# ```
#
# After some more initializations, the function enters a loop that iteratively updates the model as follows:
#
# ```python
# output, hidden = decoder(inp, hidden)
# ```
#
# where `inp` is the "input", i.e., the previous character, represented (for technical reasons) as an integer PyTorch tensor of length `1`.
# The above model update returns `output` and the updated `hidden` state.
# On the next line of code, `output` gets scaled by some temperature and exponentiated before it is interpreted as an (unnormalized) probability distribution by being passed into `torch.multinomial`.
# Thus, `output` seems to be a tensor of logits, defining the probability distribution for the next character.
# Finally, after drawing a sample `top_i` from the `torch.multinomial` distribution, the function maps the sampled integer to a character as follows:
#
# ```python
# predicted_char = all_characters[top_i]
# ```
#
# Thus, there seems to be some fixed string `all_characters` that has the character with integer representation `i` at its `i`'th position.
# Let's bring this string into scope (and while we're at it, let's also import some other stuff we'll need below):
# +
import constriction
import torch
import numpy as np
from tqdm import tqdm
# Add subdirectory `char-rnn.pytorch` to Python's path so we can import some stuff from it.
import sys
import os
sys.path.append(os.path.join(os.getcwd(), "char-rnn.pytorch"))
from model import *
from helpers import read_file, all_characters
# -
# Try it out:
print(all_characters)
# ### Implement The Encoder
#
# We now have everything we need to know to implement the encoder and decoder.
# Let's start with the encoder, and define a function `compress_file` that takes the path to a file, compresses it, and writes the output to a different file.
# Since our compression method will turn out to be very slow (see introduction), we'll also introduce an optional argument `max_chars` that will allow the caller to compress only the first `max_chars` characters from the input file and stop after that.
# The function `compress_file` will also expect a `model` argument where the caller will have to pass in the trained model (this was called `decoder` in the file `generate.py` discussed above, but we'll call it `model` here to avoid confusion).
def compress_file(model, in_filename, out_filename, max_chars=None):
message, _ = read_file(in_filename) # (`read_file` defined in `char-rnn.pytorch/helpers.py`)
if max_chars is not None:
message = message[:max_chars] # Truncate message to at most `max_chars` characters.
# Initialize the hidden state and model input as discussed above:
hidden = model.init_hidden(1) # (same as in `generate.py` discussed above)
input_char = torch.tensor([all_characters.index('\n')], dtype=torch.int64) # "fake" character that we pretend precedes the message
# Instantiate an empty Range Coder onto which we'll accumulate compressed data:
encoder = constriction.stream.queue.RangeEncoder()
# Iterate over the message and encode it character by character, updating the model as we go:
for char in tqdm(message):
output, hidden = model(input_char, hidden) # update the model (as in `generate.py`)
# Turn the `output` into an entropy model and encode the character with it:
logits = output.data.view(-1)
logits = logits - logits.max() # "Log-Sum-Exp trick" for numerical stability
unnormalized_probs = logits.exp().numpy().astype(np.float64)
entropy_model = constriction.stream.model.Categorical(unnormalized_probs)
char_index = all_characters.index(char)
encoder.encode(char_index, entropy_model)
# Prepare for next model update:
input_char[0] = char_index
# Save the compressed data to a file
print(f"Compressed {len(message)} characters into {encoder.num_bits()} bits ({encoder.num_bits() / len(message):.2f} bits per character).")
compressed = encoder.get_compressed()
if sys.byteorder != "little":
# Let's always save data in the same byte order so compressed files can be transferred across computer architectures.
compressed.byteswap(inplace=True)
compressed.tofile(out_filename)
print(f'Wrote compressed data to file "{out_filename}".')
# The main part that distinguishes our function `compress_file` from the function `generate` in `generate.py` discussed above is that, after each model update, we use the model to encode an (already given) character rather than to sample a character.
# There are two slightly subtle steps here:
# first, we subtract the constant `logits.max()` from all elements of `logits` before exponentiating them.
# Such a global shift in logit-space has no effect (apart from rounding errors) on the resulting probability distribution since it will correspond to a global scaling factor after exponentiation.
# We perform this operation out of an abundance of caution to prevent numerical overflow when we exponentiate `logits` on the next line.
# Second, we construct the `Categorical` entropy model from a tensor of *unnormalized* probabilities.
# That's OK according to [the documentation](https://bamler-lab.github.io/constriction/apidoc/python/stream/model.html#constriction.stream.model.Categorical), `constriction` will have to make sure the distribution is exactly normalize in fixed-point arithmetic anyway.
#
# ### Implement The Decoder
#
# Let's also implement a function `decompress_file` that recovers the message from its compressed representation so that we can prove that the encoder did not discard any information.
# The decoder operates very similar to the encoder, except that it starts by loading compressed data from a file instead of the message, and it initializes a `RangeDecoder` from it, from which it then decodes one symbol at a time in the iteration.
# One important difference to the encoder is that, with our current autoregressive model, the decoder cannot detect the end of the message.
# Therefore, we have to provide the message length (`num_chars`) as an argument to the decoder function.
# In a real deployment, you'll probably want to either transmit the message length as an explicit initial symbol, or you could add an "End of File" sentinel symbol to the alphabet (`all_characters`) and append this symbol to the message on the encoder side to signal to the decoder that it should stop processing.
def decompress_file(model, in_filename, out_filename, num_chars):
# Load the compressed data into a `RangeDecoder`:`
compressed = np.fromfile(in_filename, dtype=np.uint32)
if sys.byteorder != "little":
compressed.byteswap(inplace=True) # restores native byte order ("endianness").
print(f"Loaded {32 * len(compressed)} bits of compressed data.")
decoder = constriction.stream.queue.RangeDecoder(compressed)
# Initialize the hidden state and model input exactly as in the encoder:
hidden = model.init_hidden(1) # (same as in `generate.py` discussed above)
input_char = torch.tensor([all_characters.index('\n')], dtype=torch.int64) # "fake" character that we pretend precedes the message
# Decode the message character by character, updating the model as we go:
with open(out_filename, "w") as out_file:
for _ in tqdm(range(num_chars)):
# Update model and optain (unnormalized) probabilities, exactly as in the encoder:
output, hidden = model(input_char, hidden)
logits = output.data.view(-1)
logits = logits - logits.max()
unnormalized_probs = logits.exp().numpy().astype(np.float64)
entropy_model = constriction.stream.model.Categorical(unnormalized_probs)
# This time, use the `entropy_model` for *decoding* to obtain the next character:
char_index = decoder.decode(entropy_model)
char = all_characters[char_index]
out_file.write(char)
# Prepare for next model update, exactly as in the encoder:
input_char[0] = char_index
print(f'Wrote decompressed data to file "{out_filename}".')
# ## Step 5: Try It Out
#
# If you've followed along and taken the time to understand the encoder/decoder implementation in Step 4 above then the model should have finished training by now.
# Load it into memory:
model = torch.load("shakespeare_train.pt")
# Now, try out if our implementation can indeed compress and decompress a text file with this model.
# We'll compress the *test* subset of our data set so that we test our method on text that was not used for training (albeit, admittedly, the test data is very similar to the training data since both were written by the same author):
compress_file(model, "shakespeare_test.txt", "shakespeare_test.txt.compressed", max_chars=10_000)
# If you didn't change anything in the training schedule then you should get a bitrate of about 2.1 bits per character.
# Before we compare this bitrate to that of general-purpose compression methods, let's first verify that the compression method is actually correct.
# Decode the compressed file again:
decompress_file(model, "shakespeare_test.txt.compressed", "shakespeare_test.txt.decompressed", 10_000)
# Let's take a quick peak in the original and reconstructed text:
# !head shakespeare_test.txt shakespeare_test.txt.decompressed
# The beginnings certainly look similar.
# But let's check more thoroughly.
# Remember that we only encoded and decoded the first 10,000 characters of the test data, so that's what we have to compare to (note: the test turns out to be pure ASCII, so the first 10,000 characters map exactly to the first 10,000 bytes):
#
# !head -c 10000 shakespeare_test.txt | diff - shakespeare_test.txt.decompressed # If this prints no output we're good.
# ## Step 6: Evaluation
#
# There's a lot we could analyze now:
#
# - How do the bitrates of our method compare to general-purpose compression methods like `gzip`, `bzip2`, and `xz`?
# - How well does our method generalize to other text, ranging from other English text by a different author all the way to text in a different language?
# - Where do the encoder and decoder spend most of their runtime?
#
# We'll just address the first question here and leave the others to the reader.
# Let's compress the same first 10,000 characters of the test data with `gzip`, `bzip2`, and `xz` (if installed on your system):
# !head -c 10000 shakespeare_test.txt | gzip --best > shakespeare_test.txt.gzip
# !head -c 10000 shakespeare_test.txt | bzip2 --best > shakespeare_test.txt.bz2
# !head -c 10000 shakespeare_test.txt | xz --best > shakespeare_test.txt.xz
# Then let's compare the sizes of the compressed files:
# !ls -l shakespeare_test.txt.*
# Despite using a very simple model that we took from a tutorial completely unrelated to data compression, our compression method reduces the bitrate compared to `bzip2` by 40%.
# Of course, we shouldn't read too much into this since we trained the model on data that is very similar to the test data.
# ## Bonus 1: Getting It *Almost* Right and Yet Fatally Wrong
#
# The [API reference for `constriction`'s entropy models](https://bamler-lab.github.io/constriction/apidoc/python/stream/model.html) highlights that entropy models are brittle: event tiny discrepancies in rounding operations between encoder and decoder can have catastrophic effects for entropy coding.
# The models provided by `constriction` are implemented in exact fixed point arithmetic to allow for well-defined and consistent rounding operations when, e.g., inverting the CDF.
# However, `constriction` can only guarantee consistent rounding operations in its internal operations.
# You have to ensure yourself that any probabilities you provide to `constriction` are *exactly* the same on the encoder and decoder side.
#
# The following example illustrates how even tiny discrepancies between rounding operations on the encoder and decoder side can completely derail the entropy coder.
# Recall that, in both functions `compress_file` and `decompress_file` above, we subtract `logits.max()` from `logits` before we exponentiate them.
# This can prevent numerical overflow in the exponentiation but one might expect that it has otherwise no effect on the resulting probability distribution since it only leads to a global scaling of `unnormalized_probs`, which should drop out once `constriction` normalizes the probabilities—*except* that this is not quite correct:
# even if there's no numerical overflow, the different scaling affects all subsequent rounding operations that are implicitly performed by the CPU in any floating point operation.
# In and of itself, these implicit rounding operations are not a big issue and unavoidable in floating point calculations.
# However, they do become an issue when they are done inconsistently between the encoder and the decoder, as we show in the next example.
#
# Let's keep the encoder as it is, but let's slightly modify the decoder by commenting out the line that subtracts `logits.max()` from `logits`, as highlighted by the string `<--- COMMENTED OUT` in the following example:
def decompress_file_almost_right(model, in_filename, out_filename, num_chars):
# Load the compressed data into a `RangeDecoder`:`
compressed = np.fromfile(in_filename, dtype=np.uint32)
if sys.byteorder != "little":
compressed.byteswap(inplace=True) # restores native byte order ("endianness").
print(f"Loaded {32 * len(compressed)} bits of compressed data.")
decoder = constriction.stream.queue.RangeDecoder(compressed)
# Initialize the hidden state and model input exactly as in the encoder:
hidden = model.init_hidden(1) # (same as in `generate.py` discussed above)
input_char = torch.tensor([all_characters.index('\n')], dtype=torch.int64) # "fake" character that we pretend precedes the message
# Decode the message character by character, updating the model as we go:
with open(out_filename, "w") as out_file:
for _ in tqdm(range(num_chars)):
# Update model and optain (unnormalized) probabilities, exactly as in the encoder:
output, hidden = model(input_char, hidden)
logits = output.data.view(-1)
# logits = logits - logits.max() <--- COMMENTED OUT
unnormalized_probs = logits.exp().numpy().astype(np.float64)
entropy_model = constriction.stream.model.Categorical(unnormalized_probs)
# This time, use the `entropy_model` for *decoding* to obtain the next character:
char_index = decoder.decode(entropy_model)
char = all_characters[char_index]
out_file.write(char)
# Prepare for next model update, exactly as in the encoder:
input_char[0] = char_index
print(f'Wrote decompressed data to file "{out_filename}".')
# If we use this slightly modified decoder to decode data that was encoded with the original encoder `compress_file`, we *might* run into an issue.
#
# **Note:** The following example may or may not work on your setup, depending on the random seeds used for training the model.
# But if it fails, as in my setup below, then it will fail catastrophically and either throw an error (as it does here) or (if we're not quite as lucky) silently continue decoding but decode complete gibberish after some point in the stream.
decompress_file_almost_right(model, "shakespeare_test.txt.compressed", "shakespeare_test.txt.decompressed_wrong", 10_000)
# Notice that we were able to decode the first 883 characters just fine (you might get a different number here).
# Issues due to tiny discrepancies in rounding operations are very unlikely to occur, but when they occur in an entropy coder, they are fatal.
# We actually got lucky here: it's also possible that the decoder does not detect any errors but that it starts decoding complete gibberish after some point (in fact, had we used an ANS coder instead of a Range Coder, then decoding would have been infallible but could still produce wrong results when misused).
#
# Due to this brittleness, entropy models have to be implemented with care, and we consider `constriction`'s implementations of entropy models an important part of the library, in addition to `constriction`'s entropy coders.
# Yet, as the above example shows, even the most careful implementation of an entropy model cannot protect from errors when misused.
# ## Bonus 2: Huffman Coding
#
# Our above compression method uses Range Coding for the entropy coder.
# The `constriction` library also provides another entropy coder with "queue" semantics: Huffman coding.
# The API for Huffman coding is somewhat different to that of Range Coding because of the very different nature of the two algorithms, but it's easy to port our encoder and decoder to Huffman coding:
#
# ### Encoder
def compress_file_huffman(model, in_filename, out_filename, max_chars=None):
message, _ = read_file(in_filename) # (`read_file` defined in `char-rnn.pytorch/helpers.py`)
if max_chars is not None:
message = message[:max_chars] # Truncate message to at most `max_chars` characters.
# Initialize the hidden state and model input as discussed above:
hidden = model.init_hidden(1) # (same as in `generate.py` discussed above)
input_char = torch.tensor([all_characters.index('\n')], dtype=torch.int64) # "fake" character that we pretend precedes the message
# Instantiate an empty `QueueEncoder` onto which we'll accumulate compressed data:
encoder = constriction.symbol.QueueEncoder() # <-- CHANGED LINE
# Iterate over the message and encode it character by character, updating the model as we go:
for char in tqdm(message):
output, hidden = model(input_char, hidden) # update the model (as in `generate.py`)
# Turn the `output` into an entropy model and encode the character with it:
logits = output.data.view(-1)
logits = logits - logits.max() # "Log-Sum-Exp trick" for numerical stability
unnormalized_probs = logits.exp().numpy().astype(np.float64)
codebook = constriction.symbol.huffman.EncoderHuffmanTree(unnormalized_probs) # <-- CHANGED LINE
char_index = all_characters.index(char)
encoder.encode_symbol(char_index, codebook) # <-- CHANGED LINE
# Prepare for next model update:
input_char[0] = char_index
# Save the compressed data to a file
compressed, num_bits = encoder.get_compressed() # <-- CHANGED LINE
print(f"Compressed {len(message)} characters into {num_bits} bits ({num_bits / len(message):.2f} bits per character).")
if sys.byteorder != "little":
# Let's always save data in the same byte order so compressed files can be transferred across computer architectures.
compressed.byteswap(inplace=True)
compressed.tofile(out_filename)
print(f'Wrote compressed data to file "{out_filename}".')
# ### Decoder
def decompress_file_huffman(model, in_filename, out_filename, num_chars):
# Load the compressed data into a `RangeDecoder`:`
compressed = np.fromfile(in_filename, dtype=np.uint32)
if sys.byteorder != "little":
compressed.byteswap(inplace=True) # restores native byte order ("endianness").
print(f"Loaded {32 * len(compressed)} bits of compressed data.")
decoder = constriction.symbol.QueueDecoder(compressed) # <-- CHANGED LINE
# Initialize the hidden state and model input exactly as in the encoder:
hidden = model.init_hidden(1) # (same as in `generate.py` discussed above)
input_char = torch.tensor([all_characters.index('\n')], dtype=torch.int64) # "fake" character that we pretend precedes the message
# Decode the message character by character, updating the model as we go:
with open(out_filename, "w") as out_file:
for _ in tqdm(range(num_chars)):
# Update model and optain (unnormalized) probabilities, exactly as in the encoder:
output, hidden = model(input_char, hidden)
logits = output.data.view(-1)
logits = logits - logits.max()
unnormalized_probs = logits.exp().numpy().astype(np.float64)
codebook = constriction.symbol.huffman.DecoderHuffmanTree(unnormalized_probs) # <-- CHANGED LINE
# This time, use the `codebook` for *decoding* to obtain the next character:
char_index = decoder.decode_symbol(codebook) # <-- CHANGED LINE
char = all_characters[char_index]
out_file.write(char)
# Prepare for next model update, exactly as in the encoder:
input_char[0] = char_index
print(f'Wrote decompressed data to file "{out_filename}".')
# ### Try it Out Again
compress_file_huffman(model, "shakespeare_test.txt", "shakespeare_test.txt.compressed-huffman", max_chars=10_000)
# For comparison: when we used a Range Coder we got a better bitrate of only about 2.1 bits per character.
decompress_file_huffman(model, "shakespeare_test.txt.compressed-huffman", "shakespeare_test.txt.decompressed-huffman", 10_000)
# Verify correctness again:
# !head -c 10000 shakespeare_test.txt | diff - shakespeare_test.txt.decompressed-huffman # If this prints no output we're good.
# ## Conclusion
#
# We've discussed how you can use `constriction`'s entropy coders with an entropy model that is an autoregressive machine-learning model.
# Autoregressive models allow you to model correlations between symbols, and exploit them to improve compression performance.
# An alternative method for exploiting correlations in data compression is the so-called bits-back technique, which applies to latent variable models that tend to be better parallelizable than autoregressive models.
# An example of bits-back coding with `constriction` is provided in [this problem set](https://robamler.github.io/teaching/compress21/problem-set-05.zip) (with [solutions](https://robamler.github.io/teaching/compress21/problem-set-05-solutions.zip)).
|
examples/python/03-tutorial-autoregressive-nlp-compression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
COLLECTION_NAME = "SVS"
BUCKET_WIDTH = 250
TOKENS = 8888
sales = pd.read_csv("../data/pre-reveal_sales.csv")
rarity_dict = {}
bins = int(TOKENS / BUCKET_WIDTH)
total_sales = 0
for i in range(0, bins):
upper = (i + 1) * BUCKET_WIDTH
lower = i * BUCKET_WIDTH
hist_bin = sales[ (sales["RANK"] > lower) & (sales["RANK"] < upper)]
rarity_dict[lower] = hist_bin["PRICE"].sum() / len(hist_bin)
plt.bar(rarity_dict.keys(), rarity_dict.values(), width = 50, color='g', align = "edge", edgecolor ='black')
plt.title("Pre-reveal Sales by Rarity - {}".format(COLLECTION_NAME))
plt.xlabel("Rarity Rank (Bucket Width = {} Tokens)".format(BUCKET_WIDTH), fontsize = 10)
plt.ylabel("Average Price (ETH)", fontsize = 10)
plt.savefig('../figures/{}_average_price_vs_rank.png'.format(COLLECTION_NAME), dpi = 300)
plt.show()
# -
|
case_studies/SVS/code/average_price.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Last updated on Apr 9, 2020
# # Demo script for the analyses done in Nakamura and Huang (2018, Science)
#
# This is a complimentary demo script that can be used to implement the local wave activity, fluxes and flux convergence/divergence computation required in the analyses presented in Nakamura and Huang, Atmospheric Blocking as a Traffic Jam in the Jet Stream. Science. (2018)
#
# This notebook demonstrate how to compute local wave activity and all the flux terms in equations (2) and (3) in NH2018 with the updated functionality in the python package `hn2016_falwa`. To run the script, please install the
# package `hn2016_falwa` using
# ```
# python setup.py develop
# ```
# after cloning the [GitHub repo](http://github.com/csyhuang/hn2016_falwa).
#
# The functionalities are enhanced and included in the class object `QGField` under `hn2016_falwa.oopinterface`. Please refer to the [documentation](http://hn2016-falwa.readthedocs.io/) (search `QGField`) or the end of this notebook for the input/methods this class provides.
#
# Please [raise an issue in the GitHub repo](https://github.com/csyhuang/hn2016_falwa/issues) or contact <NAME> (<EMAIL>) if you have any questions or suggestions regarding the package.
import numpy as np
from numpy import dtype
from math import pi
from netCDF4 import Dataset
import matplotlib.pyplot as plt
import datetime as dt
# %matplotlib inline
from hn2016_falwa.oopinterface import QGField
import hn2016_falwa.utilities as utilities
import datetime as dt
# # Load ERA-Interim reanalysis data retrieved from ECMWF server
# The sample script in this directory `download_example.py` include the code to retrieve zonal wind field U, meridional
# wind field V and temperature field T at various pressure levels. Given that you have an account on ECMWF server and
# have the `ecmwfapi` package installed, you can run the scripts to download data from there:
# ```
# python download_example.py
# ```
# +
# --- Load the zonal wind and QGPV at 240hPa --- #
u_file = Dataset('2005-01-23_to_2005-01-30_u.nc', mode='r')
v_file = Dataset('2005-01-23_to_2005-01-30_v.nc', mode='r')
t_file = Dataset('2005-01-23_to_2005-01-30_t.nc', mode='r')
time_array = u_file.variables['time'][:]
time_units = u_file.variables['time'].units
time_calendar = u_file.variables['time'].calendar
ntimes = time_array.shape[0]
print('Dimension of time: {}'.format(time_array.size))
# -
# # Load the dimension arrays
# In this version, the `QGField` object takes only:
# - latitude array in degree ascending order, and
# - pressure level in hPa in decending order (from ground to aloft).
# +
xlon = u_file.variables['longitude'][:]
# latitude has to be in ascending order
ylat = u_file.variables['latitude'][:]
if np.diff(ylat)[0]<0:
print('Flip ylat.')
ylat = ylat[::-1]
# pressure level has to be in descending order (ascending height)
plev = u_file.variables['level'][:]
if np.diff(plev)[0]>0:
print('Flip plev.')
plev = plev[::-1]
nlon = xlon.size
nlat = ylat.size
nlev = plev.size
# -
clat = np.cos(np.deg2rad(ylat)) # cosine latitude
p0 = 1000. # surface pressure [hPa]
kmax = 49 # number of grid points for vertical extrapolation (dimension of height)
dz = 1000. # differential height element
height = np.arange(0,kmax)*dz # pseudoheight [m]
dphi = np.diff(ylat)[0]*pi/180. # differential latitudinal element
dlambda = np.diff(xlon)[0]*pi/180. # differential latitudinal element
hh = 7000. # scale height
cp = 1004. # heat capacity of dry air
rr = 287. # gas constant
omega = 7.29e-5 # rotation rate of the earth
aa = 6.378e+6 # earth radius
prefactor = np.array([np.exp(-z/hh) for z in height[1:]]).sum() # integrated sum of density from the level
#just above the ground (z=1km) to aloft
npart = nlat # number of partitions to construct the equivalent latitude grids
maxits = 100000 # maximum number of iteration in the SOR solver to solve for reference state
tol = 1.e-5 # tolerance that define convergence of solution
rjac = 0.95 # spectral radius of the Jacobi iteration in the SOR solver.
jd = nlat//2+1 # (one plus) index of latitude grid point with value 0 deg
# This is to be input to fortran code. The index convention is different.
# # Create a netCDF file to store output
#
# A netCDF file `2005-01-23_to_2005-01-30_output.nc` with same number of time steps in the input file is created to store all the computed quantities.
# === Outputing files ===
output_fname = '2005-01-23_to_2005-01-30_output.nc'
output_file = Dataset(output_fname, 'w')
output_file.createDimension('levelist',kmax)
output_file.createDimension('latitude',nlat)
output_file.createDimension('longitude',nlon)
output_file.createDimension('time',ntimes)
plevs = output_file.createVariable('levelist',dtype('float32').char,('levelist',)) # Define the coordinate variables
lats = output_file.createVariable('latitude',dtype('float32').char,('latitude',)) # Define the coordinate variables
lons = output_file.createVariable('longitude',dtype('float32').char,('longitude',))
times = output_file.createVariable('time',dtype('int').char,('time',))
plevs.units = 'hPa'
lats.units = 'degrees_north'
lons.units = 'degrees_east'
times.units = time_units
times.calendar = time_calendar
plevs[:] = p0 * np.exp(-height/hh)
lats[:] = ylat
lons[:] = xlon
times[:] = time_array
qgpv = output_file.createVariable('qgpv',dtype('float32').char,('time','levelist','latitude','longitude'))
qgpv.units = '1/s'
interpolated_u = output_file.createVariable('interpolated_u',dtype('float32').char,('time','levelist','latitude','longitude'))
interpolated_u.units = 'm/s'
interpolated_v = output_file.createVariable('interpolated_v',dtype('float32').char,('time','levelist','latitude','longitude'))
interpolated_v.units = 'm/s'
interpolated_theta = output_file.createVariable('interpolated_theta',dtype('float32').char,('time','levelist','latitude','longitude'))
interpolated_theta.units = 'K'
qref = output_file.createVariable('qref',dtype('float32').char,('time','levelist','latitude'))
qref.units = '1/s'
uref = output_file.createVariable('uref',dtype('float32').char,('time','levelist','latitude'))
uref.units = 'm/s'
ptref = output_file.createVariable('ptref',dtype('float32').char,('time','levelist','latitude'))
ptref.units = 'K'
lwa = output_file.createVariable('lwa',dtype('float32').char,('time','levelist','latitude','longitude'))
lwa.units = 'm/s'
adv_flux_f1 = output_file.createVariable('Zonal advective flux F1',dtype('float32').char,('time','latitude','longitude'))
adv_flux_f1.units = 'm**2/s**2'
adv_flux_f2 = output_file.createVariable('Zonal advective flux F2',dtype('float32').char,('time','latitude','longitude'))
adv_flux_f2.units = 'm**2/s**2'
adv_flux_f3 = output_file.createVariable('Zonal advective flux F3',dtype('float32').char,('time','latitude','longitude'))
adv_flux_f3.units = 'm**2/s**2'
adv_flux_conv = output_file.createVariable('Zonal advective flux Convergence -Div(F1+F2+F3)',dtype('float32').char,('time','latitude','longitude'))
adv_flux_conv.units = 'm/s**2'
divergence_eddy_momentum_flux = output_file.createVariable('Eddy Momentum Flux Divergence',dtype('float32').char,('time','latitude','longitude'))
divergence_eddy_momentum_flux.units = 'm/s**2'
meridional_heat_flux = output_file.createVariable('Low-level Meridional Heat Flux',dtype('float32').char,('time','latitude','longitude'))
meridional_heat_flux.units = 'm/s**2'
lwa_baro = output_file.createVariable('lwa_baro',dtype('float32').char,('time','latitude','longitude'))
lwa_baro.units = 'm/s'
u_baro = output_file.createVariable('u_baro',dtype('float32').char,('time','latitude','longitude'))
u_baro.units = 'm/s'
# # Set the level of pressure and the timestamp to display below
tstamp = [dt.datetime(2005,1,23,0,0) + dt.timedelta(seconds=6*3600) * tt for tt in range(ntimes)]
plev_selected = 10 # selected pressure level to display
tstep_selected = 0
# # Set names of the variables to display
# # Loop through the input file and store all the computed quantities in a netCDF file
# +
for tstep in range(32): # or ntimes
uu = u_file.variables['u'][tstep, ::-1, ::-1, :].data
vv = v_file.variables['v'][tstep, ::-1, ::-1, :].data
tt = t_file.variables['t'][tstep, ::-1, ::-1, :].data
qgfield_object = QGField(xlon, ylat, plev, uu, vv, tt)
qgpv[tstep, :, :, :], interpolated_u[tstep, :, :, :], interpolated_v[tstep, :, :, :], \
interpolated_theta[tstep, :, :, :], static_stability = qgfield_object.interpolate_fields()
qref[tstep, :, :], uref[tstep, :, :], ptref[tstep, :, :] = \
qgfield_object.compute_reference_states(northern_hemisphere_results_only=False)
adv_flux_f1[tstep, :, :], \
adv_flux_f2[tstep, :, :], \
adv_flux_f3[tstep, :, :], \
adv_flux_conv[tstep, :, :], \
divergence_eddy_momentum_flux[tstep, :, :], \
meridional_heat_flux[tstep, :, :], \
lwa_baro[tstep, :, :], \
u_baro[tstep, :, :], \
lwa[tstep, :, :, :] \
= qgfield_object.compute_lwa_and_barotropic_fluxes(northern_hemisphere_results_only=False)
if tstep == tstep_selected:
# === Below demonstrate another way to access the computed variables ===
# 3D Variables that I would choose one pressure level to display
variables_3d = [
(qgfield_object.qgpv, 'Quasigeostrophic potential vorticity (QGPV)'),
(qgfield_object.lwa, 'Local wave activity (LWA)'),
(qgfield_object.interpolated_u, 'Interpolated zonal wind (u)'),
(qgfield_object.interpolated_v, 'Interpolated meridional wind (v)')]
# Reference states to be displayed on y-z plane
variables_yz = [
(qgfield_object.qref, 'Qref'),
(qgfield_object.uref, 'Uref'),
(qgfield_object.ptref, 'PTref')]
# Vertically averaged variables to be displayed on x-y plane
variables_xy = [
(qgfield_object.adv_flux_f1, 'Advective flux F1'),
(qgfield_object.adv_flux_f2, 'Advective flux F2'),
(qgfield_object.adv_flux_f3, 'Advective flux F3'),
(qgfield_object.convergence_zonal_advective_flux, 'Advective flux convergence -Div(F1+F2+F3)'),
(qgfield_object.divergence_eddy_momentum_flux, 'divergence_eddy_momentum_flux'),
(qgfield_object.meridional_heat_flux, 'meridional_heat_flux')
]
# Plot 240 hPa of 3D-variables
for variable, name in variables_3d:
plt.figure(figsize=(12,6))
plt.contourf(xlon, ylat[1:-1], variable[plev_selected, 1:-1, :], 50, cmap='jet')
if name=='Local wave activity (LWA)':
plt.axhline(y=0, c='w', lw=30)
plt.colorbar()
plt.ylabel('Latitude (deg)')
plt.xlabel('Longitude (deg)')
plt.title(name + ' at 240hPa | ' + str(tstamp[tstep]))
plt.show()
# Plot reference states
for variable, name in variables_yz:
plt.figure(figsize=(6,4))
plt.contourf(ylat[1:-1], height, variable[:, 1:-1], 50, cmap='jet')
plt.axvline(x=0, c='w', lw=2)
plt.xlabel('Latitude (deg)')
plt.ylabel('Pseudoheight (m)')
plt.colorbar()
plt.title(name + ' | ' + str(tstamp[tstep]))
plt.show()
# Plot barotropic (2D-)variables
for variable, name in variables_xy:
plt.figure(figsize=(12,6))
plt.contourf(xlon, ylat[1:-1], variable[1:-1, :], 50, cmap='jet')
plt.axhline(y=0, c='w', lw=30)
plt.ylabel('Latitude (deg)')
plt.xlabel('Longitude (deg)')
plt.colorbar()
plt.title(name + ' | ' + str(tstamp[tstep]))
plt.show()
print('tstep = {}/{}\n'.format(tstep, ntimes))
output_file.close()
print('Output {} timesteps of data to the file {}'.format(tstep + 1, output_fname))
|
examples/nh2018_science/demo_script_for_nh2018.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="lMWLFva9WIaG"
import os
import datetime
import IPython
import IPython.display
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
from keras import Input
from keras.models import Sequential
from keras.layers import Flatten
from keras.layers import Dense, LSTM, GRU
from keras.layers import Conv1D
from keras.layers import MaxPooling1D
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
mpl.rcParams['figure.figsize'] = (8, 6)
mpl.rcParams['axes.grid'] = False
# + id="b0wRRC1MWI_O"
zip_path = tf.keras.utils.get_file(
origin='https://storage.googleapis.com/tensorflow/tf-keras-datasets/jena_climate_2009_2016.csv.zip',
fname='jena_climate_2009_2016.csv.zip',
extract=True)
csv_path, _ = os.path.splitext(zip_path)
# + id="oS_-CU_-WJBi"
df = pd.read_csv(csv_path)
# slice [start:stop:step], starting from index 5 take every 6th record.
df = df[5::6]
date_time = pd.to_datetime(df.pop('Date Time'), format='%d.%m.%Y %H:%M:%S')
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="ydWH0hKfWJD0" outputId="d3c3e6b8-a531-4ee0-8cb8-13f6d11685cc"
df
# + id="3qVk45uXWJIX"
# take three day of datas to predict one day
timestep = 72
# size of training data - 500 days (12000 hours)
training_num = 12000
epoch = 10
batch_size = 200
# + colab={"base_uri": "https://localhost:8080/"} id="WVcWIHxFWJM5" outputId="e7c15458-bbad-4571-bf16-d0a7bef1b8b8"
dt = df['T (degC)']
dt = np.array(dt).reshape(-1,1)
dt.shape
# + [markdown] id="78Tb9hAC6zEC"
# # <font color = purple>Normalize the data to (0, 1)</font>
# + colab={"base_uri": "https://localhost:8080/"} id="dD0c9Lj1hNxV" outputId="b44dd060-5552-4ad8-8b82-b34121ed8d2c"
sc = MinMaxScaler(feature_range = (0, 1))
dt_nor = sc.fit_transform(dt)
dt_nor
# + [markdown] id="_gTGzxOL9M6C"
# # <font color = purple>Find the best width</font>
# + id="zdLlPJOk9RX6"
# take three day of datas to predict one day
timestep
# size of training data - 500 days
training_num = 12000
epoch = 10
batch_size = 200
def width(timestep,model_kind):
xTrainSet = dt_nor[:training_num]
yTrainSet = dt_nor[1:training_num+1]
xTrain = []
for i in range(timestep, training_num):
xTrain.append(xTrainSet[i-timestep : i])
xTrain = np.array(xTrain)
xTrain = np.reshape(xTrain, (xTrain.shape[0], xTrain.shape[1], 1))
yTrain = []
for i in range(timestep, training_num):
yTrain.append(yTrainSet[i])
yTrain = np.array(yTrain)
if model_kind == 'model_rnn':
model = Sequential()
model.add(LSTM(128, return_sequences = True, input_shape = (xTrain.shape[1],1)))
model.add(GRU(64))
model.add(Dense(1))
if model_kind == 'model_dense':
model = Sequential()
model.add(Input(shape = (xTrain.shape[1])))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(64, activation='relu'))
model.add(Dense(1))
if model_kind == 'model_cnn':
conv_width = 3
model = Sequential()
model.add(Conv1D(64, kernel_size=(conv_width), input_shape = (xTrain.shape[1],1), activation='relu'))
# model.add(MaxPooling1D(pool_size=(8)))
model.add(Conv1D(32, kernel_size=(conv_width), activation='relu'))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(1))
model.compile(optimizer = 'adam',
loss = 'mean_squared_error',
metrics = [tf.metrics.MeanAbsoluteError()])
model.fit(x = xTrain, y = yTrain, epochs = epoch, batch_size = batch_size, verbose=0)
xTestSet = dt_nor[training_num : 40800-2]
xTestSet = np.array(xTestSet)
yTestSet = dt_nor[training_num+1 : 40800-1]
yTestSet = np.array(yTestSet)
xTest = []
for i in range(timestep, len(xTestSet)):
xTest.append(xTestSet[i-timestep : i])
xTest = np.array(xTest)
yTest = []
for i in range(timestep, len(xTestSet)):
yTest.append(yTestSet[i])
yTest = np.array(yTest)
yTest = sc.inverse_transform(yTest)
yPredictes = model.predict(x=xTest)
yPredictes = sc.inverse_transform(yPredictes)
r2 = r2_score(yTest, yPredictes)
return r2
# + colab={"base_uri": "https://localhost:8080/"} id="32QZ0loO-unc" outputId="c0763375-4400-4c52-a84e-545b26e891fa"
rnn_width_dict = {}
for step in range(5,51):
rnn_width_dict[step] = width(step,'model_rnn')
print(step,end="-")
if step%10 == 0:
print()
rnn_width = max(rnn_width_dict,key=rnn_width_dict.get)
rnn_width
# + colab={"base_uri": "https://localhost:8080/"} id="1DQuIqJoHrvP" outputId="aa229652-5cce-46c6-880f-55bb73d696f7"
dense_width_dict = {}
for step in range(5,51):
dense_width_dict[step] = width(step,'model_dense')
print(step,end="-")
if step%10 == 0:
print()
dense_width = max(dense_width_dict,key=dense_width_dict.get)
dense_width
# + colab={"base_uri": "https://localhost:8080/"} id="X_ys9ObeHr5O" outputId="7d244c23-a772-421d-f486-eb3302fc8b00"
cnn_width_dict = {}
for step in range(5,51):
cnn_width_dict[step] = width(step,'model_cnn')
print(step,end="-")
if step%10 == 0:
print()
cnn_width = max(cnn_width_dict,key=cnn_width_dict.get)
cnn_width
# + [markdown] id="Hyjbp5_Oau0G"
# # <font color = purple>Rnn</font>
# + id="MtrN2VVVFZuR"
timestep = rnn_width
# + id="Xd2y306oaB8U"
xTrainSet = dt_nor[:training_num]
yTrainSet = dt_nor[1:training_num+1]
# + colab={"base_uri": "https://localhost:8080/"} id="iKn3ZKbKWJRZ" outputId="cfb53cf7-9e80-4443-fb03-cee1f45e3c42"
xTrain = []
for i in range(timestep, training_num):
xTrain.append(xTrainSet[i-timestep : i])
xTrain = np.array(xTrain)
xTrain = np.reshape(xTrain, (xTrain.shape[0], xTrain.shape[1], 1))
print(xTrain.shape)
yTrain = []
for i in range(timestep, training_num):
yTrain.append(yTrainSet[i])
yTrain = np.array(yTrain)
print(yTrain.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="iLCdrDaSKFoQ" outputId="f104564f-e608-4b55-904c-d252c4952dca"
model_rnn = Sequential()
model_rnn.add(LSTM(128, return_sequences = True, input_shape = (xTrain.shape[1],1)))
model_rnn.add(GRU(64))
model_rnn.add(Dense(1))
model_rnn.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="nfTQXBO8dTke" outputId="666dae66-1f50-4d3b-8464-3e7e0921fc1a"
model_rnn.compile(optimizer = 'adam',
loss = 'mean_squared_error',
metrics = [tf.metrics.MeanAbsoluteError()])
model_rnn.fit(x = xTrain, y = yTrain, epochs = epoch, batch_size = batch_size)
# + [markdown] id="cP8Pn4zJ7vw7"
# # <font color = purple>Test model's accuracy by r2_score (1200 days)</font>
# + id="EzxEdlTPiTE5"
xTestSet = dt_nor[training_num : 40800-2]
xTestSet = np.array(xTestSet)
yTestSet = dt_nor[training_num+1 : 40800-1]
yTestSet = np.array(yTestSet)
# + colab={"base_uri": "https://localhost:8080/"} id="KQbW3X54iTIF" outputId="8a476ac7-e0a3-47ac-80cd-4ff47945eb3e"
xTest = []
for i in range(timestep, len(xTestSet)):
xTest.append(xTestSet[i-timestep : i])
xTest = np.array(xTest)
print(len(xTest))
yTest = []
for i in range(timestep, len(xTestSet)):
yTest.append(yTestSet[i])
yTest = np.array(yTest)
yTest = sc.inverse_transform(yTest)
len(yTest)
# + colab={"base_uri": "https://localhost:8080/"} id="_tTE75kxiTNf" outputId="657f6094-59a2-4af2-c990-7e828d9aeba9"
yPredictes = model_rnn.predict(x=xTest)
yPredictes = sc.inverse_transform(yPredictes)
yPredictes
# + id="UcSj9Hg_HMpC"
r2_value = {}
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="o8VQ7teLp0lB" outputId="640a90e4-a316-460e-d979-b2496f470f25"
plt.plot(yTest, 'c-', label='Real')
plt.plot(yPredictes, 'm-', label='Predict')
# plt.plot(data_original, color='red', label='Real')
# plt.plot(range(len(y_train)),yPredicts, color='blue', label='Predict')
plt.title(label='Prediction')
plt.xlabel(xlabel='Time')
plt.ylabel(ylabel='T')
plt.legend()
plt.show()
r2 = r2_score(yTest, yPredictes)
r2_value['RNN'] = r2
print(r2)
# + [markdown] id="VZfrf8pP3nit"
# # <font color=purple>Dense</font>
# + id="zYI4xYFK3vOg"
timestep = dense_width
# + id="Z1azZOyM3vQ3"
xTrainSet = dt_nor[:training_num]
yTrainSet = dt_nor[1:training_num+1]
# + colab={"base_uri": "https://localhost:8080/"} id="E-VUIv-93vTS" outputId="c46ba3c1-9fd0-4208-c67d-631dce50a2e3"
xTrain = []
for i in range(timestep, training_num):
xTrain.append(xTrainSet[i-timestep : i])
xTrain = np.array(xTrain)
#xTrain = np.squeeze(xTrain)
xTrain = np.reshape(xTrain, (xTrain.shape[0], xTrain.shape[1], 1))
print(xTrain.shape)
yTrain = []
for i in range(timestep, training_num):
yTrain.append(yTrainSet[i])
yTrain = np.array(yTrain)
#yTrain = np.reshape(yTrain, (yTrain.shape[0], 1))
print(yTrain.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="GUGkjUTkKN9I" outputId="319aed0f-9e21-4ce0-9461-e01ac9a3c65e"
model_dense = Sequential()
model_dense.add(Input(shape = (xTrain.shape[1])))
model_dense.add(Flatten())
model_dense.add(Dense(128, activation='relu'))
model_dense.add(Dense(64, activation='relu'))
model_dense.add(Dense(1))
model_dense.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="Y3gAF-8L3vYC" outputId="00daeeee-fb43-4590-b421-06b71cdedde8"
model_dense.compile(optimizer = 'adam',
loss = 'mean_squared_error',
metrics = [tf.metrics.MeanAbsoluteError()])
model_dense.fit(x = xTrain, y = yTrain, epochs = epoch, batch_size = batch_size)
# + [markdown] id="AXz4r4Gk7g-y"
# # <font color=purple>Test model's accuracy by r2_score (1200 days)</font>
# + id="QBPdG6rf3vaq"
xTestSet = dt_nor[training_num : 40800-2]
xTestSet = np.array(xTestSet)
yTestSet = dt_nor[training_num+1 : 40800-1]
yTestSet = np.array(yTestSet)
# + colab={"base_uri": "https://localhost:8080/"} id="A6q9mX1i6Eke" outputId="69490693-930e-49d1-8c25-f9f41ac87b2f"
xTest = []
for i in range(timestep, len(xTestSet)):
xTest.append(xTestSet[i-timestep : i])
xTest = np.array(xTest)
#xTest = np.squeeze(xTest)
yTest = []
for i in range(timestep, len(xTestSet)):
yTest.append(yTestSet[i])
yTest = np.array(yTest)
yTest = sc.inverse_transform(yTest)
len(xTest)
# + colab={"base_uri": "https://localhost:8080/"} id="sP2mCgPv8fm5" outputId="6acb498f-5b8d-4add-84df-7d1c29abe4ef"
yTest.shape
# + colab={"base_uri": "https://localhost:8080/"} id="B1R_0pKF6Em0" outputId="c584f58b-2a4e-4156-b04e-91142a14f6cf"
yPredictes = model_dense.predict(x=xTest)
yPredictes = sc.inverse_transform(yPredictes)
yPredictes
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="Ropf9xE06EpC" outputId="776d8949-7a67-4370-894b-4af506663879"
plt.plot(yTest, 'c-', label='Real')
plt.plot(yPredictes, 'm-', label='Predict')
# plt.plot(data_original, color='red', label='Real')
# plt.plot(range(len(y_train)),yPredicts, color='blue', label='Predict')
plt.title(label='Prediction')
plt.xlabel(xlabel='Time')
plt.ylabel(ylabel='T')
plt.legend()
plt.show()
r2 = r2_score(yTest, yPredictes)
r2_value['Dense'] = r2
print(r2)
# + [markdown] id="58nqhD7H4lcf"
# # <font color=purple>Cnn</font>
# + id="u1-tI1mhINw9"
timestep = cnn_width
# + id="LbDV9Xd23vdA"
xTrainSet = dt_nor[:training_num]
yTrainSet = dt_nor[1:training_num+1]
# + colab={"base_uri": "https://localhost:8080/"} id="jSaV4zYq3vgW" outputId="a45a2092-08a2-433b-de49-370a500935ff"
xTrain = []
for i in range(timestep, training_num):
xTrain.append(xTrainSet[i-timestep : i])
xTrain = np.array(xTrain)
# xTrain = np.squeeze(xTrain)
xTrain = np.reshape(xTrain, (xTrain.shape[0], xTrain.shape[1], 1))
print(xTrain.shape)
yTrain = []
for i in range(timestep, training_num):
yTrain.append(yTrainSet[i])
yTrain = np.array(yTrain)
#yTrain = np.reshape(yTrain, (yTrain.shape[0], 1))
print(yTrain.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="Jddy-dawKYbm" outputId="2ec4e577-527f-4012-c8d0-267cde1e23bd"
conv_width = 3
model_cnn = Sequential()
model_cnn.add(Conv1D(64, kernel_size=(conv_width), input_shape = (xTrain.shape[1],1), activation='relu'))
model_cnn.add(Conv1D(32, kernel_size=(conv_width), activation='relu'))
model_cnn.add(Flatten())
model_cnn.add(Dense(32, activation='relu'))
model_cnn.add(Dense(1))
model_cnn.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="YosJ3W-48aE9" outputId="582b8c95-c080-4a66-c6e8-8c3ffa75d4cc"
model_cnn.compile(optimizer = 'adam',
loss = 'mean_squared_error',
metrics = [tf.metrics.MeanAbsoluteError()])
model_cnn.fit(x = xTrain, y = yTrain, epochs = epoch, batch_size = batch_size)
# + [markdown] id="R1kjq6i77qrp"
# # <font color=purple>Test model's accuracy by r2_score (1200 days)</font>
# + id="2HcWUyDc8aHY"
xTestSet = dt_nor[training_num : 40800-2]
xTestSet = np.array(xTestSet)
yTestSet = dt_nor[training_num+1 : 40800-1]
yTestSet = np.array(yTestSet)
# + colab={"base_uri": "https://localhost:8080/"} id="xIII9cjH8aJh" outputId="0959322e-879b-4580-ceb0-f12d6604da7e"
xTest = []
for i in range(timestep, len(xTestSet)):
xTest.append(xTestSet[i-timestep : i])
xTest = np.array(xTest)
# xTest = np.squeeze(xTest)
yTest = []
for i in range(timestep, len(xTestSet)):
yTest.append(yTestSet[i])
yTest = np.array(yTest)
yTest = sc.inverse_transform(yTest)
len(xTest)
# + colab={"base_uri": "https://localhost:8080/"} id="Hc-KT-zY8aLm" outputId="5c66d751-0241-4873-c7a6-4b7b1339c645"
yPredictes = model_cnn.predict(x=xTest)
yPredictes = sc.inverse_transform(yPredictes)
yPredictes.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="Jf1Jsdpn8aNw" outputId="a944f65e-6eaf-4111-81c8-50f4b4e0aa0b"
plt.plot(yTest, 'c-', label='Real')
plt.plot(yPredictes, 'm-', label='Predict')
# plt.plot(data_original, color='red', label='Real')
# plt.plot(range(len(y_train)),yPredicts, color='blue', label='Predict')
plt.title(label='Prediction')
plt.xlabel(xlabel='Time')
plt.ylabel(ylabel='T')
plt.legend()
plt.show()
r2 = r2_score(yTest, yPredictes)
r2_value['CNN'] = r2
print(r2)
# -
# # <font color=purple>Compare</font>
# + colab={"base_uri": "https://localhost:8080/"} id="JFDYLKRR8aRU" outputId="45f3f16d-6246-4142-c3db-7cba14e0db4f"
r2_value
# + colab={"base_uri": "https://localhost:8080/", "height": 448} id="ICAFX9ZaHzSv" outputId="1dc646a3-574d-4715-b0e5-c098a453a610"
x = np.arange(3)
width = 0.2
val_r2 = r2_value.values()
plt.figure(figsize=(4,7))
plt.ylabel('r2_score [T (degC)]')
plt.bar(x , val_r2, 0.4, label='Test')
plt.xticks(ticks=x, labels=r2_value.keys(), rotation=45)
_ = plt.legend()
# + [markdown] id="IUfXqKYFg-Z3"
# # <font color=purple>Use predict data to predict future</font>
# + [markdown] id="ePrX7u4OVqtK"
# # <font color=purple>Rnn</fnt>
# + id="7eM2Zs3ThDrK"
# Take last 24 hours to predict
Predict_hours = 24
xPred = dt_nor[-Predict_hours-rnn_width:-Predict_hours]
xPred = np.array(xPred)
xPred_in = sc.inverse_transform(xPred)
xPred = np.reshape(xPred, (xPred.shape[1], xPred.shape[0], 1))
yFutureTest = dt_nor[-Predict_hours:]
yFutureTest = np.array(yFutureTest)
yFutureTest = sc.inverse_transform(yFutureTest)
real = []
real = np.append(xPred_in, yFutureTest, axis = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="RQkZ9apafqqm" outputId="63152061-ed59-460d-c405-65d9ee6c621c"
xPred.shape
# + id="Le1hsun3i-qm"
def PredFuture_rnn(xPred):
yPred = model_rnn.predict(x=xPred)
yPred = np.reshape(yPred, (1, 1, 1))
data = np.append(xPred, yPred, axis = 1)
data = data[:, -(rnn_width):, :]
return data
yModelPred = []
for i in range (Predict_hours):
xPred = PredFuture_rnn(xPred)
yModelPred.append(xPred[0][-1])
# + id="6D6FFRIuttNG"
yModelPred = np.array(yModelPred)
yModelPred = sc.inverse_transform(yModelPred)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="qwRIuIrywg6f" outputId="3baff5b5-57e3-4a95-f0b7-a7cbc3fc8e2f"
plt.plot(real, 'y-', label='train')
plt.plot(range(rnn_width, Predict_hours + rnn_width), yFutureTest, 'c-', label='Real')
plt.plot(range(rnn_width, Predict_hours + rnn_width), yModelPred, 'm-', label='Predict')
plt.title(label='Prediction')
plt.xlabel(xlabel='Time')
plt.ylabel(ylabel='T')
plt.legend()
plt.show()
r2_future = {}
r2 = r2_score(yFutureTest, yModelPred)
r2_future['RNN'] = r2
print(r2)
# + [markdown] id="DtQD4DoNbbHp"
# # <font color=purple>Cnn</font>
# + id="l8048wtcbdJG"
# Take last 24 hours to predict
Predict_hours = 24
xPred = dt_nor[-Predict_hours-cnn_width:-Predict_hours]
xPred = np.array(xPred)
xPred_in = sc.inverse_transform(xPred)
# #xPred = np.reshape(xPred, (1,xPred.shape[0],xPred.shape[1]))
xPred = np.reshape(xPred, (xPred.shape[1], xPred.shape[0], 1))
yFutureTest = dt_nor[-Predict_hours:]
yFutureTest = np.array(yFutureTest)
yFutureTest = sc.inverse_transform(yFutureTest)
real = []
real = np.append(xPred_in, yFutureTest, axis = 0)
# + id="SvEEMRkYbdP9"
def PredFuture_cnn(xPred):
yPred = model_cnn.predict(xPred)
yPred = np.reshape(yPred, (1, 1, 1))
data = np.append(xPred, yPred, axis = 1)
data = data[:, -(cnn_width):, :]
return data
yModelPred = []
for i in range (Predict_hours):
xPred = PredFuture_cnn(xPred)
yModelPred.append(xPred[0][-1])
# + id="Yyrt7gXBbdSo"
yModelPred = np.array(yModelPred)
yModelPred = sc.inverse_transform(yModelPred)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="spwbdFjTbdV9" outputId="ea9919b8-01a4-452d-c29e-00a4e4461ecd"
plt.plot(real, 'y-', label='train')
plt.plot(range(cnn_width, Predict_hours + cnn_width), yFutureTest, 'c-', label='Real')
plt.plot(range(cnn_width, Predict_hours + cnn_width), yModelPred, 'm-', label='Predict')
plt.title(label='Prediction')
plt.xlabel(xlabel='Time')
plt.ylabel(ylabel='T')
plt.legend()
plt.show()
r2 = r2_score(yFutureTest, yModelPred)
r2_future['CNN'] = r2
print(r2)
# + [markdown] id="aBzs_D3z4ZRP"
# # <font color=purple>Dense</font>
# + id="qbSXOc1F4bXW"
# Take last 24 hours to predict
Predict_hours = 24
xPred = dt_nor[-Predict_hours-dense_width:-Predict_hours]
xPred = np.array(xPred)
xPred_in = sc.inverse_transform(xPred)
# #xPred = np.reshape(xPred, (1,xPred.shape[0],xPred.shape[1]))
xPred = np.reshape(xPred, (xPred.shape[1], xPred.shape[0], 1))
yFutureTest = dt_nor[-Predict_hours:]
yFutureTest = np.array(yFutureTest)
yFutureTest = sc.inverse_transform(yFutureTest)
real = []
real = np.append(xPred_in, yFutureTest, axis = 0)
# + id="G_ZCE2YO4bZ_"
def PredFuture_dense(xPred):
yPred = model_dense.predict(xPred)
yPred = np.reshape(yPred, (1, 1, 1))
data = np.append(xPred, yPred, axis = 1)
data = data[:, -(dense_width):, :]
return data
yModelPred = []
for i in range (Predict_hours):
xPred = PredFuture_dense(xPred)
yModelPred.append(xPred[0][-1])
# + id="U5J64VO34bdd"
yModelPred = np.array(yModelPred)
yModelPred = sc.inverse_transform(yModelPred)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="a7GCw57B4bf0" outputId="3b5f5452-4af5-447a-ba6f-c79fc15eff2d"
plt.plot(real, 'y-', label='train')
plt.plot(range(dense_width, Predict_hours + dense_width), yFutureTest, 'c-', label='Real')
plt.plot(range(dense_width, Predict_hours + dense_width), yModelPred, 'm-', label='Predict')
plt.title(label='Prediction')
plt.xlabel(xlabel='Time')
plt.ylabel(ylabel='T')
plt.legend()
plt.show()
r2 = r2_score(yFutureTest, yModelPred)
r2_future['Dense'] = r2
print(r2)
# -
# # <font color=purple>Compare</font>
# + colab={"base_uri": "https://localhost:8080/"} id="2BUqUqJd6W4P" outputId="47851857-b854-4fbc-d1ba-b6185053b64f"
r2_future
# + colab={"base_uri": "https://localhost:8080/", "height": 448} id="To-Pb2R_6Y0h" outputId="5ec6b25a-d356-4652-ea82-bf1134716e03"
x = np.arange(3)
width = 0.2
val_r2 = r2_future.values()
plt.figure(figsize=(4,7))
plt.ylabel('r2_score [T (degC)]')
plt.axhline(0, color= 'r')
plt.bar(x , val_r2, 0.4, label='Test')
plt.xticks(ticks=x, labels=r2_future.keys(), rotation=45)
_ = plt.legend()
|
Jena_climate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://cocl.us/pytorch_link_top">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
# </a>
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
# <h1>Test Uniform, Default and He Initialization on MNIST Dataset with Relu Activation</h1>
# <h2>Table of Contents</h2>
# <p>In this lab, you will test the Uniform Initialization, Default Initialization and He Initialization on the MNIST dataset with Relu Activation</p>
#
# <ul>
# <li><a href="#Model">Neural Network Module and Training Function</a></li>
# <li><a href="#Makeup_Data">Make Some Data</a></li>
# <li><a href="#Cost">Define Several Neural Network, Criterion function, Optimizer</a></li>
# <li><a href="#Train">Test Uniform, Default and He Initialization</a></li>
# <li><a href="#Result">Analyze Results</a></li>
# </ul>
# <p>Estimated Time Needed: <strong>25 min</strong></p>
#
# <hr>
# <h2>Preparation</h2>
# We'll need the following libraries:
# +
# Import the libraries we need to use in this lab
# Using the following line code to install the torchvision library
# # !conda install -y torchvision
import torch
import torch.nn as nn
import torchvision.transforms as transforms
import torchvision.datasets as dsets
import torch.nn.functional as F
import matplotlib.pylab as plt
import numpy as np
torch.manual_seed(0)
# -
# <!--Empty Space for separating topics-->
# <h2 id="Model">Neural Network Module and Training Function</h2>
# Define the neural network module or class with He Initialization
# +
# Define the class for neural network model with He Initialization
class Net_He(nn.Module):
# Constructor
def __init__(self, Layers):
super(Net_He, self).__init__()
self.hidden = nn.ModuleList()
for input_size, output_size in zip(Layers, Layers[1:]):
linear = nn.Linear(input_size, output_size)
torch.nn.init.kaiming_uniform_(linear.weight, nonlinearity='relu')
self.hidden.append(linear)
# Prediction
def forward(self, x):
L = len(self.hidden)
for (l, linear_transform) in zip(range(L), self.hidden):
if l < L - 1:
x = F.relu(linear_transform(x))
else:
x = linear_transform(x)
return x
# -
# Define the class or neural network with Uniform Initialization
# +
# Define the class for neural network model with Uniform Initialization
class Net_Uniform(nn.Module):
# Constructor
def __init__(self, Layers):
super(Net_Uniform, self).__init__()
self.hidden = nn.ModuleList()
for input_size, output_size in zip(Layers, Layers[1:]):
linear = nn.Linear(input_size,output_size)
linear.weight.data.uniform_(0, 1)
self.hidden.append(linear)
# Prediction
def forward(self, x):
L = len(self.hidden)
for (l, linear_transform) in zip(range(L), self.hidden):
if l < L - 1:
x = F.relu(linear_transform(x))
else:
x = linear_transform(x)
return x
# -
# Class or Neural Network with PyTorch Default Initialization
# +
# Define the class for neural network model with PyTorch Default Initialization
class Net(nn.Module):
# Constructor
def __init__(self, Layers):
super(Net, self).__init__()
self.hidden = nn.ModuleList()
for input_size, output_size in zip(Layers, Layers[1:]):
linear = nn.Linear(input_size, output_size)
self.hidden.append(linear)
def forward(self, x):
L=len(self.hidden)
for (l, linear_transform) in zip(range(L), self.hidden):
if l < L - 1:
x = F.relu(linear_transform(x))
else:
x = linear_transform(x)
return x
# -
# Define a function to train the model, in this case the function returns a Python dictionary to store the training loss and accuracy on the validation data
# +
# Define function to train model
def train(model, criterion, train_loader, validation_loader, optimizer, epochs = 100):
i = 0
loss_accuracy = {'training_loss': [], 'validation_accuracy': []}
#n_epochs
for epoch in range(epochs):
for i, (x, y) in enumerate(train_loader):
optimizer.zero_grad()
z = model(x.view(-1, 28 * 28))
loss = criterion(z, y)
loss.backward()
optimizer.step()
loss_accuracy['training_loss'].append(loss.data.item())
correct = 0
for x, y in validation_loader:
yhat = model(x.view(-1, 28 * 28))
_, label = torch.max(yhat, 1)
correct += (label == y).sum().item()
accuracy = 100 * (correct / len(validation_dataset))
loss_accuracy['validation_accuracy'].append(accuracy)
print('epoch: '+ str(epoch) +'/'+str(epochs) + ' training_loss: '+ str(loss.data.item()))
return loss_accuracy
# -
# <!--Empty Space for separating topics-->
# <h2 id="Makeup_Data">Make some Data</h2>
# Load the training dataset by setting the parameters <code>train </code> to <code>True</code> and convert it to a tensor by placing a transform object int the argument <code>transform</code>
# +
# Create the training dataset
train_dataset = dsets.MNIST(root='./data', train=True, download=True, transform=transforms.ToTensor())
# -
# Load the testing dataset by setting the parameters train <code>False</code> and convert it to a tensor by placing a transform object int the argument <code>transform</code>
# +
# Create the validation dataset
validation_dataset = dsets.MNIST(root='./data', train=False, download=True, transform=transforms.ToTensor())
# -
# Create the training-data loader and the validation-data loader object
# +
# Create the data loader for training and validation
train_loader = torch.utils.data.DataLoader(dataset=train_dataset, batch_size=2000, shuffle=True)
validation_loader = torch.utils.data.DataLoader(dataset=validation_dataset, batch_size=5000, shuffle=False)
# -
# <!--Empty Space for separating topics-->
# <h2 id="Cost">Define Neural Network, Criterion function, Optimizer and Train the Model</h2>
# Create the criterion function
# +
# Create the criterion function
criterion = nn.CrossEntropyLoss()
# -
# Create a list that contains layer size
# +
# Create the parameters
input_dim = 28 * 28
output_dim = 10
layers = [input_dim, 100, 200, 100, output_dim]
# -
# <!--Empty Space for separating topics-->
# <h2 id="Train">Test PyTorch Default Initialization, Xavier Initialization and Uniform Initialization</h2>
# Train the network using PyTorch Default Initialization
# +
# Train the model with the default initialization
model = Net(layers)
learning_rate = 0.01
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
training_results = train(model, criterion, train_loader,validation_loader, optimizer, epochs=30)
# -
# Train the network using He Initialization function
# +
# Train the model with the He initialization
model_He = Net_He(layers)
optimizer = torch.optim.SGD(model_He.parameters(), lr=learning_rate)
training_results_He = train(model_He, criterion, train_loader, validation_loader, optimizer, epochs=10)
# -
# Train the network using Uniform Initialization function
# +
# Train the model with the Uniform initialization
model_Uniform = Net_Uniform(layers)
optimizer = torch.optim.SGD(model_Uniform.parameters(), lr=learning_rate)
training_results_Uniform = train(model_Uniform, criterion, train_loader, validation_loader, optimizer, epochs=10)
# -
# <!--Empty Space for separating topics-->
# <h2 id="Result">Analyze Results</h2>
# Compare the training loss for each activation
# +
# Plot the loss
plt.plot(training_results_He['training_loss'], label='He')
plt.plot(training_results['training_loss'], label='Default')
plt.plot(training_results_Uniform['training_loss'], label='Uniform')
plt.ylabel('loss')
plt.xlabel('iteration ')
plt.title('training loss iterations')
plt.legend()
# -
# Compare the validation loss for each model
# +
# Plot the accuracy
plt.plot(training_results_He['validation_accuracy'], label='He')
plt.plot(training_results['validation_accuracy'], label='Default')
plt.plot(training_results_Uniform['validation_accuracy'], label='Uniform')
plt.ylabel('validation accuracy')
plt.xlabel('epochs ')
plt.legend()
plt.show()
# -
# <!--Empty Space for separating topics-->
# <a href="http://cocl.us/pytorch_link_bottom">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
# </a>
# <h2>About the Authors:</h2>
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
# Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/"><NAME></a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a"><NAME></a>
# <hr>
# Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
|
IBM_AI_Engineering/Course-4-deep-neural-networks-with-pytorch/Week-5-Deep-Networks/8.3.3.He_Initialization_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
a=open('comments.txt')
b=a.read()
d=b.split()
e={}
for i in d:
if i in e:
d[i] +=1
else:
d[i]=1
print(e)
|
Wordscount in commentstxt.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # Make custom ARIBA database
#
# For the *N. gonorrhoeae* data of the manuscript, we were interested in particular sequences and SNPs. This means using custom reference data as opposed to one of the public reference sets (the use of which is described [in the ARIBA wiki page](https://github.com/sanger-pathogens/ariba/wiki/Task:-getref)). If you just want to use a public database, then use the command "[ariba getref](https://github.com/sanger-pathogens/ariba/wiki/Task:-getref)" instead.
#
#
# We are interested in several reference sequences, some of which are coding seqeunces and others are not, and particular variants in those sequences. The idea is to generate input files to ARIBA that contain all the sequences and variants of interest, using the ARIBA function [aln2meta](https://github.com/sanger-pathogens/ariba/wiki/Task:-aln2meta).
#
# First, let's change to the directory with the reference data.
cd data/Ref/
# ## Using one reference sequence
#
# We start by describing *folP*, but the method is (nearly) the same for all other sequences.
#
# There are several alleles of *folP* and we want to include them all the ARIBA database. We have a SNP of interest R228S in the sequence in the allele called "folP.WHO_F_01537c", which confers resistance to sulphonamides. When we run ARIBA, a particular sample may have a different allele from folP.WHO_F_01537c, but we would still like to know whether it has the SNP R228S. However, the position may not be 228 because of insertion or deletions. So we use a multiple alignment of all the reference alleles, and just supply the SNP to ARIBA in one of the alleles, in this case folP.WHO_F_01537c.
#
# The "aln2meta" function of ARIBA needs two input files: a multiple alignment file of the alleles, and a tab-delimited file of the SNPs of interest. In this case, the SNPs file simply contains one line:
cat aln2meta_input/folP_in.tsv
# There are four columns:
#
# 1. Sequence name
# 2. SNP (an amino acid change at position 228, where R is the wild type and S is the variant)
# 3. A "group name" for this SNP. This is optional and a dot "." means no group name. Putting SNPs into the same group allows ARIBA to report them together later on.
# 4. A description of the SNP. This will appear in ARIBA's output files to save looking up the reason the SNP is of interest.
#
# This file is used together with the mulitple alignment file to generate input files to ARIBA when making the database. This is the command to run:
ariba aln2meta --variant_only aln2meta_input/folP.aln \
aln2meta_input/folP_in.tsv coding aln2meta_output/folP
# A few things to note about the above command:
#
# 1. The option `--variant_only` was used, which affects how ARIBA reports later on when summarizing across all samples. We are only interested in this gene being present if it has a variant that causes resistance.
#
# 2. `aln2meta_input/folP.aln` is the name of the multiple alignment file.
#
# 3. The sequence is "coding", which makes ARIBA treat it as such, in particular it will interpret the variant R228S as an amino acid change at position 228 in the translated amino acid sequence. The input sequence is still in nucleotides, not amino acids.
#
# 4. The command output three files, which can be used as input to the command `ariba prepareref` (see later).
ls aln2meta_output/folP*
# Although we have many more reference sequences of interest to deal with for the complete analysis, for illustrative purposes here we can use the three files `aln2meta_output/folP*` to make an ARIBA reference database:
ariba prepareref -f aln2meta_output/folP.fa \
-m aln2meta_output/folP.tsv --cdhit_clusters \
aln2meta_output/folP.cluster test.aribadb
# This made an ARIBA database of just those sequences and the SNP R228S in a new directory called `test.aribadb`. Let's check that it was made:
ls test.aribadb
# You do not need to worry about the contents of the new directory `test.aribadb`, just know that it can be used as input to run ARIBA. However, this was for just one of the many reference sequences of interest, so we will delete it.
rm -r test.aribadb
# ## Using all reference sequences
#
# We need to deal with each of the reference sequences in turn by running `ariba aln2meta` on each, like in the above example with *folP*. The only difference is that some of them are non-coding sequences, which means that the command must have `noncoding` instead of `coding`. For example:
ariba aln2meta --variant_only aln2meta_input/16S.aln \
aln2meta_input/16S_in.tsv noncoding aln2meta_output/16S
# There are 10 coding sequences and two non-coding sequences. Instead of writing 12 commands, we will use two 'for loops'. First, the coding sequences:
for x in folP gyrA mtrR parC parE penA ponA porB1b rpoB rpsJ
do
ariba aln2meta --variant_only aln2meta_input/$x.aln \
aln2meta_input/$x\_in.tsv coding aln2meta_output/$x
done
# And now the two non-coding sequences:
for x in 16S 23S
do
ariba aln2meta --variant_only aln2meta_input/$x.aln \
aln2meta_input/$x\_in.tsv noncoding aln2meta_output/$x
done
# This has generated three files for each sequence. We will combine these to make input files for running `ariba prepareref`.
cat aln2meta_output/*.fa presence_absence/*.fa > Ngo_ARIBA.fa
cat aln2meta_output/*.tsv presence_absence/presence_absence.tsv \
> Ngo_ARIBA.tsv
cat aln2meta_output/*.cluster \
presence_absence/presence_absence.clusters \
> Ngo_ARIBA.clusters
# Finally, we have the three input files needed to make a single ARIBA database that has information on all the sequences and SNPs of interest. In case the directory is already there, we delete it first, then generate the database:
rm -rf Ngo_ARIBAdb
ariba prepareref -f Ngo_ARIBA.fa -m Ngo_ARIBA.tsv \
--cdhit_clusters Ngo_ARIBA.clusters Ngo_ARIBAdb
# We now have a directory `Ngo_ARIBAdb` that can be used as the reference database when running `ariba run` on each sample.
#
# Now move on to the next part of the tutorial where we [run ARIBA using the custom reference data](run_ariba.ipynb), or [return to the index](index.ipynb).
|
ARIBA/make_custom_db.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring the *RMS Titanic* sinking in Neo4j
#
# The Titanic dataset is very well known among the data science and analytics community. This notebook outlines the initial preprocessing steps in the pipeline to prepare the data for import into a property graph database such as Neo4j. Once the data is in Neo4j the relationships between entities in the data, such as passengers, lifeboats, destinations, etc. can be easily analyzed and visualized. Graph data science can offer an understanding of context within data in a way that tabular data does not do easily.
#
# ## Setting up resources
# 1) Configure environment using Anaconda or virtualenv.
#
# 2) Deploy a local Neo4j Docker instance:
# ```bash
# # cd neo4j-titanic \
# # && docker build -t neo4j-titanic:neo4j_db ./neo4j \
# # && docker run --name neo4j_db -d -p 7474:7474 -p 7473:7473 -p 7687:7687 \
# # -v $PWD/data/interim:/var/lib/neo4j/import neo4j-titanic:neo4j_db
# ```
#
# 3) (optional) If geoparsing is desired, the Mordecai Python package requires an ElasticSearch service to be running on the correct port with the correct index. This can be set up from the command line:
# ```bash
# docker pull elasticsearch:5.5.2 \
# && wget https://s3.amazonaws.com/ahalterman-geo/geonames_index.tar.gz --output-file=wget_log.txt \
# && tar -xzf geonames_index.tar.gz \
# && docker run -d -p 127.0.0.1:9200:9200 -v "$(pwd)/es/geonames_index/:/usr/share/elasticsearch/data" elasticsearch:5.5.2
# ```
#
# ## Running the pipeline
# The ```src.preprocess``` module contains functions for cleaning, feature engineering and batch geoparsing with NLP analysis.
# ## Import Data
# +
import sys
# Set paths for modules
sys.path[0] = '../'
import pandas as pd
pd.options.display.max_rows = None # display all rows
pd.options.display.max_columns = None # display all columns
import py2neo
# import pipeline
from src.preprocess import clean_data, remap_abbrev
from src.eda import get_snapshot
# +
# Define paths --> move to .env
URL = 'https://query.data.world/s/xjk6hp7t7w3553bfpkfshr2bjd67a4'
RAW_PATH = sys.path[0] + 'data/raw/titanic.csv'
INTERIM_PATH = sys.path[0] + 'data/interim/titanic_clean.csv'
GEOPARSED_PATH = sys.path[0] + 'data/processed/titanic_final.csv'
# import
data = pd.read_csv(URL)
# -
# ## Preprocessing
#
# For the purpose of creating a property graph it is useful to correct errors, fill NaN values with useful information and update values to improve readability. Creating new feature columns can expedite node creation when the graph is created. Preprocessing makes use of the ```clean_data``` function.
#
# ### Cleaning Steps:
# * ***embarked*** - Fix NaN & Replace letters with place names for readability
# * ***home.dest*** - Fill NaN with 'Unspecified Destination'; Replace abbreviations with names.
#
# ### Feature Engineering Steps:
# * ***family.size*** - Combines *sibsb* and *parch* for total size of family including passenger
# * ***surname*** - Extracts *surname* from *name*. This will make it much easier to define family relationships.
# * ***deck*** - Extracts the *deck* from *cabin* in order to make this into a node.
# Clean data and save
data = clean_data(data)
data.to_csv(INTERIM_PATH, index=False)
data.head()
get_snapshot(data)
# ## Geoparsing
#
# The goal of data analysis is to extract as much useful information as possible. In this case it would be useful to create nodes from the destination countries of passengers. In order to do this we have to extract that data from the unstructured text data in *home.dest* using Natural Language Processing (NLP).
#
# The `geoparse_data` function uses the Mordecai package to extract geopolitical entities from unstructured text. Applying this to the *home.dest* column returns the country ISO values, and the Pycountry package is used to convert these into country names for our *home.country* nodes. These steps can be viewed in detail in the *0.1-process-data* notebook.
# Replace home.dest abbreviations with full names for NLP step
data['home.dest'] = remap_abbrev(data['home.dest'])
# +
# Run geoparser. This step can take some time so prepare to wait.
#data = geoparse_data(data[:5])
# +
# Save to processed data folder. Careful not to overwrite accidentally.
#data.to_csv(GEOPARSED_PATH,index=False)
# -
# Inspect
data.head()
# ## Load to Neo4j
#
# There are a number of options for getting data into Neo4j depending on the size of the import. The simplest way to load data into a container with Neo4j is through py2neo or a shell command to run a query that reads a preprocessed CSV, although this may not be the quickest option. The fastest method is use the `neo4j-admin import` tool which works especially well with large datasets. The drawback is that it requires specially formatted CSV files which means additional preprocessing steps.
# +
# Method 1: use shell to load
# #!cat neo4j/create_db.cyp | docker exec --interactive neo4j_db bin/cypher-shell -u neo4j -p test
# +
# Method 2: use py2neo
file = sys.path[0] + '/neo4j/create_db.cyp'
with open(file, 'r') as f:
query = f.read()
# Connect to local running neo4j instance
graph = py2neo.Graph(host='127.0.0.1', password='<PASSWORD>')
# Split queries into individual statements
queries = query.split(';')
# Run each query
for cypher in queries[:-1]:
graph.run(cypher)
# -
|
notebooks/0.2-pipeline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy.stats import uniform
import matplotlib
import seaborn
import matplotlib.pyplot as plt
# -
seaborn.set_style(fig)
# ## STANDARD UNIFORM DISTRIBUTION (a = 0, b =1)
# ### Instantiating a Random continuous variable following a uniform distribution
U_rv = uniform()
# Looking at 10 variates (sample draws) from this distribution.
# Setting a random state for reproducibility
x = U_rv.rvs(size=100000, random_state=42)
x[0:10]
# ### Frequency Distribution of uniformally distributed random variables.
# +
plt.figure(figsize=(20,8))
seaborn.distplot(x, kde=False, bins = 30)
plt.xlabel('Possible continous outcomes of the random variable')
plt.ylabel('Count of Random variates')
plt.title('Frequency distributon of a random variable following a uniform distribution')
# -
U_rv.mean()
U_rv.median()
# <br/>
#
# ### Probability Distribution Function (PDF)
# +
plt.figure(figsize=(20,8))
#fig, ax = plt.subplots(1, 1)
plt.hist(x, density=True)
plt.plot(x, uniform.pdf(x),'red', lw=5, alpha=0.2, label='uniform PDF')
plt.xlabel('Numerical range of the continuous random variable')
plt.ylabel('PDF')
plt.title('Probability density function values (can be > 1) for a uniformly distributed continuous random variable')
plt.legend(loc='upper right', frameon=False)
plt.show()
# -
# <br/>
#
# ### Cumulative Distribution Function (CDF)
# +
plt.figure(figsize=(20,8))
#fig, ax = plt.subplots(1, 1)
plt.hist(x, density=True)
plt.plot(x, uniform.cdf(x),'g', lw=10, alpha=0.5, label='uniform CDF')
plt.xlabel('Numerical range of the continuous random variable')
plt.ylabel('PDF')
plt.title('Probability density function values (can be > 1) for a uniformly distributed continuous random variable')
plt.legend(loc='upper right', frameon=False)
plt.show()
# -
# ---
#
# What is the 20th percentile of this uniform distribution ? i.e. What is the value of the variable such that, the AUC to the left of which is 0.2.
U_rv.ppf(0.2)
# 50th percentile. -- Its the same as rv.median as seen above.
U_rv.ppf(0.5)
# ### Upper and lower bounds (support) of the uniform distribution
uniform.a
uniform.b
# <br/>
# <br/>
#
# ## Using a different range for a fixed distribution
U_rv_fixed = uniform(1, scale=4)
x = U_rv_fixed.rvs(size=100000, random_state=42)
x[0:10]
# ### Frequency Distribution of uniformally distributed random variables.
# +
plt.figure(figsize=(20,8))
seaborn.distplot(x, kde=False, bins = 30)
plt.xlabel('Possible continous outcomes of the random variable')
plt.ylabel('Count of Random variates')
plt.title('Frequency distributon of a random variable following a uniform distribution')
# -
U_rv_fixed.mean()
U_rv_fixed.median()
# See how `loc` and `scale` change the `mean` and `median` of the U_rv (standard u distribution) and U_rv_fixed.
|
Data_Science_Utils/Uniform_Distribution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import torch
import UnarySim
from UnarySim.sw.stream.shuffle_int import SkewedSyncInt
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
ss = SkewedSyncInt(depth=3).to(device)
a = torch.tensor([[0, 0]]).type(torch.float).to(device)
b = torch.tensor([[1, 1]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[4, 4]]).type(torch.float).to(device)
b = torch.tensor([[0, 0]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[4, 4]]).type(torch.float).to(device)
b = torch.tensor([[1, 1]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[0, 0]]).type(torch.float).to(device)
b = torch.tensor([[0, 0]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[4, 4]]).type(torch.float).to(device)
b = torch.tensor([[1, 1]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[4, 4]]).type(torch.float).to(device)
b = torch.tensor([[0, 0]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[4, 4]]).type(torch.float).to(device)
b = torch.tensor([[0, 0]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[4, 4]]).type(torch.float).to(device)
b = torch.tensor([[0, 0]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
a = torch.tensor([[2, 2]]).type(torch.float).to(device)
b = torch.tensor([[0, 0]]).type(torch.float).to(device)
out_a, out_b = ss(a,b)
print(out_a)
print(out_b)
print(ss.cnt)
|
sw/test/stream/test_stream_shuffle_skewedsyncint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="0DHmilV6qlC-"
# ## Index ##
# - Einleitung
# - Einfuehrung des SEIR Models
# - Erweiterung des SEIR Models: Mathematische Erklärung
# - Das SEIIRHQ-Modell
# - Das SSSEIIRHQ-Modell
# - Umsetzung des Modells in Python
# - Vorlage (Beschreibung der Vorlage, Euler etc.)
# - Anpassung der Vorlage (code)
# - Kommentierung der Ergebnisse
# - Vergleich mit reellen Daten
# - Quellen
#
# + [markdown] id="Ed-sUGmf_EER"
# ## Einleitung
#
# In diesem Jupyter-Notebook beschäftigen wir uns mit dem sogenannten SEIR-Modell. Vorrangig geht es hierbei um die algorithmische Umsetzung dieses Modells in Python, unter besonderer Berücksichtigung von Python-Erweiterungen, die im Bereich wissenschaftlicher Programmierung genutzt werden.
# Zunächst soll kurz vorgestellt werden, worum es sich bei SEIR handelt. Anschließend werden Code-Beispiele präsentiert, die die Umsetzung des SEIR-Algorithmus in Python verdeutlichen sollen. Zum Abschluss werden eigene Erweiterungen für das SEIR-Modell präsentiert, die eine bessere Abbildung des Pandemiegeschehens durch Maßnahmen wie Impfung, Qurantäne-Anordnungen etc.leisten sollen. Teile des Codes wurden von https://github.com/m-schmidt-math-opt/covid-19-extended-seir-model übernommen und überarbeitet. Die theoretische Grundlage der Überarbeitung stellt zudem das Paper von He et. al. 2020 (https://www.nature.com/articles/s41598-021-83540-2) dar. Aus diesen Paper wurden die Ideen für die Erweiterungen des SEIR Modells bezogen und dann in eigene Differenzialgleichungen und Python-Code überführt.
#
#
# + [markdown] id="XIPBwqfEsSUK"
#
# # Einführung SEIR Modell
#
# Das sogenannte SEIR-Modell stellt eine Möglichkeit dar, Pandemie-Verläufe innerhalb von Gesellschaften, wie beispielsweise der deutschen Bevölkerung, zu modellieren und wurde auch für den Verlauf der aktuellen Covid-19-Pandemie benutzt, um Vorhersagen über die Entwicklung und den weiteren Verlauf sowie die Wirksamkeit von Gegenmaßnahmen treffen zu können.
#
# Hierbei steht die Abkürzung SEIR für:
#
# **S**usceptible (Gefährdete, Nicht-Infizierte)
#
# **E**xposed (Infizierte, aber selbst noch nicht infektiös)
#
# **I**nfectious (Infektiöse)
#
# **R**ecovered (Genesene, nicht mehr infektiös)
#
#
# Die gesamte Bevölkerung einer Gesellschaft lässt sich in eine dieser Gruppen einteilen und die Summe S+E+I+R=N entspricht dabei der Gesamtzahl N der Individuen ebendieser. Mithilfe von vier Differenzialgleichungen lässt sich die Zunahme und Abnahme innerhalb dieser vier Gruppen abbilden. Das Lösen besagter Differenzialgleichungen erfolgt in unseren Beispielen sowie in den in der Literatur gefundenen Code-Sampeln mithilfe des expliziten Euler-Vefahrens, einem simplen Algorithmus zur numerischen Lösung von Anfangswertproblemen.
#
#
# Die unten aufgeführten Formeln entsprechen jeweils der Änderungsrate , also der Ableitung, der vier Kategorien (**S, E, I, R**) pro Zeitschritt.
#
# Von der Gruppe der anfälligen Personen, **Kategorie S**, wird die Zahl der Personen, die sich innerhalb der Zeiteinheit durchschnittlich aufgrund ihrer Kontakte (entspricht Parameter **β**) infizieren, subtrahiert.
#
# Diese von der Gruppe S abgezogenen, also infizierten Personen werden im gleichen Schritt der **Kategorie E**, was der Zahl der infizierten, aber noch nicht infektiösen Zahl an Menschen betrifft, zugeschlagen, wobei wiederum diejenigen, die gerade infektiös geworden sind (abhängig vom Parameter **θ**), von dieser Gruppe abgezogen werden.
#
# Besagte infektiös Infizierte werden nun der **Kategorie I** zugeschlagen, von der in einem weiteren Schritt alle Genesenen (ermittelt durch Parameter **γ**) abgezogen werden.
#
# Die Genesenen landen dauerhaft in **Kategorie R**.
#
# Die Parameter $\beta$, $\theta$ und $\gamma$ geben bei allen Übergängen jeweil an, wie viele Personen von einer Kategorie im nächsten Zeitschritt durchschnittlich in eine andere Kategorie übergehen. Die genauen Werte für diese Parameter müssen empirisch durch Fachwissenschaftler ermittelt werden und unterliegen einer gewissen Unsicherheit. Allerdings lässt sich jeder Wert innerhalb unserer Algorithmen beliebig anpassen.
#
# ##Formeln
#
# $\dot S = - \frac{1}{N} \beta S I$
#
# $\dot E = \frac{1}{N} \beta S I - \theta E$
#
# $\dot I = \theta E - \gamma I$
#
# $\dot R = \gamma I $
#
# wobei:
#
# |Variable|Beschreibug|
# |--------|-----------|
# |S(t)|Anteil der Anfälligen, engl. susceptible. Noch nicht infiziert.|
# |E(t)|Anteil der Exponierten, engl. exposed. Infiziert, aber noch nicht infektiös.|
# |I(t)|Anteil der Infektiösen, engl. infectious.|
# |R(t)|Anteil der Erholten, engl. recovered oder resistant. Bzw. verstorben oder nach Symptomen in Quarantäne.|
#
#
# und
#
# |Parameter|Beschreibung|
# |---------|------------|
# |$\beta$|Transmissionsrate. Der Kehrwert ist die mittlere Zeit zwischen Kontakten mit Übertragung.|
# |$\gamma$|Erholungsrate. Der Kehrwert ist die mittlere infektiöse Zeit. |
# |$\theta$|Übergangsrate. Der Kehrwert ist die mittlere Latenzzeit.|
#
# + id="ng-82pCjcK1N"
from numpy import array as vector
import numpy as np
class SEIR_Model:
"""
SEIR Basismodell: Die Einfachste Form des SEIR Modells als Baseline, ohne weitere Differenzierung
der Gruppen
"""
def __init__(self, packed_parameters):
if test: print("Running in test mode.")
print("Instantiate the SEIR model ...")
self.n = packed_parameters[0]
self.beta = packed_parameters[1]
self.theta = packed_parameters[2]
self.gamma = packed_parameters[3]
self.x0 = packed_parameters[4]
self.numerical_tolerance_fine = 1e-2 # todo
self.numerical_tolerance_coarse = 1e-2
def eval_rhs(self, x_t):
s, e, i, r = x_t
# Sanity checks
if test: assert(abs(sum(x_t) - sum(self.x0)) < self.numerical_tolerance_coarse)
x_t_1 = vector([
-self.beta*(s/self.n)*i,
self.beta*(s/self.n)*i - self.theta*e,
self.theta*e - self.gamma*i,
self.gamma*i
])
# Sanity checks
if test:
assert(abs(sum(x_t_1)) < self.numerical_tolerance_fine)
return x_t_1
# class SEIR_Model
# + [markdown] id="owlAL3Knf_JO"
# ## Erweiterungen des SEIR-Modells
#
# Das SEIR-Modell stellt eine erste Annäherung an die Entwicklung des Geschehens einer Pandemie wie Covid-19 dar. Die drei Parameter (β, γ und θ) können dabei beliebig modifiziert werden, je nach dem, welche Werte dafür von wissenschaftlicher Seite präferiert werden. Doch das Modell stößt schnell an seine Grenzen und wird der Komplexität einer Pandemie nicht vollständig gerecht. Es ergeben sich folgende Beschränkungen:
#
# - Es wird nicht zwischen den Parametern verschiedener sozialer Gruppen unterschieden.
# - Die Möglichkeit einer Impfung fließt nicht in die Berechnungen mit ein.
# - Genese gelten dauerhaft als immun gegen die Krankheit.
# - Die Möglichkeit, gefährdete oder infizierte Personen in Quarantäne zu schicken, wird nicht berücksichtigt.
# - Der Grad der Hospitalisierung, d.h. der Anteil der Menschen mit einer ernstzunehmenden Erkrankung, findet keine Beachtung
#
# Im Folgenden soll nun das SEIR-Modell um einige Zusatzannahmen erweitert werden und algorithmisch in Python umgesetzt werden. Die Anregungen für die Erweiterung des Modells übernehmen wir aus: (https://link.springer.com/content/pdf/10.1007/s11071-020-05743-y.pdf). Die algorithmische Umsetzung in Python erfolgt durch uns. Es ergeben sich dann die unten aufgeführten (modifizierten und komplexeren) Differenzialgleichungen, die wiederum numerisch, z. B. mit dem Euler-Verfahren, gelöst werden können.
# + [markdown] id="qAvcLKEac4JQ"
# ## Das SEIIRHQ Modell
#
# Im SEIR-Modell wird die Anzahl der hospitalisiserter Patienten berechnet. Außerdem wir die Einführung von Quarantänemaßnahmen berücksichtig. Es wird daher unterschieden zwischen Personen mit "Intervention" (Quarantäne/Hospitalisierung) sowie Patienten ohne Intervention. Diese Patienten nehmen weiter am gesellschaftlichen Leben teil, obwohl sie krank und ansteckend sind, und stecken daher andere Personen mit erhöhter Wahrscheinlichkeit an.
#
# Weiterhin wird die Tatsache berücksichtigt, dass Patienten, die bereits angesteckt waren und wieder genesen sind ("recovered"), sich ein weiteres Mal anstecken können. Diese Übergangsrate wird mit dem Parameter **$\alpha$** simuliert
#
#
# ##Formeln
#
#
# $\dot S = - \frac{S}{N} (\beta_1 I_1 + \beta_2 I_2 + \chi E) + \rho_1 Q - \rho_2 S + \alpha R$
#
# $\dot E = \frac{S}{N} (\beta_1 I_1 + \beta_2 I_2 + \chi E) - \theta_1 E - \theta_2 E$
#
# $\dot I_1 = \theta_1 E - \gamma_1 I_1$
#
# $\dot I_2 = \theta_2 E - \gamma_2 I_2 - \phi I_2 + \lambda (\Lambda + Q)$
#
# $\dot R = \gamma_1 I_1 + \gamma_2 I_2 + \psi H - \alpha R$
#
# $\dot H = \phi I_2 - \psi H$
#
# $\dot Q = \Lambda + \rho_2 S - \lambda (\Lambda + Q) - \rho_1 Q$
#
#
# wobei:
#
# |Variable|Beschreibug|
# |--------|-----------|
# |$S$|Anfällige|
# |E|Exponierte|
# |$I_1$|Infizierte ohne Behandlung|
# |$I_2$|Infizierte mit Behandlung|
# |R|Genesen|
# |H|Hospitalisiert|
# |Q|Unter Quarantäne gestellt|
#
# und
#
# |Parameter|Beschreibung|
# |---------|------------|
# |$\alpha$|Vorübergehende Immunitätsrate|
# |$\beta_1, \beta_2$|Die Kontakt- und Infektionsrate der Übertragung pro Kontakt mit einer infizierten Klasse|
# |$\gamma_1, \gamma_2$|Genesungsrate von symptomatischen infizierten Personen zu genesenen|
# |$\chi$|Wahrscheinlichkeit der Übertragung pro Kontakt durch exponierte Personen bei ungeimpften Personen|
# |$\lambda$|Verhältnis zwischen der unter Quarantäne gestellten Gruppe und der genesenen Gruppe|
# |$\Lambda$|Externer Beitrag aus dem Ausland|
# |$\rho_1, \rho_2$|Übergangsrate von den unter Quarantäne gestellt exponierten Personen zwischen der unter Quarantäne gestellten infizierten Gruppe und der Allgemeinheit|
# |$\theta_1, \theta_2$|Übergangsrate von exponierten Personen in die Klasse der Infizierten|
# |$\phi$|Anteil der infektiösen Personen mit Symptomen, die ins Krankenhaus eingeliefert werden|
# |$\psi$|Genesungsrate der unter Quarantäne stehenden infizierten Personen|
#
# + id="7mjqZJ_knBOe"
from numpy import array as vector
import numpy as np
import random
class SEIIRHQ_Model:
"""
Anpassung des SEIR Modells. Zusätzlich zum SEIR Modell wird hier die Anzahl hospitalisierter Patienten berücksichtig (H). Außerdem werden
die Infizierten Personen (I) nach Behandlungsbedürftigen Patienten und nicht behandlungsbedürftigen Patienten unterteilt
"""
def __init__(self, packed_parameters):
if test: print("Running in test mode.")
print("Instantiate the SEIIRHQ model ...")
self.n = packed_parameters[0] # total of individuals
self.beta_no_intv = packed_parameters[1] # The contact and infection rate of transmission per contact from the infected without intervention class
self.beta_intv = packed_parameters[2] # The contact and infection rate of transmission per contact from the infected with intervention class
self.theta_no_intv = packed_parameters[3] # Transition rate of exposed individuals to the infected without intervention class
self.theta_intv = packed_parameters[4] # Transition rate of exposed individuals to the infected with intervention class
self.gamma_no_intv = packed_parameters[5] # Recovery rate of symptomatic infected without intervention to recovered
self.gamma_intv = packed_parameters[6] # Recovery rate of symptomatic infected with intervention to recovered
self.alpha = packed_parameters[7] # Temporary immunity rate
self.rho_in = packed_parameters[8] # Transition rate between the subsceptible class and quarantined class
self.rho_out = packed_parameters[9] # Transition rate between the quarantined class and the susceptible class
self.chi = packed_parameters[10] # Probability of transmission per contact from exposed individuals
self.phi = packed_parameters[11] # Rate of infectious with symptoms to hospitalized
self.lbda = packed_parameters[12] # Rate of the quarantined class to the infectious class with intervention
self.psi = packed_parameters[13] # Recovered rate of hospitalized / quarantined infected individuals
self.x0 = packed_parameters[14] # seiirhq at time 0
self.total_immigrants=0
self.i=0
self.numerical_tolerance_fine = 10 #
self.numerical_tolerance_coarse = 1e-2
def eval_rhs(self, x_t):
s, e, i_no_intv, i_intv, r, h, q = x_t
immigrants=random.uniform(1000, 10000)
self.total_immigrants=self.total_immigrants+immigrants
factor = (s / self.n) * (self.beta_no_intv * i_no_intv + self.beta_intv * i_intv + self.chi * e)
# [s, e, i_no_intv, i_intv, r, h, q]
x_t_1 = vector([
-factor + self.rho_out*q - self.rho_in*s + self.alpha*r, # S
factor - self.theta_no_intv*e - self.theta_intv*e, # E
self.theta_no_intv*e - self.gamma_no_intv*i_no_intv, # I without intervention
self.theta_intv*e - self.gamma_intv*i_intv - self.phi*i_intv + self.lbda*(immigrants + q), # I with intervention
self.gamma_no_intv*i_no_intv + self.gamma_intv*i_intv + self.psi*h - self.alpha*r, # R
self.phi*i_intv-self.psi*h, # H
immigrants + self.rho_in*s - self.rho_out*q - self.lbda*q # Q
])
# Sanity checks
if test: assert(abs(sum(x_t_1)-(immigrants+(self.lbda*(immigrants)))) < self.numerical_tolerance_fine)
return x_t_1
# class SEIIRHQ_Model
# + [markdown] id="GvBuivlCAJ8Q"
#
#
# ```
# # Als Code formatiert
# ```
#
#
# ## Das SSSEIIRHQ-Modell
# Das SEIIRHQ-Modell wird nun um weitere Kategorien erweitert. Der Gruppe der anfälligen Personen wird nun dreigeteilt: zum einen wird zwischen ungeimpften (**S1**) und geimpften Erwachsenen (**S2**) unterschieden, zum anderen wird die Kategorie 'Kinder' (**S3**) hinzugefügt. Diese neue Unterteilung macht Sinn, da nun verschiedenen sozialen Gruppen unterschiedliche Parameter zugewiesen werden können. Die Zahl und die Komplexität der benutzten Differenzialgleichungen nimmt dadurch zu. Die hierzu verwendeten Formeln sind hier aufgeführt und sollen durch das Schaubild darunter verdeutlicht werden.
#
# ##Formeln
#
#
# $\dot S_1 = - \frac{S_1}{N} (\beta_{1_{novaccined}} I_1 + \beta_{2_{novaccined}} I_2 + \chi_{novaccined} E) + \rho_1 Q - \rho_2 S_1 + \epsilon S_2 - \epsilon S_1 + \alpha (1 - \eta) R $
#
# $\dot S_2 = - \frac{S_2}{N} (\beta_{1_{vaccined}} I_1 + \beta_{2_{vaccined}} I_2 + \chi_{vaccined} E) + \rho_1 Q - \rho_2 S_2 - \epsilon S_2 + \epsilon S_1 $
#
# $\dot S_3 = - \frac{S_3}{N} (\beta_{1_{children}} I_1 + \beta_{2_{children}} I_2 + \chi_{children} E) + \rho_1 Q - \rho_2 S_3 + \alpha \eta R$
#
#
# $\dot E = \frac{(S_1+S_2+S_3)}{N} ((\beta_{1_{novaccined}} + \beta_{1_{vaccined}} + \beta_{1_{children}}) I_1 + (\beta_{2_{novaccined}} + \beta_{2_{vaccined}} + \beta_{2_{children}}) I_2 + (\chi_{novaccined} + \chi_{vaccined} + \chi_{children}) E) - \theta_1 E - \theta_2 E$
#
# $\dot I_1 = \theta_1 E - \gamma_1 I_1$
#
# $\dot I_2 = \theta_2 E - \gamma_2 I_2 - \phi I_2 + \lambda (\Lambda + Q)$
#
# $\dot R = \gamma_1 I_1 + \gamma_2 I_2 + \psi H - \alpha R$
#
# $\dot H = \phi I_2 - \psi H$
#
# $\dot Q = \rho_2 (S_1 + S_2 + S_3) - \lambda Q - \rho_1 Q$
#
#
# wobei:
#
# |Variable|Beschreibug|
# |--------|-----------|
# |$S_1$|Anfällige Klasse der ungeimpften Erwachsenen|
# |$S_2$|Anfällige Klasse der geimpften Erwachsenen|
# |$S_3$|Anfällige Klasse der ungeimpften Kindern|
# |E|Exponierte|
# |$I_1$|Infizierte ohne Behandlung|
# |$I_2$|Infizierte mit Behandlung|
# |R|Genesen|
# |H|Hospitalisiert|
# |Q|Unter Quarantäne gestellt|
#
# und
#
# |Parameter|Beschreibung|
# |---------|------------|
# |$\alpha$|Vorübergehende Immunitätsrate|
# |$\beta_1, \beta_2$|Die Kontakt- und Infektionsrate der Übertragung pro Kontakt mit einer infizierten Klasse|
# |$\gamma_1, \gamma_2$|Genesungsrate von symptomatischen infizierten Personen zu genesenen|
# |$\chi$|Wahrscheinlichkeit der Übertragung pro Kontakt durch exponierte Personen bei ungeimpften Personen|
# |$\lambda$|Verhältnis zwischen der unter Quarantäne gestellten Gruppe und der genesenen Gruppe|
# |$\rho_1, \rho_2$|Übergangsrate von den unter Quarantäne gestellt exponierten Personen zwischen der unter Quarantäne gestellten infizierten Gruppe und der Allgemeinheit|
# |$\theta_1, \theta_2$|Übergangsrate von exponierten Personen in die Klasse der Infizierten|
# |$\phi$|Anteil der infektiösen Personen mit Symptomen, die ins Krankenhaus eingeliefert werden|
# |$\psi$|Genesungsrate der unter Quarantäne stehenden infizierten Personen|
# |$\epsilon$|Impfungsrate|
# |$\eta$|Prozent von Kindern insgesamt|
#
# + colab={"base_uri": "https://localhost:8080/", "height": 743} id="11Knw1FffBL6" outputId="3d904cfd-5c74-423d-8581-9f580563a2ba"
from IPython.display import Image
Image('./images/COVID_Math_Model.png')
# + id="CkqjeOUORwQl"
from numpy import array as vector
import numpy as np
class SSSEIIRHQ_Model:
"""
Die Klasse SSSEIIRHQ_Modell repräsentiert das SSSEIIRHQModell innerhalb des Programms.
In ihr sind alle Formeln(Differenzialgleichungen) notiert, die wichtig sind für dieses Modell
und die die Übergänge zwischen den Kategorien beschreiben.
Sie wird der Klasse Explizit_Euler übergeben für die weitere Berechnung.
"""
#Im Konstruktor werden die gepackten Werte für die Berechnungsparamter (alpha, beta, gamma etc.) sowie die jeweiligen Gruppengrößen
#für die einzelnen Kategorien übergeben und entsprechenden Klassen-Variablen zugeordnet.
def __init__(self, packed_parameters):
print("Instantiate the SSSEIIRHQ model ...")
if test: print("Running in test mode.")
self.n = packed_parameters[0] # total of individuals
self.beta_no_intv = packed_parameters[1] # The contact and infection rate of transmission per contact from the infected without intervention class
self.beta_intv = packed_parameters[2] # The contact and infection rate of transmission per contact from the infected with intervention class
self.theta_no_intv = packed_parameters[3] # Transition rate of exposed individuals to the infected without intervention class
self.theta_intv = packed_parameters[4] # Transition rate of exposed individuals to the infected with intervention class
self.gamma_no_intv = packed_parameters[5] # Recovery rate of symptomatic infected without intervention to recovered
self.gamma_intv = packed_parameters[6] # Recovery rate of symptomatic infected with intervention to recovered
self.alpha = packed_parameters[7] # Temporary immunity rate
self.rho_in = packed_parameters[8] # Transition rate between the subsceptible class and quarantined class
self.rho_out = packed_parameters[9] # Transition rate between the quarantined class and the susceptible class
self.chi = packed_parameters[10] # Probability of transmission per contact from exposed individuals
self.phi = packed_parameters[11] # Rate of infectious with symptoms to hospitalized
self.lbda = packed_parameters[12] # Rate of the quarantined class to the infectious class with intervention
self.psi = packed_parameters[13] # Recovered rate of hospitalized / quarantined infected individuals
self.epsilon = packed_parameters[14] # Impfungsrate
self.eta = packed_parameters[15] # Prozent von Kindern
self.x0 = packed_parameters[16] # ssseiirhq at time 0
self.numerical_tolerance_fine = 1e-2 # todo
self.numerical_tolerance_coarse = 1e-2
#Die Werte für die einzelnen Parameter und die Gruppengröße der Kategorien werden in Form eines Vektors übergeben.
#Die Werte für die Kategorien werden entsprechend der hinterlegten Formeln aktualisiert und in Form eines Vektors zurückgegeben.
def eval_rhs(self, x_t):
s_no_v, s_v, s_ch, e, i_no_intv, i_intv, r, h, q = x_t
# Sanity checks
if test: assert(abs(sum(x_t) - sum(self.x0)) < self.numerical_tolerance_coarse)
factor_no_v = (s_no_v / self.n) * (self.beta_no_intv[0] * i_no_intv + self.beta_intv[0] * i_intv + self.chi[0] * e)
factor_v = (s_v / self.n) * (self.beta_no_intv[1] * i_no_intv + self.beta_intv[1] * i_intv + self.chi[1] * e)
factor_ch = (s_ch / self.n) * (self.beta_no_intv[2] * i_no_intv + self.beta_intv[2] * i_intv + self.chi[2] * e)
# [s_no_v, s_v, s_ch, e, i_no_intv, i_intv, r, h, q]
x_t_1 = vector([
-factor_no_v + self.rho_out*s_no_v/(s_no_v+s_v+s_ch)*q - self.rho_in*s_no_v + (self.alpha*r*(1.00 - self.eta)) - self.epsilon*s_no_v + self.epsilon*s_v, # S not vaccined
-factor_v + self.rho_out*s_v/(s_no_v+s_v+s_ch)*q - self.rho_in*s_v - self.epsilon*s_v + self.epsilon * s_no_v, # S vaccined
-factor_ch + self.rho_out*s_ch/(s_no_v+s_v+s_ch)*q - self.rho_in*s_ch + self.alpha*self.eta*r, # S children
(factor_no_v + factor_v + factor_ch) - self.theta_no_intv*e - self.theta_intv*e, # E
self.theta_no_intv*e - self.gamma_no_intv*i_no_intv, # I without intervention
self.theta_intv*e - self.gamma_intv*i_intv - self.phi*i_intv + self.lbda*q, # I with intervention
self.gamma_no_intv*i_no_intv + self.gamma_intv*i_intv + self.psi*h - self.alpha*r, # R
self.phi*i_intv-self.psi*h, # H
self.rho_in*(s_no_v + s_v + s_ch) - self.rho_out*q - self.lbda*q # Q
])
# Sanity checks
if test:
assert(abs(sum(x_t_1)) < self.numerical_tolerance_fine) #Test, ob Abgänge und Zugänge über alle Gruppen hinweg ausgleichen
return x_t_1
# class SSSEIIRHQ_Model
# + [markdown] id="ZCbKh3EYQr-_"
# Die **Klasse Data_Reader** stellt die Funktionalitäten für das Auslesen der Datensätze aus einer cSV-Datei zur Verfügung.
# + id="MfmuteWZcNCI"
from numpy.lib.function_base import hamming
from numpy import array as vector
from numpy import matrix as matrix
import numpy as np
import csv
class Data_Reader:
"""
Die Data_Reader-Klasse zum Auslesen der Daten aus einer csv-Datei. Für jedes Modell
existiert eine eigene Funktion, die die entsprechenden Daten, die das Modell benötigt,
zurückliefert.
"""
#Auslesen und Zurückgeben der Daten für das SEIR-Modell
def read_seir_from_csv_file(self, filename):
print("Parse SEIR data from CSV file " + filename + " ...")
with open(filename, encoding='utf-8-sig') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=";")
for row in csv_reader:
#Parameters
if row[0] == "beta":
beta = float(row[1].replace(",","."))
elif row[0] == "theta":
theta = float(row[1].replace(",","."))
elif row[0] == "gamma":
gamma = float(row[1].replace(",","."))
#SEIR
elif row[0] == "N":
n = float(row[1].replace(",","."))
elif row[0] == "S":
s = float(row[1].replace(",","."))
elif row[0] == "E":
e = float(row[1].replace(",","."))
elif row[0] == "I":
i = float(row[1].replace(",","."))
elif row[0] == "R":
r = float(row[1].replace(",","."))
else:
print("Unknown data field: " + str(row[0]))
assert(False)
x0 = (s, e, i, r)
packed_data = [ n,
beta,
theta,
gamma,
x0 ]
return packed_data
# Read SEIIRHQ Model
def read_seiirhq_from_csv_file(self, filename):
print("Parse SEIIRHQ data from CSV file " + filename + " ...")
with open(filename, encoding='utf-8-sig') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=";")
for row in csv_reader:
#Parameters
if row[0] == "beta_no_intv":
beta_no_intv = float(row[1].replace(",","."))
elif row[0] == "beta_intv":
beta_intv = float(row[1].replace(",","."))
elif row[0] == "theta_no_intv":
theta_no_intv = float(row[1].replace(",","."))
elif row[0] == "theta_intv":
theta_intv = float(row[1].replace(",","."))
elif row[0] == "gamma_no_intv":
gamma_no_intv = float(row[1].replace(",","."))
elif row[0] == "gamma_intv":
gamma_intv = float(row[1].replace(",","."))
elif row[0] == "alpha":
alpha = float(row[1].replace(",","."))
elif row[0] == "rho_in":
rho_in = float(row[1].replace(",","."))
elif row[0] == "rho_out":
rho_out = float(row[1].replace(",","."))
elif row[0] == "chi":
chi = float(row[1].replace(",","."))
elif row[0] == "phi":
phi = float(row[1].replace(",","."))
elif row[0] == "lambda":
lbda = float(row[1].replace(",","."))
elif row[0] == "psi":
psi = float(row[1].replace(",","."))
#SEIIRHQ
elif row[0] == "N":
n = float(row[1].replace(",","."))
elif row[0] == "S":
s = float(row[1].replace(",","."))
elif row[0] == "E":
e = float(row[1].replace(",","."))
elif row[0] == "I_no_intv":
i_no_intv = float(row[1].replace(",","."))
elif row[0] == "I_intv":
i_intv = float(row[1].replace(",","."))
elif row[0] == "R":
r = float(row[1].replace(",","."))
elif row[0] == "H":
h = float(row[1].replace(",","."))
elif row[0] == "Q":
q = float(row[1].replace(",","."))
else:
print("Unknown data field: " + str(row[0]))
assert(False)
x0 = (s, e, i_no_intv, i_intv, r, h, q)
packed_data = [n, # total of individuals
beta_no_intv, # The contact and infection rate of transmission per contact from the infected without intervention class
beta_intv, # The contact and infection rate of transmission per contact from the infected with intervention class
theta_no_intv, # Transition rate of exposed individuals to the infected without intervention class
theta_intv, # Transition rate of exposed individuals to the infected with intervention class
gamma_no_intv, # Recovery rate of symptomatic infected without intervention to recovered
gamma_intv, # Recovery rate of symptomatic infected with intervention to recovered
alpha, # Temporary immunity rate
rho_in, # Transition rate between the subsceptible class and quarantined class
rho_out, # Transition rate between the quarantined class and the susceptible class
chi, # Probability of transmission per contact from exposed individuals
phi, # Rate of infectious with symptoms to hospitalized
lbda, # Rate of the quarantined class to the infectious class with intervention
psi, # Recovered rate of hospitalized / quarantined infected individuals
x0] # time 0
return packed_data
# Read SSSEIIRHQ Model
def read_ssseiirhq_from_csv_file(self, filename):
print("Parse SSSEIIRQH data from CSV file " + filename + " ...")
with open(filename, encoding='utf-8-sig') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=";")
for row in csv_reader:
#Parameters
if row[0] == "beta_no_intv":
beta_no_intv = vector(self._to_float(row[1:4]))
elif row[0] == "beta_intv":
beta_intv = vector(self._to_float(row[1:4]))
elif row[0] == "theta_no_intv":
theta_no_intv = float(row[1].replace(",","."))
elif row[0] == "theta_intv":
theta_intv = float(row[1].replace(",","."))
elif row[0] == "gamma_no_intv":
gamma_no_intv = float(row[1].replace(",","."))
elif row[0] == "gamma_intv":
gamma_intv = float(row[1].replace(",","."))
elif row[0] == "alpha":
alpha = float(row[1].replace(",","."))
elif row[0] == "rho_in":
rho_in = float(row[1].replace(",","."))
elif row[0] == "rho_out":
rho_out = float(row[1].replace(",","."))
elif row[0] == "chi":
chi = vector(self._to_float(row[1:4]))
elif row[0] == "phi":
phi = float(row[1].replace(",","."))
elif row[0] == "lambda":
lbda = float(row[1].replace(",","."))
elif row[0] == "psi":
psi = float(row[1].replace(",","."))
elif row[0] == "epsilon":
epsilon = float(row[1].replace(",","."))
elif row[0] == "eta":
eta = float(row[1].replace(",","."))
#SSSEIIRHQ
elif row[0] == "N":
n = float(row[1].replace(",","."))
elif row[0] == "S_not_vaccined":
s_no_v = float(row[1].replace(",","."))
elif row[0] == "S_vaccined":
s_v = float(row[1].replace(",","."))
elif row[0] == "S_children":
s_ch = float(row[1].replace(",","."))
elif row[0] == "E":
e = float(row[1].replace(",","."))
elif row[0] == "I_no_intv":
i_no_intv = float(row[1].replace(",","."))
elif row[0] == "I_intv":
i_intv = float(row[1].replace(",","."))
elif row[0] == "R":
r = float(row[1].replace(",","."))
elif row[0] == "H":
h = float(row[1].replace(",","."))
elif row[0] == "Q":
q = float(row[1].replace(",","."))
else:
print("Unknown data field: " + str(row[0]))
assert(False)
x0 = (s_no_v, s_v, s_ch, e, i_no_intv, i_intv, r, h, q)
packed_data = [n, # total of individuals
beta_no_intv, # The contact and infection rate of transmission per contact from the infected without intervention class
beta_intv, # The contact and infection rate of transmission per contact from the infected with intervention class
theta_no_intv, # Transition rate of exposed individuals to the infected without intervention class
theta_intv, # Transition rate of exposed individuals to the infected with intervention class
gamma_no_intv, # Recovery rate of symptomatic infected without intervention to recovered
gamma_intv, # Recovery rate of symptomatic infected with intervention to recovered
alpha, # Temporary immunity rate
rho_in, # Transition rate between the subsceptible class and quarantined class
rho_out, # Transition rate between the quarantined class and the susceptible class
chi, # Probability of transmission per contact from exposed individuals
phi, # Rate of infectious with symptoms to hospitalized
lbda, # Rate of the quarantined class to the infectious class with intervention
psi, # Recovered rate of hospitalized / quarantined infected individuals
epsilon, # Impfungsrate
eta, # Prozent von Kindern
x0] # time 0
return packed_data
def _to_float(self, my_list):
return [float(x.replace(",",".")) for x in my_list]
# class Data_Reader
# + [markdown] id="RN58N1GCWzua"
# ## Explizites Euler-Verfahren
#
#
# Gelöst werden diese Differenzialgleichungen nun entweder analytisch, wenn klar ist, ob eine Stammfunktion hierzu existiert, oder mithilfe numerischer Verfahren, wenn letzteres unmöglich oder zu aufwendig ist. Das simpelste Verfahren zur Annäherung an die exakte Lösung einer Differenzialgleichung, stellt das Euler-Verfahren dar - welches auch bei unseren Berechnungen Anwendung findet. Bei diesem Algorithmus wird zunächst eine Schrittweite festgelegt, die die Anzahl der Rechenschritte festlegt und gleichzeitig den Grad der Genauigkeit der Lösung bestimmt. Dann werden ausgehend von dem Anfangswert die Punkte eines Graphen ermittelt. Dies geschieht, indem immer von einem Punkt der Lösungsmenge mithilfe des Produktes aus Schrittweite und Ableitung zu dem nächsten Punkt übergegangen wird.
#
# Es ergibt sich folgendes Schema:
#
# h = Schrittweite
# n = Anzahl der Schritte
#
# x0 = X-Wert des Anfangswertproblems
# y0 = Y-Wert des Anfangswertproblems
#
# k = 0
#
# Wiederhole:
#
# xk+1 = x0 + k*h
#
# yk+1 = yk + h* ḟ(xk)
# bis k = n
#
# + id="WAdzV8A9cF3-"
class Explicit_Euler:
"""
Klasse zur numerischen Berechnung von Differenzialgleichungen.
Mittels Numpy-Vektor werden in einem Berechnungsschritt immer
alle Differenzialgleichungen gleichzeitig berechnet
in:
ode = das jeweilige Modell der Differenzialgleichungen (ordinary differential equation): seir, seiirhq, ssseiirhq
stepsize = Schrittgröße
"""
def __init__(self, ode, stepsize):
self.ode = ode
self.stepsize = stepsize
#eigentlicher Euler-Algorithmus
#in:
#
#t_start = Beginn des Zeitraums
#x_start = Vektor mit Startwerten für jede einzelne Kategorie
#t_end = Ende des Zeitraums
#
#out:
# results = mehrdimensionales Array mit Werten für jede Kategorie pro Zeiteinheit
def solve(self, t_start, x_start, t_end):
print("Solve model with explicit Euler scheme ...")
t = t_start
x = x_start
results = [[t, x]]
nr_of_steps = int((t_end - t_start) / self.stepsize) + 1
print("Number of steps: " + str(nr_of_steps))
for k in range(0, nr_of_steps):
x = x + self.stepsize * self.ode.eval_rhs(x)
t = t_start + k * self.stepsize
results.append([t, x])
return results
# class Explicit_Euler
# + [markdown] id="KiKZLmHVQI_a"
# Mithilfe der Klassen **SEIR_Result_Analyzer, SEIIRHQ_Result_Analyzer und SSSEIIRHQ_Result_Analyzer** werden die vorher berechneten Daten aufbereitet und für die Ausgabe vorbereitet.
# + id="TW5Ctq6WlSyk"
class SEIR_Result_Analyzer:
"""
Die Klasse SEIR_Result_Analyzer erhält die vorher ermittelten Daten aus den Euler-Berechnungen und bereitet sie für die
weitere Verwendung auf, d.h. für jeden Graphen (S, E, I, R), der später zu sehen sein wird eine Punktmenge generiert, die via matplot angezeigt
werden kann. Außerdem wird noch die Dauer der Pandemie berechnet.
in:
results = mehrdimensionales Array der per Euler-Verfahren ermittelten Werte für SEIR, jeweils pro Tag
stepsize = Schrittgröße
n = Bevölkerung in Deutschland
t_start = Beginn des Zeitraums
t_end = Ende des Zeitraums
"""
def __init__(self, results, stepsize, n, t_start, t_end):
# Store given values
self.results = results
self.stepsize = stepsize
self.t_start = t_start
self.t_end = t_end
self.n = n
# Sanity check
if test: assert(self.t_start < self.t_end)
# Preparing empty lists (of lists)
self.t_vals = []
self.s_vals = []
self.e_vals = []
self.i_vals = []
self.r_vals = []
self.i_total = []
#Rückgabe der Werte für S, E, I, R und die Zeitschritte t, als Set of lists
def get_extracted_results_as_dict(self):
result_dict = {}
result_dict["t_vals"] = self.t_vals
result_dict["s_vals"] = self.s_vals
result_dict["e_vals"] = self.e_vals
result_dict["i_vals"] = self.i_vals
result_dict["r_vals"] = self.r_vals
return result_dict
#Aufteilen der Werte für s, e, i, r und die Zeitschritte t auf Listen
def extract_results(self):
# Extract and separate results
for res in self.results:
if test: assert(len(res) == 2) # entry is of type [t, x]
self.t_vals.append(res[0])
x = res[1]
if test: assert(len(x) == 4)
if test: assert(x[0] >= 0.0)
self.s_vals.append(x[0])
if test: assert(x[1] >= 0.0)
self.e_vals.append(x[1])
if test: assert(x[2] >= 0.0)
self.i_vals.append(x[2])
if test: assert(x[3] >= 0.0)
self.r_vals.append(x[3])
self.i_total = self._compute_total_i_cases()
def _compute_total_i_cases(self):
i_total = []
for index, t in enumerate(self.t_vals):
i_total_at_t = 0.0
if test: assert(len(self.t_vals) == len(self.i_vals))
if test: assert(self.i_vals[index] >= 0.0)
if test: assert(self.i_vals[index] <= self.n)
i_total_at_t += self.i_vals[index]
i_total.append(i_total_at_t)
if test: assert(len(i_total) == len(self.t_vals))
return i_total
def _compute_duration_of_pandemic(self):
# Compute duration of pandemic
threshold = 100.0
time_points_below_threshold = 0
i_below_at_end = (self.i_total[-1] < threshold)
for val in reversed(self.i_total):
if val < threshold:
time_points_below_threshold += 1
else:
break
number_of_days = self.t_end - self.t_start
days_below_threshold = time_points_below_threshold * self.stepsize
duration_of_pandemic = number_of_days - days_below_threshold
return duration_of_pandemic, i_below_at_end
def compute_and_write_results(self):
duration_of_pandemic, i_below_at_end = self._compute_duration_of_pandemic()
print('Dauer der Simulation in Tagen:', self.t_end)
if duration_of_pandemic < self.t_end:
print('Dauer der Pandemie:', duration_of_pandemic,'Tage')
else:
print('Dauer der Pandemie: mehr als ', duration_of_pandemic,'Tage')
if i_below_at_end:
print('Die Pandemie ist am letzen Simulationstag schon beendet.')
else:
print('Die Pandemie ist am letzen Simulationstag noch nicht zu Ende')
class SEIIRHQ_Result_Analyzer:
def __init__(self, results, stepsize, n, t_start, t_end):
# Store given values
self.results = results
self.stepsize = stepsize
self.t_start = t_start
self.t_end = t_end
self.n = n
# Sanity check
if test: assert(self.t_start < self.t_end)
# Preparing empty lists (of lists)
self.t_vals = []
self.s_vals = []
self.e_vals = []
self.i_no_intv_vals = []
self.i_intv_vals = []
self.r_vals = []
self.h_vals = []
self.q_vals = []
self.i_no_intv_total = []
def get_extracted_results_as_dict(self):
result_dict = {}
result_dict["t_vals"] = self.t_vals
result_dict["s_vals"] = self.s_vals
result_dict["e_vals"] = self.e_vals
result_dict["i_no_intv_vals"] = self.i_no_intv_vals
result_dict["i_intv_vals"] = self.i_intv_vals
result_dict["r_vals"] = self.r_vals
result_dict["h_vals"] = self.h_vals
result_dict["q_vals"] = self.q_vals
return result_dict
def extract_results(self):
# Extract and separate results
for res in self.results:
if test: assert(len(res) == 2) # entry is of type [t, x]
self.t_vals.append(res[0])
x = res[1]
if test: assert(len(x) == 7)
if test: assert(x[0] >= 0.0)
self.s_vals.append(x[0])
if test: assert(x[1] >= 0.0)
self.e_vals.append(x[1])
if test: assert(x[2] >= 0.0)
self.i_no_intv_vals.append(x[2])
if test: assert(x[3] >= 0.0)
self.i_intv_vals.append(x[3])
if test: assert(x[4] >= 0.0)
self.r_vals.append(x[4])
if test: assert(x[5] >= 0.0)
self.h_vals.append(x[5])
if test: assert(x[6] >= 0.0)
self.q_vals.append(x[6])
self.i_no_intv_total = self._compute_total_i_no_intv_cases()
def compute_and_write_results(self):
duration_of_pandemic, i_not_intv_below_at_end = self._compute_duration_of_pandemic()
print('Dauer der Simulation in Tagen:', self.t_end)
if duration_of_pandemic < self.t_end:
print('Dauer der Pandemie:', duration_of_pandemic,'Tage')
else:
print('Dauer der Pandemie: mehr als ', duration_of_pandemic,'Tage')
if i_not_intv_below_at_end:
print('Die Pandemie ist am letzen Simulationstag schon beendet.')
else:
print('Die Pandemie ist am letzen Simulationstag noch nicht zu Ende')
def _compute_total_i_no_intv_cases(self):
i_no_intv_total = []
for index, t in enumerate(self.t_vals):
i_no_intv_total_at_t = 0.0
if test: assert(len(self.t_vals) == len(self.i_no_intv_vals))
if test: assert(self.i_no_intv_vals[index] >= 0.0)
if test: assert(self.i_no_intv_vals[index] <= self.n)
i_no_intv_total_at_t += self.i_no_intv_vals[index]
i_no_intv_total.append(i_no_intv_total_at_t)
if test: assert(len(i_no_intv_total) == len(self.t_vals))
return i_no_intv_total
def _compute_duration_of_pandemic(self):
# Compute duration of pandemic
threshold = 100.0
time_points_below_threshold = 0
i_not_intv_below_at_end = (self.i_no_intv_total[-1] < threshold)
for val in reversed(self.i_no_intv_total):
if val < threshold:
time_points_below_threshold += 1
else:
break
number_of_days = self.t_end - self.t_start
days_below_threshold = time_points_below_threshold * self.stepsize
duration_of_pandemic = number_of_days - days_below_threshold
return duration_of_pandemic, i_not_intv_below_at_end
class SSSEIIRHQ_Result_Analyzer:
def __init__(self, results, stepsize, n, t_start, t_end):
# Store given values
self.results = results
self.stepsize = stepsize
self.t_start = t_start
self.t_end = t_end
self.n = n
# Sanity check
if test: assert(self.t_start < self.t_end)
# Preparing empty lists (of lists)
self.t_vals = []
self.s_no_v_vals = []
self.s_v_vals = []
self.s_ch_vals = []
self.e_vals = []
self.i_no_intv_vals = []
self.i_intv_vals = []
self.r_vals = []
self.h_vals = []
self.q_vals = []
self.i_no_intv_total = []
def get_extracted_results_as_dict(self):
result_dict = {}
result_dict["t_vals"] = self.t_vals
result_dict["s_no_v_vals"] = self.s_no_v_vals
result_dict["s_v_vals"] = self.s_v_vals
result_dict["s_ch_vals"] = self.s_ch_vals
result_dict["e_vals"] = self.e_vals
result_dict["i_no_intv_vals"] = self.i_no_intv_vals
result_dict["i_intv_vals"] = self.i_intv_vals
result_dict["r_vals"] = self.r_vals
result_dict["h_vals"] = self.h_vals
result_dict["q_vals"] = self.q_vals
return result_dict
def extract_results(self):
# Extract and separate results
for res in self.results:
assert(len(res) == 2) # entry is of type [t, x]
self.t_vals.append(res[0])
x = res[1]
if test: assert(len(x) == 9)
if test: assert(x[0] >= 0.0)
self.s_no_v_vals.append(x[0])
if test: assert(x[1] >= 0.0)
self.s_v_vals.append(x[1])
if test: assert(x[2] >= 0.0)
self.s_ch_vals.append(x[2])
if test: assert(x[3] >= 0.0)
self.e_vals.append(x[3])
if test: assert(x[4] >= 0.0)
self.i_no_intv_vals.append(x[4])
if test: assert(x[5] >= 0.0)
self.i_intv_vals.append(x[5])
if test: assert(x[6] >= 0.0)
self.r_vals.append(x[6])
if test: assert(x[7] >= 0.0)
self.h_vals.append(x[7])
if test: assert(x[8] >= 0.0)
self.q_vals.append(x[8])
self.i_no_intv_total = self._compute_total_i_no_intv_cases()
def compute_and_write_results(self):
duration_of_pandemic, i_not_intv_below_at_end = self._compute_duration_of_pandemic()
print('Dauer der Simulation in Tagen:', self.t_end)
if (duration_of_pandemic < self.t_end ):
print('Dauer der Pandemie:', duration_of_pandemic,'Tage')
else:
print('Dauer der Pandemie: mehr als ', duration_of_pandemic,'Tage')
if i_not_intv_below_at_end:
print('Die Pandemie ist am letzen Simulationstag schon beendet.')
else:
print('Die Pandemie ist am letzen Simulationstag noch nicht zu Ende')
def _compute_total_i_no_intv_cases(self):
i_no_intv_total = []
for index, t in enumerate(self.t_vals):
i_no_intv_total_at_t = 0.0
if test: assert(len(self.t_vals) == len(self.i_no_intv_vals))
if test: assert(self.i_no_intv_vals[index] >= 0.0)
if test: assert(self.i_no_intv_vals[index] <= self.n)
i_no_intv_total_at_t += self.i_no_intv_vals[index]
i_no_intv_total.append(i_no_intv_total_at_t)
if test: assert(len(i_no_intv_total) == len(self.t_vals))
return i_no_intv_total
def _compute_duration_of_pandemic(self):
# Compute duration of pandemic
threshold = 100.0
time_points_below_threshold = 0
i_not_intv_below_at_end = (self.i_no_intv_total[-1] < threshold)
for val in reversed(self.i_no_intv_total):
if val < threshold:
time_points_below_threshold += 1
else:
break
number_of_days = self.t_end - self.t_start
days_below_threshold = time_points_below_threshold * self.stepsize
duration_of_pandemic = number_of_days - days_below_threshold
return duration_of_pandemic, i_not_intv_below_at_end
# + [markdown] id="CbBaPCfGPs4d"
# Die folgenden Klassen (**SEIR_Visualizer, SEIIRHQ_Visualizer und SSEIIRHQ_Visualizer**) sind für die grafische Ausgabe der Daten zutsändig.
# + id="UZYpTJ--1lqn"
import matplotlib.pyplot as plot
import numpy as np
import pandas as pd
class SEIR_Visualizer:
def __init__(self, result_dict, n, t_start, t_end):
# Store given values
self.result_dict = result_dict
self.n = n
self.t_start = t_start
self.t_end = t_end
self.dfa=self.read_actuals()
# Sanity checks
assert(self.t_start < self.t_end)
# Unpack results
self.t_vals = self.result_dict["t_vals"]
self.s_vals = self.result_dict["s_vals"]
self.e_vals = self.result_dict["e_vals"]
self.i_vals = self.result_dict["i_vals"]
self.r_vals = self.result_dict["r_vals"]
def plot_curves(self):
plot.style.use('fivethirtyeight')
figure,axes = plot.subplots(figsize=(8, 6))
figure.subplots_adjust(bottom = 0.15)
axes.grid(linestyle = ':', linewidth = 0.2, color = "#808080")
axes.set_xlabel("Days", fontsize=14)
axes.set_ylabel("Individuals", fontsize=14)
axes.tick_params(axis='both', which='major', labelsize=14)
axes.plot(self.t_vals, self.s_vals, label='Subsceptible', color = "#0000cc")
axes.plot(self.t_vals, self.e_vals, color = "#ffb000", label='Exposed', linestyle = '--')
axes.plot(self.t_vals, self.i_vals, label='Infectious', color = "#a00060")
axes.plot(self.t_vals, self.r_vals, color = "#008000", label='Recovered', linestyle = '--')
axes.plot(self.dfa.days, self.dfa['Individuals cumulative'], color = "wheat", label='actual infectious', linestyle = '--')
plot.yticks(np.arange(0, 8e7, 1e7), ['0',
r'$10\,$M',
r'$20\,$M',
r'$30\,$M',
r'$40\,$M',
r'$50\,$M',
r'$60\,$M',
r'$70\,$M'])
axes.set_xlim([0,t_end ])
plot.legend()
plot.show()
def read_actuals(self):
dfa = pd.read_excel(
os.path.join('./data', 'actual_numbers', 'Nowcast_R_aktuell.xlsx'),
engine='openpyxl')
dfa['days'] = np.arange(len(dfa))
dfa.rename(columns={'PS_COVID_Faelle': 'Individuals'}, inplace=True)
dfa['Individuals cumulative'] = dfa.Individuals.cumsum()
return dfa
class SEIIRHQ_Visualizer:
def __init__(self, result_dict, n, t_start, t_end):
# Store given values
self.result_dict = result_dict
self.n = n
self.t_start = t_start
self.t_end = t_end
self.dfa=self.read_actuals()
# Sanity checks
assert(self.t_start < self.t_end)
# Unpack results
self.t_vals = self.result_dict["t_vals"]
self.s_vals = self.result_dict["s_vals"]
self.e_vals = self.result_dict["e_vals"]
self.i_no_intv_vals = self.result_dict["i_no_intv_vals"]
self.i_intv_vals = self.result_dict["i_intv_vals"]
self.r_vals = self.result_dict["r_vals"]
self.h_vals = self.result_dict["h_vals"]
self.q_vals = self.result_dict["q_vals"]
def plot_curves(self):
"""
Um die hohen Zahlen der Gruppe 'Subsceptible' anzuzeigen, und gleichzeitig eine
detaillierte wird im Folgenden eine Darstellung gewählt mit einer Lücke in der Y - Achse. Technisch
umgesetzt wird dieser Graph in Form von 2 Subplots, die direkt übereinander gelegt werden
"""
plot.style.use('fivethirtyeight')
figure, (axes1, axes2) = plot.subplots(2, 1, sharex=True,figsize=(10, 16))
figure.subplots_adjust(hspace=0.05)
figure.subplots_adjust(bottom = 0.15)
axes1.grid(linestyle = ':', linewidth = 0.2, color = "#808080")
axes1.set_xlabel("Days", fontsize=14)
axes1.set_ylabel("Individuals", fontsize=14)
axes1.tick_params(axis='both', which='major', labelsize=14)
axes2.grid(linestyle = ':', linewidth = 0.2, color = "#808080")
axes2.set_xlabel("Days", fontsize=14)
axes2.set_ylabel("Individuals", fontsize=14)
axes2.tick_params(axis='both', which='major', labelsize=14)
axes1.plot(self.t_vals, self.s_vals, label='Subsceptible', color = "olive", linewidth = 0.5)
axes1.plot(self.t_vals, self.e_vals, color = "orange", label='Exposed', linewidth = 0.5)
axes1.plot(self.t_vals, self.i_no_intv_vals, label='Infectious without interv.', color = "red", linewidth = 0.5)
axes1.plot(self.t_vals, self.i_intv_vals, label='Infectious with interv.', color = "maroon", linewidth = 0.5)
axes1.plot(self.t_vals, self.r_vals, color = "green", label='Recovered', linewidth = 0.5)
axes1.plot(self.t_vals, self.h_vals, color = "black", label='Hospitalized', linewidth = 0.5)
axes1.plot(self.t_vals, self.q_vals, color = "gray", label='Quarantined', linewidth = 0.5)
axes1.plot(self.dfa.days, self.dfa['Individuals cumulative'], color = "wheat", label='Actual infectious', linestyle = '--')
axes1.plot(self.t_vals, np.add(self.i_no_intv_vals, self.i_intv_vals), label='Infectious (Total)', color = "magenta", linestyle = '--', linewidth = 0.7)
axes2.plot(self.t_vals, self.e_vals, color = "orange", label='Exposed', linewidth = 0.5)
axes2.plot(self.t_vals, self.i_no_intv_vals, label='Infectious without interv.', color = "red", linewidth = 0.5)
axes2.plot(self.t_vals, self.i_intv_vals, label='Infectious with interv.', color = "maroon", linewidth = 0.5)
axes2.plot(self.t_vals, self.r_vals, color = "green", label='Recovered', linewidth = 0.5)
axes2.plot(self.t_vals, self.h_vals, color = "black", label='Hospitalized', linewidth = 0.5)
axes2.plot(self.t_vals, self.q_vals, color = "gray", label='Quarantined', linewidth = 0.5)
axes2.plot(self.dfa.days, self.dfa['Individuals cumulative'], color = "wheat", label='Actual infectious', linestyle = '--')
axes2.plot(self.t_vals, np.add(self.i_no_intv_vals, self.i_intv_vals), label='Infectious (Total)', color = "magenta", linestyle = '--', linewidth = 0.7)
axes1.set_ylim(57000000, 70000000) # nur Susceptible
axes2.set_ylim(0, 22000000) # Alle anderen Gruppen
axes1.spines['bottom'].set_visible(False)
axes2.spines['top'].set_visible(False)
axes1.xaxis.tick_top()
axes1.tick_params(labeltop=False)
axes2.xaxis.tick_bottom()
x_axis = axes1.axes.get_xaxis()
x_axis.set_visible(False)
axes1.set_xlim([0,t_end ])
axes2.set_xlim([0,t_end ])
d = .5
kwargs = dict(marker=[(-1, -d), (1, d)], markersize=12,
linestyle="none", color='k', mec='k', mew=1, clip_on=False)
axes1.plot([0, 1], [0, 0], transform=axes1.transAxes, **kwargs)
axes2.plot([0, 1], [1, 1], transform=axes2.transAxes, **kwargs)
plot.sca(axes2)
plot.yticks(np.arange(0, 3e7, 1e7), ['0',
r'$10\,$M',
r'$20\,$M'])
plot.sca(axes1)
plot.yticks(np.arange(6e7, 8e7, 1e7), [
r'$60\,$M',
r'$70\,$M'])
plot.legend(loc='lower right')
plot.show()
def read_actuals(self):
"""
lese die Daten zum tatsächlichen Pandemieverlauf ein, um diese als Referenz zu plotten
"""
dfa = pd.read_excel(
os.path.join('./data', 'actual_numbers', 'Nowcast_R_aktuell.xlsx'),
engine='openpyxl')
dfa['days'] = np.arange(len(dfa))
dfa.rename(columns={'PS_COVID_Faelle': 'Individuals'}, inplace=True)
dfa['Individuals cumulative'] = dfa.Individuals.cumsum()
return dfa
class SSSEIIRHQ_Visualizer:
def __init__(self, result_dict, n, t_start, t_end):
# Store given values
self.result_dict = result_dict
self.n = n
self.t_start = t_start
self.t_end = t_end
self.dfa=self.read_actuals()
# Sanity checks
assert(self.t_start < self.t_end)
# Unpack results
self.t_vals = self.result_dict["t_vals"]
self.s_no_v_vals = self.result_dict["s_no_v_vals"]
self.s_v_vals = self.result_dict["s_v_vals"]
self.s_ch_vals = self.result_dict["s_ch_vals"]
self.e_vals = self.result_dict["e_vals"]
self.i_no_intv_vals = self.result_dict["i_no_intv_vals"]
self.i_intv_vals = self.result_dict["i_intv_vals"]
self.r_vals = self.result_dict["r_vals"]
self.h_vals = self.result_dict["h_vals"]
self.q_vals = self.result_dict["q_vals"]
def plot_curves(self):
plot.style.use('fivethirtyeight')
figure,axes = plot.subplots(figsize=(8, 16))
figure.subplots_adjust(bottom = 0.15)
axes.grid(linestyle = ':', linewidth = 0.2, color = "#808080")
axes.set_xlabel("Days", fontsize=14)
axes.set_ylabel("Individuals", fontsize=14)
axes.tick_params(axis='both', which='major', labelsize=14)
axes.plot(self.t_vals, self.s_no_v_vals, label='Subsceptible adults not vaccined', color = "olive",linewidth = 0.5)
axes.plot(self.t_vals, self.s_v_vals, label='Subsceptible adults vaccined', color = "violet",linewidth = 0.5)
axes.plot(self.t_vals, self.s_ch_vals, label='Subsceptible children', color = "cyan",linewidth = 0.5)
axes.plot(self.t_vals, self.e_vals, color = "orange", label='Exposed',linewidth = 0.5)
axes.plot(self.t_vals, self.i_no_intv_vals, label='Infectious without interv.', color = "red",linewidth = 0.5)
axes.plot(self.t_vals, self.i_intv_vals, label='Infectious with interv.', color = "maroon",linewidth = 0.5)
axes.plot(self.t_vals, self.r_vals, color = "green", label='Recovered',linewidth = 0.5)
axes.plot(self.t_vals, self.h_vals, color = "black", label='Hospitalized',linewidth = 0.5)
axes.plot(self.t_vals, self.q_vals, color = "gray", label='Quarantined',linewidth = 0.5)
axes.plot(self.t_vals, np.add(self.i_no_intv_vals, self.i_intv_vals), label='Infectious (Total)', color = "magenta", linestyle = '--', linewidth = 0.7)
axes.plot(self.dfa.days, self.dfa['Individuals cumulative'], color = "wheat", label='Actual infectious', linestyle = '--')
plot.yticks(np.arange(0, 8e7, 1e7), ['0',
r'$10\,$M',
r'$20\,$M',
r'$30\,$M',
r'$40\,$M',
r'$50\,$M',
r'$60\,$M',
r'$70\,$M'])
axes.set_xlim([0,t_end ])
plot.legend()
plot.show()
def read_actuals(self):
dfa = pd.read_excel(
os.path.join('./data', 'actual_numbers', 'Nowcast_R_aktuell.xlsx'),
engine='openpyxl')
dfa['days'] = np.arange(len(dfa))
dfa.rename(columns={'PS_COVID_Faelle': 'Individuals'}, inplace=True)
dfa['Individuals cumulative'] = dfa.Individuals.cumsum()
return dfa
# + [markdown] id="2Y5f4WkZPdop"
# Ab hier beginnt der Hauptteil des Programmes, **'Main'**, von dem aus die anderen Klassen aufgerufen werden.
# + id="8JXUiy4Aci8Z"
# Global imports
import os
import time
# Start and end time, stepsize
t_start = 0
t_end = 365 * 3
stepsize = 1e-2
data_reader = Data_Reader()
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="5H27lU-3oSwt" outputId="5331bb86-f1a9-40eb-a1f9-e95d720f23d9"
# Preparing data parsing for SEIR
data_set_name = "SEIR"
data_directory_name = "./data/" + data_set_name + "/"
data_directory = os.fsencode(data_directory_name)
test = True
for file in os.listdir(data_directory):
data_filename = os.fsdecode(file)
if data_filename.endswith(".csv"):
# Parsing the seir data
data_filename_prefix = data_filename.split(".")[0]
packed_data = data_reader.read_seir_from_csv_file(data_directory_name + data_filename_prefix + ".csv")
# Unpacking some of the parsed data
n = packed_data[0]
x0 = packed_data[-1]
print("Setting up the SEIR model")
ode_system = SEIR_Model(packed_data)
# Solve the ODE system
explicit_euler = Explicit_Euler(ode_system, stepsize)
start_time = time.time()
results = explicit_euler.solve(t_start, x0, t_end)
end_time = time.time()
print("Required CPU time = " + str(round(end_time - start_time, 3)) + " seconds")
# Analyze results
result_analyzer = SEIR_Result_Analyzer(results, stepsize, n, t_start, t_end)
result_analyzer.extract_results()
result_dict = result_analyzer.get_extracted_results_as_dict()
result_analyzer.compute_and_write_results()
# Visualize results
visualizer = SEIR_Visualizer(result_dict, n, t_start, t_end)
visualizer.plot_curves()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="YlNFUa6l-bbS" outputId="d8f3f8cd-d5fb-499b-9884-c7f57e73666b"
# Preparing data parsing for SEIIRHQ
data_set_name = "SEIIRHQ"
data_directory_name = "./data/" + data_set_name + "/"
data_directory = os.fsencode(data_directory_name)
test = True
for file in os.listdir(data_directory):
data_filename = os.fsdecode(file)
if data_filename.endswith(".csv"):
# Parsing the seiirhq data
data_filename_prefix = data_filename.split(".")[0]
packed_data = data_reader.read_seiirhq_from_csv_file(data_directory_name + data_filename_prefix + ".csv")
# Unpacking some of the parsed data
n = packed_data[0]
x0 = packed_data[-1]
print("Setting up the SEIIRHQ Model model")
ode_system = SEIIRHQ_Model(packed_data)
# Solve the ODE system
explicit_euler = Explicit_Euler(ode_system, stepsize)
start_time = time.time()
results = explicit_euler.solve(t_start, x0, t_end)
end_time = time.time()
print("Required CPU time = " + str(round(end_time - start_time, 3)) + " seconds")
# Analyze results
result_analyzer = SEIIRHQ_Result_Analyzer(results, stepsize, n, t_start, t_end)
result_analyzer.extract_results()
result_dict = result_analyzer.get_extracted_results_as_dict()
result_analyzer.compute_and_write_results()
# Visualize results
visualizer = SEIIRHQ_Visualizer(result_dict, n, t_start, t_end)
visualizer.plot_curves()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="dJ2wPT11Vzdf" outputId="0939c7e0-1b91-4775-c554-ba6decf58c09"
# Preparing data parsing for SSSEIIRHQ
data_set_name = "SSSEIIRHQ"
data_directory_name = "./data/" + data_set_name + "/"
data_directory = os.fsencode(data_directory_name)
test = True
for file in os.listdir(data_directory):
data_filename = os.fsdecode(file)
if data_filename.endswith(".csv"):
# Parsing the ssseiirhq data
data_filename_prefix = data_filename.split(".")[0]
packed_data = data_reader.read_ssseiirhq_from_csv_file(data_directory_name + data_filename_prefix + ".csv")
# Unpacking some of the parsed data
n = packed_data[0]
x0 = packed_data[-1]
print("Setting up the SSSEIRHQ model")
ode_system = SSSEIIRHQ_Model(packed_data)
# Solve the ODE system
explicit_euler = Explicit_Euler(ode_system, stepsize)
start_time = time.time()
results = explicit_euler.solve(t_start, x0, t_end)
end_time = time.time()
print("Required CPU time = " + str(round(end_time - start_time, 3)) + " seconds")
# Analyze results
result_analyzer = SSSEIIRHQ_Result_Analyzer(results, stepsize, n, t_start, t_end)
result_analyzer.extract_results()
result_dict = result_analyzer.get_extracted_results_as_dict()
result_analyzer.compute_and_write_results()
# Visualize results
visualizer = SSSEIIRHQ_Visualizer(result_dict, n, t_start, t_end)
visualizer.plot_curves()
# + [markdown] id="NYvlMYPsLmzK"
# ## Diskussion und Fazit
# Eine zunehmend detailliertere Betrachtung der Deutschen Bevölkerung ermöglich ein genaueres Verständnis des Pandemieverlaufes im Vergleich zum "einfachen" SEIR Modell. Insbesondere die Modellierung verschiedener Krankheitsverläufe, sowie die Differenzierung nach unterschiedlich gefährdeten Bevölkerungsgruppen wird der Gesamtgesellschaftlichen Debatte um das Coronavirus deutlich besser gerecht, als das ursprüngliche SEIR Modell.
# Im Umgang mit dem Coronavirus gibt es 3 erstrebenswerte Ziele, die sich aber gegenseitig ausschließen:
# 1. Ein schnelles Ende der Pandemie
# 2. Die Überlastung der Intensivstationen verhindern
# 3. Möglichst geringe Todeszahlen
#
# Maßnahmen der Pandemieeinämmung ("Lockdowns") verlängern dabei die Dauer der Pandemie, senken aber im Gegenzug die Todeszahlen und verhindern eine Überlastund des Gesundheitssystems. Im Vergleich mit der tatsächlichen Realität zeigt sich dabei, dass die Pandemie bereits deutlich länger andauert, als die verschiedenen SEIR Modelle das vorhergesagt haben, und es aber insgesamt deutlich weniger erkranke Personen gibt.
# So gibt es im ursprünglichen SEIR Modell in Spitze über 50 Millionen "Exposed" Personen. Diese Vorstellung wäre in der Realität eine gesellschaftliche Katastrophe, mit unseren Anpassungen des Modells konnten wir der Realität deutlich näher kommen
# Gründe für längere Dauer der Pandemie sind vermutlich neben den Lockdown-Maßnahmen insbesondere die verschiedenen Corona Varianten, sowie die Möglichkeit, mehrfach an Corona zu erkranken. Diese beiden Aspekte sind allerdings jenseits des Horizontes dieser Ausarbeitung
#
#
# + [markdown] id="-q3WtoSqbjQ3"
# ## Quellen:
# - https://de.wikipedia.org/wiki/SEIR-Modell
# - https://de.wikipedia.org/wiki/Explizites_Euler-Verfahren
# - https://link.springer.com/article/10.1007/s11071-020-05743-y
# - https://www.nature.com/articles/s41598-021-83540-2
# - https://github.com/m-schmidt-math-opt/covid-19-extended-seir-model
#
|
SEIR_Erweiterung_Corona.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Support vector machine applicate a XOR
# +
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# -
import numpy as np
from sklearn import svm
from sklearn.kernel_approximation import RBFSampler
from sklearn.linear_model import SGDClassifier
# +
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
from matplotlib import cm
plt.style.use('fivethirtyeight')
plt.rcParams['font.family'] = 'sans-serif'
plt.rcParams['font.serif'] = 'Ubuntu'
plt.rcParams['font.monospace'] = 'Ubuntu Mono'
plt.rcParams['font.size'] = 10
plt.rcParams['axes.labelsize'] = 10
plt.rcParams['axes.labelweight'] = 'bold'
plt.rcParams['axes.titlesize'] = 10
plt.rcParams['xtick.labelsize'] = 8
plt.rcParams['ytick.labelsize'] = 8
plt.rcParams['legend.fontsize'] = 10
plt.rcParams['figure.titlesize'] = 12
plt.rcParams['image.cmap'] = 'jet'
plt.rcParams['image.interpolation'] = 'none'
plt.rcParams['figure.figsize'] = (16, 8)
plt.rcParams['lines.linewidth'] = 2
plt.rcParams['lines.markersize'] = 8
colors = ['#008fd5', '#fc4f30', '#e5ae38', '#6d904f', '#8b8b8b', '#810f7c',
'#137e6d', '#be0119', '#3b638c', '#af6f09', '#008fd5', '#fc4f30', '#e5ae38', '#6d904f', '#8b8b8b',
'#810f7c', '#137e6d', '#be0119', '#3b638c', '#af6f09']
cmap = mcolors.LinearSegmentedColormap.from_list("", ["#82cafc", "#069af3", "#0485d1", colors[0], colors[8]])
cmap_big = cm.get_cmap('Spectral', 512)
cmap = mcolors.ListedColormap(cmap_big(np.linspace(0.5, 1, 128)))
# -
xx, yy = np.meshgrid(np.linspace(-3, 3, 500),
np.linspace(-3, 3, 500))
np.random.seed(0)
X = np.random.randn(300, 2)
Y = np.logical_xor(X[:, 0] > 0, X[:, 1] > 0)
fig = plt.figure(figsize=(16,8))
fig.patch.set_facecolor('white')
for i in range(2):
idx = np.where(Y == i)
plt.scatter(X[idx, 0], X[idx, 1], c=colors[i], s=40, edgecolors='k', alpha = .9, label='Class {0:d}'.format(i),cmap=cmap)
plt.xlabel('$x_1$', fontsize=14)
plt.ylabel('$x_2$', fontsize=14)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.xlim(-3, 3)
plt.ylim(-3, 3)
plt.legend()
plt.show()
# fit the model
#clf= svm.SVC(gamma=10)
clf=svm.SVC(kernel='linear')
#clf=svm.SVC(kernel='poly', degree=3, coef0=1)
#clf=svm.SVC(kernel='sigmoid', gamma=15)
clf = clf.fit(X, Y)
# +
# plot the decision function for each datapoint on the grid
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
fig = plt.figure(figsize=(16,8))
fig.patch.set_facecolor('white')
ax = fig.gca()
imshow_handle = plt.imshow(Z, interpolation='nearest',
extent=(xx.min(), xx.max(), yy.min(), yy.max()), aspect='auto',
origin='lower', alpha=.5, cmap=cmap)
contours = plt.contour(xx, yy, Z, levels=[0], linewidths=2,
linetypes='--', colors=[colors[9]])
for i in range(2):
idx = np.where(Y == i)
plt.scatter(X[idx, 0], X[idx, 1], c=colors[i], edgecolors='k', s=40,
label='Class {0:d}'.format(i),cmap=cmap)
plt.xlabel('$x_1$', fontsize=14)
plt.ylabel('$x_2$', fontsize=14)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.xlim(-3, 3)
plt.ylim(-3, 3)
plt.legend()
plt.show()
# -
print('Accuracy: {0:3.5f}'.format(np.sum(Y==clf.predict(X))/float(X.shape[0])*100))
# # Gradient descent con hinge loss
rbf_feature = RBFSampler(gamma=10, random_state=1)
X_features = rbf_feature.fit_transform(X)
clf = SGDClassifier(max_iter=1000, penalty='l2', alpha=.001)
clf = clf.fit(X_features, Y)
print('Accuracy: {0:3.5f}'.format(np.sum(Y==clf.predict(X_features))/float(X_features.shape[0])*100))
# +
Z = clf.predict(rbf_feature.fit_transform(np.c_[xx.ravel(), yy.ravel()]))
Z = Z.reshape(xx.shape)
fig = plt.figure()
fig.patch.set_facecolor('white')
ax = fig.gca()
imshow_handle = plt.imshow(Z, interpolation='nearest',
extent=(xx.min(), xx.max(), yy.min(), yy.max()), aspect='auto',
origin='lower', alpha=.3)
contours = plt.contour(xx, yy, Z, levels=[0], linewidths=1,
linetypes='--')
for i in range(2):
idx = np.where(Y == i)
plt.scatter(X[idx, 0], X[idx, 1], c=colors[i], edgecolors='k', s=40,
label='Class {0:d}'.format(i),cmap=cmap)
plt.xlabel('$x_1$', fontsize=14)
plt.ylabel('$x_2$', fontsize=14)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.xlim(-3, 3)
plt.ylim(-3, 3)
plt.legend()
plt.show()
# -
|
codici/.ipynb_checkpoints/svm_xor-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tf-gpu)
# language: python
# name: tf-gpu
# ---
import os, sys
os.environ["CUDA_VISIBLE_DEVICES"]="1"
import cv2
from matplotlib import pyplot as plt
from IPython.display import SVG, display
from keras.utils.vis_utils import model_to_dot
import pandas as pd
import numpy as np
from keras.layers.merge import _Merge
from numpy.random import randn
from keras.models import Sequential, Model
from keras.layers import Activation, Dropout, Multiply, LSTM,Embedding, Conv2D, Dense, \
Conv2DTranspose, Lambda, Input, Concatenate, TimeDistributed, \
MaxPooling2D, Flatten, BatchNormalization, GlobalAveragePooling2D, Reshape, LeakyReLU
from keras.initializers import RandomNormal
from keras.callbacks import ModelCheckpoint, TensorBoard
import tensorflow as tf
from keras import backend as k
from keras.applications import InceptionResNetV2
from keras.optimizers import Adadelta,Adam,RMSprop
from tqdm import tqdm_notebook
from keras.utils import to_categorical
from keras.datasets import fashion_mnist
import keras
from SpectralNormalization import ConvSN2D, DenseSN, ConvSN2DTranspose
from self_attention import Attention
from functools import partial
gpu_options = tf.GPUOptions(allow_growth=True)
sess = tf.Session(config=tf.ConfigProto(gpu_options=gpu_options))
keras.backend.tensorflow_backend.set_session(sess)
def load_real_samples(class_number=0, categorical = False):
# load dataset
(trainX, trainy), (testX, testy) = fashion_mnist.load_data()
if class_number>-1:
selected_ix = (trainy == class_number)
selected_ix = np.squeeze(selected_ix)
trainX = trainX[selected_ix]
trainy = trainy[selected_ix]
else:
pass
# expand to 3d, e.g. add channels
# X = np.expand_dims(trainX, axis=-1)
# convert from ints to floats
trainX = trainX.astype('float32')
testX = testX.astype('float32')
trainX = np.repeat(np.expand_dims(trainX,axis=-1),repeats=3,axis=-1)
testX = np.repeat(np.expand_dims(testX,axis=-1),repeats=3,axis=-1)
# scale from [0,255] to [-1,1]
trainX = (trainX-127.5)/127.5
testX = (testX-127.5)/127.5
if categorical:
trainy = to_categorical(trainy)
testy = to_categorical(testy)
return trainX, trainy, testX, testy
def generate_real_samples(dataset, conditions, n_samples):
# choose random instances
ix = np.random.randint(0, dataset.shape[0], n_samples)
# print(ix)
# select images
X = dataset[ix]
X_cond = conditions[ix]
# generate class labels, -1 for 'real'
# y = -np.ones((n_samples, 1))
return X, X_cond
X, X_cond, T, T_cond = load_real_samples(class_number=-1, categorical=True)
# +
# tg = dataGenerator(mode='train')
# vg = dataGenerator(mode='val')
# -
X.shape, X_cond.shape,T.shape, T_cond.shape
x,c = generate_real_samples(T,T_cond,100)
plt.imshow((x[0]+1)/2)
c[0]
# +
randomDim = 90
# Optimizer Hyperparameters
gen_lr = 0.0001
gen_beta1 = 0.0
gen_beta2 = 0.999
disc_lr = 0.0004
disc_beta1 = 0.0
disc_beta2 = 0.999
# The training ratio is the number of discriminator updates
# per generator update. The paper uses 5.
TRAINING_RATIO = 1
GRADIENT_PENALTY_WEIGHT = 10 # As per the paper
# Iteration Hyperparameters
batch_size = 64
eval_batch_size = 10000
start_iter = 1
max_iters = 300000
eval_iters = 1000
save_evals = 10
# +
g_opt = Adam(lr=gen_lr, beta_1=gen_beta1, beta_2=gen_beta2)
d_opt = Adam(lr=disc_lr, beta_1=disc_beta1, beta_2=disc_beta2)
# +
def plot_data(x,ax):
# x = x.reshape((size,size))
ax.imshow((x+1)/2, cmap='gray')
# if y is not None:
# ax.scatter(y[0::2] , y[1::2] , marker='x', s=10)
def plot_images(x,batch_size=100):
fig = plt.figure(figsize=(5,5))
fig.subplots_adjust(left=0, right=1, bottom=0, top=1, hspace=0.01, wspace=0.01)
for i in range(batch_size):
ax = fig.add_subplot(10, 10, i + 1, xticks=[], yticks=[])
# print(y['classes'][i])
# print(np.argmax(c[i]))
plot_data(x[i], ax)
plot_images(X)
# +
import keras.backend as K
def hinge_loss(y_true, y_pred):
return K.mean(K.square(K.maximum(1. - (y_true * y_pred), 0.)), axis=-1)
# +
dLosses = []
gLosses = []
from keras.applications.inception_v3 import InceptionV3
from scipy.linalg import sqrtm
class RandomWeightedAverage(_Merge):
"""Takes a randomly-weighted average of two tensors. In geometric terms, this
outputs a random point on the line between each pair of input points.
Inheriting from _Merge is a little messy but it was the quickest solution I could
think of. Improvements appreciated."""
def _merge_function(self, inputs):
weights = K.random_uniform((batch_size, 1, 1, 1))
return (weights * inputs[0]) + ((1 - weights) * inputs[1])
def plotLoss(epoch):
plt.figure(figsize=(10, 8))
plt.plot(dLosses, label='Discriminitive loss')
plt.plot(gLosses, label='Generative loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend()
plt.savefig('plots/fmnist_cond_hinge_sn_sa_gp/dcgan_%d_loss_epoch.png' % epoch)
plt.close()
def plotGeneratedImages(epoch, examples=100, dim=(1, 1), figsize=(2, 2)):
n_classes = 10
z = np.repeat(np.random.uniform(-1, 1, (10, randomDim)),10,axis=0)
labels = np.tile(np.eye(n_classes), [10, 1])
# labels = np.repeat(np.arange(10).reshape(1,-1),10, axis=0).reshape((-1))
generatedImages = generator.predict([labels, z])
# print(generatedImages.shape)
# generatedImages = generatedImages*255.0 + 255.0
# print(generatedImages.min(),generatedImages.max())
fig = plt.figure(figsize=(5,5))
fig.subplots_adjust(left=0, right=1, bottom=0, top=1, hspace=0.01, wspace=0.01)
for i in range(examples):
ax = fig.add_subplot(10, 10, i + 1, xticks=[], yticks=[])
# print(y['classes'][i])
plot_data(generatedImages[i], ax)
fol = 'generatedimages/fmnist_cond_hinge_sn_sa_gp/'
if not os.path.exists(fol):
os.makedirs(fol)
plt.savefig(fol+'random_{:05d}.png'.format(epoch))
plt.close()
def saveModels(epoch):
fol = 'models/fmnist_cond_hinge_sn_sa_gp/'
if not os.path.exists(fol):
os.makedirs(fol)
generator.save(fol+'dcgan_generator_epoch_%d.h5' % epoch)
discriminator.save(fol+'dcgan_discriminator_epoch_%d.h5' % epoch)
# scale an array of images to a new size
def scale_images(images, new_shape):
images_list = list()
for image in images:
# resize with nearest neighbor interpolation
new_image = cv2.resize(image,(new_shape),interpolation=cv2.INTER_LINEAR)
# store
images_list.append(new_image)
return np.asarray(images_list)
# calculate frechet inception distance
def calculate_fid(model, images1, images2):
# calculate activations
act1 = model.predict(images1)
act2 = model.predict(images2)
# calculate mean and covariance statistics
mu1, sigma1 = act1.mean(axis=0), np.cov(act1, rowvar=False)
mu2, sigma2 = act2.mean(axis=0), np.cov(act2, rowvar=False)
# calculate sum squared difference between means
ssdiff = np.sum((mu1 - mu2)**2.0)
# calculate sqrt of product between cov
covmean = sqrtm(sigma1.dot(sigma2))
# check and correct imaginary numbers from sqrt
if np.iscomplexobj(covmean):
covmean = covmean.real
# calculate score
fid = ssdiff + np.trace(sigma1 + sigma2 - 2.0 * covmean)
return fid
def gradient_penalty_loss(y_true, y_pred, averaged_samples,
gradient_penalty_weight):
"""Calculates the gradient penalty loss for a batch of "averaged" samples.
In Improved WGANs, the 1-Lipschitz constraint is enforced by adding a term to the
loss function that penalizes the network if the gradient norm moves away from 1.
However, it is impossible to evaluate this function at all points in the input
space. The compromise used in the paper is to choose random points on the lines
between real and generated samples, and check the gradients at these points. Note
that it is the gradient w.r.t. the input averaged samples, not the weights of the
discriminator, that we're penalizing!
In order to evaluate the gradients, we must first run samples through the generator
and evaluate the loss. Then we get the gradients of the discriminator w.r.t. the
input averaged samples. The l2 norm and penalty can then be calculated for this
gradient.
Note that this loss function requires the original averaged samples as input, but
Keras only supports passing y_true and y_pred to loss functions. To get around this,
we make a partial() of the function with the averaged_samples argument, and use that
for model training."""
# first get the gradients:
# assuming: - that y_pred has dimensions (batch_size, 1)
# - averaged_samples has dimensions (batch_size, nbr_features)
# gradients afterwards has dimension (batch_size, nbr_features), basically
# a list of nbr_features-dimensional gradient vectors
gradients = K.gradients(y_pred, averaged_samples)[0]
# compute the euclidean norm by squaring ...
gradients_sqr = K.square(gradients)
# ... summing over the rows ...
gradients_sqr_sum = K.sum(gradients_sqr,
axis=np.arange(1, len(gradients_sqr.shape)))
# ... and sqrt
gradient_l2_norm = K.sqrt(gradients_sqr_sum)
# compute lambda * (1 - ||grad||)^2 still for each single sample
gradient_penalty = gradient_penalty_weight * K.square(1 - gradient_l2_norm)
# return the mean as loss over all the batch samples
return K.mean(gradient_penalty)
# -
model = InceptionV3(include_top=False, pooling='avg', input_shape=(299,299,3))
model.trainable = False
model.trainable_weights
def build_generator():
condition = Input(shape=(10,),name='GeneratorCondition')
noiseInput = Input(shape=(randomDim,),name='GeneratorInput')
condition_emb = Embedding(input_dim=10,input_length=10,output_dim=16)(condition)
condition_emb = Flatten()(condition_emb)
model = Concatenate()([condition_emb,noiseInput])
model = Dense(1*1*(160+90))(model)
# model = LeakyReLU(0.1)(model)
model = Reshape((1, 1, (250)))(model)
model = ConvSN2DTranspose(1024, strides=(2,2), kernel_size=(1, 1), padding='valid')(model)
model = BatchNormalization()(model)
model = LeakyReLU(0.1)(model)
model = ConvSN2DTranspose(512, strides=(2,2), kernel_size=(5, 5), padding='valid')(model)
model = BatchNormalization()(model)
model = LeakyReLU(0.1)(model)
model = ConvSN2DTranspose(256, strides=(2,2), kernel_size=(3, 3), padding='same')(model)
model = BatchNormalization()(model)
model = LeakyReLU(0.1)(model)
model = ConvSN2DTranspose(256, strides=(2,2), kernel_size=(4, 4), padding='same')(model)
model = BatchNormalization()(model)
model = LeakyReLU(0.1)(model)
model, beta, gamma = Attention(256)(model)
model = ConvSN2DTranspose(3, strides=(1,1), kernel_size=(4,4),padding='same',activation='tanh',name='G')(model)
g = Model([condition,noiseInput],model, name='Generator')
g.summary()
# display(SVG(model_to_dot(generator,show_layer_names=True,show_shapes=True).create(prog='dot', format='svg')))
return g
generator = build_generator()
# +
# Discriminator
def build_discriminator():
condition = Input(shape=(10,), name='DiscriminatorCondition')
condition_emb = Embedding(input_dim=10,input_length=10,output_dim=32)(condition)
condition_emb = Flatten()(condition_emb)
# print(condition_emb)
inpImage = Input(shape=(28,28,3), name = 'DiscriminatorInput')
d_model = ConvSN2D(filters=128, kernel_size=(4, 4), strides=(2, 2), padding='same')(inpImage)
d_model = BatchNormalization()(d_model)
d_model = LeakyReLU(0.1)(d_model)
d_model, beta, gamma = Attention(128)(d_model)
d_model = ConvSN2D(512, kernel_size=(4, 4), strides=(2, 2), padding='same')(d_model)
d_model = BatchNormalization()(d_model)
d_model = LeakyReLU(0.1)(d_model)
d_model = ConvSN2D(1024, kernel_size=(4, 4), strides=(2, 2), padding='same')(d_model)
d_model = BatchNormalization()(d_model)
d_model = LeakyReLU(0.1)(d_model)
d_model = ConvSN2D(1024, kernel_size=(4, 4), strides=(2, 2), padding='same')(d_model)
d_model = BatchNormalization()(d_model)
d_model = LeakyReLU(0.1)(d_model)
d_model = Flatten()(d_model)
d_model = Dense(320, activation='relu')(d_model)
d_model = Multiply()([d_model,condition_emb])
# d_model = Lambda(lambda x: tf.expand_dims(x,axis=-1))(d_model)
d_model = Dense(1, activation='linear',name='D')(d_model)
d = Model( [condition,inpImage], d_model, name='Discriminator')
d.compile(loss=hinge_loss, optimizer=d_opt, metrics=['accuracy'])
d.summary()
# display(SVG(model_to_dot(discriminator,show_layer_names=True,show_shapes=True).create(prog='dot', format='svg')))
return d
discriminator = build_discriminator()
# +
# def gradient_penalty(real, fake, classes):
# alpha = tf.random_uniform(shape=[self.batch_size, 1, 1, 1], minval=0., maxval=1.)
# interpolated = alpha*real + (1. - alpha)*fake
# logit = discriminator([interpolated, classes], reuse=True)
# grad = tf.gradients(logit, interpolated)[0] # gradient of D(interpolated)
# grad_norm = tf.norm(flatten(grad), axis=1) # l2 norm
# GP = 0
# GP = self.ld * tf.reduce_mean(tf.square(grad_norm - 1.))
# return GP
# +
# discriminator.trainable = False
# g_cond_inp,latent_inp = generator.inputs
# g = generator([g_cond_inp,latent_inp])
# # d_cond_inp = Input((10,))
# # print(x.shape)
# ganOutput = discriminator([g_cond_inp, g])
# gan = Model(inputs=[g_cond_inp,latent_inp], outputs=ganOutput, name='GAN')
# # gan.layers[1].name = "Generator_Network"
# # gan.layers[2].name = "Discriminator_Network"
# gan.compile(loss=hinge_loss, optimizer=g_opt, metrics=['accuracy'])
# gan.summary()
# display(SVG(model_to_dot(model=gan,show_layer_names=True,show_shapes=True).create(prog='dot', format='svg')))
# +
# The generator_model is used when we want to train the generator layers.
# As such, we ensure that the discriminator layers are not trainable.
# Note that once we compile this model, updating .trainable will have no effect within
# it. As such, it won't cause problems if we later set discriminator.trainable = True
# for the discriminator_model, as long as we compile the generator_model first.
for layer in discriminator.layers:
layer.trainable = False
discriminator.trainable = False
generator_input = Input(shape=(90,),name='generator_input')
generator_cond = Input(shape=(10,),name='generator_cond')
discriminator_cond = Input(shape=(10,),name='discriminator_cond')
generator_layers = generator([generator_cond, generator_input])
discriminator_layers_for_generator = discriminator([discriminator_cond,generator_layers])
generator_model = Model(inputs=[discriminator_cond,generator_cond, generator_input],
outputs=[discriminator_layers_for_generator])
# We use the Adam paramaters from Gulrajani et al.
generator_model.compile(optimizer=g_opt,
loss=hinge_loss)
# +
generator_model.summary()
display(SVG(model_to_dot(model=generator_model,show_layer_names=True,show_shapes=True).create(prog='dot', format='svg')))
# +
# Now that the generator_model is compiled, we can make the discriminator
# layers trainable.
for layer in discriminator.layers:
layer.trainable = True
for layer in generator.layers:
layer.trainable = False
discriminator.trainable = True
generator.trainable = False
# The discriminator_model is more complex. It takes both real image samples and random
# noise seeds as input. The noise seed is run through the generator model to get
# generated images. Both real and generated images are then run through the
# discriminator. Although we could concatenate the real and generated images into a
# single tensor, we don't (see model compilation for why).
real_samples = Input(shape=(28,28,3),name='real_samples')
generator_input_for_discriminator = Input(shape=(90,),name='generator_input_for_discriminator')
generator_cond_for_discriminator = Input(shape=(10,),name='generator_cond_for_discriminator')
discriminator_cond_for_discriminator = Input(shape=(10,), name = 'discriminator_cond_for_discriminator')
generated_samples_for_discriminator = generator([generator_cond_for_discriminator,generator_input_for_discriminator])
discriminator_output_from_generator = discriminator([discriminator_cond_for_discriminator,generated_samples_for_discriminator])
discriminator_output_from_real_samples = discriminator([discriminator_cond_for_discriminator,real_samples])
# We also need to generate weighted-averages of real and generated samples,
# to use for the gradient norm penalty.
averaged_samples = RandomWeightedAverage()([real_samples,
generated_samples_for_discriminator])
# We then run these samples through the discriminator as well. Note that we never
# really use the discriminator output for these samples - we're only running them to
# get the gradient norm for the gradient penalty loss.
averaged_samples_out = discriminator([discriminator_cond_for_discriminator,averaged_samples])
# The gradient penalty loss function requires the input averaged samples to get
# gradients. However, Keras loss functions can only have two arguments, y_true and
# y_pred. We get around this by making a partial() of the function with the averaged
# samples here.
partial_gp_loss = partial(gradient_penalty_loss,
averaged_samples=averaged_samples,
gradient_penalty_weight=GRADIENT_PENALTY_WEIGHT)
# Functions need names or Keras will throw an error
partial_gp_loss.__name__ = 'gradient_penalty'
# Keras requires that inputs and outputs have the same number of samples. This is why
# we didn't concatenate the real samples and generated samples before passing them to
# the discriminator: If we had, it would create an output with 2 * BATCH_SIZE samples,
# while the output of the "averaged" samples for gradient penalty
# would have only BATCH_SIZE samples.
# If we don't concatenate the real and generated samples, however, we get three
# outputs: One of the generated samples, one of the real samples, and one of the
# averaged samples, all of size BATCH_SIZE. This works neatly!
discriminator_model = Model(inputs=[generator_cond_for_discriminator \
,discriminator_cond_for_discriminator \
,real_samples \
,generator_input_for_discriminator],
outputs=[discriminator_output_from_real_samples \
,discriminator_output_from_generator \
,averaged_samples_out])
# We use the Adam paramaters from Gulrajani et al. We use the Wasserstein loss for both
# the real and generated samples, and the gradient penalty loss for the averaged samples
discriminator_model.compile(optimizer=d_opt,
loss=[hinge_loss,
hinge_loss,
partial_gp_loss])
# -
discriminator_model.summary()
display(SVG(model_to_dot(model=discriminator_model,show_layer_names=True,show_shapes=True).create(prog='dot', format='svg')))
tensorboard = TensorBoard(
log_dir='log/fmnist_cond_hinge_sn_sa_gp',
histogram_freq=0,
batch_size=32,
write_graph=True,
write_grads=True,
)
tensorboard.set_model(generator_model)
# +
# generator.load_weights('models/cifar10_cond_hinge_sn_sa_gp/dcgan_generator_epoch_15000.h5')
# discriminator.load_weights('models/cifar10_cond_hinge_sn_sa_gp/dcgan_discriminator_epoch_15000.h5')
# -
# +
def train(initepoch=1,epochs=1, batch_size=128, steps = 10):
print ('Epochs:', epochs)
print ('Batch size:', batch_size)
print ('steps per epoch:',steps)
fids = []
fid = 0.0
prev_fid = 100000
counter = 0
real_labels = np.ones((batch_size,1),dtype=np.float32)
fake_labels = -np.ones((batch_size,1),dtype=np.float32)
dummy_labels = np.zeros((batch_size, 1), dtype=np.float32)
for e in tqdm_notebook(range(initepoch, epochs+1)):
# for step in range(steps):
# Get a random set of input noise and images
for l in discriminator.layers: l.trainable = True
discriminator.trainable = True
for l in generator.layers: l.trainable = False
generator.trainable = False
for i in range(TRAINING_RATIO):
z = np.random.uniform(-1, 1, (batch_size, randomDim))
imageBatch, conds = generate_real_samples(dataset=X,conditions=X_cond, n_samples=batch_size)
# generatedImages = generator.predict([conds, z])
d_loss = discriminator_model.train_on_batch([conds, conds, imageBatch, z], [real_labels, fake_labels, dummy_labels])
# d_real = discriminator.train_on_batch([conds, imageBatch], real_labels)
# #Clip discriminator weights
# for l in discriminator.layers:
# weights = l.get_weights()
# weights = [np.clip(w, -0.01,0.01) for w in weights]
# l.set_weights(weights)
# print(dloss)
# Train generator
_, conds = generate_real_samples(dataset=X,conditions=X_cond, n_samples=batch_size)
z = np.random.uniform(-1, 1, (batch_size, randomDim))
for l in discriminator.layers: l.trainable = False
discriminator.trainable = False
for l in generator.layers: l.trainable = True
generator.trainable = True
ganloss = generator_model.train_on_batch([conds, conds, z], real_labels)
# Store loss of most recent batch from this epoch
# dLosses.append(dloss)
# gLosses.append(gloss)
if e == 1 or e % 1000 == 0:
# test_range = np.arange(0, T.shape[0])
# fids = []
# gi = 0
# while gi < T.shape[0]:
z = np.random.uniform(-1, 1, (eval_batch_size, randomDim))
# idxs = test_range[gi:gi+eval_batch_size]
# r_images, r_conds = get_real_samples(T,T_cond,idxs)
g_images = generator.predict([T_cond, z])
r_images = scale_images(T, (299,299))
g_images = scale_images(g_images, (299,299))
fid = calculate_fid(model,r_images,g_images)
tensorboard.on_epoch_end(e, {
"fid":fid \
,"d_loss_0":d_loss[0] \
,"d_loss_1":d_loss[1] \
,"d_loss_2":d_loss[2] \
,"d_loss_3":d_loss[3] \
# ,"D_fake_acc":d_fake[1]
# ,"D_fake_loss":d_fake[0]
# ,"GAN_acc":ganloss[1]
,"gan_loss":ganloss})
plotGeneratedImages(e)
if prev_fid>fid:
saveModels(e)
prev_fid = fid
else:
fid-=0.001
tensorboard.on_epoch_end(e, {
"fid":fid \
,"d_loss_0":d_loss[0] \
,"d_loss_1":d_loss[1] \
,"d_loss_2":d_loss[2] \
,"d_loss_3":d_loss[3] \
# ,"D_fake_acc":d_fake[1]
# ,"D_fake_loss":d_fake[0]
# ,"GAN_acc":ganloss[1]
,"gan_loss":ganloss})
return np.asarray(fids)
# -
fids = train(initepoch=1, epochs=300000, batch_size=batch_size, steps=1)
# noise = randn(randomDim*batchSize)
# noise = noise.reshape((batchSize,randomDim))
# imageBatch , conds= generate_real_samples(dataset=X, conditions=X_cond,n_samples=64)
# imagesCount = 64
# generatorModel.load_weights('models/cifar10_cond_hingegan_sn_wt_clip/dcgan_generator_epoch_353000.h5')
# images2 = generatorModel.predict([conds,noise])
# images1 = scale_images(images1, (140,140))
# images2 = scale_images(images2, (140,140))
# fid = calculate_fid(model, images1, images2)
d_m = Model(discriminator.inputs,discriminator.get_layer(index=6).output)
d_m.summary()
# g_m = Model(generatorModel.inputs,generatorModel.get_layer(index=-2).output)
# g_m.summary()
# +
# for l in discriminator.layers:
# weights = l.get_weights()
# weights = [np.clip(w, -0.01,0.01) for w in weights]
# l.set_weights(weights)
# for l in discriminator.layers:
# weights = l.get_weights()
# print(weights)
# print(l.name,':',np.shape(weights))
# +
# for w in discriminator.layers[4].get_weights():
# print(np.min(w),np.max(w))
# +
# Layer (type) Output Shape Param # Connected to
# ==================================================================================================
# DiscriminatorInput (InputLayer) (None, 32, 32, 3) 0
# __________________________________________________________________________________________________
# conv_s_n2d_1 (ConvSN2D) (None, 16, 16, 256) 12800 DiscriminatorInput[0][0]
# __________________________________________________________________________________________________
# batch_normalization_5 (BatchNor (None, 16, 16, 256) 1024 conv_s_n2d_1[0][0]
# __________________________________________________________________________________________________
# leaky_re_lu_5 (LeakyReLU) (None, 16, 16, 256) 0 batch_normalization_5[0][0]
# __________________________________________________________________________________________________
# attention_2 (Attention) [(None, 16, 16, 256) 82241 leaky_re_lu_5[0][0]
# __________________________________________________________________________________________________
# conv_s_n2d_2 (ConvSN2D) (None, 8, 8, 512) 2098176 attention_2[0][0]
# __________________________________________________________________________________________________
# batch_normalization_6 (BatchNor (None, 8, 8, 512) 2048 conv_s_n2d_2[0][0]
# __________________________________________________________________________________________________
# leaky_re_lu_6 (LeakyReLU) (None, 8, 8, 512) 0 batch_normalization_6[0][0]
# __________________________________________________________________________________________________
# conv_s_n2d_3 (ConvSN2D) (None, 4, 4, 1024) 8390656 leaky_re_lu_6[0][0]
# __________________________________________________________________________________________________
# batch_normalization_7 (BatchNor (None, 4, 4, 1024) 4096 conv_s_n2d_3[0][0]
# __________________________________________________________________________________________________
# leaky_re_lu_7 (LeakyReLU) (None, 4, 4, 1024) 0 batch_normalization_7[0][0]
# __________________________________________________________________________________________________
# conv_s_n2d_4 (ConvSN2D) (None, 2, 2, 1024) 16779264 leaky_re_lu_7[0][0]
# __________________________________________________________________________________________________
# batch_normalization_8 (BatchNor (None, 2, 2, 1024) 4096 conv_s_n2d_4[0][0]
# __________________________________________________________________________________________________
# leaky_re_lu_8 (LeakyReLU) (None, 2, 2, 1024) 0 batch_normalization_8[0][0]
# __________________________________________________________________________________________________
# flatten_1 (Flatten) (None, 4096) 0 leaky_re_lu_8[0][0]
# __________________________________________________________________________________________________
# DiscriminatorCondition (InputLa (None, 10) 0
# __________________________________________________________________________________________________
# concatenate_2 (Concatenate) (None, 4106) 0 flatten_1[0][0]
# DiscriminatorCondition[0][0]
# __________________________________________________________________________________________________
# D (Dense) (None, 1) 4107 concatenate_2[0][0]
# ==================================================================================================
# Total params: 27,378,508
# Trainable params: 27,370,060
# Non-trainable params: 8,448
# +
z = np.random.uniform(-1, 1, (100, randomDim))
f_conds = to_categorical(np.zeros(100)+9,num_classes=10)
generatedImages = generator.predict([f_conds,z])
realImages, conds = generate_real_samples(dataset=X,conditions=X_cond, n_samples=100)
df, da, dsma, dg = d_m.predict([f_conds,generatedImages])
# gf, ga, gsma, gg = g_m.predict([f_conds,z])
# -
plot_images(generatedImages)
dg.min(),dg.max()
np.unique(gg).shape
pd.DataFrame(fids)
_, conds = generate_real_samples(dataset=X,conditions=X_cond, n_samples=batch_size)
for i in range(10):
print(np.argmax(conds[i]))
plt.imshow(_[i])
plt.show()
np.repeat(a=np.arange(10).reshape(1,-1),repeats=10, axis=0).reshape((-1))
np.arange(10).reshape(1,-1)
# +
# https://arxiv.org/pdf/1611.06355.pdf - invertible
# https://arxiv.org/pdf/1411.1784.pdf - cGANs
# https://arxiv.org/pdf/1802.05957.pdf - SN GANs
# https://www.quora.com/How-does-Conditional-Batch-normalization-work-and-how-is-it-different-from-regular-Batch-normalization
# -
imagesCount = 100
z = np.repeat(np.random.uniform(-1, 1, (10, randomDim)),10,axis=0)
labels = np.tile(np.eye(10), [10, 1])
# labels = np.repeat(a=np.arange(10).reshape(1,-1),repeats=10, axis=0).reshape((-1))
images1, _ = generate_real_samples(dataset=X,conditions=X_cond,n_samples=imagesCount) # Here X is test data of CIFAR10
images2 = generator.predict([labels, z])
images1 = scale_images(images1, (140,140))
images2 = scale_images(images2, (140,140))
# fid = calculate_fid(model, images1, images2)
# fids.append(fid)
images1.min(),images1.max()
images2.min(),images2.max()
from PIL import Image
def tile_images(image_stack):
"""Given a stacked tensor of images, reshapes them into a horizontal tiling for
display."""
print(image_stack.shape)
assert len(image_stack.shape) == 4
image_list = [image_stack[i, :, :] for i in range(image_stack.shape[0])]
print(np.shape(image_list))
image_list = np.concatenate(image_list, axis=1)
print(image_list.shape)
return image_list
def generate_images(generator_model):
"""Feeds random seeds into the generator and tiles and saves the output to a PNG
file."""
test_image_stack = generator_model.predict([np.tile(np.eye(10),reps=[10,1]),np.repeat(np.random.uniform(low=-1,high=1,size=(10, 90)),repeats=10,axis=0)])
test_image_stack = (test_image_stack * 127.5) + 127.5
# test_image_stack = np.squeeze(np.round(test_image_stack).astype(np.uint8))
tiled_output = tile_images(test_image_stack)
print(tiled_output.shape)
tiled_output = plt.imshow(tiled_output) # L specifies greyscale
display(tiled_output)
# outfile = os.path.join(output_dir, 'epoch_{}.png'.format(epoch))
# tiled_output.save(outfile)
generate_images(generator)
np.tile(np.eye(10),reps=[10,1])
|
Keras-cSAWGAN/Cond_HingeGAN_SpectralNorm_SelfAttention_GP-Fmnist_Proj.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mbrady4/DS-Unit-1-Sprint-3-Data-Storytelling/blob/master/module3-make-explanatory-visualizations/LS_DS_223_Make_explanatory_visualizations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="-8-trVo__vRE"
# _Lambda School Data Science_
#
# # Make explanatory visualizations
#
#
#
#
# Tody we will reproduce this [example by FiveThirtyEight:](https://fivethirtyeight.com/features/al-gores-new-movie-exposes-the-big-flaw-in-online-movie-ratings/)
# + colab_type="code" id="ya_w5WORGs-n" colab={"base_uri": "https://localhost:8080/", "height": 355} outputId="ccd23002-0688-4591-c61d-1372167986ec"
from IPython.display import display, Image
url = 'https://fivethirtyeight.com/wp-content/uploads/2017/09/mehtahickey-inconvenient-0830-1.png'
example = Image(url=url, width=400)
display(example)
# + [markdown] colab_type="text" id="HP4DALiRG3sC"
# Using this data: https://github.com/fivethirtyeight/data/tree/master/inconvenient-sequel
# + [markdown] colab_type="text" id="HioPkYtUG03B"
# Objectives
# - add emphasis and annotations to transform visualizations from exploratory to explanatory
# - remove clutter from visualizations
#
# Links
# - [Strong Titles Are The Biggest Bang for Your Buck](http://stephanieevergreen.com/strong-titles/)
# - [Remove to improve (the data-ink ratio)](https://www.darkhorseanalytics.com/blog/data-looks-better-naked)
# - [How to Generate FiveThirtyEight Graphs in Python](https://www.dataquest.io/blog/making-538-plots/)
# + [markdown] colab_type="text" id="0w_iMnQ6-VoQ"
# ## Make prototypes
#
# This helps us understand the problem
# + colab_type="code" id="5uz0eEaEN-GO" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="afb8e3cd-7097-4b66-e1a0-0f8bed9af17f"
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
plt.style.use('fivethirtyeight')
fake = pd.Series([38, 3, 2, 1, 2, 4, 6, 5, 5, 33],
index=range(1,11))
fake.plot.bar(color='C1', width=0.9);
# + colab_type="code" id="KZ0VLOV8OyRr" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="87ee135a-e70b-438f-d424-6b3735cddc3b"
fake2 = pd.Series(
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
2, 2, 2,
3, 3, 3,
4, 4,
5, 5, 5,
6, 6, 6, 6,
7, 7, 7, 7, 7,
8, 8, 8, 8,
9, 9, 9, 9,
10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10])
fake2.value_counts().sort_index().plot.bar(color='C1', width=0.9);
# + [markdown] colab_type="text" id="mZb3UZWO-q05"
# ## Annotate with text
# + colab_type="code" id="f6U1vswr_uWp" colab={"base_uri": "https://localhost:8080/", "height": 350} outputId="3e916416-c17e-4c80-ae2f-55cf8ec04c62"
plt.style.use('fivethirtyeight')
fig = plt.figure()
fig.patch.set_facecolor('white')
fake = pd.Series([38, 3, 2, 1, 2, 4, 6, 5, 5, 33],
index=range(1,11))
ax = fake.plot.bar(color='#ef7030', width=0.9)
#plt.title("An Inconvenient Sequal: Trust To Power' is Divisive")
#plt.xlabel('Percent of Total Votes')
#plt.ylabel('Rating')
ax.set(yticks=range(0,50,10),
facecolor='white')
plt.ylabel('Percent of Total Votes', fontsize=10, fontweight='bold')
plt.xlabel('Rating', fontsize=10, fontweight='bold')
ax.tick_params(labelrotation=0,
labelsize=10)
ax.text(x=-1.75,
y=44.5,
s="An Inconvenient Sequal: Trust To Power' is Divisive",
fontsize=14,
fontweight='bold')
ax.text(x=-1.75,
y=42,
s="IMBD ratings for the film as of Aug. 29",
fontsize=12,
fontweight='normal');
ax.text(x=8.25,
y=-8,
s="Source: IBMD",
fontsize=9,
fontweight='normal');
ax.text(x=-1.75,
y=-8,
s="Lambda School",
fontsize=9,
fontweight='normal');
# + [markdown] colab_type="text" id="x8jRZkpB_MJ6"
# ## Reproduce with real data
# + colab_type="code" id="3SOHJckDUPI8" colab={}
df = pd.read_csv('https://raw.githubusercontent.com/fivethirtyeight/data/master/inconvenient-sequel/ratings.csv')
# + colab_type="code" id="cDltXxhC_yG-" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="23622d31-af1e-4e24-e16c-474bf2664a29"
df.shape
# + id="k8VzKmzIG9is" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 329} outputId="1944e841-ef25-4475-b561-67052cbe72f5"
pd.set_option('display.max_columns', 500)
df.head()
# + id="oUGVU3JYJ1cI" colab_type="code" colab={}
df['timestamp'] = pd.to_datetime(df['timestamp'])
# + id="qeXkeCxYJn8m" colab_type="code" colab={}
df.set_index('timestamp', inplace=True)
# + id="fz4Bt8PwJyAc" colab_type="code" colab={}
last_day = df['2017-08-29']
# + id="0eLaSR8UMFx9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="f62e806a-35b3-4a73-bd32-f8e39632dab4"
last_day[ last_day['category'] == 'IMDb users'].respondents.plot()
# + id="aTJQW0qSKF_n" colab_type="code" colab={}
final = last_day.tail(1)
# + id="n0Fce3XPM5D7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="f417ef61-9df1-4da3-c79a-dcfa39ef30d2"
final
# + id="1B-ZhiSfNEaj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="e9031b5b-4485-4960-f559-98a53d3f7a90"
pct_columns = ([
'1_pct', '2_pct', '3_pct', '4_pct', '5_pct', '6_pct', '7_pct', '8_pct', '9_pct', '10_pct'
])
data = final[pct_columns].T
data
# + id="pRjl_tgWObU3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="87c68c2d-da15-45af-a730-fcb009051024"
plt.style.use('fivethirtyeight')
fig = plt.figure()
fig.patch.set_facecolor('white')
fake = pd.Series([38, 3, 2, 1, 2, 4, 6, 5, 5, 33],
index=range(1,11))
ax = data.plot.bar(color='#ef7030', width=0.9)
#plt.title("An Inconvenient Sequal: Trust To Power' is Divisive")
#plt.xlabel('Percent of Total Votes')
#plt.ylabel('Rating')
ax.set(yticks=range(0,50,10),
facecolor='white')
plt.ylabel('Percent of Total Votes', fontsize=10, fontweight='bold')
plt.xlabel('Rating', fontsize=10, fontweight='bold')
ax.tick_params(labelrotation=0,
labelsize=10)
ax.text(x=-1.75,
y=44.5,
s="An Inconvenient Sequal: Trust To Power' is Divisive",
fontsize=14,
fontweight='bold')
ax.text(x=-1.75,
y=42,
s="IMBD ratings for the film as of Aug. 29",
fontsize=12,
fontweight='normal');
ax.text(x=8.25,
y=-8,
s="Source: IBMD",
fontsize=9,
fontweight='normal');
ax.text(x=-1.75,
y=-8,
s="Lambda School",
fontsize=9,
fontweight='normal');
# + [markdown] colab_type="text" id="NMEswXWh9mqw"
# # ASSIGNMENT
#
# Replicate the lesson code. I recommend that you [do not copy-paste](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit).
#
# # STRETCH OPTIONS
#
# #### Reproduce another example from [FiveThityEight's shared data repository](https://data.fivethirtyeight.com/).
#
# For example:
# - [thanksgiving-2015](https://fivethirtyeight.com/features/heres-what-your-part-of-america-eats-on-thanksgiving/) (try the [`altair`](https://altair-viz.github.io/gallery/index.html#maps) library)
# - [candy-power-ranking](https://fivethirtyeight.com/features/the-ultimate-halloween-candy-power-ranking/) (try the [`statsmodels`](https://www.statsmodels.org/stable/index.html) library)
# - or another example of your choice!
#
# #### Make more charts!
#
# Choose a chart you want to make, from [Visual Vocabulary - Vega Edition](http://ft.com/vocabulary).
#
# Find the chart in an example gallery of a Python data visualization library:
# - [Seaborn](http://seaborn.pydata.org/examples/index.html)
# - [Altair](https://altair-viz.github.io/gallery/index.html)
# - [Matplotlib](https://matplotlib.org/gallery.html)
# - [Pandas](https://pandas.pydata.org/pandas-docs/stable/visualization.html)
#
# Reproduce the chart. [Optionally, try the "Ben Franklin Method."](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit) If you want, experiment and make changes.
#
# Take notes. Consider sharing your work with your cohort!
#
#
#
#
#
#
#
#
|
module3-make-explanatory-visualizations/LS_DS_223_Make_explanatory_visualizations.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,py
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:core_acc] *
# language: python
# name: conda-env-core_acc-py
# ---
# # Relationships using expression distance
#
# This notebook is performing the same analysis as seen in [all_gene_relationships.ipynb](archive/all_gene_relationships.ipynb), where we are examining who is related to who. Previously we started with an accessory gene and asked: is the highest correlated gene another accessory gene or a core gene? For this analysis, we are starting with the most stable core genes and asking the same question: is the highest correlated gene core or accessory?
#
# Note: We do not have the genome location metric here because this would require a significant effort to figure out how to modify the existing code to only focus on a subset of genes.
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
import random
import scipy
import pandas as pd
import numpy as np
import textwrap
import seaborn as sns
import matplotlib.pyplot as plt
from statsmodels.stats.multitest import multipletests
from plotnine import (
ggplot,
labs,
geom_hline,
geom_bar,
geom_errorbar,
positions,
aes,
ggsave,
theme_bw,
theme,
theme_seaborn,
facet_wrap,
scale_fill_manual,
scale_x_discrete,
xlim,
ylim,
guides,
guide_legend,
element_blank,
element_text,
element_rect,
element_line,
coords,
)
from scripts import utils, paths, gene_relationships, annotations
random.seed(1)
# +
# User params
offset_to_bin = 10
use_operon = True
sum_increment_to_use = 1
# Output filename
pao1_figure_filename = (
"PAO1_stablility_expression_relationships_operon_corrected_spell.svg"
)
pa14_figure_filename = (
"PA14_stability_expression_relationships_operon_corrected_spell.svg"
)
# -
# ### Import gene ids
# +
# Import correlation matrix to get gene ids
pao1_corr_filename = paths.PAO1_CORR_LOG_SPELL
pa14_corr_filename = paths.PA14_CORR_LOG_SPELL
pao1_corr = pd.read_csv(pao1_corr_filename, sep="\t", index_col=0, header=0)
pa14_corr = pd.read_csv(pa14_corr_filename, sep="\t", index_col=0, header=0)
# -
# Make a dataframe with gene ids
pao1_membership = pd.DataFrame(data=[], index=pao1_corr.index)
print(pao1_membership.shape)
pao1_membership.head()
pa14_membership = pd.DataFrame(data=[], index=pa14_corr.index)
print(pa14_membership.shape)
pa14_membership.head()
# ### Import and format operon data
pao1_operon_filename = paths.PAO1_OPERON
pa14_operon_filename = paths.PA14_OPERON
pao1_operon = annotations.load_format_operons(pao1_operon_filename)
pa14_operon = annotations.load_format_operons(pa14_operon_filename)
print(pao1_operon.shape)
pao1_operon.head()
if use_operon:
pao1_operon_expression_to_use = pao1_operon
pa14_operon_expression_to_use = pa14_operon
else:
pao1_operon_expression_to_use = None
pa14_operon_expression_to_use = None
# ### Map core/accessory labels to genes
# Read in expression data
pao1_expression_filename = paths.PAO1_COMPENDIUM
pa14_expression_filename = paths.PA14_COMPENDIUM
pao1_annot_filename = paths.GENE_PAO1_ANNOT
pa14_annot_filename = paths.GENE_PA14_ANNOT
(
pao1_arr,
pa14_arr,
pao1_core,
pao1_acc,
pa14_core,
pa14_acc,
) = annotations.map_core_acc_annot(
pao1_membership,
pa14_membership,
pao1_expression_filename,
pa14_expression_filename,
pao1_annot_filename,
pa14_annot_filename,
)
print(pao1_arr.shape)
pao1_arr.head()
pao1_arr.tail()
print(pa14_arr.shape)
pa14_arr.head()
pa14_arr.tail()
# ## Find relationships using expression distance
# Correlation matrix files
pao1_corr_filename = paths.PAO1_CORR_LOG_SPELL
pa14_corr_filename = paths.PA14_CORR_LOG_SPELL
# Load correlation data
pao1_corr = pd.read_csv(pao1_corr_filename, sep="\t", index_col=0, header=0)
pa14_corr = pd.read_csv(pa14_corr_filename, sep="\t", index_col=0, header=0)
# +
# Load transcriptional similarity df
# These are the subset of genes that we will consider
pao1_similarity_scores_filename = (
"../3_core_core_analysis/pao1_core_similarity_associations_final_spell.tsv"
)
pa14_similarity_scores_filename = (
"../3_core_core_analysis/pa14_core_similarity_associations_final_spell.tsv"
)
pao1_similarity_scores = pd.read_csv(
pao1_similarity_scores_filename, sep="\t", header=0, index_col=0
)
pa14_similarity_scores = pd.read_csv(
pa14_similarity_scores_filename, sep="\t", header=0, index_col=0
)
# +
# Get most and least stable core genes
pao1_most_stable_genes = list(
pao1_similarity_scores[pao1_similarity_scores["label"] == "most stable"].index
)
pao1_least_stable_genes = list(
pao1_similarity_scores[pao1_similarity_scores["label"] == "least stable"].index
)
pa14_most_stable_genes = list(
pa14_similarity_scores[pa14_similarity_scores["label"] == "most stable"].index
)
pa14_least_stable_genes = list(
pa14_similarity_scores[pa14_similarity_scores["label"] == "least stable"].index
)
# -
# %%time
expression_dist_counts_pao1_most = (
gene_relationships.get_relationship_in_expression_space(
pao1_corr,
pao1_most_stable_genes,
pao1_arr,
offset_to_bin,
pao1_operon_expression_to_use,
sum_increment_to_use,
)
)
# %%time
expression_dist_counts_pao1_least = (
gene_relationships.get_relationship_in_expression_space(
pao1_corr,
pao1_least_stable_genes,
pao1_arr,
offset_to_bin,
pao1_operon_expression_to_use,
sum_increment_to_use,
)
)
# %%time
expression_dist_counts_pa14_most = (
gene_relationships.get_relationship_in_expression_space(
pa14_corr,
pa14_most_stable_genes,
pa14_arr,
offset_to_bin,
pa14_operon_expression_to_use,
sum_increment_to_use,
)
)
# %%time
expression_dist_counts_pa14_least = (
gene_relationships.get_relationship_in_expression_space(
pa14_corr,
pa14_least_stable_genes,
pa14_arr,
offset_to_bin,
pa14_operon_expression_to_use,
sum_increment_to_use,
)
)
expression_dist_counts_pao1_most.head()
expression_dist_counts_pao1_least.head()
expression_dist_counts_pa14_most.head()
expression_dist_counts_pa14_least.head()
# ### Format data for plotting
#
# Here we will calculate the proportion of gene types per offset and then normalize by the proportion of core and accessory genes. This will return an oddsratio type value where if the value is >1 than the proportion of genes of that type are more than expected.
# Calculate the percentages per offset
expression_dist_counts_pao1_most["percent"] = expression_dist_counts_pao1_most[
"total"
] / len(pao1_most_stable_genes)
expression_dist_counts_pao1_least["percent"] = expression_dist_counts_pao1_least[
"total"
] / len(pao1_least_stable_genes)
expression_dist_counts_pa14_most["percent"] = expression_dist_counts_pa14_most[
"total"
] / len(pa14_most_stable_genes)
expression_dist_counts_pa14_least["percent"] = expression_dist_counts_pa14_least[
"total"
] / len(pa14_least_stable_genes)
# Baseline/expected proportions for PAO1
pao1_total = len(pao1_core) + len(pao1_acc)
pao1_acc_expected = len(pao1_acc) / pao1_total
pao1_core_expected = len(pao1_core) / pao1_total
print("total pao1 genes", pao1_total)
print("pao1 acc baseline", pao1_acc_expected)
print("pao1 core baseline", pao1_core_expected)
# Baseline/expected proportions for PA14
pa14_total = len(pa14_core) + len(pa14_acc)
pa14_acc_expected = len(pa14_acc) / pa14_total
pa14_core_expected = len(pa14_core) / pa14_total
print("total pa14 genes", pa14_total)
print("pa14 acc baseline", pa14_acc_expected)
print("pa14 core baseline", pa14_core_expected)
# +
# Normalize by baseline PAO1 most stable
pao1_acc_most_ids = expression_dist_counts_pao1_most.loc[
expression_dist_counts_pao1_most["gene type"] == "acc"
].index
pao1_core_most_ids = expression_dist_counts_pao1_most.loc[
expression_dist_counts_pao1_most["gene type"] == "core"
].index
expression_dist_counts_pao1_most.loc[pao1_acc_most_ids, "normalized"] = (
expression_dist_counts_pao1_most.loc[pao1_acc_most_ids, "percent"]
/ pao1_acc_expected
)
expression_dist_counts_pao1_most.loc[pao1_core_most_ids, "normalized"] = (
expression_dist_counts_pao1_most.loc[pao1_core_most_ids, "percent"]
/ pao1_core_expected
)
# +
# Normalize by baseline PAO1 least stable
pao1_acc_least_ids = expression_dist_counts_pao1_least.loc[
expression_dist_counts_pao1_least["gene type"] == "acc"
].index
pao1_core_least_ids = expression_dist_counts_pao1_least.loc[
expression_dist_counts_pao1_least["gene type"] == "core"
].index
expression_dist_counts_pao1_least.loc[pao1_acc_least_ids, "normalized"] = (
expression_dist_counts_pao1_least.loc[pao1_acc_least_ids, "percent"]
/ pao1_acc_expected
)
expression_dist_counts_pao1_least.loc[pao1_core_least_ids, "normalized"] = (
expression_dist_counts_pao1_least.loc[pao1_core_least_ids, "percent"]
/ pao1_core_expected
)
# +
# Normalize by baseline PA14 most stable
pa14_acc_most_ids = expression_dist_counts_pa14_most.loc[
expression_dist_counts_pa14_most["gene type"] == "acc"
].index
pa14_core_most_ids = expression_dist_counts_pao1_most.loc[
expression_dist_counts_pa14_most["gene type"] == "core"
].index
expression_dist_counts_pa14_most.loc[pa14_acc_most_ids, "normalized"] = (
expression_dist_counts_pa14_most.loc[pa14_acc_most_ids, "percent"]
/ pa14_acc_expected
)
expression_dist_counts_pa14_most.loc[pa14_core_most_ids, "normalized"] = (
expression_dist_counts_pa14_most.loc[pa14_core_most_ids, "percent"]
/ pa14_core_expected
)
# +
# Normalize by baseline PA14 least stable
pa14_acc_least_ids = expression_dist_counts_pa14_least.loc[
expression_dist_counts_pa14_least["gene type"] == "acc"
].index
pa14_core_least_ids = expression_dist_counts_pa14_least.loc[
expression_dist_counts_pa14_least["gene type"] == "core"
].index
expression_dist_counts_pa14_least.loc[pa14_acc_least_ids, "normalized"] = (
expression_dist_counts_pa14_least.loc[pa14_acc_least_ids, "percent"]
/ pa14_acc_expected
)
expression_dist_counts_pa14_least.loc[pa14_core_least_ids, "normalized"] = (
expression_dist_counts_pa14_least.loc[pa14_core_least_ids, "percent"]
/ pa14_core_expected
)
# -
# Combine PAO1 dataframes
expression_dist_counts_pao1_most.loc[pao1_acc_most_ids, "label"] = "most stable acc"
expression_dist_counts_pao1_most.loc[pao1_core_most_ids, "label"] = "most stable core"
expression_dist_counts_pao1_least.loc[pao1_acc_least_ids, "label"] = "least stable acc"
expression_dist_counts_pao1_least.loc[
pao1_core_least_ids, "label"
] = "least stable core"
# Combine PA14 dataframes
expression_dist_counts_pa14_most.loc[pa14_acc_most_ids, "label"] = "most stable acc"
expression_dist_counts_pa14_most.loc[pa14_core_most_ids, "label"] = "most stable core"
expression_dist_counts_pa14_least.loc[pa14_acc_least_ids, "label"] = "least stable acc"
expression_dist_counts_pa14_least.loc[
pa14_core_least_ids, "label"
] = "least stable core"
# ### Add confidence interval
# +
# Import confidence interval data
pao1_most_ci = pd.read_csv("pao1_most_ci.tsv", sep="\t", index_col=0, header=0)
pao1_least_ci = pd.read_csv("pao1_least_ci.tsv", sep="\t", index_col=0, header=0)
pa14_most_ci = pd.read_csv("pa14_most_ci.tsv", sep="\t", index_col=0, header=0)
pa14_least_ci = pd.read_csv("pa14_least_ci.tsv", sep="\t", index_col=0, header=0)
# -
expression_dist_counts_pao1_most = expression_dist_counts_pao1_most.merge(
pao1_most_ci[["ymin", "ymax"]], left_index=True, right_index=True
)
expression_dist_counts_pao1_least = expression_dist_counts_pao1_least.merge(
pao1_least_ci[["ymin", "ymax"]], left_index=True, right_index=True
)
expression_dist_counts_pa14_most = expression_dist_counts_pa14_most.merge(
pa14_most_ci[["ymin", "ymax"]], left_index=True, right_index=True
)
expression_dist_counts_pa14_least = expression_dist_counts_pa14_least.merge(
pa14_least_ci[["ymin", "ymax"]], left_index=True, right_index=True
)
expression_dist_counts_pao1_all = pd.concat(
[expression_dist_counts_pao1_most, expression_dist_counts_pao1_least]
)
expression_dist_counts_pao1_all
expression_dist_counts_pa14_all = pd.concat(
[expression_dist_counts_pa14_most, expression_dist_counts_pa14_least]
)
expression_dist_counts_pa14_all
# ### Plot
# +
pao1_subset = expression_dist_counts_pao1_all[
(expression_dist_counts_pao1_all["gene type"] == "acc")
]
pao1_subset["offset"] = list(pao1_subset["offset"].astype("str"))
pao1_subset["offset"].replace("+10", ">10", inplace=True)
x_ticks = [">10", "1", "2", "3", "4", "5", "6", "7", "8", "9", "10"]
fig_pao1 = (
ggplot(pao1_subset, aes(x="offset", y="normalized", fill="label"))
+ geom_bar(stat="identity", position="dodge", width=0.8)
+ geom_errorbar(
pao1_subset,
aes(x="offset", ymin="ymin", ymax="ymax"),
position=positions.position_dodge(0.8),
color="black",
)
+ geom_hline(aes(yintercept=1.0), linetype="dashed")
+ labs(
x="Rank co-expression",
y="observed/expected",
title="Stability vs accessory gene relationship (PAO1)",
)
+ theme_seaborn("white")
+ theme(
panel_grid_major_x=element_line(color="lightgrey"),
panel_grid_major_y=element_line(color="lightgrey"),
axis_line=element_line(color="grey"),
legend_title=element_blank(),
legend_text=element_text(family="sans-serif", size=12),
plot_title=element_text(family="sans-serif", size=16),
axis_text=element_text(family="sans-serif", size=12),
axis_title=element_text(family="sans-serif", size=14),
)
+ scale_fill_manual(
values=["#a6aed0ff", "#4e1c80"],
labels=[
"least stable",
"most stable",
],
)
+ scale_x_discrete(limits=x_ticks, labels=x_ticks)
+ ylim(0, 3.5)
)
print(fig_pao1)
# +
pa14_subset = expression_dist_counts_pa14_all[
(expression_dist_counts_pa14_all["gene type"] == "acc")
]
pa14_subset["offset"] = list(pa14_subset["offset"].astype("str"))
pa14_subset["offset"].replace("+10", ">10", inplace=True)
fig_pa14 = (
ggplot(pa14_subset, aes(x="offset", y="normalized", fill="label"))
+ geom_bar(stat="identity", position="dodge", width=0.8)
+ geom_errorbar(
pa14_subset,
aes(x="offset", ymin="ymin", ymax="ymax"),
position=positions.position_dodge(0.8),
color="black",
)
+ geom_hline(aes(yintercept=1.0), linetype="dashed")
+ labs(
x="Rank co-expression",
y="observed/expected",
title="Stability vs accessory gene relationship (PA14)",
)
+ theme_seaborn("white")
+ theme(
panel_grid_major_x=element_line(color="lightgrey"),
panel_grid_major_y=element_line(color="lightgrey"),
axis_line=element_line(color="grey"),
legend_title=element_blank(),
legend_text=element_text(family="sans-serif", size=12),
plot_title=element_text(family="sans-serif", size=16),
axis_text=element_text(family="sans-serif", size=12),
axis_title=element_text(family="sans-serif", size=14),
)
+ scale_fill_manual(
values=["#a6aed0ff", "#4e1c80"],
labels=[
"least stable",
"most stable",
],
)
+ scale_x_discrete(limits=x_ticks, labels=x_ticks)
+ ylim(0, 3.5)
)
print(fig_pa14)
# +
# Calculate statistical test between the distribution of the top 10 co-expressed
# genes related to the least stable vs the most stable core genes
# Test: mean number of co-expressed accessory genes in least stable group vs mean number of
# co-expressed accessory genes in most stable group
# (compare dark blue and light blue bars)
pao1_least_df = pao1_subset[pao1_subset["label"] == "least stable acc"]
pao1_least_df = pao1_least_df[pao1_least_df.offset != "+10"]
pao1_least_vals = pao1_least_df["normalized"].values
pao1_most_df = pao1_subset[pao1_subset["label"] == "most stable acc"]
pao1_most_df = pao1_most_df[pao1_most_df.offset != "+10"]
pao1_most_vals = pao1_most_df["normalized"].values
# Independent t-test
# Test the null hypothesis such that the means of two populations are equal
(pao1_stats, pao1_pvalue) = scipy.stats.ttest_ind(pao1_least_vals, pao1_most_vals)
print(pao1_stats, pao1_pvalue)
# Non-parametric test
# nonparametric test of the null hypothesis that, for randomly selected values X and Y from two populations,
# the probability of X being greater than Y is equal to the probability of Y being greater than X.
(pao1_stats, pao1_pvalue) = scipy.stats.mannwhitneyu(pao1_least_vals, pao1_most_vals)
print(pao1_stats, pao1_pvalue)
# +
pa14_least_df = pa14_subset[pa14_subset["label"] == "least stable acc"]
pa14_least_df = pa14_least_df[pa14_least_df.offset != "+10"]
pa14_least_vals = pa14_least_df["normalized"].values
pa14_most_df = pa14_subset[pa14_subset["label"] == "most stable acc"]
pa14_most_df = pa14_most_df[pa14_most_df.offset != "+10"]
pa14_most_vals = pa14_most_df["normalized"].values
# Independent t-test
(pa14_stats, pa14_pvalue) = scipy.stats.ttest_ind(pa14_least_vals, pa14_most_vals)
print(pa14_stats, pa14_pvalue)
# Non-parametric test
(pa14_stats, pa14_pvalue) = scipy.stats.mannwhitneyu(pa14_least_vals, pa14_most_vals)
print(pa14_stats, pa14_pvalue)
# -
# Based on the bar plots we can be confident in our trend (as seen by the confidence intervals) that least stable genes are more co-expressed with accessory genes compared to most stable genes. This difference between least and most stable genes is further quantified by the t-test comparing the distribution of accessory genes related least vs most genes.
ggsave(plot=fig_pao1, filename=pao1_figure_filename, device="svg", dpi=300)
ggsave(plot=fig_pa14, filename=pa14_figure_filename, device="svg", dpi=300)
# **Takeaway:**
#
# * Least stable core genes have more accessory gene neighbors compared to most stable core genes
# * Previous evidence found that insertion sequences (type of accessory gene) can change the expression of existing genes once it is integrated into the genome. So perhaps these least stable core genes transcriptional behavior is modified by the accessory genes.
|
5_core_acc_analysis/1_stable_gene_relationships.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="iKEPfQAGW_go"
# !rm -rf *
# + colab={"base_uri": "https://localhost:8080/"} id="Rx7hgw1QR6GZ" outputId="16a1f804-e1ee-4b75-cfb4-81f003897ce0"
from google_drive_downloader import GoogleDriveDownloader as gdd
gdd.download_file_from_google_drive(file_id='1-LxbG3N1Eem7wrBYALvoSfMcq_1berKt',dest_path='./data.zip')
gdd.download_file_from_google_drive(file_id='1AqfOtvDZuDyvFo1eUpmcMqfUsDApq5Nz',dest_path='./models.zip')
# download pretrained models
# + colab={"base_uri": "https://localhost:8080/"} id="V3afdUvMRxhb" outputId="7d2b23a8-c4ac-4465-e7af-919d5a3a3c85"
# !git clone https://github.com/ZhengJun-AI/Spinal-Disease-Detection.git
# !unzip data.zip
# !unzip models.zip
# + id="gY_FZw9QSHLI" colab={"base_uri": "https://localhost:8080/"} outputId="f6414d40-17c2-474f-bbd0-a5dac99cea8c"
# !python ./Spinal-Disease-Detection/demo/demo.py -n ./label-all-npy -l ./Spinal-Disease-Detection/demo/new_data-bbox-cls.json -i 123 --show_label
# use -i to choose different data
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="45VZbHHfX1T4" outputId="49df2de3-7abd-4d98-c667-f96f6bf6be7f"
import matplotlib.pyplot as plt
import os
img_root = './result'
imgs = os.listdir(img_root)
img = plt.imread(os.path.join(img_root, imgs[-1]))
plt.imshow(img)
plt.show()
# + id="8BMlDRM4Y4Zr"
|
demo/demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TinyFPGAで音源を作ってみる(計算メモ)
#
# [TinyFPGA BX](https://tinyfpga.com/bx/guide.html)で音源を作ってみる。
# そのために必要だった計算等のメモ
# 原理は至極単純でFPGAの組込みRAMに1周期分の波形データを初期値として保存しておいて音程(pitch)に対応したクロック周期(マスタクロックの分周で作る)で逐次データ読みだしてPWM変調出力してオーディオアンプに入力して音を出す。それだけ、
#
#
# <概略仕様>
# 波形RAMデータ分解能:16bit
# 波形RAMデータ長:512
#
# マスタクロック周波数:64MHz
#
# PWMキャリア周波数 : 44.1kHz
# PWM分解能 :9bit
# PWM電圧レベル:3.3V
#
# ## 平均律の周波数
#
# A4 = 440Hzを基準に平均律の周波数とFPGAの波形再生用基準クロック(64MHz)の分周器の分周率div_num(正確には分周器の設計上分周率-1)を求める。
#
# まずA4を基準に半音ずつ低い音 A4→G3#→G3→F3#→F3→E3、、、、の周波数をもとめる。
# ($2^{1/12}$を逐次割っていくことで求まる)
pitch=[440]
last_pitch=440
for i in range(26):
last_pitch/=2.**(1./12)
pitch+=[last_pitch]
pitch
# もとめた各ピッチの周波数とマスタクロック周波数64MHz,波形メモリ長512より、対応する分周比-1をもとめる。
# この値をVerilogのパラメータとして持たせておいて分周器の入力としての分周率を可変にすることでピッチ調整のメカニズムにする。
#
# 例えばA4を出す場合は64MHzを282+1=284分周したクロックで波形データを読む。
div_num=[]
for f in pitch:
div_num+=[int(64000000./(512.*f)-0.5)]
div_num
# まずA4より高い音に対しても同様にもとめる。高い音は分周率を求める際の丸め誤差が相対的に大きくなるのでマスタクロックの周波数を上げないと音痴になる。
# 64MHz以上にしたかったがPLLが安定しないのであきらめた。
#
pitch=[440]
last_pitch=440
for i in range(26):
last_pitch*=2.**(1./12)
pitch+=[last_pitch]
pitch
div_num=[]
for f in pitch:
div_num+=[int(64000000./(512.*f)-0.5)]
div_num
# ## TinyFPGAのRAM初期値格納データ(Verilog HDL)生成コード
#
#
# TinyFPGAで音声データを出力する原波形データを作成する。
#
# 512長のRAM(16bit)に一周期分のデータを保存する。
#
# 16bit符号なし整数で0.5オフセットした振幅0.5の正弦波を生成する。
# プロットは3周期分プロットしているがメモリデータとするのは1周期分512データ
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
f_name="ramdata_init.v"
instans_name="RAM_inst"
x=np.linspace(0,2*np.pi*4,512*4,endpoint=False)
y=0.5+0.5*np.sin(x)
#y=0.5+0.17*(np.sin(x)/5.0+np.sin(2*x)/1.5++np.sin(3*x)/1.0+np.sin(4*x)/1.3+np.sin(5*x)/2.0++np.sin(6*x)/3.0)
#y=1.0-(np.exp(-0.35*x))
# -
plt.plot(x,y)
# 0-1の範囲を16bit符号なし整数データ(0-65535)に変換
y_data=(y*(2**16-1)).astype(np.uint16)
np.max(y_data),np.min(y_data)
plt.plot(x,y_data)
# FPGA([iCE40](https://www.latticesemi.com/ja-JP/Products/FPGAandCPLD/iCE40))のメモリインスタンスの初期値を設定するVerilog HDLコードを生成する。
# iCE40のRAMブロックは1ブロック4kbit(256x16bit)なので2ブロック分のRAMブロックインスタンス(RAM_inst0 , RAM_inst1)のデータを生成する。
# +
mem_data=[]
#4kbit RAM block 1
for i in range(16):
str_buf="defparam "+instans_name+"0.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf += "{:04X}".format(y_data[i*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
mem_data += [""]
#4kbit RAM block 2
for i in range(16):
str_buf="defparam "+instans_name+"1.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf+="{:04X}".format(y_data[(i+16)*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
# -
mem_data
# あとは生成したコードを出力してCopy and PasteでVerilog HDLとして使うだけ、、
# ファイル出力してもいいと思ったけど特に必要性も感じないのでこの程度に、、、
# +
for line in mem_data:
print(line)
# -
# ## 実際のwavデータから
#
# 正弦波の音はピーピーって感じで味気ないので倍音成分を加えて楽器みたいにしたい、、とおもい適当な関数とかフーリエ級数でいろいろな波形を適当に作ってやってみても思うようにできなかった。
# なので生音の波形を調べて、それをもとに波形を合成してみるアプローチをとることにした。
#
#
# 生音を保存されたwavファイルのデータ数値化は[PySoundFile](https://pypi.org/project/SoundFile/)でできるようである。
# (anacondaのパッケージには入ってなかったのでpipでインストールした。)
#
# 音源のwavファイルは以下サイトに適当なのがあった。
#
# [https://www.cs.miyazaki-u.ac.jp/~date/lectures/2009ics/kadai/sound2009ics.html](https://www.cs.miyazaki-u.ac.jp/~date/lectures/2009ics/kadai/sound2009ics.html)
#
# #### トランペット音
#
# まずはトランペットの音のwavファイルの波形を調べて、フーリエ級数による合成を試みる。
import soundfile as sf
wav_file_path="target_data/doremi001t.wav"
data, samplerate = sf.read(wav_file_path)
sample_period=1./samplerate
samplerate,sample_period
t=np.linspace(0,data.shape[0]*sample_period,data.shape[0],endpoint=False)
t.shape
# +
wav_data=np.ndarray((2,data.shape[0]),dtype=float)
wav_data[0,:]=t
wav_data[1,:]=data
plt.plot(wav_data[0],wav_data[1])
plt.xlim(0.398,0.408)
# -
# これはC2音(261.6Hz)発音部分の波形を拡大したもの
#
# 実際の波形は上記のようにとんがった波形になる模様
#
# Scipyのfftモジュールをつかってフーリエ変換により周波数成分を調べる。
#
# 解析対象はC2音を出している以下の範囲(2周期分)とする
plt.plot(wav_data[0],wav_data[1])
plt.xlim(0.3993,0.40699)
#plt.xlim(0.6456,0.65334)
# ターゲットとなるデータを切り出す。
#
# +
target_wav_data=wav_data[:,int(0.3993/sample_period):int(0.40699/sample_period)]
t0=target_wav_data[0,0]
target_wav_data[0,:]-=t0
plt.plot(target_wav_data[0],target_wav_data[1])
# -
target_wav_data.shape
target_wav_data[1].mean()
# ターゲット部分に離散時間フーリエ変換を実施。FFTライブラリとして[scipy.fft](https://docs.scipy.org/doc/scipy/reference/fft.html)を利用する。
# データ数nはだいたい1周期分である85を使う。
# 結果は複素数の配列でとりあえず実部、虚部をプロットしてみる。。
# +
import scipy.fft as fft
spectrum_data=fft.fft(target_wav_data[1],85)
# -
type(spectrum_data)
spectrum_data.dtype
spectrum_data.shape
plt.plot(spectrum_data.real)
plt.plot(spectrum_data.imag)
# もうちょっと突っ込んでみる絶対値と偏角計算し、横軸を実周波数にしたプロットをしてみる。。。
spectrum=np.ndarray((3,spectrum_data.shape[0]),dtype=float)
spectrum[0]=np.linspace(0,samplerate,spectrum_data.shape[0])
spectrum[1]=np.abs(spectrum_data)
spectrum[2]=np.angle(spectrum_data)
# +
fig, axs = plt.subplots(2, 1,figsize=[7.,7.]) #2x1のマルチっプロットの作成
axs[0].plot(spectrum[0],spectrum[1])
axs[0].set_title("level")
#axs[0].set_xlim(100,2000)
axs[1].plot(spectrum[0],spectrum[2])
axs[1].set_title("angle")
#axs[1].set_xlim(100,2000)
# -
# 横軸データは実周波数ではなく普通にn番目データとしてみたほうが今回の目的からは扱いやすそうなので横軸は無視すして
# 基本周波数、倍音成分のあるポイントを調べる。
#
# +
fig, axs = plt.subplots(2, 1,figsize=[7.,7.]) #2x1のマルチっプロットの作成
axs[0].plot(spectrum[1])
axs[0].set_title("level")
axs[1].plot(spectrum[2])
axs[1].set_title("angle")
# +
fig, axs = plt.subplots(2, 1,figsize=[7.,7.]) #2x1のマルチっプロットの作成
fc=10
axs[0].plot(spectrum[1])
axs[0].set_title("level")
axs[0].set_xlim(fc-5,fc+5)
axs[1].plot(spectrum[2])
axs[1].set_title("angle")
axs[1].set_xlim(fc-5,fc+5)
# -
# 以上のようにnを1周期分になるようにしたので
#
# n倍高調波の成分はn番目のフーリエ変換データになる。
#
# 40倍高調波の成分のフーリエ級数のパラメータ(振幅、位相)を以下のように取り出す。
#
freq=[]
f_param_amp=[]
f_param_ang=[]
for i in range(1,41):
f_param_amp+=[spectrum[1,i]]
f_param_ang+=[spectrum[2,i]]
#f_param_ang+=[(spectrum[2,i-4:i+5]*spectrum[1,i-4:i+5]).mean()/spectrum[1,i]]
freq+=[spectrum[0][i]]
f_param_amp
f_param_ang
# 試しに波形を再現してみる。
# 30倍高調波までの再現
#
x=np.linspace(0,2*np.pi*2,512*2,endpoint=False)
y=np.zeros(512*2)
for i in range(1,31):
y+=f_param_amp[i-1]*np.cos(i*x+f_param_ang[i-1])
plt.plot(x,y)
plt.plot(target_wav_data[0],target_wav_data[1])
# 元の波形を再掲
plt.plot(target_wav_data[0],target_wav_data[1])
# 見た目思いのほか高い再現度
# 波形のレンジをノーマライズしてメモリ格納形式に変換する。
ymax=np.max(np.abs(y))
y_norm=y/(ymax*2.0)+0.5
y_data=(y_norm*(2**16-1)).astype(np.uint16)
plt.plot(x,y_norm)
y_data=(y_norm*(2**16-1)).astype(np.uint16)
plt.plot(y_data)
# +
mem_data=[]
#4kbit RAM block 1
for i in range(16):
str_buf="defparam "+instans_name+"0.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf += "{:04X}".format(y_data[i*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
mem_data += [""]
#4kbit RAM block 2
for i in range(16):
str_buf="defparam "+instans_name+"1.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf+="{:04X}".format(y_data[(i+16)*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
for line in mem_data:
print(line)
# -
# と波形データは作れたのだが思うような音として再生できてない、、、、
# #### ストリングス音
#
# ストリングス音としてバイオリンの音の波形も同様に調べてみた。
#
# +
wav_file_path="target_data/doremi001v.wav"
data, samplerate = sf.read(wav_file_path)
sample_period=1./samplerate
t=np.linspace(0,data.shape[0]*sample_period,data.shape[0],endpoint=False)
wav_data=np.ndarray((2,data.shape[0]),dtype=float)
wav_data[0,:]=t
wav_data[1,:]=data
plt.plot(wav_data[0],wav_data[1])
#plt.xlim(0.5,0.7)
# -
# バイオリンはトランペットと違いビブラートがあって低周波の揺らぎが強い感じがするのだが、、、それはひとまず無視して先ほど同様2波長分のデータを使うこと位下。
plt.plot(wav_data[0],wav_data[1])
plt.xlim(0.5521,0.5598)
# データ切り出し
# +
target_wav_data=wav_data[:,int(0.5521/sample_period):int(0.5598/sample_period)]
t0=target_wav_data[0,0]
target_wav_data[0,:]-=t0
plt.plot(target_wav_data[0],target_wav_data[1])
# -
target_wav_data.shape
# 先ほどと同様fftを施して振幅、位相プロットする。(nも同様170/2=85)とする
# +
spectrum_data=fft.fft(target_wav_data[1],85)
spectrum=np.ndarray((3,spectrum_data.shape[0]),dtype=float)
spectrum[0]=np.linspace(0,samplerate,spectrum_data.shape[0])
spectrum[1]=np.abs(spectrum_data)
spectrum[2]=np.angle(spectrum_data)
# +
fig, axs = plt.subplots(2, 1,figsize=[7.,7.]) #2x1のマルチっプロットの作成
axs[0].plot(spectrum[1])
axs[0].set_title("level")
axs[1].plot(spectrum[2])
axs[1].set_title("angle")
# -
# 先ほどと同様、40倍高調波の成分のフーリエ級数のパラメータ(振幅、位相)を取り出し25倍高調波までで再現してみる。
#
freq=[]
f_param_amp=[]
f_param_ang=[]
for i in range(1,41):
f_param_amp+=[spectrum[1,i]]
f_param_ang+=[spectrum[2,i]]
#f_param_ang+=[(spectrum[2,i-4:i+5]*spectrum[1,i-4:i+5]).mean()/spectrum[1,i]]
freq+=[spectrum[0][i]]
x=np.linspace(0,2*np.pi*2,512*2,endpoint=False)
y=np.zeros(512*2)
for i in range(1,26):
y+=f_param_amp[i-1]*np.cos(i*x+f_param_ang[i-1])
plt.plot(x,y)
plt.plot(target_wav_data[0],target_wav_data[1])
# 元の波形を再掲
plt.plot(target_wav_data[0],target_wav_data[1])
# 波形のレンジをノーマライズしてメモリ格納形式に変換する。
ymax=np.max(np.abs(y))
y_norm=y/(ymax*2.0)+0.5
y_data=(y_norm*(2**16-1)).astype(np.uint16)
plt.plot(y_data)
# +
mem_data=[]
#4kbit RAM block 1
for i in range(16):
str_buf="defparam "+instans_name+"0.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf += "{:04X}".format(y_data[i*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
mem_data += [""]
#4kbit RAM block 2
for i in range(16):
str_buf="defparam "+instans_name+"1.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf+="{:04X}".format(y_data[(i+16)*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
for line in mem_data:
print(line)
# -
# #### クラリネット音
#
# 木管音としてクラリネットの音の波形も、、、
# +
wav_file_path="target_data/doremi001c.wav"
data, samplerate = sf.read(wav_file_path)
sample_period=1./samplerate
t=np.linspace(0,data.shape[0]*sample_period,data.shape[0],endpoint=False)
wav_data=np.ndarray((2,data.shape[0]),dtype=float)
wav_data[0,:]=t
wav_data[1,:]=data
plt.plot(wav_data[0],wav_data[1])
#plt.xlim(0.5,0.7)
# -
# 感覚としては意外と電子音でも行けそうな感じ
plt.plot(wav_data[0],wav_data[1])
plt.xlim(0.5551,0.5637)
# +
target_wav_data=wav_data[:,int(0.5551/sample_period):int(0.5637/sample_period)]
t0=target_wav_data[0,0]
target_wav_data[0,:]-=t0
plt.plot(target_wav_data[0],target_wav_data[1])
# -
target_wav_data.shape
# 190??
#
# ちょっとこれまでのとピッチが違うような、、
#
# 一応サンプリングレートも確認してみるても同じなので2周期190サンプルでこれまで(170)とちょっと違う模様
#
# フーリエ変換はn=95にすればいいので特に問題はないけど少し気になる。・
samplerate
# +
spectrum_data=fft.fft(target_wav_data[1],95)
spectrum=np.ndarray((3,spectrum_data.shape[0]),dtype=float)
spectrum[0]=np.linspace(0,samplerate,spectrum_data.shape[0])
spectrum[1]=np.abs(spectrum_data)
spectrum[2]=np.angle(spectrum_data)
fig, axs = plt.subplots(2, 1,figsize=[7.,7.]) #2x1のマルチっプロットの作成
axs[0].plot(spectrum[1])
axs[0].set_title("level")
axs[1].plot(spectrum[2])
axs[1].set_title("angle")
# -
# 40倍高調波の成分のフーリエ級数のパラメータ(振幅、位相)を取り出し25倍高調波までで再現。
# +
freq=[]
f_param_amp=[]
f_param_ang=[]
for i in range(1,41):
f_param_amp+=[spectrum[1,i]]
f_param_ang+=[spectrum[2,i]]
#f_param_ang+=[(spectrum[2,i-4:i+5]*spectrum[1,i-4:i+5]).mean()/spectrum[1,i]]
freq+=[spectrum[0][i]]
x=np.linspace(0,2*np.pi*2,512*2,endpoint=False)
y=np.zeros(512*2)
for i in range(1,26):
y+=f_param_amp[i-1]*np.cos(i*x+f_param_ang[i-1])
plt.plot(x,y)
plt.plot(target_wav_data[0],target_wav_data[1])
# -
# 元の波形を再掲
plt.plot(target_wav_data[0],target_wav_data[1])
# +
ymax=np.max(np.abs(y))
y_norm=y/(ymax*2.0)+0.5
y_data=(y_norm*(2**16-1)).astype(np.uint16)
plt.plot(y_data)
mem_data=[]
#4kbit RAM block 1
for i in range(16):
str_buf="defparam "+instans_name+"0.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf += "{:04X}".format(y_data[i*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
mem_data += [""]
#4kbit RAM block 2
for i in range(16):
str_buf="defparam "+instans_name+"1.INIT_"+"{:1X} = 256'h".format(i)
for j in range(16):
str_buf+="{:04X}".format(y_data[(i+16)*16+15-j])
if(j!=15):
str_buf += "_"
str_buf += ";"
mem_data+=[str_buf]
for line in mem_data:
print(line)
# -
|
.ipynb_checkpoints/memdata-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# +
import torch
import torch.nn as nn
import numpy as np
import matplotlib.pyplot as plt
import random
import time
import copy
from collections import namedtuple, deque
# %matplotlib notebook
# GPU if available
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
BOARD_WIDTH = 10
BOARD_HEIGHT = 20
BLANK = 0
TEMPLATE_WIDTH = 5
TEMPLATE_HEIGHT = 5
S_SHAPE_TEMPLATE = [['.....',
'.....',
'..OO.',
'.OO..',
'.....'],
['.....',
'..O..',
'..OO.',
'...O.',
'.....']]
Z_SHAPE_TEMPLATE = [['.....',
'.....',
'.OO..',
'..OO.',
'.....'],
['.....',
'..O..',
'.OO..',
'.O...',
'.....']]
I_SHAPE_TEMPLATE = [['..O..',
'..O..',
'..O..',
'..O..',
'.....'],
['.....',
'.....',
'OOOO.',
'.....',
'.....']]
O_SHAPE_TEMPLATE = [['.....',
'.....',
'.OO..',
'.OO..',
'.....']]
J_SHAPE_TEMPLATE = [['.....',
'.O...',
'.OOO.',
'.....',
'.....'],
['.....',
'..OO.',
'..O..',
'..O..',
'.....'],
['.....',
'.....',
'.OOO.',
'...O.',
'.....'],
['.....',
'..O..',
'..O..',
'.OO..',
'.....']]
L_SHAPE_TEMPLATE = [['.....',
'...O.',
'.OOO.',
'.....',
'.....'],
['.....',
'..O..',
'..O..',
'..OO.',
'.....'],
['.....',
'.....',
'.OOO.',
'.O...',
'.....'],
['.....',
'.OO..',
'..O..',
'..O..',
'.....']]
T_SHAPE_TEMPLATE = [['.....',
'..O..',
'.OOO.',
'.....',
'.....'],
['.....',
'..O..',
'..OO.',
'..O..',
'.....'],
['.....',
'.....',
'.OOO.',
'..O..',
'.....'],
['.....',
'..O..',
'.OO..',
'..O..',
'.....']]
PIECES = {'S': S_SHAPE_TEMPLATE,
'Z': Z_SHAPE_TEMPLATE,
'J': J_SHAPE_TEMPLATE,
'L': L_SHAPE_TEMPLATE,
'I': I_SHAPE_TEMPLATE,
'O': O_SHAPE_TEMPLATE,
'T': T_SHAPE_TEMPLATE}
PIECES_IND = {'S': 0,
'Z': 1,
'J': 2,
'L': 3,
'I': 4,
'O': 5,
'T': 6}
PIECES_MARGINS = {'S': [[1, 1, 0], [0, 1, 1]],
'Z': [[1, 1, 0], [1, 0, 1]],
'J': [[1, 1, 1], [0, 1, 1], [1, 1, 0], [1, 0, 1]],
'L': [[1, 1, 1], [0, 1, 1], [1, 1, 0], [1, 0, 1]],
'I': [[0, 0, 2], [2, 1, 0]],
'O': [[1, 0, 0]],
'T': [[1, 1, 1], [0, 1, 1], [1, 1, 0], [1, 0, 1]]}
class Tetris:
def __init__(self):
self.board = self.getBlankBoard()
self.current_piece = self.getNewPiece()
def reset(self):
"""
Restarts the game with a blank board and new piece.
@rtype: torch tensor
A tensor representing the state.
"""
self.board = self.getBlankBoard()
self.current_piece = self.getNewPiece()
return self.convertToFeatures(self.board)
def isOnBoard(self, x, y):
"""
Checks if the position (x,y) is on the board.
@type x: int
The x position
@type y: int
The y position
@rtype: Boolean
If (x,y) is on the board.
"""
return 0 <= x < BOARD_WIDTH and 0 <= y < BOARD_HEIGHT
def getBlankBoard(self):
"""
Returns a blank board.
"""
return np.zeros((BOARD_WIDTH, BOARD_HEIGHT))
def isValidPosition(self, board, piece, x, y, rotation):
"""
Checks if a piece has a valid position on the board.
@type board: np.array
A np array representing the state of the board.
@type piece: string
A string representing the shape of the piece.
@type x: int
The x position of the piece.
@type y: int
The y position of the piece.
@rotation: int
The rotation of the piece.
@rtype: Boolean
If the piece has a valid position on the board.
"""
for dx in range(TEMPLATE_WIDTH):
for dy in range(TEMPLATE_HEIGHT):
template = PIECES[piece][rotation % len(PIECES[piece])]
if template[dy][dx] == 'O':
board_x_pos, board_y_pos = x + (dx - 2), y - (dy - 2)
if not self.isOnBoard(board_x_pos, board_y_pos) or board[board_x_pos][board_y_pos]:
return False
return True
def getNewPiece(self):
"""
Gets a new piece.
@rtype: string
A string representing the shape of the new piece.
"""
return random.choice(list(PIECES.keys()))
def findXYCoordinate(self, piece, action, board):
"""
Find the x and y coordinates to place a piece given an action.
@type piece: string
A letter representing the shape of the piece to be placed.
@type action: int
An integer representing the action.
@type board: np.array
A np array representing the state of the board.
@rtype: tuple[int]
A tuple (x, y, rotation) representing the (x,y) coordinates of the
piece if it were to be placed on the board as well as the rotation
of the piece. Note that this does not actually place the piece.
"""
rotation = action % 4
left_margin, right_margin, top_margin = PIECES_MARGINS[piece][
rotation % len(PIECES_MARGINS[piece])]
x = max(left_margin, min(action // 4, BOARD_WIDTH - right_margin - 1))
# Finding y coordinate to place the piece
valid_y = None
flag = False
for y in range(BOARD_HEIGHT - top_margin - 1, -2, -1):
if self.isValidPosition(board, piece, x, y, rotation):
flag = True
else:
if flag:
valid_y = y + 1
break
return x, valid_y, rotation
def transitionState(self, action):
"""
Returns the next state given the action.
@type action: int
An integer representing the action chosen.
In total, there are BOARD_WIDTH x 4 actions, representing
choices in the x coordinate and rotation of the piece.
For a chosen x and rotation r, the action is 4 * x + r.
@rtype: tuple
A tuple (reward, next_state, done) representing the reward, next state,
and if the game has finished.
"""
x, y, rotation = self.findXYCoordinate(self.current_piece, action, self.board)
if y != None:
self.board = self.placeOnBoard(self.current_piece, x, y, rotation, self.board)
self.current_piece = self.getNewPiece()
next_state = self.convertToFeatures(self.board)
self.board, lines_cleared = self.clearLines(self.board)
#delta_r, delta_c = self.countHoles(self.board)
#reward = lines_cleared**2 - delta_r - delta_c + 0.1
reward = 0.1 + (lines_cleared+1)**2
return reward, next_state, False, lines_cleared
return -1, self.convertToFeatures(self.board), True, 0
def getAllNextStates(self):
"""
Get all of the next states corresponding to all possible next actions.
@rtype: list[tuple]
A list of tuples (action, features) representing the features of the
next state if an action is taken.
"""
data = []
for action in range(BOARD_WIDTH * 4):
x, y, rotation = self.findXYCoordinate(self.current_piece, action, self.board)
if y != None:
self.board = self.placeOnBoard(self.current_piece, x, y, rotation, self.board)
features = self.convertToFeatures(self.board)
data.append((action, features))
self.board = self.removeFromBoard(self.current_piece, x, y, rotation, self.board)
return data
def removeFromBoard(self, piece, x, y, rotation, board):
"""
removes the current piece on the board.
@type piece: string
A letter representing the shape of the piece.
@type x: int
The x position of the piece.
@type y: int
The y position of the piece.
@type rotation: int
The rotation of the piece.
@type board: np.array
A np array representing the board.
@type: np.array
A np array representing the board after the piece has been removed.
"""
template = PIECES[piece][rotation % len(PIECES[piece])]
for dx in range(TEMPLATE_WIDTH):
for dy in range(TEMPLATE_HEIGHT):
if template[dy][dx] == 'O':
board_x_pos, board_y_pos = x + (dx - 2), y - (dy - 2)
board[board_x_pos][board_y_pos] = 0.0
return board
def placeOnBoard(self, piece, x, y, rotation, board):
"""
Places the current piece on the board. Assumes that the piece
is in a valid position.
@type piece: string
A letter representing the shape of the piece.
@type x: int
The x position of the piece.
@type y: int
The y position of the piece.
@type rotation: int
The rotation of the piece.
@type board: np.array
A np array representing the board.
@type: np.array
A np array representing the board after the piece has been placed.
"""
template = PIECES[piece][rotation % len(PIECES[piece])]
for dx in range(TEMPLATE_WIDTH):
for dy in range(TEMPLATE_HEIGHT):
if template[dy][dx] == 'O':
board_x_pos, board_y_pos = x + (dx - 2), y - (dy - 2)
board[board_x_pos][board_y_pos] = 1.0
return board
def clearLines(self, board):
"""
Removes completed lines from the board.
@rtype: int
The number of lines removed.
"""
lines_removed = 0
y = 0 # start y at the bottom of the board
while y < BOARD_HEIGHT:
if self.isCompleteLine(y, board):
# Remove the line and pull boxes down by one line.
for pull_down_Y in range(y, BOARD_HEIGHT - 1):
for x in range(BOARD_WIDTH):
board[x][pull_down_Y] = board[x][pull_down_Y + 1]
# Set very top line to blank.
for x in range(BOARD_WIDTH):
board[x][BOARD_HEIGHT - 1] = BLANK
lines_removed += 1
# Note on the next iteration of the loop, y is the same.
# This is so that if the line that was pulled down is also
# complete, it will be removed.
else:
y += 1 # move on to check next row up
return board, lines_removed
def countCompleteLines(self, board):
"""
Counts the number of completed lines.
@type board: np.array
An np array representing the board.
@rtype: int
The number of completed lines on the board.
"""
completed_lines = 0
for y in range(BOARD_HEIGHT):
if self.isCompleteLine(y, board):
completed_lines += 1
return completed_lines
def isCompleteLine(self, y, board):
"""
Checks if the line at height y is complete.
@type y: int
The height of the row to check.
@rtype : Boolean
True if the row is complete.
"""
for x in range(BOARD_WIDTH):
if board[x][y] == 0.0: return False
return True
def convertToFeatures(self, board):
"""
Converts the current board position and falling piece to a
list of features.
The features consist of:
- Number of holes along the vertical and horizontal directions.
- Total height of all columns.
- Bumpiness.
- Number of completed lines on the board.
@rtype: torch tensor
Torch tensor of the features described above. Values normalized to be between -1 and 1.
"""
delta_r, delta_c = self.countHoles(board)
total_height, bumpiness = self.scoreBumpiness(board)
completed_lines = self.countCompleteLines(board)
return torch.tensor([[delta_r/BOARD_HEIGHT, delta_c/BOARD_WIDTH,
total_height/BOARD_WIDTH,
bumpiness/BOARD_WIDTH, completed_lines*3]],
dtype=torch.float32).to(device)
def countHoles(self, board):
"""
Counts the number of transitions from filled to empty or vice
versa in the rows and columns.
@rtype: tuple[int]
A tuple (delta_r, delta_c) representing the number of transitions
from filled to empty squares or vice versa across rows and columns respectively.
"""
# Across rows:
delta_r = 0
for y in range(BOARD_HEIGHT):
for x in range(BOARD_WIDTH - 1):
if board[x][y] != board[x + 1][y]:
delta_r += 1
# Across columns:
delta_c = 0
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT - 1):
if board[x][y] != board[x][y + 1]:
delta_c += 1
return delta_r, delta_c
def scoreBumpiness(self, board):
"""
Calculates the "bumpiness" of the board, defined as the
sum of the absolute differences in heights of adjacent columns,
except for the largest difference.
@type board: np.array
A np array representing the board.
@rtype: tuple[int]
A tuple (max_height, bumpiness) representing the
maximum height of any column and the bumpiness.
"""
heights = []
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT-1, -1, -1):
if board[x][y]: break
heights.append(y)
diffs = [abs(heights[i]-heights[i-1]) for i in range(1, len(heights))]
diffs.sort()
return sum(heights), sum(diffs)-diffs[-1]
Transition = namedtuple('Transition', ('state', 'next_state', 'reward', 'done'))
class ReplayMemory:
def __init__(self, capacity):
"""
Initializes the replay memory.
@type capacity: int
The capacity of the memory.
"""
self.memory = deque([], maxlen=capacity)
def push(self, *args):
"""Saves a transition.
"""
self.memory.append(Transition(*args))
def sample(self, batch_size):
"""
Samples a batch of transitions of size batch_size
randomly.
@type batch_size: int
The size of the batch.
@rtype: list[tuple]
A list of the sampled transitions.
"""
return random.sample(self.memory, batch_size)
def __len__(self):
return len(self.memory)
class Agent:
""" Agent object that uses the actor-critic network to find the
optimal policy.
"""
def __init__(self, env, NN, optimizer, criterion):
""" Initializes the agent.
@type env: Tetris
The Tetris environment.
@type NN: NeuralNet
Neural network for computing the state-action values.
@type optimizer: torch.optim
Torch optimizer object.
@type criterion: nn loss
Neural network loss function.
"""
self.env = env
self.NN = NN
self.optimizer = optimizer
self.criterion = criterion
def chooseAction(self, epsilon):
""" Chooses action. With probability epsilon it will choose
an exploratory action. Otherwise, it will choose
an action which maximizes the estimated reward of all
possible next states.
@type epsilon: float
Exploration probability.
@rtype: int
An integer representing the action.
"""
if random.uniform(0, 1) < epsilon:
return random.randrange(BOARD_WIDTH * 4)
cur_best_val = -float('inf')
cur_best_action = 0
data = env.getAllNextStates()
for action, state in data:
value = self.NN(state).item()
if value > cur_best_val:
cur_best_val = value
cur_best_action = action
return cur_best_action
def optimizeModel(self, memory, batch_size, gamma):
"""
Performs one step of mini-batch gradient descent.
@type memory: ReplayMemory
The replay memory from which to draw the experience.
@type batch_size: int
The mini-batch size.
@type gamma: float
The discount factor.
"""
batch_size = min(len(memory), batch_size)
# Sampling experiences
transitions = memory.sample(batch_size)
batch = Transition(*zip(*transitions))
# Unpacking data
state_batch = torch.cat(batch.state)
next_state_batch = torch.cat(batch.next_state)
reward_batch = torch.cat(batch.reward)
done_batch = torch.cat(batch.done)
# Predictions and targets
predictions = self.NN(state_batch)
with torch.no_grad():
targets = reward_batch + gamma * self.NN(next_state_batch) * done_batch
# Loss and gradient descent
loss = self.criterion(predictions, targets)
loss.backward()
self.optimizer.step()
self.optimizer.zero_grad()
def train(self, episodes, epsilon_initial, epsilon_min, epsilon_stop_episode, gamma, memory_capacity, batch_size):
""" Trains the agent using the actor-critic method with eligibility traces.
@type episodes: int
The number of episodes to train.
@type epsilon: float
The exploration probability.
@type gamma: float
The discount factor.
@type memory_capacity: int
The capacity of the replay memory.
@type batch_size: int
Mini-batch size for training.
"""
# %matplotlib
fig = plt.gcf()
fig.show()
fig.canvas.draw()
plt.grid()
plt.xlim(-0.5, BOARD_WIDTH - 0.5)
plt.ylim(BOARD_HEIGHT - 0.5, -0.5)
memory = ReplayMemory(memory_capacity)
tot_steps = 0
LC = 0
depsilon = (epsilon_initial-epsilon_min)/epsilon_stop_episode
for episode in range(episodes):
if epsilon_initial > epsilon_min:
epsilon_initial -= depsilon
if (episode + 1) % 10 == 0:
print(f'Episode {episode + 1}/{episodes} completed!')
torch.save(self.NN.state_dict(), 'tetris_NN_value_model')
print(f'Average steps per episode: {tot_steps / 10}')
print(f'Average lines cleared per episode: {LC / 10}')
tot_steps = 0
LC = 0
state, done = self.env.reset(), False
# Initialize eligibility traces
while not done:
tot_steps += 1
time.sleep(0.2)
plt.imshow(np.transpose(self.env.board)[::-1], cmap=plt.cm.binary, interpolation='none', origin='lower')
ax = plt.gca()
ax.set_xticks(np.arange(-0.5, BOARD_WIDTH - 0.5, 1))
ax.set_yticks(np.arange(0.5, BOARD_HEIGHT - 0.5, 1))
fig.canvas.draw()
action = self.chooseAction(epsilon_initial)
reward, next_state, done, lines_cleared = self.env.transitionState(action)
reward = torch.tensor([[reward]], device=device)
done = torch.tensor([[done]], device=device)
# Saves the transition
memory.push(state, next_state, reward, done)
LC += lines_cleared
# Perform one step of batch gradient descent
self.optimizeModel(memory, batch_size, gamma)
state = next_state
class QNetwork(nn.Module):
def __init__(self, input_size, hidden_size1, hidden_size2):
super(QNetwork, self).__init__()
self.l1 = nn.Linear(input_size, hidden_size1)
self.l2 = nn.Linear(hidden_size1, hidden_size2)
self.l3 = nn.Linear(hidden_size2, 1)
def forward(self, x):
x = torch.relu(self.l1(x))
x = torch.relu(self.l2(x))
x = self.l3(x)
return x
if __name__ == "__main__":
# Network parameters
input_size = 5
hidden_size1 = 32
hidden_size2 = 16
# Training parameters
episodes = 1000000
gamma = 0.8
learning_rate = 2e-3
epsilon_initial = 0.1
epsilon_min = 0.1
epsilon_stop_episode = 1500
memory_capacity = 10000
batch_size = 16
env = Tetris()
model_value = QNetwork(input_size, hidden_size1, hidden_size2).to(device)
criterion = nn.MSELoss()
optimizer = torch.optim.SGD(model_value.parameters(), lr=learning_rate)
model_value.load_state_dict(torch.load('tetris_NN_value_model'))
tetris_agent = Agent(env, model_value, optimizer, criterion)
tetris_agent.train(episodes, epsilon_initial, epsilon_min, epsilon_stop_episode, gamma, memory_capacity, batch_size)
# -
|
Testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Plotting distributions on the same graph
# +
# libraries & dataset
import seaborn as sns
import matplotlib.pyplot as plt
# set a grey background (use sns.set_theme() if seaborn version 0.11.0 or above)
sns.set(style="darkgrid")
df = sns.load_dataset("iris")
sns.histplot(data=df, x="sepal_length", color="skyblue", label="Sepal Length", kde=True)
sns.histplot(data=df, x="sepal_width", color="red", label="Sepal Width", kde=True)
plt.legend()
plt.show()
# -
# ## Splitting the figure in as much axes as needed
# +
# libraries & dataset
import seaborn as sns
import matplotlib.pyplot as plt
# set a grey background (use sns.set_theme() if seaborn version 0.11.0 or above)
sns.set(style="darkgrid")
df = sns.load_dataset("iris")
fig, axs = plt.subplots(2, 2, figsize=(7, 7))
sns.histplot(data=df, x="sepal_length", kde=True, color="skyblue", ax=axs[0, 0])
sns.histplot(data=df, x="sepal_width", kde=True, color="olive", ax=axs[0, 1])
sns.histplot(data=df, x="petal_length", kde=True, color="gold", ax=axs[1, 0])
sns.histplot(data=df, x="petal_width", kde=True, color="teal", ax=axs[1, 1])
plt.show()
|
src/notebooks/25-histogram-with-several-variables-seaborn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
# # Simple Demo for tf.string_split
#
# Reference:
# * [Tensorflow tf.string_split Demo](https://github.com/tensorflow/transform/blob/master/tensorflow_transform/mappers_test.py#L117)
string_tensor = tf.constant(['One was Johnny', 'Two was a rat'])
tokenized_tensor = tf.string_split(string_tensor, delimiter=' ')
with tf.Session() as sess:
print(tokenized_tensor.eval())
# # How to convert [tf.string] to [np.array]
#
# 1. as **tf.map_fn** has to keep dimension, just use tf.map_fn for string process and keep_dim
# 2. **keypoints:**:
# * using tf.string_split for split for all, then to SparseTensorValue
# * convert SparseTensorValue to dense matrix, via tf.sparse_tensor_to_dense
# * convert all of element of dense from string to numeric
# +
## 1. example 1 : just for converting to qualified string matrix
x = tf.placeholder(tf.string)
def parse(x):
x = tf.regex_replace(x, "\[", "")
x = tf.regex_replace(x, "\]", "")
return x
output_strs = tf.map_fn(parse, x)
t1 = tf.string_split(output_strs, delimiter=",")
target_indices, target_values = t1.indices, tf.strings.to_number(t1.values)
with tf.Session() as sess:
print(sess.run(target_values, feed_dict={x: ["[1.0,2.0]", "[2.0,3.0]"]}))
# +
## 2. example 2 : matrix conversion for follow-ups
x = tf.placeholder(tf.string)
y = tf.map_fn(lambda elem: tf.regex_replace(elem, "[\[|\]]", ""), x, dtype=tf.string)
y = tf.string_split(y, delimiter=",")
y = tf.sparse_tensor_to_dense(y, default_value="")
y = tf.strings.to_number(y)
with tf.Session() as sess:
print(sess.run(y, feed_dict={x: ["[1.0,2.0]", "[2.0,3.0]"]}))
# -
|
05_basic/string_list_tensor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append("../../")
import warnings
import numpy as np
import astropy.units as u
from astropy.constants import h, e, m_e, c, sigma_T
from agnpy.synchrotron import R, epsilon_equivalency
from agnpy.compton import compton_kernel
from agnpy.emission_regions import Blob
from agnpy.targets import PointSourceBehindJet
import matplotlib.pyplot as plt
e = e.gauss
# -
# ## my implementation of the trapezoidal rule in log-log space
np.seterr(all='print')
warnings.filterwarnings('error')
# +
def log(x):
# smallest positive float (before 0)
float_tiny = np.finfo(np.float64).tiny
# largest positive float
float_max = np.finfo(np.float64).max
values = np.clip(x, float_tiny, float_max)
return np.log(values)
def power(x):
try:
x ** m
except Warning:
print("too big power!")
# -
def trapz_loglog(y, x, axis=0):
"""
Integrate along the given axis using the composite trapezoidal rule in
loglog space.
Integrate `y` (`x`) along given axis in loglog space.
Parameters
----------
y : array_like
Input array to integrate.
x : array_like, optional
Independent variable to integrate over.
axis : int, optional
Specify the axis.
Returns
-------
trapz : float
Definite integral as approximated by trapezoidal rule in loglog space.
"""
try:
y_unit = y.unit
y = y.value
except AttributeError:
y_unit = 1.0
try:
x_unit = x.unit
x = x.value
except AttributeError:
x_unit = 1.0
slice_low = [slice(None)] * y.ndim
slice_up = [slice(None)] * y.ndim
# multi-dimensional equivalent of x_low = x[:-1]
slice_low[axis] = slice(None, -1)
# multi-dimensional equivalent of x_up = x[1:]
slice_up[axis] = slice(1, None)
slice_low = tuple(slice_low)
slice_up = tuple(slice_up)
# reshape x to be broadcasted with y
if x.ndim == 1:
shape = [1] * y.ndim
shape[axis] = x.shape[0]
x = x.reshape(shape)
x_low = x[slice_low]
x_up = x[slice_up]
y_low = y[slice_low]
y_up = y[slice_up]
log_x_low = log(x_low)
log_x_up = log(x_up)
log_y_low = log(y_low)
log_y_up = log(y_up)
# index in the bin
m = (log_y_low - log_y_up) / (log_x_low - log_x_up)
vals = y_low / (m + 1) * (x_up * (x_up / x_low) ** m - x_low)
# value of y very close to zero will make m large and explode the exponential
tozero = (
np.isclose(y_low, 0, atol=0, rtol=1e-10) +
np.isclose(y_up, 0, atol=0, rtol=1e-10) +
np.isclose(x_low, x_up, atol=0, rtol=1e-10)
)
vals[tozero] = 0.0
return np.add.reduce(vals, axis) * x_unit * y_unit
# ## a simple test with a straight line in log-log scale
# +
def line_loglog(x, m, n):
"""a straight line in loglog-space"""
return x ** m * np.e ** n
def integral_line_loglog(x_min, x_max, m, n):
"""analytical integral of the line in log-log space"""
f_low = line_loglog(x_min, m + 1, n) / (m + 1)
f_up = line_loglog(x_max, m + 1, n) / (m + 1)
return f_up - f_low
# -
m = 1.5
n = -2.0
x = np.logspace(2, 5)
y = line_loglog(x, m, n)
y = np.asarray([y, y])
trapz_loglog(y.T, x, axis=0)
integral_line_loglog(x[0], x[-1], m, n)
np.trapz(y.T, x, axis=0)
1 - trapz_loglog(y.T, x, axis=0) / integral_line_loglog(x[0], x[-1], m, n)
1 - np.trapz(y.T, x, axis=0) / integral_line_loglog(x[0], x[-1], m, n)
# ## a test with synchrotron radiation
# +
blob = Blob()
nu = np.logspace(9, 20, 20) * u.Hz
# check the blob
print(blob)
# -
def sed_synch(nu, integration):
"""compute the synchrotron SED"""
epsilon = nu.to("", equivalencies=epsilon_equivalency)
# correct epsilon to the jet comoving frame
epsilon_prime = (1 + blob.z) * epsilon / blob.delta_D
# electrond distribution lorentz factor
gamma = blob.gamma
N_e = blob.N_e(gamma)
prefactor = np.sqrt(3) * epsilon * np.power(e, 3) * blob.B_cgs / h
# for multidimensional integration
# axis 0: electrons gamma
# axis 1: photons epsilon
# arrays starting with _ are multidimensional and used for integration
_gamma = np.reshape(gamma, (gamma.size, 1))
_N_e = np.reshape(N_e, (N_e.size, 1))
_epsilon = np.reshape(epsilon, (1, epsilon.size))
x_num = 4 * np.pi * _epsilon * np.power(m_e, 2) * np.power(c, 3)
x_denom = 3 * e * blob.B_cgs * h * np.power(_gamma, 2)
x = (x_num / x_denom).to_value("")
integrand = _N_e * R(x)
integral = integration(integrand, gamma, axis=0)
emissivity = (prefactor * integral).to("erg s-1")
sed_conversion = np.power(blob.delta_D, 4) / (
4 * np.pi * np.power(blob.d_L, 2)
)
return (sed_conversion * emissivity).to("erg cm-2 s-1")
# %%timeit
sed_synch(nu, np.trapz)
# %%timeit
sed_synch(nu, trapz_loglog)
sed_trapz = sed_synch(nu, np.trapz)
sed_trapz_loglog = sed_synch(nu, trapz_loglog)
plt.loglog(nu, sed_trapz, marker="o")
plt.loglog(nu, sed_trapz_loglog, ls="--", marker=".")
plt.show()
# ## a test with inverse Compton radiation
# ### EC on point-like source
def sed_flux_point_source(nu, target, r, integrate):
"""SED flux for EC on a point like source behind the jet
Parameters
----------
nu : `~astropy.units.Quantity`
array of frequencies, in Hz, to compute the sed, **note** these are
observed frequencies (observer frame).
"""
# define the dimensionless energy
epsilon_s = nu.to("", equivalencies=epsilon_equivalency)
# transform to BH frame
epsilon_s *= 1 + blob.z
# for multidimensional integration
# axis 0: gamma
# axis 1: epsilon_s
# arrays starting with _ are multidimensional and used for integration
gamma = blob.gamma_to_integrate
transformed_N_e = blob.N_e(gamma / blob.delta_D).value
_gamma = np.reshape(gamma, (gamma.size, 1))
_N_e = np.reshape(transformed_N_e, (transformed_N_e.size, 1))
_epsilon_s = np.reshape(epsilon_s, (1, epsilon_s.size))
# define integrating function
# notice once the value of mu = 1, phi can assume any value, we put 0
# convenience
_kernel = compton_kernel(
_gamma, _epsilon_s, target.epsilon_0, blob.mu_s, 1, 0
)
_integrand = np.power(_gamma, -2) * _N_e * _kernel
integral_gamma = integrate(_integrand, gamma, axis=0)
prefactor_num = (
3
* sigma_T
* target.L_0
* np.power(epsilon_s, 2)
* np.power(blob.delta_D, 3)
)
prefactor_denom = (
np.power(2, 7)
* np.power(np.pi, 2)
* np.power(blob.d_L, 2)
* np.power(r, 2)
* np.power(target.epsilon_0, 2)
)
sed = prefactor_num / prefactor_denom * integral_gamma
return sed.to("erg cm-2 s-1")
# +
# target and distance
r = 1e16 * u.cm
L_0 = 2e46 * u.Unit("erg s-1")
epsilon_0 = 1e-3
ps = PointSourceBehindJet(L_0, epsilon_0)
nu = np.logspace(20, 30) * u.Hz
# increase the size of the gamma grid
blob.set_gamma_size(500)
# -
# %%timeit
sed_flux_point_source(nu, ps, r, np.trapz)
# %%timeit
sed_flux_point_source(nu, ps, r, trapz_loglog)
sed_trapz = sed_flux_point_source(nu, ps, r, np.trapz)
sed_trapz_loglog = sed_flux_point_source(nu, ps, r, trapz_loglog)
plt.loglog(nu, sed_trapz, marker="o")
plt.loglog(nu, sed_trapz_loglog, ls="--", marker=".")
plt.show()
|
experiments/basic/trapz_loglog_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Using bokeh with pandas
# Open / plot data from csv files
# +
from bokeh.plotting import figure # Container to add plotting information
from bokeh.io import output_file, show
import numpy as np
import os
import pandas as pd
direc = os.path.join('/Users','elena','Documents','ProfessionalDevelopment',
'Udemy','DataVisualization-Bokeh')
# Data on number of female graduates
# each year in different fields
data = pd.read_csv(os.path.join(direc,'bachelors.csv'))
# -
data.columns
f = figure()
f.line(data['Year'],data['Engineering'])
show(f)
|
Bokeh_and_Pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from matplotlib import pyplot as plt
plt.style.use('seaborn')
data = pd.read_csv('data4.csv')
ages = data['Age']
dev_salaries = data['All_Devs']
py_salaries = data['Python']
js_salaries = data['JavaScript']
plt.plot(ages, py_salaries, label='Python')
plt.plot(ages, js_salaries, label='JavaScript')
plt.plot(ages, dev_salaries, color='#444444',
linestyle='--', label='All Devs')
plt.legend()
plt.title('Median Salary (USD) by Age')
plt.xlabel('Ages')
plt.ylabel('Median Salary (USD)')
plt.tight_layout()
plt.show()
# +
import pandas as pd
from matplotlib import pyplot as plt
plt.style.use('seaborn')
data = pd.read_csv('data4.csv')
ages = data['Age']
dev_salaries = data['All_Devs']
py_salaries = data['Python']
js_salaries = data['JavaScript']
fig, ax = plt.subplots()
ax.plot(ages, py_salaries, label='Python')
ax.plot(ages, js_salaries, label='JavaScript')
ax.plot(ages, dev_salaries, color='#444444',
linestyle='--', label='All Devs')
ax.legend()
ax.set_title('Median Salary (USD) by Age')
ax.set_xlabel('Ages')
ax.set_ylabel('Median Salary (USD)')
plt.tight_layout()
plt.show()
# +
import pandas as pd
from matplotlib import pyplot as plt
plt.style.use('seaborn')
data = pd.read_csv('data4.csv')
ages = data['Age']
dev_salaries = data['All_Devs']
py_salaries = data['Python']
js_salaries = data['JavaScript']
fig, (ax1,ax2) = plt.subplots(nrows=2,ncols=1)
ax1.plot(ages, py_salaries, label='Python')
ax2.plot(ages, js_salaries, label='JavaScript')
ax2.plot(ages, dev_salaries, color='#444444',
linestyle='--', label='All Devs')
ax1.legend()
ax1.set_title('Median Salary (USD) by Age')
ax1.set_ylabel('Median Salary (USD)')
ax2.legend()
ax2.set_xlabel('Ages')
ax2.set_ylabel('Median Salary (USD)')
plt.tight_layout()
plt.show()
# +
import pandas as pd
from matplotlib import pyplot as plt
plt.style.use('seaborn')
data = pd.read_csv('data4.csv')
ages = data['Age']
dev_salaries = data['All_Devs']
py_salaries = data['Python']
js_salaries = data['JavaScript']
fig, (ax1,ax2) = plt.subplots(nrows=2,ncols=1,sharex=True)
ax1.plot(ages, py_salaries, label='Python')
ax2.plot(ages, js_salaries, label='JavaScript')
ax2.plot(ages, dev_salaries, color='#444444',
linestyle='--', label='All Devs')
ax1.legend()
ax1.set_title('Median Salary (USD) by Age')
ax1.set_ylabel('Median Salary (USD)')
ax2.legend()
ax2.set_xlabel('Ages')
ax2.set_ylabel('Median Salary (USD)')
plt.tight_layout()
plt.show()
# +
import pandas as pd
from matplotlib import pyplot as plt
plt.style.use('seaborn')
data = pd.read_csv('data4.csv')
ages = data['Age']
dev_salaries = data['All_Devs']
py_salaries = data['Python']
js_salaries = data['JavaScript']
fig1,ax1 = plt.subplots()
fig2,ax2 = plt.subplots()
ax1.plot(ages, py_salaries, label='Python')
ax2.plot(ages, js_salaries, label='JavaScript')
ax2.plot(ages, dev_salaries, color='#444444',
linestyle='--', label='All Devs')
ax1.legend()
ax1.set_title('Median Salary (USD) by Age')
ax1.set_ylabel('Median Salary (USD)')
ax2.legend()
ax2.set_xlabel('Ages')
ax2.set_ylabel('Median Salary (USD)')
plt.tight_layout()
plt.show()
# +
import pandas as pd
from matplotlib import pyplot as plt
plt.style.use('seaborn')
data = pd.read_csv('data4.csv')
ages = data['Age']
dev_salaries = data['All_Devs']
py_salaries = data['Python']
js_salaries = data['JavaScript']
fig1,ax1 = plt.subplots()
fig2,ax2 = plt.subplots()
ax1.plot(ages, py_salaries, label='Python')
ax2.plot(ages, js_salaries, label='JavaScript')
ax2.plot(ages, dev_salaries, color='#444444',
linestyle='--', label='All Devs')
ax1.legend()
ax1.set_title('Median Salary (USD) by Age')
ax1.set_ylabel('Median Salary (USD)')
ax2.legend()
ax2.set_xlabel('Ages')
ax2.set_ylabel('Median Salary (USD)')
plt.tight_layout()
plt.show()
fig1.savefig('fig1.png')
# -
|
05-Machine-Learning-Code/数据分析工具/Matplotlib/.ipynb_checkpoints/10_subplot-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# name: python391jvsc74a57bd0f727a3ef71f8e000ac8e615c9588a9ba2bacd5f4ec63f2919734c7064d9d0e56
# ---
# # Tutorial
#
# > Tutorial details
from nbdev.export import notebook2script; notebook2script()
|
02_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import fastText
import math
import numpy as np
from numpy import random
from keras.models import Sequential, Model
from keras.layers import LSTM, Dense, Conv1D, Conv2D, Dropout, MaxPooling1D, GlobalMaxPooling1D, Bidirectional, Input, Masking, Flatten, Concatenate
from keras import regularizers
import os
import re
import threading
line_number=0
lock = threading.Lock()
os.getcwd()
# +
ft = fastText.load_model("/home/jindal/notebooks/fastText/wiki.de.bin")
nb_embedding_dims = ft.get_dimension()
nb_sequence_length = 75
# +
# words = ["mein", "dein", "unser", "mein"]
# for w in words:
# if w in word_vectors:
# print("I know " + w)
# else:
# wv = ft.get_word_vector(w)
# print(wv)
# word_vectors[w] = wv
# -
print(nb_embedding_dims)
print(ft.get_word_vector("mein"))
def my_generator(features, labels, batch_size):
global line_number, lock
batch_features = np.zeros((batch_size, nb_sequence_length, nb_embedding_dims)) # initializing features with zeros
batch_labels = np.zeros((batch_size, 2)) # 2 as one hot
while True:
# print(len(features))
for i in range(batch_size):
lock.acquire()
index = line_number%len(features)
line_number+=1
lock.release()
# index = random.choice(len(features), 1)[0]
# print(index)
batch_features[i] = process_features(features[index], nb_sequence_length, nb_embedding_dims)
# print(batch_features[i])
# print(batch_features[i].shape)
batch_labels[i] = labels[index]
yield batch_features, batch_labels
word_vectors = {}
def process_features(textline, nb_sequence_length, nb_embedding_dims): # given a sentence, returns the embedding for it of fixed size
words = re.compile('[\w-]+|[\W ]+', re.UNICODE).findall(textline.strip())
words = [w.strip() for w in words if w.strip() != '']
# print(words)
features = np.zeros((nb_sequence_length, nb_embedding_dims))
max_words = min(len(words), nb_sequence_length)
idx = nb_sequence_length - len(words[:max_words])
for w in words[:max_words]:
if w in word_vectors:
wv = word_vectors[w]
else:
wv = ft.get_word_vector(w.lower())
word_vectors[w] = wv
features[idx] = wv
# print(str(idx) + " " + w)
idx = idx + 1
return features
train_lines = [line.strip().split("\t") for line in open('/home/jindal/notebooks/Resources/OffLang/sample_train.txt', encoding = "UTF-8")]
dev_lines = [line.strip().split("\t") for line in open('/home/jindal/notebooks/Resources/OffLang/sample_dev.txt', encoding = "UTF-8")]
# +
from keras.utils import to_categorical
train_sentences = [x[0] for x in train_lines]
train_labels = to_categorical([0 if x[1] == "OTHER" else 1 for x in train_lines])
# train_labels = [0 if x[1] == "OTHER" else 1 for x in train_lines]
dev_sentences = [x[0] for x in dev_lines]
dev_labels = to_categorical([0 if x[1] == "OTHER" else 1 for x in dev_lines])
# dev_labels = [0 if x[1] == "OTHER" else 1 for x in dev_lines]
# +
# print(train_labels)
# -
model = Sequential([
LSTM(64, recurrent_dropout = 0.5, dropout = 0.5, activation = 'relu', input_shape=(nb_sequence_length, nb_embedding_dims)),
Dense(32, activation = 'relu'),
Dropout(0.2),
Dense(2, activation = 'softmax')
])
model.compile(loss='sparse_categorical_crossentropy', optimizer='nadam', metrics = ['accuracy'])
# +
# model = Sequential([
# Conv1D(128, kernel_size = 3, padding = 'valid', input_shape=(nb_sequence_length, nb_embedding_dims), activation = 'relu'),
# MaxPooling1D(5),
# Flatten(),
# Dense(64, activation = 'relu'),
# Dropout(0.2),
# Dense(2, activation = 'softmax')
# ])
# model.compile(loss='categorical_crossentropy', optimizer='nadam', metrics = ['accuracy'])
# -
filter_sizes = (3, 4, 5)
model_input = Input(shape = (nb_sequence_length, nb_embedding_dims))
model_layers = Dropout(0.8)(model_input)
conv_blocks = []
for sz in filter_sizes:
conv = Conv1D(filters = 100,
kernel_size = sz,
padding = "valid",
activation = "relu",
strides = 1,
kernel_regularizer = regularizers.l2(0.0001))(model_layers)
conv = GlobalMaxPooling1D()(conv)
# conv = Flatten()(conv)
conv_blocks.append(conv)
model_concatenated = Concatenate()(conv_blocks)
model_concatenated = Dropout(0.8)(model_concatenated)
model_concatenated = Dense(64, activation = "relu")(model_concatenated)
model_output = Dense(2, activation = "softmax")(model_concatenated)
model = Model(model_input, model_output)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics = ['accuracy'])
samples_per_epoch = len(train_sentences)
epochs = 50
batch_size = 32
steps_per_epoch = math.ceil(samples_per_epoch / batch_size)
model.fit_generator(
my_generator(train_sentences, train_labels, batch_size),
steps_per_epoch=steps_per_epoch, nb_epoch=epochs,
validation_data = my_generator(dev_sentences, dev_labels, batch_size),
validation_steps = math.ceil(len(dev_sentences) / batch_size)
)
|
language_model/TextClassification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### <span style="color:green"> Exercise: Creator </span>
#
# Complete the code below to print the creator of each item.
# You will need to browse the [data](https://api.nb.no/catalog/v1/items?digitalAccessibleOnly=true&size=3&filter=mediatype:bøker&q=Bing,Jon).
#
# This exercise is a bit vague. The `creators` field has a list containing one or more creators.
# Should we print the list, which includes Python syntax, or make it human readable?
# If we encounter such vague specifications when programming, we must *stop* and clarify them before continuing. IT projects often go awry because of vague or incorrect specifications.
# +
import requests
import json
URL = "https://api.nb.no/catalog/v1/items?digitalAccessibleOnly=true&size=3&filter=mediatype:bøker&q=Bing,Jon"
data = requests.get(URL).json()
embedded = data['_embedded']
items = embedded['items']
for item in items:
metadata = item['metadata']
print("Item title:", metadata['title'])
creators = metadata['creators']
# version 1, print the list
print('Creators:', creators)
# version 2, looping over the list
for creator in creators:
print('Creator:', creator)
print() # insert empty line
# -
# ### <span style="color:green"> Exercise: Presentation and URN </span>
#
# The field `presentation` contains a link to the full text.
# Complete the code below to print the `presentation` URL and the 'URN' of each item.
#
# You will need to browse the [data](https://api.nb.no/catalog/v1/items?digitalAccessibleOnly=true&size=3&filter=mediatype:bøker&q=Bing,Jon).
#
# The `presentation` URL is located below `_links`, while the `urn` is in `metadata`.
# +
import requests
import json
URL = "https://api.nb.no/catalog/v1/items?digitalAccessibleOnly=true&size=3&filter=mediatype:bøker&q=Bing,Jon"
data = requests.get(URL).json()
embedded = data['_embedded']
items = embedded['items']
for item in items:
metadata = item['metadata']
print("Item title:", metadata['title'])
links = item['_links']
presentation = links['presentation']
href = presentation['href']
print("Presentation URL:", href)
identifiers = metadata['identifiers']
urn = identifiers['urn']
print(urn)
print()
|
solutions/01_intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
#Plot Tools
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
#Model Building
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
import sklearn
import keras
from keras.models import Sequential
from keras.layers import InputLayer,Dense
import tensorflow as tf
#Model Validation
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# ## Load Dataset
data = pd.read_csv(r"E:\ExcelR Assignment\Assignment 16 - Neural Networks\forestfires.csv")
data.head()
# ### Drop month and day as we already have one hot encoded values
data = data.drop(['month','day','daysun','monthdec'],axis=1)
data.head(2)
data.dtypes
# ## Data Analysis & Data Visualization
import warnings
warnings.filterwarnings("ignore")
fig, axes = plt.subplots(3, 3, figsize=(20, 15))
fig.suptitle('Univariate Analysis',fontsize=20)
sns.distplot(data['FFMC'],ax=axes[0,0],color='indigo')
sns.distplot(data['DMC'],ax=axes[0,1],color='orange')
sns.distplot(data['DC'],ax=axes[0,2],color='olive')
sns.distplot(data['ISI'],ax=axes[1,0],color='indigo')
sns.distplot(data['temp'],ax=axes[1,1],color='orange')
sns.distplot(data['RH'],ax=axes[1,2],color='olive')
sns.distplot(data['wind'],ax=axes[2,0],color='indigo')
sns.distplot(data['rain'],ax=axes[2,1],color='orange')
sns.distplot(data['area'],ax=axes[2,2],color='olive')
# ### Inferences:
#
# 1. Left Skewness : DMC | ISI | RH | rain | area
# 2. Right Skewness : FFMC | DC
# 3. temp | wind seems to have normally distribution
label_encoder = LabelEncoder()
data['size_category'] = label_encoder.fit_transform(data['size_category'])
data.head()
# #### So small = 1 and large = 0 as per Label Encoding
# ## Splitting the variables
X = data.iloc[:,0:-1]
Y = data.iloc[:,-1]
data['size_category'].value_counts()
# ## Up-sampling as the Target Variable is not balanced
# +
from imblearn.combine import SMOTETomek
# Performing Oversampling method to handle imbalanced data
smk = SMOTETomek(random_state=42)
X_res,y_res = smk.fit_resample(X,Y)
# -
(X.shape,Y.shape),(X_res.shape,y_res.shape)
# ### Create Train & Validation Set
X_train,X_test,y_train,y_test = train_test_split(X_res,y_res,stratify=y_res,random_state=42,test_size=0.3)
# ### Standardizing the first few columns till area
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# ## Define Architecture
input_neurons = X_train.shape[1]
output_neurons = 1
Hidden_Layers = 2
hidden_layer_1 = 20
hidden_layer_2 = 8
model = Sequential()
model.add(InputLayer(input_shape=(input_neurons,)))
model.add(Dense(units=hidden_layer_1,activation='relu'))
model.add(Dense(units=hidden_layer_2,activation='relu'))
model.add(Dense(units=output_neurons,activation='sigmoid'))
model.summary()
model.compile(loss='binary_crossentropy',optimizer='Adam',metrics=['accuracy'])
# ## Train the model
model_history = model.fit(X_train,y_train,validation_data=(X_test,y_test),epochs=100)
# ## Evaluate Model Performance
# #### Summarizing history for loss
plt.plot(model_history.history['loss'])
plt.plot(model_history.history['val_loss'])
plt.title('model_loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','validation'],loc='upper left')
plt.show()
# #### Summarizing history for accuracy
plt.plot(model_history.history['accuracy'])
plt.plot(model_history.history['val_accuracy'])
plt.title('model_accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train','validation'],loc='upper left')
plt.show()
# ## Model Performance on Validation set
prediction = model.predict_classes(X_test)
accuracy_score(y_test,prediction)
# ### So our model has good Test Accuracy
|
Assignment 16 Part 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="lckWPMSNtmmn"
# https://openreview.net/pdf?id=SkZxCk-0Z
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="x_3L4cW4Ywou"
import parser
import tensorflow as tf
tf.enable_eager_execution()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="HiiMXz-oY7PZ"
def read_data(fname):
"""
Reads the data files.
"""
with open(fname, 'r') as f:
data = f.read()
data = data.split('\n')
new_data = []
for d in data[:-1]:
# print('\r {}'.format(d), end='', flush=True)
a, b, e, _, _, _ = tuple(d.split(','))
new_data.append([a, b, int(e)])
return new_data
def batch_data(data, batch_size):
n = len(data)
data = list(zip(*data[0:-1])) # transpose the data
for i in range(n//batch_size-1):
A = data[0][i*batch_size:(i+1)*batch_size]
B = data[1][i*batch_size:(i+1)*batch_size]
E = data[2][i*batch_size:(i+1)*batch_size]
yield A, B, E
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 374} colab_type="code" executionInfo={"elapsed": 707, "status": "ok", "timestamp": 1528060779089, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-bOBnzCbRWgs/AAAAAAAAAAI/AAAAAAAAABw/Sd6a7MwcUDU/s50-c-k-no/photo.jpg", "userId": "112995735588747661471"}, "user_tz": -720} id="bI2KLSu6x3Qt" outputId="9f81bef4-fcd7-4577-f4c9-548d04220628"
data = read_data('../logical_entailment_dataset/data/train.txt')
A, B, E = next(batch_data(data, 10))
A, B, E
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="mETRZKx-eQvz"
class Parser():
def __init__(self, language):
self.language = language
self.parser = parser.Parser(language)
self.vocabulary = {op: i for i, op in enumerate(language.symbols)}
def __call__(self, s):
parse_result = self.parser.parse(s)
ops = [self.vocabulary[op.decode("utf-8")] for op in parse_result.ops]
return ops, parse_result.inputs
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 357} colab_type="code" executionInfo={"elapsed": 737, "status": "ok", "timestamp": 1528060782711, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-bOBnzCbRWgs/AAAAAAAAAAI/AAAAAAAAABw/Sd6a7MwcUDU/s50-c-k-no/photo.jpg", "userId": "112995735588747661471"}, "user_tz": -720} id="g9Hf9RzTd2I-" outputId="6246582c-9a74-45da-ebb8-5f8deabbc9ca"
prop_parser = Parser(parser.propositional_language())
tree = prop_parser('((m>m)&(((m>m)>(m>m))&((m>m)>(m|m))))')
tree
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="cxCd0T8EV0cr"
d_world = 30
n_worlds=24
d_embed = 50
batch_size = 10
n_ops = len(prop_parser.language.symbols)
# -
n_ops
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="4i0p8m_0v6ux"
class Sat3Cell():
"""
Real valued evaluation of satisfiability.
Given a real valued truth assignment, aka the world you are in,
check if it satisfies the given equation.
"""
def __init__(self, d_world, n_ops, d_embed):
num_units = d_embed
self.op_embeddings = tf.get_variable(shape=(n_ops, d_world, num_units), dtype=tf.float32, name='operation_embeddings')
self.W4 = tf.get_variable(shape=(n_ops, 2*d_embed, num_units), dtype=tf.float32, name='W4')
self.b4 = tf.get_variable(shape=(n_ops, num_units), dtype=tf.float32, name='b4')
def __call__(self, w, op, l=None, r=None, scope=None):
"""
Args:
w (tf.tensor): [1, d_world]
TODO op (list):
"""
# TODO change so __call__ can recieve a batch.
# then bundle all embed/matmul calls
# but op will be varing length. need to stack them!?
with tf.variable_scope(scope or type(self).__name__):
# nullary ops
if l is None and r is None:
# look up their embeddings
h = tf.matmul(w, self.op_embeddings[op])
else:
# unary and binary ops
if l is not None and r is None:
r = tf.zeros_like(l) # just fake it
x = tf.concat([l, r], axis=1)
h = tf.matmul(x, self.W4[op]) + self.b4[op]
return tf.nn.l2_normalize(h, axis=1)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1646, "status": "ok", "timestamp": 1528060790466, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-bOBnzCbRWgs/AAAAAAAAAAI/AAAAAAAAABw/Sd6a7MwcUDU/s50-c-k-no/photo.jpg", "userId": "112995735588747661471"}, "user_tz": -720} id="IPF7b7gNvtZx" outputId="cf448fcb-68e0-447c-83cc-f07485dd08e4"
sat3 = Sat3Cell(d_world, n_ops, d_embed)
w = tf.random_normal([1, d_world])
l = tf.random_normal((1, d_embed))
r = tf.random_normal((1, d_embed))
h = sat3(w, 0)
print(h.shape)
h = sat3(w, 12, l, r)
print(h.shape)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="SXi2C4JLsmOu"
class TreeNN():
def __init__(self, cell, parser):
self.cell = cell
self.parser = parser
# !? what about learning to parse the inputs into a tree!?
def __call__(self, w, s):
"""
Because each parse will be different!?
Args:
w: a world
s: a string
Returns: (1, n)
"""
# NOTE Can only handle a single element of a batch at a time
tree = self.parser(s)
return self.apply(tree, [])
def apply(self, tree, results, i=0):
"""
Applies self.cell in a recursive manner.
Args:
tree (tuple): (ops, args)
ops (list): nodes in depth first order
args (list): the children of ops in depth first order
"""
ops, args = tree
# if the current node has children, fetch them from results
l = None
r = None
if len(args[0]) == 1:
l = results[i+args[0][0]]
elif len(args[0]) == 2:
l = results[i+args[0][0]]
r = results[i+args[0][1]]
if len(tree[1]) == 1:
return self.cell(w, ops[i], l, r)
else:
results.append(self.cell(w, ops[i], l, r))
tree = (ops, args[1:])
return self.apply(tree, results, i+1)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 221} colab_type="code" executionInfo={"elapsed": 968, "status": "ok", "timestamp": 1528060794097, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-bOBnzCbRWgs/AAAAAAAAAAI/AAAAAAAAABw/Sd6a7MwcUDU/s50-c-k-no/photo.jpg", "userId": "112995735588747661471"}, "user_tz": -720} id="gWvMnNwPvr1F" outputId="b5671bde-1c65-4d4f-a914-f31b40cffddd"
treenn = TreeNN(Sat3Cell(d_world, n_ops, d_embed), prop_parser)
treenn(w, '((m>m)&(((m>m)>(m>m))&((m>m)>(m|m))))')
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Mu58VBNtly-O"
class PossibleWorlds():
"""
A NN designed specifically for predicting entailment.
"""
def __init__(self, encoder, num_units, n_worlds, d_world):
self.encoder = encoder
self.n_worlds = n_worlds
self.worlds = tf.get_variable(shape=(n_worlds, d_world), dtype=tf.float32, name='worlds')
self.dense = tf.keras.layers.Dense(num_units)
def inner(self, a, b):
"""
Convolve over possible worlds.
For each random direction, do !??!
"""
p = tf.constant(1.0, dtype=tf.float32)
for i in range(self.n_worlds):
x = tf.concat([self.encoder(self.worlds[i:i+1], a),
self.encoder(self.worlds[i:i+1], b)], axis=1)
p *= self.dense(x) # in the paper this isnt actually a dense layer....
return p
def __call__(self, A, B):
"""
For each element of a batch.
"""
return tf.concat([self.inner(a, b) for a, b in zip(A, B)], axis=0)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="h3ZtKa-gu-re"
possibleworldsnet = PossibleWorlds(
encoder=TreeNN(Sat3Cell(d_world, n_ops, d_embed), prop_parser),
num_units=1,
n_worlds=n_worlds,
d_world=d_world
)
variables = (possibleworldsnet.dense.variables +
[possibleworldsnet.encoder.cell.b4,
possibleworldsnet.encoder.cell.op_embeddings,
possibleworldsnet.encoder.cell.W4])
opt = tf.train.AdamOptimizer()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="vdu18mMYvzzp"
def gradients(A, B, E):
with tf.GradientTape() as tape:
y = possibleworldsnet(A, B)
loss = tf.losses.sigmoid_cross_entropy(multi_class_labels=tf.constant(E, dtype=tf.float32, shape=(batch_size, 1)),
logits=y)
step = tf.train.get_or_create_global_step().numpy()
print('\rstep: {} loss {}'.format(step, tf.reduce_mean(loss)), end='', flush=True)
return tape.gradient(loss, variables)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="aka3mR6M5uoD" outputId="ece98d6a-52dc-439e-f821-9b6baf39fbe3"
for A, B, E in batch_data(data, batch_size):
gnvs = zip(gradients(A, B, E), variables)
opt.apply_gradients(gnvs, global_step=tf.train.get_or_create_global_step())
# + [markdown] colab_type="text" id="rXB2xksk9nJo"
# Argh. It is soo slow... A problem for another day.
|
entailment/possibleworldnet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''base'': conda)'
# language: python
# name: python397jvsc74a57bd0f16b4f9c118091e521975b08f7e11c780453654917dc239e484e68cca92583fe
# ---
session.close()
# %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
# create engine to hawaii.sqlite
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
# reflect an existing database into a new model
Base = automap_base()
Base.prepare(engine, reflect=True)
Base.classes.keys()
# reflect the tables
# View all of the classes that automap found
# equivalent of writing the classes by hand
Measurement = Base.classes.measurement
Station = Base.classes.station
# +
# Save references to each table
# -
# Create our session (link) from Python to the DB
session = Session(engine)
# # Exploratory Precipitation Analysis
# Find the most recent date in the data set.
recent_date = session.query(Measurement.date).order_by(Measurement.date.desc()).first()
recent_date
# 365 days later
year_later = dt.date(2017, 8, 23) - dt.timedelta(days=365)
year_later
# +
# date from recent_date 2017-8-23 until 2016-8-23
precips = session.query(Measurement.prcp, Measurement.date).filter(Measurement.date >= dt.datetime(2016, 8, 23)).all()
precipitation = []
date = []
# precip[0] is precipitation, precip[1] is date
for precip in precips:
# precip_dict = {}
# precip_dict["date"]: "precip[1]"
# precip_dict["precipitation"] = precip[0]
# print(precip[1])
date.append(precip[1])
precipitation.append(precip[0])
precip_date_df = pd.DataFrame(list(zip(date, precipitation)), columns=["Date", "Precipitation"])
# Starting from the most recent data point in the database.
# Calculate the date one year from the last date in data set.
# Perform a query to retrieve the data and precipitation scores
# Save the query results as a Pandas DataFrame
# precip_date_df = pd.DataFrame(data=precip_dict)
# # Sort the dataframe by date and drop nulls
precip_date_df = precip_date_df.sort_values("Date")
precip_date_df.dropna()
# am i grabbing extra data? Should be 365 rows but i have 2015
# -
# didn't end up being able to figure out how to implement
# x-axis labels instead of all dates
x_labels = ["2016-08-23", "2016-10-01", "2016-11-09", "2016-12-19", "2017-01-28", "2017-03-09", "2017-04-18", "2017-05-31", "2017-07-01"]
x_labels
# +
# Use DataFrame.plot() in order to create a bar chart of the data
fig = precip_date_df.plot(x="Date", y="Precipitation", subplots=True, sharex=True, kind="bar", rot=45)
# Set a title for the chart
plt.title("Precipitation Data")
plt.xlabel("Date")
plt.ylabel("Inches")
plt.ylim(0, max(precip_date_df["Precipitation"])+10)
# Cannot figure out how to only show a subset of xlabels so it looks ugly
plt.show()
# +
# matplotlib method - better but i still am having an issue somewhere
x_axis = np.arange(len(precip_date_df))
plt.figure(figsize=(20,3))
plt.bar(x_axis, precip_date_df["Precipitation"], color='r', alpha=0.5, align="center")
# plt.xticks(tick_locations, precip_date_df["Date"], rotation="vertical")
# plt.xticks(np.arange(9), x_labels)
# Set x and y limits
plt.xlim(-0.75, len(x_axis))
plt.ylim(0, max(precip_date_df["Precipitation"]))
# Set a Title and labels
plt.title("Precipitation")
plt.xlabel("Date")
plt.ylabel("Precipitation")
# Save our graph and show the grap
# plt.tight_layout()
plt.show()
# -
# Use Pandas to calcualte the summary statistics for the precipitation data
precip_date_df.describe()
# # Exploratory Station Analysis
# Design a query to calculate the total number stations in the dataset
session.query(func.count(Station.station)).all()
# Design a query to find the most active stations (i.e. what stations have the most rows?)
# List the stations and the counts in descending order.
# query the stations and number of times they appear, then group by station and desc order by the counts
stations = session.query(Measurement.station, func.count(Measurement.station))\
.group_by(Measurement.station).order_by(func.count(Measurement.station).desc()).all()
for station in stations:
print(station)
# Using the most active station id from the previous query, calculate the lowest, highest, and average temperature. (tobs column)
active_station = stations[0][0]
session.query(func.min(Measurement.tobs), func.max(Measurement.tobs), func.avg(Measurement.tobs))\
.filter(Measurement.station == active_station).all()
# +
# Using the most active station id
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
# date from recent_date 2017-8-23 until 2016-8-23
temp_data = session.query(Measurement.date, Measurement.tobs).filter(Measurement.date >= "2016-08-23")\
.filter(Measurement.station == active_station).all()
# make df
temp_df = pd.DataFrame(temp_data)
# plot histogram
temp_df.plot.hist(bins=12)
# -
# # Close session
# Close Session
session.close()
|
.ipynb_checkpoints/climate_starter-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pyspark
# ---
from pyspark.sql.session import *
from pyspark import __version__
if __version__ < '2.4.0':
raise Exception("Require version 2.4.0 or higher")
# Note: gs requires the GCS connector JARs are present on the class path
session = SparkSession.builder.getOrCreate()
sc = session.sparkContext
sc.stop()
session.stop()
session = (SparkSession.builder
.appName("debuggingIsCool")
.config("spark.executor.instances", "40")
.config("spark.driver.memoryOverhead", "0.25")
.config("spark.files", "gs://boo-stuff/alt_worker_daemon.py")
.config("spark.pyfiles", "gs://boo-stuff/remote-pdb-1.2.0.zip")
.config("spark.python.use.daemon", "true")
.config("spark.python.daemon.module", "alt_worker_daemon.py")
).getOrCreate()
sc = session.sparkContext
conf = sc.getConf()
conf.get("spark.python.daemon.module")
rdd = sc.parallelize(range(200), 10)
def wait_fun(elem):
import time
time.sleep(0.5)
return 1
rdd.map(wait_fun).count()
sc.stop()
session.stop()
|
Debugging is Fun1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Multi-Layer Perceptron
# - Dense layer, fully connected layer....
# - Hidden layer와 그 안에 activation을 추가한다.
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
print("Current Tensorflow is [%s]" % (tf.__version__))
print("Packages Loaded.")
mnist = input_data.read_data_sets('data/', one_hot=True)
# ##### 1. Parameter Setting
# +
n_input = 784
n_hidden_1 = 256
n_hidden_2 = 128
n_classes = 10
# set input and output
x = tf.placeholder(dtype='float', shape=[None, n_input])
y = tf.placeholder(dtype='float', shape=[None, n_classes])
# set network parameters(weights, biases)
stddev = 0.1
weights = {
# 가중치의 초기값은
# 평균 : 0(default), 표준편차 : 0.1 인 정규분포에서 random으로 뽑는다
# hidden layer1의 노드 수는 256개, hidden layer2의 노드 수는 128개
# out layer의 노드 수 = label 갯수 = 10개(0~9, 숫자 10개)
'h1' : tf.Variable(initial_value=tf.random_normal(shape=[n_input, n_hidden_1],stddev=stddev)), # 784 x 256 matrix
'h2' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_1, n_hidden_2], stddev=stddev)), # 256 x 128 matrix
'out' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_2, n_classes], stddev=stddev)), # 128 x 10 matrix
}
biases = {
'b1' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_1])), # 256개
'b2' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_2])),
'out' : tf.Variable(initial_value=tf.random_normal(shape=[n_classes])),
}
print("Network Ready!!!")
# -
# ##### 2. Define Graph
# +
# model
def multilayer_perceptron(_x, _weights, _biases):
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(_x, _weights['h1']), _biases['b1'])) # 1번째 layer 통과
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, _weights['h2']), _biases['b2'])) # 2번째 layer 통과
# return은 logit을 뽑아야 한다.(softmax 취하기 전 형태)
# softmax취해서 return하면 성능 떨어짐...
return (tf.matmul(layer_2, _weights['out']) + _biases['out'])
# prediction
pred = multilayer_perceptron(x, weights, biases)
# Loss and Optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(labels=y, logits=pred))
optimizer = tf.train.AdamOptimizer(learning_rate=0.001).minimize(cost)
correct = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct, 'float'))
# Initialize
init = tf.global_variables_initializer()
print("Function Ready!!!")
# -
# ##### 3. Run
# +
training_epochs = 20
batch_size = 100
display_step = 4
# Launch the Graph
sess = tf.Session()
sess.run(init)
# Optimize
for epoch in range(training_epochs):
avg_cost = 0
total_batch = int(mnist.train.num_examples / batch_size)
# Iteration
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size=batch_size)
feeds = {x: batch_xs, y: batch_ys}
sess.run(optimizer, feed_dict=feeds)
avg_cost += sess.run(cost, feed_dict=feeds)
avg_cost = avg_cost / total_batch
# Display
if (epoch+1) % display_step == 0:
print("Epoch: %03d/%03d cost: %.9f" % (epoch, training_epochs, avg_cost))
feeds = {x: batch_xs, y: batch_ys}
train_acc = sess.run(accuracy, feed_dict=feeds)
print("Train Accuracy: %.3f" % (train_acc))
feeds = {x: mnist.test.images, y: mnist.test.labels}
test_acc = sess.run(accuracy, feed_dict=feeds)
print("Test Accuracy: %.3f" % (test_acc))
print("Optimization Finished!!")
# -
# -----
# -----
# ### Layer를 하나 더 추가해보자!!
# +
n_input = 784
n_hidden_1 = 256
n_hidden_2 = 128
n_hidden_3 = 64
n_classes = 10
# set input and output
x = tf.placeholder(dtype='float', shape=[None, n_input])
y = tf.placeholder(dtype='float', shape=[None, n_classes])
# set network parameters(weights, biases)
stddev = 0.1
weights = {
'h1' : tf.Variable(initial_value=tf.random_normal(shape=[n_input, n_hidden_1],stddev=stddev)),
'h2' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_1, n_hidden_2], stddev=stddev)),
'h3' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_2, n_hidden_3], stddev=stddev)),
'out' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_3, n_classes], stddev=stddev)),
}
biases = {
'b1' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_1])),
'b2' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_2])),
'b3' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_3])),
'out' : tf.Variable(initial_value=tf.random_normal(shape=[n_classes])),
}
print("Network Ready!!!")
# -
# ##### 2. Define Graph
# +
# model
def multilayer_perceptron(_x, _weights, _biases):
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(_x, _weights['h1']), _biases['b1']))
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, _weights['h2']), _biases['b2']))
layer_3 = tf.nn.sigmoid(tf.add(tf.matmul(layer_2, _weights['h3']), _biases['b3']))
# return은 logit을 뽑아야 한다.(softmax 취하기 전 형태)
# softmax취해서 return하면 성능 떨어짐...
return (tf.matmul(layer_3, _weights['out']) + _biases['out'])
# prediction
pred = multilayer_perceptron(x, weights, biases)
# Loss and Optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(labels=y, logits=pred))
optimizer = tf.train.AdamOptimizer(learning_rate=0.001).minimize(cost)
correct = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct, 'float'))
# Initialize
init = tf.global_variables_initializer()
print("Function Ready!!!")
# -
# ##### 3. Run
# +
training_epochs = 20
batch_size = 100
display_step = 4
# Launch the Graph
sess = tf.Session()
sess.run(init)
# Optimize
for epoch in range(training_epochs):
avg_cost = 0
total_batch = int(mnist.train.num_examples / batch_size)
# Iteration
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size=batch_size)
feeds = {x: batch_xs, y: batch_ys}
sess.run(optimizer, feed_dict=feeds)
avg_cost += sess.run(cost, feed_dict=feeds)
avg_cost = avg_cost / total_batch
# Display
if (epoch+1) % display_step == 0:
print("Epoch: %03d/%03d cost: %.9f" % (epoch, training_epochs, avg_cost))
feeds = {x: batch_xs, y: batch_ys}
train_acc = sess.run(accuracy, feed_dict=feeds)
print("Train Accuracy: %.3f" % (train_acc))
feeds = {x: mnist.test.images, y: mnist.test.labels}
test_acc = sess.run(accuracy, feed_dict=feeds)
print("Test Accuracy: %.3f" % (test_acc))
print("Optimization Finished!!")
# -
# ---
# ---
# ### Activation Funtion을 ReLU로 바꿔보자!!
# +
n_input = 784
n_hidden_1 = 256
n_hidden_2 = 128
n_hidden_3 = 64
n_classes = 10
# set input and output
x = tf.placeholder(dtype='float', shape=[None, n_input])
y = tf.placeholder(dtype='float', shape=[None, n_classes])
# set network parameters(weights, biases)
stddev = 0.1
weights = {
'h1' : tf.Variable(initial_value=tf.random_normal(shape=[n_input, n_hidden_1],stddev=stddev)),
'h2' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_1, n_hidden_2], stddev=stddev)),
'h3' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_2, n_hidden_3], stddev=stddev)),
'out' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_3, n_classes], stddev=stddev)),
}
biases = {
'b1' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_1])),
'b2' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_2])),
'b3' : tf.Variable(initial_value=tf.random_normal(shape=[n_hidden_3])),
'out' : tf.Variable(initial_value=tf.random_normal(shape=[n_classes])),
}
print("Network Ready!!!")
# -
# ##### 2. Define Graph
# +
# model
def multilayer_perceptron(_x, _weights, _biases):
layer_1 = tf.nn.relu(tf.add(tf.matmul(_x, _weights['h1']), _biases['b1']))
layer_2 = tf.nn.relu(tf.add(tf.matmul(layer_1, _weights['h2']), _biases['b2']))
layer_3 = tf.nn.relu(tf.add(tf.matmul(layer_2, _weights['h3']), _biases['b3']))
# return은 logit을 뽑아야 한다.(softmax 취하기 전 형태)
# softmax취해서 return하면 성능 떨어짐...
return (tf.matmul(layer_3, _weights['out']) + _biases['out'])
# prediction
pred = multilayer_perceptron(x, weights, biases)
# Loss and Optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(labels=y, logits=pred))
optimizer = tf.train.AdamOptimizer(learning_rate=0.001).minimize(cost)
correct = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct, 'float'))
# Initialize
init = tf.global_variables_initializer()
print("Function Ready!!!")
# -
# ##### 3. Run
# +
training_epochs = 20
batch_size = 100
display_step = 4
# Launch the Graph
sess = tf.Session()
sess.run(init)
# Optimize
for epoch in range(training_epochs):
avg_cost = 0
total_batch = int(mnist.train.num_examples / batch_size)
# Iteration
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size=batch_size)
feeds = {x: batch_xs, y: batch_ys}
sess.run(optimizer, feed_dict=feeds)
avg_cost += sess.run(cost, feed_dict=feeds)
avg_cost = avg_cost / total_batch
# Display
if (epoch+1) % display_step == 0:
print("Epoch: %03d/%03d cost: %.9f" % (epoch, training_epochs, avg_cost))
feeds = {x: batch_xs, y: batch_ys}
train_acc = sess.run(accuracy, feed_dict=feeds)
print("Train Accuracy: %.3f" % (train_acc))
feeds = {x: mnist.test.images, y: mnist.test.labels}
test_acc = sess.run(accuracy, feed_dict=feeds)
print("Test Accuracy: %.3f" % (test_acc))
print("Optimization Finished!!")
# -
# 다른거 손대지 않고 Graph만 바꿔주면 된다는 것이 TensorFlow의 장점!
|
5.MLP_NumberClassifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Test-bed notebook to find taxonomic groupings for pheromoneFinder
# ### Authored by <NAME>, Murray and Gaudet Labs, Harvard
# Modified to run on my sager since my xps15 died, installed the following packages with potential for version issues.
# 1. biopython 1.78
# 2. dendropy 4.5.1
# +
##This is to test code snippets for the pheromoneFinder codebase
import sys
import os
from os import listdir
from os.path import isfile, join
import traceback
import re
from Bio import SeqIO
from Bio.Seq import Seq
##from Bio.Alphabet import IUPAC ##deprecated in biopython 1.78
from collections import defaultdict
from Bio.SeqUtils.ProtParam import ProteinAnalysis
from Bio.Blast.Applications import NcbiblastxCommandline
from Bio.Blast.Applications import NcbiblastnCommandline
from Bio.Blast import NCBIXML
import xml.etree.ElementTree as ET
import dendropy
import numpy as np
import seaborn as sns
from pylab import *
import matplotlib.pyplot as plt
import csv
import math
import operator
from operator import attrgetter
from pprint import pprint
data_dir = os.path.join("..", "..", "..", "pheromoneFinder_data", "Cluster")
figure_dir = os.path.join(data_dir, "figures")
#Setting matplotlib to be inline
# %matplotlib inline
# -
tree_dir = os.path.join(data_dir, "tree")
#Printing file list
fileList = [f for f in os.listdir(tree_dir) if os.path.isfile(os.path.join(tree_dir, f))]
print("List of files:")
print("\n".join(fileList))
##Importing Newick phylogenetic trees
phyloTree = dendropy.Tree.get(file=open(os.path.join(tree_dir, "332_2408OGs_timetree_reltime.nwk"), 'r'), schema="newick")
# ### Code blocks below are to calculate and plot the pair-wise distances of species
# These are using the dendropy.<i>Tree</i>.pdc(), to read in the Newick reltime tree from the Shen XX, Cell (2018) paper to identify clades of species that have diverged within a time horizon.
# > Written originally in Windows and moved to WSL Ubuntu 18.04 for further work and testing to move to Odyssey.
# +
distMatrix = np.zeros((len(phyloTree.taxon_namespace), len(phyloTree.taxon_namespace)))
pdc = phyloTree.phylogenetic_distance_matrix()
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
#print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
distMatrix[i,j] = pdc(t1, t2);
#print(distMatrix)
print(phyloTree.taxon_namespace[0].label, "\t", phyloTree.taxon_namespace[-1].label, "\n",
pdc.distance(phyloTree.taxon_namespace[0], phyloTree.taxon_namespace[-1]))
print(len(distMatrix.flatten()))
mask = np.zeros_like(distMatrix, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
#print(pdc.distance(phyloTree.taxon_namespace[:], phyloTree.taxon_namespace[:]))
fig1 = figure(figsize=(16,6))
ax1 = subplot(121)
#ax = sns.heatmap(distMatrix[0:99, 0:99], yticklabels=phyloTree.taxon_namespace[0:99])
ax1 = sns.heatmap(distMatrix, mask=mask, cmap="Greys_r")
ax2 = subplot(122)
plot = list(filter(lambda a: a != 0, distMatrix.flatten()))
ax2 = sns.histplot(plot, kde=True)
##ax2 = sns.distplot(plot, kde=True, rug=True) ##distplot is being deprecated and replacing with axis-level histplot
pairwise_dist_title = " ".join([str(len(plot)), "pairwise distances in tree"])
plt.title(pairwise_dist_title);
#savefig(os.path.join(figure_dir, "Shen_332yeast_pairwise-dist_reltime.pdf"), dpi=300, transparent=True)
# -
# #### Code-base to identify the time-horizon for the conservation of a-factor-like (lipidated) fungal pheromones
# I am going to assume that the "conservation" here corresponds to pheromones that can be recognized from one species to another. The best examples I have are S. cerevisiae and K. lactis, within the Saccharomycetaceae and the 5 species in the Yarrowia clade.
##Identifying the evolutionary time horizon for the divergence of all species in the Yarrowia clade.
yarrowiaPair = []
yarrowiaDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("Yarrowia" in t1.label) and ("Yarrowia" in t2.label)) or (("Yarrowia" in t2.label) and ("Yarrowia" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
yarrowiaPair.append([t1.label, t2.label])
yarrowiaDist.append(pdc(t1, t2))
index, value = max(enumerate(yarrowiaDist), key=operator.itemgetter(1))
print("Max. divergence time of Yarrowia is between: ", yarrowiaPair[index], "\t of time: ", yarrowiaDist[index])
print(max(yarrowiaDist))
##Identifying the evolutionary divergence time between the Saccharomycetaceae (K. lactis) and Saccharomycodaceae (Hanseniaspora)
speciesPair = []
speciesDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("lactis" in t1.label) and ("Hanseniaspora" in t2.label)) or (("lactis" in t2.label) and ("Hanseniaspora" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
speciesPair.append([t1.label, t2.label])
speciesDist.append(pdc(t1, t2))
index, value = max(enumerate(speciesDist), key=operator.itemgetter(1))
print("Max. divergence time of Saccharomyces MFA1/2-like is between: ", speciesPair[index], "\t of time: ", speciesDist[index])
print(max(speciesDist))
##Identifying the evolutionary time horizon for the divergence of all species in the Saccharomycetaceae clade (tested as between S. cerevisiae and K. lactis).
speciesPair = []
speciesDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("cere" in t1.label) and ("lactis" in t2.label)) or (("cere" in t2.label) and ("lactis" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
speciesPair.append([t1.label, t2.label])
speciesDist.append(pdc(t1, t2))
index, value = max(enumerate(speciesDist), key=operator.itemgetter(1))
print("Max. divergence time of Saccharomyces MFA1/2-like is between: ", speciesPair[index], "\t of time: ", speciesDist[index])
print(max(speciesDist))
##Identifying the distance of closest neighbor to the Yarrowia lineage. As seen it is already more than the value expected to share pheromones; though the divergence time
##is not a hard limit that arises naturally.
speciesPair = []
speciesDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("Nadsonia" in t1.label) and ("Yarrowia" in t2.label)) or (("Nadsonia" in t2.label) and ("Yarrowia" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
speciesPair.append([t1.label, t2.label])
speciesDist.append(pdc(t1, t2))
index, value = max(enumerate(speciesDist), key=operator.itemgetter(1))
print("Max. divergence time of: ", speciesPair[index], "\t of time: ", speciesDist[index])
print(max(speciesDist))
# #### Code-block to identify the time-horizon for the conservation of the regulation of mating genes.
# This is to identify the time-horizon of evolutionary conservation of the regulation of mating genes. The Sorrells TR, Nature (2015) [doi: 10.1038/nature14613] paper inspired the basis of looking for an evolutionary relationship to the motifs for mating regulatory TFs, specifically STE12 in the Saccharomyces clade.
##Identifying the evolutionary time horizon for the divergence of all species in the Saccharomycetaceae clade (tested as between S. cerevisiae and K. lactis).
speciesPair = []
speciesDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("cerevisiae" in t1.label) and ("polyspora" in t2.label)) or (("cerevisiae" in t2.label) and ("polyspora" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
speciesPair.append([t1.label, t2.label])
speciesDist.append(pdc(t1, t2))
index, value = max(enumerate(speciesDist), key=operator.itemgetter(1))
print("Max. divergence time of first clade (Saccharomyces) described in Sorrells et al is between: ", speciesPair[index], "\t of time: ", speciesDist[index])
print(max(speciesDist))
##Identifying the evolutionary time horizon for the divergence of all species in the Saccharomycetaceae clade (tested as between S. cerevisiae and K. lactis).
speciesPair = []
speciesDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("kluyveri" in t1.label) and ("lactis" in t2.label)) or (("kluyveri" in t2.label) and ("lactis" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
speciesPair.append([t1.label, t2.label])
speciesDist.append(pdc(t1, t2))
index, value = max(enumerate(speciesDist), key=operator.itemgetter(1))
print("Max. divergence time of second clade (Kluyveromyces) described in Sorrells et al is between: ", speciesPair[index], "\t of time: ", speciesDist[index])
print(max(speciesDist))
##Identifying the evolutionary time horizon for the divergence of all species in the Saccharomycetaceae clade (tested as between S. cerevisiae and K. lactis).
speciesPair = []
speciesDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("bicuspidata" in t1.label) and ("guilliermondii" in t2.label)) or (("bicuspidata" in t2.label) and ("guilliermondii" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
speciesPair.append([t1.label, t2.label])
speciesDist.append(pdc(t1, t2))
index, value = max(enumerate(speciesDist), key=operator.itemgetter(1))
print("Max. divergence time of third clade (Candida) described in Sorrells et al is between: ", speciesPair[index], "\t of time: ", speciesDist[index])
print(max(speciesDist))
##Identifying the evolutionary time horizon for the divergence of all species in the Saccharomycetaceae clade (tested as between S. cerevisiae and K. lactis).
###This doesn't seem like the right pair. I am going to ignore this for now.
speciesPair = []
speciesDist = []
for i, t1 in enumerate(phyloTree.taxon_namespace):
for j, t2 in enumerate(phyloTree.taxon_namespace[:i]):
if ((("polymorpha" in t1.label) and ("tannophilus" in t2.label)) or (("polymorpha" in t2.label) and ("tannophilus" in t1.label))):
print("Distance between '%s' and '%s': %s" % (t1.label, t2.label, pdc(t1, t2)))
speciesPair.append([t1.label, t2.label])
speciesDist.append(pdc(t1, t2))
index, value = max(enumerate(speciesDist), key=operator.itemgetter(1))
print("Max. divergence time of fourth clade (Pichia) described in Sorrells et al is between: ", speciesPair[index], "\t of time: ", speciesDist[index])
print(max(speciesDist))
# ----
# Inferences from the runs above:
# 1. From the 4 clades, it seems like a good time horizon of conservation of mating gene motifs are 180, since I am hoping for a conservative estimate, i.e. the horizon that gives me maximum confidence of shared motifs.
# 2. This is also consistent with the fact that the only time-horizon of STE12 motif conserved in a-specific gene promoters is for the clade of Saccharomyces.
# 3. Another possible thought is to restrict this search within the major clades that I have. But would this work?
# ----
# ----
# ### First-pass taxonomic grouping based on the divergence times
# Let us assume that the list of taxons are ordered such that neighbors in the list are the closest and we can therefore limit are check of relatedness to the immediate neighbor and only move on once we classify as within the group or out of the group.
# > I'm not sure if this is true for an arbitrary tree. I am going to think about how to generalize based on just slicing lists based on the distance measures. I can move down a list of taxon_names and then remove members that are within a particular divergence time.
# >
# > {191026/1628hrs} I've managed to code a more generalized classifier 3 cells down.
#
# Also it is worth considering that the divergenceLimit that is appropriate is that which separates the Saccharomyces cerevisiae and Kluyveromyces lactis species on the tree since the pheromones can be found by sequence homology. Need to confirm this by tBLASTN.
# +
##Trying to build groupings of strains that are within a known value of divergence time.
##I am going to round up from the value estimated in the yarrowia group.
divergenceLimit = 50.0;
taxoGroups = [[phyloTree.taxon_namespace[0].label]]
for i, t1 in enumerate(phyloTree.taxon_namespace[:-2]):
#print(i)
if pdc(t1, phyloTree.taxon_namespace[i+1]) <= divergenceLimit:
taxoGroups[-1].append(phyloTree.taxon_namespace[i+1].label)
else:
taxoGroups.append([phyloTree.taxon_namespace[i+1].label])
print(len(taxoGroups))
##Hist of group sizes
groupSize = []
for taxoGroup in taxoGroups:
groupSize.append(len(taxoGroup))
binList = list(range(max(groupSize)))
print(max(groupSize))
ax1 = sns.histplot(groupSize, kde=False, bins=binList, stat='count', element='step', fill=False);
##ax1 = sns.distplot(groupSize, kde=False, bins=binList, hist_kws={"histtype": "step", "linewidth": 3, "alpha": 1});
#plt.ylim(0, 50)
#ax2 = sns.swarmplot(groupSize);
# with open(os.path.join(tree_dir, "taxonGroups.csv"), 'w') as writeFile:
# writer = csv.writer(writeFile)
# writer.writerows(taxoGroups)
# +
##Trying to build groupings of strains that are within a known value of divergence time.
##I am going to be as loose about including more species by doubling the divergence time from the yarrowia group (above).
divergenceLimit = 100.0;
taxoGroups = [[phyloTree.taxon_namespace[0].label]]
for i, t1 in enumerate(phyloTree.taxon_namespace[:-2]):
#print(i)
if pdc(t1, phyloTree.taxon_namespace[i+1]) <= divergenceLimit:
taxoGroups[-1].append(phyloTree.taxon_namespace[i+1].label)
else:
taxoGroups.append([phyloTree.taxon_namespace[i+1].label])
print(len(taxoGroups))
##Hist of group sizes
groupSize = []
for taxoGroup in taxoGroups:
groupSize.append(len(taxoGroup))
binList = list(range(max(groupSize)))
#print(binList)
ax1 = sns.histplot(groupSize, kde=False, bins=binList, stat='count', element='step', fill=False);
# +
## Building a loop through leaf names that can sort taxonomic groups. This is a generalized search that should identify species groupings for
divergenceLimit = 240.0;
taxoGroups = []
taxonList = []
for taxon in phyloTree.taxon_namespace:
taxonList.append(taxon.label)
#treeClone = phyloTree.clone(depth=1)
for taxon in phyloTree.taxon_namespace:
while taxon.label in taxonList:
taxoGroups.append([taxon.label])
for taxonCompare in phyloTree.taxon_namespace:
if (pdc(taxon, taxonCompare) <= divergenceLimit) and (taxonCompare != taxon):
taxoGroups[-1].append(taxonCompare.label)
taxonList.remove(taxonCompare.label)
taxonList.remove(taxon.label)
print("Total number of monophyletic groups:\t", len(taxoGroups))
##Hist of group sizes
groupSize = []
for taxoGroup in taxoGroups:
groupSize.append(len(taxoGroup))
binList = list(range(max(groupSize)))
print("Number of species in largest monophyletic group:\t", max(groupSize))
ax1 = sns.histplot(groupSize, kde=False, bins=binList, stat='count', element='step', fill=False);
#ax1 = sns.distplot(groupSize, kde=False, bins=binList, hist_kws={"histtype": "step", "linewidth": 3, "alpha": 1});
#plt.ylim(0, 50)
#ax2 = sns.swarmplot(groupSize);
with open(os.path.join(tree_dir, "taxonGroups_240max-dist.csv"), 'w') as writeFile:
writer = csv.writer(writeFile)
writer.writerows(taxoGroups)
# -
###Phylogenetic groups that correspond to infered time-horizon of lipidated pheromone conservation.
print(*[group for group in taxoGroups if len(group)>=1], sep = "\n")
#print(phyloTree.taxon_namespace)
test = [];
taxonList = phyloTree.taxon_namespace;
for taxon in phyloTree.taxon_namespace:
if taxon in taxonList:
#print(taxon.label)
for taxonCompare in phyloTree.taxon_namespace:
test.append(pdc(taxon, taxonCompare))
print(len(test))
# #### Code to print out Phylo-groups within a certain evolutionary distance threshold where the pheromones are expected to be conserved
# Work started on 20200204 at 1545hrs. I am just going to publish groups that have 2 or more species within the group.
##Making sure the species names have underscores instead of white space analogous to what is listed in the dictionary "332taxa_index.txt" (and "332genomes_Shen2017.csv").
iterCount = 0;
for iterCount in range(len(taxoGroups)):
#print(len(taxoGroups[iterCount]))
iterCount1 = 0;
for iterCount1 in range(len(taxoGroups[iterCount])):
taxoGroups[iterCount][iterCount1] = taxoGroups[iterCount][iterCount1].replace(" ", "_")
#print(taxoGroups[iterCount])
#print("Done modifying species names in ", len(taxoGroups), "taxon groups. Good Luck!")
# +
##I need to import the csv from 332taxa_index into a dictionary with the columns {"old_speceis_names" : "original_genome_id"}. I think there is a direct way to do this.
with open(os.path.join(tree_dir, "332genomes_Shen2017.csv"), mode='r') as infile:
reader = csv.reader(infile)
fileDict = {rows[1]:rows[0] for rows in reader}
#print(len(fileDict))
#print(taxoGroups[0][0], fileDict[taxoGroups[0][0]])
##I am going to print out the species lits in the Phylo groups of Fungi and I hope to pool these species together to judge the copy number in these groups.
if not(os.path.isdir(os.path.join(tree_dir, "PhyloGroups_pheromones"))):
os.mkdir(os.path.join(tree_dir, "PhyloGroups_pheromones"))
groupCount = 0
for groupCount in range(len(taxoGroups)):
if len(taxoGroups[groupCount]) >= 2:
outputHandle = open(os.path.join(tree_dir, "PhyloGroups_pheromones", "phyloGroup_"+str(groupCount+1)+".txt"), "w")
for species in taxoGroups[groupCount]:
#print(fileDict[species])
outputHandle.write(fileDict[species]+".fas.caax.asn"+"\n")
outputHandle.close()
# -
# #### Code-chunk to identify time-horizon of mating regulatory motifs in the yeast lineage
# +
## Building a loop through leaf names that can sort taxonomic groups. This is a generalized search that should identify species groupings for
divergenceLimit = 180.0;
taxonGroups_regulation = []
taxonList_regulation = []
for taxon in phyloTree.taxon_namespace:
taxonList_regulation.append(taxon.label)
#treeClone = phyloTree.clone(depth=1)
for taxon in phyloTree.taxon_namespace:
while taxon.label in taxonList_regulation:
taxonGroups_regulation.append([taxon.label])
for taxonCompare in phyloTree.taxon_namespace:
if (pdc(taxon, taxonCompare) <= divergenceLimit) and (taxonCompare != taxon):
taxonGroups_regulation[-1].append(taxonCompare.label)
taxonList_regulation.remove(taxonCompare.label)
taxonList_regulation.remove(taxon.label)
print("Total number of monophyletic groups:\t", len(taxonGroups_regulation))
##Hist of group sizes
groupSize = []
for taxonGroup_regulation in taxonGroups_regulation:
groupSize.append(len(taxonGroup_regulation))
binList = list(range(max(groupSize)))
print("Number of species in largest monophyletic group:\t", max(groupSize))
ax1 = sns.histplot(groupSize, kde=False, bins=binList, stat='count', element='step', fill=False);
#plt.ylim(0, 50)
#ax2 = sns.swarmplot(groupSize);
with open(os.path.join(tree_dir, "taxonGroups_180max-dist_mating-regulation.csv"), 'w') as writeFile:
writer = csv.writer(writeFile)
writer.writerows(taxoGroups)
# -
#print(phyloTree.taxon_namespace)
test = [];
taxonList_regulation = phyloTree.taxon_namespace;
for taxon in phyloTree.taxon_namespace:
if taxon in taxonList_regulation:
#print(taxon.label)
for taxonCompare in phyloTree.taxon_namespace:
test.append(pdc(taxon, taxonCompare))
print(len(test))
print(*[group for group in taxonGroups_regulation if len(group)>=1], sep = "\n")
##Making sure the species names have underscores instead of white space analogous to what is listed in the dictionary "332taxa_index.txt" (and "332genomes_Shen2017.csv").
iterCount = 0;
for iterCount in range(len(taxonGroups_regulation)):
#print(len(taxoGroups_regulation[iterCount]))
iterCount1 = 0;
for iterCount1 in range(len(taxonGroups_regulation[iterCount])):
taxonGroups_regulation[iterCount][iterCount1] = taxonGroups_regulation[iterCount][iterCount1].replace(" ", "_")
#print(taxonGroups_regulation[iterCount])
print("Done modifying species names in ", len(taxonGroups_regulation), "taxon groups. Good Luck!")
# +
##I need to import the csv from 332taxa_index into a dictionary with the columns {"old_speceis_names" : "original_genome_id"}. I think there is a direct way to do this.
with open(os.path.join(tree_dir, "332genomes_Shen2017.csv"), mode='r') as infile:
reader = csv.reader(infile)
fileDict = {rows[1]:rows[0] for rows in reader}
#print(len(fileDict))
#print(taxonGroups_regulation[0][0])
#print(taxonGroups_regulation[0][0], fileDict[taxonGroups_regulation[0][0]])
##I am going to print out the species lits in the Phylo groups of Fungi and I hope to pool these species together to judge the copy number in these groups.
if not(os.path.isdir(os.path.join(tree_dir, "PhyloGroups_regulation"))):
os.mkdir(os.path.join(tree_dir, "PhyloGroups_regulation"))
groupCount = 0
for groupCount in range(len(taxonGroups_regulation)):
if len(taxonGroups_regulation[groupCount]) >= 2:
outputHandle = open(os.path.join(tree_dir, "PhyloGroups_regulation", "phyloGroup_"+str(groupCount+1)+".txt"), "w")
for species in taxonGroups_regulation[groupCount]:
#print(fileDict[species])
outputHandle.write("promoters_0_"+fileDict[species]+".fasta"+"\n")
outputHandle.close()
# +
##I need to import the csv from 332taxa_index into a dictionary with the columns {"old_speceis_names" : "original_genome_id"}. I think there is a direct way to do this.
with open(os.path.join(tree_dir, "332genomes_Shen2017.csv"), mode='r') as infile:
reader = csv.reader(infile)
fileDict = {rows[1]:rows[0] for rows in reader}
#print(len(fileDict))
#print(taxonGroups_regulation[0][0])
#print(taxonGroups_regulation[0][0], fileDict[taxonGroups_regulation[0][0]])
##I am going to print out the species lits in the Phylo groups of Fungi and I hope to pool these species together to judge the copy number in these groups.
if not(os.path.isdir(os.path.join(tree_dir, "PhyloGroups_candidate_regulation"))):
os.mkdir(os.path.join(tree_dir, "PhyloGroups_candidate_regulation"))
groupCount = 0
for groupCount in range(len(taxonGroups_regulation)):
if len(taxonGroups_regulation[groupCount]) >= 2:
outputHandle = open(os.path.join(tree_dir, "PhyloGroups_candidate_regulation", "phyloGroup_"+str(groupCount+1)+".txt"), "w")
for species in taxonGroups_regulation[groupCount]:
#print(fileDict[species])
outputHandle.write(fileDict[species]+".fas.caax.asn.promoters"+"\n")
outputHandle.close()
# -
# ## Code to sort all CAAX-Asn candidates for unique loci
# >Date started 191211, 1500 IST
#
# This is a small code block to accept a fasta list of all the candidates I identify from a genome and then try to get a unique list for finding multiple copies of a candidate at different genome loci (obviously).
# +
##Importing the FASTA file, this won't be used when I add this to the python codebase for functionality. Remember to run the first two blocks for all library imports.
inputFile = "Yarli1_AssemblyScaffolds-put-v2p3_180508.fasta"
fileHandle = open(data_dir+inputFile, "r")
candidates = list(SeqIO.parse(fileHandle, "fasta"))
fileHandle.close()
print(candidates[0].id)
#print(candidates[0].id.split("||")[1].split("[")[3].split("-")[:])
print("There are %i candidates" % len(candidates))
# -
# #### Algorthm for stack search
# - The simplest approach is to brute-force the search and basically compare the current candidate to with all others to see if there is something that shares the chromID, direction and CAAX-STOP positions while having a higher candidate number.
# - Another option is to just invert the stack and pull candidates one after another and since the first candidate from the back would be the largest ORF at any given CAAX-STOP, then all I would have to do is compare the current ID to the last in the new list I am making, and if it is the same keep moving, else append to the list I am making.
# +
##Function to parse candidates and eliminate those that share a CAAX tag (retain the largest ORF among those that share CAAX-STOP).
##Trying the cheat version with a reversed iterator.
uniqueCandidates = []
skipCount = 0
for candidate in reversed(candidates):
#print(candidate.id)
if (uniqueCandidates != []):
if (candidate.id.split("||")[0] == uniqueCandidates[-1].id.split("||")[0]) and (candidate.id.split("||")[1].split("[")[1] == uniqueCandidates[-1].id.split("||")[1].split("[")[1]):
if candidate.id.split("||")[1].split("[")[3].split("-")[2] == uniqueCandidates[-1].id.split("||")[1].split("[")[3].split("-")[2]:
skipCount += 1
else:
uniqueCandidates.append(candidate)
else:
uniqueCandidates.append(candidate)
else:
uniqueCandidates.append(candidate)
print("Skipped %i candidates" % skipCount)
uniqueCandidates.reverse()
print(len(uniqueCandidates))
##printing output.
outputHandle = open(data_dir+inputFile.split(".f")[0]+"_unique.fasta", "w")
for candidate in uniqueCandidates:
outputHandle.write(">"+str(candidate.id)+"\n"+str(candidate.seq)+"\n")
outputHandle.close()
# -
for candidate in uniqueCandidates:
print(candidate.id)
|
notebooks/pheromoneFinder_py3-testbed_210102.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import json
import sys
sys.path.append('../.venv/lib/python3.7/site-packages')
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# +
metricProps = {
'messageID': '',
'originatorHostID': '',
'totalNanoTime': '',
'lastDeliveryHop': '',
'relativeMessageRedundancy': ''
}
class Metric(object):
def __init__(self, props = metricProps):
self.messageID = props['messageID']
self.originatorHostID = props['originatorHostID']
self.totalNanoTime = int(props['totalNanoTime'])
self.lastDeliveryHop = float(props['lastDeliveryHop'])
self.relativeMessageRedundancy = float(props['relativeMessageRedundancy'])
# -
def computeMetrics(algorithm = 'discovery'):
metrics = []
nanoTimes = []
ldh = []
rmr = []
pubsubAnalysisFiles = glob.glob('../data/gossip/{}/analyses/*.json'.format(algorithm))
for pubsubAnalysisFile in pubsubAnalysisFiles:
tmpMetrics = []
with open(pubsubAnalysisFile) as json_file:
data = json.load(json_file)
for metric in data:
m = Metric(metric)
tmpMetrics.append(m)
nanoTimes.append(m.totalNanoTime)
ldh.append(m.lastDeliveryHop)
rmr.append(m.relativeMessageRedundancy)
metrics.append(tmpMetrics)
_ = plt.hist(nanoTimes, bins='auto', histtype='step')
plt.title("{} - Total Nano Time Histogram".format(algorithm))
plt.xlabel('Total Nano Time')
plt.ylabel('Occurrence')
plt.show()
_ = plt.hist(ldh, bins='auto', histtype='step')
plt.title("{} - Last Delivery Hop Histogram".format(algorithm))
plt.xlabel('Last Delivery Hop')
plt.ylabel('Occurrence')
plt.show()
_ = plt.hist(rmr, bins='auto', histtype='step')
plt.title("{} - Relative Message Redundancy Histogram".format(algorithm))
plt.xlabel('Relative Message Redundancy')
plt.ylabel('Occurrence')
plt.show()
nanoMean = np.mean(nanoTimes)
nanoMedian = np.median(nanoTimes)
nanoStd = np.std(nanoTimes)
ldhMean = np.mean(ldh)
ldhMedian = np.median(ldh)
ldhStd = np.std(ldh)
rmrMean = np.mean(rmr)
rmrMedian = np.median(rmr)
rmrStd = np.std(rmr)
print('Messages published: {}'.format(len(nanoTimes)))
print('Total Nano Times - mean: {}, median: {}, std: {}'.format(nanoMean, nanoMedian, nanoStd))
print('Last Delivery Hop - mean: {}, median: {}, std: {}'.format(ldhMean, ldhMedian, ldhStd))
print('Relative Message Redundancy - mean: {}, median: {}, std: {}'.format(rmrMean, rmrMedian, rmrStd))
computeMetrics('discovery')
computeMetrics('no-discovery')
|
examples/benchmark-tests-3/jupyter/Analyses.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Load the Dataset
# +
import pandas as pd
import warnings
warnings.filterwarnings("ignore")
# -
df = pd.read_csv('No2 dataset.csv')
df.head()
df['Date_Time'].max(), df['Date_Time'].min()
df.dtypes
# ### Convert to datetime
df['Date_Time'] = pd.to_datetime(df['Date_Time'], format= '%d/%m/%Y %H.%M.%S')
df.dtypes
# The format depends on your data. Here are a few examples
#
# 1. **1/12/2001**: %d/%m/%Y
# 2. **2001/12/1**: %Y/%m/%d
# 3. **2001-12-01**: %Y-%m-%d
# 4. **2001-12-01 09:00**: %Y-%m-%d %H:%M
df['Date_Time'].max(), df['Date_Time'].min()
# ### Extract hour and minute
df['Date_Time'].dt.hour.head()
df['Date_Time'].dt.minute.head()
# ### The day of week - Monday (0) to Sunday (6)
df['Date_Time'].dt.dayofweek.head(7)
df['Date_Time'].dt.weekday_name.head(7)
# ### Extracting the month from the date
df['Date_Time'].dt.month.head(7)
df['Date_Time'].dt.is_month_end.head(7)
# ### Put it all together
new_df = pd.DataFrame({"year": df['Date_Time'].dt.year,
"month": df['Date_Time'].dt.month,
"day": df['Date_Time'].dt.day,
"hour": df['Date_Time'].dt.hour,
"dayofyear": df['Date_Time'].dt.dayofyear,
"week": df['Date_Time'].dt.week,
"dayofweek": df['Date_Time'].dt.dayofweek,
"dayofweekname": df['Date_Time'].dt.weekday_name,
"quarter": df['Date_Time'].dt.quarter,
})
new_df.head()
complete_data = pd.concat([df, new_df], axis=1)
complete_data.head()
# ### is_weekday? (Create using the dayofweek numbers)
# +
complete_data['is_weekday']=0
for i in range(0, len(complete_data)):
if ((complete_data['dayofweek'][i] == 5) | (complete_data['dayofweek'][i] == 6)):
complete_data['is_weekday'][i] = 0
else:
complete_data['is_weekday'][i] = 1
# -
complete_data[['dayofweek', 'is_weekday']].head()
# ## Difference Between Dates
agent = pd.read_csv('agent_classification.csv')
agent.head()
agent[['Application_Receipt_Date','Applicant_BirthDate']].head()
agent['Application_Receipt_Date'] = pd.to_datetime(agent['Application_Receipt_Date'], format='%m/%d/%Y')
agent['Applicant_BirthDate'] = pd.to_datetime(agent['Applicant_BirthDate'], format='%m/%d/%Y')
(agent['Application_Receipt_Date'][0] - agent['Applicant_BirthDate'][0]).days
agent['Applicant_age'] = agent.apply(lambda x: (x['Application_Receipt_Date'] - x['Applicant_BirthDate']).days, axis=1)
|
Feature Engineering/Implementing DateTime Features/Datetime Features.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="_T81rYsZbh4x" colab_type="text"
# # ASSIGNMENT 1
# Using Keras to build a CNN network for CIFAR-10 dataset. Each record is of size
# 1*3072. Building a CNN network to classify the data into the 10
# classes.
# + [markdown] id="E0sxAlBqyU7g" colab_type="text"
# # Dataset
# CIFAR-10 dataset The CIFAR-10 dataset consists of 60000 32x32 colour images in 10 classes, with 6000 images per class. There are 50000 training images and 10000 test images.
#
# The dataset is divided into five training batches and one test batch, each with 10000 images. The test batch contains exactly 1000 randomly-selected images from each class. The training batches contain the remaining images in random order, but some training batches may contain more images from one class than another. Between them, the training batches contain exactly 5000 images from each class.
#
# http://www.cs.utoronto.ca/~kriz/cifar.html
# + [markdown] id="U_1YfYEjg0yk" colab_type="text"
# # Importing libraries
# + id="Fjtih1xkgxRu" colab_type="code" colab={}
from __future__ import print_function
import keras
from keras.datasets import cifar10
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Conv2D, MaxPooling2D
import os
# + [markdown] id="uEFkUN-XjvmV" colab_type="text"
# # Loading the CIFAR 10 dataset
# + id="GQFbwb4mgx8Z" colab_type="code" colab={}
batch_size = 32
num_classes = 10
epochs = 50
data_augmentation = True
num_predictions = 20
save_dir = os.path.join(os.getcwd(), 'saved_models')
model_name = 'keras_cifar10_trained_model.h5'
# + [markdown] id="MRPVjoxl2NOq" colab_type="text"
# # The data, split between train and test sets:
# + id="7QfiFe7gl79d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="53dacdc2-6263-4fb9-8ffb-55d749a66a9e"
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# + [markdown] id="KrLhw1HT2jIG" colab_type="text"
# # Convert class vectors to binary class matrices.
# + id="pIG9OWT2mDkY" colab_type="code" colab={}
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# + [markdown] id="IiwfGA8JSHLX" colab_type="text"
# # Defining the model
# + id="amUoQATgmKgu" colab_type="code" colab={}
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same',
input_shape=x_train.shape[1:]))
model.add(Activation('relu'))
model.add(Conv2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
# + id="bUJxHDcVgyCA" colab_type="code" colab={}
model.add(Conv2D(64, (3, 3), padding='same'))
model.add(Activation('relu'))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
# + id="QfdVdkkAgyEp" colab_type="code" colab={}
model.add(Flatten())
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
# + [markdown] id="eGUcCDc-STru" colab_type="text"
# # Initiating RMSprop optimizer
# + id="P2CPDxBXgyIx" colab_type="code" colab={}
opt = keras.optimizers.rmsprop(lr=0.0001, decay=1e-6)
# + [markdown] id="Oi0Yi_P4SX74" colab_type="text"
# # Training the model using RMSprop
# + id="GsdpV_KHgyLd" colab_type="code" colab={}
model.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
# + id="5oBMxyPEgyN9" colab_type="code" colab={}
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
# + [markdown] id="sCmYW7-MSjSx" colab_type="text"
# # Fitting the model
# + id="55OSWYPkgySk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1734} outputId="98e0c476-23c3-491e-e48f-04298cf5046f"
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# + [markdown] id="9nT8A_eySpdJ" colab_type="text"
# # Scoring and Evaluating trained model.
# + id="RH-6DMLzgyHM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="771704b0-8b79-48ef-90af-06b007eb8eec"
scores = model.evaluate(x_test, y_test, verbose=1)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# + [markdown] id="jAKe7aewS2kS" colab_type="text"
# # Printing the accuracy
# + id="q6Cf7UyE4rdb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c0563ae0-961f-4803-9d34-f6823c651bf2"
scores = model.evaluate(x_test, y_test, verbose=0)
print("Accuracy: %.2f%%" % (scores[1]*100))
# + [markdown] id="NRv9W64QSxy5" colab_type="text"
# # Plot training & validation accuracy values
# + id="rLXTbGJwCerc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 735} outputId="040199bc-7345-4def-cb79-2f762d771635"
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# + id="KD2AXXHhjbho" colab_type="code" colab={}
# plot graph
plot_model(model, to_file='convolutional_neural_network.png')
# + [markdown] id="6P5qLfk03M7s" colab_type="text"
# # Results
# 1) Provide a recommendation for the best model you would
# recommend for classification. Which model (with parameter values) would you choose
# and why?
# According to me, the best model for classification is cnn model 1. The best parameters that helped improve the model are: epochs = 50
# batch_size = 32
# keep_probability = 0.7
# learning_rate = 0.001
# Optimizer
#
# • Comment on how good your model is ? Does it overfit/underfit data ? What could you
# do to improve the model
# I ran it for 50 epochs and got almost 78% accuracy. It can surely go much further since it was still undertrained! To improve the model the learning rate should be improved. Also, number of neurons can be more complicated for a better fit.
# + [markdown] id="iVA_z0aSIZdi" colab_type="text"
# # References
# [1] https://github.com/fchollet/keras/blob/master/examples/cifar10_cnn.py
#
# [2] http://www.cs.utoronto.ca/~kriz/cifar.html
#
# [3] https://towardsdatascience.com/cifar-10-image-classification-in-tensorflow-5b501f7dc77c
#
|
Assignment 1 Keras/CIFAR10_using_cnn_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py352]
# language: python
# name: conda-env-py352-py
# ---
# # 3x Options for Geo-Coding using API Calls
import requests
import ujson
# # TAMU (Texas A&M... WHOOP!) GeoSciences API
# +
tamu_creds = ujson.load(open("D:\\Python\\tamu_geo_credentials.json"))
api_url = "https://geoservices.tamu.edu/Services/Geocode/WebService/GeocoderWebServiceHttpNonParsed_V04_01.aspx?"
api_data = {'apiKey':tamu_creds['api_key'],'version':'4.01','streetAddress':'1325 West Walnut Hill Lane','city':'Irving','state':'TX','zip':'75038'}
r = requests.get(api_url, params = api_data)
print(r.url)
print(r)
print(r.text)
# -
# # Google API
# +
api_data={'address':"1325+West+Walnut+Hill+Lane,+Irving,+TX",'key':ujson.load(open('D:\\Python\\google_maps_api.json'))['api_key']}
url="https://maps.googleapis.com/maps/api/geocode/json"
r = requests.get(url, params = api_data)
print(r.url)
print(r)
print(r.json())
# -
# # GeoPy
# +
from geopy.geocoders import Nominatim
geolocator = Nominatim()
location = geolocator.geocode("1325 West Walnut Hill Lane, Irving TX 75038")
print(location.address)
print(location.latitude,location.longitude)
print(location.raw)
|
Geocoding_Examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def fun(x, y):
print("x=" + str(x) + ", y=" + str(y))
if y == 0:
return 0
return x+fun(x,y-1)
def fun2(a, b):
print("a=" + str(a) + ", b=" + str(b))
if b == 0:
return 1
return fun(a, fun2(a, b-1))
fun2(3, 3)
def recMethod(x):
if x == 40:
return x
else:
return x + recMethod(x*2)
recMethod(5)
def mystery(z, n):
if n == z:
return z
elif n > z:
return mystery(z, n-z)
else:
return mystery(z-n, n)
mystery(6,4)
def digits(n):
if n < 0:
n = -n
if n < 10 and n >= 0:
return n
return n%10 + digits(n//10)
digits(-24684)
def Side(x):
if x > 5:
print(x)
else:
print(x)
Side(x - 1)
Side(10)
def search2(seq, v, low , high):
if low > high:
return None
mid = (low + high) // 2
if v == seq[mid]:
return mid
print(seq, v)
elif v > seq[mid]:
return search2(seq, v, mid + 1, high)
else:
return search2(seq, v, low, mid - 1)
seq = [4,6,9,2,8,3,7]
search2(seq, 3, 0, len(seq))
|
A2ComputerScience/Recursion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Neural network hybrid recommendation system on Google Analytics data preprocessing
#
# This notebook demonstrates how to implement a hybrid recommendation system using a neural network to combine content-based and collaborative filtering recommendation models using Google Analytics data. We are going to use the learned user embeddings from [wals.ipynb](wals.ipynb) and combine that with our previous content-based features from [content.ipynb](content.ipynb)
#
# First we are going to preprocess our data using BigQuery and Cloud Dataflow to be used in our later neural network hybrid recommendation model.
# + [markdown] deletable=true editable=true
# Apache Beam only works in Python 2 at the moment, so we're going to switch to the Python 2 kernel. In the above menu, click the dropdown arrow and select `python2`.
# + deletable=true editable=true language="bash"
# source activate py2env
# pip uninstall -y google-cloud-dataflow
# conda install -y pytz==2018.4
# pip install apache-beam[gcp]
# + [markdown] deletable=true editable=true
# Now restart notebook's session kernel!
# + deletable=true editable=true
# Import helpful libraries and setup our project, bucket, and region
import os
PROJECT = 'cloud-training-demos' # REPLACE WITH YOUR PROJECT ID
BUCKET = 'cloud-training-demos-ml' # REPLACE WITH YOUR BUCKET NAME
REGION = 'us-central1' # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
# do not change these
os.environ['PROJECT'] = PROJECT
os.environ['BUCKET'] = BUCKET
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = '1.8'
# + deletable=true editable=true
# %bash
gcloud config set project $PROJECT
gcloud config set compute/region $REGION
# + [markdown] deletable=true editable=true
# <h2> Create ML dataset using Dataflow </h2>
# Let's use Cloud Dataflow to read in the BigQuery data, do some preprocessing, and write it out as CSV files.
#
# First, let's create our hybrid dataset query that we will use in our Cloud Dataflow pipeline. This will combine some content-based features and the user and item embeddings learned from our WALS Matrix Factorization Collaborative filtering lab that we extracted from our trained WALSMatrixFactorization Estimator and uploaded to BigQuery.
# + deletable=true editable=true
query_hybrid_dataset = """
WITH CTE_site_history AS (
SELECT
fullVisitorId as visitor_id,
(SELECT MAX(IF(index = 10, value, NULL)) FROM UNNEST(hits.customDimensions)) AS content_id,
(SELECT MAX(IF(index = 7, value, NULL)) FROM UNNEST(hits.customDimensions)) AS category,
(SELECT MAX(IF(index = 6, value, NULL)) FROM UNNEST(hits.customDimensions)) AS title,
(SELECT MAX(IF(index = 2, value, NULL)) FROM UNNEST(hits.customDimensions)) AS author_list,
SPLIT(RPAD((SELECT MAX(IF(index = 4, value, NULL)) FROM UNNEST(hits.customDimensions)), 7), '.') AS year_month_array,
LEAD(hits.customDimensions, 1) OVER (PARTITION BY fullVisitorId ORDER BY hits.time ASC) AS nextCustomDimensions
FROM
`cloud-training-demos.GA360_test.ga_sessions_sample`,
UNNEST(hits) AS hits
WHERE
# only include hits on pages
hits.type = "PAGE"
AND
fullVisitorId IS NOT NULL
AND
hits.time != 0
AND
hits.time IS NOT NULL
AND
(SELECT MAX(IF(index = 10, value, NULL)) FROM UNNEST(hits.customDimensions)) IS NOT NULL
),
CTE_training_dataset AS (
SELECT
(SELECT MAX(IF(index=10, value, NULL)) FROM UNNEST(nextCustomDimensions)) AS next_content_id,
visitor_id,
content_id,
category,
REGEXP_REPLACE(title, r",", "") AS title,
REGEXP_EXTRACT(author_list, r"^[^,]+") AS author,
DATE_DIFF(DATE(CAST(year_month_array[OFFSET(0)] AS INT64), CAST(year_month_array[OFFSET(1)] AS INT64), 1), DATE(1970, 1, 1), MONTH) AS months_since_epoch
FROM
CTE_site_history
WHERE (SELECT MAX(IF(index=10, value, NULL)) FROM UNNEST(nextCustomDimensions)) IS NOT NULL)
SELECT
CAST(next_content_id AS STRING) AS next_content_id,
CAST(training_dataset.visitor_id AS STRING) AS visitor_id,
CAST(training_dataset.content_id AS STRING) AS content_id,
CAST(IFNULL(category, 'None') AS STRING) AS category,
CONCAT("\\"", REPLACE(TRIM(CAST(IFNULL(title, 'None') AS STRING)), "\\"",""), "\\"") AS title,
CAST(IFNULL(author, 'None') AS STRING) AS author,
CAST(months_since_epoch AS STRING) AS months_since_epoch,
IFNULL(user_factors._0, 0.0) AS user_factor_0,
IFNULL(user_factors._1, 0.0) AS user_factor_1,
IFNULL(user_factors._2, 0.0) AS user_factor_2,
IFNULL(user_factors._3, 0.0) AS user_factor_3,
IFNULL(user_factors._4, 0.0) AS user_factor_4,
IFNULL(user_factors._5, 0.0) AS user_factor_5,
IFNULL(user_factors._6, 0.0) AS user_factor_6,
IFNULL(user_factors._7, 0.0) AS user_factor_7,
IFNULL(user_factors._8, 0.0) AS user_factor_8,
IFNULL(user_factors._9, 0.0) AS user_factor_9,
IFNULL(item_factors._0, 0.0) AS item_factor_0,
IFNULL(item_factors._1, 0.0) AS item_factor_1,
IFNULL(item_factors._2, 0.0) AS item_factor_2,
IFNULL(item_factors._3, 0.0) AS item_factor_3,
IFNULL(item_factors._4, 0.0) AS item_factor_4,
IFNULL(item_factors._5, 0.0) AS item_factor_5,
IFNULL(item_factors._6, 0.0) AS item_factor_6,
IFNULL(item_factors._7, 0.0) AS item_factor_7,
IFNULL(item_factors._8, 0.0) AS item_factor_8,
IFNULL(item_factors._9, 0.0) AS item_factor_9,
FARM_FINGERPRINT(CONCAT(CAST(visitor_id AS STRING), CAST(content_id AS STRING))) AS hash_id
FROM CTE_training_dataset AS training_dataset
LEFT JOIN `cloud-training-demos.GA360_test.user_factors` AS user_factors
ON CAST(training_dataset.visitor_id AS FLOAT64) = CAST(user_factors.user_id AS FLOAT64)
LEFT JOIN `cloud-training-demos.GA360_test.item_factors` AS item_factors
ON CAST(training_dataset.content_id AS STRING) = CAST(item_factors.item_id AS STRING)
"""
# + [markdown] deletable=true editable=true
# Let's pull a sample of our data into a dataframe to see what it looks like.
# + deletable=true editable=true
import google.datalab.bigquery as bq
df_hybrid_dataset = bq.Query(query_hybrid_dataset + "LIMIT 100").execute().result().to_dataframe()
df_hybrid_dataset.head()
# + deletable=true editable=true
df_hybrid_dataset.describe()
# + deletable=true editable=true
import apache_beam as beam
import datetime, os
def to_csv(rowdict):
# Pull columns from BQ and create a line
import hashlib
import copy
CSV_COLUMNS = 'next_content_id,visitor_id,content_id,category,title,author,months_since_epoch'.split(',')
FACTOR_COLUMNS = ["user_factor_{}".format(i) for i in range(10)] + ["item_factor_{}".format(i) for i in range(10)]
# Write out rows for each input row for each column in rowdict
data = ','.join(['None' if k not in rowdict else (rowdict[k].encode('utf-8') if rowdict[k] is not None else 'None') for k in CSV_COLUMNS])
data += ','
data += ','.join([str(rowdict[k]) if k in rowdict else 'None' for k in FACTOR_COLUMNS])
yield ('{}'.format(data))
def preprocess(in_test_mode):
import shutil, os, subprocess
job_name = 'preprocess-hybrid-recommendation-features' + '-' + datetime.datetime.now().strftime('%y%m%d-%H%M%S')
if in_test_mode:
print('Launching local job ... hang on')
OUTPUT_DIR = './preproc/features'
shutil.rmtree(OUTPUT_DIR, ignore_errors=True)
os.makedirs(OUTPUT_DIR)
else:
print('Launching Dataflow job {} ... hang on'.format(job_name))
OUTPUT_DIR = 'gs://{0}/hybrid_recommendation/preproc/features/'.format(BUCKET)
try:
subprocess.check_call('gsutil -m rm -r {}'.format(OUTPUT_DIR).split())
except:
pass
options = {
'staging_location': os.path.join(OUTPUT_DIR, 'tmp', 'staging'),
'temp_location': os.path.join(OUTPUT_DIR, 'tmp'),
'job_name': job_name,
'project': PROJECT,
'teardown_policy': 'TEARDOWN_ALWAYS',
'no_save_main_session': True
}
opts = beam.pipeline.PipelineOptions(flags = [], **options)
if in_test_mode:
RUNNER = 'DirectRunner'
else:
RUNNER = 'DataflowRunner'
p = beam.Pipeline(RUNNER, options = opts)
query = query_hybrid_dataset
if in_test_mode:
query = query + ' LIMIT 100'
for step in ['train', 'eval']:
if step == 'train':
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hash_id), 10) < 9'.format(query)
else:
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hash_id), 10) = 9'.format(query)
(p
| '{}_read'.format(step) >> beam.io.Read(beam.io.BigQuerySource(query = selquery, use_standard_sql = True))
| '{}_csv'.format(step) >> beam.FlatMap(to_csv)
| '{}_out'.format(step) >> beam.io.Write(beam.io.WriteToText(os.path.join(OUTPUT_DIR, '{}.csv'.format(step))))
)
job = p.run()
if in_test_mode:
job.wait_until_finish()
print("Done!")
preprocess(in_test_mode = False)
# + [markdown] deletable=true editable=true
# Let's check our files to make sure everything went as expected
# + deletable=true editable=true
# %bash
# rm -rf features
# mkdir features
# + deletable=true editable=true
# !gsutil -m cp -r gs://{BUCKET}/hybrid_recommendation/preproc/features/*.csv* features/
# + deletable=true editable=true
# !head -3 features/*
# + [markdown] deletable=true editable=true
# <h2> Create vocabularies using Dataflow </h2>
#
# Let's use Cloud Dataflow to read in the BigQuery data, do some preprocessing, and write it out as CSV files.
#
# Now we'll create our vocabulary files for our categorical features.
# + deletable=true editable=true
query_vocabularies = """
SELECT
CAST((SELECT MAX(IF(index = index_value, value, NULL)) FROM UNNEST(hits.customDimensions)) AS STRING) AS grouped_by
FROM `cloud-training-demos.GA360_test.ga_sessions_sample`,
UNNEST(hits) AS hits
WHERE
# only include hits on pages
hits.type = "PAGE"
AND (SELECT MAX(IF(index = index_value, value, NULL)) FROM UNNEST(hits.customDimensions)) IS NOT NULL
GROUP BY
grouped_by
"""
# + deletable=true editable=true
import apache_beam as beam
import datetime, os
def to_txt(rowdict):
# Pull columns from BQ and create a line
# Write out rows for each input row for grouped by column in rowdict
return '{}'.format(rowdict['grouped_by'].encode('utf-8'))
def preprocess(in_test_mode):
import shutil, os, subprocess
job_name = 'preprocess-hybrid-recommendation-vocab-lists' + '-' + datetime.datetime.now().strftime('%y%m%d-%H%M%S')
if in_test_mode:
print('Launching local job ... hang on')
OUTPUT_DIR = './preproc/vocabs'
shutil.rmtree(OUTPUT_DIR, ignore_errors=True)
os.makedirs(OUTPUT_DIR)
else:
print('Launching Dataflow job {} ... hang on'.format(job_name))
OUTPUT_DIR = 'gs://{0}/hybrid_recommendation/preproc/vocabs/'.format(BUCKET)
try:
subprocess.check_call('gsutil -m rm -r {}'.format(OUTPUT_DIR).split())
except:
pass
options = {
'staging_location': os.path.join(OUTPUT_DIR, 'tmp', 'staging'),
'temp_location': os.path.join(OUTPUT_DIR, 'tmp'),
'job_name': job_name,
'project': PROJECT,
'teardown_policy': 'TEARDOWN_ALWAYS',
'no_save_main_session': True
}
opts = beam.pipeline.PipelineOptions(flags = [], **options)
if in_test_mode:
RUNNER = 'DirectRunner'
else:
RUNNER = 'DataflowRunner'
p = beam.Pipeline(RUNNER, options = opts)
def vocab_list(index, name):
query = query_vocabularies.replace("index_value", "{}".format(index))
(p
| '{}_read'.format(name) >> beam.io.Read(beam.io.BigQuerySource(query = query, use_standard_sql = True))
| '{}_txt'.format(name) >> beam.Map(to_txt)
| '{}_out'.format(name) >> beam.io.Write(beam.io.WriteToText(os.path.join(OUTPUT_DIR, '{0}_vocab.txt'.format(name))))
)
# Call vocab_list function for each
vocab_list(10, 'content_id') # content_id
vocab_list(7, 'category') # category
vocab_list(2, 'author') # author
job = p.run()
if in_test_mode:
job.wait_until_finish()
print("Done!")
preprocess(in_test_mode = False)
# + [markdown] deletable=true editable=true
# Also get vocab counts from the length of the vocabularies
# + deletable=true editable=true
import apache_beam as beam
import datetime, os
def count_to_txt(rowdict):
# Pull columns from BQ and create a line
# Write out count
return '{}'.format(rowdict['count_number'])
def mean_to_txt(rowdict):
# Pull columns from BQ and create a line
# Write out mean
return '{}'.format(rowdict['mean_value'])
def preprocess(in_test_mode):
import shutil, os, subprocess
job_name = 'preprocess-hybrid-recommendation-vocab-counts' + '-' + datetime.datetime.now().strftime('%y%m%d-%H%M%S')
if in_test_mode:
print('Launching local job ... hang on')
OUTPUT_DIR = './preproc/vocab_counts'
shutil.rmtree(OUTPUT_DIR, ignore_errors=True)
os.makedirs(OUTPUT_DIR)
else:
print('Launching Dataflow job {} ... hang on'.format(job_name))
OUTPUT_DIR = 'gs://{0}/hybrid_recommendation/preproc/vocab_counts/'.format(BUCKET)
try:
subprocess.check_call('gsutil -m rm -r {}'.format(OUTPUT_DIR).split())
except:
pass
options = {
'staging_location': os.path.join(OUTPUT_DIR, 'tmp', 'staging'),
'temp_location': os.path.join(OUTPUT_DIR, 'tmp'),
'job_name': job_name,
'project': PROJECT,
'teardown_policy': 'TEARDOWN_ALWAYS',
'no_save_main_session': True
}
opts = beam.pipeline.PipelineOptions(flags = [], **options)
if in_test_mode:
RUNNER = 'DirectRunner'
else:
RUNNER = 'DataflowRunner'
p = beam.Pipeline(RUNNER, options = opts)
def vocab_count(index, column_name):
query = """
SELECT
COUNT(*) AS count_number
FROM ({})
""".format(query_vocabularies.replace("index_value", "{}".format(index)))
(p
| '{}_read'.format(column_name) >> beam.io.Read(beam.io.BigQuerySource(query = query, use_standard_sql = True))
| '{}_txt'.format(column_name) >> beam.Map(count_to_txt)
| '{}_out'.format(column_name) >> beam.io.Write(beam.io.WriteToText(os.path.join(OUTPUT_DIR, '{0}_vocab_count.txt'.format(column_name))))
)
def global_column_mean(column_name):
query = """
SELECT
AVG(CAST({1} AS FLOAT64)) AS mean_value
FROM ({0})
""".format(query_hybrid_dataset, column_name)
(p
| '{}_read'.format(column_name) >> beam.io.Read(beam.io.BigQuerySource(query = query, use_standard_sql = True))
| '{}_txt'.format(column_name) >> beam.Map(mean_to_txt)
| '{}_out'.format(column_name) >> beam.io.Write(beam.io.WriteToText(os.path.join(OUTPUT_DIR, '{0}_mean.txt'.format(column_name))))
)
# Call vocab_count function for each column we want the vocabulary count for
vocab_count(10, 'content_id') # content_id
vocab_count(7, 'category') # category
vocab_count(2, 'author') # author
# Call global_column_mean function for each column we want the mean for
global_column_mean('months_since_epoch') # months_since_epoch
job = p.run()
if in_test_mode:
job.wait_until_finish()
print("Done!")
preprocess(in_test_mode = False)
# + [markdown] deletable=true editable=true
# Let's check our files to make sure everything went as expected
# + deletable=true editable=true
# %bash
# rm -rf vocabs
# mkdir vocabs
# + deletable=true editable=true
# !gsutil -m cp -r gs://{BUCKET}/hybrid_recommendation/preproc/vocabs/*.txt* vocabs/
# + deletable=true editable=true
# !head -3 vocabs/*
# + deletable=true editable=true
# %bash
# rm -rf vocab_counts
# mkdir vocab_counts
# + deletable=true editable=true
# !gsutil -m cp -r gs://{BUCKET}/hybrid_recommendation/preproc/vocab_counts/*.txt* vocab_counts/
# + deletable=true editable=true
# !head -3 vocab_counts/*
|
courses/machine_learning/deepdive/10_recommend/labs/hybrid_recommendations/hybrid_recommendations_preproc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7c263afb-de80-4220-8ed9-c8e87c5af9f1"}
# 
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "328c339e-a337-40e3-a8fb-7377c176690f"}
# # Training and Reusing Clinical Named Entity Recognition Models
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "adba3bd3-4761-44e0-aa9c-4d817e88e9fe"}
# Please make sure that your cluster is setup properly according to https://nlp.johnsnowlabs.com/docs/en/licensed_install#install-spark-nlp-for-healthcare-on-databricks
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "2cabc9d7-b5ae-43a5-992a-eea4b6a40763"}
# ## Blogposts and videos:
#
# https://towardsdatascience.com/named-entity-recognition-ner-with-bert-in-spark-nlp-874df20d1d77
#
# https://www.youtube.com/watch?v=YM-e4eOiQ34
#
# https://medium.com/spark-nlp/named-entity-recognition-for-healthcare-with-sparknlp-nerdl-and-nercrf-a7751b6ad571
#
# https://medium.com/atlas-research/ner-for-clinical-text-7c73caddd180
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "637c75c5-9374-4a6d-8f65-84a71b48586a"}
import os
import json
import string
import numpy as np
import pandas as pd
import sparknlp
import sparknlp_jsl
from sparknlp.base import *
from sparknlp.util import *
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.pretrained import ResourceDownloader
from pyspark.sql import functions as F
from pyspark.ml import Pipeline, PipelineModel
pd.set_option('max_colwidth', 100)
pd.set_option('display.max_columns', 100)
pd.set_option('display.expand_frame_repr', False)
print('sparknlp.version : ',sparknlp.version())
print('sparknlp_jsl.version : ',sparknlp_jsl.version())
spark
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7736d0b6-ebc9-4793-8a97-0ebbeb3b49f3"}
# # Clinical NER Pipeline (with pretrained models)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "33d09296-d6b7-4053-ab34-fb3e9487d51b"}
# Annotator that transforms a text column from dataframe into an Annotation ready for NLP
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
# Sentence Detector annotator, processes various sentences per line
sentenceDetector = SentenceDetectorDLModel.pretrained("sentence_detector_dl_healthcare","en","clinical/models") \
.setInputCols(["document"]) \
.setOutputCol("sentence")
#sentenceDetector = SentenceDetector()\
# .setInputCols(["document"])\
# .setOutputCol("sentence")
# Tokenizer splits words in a relevant format for NLP
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
# Clinical word embeddings trained on PubMED dataset
word_embeddings = WordEmbeddingsModel.pretrained("embeddings_clinical", "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
# NER model trained on i2b2 (sampled from MIMIC) dataset
clinical_ner = MedicalNerModel.pretrained("ner_clinical_large", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")\
.setLabelCasing("upper")
ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "ner"]) \
.setOutputCol("ner_chunk")
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
clinical_ner,
ner_converter])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "408b6428-e3a5-49b7-a92c-32939c3de059"}
#checking the stages in pipeline
model.stages
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "be25425f-c1b2-4efa-9142-3592b8924c92"}
#getting the classes in pretrained model
clinical_ner.getClasses()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "4aa9b7f9-5a7d-44d3-b841-68497d3c6635"}
#extracting the embedded default param values
clinical_ner.extractParamMap()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "b2e5d9e3-a0ae-4bc8-90a6-a282d9487c78"}
#checking the embeddings
clinical_ner.getStorageRef()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "e86347f2-3d0b-4465-8bfc-4c03c989eda2"}
#downloading the sample dataset
# ! wget -q https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/en/pubmed/pubmed_sample_text_small.csv
dbutils.fs.cp("file:/databricks/driver/pubmed_sample_text_small.csv", "dbfs:/")
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "1c6c4b30-8ad2-4684-afc3-a74bcea7b4f4"}
# %fs ls file:/databricks/driver
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "6c8932c9-b85c-4714-8190-24f2f2be27e9"}
import pyspark.sql.functions as F
pubMedDF = spark.read\
.option("header", "true")\
.csv("dbfs:/pubmed_sample_text_small.csv")\
pubMedDF.show(truncate=80)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "20fcb450-e7c5-40df-ac4f-681049346cf6"}
pubMedDF.printSchema()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "49f42e5b-6b93-436f-8ebe-bbf4652adce6"}
result = model.transform(pubMedDF.limit(100))
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "a039d3b6-ea26-40a6-a6d2-1dd7a1fd7c89"}
result.show()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "1e5055e9-b3c6-42ec-a7b1-0008d13a40d3"}
result.select('token.result','ner.result').show(truncate=80)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "48a25bc8-2cf8-4681-92d6-47bc93202a31"}
result_df= result.select(F.explode(F.arrays_zip(result.token.result, result.ner.result)).alias("cols"))\
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ner_label"))
result_df.show(50, truncate=100)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "d62bc5a9-2177-4f32-9fce-493617109e0c"}
result_df.select("token", "ner_label").groupBy('ner_label').count().orderBy('count', ascending=False).show(truncate=False)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "cac8e54f-f77b-43e4-8c7c-ce6e94d0e551"}
result.select('ner_chunk').take(1)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "35f71278-bc4b-4dc5-bcaf-75ab4fd6a4a9"}
result.select(F.explode(F.arrays_zip(result.ner_chunk.result, result.ner_chunk.metadata)).alias("cols"))\
.select(F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']['entity']").alias("ner_label")).show(truncate=False)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "468a9b55-5516-4bd7-bb92-ed5f1ec4fb4b"}
# fullAnnotate in LightPipeline
text = '''
A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and subsequent type two diabetes mellitus ( T2DM ), one prior episode of HTG-induced pancreatitis three years prior to presentation , associated with an acute hepatitis , and obesity with a body mass index ( BMI ) of 33.5 kg/m2 , presented with a one-week history of polyuria , polydipsia , poor appetite , and vomiting . Two weeks prior to presentation , she was treated with a five-day course of amoxicillin for a respiratory tract infection . She was on metformin , glipizide , and dapagliflozin for T2DM and atorvastatin and gemfibrozil for HTG . She had been on dapagliflozin for six months at the time of presentation . Physical examination on presentation was significant for dry oral mucosa ; significantly , her abdominal examination was benign with no tenderness , guarding , or rigidity . Pertinent laboratory findings on admission were : serum glucose 111 mg/dl , bicarbonate 18 mmol/l , anion gap 20 , creatinine 0.4 mg/dL , triglycerides 508 mg/dL , total cholesterol 122 mg/dL , glycated hemoglobin ( HbA1c ) 10% , and venous pH 7.27 . Serum lipase was normal at 43 U/L . Serum acetone levels could not be assessed as blood samples kept hemolyzing due to significant lipemia . The patient was initially admitted for starvation ketosis , as she reported poor oral intake for three days prior to admission . However , serum chemistry obtained six hours after presentation revealed her glucose was 186 mg/dL , the anion gap was still elevated at 21 , serum bicarbonate was 16 mmol/L , triglyceride level peaked at 2050 mg/dL , and lipase was 52 U/L . The β-hydroxybutyrate level was obtained and found to be elevated at 5.29 mmol/L - the original sample was centrifuged and the chylomicron layer removed prior to analysis due to interference from turbidity caused by lipemia again . The patient was treated with an insulin drip for euDKA and HTG with a reduction in the anion gap to 13 and triglycerides to 1400 mg/dL , within 24 hours . Her euDKA was thought to be precipitated by her respiratory tract infection in the setting of SGLT2 inhibitor use . The patient was seen by the endocrinology service and she was discharged on 40 units of insulin glargine at night , 12 units of insulin lispro with meals , and metformin 1000 mg two times a day . It was determined that all SGLT2 inhibitors should be discontinued indefinitely . She had close follow-up with endocrinology post discharge .
'''
print (text)
light_model = LightPipeline(model)
light_result = light_model.fullAnnotate(text)
chunks = []
entities = []
sentence= []
begin = []
end = []
for n in light_result[0]['ner_chunk']:
begin.append(n.begin)
end.append(n.end)
chunks.append(n.result)
entities.append(n.metadata['entity'])
sentence.append(n.metadata['sentence'])
import pandas as pd
df_clinical = pd.DataFrame({'chunks':chunks, 'begin': begin, 'end':end,
'sentence_id':sentence, 'entities':entities})
df_clinical.head(20)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "b9fd11f8-281b-49fe-a1f9-586395440831"}
from sparknlp_display import NerVisualizer
visualiser = NerVisualizer()
ner_vis = visualiser.display(light_result[0], label_col='ner_chunk', document_col='document', return_html=True)
# Change color of an entity label
#visualiser.set_label_colors({'PROBLEM':'#008080', 'TEST':'#800080', 'TREATMENT':'#808080'})
#visualiser.display(light_result[0], label_col='ner_chunk')
# Set label filter
# visualiser.display(light_result, label_col='ner_chunk', document_col='document',
#labels=['PROBLEM','TEST'])
displayHTML(ner_vis)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "89748ed3-3e5d-4098-9597-61ead91cefda"}
# ## NER JSL Model
#
# Let's show an example of `ner_jsl` model that has about 80 labels by changing just only the model name.
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "a2a74fc8-d9d9-4d22-85a5-3e6a71cec724"}
# **Entities**
#
# | | | | | |
# |-|-|-|-|-|
# |Injury_or_Poisoning|Direction|Test|Admission_Discharge|Death_Entity|
# |Relationship_Status|Duration|Respiration|Hyperlipidemia|Birth_Entity|
# |Age|Labour_Delivery|Family_History_Header|BMI|Temperature|
# |Alcohol|Kidney_Disease|Oncological|Medical_History_Header|Cerebrovascular_Disease|
# |Oxygen_Therapy|O2_Saturation|Psychological_Condition|Heart_Disease|Employment|
# |Obesity|Disease_Syndrome_Disorder|Pregnancy|ImagingFindings|Procedure|
# |Medical_Device|Race_Ethnicity|Section_Header|Symptom|Treatment|
# |Substance|Route|Drug_Ingredient|Blood_Pressure|Diet|
# |External_body_part_or_region|LDL|VS_Finding|Allergen|EKG_Findings|
# |Imaging_Technique|Triglycerides|RelativeTime|Gender|Pulse|
# |Social_History_Header|Substance_Quantity|Diabetes|Modifier|Internal_organ_or_component|
# |Clinical_Dept|Form|Drug_BrandName|Strength|Fetus_NewBorn|
# |RelativeDate|Height|Test_Result|Sexually_Active_or_Sexual_Orientation|Frequency|
# |Time|Weight|Vaccine|Vital_Signs_Header|Communicable_Disease|
# |Dosage|Overweight|Hypertension|HDL|Total_Cholesterol|
# |Smoking|Date||||
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "3675ab77-3a9a-4077-a464-bc30ff1c908b"}
# Annotator that transforms a text column from dataframe into an Annotation ready for NLP
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
# Sentence Detector annotator, processes various sentences per line
sentenceDetector = SentenceDetectorDLModel.pretrained("sentence_detector_dl_healthcare","en","clinical/models") \
.setInputCols(["document"]) \
.setOutputCol("sentence")
#sentenceDetector = SentenceDetector()\
# .setInputCols(["document"])\
# .setOutputCol("sentence")
# Tokenizer splits words in a relevant format for NLP
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
# Clinical word embeddings trained on PubMED dataset
word_embeddings = WordEmbeddingsModel.pretrained("embeddings_clinical", "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
# NER model trained on i2b2 (sampled from MIMIC) dataset
jsl_ner = MedicalNerModel.pretrained("ner_jsl", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("jsl_ner")\
.setLabelCasing("upper")
jsl_ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "jsl_ner"]) \
.setOutputCol("jsl_ner_chunk")
jsl_ner_pipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
jsl_ner,
jsl_ner_converter])
empty_data = spark.createDataFrame([[""]]).toDF("text")
jsl_ner_model = jsl_ner_pipeline.fit(empty_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "22bf3135-f94e-46ba-be8c-2cb64de886ec"}
print (text)
jsl_light_model = LightPipeline(jsl_ner_model)
jsl_light_result = jsl_light_model.fullAnnotate(text)
chunks = []
entities = []
sentence= []
begin = []
end = []
for n in jsl_light_result[0]['jsl_ner_chunk']:
begin.append(n.begin)
end.append(n.end)
chunks.append(n.result)
entities.append(n.metadata['entity'])
sentence.append(n.metadata['sentence'])
import pandas as pd
jsl_df = pd.DataFrame({'chunks':chunks, 'begin': begin, 'end':end,
'sentence_id':sentence, 'entities':entities})
jsl_df.head(20)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "adea1241-2f39-446c-ac72-71f3cadb8b70"}
from sparknlp_display import NerVisualizer
visualiser = NerVisualizer()
ner_vis = visualiser.display(jsl_light_result[0], label_col='jsl_ner_chunk', document_col='document', return_html=True)
displayHTML(ner_vis)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "a891f65a-75ae-488e-a155-0062b79ef023"}
# ## Posology NER
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "6154e55b-fd2d-43b8-811b-7667fbfe1b0b"}
# NER model trained on i2b2 (sampled from MIMIC) dataset
posology_ner = MedicalNerModel.pretrained("ner_posology", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")\
.setLabelCasing("upper")
ner_converter = NerConverter()\
.setInputCols(["sentence","token","ner"])\
.setOutputCol("ner_chunk")
# greedy model
posology_ner_greedy = MedicalNerModel.pretrained("ner_posology_greedy", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner_greedy")
ner_converter_greedy = NerConverter()\
.setInputCols(["sentence","token","ner_greedy"])\
.setOutputCol("ner_chunk_greedy")
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
posology_ner,
ner_converter,
posology_ner_greedy,
ner_converter_greedy])
empty_data = spark.createDataFrame([[""]]).toDF("text")
posology_model = nlpPipeline.fit(empty_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "f1dfc92d-7cd8-46f5-beb0-c73b7d8dedb9"}
posology_ner.getClasses()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "cef14918-059d-4179-b4c6-1d216cbb0e0a"}
posology_result = posology_model.transform(pubMedDF.limit(100))
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "c90f7c32-5d67-48ec-9089-c1c7297d73c7"}
posology_result.show(10)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "b0dec0f6-3b6b-428b-84d7-e65cbf877e54"}
posology_result.printSchema()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "91684cd2-e80f-42b5-a1ef-b250eb8b204f"}
from pyspark.sql.functions import monotonically_increasing_id
# This will return a new DF with all the columns + id
posology_result = posology_result.withColumn("id", monotonically_increasing_id())
posology_result.show(3)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "a2690ad2-c595-4e02-b4b0-7924e37da39d"}
posology_result.select('token.result','ner.result').show(truncate=100)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "5a00cb8a-0c23-4da4-89ed-51d6f40aba5b"}
from pyspark.sql import functions as F
posology_result_df = posology_result.select(F.explode(F.arrays_zip(posology_result.token.result, posology_result.ner.result )).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ner_label"))\
.filter("ner_label!='O'")
posology_result_df.show(20, truncate=100)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "56f479cf-2bfc-434e-8b1b-76b54a8205ba"}
posology_greedy_result_df = posology_result.select(F.explode(F.arrays_zip(posology_result.token.result,
posology_result.ner_greedy.result)).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ner_label"))\
.filter("ner_label!='O'")
posology_greedy_result_df.show(20, truncate=100)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "036d073d-9275-4617-a71b-04e22984e752"}
posology_result.select('id',F.explode(F.arrays_zip(posology_result.ner_chunk.result, posology_result.ner_chunk.begin, posology_result.ner_chunk.end,
posology_result.ner_chunk.metadata)).alias("cols")) \
.select('id', F.expr("cols['3']['sentence']").alias("sentence_id"),
F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']").alias("begin"),
F.expr("cols['2']").alias("end"),
F.expr("cols['3']['entity']").alias("ner_label"))\
.filter("ner_label!='O'")\
.show(truncate=False)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "67843214-ad73-4469-b57d-14ee4991bb0b"}
posology_result.select('id',F.explode(F.arrays_zip(posology_result.ner_chunk_greedy.result, posology_result.ner_chunk_greedy.begin,
posology_result.ner_chunk_greedy.end, posology_result.ner_chunk_greedy.metadata)).alias("cols")) \
.select('id', F.expr("cols['3']['sentence']").alias("sentence_id"),
F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']").alias("begin"),
F.expr("cols['2']").alias("end"),
F.expr("cols['3']['entity']").alias("ner_label"))\
.filter("ner_label!='O'")\
.show(truncate=False)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "cf720338-7277-449b-bd74-93a96fe02519"}
posology_result.select('ner_chunk').take(2)[1][0][0].result
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "6e2fe2e9-5891-4251-b61d-2dbd08872078"}
posology_result.select('ner_chunk').take(2)[1][0][0].metadata
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "52aefb15-d3eb-4673-a6f3-d153546e8c42"}
# ### with LightPipelines
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "04cc3646-c153-4bab-9b63-577f5833db9a"}
light_model = LightPipeline(posology_model)
text ='The patient was prescribed 1 capsule of Advil for 5 days . He was seen by the endocrinology service and she was discharged on 40 units of insulin glargine at night , 12 units of insulin lispro with meals , and metformin 1000 mg two times a day . It was determined that all SGLT2 inhibitors should be discontinued indefinitely for 3 months .'
light_result = light_model.annotate(text)
list(zip(light_result['token'], light_result['ner']))
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "a086fb49-2445-4d18-a11b-88a4563b4271"}
list(zip(light_result['token'], light_result['ner_greedy']))
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "e5f03ae7-fc68-4221-b545-7e4a52b6f56a"}
light_result = light_model.fullAnnotate(text)
chunks = []
entities = []
begin =[]
end = []
for n in light_result[0]['ner_chunk']:
begin.append(n.begin)
end.append(n.end)
chunks.append(n.result)
entities.append(n.metadata['entity'])
import pandas as pd
df = pd.DataFrame({'chunks':chunks, 'entities':entities,
'begin': begin, 'end': end})
df
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "ba948790-7bbd-4054-b010-b77aa2bff083"}
# #### NER Visualization
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "028d47a8-1a2a-4473-bc85-55c555e4e1b5"}
from sparknlp_display import NerVisualizer
visualiser = NerVisualizer()
ner_vis = visualiser.display(light_result[0], label_col='ner_chunk', document_col='document', return_html=True)
displayHTML(ner_vis)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "d1760464-8289-479d-855e-43626fadfa7c"}
# ner_greedy
visualiser_greedy = NerVisualizer()
ner_greedy_vis = visualiser_greedy.display(light_result[0], label_col='ner_chunk_greedy', document_col='document', return_html=True)
displayHTML(ner_greedy_vis)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "941abf71-9254-4510-9579-57a55d8dcc96"}
# ## Writing a generic NER function
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "938606fe-b8f1-4258-a9ed-73c69815caf3"}
def get_base_pipeline (embeddings = 'embeddings_clinical'):
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
# Sentence Detector annotator, processes various sentences per line
sentenceDetector = SentenceDetector()\
.setInputCols(["document"])\
.setOutputCol("sentence")
# Tokenizer splits words in a relevant format for NLP
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
# Clinical word embeddings trained on PubMED dataset
word_embeddings = WordEmbeddingsModel.pretrained(embeddings, "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
base_pipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings
])
return base_pipeline
def get_clinical_entities (embeddings, spark_df, nrows = 100, model_name = 'ner_clinical'):
# NER model trained on i2b2 (sampled from MIMIC) dataset
loaded_ner_model = MedicalNerModel.pretrained(model_name, "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")
ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "ner"]) \
.setOutputCol("ner_chunk")
base_pipeline = get_base_pipeline (embeddings)
nlpPipeline = Pipeline(stages=[
base_pipeline,
loaded_ner_model,
ner_converter])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
result = model.transform(spark_df.limit(nrows))
result = result.withColumn("id", monotonically_increasing_id())
result_df = result.select('id',F.explode(F.arrays_zip(result.ner_chunk.result, result.ner_chunk.begin, result.ner_chunk.end,
result.ner_chunk.metadata)).alias("cols")) \
.select('id', F.expr("cols['3']['sentence']").alias("sentence_id"),
F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']").alias("begin"),
F.expr("cols['2']").alias("end"),
F.expr("cols['3']['entity']").alias("ner_label"))\
.filter("ner_label!='O'")
return result_df
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "b352d88a-3812-467d-b28e-dec4bdd8c23f"}
embeddings = 'embeddings_clinical'
model_name = 'ner_clinical'
nrows = 100
ner_df = get_clinical_entities (embeddings, pubMedDF, nrows, model_name)
ner_df.show()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "0cbe1302-146a-4394-8da8-af6e5885b429"}
embeddings = 'embeddings_clinical'
model_name = 'ner_posology'
nrows = 100
ner_df = get_clinical_entities (embeddings, pubMedDF, nrows, model_name)
ner_df.show()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "b7796e49-23ca-44f7-9be4-0600588414c5"}
import pandas as pd
def get_clinical_entities_light (light_model, text):
light_result = light_model.fullAnnotate(text)
chunks = []
entities = []
for n in light_result[0]['ner_chunk']:
chunks.append(n.result)
entities.append(n.metadata['entity'])
df = pd.DataFrame({'chunks':chunks, 'entities':entities})
return df
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "4c9ab2f0-e142-4aee-9d8d-d875b6c4f6ab"}
text ='The patient was prescribed 1 capsule of Parol with meals . He was seen by the endocrinology service and she was discharged on 40 units of insulin glargine at night , 12 units of insulin lispro with meals , and metformin 1000 mg two times a day . It was determined that all SGLT2 inhibitors should be discontinued indefinitely fro 3 months .'
light_model = LightPipeline(posology_model)
get_clinical_entities_light (light_model, text)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "c440e27c-9781-4671-b545-2633f4788207"}
# ## PHI NER
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "74a38c17-b6af-4510-80fa-ac03c574685f"}
embeddings = 'embeddings_clinical'
model_name = 'ner_deid_large'
# deidentify_dl
# ner_deid_large
nrows = 100
ner_df = get_clinical_entities (embeddings, pubMedDF, nrows, model_name)
pd_ner_df = ner_df.toPandas()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7c6f8351-8a98-494a-bc1d-779540319b0c"}
pd_ner_df.sample(20)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "aa02ac63-790b-4001-ad7c-9eb2a79c1c5a"}
pd_ner_df.ner_label.value_counts()
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7dc27c6b-7b15-4fe0-a54c-75d3c951e7a1"}
# ## BioNLP (Cancer Genetics) NER
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "d2478dc7-9c20-41bd-bf77-9014596c4129"}
embeddings = 'embeddings_clinical'
model_name = 'ner_bionlp'
nrows = 100
ner_df = get_clinical_entities (embeddings, pubMedDF, nrows, model_name)
ner_df.show(truncate = False)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "829e86fd-8255-48f2-b62d-453c91163b69"}
# # NER Chunker
# We can extract phrases that fits into a known pattern using the NER tags. NerChunker would be quite handy to extract entity groups with neighboring tokens when there is no pretrained NER model to address certain issues. Lets say we want to extract clinical findings and body parts together as a single chunk even if there are some unwanted tokens between.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "2d5c4222-7e4b-4a1e-82ee-e4dd4c0afcb5"}
posology_ner = MedicalNerModel.pretrained("ner_posology", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")
ner_chunker = NerChunker()\
.setInputCols(["sentence","ner"])\
.setOutputCol("ner_chunk")\
.setRegexParsers(["<DRUG>.*<FREQUENCY>"])
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
posology_ner,
ner_chunker])
empty_data = spark.createDataFrame([[""]]).toDF("text")
ner_chunker_model = nlpPipeline.fit(empty_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "739f4e3e-8151-4697-9bf7-77d08a8989fc"}
posology_ner.getClasses()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "457e97d7-0fc5-43ef-ae89-95cf8cc1fd26"}
light_model = LightPipeline(ner_chunker_model)
text ='The patient was prescribed 1 capsule of Advil for 5 days . He was seen by the endocrinology service and she was discharged on 40 units of insulin glargine at night , 12 units of insulin lispro with meals , metformin 1000 mg two times a day . It was determined that all SGLT2 inhibitors should be discontinued indefinitely fro 3 months .'
light_result = light_model.annotate(text)
list(zip(light_result['token'], light_result['ner']))
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "455500e0-2c25-4b90-980b-28181025195e"}
light_result["ner_chunk"]
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "9c38e21e-86bc-4666-a7a7-c6c17e0ce94b"}
# #Chunk Filterer
# ChunkFilterer will allow you to filter out named entities by some conditions or predefined look-up lists, so that you can feed these entities to other annotators like Assertion Status or Entity Resolvers. It can be used with two criteria: isin and regex.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "ce636d1f-7cd4-45af-8c16-80be5cb533a3"}
posology_ner = MedicalNerModel.pretrained("ner_posology", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")
ner_converter = NerConverter()\
.setInputCols(["sentence","token","ner"])\
.setOutputCol("ner_chunk")
chunk_filterer = ChunkFilterer()\
.setInputCols("sentence","ner_chunk")\
.setOutputCol("chunk_filtered")\
.setCriteria("isin")\
.setWhiteList(['Advil','metformin', 'insulin lispro'])
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
posology_ner,
ner_converter,
chunk_filterer])
empty_data = spark.createDataFrame([[""]]).toDF("text")
chunk_filter_model = nlpPipeline.fit(empty_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "bd3ebc5c-5893-4bf2-b3f2-920f4e0faa4f"}
light_model = LightPipeline(chunk_filter_model)
text ='The patient was prescribed 1 capsule of Advil for 5 days . He was seen by the endocrinology service and she was discharged on 40 units of insulin glargine at night , 12 units of insulin lispro with meals , metformin 1000 mg two times a day . It was determined that all SGLT2 inhibitors should be discontinued indefinitely fro 3 months .'
light_result = light_model.annotate(text)
light_result.keys()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "b03f3fb1-ec01-4917-9536-b3c58f530603"}
light_result['ner_chunk']
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "acbfe607-af86-4183-b382-940fd8d7f646"}
light_result["chunk_filtered"]
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "55ea3fd0-594d-4279-bc58-e8d1fcf3c753"}
ner_model = MedicalNerModel.pretrained("ner_clinical", "en", "clinical/models")\
.setInputCols("sentence","token","embeddings")\
.setOutputCol("ner")
chunk_filterer = ChunkFilterer()\
.setInputCols("sentence","ner_chunk")\
.setOutputCol("chunk_filtered")\
.setCriteria("isin")\
.setWhiteList(['severe fever','sore throat'])
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
ner_model,
ner_converter,
chunk_filterer])
empty_data = spark.createDataFrame([[""]]).toDF("text")
chunk_filter_model = nlpPipeline.fit(empty_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "1e2935d4-ba38-4fb2-954a-4a6ee95aca3e"}
text = 'Patient with severe fever, severe cough, sore throat, stomach pain, and a headache.'
filter_df = spark.createDataFrame([[text]]).toDF('text')
chunk_filter_result = chunk_filter_model.transform(filter_df)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "76098683-4f06-487a-b764-b9eafde42f58"}
chunk_filter_result.select('ner_chunk.result', 'chunk_filtered.result').show(truncate=False)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "05db50e9-2bab-4deb-95ae-de242c6143e4"}
# ## Changing entity labels with `NerConverterInternal()`
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "24a1463f-1b45-420c-b389-927f8e0e4cfc"}
replace_dict = """Drug_BrandName, Drug
Frequency, Drug_Frequency
Dosage, Drug_Dosage
Strength, Drug_Strength
"""
with open('replace_dict.csv', 'w') as f:
f.write(replace_dict)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "aaa64ddd-f4e1-41d4-8c49-40eefb393bb1"}
# %fs ls file:/databricks/driver
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "0482c1ec-68e9-49e6-ac84-b640a42861cf"}
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentenceDetector = SentenceDetectorDLModel.pretrained("sentence_detector_dl_healthcare","en","clinical/models") \
.setInputCols(["document"]) \
.setOutputCol("sentence")
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
word_embeddings = WordEmbeddingsModel.pretrained("embeddings_clinical", "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
jsl_ner = MedicalNerModel.pretrained("ner_jsl", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("jsl_ner")
jsl_ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "jsl_ner"]) \
.setOutputCol("jsl_ner_chunk")
jsl_ner_converter_internal = NerConverterInternal()\
.setInputCols(["sentence","token","jsl_ner"])\
.setOutputCol("replaced_ner_chunk")\
.setReplaceDictResource("file:/databricks/driver/replace_dict.csv","text", {"delimiter":","})
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
jsl_ner,
jsl_ner_converter,
jsl_ner_converter_internal
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
ner_converter_model = nlpPipeline.fit(empty_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "742794e8-1154-4506-b497-f4a567dd5fa7"}
import pandas as pd
def get_clinical_entities_light (light_model, text, chunk_name="ner_chunk"):
light_result = light_model.fullAnnotate(text)
chunks = []
entities = []
for n in light_result[0][chunk_name]:
chunks.append(n.result)
entities.append(n.metadata['entity'])
df = pd.DataFrame({'chunks':chunks, 'entities':entities})
return df
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "8c8f0228-3c0d-46c1-b549-431f916c57a8"}
text ='The patient was prescribed 1 capsule of Parol with meals . He was seen by the endocrinology service and she was discharged on 40 units of insulin glargine at night , 12 units of insulin lispro with meals , and metformin 1000 mg two times a day . It was determined that all SGLT2 inhibitors should be discontinued indefinitely fro 3 months .'
light_model = LightPipeline(ner_converter_model)
jsl_ner_chunk_df = get_clinical_entities_light (light_model, text, chunk_name='jsl_ner_chunk')
replaced_ner_chunk_df = get_clinical_entities_light (light_model, text, chunk_name='replaced_ner_chunk')
pd.concat([jsl_ner_chunk_df, replaced_ner_chunk_df.iloc[:,1:].rename(columns= {'entities':'replaced'})], axis=1)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "f14c82b8-66fd-4900-856e-90624089c119"}
# ## Training a Clinical NER (NCBI Disease Dataset)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "276cd261-d73e-492a-829c-88db1bb34482"}
# **WARNING:** You should use TensorFlow version 2.3 for training a Clinical Ner.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "4a2fdf79-45a0-4ece-adef-6c68f8f5f339"}
# !wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/NER_NCBIconlltrain.txt
# !wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/NER_NCBIconlltest.txt
dbutils.fs.cp("file:/databricks/driver/NER_NCBIconlltest.txt", "dbfs:/")
dbutils.fs.cp("file:/databricks/driver/NER_NCBIconlltrain.txt", "dbfs:/")
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "0ed550e4-daa2-486a-9f31-3b93ab76e240"}
from sparknlp.training import CoNLL
conll_data = CoNLL().readDataset(spark, 'file:/databricks/driver/NER_NCBIconlltrain.txt')
conll_data.show(3)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "58e4d47f-88dd-4e36-9bab-03a9708b7c8e"}
conll_data.count()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "d7f07bf6-5595-4c91-8993-8d2b63cea7c6"}
from pyspark.sql import functions as F
conll_data.select(F.explode(F.arrays_zip(conll_data.token.result, conll_data.label.result)).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth")).groupBy('ground_truth').count().orderBy('count', ascending=False).show(100,truncate=False)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "8a8e883e-527c-451a-af9b-d0e7f6735ade"}
conll_data.select("label.result").distinct().count()
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "4e129512-bcca-4671-a445-b486e07c17fb"}
import pyspark.sql.functions as F
#conll_data.select(F.countDistinct("label.result")).show()
#conll_data.groupBy("label.result").count().show(truncate=False)
conll_data = conll_data.withColumn('unique', F.array_distinct(conll_data.label.result))\
.withColumn('c', F.size('unique'))\
.filter(F.col('c')>1)
conll_data.select(F.explode(F.arrays_zip(conll_data.token.result, conll_data.label.result)).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth")).groupBy('ground_truth').count().orderBy('count', ascending=False).show(100,truncate=False)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "0a814c66-180a-4fd9-a399-dc8fafd149f6"}
# Clinical word embeddings trained on PubMED dataset
clinical_embeddings = WordEmbeddingsModel.pretrained('embeddings_clinical', "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "0c7b40de-0660-4b28-bfe1-a2ff85d23dc2"}
test_data = CoNLL().readDataset(spark, "file:/databricks/driver/NER_NCBIconlltest.txt")
test_data = clinical_embeddings.transform(test_data)
test_data.write.parquet('dbfs/NER_NCBIconlltest.parquet')
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "c784547c-a240-4ae4-bff9-0bb8c6988cdc"}
# %fs ls file:/databricks/driver
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "6b74cc4f-ca46-488f-9f52-45179be7ca3e"}
# ## MedicalNER Graph
#
# **WARNING:** For training an NER model with custom graph, please use TensorFlow version 2.3
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "1b48f68b-87a8-4ea6-848d-137bce7ec487"}
import tensorflow
from sparknlp_jsl.training import tf_graph
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "8a2a3c4b-c4a2-488c-b65b-f4821dbd2511"}
tensorflow.__version__
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "37d46a82-7f95-4135-90a8-e24e971c9918"}
# Firstly, we will create graph and log folder.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7d81083f-ac04-497b-bc6a-516c75870672"}
# %fs mkdirs file:/dbfs/ner/ner_logs
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "615bf37f-88cd-4c85-8a43-7f50678074f1"}
# %fs mkdirs file:/dbfs/ner/medical_ner_graphs
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "77aedd68-7552-4b35-8a97-08ea7ddfab07"}
# %fs ls file:/dbfs/ner
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "920512a3-e09f-4d13-b324-b3d4a92cdd76"}
# We created ner log and graph files.
# Now, we will create graph and fit the model.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "9b7b3d2c-8c15-48c5-b4a4-f96e49ba631d"}
tf_graph.print_model_params("ner_dl")
tf_graph.build("ner_dl", build_params={"embeddings_dim": 200, "nchars": 85, "ntags": 12, "is_medical": 1}, model_location="/dbfs/ner/medical_ner_graphs", model_filename="auto")
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "e4867545-50e8-4265-841a-0b833f6e6f80"}
# for open source users
'''
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jupyter/training/english/dl-ner/nerdl-graph/create_graph.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jupyter/training/english/dl-ner/nerdl-graph/dataset_encoder.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jupyter/training/english/dl-ner/nerdl-graph/ner_model.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jupyter/training/english/dl-ner/nerdl-graph/ner_model_saver.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jupyter/training/english/dl-ner/nerdl-graph/sentence_grouper.py
!pip -q install tensorflow==1.15.0
import create_graph
ntags = 3 # number of labels
embeddings_dim = 200
nchars =83
create_graph.create_graph(ntags, embeddings_dim, nchars)
'''
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "c316cb83-0440-4942-beec-a53ba2a8d573"}
# %fs ls file:/dbfs/ner/medical_ner_graphs/
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "a12b3d18-e3c2-4d41-aae1-79f5e48ce7f3"}
nerTagger = MedicalNerApproach()\
.setInputCols(["sentence", "token", "embeddings"])\
.setLabelColumn("label")\
.setOutputCol("ner")\
.setMaxEpochs(2)\
.setBatchSize(64)\
.setRandomSeed(0)\
.setVerbose(1)\
.setValidationSplit(0.2)\
.setEvaluationLogExtended(True) \
.setEnableOutputLogs(True)\
.setIncludeConfidence(True)\
.setTestDataset("/NER_NCBIconlltest.parquet")\
.setOutputLogsPath('dbfs:/ner/ner_logs')\
.setGraphFolder('dbfs:/ner/medical_ner_graphs') # control your graph folder
ner_pipeline = Pipeline(stages=[
clinical_embeddings,
nerTagger
])
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "758948a8-a070-461f-bfc4-30fdc685269c"}
# 2 epochs 39 sec
ner_model = ner_pipeline.fit(conll_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "f7ca7dab-3e6f-4471-96d0-02c28a32d335"}
# %sh cd /dbfs/ner/ && ls -la
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "880203c2-cfe4-431e-bf2a-a2d0badb9cd1"}
# %sh cd /dbfs/ner/ner_logs && ls -lt
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "c92d2fe8-8eb9-412a-840f-972d17cc09bf"}
# %sh cat /dbfs/ner/ner_logs/MedicalNerApproach_73f4f0678ff4.log
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "5433c79b-6437-4f17-99d1-467d7e66001d"}
# ### Evaluate your model
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "011ba1cd-6135-4e5b-b741-a5b7d6f3e7e6"}
pred_df = ner_model.stages[-1].transform(test_data)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "cefc4714-a914-40a8-8003-9a0f2e22406b"}
pred_df.columns
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "afbd7dc9-e258-41f9-b393-27cbfa2e7f0c"}
from sparknlp_jsl.eval import NerDLMetrics
import pyspark.sql.functions as F
evaler = NerDLMetrics(mode="full_chunk", dropO=True)
eval_result = evaler.computeMetricsFromDF(pred_df.select("label","ner"), prediction_col="ner", label_col="label").cache()
eval_result.withColumn("precision", F.round(eval_result["precision"],4))\
.withColumn("recall", F.round(eval_result["recall"],4))\
.withColumn("f1", F.round(eval_result["f1"],4)).show(100)
print(eval_result.selectExpr("avg(f1) as macro").show())
print (eval_result.selectExpr("sum(f1*total) as sumprod","sum(total) as sumtotal").selectExpr("sumprod/sumtotal as micro").show())
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7a9469fb-15be-46e1-b16f-aaf12a47b908"}
evaler = NerDLMetrics(mode="partial_chunk_per_token", dropO=True)
eval_result = evaler.computeMetricsFromDF(pred_df.select("label","ner"), prediction_col="ner", label_col="label").cache()
eval_result.withColumn("precision", F.round(eval_result["precision"],4))\
.withColumn("recall", F.round(eval_result["recall"],4))\
.withColumn("f1", F.round(eval_result["f1"],4)).show(100)
print(eval_result.selectExpr("avg(f1) as macro").show())
print (eval_result.selectExpr("sum(f1*total) as sumprod","sum(total) as sumtotal").selectExpr("sumprod/sumtotal as micro").show())
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "0b0192f1-2002-4b7a-b18e-daf956744335"}
ner_model.stages[1].write().overwrite().save('dbfs:/databricks/driver/models/custom_Med_NER_2e')
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "ed434753-3fd1-443a-8c8a-90d1e28b649e"}
document = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentence = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentence')
token = Tokenizer()\
.setInputCols(['sentence'])\
.setOutputCol('token')
loaded_ner_model = MedicalNerModel.load("dbfs:/databricks/driver/models/custom_Med_NER_2e")\
.setInputCols(["sentence", "token", "embeddings"])\
.setOutputCol("ner")
converter = NerConverter()\
.setInputCols(["document", "token", "ner"])\
.setOutputCol("ner_span")
ner_prediction_pipeline = Pipeline(
stages = [
document,
sentence,
token,
clinical_embeddings,
loaded_ner_model,
converter])
empty_data = spark.createDataFrame([['']]).toDF("text")
prediction_model = ner_prediction_pipeline.fit(empty_data)
from sparknlp.base import LightPipeline
light_model = LightPipeline(prediction_model)
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "d6fc3f2e-2dfa-425d-94f8-e9e40e384a93"}
text = "She has a metastatic breast cancer to lung"
def get_preds(text, light_model):
result = light_model.fullAnnotate(text)[0]
return [(i.result, i.metadata['entity']) for i in result['ner_span']]
get_preds(text, light_model)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "af43603e-4b9b-40bf-8865-694a2d27b1d1"}
# End of Notebook #
|
tutorials/Certification_Trainings/Healthcare/databricks_notebooks/2. Training and Reusing Clinical Named Entity Recognition Models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# Autorzy: <NAME>, <NAME>
#
# # Na jakie dane będziemy zwracać uwagę?
#
# Ponieważ naszym zadaniem jest skonstruowanie systemu rekomendującego **produkty użytkownikom na podstawie ich sesji**,
# będziemy potrzebowali danych użytkowników obecnych w systemie sklepu eSzoppping, wraz z ich listą interakcji.
#
# ---
#
# ### Dane użytkowników
#
# Dane użytkowników zawarte są w pliku **users.jsonl**.
#
# Pojedynczy rekord opisujący użytkownika składa się z następujących kolumn:
# - id użytkownika w systemie
# - imię użytkownika
# - miasto zamieszkania
# - miejsce zamieszkania
#
# Przypuszczalnie, nie wszystkie informacje zawarte w rekordach będą dla nas użyteczne. Dalsza analiza oceni przydatność poszczególnych atrybutów.
#
# ---
#
# ### Dane produktów
#
# Dane związane z produktami zawarte są w pliku **products.jsonl**.
#
# Pojedynczy rekord opisujący produkt składa się z następujących kolumn:
# - id produktu w systemie
# - nazwa produktu
# - kategoria przynależności produktu
# - cena produktu
# - ocena produktu
#
# Przypuszczalnie, nie wszystkie informacje zawarte w rekordach będą dla nas użyteczne. Dalsza analiza oceni przydatność poszczególnych atrybutów.
#
# ---
#
#
# ### Dane sesji
#
# Dane łączące użytkowników z produktami zawarte są w pliku **sessions.jsonl**.
# Plik ten zawiera rekordy opisujące aktywności użytkowników w obrębie strony eSzoppping.
#
# Pojedynczy rekord opisujący akcję użytkownika w systemie składa się z następujących kolumn:
# - id sesji w systemie
# - punkt w czasie odbycia się aktywności
# - id użytkownika, którego dotyczy wydarzenie
# - id produktu, którego dotyczy wydarzenie
# - typ wydarzenia
# - informacja o oferowanej zniżce
# - id dokonanego zakupu
#
# Aby wydobyć "pełnię" informacji o każdej z sesji, będziemy musieli uwzględnić pliki opisujące **produkty i użytkowników** w analizie danych sesji.
#
# Przypuszczalnie, nie wszystkie informacje zawarte w rekordach będą dla nas użyteczne. Dalsza analiza oceni przydatność poszczególnych atrybutów.
#
# ---
#
# ### Nieistotne dane
#
# Dane, których nie użyjemy w trakcie analizy, to te zawarte w pliku **deliveries.jsonl**. Plik ten zawiera rekordy opisujące zdarzenia dowozu produktu do kupującego. Zakładamy, iż dane te są nieistotne z punktu widzenia interakcji użytkownika ze sklepem internetowym.
# + pycharm={"name": "#%%\n"}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# Used for verbose data presentation.
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
pd.set_option('max_colwidth', None)
users_data_path = '../data/users.jsonl'
products_data_path = '../data/products.jsonl'
sessions_data_path = '../data/sessions.jsonl'
users_df = pd.read_json(users_data_path, lines=True)
sessions_df = pd.read_json(sessions_data_path, lines=True)
products_df = pd.read_json(products_data_path, lines=True)
# + [markdown] pycharm={"name": "#%% md\n"}
# # Analiza samodzielnych danych
#
# Teraz dokładnie przyjrzymy się danym, które mogą być analizowane samodzielnie. Zbadamy typy ich atrybutów, rozkłady oraz podejmiemy pierwsze decyzje modelowania danych.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Dane produktów
#
# Aby mieć lepsze "poczucie" analizowanych danych, poniżej prezentujemy 10 pierwszych rekordów obecnych w dostarczonych danych.
# + pycharm={"name": "#%%\n"}
products_df.head(n=10)
# + [markdown] pycharm={"name": "#%% md\n"}
# Już z tych przykładowych danych możemy wyciągnąć kilka interesujących wniosków. Po pierwsze, ceny produktów mogę w sposób znaczący różnić się między sobą (58.97 kontra 7639.00). Po drugie, nazwa produktu wydaje się zbędna w momencie posiadania jego identyfikatora (jest ona po prostu kolejnym unikatowym ciągiem znaków). Po trzecie, i to jest najbardziej istotny wniosek, kolumna **category_path** zawiera informację o kategorii produktu w formie "doprecyzowywania" tzn. kategorie rozdzielane są znakiem **;** a najistotniejsza kategoria umieszczona jest na samym początku napisu.
# + pycharm={"name": "#%%\n"}
products_df.info()
# + [markdown] pycharm={"name": "#%% md\n"}
# Widzimy, iż dane produktów nie zawierają żadnych brakujących wartości.
#
# Dane produktów zawierają dwie kolumny o typie napisowym (product_name oraz category_path), jedną o typie całkowitym (product_id) oraz dwie o typie zmiennoprzecinkowym (price i user_rating).
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Identyfikatory produktu
#
# W celu lepszego zaznajomienia się z danymi opisującymi identyfikator produktu dokonujemy dalszej analizy kolumny **product_id**.
# + pycharm={"name": "#%%\n"}
print('Max id of the product is: {}'.format(products_df['product_id'].max()))
print('Min id of the product is: {}'.format(products_df['product_id'].min()))
print('Distinct id count is: {}'.format(products_df['product_id'].nunique()))
# + [markdown] pycharm={"name": "#%% md\n"}
# Jak widzimy, minimalnym identyfikatorem produktu jest wartość **1001** a maksymalnym wartość **1319**. Ponadto wszystkie wartości pomiędzy **1001 a 1319** są osiągane dokładnie 1 raz.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Nazwy produktu
#
# Następnie analizie poddamy kolumnę opisującą nazwę produktu **product_name**.
# + pycharm={"name": "#%%\n"}
print('Distinct name count is: {}'.format(products_df['product_name'].nunique()))
# + [markdown] pycharm={"name": "#%% md\n"}
# Widzimy, iż w naszych danych nie występują produkty o tych samych nazwach.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Ścieżka kategorii
#
# Teraz analizie poddamy kolumnę opisującą ścieżkę kategorii produktu **category_path**. W tym celu przyjrzymy się dostępnym ścieżkom kategorii oraz liczności każdej z nich tzn. ile produktów należy do danej ścieżki.
# + pycharm={"name": "#%%\n"}
print(products_df['category_path'].drop_duplicates().reset_index(drop=True).sort_values().to_string(index=False))
# + [markdown] pycharm={"name": "#%% md\n"}
# Zauważamy, że ścieżki kategorii tworzą hierarchię. Każda kategoria ma swój korzeń, który jest rozwijany poprzez ewentualne dopisywanie nowych podkategorii.
# + pycharm={"name": "#%%\n"}
products_df.groupby('category_path')['product_id'].count().sort_values(ascending=False)
# + [markdown] pycharm={"name": "#%% md\n"}
# Powyższy wynik ukazuje nam, że liczba produktów w kategoriach nie jest rozłożona w sposób równomierny. Niektóre kategorie posiadają jedynie jeden zakwalifikowany produkt, podczas gdy inne zawierają ich aż kilkadziesiąt lub kilkaset. Proponowanym rozwiązaniem będzie zmniejszenie liczby dostępnych kategorii poprzez rzutowanie danych na odpowiedni korzeń hierarchii kategorii.
#
# Poprzez rzutowanie moglibyśmy otrzymać następujące wyniki:
# - "Gry komputerowe": 202
# - "Gry na konsole": 41 (Xbox and PS3)
# - "Sprzęt RTV": 36
# - "Komputery": 28
# - "Telefony i akcesoria": 12
#
# Liczba i liczność wytworzonych kategorii wydaje się zadowalająca na ten moment.
# + pycharm={"name": "#%%\n"}
separator = ';'
new_groups = ['Gry komputerowe', 'Gry na konsole', 'Sprzęt RTV', 'Komputery', 'Telefony i akcesoria']
def cast_category_path(category_path):
categories = category_path.split(separator)
found_groups = [group for group in new_groups if group in categories]
if len(found_groups) != 1:
raise RuntimeError('wrong group cast: {}'.format(found_groups))
return found_groups[0]
transformed = products_df['category_path'].apply(cast_category_path)
print(transformed.value_counts())
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Cena produktu
#
# Poniżej przedstawiamy histogram prezentujący rozkład wartości cen zawartych w kolumnie **price**.
# + pycharm={"name": "#%%\n"}
print('Max price is: {}'.format(products_df['price'].max()))
print('Min price is: {}'.format(products_df['price'].min()))
sns.displot(data=products_df, x="price", kind="kde")
# + pycharm={"name": "#%%\n"}
sns.violinplot(data=products_df, x='price')
# + [markdown] pycharm={"name": "#%% md\n"}
# Widzimy, że zdecydowana większość cen mieści się poniżej kwoty 100. Niestety, wartości odstające zakłócają wygląd wykresów. Decydujemy się rozbić produkty na tańsze i droższe. Jako punkt podziału przyjmujemy wartość 100.
# + pycharm={"name": "#%%\n"}
threshold = 100
cheap_products = (products_df.loc[products_df['price'] < threshold], 'Cheap')
expensive_products = (products_df.loc[products_df['price'] >= threshold], 'Expensive')
products_split = [cheap_products, expensive_products]
for ps in products_split:
print('{} products amount is: {}'.format(ps[1], ps[0]['price'].count()))
sns.displot(data=ps[0], x="price", kind="kde")
# + [markdown] pycharm={"name": "#%% md\n"}
# Jak widzimy, wartości zmiennej opisującej ceny produktu nie są rozłożone w sposób równomierny. Przypominają jednak złożenia kilku rozkładów normalnych, których wartości oczekiwane i wariancje są różne. Interesujące wydają nam się również produkty z ceną 1.0. Postanawiamy przyjrzeć się rekordom z taką ceną.
# + pycharm={"name": "#%%\n"}
products_df.loc[products_df["price"] == products_df["price"].min()]
# + [markdown] pycharm={"name": "#%% md\n"}
# Powyższe pozycje, choć zastanawiające, są możliwe.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Ocena produktu
#
# Poniżej prezentujemy wykres przedstawiający rozkład wartości zmiennych opisujących opinie o produkcie zawartych w kolumnie **user_rating**.
# + pycharm={"name": "#%%\n"}
print('Max rating is: {}'.format(products_df['user_rating'].max()))
print('Min rating is: {}'.format(products_df['user_rating'].min()))
sns.displot(data=products_df, x='user_rating', kind='kde')
# + [markdown] pycharm={"name": "#%% md\n"}
# Jak widać, opinie o produktach rozłożone są w sposób bardziej równomierny niż ceny produktów. Ponadto, wszystkie wartości z kolumny **user_rating** zawarte są w przedziale (0,5).
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Dane użytkowników
#
# Podobnie jak w przypadku analizy danych o produktach, przyjrzyjmy się przykładowym wpisom w dostępnych danych:
# + pycharm={"name": "#%%\n"}
users_df.head(n=10)
# + [markdown] pycharm={"name": "#%% md\n"}
# Widzimy, że praktycznie wszystkie kolumny posiadają typ napisowy (**name, city, street**). Ponadto, kolumna **street** w rzeczywistości zawiera adres użytkownika, a nie ulicę, na której mieszka. Pozostałe kolumny wydają się standardowe w kontekście zawieranych danych.
# + pycharm={"name": "#%%\n"}
users_df.info()
# + [markdown] pycharm={"name": "#%% md\n"}
# Widzimy, że dane użytkowników nie zawierają żadnych brakujących wartości. Wszystkie wpisy są kompletne.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Identyfikator użytkownika
#
# Teraz przyjrzymy się wartościom zawartym w kolumnie **user_id**. Interesuje nas, czy wszystkie wartości są unikalne. Ponadto dobrze byłoby znać zakres identyfikatorów oraz czy jest on w pełni zapełniony.
# + pycharm={"name": "#%%\n"}
print('Max user id is: {}'.format(users_df['user_id'].max()))
print('Min user id is: {}'.format(users_df['user_id'].min()))
print('Unique identifiers number is: {}'.format(users_df['user_id'].nunique()))
# + [markdown] pycharm={"name": "#%% md\n"}
# Z powyższych wyników wnioskujemy, że zakres identyfikatorów wynosi [102, 301]. Ponadto, każda wartość w tym zakresie przyjmowana jest dokładnie jeden raz.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Imię użytkownika
#
# Następną kolumną, jaką poddamy dalszej analizie, jest ta zawierająca informacje o imieniu użytkownika. Jesteśmy zainteresowani, czy w dostępnych danych istnieje duplikacja użytkownika (różne identyfikatory, ale to samo imię i nazwisko).
# + pycharm={"name": "#%%\n"}
print('Unique name count is: {}'.format(users_df['name'].nunique()))
# + [markdown] pycharm={"name": "#%% md\n"}
# Jak widać, wszystkie imiona użytkowników są unikatowe. Dzięki temu jesteśmy w stanie stwierdzić, że w zbiorze danych nie występują potencjalne anomalie. Poza tym zakładamy, że kolumna o nazwie **name** nie wnosi żadnych nowych informacji do modelu i rozwiązywanego zagadnienia.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### <NAME>
#
# Kolumna zawierająca miasto zamieszkania użytkownika (zakładamy, że jest to miejsce zamieszkania, jednak nie mamy tutaj żadnej pewności co do tego) wydaje się nieść potencjalnie wiele przydatnych informacji. Przede wszystkim dzięki niej jesteśmy w stanie stwierdzić, czy zbiór użytkowników, na których operujemy, jest reprezentatywny. Ponadto potencjalnie może zachodzić korelacja pomiędzy miastem zamieszkania a zainteresowaniem pewnymi produktami przez grupy użytkowników. Poniżej prezentujemy histogram z uzyskanymi wynikami.
# + pycharm={"name": "#%%\n"}
sns.catplot(data=users_df, kind='count', x='city', height=8)
# + [markdown] pycharm={"name": "#%% md\n"}
# Z powyższego histogramu widzimy, iż pomimo pewnych nieregularności, dane są rozłożone dość równomiernie. Dzięki temu jesteśmy w stanie stwierdzić, że dane, na których pracujemy, są reprezentatywne
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Miejsce zamieszkania
#
# Ostatnią kolumną do przeanalizowania jest ta zawierająca dane o adresach użytkowników. Zakładamy, iż prawdopodobnie kolumna to nie będzie dla nas w żaden sposób informatywna. Niemniej jednak jesteśmy zainteresowani, czy w danych nie występuje duplikacja adresu.
# + pycharm={"name": "#%%\n"}
print('Unique address count is: {}'.format(users_df['street'].nunique()))
# + [markdown] pycharm={"name": "#%% md\n"}
# Jak widać, wszystkie adresu są unikatowe.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Dane sesji
#
# Przyjrzyjmy się nieco bliżej danym sesji.
# + pycharm={"name": "#%%\n"}
sessions_df.head(10)
# + [markdown] pycharm={"name": "#%% md\n"}
# Widzimy, że dane sesji nie zawierają bezpośrednio informacji o użytkownikach i produktach, a jedynie ich unikalne identyfikatory. Stwierdzamy, iż w celu dokonania lepszej analizy powinniśmy dołączyć do danych sesji odpowiadające dane z tabeli o użytkownikach i produktach.
#
# W tej sekcji postanowiliśmy odpowiedzieć sobie na kilka pytań istotnych z punktu widzenia zadania:
# - Czy istnieje użytkownik nieposiadający żadnej sesji?
# - Czy istnieje produkt, który nie został przez nikogo wyświetlony?
# - Czy dane użytkowników i produktów są pełne, tj. czy w danych sesji jest produkt bądź użytkownik, który nie widnieje w innych dostarczonych danych?
#
# Przy łączeniu danych wykorzystaliśmy operację **outer join**, by wszelkie braki w danych były proste do wychwycenia.
# + pycharm={"name": "#%%\n"}
# Use outer join in order to keep all values from data frames.
merged_df = pd.merge(sessions_df, users_df, how='outer', on='user_id')
merged_df = pd.merge(merged_df, products_df, how='outer', on='product_id')
print(sessions_df.shape[0] == merged_df.shape[0])
merged_df.info()
# + [markdown] pycharm={"name": "#%% md\n"}
# Stwierdzamy, iż dane nie zawierają anomalii (brak nieznanych użytkowników i produktów).
# Dodatkowo obserwujemy, że użytkownicy częściej przeglądają produkty, niż je kupują. Stosunek obu rodzajów aktywności wynosi 105358:10692.
# + [markdown] pycharm={"name": "#%% md\n"}
# Obszary, którym przyjrzymy się nieco dokładniej w dalszej części:
# - session_id — stwierdzenie liczby unikalnych sesji. Sprawdzenie, czy nie występują błędne dane (np. ujemne identyfikatory)
# - timestamp — sprawdzenie przedziału czasu, z jakiego pochodzą dane
# - analiza danych sesji — jakie są najpopularniejsze kategorie, produkty itp.
# - gęstość macierzy interakcji
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Identyfikator sesji
#
# Poszukujemy błędów w atrybucie **session_id**, takich jak wartości ujemne czy duplikacja.
# + pycharm={"name": "#%%\n"}
print(f"Minimum session id: {sessions_df['session_id'].min()}")
print(f"Maximum session id: {sessions_df['session_id'].max()}")
print(f"Number of unique sessions: {sessions_df['session_id'].nunique()}")
# + [markdown] pycharm={"name": "#%% md\n"}
# Wartości w kolumnie **session_id** są poprawne. Liczba 19603 unikalnych sesji jest również pokaźnym zbiorem danych.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Sprawdzenie przedziału czasowego
#
# Dla naszego problemu istotne jest sprawdzenie, czy dane pochodzą z odpowiednio długiego okresu. Uważamy, że minimalny przedział pokryty przez dane powinien wynosić 1 rok. Dodatkowo zweryfikujemy czy okres pokryty jest w całości (tzn. czy nie występują okresy z brakiem danych).
# + pycharm={"name": "#%%\n"}
session_by_day_df = sessions_df.groupby(pd.Grouper(key='timestamp', axis=0, freq='1D', sort=True))
print(f'Days count in data: {session_by_day_df.size().count()}')
print(f'first date: {session_by_day_df.count().head(1)}\nlast date: {session_by_day_df.count().tail(1)}')
# + [markdown] pycharm={"name": "#%% md\n"}
# Dane pochodzą z 2 lat, a dokładnie z okresu od **11.12.2019 do 10.12.2022**, w danych nie występują przerwy. Oznacza to, że dane są reprezentatywne i mogą być wykorzystane do rozwiązania zadania.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Popularność produktów i kategorii
#
# Poniżej sprawdzamy, jak często każdy z produktów występuję w danych. Można rozumieć to jako analizę "popularności" produktów w sklepie.
# + pycharm={"name": "#%%\n"}
sns.catplot(data=sessions_df, kind='count', x='product_id', height=8)
# + [markdown] pycharm={"name": "#%% md\n"}
# Jak widać, w częstości interakcji z produktami istnieje duża dysproporcja. Widzimy, że jedynie niewielki odsetek produktów przyciąga znaczną uwagę użytkowników, podczas gdy większość produktów nie budzi większego zainteresowanie.
#
# Co istotne, każdy z produktów został obejrzany przez choć jednego użytkownika.
#
# Zainteresowało nas, jakie dokładnie produkty są najpopularniejsze i do jakich kategorii należą. Za poziom odcięcia przyjęliśmy 95 percentyl.
# + pycharm={"name": "#%%\n"}
sub_session_df = sessions_df.groupby('product_id').filter(lambda x : len(x) > sessions_df['product_id'].value_counts().quantile(q=0.95)).copy()
sub_session_product_df = pd.merge(sub_session_df, products_df, how='inner', on='product_id')
sns.catplot(data=sub_session_product_df, kind='count', x='product_name', height=8, aspect=2.2)
plt.xticks(rotation=90) # Rotate labels to make plot more readable
sns.catplot(data=sub_session_product_df, kind='count', x='category_path', height=8, aspect=2.2)
top_products_df = products_df.loc[products_df['product_id'].isin(sub_session_df['product_id'].unique())]
print(top_products_df['user_rating'].agg(['count', 'mean', 'max', 'min']))
top_products_df
# + [markdown] pycharm={"name": "#%% md\n"}
# Okazuje się, że najpopularniejszymi produktami w sklepie są **Gry komputerowe**. Drugą najczęściej występującą kategorią są **Gry na konsolę**.
#
# Bardzo popularne okazały się produkty o bardzo niskiej ocenie np. **Gra Call of Duty Black Ops**: ~4000 interakcji, ocena ~0.4.
#
# Wśród 16 najpopularniejszych produktów średnia ocena wynosi ~3.1.
#
# Spojrzymy teraz całościowo na sesje i przeanalizujemy, jakie są najczęściej oglądane i kupowane kategorie produktów. Ze względu, że "pełna kategoria" wprowadza zbyt dużą gradację, na potrzeby tej analizy kategorie zostały zredukowane do 5 (zaproponowanych wcześniej podczas analizy atrybutu **category_path**).
#
# Dane sesji podzieliśmy na dwa rozłączne podzbiory, w jednym znajdują się tylko obejrzane produkty, w drugim tylko produkty kupione (**event_type** odpowiednio **VIEW_PRODUCT** i **BUY_PRODUCT**).
# + pycharm={"name": "#%%\n"}
session_product_df = pd.merge(sessions_df, products_df, how='inner', on='product_id')
no_buy_sessions = session_product_df.loc[session_product_df['event_type'] == 'VIEW_PRODUCT'].copy()
buy_sessions = session_product_df.loc[session_product_df['event_type'] == 'BUY_PRODUCT'].copy()
# Reduce the number of product categories
no_buy_sessions['category_path'] = no_buy_sessions['category_path'].apply(cast_category_path)
buy_sessions['category_path'] = buy_sessions['category_path'].apply(cast_category_path)
sns.catplot(data=no_buy_sessions, kind='count', x='category_path', height=8)
plt.title("Sessions with no buy action")
sns.catplot(data=buy_sessions, kind='count', x='category_path', height=8)
plt.title("Sessions with buy action")
# + [markdown] pycharm={"name": "#%% md\n"}
# Jak widzimy, zarówno najczęściej oglądaną kategorią, jak i najchętniej kupowaną są **Gry komputerowe**. Następnie praktycznie na równej pozycji znajdują się **Gry na konsole**, **Sprzęt RTV** i **Komputery**. Pomiędzy tą trójką a faworytem istnieje jednak znacząca przepaść. Na ostatniej pozycji znajdziemy kategorię **Telefony i akcesoria**.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Współczynnik informacji wzajemnej
#
# Informacyjność danych jest istotna dla każdego problemu modelowania. Dla nas szczególnie istotne jest sprawdzenie, czy istnieje jakiś związek pomiędzy użytkownikiem a przeglądanymi produktami.
#
# Skalę makroskopową, tzn. czy użytkownik posiada jakieś indywidualne preferencje możemy sprawdzić licząc współczynnik informacji wzajemnej dla atrybutów **user_id** oraz **category_path**.
# Współczynnik policzyliśmy dla czterech formatów danych:
# - sesje oglądania produktu, ścieżka kategorii rzutowana do 5 kategorii głównych
# - sesje oglądania produktu, pełna ścieżka kategorii
# - sesje kupowania produktu, ścieżka kategorii rzutowana do 5 kategorii głównych
# - sesje kupowania produktu, pełna ścieżka kategorii
# + pycharm={"name": "#%%\n"}
from sklearn import metrics
no_buy_session_product_df = session_product_df.drop(session_product_df.index[session_product_df['event_type'] == 'BUY_PRODUCT']).copy()
buy_session_product = session_product_df.dropna(subset=['purchase_id']).copy()
cast_no_buy_session_product_df = no_buy_session_product_df.copy()
cast_no_buy_session_product_df['category_path'] = no_buy_session_product_df['category_path'].apply(cast_category_path)
cast_buy_session_product_df = buy_session_product.copy()
cast_buy_session_product_df['category_path'] = buy_session_product['category_path'].apply(cast_category_path)
print(f"Mutual information score for user_id and casted category_path (view events): {metrics.normalized_mutual_info_score(cast_no_buy_session_product_df['user_id'], cast_no_buy_session_product_df['category_path'])}")
print(f"Mutual information score for user_id and category_path (view events): {metrics.normalized_mutual_info_score(no_buy_session_product_df['user_id'], no_buy_session_product_df['category_path'])}")
print(f"Mutual information score for user_id and casted category_path (buy events): {metrics.normalized_mutual_info_score(cast_buy_session_product_df['user_id'], cast_buy_session_product_df['category_path'])}")
print(f"Mutual information score for user_id and category_path (buy events): {metrics.normalized_mutual_info_score(buy_session_product['user_id'], buy_session_product['category_path'])}")
# + [markdown] pycharm={"name": "#%% md\n"}
# Wyznaczony współczynnik informacji wzajemnej (znormalizowany) ma bardzo niską wartość. Tylko jeden współczynnik ma wartość powyżej 0.02 i wynosi on ~0.036. Co ciekawe, okazuje się on być wyższy dla bardziej szczegółowych kategorii.
#
# Po dłuższym zastanowieniu doszliśmy do wniosku, że nie jesteśmy pewni czy wskaźnik ten powinien mieć wysoką wartość. Na ten moment przyjmujemy te dane takimi, jake są i ocenimy je w późniejszym etapie.
# Kolejnym pomysłem było sprawdzenie, czy istnieje zależność między ilością interakcji (popularnością) z produktem a jego oceną.
# Tym razem, ze względu na ciągłość atrybutów i chęć wychwycenia bardziej liniowej zależności, policzyliśmy korelację liniową.
# Podobnie jak wcześniej policzyliśmy ją oddzielnie dla sesji przeglądania i zakupowych.
# + pycharm={"name": "#%%\n"}
no_buy_session_product_df['user_rating'] = no_buy_session_product_df['user_rating'].apply(lambda x : round(x, 1))
no_buy_prod_count_series = no_buy_session_product_df.groupby(['product_id'])['user_id'].count().copy()
no_buy_rating_series = no_buy_session_product_df.groupby(['product_id'])['user_rating'].unique().copy()
no_buy_rating_series = no_buy_rating_series.apply(lambda x : x[0])
print(f"Linear correlation between popularity of a product and user_rating (view events): {no_buy_prod_count_series.corr(no_buy_rating_series)}")
buy_session_product['user_rating'] = buy_session_product['user_rating'].apply(lambda x : round(x, 1))
buy_prod_count_series = buy_session_product.groupby(['product_id'])['user_id'].count().copy()
buy_rating_series = buy_session_product.groupby(['product_id'])['user_rating'].unique().copy()
buy_rating_series = buy_rating_series.apply(lambda x : x[0])
print(f"Linear correlation between popularity of a product and user_rating (buy events): {buy_prod_count_series.corr(buy_rating_series)}")
# + [markdown] pycharm={"name": "#%% md\n"}
# Okazuje się, że i w tym wypadku obserwujemy dość słabą korelację. Dla zdarzeń przeglądania jest ona bliska zeru. Początkowo zakładaliśmy, że spodziewamy się tutaj silnej korelacji. Po przemyśleniach uważamy, że nie musi to być prawdą.
# Ocena produktu jest dobrowolna, stąd nie każdy użytkownik się na nią decyduje. Dodatkowo użytkownicy mogą przy swoich wyborach kierować się innymi (np. zewnętrznymi) kryteriami, a nie średnią oceną produktu w sklepie eSzoppping.
# Wszystko to sprawia, że ocena nie musi być reprezentatywna co do popularności produktu.
#
# Korelacja dla zdarzeń zakupu jest wielokrotnie wyższa, co sugeruje, że ocena produktu w sklepie ma jakiś wpływ przy decyzji zakupowej użytkownika.
# + [markdown] pycharm={"name": "#%% md\n"}
# Postanowiliśmy jeszcze sprawdzić, czy zachodzi korelacja liniowa pomiędzy ilością interakcji (popularnością) z danym produktem a jego ceną. Jak wcześniej współczynnik, policzyliśmy niezależenie dla sesji przeglądania i zakupowych.
# + pycharm={"name": "#%%\n"}
no_buy_price_series = no_buy_session_product_df.groupby(['product_id'])['price'].unique().copy()
no_buy_price_series = no_buy_price_series.apply(lambda x : x[0])
print(f"Linear correlation between popularity of a product and price (view events): {no_buy_prod_count_series.corr(no_buy_price_series)}")
buy_price_series = buy_session_product.groupby(['product_id'])['price'].unique().copy()
buy_price_series = buy_price_series.apply(lambda x : x[0])
print(f"Linear correlation between popularity of a product and price (buy events): {buy_prod_count_series.corr(buy_price_series)}")
# + [markdown] pycharm={"name": "#%% md\n"}
# Wyznaczone współczynniki ponownie są bliskie zeru.
# Sugeruje to, że w przypadku użytkowników sklepu eSzoppping cena produktu nie gra większej roli, zarówno przy oglądaniu produktów, jak i ich zakupie.
# Oznacza to, że ten argument jest potencjalnie zbędny przy tworzeniu modelu.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Badanie gęstości macierzy interakcji
# Postanowiliśmy sprawdzić również, jak wygląda gęstość otrzymanych danych dla zadania predykcji. Utworzyliśmy macierz, której kolumny odpowiadają wszystkim produktom w ofercie, a wiersze poszczególnym użytkownikom.
# Jeżeli użytkownik wszedł w interakcje z danym produktem (obejrzał go lub kupił), w odpowiadającej komórce będzie 1, w przeciwnym wypadku komórka pozostanie pusta. Dla tak utworzonej macierzy policzyliśmy gęstość.
# + pycharm={"name": "#%%\n"}
df = sessions_df.drop(columns=["session_id", "timestamp", "event_type", "offered_discount", "purchase_id"])
df["hit"] = 1
heat_map_df = pd.pivot_table(df, index="user_id", columns="product_id", values="hit")
print("Sparse matrix density: " + str((heat_map_df.size - heat_map_df.isna().sum().sum()) / heat_map_df.size))
# + [markdown] pycharm={"name": "#%% md\n"}
# Gęstość powyższej macierzy wynosi ~44,8%, co jest wartością wystarczającą dla zadania rekomendacji.
# + [markdown] pycharm={"name": "#%% md\n"}
# # Podsumowanie
#
# Podsumowując powyższą analizę:
# * W dostarczonych danych nie wykryto oczywistych błędów, takich jak: braki wartości, błędne wartości atrybutów. Jedyne co zwróciło naszą uwagę to trzy produkty, których koszt wynosi 1.
# * Dostarczone dane pochodzą z okresu 24 miesięcy, co daje nam reprezentatywny przedział czasowy.
# * Kategorie produktów są bardzo nierównomierne. Nawet po zredukowaniu ilości grup do 5 najgorszy współczynnik niezbilansowania wynosi ~1:14. Możemy próbować temu zaradzić, rozbijając kategorię większościową (Gry komputerowe), na mniejsze np. na gatunki gier. Ostateczną decyzję podejmiemy poprzez analizę zachowanie modeli.
# * W danych zachodzi również duża nierównomierność w popularności produktów mierzonej jako liczba interakcji. Podobnie sytuacja wygląda z kategoriami, widzimy bezpośrednią korelację między liczebnością grupy (zredukowanej), a jej popularnością.
# * Współczynnik informacji wzajemnej między *user_id*, a *category_path* pokazał, że użytkownicy nie wydają się mieć z góry określony preferencji.
# * Współczynniki korelacji liniowej dla ilości interakcji z danym produktem, a jego oceną są bliskie zeru, podobnie jak dla współczynnika między ilością interakcji a ceną produkty.
# Oznacza to, że te atrybuty te są dla nas mało interesujące (niska informacyjność).
# * Policzone współczynniki sugerują brak spodziewanych zależności. Nie oznacza to koniecznie błędów w danych. Ponowną iterację przeprowadzimy po wytworzeniu pierwszych modeli.
# * Sprawdziliśmy również, jak wygląda gęstość macierzy interakcji, kluczowy wskaźnik dla problemu rekomendacji — nie budzi on żadnych zastrzeżeń.
|
notebooks/data_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from math import exp, log, factorial
import matplotlib.pyplot as plt
a = 1.0
b = 2.0
N = 10
delta = (b - a) / N
alpha = 1.7
points = [a + i * delta for i in range(N + 1)]
def f(x):
return alpha * exp(-x) + (1 - alpha) * log(x)
def fDerivN1(x):
return (-1) ** (N + 1) * alpha * exp(-x) + (1 - alpha) * ((-1) ** (N)) * factorial(N) / x ** (N + 1)
def maxDerivN1(samples):
space = np.linspace(a, b, samples)
return np.max(np.abs(np.array([(fDerivN1(x)) for x in space], dtype=np.double)))
def omega(k, x):
global points
result = 1
for i in range(N + 1):
if i != k:
result *= (x - points[i])
return result
def denominator(k):
global points
result = 1
for i in range(N + 1):
if i != k:
result *= (points[k] - points[i])
return result
def l(k):
return lambda x: omega(k, x) / denominator(k)
def LagrangePolynomial(x):
result = 0
for i in range(N + 1):
result += l(i)(x) * f(points[i])
return result
def deficiency(x):
return maxDerivN1(10000) * omega(-1, x) / factorial(N + 1)
def plotDifference(samples):
space = np.linspace(a, b, samples)
plt.plot(space, np.zeros(np.shape(space)))
plt.plot(space, np.array([LagrangePolynomial(x) - f(x) for x in space], dtype=np.double))
plt.savefig("../TeX/Interpolation/LagrangeDiff.png")
plt.show()
if __name__ == '__main__':
check = [points[0] + delta / 2.6,
points[5] + delta / 2.6,
points[9] + delta / 2.6]
[print("Pn({0}) = {1}".format(x, LagrangePolynomial(x))) for x in check]
print()
[print("rn({0}) = {1}".format(x, LagrangePolynomial(x) - f(x))) for x in check]
print()
print("M: " + str(maxDerivN1(10000)))
print()
print("Expected deficiency: " +
str(np.max(np.abs(np.array([(deficiency(x)) for x in check], dtype=np.double)))))
space = np.linspace(a, b, 1000)
print("Real deficiency on whole interval: " +
str(np.max(np.abs(np.array([(LagrangePolynomial(x) - f(x)) for x in space], dtype=np.double)))))
print()
print("Real deficiency on control points: " +
str(np.max(np.abs(np.array([(LagrangePolynomial(x) - f(x)) for x in check], dtype=np.double)))))
plotDifference(1000)
|
Interpolation/Lagrange.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
data = pd.read_csv('A:\\Data Analysis Jupyter\\Water-Quality-Analysis\\Dataset\\water_dataX.csv', encoding="ISO-8859-1")
data.head()
data.fillna(0, inplace=True)
data.head()
data.dtypes
data.columns
data['Temp'] = pd.to_numeric(data['Temp'], errors = 'coerce')
data['D.O. (mg/l)'] = pd.to_numeric(data['D.O. (mg/l)'], errors = 'coerce')
data['PH'] = pd.to_numeric(data['PH'], errors='coerce')
data['B.O.D. (mg/l)'] = pd.to_numeric(data['B.O.D. (mg/l)'], errors='coerce')
data['CONDUCTIVITY (µmhos/cm)'] = pd.to_numeric(data['CONDUCTIVITY (µmhos/cm)'], errors='coerce')
data['NITRATENAN N+ NITRITENANN (mg/l)'] = pd.to_numeric(data['NITRATENAN N+ NITRITENANN (mg/l)'], errors='coerce')
data['TOTAL COLIFORM (MPN/100ml)Mean'] = pd.to_numeric(data['TOTAL COLIFORM (MPN/100ml)Mean'], errors='coerce')
data.dtypes
# +
start = 1
end = 1779
station = data.iloc[start:end, 0]
location = data.iloc[start:end ,1]
state = data.iloc[start:end, 2]
do = data.iloc[start:end, 4].astype(np.float64)
value=0
ph = data.iloc[ start:end, 5]
co = data.iloc [start:end, 6].astype(np.float64)
year = data.iloc[start:end, 11]
tc = data.iloc[2:end, 10].astype(np.float64)
bod = data.iloc[start:end, 7].astype(np.float64)
na = data.iloc[start:end, 8].astype(np.float64)
na.dtype
# -
data.head()
# +
data = pd.concat([station,location,state,do,ph,co,bod,na,tc,year], axis=1)
data.columns = ['station','location','state','do','ph','co','bod','na','tc','year']
# -
data.head()
#calulation of Ph
data['npH']=data.ph.apply(lambda x: (100 if (8.5>=x>=7)
else(80 if (8.6>=x>=8.5) or (6.9>=x>=6.8)
else(60 if (8.8>=x>=8.6) or (6.8>=x>=6.7)
else(40 if (9>=x>=8.8) or (6.7>=x>=6.5)
else 0)))))
#calculation of dissolved oxygen
data['ndo']=data.do.apply(lambda x:(100 if (x>=6)
else(80 if (6>=x>=5.1)
else(60 if (5>=x>=4.1)
else(40 if (4>=x>=3)
else 0)))))
#calculation of total coliform
data['nco']=data.tc.apply(lambda x:(100 if (5>=x>=0)
else(80 if (50>=x>=5)
else(60 if (500>=x>=50)
else(40 if (10000>=x>=500)
else 0)))))
#calc of B.D.O
data['nbdo']=data.bod.apply(lambda x:(100 if (3>=x>=0)
else(80 if (6>=x>=3)
else(60 if (80>=x>=6)
else(40 if (125>=x>=80)
else 0)))))
#calculation of electrical conductivity
data['nec']=data.co.apply(lambda x:(100 if (75>=x>=0)
else(80 if (150>=x>=75)
else(60 if (225>=x>=150)
else(40 if (300>=x>=225)
else 0)))))
#Calulation of nitrate
data['nna']=data.na.apply(lambda x:(100 if (20>=x>=0)
else(80 if (50>=x>=20)
else(60 if (100>=x>=50)
else(40 if (200>=x>=100)
else 0)))))
data.head()
data.dtypes
data['wph']=data.npH * 0.165
data['wdo']=data.ndo * 0.281
data['wbdo']=data.nbdo * 0.234
data['wec']=data.nec* 0.009
data['wna']=data.nna * 0.028
data['wco']=data.nco * 0.281
data['wqi']=data.wph+data.wdo+data.wbdo+data.wec+data.wna+data.wco
data.head()
ag = data.groupby('year').mean()
ag.head()
data = ag.reset_index(level = 0, inplace = False)
data.head()
year = data['year'].values
AQI = data['wqi'].values
data['wqi'] = pd.to_numeric(data['wqi'], errors = 'coerce')
data['wqi'] = pd.to_numeric(data['wqi'], errors = 'coerce')
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (20.0, 10.0)
# -
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = Axes3D(fig)
ax.scatter(year, AQI, color = 'red')
plt.show()
data
data = data[np.isfinite(data['wqi'])]
data.head()
# +
cols = ['year']
y = data['wqi']
x = data[cols]
plt.scatter(x, y)
plt.show()
# -
from sklearn import neighbors, datasets
data = data.reset_index(level = 0, inplace = False)
data
from sklearn import linear_model
from sklearn.cross_validation import train_test_split
cols = ['year']
y = data['wqi']
x = data[cols]
reg = linear_model.LinearRegression()
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2, random_state = 4)
reg.fit(x_train, y_train)
a = reg.predict(x_test)
a
y_test
from sklearn.metrics import mean_squared_error
print('mse:%.2f'%mean_squared_error(y_test,a))
dt = pd.DataFrame({'Actual': y_test, 'Predicted': a})
#using gradient descent to optimize it further
x = (x - x.mean()) / x.std()
x = np.c_[np.ones(x.shape[0]), x]
x
import matplotlib.pyplot as plt
data=data.set_index('year')
data.plot(figsize=(15,6))
plt.show()
# +
alpha = 0.1 #Step size
iterations = 3000 #No. of iterations
m = y.size #No. of data points
np.random.seed(4) #Setting the seed
theta = np.random.rand(2) #Picking some random values to start with
def gradient_descent(x, y, theta, iterations, alpha):
past_costs = []
past_thetas = [theta]
for i in range(iterations):
prediction = np.dot(x, theta)
error = prediction - y
cost = 1/(2*m) * np.dot(error.T, error)
past_costs.append(cost)
theta = theta - (alpha * (1/m) * np.dot(x.T, error))
past_thetas.append(theta)
return past_thetas, past_costs
past_thetas, past_costs = gradient_descent(x, y, theta, iterations, alpha)
theta = past_thetas[-1]
#Print the results...
print("Gradient Descent: {:.2f}, {:.2f}".format(theta[0], theta[1]))
# -
plt.title('Cost Function J')
plt.xlabel('No. of iterations')
plt.ylabel('Cost')
plt.plot(past_costs)
plt.show()
import numpy as np
newB=[74.76, 2.13]
def rmse(y,y_pred):
rmse= np.sqrt(sum(y-y_pred))
return rmse
# +
y_pred=x.dot(newB)
dt = pd.DataFrame({'Actual': y, 'Predicted': y_pred})
dt=pd.concat([data, dt], axis=1)
dt
# +
#testing the accuracy of the model
from sklearn import metrics
print(np.sqrt(metrics.mean_squared_error(y,y_pred)))
# +
#plotting the actual and predicted results
x_axis=dt.year
y_axis=dt.Actual
y1_axis=dt.Predicted
plt.scatter(x_axis,y_axis)
plt.plot(x_axis,y1_axis,color='r')
plt.title("linear regression")
plt.show()
|
Code/Water-Quaity-Kernel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# $\int \frac{1}{x^4 - x^3}dx$
# $\int \frac{1}{x^4 - x^3} = \frac{1}{x^3(x - 1)}$
# $\frac{1}{x^3(x - 1)} = \frac{A}{x} + \frac{B}{x^2} + \frac{C}{x^3} + \frac{D}{(x-1)}$
# $1 = A(x^2)(x-1) + Bx(x-1) + C(x-1) + Dx^3$
# $x = 1$
# $1 = A(1^2)(1-1) + B\cdot1\cdot(1-1) + C(1-1) + D\cdot 1^3$
# $D = 1$
# $x = 0$
# $1 = A(0^2)(0-1) + B\cdot0\cdot(0-1) + C(0-1) + 1\cdot 0^3$
# $C = -1$
# $x = 2$
# $1 = A(2^2)(2-1) + B\cdot 2 \cdot (2-1) + (-1)\cdot(2-1) + 2^3$
# $1 = 4A + 2B + 7$
# $-3 = 2A + B$
# $B = -3 - 2A$
# $1 = 4A + 2(-3 - 2A) + 7$
# $1 = 4A -6 - 4A + 7$
# $1 = $
|
Problemas 7.4/05.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import keras
from math import ceil
from datetime import datetime
import json
DATA_PATH = '/data/'
MODEL_PATH = 'models/'
BATCH_SIZE = 128
# ## Load data
from keras.applications.xception import Xception
from keras.preprocessing import image
from keras.models import Model
from keras.layers import Dense, GlobalAveragePooling2D
from keras.optimizers import Adam
train_generator = image.ImageDataGenerator(
rotation_range=10,
width_shift_range=0.1,
height_shift_range=0.1,
zoom_range=0.1,
horizontal_flip=True,
vertical_flip=True,
)
train_batches = train_generator.flow_from_directory(
DATA_PATH + 'train/',
target_size=(299, 299),
shuffle=True,
batch_size=BATCH_SIZE,
class_mode='categorical',
)
valid_batches = image.ImageDataGenerator().flow_from_directory(
DATA_PATH + 'valid/',
target_size=(299, 299),
shuffle=True,
batch_size=BATCH_SIZE,
class_mode='categorical',
)
num_classes = train_batches.num_classes
base_model = Xception(weights='imagenet', include_top=False)
x = base_model.output
x = GlobalAveragePooling2D()(x)
predictions = Dense(num_classes, activation='softmax')(x)
model = Model(inputs=base_model.input, outputs=predictions)
for layer in base_model.layers:
layer.trainable = False
model.compile(optimizer=Adam(), loss='categorical_crossentropy', metrics=['accuracy'])
model.summary()
model.fit_generator(
train_batches,
steps_per_epoch=int(ceil(train_batches.samples/BATCH_SIZE)),
validation_data=valid_batches,
validation_steps=int(ceil(valid_batches.samples/BATCH_SIZE)),
epochs=1,
)
model.optimizer.lr = 0.01
model.fit_generator(
train_batches,
steps_per_epoch=int(ceil(train_batches.samples/BATCH_SIZE)),
validation_data=valid_batches,
validation_steps=int(ceil(valid_batches.samples/BATCH_SIZE)),
epochs=10,
)
def save_model(model, train_batches):
model_name = MODEL_PATH + f'carnet-{datetime.now().isoformat()}'
model.save_weights(model_name + '-weights.h5')
with open(model_name + '-model.json', 'w') as f:
f.write(model.to_json())
with open(model_name + '-classes.json', 'w') as f:
json.dump(train_batches.class_indices, f)
from keras.models import model_from_json
def load_model():
with open(MODEL_PATH + 'carnet-2017-11-12T00:04:15.433428-model.json') as f:
model = model_from_json(f.read())
model.load_weights(MODEL_PATH + 'carnet-2017-11-12T00:04:15.433428-weights.h5')
return model
model = load_model()
model.optimizer.lr = 0.001
model.fit_generator(
train_batches,
steps_per_epoch=int(ceil(train_batches.samples/BATCH_SIZE)),
validation_data=valid_batches,
validation_steps=int(ceil(valid_batches.samples/BATCH_SIZE)),
epochs=1,
)
save_model(model, train_batches)
for i in range(20):
print(f'{i+1}/10')
model.fit_generator(
train_batches,
steps_per_epoch=int(ceil(train_batches.samples/BATCH_SIZE)),
validation_data=valid_batches,
validation_steps=int(ceil(valid_batches.samples/BATCH_SIZE)),
epochs=10,
)
save_model(model, train_batches)
|
Car brand detector.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
#
# _You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-data-analysis/resources/0dhYG) course resource._
#
# ---
# # Merging Dataframes
#
# +
import pandas as pd
df = pd.DataFrame([{'Name': 'Chris', 'Item Purchased': 'Sponge', 'Cost': 22.50},
{'Name': 'Kevyn', 'Item Purchased': 'Kitty Litter', 'Cost': 2.50},
{'Name': 'Filip', 'Item Purchased': 'Spoon', 'Cost': 5.00}],
index=['Store 1', 'Store 1', 'Store 2'])
df
# -
df['Date'] = ['December 1', 'January 1', 'mid-May']
df
df['Delivered'] = True
df
df['Feedback'] = ['Positive', None, 'Negative']
df
adf = df.reset_index()
adf['Date'] = pd.Series({0: 'December 1', 2: 'mid-May'})
adf
staff_df = pd.DataFrame([{'Name': 'Kelly', 'Role': 'Director of HR'},
{'Name': 'Sally', 'Role': 'Course liasion'},
{'Name': 'James', 'Role': 'Grader'}])
staff_df = staff_df.set_index('Name')
student_df = pd.DataFrame([{'Name': 'James', 'School': 'Business'},
{'Name': 'Mike', 'School': 'Law'},
{'Name': 'Sally', 'School': 'Engineering'}])
student_df = student_df.set_index('Name')
print(staff_df.head())
print()
print(student_df.head())
pd.merge(staff_df, student_df, how='outer', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='inner', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='left', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='right', left_index=True, right_index=True)
staff_df = staff_df.reset_index()
student_df = student_df.reset_index()
pd.merge(staff_df, student_df, how='left', left_on='Name', right_on='Name')
staff_df = pd.DataFrame([{'Name': 'Kelly', 'Role': 'Director of HR', 'Location': 'State Street'},
{'Name': 'Sally', 'Role': 'Course liasion', 'Location': 'Washington Avenue'},
{'Name': 'James', 'Role': 'Grader', 'Location': 'Washington Avenue'}])
student_df = pd.DataFrame([{'Name': 'James', 'School': 'Business', 'Location': '1024 Billiard Avenue'},
{'Name': 'Mike', 'School': 'Law', 'Location': 'Fraternity House #22'},
{'Name': 'Sally', 'School': 'Engineering', 'Location': '512 Wilson Crescent'}])
pd.merge(staff_df, student_df, how='left', left_on='Name', right_on='Name')
staff_df = pd.DataFrame([{'First Name': 'Kelly', 'Last Name': 'Desjardins', 'Role': 'Director of HR'},
{'First Name': 'Sally', 'Last Name': 'Brooks', 'Role': 'Course liasion'},
{'First Name': 'James', 'Last Name': 'Wilde', 'Role': 'Grader'}])
student_df = pd.DataFrame([{'First Name': 'James', 'Last Name': 'Hammond', 'School': 'Business'},
{'First Name': 'Mike', 'Last Name': 'Smith', 'School': 'Law'},
{'First Name': 'Sally', 'Last Name': 'Brooks', 'School': 'Engineering'}])
staff_df
student_df
pd.merge(staff_df, student_df, how='inner', left_on=['First Name','Last Name'], right_on=['First Name','Last Name'])
# # Idiomatic Pandas: Making Code Pandorable
import pandas as pd
df = pd.read_csv('census.csv')
df
(df.where(df['SUMLEV']==50)
.dropna()
.set_index(['STNAME','CTYNAME'])
.rename(columns={'ESTIMATESBASE2010': 'Estimates Base 2010'}))
df = df[df['SUMLEV']==50]
df.set_index(['STNAME','CTYNAME'], inplace=True)
df.rename(columns={'ESTIMATESBASE2010': 'Estimates Base 2010'})
import numpy as np
def min_max(row):
data = row[['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']]
return pd.Series({'min': np.min(data), 'max': np.max(data)})
df.apply(min_max, axis=1)
import numpy as np
def min_max(row):
data = row[['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']]
row['max'] = np.max(data)
row['min'] = np.min(data)
return row
df.apply(min_max, axis=1)
rows = ['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']
df.apply(lambda x: np.max(x[rows]), axis=1)
# # Group by
import pandas as pd
import numpy as np
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df
# %%timeit -n 10
for state in df['STNAME'].unique():
avg = np.average(df.where(df['STNAME']==state).dropna()['CENSUS2010POP'])
print('Counties in state ' + state + ' have an average population of ' + str(avg))
# %%timeit -n 10
for group, frame in df.groupby('STNAME'):
avg = np.average(frame['CENSUS2010POP'])
print('Counties in state ' + group + ' have an average population of ' + str(avg))
df.head()
# +
df = df.set_index('STNAME')
def fun(item):
if item[0]<'M':
return 0
if item[0]<'Q':
return 1
return 2
for group, frame in df.groupby(fun):
print('There are ' + str(len(frame)) + ' records in group ' + str(group) + ' for processing.')
# -
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df.groupby('STNAME').agg({'CENSUS2010POP': np.average})
print(type(df.groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']))
print(type(df.groupby(level=0)['POPESTIMATE2010']))
(df.set_index('STNAME').groupby(level=0)['CENSUS2010POP']
.agg({'avg': np.average, 'sum': np.sum}))
(df.set_index('STNAME').groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']
.agg({'avg': np.average, 'sum': np.sum}))
(df.set_index('STNAME').groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']
.agg({'POPESTIMATE2010': np.average, 'POPESTIMATE2011': np.sum}))
# # Scales
df = pd.DataFrame(['A+', 'A', 'A-', 'B+', 'B', 'B-', 'C+', 'C', 'C-', 'D+', 'D'],
index=['excellent', 'excellent', 'excellent', 'good', 'good', 'good', 'ok', 'ok', 'ok', 'poor', 'poor'])
df.rename(columns={0: 'Grades'}, inplace=True)
df
df['Grades'].astype('category').head()
grades = df['Grades'].astype('category',
categories=['D', 'D+', 'C-', 'C', 'C+', 'B-', 'B', 'B+', 'A-', 'A', 'A+'],
ordered=True)
grades.head()
grades > 'C'
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df = df.set_index('STNAME').groupby(level=0)['CENSUS2010POP'].agg({'avg': np.average})
pd.cut(df['avg'],10)
# # Pivot Tables
#http://open.canada.ca/data/en/dataset/98f1a129-f628-4ce4-b24d-6f16bf24dd64
df = pd.read_csv('cars.csv')
df.head()
df.pivot_table(values='(kW)', index='YEAR', columns='Make', aggfunc=np.mean)
df.pivot_table(values='(kW)', index='YEAR', columns='Make', aggfunc=[np.mean,np.min], margins=True)
# # Date Functionality in Pandas
import pandas as pd
import numpy as np
# ### Timestamp
pd.Timestamp('9/1/2016 10:05AM')
# ### Period
pd.Period('1/2016')
pd.Period('3/5/2016')
# ### DatetimeIndex
t1 = pd.Series(list('abc'), [pd.Timestamp('2016-09-01'), pd.Timestamp('2016-09-02'), pd.Timestamp('2016-09-03')])
t1
type(t1.index)
# ### PeriodIndex
t2 = pd.Series(list('def'), [pd.Period('2016-09'), pd.Period('2016-10'), pd.Period('2016-11')])
t2
type(t2.index)
# ### Converting to Datetime
d1 = ['2 June 2013', 'Aug 29, 2014', '2015-06-26', '7/12/16']
ts3 = pd.DataFrame(np.random.randint(10, 100, (4,2)), index=d1, columns=list('ab'))
ts3
ts3.index = pd.to_datetime(ts3.index)
ts3
pd.to_datetime('4.7.12', dayfirst=True)
# ### Timedeltas
pd.Timestamp('9/3/2016')-pd.Timestamp('9/1/2016')
pd.Timestamp('9/2/2016 8:10AM') + pd.Timedelta('12D 3H')
# ### Working with Dates in a Dataframe
dates = pd.date_range('10-01-2016', periods=9, freq='2W-SUN')
dates
df = pd.DataFrame({'Count 1': 100 + np.random.randint(-5, 10, 9).cumsum(),
'Count 2': 120 + np.random.randint(-5, 10, 9)}, index=dates)
df
df.index.weekday_name
df.diff()
df.resample('M').mean()
df['2017']
df['2016-12']
df['2016-12':]
df.asfreq('W', method='ffill')
# +
import matplotlib.pyplot as plt
# %matplotlib inline
df.plot()
# -
|
Week+3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:automl]
# language: python
# name: conda-env-automl-py
# ---
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
from datetime import date, datetime, timedelta
import pickle
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
import cufflinks as cf
import seaborn as sns
from scipy import signal, stats
from scipy.stats import pearsonr
turbine_readings = pd.read_csv("./data/WindData_scrubbed.csv")
weather_data = pd.read_csv("hourly_weather_data.csv")
turbine_readings = turbine_readings.iloc[:, 0:74]
mean_readings = turbine_readings.apply(lambda x: np.mean(x), axis=1)
median_readings = turbine_readings.apply(lambda x: np.median(x), axis=1)
print(turbine_readings.shape)
weather_data = weather_data.dropna(axis=1, how="all")
weather_data.fillna(method="ffill", inplace=True)
weather_data["timestamp"] = pd.to_datetime(weather_data["valid_time_gmt"], unit="s") - pd.Timedelta("08:00:00")
weather_data["weather_date"] = weather_data["timestamp"].dt.date
weather_data["weather_time"] = weather_data["timestamp"].dt.time
weather_data["weather_month"] = weather_data["timestamp"].dt.month
weather_data["weather_day_of_month"] = weather_data["timestamp"].dt.day
weather_data["weather_day_of_week"] = weather_data["timestamp"].dt.dayofweek
weather_data.set_index("timestamp", inplace=True)
windspeed = weather_data["wspd"]
windspeed_cubed = windspeed**3
def plot_cross_correlation(feature, readings, interval):
feature = feature.resample(interval).asfreq()
feature.interpolate(method="linear", inplace=True)
# signal.correlate calculates the integral(area) of the product of shifting time series
cross_corrs_valid = signal.correlate(feature, readings, mode="valid", method="direct")
print("Valid Shape: ", cross_corrs_valid.shape)
cross_corrs_full = signal.correlate(feature, readings, mode="full", method="direct")
print("Full Shape: ", cross_corrs_full.shape)
#print(len(feature.index), cross_corrs_valid.shape)
trace = go.Scatter(
x = feature.index,
y = cross_corrs_valid
)
data = [trace]
py.iplot(data)
plt.figure(figsize=(15,8))
plt.plot(cross_corrs_full)
plt.axvline(x=readings.shape[0]-1, color="red")
plt.axvline(x=cross_corrs_full.shape[0]-readings.shape[0], color="red")
def plot_pearson_correlation(feature, readings, interval):
feature = feature.resample(interval).asfreq()
feature.interpolate(method="linear", inplace=True)
# signal.correlate calculates the integral(area) of the product of shifting time series
pears_corrs = pearsonr(feature, readings)
print("pearsonr Shape: ", pears_corrs.shape)
cross_corrs_full = signal.correlate(feature, readings, mode="full", method="direct")
print("Full Shape: ", cross_corrs_full.shape)
#print(len(feature.index), cross_corrs_valid.shape)
trace = go.Scatter(
x = feature.index,
y = pears_corrs[0]
)
data = [trace]
py.iplot(data)
plt.figure(figsize=(15,8))
plt.plot(cross_corrs_full)
plt.axvline(x=readings.shape[0]-1, color="red")
plt.axvline(x=cross_corrs_full.shape[0]-readings.shape[0], color="red")
plot_cross_correlation(windspeed_cubed, mean_readings, "1Min")
plot_cross_correlation(windspeed_cubed, mean_readings, "5Min")
plot_cross_correlation(windspeed_cubed, mean_readings, "20Min")
plot_cross_correlation(windspeed_cubed, mean_readings, "1H")
plot_cross_correlation(windspeed_cubed, mean_readings, "100Min")
print(stats.pearsonr(turbine_readings.iloc[:, 0], turbine_readings.iloc[:, 1]))
print(np.corrcoef(turbine_readings.iloc[:, 0], turbine_readings.iloc[:, 1]))
windspeed_5min = windspeed_cubed.resample("5Min").asfreq()
windspeed_5min.interpolate(method="linear", inplace=True)
correlations = []
for i in range(0, (len(windspeed_5min)-len(mean_readings)+1)):
correlations.append(stats.pearsonr(mean_readings, windspeed_5min.iloc[i:i+mean_readings.shape[0]])[0])
print(len(correlations))
trace = go.Scatter(
x = windspeed_5min.index,
y = correlations
)
data = [trace]
py.iplot(data)
# # Modelling the GE output (Regression)
#
feature = windspeed_cubed
feature = feature.resample('5Min').asfreq()
feature.interpolate(method="linear", inplace=True)
# +
corr_array = signal.correlate(feature, mean_readings, mode='valid', method = 'direct')
# -
peak_timestamp = feature.index[np.argmax(corr_array)]
peak_timestamp
ranga = pd.date_range(start= peak_timestamp, periods = 2600, freq='5T')
ranga
time_turbine = pd.DataFrame(list(mean_readings), index=ranga)
ge_data = time_turbine
# +
ge_data['wspd'] = feature[time_turbine.index]
# +
feature_temp = weather_data.temp
feature_temp = feature_temp.resample('5Min').asfreq()
feature_temp.interpolate(method="linear", inplace=True)
ge_data['temp'] = feature_temp[time_turbine.index]
# +
feature_wdir = weather_data.wdir
feature_wdir = feature_wdir.resample('5Min').asfreq()
feature_wdir.interpolate(method="linear", inplace=True)
ge_data['wdir'] = feature_wdir[time_turbine.index]
# -
ge_data.rename(columns = {0:'output'}, inplace=True)
ge_data['dt_col'] = pd.to_datetime(ge_data.index)
ge_data = ge_data.set_index('dt_col')
# +
ge_output = ge_data[['output']]
# -
ge_output = pd.Series(ge_data.output)
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot, plot_mpl
init_notebook_mode(connected=True)
#from plotly.offline import plot_mpl
#from plotly.plotly import plot_mpl
from statsmodels.tsa.seasonal import seasonal_decompose
result = seasonal_decompose(ge_output, freq = 288)
fig = result.plot()
plot_mpl(fig)
#fig = go.Figure(d
# -
from statsmodels.tsa.vector_ar.vecm import coint_johansen
#since the test works for only 12 variables, I have randomly dropped
#in the next iteration, I would drop another and check the eigenvalues
#johan_test_temp = data.drop([ 'CO(GT)'], axis=1)
coint_johansen(ge_data,-1,1).eig
coint_johansen(ge_data[["output","wspd"]],-1,1).eig
# +
from sklearn.preprocessing import MinMaxScaler, StandardScaler
scaler = StandardScaler()
minmax = MinMaxScaler(feature_range=(0,1))
ge_data.wspd = scaler.fit_transform(np.array(ge_data.wspd.values).reshape(-1,1))
ge_data.wdir = scaler.fit_transform(np.array(ge_data.wdir.values).reshape(-1,1))
ge_data.temp = scaler.fit_transform(np.array(ge_data.temp.values).reshape(-1,1))
ge_data.output = minmax.fit_transform(np.array(ge_data.output.values).reshape(-1,1))
# +
#creating the train and validation set
train = ge_data[:int(0.8*(len(ge_data)))]
valid = ge_data[int(0.8*(len(ge_data))):]
#fit the model
from statsmodels.tsa.vector_ar.var_model import VAR
model1 = VAR(endog=train)
model_fit = model1.fit()
# make prediction on validation
prediction = model_fit.forecast(model_fit.y, steps=len(valid))
# +
# Baseline Model Random Forest
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import RandomizedSearchCV
# +
param_grid = {'bootstrap': [True, False],
'max_depth': [10, 20, 30, 40, 50, 60, 70, 80, 90, 100, None],
'max_features': ['auto', 'sqrt'],
'min_samples_leaf': [1, 2, 4],
'min_samples_split': [2, 5, 10],
'n_estimators': [200, 400, 600, 800, 1000, 1200, 1400, 1600, 1800, 2000]}
rf = RandomForestRegressor()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
rf_random = RandomizedSearchCV(estimator = rf, param_distributions = param_grid, n_iter = 100, cv = 4, verbose=2, random_state=42, n_jobs = -1)
# Fit the random search model
rf_random.fit(train[['wspd','temp','wdir']], train.output)
# -
rf_random.score(valid[['wspd','temp','wdir']], valid.output)
print("The RMSE for baseline RF model is: " ,np.sqrt(mean_squared_error( y_pred=rf_random.predict(valid[['wspd','temp','wdir']]), y_true=valid.output)))
# +
train_X = train[["wspd","temp","wdir"]]
train_y = train.output
val_X = valid[["wspd","temp","wdir"]]
val_y = valid.output
train_X = train_X.values
val_X = val_X.values
# +
train_X = train_X.reshape((train_X.shape[0], 1, train_X.shape[1]))
#test_X = test_X.reshape((test_X.shape[0], 1, test_X.shape[1]))
val_X = val_X.reshape((val_X.shape[0], 1, val_X.shape[1]))
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM, GRU
# design network
model = Sequential()
#model.add(LSTM())
model.add(GRU(24, input_shape=(train_X.shape[1], train_X.shape[2])))
#model.add(GRU(100))
model.add(Dense(1))
model.compile(loss='mse', optimizer='adam')
# fit network
history = model.fit(train_X, train_y, epochs=75, batch_size=300, validation_data=(val_X, val_y), verbose=2, shuffle=False)
# plot history
#import matplotlib.pyplot as pyplot
#pyplot.plot(history.history['loss'], label='train')
#pyplot.plot(history.history['val_loss'], label='test')
#pyplot.legend()
#pyplot.show()
#y_pred = model.predict(test_X)
# -
y_pred = model.predict(val_X)
# +
cols = ge_data.columns
from sklearn.metrics import mean_squared_error
#converting predictions to dataframe
pred = pd.DataFrame(index=range(0,len(prediction)),columns=[cols])
for j in range(0,3):
for i in range(0, len(prediction)):
pred.iloc[i][j] = prediction[i][j]
#check rmse
for i in cols:
print('rmse value for', i, 'is : ', np.sqrt(mean_squared_error(pred[i], valid[i])))
# -
pred.output = pred.output.astype('float64')
pred.output = np.array(pred.output).reshape(-1,)
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
#traces = []
#random_turbines = np.random.choice(turbine_names, size=3, replace=False)
trace1 = go.Scatter(x = valid.index,
y = list(100*val_y),
mode = 'lines',
name = 'test')
trace2 = go.Scatter(x = valid.index,
y = np.array(100*y_pred).reshape(-1,),
mode = 'lines',
name = 'pred')
traces=[trace1, trace2]
layout = go.Layout(
title='GE data LSTM model'
)
fig = go.Figure(data=traces, layout=layout)
#plot_url = py.plot(fig, filename='multiple-axes-double')
py.iplot(fig, filename='line-mode')
# -
np.sqrt(mean_squared_error(y_pred, val_y))
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
#traces = []
#random_turbines = np.random.choice(turbine_names, size=3, replace=False)
trace1 = go.Scatter(x = valid.index,
y = list(valid.output),
mode = 'lines',
name = 'test')
trace2 = go.Scatter(x = valid.index,
y = np.array(pred.output).reshape(-1,),
mode = 'lines',
name = 'pred')
traces=[trace1, trace2]
layout = go.Layout(
title='VAR model'
)
fig = go.Figure(data=traces, layout=layout)
#plot_url = py.plot(fig, filename='multiple-axes-double')
py.iplot(fig, filename='line-mode')
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
#traces = []
#random_turbines = np.random.choice(turbine_names, size=3, replace=False)
trace1 = go.Scatter(x = valid.index,
y = list(100*valid.output),
mode = 'lines',
name = 'Actual Output')
trace2 = go.Scatter(x = valid.index,
y = 100*rf_random.predict(valid[['wspd','temp','wdir']]),
mode = 'lines',
name = 'Predicted Output')
traces=[trace1, trace2]
layout = go.Layout(
title='Random Forest model'
)
fig = go.Figure(data=traces, layout=layout)
#plot_url = py.plot(fig, filename='multiple-axes-double')
py.iplot(fig, filename='line-mode')
# -
# # Sotavento Model
import pandas as pd
#soluto_data = pd.read_csv('soluto_windfarm.csv', encoding='latin1')
#daily_soluto = pd.read_excel('soluto_daily.xlsx', sheet_name='daily')
hourly_soluto = pd.read_excel('soluto_daily.xlsx', sheet_name='hourly')
#soluto_data.Date = pd.to_datetime(soluto_data.Date)
hourly_soluto.Date = pd.to_datetime(hourly_soluto.Date)
hourly_soluto = hourly_soluto.sort_values("Date")
hourly_soluto=hourly_soluto.set_index('Date')
hourly_soluto.head()
hourly_soluto = hourly_soluto[~hourly_soluto.index.duplicated(keep='first')]
# +
# Converting from european format to regular float
hourly_soluto['Energy'] = hourly_soluto['Energy'].apply(lambda x: x.replace('.','').replace(',', '.'))
hourly_soluto['Speed'] = hourly_soluto['Speed'].apply(lambda x: x.replace('.','').replace(',', '.'))
#hourly_soluto['Direction'] = hourly_soluto['Direction'].apply(lambda x: x.replace('.','').replace(",","."))
# -
hourly_soluto1 = hourly_soluto.replace({'-': 0.000001})
hourly_soluto = hourly_soluto.replace({'-': np.nan})
hourly_soluto_p = hourly_soluto.dropna()
hourly_soluto_p.Energy = hourly_soluto_p.Energy.astype('float64')
hourly_soluto_p.Speed = hourly_soluto_p.Speed.astype('float64')
subset = hourly_soluto['2013-01-01 00:00:00' : '2015-12-31 23:00:00']
subset.Energy = subset.Energy.astype('float64')
subset.Speed = subset.Speed.astype('float64')
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot, plot_mpl
init_notebook_mode(connected=True)
#from plotly.offline import plot_mpl
#from plotly.plotly import plot_mpl
from statsmodels.tsa.seasonal import seasonal_decompose
result = seasonal_decompose(subset.Speed.values, model='additive', freq = 24*365)
fig = result.plot()
plot_mpl(fig)
#fig = go.Figure(data=traces, layout=layout)
#plot_url = py.plot(fig, filename='multiple-axes-double')
#py.iplot(fig, filename='line-mode')
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
data = [go.Scatter(x=hourly_soluto.index, y=hourly_soluto.Energy )]
py.iplot(data)
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
traces = []
#random_turbines = np.random.choice(turbine_names, size=3, replace=False)
for feature in soluto_data.columns[1:]:
#filtered_turbine = daily_mean_turbine_reading.loc[daily_mean_turbine_reading["turbine_num"] == turbine]
trace = go.Scatter(x = soluto_data.Date,
y = float(soluto_data[feature])/max(float(soluto_data[feature])),
mode = 'lines',
name = feature)
traces.append(trace)
layout = go.Layout(
title='Time Series - Wind Speed (NOAA data), Turbine Output',
yaxis=dict(
title='Turbine Output (kWh)',
#title='Wind Speed (mph)',
titlefont=dict(
color='blue'
),
tickfont=dict(
color='blue'
),
),
yaxis2=dict(
title='Wind Speed (mph)',
titlefont=dict(
color='orange'
),
tickfont=dict(
color='orange'
),
overlaying='y',
side='right'
)
)
fig = go.Figure(data=traces, layout=layout)
#plot_url = py.plot(fig, filename='multiple-axes-double')
py.iplot(fig, filename='line-mode')
# -
import statsmodels.api as sm
print(sm.tsa.stattools.grangercausalitytests(subset[['Energy','Speed']],maxlag=1000))
print(sm.tsa.stattools.grangercausalitytests(subset[['Speed','Energy']],1))
# ## LSTM
from sklearn.preprocessing import StandardScaler, MinMaxScaler
# +
def series_to_supervised(data, n_in=1, n_out=1, dropnan=True, feat_names = None):
n_vars = 1 if type(data) is list else data.shape[1]
df = pd.DataFrame(data)
cols, names = list(), list()
# input sequence (t-n, ... t-1)
for i in range(n_in, 0, -1):
cols.append(df.shift(i))
names += [(feat_names[j] + '(t-%d)' % i) for j in range(n_vars)]
# forecast sequence (t, t+1, ... t+n)
for i in range(0, n_out):
cols.append(df.shift(-i))
if i == 0:
names += [(feat_names[j] + '(t)' ) for j in range(n_vars)]
else:
names += [('var%d(t+%d)' % (j+1, i)) for j in range(n_vars)]
# put it all together
agg = pd.concat(cols, axis=1)
agg.columns = names
# drop rows with NaN values
if dropnan:
agg.dropna(inplace=True)
return agg
# -
values = dataset.values
# integer encode direction
encoder = LabelEncoder()
values[:,4] = encoder.fit_transform(values[:,4])
# ensure all data is float
values = values.astype('float32')
# normalize features
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(values)
# frame as supervised learning
reframed = series_to_supervised(scaled, 1, 1)
# drop columns we don't want to predict
reframed.drop(reframed.columns[[9,10,11,12,13,14,15]], axis=1, inplace=True)
print(reframed.head())
val = hourly_soluto_p.values
scaler = StandardScaler()
scaled = scaler.fit_transform(val)
reframed = series_to_supervised(scaled, n_in= 24,n_out= 1, feat_names= hourly_soluto_p.columns)
# split into train and test sets
values = reframed.values
n_train_hours = 365 * 24
train = values[:n_train_hours, :]
test = values[n_train_hours:, :]
# split into input and outputs
train_X, train_y = train[:, :-1], train[:, -1]
test_X, test_y = test[:, :-1], test[:, -1]
# reshape input to be 3D [samples, timesteps, features]
train_X = train_X.reshape((train_X.shape[0], 1, train_X.shape[1]))
test_X = test_X.reshape((test_X.shape[0], 1, test_X.shape[1]))
print(train_X.shape, train_y.shape, test_X.shape, test_y.shape)
# +
X = hourly_soluto_p[['Speed', 'Direction']]
y = hourly_soluto_p[['Energy']]
X_train = X['2006-01-01 01:00:00' : '2016-12-30 23:00:00']
X_val = X['2017-01-01 01:00:00' : '2017-12-31 23:00:00']
X_test = X['2018-01-01 01:00:00' : '2018-11-30 23:00:00']
y_train = y['2006-01-01 01:00:00' : '2016-12-30 23:00:00']
y_val = y['2017-01-01 01:00:00' : '2017-12-31 23:00:00']
y_test = y['2018-01-01 01:00:00' : '2018-11-30 23:00:00']
# +
train_X = X_train.values
train_y = y_train.values
test_X = X_val.values
test_y = y_val.values
minmax = MinMaxScaler(feature_range=(0,1))
train_X = scaler.fit_transform(train_X)
train_y = minmax.fit_transform(train_y)
test_X = scaler.fit_transform(test_X)
test_y = minmax.fit_transform(test_y)
# +
train_X = train_X.reshape((train_X.shape[0], 1, train_X.shape[1]))
test_X = test_X.reshape((test_X.shape[0], 1, test_X.shape[1]))
# -
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM, GRU
# +
# design network
model = Sequential()
#model.add(LSTM())
model.add(GRU(75, input_shape=(train_X.shape[1], train_X.shape[2])))
model.add(Dense(1))
model.compile(loss='mae', optimizer='adam')
# fit network
history = model.fit(train_X, train_y, epochs=100, batch_size=300, validation_data=(test_X, test_y), verbose=2, shuffle=False)
# plot history
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.legend()
pyplot.show()
# +
val_X = X_test.values
val_y = y_test.values
#minmax = MinMaxScaler(feature_range=(0,1))
val_X = scaler.fit_transform(val_X)
val_y = minmax.fit_transform(val_y)
# +
val_X = val_X.reshape((val_X.shape[0], 1, val_X.shape[1]))
# +
# make a prediction
y_pred = model.predict(val_X)
# -
pyplot.plot(pd.DataFrame(100*y_pred[1000:1200], index= X_val.index[1000 : 1200]), label='Predction')
pyplot.plot(pd.DataFrame(100*val_y[1000:1200], index= X_val.index[1000 : 1200]), label='Actual')
pyplot.legend()
pyplot.xlabel('time stamps')
pyplot.ylabel('% utilization')
pyplot.show()
pyplot.plot(100*y_pred[4000:4200], label='Predction')
pyplot.plot(100*val_y[4000:4200], label='Actual')
pyplot.legend()
pyplot.show()
from sklearn.metrics import mean_absolute_error
100*mean_absolute_error(y_pred, val_y)
# +
import plotly.offline as py
import plotly.graph_objs as go
from plotly import __version__
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
traces = []
#for turbine in random_turbines:
trace = go.Scatter(x = X_val.index,
y = y_pred,
mode = 'lines',
name = 'Prediction')
traces.append(trace)
py.iplot(traces, filename='line-mode')
# -
|
13 - Model Fitting Copy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
# # Using activation function
# You can use activation function which prepared by tensorflow.
# It is in tf.nn library!
sess = tf.Session()
# Relu
print(sess.run(tf.nn.relu([-3., 3., 10.])))
# Relu6
# Set upper limit as 6
print(sess.run(tf.nn.relu6([-3., 3., 10.])))
# sigmoid
print(sess.run(tf.nn.sigmoid([-1., 0., 1.])))
# Hyperbolic tangent
print(sess.run(tf.nn.tanh([-1., 0., 1.])))
# Softsign function
print(sess.run(tf.nn.softsign([-1., 0., 1.])))
# Softplus function
print(sess.run(tf.nn.softplus([-1., 0., 1.])))
# ELU(Exponential Linear Unit)
print(sess.run(tf.nn.elu([-1., 0., 1.])))
|
chapter1/Activation functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # 循环神经网络的从零开始实现
#
# 在本节中,我们将从零开始实现一个基于字符级循环神经网络的语言模型,并在周杰伦专辑歌词数据集上训练一个模型来进行歌词创作。首先,我们读取周杰伦专辑歌词数据集。
# + attributes={"classes": [], "id": "", "n": "1"}
import d2lzh as d2l
import math
from mxnet import autograd, nd
from mxnet.gluon import loss as gloss
import time
(corpus_indices, char_to_idx, idx_to_char,
vocab_size) = d2l.load_data_jay_lyrics()
# -
# ## one-hot向量
#
# 为了将词表示成向量输入到神经网络,一个简单的办法是使用one-hot向量。假设词典中不同字符的数量为$N$(即词典大小`vocab_size`),每个字符已经同一个从0到$N-1$的连续整数值索引一一对应。如果一个字符的索引是整数$i$, 那么我们创建一个全0的长为$N$的向量,并将其位置为$i$的元素设成1。该向量就是对原字符的one-hot向量。下面分别展示了索引为0和2的one-hot向量,向量长度等于词典大小。
# + attributes={"classes": [], "id": "", "n": "2"}
nd.one_hot(nd.array([0, 2]), vocab_size)
# -
# 我们每次采样的小批量的形状是(批量大小, 时间步数)。下面的函数将这样的小批量变换成数个可以输入进网络的形状为(批量大小, 词典大小)的矩阵,矩阵个数等于时间步数。也就是说,时间步$t$的输入为$\boldsymbol{X}_t \in \mathbb{R}^{n \times d}$,其中$n$为批量大小,$d$为输入个数,即one-hot向量长度(词典大小)。
# + attributes={"classes": [], "id": "", "n": "3"}
def to_onehot(X, size): # 本函数已保存在d2lzh包中方便以后使用
return [nd.one_hot(x, size) for x in X.T]
X = nd.arange(10).reshape((2, 5))
inputs = to_onehot(X, vocab_size)
len(inputs), inputs[0].shape
# -
# ## 初始化模型参数
#
# 接下来,我们初始化模型参数。隐藏单元个数 `num_hiddens`是一个超参数。
# + attributes={"classes": [], "id": "", "n": "4"}
num_inputs, num_hiddens, num_outputs = vocab_size, 256, vocab_size
ctx = d2l.try_gpu()
print('will use', ctx)
def get_params():
def _one(shape):
return nd.random.normal(scale=0.01, shape=shape, ctx=ctx)
# 隐藏层参数
W_xh = _one((num_inputs, num_hiddens))
W_hh = _one((num_hiddens, num_hiddens))
b_h = nd.zeros(num_hiddens, ctx=ctx)
# 输出层参数
W_hq = _one((num_hiddens, num_outputs))
b_q = nd.zeros(num_outputs, ctx=ctx)
# 附上梯度
params = [W_xh, W_hh, b_h, W_hq, b_q]
for param in params:
param.attach_grad()
return params
# -
# ## 定义模型
#
# 我们根据循环神经网络的计算表达式实现该模型。首先定义`init_rnn_state`函数来返回初始化的隐藏状态。它返回由一个形状为(批量大小, 隐藏单元个数)的值为0的`NDArray`组成的元组。使用元组是为了更便于处理隐藏状态含有多个`NDArray`的情况。
# + attributes={"classes": [], "id": "", "n": "5"}
def init_rnn_state(batch_size, num_hiddens, ctx):
return (nd.zeros(shape=(batch_size, num_hiddens), ctx=ctx), )
# -
# 下面的`rnn`函数定义了在一个时间步里如何计算隐藏状态和输出。这里的激活函数使用了tanh函数。[“多层感知机”](../chapter_deep-learning-basics/mlp.ipynb)一节中介绍过,当元素在实数域上均匀分布时,tanh函数值的均值为0。
# + attributes={"classes": [], "id": "", "n": "6"}
def rnn(inputs, state, params):
# inputs和outputs皆为num_steps个形状为(batch_size, vocab_size)的矩阵
W_xh, W_hh, b_h, W_hq, b_q = params
H, = state
outputs = []
for X in inputs:
H = nd.tanh(nd.dot(X, W_xh) + nd.dot(H, W_hh) + b_h)
Y = nd.dot(H, W_hq) + b_q
outputs.append(Y)
return outputs, (H,)
# -
# 做个简单的测试来观察输出结果的个数(时间步数),以及第一个时间步的输出层输出的形状和隐藏状态的形状。
# + attributes={"classes": [], "id": "", "n": "7"}
state = init_rnn_state(X.shape[0], num_hiddens, ctx)
inputs = to_onehot(X.as_in_context(ctx), vocab_size)
params = get_params()
outputs, state_new = rnn(inputs, state, params)
len(outputs), outputs[0].shape, state_new[0].shape
# -
# ## 定义预测函数
#
# 以下函数基于前缀`prefix`(含有数个字符的字符串)来预测接下来的`num_chars`个字符。这个函数稍显复杂,其中我们将循环神经单元`rnn`设置成了函数参数,这样在后面小节介绍其他循环神经网络时能重复使用这个函数。
# + attributes={"classes": [], "id": "", "n": "8"}
# 本函数已保存在d2lzh包中方便以后使用
def predict_rnn(prefix, num_chars, rnn, params, init_rnn_state,
num_hiddens, vocab_size, ctx, idx_to_char, char_to_idx):
state = init_rnn_state(1, num_hiddens, ctx)
output = [char_to_idx[prefix[0]]]
for t in range(num_chars + len(prefix) - 1):
# 将上一时间步的输出作为当前时间步的输入
X = to_onehot(nd.array([output[-1]], ctx=ctx), vocab_size)
# 计算输出和更新隐藏状态
(Y, state) = rnn(X, state, params)
# 下一个时间步的输入是prefix里的字符或者当前的最佳预测字符
if t < len(prefix) - 1:
output.append(char_to_idx[prefix[t + 1]])
else:
output.append(int(Y[0].argmax(axis=1).asscalar()))
return ''.join([idx_to_char[i] for i in output])
# -
# 我们先测试一下`predict_rnn`函数。我们将根据前缀“分开”创作长度为10个字符(不考虑前缀长度)的一段歌词。因为模型参数为随机值,所以预测结果也是随机的。
# + attributes={"classes": [], "id": "", "n": "9"}
predict_rnn('分开', 10, rnn, params, init_rnn_state, num_hiddens, vocab_size,
ctx, idx_to_char, char_to_idx)
# -
# ## 裁剪梯度
#
# 循环神经网络中较容易出现梯度衰减或梯度爆炸。我们会在[“通过时间反向传播”](bptt.ipynb)一节中解释原因。为了应对梯度爆炸,我们可以裁剪梯度(clip gradient)。假设我们把所有模型参数梯度的元素拼接成一个向量 $\boldsymbol{g}$,并设裁剪的阈值是$\theta$。裁剪后的梯度
#
# $$ \min\left(\frac{\theta}{\|\boldsymbol{g}\|}, 1\right)\boldsymbol{g}$$
#
# 的$L_2$范数不超过$\theta$。
# + attributes={"classes": [], "id": "", "n": "10"}
# 本函数已保存在d2lzh包中方便以后使用
def grad_clipping(params, theta, ctx):
norm = nd.array([0], ctx)
for param in params:
norm += (param.grad ** 2).sum()
norm = norm.sqrt().asscalar()
if norm > theta:
for param in params:
param.grad[:] *= theta / norm
# -
# ## 困惑度
#
# 我们通常使用困惑度(perplexity)来评价语言模型的好坏。回忆一下[“softmax回归”](../chapter_deep-learning-basics/softmax-regression.ipynb)一节中交叉熵损失函数的定义。困惑度是对交叉熵损失函数做指数运算后得到的值。特别地,
#
# * 最佳情况下,模型总是把标签类别的概率预测为1,此时困惑度为1;
# * 最坏情况下,模型总是把标签类别的概率预测为0,此时困惑度为正无穷;
# * 基线情况下,模型总是预测所有类别的概率都相同,此时困惑度为类别个数。
#
# 显然,任何一个有效模型的困惑度必须小于类别个数。在本例中,困惑度必须小于词典大小`vocab_size`。
#
# ## 定义模型训练函数
#
# 与之前章节的模型训练函数相比,这里的模型训练函数有以下几点不同:
#
# 1. 使用困惑度评价模型。
# 2. 在迭代模型参数前裁剪梯度。
# 3. 对时序数据采用不同采样方法将导致隐藏状态初始化的不同。相关讨论可参考[“语言模型数据集(周杰伦专辑歌词)”](lang-model-dataset.ipynb)一节。
#
# 另外,考虑到后面将介绍的其他循环神经网络,为了更通用,这里的函数实现更长一些。
# + attributes={"classes": [], "id": "", "n": "11"}
# 本函数已保存在d2lzh包中方便以后使用
def train_and_predict_rnn(rnn, get_params, init_rnn_state, num_hiddens,
vocab_size, ctx, corpus_indices, idx_to_char,
char_to_idx, is_random_iter, num_epochs, num_steps,
lr, clipping_theta, batch_size, pred_period,
pred_len, prefixes):
if is_random_iter:
data_iter_fn = d2l.data_iter_random
else:
data_iter_fn = d2l.data_iter_consecutive
params = get_params()
loss = gloss.SoftmaxCrossEntropyLoss()
for epoch in range(num_epochs):
if not is_random_iter: # 如使用相邻采样,在epoch开始时初始化隐藏状态
state = init_rnn_state(batch_size, num_hiddens, ctx)
l_sum, n, start = 0.0, 0, time.time()
data_iter = data_iter_fn(corpus_indices, batch_size, num_steps, ctx)
for X, Y in data_iter:
if is_random_iter: # 如使用随机采样,在每个小批量更新前初始化隐藏状态
state = init_rnn_state(batch_size, num_hiddens, ctx)
else: # 否则需要使用detach函数从计算图分离隐藏状态
for s in state:
s.detach()
with autograd.record():
inputs = to_onehot(X, vocab_size)
# outputs有num_steps个形状为(batch_size, vocab_size)的矩阵
(outputs, state) = rnn(inputs, state, params)
# 连结之后形状为(num_steps * batch_size, vocab_size)
outputs = nd.concat(*outputs, dim=0)
# Y的形状是(batch_size, num_steps),转置后再变成长度为
# batch * num_steps 的向量,这样跟输出的行一一对应
y = Y.T.reshape((-1,))
# 使用交叉熵损失计算平均分类误差
l = loss(outputs, y).mean()
l.backward()
grad_clipping(params, clipping_theta, ctx) # 裁剪梯度
d2l.sgd(params, lr, 1) # 因为误差已经取过均值,梯度不用再做平均
l_sum += l.asscalar() * y.size
n += y.size
if (epoch + 1) % pred_period == 0:
print('epoch %d, perplexity %f, time %.2f sec' % (
epoch + 1, math.exp(l_sum / n), time.time() - start))
for prefix in prefixes:
print(' -', predict_rnn(
prefix, pred_len, rnn, params, init_rnn_state,
num_hiddens, vocab_size, ctx, idx_to_char, char_to_idx))
# -
# ## 训练模型并创作歌词
#
# 现在我们可以训练模型了。首先,设置模型超参数。我们将根据前缀“分开”和“不分开”分别创作长度为50个字符(不考虑前缀长度)的一段歌词。我们每过50个迭代周期便根据当前训练的模型创作一段歌词。
# + attributes={"classes": [], "id": "", "n": "12"}
num_epochs, num_steps, batch_size, lr, clipping_theta = 250, 35, 32, 1e2, 1e-2
pred_period, pred_len, prefixes = 50, 50, ['分开', '不分开']
# -
# 下面采用随机采样训练模型并创作歌词。
# + attributes={"classes": [], "id": "", "n": "13"}
train_and_predict_rnn(rnn, get_params, init_rnn_state, num_hiddens,
vocab_size, ctx, corpus_indices, idx_to_char,
char_to_idx, True, num_epochs, num_steps, lr,
clipping_theta, batch_size, pred_period, pred_len,
prefixes)
# -
# 接下来采用相邻采样训练模型并创作歌词。
# + attributes={"classes": [], "id": "", "n": "19"}
train_and_predict_rnn(rnn, get_params, init_rnn_state, num_hiddens,
vocab_size, ctx, corpus_indices, idx_to_char,
char_to_idx, False, num_epochs, num_steps, lr,
clipping_theta, batch_size, pred_period, pred_len,
prefixes)
# -
# ## 小结
#
# * 可以用基于字符级循环神经网络的语言模型来生成文本序列,例如创作歌词。
# * 当训练循环神经网络时,为了应对梯度爆炸,可以裁剪梯度。
# * 困惑度是对交叉熵损失函数做指数运算后得到的值。
#
#
# ## 练习
#
# * 调调超参数,观察并分析对运行时间、困惑度以及创作歌词的结果造成的影响。
# * 不裁剪梯度,运行本节中的代码,结果会怎样?
# * 将`pred_period`变量设为1,观察未充分训练的模型(困惑度高)是如何创作歌词的。你获得了什么启发?
# * 将相邻采样改为不从计算图分离隐藏状态,运行时间有没有变化?
# * 将本节中使用的激活函数替换成ReLU,重复本节的实验。
#
#
#
# ## 扫码直达[讨论区](https://discuss.gluon.ai/t/topic/989)
#
# 
|
深度学习/d2l-zh-1.1/chapter_recurrent-neural-networks/rnn-scratch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="iSemvd0ljKHS"
# # 4. Network Architecture & Training
# + [markdown] id="msYjbPfCjKHU"
# Welcome to the fourth notebook of our six part series part of our tutorial on Deep Learning for Human Activity Recognition. Within the last notebook you learned:
#
# - What are common evaluation metrics when evaluating the performance of an Human Activity Recognition model?
# - How are they defined? How are they computed? How do they differ from each other?
#
# This notebook will teach you everything you need to know about how neural networks are defined and trained using [PyTorch](https://pytorch.org/). As mentioned during the [theoretical part](https://https://mariusbock.github.io/dl-for-har/) of this session, we will not go into detail about each building block of a neural network and how a network is trained, but rather stick to a basic level of understanding. If you want to dig deeper, we recommend you checking out other sources, like [Coursera](https://www.coursera.org/courses?query=deep%20learning) and [YouTube](https://www.youtube.com/results?search_query=deep+learning), as there are plenty of well written tutorials on the fundamentals of Deep Learning. After working through this notebook you will be able to answer the following questions:
#
# - How do I define a sample neural network architecture in PyTorch?
# - What additional preprocessing do I need to apply to my data to fed it into my network?
# - How do I define a train loop which trains my neural network?
# + [markdown] id="s9yp-QySq02H"
# ## 4.1. Important Remarks
# + [markdown] id="uwUayaWhq8Gu"
# If you are accessing this tutorial via [Google Colab](https://colab.research.google.com/github/mariusbock/dl-for-har/blob/main/tutorial_notebooks/training.ipynb), first make sure to use Google Colab in English. This will help us to better assist you with issues that might arise during the tutorial. There are two ways to change the default language if it isn't English already:
# 1. On Google Colab, go to `Help` -> `View in English`
# 2. Change the default language of your browser to `English`.
#
# To also ease the communication when communicating errors, enable line numbers within the settings of Colab.
#
# 1. On Google Colab, go to `Tools` -> `Settings` -> `Editor` -> `Show line numbers`
#
# In general, we strongly advise you to use Google Colab as it provides you with a working Python distribution as well as free GPU resources. To make Colab use GPUs, you need to change the current notebooks runtime type via:
#
# - `Runtime` -> `Change runtime type` -> `Dropdown` -> `GPU` -> `Save`
#
# **Hint:** you can auto-complete code in Colab via `ctrl` + `spacebar`
#
# For the live tutorial, we require all participants to use Colab. If you decide to rerun the tutorial at later points and rather want to have it run locally on your machine, feel free to clone our [GitHub repository](https://github.com/mariusbock/dl-for-har).
#
# To get started with this notebook, you need to first run the code cell below. Please set `use_colab` to be `True` if you are accessing this notebook via Colab. If not, please set it to `False`. This code cell will make sure that imports from our GitHub repository will work.
# + id="5rXaaTKSjKHV"
import os, sys
use_colab = True
module_path = os.path.abspath(os.path.join('..'))
if use_colab:
# move to content directory and remove directory for a clean start
# %cd /content/
# %rm -rf dl-for-har
# clone package repository (will throw error if already cloned)
# !git clone https://github.com/mariusbock/dl-for-har.git
# navigate to dl-for-har directory
# %cd dl-for-har/
else:
os.chdir(module_path)
# this statement is needed so that we can use the methods of the DL-ARC pipeline
if module_path not in sys.path:
sys.path.append(module_path)
# + [markdown] id="09SQsOe6jKHV"
# ## 4.2. Defining a Network Architecture
# + [markdown] id="pWrCb-ypjKHW"
# During this tutorial we will use [PyTorch](https://pytorch.org/) as our Deep Learning framework of choice. The open source library is one of the most popular frameworks out there for applying Deep Learning. It has all the necessary building blocks found in neural networks pre-implemented as well as offers a variety of helpful functions which can be used to easily implement your first Deep Learning script with just a few lines of code.
#
# In the following we will define our neural network architecture. Once defined we can use our previously preprocessed sensor-data to train a network which will be able to predict the type of activities being performed for a given sliding window.
#
# As mentioned during the introduction to this chapter, the architecture which we will used is called **DeepConvLSTM** [[1]](#1). The architecture was introduced by <NAME> and <NAME> in 2016 and is to this date a state-of-the-art architecture for applying Deep Learning on Human Activity Recognition. The architecture combines both convolutional and recurrent layers.
#
# The architecture is made of three main parts:
#
# 1. **Convolutional layers:** Convolutional layers are based on filters (e.g. a 2 by 1 matrix) shifting over some input (e.g. a sliding window) resulting in activation feature map. The main idea of convolutions is that they are able to detect a specific type of feature anywhere within the input. Within the original architecture Ordonez and Roggen apply 4 convolutional layers each with 64 filters of size 5 by 1.
# 2. **LSTM layer(s):** After applying convolutional layers, Ordonez and Roggen make us of an LSTM in order to capture time dependencies on features extracted by convolutional operations. An LSTM is a type of neural network which is able to learn temporal dependencies in data via gated mechanisms. The LSTM itself is structured into layers. Within the original architecture Ordonez and Roggen employ a 2-layered LSTM with 128 hidden units.
# 3. **Classification layer:** The output of the LSTM is finally fed into a classifier which is a fully-connected layer and produces the final predictions. Preceeding the classifier, Ordonez and Roggen additionally put a dropout layer, which is a form of regularization. A dropout layer randomly deactivates neurons according to a dropout probability and thus prevents the probability of your network overfitting.
#
# Contradicting to popular belief that one needs at least a 2-layered LSTM when dealing with sequential data, within a recent work of ours, we exhibited that a 1-layered LSTM might be a better suited option when dealing with raw sensor-data [[2]](#2). Therefore, within the next code block, we will define the altered DeepConvLSTM architecture as presented in our paper which **employs a 1-layered instead of 2-layered LSTM**.
#
# In order to give you a better idea of how to define your PyTorch implementation of the DeepConvLSTM, we already defined a [PyTorch module](https://pytorch.org/docs/stable/generated/torch.nn.Module.html) called **DeepConvLSTM** for you to start out with. A PyTorch module typically consists of two main functions - the `init()` and `forward()` function. Within the former all relevant parameters and building blocks of the neural network are defined. Within the latter the parameters and building blocks are put together, i.e. the computation of the network defined. Within the next tasks you will be asked to fill in some of the missing parts of said module function.
# + [markdown] id="qPb2NTEZ7YA5"
# ### Task 1: Implementing the DeepConvLSTM
# 1. Within the `init()` function define the activation function. Use [PyTorch's implementation](https://pytorch.org/docs/stable/generated/torch.nn.ReLU.html) of the ReLU activation function called `ReLU`. Set `inplace=True`. (`lines 17-18`)
# 2. Within the `init()` function define the four convolution layers. Use [PyTorch's implementation](https://pytorch.org/docs/stable/generated/torch.nn.Conv2d.html) of a 2d-convolution called `Conv2d`. Hints on the input and dimensions are given as comments within the code. The filter size should be of size (`filter_width x 1`) (`lines 20-24`)
# 3. Within the `init()` function define the LSTM. Use [PyTorch's implementation](https://pytorch.org/docs/stable/generated/torch.nn.LSTM.html) of a LSTM called `LSTM`. Hints on the input size of the LSTM is given as comments within the code. The `hidden_size` and `num_layers` are given as attributes within the `init()` function. (`lines 26-27`)
# 4. Within the `init()` define the dropout layer. Use [PyTorch's implementation](https://pytorch.org/docs/stable/generated/torch.nn.Dropout.html) of a dropout layer called `Dropout`. Pass the `Dropout` object the `drop_prob` variable defined within the `init()` function (`lines 29-30`)
# 5. Within the `init()` define the classifier, i.e. fully connected layer. Use [PyTorch's implementation](https://pytorch.org/docs/stable/generated/torch.nn.Linear.html) of a fully-connected layer called `Linear`. (`lines 32-33`)
# 6. Fill in the blanks within the `forward()` function. Apply each of the building blocks you defined in the `init()` on your input `x`. (`lines 39-43, 52-53 and 58-60`)
# + id="acRTdIHDjKHW"
from torch import nn
class DeepConvLSTM(nn.Module):
def __init__(self, config):
super(DeepConvLSTM, self).__init__()
# parameters
self.window_size = config['window_size']
self.drop_prob = config['drop_prob']
self.nb_channels = config['nb_channels']
self.nb_classes = config['nb_classes']
self.seed = config['seed']
self.nb_filters = config['nb_filters']
self.filter_width = config['filter_width']
self.nb_units_lstm = config['nb_units_lstm']
self.nb_layers_lstm = config['nb_layers_lstm']
# define activation function
self.relu = nn.ReLU(inplace=True)
# define conv layers
self.conv1 = nn.Conv2d(1, self.nb_filters, (self.filter_width, 1))
self.conv2 = nn.Conv2d(self.nb_filters, self.nb_filters, (self.filter_width, 1))
self.conv3 = nn.Conv2d(self.nb_filters, self.nb_filters, (self.filter_width, 1))
self.conv4 = nn.Conv2d(self.nb_filters, self.nb_filters, (self.filter_width, 1))
# define lstm layers
self.lstm = nn.LSTM(input_size=self.nb_filters * self.nb_channels, hidden_size=self.nb_units_lstm, num_layers=self.nb_layers_lstm)
# define dropout layer
self.dropout = nn.Dropout(self.drop_prob)
# define classifier
self.fc = nn.Linear(self.nb_units_lstm, self.nb_classes)
def forward(self, x):
# reshape data for convolutions
x = x.view(-1, 1, self.window_size, self.nb_channels)
# apply convolution and the activation function
x = self.relu(self.conv1(x))
x = self.relu(self.conv2(x))
x = self.relu(self.conv3(x))
x = self.relu(self.conv4(x))
# sets the final sequence length
final_seq_len = x.shape[2]
# permute dimensions and reshape for LSTM
x = x.permute(0, 2, 1, 3)
x = x.reshape(-1, final_seq_len, self.nb_filters * self.nb_channels)
# apply LSTM (note: it has two outputs!)
x, _ = self.lstm(x)
# reshape data for classifier
x = x.view(-1, self.nb_units_lstm)
# apply dropout and feed data through classifier
x = self.dropout(x)
x = self.fc(x)
# reshape data and return predicted label of last sample within final sequence (determines label of window)
out = x.view(-1, final_seq_len, self.nb_classes)
return out[:, -1, :]
# + [markdown] id="6k2G6nifjKHX"
# ## 4.3. Preparing your data
# + [markdown] id="j56XD_tJjKHX"
# Great, we now have a neural network defined which we can call and use for training! But, there is one essential step missing before moving on towards the training loop - your data needs to be put into the correct format (again). In addition to the preprocessing steps that you know from the previous notebook, we also need to **make sure that our dataset is using the correct data types which are compatible with a GPU**. Furthermore, our data needs to be split into a **training** and **validation** dataset. As you know from the [theoretical part of this section](https://mariusbock.github.io/dl-for-har), within Deep Learning we essentially try to approximate a function. To judge whether the parameterized function we came up with appropriately approximates such underlying function, we validate our network's perfomance on unseen data. If the algorithm still performs well, i.e. predicts the correct labels for the unseen data, we say that we have found a **generalized function**. The next notebook will cover in more detail what different validation methods exist and go into detail why we need and what common pitfalls exist.
#
# The following task will guide you through the necessary preprocessing one needs to apply on top of the [RealWorld (HAR) dataset](https://sensor.informatik.uni-mannheim.de/#dataset_realworld). The first step of loading the data will already be filled out for you. As you can see, we used a predefined method called `load_dataset()`, which is part of the DL-ARC feature stack.
#
# The preprocessing consists of **four essential parts**:
#
# 1. Split the data into a training and validation dataset. The validation dataset is used to gain feedback on the perfomance of the model and functions as unseen data. Results obtained on the validation dataset can be used as an indicator whether the changes you make to a network and/ or its training process are improving or worsening results.
# 2. Apply the sliding window approach on top of the training and validation dataset. As you learned in the previous notebook, we do not classify a single record, but a window of records. The label of the last record within a window defines the label of the window and is our ultimate goal to predict.
# 3. (Optional) Omit the subject identifier column.
# 4. Convert the two datasets into the correct data format so that they are compatible with the GPU.
# + [markdown] id="-OKmg7nmaWKv"
# ### Task 2: Getting your data ready for training
# 1. Split the data into a train and validation dataset. The train dataset shall consist of the data of the first two subjects. The validation dataset shall be the data of the third subject. (`lines 16-19`)
# 2. Segment your train and validation data into windows. Instead of going back to your defined function within the last notebook, you can use our [predefined method](https://github.com/mariusbock/dl-for-har/blob/main/data_processing/sliding_window.py) which is part of the DL-ARC feature stack called `apply_sliding_window`. It is already imported for you. (`lines 26-29`)
# 3. (*Optional*) Omit the first feature column (subject_identifier) from the train and validation dataset. (`lines 35-36`)
# 4. Convert the feature columns of the train and validation to `float32` and label column to `uint8` for GPU compatibility. Use the [built-in function](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.astype.html) of a pandas dataframe called `astype()`. (`lines 41-43`)
#
# + id="zWgC9soBjKHY"
import pandas as pd
import numpy as np
import warnings
from data_processing.sliding_window import apply_sliding_window
from data_processing.preprocess_data import load_dataset
warnings.filterwarnings('ignore')
# data loading (we are using a predefined method called load_dataset, which is part of the DL-ARC feature stack)
X, y, num_classes, class_names, sampling_rate, has_null = load_dataset('rwhar_3sbjs', include_null=True)
# since the method returns features and labels separatley, we need to concat them
data = np.concatenate((X, y[:, None]), axis=1)
# define the train data to be all data belonging to the first two subjects
train_data = data[data[:, 0] <= 1]
# define the validation data to be all data belonging to the third subject
valid_data = data[data[:, 0] == 2]
# settings for the sliding window (change them if you want to!)
sw_length = 50
sw_unit = 'units'
sw_overlap = 50
# apply a sliding window on top of both the train and validation data; you can use our predefined method
# you can import it via from preprocessing.sliding_window import apply_sliding_window
X_train, y_train = apply_sliding_window(train_data[:, :-1], train_data[:, -1], sliding_window_size=sw_length, unit=sw_unit, sampling_rate=50, sliding_window_overlap=sw_overlap)
X_valid, y_valid = apply_sliding_window(valid_data[:, :-1], valid_data[:, -1], sliding_window_size=sw_length, unit=sw_unit, sampling_rate=50, sliding_window_overlap=sw_overlap)
print("\nShape of the train and validation datasets after splitting and windowing: ")
print(X_train.shape, y_train.shape)
print(X_valid.shape, y_valid.shape)
# (optional) omit the first feature column (subject_identifier) from the train and validation dataset
X_train, X_valid = X_train[:, :, 1:], X_valid[:, :, 1:]
print("\nShape of the train and validation feature dataset after splitting and windowing: ")
print(X_train.shape, X_valid.shape)
# convert the features of the train and validation to float32 and labels to uint8 for GPU compatibility
X_train, y_train = X_train.astype(np.float32), y_train.astype(np.uint8)
X_valid, y_valid = X_valid.astype(np.float32), y_valid.astype(np.uint8)
# + [markdown] id="xBiCDiSDjKHY"
# ## 4.4. Training Your Network
# + [markdown] id="vNNnPX2_jKHZ"
# Since we now have brought the data into the correct format, let's train our network with it!
#
# A typical training loop can be divided into three steps:
#
# 1. **Definition:** You define your network, optimizer and loss
# 2. **Training:** Iterating over the number of epochs: you chunk your training data into so-called batches and iteratively feed them through your network. After a batch has been fed through the network, you compute the loss said batch produced. Using the loss you backprogate it through the network using the optimizer which adjusts the weights accordingly.
# 3. **Validation:** After you have processed your whole training dataset, you go on to validate the predictive performance of the network. To do so you again chunk your training and validation data into batches. Iterating over all batches of both all datasets, fed the batches through the trained network and obtain its predictions. **Note:** you only want to obtain predictions and not backpropagate any loss.
#
# The obtained predictions can now be used to calculate standard evaluation metrics such as **precision** and **recall**. Due to being limited in time we will not talk about their computation in great detail during the tutorial. Nevertheless, we created a [separate notebook](https://colab.research.google.com/github/mariusbock/dl-for-har/blob/main/tutorial_notebooks/evaluation.ipynb) for you which covers the most essential evaluation metrics used in HAR. Feel free to work through it if you want to accustom yourself with how each of them is calculated.
#
# The next task will guide you through **implementing your training and validation loop**. It will again have parts missing which you need to fill out, but will already provide you with certain code segments, to ease the task and focus on the essential parts.
# + [markdown] id="vcKPnZBpjKHZ"
# ### Task 3: Define your own train loop
# + [markdown] id="f2qXFHVojKHZ"
# 1. You'll see that we already defined a `config` object which you can use to pass to your network. Nevertheless, there are three values missing, namely the `window_size`, `nb_channels` and `nb_classes`. Define them correctly. (`lines 27-33`)
# 2. Define your DeepConvLSTM network by calling the object we previously defined. Also define the `optimizer` being the [Adam optimizer](https://pytorch.org/docs/stable/optim.html) and `criterion` being the [Cross-Entropy Loss](https://pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html) (`lines 35-36 and 42-44`)
# 3. Define the `DataLoader` objects. The `DataLoader` objects only work with [PyTorch tensor datasets](https://pytorch.org/docs/stable/data.html), which we already defined for you as the `val_dataset` and `train_dataset` variables. Pass the `DataLoader` object the dataset variables, the `batch_size` you want to use and set `shuffle=True`. (`lines 43-49`)
# 4. Further define the training loop by iterating over the training `DataLoader` object. We already defined parts for you. In a nutshell: for each batch, compute the loss by passing it through the network; backprogate the computed loss using your optimizer object. Use the [.backward()](https://pytorch.org/docs/stable/autograd.html) of the loss object and [.step()](https://pytorch.org/docs/stable/optim.html) of the optimizer to do so. (`lines 69-70 and 75-78`)
# 5. While training obtain predictions for the train dataset. To do so obtain the final predicitons for each batch by applying the PyTorch `softmax` [function](https://pytorch.org/docs/stable/generated/torch.nn.functional.softmax.html) on top of the network output. (`lines 80-81`)
# 6. After training obtain predictions for the validation dataset using the resulting trained network of the current epoch. Iterate again over the validation `DataLoader` object and fed each batch through the network. In addition to calculating the [cross-entropy loss](https://pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html), obtain the final predicitons for each batch by applying the PyTorch `softmax` [function](https://pytorch.org/docs/stable/generated/torch.nn.functional.softmax.html) on top of the network output. Using the predictions the script will calculate [accuracy](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html), [precision](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_score.html), [recall](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.recall_score.html) and [F1-score](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html) on both your training and validation set. (`lines 113-114 and 119-120`)
# 7. Play around with different values for the parameters within the `config` file. How does each one of them influence your training loop? Feel free to also use a completly different optimizer - you can find all the different options on the [PyTorch website](https://pytorch.org/docs/stable/optim.html). (`lines 10-25`)
# + id="3hqs2TL0jKHa"
import torch
from torch.utils.data import DataLoader
from sklearn.metrics import precision_score, recall_score, f1_score, jaccard_score
from misc.torchutils import seed_torch
import time
# this is the config object which contains all relevant settings. Feel free to change them and see how it influences
# your results. Parameters which shouldn't be changed are marked.
config = {
'nb_filters': 64,
'filter_width': 11,
'nb_units_lstm': 128,
'nb_layers_lstm': 1,
'drop_prob': 0.5,
'seed': 1,
'epochs': 20,
'batch_size': 100,
'learning_rate': 1e-4,
'weight_decay': 1e-6,
'gpu_name': 'cuda:0',
'print_counts': False
}
# in order to get reproducible results, we need to seed torch and other random parts of our implementation
seed_torch(config['seed'])
# define the missing parameters within the config file.
# window_size = size of the sliding window in units
# nb_channels = number of feature channels
# nb_classes = number of classes that can be predicted
config['window_size'] = X_train.shape[1]
config['nb_channels'] = X_train.shape[2]
config['nb_classes'] = len(class_names)
# initialize your DeepConvLSTM object
network = DeepConvLSTM(config)
# sends network to the GPU and sets it to training mode
network.to(config['gpu_name'])
network.train()
# initialize the optimizer and loss
optimizer = torch.optim.Adam(network.parameters(), lr=config['learning_rate'], weight_decay=config['weight_decay'])
criterion = nn.CrossEntropyLoss()
# initializes the train and validation dataset in Torch format
train_dataset = torch.utils.data.TensorDataset(torch.from_numpy(X_valid).float(), torch.from_numpy(y_valid))
val_dataset = torch.utils.data.TensorDataset(torch.from_numpy(X_train), torch.from_numpy(y_train))
# define the train- and valloader; use from torch.utils.data import DataLoader
trainloader = DataLoader(train_dataset, batch_size=config['batch_size'], shuffle=True)
valloader = DataLoader(val_dataset, batch_size=config['batch_size'], shuffle=True)
# define your training loop; iterates over the number of epochs
for e in range(config['epochs']):
# helper objects needed for proper documentation
train_losses = []
train_preds = []
train_gt = []
start_time = time.time()
batch_num = 1
# iterate over the trainloader object (it'll return batches which you can use)
for i, (x, y) in enumerate(trainloader):
# sends batch x and y to the GPU
inputs, targets = x.to(config['gpu_name']), y.to(config['gpu_name'])
optimizer.zero_grad()
# send inputs through network to get predictions
train_output = network(inputs)
# calculates loss
loss = criterion(train_output, targets.long())
# backprogate your computed loss through the network
# use the .backward() and .step() function on your loss and optimizer
loss.backward()
optimizer.step()
# calculate actual predictions (i.e. softmax probabilites); use torch.nn.functional.softmax()
train_output = torch.nn.functional.softmax(train_output, dim=1)
# appends the computed batch loss to list
train_losses.append(loss.item())
# creates predictions and true labels; appends them to the final lists
y_preds = np.argmax(train_output.cpu().detach().numpy(), axis=-1)
y_true = targets.cpu().numpy().flatten()
train_preds = np.concatenate((np.array(train_preds, int), np.array(y_preds, int)))
train_gt = np.concatenate((np.array(train_gt, int), np.array(y_true, int)))
# prints out every 100 batches information about the current loss and time per batch
if batch_num % 100 == 0 and batch_num > 0:
cur_loss = np.mean(train_losses)
elapsed = time.time() - start_time
print('| epoch {:3d} | {:5d} batches | ms/batch {:5.2f} | train loss {:5.2f}'.format(e, batch_num, elapsed * 1000 / config['batch_size'], cur_loss))
start_time = time.time()
batch_num += 1
# helper objects
val_preds = []
val_gt = []
val_losses = []
# sets network to eval mode and
network.eval()
with torch.no_grad():
# iterate over the valloader object (it'll return batches which you can use)
for i, (x, y) in enumerate(valloader):
# sends batch x and y to the GPU
inputs, targets = x.to(config['gpu_name']), y.to(config['gpu_name'])
# send inputs through network to get predictions
val_output = network(inputs)
# calculates loss by passing criterion both predictions and true labels
val_loss = criterion(val_output, targets.long())
# calculate actual predictions (i.e. softmax probabilites); use torch.nn.functional.softmax() on dim=1
val_output = torch.nn.functional.softmax(val_output, dim=1)
# appends validation loss to list
val_losses.append(val_loss.item())
# creates predictions and true labels; appends them to the final lists
y_preds = np.argmax(val_output.cpu().numpy(), axis=-1)
y_true = targets.cpu().numpy().flatten()
val_preds = np.concatenate((np.array(val_preds, int), np.array(y_preds, int)))
val_gt = np.concatenate((np.array(val_gt, int), np.array(y_true, int)))
# print epoch evaluation results for train and validation dataset
print("\nEPOCH: {}/{}".format(e + 1, config['epochs']),
"\nTrain Loss: {:.4f}".format(np.mean(train_losses)),
"Train Acc: {:.4f}".format(jaccard_score(train_gt, train_preds, average='macro')),
"Train Prec: {:.4f}".format(precision_score(train_gt, train_preds, average='macro')),
"Train Rcll: {:.4f}".format(recall_score(train_gt, train_preds, average='macro')),
"Train F1: {:.4f}".format(f1_score(train_gt, train_preds, average='macro')),
"\nVal Loss: {:.4f}".format(np.mean(val_losses)),
"Val Acc: {:.4f}".format(jaccard_score(val_gt, val_preds, average='macro')),
"Val Prec: {:.4f}".format(precision_score(val_gt, val_preds, average='macro')),
"Val Rcll: {:.4f}".format(recall_score(val_gt, val_preds, average='macro')),
"Val F1: {:.4f}".format(f1_score(val_gt, val_preds, average='macro')))
# if chosen, print the value counts of the predicted labels for train and validation dataset
if config['print_counts']:
print('Predicted Train Labels: ')
print(np.vstack((np.nonzero(np.bincount(train_preds))[0], np.bincount(train_preds)[np.nonzero(np.bincount(train_preds))[0]])).T)
print('Predicted Val Labels: ')
print(np.vstack((np.nonzero(np.bincount(val_preds))[0], np.bincount(val_preds)[np.nonzero(np.bincount(val_preds))[0]])).T)
# set network to train mode again
network.train()
# + [markdown] id="A0pirE87l6vK"
# # References
# + [markdown] id="FUVSpp-7l-QI"
# <a id="1">[1]</a> <NAME> and <NAME>. 2016.
# Deep Convolutional and LSTM Recurrent Neural Networks for Multimodal Wearable Activity Recognition.
# Sensors 16, 1 (2016). https://doi.org/10.3390/s16010115
#
# <a id="2">[2]</a> <NAME>, <NAME>, <NAME>, and <NAME>. 2021. Improving Deep Learning for HAR with Shallow LSTMs. In Proceedings of the 2021 International Symposium on Wearable Computers, ISWC 2021, September 21-26, 2021, ACM. https://doi.org/10.1145/3460421.3480419
|
tutorial_notebooks/solutions/training_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
#skip
! [ -e /content ] && pip install -Uqq self-supervised
# +
#default_exp vision.dino
# -
# # DINO
#
# > DINO: [Emerging Properties in Self-Supervised Vision Transformers](https://arxiv.org/pdf/2104.14294.pdf)
#export
from fastai.vision.all import *
from self_supervised.augmentations import *
from self_supervised.layers import *
from self_supervised.models.vision_transformer import *
# ## Algorithm
# #### DINO
# 
# **Abstract**: In this paper, we question if self-supervised learning provides
# new properties to Vision Transformer (ViT) [18] that
# stand out compared to convolutional networks (convnets).
# Beyond the fact that adapting self-supervised methods to this
# architecture works particularly well, we make the following
# observations: first, self-supervised ViT features contain
# explicit information about the semantic segmentation of an
# image, which does not emerge as clearly with supervised
# ViTs, nor with convnets. Second, these features are also excellent
# k-NN classifiers, reaching 78.3% top-1 on ImageNet
# with a small ViT. Our study also underlines the importance of
# momentum encoder [31], multi-crop training [10], and the
# use of small patches with ViTs. We implement our findings
# into a simple self-supervised method, called DINO, which
# we interpret as a form of self-distillation with no labels.
# We show the synergy between DINO and ViTs by achieving
# 80.1% top-1 on ImageNet in linear evaluation with ViT-Base.
# **Own Summary**: In this paper authors show effectiveness of the combination of DINO framework and ViT based architectures such as ViT and DEIT. There is no contrastive training nor negative pairs, rather ideas such as momentum encoder and multi-crop augmention from `BYOL` and `SWAV` respectively are adapted. They use distillation with a teacher-student configuration, and avoid representation collapse by centering and sharpening target distributions generated by the teacher. 2 large views (~50%) are used as targets and all views (2 large, 4 small) are used for predictions similar to `SWAV`. Centering values and teacher parameters are updated via ema (exponential moving average).
#export
class DINOHead(nn.Module):
'''
copy.deepcopy:
RuntimeError: Only Tensors created explicitly by the user (graph leaves)
support the deepcopy protocol at the moment
https://pytorch.org/docs/stable/generated/torch.nn.utils.weight_norm.html
https://pytorch.org/docs/stable/generated/torch.nn.GELU.html
'''
def __init__(self, in_dim, out_dim, use_bn=False, norm_last_layer=True, nlayers=3, hidden_dim=2048, bottleneck_dim=256):
super().__init__()
nlayers = max(nlayers, 1)
if nlayers == 1:
self.mlp = nn.Linear(in_dim, bottleneck_dim)
else:
layers = [nn.Linear(in_dim, hidden_dim)]
if use_bn:
layers.append(nn.BatchNorm1d(hidden_dim))
layers.append(nn.GELU())
for _ in range(nlayers - 2):
layers.append(nn.Linear(hidden_dim, hidden_dim))
if use_bn:
layers.append(nn.BatchNorm1d(hidden_dim))
layers.append(nn.GELU())
layers.append(nn.Linear(hidden_dim, bottleneck_dim))
self.mlp = nn.Sequential(*layers)
self.apply(self._init_weights)
self.last_layer = nn.utils.weight_norm(nn.Linear(bottleneck_dim, out_dim, bias=False))
self.last_layer.weight_g.data.fill_(1)
if norm_last_layer:
self.last_layer.weight_g.requires_grad = False
def _init_weights(self, m):
if isinstance(m, nn.Linear):
trunc_normal_(m.weight, std=.02)
if isinstance(m, nn.Linear) and m.bias is not None:
nn.init.constant_(m.bias, 0)
def forward(self, x):
x = self.mlp(x)
x = nn.functional.normalize(x, dim=-1, p=2)
x = self.last_layer(x)
return x
#export
@delegates(get_multi_aug_pipelines, but=['n', 'size', 'resize_scale'])
def get_dino_aug_pipelines(num_crops=(2,4), crop_sizes=(224,96), min_scales=(0.4,0.05), max_scales=(1.,0.4), **kwargs):
aug_pipelines = []
for nc, size, mins, maxs in zip(num_crops, crop_sizes, min_scales, max_scales):
aug_pipelines += get_multi_aug_pipelines(n=nc, size=size, resize_scale=(mins,maxs), **kwargs)
return aug_pipelines
aug_pipelines = get_dino_aug_pipelines()
#export
class DINOModel(Module):
def __init__(self, student, teacher):
"A module for loading and saving all training params together"
self.student,self.teacher = student,teacher
self.teacher.load_state_dict(student.state_dict())
for p in self.teacher.parameters(): p.requires_grad = False
self.register_buffer('C', torch.zeros(1,num_features_model(teacher)))
def forward(self,x): return self.student(x)
bs = 4
x_large = [torch.randn(4,3,224,224)]*2
x_small = [torch.randn(4,3,96,96)]*4
# +
deits16 = deit_small(patch_size=16, drop_path_rate=0.1)
deits16 = MultiCropWrapper(deits16)
dino_head = DINOHead(deits16.encoder.embed_dim, 2**16, norm_last_layer=True)
student_model = nn.Sequential(deits16,dino_head)
deits16 = deit_small(patch_size=16)
deits16 = MultiCropWrapper(deits16)
dino_head = DINOHead(deits16.encoder.embed_dim, 2**16, norm_last_layer=True)
teacher_model = nn.Sequential(deits16,dino_head)
dino_model = DINOModel(student_model, teacher_model)
# -
dino_model.student[1]
#export
class DINO(Callback):
order,run_valid = 9,True
def __init__(self, aug_pipelines, large_crop_ids=[0,1],
cmom=0.9,
tmom_start=0.996, tmom_end=1., tmom_sched=SchedCos,
tpt_start=0.04, tpt_end=0.04, tpt_warmup_pct=0., tpt_sched=SchedLin,
tps=0.1,
freeze_last_layer=1,
print_augs=False):
"""
DINO teacher student training with distillation.
Refer to original repo:
https://github.com/facebookresearch/dino/blob/0be6e112dd579203caaa1d0f066e29ca536f76dd/main_dino.py#L41
cmom: Center update momentum.
tmom: Teacher update momentum. Set larger, e.g. 0.9995, for small batches or 0.996 for large batches (256+).
tpt_warmup: Warm up starting temperature
tpt_warmup_pct: Percentage of training for warmup
tpt_sched: Warm up scheduler, e.g. SchedLin, SchedCos, SchedExp
tpt: Teacher temperature after warm up. Decrease if training loss does not decrease.
Smaller temperature means more sharpening.
tps: Student temperature.
freeze_last_layer: How many epochs to freeze the last layer
"""
store_attr('large_crop_ids,cmom,freeze_last_layer,tps')
self.augs = aug_pipelines
self.tpt_scheduler = combine_scheds([tpt_warmup_pct,1-tpt_warmup_pct],
[tpt_sched(tpt_start,tpt_end),SchedNo(tpt_end,tpt_end)])
self.tmom_scheduler = tmom_sched(tmom_start, tmom_end)
if print_augs:
for aug in self.augs: print(aug)
def before_fit(self):
"Create teacher model as a copy of student"
self.learn.loss_func = self.lf
self.tpt = self.tpt_scheduler(0.)
self.tmom = self.tmom_scheduler(0.)
self.model.teacher.eval()
for n,p in self.learn.model.student[1].last_layer.named_parameters():
if n == 'weight_v' : p.requires_grad = False
def before_batch(self):
"Augment multi crop views"
self.bs = self.x.size(0)
self.learn.xb = ([aug(self.x) for aug in self.augs],)
x_large = [self.learn.xb[0][i] for i in self.large_crop_ids]
# TODO: Do we need to put the teacher in eval(), not it original repo?
with torch.no_grad():
targs = self.model.teacher(x_large)
self.learn.yb = (targs,)
self.cb = targs.mean(0, keepdim=True)
def _momentum_update_teacher(self):
for param_s, param_t in zip(self.learn.model.student.parameters(), self.model.teacher.parameters()):
param_t.data = param_t.data * self.tmom + param_s.data * (1. - self.tmom)
def _momentum_update_center(self):
self.model.C = self.model.C*self.cmom + self.cb*(1-self.cmom)
def after_step(self):
"Center and teacher updates"
self._momentum_update_teacher(); self._momentum_update_center()
def after_epoch(self):
"Update tpt at the end of each epoch"
self.tpt = self.tpt_scheduler(self.pct_train)
self.tmom = self.tmom_scheduler(self.pct_train)
if self.epoch == self.freeze_last_layer:
print("Setting last layer to trainable")
for n,p in self.learn.model.student[1].last_layer.named_parameters():
if n == 'weight_v' : p.requires_grad = True
def lf(self, pred, *yb):
"Multi crop cross entropy loss: -qlog(p)"
yb = yb[0]
pred = F.log_softmax(pred / self.tps, dim=-1)
yb = F.softmax((yb - self.model.C) / self.tpt, dim=-1)
n_targs, n_preds = yb.size(0)//self.bs, pred.size(0)//self.bs
yb, pred = yb.chunk(n_targs), pred.chunk(n_preds)
loss, npairs = 0, n_targs*(n_preds-1)
for ti in range(n_targs):
for pi in range(n_preds):
if ti != pi:
loss += (-yb[ti]*pred[pi]).sum(-1).mean() / npairs
return loss
@torch.no_grad()
def show(self, n=1):
xbs = self.learn.xb[0]
idxs = np.random.choice(range(self.bs), n, False)
images = [aug.decode(xb.to('cpu').clone()).clamp(0, 1)[i]
for i in idxs
for xb, aug in zip(xbs, self.augs)]
return show_batch(images[0], None, images, max_n=len(images), nrows=n)
# Training schedule for DINO
fig,ax = plt.subplots(1,2,figsize=(15,5))
lr_sched = combine_scheds([0.1,0.9], [SchedLin(0.,1e-3), SchedCos(1e-3,1e-6)])
ax[0].plot([lr_sched(i) for i in np.linspace(0,1,100)]);ax[0].set_title('lr')
wd_sched = SchedCos(0.04,0.4)
ax[1].plot([wd_sched(i) for i in np.linspace(0,1,100)]);ax[1].set_title('wd');
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
|
nbs/15 - dino.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
df = pd.read_stata('text_example_final.dta')
journal1 = df[df['source']==u'管理世界']
journal2 = df[df['source']==u'经济研究']
df.head()
# +
def get_keywords(df):
dic = {}
for keywords in df['keyword']:
keywords = keywords.split(';;')
for keyword in keywords:
if(keyword == ""):
continue
if keyword not in dic:
dic[keyword] = 0
dic[keyword] = dic[keyword] + 1
lis = []
for keyword in dic.keys():
lis.append((dic[keyword], keyword))
lis = sorted(lis)
lis.reverse()
return dic, lis
d2017, l2017 = get_keywords(df[df['year']==2017])
d2018, l2018 = get_keywords(df[df['year']==2018])
d2019, l2019 = get_keywords(df[df['year']==2019])
keyword_dic = {}
for _,keyword in get_keywords(df)[1][0:10]:
keyword_dic[keyword] = 0
# first = 5
# for _,keyword in l2017[0:first]:
# keyword_dic[keyword] = 0
# for _,keyword in l2018[0:first]:
# keyword_dic[keyword] = 0
# for _,keyword in l2019[0:first]:
# keyword_dic[keyword] = 0
pd.DataFrame(l2017).head()
# +
from matplotlib import pyplot as plt
import matplotlib as mpl
import numpy as np
plt.rcParams['font.family']='sans-serif'
plt.rcParams['font.sans-serif']=['SimHei'] #用来正常显示中文标签
plt.rcParams['axes.unicode_minus']=False #用来正常显示负号
dpi = 200
fig = plt.figure(figsize=(1600/dpi, 1600/dpi), dpi=dpi)
for keyword in keyword_dic.keys():
x = [2017, 2018, 2019]
print(keyword)
y = [d2017.get(keyword) or 0,d2018.get(keyword) or 0, d2019.get(keyword) or 0]
plt.plot(x,y, label=keyword)
plt.legend(loc=[1.1, 0])
plt.xticks([2017, 2018, 2019])
plt.yticks(np.arange(0, 16, 1))
plt.savefig('p3_1a.png', figsize=(1600/dpi, 1600/dpi), dpi=dpi)
fig.set_facecolor("#FFFFFF")
print(journal1['year'].min(), journal1['year'].max())
print(journal2['year'].min(), journal2['year'].max())
# -
data = get_keywords(df)[1][:20]
dpi = 200
fig = plt.figure(figsize=(1600/dpi, 400/dpi), dpi=dpi)
plt.xticks(rotation=90)
plt.bar([d[1] for d in data],[d[0] for d in data])
fig.set_facecolor("#FFFFFF")
|
NLTK/p3_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2 with Spark 2.0
# language: python
# name: python2-spark20
# ---
# !pip install --user xlrd
# +
import Common_Functions
fundDF = pd.read_excel(Common_Functions.getFileFromObjectStorage('MizuhoPOC', 'Funds.xlsm'),header=[0]).rename(index=str, columns={"ALADDIN": "ID"})
fundDF.head(5)
# +
import pandas as pd
from io import BytesIO
import requests
import json
import xlrd
from pyspark.sql.functions import *
from datetime import datetime
from dateutil.parser import parse
from ingest.Connectors import Connectors
# +
# The code was removed by DSX for sharing.
# +
# The code was removed by DSX for sharing.
# +
fundDF = pd.read_excel(getFileFromObjectStorage('MizuhoPOC', 'Funds.xlsm'),header=[0]).rename(index=str, columns={"ALADDIN": "ID"})
fundDF.head(5)
# +
spark = SparkSession.builder.getOrCreate()
mhcbSparkDF = spark.createDataFrame(fundDF)
mhcbSparkDF.printSchema()
# Connection to Dash DB for writing the data
dashdbsaveoption = {
Connectors.DASHDB.HOST : dashCredentials["host"],
Connectors.DASHDB.DATABASE : dashCredentials["db"],
Connectors.DASHDB.USERNAME : dashCredentials["username"],
Connectors.DASHDB.PASSWORD : dashCredentials["password"],
Connectors.DASHDB.TARGET_TABLE_NAME : dashCredentials["tableName"],
Connectors.DASHDB.TARGET_WRITE_MODE : 'merge'
}
mhcbDashDBDF = mhcbSparkDF.write.format("com.ibm.spark.discover").options(**dashdbsaveoption).save()
# -
|
clients/Mizuho/Reporting/src/main/resources/Load_Funds_Ref_Table.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# import csv as c
#
# -
enrollment_filename = "/home/pri/Downloads/IntroToDataAnalysis/enrollments.csv"
daily_engagement_filename = "/home/pri/Downloads/IntroToDataAnalysis/daily_engagement.csv"
project_submissions_filename = "/home/pri/Downloads/IntroToDataAnalysis/project_submissions.csv"
def returnFileset(filename):
f = open(filename)
content = c.reader(f)
return content
f.close()
setvalue_daily_engagement_filename = list(returnFileset(daily_engagement_filename))
print("No of rows in the Daily Engagement File",len(setvalue_daily_engagement_filename)-1)
element = setvalue_daily_engagement_filename[1]
uniquecount=-1
for row in setvalue_daily_engagement_filename:
if row[0] != element[0]:
#print("row",row[0])
#print("element",element[0])
element[0] = row[0]
uniquecount=uniquecount+1
#print(uniquecount)
print("Unique account numbers",uniquecount)
setvalue_enrollment_filename = list(returnFileset(enrollment_filename))
print("No of rows in the enrollment File",len(setvalue_enrollment_filename)-1)
elemente = setvalue_enrollment_filename[1]
uniquecount_e=-1
for row in setvalue_enrollment_filename:
if row[0] != elemente[0]:
#print("row",row[0])
#print("element",elemente[0])
elemente[0] = row[0]
uniquecount_e=uniquecount_e+1
#print(uniquecount_e)
print("Unique account numbers",uniquecount_e)
setvalue_project_submissions_filename = list(returnFileset(project_submissions_filename))
print("No of rows in the project submissions File",len(setvalue_project_submissions_filename)-1)
elements = setvalue_project_submissions_filename[1]
uniquecount_s=-1
for row in setvalue_project_submissions_filename:
if row[3] != elements[3]:
# print("row",row[3])
# print("element",elements[3])
elements[3] = row[3]
uniquecount_s=uniquecount_s+1
# print(uniquecount_s)
print("Unique account numbers",uniquecount_s)
|
Data Exploration - Python practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dynamic programming
# This notebook is based on the tutorial on Dynamic programming on this webpage (in Russian): https://bestprogrammer.ru/izuchenie/uchebnik-po-dinamicheskomu-programmirovaniyu-sozdanie-effektivnyh-programm-na-python
# ## Imports
import time
import matplotlib.pyplot as plt
# ## Recursive algorithm for Fibonacci numbers
# +
# %%timeit
def fib(n):
if n <= 0:
return 0
if n == 1:
return 1
else:
return fib(n-1) + fib(n-2)
fib(10)
# -
# ## Recursive algorithm with memoization for Fibonacci numbers
# +
# %%timeit
cache = {}
def fib(n):
if n <= 0:
return 0
if n == 1:
return 1
elif n in cache:
return cache[n]
else:
cache[n] = fib(n-1) + fib(n-2)
return cache[n]
fib(10)
# -
# We can see that with memoization, the runtime decreases from 18 to 3 microseconds, that is, we get a six-fold improvement.
# ## Knapsack problem
# Based on the description from https://en.wikipedia.org/wiki/Knapsack_problem as the original statement is incomprehensible.
#
# We consider the problem of choosing some items from the given $n$ items such that the total value of these items is maximized but the total weight of these items does not exceed a given maximum capacity. This is called **0-1 knapsack problem** because we either take an item (1) or do not (0); we are not allowed to take fractions of the items.
#
# Let's denote by $x_i$ if we take the $ith$ item or not: $x_i \in \{0, 1\}$. The value and the weight of the $i$th item are $v_i$ and $w_i$, respectively. The maximum allowed capacity is $M$.
#
# Then, mathematically we solve the following problem:
# $$
# \begin{align}
# \text{maximize} & \sum_{i=1}^n x_i v_i \\
# \text{subject to} & \sum_{i=1}^n x_i w_i \leq M,
# \end{align}
# $$
# where $x_i \in \{0, 1\}$.
# The following implementation and test problem follows https://en.wikipedia.org/wiki/Knapsack_problem
def solve_knapsack_problem(v, w, M):
"""Solve knapsack problem with values `v`, weights `w`, and capacity `M`."""
n = len(v)
assert len(v) == len(w)
cache = {}
for i in (range(0, n)):
cache[i, 0] = 0
for j in (range(0, M+1)):
cache[0, j] = 0
for i in range(1, n):
for j in range(0, M+1):
if w[i] > j:
cache[i, j] = cache[i-1, j]
else:
cache[i, j] = max(
cache[i-1, j],
cache[i-1, j-w[i]] + v[i]
)
return cache[n-1, M]
# +
v = [5, 4, 3, 2]
w = [4, 3, 2, 1]
M = 6
solve_knapsack_problem(v, w, M)
# + [markdown] tags=[]
# ## Coin change problem
# -
|
2021-12-18-dynamic-programming.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Randomness and reproducibility
# + [markdown] raw_mimetype="text/restructuredtext"
# Random numbers and [stochastic processes](http://www2.econ.iastate.edu/tesfatsi/ace.htm#Stochasticity)
# are essential to most agent-based models.
# [Pseudo-random number generators](https://en.wikipedia.org/wiki/Pseudorandom_number_generator)
# can be used to create numbers in a sequence that appears
# random but is actually a deterministic sequence based on an initial seed value.
# In other words, the generator will produce the same pseudo-random sequence
# over multiple runs if it is given the same seed at the beginning.
# Note that is possible that the generators will draw the same number repeatedly,
# as illustrated in this [comic strip](https://dilbert.com/strip/2001-10-25) from <NAME>:
#
# 
# -
import agentpy as ap
import numpy as np
import random
# ## Random number generators
# Agentpy models contain two internal pseudo-random number generators with different features:
#
# - `Model.random` is an instance of `random.Random` (more info [here](https://realpython.com/python-random/))
# - `Model.nprandom` is an instance of `numpy.random.Generator` (more info [here](https://numpy.org/devdocs/reference/random/index.html))
# To illustrate, let us define a model that uses both generators to draw a random integer:
class RandomModel(ap.Model):
def setup(self):
self.x = self.random.randint(0, 99)
self.y = self.nprandom.integers(99)
self.report(['x', 'y'])
self.stop()
# If we run this model multiple times, we will likely get a different series of numbers in each iteration:
exp = ap.Experiment(RandomModel, iterations=5)
results = exp.run()
results.reporters
# ## Defining custom seeds
# + [markdown] raw_mimetype="text/restructuredtext"
# If we want the results to be reproducible,
# we can define a parameter `seed` that
# will be used automatically at the beginning of a simulation
# to initialize both generators.
# -
parameters = {'seed': 42}
exp = ap.Experiment(RandomModel, parameters, iterations=5)
results = exp.run()
# By default, the experiment will use this seed to generate different random seeds for each iteration:
results.reporters
# Repeating this experiment will yield the same results:
exp2 = ap.Experiment(RandomModel, parameters, iterations=5)
results2 = exp2.run()
results2.reporters
# Alternatively, we can set the argument `randomize=False` so that the experiment will use the same seed for each iteration:
exp3 = ap.Experiment(RandomModel, parameters, iterations=5, randomize=False)
results3 = exp3.run()
# Now, each iteration yields the same results:
results3.reporters
# ## Sampling seeds
# For a sample with multiple parameter combinations, we can treat the seed like any other parameter.
# The following example will use the same seed for each parameter combination:
parameters = {'p': ap.Values(0, 1), 'seed': 0}
sample1 = ap.Sample(parameters, randomize=False)
list(sample1)
# If we run an experiment with this sample,
# the same iteration of each parameter combination will have the same seed (remember that the experiment will generate different seeds for each iteration by default):
exp = ap.Experiment(RandomModel, sample1, iterations=2)
results = exp.run()
results.reporters
# Alternatively, we can use `Sample` with `randomize=True` (default)
# to generate random seeds for each parameter combination in the sample.
sample3 = ap.Sample(parameters, randomize=True)
list(sample3)
# This will always generate the same set of random seeds:
sample3 = ap.Sample(parameters)
list(sample3)
# An experiment will now have different results for every parameter combination and iteration:
exp = ap.Experiment(RandomModel, sample3, iterations=2)
results = exp.run()
results.reporters
# Repeating this experiment will yield the same results:
exp = ap.Experiment(RandomModel, sample3, iterations=2)
results = exp.run()
results.reporters
# ## Stochastic methods of AgentList
# Let us now look at some stochastic operations that are often used in agent-based models.
# To start, we create a list of five agents:
model = ap.Model()
agents = ap.AgentList(model, 5)
agents
# If we look at the agent's ids, we see that they have been created in order:
agents.id
# + [markdown] raw_mimetype="text/restructuredtext"
# To shuffle this list, we can use `AgentList.shuffle`:
# -
agents.shuffle().id
# + [markdown] raw_mimetype="text/restructuredtext"
# To create a random subset, we can use `AgentList.random`:
# -
agents.random(3).id
# And if we want it to be possible to select the same agent more than once:
agents.random(6, replace=True).id
# ## Agent-specific generators
# + [markdown] raw_mimetype="text/restructuredtext"
# For more advanced applications, we can create separate generators for each object.
# We can ensure that the seeds of each object follow a controlled pseudo-random sequence by using the models' main generator to generate the seeds.
# +
class RandomAgent(ap.Agent):
def setup(self):
seed = self.model.random.getrandbits(128) # Seed from model
self.random = random.Random(seed) # Create agent generator
self.x = self.random.random() # Create a random number
class MultiRandomModel(ap.Model):
def setup(self):
self.agents = ap.AgentList(self, 2, RandomAgent)
self.agents.record('x')
self.stop()
# -
parameters = {'seed': 42}
exp = ap.Experiment(
MultiRandomModel, parameters, iterations=2,
record=True, randomize=False)
results = exp.run()
results.variables.RandomAgent
# Alternatively, we can also have each agent start from the same seed:
# +
class RandomAgent2(ap.Agent):
def setup(self):
self.random = random.Random(self.p.agent_seed) # Create agent generator
self.x = self.random.random() # Create a random number
class MultiRandomModel2(ap.Model):
def setup(self):
self.agents = ap.AgentList(self, 2, RandomAgent2)
self.agents.record('x')
self.stop()
# -
parameters = {'agent_seed': 42}
exp = ap.Experiment(
MultiRandomModel2, parameters, iterations=2,
record=True, randomize=False)
results = exp.run()
results.variables.RandomAgent2
|
docs/guide_random.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3
# language: python
# name: python3
# ---
# # Calculating Multi-Modal Concordance
from azureml.core import Workspace
from model_drift import settings
from model_drift.azure_utils import download_metrics_file
from model_drift.drift.io import load_metrics_file, load_weights, load_stats
from model_drift.drift.unify import calculate_mmc
from model_drift.helpers import filter_columns
# ## Calculating MMC
#
# We first load the results from the csv file, and pick which aggregate measure we wish to use. Options are:
# - `"mean"` (default) the aggregate mean metric after N over-sampled trials
# - `"median"` the aggregate mean metric after N over-sampled trials
# - `"obs"` for observed metrics (no resampling)
#
# From these metrics, we pick out the performance columns and filter the metrics dataframe to only include the desired individual metrics.
# In this case, we use only the `"distance"` metrics from KS test and Chi-Square tests.
#
# Next, we load our values for shift ($\zeta_i$) and scale ($\eta_i$), as well as our metric weights ($\alpha_i$).
#
# Finally, we calculate the MMC from these metrics using the helper function `model_drift.drift.unify.calculate_mmc`.
# This function uses the weights and standardization values to calculate MMC as follows:
# $$
# \mathit{MMC}(\omega) = \sum_{i=1}^{L} \alpha_i \cdot \Gamma_i\bigg(\hat{\psi}_i(\omega)\bigg) = \sum_{i=1}^{L}\frac{\alpha_i}{\eta_i}\bigg(\hat{\psi}_i(\omega)-\zeta_i\bigg)
# $$
# For $\Gamma(\cdot)$ we used `model_drift.drift.unify.standardize`, and the weighted sum is handled by `model_drift.drift.unify.w_avg`
#
# Results Files
res_unmod = "tough_balloon_jgphht2z" # Unmodified PadChest
res_q100 = "serene_net_cb97mxt8" # q100
res_q25 = "busy_sun_f8s4yt3h" # q25
res_q5 = "good_stamp_bdz1lskm" #q5
res_lateral = "shy_shampoo_rf8w34nf" # lateral
# +
res = res_q5
results_csv = settings.RESULTS_DIR.joinpath('drift', res+".csv")
# Parameters
performance_col = ("performance", "micro avg", "auroc")
weights_file = settings.MODEL_DIR.joinpath("weights", "metric_weights.csv")
stats_file = settings.MODEL_DIR.joinpath("weights", "std_stats.csv")
include_metrics = ["distance"]
# Load individual metrics
error_df, results_df = load_metrics_file(results_csv, which="mean")
# Get performance from results
perf_df = results_df[performance_col]
error_perf_df = error_df[performance_col]
# Filter metrics to only desired individual metrics
metrics_df = filter_columns(results_df, exclude=['performance', 'count'])
metrics_df = filter_columns(metrics_df, include=include_metrics)
# Load weights, shifts and offsets
weights = load_weights(weights_file)
std_stats = load_stats(stats_file)
#calculate mmc
mmc = calculate_mmc(metrics_df, weights=weights, std_stats=std_stats)
mmc.plot(figsize=(15, 6))
# -
# ## Interactive Plot using Plotly
# +
from model_drift.figure_helper import FigureHelper
import pandas as pd
# We smooth for aesthetics when plotting
def smooth(y: pd.DataFrame, span=7):
if span > 0:
ys = y.ewm(span=span, ignore_na=False).mean()
ys[y.isna()] = None
else:
ys = y
return ys
span = 7
graph_start = "2014-04-01"
graph_end = "2015-02-01"
x = pd.date_range(mmc.index.min(), mmc.index.max())
fh = FigureHelper(x, color_list=['#636EFA', '#EF553B', '#00CC96', '#AB63FA', '#8C564B', '#7f7f7f'])
perf_df = perf_df.reindex(x)
error_perf_df = error_perf_df.reindex(x)
mmc = mmc.reindex(x)
fh.add_trace(y=smooth(perf_df, span=span),
yu=smooth(perf_df+error_perf_df, span=span),
yl=smooth(perf_df-error_perf_df, span=span), name=run, connectgaps=False, row=1, col=1, showlegend=True)
fh.add_trace(y=smooth(mmc, span=span), name=run, connectgaps=False, row=2, col=1)
fig = fh.make_fig(shared_xaxes=True, vertical_spacing=0.025)
fig.update_xaxes(showspikes=True, spikecolor="black", spikesnap="cursor", spikemode="across", spikethickness=1,
range=[graph_start, graph_end])
fig.update_layout(spikedistance=1000, height=500, plot_bgcolor="#E8E8EA", title="Multi-Modal Data Concordance")
xaxis = dict(
tickformat='%Y-%m-%d',
tickmode='linear',
dtick="M1"
)
fig.update_layout(
xaxis1=xaxis,
xaxis2=xaxis,
)
fig.show()
# -
# ## Download results from AzureML
# Skip this step if you are running locally.
# Get results file from azure ml
workspace = Workspace.from_config(settings.AZUREML_CONFIG)
experiment = "generate-drift-metrics"
run = "<run_name>"
results_csv = download_metrics_file(run, settings.RESULTS_DIR.joinpath('drift'), experiment, workspace, overwrite=False)
|
notebooks/calculate_drift/calculate-mmc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Arbitrage Pricing Theory
#
# By <NAME>" Nitishinskaya, <NAME>, and <NAME>.
#
# Part of the Quantopian Lecture Series:
#
# * [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
# * [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
#
# Notebook released under the Creative Commons Attribution 4.0 License.
#
# ---
#
# Arbitrage pricing theory is a major asset pricing theory that relies on expressing the returns using a linear factor model:
#
# $$R_i = a_i + b_{i1} F_1 + b_{i2} F_2 + \ldots + b_{iK} F_K + \epsilon_i$$
#
# This theory states that if we have modelled our rate of return as above, then the expected returns obey
#
# $$ E(R_i) = R_F + b_{i1} \lambda_1 + b_{i2} \lambda_2 + \ldots + b_{iK} \lambda_K $$
#
# where $R_F$ is the risk-free rate, and $\lambda_j$ is the risk premium - the return in excess of the risk-free rate - for factor $j$. This premium arises because investors require higher returns to compensate them for incurring risk. This generalizes the capital asset pricing model (CAPM), which uses the return on the market as its only factor.
#
# We can compute $\lambda_j$ by constructing a portfolio that has a sensitivity of 1 to factor $j$ and 0 to all others (called a <i>pure factor portfolio</i> for factor $j$), and measure its return in excess of the risk-free rate. Alternatively, we could compute the factor sensitivities for $K$ well-diversified (no asset-specific risk, i.e. $\epsilon_p = 0$) portfolios, and then solve the resulting system of linear equations.
# ## Arbitrage
#
# There are generally many, many securities in our universe. If we use different ones to compute the $\lambda$s, will our results be consistent? If our results are inconsistent, there is an <i>arbitrage opportunity</i> (in expectation). Arbitrage is an operation that earns a profit without incurring risk and with no net investment of money, and an arbitrage opportunity is an opportunity to conduct such an operation. In this case, we mean that there is a risk-free operation with <i>expected</i> positive return that requires no net investment. It occurs when expectations of returns are inconsistent, i.e. risk is not priced consistently across securities.
#
# For instance, there is an arbitrage opportunity in the following case: say there is an asset with expected rate of return 0.2 for the next year and a $\beta$ of 1.2 with the market, while the market is expected to have a rate of return of 0.1, and the risk-free rate on 1-year bonds is 0.05. Then the APT model tells us that the expected rate of return on the asset should be
#
# $$ R_F + \beta \lambda = 0.05 + 1.2 (0.1 - 0.05) = 0.11$$
#
# This does not agree with the prediction that the asset will have a rate of return of 0.2. So, if we buy \$100 of our asset, short \$120 of the market, and buy \$20 of bonds, we will have invested no net money and are not exposed to any systematic risk (we are market-neutral), but we expect to earn $0.2 \cdot 100 - 0.1 \cdot 120 + 20 \cdot 0.05 = 9$ dollars at the end of the year.
#
# The APT assumes that these opportunities will be taken advantage of until prices shift and the arbitrage opportunities disappear. That is, it assumes that there are arbitrageurs who have sufficient amounts of patience and capital. This provides a justification for the use of empirical factor models in pricing securities: if the model were inconsistent, there would be an arbitrage opportunity, and so the prices would adjust.
# ##Goes Both Ways
#
# Often knowing $E(R_i)$ is incredibly difficult, but notice that this model tells us what the expected returns should be if the market is fully arbitraged. This lays the groundwork for long-short equity strategies based on factor model ranking systems. If you know what the expected return of an asset is given that the market is arbitraged, and you hypothesize that the market will be mostly arbitraged over the timeframe on which you are trading, then you can construct a ranking.
#
# ##Long-Short Equity
#
# To do this, estimate the expected return for each asset on the market, then rank them. Long the top percentile and short the bottom percentile, and you will make money on the difference in returns. Said another way, if the assets at the top of the ranking on average tend to make $5\%$ more per year than the market, and assets at the bottom tend to make $5\%$ less, then you will make $(M + 0.05) - (M - 0.05) = 0.10$ or $10\%$ percent per year, where $M$ is the market return that gets canceled out.
#
# Long-short equity accepts that any individual asset is very difficult to model, relies on broad trends holding true. We can't accurately predict expected returns for an asset, but we can predict the expected returns for a group of 1000 assets as the errors average out.
#
# We will have a full lecture on long-short models later.
#
# ##How many factors do you want?
#
# As discussed in other lectures, noteably Overfitting, having more factors will explain more and more of your returns, but at the cost of being more and more fit to noise in your data. Do discover true signals and make good predictions going forward, you want to select as few parameters as possible that still explain a large amount of the variance in returns.
# ##Example: Computing Expected Returns for Two Assets
import numpy as np
import pandas as pd
from statsmodels import regression
import matplotlib.pyplot as plt
# Let's get some data.
# +
start_date = '2014-06-30'
end_date = '2015-06-30'
# We will look at the returns of an asset one-month into the future to model future returns.
offset_start_date = '2014-07-31'
offset_end_date = '2015-07-31'
# Get returns data for our assets
asset1 = get_pricing('HSC', fields='price', start_date=offset_start_date, end_date=offset_end_date).pct_change()[1:]
asset2 = get_pricing('MSFT', fields='price', start_date=offset_start_date, end_date=offset_end_date).pct_change()[1:]
# Get returns for the market
bench = get_pricing('SPY', fields='price', start_date=start_date, end_date=end_date).pct_change()[1:]
# Use an ETF that tracks 3-month T-bills as our risk-free rate of return
treasury_ret = get_pricing('BIL', fields='price', start_date=start_date, end_date=end_date).pct_change()[1:]
# +
# Define a constant to compute intercept
constant = pd.TimeSeries(np.ones(len(asset1.index)), index=asset1.index)
df = pd.DataFrame({'R1': asset1,
'R2': asset2,
'SPY': bench,
'RF': treasury_ret,
'Constant': constant})
df = df.dropna()
# -
# We'll start by computing static regressions over the whole time period.
# +
OLS_model = regression.linear_model.OLS(df['R1'], df[['SPY', 'RF', 'Constant']])
fitted_model = OLS_model.fit()
print 'p-value', fitted_model.f_pvalue
print fitted_model.params
R1_params = fitted_model.params
OLS_model = regression.linear_model.OLS(df['R2'], df[['SPY', 'RF', 'Constant']])
fitted_model = OLS_model.fit()
print 'p-value', fitted_model.f_pvalue
print fitted_model.params
R2_params = fitted_model.params
# -
# As we've said before in other lectures, these numbers don't tell us too much by themselves. We need to look at the distribution of estimated coefficients and whether it's stable. Let's look at the rolling 100-day regression to see how it looks.
# +
model = pd.stats.ols.MovingOLS(y = df['R1'], x=df[['SPY', 'RF']],
window_type='rolling',
window=100)
rolling_parameter_estimates = model.beta
rolling_parameter_estimates.plot();
plt.hlines(R1_params['SPY'], df.index[0], df.index[-1], linestyles='dashed', colors='blue')
plt.hlines(R1_params['RF'], df.index[0], df.index[-1], linestyles='dashed', colors='green')
plt.hlines(R1_params['Constant'], df.index[0], df.index[-1], linestyles='dashed', colors='red')
plt.title('Asset1 Computed Betas');
plt.legend(['Market Beta', 'Risk Free Beta', 'Intercept', 'Market Beta Static', 'Risk Free Beta Static', 'Intercept Static']);
# +
model = pd.stats.ols.MovingOLS(y = df['R2'], x=df[['SPY', 'RF']],
window_type='rolling',
window=100)
rolling_parameter_estimates = model.beta
rolling_parameter_estimates.plot();
plt.hlines(R2_params['SPY'], df.index[0], df.index[-1], linestyles='dashed', colors='blue')
plt.hlines(R2_params['RF'], df.index[0], df.index[-1], linestyles='dashed', colors='green')
plt.hlines(R2_params['Constant'], df.index[0], df.index[-1], linestyles='dashed', colors='red')
plt.title('Asset2 Computed Betas');
plt.legend(['Market Beta', 'Risk Free Beta', 'Intercept', 'Market Beta Static', 'Risk Free Beta Static', 'Intercept Static']);
# -
# It might seem like the market betas are stable here, but let's zoom in to check.
# +
model = pd.stats.ols.MovingOLS(y = df['R2'], x=df[['SPY', 'RF']],
window_type='rolling',
window=100)
rolling_parameter_estimates = model.beta
rolling_parameter_estimates['SPY'].plot();
plt.hlines(R2_params['SPY'], df.index[0], df.index[-1], linestyles='dashed', colors='blue')
plt.title('Asset2 Computed Betas');
plt.legend(['Market Beta', 'Market Beta Static']);
# -
# As you can see, the plot scale massively affects how we perceive estimate quality.
# ##Predicting the Future
#
# Let's use this model to predict future prices for these assets.
# +
start_date = '2014-07-25'
end_date = '2015-07-25'
# We will look at the returns of an asset one-month into the future to model future returns.
offset_start_date = '2014-08-25'
offset_end_date = '2015-08-25'
# Get returns data for our assets
asset1 = get_pricing('HSC', fields='price', start_date=offset_start_date, end_date=offset_end_date).pct_change()[1:]
# Get returns for the market
bench = get_pricing('SPY', fields='price', start_date=start_date, end_date=end_date).pct_change()[1:]
# Use an ETF that tracks 3-month T-bills as our risk-free rate of return
treasury_ret = get_pricing('BIL', fields='price', start_date=start_date, end_date=end_date).pct_change()[1:]
# Define a constant to compute intercept
constant = pd.TimeSeries(np.ones(len(asset1.index)), index=asset1.index)
df = pd.DataFrame({'R1': asset1,
'SPY': bench,
'RF': treasury_ret,
'Constant': constant})
df = df.dropna()
# -
# We'll perform a historical regression to get our model parameter estimates.
# +
OLS_model = regression.linear_model.OLS(df['R1'], df[['SPY', 'RF', 'Constant']])
fitted_model = OLS_model.fit()
print 'p-value', fitted_model.f_pvalue
print fitted_model.params
b_SPY = fitted_model.params['SPY']
b_RF = fitted_model.params['RF']
a = fitted_model.params['Constant']
# -
# Get the factor data for the last month so we can predict the next month.
# +
start_date = '2015-07-25'
end_date = '2015-08-25'
# Get returns for the market
last_month_bench = get_pricing('SPY', fields='price', start_date=start_date, end_date=end_date).pct_change()[1:]
# Use an ETF that tracks 3-month T-bills as our risk-free rate of return
last_month_treasury_ret = get_pricing('BIL', fields='price', start_date=start_date, end_date=end_date).pct_change()[1:]
# -
# Make our predictions.
predictions = b_SPY * last_month_bench + b_RF * last_month_treasury_ret + a
predictions.index = predictions.index + pd.DateOffset(months=1)
plt.plot(asset1.index[-30:], asset1.values[-30:], 'b-')
plt.plot(predictions.index, predictions, 'b--')
plt.ylabel('Returns')
plt.legend(['Actual', 'Predicted']);
# Of course, this analysis hasn't yet told us anything about the quality of our predictions. To check the quality of our predictions we need to use techniques such as out of sample testing or cross-validation. For the purposes of long-short equity ranking systems, the Spearman Correlation lecture details a way to check the quality of a ranking system.
#
# ##Important Note!
#
# Again, any of these individual predictions will probably be inaccurate. Industry-quality modeling makes predictions for thousands of assets and relies on broad tends holding. If I told you that I have a predictive model with a 51% success rate, you would not make one prediction and bet all your money on it. You would make thousands of predictions and divide your money between them.
|
lectures/Arbitrage Pricing Theory.ipynb
|