code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Домашняя работа №8
# # Студент: <NAME>
import numpy as np
import matplotlib.pyplot as plt
# Есть уравнение ЗШЛ: $-\phi''(r) + l(l+1)r^{-2}\phi(r) - 2r^{-1}\phi(r) = 2E_{nl}\phi(r)$, $\phi(0) = \phi(\infty) = 0$
#
# Перепишем его, сделав замену и приведя к виду, схожему с тем, что был на лекции:
#
# $\lambda = -2E_{nl} = -2 * \frac{-1}{2 * (n +l+1)^2} = \frac{1}{(n + l + 1)^2}$
#
# $\phi''(r) - (l(l+1)r^{-2} - 2r^{-1})\phi(r) = \lambda\phi(r)$
#
# $\phi''(r) - p(r)\phi(r) = \lambda\rho(r)\phi(r)$
#
# Где $p(r) = l(l+1)r^{-2} - 2r^{-1}, \rho(r) = 1$
#
# $\phi(0) = \phi(R) = 0$
# Используйте сеточкую аапроксимацию второго порядка для решения данной ЗШЛ.
#
# (Спектр матрицы можно найти с помощью библеотечных функций)
#
# Вычислите 5 первых с.з. для $l = 0$ с точностью $\epsilon = 10^{-5}$
#
# (Начните высиления со значения R = 10)
#
# Постройте первые 5 собственных функций
# Для этого надо построить матрицу и найти ее спектр.
# +
def rho(x):
return 1
def p(x, l):
return l * (l + 1) * x ** (-2) - 2 * x ** (-1)
def get_matrix(l, R, N):
h = R / N
rhos = [rho(i * h) for i in range(1, N)]
ps = [p(i * h, l) for i in range(1, N)]
A = np.zeros((N - 1, N - 1))
for i in range(0, N - 1):
if i != 0:
A[i][i - 1] = 1 / h ** 2
A[i][i] = - (2 / h ** 2 + ps[i])
if i != N - 2:
A[i][i + 1] = 1 / h ** 2
return -A / 2
# -
# С лекции мы знаем, что сходимость сеточных методов это $O(h^2)$, значит чтобы погрешность была порядка $10^{-5}$, надо решить:
#
# $h^2 = 10^{-5} <=> h = 10^{-2.5} <=> R / N = 10^{-2.5} <=> N = 10^{2.5} * R$
#
# Если R = 10, то $N = 10^{3.5}$
#
# Поэтому возьмем $N = 4000$
# Но так как мы используем O нотацию, то она скрывает в себе константы, которые могут сильно влиять. В итоге более менее хороший ответ не достигается при R = 10, N = 4000. Если же взять при R = 100, N = 5000, то заданная погрешность в $10^{-5}$ достигается.
# +
R = 100
A = get_matrix(0, R, 5000)
eig = np.linalg.eig(A)
spectrum = eig[0]
eig_vectors = eig[1]
indexes = np.argsort(spectrum)[:5]
print("5 первых собственных значений")
print(spectrum[indexes])
print("---")
print("5 первых собственных функций")
print(eig_vectors[indexes])
# -
def draw_eig_func(data, R):
N = (len(data) + 1)
h = R / N
data_y = np.zeros(N + 1)
data_y[1:N] += data
data_x = [i * h for i in range(N + 1)]
plt.subplot(211)
plt.plot(data_x, data_y)
plt.ylabel("eigen function(N)")
plt.xlabel("N")
plt.figure(figsize=(10, 10), dpi=180)
draw_eig_func(eig_vectors[indexes][0], R)
plt.title("eigen function for lambda1 = " + str(spectrum[indexes][0]))
plt.show()
plt.figure(figsize=(10, 10), dpi=180)
draw_eig_func(eig_vectors[indexes][1], R)
plt.title("eigen function for lambda2 = " + str(spectrum[indexes][1]))
plt.show()
plt.figure(figsize=(10, 10), dpi=180)
draw_eig_func(eig_vectors[indexes][2], R)
plt.title("eigen function for lambda3 = " + str(spectrum[indexes][2]))
plt.show()
plt.figure(figsize=(10, 10), dpi=180)
draw_eig_func(eig_vectors[indexes][3], R)
plt.title("eigen function for lambda4 = " + str(spectrum[indexes][3]))
plt.show()
plt.figure(figsize=(10, 10), dpi=180)
draw_eig_func(eig_vectors[indexes][4], R)
plt.title("eigen function for lambda5 = " + str(spectrum[indexes][4]))
plt.show()
# Видимо чем меньше собственное число, тем меньше период этого "глаза", поэтому уже начиная с 4 собственного числа получается чисто синий цвет - график слишком скачет.
# +
def get_A_numerov(l, R, N):
h = R / N
rhos = [rho(i * h) for i in range(1, N)]
ps = [p(i * h, l) for i in range(1, N)]
A = np.zeros((N - 1, N - 1))
for i in range(0, N - 1):
if i != 0:
A[i][i - 1] = 1 / h ** 4 - 1 / 12 * ps[i - 1] / h ** 2
A[i][i] = - (2 / h ** 4 + ps[i] - 1 / 6 * ps[i] / h ** 2)
if i != N - 2:
A[i][i + 1] = 1 / h ** 4 - 1 / 12 * ps[i + 1] / h ** 2
return A
def get_B_numerov(l, R, N):
h = R / N
rhos = [rho(i * h) for i in range(1, N)]
B = np.zeros((N - 1, N - 1))
for i in range(0, N - 1):
if i != 0:
B[i][i - 1] = rhos[i - 1] / h ** 2 / 12
B[i][i] = rhos[i] - 1/ 6 * rhos[i] / h ** 2
if i != N - 2:
B[i][i + 1] = rhos[i + 1] / h ** 2 / 12
return B
def get_matrix_numerov(l, R, N):
return -np.matmul(np.linalg.inv(get_B_numerov(l, R, N)), get_A_numerov(l, R, N)) / 2
# +
def get_first_5(A):
return np.sort(np.linalg.eig(A)[0])[:5]
def draw_error(method, R, N_min, N_max):
real = [-1 / (2 * (n + 1) ** 2) for n in range(5)]
data_x = [i for i in range(N_min, N_max + 1)]
data_y = []
for N in range(N_min, N_max + 1):
matrix = method(0, R, N)
data_y.append(np.log10(max(abs(get_first_5(matrix) - real))))
plt.subplot(211)
plt.plot(data_x, data_y)
plt.ylabel("log10(max error)")
plt.xlabel("N")
# -
plt.figure(figsize=(10, 10), dpi=180)
R = 300
N_min = 6
N_max = 200
draw_error(get_matrix, R, N_min, N_max)
draw_error(get_matrix_numerov, R, N_min, N_max)
plt.title("Max error N")
plt.legend(("mesh", "numerov"))
plt.show()
# Для более менее адевкватной ошибки пришлось увеличить R. Хорошо видно, что mesh дает прямую, а значит ошибка действительно ведет себя как $O(h^2)$. Видим, что Нумеров справляется при больших N лучше. Что и следовало ожидать, ведь он точнее. Лучше, чем $h^4$.
|
hw08/hw08.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import numpy.testing as npt
import pandas as pd
import diff_classifier.msd as msd
import pandas.util.testing as pdt
from scipy import interpolate
import diff_classifier.features as ft
import math
frames = 10
d = {'Frame': np.linspace(0, frames, frames),
'X': np.sin(np.linspace(0, frames, frames)+3),
'Y': np.cos(np.linspace(0, frames, frames)+3),
'Track_ID': np.ones(frames)}
df = pd.DataFrame(data=d)
df = msd.all_msds2(df, frames=frames+1)
ft.msd_ratio(df, 1, 9)
d4
frames = 10
d = {'Frame': np.linspace(0, frames, frames),
'X': np.linspace(0, frames, frames)+5,
'Y': np.linspace(0, frames, frames)+3,
'Track_ID': np.ones(frames)}
df = pd.DataFrame(data=d)
df = msd.all_msds2(df, frames=frames+1)
ft.efficiency(df)
d = {'Frame': [0, 1, 2, 3, 4, 0, 1, 2, 3, 4],
'Track_ID': [1, 1, 1, 1, 1, 2, 2, 2, 2, 2],
'X': [0, 0, 1, 1, 2, 1, 1, 2, 2, 3],
'Y': [0, 1, 1, 2, 2, 0, 1, 1, 2, 2]}
df = pd.DataFrame(data=d)
dfi = msd.all_msds2(df, frames = 5)
feat = ft.calculate_features(dfi)
dfi
feat
frames = 6
d = {'Frame': np.linspace(0, frames, frames),
'X': [0, 1, 1, 2, 2, 3],
'Y': [0, 0, 1, 1, 2, 2],
'Track_ID': np.ones(frames)}
df = pd.DataFrame(data=d)
df = msd.all_msds2(df, frames=frames+1)
assert ft.aspectratio(df)[0:2] == (3.9000000000000026, 0.7435897435897438)
npt.assert_almost_equal(ft.aspectratio(df)[2], np.array([1.5, 1. ]))
ft.aspectratio(df)[2]
# +
frames = 10
d = {'Frame': np.linspace(0, frames, frames),
'X': np.linspace(0, frames, frames)+5,
'Y': np.linspace(0, frames, frames)+3,
'Track_ID': np.ones(frames)}
df = pd.DataFrame(data=d)
df = msd.all_msds2(df, frames=frames+1)
d1, d2, d3, d4, d5, d6 = ft.minBoundingRect(df)
o1, o2, o3, o4 = (-2.356194490192, 0, 14.142135623730, 0)
o5 = np.array([10, 8])
o6 = np.array([[5., 3.], [15., 13.], [15., 13.], [5., 3.]])
assert math.isclose(d1, o1, abs_tol=1e-10)
assert math.isclose(d2, o2, abs_tol=1e-10)
assert math.isclose(d3, o3, abs_tol=1e-10)
assert math.isclose(d4, o4, abs_tol=1e-10)
npt.assert_almost_equal(d5, o5)
npt.assert_almost_equal(d6, o6)
# -
d1
o1
|
notebooks/development/02_21_18_updating_test_functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %config InlineBackend.figure_formats = ['svg']
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import math
from tqdm import tqdm_notebook as tqdm
import pickle
import os
from typing import List, Dict
import seaborn as sns
sns.set_palette("dark")
# -
os.chdir("../../java")
colormap = {
"cooperative": "C0",
"truncation": "C1",
"pps": "C2",
"random_sample": "C3",
"dyadic_truncation": "C6",
"dyadic_b4": "C1",
"dyadic_b10": "C2",
"q_cooperative": "C0",
"q_truncation": "C1",
"q_pps": "C2",
"q_random_sample": "C3",
"q_dyadic_b2": "C6",
"spacesaving": "C4",
"cms_min": "C5",
"kll": "C4",
"low_discrep": "C5",
"yahoo_mg": "C7",
}
markers = {
"cooperative": "x",
"truncation": "^",
"pps": "s",
"random_sample": "+",
"dyadic_truncation": "<",
"dyadic_b4": "^",
"dyadic_b10": "+",
"q_cooperative": "x",
"q_truncation": "^",
"q_pps": "s",
"q_random_sample": "+",
"q_dyadic_b2": "<",
"spacesaving": "D",
"cms_min": "o",
"kll": "D",
"low_discrep": "o",
"yahoo_mg": "*",
}
alg_display_name = {
"cooperative": "Cooperative",
"truncation": "Truncation",
"pps": "PPS",
"random_sample": "USample",
"dyadic_truncation": "Hierarchy",
"dyadic_b4": "Hierarchy $b=4$",
"dyadic_b10": "Hierarchy $b=10$",
"q_cooperative": "Cooperative",
"q_truncation": "Truncation",
"q_pps": "PPS",
"q_random_sample": "USample",
"q_dyadic_b2": "Hierachy",
"spacesaving": "SpaceSaving",
"cms_min": "CMS",
"kll": "KLL",
"low_discrep": "LowDiscrep",
"yahoo_mg": "MG"
}
data_display_name = {
"caida_10M": "CAIDA",
"zipf1p1_10M": "Zipf",
"msft_network_10M": "Provider",
"msft_os_10M": "OSBuild",
"power_2M": "Power",
"uniform_1M": "Uniform",
"msft_records_10M": "Traffic",
}
def get_error_file(experiment_name):
return os.path.join(
"output/results/{}".format(experiment_name),
"errors.csv"
)
def query_length_plot(
experiment_name,
sketch_names: List,
item_agg="max",
query_agg="mean",
ax = None,
absolute=False,
):
e_combined = pd.read_csv(get_error_file(experiment_name))
e_combined["e_norm"] = e_combined[item_agg] / e_combined["total"]
if absolute:
e_combined["e_norm"] = e_combined[item_agg]
eg = e_combined.groupby(["sketch", "query_len"]).aggregate({
"e_norm": ["mean", "std", "max", "count"],
})
eg["err"] = eg[("e_norm", query_agg)]
eg["err_std"] = eg["e_norm", "std"] / np.sqrt(eg["e_norm", "count"])
if ax is None:
f = plt.figure(figsize=(6,4.5))
ax = f.gca()
for method in sketch_names:
eg_cur = eg.loc[method]
ax.errorbar(
eg_cur.index,
eg_cur["err"],
yerr=eg_cur["err_std"],
color=colormap[method],
markersize=5,
lw=.5,
)
ax.plot(
eg_cur.index,
eg_cur["err"],
label=alg_display_name[method],
marker=markers[method],
color=colormap[method],
markersize=5,
fillstyle="none",
lw=.5,
)
ax.set_xscale("log")
ax.set_yscale("log")
# ax.grid(axis="y", linestyle=(0, (2, 10)))
return ax, eg
def query_time_plot(
experiment_name,
sketch_names: List,
ax = None,
):
e_combined = pd.read_csv(get_error_file(experiment_name))
eg = e_combined.groupby(["sketch", "query_len"]).aggregate({
"query_time": ["mean", "std", "max", "count"],
})
eg["time"] = eg[("query_time", "mean")]
for method in sketch_names:
eg_cur = eg.loc[method]
ax.plot(
eg_cur.index,
eg_cur["time"],
label=alg_display_name[method],
marker=markers[method],
color=colormap[method],
markersize=5,
fillstyle="none",
lw=.5,
)
ax.set_xscale("log")
ax.set_yscale("log")
# ax.grid(axis="y", linestyle=(0, (2, 10)))
return ax, eg
# +
sketch_names = [
"cooperative",
"dyadic_truncation",
"random_sample",
"cms_min",
"truncation",
"yahoo_mg",
"pps"
]
sketch_size = 64
experiments = [
"l_zipf_f",
"l_caida_f",
"l_mos_f",
"l_mnetwork_f",
]
cur_experiment = experiments[0]
fig = plt.figure(figsize=(4,3), dpi=100)
ax = fig.gca()
query_length_plot(
cur_experiment,
sketch_names,
item_agg="max",
query_agg="mean",
ax=ax,
absolute=False,
)
ax.set_title(cur_experiment)
ax.set_xlabel("Query Length (Segments)")
ax.set_ylabel("Relative Error")
fig.tight_layout()
lgd = ax.legend(frameon=False, loc='upper center', bbox_to_anchor=(.5, 1.5), ncol=5)
fig.show()
# fname = "output/plots/linear_freq.pdf"
# fname = "output/{}/query_error.png".format(cur_experiment)
# fig.savefig(fname, bbox_extra_artists=(lgd,), bbox_inches='tight')
fig = plt.figure(figsize=(4,3), dpi=100)
ax = fig.gca()
query_time_plot(
cur_experiment,
sketch_names,
ax=ax,
)
ax.set_title(cur_experiment)
ax.set_xlabel("Query Length (Segments)")
ax.set_ylabel("Query Time (ms)")
fig.tight_layout()
lgd = ax.legend(frameon=False, loc='upper center', bbox_to_anchor=(.5, 1.4), ncol=4)
fig.show()
# fname = "output/{}/query_time.png".format(cur_experiment)
# fig.savefig(fname, dpi=200, bbox_extra_artists=(lgd,), bbox_inches='tight')
# +
sketch_names = [
"cooperative",
"dyadic_truncation",
# "dyadic_b3",
"random_sample",
"truncation",
"kll",
"low_discrep",
"pps"
]
sketch_size = 64
experiments = [
"l_power_q",
"l_mrecords_q",
"l_uniform_q",
]
cur_experiment = experiments[0]
fig = plt.figure(figsize=(4,3), dpi=100)
ax = fig.gca()
query_length_plot(
cur_experiment,
sketch_names,
item_agg="max",
query_agg="mean",
ax=ax,
absolute=False,
)
ax.set_title(cur_experiment)
ax.set_xlabel("Query Length (Segments)")
ax.set_ylabel("Relative Error")
fig.tight_layout()
lgd = ax.legend(frameon=False, loc='upper center', bbox_to_anchor=(.5, 1.5), ncol=5)
fig.show()
# fname = "output/plots/linear_freq.pdf"
# fig.savefig(fname, bbox_extra_artists=(lgd,), bbox_inches='tight')
fig = plt.figure(figsize=(4,3), dpi=100)
ax = fig.gca()
query_time_plot(
cur_experiment,
sketch_names,
ax=ax,
)
ax.set_title(cur_experiment)
ax.set_xlabel("Query Length (Segments)")
ax.set_ylabel("Query Time (ms)")
fig.tight_layout()
lgd = ax.legend(frameon=False, loc='upper center', bbox_to_anchor=(.5, 1.5), ncol=5)
fig.show()
# -
# # Varying Segments
enames = [
"l_caida_f",
"l_caidaseg1_f",
"l_caidaseg2_f"
]
dfs = [
pd.read_csv(get_error_file(en))
for en in enames
]
df = pd.concat(dfs).reset_index(drop=True)
df["adj_query_len"] = df["granularity"] / df["query_len"]
df_sel = df[df["adj_query_len"] == 4.0]
df_sel.groupby(["sketch", "granularity"]).aggregate({
"max": ["mean"]
})
# # Varying Size
fnames = [
"l_zipfsize_f",
"l_caidasize_f",
]
df = pd.read_csv(get_error_file(fnames[1]))
df_sel = df[df["query_len"] == 512]
df_sel.groupby(["sketch", "size"]).aggregate({
"max": ["mean"]
})
# # Accumulator
df = pd.read_csv(get_error_file("l_poweracc_q"))
df_sel = df[df["query_len"] == 512]
df_sel.groupby(["sketch", "accumulator_size"]).aggregate({
"max": ["mean"]
})
# # Cubes
df = pd.read_csv(get_error_file("c_bcube_f"))
df["err_n"] = df["max"] / df["total"]
dfg = df.groupby(["sketch"]).aggregate({"err_n": "mean"})
dfg
df = pd.read_csv(get_error_file("c_bcube_lesion_f"))
df["err_n"] = df["max"] / df["total"]
dfg = df.groupby(["sketch", "workload_query_prob"]).aggregate({"err_n": "mean"})
dfg
# # Cube Runtime
df = pd.read_csv(get_error_file("c_mrecords_q"))
dfg = df.groupby(["sketch", "query_len"]).aggregate({"query_time": "mean"})
dfg
|
python/notebooks/RemoteExperiment.ipynb
|
'''
Example implementations of HARK.ConsumptionSaving.ConsPortfolioModel
'''
from HARK.ConsumptionSaving.ConsPortfolioModel import PortfolioConsumerType, init_portfolio
from HARK.ConsumptionSaving.ConsIndShockModel import init_lifecycle
from HARK.utilities import plot_funcs
from copy import copy
from time import time
import numpy as np
import matplotlib.pyplot as plt
# Make and solve an example portfolio choice consumer type
print('Now solving an example portfolio choice problem; this might take a moment...')
MyType = PortfolioConsumerType()
MyType.cycles = 0
t0 = time()
MyType.solve()
t1 = time()
MyType.cFunc = [MyType.solution[t].cFuncAdj for t in range(MyType.T_cycle)]
MyType.ShareFunc = [MyType.solution[t].ShareFuncAdj for t in range(MyType.T_cycle)]
print('Solving an infinite horizon portfolio choice problem took ' + str(t1-t0) + ' seconds.')
# Plot the consumption and risky-share functions
print('Consumption function over market resources:')
plot_funcs(MyType.cFunc[0], 0., 20.)
print('Risky asset share as a function of market resources:')
print('Optimal (blue) versus Theoretical Limit (orange)')
plt.xlabel('Normalized Market Resources')
plt.ylabel('Portfolio Share')
plt.ylim(0.0,1.0)
# Since we are using a discretization of the lognormal distribution,
# the limit is numerically computed and slightly different from
# the analytical limit obtained by Merton and Samuelson for infinite wealth
plot_funcs([MyType.ShareFunc[0]
# ,lambda m: RiskyShareMertSamLogNormal(MyType.RiskPrem,MyType.CRRA,MyType.RiskyVar)*np.ones_like(m)
,lambda m: MyType.ShareLimit*np.ones_like(m)
] , 0., 200.)
# Now simulate this consumer type
MyType.track_vars = ['cNrm', 'Share', 'aNrm', 't_age']
MyType.T_sim = 100
MyType.initialize_sim()
MyType.simulate()
print('\n\n\n')
print('For derivation of the numerical limiting portfolio share')
print('as market resources approach infinity, see')
print('http://www.econ2.jhu.edu/people/ccarroll/public/lecturenotes/AssetPricing/Portfolio-CRRA/')
""
# Make another example type, but this one optimizes risky portfolio share only
# on the discrete grid of values implicitly chosen by RiskyCount, using explicit
# value maximization.
init_discrete_share = init_portfolio.copy()
init_discrete_share['DiscreteShareBool'] = True
init_discrete_share['vFuncBool'] = True # Have to actually construct value function for this to work
# Make and solve a discrete portfolio choice consumer type
print('Now solving a discrete choice portfolio problem; this might take a minute...')
DiscreteType = PortfolioConsumerType(**init_discrete_share)
DiscreteType.cycles = 0
t0 = time()
DiscreteType.solve()
t1 = time()
DiscreteType.cFunc = [DiscreteType.solution[t].cFuncAdj for t in range(DiscreteType.T_cycle)]
DiscreteType.ShareFunc = [DiscreteType.solution[t].ShareFuncAdj for t in range(DiscreteType.T_cycle)]
print('Solving an infinite horizon discrete portfolio choice problem took ' + str(t1-t0) + ' seconds.')
# Plot the consumption and risky-share functions
print('Consumption function over market resources:')
plot_funcs(DiscreteType.cFunc[0], 0., 50.)
print('Risky asset share as a function of market resources:')
print('Optimal (blue) versus Theoretical Limit (orange)')
plt.xlabel('Normalized Market Resources')
plt.ylabel('Portfolio Share')
plt.ylim(0.0,1.0)
# Since we are using a discretization of the lognormal distribution,
# the limit is numerically computed and slightly different from
# the analytical limit obtained by Merton and Samuelson for infinite wealth
plot_funcs([DiscreteType.ShareFunc[0]
,lambda m: DiscreteType.ShareLimit*np.ones_like(m)
] , 0., 200.)
print('\n\n\n')
""
# Make another example type, but this one can only update their risky portfolio
# share in any particular period with 15% probability.
init_sticky_share = init_portfolio.copy()
init_sticky_share['AdjustPrb'] = 0.15
# Make and solve a discrete portfolio choice consumer type
print('Now solving a portfolio choice problem with "sticky" portfolio shares; this might take a moment...')
StickyType = PortfolioConsumerType(**init_sticky_share)
StickyType.cycles = 0
t0 = time()
StickyType.solve()
t1 = time()
StickyType.cFuncAdj = [StickyType.solution[t].cFuncAdj for t in range(StickyType.T_cycle)]
StickyType.cFuncFxd = [StickyType.solution[t].cFuncFxd for t in range(StickyType.T_cycle)]
StickyType.ShareFunc = [StickyType.solution[t].ShareFuncAdj for t in range(StickyType.T_cycle)]
print('Solving an infinite horizon sticky portfolio choice problem took ' + str(t1-t0) + ' seconds.')
# Plot the consumption and risky-share functions
print('Consumption function over market resources when the agent can adjust his portfolio:')
plot_funcs(StickyType.cFuncAdj[0], 0., 50.)
print("Consumption function over market resources when the agent CAN'T adjust, by current share:")
M = np.linspace(0., 50., 200)
for s in np.linspace(0.,1.,21):
C = StickyType.cFuncFxd[0](M, s*np.ones_like(M))
plt.plot(M,C)
plt.xlim(0.,50.)
plt.ylim(0.,None)
plt.show()
print('Risky asset share function over market resources (when possible to adjust):')
print('Optimal (blue) versus Theoretical Limit (orange)')
plt.xlabel('Normalized Market Resources')
plt.ylabel('Portfolio Share')
plt.ylim(0.0,1.0)
plot_funcs([StickyType.ShareFunc[0]
,lambda m: StickyType.ShareLimit*np.ones_like(m)
] , 0., 200.)
""
# Make another example type, but this one has *age-varying* perceptions of risky asset returns.
# Begin by making a lifecycle dictionary, but adjusted for the portfolio choice model.
init_age_varying_risk_perceptions = copy(init_lifecycle)
init_age_varying_risk_perceptions['RiskyCount'] = init_portfolio['RiskyCount']
init_age_varying_risk_perceptions['ShareCount'] = init_portfolio['ShareCount']
init_age_varying_risk_perceptions['aXtraMax'] = init_portfolio['aXtraMax']
init_age_varying_risk_perceptions['aXtraCount'] = init_portfolio['aXtraCount']
init_age_varying_risk_perceptions['aXtraNestFac'] = init_portfolio['aXtraNestFac']
init_age_varying_risk_perceptions['BoroCnstArt'] = init_portfolio['BoroCnstArt']
init_age_varying_risk_perceptions['CRRA'] = init_portfolio['CRRA']
init_age_varying_risk_perceptions['DiscFac'] = init_portfolio['DiscFac']
init_age_varying_risk_perceptions['RiskyAvg'] = [1.08]*init_lifecycle['T_cycle']
init_age_varying_risk_perceptions['RiskyStd'] = list(np.linspace(0.20,0.30,init_lifecycle['T_cycle']))
init_age_varying_risk_perceptions['RiskyAvgTrue'] = 1.08
init_age_varying_risk_perceptions['RiskyStdTrue'] = 0.20
AgeVaryingRiskPercType = PortfolioConsumerType(**init_age_varying_risk_perceptions)
AgeVaryingRiskPercType.cycles = 1
# Solve the agent type with age-varying risk perceptions
#print('Now solving a portfolio choice problem with age-varying risk perceptions...')
t0 = time()
AgeVaryingRiskPercType.solve()
AgeVaryingRiskPercType.cFunc = [AgeVaryingRiskPercType.solution[t].cFuncAdj for t in range(AgeVaryingRiskPercType.T_cycle)]
AgeVaryingRiskPercType.ShareFunc = [AgeVaryingRiskPercType.solution[t].ShareFuncAdj for t in range(AgeVaryingRiskPercType.T_cycle)]
t1 = time()
print('Solving a ' + str(AgeVaryingRiskPercType.T_cycle) + ' period portfolio choice problem with age-varying risk perceptions took ' + str(t1-t0) + ' seconds.')
# Plot the consumption and risky-share functions
print('Consumption function over market resources in each lifecycle period:')
plot_funcs(AgeVaryingRiskPercType.cFunc, 0., 20.)
print('Risky asset share function over market resources in each lifecycle period:')
plot_funcs(AgeVaryingRiskPercType.ShareFunc, 0., 200.)
# The code below tests the mathematical limits of the model.
# +
# Create a grid of market resources for the plots
mMin = 0 # Minimum ratio of assets to income to plot
mMax = 5*1e2 # Maximum ratio of assets to income to plot
mPts = 1000 # Number of points to plot
eevalgrid = np.linspace(0,mMax,mPts) # range of values of assets for the plot
# Number of points that will be used to approximate the risky distribution
risky_count_grid = [5,200]
# Plot by ages (time periods) at which to plot. We will use the default life-cycle calibration.
ages = [2, 4, 6, 8]
#Create a function to compute the Merton-Samuelson limiting portfolio share.
def RiskyShareMertSamLogNormal(RiskPrem,CRRA,RiskyVar):
return RiskPrem/(CRRA*RiskyVar)
# + Calibration and solution
for rcount in risky_count_grid:
# Create a new dictionary and replace the number of points that
# approximate the risky return distribution
# Create new dictionary copying the default
merton_dict = init_lifecycle.copy()
merton_dict['RiskyCount'] = rcount
# Create and solve agent
agent = PortfolioConsumerType(**merton_dict)
agent.solve()
# Compute the analytical Merton-Samuelson limiting portfolio share
RiskyVar = agent.RiskyStd**2
RiskPrem = agent.RiskyAvg - agent.Rfree
MS_limit = RiskyShareMertSamLogNormal(RiskPrem,
agent.CRRA,
RiskyVar)
# Now compute the limiting share numerically, using the approximated
# distribution
agent.update_ShareLimit()
NU_limit = agent.ShareLimit
plt.figure()
for a in ages:
plt.plot(eevalgrid,
agent.solution[a]\
.ShareFuncAdj(eevalgrid),
label = 't = %i' %(a))
plt.axhline(NU_limit, c='k', ls='-.', label = 'Exact limit as $m\\rightarrow \\infty$.')
plt.axhline(MS_limit, c='k', ls='--', label = 'M&S Limit without returns discretization.')
plt.ylim(0,1.05)
plt.xlim(eevalgrid[0],eevalgrid[-1])
plt.legend()
plt.title('Risky Portfolio Share by Age\n Risky distribution with {points} equiprobable points'.format(points = rcount))
plt.xlabel('Wealth (m)')
plt.ioff()
plt.draw()
# -
|
examples/ConsumptionSaving/example_ConsPortfolioModel.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,.md//md
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Flow Chart Generator
from jp_flowchartjs.jp_flowchartjs import FlowchartWidget
# *An [issue](https://github.com/adrai/flowchart.js/issues/186) has been filed regarding the misaligned return path in the flow diagram in `jp_flowchartjs`.*
# ## 03_1_1
#
# Mermaid code https://mermaid-js.github.io/mermaid-live-editor/
# The mermaid editor is supposed to encode/decode base 64 strings
# but I can't get my own encoder/decoder to work to encode/decode things the same way?
#
# graph LR
# A(Start) --> B[Set the count to 0]
# B --> C[Swim a length]
# C --> D[Add 1 to the count]
# D --> E{Is the<br/>count less<br/>than 20?}
# E --> |Yes| C
# E --> |No| F(End)
#
# Long description:
#
# A flow chart for a person swimming 20 lengths of a pool. The flow chart starts with an oval shape labelled ‘Start’. From here there are sequences of boxes connected by arrows: first ‘Set the count to 0’, then ‘Swim a length’ and last ‘Add 1 to the count’. From this box an arrow leads to a decision diamond labelled ‘Is the count less than 20?’ Two arrows lead from this: one labelled ‘Yes’, the other ‘No’. The ‘Yes’ branch leads back to rejoin the box ‘Swim a length’. The ‘No’ branch leads directly to an oval shape labelled ‘End’. There is thus a loop in the chart which includes the steps ‘Swim a length’ and ‘Add 1 to the count’ and ends with the decision ‘Is the count less than 20?’
# %%flowchart_magic
st=>start: Start
e=>end: End
op1=>operation: Set the count to 0
op2=>operation: Swim a length
op3=>operation: Add 1 to the count
cond=>condition: Is the count less than 20?
st(right)->op1(right)->op2(right)->op3(right)->cond
cond(yes,right)->op2
cond(no)->e
# ### Ironing
#
# Mermaid drawing (this is far from ideal...Maybe better to use draw.io?)<font color='red'>JD: To be resolved.</font>
#
# graph LR
# A(Start) --> B{Any clothes<br>left in<br/>basket?}
# B --> |Yes| C[Take out<br/>item and<br/>iron]
# C --> D[Put it on pile<br/>of ironed<br/>clothes]
# D --> B
# B --> |No| E(End)
#
# Long description:
#
# A flow chart for a person ironing clothes. This starts with an oval ‘Start’. An arrow leads to a decision diamond ‘Any clothes left in basket?’ Two arrows lead from this: one labelled ‘Yes’, the other ‘No’. The ‘Yes’ branch continues in turn to two boxes ‘Take out item and iron’ and ‘Put it on pile of ironed clothes’; an arrow leads back from this box to rejoin the decision ‘Any clothes left in basket?’ The ‘No’ branch leads directly to an oval ‘End’. There is thus a loop in the chart which begins with the decision ‘Any clothes left in basket?’ and includes the steps ‘Take out item and iron’ and ‘Put it on pile of ironed clothes’.
# %%flowchart_magic
st=>start: Start
e=>end: End
cond=>condition: Any clothes left in basket?
op2=>operation: Take out item and iron
op3=>operation: Put it on pile of ironed clothes
st(right)->cond
cond(yes, right)->op2
op2(right)->op3(top)->cond
cond(no, bottom)->e
# graph TD
# A(Start) --> B[Set counter to 0]
# B --> C[# Draw side<br/>...code...]
# C --> D[# Turn ninety degrees<br/>...code...]
# D --> E[Add 1 to counter]
# E --> F{Is counter < 4?}
# F --> |Yes| C
# F --> |No| G(End)
#
# Long description:
#
# A flow chart for a robot program with a loop. This starts with an oval ‘Start’. An arrow leads first to a box ‘Set counter to 0’ and then to a sequence of further boxes: ‘# Draw side’ with an implication of the code associated with that, ‘# Turn ninety degrees’, again with a hint regarding the presence of code associated with that activity, and lastly ‘Add 1 to counter’. The arrow from this box leads to a decision diamond ‘Is counter < 4?’ Two arrows lead from this, one labelled ‘Yes’, the other ‘No’. The ‘Yes’ branch loops back to rejoin the sequence at ‘# Draw side code’. The ‘No’ branch leads directly to an oval ‘End’. There is thus a loop in the chart which includes the sequence of motor control commands, incrementing the counter and ends with the decision ‘Is counter < 4?’.
# %%flowchart_magic
st=>start: Start
e=>end: End
op1=>operation: Set counter to 0
op2=>operation: Draw side code
op3=>operation: Turn ninety degrees code
op4=>operation: Add 1 to counter
cond=>condition: Is counter < 4?
st(right)->op1(right)->op2->op3->cond
cond(yes, right)->op2
cond(no, bottom)->e
# ### 02_Robot_Lab/Section_00_02.md
#
# 
# +
# %%flowchart_magic
st=>start: Start
e=>end: End
op1=>operation: Start moving forwards
cond1=>condition: Light < 100?
op2=>parallel: Display reading (continue drive forwards)
op3=>operation: Drive forward a short way
cond2=>condition: Light < 50?
op4=>operation: Say "grey"
op5=>operation: Say "black"
st(right)->op1(right)->cond1({'flowstate':{'cond1':{'yes-text' : 'ssno', 'no-text' : 'yes'}}} )
cond1(no,right)->op2(path1, top)->cond1
cond1(yes)->op3->cond2
cond2(no)->op4->e
cond2(yes)->op5->e
# -
# <div class='alert alert-warning'>The flowchart.js layout algorithm will allow us to change yes/no labels on conditions, but the magic does not currently provide a way to pass alternative values in (eg swapping yes/no labels to change the sense of the decision as drawn). Also, the layout algorithm seems to be broken when swapping yes and no.</div>
# +
## # Mermaid.js code
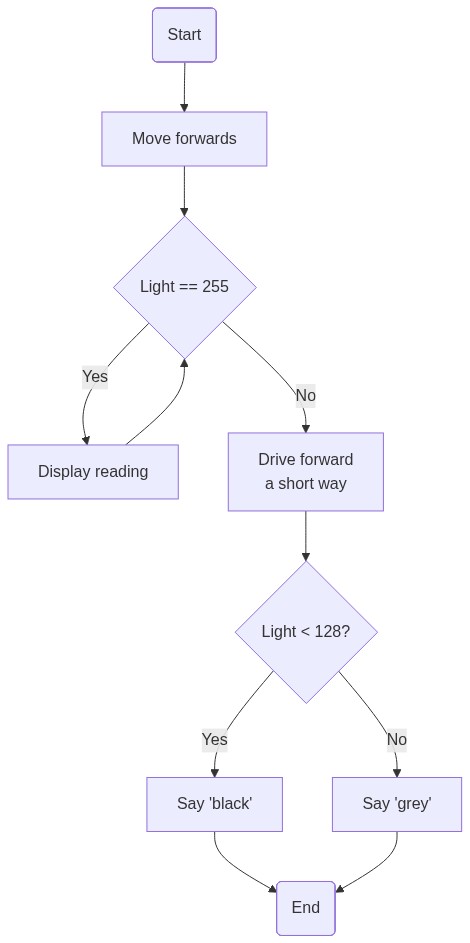
graph TD
A(Start) --> B[Move forwards]
B --> C{Light == 100}
C --> |Yes| D[Display reading]
D --> C
C --> |No| E[Drive forward<br/>a short way]
E --> F{Light < 50?}
F --> |Yes| G[Say 'black']
F --> |No| H[Say 'grey']
G --> I(End)
H --> I
# -
# We can do checklists in notebooks:
#
# - [ ] item 1
# - [ ] item 2
# +
# # %%flowchart_magic
st=>start: Start
e=>end: End
op1=>operation: Start moving forwards
cond1=>condition: Light < 100?
op2=>parallel: Display reading (continue drive forwards)
op3=>operation: Drive forward a short way
cond2=>condition: Light < 50?
op4=>operation: Say "grey"
op5=>operation: Say "black"
st(right)->op1(right)->cond1({'flowstate':{'cond1':{'yes-text' : 'ssno', 'no-text' : 'yes'}}} )
cond1(no,right)->op2(path1, top)->cond1
cond1(yes)->op3->cond2
cond2(no)->op4->e
cond2(yes)->op5->e
# -
# The flow chart starts with a *Start* label followed by a rectangular *Start moving forwards* step. An initial diamond shaped decision block tests the condition "Light < 100". If false ("no"), the program continues with a rectangular "Display reading (continue drive forwards)" step which then returns control back to the first decision block. If true ("yes") control progresses via a rectangular "Drive forward a short way" step and then another diamond shaped decision step which tests the condition "Light < 50". If "no", control passes to a rectangular "Say 'grey'" step and then a terminating "End" condition. If "yes", control passes to a rectangular "Say 'black'" step and then also connects to the terminating "End" condition.
# ## Design cycle
# + active=""
# # st=>start: Start
# e=>end: End
# op1=>operation: Generate
# op2=>parallel: Evaluate
# st(right)->op1(right)->op2
# op2(path1, top)->op1
# op2(path2, right)->e
|
content/backgrounds/Flowchart Generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn import datasets
from sklearn import metrics
from sklearn.datasets import load_iris
iris = load_iris()
x = iris.data # independent features
y = iris.target # dependent features
#print(x)
#print(y)
y
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split (x, y, test_size = 0.20,random_state = 0)
# +
#GaussianNB is specifically used when the features have continuous values.
from sklearn.naive_bayes import GaussianNB
model = GaussianNB()
model.fit(X_train, y_train)
prediction = model.predict(X_test)
print(prediction)
print(y_test)
# -
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test,prediction)
TPN=11+13+5
Total=30
accuracy =TPN/Total
accuracy
# +
from sklearn.metrics import accuracy_score
print(accuracy_score(y_test, prediction))
# +
from sklearn.metrics import f1_score
print(f1_score(y_test, prediction))
# +
print(metrics.confusion_matrix(y_test, prediction))
prediction
|
Lec 15 Naive Bayes Questions/Ma'am/gaussian.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A software bug
# ## Problem Definition
# You are using a legacy software to solve a production mix Continuous Linear Programming (CLP) problem in your company where the objective is to maximise profits. The following table provides the solution of the primal, dual and a sensitivity analysis for the three decision variables that represent the quantities to produce of each product
#
# |Variables | Solution | Reduced cost | Objective Coefficient | Lower bound | Upper bound|
# |---|---|---|---|---|---|
# |$x_1$ | 300.00 | ? | 30.00 | 24.44 | Inf|
# |$x_2$ | 33.33 | ? | 20.00 | -0.00 | 90.00|
# |$x_3$ | 0 | -8.33 | 40.00 | -Inf | 48.33 |
#
# Answer the following questions:
#
# **a.** Notice that there are some values missing (a bug in the software shows a ? sign instead of a numerical value, remember to use Python the next time around). Fill the missing values and explain your decision
# The reduced costs of $x_1$ and $x_2$ must be zero, since the two variables are basic, the reduced cost is zero.
#
# **b.** According to the provided solution, which of the three products has the highest impact in your objective function? Motivate your response
# The product of the objective coefficient times the solution determines the impact in the objective function. In this case, the variable that has the highest impact in the objective function is $x_1$
# **c.** What does the model tell you about product $x_3$. Is it profitable to manufacture under the current conditions modeled in the problem? Provide quantitative values to motivate your response
# In this solution, it is not profitable to manufacture $x_3$. In fact, manufacturing 1 unit would have a negative impact of 8.33 in the objective function.
#
# **d.** Recall that in this type of problem, the objective coefficient represents the marginal profit per unit of product. What would happen if the objective function of variable $x_3$ is increased over 50? Describe what would be the impact in the obtained solution (1 point)
# The upper bound of the objective coefficient is 48.33, meaning that if the objective coefficient goes over this value, we would experience a change in the FSB. Most likely, variable The objective coefficient would need to increase up to 48.33 to make this product profitable $x_3$ would enter the solution.
#
#
#
# The following table represents the value obtained for the decision variables related to the constraints:
#
# | Constraint | Right Hand Side | Shadow Price | Slack | Min RHS | Max RHS |
# |-----|-----|-----|----|-----|----|
# | $s_1$ | 400.00 | 6.67 | 0.00 | 300.00 | 525.00 |
# | $s_2$ | 600.00 | 11.67 | 0.00 | 0.00 | 800.00 |
# | $s_3$ | 600.00 | 0.00 | 166.67 | 433.33 | Inf |
#
# Bearing in mind that the constraints represent the availability of 3 limited resources (operating time in minutes) and that the type of constraints is in every case "less or equal" answer the following questions:
#
# **e.** Which decision variables are basic? How many are there and, is this result expected? Motivate your response
# The basic decision variables are $x_1$, $x_2$, and $s_3$, as many basic decision variables as constraints, since this is a Feasible Basic Solution (FBS).
#
# **f.** Imagine that you had to cut down costs by reducing the availability of the limited resources in the model. Which one would you select? How much could you cut down without changing the production mix? Motivate your response
# If the Right Hand Sides represent availability of resources, we could reduce the availability of the third constraint in 166.7 units (slack) without any change in the base.
#
# **g.** Now, with the money saved, you could invest to increase the availability of other limited resources. Again, indicate which one would you select and by how much you would increase the availability of this resource without changing the production mix. Motivate your response
# The best option is to invest in the resource of the second constraint, since it has the largest shadow price. The RHS can increase up to 800 units without a change in the base.
#
# **h.** How do the columns of the two tables relate to the primal and dual problem? Moreover, for each column, not considering upper and lower bounds, write down a brief description of each column and the relationship it has with the primal and dual problem
# In the first table, the column solution contains the values of the decision variables of the primal and the column reduced costs represents the slack variables of the dual.
# In the second table, the column slack represents the values of the slack variables in the primal solution and the column shadow prices, represent the values of the decision variables of the dual.
# + [markdown] pycharm={"name": "#%% md\n"}
#
|
docs/source/CLP/solved/A software bug (Solved).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ml] *
# language: python
# name: conda-env-ml-py
# ---
# # Early Stopping
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import sys
sys.path.insert(0, "../src")
# +
import numpy as np
import pandas as pd
from sklearn import metrics
from sklearn import model_selection
import albumentations as A
import torch
import callbacks
import config
import dataset
import models
import engine
# +
import numpy as np
import torch
class EarlyStopping:
def __init__(self, patience=7, mode="max", delta=0.0001):
self.patience = patience
self.mode = mode
self.delta = delta
self.best_score = None
self.counter = 0
self.early_stop = False
if mode == "max":
self.val_score = -np.inf
elif mode == "min":
self.val_score = np.inf
def __call__(self, epoch_score, model, model_path):
if self.mode == "max":
score = np.copy(epoch_score)
elif self.mode == "min":
score = -1.0 * epoch_score
if self.best_score is None:
self.best_score = score
self.save_checkpoint(epoch_score, model, model_path)
elif score < self.best_score + self.delta:
self.counter += 1
print(f"EarlyStopping counter: {self.counter} out of {self.patience}")
if self.counter >= self.patience:
self.early_stop = True
else:
self.best_score = score
self.counter = 0
self.save_checkpoint(epoch_score, model, model_path)
def save_checkpoint(self, epoch_score, model, model_path):
if epoch_score not in (np.inf, -np.inf, -np.nan, np.nan):
print(f"Validation score improved ({self.val_score} --> {epoch_score}). Saving model!")
torch.save(model.state_dict(), model_path)
self.val_score = epoch_score
# -
df = pd.read_csv(config.TRAIN_CSV)
# df_train, df_valid = model_selection.train_test_split(df, test_size=0.1, stratify=df.digit)
train_idx, valid_idx = model_selection.train_test_split(np.arange(len(df)), test_size=0.1, stratify=df.digit)
train_dataset = dataset.EMNISTDataset(df, train_idx)
valid_dataset = dataset.EMNISTDataset(df, valid_idx)
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=config.TRAIN_BATCH_SIZE, shuffle=True)
valid_loader = torch.utils.data.DataLoader(valid_dataset, batch_size=config.TEST_BATCH_SIZE)
# +
EPOCHS = 200
device = torch.device(config.DEVICE)
model = models.SpinalVGG()
# model = models.Model()
model.to(device)
optimizer = torch.optim.Adam(model.parameters())
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(
optimizer, mode='max', verbose=True, patience=10, factor=0.5
)
early_stop = callbacks.EarlyStopping(patience=15, mode="max")
for epoch in range(EPOCHS):
engine.train(train_loader, model, optimizer, device)
predictions, targets = engine.evaluate(valid_loader, model, device)
predictions = np.array(predictions)
predictions = np.argmax(predictions, axis=1)
accuracy = metrics.accuracy_score(targets, predictions)
print(f"Epoch: {epoch}, Valid accuracy={accuracy}")
model_path = "./test.pt"
early_stop(accuracy, model, model_path)
if early_stop.early_stop:
print(f"Early stopping. Best score {early_stop.best_score}. Loading weights...")
model.load_state_dict(torch.load(model_path))
break
scheduler.step(accuracy)
# -
model.load_state_dict(torch.load("./test.pt"))
torch.save(model.state_dict(), "../models/spinalvgg.pt")
model = models.SpinalVGG()
model.load_state_dict(torch.load("../models/spinalvgg.pt"))
model.to(device)
df_test = pd.read_csv(config.TEST_CSV)
test_dataset = dataset.EMNISTTestDataset(df_test)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=config.TEST_BATCH_SIZE)
predictions = engine.infer(test_loader, model, device)
predictions = np.array(predictions)
predictions = np.argmax(predictions, axis=1)
submission = pd.DataFrame({"id": df_test.id, "digit": predictions})
submission.to_csv("../output/spinalvgg.csv", index=False)
submission.head()
|
notebooks/04-early-stopping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
from collections import Counter
import dill
import glob
import igraph as ig
import itertools
import leidenalg
#import magic
import matplotlib
from matplotlib import pyplot
import numba
import numpy
import os
import pickle
from plumbum import local
import random
import re
import scipy
from scipy.cluster import hierarchy
import scipy.sparse as sps
from scipy.spatial import distance
import scipy.stats as stats
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.decomposition import TruncatedSVD
from sklearn import neighbors
from sklearn import metrics
import sys
import umap
#from plotly import tools
#import plotly.offline as py
#import plotly.graph_objs as go
#py.init_notebook_mode(connected=True)
# +
def find_nearest_genes(peak_files, out_subdir, refseq_exon_bed):
#get unix utilities
bedtools, sort, cut, uniq, awk = local['bedtools'], local['sort'], local['cut'], local['uniq'], local['awk']
#process the peak files to find nearest genes
nearest_genes = []
for path in sorted(peak_files):
out_path = os.path.join(out_subdir, os.path.basename(path).replace('.bed', '.nearest_genes.txt'))
cmd = (bedtools['closest', '-D', 'b', '-io', '-id', '-a', path, '-b', refseq_exon_bed] |
cut['-f1,2,3,5,9,12'] | #fields are chrom, start, stop, peak sum, gene name, distance
awk['BEGIN{OFS="\t"}{if($6 > -1200){print($1, $2, $3, $6, $5, $4);}}'] |
sort['-k5,5', '-k6,6nr'] |
cut['-f5,6'])()
with open(out_path, 'w') as out:
prev_gene = None
for idx, line in enumerate(str(cmd).strip().split('\n')):
if prev_gene is None or not line.startswith(prev_gene):
# print(line)
line_split = line.strip().split()
prev_gene = line_split[0]
out.write(line + '\n')
nearest_genes.append(out_path)
return nearest_genes
def load_expr_db(db_path):
if os.path.basename(db_path) == 'RepAvgGeneTPM.csv':
with open(db_path) as lines_in:
db_headers = lines_in.readline().strip().split(',')[1:]
db_vals = numpy.loadtxt(db_path, delimiter=',', skiprows=1, dtype=object)[:,1:]
else:
with open(db_path) as lines_in:
db_headers = lines_in.readline().strip().split('\t')
db_vals = numpy.loadtxt(db_path, delimiter='\t', skiprows=1, dtype=object)
print('Loaded DB shape: {!s}'.format(db_vals.shape))
return (db_headers, db_vals)
TOPN=500
def get_gene_data(genes_path, gene_expr_db, topn=TOPN):
if isinstance(genes_path, list):
genes_list = genes_path
else:
with open(genes_path) as lines_in:
genes_list = [elt.strip().split()[:2] for elt in lines_in]
gene_idx = [(numpy.where(gene_expr_db[:,0] == elt[0])[0],elt[1]) for elt in genes_list]
gene_idx_sorted = sorted(gene_idx, key=lambda x:float(x[1]), reverse=True)
gene_idx, gene_weights = zip(*[elt for elt in gene_idx_sorted if len(elt[0]) > 0][:topn])
gene_idx = [elt[0] for elt in gene_idx]
gene_data = gene_expr_db[:,1:].astype(float)[gene_idx,:]
denom = numpy.sum(gene_data, axis=1)[:,None] + 1e-8
gene_norm = gene_data/denom
return gene_idx, gene_data, gene_norm, len(genes_list), numpy.array(gene_weights, dtype=float)
def sample_db(data_norm, expr_db, data_weights=None, nsamples=1000):
samples = []
rs = numpy.random.RandomState(15321)
random_subset = numpy.arange(expr_db.shape[0])
num_to_select = data_norm.shape[0]
for idx in range(nsamples):
rs.shuffle(random_subset)
db_subset = expr_db[random_subset[:num_to_select]][:,1:].astype(float)
denom = numpy.sum(db_subset, axis=1)[:None] + 1e-8
db_subset_norm = numpy.mean((db_subset.T/denom).T, axis=0)
if data_weights is not None:
samples.append(numpy.log2(numpy.average(data_norm, axis=0, weights=gene_weights)/db_subset_norm))
else:
samples.append(numpy.log2(numpy.average(data_norm, axis=0, weights=None)/db_subset_norm))
samples = numpy.vstack(samples)
samples_mean = numpy.mean(samples, axis=0)
samples_sem = stats.sem(samples, axis=0)
conf_int = numpy.array([stats.t.interval(0.95, samples.shape[0]-1,
loc=samples_mean[idx], scale=samples_sem[idx])
for idx in range(samples.shape[1])]).T
conf_int[0] = samples_mean - conf_int[0]
conf_int[1] = conf_int[1] - samples_mean
return samples_mean, conf_int
def plot_l2_tissues(nearest_genes_glob, refdata, expr_db=None, expr_db_headers=None, ncols=3,
topn=TOPN, weights=False, nsamples=100, savefile=None, display_in_notebook=True):
if expr_db is None:
#Get all L2 tissue expression data to normalize the distribution of genes from peaks
l2_tissue_db_path = os.path.join(refdata,'gexplore_l2_tissue_expr.txt')
expr_db_headers, expr_db = load_expr_db(l2_tissue_db_path)
gene_lists = glob.glob(nearest_genes_glob)
if os.path.basename(gene_lists[0]).startswith('peaks'):
gene_lists.sort(key=lambda x:int(os.path.basename(x).split('.')[0].replace('peaks', '')))
elif os.path.basename(gene_lists[0]).startswith('topic'):
gene_lists.sort(key=lambda x:int(os.path.basename(x).split('.')[1].replace('rank', '')))
else:
gene_lists.sort(key=lambda x:os.path.basename(x).split('.')[0])
gene_list_data = [(os.path.basename(path).split('.')[0], get_gene_data(path, expr_db, topn=topn)) for path in gene_lists]
print('\n'.join(['{!s} nearest genes: found {!s} out of {!s} total'.format(fname, data.shape[0], gene_list_len)
for (fname, (data_idx, data, data_norm, gene_list_len, gene_weights)) in gene_list_data]))
l2_tissue_colors = [('Body wall muscle', '#e51a1e'),
('Intestinal/rectal muscle', '#e51a1e'),
('Pharyngeal muscle', '#377db8'),
('Pharyngeal epithelia', '#377db8'),
('Pharyngeal gland', '#377db8'),
('Seam cells', '#4eae4a'),
('Non-seam hypodermis', '#4eae4a'),
('Rectum', '#4eae4a'),
('Ciliated sensory neurons', '#984ea3'),
('Oxygen sensory neurons', '#984ea3'),
('Touch receptor neurons', '#984ea3'),
('Cholinergic neurons', '#984ea3'),
('GABAergic neurons', '#984ea3'),
('Pharyngeal neurons', '#984ea3'),
('flp-1(+) interneurons', '#984ea3'),
('Other interneurons', '#984ea3'),
('Canal associated neurons', '#984ea3'),
('Am/PH sheath cells', '#ff8000'),
('Socket cells', '#ff8000'),
('Excretory cells', '#ff8000'),
('Intestine', '#fcd800'),
('Germline', '#f97fc0'),
('Somatic gonad precursors', '#f97fc0'),
('Distal tip cells', '#f97fc0'),
('Vulval precursors', '#f97fc0'),
('Sex myoblasts', '#f97fc0'),
('Coelomocytes', '#a75629')]
idx_by_color = {}
for idx, (name, color) in enumerate(l2_tissue_colors):
try:
idx_by_color[color][1].append(idx)
except KeyError:
idx_by_color[color] = [name, [idx]]
# rs = numpy.random.RandomState(15321)
# random_subset = numpy.arange(expr_db.shape[0])
# rs.shuffle(random_subset)
# #num_to_select = int(numpy.mean([neuron_data.shape[0], emb_muscle_data.shape[0], l2_muscle_data.shape[0]]))
# num_to_select = len(random_subset)
# l2_tissue_db_subset = expr_db[random_subset[:num_to_select]][:,1:].astype(float)
# denom = numpy.sum(l2_tissue_db_subset, axis=1)[:,None] + 1e-8
# l2_tissue_db_norm = numpy.mean(l2_tissue_db_subset/denom, axis=0)
print('Tissue DB norm shape: {!s}'.format(expr_db.shape))
pyplot.rcParams.update({'xtick.labelsize':14,
'ytick.labelsize':14,
'xtick.major.pad':8})
ind = numpy.arange(len(expr_db_headers) - 1)
width = 0.66
axis_fontsize = 18
title_fontsize = 19
nrows = int(numpy.ceil(len(gene_list_data)/float(ncols)))
fig, axes = pyplot.subplots(nrows=nrows, ncols=ncols, figsize=(7 * ncols, 7 * nrows), sharey=True)
for idx, (fname, (data_idx, data, data_norm, gene_list_len, gene_weights)) in enumerate(gene_list_data):
ax_idx = (idx//ncols, idx%ncols) if nrows > 1 else idx
# to_plot = numpy.log2(numpy.mean(data_norm, axis=0)/l2_tissue_db_norm)
# import pdb; pdb.set_trace()
if weights is True:
# to_plot = numpy.log2(numpy.average(data_norm, axis=0, weights=gene_weights)/l2_tissue_db_norm)
to_plot, errs = sample_db(data_norm, expr_db, data_weights=gene_weights, nsamples=nsamples)
else:
# to_plot = numpy.log2(numpy.average(data_norm, axis=0, weights=None)/l2_tissue_db_norm)
to_plot, errs = sample_db(data_norm, expr_db, data_weights=None, nsamples=nsamples)
for idx, (name, color) in enumerate(l2_tissue_colors):
axes[ax_idx[0],ax_idx[1]].bar(ind[idx], to_plot[idx], width, yerr=errs[:,idx][:,None], color=color, label=name)
axes[ax_idx[0],ax_idx[1]].axhline(0, color='k')
axes[ax_idx[0],ax_idx[1]].set_xlim((-1, len(expr_db_headers)))
axes[ax_idx[0],ax_idx[1]].set_title('{!s}\n({!s} genes)\n'.format(fname, data.shape[0]), fontsize=title_fontsize)
axes[ax_idx[0],ax_idx[1]].set_ylabel('Log2 ratio of mean expr proportion\n(ATAC targets:Random genes)', fontsize=axis_fontsize)
axes[ax_idx[0],ax_idx[1]].set_xlabel('L2 tissues', fontsize=axis_fontsize)
axes[ax_idx[0],ax_idx[1]].set_xticks(ind + width/2)
axes[ax_idx[0],ax_idx[1]].set_xticklabels([])
#axes[0].set_xticklabels(expr_db_headers[1:], rotation=90)
if nrows > 1:
axes[0,ncols-1].legend(bbox_to_anchor=[1.0,1.0])
else:
axes[-1].legend(bbox_to_anchor=[1.0,1.0])
if display_in_notebook is True:
fig.tight_layout()
if savefile is not None:
fig.savefig(savefile, bbox_inches='tight')
def plot_stages(nearest_genes_glob, refdata, expr_db=None, expr_db_headers=None, ncols=3, topn=TOPN, weights=False):
if expr_db is None:
#Get all stages expression data to normalize the distribution of genes from peaks
stage_db_path = os.path.join(refdata,'gexplore_stage_expr.txt')
expr_db_headers, expr_db = load_expr_db(stage_db_path)
gene_lists = glob.glob(nearest_genes_glob)
if os.path.basename(gene_lists[0]).startswith('peaks'):
gene_lists.sort(key=lambda x:int(os.path.basename(x).split('.')[0].replace('peaks', '')))
elif os.path.basename(gene_lists[0]).startswith('topic'):
gene_lists.sort(key=lambda x:int(os.path.basename(x).split('.')[1].replace('rank', '')))
else:
gene_lists.sort(key=lambda x:os.path.basename(x).split('.')[0])
gene_list_data = [(os.path.basename(path).split('.')[0], get_gene_data(path, expr_db, topn=topn)) for path in gene_lists]
print('\n'.join(['{!s} nearest genes: found {!s} out of {!s} total'.format(fname, data.shape[0], gene_list_len)
for (fname, (data_idx, data, data_norm, gene_list_len, gene_weights)) in gene_list_data]))
rs = numpy.random.RandomState(15321)
random_subset = numpy.arange(expr_db.shape[0])
rs.shuffle(random_subset)
#num_to_select = int(numpy.mean([neuron_data.shape[0], emb_muscle_data.shape[0], l2_muscle_data.shape[0]]))
num_to_select = len(random_subset)
stage_db_subset = expr_db[random_subset[:num_to_select]][:,1:].astype(float)
denom = numpy.sum(stage_db_subset, axis=1)[:,None] + 1e-8
stage_db_norm = numpy.mean(stage_db_subset/denom, axis=0)
print('Stage DB norm shape: {!s}'.format(stage_db_norm.shape))
emb_idx = [expr_db_headers[1:].index(elt) for elt in expr_db_headers[1:]
if elt.endswith('m') or elt == '4-cell']
larva_idx = [expr_db_headers[1:].index(elt) for elt in expr_db_headers[1:]
if elt.startswith('L')]
adult_idx = [expr_db_headers[1:].index(elt) for elt in expr_db_headers[1:]
if 'adult' in elt]
dauer_idx = [expr_db_headers[1:].index(elt) for elt in expr_db_headers[1:]
if 'dauer' in elt]
# rest_idx = [expr_db_headers[1:].index(elt) for elt in expr_db_headers[1:]
# if not elt.endswith('m') and not elt.startswith('L') and elt != '4-cell']
pyplot.rcParams.update({'xtick.labelsize':20,
'ytick.labelsize':20,
'xtick.major.pad':8})
ind = numpy.arange(len(expr_db_headers) - 1)
width = 0.66
axis_fontsize = 25
title_fontsize = 27
nrows = int(numpy.ceil(len(gene_list_data)/float(ncols)))
fig, axes = pyplot.subplots(nrows=nrows, ncols=ncols, figsize=(7 * ncols, 7 * nrows), sharey=True)
for idx, (fname, (data_idx, data, data_norm, gene_list_len, gene_weights)) in enumerate(gene_list_data):
ax_idx = (idx//ncols, idx%ncols) if nrows > 1 else idx
# to_plot = numpy.log2(numpy.mean(data_norm, axis=0)/stage_db_norm)
if weights is True:
to_plot = numpy.log2(numpy.average(data_norm, axis=0, weights=gene_weights)/stage_db_norm)
else:
to_plot = numpy.log2(numpy.average(data_norm, axis=0, weights=None)/stage_db_norm)
axes[ax_idx].bar(ind[emb_idx], to_plot[emb_idx], width, color='orange', label='Embryo')
axes[ax_idx].bar(ind[larva_idx], to_plot[larva_idx], width, color='blue', label='Larva')
axes[ax_idx].bar(ind[adult_idx], to_plot[adult_idx], width, color='red', label='Adult')
axes[ax_idx].bar(ind[dauer_idx], to_plot[dauer_idx], width, color='green', label='Dauer')
# axes[ax_idx].bar(ind[rest_idx], to_plot[rest_idx], width, color='grey', label='Other')
axes[ax_idx].axhline(0, color='k')
axes[ax_idx].set_xlim((-1, len(expr_db_headers)))
axes[ax_idx].set_title('{!s}\n({!s} genes)\n'.format(fname, data.shape[0]), fontsize=title_fontsize)
axes[ax_idx].set_ylabel('Log2 Ratio of Mean Expr Proportion\n(ATAC Targets:All Genes)', fontsize=axis_fontsize)
axes[ax_idx].set_xlabel('Developmental Stage', fontsize=axis_fontsize)
axes[ax_idx].set_xticks(ind + width/2)
axes[ax_idx].set_xticklabels([])
fig.tight_layout()
def leiden_clustering(umap_res, resolution_range=(0,1), random_state=2, kdtree_dist='euclidean'):
tree = neighbors.KDTree(umap_res, metric=kdtree_dist)
vals, i, j = [], [], []
for idx in range(umap_res.shape[0]):
dist, ind = tree.query([umap_res[idx]], k=25)
vals.extend(list(dist.squeeze()))
j.extend(list(ind.squeeze()))
i.extend([idx] * len(ind.squeeze()))
print(len(vals))
ginput = sps.csc_matrix((numpy.array(vals), (numpy.array(i),numpy.array(j))),
shape=(umap_res.shape[0], umap_res.shape[0]))
sources, targets = ginput.nonzero()
edgelist = zip(sources.tolist(), targets.tolist())
G = ig.Graph(edges=list(edgelist))
optimiser = leidenalg.Optimiser()
optimiser.set_rng_seed(random_state)
profile = optimiser.resolution_profile(G, leidenalg.CPMVertexPartition, resolution_range=resolution_range, number_iterations=0)
print([len(elt) for elt in profile])
return profile
def write_peaks_and_map_to_genes(data_array, row_headers, c_labels, out_dir, refseq_exon_bed,
uniqueness_threshold=3, num_peaks=1000):
#write the peaks present in each cluster to bed files
if not os.path.isdir(out_dir):
os.makedirs(out_dir)
else:
local['rm']('-r', out_dir)
os.makedirs(out_dir)
#write a file of peaks per cluster in bed format
peak_files = []
for idx, cluster_name in enumerate(sorted(set(c_labels))):
cell_coords = numpy.where(c_labels == cluster_name)
peak_sums = numpy.mean(data_array[:,cell_coords[0]], axis=1)
peak_sort = numpy.argsort(peak_sums)
# sorted_peaks = peak_sums[peak_sort]
# print('Cluster {!s} -- Present Peaks: {!s}, '
# 'Min Peaks/Cell: {!s}, '
# 'Max Peaks/Cell: {!s}, '
# 'Peaks in {!s}th cell: {!s}'.format(cluster_name, numpy.sum(peak_sums > 0),
# sorted_peaks[0], sorted_peaks[-1],
# num_peaks, sorted_peaks[-num_peaks]))
out_tmp = os.path.join(out_dir, 'peaks{!s}.tmp.bed'.format(cluster_name))
out_path = out_tmp.replace('.tmp', '')
peak_indices = peak_sort[-num_peaks:]
with open(out_tmp, 'w') as out:
out.write('\n'.join('chr'+'\t'.join(elt) if not elt[0].startswith('chr') else '\t'.join(elt)
for elt in numpy.hstack([row_headers[peak_indices],
peak_sums[peak_indices,None].astype(str)])) + '\n')
(local['sort']['-k1,1', '-k2,2n', out_tmp] > out_path)()
os.remove(out_tmp)
peak_files.append(out_path)
bedtools, sort, cut, uniq, awk = local['bedtools'], local['sort'], local['cut'], local['uniq'], local['awk']
out_subdir = os.path.join(out_dir, 'nearest_genes')
if not os.path.isdir(out_subdir):
os.makedirs(out_subdir)
nearest_genes = []
for path in sorted(peak_files):
out_path = os.path.join(out_subdir, os.path.basename(path).replace('.bed', '.nearest_genes.txt'))
cmd = (bedtools['closest', '-D', 'b', '-io', '-id', '-a', path, '-b', refseq_exon_bed] |
cut['-f1,2,3,5,9,12'] | #fields are chrom, start, stop, peak sum, gene name, distance
awk['BEGIN{OFS="\t"}{if($6 > -1200){print($1, $2, $3, $6, $5, $4);}}'] |
sort['-k5,5', '-k6,6nr'] |
cut['-f5,6'])()
with open(out_path, 'w') as out:
prev_gene = None
for idx, line in enumerate(str(cmd).strip().split('\n')):
if prev_gene is None or not line.startswith(prev_gene):
# print(line)
line_split = line.strip().split()
prev_gene = line_split[0]
out.write(line + '\n')
nearest_genes.append(out_path)
all_genes = []
# for idx in range(len(nearest_genes)):
# nearest_genes_path = os.path.join(out_subdir, 'peaks{!s}.nearest_genes.txt'.format(idx))
for nearest_genes_path in nearest_genes:
with open(nearest_genes_path) as lines_in:
all_genes.append([elt.strip().split() for elt in lines_in.readlines()])
# count_dict = Counter([i[0] for i in itertools.chain(*[all_genes[elt] for elt in range(len(nearest_genes))])])
count_dict = Counter([i[0] for i in itertools.chain(*all_genes)])
#print unique genes
for idx, nearest_genes_path in enumerate(nearest_genes):
unique_genes = [elt for elt in all_genes[idx] if count_dict[elt[0]] < uniqueness_threshold]
print(idx, len(unique_genes))
# unique_genes_path = os.path.join(out_subdir, 'peaks{!s}.nearest_genes_lt_{!s}.txt'.
# format(idx, uniqueness_threshold))
unique_genes_path = os.path.splitext(nearest_genes_path)[0] + '_lt_{!s}.txt'.format(uniqueness_threshold)
with open(unique_genes_path, 'w') as out:
out.write('\n'.join(['\t'.join(elt) for elt in unique_genes]) + '\n')
#print shared genes
shared_genes_by_cluster = []
all_genes = [dict([(k,float(v)) for k,v in elt]) for elt in all_genes]
for gene_name in sorted(count_dict.keys()):
if count_dict[gene_name] < uniqueness_threshold:
continue
shared_genes_by_cluster.append([gene_name])
for cluster_dict in all_genes:
shared_genes_by_cluster[-1].append(cluster_dict.get(gene_name, 0.0))
shared_out = os.path.join(out_subdir, 'non-unique_genes_lt_{!s}.txt'.
format(uniqueness_threshold))
numpy.savetxt(shared_out, shared_genes_by_cluster, fmt='%s')
# fmt=('%s',)+tuple('%18f' for _ in range(len(all_genes))))
return
def write_peaks_and_map_to_genes2(data_array, peak_topic_specificity, row_headers, c_labels, out_dir,
refseq_exon_bed, uniqueness_threshold=3, num_peaks=1000):
# import pdb; pdb.set_trace()
#write the peaks present in each cluster to bed files
if not os.path.isdir(out_dir):
os.makedirs(out_dir)
else:
local['rm']('-r', out_dir)
os.makedirs(out_dir)
#write a file of peaks per cluster in bed format
peak_files = []
for idx, cluster_name in enumerate(sorted(set(c_labels))):
cell_coords = numpy.where(c_labels == cluster_name)
peaks_present = numpy.sum(data_array[cell_coords[0],:], axis=0)
out_tmp = os.path.join(out_dir, 'peaks{!s}.tmp.bed'.format(cluster_name))
out_path = out_tmp.replace('.tmp', '')
# peak_indices = peak_sort[-num_peaks:]
peak_scores = (peak_topic_specificity ** 2) * peaks_present
sort_idx = numpy.argsort(peak_scores[peaks_present.astype(bool)])
peak_indices = sort_idx[-num_peaks:]
with open(out_tmp, 'w') as out:
# out.write('\n'.join('chr'+'\t'.join(elt) if not elt[0].startswith('chr') else '\t'.join(elt)
# for elt in numpy.hstack([row_headers[peaks_present.astype(bool)][peak_indices],
# peak_scores[peaks_present.astype(bool)][peak_indices,None].astype(str)])) + '\n')
out.write('\n'.join('\t'.join(elt) for elt in
numpy.hstack([row_headers[peaks_present.astype(bool)][peak_indices],
peak_scores[peaks_present.astype(bool)][peak_indices,None].astype(str)])) + '\n')
(local['sort']['-k1,1', '-k2,2n', out_tmp] > out_path)()
os.remove(out_tmp)
peak_files.append(out_path)
bedtools, sort, cut, uniq, awk = local['bedtools'], local['sort'], local['cut'], local['uniq'], local['awk']
out_subdir = os.path.join(out_dir, 'nearest_genes')
if not os.path.isdir(out_subdir):
os.makedirs(out_subdir)
nearest_genes = []
for path in sorted(peak_files):
out_path = os.path.join(out_subdir, os.path.basename(path).replace('.bed', '.nearest_genes.txt'))
cmd = (bedtools['closest', '-D', 'b', '-io', '-id', '-a', path, '-b', refseq_exon_bed] |
cut['-f1,2,3,5,9,12'] | #fields are chrom, start, stop, peak sum, gene name, distance
awk['BEGIN{OFS="\t"}{if($6 > -1200){print($1, $2, $3, $6, $5, $4);}}'] |
sort['-k5,5', '-k6,6nr'] |
cut['-f5,6'])()
with open(out_path, 'w') as out:
prev_gene = None
for idx, line in enumerate(str(cmd).strip().split('\n')):
if prev_gene is None or not line.startswith(prev_gene):
# print(line)
line_split = line.strip().split()
prev_gene = line_split[0]
out.write(line + '\n')
nearest_genes.append(out_path)
all_genes = []
# for idx in range(len(nearest_genes)):
# nearest_genes_path = os.path.join(out_subdir, 'peaks{!s}.nearest_genes.txt'.format(idx))
for nearest_genes_path in nearest_genes:
with open(nearest_genes_path) as lines_in:
all_genes.append([elt.strip().split() for elt in lines_in.readlines()])
# count_dict = Counter([i[0] for i in itertools.chain(*[all_genes[elt] for elt in range(len(nearest_genes))])])
count_dict = Counter([i[0] for i in itertools.chain(*all_genes)])
#print unique genes
for idx, nearest_genes_path in enumerate(nearest_genes):
unique_genes = [elt for elt in all_genes[idx] if count_dict[elt[0]] < uniqueness_threshold]
print(idx, len(unique_genes))
# unique_genes_path = os.path.join(out_subdir, 'peaks{!s}.nearest_genes_lt_{!s}.txt'.
# format(idx, uniqueness_threshold))
unique_genes_path = os.path.splitext(nearest_genes_path)[0] + '_lt_{!s}.txt'.format(uniqueness_threshold)
with open(unique_genes_path, 'w') as out:
out.write('\n'.join(['\t'.join(elt) for elt in unique_genes]) + '\n')
#print shared genes
shared_genes_by_cluster = []
all_genes = [dict([(k,float(v)) for k,v in elt]) for elt in all_genes]
for gene_name in sorted(count_dict.keys()):
if count_dict[gene_name] < uniqueness_threshold:
continue
shared_genes_by_cluster.append([gene_name])
for cluster_dict in all_genes:
shared_genes_by_cluster[-1].append(cluster_dict.get(gene_name, 0.0))
shared_out = os.path.join(out_subdir, 'non-unique_genes_lt_{!s}.txt'.
format(uniqueness_threshold))
numpy.savetxt(shared_out, shared_genes_by_cluster, fmt='%s')
# fmt=('%s',)+tuple('%18f' for _ in range(len(all_genes))))
return
def write_peaks_and_map_to_genes3(data_array, row_headers, c_labels, out_dir,
refseq_exon_bed, uniqueness_threshold=3, num_peaks=1000):
# import pdb; pdb.set_trace()
#write the peaks present in each cluster to bed files
if not os.path.isdir(out_dir):
os.makedirs(out_dir)
else:
local['rm']('-r', out_dir)
os.makedirs(out_dir)
agg_clusters = numpy.vstack([numpy.sum(data_array[numpy.where(c_labels == cluster_idx)[0]], axis=0)
for cluster_idx in sorted(set(c_labels))])
tfidf = TfidfTransformer(norm='l2', use_idf=True, smooth_idf=True, sublinear_tf=False)
agg_clusters_tfidf = tfidf.fit_transform(agg_clusters).toarray()
#write a file of peaks per cluster in bed format
peak_files = []
for idx, cluster_name in enumerate(sorted(set(c_labels))):
out_tmp = os.path.join(out_dir, 'peaks{!s}.tmp.bed'.format(cluster_name))
out_path = out_tmp.replace('.tmp', '')
sort_idx = numpy.argsort(agg_clusters_tfidf[idx])
peak_indices = sort_idx[-num_peaks:]
with open(out_tmp, 'w') as out:
# out.write('\n'.join('chr'+'\t'.join(elt) if not elt[0].startswith('chr') else '\t'.join(elt)
# for elt in numpy.hstack([row_headers[peaks_present.astype(bool)][peak_indices],
# peak_scores[peaks_present.astype(bool)][peak_indices,None].astype(str)])) + '\n')
out.write('\n'.join('\t'.join(elt) for elt in
numpy.hstack([row_headers[peak_indices],
agg_clusters_tfidf[idx][peak_indices,None].astype(str)])) + '\n')
(local['sort']['-k1,1', '-k2,2n', out_tmp] > out_path)()
os.remove(out_tmp)
peak_files.append(out_path)
bedtools, sort, cut, uniq, awk = local['bedtools'], local['sort'], local['cut'], local['uniq'], local['awk']
out_subdir = os.path.join(out_dir, 'nearest_genes')
if not os.path.isdir(out_subdir):
os.makedirs(out_subdir)
nearest_genes = []
for path in sorted(peak_files):
out_path = os.path.join(out_subdir, os.path.basename(path).replace('.bed', '.nearest_genes.txt'))
cmd = (bedtools['closest', '-D', 'b', '-io', '-id', '-a', path, '-b', refseq_exon_bed] |
cut['-f1,2,3,5,9,12'] | #fields are chrom, start, stop, peak sum, gene name, distance
awk['BEGIN{OFS="\t"}{if($6 > -1200){print($1, $2, $3, $6, $5, $4);}}'] |
sort['-k5,5', '-k6,6nr'] |
cut['-f5,6'])()
with open(out_path, 'w') as out:
prev_gene = None
for idx, line in enumerate(str(cmd).strip().split('\n')):
if prev_gene is None or not line.startswith(prev_gene):
# print(line)
line_split = line.strip().split()
prev_gene = line_split[0]
out.write(line + '\n')
nearest_genes.append(out_path)
all_genes = []
# for idx in range(len(nearest_genes)):
# nearest_genes_path = os.path.join(out_subdir, 'peaks{!s}.nearest_genes.txt'.format(idx))
for nearest_genes_path in nearest_genes:
with open(nearest_genes_path) as lines_in:
all_genes.append([elt.strip().split() for elt in lines_in.readlines()])
# count_dict = Counter([i[0] for i in itertools.chain(*[all_genes[elt] for elt in range(len(nearest_genes))])])
count_dict = Counter([i[0] for i in itertools.chain(*all_genes)])
#print unique genes
for idx, nearest_genes_path in enumerate(nearest_genes):
unique_genes = [elt for elt in all_genes[idx] if count_dict[elt[0]] < uniqueness_threshold]
print(idx, len(unique_genes))
# unique_genes_path = os.path.join(out_subdir, 'peaks{!s}.nearest_genes_lt_{!s}.txt'.
# format(idx, uniqueness_threshold))
unique_genes_path = os.path.splitext(nearest_genes_path)[0] + '_lt_{!s}.txt'.format(uniqueness_threshold)
with open(unique_genes_path, 'w') as out:
out.write('\n'.join(['\t'.join(elt) for elt in unique_genes]) + '\n')
#print shared genes
shared_genes_by_cluster = []
all_genes = [dict([(k,float(v)) for k,v in elt]) for elt in all_genes]
for gene_name in sorted(count_dict.keys()):
if count_dict[gene_name] < uniqueness_threshold:
continue
shared_genes_by_cluster.append([gene_name])
for cluster_dict in all_genes:
shared_genes_by_cluster[-1].append(cluster_dict.get(gene_name, 0.0))
shared_out = os.path.join(out_subdir, 'non-unique_genes_lt_{!s}.txt'.
format(uniqueness_threshold))
numpy.savetxt(shared_out, shared_genes_by_cluster, fmt='%s')
# fmt=('%s',)+tuple('%18f' for _ in range(len(all_genes))))
return
# -
# ## Peaks model
# +
#read in sc peak table
peaktable_path = '../tissue_analysis/glia/filtered_peaks_iqr4.0_low_cells.bow'
peak_data_sparse = numpy.loadtxt(peaktable_path, dtype=int, skiprows=3)
peak_data = sps.csr_matrix((peak_data_sparse[:,2], (peak_data_sparse[:,0] - 1, peak_data_sparse[:,1] - 1)))
cell_names_path = '../tissue_analysis/glia/filtered_peaks_iqr4.0_low_cells.indextable.txt'
cell_names = numpy.loadtxt(cell_names_path, dtype=object)[:,0]
peak_names_path = '../tissue_analysis/glia/filtered_peaks_iqr4.0_low_cells.extra_cols.bed'
peak_row_headers = numpy.loadtxt(peak_names_path, dtype=object)
#chr_regex = re.compile('[:-]')
peak_row_headers = numpy.hstack([peak_row_headers, numpy.array(['name'] * peak_row_headers.shape[0])[:,None]])
print(peak_data.shape)
orig_peaktable_path = '../tissue_analysis/glia/glia_all_peaks.bow'
orig_peak_data_sparse = numpy.loadtxt(orig_peaktable_path, dtype=int, skiprows=3)
orig_peak_data = sps.csr_matrix((orig_peak_data_sparse[:,2],
(orig_peak_data_sparse[:,0] - 1, orig_peak_data_sparse[:,1] - 1)))
orig_cell_names_path = '../tissue_analysis/glia/glia_all_peaks.zeros_filtered.indextable.txt'
orig_cell_names = numpy.loadtxt(orig_cell_names_path, dtype=object)[:,0]
orig_peak_names_path = '../tissue_analysis/glia/glia_all_peaks.zeros_filtered.bed'
orig_peak_row_headers = numpy.loadtxt(orig_peak_names_path, dtype=object)
#chr_regex = re.compile('[:-]')
orig_peak_row_headers = numpy.hstack([orig_peak_row_headers,
numpy.array(['name'] * orig_peak_row_headers.shape[0])[:,None]])
print(orig_peak_data.shape)
#peak_binary = sps.csr_matrix((numpy.ones((peak_data_sparse.shape[0],)),
# (peak_data_sparse[:,0], peak_data_sparse[:,1])))
#print(peak_binary.shape)
#make a cells x peaks array
peak_data_array = peak_data.toarray().astype(numpy.int8)
print(peak_data_array.shape)
del(peak_data)
orig_peak_data_array = orig_peak_data.toarray().astype(numpy.int8)
print(orig_peak_data_array.shape)
del(orig_peak_data)
# -
cell_idx_map = [numpy.where(orig_cell_names == elt)[0][0] for elt in cell_names]
orig_peak_data_array_filtered = orig_peak_data_array[cell_idx_map]
print(orig_peak_data_array_filtered.shape)
numpy.median(numpy.sum(peak_data_array, axis=1))
numpy.median(numpy.sum(peak_data_array, axis=0))
numpy.max(peak_data_array)
# ## Analysis functions
# +
REFDATA = 'ATAC_sequencing/2018_worm_atac/ref_data/WS235'
refseq_exon_bed = os.path.join(REFDATA, 'c_elegans.WS272.canonical_geneset.genes.common_names.sorted.bed.gz')
import gzip
ucsc = True if peak_row_headers[0][0].startswith('chr') else False
with gzip.open(refseq_exon_bed, 'rb') as lines_in:
exon_locs = []
for line in lines_in:
line = line.decode()[3:].strip().split()
if ucsc is True:
line[0] = 'chr{!s}'.format(line[0])
line[1] = int(line[1])
line[2] = int(line[2])
exon_locs.append(line)
gene_locs = {}
for exon in exon_locs:
gene_locs.setdefault(exon[3], []).append(exon)
for gene, locs in gene_locs.items():
gene_locs[gene] = sorted(locs, key=lambda x:(x[1],x[2]))
# +
class DistanceException(Exception):
pass
class NoPeakException(Exception):
pass
def get_closest_peaks(gene_name, row_headers, verbose=False, dist_threshold=1200, dist_excpt=False):
gene_coord = gene_locs[gene_name][0] if gene_locs[gene_name][0][-1] == '+' else gene_locs[gene_name][-1]
if verbose:
print(gene_coord)
if gene_coord[-1] == '+':
try:
nearest_peak = numpy.where(numpy.logical_and(row_headers[:,0] == gene_coord[0],
row_headers[:,1].astype(int) <= gene_coord[1]))[0][-1]
except IndexError:
raise NoPeakException()
alt_peak = nearest_peak - 1
# peak_dist = numpy.absolute(gene_coord[1] - row_headers[[nearest_peak, alt_peak],1].astype(int))
peak_dist = gene_coord[1] - row_headers[[nearest_peak, alt_peak],2].astype(int)
if verbose:
print(row_headers[[nearest_peak, alt_peak]])
print(peak_dist)
else:
try:
nearest_peak = numpy.where(numpy.logical_and(row_headers[:,0] == gene_coord[0],
row_headers[:,2].astype(int) >= gene_coord[2]))[0][0]
except IndexError:
raise NoPeakException()
alt_peak = nearest_peak + 1
# peak_dist = numpy.absolute(gene_coord[2] - row_headers[[nearest_peak, alt_peak],2].astype(int))
peak_dist = row_headers[[nearest_peak, alt_peak],1].astype(int) - gene_coord[2]
if verbose:
print(row_headers[[nearest_peak, alt_peak]])
print(peak_dist)
if peak_dist[0] > dist_threshold:
msg = 'Warning: nearest peak to {!s} is far away! ({!s} bp)'.format(gene_name, peak_dist[0])
if dist_excpt:
raise DistanceException(msg)
else:
print(msg)
return nearest_peak, alt_peak
def get_closest_peaks2(gene_name, row_headers, verbose=False, dist_threshold=1200, gene_end_threshold=100, dist_excpt=False):
gene_coord = gene_locs[gene_name][0] if gene_locs[gene_name][0][-1] == '+' else gene_locs[gene_name][-1]
# gene_coord = gene_locs[gene_name][0]
# gene_coord[2] = gene_locs[gene_name][-1][2]
if verbose:
print(gene_coord)
if gene_coord[-1] == '+':
try: #same chromosome
nearest_peaks = numpy.where(numpy.logical_and(row_headers[:,0] == gene_coord[0],
#peak start is before gene stop
#peak stop is after gene start - dist threshold and close to gene end
numpy.logical_and(row_headers[:,1].astype(int) <= (gene_coord[2] + gene_end_threshold),
row_headers[:,2].astype(int) >= (gene_coord[1] - dist_threshold))))[0]
# numpy.logical_and(row_headers[:,2].astype(int) >= (gene_coord[1] - dist_threshold),
# row_headers[:,2].astype(int) <= (gene_coord[2] + gene_end_threshold)))))[0]
except IndexError:
raise NoPeakException()
# alt_peak = nearest_peak - 1
# peak_dist = numpy.absolute(gene_coord[1] - row_headers[[nearest_peak, alt_peak],1].astype(int))
peak_dist = gene_coord[1] - row_headers[nearest_peaks,1].astype(int)
if verbose:
print(row_headers[nearest_peaks])
print(peak_dist)
else:
try:
nearest_peaks = numpy.where(numpy.logical_and(row_headers[:,0] == gene_coord[0],
numpy.logical_and(row_headers[:,2].astype(int) >= (gene_coord[1] - gene_end_threshold),
row_headers[:,1].astype(int) <= (gene_coord[2] + dist_threshold))))[0]
# numpy.logical_and(row_headers[:,1].astype(int) <= (gene_coord[2] + dist_threshold),
# row_headers[:,1].astype(int) >= (gene_coord[1] - gene_end_threshold)))))[0]
except IndexError:
raise NoPeakException()
# alt_peak = nearest_peak + 1
# peak_dist = numpy.absolute(gene_coord[2] - row_headers[[nearest_peak, alt_peak],2].astype(int))
peak_dist = row_headers[nearest_peaks,2].astype(int) - gene_coord[2]
if verbose:
print(row_headers[nearest_peaks])
print(peak_dist)
# if peak_dist[0] > dist_threshold:
# msg = 'Warning: nearest peak to {!s} is far away! ({!s} bp)'.format(gene_name, peak_dist[0])
# if dist_excpt:
# raise DistanceException(msg)
# else:
# print(msg)
return nearest_peaks
def get_gene_cells(gene_name, row_headers, peak_data_array, **kwargs):
nearest_peaks = get_closest_peaks2(gene_name, row_headers, **kwargs)
cells_idx = numpy.any(peak_data_array[:,nearest_peaks], axis=1)
return cells_idx
def get_gene_idx(gene_name, row_headers, peaktopic_frac, topic_prob_threshold=0.5, **kwargs):
nearest_peak, alt_peak = get_closest_peaks(gene_name, row_headers, **kwargs)
topic_idx = numpy.argsort(peaktopic_frac[nearest_peak])[::-1]
num_to_get = numpy.where(numpy.cumsum(peaktopic_frac[nearest_peak][topic_idx]) > topic_prob_threshold)[0][0] + 1
return nearest_peak, topic_idx[:num_to_get]
def get_gene_topn_topics(gene_name, row_headers, peaktopic_frac, ntopics=1, **kwargs):
nearest_peak, alt_peak = get_closest_peaks(gene_name, row_headers, **kwargs)
topic_idx = numpy.argsort(peaktopic_frac[nearest_peak])[::-1]
return nearest_peak, topic_idx[:ntopics]
# -
# ## Topic Mode
# +
doctopic_path = '../tissue_analysis/glia/0000_topics8_alpha3.000_beta2000.000/topic_mode.theta'
doctopic_peaks = numpy.loadtxt(doctopic_path, delimiter=',', dtype=float)
print(doctopic_peaks.shape)
#center and scale the topic values
#col_means = numpy.mean(doctopic.T, axis=0)
#doctopic_norm = doctopic.T - col_means
#doctopic_norm = doctopic_norm / numpy.std(doctopic_norm, axis=0)
#doctopic_norm = doctopic_norm.T
#print(doctopic_norm.shape)
col_means = numpy.mean(doctopic_peaks.T, axis=0)
doctopic_peaks_norm = doctopic_peaks.T - col_means
l2_for_norm = (doctopic_peaks_norm ** 2).sum(axis=0).flatten() ** 0.5
doctopic_peaks_norm /= l2_for_norm
doctopic_peaks_norm = doctopic_peaks_norm.T
print(doctopic_peaks_norm.shape)
doctopic_peaks_frac = (doctopic_peaks.T/doctopic_peaks.sum(axis=1).astype(float)).T
print(doctopic_peaks_frac.shape)
# +
peaktopic_path = '../tissue_analysis/glia/0000_topics8_alpha3.000_beta2000.000/topic_mode.wordTopic'
peaktopic = numpy.loadtxt(peaktopic_path, delimiter=',', dtype=float)
print(peaktopic.shape)
#center and scale the topic values
#col_means = numpy.mean(doctopic.T, axis=0)
#doctopic_norm = doctopic.T - col_means
#doctopic_norm = doctopic_norm / numpy.std(doctopic_norm, axis=0)
#doctopic_norm = doctopic_norm.T
#print(doctopic_norm.shape)
nonzero_idx = numpy.where(numpy.sum(peaktopic, axis=1) > 0)[0]
peaktopic = peaktopic[nonzero_idx]
peak_row_headers = peak_row_headers[nonzero_idx]
peak_data_array = peak_data_array[:,nonzero_idx]
col_means = numpy.mean(peaktopic.T, axis=0)
peaktopic_norm = peaktopic.T - col_means
l2_for_norm = (peaktopic_norm ** 2).sum(axis=0).flatten() ** 0.5
peaktopic_norm /= l2_for_norm
peaktopic_norm = peaktopic_norm.T
print(peaktopic_norm.shape)
peaktopic_frac = (peaktopic.T/peaktopic.sum(axis=1).astype(float)).T
print(peaktopic_frac.shape)
# -
doctopic_peaks_umap3_obj = umap.UMAP(n_components=3, random_state=253)
doctopic_peaks_umap3_res = doctopic_peaks_umap3_obj.fit_transform(doctopic_peaks_norm)
print(doctopic_peaks_umap3_res.shape)
doctopic_peaks_umap2_obj = umap.UMAP(n_components=2, n_neighbors=15, random_state=1)
doctopic_peaks_umap2_res = doctopic_peaks_umap2_obj.fit_transform(doctopic_peaks_norm)
print(doctopic_peaks_umap2_res.shape)
ncols=4
nrows = int(numpy.ceil(doctopic_peaks_frac.shape[1]/ncols))
fig, axes = pyplot.subplots(nrows=nrows, ncols=ncols, figsize=(3.5*ncols,3*nrows))
for idx, topic in enumerate(numpy.arange(doctopic_peaks_frac.shape[1])):
row_idx, col_idx = int(idx/ncols), int(idx%ncols)
if nrows > 1 and ncols > 1:
ax = axes[row_idx, col_idx]
elif nrows > 1 or ncols > 1:
ax = axes[idx]
else:
ax = axes
s = ax.scatter(doctopic_peaks_umap2_res[:,0],
doctopic_peaks_umap2_res[:,1],
cmap='viridis',
c=doctopic_peaks_frac[:,topic],
s=2)
ax.set_ylabel('UMAP2')
ax.set_xlabel('UMAP1')
ax.set_title('Topic {!s}'.format(topic))
fig.colorbar(s, ax=ax)
#cbar = fig.colorbar(s, ticks=numpy.arange(len(gene_names)))
#cbar.ax.set_yticklabels(gene_names, fontsize=10)
fig.tight_layout()
# ## Construct an AnnData object and save it in loom format
# +
def add_lda_result_to_anndata_obj(anndata_obj, lda_base, lda_cellnames, lda_peak_bed):
filt_cellnames = numpy.loadtxt(lda_cellnames, dtype=object)[:,0]
filt_cellnames_set = set(filt_cellnames)
filt_cellnames_map = [(True, idx, numpy.where(filt_cellnames == elt)[0][0])
if elt in filt_cellnames_set else (False, idx, numpy.nan)
for idx, elt in enumerate(anndata_obj.obs.index.values)]
unfilt_cellnames_idx, filt_cellnames_idx = [list(idx_tuple) for idx_tuple in zip(*[(elt[1], elt[2]) for elt in filt_cellnames_map if elt[0] is True])]
anndata_obj.obs['lda_cell'] = [elt[0] for elt in filt_cellnames_map]
theta = numpy.loadtxt(lda_base + '.theta', delimiter=',', dtype=float)
unfiltered_theta = numpy.ones((len(filt_cellnames_map), theta.shape[1])) * numpy.nan
unfiltered_theta[unfilt_cellnames_idx] = theta[filt_cellnames_idx]
anndata_obj.obsm['lda_theta'] = unfiltered_theta
col_means = numpy.mean(theta.T, axis=0)
theta_norm = theta.T - col_means
l2_for_norm = (theta_norm ** 2).sum(axis=0).flatten() ** 0.5
theta_norm /= l2_for_norm
theta_norm = theta_norm.T
unfiltered_theta_norm = numpy.ones(unfiltered_theta.shape) * numpy.nan
unfiltered_theta_norm[unfilt_cellnames_idx] = theta_norm[filt_cellnames_idx]
anndata_obj.obsm['lda_theta_norm'] = unfiltered_theta_norm
doctopic = numpy.loadtxt(lda_base + '.docTopic', delimiter=',', dtype=float)
unfiltered_doctopic = numpy.ones(unfiltered_theta.shape) * numpy.nan
unfiltered_doctopic[unfilt_cellnames_idx] = doctopic[filt_cellnames_idx]
anndata_obj.obsm['lda_doctopic'] = unfiltered_doctopic
doctopic_frac = (doctopic.T/doctopic.sum(axis=1)).T
unfiltered_doctopic_frac = numpy.ones(unfiltered_theta.shape) * numpy.nan
unfiltered_doctopic_frac[unfilt_cellnames_idx] = doctopic_frac[filt_cellnames_idx]
anndata_obj.obsm['lda_doctopic_frac'] = unfiltered_doctopic_frac
filt_peaks = numpy.loadtxt(lda_peak_bed, dtype=object)
filt_peaks_str = ['{!s}\t{!s}\t{!s}'.format(*filt_peaks[idx,:3])
for idx in range(filt_peaks.shape[0])]
filt_peaks_set = set(filt_peaks_str)
unfilt_peaks_str = ['{!s}\t{!s}\t{!s}'.format(anndata_obj.var['chr'][idx],
anndata_obj.var['start'][idx],
anndata_obj.var['stop'][idx])
for idx in range(anndata_obj.var.shape[0])]
filt_peaks_map = [(True, idx, filt_peaks_str.index(elt))
if elt in filt_peaks_set else (False, idx, numpy.nan)
for idx, elt in enumerate(unfilt_peaks_str)]
unfilt_peaks_idx, filt_peaks_idx = [list(idx_tuple) for idx_tuple in zip(*[(elt[1], elt[2]) for elt in filt_peaks_map if elt[0] is True])]
anndata_obj.var['lda_peak'] = [elt[0] for elt in filt_peaks_map]
phi = numpy.loadtxt(lda_base + '.phi', delimiter=',', dtype=float).T
unfiltered_phi = numpy.ones((anndata_obj.var.shape[0], phi.shape[1])) * numpy.nan
unfiltered_phi[unfilt_peaks_idx] = phi[filt_peaks_idx]
anndata_obj.varm['lda_phi'] = unfiltered_phi
wordtopic = numpy.loadtxt(lda_base + '.wordTopic', delimiter=',', dtype=float)
unfiltered_wordtopic = numpy.ones(unfiltered_phi.shape) * numpy.nan
unfiltered_wordtopic[unfilt_peaks_idx] = wordtopic[filt_peaks_idx]
anndata_obj.varm['lda_wordtopic'] = unfiltered_wordtopic
wordtopic_frac = (wordtopic.T/wordtopic.sum(axis=1)).T
unfiltered_wordtopic_frac = numpy.ones(unfiltered_phi.shape) * numpy.nan
unfiltered_wordtopic_frac[unfilt_peaks_idx] = wordtopic_frac[filt_peaks_idx]
anndata_obj.varm['lda_wordtopic_frac'] = unfiltered_wordtopic_frac
return
def read_in_bow(bow_path, cell_names_path, feature_info_path, gene_name_map=None):
try:
bow_data_sparse = numpy.loadtxt(bow_path, dtype=int, skiprows=3)
except StopIteration:
#probably NFS lag; just wait a few seconds and try again
time.sleep(10)
bow_data_sparse = numpy.loadtxt(bow_path, dtype=int, skiprows=3)
open_func, open_mode = (gzip.open, 'rb') if bow_path.endswith('.gz') else (open, 'rb')
with open_func(bow_path, open_mode) as lines_in:
cellnum = int(lines_in.readline().decode().strip())
featnum = int(lines_in.readline().decode().strip())
bow_data = sps.csr_matrix((bow_data_sparse[:,2],
(bow_data_sparse[:,0] - 1,
bow_data_sparse[:,1] - 1)),
shape=(cellnum, featnum))
try:
cell_names = numpy.loadtxt(cell_names_path, dtype=object)[:,0]
except StopIteration:
time.sleep(10)
cell_names = numpy.loadtxt(cell_names_path, dtype=object)[:,0]
try:
feature_info_bed = numpy.loadtxt(feature_info_path, dtype=object)
except StopIteration:
time.sleep(10)
feature_info_bed = numpy.loadtxt(feature_info_path, dtype=object)
col_names = ['chr', 'start', 'stop', 'name', 'score', 'strand']
num_cols = feature_info_bed.shape[1]
if num_cols <= 6:
col_names = col_names[:num_cols]
else:
col_names = col_names + list(numpy.arange(6, num_cols).astype(str))
anndata_obj = anndata.AnnData(X=bow_data,
obs=pandas.DataFrame(index=cell_names),
var=pandas.DataFrame(data=feature_info_bed,
columns=col_names))
if gene_name_map is not None:
gene_name_map = numpy.loadtxt(gene_name_map, delimiter='\t', dtype=object)
gene_name_map = dict([gene_name_map[idx] for idx in range(gene_name_map.shape[0])])
anndata_obj.var['common_name'] = [gene_name_map.get(elt, elt) for elt in anndata_obj.var.name]
return anndata_obj
# +
#make and save anndata object
import anndata
import pandas
#first, read in the original BOW data (the unfiltered data)
orig_peaktable_path = '../tissue_analysis/glia/glia_all_peaks.bow'
orig_cell_names_path = '../tissue_analysis/glia/glia_all_peaks.zeros_filtered.indextable.txt'
orig_peak_names_path = '../tissue_analysis/glia/glia_all_peaks.zeros_filtered.bed'
anndata_obj = read_in_bow(orig_peaktable_path, orig_cell_names_path, orig_peak_names_path)
#next, add the LDA results, taking into account that some of the cells/peaks were filtered out before running LDA
lda_base = '../tissue_analysis/glia/0000_topics8_alpha3.000_beta2000.000/topic_mode'
lda_cellnames = '../tissue_analysis/glia/filtered_peaks_iqr4.0_low_cells.indextable.txt'
lda_peak_bed = '../tissue_analysis/glia/filtered_peaks_iqr4.0_low_cells.extra_cols.bed'
add_lda_result_to_anndata_obj(anndata_obj, lda_base, lda_cellnames, lda_peak_bed)
print(anndata_obj)
# +
umap_to_add = numpy.zeros((anndata_obj.shape[0], doctopic_peaks_umap2_res.shape[1])) * numpy.nan
anndata_idx = numpy.where(anndata_obj.obs.lda_cell)[0]
umap_idx = [numpy.where(cell_names == elt)[0][0] for elt in anndata_obj.obs[anndata_obj.obs.lda_cell].index.values]
umap_to_add[anndata_idx] = doctopic_peaks_umap2_res[umap_idx]
print(umap_to_add.shape)
anndata_obj.obsm['umap2'] = umap_to_add
umap_to_add = numpy.zeros((anndata_obj.shape[0], doctopic_peaks_umap3_res.shape[1])) * numpy.nan
anndata_idx = numpy.where(anndata_obj.obs.lda_cell)[0]
umap_idx = [numpy.where(cell_names == elt)[0][0] for elt in anndata_obj.obs[anndata_obj.obs.lda_cell].index.values]
umap_to_add[anndata_idx] = doctopic_peaks_umap3_res[umap_idx]
print(umap_to_add.shape)
anndata_obj.obsm['umap3'] = umap_to_add
# -
anndata_obj.write_loom('../tissue_analysis/glia/primary_lda_results.loom',
write_obsm_varm=True)
|
notebooks/glia_analysis.primary_lda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from scratch.logistic_regression import tuples
from matplotlib import pyplot as plt
data = tuples
xs = [[1.0] + list(row[:2]) for row in data]
ys = [row[2] for row in data]
# -
# salary vs years experience vs paid / unpaid users
plt.scatter([row[1]
for row, y in zip(xs, ys)
if y],
[row[2]
for row, y in zip(xs, ys)
if y], label='paid')
plt.scatter([row[1]
for row, y in zip(xs, ys)
if not y],
[row[2]
for row, y in zip(xs, ys)
if not y],
marker='+', label='unpaid');
plt.title('Paid and unpaid users')
plt.xlabel('years experience')
plt.ylabel('annual salary')
plt.legend();
# +
# lets model the problem as a linear regression
# can we predict who is a paid user in the form paid = B * X ?
# we can model it for sure!
from scratch.working_with_data import rescale
from scratch.multiple_regression import least_squares_fit, predict
from scratch.gradient_descent import gradient_step
import math
# -
learning_rate = 0.001
rescaled_xs = rescale(xs)
# issues with negative or over 1 values of predicted
# biased values for large experience values, larger error
beta = least_squares_fit(rescaled_xs, ys, learning_rate, 1000, 1)
predictions = [predict(xi, beta) for xi in rescaled_xs]
plt.scatter(predictions, ys)
plt.xlabel("predicted")
plt.ylabel("actual")
plt.show()
# +
# lets create a link function
def logistic(x: float)->float:
return 1.0 / (1 + math.exp(-x))
# -
xs2 = [xi / 10 for xi in range(-100, 100)]
ys2 = [logistic(xi) for xi in xs2]
plt.plot(xs2, ys2)
plt.title('Logistic function');
# +
# as the input gets lower --> output approx 0
# as the input gets higher --> output approx 1
# th elogistic fn also as a convineint derivative
def logistic_prime(x: float) ->float:
y = logistic(x)
return y * (1 - y)
# we'll use it to fit the model:
# y = fn(X * B) + err
# where f is the logistic fn
# in least squares min the sum squared err and maximizing likelihood of data is the same
# in log regression this is not the case so
# we'll use the gradient that maximizes the likelihood directly
# the pdf for yi can be written
# p(y| xi, B) = fn(xi * B)** yi * (1 - fn(xi * B)) ** (1 - yi)
# turns out it's easier to maximize the log likelihood
# log L(B|xi, yi) = yi * log(fn(xi * B)) + (1 - yi) * log(1 - fn(xi * B))
# because log is strictly increasin, any beta that maximizes the log also
# maximizes the likelihood
from scratch.linear_algebra import Vector, dot
# grad descent minimizes,
# so we will maximize the negative log likelihood
def _negative_log_likelihood(x: Vector, y:float, beta:Vector) -> float:
"""
The negative log likelihood for one data point
"""
if y == 1:
return -math.log(logistic(dot(x,beta)))
else:
return -math.log(1 - logistic(dot(x, beta)))
# as we assume data points are independent in this model
# overall likelihood is the product of individual likelihoods
# in log terms that means we can sum the likelihoods
def negative_log_likelihood(xs: list[Vector],
ys: list[float],
beta: Vector) -> float:
return sum(_negative_log_likelihood(xi, yi, beta)
for xi, yi in zip(xs, ys))
# now for the gradient
from scratch.linear_algebra import vector_sum
def _negative_log_partial(x: Vector, y: float,
beta: Vector, j:int) -> float:
"""
The jth partial derivative for one data point
ith is the index of the data point
"""
return -(y - logistic(dot(x, beta))) * x[j]
def _negative_log_gradient(x: Vector, y: float,
beta: Vector)-> Vector:
"""
The Gradient for one data point
"""
return [_negative_log_partial(x, y, beta, j)
for j in range(len(beta))]
def negative_log_gradient(xs: list[Vector],
ys: list[float],
beta: Vector)-> Vector:
return vector_sum([_negative_log_gradient(xi, yi, beta)
for xi, yi in zip(xs, ys)])
# applying the model
from scratch.machine_learning import train_test_split
import random
import tqdm
random.seed(0)
x_train, x_test, y_train, y_test = train_test_split(rescaled_xs, ys, 0.33)
# +
learning_rate = 0.01
# random starting point for beta
beta = [random.random() for _ in rescaled_xs[0]]
with tqdm.trange(5_000) as t:
for epoch in t:
gradient = negative_log_gradient(x_train, y_train, beta)
beta = gradient_step(beta, gradient, -learning_rate)
loss = negative_log_likelihood(x_train, y_train, beta)
t.set_description(f"loss: {loss:.3f} beta: {beta}")
# +
# these are the coef for the rescaled data
# lets convert them back to original scale
from scratch.working_with_data import scale
means, std = scale(xs)
beta_0 = (beta[0] - beta[1] * means[1] / std[1]) - (beta[2] * means[2] / std[2])
beta_unscaled = [beta_0,
beta[1] / std[1],
beta[2] / std[2]]
beta_unscaled
# not as easy to interpret as liner regression
# all else being equal an extra year of experience
# adds 1.6 to the input of logistic fn
#the impact on the output also depends on the other inputs as well
# if dot(beta, x_i) is already large/ small increasing bi a lot or little will
# not affectc the outcome of the probability
# all else being equal - people with more experience
# are more likly to pay for accounts
# and people of larger salary less likely
# to pay for accounts
# +
# Goodness of Fit
# ie how good is this model
tp = fp = tn = fn = 0
# nice implicit control flow
for xi, yi in zip(x_test, y_test):
prediction = logistic(dot(beta, xi))
if yi == 1 and prediction >= .5:
tp += 1
elif yi == 1:
fn += 1
elif prediction >= .5:
fp += 1
else:
tn += 1
precision = tp / (tp + fp)
recall = tp / (tp + fn)
precision, recall
# -
predictions = [logistic(dot(beta, xi))
for xi in x_test]
plt.scatter(predictions, y_test, marker='+')
plt.title('Logistic Regression Predicted vs. Actual')
plt.xlabel('predicted probability')
plt.ylabel('actual outcome');
|
Logistic_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project Milestone -- 2
#
# ## Group 1 -- SARSA
#
#
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete by Next Milestone
# | --- | --- | --- | --- |
# | Code | 20% | Apr 21 | 60% |
# | Paper | 10% | Apr 21 | 30% |
# | Demo | 10% | May 01 | 30% |
# | Presentation | 10% | May 05 | 20% |
#
#
# 1. What deliverable goals established in the last milestone report were accomplished to the anticipated percentage?
# -- Our goals for the code and paper that were set from last week were met. We also managed to get started on our demo and our presentation.
#
# 2. What deliverable goals established in the last milestone report were not accomplished to the anticipated percentage?
# -- We met all of our goals established in the last milestone report and exceeded some as well.
#
# 3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?
#
# <NAME> -- Continue working on my part of the YOLO V3 network, and build a mini presentation.
#
# <NAME> --
#
# <NAME> --
#
# <NAME> --
#
# <NAME> --
#
# <NAME> --
|
Milestone2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
df= pd.read_csv('Iris.csv')
df
df.drop(columns='Id',inplace= True)
# # Utilizing info() and describe to check the type of Data and to see probable missing values and outliers:
df.info()
df.describe()
# Observations:
# No outliers or missing values noticed.
# # Visualizing the features of the dataset with respect to the target feature inorder visualize the clusters formations
sns.pairplot(df,hue='Species')
# Observations:
# The setosa category seems to be forming a separate cluster throughout while overlapping is observed for other two categories
df.replace(['Iris-setosa','Iris-versicolor','Iris-virginica'],['setosa','versicolor','virginica'],inplace=True)
# # Checking if the categories are balanced or imbalanced using groupby and barplot:
df.groupby('Species').count()
(df['Species'].value_counts()).plot(kind='bar',color = ['red', 'green','blue'])
# # Visualizing the features indivually using barplot
# +
#axs = plt.subplots(1,4)
#sns.barplot(x='Species',y='PetalWidthCm',data=df,ax=axs[0,0])
#sns.barplot(x='Species',y='PetalLengthCm',data=df,ax=axs[0,1])
#sns.barplot(x='Species',y='SepalWidthCm',data=df,ax=axs[0,2])
#sns.barplot(x='Species',y='SepalLengthCm',data=df,ax=axs[0,3])
# -
sns.barplot(x='Species',y='SepalLengthCm',data=df)
sns.barplot(x='Species',y='PetalLengthCm',data=df)
sns.barplot(x='Species',y='SepalWidthCm',data=df)
sns.barplot(x='Species',y='SepalLengthCm',data=df)
# # Observations:
# The length and width of petal and length Sepal seems to be in the incresing from setosa to virginica while incase sepal width Setosa seems to be higher
# # KNN model
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split,cross_val_score
from sklearn.preprocessing import LabelEncoder, StandardScaler
from sklearn.feature_extraction.text import CountVectorizer
# # Encoding the target feature into numbers using label encoding and then converting the feature into categorical data type
df_encoded= df.copy()
label= LabelEncoder()
df_encoded['labels']= label.fit_transform(df['Species'])
print(df_encoded)
list(label.inverse_transform([0, 1, 2]))
df_encoded['labels'].astype('category')
# # Splitting the dataset into dependent (Y) and independent (X) variables
x= df_encoded.drop(columns=['labels','Species','Id'])
y=df_encoded['labels']
#x_scaled= StandardScaler().fit_transform(x)
x_train,x_test,y_train,y_test= train_test_split(x,y,test_size= 0.3,random_state=101)
# # Choosing the value of k by calculating the rate of accuracy using cross-validaion
accuracy=[]
for i in range(1,40):
knn= KNeighborsClassifier(n_neighbors=i)
score= cross_val_score(knn,df_encoded.drop(columns=['Species']),df_encoded['labels'],cv=10)
accuracy.append(score.mean())
plt.figure(figsize=(10,6))
plt.plot(range(1,40),accuracy,color='black',linestyle='dashed',marker='o',markerfacecolor='red',markersize=10)
plt.xlabel('K')
plt.ylabel('accuracy rate')
# # ERROR Rate
# +
error_rate = []
for i in range(1,40):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(x_train,y_train)
pred_i = knn.predict(x_test)
error_rate.append(np.mean(pred_i != y_test))
plt.figure(figsize=(10,6))
plt.plot(range(1,40),error_rate,color='blue', linestyle='dashed',
marker='o',markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
print("Minimum error:-",min(error_rate),"at K =",error_rate.index(min(error_rate)))
# -
# # Accuracy Rate
# +
acc = []
# Will take some time
from sklearn import metrics
for i in range(1,40):
neigh = KNeighborsClassifier(n_neighbors = i).fit(x_train,y_train)
yhat = neigh.predict(x_test)
acc.append(metrics.accuracy_score(y_test, yhat))
plt.figure(figsize=(10,6))
plt.plot(range(1,40),acc,color = 'blue',linestyle='dashed',
marker='o',markerfacecolor='red', markersize=10)
plt.title('accuracy vs. K Value')
plt.xlabel('K')
plt.ylabel('Accuracy')
print("Maximum accuracy:-",max(acc),"at K =",acc.index(max(acc)))
# -
# Observation:
#
# Both error rate and Accuracy rate K=0 as the optimum value
neigh1 = KNeighborsClassifier(n_neighbors = i).fit(x_train,y_train)
pred1 = neigh1.predict(x_test)
print(metrics.confusion_matrix(y_test,pred1))
print(metrics.classification_report(y_test,pred1))
print(x_train)
# # Now lets try and classify a value
def predict(x_input):
neigh2 = KNeighborsClassifier(n_neighbors = 6).fit(x_train,y_train)
pred2 = neigh2.predict(x_input)
if pred2.flatten()==0:
print('Belongs to species : Setosa')
elif pred2.flatten()==1:
print('Belongs to species : Versicolor')
else:
print('Belongs to species : Virginica')
# # for user-input independent features the below can be used:
x_input=[]
for i in ['SepalLengthCm','SepalWidthCm','PetalLengthCm','PetalWidthCm']:
a = float(input(f'Enter the value for {i}:'))
x_input.append(a)
x_input= (np.array(x_input)).reshape(1,-1)
predict(x_input)
# completed
#
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://i.creativecommons.org/l/by-nc-nd/3.0/88x31.png" alt="Drawing" style="width: 150px;"/>
#
#
# **Auteurs originaux** : <NAME> et <NAME> (auteurs du MOOC *Python 3 : des fondamentaux aux concepts avancés du langage* disponible sur la plateforme FUN).
#
# **Modifié et adapté par** : <NAME> (08/09/2019)
# # Rôle de l'informatique et des nouvelles technologies de l'information
# L'informatique occupe une place de plus en plus importante dans nos vies quotidiennes, que ce soit dans son enseignement ou dans la recherche. En effet, du point de vue de notre accés à l'information, les **données** à traiter sont de plus en plus nombreuses et leur analyse requiert une forme de logique, que les ordinateurs et les langages de programmation peuvent très simplement implémenter et traiter.
#
# Qu'il s'agisse de physique, de sociologie, de marketing, ou de finances, la simulation numérique permet de résoudre des problèmes qui seraient partiellement ou totalement inaccessibles en calculant « à la main ».
#
# L'enseignement de l'informatique en tant que science (les anglosaxons parlent de _computer science_) se développe et l'apprentissage d'un langage de programmation est devenu un « passage obligé » dans l'enseignement dès l'entrée au Lycée.
#
# Dans l'état actuel des choses, le langage scientifique le plus répandu en France et dans les pays anglo-saxons est le langage [Python](https://www.python.org/). Son utilisation se développe dès la seconde et il sert, en particulier, de support à l'enseignement de l'informatique en classes préparatoires aux grandes écoles (CPGE) qui sont une voie d'admission à des métiers dans de nombreux domaines.
# # L'informatique dans le cours de SNT à NDO
# Nous avons choisi, dans ce cours, de faire découvrir Python en ciblant quelques utilisations possibles de ce langage en Physique. Nous pensons néanmoins qu'il est important de mettre en évidence l'importance du recours à l'informatique pour développer des modèles qui seront vus ensuite en cours de Physique pendant les traveaux partiques par exemple.
#
# Alors comment va-t-on faire ? Mettons nous dans la peau d'un voyageur qui arriverait dans un pays dont il ne connaît pas la langue. Au début, il ne comprend rien. Il se débrouille, il fait des signes, il finit par identifier des mots ou des phrases qui reviennent sans cesse et dont il finit par _deviner_ le sens (en se trompant parfois).
#
# Ce voyageur sera pris en charge à son arrivée par un professeur qui parle les deux langues, qui prendra soin de te traduire _la plupart_ des mots de la langue étrangère (Python), qui expliquera un peu la grammaire (la syntaxe), qui expliquera les coutumes locales (les bonnes habitudes ou _best practice_) et qui fournira, au cas où, un dictionnaire (la documentation).
# # Jupyter et un portable
# Pour utiliser Python, il faut disposer au moins d'un petit ordinateur portable, voire d'une tablette avec un clavier, voire d'un smartphone. Nous désigneront sous le nom de "portable" dans ce cours, le terminal avec lequel l'élève accéde à l'ordinateur du professeur.
#
# Le choix de l'établissement est d'avoir un programme appellé [edupython](https://edupython.tuxfamily.org/) qui est installé sur les ordinateurs accéssibles dans les salles équipés. Malheureusement nous ne disposons pas d'ordinateurs pour tout le monde et il conviendra donc de s'équiper avec un terminal personnel disposant d'un clavier et d'un accés wifi.
# La solution idéale serait d'installer la distribution Python appelée [anaconda](https://www.anaconda.com/download/), mais cela impose d'avoir beaucoup de place disponible et donc un ordinateur qui n'est pas facile à transporter. Une autre solution, pour des utilisateurs experts, est [miniconda](https://docs.conda.io/en/latest/miniconda.html). Dans tous les cas nous allons travailler avec la version 3 (ou supérieure) de python et chaque élève sera tenu pour résponsable du matériel qu'il apportera au cours de SNT. En cas de perte, ou dégradation dudit matériel, l'enseignant et l'établissement ne peuvent en aucun cas être considérés comme légalement impliqués.
#
# On notera aussi que le fait d'installer un programme n'est pas nécessaire, et comme cela nécessite quelques connaissances cela peut aboutir à quelques difficultés techniques (compatibilité) que nous voulons absolument éviter.
#
# Aussi avons-nous avons choisi d'illustrer le recours au langage Python dans ce cours en utilisant [Jupyter](http://jupyter.org/). Très schématiquement, cela permet de modifier et d'exécuter des scripts Python, **directement dans le navigateur, sans aucune installation sur votre portable**. Cela nous a semblé beaucoup plus simple et motivant. Vous pouvez donc suivre le cours avec n'importe quel portable pourvu qu'il se connecte au réseau local qui sera accéssible pendant le cours.
# # _Notebooks_
# Pour pouvoir utiliser Jupyter, il faut rédiger des documents « mixtes », contenant du texte et du code Python, que l'on appelle des _notebooks_. Ces lignes font partie d'un _notebook_ où l'on entre les définitions et les grand concepts qui seront abordés en cours.
#
# Dans la suite de ce _notebook_, on va utiliser du code Python. Ce code est uniquement destiné à valider le fonctionnement des notebooks et il est très simple.
# ## Avantages des _notebooks_
# <span style="color:red"> Ce support permet un format plus lisible et la première chose à faire sera de renomer le _Notebook_ donné en cours pour le personnaliser, la manière de garder une trace ecrite en pensant que l'on peut la partager avec le reste de la classe. Pour effectuer cette opération le plus simple est d'aller dans l'onglet _file_ puis de choisir _Save as_ et enfin de sauvegarder le cours sous forme : [TITRECOURS-NOM], on observe que le nom d'un fichier ne comporte pas d'espaces (se sont des caractères spéciaux) il doit impérativement contennir le nom de famille.
# Le Titre est donné par le professeur au début de chaque cours. Celui de ce cours est "Methodologie" là encore pas d'accents car se sont des caractères spéciaux.
#
# Il est très important d'être rigoureux en informatique, autrement on peut vite faire de gros dégats.</span>
# <span style="color:red"> Une partie de la note de chaque examen portera sur la rigueur avec laquelle on à tennu compte de ces consignes. Un élève dont le nom de famille est trop long peut demander à le raccourcir, mais il gardera le même raccourci tout au long de l'année. </span>
# J'attire votre attention sur le fait que **les fragments de code peuvent être testés et modifiés** (la page qui s'affiche dans le navigateur n'est pas une page statique : elle est dite _dynamique_). Ainsi il est facile d'essayer des variantes autour du _notebook_ original.
#
# Notez bien également que le code Python est interprété **sur une machine distante** (donc pas sur votre portable), ce qui permet de faire des premiers pas avant même d'avoir procédé à l'installation de Python sur l'ordinateur personnel (c'est tout l'intérêt de la chose).
# ## Comment utiliser les _notebooks_
# En haut du _notebook_, on peut voire une barre, contenant :
# * un titre pour le _notebook_, avec une date entre parenthèses, si une version antérieure doit être consérvée on peut ajouter "_V1" par exemple à la date ;
# * une barre de menus avec les entrées, nous allons nous concentrer sur les entrés du menu qui sont appellées :`File`, `Insert`, `Cell`, `Kernel`;
# * une barre de boutons qui sont des raccourcis vers certains menus fréquemment utilisés. Si tu laisses ta souris au dessus d'un bouton, un petit texte apparaît, indiquant à quelle fonction correspond ce bouton, mais cette aide est souvent donnée en anglais qui est la langue de la pluspart des programmeurs.
#
# Un _notebook_ est constitué d'une suite de cellules, soit textuelles (Markdown), soit contenant du code (Code). Les cellules de code sont facilement reconnaissables, elles sont précédées de
#
# `In [ ]:`
#
# La cellule qui suit celle que tu es en train de lire est une cellule de code.
#
# Pour commencer, sélectionne la cellule de code ci-dessous avec ta souris et appuie dans la barre de boutons sur celui en forme de flèche triangulaire vers la droite (Run).
20 * 3
# En utilisant la fleche qui symbolise la lecture, la cellule est « exécutée » (on dira plus volontiers « évaluée »), et on passe à la cellule suivante.
#
# Alternativement on peux simplement taper au clavier ***Shift+Enter***, ou, selon les claviers, ***Maj-Entrée***, pour obtenir le même effet (<kbd>SHIFT</kbd>+<kbd>ENTER</kbd> ou <kbd>MAJ</kbd>+<kbd>ENTRÉE</kbd>). D'une manière générale, il est important d'apprendre et d'utiliser les raccourcis clavier, cela fera gagner beaucoup de temps par la suite. Et il va de soi que les portables devront avoir un clavier reconnu pour permetre la saisie des notes de cours.
#
# La façon habituelle d'*exécuter* l'ensemble du notebook consiste à partir de la première cellule, et à taper <kbd>SHIFT</kbd>+<kbd>ENTER</kbd> (ou <kbd>MAJ</kbd>+<kbd>ENTRÉE</kbd>) jusqu'au bout du notebook, en n'allant pas trop vite, c'est-à-dire en attendant le résultat de l'exécution de chaque cellule.
# Lorsqu'une cellule de code a été évaluée, Jupyter ajoute sous la cellule `In` une cellule `Out` qui donne le résultat du fragment Python, soit ci-dessus 60.
#
# Jupyter ajoute également un nombre entre les crochets pour afficher, par exemple ci-dessus.
#
# `In [1]:`
#
# Ce nombre te permet de retrouver l'ordre dans lequel les cellules ont été évaluées.
# On peux naturellement modifier ces cellules de code pour faire des essais. Ainsi, il est possible de se servir du modèle ci-dessous pour calculer 10 à la puissance 5
# math.pow (pour power) calcule la puissance
import math
math.pow(10,5)
# En fait, comme sur une calculatrice cette ecriture est semblable (mais plus sophistiquée) à celle la :
1E5
# On remarque d'emblée que python aime bien donner des chifres avec une précision décimale. Il faudra en tenir compte dans certaines opérations simples comme les rapports.
# Il peut être parfois utile d'évaluer tout le _notebook_ en une seule fois en utilisant le menu *Cell -> Run All*.
# ## Attention à bien évaluer les cellules dans l'ordre
# Il est important que les cellules de code soient évaluées dans le bon ordre. Si on ne respecte pas l'ordre dans lequel les cellules de code sont présentées, le résultat peut être inattendu : C'est comme mettre la charue avant les bœufs
#
# En fait, évaluer un programme sous forme de notebook revient à le découper en petits fragments. Si on exécute ces fragments dans le désordre, on obtient naturellement un programme différent.
# On le voit sur cet exemple :
message = "Il faut faire attention à l'ordre dans lequel on évalue les notebooks"
print(message)
# Si un peu plus loin dans le notebook on fait par exemple :
# ceci a pour effet d'effacer la variable 'message'
del message
# qui rend le symbole `message` indéfini, alors, bien sûr, tu ne peux plus évaluer la cellule qui fait `print` puisque la variable `message` n'est plus connue de l'interpréteur.
# ## Réinitialiser l'interpréteur
# Si trop de modifications sont faites, ou si le fil de ce qui doit être évalué est perdu, il peut être utile de redémarrer l'interpréteur. Le menu *Kernel → Restart* permet de faire cela.
# Le menu *Kernel → Interrupt* peut être quant à lui utilisé si le fragment prend trop longtemps à s'exécuter (par exemple quand on écrit une boucle dont la logique est cassée et qui ne termine pas). Nous verons comment éviter ses erreurs et dans la mesure ou nous progresseront ces erreurs seront mieux gérés (elles ne sont pas problématique car on peut toujours executer son notebook sur un serveur distant: par exemple [jupyter.org/try](https://jupyter.org/try)
# ## Il faut travailler sur sa copie personnelle
# Un des avantages principaux des _notebooks_ est de permettre de modifier le code écrit par le professeur et de voir, par soi-même, comment se comporte le code modifié.
#
# Pour cette raison, chaque participant dispose de sa **propre copie** de chaque _notebook_. Dans la première séance nous verrons comment ces copies apparaissent au fûr et a mesure. Apporter toutes les modifications, souhaitables aux notebooks sans affecter les autres étudiants, est une attitude qui sera très bien valorisée par le professeur.
# ## Revenir à la version du cours
# On peut toujours faire le point avant de changer la version « du cours » grâce au menu
# *File → Save and Checkpoint*.
# Ensuite *File → Revert to Checkpoint* permet de retrouver la version qui à été mise de côté.
# Attention, avec cette fonction, on restaures **tout le _notebook_** et donc **on perds tes modifications que tu as faites sur ce _notebook_**.
# ## Télécharger au format Python
# On peux télécharger un _notebook_ au format Python sur l'ordinateur grâce au menu
# *File → Download as → Python*
# Les cellules de texte sont préservées dans le résultat sous forme de commentaires Python.
# ## Partager un _notebook_ en lecture seule
# Enfin, avec le menu *File → Share static version*, on peux publier une version en lecture seule de ton _notebook_ ; tu obtiens une URL que tu peux publier par exemple pour demander de l'aide sur le forum. Ainsi, les autres participants peuvent accéder en lecture seule à ton code.
# ## Ajouter des cellules
# On peux ajouter une cellule n'importe où dans le document avec le bouton **+** de la barre de boutons.
#
# Lorsque l'on arrive à la fin du _notebook_, une nouvelle cellule est créée chaque fois que l'on évalue la dernière cellule. De cette façon, on dispose d'un brouillon pour ses propres essais.
# # En cas de problème...
# Encore une chose... Si tout ne se passe pas comme prévu ou si tu rencontres des difficultés, n'hésite pas à en parler au professeur, voire même à poser la question sur un forum
#
# Attention néanmoins... Pour que les participants du forum (et à plus forte raison le professeur) puissent t'aider à résoudre ton problème, il faut qu'il soit décrit correctement, d'abord avec des mots (on évite des messages du style « Ça marche pas. ») et éventuellement avec des images (parfois, cela vaut mieux qu'un long discours).
#
# Aussi, il faut être le plus précis possible dans la description de ton problème et à joindre une ou plusieures captures d'écran qui te permette de l'illustrer.
# ## Un exercice interessant
# +
x=-80538738812075974
y=80435758145817515
z=12602123297335631
x3=pow(x,3)
y3=pow(y,3)
z3=pow(z,3)
print("x au cube = {} ".format(x3))
print("y au cube = {} ".format(y3))
print("z au cube = {} ".format(z3))
# -
x3+y3+z3
|
Methodologie-DILLMANN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><img src="http://i.imgur.com/sSaOozN.png" width="500"></center>
# ## Course: Computational Thinking for Governance Analytics
#
# ### Prof. <NAME>, PhD
# * Visiting Professor of Computational Policy at Evans School of Public Policy and Governance, and eScience Institute Senior Data Science Fellow, University of Washington.
# * Professor of Government and Political Methodology, Pontificia Universidad Católica del Perú.
#
# _____
#
# # Session 1: Programming Fundamentals
#
# ## Part A: Data Structures in Python
# <a id='beginning'></a>
# Programming languages use data structures to tell the computer how to organize the data we are working with. That is, data structures provided by a programming language are not the same in another one. However, in most cases, a name given to a data structure in one programming language should generally be the same in other one. It is worth keeping in mind, that a particular data structure may serve for one purpose, but not for other ones.
#
# In everyday life, a book can be considered a data structure: we use it to store some kind of information. It has some advantages: it has a table of contents; it has numbers on the pages; you can take it with you; read it as long as you can see the words; and read it again as many times as you want. It has some disadvantages: you can lose it, and need to buy it again; it can deteriorate; get eaten by an insect; and so on.
#
# We are going to talk about 3 data structures in Python:
#
#
# 1. [List](#part1)
# 2. [Tuple](#part2)
# 3. [Dictionary](#part3)
# 4. [Data Frame](#part4)
#
# **Lists** and **tuples** are basic containers, while **dictionaries** (a.k.a **dicts**) could be considered less simple and with a different 'philosophy'. **Data frames** are complex structures not directly supported by base Python, but easily managed with an additional package.
# ____
# <a id='part1'></a>
#
# ## List
# Lists in Python are containers of values as in **R**. The values can be of any kind (numbers or non-numbers), and even other containers (simple or complex). If we have an spreadsheet as a reference, a row is a 'natural' list. Different from R, you can not give names to the list elements.
DetailStudent=["<NAME>",40,"False"]
# [ ] creates a list, = is assignment
# The *object* 'DetailStudent' serves to store temporarily the list. To name a list, use combinations of letters and numbers (never start with a number) in a meaningful way. Typing the name of the object, now a list, will give you all the contents you saved in there:
DetailStudent
# Python's lists are similar to vectors in R, but Python does not coerce the values (40 is still a number). Lists in Python are so flexible and simple, that it is common to have nested lists:
DetailStudentb=['<NAME>',60,'True']
Classroom=[DetailStudent,DetailStudentb] # list of lists
Classroom
# You can access individual elements like this:
Classroom[1]
# From the last result, you must always remember that Python positions start in **0**, see more examples of accessing:
DetailStudentb[0] # first element
DetailStudentb[:2] # before the index 2, that is position 0 and 1 / In R: DetailStudentb[1:2] (both limits needed)
# : means before, 2 is the index
DetailStudent[-1] # R does not work like this to get you the last element of a list...This will erase the first one
#[] means position, -1 means to give me the last element of the list
# You can alter lists like in R (just remember positions start from 0 in Python):
DetailStudent[0]='<NAME>'
DetailStudent
#altering the first element to be Alfred rather than Fred
#Makes changes without "warning" so you need to be careful, pay attention
# Deleting elements is easy, and we can do it:
#
# * By position
# * By value
#
# Let's see. If we have these lists:
elementsA=[1,2,3,4]
elementsB=[1,2,3,4]
# Then:
## DELETING BY POSITION
del elementsA[2] #delete third element
# then:
elementsA # alternative: elements[:2]+elements[3:]
# DELETING BY VALUE
elementsB.remove(2)
elementsB
# Getting rid of your list:
# +
newList=['a','b']
del newList
newList # becareful!... it is gone!
#Python points out the line in which there is an error! this is really nice
# -
# It is important to know how to get **unique values**:
weekdays=['M','T','W','Th','S','Su','Su']
weekdays
#then:
weekdays=list(set(weekdays))
weekdays
#set gives us the unique values
# ### Doesn't Python have vectors?
# Vectors are NOT part of the basic Python, you need to use a mathematical module like **numpy**. When working with vectors, the operations of comparison ('>', '<', etc.) will work **element by element** as in R:
# +
# For Python to work as R with vectors, you need to use the
# mathematical structure offered by numpy:
import numpy as np
#import is same as library in R - means to "activate" the thing that is on your computer already
#np.array means "array" is a function in np - the name we gave to numpy when we imported it
vector1=np.array(['b','c','d'])
vector2=np.array(['a','b','d'])
vector1>vector2
#numpy is how you do algebra, math, stats - R is ready to go because it was designed for use, Python needs a bit more coaxing
# -
# If vectors have different sizes, comparison works if one has ONE element:
vector3=np.array(['a'])
vector1>vector3 # each element of vector1 compared to the only one in vector3
#not very important to learn
vector3
# But, this confuses vectors:
vector4=np.array(['a','b'])
vector1>vector4
# This is also valid for numbers:
# If these are our vectors:
numbers1=np.array([1,2,3])
numbers2=np.array([1,2,3])
numbers3=np.array([1])
numbers4=np.array([10,12])
# Then, these work well:
# adding element by element:
numbers1+numbers2
# adding one value to all the elements of other vector:
numbers1+numbers3
# multiplication (element by element)!
numbers1*numbers2
# and this kind of multiplication:
numbers1*3
# This will not work (it does not work in R either):
numbers1+numbers4
# When dealing with vectors, the elements must share the same type. Otherwise, elements will be coerced into the same type:
numbers5=np.array([1,2,'3'])
numbers5
numbers6=np.array([1,2,3.0])
numbers6
# [Go to page beginning](#beginning)
# _____
# <a id='part2'></a>
#
# ## Tuples
# Tuples are similar to lists. They can store any kind value, and even other structures:
DetailStudentaTuple=("<NAME>",40,"False")
#parantheses create the tuple
# To create tuples, you can use '()', the command *tuple()* or nothing:
DetailStudentbTuple='<NAME>',60,'True'
#by default Python creates tuples
# So, **why do we need *tuples*?** When you do not want that your object be altered:
DetailStudentbTuple[1]=50
# [Go to page beginning](#beginning)
# ____
# <a id='part3'></a>
# ## Dicts
# Dicts, on the surface, are very similar to lists in R:
# creating dict:
DetailStudentDict={'fullName':"<NAME>",
'age':40,
'female':False}
# seeing it:
DetailStudentDict
# But you realize soon a difference:
DetailStudentDict[0]
# Dicts _only_ use their **keys** to access the elements:
DetailStudentDict['age']
# Dicts do allow changing values:
DetailStudentDict['age']=41
# then:
DetailStudentDict['age']
# ## Lists versus Tuples vs Dicts?
# __A) Make sure what you have:__
#
# You can easily know what structure you have like this:
type(DetailStudentDict)
type(DetailStudent)
type(DetailStudentaTuple)
# __B) Make sure functions are shareable__
#
# They share many basic functions:
listTest=[1,2,3,3]
tupleTest=(1,2,3,4,4)
dictTest={'a':1,'b':2,'c':2}
len(listTest), len(tupleTest), len(dictTest)
# Some may work slightly different:
# using set to keep unique values:
set(listTest)
set(tupleTest) # so far so good...
set(dictTest) # this MAY not be what you expected.
#The set for a dictionary lists the "headers" rather than the values
# Notice the use of comparissons between lists and vectors:
numbers4=np.array([2])
numbers1<numbers4
# This will work the same for text:
list1=np.array(['b','c','d'])
list2=np.array(['a','b','d'])
list1>list2
# If we used lists, you get a similar bahavior (not implemented in base R):
list1=['b','c','d']
list2=['a','b','d']
list1>list2
# Python is doing a simple _lexicographical ordering_, that is, they compare the first element of each list (from left to right), and report _True_ or _False_ if they differ using '>' (or '<'). It is like comparing two words:
np.array([1,2,4]) > np.array([1,2,3]) # this is true because 4>3, and the previous are equal.
[1,2,4] > [1,2,3]
# this is true because 9>8, and the previous are equal, when a difference is detected, the comparisson stops.
(1,2,9,1) > (1,2,8,9,9)
# while you can not compare if sizes differ:
np.array([1,2,9,1]) > np.array([1,2,8,9,9])
# Math operations should be taken with care:
# +
# This will CONCATENATE:
numbersL1=[1,2,3]
numbersL2=[1,2,3]
numbersL1+numbersL2
#You're not adding the values, you are concatenating the two lists
# -
# this won't work:
numbersL1 * numbersL2
# this will:
numbersL1 * 3
# Due to its flexibility, lists are used pervasively in simple Python code.
# [Go to page beginning](#beginning)
# ____
# <a id='part4'></a>
# ## Data Frames
# Data frames are containers of values. The most common analogy is an spreadsheet. To create a data frame, we need to call **pandas**:
import pandas
# We can prepare the data frame now:
# columns of the data frame (as lists):
names=["Qing", "Françoise", "Raúl", "Bjork"]
ages=[32,33,28,30]
country=["China", "Senegal", "Spain", "Norway"]
education=["Bach", "Bach", "Master", "PhD"]
# now in a dict:
data={'names':names, 'ages':ages, 'country':country, 'education':education}
data
# ...and from dict to DataFrame:
students=pandas.DataFrame.from_dict(data)
# seeing it:
students
# Sometimes, Python users code like this:
# +
import pandas as pd # renaming the library
students=pd.DataFrame.from_dict(data)
students
# -
# Or like this:
# +
from pandas import DataFrame as df # calling a function from the library and renaming the function name
students=df.from_dict(data)
students
# -
# You can set a particular column as **row name**:
students.set_index('names') # You have not changed until: students.set_index('names',inplace=True)
# The command *type()* still works here:
type(students)
# You can get more information on the data types like this (as _str()_ in R):
students.dtypes
# The _info()_ function can get you more details:
students.info()
# The data frames in pandas behave much like in R:
#one particular column
students.names
# or
students['names'] # it is not the same as: students[['names']]
# it is not the same as:
students[['names']] # a data frame, not a column (or series)
# two columns
students.iloc[:,[1,3]]
# thie is also a DF
students[['country','names']]
## Using positions is the best way to get several columns:
students.iloc[:,1:4]
# Deleting a column:
# +
# This is what you want get rid of:
byeColumns=['education']
#this would change the original: students.drop(byeColumns,axis=1,inplace=False)
studentsNoEd=students.drop(byeColumns,axis=1)
# this is a new DF
studentsNoEd
# -
# You can modify any values in a data frame. Let me create a **deep** copy of this data frame to play with:
studentsCopy=students.copy()
studentsCopy
# Then,
# I can change the age of Qing to 23 replacing 32:
studentsCopy.iloc[0,0]=23 # change is immediate! (no warning)
#[row,column]
# I can reset a column as **missing**:
studentsCopy.country=None
# And, delete a column by droping it:
studentsCopy.drop(['ages'],1,inplace=True) # axis=1 is column
# Then, our copy looks like this:
studentsCopy
# One important detail when erasing rows, is to reset the indexes:
# another copy for you to see the difference:
studentsCopy2=students.copy()
studentsCopy2
# drop third row (axis=0)
studentsCopy2.drop(2)
# resetting index
studentsCopy2.drop(2).reset_index()
#better resetting index
studentsCopy2.drop(2).reset_index(drop=True)
# Pandas offers some practical functions:
# rows and columns
students.shape # dim(meals) in R
# length:
len(students) # length in R gives number of columns, here you get number of rows.
# There is no specific function to get number of rows/columns in pandas, but **len** is useful:
len(students.index) # or students.shape[0]
len(students.columns) # or students.shape[1]
# Remember that you can use len with list, tuples and data frames!...and even dictionaries (notice it gives you the count at the top level, it is not smart to report the count inside of an composite element).
aDict={'name':'John', "language_spoken":['Spanish','English']}
len(aDict)
# You also have _tail_ and _head_ functions in Pandas, to get some top or bottom rows:
students.head(2) #and students.tail(2)
# You can also see the column names like this:
# similar to names() in R
students.columns
# It may look like a list, but it is not:
type(students.columns) # index type...but list functions work here!
# If you needed a list:
# +
students.columns.values.tolist()
# or:
# students.columns.tolist()
# this is the easiest:
# list(students)
# -
# ### Querying Data Frames:
# Once you have a data frame you can start writing interesting queries:
# Who is the oldest in the group?
students[students.ages==max(students.ages)].names
#Within the students dataframe, look at the ages column, find the max of the ages, and give me the name
# Who is above 30 and from China?
students[(students.ages>30) & (students.country=='China')] # parenthesis are important with '&' in Pandas!!!
#Within students DF, who is above 30 and from China - gives the entire row
# Who is not from Norway?
students[students.country!="Norway"]
# +
# Who is from one of these?
DangeourousPlaces=["Peru", "USA", "Spain"]
students[students.country.isin(DangeourousPlaces)]
#who has a country that is within the DangerousPlaces list
# -
students[~students.country.isin(DangeourousPlaces)] # the opposite
#tild ~ gives you "not"
# The education level of who is above 30 and from China?
students[(students.ages>30) & (students.country=='China')].education
# **Show me the data ordered by age (decreasing)?**
toSort=["ages"]
Order=[False]
students.sort_values(by=toSort,ascending=Order)
# Show who is the oldest person with a Bachelor:
students[students.education=='Bach'].sort_values('ages',ascending=True).tail(1)
#Among students who have BA, sort by ages in increasing value, and then gives the "tail" - who ever is in the furthest down row
# ## Class exercises:
# In a new Jupyter notebook solve each excercise, and then upload them to GitHub. Name the notebook as 'ex_data_structures':
# A. Turn this into a Data Frame name "friends":
names=["Tomás", "Pauline", "Pablo", "Bjork","Alan","Juana"]
woman=[False,True,False,False,False,True]
ages=[32,33,28,30,32,27]
country=["Chile", "Senegal", "Spain", "Norway","Peru","Peru"]
education=["Bach", "Bach", "Master", "PhD","Bach","Master"]
# B. Answer the following:
# +
# Who is the oldest person in this group of friends?
# +
# How many people are 32?
# +
# How many are not Peruvian? (use two different codes)
# +
# Who is the person with the highest level of education?
# +
# what is the sex of the oldest person in the group?
# -
# ### Homework
# If you have the query:
# +
# where is the youngest male in the group from?
# -
# a. Find the answer using *sort_values()*
#
# b. Do some research and find the answer using *[where()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.where.html)* and *[min()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.min.html)*
#
# c. Do some research and find the answer using *[query()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.query.html)* and *[min()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.min.html)*
# Solve this in a new Jupyter notebook, and then upload it to GitHub. Name the notebook as 'hw_data_structures'.
# ____
#
# * [Go to page beginning](#beginning)
# * [Go to REPO in Github](https://github.com/EvansDataScience/ComputationalThinking_Gov_1)
# * [Go to Course schedule](https://evansdatascience.github.io/GovernanceAnalytics/)
|
S1_A_Py_dataStructures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import autoreg
import GPy
import numpy as np
import scipy.io
from pylab import *
# %matplotlib inline
def comp_RMSE(a,b):
return np.sqrt(np.square(a-b).mean())
data = scipy.io.loadmat('./identificationExample.mat')
data_in = data['u'][:,None]
data_out = data['y'][:,None]
win_in = int(data['lu'])
win_out = int(data['ly'])
data_in_train = data_in[:150]
data_out_train = data_out[:150]
data_in_test = data_in[150:]
data_out_test = data_out[150:]
# One hidden layer
m = autoreg.DeepAutoreg([0,win_out],data_out_train, U=data_in_train, U_win=win_in,X_variance=0.01,
num_inducing=50)
m.layer_0.likelihood.variance[:] = data_out_train.var()*0.01
m.layer_1.likelihood.variance[:] = 0.01
m.layer_0.likelihood.fix(warning=False)
m.layer_1.likelihood.fix(warning=False)
print m
m.optimize(messages=1,max_iters=50)
m.layer_0.likelihood.constrain_positive(warning=False)
m.layer_1.likelihood.constrain_positive(warning=False)
m.optimize(messages=1,max_iters=1000)
m.layer_1.kern.lengthscale
m.layer_0.kern.lengthscale
pd = m.freerun(U=data_in_test)
_=plot(pd,'b',label='pred')
_=plot(data_out_test[1:],'r',label='ground-truth')
legend(loc='best')
print comp_RMSE(pd,data_out_test[1:])
|
examples/example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Question Detector Notebook Contents
# - [How can I create a QuestionDetector?](#How-can-I-create-a-QuestionDetector?)
# - [So how does the QuestionDetector work? ](#So-how-does-the-QuestionDetector-work?)
# - [What is the difference between IssueQuestion vs EmailQuestion vs IssueCommentQuestion? ](#What-is-the-difference-between-IssueQuestion-vs-EmailQuestion-vs-IssueCommentQuestion?)
# - [What is this context attribute I'm seeing?](#What-is-this-context-attribute-I'm-seeing-in-the-Question-objects?)
# - [Can I use the QuestionDetector for my projects that aren't issue/email/comment related?](#Can-I-use-the-QuestionDetector-for-my-projects-that-aren't-issue/email/comment-related?)
# ## How can I create a QuestionDetector?
# Donkeybot's `QuestionDetector` must be one of the following types : "email", "issue" or "comment"
# This is so that the `QuestionDetector` creates the correct type of Question objects.
# Be it an `EmailQuestion`, `IssueQuestion`, `CommentQuestion`.
# Let's create one for `IssueQuestions`!
from bot.question.detector import QuestionDetector
detector = QuestionDetector("issue")
text = """
What is this 'text', you ask?
Well, it's a monologue I'm having... can it help with something you still ask?
In testing the QuesitonDetector of course!
Did that answer all your questions?
I sure hope so..."
"""
# ## So how does the QuestionDetector work?
# Simply use the .detect() method!
# The results are going to be a list of `Question` objects.
# In this specific example `IssueQuestion` objects.
results = detector.detect(text)
results
# And all 3 questions from the sample text above have been identified!
[(question.question) for question in results]
# ## What is the difference between `IssueQuestion` vs `EmailQuestion` vs `IssueCommentQuestion`?
# The only difference is their `origin` and how they get their `context` attributes.
results[1].__dict__
# ## What is this `context` attribute I'm seeing in the Question objects?
# Well, that's what the AnswerDetector uses to try and answer each question!
# ~To be more specific~
#
# 1) When a new User Question is asked and is very similar or identical to the questions archived by using the .detect() method.
# 2) Then the context of these archived questions is used as context for the new User Question.
# 3) Donkeybot's AnswerDetector tries to find suitable answers!
# For `IssueQuestions` the context are any comments that are part of the same GitHub issue.
# For `IssueCommentQuestion` the context are comments after this specific one where the Question was detected.
# For `EmailQuestions` the context are the bodies of the reply emails to the email where the Question was detected.
# Each different Question object has it's own unique `find_context_from_table()`
# method that sets the attribute by following the logic explained above.
#
# Basically go into the table in our Data Storage and SELECT the context we want.
# ## Can I use the QuestionDetector for my projects that aren't issue/email/comment related?
# Yes!
#
# But, if you aren't following the issue, email, comment logic Donkeybot follows at the point of writing this.
# (end of GSoC '20').
#
# Then, Donkeybot needs to be expanded to have a `Question` superclass and a `set_contexT()` method fo you to simple set the context without going into some dependand Data Storage.
#
# If you want to see this in Donkeybot [open an issue](https://github.com/rucio/donkeybot/issues) and suggest it.
# I'll see that you've been reading the documentation and that this functionality is needed :D
#
#
|
examples/question_detector.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <a href="https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv" target="_blank"><img align="left" src="data/cover.jpg" style="width: 76px; height: 100px; background: white; padding: 1px; border: 1px solid black; margin-right:10px;"></a>
# *This notebook contains an excerpt from the book [Machine Learning for OpenCV](https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv) by <NAME>.
# The code is released under the [MIT license](https://opensource.org/licenses/MIT),
# and is available on [GitHub](https://github.com/mbeyeler/opencv-machine-learning).*
#
# *Note that this excerpt contains only the raw code - the book is rich with additional explanations and illustrations.
# If you find this content useful, please consider supporting the work by
# [buying the book](https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv)!*
# <!--NAVIGATION-->
# < [Using Decision Trees to Diagnose Breast Cancer](05.02-Using-Decision-Trees-to-Diagnose-Breast-Cancer.ipynb) | [Contents](../README.md) | [Detecting Pedestrians with Support Vector Machines](06.00-Detecting-Pedestrians-with-Support-Vector-Machines.ipynb) >
# # Using Decision Trees for Regression
#
# Although we have so far focused on using decision trees in classification tasks, you can also use them for regression. But, you will need to use Scikit-Learn again, as OpenCV does not provide this flexibility. We therefore only briefly review its functionality here.
#
# Let's say we wanted to use a decision tree to fit a sine wave. To make things interesting, we will also add some noise to the data points using NumPy's random number generator:
import numpy as np
rng = np.random.RandomState(42)
# We then create 100 x values between 0 and 5, and calculate the corresponding sine values:
X = np.sort(5 * rng.rand(100, 1), axis=0)
y = np.sin(X).ravel()
# We then add noise to every other data point in y (using y[::2]), scaled by 0.5 so we don't introduce to much jitter:
y[::2] += 0.5 * (0.5 - rng.rand(50))
# You can then create regression tree like any other tree before. We will build two trees, one with a depth of 2, and one with a depth of 5:
from sklearn import tree
regr1 = tree.DecisionTreeRegressor(max_depth=2, random_state=42)
regr1.fit(X, y)
regr2 = tree.DecisionTreeRegressor(max_depth=5, random_state=42)
regr2.fit(X, y)
# We can then use the decision tree like a linear regressor from Chapter 3, First Steps in Supervised Learning. For this we create a test set with x values densely sampled in the whole range from 0 through 5:
X_test = np.arange(0.0, 5.0, 0.01)[:, np.newaxis]
# The predicted y values can then be obtained with the predict method:
y_1 = regr1.predict(X_test)
y_2 = regr2.predict(X_test)
# If we plot all of these together, we can see how the decision trees differ:
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
plt.figure(figsize=(10, 6))
plt.scatter(X, y, c='k', s=50, label='data')
plt.plot(X_test, y_1, label="max_depth=2", linewidth=5)
plt.plot(X_test, y_2, label="max_depth=5", linewidth=3)
plt.xlabel("data")
plt.ylabel("target")
plt.legend()
# -
# <!--NAVIGATION-->
# < [Using Decision Trees to Diagnose Breast Cancer](05.02-Using-Decision-Trees-to-Diagnose-Breast-Cancer.ipynb) | [Contents](../README.md) | [Detecting Pedestrians with Support Vector Machines](06.00-Detecting-Pedestrians-with-Support-Vector-Machines.ipynb) >
|
notebooks/05.03-Using-Decision-Trees-for-Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Extract samples of gambiae and coluzzii from sequencing dataset used in Sample ID, align and convert to zarr genotype calls relative to PEST genome. Also subset to Ag1000g biallelic positions.
#
# Results:
# - sequences converted to genotype calls & nalts, subsetted and visualised
# - lots of variants are lost with subsetting to Ag1000g biallelic sites. These mostly come from large inaccessible regions, but novel variants in sequencing are quite common as well.
#
#
#
# Limitations:
# - not accounting for various ALTs - all substitutions get 1, all deletions get -1 (missing), insertions are ignored.
# - ignoring multiallelic calls
# - manual POS margin adjustment to fit with Ag1000g records
#
# TODO: - in future conversions, check REF/ALT for a) ALT recoding b) POS consistenct checks
import os
import sys
import zarr
import allel
import numpy as np
import pandas as pd
from Bio import AlignIO
import matplotlib.pyplot as plt
# +
#in
# sequencing data and metadata used for Species ID
SEQ_FILE = '../7_species_id/data/0_haplotypes.csv'
SEQ_META_FILE = '../7_species_id/data/0_samples.csv'
# reference data and metadata
REF_FILE = '../4_ref_extraction/data/refs.csv'
REF_META_FILE = '../4_ref_extraction/data/samples_ref.csv'
# panel metadata - for positions
PANEL = '../../data/panel_info.csv'
# out
WD = '../../../data/phylo_ampl_ag1k/seq_samples'
ALN_FA = os.path.join(WD, '{}.fa')
ZARR = os.path.join(WD, 'seq_samples.zarr')
SPP_META = (os.path.join(WD, 'seq_samples.meta.csv'))
# ag1000g zarr for positions extraction
AG1K_ZARR = os.path.join(WD, '../phase2/AR1')
AG1K_META = os.path.join(WD, '../phase2/AR1/samples/samples.meta.txt')
# params
# species to extract
SPP = ['Anopheles_gambiae','Anopheles_coluzzii']
# amplicons
AMPLS = [str(i) for i in range(62)]
# reference genome sample name - used as REF allele state
REF_SAMPLE = 'anopheles-gambiae-pestchromosomesagamp4'
# -
# ! mkdir -p {WD}
panel = pd.read_csv(PANEL, index_col='amplicon')
panel.head()
# ## Read and combine sequencing and reference data
all_seq = pd.read_csv(SEQ_FILE, dtype=str)
all_meta = pd.read_csv(SEQ_META_FILE, dtype=str)
all_ref_seq = pd.read_csv(REF_FILE, dtype=str)
all_ref_meta = pd.read_csv(REF_META_FILE, dtype=str)
comb_seq = pd.concat([all_seq, all_ref_seq], sort=False)
comb_seq.head(1)
comb_meta = pd.concat([all_meta, all_ref_meta], sort=False)
comb_meta.head(1)
# ## Subset to species of interest
spp_meta = comb_meta[comb_meta.Species.isin(SPP)].copy()
spp_meta
spp_seq = comb_seq[comb_seq.s_Sample.isin(spp_meta.s_Sample)].copy()
spp_seq.shape
# assign IDs to unique sequences
combuids = dict()
for tgt, group in spp_seq.groupby(['target']):
for (i, cons) in enumerate(group['consensus'].unique()):
combuids[tgt+cons] = '{}-{}'.format(tgt, i)
spp_seq['combUID'] = (spp_seq.target + spp_seq.consensus).replace(combuids)
spp_seq.sample(3)
# ## Ag1k compatible metadata
spp_meta.rename(
{'s_Sample':'ox_code',
'Source':'population',
'Country':'country',
'Specimen Sex':'sex'},
axis=1,
inplace=True)
# population labels
spp_meta['pop_label'] = spp_meta.population.replace(pop_labels)
spp_meta['m_s'] = spp_meta.Species.replace(
{'Anopheles_gambiae':'S',
'Anopheles_coluzzii':'M'})
spp_meta = spp_meta[['ox_code','country','population','sex','m_s','pop_label']]
spp_meta.head()
# write meta
spp_meta.to_csv(SPP_META, index=False)
# ! head -2 {SPP_META}
# ## Align each amplicon
# generate alignments
for ampl in AMPLS:
sys.stdout.write('\r' + ampl)
out_fa = ALN_FA.format(ampl)
# do not re-generate existing alignments
if os.path.isfile(out_fa):
continue
# subset amplicon data
ampl_data = spp_seq[spp_seq.target == ampl]
# get only unique sequences
ampl_data = ampl_data[~ampl_data.combUID.duplicated()]
# write under combUID name
with open('temp.fa', 'w') as o:
for (i, row) in ampl_data.iterrows():
o.write('>{}\n{}\n'.format(row.combUID,
row.consensus))
# align
# ! mafft temp.fa > {out_fa} 2> /dev/null
# ! rm temp.fa
print('\nDone!')
# add alignments to sequence table
aln_dict = {}
for ampl in AMPLS:
aln = AlignIO.read(ALN_FA.format(ampl), 'fasta')
ampl_aln_dict = {seq.name:str(seq.seq) for seq in aln}
aln_dict.update(ampl_aln_dict)
spp_seq['aln'] = spp_seq.combUID.replace(aln_dict)
spp_seq.sample(3)
# ## Covert, subset & store
# +
def call_haploid(seq, ref_seq):
'''
Simple genotype inference relative to reference:
match - 0,
mismatch - 1
indel - -1
Not vectorised
Not accounting for various ALT's
'''
assert len(seq) == len(ref_seq), 'sequences unaligned'
gt = []
for c, ref_c in zip(seq, ref_seq):
# skip insertions
if ref_c == '-':
continue
# deletion - missing data
if c == '-':
gt.append(-1)
# match
elif c == ref_c:
gt.append(0)
# mismatch
else:
gt.append(1)
return gt
call_haploid('a-tt','ac-c')
# +
# mandatory pre-clean
# ! rm -r {ZARR}
for ampl in AMPLS:
sys.stdout.write('\r' + ampl)
# amplicon sequence data
ampl_seq = spp_seq[spp_seq.target == ampl].copy()
# reference alignment
ampl_ref_aln = ampl_seq.loc[ampl_seq.s_Sample == REF_SAMPLE, 'aln']
assert ampl_ref_aln.shape[0] == 1, 'non-unique or missing REF seq in {}'.format(ampl)
ampl_ref_aln = ampl_ref_aln.iloc[0]
# gapless ref length
ampl_ref_len = len(ampl_ref_aln.replace('-',''))
# convert haploid genotypes
ampl_seq['gt'] = ampl_seq.aln.apply(lambda x: call_haploid(x, ampl_ref_aln))
# diploid sample genotypes
# output gt matrix axes samples:sites:ploidy
ampl_gt = []
# order of samples as in metadata
for sample in spp_meta.ox_code:
sample_ampl_gt = ampl_seq.loc[ampl_seq.s_Sample == sample, 'gt']
# missing
if sample_ampl_gt.shape[0] == 0:
diploid_gt = [(-1,-1) for i in range(ampl_ref_len)]
# homozygous
elif sample_ampl_gt.shape[0] == 1:
diploid_gt = [(x,x) for x in sample_ampl_gt.iloc[0]]
# heterozygous - take first two alleles
else:
diploid_gt = [(x,y) for x,y in zip(sample_ampl_gt.iloc[0], sample_ampl_gt.iloc[1])]
if sample_ampl_gt.shape[0] > 2:
print('Warning: more than two alleles in sample {} amplicon {}'.format(sample, ampl))
ampl_gt.append(diploid_gt)
# reshape to sites:samples:ploidy
ampl_gt = np.swapaxes(ampl_gt, 0, 1)
# get amplicon coordinates in PEST
# NB corrected to be compatible with Ag1k coordinates
ampl_pos = np.arange(panel.loc[ampl,'start_insert'] + 1, panel.loc[ampl,'end_insert'] + 1)
# get positions of ag1k biallelic sites
ag_root = zarr.open_group(AG1K_ZARR, "r")
ag_ampl_pos = ag_root[ampl]['POS'][:]
# mask sequenced sample positions
mask = np.in1d(ampl_pos, ag_ampl_pos)
# subset along axis 0 (variants)
agpos_ampl_gt = ampl_gt[mask,:,:]
assert ag_ampl_pos.shape[0] == agpos_ampl_gt.shape[0], 'Position subsetting failed in {}'.format(ampl)
# compute nalts
agpos_ampl_nalt = allel.GenotypeArray(agpos_ampl_gt).to_n_alt()
# write
root = zarr.open_group(ZARR, "a")
g = root.create_group(ampl)
g.create_dataset('genotype', data=agpos_ampl_gt)
g.create_dataset('NALT', data=agpos_ampl_nalt)
ampl_nalt = allel.GenotypeArray(ampl_gt).to_n_alt()
g.create_dataset('raw_genotype', data=ampl_gt)
g.create_dataset('raw_nalt', data=ampl_nalt)
print('\nDone!')
# -
# ## Plot variant/invariant sites
root = zarr.open_group(ZARR, "r")
fig, ax = plt.subplots(1,1, figsize=(15,4))
# how many variants in each amplicon?
for ampl in AMPLS:
raw_isvar = np.any((root[ampl]['raw_nalt'][:] > 0), axis=1) # not considering all ALT - PEST is all REF
isvar = np.any((root[ampl]['NALT'][:] > 0), axis=1) # not considering all ALT - PEST is all REF
p0 = ax.bar(ampl, isvar.shape[0], color='b')
p1 = ax.bar(ampl, raw_isvar.sum(), color='g')
p2 = ax.bar(ampl, isvar.sum(), color='r')
plt.legend((p0[0], p1[0], p2[0], ),
('Total Ag1000g biallelic sites in amplicon',
'Total variants in seq',
'Ag1000g biallelic variants in seq',))
ax.set_title('Variant filtering by Ag1000g biallelic positions');
# compare to selection of samples samples of ag1k
root = zarr.open_group(AG1K_ZARR, "r")
fig, ax = plt.subplots(1,1, figsize=(15,4))
# how many variants in each amplicon?
for ampl in AMPLS:
sampleset = slice(953,962)
isvar = np.any((root[ampl]['NALT'][:, sampleset] > 0), axis=1) # not considering all ALT - PEST is all REF
p1 = ax.bar(ampl, isvar.sum(), color='g')
p2 = ax.bar(ampl, (~isvar).sum(), bottom=isvar.sum(), color='r')
plt.legend((p1[0], p2[0]), ('Variant', 'Invariant'))
ax.set_title('Variation in Ag1000g biallelic sites within amplicons for first 10 Ag1000g samples');
raise Exception('Analysis ended!')
# ## Variant filtering sandbox
ag1k_meta = pd.read_csv(AG1K_META, sep='\t')
# visualise sequencing variants and ag1k biallelic sites for a given amplicon
# data
def plot_ampl_var_pattern(ampl):
# seq data
seq_root = zarr.open_group(ZARR, "r")
# mask
raw_isvar = np.any((seq_root[ampl]['raw_nalt'][:] > 0), axis=1)
# individual variants
ind_var = np.sum(seq_root[ampl]['raw_genotype'][:], axis=2)
ind_var = np.swapaxes(ind_var, 0,1)
# individual nsites
ind_nalt = seq_root[ampl]['raw_nalt']
ind_nalt = np.swapaxes(ind_nalt, 0,1)
seq_len = ind_var.shape[1]
# ref data
ag_root = zarr.open_group(AG1K_ZARR, "r")
# coordinate corrected to match POS in Ag1000g
ampl_pos = np.arange(panel.loc[ampl,'start_insert'] + 1, panel.loc[ampl,'end_insert'] + 1)
ag_ampl_pos = ag_root[ampl]['POS'][:]
ag_ampl_pos_mask = np.in1d(ampl_pos, ag_ampl_pos)
# sample 5 AOcol and GAgam
sampleset = slice(953,962)
ags_nalt = ag_root[ampl]['NALT'][:,sampleset]
ags_nalt = np.swapaxes(ags_nalt, 0,1)
# expand to sequence length (fill with REF)
large_ags_nalt = np.zeros([ags_nalt.shape[0],seq_len])
large_ags_nalt[:, ag_ampl_pos_mask] = ags_nalt
# comb data
d = [raw_isvar,ag_ampl_pos_mask]
# plot
fig_w = seq_len // 12
fig, axs = plt.subplots(4,1,figsize=(fig_w,10))
# ind var
axs[0].imshow(ind_var, aspect='auto', cmap=plt.cm.gray);
axs[0].set_yticks(range(ind_var.shape[0]))
axs[0].set_yticklabels(spp_meta.ox_code)
axs[0].set_title('Individual genotypes in amplicon {}'.format(ampl))
axs[0].set_xticks([])
# ind nalt
axs[1].imshow(ind_nalt, aspect='auto', cmap=plt.cm.gray);
axs[1].set_yticks(range(ind_var.shape[0]))
axs[1].set_yticklabels(spp_meta.ox_code)
axs[1].set_title('Individual nalts in amplicon {}'.format(ampl))
axs[1].set_xticks([])
# masks
axs[3].imshow(d, aspect='auto', cmap=plt.cm.gray);
axs[3].set_yticks(range(len(d)))
axs[3].set_yticklabels(['seq_var','ag1k_biallelic'])
axs[3].set_title('Sequence variation and Ag1000g biallelic sites in amplicon {}'.format(ampl))
axs[3].set_xticks([])
# ag1k variation
axs[2].imshow(large_ags_nalt, aspect='auto', cmap=plt.cm.gray)
axs[2].set_yticks(range(ags_nalt.shape[0]))
axs[2].set_yticklabels(ag1k_meta.loc[sampleset, 'population'])
axs[2].set_title('Sequence variation in Ag1000g samples in amplicon {}'.format(ampl))
axs[2].set_xticks([])
plot_ampl_var_pattern('2')
plot_ampl_var_pattern('43')
plot_ampl_var_pattern('25')
plot_ampl_var_pattern('17')
# ## Convert to zarr sandbox
# ! rm -r {ZARR}
ampl = '6'
ampl_seq = spp_seq[spp_seq.target == ampl].copy()
ampl_seq
ampl_ref_aln = ampl_seq.loc[ampl_seq.s_Sample == REF_SAMPLE, 'aln']
assert ampl_ref_aln.shape[0] == 1, 'non-unique or missing REF seq in {}'.format(ampl)
ampl_ref_aln = ampl_ref_aln.iloc[0]
ampl_ref_aln
# gapless ref length
ampl_ref_len = len(ampl_ref_aln.replace('-',''))
ampl_ref_len
# +
def call_haploid(seq, ref_seq):
'''
Simple genotype inference relative to reference:
match - 0,
mismatch - 1
indel - -1
Not vectorised
Not accounting for various ALT's
'''
assert len(seq) == len(ref_seq), 'sequences unaligned'
gt = []
for c, ref_c in zip(seq, ref_seq):
# skip insertions
if ref_c == '-':
continue
# deletion - missing data
if c == '-':
gt.append(-1)
# match
elif c == ref_c:
gt.append(0)
# mismatch
else:
gt.append(1)
return gt
call_haploid('a-tt','ac-c')
# -
ampl_seq['gt'] = ampl_seq.aln.apply(lambda x: call_haploid(x, ampl_ref_aln))
ampl_seq['gt'].apply(lambda x: sum(x))
# output gt matrix axes samples:sites:ploidy
ampl_gt = []
# order of samples as in metadata
for sample in spp_meta.ox_code:
sample_ampl_gt = ampl_seq.loc[ampl_seq.s_Sample == sample, 'gt']
# missing
if sample_ampl_gt.shape[0] == 0:
diploid_gt = [(-1,-1) for i in range(ampl_ref_len)]
# homozygous
elif sample_ampl_gt.shape[0] == 1:
diploid_gt = [(x,x) for x in sample_ampl_gt.iloc[0]]
# heterozygous - take first two alleles
else:
diploid_gt = [(x,y) for x,y in zip(sample_ampl_gt.iloc[0], sample_ampl_gt.iloc[1])]
if sample_ampl_gt.shape[0] > 2:
print('Warning: more than two alleles in sample {} amplicon {}'.format(sample, ampl))
ampl_gt.append(diploid_gt)
# reshape to sites:samples:ploidy
ampl_gt = np.swapaxes(ampl_gt, 0, 1)
ampl_gt.shape
# get coordinates in PEST
ampl_pos = np.arange(panel.loc[ampl,'start_insert'], panel.loc[ampl,'end_insert'])
len(ampl_pos)
ampl_pos
# get positions used in ag1k
ag_root = zarr.open_group(AG1K_ZARR, "r")
ag_ampl_pos = ag_root[ampl]['POS'][:]
ag_ampl_pos
ag_root[ampl].tree()
# mask with positions in Ag1000g dataset
mask = np.in1d(ampl_pos, ag_ampl_pos)
# subset along axis 1 (variants)
ag_ampl_gt = ampl_gt[mask,:,:]
assert ag_ampl_pos.shape[0] == ag_ampl_gt.shape[0], 'Position subsetting failed in {}'.format(ampl)
# nalts
ag_ampl_nalt = allel.GenotypeArray(ag_ampl_gt).to_n_alt()
ag_ampl_nalt.shape
# write
root = zarr.open_group(ZARR, "a")
g = root.create_group(ampl)
g.create_dataset('genotype', data=ag_ampl_gt)
g.create_dataset('NALT', data=ag_ampl_nalt)
|
work/8_ag1k_analysis/0_import_seq_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 教學目標:
# 了解 Padding 的作法
#
# 了解 Pooling 的作法
# # 範例說明:
# (1)zero_pad - feature map 跟原圖的比對
#
# (2)pool_forward - 前行網路的 pooling layer
# +
import numpy as np
import h5py
import matplotlib.pyplot as plt
# 繪圖結果直接顯示在Jupyter cell 之內
# %matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # 設定繪圖板的大小
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# autoreload。可以讓我們不退出IPython就動態修改代碼,在執行代碼前IPython會幫我們自動重載改動的模塊
# %load_ext autoreload
# %autoreload 2
np.random.seed(1)
# -
# GRADED FUNCTION: zero_pad
def zero_pad(X, pad):
"""
對image X 做 zero-padding.
參數定義如下:
X -- python numpy array, 呈現維度 (m, n_H, n_W, n_C), 代表一批 m 個圖像
n_H: 圖高, n_W: 圖寬, n_C: color channels 數
pad -- 整數, 加幾圈的 zero padding.
Returns:
X_pad -- image of shape (m, n_H + 2*pad, n_W + 2*pad, n_C) 做完zero-padding 的結果
"""
### Code 起始位置
X_pad = np.pad(X, ((0, 0), (pad, pad), (pad, pad), (0, 0)), 'constant', constant_values=(0, 0))
return X_pad
# +
'''
seed( ) 用於指定隨機數生成時所用算法開始的整數值,
如果使用相同的seed( )值,則每次生成的隨即數都相同,
如果不設置這個值,則係統根據時間來自己選擇這個值,
此時每次生成的隨機數因時間差異而不同。
'''
np.random.seed(1)
x = np.random.randn(4, 3, 3, 2) #產生gray image
x_pad = zero_pad(x, 2) # 加兩圈 Pad
print ("x.shape =", x.shape)
print ("x_pad.shape =", x_pad.shape)
print ("x[1,1] =", x[1,1])
print ("x_pad[1,1] =", x_pad[1,1])
fig, axarr = plt.subplots(1, 2)
axarr[0].set_title('x')
axarr[0].imshow(x[0,:,:,0])
axarr[1].set_title('x_pad')
axarr[1].imshow(x_pad[0,:,:,0])
# -
# GRADED FUNCTION: pool_forward
def pool_forward(A_prev, hparameters, mode = "max"):
"""
設計一個前行網路的池化層
參數定義如下:
A_prev -- 輸入的numpy 陣列, 維度 (m, n_H_prev, n_W_prev, n_C_prev)
hparameter 超參數 -- "f" and "stride" 所形成的python 字典
mode -- 池化的模式: "max" or "average"
返回:
A -- 輸出的池化層, 維度為 (m, n_H, n_W, n_C) 的 numpy 陣列
cache -- 可以應用在 backward pass pooling layer 資料, 包含 input and hparameter
"""
# 檢索尺寸 from the input shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# 檢索超參數 from "hparameters"
f = hparameters["f"]
stride = hparameters["stride"]
# 定義輸出的dimensions
n_H = int(1 + (n_H_prev - f) / stride)
n_W = int(1 + (n_W_prev - f) / stride)
n_C = n_C_prev
# 初始化輸出的 matrix A
A = np.zeros((m, n_H, n_W, n_C))
### 程式起始位置 ###
for i in range(m): # 訓練樣本的for 迴圈
for h in range(n_H): # 輸出樣本的for 迴圈, 針對vertical axis
for w in range(n_W): # 輸出樣本的for 迴圈, 針對 horizontal axis
for c in range (n_C): # 輸出樣本的for 迴圈, 針對channels
# 找出特徵圖的寬度跟高度四個點
vert_start = h * stride
vert_end = h * stride+ f
horiz_start = w * stride
horiz_end = w * stride + f
# 定義第i個訓練示例中
a_prev_slice = A_prev[i, vert_start:vert_end, horiz_start:horiz_end,c]
# 計算輸入data 的池化結果. 使用 if statment 去做分類
if mode == "max":
A[i, h, w, c] = np.max(a_prev_slice)
elif mode == "average":
A[i, h, w, c] = np.mean(a_prev_slice)
### 程式結束 ###
# 儲存輸入的特徵圖跟所設定的超參數, 可以用在 pool_backward()
cache = (A_prev, hparameters)
# 確認輸出的資料維度
assert(A.shape == (m, n_H, n_W, n_C))
return A, cache
np.random.seed(1)
A_prev = np.random.randn(2, 4, 4, 3)
hparameters = {"stride" : 2, "f": 3}
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A =", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A =", A)
|
homeworks/D095/Day95-CNN_Pooling_Padding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.svm import SVC
from sklearn import metrics
from ast import literal_eval
from mlxtend.plotting import plot_decision_regions
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
import warnings
import numpy as np
from collections import OrderedDict
from lob_data_utils import lob, db_result, model
from lob_data_utils.svm_calculation import lob_svm
sns.set_style('whitegrid')
warnings.filterwarnings('ignore')
# -
data_length = 15000
stocks = ['9064', '9061', '9265']
should_save_fig = False # TODO: change plot names if changing that value
d_stocks = {}
d_cv_stocks = {}
d_test_stocks = {}
for s in stocks:
d, d_test = lob.load_prepared_data(
s, data_dir='../queue_imbalance/data/prepared', cv=False, length=data_length)
d.index = pd.to_datetime(d['Unnamed: 0'].values)
d_test.index = pd.to_datetime(d_test['Unnamed: 0'].values)
d['prev_queue_imbalance'] = [None] + d['queue_imbalance'].iloc[0:len(d)-1].values.tolist()
d.dropna(inplace=True)
d_test['prev_queue_imbalance'] = [None] + d_test['queue_imbalance'].iloc[0:len(d_test)-1].values.tolist()
d_test.dropna(inplace=True)
d_stocks[s] = d
d_test_stocks[s] = d_test
d_stocks['9061'].head()
# ## SVM with queue imbalance
df_res = {}
for s in stocks:
df_res_temp = pd.read_csv('res_{}_prev_queue_imbalance.csv'.format(s))
df_res_temp = df_res_temp[df_res_temp['features'] == 'queue_imbalance']
df_res_temp = df_res_temp[df_res_temp['method'] == 'svm_linear']
df_res_temp['stock'] = [s for i in range(len(df_res_temp))]
df_res[s] = df_res_temp
columns_to_mean = ['f1', 'kappa', 'matthews',
'precision', 'recall', 'roc_auc', 'train_f1', 'train_kappa',
'train_matthews', 'train_precision', 'train_recall', 'train_roc_auc']
for c in columns_to_mean:
cc = []
for i, row in df_res[s].iterrows():
cc.append(np.array(literal_eval(row[c])).mean())
df_res[s][c] = cc
df_res['9265'].sort_values(by='matthews', ascending=False).head()
df_best_svm = pd.DataFrame()
for s in stocks:
idx_max = df_res[s]['matthews'].idxmax()
df_best_svm = df_best_svm.append(df_res[s].loc[idx_max])
df_best_svm
df_res['9064']
print(df_best_svm[['stock', 'method', 'c', 'gamma', 'coef0']].to_latex())
def fit_best_svm_classifier(df_best_svm, df, stock=None):
gamma = df_best_svm[df_best_svm['stock'] == stock]['gamma'].values[0]
coef0 = df_best_svm[df_best_svm['stock'] == stock]['coef0'].values[0]
c = df_best_svm[df_best_svm['stock'] == stock]['c'].values[0]
kernel = df_best_svm[df_best_svm['stock'] == stock]['method'].values[0].split('_')[1]
X = df['queue_imbalance'].values.reshape(-1, 1)
y = df['mid_price_indicator']
clf = SVC(gamma=gamma, C=c, coef0=coef0, kernel=kernel, random_state=12313)
clf.fit(X, y)
return clf
# +
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import TimeSeriesSplit
f, ax = plt.subplots(1, 3, figsize=(15,4))
i = 0
for stock in stocks:
log_clf = fit_best_svm_classifier(df_best_svm, d_stocks[stock], stock=stock)
lob.plot_learning_curve(
log_clf, d_stocks[stock]['queue_imbalance'].values.reshape(-1, 1),
d_stocks[stock]['mid_price_indicator'],
cv=TimeSeriesSplit(n_splits=10, max_train_size=9000), ax=ax[i], scoring='f1')
ax[i].set_title('Learning Curve for {}'.format(stock))
i += 1
if should_save_fig:
print('Saving')
plt.savefig('svm_learning_curves.png')
# -
f, (ax1, ax3) = plt.subplots(1, 2, figsize=(21,6))
for stock in stocks:
log_clf = fit_best_svm_classifier(df_best_svm, d_stocks[stock], stock=stock)
pred_train = log_clf.predict(d_stocks[stock]['queue_imbalance'].values.reshape(-1, 1))
pred_test = log_clf.predict(d_test_stocks[stock]['queue_imbalance'].values.reshape(-1, 1))
d_stocks[stock]['pred_log'] = pred_train
d_test_stocks[stock]['pred_log'] = pred_test
lob.plot_f1(d_stocks[stock], log_clf, stock=stock, label='', ax=ax1,
title='Precision vs Recall for 3 stocks on training data')
lob.plot_f1(d_test_stocks[stock], log_clf, stock=stock, label='', ax=ax3,
title='Precision vs Recall for 3 stocks on testing data')
if should_save_fig:
print('Saving')
plt.savefig('svm_results.png')
# +
def get_scores_dict_for_data(functions_to_run, dfs, log_clf, stock):
scores = {'stock': stock}
for func_name, func in functions_to_run.items():
for df_name, df in dfs.items():
pred = log_clf.predict(df['queue_imbalance'].values.reshape(-1, 1))
scores['{}_{}'.format(df_name, func_name)] = func(df['mid_price_indicator'], pred)
return scores
functions_to_run = {'precision': metrics.precision_score, 'roc_auc': metrics.roc_auc_score,
'f1_score': metrics.f1_score, 'recall': metrics.recall_score,
'matthews': metrics.matthews_corrcoef, 'kappa': metrics.cohen_kappa_score}
scores = []
for stock in stocks:
log_clf = fit_best_svm_classifier(df_best_svm, d_stocks[stock], stock=stock)
dfs = {'train': d_stocks[stock], 'test': d_test_stocks[stock], }
res_train = model.validate_model(fit_best_svm_classifier(df_best_svm, d_stocks[stock], stock=stock),
d_stocks[stock][['queue_imbalance']], d_stocks[stock]['mid_price_indicator'])
res = get_scores_dict_for_data(functions_to_run, dfs, log_clf, stock)
res = {**res, **res_train}
scores.append(res)
df_scores = pd.DataFrame(scores, index=stocks)
# -
df_scores[['train_precision', 'precision', 'test_precision', 'train_recall', 'recall', 'test_recall']]
df_scores[['train_f1', 'f1', 'test_f1_score', 'train_roc_auc', 'roc_auc', 'test_roc_auc']]
f, ax = plt.subplots(1, 3, figsize=(27,6))
for i in range(len(stocks)):
s = stocks[i]
d_stocks[s]['Predicition of Mid Price Indicator'] = d_stocks[s]['pred_log']
d_stocks[s]['Mid Price Indicator'] = d_stocks[s]['mid_price_indicator']
d_stocks[s][['Predicition of Mid Price Indicator', 'Mid Price Indicator']].plot(
kind='kde', ax=ax[i])
ax[i].set_title('Density of Mid Price Indicator and its prediction {} on training data'.format(s))
ax[i].legend(loc='lower right')
if should_save_fig:
print('Saving')
plt.savefig('density_of_mid_price_and_prediction_training_data_svm.png')
for s in stocks:
d_stocks[s]['queue_imbalance'].plot(kind='kde')
d_stocks[s]['mid_price_indicator'].plot(kind='kde')
# +
def convert_scores(df, column):
scores = []
for i, row in df.iterrows():
scores.append(np.mean(row[column]))
return scores
scores_columns = ['f1', 'kappa', 'matthews', 'precision', 'recall', 'roc_auc', 'train_f1', 'train_kappa',
'train_matthews', 'train_precision', 'train_recall', 'train_roc_auc']
for col in scores_columns:
df_scores[col] = convert_scores(df_scores, col)
df_scores
# -
print('linear kernel')
df_scores[['matthews', 'test_matthews']]
print('linear kernel')
df_scores[['roc_auc', 'test_roc_auc']]
print(df_scores[['matthews', 'test_matthews', 'roc_auc', 'test_roc_auc']].to_latex())
print(df_scores[['f1', 'test_f1_score', 'precision', 'test_precision', 'recall', 'test_recall']].to_latex())
df_best_svm
# +
f, ax = plt.subplots(3, 1, figsize=(35, 15), sharex=True)
i = 0
for i in range(len(stocks)):
s = stocks[i]
df = d_stocks[s]
X = d_stocks[s][['queue_imbalance']].values
y = d_stocks[s]['mid_price_indicator'].values.astype(np.integer)
clf = fit_best_svm_classifier(df_best_svm, d_stocks[s], stock=s)
# plot_decision_regions(X[0:1500], y[0:1500], clf=clf,ax=ax[i], colors=','.join(['orange', 'blue']))
ax[i].set_xlabel('Queue Imbalance')
ax[i].set_title('SVM Decision Regions for {} on training data'.format(s))
ax[i].set_xlim(-1.01, 1.01)
if should_save_fig:
print('Saving')
plt.savefig('svm_decision_region.png')
# -
# ## Compare Queue Imbalance vs Prev Queue Imblanace
|
overview_val10/overview_all_three_svm-linear.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="0v8XUs2a_67p"
#6 - temperature
hora = 0
temperaturas_C = []
suma_temperaturas_C = 0
while hora < 5:
temperatura_F = int(input('Introduce los valores de las temperaturas para cada hora: '))
temperatura_C = round(( temperatura_F - 32) / 1.8,1)
temperaturas_C.append (temperatura_C)
suma_temperaturas_C = suma_temperaturas_C + temperatura_C
hora+=1
temperatura_media_C = suma_temperaturas_C / hora
if (temperatura_media_C > 65) or (temperatura_C > 80) :
print(temperaturas_C)
print ('Cambiar el sistema de refrigeración')
break
elif (temperatura_C >= 70) and (temperaturas_C [hora-2] >= 70) and (temperaturas_C [hora-3] >= 70) and (temperaturas_C [hora-4] >= 70):
print(temperaturas_C)
print ('Cambiar el sistema de refrigeración')
break
print(temperaturas_C)
print('No es necesario cambiar el sistema de refrigeración')
#Me encuentro con el problema de que si la primera temperatura es superior a 65ºC ya me indica que tengo que cambiar el sistema de refrigeración, porque la media va a ser superior a 65ºC
|
01_RampUp/week2/practices/06-temperature/your-solution-here/W206_Temperature-Cris.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false}
# %matplotlib inline
# -
#
# Similarity Queries
# ==================
#
# Demonstrates querying a corpus for similar documents.
#
#
# + jupyter={"outputs_hidden": false}
import logging
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
# -
# Creating the Corpus
# -------------------
#
# First, we need to create a corpus to work with.
# This step is the same as in the previous tutorial;
# if you completed it, feel free to skip to the next section.
#
#
# + jupyter={"outputs_hidden": false}
from collections import defaultdict
from gensim import corpora
import pprint
documents = [
"Human machine interface for lab abc computer applications",
"A survey of user opinion of computer system response time",
"The EPS user interface management system",
"System and human system engineering testing of EPS",
"Relation of user perceived response time to error measurement",
"The generation of random binary unordered trees",
"The intersection graph of paths in trees",
"Graph minors IV Widths of trees and well quasi ordering",
"Graph minors A survey",
]
# -
# remove common words and tokenize
stoplist = set('for a of the and to in'.split())
texts = [
[word for word in document.lower().split() if word not in stoplist]
for document in documents
]
pprint.pprint(stoplist)
pprint.pprint(texts)
# +
# remove words that appear only once
frequency = defaultdict(int)
for text in texts:
for token in text:
frequency[token] += 1
pprint.pprint(frequency)
texts = [
[token for token in text if frequency[token] > 1]
for text in texts
]
pprint.pprint(texts)
# +
dictionary = corpora.Dictionary(texts)
corpus = [dictionary.doc2bow(text) for text in texts]
pprint.pprint(dictionary)
pprint.pprint(corpus)
# -
# Similarity interface
# --------------------
#
# In the previous tutorials on
# `sphx_glr_auto_examples_core_run_corpora_and_vector_spaces.py`
# and
# `sphx_glr_auto_examples_core_run_topics_and_transformations.py`,
# we covered what it means to create a corpus in the Vector Space Model and how
# to transform it between different vector spaces. A common reason for such a
# charade is that we want to determine **similarity between pairs of
# documents**, or the **similarity between a specific document and a set of
# other documents** (such as a user query vs. indexed documents).
#
# To show how this can be done in gensim, let us consider the same corpus as in the
# previous examples (which really originally comes from Deerwester et al.'s
# `"Indexing by Latent Semantic Analysis" <http://www.cs.bham.ac.uk/~pxt/IDA/lsa_ind.pdf>`_
# seminal 1990 article).
# To follow Deerwester's example, we first use this tiny corpus to define a 2-dimensional
# LSI space:
#
#
# + jupyter={"outputs_hidden": false}
from gensim import models
lsi = models.LsiModel(corpus, id2word=dictionary, num_topics=2)
# -
# For the purposes of this tutorial, there are only two things you need to know about LSI.
# First, it's just another transformation: it transforms vectors from one space to another.
# Second, the benefit of LSI is that enables identifying patterns and relationships between terms (in our case, words in a document) and topics.
# Our LSI space is two-dimensional (`num_topics = 2`) so there are two topics, but this is arbitrary.
# If you're interested, you can read more about LSI here: `Latent Semantic Indexing <https://en.wikipedia.org/wiki/Latent_semantic_indexing>`_:
#
# Now suppose a user typed in the query `"Human computer interaction"`. We would
# like to sort our nine corpus documents in decreasing order of relevance to this query.
# Unlike modern search engines, here we only concentrate on a single aspect of possible
# similarities---on apparent semantic relatedness of their texts (words). No hyperlinks,
# no random-walk static ranks, just a semantic extension over the boolean keyword match:
#
#
# + jupyter={"outputs_hidden": false}
doc = "Human computer interaction"
vec_bow = dictionary.doc2bow(doc.lower().split())
vec_lsi = lsi[vec_bow] # convert the query to LSI space
print(vec_lsi)
# -
# In addition, we will be considering `cosine similarity <http://en.wikipedia.org/wiki/Cosine_similarity>`_
# to determine the similarity of two vectors. Cosine similarity is a standard measure
# in Vector Space Modeling, but wherever the vectors represent probability distributions,
# `different similarity measures <http://en.wikipedia.org/wiki/Kullback%E2%80%93Leibler_divergence#Symmetrised_divergence>`_
# may be more appropriate.
#
# Initializing query structures
# ++++++++++++++++++++++++++++++++
#
# To prepare for similarity queries, we need to enter all documents which we want
# to compare against subsequent queries. In our case, they are the same nine documents
# used for training LSI, converted to 2-D LSA space. But that's only incidental, we
# might also be indexing a different corpus altogether.
#
#
# + jupyter={"outputs_hidden": false}
from gensim import similarities
index = similarities.MatrixSimilarity(lsi[corpus]) # transform corpus to LSI space and index it
# -
# <div class="alert alert-danger"><h4>Warning</h4><p>The class :class:`similarities.MatrixSimilarity` is only appropriate when the whole
# set of vectors fits into memory. For example, a corpus of one million documents
# would require 2GB of RAM in a 256-dimensional LSI space, when used with this class.
#
# Without 2GB of free RAM, you would need to use the :class:`similarities.Similarity` class.
# This class operates in fixed memory, by splitting the index across multiple files on disk, called shards.
# It uses :class:`similarities.MatrixSimilarity` and :class:`similarities.SparseMatrixSimilarity` internally,
# so it is still fast, although slightly more complex.</p></div>
#
# Index persistency is handled via the standard :func:`save` and :func:`load` functions:
#
#
# + jupyter={"outputs_hidden": false}
index.save('./tmp/deerwester.index')
index = similarities.MatrixSimilarity.load('./tmp/deerwester.index')
# -
# This is true for all similarity indexing classes (:class:`similarities.Similarity`,
# :class:`similarities.MatrixSimilarity` and :class:`similarities.SparseMatrixSimilarity`).
# Also in the following, `index` can be an object of any of these. When in doubt,
# use :class:`similarities.Similarity`, as it is the most scalable version, and it also
# supports adding more documents to the index later.
#
# Performing queries
# ++++++++++++++++++
#
# To obtain similarities of our query document against the nine indexed documents:
#
#
# + jupyter={"outputs_hidden": false}
sims = index[vec_lsi] # perform a similarity query against the corpus
print(list(enumerate(sims))) # print (document_number, document_similarity) 2-tuples
# -
# Cosine measure returns similarities in the range `<-1, 1>` (the greater, the more similar),
# so that the first document has a score of 0.99809301 etc.
#
# With some standard Python magic we sort these similarities into descending
# order, and obtain the final answer to the query `"Human computer interaction"`:
#
#
# + jupyter={"outputs_hidden": false}
sims = sorted(enumerate(sims), key=lambda item: -item[1])
for i, s in enumerate(sims):
print(s, documents[i])
# -
# The thing to note here is that documents no. 2 (``"The EPS user interface management system"``)
# and 4 (``"Relation of user perceived response time to error measurement"``) would never be returned by
# a standard boolean fulltext search, because they do not share any common words with ``"Human
# computer interaction"``. However, after applying LSI, we can observe that both of
# them received quite high similarity scores (no. 2 is actually the most similar!),
# which corresponds better to our intuition of
# them sharing a "computer-human" related topic with the query. In fact, this semantic
# generalization is the reason why we apply transformations and do topic modelling
# in the first place.
#
# Where next?
# ------------
#
# Congratulations, you have finished the tutorials -- now you know how gensim works :-)
# To delve into more details, you can browse through the `apiref`,
# see the `wiki` or perhaps check out `distributed` in `gensim`.
#
# Gensim is a fairly mature package that has been used successfully by many individuals and companies, both for rapid prototyping and in production.
# That doesn't mean it's perfect though:
#
# * there are parts that could be implemented more efficiently (in C, for example), or make better use of parallelism (multiple machines cores)
# * new algorithms are published all the time; help gensim keep up by `discussing them <http://groups.google.com/group/gensim>`_ and `contributing code <https://github.com/piskvorky/gensim/wiki/Developer-page>`_
# * your **feedback is most welcome** and appreciated (and it's not just the code!):
# `bug reports <https://github.com/piskvorky/gensim/issues>`_ or
# `user stories and general questions <http://groups.google.com/group/gensim/topics>`_.
#
# Gensim has no ambition to become an all-encompassing framework, across all NLP (or even Machine Learning) subfields.
# Its mission is to help NLP practitioners try out popular topic modelling algorithms
# on large datasets easily, and to facilitate prototyping of new algorithms for researchers.
#
#
|
notebooks/04_run_similarity_queries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
d=int(input('Quantos dias o carro foi alugado? '))
r=float(input('Quantos KM rodados? '))
d1=d*60
r1=r*0.15
print('O total a pagar é de R${:.2f}'.format(d1+r1))
|
.ipynb_checkpoints/EX015 - Aluguel de Carros-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PROJ_irox_oer] *
# language: python
# name: conda-env-PROJ_irox_oer-py
# ---
# # Analayze local coordination environment of bulk IrOx polymorphs
# ---
#
# This will determine which structures to select for further processing
# # Import Modules
# +
import os
print(os.getcwd())
import sys
from ase.db import connect
import pandas as pd
# #########################################################
from methods import get_df_dft
# +
# Contents will be saved to json
out_dict = dict()
directory = "out_data"
if not os.path.exists(directory):
os.makedirs(directory)
# -
# # Read file
# +
FinalStructuresdb_file = os.path.join(
os.environ["PROJ_irox_oer_data"],
"active_learning_proj_data/FinalStructures_1.db")
db = connect(FinalStructuresdb_file)
data_list = []
for row in db.select():
row_dict = dict(
energy=row.get("energy"),
**row.key_value_pairs)
data_list.append(row_dict)
df = pd.DataFrame(data_list)
df = df[~df["stoich"].isna()]
df = df.set_index("structure_id")
df = df.drop(columns=["energy", "id_old", ])
# -
df_dft = get_df_dft()
# +
df_i = df[df.coor_env == "O:6"]
print("Number of octahedral AB2:", df_i[df_i.stoich == "AB2"].shape[0])
print("Number of octahedral AB3:", df_i[df_i.stoich == "AB3"].shape[0])
# +
df_dft_i = df_dft.loc[
df_dft.index.intersection(
df_i.index.tolist()
)
]
out_dict["bulk_ids__octa_unique"] = df_dft_i.index.tolist()
df_dft_i.head()
# +
import plotly.express as px
fig = px.histogram(df_dft_i, x="num_atoms", nbins=20)
fig.update_layout(title="Number of atoms for unique octahedral IrOx bulk structures")
fig.show()
# +
from plotting.my_plotly import my_plotly_plot
my_plotly_plot(
figure=fig,
plot_name="atom_count_histogram_octahedral",
write_html=True,
write_png=False,
png_scale=6.0,
write_pdf=False,
write_svg=False,
try_orca_write=False,
)
# -
# # Saving data
# #######################################################################
import json
data_path = os.path.join("out_data/data.json")
with open(data_path, "w") as fle:
json.dump(out_dict, fle, indent=2)
# #######################################################################
|
workflow/creating_slabs/selecting_bulks/select_bulks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from random import shuffle
import cv2
import uuid
from tqdm import tqdm
import os
import pydicom
import numpy as np
# -
# +
# dir Setup
data_dir = "/media/mic/ML/Training_Data/DEC-17-17/train/GRADE_3/"
output_dir = "/media/mic/ML/Training_Data/DEC-17-17/train/grade_3/"
patients = os.listdir(data_dir)
# -
print(patients)
print("")
print(str("Patients - ") + str(len(patients)))
def process_training_data():
# Define Vars
PX_SIZE = 256
IMAGES_FOUND = 0
MAX_IMAGES_COUNT = 10000
training_data = []
label = [0,0,1,0]
# Patients by ID
for patient_id in patients:
for user in tqdm(os.listdir(data_dir + patient_id)):
patient_path = data_dir + patient_id + '/' + user
patient_scans = os.listdir(patient_path)
# Patient each Scans
for patient_scan in patient_scans:
data_path = patient_path + '/' + patient_scan
# Scan Images .dcm
for s in os.listdir(data_path):
if IMAGES_FOUND <= MAX_IMAGES_COUNT:
slice = pydicom.read_file(data_path + '/' + s)
if slice.pixel_array.shape[0] >= PX_SIZE and slice.pixel_array.shape[1] >= PX_SIZE:
# Resize
img = cv2.resize(np.array(slice.pixel_array),(PX_SIZE,PX_SIZE))
# Save As PNG
#cv2.imwrite( str(output_dir) + str(uuid.uuid1()) + ".jpg", img)
# Add to Array
training_data.append([np.array(img),np.array(label)])
IMAGES_FOUND += 1
else:
print("Low Resolution Image", slice.pixel_array.shape)
elif IMAGES_FOUND > MAX_IMAGES_COUNT:
return training_data
if IMAGES_FOUND % 100 == 0:
print(IMAGES_FOUND)
shuffle(training_data)
np.save('train_data_256X256_grade_3.npy', training_data)
return training_data
training_data = process_training_data()
|
prepare_data/GRADE_3_PREPROCESSING.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# INFO
#
# ------------------------------------------------------------------------------
# This is a script for generating corpus from several txt files and filtering
#
# ----------------------------------------------------------------------
#
#
# Note that you might need to change filtering options depending on your data since this script
# was originally written for data in the Azerbaijani language
#
# -------------------------------------------------------------------------------------------
#
# simplest way to change filtering is to switch next regex with characters in your alphabet (and optionally few more puntuations or numbers)
# in following line. It will erase any other character that is not mentioned in regex.
# text =re.sub("[^qüertyuiopöğasdfghjklıəzxcvbnmçşi. \.]","",text)
#
# ----------------------------------------------------------------------------------------
#
import re, string, os
files = []
# +
# every txt file within directory one by one, filters them and appends them to the <files> list
for file in os.listdir(_directory_name_):
if file.endswith('.txt'):
text = (open("raw/" + i, encoding ="utf-8")).read()
text = text.replace(u'\xa0', u'').replace(u'\n', u' ').replace("Ģ","ş").replace("Ġ","i").replace(","," ").replace(":"," ").lower()
text = text.replace(u'...', u'.').replace(u'...', u'.').replace(u'..', u'.').replace(u".", u" . ")
text =re.sub("[^qüertyuiopöğasdfghjklıəzxcvbnmçşi. \.]","",text)
files.append(text)
# -
for i in os.listdir("filter"):
if i.endswith('.txt'):
a = (open("filter/" + i, encoding ="utf-8")).read()
a = a.replace(u'\xa0', u'').replace(u'\n', u'. ').replace("Ģ","ş").replace("Ġ","i").replace(","," ").replace(":"," ").lower()
a = a.replace(u'...', u'.').replace(u'...', u'.').replace(u'..', u'.').replace(u".", u" . ")
a =re.sub("[^qüertyuiopöğasdfghjklıəzxcvbnmçşi. \.]","",a)
files.append(a)
# sample for filtering one file instead of whole directory
a = (open(_your_file_, encoding ="utf-8")).read()
a = a.replace(u'\xa0', u'').replace(u'\n', u'. ').replace("Ģ","ş").replace("Ġ","i").replace(","," ").replace(":"," ").lower()
a = a.replace(u'...', u'.').replace(u'...', u'.').replace(u'..', u'.').replace(u".", u" . ")
a =re.sub("[^qüertyuiopöğasdfghjklıəzxcvbnmçşi. \.]","",a)
files.append(a)
#merging all filtered texts into one corpus
corpus = ' '.join(files)
# +
# creating file to upload corpus
file = open("corpus.txt","w",encoding = "utf-8")
file.write(corpus)
file.close()
# -
|
corpus_generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Xc53ZNVtt2DK" colab_type="text"
# **Imports**
# + id="-tBvcSHDt40H" colab_type="code" colab={}
import tensorflow as tf
import numpy as np
import rcwa_utils
import tensor_utils
import solver
import matplotlib.pyplot as plt
# + [markdown] id="ZZsj6Huot-sn" colab_type="text"
# **Loss Function Definition**
# + id="-ZzTt9P8t-CQ" colab_type="code" colab={}
def loss_func():
# Global parameters dictionary.
global params
# Generate permittivity and permeability distributions.
ER_t, UR_t = solver.generate_cylindrical_nanoposts(var_duty, params)
# Simulate the system.
outputs = solver.simulate(ER_t, UR_t, params)
# Maximize the reflectance.
ref_lambda1 = outputs['REF'][0, 0, 0]
return (1 - ref_lambda1)
# + [markdown] id="nPtzCzc6uD-3" colab_type="text"
# **Setup and Initialize Variables**
# + id="fOKOZVWDuEI-" colab_type="code" colab={}
# Initialize global `params` dictionary storing optimization and simulation settings.
params = solver.initialize_params(wavelengths = [632.0], thetas = [0.0])
params['erd'] = 6.76 # Grating layer permittivity.
params['ers'] = 2.25 # Subtrate layer permittivity.
params['PQ'] = [11, 11] # Fourier Harmonics.
# Initialize grating duty cycle variable.
var_shape = (1, params['pixelsX'], params['pixelsY'])
duty_initial = 0.6 * np.ones(shape = var_shape)
var_duty = tf.Variable(duty_initial, dtype = tf.float32)
# + [markdown] id="nZ8sGxj9uWVf" colab_type="text"
# **Optimize**
# + id="9XyygRg0uWdv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="85f89e7f-a722-4395-d1fa-f505438fbc95"
# Number of optimization iterations.
N = 49
# Define an optimizer and data to be stored.
opt = tf.keras.optimizers.Adam(learning_rate = 1E-3)
loss = np.zeros(N + 1)
duty = np.zeros(N + 1)
length = np.zeros(N + 1)
# Compute the initial loss.
loss[0] = loss_func().numpy()
# Optimize.
print('Optimizing...')
for i in range(N):
opt.minimize(loss_func, var_list = [var_duty])
loss[i + 1] = loss_func().numpy()
# + [markdown] id="HAD2a8oludGP" colab_type="text"
# **Display Learning Curve**
# + id="Axql3myAaZ2V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="a9da9e39-361f-4c90-a583-a33aeb54c043"
plt.plot(loss)
plt.xlabel('Iterations')
plt.ylabel('Transmittance')
plt.xlim(0, N)
plt.show()
# + id="60hyJXTYuoUH" colab_type="code" colab={}
|
examples/gratings/reflective_grating.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Grover's algorithm examples
#
# This notebook has examples demonstrating how to use the Qiskit [Grover](https://qiskit.org/documentation/stubs/qiskit.aqua.algorithms.Grover.html) search algorithm, with different oracles.
import pylab
import numpy as np
from qiskit import BasicAer
from qiskit.tools.visualization import plot_histogram
from qiskit.aqua import QuantumInstance
from qiskit.aqua.algorithms import Grover
from qiskit.aqua.components.oracles import LogicalExpressionOracle, TruthTableOracle
# ## Finding solutions to 3-SAT problems
#
# Let's look at an example 3-Satisfiability (3-SAT) problem and walk-through how we can use Quantum Search to find its satisfying solutions. 3-SAT problems are usually expressed in [Conjunctive Normal Forms (CNF)](https://en.wikipedia.org/wiki/Conjunctive_normal_form) and written in the [DIMACS-CNF](http://www.satcompetition.org/2009/format-benchmarks2009.html) format. For example:
input_3sat_instance = '''
c example DIMACS-CNF 3-SAT
p cnf 3 5
-1 -2 -3 0
1 -2 3 0
1 2 -3 0
1 -2 -3 0
-1 2 3 0
'''
# The CNF of this 3-SAT instance contains 3 variables and 5 clauses:
#
# $(\neg v_1 \vee \neg v_2 \vee \neg v_3) \wedge (v_1 \vee \neg v_2 \vee v_3) \wedge (v_1 \vee v_2 \vee \neg v_3) \wedge (v_1 \vee \neg v_2 \vee \neg v_3) \wedge (\neg v_1 \vee v_2 \vee v_3)$
#
# It can be verified that this 3-SAT problem instance has three satisfying solutions:
#
# $(v_1, v_2, v_3) = (T, F, T)$ or $(F, F, F)$ or $(T, T, F)$
#
# Or, expressed using the DIMACS notation:
#
# `1 -2 3`, or `-1 -2 -3`, or `1 2 -3`.
#
# With this example problem input, we then create the corresponding `oracle` for our `Grover` search. In particular, we use the `LogicalExpressionOracle` component, which supports parsing DIMACS-CNF format strings and constructing the corresponding oracle circuit.
oracle = LogicalExpressionOracle(input_3sat_instance)
# The `oracle` can now be used to create an Grover instance:
grover = Grover(oracle)
# We can then configure the backend and run the Grover instance to get the result:
backend = BasicAer.get_backend('qasm_simulator')
quantum_instance = QuantumInstance(backend, shots=1024)
result = grover.run(quantum_instance)
print(result.assignment)
# As seen above, a satisfying solution to the specified 3-SAT problem is obtained. And it is indeed one of the three satisfying solutions.
#
# Since we used the `'qasm_simulator'`, the complete measurement result is also returned, as shown in the plot below, where it can be seen that the binary strings `000`, `011`, and `101` (note the bit order in each string), corresponding to the three satisfying solutions all have high probabilities associated with them.
# + tags=["nbsphinx-thumbnail"]
plot_histogram(result.measurement)
# -
# ## Boolean Logical Expressions
#
# Qiskit's `Grover` can also be used to perform Quantum Search on an `Oracle` constructed from other means, in addition to DIMACS. For example, the `LogicalExpressionOracle` can actually be configured using arbitrary Boolean logical expressions, as demonstrated below.
expression = '(w ^ x) & ~(y ^ z) & (x & y & z)'
oracle = LogicalExpressionOracle(expression)
grover = Grover(oracle)
result = grover.run(QuantumInstance(BasicAer.get_backend('qasm_simulator'), shots=1024))
plot_histogram(result.measurement)
# In the example above, the input Boolean logical expression `'(w ^ x) & ~(y ^ z) & (x & y & z)'` should be quite self-explanatory, where `^`, `~`, and `&` represent the Boolean logical XOR, NOT, and AND operators, respectively. It should be quite easy to figure out the satisfying solution by examining its parts: `w ^ x` calls for `w` and `x` taking different values; `~(y ^ z)` requires `y` and `z` be the same; `x & y & z` dictates all three to be `True`. Putting these together, we get the satisfying solution `(w, x, y, z) = (False, True, True, True)`, which our `Grover`'s result agrees with.
# ## TruthTable Oracles
#
# With Qiskit, `Oracle`s can also be constructed from truth tables, meaning we can also perform Quantum Search on truth tables. Even though this might seem like a moot point as we would be essentially searching for entries of a truth table with the $1$ value, it's a good example for demonstrative purpose.
truthtable = '1000000000000001'
# As shown, the `truthtable` is specified with a bitstring containing values of all entries in the table. It has length $16$, so the corresponding truth table is of $4$ input bits. Since the very first and last values are $1$, the corresponding truth table target entries are `0000` and `1111`.
#
# Next, we can setup the `Oracle` and `Grover` objects to perform Quantum Search as usual.
oracle = TruthTableOracle(truthtable)
grover = Grover(oracle)
result = grover.run(QuantumInstance(BasicAer.get_backend('qasm_simulator'), shots=1024))
plot_histogram(result.measurement)
# As seen in the above plot the search result coincides with our expectation.
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
|
tutorials/algorithms/08_grover_examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="01kP0XEgTJ-c" outputId="439d43e3-a1e1-4c08-c114-6b0f520557a7"
# !pip install gdown
# + colab={"base_uri": "https://localhost:8080/"} id="3LxRMP3eTRzS" outputId="151d0083-fca2-450d-a0ea-ecb4434ff36d"
# !gdown https://drive.google.com/uc?id=1nI47j3kVW-ZFcUAUSYJVp17wwIs-EpHC
# !unzip Scrapping.zip
# !rm -rf Scrapping.zip
# + colab={"base_uri": "https://localhost:8080/"} id="qzUBUz-OsSSm" outputId="881de6d1-131c-4bf9-935a-29c238c5a3b3"
import os
BASE_PATH = os.getcwd()
if not os.path.exists('/model'):
# !mkdir model
if not os.path.exists('/logs'):
# !mkdir logs
# + colab={"base_uri": "https://localhost:8080/"} id="ircwCxcEThmt" outputId="b6a64ec7-c303-4883-85f9-f2309c730f89"
import os
train_dir = os.path.join('/content/Train')
test_dir = os.path.join('/content/Test')
print("===TRAINING===")
for i in os.listdir(train_dir):
print(f'Total Training {i}: {len(os.listdir(os.path.join(train_dir, i)))}')
print("\n===VALIDATION===")
for i in os.listdir(test_dir):
print(f'Total Validation {i}: {len(os.listdir(os.path.join(test_dir, i)))}')
print(f'\nTotal Class: {len(os.listdir(train_dir))}')
# + id="cpeUzokcVeGW"
import datetime
import tensorflow as tf
import keras_preprocessing
from tensorflow.keras import layers
from keras_preprocessing import image
from keras_preprocessing.image import ImageDataGenerator
# + [markdown] id="vw7CuNZrTo0x"
# # Pre Processing
# + colab={"base_uri": "https://localhost:8080/"} id="rIy68hFJUQRU" outputId="7d5fc6d1-24ef-4f7f-a6b8-62cfade2e5cc"
TRAINING_DIR = "/content/Train"
training_datagen = ImageDataGenerator(rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
VALIDATION_DIR = "/content/Test"
validation_datagen = ImageDataGenerator(rescale=1./255)
train_generator = training_datagen.flow_from_directory(TRAINING_DIR,
target_size=(160, 160),
class_mode='categorical',
interpolation='nearest')
validation_generator = validation_datagen.flow_from_directory(VALIDATION_DIR,
target_size=(160, 160),
class_mode='categorical',
interpolation='nearest')
# + [markdown] id="ZO15maSnT84E"
# # Transfer Learning: Arsitektur & Config
# + id="w9kULVLaUDVD"
base_model = tf.keras.applications.MobileNetV2(input_shape=(160,160,3))
# + id="PRH_ZX9XYAYh"
# Frezee The Model
base_model.trainable = False
# + colab={"base_uri": "https://localhost:8080/"} id="rPNfJsEVUIus" outputId="0a167a21-03f1-4ef7-da2c-17832d052f3b"
base_model.summary()
# + id="PjvxLmlIUM8F"
base_input = base_model.layers[0].input ##input
base_output = base_model.layers[-2].output
# Flat_layer = layers.Flatten()(base_output)
# final_output = layers.Dense(1280)(Flat_layer)
final_output = layers.Dense(9, activation='softmax')(base_output)
# final_output = layers.Activation('softmax')(final_output)
# + id="EKrqJxzPUc9v"
model = tf.keras.Model(inputs= base_input, outputs= final_output)
# + colab={"base_uri": "https://localhost:8080/"} id="Q3WtTozSUfPX" outputId="8efbfad5-fda9-4c82-9d6d-5f5ec481cb6b"
model.summary()
# + id="beXqedrRUvhf"
MODEL_PATH = os.path.join('/content/model')
tensorboard_path = os.path.join(
BASE_PATH, "logs", datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
)
checkpoint_path = os.path.join(MODEL_PATH, "save_at_{epoch}")
callbacks = [
tf.keras.callbacks.ModelCheckpoint(checkpoint_path),
tf.keras.callbacks.TensorBoard(log_dir=tensorboard_path,
histogram_freq=1),
tf.keras.callbacks.EarlyStopping(monitor="val_loss",
patience=2,
mode="auto",
restore_best_weights=True),]
# + [markdown] id="ApphNqhxZGsm"
# # Adaptation Phase
# + id="QjMO29HxUrrm"
lr = 0.0001
model.compile(loss=tf.keras.losses.CategoricalCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.Adam(lr),
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="DZ-tu8hfziYb" outputId="610883d5-fd58-479e-e224-5e2f25e8c4bb"
# Initial Epochs
initial_epochs = 10
loss0, accuracy0 = model.evaluate(validation_generator)
# + id="KB6VNj8i3Jem"
# Error Handling Image Size Inconsistent
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
# + colab={"base_uri": "https://localhost:8080/"} id="C-GnrT7bnEVa" outputId="96772707-0e27-46b1-b713-abe625901be2"
history = model.fit(train_generator,
epochs=100,
validation_data=validation_generator,
callbacks=callbacks)
# + id="4_yohFROoow9"
model.save('T_house_model_before_fine_tune.h5')
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="L1RpYTr-JHh4" outputId="76c256f8-1f8a-47f9-db77-3a2c6eebb3c8"
import matplotlib.pyplot as plt
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.ylabel('Cross Entropy')
plt.ylim([0,1.0])
plt.title('Training and Validation Loss')
plt.xlabel('epoch')
plt.show()
# + [markdown] id="rmBI6yv6Kay6"
# # Fine Tuning (Coming Soon)
# + id="Povaomx_UPrc"
base_model.trainable = True
# + colab={"base_uri": "https://localhost:8080/"} id="ffOC6-jgVWgQ" outputId="26babf79-77b0-4f59-c21b-377ced2cc7b7"
model.summary()
# + id="XeHotuTkVWmy"
callbacks = [
tf.keras.callbacks.ModelCheckpoint(checkpoint_path),
tf.keras.callbacks.TensorBoard(log_dir=tensorboard_path,
histogram_freq=1),
tf.keras.callbacks.EarlyStopping(monitor="val_loss",
patience=5,
mode="auto",
restore_best_weights=True),]
# + id="Qn2Vxs_RVWs8"
lr = 0.0001
model.compile(loss=tf.keras.losses.CategoricalCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.Adam(lr/10),
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="PnDaWvb2ctAt" outputId="3446f2ce-1220-437c-9294-47abc66b0610"
history_fine = model.fit(train_generator,
epochs=200,
initial_epoch=history.epoch[-1],
validation_data=validation_generator,
callbacks=callbacks,
verbose=1)
# + id="s-strCftctG-"
model.save('T_house_model_after_fine_tune.h5')
# + id="6FVMkU-gctNf"
acc = history.history['accuracy'][0:121]
val_acc = history.history['val_accuracy'][0:121]
loss = history.history['loss'][0:121]
val_loss = history.history['val_loss'][0:121]
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="aaFbONH0i5EA" outputId="f46ebd41-ffa7-43d5-bf48-cda018fea16d"
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.plot([85,85],
plt.ylim(), label='Start Fine Tuning')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.plot([85,85],
plt.ylim(), label='Start Fine Tuning')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.xlabel('epoch')
plt.show()
# + [markdown] id="y8Uylof24vk0"
# #Optional Step
# + colab={"base_uri": "https://localhost:8080/"} id="dWCMIarLJZeB" outputId="3187f58b-b3df-4bbd-fdc1-045d0ccff6ca"
# !zip -r "/content/model_24_04_2021.zip" "/content/model"
# !zip -r "/content/logs_24_04_2021.zip" "/content/logs"
|
TF_Traditional_House_Classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="6coq6VkPKjBn"
# #### Основы программирования в Python для социальных наук
#
# ## Web-scraping таблиц. Подготовка к самостоятельной
#
# Семинар 7
#
# *Автор: <NAME>, <NAME>*
#
# + [markdown] colab_type="text" id="_51vTUulKjBw"
# Этот блокнот поможет вам разобраться, как подходить к самостоятельной работе. Один из пунктов - это скрейпинг таблицы из википедии. Посмотрим на примере, как это делать. Вы знаете из онлайн-курса как пользоваться библиотеками для доступа к сайтам по ссылкам и библиотекой BS для поиска тегов. Сегодня посмотрим пример, как сохранить таблицу из вики.
# + [markdown] colab_type="text" id="B9wlRSI7KjB2"
# **Задание 1.**
# *5 баллов*
#
# 1. На странице в wiki https://en.wikipedia.org/wiki/List_of_nuclear_weapons_tests нужно найти таблицу под названием "Worldwide nuclear test with a yield of 1.4 Mt TNT equivalent and more".
# 2. С помощью поиска по тегам, нужно сохранить из таблицы следующие колонки: 'Date (GMT)', 'Yield (megatons)', 'Country'. Каждая колонка таблицы должна быть сохранена в отдельную переменную, внутри которой лежит список, где первое значение - название колонки. Например, для колонки 'Date (GMT)' список будет выглядеть так:
# ['Date (GMT)', 'October 31, 1952', ...остальные значения..., 'November 17, 1976']
# 3. Выведите эти три списка командой
# print(Dates)
# print(Yield)
# print(Country)
# + colab={} colab_type="code" id="prG9uZRkKjB6"
import requests
from bs4 import BeautifulSoup
website_url = requests.get('https://en.wikipedia.org/wiki/List_of_nuclear_weapons_tests').text
soup = BeautifulSoup(website_url,'lxml')
My_table = soup.find_all('table',{'class':'wikitable sortable'})
rows = My_table[1].find_all('tr')
Country = []
Country.append(rows[0].find_all('th')[3].get_text().strip())
for row in rows[1:]:
r = row.find_all('td')
Country.append(r[3].get_text().strip())
print(Dates)
print(Yield)
print(Country)
# + [markdown] colab_type="text" id="K7oSd3u9KjCJ"
# **Задание 2.**
# *5 баллов (каждыый шаг 1 балл)*
#
# 1. Напишите функцию, которая берет аргументом название страны и возвращает (return) среднюю мощность взрыва для этой страны (нужно сложить все значения из колонки 'Yield (megatons)', которым соответствует страна, например, США, и раделить на количество этих значений). Для подсчета используйте списки, которые вы извлекли в Задании 1.
# 2. Из списка Country оставьте только уникальные значения для стран и запустите вашу функцию в цикле для каждого значения Country. Внутри цикла сделайте следующий вывод "{название страны}: средняя мощность взрыва {средняя мощность} мегатон"
# 3. Отдельно сохраните в переменную и выведите среднюю мощность взрыва (Yield (megatons) для бомб, которые тестировались в USA.
# 4. Отдельно сохраните в переменную и выведите среднюю мощность взрыва (Yield (megatons) для бомб, которые тестировались в Soviet Union.
# 5. Сравните эти значения и выведите название страны, для которой средняя мощность взрыва выше.
#
#
# Задание, выполненное без использования автоматически собранных данных, не засчитывается (например, если вы скопировали все значения из таблицы вручную и нашли их среднее).
# + colab={} colab_type="code" id="y0PaUmqZKjCM"
def average_yield(country):
yield_sum = 0
yield_count = 0
for idx in range(len(Country)):
if Country[idx] == country:
yield_sum += Yield[idx]
yield_count += 1
return yield_sum / yield_count
# + colab={} colab_type="code" id="te9JpvMlKjCY"
for country in set(Country[1:]):
print(country, ': средняя мощность взрыва', average_yield(country), 'мегатон')
# + colab={} colab_type="code" id="uK9STrHDKjCh"
average_yield('USA')
# + colab={} colab_type="code" id="k5OXKLBdKjCu"
average_yield('Soviet Union')
# + colab={} colab_type="code" id="WV6-klCcKjC9"
if average_yield('USA') > average_yield('Soviet Union'):
print('USA')
else:
print('Soviet Union')
# + [markdown] colab_type="text" id="FLvp-KL7KjEM"
# # Пример решения Задания 1.
# + [markdown] colab_type="text" id="4V_Cb2t2KjEN"
# Сначала мы импортируем библиотеку `requests`. Она позволяет нам просто и удобно посылать HTTP/1.1 запросы, не утруждаясь ручным трудом.
# + colab={} colab_type="code" id="GCHfTwnWKjEO"
import requests
# + [markdown] colab_type="text" id="PyXOofPIKjET"
# Теперь мы должны указать адрес страницы с которой мы будем скрейпить данные и сохраним ее в переменную `website_url`.
# `requests.get(url).text` обратиться к сайту и вернет `HTML` код сайта.
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="x2WEjHmOKjEU" outputId="12930b62-1106-4505-c841-ff4660314201"
website_url = requests.get('https://en.wikipedia.org/wiki/List_of_nuclear_weapons_tests').text
website_url
# + [markdown] colab_type="text" id="bT9LVvyEKjEX"
# Как мы видим, весь код представлен просто блоком текста, который не удобно читать и разбирать. Поэтому мы создадим объект `BeautifulSoup` с помощью функциии `BeautifulSoup`, предварительно импортировав саму библиотеку. `Beautiful Soup` это библиотека для парсинга `HTML` и `XML` документов. Она создает дерево из `HTML` кода, что очень полезно при скрейпинге. Функция `prettify()` позволяет видеть код в более удобном виде, в том числе с разбивкой по тегам.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="hRGbPlOgKjEZ" outputId="2aa7d599-9c1e-4b4f-9e20-dc9a30022fac"
from bs4 import BeautifulSoup
soup = BeautifulSoup(website_url,'lxml')
print(soup.prettify())
# + [markdown] colab_type="text" id="KzzCs4YwKjEe"
# Если внимательно изучить код `HTML` искомой таблицы, то можно обнаружить что вся таблица находится в классе `Wikitable Sortable`. (Для включения отображения кода сайта в вашем браузере можно нажать правкой кнопкой мыши на таблицы и выбрать пункт *Исследовать элемент*).
# + [markdown] colab_type="text" id="hCvv4E0IKjEf"
# 
# + [markdown] colab_type="text" id="F6ggqHkBKjEg"
# Поэтому первой задачей будет найти класс *wikitable sortable* в коде `HTML`. Это можно сделать с помощью функции `find_all`, указав в качестве аргументов, что мы ищем тэг `table` с классом `wikitable sortable`.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="JxwjvVxxKjEh" outputId="de27b6e0-0669-411f-999c-5289774e175a"
My_table = soup.find_all('table',{'class':'wikitable sortable'})
My_table
# + [markdown] colab_type="text" id="nQZZEFpUKjEn"
# Но как вы могли заметить, то на страницы есть две таблицы, которые принадлежат этому классу. Функция `find_all` вернет все найденные объекты в виде списка. Поэтому проверим второй найденный элемент.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="p3SqtX1DKjEp" outputId="e12724cd-15d1-4df1-82db-1c59c0cfd688"
My_table[1]
# + [markdown] colab_type="text" id="FkYcIKRTKjEv"
# Все верно, это наша искомая таблица. Если дальше изучить содержимое таблицы, то станет понятно что внутри тега `th` находится заголовок таблицы, а внутри `td` строки таблицы. А оба этих тега находятся внутри тегов `tr` что является по факту строкой таблицы. Давайте извлечем все строки таблицы также используя функцию `find_all`.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="F5QYpbNzKjEw" outputId="2e60d107-6b57-4221-a612-a6899fa07bc1"
rows = My_table[1].find_all('tr')
rows
# + [markdown] colab_type="text" id="mZQYvyv4KjE2"
# Давайте внимательно изучим содержимое одной строки, вытащим все `td`. Отобразим вторую строчку:
# + colab={"base_uri": "https://localhost:8080/", "height": 137} colab_type="code" id="OYHYLlvKKjE3" outputId="a6d852aa-5ae5-4488-bacd-a2d93fe2691c"
rows[1].find_all('td')
# + [markdown] colab_type="text" id="ytmoYTZfKjE8"
# Мы видим нужные нам данные между тегов `<td><\td>`, а также ссылки с тегом `<a>` и даже смешанные ячейки с обоими этими вариантами. Давайте сначала извлечем просто данные. Для этого используем функцию `get_text()` - она вернет все что между тегами.
# + [markdown] colab_type="text" id="qCNsBY7FKjE9"
# Возьмем, например, дату (она будет первым элементом):
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="7N3ERSqEKjE-" outputId="0c36a552-5199-42c5-babb-19034055989b"
rows[1].find_all('td')[0].get_text()
# + [markdown] colab_type="text" id="NKExPVoTKjFC"
# Единственное, нам нужно отдельно обработать, это первую строку, в которой хранится заголовок ряда (table header)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="D6FMSQ24KjFD" outputId="67c3f383-2f87-4568-e655-028de61c7964"
rows[0].find_all('th')[0].get_text()
# + [markdown] colab_type="text" id="fF_onkH1KjFH"
# Все классно, только довайте избавимся от знака переноса строки.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="o7VP8de2KjFI" outputId="802f3948-1b76-4040-c081-10e2f08c3904"
rows[0].find_all('th')[0].get_text().strip()
# + [markdown] colab_type="text" id="T6KWXg1DKjFM"
# Вообще хорошая идея всегда использовать метод strip(), чтобы удалять такие знаки (если удалять нечего, ошибку он не выдаст).
# + [markdown] colab_type="text" id="UzfvB4coKjFN"
# Давайте теперь извлечем все даты. Создадим список для их хранения `Dates` и будет итерироваться по всем элементам:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="TaORSTo3KjFO" outputId="98f1d75e-7588-42e1-8b53-9794c0eae984"
Dates = []
Dates.append(rows[0].find_all('th')[0].get_text().strip()) # отдельно добавляем заголовок
for row in rows[1:]: # начинаем со второго ряда таблицы, потому что 0 уже обработали выше
r = row.find_all('td') # находим все теги td для строки таблицы
Dates.append(r[0].get_text().strip()) # сохраняем данные в наш список
Dates
# + [markdown] colab_type="text" id="lrWAxB80KjFR"
# Ок! Следующие колонки, которые нам нужны - мощность взрыва и страна. Давайте поймем, где их искать.
# + colab={"base_uri": "https://localhost:8080/", "height": 241} colab_type="code" id="yxBedDPLKjFS" outputId="e284fd84-b244-477d-d8a1-5e43fa723ab1"
rows[0]
# + [markdown] colab_type="text" id="qWwWexaeKjFV"
# Видим, что Yield вторая колонка, а страна третья. Соберем их в отдельные списки по той же схеме, что дату. Но сначала проверим, что правильно посчитали номера.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="U_wHxWlqKjFX" outputId="8a4c6e2d-8d52-4400-befd-e064801c9c84"
rows[0].find_all('th')[1]
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="HcuXz-LsKjFa" outputId="3c310980-37d9-440a-d48d-2abb1ef672a8"
rows[0].find_all('th')[3]
# + [markdown] colab_type="text" id="Fh9zlgKKKjFe"
# Вроде все правильно. Единственно, не забудем хранить числа как float
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="iHUOvwyLKjFf" outputId="8495bf4c-e9ee-4b18-cbc7-0cc010e14062"
Yield = []
Yield.append(rows[0].find_all('th')[1].get_text().strip()) # отдельно добавляем заголовок
for row in rows[1:]: # начинаем со второго ряда таблицы, потому что 0 уже обработали выше
r = row.find_all('td') # находим все теги td для строки таблицы
Yield.append(float(r[1].get_text().strip())) # сохраняем данные в наш список и переводим в float
Yield
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="cAjfGoVAKjFh" outputId="d929d3ea-d267-4f1c-bf48-181ccc794e73"
Country = []
Country.append(rows[0].find_all('th')[3].get_text().strip()) # отдельно добавляем заголовок
for row in rows[1:]: # начинаем со второго ряда таблицы, потому что 0 уже обработали выше
r = row.find_all('td') # находим все теги td для строки таблицы
Country.append(r[3].get_text().strip()) # сохраняем данные в наш список и переводим в float
Country
# + colab={"base_uri": "https://localhost:8080/", "height": 89} colab_type="code" id="MHhQI6fWKjFj" outputId="882ac651-3270-4007-a59e-f5abf44a3660"
print(Dates)
print(Yield)
print(Country)
# + [markdown] colab_type="text" id="sOGC8QvBKjFl"
# # Пример решения задания 2
# + [markdown] colab_type="text" id="Lp5wi55DKjFm"
# 1. Напишите функцию, которая берет аргументом название страны и возвращает (return) среднюю мощность взрыва для этой страны (нужно сложить все значения из колонки 'Yield (megatons)', которым соответствует страна, например, США, и раделить на количество этих значений). Для подсчета используйте списки, которые вы извлекли в Задании 1.
# 2. Из списка Country оставьте только уникальные значения для стран и запустите вашу функцию в цикле для каждого значения Country. Внутри цикла сделайте следующий вывод "{название страны}: средняя мощность взрыва {средняя мощность} мегатон"
# 3. Отдельно сохраните в переменную и выведите среднюю мощность взрыва (Yield (megatons) для бомб, которые тестировались в USA.
# 4. Отдельно сохраните в переменную и выведите среднюю мощность взрыва (Yield (megatons) для бомб, которые тестировались в Soviet Union.
# 5. Сравните эти значения и выведите название страны, для которой средняя мощность взрыва выше.
# + colab={} colab_type="code" id="AjvuRuV5KjFn"
# 1
def average_yield(country):
yield_sum = 0 # создаем счетчитк, в который будем приплюсовывать мощность каждого испытания в заданной стране
yield_count = 0 # создаем счетчик, в котором будем хранить количество испытаний
for idx in range(len(Country)): # запускаем цикл для всех значений индексов списка Country
if Country[idx] == country: # проверяем, равно ли значение в списке Country стране, для которой вызвана функция
yield_sum += Yield[idx] # если да, то добавляем мощность взрыва под этим же индексом
yield_count += 1 # считаем это исптание
return yield_sum / yield_count # после окончания работы цикла возвращаем среднюю мощность
# + colab={"base_uri": "https://localhost:8080/", "height": 103} colab_type="code" id="9SCRs4utYkM_" outputId="e3cb3341-a75d-4b13-a029-ed27d62f178e"
# 2
for country in set(Country[1:]): # чтобы оставить только уникальные значения - делаем множество из списка + с помощью среза избавляемся от от заголовка колонки под индексом [0]
print(country, ': средняя мощность взрыва', average_yield(country), 'мегатон')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="gcmB_lfTYq5D" outputId="06f7ebfb-f12c-4550-ce57-cefac7114c1c"
# 3, 4
yield_ussr = average_yield('Soviet Union')
yield_usa = average_yield('USA')
print(yield_ussr, yield_usa)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="shRI3eW3ZkTz" outputId="141b3dc5-8d63-46fc-8750-d56dbf6120e7"
# 5
if yield_ussr > yield_usa:
print('Soviet Union')
else:
print('USA')
# -
|
5. Web-scraping/web_scraping_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This notebook provides the functionality to build, train, and test a CNN for predicting mosquito age, grouped age, species, and status.
#
# ## Structure:
# * Import packages to be used.
# * Load mosquito data.
# * Define fucntions for plotting, visualisation, and logging.
# * Define a function to build the CNN.
# * Define a function to train the CNN.
# * Main section to organise data, define the CNN, and call the building and training of the CNN.
#
# +
import pylab as pl
import datetime
import pandas as pd
import itertools
from itertools import cycle
import pickle
import random as rn
import os
from time import time
from tqdm import tqdm
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.lines import Line2D
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.model_selection import train_test_split, KFold
from sklearn.metrics import confusion_matrix
from sklearn.preprocessing import normalize, StandardScaler
from sklearn.utils import resample
import tensorflow as tf
import keras
from keras.models import Sequential, Model
from keras import layers, metrics
from keras.layers import Input
from keras.layers.merge import Concatenate
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers.normalization import BatchNormalization
from keras.layers.convolutional import Conv1D, MaxPooling1D
from keras.models import model_from_json, load_model
from keras.regularizers import *
from keras.callbacks import CSVLogger
from keras import backend as K
# -
# rand_seed = np.random.randint(low=0, high=100)
rand_seed = 16
print(rand_seed)
# +
os.environ['PYTHONHASHSEED'] = '0'
## The below is necessary for starting Numpy generated random numbers in a well-defined initial state.
np.random.seed(42)
## The below is necessary for starting core Python generated random numbers in a well-defined state.
rn.seed(12345)
## Force TensorFlow to use single thread.
## Multiple threads are a potential source of
## non-reproducible results.
## For further details, see: https://stackoverflow.com/questions/42022950/which-seeds-have-to-be-set-where-to-realize-100-reproducibility-of-training-res
# session_conf = tf.ConfigProto(device_count = {'GPU':0}, intra_op_parallelism_threads=4) #session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
# session_conf = tf.ConfigProto(device_count = {'GPU':0}) #session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
#session_conf.gpu_options.per_process_gpu_memory_fraction = 0.5
## The below tf.set_random_seed() will make random number generation
## in the TensorFlow backend have a well-defined initial state.
## For further details, see: https://www.tensorflow.org/api_docs/python/tf/set_random_seed
tf.set_random_seed(1234)
gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.35)
sess = tf.Session(graph=tf.get_default_graph(), config=tf.ConfigProto(gpu_options=gpu_options))
K.set_session(sess)
# -
# ## Function used to create a new folder for the CNN outputs.
# Useful to stop forgetting to name a new folder when trying out a new model varient and overwriting a days training.
def build_folder(fold, to_build = False):
if not os.path.isdir(fold):
if to_build == True:
os.mkdir(fold)
else:
print('Directory does not exists, not creating directory!')
else:
if to_build == True:
raise NameError('Directory already exists, cannot be created!')
# ## Function for plotting confusion matrcies
# This normalizes the confusion matrix and ensures neat plotting for all outputs.
def plot_confusion_matrix(cm, classes, output, save_path, model_name, fold,
normalize=True,
title='Confusion matrix',
cmap=plt.cm.Blues,
printout=False):
font = {'weight' : 'normal',
'size' : 18}
matplotlib.rc('font', **font)
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
if printout:
print("Normalized confusion matrix")
else:
if printout:
print('Confusion matrix, without normalization')
if printout:
print(cm)
plt.figure(figsize=(8,8))
plt.imshow(cm, interpolation='nearest', cmap=cmap, vmin=0, vmax=1) # np.max(np.sum(cm, axis=1)))
# plt.title([title+' - '+model_name])
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout(pad=2)
# plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.savefig((save_path+"Confusion_Matrix_"+model_name+"_"+fold+"_"+output[1:]+".png"))
plt.close()
# ## Function used for visualizing outputs
# This splits the output data into the four categories before plotting the confusion matricies.
## for visualizing losses and metrics once the neural network fold is trained
def visualize(histories, save_path, model_name, fold, classes, outputs, predicted, true):
# Sort out predictions and true labels
for label_predictions_arr, label_true_arr, classes, outputs in zip(predicted, true, classes, outputs):
classes_pred = np.argmax(label_predictions_arr, axis=-1)
classes_true = np.argmax(label_true_arr, axis=-1)
cnf_matrix = confusion_matrix(classes_true, classes_pred)
plot_confusion_matrix(cnf_matrix, classes, outputs, save_path, model_name, fold)
# ## Data logging
## for logging data associated with the model
def log_data(log, name, fold, save_path):
f = open((save_path+name+'_'+str(fold)+'_log.txt'), 'w')
np.savetxt(f, log)
f.close()
# ## Fucntion for graphing the training data
# This fucntion creates tidy graphs of loss and accuracy as the models are training.
def graph_history(history, model_name, model_ver_num, fold, save_path):
font = {'weight' : 'normal',
'size' : 18}
matplotlib.rc('font', **font)
#not_validation = list(filter(lambda x: x[0:3] != "val", history.history.keys()))
# print('history.history.keys : {}'.format(history.history.keys()))
filtered = filter(lambda x: x[0:3] != "val", history.history.keys())
not_validation = list(filtered)
for i in not_validation:
plt.figure(figsize=(15,7))
# plt.title(i+"/ "+"val_"+i)
plt.plot(history.history[i], label=i)
plt.plot(history.history["val_"+i], label="val_"+i)
plt.legend()
plt.xlabel("epoch")
plt.ylabel(i)
plt.savefig(save_path +model_name+"_"+str(model_ver_num)+"_"+str(fold)+"_"+i)
plt.close()
# ## funciton to create the CNN
# This function takes as an input a list of dictionaries. Each element in the list is a new hidden layer in the model. For each layer the dictionary defines the layer to be used.
#
# ### Available options are:
# Convolutional Layer:
# * type = 'c'
# * filter = optional number of filters
# * kernel = optional size of the filters
# * stride = optional size of stride to take between filters
# * pooling = optional width of the max pooling
# * {'type':'c', 'filter':16, 'kernel':5, 'stride':1, 'pooling':2}
#
# dense layer:
# * type = 'd'
# * width = option width of the layer
# * {'type':'d', 'width':500}
# +
def create_models(model_shape, input_layer_dim):
regConst = 0.02
sgd = keras.optimizers.SGD(lr=0.003, decay=1e-5, momentum=0.9, nesterov=True, clipnorm=1.)
cce = 'categorical_crossentropy'
input_vec = Input(name='input', shape=(input_layer_dim,1))
for i, layerwidth in zip(range(len(model_shape)),model_shape):
if i == 0:
if model_shape[i]['type'] == 'c':
xd = Conv1D(name=('Conv'+str(i+1)), filters=model_shape[i]['filter'],
kernel_size = model_shape[i]['kernel'], strides = model_shape[i]['stride'],
activation = 'relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(input_vec)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
xd = MaxPooling1D(pool_size=(model_shape[i]['pooling']))(xd)
elif model_shape[i]['type'] == 'd':
xd = Dense(name=('d'+str(i+1)), units=model_shape[i]['width'], activation='relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(input_vec)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
xd = Dropout(name=('dout'+str(i+1)), rate=0.5)(xd)
else:
if model_shape[i]['type'] == 'c':
xd = Conv1D(name=('Conv'+str(i+1)), filters=model_shape[i]['filter'],
kernel_size = model_shape[i]['kernel'], strides = model_shape[i]['stride'],
activation = 'relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(xd)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
xd = MaxPooling1D(pool_size=(model_shape[i]['pooling']))(xd)
elif model_shape[i]['type'] == 'd':
if model_shape[i-1]['type'] == 'c':
xd = Flatten()(xd)
xd = Dropout(name=('dout'+str(i+1)), rate=0.5)(xd)
xd = Dense(name=('d'+str(i+1)), units=model_shape[i]['width'], activation='relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(xd)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
# xAge = Dense(name = 'age', units = 17,
# activation = 'softmax',
# kernel_regularizer = l2(regConst),
# kernel_initializer = 'he_normal')(xd)
xAgeGroup = Dense(name = 'age_group', units = 3,
activation = 'softmax',
kernel_regularizer = l2(regConst),
kernel_initializer = 'he_normal')(xd)
xSpecies = Dense(name ='species', units = 3,
activation = 'softmax',
kernel_regularizer = l2(regConst),
kernel_initializer = 'he_normal')(xd)
outputs = []
# for i in ['xAge', 'xAgeGroup', 'xSpecies']:
for i in ['xAgeGroup', 'xSpecies']:
outputs.append(locals()[i])
model = Model(inputs = input_vec, outputs = outputs)
model.compile(loss=cce, metrics=['acc'],
optimizer=sgd)
# model.summary()
return model
# -
# ## Function to train the model
#
# This function will split the data into training and validation and call the create models function. This fucntion returns the model and training history.
# +
def train_models(model_to_test, save_path, SelectFreqs=False):
model_shape = model_to_test["model_shape"][0]
model_name = model_to_test["model_name"][0]
# input_layer_dim = model_to_test["input_layer_dim"][0]
model_ver_num = model_to_test["model_ver_num"][0]
fold = model_to_test["fold"][0]
label = model_to_test["labels"][0]
features = model_to_test["features"][0]
classes = model_to_test["classes"][0]
outputs = model_to_test["outputs"][0]
compile_loss = model_to_test["compile_loss"][0]
compile_metrics = model_to_test["compile_metrics"][0]
## Kfold training
seed = rand_seed
kfold = KFold(n_splits=10, shuffle=True, random_state=seed)
## Split data into test and train
model_ver_num = 0
cv_scores = []
best_score = 0
for train_index, val_index in kfold.split(features):
print('Fold {} Running'.format(model_ver_num))
X_train, X_val = features[train_index], features[val_index]
y_train, y_val = list(map(lambda y:y[train_index], label)), list(map(lambda y:y[val_index], label))
model = create_models(model_shape, input_layer_dim)
if model_ver_num == 0:
model.summary()
modsavedir = save_path+"Model_"+str(model_ver_num)+"/"
build_folder(modsavedir, True)
history = model.fit(x = X_train,
y = y_train,
batch_size = 128*16,
verbose = 0,
epochs = 8000,
validation_data = (X_val, y_val),
callbacks = [keras.callbacks.EarlyStopping(monitor='val_loss',
patience=400, verbose=0, mode='auto'),
CSVLogger(modsavedir+model_name+"_"+str(model_ver_num)+'.csv', append=True, separator=';')])
scores = model.evaluate(X_val, y_val)
model.save((modsavedir+model_name+"_"+str(model_ver_num)+"_"+'Model.h5'))
graph_history(history, model_name, model_ver_num, 0, modsavedir)
if (scores[3] + scores[4]) > best_score:
out_model = model
out_history = history
model_ver_num += 1
# # Clear the Keras session, otherwise it will keep adding new
# # models to the same TensorFlow graph each time we create
# # a model with a different set of hyper-parameters.
# K.clear_session()
# # Delete the Keras model with these hyper-parameters from memory.
# del model
out_model.save((save_path+model_name+"_"+'Model.h5'))
graph_history(out_history, model_name, 0, 0, save_path)
return out_model, out_history
# -
# ## Load the data
#
# The data file is created using Loco Mosquito:
# https://github.com/magonji/MIMI-project/blob/master/Loco%20mosquito%204.0.ipynb
#
# ### The data file has headings: Species - Status - RearCnd - Age - Country- Frequencies
# +
df = pd.read_csv("/home/josh/Documents/Mosquito_Project/New_Data/Data/MIMIdata_update_19_02/mosquitoes_country_LM_5_0.dat", '\t')
df.head(10)
RearCnd_counts = df.groupby('RearCnd').size()
df['AgeGroup'] = 0
df['AgeGroup'] = np.where(df['Age']>10, 2, np.where(df['Age']>4, 1, 0))
df_vf = df[df['RearCnd']=='VF']
df_vf = df_vf[df_vf['Status']=='UN']
df = df[df['RearCnd']!='VF']
df = df[df['Status']!='UN']
df_l = df[df['RearCnd']=='TL']
df_l_g = df_l[df_l['Country']=='S']
df_l_g_a = df_l_g[df_l_g['Species']=='AA']
age_counts = df_l_g_a.groupby('AgeGroup').size()
df_l_g_g = df_l_g[df_l_g['Species']=='AG']
age_counts = df_l_g_g.groupby('AgeGroup').size()
df_l_g_c = df_l_g[df_l_g['Species']=='AC']
age_counts = df_l_g_c.groupby('AgeGroup').size()
df_l_t = df_l[df_l['Country']=='T']
df_l_t_a = df_l_t[df_l_t['Species']=='AA']
age_counts = df_l_t_a.groupby('AgeGroup').size()
df_l_t_g = df_l_t[df_l_t['Species']=='AG']
age_counts = df_l_t_g.groupby('AgeGroup').size()
df_l_b = df_l[df_l['Country']=='B']
df_l_b_g = df_l_b[df_l_b['Species']=='AG']
age_counts = df_l_b_g.groupby('AgeGroup').size()
df_l_b_c = df_l_b[df_l_b['Species']=='AC']
age_counts = df_l_b_c.groupby('AgeGroup').size()
df_f = df[df['RearCnd']=='TF']
df_f_t = df_f[df_f['Country']=='T']
df_f_t_a = df_f_t[df_f_t['Species']=='AA']
age_counts = df_f_t_a.groupby('AgeGroup').size()
# df_f_t_g = df_f_t[df_f_t['Species']=='AG'] #There isn't any
df_f_b = df_f[df_f['Country']=='B']
df_f_b_g = df_f_b[df_f_b['Species']=='AG']
age_counts = df_f_b_g.groupby('AgeGroup').size()
df_f_b_c = df_f_b[df_f_b['Species']=='AC']
age_counts = df_f_b_c.groupby('AgeGroup').size()
df_vf_t = df_vf[df_vf['Country']=='T']
df_vf_t_a = df_vf_t[df_vf_t['Species']=='AA']
age_counts = df_vf_t_a.groupby('AgeGroup').size()
df_vf_t_g = df_vf_t[df_vf_t['Species']=='AG']
age_counts = df_vf_t_g.groupby('AgeGroup').size()
df_vf_b = df_vf[df_vf['Country']=='B']
df_vf_b_g = df_vf_b[df_vf_b['Species']=='AG']
age_counts = df_vf_b_g.groupby('AgeGroup').size()
df_vf_b_c = df_vf_b[df_vf_b['Species']=='AC']
age_counts = df_vf_b_c.groupby('AgeGroup').size()
size_inc = 400
for age in range(3):
df_temp = df_l_t_a[df_l_t_a['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
if age == 0:
df_train = df_temp.iloc[index_df_temp_inc]
# df_test = df_temp.iloc[index_df_temp_not_inc]
else:
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_l_t_g[df_l_t_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 400
for age in range(3):
df_temp = df_l_b_g[df_l_b_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_l_b_c[df_l_b_c['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 300 # 50
for age in range(3):
df_temp = df_f_t_a[df_f_t_a['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
if age == 0:
df_trainf = df_temp.iloc[index_df_temp_inc]
# df_test = df_temp.iloc[index_df_temp_not_inc]
else:
df_trainf = pd.concat([df_trainf, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_f_b_g[df_f_b_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_trainf = pd.concat([df_trainf, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_f_b_c[df_f_b_c['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_trainf = pd.concat([df_trainf, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 0
for age in range(3):
df_temp = df_vf_t_a[df_vf_t_a['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
if age == 0:
df_test = df_temp.iloc[index_df_temp_not_inc]
else:
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_vf_t_g[df_vf_t_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 0
for age in range(3):
df_temp = df_vf_b_g[df_vf_b_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_vf_b_c[df_vf_b_c['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
X = df_train.iloc[:,6:-1]
y_age = df_train["Age"]
y_age_groups = df_train["AgeGroup"]
y_species = df_train["Species"]
y_status = df_train["Status"]
X = np.asarray(X)
y_age = np.asarray(y_age)
y_age_groups = np.asarray(y_age_groups)
y_species = np.asarray(y_species)
y_status = np.asarray(y_status)
print('shape of X : {}'.format(X.shape))
print('shape of y age : {}'.format(y_age.shape))
print('shape of y age groups : {}'.format(y_age_groups.shape))
print('shape of y species : {}'.format(y_species.shape))
print('shape of y status : {}'.format(y_status.shape))
Xf = df_trainf.iloc[:,6:-1]
y_agef = df_trainf["Age"]
y_age_groupsf = df_trainf["AgeGroup"]
y_speciesf = df_trainf["Species"]
y_statusf = df_trainf["Status"]
Xf = np.asarray(Xf)
y_agef = np.asarray(y_agef)
y_age_groupsf = np.asarray(y_age_groupsf)
y_speciesf = np.asarray(y_speciesf)
y_statusf = np.asarray(y_statusf)
print('shape of X f : {}'.format(Xf.shape))
print('shape of y age f : {}'.format(y_agef.shape))
print('shape of y age groups f : {}'.format(y_age_groupsf.shape))
print('shape of y species f : {}'.format(y_speciesf.shape))
print('shape of y status f : {}'.format(y_statusf.shape))
X_vf = df_test.iloc[:,6:-1]
y_age_vf = df_test["Age"]
y_age_groups_vf = df_test["AgeGroup"]
y_species_vf = df_test["Species"]
y_status_vf = df_test["Status"]
X_vf = np.asarray(X_vf)
y_age_vf = np.asarray(y_age_vf)
y_age_groups_vf = np.asarray(y_age_groups_vf)
y_species_vf = np.asarray(y_species_vf)
y_status_vf = np.asarray(y_status_vf)
# -
# ## Main section
#
# Functionality:
# * Oganises the data into a format of lists of data, classes, labels.
# * Define the CNN to be built.
# * Define the KFold validation to be used.
# * Build a folder to output data into.
# * Standardize and oragnise data into training/testing.
# * Call the model training.
# * Organize outputs and call visualization for plotting and graphing.
#
# +
input_layer_dim = len(Xf[0])
y_age_groups_list_l = [[age] for age in y_age_groups]
y_species_list_l = [[species] for species in y_species]
age_groups_l = MultiLabelBinarizer().fit_transform(np.array(y_age_groups_list_l))
age_group_classes = ["1-4", "5-10", "11-17"]
species_l = MultiLabelBinarizer().fit_transform(np.array(y_species_list_l))
species_classes = list(np.unique(y_species_list_l))
y_age_groups_list_f = [[age] for age in y_age_groupsf]
y_species_list_f = [[species] for species in y_speciesf]
age_groups_f = MultiLabelBinarizer().fit_transform(np.array(y_age_groups_list_f))
species_f = MultiLabelBinarizer().fit_transform(np.array(y_species_list_f))
outdir = "Results_Paper/"
build_folder(outdir, False)
SelectFreqs = False
## Labels default - all classification
labels_default_f, labels_default_l, classes_default, outputs_default = [age_groups_f, species_f], [age_groups_l, species_l], [age_group_classes, species_classes], ['xAgeGroup', 'xSpecies']
## Declare and train the model
model_size = [{'type':'c', 'filter':16, 'kernel':8, 'stride':1, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':8, 'stride':2, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':3, 'stride':1, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':6, 'stride':2, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':5, 'stride':1, 'pooling':2},
{'type':'d', 'width':500}]
## Name the model
model_name = 'Baseline_CNN'
histories = []
fold = 1
train_model = True
## Name a folder for the outputs to go into
savedir = (outdir+"Trian_Lab_Field_V2/")
build_folder(savedir, True)
start_time = time()
save_predicted = []
save_true = []
## Scale train, test
scl = StandardScaler()
features_scl = scl.fit(X=np.vstack((X,Xf)))
features_l = features_scl.transform(X=X)
features_f = features_scl.transform(X=Xf)
## Split into training / testing
test_splits = train_test_split(features_f, *(labels_default_f), test_size=0.1, shuffle=True, random_state=rand_seed)
## Pack up data
X_train = test_splits.pop(0)
X_test = test_splits.pop(0)
y_train = test_splits[::2]
y_test = test_splits[1::2]
X_train = np.vstack((X_train, features_l))
y_train = [np.vstack((y_train[0], labels_default_l[0])), np.vstack((y_train[1], labels_default_l[1]))]
if not SelectFreqs:
X_train = np.expand_dims(X_train, axis=2)
X_test = np.expand_dims(X_test, axis=2)
model_to_test = {
"model_shape" : [model_size], # defines the hidden layers of the model
"model_name" : [model_name],
"input_layer_dim" : [input_layer_dim], # size of input layer
"model_ver_num" : [0],
"fold" : [fold], # kf.split number on
"labels" : [y_train],
"features" : [X_train],
"classes" : [classes_default],
"outputs" : [outputs_default],
"compile_loss": [{'age': 'categorical_crossentropy'}],
"compile_metrics" :[{'age': 'accuracy'}]
}
## Call function to train all the models from the dictionary
model, history = train_models(model_to_test, savedir, SelectFreqs=SelectFreqs)
histories.append(history)
predicted_labels = list([] for i in range(len(y_train)))
true_labels = list([] for i in range(len(y_train)))
y_predicted = model.predict(X_test)
predicted_labels = [x+[y] for x,y in zip(predicted_labels,y_predicted)]
true_labels = [x+[y] for x,y in zip(true_labels,y_test)]
predicted_labels = [predicted_labels[i][0].tolist() for i in range(len(predicted_labels))]
true_labels = [true_labels[i][0].tolist() for i in range(len(true_labels))]
for pred, tru in zip(predicted_labels, true_labels):
save_predicted.append(pred)
save_true.append(tru)
## Visualize the results
visualize(histories, savedir, model_name, str(fold), classes_default, outputs_default, predicted_labels, true_labels)
# Clear the Keras session, otherwise it will keep adding new
# models to the same TensorFlow graph each time we create
# a model with a different set of hyper-parameters.
K.clear_session()
# Delete the Keras model with these hyper-parameters from memory.
del model
# visualize(1, savedir, model_name, "Averaged", classes_default, outputs_default, save_predicted, save_true)
end_time = time()
print('Run time : {} s'.format(end_time-start_time))
print('Run time : {} m'.format((end_time-start_time)/60))
print('Run time : {} h'.format((end_time-start_time)/3600))
# -
# ## Testing Phase
# ### Process testing data
def test_data_extract(RearCnd, Country, Species):
df_RearCnd = df_test[df_test['RearCnd']==RearCnd]
df_Country = df_RearCnd[df_RearCnd['Country']==Country]
df_Species = df_Country[df_Country['Species']==Species]
X_test_extract = df_Species.iloc[:,6:-1]
y_age_test_extract = df_Species["Age"]
y_age_groups_test_extract = df_Species["AgeGroup"]
y_species_test_extract = df_Species["Species"]
y_status_test_extract = df_Species["Status"]
print('shape of X_vf : {}'.format(X_test_extract.shape))
print('shape of y_age_vf : {}'.format(y_age_test_extract.shape))
print('shape of y_age_groups_test_extract : {}'.format(y_age_groups_test_extract.shape))
print('shape of y y_species_vf : {}'.format(y_species_test_extract.shape))
print('shape of y y_status_vf : {}'.format(y_status_test_extract.shape))
X_test_extract = np.asarray(X_test_extract)
y_age_test_extract = np.asarray(y_age_test_extract)
y_age_groups_test_extract = np.asarray(y_age_groups_test_extract)
y_species_test_extract = np.asarray(y_species_test_extract)
y_status_test_extract = np.asarray(y_status_test_extract)
return X_test_extract, y_age_test_extract, y_age_groups_test_extract, y_species_test_extract, y_status_test_extract
def test_data_format(y_age_groups, y_species):
y_age_groups_list = [[age] for age in y_age_groups]
y_species_list = [[species] for species in y_species]
age_groups = MultiLabelBinarizer().fit([(0,), (1,), (2,)])
age_groups = age_groups.transform(np.array(y_age_groups_list))
age_group_classes = ["1-4", "5-10", "11-17"]
species = MultiLabelBinarizer().fit([set(['AA']), set(['AC']), set(['AG'])])
species = species.transform(np.array(y_species_list))
species_classes = list(np.unique(y_species_list))
# labels_default, classes_default, outputs_default = [age_groups, species], [age_group_classes, species_classes], ['xAgeGroup', 'xSpecies']
labels_default, classes_default, outputs_default = [species], [species_classes], ['xSpecies']
return labels_default, classes_default, outputs_default
# +
## Set up folders
outdir = "output_data_update_19_02/"
build_folder(outdir, False)
loaddir = (outdir+"Train_Field/Trian_Species_Only/")
for i in range(0,10):
savedir = (outdir+"Train_Field/Trian_Species_Only/Testing/model"+str(i+1)+"/")
build_folder(savedir, True)
## Load model
model = load_model((loaddir+"Baseline_CNN_0_"+str(i+1)+"_Model.h5"))
print('Model loaded successfully')
## Set up testing data choice
RearCnd = 'TL'
Country = 'T'
Species = 'AA'
for RearCnd in ['TL', 'TF', 'VF']:
for Country, Species in [['T', 'AA'], ['T', 'AG'], ['B', 'AG'], ['B', 'AC']]:
## Extract data
X_test, y_age_test, y_age_groups_test, y_species_test, y_status_test = test_data_extract(RearCnd, Country, Species)
if len(y_age_test) == 0:
pass
else:
## Format data
labels_default, classes_default, outputs_default = test_data_format(y_age_groups_test, y_species_test)
model_name = 'Testing_'+RearCnd+'_'+Country+'_'+Species
## Scale train, test
scl = StandardScaler()
scl_fit = scl.fit(X=X)
features = scl_fit.transform(X=X)
features_test = scl_fit.transform(X=X_test)
## Split data into test and train
X_test = features_test[:]
y_test = list(map(lambda y:y[:], labels_default))
X_test = np.expand_dims(X_test, axis=2)
## Prediction
predicted_labels = list([] for i in range(len(y_test)))
true_labels = list([] for i in range(len(y_test)))
y_predicted = model.predict(X_test)
predicted_labels = [x+[y] for x,y in zip(predicted_labels,y_predicted)]
true_labels = [x+[y] for x,y in zip(true_labels,y_test)]
predicted_labels = [predicted_labels[i][0].tolist() for i in range(len(predicted_labels))]
true_labels = [true_labels[i][0].tolist() for i in range(len(true_labels))]
species_classes_pred_temp = np.unique(np.argmax(y_predicted[1], axis=-1))
species_classes_pred = []
for spec in species_classes_pred_temp:
if spec == 0:
species_classes_pred.append('AA')
elif spec == 1:
species_classes_pred.append('AC')
elif spec == 2:
species_classes_pred.append('AG')
species_classes_pred = np.array(species_classes_pred)
classes_pred = [classes_default[0], species_classes_pred]
## Visualize the results
# classes_default_vf_B = [list(np.unique(y_predicted[0])), age_group_classes_vf_B, species_classes_vf_B]
# visualize(histories, savedir, (model_name+'_B_AG'), str(fold), classes_pred, outputs_default_vf_B_AG, predicted_labels, true_labels)
visualize(histories, savedir, model_name, str(0), classes_pred, outputs_default, predicted_labels, true_labels)
print('Testing of {} - {} - {} completed'.format(RearCnd, Country, Species))
# -
# ## Sensitivity Plots
#
# +
## Functions for Z-score and sensitivity for input-output
def generate_sensitivity_Z_score(model, layer_name, age, size=1625):
layer_output = model.get_layer(layer_name).output
if layer_name == 'age':
df_1 = df[df['Age']==age]
loss = layer_output[:, age-1]
elif layer_name == 'species':
df_1 = df[df['Species']==age]
if age == 'AA':
loss = layer_output[:, 0]
elif age == 'AC':
loss = layer_output[:, 1]
elif age == 'AG':
loss = layer_output[:, 2]
elif layer_name == 'age_group':
df1 = df_train[df_train['AgeGroup']==age]
loss = layer_output[:, age]
X = df1.iloc[:,6:-1]
X = np.asarray(X)
grads = K.gradients(loss, model.input)[0]
iterate = K.function([model.input], [loss, grads])
gradients = []
for i in range(len(X)):
input_img_data = X[i]
input_img_data = np.expand_dims(input_img_data, axis=0)
input_img_data = np.expand_dims(input_img_data, axis=2)
loss_value, grads_value = iterate([input_img_data])
gradients.append(np.squeeze(np.abs(grads_value)))
sensitivity = 1/len(gradients) * np.sum(gradients, axis=0)
return sensitivity/np.linalg.norm(sensitivity)
def sensitivites_for_age(age):
outdir = "Results_Paper/"
loaddir = (outdir+"Trian_Lab_Field_V2/")
sensitivities = []
for count in tqdm(range(10)):
model = load_model((loaddir+"Model_"+str(count)+"/Baseline_CNN_"+str(count)+"_Model.h5"))
# model.summary()
for layer in model.layers:
layer.trainable = False
sensitivity = generate_sensitivity_Z_score(model, 'age_group', age)
sensitivities.append(sensitivity)
del model
return sensitivities
# +
## Generates outputs of Z-score and sensitivty for input-ouput
outdir = "Results_Paper/"
savedir = (outdir+"Trian_Lab_Field_V2/")
build_folder(savedir, False)
sensitivities_save = []
for age in tqdm(range(0,3)):
sensitivities = sensitivites_for_age(age)
sensitivities_save.append(sensitivities)
sensitivities = [sensitivities_save[0][i] + sensitivities_save[1][i] + sensitivities_save[2][i] for i in range(10)]
Z_scores = []
m_signals = []
for sens1 in range(10):
for sens2 in range(10):
s_signal = (sensitivities[sens1] + sensitivities[sens2]) / np.sqrt(2)
mean_b = np.mean(s_signal)
sigma_b = np.std(s_signal)
for sig in s_signal:
Z_b = (sig-mean_b)/sigma_b
Z_scores.append(Z_b)
m_signals.append(sig)
fig = plt.figure()
plt.scatter(m_signals, Z_scores)
poly_index = 3
plt.plot(np.unique(m_signals), np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals)), color='k', linewidth=3)
index_95 = (np.where(np.logical_and(np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals)) < 1.7, np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals)) > 1.6)))
index_95 = index_95[0][int(len(index_95)/2)]
y_value = (np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals))[index_95])
x_value = (np.unique(m_signals)[index_95])
plt.plot([0, x_value], [y_value, y_value], 'k--')
plt.plot([x_value, x_value], [-4, y_value], 'k--')
plt.ylim([-4, 6])
plt.xlabel('Signal value')
plt.ylabel('Z-score')
plt.title(('Z-score Calculation - Age '+str(age)))
plt.tight_layout()
plt.savefig((savedir+'Z_Score_Grouped_Age_'+str(age)+'.png'))
## Start of individual age Sensitivity plots
font = {'family' : 'normal',
'weight' : 'normal',
'size' : 14}
matplotlib.rc('font', **font)
fig = plt.figure(figsize=(8,4))
ax = fig.add_subplot(1,1,1)
sens_vals = np.sum(np.array(sensitivities), axis=0)/10
print(sens_vals.shape)
l1 = plt.plot(np.arange(3800, 550, -2), np.squeeze(sensitivities[0]), c='b')
l3 = plt.plot([3800, 550], [x_value, x_value], 'k--')
ax.set_xlim(3800, 550)
ax.set_ylim(0, 0.3)
ax.set_xlabel('Wavenumber $cm^{-1}$', fontsize=18)
ax.set_ylabel('Sensitivity', fontsize=18)
# ax.set_title(('Sensitivity map'))
for mol in [3400, 3276, 2923, 2859, 1901, 1746, 1636, 1539, 1457, 1307, 1154, 1076, 1027, 880]:
l2 = plt.plot([mol, mol], [0, 0.3], 'k', linewidth=1)
ax2 = ax.twiny()
new_tick_loc = [3400, 3276, 2923, 2859, 1901, 1746, 1636, 1539, 1457, 1307, 1154, 1076, 1027, 880]
ax2.set_xlim(ax.get_xlim())
ax2.set_xticks(new_tick_loc)
ax2.set_xticklabels(['$O-H$', '$N-H$', '$C-H_2$', '$C-H_2$', '', '$C=O$', '$C=O$', '$O=C-N$', '$C-CH_3$', '$C-N$',
'$C-O-C$', '$C-O$', '$C-O$', '', '$C-C$'])
plt.setp(ax2.get_xticklabels(), rotation=90)
plt.tight_layout()
plt.savefig((savedir+'Sensitivity_Map_Grouped_Age.png'))
# -
|
CNN/CNN-model/MIMI-Paper-CNN-TCField.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import pandas as pd
from langdetect import detect
import numpy as np
import nltk
from nltk.stem import WordNetLemmatizer
import string
from sklearn.feature_extraction.text import CountVectorizer
import math
import matplotlib.pyplot as plt
lem = WordNetLemmatizer() #create lemmatizer
# +
import ssl
try:
_create_unverified_https_context = ssl._create_unverified_context
except AttributeError:
pass
else:
ssl._create_default_https_context = _create_unverified_https_context
nltk.download('wordnet')
# -
dictionary = pd.read_csv('../data/processedDict.csv')
dictionary['word'] = dictionary['word'].apply(lambda x: lem.lemmatize(x, pos='n'))
dictionary
# +
filepath = '../data/en_reviews/Manchester.csv'
reviews = pd.read_csv(filepath)
# reviews = reviews['date']
reviews
table = str.maketrans('', '', string.punctuation) #mapping to strip punctuation in review
#strip punct of each review -> lemmatise -> output is list of words so join into sentences
reviews['comments'] = reviews.comments.apply(lambda review: ' '.join(map(str, [lem.lemmatize(word.translate(table), pos='n') for word in review.lower().split()])))
reviews
reviews['date'] = pd.to_datetime(reviews['date'])
# -
#### DELETE THIS LATER ###
mask = (reviews['date'] >= '2014-01-01') & (reviews['date'] < '2018-01-01')
reviews = reviews.loc[mask].copy()
reviews
reviews
def get_trends_nice(category, subcats):
years = [2010, 2011, 2012, 2013, 2014, 2015, 2016, 2017, 2018, 2019]
allwords = reviews['comments'].tolist()
allwords = " ".join(allwords)
unique_words = set(allwords.split(' '))
len(unique_words)
unique_words = list(unique_words)
unique_words = [string for string in unique_words if string != ""]
# len(unique_words)
ls = []
for word in unique_words:
word = ''.join([i for i in word if not i.isdigit()])
ls += [word]
unique_words= ls
unique_words = [string for string in unique_words if string != ""]
unique_words = list(dict.fromkeys(unique_words))
def countWords(word, review):
count = 0
for i in review:
if i == word:
count+=1
return count
def getDenom(review, unique_words):
count = 0
den = 0
ls = []
review = review.split()
for word in review:
kmp = countWords(word, review)
if (kmp > 0 and word not in ls):
ls += [word]
den += math.log(1 + kmp)
return den
reviews['den'] = reviews['comments'].apply(lambda x: getDenom(x, unique_words))
def getNom(category, review, dictionary, cat_levl="cat_lev1"):
nom = 0
review = review.split()
dictionaryWords = dictionary[(dictionary[cat_levl] == category)]
dictionaryWords = dictionaryWords['word']
for word in dictionaryWords:
nom += math.log(1 + review.count(word))
return nom
reviews['temp'] = reviews['comments'].apply(lambda x: getNom(category, x, dictionary))
reviews[category] = reviews['temp']*100/reviews['den']
k = {}
for subcat in subcats:
temp = reviews['comments'].apply(lambda x: getNom(subcat, x, dictionary, cat_levl="cat_lev3"))
reviews[subcat] = temp * 100 / reviews["den"]
k[subcat] = reviews[subcat].loc[reviews[subcat] > 0].min()
print(k)
k_business= reviews[category].loc[reviews[category] > 0]
k_business = k_business.min()
print(k_business)
def adoptionForSetOfReviews(category, setOfReviews, dictionary, startDate, endDate, k):
adoption = 1
mask = (setOfReviews['date'] >= startDate) & (setOfReviews['date'] < endDate)
setOfReviews = setOfReviews.loc[mask]
setOfReviews = setOfReviews[category]
if (len(setOfReviews) == 0):
return 0
else:
b = 1/len(setOfReviews)
for review in setOfReviews:
adoption *= math.pow((review + k),b)
adoption = adoption - k
return adoption
d2 = {'year' : years, 'value':0.0}
out = pd.DataFrame(data=d2)
for i in range(len(years)):
out.at[i, "value_{}".format(category)] = adoptionForSetOfReviews(category, reviews, dictionary, "{}-01-01".format(years[i]), "{}-01-01".format(years[i] + 1), k_business)
for subcat in subcats:
out.at[i, "value_{}_{}".format(category, subcat)] = adoptionForSetOfReviews(subcat, reviews, dictionary, "{}-01-01".format(years[i]), "{}-01-01".format(years[i] + 1), k[subcat])
return out
subcats = ["meal", "people", "personality_host", "talking", "sharing"]
soci = get_trends_nice("social", subcats)
soci['coef'] = soci['value_social'] / (soci['value_social_meal'] + soci['value_social_people']+soci['value_social_personality_host'] + soci['value_social_talking'] + soci['value_social_sharing'])
soci
# +
soci['value_social_meal'] = soci['value_social_meal']*soci['coef']
soci['value_social_people'] = soci['value_social_people']*soci['coef']
soci['value_social_personality_host'] = soci['value_social_personality_host']*soci['coef']
soci['value_social_talking'] = soci['value_social_talking']*soci['coef']
soci['value_social_sharing'] = soci['value_social_sharing']*soci['coef']
soci
# +
a = pd.DataFrame(columns=["year", "subcat", "val"])
for subcat in subcats:
ap = {"year": soci["year"], "val": soci["value_social_{}".format(subcat)]}
ap = pd.DataFrame(data=ap)
ap["subcat"] = subcat
a = a.append(ap)
a
# -
pivot_social = a.pivot(index='year', columns='subcat', values='val')
pivot_social
ax = pivot_social.loc[:,['meal','people', 'personality_host', 'sharing', 'talking']].plot.bar(stacked=True, figsize=(10,7), color=['#351800','#D85907','#D88A07','#FFA900','#FFD70F'])
plt.title("Manchester social")
ax.get_legend().remove()
business_subcats = ['location', 'communication', 'personality_host2', 'property_type', 'logistic', 'advice', 'facilities', 'interiors']
busi = get_trends_nice("business", business_subcats)
busi['coef'] = busi['value_business'] / (busi['value_business_location'] + busi['value_business_communication']+busi['value_business_personality_host2'] + busi['value_business_property_type'] + busi['value_business_logistic']+busi['value_business_facilities'] + busi['value_business_advice']+busi['value_business_interiors'])
busi
# +
busi['value_business_communication'] = busi['value_business_communication']*busi['coef']
busi['value_business_location'] = busi['value_business_location']*busi['coef']
busi['value_business_personality_host2'] = busi['value_business_personality_host2']*busi['coef']
busi['value_business_advice'] = busi['value_business_advice']*busi['coef']
busi['value_business_property_type'] = busi['value_business_property_type']*busi['coef']
busi['value_business_facilities'] = busi['value_business_facilities']*busi['coef']
busi['value_business_interiors'] = busi['value_business_interiors']*busi['coef']
busi['value_business_logistic'] = busi['value_business_logistic']*busi['coef']
busi
# +
a = pd.DataFrame(columns=["year", "subcat", "val"])
for subcat in subcats:
ap = {"year": busi["year"], "val": busi["value_business_{}".format(subcat)]}
ap = pd.DataFrame(data=ap)
ap["subcat"] = subcat
a = a.append(ap)
a
# -
pivot_business = a.pivot(index='year', columns='subcat', values='val')
ax = pivot_business.loc[:, ['location', 'communication', 'personality_host2', 'property_type', 'logistic', 'advice', 'facilities', 'interiors']].plot.bar(stacked=True, figsize=(10,7), color=['#143258','#235BA0','#70A9C1','#B1CEDB','#16B9FF','#BCEBFF','#119DAF', '#DFC3D4'])
plt.title("Manchester business")
ax.get_legend().remove()
|
Scripts/categories_pretty.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center>Models and Pricing of Financial Derivatives HW_02</center>
#
# **<center>11510691 程远星$\DeclareMathOperator*{\argmin}{argmin}
# \DeclareMathOperator*{\argmax}{argmax}
# \DeclareMathOperator*{\plim}{plim}
# \newcommand{\using}[1]{\stackrel{\mathrm{#1}}{=}}
# \newcommand{\ffrac}{\displaystyle \frac}
# \newcommand{\asim}{\overset{\text{a}}{\sim}}
# \newcommand{\space}{\text{ }}
# \newcommand{\bspace}{\;\;\;\;}
# \newcommand{\QQQ}{\boxed{?\:}}
# \newcommand{\void}{\left.\right.}
# \newcommand{\Tran}[1]{{#1}^{\mathrm{T}}}
# \newcommand{\d}[1]{\displaystyle{#1}}
# \newcommand{\CB}[1]{\left\{ #1 \right\}}
# \newcommand{\SB}[1]{\left[ #1 \right]}
# \newcommand{\P}[1]{\left( #1 \right)}
# \newcommand{\abs}[1]{\left| #1 \right|}
# \newcommand{\norm}[1]{\left\| #1 \right\|}
# \newcommand{\dd}{\mathrm{d}}
# \newcommand{\Exp}{\mathrm{E}}
# \newcommand{\RR}{\mathbb{R}}
# \newcommand{\EE}{\mathbb{E}}
# \newcommand{\NN}{\mathbb{N}}
# \newcommand{\ZZ}{\mathbb{Z}}
# \newcommand{\QQ}{\mathbb{Q}}
# \newcommand{\AcA}{\mathscr{A}}
# \newcommand{\FcF}{\mathscr{F}}
# \newcommand{\Var}[2][\,\!]{\mathrm{Var}_{#1}\left[#2\right]}
# \newcommand{\Avar}[2][\,\!]{\mathrm{Avar}_{#1}\left[#2\right]}
# \newcommand{\Cov}[2][\,\!]{\mathrm{Cov}_{#1}\left(#2\right)}
# \newcommand{\Corr}[2][\,\!]{\mathrm{Corr}_{#1}\left(#2\right)}
# \newcommand{\I}[1]{\mathrm{I}\left( #1 \right)}
# \newcommand{\N}[1]{\mathcal{N} \left( #1 \right)}
# \newcommand{\ow}{\text{otherwise}}
# \void^\dagger$</center>**
# ## Question 1
#
# A stock price is currently $\$40$. It is known that at the end of $3$ months it will be either $\$45$ or $\$35$. The risk-free rate of interest with ***quarterly compounding*** is $8\%$ per annum. Calculate the value of a $3$-month European put option on the stock with an exercise price of $\$40$. Verify that no-arbitrage arguments and risk-neutral valuation arguments give the same answers.
#
# $\bspace Answer$
#
# >Consider a portfolio with $x$ much bounds and $y$ units of asset which is used to replicate the put, then
# >
# >$$\begin{cases}
# 1.02x + 45y = \P{40 - 45}^+ = 0\\
# 1.02x + 35y = \P{40 - 35}^+ = 5
# \end{cases}$$
# >
# >$$x = \ffrac{5\times45-0\times35} {1.02\times\P{45 - 35}}=22.059,y = \ffrac{0-5} {45-35} =-0.5$$
# >
# >Or, first find the up and down rate, $u=45/40 = 1.125$ and $d = 35/40 = 0.857$, then
# >
# >$$x = \ffrac{1.125\times5 - 0.875\times0} {\P{1.125-0.875} 1.02}=22.059,y = \ffrac{0-5} {\P{1.125-0.875}40} =-0.5$$
# >
# >So the price of the put option: $p = x+yS_0 = 22.059-0.5\times40 = 2.059$
# >
# >***
# >
# >And using the riskless hedging principle, suppose to hedge, we need to *short* $\Delta$ shares, then the terminal payoff would be either $-35\Delta+5$ or $-45\Delta$. If
# >
# >$$-35\Delta + 5 = -45\Delta$$
# >
# >the portfolio would be riskless and the terminal payoff would be $22.5$ with $\Delta = -0.5$. And discount that to time $0$, so we have
# >
# >$$-40\Delta + p = \ffrac{22.5}{1.02} \Rightarrow p = 22.5/1.02 + 40\P{-0.5} = 2.059$$
# >
# >Or by risk-neutral method, first we find the risk-neutral probability $q$:
# >
# >$$q = \ffrac{1.02-0.875}{1.125-0.875} =0.58$$
# >
# >Then the put option price: $p = \ffrac{1}{1.02}\P{0\times 0.58+5\times 0.42}=2.059$
#
# $Remark$
#
# >The risk-free rate of interest with quarterly compounding is $8\%$ per annum, meaning that $e^{rT} = 1.02$.
# ## Question 2
#
# Consider the situation in which stock price movements during the life of a European option are governed by a *two-step binomial tree*. Explain why it is not possible to set up a position in the stock and the option that remains *riskless* for the *whole life* of the option.
#
# $\bspace Answer$
#
# >When the stock price enters the second phase, the stock price will change, usually, so that the riskless hedging principle fails after that. To continue doing so, we need to find the new value of the number of shares to hold.
# ## Question 3
#
# A stock price is currently $\$50$. Over each of the next two $3$-month periods it is expected to go up by $6\%$ or down by $5\%$. The risk-free interest rate is $5\%$ per annum with ***continuous compounding***. What is the value of a $6$-month European call option with a strike price of $\$51$?
#
# $\bspace Answer$
#
# >The risk-neutral probability of an *up move*, $q$, is given by
# >
# >$$q = \ffrac{e^{rT} - d}{u-d} = \ffrac{e^{0.05\times 0.25-0.95}} {1.06-0.95} = 0.569$$
# >
# >And its value keeped for each perioed since the parameters never change. So at the end of the first phase, the option price would be
# >
# >$$c_1\P{1} = e^{-0.05\times0.25}\SB{\P{50\times1.06^2-51}\times q} = 2.911$$
# >
# >And then the first phase, we have
# >
# >$$c_0\P{0} = e^{-0.05\times0.25}\SB{c_1\P{1}\times q} = 1.636$$
# >
# >And about the formula for $q$, here's the proof:
# >
# >$$\begin{cases}
# xe^{rT} + yuS_0 = \P{yuS_0 - K}^+ = c_u\\
# xe^{rT} + ydS_0 = \P{ydS_0 - K}^+ = c_d
# \end{cases}\Rightarrow \begin{cases}
# x = \ffrac{uc_d - dc_u}{\P{u-d}e^{rT}}\leq 0\\
# y = \ffrac{c_u-c_d}{\P{u-d}S_0}\geq 0
# \end{cases}$$
# >
# >Then $c = x+yS_0 = e^{-rT}\P{q\cdot c_u + \P{1-q}c_d}$ where $q = \ffrac{e^{rT} - d}{u-d}$.
# ## Question 4
#
# For the situation considered in HW_2.3, what is the value of a $6$-month European *put* option with a strike price of $\$51$? Verify that the *European call* and *European put* prices satisfy put-call parity. If the put option were *American*, would it ever be optimal to exercise it early at any of the nodes on the tree?
#
# $\bspace Answer$
#
# >We now share the $q$ with the last question so we can directly use the path probability method:
# >
# >$$p = e^{-0.05\times0.5}\SB{\P{1-q}^2\P{51 - 50\times0.95^2} + \binom{2} {1} q\P{1-q}\P{51-50 \times 0.95 \times 1.06}} = 1.375$$
# >
# >And to test the put-call parity:
# >
# >$$1.636 + 51 \times e^{-0.05\times0.5}-1.375-50 \approx 0$$
# >
# >That's it. And if it's American call options (without dividend), we won't early exercise cause it's never optimal while if it's American put option, first, we won't do that if the price goes up, or at the begining. Second, if the price goes down in the first period, at that time the payoff from early exercise is $51-50\times 0.95 = 3.5$ while the American put option value at that time is
# >
# >$$P_1\P{0} = e^{-0.05\times0.25}\SB{\P{1-q}\P{51 - 50\times0.95^2}+q\times\P{51-50 \times 0.95 \times 1.06}} = 2.866 < 3.5$$
# >
# >That's to say that it's better to early exercise the American Put option if the stock price goes down in the first phase.
# ## Question 5
#
# A stock price is currently $25$ bucks. It is known that at the end of $2$ months it will be either $\$23$ or $\$27$. The risk-free interest rate is $10\%$ per annum with continuous compounding. Suppose $S_T$ is the stock price at the end of $2$ months. What is the value of a derivative that pays off $S^2_T$ at this time?
#
# $\bspace Answer$
#
# >We replicate that portfolio with a claim which is priced as $V_t^h$, with $x$ amount of cash and $y$ amount of the stocks. Then using no arbitrage pricing, we have
# >
# >$$\begin{cases}
# x\cdot e^{0.1\times 1/6} + y \times 27 = 27^2 \\
# x\cdot e^{0.1\times 1/6} + y \times 23 = 23^2
# \end{cases} \\[2em]
# \Rightarrow x= \ffrac{-27\times23} {e^{0.1 \times 1/6}},\bspace y = 50$$
# >
# >So that the price of that claim would be
# >
# >$$\Pi\P{\ffrac{1} {6}, U} = V_{1/6}^{h} = x + y\cdot S_0 = \ffrac{-27\times23} {e^{0.1 \times 1/6}} + 50 \times 25 = 639.264$$
# >
# >***
# >
# >Or using the riskless hedging principle, consider portfolio that longs this derivative and short $\Delta$ shares. The value at time $T$ is $27^2 - 27\Delta = 23^2 - 23\Delta$ and thus $\Delta = 50$, the value is $-621$. Let $f$ be the price of the derivative at time $0$, then
# >
# >$$\P{f - 25\Delta}e^{rT} = \P{f- 50\times 25}e^{0.1\times 1/6} = -621 \Rightarrow f = 639.264$$
# ## Question 6
#
# A stock price is currently $\$40$. Over each of the next two $3$-month periods it is expected to go up by $10\%$ or down by $10\%$. The risk-free interest rate is $12\%$ per annum with continuous compounding.
#
# $\P{\text a}$ What is the value of a $6$-month European put option with a strike price of $\$42$?
#
# $\bspace Answer$
#
# >$$q = \ffrac{e^{0.12 \times 0.25} - 0.9} {1.1-0.9} = 0.652$$
# >
# >The European put option can be find similarly:
# >
# >$$p = e^{-0.12\times 0.5} \SB{\P{1-q}^2 \P{42-40\times0.9^2} + \binom{2} {1} \P{1-q}q\P{42-40 \times 0.9 \times 0.1}}=2.121$$
#
# $\P{\text b}$ What is the value of a $6$-month American put option with a strike price of $\$42$?
#
# $\bspace Answer$
#
# >We will calculate the American Put option backwards.
# >
# >$$\begin{align}
# P_{1}\P{1} &= \max\P{e^{-0.12\times 0.25}\SB{\P{1-q}\P{42-40 \times 0.9 \times 1.1}},0} = 0.811 \\
# P_{1}\P{0} &= \max\P{e^{-0.12\times 0.25}\SB{q\P{42-40 \times 0.9 \times 1.1} + \P{1-q}\P{42-40\times0.9^2}},42-40\times0.9} = 6 \\
# P_{0}\P{0} &= \max\P{e^{-0.12\times 0.25}\SB{q\cdot P_{1}\P{1} + \P{1-q}\cdot P_{1}\P{0}},42-40} = 2.539
# \end{align}
# $$
# ## Question 7
#
# A stock price is currently $30$ bucks. During each $2$-month period for the next $4$ months it will increase by $8\%$ or reduce by $10\%$. The risk-free interest rate is $5\%$. Use a two-step tree to calculate the value of a derivative that pays off $\P{\max\P{30 − S_T , 0}}^2$ , where $S_T$ is the stock price in $4$ months. If the derivative is American-style, should it be exercised early?
#
# $\bspace Answer$
#
# >It's not hard to find the value of the tree in the last column: $0$, $\P{30-29.16}^2 = 0.7056$ and $\P{30-24.3}^2 = 32.49$. Then we calculate the probability $q$:
# >
# >$$q = \ffrac{e^{0.05 \times 1/6}-0.9} {1.08-0.9} = 0.602$$
# >
# >Then for the European styled derivatives, the option value is:
# >
# >$$e^{-0.05\times1/3}\SB{\P{1-q}^2\times 32.49+\binom{2} {1}\P{1-q}q\times0.7056} = 5.394$$
# >
# >While for the American styled, firstly, like the same, it won't happen at the end of second phase, nor the begining. Then at the end of the first phase, if the stock price goes up in the first period, then still nothing should be done. If it goes down, then the payoff from early exercise is $\P{30-30\times0.9}^2=9$ however the value of American Put option at that time is
# >
# >$$P_{1}\P{0} = \max\P{e^{-0.05\times1/6}\SB{q\times 0.7056 + \P{1-q}\times 32.49},9} = 13.245>9$$
# >
# >So, even if it's an American styled derivatives, at no point of time should you early exercise it.
|
FinMath/Models and Pricing of Financial Derivatives/HW/HW_02_11510691_fixed.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
s1 = np.random.randint(10, size = 5)
s2 = np.random.randint(10, size = 5)
s3 = np.random.randint(10, size = 5)
# +
sozluk = {"var1" : s1,
"var2" : s2,
"var3" : s3
}
sozluk
# -
df = pd.DataFrame(sozluk) #sozluk uzerinden dataFrame olusturma;
df
df[0:1]
df.index = ["a", "b", "c", "d", "e"]
df
# +
#silme
# -
df.drop("a", axis = 0) #a satırını sil. AMA ana yapı üzerinde degisiklik yapılmadı
#orjinal df aynı
df
# +
#ancak kalıcı olarak gitsin istiyorsa inplace = true kullanırız
# -
df.drop("a", axis = 0, inplace = True)#inplace true old. icin a satırı kalıcı olarak gitti
df
# +
#fancy
# -
silinecek_indexler = ["c", "e"]
df.drop(silinecek_indexler, axis = 0)
# +
#degsikenler icin
# -
"var1" in df
degiskenler = ["var1", "var4", "var2"]
for i in degiskenler:
print(i in df) #df'in icinde gez, degiskenler listesindeki her bir degiskeni tek tek sor var mı diye
df
# +
#yeni degisken olusturma
# -
df["var4"] = df["var1"]*df["var2"]# !!! var4 olmadığı icin olusturmaya karar verir.
df["var5"] = df["var1"]*df["var3"]# !!! var5 olmadığı icin olusturmaya karar verir.
df
# +
#degisken silme;
# -
df.drop("var4", axis = 1)
df
df.drop("var4", axis = 1, inplace=True)
silinecekler = ["var1" ,"var2"] #fancy index
df.drop(silinecekler, axis = 1)#fancy index
|
pandas_dataFrame_eleman_islemleri.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:hetmech] *
# language: python
# name: conda-env-hetmech-py
# ---
# # Metapath Rank Comparison
# Compare the rankings the unsupervised methods in hetmech assign to different epilepsy/drug metapaths to the supervised ones from rephetio
# +
import math
from matplotlib import pyplot as plt
import pandas
import plotnine as p9
from scipy import stats
import sys
sys.path.append("..")
from src.database_utils import get_db_connection
# -
# %%time
epilepsy_id = 'DOID:1826'
# Get top ten most important metapaths for epilepsy (which are all compound-disease pairs)
query = f'''
SELECT
outer_pc.dwpc AS dwpc,
outer_pc.p_value AS p_value,
outer_pc.metapath_id AS metapath_id,
top_ids.source_name AS source_name,
top_ids.source_identifier AS source_identifier,
top_ids.target_name AS target_name,
top_ids.target_identifier AS target_identifier
FROM
(SELECT dwpc, p_value, metapath_id, source_id, target_id, n1.name AS source_name, n1.identifier AS source_identifier,
n2.name AS target_name, n2.identifier AS target_identifier
FROM dj_hetmech_app_pathcount pc
JOIN dj_hetmech_app_node join_node
ON pc.target_id=join_node.id OR pc.source_id=join_node.id
JOIN dj_hetmech_app_node n1
ON pc.source_id = n1.id
JOIN dj_hetmech_app_node n2
ON pc.target_id = n2.id
WHERE join_node.identifier='{epilepsy_id}' AND (n1.metanode_id = 'Compound' OR n2.metanode_id = 'Compound')
ORDER BY pc.p_value) AS top_ids
JOIN dj_hetmech_app_pathcount outer_pc
ON (top_ids.source_id = outer_pc.source_id AND
top_ids.target_id = outer_pc.target_id) OR
(top_ids.source_id = outer_pc.target_id AND
top_ids.target_id = outer_pc.source_id)
ORDER BY outer_pc.p_value;
'''
with get_db_connection() as connection:
metapaths = pandas.read_sql(query, connection)
# +
metapaths = metapaths.sort_values(by=['source_name', 'metapath_id'])
# Ensure that you only have one copy of each (source_name, metapath_id) pair
metapaths = metapaths.drop_duplicates(subset=['source_name', 'metapath_id'])
# Remove any rows with NaN values
metapaths = metapaths.dropna()
min_p_value = metapaths[metapaths.p_value != 0].p_value.min()
metapaths.loc[metapaths.p_value == 0, 'p_value'] = min_p_value
metapaths['neg_log_p_value'] = metapaths.p_value.apply(lambda x: -math.log10(x))
metapaths.head()
# -
# %%time
path = 'https://github.com/dhimmel/learn/raw/8792c2e408e790cd8d77adb34d013961f4d5c4f0/prediction/predictions/term-contribution.tsv.bz2'
supervised_df = pandas.read_table(path)
supervised_df = supervised_df[supervised_df['disease_id'] == epilepsy_id]
supervised_df.head()
reformatted_supervised_df = pandas.melt(
frame=supervised_df,
id_vars=['compound_id', 'disease_id'],
var_name='metapath_id',
value_name='metapath_weight',
)
reformatted_supervised_df.head()
# +
# Drop all rows with negative or zero weights
positive_supervised_df = reformatted_supervised_df[(reformatted_supervised_df['metapath_id'].str.contains('dwpc_')) &
(reformatted_supervised_df['metapath_weight'] > 0)].copy()
positive_supervised_df['metapath_id'] = reformatted_supervised_df['metapath_id'].str.replace('dwpc_', '')
# Remove CtDrD, there are only 7 results in the merged dataset and they break the loess curves later
positive_supervised_df = positive_supervised_df[~ positive_supervised_df['metapath_id'].str.contains('CtDrD')]
# It's worth noting that there aren't any length one paths in the term contributions dataset
positive_supervised_df.head()
# -
merged_df = metapaths.merge(
positive_supervised_df,
left_on=['metapath_id', 'source_identifier', 'target_identifier'],
right_on=['metapath_id', 'compound_id', 'disease_id'],
)
merged_df.head()
plt.scatter(merged_df.neg_log_p_value, merged_df.metapath_weight, alpha=.2, s=5)
plt.title('Rephetio Metapath Weights vs Metapath Negative Log P-Values')
plt.xlabel('Metapath Negative Log P Value')
plt.ylabel('Rephetio Metapath Weight')
plt.scatter(merged_df.p_value, merged_df.metapath_weight, alpha=.2, s=5)
plt.title('Rephetio Metapath Weights vs Metapath P-Values')
plt.xlabel('Metapath P-Value')
plt.ylabel('Rephetio Metapath Weight')
colors = p9.scale_color_hue().palette(9)
show_metapaths = ['CiPCiCtD', 'CrCtD', 'CbGaD', 'CbGbCtD', 'CrCrCtD', 'CpDpCtD', 'CcSEcCtD', 'CbGeAlD', 'CrCbGaD']
(
p9.ggplot(merged_df)
+ p9.geom_point(p9.aes('neg_log_p_value', 'metapath_weight', color='metapath_id'), alpha=.4)
+ p9.scale_color_manual(colors, breaks=show_metapaths)
+ p9.ggtitle('Rephetio Metapath Weights vs Metapath P-Values')
)
(
p9.ggplot(merged_df, p9.aes('neg_log_p_value', 'metapath_weight', color='metapath_id', fill='metapath_id'))
+ p9.geom_smooth(method="loess")
+ p9.scale_color_manual(colors, breaks=show_metapaths)
+ p9.ggtitle('Rephetio Metapath Weights vs Metapath P-Values')
+ p9.guides(fill=False)
)
|
explore/metapath_rank_comparison.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Monitoring Python Applications with Elastic APM
# ## Medium Link: <https://medium.com/p/33237a39d7b6>
# ## Monitoring Flask / Flask-RESTPlus Applications
# ### Installation
# ```bash
# pip install elastic-apm[flask]
# ```
# ### Implementation
# ```python
# from flask import Flask
# from elasticapm.contrib.flask import ElasticAPM
# import elasticapm
#
# server_url = 'http://localhost:8200'
# service_name = 'DemoFlask'
# environment = 'dev'
#
# app = Flask(__name__)
# apm = ElasticAPM(app, server_url=server_url, service_name=service_name, environment=environment)
#
#
# @app.before_request
# def apm_log():
# elasticapm.label(platform = 'DemoPlatform',
# application = 'DemoApplication')
#
#
# @app.route('/hello-world/')
# def helloWorld():
# return "Hello World"
#
#
# app.run()
# ```
# ## Monitoring FastAPI Applications
# ### Installation
# ```bash
# pip install elastic-apm
# ```
# ### Implementation
# ```python
# import uvicorn
# from fastapi import FastAPI
#
# from elasticapm.contrib.starlette import make_apm_client, ElasticAPM
#
#
# apm_config = {
# 'SERVICE_NAME': 'DemoFastAPI',
# 'SERVER_URL': 'http://localhost:8200',
# 'ENVIRONMENT': 'dev',
# 'GLOBAL_LABELS': 'platform=DemoPlatform, application=DemoApplication'
# }
#
# apm = make_apm_client(apm_config)
#
# app = FastAPI()
# app.add_middleware(ElasticAPM, client=apm)
#
#
# @app.get('/hello-world/')
# def hello_world():
# return "Hello World"
#
#
# uvicorn.run(app)
# ```
# ## Monitoring Python Applications
# ### Installation
# ```bash
# pip install elastic-apm
# ```
# ### Implementation
# ```python
# import time
# from apscheduler.schedulers.background import BackgroundScheduler
#
# from elasticapm import Client
# import elasticapm
#
# client = Client(
# {'SERVICE_NAME': 'DemoPython',
# 'SERVER_URL': 'http://localhost:8200',
# 'ENVIRONMENT': 'dev'}
# )
#
# elasticapm.instrumentation.control.instrument()
#
# def hello_world():
# client.begin_transaction('schedule')
# elasticapm.label(platform='DemoPlatform', application='DemoApplication')
#
# print("Hello World")
#
# client.end_transaction('demo-transaction', 'success')
#
#
# if __name__ == "__main__":
# scheduler = BackgroundScheduler()
# scheduler.add_job(hello_world, 'interval', seconds=5)
# scheduler.start()
# print('Started scheduler..')
# print('Press Ctrl+C to exit')
# try:
# # Necessary to simulate application activity (which keeps the main thread alive).
# while True:
# time.sleep(2)
# except (KeyboardInterrupt, SystemExit):
# print('Shutting down scheduler')
# scheduler.shutdown()
# ```
|
Monitoring Python Applications with Elastic APM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import axon
from decimal import Decimal
from datetime import datetime, time, date
text = axon.dumps([['abc абв', 1, 3.14, True], [datetime.now(), Decimal('3.14')]])
print(text)
vals = [
{'id':1, 'nickname':'nick', 'time':time(12, 31, 34), 'text':'hello!'},
{'id':2, 'nickname':'mark', 'time':time(12, 32, 3), 'text':'hi!'}
]
text = axon.dumps(vals)
print(text)
text = axon.dumps(vals, pretty=1)
print(text)
vals == axon.loads(text)
vals = [[{'a':1, 'b':2, 'c':3}, {'a':[1,2,3], 'b':(1,2,3), 'c':{1,2,3}}]]
text = axon.dumps(vals)
print(text)
text = axon.dumps(vals, pretty=1)
print(text)
vals == axon.loads(text)
vals = axon.loads('person{name:"nick" age:32 email:"<EMAIL>"}')
print(type(vals[0]))
print(vals[0])
text = axon.dumps(vals)
print(text)
text = axon.dumps(vals, pretty=1)
print(text)
text = axon.dumps(vals, pretty=1, braces=1)
print(text)
# +
class Person:
def __init__(self, name, age, email):
self.name = name
self.age = age
self.email = email
def __str__(self):
return "Person(name=%r, age=%r, email=%r)" % (self.name, self.age, self.email)
@axon.reduce(Person)
def reduce_Person(p):
return axon.node('person', {'name':p.name, 'age':p.age, 'email':p.email})
@axon.factory('person')
def factory_Person(attrs, vals):
return Person(name=attrs['name'], age=attrs['age'], email=attrs['email'])
# -
p = Person('nick', 32, '<EMAIL>')
text = axon.dumps([p])
print(text)
val = axon.loads(text, mode='strict')[0]
print(val)
print(val.name==p.name, val.age==p.age, val.email==p.email)
|
examples/readme.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# name: ir
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/anicelysantos/tutorial_r/blob/main/pacotes_carga_dados.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="re-0qzuXW7yx"
library(readr)
# + [markdown] id="5GLIG-f__Pj4"
# # Arquivos .csv e .txt
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="MRjaJYWmbIkK" outputId="87a8d223-32a7-4a55-c3df-426b2a18eacd"
#arquivos separados por vírgula
imdb_csv <- read.csv(file = '/content/sample_data/imdb.csv')
head(imdb_csv)
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="-jgupAHK9sA1" outputId="ca8ee839-ab3f-4208-bd5e-796e809323fe"
#arquivos separados por ponto e virgula
imdb_csv2 <- read.csv2('/content/sample_data/imdb2.csv')
head(imdb_csv2)
# + colab={"base_uri": "https://localhost:8080/", "height": 688} id="SDRSmXkq-Wrb" outputId="8fe049fa-0302-4b0b-a761-e2c398610cf5"
#Arquivos txt, o \t é a tabulação
imdb_txt <- read_delim('/content/sample_data/imdb.txt')
head(imdb_txt)
# + [markdown] id="OZ-YcZdB_KwX"
# # Locale
# + colab={"base_uri": "https://localhost:8080/", "height": 226} id="V-SsKqQs_BEF" outputId="1108caef-8691-428c-ace6-626aaf42510f"
locale()
# + colab={"base_uri": "https://localhost:8080/", "height": 226} id="mPvSHCUn_bmi" outputId="6b92bc9f-dedd-4032-d4fa-64d745b1669c"
locale(date_names = 'pt')
# + colab={"base_uri": "https://localhost:8080/", "height": 226} id="N2RdfDeS_jIR" outputId="44dcc79d-88eb-4809-cee2-946a815c225b"
locale(decimal_mark = ',')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="eq1YaT5n_qiq" outputId="48dc661c-a7b9-41ae-e19d-267e4f7796d1"
frase_com_acentos <- 'você comerá uma maçã amanhã à tarde'
Encoding(frase_com_acentos)
# + id="B8hBqPeL__6b"
Encoding(frase_com_acentos) <- 'latin1'
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="PB685eoqAICJ" outputId="76a5c553-3e6f-4634-be0d-34fac1b23d58"
frase_com_acentos
# + colab={"base_uri": "https://localhost:8080/", "height": 688} id="WjurZd9AAP_q" outputId="cabf7085-53b3-4b08-e7c7-61255639bc05"
outro_encoding <- read_csv('/content/sample_data/imdb.csv', locale = locale(encoding = 'latin1'))
head(outro_encoding)
# + [markdown] id="mw7Zc6opA0o7"
# # Parseando valores
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="m2PrW6NIAiyy" outputId="c73044eb-8ab5-4635-8458-8ff3c7a16a91"
parse_number(c('5','5.0','5,0','R$5.00','5a'))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="smkFN9ItBMQe" outputId="7dce3caf-fd92-4561-f7f0-b2ffccfc03a3"
parse_number('5,0', locale = locale(decimal_mark = ','))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="caH9nlwTBc0a" outputId="988e2fd5-21e9-4278-824d-0c54732fb1cf"
#Inglês
parse_date(
'01/june/2010',
format = '%d/%B/%Y'
)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="wWY8qsVrBvya" outputId="633cf00f-d174-4f03-fbc2-e7aa2fd598b5"
#Português
parse_date(
'01/junho/2021',
format = '%d/%B/%Y',
locale = locale(date_names = 'pt')
)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="RzT59YvMCByy" outputId="28cc4cd3-6213-4e6e-cb45-625c6309f94f"
#Pode especificar NAs
parse_number(c('5', '5.0', '5,0', 'R$5.00', '5 a'), na = '5 a')
# + [markdown] id="Ahpp57U_CoAo"
# Outras funções úteis:
# * `readr::parse_integer()`, para parsear números inteiros
# * `readr::parse_character()`, para parsear strings
# * `readr::parse_date()`, `readr::parse_time()`, `readr::parse_datetime()` para parsear datas, horas e data/horas.
# + [markdown] id="qrDejRt9DP-o"
# # Salvando arquivos de texto
# + id="kswVJO0-ClBM"
#arquivo .csv
write_csv(x = mtcars, file='/content/sample_data/mtcars.csv')
# + id="esWrFaNqDmVp"
#Base separada por tabulação
write_delim(x = mtcars, file='/content/sample_data/mtcars.txt', delim='\t')
# + [markdown] id="HOlUVzkMD5hS"
# # Arquivos .rds
# + id="c0rtTLUgD3QE"
#Esses arquivos só podem ser lidos dentro do R
write_rds(mtcars, file='mtcars.rds', compress='gz')
# + colab={"base_uri": "https://localhost:8080/", "height": 375} id="wvRXRV8gEO2q" outputId="0e214655-9f29-401b-847a-729b50fce5c2"
imdb_rds <- read_rds(file='mtcars.rds')
head(imdb_rds)
# + [markdown] id="WiEBOhkHEjWc"
# # Exercicios
# + [markdown] id="b_rPJxmjEsmU"
# 1. Qual a diferença entre as funções `read_csv()` e `read_csv2()`?
#
# A primeira lê arquivos separados por vígula e a segunda separados por ponto e vírgula
# + [markdown] id="u7G8eg2sE7_h"
# 2. Leia o arquivo imdb.csv utilizando a função read_delim().
# + colab={"base_uri": "https://localhost:8080/", "height": 688} id="tLfdyDuxEq_Q" outputId="41f8df6e-80ca-435c-a868-5d78991fd6df"
leia <- read_delim('/content/sample_data/imdb.csv', delim = ',')
head(leia)
# + [markdown] id="59uv7tw1FPeg"
# 3. Escreva a base `mtcars` em um arquivo `mtcars.csv` que não contenha o nome das colunas.
# + id="PuwyqumHEbRL"
write_csv(mtcars,'sc_mtcars.csv', col_names=FALSE)
# + [markdown] id="QnSOJY4iF-85"
# 4. Use a função `write_rds()` para salvar em arquivos:
#
# a) um número
# + id="sh3jXyKIFqW0"
numero <- 5
write_rds(numero,'numero.rds')
# + [markdown] id="yylacixYG24I"
# b) um vetor de strings
# + id="8PWEOy84G2Gj"
estados <- c('PE', 'CE', 'PB','RN')
write_rds(estados, 'estados.rds')
# + [markdown] id="h3sUCgbOHMqY"
# Uma lista com valores numéricos, textuais e lógicos
# + id="cUSVpAVcHHmg"
lista <- list(numero = 1, texto = c('M', 'F', 'F'), logico = FALSE)
write_rds(lista,'lista.rds')
# + [markdown] id="aDQzOfx8HiVk"
# d) As 3 primeiras colunas da base `mtcars`
#
# + id="Iuxg7TDCHgSc"
write_rds(mtcars[,1:3], 'mtcars_3_colunas.rds')
# + [markdown] id="dfxbOe5THyV1"
# 5. Utilize a função `read_rds()` para importar de volta para o R os arquivos criados no exercício 4.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="8Q1iR8oQHwz0" outputId="8e5fd848-04f4-4900-dfcb-d2511dbdb93c"
read_rds('numero.rds')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="ZEHd9KfAH69z" outputId="66ebbcaa-abaf-4a73-87df-b596c6ab985e"
read_rds('estados.rds')
# + colab={"base_uri": "https://localhost:8080/", "height": 149} id="O2ncibiXH9Zl" outputId="ec2ee8a1-d3ed-494b-aa83-17d4c74b94ab"
read_rds('lista.rds')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Tp_gSxXTIEm1" outputId="0c316782-1ea2-4d69-9823-1612735d0499"
read_rds('mtcars_3_colunas.rds')
|
pacotes_carga_dados.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lmcanavals/algorithmic_complexity/blob/main/notebooks/bt_edmonds_karp.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="mmHU8WqamaVD" outputId="e9f1139a-8ecf-43c2-ad78-8c54fe9dc0e3"
# !git clone https://github.com/lmcanavals/algorithmic_complexity.git
# + id="rWiDhwV_mqNl"
from algorithmic_complexity.aclib import graphstuff as gs
import networkx as nx
import numpy as np
# + id="gZkf-rO5mx50"
def bfs(G, s, t):
n = len(G)
visited = [False]*n
path = [-1]*n
q = [s]
visited[s] = True
while q:
u = q[0]
if u == t:
break
q = q[1:]
for v in range(n):
if G[u][v] > 0 and not visited[v]:
visited[v] = True
path[v] = u
q.append(v)
if path[t] == -1:
return None, 0
bottleneck = np.Inf
cpath = []
while t != -1:
cpath = [t] + cpath
if path[t] > -1:
bottleneck = bottleneck if bottleneck < G[path[t], t] else G[path[t], t]
t = path[t]
return cpath, bottleneck
# + id="IvSamUlwtFPk"
def edmondsKarp(G, s, t):
n = len(G)
Gres = G.copy()
Gflow = np.zeros((n, n))
while True:
path, bottleneck = bfs(Gres, s, t)
if path != None:
for i in range(1, len(path)):
u = path[i - 1]
v = path[i]
Gres[u][v] -= bottleneck
Gres[v][u] += bottleneck
Gflow[u][v] = Gflow[u][v] - Gflow[v][u] + bottleneck
else:
break
return Gflow, np.sum(Gflow[s])
# + [markdown] id="8TEfn9ejoZ6q"
# Node | Number
# -- | --
# S | 0
# a | 1
# b | 2
# c | 3
# d | 4
# t | 5
# + colab={"base_uri": "https://localhost:8080/"} id="b61_7vU8oP7s" outputId="e3e635ea-3c0a-4d52-ab52-d0d4f7bf2332"
# %%file 1.adjmatrix
0 16 0 13 0 0
0 0 12 10 0 0
0 0 0 9 0 20
0 4 0 0 14 0
0 0 7 0 0 4
0 0 0 0 0 0
# + colab={"base_uri": "https://localhost:8080/"} id="XKfHyM-fpbFU" outputId="318939cb-af1f-4054-e08d-36a7a72acfa1"
G = np.fromfile("1.adjmatrix", sep=" ").reshape((6, 6))
G
# + colab={"base_uri": "https://localhost:8080/", "height": 209} id="JJh0qv0mpniL" outputId="6420b908-530f-425a-84f0-26b3bae309a5"
Gnx = nx.from_numpy_matrix(G, create_using=nx.DiGraph)
gs.nx2gv(Gnx, weighted=True)
# + colab={"base_uri": "https://localhost:8080/"} id="1ahyTikYpzBe" outputId="f984b3ce-ab53-42df-9ea2-0c7b686f92e0"
bfs(G, 0, 5)
# + colab={"base_uri": "https://localhost:8080/", "height": 139} id="JSQ8KJBcqRW1" outputId="6fc96cbc-0ed6-4b04-d0f4-4853a69d8bc9"
Gf, fm = edmondsKarp(G, 0, 5)
print(fm)
Gnx = nx.from_numpy_matrix(Gf, create_using=nx.DiGraph)
gs.nx2gv(Gnx, weighted=True)
# + id="xwF3X-otvHGM"
|
notebooks/bt_edmonds_karp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_style(style="whitegrid")
sns.set(font_scale=2)
import pandas as pd
import re
# -
df = pd.read_csv("datas/clinvar_conflicting.csv", dtype= {"CHROM":str, 38:str, 40:object})
df.head()
df.CHROM.nunique() # Total Number of Chromosome inside VCF File
df.CHROM.unique()
# list of Unique Chromosomes
df.shape
a = df.groupby(["CHROM","POS", "REF", "ALT"])
a.ngroups
pd.DataFrame(a)
df.CHROM.value_counts()
# +
## The class distribution is skewed a bit to the 0 class,
# meaning there are fewer variants with conflicting submissions.
# -
ax = sns.countplot(x="CLASS", data=df)
ax.set(xlabel = "CLASS", ylabel = "Number of Variants")
# +
# From the above Figure Its clear that conflicting variants are more common in some genens.
# -
gene_ct = pd.crosstab(df.SYMBOL, df.CLASS, margins= True)
gene_ct
gene_ct = pd.crosstab(df.SYMBOL, df.CLASS, margins=True)
gene_ct
df.head()
df.columns
# +
gene_ct = pd.crosstab(df.SYMBOL, df.CLASS, margins=True)
gene_ct.drop("All", axis = 0, inplace = True)
# Limit to the 50 Most submitted genes for visualization
gene_ct = gene_ct.sort_values(by = "All", ascending = False).head(20)
gene_ct.drop("All", axis = 1, inplace = True)
gene_ct.plot.bar(stacked = True, figsize = (12,4))
# -
# ### CLNVC (Variant Type)
# +
vt_ct = pd.crosstab(df.CLNVC, df.CLASS, margins= True)
vt_ct.drop("All", axis = 0, inplace = True)
# Limit to the 50 most submitted genes for visualization
vt_ct = vt_ct.sort_values(by = "All", ascending = False)
vt_ct.drop("All", axis = 1, inplace = True)
vt_ct.plot.bar(stacked = True, figsize = (12,4))
# -
# Exons are features of genes that map sequence nucleotides functional parts of DNA. Genes have differing numbers of exons, some have few , some have many. Lets see if , regardless of gene wheather or not conflicting variantgs are enriched in a general exon location
df.EXON.head()
print(df.shape)
x = (lambda x: [int(s) for s in re.findall(r"\b\d+\b",x)][0])
x(df.EXON[1])
df.EXON.fillna("0", inplace=True)
df["variant_exon"] = df.EXON.apply(lambda x: [int(s) for s in re.findall(r"\b\d+\b",x)][0])
# `variant_exon` = 0 represents that the variant is located in an Intron variants seem to be conflicting much more frequently thatn exon variants
df.columns
exondf = pd.crosstab(df["variant_exon"], df["CLASS"])
exondf.plot.bar(stacked = True, figsize = (20,5))
plt.xlim(-0.5, 20.5)
# parse and encode the MC (Molecular Consequence) fild
# +
MC_list = df.MC.dropna().str.split(",").apply(lambda row: list(c.split("|")[1] for c in row))
MC_encoded = pd.get_dummies(MC_list.apply(pd.Series).stack()).sum(level = 0)
MC_encoded = MC_encoded.reindex(index = MC_list.index)
# Incorporate the transformed MC feature into the existinf Data Frame
df = df.join(MC_encoded).drop(columns=["MC"])
# Transformed MC feature
MC_encoded.head()
# -
# Manually generate crosstab, there is probably a faster method via pandas
# +
mccounts = {0:{},
1:{},
"All": {}
}
for col in MC_encoded.columns:
for class_ in [0,1]:
mccounts[class_][col] = df.loc[df["CLASS"] == class_][col].sum()
mccounts["All"][col] = df[col].sum
mc_ct = pd.DataFrame.from_dict(mccounts)
mc_ct_all = mc_ct.sum(axis = 0)
mc_ct_all.name = "All"
mc_ct = mc_ct.append(mc_ct_all, ignore_index = False)
mc_ct.drop("All", axis = 1, inplace = True)
mc_ct.plot.bar(stacked = True, figsize = (12,4))
# -
# results from SIFT and PolyPhen Software that predict the severity of a variant, in-silico.
# +
sift_ct = pd.crosstab(df.SIFT, df.CLASS, margins=True)
sift_ct.drop("All", axis = 0, inplace = True)
# Limit to the 50 Most submitted genes for Visualization
sift_ct = sift_ct.sort_values(by = "All", ascending = False)
sift_ct.drop("All", axis = 1, inplace = True)
sift_ct.plot.bar(stacked = True, figsize = (12,4))
# -
# ##### PolyPhen
# +
pp_ct = pd.crosstab(df.PolyPhen, df.CLASS, margins= True)
pp_ct.drop("All", axis = 0, inplace = True)
# Limit to the 50 most submitted genes for visualization
pp_ct = pp_ct.sort_values(by = "All", ascending = False)
pp_ct.drop("All", axis = 1, inplace = True)
pp_ct.plot.bar(stacked = True, figsize = (12,4))
# -
# #### Encode Sift and PolyPhen
df = pd.get_dummies(df, columns=["SIFT", "PolyPhen"])
# Correlation for categorical features by way of chi-square test
from itertools import combinations
from scipy.stats import chi2_contingency
# Select a few categorical Features
categoricals_index = pd.MultiIndex.from_tuples(combinations(["CHROM", "REF", "ALT", "IMPACT","Consequence",\
"SYMBOL", "CLASS"],2))
categorical_corr = pd.DataFrame(categoricals_index, columns= ["cols"])
def chisq_of_df_cols(row):
c1, c2 = row[0], row[1]
groupsizes = df.groupby([c1,c2]).size()
ctsum = groupsizes.unstack(c1)
# fillna (0) is necessary to remove any NAs which will cause exceptions
return chi2_contingency(ctsum.fillna(0))[1]
categorical_corr["chi2_p"] = categorical_corr.cols.apply(chisq_of_df_cols)
categorical_corr
categorical_corr.index = categoricals_index
categorical_corr = categorical_corr.chi2_p.unstack()
# I tried plotting a heatmap with -np.log(p) but it didnot liik good as visualization.
categorical_corr
# THis Seems to meaning less.
# +
# The dark blue box in the heatmap highlights the negative correlation with the
# allele frequency features. Common alleles are less likely to pathogenic (cause disease)
# therefore most labs agrees they should be benign
# +
import numpy as np
corr = df.select_dtypes(exclude="object").corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype = np.bool)
mask[np.triu_indices_from(mask)] = True
# Setup the matplotlib figure
f, ax = plt.subplots(figsize = (15,12))
## Generating a custom diverging_paletter(220, 10, as_cmap = True)
cmap = sns.diverging_palette(220,10, as_cmap=True)
## Draw the heatmap with the mask and correct aspect ratio
g = sns.heatmap(corr, mask = mask, cmap=cmap, vmax=0.5, center= 0,
square=True, linewidths=0.5, cbar_kws={"shrink":0.5})
from matplotlib.patches import Rectangle
g.add_patch(Rectangle((1,6),3,1, fill = False, edgecolor = "blue", lw = 4))
# -
# One of the ways variants can be classified is by the amount (and type) of sequence change.<br>
# A substitution of a nucleotide (letter) is considered a single nucleotide variant (SNV), these are sometimes referred to as <br> **Single Nucleotide Polymorphisms(SNP).** <br><br>
#
# When one or more nucleotides are inserted or deleted the variant is considered an insertion or deletion. There fore, if the <br> if the length of `REF` or `ALT` is >1 then the variant can be considered an Insertion or Deletion (indel), other wise it can be considered a SNV.
snvs = df.loc[(df.REF.str.len() == 1) & (df.ALT.str.len()== 1)]
indels = df.loc[df.REF.str.len()>1 | (df.ALT.str.len()>1)]
# +
print(len(snvs) + len(indels))
print(len(df))
# Generally The len of SNV Plus Indels will be the same
len(df) == len(snvs) + len(indels)
# -
df.loc[(df.REF.str.len() >1)]
# SNVs are more likely to be conflicting than Indels
snp_indel = pd.concat([
snvs.CLASS.value_counts(normalize = True).rename("snv_class"),
indels.CLASS.value_counts(normalize = True).rename("indel_class")],
axis=1).T
snp_indel
snp_indel.plot.area()
snp_indel.plot.bar(stacked = True, figsize = (12,4))
# `CLNDN` are lists of diseases associated with the variant. It may be beneficial to treat both `not_specified`
# and / or `not_provided` as the same category.
clndn = pd.concat([df.CLASS.loc[(df.CLNDN == "not_specified") | (df.CLNDN == "not_provided") | (df.CLNDN == "not_specified|notprovided")].value_counts(normalize=True)
.rename("Disease_not_specified"), df.CLASS.loc[(df.CLNDN != "not_specified") | (df.CLNDN != "not_Provided") | (df.CLNDN != "not_specified|not_provided")].value_counts(normalize=True).rename("Some_disease_specified")], axis=1).T
clndn
clndn.plot.bar(stacked=True, figsize = (12,4))
# +
## Most AF values are vey low
# -
sns.distplot(df.AF_ESP, label = "AF_ESP")
sns.distplot(df.AF_EXAC, label="AF_EXAC")
sns.distplot(df.AF_TGP, label= "AF_TGP")
plt.legend()
|
Kaggle_Github/Genetic Variant Classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
from newsapi import NewsApiClient
import nltk
# %matplotlib inline
from nltk.sentiment.vader import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
# # News Headlines Sentiment
#
# Use the news api to pull the latest news articles for bitcoin and ethereum and create a DataFrame of sentiment scores for each coin.
#
# Use descriptive statistics to answer the following questions:
# 1. Which coin had the highest mean positive score?
# 2. Which coin had the highest negative score?
# 3. Which coin had the highest positive score?
# Read your api key environment variable
api_key = os.getenv("news_api")
# Create a newsapi client
newsapi = NewsApiClient(api_key=api_key)
# +
# Fetch the Bitcoin news articles
bitcoin_headlines = newsapi.get_everything(
q="bitcoin",
language="en",
page_size=100,
sort_by="relevancy"
)
# Print total articles
print(f"Total articles about Bitcoin: {bitcoin_headlines['totalResults']}")
# Show sample article
bitcoin_headlines["articles"][20]
# +
# Fetch the Ethereum news articles
ethereum_headlines = newsapi.get_everything(
q="ethereum",
language="en",
page_size=100,
sort_by="relevancy"
)
# Print total articles
print(f"Total articles about Ethereum: {ethereum_headlines['totalResults']}")
# Show sample article
ethereum_headlines["articles"][20]
# +
# Create the Bitcoin sentiment scores DataFrame
bitcoin_sentiments = []
for article in bitcoin_headlines["articles"]:
try:
text = article["content"]
sentiment = analyzer.polarity_scores(text)
compound = sentiment["compound"]
pos = sentiment["pos"]
neu = sentiment["neu"]
neg = sentiment["neg"]
bitcoin_sentiments.append({
"text": text,
"compound": compound,
"positive": pos,
"negative": neg,
"neutral": neu
})
except AttributeError:
pass
# Create DataFrame
bitcoin_df = pd.DataFrame(bitcoin_sentiments)
# Reorder DataFrame columns
cols = ["compound", "positive", "negative", "neutral", "text"]
bitcoin_df = bitcoin_df[cols]
bitcoin_df.head()
# +
# Create the ethereum sentiment scores DataFrame
ethereum_sentiments = []
for article in ethereum_headlines["articles"]:
try:
text = article["content"]
sentiment = analyzer.polarity_scores(text)
compound = sentiment["compound"]
pos = sentiment["pos"]
neu = sentiment["neu"]
neg = sentiment["neg"]
ethereum_sentiments.append({
"text": text,
"compound": compound,
"positive": pos,
"negative": neg,
"neutral": neu
})
except AttributeError:
pass
# Create DataFrame
ethereum_df = pd.DataFrame(ethereum_sentiments)
# Reorder DataFrame columns
cols = ["compound", "positive", "negative", "neutral", "text"]
ethereum_df = ethereum_df[cols]
ethereum_df.head()
# -
# Describe the Bitcoin Sentiment
bitcoin_df.describe()
# Describe the Ethereum Sentiment
ethereum_df.describe()
# ### Questions:
#
# Q: Which coin had the highest mean positive score?
#
# A: Bitcoin but by a mere .00003 so it is not significant and both are by all standards the same.
#
# Q: Which coin had the highest compound score?
#
# A: Ethereum has a higher mean compound score.
#
# Q. Which coin had the highest positive score?
#
# A: Bitcoin has the highest postive score.
#
# Q: Which coin had the highest negative score?
#
# A: Bitcoin has the highest negative score.
# ---
# # Tokenizer
#
# In this section, you will use NLTK and Python to tokenize the text for each coin. Be sure to:
# 1. Lowercase each word
# 2. Remove Punctuation
# 3. Remove Stopwords
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer, PorterStemmer
from string import punctuation
import re
# +
# Expand the default stopwords list if necessary
# -
import nltk
nltk.download('wordnet')
# Complete the tokenizer function
lemmatizer = WordNetLemmatizer()
def tokenizer(text):
"""Tokenizes text."""
sw = set(stopwords.words('english'))
regex = re.compile("[^a-zA-Z ]")
re_clean = regex.sub('', str(text))
words = word_tokenize(re_clean)
lem = [lemmatizer.lemmatize(word) for word in words]
tokens = [word.lower() for word in lem if word.lower() not in sw]
return tokens
# Create a new tokens column for bitcoin
bitcoin_text = bitcoin_df['text'].tolist()
sentence_tokenized = [sent_tokenize(i) for i in bitcoin_text]
word_tokenized = []
for story in sentence_tokenized:
words = []
for sent in story:
words = tokenizer(story)
# append all words for each article to the word_tokenized list
word_tokenized.append(words)
bitcoin_df['bitcoin_tokens'] = word_tokenized
bitcoin_df
# Create a new tokens column for ethereum
ethereum_text = ethereum_df['text'].tolist()
sentence_tokenized = [sent_tokenize(i) for i in ethereum_text]
word_tokenized = []
for story in sentence_tokenized:
words = []
for sent in story:
words = tokenizer(story)
# append all words for each article to the word_tokenized list
word_tokenized.append(words)
ethereum_df['ethereum_tokens'] = word_tokenized
ethereum_df
# ---
# # NGrams and Frequency Analysis
#
# In this section you will look at the ngrams and word frequency for each coin.
#
# 1. Use NLTK to produce the n-grams for N = 2.
# 2. List the top 10 words for each coin.
from collections import Counter
from nltk import ngrams
# Generate the Bitcoin N-grams where N=2
def bigram_counter(corpus):
# Combine all articles in corpus into one large string
big_string = ' '.join(corpus)
processed = tokenizer(big_string)
bigrams = ngrams(processed, n=2)
count = dict(Counter(bigrams))
return pd.DataFrame(list(count.items()), columns=['bigram', 'count']).sort_values(by=['count'], ascending=False)
bitcoin_bigrams = bigram_counter(bitcoin_df['text'])
bitcoin_bigrams
# Generate the Ethereum N-grams where N=2
ethereum_bigrams = bigram_counter(ethereum_df['text'])
ethereum_bigrams
# Use the token_count function to generate the top 10 words from each coin
def token_count(tokens, N=10):
"""Returns the top N tokens from the frequency count"""
big_string = ' '.join(tokens)
tokens = tokenizer(big_string)
return Counter(tokens).most_common(N)
# Get the top 10 words for Bitcoin
bitcoin_10 = token_count(bitcoin_df['text'])
bitcoin_10
# Get the top 10 words for Ethereum
ethereum_10 = token_count(ethereum_df['text'])
ethereum_10
# # Word Clouds
#
# In this section, you will generate word clouds for each coin to summarize the news for each coin
from wordcloud import WordCloud
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = [20.0, 10.0]
bitcoin_df['bitcoin_tokens']
# Generate the Bitcoin word cloud
bt_entities = bitcoin_df['bitcoin_tokens'].astype(str)
wc = WordCloud().generate(' '.join(bt_entities))
plt.imshow(wc)
# Generate the Ethereum word cloud
et_entities = ethereum_df['ethereum_tokens'].astype(str)
wc = WordCloud().generate(' '.join(et_entities))
plt.imshow(wc)
# # Named Entity Recognition
#
# In this section, you will build a named entity recognition model for both coins and visualize the tags using SpaCy.
import spacy
from spacy import displacy
# +
# Optional - download a language model for SpaCy
# # !python -m spacy download en_core_web_sm
# -
# Load the spaCy model
nlp = spacy.load('en_core_web_sm')
# ## Bitcoin NER
# Concatenate all of the bitcoin text together
bitcoin_all_text = bitcoin_df['text'].str.cat(sep=', ')
bitcoin_all_text[:3000]
# Run the NER processor on all of the text
bt_doc = nlp(bitcoin_all_text)
# Add a title to the document
bt_doc.user_data["title"] = "Bitcoin NER"
# Render the visualization
displacy.render(bt_doc, style='ent')
# +
# List all Entities
bt_entities = [(ent.text, ent.label_) for ent in bt_doc.ents]
for x in bt_entities:
print(x[0], x[1])
# -
# ---
# ## Ethereum NER
# Concatenate all of the bitcoin text together
ethereum_all_text = ethereum_df['text'].str.cat(sep=', ')
ethereum_all_text[:3000]
# Run the NER processor on all of the text
et_doc = nlp(ethereum_all_text)
# Add a title to the document
et_doc.user_data["title"] = "Ethereum NER"
# Render the visualization
displacy.render(et_doc, style='ent')
# +
# List all Entities
et_entities = [(ent.text, ent.label_) for ent in et_doc.ents]
for x in et_entities:
print(x[0], x[1])
# -
|
code/crypto_sentiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Language Filtering
# import packages
import pandas as pd
from langdetect import detect_langs
import multiprocessing as mp
import tqdm
import re
# import data
data = pd.read_csv('clean_data_odd.csv')
data.head()
# filter out coments where there are no alphabetical characters
# will mess with language detection if left in!
data['has_char'] = data['body'].apply(str).str.contains('[a-zA-Z]+')
data.head()
# see how many returned
data['has_char'].value_counts()
# +
# filter out the flase results
data = data.loc[data['has_char'] == True]
#check to make sure that they were filtered out
data['has_char'].value_counts()
# -
# reindex since data has been filtered out
data = data.reset_index(drop = True)
# ## method 1
# - #### filter via subreddit using R
# - #### go to part 2_clean_reddit_comments.md
# ## > method 2
# - #### filter via language detection
# ```Python
# # create new ['language'] column
# data['language'] = data['body'].apply(detect_langs)
#
# # new column ['english'] returns 'True' if english has any probability of being the language
# data['english'] = data['language'].apply(str).str.contains('en')
#
# # see how many were classified as each
# data['english'].value_counts()
#
# # see what was classified as false
# data[data['english'] == False].head()
#
# # parse data to only return values where ['english'] is True
# data = data[(data['english'] == True)]
#
# #reset index as indexes of filtered data are deleted
# data = data.reset_index(drop = True)
#
# # drop data that we don't need anymore
# del data['language']
# del data['english']
# ```
# ## > method 3
# - #### method 2 but as a function
# ```Python
# # Arguments: (df = your dataframe), (series = name of column in df as string), (language_select = two letter string of language code that you want)
#
# def language_filter(df, series = str, language_select = str):
#
# # create copied df
# df_copy = df.copy()
#
# # create ['languague'] from output of detect_langs
# df_copy['language'] = df_copy[series].apply(detect_langs)
#
# # new column ['contains_your_language'] returns 'True' if ['language'] contains any probability of your language
# df_copy['contains_your_language'] = df_copy['language'].apply(str).str.contains(language_select)
#
# # parse data to only return values where ['contains_your_language'] is True
# df_copy = df_copy.loc[df_copy['contains_your_language'] == True]
#
# # remove ['language'] and ['contains_your_language'] as they are no longer needed
# del df_copy['language']
# del df_copy['contains_your_language']
#
# # reindex df
# df_copy = df_copy.reset_index(drop = True)
#
# # return your new filtered dataframe
# return df_copy
# ```
# ```Python
# # run function
# data = language_filter(df = data, series = 'body', language_select = 'en')
# data.head()
# ```
# ## > method 4
# - #### parallel processing
# The parallel processing package doesn't have a version of pandas `.apply`, it has a<br>
# version of `map` (which is a list's version of apply). This means we need to transform <br>
# our data into a list and then use the map function. below are some examples:
#
# ```Python
# # list comprehension = this is the function that is going to be parallel processed
# test_list = [i for i in map(detect_langs, data['body'])]
#
# # same thing as list comprehension above
# test_list = []
# for i in data['body']:
# results = detect_langs(i)
# test_list.append(results)
# ```
# number of parallel processes to create
# should be number of cpu cores that your computer has
num_processes = mp.cpu_count()
num_processes
# Below, `pool.map` works just the same as `map`. Chunksize is the amount of data<br>
# that each process will work on at a time. To get the progress bar, simply wrap<br>
# `tqdm.tqdm` around the object that you are iterating over, which in this case<br>
# is `data['body']`
# parallel process with progress bar
if __name__ == '__main__':
with mp.Pool(num_processes) as pool:
data['language'] = [i for i in pool.map(detect_langs, tqdm.tqdm(data['body']), chunksize = 10)]
# new column ['english'] returns 'True' if english has any probability of being the language
data['english'] = data['language'].apply(str).str.contains('en')
data.head()
# see how many were classified as each
data['english'].value_counts()
# parse data to only return values where ['english'] is True
data = data[(data['english'] == True)]
#reset index as indexes of filtered data are deleted
data = data.reset_index(drop = True)
# drop data that we don't need anymore
del data['language']
del data['english']
data.write_csv('filtered_data_odd.csv')
|
4_language_filtering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D3_OptimalControl/W3D3_Tutorial2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>[](https://kaggle.com/kernels/welcome?src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D3_OptimalControl/W3D3_Tutorial2.ipynb)
# -
# # Tutorial 2: Optimal Control for Continuous State
# **Week 3, Day 3: Optimal Control**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and <NAME>
#
# + [markdown] colab_type="text"
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# -
# ---
# # Tutorial Objectives
# In this tutorial, we will implement a continuous control task: you will design control inputs for a linear dynamical system to reach a target state. The state here is continuous-valued, i.e. takes on any real number from $-\infty$ to $\infty$.
#
# You have already learned about control for binary states in Tutorial 1, and you have learned about stochastic dynamics, latent states, and measurements yesterday. Now we introduce you to the new concepts of designing a controller with full observation of the state (linear qudratic regulator - LQR), and under partial observability of the state (linear quadratic gaussian - LQG).
#
# The running example we consider throughout the tutorial is a cat trying to catch a mouse in space, using its handy little jet pack to navigate.
# ---
# # Setup
# + cellView="both" pycharm={"is_executing": false}
# Imports
import numpy as np
import scipy
import matplotlib.pyplot as plt
from matplotlib import gridspec
from math import isclose
# + cellView="form"
#@title Figure Settings
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import ipywidgets as widgets
from ipywidgets import interact, fixed, HBox, Layout, VBox, interactive, Label
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form"
#@title Helper functions
# Helper function for plotting
def plot_vs_time(s, slabel, color, goal=None, ylabel=None):
plt.plot(s, color, label=slabel)
if goal is not None:
plt.plot(goal, 'm', label='goal $g$')
plt.xlabel("Time", fontsize=14)
plt.legend(loc="upper right")
if ylabel:
plt.ylabel(ylabel, fontsize=14)
class ExerciseError(AssertionError):
pass
def test_lds_class(lds_class):
from math import isclose
ldsys = lds_class(T=2, ini_state=2., noise_var=0.)
if not isclose(ldsys.dynamics(.9)[1], 1.8):
raise ExerciseError("'dynamics' method is not correctly implemented!")
if not isclose(ldsys.dynamics_openloop(.9, 2., np.zeros(ldsys.T)-1.)[1], -0.2):
raise ExerciseError("'dynamics_openloop' method is not correctly implemented!")
if not isclose(ldsys.dynamics_closedloop(.9, 2., np.zeros(ldsys.T)+.3)[0][1], 3.):
raise ExerciseError("s[t] in 'dynamics_closedloop' method is not correctly implemented!")
if not isclose(ldsys.dynamics_closedloop(.9, 2., np.zeros(ldsys.T)+.3)[1][0], .6):
raise ExerciseError("a[t] in 'dynamics_closedloop' method is not correctly implemented!")
ldsys.noise_var = 1.
if isclose(ldsys.dynamics(.9)[1], 1.8):
raise ExerciseError("Did you forget to add noise to your s[t+1] in 'dynamics'?")
if isclose(ldsys.dynamics_openloop(.9, 2., np.zeros(ldsys.T)-1.)[1], -0.2):
raise ExerciseError("Did you forget to add noise to your s[t+1] in 'dynamics_openloop'?")
if isclose(ldsys.dynamics_closedloop(.9, 2., np.zeros(ldsys.T)+.3)[0][1], 3.):
raise ExerciseError("Did you forget to add noise to your s[t+1] in 'dynamics_closedloop'?")
if not isclose(ldsys.dynamics_closedloop(.9, 2., np.zeros(ldsys.T)+.3)[1][0], .6):
raise ExerciseError("Your input a[t] should not be noisy in 'dynamics_closedloop'.")
print('Well Done!')
def test_lqr_class(lqr_class):
from math import isclose
lqreg = lqr_class(T=2, ini_state=2., noise_var=0.)
lqreg.goal = np.array([-2, -2])
s = np.array([1, 2])
a = np.array([3, 4])
if not isclose(lqreg.calculate_J_state(s), 25):
raise ExerciseError("'calculate_J_state' method is not correctly implemented!")
if not isclose(lqreg.calculate_J_control(a), 25):
raise ExerciseError("'calculate_J_control' method is not correctly implemented!")
print('Well Done!')
# -
# ---
# # Section 1: Exploring a Linear Dynamical System (LDS) with Open-Loop and Closed-Loop Control
# + cellView="form"
# @title Video 1: Flying Through Space
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="MLUTR8z16jI", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
#
# In this example, a cat is trying to catch a mouse in space. The location of the mouse is the goal state $g$, here a static goal. Later on, we will make the goal time varying, i.e. $g(t)$. The cat's location is the state of the system $s_t$. The state has its internal dynamics: think of the cat drifting slowly in space. These dynamics are such that the state at the next time step $s_{t+1}$ are a linear function of the current state $s_t$. There is some environmental noise (think: meteorites) affecting the state, here modeled as gaussian noise $w_t$.
#
# The control input or action $a_t$ is the action of the jet pack, which has an effect $Ba_t$ on the state at the next time step $s_{t+1}$. In this tutorial, we will be designing the action $a_t$ to reach the goal $g$, with known state dynamics.
#
# Thus, our linear discrete-time system evolves according to the following equation:
#
# \begin{eqnarray*}
# s_{t+1} &=& Ds_t + Ba_t + w_t \tag{1}\\
# s_{0} &=& s_{init}
# \end{eqnarray*}
#
# with
#
# $t$: time step, ranging from $1$ to $T$, where $T$ is the time horizon.
#
# $s_t$: state at time $t$
#
# $a_t$: action at time $t$ (also known as control input)
#
# $w_t$: gaussian noise at time $t$
#
# $D$ and $B$: parameters of the linear dynamical system.
#
# For simplicity, we will consider the 1D case, where the matrices reduce to scalars, and the states, control and noise are one-dimensional as well. Specifically, $D$ and $B$ are scalars.
#
# We will consider the goal $g$ to be the origin, i.e. $g=0$, for Exercises 1 and 2. Note that if the state dynamics are stable, the state reaches $0$ in any case. This is a slightly unrealistic situation for the purposes of simplicity, but we will see more realistic cases later on with $g \neq 0$ in Exercise 3.
#
# **Stability** \\
# The system is stable, i.e. the output remains finite for any finite initial condition $s_{init}$, if $|D|<1$.
#
# **Control** \\
# In *open-loop control*, $a_t$ is not a function of $s_t$. In *closed-loop linear control*, $a_t$ is a linear function of the state $s_t$. Specifically, $a_t$ is the control gain $L_t$ multiplied by $s_t$, i.e. $a_t=L_t s_t$. For now, you will explore these equations, and later on, you will *design* $L_t$ to reach the goal $g$.
# ### Exercise 1: Implement state evolution equations
#
#
# Implement the state evolution equations in the class methods as provided below, for the following cases: \\
# (a) no control: `def dynamics` \\
# (b) open-loop control: `def dynamics_openloop` \\
# (c) closed-loop control: `def dynamics_closedloop` \\
#
# *Tip: refer to Equation (1) above. The provided code uses the same notation*
# + cellView="both" pycharm={"is_executing": false}
class LDS:
def __init__(self, T: int, ini_state: float, noise_var: float):
self.T = T # time horizon
self.ini_state = ini_state
self.noise_var = noise_var
def dynamics(self, D: float):
s = np.zeros(self.T) # states initialization
s[0] = self.ini_state
noise = np.random.normal(0, self.noise_var, self.T)
for t in range(self.T - 1):
####################################################################
## Insert your code here to fill with the state dynamics equation
## without any control input
## complete the function and remove
raise NotImplementedError("Exercise: Please complete 'dynamics'")
####################################################################
# calculate the state of t+1
s[t + 1] = ...
return s
def dynamics_openloop(self, D: float, B: float, a: np.ndarray):
s = np.zeros(self.T) # states initialization
s[0] = self.ini_state
noise = np.random.normal(0, self.noise_var, self.T)
for t in range(self.T - 1):
####################################################################
## Insert your code here to fill with the state dynamics equation
## with open-loop control input a[t]
## complete the function and remove
raise NotImplementedError("Please complete 'dynamics_openloop'")
####################################################################
# calculate the state of t+1
s[t + 1] = ...
return s
def dynamics_closedloop(self, D: float, B: float, L: np.ndarray):
s = np.zeros(self.T) # states initialization
s[0] = self.ini_state
noise = np.random.normal(0, self.noise_var, self.T)
a = np.zeros(self.T - 1)
for t in range(self.T - 1):
####################################################################
## Insert your code here to fill with the state dynamics equation
## with closed-loop control input as a function of control gain L.
## complete the function and remove
raise NotImplementedError("Please complete 'dynamics_closedloop'")
####################################################################
# calculate the current action
a[t] = ...
# calculate the next state
s[t + 1] = ...
return s, a
# uncomment the line below to test your class
# test_lds_class(LDS)
# +
# to_remove solution
class LDS:
def __init__(self, T: int, ini_state: float, noise_var: float):
self.T = T # time horizon
self.ini_state = ini_state
self.noise_var = noise_var
def dynamics(self, D: float):
s = np.zeros(self.T) # states initialization
s[0] = self.ini_state
noise = np.random.normal(0, self.noise_var, self.T)
for t in range(self.T - 1):
# calculate the state of t+1
s[t + 1] = D * s[t] + noise[t]
return s
def dynamics_openloop(self, D: float, B: float, a: np.ndarray):
s = np.zeros(self.T) # states initialization
s[0] = self.ini_state
noise = np.random.normal(0, self.noise_var, self.T)
for t in range(self.T - 1):
# calculate the state of t+1
s[t + 1] = D * s[t] + B * a[t] + noise[t]
return s
def dynamics_closedloop(self, D: float, B: float, L: np.ndarray):
s = np.zeros(self.T) # states initialization
s[0] = self.ini_state
noise = np.random.normal(0, self.noise_var, self.T)
a = np.zeros(self.T - 1)
for t in range(self.T - 1):
# calculate the current action
a[t] = L[t] * s[t]
# calculate the next state
s[t + 1] = D * s[t] + B * a[t] + noise[t]
return s, a
test_lds_class(LDS)
# -
# ### Interactive Demo 1.1: Explore no control vs. open-loop control vs. closed-loop control
#
# Once your code above passes the tests, use the interactive demo below to visualize the effects of different kinds of control inputs.
#
# (a) For the no-control case, can you identify two distinct outcomes, depending on the value of D? Why?
#
# (b) The open-loop controller works well--or does it? Run the simulation multiple times and see if there are any problems, especially in challenging (high noise) conditions.
#
# (c) Does the closed-loop controller fare better with the noise? Vary the values of $L$ and find a range where it quickly reaches the goal.
#
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
#@markdown Play around (attentively) with **`a`** and **`L`** to see the effect on the open-loop controlled and closed-loop controlled state.
def simulate_lds(D=0.95, L=-0.3, a=-1., B=2., noise_var=0.1,
T=50, ini_state=2.):
# linear dynamical system
lds = LDS(T, ini_state, noise_var)
# No control
s_no_control=lds.dynamics(D)
# Open loop control
at = np.append(a, np.zeros(T - 1))
s_open_loop = lds.dynamics_openloop(D, B, at)
# Closed loop control
Lt = np.zeros(T) + L
s_closed_loop, a_closed_loop = lds.dynamics_closedloop(D, B, Lt)
plt.figure(figsize=(10, 6))
plt.plot(s_no_control, 'b', label='No control')
plt.plot(s_open_loop, 'g', label='Open Loop with a = {}'.format(a))
plt.plot(s_closed_loop, 'r', label='Closed Loop with L = {}'.format(L))
plt.plot(np.zeros(T), 'm', label='goal')
plt.title('LDS State Evolution')
plt.ylabel('State', fontsize=14)
plt.xlabel('Time', fontsize=14)
plt.legend(loc="upper right")
plt.show()
widget=interactive(simulate_lds, {'manual': True},
D=(.85, 1.05, .1),
L=(-0.6, 0., .15),
a=(-2., 1., 1.),
B=(1., 3., 1.),
noise_var=(0., 0.2, .1),
T=fixed(50),
ini_state=(2., 10., 4.))
widget.children[-2].description='Run Simulation'
widget.children[-2].style.button_color='lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
#to_remove explanation
"""
In Exercise 1.2, you should have noticed the following:
* No control (blue): the dynamics parameter D controls how fast the dynamics decay
towards 0. For -1<D<1, the system is stable and therefore approaches zero quickly.
However, D>1 produces an unstable system, causing , you should have
noticed that the 'no control' state (blue curve) rapidly explodes
(i.e., heads off to infinity)
* Open-loop control: While the open-loop state (green curve) often reachs the goal
quickly, it may not stay there. Under high noise conditions, it tends to
drift away from the goal, though you may not see this in every simulation.
* Closed-loop control: The closed-loop state (red curve) reaches the goal and
stays there even in the presence of noise. It converges especially quickly for
Ls around 0.45
Remember that in closed-loop control,
we have a[t]=L[t] * s[t] $. Note that with a constant control gain $L[t]=L,
the state evolution equations can be rearranged to show that the stability of
the closed-loop system now depends on the value of D+BL. (See Equation 2, below).
If $|D+BL|<1$, our closed-loop system will be stable. More generally, you can
view the role of a closed-loop control input as changing the system *dynamics*
in an optimal way to reach the goal.
""";
# ### Interactive Demo 1.2: Exploring the closed-loop setting further
# Execute the cell below to visualize the MSE between the state and goal, as a function of control gain $L$. You should see a U-shaped curve, with a clear minimum MSE. The control gain at which the minimum MSE is reached, is the 'optimal' constant control gain for minimizing MSE, here called the numerical optimum.
#
# A green dashed line is shown $L = -\frac{D}{B}$ with $D=0.95$ and $B=2$. Consider how Why is this the theoretical optimal control gain for minimizing MSE of the state $s$ to the goal $g=0$? Examine how the states evolve with a constant gain $L$
# $$
# \begin{eqnarray*}
# s_{t+1} &=& Ds_t + Ba_t + w_t \\
# &=& Ds_t + B(Ls_t) + w_t \\
# &=& (D+BL)s_t + w_t \tag{2}
# \end{eqnarray*}
# $$
#
# Now, let's visualize the evolution of the system as we change the control gain. We will start with the optimal gain (the control gain that gets us the minimum MSE), and then explore over- and under- ambitious values.
# + cellView="form"
#@markdown Execute this cell to visualize MSE between state and goal, as a function of control gain
def calculate_plot_mse():
D, B, noise_var, T, ini_state = 0.95, 2., 0.1, 50, 2.
control_gain_array = np.linspace(0.1, 0.9, T)
mse_array = np.zeros(control_gain_array.shape)
for i in range(len(control_gain_array)):
lds = LDS(T, ini_state, noise_var)
L = - np.ones(T) * control_gain_array[i]
s, a = lds.dynamics_closedloop(D, B, L)
mse_array[i] = np.sum(s**2)
plt.figure()
plt.plot(-control_gain_array, mse_array, 'b')
plt.axvline(x=-D/B, color='g', linestyle='--')
plt.xlabel("control gain (L)", fontsize=14)
plt.ylabel("MSE between state and goal" , fontsize=14)
plt.title("MSE vs control gain", fontsize=20)
plt.show()
calculate_plot_mse()
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
#@markdown Explore different values of control gain **`L`** (close to optimal, over- and under- ambitious) \\
def simulate_L(L:float=-0.45):
D, B, noise_var, T, ini_state = 0.95, 2., 0.1, 50, 2.
lds = LDS(T, ini_state, noise_var)
# Closed loop control with the numerical optimal control gain
Lt = np.ones(T) * L
s_closed_loop_choice, _ = lds.dynamics_closedloop(D, B, Lt)
# Closed loop control with the theoretical optimal control gain
L_theory = - D / B * np.ones(T)
s_closed_loop_theoretical, _ = lds.dynamics_closedloop(D, B, L_theory)
# Plotting closed loop state evolution with both theoretical and numerical optimal control gains
plt.figure(figsize=(10, 6))
plot_vs_time(s_closed_loop_theoretical,
'Closed Loop (Theoretical optimal control gain)','b')
plot_vs_time(s_closed_loop_choice,
'Closed Loop (your choice of L = {})'.format(L), 'g',
goal=np.zeros(T), ylabel="State")
plt.title('Closed Loop State Evolution')
plt.show()
widget=interactive(simulate_L, {'manual': True}, L=(-1.05, 0.051, .1))
widget.children[-2].description='Run Simulation'
widget.children[-2].style.button_color='lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
#to_remove explanation
"""
In Demo 1.2, you should have seen that the optimal control gain (L = -0.45) takes a
short amount of time to get to the goal, and then stays there. We can try to get
to the goal in an even shorter time using an 'over-ambitious' control gain (L < -0.45), but
this may actually overshoot the goal and may cause oscillations in the system,
thus increasing the MSE. On the other hand, an 'under-ambitious' control gain
takes a longer time to get to the goal and thus increases the MSE.
Finally, at L>0, the system runs away to infinity.
Why is L=-D/B optimal for reaching our goal? Recall that our next state is
(D+B*L)*s[t] + noise. Plugging that L=-D/B causes that leading term to become zero,
which is our goal. Since the noise has zero mean, it's not possible to do any better!
""";
# ---
# # Section 2: Designing an optimal control input using a linear quadratic regulator (LQR)
# + cellView="form"
# @title Video 2: Linear quadratic regulator (LQR)
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="NZSwDy7wtIs", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Section 2.1 Constraints on the system
# Now we will start imposing additional constraints on our system. For example.
# if you explored different values for $s_{init}$ above, you would have seen very large values for $a_t$ in order to get to the mouse in a short amount of time. However, perhaps the design of our jetpack makes it dangerous to use large amounts of fuel in a single timestep. We certainly do not want to explode, so we would like to keep the actions $a_t$ as small as possible while still mantaining good control.
#
# Moreover, in Exercise 1, we had restricted ourselves to a static control gain $L_t \equiv L$. How would we vary it if we could?
#
# This leads us to a more principled way of designing the optimal control input.
# ### Setting up a cost function
#
# In a finite-horizon LQR problem, the cost function is defined as:
#
# \begin{eqnarray}
# J({\bf s},{\bf a}) &=& J_{state}({\bf s}) + \rho J_{control}({\bf a}) \\
# &=& \sum_{t = 0}^{T} (s_{t}-g)^2 + \rho \sum_{t=0}^{T-1}a_{t}^2 \tag{3}
# \end{eqnarray}
#
# where $\rho$ is the weight on the control effort cost, as compared to the cost of not being at the goal. Here, ${\bf a} = \{a_t\}_{t=0}^{T-1}$, ${\bf s} = \{s_t\}_{t=0}^{T}$. This is a quadratic cost function. In Exercise $2$, we will only explore $g=0$, in which case $J_{state}({\bf s})$ can also be expressed as $\sum_{t = 0}^{T} s_{t}^2$. In Exercise $3$, we will explore a non-zero time-varying goal.
#
# The goal of the LQR problem is to find control ${\bf a}$ such that $J({\bf s},{\bf a})$ is minimized. The goal is then to find the control gain at each time point, i.e.,
#
# $$ \text{argmin} _{\{L_t\}_{t=0}^{T-1}} J({\bf s},{\bf a}) \tag{4} $$
#
# where $a_t = L_t s_t$.
#
# ## Section 2.2 Solving LQR
# The solution to Equation (4), i.e. LQR for a finite time horizon, can be obtained via Dynamic Programming. For details, check out [this lecture by <NAME>](https://stanford.edu/class/ee363/lectures/dlqr.pdf).
#
# For an infinite time horizon, one can obtain a closed-form solution using Riccati equations, and the solution for the control gain becomes time-invariant, i.e. $L_t \equiv L$. We will use this in Exercise 4. For details, check out [this other lecture by <NAME>](https://stanford.edu/class/ee363/lectures/dlqr-ss.pdf).
#
# Additional reference for entire section: \\
# [<NAME>. "Dynamic programming and optimal control". Vol. 1. No. 2. Belmont, MA: Athena scientific, 1995.](http://www.athenasc.com/dpbook.html)
#
# ### Exercise 2.1: Implement the cost function
# The cost function $J_{control}({\bf s}, {\bf a})$ can be divided into two parts: $J_{state}({\bf s})$ and $J_{control}({\bf a})$.
#
# Code up these two parts in the class methods `def calculate_J_state` and `def calculate_J_control` in the following helper class for LQR.
#
# +
class LQR(LDS):
def __init__(self, T, ini_state, noise_var):
super().__init__(T, ini_state, noise_var)
self.goal = np.zeros(T) # The class LQR only supports g=0
def control_gain_LQR(self, D, B, rho):
P = np.zeros(self.T) # Dynamic programming variable
L = np.zeros(self.T - 1) # control gain
P[-1] = 1
for t in range(self.T - 1):
P_t_1 = P[self.T - t - 1]
P[self.T - t-2] = (1 + P_t_1 * D**2 - D * P_t_1 * B / (
rho + P_t_1 * B) * B**2 * P_t_1 * D)
L[self.T - t-2] = - (1 / (rho + P_t_1 * B**2) * B * P_t_1 * D)
return L
def calculate_J_state(self, s:np.ndarray):
########################################################################
## Insert your code here to calculate J_state(s) (see Eq. 3)
## complete the function and remove
raise NotImplementedError("Please complete 'calculate_J_state'")
########################################################################
# calculate the state
J_state = ...
return J_state
def calculate_J_control(self, a:np.ndarray):
########################################################################
## Insert your code here to calculate J_control(a) (see Eq. 3).
## complete the function and remove
raise NotImplementedError("Please complete 'calculate_J_control'")
########################################################################
# calculate the control
J_control = ...
return J_control
# uncomment the line below to test your class
# test_lqr_class(LQR)
# +
# to_remove solution
class LQR(LDS):
def __init__(self, T, ini_state, noise_var):
super().__init__(T, ini_state, noise_var)
self.goal = np.zeros(T) # The class LQR only supports g=0
def control_gain_LQR(self, D, B, rho):
P = np.zeros(self.T) # Dynamic programming variable
L = np.zeros(self.T - 1) # control gain
P[-1] = 1
for t in range(self.T - 1):
P_t_1 = P[self.T - t - 1]
P[self.T - t-2] = (1 + P_t_1 * D**2 - D * P_t_1 * B / (
rho + P_t_1 * B) * B**2 * P_t_1 * D)
L[self.T - t-2] = - (1 / (rho + P_t_1 * B**2) * B * P_t_1 * D)
return L
def calculate_J_state(self, s:np.ndarray):
# calculate the state
J_state = np.sum((s - self.goal)**2)
return J_state
def calculate_J_control(self, a:np.ndarray):
# calculate the control
J_control = np.sum(a**2)
return J_control
test_lqr_class(LQR)
# -
# ### Interactive Demo 2: LQR to the origin
#
# In this exercise, we will use your new LQR controller to track a static goal at $g=0$. Here, we will explore how varying $\rho$ affects its actions by\\
#
# 1. Using Equation 3, find a value for $\rho$ that will get you the same cost and control gain as Exercise 1.
# 2. Pick a larger value for $\rho$ and see the effect on the action.
# 3. Try increasing the rho to 2. What do you notice? \\
# 4. For different values of $\rho$, how does the control gain vary?
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
def simulate_rho(rho=1.):
D, B, T, ini_state, noise_var = 0.9, 2., 50, 2., .1 # state parameter
lqr = LQR(T, ini_state, noise_var)
L = lqr.control_gain_LQR(D, B, rho)
s_lqr, a_lqr = lqr.dynamics_closedloop(D, B, L)
plt.figure(figsize=(14, 4))
plt.suptitle('LQR Control for rho = {}'.format(rho), y=1.05)
plt.subplot(1, 3, 1)
plot_vs_time(s_lqr,'State evolution','b',goal=np.zeros(T))
plt.ylabel('State $s_t$')
plt.subplot(1, 3, 2)
plot_vs_time(a_lqr,'LQR Action','b')
plt.ylabel('Action $a_t$')
plt.subplot(1, 3, 3)
plot_vs_time(L,'Control Gain','b')
plt.ylabel('Control Gain $L_t$')
plt.tight_layout()
plt.show()
widget=interactive(simulate_rho, {'manual': True}, rho=(0., 2., 0.5))
widget.children[-2].description = 'Run Simulation'
widget.children[-2].style.button_color = 'lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
#to_remove explanation
"""
* rho=0 will get you the same cost and control gain as in Exercise 1 by
zeroing out the cost term.
* A small value for rho will have a similar solution as in (a), but with
potentially large values for |a[t]|.
* A large value for rho, like 2 will lead to small values for |a[t]|.
* The control gain becomes more time-varying (as opposed to fairly static)
for large rho. For some parameter values, L[t] oscillates during the entire
trajectory in order to keep $|a_t|$ low. Try D = 0.9, B = 2 and rho = 2.
""";
# ## Section 2.3: The tradeoff between state cost and control cost
#
# In Exercise 2.1, you implemented code to calculate for $J_{state}$ and $J_{control}$ in the class methods for the class LQR.
#
# We will now plot them against each other for varying values of $\rho$ to explore the tradeoff between state cost and control cost.
# + cellView="form"
#@markdown Execute this cell to visualize the tradeoff between state and control cost
def calculate_plot_costs():
D, B, noise_var, T, ini_state = 0.9, 2., 0.1, 50, 2.
rho_array = np.linspace(0.2, 40, 100)
J_state = np.zeros(rho_array.shape)
J_control = np.zeros(rho_array.shape)
for i in np.arange(len(rho_array)):
lqr = LQR(T, ini_state, noise_var)
L = lqr.control_gain_LQR(D, B, rho_array[i])
s_lqr, a_lqr = lqr.dynamics_closedloop(D, B, L)
J_state[i] = lqr.calculate_J_state(s_lqr)
J_control[i] = lqr.calculate_J_control(a_lqr)
fig = plt.figure(figsize=(6, 6))
plt.plot(J_state, J_control, '.b')
plt.xlabel("$J_{state} = \sum_{t = 0}^{T} (s_{t}-g)^2$", fontsize=14)
plt.ylabel("$J_{control} = \sum_{t=0}^{T-1}a_{t}^2$" , fontsize=14)
plt.title("Error vs control effort", fontsize=20)
plt.show()
calculate_plot_costs()
# -
# You should notice the bottom half of a 'C' shaped curve, forming the tradeoff between the state cost and the control cost under optimal linear control.
# For a desired value of the state cost, we cannot reach a lower control cost than the curve in the above plot. Similarly, for a desired value of the control cost, we must accept that amount of state cost. For example, if you know that you have a limited amount of fuel, which determines your maximum control cost to be $J_{control}^{max}$.
#
# You will be able to show that you will not be able to track your state with a higher accuracy than the corresponding $J_{state}$ as given by the graph above. This is thus an important curve when designing a system and exploring its control.
#
# ---
# # Section 3: LQR for tracking a time-varying goal
# + cellView="form"
# @title Video 3: Tracking a moving goal
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="HOoqM7kBWSY", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# In a more realistic situation, the mouse would move around constantly. Suppose you were able to predict the movement of the mouse as it bounces from one place to another. This becomes your goal trajectory $g_t$.
#
# When the target state, denoted as $g_t$, is not $0$, the cost function becomes
# $$ J({\bf a}) = \sum_{t = 0}^{T} (s_{t}- g_t) ^2 + \rho \sum_{t=0}^{T-1}(a_{t}-\bar a_t)^2$$
# Here, $\bar a_t$ is the desired action based on the goal trajectory. In other words, the controller considers the goal for the next time step, and designs a preliminary control action that gets the state at the next time step to the desired goal. Specifically, without taking into account noise $w_t$, we would like to design $\bar a_t$ such that $s_{t+1}=g_{t+1}$. Thus, from Equation $(1)$,
#
# \begin{eqnarray*}
# g_{t+1} &=& Ds_t + B \bar a_t\\
# \bar a_{t} &=& \frac{- Ds_t + g_{t+1}}{B}\\
# \end{eqnarray*}
#
# The final control action $a_t$ is produced by adding this desired action $\bar a_t$ with the term with the control gain $L_t(s_t - g_t)$.
# + cellView="form"
#@markdown Execute this cell to include class
#@markdown for LQR control to desired time-varying goal
class LQR_tracking(LQR):
def __init__(self, T, ini_state, noise_var, goal):
super().__init__(T, ini_state, noise_var)
self.goal = goal
def dynamics_tracking(self, D, B, L):
s = np.zeros(self.T) # states intialization
s[0] = self.ini_state
noise = np.random.normal(0, self.noise_var, self.T)
a = np.zeros(self.T) # control intialization
a_bar = np.zeros(self.T)
for t in range(self.T - 1):
a_bar[t] = ( - D * s[t] + self.goal[t + 1]) / B
a[t] = L[t] * (s[t] - self.goal[t]) + a_bar[t]
s[t + 1] = D * s[t] + B * a[t] + noise[t]
return s, a, a_bar
def calculate_J_state(self,s):
J_state = np.sum((s-self.g)**2)
return J_state
def calculate_J_control(self, a, a_bar):
J_control = np.sum((a-a_bar)**2)
return J_control
# -
# ### Interactive Demo 3: LQR control to desired time-varying goal
# Use the demo below to explore how LQR tracks a time-varying goal.
# Starting with the sinusoidal goal function `sin`, investigate how the system reacts with different values of $\rho$ and process noise variance. Next, explore other time-varying goal, such as a step function and ramp.
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
def simulate_tracking(rho=20., noise_var=0.1, goal_func='sin'):
D, B, T, ini_state = 0.9, 2., 100, 0.
if goal_func == 'sin':
goal = np.sin(np.arange(T) * 2 * np.pi * 5 / T)
elif goal_func == 'step':
goal = np.zeros(T)
goal[int(T / 3):] = 1.
elif goal_func == 'ramp':
goal = np.zeros(T)
goal[int(T / 3):] = np.arange(T - int(T / 3)) / (T - int(T / 3))
lqr_time = LQR_tracking(T, ini_state, noise_var, goal)
L = lqr_time.control_gain_LQR(D, B, rho)
s_lqr_time, a_lqr_time, a_bar_lqr_time = lqr_time.dynamics_tracking(D, B, L)
plt.figure(figsize=(13, 5))
plt.suptitle('LQR Control for time-varying goal', y=1.05)
plt.subplot(1, 2, 1)
plot_vs_time(s_lqr_time,'State evolution $s_t$','b',goal, ylabel="State")
plt.subplot(1, 2, 2)
plot_vs_time(a_lqr_time, 'Action $a_t$', 'b', ylabel="Action")
plt.show()
widget=interactive(simulate_tracking, {'manual': True},
rho=(0., 40., 10.),
noise_var=(0., 1., .2),
goal_func=['sin', 'step', 'ramp']
)
widget.children[-2].description = 'Run Simulation'
widget.children[-2].style.button_color = 'lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# +
#to_remove explanation
"""
In Exercise 3, you should have noticed that:
* The system follows time varying goals rather well, with little change to the
cost function and the control equations.
* Setting rho=0 leads to noise in the first part of the time series.
Here, we see that the control cost in fact acts as a regularizer.
* Larger values of the process noise variance lead to a higher MSE between the
state and the desired goal.
""";
# -
# ---
# # Section 4: Control of an partially observed state using a Linear Quadratic Gaussian (LQG) controller
#
# ## Section 4.1 Introducing the LQG Controller
# + cellView="form"
# @title Video 4: Linear Quadratic Gaussian (LQG) Control
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="c_D7iDLT_bw", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# In practice, the controller does not have full access to the state. For example, your jet pack in space may be controlled by Mission Control back on earth! In this case, noisy measurements $m_t$ of the state $s_t$ are taken via radar, and the controller needs to (1) estimate the true state, and (2) design an action based on this estimate.
#
# Fortunately, the separation principle tells us that it is optimal to do (1) and (2) separately. This makes our problem much easier, since we already know how to do each step.
#
# 1) *State Estimation*
# Can we recover the state from the measurement?
# From yesterday's lecture, it is known that the states $\hat{s}_t$ can be estimated from the measurements $m_t$ using the __Kalman filter__.
#
# 2) *Design Action*
# In Sections 2 and 3 above, we just learnt about the LQR controller which designs an action based on the state. The separation principle tells us that it is sufficient to replace the use of the state in LQR with the *estimated* state, i.e.
#
# $$a_t = L_t \hat s_t$$
#
# The state dynamics will then be:
# $$s_{t+1} = D s_t + B a_t + w_t$$
# where $w_t$ is the process noise (proc_noise), and the observation / measurement is:
# $$ y_t = C s_t + v_t$$
# with $v_t$ being the measurement noise (meas_noise).
#
# The combination of (1) state estimation and (2) action design using LQR is known as a **linear quadratic gaussian (LQG)**. Yesterday, you completed the code for Kalman filter. Based on that, you will code up the LQG controller. For these exercises, we will resturn to using the goal $g=0$, as in Section 2.
#
# ### Interactive Demo 4.1: The Kalman filter in conjunction with a linear closed-loop controller (LQG Control)
# In the `MyKalmanFilter` class, the method `filter_control` implements filtering in closed-loop feedback. It is a combination of generating samples (states $s_t$) and filtering (generating state estimates $\hat s_t$), as you have seen in yesterday's tutorial. The only difference from yesterday is that today's Kalman filter is in closed loop with the controller. Thus, each $s_{t+1}$ gets an input $a_t$, which itself depends on the state estimate of the last time step $\hat s_t$.
#
# Below you find the code snipets for the Kalman filter in closed loop (`MyKalmanFilter`) class that provide you an insight in action update (`control_policy_LQG`) and state estimation (`state_dynamics_LQG`). Please feel free to inspect the helper functions and classes for the details.
#
# You should have seen the next cell containing `MyKalmanFilter` class yesterday, with the exception of the controller acting on the state estimate in feedback, using the methods/equations you will find below.
# + cellView="form"
#@markdown Execute this cell to include MyKalmanFilter class
class MyKalmanFilter():
def __init__(self, n_dim_state, n_dim_obs, transition_matrices, transition_covariance, observation_matrices,
observation_covariance, initial_state_mean, initial_state_covariance, control_matrices):
"""
@param n_dim_state: dimension of the latent variables
@param n_dim_obs: dimension of the observed variables
@param transition_matrices: D
@param transition_covariance: process noise
@param observation_matrices: C
@param observation_covariance: measurement noise
@param initial_state_mean: initial state estimate
@param initial_state_covariance: initial estimate on state variance
@param control_matrices: B
"""
self.n_dim_state = n_dim_state
self.n_dim_obs = n_dim_obs
self.transition_matrices = transition_matrices
self.transition_covariance = transition_covariance
self.observation_matrices = observation_matrices
self.observation_covariance = observation_covariance
self.initial_state_mean = initial_state_mean
self.initial_state_covariance = initial_state_covariance
self.control_matrices = control_matrices
def filter_control(self, n_timesteps, control_gain, use_myfilter=True):
"""
Method that performs Kalman filtering with a controller in feedback
@param n_timesteps: length of the data sample
@param control_gain: a numpy array whose dimension is [n_timesteps, self.n_dim_state]
@output: filtered_state_means: a numpy array whose dimension is [n_timesteps, self.n_dim_state]
@output: filtered_state_covariances: a numpy array whose dimension is [n_timesteps, self.n_dim_state, self.n_dim_state]
@output: latent_state: a numpy array whose dimension is [n_timesteps, self.n_dim_state]
@output: observed_state: a numpy array whose dimension is [n_timesteps, self.n_dim_obs]
@output: control: a numpy array whose dimension is [n_timesteps, self.n_dim_state]
"""
# validate inputs
# assert observed_dim == self.n_dim_obs
n_example = n_timesteps
observed_dim = self.n_dim_obs
latent_state = []
observed_state = []
control = []
current_latent_state = self.initial_state_mean #initial_state
control.append(self.initial_state_mean)
latent_state.append(current_latent_state)
observed_state.append(np.dot(self.observation_matrices, current_latent_state) +
np.random.multivariate_normal(np.zeros(self.n_dim_obs), self.observation_covariance))
# create holders for outputs
filtered_state_means = np.zeros([n_example, self.n_dim_state])
filtered_state_covariances = np.zeros([n_example, self.n_dim_state, self.n_dim_state])
if use_myfilter:
# the first state mean and state covar is the initial expectation
filtered_state_means[0] = self.initial_state_mean
filtered_state_covariances[0] = self.initial_state_covariance
# initialize internal variables
current_state_mean = self.initial_state_mean.copy()
current_state_covar = self.initial_state_covariance.copy()
self.p_n_list = np.zeros((n_example, self.n_dim_obs, self.n_dim_obs))
for i in range(1, n_example):
## Use the code in Exercise 4.1 to get the current action
current_action = control_policy_LQG(self,current_state_mean,control_gain[i])
control.append(current_action)
## Use the code in Exercise 4.1 to update the state
current_latent_state = state_dynamics_LQG(self,current_latent_state, current_action)
latent_state.append(current_latent_state)
# use observation_matrices and observation_covariance to calculate next observed state
observed_state.append(np.dot(self.observation_matrices, current_latent_state
) + np.random.multivariate_normal(np.zeros(self.n_dim_obs), self.observation_covariance))
current_observed_data = observed_state[-1]
# run a single step forward filter
# prediction step
predicted_state_mean = np.dot(self.transition_matrices, current_state_mean
) + np.dot(self.control_matrices, current_action)
predicted_state_cov = np.matmul(np.matmul(self.transition_matrices, current_state_covar),
np.transpose(self.transition_matrices)) + self.transition_covariance
# observation step
innovation = current_observed_data - np.dot(self.observation_matrices, predicted_state_mean)
innovation_covariance = np.matmul(np.matmul(self.observation_matrices, predicted_state_cov),
np.transpose(self.observation_matrices)) + self.observation_covariance
# update step
kalman_gain = np.matmul(np.matmul(predicted_state_cov, np.transpose(self.observation_matrices)),
np.linalg.inv(innovation_covariance))
current_state_mean = predicted_state_mean + np.dot(kalman_gain, innovation)
current_state_covar = np.matmul((np.eye(current_state_covar.shape[0]) -
np.matmul(kalman_gain, self.observation_matrices)),
predicted_state_cov)
# populate holders
filtered_state_means[i, :] = current_state_mean
filtered_state_covariances[i, :, :] = current_state_covar
self.p_n_list[i, :, :] = predicted_state_cov
# self.p_n_list[i-1, :, :] = predicted_state_cov
# new
# self.p_n_list[-1, :, :] = np.matmul(np.matmul(self.transition_matrices, filtered_state_covariances[-1,:,:]),
# np.linalg.inv(self.transition_matrices)) + self.transition_covariance
# else:
# #################################################################################
# # below: this is an alternative if you do not have an implementation of filtering
# kf = KalmanFilter(n_dim_state=self.n_dim_state, n_dim_obs=self.n_dim_obs)
# need_params = ['transition_matrices', 'observation_matrices', 'transition_covariance',
# 'observation_covariance', 'initial_state_mean', 'initial_state_covariance']
# for param in need_params:
# setattr(kf, param, getattr(self, param))
# filtered_state_means, filtered_state_covariances = kf.filter(X)
# #################################################################################
filtered_state_means = np.squeeze(np.array(filtered_state_means))
filtered_state_covariances = np.squeeze(np.array(filtered_state_covariances))
latent_state = np.squeeze(np.array(latent_state))
observed_state = np.squeeze(np.array(observed_state))
control = np.squeeze(np.array(control))
return filtered_state_means, filtered_state_covariances, latent_state, observed_state, control
def plot_state_vs_time(self, n_timesteps, control_gain, title, use_myfilter=True, goal=None):
filtered_state_means_impl, filtered_state_covariances_impl, latent, measurement, control = self.filter_control(
n_timesteps, control_gain)
fig = plt.figure(figsize=(12, 4))
plt.suptitle(title, y=1.05)
gs = gridspec.GridSpec(1, 2, width_ratios=[1, 2])
ax0 = plt.subplot(gs[0])
ax0.plot(latent,filtered_state_means_impl, 'b.')
ax0.set_xlabel('Latent State')
ax0.set_ylabel('Estimated State')
ax0.set_aspect('equal')
ax1 = plt.subplot(gs[1])
ax1.plot(latent, 'b', label = 'Latent State')
ax1.plot(filtered_state_means_impl, 'r', label = 'Estimated State')
if goal is not None:
ax1.plot(goal, 'm', label = 'goal')
ax1.set_xlabel('Time')
ax1.set_ylabel('State')
ax1.legend(loc="upper right")
plt.tight_layout()
plt.show()
# +
# inspect the 'control_policy_LQG' and 'state_dynamics_LQG' methods:
def control_policy_LQG(self, mean_estimated_state, control_gain):
current_action = control_gain * mean_estimated_state
return current_action
def state_dynamics_LQG(self, current_latent_state, current_action):
current_latent_state = np.dot(self.transition_matrices, current_latent_state)\
+ np.dot(self.control_matrices, current_action)\
+ np.random.multivariate_normal(np.zeros(self.n_dim_state),
self.transition_covariance)
return current_latent_state
# -
# Take a look at the helper code for the `MyKalmanFilter` class above. In the following exercises, we will use the same notation that we have been using in this tutorial; adapter code has been provided to convert it into the representation `MyKalmanFilter expects`.
#
# Use interactive demo below to refresh your memory of how a Kalman filter estimates state. `C` scales the observation matrix.
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
def simulate_kf_no_control(D=0.9, B=2., C=1., L=0., T=50, ini_state=5,
proc_noise = 0.1, meas_noise = 0.2):
control_gain = np.ones(T) * L
# Format the above variables into a format acccepted by the Kalman Filter
n_dim_state = 1
n_dim_obs = 1
n_timesteps = T
transition_matrices = np.eye(n_dim_state) * D
transition_covariance = np.eye(n_dim_obs) * proc_noise # process noise
observation_matrices = np.eye(n_dim_state) * C
observation_covariance = np.eye(n_dim_obs) * meas_noise
initial_state_mean = np.ones(n_dim_state) * ini_state
initial_state_covariance = np.eye(n_dim_state) * .01
control_matrices = np.eye(n_dim_state) * B
my_kf = MyKalmanFilter(n_dim_state, n_dim_obs, transition_matrices,
transition_covariance, observation_matrices,
observation_covariance, initial_state_mean,
initial_state_covariance, control_matrices)
my_kf.plot_state_vs_time(n_timesteps, control_gain,
'State estimation with KF (no control input)')
widget=interactive(simulate_kf_no_control, {'manual': True},
D=fixed(.95),
B=fixed(2.),
C=(0., 3., 1.),
proc_noise=(0., 1., .1),
meas_noise=(0.1, 1., .1),
T=fixed(50),
L=fixed(0),
ini_state=fixed(5.))
widget.children[-2].description = 'Run Simulation'
widget.children[-2].style.button_color = 'lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
#to_remove explanation
"""
You should have seen that the Kalman filter generally estimates the latent state
accurately, even with fairly high noise levels, except when C=0.
"""
# ### Interactive Demo 4.2: LQG controller output with varying control gains
#
# Now let's implement the Kalman filter with closed-loop feedback with the controller. We will first use an arbitary control gain and a fixed value for measurement noise. We will then use the control gain from the LQR for optimal performance, with varying values for $\rho$.
#
# (a) Visualize the system dynamics $s_t$ in closed-loop control with an arbitrary constant control gain. Vary this control gain.
#
# (b) Vary $\rho$ to visualize the output of the optimal LQG controller. Here, we will use an optimal *constant* control gain, which is optimal in the case of an infinite time horizon (get to the goal and stay there forever).
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
def simulate_kf_with_control(D=0.9, B=2., C=1., L=-0.1, T=50, ini_state=5,
proc_noise = 0.1, meas_noise = 0.2):
control_gain = np.ones(T)*L
# Format the above variables into a format acccepted by the Kalman Filter
n_dim_state = 1
n_dim_obs = 1
n_timesteps = T
transition_matrices = np.eye(n_dim_state) * D
transition_covariance = np.eye(n_dim_obs) * proc_noise # process noise
observation_matrices = np.eye(n_dim_state) * C
observation_covariance = np.eye(n_dim_obs) * meas_noise
initial_state_mean = np.ones(n_dim_state) * ini_state
initial_state_covariance = np.eye(n_dim_state) * .01
control_matrices = np.eye(n_dim_state) * B
my_kf = MyKalmanFilter(n_dim_state, n_dim_obs, transition_matrices,
transition_covariance, observation_matrices,
observation_covariance, initial_state_mean,
initial_state_covariance, control_matrices)
my_kf.plot_state_vs_time(n_timesteps, control_gain, goal = np.zeros(T),
title='State estimation with KF (controller gain = {})'.format(L))
widget=interactive(simulate_kf_with_control, {'manual': True},
D=fixed(.9),
B=fixed(2.),
C=(0., 3., 1.),
proc_noise=(0., 1., .1),
meas_noise=(0.1, 1., .1),
T=fixed(50),
L=(-0.5, 0., .1),
ini_state=fixed(5.))
widget.children[-2].description = 'Run Simulation'
widget.children[-2].style.button_color = 'lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
# ### Interactive Demo 4.3: LQG with varying control effort costs
#
# Now let's see the performance of the LQG controller. We will use an LQG controller gain, where the control gain is from a system with an infinite-horizon. In this case, the optimal control gain turns out to be a constant.
#
# Vary the value of $\rho$ from $0$ to large values, to see the effect on the state.
# + cellView="form"
#@markdown Execute this cell to include helper function for LQG
class LQG(MyKalmanFilter, LQR):
def __init__(self, T, n_dim_state, n_dim_obs,
transition_matrices, transition_covariance, observation_matrices,
observation_covariance, initial_state_mean, initial_state_covariance, control_matrices):
MyKalmanFilter.__init__(self,n_dim_state, n_dim_obs,
transition_matrices, transition_covariance,
observation_matrices,observation_covariance,
initial_state_mean, initial_state_covariance, control_matrices)
LQR.__init__(self,T, initial_state_mean, transition_covariance)
def control_gain_LQR_infinite(self, rho):
control_gain_LQR_finite = self.control_gain_LQR(self.transition_matrices, self.control_matrices, rho)
return control_gain_LQR_finite[0]
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
def simulate_kf_with_lqg(D=0.9, B=2., C=1., T=50, ini_state=5,
proc_noise=0.1, meas_noise=0.2, rho=1.):
# Format the above variables into a format acccepted by the Kalman Filter
n_dim_state = 1
n_dim_obs = 1
n_timesteps = T
transition_matrices = np.eye(n_dim_state) * D
transition_covariance = np.eye(n_dim_obs) * proc_noise # process noise
observation_matrices = np.eye(n_dim_state) * C
observation_covariance = np.eye(n_dim_obs) * meas_noise
initial_state_mean = np.ones(n_dim_state) * ini_state
initial_state_covariance = np.eye(n_dim_state) * .01
control_matrices = np.eye(n_dim_state) * B
my_kf = MyKalmanFilter(n_dim_state, n_dim_obs, transition_matrices,
transition_covariance, observation_matrices,
observation_covariance, initial_state_mean,
initial_state_covariance, control_matrices)
lqg = LQG(n_timesteps, n_dim_state, n_dim_obs,
transition_matrices, transition_covariance, observation_matrices,
observation_covariance, initial_state_mean, initial_state_covariance,
control_matrices)
control_gain_lqg = lqg.control_gain_LQR_infinite(rho) * np.ones(n_timesteps)
lqg.plot_state_vs_time(n_timesteps, control_gain_lqg, goal = np.zeros(T),
title='State estimation with KF (LQG controller)')
widget=interactive(simulate_kf_with_lqg, {'manual': True},
D = fixed(.9),
B = fixed(2.),
C = fixed(1.),
proc_noise = fixed(.1),
meas_noise = fixed(.2),
T = fixed(50),
ini_state = fixed(5.),
rho=(0., 5., 1.))
widget.children[-2].description = 'Run Simulation'
widget.children[-2].style.button_color = 'lightgreen'
controls = HBox(widget.children[:-1], layout = Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]));
# -
# ### Interactive Demo 4.4: How does the process noise and the measurement noise influence the controlled state and desired action?
#
# Process noise $w_t$ (proc_noise) and measurement noise $v_t$ (meas_noise) have very different effects on the controlled state.
#
# (a) To visualize this, play with the sliders to get an intuition for how process noise and measurement noise influences the controlled state. How are these two sources of noise different?
#
# (b) Next, for varying levels of process noise and measurement noise (note that the control policy is exactly the same for all these values), plot the mean squared error (MSE) between state and the goal, as well as the control cost. What do you notice?
#
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
def lqg_slider(D=0.9, B=2., C=1., T=50, ini_state=5,
proc_noise=2.9, meas_noise=0., rho=1.):
# Format the above variables into a format acccepted by the Kalman Filter
# Format the above variables into a format acccepted by the Kalman Filter
n_dim_state = 1
n_dim_obs = 1
n_timesteps = T
transition_matrices = np.eye(n_dim_state) * D
transition_covariance = np.eye(n_dim_obs) * proc_noise # process noise
observation_matrices = np.eye(n_dim_state) * C
observation_covariance = np.eye(n_dim_obs) * meas_noise
initial_state_mean = np.ones(n_dim_state) * ini_state
initial_state_covariance = np.eye(n_dim_state) * .01
control_matrices = np.eye(n_dim_state) * B
rho = 1
lqg = LQG(n_timesteps, n_dim_state, n_dim_obs,
transition_matrices, transition_covariance, observation_matrices,
observation_covariance, initial_state_mean, initial_state_covariance, control_matrices)
control_gain_lqg = lqg.control_gain_LQR_infinite(rho) * np.ones(n_timesteps)
lqg.plot_state_vs_time(n_timesteps, control_gain_lqg, goal = np.zeros(n_timesteps),
title='State estimation with KF (LQG controller)')
widget=interactive(lqg_slider, {'manual': True},
D = fixed(.9),
B = fixed(2.),
C = fixed(1.),
proc_noise = (0., 3., .1),
meas_noise = (0.1, 3., .1),
T = fixed(50),
ini_state = fixed(5.),
rho=fixed(1.))
widget.children[-2].description = 'Run Simulation'
widget.children[-2].style.button_color = 'lightgreen'
controls = HBox(widget.children[:-1], layout = Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]));
# -
#to_remove explanation
"""
As you increase the process noise, you will notice that it becomes more
difficult to keep the state close to the goal g=0, even though we may have very
little measurement noise (thus can estimate the state exactly).
On the other hand, as you increase the measurement noise, you will notice that
it is harder to estimate the states, and this also may make it harder to keep the
state close to the goal.
Which has a larger effect? How does this effect the required action a[t]?
We will quantify these in the next section.
""";
# ## Section 4.2 Noise effects on the LQG
#
# We can now quantify how the state cost and control costs changes when we change the process and measurement noise levels. To do so, we will run many simulations, stepping through levels of process and measurement noise, tracking MSE and cost of control for each. Run the cell below to perform this simulations and plot them. How do you interpret the results?
# + cellView="form"
#@markdown Execute this cell to to quantify the dependence of state and control
#@markdown cost on process and measurement noise (takes ~20 seconds)
D = 0.9 # state parameter
B = 2 # control parameter
C = 1 # measurement parameter
noise_var = 0.1
T = 200 # time horizon
ini_state = 5 # initial state
process_noise_var = 0.1 # process noise
measurement_noise_var = 0.2 # measurement noise
rho = 1
# Format the above variables into a format acccepted by the Kalman Filter
n_dim_state = 1
n_dim_obs = 1
n_timesteps = T
transition_matrices = np.eye(n_dim_state) * D
transition_covariance = np.eye(n_dim_obs) * noise_var # process noise
observation_matrices = np.eye(n_dim_state) * C
observation_covariance = np.eye(n_dim_obs) * measurement_noise_var
initial_state_mean = np.ones(n_dim_state) * ini_state
initial_state_covariance = np.eye(n_dim_state) * .01
control_matrices = np.eye(n_dim_state) * B
# Implement LQG control over n_iter iterations, and record the MSE between state and goal
MSE_array_N_meas = []
MSE_array_N_proc = []
Jcontrol_array_N_meas = []
Jcontrol_array_N_proc = []
n_iter = 10
meas_noise_array = np.linspace(0,3,20)
proc_noise_array = np.linspace(0.1,3,20)
for i in range(n_iter):
MSE_array = np.zeros(proc_noise_array.shape)
Jcontrol_array = np.zeros(meas_noise_array.shape)
for i in range(len(proc_noise_array)):
transition_covariance = np.eye(n_dim_obs) * proc_noise_array[i]
observation_covariance = np.eye(n_dim_obs) * measurement_noise_var
lqg = LQG(n_timesteps, n_dim_state, n_dim_obs,
transition_matrices, transition_covariance, observation_matrices,
observation_covariance, initial_state_mean, initial_state_covariance, control_matrices)
control_gain_lqg = lqg.control_gain_LQR_infinite(rho) * np.ones(n_timesteps) # Get the control gain
filtered_state_means_impl, filtered_state_covariances_impl, latent, measurement, control = lqg.filter_control(
n_timesteps, control_gain_lqg)
MSE_array[i] = lqg.calculate_J_state(latent)
Jcontrol_array[i] = lqg.calculate_J_control(control)
MSE_array_N_proc.append(MSE_array)
Jcontrol_array_N_proc.append(Jcontrol_array)
MSE_array = np.zeros(meas_noise_array.shape)
Jcontrol_array = np.zeros(meas_noise_array.shape)
for i in range(len(meas_noise_array)):
observation_covariance = np.eye(n_dim_obs) * meas_noise_array[i]
transition_covariance = np.eye(n_dim_obs) * noise_var
lqg = LQG(n_timesteps, n_dim_state, n_dim_obs,
transition_matrices, transition_covariance, observation_matrices,
observation_covariance, initial_state_mean, initial_state_covariance, control_matrices)
control_gain_lqg = lqg.control_gain_LQR_infinite(rho) * np.ones(n_timesteps) # Get the control gain
filtered_state_means_impl, filtered_state_covariances_impl, latent, measurement, control = lqg.filter_control(
n_timesteps, control_gain_lqg)
MSE_array[i] = lqg.calculate_J_state(latent)
Jcontrol_array[i] = lqg.calculate_J_control(control)
MSE_array_N_meas.append(MSE_array)
Jcontrol_array_N_meas.append(Jcontrol_array)
MSE_array_proc_mean = np.mean(np.array(MSE_array_N_proc), axis = 0)
MSE_array_proc_std = np.std(np.array(MSE_array_N_proc), axis = 0)
MSE_array_meas_mean = np.mean(np.array(MSE_array_N_meas), axis = 0)
MSE_array_meas_std = np.std(np.array(MSE_array_N_meas), axis = 0)
Jcontrol_array_proc_mean = np.mean(np.array(Jcontrol_array_N_proc), axis = 0)
Jcontrol_array_proc_std = np.std(np.array(Jcontrol_array_N_proc), axis = 0)
Jcontrol_array_meas_mean = np.mean(np.array(Jcontrol_array_N_meas), axis = 0)
Jcontrol_array_meas_std = np.std(np.array(Jcontrol_array_N_meas), axis = 0)
# Visualize the quantification
f, axs = plt.subplots(2, 2, sharex=True, sharey=True, figsize=(10, 8))
axs[0,0].plot(proc_noise_array, MSE_array_proc_mean, 'r-')
axs[0,0].fill_between(proc_noise_array, MSE_array_proc_mean+MSE_array_proc_std,
MSE_array_proc_mean-MSE_array_proc_std, facecolor='tab:gray', alpha=0.5)
axs[0,0].set_title('Effect of process noise')
axs[0,0].set_ylabel('State Cost (MSE between state and goal)')
axs[0,1].plot(meas_noise_array, MSE_array_meas_mean, 'r-')
axs[0,1].fill_between(meas_noise_array, MSE_array_meas_mean+MSE_array_meas_std,
MSE_array_meas_mean-MSE_array_meas_std, facecolor='tab:gray', alpha=0.5)
axs[0,1].set_title('Effect of measurement noise')
axs[1,0].plot(proc_noise_array, Jcontrol_array_proc_mean, 'r-')
axs[1,0].fill_between(proc_noise_array, Jcontrol_array_proc_mean+Jcontrol_array_proc_std,
Jcontrol_array_proc_mean-Jcontrol_array_proc_std, facecolor='tab:gray', alpha=0.5)
axs[1,0].set_xlabel('Process Noise')
axs[1,0].set_ylabel('Cost of Control')
axs[1,1].plot(meas_noise_array, Jcontrol_array_meas_mean, 'r-')
axs[1,1].fill_between(meas_noise_array, Jcontrol_array_meas_mean+Jcontrol_array_meas_std,
Jcontrol_array_meas_mean-Jcontrol_array_meas_std, facecolor='tab:gray', alpha=0.5)
axs[1,1].set_xlabel('Measurement Noise')
plt.show()
# -
#to_remove explanation
"""
While both sources of noise have an effect on the controlled state, the
process noise has a much larger effect. As the process noise w[t] increases,
state cost (MSE between state and goal) and control cost increase drastically.
You can get an intuition as to why using the sliders in the demo above. To make
matters worse, as the process noise gets larger, you will also need to put in
more effort to keep the system close to the goal.
The measurement noise v[t] also has an effect on the accuracy of the
controlled state. As this noise increases, the MSE between the state and goal
increases. The cost of control in this case remains fairly constant with
increasing levels of measurement noise.
""";
|
tutorials/W3D3_OptimalControl/W3D3_Tutorial2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## <b> Python basics <b>
# #### <b> Variables <b>
# Integer
a = 5
print(a)
a
type(a)
# Float
f = 10.7
f
type(f)
# String
b = '2'
b
# Restricitions for different types, e.g.:
a+b
# But you can do:
a*b
# Combining strings:
c = ' whales swim in the sea'
c
b+c
# You can convert types:
str(a)+c
# #### <b> Everything is an object <b>
# IPython give you list of object's methods, typing dot and pressing TAB:
c.
# Methods are basically default functions that can be applied to our variable:
c.upper()
c.title()
c.count('a')
c.find('the')
# You will get assistance by typing:
# +
# c.find?
# -
# Or byusing an open bracket and press <b>SHIFT+TAB<b>:
c.find(
# Integer variables are also objects:
a.bit_length()
# And you can combine methods:
c.title().count('Wh').bit_length()
# #### <b>Lists<b>
# There are several other interesting variable types in Python, but the one we would need the most is the list.
# In order to create list put coma separated values in square brackets:
l = [1,2,3,4,5]
l
# Sort of similar to Matlab variables, but not exactly.
# Values in list can be any type:
l = ['one', 'two', 'three', 'four', 'five']
l
# Combined
l = ['one', 2, 'three', 4.0, 3+2]
l
# Any type means ANY type:
l = ['one', 2, 'three', [1,2,3,4,5], 3+2]
l
# You can access list values by index:
l[0]
# Yes, zero based numbering! [Look here](http://en.wikipedia.org/wiki/Zero-based_numbering).
l[1]
# Let's have a look at the 4th element of our list:
l[3]
# It's also a list, and its values can be accessed by indexes as well:
l[3][4]
# You also can acces multiple elements of the list using slices:
l
l[1:3]
# Slice will start with the first slice index and go up to but not including the second slice index.
l[3]
# ### <b>Control Structures<b>
# For loop:
# This loop will print all elements from the list *l*
# +
l = ['one', 2, 'three', [1,2,3,4,5], 3+2]
for element in l:
print(element)
# -
# Two interesting thins here. First: indentation, it's in the code, you must use it, otherwise code will not work:
for element in l:
print(element)
# Second - you can iterate through the elements of the list. There is an option to iterate through a bunch of numbers as we used to in Matlab:
for index in range(5):
print(l[index])
# where *range* is just generating a sequence of numbers:
list(range(5))
# ### Branches
# We are not going to use branches in this notes, but this is how they look like just as another example of indentation use:
x = -1
if x > 0:
print("Melting")
elif x == 0:
print("Zero")
else:
print("Freezing")
# ### Modules
# Pure python does not do much. To do some specific tasks you need to import modules. Here I am going to demonstrate several ways to do so.
# The most common one is to import complete library. In this example we import *urllib2* - a library for opening URLs using a variety of protocols.
import requests
# Here we get information from [FESOM](http://fesom.de/) website site. Note how function *get* is called. We have to use name of the library, then dot, then name of the function from the library:
response = requests.get('http://fesom.de/')
response.headers
# Another option is to import it like this:
from requests import *
# In this case all functions will be imported in to the name-space and you can use *get* directly, without typing the name of the library first:
response = get('http://fesom.de/')
response.headers
# But generally this is very bad idea and is not recomended, because your name-space is populated by things that you don't really need and it's hard to tell where the function comes from.
whos
# You can import only function that you need:
from requests import get
response = get('http://fesom.de/')
response.headers
# Or import library as alias in order to avoid extensive typing:
import requests as rq
response = rq.get('http://fesom.de/')
response.headers
# ## Links:
# [Dive Into Python](https://diveintopython3.problemsolving.io/)
|
2_Geospatial_Python/notebooks/Geoscience_intro/02_basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Load and Explore Cifar10 Dataset
# *by <NAME>*
# <img src="../../images/keras-tensorflow-logo.jpg" width="400">
# # Image Classification Task
# Cifar10 is a famous computer-vision dataset used for object recognition.
#
# The dataset consists of:
# - 32x32 pixel colored images
# - 10 classes
# - 6,000 images per classes
# - 50,000 images in the training set
# - 10,000 images in the test set
#
# <img src="../../images/cifar10.png" width="400">
# # Imports
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tensorflow as tf
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
# -
# # Download and Load Cifar10 Dataset
(x_train, y_train), (x_test, y_test) = tf.contrib.keras.datasets.cifar10.load_data()
# # Training Tensor Shape
x_train.shape
# # Testing Tensor Shape
x_test.shape
# # Ploting Helper Function
def plot_10_by_10_images(images):
# figure size
fig = plt.figure(figsize=(10,10))
# plot image grid
for x in range(10):
for y in range(10):
ax = fig.add_subplot(10, 10, 10*y+x+1)
plt.imshow(images[10*y+x])
plt.xticks(np.array([]))
plt.yticks(np.array([]))
plt.show()
# # Explore Cifar10 Dataset
plot_10_by_10_images(x_train[:100])
# ## Next Lesson
# ### SqueezeNet Architecture
# - AlexNet-level accuracy with 50x fewer parameters
# - CNN Squeeze layers and delayed downsampling
#
# <img src="../../images/divider.png" width="100">
|
Advanced Computer Vision with TensorFlow/Code/8609_course-working-files/Section 1/Lesson 1.1/Dataset-Cifar10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: angr
# language: python
# name: angr
# ---
# +
import angr
import angrcli.plugins.context_view
import angrcli.plugins.watches
from angrcli.interaction.explore import ExploreInteractive
# +
proj = angr.Project("./simproc_demo.elf", load_options={'auto_load_libs': False})
cfg = proj.analyses.CFGFast()
argv1 = "FOOBAR"
state = proj.factory.entry_state()
e = ExploreInteractive(proj,state)
# -
|
src/angr/tutorial/angr-workshop/hands_on/simproc_demo-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. CSS intro
# ## 1.1 Show css in markdown
# ```css
# /* Comments in css */
#
# /* Style for H1 */
# h1{
# font-family: 'Courier New', Courier, monospace;
# color: aqua;
# }
#
# /* Style for P*/
# p {
# font-family: 'Times New Roman', Times, serif;
# }
#
# /* .SOMELINK links to class vars*/
# /* .In this example: <p class="Link_specific_HTML_elements"> */
#
# .Link_specific_HTML_elements{
# font-size: larger; color: brown;
# }
#
#
# #Footer_id{
# font-family: 'Courier New', Courier, monospace;
# color: aqua;
# }
#
# .title {
# color:pink;
# }
#
#
# .cursive {
# font-family: cursive;
# }
#
# .capitalize {
# text-transform: capitalize;
# }
#
# h5.destination{
# font-family:Verdana, Geneva, Tahoma, sans-serif;
# }
#
# .Nested li{
# color: seagreen;
# }
#
# ```
# ## 1.2 Write 1.1 into css file in local directory
# +
# %%writefile style.css
/* Comments in css */
/* Style for H1 */
h1{
font-family: 'Courier New', Courier, monospace;
color: aqua;
}
/* Style for P*/
p {
font-family: 'Times New Roman', Times, serif;
}
/* .SOMELINK: links to class variables*/
/* .In this example: <p class="Link_specific_HTML_elements"> */
.Link_specific_HTML_elements{
font-size: larger; color: brown;
}
/* #SOMELINK: links to id variable*/
#Footer_id{
font-family: 'Courier New', Courier, monospace;
color: aqua;
}
/* class variable link*/
.title {
color:pink;
}
/* class variable link*/
.cursive {
font-family: cursive;
}
/* class variable link*/
.capitalize {
text-transform: capitalize;
}
/* class variable link*/
h5.destination{
font-family:Verdana, Geneva, Tahoma, sans-serif;
}
/* class variable link*/
.Nested li{
color: seagreen;
}
# -
# ## 1.3 HTML and CSS
# - In order to use css you need to use the link tag
# - `<link rel="stylesheet" href="./style.css", type="text/css">`
# - CSS path referenced in href="./CSSFILENAME"
# + language="html"
# <!DOCTYPE html>
# <html>
# <head>
# <title>
# Example: Inline styles
# </title>
# <link rel="stylesheet" href="./style.css", type="text/css">
# <body>
# <h1>
# Inline examples
# </h1>
# <p>
# This is how you change the color, font, and size inline
# </p>
# <h2>
# <p class="Link_specific_HTML_elements">
# One class link
# </p>
# </h2>
#
# <h3 class="title cursive capitalize">
# Multiple class link
# </h3>
#
# <h5>
# Chaining
# </h5>
# <ul class="Nested">
# <li>One</li>
# <li>Two</li>
# </ul>
# <h6>
# This is another link
# </h6>
# </body>
# <footer id="Footer_id">
# ID link
# </footer>
# </head>
# </html>
|
CSS/CSS_intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2021년 10월 12일 화요일
# ### BaekJoon - 단어순서 뒤집기 (Python)
# ### 문제 : https://www.acmicpc.net/problem/12605
# ### 블로그 : https://somjang.tistory.com/entry/BaekJoon-12605%EB%B2%88-%EB%8B%A8%EC%96%B4%EC%88%9C%EC%84%9C-%EB%92%A4%EC%A7%91%EA%B8%B0-Python
# ### Solution
# +
def filp_words(idx, word_list):
word_list = word_list.split()
word_list = word_list[::-1]
answer = " ".join(word_list)
return f"Case #{idx + 1}: {answer}"
if __name__ == "__main__":
for i in range(int(input())):
word_list = input()
print(filp_words(i, word_list))
|
DAY 501 ~ 600/DAY510_[BaekJoon] 단어순서 뒤집기 (Python).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Boolean Indexing
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pandas_datareader import DataReader
# %matplotlib inline
# -
# # Calculating boolean statistics
pd.options.display.max_columns = 50
movie = pd.read_csv('data/movie.csv', index_col='movie_title')
movie.head()
movie_2_hours = movie['duration'] > 120
movie_2_hours.head(10)
movie_2_hours.sum()
movie_2_hours.mean()
movie_2_hours.describe()
movie['duration'].dropna().gt(120).mean()
movie_2_hours.value_counts() / movie_2_hours.size
actors = movie[['actor_1_facebook_likes', 'actor_2_facebook_likes']].dropna()
(actors['actor_1_facebook_likes'] > actors['actor_2_facebook_likes']).mean()
# # Constructing multiple boolean conditions
movie = pd.read_csv('data/movie.csv', index_col='movie_title')
movie.head()
# +
criteria1 = movie.imdb_score > 8
criteria2 = movie.content_rating == 'PG-13'
criteria3 = (movie.title_year < 2000) | (movie.title_year >= 2010)
criteria2.head()
# -
criteria_final = criteria1 & criteria2 & criteria3
criteria_final.head()
# # There's more...
movie.title_year < 2000 | movie.title_year > 2009
# # Filtering with boolean indexing
# +
movie = pd.read_csv('data/movie.csv', index_col='movie_title')
crit_a1 = movie.imdb_score > 8
crit_a2 = movie.content_rating == 'PG-13'
crit_a3 = (movie.title_year < 2000) | (movie.title_year > 2009)
final_crit_a = criteria1 & criteria2 & criteria3
# -
crit_b1 = movie.imdb_score < 5
crit_b2 = movie.content_rating == 'R'
crit_b3 = (movie.title_year >= 2000) & (movie.title_year <= 2010)
final_crit_b = crit_b1 & crit_b2 & crit_b3
final_crit_all = final_crit_a | final_crit_b
final_crit_all.head()
movie[final_crit_all].head()
movie_filtered = movie.loc[final_crit_all, ['imdb_score', 'content_rating', 'title_year']]
movie_filtered.head(10)
# # There's more...
final_crit_a2 = (movie.imdb_score > 8) & \
(movie.content_rating == 'PG-13') & \
((movie.title_year < 2000) | (movie.title_year > 2009))
final_crit_a2.equals(final_crit_a)
# # Replicating boolean indexing with index selection
college = pd.read_csv('data/college.csv')
college[college['STABBR'] == 'TX'].head()
college2 = college.set_index('STABBR')
college2.loc['TX'].head()
# %timeit college[college['STABBR'] == 'TX']
# %timeit college2.loc['TX']
# %timeit college2 = college.set_index('STABBR')
# ## There's more...
states =['TX', 'CA', 'NY']
college[college['STABBR'].isin(states)]
college2.loc[states].head()
# # Selecting with unique and sorted indexes
college = pd.read_csv('data/college.csv')
college2 = college.set_index('STABBR')
college2.index.is_monotonic
college3 = college2.sort_index()
college3.index.is_monotonic
# %timeit college[college['STABBR'] == 'TX']
# %timeit college2.loc['TX']
# %timeit college3.loc['TX']
college_unique = college.set_index('INSTNM')
college_unique.index.is_unique
college[college['INSTNM'] == 'Stanford University']
college_unique.loc['Stanford University']
# %timeit college[college['INSTNM'] == 'Stanford University']
# %timeit college_unique.loc['Stanford University']
# ## There's more...
college.index = college['CITY'] + ', ' + college['STABBR']
college = college.sort_index()
college.head()
college.loc['Miami, FL'].head()
# %timeit college[(college['CITY'] == 'Miami') & (college['STABBR'] == 'FL')]
# %timeit college.loc['Miami, FL']
college[(college['CITY'] == 'Miami') & (college['STABBR'] == 'FL')].equals(college.loc['Miami, FL'])
# # Gaining perspective on stock prices
slb = pd.read_csv('data/slb_stock.csv', index_col='Date', parse_dates=['Date'])
slb.head()
slb_close = slb['Close']
slb_summary = slb_close.describe(percentiles=[.1, .9])
slb_summary
upper_10 = slb_summary.loc['90%']
lower_10 = slb_summary.loc['10%']
criteria = (slb_close < lower_10) | (slb_close > upper_10)
slb_top_bottom_10 = slb_close[criteria]
# +
import matplotlib.pyplot as plt
slb_close.plot(color='b', figsize=(12,6))
slb_filtered.plot(marker='o', style=' ', ms=4, color='r')
xmin = criteria.index[0]
xmax = criteria.index[-1]
plt.hlines(y=[lower_10, upper_10], xmin=xmin, xmax=xmax,color='r')
# -
slb_close.plot(color='b', figsize=(12,6))
plt.hlines(y=[lower_10, upper_10], xmin=xmin, xmax=xmax,color='r')
plt.fill_between(criteria.index, lower_10, slb_close.values)
plt.fill_between(criteria.index, lower_10, slb_close.values, where= slb_close < lower_10, color='y')
plt.fill_between(criteria.index, upper_10, slb_close.values, where= slb_close > upper_10, color='y')
# # Translating SQL WHERE clauses
employee = pd.read_csv('data/employee.csv')
employee.DEPARTMENT.value_counts().head()
employee.GENDER.value_counts()
employee.BASE_SALARY.describe().astype(int)
criteria_dept = employee.DEPARTMENT.isin(['Houston Police Department-HPD', 'Houston Fire Department (HFD)'])
criteria_gender = employee.GENDER == 'Female'
criteria_sal = (employee.BASE_SALARY >= 80000) & (employee.BASE_SALARY <= 120000)
criteria_final = criteria_dept & criteria_gender & criteria_sal
select_columns = ['UNIQUE_ID', 'DEPARTMENT', 'GENDER', 'BASE_SALARY']
employee.loc[criteria_final, select_columns].head()
# ## There's more...
criteria_sal = employee.BASE_SALARY.between(80000, 120000)
top_5_depts = employee.DEPARTMENT.value_counts().index[:5]
criteria = ~employee.DEPARTMENT.isin(top_5_depts)
employee[criteria].head()
# # Determing normality of stock market returns
amzn = pd.read_csv('data/amzn_stock.csv', index_col='Date', parse_dates=['Date'])
amzn.head()
amzn_daily_return = amzn.Close.pct_change()
amzn_daily_return.head()
amzn_daily_return = amzn_daily_return.dropna()
amzn_daily_return.hist(bins=20)
mean = amzn_daily_return.mean()
std = amzn_daily_return.std()
abs_z_score = amzn_daily_return.sub(mean).abs().div(std)
within_1 = abs_z_score.lt(1).mean()
within_2 = abs_z_score.lt(2).mean()
within_3 = abs_z_score.lt(3).mean()
print('{:.3f} fall within 1 standard deviation. '
'{:.3f} within 2 and {:.3f} within 3'.format(within_1, within_2, within_3))
def test_return_normality(stock_data):
close = stock_data['Close']
daily_return = close.pct_change().dropna()
daily_return.hist(bins=20)
mean = daily_return.mean()
std = daily_return.std()
abs_z_score = abs(daily_return - mean) / std
within_1 = (abs_z_score < 1).mean()
within_2 = (abs_z_score < 2).mean()
within_3 = (abs_z_score < 3).mean()
print('{:.3f} fall within 1 standard deviation. '
'{:.3f} within 2 and {:.3f} within 3'.format(within_1, within_2, within_3))
slb = pd.read_csv('data/slb_stock.csv', index_col='Date', parse_dates=['Date'])
test_return_normality(slb)
# # Improving readability of boolean indexing with the query method
employee = pd.read_csv('data/employee.csv')
depts = ['Houston Police Department-HPD', 'Houston Fire Department (HFD)']
select_columns = ['UNIQUE_ID', 'DEPARTMENT', 'GENDER', 'BASE_SALARY']
# +
query_string = "DEPARTMENT in @depts " \
"and GENDER == 'Female' " \
"and 80000 <= BASE_SALARY <= 120000"
emp_filtered = employee.query(query_string)
emp_filtered[select_columns].head()
# -
# # There's more...
top10_depts = employee.DEPARTMENT.value_counts().index[:10].tolist()
employee_filtered2 = employee.query("DEPARTMENT not in @top10_depts and GENDER == 'Female'")
employee_filtered2[['DEPARTMENT', 'GENDER']].head()
# # Preserving DataFrames with the where method
movie = pd.read_csv('data/movie.csv', index_col='movie_title')
facebook_likes = movie['actor_1_facebook_likes'].dropna()
facebook_likes.head()
facebook_likes.describe(percentiles=[.1, .25, .5, .75, .9]).astype(int)
facebook_likes.describe(percentiles=[.1,.25,.5,.75,.9])
facebook_likes.hist()
criteria_high = facebook_likes < 20000
criteria_high.mean()
facebook_likes.where(criteria_high).head()
facebook_likes.where(criteria_high, other=20000).head()
criteria_low = facebook_likes > 300
facebook_likes_cap = facebook_likes.where(criteria_high, other=20000)\
.where(criteria_low, 300)
facebook_likes_cap.head()
len(facebook_likes), len(facebook_likes_cap)
facebook_likes_cap.hist()
facebook_likes_cap2 = facebook_likes.clip(lower=300, upper=20000)
facebook_likes_cap2.equals(facebook_likes_cap)
# # Masing DataFrame rows
movie = pd.read_csv('data/movie.csv', index_col='movie_title')
criteria = (movie['title_year'] >= 2010) | (movie['title_year'].isnull())
movie.mask(criteria).head()
movie_mask_filtered = movie.mask(criteria).dropna(how='all')
movie_mask_filtered.head()
movie_boolean_filtered = movie[movie['title_year'] < 2010]
movie_boolean_filtered.head()
movie_mask_filtered.equals(movie_boolean_filtered)
movie_mask_filtered.shape == movie_boolean_filtered.shape
movie_mask_filtered.dtypes == movie_boolean_filtered.dtypes
from pandas.testing import assert_frame_equal
assert_frame_equal(movie_boolean_filtered, movie_mask_filtered, check_dtype=False)
# %timeit movie.mask(criteria).dropna(how='all')
# %timeit movie[movie['title_year'] < 2010]
# # Selecting with booleans, integer location and labels
movie = pd.read_csv('data/movie.csv', index_col='movie_title')
criteria = (movie['content_rating'] == 'G') & (movie['imdb_score'] < 4)
movie_loc = movie.loc[criteria]
movie_loc.head()
movie_loc.equals(movie[criteria])
movie_iloc = movie.iloc[criteria]
movie_iloc = movie.iloc[criteria.values]
movie_iloc.equals(movie_loc)
movie.loc[criteria.values]
criteria_col = movie.dtypes == np.int64
criteria_col.head()
movie.loc[:, criteria_col].head()
movie.iloc[:, criteria_col.values].head()
movie.loc[criteria, ['content_rating', 'imdb_score', 'title_year', 'gross']].sort_values('imdb_score')
cols = ['content_rating', 'imdb_score', 'title_year', 'gross']
col_index = [movie.columns.get_loc(col) for col in cols]
col_index
movie.iloc[criteria.values, col_index].sort_values('imdb_score')
# ## How it works
a = criteria.values
a[:5]
len(a), len(criteria)
# # There's more...
movie.loc[[True, False, True], [True, False, False, True]]
|
Module 3/Chapter05/Chapter 5 Boolean Indexing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ruhan-dave/gradio_wonders_dpl/blob/main/wonders_notebook_final.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="thKdREVzxkHu"
# !pip install fastai2
# !pip install nbdev
import fastai2
from fastai2.vision import *
from nbdev.showdoc import *
# + colab={"base_uri": "https://localhost:8080/"} id="XWmHN_0lv_Nq" outputId="d6e88aa5-2326-4e1c-af96-119746b2af0d"
# !pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
# !ls
# + id="ZI96BmJm9EOp"
from fastbook import *
from fastai.vision.all import *
from fastai.vision.widgets import *
# + id="eMpijX-Gi6RQ"
classes = ["Ch<NAME>", "Christ The Redeemer", "Colosseum", "Machu Picchu", "Petra", "Taj Mahal", "The Great Wall of China"]
# + colab={"base_uri": "https://localhost:8080/", "height": 124} id="SEBFG9Kw2527" outputId="f007fb07-9854-4693-8a35-8fd547ee71c8"
# just for fun, checking out the colab GPU
"""
gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
if gpu_info.find('failed') >= 0:
print('Not connected to a GPU')
else:
print(gpu_info)
["<NAME>", "<NAME>", "<NAME>", "Christ The Redeemer", "Chrysler Building", "CN Tower", "Colosseum", "Disneyland Castle",
"Dome of the Rock", "Eiffel Tower", "Great Mosque of Djenné", "Golen Gate Bridge", "Guggenheim Museum", "House of Parliament and Elizabeth Tower",
"Le Mont-Saint-Michel", "Leaning Tower of Pisa", "Louvre", "<NAME>", "Parthenon", "Petra", "Potala Palace", "Sagrada Família",
"Space Needle", "St. Basil’s Cathedral", "Sultan Ahmed Mosque", "Sydney Opera House", "Taipei 101", "<NAME>", "Temple of Heaven",
"The Flatiron Building", "The Forbidden City", "The Gateway Arch", "The Great Wall of China", "The Lotus Temple", "The Sphinx"]
"""
# + id="7Do7jDWHmVUW"
from google.colab import drive
# + colab={"base_uri": "https://localhost:8080/"} id="GC1LFrT3DvQu" outputId="45ec31cc-e4b4-4dcc-f8d9-cbdd34fff866"
path = 'gdrive/MyDrive/AI_projects/wonders'
p_path = Path(path)
fns = get_image_files(path)
fns
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="1TmXXrV4Sk74" outputId="fe9bf3bb-5054-4206-d4e4-8335d20b6b4e"
path
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="a6gZB50AD-sZ" outputId="f72ed19f-8423-43b9-fea6-9f2f1fe012e2"
"""
failed = verify_images(fns)
failed.map(Path.unlink);
"""
# + colab={"base_uri": "https://localhost:8080/"} id="X1lZGRWazhww" outputId="72e3a13a-fcd8-45f1-a494-10440bbc1b21"
Path.BASE_PATH = path
p_path.ls()
# + colab={"base_uri": "https://localhost:8080/", "height": 107} id="AiN5mpCslmok" outputId="84783e8c-df96-4583-ba5a-94bf9051b894"
# This is just for testing out and understanding regular expressions
"""
import re
string = "wang/ruhan-200<EMAIL>"
re.findall(r"[a-z]+/[a-z]+-[0-9]+@", string) # returns ['wang/ruhan-2001@']
re.findall(r"[0-9]+@[a-z]+.com", string) # returns ['<EMAIL>']
re.findall(r"\d+@[a-z]+", string) # returns ['2<EMAIL>']
fname = 'images/Bengal_82.jpg'
re.findall(r'/(.+)_\d+.jpg$', fname) # if you specify (sth) inside r"...", then only the re instructions inside () will be returned.
link = "images/mechanic/00000052.jpg"
re.findall(r"[a-z]+/[0-9]+.jpg$", link) # returns ['mechanic/00000052.jpg']
re.findall(r"/([a-z]+)/[0-9]+.jpg$", link) # returns ['mechanic']
"""
# + id="X1rfv6RfFQe6"
# continue here
# + id="hEaCzTH5fTc5"
world_wonders = DataBlock(blocks = (ImageBlock, CategoryBlock),
get_items=get_image_files,
splitter=RandomSplitter(valid_pct=0.2, seed=42),
get_y=parent_label,
item_tfms=Resize(460),
batch_tfms=aug_transforms(size=225, mult=1.0, do_flip=True, flip_vert=False, max_rotate=15.0, min_zoom=0.9, max_zoom=1.1,
max_lighting=0.2, max_warp=0.2, p_affine=0.75, p_lighting=0.75, xtra_tfms=None, mode='bilinear', pad_mode='reflection',
align_corners=True, batch=False, min_scale=0.75))
# + id="fkpau0MEcJ8h"
# We need our images to have the same dimensions, so that they can collate into tensors to be passed to the GPU.
# We also want to minimize the number of distinct augmentation computations we perform. The performance requirement suggests that we should,
# where possible, compose our augmentation transforms into fewer transforms (to reduce the number of computations and the number of lossy operations)
# and transform the images into uniform sizes (for more efficient processing on the GPU).
# To work around the challenge of resizing images to larger pixels but with lower resolution, spurious empty zones, and degraded information,
#presizing adopts two strategies that are shown:
# Resize images to relatively "large" dimensions—that is, dimensions significantly larger than the target training dimensions.
# Compose all of the common augmentation operations (including a resize to the final target size) into one, and perform the combined operation
#on the GPU only once at the end of processing, rather than performing the operations individually and interpolating multiple times.
# + id="gx-MmsEFdPzz"
# Crop full width or height: This is in item_tfms, so it's applied to each individual image before it is copied to the GPU.
# It's used to ensure all images are the same size. On the training set, the crop area is chosen randomly.
# On the validation set, the center square of the image is always chosen.
# Random crop and augment: This is in batch_tfms, so it's applied to a batch all at once on the GPU, which means it's fast.
# On the validation set, only the resize to the final size needed for the model is done here.
# On the training set, the random crop and any other augmentations are done first.
# + colab={"base_uri": "https://localhost:8080/"} id="fioO36FNCEa8" outputId="b1306aea-dc43-4f2a-d9dc-5a37633a99bc"
dls = world_wonders.dataloaders(path)
# + id="OtHJIWhgI0TK"
# dls.valid.show_batch(max_n=4, nrows=1)
# + id="cyh8KsSUZIrl"
# world_wonders.summary(path)
# + id="yI-VvF9VOwM6"
# We will compare a few of the small models (resnet 34 and resnet 50, before experimenting with the resnet 101 and 152 to save time
# and we compare these 2 models to see how well they fit the images because having more layers does not necessarily mean lower error rates.
# + id="Dx7cMRpHeoeT"
#learn = cnn_learner(dls, resnet34, metrics=error_rate)
#learn.fine_tune(2)
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="MXpcKBUEsm9O" outputId="32bbac6d-403f-4054-f55c-96bda5167b04"
learn = cnn_learner(dls, resnet34, metrics=error_rate)
learn.fine_tune(4)
# + id="vaaEwVmMtJ8o"
# The table shown when we fit a model shows us the results after each epoch of training.
# Remember, an epoch is one complete pass through all of the images in the data.
# The columns shown are the average loss over the items of the training set, the loss on the validation set,
#and any metrics that we requested—in this case, the error rate.
# + id="ELj0ObOX4y1D"
#cleaner = ImageClassifierCleaner(cnn50)
#cleaner
# for idx in cleaner.delete(): cleaner.fns[idx].unlink()
# for idx, cat in cleaner.change(): shutil.move(str(cleaner.fns[idx]), path/cat)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="l5AkoVTu3SJQ" outputId="7c2cee9a-934a-4bdb-85ab-13d97c5092ea"
steep = learn.lr_find()
# + colab={"base_uri": "https://localhost:8080/"} id="ATJS0hqD4xeh" outputId="a1eb1136-4245-4071-b85b-967a04d644fc"
steep
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="e4oC9Z1h59nr" outputId="a2ac6d63-8710-44e8-b699-bcf0ed00afcd"
learn.fit_one_cycle(2, 2e-5)
# + id="y1TtAuEbJmRQ"
# Yay! This definitely helped the model improve. It looks like we should pick learning rate of "base_lr=5e-3".
# Now we will do transfer learning.
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="ImMARBQWLUjj" outputId="a647b030-6361-4da7-e5a9-5cebc9280a5a"
learn.unfreeze()
learn.lr_find()
# + colab={"base_uri": "https://localhost:8080/", "height": 364} id="VQAxrCpPrMLB" outputId="43ce1b90-a382-4fba-d3a1-85583853664c"
learner = cnn_learner(dls, resnet34, metrics=error_rate)
learner.fit_one_cycle(3, 1e-4)
learner.unfreeze()
learner.fit_one_cycle(6, cbs=EarlyStoppingCallback(monitor='error_rate', patience=2), lr_max=slice(1e-6, 1e-3))
# + colab={"base_uri": "https://localhost:8080/", "height": 620} id="jlr-GgTSZhCy" outputId="9719e86d-4741-4440-aae8-cd6b87761600"
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix(figsize=(10, 10), dpi=60)
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="ZYjG2HPqFsiZ" outputId="6c80960f-f688-458c-c0a9-71b9eb133f3b"
learn = cnn_learner(dls, resnet34, metrics=error_rate)
learn.fine_tune(6)
# + colab={"base_uri": "https://localhost:8080/", "height": 620} id="liu5XsifEOcX" outputId="09c3e5f5-83cd-451c-a7b3-3c0277b5f126"
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix(figsize=(10, 10), dpi=60)
# + id="84sQ3pQlBt8m"
# I'm gonna settle with the model with 3% error rate.
# + id="iogAiCZ-Xotn"
learn.path = Path("gdrive/MyDrive/AI_projects/wonders") # have to include the / to specify where to save it to.
# + id="6cq-msdVSuI4"
learn.export("wonder_app_model.pkl")
#path = Path("gdrive/MyDrwonder_appive/AI_projects/wonders")
#path.ls(file_exts='.pkl')
# + colab={"base_uri": "https://localhost:8080/"} id="mqQwA1geR1va" outputId="5e7a765d-8588-4277-aee7-e9a3e915bd71"
import os
filename = f"{path}/wonder_app_model.pkl"
os.makedirs(os.path.dirname(filename), exist_ok=True)
learn = load_learner(fname=filename)
learn.dls.vocab
# + id="32POgBXGGIYa"
# !pip install gradio
# + colab={"base_uri": "https://localhost:8080/", "height": 663} id="1nfPBrR1xJaI" outputId="c8bdba82-1f69-404d-cac7-0f9b50eb4eb3"
import gradio as gr
def predict_image(img):
prediction = learn.predict(PILImage.create(img))
classes = learn.dls.vocab
probs_list = prediction[2].numpy()
return {c: f"{round(float(probs_list[i]), 4)}" for (i, c) in enumerate(classes)}
image = gr.inputs.Image(shape=(225,225))
label = gr.outputs.Label(num_top_classes=7)
interface = gr.Interface(
fn=predict_image,
inputs=image,
outputs=label
)
interface.launch(debug=True, enable_queue=True)
# 'category': classes[prediction[1].item()],
# + id="rRHYACmhK1pU" colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["d4917bd874934d9a8ce64c27e9bcfec6", "ee9c9807c3ba478a91122616a6b5be3f", "9479fc4813bd413b9ff48a2733415f42"]} outputId="26a2df60-6b01-420d-ded1-1a0d055be648"
upload = widgets.FileUpload()
upload
# + id="ObekhG-_eyNM"
upload.data
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="Dd1kKUR-eRBx" outputId="c4ceb14a-057f-4f85-f9c5-1aafc8ec2bbc"
from fastai.vision.core import *
img = PILImage.create(upload.data[-1])
img
# + colab={"base_uri": "https://localhost:8080/", "height": 150, "referenced_widgets": ["8a203eea78034939beb675a6f19064a9", "20f2f5a5e8f9452f8e3c4fc04870cb03"]} id="Og6qejd8e1f2" outputId="dacab970-73b2-436d-d7c7-941da015d6fa"
out = widgets.Output()
out.clear_output()
with out: display(img.to_thumb(200, 200))
out
# + colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["0c17b7d36d0844a084e9633a4ab3f230", "1c37d793bd8c4266ba5bee9e9c9847f8", "76760b3c911344f1a18d001aaef23deb"]} id="TXt0B4UGgCzi" outputId="9f5f8638-f015-4340-cb35-2d7fd94b1312"
run = widgets.Button(description="Go!")
run
# + [markdown] id="1YpRaHGehv6W"
#
# + id="QVFxwZbThLNs"
def classify(change):
img = PILImage.create(upload.data[-1])
out.clear_output()
with out: display(img.to_thumb(200, 200))
pred, idx, probs = learn.predict(img)
val = f"Looks most like: {pred} with probability: {round(float(probs[idx]), 2)}"
# + id="RJ2qKKNymGxW"
run.on_click(classify)
# + colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["b31b56c3b40345acb42e06c70b4e1bf4", "d21fbd3fce5d4d8687a058ba6af3152f", "21563ef0fdb14f9cab9c719829e7f407"]} id="9Y5ekpjGoO-y" outputId="32dcc524-c951-4ce4-899e-44b7a57a8852"
pred, idx, probs = learn.predict(img)
predict = widgets.Label()
predict.value = f"Looks most like: {pred}. Probability: {100*round(float(probs[idx]), 4)}%"
predict
# + id="HenH31miqsSR"
#{c: f"{round(100*float(probs[i]), 4)}%" for (i, c) in enumerate(classes)}
# + colab={"base_uri": "https://localhost:8080/", "height": 278, "referenced_widgets": ["57fbcfd968dd4d7eb38b4f900525dd75", "daa598ffe8a943649b03e785aeec01eb", "45833638e4dc415fa5b5e80bb2d787a2", "d4917bd874934d9a8ce64c27e9bcfec6", "0c17b7d36d0844a084e9633a4ab3f230", "8a203eea78034939beb675a6f19064a9", "b31b56c3b40345acb42e06c70b4e1bf4", "d494fbf87e414474accddfda25a55f16", "570853e9888249bea782e2eebd8f54c4", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "d21fbd3fce5d4d8687a058ba6af3152f", "<KEY>", "<KEY>"]} id="A862KHHbnoNT" outputId="4ce82494-6ca2-4956-c43a-83b0161edcb1"
VBox([widgets.Label('Select your wonder!'),
upload, run, out, predict])
# + id="c_BxcwdOqkK2"
# !pip install voila
# !jupyter serverextension enable --sys-prefix voila
# + id="UabG78UisC3V"
file = f"/Users/owner/Desktop/wonder_app/wonder_app_model.pkl"
test = load_learner(fname=file)
|
wonders_notebook_final.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linearna regresija
# **Linearna regresija** predstavlja nalaženje linearne veze između nezavisne promenljive $x$ i zavisne promenljive $y$. Ukoliko je promenljiva $x$ skalar, radi se o prostoj linearnoj regresiji, a ako je promenljiva $x$ vektor, reč je o linearnoj regresiji za funkciju više promenljivih. U oba slučaja, pretpostavka je da je zavisna promenljiva $y$ skalar.
# **Prosta linearna regresija** predstavlja nalaženje linearne veze između parova skalara $(x_i, y_i)$, $1 \le i \le n$. Veza je data linearnom funkcijom $f(x) = \beta_0 + \beta_1 x$, gde su $\beta_0$ i $\beta_1$ parametri koje treba odrediti. Ako je $X=[x_1, \dots, x_n]$ i $Y=[y_1, \dots, y_n]$, izraz se može zapisati u matričnom obliku kao $\begin{bmatrix}1&X\end{bmatrix} \cdot \begin{bmatrix}\beta_0\\\beta_1\end{bmatrix} = Y$, gde je $\begin{bmatrix}1&X\end{bmatrix}$ matrica čija se prva kolona sastoji samo od jedinica, a druga kolona je vektor $X$.
# **Linearna regresija za funkciju više promenljivih** predstavlja nalaženje linearne veze između parova vektora i skalara $(X_i, y_i)$, $1 \le i \le n$. Veza je data linearnom funkcijom $f(x_1,\dots,x_m) = \beta_0 + \beta_1 x_1 + \dots + \beta_m x_m$, gde su $\beta_0, \dots, \beta_1$ parametri koje treba odrediti. Izraz se zapisati u matričnom obliku kao $\begin{bmatrix}1&X_1&\cdots&X_m\end{bmatrix} \cdot \begin{bmatrix}\beta_0\\ \vdots \\ \beta_m\end{bmatrix} = Y$, gde je $\begin{bmatrix}1&X_1&\cdots&X_m\end{bmatrix}$ matrica čija se prva kolona sastoji samo od jedinica, a ostale kolone su redom vektori $X_1,\dots,X_m$.
# Sistem jednačina $\begin{bmatrix}1&X\end{bmatrix} \cdot \begin{bmatrix}\beta_0\\\beta_1\end{bmatrix} = Y$ kod proste linearne regresije, odnosno $\begin{bmatrix}1&X_1&\cdots&X_m\end{bmatrix} \cdot \begin{bmatrix}\beta_0\\ \vdots \\ \beta_m\end{bmatrix} = Y$ kod linearne regresije za funkciju više promenljivih, se najčešće rešava **metodom najmanjih kvadrata**. Takav sistem u praksi obično ima više jednačina nego promenljivih i najčešće nema egzaktno rešenje.
# Ako sistem napišemo u obliku $A\beta = Y$, njegovo rešavanje predstavlja nalaženje onog vektora $\beta$ za koji je vrednost izraza $||Y - \beta A||_2^2$ najmanja. Ovde je sa $||a||_2 = \sqrt{a_1^2 + a_2^2 + \dots + a_n^2}$ označena 2-norma vektora $a = (a_1, a_2, \dots, a_n)$. Rešenje je dato u obliku $\beta=(A^{T}A)^{-1}A^{T}Y$, gde se matrica $(A^{T}A)^{-1}A^{T}$ se naziva pseudoinverz matrice $A$.
# U slučaju proste linearne regresije, može se pokazati da je $$\beta_1 = \frac{\sum_{i=1}^n (x_i-\bar{x})(y_i-\bar{y})}{\sum_{i=1}^n (x_i-\bar{x})^2}$$ i $\beta_0 = \bar{y} - \beta_1 \bar{x}$, gde je $\bar{x} = \frac{1}{n} \sum_{i=1}^n x_i$ aritmetička sredina nezavisnih, a $\bar{y} = \frac{1}{n} \sum_{i=1}^n y_i$ zavisnih promenljivih.
# Funkcija `lstsq` iz `numpy.linalg` paketa određuje rešenje sistema jednačina metodom najmanjih kvadrata. Ukoliko postoji rešenje, vraća se njegova vrednost. Inače, vrši se minimizacija $\min_{\beta} ||Y - \beta A||_2^2$. Prva dva argumenta su redom matrica $A$ i vektor $Y$, a treći argument se odnosi na odbacivanje malih singularnih vrednosti matrice. On će u narednim primerima biti postavljen na `None`. Povratne vrednosti su, redom, rešenje, ostatak, rang matrice i singularne vrednosti.
# **Primeri**
# **1.** Rešiti sledeći sistem jednačina dat u matričnom obliku: $$\begin{bmatrix}2&0\\-1&1\\0&2\end{bmatrix} \begin{bmatrix}x_1\\x_2\end{bmatrix} = \begin{bmatrix}2\\0\\-2\end{bmatrix}.$$
# **Rešenje.** Iz prve jednačine sledi da je $x_1 = 1$, iz poslednje $x_2 = -1$, a iz druge $x_1 = x_2$, što je kontradikcija, pa sistem nema rešenja. Rešavanje ovog sistema predstavlja nalaženje vrednosti $x_1$ i $x_2$ koja minimizuje ostatak u smislu metode najmanjih kvadrata. Zadatak će biti urađen na dva načina $-$ direktno preko formule i korišćenjem funkcije `lstsq`.
# Prvi način:
import numpy as np
from numpy import linalg as LA
A = np.array([[2,0], [-1,1], [0,2]])
b = np.array([2, 0, -2]).T
x = LA.inv(A.T.dot(A)).dot(A.T).dot(b)
# rss je skracenica od residual square sum
rss = LA.norm(b - A.dot(x)) ** 2
print('Resenje: x1 =', x[0], 'x2 =', x[1])
print('Ostatak:', rss)
# Drugi način:
x, rss, _, _ = LA.lstsq(A, b, rcond=None)
print('Resenje: x1 =', x[0], 'x2 =', x[1])
# ostatak je dat inace u obliku niza
print('Ostatak:', rss[0])
# **2.** Odrediti koeficijente $\beta_0$ i $\beta_1$ tako da funkcija $f(x) = \beta_0 + \beta_1 x$ u smislu metode najmanjih kvadrata najbolje aproksimira skup tačaka $(0, 1.2)$, $(0.5, 2.05)$, $(1, 2.9)$ i $(-0.5, 0.1)$ u ravni.
# **Rešenje.** Neka su $(x_i, y_i)$, $1 \le i \le 4$ redom parovi tačaka. Nalaženje vrednosti koeficijenata predstavlja rešavanje sistema jednačina $A\beta = y$, gde je $A = \begin{bmatrix}1&x_1\\1&x_2\\1&x_3\\1&x_4\end{bmatrix}$, $\beta = \begin{bmatrix}\beta_0\\ \beta_1\end{bmatrix}$ i $y = \begin{bmatrix}y_1\\y_2\\y_3\\y_4\end{bmatrix}$. Zadatak će biti rešen na tri načina $-$ koristeći funkciju `lstsq`, nalaženjem pseudoinverza i eksplicitnim nalaženjem vrednosti koeficijenata prema obrascu za prostu linearnu regresiju.
# Prvi način:
# +
points = [[0, 1.2], [0.5, 2.05], [1, 2.9], [-0.5, 0.1]]
x = np.array([p[0] for p in points])
y = np.array([p[1] for p in points])
n = x.shape[0]
A = np.vstack((np.ones(n), x)).T
beta0, beta1 = LA.lstsq(A, y.T, rcond=None)[0]
(beta0, beta1)
# -
beta0, beta1 = LA.inv(A.T.dot(A)).dot(A.T).dot(y.T)
(beta0, beta1)
beta1 = np.sum((x - x.mean()) * (y - y.mean())) / np.sum((x - x.mean()) ** 2)
beta0 = y.mean() - beta1 * x.mean()
(beta0, beta1)
# **3.** Odrediti koeficijente $a$, $b$ i $c$ funkcije $f(x,y) = a + b x + c y$ tako da u smislu metode najmanjih kvadrata aproksimira skup tacaka $(3, 3, -1)$, $(5, 3, 1)$ i $(3, 4, 2)$ u prostoru.
# **Rešenje.** Neka su $(x_i, y_i, z_i)$, $1 \le i \le 3$ redom parovi tačaka. Nalaženje vrednosti koeficijenata predstavlja rešavanje sistema jednačina $A\beta = z$, gde je $A = \begin{bmatrix}1&x_1&y_1\\1&x_2&y_2\\1&x_3&y_3\end{bmatrix}$, $\beta = \begin{bmatrix}a\\ b\\ c\end{bmatrix}$ i $z = \begin{bmatrix}z_1\\z_2\\z_3\end{bmatrix}$.
# +
points = [[3,3,-1], [5,3,1], [3,4,2]]
x = np.array([p[0] for p in points])
y = np.array([p[1] for p in points])
z = np.array([p[2] for p in points])
n = x.shape[0]
A = np.vstack((np.ones(n), x, y)).T
solution = LA.lstsq(A, z.T, rcond=None)
a, b, c = solution[0]
(a, b, c)
# -
# **4.** Odrediti vrednosti koeficijenata $a$ i $b$ tako da funkcija $f(x) = a + b \sin x$ u smislu metode najmanjih kvadrata aproksimira skup tacaka $(2, 2.6)$, $(-1.22, -1.7)$, $(8.32, 2.5)$ i $(4.23, -1.6)$ u ravni. Dati ocenu greske. Prikazati skup tačaka i nacrtati rezultujucu funkciju.
# **Rešenje.** Neka su $(x_i, y_i)$, $1 \le i \le 4$ redom parovi tačaka. Nalaženje vrednosti koeficijenata predstavlja rešavanje sistema jednačina $A\beta = y$, gde je $A = \begin{bmatrix}1&\sin x_1\\1&\sin x_2\\1&\sin x_3\\1&\sin x_4\end{bmatrix}$, $\beta = \begin{bmatrix}a\\ b\end{bmatrix}$ i $y = \begin{bmatrix}y_1\\y_2\\y_3\\y_4\end{bmatrix}$.
# +
import matplotlib.pyplot as plt
points = [[2, 2.6], [-1.22, -1.7], [8.32, 2.5], [4.23, -1.6]]
x = np.array([p[0] for p in points])
y = np.array([p[1] for p in points])
A = np.vstack([np.ones(x.shape[0]), np.sin(x)]).T
solution = LA.lstsq(A, y.T, rcond=None)
a, b = solution[0]
rss = solution[1][0]
print('a =', a, 'b =', b)
print('rss = ', rss)
t_points = np.linspace(-2, 10, 100)
t_values = a + b * np.sin(t_points)
plt.plot(t_points, t_values, color='red')
plt.plot(x, y, 'o', color='blue')
plt.show()
# -
# **5.** U datoteci *social_reach.csv* se nalaze cene reklamiranja za različite demografske grupe, koje su date u hiljadama evra za 1000 pregleda. Svaka od tri kolone označava različitu platformu za reklamiranje (na primer, platforme mogu biti Facebook, Instagram ili YouTube). Svaki red označava različitu demografsku grupu, koja može npr. biti posebna država u kojoj se reklama plasira. Potrebno je odrediti iznos sredstava da se ostvari približno milion pregleda za svaku demografsku grupu, gledajući po svim platformama ukupno.
# **Rešenje.** Najpre ćemo učitati i prikazati CSV fajl:
# +
import numpy as np
from numpy import linalg as LA
import matplotlib.pyplot as plt
import pandas as pd
data = pd.read_csv('social_reach.csv')
data
# -
# Neka je $a_{ij}$, $0 \le i \le 9$, $0 \le j \le 2$ cena u hiljadama evra za 1000 pregleda za svaku od 10 demografskih grupa i svaku platformu za reklamiranje. Kako je potrebno odrediti ukupne cene za reklamiranje, potrebno je rešiti sledeći sistem jednačina:
#
# $$\begin{bmatrix}a_{00}&a_{01}&a_{02}\\a_{10}&a_{11}&a_{12}\\ \vdots & \vdots & \vdots\\a_{90}&a_{91}&a_{92}\end{bmatrix}\begin{bmatrix}x_1\\x_2\\x_3\end{bmatrix} = \begin{bmatrix}1000\\1000\\\vdots\\1000\end{bmatrix}.$$
#
# Elementi rezultujućeg vektora su 1000, jer su cene u matrici date za 1000 pregleda, a potrebno je ostvariti milion pregleda. Za svaku demografsku grupu $i$, potrebno je odrediti koeficijente $x_1,x_2,x_3$ tako da važi $a_{i0}x_1+a_{i1}x_2+a_{i2}x_3 \approx 1000$, pa se koeficijenti mogu približno odrediti metodom najmanjih kvadrata.
A = data[['web1', 'web2', 'web3']]
b = 1000 * np.ones(A.shape[0])
x, _, _, _ = LA.lstsq(A, b, rcond=None)
result = pd.Series(np.dot(A, x))
plt.xticks(np.linspace(0, 9, 10))
plt.bar(result.index, result.values)
plt.show()
# Na prethodnom grafiku je prikazan ukupan broj u hiljadama pregleda za svaku demografsku grupu. Ukupna cena koja je plaćena za demografsku grupu $i$ iznosi $a_{i0}x_1+a_{i1}x_2+a_{i2}x_3$ hiljada dolara, gde su vrednosti koordinata vektora $x$ jednake:
print('x1 =', x[0])
print('x2 =', x[1])
print('x3 =', x[2])
|
2019_2020/LinearRegression/Materials/linearna_regresija.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# Import py_entitymatching package
import py_entitymatching as em
import os
import pandas as pd
import pandas_profiling
# Then, read the (sample) input tables
# +
# Get the datasets directory
datasets_dir = em.get_install_path() + os.sep + 'datasets'
# Get the paths of the input tables
path_A = datasets_dir + os.sep + 'dblp_demo.csv'
# -
# Read the CSV files and set 'ID' as the key attribute
A = em.read_csv_metadata(path_A, key='id')
A.head()
# # Data Profiling
pandas_profiling.ProfileReport(A)
# ## Saving the Data Profiling Report to an HTML File
pfr = pandas_profiling.ProfileReport(A)
pfr.to_file("/tmp/example.html")
pfr
|
notebooks/guides/step_wise_em_guides/Data Profiling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple MNIST
import tensorflow as tf
print(tf.__version__)
# ## Download MNIST Dataset
# +
from tensorflow.examples.tutorials.mnist import input_data
# Supress warning and informational messages
old_v = tf.logging.get_verbosity()
tf.logging.set_verbosity(tf.logging.ERROR)
mnist = input_data.read_data_sets("../MNIST_data/", one_hot=True)
# Restore warning and informational messages
tf.logging.set_verbosity(old_v)
print('images shape:', mnist.train.images.shape)
# +
import numpy as np
import matplotlib.pyplot as plt
x_train = mnist.train.images.reshape(mnist.train.images.shape[0], 28, 28)
img_index = 2
print("label: ", mnist.train.labels[img_index])
plt.imshow(1-x_train[img_index], cmap='gray')
# -
# ## Define Model
# +
# x is placeholder for the 28 X 28 image data
x = tf.placeholder(tf.float32, shape=[None, 784])
# y_ is called "y bar" and is a 10 element vector, containing the predicted probability of each
# digit(0-9) class. Such as [0.14, 0.8, 0,0,0,0,0,0,0, 0.06]
y_ = tf.placeholder(tf.float32, [None, 10])
# define weights and balances
W = tf.Variable(tf.zeros([784, 10]))
b = tf.Variable(tf.zeros([10]))
# define our inference model
y = tf.nn.softmax(tf.matmul(x, W) + b)
# loss is cross entropy
#cross_entropy = -tf.reduce_sum(y_ * tf.log(y))
#train_step = tf.train.GradientDescentOptimizer(0.01).minimize(cross_entropy)
cross_entropy = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits_v2(labels=y_, logits=y))
train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
# -
# ## Train
# +
# initialize the global variables
init = tf.global_variables_initializer()
with tf.Session() as sess:
# perform the initialization which is only the initialization of all global variables
sess.run(init)
# Perform 1000 training steps
for i in range(1000):
batch_xs, batch_ys = mnist.train.next_batch(100) # get 100 random data points from the data. batch_xs = image,
# batch_ys = digit(0-9) class
sess.run(train_step, feed_dict={x: batch_xs, y_: batch_ys}) # do the optimization with this data
# Evaluate how well the model did. Do this by comparing the digit with the highest probability in
# actual (y) and predicted (y_).
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
test_accuracy = sess.run(accuracy, feed_dict={x: mnist.test.images, y_: mnist.test.labels})
print("Test Accuracy: {0}%".format(test_accuracy * 100.0))
|
notebooks/04 Classification (Simple MNIST).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: entropy
# language: python
# name: entropy
# ---
import os
import qm
from entropylab import *
import numpy as np
import matplotlib.pyplot as plt
# # Using the HDF5 backend
#
# The data persistence layer of Entropy supports saving your experiment results to either a SQL based database (most simply to a sqlite db) or to the [HDF5 data format](https://en.wikipedia.org/wiki/Hierarchical_Data_Format).
#
# By default, experiment results are saved to an HDF5 file and additional data is saved to the SQL database.
#
# **Note** this is a change from earlier version behavior where all data was saved to a single SQL file.
#
# This notebook shows this feature, how to to deactivate it and how to migrate your existing databses.
#
# We start by setting up the database files. The data base entry point is calling the `SqlAlchemyDB` function with the db file path.
# +
db_file='docs_cache/tutorial.db'
hdf5_file='docs_cache/tutorial.hdf5'
if os.path.exists(db_file):
os.remove(db_file)
os.remove(hdf5_file)
db = SqlAlchemyDB(db_file)
# -
# This creates two files: one is the `.db` file and a second file with the `.hdf5` extension is created next to it, in the same folder.
#
# You can view the contenst of HDF files directly by using programs such as [hdf5 view](https://www.hdfgroup.org/downloads/hdfview/), or programmatically. In python this can be done with the h5py and PyTables packages, but many programming languages and environments (e.g. MATLAB) have tools to work with HDF5 files.
# ## Turning the feature off
#
# As mentioned above, the HDF5 feature is turned on by default.
#
# You can turn it on by using the `enable_hdf5_storage` flag on the `SqlAlchemyDB` function
#
#
db=SqlAlchemyDB(path="mydb.db",enable_hdf5_storage=False) #via ctor
# **Note**:Turning this feature off build a DB with results all contained in the SQL file. If you then want to turn it on, you will need to migrate the DB using the upgrade tool we supply (introduced later on this notebook).
# ### Turning HDF5 off using a configuration file
#
# You can turn off the HDF5 feature by default (instead of using a feature flag).
# To do this, you need to create a file called `setting.toml` next to the `.py` file running your entropy graph.
# The `.toml` file should have the following contents:
# ```toml
# [toggles]
# hdf5_storage = true
# ```
# Even if the `.toml` file is present, the `SqlAlchemyDB` feature flag value overrides the behavior.
# ## Seeing experiment results as they are saved in HDF5
# The following example graph generates a results which is then saved to the HDF5 file
# +
db=SqlAlchemyDB(path=db_file,enable_hdf5_storage=True)
er = ExperimentResources(db)
def node_operation():
return {'res':np.array([1,2,3,4])}
node1 = PyNode(label="first_node", program=node_operation,output_vars={'res'})
experiment = Graph(resources=er, graph={node1}, story="run_a")
handle = experiment.run()
# -
# ## Migration from sqlite
# To migrate an existing DB to HDF5 use the following snippet
file_to_migrate = 'some_sqlite_db.db'
results_backend.sqlalchemy.upgrade_db(path=file_to_migrate)
# The migtation tool copies the data from sqlite file to the hdf5 file. This may take a while to complete on larger database files. Get a coffee while it's working.
#
|
docs/hdf5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Distributions
# +
# #!pip install empiricaldist
# -
from empiricaldist import Pmf
# ### Probability Mass Functions (PMF)
# initiate a Pmf object
coin = Pmf()
coin['heads'] = 1/2
coin['tails'] = 1/2
coin
# initiate a Pmf from a sequence
die = Pmf.from_seq([1,2,3,4,5,6])
die
# Get probs from index
die([1,4,7])
# Initiate from list
letters = Pmf.from_seq(list("Mississippi"))
letters
letters['s'], letters('s'), letters('t'),
#letters['t'] => ERROR => May use default_dict???
|
51_bayes_distributions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.chdir("/gpfs/milgram/project/turk-browne/projects/rtTest/kp_scratch")
print(f"conda env={os.environ['CONDA_DEFAULT_ENV']}")
import sys,pickle
import numpy as np
from sklearn.linear_model import LogisticRegression
import nibabel as nib
def save_obj(obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name):
with open(name + '.pkl', 'rb') as f:
return pickle.load(f)
def Class(data, bcvar):
metas = bcvar[0]
data4d = data[0]
print(data4d.shape)
accs = []
for run in range(6):
print(f"run={run}")
testX = data4d[run]
testY = metas[run]
trainX = data4d[np.arange(6) != run]
trainX = trainX.reshape(trainX.shape[0]*trainX.shape[1], -1)
trainY = []
for meta in range(6):
if meta != run:
trainY.extend(metas[run])
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
# Monitor progress by printing accuracy (only useful if you're running a test set)
acc = clf.score(testX, testY)
accs.append(acc)
return np.mean(accs)
def getMask(topN, subject):
workingDir="/gpfs/milgram/project/turk-browne/projects/rtTest/"
for pn, parc in enumerate(topN):
_mask = nib.load(workingDir+"/{}/{}/{}".format(roiloc, subject, parc))
aff = _mask.affine
_mask = _mask.get_data()
_mask = _mask.astype(int)
# say some things about the mask.
mask = _mask if pn == 0 else mask + _mask
mask[mask>0] = 1
return mask
# tmpFile = sys.argv[1]
tmpFile = "./tmp__folder/0119173_25_schaefer2018_neurosketch_11_0"
print(f"tmpFile={tmpFile}")
[_topN,subject,dataSource,roiloc,N] = load_obj(tmpFile)
_topN=('225.nii.gz',
'232.nii.gz',
'108.nii.gz',
'107.nii.gz',
'238.nii.gz',
'190.nii.gz',
'138.nii.gz',
'221.nii.gz',
'280.nii.gz',
'243.nii.gz')
[bcvar,runs] = load_obj(f"./tmp__folder/{subject}_{dataSource}_{roiloc}_{N}")
_mask=getMask(_topN,subject) ; print('mask dimensions: {}'. format(_mask.shape)) ; print('number of voxels in mask: {}'.format(np.sum(_mask)))
_runs = [runs[:,:,_mask==1]] ; print("Runs shape", _runs[0].shape)
# [_runs,bcvar] = load_obj(tmpFile)
sl_result = Class(_runs, bcvar)
# np.save(tmpFile+'_result',sl_result)
print(f"sl_result={sl_result}")
# -
bcvar[0][0]
# +
'''
这个code的目的是用neurosketch 的数据来检测现在在realtime data里面发现的issue:也就是ceiling有时候竟然比floor更小
这个code的运行逻辑是
用neurosketch前五个run训练2 way classifiers,然后用最后一个run来计算ceiling和floor的值,看是否合理
'''
'''
purpose:
find the best performed mask from the result of aggregate_greedy.py and save as chosenMask
train all possible pairs of 2way classifiers and save for evidence calculation
load saved classifiers and calculate different forms of evidence
steps:
load the result of aggregate_greedy.py
display the result of aggregate_greedy.py
find the best performed ROI for each subject and display the accuracy of each subject, save the best performed ROI as chosenMask
load the functional and behavior data and choseMask and train all possible pairs of 2way classifiers
calculate the evidence floor and ceil for each subject and display different forms of evidences.
'''
'''
load the result of aggregate_greedy.py
'''
# To visualize the greedy result starting for 31 ROIs, in total 25 subjects.
import os
os.chdir("/gpfs/milgram/project/turk-browne/projects/rtTest/kp_scratch/")
from glob import glob
import matplotlib.pyplot as plt
from tqdm import tqdm
import pickle5 as pickle
import subprocess
import numpy as np
import os
print(f"conda env={os.environ['CONDA_DEFAULT_ENV']}")
import numpy as np
import nibabel as nib
import sys
import time
import pandas as pd
from sklearn.linear_model import LogisticRegression
import itertools
import pickle
import subprocess
from subprocess import call
workingDir="/gpfs/milgram/project/turk-browne/projects/rtTest/"
def save_obj(obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name):
with open(name + '.pkl', 'rb') as f:
return pickle.load(f)
roiloc="schaefer2018"
dataSource="neurosketch"
subjects_correctly_aligned=['1206161','0119173','1206162','1130161','1206163','0120171','0111171','1202161','0125172','0110172','0123173','0120173','0110171','0119172','0124171','0123171','1203161','0118172','0118171','0112171','1207162','0117171','0119174','0112173','0112172']
subjects=subjects_correctly_aligned
N=25
workingPath="/gpfs/milgram/project/turk-browne/projects/rtTest/"
GreedyBestAcc=np.zeros((len(subjects),N+1))
GreedyBestAcc[GreedyBestAcc==0]=None
GreedyBestAcc={}
numberOfROIs={}
for ii,subject in enumerate(subjects):
# try:
# GreedyBestAcc[ii,N]=np.load(workingPath+"./{}/{}/output/uniMaskRanktag2_top{}.npy".format(roiloc, subject, N))
# except:
# pass
t=np.load(workingPath+"./{}/{}/output/uniMaskRanktag2_top{}.npy".format(roiloc, subject, N))
GreedyBestAcc[subject]=[np.float(t)]
numberOfROIs[subject]=[N]
# for len_topN_1 in range(N-1,0,-1):
for len_topN in range(1,N):
# Wait(f"./tmp/{subject}_{N}_{roiloc}_{dataSource}_{len_topN_1}.pkl")
try:
# {当前的被试}_{greedy开始的ROI数目,也就是25}_{mask的种类schaefer2018}_{数据来源neurosketch}_{当前的 megaROI 包含有的数目}
di = load_obj(f"./tmp_folder/{subject}_{N}_{roiloc}_{dataSource}_{len_topN}")
GreedyBestAcc[subject].append(np.float(di['bestAcc']))
numberOfROIs[subject].append(len_topN)
# GreedyBestAcc[ii,len_topN] = di['bestAcc']
except:
pass
# '''
# to load the imtermediate results from greedy code to examine the system
# '''
# def wait(tmpFile):
# while not os.path.exists(tmpFile+'_result.npy'):
# time.sleep(5)
# print(f"waiting for {tmpFile}_result.npy\n")
# return np.load(tmpFile+'_result.npy')
# subject= '0119173' #sys.argv[1]
# sub_id = [i for i,x in enumerate(subjects) if x == subject][0]
# intermediate_result=np.zeros((N+1,N+1))
# # 应该有多少?25个24ROI,2个1ROI,24个
# for i in range(N,1,-1):
# for j in range(i):
# tmpFile=f"./tmp_folder/{subject}_{N}_{roiloc}_{dataSource}_{i}_{j}"
# sl_result=wait(tmpFile)
# intermediate_result[i,j]=sl_result
# # _=plt.imshow(intermediate_result)
# #最后一行是25个24ROI,第2行是2个1ROI
'''
display the result of aggregate_greedy.py
'''
# GreedyBestAcc=GreedyBestAcc.T
# plt.imshow(GreedyBestAcc)
# _=plt.figure()
# for i in range(GreedyBestAcc.shape[0]):
# plt.scatter([i]*GreedyBestAcc.shape[1],GreedyBestAcc[i,:],c='g',s=2)
# plt.plot(np.arange(GreedyBestAcc.shape[0]),np.nanmean(GreedyBestAcc,axis=1))
# # plt.ylim([0.19,0.36])
# # plt.xlabel("number of ROIs")
# # plt.ylabel("accuracy")
# _=plt.figure()
# for j in range(GreedyBestAcc.shape[1]):
# plt.plot(GreedyBestAcc[:,j])
# GreedyBestAcc=GreedyBestAcc.T
# _=plt.figure()
# plt.imshow(GreedyBestAcc)
'''
find the best performed ROI for each subject and display the accuracy of each subject, save the best performed ROI as chosenMask
'''
#find best ID for each subject
bestID={}
for ii,subject in enumerate(subjects):
t=GreedyBestAcc[subject]
bestID[subject] = numberOfROIs[subject][np.where(t==np.nanmax(t))[0][0]] #bestID 指的是每一个subject对应的最好的megaROI包含的ROI的数目
chosenMask={}
for subject in bestID:
# best ID
# {当前的被试}_{greedy开始的ROI数目,也就是25}_{mask的种类schaefer2018}_{数据来源neurosketch}_{最好的megaROI 包含有的数目}
di = load_obj(f"./tmp_folder/{subject}_{N}_{roiloc}_{dataSource}_{bestID[subject]}")
chosenMask[subject] = di['bestROIs']
def getMask(topN, subject):
workingDir="/gpfs/milgram/project/turk-browne/projects/rtTest/"
for pn, parc in enumerate(topN):
_mask = nib.load(workingDir+"/{}/{}/{}".format(roiloc, subject, parc))
aff = _mask.affine
_mask = _mask.get_data()
_mask = _mask.astype(int)
# say some things about the mask.
mask = _mask if pn == 0 else mask + _mask
mask[mask>0] = 1
return mask
for sub in chosenMask:
mask=getMask(chosenMask[sub], sub)
# if not os.path.exists(f"{workingDir}/{roiloc}/{sub}/chosenMask.npy"):
np.save(f"{workingDir}/{roiloc}/{sub}/chosenMask",mask)
from scipy.stats import zscore
def normalize(X):
_X=X.copy()
_X = zscore(_X, axis=0)
_X[np.isnan(_X)]=0
return _X
def mkdir(folder):
if not os.path.isdir(folder):
os.mkdir(folder)
'''
load the functional and behavior data and choseMask and train all possible pairs of 2way classifiers
'''
subject= '0119173' #sys.argv[1]
sub_id = [i for i,x in enumerate(subjects) if x == subject][0]
print("best 4way classifier accuracy = ",GreedyBestAcc[subject][bestID[subject]])
'''
purpose:
train offline models
steps:
load preprocessed and aligned behavior and brain data
select data with the wanted pattern like AB AC AD BC BD CD
train correspondng classifier and save the classifier performance and the classifiers themselves.
'''
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn
import joblib
import nibabel as nib
import itertools
from sklearn.linear_model import LogisticRegression
def gaussian(x, mu, sig):
# mu and sig is determined before each neurofeedback session using 2 recognition runs.
return round(1+18*(1 - np.exp(-np.power(x - mu, 2.) / (2 * np.power(sig, 2.))))) # map from (0,1) -> [1,19]
def jitter(size,const=0):
jit = np.random.normal(0+const, 0.05, size)
X = np.zeros((size))
X = X + jit
return X
def other(target):
other_objs = [i for i in ['bed', 'bench', 'chair', 'table'] if i not in target]
return other_objs
def red_vox(n_vox, prop=0.1):
return int(np.ceil(n_vox * prop))
def get_inds(X, Y, pair, testRun=None):
inds = {}
# return relative indices
if testRun:
trainIX = Y.index[(Y['label'].isin(pair)) & (Y['run_num'] != int(testRun))]
else:
trainIX = Y.index[(Y['label'].isin(pair))]
# pull training and test data
trainX = X[trainIX]
trainY = Y.iloc[trainIX].label
# Main classifier on 5 runs, testing on 6th
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
B = clf.coef_[0] # pull betas
# retrieve only the first object, then only the second object
if testRun:
obj1IX = Y.index[(Y['label'] == pair[0]) & (Y['run_num'] != int(testRun))]
obj2IX = Y.index[(Y['label'] == pair[1]) & (Y['run_num'] != int(testRun))]
else:
obj1IX = Y.index[(Y['label'] == pair[0])]
obj2IX = Y.index[(Y['label'] == pair[1])]
# Get the average of the first object, then the second object
obj1X = np.mean(X[obj1IX], 0)
obj2X = np.mean(X[obj2IX], 0)
# Build the importance map
mult1X = obj1X * B
mult2X = obj2X * B
# Sort these so that they are from least to most important for a given category.
sortmult1X = mult1X.argsort()[::-1]
sortmult2X = mult2X.argsort()
# add to a dictionary for later use
inds[clf.classes_[0]] = sortmult1X
inds[clf.classes_[1]] = sortmult2X
return inds
if 'milgram' in os.getcwd():
main_dir='/gpfs/milgram/project/turk-browne/projects/rtTest/'
else:
main_dir='/Users/kailong/Desktop/rtTest'
working_dir=main_dir
os.chdir(working_dir)
objects = ['bed', 'bench', 'chair', 'table']
if dataSource == "neurosketch":
funcdata = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/subjects/{sub}_neurosketch/data/nifti/realtime_preprocessed/{sub}_neurosketch_recognition_run_{run}.nii.gz"
metadata = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/data/features/recog/metadata_{sub}_V1_{phase}.csv"
anat = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/subjects/{sub}_neurosketch/data/nifti/{sub}_neurosketch_anat_mprage_brain.nii.gz"
elif dataSource == "realtime":
funcdata = "/gpfs/milgram/project/turk-browne/projects/rtcloud_kp/subjects/{sub}/ses{ses}_recognition/run0{run}/nifti/{sub}_functional.nii.gz"
metadata = "/gpfs/milgram/project/turk-browne/projects/rtcloud_kp/subjects/{sub}/ses{ses}_recognition/run0{run}/{sub}_0{run}_preprocessed_behavData.csv"
anat = "$TO_BE_FILLED"
else:
funcdata = "/gpfs/milgram/project/turk-browne/projects/rtTest/searchout/feat/{sub}_pre.nii.gz"
metadata = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/data/features/recog/metadata_{sub}_V1_{phase}.csv"
anat = "$TO_BE_FILLED"
# print('mask dimensions: {}'. format(mask.shape))
# print('number of voxels in mask: {}'.format(np.sum(mask)))
phasedict = dict(zip([1,2,3,4,5,6],["12", "12", "34", "34", "56", "56"]))
imcodeDict={"A": "bed", "B": "Chair", "C": "table", "D": "bench"}
chosenMask = np.load(f"/gpfs/milgram/project/turk-browne/projects/rtTest/schaefer2018/{subject}/chosenMask.npy")
print(f"np.sum(chosenMask)={np.sum(chosenMask)}")
# Compile preprocessed data and corresponding indices
metas = []
for run in range(1, 7):
print(run, end='--')
# retrieve from the dictionary which phase it is, assign the session
phase = phasedict[run]
# Build the path for the preprocessed functional data
this4d = funcdata.format(run=run, phase=phase, sub=subject)
# Read in the metadata, and reduce it to only the TR values from this run, add to a list
thismeta = pd.read_csv(metadata.format(run=run, phase=phase, sub=subject))
if dataSource == "neurosketch":
_run = 1 if run % 2 == 0 else 2
else:
_run = run
thismeta = thismeta[thismeta['run_num'] == int(_run)]
if dataSource == "realtime":
TR_num = list(thismeta.TR.astype(int))
labels = list(thismeta.Item)
labels = [imcodeDict[label] for label in labels]
else:
TR_num = list(thismeta.TR_num.astype(int))
labels = list(thismeta.label)
print("LENGTH OF TR: {}".format(len(TR_num)))
# Load the functional data
runIm = nib.load(this4d)
affine_mat = runIm.affine
runImDat = runIm.get_fdata()
# Use the TR numbers to select the correct features
features = [runImDat[:,:,:,n+3] for n in TR_num] # here shape is from (94, 94, 72, 240) to (80, 94, 94, 72)
features = np.array(features)
features = features[:, chosenMask==1]
print("shape of features", features.shape, "shape of chosenMask", chosenMask.shape)
features = normalize(features)
# features = np.expand_dims(features, 0)
# Append both so we can use it later
# metas.append(labels)
# metas['label']
t=pd.DataFrame()
t['label']=labels
t["run_num"]=run
behav_data=t if run==1 else pd.concat([behav_data,t])
runs = features if run == 1 else np.concatenate((runs, features))
dimsize = runIm.header.get_zooms()
brain_data = runs
print(brain_data.shape)
print(behav_data.shape)
FEAT=brain_data
print(f"FEAT.shape={FEAT.shape}")
META=behav_data
def Class(brain_data,behav_data):
accs = []
for run in range(1,7):
trainIX = behav_data['run_num']!=int(run)
testIX = behav_data['run_num']==int(run)
trainX = brain_data[trainIX]
trainY = behav_data.iloc[np.asarray(trainIX)].label
testX = brain_data[testIX]
testY = behav_data.iloc[np.asarray(testIX)].label
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
# Monitor progress by printing accuracy (only useful if you're running a test set)
acc = clf.score(testX, testY)
accs.append(acc)
accs
return np.mean(accs)
accs=Class(brain_data,behav_data)
print(f"new trained 4 way classifier accuracy={accs}")
# +
for run in range(6):
t=brain_data[run*80:(run+1)*80,:]
t=np.expand_dims(t,0)
a= t if run == 0 else np.concatenate((a, t))
a=[a]
b=[]
for run in range(6):
b.append(list(behav_data[behav_data['run_num']==(run+1)].label))
b=[b]
def Class(data, bcvar):
metas = bcvar[0]
data4d = data[0]
print(data4d.shape)
accs = []
for run in range(6):
print(f"run={run}")
testX = data4d[run]
testY = metas[run]
trainX = data4d[np.arange(6) != run]
trainX = trainX.reshape(trainX.shape[0]*trainX.shape[1], -1)
trainY = []
for meta in range(6):
if meta != run:
trainY.extend(metas[run])
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
# Monitor progress by printing accuracy (only useful if you're running a test set)
acc = clf.score(testX, testY)
accs.append(acc)
return np.mean(accs)
Class(a,b)
# +
for run in range(6):
t=brain_data[run*80:(run+1)*80,:]
t=np.expand_dims(t,0)
a= t if run == 0 else np.concatenate((a, t))
a=[a]
b=[]
for run in range(6):
b.append(list(behav_data[behav_data['run_num']==(run+1)].label))
b=[b]
def Class(data, bcvar):
metas = bcvar[0]
data4d = data[0]
print(data4d.shape)
accs = []
for run in range(6):
print(f"run={run}")
testX = data4d[run]
testY = metas[run]
trainX = data4d[np.arange(6) != run]
trainX = trainX.reshape(trainX.shape[0]*trainX.shape[1], -1)
trainY = []
for meta in range(6):
if meta != run:
trainY.extend(metas[meta])
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
# Monitor progress by printing accuracy (only useful if you're running a test set)
acc = clf.score(testX, testY)
accs.append(acc)
return np.mean(accs)
print(Class(a,b))
def Class(brain_data,behav_data):
accs = []
for run in range(1,7):
trainIX = behav_data['run_num']!=int(run)
testIX = behav_data['run_num']==int(run)
trainX = brain_data[trainIX]
trainY = behav_data[np.asarray(trainIX)].label
testX = brain_data[testIX]
testY = behav_data[np.asarray(testIX)].label
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
# Monitor progress by printing accuracy (only useful if you're running a test set)
acc = clf.score(testX, testY)
accs.append(acc)
accs
return np.mean(accs)
accs=Class(brain_data,behav_data)
print(f"new trained 4 way classifier accuracy={accs}")
# +
for run in range(6):
t=brain_data[run*80:(run+1)*80,:]
t=np.expand_dims(t,0)
a= t if run == 0 else np.concatenate((a, t))
a=[a]
b=[]
for run in range(6):
b.append(list(behav_data[behav_data['run_num']==(run+1)].label))
b=[b]
data, bcvar=a,b
metas = bcvar[0]
data4d = data[0]
print(data4d.shape)
accs = []
for run in range(6):
print(f"run={run}")
testX = data4d[run]
testY = metas[run]
trainX = data4d[np.arange(6) != run]
trainX = trainX.reshape(trainX.shape[0]*trainX.shape[1], -1)
trainY = []
for meta in range(6):
if meta != run:
trainY.extend(metas[meta])
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
# Monitor progress by printing accuracy (only useful if you're running a test set)
acc = clf.score(testX, testY)
accs.append(acc)
accs
# +
# for a,b in zip(trainY,_trainY):
# print(a==b)
run=2
behav_data['run_num']==run
# +
trainIX = behav_data['run_num']!=run
testIX = behav_data['run_num']==run
trainX = brain_data[trainIX]
trainY = list(behav_data[trainIX].label)
trainY
# +
'''
purpose:
according to the given number of ROIs N, pick up the top N ROIs accuracy and combine them for a combined mask and retrain the model and getting result
steps:
load accuracy for all the ROIs for given subject
pick up the top N ROIs
combine these top N masks
retrain the model and get the accuracy.
get the N combinations of N-1 ROIs
retrain the model and get the accuracy for these N combinations
get the N-1 combinations of N-2 ROIs
retrain the model and get the accuracy for these N-1 combinations
'''
'''
you could try to see whether combining parcels improves performance.
That's going to be the most important bit, because we'll want to decide on a tradeoff between number of voxels and accuracy.
The script of interest here is aggregate.sh which is just a feeder for aggregate.py.
This will use the .npy outputs of classRegion.py to select and merge the top N ROIs/parcels, and will return the list of ROI names, the number of voxels, and the cross-validated classifier accuracy
in this newly combined larger mask. An example run of this is as follows:
sbatch aggregate.sh 0111171 neurosketch schaefer2018 15
'''
import os
print(f"conda env={os.environ['CONDA_DEFAULT_ENV']}")
import numpy as np
import nibabel as nib
import sys
import time
import pandas as pd
from sklearn.linear_model import LogisticRegression
import itertools
# from tqdm import tqdm
import pickle
import subprocess
from subprocess import call
def save_obj(obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name):
with open(name + '.pkl', 'rb') as f:
return pickle.load(f)
# What subject are you running
'''
Takes args (in order):
subject (e.g. 0111171)
dataSource (e.g. neurosketch, but also realtime)
roiloc (wang2014 or schaefer2018)
N (the number of parcels or ROIs to start with)
'''
global subject,dataSource,roiloc,N
subject="0119173" #sys.argv[1]
# dataSource,roiloc,N=sys.argv[2],sys.argv[3],int(sys.argv[4])
dataSource,roiloc,N="neurosketch","schaefer2018",25
print("Running subject {}, with {} as a data source, {}, starting with {} ROIs".format(subject, dataSource, roiloc, N))
# dataSource depending, there are a number of keywords to fill in:
# ses: which day of data collection
# run: which run number on that day (single digit)
# phase: 12, 34, or 56
# sub: subject number
if dataSource == "neurosketch":
funcdata = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/subjects/{sub}_neurosketch/data/nifti/realtime_preprocessed/{sub}_neurosketch_recognition_run_{run}.nii.gz"
metadata = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/data/features/recog/metadata_{sub}_V1_{phase}.csv"
anat = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/subjects/{sub}_neurosketch/data/nifti/{sub}_neurosketch_anat_mprage_brain.nii.gz"
elif dataSource == "realtime":
funcdata = "/gpfs/milgram/project/turk-browne/projects/rtcloud_kp/subjects/{sub}/ses{ses}_recognition/run0{run}/nifti/{sub}_functional.nii.gz"
metadata = "/gpfs/milgram/project/turk-browne/projects/rtcloud_kp/subjects/{sub}/ses{ses}_recognition/run0{run}/{sub}_0{run}_preprocessed_behavData.csv"
anat = "$TO_BE_FILLED"
else:
funcdata = "/gpfs/milgram/project/turk-browne/projects/rtTest/searchout/feat/{sub}_pre.nii.gz"
metadata = "/gpfs/milgram/project/turk-browne/jukebox/ntb/projects/sketchloop02/data/features/recog/metadata_{sub}_V1_{phase}.csv"
anat = "$TO_BE_FILLED"
workingDir="/gpfs/milgram/project/turk-browne/projects/rtTest/"
starttime = time.time()
tag="tag2"
# '1201161', '1121161','0115172','0112174' #these subject have nothing in output folder
subjects_correctly_aligned=['1206161','0119173','1206162','1130161','1206163','0120171','0111171','1202161','0125172','0110172','0123173','0120173','0110171','0119172','0124171','0123171','1203161','0118172','0118171','0112171','1207162','0117171','0119174','0112173','0112172']
if roiloc == "schaefer2018":
RESULT=np.empty((len(subjects_correctly_aligned),300))
topN = []
for ii,sub in enumerate(subjects_correctly_aligned):
outloc = workingDir+"/{}/{}/output".format(roiloc, sub)
for roinum in range(1,301):
# result = np.load("{}/{}.npy".format(outloc, roinum))
result = np.load(f"{outloc}/{tag}_{roinum}.npy")
RESULT[ii,roinum-1]=result
# RESULT = result if roinum == 1 else np.vstack((RESULT, result))
RESULT = np.mean(RESULT,axis=0)
print(f"RESULT.shape={RESULT.shape}")
RESULTix = RESULT[:].argsort()[-N:]
for idx in RESULTix:
topN.append("{}.nii.gz".format(idx+1))
# print(topN[-1])
else:
RESULT_all=[]
topN = []
for ii,sub in enumerate(subjects_correctly_aligned):
outloc = workingDir+"/{}/{}/output".format(roiloc, sub)
for hemi in ["lh", "rh"]:
for roinum in range(1, 26):
# result = np.load("{}/roi{}_{}.npy".format(outloc, roinum, hemi))
result = np.load(f"{outloc}/{tag}_roi{roinum}_{hemi}.npy")
Result = result if roinum == 1 else np.vstack((Result, result))
RESULT = Result if hemi == "lh" else np.hstack((RESULT, Result))
RESULT_all.append(RESULT)
RESULT_all=np.asarray(RESULT_all)
print(f"RESULT_all.shape={RESULT_all.shape}")
RESULT_all=np.mean(RESULT_all,axis=0)
print(f"RESULT_all.shape={RESULT_all.shape}")
RESULT1d = RESULT.flatten()
RESULTix = RESULT1d.argsort()[-N:]
x_idx, y_idx = np.unravel_index(RESULTix, RESULT.shape)
# Check that we got the largest values.
for x, y, in zip(x_idx, y_idx):
print(x,y)
if y == 0:
topN.append("roi{}_lh.nii.gz".format(x+1))
else:
topN.append("roi{}_rh.nii.gz".format(x+1))
# print(topN[-1])
print(f"len(topN)={len(topN)}")
print(f"topN={topN}")
def Wait(waitfor, delay=1):
while not os.path.exists(waitfor):
time.sleep(delay)
print('waiting for {}'.format(waitfor))
def normalize(X):
from scipy.stats import zscore
# X = X - X.mean(0)
X = zscore(X, axis=0)
X[np.isnan(X)]=0
return X
def Class(data, bcvar):
metas = bcvar[0]
data4d = data[0]
print(data4d.shape)
accs = []
for run in range(6):
testX = data4d[run]
testY = metas[run]
trainX = data4d[np.arange(6) != run]
trainX = trainX.reshape(trainX.shape[0]*trainX.shape[1], -1)
trainY = []
for meta in range(6):
if meta != run:
trainY.extend(metas[meta])
clf = LogisticRegression(penalty='l2',C=1, solver='lbfgs', max_iter=1000,
multi_class='multinomial').fit(trainX, trainY)
# Monitor progress by printing accuracy (only useful if you're running a test set)
acc = clf.score(testX, testY)
accs.append(acc)
return np.mean(accs)
phasedict = dict(zip([1,2,3,4,5,6],["12", "12", "34", "34", "56", "56"]))
imcodeDict={"A": "bed", "B": "Chair", "C": "table", "D": "bench"}
def getMask(topN, subject):
workingDir="/gpfs/milgram/project/turk-browne/projects/rtTest/"
for pn, parc in enumerate(topN):
_mask = nib.load(workingDir+"/{}/{}/{}".format(roiloc, subject, parc))
aff = _mask.affine
_mask = _mask.get_data()
_mask = _mask.astype(int)
# say some things about the mask.
mask = _mask if pn == 0 else mask + _mask
mask[mask>0] = 1
return mask
mask=getMask(topN, subject)
print('mask dimensions: {}'. format(mask.shape))
print('number of voxels in mask: {}'.format(np.sum(mask)))
# Compile preprocessed data and corresponding indices
metas = []
for run in range(1, 7):
print(run, end='--')
# retrieve from the dictionary which phase it is, assign the session
phase = phasedict[run]
ses = 1
# Build the path for the preprocessed functional data
this4d = funcdata.format(ses=ses, run=run, phase=phase, sub=subject)
# Read in the metadata, and reduce it to only the TR values from this run, add to a list
thismeta = pd.read_csv(metadata.format(ses=ses, run=run, phase=phase, sub=subject))
if dataSource == "neurosketch":
_run = 1 if run % 2 == 0 else 2
else:
_run = run
thismeta = thismeta[thismeta['run_num'] == int(_run)]
if dataSource == "realtime":
TR_num = list(thismeta.TR.astype(int))
labels = list(thismeta.Item)
labels = [imcodeDict[label] for label in labels]
else:
TR_num = list(thismeta.TR_num.astype(int))
labels = list(thismeta.label)
print("LENGTH OF TR: {}".format(len(TR_num)))
# Load the functional data
runIm = nib.load(this4d)
affine_mat = runIm.affine
runImDat = runIm.get_data()
# Use the TR numbers to select the correct features
features = [runImDat[:,:,:,n+3] for n in TR_num]
features = np.array(features)
# features = features[:, mask==1]
print("shape of features", features.shape, "shape of mask", mask.shape)
# featmean = features.mean(1).mean(1).mean(1)[..., None,None,None] #features.mean(1)[..., None]
# features = features - featmean
# features = features - features.mean(0)
features = normalize(features)
features = np.expand_dims(features, 0)
# Append both so we can use it later
metas.append(labels)
runs = features if run == 1 else np.concatenate((runs, features))
dimsize = runIm.header.get_zooms()
# Preset the variables
print("Runs shape", runs.shape)
bcvar = [metas]
save_obj([bcvar,runs],f"./tmp__folder/{subject}_{dataSource}_{roiloc}_{N}") #{len(topN)}_{i}
# # Distribute the information to the searchlights (preparing it to run)
# _runs = [runs[:,:,mask==1]]
# print("Runs shape", _runs[0].shape)
# slstart = time.time()
# sl_result = Class(_runs, bcvar)
# print("results of classifier: {}, type: {}".format(sl_result, type(sl_result)))
# SL = time.time() - slstart
# tot = time.time() - starttime
# print('total time: {}, searchlight time: {}'.format(tot, SL))
def wait(tmpFile):
while not os.path.exists(tmpFile+'_result.npy'):
time.sleep(5)
print(f"waiting for {tmpFile}_result.npy\n")
return np.load(tmpFile+'_result.npy')
def numOfRunningJobs():
# subprocess.Popen(['squeue -u kp578 | wc -l > squeue.txt'],shell=True) # sl_result = Class(_runs, bcvar)
randomID=str(time.time())
# print(f"squeue -u kp578 | wc -l > squeue/{randomID}.txt")
call(f'squeue -u kp578 | wc -l > squeue/{randomID}.txt',shell=True)
numberOfJobsRunning = int(open(f"squeue/{randomID}.txt", "r").read())
print(f"numberOfJobsRunning={numberOfJobsRunning}")
return numberOfJobsRunning
if not os.path.exists(f"./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_{len(topN)}.pkl"):
_runs = [runs[:,:,mask==1]]
print("Runs shape", _runs[0].shape)
slstart = time.time()
sl_result = Class(_runs, bcvar)
save_obj({"subject":subject,
"startFromN":N,
"currNumberOfROI":len(topN),
"bestAcc":sl_result, # this is the sl_result for the topN, not the bestAcc, bestAcc is for the purpose of keeping consistent with others
"bestROIs":topN},# this is the topN, not the bestROIs, bestROIs is for the purpose of keeping consistent with others
f"./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_{len(topN)}"
)
# ./tmp__folder/0125171_40_schaefer2018_neurosketch_39.pkl
if os.path.exists(f"./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_{1}.pkl"):
print(f"./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_1.pkl exists")
raise Exception('runned or running')
# N-1
def next(topN):
print(f"len(topN)={len(topN)}")
print(f"topN={topN}")
if len(topN)==1:
return None
else:
try:
allpairs = itertools.combinations(topN,len(topN)-1)
topNs=[]
sl_results=[]
tmpFiles=[]
while os.path.exists("./tmp__folder/holdon.npy"):
time.sleep(10)
print("sleep for 10s ; waiting for ./tmp__folder/holdon.npy to be deleted")
# np.save("./tmp__folder/holdon",1)
for i,_topN in enumerate(allpairs):
tmpFile=f"./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_{len(topN)}_{i}"
print(f"tmpFile={tmpFile}")
topNs.append(_topN)
tmpFiles.append(tmpFile)
if not os.path.exists(tmpFile+'_result.npy'):
# prepare brain data(runs) mask and behavior data(bcvar)
# save_obj([_topN,subject,dataSource,roiloc,N], tmpFile)
print("kp2")
# numberOfJobsRunning = numOfRunningJobs()
print("kp3")
# while numberOfJobsRunning > 400: # 300 is not filling it up
# print("kp4 300")
# print("waiting 10, too many jobs running") ; time.sleep(10)
# numberOfJobsRunning = numOfRunningJobs()
# print("kp5")
# get the evidence for the current mask
# print(f'sbatch class.sh {tmpFile}')
# proc = subprocess.Popen([f'sbatch --requeue class.sh {tmpFile}'],shell=True) # sl_result = Class(_runs, bcvar)
tmpFile = tmpFile #sys.argv[1]
print(f"tmpFile={tmpFile}")
# [_topN,subject,dataSource,roiloc,N] = load_obj(tmpFile)
# [bcvar,runs] = load_obj(f"./tmp__folder/{subject}_{dataSource}_{roiloc}_{N}")
_mask=getMask(_topN,subject) ; print('mask dimensions: {}'. format(_mask.shape)) ; print('number of voxels in mask: {}'.format(np.sum(_mask)))
_runs = [runs[:,:,_mask==1]] ; print("Runs shape", _runs[0].shape)
# [_runs,bcvar] = load_obj(tmpFile)
sl_result = Class(_runs, bcvar)
np.save(tmpFile+'_result',sl_result)
print(f"sl_result={sl_result}")
# outs, errs = proc.communicate(timeout=5)
# print(f"outs={outs}")
# print(f"errs={errs}")
print("kp6")
else:
print(tmpFile+'_result.npy exists!')
# os.remove("./tmp__folder/holdon.npy")
sl_results=[]
for tmpFile in tmpFiles:
sl_result=wait(tmpFile)
sl_results.append(sl_result)
print(f"sl_results={sl_results}")
print(f"max(sl_results)=={max(sl_results)}")
maxID=np.where(sl_results==max(sl_results))[0][0]
save_obj({"subject":subject,
"startFromN":N,
"currNumberOfROI":len(topN)-1,
"bestAcc":max(sl_results),
"bestROIs":topNs[maxID]},
f"./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_{len(topN)-1}"
)
print(f"bestAcc={max(sl_results)} For {len(topN)-1} = ./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_{len(topN)-1}")
tmpFiles=next(topNs[maxID])
except:
return tmpFiles
tmpFiles=next(topN)
# +
'''
load the result of aggregate_greedy.py
'''
# To visualize the greedy result starting for 31 ROIs, in total 25 subjects.
import os
os.chdir("/gpfs/milgram/project/turk-browne/projects/rtTest/kp_scratch/")
from glob import glob
import matplotlib.pyplot as plt
from tqdm import tqdm
import pickle5 as pickle
import subprocess
import numpy as np
import os
print(f"conda env={os.environ['CONDA_DEFAULT_ENV']}")
import numpy as np
import nibabel as nib
import sys
import time
import pandas as pd
from sklearn.linear_model import LogisticRegression
import itertools
import pickle
import subprocess
from subprocess import call
workingDir="/gpfs/milgram/project/turk-browne/projects/rtTest/"
def save_obj(obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name):
with open(name + '.pkl', 'rb') as f:
return pickle.load(f)
roiloc="schaefer2018"
dataSource="neurosketch"
subjects_correctly_aligned=['1206161','0119173','1206162','1130161','1206163','0120171','0111171','1202161','0125172','0110172','0123173','0120173','0110171','0119172','0124171','0123171','1203161','0118172','0118171','0112171','1207162','0117171','0119174','0112173','0112172']
subjects=subjects_correctly_aligned
N=25
workingPath="/gpfs/milgram/project/turk-browne/projects/rtTest/"
GreedyBestAcc=np.zeros((len(subjects),N+1))
GreedyBestAcc[GreedyBestAcc==0]=None
GreedyBestAcc={}
numberOfROIs={}
for ii,subject in enumerate(subjects):
# try:
# GreedyBestAcc[ii,N]=np.load(workingPath+"./{}/{}/output/uniMaskRanktag2_top{}.npy".format(roiloc, subject, N))
# except:
# pass
t=np.load(workingPath+"./{}/{}/output/uniMaskRanktag2_top{}.npy".format(roiloc, subject, N))
GreedyBestAcc[subject]=[np.float(t)]
numberOfROIs[subject]=[N]
for len_topN in range(N-1,0,-1):
# for len_topN in range(1,N):
# Wait(f"./tmp/{subject}_{N}_{roiloc}_{dataSource}_{len_topN_1}.pkl")
try:
# {当前的被试}_{greedy开始的ROI数目,也就是25}_{mask的种类schaefer2018}_{数据来源neurosketch}_{当前的 megaROI 包含有的数目}
di = load_obj(f"./tmp__folder/{subject}_{N}_{roiloc}_{dataSource}_{len_topN}")
GreedyBestAcc[subject].append(np.float(di['bestAcc']))
numberOfROIs[subject].append(len_topN)
# GreedyBestAcc[ii,len_topN] = di['bestAcc']
except:
pass
# '''
# to load the imtermediate results from greedy code to examine the system
# '''
# def wait(tmpFile):
# while not os.path.exists(tmpFile+'_result.npy'):
# time.sleep(5)
# print(f"waiting for {tmpFile}_result.npy\n")
# return np.load(tmpFile+'_result.npy')
# subject= '0119173' #sys.argv[1]
# sub_id = [i for i,x in enumerate(subjects) if x == subject][0]
# intermediate_result=np.zeros((N+1,N+1))
# # 应该有多少?25个24ROI,2个1ROI,24个
# for i in range(N,1,-1):
# for j in range(i):
# tmpFile=f"./tmp_folder/{subject}_{N}_{roiloc}_{dataSource}_{i}_{j}"
# sl_result=wait(tmpFile)
# intermediate_result[i,j]=sl_result
# # _=plt.imshow(intermediate_result)
# #最后一行是25个24ROI,第2行是2个1ROI
'''
display the result of aggregate_greedy.py
'''
print(numberOfROIs)
# GreedyBestAcc=GreedyBestAcc.T
_GreedyBestAcc=np.zeros((len(subjects),N+1))
_GreedyBestAcc[_GreedyBestAcc==0]=None
for ii,sub in enumerate(GreedyBestAcc):
_GreedyBestAcc[ii,0:len(GreedyBestAcc[sub])]=GreedyBestAcc[sub]
if len(GreedyBestAcc[sub])<5:
print(f"sbatch --requeue aggregate_greedy.sh {sub} neurosketch schaefer2018 25")
plt.imshow(_GreedyBestAcc)
_=plt.figure()
for j in range(_GreedyBestAcc.shape[1]):
plt.scatter([j]*_GreedyBestAcc.shape[0],_GreedyBestAcc[:,j],c='g',s=2)
plt.plot(np.arange(_GreedyBestAcc.shape[1]),np.nanmean(_GreedyBestAcc,axis=0))
plt.xticks(numberOfROIs['0112173'])
# plt.ylim([0.19,0.36])
# plt.xlabel("number of ROIs")
# plt.ylabel("accuracy")
_=plt.figure()
for j in range(_GreedyBestAcc.shape[0]):
plt.plot(_GreedyBestAcc[j,:])
_=plt.figure()
plt.imshow(_GreedyBestAcc)
|
archive/Untitled7.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gaussian Process
# #### Books / blog posts
# Gaussian Process Book http://www.gaussianprocess.org/gpml/ <br />
# A Visual Exploration of Gaussian Processes https://distill.pub/2019/visual-exploration-gaussian-processes/ <br />
# <NAME> Ph.D. Thesis on GPyTorch https://geoffpleiss.com/static/media/gpleiss_thesis.d218bc00.pdf
#
# #### Lectures / tutorials
# Probebelistic ML https://www.youtube.com/playlist?list=PL05umP7R6ij1tHaOFY96m5uX3J21a6yNd <br />
# ML Tutorial: Gaussian Processes (<NAME>) https://www.youtube.com/watch?v=92-98SYOdlY
#
# #### Beyond basics
# Sparse Gaussian Processes (https://proceedings.neurips.cc/paper/2005/file/4491777b1aa8b5b32c2e8666dbe1a495-Paper.pdf) <br />
# Gaussian processes for big data (https://arxiv.org/abs/1309.6835) <br />
# Scalable Variational Gaussian Process Classification (http://proceedings.mlr.press/v38/hensman15.pdf) <br />
# Robust Deep Gaussian Processes (https://arxiv.org/abs/1904.02303) <br />
# Parametric Gaussian Process Regressors (https://arxiv.org/pdf/1910.07123.pdf) <br />
# Neural Processes (https://arxiv.org/abs/1807.01622) <br />
import numpy as np
from numpy.random import multivariate_normal
from scipy.linalg import cho_solve, cho_factor
import scipy
import matplotlib.pyplot as plt
np.random.seed(100)
# ### Plot Functions
n = 500 # number of grip-points for plotting
plot_window = (-8, 8) # size of data/plot window
x_grid = np.linspace(-plot_window[0], plot_window[0], 7)[:, np.newaxis]
from pylab import rcParams
rcParams['figure.figsize'] = 15, 5
def plot_data(X,Y,x_grid,y_true):
plt.scatter(X, Y, color='b')
plt.errorbar(X, Y, yerr=noise_scale, fmt='o')
plt.plot(x_grid[:,0], y_true[:,0], color='lightblue')
plt.fill_between(x_grid[:,0], y_true[:,0] - noise_scale / 2, y_true[:,0] + noise_scale / 2, alpha=0.1, color='lightblue')
plt.ylim(-1.5,6.5)
def plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo):
fig, ax = plt.subplots(2,1, gridspec_kw={'height_ratios': [3, 1]})
ax[0].scatter(X, Y, color='b')
ax[0].errorbar(X, Y, yerr=noise_scale, fmt='o')
ax[0].plot(x_grid[:,0], y_true[:,0], color='lightblue')
ax[0].fill_between(x_grid[:,0], y_true[:,0] - noise_scale / 2, y_true[:,0] + noise_scale / 2, alpha=0.1, color='lightblue')
ax[0].set_ylim(-1.5,6.5)
ax[0].plot(x_grid[:,0], mpost[:,0], color="r")
for i in range(spost.shape[1]):
ax[0].plot(x_grid[:,0], spost[:,i], "--", color="r", alpha=0.2)
ax[1].plot(x_grid[:,0], stdpo[:,0], "-", color="k")
# +
def GaussPDFscaled(y, m, s):
return np.exp(-0.5 * (y - m.T) ** 2 / (s ** 2).T)
def plot_prior_posteriori(x_grid, m, s, stdpi, mpost, spost, stdpo):
fig, ax = plt.subplots(1,2)
yy = np.linspace(-5, 10, 200).reshape([200,1])
P = GaussPDFscaled(yy, m[::-1], stdpi)
ax[0].imshow(P, extent=[-8, 8, -5, 10], aspect="auto", origin="lower", cmap="Greys", alpha=0.4)
ax[0].plot(x_grid, phi(x_grid), "-", color="grey")
ax[0].plot(x_grid, s, ":", color="r")
ax[0].plot(x_grid, m, "-", color="r")
ax[0].plot(x_grid, m + 2 * stdpi, "-", color="lightcoral")
ax[0].plot(x_grid, m - 2 * stdpi, "-", color="lightcoral")
ax[0].set(xlim=[-8,8], ylim=[-5,10], title="prior")
Ppost = GaussPDFscaled(yy, mpost[::-1], stdpo)
ax[1].imshow(Ppost, extent=[-8, 8, -5, 10], aspect="auto", origin="lower", cmap="Greys", alpha=0.4)
ax[1].errorbar(X, Y, yerr=noise_scale, fmt='ok')
#ax[1].plot(x_grid, spost, ":", color="r")
ax[1].plot(x_grid, mpost, "-", color="r")
ax[1].plot(x_grid, mpost + 2 * stdpo, "-", color="lightcoral")
ax[1].plot(x_grid, mpost - 2 * stdpo, "-", color="lightcoral")
ax[1].plot(x_grid, mpost + 2 * stdpo + 2 * noise_scale, "-", color="gold")
ax[1].plot(x_grid, mpost - 2 * stdpo - 2 * noise_scale, "-", color="gold")
ax[1].set(xlim=[-8,8], ylim=[-5,10], title="posterior")
# -
# # Background
# ## Gaussian Distributions
# Definition: $p(x)=\frac{1}{\sigma \sqrt[]{2\pi}} e^{-\frac{(x-\mu)^2}{2\sigma^2}} := \mathcal{N}(x;\mu,\,\sigma^{2})$
#
samples = np.random.normal(0,1,10000) # $X \sim \mathcal{N}(\mu,\,\sigma^{2})$
def pdf(x, mean, sigma):
return 1/(sigma*np.sqrt(2*np.pi)) * np.exp(- (x-mean)**2 / (2*sigma**2))
p_x = pdf(x_grid, 0, 1)
plt.plot(x_grid, p_x[:,0], color='b')
plt.hist(samples, 50, density=True, facecolor='lightblue', alpha=0.75)
plt.show()
# products of Gaussian are Gaussians: $\mathcal{N}(x;a,A)\mathcal{N}(x;b,B)=\mathcal{N}(x;c,C)\mathcal{N}(x;a,A) \;\; C:=(A^{-1}+B^{-1})^{-1} \;\; c:= C(A^{-1}a+B^{-1}b)$
# multiply by a constant: $c \mathcal{N}(x;a,A)=\mathcal{N}(x;c a,c A c^T)$
# ## Bayes' Rule
# $p(x|y) = \frac{p(y|x)p(x)}{\int_{}^{} p(x)p(y|x) \,dx}$
# Let $p(x)=\mathcal{N}(x; \mu,\,\sigma^{2})$ and $p(y|x)=\mathcal{N}(y; x, \mathcal{v}^2)$
# Then $p(x|y)=\mathcal{N}(x; m, s^2)$ with $s^2 := \frac{1}{\sigma^{-2} + \mathcal{v}^{-2}}$ and $m := \frac{\sigma^{-2}\mu + \mathcal{v}^{-2}y}{\sigma^{-2} + \mathcal{v}^{-2}}$
X = np.linspace(0, 4, 9)
Y = pdf(X, 2, 0.5)
p_x = pdf(x_grid, 0, 1)
ss = 1/(1+1/0.5**2)
m =(1/0.5**2 * Y).sum()/ 1/(1+1/0.5**2)
p_xy = pdf(x_grid, m, ss)
plt.scatter(X,Y, color='r')
plt.plot(x_grid, p_x, color='b')
plt.plot(x_grid, p_xy, color='purple')
plt.show()
# # Gaussian Process for Regression
# Following the "weight-space view" in the Gaussian Process for ML book [1] and the Probabelistic ML Class of Prof. <NAME> [2].
#
# [1] http://www.gaussianprocess.org/gpml/ <br />
# [2] https://www.youtube.com/playlist?list=PL05umP7R6ij1tHaOFY96m5uX3J21a6yNd <br />
# ## Generate Regression Data
# We generate 12 data points following a multipolynomial function plus gaussian noise.
def data_function(x, noise_scale=0):
true_Y = -0.0005*np.power(x, 4) + -0.008*np.power(x, 3) + 0.03*np.power(x, 2) + 0.7 * np.power(x, 1) + 2
return true_Y + np.random.normal(0, scale=noise_scale, size=true_Y.shape)
data_points = 12
noise_scale = 0.4
data_window = (plot_window[0] +1, plot_window[1] -1)
X = np.random.uniform(*data_window, size=data_points)[:, np.newaxis]
Y = data_function(X, noise_scale)
y_true = data_function(x_grid)
plot_data(X,Y,x_grid,y_true)
# ## Parametric Regression
# $f(x) = w_2*x + w_1 = \phi(x)^T\textbf{w} $ with feature $\phi_x := \phi(x) = [1, x]^T$
def phi(x):
return np.power(x, range(3)) # 1 + x + x^2 + ...
xx_inv = np.linalg.inv(phi(X).T @ phi(X))
xy = phi(X).T @ Y
weights = xx_inv @ xy
y_pred = phi(x_grid) @ weights
weights
plot_data(X,Y,x_grid,y_true)
p = plt.plot(x_grid[:,0], y_pred[:,0])
# ## Gaussian Parametric Regression
# Drawing weights from a weight distribution $p(w) = \mathcal{N}(w; \mu, \Sigma)$
# $p(f) = \mathcal{N}(f; \phi_x^T \mu,\, \phi_x^T \Sigma \phi_x) \;\; \Leftrightarrow \;\; f(x)= \phi_x^T\textbf{w} \;\; \forall \;\; \textbf{w} \sim \mathcal{N}(w; \mu, \Sigma)$
# Gaussian inference on a linear function
# prior: $p(w) = \mathcal{N}(w; \mu, \Sigma)$
# likelihood: $ p(y|\,w,\phi_x) = \mathcal{N}(y; \phi_x^T w, \sigma^2 I) = \mathcal{N}(y; f_x, \sigma^2 I) $
# posterior on $\textbf{w}$: $p(w|\,y,\phi_x) = \frac{p(y|\,w,\phi_x)p(w)}{\int_{}^{} p(y|\,w,\phi_x)p(w) \,dw}$
# posterior on $\textbf{w}$: $p(w|\,y,\phi_x) = \mathcal{N}(w; (\Sigma^{-1} + \sigma^{-1} \phi_x^T \phi_x)^{-1} (\Sigma^{-1} \mu + \sigma^{-1} \phi_X \textbf{y}) , (\Sigma^{-1} + \sigma^{-1} \phi_x^T \phi_x)^{-1} ) $
# $p(f_{x^*}|\,y,\phi_x) = \mathcal{N}(w; \phi_{x^*}(\Sigma^{-1} + \sigma^{-2} \phi_x^T \phi_x)^{-1} (\Sigma^{-1} \mu + \sigma^{-1} \phi_X \textbf{y}) , \phi_{x^*}(\Sigma^{-1} + \sigma^{-1} \phi_x^T \phi_x)^{-1} )\phi_{x^*}^T $
# $p(f_{x^*}|\,y,\phi_x) = \mathcal{N}(w; \phi_{x^*}(A)^{-1} (\Sigma^{-1} \mu + \sigma^{-1} \phi_X \textbf{y}) , \phi_{x^*}(A)^{-1} )\phi_{x^*}^T $ with $A = \Sigma^{-1} + \sigma^{-2} \phi_x^T \phi_x$
# +
F = len(phi(0)) # number of features
mu = np.zeros((F, 1)) # prior mean
Sigma = np.eye(F) # prior Sigma
# prior
m = phi(x_grid) @ mu
kxx = phi(x_grid) @ Sigma @ phi(x_grid).T
s = multivariate_normal(m.flatten(), kxx, size=5).T
stdpi = np.sqrt(np.diag(kxx))[:, np.newaxis]
#posteriori
A = Sigma + 1/(noise_scale**2) * phi(X).T @ phi(X)
mu_weight = 1/(noise_scale**2) * np.linalg.inv(A) @ phi(X).T @ Y
sigma_weight = np.linalg.inv(A)
mpost = phi(x_grid) @ mu_weight
vpost = phi(x_grid) @ sigma_weight @ phi(x_grid).T
spost = multivariate_normal(mpost.flatten(), vpost, size=10).T
stdpo = np.sqrt(np.diag(vpost))[:, np.newaxis]
# -
sigma_weight
plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo)
plot_prior_posteriori(x_grid, m, s, stdpi, mpost, spost, stdpo)
# #### replace $\phi(x)$
def phi(x):
ell = 1.0
return 3 * np.exp(-((x-np.linspace(-8,8,16).T) ** 2) / (ell ** 2) / 2.0)
# +
F = len(phi(0)) # number of features
mu = np.zeros((F, 1)) # prior mean
Sigma = np.eye(F) # prior Sigma
# prior
m = phi(x_grid) @ mu
kxx = phi(x_grid) @ Sigma @ phi(x_grid).T
s = multivariate_normal(m.flatten(), kxx, size=5).T
stdpi = np.sqrt(np.diag(kxx))[:, np.newaxis]
#posteriori
A = Sigma + 1/(noise_scale**2) * phi(X).T @ phi(X)
mu_weight = 1/(noise_scale**2) * np.linalg.inv(A) @ phi(X).T @ Y
sigma_weight = np.linalg.inv(A)
mpost = phi(x_grid) @ mu_weight
vpost = phi(x_grid) @ sigma_weight @ phi(x_grid).T
spost = multivariate_normal(mpost.flatten(), vpost, size=10).T
stdpo = np.sqrt(np.diag(vpost))[:, np.newaxis]
# -
A.shape
plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo)
plot_prior_posteriori(x_grid, m, s, stdpi, mpost, spost, stdpo)
# #### kernel trick
# $p(f_{x^*}|\,y,\phi_x) = \mathcal{N}(w; \phi_{x^*}(\Sigma^{-1} + \sigma^{-2} \phi_x^T \phi_x)^{-1} (\Sigma^{-1} \mu + \sigma^{-1} \phi_X \textbf{y}) , \phi_{x^*}(\Sigma^{-1} + \sigma^{-1} \phi_x^T \phi_x)^{-1} )\phi_{x^*}^T $
# $A = \Sigma^{-1} + \sigma^{-2} \phi_x^T \phi_x$
# $p(f_{x^*}|\,y,\phi_x) = \mathcal{N}(w; \phi_{x^*}(A)^{-1} (\Sigma^{-1} \mu + \sigma^{-1} \phi_X \textbf{y}) , \phi_{x^*}(A)^{-1} )\phi_{x^*}^T $
# +
#posteriori
A = Sigma + 1/(noise_scale**2) * phi(X).T @ phi(X)
mpost = 1/(noise_scale**2) * phi(x_grid) @ np.linalg.inv(A) @ phi(X).T @ Y
vpost = phi(x_grid) @ np.linalg.inv(A) @ phi(x_grid).T
spost = multivariate_normal(mpost.flatten(), vpost, size=10).T
stdpo = np.sqrt(np.diag(vpost))[:, np.newaxis]
# -
print("invert shape:", A.shape)
plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo)
# $K = \phi_x^T \Sigma \phi_x$
# $p(f_{x^*}|\,y,\phi_x) = \mathcal{N}(w; \phi_{x^*}^T \Sigma \phi_{x} (K + \sigma^{-2} I)^{-1} \textbf{y} , \phi_{x^*}^T \Sigma \phi_{x^*} - \phi_{x^*}^T \Sigma (K + \sigma^{-2} I)^{-1} \phi_x^T \Sigma \phi_{x^*} $
# +
#posteriori
K = phi(X) @ Sigma @ phi(X).T
m_pre = phi(x_grid) @ Sigma @ phi(X).T @ np.linalg.inv(K + noise_scale**2 * np.eye(data_points))
mpost = m_pre @ Y
vpost = phi(x_grid) @ Sigma @ phi(x_grid).T - m_pre @ (phi(X) @ Sigma @ phi(x_grid).T)
spost = multivariate_normal(mpost.flatten(), vpost, size=10).T
stdpo = np.sqrt(np.diag(vpost))[:, np.newaxis]
# -
print("invert shape:", (K + noise_scale**2 * np.eye(data_points)).shape)
plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo)
# #### Define a kernel
def kernel(a, b):
return phi(a) @ Sigma @ phi(b).T
# a kxl , b kxh -> c lxh
# +
#posteriori
kXX = kernel(X, X) # K
kxX = kernel(x_grid, X)
kxx = kernel(x_grid, x_grid)
m_pre = kxX @ np.linalg.inv(kXX + noise_scale**2 * np.eye(data_points))
mpost = m_pre @ Y
vpost = kxx - m_pre @ kxX.T
spost = multivariate_normal(mpost.flatten(), vpost, size=10).T
stdpo = np.sqrt(np.diag(vpost))[:, np.newaxis]
# -
plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo)
# ## Gaussian Process for Regression
def mean_function(x):
return x * 0.0 +2
def kernel_builder(f):
return lambda a, b: np.array(
[[ np.float(f(a[i], b[j])) for j in range(b.size)] for i in range(a.size)]
)
X.shape
# +
def SE(a, b, ell, sigma_f=3): # squared exponential kernel
return sigma_f ** 2 * np.exp(-((a-b) ** 2) / (2.0 * ell ** 2))
kernel = kernel_builder(lambda a, b: SE( a, b, 0.5))
# +
# construct implied prior on f_x
mx = mean_function(x_grid)
kxx = kernel(x_grid, x_grid)
s = multivariate_normal(mx.flatten(), kxx, size=5).T
stdpi = np.sqrt(np.diag(kxx))[:, np.newaxis]
mX = mean_function(X)
kXX = kernel(X, X)
kxX = kernel(x_grid, X)
m_pre = kxX @ np.linalg.inv(kXX + noise_scale ** 2 * np.eye(data_points))
mpost = mx + m_pre @ (Y - mX)
vpost = kxx - m_pre @ kxX.T
spost = multivariate_normal(mpost.flatten(), vpost, size=5).T
stdpo = np.sqrt(np.diag(vpost))[:, np.newaxis]
# -
plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo)
plot_prior_posteriori(x_grid, mx, s, stdpi, mpost, spost, stdpo)
# use cholesky decomposition to reduce computation: $O(n^3) \Rightarrow O(n^3 / 3)$
# +
# construct implied prior on f_x
m = mean_function(x_grid)
kxx = kernel(x_grid, x_grid)
s = multivariate_normal(m.flatten(), kxx, size=5).T
stdpi = np.sqrt(np.diag(kxx))[:, np.newaxis]
G = kXX + noise_scale ** 2 * np.eye(data_points)
G = cho_factor(G)
kxX = kernel(x_grid, X)
A = cho_solve(G, kxX.T).T
# posterior P(f_x|Y) = N(f_x, mpost, vpost)
mpost = mean_function(x_grid ) + A @ (Y - mean_function(X))
vpost = kernel(x_grid, x_grid) - A @ kxX.T
spost = multivariate_normal(mpost.flatten(), vpost, size=5).T
stdpo = np.sqrt(np.diag(vpost))[:, np.newaxis]
# -
plot_prediction(X,Y, y_true, x_grid, mpost, spost, stdpo)
plot_prior_posteriori(x_grid, m, s, stdpi, mpost, spost, stdpo)
# ## Model Selection (aka. Hyperparameter Tuning)
#
# Implementation of Eq. 5.9 in _Gaussian Processes for Machine Learning_ (which optimizes Eq. 5.8). The covariance function is defined in Eq. 5.1.
#
# ### Inputs
#
# * ~~Covariance matrix $K$; `kXX` in the notebook~~
# * Kernel function $k$; `kernel` in the notebook
# * Data points $X$; `X` in the notebook
# * Hyperparameters (for the squared exponential kernel) $\theta=\begin{bmatrix}l & \sigma_n & \sigma_f\end{bmatrix}$; corresponding to `[ell, noise_scale, sigma_f]` in the notebook
# * Targets $y$; `Y` in the notebook
|
gaussian-processes/regressor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loss Function
#
# We can create a probability distrbution over the outputs by using a softmax function which is defined as:
#
# $$\frac{e^{x}}{\sum_{x'}e^{x'}}$$
#
# We can then use that probability distribution and knowledge of the correct categories to evaluate the classifier.
#
# A classifier with high confidence in an incorrect category is a poor classifier. A classifier with low confidence in an incorrect category and high confidence in the correct category is a good classifier.
#
# We show below how the softmax function is applied to the output values to turn them into probabilities, and how a measure of the incorrectness, the cross-entropy loss, is computed from them.
# +
import torch
import torch.nn.functional as F
data = torch.Tensor([[6, 2, 1.9]])
weights = torch.Tensor([[1, 0],[0, 1],[0, 1]])
c = torch.mm(data, weights)
print("c1 and c2: " + str(c))
exp = torch.exp(c)
print("e to the power of c: "+str(exp))
soft = exp / torch.sum(exp)
print("e to the power of c normalized: "+str(soft))
result = F.softmax(torch.autograd.Variable(c), dim=1)
print("softmax: " + str(result))
# -
# Convince yourself that the output of the 'softmax' function is the same as 'e to the power of c normalized'.
# +
result = torch.log(result)
print("log(softmax): " + str(result))
# The correct category
target = torch.LongTensor(1)
target[0] = 0
loss = F.nll_loss(result, target)
print("Loss: "+str(loss.data.item()))
# -
# Pytorch also has a ready-made function to compute cross entropy directly from the neural network's outputs.
# Convince yourself that the loss calculated by either method is the same.
loss = F.cross_entropy(c, target)
print("Loss: "+str(loss.data.item()))
# Try this for different values of input data.
#
# You could try them in the following order - in order of increasing difficulty
#
# $\begin{bmatrix}6 & 2 & 1.9\end{bmatrix}$
# $\begin{bmatrix}5 & 2 & 1.9\end{bmatrix}$
# $\begin{bmatrix}4 & 2 & 1.9\end{bmatrix}$
#
# You should see the loss increase as the input data points become more difficult to decide about (as $f_{1}$ and $f_{2} + f_{3}$ get increasingly closer. Remember, we're still using the weight matrix for Toy Problem 2 and the decision that classifier has to take is whether $f_{1} < f_{2} + f_{3}$).
|
exercise_430.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cityscan
# language: python
# name: cityscan
# ---
# # Betweenness Centrality
#
# Betweenness centrality can be calculated for either nodes or edges. It is the sum of the fraction of all-pairs shortest paths that pass through either nodes or edges (depending on which you are running it on). The algorithm we use is from NetworkX, read more here: https://networkx.org/documentation/networkx-1.10/reference/generated/networkx.algorithms.centrality.betweenness_centrality.html
#
# note: betweenness_centrality doesn't work for MultiDiGraph, see: https://github.com/networkx/networkx/issues/4270
#
# also see: https://github.com/networkx/networkx/issues/3432
#
# ## Data Prep
#
# This notebook expects to see an 'admin' folder and a 'data' folder in your working directory. Your admin AOI shapefiles go inside the admin folder. When you call the main function, make sure the city argument is a string tha matches the name of the admin AOI shapefile you want to process (without the '.shp' part). The outputs will be saved in the 'data' folder in a folder with the same city name as well.
import sys
import osmnx as ox
import networkx as nx
import matplotlib.cm as cm
import matplotlib.colors as colors
import pandas as pd
import datetime
import numpy as np
import geocoder
from shapely.ops import unary_union
import geopandas as gpd
from glob import glob
import os
import pylab as pl
import matplotlib.pyplot as plt
import math
ox.config(log_console=True, use_cache=True)
def get_time(t1, t2):
diff = t2 - t1
c = round(diff.total_seconds() / 60, 2)
return c
def get_polygon(string):
admin_folder = './admin'
for file in os.listdir(admin_folder):
if file.endswith(f"{string}.shp"):
print(file)
print('attempt to read file')
# 1. need to transform the input shp to correct prj
boundary_poly = gpd.read_file(admin_folder + '/' + file).set_crs(epsg=4326)
print('print admin_file crs')
print(boundary_poly.crs)
if not boundary_poly.crs == 'epsg:4326':
sys.exit('admin file crs is not 4326, provide an admin file with crs than 4326 and re-run')
#adm = adm.to_crs(epsg=4326)
pol = [i for i in boundary_poly.geometry]
boundary_poly = unary_union(pol)
# if boundary_poly not defined
#else:
#boundary_poly = -1
return boundary_poly
def get_graph(place):
string = place.split(',')[0]
try:
os.mkdir('data/{}'.format(string))
except FileExistsError:
pass
print('Fetching graph data for {}'.format(place))
poly = get_polygon(string)
poly = poly.buffer(0)
#if poly == -1:
#print('poly not defined')
#else:
try:
G = nx.read_gpickle('data/{a}/{a}'.format(a=string))
val = 1
except FileNotFoundError:
print("no pickle file found, retrieving new graph via OSMNX")
G = ox.graph_from_polygon(poly, network_type='drive')
val = 0
# not sure if projecting is needed
# Project a shapely geometry from its current CRS to another.
#If to_crs is None, project to the UTM CRS for the UTM zone in which the
#geometry's centroid lies.
#G = ox.project_graph(G)
print('Writing graph file')
if val != 1:
nx.write_gpickle(G, path='data/{a}/{a}'.format(a=string))
return G
def get_centrality_stats(place):
import numpy as np
string = place.split(',')[0]
try:
edges = gpd.read_file("data/{}/edges.shp".format(string))
if 'edge_centr' in edges.columns:
df = pd.DataFrame()
df['edge_centr'] = edges.edge_centr.astype(float)
df['edge_centr_avg'] = np.nansum(df.edge_centr.values)/len(df.edge_centr)
df.to_csv("data/{a}/Extended_stats_{a}.csv".format(a=string))
except FileNotFoundError:
print("Edges file doesn't exist. Running edge_centrality function.")
G = get_graph(G)
extended_stats = ox.extended_stats(G, bc=True)
dat = pd.DataFrame.from_dict(extended_stats)
dat.to_csv('data/{a}/Extended_Stats_{b}.csv'.format(a=string, b=string))
except Exception as e:
print('Exception Occurred', e)
def get_centrality(place, centrality_type = "both"):
# centrality_type can be either node, edge, or both
t1 = datetime.datetime.now()
string = place.split(',')[0]
# download and project a street network
G = get_graph(place)
#G = ox.graph_from_place('Davao City, Philippines')
#G = ox.graph_from_place(place, which_result=None)
t2 = datetime.datetime.now()
print('{} minutes elapsed!'.format(get_time(t1, t2)))
G = nx.DiGraph(G)
if centrality_type == "node" or centrality_type == "both":
print('Getting node centrality')
node_centrality = nx.betweenness_centrality(G)
t3 = datetime.datetime.now()
print('{} minutes elapsed!'.format(get_time(t1, t3)))
nx.set_node_attributes(G, node_centrality, 'node_centrality')
if centrality_type == "edge" or centrality_type == "both":
print('Getting edge centrality')
# edge closeness centrality: convert graph to a line graph so edges become nodes and vice versa
edge_centrality = nx.edge_betweenness_centrality(G)
t4 = datetime.datetime.now()
print('{} minutes elapsed!'.format(get_time(t1, t4)))
new_edge_centrality = {}
for u,v in edge_centrality:
#new_edge_centrality[(u,v,0)] = edge_centrality[u,v]
new_edge_centrality[(u,v)] = edge_centrality[u,v]
nx.set_edge_attributes(G, new_edge_centrality, 'edge_centrality')
print('Saving output gdf')
print('print string')
print(string)
G = nx.MultiDiGraph(G)
if centrality_type == "node":
ox.save_graph_shapefile(G, filepath='data/{}'.format(string))
elif centrality_type == "edge":
ox.save_graph_shapefile(G, filepath='data/{}'.format(string))
else:
ox.save_graph_shapefile(G, filepath='data/{}'.format(string))
t5 = datetime.datetime.now()
print('{} minutes elapsed!'.format(get_time(t1, t5)))
print('Getting basic stats')
basic_stats = ox.basic_stats(G)
dat = pd.DataFrame.from_dict(basic_stats)
#dat.to_csv('data/{a}/Basic_Stats_{b}.csv'.format(a=string, b=string))
dat.to_csv('data/{a}/Basic_Stats_{b}.csv'.format(a=string, b=string))
t6 = datetime.datetime.now()
print('{} minutes elapsed!'.format(get_time(t1, t6)))
#print('Getting extended stats')
#extended_stats = ox.extended_stats(G, bc=True)
get_centrality_stats(string)
#dat = pd.DataFrame.from_dict(extended_stats)
#dat.to_csv('data/{a}/Extended_Stats_{b}.csv'.format(a=string, b=string))
t7 = datetime.datetime.now()
print('Completed with total time of {} minutes'.format(get_time(t1, t6)))
return
def get_bc_graph_plots(place):
string = place.split(',')[0]
G = nx.read_gpickle("data/{a}/{b}".format(a=string, b=string))
b = ox.basic_stats(G)
#G_projected = ox.project_graph(G)
node_lis = glob('data/{}/nodes.shp'.format(string))
extended_path_lis = glob('data/{}/Extended_*.csv'.format(string))
gdf_node = gpd.GeoDataFrame.from_file(node_lis[0])
exten = pd.read_csv(extended_path_lis[0])
exten= exten.rename(columns={'Unnamed: 0':'osmid'})
exten['betweenness_centrality'] = exten['edge_centr']*100
max_node = exten[exten.betweenness_centrality == max(exten.betweenness_centrality)]['osmid'].values[0]
max_bc = max(exten.betweenness_centrality)
nc = ['r' if node==max_node else '#336699' for node in G.nodes()]
ns = [80 if node==max_node else 8 for node in G.nodes()]
print('{}: The most critical node has {:.2f}% of shortest journeys passing through it. \n'.format(place, max_bc))
print('The road network of {} has {} nodes and {} edges \n\n'.format(string, b['n'], b['m']))
fig, ax = ox.plot_graph(G, node_size=ns, node_color=nc, node_zorder=2, node_alpha=0.8, edge_alpha=0.8,
figsize=(8,8))
gdf_node[gdf_node.osmid == max_node].plot(ax=ax, color='red', zorder = 3)
#ax.set_title('{}: {:.2f}% of shortest paths between all nodes \n in the network through this node'.format(string, max_bc), fontsize=15)
print('\n\n\n')
fig.savefig('data/{}/{}_bc_graph_plot.png'.format(string, string), dpi=300)
return
def get_network_plots(city):
string = city.split(',')[0]
G = get_graph(string)
fig, ax = ox.plot_graph(G, bgcolor = '#ffffff', node_color = '#336699', node_zorder = 2, node_size = 5)
fig.savefig('data/{}/{}_network_plot.png'.format(string, string), dpi=300)
return
def plot_radar(city):
string = city.split(',')[0]
print('print string')
print(string)
#G = ox.graph_from_place(city, network_type='drive')
G = get_graph(city)
try:
if G.graph['crs'].is_projected:
raise Exception("Graph seems to be projected, bearings will not generated if x and y are not in decimal degrees")
except:
print("graph seems to be unprojected, this is ok, continue")
G = ox.add_edge_bearings(G)
# return G
# for debugging: gn.example_edge(G)
#export edges with bearings
#ox.io.save_graph_shapefile(G, filepath='data/{a}/shp_w_bearings'.format(a=string), encoding='utf-8')
bearings = pd.Series([data.get('bearing', np.nan) for u, v, k, data in G.edges(keys=True, data=True)])
# u = from node
# v = to node
# k = index for multiple edges between 2 nodes (0-indexed)
# bearing = angle
# save bearings as csv
bearings.to_csv('data/{a}/{a}_bearings.csv'.format(a=string))
fig = plt.figure() # an empty figure with no axes
ax = fig.add_subplot(1, 1, 1, projection='polar')
polar_plot(ax, bearings)
plt.show()
fig.savefig('data/{a}/{a}_radar_plot.png'.format(a=string), dpi=300)
plt.close()
return
def get_crp_cities_stats(cities):
"""
Input:
cities: list of cities
Output:
CSV file with CRP stats for cities provided as input
"""
dat = pd.DataFrame(columns = ['Number of Intersections', 'Number of Roads', 'Total Length of network (in km)',
'Maximum Betweenness Centrality', 'Average Betweenness Centrality'])
num_intersect, num_roads, tot_len, max_bw, avg_bw, city = [], [], [], [], [], []
for i in cities:
bas = pd.read_csv("data/{a}/Basic_Stats_{a}.csv".format(a=i))
ext = pd.read_csv("data/{a}/Extended_stats_{a}.csv".format(a=i))
num_intersect.append(bas['n'].unique()[0])
num_roads.append(bas['m'].unique()[0])
tot_len.append(float(bas.edge_length_total.unique()[0]) / 1000)
max_bw.append(ext.betweenness_centrality.max())
avg_bw.append(ext.betweenness_centrality_avg.iloc[0])
city.append(i)
dat['Number of Intersections'] = num_intersect
dat['Number of Roads'] = num_roads
dat['Total Length of network (in km)'] = tot_len
dat['Maximum Betweenness Centrality'] = max_bw
dat['Average Betweenness Centrality'] = avg_bw
dat.index = city
dat.to_csv('data/CRP_Stats.csv')
def count_and_merge(n, bearings):
# make twice as many bins as desired, then merge them in pairs
# prevents bin-edge effects around common values like 0° and 90°
n = n * 2
bins = np.arange(n + 1) * 360 / n
count, _ = np.histogram(bearings, bins=bins)
# move the last bin to the front, so that 0° to 5° and 355° to 360° will be binned together
count = np.roll(count, 1)
return count[::2] + count[1::2]
# function to draw a polar histogram for a set of edge bearings
# This function was adopted from an OSMNX example: https://github.com/gboeing/osmnx-examples/blob/9583920c08558662ef77d99529f13df757015172/notebooks/17-street-network-orientations.ipynb
def polar_plot(ax, bearings, n=36, title=''):
bins = np.arange(n + 1) * 360 / n
count = count_and_merge(n, bearings)
_, division = np.histogram(bearings, bins=bins)
frequency = count / count.sum()
division = division[0:-1]
width = 2 * np.pi / n
#print('print division')
#print(division)
ax.set_theta_zero_location('N')
ax.set_theta_direction('clockwise')
x = division * np.pi / 180
bars = ax.bar(x, height=frequency, width=width, align='center', bottom=0, zorder=2,
color='#003366', edgecolor='k', linewidth=0.5, alpha=0.7)
ax.set_ylim(top=frequency.max())
title_font = {'family':'DejaVu Sans', 'size':24, 'weight':'bold'}
xtick_font = {'family':'DejaVu Sans', 'size':10, 'weight':'bold', 'alpha':1.0, 'zorder':3}
ytick_font = {'family':'DejaVu Sans', 'size': 9, 'weight':'bold', 'alpha':0.2, 'zorder':3}
ax.set_title(title.upper(), y=1.05, fontdict=title_font)
ax.set_yticks(np.linspace(0, max(ax.get_ylim()), 5))
yticklabels = ['{:.2f}'.format(y) for y in ax.get_yticks()]
yticklabels[0] = ''
ax.set_yticklabels(labels=yticklabels, fontdict=ytick_font)
xticklabels = ['N', '', 'E', '', 'S', '', 'W', '']
ax.set_xticks(ax.get_xticks())
ax.set_xticklabels(labels=xticklabels, fontdict=xtick_font)
ax.tick_params(axis='x', which='major', pad=-2)
# +
def main(city, centrality_type="both"):
# calculate either 'node' centrality, 'edge' centrality, or 'both'
get_centrality(city, centrality_type = centrality_type)
# may need to debug the betweeness centrality road network plots
#get_bc_graph_plots(city)
# plot the road network
get_network_plots(city)
# generate the road bearing polar plots
plot_radar(city)
if __name__ == "__main__":
if len(sys.argv) == 2:
main(sys.argv[1])
# -
import time
start = time.time()
AOI_name = 'Nis_AOI'
main(AOI_name, centrality_type = "edge")
end = time.time()
time_elapsed = end - start
time_elapsed
# ## Example of calling get_network_plus_building_footprints_plot
def get_network_plus_building_footprints_plot(place, zoom = 1, network_type='drive', bldg_color='orange', dpi=90,
default_width=1, street_widths=None):
# https://github.com/gboeing/osmnx-examples/blob/master/notebooks/10-building-footprints.ipynb
# notes: The preview in Jupyter Notebook is only showing the roads, however the saved PNG file shows both the roads
# and the building footprints
string = place.split(',')[0]
admin_folder = './admin'
fp = f'data/{string}/{string}_network_plus_building_footprints.png'
# Find centroid
poly = gpd.read_file(admin_folder + '/' + string + '.shp').set_crs(epsg = 4326).geometry.buffer(0)
centroid = poly.centroid
x = centroid[0].coords.xy[0][0]
y = centroid[0].coords.xy[1][0]
# Find UTM
utm_zone = math.floor((x + 180) / 6) + 1
utm_crs = f"+proj=utm +zone={utm_zone} +ellps=WGS84 +datum=WGS84 +units=m +no_defs"
# Find polygon length to set dist
poly = poly.to_crs(crs = utm_crs)
length = poly.length[0] / 2 / zoom
print('distance:', length)
gdf = ox.geometries.geometries_from_point((y, x), {'building':True}, dist = length)
# figsize is in inches and can be adjusted to increase the size of the figure
fig, ax = ox.plot_figure_ground(point=(y, x), figsize=(14, 14), dist=length,
network_type=network_type,
default_width=default_width,
street_widths=street_widths,
save=False, show=True, close=True)
fig, ax = ox.plot_footprints(gdf, ax=ax, filepath=fp, color=bldg_color, dpi=dpi,
save=True, show=True, close=True)
# cover page
get_network_plus_building_footprints_plot(AOI_name,
zoom = 4,
network_type = 'drive',
bldg_color = 'orange',
dpi = 90,
default_width = 1,
street_widths = {'secondary': 1, 'primary': 1})
|
road_network_analysis/CityScan_Betweenness_Centrality_and_Bearings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from tkinter.filedialog import askopenfilename
import pandas as pd
import tkinter
import os
import numpy as np
tk_window = tkinter.Tk()
cwd = os.getcwd()
tr_data = pd.read_csv(askopenfilename(title='Choose your training data'))
tr_ans = pd.read_csv(askopenfilename(title='Choose your training answer'))
tr_ans = tr_ans.iloc[:, 0]
ts_data = pd.read_csv(askopenfilename(title='Choose your test data'))
tk_window.destroy()
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
# wine
model = GradientBoostingClassifier(n_estimators = 1050, learning_rate=0.11, max_depth = 3, random_state=0)
model = model.fit(tr_data, tr_ans) # iris 불필요
y_pred = model.predict(ts_data) # iris 불필요
pred_df = pd.DataFrame(y_pred)
pred_df.to_csv("output.csv", mode='w')
# +
def accuracy(real, predict):
return sum(real == predict) / float(real.shape[0])
tk_window = tkinter.Tk()
cwd = os.getcwd()
ts_ans = pd.read_csv(askopenfilename(title='Choose your test answer'))
ts_ans = ts_ans.iloc[:, 0]
tk_window.destroy()
print(accuracy(ts_ans, y_pred))
# -
|
code/wine_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3
# language: python
# name: py3
# ---
# # Basic usage of `pdpipe`
# So how does using `pdpipe` looks like? Let's first import `pandas` and `pdpipe`, an intialize a nice little dataframe:
#
import pandas as pd
raw_df = pd.DataFrame(
data=[
[42, 23, 'Jo', 'M', True, False, 0.07, 'USA', 'Living life to its fullest'],
[81, 23, 'Dana', 'F', True, True, 0.3, 'USA', 'the pen is mightier then the sword'],
[11, 25, 'Bo', 'M', False, True, 2.3, 'Greece', 'all for one and one for all'],
[14, 44, 'Derek', 'M', True, True, 1.1, 'Denmark', 'every life is precious'],
[22, 72, 'Regina', 'F', True, False, 7.1, 'Greece', 'all of you get off my porch'],
[48, 50, 'Jim', 'M', False, False, 0.2, 'Germany', 'boy do I love dogs and cats'],
[50, 80, 'Richy', 'M', False, True, 100.2, 'Finland', 'I love Euro bills'],
[80, 80, 'Wealthus', 'F', False, True, 123.2, 'Finland', 'In Finance We Trust'],
],
columns=['Id', 'Age', 'Name', 'Gender', 'Smoking', 'Runs', 'Savings', 'Country', 'Quote'],
)
# This results in the following dataframe:
raw_df
# ## Constructing pipelines
# We can create different pipeline stage object by calling their constructors,
# which can be of course identified by their camel-cased names, such as
# `pdp.ColDrop` for dropping columns and `pdp.Encode` to encode them, etc.
#
# To build a pipeline, we will usually call the `PdPipeline` class constructor,
# and provide it with a list of pipeline stage objects:
# +
import pdpipe as pdp
from pdpipe import df
pipeline = pdp.PdPipeline([
df.set_index('Id'),
pdp.ColDrop('Name'),
df.drop_rows_where['Savings'] > 100,
df['Healthy'] << df['Runs'] & ~df['Smoking'],
pdp.Bin({'Savings': [1]}, drop=False),
pdp.Scale('StandardScaler'),
pdp.TokenizeText('Quote'),
pdp.SnowballStem('EnglishStemmer', columns=['Quote']),
pdp.RemoveStopwords('English', 'Quote'),
pdp.Encode('Gender'),
pdp.OneHotEncode('Country'),
])
# -
# Printing the pipeline object displays it in order.
pipeline
# The numbers presented in square brackets are the indices of the
# corresponding pipeline stages, and they can be used to retrieve either the
# specific pipeline stage objects composing the pipeline, e.g. with
# `pipeline[5]`, or sub-pipelines composed of sub-sequences of
# the pipeline, e.g. with `pipeline[2:6]`.
# ## Applying pipelines
# The pipeline can now be applied to an input dataframe using the `apply`
# method. We will also provide the `verbose` keyword with `True` to have a
# informative prints or the progress of dataframe processing, stage by stage:
res = pipeline.apply(raw_df, verbose=True)
# We will thus get the dataframe below. We can see all numerical columns were scaled, the `Country` column was one-hot-encoded, `Savings` also got a binned version and the textual `Quote` column underwent some word-level manipulations:
res
# ## Fit and transform
# Pipelines are also callable objects themselves, so calling `pipeline(df)` is
# equivalent to calling `pipeline.apply(df)`.
#
# Additionally, pipelines inherently have a fit state. If none of the stages
# composing them is fittable in nature this doesn't make a lot of a difference,
# but many stage have a `fit_transform` vs `transform` logic, like encoders,
# scalers and so forth.
#
# > *The `apply` pipeline method uses either `fit_transform` and `transform`
# in an intelligent and sensible way: If the pipeline is not fitted, calling
# it is equivalent to calling `fit_transform`, while if it is fitted, the
# call is practically a `transform` call.*
#
# Let's say we want to utilize pdpipe's powerful slicing syntax to apply only
# *some* of the pipeline stages to the raw dataframe. We will now use the
# `fit_transform` method of the pipeline itself to force all encompassed pipeline
# stages to fit-transform themselves.
#
# Here, we will use `pipeline[4:7]` to apply the binning, scaling and tokenization stages only:
pipeline[4:7].fit_transform(raw_df)
|
notebooks/basic_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Build and train a simple model
#
# This is a notebook that builds and trains a simple model as an example of how to use Sparana. It uses ReLu activations with a linear final layer, Xavier initialization and the Adam optimizer. The layers, initialization and optimizer are interchangable with some other types in the library. You will need MNIST pickle files to run this.
# +
# Numpy and cupy are imported, because I use them in all of my experiments, you don't need them to run these cells
import numpy as np
import cupy as cp
# This is needed to load the MNIST files for training and testing
import pickle
# These are the Sparana objects needed to build a model
from sparana.model import model
from sparana.layers import full_relu_layer
from sparana.layers import full_linear_layer
# This is the optimizer used to train the model
from sparana.optimizer import adam_optimizer
# Load the data into this object, which will return optimized minibatches, and track how many minibatches/epochs
# batches have been loaded
from sparana.data_loader import loader
# This is not needed to train and test a simple model, but I am going to demonstrate it here too
from sparana.saver import model_saver
# Put your own path in here
path = 'path'
# This initializes the model object, the 2 things that are required are the input size, and a list of layer objects.
mymodel = model(input_size = 784,
# These are the layers input as a list, the final layer size is the number of classes the model will have.
layers = [full_relu_layer(size = 1000),
full_relu_layer(size = 800),
full_relu_layer(size = 400),
full_linear_layer(size = 10)],
# This is set automatically, but I keep it in here as a demonstration
comp_type = 'GPU')
# Initialize the weights here, after this randomly generated matrices are now in the GPU memory
mymodel.initialize_weights('Xavier', bias_constant = 0.1)
# Initialize the Adam optimizer, the associated matrices are now in GPU memory
opt = adam_optimizer(mymodel, 0.0001, epsilon = 0.001)
#Initialize the saver
mysaver = model_saver(mymodel)
# Initialize the loader object and load the MNIST dataset from pickle files using pickle
myloader = loader(pickle.load(open('MNIST_train_images.p', 'rb')),
pickle.load(open('MNIST_train_labels.p', 'rb')),
pickle.load(open('MNIST_test_images.p', 'rb')),
pickle.load(open('MNIST_test_labels.p', 'rb')))
# -
# # Training
#
# This cell just trains, then stores the model on RAM. The saver object can save the model to the hard disk with the line:
# ```python
# mysaver.pickle_model('filename.p')
# ```
# Demonstrated in the Demo-Lobotomizer notebook where I have a practical reason to do that.
# +
for i in range(20000):
images, labels = myloader.minibatch(250)
opt.train_step(images, labels)
print(mymodel.get_accuracy(myloader.test_data(), myloader.test_labels()))
mysaver.store_model()
# -
# # Outputs
#
# Just a couple of lines here displaying outputs of the model class. I am looking at 10 datapoints from the training set. The outputs will look the same as test set outputs, and will not clutter up your screen.
#
# Quick not here, I am using a linear final layer here, so all of the output values should be either very close to 1 or very close to 0, they do not sum to 1 like they would for a softmax layer.
# +
images, labels = myloader.minibatch(10)
outputs = mymodel.outputs(images)
print('Model outputs')
print(outputs)
# I didn't put this in the library, it's only 1 line, didn't seem necessary
one_hot = np.argmax(outputs, axis = 1)
print('Argmax of model outputs')
print(one_hot)
#We can compare this to the correct classes, the labels
print('Labels')
print(labels)
# I hope these are all correct, won't look good if they are wrong ¯\_(ツ)_/¯
# Or I can just get the accuracy calling get_accuracy from the model object, with the test set
accuracy = mymodel.get_accuracy(myloader.test_data(), myloader.test_labels())
print('Accuracy')
print(accuracy)
# -
|
Demo-Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ' Zipline environment'
# language: python
# name: zipline
# ---
# <img alt="QuantRocket logo" src="https://www.quantrocket.com/assets/img/notebook-header-logo.png">
#
# © Copyright Quantopian Inc.<br>
# © Modifications Copyright QuantRocket LLC<br>
# Licensed under the [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/legalcode).
#
# <a href="https://www.quantrocket.com/disclaimer/">Disclaimer</a>
# + [markdown] deletable=true editable=true
# # Maximum Likelihood Estimates (MLEs)
#
# By <NAME> and <NAME> developed as part of the Masters of Finance curriculum at MIT Sloan.
#
# + [markdown] deletable=true editable=true
# In this tutorial notebook, we'll do the following things:
# 1. Compute the MLE for a normal distribution.
# 2. Compute the MLE for an exponential distribution.
# 3. Fit a normal distribution to asset returns using MLE.
# + [markdown] deletable=true editable=true
# First we need to import some libraries
# + deletable=true editable=true
import math
import matplotlib.pyplot as plt
import numpy as np
import scipy
import scipy.stats
# + [markdown] deletable=true editable=true
# ## Normal Distribution
# We'll start by sampling some data from a normal distribution.
# + deletable=true editable=true jupyter={"outputs_hidden": false}
TRUE_MEAN = 40
TRUE_STD = 10
X = np.random.normal(TRUE_MEAN, TRUE_STD, 1000)
# + [markdown] deletable=true editable=true
# Now we'll define functions that, given our data, will compute the MLE for the $\mu$ and $\sigma$ parameters of the normal distribution.
#
# Recall that
#
# $$\hat\mu = \frac{1}{T}\sum_{t=1}^{T} x_t$$
#
# $$\hat\sigma = \sqrt{\frac{1}{T}\sum_{t=1}^{T}{(x_t - \hat\mu)^2}}$$
# + deletable=true editable=true jupyter={"outputs_hidden": false}
def normal_mu_MLE(X):
# Get the number of observations
T = len(X)
# Sum the observations
s = sum(X)
return 1.0/T * s
def normal_sigma_MLE(X):
T = len(X)
# Get the mu MLE
mu = normal_mu_MLE(X)
# Sum the square of the differences
s = sum( np.power((X - mu), 2) )
# Compute sigma^2
sigma_squared = 1.0/T * s
return math.sqrt(sigma_squared)
# + [markdown] deletable=true editable=true
# Now let's try our functions out on our sample data and see how they compare to the built-in `np.mean` and `np.std`
# + deletable=true editable=true jupyter={"outputs_hidden": false}
print("Mean Estimation")
print(normal_mu_MLE(X))
print(np.mean(X))
print("Standard Deviation Estimation")
print(normal_sigma_MLE(X))
print(np.std(X))
# + [markdown] deletable=true editable=true
# Now let's estimate both parameters at once with scipy's built in `fit()` function.
# + deletable=true editable=true jupyter={"outputs_hidden": false}
mu, std = scipy.stats.norm.fit(X)
print("mu estimate:", str(mu))
print("std estimate:", str(std))
# + [markdown] deletable=true editable=true
# Now let's plot the distribution PDF along with the data to see how well it fits. We can do that by accessing the pdf provided in `scipy.stats.norm.pdf`.
# + deletable=true editable=true jupyter={"outputs_hidden": false}
pdf = scipy.stats.norm.pdf
# We would like to plot our data along an x-axis ranging from 0-80 with 80 intervals
# (increments of 1)
x = np.linspace(0, 80, 80)
plt.hist(X, bins=x, density='true')
plt.plot(pdf(x, loc=mu, scale=std))
plt.xlabel('Value')
plt.ylabel('Observed Frequency')
plt.legend(['Fitted Distribution PDF', 'Observed Data', ]);
# + [markdown] deletable=true editable=true
# ## Exponential Distribution
# Let's do the same thing, but for the exponential distribution. We'll start by sampling some data.
# + deletable=true editable=true
TRUE_LAMBDA = 5
X = np.random.exponential(TRUE_LAMBDA, 1000)
# + [markdown] deletable=true editable=true
# `numpy` defines the exponential distribution as
# $$\frac{1}{\lambda}e^{-\frac{x}{\lambda}}$$
#
# So we need to invert the MLE from the lecture notes. There it is
#
# $$\hat\lambda = \frac{T}{\sum_{t=1}^{T} x_t}$$
#
# Here it's just the reciprocal, so
#
# $$\hat\lambda = \frac{\sum_{t=1}^{T} x_t}{T}$$
# + deletable=true editable=true jupyter={"outputs_hidden": false}
def exp_lamda_MLE(X):
T = len(X)
s = sum(X)
return s/T
# + deletable=true editable=true jupyter={"outputs_hidden": false}
print("lambda estimate:", str(exp_lamda_MLE(X)))
# + deletable=true editable=true jupyter={"outputs_hidden": false}
# The scipy version of the exponential distribution has a location parameter
# that can skew the distribution. We ignore this by fixing the location
# parameter to 0 with floc=0
_, l = scipy.stats.expon.fit(X, floc=0)
# + deletable=true editable=true jupyter={"outputs_hidden": false}
pdf = scipy.stats.expon.pdf
x = range(0, 80)
plt.hist(X, bins=x, density='true')
plt.plot(pdf(x, scale=l))
plt.xlabel('Value')
plt.ylabel('Observed Frequency')
plt.legend(['Fitted Distribution PDF', 'Observed Data', ]);
# + [markdown] deletable=true editable=true
# ## MLE for Asset Returns
#
# Now we'll fetch some real returns and try to fit a normal distribution to them using MLE.
# + deletable=true editable=true jupyter={"outputs_hidden": false}
from quantrocket.master import get_securities
from quantrocket import get_prices
aapl_sid = get_securities(symbols="AAPL", vendors='usstock').index[0]
prices = get_prices('usstock-free-1min', data_frequency='daily', sids=aapl_sid, fields='Close', start_date='2014-01-01', end_date='2015-01-01')
prices = prices.loc['Close'][aapl_sid]
# This will give us the number of dollars returned each day
absolute_returns = np.diff(prices)
# This will give us the percentage return over the last day's value
# the [:-1] notation gives us all but the last item in the array
# We do this because there are no returns on the final price in the array.
returns = absolute_returns/prices[:-1]
# + [markdown] deletable=true editable=true
# Let's use `scipy`'s fit function to get the $\mu$ and $\sigma$ MLEs.
# + deletable=true editable=true jupyter={"outputs_hidden": false}
mu, std = scipy.stats.norm.fit(returns)
pdf = scipy.stats.norm.pdf
x = np.linspace(-1,1, num=100)
h = plt.hist(returns, bins=x, density='true')
l = plt.plot(x, pdf(x, loc=mu, scale=std))
# + [markdown] deletable=true editable=true
# Of course, this fit is meaningless unless we've tested that they obey a normal distribution first. We can test this using the Jarque-Bera normality test. The Jarque-Bera test will reject the hypothesis of a normal distribution if the p-value is under a c.
# + deletable=true editable=true jupyter={"outputs_hidden": false}
from statsmodels.stats.stattools import jarque_bera
jarque_bera(returns)
# + deletable=true editable=true jupyter={"outputs_hidden": false}
jarque_bera(np.random.normal(0, 1, 100))
# -
# ---
#
# **Next Lecture:** [Regression Model Instability](Lecture14-Regression-Model-Instability.ipynb)
#
# [Back to Introduction](Introduction.ipynb)
# ---
#
# *This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian") or QuantRocket LLC ("QuantRocket"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, neither Quantopian nor QuantRocket has taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information believed to be reliable at the time of publication. Neither Quantopian nor QuantRocket makes any guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
|
quant_finance_lectures/Lecture13-Maximum-Likelihood-Estimation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("../../")
import os
os.environ["CUDA_VISIBLE_DEVICES"]="2"
import glob
from pathlib import Path
import numpy as np
from tifffile import imread, imwrite
from skimage.measure import label
from skimage.morphology import remove_small_objects
WholeWingdir = '/home/sancere/Kepler/CurieTrainingDatasets/Dalmiro_Laura/WingCompartments/WholeWing/MaskResults/'
Compartmentdir = '/home/sancere/Kepler/CurieTrainingDatasets/Dalmiro_Laura/WingCompartments/WholeWing/MaskCompartmentResults/'
Savedir = '/home/sancere/Kepler/CurieTrainingDatasets/Dalmiro_Laura/WingCompartments/WholeWing/VeinResults/'
Path(Savedir).mkdir(exist_ok=True)
# +
Raw_path = os.path.join(WholeWingdir, '*tif')
filesRaw = glob.glob(Raw_path)
Comp_path = os.path.join(Compartmentdir, '*tif')
filesComp = glob.glob(Comp_path)
for fname in filesRaw:
Name = os.path.basename(os.path.splitext(fname)[0])
for secondfname in filesComp:
SecondName = os.path.basename(os.path.splitext(secondfname)[0])
if Name == SecondName:
WholeWing = imread(fname)
CompartmentWing = imread(secondfname)
VeinWing = np.ones_like(WholeWing)
indiceszero = np.where(WholeWing > 0) and np.where(CompartmentWing > 0)
VeinWing[indiceszero] = 0
VeinWing = VeinWing * WholeWing
VeinWing = label(VeinWing)
VeinWing = remove_small_objects(VeinWing, 500)
VeinWing = VeinWing > 0
imwrite(Savedir + '/' + Name + '.tif', VeinWing.astype('uint8'))
# -
|
examples/Train/CreateVeinsCompartment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:st]
# language: python
# name: conda-env-st-py
# ---
# +
import scanpy as sc, scanpy
import anndata
import numpy as np , pandas as pd
# TensorFlow and tf.keras
import sklearn
from tqdm import tqdm
import tensorflow as tf , tensorflow
from tensorflow import keras
import anndata
import scipy.io
import re
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import scanpy.api as sc
from keras.utils import to_categorical
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import classification_report, accuracy_score
from sklearn.utils import class_weight
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import RobustScaler, StandardScaler, MinMaxScaler , MaxAbsScaler
#import sklearn.model_selection , sklearn.cross_validation
import os,glob
print(tf.__version__)
from sklearn.utils import resample
import copy
sc.settings.set_figure_params(dpi=200)
sc.settings.verbosity = 3 # verbosity: errors (0), warnings (1), info (2), hints (3)
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#import pandas_ml
#from pandas_ml import ConfusionMatrix
import pickle
from matplotlib import rcParams
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
import diffxpy.api as de
import tensorflow as tf
import matplotlib
#matplotlib.use('PS')
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
#np.random.seed(0)
from scipy import stats
from tensorflow.keras.layers import Dense, Input, Concatenate
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adadelta , Adam
from tensorflow.keras import backend as K
# +
result_prefix = 'nasal/' #'internal_human_lung/' , 'gut/' , 'lung_european/', 'internal_fibroblast_only/' , 'nasal/'
sc.settings.figdir = result_prefix
gene1='ACE2'
gene2='TMPRSS2'
# -
def set_fig_params() :
rcParams.update(matplotlib.rcParamsDefault)
rcParams['pdf.fonttype'] = 42
sns.set(context = 'paper' , style='ticks' ,
rc = {
'figure.autolayout' : True,
'axes.titlesize' : 8 ,
'axes.titleweight' :'bold',
'figure.titleweight' : 'bold' ,
'figure.titlesize' : 8 ,
'axes.labelsize' : 8 ,
'axes.labelpad' : 2 ,
'axes.labelweight' : 'bold' ,
'axes.spines.top' : False,
'axes.spines.right' : False,
'xtick.labelsize' : 7 ,
'ytick.labelsize' : 7 ,
'legend.fontsize' : 7 ,
'figure.figsize' : (3.5, 3.5/1.6 ) ,
'xtick.direction' : 'out' ,
'ytick.direction' : 'out' ,
'xtick.major.size' : 2 ,
'ytick.major.size' : 2 ,
'xtick.major.pad' : 2,
'ytick.major.pad' : 2,
#'lines.linewidth' : 1
} )
# ### Patient Specific Features (one hot encode patient id into input)
# ### Add other covariates that people trust as true
#
# +
###################################################################################################################
if result_prefix == 'internal_human_lung/':
if 0 : # DO NOT TOUCH THIS FLAG.
full_adata_annotated = sc.read('all-regev-tsankov-lung-newlabels.h5ad')
barcodes = [barcode.split('-')[0]+"_"+barcode.split('-')[1].split("_")[1] for barcode in full_adata.obs_names]
full_adata.obs_names = barcodes
barcodes_annotated = [barcode.split('-')[0:-1][0] for barcode in full_adata_annotated.obs_names]
full_adata_annotated.obs_names = barcodes_annotated
full_adata.obs['Cell Type'] = full_adata_annotated.obs['pd_celltype']
full_adata.obs['pd_distalproximal'] = full_adata_annotated.obs['pd_distalproximal']
full_adata[full_adata_annotated.obs_names] = full_adata[full_adata_annotated.obs_names].obs
full_adata.write('all-regev-tsankov-lung-newlabels.h5ad')
full_adata = sc.read('tsankov-mitofilter-processed.h5ad')
corona_plus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()>0) & (full_adata[:,gene2].X.todense() >0)))
corona_minus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()==0)&(full_adata[:,gene2].X.todense() ==0)))
full_adata.obs['corona_plus']= corona_plus
full_adata.obs['corona_minus']= corona_minus
#full_adata.X = np.asarray(full_adata.X.todense())
raw_adata = copy.deepcopy(full_adata )
raw_adata.X = full_adata.layers['counts']
sc.pp.filter_cells(raw_adata, min_genes=200)
sc.pp.filter_genes(raw_adata, min_cells=3)
v2_batches = ['10x_HU28_180121', '10x_HU30_180121', '10x_HU32_180121',
'10x_HU37_180325', '10x_HU39_180325', '10x_HU40_180901',
'10x_HU48_180901', '10x_HU49_180901']
v3_batches = ['10x_HU52_190405', '10x_HU62_190901']
sc.pp.normalize_total(raw_adata, target_sum=1e4)
sc.pp.log1p(raw_adata)
sc.pp.highly_variable_genes(raw_adata, n_top_genes=20000 )
raw_adata.X = np.asarray(raw_adata.X.todense())
#raw_adata = raw_adata[((raw_adata.obs.batch.isin(v2_batches) & (raw_adata.obs.percent_mito < 0.10)) |
# (raw_adata.obs.batch.isin(v3_batches) & (raw_adata.obs.percent_mito < 0.20)) )]
raw_adata = raw_adata[:, raw_adata.var.highly_variable]
adata = scanpy.AnnData(X = raw_adata.X,
obs = raw_adata.obs,
var = raw_adata.var,
varm = raw_adata.varm,
obsm = raw_adata.obsm,
uns = raw_adata.uns
)
##For celltype level analysis
celltype_summary = adata.obs[['Cell Type','corona_plus','corona_minus']].groupby('Cell Type').sum().corona_plus
celltype_list = list(celltype_summary[(celltype_summary > 10)].index)
###################################################################################################################
if result_prefix == 'gut/':
full_adata = sc.read(result_prefix+'gut_adata.h5ad')
corona_plus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()>0) & (full_adata[:,gene2].X.todense() >0)))
corona_minus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()==0)&(full_adata[:,gene2].X.todense() ==0)))
full_adata.obs['corona_plus']= corona_plus
full_adata.obs['corona_minus']= corona_minus
full_adata.X = np.asarray(full_adata.X.todense())
full_adata.obs['nUMI'] = np.sum(full_adata.X,1)
raw_adata = copy.deepcopy(full_adata )
sc.pp.filter_cells(raw_adata, min_genes=200)
sc.pp.filter_genes(raw_adata, min_cells=3)
sc.pp.normalize_total(raw_adata, target_sum=1e4)
sc.pp.log1p(raw_adata)
sc.pp.highly_variable_genes(raw_adata, n_top_genes=20000 )##20000
raw_adata = raw_adata[:, raw_adata.var.highly_variable]
adata = scanpy.AnnData(X = raw_adata.X,
obs = raw_adata.obs,
var = raw_adata.var,
varm = raw_adata.varm,
obsm = raw_adata.obsm,
uns = raw_adata.uns
)
celltype_summary = adata.obs[['Cell Type','corona_plus','corona_minus']].groupby('Cell Type').sum().corona_plus
celltype_list = list(celltype_summary[(celltype_summary > 10)].index)
###################################################################################################################
if result_prefix == 'lung_european/':
full_adata = sc.read(result_prefix+'lca-processed.h5ad')
corona_plus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()>0) & (full_adata[:,gene2].X.todense() >0)))
corona_minus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()==0)&(full_adata[:,gene2].X.todense() ==0)))
full_adata.obs['corona_plus']= corona_plus
full_adata.obs['corona_minus']= corona_minus
raw_adata = copy.deepcopy(full_adata )
#adata = raw_adata
raw_adata.X = full_adata.layers['counts']
sc.pp.filter_cells(raw_adata, min_genes=200)
sc.pp.filter_genes(raw_adata, min_cells=3)
sc.pp.normalize_total(raw_adata, target_sum=1e4)
sc.pp.log1p(raw_adata)
sc.pp.highly_variable_genes(raw_adata, n_top_genes=10000 )
raw_adata.X = np.asarray(raw_adata.X.todense())
raw_adata = raw_adata[:, raw_adata.var.highly_variable]
adata = scanpy.AnnData(X = raw_adata.X,
obs = raw_adata.obs,
var = raw_adata.var,
varm = raw_adata.varm,
obsm = raw_adata.obsm,
uns = raw_adata.uns
)
adata.obs['Cell Type'] = adata.obs['ann_level_3']
adata.obs['Cell Type'].replace('Ciliated lineage' , 'Ciliated' ,inplace= True)
celltype_summary = adata.obs[['Cell Type','corona_plus','corona_minus']].groupby('Cell Type').sum().corona_plus
celltype_list = list(celltype_summary[(celltype_summary > 10)].index)
###################################################################################################################
if result_prefix == 'nasal/':
full_adata = sc.read('nawijn_nasal.h5ad')
corona_plus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()>0) & (full_adata[:,gene2].X.todense() >0)))
corona_minus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()==0)&(full_adata[:,gene2].X.todense() ==0)))
full_adata.obs['corona_plus']= corona_plus
full_adata.obs['corona_minus']= corona_minus
full_adata.obs['nUMI'] = np.sum(full_adata.X,1)
raw_adata = copy.deepcopy(full_adata )
#adata = raw_adata
sc.pp.filter_cells(raw_adata, min_genes=200)
sc.pp.filter_genes(raw_adata, min_cells=3)
sc.pp.normalize_total(raw_adata, target_sum=1e4)
sc.pp.log1p(raw_adata)
sc.pp.highly_variable_genes(raw_adata, n_top_genes=20000 )
raw_adata.X = np.asarray(raw_adata.X.todense())
raw_adata = raw_adata[:, raw_adata.var.highly_variable]
adata = scanpy.AnnData(X = raw_adata.X,
obs = raw_adata.obs,
var = raw_adata.var,
varm = raw_adata.varm,
obsm = raw_adata.obsm,
uns = raw_adata.uns
)
adata.obs['Cell Type'] = adata.obs['ann_level_4']
#adata.obs['Cell Type'].replace('Secretory_2' , 'Sec_Ser_Sup' ,inplace= True)
adata.obs['Cell Type'].replace('Goblet-1' , 'Goblet' ,inplace= True)
adata.obs['Cell Type'].replace('Goblet-2' , 'Goblet' ,inplace= True)
celltype_summary = adata.obs[['Cell Type','corona_plus','corona_minus']].groupby('Cell Type').sum().corona_plus
celltype_list = list(celltype_summary[(celltype_summary > 1)].index)
###################################################################################################################
if result_prefix == 'internal_fibroblast_only/':
if 0 : # DO NOT TOUCH THIS FLAG.
full_adata_annotated = sc.read('all-regev-tsankov-lung-newlabels.h5ad')
barcodes = [barcode.split('-')[0]+"_"+barcode.split('-')[1].split("_")[1] for barcode in full_adata.obs_names]
full_adata.obs_names = barcodes
barcodes_annotated = [barcode.split('-')[0:-1][0] for barcode in full_adata_annotated.obs_names]
full_adata_annotated.obs_names = barcodes_annotated
full_adata.obs['Cell Type'] = full_adata_annotated.obs['pd_celltype']
full_adata.obs['pd_distalproximal'] = full_adata_annotated.obs['pd_distalproximal']
full_adata[full_adata_annotated.obs_names] = full_adata[full_adata_annotated.obs_names].obs
full_adata.write('all-regev-tsankov-lung-newlabels.h5ad')
full_adata = sc.read('all-regev-tsankov-lung-newlabels.h5ad')
corona_plus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()>0) & (full_adata[:,gene2].X.todense() >0)))
corona_minus = np.ndarray.flatten(np.asarray((full_adata[:,gene1].X.todense()==0)&(full_adata[:,gene2].X.todense() ==0)))
full_adata.obs['corona_plus']= corona_plus
full_adata.obs['corona_minus']= corona_minus
full_adata.X = np.asarray(full_adata.X.todense())
raw_adata = copy.deepcopy(full_adata )
raw_adata.X = full_adata.layers['counts']
sc.pp.filter_cells(raw_adata, min_genes=200)
sc.pp.filter_genes(raw_adata, min_cells=3)
v2_batches = ['10x_HU28_180121', '10x_HU30_180121', '10x_HU32_180121',
'10x_HU37_180325', '10x_HU39_180325', '10x_HU40_180901',
'10x_HU48_180901', '10x_HU49_180901']
v3_batches = ['10x_HU52_190405', '10x_HU62_190901']
sc.pp.normalize_total(raw_adata, target_sum=1e4)
sc.pp.log1p(raw_adata)
sc.pp.highly_variable_genes(raw_adata, n_top_genes=20000 )
#raw_adata.X = raw_adata.X.todense()
adata = raw_adata[((raw_adata.obs.batch.isin(v2_batches) & (raw_adata.obs.percent_mito < 0.10)) |
(raw_adata.obs.batch.isin(v3_batches) & (raw_adata.obs.percent_mito < 0.20)) )]
adata = adata[:, adata.var.highly_variable]
adata = adata[adata.obs['Cell Type']=='Fibroblast']
# +
def get_deg_edv(label,deg_adata, obs , n_obs = 1000 ,return_genes=100 , log2fc_min =None) :
obs_with_label = deg_adata.obs[obs] == label
subsampled_ref = sc.pp.subsample(deg_adata[~(obs_with_label)] ,n_obs=n_obs , copy= 1)
subsampled_ref.obs[obs] = 'rest'
subsampled_ref = subsampled_ref[:,deg_adata.var.highly_variable]
subsampled_test = sc.pp.subsample(deg_adata[(obs_with_label)] ,n_obs=n_obs , copy= 1)
subsampled_test = subsampled_test[:,deg_adata.var.highly_variable]
deg_concat_adata = subsampled_test.concatenate(subsampled_ref)
sc.tl.rank_genes_groups(deg_concat_adata, groupby = obs, method='wilcoxon' , n_genes= return_genes)
return scanpy.get.rank_genes_groups_df(deg_concat_adata , group=label ,log2fc_min =log2fc_min)
mito_ribo_genes = np.asarray([i for i in adata.var.index.values if ('MT-' in i) |('RPL' in i) |('RPS' in i) ])
# +
if 0 :
deg_df_dict = dict()
deg_genes_list = list()
for celltype in tqdm(celltype_list):
deg_df = get_deg_edv(label = celltype,deg_adata = adata,obs='Cell Type',
n_obs = (adata.obs['Cell Type'] == celltype).sum() ,
return_genes=100);
deg_df_dict[celltype] = deg_df;
deg_genes_list = np.union1d(deg_genes_list, deg_df.names.values);
deg_df_dict[celltype].to_csv(result_prefix+celltype+'_deg.csv')
# +
celltype_genelists_dict = dict()
for celltype in tqdm(celltype_list):
adata_celltype = adata[adata.obs['Cell Type']==celltype]
classifier_adata = adata_celltype[(adata_celltype.obs['corona_plus'] | adata_celltype.obs['corona_minus'] )]
classifier_adata = adata_celltype[(adata_celltype.obs['corona_plus'] | adata_celltype.obs['corona_minus'] )]
classifier_adata.obs['DP_vs_DN'] = 'bla'
classifier_adata.obs.loc[classifier_adata.obs['corona_plus']==1,'DP_vs_DN'] = 'DP'
classifier_adata.obs.loc[classifier_adata.obs['corona_plus']==0,'DP_vs_DN'] = 'DN'
DP_genes_list = get_deg_edv(label = 'DP',deg_adata = classifier_adata,obs='DP_vs_DN' ,
n_obs = (classifier_adata.obs['DP_vs_DN'] == 'DP').sum() ,
return_genes = 10000 , log2fc_min=0).names.values
classifier_adata = classifier_adata[:,
np.setdiff1d(
np.setdiff1d(DP_genes_list,[gene1,gene2]) ,
mito_ribo_genes)] #np.setdiff1d(, deg_genes_list)
#classifier_adata = get_subsampled_adata(classifier_adata) ## Subsamplign because of slack msges
### Run the Classifier for each celltype and save gene list
X = classifier_adata.X
y = classifier_adata.obs['corona_plus']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, stratify=y, random_state=0 )
clf = sklearn.ensemble.RandomForestClassifier(n_estimators=100, criterion='gini',random_state=0 ,
n_jobs = 50, class_weight='balanced_subsample') #max_depth=4 #
clf.fit(X_train, y_train)
print(clf.score(X_test, y_test))
#clf.predict_proba(X_test )
importances = clf.feature_importances_
indices = np.argsort(importances)[::-1]
ordered_importances = importances[indices]
ordered_genes = classifier_adata.var.index[indices]
std = np.std([tree.feature_importances_ for tree in clf.estimators_],
axis=0)
feature_importance_df = pd.DataFrame( data=ordered_importances , columns = {'Feature Importance'},
index = ordered_genes )
feature_importance_df.to_csv(result_prefix+celltype+'_gene_list.csv')
celltype_genelists_dict[celltype] = feature_importance_df
go_top_n_genes =500
signature_genes = list(ordered_genes[:go_top_n_genes])
sc.tl.score_genes(full_adata, signature_genes, score_name='ACE2+TMPRSS2+ signature score')
go_df = sc.queries.enrich(signature_genes, org="hsapiens", gprofiler_kwargs=dict(no_evidences=False))
go_df['neg_log10_p_value'] = -np.log10(go_df['p_value'])
go_df.to_csv(result_prefix+celltype+'go_enrichment.csv')
n_genes = 100
feature_importance_df = pd.DataFrame( data=ordered_importances[:n_genes] , columns = {'Feature Importance'},
index = ordered_genes[0:n_genes] )
set_fig_params()
fig = plt.figure( figsize =(7.2*2 , 7.2*2) , dpi=200)
ax = sns.barplot(x='Feature Importance',y=ordered_genes[:n_genes],data=feature_importance_df ,
edgecolor='black',linewidth = 0.25 , palette='RdPu_r')
ax.grid(False)
plt.title('ACE2+TMPRSS2+ | Top Enriched Genes')
plt.savefig(result_prefix+celltype+'ACE2+TMPRSS2+_top_enriched_genes.pdf')
if 0 :
set_fig_params()
for source in np.unique(go_df['source']) :
fig = plt.figure( figsize =(7.2*2 , 7.2) , dpi=200)
ax = sns.barplot(x='neg_log10_p_value',y='name', data= go_df[go_df['source']==source] ,
edgecolor='black',linewidth = 0.25 , palette='RdPu_r')
plt.xlabel('$-log_{10}(pval)$')
plt.title('ACE2+TMPRSS2+ | Significant '+source+' Terms')
plt.savefig(result_prefix+celltype+'ACE2+TMPRSS2+_top_enriched_goterms_'+source+'.pdf')
if result_prefix == 'internal_human_lung/':
scanpy.set_figure_params(scanpy=True, dpi=80, dpi_save=200, figsize =(7.2 , 7.2) )
sc.pl.umap(full_adata , color = ['ACE2+TMPRSS2+ signature score' , 'Cell Type' ] , size=(7.2,7.2) ,
save=celltype+'_signature_score_celltypes.pdf', show = 0 )
if result_prefix == 'lung_european/':
scanpy.set_figure_params(scanpy=True, dpi=80, dpi_save=200, figsize =(7.2 , 7.2) )
sc.pl.umap(full_adata , color = ['ACE2+TMPRSS2+ signature score' , 'ann_level_2' ] , size=(7.2,7.2) ,
save=celltype+'_signature_score_celltypes.pdf', show =0)
signature_classifier_adata = full_adata[classifier_adata.obs_names]
Double_Positive = signature_classifier_adata[signature_classifier_adata.obs['corona_plus']].obs['ACE2+TMPRSS2+ signature score']
Double_Negative = signature_classifier_adata[signature_classifier_adata.obs['corona_minus']].obs['ACE2+TMPRSS2+ signature score']
ranksum_pval = scipy.stats.ranksums(Double_Positive,Double_Negative)
ks2samp_pval = scipy.stats.ks_2samp(Double_Positive,Double_Negative)
set_fig_params()
fig = plt.figure( figsize =(7.2, 7.2/1.6 ) , dpi=200)
if ((result_prefix == 'internal_human_lung/') | (result_prefix == 'lung_european/') |
(result_prefix == 'internal_fibroblast_only/') ):
sns.kdeplot((Double_Positive) , shade=1 , color=sns.color_palette("RdBu_r", 10)[-1] ,label='ACE2+TMPRSS2+ Cells' )
sns.kdeplot((Double_Negative) , shade=1, color=sns.color_palette("RdBu_r", 10)[0], label='ACE2-TMPRSS2- Cells' )
plt.xlabel('Signature Score')
if ((result_prefix == 'gut/') | (result_prefix == 'nasal/')):
sns.kdeplot(np.log2(Double_Positive) , shade=1 , color=sns.color_palette("RdBu_r", 10)[-1] ,label='ACE2+TMPRSS2+ Cells' )
sns.kdeplot(np.log2(Double_Negative) , shade=1, color=sns.color_palette("RdBu_r", 10)[0], label='ACE2-TMPRSS2- Cells' )
plt.xlabel('$log_{2}(Signature Score)$')
ax.grid(False)
plt.ylabel('Density')
plt.title('Wilcoxon Rank-sum test | Statistic = {:.3f} | p-val = {:.3f} \n \
K-S Two-sample test | Statistic = {:.3f} | p-val = {:.3f}'.format(ranksum_pval[0],ranksum_pval[1],
ks2samp_pval[0],ks2samp_pval[1]))
plt.savefig(result_prefix+celltype+'ACE2+TMPRSS2+_kde_pvals.pdf')
plt.close('all')
#plt.show()
# +
gene_list_combined = []
#top_n_genes=5000
for celltype in celltype_list:
important_gene_indices = (celltype_genelists_dict[celltype] >0).values.flatten()
gene_list = list(celltype_genelists_dict[celltype][important_gene_indices].index.values) #list(celltype_genelists_dict[celltype_list[i]].sort_values('Feature Importance',ascending=0).index[:top_n_genes])
gene_list_combined += gene_list
gene_list_combined = np.asarray(gene_list_combined)
unique_freq= np.unique(gene_list_combined,return_counts=1)
gene_list_combined_df = pd.DataFrame(index = unique_freq[0] , columns = {'freq' })
gene_list_combined_df['freq']=unique_freq[1]
gene_list_combined_df= gene_list_combined_df.sort_values('freq',ascending=0)
#ordered_genes = list(gene_list_combined_df.index)[:top_n_genes]
# -
# ### Combine Results by Running the RF again on the pooled gene list
# ### Distribution Matching - May need to divide by bin size to get proportions
# +
def get_subsampled_adata(classifier_adata) :
dp_numi_dist = classifier_adata[classifier_adata.obs['DP_vs_DN']=='DP'].obs.nUMI.astype('float32').values
dp_numi_hist = np.histogram(dp_numi_dist , bins=100)
#if result_prefix == 'internal_human_lung/' :
# dp_numi_hist = np.histogram(dp_numi_dist ,bins =100)
#if result_prefix == 'gut/' :
# dp_numi_hist = np.histogram(dp_numi_dist ,bins =100)
dn_numi = classifier_adata[classifier_adata.obs['DP_vs_DN']=='DN'].obs.nUMI.astype('float' ,copy = 1)
dn_obs_list = []
bin_min= 0
for bin_size, bin_max in zip(dp_numi_hist[0], dp_numi_hist[1]) :
dn_bin_indices = list(dn_numi[(dn_numi> bin_min) & (dn_numi < bin_max)].index.values)
if dn_bin_indices !=[] :
dn_obs_list = dn_obs_list + list(np.random.choice(dn_bin_indices ,size = bin_size ))
bin_min = bin_max
dp_obs_list = list(classifier_adata[classifier_adata.obs['DP_vs_DN']=='DP'].obs_names)
subsampled_obs_list = dn_obs_list + dp_obs_list
subsampled_classifier_adata = classifier_adata[subsampled_obs_list]
return subsampled_classifier_adata
classifier_adata = adata[adata.obs['Cell Type'].isin(celltype_list)]
classifier_adata = classifier_adata[:,gene_list_combined_df.index]
classifier_adata = classifier_adata[(classifier_adata.obs['corona_plus'] | classifier_adata.obs['corona_minus'] )]
classifier_adata.obs['DP_vs_DN'] = 'bla'
classifier_adata.obs.loc[classifier_adata.obs['corona_plus']==1,'DP_vs_DN'] = 'DP'
classifier_adata.obs.loc[classifier_adata.obs['corona_plus']==0,'DP_vs_DN'] = 'DN'
##################
dp_pre = (classifier_adata[classifier_adata.obs['DP_vs_DN']=='DP'].obs.nUMI)
dn_pre = (classifier_adata[classifier_adata.obs['DP_vs_DN']=='DN'].obs.nUMI)
##################
classifier_adata = get_subsampled_adata(classifier_adata) ## SUBSAMPLING BECAUSE OF SLACK
##################
dp_post= (classifier_adata[classifier_adata.obs['DP_vs_DN']=='DP'].obs.nUMI)
dn_post= (classifier_adata[classifier_adata.obs['DP_vs_DN']=='DN'].obs.nUMI)
##################
# -
# +
set_fig_params()
fig , ax_= plt.subplots( figsize =(3.6, 3.6/1.6) , dpi=200 , nrows = 2 , ncols=1)
sns.kdeplot(dp_pre, label = 'DP' , ax=ax_[0] , alpha = 0.75)
sns.kdeplot(dn_pre, label = 'DN', ax=ax_[0], alpha = 0.75)
sns.kdeplot(dp_post, label = 'DP' , ax=ax_[1], alpha = 0.75)
sns.kdeplot(dn_post, label = 'DN', ax=ax_[1], alpha = 0.75)
#plt.title('nUMI distribution')
#plt.show()
ax.grid(False)
#plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
#plt.xticks([])
ax_[0].set_title('nUMI Distribution | Before Subsampling')
ax_[1].set_title('nUMI Distribution | After Subsampling')
plt.savefig(result_prefix+'nUMI_distributions.pdf')
plt.show()
#plt.close(fig)
# +
X = classifier_adata.X
y = classifier_adata.obs['corona_plus']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, stratify=y, random_state=0 )
clf = RandomForestClassifier(n_estimators=100, criterion='gini',random_state=0 ,
n_jobs = 50 , class_weight='balanced_subsample') #max_depth=4
clf.fit(X_train, y_train)
print(clf.score(X_test, y_test))
#clf.predict_proba(X_test )
importances = clf.feature_importances_
indices = np.argsort(importances)[::-1]
ordered_importances = importances[indices]
ordered_genes = classifier_adata.var.index[indices]
std = np.std([tree.feature_importances_ for tree in clf.estimators_],
axis=0)
pd.DataFrame(ordered_genes).to_csv(result_prefix+'gene_list.csv')
feature_importance_df = pd.DataFrame( data=ordered_importances , columns = {'Feature Importance'},
index = ordered_genes )
# -
# ### Generate and Save Results and Plots
# +
go_top_n_genes =500
signature_genes = list(ordered_genes[:go_top_n_genes])
go_df = sc.queries.enrich(signature_genes, org="hsapiens", gprofiler_kwargs=dict(no_evidences=False))
go_df['neg_log10_p_value'] = -np.log10(go_df['p_value'])
go_df.to_csv(result_prefix+'go_enrichment.csv')
# +
n_genes = 100
feature_importance_df = pd.DataFrame( data=ordered_importances[:n_genes] , columns = {'Feature Importance'},
index = ordered_genes[0:n_genes] )
set_fig_params()
fig = plt.figure( figsize =(7.2*2 , 7.2*2) , dpi=200)
ax = sns.barplot(x='Feature Importance',y=ordered_genes[:n_genes],data=feature_importance_df ,
edgecolor='black',linewidth = 0.25 , palette='RdPu_r')
ax.grid(False)
#plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
#plt.xticks([])
plt.title('ACE2+TMPRSS2+ | Top Enriched Genes')
plt.savefig(result_prefix+'ACE2+TMPRSS2+_top_enriched_genes.pdf')
#plt.show()
#plt.close(fig)
# +
set_fig_params()
for source in np.unique(go_df['source']) :
fig = plt.figure( figsize =(7.2*8 , 7.2*2) , dpi=200)
ax = sns.barplot(x='neg_log10_p_value',y='name', data= go_df[go_df['source']==source] ,
edgecolor='black',linewidth = 0.25 , palette='RdPu_r')
#plt.axvline(-np.log10(0.05) , c = 'k' , linestyle = 'dashed')
#ax.grid(False)
#plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.xlabel('$-log_{10}(pval)$')
plt.title('ACE2+TMPRSS2+ | Significant '+source+' Terms | '+' EDV')
plt.savefig(result_prefix+'ACE2+TMPRSS2+_top_enriched_goterms_'+source+'.pdf')
#plt.show()
# +
sc.tl.score_genes(full_adata, signature_genes, score_name='ACE2+TMPRSS2+ signature score')
if result_prefix == 'internal_human_lung/':
scanpy.set_figure_params(scanpy=True, dpi=80, dpi_save=200, figsize =(7.2 , 7.2) )
sc.pl.umap(full_adata , color = ['ACE2+TMPRSS2+ signature score' , 'Cell Type' ] , size=(7.2,7.2) ,
save='_signature_score_celltypes.pdf' , show =0)
if result_prefix == 'lung_european/':
scanpy.set_figure_params(scanpy=True, dpi=80, dpi_save=200, figsize =(7.2 , 7.2) )
sc.pl.umap(full_adata , color = ['ACE2+TMPRSS2+ signature score' , 'ann_level_2' ] , size=(7.2,7.2) ,
save='_signature_score_celltypes.pdf', show=0)
# +
signature_classifier_adata = full_adata[classifier_adata.obs_names]
Double_Positive = signature_classifier_adata[signature_classifier_adata.obs['corona_plus']].obs['ACE2+TMPRSS2+ signature score']
Double_Negative = signature_classifier_adata[signature_classifier_adata.obs['corona_minus']].obs['ACE2+TMPRSS2+ signature score']
ranksum_pval = scipy.stats.ranksums(Double_Positive,Double_Negative)
ks2samp_pval = scipy.stats.ks_2samp(Double_Positive,Double_Negative)
set_fig_params()
fig = plt.figure( figsize =(2*7.2, 2*7.2/1.6 ) , dpi=200)
if ((result_prefix == 'internal_human_lung/') | (result_prefix == 'lung_european/') |
(result_prefix == 'internal_fibroblast_only/') ):
sns.kdeplot((Double_Positive) , shade=1 , color=sns.color_palette("RdBu_r", 10)[-1] ,label='ACE2+TMPRSS2+ Cells' )
sns.kdeplot((Double_Negative) , shade=1, color=sns.color_palette("RdBu_r", 10)[0], label='ACE2-TMPRSS2- Cells' )
plt.xlabel('Signature Score')
if ((result_prefix == 'gut/') | (result_prefix == 'nasal/')):
sns.kdeplot(np.log2(Double_Positive) , shade=1 , color=sns.color_palette("RdBu_r", 10)[-1] ,label='ACE2+TMPRSS2+ Cells' )
sns.kdeplot(np.log2(Double_Negative) , shade=1, color=sns.color_palette("RdBu_r", 10)[0], label='ACE2-TMPRSS2- Cells' )
plt.xlabel('$log_{2}(Signature Score)$')
ax.grid(False)
plt.ylabel('Density')
plt.title('Wilcoxon Rank-sum test | Statistic = {:.3f} | p-val = {:.3f} \n \
K-S Two-sample test | Statistic = {:.3f} | p-val = {:.3f}'.format(ranksum_pval[0],ranksum_pval[1],
ks2samp_pval[0],ks2samp_pval[1]))
plt.savefig(result_prefix+'ACE2+TMPRSS2+_kde_pvals.pdf')
plt.close('all')
#plt.show()
# -
|
Figure4_tissue_programs/distribution_matched_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# + [markdown] render=true
# # Incremental modeling with decision optimization
#
# This tutorial includes everything you need to set up decision optimization engines, build a mathematical programming model, then incrementally modify it.
# You will learn how to:
# - change coefficients in an expression
# - add terms in an expression
# - modify constraints and variables bounds
# - remove/add constraints
# - play with relaxations
#
#
# Table of contents:
#
# - [Describe the business problem](#Describe-the-business-problem:--Games-Scheduling-in-the-National-Football-League)
# * [How decision optimization (prescriptive analytics) can help](#How--decision-optimization-can-help)
# * [Use decision optimization](#Use-decision-optimization)
# * [Step 1: Set up the prescriptive model](#Step-1:-Set-up-the-prescriptive-model)
# * [Step 2: Modify the model](#Step-2:-Modify-the-model)
# * [Summary](#Summary)
# ****
# + [markdown] render=true
# ## Describe the business problem: Telephone production
#
# A possible descriptive model of the telephone production problem is as follows:
# * Decision variables:
# * Number of desk phones produced (DeskProduction)
# * Number of cellular phones produced (CellProduction)
# Objective: Maximize profit
# * Constraints:
# * The DeskProduction should be greater than or equal to 100.
# * The CellProduction should be greater than or equal to 100.
# * The assembly time for DeskProduction plus the assembly time for CellProduction should not exceed 400 hours.
# * The painting time for DeskProduction plus the painting time for CellProduction should not exceed 490 hours.
#
# This is a type of discrete optimization problem that can be solved by using either **Integer Programming** (IP) or **Constraint Programming** (CP).
#
# > **Integer Programming** is the class of problems defined as the optimization of a linear function, subject to linear constraints over integer variables.
#
# > **Constraint Programming** problems generally have discrete decision variables, but the constraints can be logical, and the arithmetic expressions are not restricted to being linear.
#
# For the purposes of this tutorial, we will illustrate a solution with mathematical programming (MP).
#
# + [markdown] render=true
# ## How decision optimization can help
#
# * Prescriptive analytics (decision optimization) technology recommends actions that are based on desired outcomes. It takes into account specific scenarios, resources, and knowledge of past and current events. With this insight, your organization can make better decisions and have greater control of business outcomes.
#
# * Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
#
# * Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
# <br/>
#
# <u>With prescriptive analytics, you can:</u>
#
# * Automate the complex decisions and trade-offs to better manage your limited resources.
# * Take advantage of a future opportunity or mitigate a future risk.
# * Proactively update recommendations based on changing events.
# * Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
#
#
# -
# ## Use decision optimization
# + [markdown] render=true
# ### Step 1: Set up the prescriptive model
# -
# #### Writing a mathematical model
# Convert the descriptive model into a mathematical model:
# * Use the two decision variables DeskProduction and CellProduction
# * Use the data given in the problem description (remember to convert minutes to hours where appropriate)
# * Write the objective as a mathematical expression
# * Write the constraints as mathematical expressions (use “=”, “<=”, or “>=”, and name the constraints to describe their purpose)
# * Define the domain for the decision variables
#
#
# #### Telephone production: a mathematical model
# To express the last two constraints, we model assembly time and painting time as linear combinations of the two productions, resulting in the following mathematical model:
#
# ```
# maximize: 12 desk_production+20 cell_production
#
# subject to:
# desk_production>=100
# cell_production>=100
# 0.2 desk_production+0.4 cell_production<=400
# 0.5 desk_production+0.4 cell_production<=490
# ```
# +
# first import the Model class from docplex.mp
from docplex.mp.model import Model
# create one model instance, with a name
m = Model(name='telephone_production')
# + [markdown] render=true
# The continuous variable desk represents the production of desk telephones.
# The continuous variable cell represents the production of cell phones.
# -
# by default, all variables in Docplex have a lower bound of 0 and infinite upper bound
desk = m.integer_var(name='desk')
cell = m.integer_var(name='cell')
# +
m.maximize(12 * desk + 20 * cell)
# write constraints
# constraint #1: desk production is greater than 100
m.add_constraint(desk >= 100, "desk")
# constraint #2: cell production is greater than 100
m.add_constraint(cell >= 100, "cell")
# constraint #3: assembly time limit
ct_assembly = m.add_constraint( 0.2 * desk + 0.4 * cell <= 400, "assembly_limit")
# constraint #4: paiting time limit
ct_painting = m.add_constraint( 0.5 * desk + 0.4 * cell <= 490, "painting_limit")
# + [markdown] render=true
# #### Solve with Decision Optimization
#
# Depending on the size of the problem, the solve stage might fail and require the Commercial Edition of CPLEX engines, which is included in the premium environments in Watson Studio.
#
# You will get the best solution found after ***n*** seconds, because of a time limit parameter.
# -
m.print_information()
msol = m.solve()
assert msol is not None, "model can't solve"
m.print_solution()
# + [markdown] render=true
# ### Step 2: Modify the model
# -
# #### Modify constraints and variables bounds
# The model object provides getters to retrieve variables and constraints by name:
# * get_var_by_name
# * get_constraint_by_name
# The variable and constraint objects both provide properties to access the right hand side (rhs) and left hand side (lhs).
# When you modify a rhs or lhs of a variable, you of course need to give a number.
# When you modify a rhs or lhs of a constraint, you can give a number or an expression based on variables.
#
# Imagine that you want to build 2000 cells and 1000 desks maximum.
#
# And you want to increase the production of both of them from 100 to 350
# +
# Access by name
m.get_var_by_name("desk").ub = 2000
# acess via the object
cell.ub = 1000
m.get_constraint_by_name("desk").rhs = 350
m.get_constraint_by_name("cell").rhs = 350
# -
msol = m.solve()
assert msol is not None, "model can't solve"
m.print_solution()
# The production plan has been updated accordingly to these small changes.
# #### Modify expressions
# You now want to introduce a new type of product: the "hybrid" telephone.
hybrid = m.integer_var(name='hybrid')
# You need to:
# - introduce it in the objective
# - introduce it in the existing painting and assembly time constraints
# - add a new constraint for its production to produce at least 350 of them.
m.add_constraint(hybrid >= 350)
;
# The objective will move from
# <code>
# maximize: 12 desk_production+20 cell_production
# </code>
# to
# <code>
# maximize: 12 desk_production+20 cell_production + 10 hybrid_prodction
# </code>
m.get_objective_expr().add_term(hybrid, 10)
;
# The time constraints will be updated from
# <code>
# 0.2 desk_production+0.4 cell_production<=400
# 0.5 desk_production+0.4 cell_production<=490
# </code>
# to
# <code>
# 0.2 desk_production+0.4 cell_production + 0.2 hybrid_production<=400
# 0.5 desk_production+0.4 cell_production + 0.2 hybrid_production<=490
# </code>
# When you add a constraint to a model, its object is returned to you by the method add_constraint.
# If you don't have it, you can access it via its name
m.get_constraint_by_name("assembly_limit").lhs.add_term(hybrid, 0.2)
ct_painting.lhs.add_term(hybrid, 0.2)
;
# You can now compute the new production plan for our 3 products
msol = m.solve()
assert msol is not None, "model can't solve"
m.print_solution()
# Now imagine that you have improved your painting process, the distribution of the coefficients in the painting limits is no longer [0.5, 0.4, 0.2] but [0.1, 0.1, 0.1]
# You can modify the coefficients, variable by variable, with set_coefficient or via a list of (variable, coeff) with set_coefficients
ct_painting.lhs.set_coefficients([(desk, 0.1), (cell, 0.1), (hybrid, 0.1)])
msol = m.solve()
assert msol is not None, "model can't solve"
m.print_solution()
# #### Relaxations
# Now introduce a new constraint: polishing time limit.
# constraint: polishing time limit
ct_polishing = m.add_constraint( 0.6 * desk + 0.6 * cell + 0.3 * hybrid <= 290, "polishing_limit")
msol = m.solve()
if msol is None:
print("model can't solve")
# The model is now infeasible. We need to handle it and dig into the infeasibilities.
# You can now use the Relaxer object. You can control the way it will relax the constraints or you can use one of the various automatic modes:
# - 'all' relaxes all constraints using a MEDIUM priority; this is the default value.
# - 'named' relaxes all constraints with a user name but not the others.
# - 'match' looks for priority names within constraint names; unnamed constraints are not relaxed.
#
# Use the 'match' mode.
# Polishing constraint is mandatory.
# Painting constraint is a nice to have.
# Assembly constraint has low priority.
ct_polishing.name = "high_"+ct_polishing.name
ct_assembly.name = "low_"+ct_assembly.name
ct_painting.name = "medium_"+ct_painting.name
# +
# if a name contains "low", it has priority LOW
# if a ct name contains "medium" it has priority MEDIUM
# same for HIGH
# if a constraint has no name or does not match any, it is not relaxable.
from docplex.mp.relaxer import Relaxer
relaxer = Relaxer(prioritizer='match', verbose=True)
relaxed_sol = relaxer.relax(m)
relaxed_ok = relaxed_sol is not None
assert relaxed_ok, "relaxation failed"
relaxer.print_information()
# -
m.print_solution()
ct_polishing_relax = relaxer.get_relaxation(ct_polishing)
print("* found slack of {0} for polish ct".format(ct_polishing_relax))
ct_polishing.rhs+= ct_polishing_relax
m.solve()
m.report()
m.print_solution()
# ## Summary
#
#
# You have learned how to set up and use the IBM Decision Optimization CPLEX Modeling for Python to formulate a Mathematical Programming model and modify it in various ways.
# + [markdown] render=true
# #### References
# * <a href="https://rawgit.com/IBMDecisionOptimization/docplex-doc/master/docs/index.html" target="_blank" rel="noopener noreferrer">Decision Optimization CPLEX Modeling for Python documentation</a>
# * <a href="https://dataplatform.cloud.ibm.com/docs/content/wsj/getting-started/welcome-main.html" target="_blank" rel="noopener noreferrer">Watson Studio documentation</a>
# -
# <hr>
# Copyright © 2017-2019. This notebook and its source code are released under the terms of the MIT License.
# <div style="background:#F5F7FA; height:110px; padding: 2em; font-size:14px;">
# <span style="font-size:18px;color:#152935;">Love this notebook? </span>
# <span style="font-size:15px;color:#152935;float:right;margin-right:40px;">Don't have an account yet?</span><br>
# <span style="color:#5A6872;">Share it with your colleagues and help them discover the power of Watson Studio!</span>
# <span style="border: 1px solid #3d70b2;padding:8px;float:right;margin-right:40px; color:#3d70b2;"><a href="https://ibm.co/wsnotebooks" target="_blank" style="color: #3d70b2;text-decoration: none;">Sign Up</a></span><br>
# </div>
|
jupyter/Cloud Pak for Data v3.0.x/Incremental modeling with decision optimization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
# ### importamos los datos:
# +
fondo = pd.read_csv( 'Fondo2.txt',decimal=',', header=0, names= ["canal_fondo","T"], delim_whitespace=True )
hg = pd.read_csv( 'hg2.txt',decimal=',', header=0, names= ["canal_hg","T"], delim_whitespace=True )
sustrato = pd.read_csv( 'itosustrato.txt',decimal=',', header=0, names= ["canal_sustrato","T"], delim_whitespace=True )
ZnMnO = pd.read_csv( 'ZnMnO.txt',decimal=',', header=0, names= ["canal_ZnMnO","T"], delim_whitespace=True )
ZnMnO2 = pd.read_csv( 'ZnMnOrecocido.txt',decimal=',', header=0, names= ["canal_ZnMnO2","T"], delim_whitespace=True )
# -
# ### ploteamos para ver la forma de los datos:
fondo.plot('canal_fondo',"T", title= "fondo")
hg.plot('canal_hg',"T", title= "Hg")
sustrato.plot('canal_sustrato',"T", title= "sustrato")
ZnMnO.plot('canal_ZnMnO',"T", title= "ZnMnO")
ZnMnO2.plot('canal_ZnMnO2',"T", title= "ZnMnO2")
# ### Ajustamos gausianas al Hg para luego hacer la calibracion
# -primer pico: [327,332]
#
# -segundo pico: [380,384]
#
# -tercer pico: [419,423]
#
# -cuarto pico: [450,455]
#
# -quinto pico: [522,526]
#
# -sexto pico: [560,564]
#
# +
x1=hg.iloc[27:33,0]
y1=hg.iloc[27:33,1]
mean = sum(x1 * y1) / sum(y1)
sigma = np.sqrt(sum(y1 * (x1 - mean)**2) / sum(y1))
print(type(x1))
def Gauss(x1, a, x0, sigma):
return a * np.exp(-(x1-x0)**2/(2*sigma**2))
popt,pcov = curve_fit(Gauss, x1, y1, p0=[max(y1), mean, sigma])
plt.plot(x1, y1, 'b+:', label='data')
plt.plot(x1, Gauss(x1, *popt), 'r-', label='fit')
plt.legend()
plt.title('Ajuste primer pico de Hg')
plt.xlabel('Canal')
plt.ylabel('Cuentas')
plt.show()
print("el centro del pico 1 es: ",popt[1]," ± ", pcov[1,1])
print(popt)
#==========================================================================================
x2=hg.iloc[79:86,0]
y2=hg.iloc[79:86,1]
mean = sum(x2 * y2) / sum(y2)
sigma = np.sqrt(sum(y2 * (x2 - mean)**2) / sum(y2))
print(type(x2))
def Gauss(x2, a, x0, sigma):
return a * np.exp(-(x2-x0)**2/(2*sigma**2))
popt,pcov = curve_fit(Gauss, x2, y2, p0=[max(y2), mean, sigma])
plt.plot(x2, y2, 'b+:', label='data')
plt.plot(x2, Gauss(x2, *popt), 'r-', label='fit')
plt.legend()
plt.title('Ajuste segundo pico de Hg')
plt.xlabel('Canal')
plt.ylabel('Cuentas')
plt.show()
print("el centro del pico 2 es: ",popt[1]," ± ", pcov[1,1])
print(popt)
#==========================================================================================
x3=hg.iloc[118:125,0]
y3=hg.iloc[118:125,1]
mean = sum(x3 * y3) / sum(y3)
sigma = np.sqrt(sum(y3 * (x3 - mean)**2) / sum(y3))
print(type(x3))
def Gauss(x3, a, x0, sigma):
return a * np.exp(-(x3-x0)**2/(2*sigma**2))
popt,pcov = curve_fit(Gauss, x3, y3, p0=[max(y3), mean, sigma])
plt.plot(x3, y3, 'b+:', label='data')
plt.plot(x3, Gauss(x3, *popt), 'r-', label='fit')
plt.legend()
plt.title('Ajuste tercer pico de Hg')
plt.xlabel('Canal')
plt.ylabel('Cuentas')
plt.show()
print("el centro del pico 3 es: ",popt[1]," ± ", pcov[1,1])
print(popt)
#==========================================================================================
x4=hg.iloc[149:156,0]
y4=hg.iloc[149:156,1]
mean = sum(x4 * y4) / sum(y4)
sigma = np.sqrt(sum(y4 * (x4 - mean)**2) / sum(y4))
print(type(x4))
def Gauss(x4, a, x0, sigma):
return a * np.exp(-(x4-x0)**2/(2*sigma**2))
popt,pcov = curve_fit(Gauss, x4, y4, p0=[max(y4), mean, sigma])
plt.plot(x4, y4, 'b+:', label='data')
plt.plot(x4, Gauss(x4, *popt), 'r-', label='fit')
plt.legend()
plt.title('Ajuste cuarto pico de Hg')
plt.xlabel('Canal')
plt.ylabel('Cuentas')
plt.show()
print("el centro del pico 4 es: ",popt[1]," ± ", pcov[1,1])
print(popt)
#==========================================================================================
x5=hg.iloc[221:228,0]
y5=hg.iloc[221:228,1]
mean = sum(x5 * y5) / sum(y5)
sigma = np.sqrt(sum(y5 * (x5 - mean)**2) / sum(y5))
print(type(x5))
def Gauss(x5, a, x0, sigma):
return a * np.exp(-(x5-x0)**2/(2*sigma**2))
popt,pcov = curve_fit(Gauss, x5, y5, p0=[max(y5), mean, sigma])
plt.plot(x5, y5, 'b+:', label='data')
plt.plot(x5, Gauss(x5, *popt), 'r-', label='fit')
plt.legend()
plt.title('Ajuste quinto pico de Hg')
plt.xlabel('Canal')
plt.ylabel('Cuentas')
plt.show()
print("el centro del pico 5 es: ",popt[1]," ± ", pcov[1,1])
print(popt)
#==========================================================================================
x6=hg.iloc[259:266,0]
y6=hg.iloc[259:266,1]
mean = sum(x6 * y6) / sum(y6)
sigma = np.sqrt(sum(y6 * (x6 - mean)**2) / sum(y6))
print(type(x6))
def Gauss(x6, a, x0, sigma):
return a * np.exp(-(x6-x0)**2/(2*sigma**2))
popt,pcov = curve_fit(Gauss, x6, y6, p0=[max(y6), mean, sigma])
plt.plot(x6, y6, 'b+:', label='data')
plt.plot(x6, Gauss(x6, *popt), 'r-', label='fit')
plt.legend()
plt.title('Ajuste sexto pico de Hg')
plt.xlabel('Canal')
plt.ylabel('Cuentas')
plt.show()
print("el centro del pico 6 es: ",popt[1]," ± ", pcov[1,1])
print(popt)
# -
# ### hacemos el ajuste lineal para la calibracion:
# +
x=np.array([329.62,
382.09,
421.22,
452.45,
562.78])
y=np.array([312.5668,365.0153,404.6568,435.8328,
546.0735])
def lineal(x,m,b):
return m*x+b
popt,pcov= curve_fit(lineal,x,y,p0=[0.01,1.005])
plt.plot(x,y,"b.")
plt.plot(x,lineal(x,*popt),"r-", label="fit",alpha=0.7)
plt.legend()
plt.xlabel("canales")
plt.ylabel("Longitud de onda [nm]")
plt.title("Ajuste longitud de onda vs Canales")
plt.show()
print ("m = ", popt[0], " y la ordenada= ", popt[1])
m=popt[0]
b=popt[1]
# -
# ### Modificamos los datos segun la calibracion:
#
# +
hg['Longitud de onda[nm]']= hg["canal_hg"]*m+b
sustrato['Longitud de onda[nm]']=sustrato["canal_sustrato"]*m+b
ZnMnO['Longitud de onda[nm]']=ZnMnO["canal_ZnMnO"]*m+b
ZnMnO2['Longitud de onda[nm]']= ZnMnO2["canal_ZnMnO2"]*m+b
fondo['Longitud de onda[nm]']=fondo["canal_fondo"]*m + b
# +
hg.plot('Longitud de onda[nm]',"T", title= "Hg")
sustrato.plot('Longitud de onda[nm]',"T", title= "sustrato")
ZnMnO.plot('Longitud de onda[nm]',"T", title= "ZnMnO")
ZnMnO2.plot('Longitud de onda[nm]',"T", title= "ZnMnO2")
fondo.plot('Longitud de onda[nm]',"T", title= "fondo")
# +
# ZnMnO
TaucMott1=pd.DataFrame()
a = (-np.log(np.array(ZnMnO["T"])/np.array(fondo["T"]))*4.1356*10**(-15)*3*10**(17)/np.array(fondo["Longitud de onda[nm]"]))**2
b = 4.1356*10**(-15)*3*10**(17)/np.array(fondo["Longitud de onda[nm]"])
TaucMott1["(A*E)**2"]=a
TaucMott1["E[eV]"]=b
x=TaucMott1.iloc[42:60,1]
y=TaucMott1.iloc[42:60,0]
def lineal(x,m1,b1):
return m1*(x-b1)
popt,pcov= curve_fit(lineal,x,y,p0=[500,3])
plt.plot(TaucMott1.iloc[0:300,1],TaucMott1.iloc[0:300,0],)
plt.plot(x,lineal(x,*popt),"r-", label="fit",alpha=0.7)
plt.legend()
plt.xlabel("E[eV]")
plt.ylabel("(A*E)**2")
plt.title("")
plt.show()
print(popt)
# +
# ZnMnO2
TaucMott2=pd.DataFrame()
a = (-np.log(np.array(ZnMnO2["T"])/np.array(fondo["T"]))*4.1356*10**(-15)*3*10**(17)/np.array(fondo["Longitud de onda[nm]"]))**2
b = 4.1356*10**(-15)*3*10**(17)/np.array(fondo["Longitud de onda[nm]"])
TaucMott2["(A*E)**2"]=a
TaucMott2["E[eV]"]=b
x=TaucMott2.iloc[55:70,1]
y=TaucMott2.iloc[55:70,0]
def lineal(x,m2,b2):
return m2*(x-b2)
popt,pcov= curve_fit(lineal,x,y,p0=[500,3])
plt.plot(TaucMott2.iloc[0:300,1],TaucMott2.iloc[0:300,0],)
plt.plot(x,lineal(x,*popt),"r-", label="fit",alpha=0.7)
plt.legend()
plt.xlabel("E[eV]")
plt.ylabel("(A*E)**2")
plt.title("")
plt.show()
print(popt)
# +
# Sustrato
TaucMott3=pd.DataFrame()
a = (-np.log(np.array(sustrato["T"])/np.array(fondo["T"]))*4.1356*10**(-15)*3*10**(17)/np.array(fondo["Longitud de onda[nm]"]))**2
b = 4.1356*10**(-15)*3*10**(17)/np.array(fondo["Longitud de onda[nm]"])
TaucMott3["(A*E)**2"]=a
TaucMott3["E[eV]"]=b
x=TaucMott3.iloc[8:25,1]
y=TaucMott3.iloc[8:25,0]
def lineal(x,m3,b3):
return m3*(x-b3)
popt,pcov= curve_fit(lineal,x,y,p0=[500,3])
plt.plot(TaucMott3.iloc[8:300,1],TaucMott3.iloc[8:300,0],)
plt.plot(x,lineal(x,*popt),"r-", label="fit",alpha=0.7)
plt.legend()
plt.xlabel("E[eV]")
plt.ylabel("(A*E)**2")
plt.title("")
plt.show()
print(popt)
|
Exp.4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fastai:Python
# language: python
# name: conda-env-fastai-py
# ---
# # Fastai Environment Validation
# This notebook validates your Fastai environment. It assumes you have a default installation of Fastai with all dependencies and datasets, and the environment contains the [fastai.yml](fastai.yml) file. Open this notebook and select the `fastai` kernel to verify your configuration.
# ## Import Packages
# Make sure you can import packages without any errors
from fastai.vision.all import *
# ### Extract Data
# Verify you can extract the sample datasets to your local directories
path = untar_data(URLs.PETS)/'images'
# ### Define a Dataloader
# Define a `fastai` dataloader and specify a validation set
def is_cat(x): return x[0].isupper()
dls = ImageDataLoaders.from_name_func(
path, get_image_files(path), valid_pct=0.2, seed=42,
label_func=is_cat, item_tfms=Resize(224))
# ### Train a Classifier
# Create a fastai learner and run a training (finetuning)job for 1 epoch using the dataloader
learn = cnn_learner(dls, resnet34, metrics=error_rate)
learn.fine_tune(1)
|
custom-environments/fastai/env_validation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 作業
# ### 用 digits (3個數字即可 , dataset.load_digits(n_class=3)) 資料嘗試跑 PCA
#
# +
from sklearn import datasets
from sklearn import decomposition
from sklearn import datasets
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
digits = datasets.load_digits(n_class=3)
X = digits.data
y = digits.target
# -
print(X.shape)
#print(X[0].reshape(8,8))
# +
len(images_and_labels)
# +
#print(images_and_labels[:10])
# -
pca = decomposition.PCA(n_components=64)
# 載入 相關套件 並 執行 PCA 實驗 ...
pca.fit(X)
XX = pca.transform(X)
print(XX.shape)
images_and_labels = list(zip(X, y))
for index, (image, label) in enumerate(images_and_labels[:10]):
plt.subplot(2, 5, index + 1)
plt.axis('off')
plt.imshow(image.reshape(8,8), cmap=plt.cm.gray_r, interpolation='nearest')
plt.title('Training: %i' % label)
images_and_labels = list(zip(XX, y))
for index, (image, label) in enumerate(images_and_labels[:10]):
plt.subplot(2, 5, index + 1)
plt.axis('off')
plt.imshow(image.reshape(8,8), cmap=plt.cm.gray_r, interpolation='nearest')
plt.title('Training: %i' % label)
fig = plt.figure(figsize=(8, 6))
ax = fig.gca(projection='3d')
ax.scatter(XX[:, 0], XX[:, 1], XX[:, 2], c=y, cmap=plt.cm.RdYlBu)
ax.set_xlabel('PC1')
ax.set_ylabel('PC2')
ax.set_zlabel('PC3')
|
Day_059_PCASample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
#sys.path.append('/content/drive/My Drive/APT/newest/TAR-project-master')
sys.path.append('/Users/patrik/PycharmProjects/TAR-project')
# +
import pandas as pd
import numpy as np
import nltk
nltk.download('stopwords')
nltk.download('punkt')
from dataset.preprocessing.preprocessing import word_stem
# +
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sentence_transformers import SentenceTransformer
from query_model.query_engines import BOWQueryEngine, W2VQueryEngine, D2VQueryEngine, BERTQueryEngine
from query_model.transformers.bm25 import BM25Transformer
# +
corpus_path = '/Users/patrik/PycharmProjects/TAR-project/data/corpus.csv'
corpus = pd.read_csv(corpus_path).drop(columns='Unnamed: 0')
corpus.head()
# -
# ## QUERIES
# +
queries = [
'Smoking effect on the severity of covid19',
'Incubation period of covid19',
'Covid19 effect on pregnancy',
'Mortality rate of covid19',
'Comorbidities and coinfections with effect on the severity of covid19',
'Temperature effect on covid19',
'Ways of covid19 transmission',
'Basic reproductive number of covid19'
]
color_list = ['black', 'red', 'blue', 'green']
query_colors = {}
for i in range(len(queries)):
query_colors[queries[i]] = color_list[i % len(color_list)]
def run_queries(query_engine, queries, k=5):
results = query_engine.run_query(queries[0], k)
if len(queries) > 1:
for q in queries[1:]:
results = pd.concat([results, query_engine.run_query(q, k)])
return results
def color_rows(val):
col = colors[val['query']]
return ['color: %s' % col]*len(val)
# -
# ## MODELS
# TODO: train and save tf-idf and bm25
stop_words = set(stopwords.words('english'))
cv = CountVectorizer(stop_words=stop_words)
transformer = TfidfTransformer(smooth_idf=True, use_idf=True)
query_engine = BOWQueryEngine(cv, transformer)
query_engine.fit(corpus)
query_engine.save('/Users/patrik/PycharmProjects/TAR-project/trained_models/', 'tf_idf')
stop_words = set(stopwords.words('english'))
cv = CountVectorizer(stop_words=None)
transformer = BM25Transformer()
query_engine = BOWQueryEngine(cv, transformer)
query_engine.fit(corpus)
query_engine.save('/Users/patrik/PycharmProjects/TAR-project/trained_models/', 'bm25')
tf_idf = BOWQueryEngine.load('/Users/patrik/PycharmProjects/TAR-project/trained_models/tf_idf.dat')
bm25 = BOWQueryEngine.load('/Users/patrik/PycharmProjects/TAR-project/trained_models/bm25.dat')
w2v = W2VQueryEngine.load('/Users/patrik/PycharmProjects/TAR-project/trained_models/w2v.dat')
w2v_tf_idf = W2VQueryEngine.load('/Users/patrik/PycharmProjects/TAR-project/trained_models/w2vtfidf.dat')
d2v = D2VQueryEngine.load('/Users/patrik/PycharmProjects/TAR-project/trained_models/d2v1.dat')
# +
models = [tf_idf, bm25, w2v, w2v_tf_idf, d2v]
model_names = ['tf_idf', 'bm25', 'w2v', 'w2v_tf_idf', 'd2v']
model_colors = {}
for i in range(len(model_names)):
model_colors[model_names[i]] = color_list[i % len(color_list)]
def color_rows(val):
col = model_colors[val['model']]
return ['color: %s' % col]*len(val)
def run_query_on_models(query, models, model_names, k=3):
results = []
for i in range(len(models)):
res = models[i].run_query(query, k)
res['model'] = model_names[i]
results.append(res)
return pd.concat(results).reset_index()[['query', 'text', 'model', 'sim']]
# -
# we had some problems with saving the model that uses BERT,
# so we saved the results for the next 8 queries,
# this is something we should fix in the future
bert_results_path = '/Users/patrik/PycharmProjects/TAR-project/data/roBERTa_results.csv'
bert_results = pd.read_csv(bert_results_path)
bert_results['model'] = 'SBERT'
bert_results = bert_results[['query', 'text', 'model', 'sim']]
# ### 'Smoking effect on the severity of covid19'
smoking_query = 'Smoking effect on the severity of covid19'
smoking_res = run_query_on_models(smoking_query, models, model_names)
smoking_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==smoking_query][:3].style.set_properties(**{'font-size': '9pt'})
# ### 'Incubation period of covid19'
incub_query = 'Incubation period of covid19'
incub_res = run_query_on_models(incub_query, models, model_names)
incub_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==incub_query][:3].style.set_properties(**{'font-size': '9pt'})
# ### 'Covid19 effect on pregnancy'
pregn_query = 'Covid19 effect on pregnancy'
pregn_res = run_query_on_models(pregn_query, models, model_names)
pregn_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==pregn_query][:3].style.set_properties(**{'font-size': '9pt'})
# ### 'Mortality rate of covid19'
mort_query = 'Mortality rate of covid19'
mort_res = run_query_on_models(mort_query, models, model_names)
mort_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==mort_query][:3].style.set_properties(**{'font-size': '9pt'})
# ### 'Comorbidities and coinfections with effect on the severity of covid19'
coco_query = 'Comorbidities and coinfections with effect on the severity of covid19'
coco_res = run_query_on_models(coco_query, models, model_names)
coco_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==coco_query][:3].style.set_properties(**{'font-size': '9pt'})
# ### 'Temperature effect on covid19'
temp_query = 'Temperature effect on covid19'
temp_res = run_query_on_models(temp_query, models, model_names)
temp_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==temp_query][:3].style.set_properties(**{'font-size': '9pt'})
# ### 'Ways of covid19 transmission'
trans_query = 'Ways of covid19 transmission'
trans_res = run_query_on_models(trans_query, models, model_names)
trans_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==trans_query][:3].style.set_properties(**{'font-size': '9pt'})
# ### 'Basic reproductive number of covid19'
repr_query = 'Basic reproductive number of covid19'
repr_res = run_query_on_models(repr_query, models, model_names)
repr_res.style.set_properties(**{'font-size': '9pt'}).apply(color_rows, axis=1)
bert_results[bert_results['query']==repr_query][:3].style.set_properties(**{'font-size': '9pt'})
|
notebooks/demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .venv
# language: python
# name: .venv
# ---
# ## Without coordinates & noop channel
#
# Certain chart types have neither measures nor dimensions on the axes such as treemaps and bubble charts. This is a case when the noop channel comes in handy for grouping and stacking elements in these kinds of charts.
#
# To get to a treemap, we have to detach all dimensions and the measure from the axes and put two of them on the size channel, whereas the other dimension is still on the color channel. Since the same dimensions are used in both cases Vizzu will be able to animate between these states.
# **Note:** The data used in this tutorial is available [here](https://github.com/vizzuhq/ipyvizzu/blob/gh-pages/docs/tutorial/music_data.json). You can read more about the available types of data in the [Adding data](./01_02_adding_data.ipynb) chapter.
# +
from ipyvizzu import Chart, Data, Config
chart = Chart()
data = Data.from_json("./music_data.json")
chart.animate(data)
chart.animate(
Config(
{
"channels": {
"y": {"set": ["Kinds", "Popularity"]},
"x": {"set": "Genres"},
"label": {"attach": "Popularity"},
},
"color": {"set": "Kinds"},
"title": "Treemap",
}
)
)
chart.animate(
Config(
{
"channels": {
"y": {
"set": None,
},
"x": {
"set": None,
},
"size": {
"attach": ["Genres", "Popularity"],
},
}
}
)
)
snapshot1 = chart.store()
# -
# Getting from a treemap to a bubble chart is simply by changing the geometry to circle. This bubble chart is stacked by the Type dimension that is on the size channel - this is why the bubbles are in separate, small groups.
# +
chart.animate(snapshot1)
chart.animate(Config({"title": "Bubble chart - stacked"}))
chart.animate(
Config(
{
"geometry": "circle",
}
)
)
snapshot2 = chart.store()
# -
# In order to show all bubbles as one group, we use the noop (no operations) channel for the Genres dimension. The noop channel enables us to have a dimension on the chart, that doesn’t affect any parameter of the elements, only their count.
# +
chart.animate(snapshot2)
chart.animate(Config({"title": "Bubble chart - grouped - using the noop channel"}))
chart.animate(
Config({"channels": {"size": {"detach": "Genres"}, "noop": {"set": "Genres"}}})
)
# -
# Next chapter: [Color palette & fonts](./01_13_color_palette_fonts.ipynb) ----- Previous chapter: [Filtering & adding new records](./01_11_filter_add_new_records.ipynb) ----- Back to the [Table of contents](../doc.ipynb#tutorial)
|
docs/tutorial/01_12_without_coordinates_noop_channel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.017579, "end_time": "2021-09-12T06:56:00.003645", "exception": false, "start_time": "2021-09-12T06:55:59.986066", "status": "completed"} tags=[]
# # 1. Parameters
# + papermill={"duration": 0.025159, "end_time": "2021-09-12T06:56:00.038277", "exception": false, "start_time": "2021-09-12T06:56:00.013118", "status": "completed"} tags=["parameters"]
cases_dir = 'cases/unset'
metadata_file = 'input/metadata-subsample-pangolin.tsv'
build_tree = False
# + papermill={"duration": 0.016096, "end_time": "2021-09-12T06:56:00.066166", "exception": false, "start_time": "2021-09-12T06:56:00.050070", "status": "completed"} tags=["injected-parameters"]
# Parameters
cases_dir = "cases/case-200"
iterations = 3
number_samples = 200
build_tree = True
# + papermill={"duration": 2.279691, "end_time": "2021-09-12T06:56:02.353202", "exception": false, "start_time": "2021-09-12T06:56:00.073511", "status": "completed"} tags=[]
from pathlib import Path
import imp
fp, pathname, description = imp.find_module('gdi_benchmark', ['../../lib'])
gdi_benchmark = imp.load_module('gdi_benchmark', fp, pathname, description)
cases_dir_path = Path(cases_dir)
case_name = str(cases_dir_path.name)
index_path = cases_dir_path / 'index'
output_api_path = cases_dir_path / 'query-api.tsv'
output_cli_path = cases_dir_path / 'query-cli.tsv'
# + [markdown] papermill={"duration": 0.01007, "end_time": "2021-09-12T06:56:02.380460", "exception": false, "start_time": "2021-09-12T06:56:02.370390", "status": "completed"} tags=[]
# # 2. Benchmark command-line
# + papermill={"duration": 0.022741, "end_time": "2021-09-12T06:56:02.410759", "exception": false, "start_time": "2021-09-12T06:56:02.388018", "status": "completed"} tags=[]
import pandas as pd
import genomics_data_index.api as gdi
def benchmark_cli_index(name: str, index_path: Path, build_tree: bool) -> pd.DataFrame:
benchmark_commands = {
'query hasa': f'gdi --project-dir {index_path} --ncores 1 query "hasa:hgvs_gn:NC_045512.2:S:p.D614G"',
'query isa': f'gdi --project-dir {index_path} --ncores 1 query "isa:Switzerland/100108/2020"',
'query --summary': f'gdi --project-dir {index_path} --ncores 1 query "hasa:hgvs_gn:NC_045512.2:S:p.D614G" --summary',
'query --features-summary': f'gdi --project-dir {index_path} --ncores 1 query --features-summary mutations',
'list samples': f'gdi --project-dir {index_path} --ncores 1 list samples',
}
if build_tree:
benchmark_commands['query isin'] = f'gdi --project-dir {index_path} --ncores 1 query --reference-name NC_045512 "isin_5_substitutions:Switzerland/100108/2020"'
db = gdi.GenomicsDataIndex.connect(index_path)
number_samples = db.count_samples()
number_features_no_unknown = db.count_mutations(reference_genome='NC_045512', include_unknown=False)
number_features_all = db.count_mutations(reference_genome='NC_045512', include_unknown=True)
iterations = 10
benchmarker = gdi_benchmark.QueryBenchmarkHandler()
return benchmarker.benchmark_cli(name=name, kind_commands=benchmark_commands, number_samples=number_samples,
number_features_no_unknown=number_features_no_unknown, number_features_all=number_features_all,
iterations=iterations)
# + papermill={"duration": 173.615845, "end_time": "2021-09-12T06:58:56.035278", "exception": false, "start_time": "2021-09-12T06:56:02.419433", "status": "completed"} tags=[]
cli_df = benchmark_cli_index(name=case_name, index_path=index_path, build_tree=build_tree)
cli_df.head(3)
# + papermill={"duration": 0.022833, "end_time": "2021-09-12T06:58:56.067239", "exception": false, "start_time": "2021-09-12T06:58:56.044406", "status": "completed"} tags=[]
cli_df.to_csv(output_cli_path, sep='\t', index=False)
# + [markdown] papermill={"duration": 0.007218, "end_time": "2021-09-12T06:58:56.085312", "exception": false, "start_time": "2021-09-12T06:58:56.078094", "status": "completed"} tags=[]
# # 3. Test query API
# + [markdown] papermill={"duration": 0.006484, "end_time": "2021-09-12T06:58:56.098705", "exception": false, "start_time": "2021-09-12T06:58:56.092221", "status": "completed"} tags=[]
# ## 3.1. Load (example) metadata
#
# The simulated data is based off of real sample names and a real tree. So I can load up real metadata and attach it to a query (though the mutations and reference genome are all simulated).
# + papermill={"duration": 1.001992, "end_time": "2021-09-12T06:58:57.107384", "exception": false, "start_time": "2021-09-12T06:58:56.105392", "status": "completed"} tags=[]
import pandas as pd
metadata_df = pd.read_csv(metadata_file, sep='\t')
metadata_df.head(2)
# + [markdown] papermill={"duration": 0.00904, "end_time": "2021-09-12T06:58:57.133527", "exception": false, "start_time": "2021-09-12T06:58:57.124487", "status": "completed"} tags=[]
# ## 3.2. Define benchmark cases
# + papermill={"duration": 0.030475, "end_time": "2021-09-12T06:58:57.171276", "exception": false, "start_time": "2021-09-12T06:58:57.140801", "status": "completed"} tags=[]
from typing import List
import genomics_data_index.api as gdi
def benchmark_api_index(name: str, index_path: Path, build_tree: bool) -> pd.DataFrame:
db = gdi.GenomicsDataIndex.connect(index_path)
q_no_join = db.samples_query(reference_name='NC_045512', universe='mutations')
q_join = db.samples_query(reference_name='NC_045512', universe='mutations').join(metadata_df, sample_names_column='strain')
q = q_join.hasa('hgvs_gn:NC_045512.2:S:p.D614G')
r = q_join.hasa('hgvs_gn:NC_045512.2:N:p.R203K')
number_samples = db.count_samples()
number_features_no_unknown = db.count_mutations(reference_genome='NC_045512', include_unknown=False)
number_features_all = db.count_mutations(reference_genome='NC_045512', include_unknown=True)
repeat = 10
benchmark_cases = {
'db.samples_query': lambda: db.samples_query(reference_name='NC_045512', universe='mutations'),
'q.join': lambda: q_no_join.join(metadata_df, sample_names_column='strain'),
'q.features_summary': lambda: q_join.features_summary(),
'q.features_comparison': lambda: q_join.features_comparison(sample_categories='lineage', categories_kind='dataframe', kind='mutations', unit='proportion'),
'q.hasa': lambda: q_join.hasa("hgvs_gn:NC_045512.2:N:p.R203K"),
'q.isa': lambda: q_join.isa("Switzerland/100112/2020"),
'q AND r': lambda: q & r,
'q.toframe': lambda: q_join.toframe(),
'q.summary': lambda: q_join.summary(),
}
if build_tree:
benchmark_cases['q.isin (distance)'] = lambda: q_join.isin("Switzerland/100108/2020", kind='distance', distance=5, units='substitutions')
benchmark_cases['q.isin (mrca)'] = lambda: q_join.isin(["Switzerland/100108/2020", "FR993751"], kind='mrca')
benchmarker = gdi_benchmark.QueryBenchmarkHandler()
return benchmarker.benchmark_api(name=name, kind_functions=benchmark_cases,
number_samples=number_samples,
number_features_no_unknown=number_features_no_unknown,
number_features_all=number_features_all,
repeat=repeat)
# + [markdown] papermill={"duration": 0.009732, "end_time": "2021-09-12T06:58:57.195468", "exception": false, "start_time": "2021-09-12T06:58:57.185736", "status": "completed"} tags=[]
# ## 3.3. Benchmark reads index
# + papermill={"duration": 39.837479, "end_time": "2021-09-12T06:59:37.040000", "exception": false, "start_time": "2021-09-12T06:58:57.202521", "status": "completed"} tags=[]
api_df = benchmark_api_index(name=case_name, index_path=index_path, build_tree=build_tree)
api_df.head(5)
# + papermill={"duration": 0.026524, "end_time": "2021-09-12T06:59:37.077297", "exception": false, "start_time": "2021-09-12T06:59:37.050773", "status": "completed"} tags=[]
api_df.to_csv(output_api_path, sep='\t', index=False)
# + papermill={"duration": 0.008385, "end_time": "2021-09-12T06:59:37.095945", "exception": false, "start_time": "2021-09-12T06:59:37.087560", "status": "completed"} tags=[]
|
evaluations/sars-cov-2/4-query.case-200.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python training
# language: python
# name: python3
# ---
# +
import sys
sys.path.append("../src")
import logging
from pathlib import Path
import numpy as np
import torch
from natsort import natsorted
from torch import nn
from matplotlib import pyplot as plt
from dataset.data_loading import BasicDataset
from dataset.dataset_interface import DatasetInterface
from networks.UNet.unet_model import UNet
import open3d as o3d
from utils.visualization_utils import visualize_depth, to_rgb
from utils.transformation_utils import imgs_to_pcd, rs_ci, unnormalize_depth
import yaml
root_path = Path("../")
dataset_path = root_path / Path("local_resources/images/calibrated_masked_aug")
logging.info(f"using dataset {dataset_path}")
files = DatasetInterface.get_files_by_path(dataset_path)
# +
idx = np.random.randint(len(files), size=1).item()
rs_rgb, rs_depth, zv_rgb, zv_depth, mask = DatasetInterface.load(files[idx])
mask = np.squeeze(mask)
nan = np.logical_or(np.isnan(rs_depth), np.isnan(zv_depth))
mask = np.logical_and(mask, ~nan)
input_rgb = rs_rgb
input_depth = rs_depth
real_rgb = zv_rgb
real_depth = zv_depth
# blackout pixel not in mask
input_depth = np.where(mask, input_depth, np.nan)
real_depth = np.where(mask, real_depth, np.nan)
it_diff = np.abs(input_depth - real_depth)
mean_it_diff = np.nansum(it_diff) / np.sum(mask)
max_it_diff = np.nanmax(it_diff)
threshold = 4
outliers_diff = np.where((it_diff > threshold * mean_it_diff)[..., None], [255, 0, 0], [0] * 3).astype(np.uint8)
outliers_diff = np.where((it_diff > threshold * mean_it_diff), input_depth, np.nan)
logging.info(f"counted outliers {np.count_nonzero(~np.isnan(outliers_diff))}")
threshold = 3
outliers_diff = np.where((it_diff > threshold * mean_it_diff)[..., None], [255, 0, 0], [0] * 3).astype(np.uint8)
outliers_diff = np.where((it_diff > threshold * mean_it_diff), input_depth, np.nan)
logging.info(f"counted outliers {np.count_nonzero(~np.isnan(outliers_diff))}")
input_depth = np.where(it_diff > threshold * mean_it_diff, np.nan, input_depth)
it_diff = (((it_diff - np.nanmin(it_diff)) / (np.nanmax(it_diff) - np.nanmin(it_diff))) * 255).astype(np.uint8)
cleaned_it_diff = np.abs(input_depth - real_depth)
cleaned_it_diff = (((cleaned_it_diff - np.nanmin(cleaned_it_diff)) / (np.nanmax(cleaned_it_diff) - np.nanmin(cleaned_it_diff))) * 255).astype(np.uint8)
logging.info(f"Mean depth Input {np.nanmean(input_depth)}")
logging.info(f"Mean Distance IT {mean_it_diff}")
logging.info(f"Max Distance IT {max_it_diff}")
diff_pcd = imgs_to_pcd(input_rgb, it_diff.astype(np.float32), rs_ci)
input_pcd = imgs_to_pcd(input_rgb, input_depth.astype(np.float32), rs_ci)
output_pcd = imgs_to_pcd(real_rgb, real_depth.astype(np.float32), rs_ci)
input_diff_pcd = imgs_to_pcd(visualize_depth(it_diff), input_depth.astype(np.float32), rs_ci)
cleaned_input_diff_pcd = imgs_to_pcd(visualize_depth(cleaned_it_diff), input_depth.astype(np.float32), rs_ci)
# outlier_diff_pcd = imgs_to_pcd(outliers_diff, input_depth.astype(np.float32), rs_ci)
outlier_diff_pcd = imgs_to_pcd(input_rgb, outliers_diff.astype(np.float32), rs_ci)
# o3d.visualization.draw_geometries([diff_pcd])
# o3d.visualization.draw_geometries([outlier_diff_pcd])
o3d.visualization.draw_geometries([input_pcd, output_pcd])
# o3d.visualization.draw_geometries([output_pcd])
# o3d.visualization.draw_geometries([input_diff_pcd])
# o3d.visualization.draw_geometries([cleaned_input_diff_pcd])
# o3d.visualization.draw_geometries([input_pcd, pred_pcd])
_, axarr = plt.subplots(1, 4, figsize=(30, 10))
axarr[0].title.set_text("Input")
axarr[0].imshow(visualize_depth(input_depth))
axarr[1].title.set_text("Label")
axarr[1].imshow(visualize_depth(real_depth))
# axarr[2].imshow(to_rgb(rs_rgb))
axarr[2].imshow(it_diff > 3 * mean_it_diff)
axarr[3].imshow(it_diff > 5 * mean_it_diff)
|
notebooks/04_evaluate_dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### FRClassifier.ipynb
#
# Copyright (C) ‹ 2019 › ‹ <NAME> - <EMAIL> ›
#
# This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version.
#
# This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details. You should have received a copy of the GNU General Public License along with this program. If not, see http://www.gnu.org/licenses/.
#
# ---
#
# **[AMS - 190910]** Notebook created for **CERN School of Computing, Cluj-Napoca, Sept 2019**<br>
#
# ---
# Keep track of your progress:
#
# - [ ] Exercise 1 (intermediate)
# - [ ] Exercise 2 (advanced)
# - [ ] Exercise 3 (basic)
# - [ ] Exercise 4 (advanced)
#
# ---
# Some standard libraries:
import matplotlib.pyplot as plt
import numpy as np
# This tutorial uses the [pytorch library](https://pytorch.org) to build its CNN, so we need to import a bunch of pytoch libraries:
import torch
import torchvision
import torchvision.transforms as transforms
from torchsummary import summary
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
# We're going to be classifying radio galaxies. So we'll import the custom class [FRDEEP](https://hongmingtang060313.github.io/FR-DEEP/), which provides a pytorch formatted dataset filled with postage stamp images of radio galaxies:
from FRDEEP import FRDEEPF
# Some code to control the size of figures in the notebook:
plt.rcParams['figure.figsize'] = [10, 5]
plt.rcParams['figure.dpi'] = 300
# Our network is defined as a class. We first specify each of the layers individually and then arrange them into a net in the <code>forward</code> function:
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 34 * 34, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
# conv1 output width: input_width - (kernel_size - 1) => 150 - (5-1) = 146
# pool 1 output width: int(input_width/2) => 73
x = self.pool(F.relu(self.conv1(x)))
# conv2 output width: input_width - (kernel_size - 1) => 73 - (5-1) = 69
# pool 2 output width: int(input_width/2) => 34
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 34 * 34)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
# Let's print a summary of what our net looks like:
net = Net()
summary(net,(1,150,150))
# At this point we've defined our net but we don't have any data to feed it yet.
#
# To feed data into a pytorch net we use a <code>DataLoader</code>, which can apply [transforms](https://pytorch.org/docs/stable/torchvision/transforms.html) to the input data on the fly. This can be used for data augmentation, but for now we'll use a couple of very simple transforms: (1) to convert the PIL images in the dataset to tensor data, and (2) to normalise the values of the image pixels to have a mean of 0.5 and a variance of 0.5.
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize([0.5],[0.5])
])
# When we run the <code>DataLoader</code> it will check whether the dataset already exists on disk and if it doesn't then it will download it. We do this twice, once for the training data and once for the test data. The designations train/test are specified within the dataset itself.
#
# Because dataset sizes can be quite large, it's normal to read the data in **batches** of a specified size.
bsize = 8
# If you also require a validation dataset it should be a subset of the **training data**. Never take your validation dataset from your test data.
# +
trainset = FRDEEPF(root='./FIRST_data', train=True, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=bsize, shuffle=True, num_workers=2)
testset = FRDEEPF(root='./FIRST_data', train=False, download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=2, shuffle=True, num_workers=2)
# -
# ---
#
# **Exercise 1:** Can you add in a dataloader for a *validation* dataset? [Hint](https://as595.github.io/hint1/)
#
# ---
# Let's specify the labels for our target classes. We could extract these from the dataset itself, but it's easier just to make a small list:
classes = ('FRI', 'FRII')
# Let's have a quick look to see how our input data appear. The images are in tensor form, so to display them we need a function that converts them back to numpy arrays:
def imshow(img):
# unnormalize
img = img / 2 + 0.5
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
# We'll pull one iteration of our data out of the dataset. The *batch size* is set to 8, so this should return two images and their labels:
# get some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
# show images
imshow(torchvision.utils.make_grid(images)) # NOT plt.imshow
# print labels
print(' '+' '.join('%5s' % classes[labels[j]] for j in range(bsize)))
# At this point we have a net and we have some data. Next we need an objective function to optimize and an optimization method to use. Here we're minimizing the [cross entropy loss](https://ml-cheatsheet.readthedocs.io/en/latest/loss_functions.html) and we'll use the [Adagrad optimizer](https://ml-cheatsheet.readthedocs.io/en/latest/optimizers.html#adagrad).
#
# We need to specify the *learning rate* for the optimizer, which basically controls the size of each update step in this iterative optimizer.
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adagrad(net.parameters(), lr=0.01)
# Now let's put it all together. We'll run for 10 **epochs**. In each epoch the whole training dataset is passed forwards and backwards through the network. The full dataset is a bit big to drop on the algorithm all at once, so we feed it through in **batches**. Ideally the **loss** should get smaller with each epoch.
#
# Why 10 epochs? *Good question...*
#
# Deciding on the right number of epochs is a balancing act. Obviously, the more epochs you use, the longer your code will take to run... but your loss will hopefully keep getting smaller. The more complicated your dataset is, the more epochs you may need to achieve the same kind of loss values. However... if you run too many epochs you run the risk of *over-fitting* your training data. This is why validation is so important. In short: there's no good/simple answer.
# +
nepoch = 10 # number of epochs
print_num = 50 # record the loss every 50 samples
epoch_history = [] # record of epoch stage
train_history = [] # record of loss at each stage
for epoch in range(nepoch): # loop over the dataset multiple times
train_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# update the training loss for this epoch
train_loss += loss.item()
# print statistics
if i % print_num == (print_num-1): # print every 50 mini-batches
epoch_history.append(epoch+float(i/len(trainloader)))
train_history.append(train_loss / print_num)
train_loss = 0.0
print('Finished Training')
# -
plt.plot(epoch_history, train_history, label="Training Loss")
plt.xlabel('No. of Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# ---
#
# **Exercise 2:** Can you amend the code to print out the validation loss after each epoch? [Hint](https://as595.github.io/hint2/)
#
# ---
# Let's see how we did with our 10 epochs. To do this we use our test dataset:
# +
correct = 0
total = 0
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 50 test images: %d %%' % (100 * correct / total))
# -
# However, this is an imbalanced dataset (i.e. there are more FRII galaxies than FRI galaxies) so the overall accuracy may not equally represent how well we were able to classify the different types of galaxy. Let's look at the accuracy *per class* instead.
# +
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(2):
label = labels[i]
class_correct[label] += c[i].item()
class_total[label] += 1
# -
for i in range(len(classes)):
print('Accuracy of %5s : %2d %%' % (classes[i], 100 * class_correct[i] / class_total[i]))
# ---
#
# **Exercise 3:** Can you amend the code to print out the validation accuracy at each training epoch and add it to your plot?
#
# **Exercise 4:** Can you use [torchvision.transforms](https://pytorch.org/docs/stable/torchvision/transforms.html) to augment the original dataset and improve your accuracy?
#
# ---
|
tutorials/FRClassifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#changed options in tkinter
# -
import tkinter as tk
from tkinter import *
import cv2
import csv
import os
import numpy as np
from PIL import Image,ImageTk
import pandas as pd
import datetime
import time
# +
#####Window is our Main frame of system
window = tk.Tk()
window.title("FSMS-Face Recognition Based Security Management System")
window.configure(borderwidth=0)
displaystring=StringVar()
status = Label(window, textvariable=displaystring,font=('times',12),fg='#444444')
status.grid(row=8,columnspan=8,sticky='W',pady=(10,0))
# +
# manually fill log
def manually_fill():
global sb
sb = tk.Tk()
sb.iconbitmap('AMS.ico')
sb.title("Manually Fill Loggg")
#sb.geometry('580x320')
def err_screen_for_subject():
def ec_delete():
ec.destroy()
global ec
ec = tk.Tk()
ec.geometry('300x100')
ec.iconbitmap('AMS.ico')
ec.title('Warning!!')
Label(ec, text='Please enter a valid room number!', font=('times', 16)).pack()
Button(ec, text='OK', command=ec_delete,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1,
font=('times', 15 )).place(x=90, y=50)
def fill_attendance():
ts = time.time()
Date = datetime.datetime.fromtimestamp(ts).strftime('%Y_%m_%d')
timeStamp = datetime.datetime.fromtimestamp(ts).strftime('%H:%M:%S')
Time = datetime.datetime.fromtimestamp(ts).strftime('%H:%M:%S')
Hour, Minute, Second = timeStamp.split(":")
#Creatting csv of attendance
#Create table for Attendance
date_for_DB = datetime.datetime.fromtimestamp(ts).strftime('%Y_%m_%d')
global subb
subb=SUB_ENTRY.get()
DB_table_name = str(subb + "_" + Date + "_Time_" + Hour + "_" + Minute + "_" + Second)
import pymysql.connections ###Connect to the database
try:
global cursor
connection = pymysql.connect(host='localhost', user='root', password='', db='manually_fill_attendance')
cursor = connection.cursor()
except Exception as e:
print(e)
sql = "CREATE TABLE " + DB_table_name + """
(ID INT NOT NULL AUTO_INCREMENT,
ENROLLMENT varchar(100) NOT NULL,
NAME VARCHAR(50) NOT NULL,
DATE VARCHAR(20) NOT NULL,
TIME VARCHAR(20) NOT NULL,
PRIMARY KEY (ID)
);
"""
try:
cursor.execute(sql) ##for create a table
except Exception as ex:
print(ex)
#### message box
if subb == '':
err_screen_for_subject()
else:
sb.destroy()
MFW = tk.Tk()
MFW.iconbitmap('AMS.ico')
MFW.title("Manual log of "+ str(subb))
#MFW.geometry('880x470')
def del_errsc2():
errsc2.destroy()
def err_screen1():
global errsc2
errsc2 = tk.Tk()
errsc2.geometry('330x100')
errsc2.iconbitmap('AMS.ico')
errsc2.title('Warning!!')
Label(errsc2, text='Please enter Name & ID!!!', font=('times', 16)).pack()
Button(errsc2, text='OK', command=del_errsc2, width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times', 15)).place(x=90, y=50)
def testVal(inStr, acttyp):
if acttyp == '1': # insert
if not inStr.isdigit():
return False
return True
ENR = tk.Label(MFW, text="Employee ID :", font=('times', 15))
ENR.grid(row=0,column=0,sticky=('S','W'),padx=10,pady=(10,0))
#ENR.place(x=30, y=100)
STU_NAME = tk.Label(MFW, text="Employee name :",font=('times', 15))
STU_NAME.grid(row=2,column=0,sticky=('S','W'),padx=10,pady=(10,0))
#STU_NAME.place(x=30, y=200)
global ENR_ENTRY
ENR_ENTRY = tk.Entry(MFW,width=20,validate='key', bg="white", font=('times', 23))
ENR_ENTRY['validatecommand'] = (ENR_ENTRY.register(testVal), '%P', '%d')
ENR_ENTRY.grid(row=1,column=0,padx=(10,0),pady=(3,10))
#ENR_ENTRY.place(x=290, y=105)
def remove_enr():
ENR_ENTRY.delete(first=0, last=22)
STUDENT_ENTRY = tk.Entry(MFW, width=20, bg="white", font=('times', 23))
STUDENT_ENTRY.grid(row=3,column=0,padx=(10,0),pady=(3,10))
#place(x=290, y=205)
def remove_student():
STUDENT_ENTRY.delete(first=0, last=22)
####get important variable
def enter_data_DB():
ENROLLMENT = ENR_ENTRY.get()
STUDENT = STUDENT_ENTRY.get()
if ENROLLMENT=='':
err_screen1() #messagebox
elif STUDENT=='':
err_screen1()
else:
time = datetime.datetime.fromtimestamp(ts).strftime('%H:%M:%S')
Hour, Minute, Second = time.split(":")
Insert_data = "INSERT INTO " + DB_table_name + " (ID,ENROLLMENT,NAME,DATE,TIME) VALUES (0, %s, %s, %s,%s)"
VALUES = (str(ENROLLMENT), str(STUDENT), str(Date), str(time))
try:
cursor.execute(Insert_data, VALUES)
except Exception as e:
print(e)
ENR_ENTRY.delete(first=0, last=22)
STUDENT_ENTRY.delete(first=0, last=22)
def create_csv():
import csv
cursor.execute("select * from " + DB_table_name + ";")
csv_name='E:/Attendace managemnt system/Attendance/Manually Attendance/'+DB_table_name+'.csv'
with open(csv_name, "w") as csv_file:
csv_writer = csv.writer(csv_file)
csv_writer.writerow([i[0] for i in cursor.description]) # write headers
csv_writer.writerows(cursor)
O="CSV created Successfully"
Notifi.configure(text=O, bg="Green", fg="white", width=33, font=('times', 19))
Notifi.place(x=180, y=380)
import csv
import tkinter
root = tkinter.Tk()
root.title("log of " + subb)
with open(csv_name, newline="") as file:
reader = csv.reader(file)
r = 0
for col in reader:
c = 0
for row in col:
# i've added some styling
label = tkinter.Label(root, width=13, height=1, fg="black", font=('times', 13, ' bold '),
bg="white", text=row, relief=tkinter.RIDGE)
label.grid(row=r, column=c)
c += 1
r += 1
root.mainloop()
Notifi = tk.Label(MFW, text="CSV created Successfully", font=('times', 19))
####### one clear button for both
c1ear_enroll = tk.Button(MFW, text="clear", command=remove_enr,activebackground='#999999',bg='#b9b9b9',borderwidth=0,font=('times', 15, ' bold '))
c1ear_enroll.grid(row=1,column=1,padx=(0,10),sticky='W',pady=(3,10))
#c1ear_enroll.place(x=690, y=100)
c1ear_student = tk.Button(MFW, text="clear", command=remove_student,activebackground='#999999',bg='#b9b9b9',borderwidth=0, font=('times', 15, ' bold '))
c1ear_student.grid(row=3,column=1,padx=(0,10),sticky='W',pady=(3,10))
#c1ear_student.place(x=690, y=200)
###
DATA_SUB = tk.Button(MFW, text="Add more",command=enter_data_DB,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1, font=('times', 15))
DATA_SUB.grid(row=4,column=0,padx=10,pady=10)
#DATA_SUB.place(x=170, y=300)
MAKE_CSV = tk.Button(MFW, text="Convert to CSV",command=create_csv,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times', 15))
MAKE_CSV.grid(row=4,column=1,padx=10,pady=10)
#MAKE_CSV.place(x=570, y=300)
#def attf():
# import subprocess
# subprocess.Popen(r'explorer /select,"E:/Attendace managemnt system\Attendance\Manually Attendance\-------Check atttendance-------"')
#attf = tk.Button(MFW, text="Check Sheets",command=attf,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1 ,font=('times', 14, ' bold '))
#attf.place(x=730, y=410)
status = Label(MFW, textvariable=displaystring,font=('times',12),fg='#444444')
status.grid(row=8,columnspan=8,sticky='W',pady=(10,0))
MFW.mainloop()
SUB = tk.Label(sb, text="Enter room number", font=('times', 15))
SUB.grid(row=0,column=0,sticky=('S','W'),padx=10,pady=(10,0))
#SUB.place(x=30, y=100)
global SUB_ENTRY
SUB_ENTRY = tk.Entry(sb, width=20, bg="white", fg="black", font=('times', 23, ' bold '))
SUB_ENTRY.grid(row=1,column=0,padx=10,pady=(0,10))
#SUB_ENTRY.place(x=250, y=105)
fill_manual_attendance = tk.Button(sb, text="Fill log",command=fill_attendance, width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1, font=('times', 15))
fill_manual_attendance.grid(row=2,column=0,sticky=('E'),padx=10,pady=5)
#fill_manual_attendance.place(x=250, y=160)
status = Label(sb, textvariable=displaystring,font=('times',12),fg='#444444')
status.grid(row=8,columnspan=8,sticky='W',pady=(10,0))
sb.mainloop()
# +
##For clear textbox
def clearall():
txt.delete(first=0, last=22)
txt2.delete(first=0, last=22)
displaystring.set("click on 'Take Images' to proceed")
def clear():
txt.delete(first=0, last=22)
def clear1():
txt2.delete(first=0, last=22)
def del_sc1():
sc1.destroy()
def err_screen():
global sc1
sc1 = tk.Tk()
sc1.geometry('300x100')
sc1.iconbitmap('AMS.ico')
sc1.title('Warning!!')
#sc1.configure(background='snow')
Label(sc1,text='ID & Name required!!!',font=('times', 16, ' bold ')).pack()
Button(sc1,text='OK',command=del_sc1 ,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times', 15)).place(x=90,y= 50)
##Error screen2
def del_sc2():
sc2.destroy()
def err_screen1():
global sc2
sc2 = tk.Tk()
sc2.geometry('300x100')
sc2.iconbitmap('AMS.ico')
sc2.title('Warning!!')
#sc2.configure(background='snow')
Label(sc2,text='Please enter the room number!!!',font=('times', 16)).pack()
Button(sc2,text='OK',command=del_sc2 ,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times', 15)).place(x=90,y= 50)
# +
# take images
def take_img():
l1 = txt.get()
l2 = txt2.get()
if l1 == '':
err_screen() #### add message boxes
elif l2 == '':
err_screen()
else:
try:
cam = cv2.VideoCapture(0)
detector = cv2.CascadeClassifier('haarcascade_frontalface_default.xml')
Enrollment = txt.get()
Name = txt2.get()
sampleNum = 0
while (True):
ret, img = cam.read()
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
faces = detector.detectMultiScale(gray, 1.3, 5)
for (x, y, w, h) in faces:
cv2.rectangle(img, (x, y), (x + w, y + h), (255, 0, 0), 2)
# incrementing sample number
sampleNum = sampleNum + 1
# saving the captured face in the dataset folder
cv2.imwrite("TrainingImage/ " + Name + "." + Enrollment + '.' + str(sampleNum) + ".jpg",
gray[y:y + h, x:x + w])
cv2.imshow('Frame', img)
# wait for 100 miliseconds
if cv2.waitKey(1) & 0xFF == ord('q'):
break
# break if the sample number is morethan 100
elif sampleNum > 70:
break
cam.release()
cv2.destroyAllWindows()
ts = time.time()
Date = datetime.datetime.fromtimestamp(ts).strftime('%Y-%m-%d')
Time = datetime.datetime.fromtimestamp(ts).strftime('%H:%M:%S')
row = [Enrollment, Name, Date, Time]
with open('StudentDetails\StudentDetails.csv', 'a+') as csvFile:
writer = csv.writer(csvFile, delimiter=',')
writer.writerow(row)
csvFile.close()
#####statusbar
res = "Images Saved for Employee with ID : " + Enrollment + " Name : " + Name
Notification.configure(text=res, bg="SpringGreen3", width=50, font=('times', 18, 'bold'))
Notification.place(x=250, y=400)
except FileExistsError as F:
####message box
f = 'Employee Data already exists'
Notification.configure(text=f, bg="Red", width=21)
Notification.place(x=450, y=400)
# +
#automatic fill log
def subjectchoose():
def Fillattendances():
sub=tx.get()
now = time.time() ###For calculate seconds of video
future = now + 20
if time.time() < future:
if sub == '':
err_screen1() ##add error message instead
else:
recognizer = cv2.face.LBPHFaceRecognizer_create() # cv2.createLBPHFaceRecognizer()
try:
recognizer.read("TrainingImageLabel\Trainner.yml")
except:
e = 'Model not found,Please train model' #add error message box
Notifica.configure(text=e, bg="red", fg="black", width=33, font=('times', 15, 'bold'))
Notifica.place(x=20, y=250)
harcascadePath = "haarcascade_frontalface_default.xml"
faceCascade = cv2.CascadeClassifier(harcascadePath)
df = pd.read_csv("StudentDetails\StudentDetails.csv")
cam = cv2.VideoCapture(0)
font = cv2.FONT_HERSHEY_SIMPLEX
col_names = ['Enrollment', 'Name', 'Date', 'Time']
attendance = pd.DataFrame(columns=col_names)
while True:
ret, im = cam.read()
gray = cv2.cvtColor(im, cv2.COLOR_BGR2GRAY)
faces = faceCascade.detectMultiScale(gray, 1.2, 5)
for (x, y, w, h) in faces:
global Id
Id, conf = recognizer.predict(gray[y:y + h, x:x + w])
if (conf <70):
print(conf)
global Subject
global aa
global date
global timeStamp
Subject = tx.get()
ts = time.time()
date = datetime.datetime.fromtimestamp(ts).strftime('%Y-%m-%d')
timeStamp = datetime.datetime.fromtimestamp(ts).strftime('%H:%M:%S')
aa = df.loc[df['Enrollment'] == Id]['Name'].values
global tt
tt = str(Id) + "-" + aa
En = '15624031' + str(Id)
attendance.loc[len(attendance)] = [Id, aa, date, timeStamp]
cv2.rectangle(im, (x, y), (x + w, y + h), (0, 260, 0), 7)
cv2.putText(im, str(tt), (x + h, y), font, 1, (255, 255, 0,), 4)
else:
Id = 'Unknown'
tt = str(Id)
cv2.rectangle(im, (x, y), (x + w, y + h), (0, 25, 255), 7)
cv2.putText(im, str(tt), (x + h, y), font, 1, (0, 25, 255), 4)
if time.time() > future:
break
attendance = attendance.drop_duplicates(['Enrollment'], keep='first')
cv2.imshow('Filling attedance..', im)
key = cv2.waitKey(15) & 0xff
if key == 27:
break
ts = time.time()
date = datetime.datetime.fromtimestamp(ts).strftime('%Y-%m-%d')
timeStamp = datetime.datetime.fromtimestamp(ts).strftime('%H:%M:%S')
Hour, Minute, Second = timeStamp.split(":")
fileName = "Attendance/" + Subject + "_" + date + "_" + Hour + "-" + Minute + "-" + Second + ".csv"
attendance = attendance.drop_duplicates(['Enrollment'], keep='first')
print(attendance)
attendance.to_csv(fileName, index=False)
##Create table for Attendance
date_for_DB = datetime.datetime.fromtimestamp(ts).strftime('%Y_%m_%d')
DB_Table_name = str( Subject + "_" + date_for_DB + "_Time_" + Hour + "_" + Minute + "_" + Second)
import pymysql.connections
###Connect to the database
try:
global cursor
connection = pymysql.connect(host='localhost', user='root', password='', db='Face_reco_fill')
cursor = connection.cursor()
except Exception as e:
print(e)
sql = "CREATE TABLE " + DB_Table_name + """
(ID INT NOT NULL AUTO_INCREMENT,
ENROLLMENT varchar(100) NOT NULL,
NAME VARCHAR(50) NOT NULL,
DATE VARCHAR(20) NOT NULL,
TIME VARCHAR(20) NOT NULL,
PRIMARY KEY (ID)
);
"""
####Now enter attendance in Database
insert_data = "INSERT INTO " + DB_Table_name + " (ID,ENROLLMENT,NAME,DATE,TIME) VALUES (0, %s, %s, %s,%s)"
VALUES = (str(Id), str(aa), str(date), str(timeStamp))
try:
cursor.execute(sql) ##for create a table
cursor.execute(insert_data, VALUES)##For insert data into table
except Exception as ex:
print(ex) #
####### create status bar for This#####
M = 'log filled Successfully'
Notifica.configure(text=M, bg="Green", fg="white", width=33, font=('times', 15, 'bold'))
Notifica.place(x=20, y=250)
cam.release()
cv2.destroyAllWindows()
import csv
import tkinter
root = tkinter.Tk()
root.title("Log of " + Subject)
cs = 'E:/Attendace managemnt system/' + fileName
with open(cs, newline="") as file:
reader = csv.reader(file)
r = 0
for col in reader:
c = 0
for row in col:
# i've added some styling
label = tkinter.Label(root, width=8, height=1, fg="black", font=('times', 15),
bg="white", text=row, relief=tkinter.RIDGE)
label.grid(row=r, column=c)
c += 1
r += 1
root.mainloop()
print(attendance)
###windo is frame for subject chooser
windo = tk.Tk()
windo.iconbitmap('AMS.ico')
windo.title("Automatically fill log")
#windo.geometry('580x320')
Notifica = tk.Label(windo, text="log filled Successfully", font=('times', 15))
#def Attf():
# import subprocess
# subprocess.Popen(r'explorer /select,"E:/Attendace managemnt system\Attendance\-------Check atttendance-------"')
#attf = tk.Button(windo, text="Check Sheets",command=Attf,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times', 14))
#attf.place(x=430, y=255)
sub = tk.Label(windo, text="Room Number", font=('times', 15))
sub.grid(row=0,column=0,sticky=('W','S'),padx=10,pady=(10,0))
#sub.place(x=30, y=100)
tx = tk.Entry(windo, width=20, font=('times', 23))
tx.grid(row=1,column=0,padx=10,pady=(3,5))
#tx.place(x=250, y=105)
fill_a = tk.Button(windo, text="Register",command=Fillattendances ,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1, font=('times', 15))
fill_a.grid(row=2,column=0,sticky='E',padx=10,pady=10)
#fill_a.place(x=250, y=160)
status = Label(windo, textvariable=displaystring,font=('times',12),fg='#444444')
status.grid(row=8,columnspan=8,sticky='W',pady=(10,0))
windo.mainloop()
def admin_panel():
win = tk.Tk()
win.iconbitmap('AMS.ico')
win.title("Admin Login")
#win.geometry('880x420')
def log_in():
username = un_entr.get()
password = pw_entr.get()
if username == 'k' :
if password == 'k':
win.destroy()
import csv
import tkinter
root = tkinter.Tk()
root.title("Employee Details")
cs = 'E:/Attendace managemnt system/StudentDetails/StudentDetails.csv'
with open(cs, newline="") as file:
reader = csv.reader(file)
r = 0
for col in reader:
c = 0
for row in col:
# i've added some styling
label = tkinter.Label(root, width=8, font=('times', 15), text=row, relief=tkinter.RIDGE)
label.grid(row=r, column=c)
c += 1
r += 1
root.mainloop()
else:
#######message box
valid = 'Incorrect ID or Password'
Nt.configure(text=valid, bg="red", fg="black", width=38, font=('times', 19, 'bold'))
Nt.place(x=120, y=350)
else:
valid ='Incorrect ID or Password'
Nt.configure(text=valid, bg="red", fg="black", width=38, font=('times', 19, 'bold'))
Nt.place(x=120, y=350)
Nt = tk.Label(win, text="log filled Successfully", font=('times', 19))
# Nt.place(x=120, y=350)
un = tk.Label(win, text="Enter username :",font=('times', 15))
un.grid(row=0,column=0,sticky=('S','W'),padx=10,pady=(10,0))
#un.place(x=30, y=50)
pw = tk.Label(win, text="Enter password :", font=('times', 15))
pw.grid(row=2,column=0,sticky=('S','W'),padx=10,pady=(10,0))
#pw.place(x=30, y=150)
def c00():
un_entr.delete(first=0, last=22)
un_entr = tk.Entry(win, width=20, font=('times', 23))
un_entr.grid(row=1,column=0,padx=(10,0),pady=(3,10))
#un_entr.place(x=290, y=55)
def c11():
pw_entr.delete(first=0, last=22)
pw_entr = tk.Entry(win, width=20,show="*", font=('times', 23))
pw_entr.grid(row=3,column=0,padx=(10,0),pady=(3,10))
#pw_entr.place(x=290, y=155)
#### make one clear button for both
c0 = tk.Button(win, text="Clear", command=c00,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=0, font=('times', 15))
c0.grid(row=1,column=1,sticky=('W'),pady=(3,10),padx=(0,10))
#c0.place(x=690, y=55)
c1 = tk.Button(win, text="Clear", command=c11,width=13 ,activebackground='#999999',bg='#b9b9b9',borderwidth=0, font=('times', 15))
c1.grid(row=3,column=1,sticky=('W'),pady=(3,10),padx=(0,10))
#c1.place(x=690, y=155)
####
Login = tk.Button(win, text="Login", command=log_in,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1, font=('times', 15))
Login.grid(row=4,column=0,sticky='E',padx=10,pady=10)
#Login.place(x=290, y=250)
status = Label(win, textvariable=displaystring,font=('times',12),fg='#444444')
status.grid(row=8,columnspan=8,sticky='W',pady=(10,0))
win.mainloop()
# +
####### For train the model ########
def trainimg():
recognizer = cv2.face.LBPHFaceRecognizer_create()
global detector
detector = cv2.CascadeClassifier("haarcascade_frontalface_default.xml")
try:
global faces,Id
faces, Id = getImagesAndLabels("TrainingImage")
except Exception as e:
## error message#############
l='please make "TrainingImage" folder & put Images'
#
Notification.configure(text=l, bg="white", width=50, font=('times', 18, 'bold'))
Notification.place(x=350, y=400)
recognizer.train(faces, np.array(Id))
try:
recognizer.save("TrainingImageLabel\Trainner.yml")
except Exception as e:
q='Please make "TrainingImageLabel" folder'
Notification.configure(text=q, bg="SpringGreen3", width=50, font=('times', 18, 'bold'))
Notification.place(x=350, y=400)
res = "Model Trained" # +",".join(str(f) for f in Id)
Notification.configure(text=res, bg="SpringGreen3", width=50, font=('times', 18, 'bold'))
Notification.grid(row=8,columnspan=8,sticky='W',pady=(5,0))
#Notification.place(x=250, y=400)
def getImagesAndLabels(path):
imagePaths = [os.path.join(path, f) for f in os.listdir(path)]
# create empth face list
faceSamples = []
# create empty ID list
Ids = []
# now looping through all the image paths and loading the Ids and the images
for imagePath in imagePaths:
# loading the image and converting it to gray scale
pilImage = Image.open(imagePath).convert('L')
# Now we are converting the PIL image into numpy array
imageNp = np.array(pilImage, 'uint8')
# getting the Id from the image
Id = int(os.path.split(imagePath)[-1].split(".")[1])
# extract the face from the training image sample
faces = detector.detectMultiScale(imageNp)
# If a face is there then append that in the list as well as Id of it
for (x, y, w, h) in faces:
faceSamples.append(imageNp[y:y + h, x:x + w])
Ids.append(Id)
return faceSamples, Ids
#############################
window.grid_rowconfigure(0, weight=1)
window.grid_columnconfigure(0, weight=1)
window.resizable(False, False)
window.iconbitmap('AMS.ico')
Notification = tk.Label(window, text="All things good", bg="Green", fg="white", width=15,
height=3, font=('times', 17, 'bold'))
lbl = tk.Label(window, text="Employee ID", font=('times', 15))
lbl.grid(row=0, column=0,sticky='W')
def testVal(inStr, acttyp):
if acttyp == '1': #insert
if not inStr.isdigit():
return False
return True
txt = tk.Entry(window, validate="key", width=20, bg="white", font=('times', 20))
txt['validatecommand'] = (txt.register(testVal),'%P','%d')
txt.grid(row=0,column=1)
lbl2 = tk.Label(window, text="Employee Name", font=('times', 15))
lbl2.grid(row=1,column=0,sticky='W')
txt2 = tk.Entry(window, width=20, bg="white",font=('times', 20))
txt2.grid(row=1,column=1)
EnterBtn = tk.Button(window, text="Register",width=13, command =clearall,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times',15))
EnterBtn.grid(row=2,column=1,sticky='E',pady=5)
clearButton = tk.Button(window,text="clear",command=clear,activebackground='#999999',bg='#b9b9b9',borderwidth=0,font=('times',15))
clearButton.grid(row=0,column=2,sticky='W',pady=5)
clearButton1 = tk.Button(window,text="clear",command=clear1,activebackground='#999999',bg='#b9b9b9',borderwidth=0,font=('times',15))
clearButton1.grid(row=1,column=2,sticky='W',pady=5)
#maybe after pressing enter, open new window with cam on and take images button
AP = tk.Button(window, text="Check log",command=admin_panel,width=13,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times',15 ))
AP.grid(row=3,column=3,pady=10,padx=(45,10))
takeImg = tk.Button(window, text="Take Images",width=13,command=take_img,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times', 15))
takeImg.grid(row=4,column=0,pady=5,padx=10)
trainImg = tk.Button(window, text="Train Images",width=13,command=trainimg,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=( 'times',15))
trainImg.grid(row=4,column=1,pady=5,padx=10)
FA = tk.Button(window, text="Automatic log",width=13,command=subjectchoose,activebackground='#999999',bg='#b9b9b9',borderwidth=1,font=('times',15 ))
FA.grid(row=4,column=2,pady=5,padx=10)
quitWindow = tk.Button(window, text="Manual log", command=manually_fill, activebackground='#999999',width=13,bg='#b9b9b9',borderwidth=1,font=('times',15))
quitWindow.grid(row=4,column=3,pady=5,padx=(45,10))
window.mainloop()
# -
|
AMS_Run_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3.8
# ---
# # Introduction to Dask
#
# In this notebook, we'll learn how to use [Dask](https://dask.org) for reading data from Azure.
#
# The main [dask](https://github.com/dask/dask) and [distributed](https://github.com/dask/distributed) themselves are small and focused. Thousands of tools, some built by the Dask organization and most not, utilize Dask for parallel or distributed processing. Some of the most useful for data science include:
#
# - [dask/adlfs](https://github.com/dask/adlfs)
# - [dask/dask-ml](https://github.com/dask/dask-ml)
# - [pydata/xarray](https://github.com/pydata/xarray)
# - [microsoft/lightgbm](https://github.com/microsoft/lightgbm)
# - [dmlc/xgboost](https://github.com/dmlc/xgboost)
# - [rapidsai/cudf](https://github.com/rapidsai/cudf)
# - [rapidsai/cuml](https://github.com/rapidsai/cuml)
# ## Install required packages
# !pip install --upgrade dask distributed bokeh adlfs fsspec fastparquet pyarrow python-snappy lz4
# ## Get AML Workspace
#
# You can use the AML workspace to retrieve datastores and keyvaults for accessing data credentials securely.
# +
from azureml.core import Workspace
ws = Workspace.from_config()
ws
# -
# ## Create a distributed client
#
# The [client](https://distributed.dask.org/en/latest/client.html) is the primary entrypoint for parallel processing with Dask. Calling it without inputs will create a local distributed scheduler, utilizing all the CPUs and cores on your machine. This can be useful for faster processing of larger in memory dataframes, or even computations on out of memory (OOM) data.
#
# When your local machine isn't powerful enough, you can provision a larger VM in Azure - the M series has 100+ CPUs and TBs of RAM. If this still isn't powerful enough, you can create a distributed Dask cluster on most hardware - see [the Dask setup guide](https://docs.dask.org/en/latest/setup.html) for details.
#
# If you still need acceleration, [RAPIDSAI](https://github.com/rapidsai) further extends the PyData APIs on GPUs.
#
# **Make sure you check out the dashboard!**
# +
from distributed import Client
c = Client()
print(c)
c
# -
# ## Reading cloud data
#
# Reading data from the cloud is as easy as reading it locally! Sorta!
#
# ### Pandas
#
# You can read directly into Pandas from most cloud storage, with a notable exception - from the [`pandas.read_csv` documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html?highlight=read_csv#pandas-read-csv):
#
# > Any valid string path is acceptable. The string could be a URL. Valid URL schemes include http, https, ftp, s3, gs, and file.
#
# ### Pythonic Filesystem for Azure
#
# Fortunately, similar protocols have been developed for Azure storage in the [ADLFS](https://github.com/dask/adlfs) package, including:
#
# - `az` or `abfs` for Azure Data Lake Storage Gen2 (ADLSv2) and Blob
# - `adl` for Azure Data Lake Storage Gen1 (ADLSv1)
#
# These are included in Python's `fsspec`. You can use the protocol directly in Dask and convert to Pandas for now.
#
# +
import pandas as pd
import dask.dataframe as dd
from adlfs import AzureBlobFileSystem
# -
# for single files in public blobs, you can use the https protocol and read directly into Pandas
df = pd.read_csv(
"https://azuremlexamples.blob.core.windows.net/datasets/iris.csv"
)
df.head()
# the same with Dask
df = dd.read_csv(
"https://azuremlexamples.blob.core.windows.net/datasets/iris.csv"
)
df.head()
# alternative syntax in Dask
storage_options = {"account_name": "azuremlexamples"}
df = dd.read_csv(f"az://datasets/iris.csv", storage_options=storage_options)
df.head()
# ## Why use Dask?
#
# Those all did the same thing...so why use Dask? There are a few scenarios:
#
# - reading multiple files
# - reading private data from Azure with credentials
# - reading directly into GPUs (with [cuDF](https://github.com/rapidsai/cudf))
#
# You can also use the classes implemented in `adlfs` to query for files, depending on permissions.
#
# To provide your own credentials, refer to the `adlfs` documentation for details - generally you can retrieve credentials from the workspace's datastore:
#
# ```python
# import dask.dataframe as dd
# from azureml.core import Workspace
#
# ws = Workspace.from_config()
# ds = ws.get_default_datastore() # ws.datastores["my-datastore-name"]
#
# storage_options = {
# "account_name": ds.account_name,
# "account_key": ds.account_key,
# }
#
# df = dd.read_parquet(f"az://{ds.container_name}/path/to/data/*.parquet", storage_options=storage_options)
# ```
#
# The basics are demonstrated below on public data.
color = "green"
container_name = "nyctlc"
storage_options = {"account_name": "azureopendatastorage"}
fs = AzureBlobFileSystem(**storage_options)
fs
fs.ls(f"{container_name}")
fs.ls(f"{container_name}/{color}")
fs.ls(f"{container_name}/{color}/puYear=2016/")
files = fs.glob(f"{container_name}/{color}/puYear=2016/puMonth=12/*.parquet")
files = [f"az://{file}" for file in files]
files[-5:]
# + tags=[]
# %%time
ddf = (
dd.read_parquet(files, storage_options=storage_options)
.repartition(npartitions=8)
.persist()
)
ddf
# -
# %%time
len(ddf)
# %%time
len(ddf)
ddf.info()
# +
import matplotlib.pyplot as plt
plt.style.use("dark_background")
ddf["tipAmount"].compute().hist(
figsize=(16, 8), bins=256, range=(0.1, 20),
)
# -
df = ddf.compute()
df.info()
# %%time
df.describe()
# %%time
gbs = round(df.memory_usage(index=True, deep=True).sum() / 1e9, 2)
print(f"df is {gbs} GBs")
# %%time
gbs = round(ddf.memory_usage(index=True, deep=True).sum().compute() / 1e9, 2)
print(f"ddf is {gbs} GBs")
|
tutorials/using-dask/1.intro-to-dask.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.insert(0, '../Libraries/')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# # Weather data
# * Source: https://www.visualcrossing.com/ Fee for uploading more than 100 lines of info. Please contact me with your needs, I will happily pay for your data
# * Units: Wind is km/h, Temperature Celsius, Solar Radiation W/m^2
#
weather = pd.read_excel("Climate/EssexJct-05012020-10312020.xlsx",header=0,date_parser=[5],)
weather.columns #To remind you what's in spreadsheet
weather = weather.set_index('Date time')
weather['Solar Radiation'] = weather['Solar Radiation'].fillna(0.0)
# # The slab problem
#
# A slab made of insulation foam panels with thermal conductivity $k = 0.022 \mathrm{W}/\mathrm{m}.\mathrm{K}$ (https://www.poliuretano.it/EN/thermal_conductivity_polyurethane.html), thickness $5\mathrm{cm}$ and dimensions $10\times10\mathrm{m}^2$ is subjected to the weather of Essex Junction, VT. The bottom surface temperature is maintained at a constant temperature $T_{s,lower}$. **Compute the weekly, monthly and total heat rate necessary to maintain** $T_{s,lower}=0^\circ\mathrm{C}$
#
# ### Assumptions:
# * The heat transfer from rain is not considered
# * The position of the sun is not considered (The plate receives the full solar irradiation)
# * The direction of the wind is always assumed to be perpendicular to one of the edges of the plate
# * The time scale of the fluctuations of wall heat flux from convection and radiation are long enough that the temperature profile can be approximated as linear at all times
# * The upper surface temperature is assumed to be spatially uniform
#
# ### Equations
#
# The heat rate necessary to keep the lower surface at $0^\circ\mathrm{C}$ is
# $$
# q''_{cond}=\frac{k}{H}\left(T_{s,upper}-T_{s,lower}\right)
# $$
#
# To determine $T_{s,upper}$, the conservation of energy is applied to the upper surface:
# $$
# \rho C_p \frac{dT_{s,upper}}{dt}AH = g(T_{s,upper},t)=\sum q_{in}-\sum q_{out}
# $$
# or using $T=T_{s,upper}$
# $$
# \frac{dT}{dt} = f(T,t)=\frac{1}{\rho C_pAH}\sum q_{in}-\sum q_{out}
# $$
# For now we consider only the effect from convection and solar irradiation, assuming that the irradiation reported in the data is absorbed by the surface with an absorptivity $\alpha_s$.
#
# $$
# \sum q_{in}= hA(T_\infty - T_{s,upper})
# $$
# $$
# \sum q_{out}=\frac{kA}{H}(T_{s,upper} - T_{s,lower})
# $$
# Note that when $T_\infty$ is lower than $T_{s,upper}$, the $\sum q_{in}$ is negative which is the correct physical process. The convection coefficient is to be determined with the appropiate correlation.
#
# For the initial condition, $t=0$, $T_{s,upper}$ is assumed to be at ambient temperature. The solution of the conservation of energy is updated numerically.
#
# The simplest, but least accurate method, is the Euler method, where
# $$
# \frac{dT}{dt} = \frac{T(t+\Delta t)-T(t)}{\Delta t}
# $$
# or
#
# $$
# T(t+\Delta t)=T(t)+(\Delta t)f(T,t)
# $$
#
# This method's error is first order, i.e. the leading term in the truncation error term is affected by $\Delta t$. It is also an unstable time integration method unless $\Delta t$ is very small. To increase accuracy and stability we propose to use the second order predictor-corrector method, which is a two step process. The first step is to estimate $T$ and $t+\Delta t$ using the information available at $t$
# * **Predictor step**
#
# $$
# T^*=T(t)+(\Delta t)f(T,t)
# $$
#
# The second is to correct this estimate by taking the average of the RHS, $f$, of our ODE between the predictor step and the RHS at $f(T^*,t+\Delta t)$
#
# * **Corrector step**
#
# $$
# T(t+\Delta t)=T(t)+\frac{(\Delta t)}{2}\left[f(T,t)+f(T^*,t+\Delta t)\right]
# $$
# This method is second order accurate in time.
#
# ### The stability problem
#
# Even with the predictor corrector, the time step of integration cannot be too large. The $15\mathrm{min}$ increment of the data is too long and leads to `NaN` quickly into the simulation.
#
# The following cell interpolates the data for a smaller time increment. Obviously the smaller the increment, the larger computational time.
#
# With the entire data, the difference between $\Delta t=60\mathrm{s}$ and $\Delta t = 120\mathrm{s}$ is negligible, however the temperature diverges for $\Delta t = 240\mathrm{s}$ around August.
#
# ### More robust approach
#
# Stiff integrators are higher order method for time dependent systems of ODE of the form
# $$
# \frac{dy}{dt}=f(t,y)
# $$
# where $y$ is a vector of variable. These integrators are available in the function `from scipy.integrate import solve_ivp`, where ivp stands for initial value problem. The following cells mirrors the solution of noetbook `Data-manipulation-2`, using `solve_ivp`
#
# Note that some of the data in the spreadsheet is missing, hence the `interpolate` step to reconstruct the missing data
# ### Berdhal and Martin (1984) model
#
# * $c\in[0,1]$ cloudiness/cloud coverage (0 = clear, 1 = total cloud coverage)
# * $T_{air}\, [^\circ\mathrm{K}]$ Air temperature
# * $T_{dp}\, [^\circ\mathrm{C}]$ Dew point temperature (note unit, important because relates to humidity)
#
# $$
# \varepsilon_{clear} = 0.711 + 0.56(T_{dp}/100.) + 0.73(T_{dp}/100.)^2
# $$
# $$
# C_a = 1. + 0.02224c + 0.0035c^2 + 0.00028c^3
# $$
# $$
# T_{sky}=(C_a\varepsilon_{clear})^{0.25}T_{air}
# $$
# $$
# \epsilon_{sky}=1
# $$
# +
import schemdraw as schem
import schemdraw.elements as e
import matplotlib.pyplot as plt
import numpy as np
import math
import scipy.constants as sc
import HT_thermal_resistance as res
R = []
R.append(res.Resistance(name="$R_{snow}$", units="W/m"))
R.append(res.Resistance(name="$R_{cond}$", units="W/m"))
R.append(res.Resistance(name="$R_{conv}$", units="W/m"))
R.append(res.Resistance(name="$R_{rad}$", units="W/m"))
d = schem.Drawing()
d.add(e.DOT, label = r"$T_0$")
d.add(e.RES, d = 'right', label = R[0].name)
d.add(e.DOT, label = r"$T_{s,lower}$")
R1 = d.add(e.RES, d = 'right', label = R[1].name)
d.add(e.DOT, rgtlabel = r"$T_{s,upper}$")
d.push()
d.add(e.LINE, d= 'up', l = 1.5)
d.add(e.RES, d='right', label = R[2].name)
d.add(e.DOT, rgtlabel="$T_{air}$")
d.pop()
d.add(e.LINE, d='down', l = 1.5)
d.add(e.RES, d='right', botlabel = R[3].name)
d.add(e.DOT, rgtlabel="$T_{sky}$")
L1 = d.add(e.LINE, toplabel = "$q$", endpts = [[-0.25, 0], [-2.25, 0]])
d.labelI(L1, arrowofst=0)
d.draw()
# +
from scipy.interpolate import interp1d
t_data = np.arange(0,weather.shape[0]*15*60,15*60) #Because data collected in spreadsheet in 15 min intervals
#interpolation to adjust for holes
weather['Wind Speed'] = weather['Wind Speed'].interpolate('cubic')
weather['Temperature'] = weather['Temperature'].interpolate('cubic')
weather['Solar Radiation'] = weather['Solar Radiation'].interpolate('cubic')
weather['Cloud Cover'] = weather['Cloud Cover'].interpolate('cubic')
weather['Dew Point'] = weather['Dew Point'].interpolate('cubic')
U_atm = np.abs(weather['Wind Speed'][:].to_numpy()/3.6)
T_atm = weather['Temperature'][:].to_numpy()
q_sun = weather['Solar Radiation'][:].to_numpy()
cc = weather['Cloud Cover'][:].to_numpy()
cc /= 100 #To make 0 to 1 because written as percent in spreadsheet
T_dp = weather['Dew Point'][:].to_numpy()
f_U_atm = interp1d(t_data,U_atm,kind='cubic')
f_T_atm = interp1d(t_data,T_atm,kind='cubic')
f_q_sun = interp1d(t_data,q_sun,kind='cubic')
f_cc = interp1d(t_data,cc,kind='cubic')
f_T_dp = interp1d(t_data,T_dp,kind='cubic')
# -
#how to get rid of outliers
cctmp = np.minimum(cc, np.ones_like(cc))
cc = np.maximum(cctmp, np.zeros_like(cc))
np.max(cctmp)
plt.plot(cc)
# +
import scipy.constants as csts
from scipy.integrate import solve_ivp
import thermodynamics as thermo
import HT_external_convection as extconv
import HT_natural_convection as natconv
#Necessary Parameters
rho = 20.
C_p = 2500.
k = 0.05
H = 0.05
alpha_s = 0.5
eps = 0.5
Lplate = 1.
A = Lplate**2
T_s_lower = 0.
eps_sky = 1.
def f(t,T): #function to integrate
global Lplate,k,H,eps,f_U_atm,f_T_atm,f_q_sun,rho,C_p,alpha_s,f_cc,f_T_dp, q_out
# film temperature between ambient and upper surface
Tinf = f_T_atm(t)
Uinf = np.max([f_U_atm(t),0.]) #wind speed?
q_irr = alpha_s*f_q_sun(t) #irradition of sun* absorbtivity of surface
T_f = (T[0]+Tinf)/2
air_f = thermo.Fluid('air',T_f,"C")
Re = np.abs(Uinf)*Lplate/air_f.nu
Gr = natconv.Gr(beta=air_f.beta,DT=np.abs(T-T_f),D=Lplate,nu=air_f.nu)
Ra = natconv.Ra(beta=air_f.beta,DT=np.abs(T-T_f),D=Lplate,nu=air_f.nu,alpha=air_f.alpha)
# Forced convection flux
if Re >0 and (Re <= 5e5):
airflow = extconv.FlatPlate('laminar','isothermal',U_infty=Uinf,nu=air_f.nu,alpha=air_f.alpha, L=Lplate,xi=0,Re_xc= 5e5)
airflow.average(Lplate)
hconv_f = airflow.Nu_ave*air_f.k/Lplate
elif Re > 5e5:
airflow = extconv.FlatPlate('mixed','isothermal',U_infty=Uinf,nu=air_f.nu,alpha=air_f.alpha, L=Lplate,xi=0,Re_xc= 5e5)
airflow.average(Lplate)
hconv_f = airflow.Nu_ave*air_f.k/Lplate
else:
hconv_f = 0
#Natural convection flux
if Ra > 1e4:
if T[0] >= Tinf:
airflow = natconv.FlatPlate(Ra,air_f.Pr,'upper','hot')
else:
airflow = natconv.FlatPlate(Ra,air_f.Pr,'upper','cold')
hconv_n = airflow.Nu*air_f.k/Lplate
else:
hconv_n = 0
#Total convection flux (here not a function of Ri)
h = hconv_n + hconv_f
qpp_conv = h*(Tinf - T[0])
#Conduction flux
qpp_cond = k*(T[0] - T_s_lower)/H
# if i > 1050:
# print(i,hconv_n, hconv_f)
# print(i,"DT",Tinf - T,T - T_s_lower)
# print(i,"qpp",qpp_conv,qpp_cond)
# print(q_irr,qpp_conv,qsky)
# print(T_f,Tsky,cc,Ca,eps_clear,csts.sigma)
#Radiation flux
T_dp = f_T_dp(t) #interpolation func of dew point temp, t is time
cc= f_cc(t)
epsclear = 0.711 + 0.56*(T_dp/100) + 0.73*(T_dp/100)**2
Ca = 1 + 0.02224*cc + 0.0035*cc**2 + 0.00028*cc**3
T_sky = (Ca*epsclear)**0.25 *Tinf
T_s_upper = T[0] #Where does T[0] get defined
q_rad = eps*csts.sigma*(thermo.C2K(np.array(T_sky))**4 - thermo.C2K(T_s_upper)**4) #- q_irr ?
q_out = (q_irr + qpp_conv - qpp_cond + q_rad)/(rho*C_p)
return q_out
# Simulation
# f(8000,[15])
# sol = solve_ivp(f,[0,t_data[-1]],[0.],t_eval=t_data,max_step=60.)
tmax = t_data[-1] #max value for integration
mask = np.where(t_data <= tmax)
sol = solve_ivp(f,[0,tmax],[10.],method='LSODA',t_eval=t_data[mask],max_step=5*60.) #actual integration process, can vary max_step but doesn't all work
# sol = solve_ivp(f,[0,100000],[0.])
# -
weather['Ts upper'] = sol.y[0]
weather['q out'] = k*A*sol.y[0]/H
weather['q irr'] = alpha_s*weather['Solar Radiation'][:]*A
weather.plot(y='Ts upper')
ax0 = weather.plot(y='q irr')
weather.plot(y='q out',ax = ax0)
# Compare the cumulative sum of the heat rate necessary to maintain the lower surface's temperature at 0C with and without the radiative heat transfer between the upper surface and the sky.
weather['q out'].sum() #[W]
# weather['Temperature'].interpolate(method='spline')
np.argwhere(np.isnan(weather['Temperature'][:].to_numpy()))
ax0 = weather.plot(y='Ts upper')
weather.plot(y='Temperature',ax = ax0)
weather_1H = weather.resample("1H").mean()
weather_1H['Energy slab'] = k*A/H*weather_1H['Ts upper']
ax0 = weather_1H.plot(y='Ts upper')
weather_1H.plot(y='Temperature',ax = ax0)
weather_1H.plot(y='Energy slab')
weather_1H['Energy slab'].sum()
|
Snow-Cooling/HW6_Slab_V4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="tDqNK9paSN_Y"
# # Saliency and Grad-CAM Examples
#
# [](https://colab.research.google.com/github/ShawnHymel/computer-vision-with-embedded-machine-learning/blob/master/2.3.1%20-%20CNN%20Visualizations/ei_saliency_and_grad_cam.ipynb)
#
# Import a trained model (.zip file) from Edge Impulse and copy in post-processed features of a single sample image. Run the notebook to see a saliency map and Grad-CAM heatmap.
#
# Saliency maps highlight which pixels in the input image were most important (i.e. most salient) in the decision making process.
#
# Grad-CAM looks at the output feature map of the final convolution layer to figure out which areas of the image were the most important in the decision making process.
#
# Author: EdgeImpulse, Inc.<br>
# Date: June 19, 2021<br>
# License: [Apache-2.0](apache.org/licenses/LICENSE-2.0)<br>
# + id="sGv808ddkd3P"
import numpy as np
import tensorflow as tf
from keras import activations, layers, models, backend
import cv2
import matplotlib.pyplot as plt
# + id="_FANkwFgQ6gi" colab={"base_uri": "https://localhost:8080/"} outputId="da014e78-9181-4618-a513-88548a745539"
### Unzip model
# Change this based on your .zip model filename
model_zip_path = "/content/ei-electronic-components-cnn-nn-classifier-tensorflow-savedmodel-model.zip"
# Unzip in place
# !unzip "{model_zip_path}"
# + id="0MdzYh9ZkmMB"
### Settings
# Image resolution
width = 28
height = 28
# Labels
labels = ["background", "capacitor", "diode", "led", "resistor"]
# Paste ground processed features from Edge Impulse and set ground-truth label
img = [0.5536, 0.5497, 0.5702, 0.5719, 0.5608, 0.5702, 0.5758, 0.5719, 0.5758, 0.5758, 0.5758, 0.5742, 0.5781, 0.5804, 0.5827, 0.5815, 0.5811, 0.5792, 0.5820, 0.5838, 0.5850, 0.5834, 0.5815, 0.5845, 0.5827, 0.5799, 0.5781, 0.5735, 0.5497, 0.5663, 0.5758, 0.5758, 0.5696, 0.5719, 0.5758, 0.5719, 0.5735, 0.5758, 0.5765, 0.5804, 0.5834, 0.5834, 0.5834, 0.5822, 0.5845, 0.5850, 0.5862, 0.5822, 0.5815, 0.5822, 0.5845, 0.5815, 0.5845, 0.5850, 0.5850, 0.5776, 0.5663, 0.5758, 0.5758, 0.5719, 0.5735, 0.5758, 0.5742, 0.5776, 0.5792, 0.5804, 0.5827, 0.5834, 0.5850, 0.5875, 0.5880, 0.5899, 0.5834, 0.5875, 0.5915, 0.5880, 0.5891, 0.5843, 0.5840, 0.5859, 0.5845, 0.5822, 0.5845, 0.5840, 0.5702, 0.5758, 0.5719, 0.5758, 0.5742, 0.5719, 0.5765, 0.5827, 0.5850, 0.5834, 0.5834, 0.5811, 0.5977, 0.6049, 0.6026, 0.5967, 0.5900, 0.5991, 0.6032, 0.5829, 0.5886, 0.5863, 0.5838, 0.5822, 0.5811, 0.5850, 0.5854, 0.5820, 0.5696, 0.5735, 0.5719, 0.5719, 0.5730, 0.5765, 0.5827, 0.5811, 0.5811, 0.5862, 0.5834, 0.5922, 0.6072, 0.6055, 0.6055, 0.6044, 0.6049, 0.6072, 0.6032, 0.5931, 0.5931, 0.5850, 0.5845, 0.5804, 0.5822, 0.5843, 0.5865, 0.5831, 0.5735, 0.5719, 0.5758, 0.5822, 0.5815, 0.5827, 0.5834, 0.5850, 0.5954, 0.6032, 0.6028, 0.6072, 0.6049, 0.6032, 0.6037, 0.6055, 0.6055, 0.6055, 0.6049, 0.6049, 0.6026, 0.5905, 0.5815, 0.5850, 0.5838, 0.5854, 0.5870, 0.5829, 0.5742, 0.5735, 0.5758, 0.5834, 0.5850, 0.5850, 0.5815, 0.5961, 0.6083, 0.6049, 0.6049, 0.6032, 0.6032, 0.6049, 0.6055, 0.6037, 0.6055, 0.6072, 0.6032, 0.6032, 0.6049, 0.5958, 0.5891, 0.5820, 0.5842, 0.5956, 0.5991, 0.5849, 0.5735, 0.5735, 0.5742, 0.5827, 0.5811, 0.5811, 0.5815, 0.5981, 0.6049, 0.6009, 0.6055, 0.6078, 0.6049, 0.6055, 0.6072, 0.6072, 0.6049, 0.6095, 0.6101, 0.6072, 0.6009, 0.6026, 0.6049, 0.5970, 0.5928, 0.6049, 0.6067, 0.5894, 0.5758, 0.5735, 0.5804, 0.5811, 0.5866, 0.5917, 0.5854, 0.5975, 0.6049, 0.6055, 0.6055, 0.6072, 0.6072, 0.6095, 0.6037, 0.6049, 0.6072, 0.6136, 0.6136, 0.6124, 0.6060, 0.6078, 0.6078, 0.6095, 0.6049, 0.6032, 0.6072, 0.5993, 0.5804, 0.5822, 0.5850, 0.5811, 0.5863, 0.6014, 0.6067, 0.6067, 0.6055, 0.6055, 0.6067, 0.6067, 0.6044, 0.6067, 0.6067, 0.6090, 0.6136, 0.6136, 0.6120, 0.6081, 0.6097, 0.6124, 0.6136, 0.6083, 0.6037, 0.6067, 0.6055, 0.6072, 0.5822, 0.5850, 0.5850, 0.5811, 0.5871, 0.6021, 0.6044, 0.5998, 0.5975, 0.6014, 0.5975, 0.5958, 0.5993, 0.6051, 0.5993, 0.6062, 0.6101, 0.6122, 0.6306, 0.6428, 0.6287, 0.6150, 0.6032, 0.6044, 0.6032, 0.6060, 0.6049, 0.6009, 0.5850, 0.5822, 0.5822, 0.5834, 0.5850, 0.5862, 0.6007, 0.6255, 0.6336, 0.6151, 0.6309, 0.6505, 0.6334, 0.6336, 0.6542, 0.6572, 0.6388, 0.6270, 0.5476, 0.4665, 0.5099, 0.5876, 0.6102, 0.6032, 0.6044, 0.6067, 0.6067, 0.5986, 0.5799, 0.5781, 0.5827, 0.5850, 0.5815, 0.5862, 0.5517, 0.3781, 0.2192, 0.3420, 0.4005, 0.4060, 0.4991, 0.5137, 0.4310, 0.4207, 0.4463, 0.3916, 0.2547, 0.0898, 0.2204, 0.3795, 0.5794, 0.6131, 0.6062, 0.6078, 0.6055, 0.6009, 0.5757, 0.5780, 0.5803, 0.5820, 0.5805, 0.6075, 0.3876, 0.1835, 0.0399, 0.2985, 0.2197, 0.0139, 0.1731, 0.3261, 0.1489, 0.1767, 0.3422, 0.3198, 0.3476, 0.5168, 0.5582, 0.3599, 0.4792, 0.6490, 0.6372, 0.6266, 0.6157, 0.6153, 0.6008, 0.5988, 0.5937, 0.5815, 0.5922, 0.5418, 0.3256, 0.2231, 0.1299, 0.3781, 0.4072, 0.3059, 0.3375, 0.3961, 0.2047, 0.2526, 0.3623, 0.3195, 0.3432, 0.4702, 0.5333, 0.3945, 0.3316, 0.4202, 0.4269, 0.5429, 0.6038, 0.6160, 0.3374, 0.3295, 0.3386, 0.3707, 0.3733, 0.2852, 0.2560, 0.2133, 0.0585, 0.2619, 0.1949, 0.0106, 0.1203, 0.3086, 0.1467, 0.1673, 0.3037, 0.2887, 0.2583, 0.1448, 0.2198, 0.3257, 0.3154, 0.4693, 0.4956, 0.5046, 0.5136, 0.5140, 0.3459, 0.3465, 0.3447, 0.3461, 0.3315, 0.2495, 0.2328, 0.1714, 0.0491, 0.2280, 0.2032, 0.0356, 0.1313, 0.2869, 0.1546, 0.1676, 0.2735, 0.2596, 0.2374, 0.1442, 0.2082, 0.3044, 0.3166, 0.4060, 0.4084, 0.3964, 0.3924, 0.3882, 0.4320, 0.4375, 0.4345, 0.4321, 0.4292, 0.3104, 0.2058, 0.1785, 0.0831, 0.2038, 0.2002, 0.0634, 0.1012, 0.1788, 0.1286, 0.1219, 0.1668, 0.1766, 0.2123, 0.1878, 0.1933, 0.2740, 0.5056, 0.5809, 0.5736, 0.5758, 0.5742, 0.5726, 0.6038, 0.6038, 0.6072, 0.6065, 0.6158, 0.5636, 0.2647, 0.1153, 0.0912, 0.1179, 0.1702, 0.1787, 0.2035, 0.2151, 0.2285, 0.2401, 0.2233, 0.1741, 0.1466, 0.1593, 0.2002, 0.3985, 0.6271, 0.6503, 0.6452, 0.6270, 0.6203, 0.6215, 0.5783, 0.5783, 0.5806, 0.5819, 0.5819, 0.5877, 0.5240, 0.3872, 0.3661, 0.4013, 0.5094, 0.5794, 0.5998, 0.6111, 0.6070, 0.6139, 0.6102, 0.5721, 0.5139, 0.5248, 0.5613, 0.6043, 0.6180, 0.6284, 0.6268, 0.6124, 0.6124, 0.6124, 0.5838, 0.5838, 0.5834, 0.5850, 0.5815, 0.5896, 0.6131, 0.6318, 0.6329, 0.6341, 0.6309, 0.6169, 0.6125, 0.6213, 0.6187, 0.6203, 0.6234, 0.6278, 0.6340, 0.6340, 0.6296, 0.6175, 0.6136, 0.6136, 0.6136, 0.6147, 0.6118, 0.6078, 0.5799, 0.5850, 0.5850, 0.5811, 0.5850, 0.6032, 0.6032, 0.6005, 0.6097, 0.6097, 0.6074, 0.6090, 0.6099, 0.6173, 0.6101, 0.6124, 0.6236, 0.6215, 0.6167, 0.6136, 0.6148, 0.6136, 0.6141, 0.6164, 0.6164, 0.6129, 0.6136, 0.6118, 0.5735, 0.5799, 0.5850, 0.5811, 0.5836, 0.5998, 0.6072, 0.6055, 0.6078, 0.6083, 0.6055, 0.6072, 0.6096, 0.6115, 0.6060, 0.6118, 0.6159, 0.6136, 0.6129, 0.6160, 0.6164, 0.6164, 0.6136, 0.6141, 0.6164, 0.6124, 0.6095, 0.6072, 0.5742, 0.5735, 0.5804, 0.5834, 0.5827, 0.5878, 0.5942, 0.6072, 0.6095, 0.6032, 0.6049, 0.6037, 0.6067, 0.6067, 0.6044, 0.6106, 0.6152, 0.6106, 0.6101, 0.6141, 0.6141, 0.6141, 0.6152, 0.6129, 0.6129, 0.6118, 0.6037, 0.6049, 0.5719, 0.5719, 0.5758, 0.5827, 0.5845, 0.5788, 0.5822, 0.5903, 0.5998, 0.6072, 0.6032, 0.6055, 0.6072, 0.6060, 0.6037, 0.6067, 0.6072, 0.6049, 0.6101, 0.6147, 0.6141, 0.6106, 0.6136, 0.6164, 0.6072, 0.6037, 0.6072, 0.6072, 0.5758, 0.5758, 0.5719, 0.5742, 0.5769, 0.5758, 0.5804, 0.5811, 0.5862, 0.5926, 0.6026, 0.6072, 0.6032, 0.6044, 0.6055, 0.6049, 0.6072, 0.6032, 0.6049, 0.6118, 0.6147, 0.6090, 0.6049, 0.6078, 0.6055, 0.6049, 0.6055, 0.6049, 0.5592, 0.5719, 0.5758, 0.5735, 0.5742, 0.5723, 0.5758, 0.5827, 0.5742, 0.5776, 0.5926, 0.6032, 0.6072, 0.6037, 0.6072, 0.6055, 0.6055, 0.6049, 0.6009, 0.6032, 0.6095, 0.6095, 0.6032, 0.6049, 0.6072, 0.6055, 0.6049, 0.6072, 0.5497, 0.5536, 0.5608, 0.5719, 0.5758, 0.5719, 0.5758, 0.5811, 0.5781, 0.5746, 0.5811, 0.5905, 0.6014, 0.6032, 0.6055, 0.6049, 0.6032, 0.6049, 0.6032, 0.6055, 0.6049, 0.6060, 0.6072, 0.6009, 0.6032, 0.6072, 0.6055, 0.6049]
true_idx = 4 # Resistor
# You probably don't need to change this
model_dir = "/content/saved_model"
# + colab={"base_uri": "https://localhost:8080/"} id="CDqyA0XRknj6" outputId="b9f6416e-415a-4883-a242-0f69c0187bd2"
### Load model file
model = tf.keras.models.load_model(model_dir)
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="CC8OsO09k0z1" outputId="5ab90d5f-4b74-413c-9dc6-0cc14cff381d"
### Reshape image
# Convert features to 2D Numpy array
img_reshape = np.reshape(np.array(img), (height, width))
# Show the ground-truth label
print("Actual label:", labels[true_idx])
# Display image (make sure we're looking at the right thing)
plt.imshow(img_reshape, cmap='gray', vmin=0.0, vmax=1.0)
# + colab={"base_uri": "https://localhost:8080/"} id="kJaVVmkNqgAj" outputId="5b7040e6-9941-49cc-c98c-298c2c39d639"
### The Keras model expects images in a 4D array with dimensions (sample, height, width, channel)
# Add extra dimension to the image (placeholder for color channels)
img_0 = img_reshape.reshape(img_reshape.shape + (1,))
# Keras expects more than one image (in Numpy array), so convert image(s) to such array
images = np.array([img_0])
# Print dimensions of inference input
print(images.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="V430dQ8Wp_b0" outputId="e99e53f0-9ff0-40d1-aa40-e359b5b06b04"
### Do a forward pass (inference) with the test image and print the predicted probabilities
# Inference
preds = model.predict(images)
# Print out predictions
for i, pred in enumerate(preds[0]):
print(labels[i] + ": " + str(pred))
# + id="qgTp5vRMqOXD"
### For either algorithm, we need to remove the Softmax activation function of the last layer
model.layers[-1].activation = None
# + [markdown] id="5PIMyTWzHGfA"
# ## Saliency Map
# + id="-jhNP53mHJBv"
### Based on: https://github.com/keisen/tf-keras-vis/blob/master/tf_keras_vis/saliency.py
def get_saliency_map(img_array, model, class_idx):
# Gradient calculation requires input to be a tensor
img_tensor = tf.convert_to_tensor(img_array)
# Do a forward pass of model with image and track the computations on the "tape"
with tf.GradientTape(watch_accessed_variables=False, persistent=True) as tape:
# Compute (non-softmax) outputs of model with given image
tape.watch(img_tensor)
outputs = model(img_tensor, training=False)
# Get score (predicted value) of actual class
score = outputs[:, true_idx]
# Compute gradients of the loss with respect to the input image
grads = tape.gradient(score, img_tensor)
# Finds max value in each color channel of the gradient (should be grayscale for this demo)
grads_disp = [np.max(g, axis=-1) for g in grads]
# There should be only one gradient heatmap for this demo
grad_disp = grads_disp[0]
# The absolute value of the gradient shows the effect of change at each pixel
# Source: https://christophm.github.io/interpretable-ml-book/pixel-attribution.html
grad_disp = tf.abs(grad_disp)
# Normalize to between 0 and 1 (use epsilon, a very small float, to prevent divide-by-zero error)
heatmap_min = np.min(grad_disp)
heatmap_max = np.max(grad_disp)
heatmap = (grad_disp - heatmap_min) / (heatmap_max - heatmap_min + tf.keras.backend.epsilon())
return heatmap.numpy()
# + id="Jv3ZPnM5HRwg"
### Generate saliency map for the given input image
saliency_map = get_saliency_map(images, model, true_idx)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="55N3wuvNNn5z" outputId="07e6e3ff-c708-4f0f-bcf1-c57d026c434d"
### Draw map
plt.imshow(saliency_map, cmap='jet', vmin=0.0, vmax=1.0)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="MYGRdsbZQfKH" outputId="fadb2e08-ee5f-41d6-c851-64be6b8880be"
### Overlay the saliency map on top of the original input image
idx = 0
ax = plt.subplot()
ax.imshow(images[idx,:,:,0], cmap='gray', vmin=0.0, vmax=1.0)
ax.imshow(saliency_map, cmap='jet', alpha=0.25)
# + [markdown] id="__J5ufo7m_-q"
# ## Grad-CAM
# + id="o462piKhlDES"
### This function comes from https://keras.io/examples/vision/grad_cam/
def make_gradcam_heatmap(img_array, model, last_conv_layer_name, pred_index=None):
# First, we create a model that maps the input image to the activations
# of the last conv layer as well as the output predictions
grad_model = tf.keras.models.Model(
[model.inputs], [model.get_layer(last_conv_layer_name).output, model.output]
)
# Then, we compute the gradient of the top predicted class for our input image
# with respect to the activations of the last conv layer
with tf.GradientTape() as tape:
last_conv_layer_output, preds = grad_model(img_array)
if pred_index is None:
pred_index = tf.argmax(preds[0])
class_channel = preds[:, pred_index]
# This is the gradient of the output neuron (top predicted or chosen)
# with regard to the output feature map of the last conv layer
grads = tape.gradient(class_channel, last_conv_layer_output)
# This is a vector where each entry is the mean intensity of the gradient
# over a specific feature map channel
pooled_grads = tf.reduce_mean(grads, axis=(0, 1, 2))
# We multiply each channel in the feature map array
# by "how important this channel is" with regard to the top predicted class
# then sum all the channels to obtain the heatmap class activation
last_conv_layer_output = last_conv_layer_output[0]
heatmap = last_conv_layer_output @ pooled_grads[..., tf.newaxis]
heatmap = tf.squeeze(heatmap)
# The absolute value of the gradient shows the effect of change at each pixel
# Source: https://christophm.github.io/interpretable-ml-book/pixel-attribution.html
heatmap = tf.abs(heatmap)
# Normalize to between 0 and 1 (use epsilon, a very small float, to prevent divide-by-zero error)
heatmap_min = np.min(heatmap)
heatmap_max = np.max(heatmap)
heatmap = (heatmap - heatmap_min) / (heatmap_max - heatmap_min + tf.keras.backend.epsilon())
return heatmap.numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="3CLU0fKpoSvv" outputId="163c6276-8990-4d32-cede-f7a8bd23e1a1"
### We need to tell Grad-CAM where to find the last convolution layer
# Print out the layers in the model
for layer in model.layers:
print(layer, layer.name)
# Go backwards through the model to find the last convolution layer
last_conv_layer = None
for layer in reversed(model.layers):
if 'conv' in layer.name:
last_conv_layer = layer.name
break
# Give a warning if the last convolution layer could not be found
if last_conv_layer is not None:
print("Last convolution layer found:", last_conv_layer)
else:
print("ERROR: Last convolution layer could not be found. Do not continue.")
# + id="U1fxqDUjnK4K"
### Generate class activation heatmap
heatmap = make_gradcam_heatmap(images, model, last_conv_layer)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="VaMUlMGYoNYi" outputId="05a7dde4-beb7-401e-be35-e6a98714fac5"
### Draw map
plt.imshow(heatmap, cmap='jet', vmin=0.0, vmax=1.0)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="lpBNKFivsKij" outputId="2cf2e9f6-f91a-4a19-935a-befea7bed6bc"
### Overlay the saliency map on top of the original input image
# The heatmap is a lot smaller than the original image, so we upsample it
big_heatmap = cv2.resize(heatmap, dsize=(height, width), interpolation=cv2.INTER_CUBIC)
# Draw original image with heatmap superimposed over it
idx = 0
ax = plt.subplot()
ax.imshow(images[idx,:,:,0], cmap='gray', vmin=0.0, vmax=1.0)
ax.imshow(big_heatmap, cmap='jet', alpha=0.25)
# + id="nYaOQKhPX4cb"
|
2.3.1 - CNN Visualizations/ei_saliency_and_grad_cam.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tsp_deep_rl
# language: python
# name: conda_tsp_deep_rl
# ---
# +
# %load_ext nb_black
import sagemaker
import boto3
# -
# # 1. Prepare Training data
# + language="bash"
# if [ -f ../data/tsp-data.tar.gz ]; then
# echo "File tsp-data.tar.gz exists."
# else
# echo "File tsp-data.tar.gz does not exist."
# gdown https://drive.google.com/uc?id=152mpCze-v4d0m9kdsCeVkLdHFkjeDeF5
# mv tsp-data.tar.gz ../
# fi
# + language="bash"
# if [ -d ../data ]; then
# echo "Folder data exists."
# else
# echo "Folder data does not exist."
# tar -xvzf ../tsp-data.tar.gz -C ../
# fi
# -
session = sagemaker.Session()
BUCKET = session.default_bucket() # Set a default S3 bucket
s3 = boto3.resource("s3")
for file in [
"tsp20_test_concorde.txt",
"tsp50_test_concorde.txt",
"tsp100_test_concorde.txt",
]:
s3.meta.client.upload_file(f"../data/tsp/{file}", BUCKET, f"data/tsp/{file}")
# # 2. Distributed Training
sagemaker_session = sagemaker.Session()
role = sagemaker.get_execution_role()
role_name = role.split(["/"][-1])
print(f"The Amazon Resource Name (ARN) of the role used for this demo is: {role}")
print(f"The name of the role used for this demo is: {role_name[-1]}")
# +
from sagemaker.pytorch import PyTorch
estimator = PyTorch(
base_job_name="pytorch-smdataparallel-tsp",
source_dir="../src",
entry_point="run.py",
role=role,
framework_version="1.8.1",
py_version="py36",
instance_count=1,
instance_type="ml.p3.16xlarge",
sagemaker_session=sagemaker_session,
distribution={"smdistributed": {"dataparallel": {"enabled": True}}},
debugger_hook_config=False,
hyperparameters={
"problem": "tsp",
"min_size": 50,
"max_size": 50,
"neighbors": 0.2,
"knn_strat": "percentage",
"n_epochs": 100,
"epoch_size": 128000,
"batch_size": 128,
"accumulation_steps": 1,
"train_dataset": "tsp20-50_train_concorde.txt",
"val_datasets": "tsp20_test_concorde.txt tsp50_test_concorde.txt tsp100_test_concorde.txt",
"val_size": 1280,
"rollout_size": 1280,
"model": "attention",
"encoder": "gnn",
"embedding_dim": 128,
"hidden_dim": 512,
"n_encode_layers": 3,
"aggregation": "max",
"normalization": "batch",
"n_heads": 8,
"tanh_clipping": 10.0,
"lr_model": 0.0001,
"lr_critic": 0.0001,
"lr_decay": 1.0,
"max_grad_norm": 1.0,
"exp_beta": 0.8,
"baseline": "rollout",
"bl_alpha": 0.05,
"bl_warmup_epochs": 0,
"seed": 1234,
"num_workers": 0,
"log_step": 100,
},
metric_definitions=[
{
"Name": "val:gap_tsp20",
"Regex": "tsp20_test_concorde.txt Validation optimality gap=(.*?)\%",
},
{
"Name": "val:gap_tsp50",
"Regex": "tsp50_test_concorde.txt Validation optimality gap=(.*?)\%",
},
{
"Name": "val:gap_tsp100",
"Regex": "tsp100_test_concorde.txt Validation optimality gap=(.*?)\%",
},
],
max_run=1 * 24 * 60 * 60,
)
# -
estimator.fit(
{"train": f"s3://{BUCKET}/data/tsp", "val": f"s3://{BUCKET}/data/tsp"}, wait=False
)
|
notebooks/pytorch_training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AMoC Hackathon Tasks 2021
#
#
# # 1. Longitudinal Visualisation Task
#
#
# Darknet markets remain a challenging issue in terms of monitoring and mitigating their practices, for example: buying and selling of drugs worldwide. While research often analyses these markets, there remains a gap in understanding the impacts of adverse events in darknet markets following, for example, DDoS attacks or targeted site take downs.
#
# In this task we are looking for an innovative visualisation representing the datasets over time, with particular attention to how significant events might affect the nature of posting on the forums.
#
#
# ## 1.1. Data
#
# For this task we make use of over 2.5 million posts drawn from over 100,000 users from 40 cybercriminal communities, drawn from a large dataset collected between 2013 and 2015. In particular, we targeted discussion
# forums within this collection, which acted as support areas for underground marketplaces dealing in a number of different illicit goods. Communities ranged from successfully established markets with thousands of users (though not all were always active posters) to small sites that never moved beyond a handful of initial users.
#
# ## 1.2. Instructions for Accessing the Data
#
# The "Fora" folder has 40 csv-files, each containing data from a specific DNM forum. After loading a target forum file, the community name, user_id, threat_id, date, subject, category, body and quotes can be accessed.
#
# ### To access a user's metadata and his/her messages posted on the targeted forum file, the code snippet below can be used.
import pandas as pd
df = pd.read_csv("../data/hackathon_dataset/fora/abraxas.csv") # because the first positive user it's from Abraxas Forums
print(len(df))
print(df.columns.values.tolist())
# ### Additionally, each user's registered information (community, user_id, title, first_seen) can be accessed in the "measure_impact/fora_registedusers.csv" using the code below.¶
# +
reg_users = pd.read_csv("../data/hackathon_dataset/measure_impact/fora_registedusers.csv")
train_positive_users = pd.read_csv("../data/hackathon_dataset/reidentification/train_positive_users.csv", header=None)
user_df = reg_users[reg_users['user_id'].str.match( train_positive_users.iloc[0][1] )]
print(user_df)
# -
# ## 1.3. Evaluation
#
# The panel will evaluate the results for this task against the following qualitative criteria:
# - Suitability of visualization chosen (30%)
# - Effectiveness of visualization in communicating long term trends (30%)
# - Ability to understand multiple facets of the longitudinal data (20%)
# - Quality of the submitted code (20%)
#
#
# ## 1.4. Code Submission
#
# We kindly ask you to submit the following:
# - the software you built for this task;
# - a README file containing requirements and external resources needed to run your system.
#
# You can choose freely among the available programming languages and among the operating systems. Please upload your software to the "Outputs" folder created for your team in Microsoft Teams.
#
# ### Deadline for submission is Wednesday 10 February 2021 - 17:00 GMT.
#
# ### Note: By submitting your software you agree to make your code available under CC-BY-NC license for use by researchers, including the AMoC team.
#
#
# # 2. DNM User Re-identification Task
#
# In recent years, Darknet Markets (DNMs) and other environments offering anonymity are becoming increasingly popular among criminals with a high degree of computer literacy and forensic awareness.
# Although none of such anonymisation techniques is entirely bulletproof, they can easily complicate or even block cybercrime investigations by law enforcement. In such cases, the communications produced on such underground forums can be one of few clues to a cyber offender’s identity.
#
# In this task we are looking for novel approaches to automatically re-identify cyber offenders using multiple identities across different underground platforms. More specifically:
# - given a large training dataset comprising of DNM users of whom “ground truth” information is available on the different identities they have on one or more Darknet forums, their communications and metadata,
# - design a system that is able to group all identities (one or more) of each user in a new dataset (the test dataset).
#
# ## 2.1. Data
#
# For this task we make use of over 10,000 users from different cybercriminal communities who produced a minimum of 5 messages, drawn from the dataset described above. The training dataset provided for this task contains two files:
# - train_positive_users.csv – contains the users who have more than one identity. Each line represents a new user with his/her matching identities.
# - train_negative_users.csv – contains the users who have only one identity in this dataset.
#
# The test dataset ("reidentification/test_dataset.csv") contains about 3,000 users without any "ground truth" information about multiple user identities. None of the users from the training dataset is included in the test dataset. Additionally, as the training dataset is highly skewed, we ensured that the test dataset shows a similar distribution of positive and negative users as the training dataset.
#
#
# ## 2.2. Instructions for Accessing the Data
#
# ### Load positive and negative users for the train dataset from train_positive_users.csv and test_dataset.csv, respectively.
train_positive_users = pd.read_csv("../data/hackathon_dataset/reidentification/train_positive_users.csv", header=None)
train_negative_users = pd.read_csv("../data/hackathon_dataset/reidentification/train_negative_users.csv", header=None)
# #### Within the positive users dataset, each list of multiple pairs indicates the different user-ids used in different communities, but they are actually the same user, and we labeled it as 1.
print(len(train_positive_users))
print(train_positive_users.iloc[0])
# #### The negative users dataset contains a list of pairs of community- and user-ids. None of these users have more than one identity in this dataset (as far as we know).
print(len(train_negative_users))
print(train_negative_users.head(10))
# #### To access a user's metadata and his/her messages posted on the targeted forum file, please see the code snippet provided above. To select a target user's relevant information from a community, especially posted messages in the 'body' column, the following code can be used.
selected_df = df[df['user_id'].str.match( train_positive_users.iloc[0][1] )]
print(len(selected_df))
print(selected_df['body'][:5])
# ## 2.3. Evaluation
#
# Once you have developed your approach for this task, your software can be tested on the test dataset ("reidentification/test_dataset.csv"). After the hackathon, the ground truth labels for the test dataset will be made available.
#
# To evaluate your systems’ performance, we will calculate mean average precision, recall and F1 score based on the labels produced by your system on the test dataset.
#
# # 3. Output and Code Submission
#
# For this task, we ask you to submit the following:
# - one csv-file containing all positive users and one csv-file with all negative users of the test dataset, similar to the train_positive_users.csv and train_negative_users.csv provided for this task;
# - the software you built for this task;
# - a README file containing requirements and external resources needed to run your system.
#
# Again, you can choose freely among the available programming languages and among the operating systems. Please upload your csv-files and software to the "Outputs" folder created for your team in Microsoft Teams.
#
# ### Deadline for submission is Wednesday 10 February 2021 - 17:00 GMT.
#
# ### Note: By submitting your software you agree to make your code available under CC-BY-NC license for use by researchers, including the AMoC team.
#
#
#
# # 4. Report
#
# Finally, we ask you to submit an overview of your approach to solving both AMoC Hackathon tasks and your results in a short report (up to 4 pages) using the IEEE conference paper templates that can be accessed here: https://www.ieee.org/conferences/publishing/templates.html. Please upload your report to the "Outputs" folder created for your team in Microsoft Teams.
#
# ### Deadline for submitting the report is Thursday 11 February 2021 - 17:00 GMT.
#
#
# # 5. Good Luck!
#
# The panel will meet on Tuesday 16 February 2021 to deliberate on which team should win the prize. The results will be communicated to all teams via email following the panel meeting.
|
notebooks/hackathon_tasks_overview.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logistic and Linear Regression
#
# In this notebook we look at plotting 1D, 2D and k-D cases of Linear and Logistic Regression models, drawing from statistical distributions.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import pandas as pd
from scipy import stats
from sklearn.linear_model import LogisticRegression, LinearRegression
from sklearn.decomposition import PCA
from sklearn.metrics import confusion_matrix
from scipy.special import expit
# -
# ## Discrete 1d case
# +
Xd1 = np.random.normal(loc=2, scale=2, size=(100,))
yd1 = pd.cut(stats.zscore(Xd1), [-np.inf, 0, np.inf], labels=[0, 1])
lr = LogisticRegression().fit(np.atleast_2d(Xd1).T, yd1)
plt.scatter(Xd1, yd1, color='k', label='actual')
plt.scatter(Xd1, lr.predict_proba(np.atleast_2d(Xd1).T)[:, 1],
marker='x', c=yd1, label='predictions')
X_t = np.linspace(-5, 8, 400)
y_t = expit(X_t * lr.coef_[0] + lr.intercept_)
plt.plot(X_t, y_t, 'r-', label=r'logit($X\beta$)')
plt.xlabel(r"$x$")
plt.ylabel(r"$y$")
plt.legend()
plt.show()
# -
# ## Discrete 2d case
# +
Xd2 = np.random.normal(loc=2, scale=2, size=(100,2))
yd2 = pd.cut(stats.zscore(np.sum(Xd2, 1)), [-np.inf, 0, np.inf], labels=[0, 1])
lr2 = LogisticRegression().fit(Xd2, yd2)
plt.scatter(Xd2[:, 0], Xd2[:, 1], alpha=.5, c=lr2.predict_proba(Xd2)[:,1], label='predictions')
plt.scatter(Xd2[:, 0], Xd2[:, 1], c=yd2, marker='x', label='actual')
G = np.mgrid[-5:8:100j, -5:8:100j]
yp = expit(np.dot(G.T, lr2.coef_.T) + lr2.intercept_).reshape(100, 100)
plt.contour(G[0], G[1], yp)
plt.xlabel(r"$x_1$")
plt.ylabel(r"$x_2$")
plt.colorbar()
plt.legend(loc='best')
plt.show()
# -
# ## Discrete $k$-case
#
# Here we use `PCA` to compress to 2d space for visualisation
# +
Xdk = np.random.normal(loc=4.2, scale=1.5, size=(100,5))
ydk = pd.cut(stats.zscore(np.sum(Xdk, 1)), [-np.inf, 0, np.inf], labels=[0, 1])
pca_dk = PCA(2).fit_transform(Xdk)
lrk = LogisticRegression().fit(pca_dk, ydk)
plt.scatter(pca_dk[:, 0], pca_dk[:, 1], alpha=.5,
c=lrk.predict_proba(pca_dk)[:,1], label='predictions')
plt.scatter(pca_dk[:, 0], pca_dk[:, 1], c=ydk, marker='x', label='actual')
G = np.mgrid[-5:7:200j, -5:7:200j]
yp = expit(np.dot(G.T, lrk.coef_[0]) + lrk.intercept_).reshape(200, 200)
plt.contour(G[0], G[1], yp)
plt.legend()
plt.xlabel("PC1")
plt.ylabel("PC2")
plt.show()
# -
# ## Continuous 1d case
# +
Xc1 = np.random.normal(loc=3, scale=2, size=(200,))
yc1 = Xc1*1.5 - 2. + np.random.normal(loc=0, scale=1.5, size=(200,))
lm1 = LinearRegression().fit(np.atleast_2d(Xc1).T, yc1)
plt.scatter(Xc1, yc1, color='k', alpha=.5)
plt.scatter(Xc1, lm1.predict(np.atleast_2d(Xc1).T), marker='x', color='r')
X_t = np.linspace(-5, 10, 400)
plt.plot(X_t, X_t*lm1.coef_ + lm1.intercept_, 'r-')
print(lm1.coef_, lm1.intercept_)
plt.xlabel(r"$x$")
plt.ylabel(r"$y$")
plt.show()
# -
# ## Continuous 2d case
# +
Xc2 = np.random.normal(loc=3, scale=2, size=(200,2))
yc2 = np.sum(Xc2,1)*1.5 + 2. + np.random.normal(loc=0, scale=1., size=(200,))
lm2 = LinearRegression().fit(Xc2, yc2)
plt.scatter(Xc2[:, 0], Xc2[:, 1], c=lm2.predict(Xc2), alpha=.5, label='predictions')
plt.scatter(Xc2[:, 0], Xc2[:, 1], c=yc2, marker='x', label='actual')
G = np.mgrid[-5:10:100j, -5:10:100j]
yp = (np.dot(G.T, lm2.coef_.T) + lm2.intercept_).reshape(100, 100)
plt.contour(G[0], G[1], yp)
plt.colorbar()
plt.xlabel(r"$x_1$")
plt.ylabel(r"$x_2$")
plt.legend()
plt.show()
# -
# ## Continuous $k$-case
# +
Xc3 = np.random.normal(loc=2, scale=2, size=(200,5))
yc3 = np.sum(Xc3,1)*2. - 2. + np.random.normal(loc=0, scale=2.5, size=(200,))
lm3 = LinearRegression().fit(Xc3, yc3)
plt.scatter(yc3, lm3.predict(Xc3), color='r', label='points', alpha=.5)
plt.plot([-10, 45], [-10, 45], 'k--', label=r"$y=x$")
plt.xlabel(r"$y$")
plt.ylabel(r"$\hat{y}$")
plt.legend()
plt.show()
|
Extras/Linear-and-Logistic-Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="S6qNJ_JSlErv"
# # Lab 00 - A - Data Analysis In Python - First Steps
#
# Machine learning and data analysis (that is usually also a pre-step for learning) deal with data. Thererefore we need tools to manipulate it and extract the information we want. In the Python environment there are two very useful packages for this: [`numpy`](https://numpy.org/doc/1.19/) and [`pandas`](https://pandas.pydata.org/docs/reference/index.html#api).
#
# In this lab, we will take the first steps into using these packages and see some of their functionalities. These will be needed throughout the course. Both packages contain many useful functionalities, of which we will introduce only a few. To find out more about the other functionalities, the documentations, **Google** and **StackOverflow** are your best friends.
# + id="Hx4v3hy-fnu9"
# Load commonly used imports (such as numpy and pandas) and several utils functions that
# are used thoughout different labs and code examples
import sys
sys.path.append("../")
from utils import *
# + [markdown] id="asLx3ujI09-B"
# # Numpy - The Basics
#
# Let us start with numpy. This package supports vector, matrix and tensor operations over numerical (but not just) data. It is very comfortable and much faster than `for` loops and classic `list` manipulations.
# + [markdown] id="fPwJrL1Bc84L"
# ## Array Creation
#
# There are multiple ways to create an array. We can create it from an existing list, load it from a file or generate a new array.
#
# + colab={"base_uri": "https://localhost:8080/"} id="wL4cbVc-pGZ9" outputId="e5e90a34-4000-4d85-8672-7559f6ecaf6d"
array_1D = np.array([6, 2, 8, 4, 5, 10, 7, 143, 9, 10, 11])
print(array_1D)
print(array_1D.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="Gt6ZB5FG1p2d" outputId="b3f5bdbf-9547-485a-bff1-29e07aecc6aa"
array_2D = np.array(
[[10, 20, 30, 40],
[100, 200, 300, 400],
[1000, 2000, 3000, 4000]])
print(array_2D)
print(array_2D.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="ICxIiGg92M7C" outputId="5c5f5333-af82-4e07-f836-202fddf7a8fa"
array_3D = np.array(
[[[10, 20, 30, 40], [100, 200, 300, 400], [1000, 2000, 3000, 4000]],
[[11, 21, 31, 41], [101, 201, 301, 401], [1001, 2001, 3001, 4001]]])
print(array_3D)
print(array_3D.shape)
# + [markdown] id="u8ikdeRnpupl"
# Using `numpy`'s functions for creating new arrays requires specifying the shape of the desired output array. This is an n-array tuple specifying the sizes of the different dimensions.
#
# * Specifying the shape `(3)` will create a 1D array with 3 entries.
# * Specifying the shape `(10, 3)` will create a 2D matrix with 10 rows and 3 columns.
# * Specifying the shape `(10, 28, 28)` will create a 3D matrix (a tensor) which we can think of in the following manner: it is an object holding 10 2D matrices of size 28x28.
#
#
#
#
# + pycharm={"name": "#%%\n"}
# Initalize arrays with built-in numpy functions
zeros_3D = np.zeros((4, 5, 2)) # Create a 3D arrays of 0's
ones_2D = np.ones((4, 5)) # Create a 2D arrays of 1's
print(zeros_3D)
print(ones_2D)
# + colab={"base_uri": "https://localhost:8080/"} id="3ke7ltD8qt8I" outputId="9de4ca81-d090-47e7-bbad-a7cd9dea2d5f"
np.arange(50) # Create the vector [0, 1, 2, 3, 4, .. , 48, 49]
# + colab={"base_uri": "https://localhost:8080/"} id="g8tf4cd4q4oE" outputId="91be65fe-ab25-455b-9fc8-8a455b07104c"
# Create a vector of random integers from 5 to 50, with shape (2, 3)
np.random.randint(5, 50, (2, 3))
# + [markdown] id="THE5lTeTrN4w"
# There are many other functions such as `np.full`, `np.eye`, `np.random.uniform`, etc. Next, let us load an existsing dataset into a numpy array. This specific dataset represents images.
# + colab={"base_uri": "https://localhost:8080/", "height": 197} id="O3ZsPny03qmZ" outputId="3d22fab1-7cb1-4180-a1c2-a5c1528f6de5"
img_array = np.loadtxt(open("../datasets/MNIST_Images.csv", "rb"), delimiter=",").reshape(-1, 28, 28)
img_array.shape
# -
px.imshow(img_array[0]).show()
# + [markdown] id="F10jW-AO8PdQ"
# ## Array indexing and slicing
#
# A great strength of `numpy` is the ease in subsetting an array to retrieve only specific parts of it. We do so by indexing and slicing the arrays. For 1D arrays these operations are very similar to those over lists. For arrays of higher dimensions, we use a comma to separate the slicing of each dimension. For example, accessing an element in the array `arr` in the first row and second column is done by: `arr[0, 1]` (recall indexing in python begins from zero). To select only the second column, over all rows write: `arr[:, 2]`
# + id="KZJfKZ-_67Yk"
array_1D = np.array([10, 11, 12, 13, 14, 15, 16])
# Select 1st element
print("Select 1st element")
print(array_1D[1])
# Select all the elements from 1st to 4th element
print("\nSelect all the elements from 1st to 4th element")
print(array_1D[1:5])
# Select elements [1, 4]
print("\nSelect elements [1, 4]")
print(array_1D[[1, 4]])
array_2D = np.array([[1, 2, 3, 4],
[5, 6, 7, 8],
[9, 10, 11, 12],
[13, 14, 15, 16],
[17, 18, 19, 20]])
# Select 1st row, 2nd column
print(array_2D[1, 2])
randint_2D = np.random.randint(5, 50, (10, 20))
print("\nPrint random array")
print(randint_2D)
# Select from 3rd row to 5th, all the columns
print("\nSelect from 3rd row to 5th, all the columns")
print(randint_2D[2:5, :])
# Select from 3rd row to 5th, columns 1 and 2
print("\nSelect from 3rd row to 5th, columns 1 and 2")
print(randint_2D[2:5, 1:3])
# Select from 3d row to 5th, columns 3, 5, 6, and 11
print("\nSelect from 3rd row to 5th, columns 3, 5, 6, and 11")
print(randint_2D[2:5, [3, 5, 6, 11]])
# + [markdown] id="kn4PwoBerrJl"
# ## Matrix Operations
# Another strength of the `numpy` package is that all matrix operations you can think of (and even more) are already implemented. For example, element-wise addition of scalar, multiplication, powering up a matrix, log-transformations and much more.
#
# As `numpy` overloads the different mathematical operators, it is easy to write mathematical expressions over 2 (or more) vectors/matrices, such as summing or multiplying and also comparing their elements.
#
# + colab={"base_uri": "https://localhost:8080/"} id="N4OXpnPKiLbg" outputId="9d066e26-47b4-4db3-971d-5e3c4ae6a1d0"
# Operations on 1 matrix
A = np.arange(1, 33).reshape(4, 8) # Create an array of numbers from 1 ot 32, and then make a 2d array of 4 rows to 8 columns
print(A)
print("\n\nA + 1:")
print(A + 1)
print("\n2 * A:")
print(2 * A)
print("\nA*A*A:")
print(np.power(A, 3))
print("\nlog(A):")
print(np.log(A))
print("\nA Transpose:")
print(A.transpose) # also np.transpose(A) and A.T are valid syntaxes
print("\n A > 10:")
print(A > 10)
# Operations on 2 matrices
A = np.arange(6).reshape([2, 3])
B = np.random.randint(1, 10, (2, 3))
print("A")
print(A)
print("\n\nB")
print(B)
print("\nA+B:")
print(A + B) # Equivalent to np.add(a_array, b_array)
print("\n A * B (element-wise multiplication):")
print(np.multiply(A, B))
print("\n AB (matrix multiplication):")
print(A @ B.T) # Equivalent to np.dot(a_array, b_array)
print("\nA > B")
print(A > B)
# + [markdown] id="WBGJkM0PvWPC"
# For many `numpy` operations we can specify the `axis` over which to perform the operation:
# + colab={"base_uri": "https://localhost:8080/"} id="gXznlk6aiNP3" outputId="09591fed-8ec1-44b6-e98e-af7ca2e0824e"
# Concatenate matrices
print("\nConcatenate 2 arrays by the rows:")
print(np.concatenate((A, B), axis = 0)) # Concatenate rows - look at the shape
print("\nConcatenate 2 arrays by the columns:")
print(np.concatenate((A, B), axis = 1)) # Concatenate columns - look at the shape
# + [markdown] id="o4O_Kvftueqx"
# ## Basic Statistics
#
# You can easily calculate a lot of basic statistics from an array, such as the sum, mean, variance, maximum, argmax, etc. All of these can be retrieved either over the entire array or over rows/columns.
#
# For each of these functions, you can get the statistic for:
# - the whole array: `np.stat(arr)`
# - by row: `np.stat(arr, axis = 1)`
# - by column: `np.stat(arr, axis = 0)`
# + id="2JhB4W8F_gdp"
# Basic statistics of a matrix
A = np.arange(6).reshape([2, 3])
print("A")
print(A)
print("\n\nSum of the array")
print(np.sum(A)) # Sum all the matrix (return a scalar)
print("\nSum of the array by column")
print(np.sum(A, axis = 0)) # Sum by column (return a vector)
print("\nSum of the array by row")
print(np.sum(A, axis = 1)) # Sum by row (return a vector)
print("\nMax of each row")
print(np.max(A, axis = 1)) # Max by column (return a vector)
print("\nMax of each column")
print(np.max(A, axis = 0)) # Max by row (return a vector)
print("\nMax of entire array")
print(np.max(A))
print("\nAverage of all the array")
print(np.mean(A))
# + [markdown] id="qfF-hmlYw2fL"
# ## Sampling From Distributions
#
# `numpy` provides a broad set of distributions to sample from. We will cover this in more depth in lab 2.
# + [markdown] id="BOzh7VvbzFcJ"
# ## Linear Algebra
# One of the most important mathematical fields in machine learning is linear algebra. You can perform many of these operations using `numpy`. You can calculate the eigenvectors of a matrix, or its inverse, the rank of the matrix and so on.
# + colab={"base_uri": "https://localhost:8080/"} id="57F3_BlBR-R3" outputId="a5c64b69-da33-451b-f54d-940f46c40cc0"
A = np.array([[1., 2.], [3., 4.]])
print("A")
print(A)
print("\n\nInverse of A")
print(np.linalg.inv(A))
B = np.diag((1, 2, 3))
print("\n\nB")
print(B)
eigvalues, eigvectors = np.linalg.eig(B)
print("\neigenvalues, eigenvectors")
print(eigvalues, eigvectors)
print("\nRank of the matrix")
print(np.linalg.matrix_rank(B))
# + [markdown] id="7mLWs10U0dxW"
# ## Reshaping
#
# You can change the shape of your array, with transpose, flattening, reshape,
# adding a new axis (see `np.newaxis`)
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="qImJs4njAm43" outputId="3e70fefb-41fa-4c1c-d40e-1e1bb68e2590"
B = np.array([[1, 2, 3], [4, 5, 6]])
print("\n\nB")
print(B)
print("\nFlatten matrix")
print(np.ravel(B))
print("\nTranspose matrix")
print(B.transpose())
# + [markdown] id="OwjDVpmk1Obt"
# ## Sorting
#
# You can sort the matrix by row or by column
# + id="F7oe-5W4Vf2C"
B = np.array([[3, 6, 1, 4, 10], [5, 1, 8, 3, 65]])
print("B")
print(B)
print("\n\nSort by row")
print(np.sort(B, axis = 1))
# + [markdown] id="TCsGskFR14ff"
# ## Indexing By Condition
#
# + colab={"base_uri": "https://localhost:8080/"} id="YSIflm-x13uf" outputId="6790bea5-8076-416c-a327-68ec2ad1c882"
A = np.random.randint(1, 30, (5, 4))
print("A")
print(A)
print("\nGet the numbers that are greater than 5:")
print(A[A > 5])
print("\nGet the numbers that are divisible by 4:")
print(np.extract(np.mod(A, 4)==0, A))
# + [markdown] id="uZHrvwPc3xHl"
# ## Let's practice!
#
# To get you a bit more accustomed to `numpy` you are encouraged to solve the following challenges. If you choose to not solve the following challenges, be sure to understand the solutions. Do not use loops or list comprehensions.
#
# + [markdown] id="a7IjEOuKwxbp"
#
# Write a program to create a `7x10` matrix that has `0` and `1` staggered:
# ```
# # 0 1 0 1 0 1 0
# # 1 0 1 0 1 0 1
# # 0 1 0 1 0 1 0
# # 1 0 1 0 1 0 1
# ```
# Hint: use slice operations on different axes.
#
# + id="y_rAGqr2eknl"
staggered = np.zeros((7, 10))
staggered[::2, 1::2] = 1
staggered[1::2, ::2] = 1
m = 8
n = 11
tmp = np.arange(m*n).reshape((m, n))
staggered = np.ones((m, n))
staggered[(tmp%n)%2 != (tmp//n)%2] = 0
print(staggered)
# + [markdown] id="sQdwsu47y0_u"
# Calculate the volume of a cylinder with the following diameters and lengths:
#
# + id="bNPOP0jU2On0"
diameters = np.array([1, 3, 5, 2, 4])
lengths = np.array([10, 20, 3, 10, 5])
# + id="C23xh9uX6Sbj"
print(np.array(np.power(diameters/2, 2)*lengths*np.pi))
# + [markdown] id="o2RXomvKz1E2"
# Write a function that receives 2 vectors and returns their cartesian product:
# ```
# def create_cartesian_product(vec1, vec2):
# pass
# ```
#
#
# + id="Sqo0fuvPmlsM"
def cartesian_product(vec1, vec2):
# np.repeat([1, 2, 3], 4) -> [1, 1, 1, 1, 2, 2, 2, 2, 3, 3, 3, 3]
# np.tile([1, 2, 3], 4) -> [1, 2, 3, 1, 2, 3, 1, 2, 3, 1, 2, 3]
return np.transpose(np.array([np.repeat(vec1, len(vec2)), np.tile(vec2, len(vec1))]))
print(cartesian_product([1, 2, 3], [4, 5, 6, 7]))
# + [markdown] id="7fmVsBdq04UL"
# Given an array `a` and a number `n`, find the closest number to `n` in `a`:
# ```
# def find_closest(a, n):
# pass
# ```
#
#
# + id="8dipzX5I8AE-"
def find_closest(a, n):
a = np.array(a)
return a[np.argmin(np.abs(a - n))]
print(find_closest([1, 24, 12, 13, 14], 10))
# + [markdown] id="J_PKVyGO2dov"
# Check if the sudoku grid is valid:
# * Check that each row contains all the numbers from 1 to 9
# * Check that each column contains all the numbers from 1 to 9
# * Check that each of the 9 non-overlapping `3x3` blocks composing grid contain 1 to 9
#
# You can assume it contains only integers and that the shape of the array is `9x9`
# ```
# def check_sudoku(grid):
# pass
# ```
#
#
# + id="gZ5WPLiagvSI"
def is_1_to_9(array_):
# Return True if the array contains all the numbers from 1 to 9, False otherwise
return np.all(np.sort(array_, axis = None) == np.arange(1, 10))
def check_sudoku(grid):
def check_block(coords_block):
return is_1_to_9(grid[coords_block[0]*3 : coords_block[0]*3+3,
coords_block[1]*3 : coords_block[1]*3+3])
grid = np.array(grid)
# Check that the grid contains only 1 to 9
if (not is_1_to_9(np.unique(grid))):
return False
# Check that each line/column contains 1 to 9:
# Sort each column. We expect it to be
# [[1, 1, 1, 1, 1, 1, 1, 1, 1], [2, 2, 2, 2, 2, 2, 2, 2] ... [9, 9, 9, 9, 9, 9, 9, 9, 9]]
# Thus we expect the sums of the rows: 1*9, 2*9....
if np.any(np.sum(np.sort(grid, axis = 0), axis = 1) != np.arange(1, 10)*9):
return False
# Make the same for the columns
if np.any(np.sum(np.sort(grid.transpose(), axis = 0), axis = 1) != np.arange(1, 10)*9):
return False
# 0,0 0,0 0,0 0,1 0,1 0,1 0,2 0,2 0,2
# 0,0 0,0 0,0 0,1 0,1 0,1 0,2 0,2 0,2
# 0,0 0,0 0,0 0,1 0,1 0,1 0,2 0,2 0,2
#
#
# 1,0 1,0 1,0 1,1 1,1 1,1 1,2 1,2 1,2
# 1,0 1,0 1,0 1,1 1,1 1,1 1,2 1,2 1,2
# 1,0 1,0 1,0 1,1 1,1 1,1 1,2 1,2 1,2
#
#
# 2,0 2,0 2,0 2,1 2,1 2,1 2,2 2,2 2,2
# 2,0 2,0 2,0 2,1 2,1 2,1 2,2 2,2 2,2
# 2,0 2,0 2,0 2,1 2,1 2,1 2,2 2,2 2,2
# For each block of 9, check if it contains all the numbers 1 to 9
blocks_are_valid = np.apply_along_axis(check_block, 1, cartesian_product([0, 1, 2], [0, 1, 2]))
return np.all(blocks_are_valid)
# + [markdown] id="wmF3gn_pmP5X"
# Given a matrix, check if some row is a scalar multplication of another
# ```
# def check_dependencies(matrix_):
# pass
# ```
#
#
# + id="IOxhEqkmhsRS"
def check_dependencies(matrix_):
def rows_are_dependent(indices):
if indices[0] == indices[1]:
return False
return np.unique(matrix_[indices[0],] / matrix_[indices[1],]).shape[0] == 1
return np.any(np.apply_along_axis(rows_are_dependent, 1,
cartesian_product(np.arange(matrix_.shape[0]), np.arange(matrix_.shape[0]))))
# + [markdown] id="dwL7kvvTkkxb"
# Write a function that gets a 1D array and check if there is no local extrema point in addition to the global one.
# ```
# def have_an_extrema(array):
# pass
# ```
#
#
# + id="jg12aIuikhh3"
def have_a_maxima(array):
argmax_arr = np.argmax(array[1:-1]) + 1
before_max_neg = np.all(array[:argmax_arr - 1] - array[1:argmax_arr] < 0)
after_max_neg = np.all(array[argmax_arr:-1] - array[argmax_arr + 1:] > 0)
return before_max_neg and after_max_neg
def have_an_extrema(array):
# Check if it is a monotonic series
if np.unique(array[:-1] - array[1:]).shape[0] == 1: return True
# If there is a minimum, we can look for a maximum in the negated array
return have_a_maxima(array) or have_a_maxima(-array)
# + [markdown] id="Cf0Y2F8SlK6W"
# Note that instead of `array[:argmax_arr - 1] - array[1:argmax_arr]`, you could use the `np.diff` function
#
# + [markdown] id="m1S1Z7cE0E_l"
# # Pandas
# Until now, we have only looked at numerical data. But in real-world problems, we also have textual and categorical data. To manipulate this type of data, we will use the `pandas` library. One of the basic data structures of `pandas` is called a `DataFrame`. Generally, in a `DataFrame`, each row is a different sample and each column is a feature.
#
# For example, each row can represent a student, with columns of the ID, birthday, and gender of the student.
#
# `pandas` has a lot of possibilities of which we are going to introduce a very small subset.
# + [markdown] id="1JGYcSz82Viq"
# ## Array Creation
#
# In addition to creating an array from lists or randomly generated data frames, we are going to use an existing dataset of house prices (you will get back to this dataset in excercise 2).
# + id="8pjllLVn1qGW"
# Load https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data?select=train.csv
df = pd.read_csv('../datasets/house_train.csv', index_col=0)
df.head()
# + id="LZZwdRpn6FIf"
print("\nRows Names")
print(df.index)
print("\nColumns Names")
print(df.columns)
print("\nDf train shape")
print(df.shape)
# + [markdown] id="ECFka5mE49Qo"
# ## Indexing And Slicing
# Just like when using `numpy` you can select a subset of rows and columns. You can do it using indices or using names of the rows and columns. You can easily add a new column based on existing ones.
# + id="svkkCAff6sYZ"
print("\ndf[['GrLivArea', 'SalePrice', 'BedroomAbvGr']]")
print(df[['GrLivArea', 'SalePrice', 'BedroomAbvGr']]) # Select GrLivArea columns SalePrice BedroomAbvGr
print("\ndf.loc[3:10,['GrLivArea', 'SalePrice', 'BedroomAbvGr'] ]")
print(df.loc[3:10,['GrLivArea', 'SalePrice', 'BedroomAbvGr'] ])
print("\ndf.iloc[[3, 4, 5]]")
print(df.iloc[[3, 4, 5]])
print("\ndf.iloc[3:10,[6, 7, 8]]")
print(df.iloc[3:10,[6, 7, 8]])
# + id="qkqshRV-No7E"
individual_df = pd.DataFrame(np.array([np.random.randint(2000000, 3000000,50), np.random.uniform(1.50, 1.70, size = 50), np.random.uniform(45, 90, size = 50)]).transpose(), columns=['ID','Height','Weight'])
individual_df["BMI"] = individual_df["Weight"] / individual_df["Height"].pow(2)
print("\nIndividual DF")
print(individual_df.head())
# + [markdown] id="eyo4hxsp5TKm"
# # Basic Statistics
# `pandas` provides different statistical functions over `DataFrame`s.
# + id="Ho-eBIBt9FQI"
print("\nMedian of the SalePrice column")
print(df.SalePrice.median())
print("\nSelecting rows by condition")
median_price = df.SalePrice.median()
print(df[df.SalePrice > median_price].head())
# + [markdown] id="Q8lEIqi8CDNk"
# ## Group-by
#
# When working with data that contains also a categorical feature, we are often interested in performing some kind of calculation over all rows containing the same categorical value. For example, given a data frame of student grades for different courses, we can calculate the students' average grade.
# + id="tvip9jGT-Oh8"
students_df = pd.DataFrame(np.array([np.random.choice(["Zohar", "Shelly", "Omer", "Avi"],50), np.random.choice(["Linearit", "Intro", "Infi", "Probabilistic"], 50), np.random.randint(80, 101, 50)]).transpose(), columns=['Name','Course','Grade'])
students_df["Grade"] = students_df["Grade"].astype(int)
print("\n\nStudents df")
print(students_df.head())
print("\n\nCalculate average by student and by course")
print(students_df.groupby(['Name', 'Course']).mean().reset_index())
# + [markdown] id="mEFa78RCK1rq"
# ## Sorting
# As in `numpy` you can sort data frame based on a column
# + id="HjdZsJab-Z3J"
students_df.sort_values(by='Grade').head()
# + pycharm={"name": "#%%\n"}
df = pd.DataFrame(
{
"A": ["foo", "bar", "foo", "bar", "foo", "bar", "foo", "foo"],
"B": ["one", "one", "two", "three", "two", "two", "one", "three"],
"C": np.random.randn(8),
"D": np.random.randn(8),
}
)
def get_letter_type(letter):
if letter.lower() in 'aeiou':
return 'vowel'
else:
return 'consonant'
grouped = df.groupby(get_letter_type, axis=1)
# + [markdown] id="4CjuO2e5LdLD"
# ## Executing Functions By Columns
#
# In `pandas`, you can select columns and apply functions to them. You also can apply functions by elements.
# + [markdown] id="dvuu9aP6Ib-G"
# ## Merging Data Frames
#
# + id="51qFkcXH9c7c"
# Concatenate data frames
import pandas as pd
# Create 2 data frames
ids_df1 = pd.DataFrame({
'ID': ['336097897', '32109678', '25976389', '32438509', '36790307'],
'name': ['Amos', 'Eran', 'Sapir', 'Amichai', 'Hadar'],
'gender': ["M", "M", "F", "M", "F"]})
ids_df2 = pd.DataFrame({
'ID': ['21370565', '34256798', '3908412', '326780578'],
'name': ['Matan', 'Gabriel', 'Anael', 'Liora'],
'gender': ["M", "M", "F", "F"]})
print("\n\ndf1")
print(ids_df1)
print("\ndf2")
print(ids_df2)
print("\n\nJoin the two dataframes along rows:")
concatenate_data = pd.concat([ids_df1, ids_df2])
print(concatenate_data)
# +
birthdates_id = pd.DataFrame({
'ID': ['336097897', '32109678', '25976389', '32438509', '36790307', '21370565', '34256798', '3908412', '326780578'],
'birth_year': [1995, 1996, 1993, 1994, 1997, 1991, 1994, 1992, 1996]})
print("\nNow join the result_data and df_exam_data along ID:")
pd.merge(concatenate_data, birthdates_id, on='ID')
# + [markdown] id="4N-gAa7bcLp5"
# ## Let's Practice!
# + [markdown] id="HflHhEFaP5ya"
# Let us create a table of flight companies' flights. Each row will represent a single flight and will have 3 features: city of departure, city of destination and price.
#
# Implement a function `create_flight_df` that recieves a collection of cities and creates a dataset of randomly selected flights and a price in the range of 100-400.
#
# ```
# def create_flight_df(cities_poss, nrows = 100):
# pass
# ```
#
# The output data frame must not have more than a single record for any pair of cities. There are no flights from a city to itself.
# + id="KWY63-sATsr9"
import numpy as np
import pandas as pd
def create_flight_df(cities, nrows = 20):
df = pd.DataFrame([], columns=["Departure", "Destination", "Price"])
while df.shape[0] < nrows:
dep, dest = np.random.choice(cities, size=2, replace=False)
price = np.random.randint(100, 400)
if not ((df["Departure"] == dep) & (df["Destination"] == dest)).any():
df = df.append({"Departure": dep, "Destination": dest, "Price": price}, ignore_index=True)
return df
cities = ["Beijing", "Moscow", "New-York", "Tokyo", "Paris", "Cairo", "Santiago", "Lima", "Kinshasa", "Singapore",
"New-Delhi", "London", "Ankara", "Nairobi", "Ottawa", "Seoul", "Tehran", "Guatemala", "Caracas", "Vienna"]
flights = create_flight_df(cities)
flights.head()
# + [markdown] id="ItISmB4RYpcJ"
# As there are pairs of cities with no direct flight between them, let us find the pairs of cities that have a single connection flight between them and calculate the total price of the flgihts. To do so merge the two data frames. This operation is often referred to as "joining" with the options of inner, outer, left, right and cross joining. For more about merging `pandas` data frames read the [documentation](https://pandas.pydata.org/pandas-docs/stable/user_guide/merging.html#database-style-dataframe-or-named-series-joining-merging).
# + id="RvJjgHEkQq6g"
df = pd.merge(flights, flights, left_on=["Destination"], right_on=["Departure"], how="inner")
df = df[df.Departure_x != df.Destination_y]
df["Total_Price"] = df["Price_x"] + df["Price_y"]
df.head()
# -
# Create a data frame with all flights of no connection and single connection.
flights = flights.append(df[["Departure_x", "Destination_y", "Total_Price"]]\
.rename(columns={"Departure_x":"Departure", "Destination_y":"Destination", "Total_Price":"Price"}))
flights
# + [markdown] id="il0mLoEzhi6_"
# Since now we might have more than one way to flight between each pair of cities, let us find the cheapest flight option, with one connection, between two cities.
# + id="nYli93a6bMjB"
min_by_group = df.groupby(["Departure_x", "Destination_y"], as_index=False)["Total_Price"].min()
min_by_group
# + [markdown] id="TI2CrtB5irgn"
# And if we want to know on average what is the most expensive city to fly to then:
# + id="DUdokeBWiq1V"
mean_by_dest = min_by_group.groupby("Destination_y")["Total_Price"].mean()
expensive_city = mean_by_dest.idxmax()
print(expensive_city)
|
lab/Lab 00 - A - Data Analysis In Python - First Steps.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predict Function
#
# After performing a parameter update, which is done after some new measurement is collected, the next step is to incorporate motion into our Gaussian calculations. Recall that, as we estimate the location of a robot or self-driving car:
# * the measurement update *increases* our estimation certainty
# * the motion update/prediction *decreases* our certainty
#
# That is because every motion has some chance of under or overshooting its goal, and since motion is not exact, we end up losing some certainty about our exact location after each motion.
#
# Let's take the formulas from the example below and use them to write a program that takes in a mean and a motion and squared variances for both of those quantities, and returns a *new*, updated mean and variance for a new gaussian. This step is called the **motion update** or the predict step.
#
# <img src='images/motion_update.png' width="50%" height="50%">
#
# Below is our usual Gaussian equation and imports.
# +
# import math functions
from math import *
import matplotlib.pyplot as plt
import numpy as np
# gaussian function
def f(mu, sigma2, x):
''' f takes in a mean and squared variance, and an input x
and returns the gaussian value.'''
coefficient = 1.0 / sqrt(2.0 * pi *sigma2)
exponential = exp(-0.5 * (x-mu) ** 2 / sigma2)
return coefficient * exponential
# -
# For convenience, you've also been given the complete `update` code that performs a parameter update when an initial belief and new measurement information are merged.
# the update function
def update(mean1, var1, mean2, var2):
''' This function takes in two means and two squared variance terms,
and returns updated gaussian parameters.'''
# Calculate the new parameters
new_mean = (var2*mean1 + var1*mean2)/(var2+var1)
new_var = 1/(1/var2 + 1/var1)
return [new_mean, new_var]
# ### QUIZ: Write a `predict` function that returns new values for the mean and squared variance of a Gaussian after a motion.
#
# This function should take in parameters for an initial belief and motion and perform the measurement update as seen in the image at the top of this notebook.
# the motion update/predict function
def predict(mean1, var1, mean2, var2):
''' This function takes in two means and two squared variance terms,
and returns updated gaussian parameters, after motion.'''
## TODO: Calculate the new parameters
new_mean = mean1 + mean2
new_var = var1 + var2
return [new_mean, new_var]
# test your implementation
new_params = predict(10, 4, 12, 4)
print(new_params)
# ### Plot a Gaussian
#
# Plot a Gaussian by looping through a range of x values and creating a resulting list of Gaussian values, `g`, as shown below. You're encouraged to see what happens if you change the values of `mu` and `sigma2`.
# +
# display a gaussian over a range of x values
# define the parameters
mu = new_params[0]
sigma2 = new_params[1]
# define a range of x values
x_axis = np.arange(0, 40, 0.1)
# create a corresponding list of gaussian values
g = []
for x in x_axis:
g.append(f(mu, sigma2, x))
# plot the result
plt.plot(x_axis, g)
|
1. Predict Function, exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf_2.3
# language: python
# name: tf_2.3
# ---
# # Run colab 2021-01-07 ELECTRA(chinese-legal-electra-small-discriminator)
|
sanic_serving/electra_small/run_colab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tfs
# language: python
# name: tfs
# ---
# # Keras Sequential Model 2
# ### Building a sequential model
# #### Way 2: Using `add` method
# - We shall load our dataset as before.
# The tweak comes when creating the model
import tensorflow as tf
mnist = tf.keras.datasets.mnist
epochs = 20
batch_size = 32
mnist_data = mnist.load_data()
(x_train, y_train), (x_test, y_test) = mnist_data
x_train, x_test = tf.cast(x_train/255., dtype=tf.float32), tf.cast(x_test/255., dtype=tf.float32)
y_train, y_test = tf.cast(y_train, dtype=tf.int64), tf.cast(y_test, dtype=tf.int64)
#
# Build the model
#
model = tf.keras.models.Sequential()
# +
model.add(tf.keras.layers.Flatten())
model.add(tf.keras.layers.Dense(512, activation=tf.nn.relu))
model.add(tf.keras.layers.Dropout(0.2))
model.add(tf.keras.layers.Dense(10, activation='softmax'))
# -
model.compile(optimizer=tf.keras.optimizers.Adam(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy']
)
#
# Train: Fit inputs to the model outputs
#
model.fit(x_train, y_train, batch_size=batch_size, epochs=epochs)
# +
# An accuracy of 99.2 is fine, a little bad ass
# -
model.evaluate(x_test, y_test)
|
notebooks/1 intro_to_tensorflow/2.1b sequential_model_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="P9bJCDjdlgG6" colab_type="text"
# # **Tame your python**
#
# Let's see some classifiers in action
#
# `Leggo`
# + id="aQwc0re5mFld" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + id="Qy5eh1WsCDx6" colab_type="code" colab={}
def evaluate(y_test, y_pred):
from sklearn.metrics import accuracy_score
print("===== Accuracy Score =====")
print(accuracy_score(y_test, y_pred))
from sklearn.metrics import classification_report
print("===== Accuracy Score =====")
class_report = classification_report(y_test, y_pred)
print(class_report)
return
# + id="I-tpfuNjbj-q" colab_type="code" colab={}
# Visualising the results
def plot_model(classifier, X_set, y_set, y_test, y_pred, text):
from matplotlib.colors import ListedColormap
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('pink', 'cyan', 'lightgreen')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('red', 'blue', 'green'))(i), label = j)
plt.title(text)
plt.xlabel('X')
plt.ylabel('y')
plt.legend()
plt.show()
# + id="Rr8oUBtew6X7" colab_type="code" colab={}
def preprocess(X_train, X_test):
from sklearn.decomposition import PCA
pca = PCA(n_components = 2)
X_train = pca.fit_transform(X_train)
X_test = pca.transform(X_test)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
return X_train, X_test
# + [markdown] id="keG44qpdQiVA" colab_type="text"
# ## Get Wine Dataset
# + id="qyw8HvOuBEZm" colab_type="code" colab={}
from sklearn.datasets import load_wine
data = load_wine()
# + id="EukWofARwuL9" colab_type="code" outputId="245be705-aea3-4931-ac8c-e6c2259eaae9" colab={"base_uri": "https://localhost:8080/", "height": 35}
data.keys()
# + id="9BjoFGYJwyS0" colab_type="code" colab={}
X = data.data
y = data.target
# + id="SCvsH_ujALp7" colab_type="code" outputId="d57e345b-fe24-44d7-b8b6-9e5c08da0cdd" colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.model_selection import KFold
kf = KFold(n_splits=4)
kf.get_n_splits(X)
print(kf)
# + id="wKrVEKcmOH6I" colab_type="code" colab={}
def logistic_regression(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "Logistic Regression")
# + id="IYf3aBXKZOxj" colab_type="code" colab={}
def ridge_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.linear_model import RidgeClassifierCV
classifier = RidgeClassifierCV()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "RidgeClassifierCV")
# + id="j41ne3yuOLSm" colab_type="code" colab={}
def svm_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.svm import SVC
classifier = SVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SVC")
# + id="G-H_ZGr5e1wk" colab_type="code" colab={}
def mlp_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.neural_network import MLPClassifier
classifier = MLPClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "MLP")
# + id="BS6V1HSJad1f" colab_type="code" colab={}
def linearsvm_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.svm import LinearSVC
classifier = LinearSVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "LinearSVC")
# + id="Cif5chCUbF3h" colab_type="code" colab={}
def rf_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "RandomForestClassifier")
# + id="vzRuKzGracLw" colab_type="code" colab={}
def dt_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.tree import DecisionTreeClassifier
classifier = DecisionTreeClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "DecisionTreeClassifier")
# + id="om8U4fa_bJop" colab_type="code" colab={}
def gb_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.ensemble import GradientBoostingClassifier
classifier = GradientBoostingClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "GradientBoostingClassifier")
# + id="gU2yGvvqbNaH" colab_type="code" colab={}
def sgd_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.linear_model import SGDClassifier
classifier = SGDClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SGDClassifier")
# + id="2Nr9fmGGcxjx" colab_type="code" colab={}
def perceptron_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.linear_model import Perceptron
classifier = Perceptron()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "Perceptron")
# + id="bhfSkakWcx_g" colab_type="code" colab={}
def nb_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "GaussianNB")
# + id="JRMDGmgucyn4" colab_type="code" colab={}
def knn_classification(X_train, X_test, y_train, y_test):
X_train, X_test = preprocess(X_train, X_test)
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "KNeighborsClassifier")
# + id="Srk_QTAVAqox" colab_type="code" outputId="61720592-12ed-42bb-d47e-1bf50084c8fd" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
logistic_regression(X_train, X_test, y_train, y_test)
# + id="mmEHy7ZGkBRd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="533bb397-5ad1-464f-b756-d5fe217edf56"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
ridge_classification(X_train, X_test, y_train, y_test)
# + id="-BBrEGVNkEuU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="31296104-f41b-448f-ded1-0e97d733b676"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
svm_classification(X_train, X_test, y_train, y_test)
# + id="cAId0-_ykG4l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b8a6535f-07be-4e47-a5f8-1f022c36b36b"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
mlp_classification(X_train, X_test, y_train, y_test)
# + id="Jx_JL7lckJls" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="22425cda-79fd-452f-b1fd-4b7fdb0c8e70"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
linearsvm_classification(X_train, X_test, y_train, y_test)
# + id="SkuYAWKWkMbl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="79572370-1234-41be-ac70-60919eaec478"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
rf_classification(X_train, X_test, y_train, y_test)
# + id="uLHmoTs7kQRt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6762d4ae-7a48-44e3-8d08-6c10984843bf"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
dt_classification(X_train, X_test, y_train, y_test)
# + id="alyJ0Kd7kSU7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3b98c103-d9e5-4220-a733-a533cc11ac99"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
gb_classification(X_train, X_test, y_train, y_test)
# + id="uslqe_KJkUdU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c4a43285-2615-437f-abbf-5942e636125b"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
sgd_classification(X_train, X_test, y_train, y_test)
# + id="OhmnznFDkXD8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c39144e0-1f2a-476c-bbc3-e9cf73e2845c"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
perceptron_classification(X_train, X_test, y_train, y_test)
# + id="i_9YaylUkZEs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="98ebabba-0c23-412b-f259-8a692b2c31f1"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
knn_classification(X_train, X_test, y_train, y_test)
# + id="UXkrIl8Nka2E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e092afa5-6671-4d2d-ae27-7a27830f2e0b"
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
nb_classification(X_train, X_test, y_train, y_test)
|
MachineLearning_DataScience/Demo19_KfoldClassificationWine.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `dscal(N, DA, DX, INCX)`
#
# Scales a vector $\mathbf{x}$ by a constant $\alpha$.
#
# Operates on double-precision real valued arrays.
#
# Input scalar $\alpha$ is given by the double precision value `DA`.
# Input/output vector $\mathbf{x}$ is represented as a [strided array](../strided_arrays.ipynb) `DX`, spaced by `INCX`.
# Vector $\mathbf{x}$ is of size `N`.
# ### Example usage
# + jupyter={"source_hidden": true}
import os
import sys
sys.path.insert(0, os.path.abspath(os.path.join(os.path.abspath(''), "..", "..")))
# -
import numpy as np
from pyblas.level1 import dscal
x = np.array([1, 2, 3], dtype=np.double)
N = len(x)
alpha = 5
incx = 1
print("x before", x)
dscal(N, alpha, x, incx)
print("x after", x)
# ### Docstring
# + jupyter={"source_hidden": true}
help(dscal)
# -
# ### Source code
# + jupyter={"source_hidden": true}
# dscal??
|
docs/level1/dscal.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Write Output to CSV
# This submodule implements the writing of all input data (and eventually results, but not yet)
# to CSV files.
from __future__ import print_function
from salib import extend, import_notebooks
from Tables import Table
from Frame2D_Base import Frame2D
import Frame2D_Input
#test:
f = Frame2D('frame-1')
f.input_all()
# ## General Table Writing
# Define a general method to write all tables. By default, a table of a given name
# will be used from the '`self.rawdata`' object. If table '`xxx`' does not exist
# there, it will be created by calling method '`.list_xxx()`' to generate the data,
# and formed by using the list of columns in class attribute '`.COLUMNS_xxx`'.
#
# `.list_xxx()` must return a list of tuples or dictionaries, each of these being one
# row of the table. If a tuple, that data is obviously given in the same order as the columns.
# If a dictionary, the keys are the column names (see 'Member Loads', below).
@extend
class Frame2D:
def write_table(self,table_name,ds_name=None,prefix=None,record=True,precision=None,args=(),makedir=False):
t = getattr(self.rawdata,table_name,None)
if t is None:
methodname = 'list_'+table_name
method = getattr(self,methodname,None)
if method and callable(method):
data = method(*args)
t = Table(table_name,data=data,columns=getattr(self,'COLUMNS_'+table_name))
if t is None:
raise ValueError("Unable to find table '{}'".format(table_name))
t.write(ds_name=ds_name,prefix=prefix,precision=precision,makedir=makedir)
if record:
setattr(self.rawdata,table_name,t)
return t
##test:
try:
f.write_table('foobarzx3')
except Exception, e:
print('**** Error:',e)
# ### Nodes
@extend
class Frame2D:
def list_nodes(self):
return [(n.id,n.x,n.y) for n in self.nodes.values()]
##test:
t = f.write_table('nodes','test-1')
t.file_name
##test:
# %cat test-1.d/nodes.csv
# Delete the `nodes` table from `.rawdata` so that we can test that `.list_nodes()` is properly called
# to regenerated the table.
##test:
if hasattr(f.rawdata,'nodes'):
delattr(f.rawdata,'nodes')
# %rm test-1.d/nodes.csv
t = f.write_table('nodes','test-1')
t.file_name
##test:
# %cat test-1.d/nodes.csv
# ### Supports
@extend
class Frame2D:
def list_supports(self):
ans = []
for node in self.nodes.values():
if node.constraints:
cl = tuple(node.constraints)
if len(cl) < 3:
cl = cl + ('',)*(3-len(cl))
ans.append((node.id,)+cl)
return ans
##test:
f.list_supports()
##test:
t = f.write_table('supports','test-1')
t.file_name
##test:
# %cat test-1.d/supports.csv
##test:
x = 'supports'
if hasattr(f.rawdata,x):
delattr(f.rawdata,x)
# %rm test-1.d/supports.csv
t = f.write_table(x,'test-1')
t.file_name
##test:
# %cat test-1.d/supports.csv
# ### Members
@extend
class Frame2D:
def list_members(self):
return [(m.id,m.nodej.id,m.nodek.id) for m in self.members.values()]
##test:
f.list_members()
##test:
f.write_table('members','test-1')
# %cat test-1.d/members.csv
# ### Releases
@extend
class Frame2D:
def list_releases(self):
return [(m.id,)+tuple(m.releases) for m in self.members.values() if m.releases]
##test:
f.list_releases()
##test:
f.write_table('releases','test-1')
# %cat test-1.d/releases.csv
# ### Properties
@extend
class Frame2D:
def list_properties(self):
return [(m.id,m.size,m.Ix,m.A) for m in self.members.values()]
##test:
f.list_properties()
##test:
f.write_table('properties','test-1')
# %cat test-1.d/properties.csv
# ### Node Loads
@extend
class Frame2D:
def list_node_loads(self):
ans = []
dirns = ['FX','FY','FZ']
for loadid,node,nload in self.nodeloads:
for i in [0,1,2]:
if nload[i]:
ans.append((loadid,node.id,dirns[i],nload[i]))
return ans
##test:
f.list_node_loads()
##test:
f.write_table('node_loads','test-1')
# %cat test-1.d/node_loads.csv
# ### Support Displacements
@extend
class Frame2D:
def list_support_displacements(self):
ans = []
dirns = ['DX','DY','RZ']
for loadid,node,nload in self.nodedeltas:
for i in [0,1,2]:
if nload[i]:
ans.append((loadid,node.id,dirns[i],nload[i]))
return ans
##test:
f.list_support_displacements()
##test:
f.write_table('support_displacements','test-1')
# %cat test-1.d/support_displacements.csv
# ### Member Loads
# +
from MemberLoads import unmakeMemberLoad
@extend
class Frame2D:
def list_member_loads(self):
ans = []
for loadid,memb,mload in self.memberloads:
ml = unmakeMemberLoad(mload)
ml['MEMBERID'] = memb.id
ml['LOAD'] = loadid
ans.append(ml)
return ans
# -
##test:
f.list_member_loads()
##test:
f.write_table('member_loads','test-1')
# %cat test-1.d/member_loads.csv
##test:
if hasattr(f.rawdata,'member_loads'):
delattr(f.rawdata,'member_loads')
# %rm test-1.d/member_loads.csv
f.write_table('member_loads','test-1')
# %cat test-1.d/member_loads.csv
# ### Load Combinations
@extend
class Frame2D:
def list_load_combinations(self):
return [(case,load,factor) for case,load,factor in self.loadcombinations]
##test:
f.list_load_combinations()
##test:
f.write_table('load_combinations','test-1')
# %cat test-1.d/load_combinations.csv
# ### File Signatures
##test:
vars(f.rawdata).keys()
@extend
class Frame2D:
COLUMNS_signatures = ['NAME','PATH','SIGNATURE']
def list_signatures(self):
return [t.signature() for tn,t in vars(self.rawdata).items() if type(t) is Table]
##test:
f.list_signatures()
##test:
f.write_table('signatures','test-1',record=False)
# %cat test-1.d/signatures.csv
# ## Write everything
# +
import os, os.path
@extend
class Frame2D:
def write_all(self,ds_name,mkdir=False):
if mkdir:
dname = ds_name + '.d'
if not os.path.exists(dname):
os.mkdir(dname)
self.write_table('nodes',ds_name)
self.write_table('supports',ds_name)
self.write_table('members',ds_name)
self.write_table('releases',ds_name)
self.write_table('properties',ds_name)
self.write_table('node_loads',ds_name)
self.write_table('support_displacements',ds_name)
self.write_table('member_loads',ds_name)
self.write_table('load_combinations',ds_name)
self.write_table('signatures',ds_name,record=False)
# -
##test:
# %rm test-1.d/*
# %ls -l test-1.d
f.reset()
f.input_all()
f.write_all('test-1')
# %ls -l test-1.d
##test:
# %cat test-1.d/signatures.csv
# ## Results Output
##test:
import Frame2D_SolveFirstOrder
f.reset()
f.input_all()
rs = f.solve('all')
# ### Node Displacements
@extend
class Frame2D:
COLUMNS_node_displacements = ['NODEID','DX','DY','RZ']
def list_node_displacements(self,rs):
if not hasattr(rs,'node_displacements'):
return []
ans = []
D = rs.node_displacements
for node in self.nodes.values():
d = D[node.dofnums]
ans.append((node.id,d[0,0],d[1,0],d[2,0]))
return ans
##test:
f.list_node_displacements(rs)
##test:
f.write_table('node_displacements',ds_name='test-1',prefix=rs.loadcase,record=False,precision=15,args=(rs,))
# %cat test-1.d/all/node_displacements.csv
# ### Reaction Forces
@extend
class Frame2D:
COLUMNS_reaction_forces = ['NODEID','FX','FY','MZ']
def list_reaction_forces(self,rs):
if not hasattr(rs,'reaction_forces'):
return []
R = rs.reaction_forces
ans = []
for node in self.nodes.values():
if node.constraints:
l = [node.id,None,None,None]
for dirn in node.constraints:
i = node.DIRECTIONS[dirn]
j = node.dofnums[i]
val = R[j-self.nfree,0]
l[i+1] = val
ans.append(l)
return ans
##test:
f.list_reaction_forces(rs)
##test:
f.write_table('reaction_forces',ds_name='test-1',prefix=rs.loadcase,record=False,precision=15,args=(rs,))
# %cat test-1.d/all/reaction_forces.csv
# ### Member End Forces
@extend
class Frame2D:
COLUMNS_member_end_forces = 'MEMBERID,FXJ,FYJ,MZJ,FXK,FYK,MZK'.split(',')
def list_member_end_forces(self,rs):
if not hasattr(rs,'member_efs'):
return []
mefs = rs.member_efs
ans = []
for memb in self.members.values():
efs = mefs[memb].fefs
ans.append((memb.id,efs[0,0],efs[1,0],efs[2,0],efs[3,0],efs[4,0],efs[5,0]))
return ans
##test:
f.list_member_end_forces(rs)
##test:
f.write_table('member_end_forces',ds_name='test-1',prefix=rs.loadcase,record=False,precision=15,args=(rs,))
# %cat test-1.d/all/member_end_forces.csv
# ### $P-\Delta$ Forces
# See module ```..._Display```
#
# ### All Results
@extend
class Frame2D:
def write_results(self,ds_name,rs):
self.write_table('node_displacements',ds_name=ds_name,prefix=rs.loadcase,record=False,
precision=15,args=(rs,),makedir=True)
self.write_table('reaction_forces',ds_name=ds_name,prefix=rs.loadcase,record=False,
precision=15,args=(rs,))
self.write_table('member_end_forces',ds_name=ds_name,prefix=rs.loadcase,record=False,
precision=15,args=(rs,))
if rs.pdelta:
self.write_table('pdelta_forces',ds_name=ds_name,prefix=rs.loadcase,record=False,
precision=15,args=(rs,))
# +
##test:
f.reset()
f.input_all()
rs = f.solve('One')
f.write_results('test-1',rs)
# %ls -l test-1.d/One
# %cat test-1.d/One/node_displacements.csv
# %cat test-1.d/One/reaction_forces.csv
# %cat test-1.d/One/member_end_forces.csv
# -
|
Devel/V05/Frame2D_Output.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Y4kmqOoK2S-I" outputId="8e0a0ca2-64e0-4a6d-bded-ef7bef9d0cbf"
from google.colab import drive
drive.mount('/content/drive')
# + id="2T_jcsvb6LIL"
import pandas as pd
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, confusion_matrix,accuracy_score
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import DecisionTreeRegressor
from sklearn.cluster import KMeans
from numpy import unique
from numpy import where
from matplotlib import pyplot
# + id="oqBrTcUN6MQi" colab={"base_uri": "https://localhost:8080/"} outputId="beb4333a-e262-4d18-e399-2c6f1d10baf7"
data = pd.read_csv('/content/drive/MyDrive/projeler/Pokemon.csv')
data.median()
# + colab={"base_uri": "https://localhost:8080/"} id="NV60AqOr5G8i" outputId="21a6a9f0-c8b4-464c-f13e-a319a5f88f54"
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
labels = encoder.fit_transform(data.iloc[:,12])
labels.shape
# + id="Q9lotgkm5etQ"
data = data.drop('#', axis=1)
data = data.drop('Name', axis=1)
data = data.drop('Type 1', axis=1)
data = data.drop('Type 2', axis=1)
data = data.drop('Legendary', axis=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 406} id="eHSIujPSjr55" outputId="f3c72544-a290-4f44-f809-7d4d81e0b999"
data
# + colab={"base_uri": "https://localhost:8080/"} id="x5pjiMQvkjBJ" outputId="cfcc3556-99d4-4950-d01d-f5d2f41e38d9"
X = data.iloc[:,:]
y = labels
X.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 610} id="EOytYdQspLRA" outputId="fa601a2b-0c23-403b-bf8a-a2c059ac90a3"
from sklearn.datasets import load_digits
from sklearn.decomposition import FactorAnalysis
X, _ = load_digits(return_X_y=True)
transformer = FactorAnalysis(n_components=2, random_state=0)
X_transformed1 = transformer.fit_transform(X)
fig=plt.figure(figsize=(10, 10))
plt.scatter(X_transformed1[:,0], X_transformed1[:,1])
plt.show()
X_transformed1.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 610} id="rcyMsg8h9xY5" outputId="b8ca0b1d-fb0e-4452-eacd-ae0a6609bc35"
from sklearn.datasets import load_digits
from sklearn.manifold import MDS
X, _ = load_digits(return_X_y=True)
X.shape
embedding = MDS(n_components=2)
X_transformed2 = embedding.fit_transform(X)
fig=plt.figure(figsize=(10, 10))
plt.scatter(X_transformed2[:,0], X_transformed2[:,1])
plt.show()
X_transformed2.shape
# + id="FmIjDCodckKg"
|
10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from pathlib import Path
import gzip
import json
import itertools
import io
import pickle
from numbers import Number
log_dir = Path('/Users/mairas/BTSync/Shared/hurma_data/sk-data-log')
log_filenames = log_dir.glob('*.log*')
def smart_open(fn):
opener = gzip.open if fn.suffix=='.gz' else open
return opener(fn)
def line_reader(f):
for line in f:
yield line if type(line)==str else line.decode('utf-8')
all_lines = itertools.chain.from_iterable(
(line_reader(smart_open(lfn)) for lfn in log_filenames))
# %%time
all_lines = list(all_lines)
len(all_lines)
l0 = all_lines[0]
print(l0)
def load_all_lines(all_lines):
for line in all_lines:
try:
json_object = json.loads(line)
except Exception as err:
continue
yield json_object
all_json = load_all_lines(all_lines)
# + active=""
# all_paths = [
# 'design.aisShipType',
# 'design.beam',
# 'design.draft',
# 'design.length',
# 'environment.depth.belowSurface',
# 'environment.depth.belowTransducer',
# 'environment.depth.surfaceToTransducer',
# 'environment.water.temperature',
# 'environment.wind.angleApparent',
# 'environment.wind.speedApparent',
# 'navigation.courseGreatCircle.bearingTrackTrue',
# 'navigation.courseGreatCircle.crossTrackError',
# 'navigation.courseGreatCircle.nextPoint.bearingTrue',
# 'navigation.courseGreatCircle.nextPoint.distance',
# 'navigation.courseGreatCircle.nextPoint.position',
# 'navigation.courseGreatCircle.nextPoint.timeToGo',
# 'navigation.courseGreatCircle.nextPoint.velocityMadeGood',
# 'navigation.courseOverGroundTrue',
# 'navigation.courseRhumbline.crossTrackError',
# 'navigation.courseRhumbline.nextPoint.position',
# 'navigation.currentRoute.name',
# 'navigation.currentRoute.waypoints',
# 'navigation.datetime',
# 'navigation.destination.commonName',
# 'navigation.gnss.antennaAltitude',
# 'navigation.gnss.geoidalSeparation',
# 'navigation.gnss.horizontalDilution',
# 'navigation.gnss.integrity',
# 'navigation.gnss.methodQuality',
# 'navigation.gnss.positionDilution',
# 'navigation.gnss.satellites',
# 'navigation.gnss.type',
# 'navigation.headingMagnetic',
# 'navigation.headingTrue',
# 'navigation.log',
# 'navigation.magneticVariation',
# 'navigation.position',
# 'navigation.rateOfTurn',
# 'navigation.speedOverGround',
# 'navigation.speedThroughWater',
# 'navigation.speedThroughWaterReferenceType',
# 'navigation.state',
# 'navigation.trip.log',
# 'notifications.instrument.PilotOffCourse',
# 'propulsion.port.coolantTemperature',
# 'propulsion.port.exhaustTemperature',
# 'propulsion.port.temperature',
# 'sensors.ais.fromBow',
# 'sensors.ais.fromCenter',
# 'sensors.engine-temp.freeMem',
# 'steering.autopilot.target.headingMagnetic',
# 'steering.rudderAngle']
# -
enabled_paths = {
'environment.depth.belowSurface',
'environment.water.temperature',
'environment.wind.angleApparent',
'environment.wind.speedApparent',
'navigation.courseOverGroundTrue',
'navigation.headingMagnetic',
'navigation.headingTrue',
'navigation.magneticVariation',
'navigation.position',
'navigation.rateOfTurn',
'navigation.speedOverGround',
'navigation.speedThroughWater',
'propulsion.port.coolantTemperature',
'propulsion.port.exhaustTemperature',
'propulsion.port.temperature',
}
# + active=""
# %%time
# all_json = list(all_json)
# -
def collect_data(all_json):
data = {}
for j in all_json:
for upd in j['updates']:
dt = np.datetime64(upd['timestamp'])
for val in upd['values']:
if val['path'] in enabled_paths:
if isinstance(val['value'], dict):
for val_k, val_v in val['value'].items():
if not isinstance(val_v, Number):
continue
k = val['path'] + '.' + val_k
data.setdefault(k, []).append((dt, val_v))
else:
if not isinstance(val['value'], Number):
continue
data.setdefault(val['path'], []).append((dt, val['value']))
return data
# %%time
data = collect_data(all_json)
data.keys()
def convert_to_pandas(data):
pddata = {}
for k, v in data.items():
t = np.array([c[0] for c in v])
d = np.array([c[1] for c in v])
ser = pd.Series(d, index=t, name=k)
pddata[k] = ser.loc[~ser.index.duplicated(keep='first')]
return pddata
pddata = convert_to_pandas(data)
pddata['environment.wind.angleApparent']
# + active=""
# del data
# -
with (log_dir / "pddata.pickle").open('wb') as f:
pickle.dump(pddata, f)
|
examples/log_parser.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import json
import csv
import requests
import time
import os
import shutil
import matplotlib.pyplot as plt
from markowitz import *
pd.set_option('display.max_rows',10)
# - Primeiro coletamos os dados do mercado em um dict:
market = coimetrics_get_whole_market()
# - depois montamos um dataframe contendo apenas moedas existentes desde a data estabelecida
mkt = merge_market_data(market, pd.Timestamp(2018,1,1))
# - Em seguida definimos nosso portfolio com as moedas em que estamos interessados
l = ['btc','ltc','eth','ada']
#todas as opções de moedas podem ser obtidas em get_assets_from_coimetrics()
portfolio = get_portfolio_data_from_market_dataframe(mkt, l)
# - Em seguida simulamos alguns portfolios, e definimos o com melhor sharpe(em vermelho) e o mais seguro(em preto)
a = markovitz_monte_carlo(portfolio)
print('sharpe',a[3])
|
Notebooks/cryptodata/markowitz_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Sparameter and Power Data
# This notebook demonstrates how to open many data sources, arrange the data and combine it into a single csv file. Note, because of the size of the data sources involved they are not included. For more information please contact [<NAME>](mailto:<EMAIL>)
#
# ## Import and transformation of data files to a database
# ### There are several different sources of structured data each one with formatting differences
# ### Check Standard data sources
# 1. One Port Raw files that have been converted using Ron Ginley's BDAT -> Ascii converter
# 2. One Port files already stored in a legacy SAS database exported into csv by Jolene Spett
# 3. Two Port Raw files that have been converted using Ron Ginley's BDAT -> Ascii converter
# 4. Two Port files already stored in a legacy SAS database exported into csv by Jolene Spett
# 5. Two Port Non-Reciprocal files that have been converted using Ron Ginley's BDAT -> Ascii converter
# 6. Power Raw files files that have been converted using Ron Ginley's BDAT -> Ascii converter
# 7. Power files already stored in a legacy SAS database exported into csv by Jolene Spett
#
# ### DUT data sources, already analyzed using various versions of the Calrep HP Basic program
# 1. One Port .asc files stored in ascii.dut folder
# 2. Two Port .asc files stored in ascii.dut folder
# 3. Power .asc files with 4 error columns per s-parameter and power value stored in ascii.dut folder
# 4. Power .asc files with 3 error columns per power value stored in ascii.dut
#
# ### Conversion of these files requires: opening, parsing, and standardization of the data
# import of needed libraries
import os
import re
import datetime
import pandas
from types import *
# import of pyMez to change to import pyMez when __init__.py is changed
from pyMez.Code.DataHandlers.NISTModels import *
from pyMez.Code.Utils.Names import *
import numpy as np
import matplotlib.pyplot as plt
# Location of the various data sources
#input data sources
CONVERTED_RAW_FILE_DIRECTORY=r'C:\Share\Ck_Std_raw_ascii'
SAS_ONE_PORT=os.path.join(TESTS_DIRECTORY,'onechks.csv')
SAS_TWO_PORT=os.path.join(TESTS_DIRECTORY,'twochks.csv')
SAS_POWER=os.path.join(TESTS_DIRECTORY,'powchks.csv')
DUT_TOP_DIRECTORY=r'C:\Share\ascii.dut'
# output data
ONE_PORT_CHKSTD_CSV=r"C:\Share\Converted_Check_Standard\One_Port_Check_Standard.csv"
TWO_PORT_CHKSTD_CSV=r"C:\Share\Converted_Check_Standard\Two_Port_Check_Standard.csv"
TWO_PORT_NR_CHKSTD_CSV=r"C:\Share\Converted_Check_Standard\Two_Port_NR_Check_Standard.csv"
POWER_CHKSTD_CSV=r"C:\Share\Converted_Check_Standard\Power_Check_Standard.csv"
COMBINED_ONE_PORT_CHKSTD_CSV=r"C:\Share\Converted_Check_Standard\Combined_One_Port_Check_Standard.csv"
COMBINED_TWO_PORT_CHKSTD_CSV=r"C:\Share\Converted_Check_Standard\Combined_Two_Port_Check_Standard.csv"
COMBINED_POWER_CHKSTD_CSV=r"C:\Share\Converted_Check_Standard\Combined_Power_Check_Standard.csv"
ONE_PORT_CALREP_CSV=r"C:\Share\Converted_DUT\One_Port_DUT.csv"
TWO_PORT_CALREP_CSV=r"C:\Share\Converted_DUT\Two_Port_DUT.csv"
POWER_3TERM_CALREP_CSV=r"C:\Share\Converted_DUT\Power_3Term_DUT.csv"
POWER_4TERM_CALREP_CSV=r"C:\Share\Converted_DUT\Power_4Term_DUT.csv"
SQL_DATABASE=r"C:\Share\Sql_DUT_Checkstandard\sqlite_sparameter_power.db"
# ### Creating import lists by type for the converted raw data sources
# 1. We read all the files in the top folder
# 2. The 5th line in the header determines the data type (python is zero indexed so it is element 4)
# 3. We create 4 lists of all files matching the various types (One-port, Two-Port, Two-PortNR, Power)
# +
# We first get all files in the desired directory
file_names=os.listdir(CONVERTED_RAW_FILE_DIRECTORY)
# The loop runs quicker if we create lists and then add to them
# We create lists of the full path name for each of the data types
raw_files=[]
one_port_raw_files=[]
two_port_raw_files=[]
two_port_NR_raw_files=[]
power_raw_files=[]
# We iterate through the fiel names using the 5 th line to sort into our types
for index,file_name in enumerate(file_names[:]):
in_file=open(os.path.join(CONVERTED_RAW_FILE_DIRECTORY,file_name),'r')
lines=[]
for line in in_file:
lines.append(line)
in_file.close()
#print index,file_name
if re.search('1-port',lines[4],re.IGNORECASE):
one_port_raw_files.append(os.path.join(CONVERTED_RAW_FILE_DIRECTORY,file_name))
elif re.search('2-port',lines[4],re.IGNORECASE) and not re.search('2-portNR',lines[4],re.IGNORECASE):
two_port_raw_files.append(os.path.join(CONVERTED_RAW_FILE_DIRECTORY,file_name))
elif re.search('2-portNR',lines[4],re.IGNORECASE):
two_port_NR_raw_files.append(os.path.join(CONVERTED_RAW_FILE_DIRECTORY,file_name))
elif re.search('Thermistor|Dry Cal',lines[4],re.IGNORECASE):
power_raw_files.append(os.path.join(CONVERTED_RAW_FILE_DIRECTORY,file_name))
# This loop takes about 10 seconds
# -
# Now we can check if the loop worked properly
print("There are %s total files"%len(file_names))
print("There are %s one port raw files"%len(one_port_raw_files))
print("There are %s two port raw files"%len(two_port_raw_files))
print("There are %s two port NR raw files"%len(two_port_NR_raw_files))
print("There are %s power raw files"%len(power_raw_files))
total_binned_files=(len(one_port_raw_files)+len(two_port_raw_files)+len(two_port_NR_raw_files)+len(power_raw_files))
if len(file_names)==total_binned_files:
print("All Files Have Been Acounted For")
else:
print("{0} out of {1} files were binned".format(total_binned_files,len(file_names)) )
# Now each data source has to be parsed and converted to a common form
# One large issue is checking for data overlap in the data sources that can be solved by analysing the timestamp
# Also after trying several ways of doing the conversion, the best seems to be create a small csv and then add on
def build_csv_from_raw_script(input_file_names_list,output_file_name,model_name):
"""Build csv from raw script takes a list of file names conforming to model and builds a single csv.
It is intentioned to accept raw files from the sparameter power project that have been converted from bdat
using Ron Ginely's convertor (modified calrep program). The output is a single csv file with metadata added
as extra columns (ie a denormalized table)"""
try:
# our current definition of metadata keys for all of the raw models
metadata_keys=["System_Id","System_Letter","Connector_Type_Calibration","Connector_Type_Measurement",
"Measurement_Type","Measurement_Date","Measurement_Time","Program_Used","Program_Revision","Operator",
"Calibration_Name","Calibration_Date","Port_Used","Number_Connects","Number_Repeats","Nbs",
"Number_Frequencies","Start_Frequency",
"Device_Description","Device_Id"]
# import the first file
model=globals()[model_name]
initial_file=model(input_file_names_list[0])
# Add the metadata columns and replace any commas with -
for column_name in metadata_keys:
initial_file.add_column(column_name=column_name,column_type='str',
column_data=[initial_file.metadata[column_name].replace(',','-')
for row in initial_file.data])
# We also add a column at the end that is Measurement_Timestamp, that is
# Measurement_Time+Measurement_Date in isoformat
timestamp=initial_file.metadata["Measurement_Date"]+" "+initial_file.metadata["Measurement_Time"]
datetime_timestamp=datetime.datetime.strptime(timestamp,'%d %b %Y %H:%M:%S')
measurement_timestamp=datetime_timestamp.isoformat(' ')
initial_file.add_column(column_name="Measurement_Timestamp",column_type='str',
column_data=[measurement_timestamp
for row in initial_file.data])
# now we save the intial file with its column names but not its header
initial_file.header=None
initial_file.save(output_file_name)
# Now we re-open this file in the append mode and read-in each new file and append it. This seems to work
# for very large data sets, where as keeping a single object in memory fails
out_file=open(output_file_name,'a')
# now we do the same thing over and over and add it to the out file
for file_name in input_file_names_list[1:]:
model=globals()[model_name]
parsed_file=model(file_name)
for column_name in metadata_keys:
parsed_file.add_column(column_name=column_name,column_type='str',
column_data=[parsed_file.metadata[column_name].replace(',','-')
for row in parsed_file.data])
timestamp=parsed_file.metadata["Measurement_Date"]+" "+parsed_file.metadata["Measurement_Time"]
datetime_timestamp=datetime.datetime.strptime(timestamp,'%d %b %Y %H:%M:%S')
measurement_timestamp=datetime_timestamp.isoformat(' ')
parsed_file.add_column(column_name="Measurement_Timestamp",column_type='str',
column_data=[measurement_timestamp
for row in parsed_file.data])
# add an endline before appending
out_file.write('\n')
# now we only want the data string
data=parsed_file.get_data_string()
out_file.write(data)
# close the file after loop
out_file.close()
# Catch any errors
except:
raise
# Now we can try it for a subset of one ports
build_csv_from_raw_script(one_port_raw_files[:10],ONE_PORT_CHKSTD_CSV,"OnePortRawModel")
# we re-import the csv using pandas to make sure it worked
one_port_raw_data_frame=pandas.read_csv(ONE_PORT_CHKSTD_CSV)
one_port_raw_data_frame[:5]
# %timeit build_csv_from_raw_script(one_port_raw_files[:10],ONE_PORT_CHKSTD_CSV,"OnePortRawModel")
# This loop takes ~ 16.3 ms to run for a file so 16.3ms * num_files
16.3*10**-3*len(one_port_raw_files)
# Let's do all of the files and time it, this loop will be the worst case senario
import_list=one_port_raw_files[:]
start_time=datetime.datetime.now()
build_csv_from_raw_script(import_list,ONE_PORT_CHKSTD_CSV,"OnePortRawModel")
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were converted to a single csv in {1} seconds".format(len(import_list),diff.total_seconds()))
# %matplotlib notebook
# Now let's check the integrity of the data by re-importing, selecting and plotting some of it
one_port_raw_data_frame=pandas.read_csv(ONE_PORT_CHKSTD_CSV)
test_subset=one_port_raw_data_frame[one_port_raw_data_frame["Device_Id"]==80]
if COMBINE_S11_S22:
test_subset.plot(x="Frequency",y="mag")
else:
test_subset.plot(x="Frequency",y="magS11")
plt.show()
# +
# now 2 port
import_list=two_port_raw_files[:]
start_time=datetime.datetime.now()
build_csv_from_raw_script(import_list,TWO_PORT_CHKSTD_CSV,"TwoPortRawModel")
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were converted to a single csv in {1} seconds".format(len(import_list),diff.total_seconds()))
# +
# now 2 port NR
import_list=two_port_NR_raw_files[:]
start_time=datetime.datetime.now()
build_csv_from_raw_script(import_list,TWO_PORT_NR_CHKSTD_CSV,"TwoPortNRRawModel")
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were converted to a single csv in {1} seconds".format(len(import_list),diff.total_seconds()))
# +
# now power
import_list=power_raw_files[:]
start_time=datetime.datetime.now()
build_csv_from_raw_script(import_list,POWER_CHKSTD_CSV,"PowerRawModel")
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were converted to a single csv in {1} seconds".format(len(import_list),diff.total_seconds()))
# -
# ### Now that the conversion of the raw data files is finished, we need to import data from the legacy SAS database, check to see if it is in the csv file, and if it is not add it. It should be noted that the data in the SAS database has a different number of columns, so that some of them need to be translated. In addition, there is no SAS equivelent to two port NR.
# ### Adition of data process
# 1. import tables of new raw and SAS types
# 2. rename any columns that are equivelent
# 3. create any columns that are converted forms (dates)
# 4. delete any extra columns
# 5. add empty columns for undefined values
# 6. exclude any that appear in new raw data set
# 7. export new joined file
#
# step 1: import data sets
raw_one_port=pandas.read_csv(ONE_PORT_CHKSTD_CSV)
sas_one_port=pandas.read_csv(SAS_ONE_PORT)
# step 2: rename any columns that are the same with different names
same={"spid":"System_Id","SP":"Port_Used","ctype":"Connector_Type_Measurement","checkid":"Device_Id",
"MGAMA":"magS11","PGAMA":"argS11","CON":"Connect","FREQ":"Frequency"}
if COMBINE_S11_S22:
same["MGAMA"]="mag"
same["PGAMA"]="arg"
new=sas_one_port.rename(columns=same)
# step 3: create derived columns
def date_conversion(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.isoformat(" ")
def to_measurement_date(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%d %b %y")
def to_measurement_time(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%H:%M:%S")
def to_calibration_date(date_sas_format):
if type(date_sas_format) is StringType:
datetime_format=datetime.datetime.strptime(str(date_sas_format),'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%d %b %y")
else:
return date_sas_format
new["Measurement_Timestamp"]=new["MEASDATE"].map(date_conversion)
new["Measurement_Date"]=new["MEASDATE"].map(to_measurement_date)
new["Measurement_Time"]=new["MEASDATE"].map(to_measurement_time)
new["Calibration_Date"]=new["CALDATE"].map(to_calibration_date)
if COMBINE_S11_S22:
pass
else:
new["magS22"]=0.0
new["argS22"]=0.0
new["Measurement_Type"]='1-port'
# step 4: delete any extra columns
del new["CALDATE"]
del new["MEASDATE"]
del new["CAL"]
# check our progress
new[:5]
# step 5: add empty columns
empty_columns=[ u'Direction', u'System_Letter',
u'Connector_Type_Calibration', u'Program_Used', u'Program_Revision', u'Operator', u'Calibration_Name',
u'Number_Connects',
u'Number_Repeats', u'Nbs', u'Number_Frequencies', u'Start_Frequency',
u'Device_Description']
for empty_column in empty_columns:
new[empty_column]=None
# Now check that the column names are the same and order them
raw_columns=raw_one_port.columns
print raw_columns
new=new[raw_columns]
new_columns=new.columns
print new_columns
raw_columns==new_columns
# step 6: exclude any files that already exist
unique_timestamps=raw_one_port["Measurement_Timestamp"].unique()
new=new[-new["Measurement_Timestamp"].isin(unique_timestamps)]
# step7: add the files and save as csv, note at this point we can write to a db also
combined=pandas.concat([raw_one_port,new])
# combined["mag"]=combined["magS11"]+combined["magS22"]
# combined["arg"]=combined["argS11"]+combined["argS22"]
# del combined["magS11"]
# del combined["magS22"]
# del combined["argS11"]
# del combined["argS22"]
# column_order=[u'Frequency', u'Direction', u'Connect', u'mag', u'arg', u'System_Id', u'System_Letter',
# u'Connector_Type_Calibration', u'Connector_Type_Measurement',
# u'Measurement_Type', u'Measurement_Date', u'Measurement_Time',
# u'Program_Used', u'Program_Revision', u'Operator', u'Calibration_Name',
# u'Calibration_Date', u'Port_Used', u'Number_Connects',
# u'Number_Repeats', u'Nbs', u'Number_Frequencies', u'Start_Frequency',
# u'Device_Description', u'Device_Id', u'Measurement_Timestamp']
combined.to_csv(COMBINED_ONE_PORT_CHKSTD_CSV,index=False)
# Finally we check that the files are all what we expect
number_measurements_raw=len(raw_one_port["Measurement_Timestamp"].unique())
number_measurements_sas=len(sas_one_port["MEASDATE"].unique())
number_new=len(new["Measurement_Timestamp"].unique())
number_combined=len(combined["Measurement_Timestamp"].unique())
print("There were {0} measurements in the raw one port files".format(number_measurements_raw))
print("There were {0} measurements in the sas one port files".format(number_measurements_sas))
print("{0} measurements did not overlap".format(number_new))
print("This resulted in {0} unique measurements".format(number_combined))
print("The statement that the number of raw + non-overlapping measurements is equal to the number of"
"combined measurements is {0}, resulting in {1} rows of"
"data".format(number_new+number_measurements_raw==number_combined,len(combined)))
# show a detailed row count, showing how many values are empty
combined.count()
# Finaly check the data by importing it
start_time=datetime.datetime.now()
combined_csv=pandas.read_csv(COMBINED_ONE_PORT_CHKSTD_CSV)
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were imported as a single csv in {1} seconds".format(len(combined_csv),diff.total_seconds()))
combined_csv.count()
number_standards=len(combined_csv["Device_Id"].unique())
print("The number of 1-port check standards is {0}".format(number_standards))
# ### Repeat for 2-ports
# todo: make this a stand alone script
# step 1: import data sets
raw_two_port=pandas.read_csv(TWO_PORT_CHKSTD_CSV)
sas_two_port=pandas.read_csv(SAS_TWO_PORT)
# step 2: rename any columns that are the same with different names
same={"spid":"System_Id","SP":"Port_Used","ctype":"Connector_Type_Measurement","checkid":"Device_Id",
"MS11":"magS11","PS11":"argS11","PS12":"argS21","MS22":"magS22","PS22":"argS22",
"CON":"Connect","FREQ":"Frequency"}
new=sas_two_port.rename(columns=same)
# step 3: create derived columns
def date_conversion(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.isoformat(" ")
def to_measurement_date(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%d %b %y")
def to_measurement_time(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%H:%M:%S")
def to_calibration_date(date_sas_format):
if type(date_sas_format) is StringType:
datetime_format=datetime.datetime.strptime(str(date_sas_format),'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%d %b %y")
else:
return date_sas_format
def to_linear(s12_sas_format):
return 10.**(-1.*s12_sas_format/20.)
new["Measurement_Timestamp"]=new["MEASDATE"].map(date_conversion)
new["Measurement_Date"]=new["MEASDATE"].map(to_measurement_date)
new["Measurement_Time"]=new["MEASDATE"].map(to_measurement_time)
new["Calibration_Date"]=new["CALDATE"].map(to_calibration_date)
new["magS21"]=new["MS12"].map(to_linear)
new["Measurement_Type"]='2-port'
# step 4: delete any extra columns
del new["CALDATE"]
del new["MEASDATE"]
del new["CAL"]
del new["MS12"]
# step 5: add empty columns
empty_columns=[ u'Direction',u'System_Letter',
u'Connector_Type_Calibration', u'Program_Used', u'Program_Revision', u'Operator', u'Calibration_Name',
u'Number_Connects',
u'Number_Repeats', u'Nbs', u'Number_Frequencies', u'Start_Frequency',
u'Device_Description']
for empty_column in empty_columns:
new[empty_column]=None
# Now check that the column names are the same and order them
raw_columns=raw_two_port.columns
print raw_columns
new=new[raw_columns]
new_columns=new.columns
print new_columns
raw_columns==new_columns
# step 6: exclude any files that already exist
unique_timestamps=raw_two_port["Measurement_Timestamp"].unique()
new=new[-new["Measurement_Timestamp"].isin(unique_timestamps)]
# step7: add the files and save as csv, note at this point we can write to a db also
combined=pandas.concat([raw_two_port,new])
combined.to_csv(COMBINED_TWO_PORT_CHKSTD_CSV,index=False)
# Finally we check that the files are all what we expect
number_measurements_raw=len(raw_two_port["Measurement_Timestamp"].unique())
number_measurements_sas=len(sas_two_port["MEASDATE"].unique())
number_new=len(new["Measurement_Timestamp"].unique())
number_combined=len(combined["Measurement_Timestamp"].unique())
print("There were {0} measurements in the raw two port files".format(number_measurements_raw))
print("There were {0} measurements in the sas two port files".format(number_measurements_sas))
print("{0} measurements did not overlap".format(number_new))
print("This resulted in {0} unique measurements".format(number_combined))
print("The statement that the number of raw + non-overlapping measurements is equal to the number of "
"combined measurements is {0}, resulting in {1} rows of"
"data".format(number_new+number_measurements_raw==number_combined,len(combined)))
# Finaly check the data by importing it
start_time=datetime.datetime.now()
combined_csv=pandas.read_csv(COMBINED_TWO_PORT_CHKSTD_CSV)
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were imported as a single csv in {1} seconds".format(len(combined_csv),diff.total_seconds()))
number_standards=len(combined_csv["Device_Id"].unique())
print("The number of 2-port check standards is {0}".format(number_standards))
# ### Repeat for power
raw_power=pandas.read_csv(POWER_CHKSTD_CSV)
sas_power=pandas.read_csv(SAS_POWER)
print raw_power.columns
print sas_power.columns
unique_cal=raw_power["Calibration_Factor"].unique()
test=sas_power[-sas_power["KP"].isin(unique_cal)]
print test
raw_power["Efficiency"].unique()
# todo: make this a stand alone script
# step 1: import data sets
raw_power=pandas.read_csv(POWER_CHKSTD_CSV)
sas_power=pandas.read_csv(SAS_POWER)
# step 2: rename any columns that are the same with different names
same={"spid":"System_Id","SP":"Port_Used","ctype":"Connector_Type_Measurement","checkid":"Device_Id",
"MGAMA":"magS11","PGAMA":"argS11","EFF":"Efficiency","KP":"Calibration_Factor",
"CON":"Connect","FREQ":"Frequency"}
new=sas_power.rename(columns=same)
# step 3: create derived columns
def date_conversion(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.isoformat(" ")
def to_measurement_date(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%d %b %y")
def to_measurement_time(date_sas_format):
datetime_format=datetime.datetime.strptime(date_sas_format,'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%H:%M:%S")
def to_calibration_date(date_sas_format):
if type(date_sas_format) is StringType:
datetime_format=datetime.datetime.strptime(str(date_sas_format),'%d%b%y:%H:%M:%S')
return datetime_format.strftime("%d %b %y")
else:
return date_sas_format
new["Measurement_Timestamp"]=new["MEASDATE"].map(date_conversion)
new["Measurement_Date"]=new["MEASDATE"].map(to_measurement_date)
new["Measurement_Time"]=new["MEASDATE"].map(to_measurement_time)
new["Calibration_Date"]=new["CALDATE"].map(to_calibration_date)
new["Measurement_Type"]='power'
# step 4: delete any extra columns
del new["CALDATE"]
del new["MEASDATE"]
del new["CAL"]
# step 5: add empty columns
empty_columns=[ u'Direction', u'magS22',
u'argS22', u'System_Letter',
u'Connector_Type_Calibration', u'Program_Used', u'Program_Revision', u'Operator', u'Calibration_Name',
u'Number_Connects',
u'Number_Repeats', u'Nbs', u'Number_Frequencies', u'Start_Frequency',
u'Device_Description']
for empty_column in empty_columns:
new[empty_column]=None
# Now check that the column names are the same and order them
raw_columns=raw_power.columns
print raw_columns
new=new[raw_columns]
new_columns=new.columns
print new_columns
raw_columns==new_columns
# step 6: exclude any files that already exist
unique_timestamps=raw_power["Measurement_Timestamp"].unique()
new=new[-new["Measurement_Timestamp"].isin(unique_timestamps)]
# step7: add the files and save as csv, note at this point we can write to a db also
combined=pandas.concat([raw_power,new])
combined.to_csv(COMBINED_POWER_CHKSTD_CSV,index=False)
# Finally we check that the files are all what we expect
number_measurements_raw=len(raw_power["Measurement_Timestamp"].unique())
number_measurements_sas=len(sas_power["MEASDATE"].unique())
number_new=len(new["Measurement_Timestamp"].unique())
number_combined=len(combined["Measurement_Timestamp"].unique())
print("There were {0} measurements in the raw power files".format(number_measurements_raw))
print("There were {0} measurements in the sas power files".format(number_measurements_sas))
print("{0} measurements did not overlap".format(number_new))
print("This resulted in {0} unique measurements".format(number_combined))
print("The statement that the number of raw + non-overlapping measurements is equal to the number of "
"combined measurements is {0}, resulting in {1} rows of "
"data".format(number_new+number_measurements_raw==number_combined,len(combined)))
# Finaly check the data by importing it
start_time=datetime.datetime.now()
combined_csv=pandas.read_csv(COMBINED_POWER_CHKSTD_CSV)
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were imported as a single csv in {1} seconds".format(len(combined_csv),diff.total_seconds()))
number_standards=len(combined_csv["Device_Id"].unique())
print("The number of power check standards is {0}".format(number_standards))
# ### Now we import and shape the DUT files (.asc)
one_port_files=[]
two_port_files=[]
power_files=[]
for root,directory,file_names in os.walk(DUT_TOP_DIRECTORY):
#print file_names
for file_name in file_names:
match=re.search('.asc',file_name,re.IGNORECASE)
try:
if re.search('.txt',file_name,re.IGNORECASE):raise
if match:
in_file=open(os.path.join(root,file_name),'r')
contents=in_file.read()
in_file.close()
if re.search('table 1',contents,re.IGNORECASE) and re.search('table 2',contents,re.IGNORECASE) and re.search('table 3',contents,re.IGNORECASE):
two_port_files.append(os.path.join(root,file_name))
elif re.search('table 1',contents,re.IGNORECASE) and re.search('table 2',contents,re.IGNORECASE):
power_files.append(os.path.join(root,file_name))
elif re.search('table 1',contents,re.IGNORECASE):
one_port_files.append(os.path.join(root,file_name))
else:
pass
except:pass
# check the files
print("There are %s one port calrep files"%len(one_port_files))
print("There are %s two port calrep files"%len(two_port_files))
print("There are %s power calrep files"%len(power_files))
# We parse the file and extract Analysis_Date and Device_Id
start_time=datetime.datetime.now()
initial_file=OnePortCalrepModel(one_port_files[0])
device_id=initial_file.header[0].rstrip().lstrip()
print("{0} is {1}".format('device_id',device_id))
analysis_date=initial_file.header[1].rstrip().lstrip()
print("{0} is {1}".format('analysis_date',analysis_date))
initial_file.options["data_delimiter"]=","
initial_file.add_column(column_name='Device_Id',column_type='str',
column_data=[device_id for row in initial_file.data[:]])
initial_file.add_column(column_name='Analysis_Date',column_type='str',
column_data=[analysis_date for row in initial_file.data[:]])
#print initial_file
initial_file.header=None
initial_file.save(ONE_PORT_CALREP_CSV)
del initial_file
out_file=open(ONE_PORT_CALREP_CSV,'a')
file_list=one_port_files[1:]
for index,file_name in enumerate(file_list):
try:
print("Processing File Number {0}, {1}".format(index,file_name))
one_port_table=OnePortCalrepModel(file_name)
device_id=one_port_table.header[0].rstrip().lstrip()
analysis_date=one_port_table.header[1].rstrip().lstrip()
one_port_table.options["data_delimiter"]=","
one_port_table.add_column(column_name='Device_Id',
column_type='str',
column_data=[device_id for row in one_port_table.data[:]])
one_port_table.add_column(column_name='Analysis_Date',
column_type='str',
column_data=[analysis_date for row in one_port_table.data[:]])
#print one_port_table
out_file.write('\n')
data=one_port_table.get_data_string()
out_file.write(data)
print one_port_table.header
if index==len(file_list)-1:
print("Last File")
else:
print("Next file is {0}".format(one_port_files[index+1]))
except DataDimensionError:
print("{0} was passed due to a data dimensioning problem".format(file_name))
pass
except AttributeError:
print("{0} was passed due to a loading issue".format(file_name))
except TypeError:
print("{0} was passed due to an unkown issue".format(file_name))
except TypeConversionError:
print("{0} was passed due to improper number of columns".format(file_name))
except ValueError:
print("{0} was passed due to improper number of columns".format(file_name))
except:raise
out_file.close()
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were converted to a single csv in {1} seconds".format(len(file_list),diff.total_seconds()))
#check the data
one_port_calrep_data_frame=pandas.read_csv(ONE_PORT_CALREP_CSV)
one_port_calrep_data_frame[:20]
# +
# We parse the file and extract Analysis_Date and Device_Id
PRINT_REPORT=False
start_time=datetime.datetime.now()
initial_file=TwoPortCalrepModel(two_port_files[0])
device_id=initial_file.joined_table.header[0].rstrip().lstrip()
if PRINT_REPORT:
print("{0} is {1}".format('device_id',device_id))
try:
analysis_date=initial_file.joined_table.header[1].rstrip().lstrip()
except:
analysis_date=""
if PRINT_REPORT:
print("{0} is {1}".format('analysis_date',analysis_date))
initial_file.joined_table.options["data_delimiter"]=","
initial_file.joined_table.add_column(column_name='Device_Id',column_type='str',
column_data=[device_id for row in initial_file.joined_table.data[:]])
initial_file.joined_table.add_column(column_name='Analysis_Date',column_type='str',
column_data=[analysis_date for row in initial_file.joined_table.data[:]])
#print initial_file
initial_file.joined_table.header=None
initial_file.joined_table.save(TWO_PORT_CALREP_CSV)
del initial_file
out_file=open(TWO_PORT_CALREP_CSV,'a')
file_list=two_port_files[1:]
for index,file_name in enumerate(file_list):
try:
if PRINT_REPORT:
print("Processing File Number {0}, {1}".format(index,file_name))
two_port_table=TwoPortCalrepModel(file_name)
device_id=two_port_table.joined_table.header[0].rstrip().lstrip()
if PRINT_REPORT:
print("{0} is {1}".format('device_id',device_id))
try:
analysis_date=two_port_table.joined_table.header[1].rstrip().lstrip()
except:
analysis_date=""
if PRINT_REPORT:
print("{0} is {1}".format('analysis_date',analysis_date))
two_port_table.joined_table.options["data_delimiter"]=","
two_port_table.joined_table.add_column(column_name='Device_Id',column_type='str',
column_data=[device_id for row in two_port_table.joined_table.data[:]])
two_port_table.joined_table.add_column(column_name='Analysis_Date',column_type='str',
column_data=[analysis_date for row in two_port_table.joined_table.data[:]])
out_file.write("\n")
data=two_port_table.joined_table.get_data_string()
out_file.write(data)
if PRINT_REPORT:
print two_port_table.joined_table.header
if index==len(file_list)-1:
print("Last File")
else:
print("Next file is {0}".format(two_port_files[index+1]))
except DataDimensionError:
print("{0} was passed due to a data dimensioning problem".format(file_name))
pass
except AttributeError:
print("{0} was passed due to a loading issue".format(file_name))
except TypeError:
print("{0} was passed due to an unkown issue".format(file_name))
except TypeConversionError:
print("{0} was passed due to improper number of columns".format(file_name))
except ValueError:
print("{0} was passed due to improper number of columns".format(file_name))
except:raise
out_file.close()
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were converted to a single csv in {1} seconds".format(len(file_list),diff.total_seconds()))
# -
#check the data
two_port_calrep_data_frame=pandas.read_csv(TWO_PORT_CALREP_CSV)
two_port_calrep_data_frame[:10]
# First we separate the power list between the two types
power_3term=[]
power_4term=[]
for file_name in power_files:
try:
new_table=PowerCalrepModel(file_name)
number_columns=len(new_table.joined_table.data[0])
if number_columns == 19:
power_3term.append(file_name)
elif number_columns ==21:
power_4term.append(file_name)
else:
print("{0} does not conform".format(file_name))
except:
print("{0} caused an error".format(file_name))
pass
print("There are {0} three term files".format(len(power_3term)))
print("There are {0} four term files".format(len(power_4term)))
def power_calrep_to_csv_script(power_file_list,output_file,print_report=False):
""" Script converts all of the files in power_file_list to a single csv file (output_file).
Option to print a detailed report print_report=True."""
PRINT_REPORT=print_report
# start timer for analysis
start_time=datetime.datetime.now()
# seed file for format and column naems
initial_file=PowerCalrepModel(power_file_list[0])
# device id assumed to be the first line of header
device_id=initial_file.joined_table.header[0].rstrip().lstrip()
if PRINT_REPORT:
print("{0} is {1}".format('device_id',device_id))
# try and find the date in the header, since some of the dates are in different lines
# flatten the header and remove the device id. If it fails just make analysis date blank.
try:
header=string_list_collapse(initial_file.joined_table.header[:],string_delimiter="")
header=header.rstrip().lstrip().replace(device_id,"")
analysis_date=header
except:
analysis_date=""
if PRINT_REPORT:
print("{0} is {1}".format('analysis_date',analysis_date))
# insure that the data delimiter is a comma.
initial_file.joined_table.options["data_delimiter"]=","
# Add columns with device id and analysis date in them
initial_file.joined_table.add_column(column_name='Device_Id',column_type='str',
column_data=[device_id for row in initial_file.joined_table.data[:]])
initial_file.joined_table.add_column(column_name='Analysis_Date',column_type='str',
column_data=[analysis_date for row in initial_file.joined_table.data[:]])
#print initial_file
# remove the header for output purposes
initial_file.joined_table.header=None
# save the seed file with column names in csv format
initial_file.joined_table.save(output_file)
# clear the object from memory
del initial_file
# now the initial write is completed open the file in append mode and append each file
out_file=open(output_file,'a')
for index,file_name in enumerate(power_file_list):
try:
if PRINT_REPORT:
print("Processing File Number {0}, {1}".format(index,file_name))
table=PowerCalrepModel(file_name)
# device id assumed to be the first line of header
device_id=table.joined_table.header[0].rstrip().lstrip()
if PRINT_REPORT:
print("{0} is {1}".format('device_id',device_id))
# try and find the date in the header, since some of the dates are in different lines
# flatten the header and remove the device id. If it fails just make analysis date blank.
try:
header=string_list_collapse(table.joined_table.header[:],string_delimiter="")
header=header.rstrip().lstrip().replace(device_id,"")
analysis_date=header
except:
analysis_date=""
if PRINT_REPORT:
print("{0} is {1}".format('analysis_date',analysis_date))
# insure that the data delimiter is a comma.
print table.joined_table.header
print table.joined_table.column_names
print("Data is {0} rows x {1} columns".format(len(table.joined_table.data),
len(table.joined_table.data[0])))
if index==len(power_file_list)-1:
print("Last File")
else:
print("Next file is {0}".format(power_files[index+1]))
table.joined_table.options["data_delimiter"]=","
# Add columns with device id and analysis date in them
table.joined_table.add_column(column_name='Device_Id',column_type='str',
column_data=[device_id for row in table.joined_table.data[:]])
table.joined_table.add_column(column_name='Analysis_Date',column_type='str',
column_data=[analysis_date for row in table.joined_table.data[:]])
# write to out_file
out_file.write("\n")
data=table.joined_table.get_data_string()
out_file.write(data)
except DataDimensionError:
print("{0} was passed due to a data dimensioning problem".format(file_name))
pass
except AttributeError:
print("{0} was passed due to a loading issue".format(file_name))
except ValueError:
print("{0} was passed due to a column size issue".format(file_name))
except:raise
# Close out the script
out_file.close()
stop_time=datetime.datetime.now()
diff=stop_time-start_time
print("{0} files were converted to a single csv in {1} seconds".format(len(power_file_list),diff.total_seconds()))
# run the script for the two power types
power_calrep_to_csv_script(power_3term,POWER_3TERM_CALREP_CSV,print_report=False)
power_calrep_to_csv_script(power_4term,POWER_4TERM_CALREP_CSV)
|
Documentation/Examples/jupyter/Data_Transformation_Check_Standard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting up to speed with Dask
#
# ## Part 0: Getting data
#
# We are using the [NYC Taxi data](https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page), which contains several publicly-available datasets about taxi and ride-share rides taken in New York City.
#
# Data is available from 2009 to 2019, but for this exercise we will use 2019 data only. Take care when using other data, as the schemas in the CSV files changed over the years. Most notably, in mid-2016 latitude and longitude fields were replaced with more generic taxi zones for privacy reasons.
#
# Files are hosted in this S3 location: `s3://nyc-tlc/trip data`. Both Dask and Pandas support reading directly from S3 with some slight nuance, but for simplicity we will download the data for the laptop examples (Parts 1 & 2). This requires about 8GB of disk space.
import s3fs
import numpy as np
from pathlib import Path
# change this path if you don't want it here
data_path = Path('data')
data_path.mkdir(exist_ok=True)
# +
fs = s3fs.S3FileSystem(anon=True)
files = fs.glob('s3://nyc-tlc/trip data/yellow_tripdata_*')
len(files), files[:5], files[-5:]
# -
# <br>
# One file per month, approximately 8GB disk size
files_2019 = fs.glob('s3://nyc-tlc/trip data/yellow_tripdata_2019-*.csv')
len(files_2019), np.sum([fs.du(f) for f in files_2019]) / 1e9
# +
# %%time
def download_file(f):
fs.get(f, data_path/Path(f).name)
for f in files_2019:
download_file(f)
# -
# # Dask sneak peak!
#
# You can parallelize this file copy using [dask.bag](https://docs.dask.org/en/latest/bag.html)
import dask.bag as db
# %%time
_ = db.from_sequence(files_2019).map(download_file).compute()
|
0_get_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.naive_bayes import GaussianNB
# Read in the data
#Data = pd.read_csv('Full_Data.csv', encoding = "ISO-8859-1")
#Data.head(1)
data = pd.read_csv('Full_Data.csv', encoding = "ISO-8859-1")
data.head(1)
train = data[data['Date'] < '20150101']
test = data[data['Date'] > '20141231']
# +
# Removing punctuations
slicedData= train.iloc[:,2:27]
slicedData.replace(to_replace="[^a-zA-Z]", value=" ", regex=True, inplace=True)
# Renaming column names for ease of access
list1= [i for i in range(25)]
new_Index=[str(i) for i in list1]
slicedData.columns= new_Index
slicedData.head(5)
# Convertng headlines to lower case
for index in new_Index:
slicedData[index]=slicedData[index].str.lower()
slicedData.head(1)
# -
headlines = []
for row in range(0,len(slicedData.index)):
headlines.append(' '.join(str(x) for x in slicedData.iloc[row,0:25]))
headlines[0]
basicvectorizer = CountVectorizer(ngram_range=(1,1))
basictrain = basicvectorizer.fit_transform(headlines)
basicmodel = GaussianNB()
basicmodel = basicmodel.fit(basictrain.toarray(), train["Label"])
testheadlines = []
for row in range(0,len(test.index)):
testheadlines.append(' '.join(str(x) for x in test.iloc[row,2:27]))
basictest = basicvectorizer.transform(testheadlines)
predictions = basicmodel.predict(basictest.toarray())
predictions
pd.crosstab(test["Label"], predictions, rownames=["Actual"], colnames=["Predicted"])
# +
print(basictrain.shape)
from sklearn.metrics import classification_report
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
print (classification_report(test["Label"], predictions))
print (accuracy_score(test["Label"], predictions))
# -
basicvectorizer2 = CountVectorizer(ngram_range=(2,2))
basictrain2 = basicvectorizer2.fit_transform(headlines)
basicmodel2 = GaussianNB()
basicmodel2 = basicmodel2.fit(basictrain2.toarray(), train["Label"])
basictest2 = basicvectorizer2.transform(testheadlines)
predictions2 = basicmodel2.predict(basictest2)
# +
print(basictrain2.shape)
pd.crosstab(test["Label"], predictions2, rownames=["Actual"], colnames=["Predicted"])
print (classification_report(test["Label"], predictions2))
print (accuracy_score(test["Label"], predictions2))
# -
basicvectorizer3 = CountVectorizer(ngram_range=(3,3))
basictrain3 = basicvectorizer3.fit_transform(headlines)
basicmodel3 = GaussianNB()
basicmodel3 = basicmodel3.fit(basictrain3, train["Label"])
basictest3 = basicvectorizer3.transform(testheadlines)
predictions3 = basicmodel3.predict(basictest3)
# +
print(basictrain3.shape)
pd.crosstab(test["Label"], predictions3, rownames=["Actual"], colnames=["Predicted"])
print (classification_report(test["Label"], predictions3))
print (accuracy_score(test["Label"], predictions3))
|
Naive Bayes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Partial Dependence Plot
#
# ## Summary
#
# Partial dependence plots visualize the dependence between the response and a set of target features (usually one or two), marginalizing over all the other features. For a perturbation-based interpretability method, it is relatively quick. PDP assumes independence between the features, and can be misleading interpretability-wise when this is not met (e.g. when the model has many high order interactions).
#
# ## How it Works
#
# The PDP module for `scikit-learn` {cite}`pedregosa2011scikit` provides a succinct description of the algorithm [here](https://scikit-learn.org/stable/modules/partial_dependence.html).
#
# <NAME>'s "Interpretable Machine Learning" e-book {cite}`molnar2020interpretable` has an excellent overview on partial dependence that can be found [here](https://christophm.github.io/interpretable-ml-book/pdp.html).
#
# The conceiving paper "Greedy Function Approximation: A Gradient Boosting Machine" {cite}`friedman2001greedy` provides a good motivation and definition.
# ## Code Example
#
# The following code will train a blackbox pipeline for the breast cancer dataset. Aftewards it will interpret the pipeline and its decisions with Partial Dependence Plots. The visualizations provided will be for global explanations.
from interpret import set_visualize_provider
from interpret.provider import InlineProvider
set_visualize_provider(InlineProvider())
# +
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.decomposition import PCA
from sklearn.pipeline import Pipeline
from interpret import show
from interpret.blackbox import PartialDependence
seed = 1
X, y = load_breast_cancer(return_X_y=True, as_frame=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=seed)
pca = PCA()
rf = RandomForestClassifier(n_estimators=100, n_jobs=-1)
blackbox_model = Pipeline([('pca', pca), ('rf', rf)])
blackbox_model.fit(X_train, y_train)
pdp = PartialDependence(predict_fn=blackbox_model.predict_proba, data=X_train)
pdp_global = pdp.explain_global()
show(pdp_global)
# -
# ## Further Resources
#
# - [Paper link to conceiving paper](https://projecteuclid.org/download/pdf_1/euclid.aos/1013203451)
# - [scikit-learn on their PDP module](https://scikit-learn.org/stable/modules/partial_dependence.html)
# ## Bibliography
#
# ```{bibliography} references.bib
# :style: unsrt
# :filter: docname in docnames
# ```
# ## API
#
# ### PartialDependence
#
# ```{eval-rst}
# .. autoclass:: interpret.blackbox.PartialDependence
# :members:
# :inherited-members:
# ```
|
docs/interpret_docs/pdp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="rNdWfPXCjTjY"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="I1dUQ0GejU8N"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="c05P9g5WjizZ"
# # 对结构化数据进行分类
# + [markdown] colab_type="text" id="zofH_gCzgplN"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://tensorflow.google.cn/tutorials/structured_data/feature_columns">
# <img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />
# 在 tensorflow.google.cn 上查看</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/structured_data/feature_columns.ipynb">
# <img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />
# 在 Google Colab 运行</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/structured_data/feature_columns.ipynb">
# <img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />
# 在 Github 上查看源代码</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/structured_data/feature_columns.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载此 notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="Zz7U1iGCwp7T"
# Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
# [官方英文文档](https://tensorflow.google.cn/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
# [tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
# [<EMAIL> Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
# + [markdown] colab_type="text" id="K1y4OHpGgss7"
# 本教程演示了如何对结构化数据进行分类(例如,CSV 中的表格数据)。我们将使用 [Keras](https://tensorflow.google.cn/guide/keras) 来定义模型,将[特征列(feature columns)](https://tensorflow.google.cn/guide/feature_columns) 作为从 CSV 中的列(columns)映射到用于训练模型的特征(features)的桥梁。本教程包括了以下内容的完整代码:
#
# * 用 [Pandas](https://pandas.pydata.org/) 导入 CSV 文件。
# * 用 [tf.data](https://tensorflow.google.cn/guide/datasets) 建立了一个输入流水线(pipeline),用于对行进行分批(batch)和随机排序(shuffle)。
# * 用特征列将 CSV 中的列映射到用于训练模型的特征。
# * 用 Keras 构建,训练并评估模型。
#
# ## 数据集
#
# 我们将使用一个小型 [数据集](https://archive.ics.uci.edu/ml/datasets/heart+Disease),该数据集由克利夫兰心脏病诊所基金会(Cleveland Clinic Foundation for Heart Disease)提供。CSV 中有几百行数据。每行描述了一个病人(patient),每列描述了一个属性(attribute)。我们将使用这些信息来预测一位病人是否患有心脏病,这是在该数据集上的二分类任务。
#
# 下面是该数据集的[描述](https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/heart-disease.names)。 请注意,有数值(numeric)和类别(categorical)类型的列。
#
# >列| 描述| 特征类型 | 数据类型
# >------------|--------------------|----------------------|-----------------
# >Age | 年龄以年为单位 | Numerical | integer
# >Sex | (1 = 男;0 = 女) | Categorical | integer
# >CP | 胸痛类型(0,1,2,3,4)| Categorical | integer
# >Trestbpd | 静息血压(入院时,以mm Hg计) | Numerical | integer
# >Chol | 血清胆固醇(mg/dl) | Numerical | integer
# >FBS |(空腹血糖> 120 mg/dl)(1 = true;0 = false)| Categorical | integer
# >RestECG | 静息心电图结果(0,1,2)| Categorical | integer
# >Thalach | 达到的最大心率 | Numerical | integer
# >Exang | 运动诱发心绞痛(1 =是;0 =否)| Categorical | integer
# >Oldpeak | 与休息时相比由运动引起的 ST 节段下降|Numerical | integer
# >Slope | 在运动高峰 ST 段的斜率 | Numerical | float
# >CA | 荧光透视法染色的大血管动脉(0-3)的数量 | Numerical | integer
# >Thal | 3 =正常;6 =固定缺陷;7 =可逆缺陷 | Categorical | string
# >Target | 心脏病诊断(1 = true;0 = false) | Classification | integer
# + [markdown] colab_type="text" id="VxyBFc_kKazA"
# ## 导入 TensorFlow 和其他库
# + colab={} colab_type="code" id="LuOWVJBz8a6G"
# !pip install sklearn
# + colab={} colab_type="code" id="9dEreb4QKizj"
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import feature_column
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
# + [markdown] colab_type="text" id="KCEhSZcULZ9n"
# ## 使用 Pandas 创建一个 dataframe
#
# [Pandas](https://pandas.pydata.org/) 是一个 Python 库,它有许多有用的实用程序,用于加载和处理结构化数据。我们将使用 Pandas 从 URL下载数据集,并将其加载到 dataframe 中。
# + colab={} colab_type="code" id="REZ57BXCLdfG"
URL = 'https://storage.googleapis.com/applied-dl/heart.csv'
dataframe = pd.read_csv(URL)
dataframe.head()
# + [markdown] colab_type="text" id="u0zhLtQqMPem"
# ## 将 dataframe 拆分为训练、验证和测试集
#
# 我们下载的数据集是一个 CSV 文件。 我们将其拆分为训练、验证和测试集。
# + colab={} colab_type="code" id="YEOpw7LhMYsI"
train, test = train_test_split(dataframe, test_size=0.2)
train, val = train_test_split(train, test_size=0.2)
print(len(train), 'train examples')
print(len(val), 'validation examples')
print(len(test), 'test examples')
# + [markdown] colab_type="text" id="84ef46LXMfvu"
# ## 用 tf.data 创建输入流水线
#
# 接下来,我们将使用 [tf.data](https://tensorflow.google.cn/guide/datasets) 包装 dataframe。这让我们能将特征列作为一座桥梁,该桥梁将 Pandas dataframe 中的列映射到用于训练模型的特征。如果我们使用一个非常大的 CSV 文件(非常大以至于它不能放入内存),我们将使用 tf.data 直接从磁盘读取它。本教程不涉及这一点。
# + colab={} colab_type="code" id="NkcaMYP-MsRe"
# 一种从 Pandas Dataframe 创建 tf.data 数据集的实用程序方法(utility method)
def df_to_dataset(dataframe, shuffle=True, batch_size=32):
dataframe = dataframe.copy()
labels = dataframe.pop('target')
ds = tf.data.Dataset.from_tensor_slices((dict(dataframe), labels))
if shuffle:
ds = ds.shuffle(buffer_size=len(dataframe))
ds = ds.batch(batch_size)
return ds
# + colab={} colab_type="code" id="CXbbXkJvMy34"
batch_size = 5 # 小批量大小用于演示
train_ds = df_to_dataset(train, batch_size=batch_size)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
test_ds = df_to_dataset(test, shuffle=False, batch_size=batch_size)
# + [markdown] colab_type="text" id="qRLGSMDzM-dl"
# ## 理解输入流水线
#
# 现在我们已经创建了输入流水线,让我们调用它来查看它返回的数据的格式。 我们使用了一小批量大小来保持输出的可读性。
# + colab={} colab_type="code" id="CSBo3dUVNFc9"
for feature_batch, label_batch in train_ds.take(1):
print('Every feature:', list(feature_batch.keys()))
print('A batch of ages:', feature_batch['age'])
print('A batch of targets:', label_batch )
# + [markdown] colab_type="text" id="OT5N6Se-NQsC"
# 我们可以看到数据集返回了一个字典,该字典从列名称(来自 dataframe)映射到 dataframe 中行的列值。
# + [markdown] colab_type="text" id="ttIvgLRaNoOQ"
# ## 演示几种特征列
# TensorFlow 提供了多种特征列。本节中,我们将创建几类特征列,并演示特征列如何转换 dataframe 中的列。
# + colab={} colab_type="code" id="mxwiHFHuNhmf"
# 我们将使用该批数据演示几种特征列
example_batch = next(iter(train_ds))[0]
# + colab={} colab_type="code" id="0wfLB8Q3N3UH"
# 用于创建一个特征列
# 并转换一批次数据的一个实用程序方法
def demo(feature_column):
feature_layer = layers.DenseFeatures(feature_column)
print(feature_layer(example_batch).numpy())
# + [markdown] colab_type="text" id="Q7OEKe82N-Qb"
# ### 数值列
# 一个特征列的输出将成为模型的输入(使用上面定义的 demo 函数,我们将能准确地看到 dataframe 中的每列的转换方式)。 [数值列(numeric column)](https://tensorflow.google.cn/api_docs/python/tf/feature_column/numeric_column) 是最简单的列类型。它用于表示实数特征。使用此列时,模型将从 dataframe 中接收未更改的列值。
# + colab={} colab_type="code" id="QZTZ0HnHOCxC"
age = feature_column.numeric_column("age")
demo(age)
# + [markdown] colab_type="text" id="7a6ddSyzOKpq"
# 在这个心脏病数据集中,dataframe 中的大多数列都是数值列。
# + [markdown] colab_type="text" id="IcSxUoYgOlA1"
# ### 分桶列
# 通常,您不希望将数字直接输入模型,而是根据数值范围将其值分成不同的类别。考虑代表一个人年龄的原始数据。我们可以用 [分桶列(bucketized column)](https://tensorflow.google.cn/api_docs/python/tf/feature_column/bucketized_column)将年龄分成几个分桶(buckets),而不是将年龄表示成数值列。请注意下面的 one-hot 数值表示每行匹配的年龄范围。
# + colab={} colab_type="code" id="wJ4Wt3SAOpTQ"
age_buckets = feature_column.bucketized_column(age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])
demo(age_buckets)
# + [markdown] colab_type="text" id="r1tArzewPb-b"
# ### 分类列
# 在此数据集中,thal 用字符串表示(如 'fixed','normal',或 'reversible')。我们无法直接将字符串提供给模型。相反,我们必须首先将它们映射到数值。分类词汇列(categorical vocabulary columns)提供了一种用 one-hot 向量表示字符串的方法(就像您在上面看到的年龄分桶一样)。词汇表可以用 [categorical_column_with_vocabulary_list](https://tensorflow.google.cn/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_list) 作为 list 传递,或者用 [categorical_column_with_vocabulary_file](https://tensorflow.google.cn/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_file) 从文件中加载。
# + colab={} colab_type="code" id="DJ6QnSHkPtOC"
thal = feature_column.categorical_column_with_vocabulary_list(
'thal', ['fixed', 'normal', 'reversible'])
thal_one_hot = feature_column.indicator_column(thal)
demo(thal_one_hot)
# + [markdown] colab_type="text" id="dxQloQ9jOoXL"
# 在更复杂的数据集中,许多列都是分类列(如 strings)。在处理分类数据时,特征列最有价值。尽管在该数据集中只有一列分类列,但我们将使用它来演示在处理其他数据集时,可以使用的几种重要的特征列。
# + [markdown] colab_type="text" id="LEFPjUr6QmwS"
# ### 嵌入列
# 假设我们不是只有几个可能的字符串,而是每个类别有数千(或更多)值。 由于多种原因,随着类别数量的增加,使用 one-hot 编码训练神经网络变得不可行。我们可以使用嵌入列来克服此限制。[嵌入列(embedding column)](https://tensorflow.google.cn/api_docs/python/tf/feature_column/embedding_column)将数据表示为一个低维度密集向量,而非多维的 one-hot 向量,该低维度密集向量可以包含任何数,而不仅仅是 0 或 1。嵌入的大小(在下面的示例中为 8)是必须调整的参数。
#
# 关键点:当分类列具有许多可能的值时,最好使用嵌入列。我们在这里使用嵌入列用于演示目的,为此您有一个完整的示例,以在将来可以修改用于其他数据集。
# + colab={} colab_type="code" id="hSlohmr2Q_UU"
# 注意到嵌入列的输入是我们之前创建的类别列
thal_embedding = feature_column.embedding_column(thal, dimension=8)
demo(thal_embedding)
# + [markdown] colab_type="text" id="urFCAvTVRMpB"
# ### 经过哈希处理的特征列
#
# 表示具有大量数值的分类列的另一种方法是使用 [categorical_column_with_hash_bucket](https://tensorflow.google.cn/api_docs/python/tf/feature_column/categorical_column_with_hash_bucket)。该特征列计算输入的一个哈希值,然后选择一个 `hash_bucket_size` 分桶来编码字符串。使用此列时,您不需要提供词汇表,并且可以选择使 hash_buckets 的数量远远小于实际类别的数量以节省空间。
#
# 关键点:该技术的一个重要缺点是可能存在冲突,不同的字符串被映射到同一个范围。实际上,无论如何,经过哈希处理的特征列对某些数据集都有效。
# + colab={} colab_type="code" id="YHU_Aj2nRRDC"
thal_hashed = feature_column.categorical_column_with_hash_bucket(
'thal', hash_bucket_size=1000)
demo(feature_column.indicator_column(thal_hashed))
# + [markdown] colab_type="text" id="fB94M27DRXtZ"
# ### 组合的特征列
# 将多种特征组合到一个特征中,称为[特征组合(feature crosses)](https://developers.google.com/machine-learning/glossary/#feature_cross),它让模型能够为每种特征组合学习单独的权重。此处,我们将创建一个 age 和 thal 组合的新特征。请注意,`crossed_column` 不会构建所有可能组合的完整列表(可能非常大)。相反,它由 `hashed_column` 支持,因此您可以选择表的大小。
# + colab={} colab_type="code" id="oaPVERd9Rep6"
crossed_feature = feature_column.crossed_column([age_buckets, thal], hash_bucket_size=1000)
demo(feature_column.indicator_column(crossed_feature))
# + [markdown] colab_type="text" id="ypkI9zx6Rj1q"
# ## 选择要使用的列
# 我们已经了解了如何使用几种类型的特征列。 现在我们将使用它们来训练模型。本教程的目标是向您展示使用特征列所需的完整代码(例如,机制)。我们任意地选择了几列来训练我们的模型。
#
# 关键点:如果您的目标是建立一个准确的模型,请尝试使用您自己的更大的数据集,并仔细考虑哪些特征最有意义,以及如何表示它们。
# + colab={} colab_type="code" id="4PlLY7fORuzA"
feature_columns = []
# 数值列
for header in ['age', 'trestbps', 'chol', 'thalach', 'oldpeak', 'slope', 'ca']:
feature_columns.append(feature_column.numeric_column(header))
# 分桶列
age_buckets = feature_column.bucketized_column(age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])
feature_columns.append(age_buckets)
# 分类列
thal = feature_column.categorical_column_with_vocabulary_list(
'thal', ['fixed', 'normal', 'reversible'])
thal_one_hot = feature_column.indicator_column(thal)
feature_columns.append(thal_one_hot)
# 嵌入列
thal_embedding = feature_column.embedding_column(thal, dimension=8)
feature_columns.append(thal_embedding)
# 组合列
crossed_feature = feature_column.crossed_column([age_buckets, thal], hash_bucket_size=1000)
crossed_feature = feature_column.indicator_column(crossed_feature)
feature_columns.append(crossed_feature)
# + [markdown] colab_type="text" id="M-nDp8krS_ts"
# ### 建立一个新的特征层
# 现在我们已经定义了我们的特征列,我们将使用[密集特征(DenseFeatures)](https://tensorflow.google.cn/versions/r2.0/api_docs/python/tf/keras/layers/DenseFeatures)层将特征列输入到我们的 Keras 模型中。
# + colab={} colab_type="code" id="6o-El1R2TGQP"
feature_layer = tf.keras.layers.DenseFeatures(feature_columns)
# + [markdown] colab_type="text" id="8cf6vKfgTH0U"
# 之前,我们使用一个小批量大小来演示特征列如何运转。我们将创建一个新的更大批量的输入流水线。
# + colab={} colab_type="code" id="gcemszoGSse_"
batch_size = 32
train_ds = df_to_dataset(train, batch_size=batch_size)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
test_ds = df_to_dataset(test, shuffle=False, batch_size=batch_size)
# + [markdown] colab_type="text" id="bBx4Xu0eTXWq"
# ## 创建,编译和训练模型
# + colab={} colab_type="code" id="_YJPPb3xTPeZ"
model = tf.keras.Sequential([
feature_layer,
layers.Dense(128, activation='relu'),
layers.Dense(128, activation='relu'),
layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'],
run_eagerly=True)
model.fit(train_ds,
validation_data=val_ds,
epochs=5)
# + colab={} colab_type="code" id="GnFmMOW0Tcaa"
loss, accuracy = model.evaluate(test_ds)
print("Accuracy", accuracy)
# + [markdown] colab_type="text" id="3bdfbq20V6zu"
# 关键点:通常使用更大更复杂的数据集进行深度学习,您将看到最佳结果。使用像这样的小数据集时,我们建议使用决策树或随机森林作为强有力的基准。本教程的目的不是训练一个准确的模型,而是演示处理结构化数据的机制,这样,在将来使用自己的数据集时,您有可以使用的代码作为起点。
# + [markdown] colab_type="text" id="SotnhVWuHQCw"
# ### 下一步
# 了解有关分类结构化数据的更多信息的最佳方法是亲自尝试。我们建议寻找另一个可以使用的数据集,并使用和上面相似的代码,训练一个模型,对其分类。要提高准确率,请仔细考虑模型中包含哪些特征,以及如何表示这些特征。
|
site/zh-cn/tutorials/structured_data/feature_columns.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Strings in Python
#
# Programming languages refer to text as "strings" since text exists in the computer as a string of characters.
#
# To define a string variable you need to enclose the text in quotes.
msg = "Good Morning" # this is a string variable
print(msg)
# You can access each character in a string individually. To do this you need to specify which location you are wanting to access. Keep in mind that, as with most things in programming, locations start counting from 0.
msg = "Good Morning"
print(msg)
print(msg[0])
print(msg[6])
# Use the `len()` command to get the length of the string. The length of the string is a count of the number of characters in the string, not the index of the last location. `len()` returns an integer value that you can store in a variable for later use.
# +
msg = "Good Morning"
print(len(msg))
s = "0123456789"
print(len(s))
s_length = len(s)
print(s_length)
# -
# If you try to access a location in a string that does not exist (such as a location after the end of the string) then you will get an error.
msg = "Good Morning"
print(msg[30])
# Python allows you to have negative indexes for string locations. That means if you use a negative value for the location in the string, then you count back from the end. The last location is -1, the second to last location is -2, and so on.
msg = "Good Morning"
print(msg[-1])
print(msg[-2])
# You can get portions of a string that are more than just one letter. This is called "slicing" (some languages call it "substring").
#
# This command returns the portion of the string `string_name` starting from location `start` and ending at the location just before location `end`. It is important to note that the substring will not include the location `end`.
#
# `string_name[start:end]`
#
# If you don't put anything before the colon then the slice will start from the beginning of the string.
#
# `string_name[:end]`
#
# If you don't put anything after the colon then the slice will continue through the end of the string.
#
# `string_name[start:]`
msg = "Good Morning"
print(msg[0:4]) # print locations 0, 1, 2, and 3
print(msg[:4]) # slices starting at the beginning of the string
print(msg[5:]) # slices all the way to the end
print(msg[5:-1]) # does not slice all the way to the end
# Python can search a string for a letter or a substring and return the location it occurs in the string. If it is not found, then a -1 is returned. Be careful, the -1 means it's not found, not an index of -1 (the last location).
#
# `find` will return the location of the first occurance of what you are searching for because it starts searching from the beginning of the string.
#
# `rfind` (reverse find) will return the location of the last occurrance of what you are searching for because it starts searching from the end of the string to the beginning.
#
# Both of these return an integer value that can be stored in a variable for later use.
# +
msg = "Good Morning"
print( msg.find("o"))
print( msg.rfind("o"))
space = msg.find(" ")
print("The space is at location", space)
print( msg.find("Mor"))
# -
# You can combine multiple strings into a single string using a process called "concatenation". The `+` is used for concatenation.
# +
one = "Computer"
two = "Science"
print(one, two)
three = one + two
print(three)
three = one + " " + two
print(three)
space = three.find(" ")
swapped = three[space+1:] + " " + three[:space]
print(swapped)
# -
# You can only concatenate strings with other strings. You cannot concatenate strings with numbers. If you try, you will get the error you see below.
print("num =", 5) # this works
s = "num = " + 5 # this doesn't work
# There are times when you will want to concatenate a string and a number. To do this you need to tell Python to treat the number as a string (this is called "casting").
s = "num = " + str(5)
print(s)
# Below are some examples of using strings in methods.
# +
# print the string s one letter on each line
def vprint( s ):
for i in range( len(s) ):
print( s[i] )
# return the string s reversed
def reverse( s ):
rev = ""
for i in range( len(s) ):
rev = rev + s[-i-1]
return rev
### main program ###
vprint("compsci")
print( reverse("compsci") )
# -
|
python_strings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Bluelord/ML_Mastery_Python/blob/main/08_Algorithm_Evaluation_Metrics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Q72eP95wY8Uo"
# # ML Algorithm Performance Matrics**
#
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="A8Nvqlhs2bML" outputId="2ec99e6a-7a65-4891-f504-18c326d3ee59"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="fhIE9wwOZGJB"
# ## Classification Metric
#
# + [markdown] id="y4t94b04c6-Y"
# ### Classification Accuracy**
#
# It is the most common evalution matrics which shows the number of correct presictions made as a ratio of all predictions made. **This is only valid for the equal number of observation in each class**.
#
# + id="nS2yOgIFY6yQ" colab={"base_uri": "https://localhost:8080/"} outputId="0ef69715-8f08-4f32-a723-08b5e6ac0590"
# Evaluation using a train and a test set
from pandas import read_csv
from sklearn.model_selection import KFold # This testing is used to demonstrate each metric (You can use other resamling methods)
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
# Load Data
filename = "/content/drive/MyDrive/Colab Notebooks/ML Mastery python/Dataset/pima-indians-diabetes.csv"
names = ['preg', 'plas','pres','skin','test','mass','pedi','age','class']
dataframe = read_csv(filename, names=names)
array = dataframe.values
# separate array into input & output
X = array[:,0:8]
Y = array[:,8]
kfold = KFold(n_splits=10)
model = LogisticRegression(solver='liblinear')
scoring = 'accuracy' # this string is used in cross-validation for scoring
results = cross_val_score(model, X, Y, cv=kfold, scoring=scoring)
print("Accuracy: % .3f (% .3f)" %(results.mean(), results.std()))
# + [markdown] id="iYe-tsZ-inZI"
# ### Logarithmic Loss
#
# This evaluate the prediction of probabilities of membership in a given class. Predictions that are correct or incorrect are rewarded or punished proportionally to the confidence of the prediction.
#
# + id="08EJUA_VihTM" colab={"base_uri": "https://localhost:8080/"} outputId="46fffe2c-d802-46bd-9154-1157e9ce0802"
# Cross Validation Classification LogLoss
#
#
scoring = 'neg_log_loss' # Logarithmic Loss string is used in cross-validation for scoring
results = cross_val_score(model, X, Y, cv=kfold, scoring=scoring)
print("Accuracy: % .3f (% .3f)" %(results.mean(), results.std()))
# + [markdown] id="EQI2ru5xinyf"
# ### Area Under ROC curve
#
# **AUC** is matric used for binary classification problems, this discriminate between positive & negative classes. ROC is devided into **Sensiitivity** & **Specificity**.
# Sensitivity is the true postive rate ( Recall): It is the number of instances from the 1st class that we are predicting as correct.
# Specificity is the true negative rate: It is the instances when 2nd class is predicted correctly.
# + id="VPPVvHwPioLf" colab={"base_uri": "https://localhost:8080/"} outputId="de188606-964a-4e9a-f7da-90f5d8e6ee92"
# Cross Validation Classification ROC AUC
#
#
scoring = 'roc_auc' # Area Under ROC curve is used in cross-validation for scoring
results = cross_val_score(model, X, Y, cv=kfold, scoring=scoring)
print("Accuracy: % .3f (% .3f)" %(results.mean(), results.std()))
# + [markdown] id="YltugKtHio6f"
# ### Confusion Matrix
#
# This Matrix is a handy presentation of the accuracy of a model with 2 or more classes.The table of the matrix shows prediction on x-axis & true outcomes on y-axis.
# + id="WnVimGw9ipU_" colab={"base_uri": "https://localhost:8080/"} outputId="835e0e8c-2a7b-4360-ada6-850048d5adac"
# Cross Validation Classification confusion Matrix
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix
# Load Data
filename = "/content/drive/MyDrive/Colab Notebooks/ML Mastery python/Dataset/pima-indians-diabetes.csv"
names = ['preg', 'plas','pres','skin','test','mass','pedi','age','class']
dataframe = read_csv(filename, names=names)
array = dataframe.values
# separate array into input & output
X = array[:,0:8]
Y = array[:,8]
test_size = 0.33
seed = 7
X_train, X_test, Y_train, Y_test = train_test_split(X,Y, test_size=test_size, random_state=seed)
model = LogisticRegression(solver= 'liblinear')
model.fit(X_train, Y_train)
####################################
predicted = model.predict(X_test)
matrix = confusion_matrix(Y_test, predicted)
print(matrix)
# + [markdown] id="x6-qqZ0jqiV2"
# ### Classification Report
#
# SKlearn provides us with a report which gives a quick idea of accuracuy of model using a number of measures, this can also be used for multiclass classification.
#
# + id="5w9jTZZ9qQGh" colab={"base_uri": "https://localhost:8080/"} outputId="c5f198a8-e769-430e-a8a3-10cf68ca942f"
# Cross Validation Classification Report
from sklearn.metrics import classification_report
#
#
#
predicted = model.predict(X_test)
report = classification_report(Y_test, predicted)
print(report)
# + [markdown] id="JyNYgnhasE2Y"
# ## **Regression Metrics**
# + [markdown] id="lE4VeIERsMZW"
# ### Mean Absolute Error
#
# MEA is a sum abasolute difference between predictions & actual values. This gives the magnitude of error, but no idea about the direction (wetheer it is less or more the value)
#
# + id="4Bi7BJkHsnUG" colab={"base_uri": "https://localhost:8080/"} outputId="3aad503c-4ee6-4046-ddbf-6d8e4d6cf285"
# Evaluation using a train and a test set
from pandas import read_csv
from sklearn.model_selection import KFold # This testing is used to demonstrate each metric (You can use other resamling methods)
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LinearRegression
# Load Data
filename = "/content/drive/MyDrive/Colab Notebooks/ML Mastery python/Dataset/Boston-housing.csv"
#names = ['CRIM','ZN','INDUS','CHAS','NOX','RM','AGE','DIS','RAD','TAX','PTRATIO','B','LSTAT','MEDV']
dataframe = read_csv(filename)
array = dataframe.values
X = array[:,0:13]
Y = array[:,13]
kfold = KFold(n_splits=10)
model = LinearRegression()
scoring = 'neg_mean_absolute_error'
result = cross_val_score(model, X, Y, cv=kfold, scoring=scoring)
print("MAE: % .3f (% .3f)" % (result.mean(), result.std()))
# + [markdown] id="2vgOSMaHqjK1"
# ### Mean Squared Error
#
# MES is much like MAE, this converts the values back to the original unit of the output variable & can be described and presented properly.
#
# + id="o-e3u6z3spK-" colab={"base_uri": "https://localhost:8080/"} outputId="f506c7ad-07d1-48cb-9a71-54221db49557"
# Cross Validation Regression MES
#
#
scoring = 'neg_mean_squared_error'
results = cross_val_score(model, X, Y, cv=kfold, scoring=scoring)
print("MES: % .3f (% .3f)" %(results.mean(), results.std()))
# + [markdown] id="iN7sdTy3spoW"
# ## R Squared
#
# R Squared Matric provides an indications of goodness of fit of a set of predictions to the actual values. More the value (0-1) more is the fitting of cure to the model.
#
# + id="oaqt5OHgsqp_" colab={"base_uri": "https://localhost:8080/"} outputId="fece56ee-c520-4ae5-acac-d346b03589ad"
# Cross Validation Regression MES
#
#
scoring = 'r2'
results = cross_val_score(model, X, Y, cv=kfold, scoring=scoring)
print("R^2: % .3f (% .3f)" %(results.mean(), results.std()))
|
Lessons/08_Algorithm_Evaluation_Metrics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/victorknox/Hate-Speech-Detection-in-Hindi/blob/main/Hate_Speech_Detection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="XeHmXPtnwN8g"
# # Subjectivity Analysis
# - We use Sentiment lexicon resource for hindi called Hindi Sentiwordnet.
# - It has around 3000 prior-polarity subjective clues with POS tag, positive score, negative score and related terms(separated by comma)
# + colab={"base_uri": "https://localhost:8080/"} id="996HeBAnhq3k" outputId="6f72748a-9d73-48e3-d460-316aeaa73a35"
SUBJCLUE = [] # SUBJCLUE Data
with open('SUBJCLUE.txt') as f: # Formatting SUBJCLUE Data
for line in f: # iterate over the lines of the file
x = line.split() # split the line into a list of words
x[4] = x[4].split(',') # split the list into a list of words
SUBJCLUE.append(x) # append the list to the list of lists
# After this, the data would be in this form:
# ['POS tag', 'SYSNET ID(Hindi WN)', 'Positive score', 'Negative score', List of related words]
# printing the first 5 rows
for key in SUBJCLUE[:5]:
print(key[4])
# + [markdown] id="AADg4qY8qoyZ"
# ## Reading the data
#
# + [markdown] id="BmgETknXXHph"
# Note: The Dataset should be a csv file with Fields corresponding to Unique ID, Post, Labels Set
# + colab={"base_uri": "https://localhost:8080/"} id="gV-RT5LurrDr" outputId="423f29d0-a889-421a-b688-20a43b89a0ad"
import csv # importing csv module
# csv file name
filename = "Dataset/valid.csv" # change this file name to whatever you want
fields = [] # defining fields
rows = [] # defining rows
with open(filename, 'r') as csvfile: # opening csv file
# creating a csv reader object
csvreader = csv.reader(csvfile)
# extracting field names through first row
fields = next(csvreader)
# extracting each data row one by one
for row in csvreader:
rows.append(row)
# get total number of rows
print("Total no. of rows: %d"%(csvreader.line_num))
# printing the field names
print('Field names are:' + ', '.join(field for field in fields))
# printing first 5 rows
# Appending a score for each row
tot = 0
for row in rows:
row.append(tot)
# print(row)
# can be accessed using row[3]
for row in rows[:5]:
print(row)
# + [markdown] id="YUHPthTzs42T"
# ## Checking score
#
# Finding positive, negative and total scores for each sentence
#
#
# + id="dEGBezazs61_" colab={"base_uri": "https://localhost:8080/"} outputId="4c0ce45a-d393-4df0-edc7-8b1840422748"
count = 0 # initialize count
for key in SUBJCLUE: # for each word in SUBJCLUE
subjlist = key[4] # get the list of subjects
# subjlist = ['इच्छा', 'आत्मा', 'इतने']
for row in rows: # for each row in the csv file
if any([subjword in row[1] for subjword in subjlist]): # if any of the words in the list are in the row's text
count += 1 # increment count
pos = float(key[2]) # get the pos value
neg = float(key[3]) # get the neg value
tot = pos - neg # calculate the total
row[3] += tot # add the total to the row's total
# printing the number of occurences of sentiment words in dataset
print(count)
# + [markdown] id="imewqNVTN0cp"
# # Hate Lexicon Growing
#
# + id="n7c6WmGyF7oY" colab={"base_uri": "https://localhost:8080/"} outputId="2a085be0-5499-4361-eca5-c6bdd92ccb47"
# Installing required modules
# !pip install stanza
# !pip install setuptools
# !pip install subzero
# !pip install inltk
# + colab={"base_uri": "https://localhost:8080/", "height": 553, "referenced_widgets": ["726428bac2554499bea783078bb6593e", "0b9f83fe498f40f295d289003ab5ede2", "550df98871f34af690e0b2ed26ec82a1", "d7e477fe951543b590f2cf6eafaab77a", "cfc25ba9617647b4b9d5c75b066d54e6", "f05541c100ed484690cd8f1b58ad11a7", "b229c205c4de44a1885f836170339816", "f3873f326dd646618515defee0898d93"]} id="8HQa7z-8CQ32" outputId="5c20696d-8d85-40a5-e32e-ce9433745b70"
SYNSET = [] # SYNSET is a list of lists
with open('Synset.txt', encoding= 'unicode_escape') as f: # opening synset.txt file
for line in f: # iterating through SYNSET
x = line.split() # splitting lines
x[3] = x[3].split(':') # [3] is the synonyms
SYNSET.append(x) # append to SYNSET
import stanza # stanza is a library for natural language processing
stanza.download('hi', processors='tokenize,pos,lemma') # download the stanza library for Hindi NLP
import csv # csv is a library for reading and writing csv files
dataset = "" # dataset is a string
for row in rows: # iterating through rows
dataset+=row[1] # appending to dataset
verbs_content = [] # verbs_content is a list of lists
nlp = stanza.Pipeline('hi',processors='tokenize,pos,lemma') # nlp is a pipeline for processing text
# pos = open('hindi_pos.txt','w') # opening hindi_pos.txt in write mode
doc = nlp(dataset) # doc is a document object
for sentence in doc.sentences: # iterating through sentences
for word in sentence.words: # iterating through words
if word.upos == 'VERB': # if word is a verb
verbs_content.append(word.text) # append to verbs_content
strongly_negative_words = [] # strongly_negative_words is a list
weakly_negative_words = [] # weakly_negative_words is a list
for line in SUBJCLUE: # iterating through SUBJCLUE
totalscore = float(line[2]) - float(line[3]) # calculating total score
if(totalscore < -0.25): # if total score is less than -0.35
for word in line[4]: # iterating through words in line[4]
strongly_negative_words.append(word) # append to strongly_negative_words
elif totalscore < 0: # if total score is less than 0
for word in line[4]: # iterating through words in line[4]
weakly_negative_words.append(word) # append to weakly_negative_words
def Getsynset(word): # Getsynset is a function
syn = [] # syn is a list
flag=0 # flag is a variable
syn.append(word) # appending word to syn
for line in SYNSET: # iterating through SYNSET
if(line[1]=="03"): # if line[1] is equal to 03
for verb in line[3]: # iterating through verbs in line[3]
if(word == verb): # if word is equal to verb
flag = 1 # flag is set to 1
break # break
if(flag): # if flag is set to 1
syn = line[3] # syn is set to line[3]
break # break
return syn # return syn
s = {} # s is a dictionary
hlex = [] # hlex is a list
slist = ["लड़ना" , "मारना" , "लूटना" , "पीटना" , "कूटना" , "भेदभाव" ,"फोड़ना", "तोड़ना", "उखाड़ना" ] # slist is a list of verbs
for word in slist: # iterating through slist
hlex.append(word) # appending to hlex
for word in slist: # iterating through slist
s = Getsynset(word) # s is set to Getsynset
for verb1 in s: # iterating through s
if verb1 in verbs_content: # if verb1 is in verbs_content
hlex.append(verb1) # appending to hlex
# + id="l8xt1a-9EB76" colab={"base_uri": "https://localhost:8080/"} outputId="f6c4b2ab-88be-46b1-f252-fa8b30ed4807"
# open themenouns.txt in read
themed_nouns = open('themenouns.txt','r')
themenouns = [] # list of theme nouns
for line in themed_nouns: # read the file line by line
themenouns.append(line.rstrip('\n')) # append the theme nouns to the list
print(themenouns) # printing the theme nouns list
# + [markdown] id="PUjlJvTANlud"
# # Hate speech Detection Algorithm
# + colab={"base_uri": "https://localhost:8080/"} id="wk_UjwC5IrJI" outputId="89ba138d-76e7-4930-ed2a-2acde821cbe5"
print(strongly_negative_words) # printing the strongly negative words
print(weakly_negative_words) # printing the weakly negative words
print(hlex) # printing the hlex words
print(themenouns) # printing the themenouns
# + [markdown] id="KWR9UfCpYFIb"
# ## Calculating Scores without Subjective Analysis
# + [markdown] id="lEYAJ25UZBgo"
# ### Only Semantic feature set
# + id="eQIDn-pKNn4V" colab={"base_uri": "https://localhost:8080/"} outputId="6212281d-7fea-4f3d-8e89-1afb3c246648"
for row in rows: # Iterate over all rows
strongcount = 0 # Initialize strong count
hlexcount = 0 # Initialize hlex count
weakcount = 0 # Initialize weak count
themecount = 0 # Initialize theme count
if any([word in row[1] for word in strongly_negative_words]): # If any of the strongly negative words are in the tweet
strongcount += 1 # Increment strong count
# if any([word in row[1] for word in hlex]): # If any of the hlex words are in the tweet
# hlexcount += 1 # Increment hlex count
if any([word in row[1] for word in weakly_negative_words]): # If any of the weakly negative words are in the tweet
weakcount += 1 # Increment weak count
# if any([word in row[1] for word in themenouns]): # If any of the theme nouns are in the tweet
# themecount += 1 # Increment theme count
if strongcount >= 2: # If strong count is greater than or equal to 2
row.append("strongly hateful") # Append strongly hateful to the row
elif strongcount == 1: # Else if strong count is 1
if hlexcount >= 1 or themecount >= 1: # If hlex count is 1 or theme count is 1
row.append("strongly hateful") # Append strongly hateful to the row
else: # Else
row.append("weakly hateful") # Append weakly hateful to the row
elif strongcount == 0: # Else if strong count is 0
if themecount >= 1 and hlexcount >= 1: # If theme count is 1 and hlex count is 1
row.append("strongly hateful") # Append strongly hateful to the row
elif themecount >=1 and weakcount >= 1: # Else if theme count is 1 and weak count is 1
row.append("weakly hateful") # Append weakly hateful to the row
elif hlexcount == 1: # Else if hlex count is 1
row.append("weakly hateful") # Append weakly hateful to the row
else: # Else
row.append("No Hate") # Append No Hate to the row
# total rows = toal number of rows
total_rows = [row for row in rows]
# no_hate_rows = number of rows that are marked to have no hate
no_hate_rows = [row for row in rows if row[4] == "No Hate"]
# correct_no_hate_rows = number of rows that have no hate speech and are correctly marked
correct_no_hate_rows = [row for row in no_hate_rows if row[4] == "No Hate" and row[2] == "non-hostile"]
# weak_hate_rows = number of rows that are marked to have weak hate
weak_hate_rows = [row for row in rows if row[4] == "weakly hateful"]
# correct_weak_hate_rows = number of rows that have weak hate speech and are correctly marked
correct_weak_hate_rows = [row for row in weak_hate_rows if row[4] == "weakly hateful" and (row[2] == "fake" or row[2] == "defamation")]
# strong_hate_rows = number of rows that are marked to have strong hate
strong_hate_rows = [row for row in rows if row[4] == "strongly hateful"]
# correct_strong_hate_rows = number of rows that have strong hate speech and are correctly marked
correct_strong_hate_rows = [row for row in strong_hate_rows if row[4] == "strongly hateful" and row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation"]
# false negatives in the no hate list
false_neg_no_hate = [row for row in no_hate_rows if row[2] == "non-hostile" and row[4] != "No Hate"]
# false negatives in the weak hate list
false_neg_weak_hate = [row for row in weak_hate_rows if row[2] == "fake" or row[2] == "defamation" and row[4] != "weakly hateful"]
# false negatives in the strong hate list
false_neg_strong_hate = [row for row in strong_hate_rows if row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation" and row[4] != "strongly hateful"]
# calculating precision
precision = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(no_hate_rows)+len(strong_hate_rows)+len(weak_hate_rows))
# calculating recall
recall = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows)+len(false_neg_no_hate)+len(false_neg_strong_hate)+len(false_neg_weak_hate))
# calculating F1 score
f1 = 2*precision*recall/(precision+recall)
print("Total no. of rows: {}".format(len(total_rows))) # total no. of rows
print("No Hate: {}".format(len(no_hate_rows))) # no hate rows
print("Actual no hate: {}".format(len(correct_no_hate_rows))) # actual no hate rows
print("Weak Hate: {}".format(len(weak_hate_rows))) # weak hate rows
print("Actual weak hate: {}".format(len(correct_weak_hate_rows))) # actual weak hate rows
print("Strong Hate: {}".format(len(strong_hate_rows))) # strong hate rows
print("Actual strong hate: {}".format(len(correct_strong_hate_rows))) # actual strong hate rows
# print precision
print("Precision: {}".format(precision))
# print recall
print("Recall: {}".format(recall))
# print f1
print("F-score: {}".format(f1))
# + [markdown] id="Hqy7pvksZLyT"
# ### Semantic + Hate Lexicon
# + colab={"base_uri": "https://localhost:8080/"} id="cmOcHTyUmB4P" outputId="c5a55ff2-1194-4f71-b505-ed557f8da634"
for row in rows: # Iterate over all rows
strongcount = 0 # Initialize strong count
hlexcount = 0 # Initialize hlex count
weakcount = 0 # Initialize weak count
themecount = 0 # Initialize theme count
if any([word in row[1] for word in strongly_negative_words]): # If any of the strongly negative words are in the tweet
strongcount += 1 # Increment strong count
if any([word in row[1] for word in hlex]): # If any of the hlex words are in the tweet
hlexcount += 1 # Increment hlex count
if any([word in row[1] for word in weakly_negative_words]): # If any of the weakly negative words are in the tweet
weakcount += 1 # Increment weak count
# if any([word in row[1] for word in themenouns]): # If any of the theme nouns are in the tweet
# themecount += 1 # Increment theme count
if strongcount >= 2: # If strong count is greater than or equal to 2
row[4] = "strongly hateful" # Append strongly hateful to the row
elif strongcount == 1: # Else if strong count is 1
if hlexcount >= 1 or themecount >= 1: # If hlex count is 1 or theme count is 1
row[4] = "strongly hateful" # Append strongly hateful to the row
else: # Else
row[4] = "weakly hateful" # Append weakly hateful to the row
elif strongcount == 0: # Else if strong count is 0
if themecount >= 1 and hlexcount >= 1: # If theme count is 1 and hlex count is 1
row[4] = "strongly hateful" # Append strongly hateful to the row
elif themecount >=1 and weakcount >= 1: # Else if theme count is 1 and weak count is 1
row[4] = "weakly hateful" # Append weakly hateful to the row
elif hlexcount == 1: # Else if hlex count is 1
row[4] = "weakly hateful" # Append weakly hateful to the row
else: # Else
row[4] = "No Hate" # Append No Hate to the row
# total rows = toal number of rows
total_rows = [row for row in rows]
# no_hate_rows = number of rows that are marked to have no hate
no_hate_rows = [row for row in rows if row[4] == "No Hate"]
# correct_no_hate_rows = number of rows that have no hate speech and are correctly marked
correct_no_hate_rows = [row for row in no_hate_rows if row[4] == "No Hate" and row[2] == "non-hostile"]
# weak_hate_rows = number of rows that are marked to have weak hate
weak_hate_rows = [row for row in rows if row[4] == "weakly hateful"]
# correct_weak_hate_rows = number of rows that have weak hate speech and are correctly marked
correct_weak_hate_rows = [row for row in weak_hate_rows if row[4] == "weakly hateful" and (row[2] == "fake" or row[2] == "defamation")]
# strong_hate_rows = number of rows that are marked to have strong hate
strong_hate_rows = [row for row in rows if row[4] == "strongly hateful"]
# correct_strong_hate_rows = number of rows that have strong hate speech and are correctly marked
correct_strong_hate_rows = [row for row in strong_hate_rows if row[4] == "strongly hateful" and row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation"]
# false negatives in the no hate list
false_neg_no_hate = [row for row in no_hate_rows if row[2] == "non-hostile" and row[4] != "No Hate"]
# false negatives in the weak hate list
false_neg_weak_hate = [row for row in weak_hate_rows if row[2] == "fake" or row[2] == "defamation" and row[4] != "weakly hateful"]
# false negatives in the strong hate list
false_neg_strong_hate = [row for row in strong_hate_rows if row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation" and row[4] != "strongly hateful"]
# calculating precision
precision = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(no_hate_rows)+len(strong_hate_rows)+len(weak_hate_rows))
# calculating recall
recall = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows)+len(false_neg_no_hate)+len(false_neg_strong_hate)+len(false_neg_weak_hate))
# calculating F1 score
f1 = 2*precision*recall/(precision+recall)
print("Total no. of rows: {}".format(len(total_rows))) # total no. of rows
print("No Hate: {}".format(len(no_hate_rows))) # no hate rows
print("Actual no hate: {}".format(len(correct_no_hate_rows))) # actual no hate rows
print("Weak Hate: {}".format(len(weak_hate_rows))) # weak hate rows
print("Actual weak hate: {}".format(len(correct_weak_hate_rows))) # actual weak hate rows
print("Strong Hate: {}".format(len(strong_hate_rows))) # strong hate rows
print("Actual strong hate: {}".format(len(correct_strong_hate_rows))) # actual strong hate rows
# print precision
print("Precision: {}".format(precision))
# print recall
print("Recall: {}".format(recall))
# print f1
print("F-score: {}".format(f1))
# + [markdown] id="HbEg764EZSlI"
# ### Semantic + Hate Lexicon + Thematic Nouns
# + colab={"base_uri": "https://localhost:8080/"} id="vG1O1mUVmCL8" outputId="c4c29c0f-daa6-4ff7-fbb7-eaa8aa577bc1"
for row in rows: # Iterate over all rows
strongcount = 0 # Initialize strong count
hlexcount = 0 # Initialize hlex count
weakcount = 0 # Initialize weak count
themecount = 0 # Initialize theme count
if any([word in row[1] for word in strongly_negative_words]): # If any of the strongly negative words are in the tweet
strongcount += 1 # Increment strong count
if any([word in row[1] for word in hlex]): # If any of the hlex words are in the tweet
hlexcount += 1 # Increment hlex count
if any([word in row[1] for word in weakly_negative_words]): # If any of the weakly negative words are in the tweet
weakcount += 1 # Increment weak count
if any([word in row[1] for word in themenouns]): # If any of the theme nouns are in the tweet
themecount += 1 # Increment theme count
if strongcount >= 2: # If strong count is greater than or equal to 2
row[4] = "strongly hateful" # Append strongly hateful to the row
elif strongcount == 1: # Else if strong count is 1
if hlexcount >= 1 or themecount >= 1: # If hlex count is 1 or theme count is 1
row[4] = "strongly hateful" # Append strongly hateful to the row
else: # Else
row[4] = "weakly hateful" # Append weakly hateful to the row
elif strongcount == 0: # Else if strong count is 0
if themecount >= 1 and hlexcount >= 1: # If theme count is 1 and hlex count is 1
row[4] = "strongly hateful" # Append strongly hateful to the row
elif themecount >=1 and weakcount >= 1: # Else if theme count is 1 and weak count is 1
row[4] = "weakly hateful" # Append weakly hateful to the row
elif hlexcount == 1: # Else if hlex count is 1
row[4] = "weakly hateful" # Append weakly hateful to the row
else: # Else
row[4] = "No Hate" # Append No Hate to the row
# total rows = toal number of rows
total_rows = [row for row in rows]
# no_hate_rows = number of rows that are marked to have no hate
no_hate_rows = [row for row in rows if row[4] == "No Hate"]
# correct_no_hate_rows = number of rows that have no hate speech and are correctly marked
correct_no_hate_rows = [row for row in no_hate_rows if row[4] == "No Hate" and row[2] == "non-hostile"]
# weak_hate_rows = number of rows that are marked to have weak hate
weak_hate_rows = [row for row in rows if row[4] == "weakly hateful"]
# correct_weak_hate_rows = number of rows that have weak hate speech and are correctly marked
correct_weak_hate_rows = [row for row in weak_hate_rows if row[4] == "weakly hateful" and (row[2] == "fake" or row[2] == "defamation")]
# strong_hate_rows = number of rows that are marked to have strong hate
strong_hate_rows = [row for row in rows if row[4] == "strongly hateful"]
# correct_strong_hate_rows = number of rows that have strong hate speech and are correctly marked
correct_strong_hate_rows = [row for row in strong_hate_rows if row[4] == "strongly hateful" and row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation"]
# false negatives in the no hate list
false_neg_no_hate = [row for row in no_hate_rows if row[2] == "non-hostile" and row[4] != "No Hate"]
# false negatives in the weak hate list
false_neg_weak_hate = [row for row in weak_hate_rows if row[2] == "fake" or row[2] == "defamation" and row[4] != "weakly hateful"]
# false negatives in the strong hate list
false_neg_strong_hate = [row for row in strong_hate_rows if row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation" and row[4] != "strongly hateful"]
# calculating precision
precision = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(no_hate_rows)+len(strong_hate_rows)+len(weak_hate_rows))
# calculating recall
recall = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows)+len(false_neg_no_hate)+len(false_neg_strong_hate)+len(false_neg_weak_hate))
# calculating F1 score
f1 = 2*precision*recall/(precision+recall)
print("Total no. of rows: {}".format(len(total_rows))) # total no. of rows
print("No Hate: {}".format(len(no_hate_rows))) # no hate rows
print("Actual no hate: {}".format(len(correct_no_hate_rows))) # actual no hate rows
print("Weak Hate: {}".format(len(weak_hate_rows))) # weak hate rows
print("Actual weak hate: {}".format(len(correct_weak_hate_rows))) # actual weak hate rows
print("Strong Hate: {}".format(len(strong_hate_rows))) # strong hate rows
print("Actual strong hate: {}".format(len(correct_strong_hate_rows))) # actual strong hate rows
# print precision
print("Precision: {}".format(precision))
# print recall
print("Recall: {}".format(recall))
# print f1
print("F-score: {}".format(f1))
# + [markdown] id="-lWHjD3FYOoQ"
# ## Calculating Scores with Subjective Analysis
# + id="erIbFiW6BR-7" colab={"base_uri": "https://localhost:8080/"} outputId="4a8cf0cd-6c5d-4d1a-bbce-11e876efd7a6"
counter = 0 # Counter for the number of tweets
subj_rows = [] # List of all the subjective tweets
for row in rows: # Iterate through each row
if row[3] <= -0.5 or row[3] >= 1: # subjective sentence condition
subj_rows.append(row) # Append the row to the list
counter += 1
print("Number of Subjective Sentences: ")
print(counter) # Print the number of subjective tweets
# + [markdown] id="j-g57Y2EZa-o"
# ### Semantic feature set
# + id="H9HwRDrzLOwJ" colab={"base_uri": "https://localhost:8080/"} outputId="f90a1305-9602-4ac0-a3a7-65c2ac9e4770"
for row in rows: # Iterate over all rows
if row[3] <= -0.5 or row[3] >= 1: # If the score is over -0.5 or 0.5
strongcount = 0 # Set strongcount to 0
hlexcount = 0 # Set hlexcount to 0
weakcount = 0 # Set weakcount to 0
themecount = 0 # Set themecount to 0
if any([word in row[1] for word in strongly_negative_words]): # If any of the strongly negative words are in the tweet
strongcount += 1 # Add 1 to strongcount
# if any([word in row[1] for word in hlex]): # If any of the hlex words are in the tweet
# hlexcount += 1 # Add 1 to hlexcount
if any([word in row[1] for word in weakly_negative_words]): # If any of the weakly negative words are in the tweet
weakcount += 1 # Add 1 to weakcount
# if any([word in row[1] for word in themenouns]): # If any of the themenouns words are in the tweet
# themecount += 1 # Add 1 to themecount
if strongcount >= 2: # If strongcount is greater than or equal to 2
row.append("strongly hateful") # Append strongly hate to the row
elif strongcount == 1: # Else if strongcount is equal to 1
if hlexcount >= 1 or themecount >= 1: # If hlexcount is greater than or equal to 1 or themecount is greater than or equal to 1
row.append("strongly hateful") # Append strongly hate to the row
else: # Else
row.append("weakly hateful") # Append weakly hate to the row
elif strongcount == 0: # Else if strongcount is equal to 0
if themecount >= 1 and hlexcount >= 1: # If themecount is greater than or equal to 1 and hlexcount is greater than or equal to 1
row.append("strongly hateful") # Append strongly hate to the row
elif themecount >=1 and weakcount >= 1: # Else if themecount is greater than or equal to 1 and weakcount is greater than or equal to 1
row.append("weakly hateful") # Append weakly hate to the row
elif hlexcount == 1: # Else if hlexcount is equal to 1
row.append("weakly hateful") # Append weakly hate to the row
else: # Else
row.append("No Hate") # Append No Hate to the row
else: # Else
row.append("No Hate") # Append No Hate to the row
total_rows = [row for row in rows]
no_hate_rows = [row for row in rows if row[5] == "No Hate"]
correct_no_hate_rows = [row for row in no_hate_rows if row[5] == "No Hate" and row[2] == "non-hostile"]
weak_hate_rows = [row for row in rows if row[5] == "weakly hateful"]
correct_weak_hate_rows = [row for row in weak_hate_rows if row[5] == "weakly hateful" and (row[2] == "fake" or row[2] == "defamation")]
strong_hate_rows = [row for row in rows if row[5] == "strongly hateful"]
correct_strong_hate_rows = [row for row in strong_hate_rows if row[5] == "strongly hateful" and row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation"]
false_neg_no_hate = [row for row in no_hate_rows if row[2] == "non-hostile" and row[5] != "No Hate"]
false_neg_weak_hate = [row for row in weak_hate_rows if row[2] == "fake" or row[2] == "defamation" and row[5] != "weakly hateful"]
false_neg_strong_hate = [row for row in strong_hate_rows if row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation" and row[5] != "strongly hateful"]
precision = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(no_hate_rows)+len(strong_hate_rows)+len(weak_hate_rows))
recall = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows)+len(false_neg_no_hate)+len(false_neg_strong_hate)+len(false_neg_weak_hate))
f1 = 2*precision*recall/(precision+recall)
print("Total no. of rows: {}".format(len(total_rows)))
print("No Hate: {}".format(len(no_hate_rows)))
print("Actual no hate: {}".format(len(correct_no_hate_rows)))
print("Weak Hate: {}".format(len(weak_hate_rows)))
print("Actual weak hate: {}".format(len(correct_weak_hate_rows)))
print("Strong Hate: {}".format(len(strong_hate_rows)))
print("Actual strong hate: {}".format(len(correct_strong_hate_rows)))
print("Precision: {}".format(precision))
print("Recall: {}".format(recall))
print("F-score: {}".format(f1))
# + [markdown] id="yXxMHvFpZdaF"
# ### Semantic + Hate Lexicon
# + colab={"base_uri": "https://localhost:8080/"} id="8wBRGSWQrL2r" outputId="c455637c-c486-4a6a-c8e9-52e9cb684c33"
for row in rows: # Iterate over all rows
if row[3] <= -0.5 or row[3] >= 1: # If the score is over -0.5 or 0.5
strongcount = 0 # Set strongcount to 0
hlexcount = 0 # Set hlexcount to 0
weakcount = 0 # Set weakcount to 0
themecount = 0 # Set themecount to 0
if any([word in row[1] for word in strongly_negative_words]): # If any of the strongly negative words are in the tweet
strongcount += 1 # Add 1 to strongcount
if any([word in row[1] for word in hlex]): # If any of the hlex words are in the tweet
hlexcount += 1 # Add 1 to hlexcount
if any([word in row[1] for word in weakly_negative_words]): # If any of the weakly negative words are in the tweet
weakcount += 1 # Add 1 to weakcount
# if any([word in row[1] for word in themenouns]): # If any of the themenouns words are in the tweet
# themecount += 1 # Add 1 to themecount
if strongcount >= 2: # If strong count is greater than or equal to 2
row[5] = "strongly hateful" # Append strongly hateful to the row
elif strongcount == 1: # Else if strong count is 1
if hlexcount >= 1 or themecount >= 1: # If hlex count is 1 or theme count is 1
row[5] = "strongly hateful" # Append strongly hateful to the row
else: # Else
row[5] = "weakly hateful" # Append weakly hateful to the row
elif strongcount == 0: # Else if strong count is 0
if themecount >= 1 and hlexcount >= 1: # If theme count is 1 and hlex count is 1
row[5] = "strongly hateful" # Append strongly hateful to the row
elif themecount >=1 and weakcount >= 1: # Else if theme count is 1 and weak count is 1
row[5] = "weakly hateful" # Append weakly hateful to the row
elif hlexcount == 1: # Else if hlex count is 1
row[5] = "weakly hateful" # Append weakly hateful to the row
else: # Else
row[5] = "No Hate" # Append No Hate to the row
total_rows = [row for row in rows]
no_hate_rows = [row for row in rows if row[5] == "No Hate"]
correct_no_hate_rows = [row for row in no_hate_rows if row[5] == "No Hate" and row[2] == "non-hostile"]
weak_hate_rows = [row for row in rows if row[5] == "weakly hateful"]
correct_weak_hate_rows = [row for row in weak_hate_rows if row[5] == "weakly hateful" and (row[2] == "fake" or row[2] == "defamation")]
strong_hate_rows = [row for row in rows if row[5] == "strongly hateful"]
correct_strong_hate_rows = [row for row in strong_hate_rows if row[5] == "strongly hateful" and row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation"]
false_neg_no_hate = [row for row in no_hate_rows if row[2] == "non-hostile" and row[5] != "No Hate"]
false_neg_weak_hate = [row for row in weak_hate_rows if row[2] == "fake" or row[2] == "defamation" and row[5] != "weakly hateful"]
false_neg_strong_hate = [row for row in strong_hate_rows if row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation" and row[5] != "strongly hateful"]
precision = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(no_hate_rows)+len(strong_hate_rows)+len(weak_hate_rows))
recall = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows)+len(false_neg_no_hate)+len(false_neg_strong_hate)+len(false_neg_weak_hate))
f1 = 2*precision*recall/(precision+recall)
print("Total no. of rows: {}".format(len(total_rows)))
print("No Hate: {}".format(len(no_hate_rows)))
print("Actual no hate: {}".format(len(correct_no_hate_rows)))
print("Weak Hate: {}".format(len(weak_hate_rows)))
print("Actual weak hate: {}".format(len(correct_weak_hate_rows)))
print("Strong Hate: {}".format(len(strong_hate_rows)))
print("Actual strong hate: {}".format(len(correct_strong_hate_rows)))
print("Precision: {}".format(precision))
print("Recall: {}".format(recall))
print("F-score: {}".format(f1))
# + [markdown] id="wBg1o7MSZeox"
# ### Semantic + Hate Lexicon + Thematic Nouns
# + colab={"base_uri": "https://localhost:8080/"} id="5WzWpPKWrN21" outputId="0a2c4bc0-18e7-4f87-f814-4b49443a5a1d"
for row in rows: # Iterate over all rows
if row[3] <= -0.5 or row[3] >= 1: # If the score is over -0.5 or 0.5
strongcount = 0 # Set strongcount to 0
hlexcount = 0 # Set hlexcount to 0
weakcount = 0 # Set weakcount to 0
themecount = 0 # Set themecount to 0
if any([word in row[1] for word in strongly_negative_words]): # If any of the strongly negative words are in the tweet
strongcount += 1 # Add 1 to strongcount
if any([word in row[1] for word in hlex]): # If any of the hlex words are in the tweet
hlexcount += 1 # Add 1 to hlexcount
if any([word in row[1] for word in weakly_negative_words]): # If any of the weakly negative words are in the tweet
weakcount += 1 # Add 1 to weakcount
if any([word in row[1] for word in themenouns]): # If any of the themenouns words are in the tweet
themecount += 1 # Add 1 to themecount
if strongcount >= 2: # If strong count is greater than or equal to 2
row[5] = "strongly hateful" # Append strongly hateful to the row
elif strongcount == 1: # Else if strong count is 1
if hlexcount >= 1 or themecount >= 1: # If hlex count is 1 or theme count is 1
row[5] = "strongly hateful" # Append strongly hateful to the row
else: # Else
row[5] = "weakly hateful" # Append weakly hateful to the row
elif strongcount == 0: # Else if strong count is 0
if themecount >= 1 and hlexcount >= 1: # If theme count is 1 and hlex count is 1
row[5] = "strongly hateful" # Append strongly hateful to the row
elif themecount >=1 and weakcount >= 1: # Else if theme count is 1 and weak count is 1
row[5] = "weakly hateful" # Append weakly hateful to the row
elif hlexcount == 1: # Else if hlex count is 1
row[5] = "weakly hateful" # Append weakly hateful to the row
else: # Else
row[5] = "No Hate" # Append No Hate to the row
total_rows = [row for row in rows]
no_hate_rows = [row for row in rows if row[5] == "No Hate"]
correct_no_hate_rows = [row for row in no_hate_rows if row[5] == "No Hate" and row[2] == "non-hostile"]
weak_hate_rows = [row for row in rows if row[5] == "weakly hateful"]
correct_weak_hate_rows = [row for row in weak_hate_rows if row[5] == "weakly hateful" and (row[2] == "fake" or row[2] == "defamation")]
strong_hate_rows = [row for row in rows if row[5] == "strongly hateful"]
correct_strong_hate_rows = [row for row in strong_hate_rows if row[5] == "strongly hateful" and row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation"]
false_neg_no_hate = [row for row in no_hate_rows if row[2] == "non-hostile" and row[5] != "No Hate"]
false_neg_weak_hate = [row for row in weak_hate_rows if row[2] == "fake" or row[2] == "defamation" and row[5] != "weakly hateful"]
false_neg_strong_hate = [row for row in strong_hate_rows if row[2] != "non-hostile" and row[2] != "fake" and row[2] != "defamation" and row[5] != "strongly hateful"]
precision = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(no_hate_rows)+len(strong_hate_rows)+len(weak_hate_rows))
recall = (len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows))/(len(correct_no_hate_rows)+len(correct_strong_hate_rows)+len(correct_weak_hate_rows)+len(false_neg_no_hate)+len(false_neg_strong_hate)+len(false_neg_weak_hate))
f1 = 2*precision*recall/(precision+recall)
print("Total no. of rows: {}".format(len(total_rows)))
print("No Hate: {}".format(len(no_hate_rows)))
print("Actual no hate: {}".format(len(correct_no_hate_rows)))
print("Weak Hate: {}".format(len(weak_hate_rows)))
print("Actual weak hate: {}".format(len(correct_weak_hate_rows)))
print("Strong Hate: {}".format(len(strong_hate_rows)))
print("Actual strong hate: {}".format(len(correct_strong_hate_rows)))
print("Precision: {}".format(precision))
print("Recall: {}".format(recall))
print("F-score: {}".format(f1))
# + [markdown] id="mzwZW-sZYxQF"
# ## Exporting results into results.csv
# + id="SVeHBwzkOL5N"
import csv # Importing the csv module
fields = ['Unique ID', 'Post', 'Labels Set', 'Total Score', 'Hate Label' ,'Subjective Hate Label'] # Defining the fields of the csv file
with open("results.csv", 'w') as csvfile: # Opening the file
# creating a csv writer object
csvwriter = csv.writer(csvfile)
# writing the fields
csvwriter.writerow(fields)
# writing the data rows
csvwriter.writerows(rows)
|
Hate_Speech_Detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyspark.ml.regression import IsotonicRegression
from pyspark.ml.tuning import ParamGridBuilder
from pyspark.ml.tuning import CrossValidator
from pyspark.sql import SparkSession
from pyspark.ml import Pipeline
from pyspark.ml.evaluation import RegressionEvaluator
import numpy as np
import matplotlib.pyplot as plt
spark_session = SparkSession\
.builder\
.appName("Spark Regression")\
.getOrCreate()
dataset = spark_session\
.read\
.format("libsvm")\
.load("libsvm2.txt")
# +
# the technique of fitting a free-form line to a sequence of observations such that the fitted
#line is non-decreasing (or non-increasing)
#everywhere, and lies as close to the observations as possible.
ir = IsotonicRegression(featuresCol = 'features', labelCol = 'label',)
dataset.printSchema()
dataset.show()
# +
# Split the data into training and test sets (30% held out for testing)
(trainingData, testData) = dataset.randomSplit([0.7, 0.3])
print(trainingData.select("label").show(10))
train_model = ir.fit(trainingData)
# +
#cvModel = train_model.fit(trainingData)
# +
predictions = train_model.transform(testData)
# +
evaluator= RegressionEvaluator(labelCol="label", predictionCol="prediction", metricName="rmse")
rmse = evaluator.evaluate(predictions)
rfPred = train_model.transform(dataset)
rfResult = rfPred.toPandas()
plt.figure(figsize=(20,10))
plt.plot(rfResult.label, rfResult.prediction, 'b+')
plt.scatter(rfResult.label, rfResult.prediction, color="red")
plt.xlabel('Close price')
plt.ylabel('Prediction')
plt.suptitle("Model Performance RMSE: %f" % rmse)
plt.show()
print(rmse)
# -
rmse
train_model.transform(dataset).show()
rmse
|
notebooks/Classification/Isotonic Regression/IsotonicRegr-libsvm.ipynb
|