code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Nuclear theory and predictive power
# ## <NAME>
# #### Department of Phyics, Chalmers University of Technology, Sweden
# + [markdown] slideshow={"slide_type": "fragment"}
# #### 2018-08-31, Euroschool on Exotic Beams
# * This presentation is based on an [ipython](https://ipython.org/) notebook. The presentation itself is a [Reveal.js](https://revealjs.com/#/) HTML slideshow created with [nbconvert](https://github.com/jupyter/nbconvert).
# * All the material and accompanying source code is safely stored in a public [git](https://git-scm.com/) repository at [github](https://github.com/cforssen/SNFcocktail). Please feel free to download and try the examples yourself.
# ```
# [~]$ git clone https://github.com/cforssen/Euroschool2018_Forssen.git
# [~]$ cd Euroschool2018_Forssen
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Preliminaries: Python installation
#
# The installation of Python, together with the modules that allow scientific computations, is not very difficult.
#
# I recommend *Anaconda*, with the package manager *conda*, the works on Linux, Mac OS X, and even Windows.
#
# - [Anaconda](https://www.continuum.io/downloads) includes both Python and conda, plus a large number of preinstalled packages. However, this distribution requires quite some disk space. [Miniconda](http://conda.pydata.org/miniconda.html) is a good light-weight option . Read also the [conda online documentation](http://conda.pydata.org/docs/)
#
# Choose a Python-3 version and install the modules that are needed for these lectures:
#
# ```
# [~]$ conda install numpy scipy pandas matplotlib seaborn jupyter
# ```
#
# Even better, create a virtual environment (the modules are listed in the file 'environment.yml'):
# ```
# [~]$ conda env create
# [~]$ source activate euroschool-env
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# Let us start a [jupyter (ipython) notebook](http://jupyter.org/):
# ```
# [~]$ jupyter notebook Forssen_lecture1.ipynb
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# and import some important modules
# + slideshow={"slide_type": "-"}
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + slideshow={"slide_type": "skip"}
# Some care is needed when removing warnings. But for the final version of this notebook it should be safe.
import warnings
warnings.simplefilter("ignore", UserWarning)
warnings.simplefilter("ignore", FutureWarning)
# + slideshow={"slide_type": "skip"}
# Not really needed, but nicer plots
import seaborn as sns
sns.set()
sns.set_context("talk")
# + [markdown] slideshow={"slide_type": "slide"}
# # Learning from data
# + [markdown] slideshow={"slide_type": "-"}
# ## Inference
#
# > the act of passing from one proposition, statement or judgment considered as true to another whose truth is believed to follow from that of the former
#
# *(Webster)*
#
# Do premises $A, B, \ldots \to$ hypothesis, $H$?
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Deductive inference:
#
# > Premises allow definite determination of truth/falsity of H (syllogisms, symbolic logic, Boolean algebra)
#
# $B(H|A,B,...) = 0$ or $1$
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Inductive inference
#
# > Premises bear on truth/falsity of H, but don’t allow its definite determination (weak syllogisms, analogies)
#
# * $A, B, C, D$ share properties $x, y, z$;
# * $E$ has properties $x, y$
# * $\to$ $E$ probably has property $z$.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Statistical Inference
# * Quantify the strength of inductive inferences from facts, in the form of data ($D$), and other premises, e.g. models, to hypotheses about the phenomena producing the data.
#
# * Quantify via probabilities, or averages calculated using probabilities. Frequentists ($\mathcal{F}$) and Bayesians ($\mathcal{B}$) use probabilities very differently for this.
#
# * To the pioneers such as Bernoulli, Bayes and Laplace, a probability represented a *degree-of-belief* or plausability: how much they thought that something as true based on the evidence at hand. This is the Bayesian approach.
#
# * To the 19th century scholars, this seemed too vague and subjective. They redefined probability as the *long run relative frequency* with which an event occurred, given (infinitely) many repeated (experimental) trials.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The Bayesian recipe
# Assess hypotheses by calculating their probabilities $p(H_i | \ldots)$ conditional on known and/or presumed information using the rules of probability theory.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Probability Theory Axioms:
# + [markdown] slideshow={"slide_type": "-"}
# #### Product (AND) rule
#
# $$p(A, B | I) = p(A|I) p(B|A, I) = p(B|I)p(A|B,I)$$
#
# Should read $p(A,B|I)$ as the probability for propositions $A$ AND $B$ being true given that $I$ is true.
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Sum (OR) rule
#
# $$p(A + B | I) = p(A | I) + p(B | I) - p(A, B | I)$$
#
# $p(A+B|I)$ is the probability that proposition $A$ OR $B$ is true given that $I$ is true.
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Normalization
#
# $$p(A|I) + p(\bar{A}|I) = 1$$
#
# $\bar{A}$ denotes the proposition that $A$ is false.
# + [markdown] slideshow={"slide_type": "subslide"}
#
# ## Bayes' theorem
# Bayes' theorem follows directly from the product rule
#
# $$
# p(A|B,I) = \frac{p(B|A,I) p(A|I)}{p(B|I)}.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# The importance of this property to data analysis becomes apparent if we replace $A$ and $B$ by hypothesis($H$) and data($D$):
#
# $$
# p(H|D,I) = \frac{p(D|H,I) p(H|I)}{p(D|I)}.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# The power of Bayes’ theorem lies in the fact that it relates the quantity of interest, the probability that the hypothesis is true given the data, to the term we have a better chance of being able to assign, the probability that we would have observed the measured data if the hypothesis was true.
#
# The various terms in Bayes’ theorem have formal names.
# * The quantity on the far right, $p(H|I)$, is called the **prior** probability; it represents our state of knowledge (or ignorance) about the truth of the hypothesis before we have analysed the current data.
# * This is modified by the experimental measurements through $p(D|H,I)$, the **likelihood** function,
# * The denominator $p(D|I)$ is called the **evidence**. It does not depend on the hypothesis and can be regarded as a normalization constant.
# * Together, these yield the **posterior** probability, $p(H|D, I )$, representing our state of knowledge about the truth of the hypothesis in the light of the data.
#
# In a sense, Bayes’ theorem encapsulates the process of learning.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The friends of Bayes' theorem
# + [markdown] slideshow={"slide_type": "-"}
# #### Normalization
# $$\sum_i p(H_i|\ldots) = 1$$
#
# In the above, $H_i$ is an exclusive and exhaustive list of hypotheses. For example,let’s imagine that there are five candidates in a presidential election; then $H_1$ could be the proposition that the first candidate will win, and so on.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Marginalization
#
# $$\sum_i p(A,H_i|I) = \sum_i p(H_i|A,I) p(A|I) = p(A|I)$$
#
# The probability that $A$ is true, for example that unemployment will be lower in a year’s time (given all relevant information $I$, but irrespective of whoever becomes president) is then given by $\sum_i p(A,H_i|I)$.
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Marginalization (continuum limit)
#
# $$\int dx p(A,H(x)|I) = p(A|I)$$
#
# In the continuum limit of propositions we must understand $p(\ldots)$ as a pdf (probability density function).
# + [markdown] slideshow={"slide_type": "fragment"}
# Marginalization is a very powerful device in data analysis because it enables us to deal with nuisance parameters; that is, quantities which necessarily enter the analysis but are of no intrinsic interest. The unwanted background signal present in many experimental measurements are examples of nuisance parameters.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Inference With Parametric Models
# Inductive inference with parametric models is a very important tool in the natural sciences.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Consider $N$ different models $M_i$ ($i = 1, \ldots, N$), each with parameters $\boldsymbol{\alpha}_i$. Each of them implies a sampling distribution (conditional predictive distribution for possible data)
# $$
# p(D|\boldsymbol{\alpha}_i, M_i)
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# * The $\boldsymbol{\alpha}_i$ dependence when we fix attention on the actual, observed data ($D_\mathrm{obs}$) is the likelihood function
# $$
# \mathcal{L}_i (\boldsymbol{\alpha}_i) \equiv p(D_\mathrm{obs}|\boldsymbol{\alpha}_i, M_i)
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# * We may be uncertain about $i$ (**model uncertainty**),
# + [markdown] slideshow={"slide_type": "fragment"}
# * or uncertain about $\boldsymbol{\alpha}_i$ (**parameter uncertainty**).
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Parameter Estimation
#
# Premise = choice of model (pick specific $i$)
#
# $\Rightarrow$ What can we say about $\boldsymbol{\alpha}_i$?
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Model comparison:
#
# Premise = $\{M_i\}$
#
# $\Rightarrow$ What can we say about model $i$ compared to model $j$?
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Model adequacy:
#
# Premise = $M_1$
#
# $\Rightarrow$ Is $M_1$ adequate?
# + [markdown] slideshow={"slide_type": "skip"}
# #### Hybrid Uncertainty:
#
# Models share some common params: $\boldsymbol{\alpha}_i = \{ \boldsymbol{\varphi}, \boldsymbol{\eta}_i\}$
#
# $\Rightarrow$ What can we say about $\boldsymbol{\varphi}$? (Systematic error is an example)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Illustrative example #1: A Bayesian billiard game
# Adapted from the blog post [Frequentism and Bayesianism II: When Results Differ](http://jakevdp.github.io/blog/2014/06/06/frequentism-and-bayesianism-2-when-results-differ/)
#
# This example of nuisance parameters dates all the way back to the posthumous [1763 paper](http://www.stat.ucla.edu/history/essay.pdf) written by <NAME> himself. The particular version of this problem used here is borrowed from [Eddy 2004](ftp://selab.janelia.org/pub/publications/Eddy-ATG3/Eddy-ATG3-reprint.pdf).
#
# The setting is a rather contrived game in which Alice and Bob bet on the outcome of a process they can't directly observe:
#
# Alice and Bob enter a room. Behind a curtain there is a billiard table, which they cannot see, but their friend Carol can. Carol rolls a ball down the table, and marks where it lands. Once this mark is in place, Carol begins rolling new balls down the table. If the ball lands to the left of the mark, Alice gets a point; if it lands to the right of the mark, Bob gets a point. We can assume for the sake of example that Carol's rolls are unbiased: that is, the balls have an equal chance of ending up anywhere on the table. The first person to reach **six points** wins the game.
#
# Here the location of the mark (determined by the first roll) can be considered a nuisance parameter: it is unknown, and not of immediate interest, but it clearly must be accounted for when predicting the outcome of subsequent rolls. If the first roll settles far to the right, then subsequent rolls will favor Alice. If it settles far to the left, Bob will be favored instead.
# + [markdown] slideshow={"slide_type": "subslide"}
# Given this setup, here is the question we ask of ourselves:
#
# > In a particular game, after eight rolls, Alice has five points and Bob has three points. What is the probability that Bob will go on to win the game?
#
# Intuitively, you probably realize that because Alice received five of the eight points, the marker placement likely favors her. And given this, it's more likely that the next roll will go her way as well. And she has three opportunities to get a favorable roll before Bob can win; she seems to have clinched it. But, **quantitatively**, what is the probability that Bob will squeak-out a win?
# + [markdown] slideshow={"slide_type": "subslide"}
# ### A Naive Frequentist Approach
# Someone following a classical frequentist approach might reason as follows:
#
# To determine the result, we need an intermediate estimate of where the marker sits. We'll quantify this marker placement as a probability $p$ that any given roll lands in Alice's favor. Because five balls out of eight fell on Alice's side of the marker, we can quickly show that the maximum likelihood estimate of $p$ is given by:
#
# $$
# \hat{p} = 5/8
# $$
#
# (This result follows in a straightforward manner from the [binomial likelihood](http://en.wikipedia.org/wiki/Binomial_distribution)). Assuming this maximum likelihood probability, we can compute the probability that Bob will win, which is given by:
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# P(B) = (1 - \hat{p})^3
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# That is, he needs to win three rolls in a row. Thus, we find that the following estimate of the probability:
# -
p_hat = 5. / 8.
freq_prob = (1 - p_hat) ** 3
print("Naive Frequentist Probability of Bob Winning: %.2f" %freq_prob)
# + [markdown] slideshow={"slide_type": "fragment"}
# In other words, we'd give Bob the following odds of winning:
# -
print("Odds against Bob winning: %i to 1" %((1. - freq_prob) / freq_prob))
# So we've estimated using frequentist ideas that Alice will win about 17 times for each time Bob wins. Let's try a Bayesian approach next.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Bayesian approach
# -
# We can also approach this problem from a Bayesian standpoint. This is slightly more involved, and requires us to first define some notation.
#
# We'll consider the following random variables:
#
# - $B$ = <NAME>
# - $D$ = observed data, i.e. $D = (n_A, n_B) = (5, 3)$
# - $p$ = unknown probability that a ball lands on Alice's side during the current game
#
# We want to compute $P(B~|~D)$; that is, the probability that Bob wins given our observation that Alice currently has five points to Bob's three.
# + [markdown] slideshow={"slide_type": "subslide"}
# The general Bayesian method of treating nuisance parameters is *marginalization*, or integrating the joint probability over the entire range of the nuisance parameter. In this case, that means that we will first calculate the joint distribution
#
# $$
# P(B,p~|~D)
# $$
#
# and then marginalize over $p$ using the following identity:
#
# $$
# P(B~|~D) \equiv \int_{-\infty}^\infty P(B,p~|~D) {\mathrm d}p
# $$
#
# This identity follows from the definition of conditional probability, and the law of total probability: that is, it is a fundamental consequence of probability axioms and will always be true. Even a frequentist would recognize this; they would simply disagree with our interpretation of $P(p)$ as being a measure of uncertainty of our own knowledge.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Building our Bayesian Expression
# -
# To compute this result, we will manipulate the above expression for $P(B~|~D)$ until we can express it in terms of other quantities that we can compute.
# We'll start by applying the following definition of [conditional probability](http://en.wikipedia.org/wiki/Conditional_probability#Definition) to expand the term $P(B,p~|~D)$:
#
# $$
# P(B~|~D) = \int P(B~|~p, D) P(p~|~D) dp
# $$
# Next we use [Bayes' rule](http://en.wikipedia.org/wiki/Bayes%27_theorem) to rewrite $P(p~|~D)$:
#
# $$
# P(B~|~D) = \int P(B~|~p, D) \frac{P(D~|~p)P(p)}{P(D)} dp
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# Finally, using the same probability identity we started with, we can expand $P(D)$ in the denominator to find:
#
# $$
# P(B~|~D) = \frac{\int P(B~|~p,D) P(D~|~p) P(p) dp}{\int P(D~|~p)P(p) dp}
# $$
# -
# Now the desired probability is expressed in terms of three quantities that we can compute. Let's look at each of these in turn:
#
# - $P(B~|~p,D)$: This term is exactly the frequentist likelihood we used above. In words: given a marker placement $p$ and the fact that Alice has won 5 times and Bob 3 times, what is the probability that Bob will go on to six wins? Bob needs three wins in a row, i.e. $P(B~|~p,D) = (1 - p) ^ 3$.
# - $P(D~|~p)$: this is another easy-to-compute term. In words: given a probability $p$, what is the likelihood of exactly 5 positive outcomes out of eight trials? The answer comes from the well-known [Binomial distribution](http://en.wikipedia.org/wiki/Binomial_distribution): in this case $P(D~|~p) \propto p^5 (1-p)^3$
# - $P(p)$: this is our prior on the probability $p$. By the problem definition, we can assume that $p$ is evenly drawn between 0 and 1. That is, $P(p)$ is a uniform probability distribution in the range from 0 to 1.
# + [markdown] slideshow={"slide_type": "subslide"}
# Putting this all together, canceling some terms, and simplifying a bit, we find
# $$
# P(B~|~D) = \frac{\int_0^1 (1 - p)^6 p^5 dp}{\int_0^1 (1 - p)^3 p^5 dp}
# $$
# where both integrals are evaluated from 0 to 1.
# -
# These integrals might look a bit difficult, until we notice that they are special cases of the [Beta Function](http://en.wikipedia.org/wiki/Beta_function):
# $$
# \beta(n, m) = \int_0^1 (1 - p)^{n - 1} p^{m - 1}
# $$
# The Beta function can be further expressed in terms of gamma functions (i.e. factorials), but for simplicity we'll compute them directly using Scipy's beta function implementation:
# +
from scipy.special import beta
bayes_prob = beta(6 + 1, 5 + 1) / beta(3 + 1, 5 + 1)
print("P(B|D) = %.2f" %bayes_prob)
# + [markdown] slideshow={"slide_type": "subslide"}
# The associated odds are the following:
# -
print("Bayesian odds against Bob winning: %i to 1" %((1. - bayes_prob) / bayes_prob))
# So we see that the Bayesian result gives us 10 to 1 odds, which is quite different than the 17 to 1 odds found using the frequentist approach. So which one is correct?
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Brute-force (Monte Carlo) approach
# -
# For this type of well-defined and simple setup, it is actually relatively easy to use a Monte Carlo simulation to determine the correct answer. This is essentially a brute-force tabulation of possible outcomes: we generate a large number of random games, and simply count the fraction of relevant games that Bob goes on to win. The current problem is especially simple because so many of the random variables involved are uniformly distributed. We can use the ``numpy`` package to do this as follows:
# +
np.random.seed(0)
# play 100000 games with randomly-drawn p, between 0 and 1
p = np.random.random(100000)
# each game needs at most 11 rolls for one player to reach 6 wins
rolls = np.random.random((11, len(p)))
# count the cumulative wins for Alice and Bob at each roll
Alice_count = np.cumsum(rolls < p, 0)
Bob_count = np.cumsum(rolls >= p, 0)
# sanity check: total number of wins should equal number of rolls
total_wins = Alice_count + Bob_count
assert np.all(total_wins.T == np.arange(1, 12))
print("(Sanity check passed)")
# + slideshow={"slide_type": "subslide"}
# determine number of games which meet our criterion of (A wins, B wins)=(5, 3)
# this means Bob's win count at eight rolls must equal 3
good_games = Bob_count[7] == 3
print("Number of suitable games: {0}".format(good_games.sum()))
# truncate our results to consider only these games
Alice_count = Alice_count[:, good_games]
Bob_count = Bob_count[:, good_games]
# determine which of these games Bob won.
# to win, he must reach six wins after 11 rolls.
bob_won = np.sum(Bob_count[10] == 6)
print("Number of these games Bob won: {0}".format(bob_won.sum()))
# compute the probability
mc_prob = bob_won.sum() * 1. / good_games.sum()
print("Monte Carlo Probability of Bob winning: {0:.2f}".format(mc_prob))
print("MC Odds against Bob winning: {0:.0f} to 1".format((1. - mc_prob) / mc_prob))
# -
# The Monte Carlo approach gives 10-to-1 odds on Bob, which agrees with the Bayesian approach. Apparently, our naive frequentist approach above was flawed.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Discussion
# + [markdown] slideshow={"slide_type": "-"}
# This example shows several different approaches to dealing with the presence of a nuisance parameter *p*. The Monte Carlo simulation gives us a close brute-force estimate of the true probability (assuming the validity of our assumptions), which the Bayesian approach matches. The naïve frequentist approach, by utilizing a single maximum likelihood estimate of the nuisance parameter $p$, arrives at the wrong result.
#
# We should emphasize that **this does not imply frequentism itself is incorrect**. The incorrect result above is more a matter of the approach being "naive" than it being "frequentist". There certainly exist frequentist methods for handling this sort of nuisance parameter – for example, it is theoretically possible to apply a transformation and conditioning of the data to isolate the dependence on $p$ – but it's hard to find any approach to this particular problem that does not somehow take advantage of Bayesian-like marginalization over $p$.
# + [markdown] slideshow={"slide_type": "skip"}
# Another potential point of contention is that the question itself is posed in a way that is perhaps unfair to the classical, frequentist approach. A frequentist might instead hope to give the answer in terms of null tests or confidence intervals: that is, they might devise a procedure to construct limits which would provably bound the correct answer in $100\times(1 - p)$ percent of similar trials, for some value of $p$ – say, 0.05 (note this is a different $p$ than the $p$ we've been talking about above). This might be classically accurate, but it doesn't quite answer the question at hand. I'll leave discussion of the meaning of such confidence intervals for my follow-up post on the subject.
#
# There is one clear common point of these two potential frequentist responses: both require some degree of effort and/or special expertise; perhaps a suitable frequentist approach would be immediately obvious to someone with a PhD in statistics, but is most definitely *not* obvious to a statistical lay-person simply trying to answer the question at hand. In this sense, I think Bayesianism provides a better approach for this sort of problem: by simple algebraic manipulation of a few well-known axioms of probability within a Bayesian framework, we can straightforwardly arrive at the correct answer without need for other special expertise.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example #2: Linear regression with data outliers
# Adapted from the blog post [Frequentism and Bayesianism II: When Results Differ](http://jakevdp.github.io/blog/2014/06/06/frequentism-and-bayesianism-2-when-results-differ/)
# + [markdown] slideshow={"slide_type": "fragment"}
# One situation where the concept of nuisance parameters can be helpful is accounting for outliers in data. Consider the following dataset, relating the observed variables $x$ and $y$, and the error of $y$ stored in $e$.
# + slideshow={"slide_type": "subslide"}
x = np.array([ 0, 3, 9, 14, 15, 19, 20, 21, 30, 35,
40, 41, 42, 43, 54, 56, 67, 69, 72, 88])
y = np.array([33, 68, 34, 34, 37, 71, 37, 44, 48, 49,
53, 49, 50, 48, 56, 60, 61, 63, 44, 71])
e = np.array([ 3.6, 3.9, 2.6, 3.4, 3.8, 3.8, 2.2, 2.1, 2.3, 3.8,
2.2, 2.8, 3.9, 3.1, 3.4, 2.6, 3.4, 3.7, 2.0, 3.5])
# + [markdown] slideshow={"slide_type": "skip"}
# We'll visualize this data below:
# -
plt.errorbar(x, y, e, fmt='.k', ecolor='gray');
# + [markdown] slideshow={"slide_type": "fragment"}
# Our task is to find a line of best-fit to the data. It's clear upon visual inspection that there are some outliers among these points, but let's start with a simple non-robust maximum likelihood approach.
# + [markdown] slideshow={"slide_type": "subslide"}
# Like we saw in the previous post, the following simple maximum likelihood result can be considered to be either frequentist or Bayesian (with uniform priors): in this sort of simple problem, the approaches are essentially equivalent.
#
# We'll propose a simple linear model, which has a slope and an intercept encoded in a parameter vector $\theta$. The model is defined as follows:
# $$
# \hat{y}(x~|~\theta) = \theta_0 + \theta_1 x
# $$
# Given this model, we can compute a Gaussian likelihood for each point:
# $$
# p(x_i,y_i,e_i~|~\theta) \propto \exp\left[-\frac{1}{2e_i^2}\left(y_i - \hat{y}(x_i~|~\theta)\right)^2\right]
# $$
# The total likelihood is the product of all the individual likelihoods. Computing this and taking the log, we have:
# $$
# \log \mathcal{L}(D~|~\theta) = \mathrm{const} - \sum_i \frac{1}{2e_i^2}\left(y_i - \hat{y}(x_i~|~\theta)\right)^2
# $$
# This should all look pretty familiar if you read through the previous post. This final expression is the log-likelihood of the data given the model, which can be maximized to find the $\theta$ corresponding to the maximum-likelihood model. Equivalently, we can minimize the summation term, which is known as the *loss*:
# $$
# \mathrm{loss} = \sum_i \frac{1}{2e_i^2}\left(y_i - \hat{y}(x_i~|~\theta)\right)^2
# $$
# This loss expression is known as a *squared loss*; here we've simply shown that the squared loss can be derived from the Gaussian log likelihood.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Standard Likelihood Approach
# + [markdown] slideshow={"slide_type": "skip"}
# Following the logic of the previous post, we can maximize the likelihood (or, equivalently, minimize the loss) to find $\theta$ within a frequentist paradigm. For a flat prior in $\theta$, the maximum of the Bayesian posterior will yield the same result. (note that there are good arguments based on the principle of maximum entropy that a flat prior is not the best choice here; we'll ignore that detail for now, as it's a very small effect for this problem).
#
# For simplicity, we'll use scipy's ``optimize`` package to minimize the loss (in the case of squared loss, this computation can be done more efficiently using matrix methods, but we'll use numerical minimization for simplicity here)
# +
from scipy import optimize
def squared_loss(theta, x=x, y=y, e=e):
dy = y - theta[0] - theta[1] * x
return np.sum(0.5 * (dy / e) ** 2)
theta1 = optimize.fmin(squared_loss, [0, 0], disp=False)
xfit = np.linspace(0, 100)
plt.errorbar(x, y, e, fmt='.k', ecolor='gray')
plt.plot(xfit, theta1[0] + theta1[1] * xfit, '-k')
plt.title('Maximum Likelihood fit: Squared Loss');
# + [markdown] slideshow={"slide_type": "skip"}
# It's clear on examination that the outliers are exerting a disproportionate influence on the fit. This is due to the nature of the squared loss function. If you have a single outlier that is, say 10 standard deviations away from the fit, its contribution to the loss will out-weigh that of 25 points which are 2 standard deviations away!
#
# Clearly the squared loss is overly sensitive to outliers, and this is causing issues with our fit. One way to address this within the frequentist paradigm is to simply adjust the loss function to be more robust.
# + [markdown] slideshow={"slide_type": "skip"}
# ### Frequentist Correction for Outliers: Huber Loss
# + [markdown] slideshow={"slide_type": "skip"}
# The variety of possible loss functions is quite literally infinite, but one relatively well-motivated option is the [Huber loss](http://en.wikipedia.org/wiki/Huber_loss_function). The Huber loss defines a critical value at which the loss curve transitions from quadratic to linear. Let's create a plot which compares the Huber loss to the standard squared loss for several critical values $c$:
# + slideshow={"slide_type": "skip"}
t = np.linspace(-20, 20)
def huber_loss(t, c=3):
return ((abs(t) < c) * 0.5 * t ** 2
+ (abs(t) >= c) * -c * (0.5 * c - abs(t)))
plt.plot(t, 0.5 * t ** 2, label="squared loss", lw=2)
for c in (10, 5, 3):
plt.plot(t, huber_loss(t, c), label="Huber loss, c={0}".format(c), lw=2)
plt.ylabel('loss')
plt.xlabel('standard deviations')
plt.legend(loc='best');
# + [markdown] slideshow={"slide_type": "skip"}
# The Huber loss is equivalent to the squared loss for points which are well-fit by the model, but reduces the loss contribution of outliers. For example, a point 20 standard deviations from the fit has a squared loss of 200, but a c=3 Huber loss of just over 55. Let's see the result of the best-fit line using the Huber loss rather than the squared loss. We'll plot the squared loss result in light gray for comparison:
# + slideshow={"slide_type": "skip"}
def total_huber_loss(theta, x=x, y=y, e=e, c=3):
return huber_loss((y - theta[0] - theta[1] * x) / e, c).sum()
theta2 = optimize.fmin(total_huber_loss, [0, 0], disp=False)
plt.errorbar(x, y, e, fmt='.k', ecolor='gray')
plt.plot(xfit, theta1[0] + theta1[1] * xfit, color='lightgray')
plt.plot(xfit, theta2[0] + theta2[1] * xfit, color='black')
plt.title('Maximum Likelihood fit: Huber loss');
# + [markdown] slideshow={"slide_type": "skip"}
# By eye, this seems to have worked as desired: the fit is much closer to our intuition!
#
# However a Bayesian might point out that the motivation for this new loss function is a bit suspect: as we showed, the squared-loss can be straightforwardly derived from a Gaussian likelihood. The Huber loss seems a bit *ad hoc*: where does it come from? How should we decide what value of $c$ to use? Is there any good motivation for using a linear loss on outliers, or should we simply remove them instead? How might this choice affect our resulting model?
# + [markdown] slideshow={"slide_type": "subslide"}
# ### A Bayesian Approach to Outliers: Nuisance Parameters
# -
# The Bayesian approach to accounting for outliers generally involves *modifying the model* so that the outliers are accounted for. For this data, it is abundantly clear that a simple straight line is not a good fit to our data. So let's propose a more complicated model that has the flexibility to account for outliers. One option is to choose a mixture between a signal and a background:
#
# $$
# \begin{array}{ll}
# p(\{x_i\}, \{y_i\},\{e_i\}~|~\theta,\{g_i\},\sigma_b) = & \frac{g_i}{\sqrt{2\pi e_i^2}}\exp\left[\frac{-\left(\hat{y}(x_i~|~\theta) - y_i\right)^2}{2e_i^2}\right] \\
# &+ \frac{1 - g_i}{\sqrt{2\pi \sigma_B^2}}\exp\left[\frac{-\left(\hat{y}(x_i~|~\theta) - y_i\right)^2}{2\sigma_B^2}\right]
# \end{array}
# $$
#
# What we've done is expanded our model with some nuisance parameters: $\{g_i\}$ is a series of weights which range from 0 to 1 and encode for each point $i$ the degree to which it fits the model.
# + [markdown] slideshow={"slide_type": "skip"}
# $g_i=0$ indicates an outlier, in which case a Gaussian of width $\sigma_B$ is used in the computation of the likelihood. This $\sigma_B$ can also be a nuisance parameter, or its value can be set at a sufficiently high number, say 50.
# + [markdown] slideshow={"slide_type": "subslide"}
# Our model is much more complicated now: it has 22 free parameters rather than 2, but the majority of these can be considered nuisance parameters, which can be marginalized-out in the end, just as we marginalized (integrated) over $p$ in the Billiard example. Let's construct a function which implements this likelihood. We'll use the [emcee](http://dan.iel.fm/emcee/current/) package to explore the parameter space.
# + [markdown] slideshow={"slide_type": "subslide"}
# To actually compute this, we'll start by defining functions describing our prior, our likelihood function, and our posterior:
# + slideshow={"slide_type": "-"}
# theta will be an array of length 2 + N, where N is the number of points
# theta[0] is the intercept, theta[1] is the slope,
# and theta[2 + i] is the weight g_i
def log_prior(theta):
#g_i needs to be between 0 and 1
if (all(theta[2:] > 0) and all(theta[2:] < 1)):
return 0
else:
return -np.inf # recall log(0) = -inf
def log_likelihood(theta, x, y, e, sigma_B):
dy = y - theta[0] - theta[1] * x
g = np.clip(theta[2:], 0, 1) # g<0 or g>1 leads to NaNs in logarithm
logL1 = np.log(g) - 0.5 * np.log(2 * np.pi * e ** 2) - 0.5 * (dy / e) ** 2
logL2 = np.log(1 - g) - 0.5 * np.log(2 * np.pi * sigma_B ** 2) - 0.5 * (dy / sigma_B) ** 2
return np.sum(np.logaddexp(logL1, logL2))
def log_posterior(theta, x, y, e, sigma_B):
return log_prior(theta) + log_likelihood(theta, x, y, e, sigma_B)
# + [markdown] slideshow={"slide_type": "subslide"}
# Now we'll run the MCMC samples to explore the parameter space:
# +
# Note that this step will take a few minutes to run!
ndim = 2 + len(x) # number of parameters in the model
nwalkers = 50 # number of MCMC walkers
nburn = 10000 # "burn-in" period to let chains stabilize
nsteps = 15000 # number of MCMC steps to take
# set theta near the maximum likelihood, with
np.random.seed(0)
starting_guesses = np.zeros((nwalkers, ndim))
starting_guesses[:, :2] = np.random.normal(theta1, 1, (nwalkers, 2))
starting_guesses[:, 2:] = np.random.normal(0.5, 0.1, (nwalkers, ndim - 2))
import emcee
sampler = emcee.EnsembleSampler(nwalkers, ndim, log_posterior, args=[x, y, e, 50])
sampler.run_mcmc(starting_guesses, nsteps)
sample = sampler.chain # shape = (nwalkers, nsteps, ndim)
sample = sampler.chain[:, nburn:, :].reshape(-1, ndim)
# + [markdown] slideshow={"slide_type": "subslide"}
# Once we have these samples, we can exploit a very nice property of the Markov chains. Because their distribution models the posterior, we can integrate out (i.e. marginalize) over nuisance parameters simply by ignoring them!
#
# We can look at the (marginalized) distribution of slopes and intercepts by examining the first two columns of the sample:
# -
plt.plot(sample[:, 0], sample[:, 1], ',k', alpha=0.1)
plt.xlabel('intercept')
plt.ylabel('slope');
# + [markdown] slideshow={"slide_type": "skip"}
# We see a distribution of points near a slope of $\sim 0.4-0.5$, and an intercept of $\sim 29-34$. We'll plot this model over the data below, but first let's see what other information we can extract from this trace.
#
# One nice feature of analyzing MCMC samples is that the choice of nuisance parameters is completely symmetric: just as we can treat the $\{g_i\}$ as nuisance parameters, we can also treat the slope and intercept as nuisance parameters! Let's do this, and check the posterior for $g_1$ and $g_2$, the outlier flag for the first two points:
# + slideshow={"slide_type": "skip"}
plt.plot(sample[:, 2], sample[:, 3], ',k', alpha=0.1)
plt.xlabel('$g_1$')
plt.ylabel('$g_2$')
print("g1 mean: {0:.2f}".format(sample[:, 2].mean()))
print("g2 mean: {0:.2f}".format(sample[:, 3].mean()))
# + [markdown] slideshow={"slide_type": "skip"}
# There is not an extremely strong constraint on either of these, but we do see that $(g_1, g_2) = (1, 0)$ is slightly favored: the means of $g_1$ and $g_2$ are greater than and less than 0.5, respecively. If we choose a cutoff at $g=0.5$, our algorithm has identified $g_2$ as an outlier.
#
# Let's make use of all this information, and plot the marginalized best model over the original data. As a bonus, we'll draw red circles to indicate which points the model detects as outliers:
# + slideshow={"slide_type": "skip"}
theta3 = np.mean(sample[:, :2], 0)
g = np.mean(sample[:, 2:], 0)
outliers = (g < 0.5)
# + slideshow={"slide_type": "subslide"}
plt.errorbar(x, y, e, fmt='.k', ecolor='gray')
plt.plot(xfit, theta1[0] + theta1[1] * xfit, color='lightgray')
plt.plot(xfit, theta2[0] + theta2[1] * xfit, color='lightgray')
plt.plot(xfit, theta3[0] + theta3[1] * xfit, color='black')
plt.scatter(x[outliers], y[outliers],marker='o',s=40,edgecolors='r',linewidths=2,c='k')
plt.title('Maximum Likelihood fit: Bayesian Marginalization');
# -
# The result, shown by the dark line, matches our intuition! Furthermore, the points automatically identified as outliers are the ones we would identify by hand. For comparison, the gray lines show the two previous approaches: the simple maximum likelihood and the frequentist approach based on Huber loss.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Discussion
# + [markdown] slideshow={"slide_type": "-"}
# Here we've dived into linear regression in the presence of outliers. A typical Gaussian maximum likelihood approach fails to account for the outliers, but we were able to correct this in the frequentist paradigm by modifying the loss function, and in the Bayesian paradigm by adopting a mixture model with a large number of nuisance parameters.
#
# Both approaches have their advantages and disadvantages: the frequentist approach here is relatively straightforward and computationally efficient, but is based on the use of a loss function which is not particularly well-motivated. The Bayesian approach is well-founded and produces very nice results, but requires a rather subjective specification of a prior. It is also much more intensive in both coding time and computational time.
# + [markdown] slideshow={"slide_type": "skip"}
# For Bayes' billiard ball example, we showed that a naïve frequentist approach leads to the wrong answer, while a naïve Bayesian approach leads to the correct answer. This doesn't mean frequentism is wrong, but it does mean we must be very careful when applying it.
#
# For the linear regression example, we showed one possible approach from both frequentism and Bayesianism for accounting for outliers in our data. Using a robust frequentist cost function is relatively fast and painless, but is dubiously motivated and leads to results which are difficult to interpret. Using a Bayesian mixture model takes more effort and requires more intensive computation, but leads to a very nice result in which multiple questions can be answered at once: in this case, marginalizing one way to find the best-fit model, and marginalizing another way to identify outliers in the data.
# -
|
Forssen_lecture1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy
import numpy as np
pesos = np.array([50.,55.5,53.4,60.10,70.1,81.,65.3])
print('Média:',pesos.mean())
print('Máximo:',pesos.max())
print('Mínimo:',pesos.min())
pesos = np.linspace(50,60,num=20)
print(pesos)
valores = np.linspace(1,30,num=20)
print(valores)
print(pesos - valores)
print(pesos * valores)
pesos -= valores
print(pesos)
np.sum(pesos)
np.std(pesos)
print('variância populacional',np.var(pesos))
print('variância amostral',np.var(pesos,ddof=1))
print('desvio padrão populacional',np.std(pesos))
print('desvio padrão amostral',np.std(pesos,ddof=1))
np.random.rand()
vetor = np.random.rand(2,2)
print('Tipo:',type(vetor))
print(vetor)
print('Primeiro elemento, segunda linha:',vetor[1,0])
# Navegando pela primeira linha
print('Primeira linha:',[vetor[0,x] for x in range(vetor.shape[0])])
# Navegando pela segund coluna
print('Segunda coluna:',[vetor[x,1] for x in range(vetor.shape[1])])
print(vetor.shape)
print(vetor.shape[0])
print(vetor.shape[1])
np.random.rand()
np.random.seed(101)
print('Primeiro:',np.random.rand())
print('Segundo:',np.random.rand())
print('Terceiro:',np.random.rand())
seq = np.random.randn(20)
print(seq)
import scipy.stats as stats
import numpy as np
seq = np.random.randn(20)
stats.describe(seq)
print('Moda',stats.mode(seq))
print('Normal',stats.normaltest(seq))
nseq = np.random.randn(1000)
print('Normal',stats.normaltest(nseq))
import pandas as pd
serie = pd.Series([np.random.randn(50)])
print('Série',serie,'Tipo',type(serie))
df = pd.DataFrame({'Idade': np.random.randint(20,high=60,size=100),
'Altura': 1 + np.random.rand(100)})
df.head()
df.describe()
# +
# Atenção para variância e desvio padrão
print('variância',df.var()) # O ddof default é 1, então é o contrário do numpy!
print('variância pupulacional',df.var(ddof=0))
print('variância',df.std()) # O ddof default é 1, então é o contrário do numpy!
print('variância pupulacional',df.std(ddof=0))
# -
df.columns
df.values
df.index
print(df)
serie = df['Altura']
print(type(serie))
serie.head()
df.T
print(type(df))
df[(df.Idade > 35) & (df.Idade <= 40)]
modelo_df = pd.read_csv('mod-preditivo.csv')
modelo_df.head()
mod2_df = pd.read_csv('mod-preditivo-original.csv',decimal=',')
mod2_df.info()
mod2_df.head()
# # Exemplo de regressão linear com Scikit-learn
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
# %matplotlib inline
# ## Obter os dados
dados_df = pd.read_csv('pesos-alturas.csv',decimal=',')
dados_df.head()
# ## Analisar os dados
dados_df.describe()
dados_df.hist()
# ## Separar os dados em Treino e Teste
# Nesta análise, vamos tentar prever o peso de uma pessoa, dada a sua altura
X_train, X_test, y_train, y_test = train_test_split(dados_df[['Alturas']],dados_df[['Pesos']],
test_size=0.33)
# ## Criando e treinando o modelo de regressão
modelo = linear_model.LinearRegression()
modelo.fit(X_train, y_train)
# ## Avaliando o modelo
print(modelo.score(X_train,y_train))
# ## Executando o teste de previsões
predicoes = modelo.predict(X_test)
print(r2_score(y_test,predicoes))
plt.scatter(X_train, y_train, color='blue',s=10)
plt.plot(X_test, predicoes, color='red', linewidth=3)
|
book/capt6/Segundo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/24moliternoa/2021_22-Ratza-Intro-CS-Sem-2/blob/main/ANTOINETTA_MOLITERNO_Copy_of_python_basics_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2cOCCaF6Q4ys"
# ### Key Terms
#
# Complete the following questions to solidify you're understanding of the key concepts. Feel free to use the lecture notebook or any other resources to help you, but do not copy and paste answers from the internet instead use your own words to present your understanding.
#
# ----
# + [markdown] id="e_bDYmbjRHNc"
# What is a general use programming language?
# + [markdown] id="ROigjjFsRHKB"
# to create games and other technology in a language humans can learn (not 1s and 0s)
# + [markdown] id="X4j6a8y4RHGG"
# Who invented Python and when?
#
#
#
#
# + [markdown] id="B5WWUhgXRHDt"
# <NAME>, released February 20, 1991
# + [markdown] id="RHkgvf39RHBm"
# What is the difference between front-end and back-end?
# + [markdown] id="cl7LFD3pRG_H"
# front end is what you can interact with and see, the back end is the code
# + [markdown] id="oTkkpwhVRG80"
# What is a GUI?
# + [markdown] id="aoVSrA67RG6D"
# graphical user interface
# + [markdown] id="nHjQwFW1RG4C"
# What is an API?
# + [markdown] id="0A6LKX_5RG1x"
# application programming interface
# + [markdown] id="RWzJmW61RGzR"
# What is Open Source Software?
# + [markdown] id="J8Lcpl46RGwu"
# where you can see the source code and edit and use it
# + [markdown] id="gob_Rgp8RGqT"
# What is a development environment?
# + [markdown] id="BCPeV2UCRGZm"
# an environment where developers can experiment without breaking a live program
# + [markdown] id="lKduQMuWSYoC"
# What is meant by local and remote in the context of computers?
# + [markdown] id="GcJMiJjFSfO2"
# local is related or restricted to where you're at, remote is when the computer is not here and accessed thru a network
# + [markdown] id="cb6Coc48Sgev"
# What is an operating system?
# + [markdown] id="586kJGbvSlw5"
# a software that supports the computers basic functions
#
# + [markdown] id="7tRiStzGSltk"
# What is a kernel?
# + [markdown] id="jp5g1ZgWSlq6"
# the core of the computers operating system
# + [markdown] id="WdOAakQJSloH"
# What is a shell?
# + [markdown] id="Gpaic8JSSlle"
# a computer shell allows you to enter code commands with your keyboard
# + [markdown] id="hn5QICK0Slio"
# What shell do we have access to in Colab notebooks?
# + [markdown] id="vYRb0JJXSlfx"
# bash
# + [markdown] id="dmgmzh-ISldB"
# What is an interpreter?
# + [markdown] id="Y79bP4GLS3bI"
#
# + [markdown] id="A2q30j02S3G3"
# What is a value in programming?
# + [markdown] id="ilES-rT6S9vm"
# a computer program that directly executes instructions written in a programming or scripting language without translating it to computer language
# + [markdown] id="KI34llQnS--5"
# What is an expression in programming?
# + [markdown] id="VHXYGhIcTBw9"
# values and functions combined to make an output
# + [markdown] id="pmKFicLMTCcD"
# What is syntax?
# + [markdown] id="sqCxeE2TTKvy"
# a set of rules for or an analysis of the syntax of a coding language
# + [markdown] id="B8m5jqm6TLHE"
# What do we call the process of discovering and resolving errors?
# + [markdown] id="0TozR78oTRK4"
# debugging (?)
# + [markdown] id="OJR_RDQpTRyR"
# ### Code
# + [markdown] id="zPvODBfiTWCP"
# Let's revisit some of the things we practiced in the lecture. In the code cell below print your name to the console without first declaring it as a variable.
# + id="mZb-v_UwTO7B" colab={"base_uri": "https://localhost:8080/"} outputId="86c28ab5-8daa-4711-e43f-2e61489a2dc4"
print("<NAME>")
# + [markdown] id="sZPksnwpTnTD"
# Now declare your first name and last name as separate variables and combine them in the print statement.
# + id="oqmZRhYLTztw" colab={"base_uri": "https://localhost:8080/"} outputId="90014413-eace-43b9-937e-4eb66750df59"
last="moliterno"
first="antoinetta"
print(first+" "+last)
# + [markdown] id="cNe3K4WZT2_0"
# In the cell below run the "Zen of Python" easter egg.
# + id="FSkN7Q52UKyU" colab={"base_uri": "https://localhost:8080/"} outputId="51e207d2-f19f-4e8a-fa9e-da5cd8eed413"
import this
# + [markdown] id="2ADI5kQAUMLI"
# ### Explore
# + [markdown] id="vchHFmicUOid"
# This portion of the assignment contains things we didn't explicitly cover in the lecture, instead encouraging you to explore and experiment on your own to discover some of the different operators and expressions in Python. For each expression first describe what you expect to happen before running the code cell.
#
# Documentation for Python's numeric operators can be found [here](https://docs.python.org/3.10/library/stdtypes.html#numeric-types-int-float-complex)
# + [markdown] id="_lTiBbJMU28S"
# #### `5 + 2 * 2`
#
# What do you expect to happen?
# + [markdown] id="P6-diOTwU_ir"
# 2 will multiply by two and then be added to 5, outputting 9
# + id="ALTC2aYRUNRe" colab={"base_uri": "https://localhost:8080/"} outputId="a9aa85fa-bfdf-4fba-82ce-fe91416fc7bb"
5+2*2
# + [markdown] id="zSMDH8osVEEN"
# #### `2 / 3`
#
# What do you expect to happen?
# + [markdown] id="MJ_mZbouVI9_"
# two will be divided by three outputting the result
# + id="FYMUlyCEVHD1" colab={"base_uri": "https://localhost:8080/"} outputId="dc9d79eb-aa0f-42e3-b5d5-4f3321bacade"
2/3
# + id="vuTFIjbE3nWV"
# + id="cb9ghUNs3nTt"
# + [markdown] id="c8LQrbNQVIID"
# #### `2.5 * 10`
#
# What do you expect to happen?
# + [markdown] id="J1Ts7WLEVPU-"
# 2.5 is multiplied by 10 making the float 25.0
# + id="IuE7GclzVOpO" colab={"base_uri": "https://localhost:8080/"} outputId="184d7a86-bf65-499f-e7e8-1664a5bb35cb"
2.5*10
# + [markdown] id="YfKfY31nVSfy"
# #### `a`
#
# What do you expect to happen? name error
# + [markdown] id="Y_FQACtgVVL8"
#
# + id="pukzPvzXVUgM" colab={"base_uri": "https://localhost:8080/", "height": 165} outputId="c5734277-6a69-4b87-a319-6dfa691f503b"
a
# + [markdown] id="x_G2qoXLVhVj"
# #### `'a'`
#
#
# What do you expect to happen? the console will output a
# + [markdown] id="IjTY1xn_VoB4"
#
# + id="e0PEkzRHVjVo" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7cf535ac-b764-4849-d167-f7fca2601f76"
'a'
# + [markdown] id="2kCnvuRvVprG"
# #### `521 // 5`
#
# What do you expect to happen?
# + [markdown] id="QWOocovcV3i6"
# 521 is divided by 5 but it rounds up
# + id="n9QgKjHxV7oX" colab={"base_uri": "https://localhost:8080/"} outputId="eed3b7fa-753b-4176-e59c-566ad5410452"
521//5
|
ANTOINETTA_MOLITERNO_Copy_of_python_basics_assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# !pip install git+https://github.com/recohut/recohut.git@S346877
# +
# default_exp evaluation.sequences
# -
# # Sequence Evaluation
# > Implementation of Sequential evaluation modules.
#hide
from nbdev.showdoc import *
from fastcore.nb_imports import *
from fastcore.test import *
# +
#export
import pandas as pd
import numpy as np
from tqdm import tqdm
import matplotlib.pyplot as plt
from recohut.evaluation.metrics import precision, recall, mrr
# -
# Dummy ItemPop model for examples
# +
from recohut.utils.data import load_dataset
from recohut.utils.filters import filter_by_time, filter_top_k
from recohut.utils.splitting import split_last_session_out
from recohut.models.itempop import ItemPop_v2
df = load_dataset('music30_sample')
df.columns = ['session_id', 'user_id', 'item_id', 'ts', 'playtime']
df['ts'] = pd.to_datetime(df['ts'], unit='s')
# let's keep only the top-1k most popular items in the last month
df = filter_by_time(df, last_months=1, ts_col='ts')
df = filter_top_k(df, topk=1000, user_col='user_id', item_col='item_id', sess_col='session_id', ts_col='ts')
train, test = split_last_session_out(df, user_col='user_id', sess_col='session_id', seq_col='sequence', time_col='ts')
poprecommender = ItemPop_v2()
poprecommender.fit(train)
# -
#exporti
def get_test_sequences(test_data, given_k, seq_col='sequence'):
# we can run evaluation only over sequences longer than abs(LAST_K)
test_sequences = test_data.loc[test_data[seq_col].map(len) > abs(given_k), seq_col].values
return test_sequences
#exporti
def get_test_sequences_and_users(test_data, given_k, train_users, seq_col='sequence', user_col='user_id'):
# we can run evaluation only over sequences longer than abs(LAST_K)
mask = test_data[seq_col].map(len) > abs(given_k)
mask &= test_data[user_col].isin(train_users)
test_sequences = test_data.loc[mask, seq_col].values
test_users = test_data.loc[mask, user_col].values
return test_sequences, test_users
#exporti
def sequential_evaluation(recommender,
test_sequences,
evaluation_functions,
users=None,
given_k=1,
look_ahead=1,
top_n=10,
scroll=True,
step=1):
"""
Runs sequential evaluation of a recommender over a set of test sequences
:param recommender: the instance of the recommender to test
:param test_sequences: the set of test sequences
:param evaluation_functions: list of evaluation metric functions
:param users: (optional) the list of user ids associated to each test sequence. Required by personalized models like FPMC.
:param given_k: (optional) the initial size of each user profile, starting from the first interaction in the sequence.
If <0, start counting from the end of the sequence. It must be != 0.
:param look_ahead: (optional) number of subsequent interactions in the sequence to be considered as ground truth.
It can be any positive number or 'all' to extend the ground truth until the end of the sequence.
:param top_n: (optional) size of the recommendation list
:param scroll: (optional) whether to scroll the ground truth until the end of the sequence.
If True, expand the user profile and move the ground truth forward of `step` interactions. Recompute and evaluate recommendations every time.
If False, evaluate recommendations once per sequence without expanding the user profile.
:param step: (optional) number of interactions that will be added to the user profile at each step of the sequential evaluation.
:return: the list of the average values for each evaluation metric
"""
if given_k == 0:
raise ValueError('given_k must be != 0')
metrics = np.zeros(len(evaluation_functions))
with tqdm(total=len(test_sequences)) as pbar:
for i, test_seq in enumerate(test_sequences):
if users is not None:
user = users[i]
else:
user = None
if scroll:
metrics += sequence_sequential_evaluation(recommender,
test_seq,
evaluation_functions,
user,
given_k,
look_ahead,
top_n,
step)
else:
metrics += evaluate_sequence(recommender,
test_seq,
evaluation_functions,
user,
given_k,
look_ahead,
top_n)
pbar.update(1)
return metrics / len(test_sequences)
#exporti
def evaluate_sequence(recommender, seq, evaluation_functions, user, given_k, look_ahead, top_n):
"""
:param recommender: which recommender to use
:param seq: the user_profile/ context
:param given_k: last element used as ground truth. NB if <0 it is interpreted as first elements to keep
:param evaluation_functions: which function to use to evaluate the rec performance
:param look_ahead: number of elements in ground truth to consider. if look_ahead = 'all' then all the ground_truth sequence is considered
:return: performance of recommender
"""
# safety checks
if given_k < 0:
given_k = len(seq) + given_k
user_profile = seq[:given_k]
ground_truth = seq[given_k:]
# restrict ground truth to look_ahead
ground_truth = ground_truth[:look_ahead] if look_ahead != 'all' else ground_truth
ground_truth = list(map(lambda x: [x], ground_truth)) # list of list format
if not user_profile or not ground_truth:
# if any of the two missing all evaluation functions are 0
return np.zeros(len(evaluation_functions))
r = recommender.recommend(user_profile, user)[:top_n]
if not r:
# no recommendation found
return np.zeros(len(evaluation_functions))
reco_list = recommender.get_recommendation_list(r)
tmp_results = []
for f in evaluation_functions:
tmp_results.append(f(ground_truth, reco_list))
return np.array(tmp_results)
#exporti
def sequence_sequential_evaluation(recommender, seq, evaluation_functions, user, given_k, look_ahead, top_n, step):
if given_k < 0:
given_k = len(seq) + given_k
eval_res = 0.0
eval_cnt = 0
for gk in range(given_k, len(seq), step):
eval_res += evaluate_sequence(recommender, seq, evaluation_functions, user, gk, look_ahead, top_n)
eval_cnt += 1
return eval_res / eval_cnt
#export
def eval_seqreveal(train_data,
test_data,
model,
top_k=10,
):
"""
Evaluation with sequentially revealed user-profiles.
Here we evaluate the quality of the recommendations in a setting in which
user profiles are revealed sequentially. The user profile starts from the
first GIVEN_K events (or, alternatively, from the last -GIVEN_K events if GIVEN_K<0).
The recommendations are evaluated against the next LOOK_AHEAD events (the ground truth).
The user profile is next expanded to the next STEP events, the ground truth is
scrolled forward accordingly, and the evaluation continues until the sequence ends.
In typical next-item recommendation, we start with GIVEN_K=1, generate a set
of alternatives that will evaluated against the next event in the sequence
(LOOK_AHEAD=1), move forward of one step (STEP=1) and repeat until the
sequence ends.
You can set the LOOK_AHEAD='all' to see what happens if you had to recommend
a whole sequence instead of a set of a set of alternatives to a user.
Note:
Metrics are averaged over each sequence first, then averaged over all test sequences.
"""
GIVEN_K = 1
LOOK_AHEAD = 1
STEP = 1
metrics=['precision', 'recall', 'mrr']
test_sequences = get_test_sequences(test_data, GIVEN_K)
print('{} sequences available for evaluation'.format(len(test_sequences)))
results = sequential_evaluation(model,
test_sequences=test_sequences,
given_k=GIVEN_K,
look_ahead=LOOK_AHEAD,
evaluation_functions=[eval(metric) for metric in metrics],
top_n=top_k,
scroll=True, # scrolling averages metrics over all profile lengths
step=STEP)
results = [results, GIVEN_K, LOOK_AHEAD, STEP]
results = {
"Model": type(model).__name__,
"GIVEN_K": results[1],
"LOOK_AHEAD": results[2],
"STEP": results[3],
f"Precision@{top_k}": results[0][0],
f"Recall@{top_k}": results[0][1],
f"MRR@{top_k}": results[0][2],
}
return results
results = eval_seqreveal(train, test, poprecommender)
results
#export
def eval_staticprofile(train_data,
test_data,
model,
top_k=10,
):
"""
Evaluation with "static" user-profiles.
Here we evaluate the quality of the recommendations in a setting in which
user profiles are instead static. The user profile starts from the first
GIVEN_K events (or, alternatively, from the last -GIVEN_K events if GIVEN_K<0).
The recommendations are evaluated against the next LOOK_AHEAD events (the ground truth).
The user profile is not extended and the ground truth doesn't move forward.
This allows to obtain "snapshots" of the recommendation performance for
different user profile and ground truth lenghts. Also here you can set the
LOOK_AHEAD='all' to see what happens if you had to recommend a whole sequence
instead of a set of a set of alternatives to a user.
"""
GIVEN_K = 1
LOOK_AHEAD = 'all'
STEP=1
metrics=['precision', 'recall', 'mrr']
test_sequences = get_test_sequences(test_data, GIVEN_K)
print('{} sequences available for evaluation'.format(len(test_sequences)))
results = sequential_evaluation(model,
test_sequences=test_sequences,
given_k=GIVEN_K,
look_ahead=LOOK_AHEAD,
evaluation_functions=[eval(metric) for metric in metrics],
top_n=top_k,
scroll=False # notice that scrolling is disabled!
)
results = [results, GIVEN_K, LOOK_AHEAD, STEP]
results = {
"Model": type(model).__name__,
"GIVEN_K": results[1],
"LOOK_AHEAD": results[2],
"STEP": results[3],
f"Precision@{top_k}": results[0][0],
f"Recall@{top_k}": results[0][1],
f"MRR@{top_k}": results[0][2],
}
return results
results = eval_staticprofile(train, test, poprecommender)
results
#export
def eval_reclength(train_data,
test_data,
model,
):
"""
Evaluation for different recommendation list lengths. Analysis of next-item recommendation.
In next-item recommendation, we analyse the performance of the recommender system in the
scenario of next-item recommendation over the following dimensions:
- the length of the recommendation list, and
- the length of the user profile.
Note:
This evaluation is by no means exhaustive, as different the hyper-parameters
of the recommendation algorithm should be carefully tuned before drawing any
conclusions. Unfortunately, given the time constraints for this tutorial, we
had to leave hyper-parameter tuning out. A very useful reference about careful
evaluation of (session-based) recommenders can be found at:
"""
GIVEN_K = 1
LOOK_AHEAD = 1
STEP = 1
topk_list = [1, 5, 10, 20, 50, 100]
res_list = []
metrics=['precision', 'recall', 'mrr']
test_sequences = get_test_sequences(test_data, GIVEN_K)
print('{} sequences available for evaluation'.format(len(test_sequences)))
for topn in topk_list:
print('Evaluating recommendation lists with length: {}'.format(topn))
res_tmp = sequential_evaluation(model,
test_sequences=test_sequences,
given_k=GIVEN_K,
look_ahead=LOOK_AHEAD,
evaluation_functions=[eval(metric) for metric in metrics],
top_n=topn,
scroll=True, # here we average over all profile lengths
step=STEP)
mvalues = list(zip(metrics, res_tmp))
res_list.append((topn, mvalues))
# show separate plots per metric
# fig, axes = plt.subplots(nrows=1, ncols=len(metrics), figsize=(15,5))
res_list_t = list(zip(*res_list))
results = []
for midx, metric in enumerate(metrics):
mvalues = [res_list_t[1][j][midx][1] for j in range(len(res_list_t[1]))]
fig, ax = plt.subplots(figsize=(5,5))
ax.plot(topk_list, mvalues)
ax.set_title(metric)
ax.set_xticks(topk_list)
ax.set_xlabel('List length')
fig.tight_layout()
results.append(fig)
plt.close()
return results
results = eval_reclength(train, test, poprecommender)
results
display(results[0])
display(results[1])
display(results[2])
#export
def eval_profilelength(train_data,
test_data,
model,
top_k=20,
):
"""
Evaluation for different user profile lengths. Analysis of next-item recommendation.
In next-item recommendation, we analyse the performance of the recommender system in the
scenario of next-item recommendation over the following dimensions:
- the length of the recommendation list, and
- the length of the user profile.
Note:
This evaluation is by no means exhaustive, as different the hyper-parameters
of the recommendation algorithm should be carefully tuned before drawing any
conclusions. Unfortunately, given the time constraints for this tutorial, we
had to leave hyper-parameter tuning out. A very useful reference about careful
evaluation of (session-based) recommenders can be found at:
"""
given_k_list = [1, 2, 3, 4]
LOOK_AHEAD = 1
STEP = 1
topk_list = [1, 5, 10, 20, 50, 100]
res_list = []
metrics=['precision', 'recall', 'mrr']
test_sequences = get_test_sequences(test_data, max(given_k_list))
print('{} sequences available for evaluation'.format(len(test_sequences)))
for gk in given_k_list:
print('Evaluating profiles having length: {}'.format(gk))
res_tmp = sequential_evaluation(model,
test_sequences=test_sequences,
given_k=gk,
look_ahead=LOOK_AHEAD,
evaluation_functions=[eval(metric) for metric in metrics],
top_n=top_k,
scroll=False, # here we stop at each profile length
step=STEP)
mvalues = list(zip(metrics, res_tmp))
res_list.append((gk, mvalues))
# show separate plots per metric
# fig, axes = plt.subplots(nrows=1, ncols=len(metrics), figsize=(15,5))
res_list_t = list(zip(*res_list))
results = []
for midx, metric in enumerate(metrics):
mvalues = [res_list_t[1][j][midx][1] for j in range(len(res_list_t[1]))]
fig, ax = plt.subplots(figsize=(5,5))
ax.plot(given_k_list, mvalues)
ax.set_title(metric)
ax.set_xticks(given_k_list)
ax.set_xlabel('Profile length')
fig.tight_layout()
results.append(fig)
plt.close()
return results
results = eval_profilelength(train, test, poprecommender)
results
display(results[0])
display(results[1])
display(results[2])
#hide
# %reload_ext watermark
# %watermark -a "<NAME>." -m -iv -u -t -d -p recohut
|
nbs/evaluation/evaluation.sequences.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="boLhh_GqlCnp" colab_type="text"
# # Install packages
# + id="5GyZ-lp3McOZ" colab_type="code" colab={}
# !pip3 install pytorch-transformers
# + id="2YIqF__rRYs_" colab_type="code" colab={}
# !pip3 install seqeval
# + id="raajdaEMot0t" colab_type="code" colab={}
# !pip3 install spacy
# + id="iEAWOeb1Lr1V" colab_type="code" outputId="2a2ad2aa-e4aa-47d9-bbc8-b6b2c73b06df" colab={"base_uri": "https://localhost:8080/", "height": 34}
import pandas as pd
import numpy as np
import torch
from tqdm import tqdm, trange
from torch.optim import Adam
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from keras.preprocessing.sequence import pad_sequences
from sklearn.model_selection import train_test_split
from pytorch_transformers import BertTokenizer, BertConfig, BertForTokenClassification, AdamW
# + [markdown] id="RukHsVjKlMMG" colab_type="text"
# # Load data
# + id="RWUNbwXvL32c" colab_type="code" colab={}
url = "https://raw.githubusercontent.com/rpw199912j/MatBERT/master/mat_ner.csv"
data = pd.read_csv(url).fillna("O")
# + id="dS3i-H2uNjln" colab_type="code" outputId="2398c357-5026-480a-e84d-10c47563da5b" colab={"base_uri": "https://localhost:8080/", "height": 359}
data.head(10)
# + [markdown] id="jmBKJcwKlWIx" colab_type="text"
# # Pre-processing
# ## Get the sentece and labels
# + id="yh58-AQKPzcC" colab_type="code" colab={}
class SentenceGetter(object):
def __init__(self, data):
self.n_sent = 1
self.data = data
self.empty = False
self.grouped = self.data.groupby("Sentence #").apply(
lambda s: [(w, t) for w, t in zip(s["Word"].values.tolist(),
s["Tag"].values.tolist())])
self.sentences = [s for s in self.grouped]
def get_next(self):
try:
s = self.grouped["Sentence: {}".format(self.n_sent)]
self.n_sent += 1
return s
except:
return None
# + id="xiUiVPw_P_Eh" colab_type="code" colab={}
getter = SentenceGetter(data)
# + [markdown] id="B570HX6gluxj" colab_type="text"
# ### Take a look at the first sentence in the data
# + id="HCqVMLzBQBtt" colab_type="code" outputId="46780011-e1bc-4090-a738-5db570a9f56c" colab={"base_uri": "https://localhost:8080/", "height": 34}
sentences = [" ".join([s[0].lower() for s in sent]) for sent in getter.sentences]
print(sentences[0])
# + [markdown] id="tGbdOVPDlzoL" colab_type="text"
# ### Get the word-level label
# + id="9NL1XLB4QJXC" colab_type="code" outputId="f7ebabd8-a3fd-474a-8aaa-ed51a93fa0f0" colab={"base_uri": "https://localhost:8080/", "height": 34}
labels = [[s[1] for s in sent] for sent in getter.sentences]
print(labels[0])
# + [markdown] id="SVSpNhWumQXt" colab_type="text"
# Create a dictionary that maps each word label into a number
# + id="li23ZlGcQMoF" colab_type="code" outputId="b8da827c-5316-4df4-b7bc-20a57e6bfe55" colab={"base_uri": "https://localhost:8080/", "height": 54}
tags_vals = list(set(data["Tag"].values))
tag2idx = {t: i for i, t in enumerate(tags_vals)}
print(tag2idx)
# + [markdown] id="C2_Kd4UDl8IW" colab_type="text"
# # Apply BERT model
# ## Set constants and GPU processor
# + id="rxvd4GboQPAN" colab_type="code" colab={}
MAX_LEN = 64
BATCH_SIZE = 32
# + id="Yb1JcroVQSb2" colab_type="code" outputId="be6949c2-b388-43aa-91b1-a4c5d2b6c2cb" colab={"base_uri": "https://localhost:8080/", "height": 34}
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
n_gpu = torch.cuda.device_count()
print("The number of GPU: {}".format(n_gpu))
# + id="A3kaYA_cQUNj" colab_type="code" outputId="7fa064eb-bfe7-468d-96ac-02f8d5b1ec54" colab={"base_uri": "https://localhost:8080/", "height": 34}
torch.cuda.get_device_name(0)
# + [markdown] id="Pw2Xrk8ymkAV" colab_type="text"
# ## Get the pre-trained uncased word embeddings
# + id="iVdF711yQXiQ" colab_type="code" outputId="ac7eae08-a529-4023-ad25-27f077f348e3" colab={"base_uri": "https://localhost:8080/", "height": 34}
tokenizer = BertTokenizer.from_pretrained("bert-base-uncased", do_lower_case=True)
tokenizer.add_tokens(["stirrer", "teflon", "autoclave", "degc"])
# + [markdown] id="L3gTiqvcnSyh" colab_type="text"
# ## Tokenize all the sentences
# + id="_03dHZlHQba7" colab_type="code" outputId="483632a2-bced-42be-fcdc-bf418cd34d01" colab={"base_uri": "https://localhost:8080/", "height": 71}
print(sentences[1431])
tokenized_text = [tokenizer.tokenize(sent) for sent in sentences]
print(tokenized_text[1431])
# + [markdown] id="gTvSqQO1nYhT" colab_type="text"
# ### Pad all the tokenized sentences and labels to the same length
# + id="zcS_TwkkQdbA" colab_type="code" outputId="85bdb13e-1f8e-4de1-d7f6-1ae049cead30" colab={"base_uri": "https://localhost:8080/", "height": 170}
input_ids = pad_sequences([tokenizer.convert_tokens_to_ids(txt) for txt in tokenized_text],
maxlen=MAX_LEN, dtype="long", truncating="post", padding="post")
tags = pad_sequences([[tag2idx.get(l) for l in lab] for lab in labels],
maxlen=MAX_LEN, value=tag2idx["O"], dtype="long", truncating="post", padding="post")
print(input_ids[0])
print(tags[0])
# + [markdown] id="gxt1rB1-nfb4" colab_type="text"
# ### Create attention masks for the attention model
# + id="ScbJIMA_QoCB" colab_type="code" colab={}
attention_masks = [[float(i > 0) for i in ii] for ii in input_ids]
# + [markdown] id="07y1_vkqnrxd" colab_type="text"
# ## Split the data into 90% training set and 10% validation set
# + id="fp-GJTh5Qzc5" colab_type="code" colab={}
tr_inputs, val_inputs, tr_tags, val_tags = train_test_split(input_ids, tags,
random_state=2018, test_size=0.1)
tr_masks, val_masks, _, _ = train_test_split(attention_masks, input_ids,
random_state=2018, test_size=0.1)
# + [markdown] id="zqhRtq6Cn4nh" colab_type="text"
# ## Convert the data into Torch tensor format for later processing
# + id="1pPXMhDnQ2Si" colab_type="code" colab={}
tr_inputs = torch.tensor(tr_inputs)
val_inputs = torch.tensor(val_inputs)
tr_tags = torch.tensor(tr_tags)
val_tags = torch.tensor(val_tags)
tr_masks = torch.tensor(tr_masks)
val_masks = torch.tensor(val_masks)
# + [markdown] id="a2Q5jaqUoGkW" colab_type="text"
# ### Define the training and validation data in the DataLoader for NLP model
# + id="uVMsxh_8Q4XV" colab_type="code" colab={}
train_data = TensorDataset(tr_inputs, tr_masks, tr_tags)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=BATCH_SIZE)
valid_data = TensorDataset(val_inputs, val_masks, val_tags)
valid_sampler = SequentialSampler(valid_data)
valid_dataloader = DataLoader(valid_data, sampler=valid_sampler, batch_size=BATCH_SIZE)
# + [markdown] id="DARbsr9XoYYt" colab_type="text"
# # Finetuning the BERT model
# + id="F3k2YoHNQ7TJ" colab_type="code" outputId="49a77473-a7a6-4847-bcf2-0a0268a45161" colab={"base_uri": "https://localhost:8080/", "height": 34}
model = BertForTokenClassification.from_pretrained("bert-base-uncased", num_labels=len(tag2idx))
model.resize_token_embeddings(len(tokenizer))
# + [markdown] id="NWzmE7N6oc14" colab_type="text"
# ## Load the data into GPU
# + id="gYb4Pb0XRCZP" colab_type="code" outputId="5990de1b-4540-45ee-a16d-7ec33c639bca" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.cuda()
# + id="-t-u8x22RLEz" colab_type="code" colab={}
FULL_FINETUNING = True
if FULL_FINETUNING:
param_optimizer = list(model.named_parameters())
no_decay = ['bias', 'gamma', 'beta']
optimizer_grouped_parameters = [
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.01},
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.0}
]
else:
param_optimizer = list(model.classifier.named_parameters())
optimizer_grouped_parameters = [{"params": [p for n, p in param_optimizer]}]
# USE ADAM for gradient descent
optimizer = Adam(optimizer_grouped_parameters, lr=3e-5)
# + [markdown] id="38XQK6LSovt_" colab_type="text"
# ## Define metrics for finetuning
# + id="WLQ3PoTbRT2H" colab_type="code" colab={}
from seqeval.metrics import f1_score
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=2).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
# + id="-wN7VWZ6Rrnh" colab_type="code" colab={}
epochs = 5
max_grad_norm = 1.0
# + id="SFnTM9KlRhXK" colab_type="code" outputId="7754d2df-4572-4c72-9613-c09e7242146b" colab={"base_uri": "https://localhost:8080/", "height": 374}
for _ in trange(epochs, desc="Epoch"):
model.train()
tr_loss = 0
nb_tr_examples, nb_tr_steps = 0, 0
for step, batch in enumerate(train_dataloader):
# Use batch training to speed up training speed
batch = tuple(t.to(device) for t in batch)
b_input_ids, b_input_mask, b_labels = batch
# Forward-prop pass and loss computing
loss, _ = model(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask, labels=b_labels)
# Back-prop
loss.backward()
# track training loss
tr_loss += loss.item()
nb_tr_examples += b_input_ids.size(0)
nb_tr_steps += 1
# Gradient clipping to prevent gradient explosion
torch.nn.utils.clip_grad_norm_(parameters=model.parameters(),
max_norm=max_grad_norm)
# Update parameters
optimizer.step()
model.zero_grad()
print("Avg Training Loss Per Epoch: {}".format(tr_loss/nb_tr_steps))
# Validation
model.eval()
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
predictions, true_labels = [], []
for batch in valid_dataloader:
batch = tuple(t.to(device) for t in batch)
b_input_ids, b_input_mask, b_labels = batch
with torch.no_grad():
outputs = model(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask, labels=b_labels)
tmp_eval_loss, logits = outputs[:2]
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to("cpu").numpy()
predictions.extend([list(p) for p in np.argmax(logits, axis=2)])
true_labels.append(label_ids)
tmp_eval_accuracy = flat_accuracy(logits, label_ids)
eval_loss += tmp_eval_loss.mean().item()
eval_accuracy += tmp_eval_accuracy
nb_eval_examples += b_input_ids.size(0)
nb_eval_steps += 1
eval_loss = eval_loss / nb_eval_steps
print("Validation loss: {}".format(eval_loss))
print("Validation accuracy: {}".format(eval_accuracy/nb_eval_steps))
pred_tags = [tags_vals[p_i] for p in predictions for p_i in p]
valid_tags = [tags_vals[l_ii] for l in true_labels for l_i in l for l_ii in l_i]
print("F1 score: {}".format(f1_score(pred_tags, valid_tags)))
# + [markdown] id="6obVhAulo9hz" colab_type="text"
# ## Evaluate the model
# + id="N5jUJa2uRnMt" colab_type="code" outputId="5e5ed7b6-61fd-46f2-9823-cdd125fc344c" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.eval()
predictions = []
true_labels = []
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
for batch in valid_dataloader:
batch = tuple(t.to(device) for t in batch)
b_input_ids, b_input_mask, b_labels = batch
print(b_input_ids)
print(b_input_ids[1,:].tolist())
with torch.no_grad():
tmp_eval_loss, logits = model(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask, labels=b_labels)[:2]
logits = logits.detach().cpu().numpy()
predictions.extend([list(p) for p in np.argmax(logits, axis=2)])
label_ids = b_labels.to('cpu').numpy()
true_labels.append(label_ids)
tmp_eval_accuracy = flat_accuracy(logits, label_ids)
eval_loss += tmp_eval_loss.mean().item()
eval_accuracy += tmp_eval_accuracy
nb_eval_examples += b_input_ids.size(0)
nb_eval_steps += 1
pred_tags = [[tags_vals[p_i] for p_i in p] for p in predictions]
valid_tags = [[tags_vals[l_ii] for l_ii in l_i] for l in true_labels for l_i in l ]
print("Validation loss: {}".format(eval_loss/nb_eval_steps))
print("Validation Accuracy: {}".format(eval_accuracy/nb_eval_steps))
print("Validation F1-Score: {}".format(f1_score(pred_tags, valid_tags)))
# + id="4wpduSozgX5H" colab_type="code" outputId="70ed54f5-32bb-4803-a3ca-61b22321ebca" colab={"base_uri": "https://localhost:8080/", "height": 54}
print(pred_tags[1][:43])
# + id="ndFXj9hegn8w" colab_type="code" outputId="eb34c43c-ab1d-4cf0-cda6-5e889f6c5ad8" colab={"base_uri": "https://localhost:8080/", "height": 54}
print(valid_tags[1][:43])
# + id="DjghZIpPtA6X" colab_type="code" colab={}
ids = [2044, 2582, 18385, 2007, 1037, 8060, 30522, 2005, 1020, 1044, 2012, 2282, 4860, 1010, 1996, 21500, 8150, 2001, 4015, 2000, 1037, 30523, 1011, 7732, 18676, 1011, 3886, 30524, 1998, 9685, 2012, 8574, 30525, 2005, 1023, 2420, 2104, 21552, 1006, 3438, 11575, 1007, 1012, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
# + id="fpuC358Y3CgJ" colab_type="code" outputId="52c1b1be-5bbd-4ef9-cd57-4bd22852952b" colab={"base_uri": "https://localhost:8080/", "height": 54}
print(tokenizer.convert_ids_to_tokens(ids[:43]))
# + [markdown] id="CDxFFpKot6af" colab_type="text"
# # NER Visualization
# + id="oNDq6_FcdOU_" colab_type="code" colab={}
from spacy import displacy
# + id="ZzugIOkbuqxN" colab_type="code" outputId="1a165b75-8b19-4cc6-b451-2b2776f8d393" colab={"base_uri": "https://localhost:8080/", "height": 71}
tags_uppercase = [tag.upper() for tag in tags_vals]
print(tags_uppercase)
print(len(tags_uppercase))
# + id="v7RFqBiJpHn5" colab_type="code" colab={}
COLORS = {"AMOUNT-MISC": "linear-gradient(90deg, #aa9cfc, #fc9ce7)",
"NUMBER": "linear-gradient(90deg, orange, cyan)",
"AMOUNT-UNIT": "linear-gradient(90deg, red, orange)",
"PROPERTY-MISC": "linear-gradient(90deg, purple 40%, yellow)",
"MATERIAL": "#aa9cfc",
"NONRECIPE-MATERIAL": "red",
"TARGET": "#a4893d",
"META": "yellow",
"UNSPECIFIED-MATERIAL": "blue",
"APPARATUS-UNIT": "linear-gradient(90deg, #e66465, #9198e5)",
"MATERIAL-DESCRIPTOR": "#9198e5",
"SOLVENT": "#e66465",
"PROPERTY-TYPE": "brown",
"PRECURSOR": "pink",
"CONDITION-MISC": "#fc9ce7",
"APPARATUS-PROPERTY-TYPE": "orange",
"PROPERTY-UNIT": "linear-gradient(217deg, rgba(255,0,0,.8), rgba(255,0,0,0) 70.71%)",
"CONDITION-UNIT": "linear-gradient(217deg, rgba(400,0,0,.8), rgba(50,0,0,0) 70.71%)",
"APPARATUS-DESCRIPTOR": "#fea49f",
"SYNTHESIS-APPARATUS": "#bf4aa8",
"OPERATION": "#9e363a",
"CHARACTERIZATION-APPARATUS": "#4f5f76",
"BRAND": "#e4decd",
"CONDITION-TYPE": "#8bf0ba",
"GAS": "#ffdc6a",
"REFERENCE": "#feda6a"
}
# + id="hU2k3Xs3uAXc" colab_type="code" colab={}
def ner_visualize(sentence, tags, colors=COLORS):
sentence_concat = " ".join(sentence)
ents = []
start = 0
end = 0
for word, tag in zip(sentence, tags):
end = start + len(word) - 1
ents.append({"start": start, "end": end+1, "label": tag.upper()})
start = end + 2
test = [{"text": sentence_concat,
"ents": ents,
"title": None}]
options = {"ents": [tag.upper() for tag in set(tags) if tag not in ["O"]], "colors": colors}
displacy.render(test, style="ent", manual=True, options=options)
# + id="KWuVx2UtuFkC" colab_type="code" colab={}
ner_visualize(["Compound A", "was", "made", "by", "compound B", "by", "heating", "in", "the furncace", "at", "300", "degree", "celsius", "."],
["TARGET", "O", "O", "O", "MATERIAL", "O", "CONDITION-MISC","O", "APPARATUS-DESCRIPTOR", "O", "NUMBER", "O", "CONDITION-UNIT", "O"])
|
MatNER_v1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Determine the molecular gas mass of a z=1 star-forming galaxy
#
# We, in the present notebook, use known star formation rate from a galaxy and use it to estimate the molecular gas mass in the galaxy.
import numpy as np
import matplotlib.pyplot as plt
import astropy.constants as con
import astropy.units as u
from astropy.cosmology import Planck15 as cosmo
# ### Deriving Luminosity from star formation rate
#
# The formula to be used is from Kennicutt (1998) and stated below:
#
# $$SFR (in \ M_\odot year^{-1}) = 4.5 \times 10^{-44} \cdot L_{FIR} (in \ erg \ s^{-1})$$
#
# $$\Rightarrow L_{FIR} (in \ erg \ s^{-1}) = \frac{SFR (in \ M_\odot year^{-1}) }{4.5 \times 10^{-44}}$$
#
# This formula is used to compute the lumminosity density from the star formation rate.
# +
sfr = 30
kappa = 4.5e-44
lfir = sfr/kappa ## in erg/sec
print('The infrared luminosity (in cgs units) is: {:.2e}'.format(lfir))
# -
# Now we want to compute the CO luminosity from this IR luminosity. We can use the following equation from Carilli & Walter (2013),
#
# $$\log L_{IR} = 1.13 \cdot \log L'_{CO} + 0.53 $$
#
# $$\Rightarrow \log L'_{CO} = \frac{\log L_{IR} - 0.53}{1.13}$$
#
# Here, $\log L_{IR}$ in in units of $L_\odot$ and $\log L'_{CO}$ would be in $K \ km \ s^{-1} \ pc^2$.
# +
lsun = ((con.L_sun).value)*(1e7)
lfir_sun = lfir/lsun
logco = (np.log10(lfir_sun)-0.53)/1.13
lco_diff_units = 10**(logco)
print('The CO luminosity is (in K km s-1 pc2): {:.2e}'.format(lco_diff_units))
# -
# ### Choose appropriate frequency to observe with ALMA
#
# We now want to choose the appropriate frequency for observation. To do this we want to first compute the observed frequency of various CO lines at redshift $z \sim 1$.
# +
lines = np.array(['CO10', 'CO21', 'CO32', 'C043', 'CO54', 'CO65', 'CO75'])
rest_frame = np.array([115.271204, 230.537990, 345.795989, 461.040770, 576.267904, 691.473090, 806.651806])
redshift = 1.036
obs_freq = rest_frame/(1+redshift)
for i in range(len(lines)):
print(lines[i] + '\t' + str(obs_freq[i]))
# -
# ### CO (3-2) luminosity
#
# It is evident from the above observed frequencies that frequency corresponding to CO(3-2) is best for ALMA observations. Hence we want to use CO(3-2). And we want to convert our $L'_{CO}$ into $L'_{3-2}$, by multiplying a factor of 0.6 (from Carilli & Walter 2013).
lco_32 = lco_diff_units*0.6
# Adding the lensing effect,
lco_32_lens = 4.3*lco_32
# ### Calculating flux density
#
# Now we want to compute the flux density from the calculated CO luminosity using a formula from Solomon et al. (1997):
#
# $$ L'_{CO} = 3.25 \times 10^7 S_{CO}\Delta V \nu_{obs}^{-2} D_L^2 (1+z)^{-3}$$
#
# with $S_{CO}\Delta V$ in $Jy \ km \ s^{-1}$, $\nu_{obs}$ in $GHz$ and $D_L$ (which is luminosity distance -- we calculated this from the `Planck18` cosmology from `astropy.cosmology` module) in $Mpc$. Since, we want to measure the flux density (that is, $S_{CO} \Delta V$), we can adjust above equation as,
#
# $$S_{CO} \Delta V = \frac{L'_{CO} \cdot \nu_{obs}^2 (1+z)^3}{3.25 \times 10^7 \cdot D_L^2}$$
#
# Above formula is adapted below,
# +
dl = (cosmo.luminosity_distance(redshift)).value
freq1 = obs_freq[2]
aa = (lco_32_lens*freq1*freq1*((1+redshift)**3))
bb = dl*dl*(3.25e7)
flux_s = aa/bb
print('Flux density (in Jy km s-1) is: {:.2e}'.format(flux_s))
# -
# Since the above computed flux density also include the velosity dispersion, we want to divide it with an appropriate value of the velosity dispersion to get the flux density in the units os $Jy$, which is done below:
flux_density = flux_s/200
print('Flus density (in Jy): {:.2e}'.format(flux_density))
# ### Mass of H2
#
# We now want to compute the mass of moleculer $H_2$ from the CO luminosity we calculated earlier. We use the conversion factor of $\alpha_{CO} = 4 M_\odot (K km s^{-1} pc^2)^{-1}$ from Carilli & Walter (2013). Note that this conversion factor will give the mass of $H_2$ for CO (1-0) transition luminosity. Hence we want to use the same luminosity for conversion.
mass_h2 = lco_diff_units*4
print('Mass of H2 (in M_sun): {:.2e}'.format(mass_h2))
|
Radio_interferometry/p1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import Counter
import tensorflow as tf
import numpy as np
import re
# -
PARAMS = {
'min_freq': 5,
'window_size': 3,
'n_sampled': 100,
'embed_dim': 200,
'sample_words': ['six', 'gold', 'japan', 'college'],
'batch_size': 1000,
'n_epochs': 20,
}
# +
def preprocess_text(text):
text = text.replace('\n', ' ')
text = re.sub('\s+', ' ', text).strip().lower()
words = text.split()
word2freq = Counter(words)
words = [word for word in words if word2freq[word] > PARAMS['min_freq']]
print("Total words:", len(words))
_words = set(words)
PARAMS['word2idx'] = {c: i for i, c in enumerate(_words)}
PARAMS['idx2word'] = {i: c for i, c in enumerate(_words)}
PARAMS['vocab_size'] = len(PARAMS['idx2word'])
print('Vocabulary size:', PARAMS['vocab_size'])
indexed = [PARAMS['word2idx'][w] for w in words]
indexed = filter_high_freq(indexed)
print("Word preprocessing completed ...")
return indexed
def filter_high_freq(int_words, t=1e-5, threshold=0.8):
int_word_counts = Counter(int_words)
total_count = len(int_words)
word_freqs = {w: c / total_count for w, c in int_word_counts.items()}
prob_drop = {w: 1 - np.sqrt(t / word_freqs[w]) for w in int_word_counts}
train_words = [w for w in int_words if prob_drop[w] < threshold]
return train_words
def make_data(int_words):
x, y = [], []
for i in range(PARAMS['window_size'], len(int_words)-PARAMS['window_size']):
inputs = get_x(int_words, i)
x.append(inputs)
y.append(int_words[i])
return np.array(x), np.array(y)
def get_x(words, idx):
left = idx - PARAMS['window_size']
right = idx + PARAMS['window_size']
return words[left: idx] + words[idx+1: right+1]
# +
def model_fn(features, labels, mode, params):
W = tf.get_variable('softmax_W', [PARAMS['vocab_size'], PARAMS['embed_dim']])
b = tf.get_variable('softmax_b', [PARAMS['vocab_size']])
E = tf.get_variable('embedding', [PARAMS['vocab_size'], PARAMS['embed_dim']])
embedded = tf.nn.embedding_lookup(E, features) # forward activation
embedded = tf.reduce_mean(embedded, [1])
if mode == tf.estimator.ModeKeys.TRAIN:
loss_op = tf.reduce_mean(tf.nn.sampled_softmax_loss(
weights = W,
biases = b,
labels = labels,
inputs = embedded,
num_sampled = PARAMS['n_sampled'],
num_classes = PARAMS['vocab_size']))
train_op = tf.train.AdamOptimizer().minimize(
loss_op, global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode=mode, loss=loss_op, train_op=train_op)
if mode == tf.estimator.ModeKeys.PREDICT:
normalized_E = tf.nn.l2_normalize(E, -1)
sample_E = tf.nn.embedding_lookup(normalized_E, features)
similarity = tf.matmul(sample_E, normalized_E, transpose_b=True)
return tf.estimator.EstimatorSpec(mode, predictions=similarity)
def print_neighbours(similarity, top_k=5):
for i in range(len(PARAMS['sample_words'])):
neighbours = (-similarity[i]).argsort()[1:top_k+1]
log = 'Nearest to [%s]:' % PARAMS['sample_words'][i]
for k in range(top_k):
neighbour = PARAMS['idx2word'][neighbours[k]]
log = '%s %s,' % (log, neighbour)
print(log)
# +
with open('temp/ptb_train.txt') as f:
x_train, y_train = make_data(preprocess_text(f.read()))
estimator = tf.estimator.Estimator(model_fn)
estimator.train(tf.estimator.inputs.numpy_input_fn(
x_train, np.expand_dims(y_train, -1),
batch_size = PARAMS['batch_size'],
num_epochs = PARAMS['n_epochs'],
shuffle = True))
sim = np.array(list(estimator.predict(tf.estimator.inputs.numpy_input_fn(
x = np.array([PARAMS['word2idx'][w] for w in PARAMS['sample_words']]),
shuffle = False))))
print_neighbours(sim)
|
nlp-models/tensorflow/tf-estimator/word2vec_cbow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="pFTxgQSXQcJb"
# # Permutation Models for Bayesian Performance Analysis
#
# In this notebook we implement several probabilistic models on permutations to be used in a Bayesian inference framework for performance analysis. The notebook is divided in the following sections:
#
# * Preliminaries section installs and configures all the dependendencies required by the notebook.
# * Syntetic Data section contains a few tests of the Bayesian analysis carried out using syntetically generated permutation datasets.
# * Real Data section contains a Bayesian analysis carried out using real data comming from the comparison of several algorithms on several instances of the FlowShop Schedule Problem. See Ceberio et al. [1] for further details.
# + [markdown] id="B_anERYeKfI8"
# ## Preliminaries
# + [markdown] id="SxukXJ8gqUs2"
# ### Install pre-requisites
#
# * BallesMallows: an R package
# * Bayes Perm: our Pyhon package
# + [markdown] id="YmC9p4Xvqfp-"
# #### Install BayesMallows
# + id="p3mi0lMiFksw"
import rpy2
import rpy2.robjects.packages as rpackages
import rpy2.robjects.numpy2ri
from rpy2.robjects.vectors import StrVector
from rpy2.robjects.packages import importr
# + id="D3UvEi-SDJ4z"
# !sudo apt-get install libmpfr-dev -qq > /dev/null
# + id="jvV5qu9XHYhW"
rpy2.robjects.numpy2ri.activate()
# + id="54_htvZzAeMp"
# !chmod -R 777 /usr/local/lib/R/site-library
# + colab={"base_uri": "https://localhost:8080/"} id="hrfi3junrtvj" outputId="cabfb491-8bf7-4ef8-f20b-9ec70cc9b52c"
utils = rpackages.importr('utils')
utils.chooseCRANmirror(ind=1)
utils.install_packages('BayesMallows', verbose=False, quiet=True)
# + [markdown] id="_nQ44nPXqjMA"
# #### Install BayesPerm
# + colab={"base_uri": "https://localhost:8080/"} id="ycY7k-ryqmYD" outputId="61619786-7c2d-4891-83bc-7e5520172417"
# !pip install BayesPermus
# + [markdown] id="olgSO1JVqL8v"
# ### General imports
# + id="LJa-SxuZqQQJ"
import matplotlib.pyplot as plt
import itertools
import pandas as pd
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="abe14iNj3Xqe" outputId="831d2496-bc33-424d-e8cd-e30c38631f3b"
from BayesPermus.models.PlackettLuce import PlackettLuceDirichlet
from BayesPermus.models.PlackettLuce import PlackettLuceGamma
from BayesPermus.models.BradleyTerry import BradleyTerry
from BayesPermus.models.MallowsModel import MallowsModel
from BayesPermus.figure.plot import Plot
# + [markdown] id="ZDrNY-7Bz2iL"
# ## Case of study
# + [markdown] id="Qm3_SsZdz6jV"
# ### Preliminaries
# + [markdown] id="8ls9oVkvOfRp"
# #### Functions to calculate the marginal probabilities
# + id="2Pvw7deM2ov_"
def calculate_top_ranking_probs(orderings, num_samples=1000):
num_instances, num_algorithms = orderings.shape
# PL Dirichlet hyper-priors
dirichlet_alpha_pl = num_algorithms * [1]
# PL Gamma hyper-priors
gamma_alpha_pl = 0.5
gamma_beta_pl = 0.5
# BT Dirichlet hyper-priors
dirichlet_alpha_bt = num_algorithms * [1]
placettLuceDirichlet = PlackettLuceDirichlet(dirichlet_alpha_pl, num_samples=num_samples)
placettLuceGamma = PlackettLuceGamma(gamma_alpha_pl, gamma_beta_pl, num_samples=num_samples)
bradleyTerry = BradleyTerry(dirichlet_alpha_bt, num_samples=num_samples)
mallowsModel = MallowsModel(num_samples=num_samples)
pld = placettLuceDirichlet.calculate_top_ranking_probs(orderings)
plg = placettLuceGamma.calculate_top_ranking_probs(orderings)
bt = bradleyTerry.calculate_top_ranking_probs(orderings)
mm = mallowsModel.calculate_top_ranking_probs(orderings)
return pld, plg, bt, mm
# + id="Fs5clNHSeS3q"
def calculate_better_than_probs(orderings, num_samples=1000):
num_instances, num_algorithms = orderings.shape
# PL Dirichlet hyper-priors
dirichlet_alpha_pl = num_algorithms * [1]
# PL Gamma hyper-priors
gamma_alpha_pl = 0.5
gamma_beta_pl = 0.5
# BT Dirichlet hyper-priors
dirichlet_alpha_bt = num_algorithms * [1]
placettLuceDirichlet = PlackettLuceDirichlet(dirichlet_alpha_pl, num_samples=num_samples)
placettLuceGamma = PlackettLuceGamma(gamma_alpha_pl, gamma_beta_pl, num_samples=num_samples)
bradleyTerry = BradleyTerry(dirichlet_alpha_bt, num_samples=num_samples)
mallowsModel = MallowsModel(num_samples=num_samples)
pld = placettLuceDirichlet.calculate_better_than_probs(orderings)
plg = placettLuceGamma.calculate_better_than_probs(orderings)
bt = bradleyTerry.calculate_better_than_probs(orderings)
mm = mallowsModel.calculate_better_than_probs(orderings)
return pld, plg, bt, mm
# + id="hbRO8Xta2h8P"
def calculate_top_k_probs(orderings, num_samples=1000):
num_instances, num_algorithms = orderings.shape
# PL Dirichlet hyper-priors
dirichlet_alpha_pl = num_algorithms * [1]
# PL Gamma hyper-priors
gamma_alpha_pl = 0.5
gamma_beta_pl = 0.5
# BT Dirichlet hyper-priors
dirichlet_alpha_bt = num_algorithms * [1]
placettLuceDirichlet = PlackettLuceDirichlet(dirichlet_alpha_pl, num_samples=num_samples)
placettLuceGamma = PlackettLuceGamma(gamma_alpha_pl, gamma_beta_pl, num_samples=num_samples)
bradleyTerry = BradleyTerry(dirichlet_alpha_bt, num_samples=num_samples)
mallowsModel = MallowsModel(num_samples=num_samples)
pld = placettLuceDirichlet.calculate_top_k_probs(orderings)
plg = placettLuceGamma.calculate_top_k_probs(orderings)
bt = bradleyTerry.calculate_top_k_probs(orderings)
mm = mallowsModel.calculate_top_k_probs(orderings)
return pld, plg, bt, mm
# + [markdown] id="DrR1gqx_OlCL"
# #### Functions to plot the marginal probabilities
# + id="Z4RC6s3Q2-9o"
def plot_top_ranking_probs(fig_name, model_names, algorithm_names, probs, empirical):
plotter = Plot()
num_samples, num_algorithms = probs[0].shape
fig = plt.figure()
fig, axs = plt.subplots(1, num_algorithms, figsize=(4 * num_algorithms, 2), sharey=True)
plotter.plot_top_ranking_probs(model_names, algorithm_names, probs, empirical, axs)
fig.savefig(fig_name + ".pdf", bbox_inches='tight')
# + id="QgoURw4FebSd"
def plot_better_than_probs(fig_name, model_names, algorithm_names, probs, empirical):
plotter = Plot()
num_samples, num_algorithms, _ = probs[0].shape
fig = plt.figure()
fig, axs = plt.subplots(len(model_names), num_algorithms, figsize=(4 * num_algorithms, 2 * len(model_names)), sharey=True)
plotter.plot_better_than_probs(model_names, algorithm_names, probs, empirical, axs)
fig.savefig(fig_name + ".pdf", bbox_inches='tight')
# + id="Qb3kQb5j22iM"
def plot_top_k_probs(fig_name, model_names, algorithm_names, probs, empirical):
plotter = Plot()
num_samples, num_algorithms, _ = probs[0].shape
fig = plt.figure()
fig, axs = plt.subplots(len(model_names), num_algorithms, figsize=(4 * num_algorithms, 2 * len(model_names)), sharey=True)
plotter.plot_top_k_probs(model_names, algorithm_names, probs, empirical, axs)
fig.savefig(fig_name + ".pdf", bbox_inches='tight')
# + [markdown] id="TAQ4hqM_Oq5C"
# #### Functions to calculate the empirical marginals
# + id="6PQRAlm8mwqY"
def empirical_top_ranking_probs(orderings):
n, m =orderings.shape
probs = []
for i in range(m):
p_empirical = 0
for order in orderings:
if order[0] == i + 1:
p_empirical += 1
probs.append(p_empirical / n)
return probs
# + id="8_jPdti3sUIO"
def empirical_better_than(orderings):
def indexOf(arr, elem):
for i, val in enumerate(arr):
if val == elem:
return i
return -1
n, m =orderings.shape
probs = np.zeros((n, m))
for i in range(m):
for j in range(m):
if i != j:
p_empirical = 0
for order in orderings:
if indexOf(order, i + 1) < indexOf(order, j + 1):
p_empirical += 1
probs[i, j] = p_empirical / n
return probs
# + id="_mhRnfym3gYn"
def empirical_top_k(orderings):
def indexOf(arr, elem):
for i, val in enumerate(arr):
if val == elem:
return i
return -1
n, m =orderings.shape
probs = np.zeros((n, m))
for i in range(m):
for j in range(m):
p_empirical = 0
for order in orderings:
if indexOf(order, j + 1) <= i:
p_empirical += 1
probs[i, j] = p_empirical / n
return probs
# + id="_JW7Ta3kGXrg"
# + [markdown] id="OiK9h1jbOvVe"
# #### Functions to get insights on the empirical distributions
# + id="bqWLh3cLOMQu"
def calculate_hist(rankings):
permus = []
count = []
m = len(rankings)
def equals(pi, eta):
for x, y in zip(pi, eta):
if x != y:
return False
return len(pi) == len(eta)
def isin(pi, list):
for eta in list:
if equals(pi, eta):
return True
return False
def indexOf(arr, elem):
for i, val in enumerate(arr):
if val == elem:
return i
return -1
def kendall(pi, eta):
pairs = itertools.combinations(set(pi + eta), 2)
distance = 0
for x, y in pairs:
a = indexOf(pi, x) - indexOf(pi, y)
b = indexOf(eta, x) - indexOf(eta, y)
if a * b < 0:
distance += 1
return distance
for i, pi in enumerate(rankings):
c = 1
if not isin(pi, permus):
for j in range(i + 1, m):
if equals(pi, rankings[j]):
c += 1
permus.append(pi)
count.append(c)
mode_idx = np.argmax(count)
mode = permus[mode_idx]
n = 4
hist = []
for pi in rankings:
hist.append(kendall(list(pi), list(mode)))
return hist
# + id="k-DMfde1OcNh"
def plot_hist(max_distance, hist, ax, title):
hist = np.array(hist)
count = []
for d in range(0, max_distance + 1):
count.append((hist == d).sum())
ax.bar(range(max_distance + 1), count, color='gray')
ax.set_title(title)
ax.set_xlabel('Distance to mode')
# + [markdown] id="GKsQ28DPzUii"
# ### Syntetic Data Analyses
# + [markdown] id="pj8VgBhyPJ8o"
# #### Simple example
# + id="19JVvGdrGa0E"
def synthetic(num_instances, mean, std):
assert(len(mean) == len(std))
num_algorithms = len(mean)
scores = np.empty((num_instances, num_algorithms))
for i in range(num_instances):
for j in range(num_algorithms):
scores[i, j] = np.random.normal(mean[j], std[j])
return scores
# + id="-Jk8eA6PbjW8"
num_instances = 1000
mean = [2.0, 4.0, 6.0, 8.0]
std = [1.0, 1.0, 1.0, 1.0]
scores = synthetic(num_instances, mean, std)
orderings = np.argsort(scores, axis=1) + 1
rankings = np.argsort(orderings, axis=1) + 1
p_top_ranking = empirical_top_ranking_probs(orderings)
p_better_than = empirical_better_than(orderings)
p_top_k = empirical_top_k(orderings)
# + id="CNWGNR8nxYX0"
# + [markdown] id="1A0CskgFltHU"
# **Probability of an algorithm to be in the first ranking:**
# + colab={"base_uri": "https://localhost:8080/"} id="kB169eLxJJ_-" outputId="75ee02a1-bc60-42ae-8f73-40db009891f5"
probs = calculate_top_ranking_probs(orderings)
# + colab={"base_uri": "https://localhost:8080/", "height": 190} id="wysquKERwocF" outputId="dbaa7169-660d-4cb8-acd9-e61914968661"
plot_top_ranking_probs(fig_name='TopOneSynthetic', model_names=['PLD', 'PLG', 'BT', 'MM'], algorithm_names=['A1', 'A2', 'A3', 'A4'], probs=probs, empirical=p_top_ranking)
# + [markdown] id="OPESPszqlz-8"
# **Probability of an algorithm to be better than another:**
# + colab={"base_uri": "https://localhost:8080/"} id="ow3p7tAvky6v" outputId="fd0dc37f-92c5-493e-9438-b3d422670549"
probs = calculate_better_than_probs(orderings)
# + colab={"base_uri": "https://localhost:8080/", "height": 516} id="7nPC1dhjk0x0" outputId="156522c6-55f2-4075-ffd9-ac7700ee5ce1"
plot_better_than_probs(fig_name='BetterThanSyntetic', model_names=['PLD', 'PLG', 'BT', 'MM'], algorithm_names=['A1', 'A2', 'A3', 'A4'], probs=probs, empirical=p_better_than)
# + [markdown] id="Mz-Ki91e2sHq"
# **Probability of an algorithm to be in the top-k ranking:**
# + colab={"base_uri": "https://localhost:8080/"} id="LHJJ1cAB2dXY" outputId="0dbbba9d-9cf3-46ca-9c94-4e53a609ee7d"
probs = calculate_top_k_probs(orderings)
# + id="G3ujOdE454gc"
# + colab={"base_uri": "https://localhost:8080/", "height": 516} id="rAdI78622y9F" outputId="72d4a838-7da6-4204-acd3-3b4c95bf501a"
plot_top_k_probs(fig_name='TopKSyntetic', model_names=['PLD', 'PLG', 'BT', 'MM'], algorithm_names=['A1', 'A2', 'A3', 'A4'], probs=probs, empirical=p_top_k)
# + [markdown] id="Luc8uS3HnjEt"
# #### Effect of multimodal distributions
# + [markdown] id="g-PEShpxQ0ck"
# **Generate empirical distributions using different standard deviations:**
# + id="tHHgOpvEsIqF"
stds = [2.0, 4.0, 12.0]
Lscores = []
Lorderings = []
Lrankings = []
for i, std in enumerate(stds):
num_instances = 1000
mean = [2.0, 4.0, 6.0, 8.0]
std = [std, 1.0, 1.0, std]
scores = synthetic(num_instances, mean, std)
orderings = np.argsort(scores, axis=1) + 1
rankings = np.argsort(orderings, axis=1) + 1
Lscores.append(scores)
Lorderings.append(orderings)
Lrankings.append(rankings)
# + [markdown] id="_FPtXnIFQr61"
# **Histograms of the empirical distributions:**
# + colab={"base_uri": "https://localhost:8080/", "height": 187} id="zWEjHk8msKWN" outputId="f0a3ce41-050f-4039-841b-cf65c57b5a41"
fig, axs = plt.subplots(1, len(stds), figsize=(3 * len(stds), 2), sharey=True)
for config, (rankings, ax) in enumerate(zip(Lrankings, axs)):
hist = calculate_hist(rankings)
plot_hist(6, hist, ax, title='Configuration ' + str(config + 1))
# + id="s-hYMiqRUjhD"
fig.savefig('hist.pdf', bbox_inches='tight')
# + [markdown] id="izbYWAuRTWHh"
# Probability of an algorithm to be in the first ranking:
# + colab={"base_uri": "https://localhost:8080/"} id="NIz4ZA0tTQsq" outputId="b4c90081-2866-4169-c275-371eb07a2a79"
probs_per_config = []
for config, orderings in enumerate(Lorderings):
probs = calculate_top_ranking_probs(orderings)
probs_per_config.append(probs)
# + colab={"base_uri": "https://localhost:8080/", "height": 537} id="5T9txF2wWzXL" outputId="aa1bc568-cbc8-4b6b-819f-418f111e0c26"
for orderings, probs in zip(Lorderings, probs_per_config):
p_top_ranking = empirical_top_ranking_probs(orderings)
plot_top_ranking_probs("TopOneUnimodalSyntetic", model_names=['PLD', 'PLG', 'BT', 'MM'], algorithm_names=['A1', 'A2', 'A3', 'A4'], probs=probs, empirical=p_top_ranking)
# + [markdown] id="OCk8B92gQJed"
# ### EDA FSP Data
# + [markdown] id="kGqD8qycJDyM"
# #### Preliminaries
#
# + colab={"base_uri": "https://localhost:8080/"} id="kcvUxGyOl7fL" outputId="5ae02222-141d-4738-9f95-8d7fd4113a1b"
# !unzip -q FSPData.zip
# + id="HLA02Mn82Gbn"
def fix_index(df):
fixed_index = []
problems = []
for problem, rep in df.index:
if type(problem) == str:
problems.append(problem)
prev = problem
fixed_index.append((prev, rep))
fixed_index = pd.MultiIndex.from_tuples(fixed_index)
return fixed_index, problems
# + id="1BzPQr3M8UuV"
def ranks_from_score(score):
# Set of linear extensions of the original ranking. The linear extensions are
# obtained when ties are resolved in all possible ways.
permus = []
weights = []
n = len(score)
rank = np.argsort(score)
# List that contains several lists that represent element's index that are
# repeated in the original score.
#
# For example, if there is a single list with two elements, e.g. [[3, 6]] it
# means that `score[3] == score[6].
ties_set = []
excluded = []
# Loop through all elements in score.
for i in range(n):
if i not in excluded:
repeated = [i]
# Check if there are any ties in the rest of the score list.
for j in range(i + 1, n):
# If there is a tie, then add the entry to the repeated list.
if score[i] == score[j]:
repeated.append(j)
excluded += repeated
# If there is any tie, then, add it to the tie set.
if len(repeated) > 1:
ties_set.append(repeated)
# List that contains several lists that represent the elements index that are
# repeated in the original score and all their permutations.
#
# For example, if there is a single list with two elements, it means that there
# is a single repeated element that appears three times in the original score.
#
# If there are two lists with three elements each, it means that there are two
# repeated elements that appear three times each in the original score.
#
# The values within each list inside this list represent the rankings of such
# repeated entries.
extensions = []
for i, repeated in enumerate(ties_set):
extensions.append(list(itertools.permutations(repeated)))
extensions = list(itertools.product(*extensions))
# Loop through all possible linear extensions.
for extension in extensions:
# Start to modify the original ranking to create the linear extension.
permu = rank
# Swap the rankings of the repeated / tie scores iteratively.
for section in extension:
# Size of the section to be replaced.
sec_size = len(section)
# Determine the starting point in ranking in which we replace
# the section.
for start, value in enumerate(permu):
if value in section:
break
# Modify the original ranking iteratively.
permu = np.concatenate((permu[:start], section, permu[start + sec_size:]))
# Add the linear extension to the permutation set.
permus.append(permu + 1)
weights.append(1.0 / len(extensions))
return permus, weights
# + id="5BHaItMW5HFk"
def load_permus_from_file(prefix, algorithms, num_instances=10, num_reps=20):
permus = []
scores = []
problems = []
weights = []
dfs = [pd.read_csv(prefix + '-' + algorithm + '.csv', header=[0],
index_col=[0,1]) for algorithm in algorithms]
for df in dfs:
index, problems = fix_index(df)
df.index = index
df = df.astype(int)
for i, problem in enumerate(problems):
for instance in range(num_instances):
for rep in range(num_reps):
# Score of each algorithm.
score = []
for df in dfs:
# Locate the score for each algorithm per problem / instance / rep.
score.append(df.loc[(problem, str(rep + 1)), str(instance)])
# Obtain the rankings, including linear extensions.
p, w = ranks_from_score(score)
scores.append(score)
permus += p
weights += w
return np.array(permus), np.array(weights), np.array(scores)
# + id="bRYjFj2iQXgJ"
def sample_permus(permus, weights, num_samples):
n = len(weights)
sample_permus = []
sample_weights = []
while True:
idx = np.random.randint(0, n)
permu = permus[idx]
w = weights[idx]
if np.random.random() < w:
sample_permus.append(permu)
sample_weights.append(w)
if len(sample_weights) == num_samples:
return np.array(sample_permus), np.array(sample_weights)
# + id="pKcZVLN3QXlx"
# + id="6V1_vvOWRSR_"
# + id="At0AEgOHV22u"
# + [markdown] id="qvSKSYLEMU4v"
# ### Joint Taillard + Random instances
# + id="JAiTL2rMMYCo"
algorithms = ['A', 'B', 'AGA', 'VNS', 'NVNS']
orderingsT, weightsT, scoresT = load_permus_from_file('FSPData/T', algorithms)
orderingsR, weightsR, scoresR = load_permus_from_file('FSPData/R', algorithms)
orderings = np.concatenate((orderingsT, orderingsR), axis=0)
weights = np.concatenate((weightsT, weightsR))
orderings, weights = sample_permus(orderings, weights, 1000)
rankings = np.argsort(orderings, axis=1) + 1
p_top_ranking = empirical_top_ranking_probs(orderings)
p_better_than = empirical_better_than(orderings)
p_top_k = empirical_top_k(orderings)
# + id="OlwylS6QVLiT"
p_top_k = empirical_top_k(orderings)
# + colab={"base_uri": "https://localhost:8080/"} id="-4v1arP4NLZA" outputId="987ec136-890f-4762-9405-f92e1017382d"
probs = calculate_top_ranking_probs(orderings)
# + colab={"base_uri": "https://localhost:8080/", "height": 190} id="06AvzaRnNL1o" outputId="20864a65-7107-4f4c-d96b-7e232890fcb5"
plot_top_ranking_probs("TopOneTaillardRI", model_names=['PLD', 'PLG', 'BT', 'MM'], algorithm_names=algorithms, probs=probs, empirical=p_top_ranking)
# + id="dYjHMB_ENO3F"
# + colab={"base_uri": "https://localhost:8080/"} id="MkmAqezKNO5y" outputId="7974cc85-8d3d-4b63-9e47-5d47fde1ad42"
probs = calculate_better_than_probs(orderings)
# + colab={"base_uri": "https://localhost:8080/", "height": 516} id="qBCGPBZONQvF" outputId="b93552ad-b9ab-46f9-ae64-21200c58ec67"
plot_better_than_probs("BetterThanTaillardRI", model_names=['PLD', 'PLG', 'BT', 'MM'], algorithm_names=algorithms, probs=probs, empirical=p_better_than)
# + colab={"base_uri": "https://localhost:8080/"} id="_GVfC3xFbde7" outputId="0770e6a4-6d3d-4109-e543-d50c179b0937"
probs = calculate_top_k_probs(orderings)
# + colab={"base_uri": "https://localhost:8080/", "height": 516} id="sK--qo8gbgmp" outputId="519cb97c-4696-44e4-950c-c3b2bc6b7e4c"
plot_top_k_probs("TopKTaillardRI", model_names=['PLD', 'PLG', 'BT', 'MM'], algorithm_names=algorithms, probs=probs, empirical=p_top_k)
|
.ipynb_checkpoints/BayesPerm_PresentationCode-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import time
from tqdm import tqdm
import torch
import torch.nn as nn
import torch.optim
import torch.utils.data
import torchvision.transforms as transforms
import torchvision.datasets as datasets
import torchvision.models as models
# -
DATA_PATH = 'data'
train_path = os.path.join(DATA_PATH, 'train')
val_path = os.path.join(DATA_PATH, 'val')
test_path = os.path.join(DATA_PATH, 'test')
normalize = transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
train_dataset = datasets.ImageFolder(train_path, transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normalize
]))
valid_dataset = datasets.ImageFolder(val_path, transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
normalize,
]))
print(len(train_dataset))
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=32,
num_workers=4
)
valid_loader = torch.utils.data.DataLoader(
valid_dataset,
batch_size=32,
num_workers=4
)
# +
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self, num_classes):
super().__init__()
self.pool = nn.MaxPool2d(2, 2)
self.conv1 = nn.Conv2d(3, 16, kernel_size=3, padding=1)
self.conv2 = nn.Conv2d(16, 32, kernel_size=3, padding=1)
self.conv2_bn = nn.BatchNorm2d(32)
self.conv3 = nn.Conv2d(32, 64, kernel_size=3, padding=1)
self.conv4 = nn.Conv2d(64, 128, kernel_size=3, padding=1)
self.conv4_bn = nn.BatchNorm2d(128)
self.conv5 = nn.Conv2d(128, 256, kernel_size=3, padding=1)
self.conv6 = nn.Conv2d(256, 512, kernel_size=3, padding=1)
self.conv6_bn = nn.BatchNorm2d(512)
self.conv7 = nn.Conv2d(512, 512, kernel_size=3, padding=1)
self.conv8 = nn.Conv2d(512, 512, kernel_size=3, padding=1)
self.conv8_bn = nn.BatchNorm2d(512)
# self.conv9 = nn.Conv2d(512, 512, kernel_size=3, padding=1)
# self.conv10 = nn.Conv2d(512, 512, kernel_size=3, padding=1)
# self.conv10_bn = nn.BatchNorm2d(512)
self.fc1 = nn.Linear(512 * 14 * 14, 2048)
self.fc2 = nn.Linear(2048, num_classes)
def forward(self, x):
in_size = x.size(0)
x = self.conv1(x)
x = F.relu(x)
x = self.pool(F.relu(self.conv2_bn(self.conv2(x))))
x = self.conv3(x)
x = F.relu(x)
x = self.pool(F.relu(self.conv4_bn(self.conv4(x))))
x = self.conv5(x)
x = F.relu(x)
x = self.pool(F.relu(self.conv6_bn(self.conv6(x))))
x = self.conv7(x)
x = F.relu(x)
x = self.pool(F.relu(self.conv8_bn(self.conv8(x))))
# x = self.conv9(x)
# x = F.relu(x)
# x = self.pool(F.relu(self.conv10_bn(self.conv10(x))))
x = x.view(-1, 512*14*14)
x = F.dropout(F.relu(self.fc1(x)), training=self.training, p=0.4)
x = self.fc2(x)
return F.log_softmax(x, dim=1)
# -
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
device
# +
from torchsummary import summary
model = Net(196)
model.to(device)
summary(model, (3, 224, 224))
# -
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), 0.01, momentum=0.9)
def train(model,device, train_loader, epoch):
model.train()
for data in tqdm(train_loader):
x,y= data
x=x.to(device)
y=y.to(device)
optimizer.zero_grad()
y_hat= model(x)
loss = criterion(y_hat, y)
loss.backward()
optimizer.step()
print ('Train Epoch: {}\t Loss: {:.6f}'.format(epoch,loss.item()))
def valid(model, device, valid_loader):
model.eval()
valid_loss = 0
correct = 0
with torch.no_grad():
for data in tqdm(valid_loader):
x,y= data
x=x.to(device)
y=y.to(device)
optimizer.zero_grad()
y_hat = model(x)
valid_loss += criterion(y_hat, y).item() # sum up batch loss
pred = y_hat.max(1, keepdim=True)[1] # get the index of the max log-probability
correct += pred.eq(y.view_as(pred)).sum().item()
valid_loss /= len(valid_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
valid_loss, correct, len(valid_dataset), 100. * correct / len(valid_dataset)))
for epoch in range(1, 20):
train(model=model, device=device, train_loader=train_loader, epoch=epoch)
valid(model=model, device=device, valid_loader=valid_loader)
1/196 vs 9/1628
|
stanford-cars-dataset/build_model_from_scratch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from NumbaMinpack import hybrd, lmdif, minpack_sig
from numba import njit, cfunc
import numpy as np
from scipy.optimize import root
# +
@cfunc(minpack_sig)
def myfunc(x, fvec, args):
fvec[0] = x[0]**2 - 30.0*x[1]
fvec[1] = x[1]**2 - 8.0*x[0]
funcptr = myfunc.address
@njit
def myfunc_scipy(x):
return np.array([x[0]**2 - 30.0*x[1],
x[1]**2 - 8.0*x[0]])
# -
x_init = np.array([10.0,10.0])
neqs = 2
args = np.array([0.0])
# +
xsol, fvec, success, info = lmdif(funcptr, x_init, neqs, args)
sol_sp = root(myfunc_scipy,x_init,method='lm')
print('NumbaMinpack (lmdif):',xsol)
print('scipy (lmdif): ',sol_sp.x)
print()
xsol, fvec, success, info = hybrd(funcptr, x_init, args)
sol_sp = root(myfunc_scipy,x_init,method='hybr')
print('NumbaMinpack (hybrd):',xsol)
print('scipy (hybrd): ',sol_sp.x)
# -
# For small problems that take very little time within Minpack, NumbaMinpack will be faster than scipy. This is because scipy sets up the optimization problem in python, which can take more time then the actual optimization. For larger optimization problems, scipy and NumbaMinpack should take about the same amount of time.
# %timeit lmdif(funcptr, x_init, neqs, args)
# %timeit root(myfunc_scipy,x_init,method='lm')
print()
# %timeit hybrd(funcptr, x_init, args)
# %timeit root(myfunc_scipy,x_init,method='hybr')
# NumbaMinpack within jit compiled function works
@njit
def test():
sol = lmdif(funcptr, x_init, neqs, args)
return sol
test()
# scipy within jit compiled function does not work
@njit
def test_sp():
sol_sp = root(myfunc_scipy,x_init,method='lm')
return sol_sp
test_sp()
|
comparison2scipy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# # !pip3 install numpy taichi numba matplotlib
# -
import numpy as np
import taichi as ti
from numba import jit
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Множество Мандельброта
# Рассмотрим в качестве примера кода для ускорения отрисовку [множества Мандельброта](https://ru.wikipedia.org/wiki/%D0%9C%D0%BD%D0%BE%D0%B6%D0%B5%D1%81%D1%82%D0%B2%D0%BE_%D0%9C%D0%B0%D0%BD%D0%B4%D0%B5%D0%BB%D1%8C%D0%B1%D1%80%D0%BE%D1%82%D0%B0) -- одного из наиболее известных фракталов.
#
# Если отрисовать Мандельброта, то должно получиться что-то похожее на данную картинку
# 
#
# Ниже представлен код для вычисления множества Мандельброта в цветном варианте.
# Если вкратце, то алгоритм вычисления множества Мандельброта таков:
#
# * для каждой точки на плоскости с координатами $(x, y)$ мы формируем комлексное число $c = x + iy$. Стоит отметить, что для того, чтобы мы увидили что-то интересное, то у следует задать пределы для $ (-2 < x < 1), (-1.5 < y < 1.5)$
# * задаем число $Z_0 = 0$
# * запускаем вычисление последовательности $ Z_n = Z_{n-1} ^ 2 + C$
# В случае, если последовательность ограничена, то точка принадлежит ко множеству Мандельброта, иначе нет.
#
# Так как мы не можем вычислять бесконечную последовательность, то мы задаем максимаьное количество итераций $N$.
# Также не имеет смысла продолжать вычисления, если $abs(Z_n) > 2$
#
# * Если на предыдущем шаге закрашивать точку в цвет, соответствующий тому, на каком шаге мы закончили вычисление последовательности $Z_n$, то у нас получится цветной вариант множества. Где само множество по-прежнему будет черным, а на его границах будет градиент.
#
def naive_mandelbrot(
w, h, min_x, max_x, min_y, max_y, max_iterations
):
# Create empty image with shape = (w, h)
image= np.zeros((h, w), dtype=np.float32)
# Compute steps around X and Y axis
dx = (max_x - min_x) / w
dy = (max_y - min_y) / h
# Iterate over X axis
for x in range(w):
real = min_x + x * dx
# Iterate over Y axis
for y in range(h):
# Compute point C and Z_0
im = min_y + y * dy
c = real + 1j * im
z = 0
# Compute the color of the point
for k in range(max_iterations):
z = z ** 2 + c
if abs(z) > 2:
image[y, x] = k / max_iterations
break
return image
# Проверим, что все работает правильно. У нас должно получиться нечто, похожее на изображение выше.
image = naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
plt.figure(figsize=(10, 10))
plt.axis('off')
plt.imshow(image, cmap='hot')
# Давайте теперь поиграемся с параметром N, отвечающим за количество итераций:
# +
n_sequence = [2, 5, 10, 20, 50, 255]
fig, axs = plt.subplots(3, 2, figsize=(20, 20))
for i, ax in enumerate(fig.get_axes()):
n = n_sequence[i]
image = naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, n)
ax.set_title(f'N={n}')
ax.axis('off')
ax.imshow(image, cmap='hot')
# -
# Очевидно, что чем болльше у нас параметр N, тем более детальное множество Мандельброта. Но и тем больше нам необходимо провести вычислений.
# Давайте теперь замерим скорость работы:
# %timeit naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 2)
# %timeit naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 5)
# %timeit naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 10)
# %timeit naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 20)
# %timeit naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 50)
# %timeit naive_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
# ## Numpy
#
# Как мы видим, отрисовка довольно медленная. Давайте попробуем переписать код с использованием Numpy.
def numpy_mandelbrot(
w, h, min_x, max_x, min_y, max_y, max_iterations
):
# Create empty image with shape = (w, h)
image= np.zeros((h, w), dtype=np.float64)
# Compute points C and Z_0
x_linspace = np.linspace(min_x, max_x, w)
y_linspace = np.linspace(min_y, max_y, h)
c = x_linspace[np.newaxis, :] +\
1j * y_linspace[:, np.newaxis]
z = np.zeros_like(c)
# Compute the colors for all points
for k in range(max_iterations):
mask = image == 0
z[mask] = z[mask] ** 2 + c[mask]
mask = (mask & (np.abs(z) > 2))
image[mask] = float(k) / max_iterations
return image
# Проверим, что рисуется то, что нужно
image = numpy_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
plt.figure(figsize=(10, 10))
plt.axis('off')
plt.imshow(image, cmap='hot')
# %timeit numpy_mandelbrot(640, 480, -1, 2, -1.5, 1.5, 255)
# ## Cython
#
# Уже намного быстрее, но все равно недостаточно. Давайте попробуем запустить то же самое, но с использованием Cython. В начале мы вообще никак не будем менять код исходной функции `naive_mandelbrot`.
# %load_ext Cython
# + language="cython"
#
# import numpy as np
#
# def cython_mandelbrot(
# w, h, min_x, max_x, min_y, max_y, max_iterations
# ):
# # Create empty image with shape = (w, h)
# image= np.zeros((h, w), dtype=np.float32)
# # Compute steps around X and Y axis
# dx = (max_x - min_x) / w
# dy = (max_y - min_y) / h
#
# # Iterate over X axis
# for x in range(w):
# real = min_x + x * dx
# # Iterate over Y axis
# for y in range(h):
# # Compute point C and Z_0
# im = min_y + y * dy
# c = real + 1j * im
# z = 0
# # Compute the color of the point
# for k in range(max_iterations):
# z = z ** 2 + c
# if abs(z) > 2:
# image[y, x] = k / max_iterations
# break
# return image
# -
# Отрисуем:
image = cython_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
plt.figure(figsize=(10, 10))
plt.axis('off')
plt.imshow(image, cmap='hot')
# %timeit cython_mandelbrot(640, 480, -1, 2, -1.5, 1.5, 255)
# Это уже быстрее, чем использование чистого Python, но все же очень медленно. Да еще и медленнее Numpy-версии.
#
# Давайте теперь скомилируем Cython с флагом `-a` или `--annotate`, который покажет нам насколько у нас активно происходит взаимодействие Python и C.
# + magic_args="-a" language="cython"
#
# import numpy as np
#
# def cython_mandelbrot(
# w, h, min_x, max_x, min_y, max_y, max_iterations
# ):
# # Create empty image with shape = (w, h)
# image= np.zeros((h, w), dtype=np.float32)
# # Compute steps around X and Y axis
# dx = (max_x - min_x) / w
# dy = (max_y - min_y) / h
#
# # Iterate over X axis
# for x in range(w):
# real = min_x + x * dx
# # Iterate over Y axis
# for y in range(h):
# # Compute point C and Z_0
# im = min_y + y * dy
# c = real + 1j * im,
# z = 0
# # Compute the color of the point
# for k in range(max_iterations):
# z = z ** 2 + c
# if abs(z) > 2:
# image[y, x] = k / max_iterations
# break
# return image
# -
# Как мы видим, здесь очень интенсивно происходят конвертации из типов Python и C. Надо менять код.
#
# В примере ниже мы использовали почти все приколы:
# - Устранили всякие проверки индексов массива
# - Используем `cdivision`
# - Добавили всяких штук типа `cdef float[:,:] view = image` и `cdef complex`
# + magic_args="-a" language="cython"
#
# import numpy as np
# import cython
#
# @cython.cdivision(True)
# def cython_mandelbrot(
# int w, int h, float min_x, float max_x,
# float min_y, float max_y, int max_iterations
# ):
# # Create empty image with shape = (w, h)
# image = np.zeros((h, w), dtype=np.float32)
# # Define c-typed variables
# cdef float[:,:] view = image
#
# cdef complex z, c, I = 1j
# cdef int x, y, k
# cdef float dx, dy, real, im
#
# dx = <float>(max_x - min_x) / w
# dy = <float>(max_y - min_y) / h
#
# for x in range(w):
# real = min_x + x * dx
# for y in range(h):
# im = min_y + y * dy
# c = real + I * im
# z = 0
# for k in range(max_iterations):
# z = z ** 2 + c
# if abs(z) > 2:
# view[y, x] = <float>(k) / max_iterations
# break
# return image
# -
# %timeit cython_mandelbrot(640, 480, -1, 2, -1.5, 1.5, 255)
# Добавим распараллеливание при помощи OpenMP (используем `prange` вместо `range`)
#
# + magic_args="-a" language="cython"
# #distutils: extra_compile_args=-fopenmp
# #distutils: extra_link_args=-fopenmp
#
# import cython
# import numpy as np
# from cython.parallel import prange
#
# @cython.boundscheck(False)
# @cython.cdivision(True)
# def cython_mandelbrot(
# int w, int h, float min_x, float max_x,
# float min_y, float max_y, int max_iterations
# ):
# image = np.zeros((h, w), dtype=np.float32)
# cdef float[:,:] view = image
#
# cdef complex z, c, I = 1j
# cdef int x, y, k
# cdef float dx, dy, real, im
#
# dx = (max_x - min_x) / w
# dy = (max_y - min_y) / h
#
# for x in prange(w, nogil=True):
# real = min_x + x * dx
# for y in range(h):
# im = min_y + y * dy
# c = real + I * im
# z = 0
# for k in range(max_iterations):
# z = z ** 2 + c
# if abs(z) > 2:
# view[y, x] = <float>(k) / max_iterations
# break
# return image
# -
image = cython_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
plt.figure(figsize=(10, 10))
plt.axis('off')
plt.imshow(image, cmap='hot')
# %timeit cython_mandelbrot(640, 480, -1, 2, -1.5, 1.5, 255)
# Хм, результаты превосходят Numpy многократно! Но все же наш код стал ближе к C++, чем к Python. Можем ли мы также ускорить код, но без вот этих всех штук?
# ## Numba
#
# Возьмем опять наш исходный код и добавим всего лишь один декоратор с разными параметрами.
@jit(nopython=True)
def numba_mandelbrot(
w, h, min_x, max_x, min_y, max_y, max_iterations
):
# Create empty image with shape = (w, h)
image= np.zeros((h, w), dtype=np.float32)
# Compute steps around X and Y axis
dx = (max_x - min_x) / w
dy = (max_y - min_y) / h
# Iterate over X axis
for x in range(w):
real = min_x + x * dx
# Iterate over Y axis
for y in range(h):
# Compute point C and Z_0
im = min_y + y * dy
c = real + 1j * im
z = 0
# Compute the color of the point
for k in range(max_iterations):
z = z ** 2 + c
if abs(z) > 2:
image[y, x] = k / max_iterations
break
return image
image = numba_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
plt.figure(figsize=(10, 10))
plt.axis('off')
plt.imshow(image, cmap='hot')
# %timeit numba_mandelbrot(640, 480, -1, 2, -1.5, 1.5, 255)
# Здорово! Мы получили такой же результат, как и при помощи Cython, но при этом добавили лишь один декоратор!
@jit(nopython=True, fastmath=True)
def numba_mandelbrot(
w, h, min_x, max_x, min_y, max_y, max_iterations
):
# Create empty image with shape = (w, h)
image= np.zeros((h, w), dtype=np.float32)
# Compute steps around X and Y axis
dx = (max_x - min_x) / w
dy = (max_y - min_y) / h
# Iterate over X axis
for x in range(w):
real = min_x + x * dx
# Iterate over Y axis
for y in range(h):
# Compute point C and Z_0
im = min_y + y * dy
c = real + 1j * im
z = 0
# Compute the color of the point
for k in range(max_iterations):
z = z ** 2 + c
if abs(z) > 2:
image[y, x] = k / max_iterations
break
return image
# %timeit numba_mandelbrot(640, 480, -1, 2, -1.5, 1.5, 255)
# Добавим параллелизма:
# +
from numba import prange
@jit(nopython=True, fastmath=True, parallel=True)
def numba_mandelbrot(
w, h, min_x, max_x, min_y, max_y, max_iterations
):
# Create empty image with shape = (w, h)
image= np.zeros((h, w), dtype=np.float32)
# Compute steps around X and Y axis
dx = (max_x - min_x) / w
dy = (max_y - min_y) / h
# Iterate over X axis
for x in prange(w):
real = min_x + x * dx
# Iterate over Y axis
for y in range(h):
# Compute point C and Z_0
im = min_y + y * dy
c = real + 1j * im
z = 0
# Compute the color of the point
for k in range(max_iterations):
z = z ** 2 + c
if abs(z) > 2:
image[y, x] = k / max_iterations
break
return image
# -
# %timeit numba_mandelbrot(640, 480, -1, 2, -1.5, 1.5, 255)
# ## Taichi
#
# Taichi это относительно диковинная библиотека, на которую я наткнулся случайно. Однако, она обладает очень впечатляющими возможностями.
# +
ti.init(arch=ti.cpu)
@ti.func
def complex_sqr(z):
return ti.Vector([z[0]**2 - z[1]**2, z[1] * z[0] * 2])
@ti.kernel
def mandelbrot_kernel(
min_x: ti.float32, max_x: ti.float32,
min_y: ti.float32, max_y: ti.float32,
max_iterations: ti.int32,
image: ti.ext_arr()
):
h, w = image.shape[:2]
dx = (max_x - min_x) / w
dy = (max_y - min_y) / h
for y, x in image:
c = ti.Vector([min_x + x * dx, min_y + y * dy])
z = ti.Vector([0.0, 0.0])
for i in range(max_iterations):
z = complex_sqr(z) + c
if z.norm() > 2:
image[y, x] = i / max_iterations
break
def taichi_mandelbrot(w, h, min_x, max_x, min_y, max_y, max_iterations):
image= np.zeros((h, w), dtype=np.float64)
mandelbrot_kernel(min_x, max_x, min_y, max_y, max_iterations, image)
return image
# -
image_np = taichi_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
plt.figure(figsize=(20, 20))
plt.imshow(image_np, cmap='hot')
# %timeit taichi_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
ti.reset()
# +
ti.init(arch=ti.gpu)
@ti.func
def complex_sqr(z):
return ti.Vector([z[0]**2 - z[1]**2, z[1] * z[0] * 2])
@ti.kernel
def mandelbrot_kernel(
min_x: ti.float32, max_x: ti.float32,
min_y: ti.float32, max_y: ti.float32,
max_iterations: ti.int32,
image: ti.ext_arr()
):
h, w = image.shape[:2]
dx = (max_x - min_x) / w
dy = (max_y - min_y) / h
for y, x in image:
c = ti.Vector([min_x + x * dx, min_y + y * dy])
z = ti.Vector([0.0, 0.0])
for i in range(max_iterations):
z = complex_sqr(z) + c
if z.norm() > 2:
image[y, x] = i / max_iterations
break
def taichi_mandelbrot(w, h, min_x, max_x, min_y, max_y, max_iterations):
image= np.zeros((h, w), dtype=np.float64)
mandelbrot_kernel(min_x, max_x, min_y, max_y, max_iterations, image)
return image
# -
# %timeit taichi_mandelbrot(640, 480, -2, 1, -1.5, 1.5, 255)
ti.reset()
|
mandelbrot_set.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas - Series
#
# Elemento mais básico (data + indices)
#
# Muito semelhante a um dicionário
#
# Pode ser criado a partir de outros objetos
#
# Tem rótulos de eixos: pode ser indexado por um rótulo, em vez de apenas uma localização numérica.
#
# Também pode conter qualquer objeto Python arbitrário
# +
# Import Library
import numpy as np
import pandas as pd
#from pandas import Series, DataFrame
# Criando uma Serie Vazia
# data: Objetos que vão ser passados
# index: forma que vai acessar esses objetos
pd.Series(data=None, index=None)
# +
# Criando Series - A partir de outros objetos
labels = ['a','b','c']
list1 = [10,20,30]
arr1 = np.array([10,20,30])
dic1 = {'a':10,'b':20,'c':30}
# Criando Series - indices numericos (default)
series1 = pd.Series(data = list1) # same as series1 = pd.Series(list1)
print(series1)
print()
# Atributos de uma serie
print(series1.dtype)
print(series1.values)
print(series1.index)
print()
# Criando Series - a partir de lista e labels
series2 = pd.Series(list1,labels) # same as series2 = pd.Series(data = list1, index = labels)
print(series2)
print()
# Criando Series - a partir de array e labels
series3 = pd.Series(arr1,labels)
print(series3)
print()
# Criando Series - a partir de um dicionario 1
series4 = pd.Series(dic1)
print(series4)
print()
# Criando Series - a partir de um dicionario 2
dic2 = {'Sp':3500,'Rj':4000,'Am':4500,'Bsb':100}
order = ['Bsb','Am','Sp','Rj','Ce']
series5 = pd.Series(dic2,index = order)
print(series5)
print()
# Criando Series - a partir de uma lista de funções (objetos)
series6 = pd.Series([sum,print,len])
print(series6)
# +
# Acessando/Filtrando Valores/Indices
series7 = pd.Series([10,20,30,40,50],['a','b','c','d','e'])
print(series7)
print()
print(series7['b'])
print()
print(series7[['a','c']])
print()
print(series7[series7<30])
print()
print(series7*2)
print()
print('c' in series7)
print('x' in series7)
# +
# Operações sobre series, utilizando numpy
series1 = pd.Series([1,2,3])
print(np.exp(series1))
print()
# Operações entre series
ser4 = pd.Series([1,2,3,4],index = ['EUA','Alemanha','URSS','Japão'])
ser5 = pd.Series([1,2,3,4],index = ['EUA','Alemanha','Itália','Japão'])
ser6 = ser4 + ser5 # soma os valores que tem mesmo index
print(ser6)
print()
# Obs: quando não há índice nas duas series: retorna "NaN"
# Obs: operações: + - * / **
|
scripts_numpy_pandas/Pandas_01_series.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# import datasets
data=pd.read_csv('./datasets/mushrooms.csv')
print(data.shape)
# +
### the data is alphabetical we need to convert it to numerical data ###
# -
# # encoding of data
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
le=LabelEncoder()
ds=data.apply(le.fit_transform)
ds=ds.values
print(ds)
X=ds[:,1:]
Y=ds[:,0]
print(Y)
print(X[:5,:])
# +
# break data into test and train
# -
xtrain,xtest,ytrain,ytest=train_test_split(X,Y,test_size=0.2)
print(xtrain.shape,ytrain.shape)
print(xtest.shape,ytest.shape)
# # BUILDING OUR CLASSIFIER
a=np.array([1,1,0,0,1,0,1,0,1,0])
np.sum(a==1)
def prior_prob(y_train,label):
total_examples=y_train.shape[0]
class_examples=np.sum(y_train==label)
return class_examples/float(total_examples)
prior_prob(a,1)
def conditional_probability(x_train,y_train,feature_col,feature_val,label):
x_filtered=x_train[y_train==label]
numerator=np.sum(x_filtered[:,feature_col]==feature_val)
denominator=np.sum(y_train==label)
return numerator/float(denominator)
def predict(x_train,y_train,xtest):
### xtest is a single example ###
classes=np.unique(y_train)
n_feature=x_train.shape[1]
post_prob=[]
for label in classes:
likelihood=1.0
for f in range(n_feature):
cond=conditional_probability(x_train,y_train,f,xtest[f],label)
likelihood*=cond
prior=prior_prob(y_train,label)
post=likelihood*prior
post_prob.append(post)
return np.argmax(post_prob)
output=predict(xtrain,ytrain,xtest[4])
print(output)
# +
ypred=[]
for i in range(xtest.shape[0]):
y=predict(xtrain,ytrain,xtest[i])
ypred.append(y)
yped=np.array(ypred)
# -
print(yped)
print(ytest)
from sklearn.metrics import accuracy_score
acc=accuracy_score(ytest,ypred)
print(acc)
|
MachineLearning/Naive Bayes Classifier/.ipynb_checkpoints/NaiveBayes-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chrishuskey/DS-Unit-2-Linear-Models/blob/master/module2-regression-2/Assignment_DS_212_Regression_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="cScljW0znyhn" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 2*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Regression 2
#
# ## Assignment
#
# You'll continue to **predict how much it costs to rent an apartment in NYC,** using the dataset from renthop.com.
#
# - [✓] Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# - [✓] Engineer at least two new features. (See below for explanation & ideas.)
# - [✓] Fit a linear regression model with at least two features.
# - [✓] Get the model's coefficients and intercept.
# - [✓] Get regression metrics RMSE, MAE, and $R^2$, for both the train and test data.
# - [✓] What's the best test MAE you can get? Share your score and features used with your cohort on Slack!
# - [✓] As always, commit your notebook to your fork of the GitHub repo.
#
#
# #### [Feature Engineering](https://en.wikipedia.org/wiki/Feature_engineering)
#
# > "Some machine learning projects succeed and some fail. What makes the difference? Easily the most important factor is the features used." — <NAME>, ["A Few Useful Things to Know about Machine Learning"](https://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf)
#
# > "Coming up with features is difficult, time-consuming, requires expert knowledge. 'Applied machine learning' is basically feature engineering." — <NAME>, [Machine Learning and AI via Brain simulations](https://forum.stanford.edu/events/2011/2011slides/plenary/2011plenaryNg.pdf)
#
# > Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work.
#
# #### Feature Ideas
# - Does the apartment have a description?
# - How long is the description?
# - How many total perks does each apartment have?
# - Are cats _or_ dogs allowed?
# - Are cats _and_ dogs allowed?
# - Total number of rooms (beds + baths)
# - Ratio of beds to baths
# - What's the neighborhood, based on address or latitude & longitude?
#
# ## Stretch Goals
# - [ ] If you want more math, skim [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapter 3.1, Simple Linear Regression, & Chapter 3.2, Multiple Linear Regression
# - [ ] If you want more introduction, watch [<NAME>, Statistics 101: Simple Linear Regression](https://www.youtube.com/watch?v=ZkjP5RJLQF4)
# (20 minutes, over 1 million views)
# - [✓] Add your own stretch goal(s) !: Experiment with better feature selection using the model coefficients (LinReg coeffs + SelectFromModel), Recursive Feature Elimination, etc.
# + id="Esmzis4XdOVz" colab_type="code" colab={}
# Import libraries:
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import datetime
from math import sqrt
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + colab_type="code" id="cvrw-T3bZOuW" colab={}
# Read New York City apartment rental listing data
rent_data = pd.read_csv(DATA_PATH+'apartments/renthop-nyc.csv')
assert rent_data.shape == (49352, 34)
# + id="b0pBg92ic_w-" colab_type="code" colab={}
# Remove the most extreme 0.1% of prices,
# the most extreme 0.1% of latitudes, &
# the most extreme 0.1% of longitudes:
rent_data = rent_data[(rent_data['price'] >= rent_data['price'].quantile(0.001)) &
(rent_data['price'] <= rent_data['price'].quantile(0.999)) &
(rent_data['latitude'] >= rent_data['latitude'].quantile(0.001)) &
(rent_data['latitude'] <= rent_data['latitude'].quantile(0.999)) &
(rent_data['longitude'] >= rent_data['longitude'].quantile(0.001)) &
(rent_data['longitude'] <= rent_data['longitude'].quantile(0.999))]
# + id="3nvVBP64fvyM" colab_type="code" colab={}
# Change to the right data types:
rent_data['created'] = pd.to_datetime(rent_data['created'], infer_datetime_format=True) # Note: Using infer_datetime_format can supposedly be up to 5-10x faster w.r.t. processing time
# + id="pPYr1ajPevIv" colab_type="code" colab={}
# Split into training and test data:
# Training data: listings from April and May 2016
# Test data: listings from June 2016
working_data = rent_data[(rent_data['created'].dt.month <= 5)]
test = rent_data[rent_data['created'].dt.month == 6]
# Split working_data into train and val sets:
train, val = train_test_split(working_data, train_size=0.75, shuffle=True, random_state=42)
# Check to make sure the resulting datasets have the right numbers of
# observations (and that we got all of them) and features:
assert (working_data.shape[0] + test.shape[0] == rent_data.shape[0]) & (
working_data.shape[1] == test.shape[1] == rent_data.shape[1])
assert (train.shape[0] + val.shape[0] + test.shape[0] == rent_data.shape[0]) & (
train.shape[1] == val.shape[1] == test.shape[1] == rent_data.shape[1])
# + id="blqkYvjlEwZO" colab_type="code" colab={}
# Function that implements all data prep. on input train/val/test datasets
# in the same way:
def data_prep(dataframe):
# Make copy to work with:
df = dataframe.copy()
# Add new feature: Total # of bedrooms and bathrooms:
df['bedrooms+bathrooms'] = df['bedrooms'] + df['bathrooms']
# Add new feature: Perks by price tiers:
# The current features for amenities are all binary 0/1
# variables better suited to classification approaches. But since we're
# required to use linear regression to predict price here instead,
# one way to improve price-predicting power would be to group perks by
# corresponding price level (based on rarity in the data set, with some
# manual adjustments based on intuition / "domain knowledge"):
# (1) Level 1 perks: slight price-boosters
# (2) Level 2 perks: higher end perks indicative of higher-rent apartments
# (3) Level 3 perks: luxury perks indicative of very expensive apartments
df['L1_price_boost_perks'] = df['elevator'] + (df['cats_allowed'] & df['dogs_allowed']) + df['laundry_in_building']
df['L2_high_end_perks'] = df['hardwood_floors'] + df['doorman'] + df['dishwasher'] + df['fitness_center'] + df['pre-war'] + df['roof_deck'] + df['high_speed_internet']
df['L3_luxury_perks'] = df['swimming_pool'] + df['laundry_in_unit'] + df['terrace'] + df['balcony'] + df['new_construction'] + df['loft']
# Pull info from datetime, bc we can't feed type datetime into sklearn models:
df['created_year'] = df['created'].dt.year
df['created_month'] = df['created'].dt.month
# Convert "interest_level" to ordinal (1/2/3 numerical representation),
# so we can work with this feature more easily:
df['interest_level'] = df['interest_level'].replace({'low': 1, 'medium': 2, 'high': 3})
# ------------------------------------------------------------------------------
# [?? TO DO all below just for practice with aspects of Pandas/Python -- these
# are all things I'm not 100% sure how to do, but should know how to do! ??]]
# [?? What to do about the warnings below? I'm getting the same warning x2
# when I use .loc instead to do the same as the above... ??]
# [?? Luxury:
# contains "luxury" in description (but this would be a 0/1 binary feature...
# only useful for classification, or can we use with regression too?) ??]
# [?? For the perks feature above, what is a better way to do this? How would
# a top tier data scientist frame this problem, if constrained to only using a
# linear regression model and this starter dataset ??]
# [?? Column selection based on conditions/criteria: Sort perks into medium/price-boost, high-end/premium perks and luxury
# perks automatically, by sorting the column names based on conditions:
# median as 1 or 0, 75% percentile as 1 or 0, 90% quantile as 1 or 0.
# Not sure how to work with columns this way, only rows!!... --> need to learn ??]
# df_a = train.copy()
# a = df_a.median() == 0
# a = pd.DataFrame(a)
# a = a.reset_index()
# a.columns = ['index', 'criterion']
# b = a[a['criterion'] == True]
# b
# # for i in a:
# # if a.loc[i, criterion] == True:
# # print(a[i].index)
# # # pd.DataFrame(data=train, column=list)
return df
# + id="xC2ev3fHEyz5" colab_type="code" colab={}
# Implement data prep on each of train, val, test:
train = data_prep(train)
val = data_prep(val)
test = data_prep(test)
# + id="-_54KfXjlR5M" colab_type="code" colab={}
# # [?? To Do: Improve new feature #2 by first weighting the following features
# # by rarity and likely level of demand, rather than just adding up all of the
# # 1's (has/doesn't have x amenity) ??]
# # Categorize as: Price premium features:
# # Median and up:
# 'elevator',
# ('cats_allowed' & 'dogs_allowed', )
# ('laundry_in_building' or 'laundry_in_unit')
# # Categorize as: High-end perks:
# # 75% and up:
# 'hardwood_floors',
# 'doorman',
# 'dishwasher',
# 'fitness_center',
# 'roof_deck'
# 'high_speed_internet',
# 'pre-war',
# # Categorize as: Luxury perks:
# # Higher %s only (not even 75% has):
# 'swimming_pool'
# 'laundry_in_unit', # means it's more likely to be a larger apt. --> higher price
# 'balcony',
# 'terrace',
# 'new_construction',
# 'loft',
# + [markdown] id="ttZRcBnnSbzo" colab_type="text"
# # **Baselines:**
# + id="0FbJAC80SfUY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="4667c0a8-5bbd-4000-f1a8-9dd6d7783c60"
# DummyRegressor (mean) model to start with:
mean_price = train[target].mean()
# Performance of our dummy regression model:
print('Performance: Baseline #1, Dummy Regression (Mean) Model:\n')
# Performance on Training Set:
print('On Training Set:')
y_true_train = y_train
y_pred_train = [mean_price] * len(y_train)
print(f'MAE: {mean_absolute_error(y_true_train, y_pred_train):.2f}')
mse_train = mean_squared_error(y_true_train, y_pred_train)
print(f'MSE: {mse_train:.2f}')
print(f'RMSE: {sqrt(mse_train):.2f}')
print(f'R^2 score: {r2_score(y_true_train, y_pred_train):.2f}\n')
# Performance on Validation Set:
print('On Validation Set:')
y_true_val = y_val
y_pred_val = [mean_price] * len(y_val)
print(f'MAE: {mean_absolute_error(y_true_val, y_pred_val):.2f}')
mse_val = mean_squared_error(y_true_val, y_pred_val)
print(f'MSE: {mse_val:.2f}')
print(f'RMSE: {sqrt(mse_val):.2f}')
print(f'R^2 score: {r2_score(y_true_val, y_pred_val):.2f}\n')
# + id="YpAOfdoUSA4m" colab_type="code" outputId="b924b785-d02e-4dd7-b5b7-1e24ad764cc6" colab={"base_uri": "https://localhost:8080/", "height": 238}
# Multiple Linear Regression Model for the above NYC apartment rent data:
# Import model class:
from sklearn.linear_model import LinearRegression
# Initiate model:
model = LinearRegression()
# Features matrix and target vector:
features = ['bedrooms+bathrooms', 'interest_level', 'L1_price_boost_perks', 'L2_high_end_perks', 'L3_luxury_perks']
target = 'price'
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
# Fit the model to our training data:
model.fit(X_train, y_train)
# Model Performance:
print('Model Performance: Model #1, LinearRegression\n')
# Error on training set:
print('On Training Set:')
y_true_train = y_train
y_pred_train = model.predict(X_train)
print(f'MAE: {mean_absolute_error(y_true_train, y_pred_train):.1f}')
mse_train = mean_squared_error(y_true_train, y_pred_train)
print(f'MSE: {mse_train:.1f}')
print(f'RMSE: {sqrt(mse_train):.1f}')
print(f'R^2 score: {r2_score(y_true_train, y_pred_train):.2f}\n')
# Error on new data: our test set:
print('On Validation Set:')
y_true_val = y_val
y_pred_val = model.predict(X_val)
print(f'MAE: {mean_absolute_error(y_true_val, y_pred_val):.2f}')
mse_test = mean_squared_error(y_true_val, y_pred_val)
print(f'MSE: {mse_test:.2f}')
print(f'RMSE: {sqrt(mse_test):.2f}')
print(f'R^2 score: {r2_score(y_true_val, y_pred_val):.2f}')
# + [markdown] id="qstF4eOqJrvJ" colab_type="text"
# # **Feature Selection:**
# + [markdown] id="Eruf0gYTg13d" colab_type="text"
# Method 1: By LinearRegression Model Coefficients:
# + id="RGmABqVsePM9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} cellView="both" outputId="816d59cf-b4bf-443a-ea0c-35d77137332a"
# Run linear regression model using all features, so we can then check which
# are most important for our predictions:
# Multiple Linear Regression Model for the above NYC apartment rent data:
# (1) Import model class:
from sklearn.linear_model import LinearRegression
# (2) Initiate model:
fs1_linreg = LinearRegression()
# (3) Define features matrix and target vector:
# Features and target:
target = 'price'
features = train.columns.tolist()
# Also need to remove the target feature, as well as features with types that
# we can't input into sklearn models (e.g., datetime, strings):
features_to_remove = [target, 'created', 'description',
'display_address', 'street_address']
for feature in features_to_remove:
features.remove(feature)
# The resulting matrices & vectors:
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
y_test = test[target]
# (4) Fit the model to our training data:
fs1_linreg.fit(X_train, y_train)
# Performance on training data:
print('Model Performance:\n')
y_true_train = y_train
y_pred_train = fs1_linreg.predict(X_train)
print('On Training Set:')
print(f'Train MAE: {mean_absolute_error(y_true_train, y_pred_train):.1f}')
mse_train = mean_squared_error(y_true_train, y_pred_train)
print(f'Train MSE: {mse_train:.1f}')
print(f'Train RMSE: {sqrt(mse_train):.1f}')
print(f'Train R^2 score: {r2_score(y_true_train, y_pred_train):.2f}\n')
# Performance on validation set:
y_true_val = y_val
y_pred_val = fs1_linreg.predict(X_val)
print('On Validation Set:')
print(f'Train MAE: {mean_absolute_error(y_true_val, y_pred_val):.1f}')
mse_val = mean_squared_error(y_true_val, y_pred_val)
print(f'Train MSE: {mse_val:.1f}')
print(f'Train RMSE: {sqrt(mse_val):.1f}')
print(f'Train R^2 score: {r2_score(y_true_val, y_pred_val):.2f}\n')
# + id="LG-KOTfoeP7U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="54afaee4-2aa1-4078-d8a8-b93f42cb4a57"
# Check linear regression coefficients to get a sense of feature importances:
fs1_linreg_coeffs = pd.Series(fs1_linreg.coef_, X_val.columns)
# Plot coefficients on chart:
n = len(X_val.columns)
plt.figure(figsize=(10, n/2))
fs1_linreg_coeffs.sort_values().plot.barh(color='darkorange')
plt.title('Relative Importance of Features: Coefficients in LinearRegression Model')
plt.show()
# + id="YkvlxawheP4W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3e42ebde-e811-451f-e44a-5f6005166362"
# [?? What?? -- how to work with coefficients in a linear regression model... these make no sense ??]
fs1_linreg_coeffs.loc['bathrooms']
# + id="IeNPkFPYpHEf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="4b18979b-6706-477d-9bd8-6f5c4959d3aa"
from sklearn.feature_selection import SelectFromModel
# Initiate SelectFromModel feature selector:
feature_selector_coeffs_raw = SelectFromModel(estimator=fs1_linreg, threshold='mean')
# Fit feature selector:
feature_selector_coeffs_raw.fit(X_train, y_train)
# + id="XnFHaS9wrLb4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="855b6af1-26a3-4645-dd6f-7e6afdbbf348"
# See which features are supported based on raw coefficients:
feature_names = X_train.columns
pd.DataFrame(feature_selector_coeffs_raw.get_support(),
index=feature_names,
columns=['include']).sort_values(by='include', ascending=False)
# + id="UX5R13PLrQVz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a728a857-1024-4298-854c-48053e201d39"
selected_features_coeffs_raw = feature_names[feature_selector_coeffs_raw.get_support()].tolist()
selected_features_coeffs_raw
# + [markdown] id="BANN2FPmt5jA" colab_type="text"
# Coeffs w/ Normalized Feature Matrices:
# + id="PiBr2AcWhuwJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="32ceee70-e4df-4665-90f7-4459b325e6a2"
# Cross-check with normalized features going into linear regression model:
# Run linear regression model using all features, so we can then check which
# are most important for our predictions:
# Multiple Linear Regression Model for the above NYC apartment rent data:
# (1) Import model class:
from sklearn.linear_model import LinearRegression
# (2) Initiate model:
fs1_linreg_normalized = LinearRegression(normalize=True)
# (3) Define features matrix and target vector:
# Already defined above.
# (4) Fit the model to our training data:
fs1_linreg_normalized.fit(X_train, y_train)
# Performance on training data:
print('Model Performance: LinearRegression w/ Normalized Inputs\n')
y_true_train = y_train
y_pred_train = fs1_linreg_normalized.predict(X_train)
print('On Training Set:')
print(f'Train MAE: {mean_absolute_error(y_true_train, y_pred_train):.1f}')
mse_train = mean_squared_error(y_true_train, y_pred_train)
print(f'Train MSE: {mse_train:.1f}')
print(f'Train RMSE: {sqrt(mse_train):.1f}')
print(f'Train R^2 score: {r2_score(y_true_train, y_pred_train):.2f}\n')
# Performance on validation set:
y_true_val = y_val
y_pred_val = fs1_linreg_normalized.predict(X_val)
print('On Validation Set:')
print(f'Train MAE: {mean_absolute_error(y_true_val, y_pred_val):.1f}')
mse_val = mean_squared_error(y_true_val, y_pred_val)
print(f'Train MSE: {mse_val:.1f}')
print(f'Train RMSE: {sqrt(mse_val):.1f}')
print(f'Train R^2 score: {r2_score(y_true_val, y_pred_val):.2f}\n')
# + id="7pC9US_6hutP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b873465b-f70c-4ddb-ade2-53d7b61f85bc"
# Check linear regression coefficients to get a sense of feature importances:
fs1_linreg_normalized_coeffs = pd.Series(fs1_linreg_normalized.coef_, X_val.columns)
# Plot coefficients on chart:
n = len(X_val.columns)
plt.figure(figsize=(10, n/2))
fs1_linreg_normalized_coeffs.sort_values().plot.barh(color='darkorange')
plt.title('Relative Importance of Features: Coefficients in LinearRegression Model w/ Normalized Inputs')
plt.show()
# + id="PrjdYIW8uIjU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="640b6d61-a5df-4b75-af68-d348e042156a"
from sklearn.feature_selection import SelectFromModel
# Initiate SelectFromModel feature selector:
feature_selector_coeffs_normalized = SelectFromModel(estimator=fs1_linreg_normalized, threshold='mean')
# Fit feature selector:
feature_selector_coeffs_normalized.fit(X_train, y_train)
# + id="rjX4m2WEuIhQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f5da629c-a6df-4b6c-ceee-2e712c752fcc"
# See which features are supported based on raw coefficients:
feature_names = X_train.columns
pd.DataFrame(feature_selector_coeffs_normalized.get_support(),
index=feature_names,
columns=['include']).sort_values(by='include', ascending=False)
# + id="KylRQVOHuIeC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="655693c1-853a-4bd5-9f67-9261e0ceeb99"
selected_features_coeffs_normalized = feature_names[feature_selector_coeffs_normalized.get_support()].tolist()
selected_features_coeffs_normalized
# + [markdown] id="zXBt4T50uJD0" colab_type="text"
# Coeffs w/ Standardized Feature Matrices:
# + id="Mj4Rj1h3J1ir" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="1931abe5-eb3f-4edf-a653-cac22be1a4ec"
# Cross-check with standardized feature values fed into linear regression model:
# (1) Import model class:
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler
# (2) Initiate model:
fs1_linreg_standardized = LinearRegression()
# (3a) Define features matrix and target vector:
# Already defined above.
# (3b) Standardize our data, so we can better interpret the coefficients
# w.r.t. what they mean about various features' importances:
scaler = StandardScaler()
X_train_standardized = scaler.fit_transform(X_train)
X_val_standardized = scaler.transform(X_val)
# (4) Fit the model to our training data:
fs1_linreg_standardized.fit(X_train_standardized, y_train)
# Performance on training data:
print('Model Performance: LinearRegression w/ Standardized Inputs\n')
y_true_train = y_train
y_pred_train = fs1_linreg_standardized.predict(X_train_standardized)
print('On Training Set:')
print(f'Train MAE: {mean_absolute_error(y_true_train, y_pred_train):.1f}')
mse_train = mean_squared_error(y_true_train, y_pred_train)
print(f'Train MSE: {mse_train:.1f}')
print(f'Train RMSE: {sqrt(mse_train):.1f}')
print(f'Train R^2 score: {r2_score(y_true_train, y_pred_train):.2f}\n')
# Performance on validation set:
y_true_val = y_val
y_pred_val = fs1_linreg_standardized.predict(X_val_standardized)
print('On Validation Set:')
print(f'Train MAE: {mean_absolute_error(y_true_val, y_pred_val):.1f}')
mse_val = mean_squared_error(y_true_val, y_pred_val)
print(f'Train MSE: {mse_val:.1f}')
print(f'Train RMSE: {sqrt(mse_val):.1f}')
print(f'Train R^2 score: {r2_score(y_true_val, y_pred_val):.2f}\n')
# + id="tV4RIr7rJuto" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6a447783-4fb2-4505-a9e8-3092c722b899"
# Check linear regression coefficients to get a sense of feature importances:
fs1_linreg_standardized_coeffs = pd.Series(fs1_linreg_standardized.coef_, X_val.columns)
# Plot coefficients on chart:
n = len(X_val.columns)
plt.figure(figsize=(10, n/2))
fs1_linreg_standardized_coeffs.sort_values().plot.barh(color='darkorange')
plt.title('Relative Importance of Features: Coefficients in LinearRegression Model')
plt.show()
# + id="sEIYFibHvqNk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="0c6c379f-9529-45bc-854e-b5de714fd845"
from sklearn.feature_selection import SelectFromModel
# Initiate SelectFromModel feature selector:
feature_selector_coeffs_standardized = SelectFromModel(estimator=fs1_linreg_standardized, threshold='median')
# Fit feature selector:
feature_selector_coeffs_standardized.fit(X_train, y_train)
# + id="AABBZzG2vqKj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="705fabc3-03b5-4c75-8a4b-670fefdd9bac"
# See which features are supported based on raw coefficients:
feature_names = X_train.columns
pd.DataFrame(feature_selector_coeffs_standardized.get_support(),
index=feature_names,
columns=['include']).sort_values(by='include', ascending=False)
# + id="WyPZuUkQvqH4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="1c66ea75-cc61-47c6-a91b-0c6cd0e3726f"
selected_features_coeffs_standardized = feature_names[feature_selector_coeffs_standardized.get_support()].tolist()
selected_features_coeffs_standardized
# + [markdown] id="Ezr-z58xgbXd" colab_type="text"
# ### Method 2: Recursive Feature Elimination:
# + id="nHVHiY9BLhIH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="67b8318c-e7c6-47c3-b5ce-2afdb119a665"
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import RFECV
from sklearn.model_selection import StratifiedKFold
# Initiate feature selector:
feature_selector_rfe = RFECV(estimator=fs1_linreg, step=1, cv=StratifiedKFold(n_splits=5, random_state=42), scoring="neg_mean_absolute_error") # Alternative scoring options: ‘neg_mean_squared_log_error’, ‘neg_root_mean_squared_error’, ‘neg_mean_squared_error’
# Fit feature selector:
feature_selector_rfe.fit(X_train, y_train)
# Performance on training data:
print('Model Performance: LinearRegression with RFECV for Feature Selection:\n')
y_true_train = y_train
y_pred_train = feature_selector_rfe.predict(X_train)
print('On Training Set:')
print(f'Train MAE: {mean_absolute_error(y_true_train, y_pred_train):.1f}')
mse_train = mean_squared_error(y_true_train, y_pred_train)
print(f'Train MSE: {mse_train:.1f}')
print(f'Train RMSE: {sqrt(mse_train):.1f}')
print(f'Train R^2 score: {r2_score(y_true_train, y_pred_train):.2f}\n')
# Performance on validation set:
y_true_val = y_val
y_pred_val = feature_selector_rfe.predict(X_val)
print('On Validation Set:')
print(f'Train MAE: {mean_absolute_error(y_true_val, y_pred_val):.1f}')
mse_val = mean_squared_error(y_true_val, y_pred_val)
print(f'Train MSE: {mse_val:.1f}')
print(f'Train RMSE: {sqrt(mse_val):.1f}')
print(f'Train R^2 score: {r2_score(y_true_val, y_pred_val):.2f}\n')
# + id="ZSAYexcpnNH_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="2e772348-7e4a-4194-d49d-7f0148edef77"
print("Optimal number of features : %d" % feature_selector_rfe.n_features_)
num_features_baseline = next(x for x, value in enumerate(feature_selector_rfe.grid_scores_) if value > -750)
print(f'Capture bulk of accuracy with: {num_features_baseline}')
# Plot number of features VS. cross-validation scores
plt.figure()
plt.xlabel("Number of features selected")
plt.ylabel("Negative Mean Absolute Error (MAE)")
plt.plot(range(1, len(feature_selector_rfe.grid_scores_) + 1),
feature_selector_rfe.grid_scores_)
plt.show()
# + id="qIKfBLl3oM_o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="e20084a8-f757-4b65-b380-d8f6365d4f32"
ranks_rfecv = pd.DataFrame(data=feature_selector_rfe.ranking_, index=feature_names, columns=['ranking'])
ranks_rfecv['include'] = feature_selector_rfe.support_
ranks_rfecv['MAE_score'] = feature_selector_rfe.grid_scores_
ranks_rfecv.sort_values(by='include', ascending=False)
# + id="6zS8wSi4oM8w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="c75ee99c-f802-4736-870d-1078b8183eb2"
ranks_rfecv.sort_values(by=['MAE_score'], ascending=True)
# + id="d4t6WVcgoM5s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="07ae1e0e-507d-457f-f538-1407b1e2c08e"
feature_names = X_train.columns
selected_features_rfe = feature_names[feature_selector_rfe.support_].tolist()
selected_features_rfe
# + id="mX56ckx8yeBJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="106d6661-4e74-4e17-c87e-9ff41fc82a11"
# Check if RFECV recommended features also include those recommended by the
# model coeffs above --> if so, we will use the RFECV recommended top features:
condition_1 = all(elem in selected_features_rfe for elem in selected_features_coeffs_raw)
condition_2 = all(elem in selected_features_rfe for elem in selected_features_coeffs_normalized)
condition_3 = all(elem in selected_features_rfe for elem in selected_features_coeffs_standardized)
(condition_1 & condition_2 & condition_3)
# + [markdown] id="ibSbmTC11BVG" colab_type="text"
# ### Final Selected Features:
# + id="SEg8Yn64yvyA" colab_type="code" colab={}
# --> OK, let's use the RFECV recommended features then:
features = selected_features_rfe
# + [markdown] id="tq6mQ-Q61oLH" colab_type="text"
# # **Final Model:**
# + id="59IGJ5qr1nzv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="77359167-18ec-4517-a951-652436a00d40"
# (1) Import model class:
from sklearn.linear_model import LinearRegression
# (2) Initiate model:
m_final = LinearRegression()
# (3) Define features matrix and target vector:
# Features and target:
target = 'price'
features = selected_features_rfe # Same as above.
# The resulting new final matrices & vectors:
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
y_test = test[target]
# (4) Fit the model to our training data:
m_final.fit(X_train, y_train)
# Performance on training data:
print('Model Performance:\n')
y_true_train = y_train
y_pred_train = m_final.predict(X_train)
print('On Training Set:')
print(f'Train MAE: {mean_absolute_error(y_true_train, y_pred_train):.1f}')
mse_train = mean_squared_error(y_true_train, y_pred_train)
print(f'Train MSE: {mse_train:.1f}')
print(f'Train RMSE: {sqrt(mse_train):.1f}')
print(f'Train R^2 score: {r2_score(y_true_train, y_pred_train):.2f}\n')
# Performance on validation set:
y_true_val = y_val
y_pred_val = m_final.predict(X_val)
print('On Validation Set:')
print(f'Train MAE: {mean_absolute_error(y_true_val, y_pred_val):.1f}')
mse_val = mean_squared_error(y_true_val, y_pred_val)
print(f'Train MSE: {mse_val:.1f}')
print(f'Train RMSE: {sqrt(mse_val):.1f}')
print(f'Train R^2 score: {r2_score(y_true_val, y_pred_val):.2f}\n')
# + [markdown] id="9tLgNe9P2hm7" colab_type="text"
# ### Final Results: Performance on New Data (Our Holdout Test Set)
# + id="Ghwl5IRoLhDP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="0db4d406-1907-46fe-85a9-ae8f4234b85a"
# Performance on new data: our test set:
print('Final Model Performance on Test (Holdout) Set:\n')
y_true_test = y_test
y_pred_test = m_final.predict(X_test)
print('On Validation Set:')
print(f'Train MAE: {mean_absolute_error(y_true_test, y_pred_test):.2f}')
mse_val = mean_squared_error(y_true_test, y_pred_test)
print(f'Train MSE: {mse_val:.2f}')
print(f'Train RMSE: {sqrt(mse_val):.2f}')
print(f'Train R^2 score: {r2_score(y_true_test, y_pred_test):.2f}\n')
|
module2-regression-2/Assignment_DS_212_Regression_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [Intermediate Machine Learning](https://www.kaggle.com/learn/intermediate-machine-learning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/cross-validation).**
#
# ---
#
# In this exercise, you will leverage what you've learned to tune a machine learning model with **cross-validation**.
#
# # Setup
#
# The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
# Set up code checking
import os
if not os.path.exists("../input/train.csv"):
os.symlink("../input/home-data-for-ml-course/train.csv", "../input/train.csv")
os.symlink("../input/home-data-for-ml-course/test.csv", "../input/test.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.ml_intermediate.ex5 import *
print("Setup Complete")
# You will work with the [Housing Prices Competition for Kaggle Learn Users](https://www.kaggle.com/c/home-data-for-ml-course) from the previous exercise.
#
# 
#
# Run the next code cell without changes to load the training and validation sets in `X_train`, `X_valid`, `y_train`, and `y_valid`. The test set is loaded in `X_test`.
#
# For simplicity, we drop categorical variables.
# +
import pandas as pd
from sklearn.model_selection import train_test_split
# Read the data
train_data = pd.read_csv('../input/train.csv', index_col='Id')
test_data = pd.read_csv('../input/test.csv', index_col='Id')
# Remove rows with missing target, separate target from predictors
train_data.dropna(axis=0, subset=['SalePrice'], inplace=True)
y = train_data.SalePrice
train_data.drop(['SalePrice'], axis=1, inplace=True)
# Select numeric columns only
numeric_cols = [cname for cname in train_data.columns if train_data[cname].dtype in ['int64', 'float64']]
X = train_data[numeric_cols].copy()
X_test = test_data[numeric_cols].copy()
# -
# Use the next code cell to print the first several rows of the data.
X.head()
# So far, you've learned how to build pipelines with scikit-learn. For instance, the pipeline below will use [`SimpleImputer()`](https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html) to replace missing values in the data, before using [`RandomForestRegressor()`](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestRegressor.html) to train a random forest model to make predictions. We set the number of trees in the random forest model with the `n_estimators` parameter, and setting `random_state` ensures reproducibility.
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
my_pipeline = Pipeline(steps=[
('preprocessor', SimpleImputer()),
('model', RandomForestRegressor(n_estimators=50, random_state=0))
])
# -
# You have also learned how to use pipelines in cross-validation. The code below uses the [`cross_val_score()`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html) function to obtain the mean absolute error (MAE), averaged across five different folds. Recall we set the number of folds with the `cv` parameter.
# +
from sklearn.model_selection import cross_val_score
# Multiply by -1 since sklearn calculates *negative* MAE
scores = -1 * cross_val_score(my_pipeline, X, y,
cv=5,
scoring='neg_mean_absolute_error')
print("Average MAE score:", scores.mean())
# -
# # Step 1: Write a useful function
#
# In this exercise, you'll use cross-validation to select parameters for a machine learning model.
#
# Begin by writing a function `get_score()` that reports the average (over three cross-validation folds) MAE of a machine learning pipeline that uses:
# - the data in `X` and `y` to create folds,
# - `SimpleImputer()` (with all parameters left as default) to replace missing values, and
# - `RandomForestRegressor()` (with `random_state=0`) to fit a random forest model.
#
# The `n_estimators` parameter supplied to `get_score()` is used when setting the number of trees in the random forest model.
# +
from sklearn.model_selection import cross_val_score
def get_score(n_estimators):
my_pipeline = Pipeline(steps=[
('preprocessor', SimpleImputer()),
('model', RandomForestRegressor(n_estimators, random_state=0))
])
scores = -1 * cross_val_score(my_pipeline, X, y,
cv=3,
scoring='neg_mean_absolute_error')
return scores.mean()
# Check your answer
step_1.check()
# +
# Lines below will give you a hint or solution code
#step_1.hint()
#step_1.solution()
# -
# # Step 2: Test different parameter values
#
# Now, you will use the function that you defined in Step 1 to evaluate the model performance corresponding to eight different values for the number of trees in the random forest: 50, 100, 150, ..., 300, 350, 400.
#
# Store your results in a Python dictionary `results`, where `results[i]` is the average MAE returned by `get_score(i)`.
# +
results = {}
for i in range(1,9):
results[50*i] = get_score(50*i) # Your code here
# Check your answer
step_2.check()
# +
# Lines below will give you a hint or solution code
#step_2.hint()
#step_2.solution()
# -
# Use the next cell to visualize your results from Step 2. Run the code without changes.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(list(results.keys()), list(results.values()))
plt.show()
# -
# # Step 3: Find the best parameter value
#
# Given the results, which value for `n_estimators` seems best for the random forest model? Use your answer to set the value of `n_estimators_best`.
# +
n_estimators_best = 200
# Check your answer
step_3.check()
# +
# Lines below will give you a hint or solution code
#step_3.hint()
#step_3.solution()
# -
# In this exercise, you have explored one method for choosing appropriate parameters in a machine learning model.
#
# If you'd like to learn more about [hyperparameter optimization](https://en.wikipedia.org/wiki/Hyperparameter_optimization), you're encouraged to start with **grid search**, which is a straightforward method for determining the best _combination_ of parameters for a machine learning model. Thankfully, scikit-learn also contains a built-in function [`GridSearchCV()`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html) that can make your grid search code very efficient!
#
# # Keep going
#
# Continue to learn about **[gradient boosting](https://www.kaggle.com/alexisbcook/xgboost)**, a powerful technique that achieves state-of-the-art results on a variety of datasets.
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161289) to chat with other Learners.*
|
Kaggle Courses/Intermediate Machine Learning/exercise-cross-validation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.model_selection import StratifiedKFold, train_test_split
from lightgbm import LGBMClassifier, Booster
from matplotlib import pyplot as plt
from feature_extraction import get_features, get_features_with_derivative
# +
painting_data = [{'name': 'Rothko', 'colors': ['#f8b335', '#ed6a29', '#f39434', '#fdc03e', '#fa3229']},
{'name': 'Monet', 'colors': ['#848aa7', '#9392a8', '#6f7ca5', '#b2918a', '#9b8d9c']},
{'name': 'Picasso', 'colors': ['#132f3a', '#224f5b', '#93a49c', '#103755', '#436160']},
{'name': 'Cuco', 'colors': ['#be8373', '#9f9646', '#8ca487', '#768cb2', '#568132']},
{'name': 'Bacon', 'colors': ['#562f4c', '#a42238', '#4a181c', '#ba252c', '#7d212c']}]
n_features = (14 + 3) # 3 - autocorr, 14 - main
samplerate = 45
max_len_autocorr = 45
# +
data_files = ['EEG_Data/data01_g.txt',
'EEG_Data/data02_g.txt',
'EEG_Data/data03_g.txt',
'EEG_Data/data04_g.txt',
'EEG_Data/data05_g.txt',
'EEG_Data/data01.txt',
'EEG_Data/data02.txt',
'EEG_Data/data03.txt',
'EEG_Data/data04.txt',
'EEG_Data/data05.txt',
'EEG_Data/data01_d.txt',
'EEG_Data/data02_d.txt',
'EEG_Data/data03_d.txt',
'EEG_Data/data04_d.txt',
'EEG_Data/data05_d.txt',
'EEG_Data/data01_a.txt',
'EEG_Data/data02_a.txt',
'EEG_Data/data03_a.txt',
'EEG_Data/data04_a.txt',
'EEG_Data/data05_a.txt',
'EEG_Data/data01_k.txt',
'EEG_Data/data02_k.txt',
'EEG_Data/data03_k.txt',
'EEG_Data/data04_k.txt',
'EEG_Data/data05_k.txt']
labels = [3, 4, 0, 1, 2, 3, 4, 0, 1, 2, 3, 4, 0, 1, 2, 3, 4, 0, 1, 2, 3, 4, 0, 1, 2]
train_dfs = [pd.read_csv(f, sep=' ', header=None) for f in data_files]
for tr_df in train_dfs:
print(tr_df.shape)
# -
usecols = list(range(5))
n_channels = len(usecols)
def get_samples_from_pd(pd, arr_len=256, step=64, n_samples=12):
output = np.zeros((n_samples, n_features * n_channels))
for i in range(n_samples):
for j in range(n_channels):
feat_list = get_features(pd[usecols].values[i * step: i * step+arr_len, j])
output[i, j*n_features:(j+1)*n_features] = feat_list
return output
use_samples = 50
Y_full = np.zeros(len(labels) * use_samples)
for i in range(len(labels)):
Y_full[i*use_samples:(i+1) * use_samples] = labels[i]
# +
train_data = np.zeros((use_samples * len(labels), n_features * n_channels))
for i in range(len(labels)):
train_data[i*use_samples:(i+1) * use_samples,:] = get_samples_from_pd(train_dfs[i],
arr_len=samplerate,
step=5, n_samples=use_samples)
# -
train_data = np.nan_to_num(train_data)
X_train, X_test, y_train, y_test = train_test_split(train_data, Y_full, test_size=0.2)
lgb = LGBMClassifier(num_leaves=10, min_data=5, min_data_in_bin=5, learning_rate=0.01, n_estimators=2000)
lgb.fit(X_train, y_train, eval_set=[(X_test, y_test)], early_stopping_rounds=100)
lgb.booster_.save_model('lightGBM_sr_45_1sec_all5_v2.txt')
|
training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Useful for debugging
# %load_ext autoreload
# %autoreload 2
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams['figure.figsize'] = (13,8)
# #%matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# # APEX Gun example
# +
from gpt import GPT
from distgen import Generator
import os
GPT_IN = 'templates/apex_gun/gpt.in'
DISTGEN_IN = 'templates/apex_gun/distgen.yaml'
# +
gen = Generator(DISTGEN_IN)
gen['n_particle'] = 1000
gen.run()
P0 = gen.particles
factor = 2
#P0.x *= factor
#P0.y *= 1/factor
P0.plot('x', 'y')
# +
from gpt import run_gpt_with_distgen
settings = {'n_particle':100,
'gun_peak_field':20e6,
'gun_relative_phase':0,
'BSOL':0.075,
'tmax': 5e-9,
'RadiusMax':.015,
'Ntout':2000,
'dtmin':0,
'GBacc':6.5,
'xacc':6.5,
'space_charge':1}
G = run_gpt_with_distgen(settings,
gpt_input_file=GPT_IN,
distgen_input_file=DISTGEN_IN,
auto_phase=True,
verbose=True)
# -
G.plot('sigma_x')
G.plot('mean_kinetic_energy')
G.particles[-1]
G.plot()
# # Plot trajectories
G.particles[0]._settable_array_keys
# +
import numpy as np
from matplotlib import pyplot as plt
# Make trajectory structure here for now, should go somewhere else as a function
rs ={}
for t in G.particles:
for ID in t['id']:
idint=int(ID)
res = np.where(t['id']==ID)
index = res[0][0]
if(ID not in rs.keys()):
rs[idint]={'x':[],'y':[],'z':[], 't':[], 'GBz':[]}
else:
rs[idint]['x'].append(t['x'][index])
rs[idint]['y'].append(t['y'][index])
rs[idint]['z'].append(t['z'][index])
rs[idint]['t'].append(t['t'][index])
# rs[idint]['GBz'].append(t['GBz'][index])
# +
for ind in rs.keys():
for var in rs[ind]:
rs[ind][var]=np.array(rs[ind][var])
for ind in rs.keys():
plt.plot(rs[ind]['z'][0],rs[ind]['x'][0]*1e2, color='red', marker='o')
plt.plot(rs[ind]['z'],rs[ind]['x']*1e2, color='black', alpha=0.1)
plt.ylim(-1.5, 1.5)
plt.xlim(0, 0.1)
plt.title('GPT tracking')
plt.xlabel('z (m)');
plt.ylabel('x (cm)');
# -
zlist = np.array([P['mean_z'] for P in G.particles])
np.argmin(abs(zlist - 0.15))
G.particles[3]['mean_z']
# +
#G.particles[3].write('$HOME/Scratch/gpt_apex_100pC_4x.h5')
# -
G.archive('gpt_apex_gun.h5')
G2 = GPT()
G2.load_archive('gpt_apex_gun.h5')
G2.particles[3]['mean_z']
G.tout
plt.plot(np.array([P['n_particle'] for P in G.particles]))
|
examples/apex_gun_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda3-pyvizenv] *
# language: python
# name: conda-env-Anaconda3-pyvizenv-py
# ---
# # BTC Prediction
# +
# Initial imports
import os
import requests
import pandas as pd
from dotenv import load_dotenv
import alpaca_trade_api as tradeapi
from MCForecastTools import MCSimulation
import json
from datetime import date
from pandas import json_normalize
# %matplotlib inline
# -
# ## Part 1 - Import Indicator data from Topfolio
# +
# Define Variables
# Format current date as ISO format
#start_date = (pd.Timestamp.now() - pd.Timedelta(3, unit='d'))
#end_date = pd.Timestamp.now()
# Define Parameters
exchange = 'Binance'
symbol = 'BTCUSDT'
indicator_type = 'orderbook'
name = '0-1%'
interval = '3600'
# Create parameterized url
base_url = "https://api.topfol.io/indicators/candle?"
parameters_url = "startDate=2021-03-02&endDate=2021-04-02&exchange="+exchange+"&symbol="+symbol+"&indicator_type="+indicator_type+"&name="+name+"&interval="+interval
request_url = base_url + parameters_url
# Submit request and format output
response_data = requests.get(request_url)
pages = int(response_data.headers['page-amount'])
print (f'Number of Pages is {pages}')
new_results = True
page = 1
df_btcusdt_data = pd.DataFrame()
topfolio_api = requests.get(request_url).json()
data = topfolio_api
for page in range(2, pages+1):
topfolio_api = requests.get(request_url + f"&page={page}").json()
data.extend(topfolio_api)
page += 1
df_btcusdt_data = json_normalize(data)
# -
df_btcusdt_data.head()
# +
#df_btcusdt_dataeeded fields
#df_btcusdt_data = df_btcusdt_data.drop(['open_price','high_price','low_price'],axis=1)
# Convert Unix Time to iso format
df_btcusdt_data['timestamp']=(pd.to_datetime(df_btcusdt_data['timestamp'],unit='s'))
# Set index to timestamp
df_btcusdt_data = df_btcusdt_data.set_index(['timestamp'])
# Sort Data
df_btcusdt_data = df_btcusdt_data.sort_index()
# Rename Column
df_btcusdt_data.columns = ['open','high','low','close']
df_btcusdt_data.columns = pd.MultiIndex.from_product([["BTCUSDT"], df_btcusdt_data.columns])
# -
df_btcusdt_data.count()
# ## Part 2 - Forecasting
#
# ### Monte Carlo Simulation
# +
# Configuring a Monte Carlo simulation to forecast 30 years cumulative returns
#Set number of simulations
num_sims = 500
# Configure a Monte Carlo simulation to forecast
MC_week = MCSimulation(
portfolio_data = df_btcusdt_data,
num_simulation = num_sims,
num_trading_days = 90
)
# -
# Running a Monte Carlo simulation to forecast a week cumulative returns
MC_week.calc_cumulative_return()
# +
# Plot simulation outcomes
line_plot = MC_week.plot_simulation()
# Save the plot for future usage
line_plot.get_figure().savefig("MC_seven_day_sim_plot.png", bbox_inches="tight")
# +
# Plot probability distribution and confidence intervals
dist_plot = MC_week.plot_distribution()
# Save the plot for future usage
dist_plot.get_figure().savefig('MC_seven_day_dist_plot.png',bbox_inches='tight')
# -
# ### Retirement Analysis
# +
# Fetch summary statistics from the Monte Carlo simulation results
tbl = MC_week.summarize_cumulative_return()
# Print summary statistics
print(tbl)
# -
# ### Calculate the expected Bitcoin range.
# +
# Crypto API URLs
btc_url = "https://api.alternative.me/v2/ticker/Bitcoin/?convert=USD"
eth_url = "https://api.alternative.me/v2/ticker/Ethereum/?convert=USD"
# Fetch current BTC price
btc_response = requests.get(btc_url)
btc_data = btc_response.json()
btc_price = btc_data['data']['1']['quotes']['USD']['price']
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $20,000
ci_lower = round(tbl[8]*btc_price,2)
ci_upper = round(tbl[9]*btc_price,2)
diff_lower = round(btc_price - ci_lower,2)
diff_upper = round(ci_upper - btc_price,2)
# Print results
print(f"Bitcoin current price is ${btc_price}")
print(f"There is a 95% chance that it will be within the following range ${ci_lower} and ${ci_upper} over the next 90 days")
print(f"Distance from upper is ${diff_upper}")
print(f"Distance from lower is ${diff_lower}")
# -
|
notebooks/billy/btc-montecarlo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="UueoL5L1qw2n"
import pandas as pd
import numpy as np
from sklearn.utils import shuffle
from sklearn.model_selection import KFold
import lightgbm as lgb
from lightgbm import LGBMRegressor
import lightgbm
from contextlib import contextmanager
import time
import gc
import random
seed = 10
random.seed(seed)
np.random.seed(seed)
# + colab={"base_uri": "https://localhost:8080/", "height": 290} colab_type="code" id="S6ux6DWjrETi" outputId="3b8c776e-92ca-4a15-99f9-fc8f47a72a0e"
def read_df():
df_train = pd.read_csv('../input/Train.csv')
print("Train shape: {}".format(df_train.shape))
return df_train
df_train = read_df()
# + colab={} colab_type="code" id="6L1qdRLlb-ng"
def train_data_cleaning(df):
outlayers = [(4384, 4770), (19469, 19739)]
columns = ['Air temperature (C)', 'Air humidity (%)', 'Pressure (KPa)',
'Wind speed (Km/h)', 'Wind gust (Km/h)', 'Wind direction (Deg)']
for c in columns:
for start, end in outlayers:
df[c][start: end] = np.nan
for i in range(start, end):
if np.isnan(df.iloc[i + 288][c]):
df[c][i] = (11 * df.iloc[i + 2 * 288][c] + 8 * df.iloc[i - 288][c] + 7 * df.iloc[i - 2 * 288][c]) / 26
else:
df[c][i] = (df.iloc[i + 288][c] + df.iloc[i + 2 * 288][c] + df.iloc[i - 288][c] + df.iloc[i - 2 * 288][c]) / 4
return df
# -
def test_data_forecasting(df):
columns = ['Air temperature (C)', 'Air humidity (%)', 'Pressure (KPa)',
'Wind speed (Km/h)', 'Wind gust (Km/h)', 'Wind direction (Deg)',
]
for (start_v, end_v, end_n) in [(0, 8914, 10067), (10067, 16083, 17236), (17236, 26301, len(df))]:
for c in columns:
for p in range(end_v, end_n, 2 * 288):
pred = df[c].iloc[p - 2 *288: p]
df[c][p: min(p + 2 * 288, end_n)] = pred[0: min(2 * 288, end_n - p)]
return df
# + colab={"base_uri": "https://localhost:8080/", "height": 104} colab_type="code" id="6cV7_lFlvHtb" outputId="2835e018-6b07-4bc2-8cb5-2eba07ae84a5"
def feature_engineering(df):
# environment features
df['D Air temperature (C)'] = df['Air temperature (C)'] - df['Air temperature (C)'].shift(1)
df['D Pressure (KPa)'] = df['Pressure (KPa)'] - df['Pressure (KPa)'].shift(1)
# control features
df['M Irrigation field'] = df['Irrigation field'] * df['Irrigation field'].rolling(window=24).sum()
df['D Air temperature (C)'] = df['Irrigation field'] * df['D Air temperature (C)']
df['D Pressure (KPa)'] = df['Irrigation field'] * df['D Pressure (KPa)']
# state features
# target
df['Velocity'] = df['Soil humidity'] - df['Soil humidity'].shift(1)
return df
# + colab={} colab_type="code" id="ktDKPQER522D"
input_columns = ['Irrigation field', 'M Irrigation field',
'D Pressure (KPa)', 'D Air temperature (C)',
]
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="aGRNnv0AzB2O" outputId="9dbecb42-39f4-4dfb-9e5a-0b79e6a66175"
def train(train_df, input_columns, target_column):
train_df = train_df[train_df[target_column].notnull()].copy()
folds = KFold(n_splits=5, shuffle=True, random_state=123)
regs = []
train_x = train_df[input_columns]
train_y = train_df[target_column]
for n_fold, (trn_idx, val_idx) in enumerate(folds.split(train_x, train_y)):
trn_x, trn_y = train_x.iloc[trn_idx], train_y.iloc[trn_idx]
val_x, val_y = train_x.iloc[val_idx], train_y.iloc[val_idx]
reg = LGBMRegressor(
n_estimators=2000,
learning_rate=0.5,
num_leaves=123,
colsample_bytree=.8,
subsample=.9,
max_depth=15,
reg_alpha=.1,
reg_lambda=.1,
min_split_gain=.01,
min_child_weight=2
)
reg.fit(trn_x, trn_y,
eval_set= [(trn_x, trn_y), (val_x, val_y)], verbose=250, early_stopping_rounds=150,
)
#lightgbm.plot_importance(reg, height=1.0, max_num_features=10)
regs.append(reg)
del reg, trn_x, trn_y, val_x, val_y
gc.collect()
return regs
# -
def predict(regs, test_df, first_state, field_id):
test_df = test_df.rename(columns={'Soil humidity': 'Values'})
indices = [-1] + list(test_df[test_df['Values'].notnull()]['Values'].index)
test_df['Velocity'] = np.mean([reg.predict(test_df[input_columns]) for reg in regs ], axis=0)
def forward_process(start, end):
preds = []
pred = first_state if start == -1 else test_df['Values'].iloc[start]
for j in range(start + 1, end):
pred += test_df['Velocity'].iloc[j]
preds.append(pred.copy())
return preds
def backward_process(start, end):
preds = []
pred = test_df['Values'].iloc[end]
for j in range(end, start + 1, -1):
pred -= test_df['Velocity'].iloc[j]
preds.append(pred.copy())
return preds[::-1]
for start, end in zip(indices[:-1], indices[1:]):
f_preds = forward_process(start, end)
b_preds = backward_process(start, end)
j = start + 1
for i in range(start + 1, end):
if test_df['Irrigation field'].iloc[i] == 0:
test_df['Values'].iloc[i] = f_preds[i - start - 1]
j = i
else:
test_df['Values'].iloc[i] = (1 - (i - j)/(end - j - 1)) * f_preds[i - start - 1] +\
((i - j) /(end - j - 1)) * b_preds[i - start - 1]
f_preds = forward_process(indices[-1], len(test_df))
if len(f_preds) > 0:
test_df['Values'][indices[-1] + 1: len(test_df)] = f_preds
test_df['ID'] = test_df['timestamp'] + ' x Soil humidity ' + str(field_id + 1)
return test_df[['ID', 'Values']]
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="s-9Xhut8yvvG" outputId="952a0b50-7ebe-4cd4-cefa-eff3f06b9cb4"
@contextmanager
def timer(title):
t0 = time.time()
yield
print("{} - done in {:.0f}s".format(title, time.time() - t0))
with timer("Importing Datasets: "):
df_train = read_df()
gc.collect();
with timer("Time Series Imputation: "):
df_train = train_data_cleaning(df_train)
df_train = test_data_forecasting(df_train)
gc.collect();
env_columns = ['Air temperature (C)', 'Air humidity (%)', 'Pressure (KPa)',
'Wind speed (Km/h)', 'Wind gust (Km/h)', 'Wind direction (Deg)',]
df_train_1 = df_train[['timestamp', 'Soil humidity 1', 'Irrigation field 1', *env_columns]]
df_train_1 = df_train_1.rename(columns={
'Soil humidity 1': 'Soil humidity', 'Irrigation field 1': 'Irrigation field'})
df_train_2 = df_train[['timestamp', 'Soil humidity 2', 'Irrigation field 2', *env_columns]]
df_train_2 = df_train_2.rename(columns={
'Soil humidity 2': 'Soil humidity', 'Irrigation field 2': 'Irrigation field'})
df_train_3 = df_train[['timestamp', 'Soil humidity 3', 'Irrigation field 3', *env_columns]]
df_train_3 = df_train_3.rename(columns={
'Soil humidity 3': 'Soil humidity', 'Irrigation field 3': 'Irrigation field'})
df_train_4 = df_train[['timestamp', 'Soil humidity 4', 'Irrigation field 4', *env_columns]]
df_train_4 = df_train_4.rename(columns={
'Soil humidity 4': 'Soil humidity', 'Irrigation field 4': 'Irrigation field'})
limits = [(8914, 10067), (26301,28048), (16083, 17236), (26301,28030)]
preds = []
for i, (df_train_i, (start, end)) in enumerate(zip([df_train_1, df_train_2, df_train_3, df_train_4], limits)):
with timer("Feature Engineering: "):
df_train_i = feature_engineering(df_train_i)
df_train_i = df_train_i.set_index('timestamp')
df_train_i, df_test_i = df_train_i.iloc[:start], df_train_i.iloc[start: end]
df_test_i = df_test_i.reset_index()
df_test_i['Irrigation field'] = df_test_i['Irrigation field'].fillna(value=0)
gc.collect();
with timer("Training"):
regs = train(df_train_i, input_columns, 'Velocity')
gc.collect()
with timer("Testing"):
first_state = df_train_i['Soil humidity'].iloc[start - 1]
prediction = predict(regs, df_test_i, first_state, i)
preds.append(prediction)
gc.collect()
preds = pd.concat(preds, ignore_index=True)
preds.to_csv("submission.csv", index= False)
|
Wazihub Soil Moisture Prediction Challenge/Solution 3/wazihub_soil_moisture.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# _Disclaimer: This code is from a tutorial taken from [<NAME>](https://towardsdatascience.com/in-12-minutes-stocks-analysis-with-pandas-and-scikit-learn-a8d8a7b50ee7)._
import pandas as pd
import datetime
import pandas_datareader.data as web
from pandas import Series, DataFrame
start = datetime.datetime(year=2010, month=1, day=1)
end = datetime.datetime(2017, 1, 11)
df = web.DataReader("AAPL", 'yahoo', start, end)
df.tail()
# ### Exploring Rolling Mean and Return Rate of Stocks
close_px = df['Adj Close']
mavg = close_px.rolling(window=100).mean()
# %matplotlib inline
import matplotlib.pyplot as plt
from matplotlib import style
import matplotlib as mpl
mpl.rc('figure', figsize=(8, 7))
mpl.__version__
# +
style.use('ggplot')
close_px.plot(label='AAPL')
mavg.plot(label='mavg')
plt.legend()
# -
# #### Return Deviation - determine risk & return
rets = close_px / close_px.shift(1) - 1
rets.plot(label='return')
# ### Analysing competitor's stocks
dfcomp = web.DataReader(['AAPL', 'GE', 'GOOG', 'IBM', 'MSFT'], 'yahoo', start=start,
end=end)['Adj Close']
dfcomp.tail()
# #### Correlation analysis - Does one competitor affect another?
retscomp = dfcomp.pct_change()
corr = retscomp.corr()
plt.scatter(retscomp.AAPL, retscomp.GE)
plt.xlabel('Returns AAPL')
plt.ylabel('Returns GE')
# Plot of AAPL and GE return distribution. Obvs: Higher AAPL return results in higher GE returns.
pd.plotting.scatter_matrix(retscomp, diagonal='kde', figsize=(10, 10))
# Using heatmaps, we can visualise correlation ranges among competing stocks. Lighter color means higher correlation between stocks.
plt.imshow(corr, cmap='hot', interpolation='none')
plt.colorbar()
plt.xticks( range(len(corr)), corr.columns )
plt.yticks( range(len(corr)), corr.columns )
# ### Stocks returns rate & risk
plt.scatter(retscomp.mean(), retscomp.std())
plt.xlabel('Expected returns')
plt.ylabel('Risk')
for label, x, y in zip(retscomp.columns, retscomp.mean(), retscomp.std()):
plt.annotate(
label,
xy = (x, y), xytext = (20, -20),
textcoords = 'offset points', ha = 'right', va = 'bottom',
bbox = dict(boxstyle = 'round,pad=0.5', fc = 'yellow', alpha = 0.5),
arrowprops = dict(arrowstyle = '->', connectionstyle = 'arc3,rad=0'))
# ### Stock price prediction
dfreg = df.loc[:, ['Adj Close', 'Volume']]
dfreg['HL_PCT'] = (df['High'] - df['Low']) / df['Close'] * 100.0
dfreg['PCT_change'] = (df['Close'] - df['Open']) / df['Open'] * 100.0
import math
dfreg.fillna(value=-99999, inplace=True)
forecast_out = int(math.ceil(0.01 * len(dfreg)))
import numpy as np
forecast_col = 'Adj Close'
dfreg['label'] = dfreg[forecast_col].shift(-forecast_out)
X = np.array(dfreg.drop(['label'], 1))
import sklearn.preprocessing as preprocessing
X = preprocessing.scale(X)
X_lately = X[-forecast_out:]
X = X[:-forecast_out]
y = np.array(dfreg['label'])
y = y[:-forecast_out]
# ### Model generation
# +
from sklearn.linear_model import LinearRegression
from sklearn.neighbors import KNeighborsRegressor
from sklearn.linear_model import Ridge
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
# -
def split_vals(a, n): return a[:n].copy(), a[n:].copy()
# +
n_valid = 525
n_trn = len(X)-n_valid
X_train, X_valid = split_vals(X, n_trn)
y_train, y_valid = split_vals(y, n_trn)
X_train.shape, y_train.shape, X_valid.shape
# -
clfreg = LinearRegression(n_jobs=-1)
clfreg.fit(X_train, y_train)
clfpoly2 = make_pipeline(PolynomialFeatures(2), Ridge())
clfpoly2.fit(X_train, y_train)
clfpoly3 = make_pipeline(PolynomialFeatures(3), Ridge())
clfpoly3.fit(X_train, y_train)
clfknn = KNeighborsRegressor(n_neighbors=2)
clfknn.fit(X_train, y_train)
confidencereg = clfreg.score(X_valid, y_valid)
confidencepoly2 = clfpoly2.score(X_valid, y_valid)
confidencepoly3 = clfpoly3.score(X_valid, y_valid)
confidenceknn = clfknn.score(X_valid, y_valid)
[confidencereg, confidencepoly2, confidencepoly3, confidenceknn]
forecast_set = clfpoly3.predict(X_lately)
dfreg['Forecast'] = np.nan
forecast_set
# +
last_date = dfreg.iloc[-1].name
last_unix = last_date
next_unix = last_unix + datetime.timedelta(days=1)
for i in forecast_set:
next_date = next_unix
next_unix += datetime.timedelta(days=1)
dfreg.loc[next_date] = [np.nan for _ in range(len(dfreg.columns) -1)] + [i]
dfreg['Adj Close'].tail(500).plot()
dfreg['Forecast'].tail(500).plot()
plt.legend(loc=4)
plt.xlabel('Date')
plt.ylabel('Price')
plt.show()
# -
|
finance/Apple stocks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id="top"></a>
# # Landsat Vegetation Phenology
#
# <hr>
#
# *Notebook compatible with DE Africa Collection 1 Sandbox
#
# # Notebook Summary
#
# This notebook calculates vegetation phenology changes using Landsat 7 or Landsat 8 data. To detect changes in plant life for Landsat, the algorithm uses either the Normalized Difference Vegetation Index (NDVI) or the Enhanced Vegetation Index (EVI), which are common proxies for vegetation growth and health. The outputs of this notebook can be used to assess differences in agriculture fields over time or space and also allow the assessment of growing states such as planting and harvesting.
# <br>
# There are two output products. The first output product is a time series boxplot of NDVI or EVI with the data potentially binned by week, month, week of year, or month of year. The second output product is a time series lineplot of the mean NDVI or EVI for each year, with the data potentially binned by week or month. This product is useful for comparing years to each other.
# <br><br>
# See this website for more information: https://phenology.cr.usgs.gov/ndvi_foundation.php
#
# <hr>
#
# # Index
#
# * [Import Dependencies and Connect to the Data Cube](#import)
# * [Choose Platforms and Products](#plat_prod)
# * [Define the Extents of the Analysis](#define_extents)
# * [Load Data from the Data Cube and Obtain the Vegetation Proxy](#load_data)
# * [Create Phenology Products](#phenology_products)
# * [Plot the Vegetation Index Over Time in a Box-and-Whisker Plot](#phenology_plot_1)
# * [Plot the Vegetation Index Over Time for Each Year](#phenology_plot_2)
# * [Export Curve Fits to a CSV File](#export)
# * [Show TIMESAT Stats](#timesat)
# ## <span id="import">Import Dependencies and Connect to the Data Cube [▴](#top)</span>
# +
import datacube
import sys
import os
# Supress Warnings.
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import xarray as xr
import pandas as pd
import matplotlib.pyplot as plt
import datetime as dt
from odc.ui import DcViewer
from datacube.helpers import write_geotiff
#import DE Africa script
sys.path.append('../../Scripts')
from deafrica_plotting import display_map
from deafrica_plotting import rgb
#import DCAL utility scripts
sys.path.append('../DCAL_utils')
from clean_mask import landsat_qa_clean_mask, landsat_clean_mask_invalid
from sort import xarray_sortby_coord
from vegetation import NDVI, EVI
from plotter_utils import xarray_time_series_plot
# -
# ### Connect to the datacube
dc = datacube.Datacube(app="DCAL Vegitation Phenology")
# ## View Extent of Avalible Data
# +
# Get available products
products_info = dc.list_products()
# List Landsat 8 products
print("Landsat 8 Products:")
products_info[["platform", "name"]]
# -
DcViewer(dc=dc,
products = ['ls8_usgs_sr_scene'],
time='2017',
center=(0.565, 38.007),
zoom=4)
# ## <span id="define_extents">Define the Extents of the Analysis [▴](#top)</span>
# <p style="color:red";><b>CHANGE INPUTS BELOW
# +
# Select an analysis region (Lat-Lon) within the extents listed above.
# Select a time period (Min-Max) within the extents listed above (Year-Month-Day)
# Tanzania Grassland / Cropland
# lat = (-4.5074, -4.4860) # North of Swaga Game Reserve
# lon = (35.1349, 35.1735) # North of Swaga Game Reserve
# Tanzania Grassland / Cropland
# lat = (-8.1541, -8.1272) # Southern Cropland
# lon = (33.2016, 33.2545) # Southern Cropland
# Aviv Coffee Farm, Tanzania (small)
# lat = (-10.6999, -10.6959)
# lon = (35.2608, 35.2662)
# Aviv Coffee Farm, Tanzania (surrounding)
# lat = (-10.855, -10.560)
# lon = (35.130, 35.400)
# Soybean Fields in Western Kenya (from Kizito)
# lat = (-0.801180, -0.483689) # entire region
# lon = (34.193792, 34.546329) # entire region
# Ghana
latitude = (5.5813, 5.6004)
longitude = (-0.5398, -0.5203)
# Time Period
start_date, end_date = dt.datetime(2013,1,1), dt.datetime(2018,12,31)
time_extents = (start_date, end_date)
# -
# **Visualize the selected area**
display_map(longitude,latitude)
# ## <span id="load_data">Load Data from the Data Cube and Obtain the Vegetation Proxy [▴](#top)</span>
# ## Select Vegetation Proxy
# Change which line is commented out in order to switch vegetation proxy. NDVI is the recomended Vegetation Proxy
veg_proxy = 'NDVI'
# once proxy is selected load data and mask out cloud
# +
measurements = []
if veg_proxy == 'NDVI':
measurements = ['red', 'nir', 'pixel_qa']
elif veg_proxy == 'EVI':
measurements = ['red', 'blue', 'nir', 'pixel_qa']
landsat_dataset = dc.load(product = 'ls8_usgs_sr_scene',
measurements = measurements,
y = latitude,
x = longitude,
time = time_extents,
output_crs='EPSG:6933',
resolution=(-30,30))
#load cloud mask and apply to dataset
cloud_mask = landsat_qa_clean_mask(landsat_dataset, platform='LANDSAT_8')
dataset = landsat_dataset.where(cloud_mask)
#view masked dataset
dataset
# +
#change Coordinate names to be compatible with modules used latter in this notebook
dataset = dataset.rename(name_dict={'x':'longitude','y':'latitude'})
dataset
# +
#Generate CHosen Vegitation Proxy
if veg_proxy == 'NDVI':
dataset[veg_proxy] = NDVI(dataset)
if veg_proxy == 'EVI':
dataset[veg_proxy] = EVI(dataset)
dataset
# -
# ## <span id="phenology_products">Create Phenology Products[▴](#top)</span>
#
# If no plots appear in the figures below, there is no data available for the region selected.
# ### <span id="phenology_plot_1">Plot the Vegetation Index Over Time in a Box-and-Whisker Plot[▴](#top)</span>
# <p style="color:red";><b>CHANGE INPUTS BELOW
# +
# Specify whether to plot a curve fit of the vegetation index along time. Input can be either TRUE or FALSE
plot_curve_fit = True
assert isinstance(plot_curve_fit, bool), "The variable 'plot_curve_fit' must be "\
"either True or False."
# Specify the target aggregation type of the curve fit. Input can be either 'mean' or 'median'.
curve_fit_target = 'median'
assert curve_fit_target in ['mean', 'median'], "The variable 'curve_fit_target' must be either "\
"'mean' or 'median'."
# The maximum number of data points that appear along time in each plot.
# If more than this number of data points need to be plotted, a grid of plots will be created.
max_times_per_plot = 40
# -
# <p style="color:red";><b>CHANGE INPUTS BELOW
# +
# Select the binning approach for the vegetation index. Set the 'bin_by' parameter.
# None = do not bin the data
# 'week' = bin the data by week with an extended time axis
# 'month' = bin the data by month with an extended time axis
# 'weekofyear' = bin the data by week and years using a single year time axis
# 'monthofyear' = bin the data by month and years using a single year time axis
# It is also possible to change some of the plotting features using the code below.
bin_by = 'monthofyear'
assert bin_by in [None, 'week', 'month', 'weekofyear', 'monthofyear'], \
"The variable 'bin_by' can only have one of these values: "\
"[None, 'week', 'month', 'weekofyear', 'monthofyear']"
aggregated_by_str = None
if bin_by is None:
plotting_data = dataset
elif bin_by == 'week':
plotting_data = dataset.resample(time='1w').mean()
aggregated_by_str = 'Week'
elif bin_by == 'month':
plotting_data = dataset.resample(time='1m').mean()
aggregated_by_str = 'Month'
elif bin_by == 'weekofyear':
plotting_data = dataset.groupby('time.week').mean(dim=('time'))
aggregated_by_str = 'Week of Year'
elif bin_by == 'monthofyear':
plotting_data = dataset.groupby('time.month').mean(dim=('time'))
aggregated_by_str = 'Month of Year'
params = dict(dataset=plotting_data, plot_descs={veg_proxy:{'none':[
{'box':{'boxprops':{'facecolor':'forestgreen'}}}]}})
if plot_curve_fit:
params['plot_descs'][veg_proxy][curve_fit_target] = [{'gaussian_filter':{}}]
xarray_time_series_plot(**params, fig_params=dict(figsize=(12,8), dpi=150),
max_times_per_plot=max_times_per_plot)
plt.title('Box-and-Whisker Plot of {0} with a Curvefit of Median {0}'.format(veg_proxy))
plt.show()
# -
# ### <span id="phenology_plot_2">Plot the Vegetation Index Over Time for Each Year[▴](#top)</span>
# Note that the curve fits here do not show where some times have no data (encoded as NaNs), as is shown in the box-and-whisker plot. Notably, the curve fits interpolate over times with missing data that are not the first or last time (e.g. January or December for monthly binned data).
# <p style="color:red";><b>CHANGE INPUTS BELOW
# +
years_with_data = []
plot_descs = {}
daysofyear_per_year = {}
plotting_data_years = {}
time_dim_name = None
for year in range(start_date.year, end_date.year+1):
year_data = dataset.sel(time=slice('{}-01-01'.format(year), '{}-12-31'.format(year)))[veg_proxy]
if len(year_data['time']) == 0: # There is nothing to plot for this year.
print("Year {} has no data, so will not be plotted.".format(year))
continue
years_with_data.append(year)
spec_ind_dayofyear = year_data.groupby('time.dayofyear').mean()
daysofyear_per_year[year] = spec_ind_dayofyear.where(~spec_ind_dayofyear.isnull()).dayofyear
# Select the binning approach for the vegetation index. Set the 'bin_by' parameter.
# 'weekofyear' = bin the data by week and years using a single year time axis
# 'monthofyear' = bin the data by month and years using a single year time axis
bin_by = 'monthofyear'
assert bin_by in ['weekofyear', 'monthofyear'], \
"The variable 'bin_by' can only have one of these values: "\
"['weekofyear', 'monthofyear']"
aggregated_by_str = None
if bin_by == 'weekofyear':
plotting_data_year = year_data.groupby('time.week').mean(dim=('time'))
time_dim_name = 'week'
elif bin_by == 'monthofyear':
plotting_data_year = year_data.groupby('time.month').mean(dim=('time'))
time_dim_name = 'month'
plotting_data_years[year] = plotting_data_year
num_time_pts = len(plotting_data_year[time_dim_name])
# Select the curve-fit type.
# See the documentation for `xarray_time_series_plot()` regarding the `plot_descs` parameter.
plot_descs[year] = {'mean':[{'gaussian_filter':{}}]}
time_dim_name = 'week' if bin_by == 'weekofyear' else 'month' if bin_by == 'monthofyear' else 'time'
num_times = 54 if bin_by == 'weekofyear' else 12
time_coords_arr = np.arange(1, num_times+1) # In xarray, week and month indices start at 1.
time_coords_da = xr.DataArray(time_coords_arr, coords={time_dim_name:time_coords_arr},
dims=[time_dim_name], name=time_dim_name)
coords = dict(list(plotting_data_years.values())[0].coords)
coords[time_dim_name] = time_coords_da
plotting_data = xr.Dataset(plotting_data_years, coords=coords)
params = dict(dataset=plotting_data, plot_descs=plot_descs)
fig, curve_fit_plotting_data = \
xarray_time_series_plot(**params, fig_params=dict(figsize=(8,4), dpi=150))
plt.title('Line Plot of {0} for Each Year'.format(veg_proxy))
plt.show()
# -
# ### <span id="export">Export Curve Fits to a CSV File [▴](#top)</span>
# +
# Convert the data to a `pandas.DataFrame`.
dataarrays = []
for (year, _, _), dataarray in curve_fit_plotting_data.items():
dataarrays.append(dataarray.rename(year))
curve_fit_df = xr.merge(dataarrays).to_dataframe()
# Convert the month floats to day ints and average by day (scale to [0,1], multiply by 364, add 1).
curve_fit_df.index.values[:] = (364/11) * (curve_fit_df.index.values - 1) + 1
curve_fit_df.index = curve_fit_df.index.astype(int)
curve_fit_df.index.name = 'day of year'
curve_fit_df = curve_fit_df.groupby('day of year').mean()
# Export the data to a CSV.
csv_output_dir = 'output/CSVs/'
if not os.path.exists(csv_output_dir):
os.makedirs(csv_output_dir)
curve_fit_df.to_csv(csv_output_dir + 'vegetation_phenology_yearly_curve_fits_landsat.csv')
# -
# ### <span id="timesat">Show [TIMESAT](http://web.nateko.lu.se/timesat/timesat.asp) Stats [▴](#top)</span>
def TIMESAT_stats(dataarray, time_dim='time'):
"""
For a 1D array of values for a vegetation index - for which higher values tend to
indicate more vegetation - determine several statistics:
1. Beginning of Season (BOS): The time index of the beginning of the growing season.
(The downward inflection point before the maximum vegetation index value)
2. End of Season (EOS): The time index of the end of the growing season.
(The upward inflection point after the maximum vegetation index value)
3. Middle of Season (MOS): The time index of the maximum vegetation index value.
4. Length of Season (EOS-BOS): The time length of the season (index difference).
5. Base Value (BASE): The minimum vegetation index value.
6. Max Value (MAX): The maximum vegetation index value (the value at MOS).
7. Amplitude (AMP): The difference between BASE and MAX.
Parameters
----------
dataarray: xarray.DataArray
The 1D array of non-NaN values to determine the statistics for.
time_dim: string
The name of the time dimension in `dataarray`.
Returns
-------
stats: dict
A dictionary mapping statistic names to values.
"""
assert time_dim in dataarray.dims, "The parameter `time_dim` is \"{}\", " \
"but that dimension does not exist in the data.".format(time_dim)
stats = {}
data_np_arr = dataarray.values
time_np_arr = dataarray[time_dim].values
data_inds = np.arange(len(data_np_arr))
# Obtain the first and second derivatives.
fst_deriv = np.gradient(data_np_arr, time_np_arr)
pos_fst_deriv = fst_deriv > 0
neg_fst_deriv = 0 > fst_deriv
snd_deriv = np.gradient(fst_deriv, time_np_arr)
pos_snd_deriv = snd_deriv > 0
neg_snd_deriv = 0 > snd_deriv
# Determine MOS.
# MOS is the index of the highest value immediately preceding a transition
# of the first derivative from positive to negative.
pos_to_neg_fst_deriv = pos_fst_deriv.copy()
for i in range(len(pos_fst_deriv)):
if i == len(pos_fst_deriv) - 1: # last index
pos_to_neg_fst_deriv[i] = False
elif pos_fst_deriv[i] and not pos_fst_deriv[i+1]: # + to -
pos_to_neg_fst_deriv[i] = True
else: # everything else
pos_to_neg_fst_deriv[i] = False
idxmos_potential_inds = data_inds[pos_to_neg_fst_deriv]
idxmos_subset_ind = np.argmax(data_np_arr[pos_to_neg_fst_deriv])
idxmos = idxmos_potential_inds[idxmos_subset_ind]
stats['Middle of Season'] = idxmos
data_inds_after_mos = np.roll(data_inds, len(data_inds)-idxmos-1)
# Determine BOS.
# BOS is the first negative inflection point of the positive values
# of the first derivative starting after and ending at the MOS.
idxbos = data_inds_after_mos[np.argmax((pos_fst_deriv & neg_snd_deriv)[data_inds_after_mos])]
stats['Beginning of Season'] = idxbos
# Determine EOS.
# EOS is the last positive inflection point of the negative values
# of the first derivative starting after and ending at the MOS.
idxeos = data_inds_after_mos[np.argmax((neg_fst_deriv & pos_snd_deriv)[data_inds_after_mos][::-1])]
stats['End of Season'] = idxeos
# Determine EOS-BOS.
stats['Length of Season'] = idxeos - idxbos
# Determine BASE.
stats['Base Value'] = data_np_arr.min()
# Determine MAX.
stats['Max Value'] = data_np_arr.max()
# Determine AMP.
stats['Amplitude'] = stats['Max Value'] - stats['Base Value']
return stats
# +
## Settings
# The minimum number of weeks or months with data for a year to have its stats calculated.
# The aggregation used to obtain the plotting data determines which of these is used.
min_weeks_per_year = 40
min_months_per_year = 9
## End Settings
for year, dataarray in plotting_data_years.items():
dataarray = dataarray.mean(['latitude', 'longitude'])
non_nan_mask = ~np.isnan(dataarray.values)
num_times = non_nan_mask.sum()
insufficient_data = False
if bin_by == 'weekofyear':
if num_times < min_weeks_per_year:
print("There are {} weeks with data for the year {}, but the " \
"minimum number of weeks is {}.\n".format(num_times, year, min_weeks_per_year))
continue
elif bin_by == 'monthofyear':
if num_times < min_months_per_year:
print("There are {} months with data for the year {}, but the " \
"minimum number of months is {}.\n".format(num_times, year, min_months_per_year))
continue
# Remove NaNs for `TIMESAT_stats()`.
dataarray = dataarray.sel({time_dim_name: dataarray[time_dim_name].values[non_nan_mask]})
stats = TIMESAT_stats(dataarray, time_dim=time_dim_name)
# Map indices to days of the year (can't use data from `daysofyear_per_year` directly
# because `xarray_time_series_plot()` can have more points for smooth curve fitting.
time_int_arr = dataarray[time_dim_name].values
orig_day_int_arr = daysofyear_per_year[year].values
day_int_arr = np.interp(time_int_arr, (time_int_arr.min(), time_int_arr.max()),
(orig_day_int_arr.min(), orig_day_int_arr.max()))
# Convert "times" in the TIMESAT stats from indices to days (ints).
stats['Beginning of Season'] = int(round(day_int_arr[stats['Beginning of Season']]))
stats['Middle of Season'] = int(round(day_int_arr[stats['Middle of Season']]))
stats['End of Season'] = int(round(day_int_arr[stats['End of Season']]))
stats['Length of Season'] = np.abs(stats['End of Season'] - stats['Beginning of Season'])
print("Year =", year)
print("Beginning of Season (BOS) day =", stats['Beginning of Season'])
print("End of Season (EOS) day =", stats['End of Season'])
print("Middle of Season (MOS) day =", stats['Middle of Season'])
print("Length of Season (abs(EOS-BOS)) in days =", stats['Length of Season'])
print("Base Value (Min) =", stats['Base Value'])
print("Max Value (Max) =", stats['Max Value'])
print("Amplitude (Max-Min) =", stats['Amplitude'])
print()
# -
# ## Notes on modifications made moving from ARDC to DE Africa
#
# - Replace dc.load fuction with DE Africa dc.load fuction
# - changed code around loading data to just load and mask. Suggest replacing with DE Africa 'load masked usgs'
# - Move all module imports to start of notebook
# - Removed code to view extent of datacube replaced with mapviewer that can be used to visualise spatial extent of data
# - replaced save to geotiff fuction with datacube helper fuction
|
DCAL/DCAL_notebooks/DCAL_Vegetation_Phenology.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nipype Quickstart
#
# **This is a very quick non-imaging introduction to Nipype workflows. For a more comprehensive introduction, check the next section of the tutorial.**
# 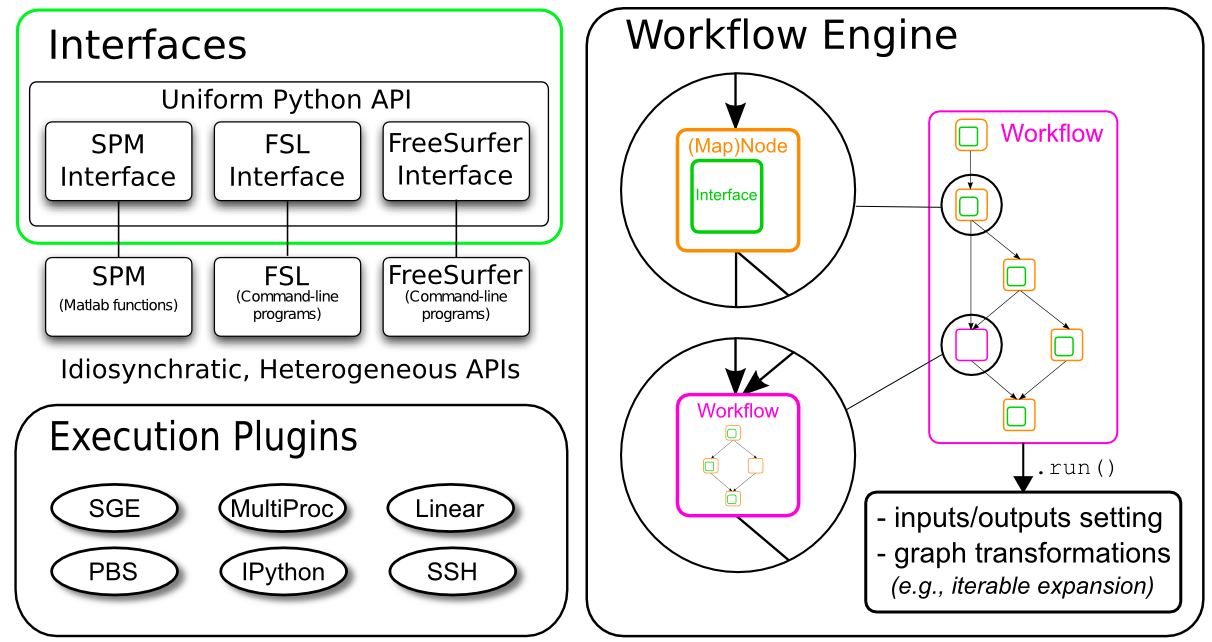
# - [Existing documentation](http://nipype.readthedocs.io/en/latest/)
#
# - [Visualizing the evolution of Nipype](https://www.youtube.com/watch?v=cofpD1lhmKU)
#
# - This notebook is taken from [reproducible-imaging repository](https://github.com/ReproNim/reproducible-imaging)
# #### Import a few things from nipype
import os
from nipype import Workflow, Node, Function
# Creating Workflow with one Node that adds two numbers
# +
def sum(a, b):
return a + b
wf = Workflow('hello')
adder = Node(Function(input_names=['a', 'b'],
output_names=['sum'],
function=sum),
name='a_plus_b')
adder.inputs.a = 1
adder.inputs.b = 3
wf.add_nodes([adder])
wf.base_dir = os.getcwd()
eg = wf.run()
list(eg.nodes())[0].result.outputs
# -
# Creating a second node and connecting to the ``hello`` Workflow
# +
def concat(a, b):
return [a, b]
concater = Node(Function(input_names=['a', 'b'],
output_names=['some_list'],
function=concat),
name='concat_a_b')
wf.connect(adder, 'sum', concater, 'a')
concater.inputs.b = 3
eg = wf.run()
print(eg.nodes())
# -
# And we can check results of our Workflow, we should see a list:
list(eg.nodes())[-1].result.outputs
# We will try to add additional Node that adds one:
# +
def plus_one(a):
return a + 1
plusone = Node(Function(input_names=['a'],
output_names=['out'],
function=plus_one),
name='add_1')
wf.connect(concater, 'some_list', plusone, 'a')
try:
eg = wf.run()
except(RuntimeError) as err:
print("RuntimeError:", err)
else:
raise
# -
# This time the workflow didn't execute cleanly and we got an error. We can use ``nipypecli`` to read the crashfile (note, that if you have multiple crashfiles in the directory you'll have to provide a full name):
# !nipypecli crash crash*
# It clearly shows the problematic Node and its input. We tried to add an integer to a list, this operation is not allowed in Python.
#
# Let's try using MapNode
# +
from nipype import MapNode
plusone = MapNode(Function(input_names=['a'],
output_names=['out'],
function=plus_one),
iterfield=['a'],
name='add_1')
wf = Workflow('hello_mapnode')
adder = Node(Function(input_names=['a', 'b'],
output_names=['sum'],
function=sum),
name='a_plus_b')
adder.inputs.a = 1
adder.inputs.b = 3
wf.connect(adder, 'sum', concater, 'a')
concater.inputs.b = 3
wf.connect(concater, 'some_list', plusone, 'a')
wf.base_dir = os.getcwd()
eg = wf.run()
print(eg.nodes())
# -
# Now the workflow finished without problems, let's see the results from ``hello.add_1``:
print(list(eg.nodes())[2].result.outputs)
# And now we will run the example with ``iterables``:
# +
adder.iterables = ('a', [1, 2])
adder.inputs.b = 2
eg = wf.run()
print(eg.nodes())
# -
# Now we have 6 nodes, we can check results for `` hello.add_1.a1``
list(eg.nodes())[5].result.outputs
wf.write_graph(graph2use='exec')
from IPython.display import Image
# We can plot a general structure of the workflow:
Image("hello_mapnode/graph.png")
# And more detailed structure with all nodes:
Image("hello_mapnode/graph_detailed.png")
# We will introduce another iterables, for the concater Node:
concater.iterables = ('b', [3, 4])
eg = wf.run()
eg.nodes();
wf.write_graph(graph2use='exec')
Image("hello_mapnode/graph_detailed.png")
# Now we will introduce JoinNode that allows us to merge results together:
# +
def merge_and_scale_data(data2):
import numpy as np
return (np.array(data2) * 1000).tolist()
from nipype import JoinNode
joiner = JoinNode(Function(input_names=['data2'],
output_names=['data_scaled'],
function=merge_and_scale_data),
name='join_scale_data',
joinsource=adder,
joinfield=['data2'])
wf.connect(plusone, 'out', joiner, 'data2')
eg = wf.run()
eg.nodes()
# -
# Let's check the output of ``hello.join_scale_data.a0`` node:
list(eg.nodes())[0].result.outputs
wf.write_graph(graph2use='exec')
Image("hello_mapnode/graph.png")
Image("hello_mapnode/graph_detailed.png")
# %time eg = wf.run(plugin='MultiProc', plugin_args={'n_procs': 2})
wf.base_dir = os.path.join(os.getcwd(), 'alt')
# %time eg = wf.run(plugin='MultiProc', plugin_args={'n_procs': 2})
# %time eg = wf.run(plugin='MultiProc', plugin_args={'n_procs': 2})
# ### Exercise 1
#
# Create a workflow to calculate a sum of factorials of numbers from a range between $n_{min}$ and $n_{max}$, i.e.:
#
# $$\sum _{k=n_{min}}^{n_{max}} k! = 0! + 1! +2! + 3! + \cdots$$
#
# if $n_{min}=0$ and $n_{max}=3$
# $$\sum _{k=0}^{3} k! = 0! + 1! +2! + 3! = 1 + 1 + 2 + 6 = 10$$
#
# + solution2="hidden" solution2_first=true
#write your code here
# 1. write 3 functions: one that returns a list of number from a specific range,
# second that returns n! (you can use math.factorial) and third, that sums the elements from a list
# 2. create a workflow and define the working directory
# 3. define 3 nodes using Node and MapNode and connect them within the workflow
# 4. run the workflow and check the results
# + solution2="hidden"
from nipype import Workflow, Node, MapNode, Function
import os
def range_fun(n_min, n_max):
return list(range(n_min, n_max+1))
def factorial(n):
# print("FACTORIAL, {}".format(n))
import math
return math.factorial(n)
def summing(terms):
return sum(terms)
wf_ex1 = Workflow('ex1')
wf_ex1.base_dir = os.getcwd()
range_nd = Node(Function(input_names=['n_min', 'n_max'],
output_names=['range_list'],
function=range_fun),
name='range_list')
factorial_nd = MapNode(Function(input_names=['n'],
output_names=['fact_out'],
function=factorial),
iterfield=['n'],
name='factorial')
summing_nd = Node(Function(input_names=['terms'],
output_names=['sum_out'],
function=summing),
name='summing')
range_nd.inputs.n_min = 0
range_nd.inputs.n_max = 3
wf_ex1.add_nodes([range_nd])
wf_ex1.connect(range_nd, 'range_list', factorial_nd, 'n')
wf_ex1.connect(factorial_nd, 'fact_out', summing_nd, "terms")
eg = wf_ex1.run()
# + [markdown] solution2="hidden"
# let's print all nodes:
# + solution2="hidden"
eg.nodes()
# + [markdown] solution2="hidden"
# the final result should be 10:
# + solution2="hidden"
list(eg.nodes())[2].result.outputs
# + [markdown] solution2="hidden"
# we can also check the results of two other nodes:
# + solution2="hidden"
print(list(eg.nodes())[0].result.outputs)
print(list(eg.nodes())[1].result.outputs)
# -
# ### Exercise 2
#
# Create a workflow to calculate the following sum for chosen $n$ and five different values of $x$: $0$, $\frac{1}{2} \pi$, $\pi$, $\frac{3}{2} \pi$, and $ 2 \pi$.
#
# $\sum _{{k=0}}^{{n}}{\frac {(-1)^{k}}{(2k+1)!}}x^{{2k+1}}\quad =x-{\frac {x^{3}}{3!}}+{\frac {x^{5}}{5!}}-\cdots $
#
# + solution2="hidden" solution2_first=true
# write your solution here
# 1. write 3 functions: one that returns a list of number from a range between 0 and some n,
# second that returns a term for a specific k, and third, that sums the elements from a list
# 2. create a workflow and define the working directory
# 3. define 3 nodes using Node and MapNode and connect them within the workflow
# 4. use iterables for 4 values of x
# 5. run the workflow and check the final results for every value of x
# + solution2="hidden"
# we can reuse function from previous exercise, but they need some edits
from nipype import Workflow, Node, MapNode, JoinNode, Function
import os
import math
def range_fun(n_max):
return list(range(n_max+1))
def term(k, x):
import math
fract = math.factorial(2 * k + 1)
polyn = x ** (2 * k + 1)
return (-1)**k * polyn / fract
def summing(terms):
return sum(terms)
wf_ex2 = Workflow('ex2')
wf_ex2.base_dir = os.getcwd()
range_nd = Node(Function(input_names=['n_max'],
output_names=['range_list'],
function=range_fun),
name='range_list')
term_nd = MapNode(Function(input_names=['k', 'x'],
output_names=['term_out'],
function=term),
iterfield=['k'],
name='term')
summing_nd = Node(Function(input_names=['terms'],
output_names=['sum_out'],
function=summing),
name='summing')
range_nd.inputs.n_max = 15
x_list = [0, 0.5 * math.pi, math.pi, 1.5 * math.pi, 2 * math.pi]
term_nd.iterables = ('x', x_list)
wf_ex2.add_nodes([range_nd])
wf_ex2.connect(range_nd, 'range_list', term_nd, 'k')
wf_ex2.connect(term_nd, 'term_out', summing_nd, "terms")
eg = wf_ex2.run()
# + [markdown] solution2="hidden"
# let's check all nodes
# + solution2="hidden"
eg.nodes()
# + [markdown] solution2="hidden"
# let's print all results of ``ex2.summing``
# + solution2="hidden"
print(list(eg.nodes())[2].result.outputs)
print(list(eg.nodes())[4].result.outputs)
print(list(eg.nodes())[6].result.outputs)
print(list(eg.nodes())[8].result.outputs)
print(list(eg.nodes())[10].result.outputs)
# + [markdown] solution2="hidden"
# Great, we just implemented pretty good Sine function! Those number should be approximately 0, 1, 0, -1 and 0. If they are not, try to increase $n_max$.
# -
# ### Exercise 2a
#
# Use JoinNode to combine results from Exercise 2 in one container, e.g. a dictionary, that takes value $x$ as a key and the result from ``summing`` Node as a value.
# + solution2="hidden" solution2_first=true
# write your code here
# 1. create an additional function that takes 2 lists and combines them into one container, e.g. dictionary
# 2. use JoinNode to define a new node that merges results from Exercise 2 and connect it to the workflow
# 3. run the workflow and check the results of the merging node
# + solution2="hidden"
def merge_results(results, x):
return dict(zip(x, results))
join_nd = JoinNode(Function(input_names=['results', 'x'],
output_names=['results_cont'],
function=merge_results),
name='merge',
joinsource=term_nd, # this is the node that used iterables for x
joinfield=['results'])
# taking the list of arguments from the previous part
join_nd.inputs.x = x_list
# connecting a new node to the summing_nd
wf_ex2.connect(summing_nd, "sum_out", join_nd, "results")
eg = wf_ex2.run()
# + [markdown] solution2="hidden"
# let's print all nodes
# + solution2="hidden"
eg.nodes()
# + [markdown] solution2="hidden"
# and results from ``merge`` Node:
# + solution2="hidden"
list(eg.nodes())[1].result.outputs
|
workshop/nipype_tutorial/notebooks/introduction_quickstart_non-neuroimaging.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from miniutils.progress_bar import progbar, parallel_progbar, iparallel_progbar
import time
def mapper(x):
time.sleep(1)
return x ** 2
def flatmapper(x):
interval = x / 5
return [i * interval for i in range(5) if time.sleep(0.2) is None]
lst = list(range(5))
long_lst = list(range(50))
print([mapper(x) for x in lst])
print([y for x in lst for y in flatmapper(x)])
# -
[mapper(x) for x in progbar(lst)]
parallel_progbar(mapper, lst, nprocs=2)
for k in iparallel_progbar(mapper, lst, nprocs=2):
print(k, flush=True)
parallel_progbar(flatmapper, lst, nprocs=2, flatmap=True)
for k in iparallel_progbar(flatmapper, lst, nprocs=len(lst), flatmap=True):
print(k, flush=True)
parallel_progbar(flatmapper, long_lst, nprocs=len(lst), flatmap=True)
|
tests/visual_tests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
import csv
import ast
import sys
import operator
# !conda install --yes --prefix {sys.prefix} pandas
import pandas as pd
lexicon_file = "lexicon/NRC-Emotion-Lexicon-v0.92/NRC-Emotion-Lexicon-Wordlevel-v0.92.txt"
lexicon_df = pd.read_csv(lexicon_file, names=["word", "emotion", "association"], sep='\t')
def lyrics_decoder(lyrics):
"""
Fixes the UTF8 encoding and decodes it
:param lyrics: invalid UTF8 encoded lyrics
:return: decoded lyrics
"""
return ast.literal_eval(lyrics).decode('utf8')
with open('lyrics/bh100.csv', 'r') as csvfile:
reader = csv.reader(csvfile, delimiter=',')
next(reader)
with open('results/sentiments.csv', 'w', newline="") as csvfile2:
filewriter = csv.writer(
csvfile2,
delimiter=',',
quotechar='|',
quoting=csv.QUOTE_MINIMAL
)
filewriter.writerow(
[
'Rank','Song','Artist','Year','Main Emotion',"anger","anticipation","disgust","fear","joy","sadness","surprise","trust","neutral","na"
]
)
for song in reader:
emotion_score = {
"anger":0,
"anticipation":0,
"disgust":0,
"fear":0,
"joy":0,
"sadness":0,
"surprise":0,
"trust":0,
"neutral":0,
"na":0,
}
song_rank = song[0]
song_name = song[1]
song_artist = song[2]
song_year = song[3]
try:
song_lyrics = lyrics_decoder(song[4])
for word in song_lyrics.split():
clean_word = word.replace("'",'')
matches = lexicon_df.loc[lexicon_df['word'] == clean_word]
if not matches.empty:
for emotions in matches.iterrows():
is_neutral = True
for emotion in emotions:
if type(emotion) != int \
and emotion.emotion !='positive'\
and emotion.emotion !='negative':
emotion_score[emotion.emotion] += emotion.association
if is_neutral:
emotion_score["neutral"] = 1
total_score = 0
for emotion in emotion_score:
total_score += emotion_score[emotion]
for emotion in emotion_score:
if total_score != 0:
emotion_score[emotion] = emotion_score[emotion]*100/total_score
else:
emotion_score['na'] = 100
main_emotion = max(emotion_score.items(), key=operator.itemgetter(1))[0]
filewriter.writerow(
[
song_rank,song_name,song_artist,song_year,main_emotion,emotion_score["anger"],
emotion_score["anticipation"], emotion_score["disgust"], emotion_score["fear"],
emotion_score["joy"], emotion_score["sadness"], emotion_score["surprise"],
emotion_score["trust"],emotion_score["neutral"], emotion_score["na"]
]
)
except:
filewriter.writerow(
[
song_rank,song_name,song_artist,song_year,"null",0,0,0,0,0,0,0,0,0,100
]
)
print("Done.")
|
2_sentiment_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Copyright 2019 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# -
# <img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
#
# # Torch-TensorRT Getting Started - CitriNet
# ## Overview
#
# [Citrinet](https://docs.nvidia.com/deeplearning/nemo/user-guide/docs/en/main/asr/models.html#citrinet) is an acoustic model used for the speech to text recognition task. It is a version of [QuartzNet](https://arxiv.org/pdf/1910.10261.pdf) that extends [ContextNet](https://arxiv.org/pdf/2005.03191.pdf), utilizing subword encoding (via Word Piece tokenization) and Squeeze-and-Excitation(SE) mechanism and are therefore smaller than QuartzNet models.
#
# CitriNet models take in audio segments and transcribe them to letter, byte pair, or word piece sequences.
#
# <img src="https://docs.nvidia.com/deeplearning/nemo/user-guide/docs/en/main/_images/jasper_vertical.png" alt="alt" width="50%"/>
#
# ### Learning objectives
#
# This notebook demonstrates the steps for optimizing a pretrained CitriNet model with Torch-TensorRT, and running it to test the speedup obtained.
#
# ## Content
# 1. [Requirements](#1)
# 1. [Download Citrinet model](#2)
# 1. [Create Torch-TensorRT modules](#3)
# 1. [Benchmark Torch-TensorRT models](#4)
# 1. [Conclusion](#5)
# <a id="1"></a>
# ## 1. Requirements
#
# Follow the steps in [README](README.md) to prepare a Docker container, within which you can run this notebook.
# This notebook assumes that you are within a Jupyter environment in a docker container with Torch-TensorRT installed, such as an NGC monthly release of `nvcr.io/nvidia/pytorch:<yy.mm>-py3` (where `yy` indicates the last two numbers of a calendar year, and `mm` indicates the month in two-digit numerical form)
#
# Now that you are in the docker, the next step is to install the required dependencies.
# +
# Install dependencies
# !pip install wget
# !apt-get update && DEBIAN_FRONTEND=noninteractive apt-get install -y libsndfile1 ffmpeg
# !pip install Cython
## Install NeMo
# !pip install nemo_toolkit[all]==1.5.1
# -
# <a id="2"></a>
# ## 2. Download Citrinet model
#
# Next, we download a pretrained Nemo Citrinet model and convert it to a Torchscript module:
# +
import nemo
import torch
import nemo.collections.asr as nemo_asr
from nemo.core import typecheck
typecheck.set_typecheck_enabled(False)
# +
variant = 'stt_en_citrinet_256'
print(f"Downloading and saving {variant}...")
asr_model = nemo_asr.models.EncDecCTCModelBPE.from_pretrained(model_name=variant)
asr_model.export(f"{variant}.ts")
# -
# ### Benchmark utility
#
# Let us define a helper benchmarking function, then benchmark the original Pytorch model.
# +
from __future__ import print_function
from __future__ import absolute_import
from __future__ import division
import argparse
import timeit
import numpy as np
import torch
import torch_tensorrt as trtorch
import torch.backends.cudnn as cudnn
def benchmark(model, input_tensor, num_loops, model_name, batch_size):
def timeGraph(model, input_tensor, num_loops):
print("Warm up ...")
with torch.no_grad():
for _ in range(20):
features = model(input_tensor)
torch.cuda.synchronize()
print("Start timing ...")
timings = []
with torch.no_grad():
for i in range(num_loops):
start_time = timeit.default_timer()
features = model(input_tensor)
torch.cuda.synchronize()
end_time = timeit.default_timer()
timings.append(end_time - start_time)
# print("Iteration {}: {:.6f} s".format(i, end_time - start_time))
return timings
def printStats(graphName, timings, batch_size):
times = np.array(timings)
steps = len(times)
speeds = batch_size / times
time_mean = np.mean(times)
time_med = np.median(times)
time_99th = np.percentile(times, 99)
time_std = np.std(times, ddof=0)
speed_mean = np.mean(speeds)
speed_med = np.median(speeds)
msg = ("\n%s =================================\n"
"batch size=%d, num iterations=%d\n"
" Median samples/s: %.1f, mean: %.1f\n"
" Median latency (s): %.6f, mean: %.6f, 99th_p: %.6f, std_dev: %.6f\n"
) % (graphName,
batch_size, steps,
speed_med, speed_mean,
time_med, time_mean, time_99th, time_std)
print(msg)
timings = timeGraph(model, input_tensor, num_loops)
printStats(model_name, timings, batch_size)
precisions_str = 'fp32' # Precision (default=fp32, fp16)
variant = 'stt_en_citrinet_256' # Nemo Citrinet variant
batch_sizes = [1, 8, 32, 128] # Batch sizes (default=1,8,32,128)
trt = False # If True, infer with Torch-TensorRT engine. Else, infer with Pytorch model.
precision = torch.float32 if precisions_str =='fp32' else torch.float16
for batch_size in batch_sizes:
if trt:
model_name = f"{variant}_bs{batch_size}_{precision}.torch-tensorrt"
else:
model_name = f"{variant}.ts"
print(f"Loading model: {model_name}")
# Load traced model to CPU first
model = torch.jit.load(model_name).cuda()
cudnn.benchmark = True
# Create random input tensor of certain size
torch.manual_seed(12345)
input_shape=(batch_size, 80, 1488)
input_tensor = torch.randn(input_shape).cuda()
# Timing graph inference
benchmark(model, input_tensor, 50, model_name, batch_size)
# -
# Confirming the GPU we are using here:
# !nvidia-smi
# <a id="3"></a>
# ## 3. Create Torch-TensorRT modules
#
# In this step, we optimize the Citrinet Torchscript module with Torch-TensorRT with various precisions and batch sizes.
# +
import torch
import torch.nn as nn
import torch_tensorrt as torchtrt
import argparse
variant = "stt_en_citrinet_256"
precisions = [torch.float, torch.half]
batch_sizes = [1,8,32,128]
model = torch.jit.load(f"{variant}.ts")
for precision in precisions:
for batch_size in batch_sizes:
compile_settings = {
"inputs": [torchtrt.Input(shape=[batch_size, 80, 1488])],
"enabled_precisions": {precision},
"workspace_size": 2000000000,
"truncate_long_and_double": True,
}
print(f"Generating Torchscript-TensorRT module for batchsize {batch_size} precision {precision}")
trt_ts_module = torchtrt.compile(model, **compile_settings)
torch.jit.save(trt_ts_module, f"{variant}_bs{batch_size}_{precision}.torch-tensorrt")
# -
# <a id="4"></a>
# ## 4. Benchmark Torch-TensorRT models
#
# Finally, we are ready to benchmark the Torch-TensorRT optimized Citrinet models.
# ### FP32 (single precision)
# +
precisions_str = 'fp32' # Precision (default=fp32, fp16)
batch_sizes = [1, 8, 32, 128] # Batch sizes (default=1,8,32,128)
precision = torch.float32 if precisions_str =='fp32' else torch.float16
trt = True
for batch_size in batch_sizes:
if trt:
model_name = f"{variant}_bs{batch_size}_{precision}.torch-tensorrt"
else:
model_name = f"{variant}.ts"
print(f"Loading model: {model_name}")
# Load traced model to CPU first
model = torch.jit.load(model_name).cuda()
cudnn.benchmark = True
# Create random input tensor of certain size
torch.manual_seed(12345)
input_shape=(batch_size, 80, 1488)
input_tensor = torch.randn(input_shape).cuda()
# Timing graph inference
benchmark(model, input_tensor, 50, model_name, batch_size)
# -
# ### FP16 (half precision)
# +
precisions_str = 'fp16' # Precision (default=fp32, fp16)
batch_sizes = [1, 8, 32, 128] # Batch sizes (default=1,8,32,128)
precision = torch.float32 if precisions_str =='fp32' else torch.float16
for batch_size in batch_sizes:
if trt:
model_name = f"{variant}_bs{batch_size}_{precision}.torch-tensorrt"
else:
model_name = f"{variant}.ts"
print(f"Loading model: {model_name}")
# Load traced model to CPU first
model = torch.jit.load(model_name).cuda()
cudnn.benchmark = True
# Create random input tensor of certain size
torch.manual_seed(12345)
input_shape=(batch_size, 80, 1488)
input_tensor = torch.randn(input_shape).cuda()
# Timing graph inference
benchmark(model, input_tensor, 50, model_name, batch_size)
# -
# <a id="5"></a>
# ## 5. Conclusion
#
# In this notebook, we have walked through the complete process of optimizing the Citrinet model with Torch-TensorRT. On an A100 GPU, with Torch-TensorRT, we observe a speedup of ~**2.4X** with FP32, and ~**2.9X** with FP16 at batchsize of 128.
#
# ### What's next
# Now it's time to try Torch-TensorRT on your own model. Fill out issues at https://github.com/NVIDIA/Torch-TensorRT. Your involvement will help future development of Torch-TensorRT.
#
|
docs/_notebooks/CitriNet-example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] heading_collapsed=true
# ## Movielens
# + hidden=true
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai.learner import *
from fastai.column_data import *
# + [markdown] hidden=true
# Data available from http://files.grouplens.org/datasets/movielens/ml-latest-small.zip
# + hidden=true
path='data/ml-latest-small/'
# + [markdown] hidden=true
# We're working with the movielens data, which contains one rating per row, like this:
# + hidden=true
ratings = pd.read_csv(path+'ratings.csv')
ratings.head()
# + [markdown] hidden=true
# Just for display purposes, let's read in the movie names too.
# + hidden=true
movies = pd.read_csv(path+'movies.csv')
movies.head()
# + [markdown] heading_collapsed=true
# ## Create subset for Excel
# + [markdown] hidden=true
# We create a crosstab of the most popular movies and most movie-addicted users which we'll copy into Excel for creating a simple example. This isn't necessary for any of the modeling below however.
# + hidden=true
g=ratings.groupby('userId')['rating'].count()
topUsers=g.sort_values(ascending=False)[:15]
g=ratings.groupby('movieId')['rating'].count()
topMovies=g.sort_values(ascending=False)[:15]
top_r = ratings.join(topUsers, rsuffix='_r', how='inner', on='userId')
top_r = top_r.join(topMovies, rsuffix='_r', how='inner', on='movieId')
pd.crosstab(top_r.userId, top_r.movieId, top_r.rating, aggfunc=np.sum)
# + [markdown] heading_collapsed=true
# ## Collaborative filtering
# + hidden=true
val_idxs = get_cv_idxs(len(ratings))
wd=2e-4
n_factors = 50
# + hidden=true
cf = CollabFilterDataset.from_csv(path, 'ratings.csv', 'userId', 'movieId', 'rating')
learn = cf.get_learner(n_factors, val_idxs, 64, opt_fn=optim.Adam)
# + hidden=true
learn.fit(1e-2, 2, wds=wd, cycle_len=1, cycle_mult=2, use_wd_sched=True)
# + [markdown] hidden=true
# Let's compare to some benchmarks. Here's [some benchmarks](https://www.librec.net/release/v1.3/example.html) on the same dataset for the popular Librec system for collaborative filtering. They show best results based on [RMSE](http://www.statisticshowto.com/rmse/) of 0.91. We'll need to take the square root of our loss, since we use plain MSE.
# + hidden=true
math.sqrt(0.776)
# + [markdown] hidden=true
# Looking good - we've found a solution better than any of those benchmarks! Let's take a look at how the predictions compare to actuals for this model.
# + hidden=true
preds = learn.predict()
# + hidden=true
y=learn.data.val_y
sns.jointplot(preds, y, kind='hex', stat_func=None);
# -
# ## Analyze results
# + [markdown] heading_collapsed=true
# ### Movie bias
# + hidden=true
movie_names = movies.set_index('movieId')['title'].to_dict()
g=ratings.groupby('movieId')['rating'].count()
topMovies=g.sort_values(ascending=False).index.values[:3000]
topMovieIdx = np.array([cf.item2idx[o] for o in topMovies])
# + hidden=true
m=learn.model; m.cuda()
# + [markdown] hidden=true
# First, we'll look at the movie bias term. Here, our input is the movie id (a single id), and the output is the movie bias (a single float).
# + hidden=true
movie_bias = to_np(m.ib(V(topMovieIdx)))
# + hidden=true
movie_bias
# + hidden=true
movie_ratings = [(b[0], movie_names[i]) for i,b in zip(topMovies,movie_bias)]
# + [markdown] hidden=true
# Now we can look at the top and bottom rated movies. These ratings are corrected for different levels of reviewer sentiment, as well as different types of movies that different reviewers watch.
# + hidden=true
sorted(movie_ratings, key=lambda o: o[0])[:15]
# + hidden=true
sorted(movie_ratings, key=itemgetter(0))[:15]
# + hidden=true
sorted(movie_ratings, key=lambda o: o[0], reverse=True)[:15]
# + [markdown] heading_collapsed=true
# ### Embedding interpretation
# + [markdown] hidden=true
# We can now do the same thing for the embeddings.
# + hidden=true
movie_emb = to_np(m.i(V(topMovieIdx)))
movie_emb.shape
# + [markdown] hidden=true
# Because it's hard to interpret 50 embeddings, we use [PCA](https://plot.ly/ipython-notebooks/principal-component-analysis/) to simplify them down to just 3 vectors.
# + hidden=true
from sklearn.decomposition import PCA
pca = PCA(n_components=3)
movie_pca = pca.fit(movie_emb.T).components_
# + hidden=true
movie_pca.shape
# + hidden=true
fac0 = movie_pca[0]
movie_comp = [(f, movie_names[i]) for f,i in zip(fac0, topMovies)]
# + [markdown] hidden=true
# Here's the 1st component. It seems to be 'easy watching' vs 'serious'.
# + hidden=true
sorted(movie_comp, key=itemgetter(0), reverse=True)[:10]
# + hidden=true
sorted(movie_comp, key=itemgetter(0))[:10]
# + hidden=true
fac1 = movie_pca[1]
movie_comp = [(f, movie_names[i]) for f,i in zip(fac1, topMovies)]
# + [markdown] hidden=true
# Here's the 2nd component. It seems to be 'CGI' vs 'dialog driven'.
# + hidden=true
sorted(movie_comp, key=itemgetter(0), reverse=True)[:10]
# + hidden=true
sorted(movie_comp, key=itemgetter(0))[:10]
# + [markdown] hidden=true
# We can draw a picture to see how various movies appear on the map of these components. This picture shows the first two components.
# + hidden=true
idxs = np.random.choice(len(topMovies), 50, replace=False)
X = fac0[idxs]
Y = fac1[idxs]
plt.figure(figsize=(15,15))
plt.scatter(X, Y)
for i, x, y in zip(topMovies[idxs], X, Y):
plt.text(x,y,movie_names[i], color=np.random.rand(3)*0.7, fontsize=11)
plt.show()
# -
# ## Collab filtering from scratch
# + [markdown] heading_collapsed=true
# ### Dot product example
# + hidden=true
a = T([[1.,2],[3,4]])
b = T([[2.,2],[10,10]])
a,b
# + hidden=true
a*b
# + hidden=true
(a*b).sum(1)
# + hidden=true
class DotProduct(nn.Module):
def forward(self, u, m): return (u*m).sum(1)
# + hidden=true
model=DotProduct()
# + hidden=true
model(a,b)
# + [markdown] heading_collapsed=true
# ### Dot product model
# + hidden=true
u_uniq = ratings.userId.unique()
user2idx = {o:i for i,o in enumerate(u_uniq)}
ratings.userId = ratings.userId.apply(lambda x: user2idx[x])
m_uniq = ratings.movieId.unique()
movie2idx = {o:i for i,o in enumerate(m_uniq)}
ratings.movieId = ratings.movieId.apply(lambda x: movie2idx[x])
n_users=int(ratings.userId.nunique())
n_movies=int(ratings.movieId.nunique())
# + hidden=true
class EmbeddingDot(nn.Module):
def __init__(self, n_users, n_movies):
super().__init__()
self.u = nn.Embedding(n_users, n_factors)
self.m = nn.Embedding(n_movies, n_factors)
self.u.weight.data.uniform_(0,0.05)
self.m.weight.data.uniform_(0,0.05)
def forward(self, cats, conts):
users,movies = cats[:,0],cats[:,1]
u,m = self.u(users),self.m(movies)
return (u*m).sum(1)
# + hidden=true
x = ratings.drop(['rating', 'timestamp'],axis=1)
y = ratings['rating'].astype(np.float32)
# + hidden=true
data = ColumnarModelData.from_data_frame(path, val_idxs, x, y, ['userId', 'movieId'], 64)
# + hidden=true
wd=1e-5
model = EmbeddingDot(n_users, n_movies).cuda()
opt = optim.SGD(model.parameters(), 1e-1, weight_decay=wd, momentum=0.9)
# + hidden=true
fit(model, data, 3, opt, F.mse_loss)
# + hidden=true
set_lrs(opt, 0.01)
# + hidden=true
fit(model, data, 3, opt, F.mse_loss)
# + [markdown] heading_collapsed=true
# ### Bias
# + hidden=true
min_rating,max_rating = ratings.rating.min(),ratings.rating.max()
min_rating,max_rating
# + hidden=true
def get_emb(ni,nf):
e = nn.Embedding(ni, nf)
e.weight.data.uniform_(-0.01,0.01)
return e
class EmbeddingDotBias(nn.Module):
def __init__(self, n_users, n_movies):
super().__init__()
(self.u, self.m, self.ub, self.mb) = [get_emb(*o) for o in [
(n_users, n_factors), (n_movies, n_factors), (n_users,1), (n_movies,1)
]]
def forward(self, cats, conts):
users,movies = cats[:,0],cats[:,1]
um = (self.u(users)* self.m(movies)).sum(1)
res = um + self.ub(users).squeeze() + self.mb(movies).squeeze()
res = F.sigmoid(res) * (max_rating-min_rating) + min_rating
return res
# + hidden=true
wd=2e-4
model = EmbeddingDotBias(cf.n_users, cf.n_items).cuda()
opt = optim.SGD(model.parameters(), 1e-1, weight_decay=wd, momentum=0.9)
# + hidden=true
fit(model, data, 3, opt, F.mse_loss)
# + hidden=true
set_lrs(opt, 1e-2)
# + hidden=true
fit(model, data, 3, opt, F.mse_loss)
# -
# ### Mini net
# + code_folding=[]
class EmbeddingNet(nn.Module):
def __init__(self, n_users, n_movies, nh=10, p1=0.05, p2=0.5):
super().__init__()
(self.u, self.m) = [get_emb(*o) for o in [
(n_users, n_factors), (n_movies, n_factors)]]
self.lin1 = nn.Linear(n_factors*2, nh)
self.lin2 = nn.Linear(nh, 1)
self.drop1 = nn.Dropout(p1)
self.drop2 = nn.Dropout(p2)
def forward(self, cats, conts):
users,movies = cats[:,0],cats[:,1]
x = self.drop1(torch.cat([self.u(users),self.m(movies)], dim=1))
x = self.drop2(F.relu(self.lin1(x)))
return F.sigmoid(self.lin2(x)) * (max_rating-min_rating+1) + min_rating-0.5
# -
wd=1e-5
model = EmbeddingNet(n_users, n_movies).cuda()
opt = optim.Adam(model.parameters(), 1e-3, weight_decay=wd)
fit(model, data, 3, opt, F.mse_loss)
set_lrs(opt, 1e-3)
fit(model, data, 3, opt, F.mse_loss)
|
courses/dl1/lesson5-movielens.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import os
import numpy as np
# %matplotlib inline
import seaborn as sb
import missingno as msn
sb.set()
# +
train = pd.read_csv('C:/Users/<NAME>/Desktop/Data/train1.csv')
train.head()
# -
msn.matrix(train)
train.isnull().sum()
# +
train['DEW_POINT'].value_counts()
# +
train = train.replace(to_replace = "M01", value =-1)
train = train.replace(to_replace = "M02", value =-2)
train = train.replace(to_replace = "CALM", value =0)
train = train.replace(to_replace = "CALM CALM", value =0)
train.head()
# -
train.info()
train.to_csv("C:\\Users\\<NAME>\\Desktop\\train.csv", index=None)
DP = pd.DataFrame(train['DEW_POINT'])
DP.isnull().sum()
DP = DP.infer_objects()
DP.info()
DP.median()
DP.fillna(9.0, inplace=True)
DP.isnull().sum()
train = train.drop('DATE', axis=1)
train = train.drop('TIME', axis=1)
train = train.drop('DT', axis=1)
train.info()
for i in range(len(train['DEW_POINT'])):
train['DEW_POINT'][i] = float(DP['DEW_POINT'][i])
train.to_csv("C:\\Users\\Jitender kumar\\Desktop\\train.csv", index=None)
train.info()
|
Data Preprocessing/Day_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv
# language: python
# name: venv
# ---
# # Введение
#
# Эти интерактивные тетрадки подразумевают последовательное выполнение.
# Нажмите `Shift+Enter` чтобы выполнить ячейку с кодом.
print("Hello","World")
# Для работы с символьными вычисленями будем пользоваться пакетом `sympy`:
from sympy import *
# Символы для вычислений создаются через `symbols`:
x, y, t = symbols("x, y, t")
alpha, beta, gamma = symbols("alpha, beta, gamma", cls=Function)
alpha(t) + x
# Матрицы создаются из списка списков:
m = Matrix([
[cos(alpha(t)), -sin(alpha(t)), 0],
[sin(alpha(t)), cos(alpha(t)), 0],
[0, 0, 1]
])
v = Matrix([
[x],
[y],
[1]
])
rotated = m * v
rotated
# `sympy` позволяет легко дифферинцировать выражения:
velocity = diff(rotated, t)
velocity[0]
velocity[1]
# Помимо этого, можно заменять части выражений.
# Например, если
# $$\alpha(t) = t$$
# получим следующие скорости:
velocity[0].replace(alpha(t), t).simplify()
velocity[1].replace(alpha(t), t).simplify()
# А если вращение происходит с постоянным угловым ускорением
# $$ \alpha(t) = t^2 $$
# получим следующие линейные скорости:
velocity[0].replace(alpha(t), t**2).simplify()
velocity[1].replace(alpha(t), t**2).simplify()
|
0 - Intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prediction of World Cup 2018 Qualifications in Final Phases
# This notebook runs simulations of the 2018 soccer world cup, taking into account the probabilities that each team has to win each pool match, as well as the "kickscore" of each team (both coming from [kickoff.ai](http://kickoff.ai/)).
#
# As the notebook has been written before the start of the WC, only the teams playing pool matches are known. For latter stages (where adversaries are not yet known), the simulator uses the "kickscore" $s_A$ and $s_B$ of two opposing teams $A$ and $B$, in order to obtain the probability $p$ that the team with the highest kickscore wins: $$p = \frac{1}{1 + e^{-c|s_A - s_B|}},$$
# where $c$ is some scaling constant.
#
# This logic should also work with other notions of "team strength"; so you can try plugging your own!
#
# Once the simulations are run, the notebook explores the quality of some predictions. Predictions here means deciding the set of teams that will be in 8th of final, 4th, etc. as well as who will be 1st, 2nd and third.
# One can use the function `points_for_preds()` in order to know the full distribution of points (for a given betting system) that results from a given set of predictions. From the simulation results, it is easy to retrieve the prediction that maximizes the expected number of points. However, there might be other predictions that bring nearly as many points on expectation but are more diverse (e.g., involving more teams in the latter stages). Such predictions reduce risk (while paying only a small price in expected reward) and are more likely to maintain the suspense high until the end for the player!
from collections import defaultdict
from tqdm import tqdm
import math
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import pprint
from random import random
matplotlib.rcParams.update({'font.size': 16})
# ## First, some data
# Here we encode the probabilities of each team winning, the level of each team (coming directly from [kickoff.ai](http://kickoff.ai)), as well as the structure of the groups and finals.
# +
# expressed in percentage (team1_win, draw, team2_win)
pools_probas = {
('Russia', 'Saudi Arabia'): (57, 25, 18),
('Egypt', 'Uruguay'): (14, 25, 61),
('Morocco', 'Iran'): (26, 30, 44),
('Portugal', 'Spain'): (18, 26, 56),
('France', 'Australia'): (61, 24, 15),
('Argentina', 'Iceland'): (73, 19, 8),
('Peru', 'Denmark'): (44, 30, 26),
('Croatia', 'Nigeria'): (56, 26, 18),
('Costa Rica', 'Serbia'): (34, 30, 36),
('Germany', 'Mexico'): (50, 28, 22),
('Brazil', 'Switzerland'): (66, 22, 12),
('Sweden', 'South Korea'): (47, 29, 24),
('Belgium', 'Panama'): (68, 22, 10),
('Tunisia', 'England'): (9, 21, 70),
('Colombia', 'Japan'): (59, 25, 16),
('Poland', 'Senegal'): (41, 30, 29),
('Russia', 'Egypt'): (42, 29, 29),
('Portugal', 'Morocco'): (56, 27, 17),
('Uruguay', 'Saudi Arabia'): (73, 19, 8),
('Iran', 'Spain'): (11, 22, 67),
('Denmark', 'Australia'): (39, 30, 31),
('France', 'Peru'): (47, 29, 24),
('Argentina', 'Croatia'): (50, 28, 22),
('Brazil', 'Costa Rica'): (78, 16, 6),
('Nigeria', 'Iceland'): (38, 30, 32),
('Serbia', 'Switzerland'): (24, 29, 47),
('Belgium', 'Tunisia'): (61, 25, 14),
('South Korea', 'Mexico'): (19, 27, 54),
('Germany', 'Sweden'): (57, 26, 17),
('England', 'Panama'): (77, 16, 7),
('Japan', 'Senegal'): (34, 30, 36),
('Poland', 'Colombia'): (21, 28, 51),
('Uruguay', 'Russia'): (54, 27, 19),
('Saudi Arabia', 'Egypt'): (24, 29, 47),
('Spain', 'Morocco'): (75, 18, 7),
('Iran', 'Portugal'): (24, 29, 47),
('Denmark', 'France'): (17, 26, 57),
('Australia', 'Peru'): (23, 29, 48),
('Nigeria', 'Argentina'): (10, 20, 70),
('Iceland', 'Croatia'): (16, 25, 59),
('Mexico', 'Sweden'): (42, 30, 28),
('South Korea', 'Germany'): (10, 20, 70),
('Switzerland', 'Costa Rica'): (54, 27, 19),
('Serbia', 'Brazil'): (7, 17, 76),
('Japan', 'Poland'): (33, 31, 36),
('Senegal', 'Colombia'): (19, 27, 54),
('Panama', 'Tunisia'): (28, 30, 42),
('England', 'Belgium'): (44, 29, 27)
}
kickscores = {
'Russia': 0.86, 'Saudi Arabia': 0.46, 'Egypt': 0.76, 'Uruguay': 1.27,
'Portugal': 1.25, 'Spain': 1.75, 'Morocco': 0.87, 'Iran': 1.02,
'France': 1.51, 'Australia': 0.86, 'Peru': 1.21, 'Denmark': 1.01,
'Argentina': 1.54, 'Iceland': 0.62, 'Croatia': 1.29, 'Nigeria': 0.98,
'Brazil': 1.97, 'Switzerland': 1.17, 'Costa Rica': 0.8, 'Serbia': 1.02,
'Germany': 1.62, 'Mexico': 1.25, 'Sweden': 1.12, 'South Korea': 0.67,
'Belgium': 1.3, 'Panama': 0.41, 'Tunisia': 0.77, 'England': 1.68,
'Poland': 1.08, 'Senegal': 0.9, 'Colombia': 1.29, 'Japan': 0.75
}
groups = {
'A': ['Russia', 'Saudi Arabia', 'Egypt', 'Uruguay'],
'B': ['Portugal', 'Spain', 'Morocco', 'Iran'],
'C': ['France', 'Australia', 'Peru', 'Denmark'],
'D': ['Argentina', 'Iceland', 'Croatia', 'Nigeria'],
'E': ['Brazil', 'Switzerland', 'Costa Rica', 'Serbia'],
'F': ['Germany', 'Mexico', 'Sweden', 'South Korea'],
'G': ['Belgium', 'Panama', 'Tunisia', 'England'],
'H': ['Poland', 'Senegal', 'Colombia', 'Japan']
}
# who will play against who?
finals_8 = {
'8A': ((1, 'F'), (2, 'E')), # match 'A' of 8th of finals will be 'F1' vs 'E2'
'8B': ((1, 'H'), (2, 'G')),
'8C': ((2, 'A'), (1, 'B')),
'8D': ((1, 'D'), (2, 'C')),
'8E': ((2, 'F'), (1, 'E')),
'8F': ((2, 'H'), (1, 'G')),
'8G': ((2, 'B'), (1, 'A')),
'8H': ((2, 'D'), (1, 'C'))
}
finals_4 = {
'4A': ('8A', '8B'),
'4B': ('8C', '8D'),
'4C': ('8E', '8F'),
'4D': ('8G', '8H')
}
finals_2 = { '2A': ('4A', '4B'), '2B': ('4C', '4D') }
final = {'1A': ('2A', '2B')}
""" The known winners of past matches (to fill in as the matches are observed)
Map match opponents to winner team (or "draw")
"""
observed_results_groups = {
('Russia', 'Saudi Arabia'): 'Russia',
('Uruguay', 'Egypt'): 'Uruguay',
('Morocco', 'Iran'): 'Iran',
('Portugal', 'Spain'): 'draw',
('France', 'Australia'): 'France',
('Argentina', 'Iceland'): 'draw',
('Peru', 'Denmark'): 'Denmark',
('Croatia', 'Nigeria'): 'Croatia',
('Costa Rica', 'Serbia'): 'Serbia',
('Germany', 'Mexico'): 'Mexico',
('Brazil', 'Switzerland'): 'draw',
('Sweden', 'South Korea'): 'Sweden',
('Belgium', 'Panama'): 'Belgium',
('Tunisia', 'England'): 'England',
('Colombia', 'Japan'): 'Japan',
('Poland', 'Senegal'): 'Senegal',
('Russia', 'Egypt'): 'Russia',
('Portugal', 'Morocco'): 'Portugal',
('Uruguay', 'Saudi Arabia'): 'Uruguay',
('Iran', 'Spain'): 'Spain',
('Denmark', 'Australia'): 'draw',
('France', 'Peru'): 'France',
('Argentina', 'Croatia'): 'Croatia',
('Brazil', 'Costa Rica'): 'Brazil',
('Nigeria', 'Iceland'): 'Nigeria',
('Serbia', 'Switzerland'): 'Switzerland',
('Belgium', 'Tunisia'): 'Belgium',
('South Korea', 'Mexico'): 'Mexico',
('Germany', 'Sweden'): 'Germany',
('England', 'Panama'): 'England',
('Japan', 'Senegal'): 'draw',
('Poland', 'Colombia'): 'Colombia',
('Saudi Arabia', 'Egypt'): 'Saudi Arabia',
('Uruguay', 'Russia'): 'Uruguay',
('Spain', 'Morocco'): 'draw',
('Iran', 'Portugal'): 'draw',
('Australia', 'Peru'): 'Peru',
('Denmark', 'France'): 'draw',
('Iceland', 'Croatia'): 'Croatia',
('Nigeria', 'Argentina'): 'Argentina',
('Mexico', 'Sweden'): 'Sweden',
('South Korea', 'Germany'): 'South Korea',
('Switzerland', 'Costa Rica'): 'draw',
('Serbia', 'Brazil'): 'Brazil',
('Senegal', 'Colombia'): 'Colombia',
('Japan', 'Poland'): 'Poland',
('Panama', 'Tunisia'): 'Tunisia',
('England', 'Belgium'): 'Belgium'
}
observed_results_finals = {
}
observed_groups_orderings = {
'A': ['Uruguay', 'Russia', 'Saudi Arabia', 'Egypt'],
'B': ['Spain', 'Portugal', 'Iran', 'Morocco'],
'C': ['France', 'Denmark', 'Peru', 'Australia'],
'D': ['Croatia', 'Argentina', 'Nigeria', 'Iceland'],
'E': ['Brazil', 'Switzerland', 'Serbia', 'Costa Rica'],
'F': ['Sweden', 'Mexico', 'South Korea', 'Germany'],
'G': ['Belgium', 'England', 'Tunisia', 'Panama'],
'H': ['Colombia', 'Japan', 'Senegal', 'Poland']
}
def get_observed_result(team1, team2, d):
""" Conveniently query the above maps
d is the map to query (groups or finals)
Return None if match has not been observed
"""
observed_result = None
if (team1, team2) in d:
observed_result = d[(team1, team2)]
elif (team2, team1) in d:
observed_result = d[(team2, team1)]
return observed_result
# -
# ## Some functions to randomly draw match winners
# Some function to simulate match winners, for both pool matches (where draw is allowed) and final rounds (where there is no draw).
# +
def play_pool_match(team1, team2):
""" Returns a dict containing a map from the team to the score it obtained (draw possible)
"""
# first, check if match has already been played, in which case return observed result
obs_winner = get_observed_result(team1, team2, observed_results_groups)
if obs_winner is not None:
if obs_winner == 'draw':
return {team1: 1 + 0.01 * kickscores[team1], team2: 1 + 0.01 * kickscores[team2]}
else:
loosing_team = team1 if obs_winner == team2 else team2
return {obs_winner: 3 + 0.01 * kickscores[obs_winner], loosing_team: 0 + 0.01 * kickscores[loosing_team]}
# Match has not been observed; retrieve correct probabilities from know pool-match probabilities
if (team1, team2) in pools_probas:
pr_team1, pr_draw, pr_team2 = pools_probas[(team1, team2)]
elif (team2, team1) in pools_probas:
pr_team2, pr_draw, pr_team1 = pools_probas[(team2, team1)]
else:
print('Unknown pool match! ({}, {})'.format(team1, team2))
# put them in convenient form for randomly drawing score
team_probas = sorted([(team1, pr_team1), ('draw', pr_draw), (team2, pr_team2)], key=lambda t: -t[1])
# decide result
prn = 100 * random()
for team, pr in team_probas:
if prn <= pr:
# "team" won
if team == 'draw':
# add a small salt proportional to kickscore, to later break ties
# note, this is not really ideal; we should randomly draw tie using logistic function instead
return {team1: 1 + 0.01 * kickscores[team1], team2: 1 + 0.01 * kickscores[team2]}
else:
loosing_team = team1 if team == team2 else team2
return {team: 3 + 0.01 * kickscores[team], loosing_team: 0 + 0.01 * kickscores[loosing_team]}
else:
prn -= pr
def logistic_fun(x):
""" x is a kickscore difference
"""
s = 1.5 # a scale parameter, more or less manually fitted/guessed
return 1. / (1. + math.exp(-x * s))
def play_match(team1, team2):
""" Returns the winner (no draw)
"""
# First, check whether the match has already been observed
observed_winner = get_observed_result(team1, team2, observed_results_finals)
if observed_winner is not None:
return observed_winner
# The match has not been played; simulate outcome
kickscore_1 = kickscores[team1]
kickscore_2 = kickscores[team2]
is_1_fav = kickscore_1 > kickscore_2
pr = logistic_fun(abs(kickscore_1 - kickscore_2))
prn = random()
if prn <= pr:
# The favorite wins
return team1 if is_1_fav else team2
return team2 if is_1_fav else team1
def play_group(group):
""" Returns a list of the group teams, ordered from 1st to 4th
"""
# First, check whether we already have an observed ordering for the group:
if group in observed_groups_orderings:
return observed_groups_orderings[group]
# We don't have an ordering; simulate it
group_teams = groups[group]
already_played = set()
team_to_pts = defaultdict(int)
for team1 in group_teams:
for team2 in group_teams:
if team1 == team2 or (team1, team2) in already_played:
continue
# play the match
scores = play_pool_match(team1, team2)
# keep track of sums of scores
for team, pts in scores.items():
team_to_pts[team] += pts
already_played.add((team1, team2))
already_played.add((team2, team1))
sorted_teams = sorted(list(team_to_pts.items()), key=lambda t: -t[1])
return list(map(lambda t: t[0], sorted_teams))
# -
# ## A function to play all matchs in a final:
def play_final(matchs, prev_winners, counts_to_update, is_8th=False):
""" Returns a map <match_id --> winner>
"""
finals_to_winner = dict()
for match in matchs.keys():
advs = matchs[match]
# pick the actual teams who won the pools
if is_8th:
# in this case prev_winners values are lists of teams
team1 = prev_winners[advs[0][1]][advs[0][0] - 1]
team2 = prev_winners[advs[1][1]][advs[1][0] - 1]
else:
team1 = prev_winners[advs[0]]
team2 = prev_winners[advs[1]]
winner = play_match(team1, team2)
counts_to_update[winner] += 1
finals_to_winner[match] = winner
return finals_to_winner
# ## The actual simulation:
# Here we simulate the WC for `nr_runs`. For each category, we store the number of times each team falls in this category. In addition, we also store the realisations, so as to be able to obtain points distributions or particular bets later on.
# +
nr_runs = 100000
# names of the categories that interest us
categories = ['8th', '4th', 'semi', 'final', 'winner', 'second', 'third']
# counts number of times we observe each of the teams in each position
# (category_name --> (team --> count))
counts = { name: defaultdict(int) for name in categories }
# keep track of the realisations
# [(category_name -> set(teams_in_this_category)), ...]
realisations = []
for run in tqdm(range(nr_runs)):
""" Play the pools
"""
group_2_teams_sorted = dict()
for group in groups.keys():
teams_sorted = play_group(group)
counts['8th'][teams_sorted[0]] += 1
counts['8th'][teams_sorted[1]] += 1
group_2_teams_sorted[group] = teams_sorted
""" Play the finals
"""
finals_8_to_winner = play_final(finals_8, group_2_teams_sorted, counts['4th'], is_8th=True)
finals_4_to_winner = play_final(finals_4, finals_8_to_winner, counts['semi'])
finals_2_to_winner = play_final(finals_2, finals_4_to_winner, counts['final'])
final_to_winner = play_final(final, finals_2_to_winner, counts['winner'])
# store score of 2nd
winner = final_to_winner['1A']
second = finals_2_to_winner['2A'] if finals_2_to_winner['2A'] != winner else finals_2_to_winner['2B']
counts['second'][second] += 1
# play match for 3rd place
semi_final_winners = set(finals_2_to_winner.values())
semi_final_loosers = list(set(finals_4_to_winner.values()) - semi_final_winners)
third = play_match(semi_final_loosers[0], semi_final_loosers[1])
counts['third'][third] += 1
""" Store the realisation
"""
teams_in_8th = set()
for group in groups.keys():
teams_sorted = group_2_teams_sorted[group]
teams_in_8th.add(teams_sorted[0])
teams_in_8th.add(teams_sorted[1])
real = {
'8th': teams_in_8th,
'4th': set(finals_8_to_winner.values()),
'semi': set(finals_4_to_winner.values()),
'final': set(finals_2_to_winner.values()),
'winner': set([winner]),
'second': set([second]),
'third': set([third])
}
realisations.append(real)
# -
""" Display an example of a realisation:
"""
realisations[0]
# ## Compute and display the probabilities to fall in each category for each team:
# Here, for each category, we compute and display the probability that each team falls in this category (over all simulation runs). Use the Juypter menu Cell --> current outputs --> toggle scrolling to see all plots without scrolling within the cell output.
# +
probas = dict() # category_name --> (team --> proba)
for cat in categories:
probas[cat] = dict()
for team, nr in counts[cat].items():
probas[cat][team] = nr / nr_runs
for cat in categories:
teams_probas = sorted(list(probas[cat].items()), key=lambda t: -t[1])
print('------- {} --------'.format(cat))
print(list(map(lambda t: (t[0], 100*t[1]), teams_probas)))
teams_probas_unz = list(zip(*teams_probas))
plt.figure(figsize=(12, 7))
plt.bar(range(len(teams_probas)), list(map(lambda v: v*100, teams_probas_unz[1])), tick_label=teams_probas_unz[0])
plt.xticks(rotation=90)
plt.ylabel('Probability (%)')
plt.title(cat)
# -
# # Evaluating bets
# In this cell, we "specify" a simple betting system as follows. For each category, we have to predict a number of teams falling in this category `category_to_nr_teams`, as well as the number of points earned for each correct such prediction `category_to_pts`.
#
# Once this is defined, we write a function that takes a "prediction" (i.e., a map from category to set of teams predicted to fall in this category), and returns the distribution of points obtained with this prediction on all the simulation runs.
# +
category_to_nr_teams = {
'8th': 16,
'4th': 8,
'semi': 4,
'final': 2,
'winner': 1,
'second': 1,
'third': 1
}
""" Number of points earned for each team correctly placed in the category
"""
category_to_pts = {
'8th': 8,
'4th': 13,
'semi': 22,
'final': 33,
'winner': 68,
'second': 18,
'third': 14
}
def points_for_preds(predictions):
""" [predictions] is a map from category name to set of team in this category.
E.g., 'second' -> set('Belgium')
Returns a list of all scores obtained over each simulation run
"""
assert(all(len(t[1]) == category_to_nr_teams[t[0]] for t in predictions.items())) # make sure predictions are valid
all_points = []
for real in realisations:
pts = 0
for cat in categories:
for team_predicted in predictions[cat]:
pts += category_to_pts[cat] if team_predicted in real[cat] else 0
all_points.append(pts)
return all_points
# -
# ## Expectation-maximizing bet
# Here, we compute the expectation-maximizing prediction, and evaluate the number of points we can get with it.
# +
""" Compute the expectation-maximizing prediction
"""
# compute expectation-maximizing predictions
def get_maxe_preds(probas, nr_teams):
return set(map(lambda t: t[0], sorted(list(probas.items()), key=lambda t: -t[1])[:nr_teams]))
maxe_preds = dict()
for cat in categories:
maxe_preds[cat] = get_maxe_preds(probas[cat], category_to_nr_teams[cat])
pp = pprint.PrettyPrinter(indent=2)
print('Expectation-maximizing prediction:')
pp.pprint(maxe_preds)
maxe_points = points_for_preds(maxe_preds)
print('Expected number of points with this prediction:', np.mean(maxe_points))
# -
# ## Plot the distribution of points obtained by a prediction
# +
def plot_prediction_points(prediction, label, cumulative=False):
points = points_for_preds(prediction)
plt.hist(points, bins=100 if cumulative else 50,
normed=cumulative, cumulative=cumulative, histtype='step', lw=2, label=label)
plt.xlabel('points')
plt.ylabel('probability to obtain less' if cumulative else 'count')
plt.legend(loc='upper center', bbox_to_anchor=(0.5, 1.2))
plt.figure(figsize=(10, 6))
plot_prediction_points(maxe_preds, 'Distribution of points for expectation-maximizing prediction')
plt.show()
# -
# ## Exploration of hand-made predictions
# Define my own prediction (at the beginning of tournament), and compare it to the expectation-maximizing one (updated dynamically). Can we for instance find something that limits risk; i.e., is less likely to bring low number of points? Note that the best predictions (in terms of average number of points) need not be consistent with any one realisation (e.g., one team could be the most likely team to be both winner and second).
# +
my_pred = {
'8th': { 'Argentina',
'Belgium',
'Brazil',
'Colombia',
'Croatia',
'England',
'France',
'Germany',
'Mexico',
'Poland',
'Portugal',
'Russia',
'Spain',
'Sweden',
'Switzerland',
'Uruguay' },
'4th': { 'Argentina',
'Belgium',
'Brazil',
'England',
'France',
'Germany',
'Portugal',
'Spain'},
'semi': {'Germany', 'Brazil', 'Spain', 'England'},
'final': {'Brazil', 'Spain'},
'winner': {'Brazil'},
'second': {'Brazil'},
'third': {'Germany'}
}
print('Expected number of points =', np.mean(points_for_preds(my_pred)))
plt.figure(figsize=(10, 6))
plot_prediction_points(my_pred, 'my prediction', cumulative=True)
plot_prediction_points(maxe_preds, 'Expectation-maximizing prediction', cumulative=True)
plt.show()
# -
|
wc2018-phases-predictions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + init_cell=true
# %logstop
# %logstart -rtq ~/.logs/ML_Time_Series.py append
# %matplotlib inline
import matplotlib
import seaborn as sns
sns.set()
matplotlib.rcParams['figure.dpi'] = 144
# -
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# # Time Series
# <!-- requirement: images/time_series_CV.png -->
# <!-- requirement: data/arima_model.pkl -->
# <!-- requirement: data/co2_weekly_mlo.txt -->
#
# A time series is a sequence of measurements of a variable made over time. The usually application of machine learning to a time series is to use past behavior to make forecasts. Since the time series are usually continuous values, forecasting is a supervised regression problem. Time series differ from the "standard" regression problems studied earlier because observations are _usually_ not independent and the only piece of data we have is the signal itself. We want to take advantage of the temporal nature of the data without the knowledge of the forces that caused those values. The general approach when working with a time series is to
#
# 1. Plot the time series; notice any overall trends and seasonality.
# 1. Detrend the time series by removing drift and seasonality.
# 1. Fit a baseline model and calculate the residuals.
# 1. Analyze the resulting residuals and generate features from the residuals.
# 1. Train a machine learning model to forecast/predict residuals and add back the baseline model.
#
# For this notebook, we will be analyzing the atmospheric carbon dioxide levels measured from the Mauna Loa Observatory in Hawaii. More information about the data can be found [here](https://www.esrl.noaa.gov/gmd/ccgg/trends/data.html).
# ## Components of a time series
#
# We can model our time series as having three components,
#
# $$ y(t) = \mathrm{drift} + \mathrm{seasonal} + \mathrm{noise}. $$
#
# The components are defined as
#
# 1. **Drift**: An overall trend present in the time series. An example of a drift model is
# $$ y(t) = \mu t. $$
# Other commonly applied drift models are quadratic and exponential.
#
# 1. **Seasonality**: A periodic behavior existing in the time series. For a given frequency $f$, a common model is
# $$ y(t) = A\sin(2\pi ft) + B\cos(2\pi ft). $$
#
# 1. **Noise**: The part of the time series remaining after removing drift and seasonality. It is the residual of a model containing drift and seasonality.
#
# Our approach will be to identify the first two terms to create a baseline model, leaving behind the residuals or noise. This [link](https://people.duke.edu/~rnau/whatuse.htm) provides a list of different transformations that are commonly applied when analyzing time series.
#
# ** Questions**
# * What are some examples of drift in real time series?
# * What are some examples of seasonality in real time series?
# ## Cross-validation of time series data
#
# Since observations are not independent and we want to use past data to predict future values, we need to apply slightly different approach when training and testing a machine learning model. Given the temporal nature of the data, we need to preserve order and have the training set occur prior to the test set. For cross-validation, two common methods are used, sliding and forward chaining.
#
# * **Sliding Window**: The model is trained with data in a fixed window size and tested with data in the following window of the same size. Then the window _slides_ where the previous test data becomes the training data and repeated for the number of chosen folds.
#
# * **Forward Chaining**: The model is _initially_ trained/tested with windows of the same size as the sliding window method. However, for each subsequent fold, the training window increases in size, encompassing both the previous training data and test data. The new test window once again follows the training window but stays the same length.
#
# 
# In `scikit-learn`, the forward chaining method is available in `sklearn.model_selection.TimeSeriesSplit`. See below for an example of using forward chaining with `GridSearchCV`. See this [link](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.TimeSeriesSplit.html) for more info on the usage.
# +
from sklearn.linear_model import Ridge
from sklearn.model_selection import GridSearchCV, TimeSeriesSplit
regressor = Ridge()
param_grid = {"alpha": np.logspace(-2, 2, 100)}
ts_cv = TimeSeriesSplit(5) # 5-fold forward chaining
grid_search = GridSearchCV(regressor, param_grid, cv=ts_cv, n_jobs=2)
# -
# ## Stationary signal
#
# Ideally, the resulting time series of the residuals will be **stationary**. A stationary signal or process is one in which statistical values such as the mean do not change with time. For our purposes, we are concerned about the special case where the mean, variance, and autocorrelation (explained more later) are not a function of time. This special case is called weakly stationary. Transforming a time series into a stationary process is crucial for time series analysis because a large number of analysis tools assume the process is stationary. It is easy to predict future values if things like the mean and variance stay the same with time. Consider a time series where new values are dependent on the past time series value and a random, uncorrelated noise $\epsilon_t$.
#
# $$
# y_t = \rho y_{t-1} + \epsilon_t.
# $$
#
# The parameter $\rho$ scales the contribution of the past value. If $\epsilon_t$ is uncorrelated and has mean zero, it is referred to as **white noise**. If the values are sampled from a normal distribution, the white noise is then called **white Gaussian noise**. The following visualization allows you to scale the contribution of past values by adjusting $\rho$. Notice the signal is stationary when $\rho < 1$ but is no longer stationary when $\rho=1$.
# +
from ipywidgets import interact, FloatSlider
def plot_signal(rho=0):
n = 1000
np.random.seed(0)
eps = np.random.randn(n)
y = np.zeros(n)
y[0] = eps[0]
var = np.zeros(n)
for i in range(1, n):
y[i] = rho*y[i-1] + eps[i]
var[i] = y[:i].var()
plt.subplot(211)
plt.plot(y)
plt.ylabel('y')
plt.subplot(212)
plt.plot(var)
plt.ylabel('$\sigma_y$')
plt.plot(var)
interact(plot_signal, rho=FloatSlider(min=0, max=1, value=0, step=0.01, description='$\\rho$'));
# -
# The case when $\rho=1$ is called a one-dimensional [random walk](https://en.wikipedia.org/wiki/Random_walk). A random walk is a stochastic/random process that describes the location of an object from successive random steps, random in both direction and size. The equation $y_t = y_{t-1} + \epsilon_t$ is a random walk because the position at time $t$ is some random distance from the previous location $y_{t-1}$. There have been extensive research on random walk processes since they occur in a wide range of subjects, from financial models to particle diffusion. There are two main consequences of having residuals as white noise.
#
# 1. You cannot predict/forecast future values because what is left is uncorrelated noise.
# 1. You have an adequate time series model since there is no signal left to model.
# ## Modeling Drift
#
# Let's load the atmospheric CO2 data set using pandas and plot the time series. The data set has weekly measurements but there are some missing values, denoted by `-999.99`. We will need to replace those missing values and create timestamps from the date info.
# +
# load data set
columns = ['year', 'month', 'day', 'decimal date', 'molfrac', 'days', '1 yr ago', '10 yrs ago', 'since 1880']
df = pd.read_csv('data/co2_weekly_mlo.txt', sep='\s+', header=None, names=columns, na_values=-999.99)
# create timestamp indices
df['date'] = pd.to_datetime(df[['year', 'month', 'day']])
df = df.set_index('decimal date')
# replace missing values
df['molfrac'] = df['molfrac'].fillna(method='ffill')
df.head()
# -
CO2 = df['molfrac']
CO2.plot()
plt.ylabel('CO2 ppm');
# **Questions**
#
# * What are some behaviors do you observe in the time series?
# * What model would you pose to remove the drift?
# Atmospheric CO2 levels have been consisting increasing at a slightly superlinear fashion. While the choice of drift is somewhat subjective, we will use a quadratic fit. The quadratic features will be provided from the `PolynomialFeatures` transformer. Let's create a simple model that only captures the drift; we will worry about the seasonality later. We will perform a train/test split at the year 2010 and define some functions and classes to help with the analysis.
# +
from sklearn.base import BaseEstimator, TransformerMixin
class IndexSelector(BaseEstimator, TransformerMixin):
def __init__(self):
"""Return indices of a data frame for use in other estimators."""
pass
def fit(self, df, y=None):
return self
def transform(self, df):
indices = df.index
return indices.values.reshape(-1, 1)
# +
def ts_train_test_split(df, cutoff, target):
"""Perform a train/test split on a data frame based on a cutoff date."""
ind = df.index < cutoff
df_train = df.loc[ind]
df_test = df.loc[~ind]
y_train = df.loc[ind, target]
y_test = df.loc[~ind, target]
return df_train, df_test, y_train, y_test
def plot_results(df, y_pred):
"""Plot predicted results and residuals."""
CO2.plot();
plt.plot(list(df.index), y_pred, '-r');
plt.xlabel('year')
plt.ylabel('CO2 ppm')
plt.legend(['true', 'predicted']);
plt.show();
plt.plot(resd)
plt.xlabel('year')
plt.ylabel('residual')
# +
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import PolynomialFeatures
# perform train/test split
cutoff = 2010
df_train, df_test, y_train, y_test = ts_train_test_split(df, cutoff, 'molfrac')
# construct and train pipeline
time = IndexSelector()
poly = PolynomialFeatures(degree=2)
lr = LinearRegression()
pipe = Pipeline([('indices', time),
('drift', poly),
('regressor', lr)])
pipe.fit(df_train, y_train)
# make predictions
y_pred = pipe.predict(df)
resd = CO2 - y_pred
print("Test set R^2: {:g}".format(pipe.score(df_test, y_test)))
plot_results(df, y_pred)
# -
# The residuals exhibit a periodic behavior; our next task is to remove the seasonal component of our data set. Atmospheric CO2 levels have a yearly cyclic behavior due to seasonal variations in the uptake of CO2 by vegetation. In this case, we may already know about this seasonal pattern, however, we need a systematic way to determine the dominant periodic behaviors in a time series.
# ## Modeling Seasonality
#
# Any signal can be represented as a linear superposition of sines and cosines of varying frequencies $f_n$ and amplitudes $A_n$ and $B_n$,
#
# $$ y(t) = \sum_n \left(A_n \sin(2\pi f_n t) + B_n\cos(2 \pi f_n t) \right). $$
#
# The **Fourier transform** decomposes a signal into a set of frequencies, allowing for us to determine the dominant frequencies that make up a time series. We are transforming our signal in the time domain into the frequency domain. Since we will be working with discrete data; the signal is sampled at discrete points in time, we will use the **discrete Fourier transform**. For $N$ uniformly sampled time series $y_n$, the transform is defined as
#
# $$ Y_k = \sum^{N-1}_{n=0} y_n e^{-\frac{2\pi i}{N} kn}, $$
#
# $$ Y_k = \sum^{N-1}_{n=0} y_n \left[\cos\left(\frac{2\pi i}{N} kn\right) - i\sin\left(\frac{2\pi i}{N} kn\right) \right], $$
#
# and $i$ is the imaginary number. The term $Y_k$ is the Fourier transform value for a frequency of $k$ cycles in $N$ samples; it is a complex number that represents both the amplitude and phase for its respective sinusoidal component. The amplitude for the frequency $k/N$ is
#
# $$ |Y_k|/N = \frac{\sqrt{\mathrm{Re}(Y_k)^2 + \mathrm{Im}(Y_k)^2}}{N}. $$
#
# The most common algorithm used to compute the discrete Fourier transform is the fast Fourier transform (FFT). The algorithm makes use of matrix factorization to have a time complexity of $O(n\log n)$ as opposed to the naive $O(n^2)$ implementation. Note, the time series needs to be uniformly sampled. The `scipy.fftpack` provides the FFT algorithm. Let's use FFT to determine the contributed frequencies in the signal below.
# +
from scipy import fftpack
def fft_plot(a=1, b=1, c=1, fourier=True):
np.random.seed(0)
N = 100
t_end = 4
t = np.linspace(0, t_end, N)
y = a*np.cos(2*np.pi*t) + b*np.sin(4*2*np.pi*t) + c*np.cos(8*2*np.pi*t) + 0.2*np.random.randn(N)
Y = fftpack.fft(y)
f = np.linspace(0, N, N)/t_end
if fourier:
plt.subplot(211)
plt.plot(t, y)
plt.xlim([0, 4])
plt.ylim([-4, 4])
plt.xlabel('time')
plt.subplot(212)
plt.plot(f, np.abs(Y)/len(Y))
plt.ylim([0, 2])
plt.xlabel('number of cyles in full window')
plt.tight_layout()
else:
plt.plot(t, y)
plt.xlim([0, 4])
plt.ylim([-4, 4])
plt.xlabel('time')
fft_plot(a=1, b=1, c=1, fourier=False);
# -
# From visual inspection, it is not apparent the frequencies that contribute to the signal but is derived from
#
# $$ y(t) = a\cos(2\pi t) + b\sin(8\pi t) + c\cos(16\pi t) + \epsilon(t). $$
#
# The signal is composed of three sines/cosines at frequencies of 1, 4, and 8 and random uncorrelated noise $\epsilon(t)$. The signal spans 4 time units and is sampled 25 times per unit of time. In the interactive visualization below, we display the signal and the resulting Fourier transform, allowing for the change in the amplitude of each of the three sinusoidal terms.
# +
from ipywidgets import interact
interact(fft_plot, a=(0, 4, 0.1), b=(0, 4, 0.1), c=(0, 4, 0.1));
# -
# One interpretation of the Fourier transform plot is that it is a histogram/distribution of the frequencies that contribute to the signal. The resulting graph has three peaks; each peak corresponds to a dominant frequency present in the signal. Notice how increasing the amplitude of one of the sinusoidal terms in the signal results in a larger value for the respective frequency in the Fourier transform plot.
#
# The $x$-axis represents frequency, where the smallest non-zero frequency is equal to $1/t_{span}$ where $t_{span}$ is the size of the window or duration of the time series. The highest frequency is the inverse of the sampling rate.
#
# **Questions**
# * Are there any interesting features in the plot of the Fourier transform?
# * What would happen if the magnitude of the noise increases? Would it be difficult to derive insight form the decomposed signal?
# A Fourier transform of a real signal, no imaginary part, is symmetric about the center of the frequency range. The symmetric part is a result of _aliasing_, the effect of not differentiating two signals from each other. The discrete Fourier transform cannot measure the contribution of frequencies greater than the half of the inverse of the sampling rate, referred to as the Nyquist frequency,
#
# $$ f_N = \frac{1}{2\Delta t}, $$
#
# where $\Delta t$ is the sampling rate. In the visualization below, we display the sampled values of two signals, one below the Nyquist frequency and its higher frequency alias. The different signals have the same sampled values but are derived from different frequencies. Notice how the signal in green is too fast to properly measure with our sampling rate. During the time before the signal is sampled again, the curve has gone up (or down) and down (or up) and reached its max (or min) value. Given our sampling frequency, we cannot distinguish sampled data from the green curve from that of the blue curve, they are aliases of one another. Because of the aliasing effect, it is customary to only display the Fourier transform for frequencies less than the Nyquist, only the first half of the plot.
# +
from ipywidgets import FloatSlider
def plot_alias(f=0.2, blue=True, green=True):
t = np.linspace(0, 10, 500)
t_sampled = np.arange(0, 11)
if blue:
plt.plot(t, np.sin(2*np.pi*f*t), 'b')
if green:
plt.plot(t, -np.sin(2*np.pi*(1-f)*t), 'g')
l, m, b = plt.stem(t_sampled,
np.sin(2*np.pi*f*t_sampled),
linefmt='r',
markerfmt='ro',
use_line_collection=True)
plt.setp(b, visible=False)
plt.ylim(-2, 2)
plt.xticks(t_sampled)
plt.legend(["f={}".format(f), "f={}".format(1-f), "sampled signal"])
interact(plot_alias, f=FloatSlider(min=0, max=1.0, step=0.05, value=0.05, description='$f$'));
# -
# For the atmospheric CO2 data, let's formally identify the most dominant frequencies. We subtract the mean before computing the Fourier transform. If not, there would be a large value at zero frequency. The Fourier transform of the residuals is plotted below.
# +
Y = fftpack.fft(resd-resd.mean())
t_span = CO2.index[-1] - CO2.index[0]
f = np.linspace(0, len(Y), len(Y))/t_span
plt.plot(f[:len(Y)//2], np.abs(Y[:len(Y)//2])/len(Y));
plt.xlabel('frequency (1/yr)')
plt.ylabel('amplitude');
# -
# It appears that there are no dominant frequencies greater than fives times a year. Let's zoom in for further inspection.
plt.plot(f[:200], np.abs(Y)[:200]);
plt.xlabel('frequency (1/yr)')
plt.ylabel('amplitude');
# We see two dominant frequencies occurring at once and twice a year. Our updated baseline model is now
#
# $$ y(t) = A + Bt + Ct^2 + D\sin(2\pi t) + E\cos(2\pi t) + F\sin(4\pi t) + G\cos(4\pi t), $$
#
# where $t$ in expressed in units of years. To incorporate the seasonal components, we will construct a custom transformer and use a combination of pipelines and feature unions to construct our baseline model.
# +
from sklearn.base import BaseEstimator, TransformerMixin
class FourierComponents(BaseEstimator, TransformerMixin):
def __init__(self, freqs):
"""Create features based on sin(2*pi*f*t) and cos(2*pi*f*t)."""
self.freqs = freqs
def fit(self, X, y=None):
return self
def transform(self, X):
Xt = np.zeros((X.shape[0], 2*len(self.freqs)))
for i, f in enumerate(self.freqs):
Xt[:, 2*i]= np.cos(2*np.pi*f*X).reshape(-1)
Xt[:, 2*i + 1] = np.sin(2*np.pi*f*X).reshape(-1)
return Xt
# +
from sklearn.pipeline import FeatureUnion
# construct and train pipeline
fourier = FourierComponents([1, 2]) # annual and biannual frequencies
union = FeatureUnion([('drift', poly), ('fourier', fourier)])
baseline = Pipeline([('indices', time),
('union', union),
('regressor', lr)])
baseline.fit(df_train, y_train)
# make predictions
y_pred = baseline.predict(df)
resd = CO2 - y_pred
print("Test set R^2: {:g}".format(baseline.score(df_test, y_test)))
plot_results(df, y_pred)
# -
# At the moment, we have a baseline model that works well but the residuals do not appear to be completely stationary. Our analysis is not done, we can focus our attention on extracting any patterns in the resulting correlated noise.
# **Questions**
# * What, if any, behavior do you observe in the current baseline model's residuals?
# * Instead of using $y(t) = A\cos(2\pi ft) + B\sin(2\pi t)$, we could have used the equivalent $y(t) = k\sin(2\pi ft - \phi). $ Why would the former be preferred?
# ## Modeling noise
#
# We can improve on our analysis by modeling the noise, the residuals of our baseline model. Specifically, we want to measure the persistence of past values on the signal. In other words, past values of our times series are correlated to current values. We expect there to be some correlation to past values but the persistence should die off for further values in the past. The **autocorrelation** will give us a measure the persistence of past values; it is a measure of how well correlated a signal is with a lag copy of itself. Let's define some important mathematical values that are crucial for understanding the autocorrelation.
#
# * **Covariance**: A measure of _joint_ variability of two variables,
# $$ \mathrm{cov}(X, Y) = E[(X - E[X])(Y - E[Y])] = \frac{1}{N} \sum^{n}_{i=1}(x_i - E[X])(y_i - E[Y]). $$
#
# * **Variance**: A measure of the variability of a variable with _itself_; the special case of the covariance,
# $$ \mathrm{var}(X) = \mathrm{cov}(X, X) = E[(X - E[X])^2] = \frac{1}{N} \sum^{n}_{i=1}(x_i - E[X])^2. $$
#
# * **Standard Deviation**: The square root of the variance,
# $$ \sigma_X = \sqrt{\mathrm{var}(X)}. $$
#
# * **Correlation**: The normalized covariance that ranges from -1 to 1,
# $$\rho(X, Y) = \frac{\mathrm{cov}(X, Y)}{\sigma_X \sigma_Y}. $$
#
# Three important values and meanings of the correlation coefficient are:
#
# 1. If $\rho(X, Y) = 1$, then the two variables are completely linear correlated; an increase in one corresponds to a linear increase of the other.
# 1. If $\rho(X, Y) = 0$, then the two variables are uncorrelated. Higher values of one variable does not necessarily correspond to higher or lower values of the other.
# 1. If $\rho(X, Y) = -1$, then the two variables are completely linear anti-correlated; an increase in one corresponds to a linear decrease of the other.
#
# With the correlation coefficient, we can now mathematically define and better understand the autocorrelation of a signal. The autocorrelation as a function of the duration of the lag is defined as
#
# $$ R(\tau) = \frac{\mathrm{cov}(y(t), y(t-\tau))}{\sigma_{y} \sigma_{y}} = \frac{\gamma(\tau)}{\sigma^2_{y}} = \rho(y(t), y(t-\tau)), $$
#
# where $\tau$ is the duration of the lag/delay and $\gamma$ is the autocovariance function. Since we are working with discrete data, we can define the lag with respect to the number of time steps $k$,
#
# $$ R(k) = \frac{\gamma(k)}{\sigma_{y}^2} = \rho(y_t, y_{t-k}). $$
#
# Since the autocorrelation is a measure of how correlated a signal is with a delayed copy of itself, plotting the autocorrelation function will reveal to us how correlated past values are. The pandas function `autocorrelation_plot` plots the autocorrelation function of the curve and includes 95% and 99% confidence values of the zero-correlation hypothesis. The point of interest in the curve is at what lag value is there no more correlation. Such value is the characteristic time scale of the process.
# +
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(resd)
plt.xlabel('Lag (weeks)')
plt.xlim([0, 1000]);
# -
# It appears that values past 400 to 500 weeks are not correlated with current values.
# ## Noise based feature generation
#
# With the noise/residual of our time series, we can generate features based on past values for each time step. These features can be
#
# * Statistics of a window of past values, such as the mean and max.
# * One hot encoded features based on things such as the days of the week and holidays.
# * External features for each time step, for example, the value of the stock market.
#
# After determining the characteristic time scale of our process, we can incorporate the time scale when deciding how to best generate features. A common statistic to calculate is the moving average. For a times series, the moving average of a point in time is some average value calculated using a subset of past values. There are different types of moving averages but two common ones are:
#
# * **Rolling Window Average**: The average is calculated for a window of $k$ previous points.
#
# $$ MA_t = \frac{1}{k} \sum^{n}_{n-k} y_k. $$
#
# * **Exponential Moving Average**: All points are included in calculating the average but are weighted using an exponential decay. In other words, values further in the past contribute less to the moving average than recent points. A nice property of the exponential moving average is that the moving average value can be calculated with only the current time series value and the previous exponential moving average value.
#
# $$ EMA_t = \alpha y_t + (1 - \alpha) EMA_{t-1}, $$
#
# where $\alpha$ ranges from 0 to 1 and scales the strength of the contribution of past values. The value of $\alpha$ is related to the half-life of the weights, the time for the weights to drop half of their value,
#
# $$ \alpha = 1 - \exp\left[-\frac{\ln(2)}{t_{1/2}}\right], $$
#
# where $t_{1/2}$ is the half-life. Note, while we have discussed rolling window and exponential moving _averages_, other values can be calculated for other statistics.
#
# In the visualizations below, you can control the window size and half-life of the rolling window and exponentially weighted average of the residuals. Notice how applying moving averages smooths out the residuals. These moving averages are sometimes used to smooth out data.
# +
def plot_rolling_window(window=10):
series = pd.Series(resd, index=df.index)
rolling_window = series.rolling(window=window).mean()
series.plot(alpha=0.5)
rolling_window.plot(linewidth=2, color='k')
plt.title('rolling window')
plt.xlabel('year')
plt.ylabel('moving average')
interact(plot_rolling_window, window=(1, 200, 1));
# +
def plot_exponential_weighted(half_life=100):
series = pd.Series(resd, index=df.index)
exponential_weighted = series.ewm(halflife=half_life).mean()
series.plot(alpha=0.5)
exponential_weighted.plot(linewidth=2, color='k')
plt.title('exponential weighted')
plt.xlabel('year')
plt.ylabel('moving average')
half_life_slider = FloatSlider(min=1, max=100, step=0.1, value=10, description="half-life")
interact(plot_exponential_weighted, half_life=half_life_slider);
# -
# **Questions**
# * How does one determine a good value to use for window size or half-life?
# * Considering computer memory, what moving average is better to use, rolling window average or exponential moving average?
# * How does increasing the half-life affect $\alpha$?
# With a baseline model and resulting residuals, our goal is to construct a model to predict atmospheric CO2 levels 20 weeks into the future. In other words, given the time series values we have currently measured at time $t$, we want to predict or forecast the value of the time series 20 time steps into the future since we sample the data weekly. For our approach, we will use the current time step residual, the prior residual, the rolling mean of the residual, and the rolling mean of the difference of the residual to predict the residual 20 weeks later. For the rolling window, we will choose a window size of 100 weeks.
class ResidualFeatures(BaseEstimator, TransformerMixin):
def __init__(self, window=100):
"""Generate features based on window statistics of past noise/residuals."""
self.window = window
def fit(self, X, y=None):
return self
def transform(self, X):
df = pd.DataFrame()
df['residual'] = pd.Series(X, index=X.index)
df['prior'] = df['residual'].shift(1)
df['mean'] = df['residual'].rolling(window=self.window).mean()
df['diff'] = df['residual'].diff().rolling(window=self.window).mean()
df = df.fillna(method='bfill')
return df
# +
from sklearn.metrics import r2_score
# create and train residual model
resd_train = y_train - baseline.predict(df_train)
residual_feats = ResidualFeatures(window=100)
residual_model = Pipeline([('residual_features', residual_feats), ('regressor', LinearRegression())])
residual_model.fit(resd_train.iloc[:-20], resd_train.shift(-20).dropna())
# evaluate model
resd_pred = residual_model.predict(resd) # prediction for all time steps
resd_pred = pd.Series(resd_pred, index=df.index)
resd_pred = resd_pred.shift(20).dropna() # shift predicted values to matching time step
resd_pred_test = resd_pred.loc[resd_pred.index > 2010] # evaluate only on 2010 values
print("Residual test set R^2: {:g}".format(r2_score(resd.loc[resd.index > 2010], resd_pred_test)))
# -
# Now with the residual model, we can combine both the baseline and residual model to make forecasts of atmospheric CO2 levels. It is best to create a custom estimator to encapsulate the process of combining both models.
# +
from sklearn.base import RegressorMixin
class FullModel(BaseEstimator, RegressorMixin):
def __init__(self, baseline, residual_model, steps=20):
"""Combine a baseline and residual model to predict any number of steps in the future."""
self.baseline = baseline
self.residual_model = residual_model
self.steps = steps
def fit(self, X, y):
self.baseline.fit(X, y)
resd = y - self.baseline.predict(X)
self.residual_model.fit(resd.iloc[:-self.steps], resd.shift(-self.steps).dropna())
return self
def predict(self, X):
y_b = pd.Series(self.baseline.predict(X), index=X.index)
resd = X['molfrac'] - y_b
resd_pred = pd.Series(self.residual_model.predict(resd), index=X.index)
resd_pred = resd_pred.shift(self.steps)
y_pred = y_b + resd_pred
return y_pred
# construct and train full model
full_model = FullModel(baseline, residual_model, steps=20)
full_model.fit(df_train, y_train)
# make predictions
y_pred = full_model.predict(df)
resd = CO2 - y_pred
ind = resd.index > 2010
print("Test set R^2: {:g}".format(r2_score(CO2.loc[ind], y_pred.loc[ind])))
plot_results(df, y_pred)
# -
# Our final model works really well at making predictions 20 weeks into the future. Let's plot the histogram and autocorrelation of the final residuals.
# +
from scipy.stats import norm
mu = resd.mean()
sigma = resd.std(ddof=1)
dist = norm(mu, sigma)
x = np.linspace(-2, 2, 100)
f = dist.pdf(x)
resd.hist(bins=40, density=True)
plt.plot(x, f, '-r', linewidth=2);
# -
autocorrelation_plot(resd.dropna())
plt.xlim([0, 100]);
# The residuals are Gaussian and while arguably past values are still correlated, they are not as correlated as they were before.
# ## Statistical time series models
#
# There are a class of statistically based models for time series. Most of these models are provided by the `statsmodels` Python package. Unfortunately, the API of models are different than `scikit-learn`. For this section, we will briefly discuss these models and demonstrate their usage in Python.
# ### Autoregressive and moving average models
#
# The autoregressive (AR) model of order $p$ states that the current time series value is linearly dependent on the past $p$ values with some white noise,
#
# $$y_t = c + \alpha_1 y_{t-1} + \alpha_2 y_{t-2} + ... \alpha_p y_{t-p} + \epsilon_t = c + \sum^{p}_{p=1} \alpha_p y_{t-p} + \epsilon_t, $$
#
# where $\alpha_p$ are the model parameters, $y_{t-p}$ are past time series values, $c$ is a constant, and $\epsilon_t$ is white noise. The name autoregressive refers to the model parameters being solved by applying regression with the time series values themselves. Our previous illustration discussing stationary signals is an autoregressive model of order one as the current value is equal to the scaled prior value plus some noise. Autoregressive models are great at capturing the mean reversion and momentum in the time series since it is based on a window of past values.
#
# Another model is the moving average (MA) model. Despite similar names, the MA model and concept of moving averages are different and should not be confused. The MA model of order $q$ says that the time series is linearly dependent on current and past shock values or noise,
#
# $$y_t = c + \epsilon_t + \beta_1 \epsilon_{t-1} + \beta_2 \epsilon_{t-2} + ... \beta_q \epsilon_{t-q} = c + \sum^{q}_{q=1} \beta_q \epsilon_{t-q} + \epsilon_t, $$
#
# where $\beta_q$ are the model parameters. The MA model captures the persisting effect of shock events on future time series values. To get the capabilities of both models, AR and MA models are added, forming a more general time series model referred to as autoregressive and moving average (ARMA) model. The coefficients of the AR models are solved using a variety of methods such as linear least squares regression. MA coefficients are more computationally intensive to solve because shock values are not directly observed, requiring non-linear fitting algorithms. When using ARMA, the order of both AR and MA need to be specified and can be different.
#
# **Question**
# * How should one identify an appropriate value for the order of either AR and MA?
# Let's demonstrate the AR model from `statsmodels` for forecasting the residuals of the baseline model.
type(df_train.date.to_numpy())
# +
from statsmodels.tsa.ar_model import AR
# create and fit AR model
lag = 200
resd_train = y_train - baseline.predict(df_train)
ar = AR(resd_train.values, dates=df_train['date'], freq='W')
ar = ar.fit(maxlag=lag)
resd_ar_train_pred = ar.predict(start=lag, end=len(df_train)-1)
# plot training set results
plt.plot(list(df_train.index), y_train - baseline.predict(df_train), alpha=0.5)
plt.plot(list(df_train.index[lag:]), resd_ar_train_pred, 'r')
plt.xlabel('year');
plt.ylabel('residual')
plt.legend(['true', 'predicted'])
plt.show();
# plot 20 step forecast of test set
steps = 20
resd_ar_test_pred = ar.predict(start=len(df_train), end=len(df_train) + steps - 1)
plt.plot(range(1, steps + 1), y_test.iloc[:steps] - baseline.predict(df_test.iloc[:steps]))
plt.plot(range(1, steps + 1), resd_ar_test_pred)
plt.xlabel('step')
plt.ylabel('residual')
plt.legend(['true', 'predicted']);
# -
# The general syntax for using the models from `statsmodels` is passing the training data when instantiating the model, fitting the model by passing the number of terms to include, and finally calling the `predict` method with the number of steps into the future to forecast. The AR model was able to capture the trends, the ups and downs, of the residuals but under predicted the magnitude of those trends.
# ## ARIMA
#
# The ARMA model only works for a stationary process. One method to arrive at a stationary process is to apply a difference transformation, $\Delta y_t = y_t - y_{t-1}$. In our example of a random walk, the series was not stationary but the time series of the difference is stationary because it only depends on white Gaussian noise. The autoregressive integrated moving average (ARIMA) model is a general form of ARMA that applies differencing to the time series in the hopes of generating a stationary process. The ARIMA model is often written as $\mathrm{ARIMA}(p, d, q)$, where
# * $p$: Number of terms to include in the AR model.
# * $d$: The degree of differencing, how many times differencing is applied to the series.
# * $q$: Number of terms to include in the MA model.
#
# Let's use the ARIMA model provided by the `statsmodels` package on the noise/residual of Mauna Loa data. Since the model takes a long time to fit, we have provided a pickle file of the trained model.
# +
import pickle
from statsmodels.tsa.arima_model import ARIMA
# arima = ARIMA(resd_train.values, order=(20, 1, 5), dates=df_train['date'], freq='W')
# arima = arima.fit()
# load pretrained model
with open('data/arima_model.pkl', 'rb') as f:
arima = pickle.load(f)
# plot 20 step forecast of test set
steps = 20
resd_arima_test_pred, _, _ = arima.forecast(steps)
plt.plot(range(1, steps + 1), resd_arima_test_pred)
plt.xlabel('step')
plt.ylabel('residual');
# -
# ## Exercises
#
# 1. Incorporate more features into the residual model. Consider including more window statistics and external features such as financial data. Measure the performance in both the residual and full model.
# 1. Since we have relatively small number of features, we were not worried about overfitting with the residual model using linear regression. However, overfitting becomes a problem with more features and more complicate models. Chose a different model than linear regression and tune the model's hyperparameters. You may need to use `TimeSeriesSplit` in conjunction with `GridSearchCV` to properly tune the model.
# 1. Use the full model to predict atmospheric CO2 levels for the first 20 weeks of 2019. Check to see how well the model performs once data has been made available.
# 1. Practice using the AR model available in `statsmodels` by generating a time series in the form of $y_t = \rho y_{t-1} + \epsilon_t$. Compare the fitted AR model coefficient(s) to the chosen value of $\rho$. The fitted AR model coefficients are stored in the `params` attribute.
# 1. Using either AR, MA, ARMA, or ARIMA for the residual model when forecasting atmospheric CO2 levels.
# *Copyright © 2020 The Data Incubator. All rights reserved.*
|
13_ML_Time_Series.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''augmenty'': virtualenv)'
# name: python3
# ---
# # Getting Started
#
# [](https://colab.research.google.com/github/kennethenevoldsen/augmenty/blob/master/docs/tutorials/introduction.ipynn)
#
#
#
# Augmenty is an augmentation library based on spaCy for augmenting texts. Augmenty differs from other augmentation libraries in that it corrects (as far as possible) the token, sentence and document labels under the augmentation.
# ## Installation
# Before we get ahead of ourselves let us just install the required packages:
#
# ``` bash
# pip install augmenty
# # install the spacy pipeline
# python -m spacy download en_core_web_sm
# ```
# ## Introduction
# Augmenty is an augmentation library for spaCy, consisting of many different augmenters. To get an idea of all the available augmenters you can always try out the use:
# +
import augmenty
augmenters = augmenty.augmenters()
for augmenter in augmenters:
print(augmenter)
# -
# To get get more information about an individual augmenter you can always simply use `help` for instance let is say you want to know more about the upper case augmenter you could run: `help(augmenters["upper_case.v1"])`.
# After you have an idea about the augmenter you wish to use loading in augmenters in augmenty is made easy using the `load` command given and given the arguments:
upper_case_augmenter = augmenty.load("upper_case.v1", level=1.00) # 100% uppercase
# ## Applying the augmentation
# Augmenters in augmenty always take in a spaCy [Language pipeline](https://spacy.io/api/language) and an spaCy [Example](https://spacy.io/api/example) so that it can be easily used while training workflows, however, augmenty also allows for easy application of augmenters to raw text and spaCy [Docs](https://spacy.io/api/doc).
#
#
# <div class="alert alert-info">
#
# **Why examples and not just raw text?**
#
#
# A spaCy example consist of two documents, the labelled document, containing all the correct labels including document classification such as whether a tweet is positive or negative and token classiification such as Part-of-speech-tags and named entities. When augmenting the Example augmenty seeks to correct these tags in accordance with the augmentation. As the raw text does not include these labels it as naturally not possible. For instance if I was to swap two tokens I would want to swap their corresponding labels as well. When swapping tokens augmenty even respect entities and sentences as to not split an entity or swap tokens across sentence borders. You can naturally turn this of if you wish to.
#
# </div>
#
# ### Applying augmentations on Docs
# +
import spacy
nlp = spacy.load("en_core_web_sm")
docs = nlp.pipe(
[
"Augmentation is a wonderful tool for obtaining higher performance on limited data.",
"You can also use it to see how robust your model is to changes.",
]
)
augmented_docs = augmenty.docs(docs, augmenter=upper_case_augmenter, nlp=nlp)
for doc in augmented_docs:
print(doc)
# -
# ### Applying augmentations on text
# We can also try it out on text. Let us also try out a new augmenter for replacing entities. Remember you can always use `help(augmenters["ents_replace.v1"])` to figure out which inputs the augmenter takes and see and example.
# +
texts = ["Augmenty is a wonderful tool for augmentation."]
ent_augmenter = augmenty.load(
"ents_replace.v1", level=1.00, ent_dict={"ORG": [["SpaCy"], ["The SpaCy Universe"]]}
)
augmented_texts = augmenty.texts(texts, augmenter=ent_augmenter, nlp=nlp)
for text in augmented_texts:
print(text)
# -
# ## Customizing augmenters
# Augmenty is more than a list of augmenters and also contains utilities for dealing with augmenters such as combining and moderating augmenters.
#
# ### Combining augmenters
# We can start of by combing the entity augmenter with an augmenter which replaces words with their synonym based on wordnet.
#
# +
synonym_augmenter = augmenty.load("wordnet_synonym.v1", level=1, lang="en")
combined_aug = augmenty.combine([ent_augmenter, synonym_augmenter])
# +
augmented_texts = augmenty.texts(texts, augmenter=combined_aug, nlp=nlp)
for text in augmented_texts:
print(text)
# -
# ### Moderating Augmenters
# Certain augmenters apply augmentation at different levels. For instance the augmenter `keystroke_error.v1` augments examples based on keyboard distances, where each character has a chance to be replaced with a neightbouring character. However, we might wish to apply this augmentation to 5% of characters, but only apply it 50% of the training samples. Using `augmenty.set_doc_level` we can add this last part to any augmenter, thus allowing for more flexibility when using the model.
# +
keystroke_augmenter = augmenty.load(
"keystroke_error.v1", keyboard="en_qwerty.v1", level=0.05
) # 5% if characters
keystroke_augmenter = augmenty.set_doc_level(
keystroke_augmenter, level=0.5
) # 50% of texts
# +
texts = [
"Augmenty is a wonderful tool for augmentation.",
"Augmentation is a wonderful tool for obtaining higher performance on limited data.",
"You can also use it to see how robust your model is to changes.",
]
augmented_texts = augmenty.texts(texts, augmenter=keystroke_augmenter, nlp=nlp)
for text in augmented_texts:
print(text)
# -
# Similarly one might wish the augment to instead of simply yielding the augmented example also yield the original, such that the trained model always see the actual data.
# +
token_swap_augmenter = augmenty.load("token_swap.v1", level=0.20)
token_swap_augmenter = augmenty.yield_original(
token_swap_augmenter
) # yield both the augmented and original example
augmented_texts = augmenty.texts(texts, augmenter=token_swap_augmenter, nlp=nlp)
for text in augmented_texts:
print(text)
# -
# ## Applying augmentation to Examples or a Corpus
# Examples consists of two docs, one containing the predictions of the model, the other containing the gold labelled document. For this example we will load the DaNE dataset. DaNE includes the Danish dependency treebank additionally tagged for named entities. Here we will use synonym replacement to augment a corpus.
#
# To load the corpus we will use [DaCy](https://centre-for-humanities-computing.github.io/DaCy/) which we will install using:
# ``` bash
# pip install dacy
# ```
#
# And then we can apply the methods:
#
# ```python
# from dacy import datasets
#
# train, dev, test = datasets.dane(splits=["train", "dev", "test"])
#
# from spacy.lang.da import Danish
#
# nlp_da = Danish()
#
# synonym_augmenter = augmenty.load("wordnet_synonym.v1", level=0.2, lang="da")
# augmented_corpus = [
# e for example in test(nlp_da) for e in synonym_augmenter(nlp_da, example)
# ]
# ```
# ## Creating and Contributing Augmenters
#
# After using augmenty you might want to create and contribute an augmenter. Most augmenters can be created based on already existing augmenters. For instance the augmenter `per_replace.v1`, which replaces names in a text is a spacial case of the augmenter `ents_replace.v1` with better handling of first and last names. If you want to create an augmenter from scratch following spaCy's [guide](https://spacy.io/usage/training#data-augmentation-custom) on creating custom augmenters is a good start. You can always use augmenters from augmenty as inspiration as well. If you find yourself in troubles feel free to ask in the [augmenty forums](missing).
#
# When you are satisfied with your augmenter feel free submit a [pull request](https://github.com/KennethEnevoldsen/augmenty/pulls) to add the augmenter to augmenty.
|
docs/tutorials/introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Regression & Classification, Module 3
#
# ## Assignment
#
# We're going back to our other **New York City** real estate dataset. Instead of predicting apartment rents, you'll predict property sales prices.
#
# But not just for condos in Tribeca...
#
# Instead, predict property sales prices for **One Family Dwellings** (`BUILDING_CLASS_CATEGORY` == `'01 ONE FAMILY DWELLINGS'`).
#
# Use a subset of the data where the **sale price was more than \\$100 thousand and less than $2 million.**
#
# The [NYC Department of Finance](https://www1.nyc.gov/site/finance/taxes/property-rolling-sales-data.page) has a glossary of property sales terms and NYC Building Class Code Descriptions. The data comes from the [NYC OpenData](https://data.cityofnewyork.us/browse?q=NYC%20calendar%20sales) portal.
#
# - [ ] Do train/test split. Use data from January — March 2019 to train. Use data from April 2019 to test.
# - [ ] Do one-hot encoding of categorical features.
# - [ ] Do feature selection with `SelectKBest`.
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Fit a ridge regression model with multiple features.
# - [ ] Get mean absolute error for the test set.
# - [ ] As always, commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
# - [ ] Add your own stretch goal(s) !
# - [ ] Instead of `RidgeRegression`, try `LinearRegression`. Depending on how many features you select, your errors will probably blow up! 💥
# - [ ] Instead of `RidgeRegression`, try [`RidgeCV`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeCV.html).
# - [ ] Learn more about feature selection:
# - ["Permutation importance"](https://www.kaggle.com/dansbecker/permutation-importance)
# - [scikit-learn's User Guide for Feature Selection](https://scikit-learn.org/stable/modules/feature_selection.html)
# - [mlxtend](http://rasbt.github.io/mlxtend/) library
# - scikit-learn-contrib libraries: [boruta_py](https://github.com/scikit-learn-contrib/boruta_py) & [stability-selection](https://github.com/scikit-learn-contrib/stability-selection)
# - [_Feature Engineering and Selection_](http://www.feat.engineering/) by Kuhn & Johnson.
# - [ ] Try [statsmodels](https://www.statsmodels.org/stable/index.html) if you’re interested in more inferential statistical approach to linear regression and feature selection, looking at p values and 95% confidence intervals for the coefficients.
# - [ ] Read [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapters 1-3, for more math & theory, but in an accessible, readable way.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab={} colab_type="code" id="o9eSnDYhUGD7"
import os, sys
in_colab = 'google.colab' in sys.modules
# If you're in Colab...
if in_colab:
# Pull files from Github repo
os.chdir('/content')
# !git init .
# !git remote add origin https://github.com/LambdaSchool/DS-Unit-2-Regression-Classification.git
# !git pull origin master
# Install required python packages
# !pip install -r requirements.txt
# Change into directory for module
os.chdir('module3')
# + colab={} colab_type="code" id="ipBYS77PUwNR"
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + colab={} colab_type="code" id="QJBD4ruICm1m"
import pandas as pd
import pandas_profiling
import numpy as np
# Read New York City property sales data
df = pd.read_csv('../data/condos/NYC_Citywide_Rolling_Calendar_Sales.csv')
# Change column names: replace spaces with underscores
df.columns = [col.replace(' ', '_') for col in df]
# SALE_PRICE was read as strings.
# Remove symbols, convert to integer
df['SALE_PRICE'] = (
df['SALE_PRICE']
.str.replace('$','')
.str.replace('-','')
.str.replace(',','')
.astype(int)
)
# -
# BOROUGH is a numeric column, but arguably should be a categorical feature,
# so convert it from a number to a string
df['BOROUGH'] = df['BOROUGH'].astype(str)
# +
# Reduce cardinality for NEIGHBORHOOD feature
# Get a list of the top 10 neighborhoods
top10 = df['NEIGHBORHOOD'].value_counts()[:10].index
# At locations where the neighborhood is NOT in the top 10,
# replace the neighborhood with 'OTHER'
df.loc[~df['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'
# -
# First glance at the data
df.head()
# Check shape (rows / columns) of the data
df.shape
df = df.drop('EASE-MENT', axis=1)
# Only the range of sales prices between $100,000 and $2 million are required
df = df[(df['SALE_PRICE']>= 100000) & (df['SALE_PRICE']<= 2000000)]
# Only need single family homes:
df = df[(df['BUILDING_CLASS_CATEGORY'] == '01 ONE FAMILY DWELLINGS')]
# Check shape to see the affect of the pruning.
df.shape
# Check column dtypes
for i, value in enumerate(df):
print(f'For {value},the type is {df[value].dtype}')
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
from sklearn.model_selection import KFold
from sklearn.linear_model import Ridge
# +
# Find mean variance differiential
ignore = ['SALE_PRICE', 'BUILDING_CLASS_CATEGORY']
pdict = {}
for x in df:
if x in ignore:
print(f'Skipping {x}.')
pass
else:
print(f'Evaluating {x}.')
# Evaluate means
meanOne = df.groupby(str(x))['SALE_PRICE'].mean()
meanTwo = df['SALE_PRICE'].mean()
if len(meanOne) < 2:
print(f'Skipping {x}, because it failed the mean length test.')
pass
# Set math for mean variance differiential
else:
totalMean = df.groupby(str(x)).mean()['SALE_PRICE'].mean()
domain = (meanOne - totalMean)
meanVar = sum([x**2 for x in domain]) / (len(meanOne) - 1)
percentVar = ((meanVar**.5)*100) / totalMean
pdict[x] = percentVar
# Calculate PVD
p2dict = {}
pdictsums = sum([x for y, x in pdict.items()])
for key, value in pdict.items():
v2 = (value / pdictsums)
p2dict[key] = v2
# Tuple list
PVD = sorted(p2dict.items(), reverse = True, key = lambda x: x[1])
print(f'Done.')
# -
import matplotlib.pyplot as plt
# +
plt.style.use('fivethirtyeight')
plt.figure(figsize = (10,8), facecolor = '#ededed')
plt.axes(facecolor = '#ededed', frameon = False)
# Plot bar graph:
for i, k in enumerate(PVD):
plt.bar(k[0], k[1], color='C0')
plt.xticks(rotation = 90)
# -
for i in PVD[:5]:
print(f'{i[0]} is type {df[i[0]].dtype}.')
print(f'{df[i[0]].describe()}')
# +
# I don't need building class at present, only at time of sale. I'll drop that
# and take square footage instead. I also want land square footage!
checks = ['GROSS_SQUARE_FEET', 'LAND_SQUARE_FEET']
for x in checks:
print(df[x].describe())
# -
# GSF is good. Need to get the LSF as an integer instead of a string.
def lsf_to_int(string):
assert type(string) == str
x = string.replace(',', '')
return int(x)
df['LAND_SQUARE_FEET'] = df['LAND_SQUARE_FEET'].apply(lsf_to_int)
df['LAND_SQUARE_FEET'].describe()
features = ['TOTAL_UNITS', 'GROSS_SQUARE_FEET', 'LAND_SQUARE_FEET',
'COMMERCIAL_UNITS', 'RESIDENTIAL_UNITS','BLOCK']
# Instantiate the ridge regression model
model = Ridge(alpha=1)
# 10 fold cross validation
kf = KFold(n_splits=10, shuffle=True, random_state=42)
# +
# Define features and target
X = features
y = 'SALE_PRICE'
# Run model through and get metrics
r2 = [0]
mse = [0]
pred = [0]
for train_index, test_index in kf.split(df[X]):
X_train, X_test = df[X].iloc[train_index], df[X].iloc[test_index]
y_train, y_test = df[y].iloc[train_index], df[y].iloc[test_index]
ridgeReg = model.fit(X_train, y_train)
y_pred = model.predict(X_test)
if mse == [0]:
mse = mean_squared_error(y_test, y_pred)
r2 = r2_score(y_test, y_pred)
pred = y_pred
else:
if mean_squared_error(y_test, y_pred) < mse:
mse = mean_squared_error(y_test, y_pred)
r2 = r2_score(y_test, y_pred)
pred = y_pred
fModel = ridgeReg
print(f'MSE of ${mse:,.2f}')
print(f'R2 of {(r2*100):,.2f}%')
print(f'The model is {fModel}.')
# +
# Submitting the assignment like this to make sure it's in. Will update later.
|
module3/assignment_regression_classification_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ana2021
# language: python
# name: ana2021
# ---
import os
import sys
import numpy as np
import pandas as pd
import talib
import matplotlib.pyplot as plt
import plotly.graph_objects as go
from tqdm import tqdm
from sklearn.preprocessing import MinMaxScaler
df1 = pd.read_csv('/Users/zed/AI_Lab/DoubleEnsembleML/Data/BTC.csv')
df1 = df1.fillna(method='backfill')
df1
df1['r_0'] = df1.Close/df1.Close.shift(1)-1
def getDailyVol(close,span0 = 100):
df0 = close/close.shift(1)-1
df0 = df0.ewm(span = span0).std()
return df0
dailyVol = getDailyVol(df1.Close)
df1['tag'] = pd.Series(map(lambda x,y: 1 if x>0.2*y else(-1 if x<-0.2*y else 0),df1.r_0,dailyVol))
df2 =df1.loc[:,['Date','High','Low','Close','Volume (BTC)','tag']].rename(columns = {'Volume (BTC)':'Volume'})
df2
dataset = df2
df2.tag
# +
import matplotlib.pyplot as plt
n, bins, patches = plt.hist(x=df2.tag, bins='auto', color='blue',
alpha=0.7, rwidth=0.85)
plt.grid(axis='y', alpha=0.75)
plt.xlabel('Value')
plt.ylabel('Frequency')
plt.title('My Very Own Histogram')
plt.text(23, 45, r'$\mu=15, b=3$')
maxfreq = n.max()
plt.savefig("tag.png",dpi = 200)
# -
dataset['Open'] = dataset['Close'].shift(1)
dataset['H-L'] = dataset['High'] - dataset['Low']
# dataset['O-C'] = dataset['Close'] - dataset['Open']
dataset['3day MA'] = dataset['Close'].shift(1).rolling(window = 3).mean()
dataset['10day MA'] = dataset['Close'].shift(1).rolling(window = 10).mean()
dataset['30day MA'] = dataset['Close'].shift(1).rolling(window = 30).mean()
dataset['CCI'] = talib.AROONOSC(dataset.High, dataset.Low, timeperiod=14)
dataset['RSI'] = talib.RSI(dataset['Close'].values, timeperiod = 9)
dataset['ATR'] = talib.ATR(dataset['High'].values, dataset['Low'].values, dataset['Close'].values, 7)
dataset['OBV'] = talib.OBV(dataset.Close, dataset.Volume)
dataset['HT_DCPERIOD'] = talib.HT_DCPERIOD(dataset.Close)
y = dataset['tag']
dataset = dataset.drop(['tag'],axis=1)
y
dataset
dataset.Close.shift(1)
for i in range(1,6):
name = 'Last'+str(i)+'Price'
dataset[name] = dataset.Close.shift(i)
dataset['tag'] = y
dataset = dataset.dropna(how = 'any')
dataset.to_csv('mybtc.csv')
dataset
r = dataset.Close/dataset.Close.shift(1)-1
r.dropna(how = "any")
plt.style.use('ggplot')
plt.style.available
plt.style.use('seaborn-paper')
fig = plt.figure(figsize = (20,15))
plt.plot(r)
|
report/preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: snowmobile2
# language: python
# name: snowmobile2
# ---
# # Advanced Examples
# ---
# ```{admonition} TODO
# :class: error
# Stuff that can't be explained easily goes here
# ```
|
docs/usage/advanced.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/anoopsanka/retinal_oct/blob/main/notebooks/03d-OCT_Kaggle_SimCLRAug.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="0nVH0wWPJdLP" colab={"base_uri": "https://localhost:8080/"} outputId="86bf970b-1b27-48cb-a000-eb7259a77d15"
# !git clone https://hisunnytang:Qv!8!rae@github.com/anoopsanka/retinal_oct
# + colab={"base_uri": "https://localhost:8080/"} id="pzGPKh095zLq" outputId="72a93e96-f5e2-4fec-d056-249336895291"
# %cd retinal_oct
# + id="4lER0MN55uQI"
import matplotlib.pyplot as plt
import numpy as np
from importlib.util import find_spec
if find_spec("core") is None:
import sys
sys.path.append('..')
import tensorflow as tf
import tensorflow_datasets as tfds
from core.datasets import RetinaDataset
# + id="BxyzdaJ_J1rV" colab={"base_uri": "https://localhost:8080/", "height": 281, "referenced_widgets": ["ef6e0f00dea3422b8f6a9680218f85a1", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "969899b868324a32ba73722b1355294c", "80ba0906872d477a848048f3faa7e417", "<KEY>", "<KEY>", "caa41d7627c341e3872171b7e26aa436", "<KEY>", "<KEY>", "a2c8e66408f2420fa9ab676434087719", "765e740c52e24eb5b9d120df791c934c", "<KEY>", "978f518459ed4155bced40030c068808", "<KEY>", "499c5d7a95a049c392a02863d4a2b475", "762190a9ce2444d0b8fb86be08be7e19", "<KEY>", "86d70b4d36d4433097884d9950d58a0b", "<KEY>", "8a42c5b20e6d42da802b16f9cca8c481", "<KEY>", "1449a84349054d188305b43d04e8793d", "<KEY>", "454d6ec90af4491db2b2a5187f8ea011", "e6c4deea7683485e915dc6d1ed992a94", "<KEY>", "<KEY>", "56fddcb5d3504b22a81b98edaa3fe14d", "<KEY>", "<KEY>", "deb2a7ed1e7243a28241577fa055daaf", "1651bab6f6e0456c8205d8ea276b8da2", "2ad9b00ffe2247d98e26a70ee3acafe2", "<KEY>", "<KEY>", "1356cfaa576247cdbe53d0d35e54a39f", "<KEY>", "ccf06a1119a34d26830280fe60ef21c2", "<KEY>", "<KEY>", "9ca6cefaf60c4f63a1da06ca08b238cc", "0c64ad7ed4d947f1be6fc7e683629320", "9244ce379e9a4bf6ad46873ee564208e", "3b498207f45443e4a891d1ba752b5003", "1f0f317f21f94d25905f2b006f417c46"]} outputId="77dd4b18-c2d8-4b30-d137-03823ce39152"
ds_train, ds_train_info = tfds.load('RetinaDataset', split='train[:98%]', shuffle_files=True, as_supervised=True,with_info=True)
ds_val, ds_val_info = tfds.load('RetinaDataset', split='train[-2%:]', shuffle_files=True, as_supervised=True,with_info=True)
ds_test, ds_test_info = tfds.load('RetinaDataset', split='test', shuffle_files=True, as_supervised=True,with_info=True)
# + colab={"base_uri": "https://localhost:8080/"} id="XENjo6W2_DCL" outputId="8ae9020e-64b2-4f8c-ad90-55da9a85a827"
ds_train_info
# + id="GD2k8b5nMksN"
#@title SimCLR DataUtils
# coding=utf-8
# Copyright 2020 The SimCLR Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific simclr governing permissions and
# limitations under the License.
# ==============================================================================
"""Data preprocessing and augmentation."""
import functools
from absl import flags
import tensorflow.compat.v2 as tf
FLAGS = flags.FLAGS
CROP_PROPORTION = 0.875 # Standard for ImageNet.
def random_apply(func, p, x):
"""Randomly apply function func to x with probability p."""
return tf.cond(
tf.less(
tf.random.uniform([], minval=0, maxval=1, dtype=tf.float32),
tf.cast(p, tf.float32)), lambda: func(x), lambda: x)
def random_brightness(image, max_delta, impl='simclrv2'):
"""A multiplicative vs additive change of brightness."""
if impl == 'simclrv2':
factor = tf.random.uniform([], tf.maximum(1.0 - max_delta, 0),
1.0 + max_delta)
image = image * factor
elif impl == 'simclrv1':
image = tf.image.random_brightness(image, max_delta=max_delta)
else:
raise ValueError('Unknown impl {} for random brightness.'.format(impl))
return image
def to_grayscale(image, keep_channels=True):
image = tf.image.rgb_to_grayscale(image)
if keep_channels:
image = tf.tile(image, [1, 1, 3])
return image
def color_jitter(image, strength, random_order=True, impl='simclrv2'):
"""Distorts the color of the image.
Args:
image: The input image tensor.
strength: the floating number for the strength of the color augmentation.
random_order: A bool, specifying whether to randomize the jittering order.
impl: 'simclrv1' or 'simclrv2'. Whether to use simclrv1 or simclrv2's
version of random brightness.
Returns:
The distorted image tensor.
"""
brightness = 0.8 * strength
contrast = 0.8 * strength
saturation = 0.8 * strength
hue = 0.2 * strength
if random_order:
return color_jitter_rand(
image, brightness, contrast, saturation, hue, impl=impl)
else:
return color_jitter_nonrand(
image, brightness, contrast, saturation, hue, impl=impl)
def color_jitter_nonrand(image,
brightness=0,
contrast=0,
saturation=0,
hue=0,
impl='simclrv2'):
"""Distorts the color of the image (jittering order is fixed).
Args:
image: The input image tensor.
brightness: A float, specifying the brightness for color jitter.
contrast: A float, specifying the contrast for color jitter.
saturation: A float, specifying the saturation for color jitter.
hue: A float, specifying the hue for color jitter.
impl: 'simclrv1' or 'simclrv2'. Whether to use simclrv1 or simclrv2's
version of random brightness.
Returns:
The distorted image tensor.
"""
with tf.name_scope('distort_color'):
def apply_transform(i, x, brightness, contrast, saturation, hue):
"""Apply the i-th transformation."""
if brightness != 0 and i == 0:
x = random_brightness(x, max_delta=brightness, impl=impl)
elif contrast != 0 and i == 1:
x = tf.image.random_contrast(
x, lower=1-contrast, upper=1+contrast)
elif saturation != 0 and i == 2:
x = tf.image.random_saturation(
x, lower=1-saturation, upper=1+saturation)
elif hue != 0:
x = tf.image.random_hue(x, max_delta=hue)
return x
for i in range(4):
image = apply_transform(i, image, brightness, contrast, saturation, hue)
# image = tf.clip_by_value(image, 0., 1.)
return image
def color_jitter_rand(image,
brightness=0,
contrast=0,
saturation=0,
hue=0,
impl='simclrv2'):
"""Distorts the color of the image (jittering order is random).
Args:
image: The input image tensor.
brightness: A float, specifying the brightness for color jitter.
contrast: A float, specifying the contrast for color jitter.
saturation: A float, specifying the saturation for color jitter.
hue: A float, specifying the hue for color jitter.
impl: 'simclrv1' or 'simclrv2'. Whether to use simclrv1 or simclrv2's
version of random brightness.
Returns:
The distorted image tensor.
"""
with tf.name_scope('distort_color'):
def apply_transform(i, x):
"""Apply the i-th transformation."""
def brightness_foo():
if brightness == 0:
return x
else:
return random_brightness(x, max_delta=brightness, impl=impl)
def contrast_foo():
if contrast == 0:
return x
else:
return tf.image.random_contrast(x, lower=1-contrast, upper=1+contrast)
def saturation_foo():
if saturation == 0:
return x
else:
return tf.image.random_saturation(
x, lower=1-saturation, upper=1+saturation)
def hue_foo():
if hue == 0:
return x
else:
return tf.image.random_hue(x, max_delta=hue)
x = tf.cond(tf.less(i, 2),
lambda: tf.cond(tf.less(i, 1), brightness_foo, contrast_foo),
lambda: tf.cond(tf.less(i, 3), saturation_foo, hue_foo))
return x
perm = tf.random.shuffle(tf.range(4))
for i in range(4):
image = apply_transform(perm[i], image)
#image = tf.clip_by_value(image, 0., 1.)
return image
def _compute_crop_shape(
image_height, image_width, aspect_ratio, crop_proportion):
"""Compute aspect ratio-preserving shape for central crop.
The resulting shape retains `crop_proportion` along one side and a proportion
less than or equal to `crop_proportion` along the other side.
Args:
image_height: Height of image to be cropped.
image_width: Width of image to be cropped.
aspect_ratio: Desired aspect ratio (width / height) of output.
crop_proportion: Proportion of image to retain along the less-cropped side.
Returns:
crop_height: Height of image after cropping.
crop_width: Width of image after cropping.
"""
image_width_float = tf.cast(image_width, tf.float32)
image_height_float = tf.cast(image_height, tf.float32)
def _requested_aspect_ratio_wider_than_image():
crop_height = tf.cast(
tf.math.rint(crop_proportion / aspect_ratio * image_width_float),
tf.int32)
crop_width = tf.cast(
tf.math.rint(crop_proportion * image_width_float), tf.int32)
return crop_height, crop_width
def _image_wider_than_requested_aspect_ratio():
crop_height = tf.cast(
tf.math.rint(crop_proportion * image_height_float), tf.int32)
crop_width = tf.cast(
tf.math.rint(crop_proportion * aspect_ratio * image_height_float),
tf.int32)
return crop_height, crop_width
return tf.cond(
aspect_ratio > image_width_float / image_height_float,
_requested_aspect_ratio_wider_than_image,
_image_wider_than_requested_aspect_ratio)
def center_crop(image, height, width, crop_proportion):
"""Crops to center of image and rescales to desired size.
Args:
image: Image Tensor to crop.
height: Height of image to be cropped.
width: Width of image to be cropped.
crop_proportion: Proportion of image to retain along the less-cropped side.
Returns:
A `height` x `width` x channels Tensor holding a central crop of `image`.
"""
shape = tf.shape(image)
image_height = shape[0]
image_width = shape[1]
crop_height, crop_width = _compute_crop_shape(
image_height, image_width, height / width, crop_proportion)
offset_height = ((image_height - crop_height) + 1) // 2
offset_width = ((image_width - crop_width) + 1) // 2
image = tf.image.crop_to_bounding_box(
image, offset_height, offset_width, crop_height, crop_width)
image = tf.image.resize([image], [height, width],
method=tf.image.ResizeMethod.BICUBIC)[0]
return image
def distorted_bounding_box_crop(image,
bbox,
min_object_covered=0.1,
aspect_ratio_range=(0.75, 1.33),
area_range=(0.05, 1.0),
max_attempts=100,
scope=None):
"""Generates cropped_image using one of the bboxes randomly distorted.
See `tf.image.sample_distorted_bounding_box` for more documentation.
Args:
image: `Tensor` of image data.
bbox: `Tensor` of bounding boxes arranged `[1, num_boxes, coords]`
where each coordinate is [0, 1) and the coordinates are arranged
as `[ymin, xmin, ymax, xmax]`. If num_boxes is 0 then use the whole
image.
min_object_covered: An optional `float`. Defaults to `0.1`. The cropped
area of the image must contain at least this fraction of any bounding
box supplied.
aspect_ratio_range: An optional list of `float`s. The cropped area of the
image must have an aspect ratio = width / height within this range.
area_range: An optional list of `float`s. The cropped area of the image
must contain a fraction of the supplied image within in this range.
max_attempts: An optional `int`. Number of attempts at generating a cropped
region of the image of the specified constraints. After `max_attempts`
failures, return the entire image.
scope: Optional `str` for name scope.
Returns:
(cropped image `Tensor`, distorted bbox `Tensor`).
"""
with tf.name_scope(scope or 'distorted_bounding_box_crop'):
shape = tf.shape(image)
sample_distorted_bounding_box = tf.image.sample_distorted_bounding_box(
shape,
bounding_boxes=bbox,
min_object_covered=min_object_covered,
aspect_ratio_range=aspect_ratio_range,
area_range=area_range,
max_attempts=max_attempts,
use_image_if_no_bounding_boxes=True)
bbox_begin, bbox_size, _ = sample_distorted_bounding_box
# Crop the image to the specified bounding box.
offset_y, offset_x, _ = tf.unstack(bbox_begin)
target_height, target_width, _ = tf.unstack(bbox_size)
image = tf.image.crop_to_bounding_box(
image, offset_y, offset_x, target_height, target_width)
return image
def crop_and_resize(image, height, width):
"""Make a random crop and resize it to height `height` and width `width`.
Args:
image: Tensor representing the image.
height: Desired image height.
width: Desired image width.
Returns:
A `height` x `width` x channels Tensor holding a random crop of `image`.
"""
bbox = tf.constant([0.0, 0.0, 1.0, 1.0], dtype=tf.float32, shape=[1, 1, 4])
aspect_ratio = width / height
image = distorted_bounding_box_crop(
image,
bbox,
min_object_covered=0.1,
aspect_ratio_range=(3. / 4 * aspect_ratio, 4. / 3. * aspect_ratio),
area_range=(0.08, 1.0),
max_attempts=100,
scope=None)
return tf.image.resize([image], [height, width],
method=tf.image.ResizeMethod.BICUBIC)[0]
def gaussian_blur(image, kernel_size, sigma, padding='SAME'):
"""Blurs the given image with separable convolution.
Args:
image: Tensor of shape [height, width, channels] and dtype float to blur.
kernel_size: Integer Tensor for the size of the blur kernel. This is should
be an odd number. If it is an even number, the actual kernel size will be
size + 1.
sigma: Sigma value for gaussian operator.
padding: Padding to use for the convolution. Typically 'SAME' or 'VALID'.
Returns:
A Tensor representing the blurred image.
"""
radius = tf.cast(kernel_size / 2, dtype=tf.int32)
kernel_size = radius * 2 + 1
x = tf.cast(tf.range(-radius, radius + 1), dtype=tf.float32)
blur_filter = tf.exp(-tf.pow(x, 2.0) /
(2.0 * tf.pow(tf.cast(sigma, dtype=tf.float32), 2.0)))
blur_filter /= tf.reduce_sum(blur_filter)
# One vertical and one horizontal filter.
blur_v = tf.reshape(blur_filter, [kernel_size, 1, 1, 1])
blur_h = tf.reshape(blur_filter, [1, kernel_size, 1, 1])
num_channels = tf.shape(image)[-1]
blur_h = tf.tile(blur_h, [1, 1, num_channels, 1])
blur_v = tf.tile(blur_v, [1, 1, num_channels, 1])
expand_batch_dim = image.shape.ndims == 3
if expand_batch_dim:
# Tensorflow requires batched input to convolutions, which we can fake with
# an extra dimension.
image = tf.expand_dims(image, axis=0)
blurred = tf.nn.depthwise_conv2d(
image, blur_h, strides=[1, 1, 1, 1], padding=padding)
blurred = tf.nn.depthwise_conv2d(
blurred, blur_v, strides=[1, 1, 1, 1], padding=padding)
if expand_batch_dim:
blurred = tf.squeeze(blurred, axis=0)
return blurred
def random_crop_with_resize(image, height, width, p=1.0):
"""Randomly crop and resize an image.
Args:
image: `Tensor` representing an image of arbitrary size.
height: Height of output image.
width: Width of output image.
p: Probability of applying this transformation.
Returns:
A preprocessed image `Tensor`.
"""
def _transform(image): # pylint: disable=missing-docstring
image = crop_and_resize(image, height, width)
return image
return random_apply(_transform, p=p, x=image)
def random_color_jitter(image, p=1.0, impl='simclrv2', color_jitter_strength=0.8):
def _transform(image):
color_jitter_t = functools.partial(
color_jitter, strength=color_jitter_strength, impl=impl)
image = random_apply(color_jitter_t, p=0.8, x=image)
return random_apply(to_grayscale, p=0.2, x=image)
return random_apply(_transform, p=p, x=image)
def random_blur(image, height, width, p=1.0):
"""Randomly blur an image.
Args:
image: `Tensor` representing an image of arbitrary size.
height: Height of output image.
width: Width of output image.
p: probability of applying this transformation.
Returns:
A preprocessed image `Tensor`.
"""
del width
def _transform(image):
sigma = tf.random.uniform([], 0.1, 2.0, dtype=tf.float32)
return gaussian_blur(
image, kernel_size=height//10, sigma=sigma, padding='SAME')
return random_apply(_transform, p=p, x=image)
def batch_random_blur(images_list, height, width, blur_probability=0.5):
"""Apply efficient batch data transformations.
Args:
images_list: a list of image tensors.
height: the height of image.
width: the width of image.
blur_probability: the probaility to apply the blur operator.
Returns:
Preprocessed feature list.
"""
def generate_selector(p, bsz):
shape = [bsz, 1, 1, 1]
selector = tf.cast(
tf.less(tf.random.uniform(shape, 0, 1, dtype=tf.float32), p),
tf.float32)
return selector
new_images_list = []
for images in images_list:
images_new = random_blur(images, height, width, p=1.)
selector = generate_selector(blur_probability, tf.shape(images)[0])
images = images_new * selector + images * (1 - selector)
images = tf.clip_by_value(images, 0., 1.)
new_images_list.append(images)
return new_images_list
def preprocess_for_train(image,
height,
width,
color_distort=True,
crop=True,
flip=True,
color_jitter_strength=0.9,
impl='simclrv2'):
"""Preprocesses the given image for training.
Args:
image: `Tensor` representing an image of arbitrary size.
height: Height of output image.
width: Width of output image.
color_distort: Whether to apply the color distortion.
crop: Whether to crop the image.
flip: Whether or not to flip left and right of an image.
impl: 'simclrv1' or 'simclrv2'. Whether to use simclrv1 or simclrv2's
version of random brightness.
Returns:
A preprocessed image `Tensor`.
"""
if crop:
image = random_crop_with_resize(image, height, width)
if flip:
image = tf.image.random_flip_left_right(image)
if color_distort:
image = random_color_jitter(image, impl=impl,
color_jitter_strength=color_jitter_strength)
image = tf.reshape(image, [height, width, 3])
# image = tf.clip_by_value(image, 0., 1.)
return image
def preprocess_for_eval(image, height, width, crop=True):
"""Preprocesses the given image for evaluation.
Args:
image: `Tensor` representing an image of arbitrary size.
height: Height of output image.
width: Width of output image.
crop: Whether or not to (center) crop the test images.
Returns:
A preprocessed image `Tensor`.
"""
if crop:
image = center_crop(image, height, width, crop_proportion=CROP_PROPORTION)
image = tf.reshape(image, [height, width, 3])
# image = tf.clip_by_value(image, 0., 1.)
return image
def preprocess_image(image, height, width, is_training=False,
color_distort=True, test_crop=True):
"""Preprocesses the given image.
Args:
image: `Tensor` representing an image of arbitrary size.
height: Height of output image.
width: Width of output image.
is_training: `bool` for whether the preprocessing is for training.
color_distort: whether to apply the color distortion.
test_crop: whether or not to extract a central crop of the images
(as for standard ImageNet evaluation) during the evaluation.
Returns:
A preprocessed image `Tensor` of range [0, 1].
"""
image = tf.image.convert_image_dtype(image, dtype=tf.float32)
if is_training:
return preprocess_for_train(image, height, width, color_distort)
else:
return preprocess_for_eval(image, height, width, test_crop)
# + id="KM1HTJ3uIKhm"
import matplotlib.pyplot as plt
import numpy as np
def view_image(ds):
image, label = next(iter(ds)) # extract 1 batch from the dataset
image = image #.numpy()
label = label #.numpy()
fig = plt.figure(figsize=(22, 22))
for i in range(16):
ax = fig.add_subplot(4, 4, i+1, xticks=[], yticks=[])
ax.imshow(image[i].astype(np.int32))
ax.set_title(f"Label: {label[i]}")
return image
# + id="TOfMpk2RIK0G"
from functools import partial
NCLASS = 4
IMG_SIZE = 224
def resize_image(img, lb):
return tf.image.resize(img, (224,224)), tf.one_hot(lb, NCLASS)
def augment_image(img, lb):
img, lb = resize_image(img, lb)
return preprocess_for_train(img, height=IMG_SIZE, width=IMG_SIZE), lb
ds_train_augment = ds_train.map(augment_image)
ds_val = ds_val.map(resize_image)
# + id="SoC1fHdQ60xS"
x = next(iter(ds_val))
# + colab={"base_uri": "https://localhost:8080/"} id="AFi8jL7zGmzn" outputId="de09b44d-dd0f-449d-bff0-eb9a74acf45b"
x[1]
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="tMJS_gM783cJ" outputId="f2b74904-430c-4124-ab3b-499d622af81a"
plt.imshow(x[0].numpy().astype(int))
# + id="eG3ol-iAY9eK"
def create_basemodel(base_model='resnet',
input_shape=(224,224,3),
output_units=4):
if base_model == 'resnet':
preprocess = tf.keras.applications.resnet_v2.preprocess_input
base_model = tf.keras.applications.ResNet50V2(include_top=False, weights='imagenet')
elif base_model == 'xception':
preprocess = tf.keras.applications.xception.preprocess_input
base_model = tf.keras.applications.Xception(include_top=False, weights='imagenet')
elif base_model == 'inception':
preprocess = tf.keras.layers.Lambda(lambda x: x)
base_model = tf.kaers.applications.inception_v3.InceptionV3(include_top=False, weights='imagenet')
else:
raise f"{base_model} not supported, choose from ['resnet', 'xception', 'inception']"
base_model.trainable = False
inputs = tf.keras.layers.Input(input_shape)
pool = tf.keras.layers.GlobalAveragePooling2D()
flatten = tf.keras.layers.Flatten()
softmax = tf.keras.layers.Dense(output_units, activation='softmax')
x = inputs
x = preprocess(x)
x = base_model(x)
x = pool(x)
x = flatten(x)
out = softmax(x)
return tf.keras.Model(inputs=inputs, outputs=out)
# + id="T6F419ru-MwB"
y_labels = []
labels = ds_train_augment.map(lambda x, y: y)
for l in labels.batch(64).as_numpy_iterator():
y_labels.append(l)
# + id="34mvOcPb-eeY"
y_labels = np.vstack(y_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="DotZX8A9vg--" outputId="3b6b465a-d8dd-4e2b-dc39-c5601f639435"
y_labels.sum(axis=0)
# + [markdown] id="FKgyYzWKkcML"
# # Compute Class Weight
# + id="Pc0dCVvZyH-Q"
from sklearn.utils.class_weight import compute_class_weight, compute_sample_weight
class_weights = compute_class_weight('balanced', [0, 1, 2, 3], y_labels.argmax(axis=1))
class_weights = {i: w for i, w in enumerate(class_weights)}
# + colab={"base_uri": "https://localhost:8080/"} id="AEp6JI6jX2nf" outputId="db9b8a4d-060a-42b2-c789-8de883291adf"
class_weights
# + [markdown] id="8weFi1g_keVg"
# # Feature Extraction
# + id="JDTEL8w0cZbf" colab={"base_uri": "https://localhost:8080/"} outputId="a3038f62-440d-4e14-e0d0-5d48acf8bddc"
resnet_base = create_basemodel('resnet')
metrics = ['accuracy']
callbacks = [tf.keras.callbacks.EarlyStopping(patience=3, monitor='val_loss', ),
tf.keras.callbacks.ModelCheckpoint(filepath='resnet_model.{epoch:02d}-{val_loss:.2f}.h5'),]
optimizer = tf.keras.optimizers.Adam(lr=1e-3)
resnet_base.compile(optimizer=optimizer,
loss='categorical_crossentropy',
metrics=metrics)
# + colab={"base_uri": "https://localhost:8080/"} id="3leGKwP0A7Bh" outputId="493d8e3f-d1d9-4306-85cf-ccbfdc727a7e"
resnet_base(tf.zeros((1,224,224,3)))
# + colab={"base_uri": "https://localhost:8080/"} id="nWrvDJrKvPju" outputId="07f2f191-6a07-4d13-ed72-be39b81b31e6"
resnet_base.fit(ds_train_augment.batch(32),
validation_data = ds_val.batch(32),
callbacks = callbacks,
class_weight = class_weights,
epochs=20)
# + id="NxhzWhwzXqY7" colab={"base_uri": "https://localhost:8080/"} outputId="9464d102-5f0c-48d4-bb2e-7eeb0719b8f4"
# %ls
# + id="ixLuJXB_H0AS"
refine_resnet_base = create_basemodel('resnet')
refine_resnet_base.load_weights('resnet_model.04-0.45.h5')
# + colab={"base_uri": "https://localhost:8080/"} id="mKKhfubbJPrx" outputId="a03c706b-ad80-4c90-8a0a-b8acc4899c21"
metrics = ['accuracy']
optimizer = tf.keras.optimizers.Adam(lr=1e-3)
refine_resnet_base.compile(optimizer=optimizer,
loss='categorical_crossentropy',
metrics=metrics)
refine_resnet_base.evaluate(ds_val.batch(32))
# + [markdown] id="4LQ5MZALJcsO"
# # Fine Tuning!
# - only fine-tune the very-last block of resnet
# - bear in mind that the weights in the `BatchNorm` layers should be freezed, i.e. in the evaluation mode
# + colab={"base_uri": "https://localhost:8080/"} id="NQQY4Oz9JYMd" outputId="3be61835-2735-4396-bf60-1e640cc49819"
refine_resnet_base.summary()
# + id="kpsbdaq1JgWL"
# Extract the base model
base_model = refine_resnet_base.layers[3]
# unfreeze it
base_model.trainable = True
# select only the last resnet block for retraining
# keeping the batchnorm layer unchanged
for l in base_model.layers:
name = l.name
if name.startswith('conv5_block3') and not isinstance(l, tf.keras.layers.BatchNormalization):
l.trainable = True
else:
l.trainable = False
optimizer = tf.keras.optimizers.Adam(lr=1e-4)
refine_resnet_base.compile(optimizer=optimizer,
loss='categorical_crossentropy',
metrics=metrics)
# + colab={"base_uri": "https://localhost:8080/"} id="CghBxAAaKBqp" outputId="7b1ef978-46dd-4bcd-8879-0d2edfd1d3bb"
callbacks = [tf.keras.callbacks.EarlyStopping(patience=3, monitor='val_loss', ),
tf.keras.callbacks.ModelCheckpoint(filepath='finetune_resnet_model.{epoch:02d}-{val_loss:.2f}.h5'),]
refine_resnet_base.fit(ds_train_augment.batch(32),
validation_data = ds_val.batch(32),
callbacks = callbacks,
class_weight = class_weights,
epochs=20)
# + [markdown] id="H1c3hJUpdp8E"
# # Restore the Best Model
# + id="LZ7PgmPOdvgr"
refine_resnet_base.load_weights('finetune_resnet_model.06-0.25.h5')
# + colab={"base_uri": "https://localhost:8080/"} id="2B2e5W8tdEnU" outputId="c051f828-8ac2-49dc-ef6f-c65d58e3dde5"
# def to_onehot(img,lb):
# return img, tf.one_hot(lb, 4)
refine_resnet_base.evaluate(ds_test.map(resize_image).batch(32))
# + colab={"base_uri": "https://localhost:8080/"} id="rf2eLTRjdlmu" outputId="ef340bef-92f7-4248-dbc5-a2f6da6bc764"
refine_resnet_base.evaluate(ds_val.batch(32))
# + id="JM3pEpFtMDBd"
# IMG_SIZE=224
# import tensorflow.keras.layers as layers
# data_augmentation = tf.keras.Sequential([
# layers.experimental.preprocessing.Resizing(IMG_SIZE, IMG_SIZE),
# layers.experimental.preprocessing.RandomFlip("horizontal_and_vertical"),
# layers.experimental.preprocessing.RandomRotation(20.),
# layers.experimental.preprocessing.RandomTranslation(height_factor=(-0.2,0.2),
# width_factor= (-0.2,0.2) ),
# layers.experimental.preprocessing.RandomZoom((-0.2,0.2)),
# ])
# ds_augment = ds_train.batch(32).map(lambda x, y: (data_augmentation(x, training=True), y))
|
notebooks/03d-OCT_Kaggle_SimCLRAug.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8 - AzureML
# language: python
# name: python38-azureml
# ---
# # Read and write from Numpy to Azure Data Lake and Azure Blob storage
# ## Install requirements
# %pip install -r requirements.txt
# ## Initialize public Blob filesystem
container_name = "datasets"
storage_options = {"account_name": "azuremlexamples"}
# +
from adlfs import AzureBlobFileSystem as abfs
fs = abfs(**storage_options)
files = fs.ls(f"{container_name}/mnist")
files
# -
# ## Define fuctions to read gzipped MNIST data
# +
import numpy as np
def read_images(f, num_images, image_size=28):
f.read(16) # magic
buf = f.read(image_size * image_size * num_images)
images = np.frombuffer(buf, dtype=np.uint8).astype(np.float32)
images = images.reshape(num_images, image_size, image_size, 1)
return images
def read_labels(f, num_labels):
f.read(8) # magic
buf = f.read(num_labels)
labels = np.frombuffer(buf, dtype=np.uint8)
return labels
# -
# ## Read in gzipped MNIST data
# +
import gzip
train_len = 60000
test_len = 10000
for f in files:
if "train-images" in f:
X_train = read_images(gzip.open(fs.open(f)), train_len)
elif "train-labels" in f:
y_train = read_labels(gzip.open(fs.open(f)), train_len)
elif "images" in f:
X_test = read_images(gzip.open(fs.open(f)), test_len)
elif "labels" in f:
y_test = read_labels(gzip.open(fs.open(f)), test_len)
# -
# ## Verify expected results
# +
from random import randint
i = randint(0, train_len)
x = X_train[i]
y = y_train[i]
# +
import matplotlib.pyplot as plt
plt.imshow(x.squeeze())
plt.title(f"Label: {y}")
# -
# ## Initialize private Blob filesystem
# +
from azureml.core import Workspace
ws = Workspace.from_config()
ds = ws.datastores["workspaceblobstore"]
container_name = ds.container_name
storage_options = {"account_name": ds.account_name, "account_key": ds.account_key}
# -
fs = abfs(**storage_options)
fs
fs.ls(f"{container_name}")
# ## Write numpy arrays using `np.save`
# +
with fs.open(f"{container_name}/example-data/mnist/X_train.npy", "wb") as f:
np.save(f, X_train)
with fs.open(f"{container_name}/example-data/mnist/y_train.npy", "wb") as f:
np.save(f, y_train)
with fs.open(f"{container_name}/example-data/mnist/X_test.npy", "wb") as f:
np.save(f, X_test)
with fs.open(f"{container_name}/example-data/mnist/y_test.npy", "wb") as f:
np.save(f, y_test)
# -
# ## Load numpy arrays using `np.load`
# +
with fs.open(f"{container_name}/example-data/mnist/X_train.npy", "rb") as f:
X_train = np.load(f)
with fs.open(f"{container_name}/example-data/mnist/y_train.npy", "rb") as f:
y_train = np.load(f)
with fs.open(f"{container_name}/example-data/mnist/X_test.npy", "rb") as f:
X_test = np.load(f)
with fs.open(f"{container_name}/example-data/mnist/y_test.npy", "rb") as f:
y_test = np.load(f)
# -
# ## Verify expected results
# +
from random import randint
i = randint(0, train_len)
x = X_train[i]
y = y_train[i]
# +
import matplotlib.pyplot as plt
plt.imshow(x.squeeze())
plt.title(f"Label: {y}")
|
notebooks/cloud-data/blob-adls-numpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow2_p36
# language: python
# name: conda_tensorflow2_p36
# ---
# ## Image classification with Tensorflow
#
import os
import numpy as np
import io
import math
import time
import seaborn as sns
from sklearn.metrics import accuracy_score,confusion_matrix
import matplotlib.pyplot as plt
train_dir = os.path.join(os.getcwd(), 'splitdata/train')
validation_dir = os.path.join(os.getcwd(), 'splitdata/val')
# +
# # !wget -q https://raw.githubusercontent.com/aws-samples/amazon-sagemaker-script-mode/master/local_mode_setup.sh
# # !wget -q https://raw.githubusercontent.com/aws-samples/amazon-sagemaker-script-mode/master/daemon.json
# # !/bin/bash ./local_mode_setup.sh
# -
# !pygmentize classification.py
# +
import sagemaker
from sagemaker.tensorflow import TensorFlow
model_dir = '/opt/ml/model'
train_instance_type = 'local'
hyperparameters = {'epochs': 1}
local_estimator = TensorFlow(
entry_point='classification.py',
model_dir=model_dir,
train_instance_type=train_instance_type,
train_instance_count=1,
hyperparameters=hyperparameters,
role=sagemaker.get_execution_role(),
base_job_name='tf-keras-clasif',
framework_version='2.0.0',
py_version='py3',
script_mode=True)
# +
inputs = {'train': f'file://{train_dir}','validation': f'file://{validation_dir}'}
local_estimator.fit(inputs)
# +
s3_prefix = 'tf-caltech-sample4'
traindata_s3_prefix = '{}/data/train'.format(s3_prefix)
validation_s3_prefix = '{}/data/validation'.format(s3_prefix)
train_s3 = sagemaker.Session().upload_data(path='./splitdata/train/', key_prefix=traindata_s3_prefix)
validation_s3 = sagemaker.Session().upload_data(path='./splitdata/val/', key_prefix=validation_s3_prefix)
inputs = {'train':train_s3,'validation':validation_s3}
print(inputs)
# +
train_instance_type = 'ml.p3.2xlarge'
hyperparameters = {'epochs': 30}
estimator = TensorFlow(
entry_point='classification.py',
model_dir=model_dir,
train_instance_type=train_instance_type,
train_instance_count=1,
hyperparameters=hyperparameters,
role=sagemaker.get_execution_role(),
base_job_name='tf-keras-clasif',
framework_version='2.0.0',
py_version='py3',
script_mode=True)
# -
estimator.fit(inputs)
# ## Create the Estimator
# I can use the estimator at any time
estimator
predictor = estimator.deploy(initial_instance_count=1, instance_type='ml.m5.xlarge')
# ## Model Evaluation
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from sklearn.metrics import accuracy_score
# +
train_data_gen_args = dict(rescale=1./255)
data_gen_args = dict(target_size=(224, 224),
batch_size=16,
shuffle=True,
#color_mode='grayscale',
class_mode='categorical')
# -
train_datagen = ImageDataGenerator(**train_data_gen_args)
test_generator = train_datagen.flow_from_directory('splitdata/test/', **data_gen_args)
# +
number_of_examples = 22
number_of_generator_calls = math.ceil(number_of_examples / (1.0 * data_gen_args['batch_size']))
test_labels = []
predictions = []
for i in range(0,int(number_of_generator_calls)):
instances = test_generator[i][0]
print(instances.shape)
for instance in instances:
array = instance.reshape((1,) + instance.shape)
payload = {
'instances': array.tolist()
}
resp = predictor.predict(payload)['predictions']
predictions.append(np.array(resp))
test_labels.extend(np.array(test_generator[i][1]))
# -
np.array(predictions).shape
predictions = np.array(predictions).reshape(22,4)
np.argmax(predictions,axis=1)
predictions = np.argmax(predictions,axis=1)
labels = np.argmax(np.array(test_labels),axis=1)
accuracy_score(labels,predictions)
test_generator.class_indices
classes = list(test_generator.class_indices.keys())
classes
# +
df_cm = confusion_matrix(labels,predictions,labels=np.unique(labels))
heatmap = sns.heatmap(df_cm, annot=True, fmt="d")
heatmap.yaxis.set_ticklabels(classes, rotation=0, ha='right')
heatmap.xaxis.set_ticklabels(classes, rotation=45, ha='right')
plt.ylabel('Valor Verdadero')
plt.xlabel('Valor Predicho');
# -
# ## Boto3
#
# (Real time predictor for Lambda)
import boto3
import json
client = boto3.client('sagemaker-runtime')
type(payload)
endpoint_name = "tf-keras-clasif-2020-10-22-23-05-15-949" # Your endpoint name.
content_type = "application/json" # The MIME type of the input data in the request body.
json_payload = json.dumps(payload)
response = client.invoke_endpoint(
EndpointName=endpoint_name,
ContentType=content_type,
Body=json_payload
)
eval(response['Body'].read())
|
week2/day3/sagemaker-tensorflow2/Classification-Train-Serve/ClassificationTFContainer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Autocompletion
#
# ## N-Gram Language Model
# +
import math
import random
import nltk
import numpy
import pandas
nltk.data.path.append(".")
# -
with open("en_US.twitter.txt", "r") as f:
data = f.read()
print("Data type:", type(data))
print("Number of letters:", len(data))
print("First 300 letters of the data")
print("-------")
data[0:300]
# ### Preprocessing
# +
import os
def split_data(data):
return (sentence.strip() for sentence in data.split(os.linesep) if len(sentence.strip()) > 0)
def tokenize_sentences(sentences):
return (nltk.word_tokenize(sentence.lower()) for sentence in sentences)
def tokenize_data(data):
return list(tokenize_sentences(split_data(data)))
# -
tokenized_data = tokenize_data(data)
tokenized_data[:2]
# ### Splitting Dataset
# +
train_set_share = 0.8
train_set_size = int(len(tokenized_data) * 0.8)
train_set = tokenized_data[:train_set_size]
test_set = tokenized_data[train_set_size:]
print(
os.linesep.join(
[f"train set size={len(train_set)}", f"test set size={len(test_set)}", f"total={len(tokenized_data)}"]
)
)
# -
# ### Using Unknown Words
# +
from collections import defaultdict
def get_vocabularies(tokenized_sentences, threshold=2):
word_map = defaultdict(lambda: 0)
for sentence in tokenized_sentences:
for word in sentence:
word_map[word] += 1
return {word: count for word, count in word_map.items() if count > threshold}
def replace_rare_words(tokenized_sentences, vocabularies, unknown_word_token="<UNK>"):
replaced_sentences = []
for sentence in tokenized_sentences:
sentence_copy = []
for word in sentence:
sentence_copy.append(word if word in vocabularies else unknown_word_token)
replaced_sentences.append(sentence_copy)
return replaced_sentences
# -
vocabularies = get_vocabularies(train_set)
train_set_2 = replace_rare_words(train_set, vocabularies)
train_set_2[10:12]
# ### Counting N-Grams
def count_n_grams(tokenized_sentences, n, starting_token="<S>", ending_token="<E>"):
n_gram_counts = defaultdict(lambda: 0)
for sentence in tokenized_sentences:
sentence = tuple([starting_token] * n + sentence + [ending_token])
for i in range(len(sentence) - n + 1):
n_gram = sentence[i : i + n]
n_gram_counts[n_gram] += 1
return n_gram_counts
bigram_counts = count_n_grams(train_set_2, 2)
trigram_counts = count_n_grams(train_set_2, 3)
# ### Estimate Probabilities
# +
def estimate_probability(word, previous_n_gram, n_gram_counts, n_plus_1_gram_counts, vocabularies_size, k=1.0):
previous_n_gram = tuple(previous_n_gram)
previous_n_gram_count = n_gram_counts.get(previous_n_gram, 0)
n_plus_1_gram = (*previous_n_gram, word)
n_plus_1_gram_count = n_plus_1_gram_counts.get(n_plus_1_gram, 0)
return (n_plus_1_gram_count + k) / (previous_n_gram_count + k * vocabularies_size)
def estimate_probabilities(
previous_n_gram,
n_gram_counts,
n_plus_1_gram_counts,
vocabularies,
ending_token="<E>",
unknown_word_token="<UNK>",
k=1.0,
):
words = list(vocabularies.keys()) + [ending_token, unknown_word_token]
return {
word: estimate_probability(word, previous_n_gram, n_gram_counts, n_plus_1_gram_counts, len(vocabularies), k)
for word in words
}
# +
probs = estimate_probabilities(("how", "are"), bigram_counts, trigram_counts, vocabularies)
from collections import Counter
most_likely = Counter(probs).most_common(1)[0][0]
print("how are (?)")
print(f"most likely: how are {most_likely}")
# -
# ## Perplexity Evaluation
def get_perplexity(
test_sentence,
n_gram_counts,
n_plus_1_gram_counts,
vocabularies_size,
starting_token="<S>",
ending_token="<E>",
k=1.0,
):
n = len(next(iter(n_gram_counts.keys())))
tokenized_test_sentence = (starting_token,) * n + tuple(nltk.word_tokenize(test_sentence.lower())) + (ending_token,)
production = 1
for i in range(n, len(tokenized_test_sentence)):
n_gram = tokenized_test_sentence[i - n : i]
n_plus_1_gram = (*n_gram, tokenized_test_sentence[i])
production /= (n_plus_1_gram_counts.get(n_plus_1_gram, 0) + k) / (
n_gram_counts.get(n_gram, 0) + k * vocabularies_size
)
return production ** (1 / len(tokenized_test_sentence))
# +
# the lower the better
print(get_perplexity("i always wonder", bigram_counts, trigram_counts, len(vocabularies)))
print(get_perplexity("wonder i always", bigram_counts, trigram_counts, len(vocabularies)))
print(get_perplexity("i wonder always", bigram_counts, trigram_counts, len(vocabularies)))
print(get_perplexity("i go to school", bigram_counts, trigram_counts, len(vocabularies)))
print(get_perplexity("go i to school", bigram_counts, trigram_counts, len(vocabularies)))
print(get_perplexity("school i go to", bigram_counts, trigram_counts, len(vocabularies)))
|
week7/conclusion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Major Imports
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
# # Some Pre-Processing Steps
#
# ### Batch SIze = 200
# ### Epochs = 10
# +
batch_size = 200
num_classes = 10
epochs = 10
# input image dimensions
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# -
# # Network Initialization
#
# ### 3 Convolution Layers of size 3x3, 5x5 and 3x3 having rectified linear Activation function followed by Max Pool Layer of size 2x2 followed by dense layer with dropout of 0.25.
# +
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(200, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
# -
# # Fit Model and compute accuracy score
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
|
Project 1 - Part A Submission Files/Project 1 - Part A - L18-1845.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Project Topic Classification
# run this statement only once to install Rake
# !pip install rake_nltk
# !pip install nltk
# +
import numpy as np
import pandas as pd
from rake_nltk import Rake
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.feature_extraction.text import CountVectorizer
import re, nltk, gensim
nltk.download('wordnet')
from nltk.tokenize import ToktokTokenizer
from nltk.stem import wordnet
from nltk.corpus import stopwords
from string import punctuation
# -
# ### Step 1: Read in and analyse the data
# +
import pandas as pd
import glob
path = r'./data/' # use your path
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
frame = pd.concat(li, axis=0, ignore_index=True)
# df = pd.read_csv('Kickstarter057.csv')
df = frame
df.head()
# +
def extract_cat(text):
text = text.split(",")
text = text[2]
text = text.replace ("/", " ")
text = text.replace ("name", "")
text = text.replace ("slug", "")
text = text.replace ('"', "")
text = text.replace ('{', "")
text = text.replace (':', "")
text = text.lower()
text = re.sub(r"\'\n", " ", text)
text = re.sub(r"\'\xa0", " ", text)
text = re.sub('\s+', ' ', text) # matches all whitespace characters
text = text.strip(' ')
return text
df['category'] = df['category'].apply(lambda x: extract_cat(x))
df.head()
# +
df = df[['name','category','blurb']]
df.head()
# +
import nltk
from nltk.tokenize import RegexpTokenizer
from nltk.stem import WordNetLemmatizer,PorterStemmer
from nltk.corpus import stopwords
import re
lemmatizer = WordNetLemmatizer()
stemmer = PorterStemmer()
def preprocess(sentence):
sentence=str(sentence)
sentence = sentence.lower()
sentence=sentence.replace('{html}',"")
cleanr = re.compile('<.*?>')
cleantext = re.sub(cleanr, '', sentence)
rem_url=re.sub(r'http\S+', '',cleantext)
rem_num = re.sub('[0-9]+', '', rem_url)
tokenizer = RegexpTokenizer(r'\w+')
tokens = tokenizer.tokenize(rem_num)
filtered_words = [w for w in tokens if len(w) > 2 if not w in stopwords.words('english')]
stem_words=[stemmer.stem(w) for w in filtered_words]
lemma_words=[lemmatizer.lemmatize(w) for w in stem_words]
return " ".join(filtered_words)
df['blurb']=df['blurb'].map(lambda s:preprocess(s))
df.head()
# +
#Tokenize everything in the category
dfcat = df['category']
dfcat.head
num_dfcat = len(dfcat)
#print(num_dfcat)
from nltk.tokenize import word_tokenize
dfcattok = dfcat.apply(word_tokenize)
dfcattok.head()
#for loop each one and collect the first value. This shows the main categories that we have derived from our own dataset.
maincat_list = []
for x in dfcattok:
if x[0] not in maincat_list:
maincat_list.append(x[0])
print(maincat_list)
# +
#Load dataset
import pandas as pd
import glob
#Dataset
from nltk.corpus import PlaintextCorpusReader
art = PlaintextCorpusReader('data/Train/Art', '.+\.txt')
tech = PlaintextCorpusReader('data/Train/Tech', '.+\.txt')
comics = PlaintextCorpusReader('data/Train/Comics', '.+\.txt')
film = PlaintextCorpusReader('data/Train/Film', '.+\.txt')
music = PlaintextCorpusReader('data/Train/Music', '.+\.txt')
photography = PlaintextCorpusReader('data/Train/Photography', '.+\.txt')
publishing = PlaintextCorpusReader('data/Train/Publishing', '.+\.txt')
art_docs1 = [art.words(fid) for fid in art.fileids()]
tech_docs1 = [tech.words(fid) for fid in tech.fileids()]
comics_docs1 = [comics.words(fid) for fid in comics.fileids()]
film_docs1 = [film.words(fid) for fid in film.fileids()]
music_docs1 = [music.words(fid) for fid in music.fileids()]
photography_docs1 = [photography.words(fid) for fid in photography.fileids()]
publishing_docs1 = [publishing.words(fid) for fid in publishing.fileids()]
print(art_docs1[0][0:20])
print(tech_docs1[0][0:20])
print(comics_docs1[0][0:20])
print(film_docs1[0][0:20])
print(music_docs1[0][0:20])
print(photography_docs1[0][0:20])
print(publishing_docs1[0][0:20])
# +
###Basically preprocessing date from dataset
# Combine the categories of the corpus
all_docs1 = art_docs1 + tech_docs1 + comics_docs1 + film_docs1 + music_docs1 + photography_docs1 + publishing_docs1
num_art_docs = len(art_docs1)
num_2 = len(art_docs1) + len(tech_docs1)
num_3 = num_2 + len(comics_docs1)
num_4 = num_3 + len(film_docs1)
num_5 = num_4 + len(music_docs1)
num_6 = num_5 + len(photography_docs1)
#For verifying the whether the output in dictionary is correct
print(num_art_docs)
print (len(tech_docs1))
print (len(comics_docs1))
print (len(film_docs1))
print (len(music_docs1))
print (len(photography_docs1))
# Processsing for stopwords, alphabetic words, Stemming
all_docs2 = [[w.lower() for w in doc] for doc in all_docs1]
import re
all_docs3 = [[w for w in doc if re.search('^[a-z]+$',w)] for doc in all_docs2]
from nltk.corpus import stopwords
stop_list = stopwords.words('english')
all_docs4 = [[w for w in doc if w not in stop_list] for doc in all_docs3]
from nltk.stem.porter import *
stemmer = PorterStemmer()
all_docs5 = [[stemmer.stem(w) for w in doc] for doc in all_docs4]
#Create dictionary
from gensim import corpora
dictionary = corpora.Dictionary(all_docs5)
print(dictionary)
# Convert all documents to TF Vectors
all_tf_vectors = [dictionary.doc2bow(doc) for doc in all_docs5]
#Label the trained data. Since the folder name is the label, I use the same labels.
all_data_as_dict = [{id:1 for (id, tf_value) in vec} for vec in all_tf_vectors]
print(type(all_data_as_dict))
#print(all_data_as_dict). The labels are generated by our own dataset and used here.
art_data = [(d, 'art') for d in all_data_as_dict[0:num_art_docs]] #First document to number of art documents, which is 4. Document 0-4
tech_data = [(d, 'tech') for d in all_data_as_dict[num_art_docs:num_2]]
comics_data = [(d, 'comics') for d in all_data_as_dict[num_2:num_3]]
film_data = [(d, 'film') for d in all_data_as_dict[num_3:num_4]]
music_data = [(d, 'music') for d in all_data_as_dict[num_4:num_5]]
photography_data = [(d, 'photography') for d in all_data_as_dict[num_5:num_6]]
publishing_data = [(d, 'publishing') for d in all_data_as_dict[num_6:]]
all_labeled_data = art_data + tech_data + comics_data + film_data + music_data + photography_data + publishing_data
#Generate the trained classifier
classifier = nltk.NaiveBayesClassifier.train(all_labeled_data)
test_doc = all_data_as_dict[200]
#print(all_data_as_dict[0])
print(classifier.classify(test_doc))
# +
### Validate
# Read the files in validate folder and preparing the validation corpus
art_validation = PlaintextCorpusReader('data/Validate/Art', '.+\.txt')
tech_validation = PlaintextCorpusReader('data/Validate/Tech', '.+\.txt')
comics_validation = PlaintextCorpusReader('data/Validate/Comics', '.+\.txt')
film_validation = PlaintextCorpusReader('data/Validate/Film', '.+\.txt')
music_validation = PlaintextCorpusReader('data/Validate/Music', '.+\.txt')
photography_validation = PlaintextCorpusReader('data/Validate/Photography', '.+\.txt')
publishing_validation = PlaintextCorpusReader('data/Validate/Publishing', '.+\.txt')
# Tokenization
art_valid_docs1 = [art_validation.words(fid) for fid in art_validation.fileids()]
tech_valid_docs1 = [tech_validation.words(fid) for fid in tech_validation.fileids()]
comics_valid_docs1 = [comics_validation.words(fid) for fid in comics_validation.fileids()]
film_valid_docs1 = [film_validation.words(fid) for fid in film_validation.fileids()]
music_valid_docs1 = [music_validation.words(fid) for fid in music_validation.fileids()]
photography_valid_docs1 = [photography_validation.words(fid) for fid in photography_validation.fileids()]
publishing_valid_docs1 = [publishing_validation.words(fid) for fid in publishing_validation.fileids()]
# Combine the two sets of documents for easy processing.
all_valid_docs = art_valid_docs1 + tech_valid_docs1 + comics_valid_docs1 + film_valid_docs1 + music_valid_docs1 + photography_valid_docs1 + publishing_valid_docs1
# This number will be used to separate the two sets of documents later.
num_art_valid_docs = len(art_valid_docs1)
num_valid_2 = num_art_valid_docs + len(tech_valid_docs1)
num_valid_3 = num_valid_2 + len(comics_valid_docs1)
num_valid_4 = num_valid_3 + len(film_valid_docs1)
num_valid_5 = num_valid_4 + len(music_valid_docs1)
num_valid_6 = num_valid_5 + len(photography_valid_docs1)
# Text pre-processing, including stop word removal, stemming, etc.
all_valid_docs2 = [[w.lower() for w in doc] for doc in all_valid_docs]
all_valid_docs3 = [[w for w in doc if re.search('^[a-z]+$',w)] for doc in all_valid_docs2]
all_valid_docs4 = [[w for w in doc if w not in stop_list] for doc in all_valid_docs3]
all_valid_docs5 = [[stemmer.stem(w) for w in doc] for doc in all_valid_docs4]
# Note that we're using the dictionary created earlier.
all_valid_tf_vectors = [dictionary.doc2bow(doc) for doc in all_valid_docs5]
# Convert documents into dict representation.
all_valid_data_as_dict = [{id:1 for (id, tf_value) in vec} for vec in all_valid_tf_vectors]
# Separate the two sets of documents and add labels.
art_valid_data_with_labels = [(d, 'art') for d in all_valid_data_as_dict[0:num_art_valid_docs]]
tech_valid_data_with_labels = [(d, 'tech') for d in all_valid_data_as_dict[num_art_valid_docs:num_valid_2]]
comics_valid_data_with_labels = [(d, 'comics') for d in all_valid_data_as_dict[num_valid_2:num_valid_3]]
film_valid_data_with_labels = [(d, 'film') for d in all_valid_data_as_dict[num_valid_3:num_valid_4]]
music_valid_data_with_labels = [(d, 'music') for d in all_valid_data_as_dict[num_valid_4:num_valid_5]]
photography_valid_data_with_labels = [(d, 'photography') for d in all_valid_data_as_dict[num_valid_5:num_valid_6]]
publishing_valid_data_with_labels = [(d, 'publishing') for d in all_valid_data_as_dict[num_valid_6:]]
# Combine the labeled documents.
all_valid_data_with_labels = art_valid_data_with_labels + tech_valid_data_with_labels + comics_valid_data_with_labels + film_valid_data_with_labels + music_valid_data_with_labels + photography_valid_data_with_labels + publishing_valid_data_with_labels
# -
print(nltk.classify.accuracy(classifier, all_valid_data_with_labels))
# ## Mode Testing - Predicting labels for other documents
# +
#Read the text files
test_corpus = PlaintextCorpusReader('data/Test', '.+\.txt')
fids = test_corpus.fileids()
# Tokenization
test_docs1 = [test_corpus.words(fid) for fid in fids]
# Text pre-processing, including stop word removal, stemming, etc.
test_docs2 = [[w.lower() for w in doc] for doc in test_docs1]
test_docs3 = [[w for w in doc if re.search('^[a-z]+$',w)] for doc in test_docs2]
test_docs4 = [[w for w in doc if w not in stop_list] for doc in test_docs3]
test_docs5 = [[stemmer.stem(w) for w in doc] for doc in test_docs4]
# Note that we're using the dictionary created earlier to create TF vectors
test_tf_vectors = [dictionary.doc2bow(doc) for doc in test_docs5]
# Convert documents into dict representation. This is document-label representation
test_data_as_dict = [{id:1 for (id, tf_value) in vec} for vec in test_tf_vectors]
#For each file, classify and print the label.
for i in range(len(fids)):
print(fids[i], '-->', classifier.classify(test_data_as_dict[i]),'and', classifier.classify(test_data_as_dict[i+1]) )
# -
|
src/Classification-NB-Model/Final.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import seaborn as sns
import metapack as mp
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import display
from dtcv import get_image
import cv2
# %matplotlib inline
sns.set_context('notebook')
mp.jupyter.init()
# +
#pkg = mp.jupyter.open_package()
#pkg = mp.jupyter.open_source_package()
pkg = mp.open_package('http://library.metatab.org/sandiegodata.org-downtown_cv-5.zip')
pkg
# -
display(pkg.resource('counts'))
counts = pkg.resource('counts').dataframe()
# +
# This may take a few minutes; it will download about 330 images and save them to the /tmp directory
counts['image'] = counts.image_url.apply(get_image)
counts['count'] = pd.to_numeric(counts['count'])
# +
def crop(row):
"""Crop the handwritten mark, and hopefully the shape around it, from the image"""
x, y, r = row.cx, row.cy, row.r
r = int(r*1.0)
return row.image[y-r:y+r, x-r:x+r ]
plt.imshow(crop(counts.iloc[60]))
# +
numbers = []
def invert(img):
# Convert the img to grayscale
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
r = cv2.resize(gray, (100,100))
_, tr = cv2.threshold (r, 70, 255, cv2.THRESH_BINARY_INV);
return tr
for name, row in counts.iterrows():
if row['count'] is not None and not np.isnan(row['count']) and row['count'] < 10 :
numbers.append((row['count'], invert(crop(row))))
print(len(numbers))
plt.imshow(numbers[0][1])
# +
'''Trains a simple convnet on the MNIST dataset.
Gets to 99.25% test accuracy after 12 epochs
(there is still a lot of margin for parameter tuning).
16 seconds per epoch on a GRID K520 GPU.
'''
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
batch_size = 128
epochs = 12
# the data, split between train and test sets
#(x_train, y_train), (x_test, y_test) = mnist.load_data()
# input image dimensions
#img_rows, img_cols = 28, 28
X = np.array([e[1] for e in numbers])
y = np.array([e[0] for e in numbers])
num_classes = len(np.unique(y))+1
l = len(X)
train_l = int(l*.9)
x_train = X[:train_l]
y_train = y[:train_l]
x_test = X[train_l:]
y_test = y[train_l:]
img_rows, img_cols = x_train[0].shape
# +
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# +
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),activation='relu',input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
|
ml-notebooks/HomelessDigits.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# name: python3
# ---
# # Build Classification Models
# +
import pandas as pd
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import accuracy_score,precision_score,confusion_matrix,classification_report, precision_recall_curve
from sklearn.svm import SVC
import numpy as np
cuisines_df = pd.read_csv("../data/cleaned_cuisines_own.csv")
cuisines_df.head()
# -
cuisines_label_df = cuisines_df['cuisine']
cuisines_label_df.head()
cuisines_features_df = cuisines_df.drop(["Unnamed: 0", "cuisine"], axis=1)
cuisines_features_df.head()
X_train, X_test, y_train, y_test = train_test_split(cuisines_features_df, cuisines_label_df, test_size=0.3)
# +
lr = LogisticRegression(multi_class="ovr", solver="lbfgs")
model = lr.fit(X_train, np.ravel(y_train))
accuracy = model.score(X_test, y_test)
print ("Accuracy is {}".format(accuracy))
# -
print(f'ingredients: {X_test.iloc[50][X_test.iloc[50]!=0].keys()}')
print(f'cuisine: {y_test.iloc[50]}')
# +
test= X_test.iloc[50].values.reshape(-1, 1).T
proba = model.predict_proba(test)
classes = model.classes_
resultdf = pd.DataFrame(data=proba, columns=classes)
topPrediction = resultdf.T.sort_values(by=[0], ascending = [False])
topPrediction.head()
# -
y_pred = model.predict(X_test)
print(classification_report(y_test,y_pred))
|
4-Classification/2-Classifiers-1/notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" jupyter={"outputs_hidden": true}
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import os
import keras
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout, BatchNormalization
from PIL import Image
import matplotlib.pyplot as plt
plt.style.use('dark_background')
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
import tensorflow as tf
from tensorflow import keras
import cv2 as cv
import glob as gb
from keras.models import Model
from sklearn.utils import shuffle
from keras.callbacks import EarlyStopping
from keras.preprocessing.image import ImageDataGenerator
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
# -
encoder = OneHotEncoder()
encoder.fit([[0], [1]])
# +
data = []
paths = []
result = []
for r, d, f in os.walk(r'../input/brain-mri-images-for-brain-tumor-detection/yes'):
for file in f:
if '.jpg' in file:
paths.append(os.path.join(r, file))
for path in paths:
img = Image.open(path)
img = img.resize((128,128))
img = np.array(img)
if(img.shape == (128,128,3)):
data.append(np.array(img))
result.append(encoder.transform([[0]]).toarray())
# -
# # Updates result list for images with tumor. (up)
# +
paths = []
for r, d, f in os.walk(r"../input/brain-mri-images-for-brain-tumor-detection/no"):
for file in f:
if '.jpg' in file:
paths.append(os.path.join(r, file))
for path in paths:
img = Image.open(path)
img = img.resize((128,128))
img = np.array(img)
if(img.shape == (128,128,3)):
data.append(np.array(img))
result.append(encoder.transform([[1]]).toarray())
# -
# # Updates result list for images without tumor. (up)
data = np.array(data)
data.shape
result = np.array(result)
result = result.reshape(139,2)
x_train,x_test,y_train,y_test = train_test_split(data, result, test_size=0.2, shuffle=True, random_state=0)
# +
model = Sequential()
model.add(Conv2D(32, kernel_size=(2, 2), input_shape=(128, 128, 3), padding = 'Same'))
model.add(Conv2D(32, kernel_size=(2, 2), activation ='relu', padding = 'Same'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, kernel_size = (2,2), activation ='relu', padding = 'Same'))
model.add(Conv2D(64, kernel_size = (2,2), activation ='relu', padding = 'Same'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(2, activation='softmax'))
model.compile(loss = "categorical_crossentropy", optimizer='Adamax')
print(model.summary())
# -
y_train.shape
history = model.fit(x_train, y_train, epochs = 10, batch_size = 40, verbose = 1,validation_data = (x_test, y_test))
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Test', 'Validation'], loc='upper right')
plt.show()
def names(number):
if number==0:
return 'Its a Tumor'
else:
return 'No, Its not a tumor'
img = Image.open(r"../input/brain-mri-images-for-brain-tumor-detection/no/19 no.jpg")
plt.axis('on')
x = np.array(img.resize((128,128)))
x = x.reshape(1,128,128,3)
res = model.predict_on_batch(x)
classification = np.where(res == np.amax(res))[1][0]
imshow(img)
print(str(res[0][classification]*100) + '% Confirmed, This Is ' + names(classification))
# +
img = Image.open(r"../input/brain-mri-images-for-brain-tumor-detection/yes/Y4.jpg")
x = np.array(img.resize((128,128)))
x = x.reshape(1,128,128,3)
res = model.predict_on_batch(x)
classification = np.where(res == np.amax(res))[1][0]
imshow(img)
print(str(res[0][classification]*100) + '% Confirmed, This Is A, ' + names(classification))
# -
|
Brain MRI Images for Brain Tumor Detection/brain-mri-images-for-brain-tumor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
from pprint import pprint
import os
from github import Github
GITHUB_USERNAME = os.environ.get("GITHUB_USERNAME")
GITHUB_PASSWORD = os.environ.get("GITHUB_PASSWORD")
session = Github(GITHUB_USERNAME, GITHUB_PASSWORD)
# -
import urllib
from urllib.parse import quote
repos = session.search_repositories("utils+language:python")
print(repos.totalCount)
repositories = []
for page in (repos.get_page(num) for num in range(10)):
repositories.extend(page)
len(repositories)
code_files = []
for repo in repositories[:3]:
code_files.extend(session.search_code(quote(f"test repository:{repo.full_name} language:python extension:.py", safe="")))
repositories
|
medusa/test_ground.py.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="baIUIfrChJcY" colab_type="text"
# ##### Copyright 2018 The TensorFlow Authors.
# + [markdown] colab_type="text" id="Ka96-ajYzxVU"
# # Train Your Own Model and Convert It to TFLite
#
# This notebook uses the [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist) dataset which contains 70,000 grayscale images in 10 categories. The images show individual articles of clothing at low resolution (28 by 28 pixels), as seen here:
#
# <table>
# <tr><td>
# <img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
# alt="Fashion MNIST sprite" width="600">
# </td></tr>
# <tr><td align="center">
# <b>Figure 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">Fashion-MNIST samples</a> (by Zalando, MIT License).<br/>
# </td></tr>
# </table>
#
# Fashion MNIST is intended as a drop-in replacement for the classic [MNIST](http://yann.lecun.com/exdb/mnist/) dataset—often used as the "Hello, World" of machine learning programs for computer vision. The MNIST dataset contains images of handwritten digits (0, 1, 2, etc.) in a format identical to that of the articles of clothing we'll use here.
#
# This uses Fashion MNIST for variety, and because it's a slightly more challenging problem than regular MNIST. Both datasets are relatively small and are used to verify that an algorithm works as expected. They're good starting points to test and debug code.
#
# We will use 60,000 images to train the network and 10,000 images to evaluate how accurately the network learned to classify images. You can access the Fashion MNIST directly from TensorFlow. Import and load the Fashion MNIST data directly from TensorFlow:
# + [markdown] colab_type="text" id="rjOAfhgd__Sp"
# # Setup
# + colab_type="code" id="pfyZKowNAQ4j" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="b5b29e83-3d06-4a02-f9e2-f029f849e14f"
# TensorFlow
import tensorflow as tf
# TensorFlow Datsets
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
# Helper Libraries
import numpy as np
import matplotlib.pyplot as plt
import pathlib
from os import getcwd
print('\u2022 Using TensorFlow Version:', tf.__version__)
print('\u2022 GPU Device Found.' if tf.test.is_gpu_available() else '\u2022 GPU Device Not Found. Running on CPU')
# + [markdown] colab_type="text" id="tadPBTEiAprt"
# # Download Fashion MNIST Dataset
#
# We will use TensorFlow Datasets to load the Fashion MNIST dataset.
# + colab_type="code" id="XcNwi6nFKneZ" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="a6a0d520-d45a-4db7-e3d4-df22be36df25"
#splits = tfds.Split.ALL.subsplit(weighted=(80, 10, 10))
filePath = f"{getcwd()}/../tmp2/"
(train_examples, validation_examples, test_examples), info = tfds.load('fashion_mnist', with_info=True, as_supervised=True, split=['train[:80%]', 'train[80%:90%]', 'train[90%:]'], data_dir=filePath)
num_examples = info.splits['train'].num_examples
num_classes = info.features['label'].num_classes
# + [markdown] id="H-Is76GzhJcq" colab_type="text"
# The class names are not included with the dataset, so we will specify them here.
# + colab_type="code" id="-eAv71FRm4JE" colab={}
class_names = ['T-shirt_top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + colab_type="code" id="hXe6jNokqX3_" colab={}
# Create a labels.txt file with the class names
with open('labels.txt', 'w') as f:
f.write('\n'.join(class_names))
# + colab_type="code" id="iubWCThbdN8K" colab={}
# The images in the dataset are 28 by 28 pixels.
IMG_SIZE = 28
# + [markdown] colab_type="text" id="ZAkuq0V0Aw2X"
# # Preprocessing Data
# + [markdown] colab_type="text" id="_5SIivkunKCC"
# ## Preprocess
# + id="xI9cm5fKhJc4" colab_type="code" colab={}
# EXERCISE: Write a function to normalize the images.
def format_example(image, label):
# Cast image to float32
image = tf.cast(image, tf.float32)
# Normalize the image in the range [0, 1]
image = image/255
return image, label
# + colab_type="code" id="HAlBlXOUMwqe" colab={}
# Specify the batch size
BATCH_SIZE = 256
# + [markdown] colab_type="text" id="JM4HfIJtnNEk"
# ## Create Datasets From Images and Labels
# + id="1UiE1cGghJdB" colab_type="code" colab={}
# Create Datasets
train_batches = train_examples.cache().shuffle(num_examples//4).batch(BATCH_SIZE).map(format_example).prefetch(1)
validation_batches = validation_examples.cache().batch(BATCH_SIZE).map(format_example)
test_batches = test_examples.map(format_example).batch(1)
# + [markdown] colab_type="text" id="M-topQaOm_LM"
# # Building the Model
# + [markdown] id="VeaNwp16hJdF" colab_type="text"
# ```
# Model: "sequential"
# _________________________________________________________________
# Layer (type) Output Shape Param #
# =================================================================
# conv2d (Conv2D) (None, 26, 26, 16) 160
# _________________________________________________________________
# max_pooling2d (MaxPooling2D) (None, 13, 13, 16) 0
# _________________________________________________________________
# conv2d_1 (Conv2D) (None, 11, 11, 32) 4640
# _________________________________________________________________
# flatten (Flatten) (None, 3872) 0
# _________________________________________________________________
# dense (Dense) (None, 64) 247872
# _________________________________________________________________
# dense_1 (Dense) (None, 10) 650
# =================================================================
# Total params: 253,322
# Trainable params: 253,322
# Non-trainable params: 0
# ```
# + id="rEUk7QdehJdF" colab_type="code" colab={}
# EXERCISE: Build and compile the model shown in the previous cell.
model = tf.keras.Sequential([
# Set the input shape to (28, 28, 1), kernel size=3, filters=16 and use ReLU activation,
tf.keras.layers.Conv2D(16, (3, 3), input_shape=(28, 28, 1), activation='relu'),
tf.keras.layers.MaxPooling2D((2, 2)),
# Set the number of filters to 32, kernel size to 3 and use ReLU activation
tf.keras.layers.Conv2D(32, (3, 3), activation='relu'),
# Flatten the output layer to 1 dimension
tf.keras.layers.Flatten(),
# Add a fully connected layer with 64 hidden units and ReLU activation
tf.keras.layers.Dense(units=64, activation='relu'),
# Attach a final softmax classification head
tf.keras.layers.Dense(units=10, activation='softmax')
])
# Set the appropriate loss function and use accuracy as your metric
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=["accuracy"])
# + [markdown] colab_type="text" id="zEMOz-LDnxgD"
# ## Train
# + colab_type="code" id="JGlNoRtzCP4_" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="1e89851e-b841-47de-a04d-12a0e0f293f8"
history = model.fit(train_batches, epochs=10, validation_data=validation_batches)
# + [markdown] colab_type="text" id="TZT9-7w9n4YO"
# # Exporting to TFLite
#
# You will now save the model to TFLite. We should note, that you will probably see some warning messages when running the code below. These warnings have to do with software updates and should not cause any errors or prevent your code from running.
# + id="DEfvdoWlhJdO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 171} outputId="6596a8c7-884f-4570-8723-027945b0e828"
# EXERCISE: Use the tf.saved_model API to save your model in the SavedModel format.
export_dir = 'saved_model/1'
# YOUR CODE HERE
tf.saved_model.save(model, export_dir)
# + cellView="form" colab_type="code" id="EDGiYrBdE6fl" colab={}
# Select mode of optimization
mode = "Speed"
if mode == 'Storage':
optimization = tf.lite.Optimize.OPTIMIZE_FOR_SIZE
elif mode == 'Speed':
optimization = tf.lite.Optimize.OPTIMIZE_FOR_LATENCY
else:
optimization = tf.lite.Optimize.DEFAULT
# + id="RzP6oFMBhJdU" colab_type="code" colab={}
# EXERCISE: Use the TFLiteConverter SavedModel API to initialize the converter
converter = tf.lite.TFLiteConverter.from_saved_model(export_dir)
# Set the optimzations
converter.optimizations = [optimization]
# Invoke the converter to finally generate the TFLite model
tflite_model = converter.convert()
# + colab_type="code" id="q5PWCDsTC3El" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="570ae85b-c02e-4942-c7b6-b2c5354c6025"
tflite_model_file = pathlib.Path('./model.tflite')
tflite_model_file.write_bytes(tflite_model)
# + [markdown] colab_type="text" id="SR6wFcQ1Fglm"
# # Test the Model with TFLite Interpreter
# + colab_type="code" id="rKcToCBEC-Bu" colab={}
# Load TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_content=tflite_model)
interpreter.allocate_tensors()
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
# + colab_type="code" id="E8EpFpIBFkq8" colab={}
# Gather results for the randomly sampled test images
predictions = []
test_labels = []
test_images = []
for img, label in test_batches.take(50):
interpreter.set_tensor(input_index, img)
interpreter.invoke()
predictions.append(interpreter.get_tensor(output_index))
test_labels.append(label[0])
test_images.append(np.array(img))
# + cellView="form" colab_type="code" id="kSjTmi05Tyod" colab={}
# Utilities functions for plotting
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
img = np.squeeze(img)
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label.numpy():
color = 'green'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks(list(range(10)))
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array[0], color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array[0])
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# + cellView="form" colab_type="code" id="ZZwg0wFaVXhZ" colab={"base_uri": "https://localhost:8080/", "height": 211} outputId="d390810d-fa6b-4c8c-ec20-968e57a0db94"
# Visualize the outputs
# Select index of image to display. Minimum index value is 1 and max index value is 50.
index = 49
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(index, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(index, predictions, test_labels)
plt.show()
# + [markdown] id="tD0IAnQShJdy" colab_type="text"
# # Click the Submit Assignment Button Above
#
# You should now click the Submit Assignment button above to submit your notebook for grading. Once you have submitted your assignment, you can continue with the optinal section below.
#
# ## If you are done, please **don't forget to run the last two cells of this notebook** to save your work and close the Notebook to free up resources for your fellow learners.
# + [markdown] colab_type="text" id="H8t7_jRiz9Vw"
# # Prepare the Test Images for Download (Optional)
# + colab_type="code" id="Fi09nIps0gBu" colab={}
# !mkdir -p test_images
# + colab_type="code" id="sF7EZ63J0hZs" colab={}
from PIL import Image
for index, (image, label) in enumerate(test_batches.take(50)):
image = tf.cast(image * 255.0, tf.uint8)
image = tf.squeeze(image).numpy()
pil_image = Image.fromarray(image)
pil_image.save('test_images/{}_{}.jpg'.format(class_names[label[0]].lower(), index))
# + colab_type="code" id="uM35O-uv0iWS" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="50076c29-8335-4e23-b65e-979004b2ddfa"
# !ls test_images
# + colab_type="code" id="aR20r4qW0jVm" colab={}
# !tar --create --file=fmnist_test_images.tar test_images
# + id="er_K3nYehJeG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="0deca614-6e04-490c-898d-580b1adc17c5"
# !ls
|
Tensorflow Data and Deployment/Course 2 - Device-based Models with TensorFlow Lite/TFLite_FashinMNIST.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Continuous Control
#
# ---
#
# You are welcome to use this coding environment to train your agent for the project. Follow the instructions below to get started!
#
# ### 1. Start the Environment
#
# Run the next code cell to install a few packages. This line will take a few minutes to run!
# !pip -q install ./python
# The environments corresponding to both versions of the environment are already saved in the Workspace and can be accessed at the file paths provided below.
#
# Please select one of the two options below for loading the environment.
# +
from unityagents import UnityEnvironment
import numpy as np
# select this option to load version 1 (with a single agent) of the environment
#env = UnityEnvironment(file_name='/data/Reacher_One_Linux_NoVis/Reacher_One_Linux_NoVis.x86_64')
# select this option to load version 2 (with 20 agents) of the environment
env = UnityEnvironment(file_name='/data/Reacher_Linux_NoVis/Reacher.x86_64')
# -
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# ### 2. Examine the State and Action Spaces
#
# Run the code cell below to print some information about the environment.
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
# -
# ### 3. Take Random Actions in the Environment
#
# In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
#
# Note that **in this coding environment, you will not be able to watch the agents while they are training**, and you should set `train_mode=True` to restart the environment.
env_info = env.reset(train_mode=True)[brain_name] # reset the environment
states = env_info.vector_observations # get the current state (for each agent)
scores = np.zeros(num_agents) # initialize the score (for each agent)
while True:
actions = np.random.randn(num_agents, action_size) # select an action (for each agent)
actions = np.clip(actions, -1, 1) # all actions between -1 and 1
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
scores += env_info.rewards # update the score (for each agent)
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
print('Total score (averaged over agents) this episode: {}'.format(np.mean(scores)))
# When finished, you can close the environment.
# ### 4. Building the DDPG model
# First we specifiy our hyper-parameters that we will use in this project.
# Then we build the neural networks that will act as the actor and critic models
# +
import numpy as np
import random
import copy
from collections import namedtuple, deque
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.nn.utils import clip_grad_norm_
# hyper-parameter definitions:
HIDDEN_LAYERS = [400, 300] # Hidden layer sizes of the actor and critic networks
LR_A = 1e-4 # learning rate for the actor
LR_C = 1e-3 # learning rate for the critic
DISCOUNT_RATE = 0.99 # discount factor
TAU = 1e-3 # soft update factor
BUFFER_SIZE = int(1e6) # buffer size for replay buffer
BATCH_SIZE = 128 # batch size
WEIGHT_DECAY = 0.0 # L2 weight decay
NOISE_INITIAL = 1.0 # Initial noise factor
NOISE_DECAY = 0.999999 # Noise factor
NOISE_MIN = 0.0 # Minimum noise factor
T_UPDATE = 20 # Number of timesteps between updating the target network
NUM_UPDATES = 10 # Amount of times to update the network every T_UPDATE timesteps
EPS_MAX = 2000 # Number of max episodes to train to TARGET_MEAN_SCORE
T_MAX = 1000 # Number of max timesteps per episode
TARGET_MEAN_SCORE = 30.0 # target score for the enviroment for 'success'
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# -
# We specify the actor and critic networks
# +
def hidden_init(layer):
fan_in = layer.weight.data.size()[0]
lim = 1. / np.sqrt(fan_in)
return (-lim, lim)
class Actor(nn.Module):
""" Actor(policy) model """
def __init__(self, state_size, action_size, seed, layers):
"""Initialize parameters for model
Parameters:
state_size (int): dimensions of the state space
action_size (int): dimensions of the action space
seed (int): random seed
layers (int list): size of the hidden layers
"""
super(Actor, self).__init__()
torch.seed = torch.manual_seed(seed)
self.bns = nn.ModuleList([nn.BatchNorm1d(t) for t in layers])
layers = [state_size] + layers + [action_size]
self.layers = nn.ModuleList([nn.Linear(fst, snd) for (fst, snd) in zip(layers[:-1], layers[1:])])
self.reset_parameters()
def reset_parameters(self):
""" Set initial weights """
for layer in self.layers[:-1]:
layer.weight.data.uniform_(*hidden_init(layer))
self.layers[-1].weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state):
"""
Builds an actor(policy) network that maps state to actions
"""
x = F.relu(self.bns[0](self.layers[0](state)))
for (layer, norm) in zip(self.layers[1:-1], self.bns[1:]):
#for layer in self.layers[1:-1]:
x = layer(x)
x = norm(x)
x = F.relu(x)
return F.tanh(self.layers[-1](x))
class Critic(nn.Module):
""" Critic(value) network """
def __init__(self, state_size, action_size, seed, layers):
"""Initialize parameters for model
Parameter:
state_size (int): dimensions of the state space
action_size (int): dimensions of the action space
seed (int): random seed
layers (int list): size of the hidden layers
"""
super(Critic, self).__init__()
self.seed = torch.manual_seed(seed)
# ensure that we have atleast 1 hidden layer for action to come after state in the network
assert len(layers) >= 1
self.bns = nn.ModuleList([nn.BatchNorm1d(t) for t in layers])
self.layers = nn.ModuleList([nn.Linear(state_size, layers[0])])
layers = layers + [1]
layers[0] += action_size
self.layers.extend([nn.Linear(fst, snd) for (fst, snd) in zip(layers[:-1], layers[1:])])
self.reset_parameters()
def reset_parameters(self):
""" Set initial weights """
for layer in self.layers[:-1]:
layer.weight.data.uniform_(*hidden_init(layer))
self.layers[-1].weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state, action):
"""
Builds an critic(value) network that maps state and actions to their value
"""
x = F.relu(self.bns[0](self.layers[0](state)))
# we concat here to attach the actions to the output of the first hidden layer
x = torch.cat((x, action), dim=1)
for (layer, norm) in zip(self.layers[1:-1], self.bns[1:]):
x = layer(x)
x = norm(x)
x = F.relu(x)
return self.layers[-1](x)
# -
# Defining the DDPG agent that will be trained using experience replay and soft updates on target networks
# +
class Agent():
"""Interacts with and learns from the enviroment"""
def __init__(self, num_agents, state_size, action_size, random_seed):
"""Initialize an agent
Parameters:
state_size (int): dimensions of state space
action_size (int): dimension of action space
random_seed (int): random seed
"""
self.state_size = state_size
self.action_size = action_size
self.seed = random.seed(random_seed)
# Actor networks with local and target
self.actor_local = Actor(state_size, action_size, random_seed, HIDDEN_LAYERS).to(device)
self.actor_target = Actor(state_size, action_size, random_seed, HIDDEN_LAYERS).to(device)
self.actor_optim = optim.Adam(self.actor_local.parameters(), lr=LR_A)
# Critic networks with local and target
self.critic_local = Critic(state_size, action_size, random_seed, HIDDEN_LAYERS).to(device)
self.critic_target = Critic(state_size, action_size, random_seed, HIDDEN_LAYERS).to(device)
self.critic_optim = optim.Adam(self.critic_local.parameters(), lr=LR_C)
# Noise process
self.noise = OUNoise(num_agents, action_size, random_seed)
self.noise_factor = NOISE_INITIAL
# Replay memory
self.memory = ReplayBuffer(action_size, BUFFER_SIZE, BATCH_SIZE, random_seed)
def step(self, timestep, state, action, reward, next_state, done):
# Save experiences in replay memory and use random sampling from buffer to learn
# Save experience / reward
self.memory.add(state, action, reward, next_state, done)
# Learn if enough samples are available in memory
if len(self.memory) > BATCH_SIZE and timestep % T_UPDATE == 0:
for _ in range(NUM_UPDATES):
experiences = self.memory.sample()
self.learn(experiences, DISCOUNT_RATE)
def act(self, state, add_noise=True):
"""Returns actions for given state as per current policy"""
state = torch.from_numpy(state).float().to(device)
self.actor_local.eval()
with torch.no_grad():
action = self.actor_local(state).cpu().data.numpy()
self.actor_local.train()
if add_noise:
action += self.noise.sample() * self.noise_factor
#self.noise_factor = max(self.noise_factor * NOISE_DECAY, NOISE_MIN)
return np.clip(action, -1, 1)
def reset(self):
""" Reset learning behavior episodically """
self.noise.reset()
def learn(self, experiences, gamma):
"""Update policy and value parameters using given batch of experience tuples:
Q_target = r + gamma * critic_target(next_state, actor_target(next_state))
where:
actor_target(state) -> action
critic_target(state, action) -> Q-value
Parameters:
experiences (Tuple[torch.Tensor]): tuple of (s, a, r, s', done) tuples
gamma (float): discount rate
"""
states, actions, rewards, next_states, dones = experiences
# Update critic
# Get predicted next-(state and actions) and Q-values from target models
actions_next = self.actor_target(next_states)
Q_targets_next = self.critic_target(next_states, actions_next)
# Compute Q-targets for current states (y_i)
Q_targets = rewards + (gamma * Q_targets_next * (1 - dones))
# Compute critic loss
Q_expected = self.critic_local(states, actions)
critic_loss = F.mse_loss(Q_expected, Q_targets)
# Minimize the loss
self.critic_optim.zero_grad()
critic_loss.backward()
# prevent gradient explosion by gradient clipping
clip_grad_norm_(self.critic_local.parameters(), 1)
self.critic_optim.step()
# Update Actor
# Compute actor loss
actions_pred = self.actor_local(states)
actor_loss = -self.critic_local(states, actions_pred).mean()
# Minimize the loss
self.actor_optim.zero_grad()
actor_loss.backward()
self.actor_optim.step()
# Update target network
self.soft_update(self.critic_local, self.critic_target, TAU)
self.soft_update(self.actor_local, self.actor_target, TAU)
# Update noise
self.noise_factor = max(self.noise_factor * NOISE_DECAY, NOISE_MIN)
self.noise.reset()
def soft_update(self, local_model, target_model, tau):
"""Soft update target model to local model by tau
theta_target = tau * theta_local + (1 - tau) * theta_target
Parameters:
local_model : PyTorch model (weights will be copied from)
target_model : PyTorch model (weights will be copied to)
tau (float): interpolation factor
"""
for target_param, local_param in zip(target_model.parameters(), local_model.parameters()):
target_param.data.copy_(tau*local_param.data + (1-tau)*target_param.data)
def model_eval(self):
self.actor_local.eval()
self.critic_local.eval()
def model_train(self):
self.actor_local.train()
self.critic_local.train()
class OUNoise:
"""Ornstein-Uhlenbeck process."""
def __init__(self, num_agents, size, seed, mu=0., theta=0.15, sigma=0.2):
"""Initialize parameters and noise process."""
self.size = (num_agents, size)
self.mu = mu * np.ones(self.size)
self.theta = theta
self.sigma = sigma
self.seed = random.seed(seed)
self.reset()
def reset(self):
"""Reset the internal state (= noise) to mean (mu)."""
self.state = copy.copy(self.mu)
def sample(self):
"""Update internal state and return it as a noise sample."""
x = self.state
dx = self.theta * (self.mu - x) + self.sigma * np.random.standard_normal(self.size)
self.state = x + dx
return self.state
class ReplayBuffer:
"""Fixed-size buffer to store experience tuples."""
def __init__(self, action_size, buffer_size, batch_size, seed):
"""Initialize a ReplayBuffer object.
Params
======
buffer_size (int): maximum size of buffer
batch_size (int): size of each training batch
"""
self.action_size = action_size
self.memory = deque(maxlen=buffer_size) # internal memory (deque)
self.batch_size = batch_size
self.experience = namedtuple("Experience", field_names=["state", "action", "reward", "next_state", "done"])
self.seed = random.seed(seed)
def add(self, state, action, reward, next_state, done):
"""Add a new experience to memory."""
e = self.experience(state, action, reward, next_state, done)
self.memory.append(e)
def sample(self):
"""Randomly sample a batch of experiences from memory."""
experiences = random.sample(self.memory, k=self.batch_size)
states = torch.from_numpy(np.vstack([e.state for e in experiences if e is not None])).float().to(device)
actions = torch.from_numpy(np.vstack([e.action for e in experiences if e is not None])).float().to(device)
rewards = torch.from_numpy(np.vstack([e.reward for e in experiences if e is not None])).float().to(device)
next_states = torch.from_numpy(np.vstack([e.next_state for e in experiences if e is not None])).float().to(device)
dones = torch.from_numpy(np.vstack([e.done for e in experiences if e is not None]).astype(np.uint8)).float().to(device)
return (states, actions, rewards, next_states, dones)
def __len__(self):
"""Return the current size of internal memory."""
return len(self.memory)
# -
# We train our networks using this trainer
def train(agents):
""" Training an DDPG model
Parameters:
agents (Agents): Collection of agents to train DDPG
"""
# setup record of mean scores per episode keeping complete record
scores_total = []
scores_ma = deque(maxlen=100)
# setup time_stamp to periodically update target models
time_stamp = 0
# setup model to train
agents.model_train()
# loop over all episodes or until having learned sufficiently
for i in range(1, EPS_MAX + 1):
# get enviroment info
env_info = env.reset(train_mode=True)[brain_name]
states = env_info.vector_observations
# reset internal learning behavior episodically
agents.reset()
# keep record of the score of each agent over all timesteps
scores = np.zeros(num_agents)
# have the agents act and train in the enviroment until max timesteps per episode of any agent finishes
for t in range(1, T_MAX + 1):
# get actions from agents for current state
actions = agents.act(states)
# get current enviroment based on current actions
env_info = env.step(actions)[brain_name]
# get rewards, next_states, and dones of all agents
rewards = env_info.rewards
next_states = env_info.vector_observations
dones = env_info.local_done
# send all relevant info to agents to train model
for s, a, r, ns, d in zip(states, actions, rewards, next_states, dones):
agents.step(time_stamp, s, a, r, ns, d)
# update current states and scores
states = next_states
scores += rewards
# update time_stamp
time_stamp += 1
# if any agent is done, exit current episode
if np.any(dones):
break
# update history of episodic scores
scores_mean = np.mean(scores)
scores_total.append(scores_mean)
scores_ma.append(scores_mean)
print('\rEpisode {}\tAverage Score: {:.2f}\tAverage 100 moving average Score: {:.2f}'.format(i, scores_mean, np.mean(scores_ma)), end="")
if i % 10 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}\tAverage 100 moving average Score: {:.2f}'.format(i, scores_mean, np.mean(scores_ma)))
if np.mean(np.mean(scores_ma)) >= TARGET_MEAN_SCORE:
print('\nEnvironment solved in {:d} episodes!\tAverage 100 moving average Score: {:.2f}'.format(i, np.mean(scores_ma)))
torch.save(agents.actor_local.state_dict(), 'checkpoint_actor.pth')
torch.save(agents.critic_local.state_dict(), 'checkpoint_critic.pth')
break
return scores_total
# Creating the agents to train and training them
agents = Agent(num_agents, state_size, action_size, 0)
scores = train(agents)
# Displaying a graph of the training process
# +
import matplotlib.pyplot as plt
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# -
# Testing the trained agent
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
agents.model_eval() # set model to evaluation
states = env_info.vector_observations # get the current state (for each agent)
scores = np.zeros(num_agents) # initialize the score (for each agent)
while True:
actions = agents.act(states, False) # select an action (for each agent)
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
scores += env_info.rewards # update the score (for each agent)
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
print('Total score (averaged over agents) this episode: {}'.format(np.mean(scores)))
env.close()
# ### 4. It's Your Turn!
#
# Now it's your turn to train your own agent to solve the environment! A few **important notes**:
# - When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
# ```python
# env_info = env.reset(train_mode=True)[brain_name]
# ```
# - To structure your work, you're welcome to work directly in this Jupyter notebook, or you might like to start over with a new file! You can see the list of files in the workspace by clicking on **_Jupyter_** in the top left corner of the notebook.
# - In this coding environment, you will not be able to watch the agents while they are training. However, **_after training the agents_**, you can download the saved model weights to watch the agents on your own machine!
|
Deep Reinforcement Learning/Project 2 - Continuous Control/Continuous_Control.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
import pandas as pd
import seaborn as sns
import scipy.stats as sp
# ### Pregunta 1
Ramy_mc=[73.5,123.5,173.5,223.5,273.5]
Ramy_ni=[5,20,50,20,5]
FA_144_mc=Ramy_mc
FA_144_ni=[10,17,31,50,12]
Ramy=[]
FA_144=[]
for i in range(0,len(Ramy_mc)):
for j in range(0,Ramy_ni[i]):
Ramy.append(Ramy_mc[i])
for i in range(0,len(Ramy_mc)):
for j in range(0,FA_144_ni[i]):
FA_144.append(FA_144_mc[i])
intervalos=[]
for i in range(0,len(Ramy_mc)):
if Ramy_mc[i]-25 not in intervalos:
intervalos.append(Ramy_mc[i]-25)
if Ramy_mc[i]+25 not in intervalos:
intervalos.append(Ramy_mc[i]+25)
# +
fig, ax = plt.subplots(1,2,figsize=(12,4))
ax[0].hist(Ramy,bins=intervalos,color='y')
ax[0].legend('r',fontsize='x-large')
ax[1].hist(FA_144,bins=intervalos,color='m',label='FA 144')
ax[1].legend('f',fontsize='x-large')
plt.show()
# -
print('El promedio de peso del tomate Ramy es de', np.mean(Ramy), '[grm]')
print('El promedio de peso del tomate FA 144 es de', np.mean(FA_144), '[grm]')
print('La varianza de peso del tomate Ramy es de', np.std(Ramy)**2, '[grm]^2')
print('La varianza de peso del tomate FA 144 es de', np.std(FA_144)**2, '[grm]^2')
print('El CV del tomate Ramy es',np.std(Ramy)/np.mean(Ramy))
print('El CV del tomate Ramy es',np.std(FA_144)/np.mean(FA_144))
# ### Pregunta 2
A=[9.8 ,10.2 ,10.1 ,9.7 ,8.8 ,10.7 ,11.1]
B=[10.1 ,10.1, 9.6 ,9.9 ,10.9 ,9.7]
C=[9.7, 9.5, 10.3, 8.9, 10.6, 10.4, 9.8, 11.0, 9.2 ]
intervalo=np.arange(9.2,11.3,.3)
intervalo
# +
fig, ax = plt.subplots(1,3,figsize=(18,4))
ax[0].hist(A,bins=intervalo,color='y')
ax[0].legend('A',fontsize='x-large')
ax[1].hist(B,bins=intervalo,color='m')
ax[1].legend('B',fontsize='x-large')
ax[2].hist(C,bins=intervalo,color='b')
ax[2].legend('C',fontsize='x-large')
plt.show()
# -
print(sp.describe(A))
print(sp.describe(B))
print(sp.describe(C))
print(np.std(A),np.mean(A))
print(np.std(B),np.mean(B))
print(np.std(C),np.mean(C))
print(np.std(A)/np.mean(A))
print(np.std(B)/np.mean(B))
print(np.std(C)/np.mean(C))
# ### Pregunta 3
S=[578,755,840,690,1015,1210,1350,670,1610,1550]
I=[450,610,790,750,1210,1150,1450,705,1350,1450]
print(np.std(S)**2,np.mean(S))
print(np.std(I)**2,np.mean(I))
np.corrcoef(S, I)
plt.scatter(S,I)
def reg(x,y):
modelo=[]
residuos=[]
for i in range(0,len(x)):
modelo.append(x[i]*0.96+48.82)
for i in range(0,len(x)):
residuos.append(y[i]-modelo[i])
return residuos
# +
#covarianza para un array de nxm
tabla_bivariada=[[1,2,2,3],[1,3,4,5],[2,3,6,9],[5,6,8,8],[5,8,9,6]]
# -
# ### Pregunta 4
Y=[20,25,30,35]
X=[10.5,12.5,14.5,16.5,18.5]
n=0
suma=0
promX=0
promY=0
for i in range(0,len(X)):
for j in range(0,len(Y)):
n+=tabla_bivariada[i][j]
promX+=tabla_bivariada[i][j]*X[i]
promY+=tabla_bivariada[i][j]*Y[j]
suma+=tabla_bivariada[i][j]*X[i]*Y[j]
print(promX,promY)
promX=promX/n
promY=promY/n
cov=suma/n -promX*promY
print(cov)
s2_x=0
for i in range(0,len(X)):
s2_x+= (X[i]**2)*sum(tabla_bivariada[i])
s2_x=s2_x/n-promX**2
print(s2_x)
beta_1=cov/s2_x
beta_0=promY-beta_1*promX
print(beta_1,beta_0)
|
MAT041/ayudantia_2/python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Quest for the Ultimate Optimizer - Episode 1
# ----------------
# The field of neural networks, both in terms of research and practical applications, has grown dramatically in recent years. This has been attributed, among other things, to the concurrent growth of computing power and training data. These have lead, in turn, to the development of more and more complex NN (Neural Network) architecture, with more and more layers, various type of cells, various ways to train them, etc.
# Yet, one central piece of the puzzle has remained relatively simple : the optimization algorithm. Don't get me wrong, there are a lot of different variations, they actually constitute a field of research all by themselves, have spurred thousands of research articles, most of them with detailed mathematical proof of convergence, extensive testing, etc. Yet most of these algorithms (at least those widely used for NN training) can be described by a couple of lines of codes ! 5 lines for the most complicated (and I'm being generous).
#
# I remember, when I first tried to learn about neural networks, around a year ago, this relative simplicity of the SGD (Stochastic Gradients Descent) algorithms used for NN optimization struck me as one of the most intriguing aspect of the field. In one of the [lecture of his online course](https://youtu.be/SJ48OZ_qlrc?t=579), <NAME> tries to give some explanation as to why we haven't yet found the perfect recipe to train NN, and more or less conclude that it's the diversity of the NNs both in their architectures and in their tasks that makes the NN optimization problem such a tough problem to crack, especially if you are looking for a "silver bullet", "one size fit all" optimization algorithm.
#
# One of the conclusion of Hinton's lecture on how to train NN is to look at whatever <NAME>'s, and his "No More Pesky Learning Rates" group's latest recipe is. The name of the group and of the [algorithm](https://arxiv.org/pdf/1206.1106.pdf) they came up with highlights one of the biggest frustrations you are faced with when training NN : each of these optimizers have at least one knob (when it's not 3 or 4) that needs tuning for your neural net (or any other system you're trying to optimize) to converge both in a reasonable time and to its lowest possible error value.
# Shouldn't we be able to design a system that is good at interpreting a series of data like successive gradients and predicting what is the best next update to use based on those past data (i.e. an SGD-like algorithm) ? Wait ... that sounds very much like what a recurrent neural network is good at, doesn't it ?
#
# Like any good idea, if you look hard enough on the internet, you'll find someone, way ahead of you, who has already investigated and perfected the concept. In our case, a team from DeepMind proposed, in 2016, an implementation of this idea in the paper [“Learning to learn by gradient descent by gradient descent”](https://arxiv.org/abs/1606.04474). This paper was pointed out to me by someone who works at ... you guessed it ... DeepMind.
# @Cyprien : thanks for that !
# It describes how you can train a (relatively) simple RNN (recurrent neural network) to act as the optimizer of another problem, be it very simple like minimizing the quadratic function, or more complex, like training a neural net on the MNIST or CIFAR-10 datasets.
#
# The target of the notebook below and hopefully of the series of notebooks/articles that will follow, is to reproduce some of the results from DeepMind’s paper and explore some of the doors it opens.
# ## RNN as the optimizer - First experiments
# I first tried to experiment with DeepMind's code ([which they have been nice enough to share on GitHub](https://github.com/deepmind/learning-to-learn)), but it turns out their code includes a lot of bells and whistles, that makes it easy to use, but hard to decipher its inner workings. Si I have opted to re-use Llion Jones's very simple, yet elegant way of implementing “Learning to learn" ideas as explained in his article ["Learning to Learn by Gradient Descent by Gradient Descent - As simple as possible in TensorFlow"](https://hackernoon.com/learning-to-learn-by-gradient-descent-by-gradient-descent-4da2273d64f2) which I encourage you to read if you want to understand how this is set-up
# Let's start by importing tensorflow and a couple of usefull tools
import tensorflow as tf
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import os
# If you are using the GPU version of tensorflow you can choose which PU to use. Here I'm using the CPU because I'll start with small problems where GPU don't really speed things up.
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"]=""
# ### Summary of Llion Jone's Notebook
#
# Just to be clear, the code in this section is, more or less, a copy paste from [Llion Jones's article](https://hackernoon.com/learning-to-learn-by-gradient-descent-by-gradient-descent-4da2273d64f2).
# +
DIMS = 2 # Dimensions of the quadratic function, the simplest application problem in DeepMind's paper
scale = tf.random_uniform([DIMS], 0.5, 1.5)
# The scale vector gives a different shape to the quadratic function at each initialization
def quadratic(x):
x = scale*x
return tf.reduce_sum(tf.square(x))
# +
# Some reference optimizers for benchmarking
def g_sgd(gradients, state, learning_rate=0.1):
# Vanilla Stochastic Gradient Descent
return -learning_rate*gradients, state
def g_rms(gradients, state, learning_rate=0.1, decay_rate=0.99):
# RMSProp
if state is None:
state = tf.zeros(DIMS)
state = decay_rate*state + (1-decay_rate)*tf.pow(gradients, 2)
update = -learning_rate*gradients / (tf.sqrt(state)+1e-5)
return update, state
# +
TRAINING_STEPS = 20 # This is 100 in the paper
initial_pos = tf.random_uniform([DIMS], -1., 1.)
def learn(optimizer):
losses = []
x = initial_pos
state = None
# The loop below unrolls the 20 steps of the optimizer into a single tensorflow graph
for _ in range(TRAINING_STEPS):
loss = quadratic(x)
losses.append(loss)
grads, = tf.gradients(loss, x)
update, state = optimizer(grads, state)
x += update
return losses
# -
sgd_losses = learn(g_sgd)
rms_losses = learn(g_rms)
# +
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
x = np.arange(TRAINING_STEPS)
for _ in range(3):
sgd_l, rms_l = sess.run([sgd_losses, rms_losses])
p1, = plt.semilogy(x, sgd_l, label='SGD')
p2, = plt.semilogy(x, rms_l, label='RMS')
plt.legend(handles=[p1, p2])
plt.title('Losses')
plt.show()
# +
# Now let's define the RNN optimizer
LAYERS = 2
STATE_SIZE = 20
cell = tf.contrib.rnn.MultiRNNCell(
[tf.contrib.rnn.LSTMCell(STATE_SIZE) for _ in range(LAYERS)])
cell = tf.contrib.rnn.InputProjectionWrapper(cell, STATE_SIZE)
cell = tf.contrib.rnn.OutputProjectionWrapper(cell, 1)
cell = tf.make_template('cell', cell)
def g_rnn(gradients, state):
# Make a `batch' of single gradients to create a
# "coordinate-wise" RNN as the paper describes.
gradients = tf.expand_dims(gradients, axis=1)
if state is None:
state = [[tf.zeros([DIMS, STATE_SIZE])] * 2] * LAYERS
update, state = cell(gradients, state)
# Squeeze to make it a single batch again.
return tf.squeeze(update, axis=[1]), state
# +
def optimize(loss, learning_rate=0.1):
# "Meta optimizer" to be applied on the RNN defined above
optimizer = tf.train.AdamOptimizer(learning_rate)
gradients, v = zip(*optimizer.compute_gradients(loss))
gradients, _ = tf.clip_by_global_norm(gradients, 1.)
return optimizer.apply_gradients(zip(gradients, v))
# The "meta" optimization is first applied to minimize the sum of the 20 RNN iterations results
rnn_losses = learn(g_rnn)
sum_losses = tf.reduce_sum(rnn_losses)
apply_update = optimize(sum_losses, learning_rate=0.0003)
# +
sess.run(tf.global_variables_initializer())
ave = 0
for i in range(4000):
err, _ = sess.run([sum_losses, apply_update])
ave += err
if i % 1000 == 0:
print(ave / 1000 if i!=0 else ave)
ave = 0
print(ave / 1000)
# -
# Graph to compare RNN to the 2 baseline optimizers defined above
x = np.arange(TRAINING_STEPS)
for _ in range(3):
sgd_l, rms_l, rnn_l = sess.run(
[sgd_losses, rms_losses, rnn_losses])
p1, = plt.semilogy(x, sgd_l, label='SGD')
p2, = plt.semilogy(x, rms_l, label='RMS')
p3, = plt.semilogy(x, rnn_l, label='RNN')
plt.legend(handles=[p1, p2, p3])
plt.title('Losses')
plt.show()
# In summary, with this first try, we get an RNN that beats SGD but doesn't match RMSProp after 5 iterations.
# <NAME> mentions, in his article, a problem associated with the approach used above, which might explain why the loss value stops decreasing at around 1e-5 or 1e-6 : the gradients get so small that the RNN isn’t able to compute sensible updates. That's actually a key feature of neural networks : to work well, they need their inputs to remain within a specific scale. This is not the case here since we are feeding into the RNN gradients that start at around 1 but could go down to a much lower value if the optimizer works well and actually find the zero machine.
# DeepMind's paper proposes a solution to this : use the log gradient and the sign instead of the gradient. See the paper for details. We will try to implement that later in this article. We will also see that there may be other ways to deal with this problem in episode 2 of this series.
# ### Some helper functions and formatting
# Before we try to improve our RNN performance, let's define a few functions to streamline the code
def print_dict(*args):
"""Prints variables in a dict format for easier reading"""
dict_name = dict((name,eval(name)) for name in args)
print(dict_name)
# Below the same optim as before but run for 20,000 iterations and with some additional variables, printed with the function above, to track the convergence progress :
# +
sess.run(tf.global_variables_initializer())
list_result = np.array([])
for it in range(20001):
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 2000 == 0 :
sum_error = '{:.2f}'.format(sum(errors)) # Parameter being optimized
optim_result = '{:.2E}'.format(errors[-1]) # Result of the RNN after 20 iterations
# And below the rolling average of the log10 result of RNN
# In other words, you get -6 if the RNN result oscillates between 1e-5 and 1e-7
average_log_result = '{:.2f}'.format(np.log10(list_result[-2000:]).mean())
print_dict('it', 'sum_error', 'optim_result', 'average_log_result')
# Let's store the convergence for later comparison
# Naming convention will be : name of the optimizer followed by the parameter used to optimize it (when applicable)
# Here "RNN_simple" is the name and "sum_res" is the parameter used in the meta optimization
RNN_simple_sum_res = list_result
# -
# Let's add a couple of standard graphs and test them
def graph_optimizers(f1, f2, f3, n=3, training_steps=TRAINING_STEPS):
# Draws n graphs to compare RNN to the 2 baseline optimizers
x = np.arange(training_steps)
for _ in range(n):
sgd_l, rms_l, rnn_l = sess.run([f1, f2, f3])
p1, = plt.semilogy(x, sgd_l, label='SGD')
p2, = plt.semilogy(x, rms_l, label='RMS')
p3, = plt.semilogy(x, rnn_l, label='RNN')
plt.legend(handles=[p1, p2, p3])
plt.title('Losses')
plt.show()
graph_optimizers(sgd_losses, rms_losses, rnn_losses)
# As we are dealing with 2 levels of iterations, we need a naming convention to distinguish them. So let's call the "base" iterations the ones performed by the RNN being optimized or the benchmarks defined at the beginning, and "meta" iterations the ones performed by tensorflow optimizer (here we are using Adam, cf. optimize() function definition) to optimize the result of the "base" iterations of the RNN.
# The convergence of the "base" convergences are displayed above, but it would be nice to check the "meta" convergence of our RNN, meaning the progression of the RNN performance over the 20,000 iterations we performed.
def rolling_log_average(array, L):
"""Rolling average of the log of the array over a length of L"""
# Since the loss decrease in logarithm scale, we will apply this to track average results
rolling_av = np.array([])
for i in range(array.size):
rolling_av = np.append(rolling_av, 10**(np.log10(array[:i+1][-L:] + 1e-38).mean()))
return rolling_av
import warnings
def draw_convergence(*args):
"""Draws the convergence of one or several meta optimizations
transparent area is the raw results, the plain line is the 500 rolling 'log average'"""
it = 0
for f in args:
it = max(eval(f).size, it)
handles = []
for f in args:
flist = eval(f)[np.logical_not(np.isnan(eval(f)))] #removes NaN
flist_rolling = rolling_log_average(flist, 500)
flist_size = flist.size
#matplotlib doesn't like graphs of different length so we fill the shorter graphs with None
if flist_size < it:
flist = np.append(flist, [None]*(it-flist_size))
flist_rolling = np.append(flist_rolling, [None]*(it-flist_size))
c1, = plt.semilogy(range(it), flist, alpha=0.3)
c2, = plt.semilogy(range(it), flist_rolling, color=c1.get_color(), label=f)
handles = handles + [c2]
plt.legend(handles=handles)
plt.title('End result of the optimizer')
#matplotlib still doesn't like graphs of different length so we filter associated warnings
warnings.filterwarnings("ignore",category =RuntimeWarning)
plt.show()
# As explained in the function docstring, the transparent area is the raw results, which allows a visualization of the scatter of results, while the plain line shows the 500 rolling "log average" to monitor overall progress.
# I'll be using this function on the end result of the RNN. So here, it basically displays the last "base" iteration result of the RNN after each iteration of the "meta optimizer", which oscillates arround 1e-6 at the end of both above and below graphs.
draw_convergence('RNN_simple_sum_res')
# Now that we are all setup, we can go back to our RNN
# ### Let's beat RMSProp
#
# RMSProp is one of the most widely used optimizer for NN, it was actually introduced by <NAME> in the lecture I refer to in the introduction. Our target will be to try to beat it with our RNN optimizer.
# Let's run our 2 baseline (SGD and RMSProp) a 1000 times to find out how far off we are.
list_sgd_errors = np.array([])
list_rms_errors = np.array([])
for it in range(1000):
sgd_errors, rms_errors = sess.run([sgd_losses, rms_losses])
list_sgd_errors = np.append(list_sgd_errors, sgd_errors[-1])
list_rms_errors = np.append(list_rms_errors, rms_errors[-1])
draw_convergence('list_sgd_errors', 'list_rms_errors', 'RNN_simple_sum_res')
# Now that we have seen what the scatter looks like, we can replace the results by lists containing only the 'log' average :
SGD = np.full(20001, rolling_log_average(list_sgd_errors, 1000)[-1])
RMS = np.full(20001, rolling_log_average(list_rms_errors, 1000)[-1])
draw_convergence('SGD', 'RMS', 'RNN_simple_sum_res')
# Looks like we have our work cut out for us : we need to cross the orange line.
# Let's rename it for later graph
Target_RMS = RMS
# Before we start meddling with the RNN inputs and outputs, there is a key parameter that we should try to fine tune :
# #### The loss function
# We are currently using as our loss function the sum of the errors across the 20 iterations. That's a good way to ensure that we both converge to 0 and get there as fast as possible, since the smaller the intermediate steps errors are, the quicker the optimizer is. However, given the problem we are trying to solve, the error at the end will be dwarfed by those from the first iterations, and their corresponding gradients will have the same behaviour.
# So, before we try to make an optimizer that gets to 0 quickly, we should try to find one that actually finds the 0 (if that's possible in 20 steps ...), or as said in the title, one that can beat RMSProp. To do that, let's focus only on the last error.
# +
rnn_losses = learn(g_rnn)
end_losses = rnn_losses[-1]
# Target is now to minimize the result at the end of the 20 iterations
apply_update = optimize(end_losses, learning_rate=0.0003)
sess.run(tf.global_variables_initializer())
list_result = np.array([])
for it in range(20001):
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 2000 == 0 :
optim_result = '{:.2E}'.format(errors[-1])
average_log_result = '{:.2f}'.format(np.log10(list_result[-2000:]).mean())
print_dict('it', 'optim_result', 'average_log_result')
# Let's store the convergence for later comparison
RNN_simple_end_res = list_result
# -
draw_convergence('Target_RMS', 'RNN_simple_sum_res', 'RNN_simple_end_res')
# Regarding the effect of minimizing the end result rather than the sum of errors along the 20 "base" iterations, we can make 2 observations :
# - We get about 1 order of magnitude improvement on the end result of the RNN after 20,000 "meta" iterations, which is not huge considering it is 1 out of 6 orders of magnitude overall
# - The "meta" optimization is slower initially but does not look to be slowing down after the 20,000 it, so let's give it another 30,000 it
# +
it = 20000
for _ in range(30000):
it += 1
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 5000 == 0 :
optim_result = '{:.2E}'.format(errors[-1])
average_log_result = '{:.2f}'.format(np.log10(list_result[-5000:]).mean())
print_dict('it', 'optim_result', 'average_log_result')
# Let's store the convergence for later comparison
RNN_simple_end_res = list_result
# -
Target_RMS = np.full(50001, Target_RMS[0])
draw_convergence('Target_RMS', 'RNN_simple_sum_res', 'RNN_simple_end_res')
# Conclusions : the optimization seems to settle around 1e-8, which is a bit more significant in terms of improvement.
# The shape of "base" convergence of the 2 RNNs are also different with the first (displayed earlier) outperforming RMSProp in the first few iterations, then flattening out, while the second (displayed below) is more consistent across the 20 iterations. This illustrates the fact that using the sum of errors as the loss function focuses the optimization on the first few steps.
graph_optimizers(sgd_losses, rms_losses, rnn_losses)
# Before switching our focus to the RNN and how it processes the gradients, there is something else we could try with the loss function:
# #### The loss function - using log scale
# One problem with our current loss function is that, in order to beat RMSProp, we have to go from loss of around 1 to around 1E-18, and that's a problem when the inputs scale doesn't follow the same scale change. Even for optimizer equiped to deal with such vanishing gradients (like adam or RMSProp), the small scale can generate some accuracy problems.
# Thanksfully, using the log of the error as our loss function adresses this problem. Let's see if it also brings some improvement to the RNN results
# +
rnn_losses = learn(g_rnn)
log_loss = tf.log(rnn_losses[-1]) # log added to the optimization target
apply_update = optimize(log_loss, learning_rate=0.0003)
sess.run(tf.global_variables_initializer())
list_result = np.array([])
for it in range(50001):
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 5000 == 0 :
optim_result = '{:.2E}'.format(errors[-1])
average_log_result = '{:.2f}'.format(np.log10(list_result[-5000:]).mean())
print_dict('it', 'optim_result', 'average_log_result')
# Let's store the convergence for later comparison
RNN_simple_end_log_res = list_result
# -
RMS = np.full(50000, rolling_log_average(list_rms_errors, 1000)[-1])
draw_convergence('Target_RMS', 'RNN_simple_sum_res', 'RNN_simple_end_res', 'RNN_simple_end_log_res')
# The answer is : apparently not ... there is a slightly better convergence rate but the end result is the same.
# This was either a terrible idea (I have a lot of those :) or there is something else preventing the RNN from improving further. So let's put aside this idea for now, and see if we can improve how the RNN is set up.
#
# #### The RNN inputs - log scale
# Let's go back to the problem mentioned at the end of the first section: the changing scale of the inputs. As explained before, having inputs to RNN going from 1 to 1E-15 is not ideal (there are probably a few reasons for that, but one of them is that the bias and the activation functions in the RNN would need to follow the same scale change for the RNN to keep the same behaviour).
# One way to deal with that, as mentioned earlier, is to use the log of the gradient instead of directly the gradient. This has the advantage of making the output less sensitive to scale changes. For example having gradients go from 100 to 1 will be interpreted similarly by the RNN as a change from 1 to 0.01. Problem is that, with the last value we get into negative values and we need to keep a way to distinguish between positive and negative gradients. To deal with that, we add another operation that rescale, this time linearly, the log(|gardient|) from [min_log_gradient - 0] to [0 - 1] and then applies the sign of the gradient to the result. The result is a casting operation from [exp(min_log_gradient), 1] to [0, 1] and from [-1, -exp(min_log_gradient)] to [-1, 0].
# As you may have noticed I have introduced the min_log_gradient parameter because, since we are doing computer science, we need to stop somwhere. I've started with an arbitrary value of -15, so that gradients between exp(-15) and 1 are casted to values between 0 and 1, but we will investigate that further later on.
#
# A few additional remarks :
# I had a look at the preprocessing proposed in Deepmind's paper, and I noticed 2 differences. They have a second formula to deal with gradients smaller than exp(min_log_gradient). I didn't bother with that because I'm not sure the RNN I'm using would be able to deal with two different preprocessing regime anyway (at least not without increasing the depth of the network to make it "smarter").
# They also seem to rescale the log(gradient) differently : I'm probably missing something here, but it looks like 10 and -0.1 would result in the same input for the RNN with their approach. So, if someone figured it out, I'm interested.
# That said, we already know their approach gives a huge performance boost compared to the most common optimizers on a variety of problems, so I'm pretty sure there are good reasons to do things the way they did.
#
# OK, so let's go back to the preprocessing approach I started with
# +
def g_rnn_log15(gradients, state):
gradients = tf.expand_dims(gradients, axis=1)
# Casting of gradients from [exp(-15), 1] to [0, 1] and [-1, -exp(-15)] to [-1, 0]
min_log_gradient = -15
log_gradients = tf.log(tf.abs(gradients) + np.exp(min_log_gradient-5)) # residual added to avoid log(0)
sign_gradients = tf.sign(gradients)
trans_gradients = tf.multiply(sign_gradients,((log_gradients - min_log_gradient) / (- min_log_gradient)))
if state is None:
state = [[tf.zeros([DIMS, STATE_SIZE])] * 2] * LAYERS
update, state = cell(trans_gradients, state)
# Casting of output from [0, 1] to [exp(-15), 1] and [-1, 0] to [-1, -exp(-15)]
abs_update = tf.abs(update)
sign_update = tf.sign(update)
update = tf.multiply(sign_update, tf.exp(abs_update * (- min_log_gradient) + min_log_gradient))
return tf.squeeze(update, axis=[1]), state
rnn_losses = learn(g_rnn_log15)
end_losses = rnn_losses[-1]
apply_update = optimize(end_losses, learning_rate=0.0003)
sess.run(tf.global_variables_initializer())
list_result = np.array([])
for it in range(50001):
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 5000 == 0 :
optim_result = '{:.2E}'.format(errors[-1])
average_log_result = '{:.2f}'.format(np.log10(list_result[-5000:]).mean())
print_dict('it', 'optim_result', 'average_log_result')
RNN_log15_end_res = list_result
# -
draw_convergence('Target_RMS', 'RNN_simple_end_res', 'RNN_log15_end_res')
# Now we do get briefly below the results of the initial RNN set-up, but the convergence is not stable.
# Maybe it's time to revisit this loss function, and see if those vanishing gradients are the problem here. To do that, let's use the loss function defined in the section "The loss function - using log scale"
# +
rnn_losses = learn(g_rnn_log15)
log_loss = tf.log(rnn_losses[-1]) # New target function
apply_update = optimize(log_loss, learning_rate=0.0003)
sess.run(tf.global_variables_initializer())
list_result = np.array([])
for it in range(50001):
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 5000 == 0 :
optim_result = '{:.2E}'.format(errors[-1])
average_log_result = '{:.2f}'.format(np.log10(list_result[-5000:]).mean())
print_dict('it', 'optim_result', 'average_log_result')
# Let's store the convergence for later comparison
RNN_log15_end_log_res = list_result
# -
draw_convergence('Target_RMS', 'RNN_simple_end_log_res', 'RNN_log15_end_res', 'RNN_log15_end_log_res')
# Now we are getting somwhere! Let's look how the "base" convergence looks like :
graph_optimizers(sgd_losses, rms_losses, rnn_losses)
# So, still not as good as RMSProp, but we are progressing.
# I said earlier that I set the min_log_gradient arbitrarily at -15 . In order to increase the range of gradients across which our rescaling of gradients works, we can lower this value to -30 :
# +
def g_rnn_log30(gradients, state):
gradients = tf.expand_dims(gradients, axis=1)
# Casting of gradients from [exp(-30), 1] to [0, 1] and [-1, -exp(-30)] to [-1, 0]
min_log_gradient = -30
log_gradients = tf.log(tf.abs(gradients) + np.exp(min_log_gradient-5))
sign_gradients = tf.sign(gradients)
trans_gradients = tf.multiply(sign_gradients,((log_gradients - min_log_gradient) / (- min_log_gradient)))
if state is None:
state = [[tf.zeros([DIMS, STATE_SIZE])] * 2] * LAYERS
update, state = cell(trans_gradients, state)
# Casting of output from [0, 1] to [exp(-30), 1] and [-1, 0] to [-1, -exp(-30)]
abs_update = tf.abs(update)
sign_update = tf.sign(update)
update = tf.multiply(sign_update, tf.exp(abs_update * (- min_log_gradient) + min_log_gradient))
return tf.squeeze(update, axis=[1]), state
rnn_losses = learn(g_rnn_log30)
log_loss = tf.log(rnn_losses[-1])
apply_update = optimize(log_loss, learning_rate=0.0003)
sess.run(tf.global_variables_initializer())
list_result = np.array([])
for it in range(50001):
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 5000 == 0 :
optim_result = '{:.2E}'.format(errors[-1])
average_log_result = '{:.2f}'.format(np.log10(list_result[-5000:]).mean())
print_dict('it', 'optim_result', 'average_log_result')
# Let's store the convergence for later comparison
RNN_log30_end_log_res = list_result
# -
draw_convergence('Target_RMS', 'RNN_simple_end_log_res', 'RNN_log15_end_log_res', 'RNN_log30_end_log_res')
# Slower to start, but it looks like we are almost there ...
# Let's add another 50,000 iterations :
# +
it = 50000
for _ in range(50000):
it += 1
errors, _ = sess.run([rnn_losses, apply_update])
list_result = np.append(list_result, errors[-1])
if it % 5000 == 0 :
optim_result = '{:.2E}'.format(errors[-1])
average_log_result = '{:.2f}'.format(np.log10(list_result[-5000:]).mean())
print_dict('it', 'optim_result', 'average_log_result')
# Let's store the convergence for later comparison
RNN_log30_end_log_res = list_result
# -
Target_RMS = np.full(100001, Target_RMS[0])
draw_convergence('Target_RMS', 'RNN_simple_end_log_res', 'RNN_log15_end_log_res', 'RNN_log30_end_log_res')
graph_optimizers(sgd_losses, rms_losses, rnn_losses)
# Looks like we can finally declare victory over RMSProp!
#
# Just to recap, to beat RMSProp :
# - we focussed the loss function on the last iteration of the "base" optimization
# - we applied a logarithmic scale on this loss to avoid the problem of vanishing gradient on the "meta" optimization
# - we also applied a logarithmic rescaling on the input of the RNN "base" optimizer to basically avoid the same problem
# ### What's next
# We seem to have gotten to an RNN set-up which can outperform RMSProp, so target reached. But we still have quite a few open questions :
# I mentioned that I chose the `min_log_gradient` parameter kind of arbitrarily. This is obviously not satisfying from ... well, any point of views. I tried to go further than -30 but the convergence becomes either too slow or doesn't even start, so we should try to understand this better and find a workaround. I also mentioned, at the end of the first section, that there might be other ways to preprocess the gradients than just log rescaling, so we can also explore that further ... in the next episode of The Quest for the Ultimate Optimizer.
#
# As a last comment, I should also mention that my declaration of "victory over RMSProp" is probably unfair, as I didn't try to optimize RMSProp's parameters like I did for the RNN. So, what kind of lame excuse could I use to avoid another ten pages of painfull tensorflow tinkering ? You're sick of reading my prose ? Life's unfair ? No wait, I know, let's explore that in the third episode of our Quest (if I ever find the time to work on it).
|
Episode1/Episode1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from surprise import SVDpp
from surprise import Dataset
from surprise import Reader
from surprise import accuracy
from surprise.model_selection import KFold
import pandas as pd
# -
# Cargar los registros de usuarios
ratings_df = pd.read_csv('ratings.dat', sep='::', header=None)
del (ratings_df[3])# borrar la tercera fila correspondiente a la fecha
reader = Reader(rating_scale=(1,5))# Clase usada 'parsear' el dataframe de ratings
data = Dataset.load_from_df(ratings_df,reader)# cargar el dataset desde dataframe y el reader
kf = KFold(n_splits=2) # iterador de validacion cruzada
algo = SVDpp()# uso del algoritmo
# ciclo de 2 iteraciones
for train_set, test_set in kf.split(data):
algo.fit(train_set)# ajuste del modelo con cada subdivision
predictions = algo.test(test_set) # predicciones con cada subdivision
accuracy.rmse(predictions, verbose=True)# medida de precision por cada iteracion
|
SVDpp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from IPython.core.debugger import Pdb; pdb = Pdb()
def get_down_centre_last_low(p_list):
zn_num = len(p_list) - 1
available_num = min(9, (zn_num - 6))
index = len(p_list) - 4
for i in range(0, available_num // 2):
if p_list[index - 2] < p_list[index]:
index = index -2
else:
return index
return index + 2
def get_down_centre_first_high(p_list):
s = max(enumerate(p_list[3:]), key=lambda x: x[1])[0]
return s + 3
def down_centre_expand_spliter(p_list):
lr0 = get_down_centre_last_low(p_list)
hl0 = get_down_centre_first_high(p_list[: lr0 - 2])
hr0 = lr0 -1
while hr0 < len(p_list) - 6:
if p_list[hr0] > p_list[hl0] and (len(p_list) - hr0) > 5:
hl0 = hr0
lr0 = (len(p_list) - 1 + hr0) // 2
if lr0 % 2 == 1:
lr0 = lr0 -1
# lr0 = hr0 + 3
break
hr0 = hr0 + 2
return [0, hl0, lr0, len(p_list) - 1], [p_list[0], p_list[hl0], p_list[lr0], p_list[-1]]
# y = [0, 100, 60, 130, 70, 120, 40, 90, 50, 140, 85, 105]
# y = [0, 100, 60, 110, 70, 72, 61, 143, 77, 91, 82, 100, 83, 124, 89, 99]
# y = [0, 100, 60, 110, 70, 115, 75, 120, 80, 125, 85, 130, 90, 135]
# y = [0, 100, 60, 110, 70, 78, 77, 121, 60, 93, 82, 141, 78, 134]
# x = list(range(0, len(y)))
# gg = [min(y[1], y[3])] * len(y)
# dd = [max(y[2], y[4])] * len(y)
# plt.figure(figsize=(len(y),4))
# plt.grid()
# plt.plot(x, y)
# plt.plot(x, gg, '--')
# plt.plot(x, dd, '--')
# sx, sy = down_centre_expand_spliter(y)
# plt.plot(sx, sy)
# plt.show()
# +
# Centre Expand Prototype
# %matplotlib inline
import matplotlib.pyplot as plt
y_base = [0, 100, 60, 130, 70, 120, 40, 90, 50, 140, 85, 105, 55, 80]
for i in range(10, len(y_base)):
y = y_base[:(i + 1)]
x = list(range(0, len(y)))
gg = [min(y[1], y[3])] * len(y)
dd = [max(y[2], y[4])] * len(y)
plt.figure(figsize=(i,4))
plt.grid()
plt.plot(x, y)
plt.plot(x, gg, '--')
plt.plot(x, dd, '--')
if i % 2 == 1:
sx, sy = down_centre_expand_spliter(y)
plt.plot(sx, sy)
plt.show()
# +
# Random Centre Generator
# %matplotlib inline
import random
import matplotlib.pyplot as plt
y_max = 150
y_min = 50
num_max = 14
def generate_next(y_list, direction):
if direction == 1:
y_list.append(random.randint(max(y_list[2], y_list[4], y_list[-1]) + 1, y_max))
elif direction == -1:
y_list.append(random.randint(y_min, min(y_list[1], y_list[3], y_list[-1]) - 1))
y_base = [0, 100, 60, 110, 70]
# y_base = [0, 110, 70, 100, 60]
# y_base = [0, 100, 60, 90, 70]
# y_base = [0, 90, 70, 100, 60]
direction = 1
for i in range(5, num_max):
generate_next(y_base, direction)
direction = 0 - direction
print(y_base)
for i in range(11, len(y_base), 2):
y = y_base[:(i + 1)]
x = list(range(0, len(y)))
gg = [min(y[1], y[3])] * len(y)
dd = [max(y[2], y[4])] * len(y)
plt.figure(figsize=(i,4))
plt.title(y)
plt.grid()
plt.plot(x, y)
plt.plot(x, gg, '--')
plt.plot(x, dd, '--')
sx, sy = down_centre_expand_spliter(y)
plt.plot(sx, sy)
plt.show()
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# Group 1
# y_base = [0, 100, 60, 110, 70, 99, 66, 121, 91, 141, 57, 111, 69, 111]
# y_base = [0, 100, 60, 110, 70, 105, 58, 102, 74, 137, 87, 142, 55, 128]
y_base = [0, 100, 60, 110, 70, 115, 75, 120, 80, 125, 85, 130, 90, 135]
# y_base = [0, 100, 60, 110, 70, 120, 80, 130, 90, 140, 50, 75]
# y_base = [0, 100, 60, 110, 70, 114, 52, 75, 54, 77, 65, 100, 66, 87, 70, 116]
# y_base = [0, 100, 60, 110, 70, 72, 61, 143, 77, 91, 82, 100, 83, 124, 89, 99, 89, 105]
# Group 2
# y_base = [0, 110, 70, 100, 60, 142, 51, 93, 78, 109, 60, 116, 50, 106]
# y_base = [0, 110, 70, 100, 60, 88, 70, 128, 82, 125, 72, 80, 63, 119]
# y_base = [0, 110, 70, 100, 60, 74, 66, 86, 57, 143, 50, 95, 70, 91]
# y_base = [0, 110, 70, 100, 60, 77, 73, 122, 96, 116, 82, 124, 69, 129]
# y_base = [0, 110, 70, 100, 60, 147, 53, 120, 77, 103, 56, 76, 74, 92]
# y_base = [0, 110, 70, 100, 60, 95, 55, 90, 50, 85, 45, 80, 40, 75]
# Group 3
# y_base = [0, 100, 60, 90, 70, 107, 55, 123, 79, 112, 64, 85, 74, 110]
# y_base = [0, 100, 60, 90, 70, 77, 55, 107, 76, 141, 87, 91, 60, 83]
# y_base = [0, 100, 60, 90, 70, 114, 67, 93, 58, 134, 53, 138, 64, 107]
# y_base = [0, 100, 60, 90, 70, 77, 66, 84, 79, 108, 87, 107, 72, 89]
# y_base = [0, 100, 60, 90, 70, 88, 72, 86, 74, 84, 76, 82, 74, 80]
# Group 4
# y_base = [0, 90, 70, 100, 60, 131, 57, 144, 85, 109, 82, 124, 87, 101]
# y_base = [0, 90, 70, 100, 60, 150, 56, 112, 63, 95, 84, 118, 58, 110]
# y_base = [0, 90, 70, 100, 60, 145, 64, 112, 69, 86, 71, 119, 54, 95]
# y_base = [0, 90, 70, 100, 60, 105, 55, 110, 50, 115, 45, 120, 40, 125]
for i in range(11, len(y_base), 2):
y = y_base[:(i + 1)]
x = list(range(0, len(y)))
gg = [min(y[1], y[3])] * len(y)
dd = [max(y[2], y[4])] * len(y)
plt.figure(figsize=(i,4))
plt.title(y)
plt.grid()
plt.plot(x, y)
plt.plot(x, gg, '--')
plt.plot(x, dd, '--')
sx, sy = down_centre_expand_spliter(y)
plt.plot(sx, sy)
plt.show()
|
chan/01_Centre_Expand_Handler_Alpha_14.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Custom functions to clean data
# You'll now practice writing functions to clean data.
#
# The tips dataset has been pre-loaded into a DataFrame called tips. It has a 'sex' column that contains the values 'Male' or 'Female'. Your job is to write a function that will recode 'Female' to 0, 'Male' to 1, and return np.nan for all entries of 'sex' that are neither 'Female' nor 'Male'.
#
# Recoding variables like this is a common data cleaning task. Functions provide a mechanism for you to abstract away complex bits of code as well as reuse code. This makes your code more readable and less error prone.
#
# As Dan showed you in the videos, you can use the .apply() method to apply a function across entire rows or columns of DataFrames. However, note that each column of a DataFrame is a pandas Series. Functions can also be applied across Series. Here, you will apply your function over the 'sex' column.
#
# ### Instructions
#
# - Define a function named recode_gender() that has one parameter: gender.
# - If gender equals 'Male', return 1.
# - Else, if gender equals 'Female', return 0.
# - If gender does not equal 'Male' or 'Female', return np.nan. NumPy has been pre-imported for you.
# - Apply your recode_gender() function over tips.sex using the .apply() method to create a new column: 'recode'. Note that when passing in a function inside the .apply() method, you don't need to specify the parentheses after the function name.
# - Hit 'Submit Answer' and take note of the new 'recode' column in the tips DataFrame!
import pandas as pd
tips=pd.read_csv('tips.txt')
# +
# Define recode_gender()
def recode_gender(gender):
# Return 0 if gender is 'Female'
if gender == 'Female':
return 0
# Return 1 if gender is 'Male'
elif gender == 'Male':
return 1
# Return np.nan
else:
return np.nan
# Apply the function to the sex column
tips['recode'] = tips.sex.apply(recode_gender) # axis=0 will create an error
# Print the first five rows of tips
print(tips.head())
|
cleaning data in python/Cleaning data for analysis/06. Custom functions to clean data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Track Data Drift between Training and Inference Data in Production
#
# With this notebook, you will learn how to enable the DataDrift service to automatically track and determine whether your inference data is drifting from the data your model was initially trained on. The DataDrift service provides metrics and visualizations to help stakeholders identify which specific features cause the concept drift to occur.
#
# Please email <EMAIL> with any issues. A member from the DataDrift team will respond shortly.
#
# The DataDrift Public Preview API can be found [here](https://docs.microsoft.com/en-us/python/api/azureml-contrib-datadrift/?view=azure-ml-py).
# 
# # Prerequisites and Setup
# ## Install the DataDrift package
#
# Install the azureml-contrib-datadrift, azureml-contrib-opendatasets and lightgbm packages before running this notebook.
# ```
# pip install azureml-contrib-datadrift
# pip install azureml-contrib-datasets
# pip install lightgbm
# ```
# ## Import Dependencies
# +
import json
import os
import time
from datetime import datetime, timedelta
import numpy as np
import pandas as pd
import requests
from azureml.contrib.datadrift import DataDriftDetector, AlertConfiguration
from azureml.contrib.opendatasets import NoaaIsdWeather
from azureml.core import Dataset, Workspace, Run
from azureml.core.compute import AksCompute, ComputeTarget
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.experiment import Experiment
from azureml.core.image import ContainerImage
from azureml.core.model import Model
from azureml.core.webservice import Webservice, AksWebservice
from azureml.widgets import RunDetails
from sklearn.externals import joblib
from sklearn.model_selection import train_test_split
# -
# ## Set up Configuraton and Create Azure ML Workspace
#
# If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, go through the [configuration notebook](../../../configuration.ipynb) first if you haven't already to establish your connection to the AzureML Workspace.
# +
# Please type in your initials/alias. The prefix is prepended to the names of resources created by this notebook.
prefix = "dd"
# NOTE: Please do not change the model_name, as it's required by the score.py file
model_name = "driftmodel"
image_name = "{}driftimage".format(prefix)
service_name = "{}driftservice".format(prefix)
# optionally, set email address to receive an email alert for DataDrift
email_address = ""
# -
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# ## Generate Train/Testing Data
#
# For this demo, we will use NOAA weather data from [Azure Open Datasets](https://azure.microsoft.com/services/open-datasets/). You may replace this step with your own dataset.
# +
usaf_list = ['725724', '722149', '723090', '722159', '723910', '720279',
'725513', '725254', '726430', '720381', '723074', '726682',
'725486', '727883', '723177', '722075', '723086', '724053',
'725070', '722073', '726060', '725224', '725260', '724520',
'720305', '724020', '726510', '725126', '722523', '703333',
'722249', '722728', '725483', '722972', '724975', '742079',
'727468', '722193', '725624', '722030', '726380', '720309',
'722071', '720326', '725415', '724504', '725665', '725424',
'725066']
columns = ['usaf', 'wban', 'datetime', 'latitude', 'longitude', 'elevation', 'windAngle', 'windSpeed', 'temperature', 'stationName', 'p_k']
def enrich_weather_noaa_data(noaa_df):
hours_in_day = 23
week_in_year = 52
noaa_df["hour"] = noaa_df["datetime"].dt.hour
noaa_df["weekofyear"] = noaa_df["datetime"].dt.week
noaa_df["sine_weekofyear"] = noaa_df['datetime'].transform(lambda x: np.sin((2*np.pi*x.dt.week-1)/week_in_year))
noaa_df["cosine_weekofyear"] = noaa_df['datetime'].transform(lambda x: np.cos((2*np.pi*x.dt.week-1)/week_in_year))
noaa_df["sine_hourofday"] = noaa_df['datetime'].transform(lambda x: np.sin(2*np.pi*x.dt.hour/hours_in_day))
noaa_df["cosine_hourofday"] = noaa_df['datetime'].transform(lambda x: np.cos(2*np.pi*x.dt.hour/hours_in_day))
return noaa_df
def add_window_col(input_df):
shift_interval = pd.Timedelta('-7 days') # your X days interval
df_shifted = input_df.copy()
df_shifted['datetime'] = df_shifted['datetime'] - shift_interval
df_shifted.drop(list(input_df.columns.difference(['datetime', 'usaf', 'wban', 'sine_hourofday', 'temperature'])), axis=1, inplace=True)
# merge, keeping only observations where -1 lag is present
df2 = pd.merge(input_df,
df_shifted,
on=['datetime', 'usaf', 'wban', 'sine_hourofday'],
how='inner', # use 'left' to keep observations without lags
suffixes=['', '-7'])
return df2
def get_noaa_data(start_time, end_time, cols, station_list):
isd = NoaaIsdWeather(start_time, end_time, cols=cols)
# Read into Pandas data frame.
noaa_df = isd.to_pandas_dataframe()
noaa_df = noaa_df.rename(columns={"stationName": "station_name"})
df_filtered = noaa_df[noaa_df["usaf"].isin(station_list)]
df_filtered.reset_index(drop=True)
# Enrich with time features
df_enriched = enrich_weather_noaa_data(df_filtered)
return df_enriched
def get_featurized_noaa_df(start_time, end_time, cols, station_list):
df_1 = get_noaa_data(start_time - timedelta(days=7), start_time - timedelta(seconds=1), cols, station_list)
df_2 = get_noaa_data(start_time, end_time, cols, station_list)
noaa_df = pd.concat([df_1, df_2])
print("Adding window feature")
df_window = add_window_col(noaa_df)
cat_columns = df_window.dtypes == object
cat_columns = cat_columns[cat_columns == True]
print("Encoding categorical columns")
df_encoded = pd.get_dummies(df_window, columns=cat_columns.keys().tolist())
print("Dropping unnecessary columns")
df_featurized = df_encoded.drop(['windAngle', 'windSpeed', 'datetime', 'elevation'], axis=1).dropna().drop_duplicates()
return df_featurized
# -
# Train model on Jan 1 - 14, 2009 data
df = get_featurized_noaa_df(datetime(2009, 1, 1), datetime(2009, 1, 14, 23, 59, 59), columns, usaf_list)
df.head()
# +
label = "temperature"
x_df = df.drop(label, axis=1)
y_df = df[[label]]
x_train, x_test, y_train, y_test = train_test_split(df, y_df, test_size=0.2, random_state=223)
print(x_train.shape, x_test.shape, y_train.shape, y_test.shape)
training_dir = 'outputs/training'
training_file = "training.csv"
# Generate training dataframe to register as Training Dataset
os.makedirs(training_dir, exist_ok=True)
training_df = pd.merge(x_train.drop(label, axis=1), y_train, left_index=True, right_index=True)
training_df.to_csv(training_dir + "/" + training_file)
# -
# ## Create/Register Training Dataset
# +
dataset_name = "dataset"
name_suffix = datetime.utcnow().strftime("%Y-%m-%d-%H-%M-%S")
snapshot_name = "snapshot-{}".format(name_suffix)
dstore = ws.get_default_datastore()
dstore.upload(training_dir, "data/training", show_progress=True)
dpath = dstore.path("data/training/training.csv")
trainingDataset = Dataset.auto_read_files(dpath, include_path=True)
trainingDataset = trainingDataset.register(workspace=ws, name=dataset_name, description="dset", exist_ok=True)
trainingDataSnapshot = trainingDataset.create_snapshot(snapshot_name=snapshot_name, compute_target=None, create_data_snapshot=True)
datasets = [(Dataset.Scenario.TRAINING, trainingDataSnapshot)]
print("dataset registration done.\n")
datasets
# -
# ## Train and Save Model
# +
import lightgbm as lgb
train = lgb.Dataset(data=x_train,
label=y_train)
test = lgb.Dataset(data=x_test,
label=y_test,
reference=train)
params = {'learning_rate' : 0.1,
'boosting' : 'gbdt',
'metric' : 'rmse',
'feature_fraction' : 1,
'bagging_fraction' : 1,
'max_depth': 6,
'num_leaves' : 31,
'objective' : 'regression',
'bagging_freq' : 1,
"verbose": -1,
'min_data_per_leaf': 100}
model = lgb.train(params,
num_boost_round=500,
train_set=train,
valid_sets=[train, test],
verbose_eval=50,
early_stopping_rounds=25)
# +
model_file = 'outputs/{}.pkl'.format(model_name)
os.makedirs('outputs', exist_ok=True)
joblib.dump(model, model_file)
# -
# ## Register Model
# +
model = Model.register(model_path=model_file,
model_name=model_name,
workspace=ws,
datasets=datasets)
print(model_name, image_name, service_name, model)
# -
# # Deploy Model To AKS
#
# ## Prepare Environment
# +
myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn', 'joblib', 'lightgbm', 'pandas'],
pip_packages=['azureml-monitoring', 'azureml-sdk[automl]'])
with open("myenv.yml","w") as f:
f.write(myenv.serialize_to_string())
# -
# ## Create Image
# +
# Image creation may take up to 15 minutes.
image_name = image_name + str(model.version)
if not image_name in ws.images:
# Use the score.py defined in this directory as the execution script
# NOTE: The Model Data Collector must be enabled in the execution script for DataDrift to run correctly
image_config = ContainerImage.image_configuration(execution_script="score.py",
runtime="python",
conda_file="myenv.yml",
description="Image with weather dataset model")
image = ContainerImage.create(name=image_name,
models=[model],
image_config=image_config,
workspace=ws)
image.wait_for_creation(show_output=True)
else:
image = ws.images[image_name]
# -
# ## Create Compute Target
# +
aks_name = 'dd-demo-e2e'
prov_config = AksCompute.provisioning_configuration()
if not aks_name in ws.compute_targets:
aks_target = ComputeTarget.create(workspace=ws,
name=aks_name,
provisioning_configuration=prov_config)
aks_target.wait_for_completion(show_output=True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
else:
aks_target=ws.compute_targets[aks_name]
# -
# ## Deploy Service
# +
aks_service_name = service_name
if not aks_service_name in ws.webservices:
aks_config = AksWebservice.deploy_configuration(collect_model_data=True, enable_app_insights=True)
aks_service = Webservice.deploy_from_image(workspace=ws,
name=aks_service_name,
image=image,
deployment_config=aks_config,
deployment_target=aks_target)
aks_service.wait_for_deployment(show_output=True)
print(aks_service.state)
else:
aks_service = ws.webservices[aks_service_name]
# -
# # Run DataDrift Analysis
# ## Send Scoring Data to Service
# ### Download Scoring Data
# +
# Score Model on March 15, 2016 data
scoring_df = get_noaa_data(datetime(2016, 3, 15) - timedelta(days=7), datetime(2016, 3, 16), columns, usaf_list)
# Add the window feature column
scoring_df = add_window_col(scoring_df)
# Drop features not used by the model
print("Dropping unnecessary columns")
scoring_df = scoring_df.drop(['windAngle', 'windSpeed', 'datetime', 'elevation'], axis=1).dropna()
scoring_df.head()
# +
# One Hot Encode the scoring dataset to match the training dataset schema
columns_dict = model.datasets["training"][0].get_profile().columns
extra_cols = ('Path', 'Column1')
for k in extra_cols:
columns_dict.pop(k, None)
training_columns = list(columns_dict.keys())
categorical_columns = scoring_df.dtypes == object
categorical_columns = categorical_columns[categorical_columns == True]
test_df = pd.get_dummies(scoring_df[categorical_columns.keys().tolist()])
encoded_df = scoring_df.join(test_df)
# Populate missing OHE columns with 0 values to match traning dataset schema
difference = list(set(training_columns) - set(encoded_df.columns.tolist()))
for col in difference:
encoded_df[col] = 0
encoded_df.head()
# -
# Serialize dataframe to list of row dictionaries
encoded_dict = encoded_df.to_dict('records')
# ### Submit Scoring Data to Service
# +
# %%time
# retreive the API keys. AML generates two keys.
key1, key2 = aks_service.get_keys()
total_count = len(scoring_df)
i = 0
load = []
for row in encoded_dict:
load.append(row)
i = i + 1
if i % 100 == 0:
payload = json.dumps({"data": load})
# construct raw HTTP request and send to the service
payload_binary = bytes(payload,encoding = 'utf8')
headers = {'Content-Type':'application/json', 'Authorization': 'Bearer ' + key1}
resp = requests.post(aks_service.scoring_uri, payload_binary, headers=headers)
print("prediction:", resp.content, "Progress: {}/{}".format(i, total_count))
load = []
time.sleep(3)
# -
# ## Configure DataDrift
# +
services = [service_name]
start = datetime.now() - timedelta(days=2)
end = datetime(year=2020, month=1, day=22, hour=15, minute=16)
feature_list = ['usaf', 'wban', 'latitude', 'longitude', 'station_name', 'p_k', 'sine_hourofday', 'cosine_hourofday', 'temperature-7']
alert_config = AlertConfiguration([email_address]) if email_address else None
# there will be an exception indicating using get() method if DataDrift object already exist
try:
datadrift = DataDriftDetector.create(ws, model.name, model.version, services, frequency="Day", alert_config=alert_config)
except KeyError:
datadrift = DataDriftDetector.get(ws, model.name, model.version)
print("Details of DataDrift Object:\n{}".format(datadrift))
# -
# ## Run an Adhoc DataDriftDetector Run
target_date = datetime.today()
run = datadrift.run(target_date, services, feature_list=feature_list, create_compute_target=True)
exp = Experiment(ws, datadrift._id)
dd_run = Run(experiment=exp, run_id=run)
RunDetails(dd_run).show()
# ## Get Drift Analysis Results
# +
children = list(dd_run.get_children())
for child in children:
child.wait_for_completion()
drift_metrics = datadrift.get_output(start_time=start, end_time=end)
drift_metrics
# +
# Show all drift figures, one per serivice.
# If setting with_details is False (by default), only drift will be shown; if it's True, all details will be shown.
drift_figures = datadrift.show(with_details=True)
# -
# ## Enable DataDrift Schedule
datadrift.enable_schedule()
|
how-to-use-azureml/data-drift/azure-ml-datadrift.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## check overlapped exons in idt target regions
# > file `idt.bed`
#
# > tools `bedtools`
# + language="bash"
#
# bedtools merge -i idt.nochr.exome.sorted.bed -c 4 -o distinct | grep ',' > idt.overlap.bed
# -
import re
with open('idt.overlap.bed') as idt:
f = open('idt.ol.bed','w')
for l in idt.readlines():
l = l.strip('\n').split('\t')
m = list(set(re.findall('\(([^(\(|\))]*)\)', l[3])))
m = '|'.join(m)
l.append(m)
f.write('\t'.join(l) + '\n')
f.close()
import re
import pandas as pd
df = pd.read_table('idt.ol.bed', names=['chr','start','stop','overlap','gene'])
df.head(20)
|
notebooks/idtbed_merge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import os
import pandas as pd
from matplotlib import pyplot as plt
from matplotlib.colors import rgb_to_hsv
from imageio import imread
images = os.listdir('../image/')
blue_hue = 240 / 360 * np.pi * 2
result = pd.DataFrame(columns=['파일명','cossim'])
for img in images[:10]:
try:
rgb = imread(f'../image/{img}')
if rgb.shape[2] != 3:
result = result.append({'파일명' : img, 'cossim' : 'Error'}, ignore_index=True)
continue
hsv = rgb_to_hsv(rgb)
hue_list = []
for i in range(500):
for h, s, v in hsv[:,:,:][i]:
if v >= 254 and s <= 2:
continue
hue = h * np.pi * 2
cossim = np.cos(hue-blue_hue)
hue_list.append(cossim)
result = result.append({'파일명' : img, 'cossim' : np.mean(hue_list)}, ignore_index=True)
except:
result = result.append({'파일명' : img, 'cossim' : 'Error2'}, ignore_index=True)
result
for img in images[10:100]:
try:
rgb = imread(f'../image/{img}')
if rgb.shape[2] != 3:
result = result.append({'파일명' : img, 'cossim' : 'Error'}, ignore_index=True)
continue
hsv = rgb_to_hsv(rgb)
hue_list = []
for i in range(500):
for h, s, v in hsv[:,:,:][i]:
if v >= 254 and s <= 2:
continue
hue = h * np.pi * 2
cossim = np.cos(hue-blue_hue)
hue_list.append(cossim)
result = result.append({'파일명' : img, 'cossim' : np.mean(hue_list)}, ignore_index=True)
except:
result = result.append({'파일명' : img, 'cossim' : 'Error2'}, ignore_index=True)
for img in images[100:500]:
try:
rgb = imread(f'../image/{img}')
if rgb.shape[2] != 3:
result = result.append({'파일명' : img, 'cossim' : 'Error'}, ignore_index=True)
continue
hsv = rgb_to_hsv(rgb)
hue_list = []
for i in range(500):
for h, s, v in hsv[:,:,:][i]:
if v >= 254 and s <= 2:
continue
hue = h * np.pi * 2
cossim = np.cos(hue-blue_hue)
hue_list.append(cossim)
result = result.append({'파일명' : img, 'cossim' : np.mean(hue_list)}, ignore_index=True)
except:
result = result.append({'파일명' : img, 'cossim' : 'Error2'}, ignore_index=True)
for img in images[500:1000]:
try:
rgb = imread(f'../image/{img}')
if rgb.shape[2] != 3:
result = result.append({'파일명' : img, 'cossim' : 'Error'}, ignore_index=True)
continue
hsv = rgb_to_hsv(rgb)
hue_list = []
for i in range(500):
for h, s, v in hsv[:,:,:][i]:
if v >= 254 and s <= 2:
continue
hue = h * np.pi * 2
cossim = np.cos(hue-blue_hue)
hue_list.append(cossim)
result = result.append({'파일명' : img, 'cossim' : np.mean(hue_list)}, ignore_index=True)
except:
result = result.append({'파일명' : img, 'cossim' : 'Error2'}, ignore_index=True)
for img in images[1000:1500]:
try:
rgb = imread(f'../image/{img}')
if rgb.shape[2] != 3:
result = result.append({'파일명' : img, 'cossim' : 'Error'}, ignore_index=True)
continue
hsv = rgb_to_hsv(rgb)
hue_list = []
for i in range(500):
for h, s, v in hsv[:,:,:][i]:
if v >= 254 and s <= 2:
continue
hue = h * np.pi * 2
cossim = np.cos(hue-blue_hue)
hue_list.append(cossim)
result = result.append({'파일명' : img, 'cossim' : np.mean(hue_list)}, ignore_index=True)
except:
result = result.append({'파일명' : img, 'cossim' : 'Error2'}, ignore_index=True)
result.head()
result.tail()
for img in images[1500:2170]:
try:
rgb = imread(f'../image/{img}')
if rgb.shape[2] != 3:
result = result.append({'파일명' : img, 'cossim' : 'Error'}, ignore_index=True)
continue
hsv = rgb_to_hsv(rgb)
hue_list = []
for i in range(500):
for h, s, v in hsv[:,:,:][i]:
if v >= 254 and s <= 2:
continue
hue = h * np.pi * 2
cossim = np.cos(hue-blue_hue)
hue_list.append(cossim)
result = result.append({'파일명' : img, 'cossim' : np.mean(hue_list)}, ignore_index=True)
except:
result = result.append({'파일명' : img, 'cossim' : 'Error2'}, ignore_index=True)
result.shape
result.to_csv('data/cossim_result_blue.csv', index=False)
# +
filenames = result['파일명'].tolist()
CategoryIDs = [f.split('-')[0] for f in filenames]
ProductIDs = []
for f in filenames:
try:
ProductIDs.append(f.split('-')[1].split('.png')[0])
except:
ProductIDs.append(f)
# -
result['ProductID'] = ProductIDs
result['CategoryID'] = CategoryIDs
result['카테고리명'] = result['CategoryID'].astype("category")
names = 'girl-RolePlay,girl-Doll,girl-Deco,girl-DIY,boy-RolePlay,boy-Action,boy-Control,boy-Car/Train,boy-Figure'.split(',')
result['카테고리명'].cat.categories = names
result.head()
result['성구분'] = [n.split('-')[0] for n in result['카테고리명'].tolist()]
result.head()
result.to_csv('data/cossim_result_blue_with_IDs_eng.csv', index=False)
result_no_error = result[result.cossim != 'Error']
result_no_error = result_no_error[result.cossim != 'Error2']
result_no_error.shape
result_no_error['cossim'] = result_no_error['cossim'].astype('float64')
result_no_error['cossim'].describe()
|
color analysis/soryeongk_blue_hue.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <table>
# <tr><td><img style="height: 150px;" src="images/geo_hydro1.jpg"></td>
# <td bgcolor="#FFFFFF">
# <p style="font-size: xx-large; font-weight: 900; line-height: 100%">AG Dynamics of the Earth</p>
# <p style="font-size: large; color: rgba(0,0,0,0.5);">Jupyter notebooks</p>
# <p style="font-size: large; color: rgba(0,0,0,0.5);"><NAME></p>
# </td>
# </tr>
# </table>
# # Dynamic systems: 8. Elastic material
# ## Plate in hole with `solidDisplacementFoam`
# ----
# *<NAME>,
# Geophysics Section,
# Institute of Geological Sciences,
# Freie Universität Berlin,
# Germany*
# **In this notebook, we will learn to**
#
# - calculate stresses and displacements in an **elastic plate with a hole**,
# using `solidDisplacementFoam` as solver.
#
# **Prerequisites:** (text)
#
# **Result:** You should get a figure similar to
# <img src="images/plateDx.jpg" style=width:10cm>
#
# <a href="#top">**Table of contents**</a>
#
# 1. [Solver and equations](#one)
# 2. [Implementation](#two)
# 3. [Running](#three)
# 4. [Post-processing](#four)
# 5. [Technical aspects](#five)
# <div id="one"></div>
#
# ----
# ## 1. Solver and equations
#
# `solidDisplacementFoam` is a
#
# - transient
# - incompressible
#
# solver for the continuity and momentum equations:
# $$
# \begin{array}{rcl}
# \frac{\partial^2 \rho \vec{r}}{\partial t^2} &=& \nabla \cdot \mathbb{\sigma} \\
# \mathbb{\sigma} &=& 2 \mu \mathbb{\epsilon} + \lambda tr(\mathbb{\epsilon}) \mathbb{I} \\
# \mathbb{\epsilon} &=& \frac{1}{2} \left[ \nabla \vec{r} + (\nabla \vec{r})^T \right]
# \end{array}
# $$
# with
# $\vec{r}$ [m] the displacement,
# $\mathbb{\sigma}$ [Pa] the Cauchy stress tensor,
# $\mathbb{\epsilon}$ [-] the strain tensor,
# $\mathbb{I}$ [-] the unity tensor,
# $\rho$ [kg/m$^3$] density,
# $\lambda$ [Pa] first Lame parameterm
# $\mu$ [Pa] second Lame parameter (shear modulus),
# $t$ [s] time,
# $\nabla$ [1/m] Nabla operator.
#
# On input first and second Lame parameters are defined as Youg modulus $E$ [Pa] and
# Poisson ratio $\nu$ [-]:
# $$
# \begin{array}{rcl}
# E &=& \frac{\mu (3\lambda + 2\mu)}{\lambda + \mu} \\
# \nu &=& \frac{\lambda}{2(\lambda + \mu)}
# \end{array}
# $$
#
# The equations are then solved by `solidDisplacementFoam`.
#
# **NOTE: `solidDisplacementFoam` returns not stresses, but stress times density!**
# <div id="two"></div>
#
# ----
# ## 2. Implementation
#
# We consider a block with 4m side length, and a hole in the center with $a=0.5$m radius.
# The stress is applied in the normal direction of the eastern and western (left and right) faces,
# $\sigma_{xx}$ [Pa].
# <img src="images/PlateInHole.jpg" style=width:15cm>
#
# As the problem is symmetric with respect to the origin of the hole,
# we only need to consider a quarter of the domain,
# with symmetry boundary conditions.
#
# ### Directory structure and files
#
# ~~~
# PlateHole_solidDisplacementFoam
# |-- 0
# |-- D
# |-- T
# |- constant
# |-- mechanicalProperties
# |-- thermalProperties
# |- system
# |-- blockMesh
# |-- controlDict
# |-- fvSchemes
# |-- fvSolution
# ~~~
#
# - `system/blockMeshDict`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# convertToMeters 1;
#
# vertices
# (
# (0.5 0 0)
# (1 0 0)
# (2 0 0)
# (2 0.707107 0)
# (0.707107 0.707107 0)
# (0.353553 0.353553 0)
# (2 2 0)
# (0.707107 2 0)
# (0 2 0)
# (0 1 0)
# (0 0.5 0)
# (0.5 0 0.5)
# (1 0 0.5)
# (2 0 0.5)
# (2 0.707107 0.5)
# (0.707107 0.707107 0.5)
# (0.353553 0.353553 0.5)
# (2 2 0.5)
# (0.707107 2 0.5)
# (0 2 0.5)
# (0 1 0.5)
# (0 0.5 0.5)
# );
#
# blocks
# (
# hex (5 4 9 10 16 15 20 21) (10 10 1) simpleGrading (1 1 1)
# hex (0 1 4 5 11 12 15 16) (10 10 1) simpleGrading (1 1 1)
# hex (1 2 3 4 12 13 14 15) (20 10 1) simpleGrading (1 1 1)
# hex (4 3 6 7 15 14 17 18) (20 20 1) simpleGrading (1 1 1)
# hex (9 4 7 8 20 15 18 19) (10 20 1) simpleGrading (1 1 1)
# );
#
# edges
# (
# arc 0 5 (0.469846 0.17101 0)
# arc 5 10 (0.17101 0.469846 0)
# arc 1 4 (0.939693 0.34202 0)
# arc 4 9 (0.34202 0.939693 0)
# arc 11 16 (0.469846 0.17101 0.5)
# arc 16 21 (0.17101 0.469846 0.5)
# arc 12 15 (0.939693 0.34202 0.5)
# arc 15 20 (0.34202 0.939693 0.5)
# );
#
# boundary
# (
# west
# {
# type symmetryPlane;
# faces
# (
# (8 9 20 19)
# (9 10 21 20)
# );
# }
# east
# {
# type patch;
# faces
# (
# (2 3 14 13)
# (3 6 17 14)
# );
# }
# bottom
# {
# type symmetryPlane;
# faces
# (
# (0 1 12 11)
# (1 2 13 12)
# );
# }
#
# top
# {
# type patch;
# faces
# (
# (7 8 19 18)
# (6 7 18 17)
# );
# }
# hole
# {
# type patch;
# faces
# (
# (10 5 16 21)
# (5 0 11 16)
# );
# }
# frontAndBack
# {
# type empty;
# faces
# (
# (10 9 4 5)
# (5 4 1 0)
# (1 4 3 2)
# (4 7 6 3)
# (4 9 8 7)
# (21 16 15 20)
# (16 11 12 15)
# (12 13 14 15)
# (15 14 17 18)
# (15 18 19 20)
# );
# }
# );
#
# mergePatchPairs
# (
# );
# ~~~
# </details>
#
# - `0/D`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# dimensions [0 1 0 0 0 0 0];
#
# internalField uniform (0 0 0);
#
# boundaryField
# {
# west
# {
# type symmetryPlane;
# }
# east
# {
# type tractionDisplacement;
# traction uniform (1e4 0 0);
# pressure uniform 0;
# value uniform (0 0 0);
# }
# bottom
# {
# type symmetryPlane;
# }
# top
# {
# type tractionDisplacement;
# traction uniform (0 0 0);
# pressure uniform 0;
# value uniform (0 0 0);
# }
# hole
# {
# type tractionDisplacement;
# traction uniform (0 0 0);
# pressure uniform 0;
# value uniform (0 0 0);
# }
# frontAndBack
# {
# type empty;
# }
# }
# ~~~
# </details>
#
# - `0/T`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# dimensions [0 0 0 1 0 0 0];
#
# internalField uniform 300;
#
# boundaryField
# {
# west
# {
# type symmetryPlane;
# }
# east
# {
# type zeroGradient;
# }
# bottom
# {
# type symmetryPlane;
# }
# top
# {
# type zeroGradient;
# }
# hole
# {
# type zeroGradient;
# }
# frontAndBack
# {
# type empty;
# }
# }
# ~~~
# </details>
#
# - `constant/mechanicalProperties`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# rho
# { type uniform;
# value 2600; }
#
# nu
# { type uniform;
# value 0.25; }
#
# E
# { type uniform;
# value 1e+10; }
#
# planeStress yes;
# ~~~
# </details>
#
# - `constant/thermalProperties`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# Cp
# { type uniform;
# value 434; }
#
# kappa
# { type uniform;
# value 60.5; }
#
# alphav
# { type uniform;
# value 1.1e-05; }
#
# thermalStress no;
# ~~~
# </details>
#
# - `system/controlDict`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# application solidDisplacementFoam;
# startFrom startTime;
# startTime 0;
# stopAt endTime;
# endTime 100;
# deltaT 1;
# writeControl timeStep;
# writeInterval 50;
# purgeWrite 0;
# writeFormat ascii;
# writePrecision 6;
# writeCompression off;
# timeFormat general;
# timePrecision 6;
# graphFormat raw;
# runTimeModifiable true;
# ~~~
# </details>
#
# - `system/fvSchemes`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# d2dt2Schemes
# {
# default steadyState;
# }
#
# ddtSchemes
# {
# default Euler;
# }
#
# gradSchemes
# {
# default leastSquares;
# grad(D) leastSquares;
# grad(T) leastSquares;
# }
#
# divSchemes
# {
# default none;
# div(sigmaD) Gauss linear;
# }
#
# laplacianSchemes
# {
# default none;
# laplacian(DD,D) Gauss linear corrected;
# laplacian(kappa,T) Gauss linear corrected;
# }
#
# interpolationSchemes
# {
# default linear;
# }
#
# snGradSchemes
# {
# default none;
# }
#
# ~~~
# </details>
#
# - `system/fvSolution`
# (text)
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# solvers
# {
# "(D|T)"
# {
# solver GAMG;
# tolerance 1e-06;
# relTol 0.9;
# smoother GaussSeidel;
# nCellsInCoarsestLevel 20;
# }
# }
#
# stressAnalysis
# {
# compactNormalStress yes;
# nCorrectors 1;
# D 1e-06;
# }
# ~~~
# </details>
# <div id="three"></div>
#
# ----
# ## 3. Running
#
# Running a particular example is done with the following set of commands:
# ~~~
# $ foamCleanTutorials
# $ blockMesh
# $ solidDisplacementFoam
# ~~~
#
# <img src="images/plateDx.jpg" style=width:10cm>
# <img src="images/plateDy.jpg" style=width:10cm>
# <div id="four"></div>
#
# ----
# ## 4. Post-processing: Analytical solution
#
# This problem has an analytical solution for stresses and displacements, based on a
# *cylindric coordinate system* with $r$ [m] the radial and $\theta$ the angular coordinate,
# and $T$ [Pa] the applied boundary stress.
#
# For the stresses, we find:
# $$
# \begin{array}{rcl}
# \sigma_{rr} &=& \frac{T}{2} \left( 1 - \frac{a^2}{r^2} \right)
# + \frac{T \cos(2\theta)}{2} \left( \frac{3a^4}{r^4} - \frac{4a^2}{r^2} + 1\right) \\
# \sigma_{\theta\theta} &=& \frac{T}{2} \left( 1 + \frac{a^2}{r^2} \right)
# - \frac{T \cos(2\theta)}{2} \left( \frac{3a^4}{r^4} + 1\right) \\
# \sigma_{r\theta} &=& \frac{T \cos(2\theta)}{2} \left( \frac{3a^4}{r^4} - \frac{2a^2}{r^2} - 1\right)
# \end{array}
# $$
#
# For the displacements, we find:
# $$
# \begin{array}{rcl}
# u_{r} &=& \frac{T r \cos(2\theta)}{2E} \left[ (1+\nu) + \frac{4a^2}{r^2} - (1+\nu) \frac{a^4}{r^4} \right]
# + \frac{T r}{2E} \left[ (1-\nu) + (1+\nu) \frac{a^2}{r^2} \right] \\
# u_{\theta} &=& -\frac{T r \sin(2\theta)}{2E} \left[ (1+\nu) + 2(1-\nu) \frac{a^2}{r^2} + (1+\nu) \frac{a^4}{r^4} \right]
# \end{array}
# $$
# with $E$ [Pa] Young modulus, $\nu$ Poisson ratio.
# ## 4. Post-processing: Profiles
#
# Extract stress and displacement from `solidDisplacementFoam` with
#
# ~~~
# $ postProcess -func 'components(sigma)'
# $ postProcess -func 'components(D)'
# $
# $ postProcess -func sampleDict -latestTime
# ~~~
#
# The **first two** commands generate components for stress and displacements, stored in the
# time directories.
#
# The **third** command uses the dictionary `system/sampleDict` to sample stress and displacement
# along the $x$ and $y$ axes:
#
# <details><summary><b>> Show code</b></summary>
#
# ~~~
# type sets;
#
# setFormat raw;
#
# interpolationScheme cell;
# //interpolationScheme cellPoint;
# //interpolationScheme cellPointFace;
#
# // Fields to sample.
# fields
# ( Dx
# Dy
# sigmaxx
# );
#
# sets
# (
# PlateWithHole_y
# {
# type uniform;
# nPoints 100;
# axis xyz;
# start ( 0 0.0 0.0);
# end ( 0 2.0 0.0);
# }
#
# PlateWithHole_x
# {
# type uniform;
# nPoints 100;
# axis xyz;
# start ( 0 0.0 0.0);
# end ( 2 0.0 0.0);
# }
#
# );
# ~~~
# </details>
# We obtain the two files
#
# - `postProcessing/sampleDict/100/PlateWithHole_x_Dx_Dy_sigmaxx.xy`
# - `postProcessing/sampleDict/100/PlateWithHole_y_Dx_Dy_sigmaxx.xy`
#
# We then use `python`to plot stresses and displacements, and compare them to the
# analytical solution:
# - We plot the stress $\sigma_{xx}$ along the $y$ axis for $x=0$. In our analytical solution,
# this is $\sigma_{\theta\theta}(\theta=90)$.
#
# - We plot the deformation $D_x$ along the $x$ axis for $y=0$. In our analytical solution,
# this is $u_r(\theta=0)$.
# +
import numpy as np
import matplotlib.pyplot as plt
# calculate analytical data
def sigma_thetatheta(r,theta=90.,T=1e4,a=0.5):
sigma_thetatheta = T/2*(1 + a**2/r**2) - T*np.cos(2*theta*np.pi/180.)/2 * (3*a**4/r**4 + 1)
return sigma_thetatheta
def ur(r,theta=90.,T=1e4,E=1e10,nu=0.25,a=0.5):
ur = (T*r*np.cos(2*theta*np.pi/180.)/2/E * ((1+nu) + 4*a**2/r**2 - (1+nu)*a**4/r**4)
+ T*r/2/E * ((1-nu) + (1+nu)*a**2/r**2))
return ur
# load laplacianFoam postprocessed data
data1x = np.loadtxt('data/PlateWithHole1_x_Dx_Dy_sigmaxx.xy')
data1y = np.loadtxt('data/PlateWithHole1_y_Dx_Dy_sigmaxx.xy')
data2x = np.loadtxt('data/PlateWithHole2_x_Dx_Dy_sigmaxx.xy')
data2y = np.loadtxt('data/PlateWithHole2_y_Dx_Dy_sigmaxx.xy')
#print(data1)
y=np.linspace(0.5,2,21)
fig, axs = plt.subplots(3, 1,figsize=(12,10))
print(axs)
axs[0].set_xlabel('y [m]')
axs[0].set_ylabel('$\sigma_{xx}$ [kPa]')
axs[0].set_title('Stress')
axs[0].plot(y,sigma_thetatheta(y)/1e3,linewidth=2,color='red',label='analytical')
axs[0].plot(data1y[:,1],data1y[:,5]/1e3,linewidth=2,color='green',label='numerical')
axs[0].plot(data2y[:,1][data2y[:,1]<2.1],data2y[:,5][data2y[:,1]<2.1]/1e3,
linewidth=2,linestyle=':',color='green',label='numerical (larger grid)')
axs[0].legend()
axs[1].set_xlabel('x [m]')
axs[1].set_ylabel('$D_x$ [$\mu$m]')
axs[1].set_title('Displacement')
axs[1].plot(y,ur(y,theta=0)*1e6,linewidth=2,color='red',label='analytical')
axs[1].plot(data1x[:,0],data1x[:,3]*1e6,linewidth=2,color='green',label='numerical')
axs[1].plot(data2x[:,0][data2x[:,0] < 2.1],data2x[:,3][data2x[:,0] < 2.1]*1e6,
linewidth=2,linestyle=':',color='green',label='numerical (larger grid)')
axs[1].legend()
axs[2].set_xlabel('y [m]')
axs[2].set_ylabel('$D_y$ [$\mu$m]')
axs[2].set_title('Displacement')
axs[2].plot(y,ur(y,theta=90)*1e6,linewidth=2,color='red',label='analytical')
axs[2].plot(data1y[:,1],data1y[:,4]*1e6,linewidth=2,color='green',label='numerical')
axs[2].plot(data2y[:,1][data2y[:,1] < 2.1],data2y[:,4][data2y[:,1] < 2.1]*1e6,
linewidth=2,linestyle=':',color='green',label='numerical (larger grid)')
axs[2].legend()
plt.tight_layout()
# -
# **NOTE: Where does the offset in $u_r$ come from?** From the finite size of the grid.
# ... done
|
Dynamics_lab08_HoleInPlate_solidDisplacementFoam.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# # Defining data types
# We can define types (i.e. data structures) ourselves using the `struct` keyword.
#
# It is a convention that type names are capitalized and [camel cased](https://en.wikipedia.org/wiki/Camel_case).
#
# (Note that types can not be redefined - you have to restart your Julia session to change a type definiton.)
struct MyType end
# To create an object of type `MyType` we have to call a [constructor](https://docs.julialang.org/en/latest/manual/constructors/). Loosely speaking, a constructor is a function that create new objects.
#
# Julia automatically creates a trivial constructors for us, which has the same name as the type.
methods(MyType)
m = MyType()
typeof(m)
m isa MyType
# Since no data is contained in our `MyType` - it is a so-called *singleton type* - we can basically only use it for dispatch.
# Most of the time, we'll want a self-defined type to hold some data. For this, we need *fields*.
struct A
x::Int64
end
A()
# The default constructor always expects values for all fields.
A(3)
a = A(3)
# a.<TAB>
a.x
# Note that types defined with `struct` are **immutable**, that is the values of it's fields cannot be changed.
a.x = 2
mutable struct B
x::Int64
end
b = B(3)
b.x
b.x = 4
b.x
# Note, however, that **immutability is not recursive**.
struct C
x::Vector{Int64}
end
c = C([1, 2, 3])
c.x
c.x = [3,4,5]
c.x[1] = 3
c.x
c.x .= [3,4,5] # dot to perform the assignment element-wise
# Abstract types are just as easy to define using the keyword `abstract type`.
abstract type MyAbstractType end
# Since abstract types don't have fields, they only (informally) define interfaces and can be used for dispatch.
struct MyConcreteType <: MyAbstractType # subtype
somefield::String
end
c = MyConcreteType("test")
c isa MyAbstractType
supertype(MyConcreteType)
subtypes(MyAbstractType)
# # Custom constructor
struct VolNaive
value::Float64
end
VolNaive(3.0)
VolNaive(-3.0)
struct VolSimple
value::Float64
function VolSimple(x) # inner constructor. function name must match the type name.
if !(x isa Real)
throw(ArgumentError("Must be real"))
end
if x < 0
throw(ArgumentError("Negative volume not allowed."))
end
new(x) # within an inner constructor, the `new` function can be used to create an object.
end
end
# ---
#
# **Side note:**
#
# ```julia
# if !(x isa Real)
# throw(ArgumentError("Must be real"))
# end
# if x < 0
# throw(ArgumentError("Negative volume not allowed."))
# end
# ```
#
# This can be written more compactly as
# ```julia
# x isa Real || throw(ArgumentError("Must be real"))
# x < 0 && throw(ArgumentError("Negative volume not allowed."))
# ```
#
# See ["short-circuit evaluation"](https://docs.julialang.org/en/latest/manual/control-flow/#Short-Circuit-Evaluation-1) for more information.
#
# ---
VolSimple(3.0)
VolSimple(-3.0)
VolSimple("test")
VolSimple(3) # implicit conversion from Int64 -> Float64
# # Parametric types
# Volumes don't have to be `Float64` values. We can easily relax our type definition to allow all sorts of internal value types.
struct VolParam{T}
value::T
function VolParam(x::T) where T # x can be of any type T
if !(x isa Real)
throw(ArgumentError("Must be real"))
end
if x < 0
throw(ArgumentError("Negative volume not allowed."))
end
new{T}(x) # Note that we need an extra {T} here
end
end
VolParam(3.0)
VolParam(3)
# Instead of checking the realness of the input `x` explicitly in the inner constructor, we can impose type constraints in the type and function signatures.
struct Vol{T<:Real} <: Real # the last <: Real tells Julia that a Vol is a subtype of Real, i.e. basically a real number
value::T
function Vol(x::T) where T<:Real # x can be of any type T<:Real
x < 0 && throw(ArgumentError("Negative volume not allowed."))
new{T}(x)
end
end
Vol(3)
Vol(3.0)
Vol("1.23")
Vol(-2)
# # Arithmetic
Vol(3) + Vol(4)
+(x::Vol, y::Vol) = Vol(x.value + y.value)
# If we want to extend or override functions that already exit, we need to `import` them first.
# +
import Base: +
+(x::Vol, y::Vol) = Vol(x.value + y.value)
# -
Vol(3) + Vol(4)
Vol(2) + Vol(8.3) # implicit conversion!
methodswith(Vol)
# +
import Base: -, *
-(x::Vol, y::Vol) = Vol(x.value - y.value)
*(x::Vol, y::Vol) = Vol(x.value * y.value)
# -
# Now that we have addition defined for our volume type, some functions already **just work**.
sum([Vol(3), Vol(4.8), Vol(1)])
M = Vol.(rand(3,3))
N = Vol.(rand(3,3))
M + N
# Whenever something doesn't work, we implement the necessary functions.
sin(Vol(3))
import Base: AbstractFloat
AbstractFloat(x::Vol{T}) where T = AbstractFloat(x.value)
sin(Vol(3))
sqrt(Vol(3))
# If we really wanted to have `Vol{T}` objects behave like real numbers in all operations, we'd have to do a bit more work like specifying [promotion and conversion rules](https://docs.julialang.org/en/latest/manual/conversion-and-promotion/).
# An important thing to note is that **user defined types are just as good as built-in types**!
#
# There is nothing special about built-in types. In fact, [they are implemented in the same way](https://github.com/JuliaLang/julia/blob/master/stdlib/LinearAlgebra/src/diagonal.jl#L5)!
# Let us quickly confirm that our volume "wrapper" type does not come with any performance overhead by benchmarking it in a simple function.
# # Benchmarking with `BenchmarkTools.jl`
using BenchmarkTools
operation(x) = x^2 + sqrt(x)
x = rand(2,2)
@time operation.(x)
function f()
x = rand(2,2)
@time operation.(x)
end
f()
# We should wrap benchmarks into functions!
#
# Fortunately, there are tools that do this for us. In addition, they also collect some statistics by running the benchmark multiple times.
@benchmark operation.(x)
# Typically we don't need all this information. Just use `@btime` instead of `@time`!
@btime operation.(x);
# However, we still have to take some care to avoid accessing global variables.
@btime operation.($x); # interpolate the value of x into the expression to avoid overhead of globals
# More information: [BenchmarkTools.jl](https://github.com/JuliaCI/BenchmarkTools.jl/blob/master/doc/manual.md).
# Finally, we can check the performance of our custom volume type.
@btime sqrt(Vol(3));
@btime sqrt(3);
# # Core messages of this Notebook
#
# * There are `mutable struct`s and immutable `struct`s and immutability is not recursive.
# * **Contructors** are functions that create objects. In an inner constructor we can use the function `new` to generate objects.
# * We can easily **extend `Base` functions** for our types to implement arithmetics and such.
# * We should always benchmark our code with **BenchmarkTools.jl's @btime and @benchmark**.
|
JuliaWorkshop/Part1-BasicJulia/10. Advanced - User Defined Types.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: slepc4
# language: python
# name: slepc4
# ---
from noisegen import NoiseGenerator
import numpy as np
import matplotlib
# %matplotlib inline
matplotlib.rcParams.update({'font.size': 18})
# In this notebook we will generate real valued samples of pink noise. First we must specify which frequencies will be included in samples. We choose to include 1001 evenly spaced frequencies between 0 Hz and 100 Hz.
n_frequencies = 1001
f_interval = 0.1
generator = NoiseGenerator(n_frequencies=n_frequencies, f_interval=f_interval)
# Next we specify the power spectral density to be pink with an infrared cutoff at 10 Hz. We take the variance of the noise samples to be 2.0.
variance = 2.0
f_ir = 1.0
generator.specify_psd('pink', f_ir=f_ir, normalization=variance)
# Then we generate 1000 time series of the noise.
n_traces = 1000
generator.generate_trace(n_traces=n_traces)
# We can now plot some of the samples we have generated.
axes = generator.samples.iloc[:,0:3].plot()
axes.set_ylabel(r'$Y(t)$')
axes.set_xlabel('$t$ (s)')
# We may wish to perform some diagnostics on our noise to check that it has the properties we desire. First we will calculate its variance.
measured_variance = np.var(generator.samples.values.flatten())
print('Variance is found to be '+str(measured_variance))
# This figure looks close to our desired value of 2.0. Next we will plot the power spectral density.
axes = generator.plot_psd()
axes.legend(['Meausred PSD','Specified PSD'])
#axes.set_ylim([0,0.015])
xlabel = axes.set_xlabel(r'$f$ (Hz)')
ylabel = axes.set_ylabel(r'$S(f)$ (Hz$^{-1}$)')
# Finally we plot the autocorrelation function of the noise.
axes = generator.plot_autocorrelation()
axes.set_xlabel(r'$t$ (s)')
axes.set_ylabel(r'$\langle Y(t) Y(0) \rangle$')
# The noise appears to have a finite correlation tiem and a variance close to 2.0.
|
docs/examples/ex_pink_noise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
import xml.etree.ElementTree as ET
import pandas as pd
import nltk
from nltk.stem import WordNetLemmatizer
from nltk.stem import PorterStemmer
from nltk.tokenize import word_tokenize
# + pycharm={"is_executing": false}
Posts_tree = ET.parse('Data_Dump/Posts.xml')
Posts_root = Posts_tree.getroot()
len(Posts_root)
# + pycharm={"is_executing": false}
for child in Posts_root[:5]:
print(child.attrib)
# + pycharm={"is_executing": false}
df = pd.DataFrame(columns = ['Title', 'Body'])
# + pycharm={"is_executing": false}
posts = []
for child in Posts_root:
posts.append(child.attrib)
print(posts[0])
# + pycharm={"is_executing": false}
def remove_html_tags(text):
"""Remove html tags from a string"""
import re
clean = re.compile('<.*?>')
return re.sub(clean, '', text)
ps = PorterStemmer()
lemmatizer = WordNetLemmatizer()
# + pycharm={"is_executing": false}
for i, post in enumerate(posts):
if 'Title' in post:
df = df.append({'Title': post['Title'], 'Body': remove_html_tags(post['Body'])}, ignore_index=True)
# + pycharm={"is_executing": false}
df.head(6)
# + pycharm={"is_executing": false}
## tokenize, stem and lemmatize
for i, j in df.iterrows():
title_tokens_stemmed = [ps.stem(word) for word in word_tokenize(j["Title"])]
title_tokens_stemmed_lemmetized = [lemmatizer.lemmatize(word) for word in title_tokens_stemmed]
title_tokens_stemmed_lemmetized = ' '.join(title_tokens_stemmed_lemmetized)
df.at[i, "Title"] = title_tokens_stemmed_lemmetized
body_tokens_stemmed = [ps.stem(word) for word in word_tokenize(j["Body"])]
body_tokens_stemmed_lemmetized = [lemmatizer.lemmatize(word) for word in body_tokens_stemmed]
body_tokens_stemmed_lemmetized = ' '.join(body_tokens_stemmed_lemmetized)
df.at[i, "Body"] = body_tokens_stemmed_lemmetized
# + pycharm={"is_executing": false}
df.head(5)
# + pycharm={"is_executing": false}
from sklearn.feature_extraction.text import CountVectorizer
count_vectorizer = CountVectorizer(stop_words='english')
# + pycharm={"is_executing": false, "name": "#%%\n"}
from sklearn.decomposition import LatentDirichletAllocation as LDA
from pyLDAvis import sklearn as sklearn_lda
import pyLDAvis
def print_topics(model, count_vectorizer, n_top_words):
words = count_vectorizer.get_feature_names()
for topic_idx, topic in enumerate(model.components_):
print('Topic {}: {}'.format(topic_idx, ' | '.join([words[i] for i in topic.argsort()[:-n_top_words - 1:-1]])))
number_topics = 10
number_words = 10
# fit the LDA model on all of the question titles for the data
post_titles = count_vectorizer.fit_transform(df['Title'])
lda_titles = LDA(n_components=number_topics)
lda_titles.fit(post_titles)
# fit the LDA model on all of the question bodies for the data
post_body = count_vectorizer.fit_transform(df['Title'])
lda_body = LDA(n_components=number_topics)
lda_body.fit(post_body)
# -
# print the top topics for titles
print("Post Titles: Top topics")
print()
print_topics(lda_titles, count_vectorizer, number_words)
# print the top topics for post contents
print("Post Content: Top topics")
print_topics(lda_body, count_vectorizer, number_words)
# + pycharm={"name": "#%%\n"}
# display the topic mappings with top words for 'titles'
visual_lda_titles = sklearn_lda.prepare(lda_titles,post_titles, count_vectorizer)
pyLDAvis.display(visual_lda_titles)
# -
# display the topic mappings with top words for 'post contents'
visual_lda_body = sklearn_lda.prepare(lda_body,post_body, count_vectorizer)
pyLDAvis.display(visual_lda_body)
|
Stack_Overflow_Data_Dump_Preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pathlib
BASE_DIR = pathlib.Path().resolve().parent
DATASET_DIR = BASE_DIR / "datasets"
ZIPS_DIR = DATASET_DIR / 'zips'
ZIPS_DIR.mkdir(exist_ok=True, parents=True)
SPAM_SMS_ZIP_PATH = ZIPS_DIR / "sms-spam-dataset.zip"
SPAM_YOUTUBE_ZIP_PATH = ZIPS_DIR / "youtube-spam-dataset.zip"
# -
SMS_SPAM_ZIP = "https://archive.ics.uci.edu/ml/machine-learning-databases/00228/smsspamcollection.zip"
YOUTUBE_SPAM_ZIP = "https://archive.ics.uci.edu/ml/machine-learning-databases/00380/YouTube-Spam-Collection-v1.zip"
# +
# !curl $SMS_SPAM_ZIP -o $SPAM_SMS_ZIP_PATH
# !curl $YOUTUBE_SPAM_ZIP -o $SPAM_YOUTUBE_ZIP_PATH
# -
|
nbs/1 - Download Datasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="b518b04cbfe0"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="906e07f6e562"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="a81c428fc2d3"
# # 迁移学习和微调
# + [markdown] id="3e5a59f0aefd"
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://tensorflow.google.cn/guide/keras/transfer_learning"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png">在 TensorFlow.org 上查看</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/guide/keras/transfer_learning.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png">在 Google Colab 中运行 </a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/guide/keras/transfer_learning.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png">在 GitHub 上查看源代码</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/guide/keras/transfer_learning.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png">下载笔记本</a></td>
# </table>
# + [markdown] id="8d4ac441b1fc"
# ## 设置
# + id="9a7e9b92f963"
import numpy as np
import tensorflow as tf
from tensorflow import keras
# + [markdown] id="00d4c41cfe2f"
# ## 简介
#
# **迁移学习**包括获取从迁移学习包括获取从一个问题中学习到的特征,然后将这些特征用于新的类似问题。例如,来自已学会识别浣熊的模型的特征可能对建立旨在识别狸猫的模型十分有用。
#
# 对于数据集中的数据太少而无法从头开始训练完整模型的任务,通常会执行迁移学习。
#
# 在深度学习情境中,迁移学习最常见的形式是以下工作流:
#
# 1. 从之前训练的模型中获取层。
# 2. 冻结这些层,以避免在后续训练轮次中破坏它们包含的任何信息。
# 3. 在已冻结层的顶部添加一些新的可训练层。这些层会学习将旧特征转换为对新数据集的预测。
# 4. 在您的数据集上训练新层。
#
# 最后一个可选步骤是**微调**,包括解冻上面获得的整个模型(或模型的一部分),然后在新数据上以极低的学习率对该模型进行重新训练。以增量方式使预训练特征适应新数据,有可能实现有意义的改进。
#
# 首先,我们将详细介绍 Keras trainable API,它是大多数迁移学习和微调工作流的基础。
#
# 随后,我们将演示一个典型工作流:先获得一个在 ImageNet 数据集上预训练的模型,然后在 Kaggle Dogs vs. Cats 分类数据集上对该模型进行重新训练。
#
# 此工作流改编自 [Python 深度学习](https://www.manning.com/books/deep-learning-with-python) [ 和 2016 年的博文“使用极少的数据构建强大的图像分类模型”](https://www.manning.com/books/deep-learning-with-python)
# + [markdown] id="fbf8630c325b"
# ## 冻结层:了解 trainable 特性
#
# 层和模型具有三个权重特性:
#
# - `weights` 是层的所有权重变量的列表。
# - `trainable_weights` 是需要进行更新(通过梯度下降)以尽可能减少训练过程中损失的权重列表。
# - `non_trainable_weights` 是不适合训练的权重列表。它们通常在正向传递过程中由模型更新。
#
# **示例:`Dense` 层具有 2 个可训练权重(内核与偏差)**
# + id="407deab1855e"
layer = keras.layers.Dense(3)
layer.build((None, 4)) # Create the weights
print("weights:", len(layer.weights))
print("trainable_weights:", len(layer.trainable_weights))
print("non_trainable_weights:", len(layer.non_trainable_weights))
# + [markdown] id="79fcb9cc960d"
# 一般而言,所有权重都是可训练权重。唯一具有不可训练权重的内置层是 `BatchNormalization` 层。在训练期间,它使用不可训练权重跟踪其输入的平均值和方差。要了解如何在您自己的自定义层中使用不可训练权重,请参阅从头开始编写新层的指南。
#
# **示例:`BatchNormalization` 层具有 2 个可训练权重和 2 个不可训练权重**
# + id="fbc87a09bc3c"
layer = keras.layers.BatchNormalization()
layer.build((None, 4)) # Create the weights
print("weights:", len(layer.weights))
print("trainable_weights:", len(layer.trainable_weights))
print("non_trainable_weights:", len(layer.non_trainable_weights))
# + [markdown] id="cddcdbf2bd5b"
# 层和模型还具有布尔特性 `trainable`。此特性的值可以更改。将 `layer.trainable` 设置为 `False` 会将层的所有权重从可训练移至不可训练。这一过程称为“冻结”层:已冻结层的状态在训练期间不会更新(无论是使用 `fit()` 进行训练,还是使用依赖于 `trainable_weights` 来应用梯度更新的任何自定义循环进行训练时)。
#
# **示例:将 `trainable` 设置为 `False`**
# + id="51bbc5d12742"
layer = keras.layers.Dense(3)
layer.build((None, 4)) # Create the weights
layer.trainable = False # Freeze the layer
print("weights:", len(layer.weights))
print("trainable_weights:", len(layer.trainable_weights))
print("non_trainable_weights:", len(layer.non_trainable_weights))
# + [markdown] id="32904f9a58db"
# 当可训练权重变为不可训练时,它的值在训练期间不再更新。
# + id="3c26c27a8291"
# Make a model with 2 layers
layer1 = keras.layers.Dense(3, activation="relu")
layer2 = keras.layers.Dense(3, activation="sigmoid")
model = keras.Sequential([keras.Input(shape=(3,)), layer1, layer2])
# Freeze the first layer
layer1.trainable = False
# Keep a copy of the weights of layer1 for later reference
initial_layer1_weights_values = layer1.get_weights()
# Train the model
model.compile(optimizer="adam", loss="mse")
model.fit(np.random.random((2, 3)), np.random.random((2, 3)))
# Check that the weights of layer1 have not changed during training
final_layer1_weights_values = layer1.get_weights()
np.testing.assert_allclose(
initial_layer1_weights_values[0], final_layer1_weights_values[0]
)
np.testing.assert_allclose(
initial_layer1_weights_values[1], final_layer1_weights_values[1]
)
# + [markdown] id="412d7d659aa1"
# 请勿将 `layer.trainable` 特性与 `layer.__call__()` 中的 `training` 参数(此参数控制层是在推断模式还是训练模式下运行其前向传递)混淆。有关更多信息,请参阅 [Keras常见问题解答](https://keras.io/getting_started/faq/#whats-the-difference-between-the-training-argument-in-call-and-the-trainable-attribute) 。
# + [markdown] id="e6ccd3c7ab1a"
# ## `trainable` 特性的递归设置
#
# 如果在模型或具有子层的任何层上设置 `trainable = False`,则所有子层也将变为不可训练。
#
# **示例:**
# + id="4235d0c69821"
inner_model = keras.Sequential(
[
keras.Input(shape=(3,)),
keras.layers.Dense(3, activation="relu"),
keras.layers.Dense(3, activation="relu"),
]
)
model = keras.Sequential(
[keras.Input(shape=(3,)), inner_model, keras.layers.Dense(3, activation="sigmoid"),]
)
model.trainable = False # Freeze the outer model
assert inner_model.trainable == False # All layers in `model` are now frozen
assert inner_model.layers[0].trainable == False # `trainable` is propagated recursively
# + [markdown] id="61535ba76727"
# ## 典型的迁移学习工作流
#
# 下面将介绍如何在 Keras 中实现典型的迁移学习工作流:
#
# 1. 实例化一个基础模型并加载预训练权重。
# 2. 通过设置 `trainable = False` 冻结基础模型中的所有层。
# 3. 根据基础模型中一个(或多个)层的输出创建一个新模型。
# 4. 在您的新数据集上训练新模型。
#
# 请注意,另一种更轻量的工作流如下:
#
# 1. 实例化一个基础模型并加载预训练权重。
# 2. 通过该模型运行新的数据集,并记录基础模型中一个(或多个)层的输出。这一过程称为特征提取。
# 3. 使用该输出作为新的较小模型的输入数据。
#
# 第二种工作流有一个关键优势,即您只需在自己的数据上运行一次基础模型,而不是每个训练周期都运行一次。因此,它的速度更快,开销也更低。
#
# 但是,第二种工作流存在一个问题,即它不允许您在训练期间动态修改新模型的输入数据,在进行数据扩充时,这种修改必不可少。当新数据集的数据太少而无法从头开始训练完整模型时,任务通常会使用迁移学习,在这种情况下,数据扩充非常重要。因此,在接下来的篇幅中,我们将专注于第一种工作流。
#
# 下面是 Keras 中第一种工作流的样子:
#
# 首先,实例化一个具有预训练权重的基础模型。
#
# ```python
# base_model = keras.applications.Xception(
# weights='imagenet', # Load weights pre-trained on ImageNet.
# input_shape=(150, 150, 3),
# include_top=False) # Do not include the ImageNet classifier at the top.
# ```
#
# 随后,冻结该基础模型。
#
# ```python
# base_model.trainable = False
# ```
#
# 根据基础模型创建一个新模型。
#
# ```python
# inputs = keras.Input(shape=(150, 150, 3))
# # We make sure that the base_model is running in inference mode here,
# # by passing `training=False`. This is important for fine-tuning, as you will
# # learn in a few paragraphs.
# x = base_model(inputs, training=False)
# # Convert features of shape `base_model.output_shape[1:]` to vectors
# x = keras.layers.GlobalAveragePooling2D()(x)
# # A Dense classifier with a single unit (binary classification)
# outputs = keras.layers.Dense(1)(x)
# model = keras.Model(inputs, outputs)
# ```
#
# 在新数据上训练该模型。
#
# ```python
# model.compile(optimizer=keras.optimizers.Adam(),
# loss=keras.losses.BinaryCrossentropy(from_logits=True),
# metrics=[keras.metrics.BinaryAccuracy()])
# model.fit(new_dataset, epochs=20, callbacks=..., validation_data=...)
# ```
# + [markdown] id="736c99aea690"
# ## 微调
#
# 一旦模型在新数据上收敛,您就可以尝试解冻全部或部分基础模型,并以极低的学习率端到端地重新训练整个模型。
#
# 这是可选的最后一个步骤,可能给您带来增量式改进。不过,它也可能导致快速过拟合,请牢记这一点。
#
# 重要的是,只有在将具有冻结层的模型训练至收敛后,才能执行此步骤。如果将随机初始化的可训练层与包含预训练特征的可训练层混合使用,则随机初始化的层将在训练过程中引起非常大的梯度更新,而这将破坏您的预训练特征。
#
# 在此阶段使用极低的学习率也很重要,因为与第一轮训练相比,您正在一个通常非常小的数据集上训练一个大得多的模型。因此,如果您应用较大的权重更新,则存在很快过拟合的风险。在这里,您只需要以增量方式重新调整预训练权重。
#
# 下面是实现整个基础模型微调的方法:
#
# ```python
# # Unfreeze the base model
# base_model.trainable = True
#
# # It's important to recompile your model after you make any changes
# # to the `trainable` attribute of any inner layer, so that your changes
# # are take into account
# model.compile(optimizer=keras.optimizers.Adam(1e-5), # Very low learning rate
# loss=keras.losses.BinaryCrossentropy(from_logits=True),
# metrics=[keras.metrics.BinaryAccuracy()])
#
# # Train end-to-end. Be careful to stop before you overfit!
# model.fit(new_dataset, epochs=10, callbacks=..., validation_data=...)
# ```
#
# **关于 `compile()` 和 `trainable` 的重要说明**
#
# 在模型上调用 `compile()` 意味着“冻结”该模型的行为。这意味着编译模型时的 `trainable` 特性值应当在该模型的整个生命周期中保留,直到再次调用 `compile`。因此,如果您更改任何 `trainable` 值,请确保在您的模型上再次调用 `compile()` 以将您的变更考虑在内。
#
# **关于 `BatchNormalization` 层的重要说明**
#
# 许多图像模型包含 `BatchNormalization` 层。该层在任何方面都是一个特例。下面是一些注意事项。
#
# - `BatchNormalization` 包含 2 个会在训练期间更新的不可训练权重。它们是跟踪输入的均值和方差的变量。
# - 当您设置 `bn_layer.trainable = False` 时,`BatchNormalization` 层将以推断模式运行,并且不会更新其均值和方差统计信息。一般而言,其他层的情况并非如此,因为[权重可训练性和推断/训练模式是两个正交概念](https://keras.io/getting_started/faq/#whats-the-difference-between-the-training-argument-in-call-and-the-trainable-attribute)。但是,对于 `BatchNormalization` 层,两者是关联的。
# - 当您解冻包含 `BatchNormalization` 层的模型以进行微调时,您应当通过在调用基础模型时传递 `training=False` 以将 `BatchNormalization` 层保持在推断模式。否则,应用于不可训练权重的更新会突然破坏模型已经学习的内容。
#
# 您将在本指南结尾处的端到端示例中看到这种模式的实际运行。
#
# + [markdown] id="bce9ffc4e290"
# ## 使用自定义训练循环进行迁移学习和微调
#
# 如果您使用自己的低级训练循环而不是 `fit()`,则工作流基本保持不变。在应用梯度更新时,您应当注意只考虑清单 `model.trainable_weights`:
#
# ```python
# # Create base model
# base_model = keras.applications.Xception(
# weights='imagenet',
# input_shape=(150, 150, 3),
# include_top=False)
# # Freeze base model
# base_model.trainable = False
#
# # Create new model on top.
# inputs = keras.Input(shape=(150, 150, 3))
# x = base_model(inputs, training=False)
# x = keras.layers.GlobalAveragePooling2D()(x)
# outputs = keras.layers.Dense(1)(x)
# model = keras.Model(inputs, outputs)
#
# loss_fn = keras.losses.BinaryCrossentropy(from_logits=True)
# optimizer = keras.optimizers.Adam()
#
# # Iterate over the batches of a dataset.
# for inputs, targets in new_dataset:
# # Open a GradientTape.
# with tf.GradientTape() as tape:
# # Forward pass.
# predictions = model(inputs)
# # Compute the loss value for this batch.
# loss_value = loss_fn(targets, predictions)
#
# # Get gradients of loss wrt the *trainable* weights.
# gradients = tape.gradient(loss_value, model.trainable_weights)
# # Update the weights of the model.
# optimizer.apply_gradients(zip(gradients, model.trainable_weights))
# ```
# + [markdown] id="4e63ba34ce1c"
# 对于微调同样如此。
# + [markdown] id="852447087ba9"
# ## 端到端示例:基于 Dogs vs. Cats 数据集微调图像分类模型
#
# 为了巩固这些概念,我们先介绍一个具体的端到端迁移学习和微调示例。我们将加载在 ImageNet 上预训练的 Xception 模型,并将其用于 Kaggle Dogs vs. Cats 分类数据集。
# + [markdown] id="ba75835e0de6"
# ### 获取数据
#
# 首先,我们使用 TFDS 来获取 Dogs vs. Cats 数据集。如果您拥有自己的数据集,则可能需要使用效用函数 `tf.keras.preprocessing.image_dataset_from_directory` 从磁盘上存档到类特定的文件夹中的一组图像来生成相似的有标签数据集对象。
#
# 使用非常小的数据集时,迁移学习最实用。为了使数据集保持较小状态,我们将原始训练数据(25,000 个图像)的 40% 用于训练,10% 用于验证,10% 用于测试。
# + id="1a99f56934f7"
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
train_ds, validation_ds, test_ds = tfds.load(
"cats_vs_dogs",
# Reserve 10% for validation and 10% for test
split=["train[:40%]", "train[40%:50%]", "train[50%:60%]"],
as_supervised=True, # Include labels
)
print("Number of training samples: %d" % tf.data.experimental.cardinality(train_ds))
print(
"Number of validation samples: %d" % tf.data.experimental.cardinality(validation_ds)
)
print("Number of test samples: %d" % tf.data.experimental.cardinality(test_ds))
# + [markdown] id="9db548603642"
# 下面是训练数据集中的前 9 个图像。如您所见,它们具有不同的大小。
# + id="00c8cbd1de88"
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 10))
for i, (image, label) in enumerate(train_ds.take(9)):
ax = plt.subplot(3, 3, i + 1)
plt.imshow(image)
plt.title(int(label))
plt.axis("off")
# + [markdown] id="168c4a10c072"
# 我们还可以看到标签 1 是“狗”,标签 0 是“猫”。
# + [markdown] id="f749203cd740"
# ### 标准化数据
#
# 我们的原始图像有各种大小。另外,每个像素由 0 到 255 之间的 3 个整数值(RGB 色阶值)组成。这不太适合馈送神经网络。我们需要做下面两件事:
#
# - 标准化为固定图像大小。我们选择 150x150。
# - 在 -1 至 1 之间归一化像素值。我们将使用 `Normalization` 层作为模型本身的一部分来进行此操作。
#
# 一般而言,与采用已预处理数据的模型相反,开发以原始数据作为输入的模型是一种良好的做法。原因在于,如果模型需要预处理的数据,则每次导出模型以在其他地方(在网络浏览器、移动应用中)使用时,都需要重新实现完全相同的预处理流水线。这很快就会变得非常棘手。因此,在命中模型之前,我们应当尽可能少地进行预处理。
#
# 在这里,我们将在数据流水线中进行图像大小调整(因为深度神经网络只能处理连续的数据批次),并在创建模型时将其作为模型的一部分进行输入值缩放。
#
# 我们将图像的大小调整为 150x150:
# + id="b3678f38e087"
size = (150, 150)
train_ds = train_ds.map(lambda x, y: (tf.image.resize(x, size), y))
validation_ds = validation_ds.map(lambda x, y: (tf.image.resize(x, size), y))
test_ds = test_ds.map(lambda x, y: (tf.image.resize(x, size), y))
# + [markdown] id="708bf9792a35"
# 此外,我们对数据进行批处理并使用缓存和预提取来优化加载速度。
# + id="53ef9e6092e3"
batch_size = 32
train_ds = train_ds.cache().batch(batch_size).prefetch(buffer_size=10)
validation_ds = validation_ds.cache().batch(batch_size).prefetch(buffer_size=10)
test_ds = test_ds.cache().batch(batch_size).prefetch(buffer_size=10)
# + [markdown] id="b60f852c462f"
# ### 使用随机数据扩充
#
# 当您没有较大的图像数据集时,通过将随机但现实的转换(例如随机水平翻转或小幅随机旋转)应用于训练图像来人为引入样本多样性是一种良好的做法。这有助于使模型暴露于训练数据的不同方面,同时减慢过拟合的速度。
# + id="40b1e355b9c0"
from tensorflow import keras
from tensorflow.keras import layers
data_augmentation = keras.Sequential(
[
layers.experimental.preprocessing.RandomFlip("horizontal"),
layers.experimental.preprocessing.RandomRotation(0.1),
]
)
# + [markdown] id="6fa8ddeda36e"
# 我们看一下经过各种随机转换后第一个批次的第一个图像是什么样:
# + id="9077f9fd022e"
import numpy as np
for images, labels in train_ds.take(1):
plt.figure(figsize=(10, 10))
first_image = images[0]
for i in range(9):
ax = plt.subplot(3, 3, i + 1)
augmented_image = data_augmentation(
tf.expand_dims(first_image, 0), training=True
)
plt.imshow(augmented_image[0].numpy().astype("int32"))
plt.title(int(labels[0]))
plt.axis("off")
# + [markdown] id="bc999e4672c3"
# ## 构建模型
#
# 现在,我们来构建一个遵循我们先前解释的蓝图的模型。
#
# 注意:
#
# - 我们添加 `Normalization` 层以将输入值(最初在 `[0, 255]` 范围内)缩放到 `[-1, 1]` 范围。
# - 我们在分类层之前添加一个 `Dropout` 层,以进行正则化。
# - 我们确保在调用基础模型时传递 `training=False`,使其在推断模式下运行,这样,即使在我们解冻基础模型以进行微调后,batchnorm 统计信息也不会更新。
# + id="07a2f9e9d817"
base_model = keras.applications.Xception(
weights="imagenet", # Load weights pre-trained on ImageNet.
input_shape=(150, 150, 3),
include_top=False,
) # Do not include the ImageNet classifier at the top.
# Freeze the base_model
base_model.trainable = False
# Create new model on top
inputs = keras.Input(shape=(150, 150, 3))
x = data_augmentation(inputs) # Apply random data augmentation
# Pre-trained Xception weights requires that input be normalized
# from (0, 255) to a range (-1., +1.), the normalization layer
# does the following, outputs = (inputs - mean) / sqrt(var)
norm_layer = keras.layers.experimental.preprocessing.Normalization()
mean = np.array([127.5] * 3)
var = mean ** 2
# Scale inputs to [-1, +1]
x = norm_layer(x)
norm_layer.set_weights([mean, var])
# The base model contains batchnorm layers. We want to keep them in inference mode
# when we unfreeze the base model for fine-tuning, so we make sure that the
# base_model is running in inference mode here.
x = base_model(x, training=False)
x = keras.layers.GlobalAveragePooling2D()(x)
x = keras.layers.Dropout(0.2)(x) # Regularize with dropout
outputs = keras.layers.Dense(1)(x)
model = keras.Model(inputs, outputs)
model.summary()
# + [markdown] id="2e8237de81e8"
# ## 训练顶层
# + id="9137b8daedad"
model.compile(
optimizer=keras.optimizers.Adam(),
loss=keras.losses.BinaryCrossentropy(from_logits=True),
metrics=[keras.metrics.BinaryAccuracy()],
)
epochs = 20
model.fit(train_ds, epochs=epochs, validation_data=validation_ds)
# + [markdown] id="aa51d4562fa7"
# ## 对整个模型进行一轮微调
#
# 最后,我们解冻基础模型,并以较低的学习率端到端地训练整个模型。
#
# 重要的是,尽管基础模型变得可训练,但在构建模型过程中,由于我们在调用该模型时传递了 `training=False`,因此它仍在推断模式下运行。这意味着内部的批次归一化层不会更新其批次统计信息。如果它们更新了这些统计信息,则会破坏该模型到目前为止所学习的表示。
# + id="3cc299505b72"
# Unfreeze the base_model. Note that it keeps running in inference mode
# since we passed `training=False` when calling it. This means that
# the batchnorm layers will not update their batch statistics.
# This prevents the batchnorm layers from undoing all the training
# we've done so far.
base_model.trainable = True
model.summary()
model.compile(
optimizer=keras.optimizers.Adam(1e-5), # Low learning rate
loss=keras.losses.BinaryCrossentropy(from_logits=True),
metrics=[keras.metrics.BinaryAccuracy()],
)
epochs = 10
model.fit(train_ds, epochs=epochs, validation_data=validation_ds)
# + [markdown] id="afa73d989302"
# 经过 10 个周期后,微调在这里为我们提供了出色的改进。
|
site/zh-cn/guide/keras/transfer_learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + tags=["hide-input"]
import panel as pn
import pandas as pd
import holoviews as hv
from sklearn.cluster import KMeans
penguins = pd.read_csv('https://raw.githubusercontent.com/mwaskom/seaborn-data/master/penguins.csv')
cols = list(penguins.columns)[:-1]
pn.extension('ace')
import hvplot.pandas
# + tags=["hide-input"]
slider = pn.widgets.IntSlider(start=0, end=10)
img = pn.pane.JPG(f"https://picsum.photos/800/300?image=0", embed=False, height=300)
slider.jscallback(args={'img': img}, value="""
img.text = '<img src="https://picsum.photos/800/300?image='+cb_obj.value+'" width=800 height=300></img>'
""")
app = pn.Column(slider, img)
ace = pn.widgets.Ace(readonly=True, width=800, height=200, language='python', theme='monokai', value=\
"""slider = pn.widgets.IntSlider(start=0, end=10)
def slideshow(index):
url = f"https://picsum.photos/800/300?image={index}"
return pn.pane.JPG(url)
output = pn.bind(slideshow, slider)
app = pn.Column(slider, output)""")
app1 = pn.Tabs(('Output', app), ('Code', ace))
penguins = pd.read_csv('https://raw.githubusercontent.com/mwaskom/seaborn-data/master/penguins.csv').dropna()
cols = list(penguins.columns)[2:6]
x = pn.widgets.Select(name='x', options=cols)
y = pn.widgets.Select(name='y', options=cols, value='bill_depth_mm')
n_clusters = pn.widgets.IntSlider(name='n_clusters', start=2, end=5, value=3)
def cluster(data, n_clusters):
kmeans = KMeans(n_clusters=n_clusters)
est = kmeans.fit(data)
return est.labels_.astype('str')
def plot(x, y, n_clusters):
penguins['labels'] = cluster(penguins.iloc[:, 2:6].values, n_clusters)
centers = penguins.groupby('labels').mean()
return (penguins.sort_values('labels').hvplot.scatter(
x, y, c='labels', hover_cols=['species'], line_width=1, size=60, frame_width=400, frame_height=400
).opts(marker=hv.dim('species').categorize({'Adelie': 'square', 'Chinstrap': 'circle', 'Gentoo': 'triangle'})) * centers.hvplot.scatter(
x, y, marker='x', color='black', size=400, padding=0.1, line_width=5
))
explanation = pn.pane.Markdown("""
This app applies k-means clustering on the Palmer Penguins dataset using scikit-learn, parameterizing the number of clusters and the variables to plot.
<br><br>
Each cluster is denoted by one color while the penguin species is indicated using markers:
<br><br>
● - Adelie, ▲ - Gentoo, ■ - Chinstrap
<br><br>
By comparing the two we can assess the performance of the clustering algorithm.
<br><br>
""")
code = pn.widgets.Ace(language='python', theme='monokai', height=360, value=\
"""x = pn.widgets.Select(name='x', options=cols)
y = pn.widgets.Select(name='y', options=cols, value='bill_depth_mm')
n_clusters = pn.widgets.IntSlider(name='n_clusters', start=2, end=5, value=3)
explanation = pn.pane.Markdown(...)
def plot_clusters(x, y, n_clusters):
...
pn.Row(
pn.WidgetBox(x, y, n_clusters, explanation),
pn.bind(plot, x, y, n_clusters)
)""", width=800)
app2 = pn.Tabs(
('Output', pn.Column(
pn.Row(
pn.WidgetBox(x, y, n_clusters, explanation),
pn.bind(plot, x, y, n_clusters)
)
)),
('Code', code)
)
pn.Row(pn.layout.HSpacer(), pn.Tabs(
('Penguin K-Means Clustering', app2),
('Slideshow', app1)
), pn.layout.HSpacer(), sizing_mode='stretch_width').embed(max_opts=4, json=True, json_prefix='json')
|
examples/homepage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# See requirements.txt to set up your dev environment.
import os
import cv2
import sys
import json
import scipy
import urllib
import datetime
import urllib3
import rasterio
import subprocess
import numpy as np
import pandas as pd
import seaborn as sns
from osgeo import gdal, ogr, osr
from planet import api
from planet.api import filters
from traitlets import link
import rasterio.tools.mask as rio_mask
from shapely.geometry import mapping, shape
from IPython.display import display, Image, HTML
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
#from scipy import ndimage
urllib3.disable_warnings()
from ipyleaflet import (
Map,
Marker,
TileLayer, ImageOverlay,
Polyline, Polygon, Rectangle, Circle, CircleMarker,
GeoJSON,
DrawControl
)
# %matplotlib inline
# will pick up api_key via environment variable PL_API_KEY
# but can be specified using `api_key` named argument
api_keys = json.load(open("apikeys.json",'r'))
client = api.ClientV1(api_key=api_keys["PLANET_API_KEY"])
# -
# # Let's pull it all together to do something cool.
# * Let's reuse a lot of our code to make a movie of our travel around Portland.
# * We'll first select a bunch of recent scenes, activate, and download them.
# * After that we'll create a mosaic, a path, and trace the path through the moasic.
# * We'll use the path to crop subregions, save them as images, and create a video.
# * First step is to trace our AOI and a path through it.
# +
# Basemap Mosaic (v1 API)
mosaicsSeries = 'global_quarterly_2017q1_mosaic'
# Planet tile server base URL (Planet Explorer Mosaics Tiles)
mosaicsTilesURL_base = 'https://tiles0.planet.com/experimental/mosaics/planet-tiles/' + mosaicsSeries + '/gmap/{z}/{x}/{y}.png'
# Planet tile server url
mosaicsTilesURL = mosaicsTilesURL_base + '?api_key=' + api_keys["PLANET_API_KEY"]
# Map Settings
# Define colors
colors = {'blue': "#009da5"}
# Define initial map center lat/long
center = [45.5231, -122.6765]
# Define initial map zoom level
zoom = 11
# Set Map Tiles URL
planetMapTiles = TileLayer(url= mosaicsTilesURL)
# Create the map
m = Map(
center=center,
zoom=zoom,
default_tiles = planetMapTiles # Uncomment to use Planet.com basemap
)
# Define the draw tool type options
polygon = {'shapeOptions': {'color': colors['blue']}}
rectangle = {'shapeOptions': {'color': colors['blue']}}
# Create the draw controls
# @see https://github.com/ellisonbg/ipyleaflet/blob/master/ipyleaflet/leaflet.py#L293
dc = DrawControl(
polygon = polygon,
rectangle = rectangle
)
# Initialize an action counter variable
actionCount = 0
AOIs = {}
# Register the draw controls handler
def handle_draw(self, action, geo_json):
# Increment the action counter
global actionCount
actionCount += 1
# Remove the `style` property from the GeoJSON
geo_json['properties'] = {}
# Convert geo_json output to a string and prettify (indent & replace ' with ")
geojsonStr = json.dumps(geo_json, indent=2).replace("'", '"')
AOIs[actionCount] = json.loads(geojsonStr)
# Attach the draw handler to the draw controls `on_draw` event
dc.on_draw(handle_draw)
m.add_control(dc)
m
# -
# # Query the API
# * Now we'll save the geometry for our AOI and the path.
# * We'll also filter and cleanup our data just like before.
# +
areaAOI = AOIs[1]["geometry"]
pathAOI = AOIs[2]["geometry"]
aoi_file ="portland.geojson"
with open(aoi_file,"w") as f:
f.write(json.dumps(areaAOI))
# build a query using the AOI and
# a cloud_cover filter that excludes 'cloud free' scenes
old = datetime.datetime(year=2017,month=1,day=1)
query = filters.and_filter(
filters.geom_filter(areaAOI),
filters.range_filter('cloud_cover', lt=5),
filters.date_range('acquired', gt=old)
)
# build a request for only PlanetScope imagery
request = filters.build_search_request(
query, item_types=['PSScene3Band']
)
# if you don't have an API key configured, this will raise an exception
result = client.quick_search(request)
scenes = []
planet_map = {}
for item in result.items_iter(limit=500):
planet_map[item['id']]=item
props = item['properties']
props["id"] = item['id']
props["geometry"] = item["geometry"]
props["thumbnail"] = item["_links"]["thumbnail"]
scenes.append(props)
scenes = pd.DataFrame(data=scenes)
display(scenes)
print len(scenes)
# -
# # Just like before we clean up our data and distill it down to just the scenes we want.
# +
# now let's clean up the datetime stuff
# make a shapely shape from our aoi
portland = shape(areaAOI)
footprints = []
overlaps = []
# go through the geometry from our api call, convert to a shape and calculate overlap area.
# also save the shape for safe keeping
for footprint in scenes["geometry"].tolist():
s = shape(footprint)
footprints.append(s)
overlap = 100.0*(portland.intersection(s).area / portland.area)
overlaps.append(overlap)
# take our lists and add them back to our dataframe
scenes['overlap'] = pd.Series(overlaps, index=scenes.index)
scenes['footprint'] = pd.Series(footprints, index=scenes.index)
# now make sure pandas knows about our date/time columns.
scenes["acquired"] = pd.to_datetime(scenes["acquired"])
scenes["published"] = pd.to_datetime(scenes["published"])
scenes["updated"] = pd.to_datetime(scenes["updated"])
scenes.head()
# Now let's get it down to just good, recent, clear scenes
clear = scenes['cloud_cover']<0.4
good = scenes['quality_category']=="standard"
recent = scenes["acquired"] > datetime.date(year=2017,month=1,day=1)
partial_coverage = scenes["overlap"] > 10
good_scenes = scenes[(good&clear&recent&partial_coverage)]
print good_scenes
# -
# # To make sure we are good we'll visually inspect the scenes in our slippy map.
# first create a list of colors
colors = ["#ff0000","#00ff00","#0000ff","#ffff00","#ff00ff","#00ffff","#ff0000","#00ff00","#0000ff","#ffff00","#ff00ff","#00ffff"]
# grab our scenes from the geometry/footprint geojson
# Chane this number as needed
footprints = good_scenes[0:10]["geometry"].tolist()
# for each footprint/color combo
for footprint,color in zip(footprints,colors):
# create the leaflet object
feat = {'geometry':footprint,"properties":{
'style':{'color': color,'fillColor': color,'fillOpacity': 0.2,'weight': 1}},
'type':u"Feature"}
# convert to geojson
gjson = GeoJSON(data=feat)
# add it our map
m.add_layer(gjson)
# now we will draw our original AOI on top
feat = {'geometry':areaAOI,"properties":{
'style':{'color': "#FFFFFF",'fillColor': "#FFFFFF",'fillOpacity': 0.5,'weight': 1}},
'type':u"Feature"}
gjson = GeoJSON(data=feat)
m.add_layer(gjson)
m
# # This is from the previous notebook. We are just activating and downloading scenes.
# +
def get_products(client, scene_id, asset_type='PSScene3Band'):
"""
Ask the client to return the available products for a
given scene and asset type. Returns a list of product
strings
"""
out = client.get_assets_by_id(asset_type,scene_id)
temp = out.get()
return temp.keys()
def activate_product(client, scene_id, asset_type="PSScene3Band",product="analytic"):
"""
Activate a product given a scene, an asset type, and a product.
On success return the return value of the API call and an activation object
"""
temp = client.get_assets_by_id(asset_type,scene_id)
products = temp.get()
if( product in products.keys() ):
return client.activate(products[product]),products[product]
else:
return None
def download_and_save(client,product):
"""
Given a client and a product activation object download the asset.
This will save the tiff file in the local directory and return its
file name.
"""
out = client.download(product)
fp = out.get_body()
fp.write()
return fp.name
def scenes_are_active(scene_list):
"""
Check if all of the resources in a given list of
scene activation objects is read for downloading.
"""
return True
retVal = True
for scene in scene_list:
if scene["status"] != "active":
print "{} is not ready.".format(scene)
return False
return True
def load_image4(filename):
"""Return a 4D (r, g, b, nir) numpy array with the data in the specified TIFF filename."""
path = os.path.abspath(os.path.join('./', filename))
if os.path.exists(path):
with rasterio.open(path) as src:
b, g, r, nir = src.read()
return np.dstack([r, g, b, nir])
def load_image3(filename):
"""Return a 3D (r, g, b) numpy array with the data in the specified TIFF filename."""
path = os.path.abspath(os.path.join('./', filename))
if os.path.exists(path):
with rasterio.open(path) as src:
b,g,r,mask = src.read()
return np.dstack([b, g, r])
def get_mask(filename):
"""Return a 1D mask numpy array with the data in the specified TIFF filename."""
path = os.path.abspath(os.path.join('./', filename))
if os.path.exists(path):
with rasterio.open(path) as src:
b,g,r,mask = src.read()
return np.dstack([mask])
def rgbir_to_rgb(img_4band):
"""Convert an RGBIR image to RGB"""
return img_4band[:,:,:3]
# -
# # Perform the actual activation ... go get coffee
to_get = good_scenes["id"][0:10].tolist()
to_get = sorted(to_get)
activated = []
# for each scene to get
for scene in to_get:
# get the product
product_types = get_products(client,scene)
for p in product_types:
# if there is a visual productfor p in labels:
if p == "visual": # p == "basic_analytic_dn"
print "Activating {0} for scene {1}".format(p,scene)
# activate the product
_,product = activate_product(client,scene,product=p)
activated.append(product)
# # Downloand the scenes
# +
tiff_files = []
asset_type = "_3B_Visual"
# check if our scenes have been activated
if scenes_are_active(activated):
for to_download,name in zip(activated,to_get):
# create the product name
name = name + asset_type + ".tif"
# if the product exists locally
if( os.path.isfile(name) ):
# do nothing
print "We have scene {0} already, skipping...".format(name)
tiff_files.append(name)
elif to_download["status"] == "active":
# otherwise download the product
print "Downloading {0}....".format(name)
fname = download_and_save(client,to_download)
tiff_files.append(fname)
print "Download done."
else:
print "Could not download, still activating"
else:
print "Scenes aren't ready yet"
print tiff_files
# -
# # Now, just like before, we will mosaic those scenes.
# * It is easier to call out using subprocess and use the command line util.
# * Just iterate through the files and drop them into a single file portland_mosaic.tif
subprocess.call(["rm","portland_mosaic.tif"])
commands = ["gdalwarp", # t
"-t_srs","EPSG:3857",
"-cutline",aoi_file,
"-crop_to_cutline",
"-tap",
"-tr", "3", "3"
"-overwrite"]
output_mosaic = "portland_mosaic.tif"
for tiff in tiff_files:
commands.append(tiff)
commands.append(output_mosaic)
print " ".join(commands)
subprocess.call(commands)
# # Let's take a look at what we got
merged = load_image3(output_mosaic)
plt.figure(0,figsize=(18,18))
plt.imshow(merged)
plt.title("merged")
# # Now we are going to write a quick crop function.
# * this function takes in a, scene, a center position, and the width and height of a window.
# * We'll use numpy slice notation to make the crop.
# * Let's pick a spot and see what we get.
def crop_to_area(scene,x_c,y_c,w,h):
tlx = x_c-(w/2)
tly = y_c-(h/2)
brx = x_c+(w/2)
bry = y_c+(h/2)
return scene[tly:bry,tlx:brx,:]
#
plt.figure(0,figsize=(3,4))
plt.imshow(crop_to_area(merged,3000,3000,640,480))
plt.title("merged")
#
# # Now to figure out how our lat/long values map to pixels.
# * The next thing we need is a way to map from a lat and long in our slippy map to the pixel position in our image.
# * We'll use what we know about the lat/long of the corners of our image to do that.
# * We'll ask GDAL to tell us the extents of our scene and the geotransofrm.
# * We'll then apply the GeoTransform from GDAL to the coordinates that are the extents of our scene.
# * Now we have the corners of our scene in Lat/Long
# +
# Liberally borrowed from this example
# https://gis.stackexchange.com/questions/57834/how-to-get-raster-corner-coordinates-using-python-gdal-bindings
def GetExtent(gt,cols,rows):
"""
Get the list of corners in our output image in the format
[[x,y],[x,y],[x,y]]
"""
ext=[]
# for the corners of the image
xarr=[0,cols]
yarr=[0,rows]
for px in xarr:
for py in yarr:
# apply the geo coordiante transform
# using the affine transform we got from GDAL
x=gt[0]+(px*gt[1])+(py*gt[2])
y=gt[3]+(px*gt[4])+(py*gt[5])
ext.append([x,y])
yarr.reverse()
return ext
def ReprojectCoords(coords,src_srs,tgt_srs):
trans_coords=[]
# create a transform object from the source and target ref system
transform = osr.CoordinateTransformation( src_srs, tgt_srs)
for x,y in coords:
# transform the points
x,y,z = transform.TransformPoint(x,y)
# add it to the list.
trans_coords.append([x,y])
return trans_coords
# -
# # Here we'll call the functions we wrote.
# * First we open the scene and get the width and height.
# * Then from the geotransorm we'll reproject those points to lat and long.
# TLDR: pixels => UTM coordiantes => Lat Long
raster=output_mosaic
# Load the GDAL File
ds=gdal.Open(raster)
# get the geotransform
gt=ds.GetGeoTransform()
# get the width and height of our image
cols = ds.RasterXSize
rows = ds.RasterYSize
# Generate the coordinates of our image in utm
ext=GetExtent(gt,cols,rows)
# get the spatial referencec object
src_srs=osr.SpatialReference()
# get the data that will allow us to move from UTM to Lat Lon.
src_srs.ImportFromWkt(ds.GetProjection())
tgt_srs = src_srs.CloneGeogCS()
extents = ReprojectCoords(ext,src_srs,tgt_srs)
print extents
# # Now we'll do a bit of hack.
# * That bit above is precise but complext, we are going to make everything easier to think about.
# * We are going to linearize our scene, which isn't perfect, but good enough for our application.
# * What this function does is take in a given lat,long, the size of the image, and the extents as lat,lon coordinates.
# * For a given pixel we map it's x and y values to the value between a given lat and long and return the results.
# * Now we can ask, for a given lat,long pair what is the corresponding pixel.
def poor_mans_lat_lon_2_pix(lon,lat,w,h,extents):
# split up our lat and longs
lats = [e[1] for e in extents]
lons = [e[0] for e in extents]
# calculate our scene extents max and min
lat_max = np.max(lats)
lat_min = np.min(lats)
lon_max = np.max(lons)
lon_min = np.min(lons)
# calculate the difference between our start point
# and our minimum
lat_diff = lat-lat_min
lon_diff = lon-lon_min
# create the linearization
lat_r = float(h)/(lat_max-lat_min)
lon_r = float(w)/(lon_max-lon_min)
# generate the results.
return int(lat_r*lat_diff),int(lon_r*lon_diff)
# # Let's check our work
# * First we'll create a draw point function that just puts a red dot at given pixel.
# * We'll get our scene, and map all of the lat/long points in our path to pixel values.
# * Finally we'll load our image, plot the points and show our results
def draw_point(x,y,img,t=40):
h,w,d = img.shape
y = h-y
img[(y-t):(y+t),(x-t):(x+t),:] = [255,0,0]
h,w,c = merged.shape
waypoints = [poor_mans_lat_lon_2_pix(point[0],point[1],w,h,extents) for point in pathAOI["coordinates"]]
print waypoints
merged = load_image3(output_mosaic)
[draw_point(pt[1],pt[0],merged) for pt in waypoints]
plt.figure(0,figsize=(18,18))
plt.imshow(merged)
plt.title("merged")
# # Now things get interesting....
# * Our path is just a few waypoint but to make a video we need just about every point between our waypoints.
# * To get all of the points between our waypoints we'll have to write a little interpolation script.
# * Interpolation is just a fancy word for nicely space points bewteen or waypoints, we'll call the space between each point as our "velocity."
# * If we were really slick we could define a heading vector and and build a spline so the camera faces the direction of heading. Our approach is fine as the top of the frame is always North, which makes reckoning easy.
# * Once we have our interpolation function all we need to do is to crop our large mosaic at each point in our interpolation point list and save it in a sequential file.
# +
def interpolate_waypoints(waypoints,velocity=10.0):
retVal = []
last_pt = waypoints[0]
# for each point in our waypoints except the first
for next_pt in waypoints[1:]:
# calculate distance between the points
distance = np.sqrt((last_pt[0]-next_pt[0])**2+(last_pt[1]-next_pt[1])**2)
# use our velocity to calculate the number steps.
steps = np.ceil(distance/velocity)
# linearly space points between the two points on our line
xs = np.array(np.linspace(last_pt[0],next_pt[0],steps),dtype='int64')
ys = np.array(np.linspace(last_pt[1],next_pt[1],steps),dtype='int64')
# zip the points together
retVal += zip(xs,ys)
# move to the next point
last_pt = next_pt
return retVal
def build_scenes(src,waypoints,window=[640,480],path="./movie/"):
count = 0
# Use opencv to change the color space of our image.
src = cv2.cvtColor(src, cv2.COLOR_BGR2RGB)
# define half our sampling window.
w2 = window[0]/2
h2 = window[1]/2
# for our source image get the width and height
h,w,d = src.shape
for pt in waypoints:
# for each point crop the area out.
# the y value of our scene is upside down.
temp = crop_to_area(src,pt[1],h-pt[0],window[0],window[1])
# If we happen to hit the border of the scene, just skip
if temp.shape[0]*temp.shape[1]== 0:
# if we have an issue, just keep plugging along
continue
# Resample the image a bit, this just makes things look nice.
temp = cv2.resize(temp, (int(window[0]*0.75), int(window[1]*.75)))
# create a file name
fname = os.path.abspath(path+"img{num:06d}.png".format(num=count))
# Save it
cv2.imwrite(fname,temp)
count += 1
# -
# # Before we generate our video frames, let's check our work
# * We'll load our image.
# * Build the interpolated waypoints list.
# * Draw the points on the image using our draw_point method.
# * Plot the results
# load the image
merged = load_image3(output_mosaic)
# interpolate the waypoints
interp = interpolate_waypoints(waypoints)
# draw them on our scene
[draw_point(pt[1],pt[0],merged) for pt in interp]
# display the scene
plt.figure(0,figsize=(18,18))
plt.imshow(merged)
plt.title("merged")
# # Now let's re-load the image and run the scene maker.
os.system("rm ./movie/*.png")
merged = load_image3(output_mosaic)
build_scenes(merged,interp)
# # Finally, let's make a movie.
# * Our friend AVConv, which is like ffmpeg is a handy command line util for transcoding video.
# * AVConv can also convert a series of images into a video and vice versa.
# * We'll set up our command and use subprocess to make the call.
# avconv -framerate 30 -f image2 -i ./movie/img%06d.png -b 65536k out.mpg
os.system("rm ./movie/*.png")
framerate = 30
output = "out.mpg"
command = ["avconv","-framerate", str(framerate), "-f", "image2", "-i", "./movie/img%06d.png", "-b", "65536k", output]
os.system(" ".join(command))
|
MovieTime.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Factoring Polynomials with SymPy
# Here is an example that uses [SymPy](http://sympy.org/en/index.html) to factor polynomials.
from ipywidgets import interact
from sympy import Symbol, Eq, factor
x = Symbol('x')
def factorit(n):
return Eq(x**n-1, factor(x**n-1))
factorit(12)
interact(factorit, n=(2,40));
|
docs/source/examples/Factoring.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ZxbUYitr0LJp"
# # CycleGAN, Image-to-Image Translation
#
# In this notebook, we're going to define and train a CycleGAN to read in an image from a set $X$ and transform it so that it looks as if it belongs in set $Y$. Specifically, we'll look at a set of images of [Yosemite national park](https://en.wikipedia.org/wiki/Yosemite_National_Park) taken either during the summer of winter. The seasons are our two domains!
#
# >The objective will be to train generators that learn to transform an image from domain $X$ into an image that looks like it came from domain $Y$ (and vice versa).
#
# Some examples of image data in both sets are pictured below.
#
# <img src='notebook_images/XY_season_images.png' width=50% />
#
# ### Unpaired Training Data
#
# These images do not come with labels, but CycleGANs give us a way to learn the mapping between one image domain and another using an **unsupervised** approach. A CycleGAN is designed for image-to-image translation and it learns from unpaired training data. This means that in order to train a generator to translate images from domain $X$ to domain $Y$, we do not have to have exact correspondences between individual images in those domains. For example, in [the paper that introduced CycleGANs](https://arxiv.org/abs/1703.10593), the authors are able to translate between images of horses and zebras, even though there are no images of a zebra in exactly the same position as a horse or with exactly the same background, etc. Thus, CycleGANs enable learning a mapping from one domain $X$ to another domain $Y$ without having to find perfectly-matched, training pairs!
#
# <img src='notebook_images/horse2zebra.jpg' width=50% />
#
# ### CycleGAN and Notebook Structure
#
# A CycleGAN is made of two types of networks: **discriminators, and generators**. In this example, the discriminators are responsible for classifying images as real or fake (for both $X$ and $Y$ kinds of images). The generators are responsible for generating convincing, fake images for both kinds of images.
#
# This notebook will detail the steps you should take to define and train such a CycleGAN.
#
# >1. You'll load in the image data using PyTorch's DataLoader class to efficiently read in images from a specified directory.
# 2. Then, you'll be tasked with defining the CycleGAN architecture according to provided specifications. You'll define the discriminator and the generator models.
# 3. You'll complete the training cycle by calculating the adversarial and cycle consistency losses for the generator and discriminator network and completing a number of training epochs. *It's suggested that you enable GPU usage for training.*
# 4. Finally, you'll evaluate your model by looking at the loss over time and looking at sample, generated images.
#
# + id="JBGm6nDO67cJ"
xy_season_images = plt.imread('XY_season_images.png')
plt.imshow(xy_season_images)
# + [markdown] id="T27mrQJ90LJu"
# ---
#
# ## Load and Visualize the Data
#
# We'll first load in and visualize the training data, importing the necessary libraries to do so.
#
# > If you are working locally, you'll need to download the data as a zip file by [clicking here](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/November/5be66e78_summer2winter-yosemite/summer2winter-yosemite.zip).
#
# It may be named `summer2winter-yosemite/` with a dash or an underscore, so take note, extract the data to your home directory and make sure the below `image_dir` matches. Then you can proceed with the following loading code.
# + id="Z7K5WAxG0LJw"
# loading in and transforming data
import os
import torch
from torch.utils.data import DataLoader
import torchvision
import torchvision.datasets as datasets
import torchvision.transforms as transforms
# visualizing data
import matplotlib.pyplot as plt
import numpy as np
import warnings
# %matplotlib inline
# + id="SCkWAkJH8VLV"
from google.colab import drive
drive.mount('/content/google')
import zipfile
zf = zipfile.ZipFile('/content/google/MyDrive/Data/summer2winter_yosemite.zip')
zf.extractall()
# + [markdown] id="ZYdYFACk0LJx"
# ### DataLoaders
#
# The `get_data_loader` function returns training and test DataLoaders that can load data efficiently and in specified batches. The function has the following parameters:
# * `image_type`: `summer` or `winter`, the names of the directories where the X and Y images are stored
# * `image_dir`: name of the main image directory, which holds all training and test images
# * `image_size`: resized, square image dimension (all images will be resized to this dim)
# * `batch_size`: number of images in one batch of data
#
# The test data is strictly for feeding to our generators, later on, so we can visualize some generated samples on fixed, test data.
#
# You can see that this function is also responsible for making sure our images are of the right, square size (128x128x3) and converted into Tensor image types.
#
# **It's suggested that you use the default values of these parameters.**
#
# Note: If you are trying this code on a different set of data, you may get better results with larger `image_size` and `batch_size` parameters. If you change the `batch_size`, make sure that you create complete batches in the training loop otherwise you may get an error when trying to save sample data.
# + id="XmC1KxPd0LJy"
def get_data_loader(image_type, image_dir='summer2winter_yosemite',
image_size=128, batch_size=16, num_workers=0):
"""Returns training and test data loaders for a given image type, either 'summer' or 'winter'.
These images will be resized to 128x128x3, by default, converted into Tensors, and normalized.
"""
# resize and normalize the images
transform = transforms.Compose([transforms.Resize(image_size), # resize to 128x128
transforms.ToTensor()])
# get training and test directories
image_path = './' + image_dir
train_path = os.path.join(image_path, image_type)
test_path = os.path.join(image_path, 'test_{}'.format(image_type))
# define datasets using ImageFolder
train_dataset = datasets.ImageFolder(train_path, transform)
test_dataset = datasets.ImageFolder(test_path, transform)
# create and return DataLoaders
train_loader = DataLoader(dataset=train_dataset, batch_size=batch_size, shuffle=True, num_workers=num_workers)
test_loader = DataLoader(dataset=test_dataset, batch_size=batch_size, shuffle=False, num_workers=num_workers)
return train_loader, test_loader
# + id="94e0fNgS0LJ0"
# Create train and test dataloaders for images from the two domains X and Y
# image_type = directory names for our data
dataloader_X, test_dataloader_X = get_data_loader(image_type='summer')
dataloader_Y, test_dataloader_Y = get_data_loader(image_type='winter')
# + [markdown] id="Ft8Cy87G0LJ1"
# ## Display some Training Images
#
# Below we provide a function `imshow` that reshape some given images and converts them to NumPy images so that they can be displayed by `plt`. This cell should display a grid that contains a batch of image data from set $X$.
# + id="h137EHvV0LJ2"
# helper imshow function
def imshow(img):
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
# get some images from X
dataiter = iter(dataloader_X)
# the "_" is a placeholder for no labels
images, _ = dataiter.next()
# show images
fig = plt.figure(figsize=(18, 10))
imshow(torchvision.utils.make_grid(images))
# + [markdown] id="JMU264eW0LJ3"
# Next, let's visualize a batch of images from set $Y$.
# + id="Xm9NPI8G0LJ5"
# get some images from Y
dataiter = iter(dataloader_Y)
images, _ = dataiter.next()
# show images
fig = plt.figure(figsize=(18,10))
imshow(torchvision.utils.make_grid(images))
# + [markdown] id="tFfh5f6h0LJ5"
# ### Pre-processing: scaling from -1 to 1
#
# We need to do a bit of pre-processing; we know that the output of our `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)
# + id="KY-WuSVd0LJ6"
# current range
img = images[0]
print('Min: ', img.min())
print('Max: ', img.max())
# + id="uCphu5h60LJ6"
# helper scale function
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-1.'''
# scale from 0-1 to feature_range
min, max = feature_range
x = x * (max - min) + min
return x
# + id="lv9-BMLB0LJ7"
# scaled range
scaled_img = scale(img)
print('Scaled min: ', scaled_img.min())
print('Scaled max: ', scaled_img.max())
# + [markdown] id="2qsIC5gy0LJ7"
# ---
# ## Define the Model
#
# A CycleGAN is made of two discriminator and two generator networks.
#
# ## Discriminators
#
# The discriminators, $D_X$ and $D_Y$, in this CycleGAN are convolutional neural networks that see an image and attempt to classify it as real or fake. In this case, real is indicated by an output close to 1 and fake as close to 0. The discriminators have the following architecture:
#
# <img src='notebook_images/discriminator_layers.png' width=80% />
#
# This network sees a 128x128x3 image, and passes it through 5 convolutional layers that downsample the image by a factor of 2. The first four convolutional layers have a BatchNorm and ReLu activation function applied to their output, and the last acts as a classification layer that outputs a prediction map with depth of one. Contrary to what the figure above indicates, the final output is not required to have a width and depth of one. In the original paper, the authors passed a 4x4 kernel with stride of 1 in the final convolutional layer. You should replicate that strategy.
#
# ### Convolutional Helper Function
#
# To define the discriminators, you're expected to use the provided `conv` function, which creates a convolutional layer + an optional batch norm layer.
# + id="nuOYxKCDCRz2"
# + id="0zhEfQzs0LJ8"
import torch.nn as nn
import torch.nn.functional as F
# helper conv function
def conv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a convolutional layer, with optional batch normalization.
"""
layers = []
conv_layer = nn.Conv2d(in_channels=in_channels, out_channels=out_channels,
kernel_size=kernel_size, stride=stride, padding=padding, bias=False)
layers.append(conv_layer)
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
# + [markdown] id="8_wgy0jb0LJ8"
# ### Define the Discriminator Architecture
#
# Your task is to fill in the `__init__` function with the specified 5 layer conv net architecture. Both $D_X$ and $D_Y$ have the same architecture, so we only need to define one class, and later instantiate two discriminators.
# > It's recommended that you use a **kernel size of 4x4** and use that to determine the correct stride and padding size for each layer. [This Stanford resource](http://cs231n.github.io/convolutional-networks/#conv) may also help in determining stride and padding sizes.
#
# * Define your convolutional layers in `__init__`
# * Then fill in the forward behavior of the network
#
# The `forward` function defines how an input image moves through the discriminator, and the most important thing is to pass it through your convolutional layers in order, with a **ReLu** activation function applied to all but the last layer.
#
# You should **not** apply a sigmoid activation function to the output, here, and that is because we are planning on using a squared error loss for training. And you can read more about this loss function, later in the notebook.
# + id="83Gyuif_0LJ9"
class Discriminator(nn.Module):
def __init__(self, conv_dim=64):
super(Discriminator, self).__init__()
# Define all convolutional layers
# Should accept an RGB image as input and output a single value
self.c1 = conv(3, conv_dim, 4, batch_norm=False)
self.c2 = conv(conv_dim, 2*conv_dim, 4)
self.c3 = conv(2*conv_dim, 4*conv_dim, 4)
self.classifier1 = nn.Linear(4*4*4*conv_dim, 4*conv_dim)
self.classifier2 = nn.Linear(4*conv_dim, conv_dim)
self.out = nn.Linear(conv_dim, 1)
self.leaky = nn.LeakyReLU(0.2)
self.relu = nn.ReLU()
self.flat = nn.Flatten()
self.drop = nn.Dropout(0.25)
def forward(self, x):
# define feedforward behavior
x = self.leaky(self.c1(x))
x = self.leaky(self.c2(x))
x = self.leaky(self.c3(x))
x = self.drop(self.relu(self.flat(x)))
x = self.drop(self.relu(self.classifier1(x)))
x = self.drop(self.relu(self.classifier2(x)))
x = self.out(x)
return x
# + [markdown] id="a6hm57FV0LJ9"
# ## Generators
#
# The generators, `G_XtoY` and `G_YtoX` (sometimes called F), are made of an **encoder**, a conv net that is responsible for turning an image into a smaller feature representation, and a **decoder**, a *transpose_conv* net that is responsible for turning that representation into an transformed image. These generators, one from XtoY and one from YtoX, have the following architecture:
#
# <img src='notebook_images/cyclegan_generator_ex.png' width=90% />
#
# This network sees a 128x128x3 image, compresses it into a feature representation as it goes through three convolutional layers and reaches a series of residual blocks. It goes through a few (typically 6 or more) of these residual blocks, then it goes through three transpose convolutional layers (sometimes called *de-conv* layers) which upsample the output of the resnet blocks and create a new image!
#
# Note that most of the convolutional and transpose-convolutional layers have BatchNorm and ReLu functions applied to their outputs with the exception of the final transpose convolutional layer, which has a `tanh` activation function applied to the output. Also, the residual blocks are made of convolutional and batch normalization layers, which we'll go over in more detail, next.
# + [markdown] id="EFaSPkAq0LJ9"
# ---
# ### Residual Block Class
#
# To define the generators, you're expected to define a `ResidualBlock` class which will help you connect the encoder and decoder portions of the generators. You might be wondering, what exactly is a Resnet block? It may sound familiar from something like ResNet50 for image classification, pictured below.
#
# <img src='notebook_images/resnet_50.png' width=90%/>
#
# ResNet blocks rely on connecting the output of one layer with the input of an earlier layer. The motivation for this structure is as follows: very deep neural networks can be difficult to train. Deeper networks are more likely to have vanishing or exploding gradients and, therefore, have trouble reaching convergence; batch normalization helps with this a bit. However, during training, we often see that deep networks respond with a kind of training degradation. Essentially, the training accuracy stops improving and gets saturated at some point during training. In the worst cases, deep models would see their training accuracy actually worsen over time!
#
# One solution to this problem is to use **Resnet blocks** that allow us to learn so-called *residual functions* as they are applied to layer inputs. You can read more about this proposed architecture in the paper, [Deep Residual Learning for Image Recognition](https://arxiv.org/pdf/1512.03385.pdf) by Kaiming He et. al, and the below image is from that paper.
#
# <img src='notebook_images/resnet_block.png' width=40%/>
#
# ### Residual Functions
#
# Usually, when we create a deep learning model, the model (several layers with activations applied) is responsible for learning a mapping, `M`, from an input `x` to an output `y`.
# >`M(x) = y` (Equation 1)
#
# Instead of learning a direct mapping from `x` to `y`, we can instead define a **residual function**
# > `F(x) = M(x) - x`
#
# This looks at the difference between a mapping applied to x and the original input, x. `F(x)` is, typically, two convolutional layers + normalization layer and a ReLu in between. These convolutional layers should have the same number of inputs as outputs. This mapping can then be written as the following; a function of the residual function and the input x. The addition step creates a kind of loop that connects the input x to the output, y:
# >`M(x) = F(x) + x` (Equation 2) or
#
# >`y = F(x) + x` (Equation 3)
#
# #### Optimizing a Residual Function
#
# The idea is that it is easier to optimize this residual function `F(x)` than it is to optimize the original mapping `M(x)`. Consider an example; what if we want `y = x`?
#
# From our first, direct mapping equation, **Equation 1**, we could set `M(x) = x` but it is easier to solve the residual equation `F(x) = 0`, which, when plugged in to **Equation 3**, yields `y = x`.
#
#
# ### Defining the `ResidualBlock` Class
#
# To define the `ResidualBlock` class, we'll define residual functions (a series of layers), apply them to an input x and add them to that same input. This is defined just like any other neural network, with an `__init__` function and the addition step in the `forward` function.
#
# In our case, you'll want to define the residual block as:
# * Two convolutional layers with the same size input and output
# * Batch normalization applied to the outputs of the convolutional layers
# * A ReLu function on the output of the *first* convolutional layer
#
# Then, in the `forward` function, add the input x to this residual block. Feel free to use the helper `conv` function from above to create this block.
# + id="Rccbwzjm0LJ-"
# residual block class
class ResidualBlock(nn.Module):
"""Defines a residual block.
This adds an input x to a convolutional layer (applied to x) with the same size input and output.
These blocks allow a model to learn an effective transformation from one domain to another.
s = 1, k = 3, x = x = ((x − k + 2p)/s) + 1
x = x - 3 + 2p + 1 --> p = 1
"""
def __init__(self, conv_dim):
super(ResidualBlock, self).__init__()
# conv_dim = number of inputs
# define two convolutional layers + batch normalization that will act as our residual function, F(x)
# layers should have the same shape input as output; I suggest a kernel_size of 3
self.relu = nn.ReLU()
self.layer1 = conv(conv_dim, conv_dim, kernel_size=3, stride=1, padding=1, batch_norm=True)
self.layer2 = conv(conv_dim, conv_dim, kernel_size=3, stride=1, padding=1, batch_norm=True)
def forward(self, x):
# apply a ReLu activation the outputs of the first layer
# return a summed output, x + resnet_block(x)
fx = x
fx = self.relu(self.layer1(fx))
fx = self.layer2(fx)
return fx + x
# + [markdown] id="kPzGaKwu0LJ-"
# ### Transpose Convolutional Helper Function
#
# To define the generators, you're expected to use the above `conv` function, `ResidualBlock` class, and the below `deconv` helper function, which creates a transpose convolutional layer + an optional batchnorm layer.
# + id="jqYziMEr0LJ_"
# helper deconv function
def deconv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a transpose convolutional layer, with optional batch normalization.
"""
layers = []
# append transpose conv layer
layers.append(nn.ConvTranspose2d(in_channels, out_channels, kernel_size, stride, padding, bias=False))
# optional batch norm layer
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
# + [markdown] id="-dh_RTD_0LJ_"
# ---
# ## Define the Generator Architecture
#
# * Complete the `__init__` function with the specified 3 layer **encoder** convolutional net, a series of residual blocks (the number of which is given by `n_res_blocks`), and then a 3 layer **decoder** transpose convolutional net.
# * Then complete the `forward` function to define the forward behavior of the generators. Recall that the last layer has a `tanh` activation function.
#
# Both $G_{XtoY}$ and $G_{YtoX}$ have the same architecture, so we only need to define one class, and later instantiate two generators.
# + id="Fz4h4nyz0LJ_"
class CycleGenerator(nn.Module):
def __init__(self, conv_dim=64, n_res_blocks=6):
super(CycleGenerator, self).__init__()
# 1. Define the encoder part of the generator
# 128 x 128 x 3 <-- in
self.encode1 = conv(3, conv_dim, kernel_size=4, stride=2, padding=1, batch_norm=True)
# 64
self.encode2 = conv(conv_dim, 2*conv_dim, kernel_size=4, stride=2, padding=1, batch_norm=True)
# 32
self.encode3 = conv(2*conv_dim, 4*conv_dim, kernel_size=4, stride=2, padding=1, batch_norm=True)
# 16 x 16 x 4*conv_dim --> out
self.relu = nn.ReLU()
self.tanh = nn.Tanh()
# 2. Define the resnet part of the generator
# in --> 16 x 16 x 4*conv_dim --> out
blocks = [ResidualBlock(4*conv_dim) for _ in range(n_res_blocks)]
# 3. Define the decoder part of the generator
#in --> 16 x 16 x 4*conv_dim
self.decode3 = deconv(4*conv_dim, 2*conv_dim, kernel_size=2, stride=2, padding=1, batch_norm=True)
# 32
self.decode2 = deconv(2*conv_dim, conv_dim, kernel_size=2, stride=2, padding=1, batch_norm=True)
# 64
self.decode1 = conv(conv_dim, 3, kernel_size=2, stride=2, padding=1, batch_norm=False)
# 128 x 128 x 3 --> out
def forward(self, x):
"""Given an image x, returns a transformed image."""
# define feedforward behavior, applying activations as necessary
# encode
x = self.relu(self.encode1(x))
x = self.relu(self.encode2(x))
x = self.relu(self.encode3(x))
# resnet blocks
for res_block in blocks:
x = res_block(x)
# decode
x = self.relu(self.decode3(x))
x = self.relu(self.decode2(x))
x = self.tanh(self.decode1(x))
return x
# + [markdown] id="l5n3l6Xc0LJ_"
# ---
# ## Create the complete network
#
# Using the classes you defined earlier, you can define the discriminators and generators necessary to create a complete CycleGAN. The given parameters should work for training.
#
# First, create two discriminators, one for checking if $X$ sample images are real, and one for checking if $Y$ sample images are real. Then the generators. Instantiate two of them, one for transforming a painting into a realistic photo and one for transforming a photo into into a painting.
# + id="WSSff0MM0LJ_"
def create_model(g_conv_dim=64, d_conv_dim=64, n_res_blocks=6):
"""Builds the generators and discriminators."""
# Instantiate generators
G_XtoY =
G_YtoX =
# Instantiate discriminators
D_X =
D_Y =
# move models to GPU, if available
if torch.cuda.is_available():
device = torch.device("cuda:0")
G_XtoY.to(device)
G_YtoX.to(device)
D_X.to(device)
D_Y.to(device)
print('Models moved to GPU.')
else:
print('Only CPU available.')
return G_XtoY, G_YtoX, D_X, D_Y
# + id="qfGDr81_0LKA"
# call the function to get models
G_XtoY, G_YtoX, D_X, D_Y = create_model()
# + [markdown] id="thcfKGNB0LKA"
# ## Check that you've implemented this correctly
#
# The function `create_model` should return the two generator and two discriminator networks. After you've defined these discriminator and generator components, it's good practice to check your work. The easiest way to do this is to print out your model architecture and read through it to make sure the parameters are what you expected. The next cell will print out their architectures.
# + id="m5NofGNU0LKA"
# helper function for printing the model architecture
def print_models(G_XtoY, G_YtoX, D_X, D_Y):
"""Prints model information for the generators and discriminators.
"""
print(" G_XtoY ")
print("-----------------------------------------------")
print(G_XtoY)
print()
print(" G_YtoX ")
print("-----------------------------------------------")
print(G_YtoX)
print()
print(" D_X ")
print("-----------------------------------------------")
print(D_X)
print()
print(" D_Y ")
print("-----------------------------------------------")
print(D_Y)
print()
# print all of the models
print_models(G_XtoY, G_YtoX, D_X, D_Y)
# + [markdown] id="Z7xzWJPw0LKA"
# ## Discriminator and Generator Losses
#
# Computing the discriminator and the generator losses are key to getting a CycleGAN to train.
#
# img src='notebook_images/CycleGAN_loss.png' width=90% height=90% />
#
# **Image from [original paper](https://arxiv.org/abs/1703.10593) by <NAME> et. al.**
#
# * The CycleGAN contains two mapping functions $G: X \rightarrow Y$ and $F: Y \rightarrow X$, and associated adversarial discriminators $D_Y$ and $D_X$. **(a)** $D_Y$ encourages $G$ to translate $X$ into outputs indistinguishable from domain $Y$, and vice versa for $D_X$ and $F$.
#
# * To further regularize the mappings, we introduce two cycle consistency losses that capture the intuition that if
# we translate from one domain to the other and back again we should arrive at where we started. **(b)** Forward cycle-consistency loss and **(c)** backward cycle-consistency loss.
#
# ## Least Squares GANs
#
# We've seen that regular GANs treat the discriminator as a classifier with the sigmoid cross entropy loss function. However, this loss function may lead to the vanishing gradients problem during the learning process. To overcome such a problem, we'll use a least squares loss function for the discriminator. This structure is also referred to as a least squares GAN or LSGAN, and you can [read the original paper on LSGANs, here](https://arxiv.org/pdf/1611.04076.pdf). The authors show that LSGANs are able to generate higher quality images than regular GANs and that this loss type is a bit more stable during training!
#
# ### Discriminator Losses
#
# The discriminator losses will be mean squared errors between the output of the discriminator, given an image, and the target value, 0 or 1, depending on whether it should classify that image as fake or real. For example, for a *real* image, `x`, we can train $D_X$ by looking at how close it is to recognizing and image `x` as real using the mean squared error:
#
# ```
# out_x = D_X(x)
# real_err = torch.mean((out_x-1)**2)
# ```
#
# ### Generator Losses
#
# Calculating the generator losses will look somewhat similar to calculating the discriminator loss; there will still be steps in which you generate fake images that look like they belong to the set of $X$ images but are based on real images in set $Y$, and vice versa. You'll compute the "real loss" on those generated images by looking at the output of the discriminator as it's applied to these _fake_ images; this time, your generator aims to make the discriminator classify these fake images as *real* images.
#
# #### Cycle Consistency Loss
#
# In addition to the adversarial losses, the generator loss terms will also include the **cycle consistency loss**. This loss is a measure of how good a reconstructed image is, when compared to an original image.
#
# Say you have a fake, generated image, `x_hat`, and a real image, `y`. You can get a reconstructed `y_hat` by applying `G_XtoY(x_hat) = y_hat` and then check to see if this reconstruction `y_hat` and the orginal image `y` match. For this, we recommend calculating the L1 loss, which is an absolute difference, between reconstructed and real images. You may also choose to multiply this loss by some weight value `lambda_weight` to convey its importance.
#
# img src='notebook_images/reconstruction_error.png' width=40% height=40% />
#
# The total generator loss will be the sum of the generator losses and the forward and backward cycle consistency losses.
# + [markdown] id="J_JygYa30LKB"
# ---
# ### Define Loss Functions
#
# To help us calculate the discriminator and gnerator losses during training, let's define some helpful loss functions. Here, we'll define three.
# 1. `real_mse_loss` that looks at the output of a discriminator and returns the error based on how close that output is to being classified as real. This should be a mean squared error.
# 2. `fake_mse_loss` that looks at the output of a discriminator and returns the error based on how close that output is to being classified as fake. This should be a mean squared error.
# 3. `cycle_consistency_loss` that looks at a set of real image and a set of reconstructed/generated images, and returns the mean absolute error between them. This has a `lambda_weight` parameter that will weight the mean absolute error in a batch.
#
# It's recommended that you take a [look at the original, CycleGAN paper](https://arxiv.org/pdf/1703.10593.pdf) to get a starting value for `lambda_weight`.
#
#
# + id="L72R64do0LKB"
def real_mse_loss(D_out):
# how close is the produced output from being "real"?
def fake_mse_loss(D_out):
# how close is the produced output from being "fake"?
def cycle_consistency_loss(real_im, reconstructed_im, lambda_weight):
# calculate reconstruction loss
# return weighted loss
# + [markdown] id="etUjIGW30LKB"
# ### Define the Optimizers
#
# Next, let's define how this model will update its weights. This, like the GANs you may have seen before, uses [Adam](https://pytorch.org/docs/stable/optim.html#algorithms) optimizers for the discriminator and generator. It's again recommended that you take a [look at the original, CycleGAN paper](https://arxiv.org/pdf/1703.10593.pdf) to get starting hyperparameter values.
#
# + id="8Vy4iUWc0LKC"
import torch.optim as optim
# hyperparams for Adam optimizers
lr=
beta1=
beta2=
g_params = list(G_XtoY.parameters()) + list(G_YtoX.parameters()) # Get generator parameters
# Create optimizers for the generators and discriminators
g_optimizer = optim.Adam(g_params, lr, [beta1, beta2])
d_x_optimizer = optim.Adam(D_X.parameters(), lr, [beta1, beta2])
d_y_optimizer = optim.Adam(D_Y.parameters(), lr, [beta1, beta2])
# + [markdown] id="MwWJOMvE0LKC"
# ---
#
# ## Training a CycleGAN
#
# When a CycleGAN trains, and sees one batch of real images from set $X$ and $Y$, it trains by performing the following steps:
#
# **Training the Discriminators**
# 1. Compute the discriminator $D_X$ loss on real images
# 2. Generate fake images that look like domain $X$ based on real images in domain $Y$
# 3. Compute the fake loss for $D_X$
# 4. Compute the total loss and perform backpropagation and $D_X$ optimization
# 5. Repeat steps 1-4 only with $D_Y$ and your domains switched!
#
#
# **Training the Generators**
# 1. Generate fake images that look like domain $X$ based on real images in domain $Y$
# 2. Compute the generator loss based on how $D_X$ responds to fake $X$
# 3. Generate *reconstructed* $\hat{Y}$ images based on the fake $X$ images generated in step 1
# 4. Compute the cycle consistency loss by comparing the reconstructions with real $Y$ images
# 5. Repeat steps 1-4 only swapping domains
# 6. Add up all the generator and reconstruction losses and perform backpropagation + optimization
#
# img src='notebook_images/cycle_consistency_ex.png' width=70% />
#
#
# ### Saving Your Progress
#
# A CycleGAN repeats its training process, alternating between training the discriminators and the generators, for a specified number of training iterations. You've been given code that will save some example generated images that the CycleGAN has learned to generate after a certain number of training iterations. Along with looking at the losses, these example generations should give you an idea of how well your network has trained.
#
# Below, you may choose to keep all default parameters; your only task is to calculate the appropriate losses and complete the training cycle.
# + id="R5RfFZFb0LKC"
# import save code
from helpers import save_samples, checkpoint
# + id="rAgIRlNC0LKC"
# train the network
def training_loop(dataloader_X, dataloader_Y, test_dataloader_X, test_dataloader_Y,
n_epochs=1000):
print_every=10
# keep track of losses over time
losses = []
test_iter_X = iter(test_dataloader_X)
test_iter_Y = iter(test_dataloader_Y)
# Get some fixed data from domains X and Y for sampling. These are images that are held
# constant throughout training, that allow us to inspect the model's performance.
fixed_X = test_iter_X.next()[0]
fixed_Y = test_iter_Y.next()[0]
fixed_X = scale(fixed_X) # make sure to scale to a range -1 to 1
fixed_Y = scale(fixed_Y)
# batches per epoch
iter_X = iter(dataloader_X)
iter_Y = iter(dataloader_Y)
batches_per_epoch = min(len(iter_X), len(iter_Y))
for epoch in range(1, n_epochs+1):
# Reset iterators for each epoch
if epoch % batches_per_epoch == 0:
iter_X = iter(dataloader_X)
iter_Y = iter(dataloader_Y)
images_X, _ = iter_X.next()
images_X = scale(images_X) # make sure to scale to a range -1 to 1
images_Y, _ = iter_Y.next()
images_Y = scale(images_Y)
# move images to GPU if available (otherwise stay on CPU)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
images_X = images_X.to(device)
images_Y = images_Y.to(device)
# ============================================
# TRAIN THE DISCRIMINATORS
# ============================================
## First: D_X, real and fake loss components ##
# 1. Compute the discriminator losses on real images
# 2. Generate fake images that look like domain X based on real images in domain Y
# 3. Compute the fake loss for D_X
# 4. Compute the total loss and perform backprop
d_x_loss =
## Second: D_Y, real and fake loss components ##
d_y_loss =
# =========================================
# TRAIN THE GENERATORS
# =========================================
## First: generate fake X images and reconstructed Y images ##
# 1. Generate fake images that look like domain X based on real images in domain Y
# 2. Compute the generator loss based on domain X
# 3. Create a reconstructed y
# 4. Compute the cycle consistency loss (the reconstruction loss)
## Second: generate fake Y images and reconstructed X images ##
# 5. Add up all generator and reconstructed losses and perform backprop
g_total_loss =
# Print the log info
if epoch % print_every == 0:
# append real and fake discriminator losses and the generator loss
losses.append((d_x_loss.item(), d_y_loss.item(), g_total_loss.item()))
print('Epoch [{:5d}/{:5d}] | d_X_loss: {:6.4f} | d_Y_loss: {:6.4f} | g_total_loss: {:6.4f}'.format(
epoch, n_epochs, d_x_loss.item(), d_y_loss.item(), g_total_loss.item()))
sample_every=100
# Save the generated samples
if epoch % sample_every == 0:
G_YtoX.eval() # set generators to eval mode for sample generation
G_XtoY.eval()
save_samples(epoch, fixed_Y, fixed_X, G_YtoX, G_XtoY, batch_size=16)
G_YtoX.train()
G_XtoY.train()
# uncomment these lines, if you want to save your model
# checkpoint_every=1000
# # Save the model parameters
# if epoch % checkpoint_every == 0:
# checkpoint(epoch, G_XtoY, G_YtoX, D_X, D_Y)
return losses
# + id="nrPTH-W30LKD"
n_epochs = 10 # keep this small when testing if a model first works, then increase it to >=1000
losses = training_loop(dataloader_X, dataloader_Y, test_dataloader_X, test_dataloader_Y, n_epochs=n_epochs)
# + [markdown] id="2HlSa7If0LKD"
# ## Tips on Training and Loss Patterns
#
# A lot of experimentation goes into finding the best hyperparameters such that the generators and discriminators don't overpower each other. It's often a good starting point to look at existing papers to find what has worked in previous experiments, I'd recommend this [DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf) in addition to the original [CycleGAN paper](https://arxiv.org/pdf/1703.10593.pdf) to see what worked for them. Then, you can try your own experiments based off of a good foundation.
#
# #### Discriminator Losses
#
# When you display the generator and discriminator losses you should see that there is always some discriminator loss; recall that we are trying to design a model that can generate good "fake" images. So, the ideal discriminator will not be able to tell the difference between real and fake images and, as such, will always have some loss. You should also see that $D_X$ and $D_Y$ are roughly at the same loss levels; if they are not, this indicates that your training is favoring one type of discriminator over the and you may need to look at biases in your models or data.
#
# #### Generator Loss
#
# The generator's loss should start significantly higher than the discriminator losses because it is accounting for the loss of both generators *and* weighted reconstruction errors. You should see this loss decrease a lot at the start of training because initial, generated images are often far-off from being good fakes. After some time it may level off; this is normal since the generator and discriminator are both improving as they train. If you see that the loss is jumping around a lot, over time, you may want to try decreasing your learning rates or changing your cycle consistency loss to be a little more/less weighted.
#
# + id="iIBRPc4F0LKD"
fig, ax = plt.subplots(figsize=(12,8))
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator, X', alpha=0.5)
plt.plot(losses.T[1], label='Discriminator, Y', alpha=0.5)
plt.plot(losses.T[2], label='Generators', alpha=0.5)
plt.title("Training Losses")
plt.legend()
# + [markdown] id="jJsfyBQo0LKE"
# ---
# ## Evaluate the Result!
#
# As you trained this model, you may have chosen to sample and save the results of your generated images after a certain number of training iterations. This gives you a way to see whether or not your Generators are creating *good* fake images. For example, the image below depicts real images in the $Y$ set, and the corresponding generated images during different points in the training process. You can see that the generator starts out creating very noisy, fake images, but begins to converge to better representations as it trains (though, not perfect).
#
# img src='notebook_images/sample-004000-summer2winter.png' width=50% />
#
# Below, you've been given a helper function for displaying generated samples based on the passed in training iteration.
# + id="mjCPHfut0LKE"
import matplotlib.image as mpimg
# helper visualization code
def view_samples(iteration, sample_dir='samples_cyclegan'):
# samples are named by iteration
path_XtoY = os.path.join(sample_dir, 'sample-{:06d}-X-Y.png'.format(iteration))
path_YtoX = os.path.join(sample_dir, 'sample-{:06d}-Y-X.png'.format(iteration))
# read in those samples
try:
x2y = mpimg.imread(path_XtoY)
y2x = mpimg.imread(path_YtoX)
except:
print('Invalid number of iterations.')
fig, (ax1, ax2) = plt.subplots(figsize=(18,20), nrows=2, ncols=1, sharey=True, sharex=True)
ax1.imshow(x2y)
ax1.set_title('X to Y')
ax2.imshow(y2x)
ax2.set_title('Y to X')
# + id="ji3YOZ880LKE"
# view samples at iteration 100
view_samples(100, 'samples_cyclegan')
# + id="aXoZco9J0LKE"
# view samples at iteration 1000
view_samples(1000, 'samples_cyclegan')
# + [markdown] id="6mtyMm-80LKE"
# ---
# ## Further Challenges and Directions
#
# * One shortcoming of this model is that it produces fairly low-resolution images; this is an ongoing area of research; you can read about a higher-resolution formulation that uses a multi-scale generator model, in [this paper](https://arxiv.org/abs/1711.11585).
# * Relatedly, we may want to process these as larger (say 256x256) images at first, to take advantage of high-res data.
# * It may help your model to converge faster, if you initialize the weights in your network.
# * This model struggles with matching colors exactly. This is because, if $G_{YtoX}$ and $G_{XtoY}$ may change the tint of an image; the cycle consistency loss may not be affected and can still be small. You could choose to introduce a new, color-based loss term that compares $G_{YtoX}(y)$ and $y$, and $G_{XtoY}(x)$ and $x$, but then this becomes a supervised learning approach.
# * This unsupervised approach also struggles with geometric changes, like changing the apparent size of individual object in an image, so it is best suited for stylistic transformations.
# * For creating different kinds of models or trying out the Pix2Pix Architecture, [this Github repository](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/) which implements CycleGAN *and* Pix2Pix in PyTorch is a great resource.
#
# **Once you are satified with your model, you are ancouraged to test it on a different dataset to see if it can find different types of mappings!**
#
# ---
#
# ### Different datasets for download
#
# You can download a variety of datasets used in the Pix2Pix and CycleGAN papers, by following instructions in the [associated Github repository](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/blob/master/README.md). You'll just need to make sure that the data directories are named and organized correctly to load in that data.
|
cycle-gan/CycleGAN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "-"}
# # Exploring convolution
#
# First, include some libraries
# +
# Run boilerplate code to set up environment
# %run ../prelude.py --style=tree --animation=spacetime
# -
# ## Convolution Inputs
#
# +
tm = TensorMaker()
W = 8
S = 3
tm.addTensor("I", rank_ids=["W"], shape=[W], density=0.60, seed=40, color="blue")
tm.addTensor("F", rank_ids=["S"], shape=[S], density=0.50, seed=10, color="green")
tm.displayControls()
# +
i = tm.makeTensor("I")
f = tm.makeTensor("F")
S = f.getShape()[0]
W = i.getShape()[0]
Q = W-S+1
print(f"W = {W}")
print(f"S = {S}")
print(f"Q = {Q}")
print("")
o_verify= Tensor(rank_ids=["Q"], shape=[Q])
o = o_verify.getRoot()
for q in range(Q):
o_ref = o.getPayloadRef(q)
for s in range(S):
w = q+s
o_ref += i.getPayload(w) * f.getPayload(s)
print("Input activations")
displayTensor(i)
print("Filter Weights")
displayTensor(f)
print("Output activations (expected)")
displayTensor(o_verify)
# -
# ## Weight Stationary
# +
o = Tensor(rank_ids=["Q"]).setName("O")
canvas = createCanvas(f, i, o)
f_s = f.getRoot()
i_w = i.getRoot()
o_q = o.getRoot()
print("Convolution")
for s, (f_val) in f_s:
print(f"Processing weight: ({s}, ({f_val}))")
for q, (o_ref, i_val) in o_q << i_w.project(lambda h: h-s, (0, Q)):
print(f" Processing output ({q}, ({o_ref}, {i_val})")
o_ref += f_val * i_val
canvas.addFrame((s,), (q+s,), (q,))
displayTensor(o)
displayCanvas(canvas)
assert o == o_verify
# -
# ## Input Stationary
# +
o = Tensor(rank_ids=["Q"]).setName("O")
canvas = createCanvas(f, i, o)
f_s = f.getRoot()
i_w = i.getRoot()
o_q = o.getRoot()
print("Convolution")
for w, (i_val) in i_w:
print(f"Processing input: ({w}, ({i_val}))")
for q, (o_ref, f_val) in o_q << f_s.project(lambda s: w-s, (0, Q)):
print(f" Processing output ({q}, ({o_ref}, {f_val})")
o_ref += f_val * i_val
canvas.addFrame((w-q,), (w,), (q,))
displayTensor(o)
displayCanvas(canvas)
assert o == o_verify
# -
# ## Output Stationary
# +
o = Tensor(rank_ids=["Q"]).setName("O")
f_s = f.getRoot()
i_w = i.getRoot()
o_q = o.getRoot()
print("Convolution")
output_shape = Fiber(coords=range(Q), initial=1)
canvas = createCanvas(f, i, o)
for q, (o_ref, _) in o_q << output_shape:
print(f"Processing output: ({q}, ({o_ref}))")
for w, (f_val, i_val) in f_s.project(lambda s: q+s) & i_w:
print(f" Processing weights and activations ({w}, ({f_val}, {i_val})")
o_ref += f_val * i_val
canvas.addFrame((w-q,), (w,), (q,))
displayTensor(o)
displayCanvas(canvas)
assert o == o_verify
# -
# ## Output Stationary - Two pass
# +
o1 = Tensor(rank_ids=["Q"]).setName("O1")
o2 = Tensor(rank_ids=["Q"]).setName("O2")
canvas = createCanvas(f, i, o1, o2)
f_s = f.getRoot()
i_w = i.getRoot()
o1_q = o1.getRoot()
o2_q = o2.getRoot()
print("Convolution")
pass1_count = 0
for s, (_) in f_s:
print(f"Processing weight: ({s}, (_))")
for q, (o_ref, _) in o1_q << i_w.project(lambda w: w-s, (0, Q)):
print(f" Calculating output ({q}, ({o_ref}, _)")
o_ref <<= 1
pass1_count += 1
canvas.addFrame((s,), (q+s,), (q,), ())
print(f"Pass1 count: {pass1_count}")
displayTensor(o1)
for q, (o_ref, _) in o2_q << o1_q:
print(f"Processing output: ({q}, ({o_ref}))")
for w, (f_val, i_val) in f_s.project(lambda s: q+s) & i_w:
print(f" Processing weights and activations ({w}, ({f_val}, {i_val})")
o_ref += f_val * i_val
canvas.addFrame((w-q,), (w,), (), (q,))
displayTensor(o)
displayCanvas(canvas)
assert o == o_verify
# -
# ## Output Stationary - Two pass - Optimized
# +
o1 = Tensor(rank_ids=["Q"]).setName("O1")
o2 = Tensor(rank_ids=["Q"]).setName("O2")
canvas = createCanvas(f, i, o1, o2)
f_s = f.getRoot()
i_w = i.getRoot()
o1_q = o1.getRoot()
o2_q = o2.getRoot()
print("Convolution")
pass1_count = 0
for s, (_) in f_s:
print(f"Processing weight: ({s}, (_))")
for q, (o1_ref, _) in o1_q << (i_w.project(lambda w: w-s, (0, Q)) - o1_q):
print(f" Calculating output ({q}, ({o_ref}, _)")
o1_ref <<= 1
pass1_count += 1
canvas.addFrame((s,), (q+s,), (q,), ())
print(f"{o1:*}")
displayTensor(o1)
print(f"Pass1 count: {pass1_count}")
for q, (o_ref, _) in o2_q << o1_q:
print(f"Processing output: ({q}, ({o_ref}))")
for w, (f_val, i_val) in f_s.project(lambda s: q+s) & i_w:
print(f" Processing weights and activations ({w}, ({f_val}, {i_val})")
o_ref += f_val * i_val
canvas.addFrame((w-q,), (w,), (), (q,))
displayTensor(o)
displayCanvas(canvas)
assert o == o_verify
# -
# ## Output Stationary - Naive - Parallel Weight Processing
#
# Assumes parallelism equal to the number of weights
# +
o = Tensor(rank_ids=["Q"]).setName("O")
f_s = f.getRoot()
i_w = i.getRoot()
o_q = o.getRoot()
print("Convolution")
output_shape = Fiber(coords=range(Q), initial=1)
canvas = createCanvas(f, i, o)
for q, (o_ref, _) in o_q << output_shape:
print(f"Processing output: ({q}, ({o_ref}))")
for w, (f_val, i_val) in f_s.project(lambda s: q+s) & i_w:
pe = f"PE{w-q}"
print(f" {pe}: Processing weights and activations ({w}, ({f_val}, {i_val})")
o_ref += f_val * i_val
canvas.addActivity((w-q,), (w,), (q,), worker=pe)
canvas.addFrame()
displayTensor(o)
displayCanvas(canvas)
assert o == o_verify
# -
# ## Testing area
#
# For running alternative algorithms
|
notebooks/sparse-dnn/convolution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: timeeval38
# language: python
# name: timeeval38
# ---
# # TimeEval parameter optimization result analysis of extra experiments (2)
#
# Extra experiments and their reason:
#
# - Random Black Forest (RR): was missing in extra1 run, because of a configuration error
# - ARIMA: Inspect (originally fixed) parameter "distance_metric"
# +
# imports
import json
import warnings
import pandas as pd
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
from pathlib import Path
from timeeval import Datasets
# -
# ## Configuration
# Define data and results folder:
# +
# constants and configuration
data_path = Path("/home/projects/akita/data") / "test-cases"
result_root_path = Path("/home/projects/akita/results")
experiment_result_folder = "2021-10-17_optim-extra2"
# build paths
result_paths = [d for d in result_root_path.iterdir() if d.is_dir()]
print("Available result directories:")
display(result_paths)
result_path = result_root_path / experiment_result_folder
print("\nSelecting:")
print(f"Data path: {data_path.resolve()}")
print(f"Result path: {result_path.resolve()}")
# -
# Load results and dataset metadata:
# +
# load results
print(f"Reading results from {result_path.resolve()}")
df = pd.read_csv(result_path / "results.csv")
# add dataset_name column
df["dataset_name"] = df["dataset"].str.split(".").str[0]
# load dataset metadata
dmgr = Datasets(data_path)
# -
# Extract target optimized parameter names that were iterated in this run (per algorithm):
# +
algo_param_mapping = {}
algorithms = df["algorithm"].unique()
param_ignore_list = ["max_anomaly_window_size", "anomaly_window_size", "neighbourhood_size", "window_size", "n_init_train", "embed_dim_range"]
for algo in algorithms:
param_sets = df.loc[df["algorithm"] == algo, "hyper_params"].unique()
param_sets = [json.loads(ps) for ps in param_sets]
param_names = np.unique([name for ps in param_sets for name in ps if name not in param_ignore_list])
search_space = set()
for param_name in param_names:
values = []
for ps in param_sets:
try:
values.append(ps[param_name])
except:
pass
values = np.unique(values)
if values.shape[0] > 1:
search_space.add(param_name)
algo_param_mapping[algo] = list(search_space)
for algo in algo_param_mapping:
print(algo, algo_param_mapping[algo])
# -
# Extract optimized parameters and their values (columns: optim_param_name and optim_param_value) for each experiment:
# +
def extract_hyper_params(algo):
param_names = algo_param_mapping[algo]
def extract(value):
params = json.loads(value)
result = ""
for name in param_names:
try:
value = params[name]
result += f"{name}={value},"
except KeyError:
pass
if result == "":
return pd.Series([np.nan, np.nan], index=["optim_param_name", "optim_param_value"])
elif len(param_names) == 1:
return pd.Series(result.rsplit(",", 1)[0].split("="), index=["optim_param_name", "optim_param_value"])
else:
return pd.Series(["", "".join(result.rsplit(",", 1))], index=["optim_param_name", "optim_param_value"])
return extract
df[["optim_param_name", "optim_param_value"]] = ""
for algo in algo_param_mapping:
df_algo = df.loc[df["algorithm"] == algo]
df.loc[df_algo.index, ["optim_param_name", "optim_param_value"]] = df_algo["hyper_params"].apply(extract_hyper_params(algo))
# -
# Define utility functions
def load_scores_df(algorithm_name, dataset_id, optim_params, repetition=1):
params_id = df.loc[(df["algorithm"] == algorithm_name) & (df["collection"] == dataset_id[0]) & (df["dataset"] == dataset_id[1]) & (df["optim_param_name"] == optim_params[0]) & (df["optim_param_value"] == optim_params[1]), "hyper_params_id"].item()
path = (
result_path /
algorithm_name /
params_id /
dataset_id[0] /
dataset_id[1] /
str(repetition) /
"anomaly_scores.ts"
)
return pd.read_csv(path, header=None)
# Define plotting functions:
# +
default_use_plotly = True
try:
import plotly.offline
except ImportError:
default_use_plotly = False
def plot_scores(algorithm_name, dataset_name, use_plotly: bool = default_use_plotly, **kwargs):
if isinstance(algorithm_name, tuple):
algorithms = [algorithm_name]
elif not isinstance(algorithm_name, list):
raise ValueError("Please supply a tuple (algorithm_name, optim_param_name, optim_param_value) or a list thereof as first argument!")
else:
algorithms = algorithm_name
# construct dataset ID
dataset_id = ("GutenTAG", f"{dataset_name}.unsupervised")
# load dataset details
df_dataset = dmgr.get_dataset_df(dataset_id)
# check if dataset is multivariate
dataset_dim = df.loc[df["dataset_name"] == dataset_name, "dataset_input_dimensionality"].unique().item()
dataset_dim = dataset_dim.lower()
auroc = {}
df_scores = pd.DataFrame(index=df_dataset.index)
skip_algos = []
algos = []
for algo, optim_param_name, optim_param_value in algorithms:
optim_params = f"{optim_param_name}={optim_param_value}"
algos.append((algo, optim_params))
# get algorithm metric results
try:
auroc[(algo, optim_params)] = df.loc[
(df["algorithm"] == algo) & (df["dataset_name"] == dataset_name) & (df["optim_param_name"] == optim_param_name) & (df["optim_param_value"] == optim_param_value),
"ROC_AUC"
].item()
except ValueError:
warnings.warn(f"No ROC_AUC score found! Probably {algo} with params {optim_params} was not executed on {dataset_name}.")
auroc[(algo, optim_params)] = -1
skip_algos.append((algo, optim_params))
continue
# load scores
training_type = df.loc[df["algorithm"] == algo, "algo_training_type"].values[0].lower().replace("_", "-")
try:
df_scores[(algo, optim_params)] = load_scores_df(algo, ("GutenTAG", f"{dataset_name}.{training_type}"), (optim_param_name, optim_param_value)).iloc[:, 0]
except (ValueError, FileNotFoundError):
warnings.warn(f"No anomaly scores found! Probably {algo} was not executed on {dataset_name} with params {optim_params}.")
df_scores[(algo, optim_params)] = np.nan
skip_algos.append((algo, optim_params))
algorithms = [a for a in algos if a not in skip_algos]
if use_plotly:
return plot_scores_plotly(algorithms, auroc, df_scores, df_dataset, dataset_dim, dataset_name, **kwargs)
else:
return plot_scores_plt(algorithms, auroc, df_scores, df_dataset, dataset_dim, dataset_name, **kwargs)
def plot_scores_plotly(algorithms, auroc, df_scores, df_dataset, dataset_dim, dataset_name, **kwargs):
import plotly.offline as py
import plotly.graph_objects as go
import plotly.figure_factory as ff
import plotly.express as px
from plotly.subplots import make_subplots
# Create plot
fig = make_subplots(2, 1)
if dataset_dim == "multivariate":
for i in range(1, df_dataset.shape[1]-1):
fig.add_trace(go.Scatter(x=df_dataset.index, y=df_dataset.iloc[:, i], name=f"channel-{i}"), 1, 1)
else:
fig.add_trace(go.Scatter(x=df_dataset.index, y=df_dataset.iloc[:, 1], name="timeseries"), 1, 1)
fig.add_trace(go.Scatter(x=df_dataset.index, y=df_dataset["is_anomaly"], name="label"), 2, 1)
for item in algorithms:
algo, optim_params = item
fig.add_trace(go.Scatter(x=df_scores.index, y=df_scores[item], name=f"{algo}={auroc[item]:.4f} ({optim_params})"), 2, 1)
fig.update_xaxes(matches="x")
fig.update_layout(
title=f"Results of {','.join(np.unique([a for a, _ in algorithms]))} on {dataset_name}",
height=400
)
return py.iplot(fig)
def plot_scores_plt(algorithms, auroc, df_scores, df_dataset, dataset_dim, dataset_name, **kwargs):
import matplotlib.pyplot as plt
# Create plot
fig, axs = plt.subplots(2, 1, sharex=True, figsize=(20, 8))
if dataset_dim == "multivariate":
for i in range(1, df_dataset.shape[1]-1):
axs[0].plot(df_dataset.index, df_dataset.iloc[:, i], label=f"channel-{i}")
else:
axs[0].plot(df_dataset.index, df_dataset.iloc[:, 1], label=f"timeseries")
axs[1].plot(df_dataset.index, df_dataset["is_anomaly"], label="label")
for item in algorithms:
algo, optim_params = item
axs[1].plot(df_scores.index, df_scores[item], label=f"{algo}={auroc[item]:.4f} ({optim_params})")
axs[0].legend()
axs[1].legend()
fig.suptitle(f"Results of {','.join(np.unique([a for a, _ in algorithms]))} on {dataset_name}")
fig.tight_layout()
return fig
# -
# ## Parameter assessment
# +
sort_by = ("ROC_AUC", "mean")
metric_agg_type = ["min", "mean", "median"]
time_agg_type = "mean"
aggs = {
"PR_AUC": metric_agg_type,
"ROC_AUC": metric_agg_type,
"train_main_time": time_agg_type,
"execute_main_time": time_agg_type,
"repetition": "count"
}
df_tmp = df.reset_index()
df_tmp = df_tmp.groupby(by=["algorithm", "optim_param_name", "optim_param_value"]).agg(aggs)
df_tmp = df_tmp.reset_index()
df_tmp = df_tmp.sort_values(by=["algorithm", "optim_param_name", sort_by], ascending=False)
df_tmp = df_tmp.set_index(["algorithm", "optim_param_name", "optim_param_value"])
with pd.option_context("display.max_rows", None, "display.max_columns", None):
display(df_tmp)
# -
# #### Selected parameters
#
# - Random Black Forest (RR):
# ```json
# "Random Black Forest (RR)": {
# "bootstrap": false,
# "n_trees": 10,
# "n_estimators": 200
# }
# ```
# - ARIMA:
# ```json
# "ARIMA": {
# "distance_metric": "twed"
# }
# ```
plot_scores([
("Random Black Forest (RR)", "", "bootstrap=False,n_trees=10,n_estimators=200"),
("Random Black Forest (RR)", "", "bootstrap=True,n_trees=10,n_estimators=200"),
("Random Black Forest (RR)", "", "bootstrap=False,n_trees=10,n_estimators=100")
], "ecg-type-variance", use_plotly=False)
plt.show()
# Failed runs
df[df["status"] != "Status.OK"].groupby(by=["algorithm", "optim_param_name", "optim_param_value", "status"])[["repetition"]].count()
algo = "Random Black Forest (RR)"
executions = [f for f in (result_path / algo).glob("**/execution.log") if not (f.parent / "anomaly_scores.ts").is_file()]
c = 0
for x in executions:
with x.open() as fh:
log = "".join(fh.readlines())
if "status code '137'" in log:
c += 1
else:
print(x.parent.parent.name)
print("---------------------------------------------------------------------------------")
print(log)
print(c)
|
notebooks/TimeEval parameter optimization analysis extra2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.5.2 64-bit
# language: python
# name: python35264bitf7dbc70352dc4e53b0cd48dd904a65a0
# ---
# + [markdown] colab_type="text" heading_collapsed=true id="Bpb2f9vJ1C-t"
# # 3. 変数と演算、代入
# + [markdown] colab_type="text" hidden=true id="Dw9zOz061C-v"
# ## 3.2 プログラムの実行の流れと情報の流れ
# + [markdown] colab_type="text" hidden=true id="daPmfH741C-w"
# ### 3.2.1 順次実行
# + colab_type="code" hidden=true id="SWFl6hiS1C-x" outputId="4e8bb05b-0493-4f3c-fca9-80e4895c3071" colab={}
a = 1 + 2
print(a)
# + [markdown] colab_type="text" hidden=true id="-upd0I4O1C-1"
# ### 3.2.2 変数を通じた情報の流れ
# + colab_type="code" hidden=true id="iVAYW_kc1C-1" outputId="e1ca5b3a-804b-493f-fa5a-ee9a787a6d62" colab={}
a = 1 + 2
a = 3 + 4
print(a)
# + [markdown] colab_type="text" hidden=true id="n0Ug-svo1C-4"
# ## 3.4 変数への代入と値の評価
# + colab_type="code" hidden=true id="6_ronT2b1C-5" outputId="6af8e39a-2895-49ff-b033-828e2985c561" colab={}
a = 1
print(a)
a = a + 1
print(a)
# + [markdown] colab_type="text" hidden=true id="5RrX53ry1C-7"
# #### 演習9. 変数の動作の説明
# + [markdown] colab_type="text" hidden=true id="TH2B9l-S1C-8"
# 以下は 1000 円の商品の 15% 引きを計算するプログラムです。
# - このプログラムには 1 箇所誤りがあり、実行するとエラーになります。どのような誤りがあるかを説明してください。
# - 誤りを修正したうえでプログラムの動作を説明してください。
# ```python
# kakaku = 1000
# nebikiritsu= 15
# kakaku = Kakaku*(100-nebikiritsu)/100
# print(kakaku)
# ```
# + [markdown] colab_type="text" hidden=true id="Ea3YpCX41C-9"
# ```python
# kakaku = Kakaku*(100-nebikiritsu)/100
# ```
# 右辺の変数`Kakaku`の先頭が大文字のため宣言されていない.
# + colab_type="code" hidden=true id="wAedC2xA1C-9" outputId="ec4893b6-84cb-457c-8812-4752a7799632" colab={}
kakaku = 1000
nebikiritsu= 15
kakaku = kakaku*(100-nebikiritsu)/100
print(kakaku)
# + [markdown] colab_type="text" hidden=true id="Wfh2jxbX1C_A"
# ## 3.6 Pythonで扱えるデータ型
# + [markdown] colab_type="text" hidden=true id="xpeGaz4d1C_B"
# #### 演習10. Pythonシェルで以下を実行してください。
# + colab_type="code" hidden=true id="KJnCJeD41C_C" outputId="6566fafd-d847-4051-f9ba-5d16d74cf2d3" colab={}
a = 1
b = 1/2
c = "ABC"
print(a)
print(b)
print(c)
print(type(a))
print(type(b))
print(type(c))
# + [markdown] colab_type="text" hidden=true id="vB2hao9h1C_E"
# ## 3.8 例題:平方根を求める
# + [markdown] colab_type="text" hidden=true id="0I0a9z0F1C_F"
# ### 3.8.2 Pythonプログラム
# + [markdown] colab_type="text" hidden=true id="_2jXI89M1C_F"
# #### 演習11. 次の表のソースコードの部分をIDELエディタで入力し, `ex1.py`という名で保存して実行してみてください。
# + colab_type="code" hidden=true id="mQuhxm6p1C_G" outputId="955d8f72-92f4-4393-d382-5f468cb8dd82" colab={}
# xの平方根を求める
x = 2
#
rnew = x
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
# + colab_type="code" hidden=true id="vruTKYoY1C_J" outputId="d0e3855d-4685-4d79-ae64-7ab5a55ee4d7" colab={}
2**(1/2)
# + [markdown] colab_type="text" hidden=true id="4AcFd2Jc1C_L"
# #### 演習12. 他の数値の平方根を求める。
# + [markdown] colab_type="text" hidden=true id="n73NRfRi1C_M"
# 1. `ex1.py`を変更して、他の正の数値の平方根を求めてください。
# + colab_type="code" hidden=true id="XEYdC7o41C_N" outputId="c1584e34-9692-48e8-ad89-e72b231718d2" colab={}
# xの平方根を求める
x = 3
#
rnew = x
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
# + colab_type="code" hidden=true id="POcqJkCI1C_P" outputId="bca21d78-8195-4dce-cf34-2f1c24c0d12a" colab={}
3**(1/2)
# + [markdown] colab_type="text" hidden=true id="ZgCkoD7_1C_R"
# 2. また、このプログラムで0の平方根を求めようとすると何が生じるか確認してください。単にエラーのメッセージを見るだけでなく、実際にプログラムをご自身で追いかけて(トレースすると言います)、どこで問題が生じるかを考えてください。
# + colab_type="code" hidden=true id="tcuPlx691C_S" outputId="e97f2522-0628-431e-8c56-1164f827fa56" colab={}
# xの平方根を求める
x = 0
#
rnew = x
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1,rnew,r2)
#
# + [markdown] colab_type="text" hidden=true id="iQbG8To01C_U"
# ```python
# r2 = x/r1
# ```
# 上でゼロ割りをおこなっている。
# + [markdown] colab_type="text" heading_collapsed=true id="bJiZd4jA1C_V"
# # 4. 制御構造
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="H_FA0zC_1C_X"
# ## 4.2 `for`文と`range()`関数を用いた一定回数の繰り返し
# + [markdown] colab_type="text" hidden=true id="q7h2P9XA1C_Y"
# #### 演習13. 次の表に示すプログラムを作成し、実行してください。
# + [markdown] colab_type="text" hidden=true id="G6yuBzCe1C_Y"
# プログラム3 平方根を求めるプログラム(その2、`ex2.py`)
# + colab_type="code" hidden=true id="NyIDtQw51C_Z" outputId="a5873876-e78a-4017-efe3-346a750d1c71" colab={}
# xの平方根を求める
x = 2
#
rnew = x
#
for i in range(10):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
# + colab_type="code" hidden=true id="_mvtT_-B1C_b" outputId="75f1f532-6df4-47a6-e507-6a9909e02c08" colab={}
2**(1/2)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="2KFc8hxu1C_d"
# ## 4.3 `for`文の書き方
# + [markdown] colab_type="text" hidden=true id="scjRlfVZ1C_e"
# #### 演習14. ブロックの確認
# + [markdown] colab_type="text" hidden=true id="5Ygkol5C1C_e"
# 先の例(`ex2.py`) の10行目を次のように左につめてブロックから外し、動作を確認し説明してください。
# + [markdown] colab_type="text" hidden=true id="j7390wiT1C_f"
# プログラム4 平方根を求めるプログラム(その2、`ex2_2.py`)
# + colab_type="code" hidden=true id="Uku2-nHh1C_f" outputId="e723f471-e1e1-471f-a32a-990cd45fc6f3" colab={}
# xの平方根を求める
x = 2
#
rnew = x
#
for i in range(10):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="sKonONud1C_i"
# #### 演習15. イタズラ
# + [markdown] colab_type="text" hidden=true id="AyUAAWkA1C_i"
# 上のプログラム(`ex2_2.py`)は端末への出力を`for`文の繰り返しから外したので、繰り返し部分は高速に実行できます。6行目の`range()`関数の添え字を10から100, 1000, 10000, 100000, 1000000, 10000000, と変化させてどの程度の時間がかかるか試してみてください。
# + colab_type="code" hidden=true id="019_E7gb1C_j" outputId="71686ec3-27c6-4edc-ee04-a4caa4ed80c1" colab={}
# xの平方根を求める
import time
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
time_list = list()
repeat_list = list()
for j in range(7):
start = time.time()
repeat_time = 10**(j+1)
repeat_list.append(repeat_time)
print("repeat time:", repeat_time)
x = 2
#
rnew = x
#
for i in range(repeat_time):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print("answer:", r2)
end = time.time()
exe_time = end-start
print("exe time:", exe_time, '秒')
time_list.append(exe_time)
plt.xlabel('repeat')
plt.ylabel('exe time')
ax = plt.gca()
ax.set_xscale('log')
ax.set_yscale('log')
plt.plot(repeat_list,time_list)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="Z_kzFgZW1C_l"
# ## 4.5 `for`文内での処理の制御
# + [markdown] colab_type="text" hidden=true id="hVmBRODu1C_m"
# プログラム5 `continue`と`break`
# + colab_type="code" hidden=true id="DICvv2KJ1C_m" outputId="d622aeef-c701-47b5-a05c-b8b2f28a56b6" colab={}
for i in range(10):
if i == 1:
continue
if i == 8:
break
print(i)
# + [markdown] colab_type="text" hidden=true id="hLCaua0R1C_o"
# #### 演習16. 上記の実行結果についてソースコードを用いて説明しなさい。
# + [markdown] colab_type="text" hidden=true id="0NJSffoV1C_p"
# `i=1`のとき`continue`が実行されてそれ以下のブロック内は実行されないため,
# ```python
# print(i)
# ```
# が実行されない.
# `i=8`のとき`break`が実行されてブロックから脱出するため, 8以降の`for`文は実行されない.
# + [markdown] colab_type="text" hidden=true id="_soncdHY1C_q"
# ## `range()`関数
# + [markdown] colab_type="text" hidden=true id="-iw20iWY1C_q"
# #### 演習17. `range()`関数
# + [markdown] colab_type="text" hidden=true id="jgJ4hmMf1C_r"
# 上で述べたように`list()`と組み合わせて`range()`関数の3通りの使い方をPythonシェルで練習してください。
# + [markdown] colab_type="text" hidden=true id="3jIwJbI41C_r"
# - 終了値を与える。
# + colab_type="code" hidden=true id="MXw1c61z1C_s" outputId="b29a5008-789f-4a1d-c2d8-3c93d467feb5" colab={}
# for (int i=0;i<10;i++)
l = list(range(10))
print(l)
# + [markdown] colab_type="text" hidden=true id="kM7AzEpX1C_u"
# - 開始値と終了値の2つを与える。
# + colab_type="code" hidden=true id="7K86r7wx1C_v" outputId="9c964fc7-819e-4214-f39b-e10168d88315" colab={}
# for (int i=1;i<10;i++)
l = list(range(1,10))
print(l)
# + [markdown] colab_type="text" hidden=true id="nSd7fwRM1C_x"
# - 開始値と終了値とステップ幅の3つを与える。
# + colab_type="code" hidden=true id="dXVmt34O1C_z" outputId="620b19cc-5315-4170-b8a1-70da66a6204d" colab={}
# for (int i=1;i<10;i+=2)
l = list(range(1,10,2))
print(l)
# + [markdown] colab_type="text" hidden=true id="Lzhu8Y4p1C_1"
# ## `for`文の入れ子
# + [markdown] colab_type="text" hidden=true id="3i9SDR3h1C_1"
# プログラム6 `for`文の入れ子
# + colab_type="code" hidden=true id="nd_lFi0R1C_2" outputId="153f685d-6f16-4d3e-dd93-5c02d73d30fd" colab={}
print("i j")
for i in range(3):
for j in range(3):
print(i,j)
# + [markdown] colab_type="text" hidden=true id="wF7Z2og81C_4"
# #### 演習18. 2行目の`range()`関数の引数に変数`i`を使う(`range(i)`をする)とどうなるか、試してみてください。
# + colab_type="code" hidden=true id="QRN79Xq51C_5" outputId="41f8b3e1-5850-42d2-c6b1-a8feffc3d31c" colab={}
print("i j")
for i in range(3):
for j in range(i):
print(i,j)
# + [markdown] colab_type="text" hidden=true id="jLNslxhh1C_7"
# プログラム7
# + colab_type="code" hidden=true id="UCVAFkMI1C_8" outputId="05459e9e-0725-4cfe-bb6e-a71354868d01" colab={}
for i in ["三条", "四条", "五条"]:
for j in ["河原町", "烏丸", "堀川"]:
cross = i+j
print(cross)
# + [markdown] colab_type="text" hidden=true id="263d52hC1C__"
# ## 4.8 `while`文による繰り返し
# + [markdown] colab_type="text" hidden=true id="Wj5-rZZN1C__"
# ### 4.8.1 精度を指定した平方根の計算
# + [markdown] colab_type="text" hidden=true id="VSOJ_Glq1DAA"
# プログラム8 平方根を計算するプログラム(その3、`ex3.py`)
# + colab_type="code" hidden=true id="V5BKebRI1DAA" outputId="91d5a2e4-f1b1-4f03-ea7b-c3c293cee5a6" colab={}
# xの平方根を求める
x = 2
#
rnew = x
#
diff = rnew - x/rnew
if (diff < 0):
diff = -diff
while (diff > 1.0E-6):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
diff = r1 - r2
if (diff < 0):
diff = -diff
# + [markdown] colab_type="text" hidden=true id="CVlfl8Pt1DAC"
# ## 4.9 `if`文による分岐
# + [markdown] colab_type="text" hidden=true id="B0tfCu-D1DAD"
# ### 4.9.1 無限ループ型での平方根の計算
# + [markdown] colab_type="text" hidden=true id="DVUWdTjP1DAE"
# プログラム9 平方根を求めるプログラム(無限ループ型、`ex3_2.py`)
# + colab_type="code" hidden=true id="PVN9ZF0I1DAE" outputId="46386944-9fca-4fa3-c4d7-0c5615c08212" colab={}
# xの平方根を求める
x = 2
#
rnew = x
#
while True:
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
diff = r1 - r2
if (diff < 0):
diff = -diff
if (diff <= 1.0E-6):
break
# + [markdown] colab_type="text" hidden=true id="YBsTVFGc1DAI"
# ## 4.11 `if`文の入れ子
# + [markdown] colab_type="text" hidden=true id="OkuZcm2T1DAI"
# プログラム10 複合的な条件を用いた分岐
# + colab_type="code" hidden=true id="vi7dX1B71DAJ" outputId="e79ecf66-4913-4570-9df1-b38f86931c80" colab={}
a = 1
b = 0
if (a == 1) and (b == 0):
print("YES a==1 and b==0")
# + [markdown] colab_type="text" hidden=true id="s_1cCiXD1DAK"
# プログラム11 `if`文を入れ子にした分岐
# + colab_type="code" hidden=true id="Rj_Myq2H1DAL" outputId="50600e9c-6aa7-4b97-b888-ae5aff721a80" colab={}
a = 1
b = 0
if a == 1:
if b == 0:
print("YES a==1 and b==0")
# + [markdown] colab_type="text" hidden=true id="K-DvcF-A1DAN"
# ## 4.12 端末からの入力
# + [markdown] colab_type="text" hidden=true id="924BBEWI1DAN"
# #### 演習20. `ex3.py`を改造して端末から平方根を求める数値を入力するようにしなさい。
# + colab_type="code" hidden=true id="Gn5wmk7Z1DAP" outputId="ea833ae9-0f6c-4503-dd1c-787fed0e8ce5" colab={}
# xの平方根を求める
x = float(input("Enter number "))
#
rnew = x
#
diff = rnew - x/rnew
if (diff < 0):
diff = -diff
while (diff > 1.0E-6):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
diff = r1 - r2
if (diff < 0):
diff = -diff
# + [markdown] colab_type="text" hidden=true id="rFgBk4li1DAR"
# ## 4.13 エラーへの対処
# + [markdown] colab_type="text" hidden=true id="XsZZyGEF1DAS"
# プログラム12 入力を得て検査するプログラム(`inputcheck.py`)
# + colab_type="code" hidden=true id="razW4MHE1DAS" outputId="5de5eb1b-b9e8-40cf-8267-714039822303" colab={"base_uri": "https://localhost:8080/", "height": 634}
while True:
x = input("正の数値を入力してください ")
try:
x = float(x)
except ValueError:
print(x, "は数値に変換できません")
continue
except:
print("予期していないエラーです")
exit()
if (x <= 0):
print(x, "は正の数値ではありません")
continue
# 以下は正しい入力が得られた時の処理
print(x)
# + [markdown] colab_type="text" hidden=true id="BTbbhzxh1DAV"
# ## 4.14 Pythonでの数学関数
# + [markdown] colab_type="text" hidden=true id="80knfC_z1DAV"
# 演習22. 上の例に従ってPython Shellで`math`モジュールを使ってみてください
# + colab_type="code" hidden=true id="KHbU_k841DAW" outputId="f8639a3d-0d56-41d6-8c2b-d46bafc405ca" colab={}
import math
pi = math.pi
print(pi)
sin = math.sin(pi/2)
print(sin)
# + [markdown] colab_type="text" hidden=true id="ZRcWyA0_1DAY"
# ## 4.15 数値を表示する際のフォーマット指定
# + colab_type="code" hidden=true id="guBdMZVI1DAY" outputId="21e7184f-a4e3-4a35-8d8e-227abc0b14e1" colab={}
c = 2.99792458E8
na = 6.02214076E23
form = '光速は{0:12.8g} m/s, アボガドロ数は {1:12.8g} mol**(-1) です'
print(form.format(c, na))
# + [markdown] colab_type="text" hidden=true id="F-REaiNG1DAa"
# ## 4.16 力試し
# + [markdown] colab_type="text" hidden=true id="bYRRIAQk1DAa"
# #### 演習23. `inputcheck.py`, `ex3.py`を組み合わせて以下の条件を満たす平方根を求めるプログラムを作成しなさい。
# + [markdown] colab_type="text" hidden=true id="F_5r-QJ91DAa"
# 1. 平方根を求める数を繰り返し端末から入力できるようにすること。
# 2. 平方根を求める数の入力が数値に変換できない場合は、その旨を示して、次の入力を求めること。
# 3. 平方根を求める数が0以下の場合は、その旨を示して、次の入力を求めること。
# できれば以下にも挑戦すること
# 5. 端末からの入力が"end"という文字列なら終了すること。
# 6. 計算精度を絶対精度ではなく、相対精度で$10^{-6}$とすること。これについて大きな数や小さな数(例えば$10^{10}$や$10^{-10}$)の平方根を求め、結果を確認すること。
# + colab_type="code" hidden=true id="xvP2nIuO1DAb" outputId="da70fa2b-907f-4be7-a348-a48a2412e4a0" colab={}
while True:
x = input("平方根を求める数 ")
if (x == 'end'):
break
try:
x = float(x)
except ValueError:
print(x, "は数値に変換できません")
continue
except:
print("予期していないエラーです")
exit()
if (x <= 0):
print(x, "は正の数値ではありません")
continue
# xの平方根を求める
x = float(x)
#
rnew = x
#
diff = rnew - x/rnew
if (diff < 0):
diff = abs(diff)
while (diff > 1.0E-6):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
form = '{0:1.6g}\t{1:1.6g}\t{2:1.6g}'
print(form.format(r1,rnew,r2))
diff = r1 - r2
if (diff < 0):
diff = abs(diff)
# + colab_type="code" hidden=true id="MWrlnHKZ1DAc" outputId="175d61e2-6f1b-43f2-fa8c-d735e9d5a057" colab={}
0.0000000001**(1/2)
# + [markdown] colab_type="text" heading_collapsed=true id="OAJHkHYS1DAf"
# # 5. 関数を使った処理のカプセル化
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="tfcD8T1a1DAf"
# ## 5.3 関数`square_root()`を実装する
# + [markdown] colab_type="text" hidden=true id="s4lYqeKm1DAg"
# プログラム13 関数`square_root()`の実装
# + colab_type="code" hidden=true id="ZrBm4Qdc1DAg" outputId="f9caf84a-7de9-4e13-b538-7f87f48cf70c" colab={"base_uri": "https://localhost:8080/", "height": 119}
#
def square_root(x):
'引数xの平方根を求める' #docstring
rnew = x
#
diff = rnew - x/rnew
if (diff < 0):
diff = abs(diff)
while (diff > 1.0E-6):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
diff = r1 - r2
if (diff < 0):
diff = abs(diff)
return rnew
# ここからメインプログラム
v = 2
r = square_root(v)
print("結果は ",r)
# + colab_type="code" hidden=true id="5spbPYzw1DAi" outputId="18cc6b20-fe0b-4fef-e8ba-62739a3984c5" colab={}
help(square_root)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="IOP2DnsH1DAj"
# #### 演習24. 繰り返し平方根を求めるプログラムを関数`square_root()`を定義して利用する形に書き換えなさい。
# + colab_type="code" hidden=true id="jTUYJGut1DAk" outputId="9163daf0-8928-44e2-eb4c-7862d4c0cca0" colab={}
#
def square_root(x):
'引数xの平方根を求める' #docstring
rnew = x
#
diff = rnew - x/rnew
if (diff < 0):
diff = abs(diff)
while (diff > 1.0E-6):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
diff = r1 - r2
if (diff < 0):
diff = abs(diff)
return rnew
while True:
v = input("平方根を求める数 ")
try:
v = float(v)
except ValueError:
print(v, "は数値に変換できません")
continue
except:
print("予期していないエラーです")
exit()
if (v <= 0):
print(v, "は正の数値ではありません")
continue
if (v > 0):
r = square_root(v)
print(r)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="XITdSW3l1DAl"
# #### 演習25. 関数`get_positive_numeral()`も構成し、繰り返し平方根を求めるプログラムをこれと`square_root()`を利用する形に書き換えなさい。
# + colab_type="code" hidden=true id="Sfm3f57t1DAm" outputId="967752ee-fa54-48a2-b5c7-e0f3aa788d7e" colab={}
def square_root(x):
'引数xの平方根を求める' #docstring
rnew = x
#
diff = rnew - x/rnew
if (diff < 0):
diff = abs(diff)
while (diff > 1.0E-6):
r1 = rnew
r2 = x/r1
rnew = (r1 + r2)/2
print(r1, rnew, r2)
diff = r1 - r2
if (diff < 0):
diff = abs(diff)
return rnew
def get_positive_numeral():
'端末から入力xを得て正の数値かどうか検査する'
x = input("平方根を求める数 ")
try:
x = float(x)
except ValueError:
print(x, "は数値に変換できません")
return ValueError
except:
print("予期していないエラーです")
return ValueError
if (x <= 0):
print(x, "は正の数値ではありません")
return ValueError
if (x > 0):
return x
while True:
x = get_positive_numeral()
if (x == ValueError):
continue
r = square_root(x)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="gldgetN21DAn"
# ## 5.6 関数内の変数の扱い
# + colab_type="code" hidden=true id="ox0D03ZH1DAo" outputId="b76f0c73-82ca-4236-a294-1b90480e56d5" colab={}
a = 10
b = 0
def f():
global b #グローバル宣言
c = a*a
b = c #グローバ宣言した変数は代入可能
f()
print(b,a)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="8DcS3wgm1DAp"
# ## 5.7 関数の利用パターン
# + colab_type="code" hidden=true id="1_foEZe-1DAp" outputId="cdbc2c8f-31ee-45ac-832c-72d1a9622300" colab={}
a = 0
def f():
global a
a = a + 1
def g(x):
x[0] = 0
f()
print(a)
b = [1,2,3]
g(b)
print(b)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="d9Lb-NTO1DAq"
# ## 5.8 関数の呼び出しと関数オブジェクトの引き渡し
# + colab_type="code" hidden=true id="29GNl3mK1DAr" outputId="ba4c4927-738e-4085-8dbd-63179bcbe4a1" colab={}
def f():
print("f says Hello")
# 関数を引数でもらって実行する関数
def F(y):
print("In F, ", end="") # `end="`末尾に出力を追加
y()
# fを実行
f()
# fをFに渡してFを実行
F(f)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="LMJmxAVP1DAs"
# ## 5.9 デフォルト引数値とキーワード引数
# + colab_type="code" hidden=true id="EA9YlyiF1DAs" outputId="f6f16ed2-ef3c-4a9b-fb2d-e9be658e3ee3" colab={}
def f(a, b=2, c=3):
return a+b+c
print(f(1,1,1))
print(f(1))
print(f(1,c=2))
# + [markdown] colab_type="text" heading_collapsed=true id="mMZYq42f1DAu"
# # 6. Turtleで遊ぶ
# + colab_type="code" hidden=true id="GJ-2kRFK1h6d" outputId="69ee4f83-7c59-486c-e9e9-351c658b10f6" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !pip3 install ColabTurtle
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="kyXUABxg1DAu"
# ## 6.4 使ってみよう
# + [markdown] colab_type="text" hidden=true id="543d653_1DAu"
# プログラム14 turtleを使う例(`turtle.py`という名前で保存してはいけない)
# + colab_type="code" hidden=true id="giv866ad1DAv" outputId="617deb00-0716-49c2-b4f4-87dad7883354" colab={"base_uri": "https://localhost:8080/", "height": 706}
from ColabTurtle.Turtle import*
initializeTurtle()
forward(100)
left(90)
forward(100)
left(90)
forward(100)
left(90)
forward(100)
left(90)
done()
# + [markdown] colab_type="text" hidden=true id="wkLdIVtU1DAz"
# #### 演習26. 以下のプログラムを完成させて正n角形を各プログラムを作成してください。
# プログラム15 n角形を描くプログラム(未完成)
# + colab_type="code" hidden=true id="0foew3SD1DAz" outputId="84fe4e97-bb59-48bb-dbf1-1cec206daf0c" colab={"base_uri": "https://localhost:8080/", "height": 521}
from ColabTurtle.Turtle import*
initializeTurtle()
n = 5
for i in range(n):
forward(100)
left(360/n)
# + [markdown] colab_type="text" hidden=true id="sdqfuwYx1DA1"
# #### 演習27. 星形はどうやって描けばいいでしょうか
# + colab_type="code" hidden=true id="9CWZXKj51DA1" outputId="9821369a-af95-499e-f150-ad5c911897b6" colab={"base_uri": "https://localhost:8080/", "height": 521}
from ColabTurtle.Turtle import*
initializeTurtle()
n = 5
for i in range(n):
forward(100)
left(360/n*2)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="ZmHxXy5G1DA3"
# ## 6.6 複数のタートルを動かす
# + [markdown] colab_type="text" hidden=true id="mLNdzO5G1DA3"
# ### 6.6.1 プログラム例
# + colab_type="code" hidden=true id="wagsBG5B1DA4" outputId="b5f993c1-70ea-474a-b41b-9a50284aeb3d" colab={"base_uri": "https://localhost:8080/", "height": 521}
from ColabTurtle.Turtle import*
initializeTurtle(10)
t1 = Turtle
t2 = Turtle
t1.color('red')
t2.color('blue')
for i in range(180):
t1.forward(5)
t2.forward(3)
t1.left(2)
t2.left(2)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="6SYRj-Z31DA5"
# ## 6.7 作品作りのためのヒント
# + [markdown] colab_type="text" hidden=true id="KGeJQh6E1DA6"
# ### 6.7.1 マウスクリックに応答する
# + [markdown] colab_type="text" hidden=true id="nDOu21oQ1DA7"
# プログラム16 タートルグラフィクスでのマウスクリックへの応答
# + colab_type="code" hidden=true id="csNpNX-11DA7" outputId="0aa3b24e-11c4-4b4f-f1ee-3404109ec29a" colab={}
from turtle import*
def come(x,y):
(xx,yy) = pos()
newx = ((xx+x)/2,(yy+y)/2)
print(x,y)
doto(newxy)
onscreenclick(come)
done()
# + [markdown] colab_type="text" hidden=true id="xs7MdHi31DA8"
# ### 6.7.2 座標を角度に変換する
# + colab_type="code" hidden=true id="dbeR4VAq1DA9" colab={}
from turtle import*
import math
y = 2
x = 1
angle = math.atan2(y,x)*180/math.pi
forward(100)
left(angle)
forward(100)
done()
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="dGAXTx1zcqxF"
# ## 6.9 課題 Turtleの作品制作
# + [markdown] colab_type="text" hidden=true id="e1ba8EyqcqxF"
# プログラム17 `random_turtle.py`
# + colab_type="code" hidden=true id="kZbKW6UCcqxF" outputId="2a12c8ba-1047-4ad7-d292-12bb7509a5df" colab={}
from turtle import*
import random
# 乱数を使うのでrandomモジュールもインポート
# 実行を停止するための変数(フラッグ)
stop_flag = False
# マウスがクリックされたときの関数、引数x, yをとるように
# しないといけないが、使わない
# 実行停止フラグをTrueにする
def clicked(x,y):
global stop_flag
stop_flag = True
#
# マウスがクリックされたときの動作を指定、clicked関数を
# 呼び出す
#
onscreenclick(clicked)
speed(0)
while(not stop_flag):
# -90度から90度の範囲でランダムに向きを変える
left(random.randint(-90,90))
forward(random.randint(5,15))
# タートルの位置が原点から一定の距離を超えれば、戻る
if (position()[0]**2+position()[1]**2 > 200**2):
forward(random.randint(-30,-20))
# + [markdown] colab_type="text" hidden=true id="11PS4Re5cqxH"
# プログラム18 `detour.py`
# + colab_type="code" hidden=true id="vuwNxcUYcqxH" colab={}
from turtle import *
def detour(L):
if L < 10:
forward(L)
else:
LL = L/3
detour(LL)
left(60)
detour(LL)
right(120)
detour(LL)
left(60)
detour(LL)
for i in range(6):
detour(100)
left(60)
# + [markdown] colab_type="text" hidden=true id="CFRUEpGTcqxI"
# プログラム19 `turtle-tree.py`
# + colab_type="code" hidden=true id="ED5TK3ercqxI" colab={}
from turtle import *
# 再帰的に木を描く
def tree(n):
# 引数が1以下なら5歩すすむ
if n <= 1:
forward(5)
else:
# 引数は1より大きいとき
# 引数の値に応じて前進(幹)
forward(5*(1.1**n))
# 今の位置と向きを記録
xx = pos()
h = heading()
# 左へ30度回転
left(30)
# 大きさn-2で木を描く(左の枝)
tree(n-2)
# ペンを上げて軌跡を残さない
up()
# 先に記録した位置(幹の先端)に戻る
setpos(xx)
setheading(h)
# ペンを降ろす
down()
# 右へ15度
right(15)
# 大きさn-1で木を描く(右の枝)
tree(n-1)
# ペンを上げてもどる
up()
setpos(xx)
setheading(h)
# ペンを降ろす
down()
# 時間がかかるので最も早い描画
speed(0)
# 大きさ12の木を描く
tree(12)
# + colab_type="code" heading_collapsed=true id="sKd3CRJm1DA-" colab={}
# 7. Tkinterで作るGUIアプリケーション(1)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="JksgHymXcqxL"
# ## 7.5 tkinterの例題 (`tkdemo-2term.py`)
# + colab_type="code" hidden=true id="c05Kwmn-cqxM" colab={}
import tkinter as tk
# 計算機能のための変数とイベント用の関数定義
# 2項演算のモデル
# 入力中の数字
current_number = 0
# 第一項
first_term = 0
# 第二項
second_term = 0
# 結果
result = 0
def do_plus():
"+キーが押されたときの計算動作、第一項の設定と入力中の数字のクリア"
global current_number
global first_term
first_term = current_number
current_number = 0
def do_eq():
"=キーが押されたときの計算動作、第二項の設定加算の実施、入力中の数字のクリア"
global second_term
global result
global current_number
second_term = current_number
result = first_term + second_term
current_number = 0
# 数字キーのcall Back関数
def key1():
key(1)
def key2():
key(2)
def key3():
key(3)
def key4():
key(4)
def key5():
key(5)
def key6():
key(6)
def key7():
key(7)
def key8():
key(8)
def key9():
key(9)
def key0():
key(0)
# 数字キーを一括処理する関数
def key(n):
global current_number
current_number = current_number * 10 + n
show_number(current_number)
def clear():
global current_number
current_number = 0
show_number(current_number)
def plus():
do_plus()
show_number(current_number)
def eq():
do_eq()
show_number(result)
def show_number(num):
e.delete(0,tk.END)
e.insert(0,str(num))
# tkinterでの画面の構成
root = tk.Tk()
f = tk.Frame(root)
f.grid()
# ウィジェットの作成
b1 = tk.Button(f,text='1', command=key1)
b2 = tk.Button(f,text='2', command=key2)
b3 = tk.Button(f,text='3', command=key3)
b4 = tk.Button(f,text='4', command=key4)
b5 = tk.Button(f,text='5', command=key5)
b6 = tk.Button(f,text='6', command=key6)
b7 = tk.Button(f,text='7', command=key7)
b8 = tk.Button(f,text='8', command=key8)
b9 = tk.Button(f,text='9', command=key9)
b0 = tk.Button(f,text='0', command=key0)
bc = tk.Button(f,text='C', command=clear)
bp = tk.Button(f,text='+', command=plus)
be = tk.Button(f,text='=', command=eq)
# Grid型ジオメトリマネージャによるウィジェットの
# 割付
b1.grid(row=3,column=0)
b2.grid(row=3,column=1)
b3.grid(row=3,column=2)
b4.grid(row=2,column=0)
b5.grid(row=2,column=1)
b6.grid(row=2,column=2)
b7.grid(row=1,column=0)
b8.grid(row=1,column=1)
b9.grid(row=1,column=2)
b0.grid(row=4,column=0)
bc.grid(row=1,column=3)
be.grid(row=4,column=3)
bp.grid(row=2,column=3)
# 数値を表示するウィジェット
e = tk.Entry(f)
e.grid(row=0,column=0,columnspan=4)
clear()
# ここからGUIがスタート
root.mainloop()
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="qgthq7cqcqxO"
# ## 7.8 `lambda`(λ)表現を使ったCall Back関数の記述
# + [markdown] colab_type="text" hidden=true id="uOJhBcGfcqxO"
# プログラム21 `lambda`式を使った引数付きコールバック関数の設定
# + colab_type="code" hidden=true id="RcakvIV-cqxP" colab={}
import tkinter as tk
# 計算機能のための変数とイベント用の関数定義
# 2項演算のモデル
# 入力中の数字
current_number = 0
# 第一項
first_term = 0
# 第二項
second_term = 0
# 結果
result = 0
def do_plus():
"+キーが押されたときの計算動作、第一項の設定と入力中の数字のクリア"
global current_number
global first_term
first_term = current_number
current_number = 0
def do_eq():
"=キーが押されたときの計算動作、第二項の設定加算の実施、入力中の数字のクリア"
global second_term
global result
global current_number
second_term = current_number
result = first_term + second_term
current_number = 0
# 数字キーを一括処理する関数
def key(n):
global current_number
current_number = current_number * 10 + n
show_number(current_number)
def clear():
global current_number
current_number = 0
show_number(current_number)
def plus():
do_plus()
show_number(current_number)
def eq():
do_eq()
show_number(result)
def show_number(num):
e.delete(0,tk.END)
e.insert(0,str(num))
# tkinterでの画面の構成
root = tk.Tk()
f = tk.Frame(root)
f.grid()
# ウィジェットの作成
b1 = tk.Button(f,text='1', command=lambda:key(1))
b2 = tk.Button(f,text='2', command=lambda:key(2))
b3 = tk.Button(f,text='3', command=lambda:key(3))
b4 = tk.Button(f,text='4', command=lambda:key(4))
b5 = tk.Button(f,text='5', command=lambda:key(5))
b6 = tk.Button(f,text='6', command=lambda:key(6))
b7 = tk.Button(f,text='7', command=lambda:key(7))
b8 = tk.Button(f,text='8', command=lambda:key(8))
b9 = tk.Button(f,text='9', command=lambda:key(9))
b0 = tk.Button(f,text='0', command=lambda:key(0))
bc = tk.Button(f,text='C', command=clear)
bp = tk.Button(f,text='+', command=plus)
be = tk.Button(f,text='=', command=eq)
# Grid型ジオメトリマネージャによるウィジェットの
# 割付
b1.grid(row=3,column=0)
b2.grid(row=3,column=1)
b3.grid(row=3,column=2)
b4.grid(row=2,column=0)
b5.grid(row=2,column=1)
b6.grid(row=2,column=2)
b7.grid(row=1,column=0)
b8.grid(row=1,column=1)
b9.grid(row=1,column=2)
b0.grid(row=4,column=0)
bc.grid(row=1,column=3)
be.grid(row=4,column=3)
bp.grid(row=2,column=3)
# 数値を表示するウィジェット
e = tk.Entry(f)
e.grid(row=0,column=0,columnspan=4)
clear()
# ここからGUIがスタート
root.mainloop()
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="2niRTJCwcqxQ"
# ## 7.9 ウィジェットの体裁の調整
# + [markdown] colab_type="text" hidden=true id="e-wcvdRXcqxR"
# #### 演習28. ウィジェットの体裁の調整
# + [markdown] colab_type="text" hidden=true id="ves79yWEcqxR"
# - 足し算電卓のフォントサイズ、ウィジェットの色を以下のように設定してください。背景色をFrameは`#ffffc0`(薄黄色)、数字キーは白、クリアキーは赤、+、=キーは緑にする。
# - ボタンの大きさは2(文字分)にする。
# - ボタンとエントリーのフォントとサイズは('Helvetica',14)にする。
# + colab_type="code" hidden=true id="bXUjXQMicqxS" colab={}
import tkinter as tk
# 計算機能のための変数とイベント用の関数定義
# 2項演算のモデル
# 入力中の数字
current_number = 0
# 第一項
first_term = 0
# 第二項
second_term = 0
# 結果
result = 0
def do_plus():
"+キーが押されたときの計算動作、第一項の設定と入力中の数字のクリア"
global current_number
global first_term
first_term = current_number
current_number = 0
def do_eq():
"=キーが押されたときの計算動作、第二項の設定加算の実施、入力中の数字のクリア"
global second_term
global result
global current_number
second_term = current_number
result = first_term + second_term
current_number = 0
# 数字キーを一括処理する関数
def key(n):
global current_number
current_number = current_number * 10 + n
show_number(current_number)
def clear():
global current_number
current_number = 0
show_number(current_number)
def plus():
do_plus()
show_number(current_number)
def eq():
do_eq()
show_number(result)
def show_number(num):
e.delete(0,tk.END)
e.insert(0,str(num))
# tkinterでの画面の構成
root = tk.Tk()
f = tk.Frame(root,bg='#ffffc0')
f.grid()
# ウィジェットの作成
b1 = tk.Button(f,text='1', command=lambda:key(1),font=('Helvetica',14),width=2,bg='#ffffff')
b2 = tk.Button(f,text='2', command=lambda:key(2),font=('Helvetica',14),width=2,bg='#ffffff')
b3 = tk.Button(f,text='3', command=lambda:key(3),font=('Helvetica',14),width=2,bg='#ffffff')
b4 = tk.Button(f,text='4', command=lambda:key(4),font=('Helvetica',14),width=2,bg='#ffffff')
b5 = tk.Button(f,text='5', command=lambda:key(5),font=('Helvetica',14),width=2,bg='#ffffff')
b6 = tk.Button(f,text='6', command=lambda:key(6),font=('Helvetica',14),width=2,bg='#ffffff')
b7 = tk.Button(f,text='7', command=lambda:key(7),font=('Helvetica',14),width=2,bg='#ffffff')
b8 = tk.Button(f,text='8', command=lambda:key(8),font=('Helvetica',14),width=2,bg='#ffffff')
b9 = tk.Button(f,text='9', command=lambda:key(9),font=('Helvetica',14),width=2,bg='#ffffff')
b0 = tk.Button(f,text='0', command=lambda:key(0),font=('Helvetica',14),width=2,bg='#ffffff')
bc = tk.Button(f,text='C', command=clear,font=('Helvetica',14),width=2,bg='#ff0000')
bp = tk.Button(f,text='+', command=plus,font=('Helvetica',14),width=2,bg='#00ff00')
be = tk.Button(f,text='=', command=eq,font=('Helvetica',14),width=2,bg='#00ff00')
# Grid型ジオメトリマネージャによるウィジェットの
# 割付
b1.grid(row=3,column=0)
b2.grid(row=3,column=1)
b3.grid(row=3,column=2)
b4.grid(row=2,column=0)
b5.grid(row=2,column=1)
b6.grid(row=2,column=2)
b7.grid(row=1,column=0)
b8.grid(row=1,column=1)
b9.grid(row=1,column=2)
b0.grid(row=4,column=0)
bc.grid(row=1,column=3)
be.grid(row=4,column=3)
bp.grid(row=2,column=3)
# 数値を表示するウィジェット
e = tk.Entry(f)
e.grid(row=0,column=0,columnspan=4)
clear()
# ここからGUIがスタート
root.mainloop()
# + [markdown] colab_type="text" hidden=true id="dxg-jY7TcqxU"
# #### 演習29. 電卓の四則演算への拡張
# + [markdown] colab_type="text" hidden=true id="e6gr-W_OcqxU"
# 足し算電卓を四則演算が可能なように拡張してください。ただし以下に留意すること。
# - ボタンの配置は適宜検討すること。
# - 割り算は0で割るエラーが発生する可能性があるので、第2項の数値が0の場合は何もしない。
# - 割り算の小数点以下は切り捨てる。Pythonで整数商を求める演算子は「`//`」です。
# + colab_type="code" hidden=true id="igQcHMVhcqxU" colab={}
import tkinter as tk
# 計算機能のための変数とイベント用の関数定義
# 2項演算のモデル
# 入力中の数字
current_number = 0
# 第一項
first_term = 0
# 第二項
second_term = 0
# 結果
result = 0
# 算術タイプ 0:plus,1:minus,2:multiple,3:divide
arithmetic_type = 0
def do_arith(type):
"算術キーが押されたときの計算動作、第一項の設定と入力中の数字のクリア、計算フラグをセット"
global current_number
global first_term
global arithmetic_type
first_term = current_number
arithmetic_type = type
current_number = 0
def do_eq():
"=キーが押されたときの計算動作、第二項の設定加算の実施、入力中の数字のクリア"
global second_term
global result
global current_number
global arithmetic_type
second_term = current_number
if arithmetic_type == 0:
result = first_term + second_term
elif arithmetic_type == 1:
if first_term >= secound_teerm:
result = first_term - second_term
else:
result = second_term - first_term
elif arithmetic_type == 2:
result = first_term * second_term
elif arithmetic_type == 3:
if second_term != 0:
result = first_term // second_term
else:
result = 0
current_number = 0
# 数字キーを一括処理する関数
def key(n):
global current_number
current_number = current_number * 10 + n
show_number(current_number)
def clear():
global current_number
current_number = 0
show_number(current_number)
def plus():
do_arith(0)
show_number(current_number)
def minus():
do_arith(1)
show_number(current_number)
def multiple():
do_arith(2)
show_number(current_number)
def divide():
do_arith(3)
show_number(current_number)
def eq():
do_eq()
show_number(result)
def show_number(num):
e.delete(0,tk.END)
e.insert(0,str(num))
# tkinterでの画面の構成
root = tk.Tk()
f = tk.Frame(root)
f.grid()
# ウィジェットの作成
b1 = tk.Button(f,text='1', command=lambda:key(1))
b2 = tk.Button(f,text='2', command=lambda:key(2))
b3 = tk.Button(f,text='3', command=lambda:key(3))
b4 = tk.Button(f,text='4', command=lambda:key(4))
b5 = tk.Button(f,text='5', command=lambda:key(5))
b6 = tk.Button(f,text='6', command=lambda:key(6))
b7 = tk.Button(f,text='7', command=lambda:key(7))
b8 = tk.Button(f,text='8', command=lambda:key(8))
b9 = tk.Button(f,text='9', command=lambda:key(9))
b0 = tk.Button(f,text='0', command=lambda:key(0))
bc = tk.Button(f,text='C', command=clear)
bp = tk.Button(f,text='+', command=plus)
bm = tk.Button(f,text='-', command=minus)
bt = tk.Button(f,text='*', command=multiple)
bd = tk.Button(f,text='/', command=divide)
be = tk.Button(f,text='=', command=eq,height=2)
# Grid型ジオメトリマネージャによるウィジェットの
# 割付
b1.grid(row=3,column=0)
b2.grid(row=3,column=1)
b3.grid(row=3,column=2)
b4.grid(row=2,column=0)
b5.grid(row=2,column=1)
b6.grid(row=2,column=2)
b7.grid(row=1,column=0)
b8.grid(row=1,column=1)
b9.grid(row=1,column=2)
b0.grid(row=4,column=0)
bc.grid(row=1,column=3)
be.grid(row=3,column=3,rowspan=2)
bp.grid(row=2,column=3)
bm.grid(row=3,column=4)
bt.grid(row=2,column=4)
bd.grid(row=1,column=4)
# 数値を表示するウィジェット
e = tk.Entry(f)
e.grid(row=0,column=0,columnspan=4)
clear()
# ここからGUIがスタート
root.mainloop()
# + [markdown] colab_type="text" hidden=true id="3ea0AeNXcqxW"
# #### 演習30. 実際の電卓との差異
# + [markdown] colab_type="text" hidden=true id="Fk1QOTx3cqxW"
# 作成したプログラムと実際の電卓(や電卓アプリ)との動作の違いを探ってください。例えば=キーの代わりに+などの演算キーを押した場合の動作など。
# + [markdown] colab_type="text" hidden=true id="cy81baDvcqxW"
# =キーの代わりに+などの演算キーを押した場合, 実際の電卓ではそれまでの値について演算されるが, 作成したプログラムでは演算が行われずに第2項の値が変わる.
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="yFoLrt0ZcqxX"
# ## 7.11 `Frame`クラスを拡張する方式での実装法
# + [markdown] colab_type="text" hidden=true id="XSxB0mbwcqxX"
# プログラム22 `Frame`クラスを拡張するtkinterの実装法
# + colab_type="code" hidden=true id="57ECKX3-cqxY" colab={}
import tkinter as tk
# 計算機能のための変数とイベント用の関数定義
# Frameのサブクラスを使った実装例
# 2項演算のモデル
# 入力中の数字
current_number = 0
# 第一項
first_term = 0
# 第二項
second_term = 0
# 結果
result = 0
def do_plus():
"+キーが押されたときの計算動作、第一項の設定と入力中の数字のクリア"
global current_number
global first_term
first_term = current_number
current_number = 0
def do_eq():
"=キーが押されたときの計算動作、第二項の設定加算の実施、入力中の数字のクリア"
global second_term
global result
global current_number
second_term = current_number
result = first_term + second_term
current_number = 0
#
# tk.Frameを継承したMyFrameというクラスを作り
# その中でウィジェットやコールバック関数(メソッド)を
# 設定する。tkinterを使う定番
#
class MyFrame(tk.Frame):
#
# __init__はクラスオブジェクトを作る際の初期化メソッド
# アンダースコアはそれぞれ2つずつ
def __init__(self,master = None):
super().__init__(master)
# あとで参照しないウィジェットの作成、ローカル変数
b1 = tk.Button(self,text='1', command=lambda:self.key(1))
b2 = tk.Button(self,text='2', command=lambda:self.key(2))
b3 = tk.Button(self,text='3', command=lambda:self.key(3))
b4 = tk.Button(self,text='4', command=lambda:self.key(4))
b5 = tk.Button(self,text='5', command=lambda:self.key(5))
b6 = tk.Button(self,text='6', command=lambda:self.key(6))
b7 = tk.Button(self,text='7', command=lambda:self.key(7))
b8 = tk.Button(self,text='8', command=lambda:self.key(8))
b9 = tk.Button(self,text='9', command=lambda:self.key(9))
b0 = tk.Button(self,text='0', command=lambda:self.key(0))
bc = tk.Button(self,text='C', command=self.clear)
bp = tk.Button(self,text='+', command=self.plus)
be = tk.Button(self,text='=', command=self.eq)
# Grid型ジオメトリマネージャによるウィジェット割付
b1.grid(row=3,column=0)
b2.grid(row=3,column=1)
b3.grid(row=3,column=2)
b4.grid(row=2,column=0)
b5.grid(row=2,column=1)
b6.grid(row=2,column=2)
b7.grid(row=1,column=0)
b8.grid(row=1,column=1)
b9.grid(row=1,column=2)
b0.grid(row=4,column=0)
bc.grid(row=1,column=3)
be.grid(row=4,column=3)
bp.grid(row=2,column=3)
# 他のメソッドで参照する数値を表示するウィジェット、クラスオブジェクトの
# 変数として作成、頭にself.がつく
self.e = tk.Entry(self)
self.e.grid(row=0,column=0,columnspan=4)
# クラスの定義では
# メソッドの最初の引数はself,中でクラスオブジェクトの変数、
# メソッドはselfを付けて参照
#
def key(self,n):
global current_number
current_number = current_number * 10 + n
self.show_number(current_number)
def clear(self):
global current_number
current_number = 0
self.show_number(current_number)
def plus(self):
do_plus()
self.show_number(current_number)
def eq(self):
do_eq()
self.show_number(result)
def show_number(self, num):
self.e.delete(0,tk.END)
self.e.insert(0,str(num))
#
# ここからメインプログラム
#
root = tk.Tk()
f = MyFrame(root)
f.pack()
f.mainloop()
# + [markdown] colab_type="text" heading_collapsed=true id="cw2aanzqcqxZ"
# # 8. Tkinterで作るGUIアプリケーション(2)
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="bG6FdZXZcqxa"
# ## 8.4 tkinterを用いたアナログ時計プログラム
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="jd7rHsEbcqxa"
# ### 8.4.1 ソースコード
# + [markdown] colab_type="text" hidden=true id="b3jtXB66cqxa"
# プログラム23 tkinterでのアナログ時計
# + colab_type="code" hidden=true id="74IGMGArcqxb" colab={}
# tkinter canvasを使ったアナログ時計
#
import tkinter as tk
import math
import time
#
# Frameを拡張したクラス
#
class MyFrame(tk.Frame):
def __init__(self, master=None):
super().__init__(master)
#
# キャンバスの作成
#
self.size = 200
self.clock = tk.Canvas(self, width=self.size, height=self.size, background="white")
self.clock.grid(row=0, column=0)
#
# 文字盤の描画
#
self.font_size = int(self.size/15)
for number in range(1,12+1):
x = self.size/2 + math.cos(math.radians(number*360/12 - 90))*self.size/2*0.85
y = self.size/2 + math.sin(math.radians(number*360/12 - 90))*self.size/2*0.85
self.clock.create_text(x,y,text=str(number),fill="black",font=("",14))
#
# 日付表示をオンオフするボタンの作成
#
self.b = tk.Button(self, text="Show Date", font=("",14),command=self.toggle)
self.b.grid(row=1, column=0)
#
# 時刻の経過確認などの動作のためのインスタンス変数
#
self.sec = time.localtime().tm_sec
self.sec2 = time.localtime().tm_sec
self.min = time.localtime().tm_min
self.hour = time.localtime().tm_hour
self.start = True
self.show_date = False
self.toggled = True
#
# ボタンが押されたときのcall back
#
def toggle(self):
if self.show_date:
self.b.configure(text="show date")
else:
self.b.configure(text="hide date")
self.show_date = not self.show_date
self.toggled = True
#
# 変化する画面の描画
#
def display(self):
#
# 秒針の描画、最初(start==True)か秒が変わったとき
#
if self.sec != time.localtime().tm_sec or self.start:
self.sec = time.localtime().tm_sec
angle = math.radians(self.sec*360/60 - 90)
x0 = self.size/2 - math.cos(angle)*self.size/2*0.1
y0 = self.size/2 - math.sin(angle)*self.size/2*0.1
x = self.size/2 + math.cos(angle)*self.size/2*0.75
y = self.size/2 + math.sin(angle)*self.size/2*0.75
#
# 前の描画をタグで検索して消してから描画
#
self.clock.delete("SEC")
self.clock.create_line(x0,y0,x,y,width=1,fill="red",tag="SEC")
#
# 分針、時針の描画、1分毎、時針は分まで考慮
#
if self.min != time.localtime().tm_min or self.start:
self.min = time.localtime().tm_min
x0 = self.size/2
y0 = self.size/2
angle = math.radians(self.min*360/60 - 90)
x = self.size/2 + math.cos(angle)*self.size/2*0.65
y = self.size/2 + math.sin(angle)*self.size/2*0.65
self.clock.delete("MIN")
self.clock.create_line(x0,y0,x,y,width=3,fill="blue",tag="MIN")
self.hour = time.localtime().tm_hour
x0 = self.size/2
y0 = self.size/2
angle = math.radians((self.hour%12 + self.min/60)*360/12 - 90)
x = self.size/2 + math.cos(angle)*self.size/2*0.55
y = self.size/2 + math.sin(angle)*self.size/2*0.55
self.clock.delete("HOUR")
self.clock.create_line(x0,y0,x,y,width=3,fill="green",tag="HOUR")
self.start = False
#
# 日付の描画、秒が変わるか、ボタンが押されたとき
#
if self.sec2 != time.localtime().tm_sec or self.toggled:
self.sec2 = time.localtime().tm_sec
self.toggled = False
x = self.size/2
y = self.size/2 + 20
text = time.strftime('%Y/%m/%d %H:%M:%S')
self.clock.delete("TIME")
if self.show_date:
self.clock.create_text(x,y,text=text,font=("",12),fill="black",tag="TIME")
#
# 100ミリ秒後に再度呼び出す
#
self.after(100, self.display)
root = tk.Tk()
f = MyFrame(root)
f.pack()
f.display()
root.mainloop()
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="pC69n2vVcqxc"
# ### 8.4.2 このプログラムのポイント
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="YhSrf9bJcqxc"
# #### 演習32. アナログ時計の改造
# + [markdown] colab_type="text" hidden=true id="BYsykqKvcqxd"
# アナログ時計のプログラムについて以下の改造を加えなさい。
# 1. 日付の表示について、日付と時刻ではなく、日付と午前、午後を表示するようにしてください。
# 2. ボタンをもう一つ追加し、秒針の表示をする、しない、を切り替えるようにしてください。
# + colab_type="code" hidden=true id="79elX_Iycqxd" colab={}
# tkinter canvasを使ったアナログ時計
#
import tkinter as tk
import math
import time
#
# Frameを拡張したクラス
#
class MyFrame(tk.Frame):
def __init__(self, master=None):
super().__init__(master)
#
# キャンバスの作成
#
self.size = 250
self.clock = tk.Canvas(self, width=self.size, height=self.size, background="white")
self.clock.grid(row=0, column=0,columnspan=2)
#
# 文字盤の描画
#
self.font_size = int(self.size/15)
for number in range(1,12+1):
x = self.size/2 + math.cos(math.radians(number*360/12 - 90))*self.size/2*0.85
y = self.size/2 + math.sin(math.radians(number*360/12 - 90))*self.size/2*0.85
self.clock.create_text(x,y,text=str(number),fill="black",font=("",14))
#
# 日付表示をオンオフするボタンの作成
#
self.bd = tk.Button(self, text="Show Date", font=("",14),command=self.toggle_date)
self.bd.grid(row=1, column=0)
#
# 秒針を非表示にするボタンの作成
#
self.bs = tk.Button(self, text="hide sec.", font=("",14),command=self.toggle_sec)
self.bs.grid(row=1, column=1)
#
# 時刻の経過確認などの動作のためのインスタンス変数
#
self.sec = time.localtime().tm_sec
self.sec2 = time.localtime().tm_sec
self.min = time.localtime().tm_min
self.hour = time.localtime().tm_hour
self.start = True
self.show_date = False
self.show_sec = True
self.toggled_date = True
self.toggled_sec = True
#
# ボタンが押されたときのcall back
#
def toggle_date(self):
if self.show_date:
self.bd.configure(text="show date")
else:
self.bd.configure(text="hide date")
self.show_date = not self.show_date
self.toggled_date = True
def toggle_sec(self):
if self.show_sec:
self.bs.configure(text="show sec.")
else:
self.bs.configure(text="hide sec.")
self.show_sec = not self.show_sec
self.toggled_sec = True
#
# 変化する画面の描画
#
def display(self):
#
# 秒針の描画、最初(start==True)か秒が変わったとき、ボタンが押されたとき
#
if self.sec != time.localtime().tm_sec or self.start or self.toggled_sec:
self.sec = time.localtime().tm_sec
angle = math.radians(self.sec*360/60 - 90)
x0 = self.size/2 - math.cos(angle)*self.size/2*0.1
y0 = self.size/2 - math.sin(angle)*self.size/2*0.1
x = self.size/2 + math.cos(angle)*self.size/2*0.75
y = self.size/2 + math.sin(angle)*self.size/2*0.75
#
# 前の描画をタグで検索して消してから描画
#
self.clock.delete("SEC")
if self.show_sec:
self.clock.create_line(x0,y0,x,y,width=1,fill="red",tag="SEC")
#
# 分針、時針の描画、1分毎、時針は分まで考慮
#
if self.min != time.localtime().tm_min or self.start:
self.min = time.localtime().tm_min
x0 = self.size/2
y0 = self.size/2
angle = math.radians(self.min*360/60 - 90)
x = self.size/2 + math.cos(angle)*self.size/2*0.65
y = self.size/2 + math.sin(angle)*self.size/2*0.65
self.clock.delete("MIN")
self.clock.create_line(x0,y0,x,y,width=3,fill="blue",tag="MIN")
self.hour = time.localtime().tm_hour
x0 = self.size/2
y0 = self.size/2
angle = math.radians((self.hour%12 + self.min/60)*360/12 - 90)
x = self.size/2 + math.cos(angle)*self.size/2*0.55
y = self.size/2 + math.sin(angle)*self.size/2*0.55
self.clock.delete("HOUR")
self.clock.create_line(x0,y0,x,y,width=3,fill="green",tag="HOUR")
self.start = False
#
# 日付の描画、秒が変わるか、ボタンが押されたとき
#
if self.sec2 != time.localtime().tm_sec or self.toggled_date:
self.sec2 = time.localtime().tm_sec
self.toggled_date = False
x = self.size/2
y = self.size/2 + 20
text = time.strftime('%Y/%m/%d %P%I:%M:%S')
self.clock.delete("TIME")
if self.show_date:
self.clock.create_text(x,y,text=text,font=("",12),fill="black",tag="TIME")
#
# 100ミリ秒後に再度呼び出す
#
self.after(100, self.display)
root = tk.Tk()
f = MyFrame(root)
f.pack()
f.display()
root.mainloop()
# + colab_type="code" hidden=true id="RwNNBSY7cqxf" colab={}
# + [markdown] colab_type="text" heading_collapsed=true id="Km02e6ghcyvk"
# # 9. クラス
# + [markdown] colab_type="text" hidden=true id="S02UqFyVc90x"
# ## 9.3 Pythonでのクラスの書き方、使い方
# + [markdown] colab_type="text" heading_collapsed=true hidden=true id="WqeHLEKRdFbS"
# ### 9.3.1 ソースコード
# + [markdown] colab_type="text" hidden=true id="dX5povDRdK8Q"
# プログラム24 CUI型電卓プログラム
# + colab_type="code" hidden=true id="CW7lqf5Oc5c2" outputId="4dc64392-608e-46df-ca18-7ce914ba4696" colab={"base_uri": "https://localhost:8080/", "height": 600}
class Dentaku():
def __init__(self):
self.first_term = 0
self.second_term = 0
self.result = 0
self_operation = "+"
def do_operation(self):
if self.operation == "+":
self.result = self.first_term + self.second_term
elif self.operation == "-":
self.result = self.first_term - self.second_term
# ここからメインプログラム
dentaku = Dentaku()
while True:
f = int(input("First term "))
dentaku.first_term = f
o = input("Operation ")
dentaku.operation = o
s = int(input("Second term "))
dentaku.second_term = s
dentaku.do_operation()
r = dentaku.result
print("Result ", r)
# + [markdown] colab_type="text" hidden=true id="LJdIrjYxkN3O"
# #### 演習33. `Dentaku`クラスのオブジェクトを複数生成して利用するプログラムを作成してみてください。
# + colab_type="code" hidden=true id="NlD1MjALh1jK" outputId="ed96c795-677c-4736-d72d-d43c9f89c660" colab={"base_uri": "https://localhost:8080/", "height": 583}
class Dentaku():
def __init__(self):
self.first_term = 0
self.second_term = 0
self.result = 0
self_operation = "+"
def do_operation(self):
if self.operation == "+":
self.result = self.first_term + self.second_term
elif self.operation == "-":
self.result = self.first_term - self.second_term
# ここからメインプログラム
dentaku1 = Dentaku()
dentaku2 = Dentaku()
while True:
f = int(input("First term "))
dentaku1.first_term = f
dentaku2.first_term = f
dentaku1.operation = "+"
dentaku2.operation = "-"
s = int(input("Second term "))
dentaku1.second_term = s
dentaku2.second_term = s
dentaku1.do_operation()
dentaku2.do_operation()
r1 = dentaku1.result
r2 = dentaku2.result
print("Result ", r1, r2)
# + [markdown] colab_type="text" hidden=true id="U-4FB1eXmNBw"
# #### 演習 34. `Dentaku`クラスを乗算、除算も扱えるように拡張しなさい。ただし、除算は整数商でかまいません。
# + colab_type="code" hidden=true id="kXXYKrH4mMrD" outputId="5301c309-aa11-40d2-818b-ad125cf06d19" colab={"base_uri": "https://localhost:8080/", "height": 756}
class Dentaku():
def __init__(self):
self.first_term = 0
self.second_term = 0
self.result = 0
self_operation = "+"
def do_operation(self):
if self.operation == "+":
self.result = self.first_term + self.second_term
elif self.operation == "-":
self.result = self.first_term - self.second_term
elif self.operation == "*":
self.result = self.first_term * self.second_term
elif self.operation == "/":
self.result = self.first_term // self.second_term
# ここからメインプログラム
dentaku = Dentaku()
while True:
f = int(input("First term "))
dentaku.first_term = f
o = input("Operation ")
dentaku.operation = o
s = int(input("Second term "))
dentaku.second_term = s
dentaku.do_operation()
r = dentaku.result
print("Result ", r)
# + [markdown] colab_type="text" hidden=true id="wfLH7Q7Tn3dQ"
# #### 演習 35. tkinterで作成した電卓プログラムについて、`Dentaku`クラスを利用するように改造しなさい
# + colab_type="code" hidden=true id="AoSRocGol_jR" colab={}
import tkinter as tk
# 計算機能のための変数とイベント用の関数定義
# Frameのサブクラスを使った実装例
# 2項演算のモデル
# 入力中の数字
current_number = 0
class Dentaku():
def __init__(self):
self.first_term = 0
self.second_term = 0
self.result = 0
self_operation = "+"
def do_operation(self):
if self.operation == "+":
self.result = self.first_term + self.second_term
elif self.operation == "-":
self.result = self.first_term - self.second_term
elif self.operation == "*":
self.result = self.first_term * self.second_term
elif self.operation == "/":
self.result = self.first_term // self.second_term
#
# tk.Frameを継承したMyFrameというクラスを作り
# その中でウィジェットやコールバック関数(メソッド)を
# 設定する。tkinterを使う定番
#
class MyFrame(tk.Frame):
#
# __init__はクラスオブジェクトを作る際の初期化メソッド
# アンダースコアはそれぞれ2つずつ
def __init__(self,master = None):
super().__init__(master)
# あとで参照しないウィジェットの作成、ローカル変数
b1 = tk.Button(self,text='1', command=lambda:self.key(1))
b2 = tk.Button(self,text='2', command=lambda:self.key(2))
b3 = tk.Button(self,text='3', command=lambda:self.key(3))
b4 = tk.Button(self,text='4', command=lambda:self.key(4))
b5 = tk.Button(self,text='5', command=lambda:self.key(5))
b6 = tk.Button(self,text='6', command=lambda:self.key(6))
b7 = tk.Button(self,text='7', command=lambda:self.key(7))
b8 = tk.Button(self,text='8', command=lambda:self.key(8))
b9 = tk.Button(self,text='9', command=lambda:self.key(9))
b0 = tk.Button(self,text='0', command=lambda:self.key(0))
bc = tk.Button(self,text='C', command=self.clear)
bp = tk.Button(self,text='+', command=lambda:self.operation("+"))
bm = tk.Button(self,text='-', command=lambda:self.operation("-"))
bt = tk.Button(self,text='*', command=lambda:self.operation("*"))
bd = tk.Button(self,text='/', command=lambda:self.operation("/"))
be = tk.Button(self,text='=', command=self.eq)
# Grid型ジオメトリマネージャによるウィジェット割付
b1.grid(row=3,column=0)
b2.grid(row=3,column=1)
b3.grid(row=3,column=2)
b4.grid(row=2,column=0)
b5.grid(row=2,column=1)
b6.grid(row=2,column=2)
b7.grid(row=1,column=0)
b8.grid(row=1,column=1)
b9.grid(row=1,column=2)
b0.grid(row=4,column=0)
bc.grid(row=1,column=3)
be.grid(row=4,column=3)
bp.grid(row=2,column=3)
bm.grid(row=3,column=4)
bt.grid(row=2,column=4)
bd.grid(row=1,column=4)
# 他のメソッドで参照する数値を表示するウィジェット、クラスオブジェクトの
# 変数として作成、頭にself.がつく
self.e = tk.Entry(self)
self.e.grid(row=0,column=0,columnspan=4)
# クラスの定義では
# メソッドの最初の引数はself,中でクラスオブジェクトの変数、
# メソッドはselfを付けて参照
#
def key(self,n):
global current_number
current_number = current_number * 10 + n
self.show_number(current_number)
def clear(self):
global current_number
current_number = 0
self.show_number(current_number)
def operation(self,o):
global current_number
dentaku.operation = o
dentaku.first_term = current_number
self.show_number(current_number)
self.clear()
def eq(self):
global current_number
dentaku.second_term = current_number
dentaku.do_operation()
self.show_number(dentaku.result)
def show_number(self, num):
self.e.delete(0,tk.END)
self.e.insert(0,str(num))
#
# ここからメインプログラム
#
dentaku = Dentaku()
root = tk.Tk()
f = MyFrame(root)
f.pack()
f.mainloop()
# + [markdown] heading_collapsed=true hidden=true id="F51FON-ZJLIW" colab_type="text"
# ## 9.4 クラスの変数とアクセスの制限
# + [markdown] hidden=true id="D2ZsA56pJLIW" colab_type="text"
# プログラム25 クラス変数とインスタンス変数
# + hidden=true id="-YK_JsbxJLIW" colab_type="code" colab={} outputId="c03ab063-557b-4bc7-e188-61cde74df9b8"
# クラスの練習
class MyClass():
# 以下はクラス変数
a = "マイクラス"
__b = 0
# 以下は生成する際に呼ばれる関数, mydataの初期値を
# 引数で与える
def __init__(self, data):
# __numberはインスタンスの通し番号
self.__number = MyClass.__b
self.mydata = data
print("MyClass Object is created, number: ", self.__number)
# クラス変数を1増やす
MyClass.__b += 1
# 通し番号を表示するメソッド
def show_number(self):
print(self.__number)
#
# ここからメインプログラム
#
if __name__ == "__main__":
print("MyClassのクラス変数 a:", MyClass.a)
instance1 = MyClass(1)
instance2 = MyClass(10)
instance1.show_number()
instance2.show_number()
print("mydata of instance1: ", instance1.mydata)
print("mydata of instance2: ", instance2.mydata)
instance1.mydata += 1
instance2.mydata += 2
print("mydata of instance1: ", instance1.mydata)
print("mydata of instance2: ", instance2.mydata)
# + hidden=true id="CRFYXduHJLIX" colab_type="code" colab={} outputId="f7e7cd2c-f4a3-443d-9861-0d92439d5d08"
print(instance1.__number)
# + [markdown] heading_collapsed=true id="M9UBFihQJLIY" colab_type="text"
# # 10. リスト
# + [markdown] heading_collapsed=true hidden=true id="BGM5wvfXJLIZ" colab_type="text"
# ## 10.2 Python Shellを用いた学習
# + hidden=true id="XdEMFUJjJLIZ" colab_type="code" colab={} outputId="1b1eb053-cf08-4383-f894-8fcbf8390ec5"
a = [1, 2, 3]
for d in a:
print(d)
# + [markdown] heading_collapsed=true hidden=true id="d_z2dzIGJLIa" colab_type="text"
# ## 10.3 リストとは
# + hidden=true id="hRsqEtBlJLIa" colab_type="code" colab={} outputId="b3200ae6-f876-4858-9f6d-2e6dadb1a90b"
a = [5, 1, 3, 4]
print(a)
print(a[0])
print(a[2])
# + [markdown] hidden=true id="0ciXSg8sJLIb" colab_type="text"
# ## 10.4 リストの生成
# + [markdown] hidden=true id="ibCcc_GxJLIb" colab_type="text"
# ### 10.4.1 要素を指定した生成
# + hidden=true id="h5J5mFNmJLIc" colab_type="code" colab={} outputId="5a6605bd-f0bc-4ac8-f3b4-df94c81b1b86"
a = [5, 1, 3, 4]
print(a)
print(a[0])
b = ['三条', '四条', '五条', '七条']
print(b)
print(b[0])
c = 5
a = [c, 1, 3, 4]
print(a)
print(a[0])
# + [markdown] hidden=true id="2zbqCvAdJLId" colab_type="text"
# ### 10.4.2 `range()`との組みあわせ
# + hidden=true id="Y37quYxDJLId" colab_type="code" colab={} outputId="d95ce1ff-4807-43c9-e245-daf0c731aecd"
n = list(range(5))
print(n)
# + [markdown] heading_collapsed=true hidden=true id="gmXf8gJxJLIf" colab_type="text"
# ### 10.4.3 文字列からの生成
# + hidden=true id="z4-op_5zJLIf" colab_type="code" colab={} outputId="af54a89e-6e5e-47bd-d4ec-f56b215521e7"
s = list('abcde')
print(s)
# + hidden=true id="OHRuCTeyJLIg" colab_type="code" colab={} outputId="1b85949c-3644-46f7-802a-fd28728da045"
t = "a textbook of Python"
tlist = t.split()
print(tlist)
# + [markdown] heading_collapsed=true hidden=true id="jepBL5aRJLIh" colab_type="text"
# ## 10.5 リストの要素へのアクセス
# + hidden=true id="oVe6IbvUJLIh" colab_type="code" colab={} outputId="e4faaf8a-6187-4628-cca9-41419814c488"
a = [5, 1, 3, 4]
print(a[0])
a[1] = 2
print(a)
print(len(a))
# + [markdown] hidden=true id="xn9U4FJtJLIi" colab_type="text"
# ## 10.6 リストを操作する`for`文
# + [markdown] heading_collapsed=true hidden=true id="XFecmraWJLIj" colab_type="text"
# ### 10.6.1 リストの長さと`range`関数を組み合わせる方法
# + hidden=true id="_0ruStaZJLIj" colab_type="code" colab={} outputId="116f1bd1-f4da-4fa6-8236-6dfdaec92755"
a = [5, 1, 3, 4]
for i in range(len(a)):
print(i, a[i])
# + [markdown] heading_collapsed=true hidden=true id="gGMiXEe5JLIk" colab_type="text"
# ### 10.6.2 リストを`for`文で直接使う方法
# + hidden=true id="fivMXenVJLIk" colab_type="code" colab={} outputId="1a3a345d-ee3d-4b58-e01a-a19caea5d7ce"
a = [5, 1, 3, 4]
for d in a:
print(d)
# + [markdown] heading_collapsed=true hidden=true id="tx8qSzVJJLIm" colab_type="text"
# #### 演習36. 平均値を求める
# + hidden=true id="Z_6Dcp3OJLIm" colab_type="code" colab={} outputId="a602f25b-0f94-4cec-ca2b-652c37e0b222"
a = [5, 1, 3, 4]
sum = 0
for i in range(len(a)):
sum += a[i]
average = sum/len(a)
print(average)
# + [markdown] heading_collapsed=true hidden=true id="BAuiSTtsJLIn" colab_type="text"
# #### 演習37. リストを直接`for`文で利用する形に上のプログラムを書き換えなさい。
# + hidden=true id="AT5wM1wBJLIo" colab_type="code" colab={} outputId="736ae062-f628-4db2-959e-1135d32c133f"
a = [5, 1, 3, 4]
sum = 0
for d in a:
sum += d
average = sum/len(a)
print(average)
# + [markdown] heading_collapsed=true hidden=true id="MJoO9bP0JLIp" colab_type="text"
# ## 10.7 負の添え字とスライス
# + [markdown] heading_collapsed=true hidden=true id="aNz-hBAJJLIp" colab_type="text"
# ### 10.7.1 負の添え字
# + hidden=true id="Ix9O6Um8JLIq" colab_type="code" colab={} outputId="691f0215-3d5c-4f9f-e9c0-e70441b5da41"
a = [5, 1, 3, 4]
print(a[-1])
# + [markdown] heading_collapsed=true hidden=true id="000tzk6wJLIr" colab_type="text"
# ### 10.7.2 スライス
# + hidden=true id="OxqZnH3yJLIr" colab_type="code" colab={} outputId="302b0189-1862-4938-e93d-b4c408b86c87"
a = [5, 1, 3, 4]
b = a[1:3]
print(b)
# + [markdown] heading_collapsed=true hidden=true id="31QFQWjdJLIt" colab_type="text"
# ## 10.8 リストへの追加、結合
# + [markdown] heading_collapsed=true hidden=true id="bmChXEzAJLIt" colab_type="text"
# ### 10.8.1 `append`
# + hidden=true id="mB6HHuUTJLIt" colab_type="code" colab={} outputId="cadba085-df05-40af-8921-689dcff112bf"
a = [5, 1, 3, 4]
a.append(2)
print(a)
# + [markdown] heading_collapsed=true hidden=true id="NzIKmk76JLIv" colab_type="text"
# ### 10.8.2 `extend`
# + hidden=true id="G3VkhoFvJLIv" colab_type="code" colab={} outputId="93c37054-57a5-4488-8ae4-dc929d107d66"
a = [5, 1, 3, 4]
b = [2, 6]
a.extend(b)
print(a)
# + hidden=true id="Af6CYGSBJLIw" colab_type="code" colab={} outputId="f695ef25-8d6f-4bb5-f0b5-736e513aff2b"
a = [5, 1, 3, 4]
b = [2, 6]
a.append(b)
print(a)
# + [markdown] heading_collapsed=true hidden=true id="S9WlQfMgJLIx" colab_type="text"
# ## 10.9 リストのリスト
# + hidden=true id="hTfaXqSQJLIx" colab_type="code" colab={} outputId="b4693c2c-4ef1-4a2a-8235-b37bfc53a7c1"
a = [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
print(a)
print(a[0])
print(a[0][1])
# + hidden=true id="D4UQclEVJLIy" colab_type="code" colab={} outputId="08c550e1-9ea2-44c3-b88b-4cfe0fd4dfa2"
a = [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
sum = 0
for i in range(len(a)):
for j in range(len(a[i])):
sum += a[i][j]
print(sum)
# + hidden=true id="EUZMvS9xJLIz" colab_type="code" colab={} outputId="f2fde06c-d82b-4cc6-a7d1-408ca3280fd7"
a = [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
sum = 0
for row in a:
for element in row:
sum += element
print(sum)
# + [markdown] heading_collapsed=true hidden=true id="EOk1--mlJLI0" colab_type="text"
# ## 10.10 内包表記
# + hidden=true id="FA2KBuONJLI0" colab_type="code" colab={} outputId="656e1082-357b-4095-8af2-710a63b708b3"
a = []
for i in range(5):
a.append(i*i)
print(a)
# + hidden=true id="381sPZNvJLI1" colab_type="code" colab={} outputId="8eeb8cb1-0b8a-412b-b2cf-54993dd3d2d8"
a = [i*i for i in range(5)]
print(a)
# + [markdown] heading_collapsed=true hidden=true id="MGu9nforJLI3" colab_type="text"
# ## 10.11 リストの代入と複製
# + hidden=true id="BEBvtJYzJLI3" colab_type="code" colab={} outputId="2e86fc7d-96c1-451b-d20e-07aac7c6b272"
a = [1, 2, 3]
b = a
print(a)
print(b)
b[0] = 0
a[1] = 0
print(a)
print(b)
print(id(a),id(b))
# + hidden=true id="nj8AVzJlJLI4" colab_type="code" colab={} outputId="70500434-c23b-4a07-fb44-b80420e538c5"
a = [1, 2, 3]
b = a.copy()
print(a)
print(b)
b[0] = 0
a[1] = 0
print(a)
print(b)
print(id(a),id(b))
# + [markdown] heading_collapsed=true hidden=true id="Y2AJJN46JLI6" colab_type="text"
# ## 10.12 イミュータブルとミュータブル
# + hidden=true id="PWPZrsUUJLI6" colab_type="code" colab={} outputId="8608ad01-ad19-4ca4-e05c-1e8fb12d5392"
a = 1
b = a
b = 2
print(a, b)
# + [markdown] heading_collapsed=true hidden=true id="_6l9nZ6kJLI7" colab_type="text"
# ### 10.12.1 数値や文字列はイミュータブル(変更不能)なオブジェクト
# + hidden=true id="4zXk-DshJLI7" colab_type="code" colab={} outputId="816a1d80-db3c-40aa-a5ee-f93b924eed43"
a = 1
b = a
print(id(a), id(b))
b= 2
print(id(a), id(b))
# + [markdown] heading_collapsed=true hidden=true id="H9nYuqq3JLI9" colab_type="text"
# ## 10.13 浅いコピー、深いコピー
# + hidden=true id="CyqSFHDyJLI9" colab_type="code" colab={} outputId="7ee36f19-0fbf-41b3-d10f-851129fd6131"
a = [[1, 2], [3, 4]]
b = a.copy()
b.append([5, 6])
print(a)
print(b)
b[0][0] = 0
print(a)
print(b)
print(id(a[0]), id(b[0]))
# + hidden=true id="9HQ2h2mMJLJB" colab_type="code" colab={} outputId="fa36bdbf-4253-474d-a422-ebba7044da3a"
import copy
a = [[1, 2], [3, 4]]
b = copy.deepcopy(a)
b.append([5, 6])
print(a)
print(b)
b[0][0] = 0
print(a)
print(b)
print(id(a[0]), id(b[0]))
# + [markdown] heading_collapsed=true id="X10nEDR-JLJC" colab_type="text"
# # 11. ファイル入出力
# + [markdown] heading_collapsed=true hidden=true id="C2e8SWJVJLJC" colab_type="text"
# ## 11.4 まずは動かしてみよう
# + [markdown] heading_collapsed=true hidden=true id="byj9qEwGJLJC" colab_type="text"
# ### 11.4.1 ソースコード
# + [markdown] hidden=true id="F2qzoynmJLJD" colab_type="text"
# プログラム26 ファイル入出力の例題
# + hidden=true id="ZEZ7WbsaJLJD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="a50fae3f-0d7f-461a-abea-892baf41cf34"
# 今のワーキングディレクトリ(作業中のフォルダ)
# を調べるために
# OSモジュールをimportします
import os
# 今のワーキングディレクトリを得て画面に表示します
#print(os.getcwd())
# 日本語ファイル.txtという名称のファイルを作成し、内容を書き出します
f = open('日本語ファイル.txt','w')
f.write('日本語\n日本語\n日本語\n')
f.close
# 日本語ファイル.txtを読み込み用にopenして、その内容を表示します
f = open('日本語ファイル.txt','r')
s = f.read()
f.close
print(s)
# + hidden=true id="ATKmE8UgJLJE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="afca31e0-4d80-49ae-eee0-fdefff7b699c"
# !cat ./日本語ファイル.txt
# + [markdown] hidden=true id="SDQY4NEdJLJF" colab_type="text"
# ## 11.6 例題1 波の近似
# + [markdown] hidden=true id="1LJK21AMJLJF" colab_type="text"
# ### 11.6.3 ソースコード
# + [markdown] hidden=true id="qJg3loI9JLJG" colab_type="text"
# プログラム27 のこぎり波の三角関数の和での近似
# + hidden=true id="h41_fee5JLJG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ab5bd801-2d76-4d0b-d1ec-2f330c1bc377"
# import tkinter as tk
# import tkinter.filedialog
import math
#
# tkinter の filedialog だけを利用する例
#
# root ウィンドウは withdrow() メソッドを読んで隠す
# root = tk.Tk()
# root.withdraw()
#
# 書き出し用の filedialog を読んでファイル名を得る
#
# filename = tkinter.filedialog.asksaveasfilename()
#@title String Field
filename = "sawtooth_wave.csv" #@param {type:"string"}
#
# ファイル名がもらえなければ終了
#
if filename:
pass
else:
print("No file specified")
exit()
#
# 正弦波の重ね合わせで鋸波を近似する
#
# w = sin(t) + sin(2t)/2 + sin(3t)/3 + sin(4t)/4 ...
#
# 2周期分、全体は1000ステップで、高調波は5番目まで
#
cycles = 2
steps = 1000
harmonics = 5
# ファイルが開けないときのエラー対応
try:
# ファイルを開く
with open(filename, 'w') as file:
for i in range(steps):
angle_in_degree = 360 * cycles * i / steps
angle = math.radians(angle_in_degree)
s = str(angle_in_degree)
w = 0
for i in range(1,harmonics+1):
w += math.sin(angle * (i)) / i
s = s + ", " + str(w)
# print(s)
file.write(s + "\n")
print("Writing to file " + filename + " is finished")
except IOError:
print("Unable to open file")
# + hidden=true id="7mSodPQBJLJH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="a2d70b97-54d1-4494-db41-4ab1d0631fdb"
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
#@title String Field
filename = "sawtooth_wave.csv" #@param {type:"string"}
df = pd.read_csv(filename, names=['wave1','wave2','wave3','wave4','wave5'])
plt.plot(df['wave1'])
plt.plot(df['wave2'])
plt.plot(df['wave3'])
plt.plot(df['wave4'])
plt.plot(df['wave5'])
plt.show()
# + [markdown] hidden=true id="AnSfMQnhJLJI" colab_type="text"
# #### 演習38 矩形波(方形波)の近似
# + [markdown] hidden=true id="_mo2qvGZJLJI" colab_type="text"
# 矩形波(方形波)($\pm 1$の値を交互にとる周期関数)は以下のように三角関数で近似できます。
# $f(x) = \frac{\sin(x)}{1} + \frac{\sin(3x)}{3} + \frac{\sin(5x)}{5} + \frac{\sin(7x)}{7} \cdots$
# + hidden=true id="IGx_RIveJLJI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9ea12f36-0da4-42d4-8502-c655a5acd223"
# import tkinter as tk
# import tkinter.filedialog
import math
#
# tkinter の filedialog だけを利用する例
#
# root ウィンドウは withdrow() メソッドを読んで隠す
# root = tk.Tk()
# root.withdraw()
#
# 書き出し用の filedialog を読んでファイル名を得る
#
# filename = tkinter.filedialog.asksaveasfilename()
#@title String Field
filename = "rectangle_wave.csv" #@param {type:"string"}
#
# ファイル名がもらえなければ終了
#
if filename:
pass
else:
print("No file specified")
exit()
#
# 正弦波の重ね合わせで矩形波を近似する
#
# w = sin(t) + sin(3t)/3 + sin(5t)/5 + sin(7t)/7 ...
#
# 2周期分、全体は1000ステップで、高調波は5番目まで
#
cycles = 2
steps = 1000
harmonics = 5
# ファイルが開けないときのエラー対応
try:
# ファイルを開く
with open(filename, 'w') as file:
for i in range(steps):
angle_in_degree = 360 * cycles * i / steps
angle = math.radians(angle_in_degree)
s = str(angle_in_degree)
w = 0
for i in range(1,harmonics+1):
w += math.sin(angle * (2*i-1)) / (2*i-1)
s = s + ", " + str(w)
# print(s)
file.write(s + "\n")
print("Writing to file " + filename + " is finished")
except IOError:
print("Unable to open file")
# + hidden=true id="_5KU6ndgJLJL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="613d3d62-4a88-408c-ca55-e612f465332d"
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
#@title String Field
filename = "rectangle_wave.csv" #@param {type:"string"}
df = pd.read_csv(filename, names=['wave1','wave2','wave3','wave4','wave5'])
plt.plot(df['wave1'])
plt.plot(df['wave2'])
plt.plot(df['wave3'])
plt.plot(df['wave4'])
plt.plot(df['wave5'])
plt.show()
# + [markdown] hidden=true id="EJVZP0QsJLJL" colab_type="text"
# #### 演習39 例題1のリストを使った実装
# + hidden=true id="XtoCX167JLJM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a8d57c9b-27fe-4ce1-caea-aeb6f40d755f"
# import tkinter as tk
# import tkinter.filedialog
import math
#
# tkinter の filedialog だけを利用する例
#
# root ウィンドウは withdrow() メソッドを読んで隠す
# root = tk.Tk()
# root.withdraw()
#
# 書き出し用の filedialog を読んでファイル名を得る
#
# filename = tkinter.filedialog.asksaveasfilename()
#@title String Field
filename = "sawtooth_wave.csv" #@param {type:"string"}
#
# ファイル名がもらえなければ終了
#
if filename:
pass
else:
print("No file specified")
exit()
#
# 正弦波の重ね合わせで鋸波を近似する
#
# w = sin(t) + sin(2t)/2 + sin(3t)/3 + sin(4t)/4 ...
#
# 2周期分、全体は1000ステップで、高調波は5番目まで
#
cycles = 2
steps = 1000
harmonics = 5
wave_list = [[]*(harmonics+1) for i in range(steps)]
for i in range(steps):
angle_in_degree = 360 * cycles * i / steps
angle = math.radians(angle_in_degree)
wave_list[i].append(angle_in_degree)
w = 0
for j in range(1,harmonics+1):
w += math.sin(angle * (j)) / (j)
wave_list[i].append(w)
# ファイルが開けないときのエラー対応
try:
# ファイルを開く
with open(filename, 'w') as file:
for i in range(steps):
s = str(wave_list[i][0])
w = 0
for j in range(1,harmonics+1):
s = s + ", " + str(wave_list[i][j])
# print(s)
file.write(s + "\n")
print("Writing to file " + filename + " is finished")
except IOError:
print("Unable to open file")
# + hidden=true id="etqu2jqnJLJM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="f09a9d22-8950-4457-c19c-708922bba530"
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
#@title String Field
filename = "sawtooth_wave.csv" #@param {type:"string"}
df = pd.read_csv(filename, names=['wave1','wave2','wave3','wave4','wave5'])
plt.plot(df['wave1'])
plt.plot(df['wave2'])
plt.plot(df['wave3'])
plt.plot(df['wave4'])
plt.plot(df['wave5'])
plt.show()
# + [markdown] hidden=true id="cR9trjXkJLJN" colab_type="text"
# ## 11.7 例題2
# + [markdown] hidden=true id="YKcXrKp7JLJO" colab_type="text"
# プログラム28 tkinter を用いた簡単なテキストエディタ
# + hidden=true id="0qinV6N0JLJP" colab_type="code" colab={}
import tkinter as tk
import tkinter.messagebox
import tkinter.filedialog
# messagebox, filedialog は明示的なインポートが必要
#
# tk.Frame を継承した MyFrame というクラスを作り
# その中でウィジェットやコールバック関数(メソッド)を
# 設定する。tkinter を使う定番
#
class MyFrame(tk.Frame):
# __init__はクラスオブジェクトを作る際の初期化メソッド
def __init__(self, master = None):
super().__init__(master)
self.master.title('Simple Editor')
# メニューを作る menubar -> filemenu -> Open, Save as, Exit
menubar = tk.Menu(self)
filemenu = tk.Menu(menubar, tearoff = 0)
filemenu.add_command(label = "Open", command = self.openfile)
filemenu.add_command(label = "Save as...", command = self.saveas)
filemenu.add_command(label = "Exit", command = self.master.destroy)
menubar.add_cascade(label = "File", menu = filemenu)
self.master.config(menu = menubar)
# 編集用 Text ウィジェットをクラスの変数 editbox としてつくる
self.editbox = tk.Text(self)
self.editbox.pack()
# ファイルを開くメソッド、関数とちがい self という引数が必要
def openfile(self):
# filedialog でファイル名を得る
filename = tkinter.filedialog.askopenfilename()
# filename が空でなければ処理
if filename:
tkinter.messagebox.showinfo("Filename", "Open: " + filename)
# with文で file という変数でファイルを開く
with open(filename, 'r') as file:
text = file.read()
# Text ウィジェット editbox にファイル内容を設定
self.editbox.delete('1.0', tk.END)
self.editbox.insert('1.0', text)
else:
tkinter.messagebox.showinfo("Filename","Canceled")
# ファイルに保存するメソッド
def saveas(self):
# with文で file という変数でファイルを開く
filename = tkinter.filedialog.asksaveasfilename()
if filename:
with open(filename,'w') as file:
text = file.write(self.editbox.get('1.0',tk.END))
tkinter.messagebox.showinfo("Filename", "Saved AS:" + filename)
else:
tkinter.messagebox.showinfo("Filename","Canceled")
# ここからメインプログラム
root = tk.Tk()
f = MyFrame(root)
f.pack()
f.mainloop()
# + [markdown] heading_collapsed=true id="SMA5OhxyJLJP" colab_type="text"
# # 12. 三目並べで学ぶプログラム開発
# + [markdown] hidden=true id="DOdoReeVJLJQ" colab_type="text"
# ## 12.4 三目並べを例にしたプログラムの設計
# + [markdown] hidden=true id="klq2MmDkJLJQ" colab_type="text"
# ### 12.4.1 三目並べ (tic-tac-toe)
# + [markdown] hidden=true id="8Ul-TFmiJLJQ" colab_type="text"
# 三目並べのルールとゲームの進行を言葉で表現してみてください。
# + [markdown] hidden=true id="ChKzWRs_JLJQ" colab_type="text"
# ルール
# - 3×3のマス目を用意する
# - プレイヤーは2人
# - プレイヤーはそれぞれの記号(〇・×)をマス目に入れる
# - どちらかの記号が縦・横・斜め方向に3つそろったら、その記号のプレイヤーが勝利
#
# 進行
# 1. プレイヤーの先手・後手を決める
# 1. 先手のプレイヤーは〇、後手のプレイヤーは×の記号を使う
# 1. 先手のプレイヤーから空いているマス目に記号を入れる
# 1. 後手のプレイヤーも同様に行い、これを交互に繰り返す
# 1. どちらかのプレイヤーが勝利するか、どちらもそろわずにすべてのマス目が埋まったら終了
# + [markdown] hidden=true id="So5EglbcJLJR" colab_type="text"
# ## 12.5 プログラムの実装
# + [markdown] hidden=true id="0tWEJDRTJLJR" colab_type="text"
# ### 2) 実装例 (`tic_tac_toe.py`)
# + hidden=true id="CuBJcaZ8JLJS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c4bcabe8-e8d8-48a1-ce0b-704d30a5d258"
#
# 三目並べ
#
# 特にインポートするモジュールはありません
#
# 定数の定義
#
#
# play(の中で棋譜を作成する(要完成)
'三目並べのプログラムです'
OPEN = 0
FIRST = 1
SECOND = 2
EVEN = 3
#
# 恒常的な変数
#
turn = 1
board = [[0,0,0],[0,0,0],[0,0,0]]
#
# テスト用の棋譜
#
log1 = [[0,0],[1,1],[1,0],[2,0],[0,2],[0,1],[2,1],[2,2],[1,2],[EVEN]]
log2 = [[0,0],[1,0],[1,1],[2,2],[0,2],[0,1],[2,0],[FIRST]]
log3 = [[0,1],[0,0],[2,1],[1,1],[2,2],[2,0],[1,0],[0,2],[SECOND]]
#
# 手番関連の関数
#
# 手番を文字列に
#
def show_turn():
'手番を示す文字列を返す'
if turn == FIRST:
return('先手')
elif turn == SECOND:
return('後手')
else:
return('手番の値が不適切です')
#
# 手番の初期化
#
def init_turn():
'手番を初期化する'
global turn
turn = 1
#
# 手番の交代
#
def change_turn():
'手番を交代する'
global turn
if turn == FIRST:
turn = SECOND
elif turn == SECOND:
turn = FIRST
#
# 手番関連の関数のテスト
#
def test_turn():
'手番をテストする'
init_turn()
print(show_turn(),"の番です")
change_turn()
print(show_turn(),"の番です")
change_turn()
print(show_turn(),"の番です")
#
# 盤面関連の関数
#
# 盤面を表示する文字列
#
def show_board():
'盤面を表す文字列を返す'
s = ' :0 1 2\n---------\n'
for i in range(3):
s = s + str(i) + ': '
for j in range(3):
cell = ''
if board[i][j] == OPEN:
cell = ' '
elif board[i][j] == FIRST:
cell = '0'
elif board[i][j] == SECOND:
cell = 'X'
else:
cell = '?'
s = s + cell + ' '
s = s + '\n'
return s
#
# 盤面の初期化
#
def init_board():
'盤面をすべて空(OPEN)に初期化する'
for i in range(3):
for j in range(3):
board[i][j] = OPEN
#
# 盤面のi, jの位置の値を返す
#
def examin_board(i,j):
'盤面のi行j列の値を返す'
return board[i][j]
#
# 盤面のi, jに手番tを登録、状態を文字列で返す
#
def set_board(i,j,t):
'''
盤面のi, jに手番tを登録、状態を文字列で返す
返す値は
'ok' 成功
'Not empty' 空いている場所ではない
'illegal turn' 手番が正しくない
'illegal slot' 指定された場所が正しくない
'''
if (i >= 0) and (i < 3) and (j >= 0) and (j < 3):
if (t > 0) and (t < 3):
if examin_board(i, j) == 0:
board[i][j] = t
return 'OK'
else:
return 'Not empty'
else:
return 'illegal turn'
else:
return 'illegal slot'
#
# 盤面のテスト関数
#
def test_board1():
'盤面についてのテストプログラムの1つめです'
init_board()
print(show_board())
print(set_board(0,0,1))
print(show_board())
print(set_board(1,1,2))
print(show_board())
print(set_board(1,1,1))
print(show_board())
#
# 水平方向での手番tの勝ちの判定
#
def check_board_horizontal(t):
'水平方向に手番tが勝ちであることを判定します'
for i in range(3):
if (board[i][0] == t) and (board[i][1] == t) and (board[i][2] == t):
return True
return False
#
# 垂直方向での手番tの勝ちの判定
#
def check_board_vertical(t):
'垂直方向に手番tが勝ちであることを判定します'
for j in range (3):
if (board[0][j] == t) and (board[1][j] == t) and (board[2][j] == t):
return True
return False
#
# 対角方向での手番tの勝ちの判定
#
def check_board_diagonal(t):
'対角方向に手番tが勝ちであることを判定します'
if (board[0][0] == t) and (board[1][1] == t) and (board[2][2] == t):
return True
return False
#
# 逆対角方向での手番tの勝ちの判定
#
def check_board_inverse_diagonal(t):
'逆対角方向に手番tが勝ちであることを判定します'
if (board[0][2] == t) and (board[1][1] == t) and (board[2][0] == t):
return True
return False
#
# 手番tの勝ちの単純な判定
#
def is_win_simple(t):
'手番tが勝ちであることを判定します。相手が勝っていることはチェックしません'
if check_board_horizontal(t):
return True
if check_board_vertical(t):
return True
if check_board_diagonal(t):
return True
if check_board_inverse_diagonal(t):
return True
return False
#
# 相手が勝っていないことを確認しての勝ちの判定
#
def is_win_actual(t):
'手番tが勝ちであることを判定します。相手が勝っていないことも確認します'
if not is_win_simple(t):
return False
if t == FIRST:
if is_win_simple(SECOND):
return False
else:
if is_win_simple(FIRST):
return Flase
return True
#
# 盤面に埋まっていることの判定
#
def is_full():
'盤面に空きがないことを確認します'
for i in range(3):
for j in range(3):
if board[i][j] == OPEN:
return False
return True
#
# 引き分けの判定
#
def is_even():
'盤面が引き分けであることを判定します'
if is_win_simple(FIRST):
return False
if is_win_simple(SECOND):
return False
if not is_full():
return False
return True
#
# 盤面のテスト関数2つめ、勝ち判定のテスト
#
def test_board2():
'盤面をテストする関数の2番目'
init_board()
board[0][0] = FIRST
board[1][0] = FIRST
board[2][0] = FIRST
print(show_board())
print("HORIZONTSL FIRST:", check_board_horizontal(FIRST))
print("HORIZONTSL SECOND:", check_board_horizontal(SECOND))
print("VERTICAL FIRST:", check_board_vertical(FIRST))
print("VERTICAL SECOND:", check_board_vertical(SECOND))
init_board()
board[0][0] = SECOND
board[1][0] = SECOND
board[2][0] = SECOND
print(show_board())
print("HORIZONTSL FIRST:", check_board_horizontal(FIRST))
print("HORIZONTSL SECOND:", check_board_horizontal(SECOND))
print("VERTICAL FIRST:", check_board_vertical(FIRST))
print("VERTICAL SECOND:", check_board_vertical(SECOND))
init_board()
board[0][0] = FIRST
board[0][1] = FIRST
board[0][2] = FIRST
print(show_board())
print("HORIZONTSL FIRST:", check_board_horizontal(FIRST))
print("HORIZONTSL SECOND:", check_board_horizontal(SECOND))
print("VERTICAL FIRST:", check_board_vertical(FIRST))
print("VERTICAL SECOND:", check_board_vertical(SECOND))
init_board()
board[0][0] = SECOND
board[0][1] = SECOND
board[0][2] = SECOND
print(show_board())
print("HORIZONTSL FIRST:", check_board_horizontal(FIRST))
print("HORIZONTSL SECOND:", check_board_horizontal(SECOND))
print("VERTICAL FIRST:", check_board_vertical(FIRST))
print("VERTICAL SECOND:", check_board_vertical(SECOND))
init_board()
board[0][0] = FIRST
board[1][1] = FIRST
board[2][2] = FIRST
print(show_board())
print("DIAGONAL FIRST:", check_board_diagonal(FIRST))
print("DIAGONAL SECOND:", check_board_diagonal(SECOND))
print("INV DIAGONAL FIRST:", check_board_inverse_diagonal(FIRST))
print("INV DIAGONAL SECOND:", check_board_inverse_diagonal(SECOND))
init_board()
board[0][0] = SECOND
board[1][1] = SECOND
board[2][2] = SECOND
print(show_board())
print("DIAGONAL FIRST:", check_board_diagonal(FIRST))
print("DIAGONAL SECOND:", check_board_diagonal(SECOND))
print("INV DIAGONAL FIRST:", check_board_inverse_diagonal(FIRST))
print("INV DIAGONAL SECOND:", check_board_inverse_diagonal(SECOND))
init_board()
board[0][2] = FIRST
board[1][1] = FIRST
board[2][0] = FIRST
print(show_board())
print("DIAGONAL FIRST:", check_board_diagonal(FIRST))
print("DIAGONAL SECOND:", check_board_diagonal(SECOND))
print("INV DIAGONAL FIRST:", check_board_inverse_diagonal(FIRST))
print("INV DIAGONAL SECOND:", check_board_inverse_diagonal(SECOND))
init_board()
board[0][2] = SECOND
board[1][1] = SECOND
board[2][0] = SECOND
print(show_board())
print("DIAGONAL FIRST:", check_board_diagonal(FIRST))
print("DIAGONAL SECOND:", check_board_diagonal(SECOND))
print("INV DIAGONAL FIRST:", check_board_inverse_diagonal(FIRST))
print("INV DIAGONAL SECOND:", check_board_inverse_diagonal(SECOND))
#
# 盤面のテスト関数3、勝ち、引き分けの判定
#
def test_board3():
init_board()
board[0][0] = FIRST
board[1][0] = FIRST
board[2][0] = SECOND
board[0][1] = SECOND
board[1][1] = SECOND
board[2][1] = FIRST
board[0][2] = FIRST
board[1][2] = FIRST
board[2][2] = SECOND
print(show_board())
print("HORIZONTSL FIRST:", check_board_horizontal(FIRST))
print("HORIZONTSL SECOND:", check_board_horizontal(SECOND))
print("VERTICAL FIRST:", check_board_vertical(FIRST))
print("VERTICAL SECOND:", check_board_vertical(SECOND))
print("DIAGONAL FIRST:", check_board_diagonal(FIRST))
print("DIAGONAL SECOND:", check_board_diagonal(SECOND))
print("INV DIAGONAL FIRST:", check_board_inverse_diagonal(FIRST))
print("INV DIAGONAL SECOND:", check_board_inverse_diagonal(SECOND))
print("IS WIN SIMPLE FIRST", is_win_simple(FIRST))
print("IS WIN SIMPLE SECOND", is_win_simple(SECOND))
print("IS WIN ACTUAL FIRST", is_win_actual(FIRST))
print("IS WIN ACTUAL SECOND", is_win_actual(SECOND))
print("IS FULL", is_full())
print("IS EVEN", is_even())
init_board()
board[0][0] = FIRST
board[1][0] = SECOND
board[2][0] = FIRST
board[0][1] = SECOND
board[1][1] = FIRST
board[2][1] = OPEN
board[0][2] = FIRST
board[1][2] = OPEN
board[2][2] = SECOND
print(show_board())
print("HORIZONTSL FIRST:", check_board_horizontal(FIRST))
print("HORIZONTSL SECOND:", check_board_horizontal(SECOND))
print("VERTICAL FIRST:", check_board_vertical(FIRST))
print("VERTICAL SECOND:", check_board_vertical(SECOND))
print("DIAGONAL FIRST:", check_board_diagonal(FIRST))
print("DIAGONAL SECOND:", check_board_diagonal(SECOND))
print("INV DIAGONAL FIRST:", check_board_inverse_diagonal(FIRST))
print("INV DIAGONAL SECOND:", check_board_inverse_diagonal(SECOND))
print("IS WIN SIMPLE FIRST", is_win_simple(FIRST))
print("IS WIN SIMPLE SECOND", is_win_simple(SECOND))
print("IS WIN ACTUAL FIRST", is_win_actual(FIRST))
print("IS WIN ACTUAL SECOND", is_win_actual(SECOND))
print("IS FULL", is_full())
print("IS EVEN", is_even())
init_board()
board[0][0] = SECOND
board[1][0] = FIRST
board[2][0] = SECOND
board[0][1] = FIRST
board[1][1] = SECOND
board[2][1] = FIRST
board[0][2] = SECOND
board[1][2] = OPEN
board[2][2] = FIRST
print(show_board())
print("HORIZONTSL FIRST:", check_board_horizontal(FIRST))
print("HORIZONTSL SECOND:", check_board_horizontal(SECOND))
print("VERTICAL FIRST:", check_board_vertical(FIRST))
print("VERTICAL SECOND:", check_board_vertical(SECOND))
print("DIAGONAL FIRST:", check_board_diagonal(FIRST))
print("DIAGONAL SECOND:", check_board_diagonal(SECOND))
print("INV DIAGONAL FIRST:", check_board_inverse_diagonal(FIRST))
print("INV DIAGONAL SECOND:", check_board_inverse_diagonal(SECOND))
print("IS WIN SIMPLE FIRST", is_win_simple(FIRST))
print("IS WIN SIMPLE SECOND", is_win_simple(SECOND))
print("IS WIN ACTUAL FIRST", is_win_actual(FIRST))
print("IS WIN ACTUAL SECOND", is_win_actual(SECOND))
print("IS FULL", is_full())
print("IS EVEN", is_even())
#
# ログのリプレイ
#
def replay_log(log):
'棋譜 log をたどります。print文で画面に出力します'
init_board()
init_turn()
print(show_board())
for m in log:
if len(m) == 2:
print(show_turn(), "の番です")
print(set_board(m[0], m[1], turn))
print(show_board())
print("IS WIN", turn, ":", is_win_actual(turn))
change_turn()
else:
print("RESULT IN LOG:", m[0])
print("IS WIN FIRST:", is_win_actual(FIRST))
print("IS WIN SECOND:", is_win_actual(SECOND))
print("IS EVEN:", is_even())
#
# ログのテスト
#
def test_log():
'棋譜をテストします'
print("LOG1")
replay_log(log1)
print("LOG2")
replay_log(log2)
print("LOG3")
replay_log(log3)
#
#すべてのテスト
#
def test_all():
'すべてのテストを行います'
test_turn()
test_board1()
test_board2()
test_board3()
test_log()
#
# 実際のプレイ
#
def play():
'端末への入出力を用いて実際に三目並べをプレイする関数です'
init_turn()
init_board()
print(show_board())
# 棋譜用の空リストを作る。play()の外側でアクセスするなら global宣言
# global log
log = []
while True:
print(show_turn(), "の番です")
while(True):
row = int(input("行を入力してください:"))
column = int(input("列を入力してください:"))
result = set_board(row, column, turn)
print(result)
if (result == "OK"):
break
print("不適切な入力です。再度、入力してください")
# ここ(内側の while の外)で log に手を追加
#
# 要追加
log.append([row, column])
#
print(show_board())
if (is_even()):
print("引き分けです")
# ここで棋譜に勝敗(引き分け)を追加
log.append([EVEN])
break
if (is_win_actual(turn)):
print(show_turn(), "の勝ちです")
# ここで棋譜に勝敗(turn の勝ち)を追加
log.append([turn])
break
change_turn()
# ここで棋譜のリプレイ
# 現在は log は空なので判定して処理
if len(log) > 0:
replay_log(log)
else:
print("棋譜は作成されていません")
if __name__ == '__main__':
print('三目並べ')
# + hidden=true id="w8-SXRumJLJU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2a68c5e1-6c6d-4b75-c882-d78a78cad5f7"
test_all()
# + hidden=true id="BD1pHemoJLJV" colab_type="code" colab={} outputId="2bf6eca7-eb02-4a09-e231-9341b49b14b5"
play()
# + [markdown] id="ToaMCzCyJLJW" colab_type="text"
# # 13. Pythonの学術利用
# + [markdown] id="Ec737e8HJLJX" colab_type="text"
# ## 13.3 NumPy
# + [markdown] id="7_RN9I9xJLJX" colab_type="text"
# ### 13.3.1 多次元配列の生成
# + [markdown] id="Wob_4v-UJLJX" colab_type="text"
# 1) リストから生成する
# + id="Lj1Y2odNJLJY" colab_type="code" colab={} outputId="36fd9cc6-c7fd-4966-b456-60f691104aac"
import numpy as np
data1 = [1, 2, 3]
arr1 = np.array(data1) #1次元のデータ
data2 = [[1,2,3],[4,5,6]]
arr2 = np.array(data2)
print(data1)
print(arr1)
print(data2)
print(arr2)
# + [markdown] id="MxVFTbKkJLJa" colab_type="text"
# 2) すべての要素が0の配列を作る
# + id="MMxZofE2JLJa" colab_type="code" colab={} outputId="ed11193a-290c-4922-ec31-9d58d1ccbb00"
np.zeros(5) #大きさ5の1次元配列
# + id="Dlmif-fMJLJb" colab_type="code" colab={} outputId="1d5342a5-76f2-42c8-eaa5-a1d7fd7f448d"
np.zeros((2,2)) #大きさ(2,2)の2次元配列、()が二重に注意
# + id="VzZS1pL9JLJb" colab_type="code" colab={} outputId="a9cb13d6-6450-4ec8-ad84-828799377d19"
a = np.array([[1,2,3],[4,5,6]])
np.zeros_like(a) #配列aと同じ大きさ
# + [markdown] id="TQw5f_1ZJLJc" colab_type="text"
# ### 13.3.2 `ndarray`の属性
# + id="NYHFf2JkJLJd" colab_type="code" colab={} outputId="0387b56f-489a-436d-8b0b-c5d0fd701ed1"
import numpy as np
arr2 = np.array([[1,2,3],[4,5,6]])
arr2.ndim
# + id="NsIQfUUuJLJe" colab_type="code" colab={} outputId="15ff01d2-8ce2-4c07-b603-25e3bafb517b"
arr2.shape
# + id="YYIQ3ZA0JLJf" colab_type="code" colab={} outputId="ab2f45ea-80bf-42e7-8bae-537cd478bace"
arr2.dtype
# + [markdown] id="xwZUQ_ZqJLJg" colab_type="text"
# ### 13.3.3 `ndarray`の要素へのアクセス
# + id="74ggCxJOJLJg" colab_type="code" colab={} outputId="8f7aa2e0-dcea-45e2-8c9e-ad15bd5561d4"
arr1 = np.array([1,2,3])
arr1[0]
# + id="6Ue0cpzfJLJh" colab_type="code" colab={} outputId="dba98b1b-5a2e-467b-f157-32b2f7163cca"
arr1[1] = 1
arr1
# + [markdown] id="YxAVizV0JLJi" colab_type="text"
# ### 13.3.4 スライス
# + id="7kynenyUJLJi" colab_type="code" colab={} outputId="c93aa8cc-5b5b-4749-8d09-60344077cf95"
arr1[2:]
# + id="DwPWwXkaJLJj" colab_type="code" colab={} outputId="f03d69b2-3f8f-4975-c483-dc38ab9458ea"
arr2[0:2,0:2]
# + [markdown] id="SpzyPymdJLJj" colab_type="text"
# ### 13.3.5 `ndarray`の演算
# + id="O5rmq3WLJLJj" colab_type="code" colab={} outputId="0e56c71c-d251-440f-8e21-ed6f0a028abe"
arr1 = np.array([1,2,3])
arr1*2
# + id="EsQDabCEJLJk" colab_type="code" colab={} outputId="b5cc7dde-7020-419e-f693-e87d2f6cabaa"
arr1 + 1
# + [markdown] id="zYAUtsGrJLJl" colab_type="text"
# ### 13.3.6 条件を満たす要素の抽出
# + id="V9jRwD55JLJl" colab_type="code" colab={} outputId="b6642403-1f9d-4ebc-db17-5d4c46747548"
arr1 = np.array([1,2,3,4,5])
cond = arr1 > 2 #条件を満たすかどうかの配列の生成
cond
# + id="49xrOMEeJLJm" colab_type="code" colab={} outputId="5e77be6b-386f-47cd-abf4-bed65eb1b54c"
arr1[cond] #条件を指定してスライス
# + [markdown] id="AHEHolnVJLJn" colab_type="text"
# ### 13.3.7 行列計算
# + [markdown] id="AfvfZ6pTJLJn" colab_type="text"
# - 行列の転置(行と列の入れ替え)
# + id="7VFv8sg-JLJo" colab_type="code" colab={} outputId="314257ce-e45d-4b50-8ded-a708ffe6204b"
arr2 = np.array([[1,2,3],[4,5,6]])
arr2.T
# + [markdown] id="xuPaX75oJLJp" colab_type="text"
# - 行列積
# + id="HU42brZSJLJp" colab_type="code" colab={} outputId="623a589d-09f4-47d1-d493-b9ad796beef5"
arr1 = np.array([1,2])
arr2 = np.array([[1,2,3],[4,5,6]])
arr1 @ arr2
# + [markdown] id="iUC4ntN7JLJq" colab_type="text"
# - `linalg`モジュールの利用:`numpy.linalg`(numpyを`np`でインポートしていれば`np.linalg`)には以下のような行列を扱う関数が定義されています。
# + [markdown] id="cvlfzat_JLJq" colab_type="text" endofcell="--"
# -
# - diag(対角要素),
# --
# + id="waN1DNlfJLJq" colab_type="code" colab={} outputId="3ec3cec0-1cf8-45b0-a2ff-d632fe031746"
arr2 = np.array([[1,2],[3,4]])
np.diag(arr2)
# + [markdown] id="LWHe2k8xJLJr" colab_type="text"
# - - trace(対角要素の和),
# + id="jOFJ5UROJLJr" colab_type="code" colab={} outputId="174508dc-0083-4633-ce8f-1dc64cc4ba20"
arr2 = np.array([[1,2],[3,4]])
np.trace(arr2)
# + [markdown] id="VWYETE1PJLJs" colab_type="text"
# - - det(行列式)
# + id="IRAPLqNrJLJs" colab_type="code" colab={} outputId="efbe6fc3-db76-4135-cf41-0689b1901fcc"
import numpy.linalg as LA
arr2 = np.array([[1,2],[3,4]])
print(arr2[0,0]*arr2[1,1]-arr2[0,1]*arr2[1,0])
print(LA.det(arr2))
# + [markdown] id="B-3wSAAsJLJt" colab_type="text"
# - - eig(固有値)
# + id="CDkRsggZJLJu" colab_type="code" colab={} outputId="b67b0720-f110-4032-c248-8cc0f282e26a"
arr2 = np.array([[1,2],[3,4]])
LA.eig(arr2)
# + [markdown] id="K8mExYaOJLJv" colab_type="text"
# - - inv(逆行列)
# + id="T4PMQB3AJLJw" colab_type="code" colab={} outputId="c61cabb3-64a4-463c-c9ad-e020476a3b9d"
arr2 = np.array([[1,2],[3,4]])
inv_arr2 = LA.inv(arr2)
print(inv_arr2)
print(arr2 @ inv_arr2)
# + [markdown] id="AEOSfKSPJLJw" colab_type="text"
# - - solve(一次方程式を解く)
# + id="0p7DaayxJLJy" colab_type="code" colab={} outputId="63176c96-b0b5-41e0-a817-7914057c5506"
arr1 = np.array([2,2])
arr2 = np.array([[1,2],[3,4]])
LA.solve(arr2,arr1)
# + [markdown] id="WNaphadBJLJ0" colab_type="text"
# ### 13.3.8 乱数
# + id="FeVvjmlqJLJ0" colab_type="code" colab={} outputId="aaf8d48e-4d0a-43e9-cfad-d100cb70c6df"
np.random.rand(10)
# + id="p4NB6wCdJLJ2" colab_type="code" colab={} outputId="8b7ebe71-6283-4673-9023-9f2ab678a97d"
np.random.randn(5,5)
# + id="Zeztv6kAJLJ4" colab_type="code" colab={} outputId="10021ffe-44f9-4028-ba92-5c6211802709"
np.random.permutation([1,2,3,4,5])
# + id="fvzwYh2zJLJ5" colab_type="code" colab={} outputId="b10570a3-42bb-466c-f844-098fb80b65fd"
np.random.randint(2,size=10)
# + [markdown] id="Xm3U-0s6JLJ6" colab_type="text"
# ## 13.4 Matplotlib
# + [markdown] id="guX1VplQJLJ6" colab_type="text"
# ### 13.4.2 日本語での文字出力
# + id="8yRmM8L_PxZI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="18a18029-2779-4140-dc5c-2340e79bdf2a"
# !apt -qy install fonts-noto-cjk
# !pip install -q --upgrade matplotlib
# + id="9KMki05xJLJ_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="d553646a-96a1-4562-c876-1ad154b8f102"
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
# matplotlib で日本語表示を可能にする
# matplotlib version 3.1 以降なら Yu Gothic が使用可能
from matplotlib import rcParams
rcParams['font.family'] = 'sans-serif'
rcParams['font.sans-serif'] = ['Noto Sans CJK JP']
# 以下はプロット例
data = [1,2,3]
plt.plot(data)
plt.title('タイトル')
plt.show()
# + [markdown] id="Y1rYeekcJLKA" colab_type="text"
# ### 13.4.4 利用例
# + [markdown] id="AWQavTO2JLKA" colab_type="text"
# プログラム37 `use_matplotlib.py`
# + id="9L-UPFkmJLKB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 927} outputId="477dbac6-6499-4d9a-b3db-3d24383e83b8"
#
# matplotlib 基本の使い方
#
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
# matplotlib で日本語表示を可能にする
# matplotlib version 3.1 以降なら Yu Gothic が使用可能
# matplotlib.rc('font', **{'family':'Yu Gothic'})
#
# 3本の線グラフを書く
#
plt.plot([1,2,3], 'k-', label='系列1')
plt.plot([2,3,4], 'r--', label='系列2')
plt.plot([3,4,5], 'b--o', label='系列3')
#
plt.title('タイトル')
plt.xlabel('横軸')
plt.ylabel('縦軸')
plt.legend() #凡例
plt.show()
# + [markdown] id="EKIA4N1XJLKB" colab_type="text"
# プログラム38 `use_matplotlib_scatter.py`
# + id="YHNjpwMCJLKB" colab_type="code" colab={} outputId="5b6dc704-1b5f-4c01-f27f-e12bb95baf55"
#
# matplotlib で散布図を描く
#
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
# matplotlib で日本語表示を可能にする
# matplotlib version 3.1 以降なら Yu Gothic が使用可能
matplotlib.rc('font', **{'family':'Yu Gothic'})
import numpy as np
#
# ランダムなデータの作成
#
datax = np.random.randn(100)
datay = datax + np.random.randn(100)*0.3
#
# 散布図の描画
#
plt.scatter(datax,datay,label='データ1')
#
# 別のデータの作成
#
datax = np.random.randn(100)
datay = 0.6*datax + np.random.randn(100)*0.4
#
# 色を指定して散布図を作成
#
plt.scatter(datax,datay,color='red',label='データ2')
#
# タイトル、軸ラベル、凡例の記入
#
plt.title('タイトル')
plt.xlabel('横軸')
plt.ylabel('縦軸')
plt.legend()
#
# 表示
#
plt.show()
# + [markdown] id="jHmdMb8qJLKC" colab_type="text"
# プログラム39 `use_matplotlib_hist.py`
# + id="8GiBLS7-JLKC" colab_type="code" colab={} outputId="78acaceb-84b4-48fe-db8a-40e94a8cec2c"
#
# matplotlib でヒストグラムを描く
#
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
# matplotlib で日本語表示を可能にする
# matplotlib version 3.1 以降なら Yu Gothic が使用可能
matplotlib.rc('font', **{'family':'Yu Gothic'})
import numpy as np
#
# ヒストグラムの作成
#
data = np.random.randn(1000)
plt.hist(data,bins=20)
#
# タイトル、軸ラベルを設定
#
plt.title('ヒストグラム')
plt.xlabel('データの値')
plt.ylabel('頻度')
#
# 表示
#
plt.show()
# + [markdown] id="NfChUzDuJLKD" colab_type="text"
# プログラム40 `use_matplotlib.py`
# + id="say2tAaIJLKD" colab_type="code" colab={} outputId="40d16da9-ac3a-436e-8021-127227d626f3"
#
# subplot を使う例
#
#
# matplotlib でヒストグラムを描く
#
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
# matplotlib で日本語表示を可能にする
# matplotlib version 3.1 以降なら Yu Gothic が使用可能
matplotlib.rc('font', **{'family':'Yu Gothic'})
import numpy as np
#
# 3つの subplot を作成、間隔を調整
#
fig = plt.figure()
ax1 = fig.add_subplot(2,2,1)
ax2 = fig.add_subplot(2,2,2)
ax3 = fig.add_subplot(2,2,3)
plt.subplots_adjust(hspace=0.5,wspace=0.5)
#
# 1つめに線グラフを出力
#
data = np.random.randn(100).cumsum()
ax1.set_title('線グラフ')
ax1.set_xlabel('時間')
ax1.set_ylabel('場所')
#
# 2つめに散布図を出力
#
datax = np.random.randn(100)
datay = datax + np.random.randn(100)*0.3
ax2.scatter(datax,datay,label='データ1')
datax = np.random.randn(100)
datay = 0.6*datax + np.random.randn(100)*0.4
ax2.scatter(datax,datay,color='red',label='データ2')
ax2.set_title('散布図')
ax2.set_xlabel('属性1')
ax2.set_ylabel('属性2')
ax2.legend()
#
# 3つめにヒストグラムを出力
#
data = np.random.randn(1000)
ax3.hist(data,bins=20)
ax3.set_title('ヒストグラム')
ax3.set_xlabel('データの値')
ax3.set_ylabel('頻度')
#
# グラフを表示
#
plt.show()
# + [markdown] id="o2_Cf2LKJLKE" colab_type="text"
# ## 13.5 pandas
# + [markdown] id="4faFu8jFJLKE" colab_type="text"
# ### 13.5.2 DataFrame を作る
# + [markdown] id="MN3EYLWuJLKE" colab_type="text"
# 1) numpy の array から作る
# + id="yggwNCXqJLKE" colab_type="code" colab={} outputId="57d14bb8-3aaf-458c-a1b4-77dceefb1d3f"
import numpy as np
import pandas as pd
d = np.array([[1,2,3],[4,5,6],[7,8,9]])
df = pd.DataFrame(d,columns=['a','b','c'])
df
# + id="i0DndDFHJLKF" colab_type="code" colab={} outputId="097fd4c7-0fd5-4bb0-8928-12bce75b1324"
df.columns
# + id="CVaQEYxmJLKF" colab_type="code" colab={} outputId="a3a2a17b-fa7a-4a85-d828-00e675aa3fc0"
df.index
# + [markdown] id="Ds_ucxWNJLKG" colab_type="text"
# 2) リスト型を値にもった辞書から作る
# + id="fiBgf4JjJLKG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="13d1cc3e-4400-46ed-d344-7a059e46dc69"
df = pd.DataFrame({'a':[1,4,7],'b':[2,5,8],'c':[3,6,9]})
df
# + id="KKEPGFQYJLKH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="64a14d20-5666-40d9-d50a-27ee3ae2bf21"
dic = {'a':1,'b':2,'c':3}
dic['a']
# + [markdown] id="RzQAtDuIJLKH" colab_type="text"
# ### 13.5.3 csvファイルを読み込む
# + [markdown] id="7CQ6M5ftJLKH" colab_type="text"
# プログラム41 `use_read_csv.py`
# + id="kZvZtobpJLKI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 447} outputId="5d9ba44e-ceaf-4736-85dc-f7e53c2d8fd8"
import numpy as np
import pandas as pd
import os
#
# データのあるフォルダに移動、文字列に r を付けて特殊な文字の解釈を止める
#
# pandas は日本語のファイル名をうまく処理できない
#
df = pd.read_csv("sample2.csv")
#
# 水平方向 (axis = 1) に総和をとり、Total という列を作る
df['Total'] = df.sum(axis=1)
# データフレーム df を表示
print(df)
# データフレーム df の要約統計量を表示
print(df.describe())
# + [markdown] id="5gisgKXcJLKI" colab_type="text"
# ### 13.5.5 Pandas のデータのプロット
# + [markdown] id="XFKTh542JLKI" colab_type="text"
# プログラム42 `use_DataFrame_plot.py`
# + id="H79t1a1WJLKJ" colab_type="code" colab={} outputId="8a8cf6b8-218a-4f83-f943-f2e27d758c60"
import numpy as np
import pandas as pd
import os
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
#
df = pd.read_csv("sample2.csv")
#
#
# 折れ線グラフ
#
df.plot()
print("次に進むにはグラフウィンドウを閉じてください")
plt.show()
#
# 積み上げ棒グラフ
#
df.plot.bar(stacked=True)
print("次に進むにはグラフウィンドウを閉じてください")
plt.show()
#
# 散布図
#
df.plot.scatter('Japanese','English')
print("次に進むにはグラフウィンドウを閉じてください")
plt.show()
#
# 水平方向 (axis = 1) に総和をとり、Total という列を作る
#
df['Total'] = df.sum(axis=1)
#
# ヒストグラム
#
df['Total'].plot.hist()
print("次に進むにはグラフウィンドウを閉じてください")
plt.show()
# + [markdown] id="MPteTme5JLKJ" colab_type="text"
# ## 13.6 課題
# + [markdown] id="xxbwWLtlJLKK" colab_type="text"
# #### 演習41. これを改造して鋸波のフーリエ近似を描くプログラムを作成しなさい。
# + [markdown] id="87yygZ58JLKK" colab_type="text"
# プログラム43 Numpy と Matplotlib で 1~4乗のグラフを描く
# + id="OwG8Mqq9JLKK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 587} outputId="7a281918-5f92-4f20-d81f-15e1d4430c92"
#
#
# Numpy のデータを plotする例題
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# matplotlib.rc('font', **{'family':'Yu Gothic'})
#
# x の1乗~4乗をプロットする
#
steps = 100
order = 4
maxx = 2
#
# 要素の値0で steps 行、order 列の行列を作成
#
datalist = np.zeros((steps, order))
#
# 凡例用のリスト
#
legend_label = []
#
# x の値を linspace で作成
#
x = np.linspace(0, maxx, steps)
#
# 各列について、一気に計算する
#
for i in range(1, order+1):
datalist[:, i-1] = x**i
legend_label.append(str(i)+'乗')
#
# プロット
#
plt.plot(x, datalist)
plt.title('xのべき乗')
plt.xlabel('x')
plt.ylabel('x**n')
plt.legend(legend_label)
plt.show()
# + id="xW6ZmsJrJLKK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="bccef82d-cc6d-4402-bfbb-9ec8d5bbbcc3"
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# Parameters
steps = 1000
order = 8
maxx = 4*np.pi
# Make data list
datalist = np.zeros((steps, order))
# Make legend list
legend_label = []
x = np.linspace(0, maxx, steps)
for i in range (1, order+1):
if (i > 1):
datalist[:, i-1] = datalist[:, i-2] + np.sin(i*x)/i
else:
datalist[:, i-1] = np.sin(i*x)/i
legend_label.append(str(i)+' harmonics')
plt.plot(x, datalist)
plt.title('sawtooth wave')
plt.xlabel('angle')
plt.ylabel('amplitude')
plt.legend(legend_label)
plt.xticks(np.arange(0, maxx, maxx/8))
plt.show()
# + id="RZj4lgS0JLKL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 600} outputId="82254516-0b12-40f8-f460-25d7736947d2"
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# Parameters
steps = 1024
order = 50
maxx = 4*np.pi
# Make data list
datalist = np.zeros((steps, order))
# Make legend list
legend_label = []
x = np.linspace(0, maxx, steps)
for i in range (1, order+1):
if (i > 1):
datalist[:, i-1] = datalist[:, i-2] + np.sin(i*x)/i
else:
datalist[:, i-1] = np.sin(i*x)/i
legend_label.append(str(i)+' harmonics')
# print(x)
# print(datalist[:,1])
# plt.plot(datalist[:,3])
# plt.show()
f = np.zeros((steps, order))
for i in range (1, order+1):
f[:,i-1] = np.fft.fft(datalist[:,i-1])
f_half = f[x < maxx/2]
plt.plot(x, datalist)
plt.title('sawtooth wave')
plt.xlabel('angle')
plt.ylabel('amplitude')
# plt.legend(legend_label)
plt.xticks(np.arange(0, maxx, maxx/8))
plt.show()
plt.plot(abs(f_half)**2)
plt.xscale('log')
plt.title('sawtooth wave FFT')
# plt.xlabel('angle')
# plt.ylabel('amplitude')
# plt.legend(legend_label)
plt.show()
# + id="UU4G2c_NJLKM" colab_type="code" colab={}
|
kyoto_python_2019.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Hosting Large Models</h1>
# In this notebook, we'll host and perform inferences using GPT-2 using remote grid node.
#
# **Requirements:**
# - [Install pytorch_transformers lib.](https://github.com/huggingface/pytorch-transformers#installation)
# - [Choose pre-trained model.](https://huggingface.co/pytorch-transformers/pretrained_models.html)
# - Run Grid Node app.
#
# **PS: In this example, we'll use GPT-2 Model (12-layer, 768-hidden, 12-heads, 117M parameters)**
#
# +
import syft as sy
import torch as th
import grid as gr
from pytorch_transformers import GPT2Tokenizer, GPT2LMHeadModel
# -
hook = sy.TorchHook(th)
# <h2>Set up Configs</h2>
# +
# Load pre-trained model tokenizer (vocabulary)
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
# Load pre-trained model (weights)
model = GPT2LMHeadModel.from_pretrained('gpt2',torchscript=True)
# -
# <h2>Setting Input</h2>
# +
# Encode a text inputs
text = "Who was <NAME> ? <NAME> was a"
indexed_tokens = tokenizer.encode(text)
# Convert indexed tokens in a PyTorch tensor
tokens_tensor = th.tensor([indexed_tokens])
# -
# <h2>Hosting GPT-2 Model</h2>
# +
traced_model = th.jit.trace(model, (tokens_tensor,))
# Grid Node
bob = gr.WebsocketGridClient(hook, "http://localhost:3000/", id="Bob")
bob.connect()
# -
# Host GPT-2 on Bob worker
bob.serve_model(traced_model, model_id="GPT-2", allow_download=True, allow_remote_inference=True)
# <h2>Running Inference</h2>
# +
# %%time
response = bob.run_remote_inference(model_id="GPT-2", data=tokens_tensor)
predictions = th.tensor(response)
predicted_index = th.argmax(predictions[0, -1, :]).item()
predicted_text = tokenizer.decode(indexed_tokens + [predicted_index])
print("Predicted text: ", predicted_text)
# -
# <h2>Text Generation</h2>
# +
def top_k_top_p_filtering(logits, top_k=0, top_p=0.0, filter_value=-float('Inf')):
""" Filter a distribution of logits using top-k and/or nucleus (top-p) filtering
Args:
logits: logits distribution shape (vocabulary size)
top_k > 0: keep only top k tokens with highest probability (top-k filtering).
top_p > 0.0: keep the top tokens with cumulative probability >= top_p (nucleus filtering).
Nucleus filtering is described in Holtzman et al. (http://arxiv.org/abs/1904.09751)
From: https://gist.github.com/thomwolf/1a5a29f6962089e871b94cbd09daf317
"""
assert logits.dim() == 1 # batch size 1 for now - could be updated for more but the code would be less clear
top_k = min(top_k, logits.size(-1)) # Safety check
if top_k > 0:
# Remove all tokens with a probability less than the last token of the top-k
indices_to_remove = logits < th.topk(logits, top_k)[0][..., -1, None]
logits[indices_to_remove] = filter_value
if top_p > 0.0:
sorted_logits, sorted_indices = th.sort(logits, descending=True)
cumulative_probs = th.cumsum(F.softmax(sorted_logits, dim=-1), dim=-1)
# Remove tokens with cumulative probability above the threshold
sorted_indices_to_remove = cumulative_probs > top_p
# Shift the indices to the right to keep also the first token above the threshold
sorted_indices_to_remove[..., 1:] = sorted_indices_to_remove[..., :-1].clone()
sorted_indices_to_remove[..., 0] = 0
indices_to_remove = sorted_indices[sorted_indices_to_remove]
logits[indices_to_remove] = filter_value
return logits
def sample_sequence(worker, length, context, num_samples=1, temperature=1, top_k=0, top_p=0.0, is_xlnet=False, device='cpu'):
context = th.tensor(context, dtype=th.long, device=device)
context = context.unsqueeze(0).repeat(num_samples, 1)
predicted_indexes = []
generated = context
with th.no_grad():
for _ in range(length):
inputs = {'input_ids': generated}
# Inference
outputs = th.tensor(worker.run_remote_inference(model_id="GPT-2", data=generated))
# Applying Filter
next_token_logits = outputs[0, -1, :] / temperature
filtered_logits = top_k_top_p_filtering(next_token_logits, top_k=top_k, top_p=top_p)
next_token = th.multinomial(F.softmax(filtered_logits, dim=-1), num_samples=1)
# Update context shifting tokens
generated = th.cat((th.tensor([generated[0][1:].tolist()]), next_token.unsqueeze(0)), dim=1)
# Save predicted word
predicted_indexes.append(th.argmax(outputs[0, -1, :]).item())
return predicted_indexes
# -
# %%time
out = sample_sequence(bob,20, indexed_tokens)
text = tokenizer.decode(indexed_tokens + out, clean_up_tokenization_spaces=True)
print(text)
# %%capture
model_copy = bob.download_model("GPT-2")
# +
# %%time
response = model_copy(tokens_tensor)
predictions = th.tensor(response[0])
predicted_index = th.argmax(predictions[0, -1, :]).item()
predicted_text = tokenizer.decode(indexed_tokens + [predicted_index])
print("Predicted text: ", predicted_text)
|
examples/experimental/Host GPT-2 (12-layer, 768-hidden, 12-heads, 117M parameters) .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ART for Tensorflow v2 - Keras API
# This notebook demonstrate applying ART with the new Tensorflow v2 using the Keras API. The code follows and extends the examples on www.tensorflow.org.
# +
import warnings
warnings.filterwarnings('ignore')
import tensorflow as tf
tf.compat.v1.disable_eager_execution()
import numpy as np
from matplotlib import pyplot as plt
from art.estimators.classification import KerasClassifier
from art.attacks.evasion import FastGradientMethod, CarliniLInfMethod
# -
if tf.__version__[0] != '2':
raise ImportError('This notebook requires Tensorflow v2.')
# # Load MNIST dataset
# +
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
x_test = x_test[0:100]
y_test = y_test[0:100]
# -
# # Tensorflow with Keras API
# Create a model using Keras API. Here we use the Keras Sequential model and add a sequence of layers. Afterwards the model is compiles with optimizer, loss function and metrics.
# +
model = tf.keras.models.Sequential([
tf.keras.layers.InputLayer(input_shape=(28, 28)),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy']);
# -
# Fit the model on training data.
model.fit(x_train, y_train, epochs=3);
# Evaluate model accuracy on test data.
loss_test, accuracy_test = model.evaluate(x_test, y_test)
print('Accuracy on test data: {:4.2f}%'.format(accuracy_test * 100))
# Create a ART Keras classifier for the Tensorflow Keras model.
classifier = KerasClassifier(model=model, clip_values=(0, 1))
# ## Fast Gradient Sign Method attack
# Create a ART Fast Gradient Sign Method attack.
attack_fgsm = FastGradientMethod(estimator=classifier, eps=0.3)
# Generate adversarial test data.
x_test_adv = attack_fgsm.generate(x_test)
# Evaluate accuracy on adversarial test data and calculate average perturbation.
loss_test, accuracy_test = model.evaluate(x_test_adv, y_test)
perturbation = np.mean(np.abs((x_test_adv - x_test)))
print('Accuracy on adversarial test data: {:4.2f}%'.format(accuracy_test * 100))
print('Average perturbation: {:4.2f}'.format(perturbation))
# Visualise the first adversarial test sample.
plt.matshow(x_test_adv[0])
plt.show()
# ## Carlini&Wagner Infinity-norm attack
# Create a ART Carlini&Wagner Infinity-norm attack.
attack_cw = CarliniLInfMethod(classifier=classifier, eps=0.3, max_iter=100, learning_rate=0.01)
# Generate adversarial test data.
x_test_adv = attack_cw.generate(x_test)
# Evaluate accuracy on adversarial test data and calculate average perturbation.
loss_test, accuracy_test = model.evaluate(x_test_adv, y_test)
perturbation = np.mean(np.abs((x_test_adv - x_test)))
print('Accuracy on adversarial test data: {:4.2f}%'.format(accuracy_test * 100))
print('Average perturbation: {:4.2f}'.format(perturbation))
# Visualise the first adversarial test sample.
plt.matshow(x_test_adv[0, :, :])
plt.show()
|
notebooks/art-for-tensorflow-v2-keras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import dask.distributed, os
import xarray, numpy
xarray.set_options(display_style='text');
# Explicitly open a Dask cluster,
cluster = dask.distributed.LocalCluster(
n_workers=4, processes=True,
local_directory=os.getenv('TMPDIR'),
ip='0.0.0.0',
)
client = dask.distributed.Client(cluster)
client
# Specify file paths,
source_path='/gxfs_work1/geomar/smomw355/model_data/ocean-only/VIKING20X.L46-KFS003/nemo/output'
example_file='1_VIKING20X.L46-KFS003_1m_19580101_19581231_grid_T.nc'
target_path='/gxfs_work1/geomar/smomw260/github/NEMO-to-cloud'
target_prefix='1_VIKING20X.L46-KFS003_1m_grid_T'
# Clean-up target file system location,
# !rm -r "$target_path/$target_prefix".nc "$target_path/$target_prefix".zarr
# !ls "$target_path"
# Check original netcdf format details,
# !ncdump -s -h "$source_path/$example_file"
# Open example file,
original_data = xarray.open_dataset(source_path+'/'+example_file)[['votemper', 'vosaline']]
original_data.encoding
original_data.votemper.encoding
original_data.vosaline.encoding
# %time original_data.votemper.isel(deptht=0).mean('time_counter').plot()
# %time original_data.vosaline.isel(deptht=0).mean('time_counter').plot()
# Specify a target chunk size,
# this one fits nicely to the original chunks and is very speedy...
original_data_rechunked = original_data.chunk({
'time_counter': 1,
'deptht': None, 'y': None, 'x': None,
})
# this one might already be useful...
original_data_rechunked_not_used_here = original_data.chunk({
'time_counter': 2,
'deptht': None, # full size
'y': 750, 'x': 750,
})
# Check that chunk size is compatible with Dask workers,
def print_chunk_size(da, dtype=numpy.float32):
target_chunk_size=numpy.prod(
[max(c) for c in list(da.chunks)]
)*numpy.zeros(1, dtype=dtype).nbytes/1e6 # in MB
print(f"{target_chunk_size} MB")
original_data_rechunked.vosaline.dtype
print_chunk_size(original_data_rechunked.vosaline, numpy.float32)
original_data_rechunked_not_used_here.vosaline.dtype
print_chunk_size(original_data_rechunked_not_used_here.vosaline, numpy.float32)
worker_mem = cluster.worker_spec[0]['options']['memory_limit']/1e9 # in GB
worker_cpus = cluster.worker_spec[0]['options']['nthreads']
this_is_several_times = 10
max_chunk_size = (worker_mem/worker_cpus)/this_is_several_times*1e3
print(f"{max_chunk_size} MB")
# Write to disk with an explicit variable encoding,
original_data_rechunked.encoding = {}
# %%time
original_data_rechunked.to_zarr(
store=target_path+'/'+target_prefix+'.zarr', mode='w',
encoding={
'votemper': {'dtype': 'float32'},
'vosaline': {'dtype': 'float32'}
},
)
# Open Zarr store,
zarr_store = xarray.open_zarr(
store=target_path+'/'+target_prefix+'.zarr'
)
zarr_store.encoding
zarr_store.votemper.encoding
zarr_store.vosaline.encoding
# %time zarr_store.votemper.isel(deptht=0).mean('time_counter').plot()
# %time zarr_store.vosaline.isel(deptht=0).mean('time_counter').plot()
# Save as netcdf again,
# %time zarr_store.to_netcdf(target_path+'/'+target_prefix+'.nc')
# !ncdump -s -h "$target_path/$target_prefix".nc
# From netcdf again,
netcdf_data = xarray.open_dataset(target_path+'/'+target_prefix+'.nc')
netcdf_data.encoding
netcdf_data.votemper.encoding
netcdf_data.vosaline.encoding
# %time netcdf_data.votemper.isel(deptht=0).mean('time_counter').plot()
# %time netcdf_data.vosaline.isel(deptht=0).mean('time_counter').plot()
# Python environment,
pip list
# !mamba list --explicit
|
exploratory/21-04-01_netcdf-zarr-conversion-tests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/davemlz/eemont/blob/master/docs/tutorials/022-Overloaded-Operators-List.ipynb)
# [](https://studiolab.sagemaker.aws/import/github/davemlz/eemont/blob/master/docs/tutorials/022-Overloaded-Operators-List.ipynb)
# [](https://pccompute.westeurope.cloudapp.azure.com/compute/hub/user-redirect/git-pull?repo=https://github.com/davemlz/eemont&urlpath=lab/tree/eemont/docs/tutorials/022-Overloaded-Operators-List.ipynb&branch=master)
# + [markdown] id="jZEthLln92Ep"
# # Overloaded Operators for the ee.List Object Class
# + [markdown] id="dNa470OZ8Oec"
# _Tutorial created by **<NAME>**_: [GitHub](https://github.com/davemlz) | [Twitter](https://twitter.com/dmlmont)
#
# - GitHub Repo: [https://github.com/davemlz/eemont](https://github.com/davemlz/eemont)
# - PyPI link: [https://pypi.org/project/eemont/](https://pypi.org/project/eemont/)
# - Conda-forge: [https://anaconda.org/conda-forge/eemont](https://anaconda.org/conda-forge/eemont)
# - Documentation: [https://eemont.readthedocs.io/](https://eemont.readthedocs.io/)
# - More tutorials: [https://github.com/davemlz/eemont/tree/master/docs/tutorials](https://github.com/davemlz/eemont/tree/master/docs/tutorials)
# + [markdown] id="CD7h0hbi92Er"
# ## Let's start!
# + [markdown] id="E0rc6Cya92Es"
# If required, please uncomment:
# + id="NYzyvKtk92Es"
# #!pip install eemont
# #!pip install geemap
# + [markdown] id="x3Rm3qt_92Et"
# Import the required packges.
# + id="H0C9S_Hh92Et"
import ee, eemont, geemap
# + [markdown] id="k1sdX2p592Eu"
# Authenticate and Initialize Earth Engine and geemap.
# + id="7QDXqVwy8Oef"
Map = geemap.Map()
# -
# Let's define some ee.List objects:
L1 = ee.List.sequence(0,4)
L2 = ee.List.sequence(5,9)
# ## Overloaded Operators
# `eemont` has overloaded the binary operators in the following list for the `ee.List` class:
#
# (+, \*\)
#
# Therefore, you can now use them for list operations!
# ### Concatenation
# Concatenate two or more ee.List objects using the `+` Overloaded Operator:
L3 = L1 + L2
# The result is stored as an ee.List class. Let's check it:
L3.getInfo()
# ### Repeat
# Repeat a list n times using the `*` Overloaded Operator:
L4 = ee.List([10]) * 5
# Check the result:
L4.getInfo()
# Here is another example:
L5 = L3 * 2
# Check the result:
L5.getInfo()
|
docs/tutorials/022-Overloaded-Operators-List.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Read Matrix Data
# Read in the gene expression data sets.
# +
## Download microarray matrix files from GEO
setwd("~/NLM_Reproducibility_Workshop/tb_and_arthritis/data")
# -
url <- c("ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE54nnn/GSE54992/matrix/GSE54992_series_matrix.txt.gz",
"ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE19nnn/GSE19435/matrix/GSE19435_series_matrix.txt.gz",
"ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE15nnn/GSE15573/matrix/GSE15573_series_matrix.txt.gz",
"ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE19nnn/GSE19444/matrix/GSE19444_series_matrix.txt.gz",
"ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE19nnn/GSE19435/matrix/GSE19435_series_matrix.txt.gz",
"ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE65nnn/GSE65517/matrix/GSE65517_series_matrix.txt.gz")
# +
dest.file <- c("GSE54992_series_matrix.txt.gz","GSE19435_series_matrix.txt.gz","GSE15573_series_matrix.txt.gz",
"GSE19444_series_matrix.txt.gz","GSE19435_series_matrix.txt.gz","GSE65517_series_matrix.txt.gz")
for (i in 1:length(url)){
utils::download.file(url[i], destfile=dest.file[i], mode="wb")
}
# +
## Run perl script to reformat the matrix and filter out low quality samples
## output *_series_matrix_networkanalyst.txt
## input reformated matrices into NetworkAnalyst Website
# +
## Meanwhile, read all matrices into R scripts below to analyze them by applying similar strategies.
# +
## Download and install different types of microarray annotation database files
working.dir <- "~/NLM_Reproducibility_Workshop/tb_and_arthritis/working"
setwd(working.dir)
install.packages("BiocManager")
BiocManager::install("plyr")
BiocManager::install("annotate")
BiocManager::install("illuminaHumanv4.db")
BiocManager::install("hgu133plus2.db")
BiocManager::install("illuminaHumanv2.db")
BiocManager::install("illuminaHumanv3.db")
# -
library(annotate)
library(illuminaHumanv3.db)
library(illuminaHumanv2.db)
library(hgu133plus2.db)
library(illuminaHumanv4.db)
library(plyr)
setwd("/home/ubuntu/NLM_Reproducibility_Workshop/tb_and_arthritis/working/")
getwd()
# +
## read in each type of microarray annotation database
## test one by one and make sure the annotation file match with the dataset
## could loop them into a list object
dat.v2 <- read.delim("GSE15573_series_matrix_networkanalyst.txt")
id.v2 <- select(illuminaHumanv2.db, as.character(dat.v2[2:nrow(dat.v2),1]),
c("SYMBOL","ENTREZID", "GENENAME"))
dat.v3.1 <- read.delim("GSE19435_series_matrix_networkanalyst.txt")
id.v3.1 <- select(illuminaHumanv3.db, as.character(dat.v3.1[2:nrow(dat.v3.1),1]),
c("SYMBOL","ENTREZID", "GENENAME"))
dat.v3.2 <- read.delim("GSE19444_series_matrix_networkanalyst.txt")
id.v3.2 <- select(illuminaHumanv3.db, as.character(dat.v3.2[2:nrow(dat.v3.2),1]),
c("SYMBOL","ENTREZID", "GENENAME"))
dat.v4 <- read.delim("GSE65517_series_matrix_networkanalyst.txt")
id.v4 <- select(illuminaHumanv4.db, as.character(dat.v4[2:nrow(dat.v4),1]),
c("SYMBOL","ENTREZID", "GENENAME"))
dat.plus2 <- read.delim("GSE4588_series_matrix_networkanalyst.txt")
id.plus2 <- select(hgu133plus2.db, as.character(dat.plus2[2:nrow(dat.plus2),1]),
c("SYMBOL","ENTREZID", "GENENAME"))
dat.plus2.2 <- read.delim("GSE54992_series_matrix_networkanalyst.txt")
id.plus2.2 <- select(hgu133plus2.db, as.character(dat.plus2.2[2:nrow(dat.plus2.2),1]),
c("SYMBOL","ENTREZID", "GENENAME"))
colnames(dat.v2)[1]=colnames(id.v2)[1]
dat.v2.all <- join(dat.v2,id.v2,by="PROBEID")
colnames(dat.v3.1)[1]=colnames(id.v3.1)[1]
dat.v3.1.all <- join(dat.v3.1,id.v3.1,by="PROBEID")
colnames(dat.v3.2)[1]=colnames(id.v3.2)[1]
dat.v3.2.all <- join(dat.v3.2,id.v3.2,by="PROBEID")
colnames(dat.v4)[1]=colnames(id.v4)[1]
dat.v4.all <- join(dat.v4,id.v4,by="PROBEID")
colnames(dat.plus2)[1]=colnames(id.plus2)[1]
dat.plus2.all <- join(dat.plus2,id.plus2,by="PROBEID")
colnames(dat.plus2.2)[1]=colnames(id.plus2.2)[1]
dat.plus2.2.all <- join(dat.plus2.2,id.plus2.2,by="PROBEID")
# -
# # Sample Filtering
# The paper used some inclusion criteria to select samples from each study. Samples without class labels are thus removed from further analyses.
datasets <- list(dat.v2.all,dat.v3.1.all,dat.v3.2.all,dat.v4.all,dat.plus2.all,dat.plus2.2.all)
for (i in 1:length(datasets)) {
dataset <- datasets[[i]]
dataset[1,(ncol(dataset)-2):ncol(dataset)] <- "Metadata"
cat("Removing",sum(is.na(dataset[1,])),"samples.\n")
datasets[[i]] <- dataset[,!is.na(dataset[1,])]
}
# # Convert into gene-based matrices
# Remove rows without gene mapping. Merge rows mapping to the same gene using the median values.
dataset.class <- data.frame()
for (i in 1:length(datasets)) {
dataset <- datasets[[i]]
dataset.class <- rbind(dataset.class,t(dataset[1,-c(1,(ncol(dataset)-2):ncol(dataset)),drop=F]))
dataset <- dataset[-1,]
cat("Number of rows without gene symbols:",sum(is.na(dataset$SYMBOL)),"\n")
dataset <- subset(dataset,!is.na(dataset$SYMBOL))
dataset.expr <- apply(dataset[,-c(1,(ncol(dataset)-2):ncol(dataset))],2,as.numeric)
#print(head(dataset.expr))
dataset <- aggregate(dataset.expr,
list(dataset$SYMBOL),median)
rownames(dataset) <- dataset$Group.1
cat("From",nrow(datasets[[i]]),"rows to",nrow(dataset),"rows\n")
datasets[[i]] <- dataset
}
# # Merge datasets
# Merge all studies into one, keeping only the genes that appear in all studies
common.genes <- unlist(sapply(datasets,rownames))
common.genes <- table(common.genes)
common.genes <- names(common.genes)[common.genes == length(datasets)]
cat("Number of common genes:",length(common.genes),"\n")
merged.dataset <- data.frame()
for (i in 1:length(datasets)) {
dataset <- datasets[[i]]
if (max(dataset[,-1]) > 100) {
tmp <- dataset[,-1]
tmp[tmp < 0] <- 0
dataset[,-1] <- log2(tmp+0.001)
}
cat(min(dataset[,-1]),":",max(dataset[,-1]),"\n")
merged.dataset <- rbind(merged.dataset,t(dataset[common.genes,]))
}
merged.dataset <- cbind(dataset.class,merged.dataset[rownames(dataset.class),])
colnames(merged.dataset)[1] <- "#CLASS"
#write.table(t(merged.dataset),file = "../data/merged.dataset.txt",sep="\t",quote = F,row.names = T,col.names = T)
head(datasets[[4]])
head(dataset.class)
# # Meta-Analysis
# Used the MetaIntegrator package to run meta-analysis based on random-effect model. For each dataset, the expression values are quantile normalized and log2 transformed.
# +
library("MetaIntegrator")
metaSet <- list()
# use the min expr across studies as offset for log2 transformation
min.expr <- min(unlist(lapply(datasets,function(x){min(x[common.genes,-1])})))
#print(min.expr)
for (i in 1:length(datasets)) {
dataset <- datasets[[i]][common.genes,-1]
# create data object
dataObj <- list()
dataObj$pheno <- data.frame(names=colnames(dataset),row.names = colnames(dataset))
dataObj.class <- setNames(rep(0,ncol(dataset)),colnames(dataset))
dataObj.class[as.character(dataset.class[colnames(dataset),1])!="Healthy"] <- 1
dataObj$class <- dataObj.class
dataObj$keys <- setNames(rownames(dataset),rownames(dataset))
dataObj$formattedName <- paste0("Study.",i)
expr <- as.matrix(dataset)
# quantile normalization
expr <- preprocessCore::normalize.quantiles(expr)
rownames(expr) <- rownames(dataset)
colnames(expr) <- colnames(dataset)
# log2 transformation
#print(min(expr))
dataObj$expr <- log2(expr+abs(min.expr))
#print(dim(dataObj$expr))
if (checkDataObject(dataObj,"Dataset")) {
metaSet[[i]] <- dataObj
names(metaSet)[i] <- dataObj$formattedName
str(metaSet[[i]], max.level = 1)
}
}
metaObj <- list(originalData = metaSet)
# run meta-analysis
metaObj <- runMetaAnalysis(metaObj,maxCores = 1)
# -
res <- metaObj$metaAnalysis$pooledResults
res <- res[order(abs(res$effectSizeFDR),decreasing = F),]
cat("Number of genes with effect size FDR < 1%:",sum(res$effectSizeFDR < 0.01),"\n")
head(res,10)
dim(res)
rownames(res)[1:10]
class(rownames(res))
length(rownames(res))
# +
## compare with top341 published gene list
#summary(rownames(res)[1:312]%in%as.character(sig341[,1]))
as.character(sig341[,1])[1:10]
summary(rownames(res)[1:312]%in%as.character(sig341[,1]))
# -
overlap <- intersect(rownames(res)[1:312],as.character(sig341[,1]))
#overlap.n <- vennCounts(overlap)
overlap
BiocManager::install("VennDiagram")
library(VennDiagram)
venn.plot <- draw.pairwise.venn(area1 = length(rownames(res[1:312])),#no MCP set
area2 = nrow(sig341),#no JPR set
cross.area = length(overlap),
c("MetaAnalysis SigDE", "Paper SigDE"), scaled = TRUE,
fill = c("green", "blue"),
cex = 1.5,
cat.cex = 1.5,
cat.pos = c(320, 25),
cat.dist = .05)
# # Differential Expression Genes Detection
# Used sva, limma package to do Combat normalization to remove the batch factor and detected the differential expressed genes between disease and healthy groups based on regression model.
# +
### to do combat normalization
# -
library(sva)
library(devtools)
# +
#head(t(merged.dataset) # should save merged.dataset
# +
## continue working on merged datasets and following the similiar strategy
# -
setwd("~/NLM_Reproducibility_Workshop/data")
merged.dataset <- read.delim("merged.dataset.txt")
head(merged.dataset)
# +
merged.dataset2 <- merged.dataset[-1,]
grps <- as.character(as.matrix(merged.dataset[1,]))
merged.dataset.num <- round(apply(merged.dataset[-1,],2,as.numeric),2)
dim(merged.dataset.num)
grp <- c("Healthy","TB","RA")
#grp.all <- c(which(merged.dataset[1,]==grp[1]),which(merged.dataset[1,]==grp[2]),which(merged.dataset[1,]==grp[3]))
#length(grp.all)
#merged.dataset.num.rord <- merged.dataset.num[,grp.all]
#rownames(merged.dataset.num.rord)=rownames(merged.dataset)[-1]
#colnames(merged.dataset.num.rord)
cl <- rep(c("green","red","blue","purple","orange","yellow"),c(10,30,30,30,30,11)) ## find the numbers of the samples in each study
# +
# combat normalize crossing all batches samples
#merged.dataset.norm <- normalizeVSN(merged.dataset.num.rord)
design <- model.matrix(~ grps)
batches <- rep(c(1:6),c(10,30,30,30,30,11))
# -
merged.dataset.norm <- ComBat(merged.dataset.num, batches, mod=design, par.prior=TRUE, prior.plots=FALSE)
rownames(merged.dataset.norm)=rownames(merged.dataset)[-1]
combat_fit = lm.fit(design,t(merged.dataset.norm))
par(mfrow=c(1,1))
hist(combat_fit$coefficients[2,],col=2,breaks=100)
par(mfrow=c(2,1))
boxplot(merged.dataset.num, col=cl)
boxplot(merged.dataset.norm, col=cl)
## PCA analysis
merged.dataset.num.pca <- prcomp(t(merged.dataset.num))
merged.dataset.norm.pca <- prcomp(t(merged.dataset.norm))
merged.dataset.num.pca.proportionvariances <- ((merged.dataset.num.pca$sdev^2) / (sum(merged.dataset.num.pca$sdev^2)))*100
merged.dataset.norm.pca.proportionvariances <- ((merged.dataset.norm.pca$sdev^2) / (sum(merged.dataset.norm.pca$sdev^2)))*100
# +
### Make PCA plots
plot(merged.dataset.num.pca$x)
plot(merged.dataset.num.pca$x, type="n", main="Principal components analysis bi-plot",
xlab=paste("PC1, ", round(merged.dataset.num.pca.proportionvariances[1], 1), "%"),
ylab=paste("PC2, ", round(merged.dataset.num.pca.proportionvariances[2], 1), "%"))
points(merged.dataset.num.pca$x, col=cl, pch=16, cex=1)
plot(merged.dataset.norm.pca$x)
plot(merged.dataset.norm.pca$x, type="n", main="Principal components analysis bi-plot",
xlab=paste("PC1, ", round(merged.dataset.norm.pca.proportionvariances[1], 1), "%"),
ylab=paste("PC2, ", round(merged.dataset.norm.pca.proportionvariances[2], 1), "%"))
points(merged.dataset.norm.pca$x, col=cl, pch=c(14,10,16), cex=1)
# -
head(merged.dataset.norm)
dim(merged.dataset.norm)
# +
library(limma)
design.limma <- model.matrix(~ 0+grps)
design.limma
fit <- lmFit(merged.dataset.norm, design.limma)
contrast.matrix=makeContrasts(grpsRA-grpsHealthy,grpsTB-grpsHealthy,
(grpsRA+grpsTB)-grpsHealthy,levels=design.limma)
colnames(contrast.matrix)=c("RA_ctrl","TB_ctrl","RATB_ctrl")
fit.contrast=contrasts.fit(fit,contrast.matrix)
efit.contrast=eBayes(fit.contrast)
# -
par(mfrow=c(1,3))
for (i in 1:ncol(efit.contrast$p.value)){
hist(efit.contrast$p.value[,i],main=colnames(efit.contrast$p.value)[i])
}
volcanoplot(efit.contrast, coef=1, highlight=10,names=rownames(efit.contrast),
xlab="Log Fold Change", ylab="Log Odds", pch=16,main="RA vs. Healthy volcanoPlot")
topTable(efit.contrast,coef=1,adjust.method="BH",n=50,p.value=0.01)
topTable(efit.contrast,coef=2,adjust.method="BH",n=50,p.value=0.01)
topTable(efit.contrast,coef=3,adjust.method="BH",n=50,p.value=0.01)
result <-decideTests(efit.contrast,adjust.method = "BH", p.value = 0.01)
summary(result)
vennDiagram(result,include="both",circle.col =c("red","blue","green") )
results <- classifyTestsF(efit.contrast, p.value=0.01)
summary(results)
vennDiagram(results,include="both",circle.col =c("red","blue","green") )
top.table <- topTable(efit.contrast, coef=3, sort.by = "P", n = 350)
head(top.table)
getwd()
sig341 <- read.table("/home/ubuntu/NLM_Reproducibility_Workshop/data/sig341Genes_inPaper.txt",header=F)
head(sig341)
dim(sig341)
summary(as.character(sig341[,1])%in%rownames(top.table))
|
scripts/.ipynb_checkpoints/MetaAnalysis-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import xarray as xr
import numpy as np
import pandas as pd
import fsspec
import io
filename = 'gs://solar-pv-nowcasting-data/PV/PVOutput.org/UK_PV_metadata.csv'
pv_metadata = pd.read_csv(filename)
# xr of coords = datetime, and data variabes are all the different pv systems
print(pv_metadata.columns)
# need to load system_id, longitude, latitude
# +
filename = "gs://solar-pv-nowcasting-data/PV/Passive/20211027_Passiv_PV_Data/system_metadata.csv"
passive_metadata = pd.read_csv(filename)
filename = "gs://solar-pv-nowcasting-data/PV/Passive/20211027_Passiv_PV_Data/llsoa_centroids.csv"
passive_llsoacd = pd.read_csv(filename)
# join llsoacdf data
passive_metadata = passive_metadata.merge(passive_llsoacd, on='llsoacd', how='left')
passive_metadata['system_id'] = passive_metadata['ss_id']
print(passive_metadata.columns)
print(passive_llsoacd.columns)
assert 'system_id' in passive_metadata.columns
assert 'longitude' in passive_metadata.columns
assert 'latitude' in passive_metadata.columns
print(passive_metadata[['system_id','longitude']])
# +
filename = "gs://solar-pv-nowcasting-data/PV/Passive/20211027_Passiv_PV_Data/system_metadata_OCF_ONLY.csv"
passive_metadata_ocf = pd.read_csv(filename)
passive_metadata_ocf['system_id'] = passive_metadata_ocf['ss_id']
passive_metadata_ocf['longitude'] = passive_metadata_ocf['longitude_rounded']
passive_metadata_ocf['latitude'] = passive_metadata_ocf['latitude_rounded']
print(passive_metadata_ocf.columns)
assert 'system_id' in passive_metadata_ocf.columns
assert 'longitude' in passive_metadata_ocf.columns
assert 'latitude' in passive_metadata_ocf.columns
print(passive_metadata_ocf[['system_id','longitude']])
# +
# check the difference
print((passive_metadata_ocf['longitude'] - passive_metadata['longitude']).abs().mean())
print((passive_metadata_ocf['latitude'] - passive_metadata['latitude']).abs().mean())
# -
|
notebooks/passive/format_metadata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Figure 1d-f. Quasi-vertical displacement
#
# 1. Run the [post-processing notebook](./FigS13_S15_MintPy_post_proc4TOI.ipynb) to prepare data
# 2. Run the following to plot the figure
# +
# %matplotlib inline
import os
import numpy as np
from matplotlib import pyplot as plt
import cartopy.crs as ccrs
from mintpy.defaults.plot import *
from mintpy.utils import readfile, utils as ut, plot as pp
from mintpy import view, tsview
work_dir = os.path.expanduser('~/Papers/2021_Kirishima/figs_src/obs')
os.chdir(work_dir)
print('Go to directory:', work_dir)
dem_file = os.path.expanduser('~/data/Kirishima/DEM/gsi10m.dem.wgs84')
## Points of Interest
lalo_list = [
# Shinmoe-dake
[31.9131, 130.8867], # point A for AlosDT73 - eastern rim
[31.9113, 130.8774], # point B for Alos2DT23 - western rim
# Iwo-yama
#[31.9467, 130.8524], # POI for Alos2AT131 -crater
[31.9465, 130.8531], # point C for Alos2DT23 - crater
[31.9450, 130.8528], # point D for Alos2DT23 - southern vent
[31.9467, 130.8506], # point E for Alos2DT23 - western vent
]
ref_lalo_Shinmoe = [31.916, 130.850]
ref_lalo_Iwo = [31.9315, 130.8733]
# +
# options for view.py
opt = ' --dem {} --contour-step 100 --contour-smooth 0.0 --shade-az 45 '.format(dem_file)
opt += ' -c cmy --wrap --wrap-range -5 5 -u cm '
opt += ' --notitle --fontsize 12 --ref-size 3 --nocbar --alpha 0.75 '
opt += ' --lalo-step 0.03 --lalo-loc 0 0 0 0 '
opt += ' --scalebar 0.2 0.13 0.04 --scalebar-pad 0.05 --noverbose '
opt += ' --sub-lat 31.895 31.955 --sub-lon 130.843 130.900 '
opt += ' --ref-lalo {} {} '.format(ref_lalo_Shinmoe[0], ref_lalo_Shinmoe[1])
figsize = [2.7, 2.85]
# -
# ### Fig. 1d. 2008-2010 from ALOS-1
# +
hv_file = os.path.expanduser('~/data/Kirishima/Model/data/KirishimaPost2008.h5')
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=figsize, subplot_kw=dict(projection=ccrs.PlateCarree()))
cmd = 'view.py {f} vertical {o} {o2} '.format(f=hv_file, o=opt, o2=' --noscalebar')
data, atr, inps = view.prep_slice(cmd)
ax, inps, im, cbar = view.plot_slice(ax, data, atr, inps)
# point of interest: Shinmoe-dake
ax.plot(lalo_list[0][1], lalo_list[0][0], "k.", mew=1., ms=6)
# point of interest: Iwo-yama
for lalo in lalo_list[2:]:
ax.plot(lalo[1], lalo[0], "ko", mfc='none', mew=1., ms=3)
ax.plot(ref_lalo_Iwo[1], ref_lalo_Iwo[0], "ks", mfc='none', mew=0.75, ms=3)
fig.subplots_adjust(left=0.05, right=0.95, top=0.95, bottom=0.05)
# output
out_file = os.path.abspath('dis_map_KirishimaPost2008Up.png')
plt.savefig(out_file, bbox_inches='tight', transparent=True, dpi=fig_dpi)
print('save figure to file', out_file)
plt.show()
# -
# ### Fig. 1e. 2015-2017 from ALOS-2
# +
hv_file = os.path.expanduser('~/data/Kirishima/Model/data/KirishimaPre2017.h5')
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=figsize, subplot_kw=dict(projection=ccrs.PlateCarree()))
cmd = 'view.py {f} vertical {o} {o2} '.format(f=hv_file, o=opt, o2=' ')
data, atr, inps = view.prep_slice(cmd)
ax, inps, im, cbar = view.plot_slice(ax, data, atr, inps)
# point of interest: Shinmoe-dake
ax.plot(lalo_list[1][1], lalo_list[1][0], "k.", mew=1., ms=6)
# point of interest: Iwo-yama
for lalo in lalo_list[2:]:
ax.plot(lalo[1], lalo[0], "ko", mfc='none', mew=1., ms=3)
ax.plot(ref_lalo_Iwo[1], ref_lalo_Iwo[0], "ks", mfc='none', mew=0.75, ms=3)
fig.subplots_adjust(left=0.05, right=0.95, top=0.95, bottom=0.05)
# output
out_file = os.path.abspath('dis_map_KirishimaPre2017Up.png')
plt.savefig(out_file, bbox_inches='tight', transparent=True, dpi=fig_dpi)
print('save figure to file', out_file)
plt.show()
# -
# ### Fig. 1f. 2017-2019 from ALOS-2
# +
hv_file = os.path.expanduser('~/data/Kirishima/Model/data/KirishimaPost2017.h5')
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=figsize, subplot_kw=dict(projection=ccrs.PlateCarree()))
cmd = 'view.py {f} vertical {o} {o2} '.format(f=hv_file, o=opt, o2=' --noscalebar')
data, atr, inps = view.prep_slice(cmd)
ax, inps, im, cbar = view.plot_slice(ax, data, atr, inps)
# point of interest: Shinmoe-dake
ax.plot(lalo_list[1][1], lalo_list[1][0], "k.", mew=1., ms=6)
# point of interest: Iwo-yama
for lalo in lalo_list[2:]:
ax.plot(lalo[1], lalo[0], "ko", mfc='none', mew=1., ms=3)
ax.plot(ref_lalo_Iwo[1], ref_lalo_Iwo[0], "ks", mfc='none', mew=0.75, ms=3)
fig.subplots_adjust(left=0.05, right=0.95, top=0.95, bottom=0.05)
# output
out_file = os.path.abspath('dis_map_KirishimaPost2017up.png')
plt.savefig(out_file, bbox_inches='tight', transparent=True, dpi=fig_dpi)
print('save figure to file', out_file)
plt.show()
# +
# colorbar
fig, cax = plt.subplots(nrows=1, ncols=1, figsize=[1.5, 0.15])
cbar = plt.colorbar(im, cax=cax, orientation='horizontal', ticks=[-5, 0, 5])
cbar.ax.tick_params(labelsize=font_size)
#cbar.set_label('vertical\ndisplacement [cm]', fontsize=font_size)
#cbar.ax.yaxis.set_label_position("left")
# output
out_file = os.path.abspath('cbar.png')
plt.savefig(out_file, bbox_inches='tight', transparent=True, dpi=fig_dpi)
print('save figure to file', out_file)
plt.show()
# -
# ## Back Ups: Interesting Displacement Time-series
#
# ### Uplift of Iwo-yama after the 2017 Shinmoe-dake eruption
## ALOS-2 asc T131
os.chdir(os.path.expanduser('~/data/Kirishima/KirishimaAlos2AT131/mintpyAll'))
scp_args = 'timeseries_ERA5_ramp_demErr.h5 --wrap --wrap-range -5 5 --ref-date 20161206 -n -2 --ylim -5 20 --lalo 31.9467 130.8504 '
scp_args += ' --dem inputs/gsi10m.dem.wgs84 --sub-lat 31.935 31.960 --sub-lon 130.84 130.87 --noverbose '
tsview.main(scp_args.split())
|
Fig1_obs_maps.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../scripts/')
from dp_policy_agent import *
class StateInfo: ###q4stateinfo
def __init__(self, action_num, epsilon=0.3):
self.q = np.zeros(action_num)
self.epsilon = epsilon
def greedy(self):
return np.argmax(self.q)
def epsilon_greedy(self, epsilon):
if random.random() < epsilon:
return random.choice(range(len(self.q)))
else:
return self.greedy()
def pi(self):
return self.epsilon_greedy(self.epsilon)
def max_q(self): #追加
return max(self.q)
class QAgent(DpPolicyAgent): ###q4qagent
def __init__(self, time_interval, estimator, puddle_coef=100, alpha=0.5, widths=np.array([0.2, 0.2, math.pi/18]).T, \
lowerleft=np.array([-4, -4]).T, upperright=np.array([4, 4]).T, dev_borders=[0.1,0.2,0.4,0.8]): #alpha追加
super().__init__(time_interval, estimator, None, puddle_coef, widths, lowerleft, upperright)
nx, ny, nt = self.index_nums
self.indexes = list(itertools.product(range(nx), range(ny), range(nt)))
self.actions = list(set([tuple(self.policy_data[i]) for i in self.indexes]))
self.ss = self.set_action_value_function()
###強化学習用変数### #追加
self.alpha = alpha
self.s, self.a = None, None
self.update_end = False
def set_action_value_function(self): #状態価値関数を読み込んで行動価値関数を初期化
ss = {} #state spaceという意味
for line in open("puddle_ignore_values.txt", "r"): #価値のファイルを読み込む
d = line.split()
index, value = (int(d[0]), int(d[1]), int(d[2])), float(d[3]) #インデックスをタプル、値を数字に
ss[index] = StateInfo(len(self.actions)) #StateInfoオブジェクトを割り当てて初期化
for i, a in enumerate(self.actions): #方策の行動価値を価値のファイルに書いてある値に。そうでない場合はちょっと引く
ss[index].q[i] = value if tuple(self.policy_data[index]) == a else value - 0.1
return ss
def policy(self, pose, goal=None): #q4policy
index = self.to_index(pose, self.pose_min, self.index_nums, self.widths)
s = tuple(index)
a = self.ss[s].pi()
return s, a #状態をタプルにしたものと、行動のインデックスで返すように変更
def decision(self, observation=None):###q4decision
###終了処理###
if self.update_end: return 0.0, 0.0
if self.in_goal: self.update_end = True #ゴールに入った後も一回だけ更新があるので即終了はしない
###カルマンフィルタの実行###
self.estimator.motion_update(self.prev_nu, self.prev_omega, self.time_interval)
self.estimator.observation_update(observation)
###行動決定と報酬の処理###
s_, a_ = self.policy(self.estimator.pose) #KFの結果から前回の状態遷移後の状態s'と次の行動a_(max_a'のa'ではないことに注意)を得る
r = self.time_interval*self.reward_per_sec() #状態遷移の報酬
self.total_reward += r
###Q学習と現在の状態と行動の保存###
self.q_update(self.s, self.a, r, s_) #self.s, self.aがQ値更新対象の状態行動対。報酬rと次の状態s_を使って更新。
self.s, self.a = s_, a_
###出力###
self.prev_nu, self.prev_omega = self.actions[a_]
return self.actions[a_]
def q_update(self, s, a, r, s_):###q4update
if s == None: return
q = self.ss[s].q[a]
q_ = self.final_value if self.in_goal else self.ss[s_].max_q()
self.ss[s].q[a] = (1.0 - self.alpha)*q + self.alpha*(r + q_)
###ログをとる(あとから削除)###
with open("log.txt", "a") as f:
f.write("{} {} {} prev_q:{:.2f}, next_step_max_q:{:.2f}, new_q:{:.2f}\n".format(s, r, s_, q, q_, self.ss[s].q[a]))
# +
def trial():
time_interval = 0.1
world = PuddleWorld(400000, time_interval, debug=False) #長時間アニメーション時間をとる
## 地図を生成して3つランドマークを追加 ##
m = Map()
for ln in [(-4,2), (2,-3), (4,4), (-4,-4)]: m.append_landmark(Landmark(*ln))
world.append(m)
##ゴールの追加##
goal = Goal(-3,-3)
world.append(goal)
##水たまりの追加##
world.append(Puddle((-2, 0), (0, 2), 0.1))
world.append(Puddle((-0.5, -2), (2.5, 1), 0.1))
##ロボットを1台登場させる##
init_pose = np.array([3, 3, 0]).T
kf = KalmanFilter(m, init_pose)
a = QAgent(time_interval, kf) #goalを削除
r = Robot(init_pose, sensor=Camera(m, distance_bias_rate_stddev=0, direction_bias_stddev=0),
agent=a, color="red", bias_rate_stds=(0,0))
world.append(r)
world.draw()
return a
a = trial()
# -
|
section_reinforcement_learning/q4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import requests
URL = "https://www.flipkart.com/realme-xt-pearl-blue-64-gb/p/itm731360fdbd273?pid=MOBFJYBE9FHXFEFJ&srno=s_1_1" \
"&otracker=AS_QueryStore_OrganicAutoSuggest_0_4_na_na_pr&otracker1" \
"=AS_QueryStore_OrganicAutoSuggest_0_4_na_na_pr&lid=LSTMOBFJYBE9FHXFEFJVA0XQF&fm=SEARCH&iid=a611c9af-350b-423d" \
"-87d8-df9bcc1987c7.MOBFJYBE9FHXFEFJ.SEARCH&ppt=sp&ppn=sp&ssid=n57aimhb7k0000001573581114720&qH=23f6a0071022557e "
realme-xt-pearl-blue-64-gb/product-reviews/itm731360fdbd273?pid=MOBFJYBE9FHXFEFJ
req_data = requests.get(URL)
review_soup = BeautifulSoup(req_data.content, 'html.parser')
print(dir(req_data))
review_soup
all_reviews = review_soup.find_all('a', {'href': 'col _39LH-M'})
all_reviews
# +
rating_list = []
review_header_list = []
detailed_review_list = []
user_list = []
likes_dislikes_list = []
for review in all_reviews:
#rating = review.find('div', {'class': 'hGSR34 E_uFuv'}).text
review_header = review.find_all('p', {'class': '_2xg6Ul'})
detailed_review = review.find_all('div', {'class': 'qwjRop'})
detailed_review=[e.get_text() for e in detailed_review]
user = review.find_all('p', {'class': '_3LYOAd _3sxSiS'})
user=[e.get_text() for e in user]
likes_dislikes = review.find_all('span', {'class': '_1_BQL8'})
likes_dislikes=[e.get_text() for e in likes_dislikes]
rating = review.find_all('div', {'class': 'hGSR34 E_uFuv'})
text = [e.get_text() for e in rating]
rating_list.append(text)
review_header=[e.get_text() for e in review_header]
review_header_list.append(review_header)
detailed_review_list.append(detailed_review)
user_list.append(user)
likes_dislikes_list.append(likes_dislikes)
# +
baseUrl = "https://www.flipkart.com"
all_reviews_link = review_soup.find('div', {'class': 'swINJg _3nrCtb'})
all_reviews_link
all_reviews_link.find_parent()
data = str(all_reviews_link.find_parent().get('href'))
print(baseUrl)
print(data)
url = baseUrl + data
# -
req_data = requests.get(url)
review_soup = BeautifulSoup(req_data.content, 'html.parser')
review_soup
# +
from bs4 import BeautifulSoup
import requests
URL = "https://www.flipkart.com/realme-xt-pearl-blue-64-gb/p/itm731360fdbd273?pid=MOBFJYBE9FHXFEFJ&srno=s_1_1" \
"&otracker=AS_QueryStore_OrganicAutoSuggest_0_4_na_na_pr&otracker1" \
"=AS_QueryStore_OrganicAutoSuggest_0_4_na_na_pr&lid=LSTMOBFJYBE9FHXFEFJVA0XQF&fm=SEARCH&iid=a611c9af-350b-423d" \
"-87d8-df9bcc1987c7.MOBFJYBE9FHXFEFJ.SEARCH&ppt=sp&ppn=sp&ssid=n57aimhb7k0000001573581114720&qH=23f6a0071022557e "
req_data = requests.get(URL)
review_soup = BeautifulSoup(req_data.content, 'html.parser')
all_reviews = review_soup.find_all('div', {'class': 'col _39LH-M'})
rating_list = []
review_header_list = []
detailed_review_list = []
user_list = []
likes_dislikes_list = []
for review in all_reviews:
#rating = review.find('div', {'class': 'hGSR34 E_uFuv'}).text
review_header = review.find_all('p', {'class': '_2xg6Ul'})
detailed_review = review.find_all('div', {'class': 'qwjRop'})
detailed_review=[e.get_text() for e in detailed_review]
user = review.find_all('p', {'class': '_3LYOAd _3sxSiS'})
user=[e.get_text() for e in user]
likes_dislikes = review.find_all('span', {'class': '_1_BQL8'})
likes_dislikes=[e.get_text() for e in likes_dislikes]
rating = review.find_all('div', {'class': 'hGSR34 E_uFuv'})
text = [e.get_text() for e in rating]
rating_list.append(text)
review_header=[e.get_text() for e in review_header]
review_header_list.append(review_header)
detailed_review_list.append(detailed_review)
user_list.append(user)
likes_dislikes_list.append(likes_dislikes)
# -
all_reviews
|
ReviewScrapper.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: healpyvenv
# language: python
# name: healpyvenv
# ---
# # Getting started
#
# The main interface to PySM is the `pysm3.Sky` class, the simplest way is to specify the required resolution as $N_{side}$ *HEALPix* parameter and the requested models as a list of strings, for example the simplest models for galactic dust and synchrotron `["d1", "s1"]`
import pysm3
import pysm3.units as u
import healpy as hp
import numpy as np
import warnings
warnings.filterwarnings("ignore")
sky = pysm3.Sky(nside=128, preset_strings=["d1", "s1"])
# PySM initializes the requested component objects (generally load the input templates maps with `astropy.utils.data` and cache them locally in `~/.astropy`) and stores them in the `components` attribute (a list):
sky.components
# PySM 3 uses `astropy.units`: http://docs.astropy.org/en/stable/units/ each input needs to have a unit attached to it, the unit just needs to be compatible, e.g. you can use either `u.GHz` or `u.MHz`.
map_100GHz = sky.get_emission(100 * u.GHz)
# The output of the `get_emission` method is a 2D `numpy` array in the usual `healpy` convention, `[I,Q,U]`, by default in $\mu K_{RJ}$:
map_100GHz[0, :3]
# Optionally convert to another unit using `astropy.units`
map_100GHz = map_100GHz.to(u.uK_CMB, equivalencies=u.cmb_equivalencies(100*u.GHz))
import matplotlib.pyplot as plt
# %matplotlib inline
hp.mollview(map_100GHz[0], min=0, max=1e2, title="I map", unit=map_100GHz.unit)
hp.mollview(np.sqrt(map_100GHz[1]**2 + map_100GHz[2]**2), title="P map", min=0, max=1e1, unit=map_100GHz.unit)
|
docs/basic_use.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
'''
DESCRIPTION
-----------
LocalOutlierFactor with trained model
RETURN
------
{DATASET}_lof_seen.png : png file
Similarity scores of seen label
{DATASET}_lof_seen.png : png file
Similarity score of unseen label
EXPORTED FILE(s) LOCATION
-------------------------
./reports/retrieval/{EXPERIMENT}/{DATASET}_lof_seen.png
./reports/retrieval/{EXPERIMENT}/{DATASET}_lof_unseen.png
'''
# importing default libraries
import os, argparse, sys
# sys.path.append('./')
ROOT_DIR = os.path.dirname(os.path.dirname(os.path.abspath('__file__')))
os.chdir(ROOT_DIR)
sys.path.append(ROOT_DIR)
# importing scripts in scripts folder
from scripts import config as src
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
from sklearn.manifold import TSNE
from sklearn.neighbors import LocalOutlierFactor
import warnings
warnings.filterwarnings('ignore')
import glob
TINY_SIZE = 8
SMALL_SIZE = 10
MEDIUM_SIZE = 16
BIGGER_SIZE = 20
plt.rc('font', size=MEDIUM_SIZE) # controls default text sizes
plt.rc('axes', titlesize=12) # fontsize of the axes title
plt.rc('axes', labelsize=12) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=TINY_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('legend', title_fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=MEDIUM_SIZE) # fontsize of the figure title
# -
# ## MELANOMA
loc_output = src.define_folder('./reports/retrieval/exper_melanoma')
loc_output
df_query = pd.read_pickle('./data/processed/exper_melanoma/query_log1p.pck')
df_reference = pd.read_pickle('./data/processed/exper_melanoma/reference_log1p.pck')
# +
X_query = df_query.iloc[:, :-1].values
y_ground_truth_query = df_query.iloc[:, -1:]
X_reference = df_reference.iloc[:, :-1].values
y_ground_truth_reference = df_reference.iloc[:, -1:]
# order_train = sorted(list(set(y_train.values.reshape(1,-1)[0])))
# -
order_plot = list(set(y_ground_truth_reference.values.reshape(1,-1)[0]))
order_plot.extend(['Neg.cell'])
order_plot
# ## FULL MODEL
model_ ='a1'
_, model_encoding = src.loading_model('./models/exper_melanoma/train_test_split/design_1_layer_signaling_reference_log1p_Adam.h5', -1)
model_encoding.summary()
encoding_query = model_encoding.predict(X_query)
encoding_reference = model_encoding.predict(X_reference)
print(encoding_query.shape)
print(encoding_reference.shape)
# +
clf = LocalOutlierFactor(novelty=True)
clf.fit(encoding_reference)
df_score_query = pd.concat([ y_ground_truth_query, pd.DataFrame(clf.score_samples(encoding_query), columns=['score'])], axis=1)
df_score_reference = pd.concat([ y_ground_truth_reference, pd.DataFrame(clf.score_samples(encoding_reference), columns=['score'])], axis=1)
# Calculated threshold value
threshold = np.mean(df_score_reference.groupby('cell_type').aggregate(['mean', 'std'])['score']['mean']
- df_score_reference.groupby('cell_type').aggregate(['mean', 'std'])['score']['std'])
print('Threshold value from reference dataset, ', threshold)
# + active=""
# fig, axes = plt.subplots(figsize=(6,4))#, dpi=100)
# sns.violinplot(x="cell_type", y="score", data=df_score)
#
# -
fig, axes = plt.subplots(ncols=2, sharey=True, figsize=(15,5))#, dpi=100)
sns.violinplot(x="cell_type", y="score", data=df_score_reference, ax=axes[0], order=order_plot[:-1])
sns.violinplot(x="cell_type", y="score", data=df_score_query, ax=axes[1], order=order_plot)
axes[0].axhline(threshold, color='crimson')
axes[1].axhline(threshold, color='crimson')
axes[0].set_title('for seen label')
axes[1].set_title('with unseen label')
axes[0].set(xlabel='', ylabel='similarity score')
axes[1].set(xlabel='', ylabel='')
fig.suptitle('Distribution of similarity of each cell type - '+model_+' design')
plt.tight_layout()
plt.savefig(os.path.join(loc_output, 'similarity_score_violin_'+model_+'.png'), dpi=300, bbox_inches = 'tight')
df_score_query['threshold'] = 'above'
df_score_query.loc[df_score_query['score']<=threshold, 'threshold'] = 'below'
df_score_query.shape
print(df_score_query.groupby(['threshold', 'cell_type']).size()) #/ df_score_query.groupby('cell_type').size()
(df_score_query.groupby(['threshold', 'cell_type']).size() / df_score_query.groupby('cell_type').size())*100
df_score_query.groupby('threshold').size() / len(df_score_query)
# ## LOGO
# +
LOGO_encoding_q = pd.DataFrame()
LOGO_encoding_r = pd.DataFrame()
model_ ='a1'
for i_ in range(5):
_, model_encoding = src.loading_model('./models/exper_melanoma/LeaveOneGroupOut/design_1_layer_signaling_reference_log1p_'+str(i_)+'_Adam.h5', -1)
encoding_prediction_q = model_encoding.predict(X_query)
encoding_prediction_r = model_encoding.predict(X_reference)
LOGO_encoding_q = pd.concat([LOGO_encoding_q, pd.DataFrame(encoding_prediction_q)], axis=1)
LOGO_encoding_r = pd.concat([LOGO_encoding_r, pd.DataFrame(encoding_prediction_r)], axis=1)
# +
clf = LocalOutlierFactor(novelty=True)
clf.fit(LOGO_encoding_r)
df_score_query = pd.concat([ y_ground_truth_query, pd.DataFrame(clf.score_samples(LOGO_encoding_q), columns=['score'])], axis=1)
df_score_reference = pd.concat([ y_ground_truth_reference, pd.DataFrame(clf.score_samples(LOGO_encoding_r), columns=['score'])], axis=1)
# Calculated threshold value
threshold = np.mean(df_score_reference.groupby('cell_type').aggregate(['mean', 'std'])['score']['mean']
- df_score_reference.groupby('cell_type').aggregate(['mean', 'std'])['score']['std'])
print('Threshold value from reference dataset, ', threshold)
# -
fig, axes = plt.subplots(ncols=2, sharey=True, figsize=(15,5))#, dpi=100)
sns.violinplot(x="cell_type", y="score", data=df_score_reference, ax=axes[0], order=order_plot[:-1])
sns.violinplot(x="cell_type", y="score", data=df_score_query, ax=axes[1], order=order_plot)
axes[0].axhline(threshold, color='crimson')
axes[1].axhline(threshold, color='crimson')
axes[0].set_title('for seen label')
axes[1].set_title('with unseen label')
axes[0].set(xlabel='', ylabel='similarity score')
axes[1].set(xlabel='', ylabel='')
fig.suptitle('Distribution of similarity of LOGO models - '+model_+' design')
plt.tight_layout()
plt.savefig(os.path.join(loc_output, 'similarity_score_LOGO_violin_'+model_+'.png'), dpi=300, bbox_inches = 'tight')
df_score_query['threshold'] = 'above'
df_score_query.loc[df_score_query['score']<=threshold, 'threshold'] = 'below'
df_score_query.shape
#A2
print(df_score_query.groupby(['threshold', 'cell_type']).size()) #/ df_score_query.groupby('cell_type').size()
(df_score_query.groupby(['threshold', 'cell_type']).size() / df_score_query.groupby('cell_type').size())*100
|
notebooks/7.1-pg-lof.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
path = r'C:\Users\24729\Desktop\nj_sports\Suning.xlsx'
Suning = pd.read_excel(path, usecols = 'A:K', index_col = '赛季')
del Suning['联赛级别']
Suning
Suning = Suning.rename(columns = {'场次':'game_round','胜':'win','平':'drawn','负':'lose','进球':'goals','失球':'goals_against','净胜球':'goal_difference','积分':'score','排名':'rank'})
del Suning['goal_difference']
Suning = Suning.loc[2009:]
path = r'C:\Users\24729\Desktop\nj_sports\Guoan.xlsx'
Guoan = pd.read_excel(path, skiprows = 1, usecols = 'A:J').rename(columns = {'Unnamed: 0':'赛季'})
Guoan = Guoan.replace('第8名','8').replace('亚军','2').replace('殿军','4').replace('季军','3').replace('第6名','6').replace('第9名','9').replace('第7名','7').replace('冠军','1').replace('第5名','5').replace('第9名','9')
Guoan = Guoan.drop([13,27])
Guoan['赛季'] = Guoan['赛季'].astype('int64')
Guoan = Guoan.set_index('赛季')
del Guoan['联赛组别']
Guoan = Guoan.rename(columns = {'赛':'game_round','胜':'win','平':'drawn','负':'lose','进球':'goals','失球':'goals_against','得分':'score','名次':'rank'})
Guoan = Guoan.loc[2009:2017].astype('int64')
Guoan
path = r'C:\Users\24729\Desktop\nj_sports\Luneng.xlsx'
Luneng = pd.read_excel(path, usecols = 'A:I')
Luneng['顶级联赛'] = Luneng['顶级联赛'].str.replace('第','').replace('冠军','1').replace('亚军','2').replace('季军','3')
Luneng = Luneng.set_index('赛季')
Luneng = Luneng.rename(columns = {'总场次':'game_round','胜':'win','平':'drawn','负':'lose','进球':'goals','失球':'goals_against','折算积分':'score','顶级联赛':'rank'})
Luneng = Luneng.loc[2009:2017]
Luneng
path = r'C:\Users\24729\Desktop\nj_sports\Shenhua.xlsx'
Shenhua = pd.read_excel(path, usecols = 'A:K')
Shenhua = Shenhua.replace('1→无1','1')
Shenhua['赛季'] = Shenhua['赛季'].str.replace('赛季','')
Shenhua = Shenhua.set_index('赛季')
del Shenhua['参赛球队数'], Shenhua['净胜球']
Shenhua = Shenhua.rename(columns = {'名次':'rank','参赛轮次':'game_round','胜':'win','平':'drawn','负':'lose','进球':'goals','失球':'goals_against','积分':'score'})
Shenhua = Shenhua.sort_index().iloc[15:-2]
Shenhua.index = Shenhua.index.astype('int64')
Shenhua
path = r'C:\Users\24729\Desktop\nj_sports\Shanggang.xlsx'
Shanggang = pd.read_excel(path, usecols = 'A:J')
Shanggang = Shanggang.iloc[3:12]
Shanggang['年份'] = Shanggang['年份'].str.replace('年','')
Shanggang = Shanggang.replace('第4名','4').replace('第9名','9').replace('第5名','5').replace('冠军','1').replace('亚军','2').replace('季军','3')
Shanggang.iloc[0] = [2009,'',0,0,0,0,0,0,0,0]
Shanggang.iloc[1] = [2010,'',0,0,0,0,0,0,0,0]
Shanggang.iloc[2] = [2011,'',0,0,0,0,0,0,0,0]
Shanggang.iloc[3] = [2012,'',0,0,0,0,0,0,0,0]
del Shanggang['赛事']
Shanggang = Shanggang.rename(columns = {'年份':'赛季','场次':'game_round','胜':'win','平':'drawn','负':'lose','得球':'goals','失球':'goals_against','积分':'score','名次':'rank'}).set_index('赛季')
path = r'C:\Users\24729\Desktop\nj_sports\Evergrand.xlsx'
Evergrand = pd.read_excel(path, usecols = 'A:L')
del Evergrand['队名'], Evergrand['胜率%']
Evergrand = Evergrand.loc[19:27]
Evergrand.iloc[1] = [2010,'中甲',0,0,0,0,0,0,0,0]
del Evergrand['联赛'], Evergrand['净胜球']
Evergrand = Evergrand.rename(columns = {'场次':'game_round','胜':'win','平':'drawn','负':'lose','进球':'goals','失球':'goals_against','积分':'score'}).set_index('赛季')
Evergrand
path = r'C:\Users\24729\Desktop\nj_sports\Taida.xlsx'
Taida = pd.read_excel(path, usecols = 'A:K')
Taida = Taida.loc[15:23].replace('第6名','6').replace('亚军','2').replace('第10名','10').replace('第8名','8').replace('第11名','11').replace('第7名','7').replace('第13名','13')
Taida['年份'] = Taida['年份'].str.replace('年','')
del Taida['联赛'], Taida['名称']
Taida = Taida.rename(columns = {'年份':'赛季','名次':'rank','场次':'game_round','胜':'win','平':'drawn','负':'lose','得球':'goals','失球':'goals_against','积分':'score'}).set_index('赛季')
Taida
path = r'C:\Users\24729\Desktop\nj_sports\Jianye.xlsx'
Jianye = pd.read_excel(path).iloc[15:24]
del Jianye['备注'], Jianye['联赛等级']
Jianye.iloc[4] = [2013,0,0,0,0,0,0]
Jianye = Jianye.rename(columns = {'年份':'赛季','赛':'game_round','胜':'win','平':'drawn','负':'lose','得分':'score','排名':'rank'}).set_index('赛季').astype('int64')
Jianye
year = np.arange(2009,2018)
plt.figure()
plt.plot(year, Suning['score'],'-', label = 'Suning')
plt.plot(year, Evergrand['score'],'-', label = 'Evergrand')
plt.plot(year, Luneng['score'],'-', label = 'Luneng')
plt.plot(year, Shanggang['score'],'-', label = 'Shanggang')
plt.plot(year, Guoan['score'],'-', label = 'Guoan')
plt.plot(year, Shenhua['score'],'-', label = 'Shenhua')
plt.plot(year, Jianye['score'],'-', label = 'Jianye')
plt.plot(year, Taida['score'],'-', label = 'Taida')
plt.legend()
plt.show()
# +
fig = plt.figure(dpi = 200, figsize = [8,6])
plt.subplots_adjust(hspace = .5,right = 2, wspace = .2)
plt.subplot(2,2,1)
plt.plot(year, Evergrand['score'],'-',c = 'purple', label = 'Evergrand')
plt.plot(year, Shanggang['score'],'-',c = 'navy', label = 'Shanggang')
plt.plot(year, Suning['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Luneng['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Guoan['score'],'-', c = 'gray', alpha = 0.5)
plt.plot(year, Shenhua['score'],'-', c = 'gray',alpha = .5)
plt.plot(year, Jianye['score'],'-', c = 'gray', alpha = .5)
plt.plot(year, Taida['score'],'-', c = 'gray',alpha = .5)
plt.ylabel('score')
plt.legend(loc = 0)
plt.title('Up-Rising Stars', fontsize=16)
plt.subplot(2,2,2)
plt.plot(year, Luneng['score'],'-',c = 'purple', label = 'Luneng')
plt.plot(year, Guoan['score'],'-', c = 'navy', label = 'Guoan')
plt.plot(year, Evergrand['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Shanggang['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Suning['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Shenhua['score'],'-', c = 'gray',alpha = .5)
plt.plot(year, Jianye['score'],'-', c = 'gray', alpha = .5)
plt.plot(year, Taida['score'],'-', c = 'gray',alpha = .5)
plt.ylabel('score')
plt.legend(loc = 0)
plt.title('Evergreen', fontsize=16)
plt.subplot(2,2,3)
plt.plot(year, Suning['score'],'-',c = 'red', label = 'Suning')
plt.plot(year, Shenhua['score'],'-', c = 'purple',label = 'Shenhua')
plt.plot(year, Taida['score'],'-', c = 'navy',label = 'Taida')
plt.plot(year, Luneng['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Guoan['score'],'-', c = 'gray', alpha = 0.5)
plt.plot(year, Evergrand['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Shanggang['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Jianye['score'],'-', c = 'gray', alpha = .5)
plt.ylabel('score')
plt.legend(loc = 0, ncol = 3)
plt.title('Mr. Average', fontsize=16)
plt.subplot(2,2,4)
plt.plot(year, Jianye['score'],'-', c = 'purple', label = 'Jianye')
plt.plot(year, Suning['score'],'-',c = 'gray', alpha = .5)
plt.plot(year, Shenhua['score'],'-', c = 'gray',alpha = .5)
plt.plot(year, Taida['score'],'-', c = 'gray',alpha = .5)
plt.plot(year, Luneng['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Guoan['score'],'-', c = 'gray', alpha = 0.5)
plt.plot(year, Evergrand['score'],'-',c = 'gray', alpha = 0.5)
plt.plot(year, Shanggang['score'],'-',c = 'gray', alpha = 0.5)
plt.ylabel('score')
plt.legend(loc = 4, ncol = 3)
plt.title('Poor Struggler', fontsize=16)
fig.suptitle('Four Types of Team in Chinese Football Association Super League',x = 1.05,y = 1.02, fontsize = 22)
plt.show()
plt.savefig(r'C:\Users\24729\Desktop\assignment4.jpg')
# -
|
Chinese Football Association Super League.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="qvW8-J6we6By"
# # Обучение нейросетей — оптимизация и регуляризация
#
# **Разработчик: <NAME>**
# + [markdown] id="1ecMva_Ge6B0"
# На это семинаре будет необходимо (1) реализовать Dropout-слой и проследить его влияние на обобщающую способность сети (2) реализовать BatchNormalization-слой и пронаблюдать его влияние на скорость сходимости обучения.
# + [markdown] id="wQZ-_wUwe6B0"
# ## Dropout
#
# Как всегда будем экспериментировать на датасете MNIST. MNIST является стандартным бенчмарк-датасетом, и его можно подгрузить средствами pytorch.
# + id="v4S5PFg5e6B1"
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import clear_output
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision.datasets as dsets
import torchvision.transforms as transforms
import torch.optim as optim
from torch.utils.data.sampler import SubsetRandomSampler
# + id="5EuePwt3e6B5"
input_size = 784
num_classes = 10
batch_size = 128
train_dataset = dsets.MNIST(root='./MNIST/',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = dsets.MNIST(root='./MNIST/',
train=False,
transform=transforms.ToTensor())
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=False)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# + [markdown] id="nUFeuDtfe6B9"
# Определим ряд стандартных функций с прошлых семинаров
# + id="a_rmaGJQe6B9"
def train_epoch(model, optimizer, batchsize=32):
loss_log, acc_log = [], []
model.train()
for batch_num, (x_batch, y_batch) in enumerate(train_loader):
data = x_batch
target = y_batch
optimizer.zero_grad()
output = model(data)
pred = torch.max(output, 1)[1]
acc = torch.eq(pred, y_batch).float().mean()
acc_log.append(acc)
loss = F.nll_loss(output, target).cpu()
loss.backward()
optimizer.step()
loss = loss.item()
loss_log.append(loss)
return loss_log, acc_log
def test(model):
loss_log, acc_log = [], []
model.eval()
for batch_num, (x_batch, y_batch) in enumerate(test_loader):
data = x_batch
target = y_batch
output = model(data)
loss = F.nll_loss(output, target).cpu()
pred = torch.max(output, 1)[1]
acc = torch.eq(pred, y_batch).float().mean()
acc_log.append(acc)
loss = loss.item()
loss_log.append(loss)
return loss_log, acc_log
def plot_history(train_history, val_history, title='loss'):
plt.figure()
plt.title('{}'.format(title))
plt.plot(train_history, label='train', zorder=1)
points = np.array(val_history)
plt.scatter(points[:, 0], points[:, 1], marker='+', s=180, c='orange', label='val', zorder=2)
plt.xlabel('train steps')
plt.legend(loc='best')
plt.grid()
plt.show()
def train(model, opt, n_epochs):
train_log, train_acc_log = [], []
val_log, val_acc_log = [], []
for epoch in range(n_epochs):
print("Epoch {0} of {1}".format(epoch, n_epochs))
train_loss, train_acc = train_epoch(model, opt, batchsize=batch_size)
val_loss, val_acc = test(model)
train_log.extend(train_loss)
train_acc_log.extend(train_acc)
steps = train_dataset.train_labels.shape[0] / batch_size
val_log.append((steps * (epoch + 1), np.mean(val_loss)))
val_acc_log.append((steps * (epoch + 1), np.mean(val_acc)))
clear_output()
plot_history(train_log, val_log)
plot_history(train_acc_log, val_acc_log, title='accuracy')
print("Epoch: {2}, val loss: {0}, val accuracy: {1}".format(np.mean(val_loss), np.mean(val_acc), epoch))
# + [markdown] id="MjhiP5h-e6CA"
# Создайте простейшую однослойную модель - однослойную полносвязную сеть и обучите ее с параметрами оптимизации, заданными ниже.
# + id="TLB2iHNke6CB"
class Flatten(nn.Module):
def forward(self, x):
return x.view(x.size()[0], -1)
model = nn.Sequential(
#<your code>
)
# + id="z8ppSMDae6CE"
opt = torch.optim.Adam(model.parameters(), lr=0.0005)
train(model, opt, 10)
# + [markdown] id="7smJP34Pe6CH"
# Параметром обученной нейросети является матрица весов, в которой каждому классу соответствует один из 784-мерных столбцов. Визуализируйте обученные векторы для каждого из классов, сделав их двумерными изображениями 28-28. Для визуализации можно воспользоваться кодом для визуализации MNIST-картинок с предыдущих семинаров.
# + id="WIwBt7cJe6CH"
weights = #<your code>
plt.figure(figsize=[10, 10])
for i in range(10):
plt.subplot(5, 5, i + 1)
plt.title("Label: %i" % i)
plt.imshow(weights[i].reshape([28, 28]), cmap='gray');
# + [markdown] id="TIzsqWU_e6CL"
# Реализуйте Dropout-слой для полносвязной сети. Помните, что этот слой ведет себя по-разному во время обучения и во время применения.
# + id="HaRFi9jqe6CL"
class DropoutLayer(nn.Module):
def __init__(self, p):
super().__init__()
#<your code>
def forward(self, input):
if self.training:
#<your code>
else:
#<your code>
# + [markdown] id="gvq4PLN_e6CO"
# Добавьте Dropout-слой в архитектуру сети, проведите оптимизацию с параметрами, заданными ранее, визуализируйте обученные веса. Есть ли разница между весами обученными с Dropout и без него? Параметр Dropout возьмите равным 0.7
# + id="YsfjKbTye6CO"
modelDp = nn.Sequential(
#<your code>
)
# + id="KMlZHpXae6CR"
opt = torch.optim.Adam(modelDp.parameters(), lr=0.0005)
train(modelDp, opt, 10)
# + id="Xzzz9Tkje6CT"
weights = #<your code>
plt.figure(figsize=[10, 10])
for i in range(10):
plt.subplot(5, 5, i + 1)
plt.title("Label: %i" % i)
plt.imshow(weights[i].reshape([28, 28]), cmap='gray');
# + [markdown] id="C5_G8wzQe6CW"
# Обучите еще одну модель, в которой вместо Dropout-регуляризации используется L2-регуляризация с коэффициентом 0.05. (Параметр weight_decay в оптимизаторе). Визуализируйте веса и сравните с двумя предыдущими подходами.
# + id="ayzHCMx8e6CX"
model = nn.Sequential(
Flatten(),
nn.Linear(input_size,num_classes),
nn.LogSoftmax(dim=-1)
)
# + id="pWYcCBZ7e6CZ"
opt = torch.optim.Adam(model.parameters(), lr=0.0005, weight_decay=0.05)
train(model, opt, 10)
# + id="BJbA2mA3e6Cd"
weights = #<your code>
plt.figure(figsize=[10, 10])
for i in range(10):
plt.subplot(5, 5, i + 1)
plt.title("Label: %i" % i)
plt.imshow(weights[i].reshape([28, 28]), cmap='gray');
# + [markdown] id="nKtMonw3e6Cf"
# ## Batch normalization
#
# Реализуйте BatchNormalization слой для полносвязной сети. В реализации достаточно только центрировать и разделить на корень из дисперсии, аффинную поправку (гамма и бета) в этом задании можно не реализовывать.
# + id="LWvaRXmOe6Cg"
class BnLayer(nn.Module):
def __init__(self, num_features):
super().__init__()
#<your code>
def forward(self, input):
if self.training:
#<your code>
else:
#<your code>
return #<your code>
# + [markdown] id="_tpNTUWVe6Ci"
# Обучите трехслойную полносвязную сеть (размер скрытого слоя возьмите 100) с сигмоидами в качестве функций активации.
# + id="xGNAd9Uae6Ci"
model = nn.Sequential(
#<your code>
)
# + id="7qD2KJq6e6Ck"
opt = torch.optim.RMSprop(model.parameters(), lr=0.01)
train(model, opt, 3)
# + [markdown] id="M2IHZt_Se6Co"
# Повторите обучение с теми же параметрами для сети с той же архитектурой, но с добавлением BatchNorm слоя (для всех трех скрытых слоев).
# + id="406kS0wFe6Co"
modelBN = nn.Sequential(
#<your code>
)
# + id="Uqt2bVKWe6Cq"
opt = torch.optim.RMSprop(modelBN.parameters(), lr=0.01)
train(modelBN, opt, 3)
# + [markdown] id="APw2lE1qe6Cs"
# Сравните кривые обучения и сделайте вывод о влиянии BatchNorm на ход обучения.
|
2020-fall/seminars/seminar4/DL20_fall_seminar4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SVD/PCA Algorithm
# ## Theory behind SVD and PCA
# The Spectral Theorem assures us that for any $(n,n)$ shaped, symmetric real matrix say $\mathrm{B}$, we can decompose it as $\mathrm{B}=\mathrm{Q}\mathrm{\Lambda}\mathrm{Q^{T}}$, where $\mathrm{Q}$ is an orthonormal matrix having as columns the eigenvectors of $\mathrm{B}$ and $\mathrm{\Lambda}$ is a diagonal matrix containing the eigenvalues of $\mathrm{B}$.\
# \
# This theorem is powerful and a crucial pillar of linear algebra, yet restrictive when it comes to data science matrices, hardly ever displaying those features required in its assumptions. What tool can we then use to do some similar transformation not limited to square matrices?
# \
# \
# What if we're dealing say with a matrix of data $\mathrm{A}=\begin{bmatrix}a_{1,1}&a_{1,2}\\a_{2,1}&a_{2,2}\\a_{3,1}&a_{3,2}\end{bmatrix}$ ? It is a rectangular matrix with having more observations that parameters, and we cannot apply the theorem above. This is a problem.
# \
# \
# The solution comes up to be the **SVD**, or short for **Singular Values Decomposition**. In fact, in this case the so called Singular Values will be the closest possible substitutes to our old eigenvalues. In short matrix notation, the SVD is used to decompose our good old data matrix $\mathrm{A}$ into the following expression:
# \
# \
# $$\mathrm{A} = \mathrm{U}\ \mathrm{\Sigma}\ \mathrm{V^{T}}$$
# \
# Where $\mathrm{A}$ is of shape $(n,m)$, $\mathrm{U}$ is of shape $(n,r)$, $\mathrm{\Sigma}$ is of shape $(r,r)$ and $\mathrm{V}^T$ is of shape $(r,m).$
# \
# \
# What will these mysterious new animals $\mathrm{U},\ \mathrm{\Sigma},\ \mathrm{V^{T}}$ be? Admittedly we cannot do much, we've lost our favorite tool, the Spectral Theorem, so what shall we do? **Simple**, we tweak around stuff in such a way that we are still able to use it! I cannot use this trick directly on $\mathrm{A}$, but what if I consider $\mathrm{A}^{T}\mathrm{A}$ or $\mathrm{A}\mathrm{A}^{T}$ ?\
# Turns out both of them are symmetric positive (at least) semi-definite matrices! And what happens when I do that in the equation above? Let's see.\
# \
# $$\mathrm{A}\mathrm{A}^{T} = (\mathrm{U}\ \mathrm{\Sigma}\ \mathrm{V^{T}})(\mathrm{U}\ \mathrm{\Sigma} \ \mathrm{V}^{T})^{T} = \mathrm{U}\ ( \mathrm{\Sigma}\ \mathrm{\Sigma}^T ) \mathrm{U}^{T} \ \ \ \ \ (1)$$
# \
# If we now allow $\mathrm{V}$ and $\mathrm{U}$ to be orthonormal, we have that by orthonormal properties, if a matrix $\mathrm{O}$ is orthonormal, then $\mathrm{O}^T = \mathrm{O}^{-1}$, which reduces the equation above to be
# \
# $$\mathrm{A}\mathrm{A}^{T} = \mathrm{U}( \mathrm{\Sigma} \mathrm{\Sigma}^T ) \mathrm{U}^{T} $$
# \
# But we've seen that form already, it's the Spectral Decomposition of the matrix $\mathrm{A}\mathrm{A}^{T}$! Thus $\mathrm{U}$ **must be the matrix containing as columns the eigenvectors of $\mathrm{A}\mathrm{A}^{T}$, and $( \mathrm{\Sigma} \mathrm{\Sigma}^T )$ must be our diagonal matrix of eigenvalues**.
# \
# \
# Same goes the other way now,
# $$\mathrm{A}^{T}\mathrm{A} = (\mathrm{V}\ \mathrm{\Sigma}^T \ \mathrm{U^{T}})(\mathrm{U}\ \mathrm{\Sigma} \ \mathrm{V}^{T}) = \mathrm{V}\ (\mathrm{\Sigma}\ \mathrm{\Sigma}^T) \mathrm{V}^{T}\ \ \ \ \ (2)$$
# \
# We have then that $\mathrm{V}$ **must be the matrix containing as columns the eigenvectors of $\mathrm{A}^T\mathrm{A}$, and $( \mathrm{\Sigma}^T \mathrm{\Sigma} )$ must be our diagonal matrix of eigenvalues**.
# \
# \
# Since the data matrix is always the same one, the eigenvalues of $\mathrm{A}^{T}\mathrm{A}$ are the same ones as $\mathrm{A}\mathrm{A}^T$, making $\mathrm{\Sigma}\ \mathrm{\Sigma}^T = \mathrm{\Sigma}^T\ \mathrm{\Sigma}$
# \
# \
# At this point we can use (1) and (2) to compute $\mathrm{U}, \mathrm{T}$ and $\mathrm{\Sigma}$.
# \
# This is all the SVD does for us. What is then the acronym **PCA**, short for **Principal Component Analysis**? It is in its simplest form a slight modification to the output of the SVD. Specifically it consists in arranging the components of $\mathrm{U}$, $\mathrm{\Sigma}$ and $\mathrm{V}^T$ in such away that the components of $\mathrm{\Sigma}$ are decreasing sliding down the diagonal. Mathematically $\mathrm{\Sigma}=\begin{bmatrix}\sigma_{1,1}&0&0\\0&\sigma_{2,2}&0\\0&0&\sigma_{3,3}\end{bmatrix}$, if we suppose it is $(3,3)$, where $\sigma_{1,1}\geq\sigma_{2,2}\geq\sigma_{2,2}$. This trick allows us to position the "most important" elements on top. An interpretation of the magnitude of $\sigma_{i,i}$ is "how much information do the column $i$ of $\mathrm{U}$ and row $i$ of $\mathrm{V}^T$ contain about the information in my matrix $\mathrm{A}$? Intuitively, the greater the magnitude of the $\sigma$ and the more info they contain.
#
# I will bring you a short example to fully interpret what's going on begind the svd. Suppose our matrices are composed by vectors $u$'s and $v$'s
#
# $$\mathrm{U} = \begin{bmatrix}|&|\\u_1&u_2\\|&|\end{bmatrix}_{(3x2)}, \mathrm{V} = \begin{bmatrix}-&v^{T}_1&-\\-&v^{T}_2&-\end{bmatrix}_{(2x3)}, \mathrm{\Sigma} = \begin{bmatrix}\sigma_1&0\\0&\sigma_2\end{bmatrix}_{(2x2)}$$
#
# The matrix $\mathrm{A}$ deriving from their product can be written as a sum of rank 1 matrices (there's a theorem proving this, we will see here practically). Since our shapes are $(3x2) (2x2) (2x3)$, the resulting matrix will be A, a $(3,3)$ matrix. And we can obtain it as follows:
#
# $$\mathrm{A} = \sigma_1 \begin{bmatrix}|\\u_1\\|\end{bmatrix}_{(3x1)}\begin{bmatrix}-&v^{T}_1&-\end{bmatrix}_{(1x3)} \ + \ \sigma_2 \begin{bmatrix}|\\u_2\\|\end{bmatrix}_{(3x1)}\begin{bmatrix}-&v^{T}_2&-\end{bmatrix}_{(1x3)}\ \ \ \ \ (3)$$
#
# Because those two elements will create two matrices shaped $(3,3)$ that are going to be summed up. And remember that the Singular Values $\sigma_{i,i}$ get progressively smaller in magnitude as i increases (as we slide down the diagonal).
# \
# \
# **Interpretation: We can think of this algorithm as decomposing a matrix in many overlapping frames (matrices). Where the first frame will give us many infos about our matrix, and every frame stacked on top (matrix added after the first one) gives us progressively less and less infos.**
# **PCA** refers instead then to the **"truncation"** of those three matrices $\mathrm{U}\ \mathrm{\Sigma}\ \mathrm{V^{T}}$ to obtain an approximation of the information contained in $\mathrm{A}$, given by the features that most contribute to its formation. Or alternatively, means: How many of the trailing elements (layers) in (3) can I cut off and still maintain a good amount of information from my initial matrix? The phenomena begs the question **at what index should I truncate?** Let's see all of it in practice. We will try to use SVD to compress an image, and we'll check whether **Pareto's Principle** holds true (80% of consequences come from 20% of the causes), in our case would be the statement: a great deal of information about a picture is given by a small fraction of the numbers used to create it.
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
import cv2
# Let's import the image which we'll use as an example through the notebook. For simplicity I will consider a B&W image, characterized by a matrix of 8-bit integers $a_{i,j} \in \{0, ..., 255\}$ stored in a matrix $\mathrm{A}$
A = cv2.imread('dudepic.jpg',0)
plt.imshow(A, cmap='gray', )
plt.axis('off')
print(f'Height of image: {A.shape[0]} px\nWidth of the image: {A.shape[1]} px')
# The height and width, i.e. the amount of pixels in height and width composing the picture can be seen as the entries of a matrix having those same dimensions.
# Here I will build my PCA algorithm, taking as input the matrix of data $\mathrm{A}$, computing its SVD, truncating it, and then recomputing the approximated output $\mathrm{\hat{A}}$.
# +
def reduce_dim(inp, n_dims = 20):
U,S,Vh = inp
#Truncation happens in the following line!! According to the variable n_dims
return U[:,:n_dims] @ (np.eye(n_dims) * S[:n_dims]) @ Vh[:n_dims,:]
#Shapes multiplied are (M x k) (k x k) (k x N) -> (M x N), size of the original matrix!
def PCA_algorithm(image, n_dims = 20):
assert isinstance(image, numpy.ndarray)
num_data,dim = image.shape
U, S, Vh = np.linalg.svd(image)
return reduce_dim((U, S, Vh), n_dims)
# -
output = PCA_algorithm(A)
# I will now proceed to plot the several outputs truncating at different dimensions. I would expect the image to get progressively better as we keep on adding dimensions.
fig, axarr = plt.subplots(2, 3)
fig.set_size_inches(12, 10)
number_dimensions = [5,15,30,100,150,300]
idx=0
for j in range(2):
for i in range(3):
img_plot = PCA_algorithm(A, n_dims=number_dimensions[idx])
axarr[j, i].imshow(img_plot)
axarr[j, i].axis('off')
axarr[j, i].set_title(f'{number_dimensions[idx]} dimensions')
idx+=1
# As we can see we have an image we can recognize almost immediately, keeping only 10 (!!) dimensions out of 300. We progressively get better, and at 100 (1/3 of the dimensions) we have a pretty good picture already. The rest is just adding the icing on top of the cake and the fine details.
# ## Best Choice for the number of dimensions
# It boils down to answering the question: How many number of dimensions should I keep in my model?\
# It seems clear even by the example of the image that very few dimensions contain lots of information about the model, and as we include more of them, the marginal information added by the inclusion of another dimension is progressively smaller. As we can see from the log of the Singular Values' graph.
def PCA_algorithm(image):
assert isinstance(image, numpy.ndarray)
num_data,dim = image.shape
U, S, Vh = np.linalg.svd(image)
return S
importance = np.log(pd.Series(PCA_algorithm(A))).reset_index().rename(columns={0:'vals'})
sns.lineplot(data=importance, x='index', y='vals')
# And the question remains, which can be translated to a tradeoff between **Complexity** and **Accuracy** of the model. Some researches came up with answers to that same question, in different ways. In particular, two papers are the relevant literature when it comes to this issue (SkLearn by default proposes the latter one):
# <ul>
# <li>The Optimal Hard Threshold for Singular Values is $\frac{4}{\sqrt{3}}$. <NAME> and <NAME>, 2014
# <li>Automatic choice of dimensionality for PCA. TP Minka, 2000
# <ul>
#
|
PCA picture/SVD_and_PCA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas Challenge
# ## HeroesOfPymoli_starter
# Dependencies and Set Up
import pandas as pd
import os
#Importing File
pd.read_csv("purchase_data.csv")
#Set Up purchase_data to store data
purchase_data_pd = pd.read_csv("purchase_data.csv")
purchase_data_pd.head()
# Review Dataframe format
purchase_data_pd.dtypes
#Format Purchase ID to Integer
purchase_id_pd =purchase_data_pd["Purchase ID"].astype("int")
purchase_id_pd.head()
# # Player Count
#
# - Display the total number of players
#Display total player
total_player = purchase_data_pd["SN"].value_counts().count()
players_df = pd.DataFrame({"Total Players": [total_player]})
players_df
#Purchasing Analysis (Total)
purchase_data_pd.describe()
# Calculate unique items
unique_items = purchase_data_pd["Item ID"].value_counts().count()
unique_items
# Calculate total revenue
total_revenue = purchase_data_pd["Price"].sum()
total_revenue
#Run Basic Calculation
number_purchases = purchase_data_pd["Purchase ID"].value_counts().sum()
number_purchases
#Calculate average price
average_price = purchase_data_pd["Price"].mean()
average_price
# ## Purchasing Analysis (Total)
#
# - Run basic calculations to obtain number of unique items, average price, etc.
#
#
# - Create a summary data frame to hold the results
#
#
# - Displayed data cleaner formatting
#
#
# - Display the summary data frame
#Create Summary Frame to hold results
basic_calculations_df = pd.DataFrame({"Number Unique Items": [unique_items],"Average Price": [average_price],"Number of Purchases":[number_purchases], "Total Revenue":[total_revenue]})
basic_calculations_df.head().style.format({"Average Price": "${:,.2f}","Total Revenue": "${:,.2f}"})
# Display Gender and Player Columns
purchase_data_pd.columns
#Create a new dataframe to hold gender and students
gender_students_df = purchase_data_pd.loc[:, ["SN", "Gender"]]
gender_students_df.head()
#Check for duplicates
duplicate_students = gender_students_df[gender_students_df.duplicated()].count()
duplicate_students
# ## Gender Demographics
#
# - Percentage and Count of Male Players
# - Percentage and Count of Female Players
# - Percentage and Count of Other / Non-Disclosed
# Use Gender as index and extract unique values
grouped_gender_df = gender_students_df.groupby(["Gender"])
unique_gender= grouped_gender_df.nunique()
unique_gender["SN"]
# create a new column and set it equal the the output of the calculation that we'd like to perform
unique_gender['Percent of Players'] = (unique_gender["SN"]/ total_player*100).map("{:,.2f}%".format)
unique_gender.head()
#Rename SN Column
unique_gender = unique_gender.rename(columns={"SN": "Total Count"})
unique_gender[["Total Count", "Percent of Players"]].head()
# Extract purchase count per gender
grouped_gender_df = purchase_data_pd.groupby(["Gender"])
purchase_gender = grouped_gender_df["Purchase ID"].nunique()
purchase_gender
#Create Purchase Gender into Dataframe
purchase_gender_df = pd.DataFrame(purchase_gender)
purchase_gender_df = purchase_gender_df.rename(columns={"Purchase ID": "Purchase Count"})
purchase_gender_df
#Calculate Total Purchase Value
value_gender = grouped_gender_df["Price"].sum()
value_gender
#Calculate Average Purchase Price
average_price = value_gender / purchase_gender
average_price.head()
# Add Average Purchase Price to Dataframe
purchase_gender_df['Average Purchase Price'] = average_price.map("${:,.2f}".format)
purchase_gender_df
# ## Purchasing Analysis (Gender)
#
# - Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
# - Create a summary data frame to hold the results
# Add Total Purchase Value to Dataframe
purchase_gender_df['Total Purchase Value'] = value_gender.map("${:,.2f}".format)
purchase_gender_df
#Create variable for numbers per gender
grouped_gender_df = gender_students_df.groupby(["Gender"])
unique_gender= grouped_gender_df.nunique()
gender_numbers = unique_gender["SN"]
gender_numbers
# Add Total Purchase Value to Dataframe
purchase_gender_df['Avg Total Purchase per Person'] = (value_gender/gender_numbers).map("${:,.2f}".format)
purchase_gender_df
# ## Age Demographics
#
# - Establish bins for ages
# - Categorize the existing players using the age bins. Hint: use pd.cut()
# - Calculate the numbers and percentages by age group
# - Create a summary data frame to hold the results
#
#Create Data frame for existing players by age
age_players_df = purchase_data_pd.loc[:, ["SN", "Age"]]
age_players_df.head()
# ## Purchasing Analysis (Age)
#
# - Bin the purchase_data data frame by age
#
# - Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
#
# - Create a summary data frame to hold the results
#
#Print min and max
print(age_players_df["Age"].max())
print(age_players_df["Age"].min())
#Create bins for ages
bins = [0,9,14,19,24,29,34,39,45]
#Create the name fo the bins
label_names = ["<10", "10-14", "15-19", "20-24", "25-29","30-34","35-39","40+"]
#Categorize the existing players using the age bins
age_players_df["Age Group"] = pd.cut(age_players_df["Age"], bins, labels=label_names)
age_players_df.head
#Group by group age
age_group = age_players_df.groupby("Age Group")
age_count = age_group['SN'].nunique()
age_demographics_df = pd.DataFrame(age_count)
age_demographics_df.head()
#Rename SN Column
age_demographics_df= age_demographics_df.rename(columns={"SN": "Total Count"})
age_demographics_df.head()
#Calculate Percentage of Players
age_demographics_df['Percent of Players']= (age_count/total_player*100).map("{:,.2f}%".format)
age_demographics_df.head(8)
#Create reduce dataframe
reduce_players_df = purchase_data_pd.loc[:, ["SN", "Age","Price", "Item ID", "Purchase ID"]]
reduce_players_df.head()
#Create bins for ages
bins = [0,9,14,19,24,29,34,39,45]
#Create the name fo the bins
label_names = ["<10", "10-14", "15-19", "20-24", "25-29","30-34","35-39","40+"]
#Categorize the existing players using the age bins
reduce_players_df["Age Group"] = pd.cut(reduce_players_df["Age"], bins, labels=label_names)
reduce_players_df
#Age Group by Purchase ID
age_price_group = reduce_players_df.groupby("Age Group")
item_count = age_price_group['Purchase ID'].nunique()
item_count
#Create Dataframe for Purchase Analysis
purchase_analysis_df = pd.DataFrame(item_count)
purchase_analysis_df.head()
#Calculate Total Purchase Value
value_age = age_price_group["Price"].sum()
value_age
#Calculate Average Purchase Price
average_age = value_age / item_count
average_age.head()
# Add Average Purchase Price to Dataframe
purchase_analysis_df['Average Purchase Price'] = average_age.map("${:,.2f}".format)
purchase_analysis_df
# Add Total Purchase Price to Dataframe
purchase_analysis_df['Total Purchase Price'] = value_age.map("${:,.2f}".format)
purchase_analysis_df
# Add Avg Total Purchase per Person to Dataframe
purchase_analysis_df['Total Purchase per Person'] = (value_age/age_count).map("${:,.2f}".format)
purchase_analysis_df
# ### Run basic calculations to obtain the results in the table below
#
#
# - Create a summary data frame to hold the results
# - Sort the total purchase value column in descending order
#
# #### Calculations
# Total Price per SN
sn_total= reduce_players_df.groupby("SN").sum()["Price"].rename("Total Purchase Value")
# Average Price per SN
sn_average= reduce_players_df.groupby("SN").mean()["Price"].rename("Average Purchase Price")
# Count Price per SN
sn_count= reduce_players_df.groupby("SN").count()["Price"].rename("Purchase Count")
# #### Create DataFrame
sn_df= pd.DataFrame({"Total Purchase Value": sn_total, "Average Purchase Price": sn_average, "Purchase Count": sn_count})
# #### Sort the total purchase value column in descending order
sorted_sn= sn_df.sort_values("Total Purchase Value", ascending= False)
sorted_sn["Total Purchase Value"].max
# #### Map Total Purchase Value and Average Purchase Price to Currency
sorted_sn["Total Purchase Value"]= sorted_sn["Total Purchase Value"].map("${:,.2f}".format)
sorted_sn["Average Purchase Price"]= sorted_sn["Average Purchase Price"].map("${:,.2f}".format)
# ### Display only top 5
sorted_sn.head()
# ## Most Popular Items
#
# - Retrieve the Item ID, Item Name, and Item Price columns
#
# - Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value
#
# - Create a summary data frame to hold the results
#
# - Sort the purchase count column in descending order
# #### Retrieve the Item ID, Item Name, and Item Price columns
items_pd= purchase_data_pd.loc[:,["Item ID", "Item Name", "Price"]]
# #### Calculations
## Total Price per id
id_total= items_pd.groupby(["Item ID", "Item Name"]).sum()["Price"].rename("Total Purchase Value")
## Average Price per id
id_average= items_pd.groupby(["Item ID", "Item Name"]).mean()["Price"].rename("Item Price")
## Number of purchases per id
id_count= items_pd.groupby(["Item ID", "Item Name"]).count()["Price"].rename("Purchase Count")
# #### Create DataFrame
id_df= pd.DataFrame({"Purchase Count": id_count,"Total Purchase Value":id_total, "Item Price": id_average })
# #### Sort Values by Purchase Count
sorted_id_df= id_df.sort_values("Purchase Count", ascending= False)
sorted_id_df.head()
# #### Map Total Purchase Value and Average Purchase Price to currency
sorted_id_df["Total Purchase Value"]= sorted_id_df["Total Purchase Value"].map("${:,.2f}".format)
sorted_id_df["Item Price"]= sorted_id_df["Item Price"].map("${:,.2f}".format)
# #### Display top 5 Items
top_5= sorted_id_df[:5]
top_5
# #### Most Profitable Items
#
# - Sort the above table by total purchase value in descending order
prof_items= id_df.sort_values("Total Purchase Value", ascending= False)
prof_items["Total Purchase Value"]= prof_items["Total Purchase Value"].map("${:,.2f}".format)
prof_items["Item Price"]= prof_items["Item Price"].map("${:,.2f}".format)
top_5_prof = prof_items.head()
top_5_prof
|
HeroesOfPymoli/HeroesOfPymoli.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: xopt
# language: python
# name: xopt
# ---
# Useful for debugging
# %load_ext autoreload
# %autoreload 2
# # Xopt class, TNK test function
#
# This is the class method for running Xopt. Use Bayesian exploration to explore the input space.
#
# TNK function
# $n=2$ variables:
# $x_i \in [0, \pi], i=1,2$
#
# Objectives:
# - $f_i(x) = x_i$
#
# Constraints:
# - $g_1(x) = -x_1^2 -x_2^2 + 1 + 0.1 \cos\left(16 \arctan \frac{x_1}{x_2}\right) \le 0$
# - $g_2(x) = (x_1 - 1/2)^2 + (x_2-1/2)^2 \le 0.5$
# +
# Import the class
from xopt import Xopt
import os
SMOKE_TEST = os.environ.get('SMOKE_TEST')
# -
# The `Xopt` object can be instantiated from a JSON or YAML file, or a dict, with the proper structure.
#
# Here we will make one
# +
import yaml
# Make a proper input file.
YAML="""
xopt: {output_path: null, verbose: true}
algorithm:
name: bayesian_exploration
options:
n_initial_samples: 5
n_steps: 25
verbose: True
generator_options: ## options for bayesian exploration acquisition function
batch_size: 1 ## batch size for parallelized optimization
#sigma: [[0.01, 0.0], [0.0,0.01]] ## proximal biasing term
use_gpu: False
simulation:
name: test_TNK
evaluate: xopt.evaluators.test_TNK.evaluate_TNK
vocs:
name: TNK_test
description: null
simulation: test_TNK
templates: null
variables:
x1: [0, 3.14159]
x2: [0, 3.14159]
objectives: {y1: None}
constraints:
c1: [GREATER_THAN, 0]
c2: ['LESS_THAN', 0.5]
linked_variables: {}
constants: {a: dummy_constant}
"""
config = yaml.safe_load(YAML)
if SMOKE_TEST:
config['algorithm']['options']['n_steps'] = 3
config['algorithm']['options']['generator_options']['num_restarts'] = 2
config['algorithm']['options']['generator_options']['raw_samples'] = 2
# -
X = Xopt(config)
X
# # Run BayesOpt
# +
# Pick one of these
from concurrent.futures import ThreadPoolExecutor as PoolExecutor
#from concurrent.futures import ProcessPoolExecutor as PoolExecutor
executor = PoolExecutor()
# This will also work.
#executor=None
# -
# Change max generations
X.run(executor=executor)
# # Plot
# - plot input space samples -> yellow points satisfy constraints and purple points do not
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# plot exploration results and path - exploration should explore the feasible region of the TNK problem - See Table V in https://www.iitk.ac.in/kangal/Deb_NSGA-II.pdf
fig, ax = plt.subplots()
results = X.results
print(results.keys())
variables = results['variables']
valid = results['variables'][results['feasibility'].flatten()]
ax.plot(variables[:, 0], variables[:, 1], '-o', label = 'all')
ax.plot(valid[:, 0], valid[:, 1], 'o', label = 'valid')
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.legend()
plt.show()
# -
|
examples/bayes_exp/xopt_class_tnk_exp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Git Stash
# Before you can `git pull`, you need to have committed any changes you have made. If you find you want to pull, but you're not ready to commit, you have to temporarily "put aside" your uncommitted changes.
# For this, you can use the `git stash` command, like in the following example:
# +
import os
top_dir = os.getcwd()
git_dir = os.path.join(top_dir, "learning_git")
working_dir = os.path.join(git_dir, "git_example")
os.chdir(working_dir)
# -
# Remind ourselves which branch we are using:
# + language="bash"
# git branch -vv
# +
# %%writefile Wales.md
Mountains In Wales
==================
* Pen y Fan
* Tryfan
* Snowdon
* <NAME>
* Fan y Big
* <NAME>
* Penygader
# + language="bash"
# git stash
# + language="bash"
# git pull
# -
# By stashing your work first, your repository becomes clean, allowing you to pull. To restore your changes, use `git stash apply`.
# + language="bash"
# git stash apply
# -
# The "Stash" is a way of temporarily saving your working area, and can help out in a pinch.
# # Tagging
#
# Tags are easy to read labels for revisions, and can be used anywhere we would name a commit.
#
# Produce real results *only* with tagged revisions.
#
# NB: we delete previous tags with the same name remotely and locally first, to avoid duplicates.
# + [markdown] attributes={"classes": [" Bash"], "id": ""}
# ``` Bash
# git tag -a v1.0 -m "Release 1.0"
# git push --tags
# ```
# -
# You can also use tag names in the place of commmit hashes, such as to list the history between particular commits:
# ``` Bash
# git log v1.0.. --graph --oneline
# ```
# If .. is used without a following commit name, HEAD is assumed.
# # Working with generated files: gitignore
# We often end up with files that are generated by our program. It is bad practice to keep these in Git; just keep the sources.
# Examples include `.o` and `.x` files for compiled languages, `.pyc` files in Python.
# In our example, we might want to make our .md files into a PDF with pandoc:
# +
# %%writefile Makefile
MDS=$(wildcard *.md)
PDFS=$(MDS:.md=.pdf)
default: $(PDFS)
%.pdf: %.md
pandoc $< -o $@
# + language="bash"
# make
# -
# We now have a bunch of output .pdf files corresponding to each Markdown file.
# But we don't want those to show up in git:
# + language="bash"
# git status
# -
# Use .gitignore files to tell Git not to pay attention to files with certain paths:
# %%writefile .gitignore
*.pdf
# + language="bash"
# git status
# + language="bash"
# git add Makefile
# git add .gitignore
# git commit -am "Add a makefile and ignore generated files"
# git push
# -
# # Git clean
# Sometimes you end up creating various files that you do not want to include in version control. An easy way of deleting them (if that is what you want) is the `git clean` command, which will remove the files that git is not tracking.
# + language="bash"
# git clean -fX
# + language="bash"
# ls
# -
# * With -f: don't prompt
# * with -d: remove directories
# * with -x: Also remote .gitignored files
# * with -X: Only remove .gitignore files
# # Hunks
#
# ## Git Hunks
#
# A "Hunk" is one git change. This changeset has three hunks:
# + [markdown] attributes={"classes": [" diff"], "id": ""}
# ```python
# # +import matplotlib
# # +import numpy as np
#
# from matplotlib import pylab
# from matplotlib.backends.backend_pdf import PdfPages
#
# # +def increment_or_add(key,hash,weight=1):
# # + if key not in hash:
# # + hash[key]=0
# # + hash[key]+=weight
# # +
# data_path=os.path.join(os.path.dirname(
# os.path.abspath(__file__)),
# -regenerate=False
# # +regenerate=True
# ```
# -
# ## Interactive add
#
# `git add` and `git reset` can be used to stage/unstage a whole file,
# but you can use interactive mode to stage by hunk, choosing
# yes or no for each hunk.
# ``` bash
# git add -p myfile.py
# ```
# + [markdown] attributes={"classes": [" diff"], "id": ""}
# ``` python
# # +import matplotlib
# # +import numpy as np
# #Stage this hunk [y,n,a,d,/,j,J,g,e,?]?
# ```
# -
# # GitHub pages
#
# ## Yaml Frontmatter
#
# GitHub will publish repositories containing markdown as web pages, automatically.
#
# You'll need to add this content:
#
# > ```
# > ---
# > ---
# > ```
#
# A pair of lines with three dashes, to the top of each markdown file. This is how GitHub knows which markdown files to make into web pages.
# [Here's why](https://jekyllrb.com/docs/front-matter/) for the curious.
# +
# %%writefile test.md
---
title: Github Pages Example
---
Mountains and Lakes in the UK
===================
Engerland is not very mountainous.
But has some tall hills, and maybe a mountain or two depending on your definition.
# + language="bash"
# git commit -am "Add github pages YAML frontmatter"
# -
# ## The gh-pages branch
#
# GitHub creates github pages when you use a special named branch.
# By default this is `gh-pages` although you can change it to something else if you prefer.
# This is best used to create documentation for a program you write, but you can use it for anything.
os.chdir(working_dir)
# + attributes={"classes": [" Bash"], "id": ""} language="bash"
#
# git checkout -b gh-pages
# git push -uf origin gh-pages
# -
# The first time you do this, GitHub takes a few minutes to generate your pages.
#
# The website will appear at `http://username.github.io/repositoryname`, for example:
#
# http://alan-turing-institute.github.io/github-example/
# ## Layout for GitHub pages
#
# You can use GitHub pages to make HTML layouts, here's an [example of how to do it](http://github.com/UCL/ucl-github-pages-example), and [how it looks](http://ucl.github.com/ucl-github-pages-example). We won't go into the detail of this now, but after the class, you might want to try this.
# + language="bash"
# # Cleanup by removing the gh-pages branch
# git checkout main
# git push
# git branch -d gh-pages
# git push --delete origin gh-pages
# git branch --remote
|
module04_version_control_with_git/04_08_git_stash.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# Load Pictures function
def LoadImages(path):
import csv
File = []
with open(path) as csvfile:
inputfile = csv.reader(csvfile)
for row in inputfile:
File.append(row)
return File
# +
# Load sensogram function
def LoadSensogram():
import csv
# Initialize the testmatrix array
testmatrix = []
# Initialize dummy variable
num = 0
# Open text file
with open('sensogram.txt') as csvfile:
inputfile = csv.reader(csvfile)
for row in inputfile:
testmatrix.append(row)
for i in range(len(row)):
testmatrix[num][i] = int(row[i])
num = num+1
# At this point we get all values in packets of data being integer, good job, had to loop through each array
# as it comes in to convert it from a string to an integer.
return testmatrix
# +
def DisplayImages(x,test_array,picmatrix,n):
import tkinter as tk
from PIL import Image, ImageTk
master = tk.Tk()
# This line of code specifies where the window opens up and how big it is
master.geometry("1900x500+300+300")
v = tk.IntVar()
# v.set(1)
tk.Label(master,
text="""Click over the image which the pattern matches most with:""",
justify = tk.LEFT,
padx = 20)#.pack()
def ShowChoice():
# print(v.get())
master.destroy()
selection = v.get()
return selection
img = []
photo = []
# f1 = tk.Frame(master)
# f1.grid(row=2, column=5, sticky="nsew")
for i in range(n):
img.append(Image.open(','.join(picmatrix[test_array[i]])))
img[i] = img[i].resize((250,250),Image.NEAREST)
photo.append(ImageTk.PhotoImage(img[i]))
but = tk.Radiobutton(master,
text="Image",image=photo[i],
padx = 20,
variable=v,indicatoron = 0,command=ShowChoice,
value=i)#.pack(side)#=tk.LEFT)
if i < 5:
but.grid(row=0,column=i)
else:
but.grid(row=1,column=i-5)
# pack(anchor=tk.W)
# But.grid(row=0, column=i)
master.mainloop()
return v.get()
# -
def DisplayChinese(x,ChineseCharac,picmatrix,time):
import tkinter as tk
from PIL import Image, ImageTk
master = tk.Tk()
# This line closes the window after 2000ms have passed
master.after(time, lambda: master.destroy())
# This line of code specifies where the window opens up and how big it is
master.geometry("600x500+100+100")
v = tk.IntVar()
# v.set(1)
img = []
photo = []
img.append(Image.open(','.join(picmatrix[x])))
img[0] = img[0].resize((250,250),Image.NEAREST)
photo.append(ImageTk.PhotoImage(img[0]))
img.append(Image.open(','.join(ChineseCharac[x])))
img[1] = img[1].resize((250,250),Image.NEAREST)
photo.append(ImageTk.PhotoImage(img[1]))
tk.Radiobutton(master,
text="Image",image=photo[0],
padx = 20,
variable=v,indicatoron = 0,
value=1).pack(side=tk.LEFT)
tk.Radiobutton(master,
text="Chinese",image=photo[1],
padx = 20,
variable=v,indicatoron = 0,
value=2).pack(side=tk.LEFT)
# pack(anchor=tk.W)
# But.grid(row=0, column=i)
master.mainloop()
return master
# import time
# picmatrix = LoadPictures()
# for i in range(3):
# DisplayChinese(i,picmatrix,picmatrix)
# time.sleep(1)
def ActivArray(testmatrix,i):
# Initialize array to store testmatrix values
activ_array = []
# Loop through the possible number of examples (7 is the possible nodes)
for q in range(7):
if (testmatrix[i][testmatrix[i][q+7]-1] == 1): # This checks if the current node is supposed to be on/off
activ_array.append(testmatrix[i][q+7]+4) # Add position of the node to array
activ_array.append((testmatrix[i][q+14]//3)+70) # Add delay of the node to array, THIS IS FOR STRENGTH
# Print the array of data we are sending
# print(activ_array)
return activ_array
# +
# This is the function to emit to Serial but without display the pictures
def TestPattern(ser,testmatrix,i):
import struct
import time
# Read the incoming line
print(ser.readline())
while(True):
# If the incoming line matches the trigger
if (ser.readline() == b'ready\r\n'):
# if ser == 'hi':
# Then send flag to Arduino to start listening
ser.write(b'1') # Need to include the b so that data can be sent
# this is python 3 syntax
# Get the current array for the pattern
activ_array = ActivArray(testmatrix,i)
# Read length of the array
Counter = len(activ_array)
# Send length of the array to Arduino so it knows how many to wait for
ser.write(struct.pack('>B', Counter)) # Convert the integers to bits to send via Serial
print(struct.pack('>B', Counter))
# Initialize dummy variable to loop through array
c = 0
# Loop through length of array sending values
for y in range(Counter//2):
ser.write(struct.pack('>B', activ_array[c]))
ser.write(struct.pack('>B', activ_array[c+1]))
# print('Im sending %d %d' % (activ_array[c], activ_array[c+1]))
# Jump two to go to the next pair
c = c+2
# Define waiting length
# time.sleep(1)
break
else:
time.sleep(1)
return 1
# +
# This is the Testing Function and Storing values into the Results Matrix
def TestFunction(x,picmatrix,order,n,t):
import time
# So I'm allowing the user to select between 1-5, because of the way they are pasted, I know which
# picture corresponds to each number. Knowing the picture name, I can correlate to the picture that
# should have appeared with the pattern by comparing the names I guess.
# Generate the random array, choose out of 20 and pick 4
test_array = PickRandomIndexes(x,order,t)
selection = DisplayImages(x,test_array,picmatrix,n)
time.sleep(1)
TestforthePicture(test_array,x,selection)
return test_array, selection
# +
# This portion asks the user to select which picture is the one they recognize and saves the data
def TestforthePicture(test_array,num,selection):
if num == test_array[selection]:
# print('hooray')
results.append(1)
else:
# print('not yet')
results.append(0)
return results
# -
def TestChineseCharacters(x,ChineseCharac):
# img = Image.open(','.join(ChineseCharac[x]))
# return img.show()
import tkinter as tk
from PIL import Image, ImageTk
master = tk.Tk()
# This line closes the window after 2000ms have passed
master.after(2000, lambda: master.destroy())
# This line of code specifies where the window opens up and how big it is
master.geometry("600x500+0+0")
v = tk.IntVar()
# v.set(1)
img = []
photo = []
img.append(Image.open(','.join(ChineseCharac[x])))
img[0] = img[0].resize((250,250),Image.NEAREST)
photo.append(ImageTk.PhotoImage(img[0]))
tk.Radiobutton(master,
text="Chinese",image=photo[0],
padx = 20,
variable=v,indicatoron = 0,
value=2).pack(side=tk.LEFT)
# pack(anchor=tk.W)
# But.grid(row=0, column=i)
master.mainloop()
return
def PickRandomIndexes(x,order,z):
test_array = []
test_array.append(x)
a = 0
length = len(order)-1
while a < z:
# This code is only for the spanish test, where for the test I only want the same
# vowels being presented, otherwise advantage on learning
randy = random.randint(0,length)
a = a+1
# if randy in test_array:
if order[randy] in test_array:
# print('one down')
a = a-1
else:
test_array.append(order[randy])
random.shuffle(test_array)
# print(test_array)
return test_array
def DicPickRandomIndexes(x,order,z):
def get_digit(number, n):
return number // 10**n % 10
test_array = []
test_array.append(x)
a = 0
b = 0
length = len(order)-1
while a < z:
# This code is only for the spanish test, where for the test I only want the same
# vowels being presented, otherwise advantage on learning
order_min = min(order)
order_max = max(order)
number = get_digit(x,0)
if number > 5:
number = number-5
# print('this is the number: %d' %number)
order_min_num = get_digit(order_min,0)
add_num = number - order_min_num
new_min = order_min + add_num
randy = new_min + b
#### ------------ ####
# randy = random.randint(0,length)
a = a+1
if randy in test_array:
# if order[randy] in test_array:
# print('one down')
a = a-1
b = b+5
else:
# test_array.append(order[randy])
test_array.append(randy)
random.shuffle(test_array)
return test_array
def DetailedInfo(results,order,x):
if x == 1:
for i in range(len(order)):
activ_array = ActivArray(testmatrix,order[i])
print(results[i],picmatrix[order[i]],len(activ_array))
elif x == 0:
for i in range(len(order)):
print(results[i],picmatrix[order[i]])
return
# DetailedInfo(results,order)
def PrintScore(results):
print('%d correct out of %d ' % ((sum(results),len(results))))
return
|
_Functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="./intro_images/MIE.PNG" width="100%" align="left" />
# <table style="float:right;">
# <tr>
# <td>
# <div style="text-align: right"><a href="https://alandavies.netlify.com" target="_blank">Dr <NAME></a></div>
# <div style="text-align: right">Senior Lecturer health data science</div>
# <div style="text-align: right">University of Manchester</div>
# </td>
# <td>
# <img src="./intro_images/alan.PNG" width="30%" />
# </td>
# </tr>
# </table>
# # 6.0 Using SQL in Python
# ****
# #### About this Notebook
# This notebook introduces the concept of using SQL in another programming language (Python). This includes using the SQL directly or via Object Relational Mapping (ORM).
# <div class="alert alert-block alert-warning"><b>Learning Objectives:</b>
# <br/> At the end of this notebook you will be able to:
#
# - Investigate key features of applying SQL in Python
#
# - Explore and practice using ORM in Python
#
# </div>
# <a id="top"></a>
#
# <b>Table of contents</b><br>
#
# 6.1 [Using SQL in Python](#sql)
#
# 6.2 [Object-Relational Mapping](#orm)
# We have looked at Python in previous notebooks and in this series at using SQL. Here we look at how we can use both Python and SQL together. This is typical of many modern programs that use databases at their core. Instead of using the SQL inline in the notebook we will now use the Python <code>sqlite3</code> library to connect to a database file.
# <div class="alert alert-success">
# <b>Note:</b> The previous knowledge gained about how SQL works will help make more sense of how to use SQL in Python. In some cases it may be easier to first write the code in SQL and transfer it into Python until you become more familiar with the Python libraries.
# </div>
# <a id="sql"></a>
# #### 6.1 Using SQL in Python
# +
import sqlite3
med_data_db = sqlite3.connect("medical_db.db")
cursor = med_data_db.cursor()
cursor.execute("SELECT * FROM med_data;")
results = cursor.fetchall()
for result in results:
print(result)
cursor.close()
med_data_db.close()
# -
# There is a lot going on here. Firstly I made a database file called <code>medical_db.db</code> and I recreated the <code>med_data</code> table we created in previous examples in that file. Next we load the <code>sqlite3</code> library. We then make a connection to the database using a variable I am calling <code>med_data_db</code>. We then make a <code>cursor</code>. This essentially lets us iterate over a set of records. We then use this to execute an SQL command. The command <code>SELECT * FROM med_data;</code> should be familiar to you by now. We then store the results in a variable and iterate through them in a loop. Finally we close the cursor and the connection to the database when we are done.
# Now we have the data stored in the <code>results</code> variable, we can access it like a regular Python data structure. So I can view all of the record or a specific part:
print(results[0])
print(results[0][1])
print("Patient:", results[0][1], "heart rate =" ,results[0][5])
# <div class="alert alert-danger">
# <b>Note:</b> We wouldn't typically try to read a whole database into a single variable. This might not be possible with a very large database and also would be very inefficient. Instead we would perform a query to return a subset of the data we are interested in using.
# </div>
# This lets us integrate data stored in a database with the programs we create. Hopefully you can see how useful such methods are for building a complete application that interacts with data, which is a core component of software systems and used frequently in both data science and informatics projects.
# + [markdown] solution2="hidden" solution2_first=true
# <div class="alert alert-block alert-info">
# <b>Task 1:</b>
# <br>
# Print out the blood pressure for patient <code><NAME></code> with the patients name and unit of measure (mmHG).
# </div>
# + solution2="hidden"
print(results[3][1], "blood pressure =", results[3][4], "mmHG")
# -
# Type your code here
# + [markdown] solution2="hidden" solution2_first=true
# <div class="alert alert-block alert-info">
# <b>Task 2:</b>
# <br>
# Write a query to return all the patients details for patients over the age of 50 years. Print out the returned values with the Python <code>print()</code> function.
# </div>
# + solution2="hidden"
med_data_db = sqlite3.connect("medical_db.db")
cursor = med_data_db.cursor()
cursor.execute("SELECT * FROM med_data WHERE Age > 50;")
results = cursor.fetchall()
for result in results:
print(result)
cursor.close()
med_data_db.close()
# -
# Type your code here
# <a id="orm"></a>
# #### 6.2 Object-Relational Mapping
# The only issue with this approach is that you tend to have a mixture of Python and SQL code intermixed in your code. This also requires you to be proficient in both SQL and Python and can also be harder to maintain. One way to overcome this is to use a technique called <code>Object-Relational Mapping (ORM)</code>. This works by converting the data using Object Orientated Programming techniques to make the conversion between otherwise incompatible systems. The mapping process maps tables in an SQL database to objects in Python.
# The figure below shows how this works. From the programmers perspective, they only ever deal with writing to and from Python objects (or whatever language they are using). The Object mapper converts this to SQL and queries the database returning data and converting it into objects. This way a developer doesn't need to mix SQL and Python in the same program. This also makes it easier to swap out database engines (say you wanted to change form SQLite to MySQL for instance) without rewriting large amounts of code.
# <img src="./intro_images/orm.PNG" width="100%" />
# Lets look at an example. First we will make the object to store the data in. We make one object per table in the database.
# +
from sqlalchemy import create_engine, MetaData, Table, Column, Integer, String, select
engine = create_engine('sqlite:///medical_db.db', echo = False)
meta = MetaData()
med_data = Table(
'med_data', meta,
Column('id', Integer, primary_key = True),
Column('Name', String),
Column('Age', Integer),
Column('Sex', String),
Column('Blood pressure', String),
Column('Heart rate', String),
)
meta.create_all(engine)
# -
# Here we import the <code>sqlalchemy</code> library and then load the database. We use the <code>Table</code> and <code>Column</code> functions to create an object to store the data from the SQL table along with its associated data type. Note that we are using an <code>SQLite</code> engine. We could swap this for a different vendor i.e. PostgreSQL or MySQL.
# <div class="alert alert-success">
# <b>Note:</b> By setting <code>echo = False</code> in the <code>create_engine</code> function, we just see the output. If we set it to <code>True</code>, we would also see the SQL statements it is executing behind the scenes. If you want to see what that looks like, change to <code>echo = True</code> and rerun the cell.
# </div>
# +
data = med_data.select()
conn = engine.connect()
result = conn.execute(data)
for each_row in result:
print(each_row)
# -
# we can then create a command and execute it. We store the result in a variable that we can iterate through as before. Now we can perform queries just using Python code. This keeps all our code in a project in the same language. Below we see an example query to find all the females in the table (note the use of the Python double equal operator for equality).
# +
females = med_data.select().where(med_data.c.Sex == 'F')
result = conn.execute(females)
for each_row in result:
print(each_row)
# -
# This query returns the 2 records where the <code>Sex</code> is female.
# + [markdown] solution2="hidden" solution2_first=true
# <div class="alert alert-block alert-info">
# <b>Task 3:</b>
# <br>
# 1. Using the <code>sqlalchemy</code> library, create a table object for the <code>drug_table</code> table that contains the following fields.<br />
# <code>
# id (primary key),
# medication (string),
# route (string),
# dose (string),
# patient_id (int),
# freq per day (int)
# </code>
# <br />
# 2. Output the results of the table (print the entire table).<br />
# 3. Write the following queries:<br />
# <ul>
# <li>Return all records containing the drug <code>WARFARIN</code></li>
# <li>Return all records where the frequency per day is more than 0 and less than 3</li>
# </ul>
# <br />
# Don't worry about making <code>patient_id</code> a foreign key for now, just treat it like a regular column.<br />
# <br />
# <strong>HINT:</strong> for the last task you will need to lookup and use the <code>and_</code> function (<code>from sqlalchemy import and_)</code>. Also for accessing columns that have spaces in their name, you would write <code>drug_table.c['freq per day']</code>.
# </div>
# + solution2="hidden"
drug_table = Table(
'drug_table', meta,
Column('id', Integer, primary_key = True),
Column('medication', String),
Column('route', String),
Column('dose', String),
Column('patient_id', Integer),
Column('freq per day', Integer)
)
meta.create_all(engine)
# + solution2="hidden"
data = drug_table.select()
conn = engine.connect()
result = conn.execute(data)
for each_row in result:
print(each_row)
# + solution2="hidden"
warfarin = drug_table.select().where(drug_table.c.medication == 'WARFARIN')
result = conn.execute(warfarin)
for each_row in result:
print(each_row)
# + solution2="hidden"
from sqlalchemy import and_
freq = drug_table.select().where(and_(drug_table.c['freq per day'] > 0, drug_table.c['freq per day'] < 3))
result = conn.execute(freq)
for each_row in result:
print(each_row)
# -
# Type your code here
# Type your code here
# Type your code here
# Type your code here
# Another way you might see a table defined is like that below. This is now the SQLAlchemy standard and is referred to as <code>Declarative Mapping</code>.
# +
from sqlalchemy import Table, Column, Integer, String
from sqlalchemy.ext.declarative import declarative_base
Base = declarative_base()
class DrugTable(Base):
__tablename__ = 'drug_table'
id = Column(Integer, primary_key = True)
medication = Column(String)
route = Column(String)
dose = Column(String)
patient_id = Column(Integer)
freq_per_day = Column(Integer)
# -
# Here we are using the standard <code>class</code> to create a table. Each table would have a class. The table name is defined, as are the variables with the columns data types. We can now set the <code>patient_id</code> to be a <code>foreign key</code> to link the two tables.
# +
from sqlalchemy import ForeignKey
from sqlalchemy.orm import relationship
Base = declarative_base()
class MedData(Base):
__tablename__ = 'med_data'
id = Column(Integer, primary_key = True)
name = Column(String)
age = Column(Integer)
sex = Column(String)
blood_pressure = Column("Blood pressure", String)
heart_rate = Column("Heart rate", String)
drug_table = relationship("DrugTable", back_populates="med_data")
class DrugTable(Base):
__tablename__ = 'drug_table'
id = Column(Integer, primary_key = True)
medication = Column(String)
route = Column(String)
dose = Column(String)
freq_per_day = Column(Integer)
patient_id = Column(Integer, ForeignKey('med_data.id'))
med_data = relationship("MedData", back_populates="drug_table")
# -
# To create a <code>one-to-many</code> relationship we added the <code>relationship()</code> function to both the <code>MedData</code> and <code>DrugTable</code> classes. The <code>medications</code> variable creates a relationship with the <code>DrugTable</code> and the <code>med_data</code> variable links this with the <code>medications</code> variable. Note that the <code>patient_id</code> is now set as a foreign key using the <code>ForeignKey()</code> function to point to the <code>id</code> column in the <code>MedData</code> class.
# To see the different types of relationship patterns, look at this link: <a href="https://docs.sqlalchemy.org/en/13/orm/basic_relationships.html" target="_blank">SQLAlchemy relationships</a>.
# <div class="alert alert-success">
# <b>Note:</b> Be careful to refer to the class name and table name in the correct places. Mixing these up is a common cause of error and the first thing you might want to check.
# </div>
# So now we have setup the classes to store the data, lets go ahead and use them. As before we start by creating an engine with the same database file we used previously.
from sqlalchemy import create_engine
engine = create_engine('sqlite:///medical_db.db', echo = False)
# We start by making an <code>instance</code> of the mapped class.
admission_data = MedData()
medication_data = DrugTable()
# We then set up a <code>session</code>. This handles multiple users accessing our database through our application (possibly even at the same time if we have a lot of users).
from sqlalchemy.orm import sessionmaker
Session = sessionmaker(bind=engine)
session = Session()
session.add(admission_data)
# We can now query the database to return all the data.
patients = session.query(MedData).all()
for pt in patients:
print(f'{pt.name}, {pt.age}, {pt.sex}, {pt.blood_pressure}, {pt.heart_rate}')
# We can also carry out specific queries as before. For example lets get all the heart rates more than 70 bpm.
patients = session.query(MedData).filter(MedData.heart_rate > 70).all()
for pt in patients:
print(f'{pt.name}, {pt.age}, {pt.sex}, {pt.blood_pressure}, {pt.heart_rate}')
# + [markdown] solution2="hidden" solution2_first=true
# <div class="alert alert-block alert-info">
# <b>Task 4:</b>
# <br>
# Write a query like the one above to return all the patients aged 30 and under.
# </div>
# + solution2="hidden"
patients = session.query(MedData).filter(MedData.age <= 30).all()
for pt in patients:
print(f'{pt.name}, {pt.age}, {pt.sex}, {pt.blood_pressure}, {pt.heart_rate}')
# -
# Type your code here
# We can use a join to find all the patients that are taking medications ending with the letters 'IN'.
# +
hypertension_meds = session.query(MedData).join(DrugTable).filter(DrugTable.medication.ilike('%IN')).all()
for med in hypertension_meds:
print(f'{med.name}, {med.age}, {med.sex}')
# -
# <div class="alert alert-success">
# <b>Note:</b> Remember that the results returned are iterable (can be iterated over) and you may need to carry out further processing on the results to get them into the correct format for subsequent use. This is one of the most common issues with using these methods in Python that beginners tend to struggle with. Try outputting (printing) the raw results so you can see what kind of data structure they are returned in. This will give you a clue as to how to process them further to extract the exact required information.
# </div>
# Finally when we have finished interacting with our database we can close the session.
session.close()
# You have come to the end of the SQL notebooks. We have introduced you to the basic SQL commands and taken you right through to using SQL in Python with Object Relational Mapping.Modern software systems are made using multiple frameworks and libraries to help with the heavy lifting and prevent us having to reinvent the wheel. To use these frameworks and libraries, one must become familiar with their APIs (Application Programing Interfaces). The best way to do this is to start with the documentation. For example here is the documentation for <a href="https://docs.sqlalchemy.org/en/13/orm/tutorial.html#connecting" target="_blank">SQLAlchemy</a>. Sometimes people (including software developers) often find the documentation less than clear or intuitive. We get round this by searching for simple examples or tutorial blogs/videos on the internet. This is really the best way to learn, and professional software engineers and data scientists do this all the time. We recommend you start to do this if your are not already to help build on your skills and extend your knowledge.
# ### Notebook details
# <br>
# <i>Notebook created by <strong>Dr. <NAME></strong>.
# <br>
# © <NAME> 2021
# ## Notes:
|
Intro to SQL/Intro to SQL Book 6 (Using SQL in Python).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import math
month = '12'
df = pd.read_csv(r'airbikedistance\abd2019'+month+'.csv')
df.head()
df = df[['tripduration', 'tripdistance', 'starttime', 'startstationid' , 'o3']]
df.head()
def get_hour(starttime):
hour = starttime.split(' ')[1]
return hour[0:2]
df['hour'] = df.starttime.apply(lambda x: get_hour(x))
df['trips'] = df.tripduration.apply(lambda x: 1)
df.head()
df = df[['trips', 'tripdistance', 'tripduration', 'hour', 'startstationid', 'starttime', 'o3']]
df.head()
df.info()
df = df.dropna()
df.info()
df.groupby('startstationid').size()
df = df[df['startstationid'] != '-']
df['startstationid'] = df.startstationid.apply(lambda x: int(float(x)))
dfgrouped= df.groupby(['hour', 'startstationid']).aggregate({'trips': 'sum',
'tripdistance': 'sum',
'tripduration': 'sum',
'starttime': 'min',
'o3': 'mean'})
dfgrouped.head()
dfgrouped.shape
dfgrouped.to_csv(r'airbike-dataset\ny2019'+month+'.csv')
|
NY/FinalTransformation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # PYTHON CRASH COURSE
# #### Nov 23, 2021
# #### Author: <NAME>
# ### STRING
# - formatting
# - immutable
# - slicing
num = 12
name= 'Chaithra'
'my num is {} and my name is {}'.format(num, name)
'my num is {one} and my name is {two}, more {one}'.format(one=num, two=name) # more suggested way
# no need to worry about formatting being in exact same order
var = 'chaithra kopparam cheluvaiah'
var[5] # indexing the string
var[0:] # all the characters starting from index 0
var[3:5] # all the characters starting from index 3 - 4
var[:] # all the characters
var[0] = 'z' # strings are immutable
var[::-1] # reversing the strinng
var[::-2] # keeps subtracting step size 2 from the end index
# var = 'chaithra kopparam cheluvaiah'
var[::3] # keeps adding step size 3 to the start index
# ### LIST
# - slicing
# - mutable
# - nested list
my_list = ['a','b','c','d','e']
my_list[1:4] # list slicing
my_list[1:5] # last index does not exist but it doesnt give any error
my_list[0] = 'New' # list is mutable
my_list
nested_list = [1,2,[3,4]]
nested_list[2]
nested_list[2][0] # in numpy, this syntax is used for 2d array selection
deep_nested = [1,2,[3,4,['target']]]
deep_nested[2]
deep_nested[2][2] # note that output is still a list with single element
deep_nested[2][2][0]
# ### DICTIONARIES
# - nested dicts
# - keys are immutable
d = {'key1':'v1', 'k2':'v2', 'k3':123}
# indexing dictionary
d['key1']
d['k3']
# dictionaries can take in any items as their values
d = {'k1':[1,2,3]}
d['k1']
d['k1'][0]
my_list = d['k1'] # better coding
my_list[0]
# nested dictionaries
d = {'k1':{'innerkey':[1,2,3]}}
# lets say, we want to access 2 from the list
d['k1']['innerkey'][1]
# ### TUPLE
# - Immutable
# tuple
t = (1,2,3)
# indexing tuple
t[0]
t[0] = 'NEW'
# ### SET
# - creating set from list
# - add()
{1,2,3,1,1,1,1,1,2,3,4} # keeps unique values
set([1,2,3,1,1,1,1,1,2,3,4]) # passing list to set constructor to grab unique elements
# add items to set
s = {1,2,3}
s.add(5)
s
s.add(5) # it wont retreive error
s # it just keeps unique elements
# ## Logical Operators
1 < 2
1 >= 2
1 == 1
1 == 2
1 != 3
'hi' == 'bye'
'hi' != 'bye'
(1 < 2) and (2 < 3) # paranthesis makes it more readable
(1 > 2) and (2 < 3)
# ### CONDITIONAL STATEMENTS
if 1 < 5:
print('yep!')
if 1 == 2:
print('First')
elif 3 == 3:
print('Middle')
else:
print('Last')
if 1 == 2:
print('First')
elif 4 == 4:
print('second') # it is going to execute only this block and exit
#even though other statements might be true below
elif 3 == 3:
print('Middle')
else:
print('Last')
# ### LOOPS
# - for
# - while
seq = [1,2,3,4,5]
for item in seq:
print(item)
# +
i = 1
while i < 5:
print('i is {}'.format(i))
i = i + 1
# -
# ### RANGE
range(0,5) # returns range object
list(range(5))
list(range(0,5)) # 0 is redundant
for num in range(7,5): # no error even though end < start. for loop will not run
print(num)
for num in range(7,10):
print(num)
# ### LIST COMPREHENSION - FOR LOOP BUT BACKWARDS
x = [1,2,3,4,5]
out = []
for num in x:
out.append(num **2)
out
[num**2 for num in x] # for loop but backwards - reduced lines of code : can be done using map() also
out = [num**2 for num in x]
out
# ### FUNCTIONS
def my_func(name): # function name starts with lower case letters
print('Hello, '+name)
my_func('Chaithra')
def my_func(name='Default Name'): # if you want default value to one of the parameters
print('Hello, '+name)
my_func()
my_func(name='Chai') # you can fully explain what you are passing to function
my_func # function object will be returned
# function returning a value
def square(num):
return num ** 2
output = square(2)
output
# functions have documentation string using triple enlosing quote
# triple enclosing quotes basically allows you to put in giant string
def square(num):
"""
This is a docstring.
can go multiple lines.
This function squares a number
"""
return num ** 2
square # shift + Tab to see the signature and docstring
# ### MAP & FILTER
def times2(num): return num * 2
# converting to lambda ( anonymous functions)
# remove redudant keywords - def, fucntion name, return
t = lambda num: num*2
t(12)
times2(6)
seq = [1,2,3,4,5]
map(times2, seq)
list(map(times2, seq)) # casting to list
list(map(lambda num: num*2, seq))
# filter for even numbers in the seq
filter(lambda num: num%2==0, seq)
list(filter(lambda num: num%2==0, seq))
# ### METHODS
# #### STRING METHODS
# - upper
# - lower
# - split
s = 'hello my name is chaithra'
s.lower()
s.upper()
s.split() # useful for text analysis
# +
# default separator is any whitespace.
txt = "welc\nome\n to the jungle" # delimiter is newline and space
x = txt.split()
print(x)
# -
tweet = 'Go Sports! #Sports'
tweet.split('#')
# #### DICTIONARY METHODS
# - keys
# - values
# - items
d = {'k1':1, 'k2':2}
d.keys()
d.items()
d.values()
vals = d.values() # dict_values object cannot be indexed
vals[0] # cannot be indexed
vals = list(d.values())
vals[0]
# #### LIST METHODS
# - pop
# - pop with index
# - append
lst = [1,2,3]
lst.pop() # change is permanent
lst
lst = [1,2,3,4,5]
item = lst.pop()
item
first = lst.pop(0) # pop with index
print(lst)
print(first)
lst.append('NEW') # append new element to end of the list
lst
# ### IN
'x' in [1,2,3,4,5]
'x' in ['x','y','z']
'dog' in 'martha is a dog!'
# #### TUPLE UNPACKING
x = [(1,2),(3,4),(5,6)]
x[0]
x[0][1]
for item in x:
print(item)
# tuple unpacking works when iterating over list of tuples
for (a,b) in x:
print(a)
for a,b in x: # paranthesis are optional
print(b)
# ### STRING FORMATTING IN JPMC INTERVIEW
# when we want to have specific decimal places
print('%.2f'%39.5)
print('%.2f'%39)
# {[argument_index_or_keyword]:[width][.precision][type]}
'{:.2f}'.format(38.9)
'{:.1f}'.format(38.28) # rounds off
'{num:.2f}'.format(num=38.9)
'{num:10.2f}'.format(num=38.9)
|
S04PythonCrashCourse/L01PythonCrashCourse.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="iCm2sd89tOdP"
# # Demo: Defining Control_M Workflows using Python
# + [markdown] id="dornvvu0tYxj"
# # Step 1 - Setup
# + [markdown] id="GKkmksYi1-Kn"
# ## Step 1A - Install the library
# -
# !pip --version
# !pip install git+https://github.com/tadinve/naga.git
# + id="6iwRhegFtcXJ"
from ctm_python_client.core.bmc_control_m import CmJobFlow
from ctm_python_client.jobs.dummy import DummyJob
# + [markdown] id="TfK9UygIu3Qg"
# # Step 2 - Instantiate, Authenticate and Schedule
#
# + [markdown] id="N6CwrQJ5Lxz1"
# ## Step 2A - Create the object
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="FDszp8dtRJi_" outputId="703ac9a8-eb2e-4840-bfc1-10d62fa4315f"
# Please change the URfrI, and ctm_user and enter ctm_password to match your environment
from ctm_python_client.session.session import Session
import getpass
ctm_uri = "https://acb-rhctmv20.centralus.cloudapp.azure.com:8443/automation-api"
ctm_user = "vtadinad"
ctm_pwd = "<PASSWORD>"
if "ctm_pwd" not in locals(): # has not been enterd once, will skip next time
ctm_pwd = <PASSWORD>pass.getpass("Enter your Control M Password ")
session = Session(endpoint=ctm_uri, username=ctm_user, password=ctm_pwd)
session.get_token()
# + id="vI4PTJddLxz1"
t1_flow = CmJobFlow(
application="Naga0.3_Examples", sub_application="Demo-OR_JOB", session=session
)
# + [markdown] id="k7HfewL42f18"
# ## Step 2B - Define the Schedule
# + id="PW4yoM6b-2v2"
t1_flow.set_run_as(username="ctmuser", host="acb-rhctmv20")
# + id="qMVzFXekvSTy"
# Define the schedule
months = ["JAN", "OCT", "DEC"]
monthDays = ["ALL"]
weekDays = ["MON", "TUE", "WED", "THU", "FRI"]
fromTime = "0300"
toTime = "2100"
t1_flow.set_schedule(months, monthDays, weekDays, fromTime, toTime)
# + [markdown] id="9PBDYvbtwWfL"
# # Step 3 - Create Folder
# + id="7MQ5so2gwWK1"
# Create Fodler
f1 = t1_flow.create_folder(name="OR-JOB")
# + [markdown] id="dLGmvMkuwdMW"
# # Step 4 - Create Tasks
# + id="shpuP7_fwiNP"
start = t1_flow.add_job(f1, DummyJob(f1, "Start-Flow"))
job1 = DummyJob(f1, "Job1")
job1.add_if_output("if-true", "*true*", "Job1-TO-Job2")
job1.add_if_output("if-flase", "*false*", "Job1-TO-Job3")
job1_id = t1_flow.add_job(f1, job1)
job2 = DummyJob(f1, "Job2")
job2_id = t1_flow.add_job(f1, job2)
job3 = DummyJob(f1, "Job3")
job3_id = t1_flow.add_job(f1, job3)
job4 = DummyJob(f1, "Job4")
job4_id = t1_flow.add_job(f1, job4)
end = t1_flow.add_job(f1, DummyJob(f1, "End-Flow"))
# + [markdown] id="8uRR5vGWx-9Q"
# # Step 5 - Chain Tasks
# + colab={"base_uri": "https://localhost:8080/"} id="oiaQR99LPIVm" outputId="11c0cf22-ad56-401e-82d5-14fe7121b2bc"
# start >> hello_world_id >> end
t1_flow.chain_jobs(f1, [start, job1_id])
t1_flow.chain_jobs(f1, [job4_id, end])
# + [markdown] id="QrQo_1Q4yG-7"
# # Step 6 - Display Workflow
# + [markdown] id="Mn5OvXGuydlJ"
# ## Step 6A - Display DAG
# + colab={"base_uri": "https://localhost:8080/"} id="m8vAW424yWXb" outputId="b46ecabb-7ae5-4b52-d7bb-8362d80b3dd5"
# View the t1_flow Details
nodes, edges = t1_flow.get_nodes_and_edges()
nodes, edges
# + id="PZ32Fm6GHLqd"
# display using graphviz
from ctm_python_client.utils.displayDAG import DisplayDAG
# sudo apt-get install graphviz (on unix)
# or
# brew install graphviz (for mac)
DisplayDAG(t1_flow).display_graphviz()
# + [markdown] id="X3gtVE7sykA2"
# ## Step 6B - Display JSON
# + colab={"base_uri": "https://localhost:8080/"} id="5XojggPS34rE" outputId="7e044bbf-d412-4d89-f6b9-f923fb32bd8f"
t1_flow.display_json()
# + [markdown] id="hBqW_iV2yKcW"
# # Step 7 - Submit Workflow to Control-M
# + colab={"base_uri": "https://localhost:8080/"} id="LNFCqGbpt-J1" outputId="146f4413-71f6-48cf-cad6-9a8cd7a2e723"
t1_flow.deploy()
# + colab={"base_uri": "https://localhost:8080/"} id="fP-WxlpJgtPW" outputId="dd7e02b3-6e94-43b2-b0a6-ab115eb857c4"
t1_flow.run()
# + id="3NzmdbQovSVH"
|
examples/python-notebooks/.ipynb_checkpoints/Example-OR-JOBs-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EV Datasets List
# <hr>
# https://ev.caltech.edu/dataset
# data gov
# https://catalog.data.gov/dataset/electric-vehicle-charging-stations
|
code/EV_datasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# ## Style transfer
# +
from fastai.conv_learner import *
from pathlib import Path
from scipy import ndimage
import time
torch.backends.cudnn.benchmark=True
PATH = Path('data/imagenet')
PATH_TRN = PATH/'train'
m_vgg = to_gpu(vgg16(True)).eval()
set_trainable(m_vgg, False)
face = 'input.jpg'
img_fn = PATH_TRN/face
img = open_image(f"{img_fn}")
plt.imshow(img);
sz=512
trn_tfms,val_tfms = tfms_from_model(vgg16, sz)
img_tfm = val_tfms(img)
img_tfm.shape
# +
opt_img = np.random.uniform(0, 1, size=img.shape).astype(np.float32)
opt_img = scipy.ndimage.filters.median_filter(opt_img, [8,8,1])
plt.imshow(opt_img);
# +
#set train image = input image
#opt_img = open_image(f"{img_fn}")
opt_img = val_tfms(opt_img)/2
opt_img_v = V(opt_img[None], requires_grad=True)
opt_img_v.shape
# +
max_iter = 1000
show_iter = 100
optimizer = optim.LBFGS([opt_img_v], lr=0.5)
def actn_loss(x): return F.mse_loss(m_vgg(x), targ_v)*1e3
def step(loss_fn):
global n_iter
optimizer.zero_grad()
loss = loss_fn(opt_img_v)
loss.backward()
n_iter+=1
if n_iter%show_iter==0: print(f'Iteration: {n_iter}, loss: {loss.data[0]}')
return loss
# -
# ## forward hook
# +
class SaveFeatures():
features=None
def __init__(self, m): self.hook = m.register_forward_hook(self.hook_fn)
def hook_fn(self, module, input, output): self.features = output
def close(self): self.hook.remove()
def get_opt():
opt_img = np.random.uniform(0, 1, size=img.shape).astype(np.float32)
opt_img = scipy.ndimage.filters.median_filter(opt_img, [8,8,1])
opt_img_v = V(val_tfms(opt_img/2)[None], requires_grad=True)
return opt_img_v, optim.LBFGS([opt_img_v])
def actn_loss2(x):
m_vgg(x)
out = V(sf.features)
return F.mse_loss(out, targ_v)*1e3
block_ends = [i-1 for i,o in enumerate(children(m_vgg))
if isinstance(o,nn.MaxPool2d)]
block_ends
# -
# ## Style match
# +
style_fn = PATH/'style'/'van-gogh.jpg'
style_img = open_image(f"{style_fn}")
style_img.shape, img.shape
plt.imshow(style_img);
# +
def scale_match(src, targ):
h,w,_ = img.shape
sh,sw,_ = style_img.shape
rat = max(h/sh,w/sw); rat
res = cv2.resize(style_img, (int(sw*rat), int(sh*rat)))
return res[:h,:w]
style = scale_match(img, style_img)
plt.imshow(style)
style.shape, img.shape
# +
opt_img_v, optimizer = get_opt()
sfs = [SaveFeatures(children(m_vgg)[idx]) for idx in block_ends]
m_vgg(VV(img_tfm[None]))
targ_vs = [V(o.features.clone()) for o in sfs]
[o.shape for o in targ_vs]
style_tfm = val_tfms(style_img)
# -
m_vgg(VV(style_tfm[None]))
targ_styles = [V(o.features.clone()) for o in sfs]
[o.shape for o in targ_styles]
# +
def gram(input):
b,c,h,w = input.size()
x = input.view(b*c, -1)
return torch.mm(x, x.t())/input.numel()*1e6
def gram_mse_loss(input, target): return F.mse_loss(gram(input), gram(target))
def style_loss(x):
m_vgg(opt_img_v)
outs = [V(o.features) for o in sfs]
losses = [gram_mse_loss(o, s) for o,s in zip(outs, targ_styles)]
return sum(losses)
# -
# ## Style transfer
opt_img_v, optimizer = get_opt()
def comb_loss(x):
m_vgg(opt_img_v)
outs = [V(o.features) for o in sfs]
losses = [gram_mse_loss(o, s) for o,s in zip(outs, targ_styles)]
cnt_loss = F.mse_loss(outs[0], targ_vs[0])*1e4 + F.mse_loss(outs[2], targ_vs[2])*1e6
style_loss = sum(losses)
return cnt_loss + style_loss
n_iter=0
while n_iter <= max_iter/2: optimizer.step(partial(step,comb_loss))
# +
x = val_tfms.denorm(np.rollaxis(to_np(opt_img_v.data),1,4))[0]
plt.figure(figsize=(9,9))
plt.imshow(x, interpolation='lanczos')
plt.axis('off');
timestr = time.strftime("%Y%m%d-%H%M%S")
out_fn = PATH/'predict'/f"{timestr} {face}"
plt.savefig(out_fn)
# -
for sf in sfs: sf.close()
|
style-transfer-win.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pytorch]
# language: python
# name: conda-env-pytorch-py
# ---
# + [markdown] slideshow={"slide_type": "slide"} toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Descente-de-Gradient" data-toc-modified-id="Descente-de-Gradient-1"><span class="toc-item-num">1 </span>Descente de Gradient</a></span></li></ul></div>
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# Descente de Gradient
# ================
#
# L'[algorithme de la descente de gradient](http://en.wikipedia.org/wiki/Gradient_descent) est un algorithme d'optimisation pour trouver un minimum local d'une fonction scalaire à partir d'un point donné, en effectuant de pas successifs dans la direction de l'inverse du gradient.
#
# Pour une fonction $f: \mathbb{R}^n \to \mathbb{R}$, partant d'un point $\mathbf{x}_0$, la méthode calcule les points successifs dans le domaine de la fonction
#
# $$
# \mathbf{x}_{n + 1} = \mathbf{x}_n - \eta \left( \nabla f \right)_{\mathbf{x}_n} \; ,
# $$
# + [markdown] cell_style="center" slideshow={"slide_type": "subslide"}
# où
#
# $\eta > 0$ est une taille de /pas/ suffisamment petite et and $\left( \nabla f \right)_{\mathbf{x}_n}$ est le [gradient](http://en.wikipedia.org/wiki/Gradient) de $f$ évaluée au point $\mathbf{x}_n$. Les valeurs successives de la fonction
#
# $$
# f(\mathbf{x}_0) \ge f(\mathbf{x}_1) \ge f(\mathbf{x}_2) \ge \dots
# $$
#
# vont décroître globalement et la séquence $\mathbf{x}_n$ converge habituellement vers un minimum local.
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# En pratique utiliser un pas de taille fixe $\eta$ est particulièrement inefficace et la plupart des algorithmes vont plutôt chercher à l'adapter à chaque itération.
#
# Le code suivant implémente la descente de gradient avec un pas de taille fixe s'arrétant quand la [norme](http://en.wikipedia.org/wiki/Norm_(mathematics)#Euclidean_norm) du gradient descend en dessous d'un certain seuil.
# + [markdown] slideshow={"slide_type": "subslide"}
# Attention par défaut, pytorch *accumule* les gradients à chaque passe inverse!
# C'est pourquoi il faut le remettre à zéro à chaque itération.
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# Commençons par importer les suspects usuels
# + cell_style="split" slideshow={"slide_type": "fragment"}
import torch
import numpy as np
import math
# + [markdown] slideshow={"slide_type": "subslide"}
# Illustrons l'accumulation du gradient
# + slideshow={"slide_type": "fragment"}
x1 = torch.empty(2, requires_grad=True)
x1
# + slideshow={"slide_type": "fragment"}
f1 = torch.pow(x1[0],2)
f1
# + code_folding=[] slideshow={"slide_type": "fragment"}
# x1.grad.zero_()
f1.backward(retain_graph=True)
x1.grad
# -
x1.data.sub_(torch.ones(2))
# + [markdown] slideshow={"slide_type": "slide"}
# Maintenant essayons d'implémenter une descente de gradient pour la fonction
# $f(X) = sin(x_1) + cos(x_2)$
# + slideshow={"slide_type": "fragment"}
x0 = torch.ones(2,requires_grad=True)
# + slideshow={"slide_type": "fragment"}
f = torch.sin(x0[0]) + torch.cos(x0[1])
f
# + [markdown] slideshow={"slide_type": "slide"}
# On va avoir besoin de :
# ```python
# f.backward(...) # Pour le calcul du gradient proprement dit
# x.grad.data.zero_() # pour la remise à zéro du gradient après une itération
# np.linalg.norm(x.grad.numpy()) # pour contrôler la convergence (norme l2)
# ```
#
# On veut une fonction gd qui prend en argument $f, x, \eta, \epsilon$
# + slideshow={"slide_type": "fragment"}
def gd(f, x, eta, epsilon):
while 1:
f.backward(retain_graph=True)
# print(np.linalg.norm(x.grad.numpy()))
if (torch.norm(x.grad) < epsilon):
break
else:
x.data.sub_(eta * x.grad.data)
x.grad.data.zero_()
# + slideshow={"slide_type": "slide"}
gd(f, x0, 0.9, 0.00001)
# + slideshow={"slide_type": "fragment"}
print(x0.data)
print(f.data)
# + [markdown] slideshow={"slide_type": "slide"}
# Cette fonction ne permet pas d'avoir la valeur de $f$ directement sur le résultat. Il vaut mieux utiliser une fonction qu'un noeud de notre graphe comme argument de notre descente de gradient.
# + slideshow={"slide_type": "fragment"}
x0 = torch.ones(2,requires_grad=True)
x0
# + slideshow={"slide_type": "fragment"}
def f(x):
return x[0].sin() + x[1].cos()
# + slideshow={"slide_type": "fragment"}
def gd(f, x, eta, epsilon):
fval = f(x)
while 1:
fval.backward(retain_graph=True) # On a pas besoin de recalculer f(x) dans ce cas
# seul le gradient nous intéresse ici.
# notez qu'en pratique ce n'est pratiquement
# jamais le cas.
if (torch.norm(x.grad) < epsilon):
break
else:
x.data.sub_(eta * x.grad.data)
x.grad.data.zero_()
# + slideshow={"slide_type": "slide"}
gd(f, x0, 0.9, 0.00001)
# + slideshow={"slide_type": "fragment"}
print(x0)
print(f(x0))
|
1_Descente de gradient.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Timeseries
# Pandas started out in the financial world, so it naturally has strong support for timeseries data.
# We'll look at some pandas data types and methods for manipulating timeseries data.
# Afterwords, we'll use [statsmodels' state space framework](http://www.statsmodels.org/stable/statespace.html) to model timeseries data.
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('default')
plt.rcParams['figure.figsize'] = (12, 6)
pd.options.display.max_rows = 10
# ## Datatypes
#
# - `pd.Timestamp` (nanosecond resolution `datetime.datetime`)
# - `pd.Timedelta` (nanosecond resolution `datetime.timedelta`)
# Pandas provides highly-performant (mostly) drop-in replacements for `datetime.datetime` (`pd.Timestamp`) and `datetime.tiemedelta` (`pd.Timedelta`).
# These have been tailored for efficient storage in NumPy arrays.
# For the most part you'll be working with `DatetimeIndex`es or `TimedeltaIndex`es, or Series / DataFrames containing these.
#
# The biggest limitation is that pandas stores `Timestamp`s at nanosecond resolution. Since they're backed by NumPy's 64-bit integer, the minimum and maximum values are
pd.Timestamp.min, pd.Timestamp.max
# If this is a problem, [there are workarounds](http://pandas.pydata.org/pandas-docs/stable/timeseries.html#representing-out-of-bounds-spans).
# We'll go back to the BTS data set on flights.
# This time I've provided the number of flights per hour for two airports in Chicago: Midway (MDW) and O'Hare (ORD). The data go back to January 1st, 2000.
df = pd.read_csv("data/flights-ts.csv.gz", index_col=0, parse_dates=True)
df.head()
# ## Resampling
#
# Resampling is similar to a groupby, but specialized for datetimes.
# Instead of specifying a column of values to group by, you specify a `rule`: the desired output frequency.
# The original data is binned into each group created by your rule.
resampler = df.resample("MS") # MS=Month Start
resampler
# There's an extensive list of frequency codes: http://pandas.pydata.org/pandas-docs/stable/timeseries.html#offset-aliases.
#
# If you examine the raw data in `df`, you'll notice that it's not at a fixed frequency.
# Hours where there weren't any flights just simply aren't present.
# This isn't a problem though; resample is perfect for going from "ragged" timeseries data to fixed-frequency data.
#
# Just like with `.groupby`, `.resample` returns a deferred object that hasn't really done any work yet.
# It has methods for aggregation, transformation, and general function application.
resampler.sum()
resampler.sum().plot();
# <div class="alert alert-success" data-title="Resample">
# <h1><i class="fa fa-tasks" aria-hidden="true"></i> Exercise: Resample</h1>
# </div>
# <p>Plot the standard deviation for the number of flights from `MDW` and `ORD` at a weekly frequency</p>
# Your solution
# %load solutions/timeseries_resample.py
# <div class="alert alert-success" data-title="Resample-Agg">
# <h1><i class="fa fa-tasks" aria-hidden="true"></i> Exercise: Resample-Agg</h1>
# </div>
# <p>Compute the the total number of flights (sum), mean, and median flights *per Quarter*.</p>
# %load solutions/timeseries_resample_agg.py
# ## Rolling, Expanding
#
# Applying functions to windows, moving through your data.
# These are very similar to groupby and resample. Let's get the daily number of flights with a `resample` quick.
daily = df.resample('D').sum()
daily
# Suppose you wanted a 30-day moving (or rolling) average.
# This is possible with the `.rolling` method. Like `groupby` and `resample`, this object is just going to store the information to know what subset of data to operate on next; it doesn't actually do any work yet:
daily.rolling(30, center=True)
# The first argument is the window size.
# Since `daily` is at daily frequency, 30 means a 30-day window.
# `center=True` says to label each window with the middle-most point.
# To actually do work, you call a method like `.mean`;
fig, ax = plt.subplots()
daily.rolling(30).mean().rename(columns=lambda x: x + " (30D MA)").plot(ax=ax, alpha=.25,
color=['C0', 'C1'])
daily.plot(ax=ax, alpha=.25, color=['C0', 'C1'], legend=False);
# It's common to combine resampling and rolling.
df.resample("D").sum().rolling(30).corr(pairwise=True).xs("MDW", level=1)['ORD'].plot(
title="O'Hare : Midway cross-correlation (30D MA)", figsize=(12, 4)
);
# ## Timezones
#
# pandas can store an array of datetimes with a common timezone.
# Right now the index for `df` is timezone naïve, but we can convert to a timezone with `tz_convert`:
df.index.tzinfo # None, timezone naïve
df.index.tz_localize("US/Central")
# Timezones, as usual, are annoying to deal with.
# We've hit a daylight savings time issue.
# As the error says, 2000-04-02T02:00:00 isn't actaully a valid time in US/Central.
# I checked the BTS website, and these timestamps are supposed to be local time, so presumably some data was recorded incorrectly.
# pandas is strict by default, so it we need to tell it to ignore those errors:
idx = df.index.tz_localize("US/Central", ambiguous="NaT", errors='coerce')
idx
pd.isnull(idx).sum() # 25 bad values
# Notice the dtype: `datetime64[ns, US/Central]`.
# That means nanosecond resolution in the US/Central time zone.
# Once you have a datetime with timezone, you can convert timezones with `tz_convert`:
idx.tz_convert("UTC")
# ## Offsets
#
# I wish the standard library `datetime` module had something like this.
# Let's generate some fake data with `pd.date_range`
dates = pd.date_range("2016-01-01", end="2016-12-31", freq='D')
dates
# There are a whole bunch of offsets available in the `pd.tseries.offsets` namespace. For example, to move 3 business days into the future:
dates + pd.tseries.offsets.BDay(3)
# Or to move to the next month end:
dates + pd.tseries.offsets.MonthEnd()
# ## Timedelta Math
#
# Being able to add columns of dates and timedeltas turns out to be quite convenient.
# Let's go all the way back to our first example with flight delays from New York airports.
flights = pd.read_csv("data/ny-flights.csv.gz", parse_dates=['dep', 'arr'])
flights.head()
# <div class="alert alert-success" data-title="Convert Timedelta">
# <h1><i class="fa fa-tasks" aria-hidden="true"></i> Exercise: Convert Timedelta</h1>
# </div>
# <p>Convert `flights.dep_delay` and `flights.arr_delay` to timedelta dtype.</p>
#
# - Hint: recall our type conversion methods: `pd.to_*`
# - Make new columns in `flights` called `dep_delay_td` and `arr_delay_td`
# - Check the `unit` argument for the conversion method. The delay columns are in *minutes*.
# Your solution
# %load solutions/timeseries_timedelta.py
# <div class="alert alert-success" data-title="Timedelta Math">
# <h1><i class="fa fa-tasks" aria-hidden="true"></i> Exercise: Timedelta Math</h1>
# </div>
# <p>Compute the actual time the flight left, but adding the departure time `dep` and the delay `dep_delay`.
# %load solutions/timeseries_departure.py
# # Modeling Timeseries
#
# Timeseries are an interesting problem to model.
# If we're lucky, we have a long history of past data that we can (maybe) use to predict the future.
# We can exploit regularity in the timeseries (seasonal patterns, periods of high values are typically followed by another high value, etc.) to better predict the future.
#
# Statsmodels has a nice framework for fitting timeseries models and evaluating their output.
import statsmodels.formula.api as smf
import statsmodels.tsa.api as smt
import statsmodels.api as sm
# Let's model Monthly flights from `ORD`.
y = daily.ORD.resample("MS").sum()
y.plot();
# That final value is odd because it's not a complete month. Let's drop it.
y = daily.ORD.resample("MS").sum().iloc[:-1]
y.head()
# It's common to estimate the parameters on *differenced* values.
# That is, make a new series $y'$ where $y_t' = y_t - y_{t-1}$. Pandas makes this simple with the `.diff` method.
y_prime = y.diff()
y_prime.head()
# We'll drop that first NaN:
y_prime = y.diff().dropna()
y_prime.plot();
# Think back to regular linear regression: Predict some variable $y$ with some matrix $X$:
#
# $y = \beta_0 + \beta_1 X_1 + \beta_2 X_2 ... + \beta_p X_p + \varepsilon$
#
# When modelling timeseries, past values of $y$ make for good components of $X$.
# We can do this with the pandas `.shift` method:
y_prime.shift()
# So the value for `2001-01-01` (-867) is now labeled `2000-02-01`. We can collect many of these with a list comprehension and a `concat`.
lagged = pd.concat([y_prime.shift(i) for i in range(9)], axis=1,
keys=['y', 'L1', 'L2', 'L3', 'L4', 'L5', 'L6', 'L7', 'L8'])
lagged
# +
mod_lagged = smf.ols('y ~ L1 + L2 + L3 + L4 + L5 + L6 + L7 + L8', lagged)
res_lagged = mod_lagged.fit()
res_lagged.summary()
# -
ax = res_lagged.fittedvalues.plot(label="predicted", figsize=(12, 4), legend=True)
y_prime.plot(label="actual", legend=True);
# In practice, you won't be doing the `shift`ing and `diff`ing yourself.
# It's more convenient to let statsmodels do that for us.
# Then we don't have to worry about un-differencing the fitted / predicted results to interpret them correctly.
# Also, the solvers we'll see next are a bit more sophisticated than a linear regression.
# ## AutoRegressive Model
#
# Predict $y_{t+1}$, given $y_0, y_1, \ldots y_t$
# Let's fit an autoregressive (AR) model. Autoregressive part just means using past values of $y$ to predict the future (like we did above).
# We'll use statsmodel's `SARIMAX` model. The AR part of SARIMAX is for autoregressive.
# It also handles seasonality (**S**), differencing (**I** for integrated), moving average (**MA**), and exogenous regressors (**X**).
#
# We'll stick to a simple AR(8) model (use the last 8 periods) with a single period of differencing.
mod = smt.SARIMAX(y, order=(8, 1, 0)) # AR(8), first difference, no MA
res = mod.fit()
# As usual with statsmodels, we get a nice summary with the fitted coefficeints and some test statistics (which we'll ignore)
res.summary()
# The results instance has all the usual attributes and methods, like `fittedvalues`.
ax = res.fittedvalues.iloc[1:].plot(label="Fitted", legend=True, figsize=(12, 4))
y.plot(ax=ax, label="Actual", legend=True);
# ## Forecasting
#
# The real value of timeseries analysis is to predict the future.
# We can use the `.get_prediction` method to get the predicted values, along with a confidence interval.
# First, we'll look at one-period-ahead forecasts.
# Basically, this simulates looking at our data the last day of the month, and making the forecast for the next month.
# Keep in mind though that we fit our parameters on the entire dataset. The isn't an out-of-sample prediction.
pred = res.get_prediction(start='2001-03-01')
pred_ci = pred.conf_int()
ax = y.plot(label='observed')
pred.predicted_mean.plot(ax=ax, label='Forecast', alpha=.7)
ax.fill_between(pred_ci.index,
pred_ci.iloc[:, 0],
pred_ci.iloc[:, 1], color='k', alpha=.2)
plt.legend()
sns.despine()
# Alternatively, we can make dynamic forecasts as of some month (January 2013 in the example below). That means the forecast from that point forward only use information available as of January 2013 (though again, we fit the model on the entire dataset). The predictions are generated in a similar way: a bunch of one-step forecasts. Only instead of plugging in the actual values beyond January 2013, we plug in the forecast values.
pred_dy = res.get_prediction(start='2002-03-01', dynamic='2013-01-01')
pred_dy_ci = pred_dy.conf_int()
# +
ax = y.plot(label='observed')
pred_dy.predicted_mean.plot(ax=ax, label='Forecast')
ax.fill_between(pred_dy_ci.index,
pred_dy_ci.iloc[:, 0],
pred_dy_ci.iloc[:, 1], color='k', alpha=.25)
ylim = ax.get_ylim()
ax.fill_betweenx(ylim, pd.Timestamp('2013-01-01'), y.index[-1],
alpha=.1, zorder=-1)
ax.set_ylim(ylim)
ax.annotate('Dynamic $\\longrightarrow$',
(pd.Timestamp('2013-02-01'), 16000))
plt.legend()
sns.despine()
plt.tight_layout()
# -
# There are *a lot* of issues we didn't cover here.
# Seasonality, non-stationarity, autocorrellation, unit roots, and more.
# Timeseries modeling is fraught with traps that will throw off your predictions.
# Still, this should give you a taste of what's possbile.
# ## Further Resources
#
# - [statsmodels state space documentation](http://www.statsmodels.org/dev/statespace.html)
# - [statsmodels state space examples](http://www.statsmodels.org/dev/examples/index.html#statespace)
# - [pyflux](http://www.pyflux.com), another time series modeling library
# - <NAME>'s [post on ARIMA](http://www.seanabu.com/2016/03/22/time-series-seasonal-ARIMA-model-in-python/)
# - <NAME>'s [talks at PyData](https://www.youtube.com/watch?v=tJ-O3hk1vRw)
# - My [blog post](http://tomaugspurger.github.io/modern-7-timeseries.html)
|
notebooks/05-Timeseries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Under- and overfitting, model selection
#
# ## Preliminaries
#
# In the first set of exercises you had to implement the training and evaluation of the linear regression and $k$-NN methods from scratch in order to practice your `numpy` skills. From this set of exercises onward, you can use the implementations provided in `scikit-learn` or other higher-level libraries. We start this set of exercises by demonstrating some of the features of `scikit-learn`.
#
# For example, implementation of linear regression model fitting with an analytical solution for the parameters is provided by the class `sklearn.linar_model.LinearRegression`. You can train a linear regression model in the following way:
# +
import numpy as np
from sklearn import datasets, linear_model
# load the diabetes dataset
diabetes = datasets.load_diabetes()
# use only one feature
X = diabetes.data[:, np.newaxis, 2]
y = diabetes.target
# split the data into training/testing sets
X_train = X[:-20]
X_test = X[-20:]
# split the targets into training/testing sets
y_train = y[:-20]
y_test = y[-20:]
# create linear regression object
model = linear_model.LinearRegression()
# train the model using the training dataset
model.fit(X_train, y_train)
# -
# Let's visualize the training dataset and the learned regression model.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
fig = plt.figure()
plt.plot(X_train, y_train, 'r.', markersize=12)
X_edge = np.array([np.min(X_train, 0), np.max(X_train, 0)])
plt.plot(X_edge, model.predict(X_edge), 'b-')
plt.legend(('Data', 'Linear regression'), loc='lower right')
plt.title('Linear regression')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.show()
# -
# Once trained, the model can be used to make predictions on the test data:
# Make predictions using the testing dataset
prediction = model.predict(X_test)
# The next step (not shown here) is to evaluate the performance of the trained model.
#
# Note that the `scikit-learn` interface works by first initializing an object from the class that implements the machine learning model (linear regression in this case) and then fitting the initialized model using the data in the training set. Finally, the trained (fitted) model can be used to make predictions on unseen data. In fact, all models implemented in this library follow the same *initialize-fit-predict* programming interface. For example, a $k$-NN classifier can be trained in the following way:
# +
from sklearn.model_selection import train_test_split
from sklearn import datasets, neighbors
breast_cancer = datasets.load_breast_cancer()
X = breast_cancer.data
y = breast_cancer.target
# make use of the train_test_split() utility function instead
# of manually dividing the data
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state=40)
# initialize a 3-NN classifier
model = neighbors.KNeighborsClassifier(n_neighbors=3)
# train the model using the training dataset
model.fit(X_train, y_train)
# make predictions using the testing dataset
prediction = model.predict(X_test)
# -
# Note that the features in the breast cancer dataset have different scales (some have on average very small absolute values, and some very large), which means that the distance metric used by $k$-NN will me dominated by the features with large values. You can use any of the number of feature transformation methods implemented in `scikit-learn` to scale the features. For example, you can use the `sklearn.preprocessing.StandardScaler` method to transform all features to a have a zero mean and unit variance:
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
# -
# The scaler has its own parameters which are the means and standard deviations of the features estimated from the training set. If you train a model with the scaled features, you will have to remember to also apply the scaling transformation every time you make a prediction on new unseen and unscaled data. This is somewhat prone to error. One option for making the code more robust is to create a processing pipeline that includes the scaling and $k$-NN models in a sequence:
# +
from sklearn.pipeline import Pipeline
knn = neighbors.KNeighborsClassifier(n_neighbors=3)
model = Pipeline([
("scaler", scaler),
("knn", knn)
])
# train the model using the training dataset
model.fit(X_train, y_train)
# make predictions using the testing dataset
prediction = model.predict(X_test)
# -
# If you are curious, more information about the design of the `scikit-learn` application programming interface (API) can be found [in this paper](https://arxiv.org/pdf/1309.0238.pdf).
# ## Exercises
#
# ### Bias-variance decomposition
#
# Show that the mean squared error of the estimate of a parameter can be decomposed into an expression that includes both the bias and variance (Eq. 5.53-5.54 in "Deep learning" by Goodfellow et al.).
# ***ANSWER***:
#
#
# $
# \begin{aligned}
# \operatorname{MSE}(\hat{\theta}) &=\mathrm{E}_{\theta}\left[(\hat{\theta}-\theta)^{2}\right] \\
# &=\mathrm{E}_{\theta}\left[\left(\hat{\theta}-\mathrm{E}_{\theta}[\hat{\theta}]+\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2}\right] \\
# &=\mathrm{E}_{\theta}\left[\left(\hat{\theta}-\mathrm{E}_{\theta}[\hat{\theta}]\right)^{2}+2\left(\hat{\theta}-\mathrm{E}_{\theta}[\hat{\theta}]\right)\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)+\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2}\right] \\
# &=\mathrm{E}_{\theta}\left[\left(\hat{\theta}-\mathrm{E}_{\theta}[\hat{\theta}]\right)^{2}\right]+\mathrm{E}_{\theta}\left[2\left(\hat{\theta}-\mathrm{E}_{\theta}[\hat{\theta}]\right)\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)\right]+\mathrm{E}_{\theta}\left[\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2}\right]
# \end{aligned}
# $
#
# With $\operatorname{Var}\left(\hat{\theta}\right) = \mathrm{E}\left[\left(\hat{\theta}-\mathrm{E}[\hat{\theta}]\right)^{2}\right]$ :
#
# $
# \begin{aligned}
# \operatorname{MSE}(\hat{\theta}) &= \operatorname{Var_\theta}\left(\hat{\theta}\right) + \mathrm{E}_{\theta}\left[2\left(\hat{\theta}-\mathrm{E}_{\theta}[\hat{\theta}]\right)\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)\right]+\mathrm{E}_{\theta}\left[\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2}\right] \\
# &= \operatorname{Var_\theta}\left(\hat{\theta}\right) + 2\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right) \mathrm{E}_{\theta}\left[\hat{\theta}-\mathrm{E}_{\theta}[\hat{\theta}]\right]+\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2} \\
# &= \operatorname{Var_\theta}\left(\hat{\theta}\right) + 2\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right) \left(\mathrm{E}_{\theta}[\hat{\theta}]-\mathrm{E}_{\theta}[\hat{\theta}]\right)+\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2} \\
# &= \operatorname{Var_\theta}\left(\hat{\theta}\right) + 2\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right) \left(0\right)+\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2} \\
# &= \operatorname{Var_\theta}\left(\hat{\theta}\right) +\left(\mathrm{E}_{\theta}[\hat{\theta}]-\theta\right)^{2} \\
# \end{aligned}
# $
#
# And since $\operatorname{Bias}(\hat{\theta}_m) = \mathrm{E}(\hat{\theta}_m) - \theta$:
#
#
# $
# \operatorname{MSE}(\hat{\theta}) = \operatorname{Var_\theta}\left(\hat{\theta}\right) +\operatorname{Bias}_{\theta}\left(\hat{\theta}\right)^{2}
# $
#
# ### Polynomial regression
#
# For this exercise we will be using generated data to better show the effects of the different polynomial orders.
# The data is created using the make_polynomial_regression function.
# +
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 1
def generate_dataset(n=100, degree=1, noise=1, factors=None):
# Generates a dataset by adding random noise to a randomly
# generated polynomial function.
x = np.random.uniform(low=-1, high=1, size=n)
factors = np.random.uniform(0, 10, degree+1)
y = np.zeros(x.shape)
for idx in range(degree+1):
y += factors[idx] * (x ** idx)
# add noise
y += np.random.normal(-noise, noise, n)
return x, y
# load generated data
np.random.seed(0)
X, y = generate_dataset(n=100, degree=4, noise=1.5)
plt.plot(X, y, 'r.', markersize=12);
# -
# Implement polynomial regression using the `sklearn.preprocessing.PolynomialFeatures` transformation. Using the `sklearn.grid_search.GridSearchCV` class, perform a grid search of the polynomial order hyperparameter space with cross-validation and report the performance on an independent test set.
#
# Plot a learning curve that show the validation accuracy as a function of the polynomial order.
#
# <p><font color='#770a0a'>Which models have a high bias, and which models have high variance? Motivate your answer.</font><p>
#
# ***ANSWER***: Low order polynomial models have high bias and low variance, while higher order polynomial models have low bias and high variance. This can be explained by thinking about how accurately a model can describe the dataset vs what happens to a model upon slight changes in the dataset. Low order models can not accurate describe all data points and therefore have incomplete/simple parameter estimates and thus high bias, however because much data is used to describe few parameters, these parameters do not significantly change due to slight data sampling changes, giving low estimate variances. This is inverse for higher order models.
#
# <p><font color='#770a0a'>Repeat this experiment, this time using the diabetes dataset instead of the generated data.</font></p>
#
# ### Pseudo-dataset:
# +
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
from sklearn.metrics import mean_squared_error, make_scorer
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
n = 1000
X, y = generate_dataset(n=n, degree=4, noise=1.5)
X_train = X[n//5:].reshape(-1,1)
X_test = X[:n//5].reshape(-1,1)
y_train = y[n//5:]
y_test = y[:n//5]
# Define the functions for polynomial features and linear regression
transform = PolynomialFeatures()
linreg = LinearRegression()
scaler = StandardScaler()
# Define the pipeline, containing the polynomial transformation and the subsequent linear regression
model = Pipeline([
("scaler", scaler),
("transf", transform),
("linreg", linreg)
])
# Define the hyperparameters for the Grid Search
params = {'transf__degree': (2,3,4,5,6)}
# Define a custom scoring system to evaluate the mean squared error (lower is better)
mse = make_scorer(mean_squared_error,greater_is_better=False)
# Define the Grid Search method
gridsearch = GridSearchCV(model, params, scoring=mse, cv=5)
# Run Grid Search
gridsearch.fit(X_train, y_train)
y_pred = gridsearch.predict(X_test)
MSE_test_set = mean_squared_error(y_test, y_pred)
print(f'Mean squared error (test set): {MSE_test_set:.4}')
# plot
fig = plt.figure();
plt.plot(X_train, y_train, 'r.', markersize=12, label='train_data');
plt.plot(X_test, y_test, 'g.', markersize=12, label='test_data');
plt.plot(X_test, y_pred, 'b.', markersize=12, label='predicted_test_data');
plt.title('Data')
plt.legend()
# plot 2
plt.figure()
plt.title('Scores for every degree (per fold different colors)')
colors = ['b.', 'r.', 'g.', 'y.', 'k.', 'c.']
for i in range(5):
mse_test_score = gridsearch.cv_results_[f'split{i}_test_score']
plt.plot(params['transf__degree'], mse_test_score, colors[i], markersize=12)
plt.xlabel('Degree')
plt.ylabel('MSE')
plt.legend(list(range(1,len(params['transf__degree'])+1)))
# plot 3
plt.figure()
plt.plot(params['transf__degree'],gridsearch.cv_results_[f'mean_test_score'], 'k.', markersize=12)
plt.title('Average scores of all folds')
plt.xlabel('Degree')
plt.ylabel('MSE');
# -
# ### Diabetes dataset (all 10 features):
# +
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
from sklearn.metrics import mean_squared_error, make_scorer
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.datasets import load_diabetes
def do_gridsearch_polyregs(X_train, y_train, X_test, y_test, degrees, crossvals, plot=True):
"""
Given a train and a test dataset, fit and evaluate polynomial regression models with given degrees using Grid Search
Parameters:
X_train: 2D (Datapoints x features) numpy array
the datapoints on which to train the models
y_train: 1D (Targets) numpy array
the targets on which to train the models
X_test: 2D (Datapoints x features) numpy array
the datapoints on which to test the models
y_test: 1D (Targets) numpy array
the targets on which to test the models
degrees: tuple or list of ints
the degrees of the polynomial regression models to fit
plot: bool
True for plotting the MSE scores, False for surpressing the plots
Returns:
a GridSearchCV object, containing the fitted models for every degree polynomial
"""
# Define the functions for polynomial features and linear regression
transform = PolynomialFeatures()
linreg = LinearRegression()
scaler = StandardScaler()
# Define the pipeline, containing the polynomial transformation and the subsequent linear regression
model = Pipeline([
("scaler", scaler),
("transf", transform),
("linreg", linreg)
])
# Define the hyperparameters for the Grid Search
params = {'transf__degree': degrees}
# Define a custom scoring system to evaluate the mean squared error (lower is better)
mse = make_scorer(mean_squared_error,greater_is_better=False)
# Define the Grid Search method
gridsearch = GridSearchCV(model, params, scoring=mse, cv=crossvals)
# Run Grid Search
gridsearch.fit(X_train, y_train)
y_pred = gridsearch.predict(X_test)
MSE_test_set = mean_squared_error(y_test, y_pred)
print(f'Mean squared error (test set): {MSE_test_set:.4}')
if plot:
# Plot test scores per fold
plt.figure()
plt.title('Scores for every degree (per fold different colors)')
colors = ['b.', 'r.', 'g.', 'y.', 'k.', 'c.']
for i in range(crossvals):
mse_test_score = gridsearch.cv_results_[f'split{i}_test_score']
plt.plot(params['transf__degree'], np.log(-mse_test_score), colors[i], markersize=12)
plt.xlabel('Degree')
plt.ylabel('log(-MSE)')
plt.legend(list(range(1,crossvals+1)))
# Plot average test score of all folds
plt.figure()
plt.plot(params['transf__degree'], np.log(-gridsearch.cv_results_[f'mean_test_score']), 'k.', markersize=12)
plt.title('Average scores of all folds')
plt.xlabel('Degree')
plt.ylabel('log(-MSE)');
return gridsearch
# Load the diabetes dataset (use all 10 features)
diabetes = load_diabetes()
X = diabetes.data
y = diabetes.target
# Split the data into training and test set
X_train, X_test, y_train, y_test = train_test_split(X, y)
# Define the polynomial degrees that are evaluated in the Grid Search
degrees_for_gridCV = (1,2,3,4,5,6)
# Run the Grid Search (5 cross-validations, plot the results)
grids = do_gridsearch_polyregs(X_train, y_train, X_test, y_test, degrees_for_gridCV, 5)
# -
# ### ROC curve analysis
# A common method to evaluate binary classifiers is the receiver operating characteristic (ROC) curve. Similar to the week one practicals, implement a $k$-NN classifier on the breast cancer dataset, however, his time use the $k$-NN pipeline from the preliminary. Train the model for different values of $k$ and evaluate their respective performance with an ROC curve, use the `sklearn.metrics.roc_curve` function.
# +
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import roc_curve, roc_auc_score, make_scorer, plot_roc_curve
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.datasets import load_breast_cancer
import matplotlib.pyplot as plt
# Load breast cancer dataset
breast_cancer = load_breast_cancer()
X = breast_cancer.data
y = breast_cancer.target
# Split data into training and test data
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y)
# Define the hyperparameters to be researched in the Grid Search
params = {'n_neighbors': (3,5,7,9,11,13), 'weights': ('uniform', 'distance')}
kNN = KNeighborsClassifier()
roc_auc_scorer = make_scorer(roc_auc_score, greater_is_better=True, needs_threshold=True)
gridsearch = GridSearchCV(kNN, params, scoring=roc_auc_scorer)
# Run Grid Search to find optimal settings
gridsearch.fit(X_train, y_train)
# Predict targets for test set using the optimal settings (found in the Grid Search)
y_pred = gridsearch.predict(X_test)
# Calculate true positive rate (TPR) and false positive rate (FPR) to plot ROC-curve.
plot_roc_curve(gridsearch, X_test, y_test);
# -
# ### $F_1$ score and Dice similarity coefficient
#
# The Dice similarity coefficient is a very popular evaluation measure for image segmentation applications. Assuming that $A$ is the ground truth segmentation of an object represented as a binary image, and $B$ is the binary output of an image segmentation method, the Dice similarity coefficient is computed as:
#
# $\text{Dice}(A,B) = \frac{2|A\cap B|}{|A| + |B|}$
#
# where $|\cdot|$ represents the cardinality of the objects (e.g. $|A|$ is the number of non-zero pixels in the ground truth segmentation).
#
# For example, the Dice similarity can be computed in the following way:
# +
# generate some test objecys
A = np.zeros((32, 32))
A[10:-10, 10:-10] = 1
B = np.zeros((32, 32))
B[5:-15, 5:-15] = 1
dice = 2*np.sum(A*B)/(np.sum(A)+np.sum(B))
# display the results
plt.plot()
plt.imshow(A)
plt.imshow(B, alpha=0.7)
print(dice)
# -
# <p><font color='#770a0a'>Show that the $F_1$ score, which is the harmonic mean of precision and recall, is equivalent to the Dice similarity coefficient</font><p>
#
# ***ANSWER:***
#
# $\text{Dice}(A,B) = \frac{2|A\cap B|}{|A| + |B|}$
#
# $F_1(A,B) = 2*\frac{\text{precision} * \text{recall}}{\text{precision} + \text{recall}}$
#
# $\text{precision} = \frac{TP}{TP + FP}$
#
# $\text{recall} = \frac{TP}{TP + FN}$
#
# $
# \begin{aligned}
# F_1(A,B) &= 2*\frac{\frac{TP}{TP + FP} * \frac{TP}{TP + FN}}{\frac{TP}{TP + FP} + \frac{TP}{TP + FN}} \\
# &= 2*\frac{\frac{TP^2}{(TP + FP)(TP+FN)}}{\frac{TP(TP+FN) + TP(TP+FP)}{(TP+FP)(TP+FN)}} \\
# &=2*\frac{TP^2}{(TP + FP)(TP+FN)} * \frac{(TP+FP)(TP+FN)}{TP(TP+FN) + TP(TP+FP)} \\
# &=2* \frac{TP^2}{(TP(TP+FN) + TP(TP+FP))} \\
# &=\frac{2*TP}{2 * TP+FN+FP} \\
# \end{aligned}
# $
#
# $|A\cap B|$ can also be described as a true positive prediction (area of ground truth that overlaps with area of prediction is a true positive). $|A| + |B|$ are both the areas of the ground truth and the prediction summed, in other words, $TP + FN$ for the true sample, and $TP+FP$ for the prediction. Plugging this into one of the equations (F1 or Dice) results in the other equation.
|
practicals/week_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manometry
#
# ## Learning outcomes
#
# * Apply knowledge of hydrostatic pressure to pressure measurement
# * Understand the manometer instrument
# * Learn how an inclined manometer increases measurement sensitivity
#
# # Introduction
#
# Manometers are a class of instruments used to measure fluid pressure which make use of the **Hydrostatic Equation**:
#
# \begin{align}
# p_2 - p_1 = -\rho g \left(z_2 - z_1 \right)
# \end{align}
#
# For a fluid of known density in a constant gravity well, a change in elevation $(z_2 - z_1)$ linearly corresponds to a change in pressure.
#
# Consider the section of pipe shown below. A liquid flows along the length of the pipe (we can assume it is much longer than shown and the flow moves entirely parallel to the pipe) and at some point along its length a hole is drilled and a small open tube is inserted normal to the upper surface. This arrangement is called a **piezometer tube**.
#
# <img src="media/2.2/piezometer1.png" alt="Piezometer" width="800" title="Piezometer" />
#
#
# Shown below is a cross sectional view of the geometry. The arrows indicate the flow direction. The small vertical tube is also shown.
#
# **What do think the value of $h$ is?**
#
# Will the flow in the main pipe squirt out of this smaller tube? Will the flow continue along its path and ignore the perpendicular tube completely?
#
# <img src="media/2.2/piezometer2.png" alt="Piezometer" width="800" title="Piezometer" />
#
# The answer is that for some constant velocity in the main pipe is that the flow will rise up the perpendicular tube some finite $h$ and stop once it's reached an equilibrium.
# If we change the velocity in the pipe the level $h$ will change accordingly. Why? In order to increase the velocity in the pipe we have too apply more pressure.
#
# If we are applying a positive gauge pressure at the pipe inlet and then let the pipe exit to the atmosphere (zero gauge pressure) then there will be a *pressure gradient* along the length of the pipe and the liquid will flow from high to low pressure — at a **constant velocity** (it's a little more complicated than that but more on that later). This result is that the hydrostatic pressure is constant perpendicular to the direction of flow (assuming a sufficiently small diameter). In fact if we place a series of these *'piezometer tubes'* along the length of the pipe we could visualise the pressure gradient along the pipe.
#
# <img src="media/2.2/piezometer3.png" alt="Piezometer" width="800" title="Piezometer" />
#
# The pressure at each location $n$ can be measured by measuring the local height $h_n$:
#
# \begin{align}
# p_{abs}(x) = p_{atm} + \rho g h_{n}
# \end{align}
#
# or
#
# \begin{align}
# p_{gauge}(x) = \rho g h_{n}
# \end{align}
#
# It is the static pressure that pushes (*piezo* is greek for push) the fluid laterally into the piezometer tube. The velocity of flow moving along the pipe does not directly play a role in the height $h$. In fact in the pipe was instead a closed pressure vessel containing a static fluid as observed in our Bourdon gauge example we would observe the same behaviour. However the Piezometer tube may need to be extremely long for any practical usage!
#
# ## Pressure head
#
# We can use the height of a column of liquid as a unit of pressure. We see this used in the Mercury Barometer where atmospheric pressure is often recorded in $mmHg$ or $inHg$. Since the height of the liquid column is dependent on the density and acceleration due to gravity (the specific weight, $\gamma = \rho g$):
#
# \begin{align}
# \text{Pressure Head} = \frac{\Delta P}{\gamma}
# \end{align}
#
# When working with a liquid in a complex flow system such as the pipe network of a chemical plant it is often convenient to work in pressure head.
# ## The U-tube Manometer
#
# We've seen various arrangements of tubes used to measure fluids so far; from Boyle's *'J-tube'* to the Piezometer tube. Now we will consider another, the *'U-tube Manometer'*
#
# If we take a U-shaped tube, open to atmosphere at both ends, and partially fill it form one side with a liquid, what happens? The fluid flows down and around and eventually reaches an equilibrium with the free surfaces in each arm perfectly level with one another.
#
# <video controls src="media/2.2/u-tube.mp4" width="400" />
# We can perhaps understand this equilibrium better if we consider the toy example shown in the video below. Owing to friction, the spheres that fall down the right side of the tube eventually come to rest. Each sphere has an identical finite mass and therefore gravity pulls on each with the same force which results in the distribution of the spheres balanced being between either side. In the case of the fluid we can, for now, think of the fluid as consisting of many many tiny spheres.
#
# <video controls src="media/2.2/u-tube_balls.mp4" width="400" />
# Gravity $g$ pulls the fluid (with density $\rho$) down evenly and atmospheric pressure which acts at each open end (which we will denote as 1 and 2) is equal. The Hydrostatic Equation is satisfied so that $p_1 = p_2$ and the height of the fluid relative to a common datum is $h_1 = h_2$.
#
# But what if we connect one end of our U-tube to a pressure vessel as shown on the left side of the image below?
#
# <img src="media/2.2/U-bend_Manometer.png" alt="Piezometer" width="800" title="Piezometer" />
#
# On the right is shown an illustration of the U-tube from the previous video and on the left is it after the left side is connected to the pressure vessel containing a fluid ($\gamma_1 = \rho_1 g$) at some unknown pressure $P_A$.
#
# **Is the pressure at point A greater or less than atmospheric?
# Which is the same as asking is the *gauge pressure* greater or less than zero?**
#
# We can use the hydrostatic equation to reason that the pressure $p_2 = p_3$ since any pressure acting on the interface at point 2 will be balanced at point 3 by the atmosphere and the weight of fluid in the column $h_2$. Working in gauge pressure we offset by atmospheric pressure so that the forces on the right side are equal to $\gamma_2 h_2$. On the left side the pressure $p_A$ is invariant along the dash-dot line so the forces acting on the left side are equal to $p_{A_\text{gauge}} + \gamma_1 h_1$. Since the sum of the forces is zero:
#
# \begin{align}
# p_{A_\text{gauge}} + \gamma_1 h_1 = \gamma_2 h_2
# \end{align}
#
# which is easily rearranged:
# \begin{align}
# p_{A_\text{gauge}} = \gamma_2 h_2 - \gamma_1 h_1
# \end{align}
#
# This tells us that by measuring the difference in height between the two sides for two fluids of differing specific weights we can compute the pressure at point $A$ — which, remember, is at the same vertical location as point 1 and therefore at the same hydrodynamic pressure.
#
# To answer the question, the pressure at point A is higher than atmospheric unless the density of the fluid in the tank is greater than the fluid we are using as our manometer gauge fluid — which would simply not work as the blue gauge fluid would float up through the red fluid.
#
# ### Gauge fluids
#
# Dense liquids like water and mercury are popular gauge fluids. See here for a list of commercial gauge fluids with various specific gravity ranges
#
# http://www.dwyer-inst.com/PDF_files/GageFluids_i.pdf
#
# Remember, ($SG = \frac{\rho}{\rho_{H_2O}}$)
#
# ## An Example
#
# A tank containing air and oil (SG 0.9) is connected to a mercury U-tube manometer (SG 13.8).
#
# When $h_1 = 914~mm$, $h_2 = 152~mm$ and $h_3 = 228~mm$ **determine the pressure of the air inside the tank**.
#
# <img src="media/2.2/U-bend_Manometer_ex.png" alt="Manometer example" width="500" title="Manometer example" />
#
# We can start by listing the forces on each side.
# On the left we have the air pressure and the hydrostaic pressure of the height of oil from the air/oil interface down to point 1.
# We will consider any hydrostatic pressure in the air volume as negligible since air has a very low density.
# Since $p_1 = p_2$ we can ignore the mercury below this level.
# On the right we simply have the weight of $h_3~mm$ of mercury.
#
# This gives:
#
# $p_{air} + \gamma_{oil} (h_2 + h_1) = \gamma_{Hg} h_3 $
#
# Rearranging:
#
# $p_{air} = \gamma_{Hg} h_3 - \gamma_{oil} (h_2 + h_1)$
#
# and expanding for $\gamma = \rho g = SG~\rho_{H_2O}~g$:
#
# $p_{air} = [SG_{Hg}~\rho_{H_2O}~g] h_3 - [SG_{oil}~\rho_{H_2O}~g] (h_2 + h_1)$
# +
# Physical properties
rho_w = 1000 # kg/m^3
SG_HG = 13.6
SG_oil = 0.9
g = 9.81 # m/s/s
h_1 = 0.914 # m
h_2 = 0.152 # m
h_3 = 0.228 # m
p_air = (SG_HG * rho_w * g * h_3) - (SG_oil * rho_w * g * (h_1 + h_2))
# print result using an f-string
print(f"Air presure is {p_air/1000:.2f} kPa")
# -
# ## Differential Manometer
#
# We can also connect both ends of our u-tube manometer to different pressure vessels and measure the differential pressure between them; the atmosphere is just a pressure vessel after all!
#
# Again we can equate the pressures at the bottom $p_2 = p_3$
#
# $p_{A_{gauge}} + \gamma_1 h_1 = p_{B_{gauge}} + \gamma_3 h_3 + \gamma_2 h_2$
#
# $p_{A_{gauge}} - p_{B_{gauge}} = \gamma_3 h_3 + \gamma_2 h_2 - \gamma_1 h_1$
#
# <img src="media/2.2/diff_U-bend_Manometer.png" alt="Differential Manometer" width="600" title="Differential Manometer" />
# ## Inclined-Tube Manometer
#
# The inclined manometer is variant of the U-tube manometer used for increased sensitivity when measuring small pressure differences. This greater sensitivity is due to the fact that the gauge fluid is moved over a greater length of tubing on the inclined side for the same vertical displacement on the vertical side. Once again we can gain some intuition by considering a toy model with some spheres in place of the fluid. Here our manometer tube is vertical on the right and inclined at an angle of $30^\circ$ on the right.
#
# <video controls src="media/2.2/inclined_manometer_balls.mp4" width="800" />
# The image below shows the final frame of the video when the system has reached equilibrium.
# The vertical height of the spheres is equal on each side. Two red laser beams marking the topmost spheres are used to illustrate this.
#
# <img src="media/2.2/inclined_manometer_ball0.png" alt="Inclined manometer" width="800" title="Inclined manometer" />
#
# Now lets consider the case where there is a force sufficient to move the right side down by one sphere diameter. Our laser beams remain fixed and clearly demonstrate that on the left side the spheres are also displaced one diameter vertically. However the topmost sphere on the left is also displaced significantly to the left. Note also that since the top level of the spheres is now one diameter lower the balance of equal force is also one diameter lower.
#
# <img src="media/2.2/inclined_manometer_ball1.png" alt="Inclined manometer" width="800" title="Inclined manometer" />
#
# The result of all of this is apparent from the measurement checker board placed next to each 'surface'. For the current angle, a 4 unit (1 diameter) vertical reduction on the right corresponds to a 12 unit (3 diameters) displacement along the inclined tube a 3:1 ratio. A regular u-tube manometer gives us a 2:1 ratio.
#
# ### Lets look at this a bit more rigorously
#
# In the image below, measurements (in this case differential pressure between $p_A$ and $p_B$) are read along a scale on the inclined tube by reading the level of the gauge fluid ($\gamma_3$)
#
# <img src="media/2.2/inclined_manometer0.png" alt="Inclined manometer" width="800" title="Inclined manometer" />
#
# In the upper image the pressure difference between $p_A$ and $p_B$ results in the gauge fluid moving a distance $l$ along the incline relative to $h_B$.
# The sum of the forces is:
#
# \begin{equation*}
# P_A + \gamma_{1}~h_A + \gamma_{3}~l \sin{\theta} = P_B + \gamma_{2}~(h_B)
# \end{equation*}
#
# If, as shown in the lower part of the image, the pressure $p_A$ is reduced by $\Delta P$ the length of gauge fluid $l$ will increase by distance $\alpha$. Accordingly the height $h_A$ will have to decrease by $\alpha \sin{\theta}$ and the the height $h_B$ will increase by $\Delta h_B = \alpha$. The sum of the forces becomes:
#
# \begin{equation*}
# [P_A - \Delta P] +
# \underbrace{\gamma_{1}~[h_A - \alpha \sin{\theta}]}_\text{fluid 1} +
# \underbrace{\gamma_{3}~[(l+\alpha) \sin{\theta} + \alpha]}_\text{gauge fluid} =
# P_B +
# \underbrace{\gamma_{2}~[h_B + \alpha]}_\text{fluid 2}
# \end{equation*}
#
# It is very important to observe that the measured distance along the inclined portion of the manometer is given by the distance $\alpha + l + \alpha/\sin{\theta}$ as the increase in $h_B$ to $h_B + \alpha$ shifts the dash-dot line of equal pressure in the gauge fluid down. This is clearly apparent in the toy example above where there is a three fold increase in the displacement of the fluid on the inclined side of the manometer.
# ### Example
# Determine the new differential reading along the inclined leg of the mercury manometer if the pressure in pipe A is decreased 10 kPa and the pressure in pipe B remains unchanged.
# The fluid in A has a specific gravity of 0.9 and the fluid in B is water.
#
# <img src="media/2.2/inclined_manometer1.png" alt="Inclined manometer" width="800" title="Inclined manometer" />
#
# We can write the force balance for both sides of the manometer in its initial state:
#
# \begin{equation}
# P_A + \gamma_{A}~(0.1) + \gamma_{Hg}~(0.05) \sin{30^\circ} = P_B + \gamma_{B}~(0.08) .
# \end{equation}
#
# Rewriting the balance with a $10kPa$ pressure reduction in $P_A$:
#
# \begin{align}
# [P_A - 10\times 10^{3}] + \gamma_{A}[({0.1-\alpha \sin{30^\circ}})] +
# \gamma_{Hg}[({\alpha \sin{30^\circ} + l\sin{30^\circ} + \alpha})] =
# P_B + \gamma_{B}[({0.08 + \alpha})]
# \end{align}
#
#
# Subtracting $2$ from $1$:
#
# \begin{equation*}
# -10\times 10^{3} - \gamma_A({~\alpha \sin{30^\circ}}) + \gamma_{Hg}({\alpha\sin{30^\circ} + \alpha}) = \gamma_{B}({\alpha})
# \end{equation*}
#
#
# and solving for $\alpha$
# \begin{equation*}
# \alpha = \frac{-10\times10^{3}}{\gamma_B + \gamma_A ({\sin{30^\circ}}) + \gamma_{Hg}({\sin{30^\circ}+1})} = 0.054~m
# \end{equation*}
#
# The differential pressure is read along the inclined portion of the manometer so we need to calculate the total change in length $L$ of mercury along it.
#
# \begin{equation*}
# L \sin{\theta} = [\alpha + l\sin(\theta) + \alpha\sin(\theta)]
# \end{equation*}
#
# \begin{equation*}
# L = \left[\frac{\alpha}{\sin{\theta}} + l + \alpha\right]
# \end{equation*}
#
# The resulting differential reading is then:
#
# \begin{equation*}
# \frac{\alpha}{\sin{30^\circ}} + \alpha + 0.05 = \frac{0.054}{\sin{30^\circ}}+0.054+0.05=0.215m
# \end{equation*}
#
#
# We can also solve this graphically, accounting for positive or negative changes in $\Delta P$.
#
# <img src="media/2.2/inclined_manometer3.png" alt="Inclined manometer" width="800" title="Inclined manometer" />
|
2.2 Manometry.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 4*
#
# ---
# # Model Interpretation
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [ ] Continue to iterate on your project: data cleaning, exploratory visualization, feature engineering, modeling.
# - [ ] Make at least 1 partial dependence plot to explain your model.
# - [ ] Make at least 1 Shapley force plot to explain an individual prediction.
# - [ ] **Share at least 1 visualization (of any type) on Slack!**
#
# If you aren't ready to make these plots with your own dataset, you can practice these objectives with any dataset you've worked with previously. Example solutions are available for Partial Dependence Plots with the Tanzania Waterpumps dataset, and Shapley force plots with the Titanic dataset. (These datasets are available in the data directory of this repository.)
#
# Please be aware that **multi-class classification** will result in multiple Partial Dependence Plots (one for each class), and multiple sets of Shapley Values (one for each class).
# ## Stretch Goals
#
# #### Partial Dependence Plots
# - [ ] Make multiple PDPs with 1 feature in isolation.
# - [ ] Make multiple PDPs with 2 features in interaction.
# - [ ] Use Plotly to make a 3D PDP.
# - [ ] Make PDPs with categorical feature(s). Use Ordinal Encoder, outside of a pipeline, to encode your data first. If there is a natural ordering, then take the time to encode it that way, instead of random integers. Then use the encoded data with pdpbox. Get readable category names on your plot, instead of integer category codes.
#
# #### Shap Values
# - [ ] Make Shapley force plots to explain at least 4 individual predictions.
# - If your project is Binary Classification, you can do a True Positive, True Negative, False Positive, False Negative.
# - If your project is Regression, you can do a high prediction with low error, a low prediction with low error, a high prediction with high error, and a low prediction with high error.
# - [ ] Use Shapley values to display verbal explanations of individual predictions.
# - [ ] Use the SHAP library for other visualization types.
#
# The [SHAP repo](https://github.com/slundberg/shap) has examples for many visualization types, including:
#
# - Force Plot, individual predictions
# - Force Plot, multiple predictions
# - Dependence Plot
# - Summary Plot
# - Summary Plot, Bar
# - Interaction Values
# - Decision Plots
#
# We just did the first type during the lesson. The [Kaggle microcourse](https://www.kaggle.com/dansbecker/advanced-uses-of-shap-values) shows two more. Experiment and see what you can learn!
# ### Links
#
# #### Partial Dependence Plots
# - [Kaggle / <NAME>: Machine Learning Explainability — Partial Dependence Plots](https://www.kaggle.com/dansbecker/partial-plots)
# - [<NAME>: Interpretable Machine Learning — Partial Dependence Plots](https://christophm.github.io/interpretable-ml-book/pdp.html) + [animated explanation](https://twitter.com/ChristophMolnar/status/1066398522608635904)
# - [pdpbox repo](https://github.com/SauceCat/PDPbox) & [docs](https://pdpbox.readthedocs.io/en/latest/)
# - [Plotly: 3D PDP example](https://plot.ly/scikit-learn/plot-partial-dependence/#partial-dependence-of-house-value-on-median-age-and-average-occupancy)
#
# #### Shapley Values
# - [Kaggle / <NAME>: Machine Learning Explainability — SHAP Values](https://www.kaggle.com/learn/machine-learning-explainability)
# - [<NAME>: Interpretable Machine Learning — Shapley Values](https://christophm.github.io/interpretable-ml-book/shapley.html)
# - [SHAP repo](https://github.com/slundberg/shap) & [docs](https://shap.readthedocs.io/en/latest/)
# +
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# !pip install eli5
# !pip install pdpbox
# !pip install shap
# If you're working locally:
else:
DATA_PATH = '../data/'
# -
|
module4-model-interpretation/LS_DS_234_assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + editable=true
# %load_ext sql
# + editable=true
# %sql postgresql://student:student@127.0.0.1/sparkifydb
# + editable=true
# %sql SELECT * FROM songplays LIMIT 5;
# + editable=true
# %sql SELECT count(*) FROM songplays
# + editable=true
# %sql SELECT * FROM users LIMIT 5;
# + editable=true
# %sql SELECT count(*) FROM users
# + editable=true
# %sql SELECT * FROM songs LIMIT 5;
# + editable=true
# %sql SELECT count(*) FROM songs
# + editable=true
# %sql SELECT * FROM artists LIMIT 5;
# + editable=true
# %sql SELECT count(*) FROM artists
# + editable=true
# %sql SELECT * FROM time LIMIT 5;
# + editable=true
# %sql SELECT count(*) FROM time
# + [markdown] editable=true
# ## REMEMBER: Restart this notebook to close connection to `sparkifydb`
# Each time you run the cells above, remember to restart this notebook to close the connection to your database. Otherwise, you won't be able to run your code in `create_tables.py`, `etl.py`, or `etl.ipynb` files since you can't make multiple connections to the same database (in this case, sparkifydb).
# + [markdown] editable=true
# # Sanity Tests
#
# Execute the cells below once you are ready to submit the project. Some basic sanity testing will be performed to esnure that your work does NOT contain any commonly found issues.
#
# Run each cell and if a cell produces an warning message is orange, you should make appropriate changes to your code before submitting. If all test in a cell pass,no warnings will be printed.
#
# The test cases assume that you are using certain column names in your tables. If you get a `IndexError: single positional indexer is out-of-bounds` you may need to change the column names being used by the test cases. Instructions for doing this appear right boefore cell that may require these changes.
#
# The tests below are only meant to help you make your work foolproof. The submission will still be graded by a human grader against the project rubric.
#
# ---
# + [markdown] editable=true
# ## Grab Table Names for Testing
# + editable=true
import sql_queries as sqlq
# + editable=true magic_args="_tablenames <<" language="sql"
# SELECT tablename
# FROM pg_catalog.pg_tables
# WHERE schemaname != 'pg_catalog' AND schemaname != 'information_schema' AND tableowner = 'student';
# + editable=true
tablenames = _tablenames.DataFrame()
# + editable=true
user_table = [name for name in list(tablenames.tablename) if name in sqlq.user_table_create][0]
song_table = [name for name in list(tablenames.tablename) if name in sqlq.song_table_create][0]
artist_table = [name for name in list(tablenames.tablename) if name in sqlq.artist_table_create][0]
songplay_table = [name for name in list(tablenames.tablename) if name in sqlq.songplay_table_create][0]
# + [markdown] editable=true
# ## Run Primary Key Tests
# + editable=true
# %sql _output << SELECT a.attname, format_type(a.atttypid, a.atttypmod) AS data_type, a.attnotnull, i.indisprimary \
# FROM pg_index i \
# JOIN pg_attribute a ON a.attrelid = i.indrelid \
# AND a.attnum = ANY(i.indkey) \
# WHERE i.indrelid = '{user_table}'::regclass
# + editable=true
if not _output:
print('\033[93m'+'[WARNING] '+ f"The {user_table} table does not have a primary key!")
# + editable=true
# %sql _output << SELECT a.attname, format_type(a.atttypid, a.atttypmod) AS data_type, a.attnotnull, i.indisprimary \
# FROM pg_index i \
# JOIN pg_attribute a ON a.attrelid = i.indrelid \
# AND a.attnum = ANY(i.indkey) \
# WHERE i.indrelid = '{artists_table}'::regclass
# + editable=true
if not _output:
print('\033[93m'+'[WARNING] '+ f"The {artists_table} table does not have a primary key!")
# + editable=true
# %sql _output << SELECT a.attname, format_type(a.atttypid, a.atttypmod) AS data_type, a.attnotnull, i.indisprimary \
# FROM pg_index i \
# JOIN pg_attribute a ON a.attrelid = i.indrelid \
# AND a.attnum = ANY(i.indkey) \
# WHERE i.indrelid = '{songplay_table}'::regclass
# + editable=true
if not _output:
print('\033[93m'+'[WARNING] '+ f"The {songplay_table} table does not have a primary key!")
# + editable=true
# %sql _output << SELECT a.attname, format_type(a.atttypid, a.atttypmod) AS data_type, a.attnotnull, i.indisprimary \
# FROM pg_index i \
# JOIN pg_attribute a ON a.attrelid = i.indrelid \
# AND a.attnum = ANY(i.indkey) \
# WHERE i.indrelid = '{song_table}'::regclass
# + editable=true
if not _output:
print('\033[93m'+'[WARNING] '+ f"The {song_table} table does not have a primary key!")
# + [markdown] editable=true
# ## Run Data Type and Constraints Check
# + editable=true
# %sql _output << SELECT * FROM information_schema.columns where table_name='{user_table}'
# + [markdown] editable=true
# **Check the column `user_id` for correct data type.**
# If you get a `IndexError: single positional indexer is out-of-bounds` error, you may be using a different column name. Change the column name below and run the cell again.
# + editable=true
output = _output.DataFrame()
_dtype = output[output.column_name == 'user_id'].data_type.iloc[0]
if _dtype not in ['integer', 'bigint']:
print('\033[93m'+'[WARNING] '+ f"Type {_dtype} may not be an appropriate data type for column 'user_id' in the '{user_table}' table.")
# + editable=true
# %sql _output << SELECT * FROM information_schema.columns where table_name='{song_table}'
# + [markdown] editable=true
# **Check the column `year` for correct data type.
# Check columns `title` and `duration` for not-NULL constraints.**
#
# If you get a `IndexError: single positional indexer is out-of-bounds` error, you may be using different column names. Change the column name(s) below and run the cell again.
# + editable=true
output = _output.DataFrame()
_dtype = output[output.column_name == 'year'].data_type.iloc[0]
if _dtype not in ['integer']:
print('\033[93m'+'[WARNING] '+ f"Type '{_dtype}' may not be an appropriate data type for column 'year' in the '{song_table}' table.")
_nullable_title = output[output.column_name == 'title'].is_nullable.iloc[0]
_nullable_duration = output[output.column_name == 'duration'].is_nullable.iloc[0]
if (_nullable_duration != 'NO') or (_nullable_title != 'NO'):
print('\033[93m'+'[WARNING] '+ f"You may want to add appropriate NOT NULL constraints to the '{song_table}' table.")
# + editable=true
# %sql _output << SELECT * FROM information_schema.columns where table_name='{artists_table}'
# + [markdown] editable=true
# **Check the columns `latitude` and `longitude` for correct data type.
# Check column `name` for not-NULL constraint.**
#
# If you get a `IndexError: single positional indexer is out-of-bounds` error, you may be using different column names. Change the column name(s) below and run the cell again.
# + editable=true
output = _output.DataFrame()
_dtype_latitude = output[output.column_name == 'latitude'].data_type.iloc[0]
if _dtype_latitude not in ['double precision']:
print('\033[93m'+'[WARNING] '+ f"Type '{_dtype_latitude}' may not be an appropriate data type for column 'latitude' in the '{artists_table}' table")
_dtype_longitude = output[output.column_name == 'longitude'].data_type.iloc[0]
if _dtype_longitude not in ['double precision']:
print('\033[93m'+'[WARNING] '+ f"Type '{_dtype_longitude}' may not be an appropriate data type for column 'longitude' in the '{artists_table}' table")
_nullable_name = output[output.column_name == 'name'].is_nullable.iloc[0]
if _nullable_name != 'NO':
print('\033[93m'+'[WARNING] '+ f"You may want to add appropriate NOT NULL constraints to the '{artists_table}' table.")
# + editable=true
# %sql _output << SELECT * FROM information_schema.columns where table_name='{songplay_table}'
# + [markdown] editable=true
# **Check the columns `start_time` and `user_id` for correct data type.
# Check columns `start_time` and `user_id` for not-NULL constraint.**
#
# If you get a `IndexError: single positional indexer is out-of-bounds` error, you may be using different column names. Change the column name(s) below and run the cell again.
# + editable=true
output = _output.DataFrame()
_dtype_start_time = output[output.column_name == 'start_time'].data_type.iloc[0]
if 'timestamp' not in _dtype_start_time:
print('\033[93m'+'[WARNING] '+ f"Type '{_dtype_start_time}' may not be an appropriate data type for column 'start_time' in the '{songplay_table}' table.")
_dtype_user_id = output[output.column_name == 'user_id'].data_type.iloc[0]
if _dtype_user_id not in ['integer', 'bigint']:
print('\033[93m'+'[WARNING] '+ f"Type '{_dtype_user_id}' may not be an appropriate data type for column 'user_id' in the '{songplay_table}' table.")
_nullable_time = output[output.column_name == 'start_time'].is_nullable.iloc[0]
_nullable_uid = output[output.column_name == 'user_id'].is_nullable.iloc[0]
if (_nullable_time != 'NO') or (_nullable_uid != 'NO'):
print('\033[93m'+'[WARNING] '+ f"You may want to add appropriate NOT NULL constraints to the '{songplay_table}' table.")
# + [markdown] editable=true
# ## Run Tests for Upsertion Check
# + editable=true
import re
# + editable=true
if not re.search('ON\s+CONFLICT',sqlq.songplay_table_insert,re.IGNORECASE) or \
not re.search('ON\s+CONFLICT',sqlq.user_table_insert,re.IGNORECASE) or \
not re.search('ON\s+CONFLICT',sqlq.song_table_insert,re.IGNORECASE) or \
not re.search('ON\s+CONFLICT',sqlq.artist_table_insert,re.IGNORECASE):
print('\033[93m'+'[WARNING]Some of your insert queries may need an "ON CONFLICT" clause.')
print('\033[93m'+' You can either skip conflicting insertions with with "ON CONFLICT DO NOTHING"')
print('\033[93m'+' OR use "ON CONFLICT DO UPDATE SET"')
print('\033[93m'+' Check this link for more details: https://www.postgresqltutorial.com/postgresql-upsert/')
# + editable=true
|
data-modeling-with-postgres/test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Generating C Code for the Scalar Wave Equation in Cartesian Coordinates
#
# ## Authors: <NAME> & <NAME>
# ### Formatting improvements courtesy <NAME>
#
# ## This module generates the C Code for the Scalarwave in Cartesian coordinates and sets up either monochromatic plane wave or spherical Gaussian [Initial Data](https://en.wikipedia.org/wiki/Initial_value_problem).
#
# **Notebook Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented below ([right-hand-side expressions](#code_validation1); [initial data expressions](#code_validation2)). In addition, all expressions have been validated against a trusted code (the [original SENR/NRPy+ code](https://bitbucket.org/zach_etienne/nrpy)).
#
# ### NRPy+ Source Code for this module:
# * [ScalarWave/ScalarWave_RHSs.py](../edit/ScalarWave/ScalarWave_RHSs.py)
# * [ScalarWave/InitialData.py](../edit/ScalarWave/InitialData.py)
#
# ## Introduction:
# ### Problem Statement
#
# We wish to numerically solve the scalar wave equation as an [initial value problem](https://en.wikipedia.org/wiki/Initial_value_problem) in Cartesian coordinates:
# $$\partial_t^2 u = c^2 \nabla^2 u \text{,}$$
# where $u$ (the amplitude of the wave) is a function of time and space: $u = u(t,x,y,...)$ (spatial dimension as-yet unspecified) and $c$ is the wave speed, subject to some initial condition
#
# $$u(0,x,y,...) = f(x,y,...)$$
#
# and suitable spatial boundary conditions.
#
# As described in the next section, we will find it quite useful to define
# $$v(t,x,y,...) = \partial_t u(t,x,y,...).$$
#
# In this way, the second-order PDE is reduced to a set of two coupled first-order PDEs
#
# \begin{align}
# \partial_t u &= v \\
# \partial_t v &= c^2 \nabla^2 u.
# \end{align}
#
# We will use NRPy+ to generate efficient C codes capable of generating both initial data $u(0,x,y,...) = f(x,y,...)$; $v(0,x,y,...)=g(x,y,...)$, as well as finite-difference expressions for the right-hand sides of the above expressions. These expressions are needed within the *Method of Lines* to "integrate" the solution forward in time.
#
# ### The Method of Lines
#
# Once we have initial data, we "evolve it forward in time", using the [Method of Lines](https://reference.wolfram.com/language/tutorial/NDSolveMethodOfLines.html). In short, the Method of Lines enables us to handle
# 1. the **spatial derivatives** of an initial value problem PDE using **standard finite difference approaches**, and
# 2. the **temporal derivatives** of an initial value problem PDE using **standard strategies for solving ordinary differential equations (ODEs)**, so long as the initial value problem PDE can be written in the form
# $$\partial_t \vec{f} = \mathbf{M}\ \vec{f},$$
# where $\mathbf{M}$ is an $N\times N$ matrix filled with differential operators that act on the $N$-element column vector $\vec{f}$. $\mathbf{M}$ may not contain $t$ or time derivatives explicitly; only *spatial* partial derivatives are allowed to appear inside $\mathbf{M}$. The scalar wave equation as written in the [previous module](Tutorial-ScalarWave.ipynb)
# \begin{equation}
# \partial_t
# \begin{bmatrix}
# u \\
# v
# \end{bmatrix}=
# \begin{bmatrix}
# 0 & 1 \\
# c^2 \nabla^2 & 0
# \end{bmatrix}
# \begin{bmatrix}
# u \\
# v
# \end{bmatrix}
# \end{equation}
# satisfies this requirement.
#
# Thus we can treat the spatial derivatives $\nabla^2 u$ of the scalar wave equation using **standard finite-difference approaches**, and the temporal derivatives $\partial_t u$ and $\partial_t v$ using **standard approaches for solving ODEs**. In [the next module](Tutorial-Start_to_Finish-ScalarWave.ipynb), we will apply the highly robust [explicit Runge-Kutta fourth-order scheme](https://en.wikipedia.org/wiki/Runge%E2%80%93Kutta_methods) (RK4), used widely for numerically solving ODEs, to "march" (integrate) the solution vector $\vec{f}$ forward in time from its initial value ("initial data").
#
# ### Basic Algorithm
#
# The basic algorithm for solving the scalar wave equation [initial value problem](https://en.wikipedia.org/wiki/Initial_value_problem), based on the Method of Lines (see section above) is outlined below, with NRPy+-based components highlighted in <font color='green'>green</font>. We will review how NRPy+ generates these core components in this module.
#
# 1. Allocate memory for gridfunctions, including temporary storage for the RK4 time integration.
# 1. <font color='green'>Set gridfunction values to initial data.</font>
# 1. Evolve the system forward in time using RK4 time integration. At each RK4 substep, do the following:
# 1. <font color='green'>Evaluate scalar wave RHS expressions.</font>
# 1. Apply boundary conditions.
#
# **We refer to the right-hand side of the equation $\partial_t \vec{f} = \mathbf{M}\ \vec{f}$ as the RHS. In this case, we refer to the $\mathbf{M}\ \vec{f}$ as the "scalar wave RHSs".** In the following sections we will
#
# 1. Use NRPy+ to cast the scalar wave RHS expressions -- in finite difference form -- into highly efficient C code,
# 1. first in one spatial dimension with fourth-order finite differences,
# 1. and then in three spatial dimensions with tenth-order finite differences.
# 1. Use NRPy+ to generate monochromatic plane-wave initial data for the scalar wave equation, where the wave propagates in an arbitrary direction.
#
# As for the $\nabla^2 u$ term, spatial derivatives are handled in NRPy+ via [finite differencing](https://en.wikipedia.org/wiki/Finite_difference).
#
# We will sample the solution $\{u,v\}$ at discrete, uniformly-sampled points in space and time. For simplicity, let's assume that we consider the wave equation in one spatial dimension. Then the solution at any sampled point in space and time is given by
# $$u^n_i = u(t_n,x_i) = u(t_0 + n \Delta t, x_0 + i \Delta x),$$
# where $\Delta t$ and $\Delta x$ represent the temporal and spatial resolution, respectively. $v^n_i$ is sampled at the same points in space and time.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# 1. [Step 1](#initializenrpy): Initialize core NRPy+ modules
# 1. [Step 2](#rhss1d): Scalar Wave RHSs in One Spatial Dimension, Fourth-Order Finite Differencing
# 1. [Step 3](#rhss3d): Scalar Wave RHSs in Three Spatial Dimensions, Tenth-Order Finite Differencing
# 1. [Step 3.a](#code_validation1): Code Validation against `ScalarWave.ScalarWave_RHSs` NRPy+ module
# 1. [Step 4](#id): Setting up Initial Data for the Scalar Wave Equation
# 1. [Step 4.a](#planewave): The Monochromatic Plane-Wave Solution
# 1. [Step 4.b](#sphericalgaussian): The Spherical Gaussian Solution (*Courtesy <NAME>*)
# 1. [Step 5](#code_validation2): Code Validation against `ScalarWave.InitialData` NRPy+ module
# 1. [Step 6](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Initialize core NRPy+ modules \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# Let's start by importing all the needed modules from NRPy+:
# Step P1: Import needed NRPy+ core modules:
import NRPy_param_funcs as par # NRPy+: Parameter interface
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import grid as gri # NRPy+: Functions having to do with numerical grids
import finite_difference as fin # NRPy+: Finite difference C code generation module
from outputC import lhrh # NRPy+: Core C code output module
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
# <a id='rhss1d'></a>
#
# # Step 2: Scalar Wave RHSs in One Spatial Dimension, Fourth-Order Finite Differencing \[Back to [top](#toc)\]
# $$\label{rhss1d}$$
#
# To minimize complication, we will first restrict ourselves to solving the wave equation in one spatial dimension, so
# $$\nabla^2 u = \partial_x^2 u.$$
# Extension of this operator to higher spatial dimensions is straightforward, particularly when using NRPy+.
#
# As was discussed in [the finite difference section of the tutorial](Tutorial-Finite_Difference_Derivatives.ipynb), NRPy+ approximates derivatives using [finite difference methods](), the second-order derivative $\partial_x^2$ accurate to fourth-order in uniform grid spacing $\Delta x$ (from fitting the unique 4th-degree polynomial to 5 sample points of $u$) is given by
# \begin{equation}
# \left[\partial_x^2 u(t,x)\right]_j = \frac{1}{(\Delta x)^2}
# \left(
# -\frac{1}{12} \left(u_{j+2} + u_{j-2}\right)
# + \frac{4}{3} \left(u_{j+1} + u_{j-1}\right)
# - \frac{5}{2} u_j \right)
# + \mathcal{O}\left((\Delta x)^4\right).
# \end{equation}
# +
# Step P2: Define the C parameter wavespeed. The `wavespeed`
# variable is a proper SymPy variable, so it can be
# used in below expressions. In the C code, it acts
# just like a usual parameter, whose value is
# specified in the parameter file.
thismodule = "ScalarWave"
wavespeed = par.Cparameters("REAL",thismodule,"wavespeed", 1.0)
# Step 1: Set the spatial dimension parameter, and then read
# the parameter as DIM.
par.set_parval_from_str("grid::DIM",1)
DIM = par.parval_from_str("grid::DIM")
# Step 2: Set the finite differencing order to 4.
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER",4)
# Step 3: Register gridfunctions that are needed as input
# to the scalar wave RHS expressions.
uu, vv = gri.register_gridfunctions("EVOL",["uu","vv"])
# Step 4: Declare the rank-2 indexed expression \partial_{ij} u,
# which is symmetric about interchange of indices i and j
# Derivative variables like these must have an underscore
# in them, so the finite difference module can parse the
# variable name properly.
uu_dDD = ixp.declarerank2("uu_dDD","sym01")
# Step 5: Define right-hand sides for the evolution.
uu_rhs = vv
vv_rhs = 0
for i in range(DIM):
vv_rhs += wavespeed*wavespeed*uu_dDD[i][i]
vv_rhs = sp.simplify(vv_rhs)
# Step 6: Generate C code for scalarwave evolution equations,
# print output to the screen (standard out, or stdout).
fin.FD_outputC("stdout",
[lhrh(lhs=gri.gfaccess("rhs_gfs","uu"),rhs=uu_rhs),
lhrh(lhs=gri.gfaccess("rhs_gfs","vv"),rhs=vv_rhs)])
# -
# **Success!** Notice that indeed NRPy+ was able to compute the spatial derivative operator,
# \begin{equation}
# \left[\partial_x^2 u(t,x)\right]_j \approx \frac{1}{(\Delta x)^2}
# \left(
# -\frac{1}{12} \left(u_{j+2} + u_{j-2}\right)
# + \frac{4}{3} \left(u_{j+1} + u_{j-1}\right)
# - \frac{5}{2} u_j \right),
# \end{equation}
# correctly (easier to read in the "Original SymPy expressions" comment block at the top of the C output. Note that `invdx0`$=1/\Delta x_0$, where $\Delta x_0$ is the (uniform) grid spacing in the zeroth, or $x_0$ direction.
# <a id='rhss3d'></a>
#
# # Step 3: Scalar Wave RHSs in Three Spatial Dimensions, Tenth-Order Finite Differencing \[Back to [top](#toc)\]
# $$\label{rhss3d}$$
#
# Let's next repeat the same process, only this time at **10th** finite difference order, for the **3-spatial-dimension** scalar wave equation, with SIMD enabled:
# +
# Step 1: Define the C parameter wavespeed. The `wavespeed`
# variable is a proper SymPy variable, so it can be
# used in below expressions. In the C code, it acts
# just like a usual parameter, whose value is
# specified in the parameter file.
wavespeed = par.Cparameters("REAL",thismodule,"wavespeed", 1.0)
# Step 2: Set the spatial dimension parameter
# to *FOUR* this time, and then read
# the parameter as DIM.
par.set_parval_from_str("grid::DIM",3)
DIM = par.parval_from_str("grid::DIM")
# Step 3: Set the finite differencing order to 10.
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER",10)
# Step 4a: Reset gridfunctions registered in 1D case above,
# to avoid NRPy+ throwing an error about double-
# registering gridfunctions, which is not allowed.
gri.glb_gridfcs_list = []
# Step 4b: Register gridfunctions that are needed as input
# to the scalar wave RHS expressions.
uu, vv = gri.register_gridfunctions("EVOL",["uu","vv"])
# Step 5: Declare the rank-2 indexed expression \partial_{ij} u,
# which is symmetric about interchange of indices i and j
# Derivative variables like these must have an underscore
# in them, so the finite difference module can parse the
# variable name properly.
uu_dDD = ixp.declarerank2("uu_dDD","sym01")
# Step 6: Define right-hand sides for the evolution.
uu_rhs = vv
vv_rhs = 0
for i in range(DIM):
vv_rhs += wavespeed*wavespeed*uu_dDD[i][i]
# Step 7: Simplify the expression for c^2 \nabla^2 u (a.k.a., vv_rhs):
vv_rhs = sp.simplify(vv_rhs)
# Step 8: Generate C code for scalarwave evolution equations,
# print output to the screen (standard out, or stdout).
fin.FD_outputC("stdout",
[lhrh(lhs=gri.gfaccess("rhs_gfs","uu"),rhs=uu_rhs),
lhrh(lhs=gri.gfaccess("rhs_gfs","vv"),rhs=vv_rhs)],params="SIMD_enable=True")
# -
# <a id='code_validation1'></a>
#
# ## Step 3.a: Code Validation against `ScalarWave.ScalarWave_RHSs` NRPy+ module \[Back to [top](#toc)\]
# $$\label{code_validation1}$$
#
# Here, as a code validation check, we verify agreement in the SymPy expressions for the RHSs of the three-spatial-dimension Scalar Wave equation (i.e., `uu_rhs` and `vv_rhs`) between
#
# 1. this tutorial and
# 2. the [NRPy+ ScalarWave.ScalarWave_RHSs](../edit/ScalarWave/ScalarWave_RHSs.py) module.
# +
# Step 10: We already have SymPy expressions for uu_rhs and vv_rhs in
# terms of other SymPy variables. Even if we reset the list
# of NRPy+ gridfunctions, these *SymPy* expressions for
# uu_rhs and vv_rhs *will remain unaffected*.
#
# Here, we will use the above-defined uu_rhs and vv_rhs to
# validate against the same expressions in the
# ScalarWave/ScalarWave_RHSs.py module,
# to ensure consistency between this tutorial
# (historically speaking, the tutorial was written first)
# and the ScalarWave_RHSs.py module itself.
#
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
# Step 11: Call the ScalarWave_RHSs() function from within the
# ScalarWave/ScalarWave_RHSs.py module,
# which should do exactly the same as in Steps 1-10 above.
import ScalarWave.ScalarWave_RHSs as swrhs
swrhs.ScalarWave_RHSs()
# Step 12: Consistency check between the tutorial notebook above
# and the ScalarWave_RHSs() function from within the
# ScalarWave/ScalarWave_RHSs.py module.
print("Consistency check between ScalarWave tutorial and NRPy+ module:")
print("uu_rhs - swrhs.uu_rhs = "+str(sp.simplify(uu_rhs - swrhs.uu_rhs))+"\t\t (should be zero)")
print("vv_rhs - swrhs.vv_rhs = "+str(sp.simplify(vv_rhs - swrhs.vv_rhs))+"\t\t (should be zero)")
# -
# <a id='id'></a>
#
# # Step 4: Setting up Initial Data for the Scalar Wave Equation \[Back to [top](#toc)\]
# $$\label{id}$$
#
# <a id='planewave'></a>
#
# ## Step 4.a: The Monochromatic Plane-Wave Solution \[Back to [top](#toc)\]
# $$\label{planewave}$$
#
# The solution to the scalar wave equation for a monochromatic (single-wavelength) wave traveling in the $\hat{k}$ direction is
# $$u(\vec{x},t) = f(\hat{k}\cdot\vec{x} - c t),$$
# where $\hat{k}$ is a unit vector. We choose $f(\hat{k}\cdot\vec{x} - c t)$ to take the form
# $$
# f(\hat{k}\cdot\vec{x} - c t) = \sin\left(\hat{k}\cdot\vec{x} - c t\right) + 2,
# $$
# where we add the $+2$ to ensure that the exact solution never crosses through zero. In places where the exact solution passes through zero, the relative error (i.e., the measure of error to compare numerical with exact results) is undefined. Also, $f(\hat{k}\cdot\vec{x} - c t)$ plus a constant is still a solution to the wave equation.
# +
# Step 1: Set parameters defined in other modules
xx = gri.xx # Sets the Cartesian coordinates xx[0]=x; xx[1]=y; xx[2]=z
# Step 2: Declare free parameters intrinsic to these initial data
time = par.Cparameters("REAL", thismodule, "time",0.0)
kk = par.Cparameters("REAL", thismodule, ["kk0", "kk1", "kk2"],[1.0,1.0,1.0])
# Step 3: Normalize the k vector
kk_norm = sp.sqrt(kk[0]**2 + kk[1]**2 + kk[2]**2)
# Step 4: Compute k.x
dot_product = sp.sympify(0)
for i in range(DIM):
dot_product += xx[i]*kk[i]
dot_product /= kk_norm
# Step 5: Set initial data for uu and vv, where vv_ID = \partial_t uu_ID.
uu_ID_PlaneWave = sp.sin(dot_product - wavespeed*time)+2
vv_ID_PlaneWave = sp.diff(uu_ID_PlaneWave, time)
# -
# Next we verify that $f(\hat{k}\cdot\vec{x} - c t)$ satisfies the wave equation, by computing
# $$\left(c^2 \nabla^2 - \partial_t^2 \right)\ f\left(\hat{k}\cdot\vec{x} - c t\right),$$
# and confirming the result is exactly zero.
sp.simplify(wavespeed**2*(sp.diff(uu_ID_PlaneWave,xx[0],2) +
sp.diff(uu_ID_PlaneWave,xx[1],2) +
sp.diff(uu_ID_PlaneWave,xx[2],2))
- sp.diff(uu_ID_PlaneWave,time,2))
# <a id='sphericalgaussian'></a>
#
# ## Step 4.b: The Spherical Gaussian Solution \[Back to [top](#toc)\]
# $$\label{sphericalgaussian}$$
#
# Here we will implement the spherical Gaussian solution, consists of ingoing and outgoing wave fronts:
# \begin{align}
# u(r,t) &= u_{\rm out}(r,t) + u_{\rm in}(r,t),\ \ \text{where}\\
# u_{\rm out}(r,t) &=\frac{r-ct}{r} \exp\left[\frac{-(r-ct)^2}{2 \sigma^2}\right] \\
# u_{\rm in}(r,t) &=\frac{r+ct}{r} \exp\left[\frac{-(r+ct)^2}{2 \sigma^2}\right] \\
# \end{align}
# where $c$ is the wavespeed, and $\sigma$ is the width of the Gaussian (i.e., the "standard deviation").
# +
# Step 1: Set parameters defined in other modules
xx = gri.xx # Sets the Cartesian coordinates xx[0]=x; xx[1]=y; xx[2]=z
# Step 2: Declare free parameters intrinsic to these initial data
time = par.Cparameters("REAL", thismodule, "time",0.0)
sigma = par.Cparameters("REAL", thismodule, "sigma",3.0)
# Step 4: Compute r
r = sp.sympify(0)
for i in range(DIM):
r += xx[i]**2
r = sp.sqrt(r)
# Step 5: Set initial data for uu and vv, where vv_ID = \partial_t uu_ID.
uu_ID_SphericalGaussianOUT = +(r - wavespeed*time)/r * sp.exp( -(r - wavespeed*time)**2 / (2*sigma**2) )
uu_ID_SphericalGaussianIN = +(r + wavespeed*time)/r * sp.exp( -(r + wavespeed*time)**2 / (2*sigma**2) )
uu_ID_SphericalGaussian = uu_ID_SphericalGaussianOUT + uu_ID_SphericalGaussianIN
vv_ID_SphericalGaussian = sp.diff(uu_ID_SphericalGaussian, time)
# -
# Since the wave equation is linear, both the leftgoing and rightgoing waves must satisfy the wave equation, which implies that their sum also satisfies the wave equation.
#
# Next we verify that $u(r,t)$ satisfies the wave equation, by computing
# $$\left(c^2 \nabla^2 - \partial_t^2 \right)\left\{u_{\rm R}(r,t)\right\},$$
#
# and
#
# $$\left(c^2 \nabla^2 - \partial_t^2 \right)\left\{u_{\rm L}(r,t)\right\},$$
#
# are separately zero. We do this because SymPy has difficulty simplifying the combined expression.
# +
print(sp.simplify(wavespeed**2*(sp.diff(uu_ID_SphericalGaussianOUT,xx[0],2) +
sp.diff(uu_ID_SphericalGaussianOUT,xx[1],2) +
sp.diff(uu_ID_SphericalGaussianOUT,xx[2],2))
- sp.diff(uu_ID_SphericalGaussianOUT,time,2)) )
print(sp.simplify(wavespeed**2*(sp.diff(uu_ID_SphericalGaussianIN,xx[0],2) +
sp.diff(uu_ID_SphericalGaussianIN,xx[1],2) +
sp.diff(uu_ID_SphericalGaussianIN,xx[2],2))
- sp.diff(uu_ID_SphericalGaussianIN,time,2)))
# -
# <a id='code_validation2'></a>
#
# # Step 5: Code Validation against `ScalarWave.InitialData` NRPy+ module \[Back to [top](#toc)\]
# $$\label{code_validation2}$$
#
# As a code validation check, we will verify agreement in the SymPy expressions for plane-wave initial data for the Scalar Wave equation between
# 1. this tutorial and
# 2. the NRPy+ [ScalarWave.InitialData](../edit/ScalarWave/InitialData.py) module.
# +
# We just defined SymPy expressions for uu_ID and vv_ID in
# terms of other SymPy variables. Here, we will use the
# above-defined uu_ID and vv_ID to validate against the
# same expressions in the ScalarWave/InitialData.py
# module, to ensure consistency between this tutorial
# (historically speaking, the tutorial was written first)
# and the PlaneWave ID module itself.
#
# Step 6: Call the InitialData(Type="PlaneWave") function from within the
# ScalarWave/InitialData.py module,
# which should do exactly the same as in Steps 1-5 above.
import ScalarWave.InitialData as swid
swid.InitialData(Type="PlaneWave")
# Step 7: Consistency check between the tutorial notebook above
# and the PlaneWave option from within the
# ScalarWave/InitialData.py module.
print("Consistency check between ScalarWave tutorial and NRPy+ module: PlaneWave Case")
if sp.simplify(uu_ID_PlaneWave - swid.uu_ID) != 0:
print("TEST FAILED: uu_ID_PlaneWave - swid.uu_ID = "+str(sp.simplify(uu_ID_PlaneWave - swid.uu_ID))+"\t\t (should be zero)")
sys.exit(1)
if sp.simplify(vv_ID_PlaneWave - swid.vv_ID) != 0:
print("TEST FAILED: vv_ID_PlaneWave - swid.vv_ID = "+str(sp.simplify(vv_ID_PlaneWave - swid.vv_ID))+"\t\t (should be zero)")
sys.exit(1)
print("TESTS PASSED!")
# Step 8: Consistency check between the tutorial notebook above
# and the SphericalGaussian option from within the
# ScalarWave/InitialData.py module.
swid.InitialData(Type="SphericalGaussian")
print("Consistency check between ScalarWave tutorial and NRPy+ module: SphericalGaussian Case")
if sp.simplify(uu_ID_SphericalGaussian - swid.uu_ID) != 0:
print("TEST FAILED: uu_ID_SphericalGaussian - swid.uu_ID = "+str(sp.simplify(uu_ID_SphericalGaussian - swid.uu_ID))+"\t\t (should be zero)")
sys.exit(1)
if sp.simplify(vv_ID_SphericalGaussian - swid.vv_ID) != 0:
print("TEST FAILED: vv_ID_SphericalGaussian - swid.vv_ID = "+str(sp.simplify(vv_ID_SphericalGaussian - swid.vv_ID))+"\t\t (should be zero)")
sys.exit(1)
print("TESTS PASSED!")
# -
# <a id='latex_pdf_output'></a>
#
# # Step 6: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ScalarWave.pdf](Tutorial-ScalarWave.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-ScalarWave")
|
Tutorial-ScalarWave.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tree_dev]
# language: python
# name: conda-env-tree_dev-py
# ---
# ## Interoperability with sklearn
# In this notebook, we demonstrate the interoperability of arboretum with sklearn.model_selection for cross-validation and parameter search. We will also use an example involving feature selection and a pipeline. We will be working with the ALS dataset. This is a wide noisy dataset that tree models struggle with.
# +
from arboretum.datasets import load_als
from arboretum import RFRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import mean_squared_error as mse
xtr, ytr, xte, yte = load_als()
rf = RandomForestRegressor(n_estimators=100, min_samples_leaf=5)
rf.fit(xtr, ytr)
myrf = RFRegressor(n_trees=100, min_leaf=5)
myrf.fit(xtr, ytr)
# -
pred = rf.predict(xte)
mypred = myrf.predict(xte)
mse(yte, pred), mse(yte, mypred)
# ### Grid Search CV
# Next, we run a one-parameter grid search for these models on the minimium leaf size. In order to speed things up in the notebook, we'll limit the maximum number of features tried to 30.
rf.max_features = 30
params = {'min_samples_leaf':[1, 5, 10, 20]}
gcv = GridSearchCV(rf, params, 'neg_mean_squared_error')
gcv.fit(xtr, ytr)
pred = gcv.predict(xte)
mse(yte, pred), gcv.best_score_, gcv.best_params_
myrf.max_features = 30
myparams = {'min_leaf':[1, 5, 10, 20]}
mygcv = GridSearchCV(myrf, myparams, 'neg_mean_squared_error')
mygcv.fit(xtr, ytr)
mypred = mygcv.predict(xte)
mse(yte, mypred), mygcv.best_score_, mygcv.best_params_
# ### Pipeline/Feature Selection
# Next we'll set up a pipeline with a simple univariate feature selection method, and our model. We'll set the models back to using all features now that feature selection is being used.
from sklearn.pipeline import Pipeline
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_regression
rf.max_features = None
skb = SelectKBest(f_regression, k=30)
pipe = Pipeline([('select', skb), ('model', rf)])
pipe.fit(xtr, ytr)
pred = pipe.predict(xte)
mse(yte, pred)
myrf.max_features = None
mypipe = Pipeline([('select', skb), ('model', myrf)])
mypipe.fit(xtr, ytr)
mypred = mypipe.predict(xte)
mse(yte, mypred)
# ### Conclusion
# A lot of the value of scikit-learn is in the 'plumbing' code for repetitive tasks like cross-validation, evaluation, and feature selection. In this notebook, we showed how to use arboretum with these parts of sklearn.
|
examples/sklearn-interop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env_p7a
# language: python
# name: env_p7a
# ---
# # Tree Shap
#
# To get a sense of the Shap library that we'll be using, let's implement the simple version of the Tree Shap algorithm. This is based on Scott Lundberg's paper [Consistent Individualized Feature Attribution for Tree
# Ensembles](https://arxiv.org/pdf/1802.03888.pdf)
# #### Training and re-using a single tree model
#
# You may notice that calculating Shap values for every feature, and for every individual data point, is very computationally expensive. For example, we would be training multiple models just to calculate the importance of one feature.
#
# With decision trees, we can actually train a decision tree on all the features, and then re-use that single tree to calculate shapley values using subsets of that single tree.
#
# 
# For some intuition, let’s look at a model that has just two features. The tree splits on feature one first, and then on feature 2.
# 
# If we wanted to see how a model would perform if it only used the first feature, we could look at the subtree that consists of the top three nodes of this tree.
# 
# Similarly, if we wanted to see how a model would make predictions if it used only feature 2, we could look at the subtree containing the bottom three nodes, starting at the node that splits on feature 2.
# 
# #### Another example with 3 features
# Now let’s look at a tree that is trained on three features. Let’s say it splits on feature 1, then on feature 2, then on feature 3.
# 
# How do we simulate the prediction of a tree that was only trained on features 1 and 3, but not on feature 2?
# 
# Well, if we didn’t split on feature 2, that means that we would include the training samples in both the left and right sub-tree of that node when making a prediction. This is how we can simulate that the tree never split on feature 2.
# 
# Let’s also think about how we handle the predictions when we do split on the feature. If we split on feature 3, and the particular data point we’re making a prediction for ends up in the left child node, then we can use the prediction based on training samples in the left sub-tree, and ignore the training samples in the right sub-tree.
# 
# We’ll walk through the algorithm to do this, and then you’ll get to practice this yourself.
# #### Algorithm
#
# Here’s the algorithm used to calculate the prediction of a tree, given a subset of features. You can check out the paper [Consistent Individualized Feature Attribution for Tree Ensembles](https://arxiv.org/pdf/1802.03888.pdf), page 4 algorithm 1.
# 
# Here, $G$ is a function that gets called recursively to walk down the tree starting at the root node. $w$ is the weight given to the predictions of each node. $v$ is the prediction of a leaf node. $r_{a_j}$ and $r_{b_j}$ are the number of data points in the left and right child nodes of node $j$. $r_j$ is the number of data points in node $j$.
#
# We can use this to walk through a decision tree that is trained on all features, and calculate the prediction of a tree that would have been created from a subset of the features.
#
# Let's look at specific parts of this algorithm in more detail.
# #### leaf nodes
# Let's look at the the line that handles leaf nodes. It takes the prediction of that leaf node and multiplies it by some weight. The weight is determined by the proportion of training data points that end up reaching that leaf node.
# 
# #### ignoring a feature
# Next, let’s look at the case when the feature that’s used at a node is not within the subset of features that we want to split on. In other words, we want to pretend that we didn’t train the model on this feature. In that case, in order to pretend that we’re not splitting on that feature, we take the sum of the weighted predictions from both its left and right subtree.
# 
# #### including a feature
#
# Finally, for cases when the feature at that node is within the subset of features that we want to use, then we can follow just the left subtree or just the right subtree, whichever path that the input data gets assigned to by the split.
# 
# #### Implement it in code!
# You’ll get to practice this algorithm!
import sys
# !{sys.executable} -m pip install numpy==1.14.5
# !{sys.executable} -m pip install scikit-learn==0.19.1
# !{sys.executable} -m pip install graphviz==0.9
# !{sys.executable} -m pip install shap==0.25.2
import sklearn.ensemble
import shap
import numpy as np
import graphviz
# ## generate sample data
#
# Feature 0 and feature 1 form the AND operator, and feature 2 does not contribute to the prediction of the label, because it's always zero.
# AND case (features 0 and 1)
N = 100
M = 3
X = np.zeros((N,M))
X.shape
y = np.zeros(N)
X[:1 * N//4, 1] = 1
X[:N//2, 0] = 1
X[N//2:3 * N//4, 1] = 1
y[:1 * N//4] = 1
# ## Train a decision tree
#
# +
# fit model
model = sklearn.tree.DecisionTreeRegressor(random_state=0)
model.fit(X, y)
# draw model
dot_data = sklearn.tree.export_graphviz(model, out_file=None, filled=True, rounded=True, special_characters=True)
graph = graphviz.Source(dot_data)
graph
# -
# ## Tree attributes
#
# [sklearn.tree.tree._tree](https://github.com/scikit-learn/scikit-learn/blob/master/sklearn/tree/_tree.pyx)
#
# ```
# The binary tree is represented as a number of parallel arrays. The i-th
# element of each array holds information about the node `i`. Node 0 is the
# tree's root.
# ```
tree0 = model.tree_
# #### left and right child nodes
# ```
# children_left : array of int, shape [node_count]
# children_left[i] holds the node id of the left child of node i.
# For leaves, children_left[i] == TREE_LEAF. Otherwise,
# children_left[i] > i. This child handles the case where
# X[:, feature[i]] <= threshold[i].
# children_right : array of int, shape [node_count]
# children_right[i] holds the node id of the right child of node i.
# For leaves, children_right[i] == TREE_LEAF. Otherwise,
# children_right[i] > i. This child handles the case where
# X[:, feature[i]] > threshold[i].
# ```
print(f"tree0.children_left: {tree0.children_left}")
print(f"tree0.children_right: {tree0.children_right}")
# #### features
# ```
# feature : array of int, shape [node_count]
# feature[i] holds the feature to split on, for the internal node i.
# threshold : array of double, shape [node_count]
# threshold[i] holds the threshold for the internal node i.
# value : array of double, shape [node_count, n_outputs, max_n_classes]
# Contains the constant prediction value of each node.
# impurity : array of double, shape [node_count]
# impurity[i] holds the impurity (i.e., the value of the splitting
# criterion) at node i.
# ```
print(f"tree0.feature: {tree0.feature}")
# For Node 0, feature 1 is used to split the data . For Node 2, feature 0 is used for splitting. For the other nodes (1, 3, 4), there are no features used for splitting.
# #### Thresholds
print(f"tree0.threshold: {tree0.threshold}")
# The threshold divides the data points using the chosen feature. <= 0.5 and the data go to the left child; > .5 and the data go in the right child. The -2 is for nodes that don't split on any feature.
# #### Value
print(f"tree0.value : \n{tree0.value}")
# `value` is the average prediction for each node. Node 0 predicts 0.25 on average. Node 2 predicts 0.5 on average.
# #### node samples
# ```
# n_node_samples : array of int, shape [node_count]
# n_node_samples[i] holds the number of training samples reaching node i.
#
# weighted_n_node_samples : array of int, shape [node_count]
# weighted_n_node_samples[i] holds the weighted number of training samples
# reaching node i.
# ```
print(f"tree0.n_node_samples : {tree0.n_node_samples}")
print(f"tree0.weighted_n_node_samples : {tree0.weighted_n_node_samples}")
# The n_node_samples counts how many data points from the parent gets passed to each child node. `weighted_n_node_samples` is the same in this case, because it's a single decision tree. If this were a tree within a random forest, 2/3 of the training data may be sampled and used to train a tree. The `weighted_n_node_samples` would then be re-scaled to equal the total sample size. We can use either in the calculations we'll do below.
# #### Quiz
# What proportion of samples went to the left child and right child of the root node?
# TODO
proportion_in_left_child = # ...
proportion_in_right_child = # ...
print(f"proportion of samples in left child of root node {proportion_in_left_child}")
print(f"proportion of samples in left child of root node {proportion_in_right_child}")
# ## Wrap with Tree class
#
# To make the tree object easier to work with, we'll wrap it inside our custom Tree class. Please complete the functions within the Tree class below.
# **Bonus Challenge:** Try implementing your own wrapper class for the tree object.
#
# Think about attributes that you may need in order to implement algorithm 1. For example, how do we know when a node is an internal or leaf node? What fraction of samples are in the left child relative to its parent node? On which node is each feature split on?
# +
"""
Challenge: try implementing your own wrapper class
"""
# -
# **If you prefer some starter code:** You can also use the starter code below if you prefer.
NO_NODE = -1
NO_FEATURE = -2
class Tree:
def __init__(self, tree):
if str(type(tree)).endswith("'sklearn.tree._tree.Tree'>"):
self.weight = 1
self.children_left = tree.children_left
self.children_right = tree.children_right
self.features = tree.feature
self.thresholds = tree.threshold
self.values = tree.value[:,0,0] # tree.value is n by 1 by 1; get the n prediction values (there are n nodes in tree)
self.n_node_samples = tree.n_node_samples # actual number of nodes
self.node_sample_weight = tree.weighted_n_node_samples #rescaled number of nodes
def is_internal(self,i):
return (self.children_left[i] != NO_NODE or self.children_right[i] != NO_NODE)
def is_leaf(self,i):
# TODO
def left_child(self,i):
return self.children_left[i]
def right_child(self,i):
# TODO
def proportion_of_samples_in_left_child(self,i):
# TODO
def proportion_of_samples_in_right_child(self,i):
# TODO
def node_prediction(self,i):
return self.values[i]
def feature_that_split_node_i(self,i):
return self.features[i]
def threshold_at_node_i(self,i):
return self.thresholds[i]
tree_wrap = Tree(tree0)
# #### Calculate the prediction of a tree model, given a subset of features
#
# We'll implement algorithm 1 of Scott Lundberg's paper. This is a way to use a single trained tree to estimate predictions of other trees that would be trained on a subset of the features.
# **Bonus challenge:** Try implementing this function completely by yourself!
"""Bonus challenge: implement f_given_S on your own!"""
def f_given_S(tree, S, x):
"""
tree: the custom Tree class
S: set of integers reprenting features that are used to train the model.
x: sample observation on which to calculate the prediction of the model.
"""
pass
# **If you prefer some starter code:** You can also use the starter code below to implement the algorithm.
"""
You can use this starter code if you get stuck while implementing the algorithm on your own
"""
def f_given_S(tree, S, x):
"""
tree: the custom Tree class
S: set of integers reprenting features that are used to train the model.
x: sample observation on which to calculate the prediction of the model.
"""
# the root node is at index 0 in the list
starting_node = 0
# When starting at the root node, the weight assigned is 1 (100%).
starting_weight = 1
def traverse_tree(node_i, weight):
"""
nested function that will be called recursively
"""
if tree.is_leaf(node_i):
# TODO: multiply the weight times the node prediction
else: # is internal node
feature_index = tree.feature_that_split_node_i(node_i)
feature_value = x[feature_index]
threshold = tree.threshold_at_node_i(node_i)
left_child = tree.left_child(node_i)
right_child = tree.right_child(node_i)
if feature_index in S:
if feature_value <= threshold:
# TODO: recursively traverse the left subtree
else:
# TODO: recursively traverse the right subtree
else: #feature is not in subset S
# TODO: traverse the left sub-tree,
# and update the weight to be the current weight times the proportion of samples in the left child node
# TODO: traverse the right sub-tree,
# and update the weight to be the current weight times the proportion of samples in the left child node
# TODO: return the sum of both sub-trees
# start traversing the tree
return traverse_tree(starting_node,starting_weight)
# #### Try out the function
sample_values = np.array([1,1,1])
S = set([2]) # if you input only feature 2, expect 0.25
f_given_S(tree_wrap, S, sample_values)
# #### try empty feature set
# +
S = set([]) #for empty set, expect output to be 0.25
f_given_S(tree_wrap, S, sample_values)
# -
# ## calculate weight on marginal contribution
#
# We'll calculate the weight placed on the marginal contribution of the feature:
# $ \frac{|S|! (M - |S| -1 )!}{M!}$
from math import factorial
def weight_on_marginal_contribution(size_S, M):
"""
size_S: number of features in set S
M: numer of total features
"""
# TODO
# ## calculate marginal contribution for a single feature
#
# $ f(S \cup i) - f(S)$
#
# Fill in the function that takes in the custom Tree object, a sample data point, a list containing the set of features in set S (excluding feature "i"), and also the feature for which we want to calculate the marginal contribution. Keep in mind that set S excludes feature "i".
#
# **Hint:** The python `set` class has the member function `.add`.
# Note that you may need to use the `.copy` function as well.
def marginal_contribution_of_feature(tree,x,S,feature_i):
"""
tree: the custom Tree object that wraps the sklearn tree_ object
x: a sample observation that contains all features
S: a list of integers, specifying the features in subset S, excluding feature i.
feature_i: an integer specifying the feature for which we're calculating the marginal contribution.
"""
# TODO: create the union of S and i
# TODO: return the difference in prediction with feature "i" and without feature "i"
# #### Try it out
#
# We'll try out the `marginal_contribution_of_feature` function.
feature_i = 0 #index of feature for which we want to calculate its marginal contribution
S = set([1]) # Set that excludes feature i
x = X[0] #grab one data point to calculate marginal contribution on
marginal_contribution_of_feature(tree_wrap,x,S,feature_i) # we expect 0.5
# The marginal contribution of feature 0 is 0.5. This means that the prediction of the model when feature 0 is present is 0.5 greater than the model's prediction when it only has feature 1.
# ## Generate all subsets
#
# Fill in a function that generates all possible subsets S.
# We'll use `itertools.combinations`, which takes in a list, and also the size of each subset. It returns an iterable object that contains tuples of all the combinations.
#
#
# +
from itertools import combinations
# try out the combinations function
tmp_combo = combinations([1,2,3,4],2)
for subset in tmp_combo:
print(subset)
# -
# #### fill in the function generate_all_subsets
#
# Keep in mind that since the iterable returned by `combinations` holds tuples, we can create sets out of the tuples by using `set(the_tuple_object)`. We'll store the S sets as `set` types, since we defined the `f_given_S` function to take S as a type `set`.
#
# Remember to also include the empty set. We can do this with `set([None])`
# ** Bonus challenge:** Try implementing this function on your own!
"""
Try implementing on your own!
"""
def generate_all_subsets(S):
"""
S: set of integers representing the features in set S
"""
pass
# **If you prefer some starter code: ** You can also use the starter code if you get stuck
# +
"""
Starter code version
"""
def generate_all_subsets(S):
"""
S: set of integers representing the features in set S
"""
sets_l = []
for size in range(1,len(S)+1):
# TODO: create a combinations iterable
# TODO: loop thru the combo iterable and append sets to the sets_l list
# TODO: also include the empty set
return sets_l
# -
# #### Try out the function
S = set([0,1,2])
generate_all_subsets(S)
# ## Calculate shapley value for one feature
#
# Implement a function that calculates the shapley value for a single feature, by iterating across all subsets S.
# **Bonus challenge:** Try implementing the function yourself!
def shap_feature_i(tree,x,feature_i):
"""
tree: the custom Tree object that wraps the tree_ from sklearn.
x: a sample data point
i: the feature for which we want to calculate its shapley value.
"""
pass
# **If you prefer starter code:** You can also fill in the starter code below if you prefer.
def shap_feature_i(tree,x,feature_i):
"""
tree: the custom Tree object that wraps the tree_ from sklearn.
x: a sample data point
i: the feature for which we want to calculate its shapley value.
"""
all_features = set(np.arange(0,x.shape[0]))
all_features_minus_i = all_features.copy()
all_features_minus_i.remove(feature_i) #remove feature "i"
# TODO: generate all subsets S
S_list = # ...
phi = # ...
num_features_total = len(all_features)
# iterate thru S_list
for S in S_list:
# TODO: calculate the number of features stored in S
# Handle the special case where S contains None, because the number of features should be 0 in that case
if None in S and len(S) == 1:
# ...
else:
# ...
# TODO: increment phi by the weigth on the marginal contribution * marginal contribution of "i"
phi += # ...
return np.round(phi,decimals=3)
return phi
# #### Try out the function
x = X[0]
shap_feature_i(tree_wrap,x,0)
# ## Calculate feature importance of all features
def shap_tree_explainer(tree_wrap, x):
shap_l = []
for i,v in enumerate(x):
shap_l.append(shap_feature_i(tree_wrap, x,i))
return np.array(shap_l)
# ## Take an sklearn tree model and calculate feature importance
def shap_tree_model_explainer(tree_model, x):
sklearn_tree = tree_model.tree_
tree_wrap = Tree(tree_model.tree_)
return shap_tree_explainer(tree_wrap,x)
# # Additive Feature Attribution
#
# Additive feature attribution methods are simple models that are used to explain complex models. You can see the formula in the same paper on page 3.
# 
# #### Explanation
#
# Think of our tree model as the complex model that we wish to explain with a simple, linear model. The above formula is saying that we can take a single data point with the three features, and the complex model makes a prediction. We can divide up that prediction among the three features, based on how important those features are to the complex model's prediction, and also based on whether the feature values push the prediction in the positive or negative direction.
#
# This is related to the ideas of coalition game theory. Imagine a team of basketball players scores 100 points in a game. We are trying to attribute part of the final score to each member of the team, based on their contributions, or "importance."
#
# When the contributions of each feature are added up to equal the complex model's prediction, this linear combination of contributions is the simple linear model that is being used to explain the complex model.
# #### Example
#
# Let's say that we've trained a complex model on 3 features. If it's given no inputs to make a prediction, then its prediction would be the equal weighted average of all its training samples.
# Let's say that equal weighted average of the training labels is **100**. In other words, if a model is given no features and asked to make a prediction, it would predict 100, which is the expected value based on the training labels.
#
# Now, let's say we give the complex model a single sample observation, with all three features, and the complex model gives a prediction of **200**.
#
# The additive feature attribution model may assign feature importances to the three features like this:
# feature 0: +50
# feature 1: +90
# feature 2: -40
#
# So this is saying that feature 0 pushed the complex model's prediction up by 50, feature 1 pushed the complex model's prediction up by 90, and feature 2 pushed the model's prediction down by 40. The end result was to go from the expected value of 100 to the prediction of 200.
#
# The shapley values that we just calculated are these values that push the model's prediction from the average of the training labels to the model's final prediction. When we add up the shapley values for all the features, they add up to the model's prediction.
# ## Compare this implementation with shap library
# #### test 1
x = np.array([0,0,0])
shap_values = shap_tree_model_explainer(model,x)
expected_value = np.mean(y)
print(f"my shap function: {shap_values}")
print(f"shap library: {shap.TreeExplainer(model).shap_values(x)}")
print(f"expected value (average of labels in y) {expected_value}")
print(f"sum of shapley values for all features: {np.sum(shap_values)}")
print(f"model prediction {model.predict(x.reshape(1,-1))}")
# #### Quiz
#
# How do you interpret the shapley values of each feature when features 0,1 and 2 are all 0?
# #### Answer
#
#
# #### test 2
x = np.array([1,0,0])
shap_values = shap_tree_model_explainer(model,x)
print(f"my shap function: {shap_values}")
print(f"shap library: {shap.TreeExplainer(model).shap_values(x)}")
print(f"expected value (average of labels in y) {expected_value}")
print(f"sum of shapley values for all features: {np.sum(shap_values)}")
print(f"model prediction {model.predict(x.reshape(1,-1))}")
# #### Quiz
#
# How do you interpret the shapley values of each feature when feature 0 is 1 and the other features are 0?
# #### Answer
#
#
# #### test 3
x = np.array([1,1,0])
shap_values = shap_tree_model_explainer(model,x)
print(f"my shap function: {shap_values}")
print(f"shap library: {shap.TreeExplainer(model).shap_values(x)}")
print(f"expected value (average of labels in y) {expected_value}")
print(f"sum of shapley values for all features: {np.sum(shap_values)}")
print(f"model prediction {model.predict(x.reshape(1,-1))}")
# #### Quiz
#
# How do we interpret the shapley values when feature 0 and 1 are both 1?
# #### Answer
#
#
# ## Solution
#
# [solution notebook](tree_shap_solution.ipynb)
|
quiz/m7/m7l6/tree_shap.ipynb
|