code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#
# MNIST
#
# @author becxer
# @email <EMAIL>
# @reference https://github.com/sjchoi86/Tensorflow-101
#
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# %matplotlib inline
print ("packages are loaded")
# +
# Get MNIST data set from tensorflow
from tensorflow.examples.tutorials.mnist import input_data
mnist_set = input_data.read_data_sets('images/MNIST/', one_hot = True)
# One hot coding is ...
# 0 = (1,0,0,0,0,0,0,0,0,0)
# 1 = (0,1,0,0,0,0,0,0,0,0)
# 2 = (0,0,1,0,0,0,0,0,0,0)
# 3 = (0,0,0,1,0,0,0,0,0,0)
# -
# Size of MNIST data set
print ("type(mnist_set) : %s" % (type(mnist_set)))
print ("len(mnist_set.train) : %s" % (mnist_set.train.num_examples))
print ("len(mnist_set.test) : %s" % (mnist_set.test.num_examples))
# +
# Shape of MNIST data set
train_img = mnist_set.train.images
train_label = mnist_set.train.labels
test_img = mnist_set.test.images
test_label = mnist_set.test.labels
print ("type(train_img) : %s" % type(train_img) )
print ("train_img.shape : %s" % str(train_img.shape))
print ("train_label.shape : %s" % str(train_label.shape))
# +
# Plot MNIST image
rand_idx = np.random.randint(train_img.shape[0], size = 1)
rand_img = np.reshape(train_img[rand_idx,:],(28,28))
rand_label = np.argmax(train_label[rand_idx,:])
plt.matshow(rand_img, cmap=plt.get_cmap('gray'))
plt.title("label : " + str(rand_label))
# +
# Convert Image rows to array image
import numpy
def scale_to_unit_interval(ndar, eps=1e-8):
""" Scales all values in the ndarray ndar to be between 0 and 1 """
ndar = ndar.copy()
ndar -= ndar.min()
ndar *= 1.0 / (ndar.max() + eps)
return ndar
def tile_raster_images(X, img_shape, tile_shape, tile_spacing=(0, 0),
scale_rows_to_unit_interval=True,
output_pixel_vals=True):
"""
Transform an array with one flattened image per row, into an array in
which images are reshaped and layed out like tiles on a floor.
This function is useful for visualizing datasets whose rows are images,
and also columns of matrices for transforming those rows
(such as the first layer of a neural net).
:type X: a 2-D ndarray or a tuple of 4 channels, elements of which can
be 2-D ndarrays or None;
:param X: a 2-D array in which every row is a flattened image.
:type img_shape: tuple; (height, width)
:param img_shape: the original shape of each image
:type tile_shape: tuple; (rows, cols)
:param tile_shape: the number of images to tile (rows, cols)
:param output_pixel_vals: if output should be pixel values (i.e. int8
values) or floats
:param scale_rows_to_unit_interval: if the values need to be scaled before
being plotted to [0,1] or not
:returns: array suitable for viewing as an image.
(See:`Image.fromarray`.)
:rtype: a 2-d array with same dtype as X.
"""
assert len(img_shape) == 2
assert len(tile_shape) == 2
assert len(tile_spacing) == 2
# The expression below can be re-written in a more C style as
# follows :
#
# out_shape = [0,0]
# out_shape[0] = (img_shape[0]+tile_spacing[0])*tile_shape[0] -
# tile_spacing[0]
# out_shape[1] = (img_shape[1]+tile_spacing[1])*tile_shape[1] -
# tile_spacing[1]
out_shape = [
(ishp + tsp) * tshp - tsp
for ishp, tshp, tsp in zip(img_shape, tile_shape, tile_spacing)
]
if isinstance(X, tuple):
assert len(X) == 4
# Create an output numpy ndarray to store the image
if output_pixel_vals:
out_array = numpy.zeros((out_shape[0], out_shape[1], 4),
dtype='uint8')
else:
out_array = numpy.zeros((out_shape[0], out_shape[1], 4),
dtype=X.dtype)
#colors default to 0, alpha defaults to 1 (opaque)
if output_pixel_vals:
channel_defaults = [0, 0, 0, 255]
else:
channel_defaults = [0., 0., 0., 1.]
for i in range(4):
if X[i] is None:
# if channel is None, fill it with zeros of the correct
# dtype
dt = out_array.dtype
if output_pixel_vals:
dt = 'uint8'
out_array[:, :, i] = numpy.zeros(
out_shape,
dtype=dt
) + channel_defaults[i]
else:
# use a recurrent call to compute the channel and store it
# in the output
out_array[:, :, i] = tile_raster_images(
X[i], img_shape, tile_shape, tile_spacing,
scale_rows_to_unit_interval, output_pixel_vals)
return out_array
else:
# if we are dealing with only one channel
H, W = img_shape
Hs, Ws = tile_spacing
# generate a matrix to store the output
dt = X.dtype
if output_pixel_vals:
dt = 'uint8'
out_array = numpy.zeros(out_shape, dtype=dt)
for tile_row in range(tile_shape[0]):
for tile_col in range(tile_shape[1]):
if tile_row * tile_shape[1] + tile_col < X.shape[0]:
this_x = X[tile_row * tile_shape[1] + tile_col]
if scale_rows_to_unit_interval:
# if we should scale values to be between 0 and 1
# do this by calling the `scale_to_unit_interval`
# function
this_img = scale_to_unit_interval(
this_x.reshape(img_shape))
else:
this_img = this_x.reshape(img_shape)
# add the slice to the corresponding position in the
# output array
c = 1
if output_pixel_vals:
c = 255
out_array[
tile_row * (H + Hs): tile_row * (H + Hs) + H,
tile_col * (W + Ws): tile_col * (W + Ws) + W
] = this_img * c
return out_array
# -
import PIL as pil
array_img = tile_raster_images( X = train_img, img_shape=(28, 28), tile_shape=(20, 20),tile_spacing=(1, 1))
pil.Image.fromarray(array_img)
|
Jupyter/001_MNIST.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Heroes Of Pymoli Data Analysis
# * Of the 1163 active players, the vast majority are male (84%). There also exists, a smaller, but notable proportion of female players (14%).
#
# * Our peak age demographic falls between 20-24 (44.8%) with secondary groups falling between 15-19 (18.60%) and 25-29 (13.4%).
# -----
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import pandas as pd
import pathlib
# File to Load (Remember to Change These)
file_to_load = pathlib.Path("Resources/purchase_data.csv")
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
purchase_data
# -
# ## Player Count
# * Display the total number of players
#
len(pd.unique(purchase_data['SN']))
# ## Purchasing Analysis (Total)
# * Run basic calculations to obtain number of unique items, average price, etc.
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
#
number_of_unique_items = len(pd.unique(purchase_data["Item ID"]))
number_of_unique_items
average_price = round(purchase_data["Price"].mean(), 2)
average_price
number_of_purchases = purchase_data["Purchase ID"].count()
number_of_purchases
total_revenue = purchase_data["Price"].sum()
total_revenue
purchase_analysis_df = pd.DataFrame({"Number of Unique Items" : number_of_unique_items,
"Average Price": (f'${average_price}'),
"Number of Purchases": number_of_purchases,
"Total Revenue": (f'${total_revenue}')
}, index = [0])
purchase_analysis_df
# ## Gender Demographics
# * Percentage and Count of Male Players
#
#
# * Percentage and Count of Female Players
#
#
# * Percentage and Count of Other / Non-Disclosed
#
#
#
gender_d_df = pd.DataFrame([{'Count': None,'Percentage': None}], index = ['Male', 'Female', 'Other/Non-Disclosed'])
gender_d_df
gender_series = purchase_data["Gender"].value_counts()
gender_series
gender_total = gender_series.sum()
gender_total
gender_d_df['Count']['Male'] = gender_series["Male"]
gender_d_df['Percentage']['Male'] = gender_series["Male"] / gender_total
gender_d_df['Count']['Female'] = gender_series["Female"]
gender_d_df['Percentage']['Female'] = gender_series["Female"] / gender_total
gender_d_df['Count']['Other/Non-Disclosed'] = gender_series["Other / Non-Disclosed"]
gender_d_df['Percentage']['Other/Non-Disclosed'] = gender_series["Other / Non-Disclosed"] / gender_total
formatted_d_gender_df = gender_d_df.style.format({'Percentage': '{:.2%}' })
formatted_d_gender_df
#
# ## Purchasing Analysis (Gender)
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
#
#
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
by_gender_group = purchase_data.groupby("Gender")
purchase_count = by_gender_group.count()["Purchase ID"]
purchase_count.name = "Purchase Count"
purchase_count
# -
average_purchase_price = by_gender_group["Price"].mean()
average_purchase_price.name = "Average Purchase Price"
average_purchase_price
by_person_group = purchase_data.groupby(["SN", "Gender"])
average_purchase_total = by_person_group.sum()
average_purchase_total = average_purchase_total["Price"].groupby(["Gender"]).mean()
average_purchase_total.name = "Average of Total of Purchases Per Person"
average_purchase_total
# +
total_purchase_value = by_gender_group["Price"].sum()
total_purchase_value.name = "Total Purchase Value"
total_purchase_value
# -
by_gender_df = pd.concat([purchase_count, average_purchase_price, average_purchase_total, total_purchase_value], axis = 1)
by_gender_df
format_dict = {'Average Purchase Price':'${0:,.2f}', "Average of Total of Purchases Per Person":'${0:,.2f}', "Total Purchase Value": '${0:,.2f}' }
formatted_by_gender_df = by_gender_df.style.format(format_dict)
formatted_by_gender_df
# ## Age Demographics
# * Establish bins for ages
#
#
# * Categorize the existing players using the age bins. Hint: use pd.cut()
#
#
# * Calculate the numbers and percentages by age group
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: round the percentage column to two decimal points
#
#
# * Display Age Demographics Table
#
purchase_data.describe()
age_df = purchase_data
age_bins = [0, 4, 8, 12, 16, 20, 24, 28, 32, 36, 40, 44, 48]
bin_names = ["0-4","5-8",'9-12','13-16','17-20','21-24','25-28','29-32','33-36','37-40','41-44','45-48']
age_df["Age Group"] = pd.DataFrame(pd.cut(purchase_data['Age'], age_bins, labels = bin_names))
age_df
count_of_unique_users_by_age_group = age_df.groupby("Age Group")["SN"].value_counts().groupby("Age Group").count()
count_of_unique_users_by_age_group.name = "Total Count"
count_of_unique_users_by_age_group
sum_of_age_groups = count_of_unique_users_by_age_group.sum()
percent_of_age_groups = count_of_unique_users_by_age_group / sum_of_age_groups
percent_of_age_groups.name = "Percentage"
percent_of_age_groups
age_demographics_df = pd.concat([count_of_unique_users_by_age_group, percent_of_age_groups], axis = 1)
format_dict = {'Percentage':'{:.2%}'}
age_demographics_formatted_df = age_demographics_df.style.format(format_dict)
age_demographics_formatted_df
# ## Purchasing Analysis (Age)
# * Bin the purchase_data data frame by age
#
#
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
age_df = purchase_data
age_bins = [0, 4, 8, 12, 16, 20, 24, 28, 32, 36, 40, 44, 48]
bin_names = ["0-4","5-8",'9-12','13-16','17-20','21-24','25-28','29-32','33-36','37-40','41-44','45-48']
age_df["Age Group"] = pd.DataFrame(pd.cut(purchase_data['Age'], age_bins, labels = bin_names))
age_df
agegroup_groupby = age_df.groupby("Age Group")
num_purchases = agegroup_groupby["Purchase ID"].count()
num_purchases.name = "Purchase Count"
num_purchases
avg_purchase_price = agegroup_groupby["Price"].sum() / num_purchases
avg_purchase_price.name = "Average Purchase Price"
avg_purchase_price
total_purchase_value = agegroup_groupby["Price"].sum()
total_purchase_value.name = 'Total Purchase Value'
total_purchase_value
avg_total_purchase_per_person = total_purchase_value / count_of_unique_users_by_age_group
avg_total_purchase_per_person.name = "Avg Total Purchase per Person"
avg_total_purchase_per_person
age_purchase_analysis_df = pd.concat([num_purchases,
avg_purchase_price,
total_purchase_value,
avg_total_purchase_per_person],
axis = 1)
format_dict = {'Average Purchase Price':'${0:,.2f}',
'Total Purchase Value': '${0:,.2f}',
'Avg Total Purchase per Person': '${0:,.2f}',
}
age_purchase_analysis_formatted_df = age_purchase_analysis_df.style.format(format_dict)
age_purchase_analysis_formatted_df
# ## Top Spenders
# * Run basic calculations to obtain the results in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the total purchase value column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
unique_users_df = purchase_data.
total_purchase_value = purchase_data.groupby("SN")["Price"].sum()
total_purchase_value
purchase_counts = purchase_data.groupby("SN")["SN"].count()
purchase_counts
average_purchase_price = total_purchase_value / purchase_counts
average_purchase_price
top_spenders_df = pd.concat([total_purchase_value,
purchase_counts,
average_purchase_price ],
axis = 1,
keys = ["Total Purchase Value",
'Purchase Count',
"Average Purchase Price",
]
)
top_spenders_df = top_spenders_df.sort_values("Total Purchase Value", ascending = False)
top_spenders_df
format_dict = {"Total Purchase Value": '${0:,.2f}',
"Average Purchase Price": '${0:,.2f}',
}
top_spenders_formatted_df = top_spenders_df.head().style.format(format_dict)
top_spenders_formatted_df
# ## Most Popular Items
# * Retrieve the Item ID, Item Name, and Item Price columns
#
#
# * Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the purchase count column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
items_df = purchase_data[["Item ID", "Item Name", "Price"]]
items_df
purchases_per_item = items_df.groupby(["Item ID", "Item Name"])["Item Name"].count()
purchases_per_item
price_per_item = test_df.groupby(["Item ID", "Item Name"])["Price"].mean()
price_per_item
total_purchase_value_per_item = test_df.groupby(["Item ID", "Item Name"])["Price"].sum()
total_purchase_value_per_item
final_items_df = pd.concat([purchases_per_item,
price_per_item,
total_purchase_value_per_item
],
axis = 1,
keys = ["Purchase Count", "Item Price", "Total Purchase Value"]
)
final_items_df
format_dict = {"Item Price": '${0:,.2f}',
"Total Purchase Value": '${0:,.2f}' }
most_popular_items_df_formatted = final_items_df.sort_values("Purchase Count", ascending = False).head().style.format(format_dict)
most_popular_items_df_formatted
# ## Most Profitable Items
# * Sort the above table by total purchase value in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the data frame
#
#
format_dict = {"Item Price": '${0:,.2f}',
"Total Purchase Value": '${0:,.2f}' }
most_profitable_items_df_formatted = final_items_df.sort_values("Total Purchase Value", ascending = False).head().style.format(format_dict)
most_profitable_items_df_formatted
|
HeroesOfPymoli/.ipynb_checkpoints/HeroesOfPymoli_hw-checkpoint.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.0
# language: julia
# name: julia-1.7
# ---
# # MATH50003 Numerical Analysis: Problem Sheet 10
#
# This problem sheet explores orthogonal polynomial roots, interpolatory quadrature and Gaussian quadrature.
#
# All questions are meant to be completed without using a computer.
# Problems are denoted A/B/C to indicate their difficulty.
#
# ## 1. Orthogonal Polynomial Roots
#
# **Problem 1.1 (C)** Compute the roots of $P_3(x)$, orthogonal with respect
# to $w(x) = 1$ on $[-1,1]$, by computing the eigenvalues of a $3 × 3$ truncation
# of the Jacobi matrix.
#
# **Problem 1.2 (B)** Give an explicit diagonalisation of
# $$
# X_n = \begin{bmatrix} 0 & 1/2 \\
# 1/2 & 0 & ⋱ \\
# & ⋱ & ⋱ & 1/2 \\
# && 1/2 & 0
# \end{bmatrix} ∈ ℝ^{n × n}
# $$
# for all $n$ by relating it to the Jacobi matrix for $U_n(x)$.
#
#
# **Problem 1.3 (A)** Give an explicit solution to heat on a graph
# $$
# \begin{align*}
# 𝐮(0) &= 𝐮_0 ∈ ℝ^n \\
# 𝐮_t &= Δ 𝐮
# \end{align*}
# $$
# where
# $$
# Δ := \begin{bmatrix} -2 & 1 \\
# 1 & -2 & ⋱ \\
# & 1 & ⋱ & 1 \\
# && ⋱ & -2 & 1 \\
# &&& 1 & -2
# \end{bmatrix} ∈ ℝ^{n \times n}
# $$
# (which corresponds to Dirichlet conditions.) Hint: use Problem 1.2 to diagonalise the problem.
#
#
#
# ## 2. Interpolatory quadrature
#
#
# **Problem 2.1 (C)** Compute the interpolatory quadrature rule for
# $w(x) = \sqrt{1-x^2}$ with the points $[-1,1/2,1]$.
#
#
# **Problem 2.2 (C)** Compute the 2-point
# interpolatory quadrature rule associated with roots of orthogonal polynomials for the weights $\sqrt{1-x^2}$, $1$,
# and $1-x$ on $[-1,1]$ by integrating the Lagrange bases.
#
#
#
# ## 3. Gaussian quadrature
#
#
# **Problem 3.1 (C)** Compute the 2-point and 3-point Gaussian quadrature rules associated with $w(x) = 1$ on $[-1,1]$.
#
# **Problem 3.2 (A)** Show for $w(x) = 1/\sqrt{1-x^2}$ that the Gaussian quadrature rule is
# $$
# {π \over n} \sum_{j=1}^n f(x_j)
# $$
# where $x_j = (j-1/2)π/n$ for all $n$.
#
# **Problem 3.3 (B)** Solve Problem 1.2 from PS8 using **Lemma (discrete orthogonality)** with
# $w(x) = 1/\sqrt{1-x^2}$ on $[-1,1]$.
|
sheets/week10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from pathlib import Path
import math
import pandas as pd
import requests
from bs4 import BeautifulSoup, SoupStrainer
from selenium.webdriver import Chrome
from selenium.webdriver.common.keys import Keys
import time
# -
DATA_PATH = Path.cwd().parent.parent / "data" / "raw"
recipes_urls = pd.read_csv(DATA_PATH / "urls.csv")
recipes_urls = recipes_urls['url'].to_list()
# ## Define function to load data
#
# need to make https://www.veganrecipeclub.org.uk/recipes/15-minute-flourless-choc-chip-banana-muffins/ work
# ### Find the different section of the web page
#Title
def get_title(soup):
return soup.find_all("h1")[0].get_text(strip=True)
#section name
def get_sections(soup):
elements = dict()
sections = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
for i, section in enumerate(sections):
elements[section.text] = i
return elements
# +
#Preparation time
def get_preparation_time(soup, elements):
time = 0
html_element = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
try:
if(html_element[elements['Preparation time']].nextSibling.nextSibling.text != None):
time = html_element[elements['Preparation time']].nextSibling.nextSibling.text
except:
return 0
return time
#Cooking time
def get_cooking_time(soup, elements):
time = 0
html_element = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
try:
if(html_element[elements['Cooking time']].nextSibling.nextSibling.text != None):
time = html_element[elements['Cooking time']].nextSibling.nextSibling.text
except:
return 0
return time
#Preparation time
def get_total_time(soup, elements):
time = 0
html_element = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
try:
if(html_element[elements['Total time']].nextSibling.nextSibling.text != None):
time = html_element[elements['Total time']].nextSibling.nextSibling.text
except:
return 0
return time
# -
#Style of cooking
def get_cooking_style(soup):
cooking_styles = ''
styles = soup.find_all("p", {"class": "intList"})
if len(styles) != 0:
for style in styles:
cooking_styles += style.get_text(strip=True) + ' '
return cooking_styles
#Type of recipe
def get_recipe_type(soup, elements):
html_element = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
recipe_types = ''
try:
types = html_element[elements['Meal']].nextSibling.nextSibling.find_all("li")
for type_ in types:
recipe_types += type_[0] + ' '
except:
return ''
return recipe_types
#Number of servings
def get_number_servings(soup, elements):
html_element = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
try:
return html_element[elements['Servings']].nextSibling.nextSibling.get_text()
except:
return 'unknown'
#Ingredients
def get_ingredients(soup, elements):
html_element = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
ingredients = []
try:
HTML_ingredients = html_element[elements['Ingredients']].nextSibling.nextSibling
for item in HTML_ingredients.find_all("li"):
ingredients.append(item.get_text())
if(len(HTML_ingredients.find_all("li")) == 0):
#Ingredients
HTML_ingredients = HTML_ingredients[elements['Ingredients']].parent.find("div", {"class": "recipe-list"}).p
for ingredient in HTML_ingredients.get_text().split("\n"):
ingredients.append(ingredient)
except:
return []
return ingredients
#Instructions
def get_instructions(soup, elements):
html_element = soup.find_all("p", {"style": "text-align: center; font-weight: bold;"})
instructions = []
try:
HTML_instructions = html_element[elements['Instructions']].nextSibling.nextSibling
for item in HTML_instructions.find_all("li"):
instructions.append(item.get_text())
except:
return []
return instructions
# ## Scrape every urls
recipes_urls[:5]
# +
# %%time
rows = []
columns = ['title', 'preparation_time', 'cooking_time', 'total_time', 'style', 'type', 'servings', 'ingredients', 'instructions']
for i, url in enumerate(recipes_urls):
time.sleep(5)
row = []
try:
page = requests.get(url)
soup = BeautifulSoup(page.content, 'html.parser')
sections = get_sections(soup)
row.append(get_title(soup))
row.append(get_preparation_time(soup, sections))
row.append(get_cooking_time(soup, sections))
row.append(get_total_time(soup, sections))
row.append(get_cooking_style(soup))
row.append(get_recipe_type(soup, sections))
row.append(get_number_servings(soup, sections))
row.append(get_ingredients(soup, sections))
row.append(get_instructions(soup, sections))
rows.append(row)
except:
print('link:', url, 'had an error')
print("Progress:", round(100*i/len(recipes_urls),2),"%", end='\r')
df = pd.DataFrame(rows, columns = columns)
# -
df
# ## Transform columns of dataframe
df.info()
# +
# for col in ['preparation_time', 'cooking_time', 'total_time']:
# df[col] = df[col].fillna(0)
# df[col] = df[col].astype(str)
# df[col] = df[col].str.extract('(\d+)', expand=False)
# df[col] = df[col].astype(int)
# -
df
df.to_csv(DATA_PATH / "recipes.csv")
|
notebooks/data_scrapping/Load_recipe_content.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ML Project Template
# # Frame the Problem and Look at the Big Picture
# ## Define the objective in business terms.
#
# ## How will your solution be used.
#
# ## What are the current solutions/workarounds (if any)?
#
# ## How should you frame this problem?
# * Supervised/unsupervised/semisupervised/reinforcement learning
# * Online / offline
#
# ## How should performance be measured?
#
# ## Is the performance measure aligned with the business objective?
#
# ## What would be the minimum performance needed to reach the business objective?
#
# ## What are comparable problems? Can you reuse experience or tools?
#
# ## Is human expertise available?
#
# ## How would you solve the problem manually?
#
# ## List the assumptions you (or others) have made so far.
#
# ## Verify the assumptions if possible.
#
# # Get the data
# Note: Automate as much as possible so you can easily get fresh data.
# ## List the data you need and how you need
#
# ## Find and document where you can get that data
#
# ## Check how much space it will take
#
# ## Check legal obligations, and get authorization if necessary.
#
# ## Get access authorizations
#
# ## Create a workspace (with enought storage space)
#
# ## Get the data.
#
# ## Convert the data to a format you can easily manipulate (without changing the data itself).
#
# ## Ensure sensitive information is deleted or protected (i.e. anonymized).
#
# ## Check the size and type of data (time series, sample, geographical, etc.)
#
# ## Sample a test, put it aside, and never look at it.
#
# # Explore the data
# Note: Try to get insights from a field expert for these steps
# ## Create a copy of the data for exploration (sampling it down to a manageable size if necessary)
#
# ## Create a Jupyter Notebook to keep a record of your data exploration.
#
# ## Study each attribute and its characteristics.
# * Name
# * Type (categorical, int/float,bounded/unbounded, text, structured, etc.)
# * % of missing
# * Noisiness and types of noise (stochastic, outliers, rounding errors, etc.)
# * Possibly useful for the task?
# * Type of distribution (Gaussian, uniform, logarithmic, etc.)
# ## For supervised learning, identify the target attributes.
#
# ## Visualize the data.
#
# ## Study the correlations between attributes.
#
# ## Study how you would solve the problem manually.
#
# ## Identify the promising transformations you may want to apply.
#
# ## Identify the data that would be useful.
#
# ## Document what you have learned.
#
# # Prepare the data
# Notes:
# * Work on copies of the data (keep the original dataset intact)
# * Write functions for all data transformations you apply, for five reasons:
# * So you can easily prepare the data the next time you get a fresh dataset
# * So you can apply these transformations in future projects
# * To clean and prepare a test set
# * To clean and prepare new instances once your solution is live
# * To make it easy to treat your preparation choices as hyperparameters
#
# ## Data Cleaning:
# * Fix or remove outliers (optional).
# * Fill in missing values (e.g. with zero, mean, median, ...) or drop their rows/columns.
# ## Feature selection (optional):
# * Drop the attributes that provide no useful information for the task.
# ## Feature engineering, where appropriate
# * Discretize continuous features
# * Decompose features (e.g. categorical, date/time, etc.)
# * Add promising transformations of features (e.g. log(x), sqrt(x), x^2, etc.)
# * Aggregate features into promising new features
#
# ## Short-listing Promising Models
# Notes:
# * If the data is huge, you may want to sample smaller training sets so you can train many different models in a reasonable time (be aware that this penalizes complex models such as large neural nets and Random Forests).
# * Once again, try to automate these steps as much as possible
# 1. Train many quick and dirty models from different categories (e.g. linear, naive Bayes, SVM, Random Forests, neural net, etc.) using standard parameters.
# 2. Measure and compare their performance.
# * For eah model, use N-fold cross validation and compute mean and standard deviation of the performance measure of the N folds.
# 3. Analyze the most significant variables for each algorithm.
# 4. Analyze the types fo errors the models make.
# * What data would a human have used to avoid the errors?
# 5. Have a quick round of feature selection and engineering.
# 6. Have one or two more quick iterations of the five previous steps.
# 7. Short-list the top three to five most promising models, preferring the models that make different types of errors.
#
# ## Fine-Tune the system.
# Notes:
# * You will want to use as much data as possible for this step, especially as you move toward the end of fine-tuning.
# * As always automate what you can.
# 1. Fine-tune the hyperparameters using cross-validation.
# * Treat your data transformation choices as hyperparameters, especially when you are not sure about them (e.g. should I replace missing values with zero or with the median value? Or just drop rows?).
# * Unless htere are very few hyperparameter values to explores, prefer random search over grid search. If training is very long, you may prefer a Bayesian optimization approach (e.g. using Gaussian process priors: https://arxiv.org/pdf/1206.2944.pdf)
# 2. Try ensemble methods. Combining your best models will often perform better than running them individually.
# 3. Once you are confident about your final model, measure its performance on the tes set to estimate the generalization error.
# !
# Don't tweak your model after measuring the generalization error: You would just start overfitting the test set.
# # Present your solution
# ## Document what you have done.
# ## Create a nice presentation.
# * Make sure you highlight the big picture first.
# ## Explain why your solution achieves the business objective.
# ## Don't forget to present interesting points you noticed along the way.
# * Describe what worked and what did not.
# * List your assumptions and your systems' limitations.
# ## Ensure your key findings are communicated through beautiful visualizations or easy to remember statements (e.g. "the median income is the number-one predictor of housing prices").
# # Launch
# ## Get your solution ready for production (plug into production data inputs, write unit tests, etc.)
# ## Write monitoring code to check your systems' live performance at regular intervals and trigger alerts when it drops.
# * Beware of slow degradation too: models tend to "rot" as data evolves.
# * Measuring performance may require a human pipeline (e.g. via crowdsourcing service).
# * Also monitor your inputs' quality (e.g. a malfunctioning sensor sending random values, or another team's output becoming stale). This is particularly important for online learning systems.
# ## Retrain your models on a regular basis on fresh data (automate as much as possible).
#
#
|
machine-learning-template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false"
# # Publication queries
#
# This notebook contains a collection of common publication queries for [Dimensions on Google BigQuery](https://docs.dimensions.ai/bigquery/).
#
# For more background, see also the [publications data model](https://docs.dimensions.ai/bigquery/datasource-publications.html).
# + [markdown] Collapsed="false"
# ## Prerequisites
#
# This notebook assumes that you have [verifed your connection](https://digital-science.github.io/dimensions-gbq-lab/cookbooks/1-Verifying-your-connection.html) to Dimensions on Google BigQuery and have basic familiarity with the [Google Cloud Platform](https://docs.dimensions.ai/bigquery/gcp-setup.html) concepts.
#
# The following code will load the Python BigQuery library and authenticate you as a valid user.
# + Collapsed="false"
# !pip install google-cloud-bigquery -U --quiet
# %load_ext google.cloud.bigquery
import sys
print("==\nAuthenticating...")
if 'google.colab' in sys.modules:
from google.colab import auth
auth.authenticate_user()
print('..done (method: Colab)')
else:
from google.cloud import bigquery
print('..done (method: local credentials)')
#
# PLEASE UPDATE USING YOUR CLOUD PROJECT ID (= the 'billing' account)
#
MY_PROJECT_ID = "ds-data-solutions-gbq"
print("==\nTesting connection..")
client = bigquery.Client(project=MY_PROJECT_ID)
test = client.query("""
SELECT COUNT(*) as pubs
from `dimensions-ai.data_analytics.publications`
""")
rows = [x for x in test.result()]
print("...success!")
print("Total publications in Dimensions: ", rows[0]['pubs'])
# + [markdown] Collapsed="false" toc-hr-collapsed=true toc-nb-collapsed=true
# ## 1. Top publications by Altmetric score and research organization
#
#
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
-- Top 5 pubs by Altmetric Score for GRID ID grid.4991.5 in the year 2020
SELECT
id,
title.preferred as title,
ARRAY_LENGTH(authors) as authors,
altmetrics.score as altmetrics_score
FROM
`dimensions-ai.data_analytics.publications`
WHERE
year = 2020 AND 'grid.4991.5' in UNNEST(research_orgs)
ORDER BY
altmetrics.score desc
LIMIT 5
# + [markdown] Collapsed="false" toc-hr-collapsed=true toc-nb-collapsed=true
# ## 1. Working with Publications dates
#
# Each publication has various dates available.
#
# * `date`, `year`, `date_normal`, `date_online`, `date_print` refer to the publication object. See the [documentation](https://docs.dimensions.ai/bigquery/datasource-publications.html) to find out more about their meaning.
# * `date_imported_gbq` refers to when this record was last added to GBQ - this date can be handy if you want to synchronize an external data source to GBQ.
# * `date_inserted`: this refers to when this records was originally added to Dimensions (if the records gets adjusted later, it doesn't change).
# + [markdown] Collapsed="false"
# ### Comparing date fields
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT doi,
date,
date_normal,
year,
date_online,
date_print,
date_imported_gbq,
date_inserted
FROM `dimensions-ai.data_analytics.publications`
WHERE year = 2010
AND journal.id = "jour.1115214"
ORDER BY citations_count DESC
LIMIT 10
# + [markdown] Collapsed="false"
# ### Number of publications added to Dimensions by month
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
DATETIME_TRUNC(DATETIME(date_inserted), MONTH) as date,
COUNT(id) as countDim
FROM
`dimensions-ai.data_analytics.publications`
GROUP BY date
ORDER BY date DESC
LIMIT 5
# + [markdown] Collapsed="false"
# ## 2. Working with NESTED fields
#
# UNNEST are implicit 'cross-join' queries, hence only records that have some value in the nested column are represented
#
# For example, the query below return less publications that then ones available, because only the ones with `research_org_country_names` are included (= cross join)
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(DISTINCT p.id) AS tot_articles
FROM
`dimensions-ai.data_analytics.publications` p,
UNNEST(research_org_country_names) AS research_org_country_names
WHERE
year = 2000
# + [markdown] Collapsed="false"
# As a test, we can run the query without the UNNEST clause
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(DISTINCT p.id) AS tot_articles
FROM
`dimensions-ai.data_analytics.publications` p
WHERE
year = 2000
# + [markdown] Collapsed="false"
# So how can we get all the records out?
#
# If you want to get all records, then **LEFT JOIN is the way to go** in this case
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(DISTINCT p.id) AS tot_articles
FROM
`dimensions-ai.data_analytics.publications` p
LEFT JOIN
UNNEST(research_org_country_names) AS research_org_country_names
WHERE
year = 2000
# + [markdown] Collapsed="false"
# ## 3. Generate a list of publication authors by flattening/concatenating nested data
#
# IE Flattening an array of objects into a string
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT p.id,
ARRAY_TO_STRING(
(
SELECT ARRAY
(
select CONCAT(first_name, " ", last_name)
from UNNEST(p.authors)) ), '; ') AS authors_list
FROM `dimensions-ai.data_analytics.publications` p
WHERE p.id = 'pub.1132070778'
# + [markdown] Collapsed="false"
# ## 4. Generate a list of publication categories by flattening/concatenating nested data
#
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT p.id,
ARRAY_TO_STRING(
(
SELECT ARRAY
(
SELECT name
FROM UNNEST(p.category_for.first_level.FULL)) ), '; ') AS categories_list
FROM `dimensions-ai.data_analytics.publications` p
WHERE p.id = 'pub.1132070778'
# + [markdown] Collapsed="false"
# ## 5. Number of publications per SDG category
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(DISTINCT p.id) AS tot,
sdg.name
FROM `dimensions-ai.data_analytics.publications` p,
UNNEST(category_sdg.full) sdg
GROUP BY sdg.name
LIMIT
5
# + [markdown] Collapsed="false"
# ## 6. Publications count per FoR category, total and percentage against total
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
cat.name,
COUNT(DISTINCT p.id) AS pubs_global,
ROUND ((COUNT(DISTINCT p.id) * 100 /(
SELECT
COUNT(*)
FROM
`dimensions-ai.data_analytics.publications`)), 2 ) AS pubs_global_pc
FROM
`dimensions-ai.data_analytics.publications` p,
UNNEST(category_for.first_level.full) cat
GROUP BY
cat.name
# + [markdown] Collapsed="false"
# ## 7. Finding Journals using string matching
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT COUNT(*) AS pubs,
journal.id,
journal.title,
journal.issn,
journal.eissn,
publisher.name
FROM
`dimensions-ai.data_analytics.publications`
WHERE
LOWER( journal.title ) LIKE CONCAT('%medicine%')
GROUP BY 2, 3, 4, 5, 6
ORDER BY pubs DESC
LIMIT 20
# + [markdown] Collapsed="false"
# ## 8. Finding articles matching a specific affiliation string
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT id,
aff.grid_id,
aff.raw_affiliation
FROM `dimensions-ai.data_analytics.publications`,
UNNEST(authors) auth,
UNNEST(auth.affiliations_address) AS aff
WHERE year = 2020
AND aff.grid_id = "grid.69566.3a"
AND LOWER(aff.raw_affiliation) LIKE "%school of medicine%"
# + [markdown] Collapsed="false"
# ### 8.1 Variant: get unique publication records with affiliation count
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(aff) AS matching_affiliations,
id,
title.preferred AS title
FROM
`dimensions-ai.data_analytics.publications`,
UNNEST(authors) auth,
UNNEST(auth.affiliations_address) AS aff
WHERE
year = 2020
AND aff.grid_id = "grid.69566.3a"
AND LOWER(aff.raw_affiliation) LIKE "%school of medicine%"
GROUP BY
id,
title
# + [markdown] Collapsed="false"
# ## 9. Select publications matching selected concepts
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
WITH tropical_diseases AS
(
SELECT *
FROM `dimensions-ai.data_analytics.publications` )
SELECT publisher.NAME AS publisher,
year,
count(*) AS num_pub
FROM tropical_diseases,
UNNEST(tropical_diseases.concepts) c
WHERE (
LOWER(c.concept) IN UNNEST(["buruli ulcer", "mycobacterium", "mycolactone", "bairnsdale ulcer"])
OR REGEXP_CONTAINS(title.preferred, r"(?i)/buruli ulcer|mycobacterium|mycolactone|bairnsdale ulcer/"))
AND year >= 2010
AND publisher IS NOT NULL
GROUP BY publisher, year
ORDER BY num_pub DESC,
year,
publisher LIMIT 10
# + [markdown] Collapsed="false"
# ## 10. Count of corresponding authors by publisher
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(DISTINCT id) AS tot,
publisher.name
FROM
`dimensions-ai.data_analytics.publications`,
UNNEST(authors) aff
WHERE
aff.corresponding IS TRUE
AND publisher.name IS NOT NULL
GROUP BY
publisher.name
ORDER BY
tot DESC
# + [markdown] Collapsed="false"
# ## 11. Counting new vs recurring authors, for a specific journal
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
WITH
authoryear AS (
SELECT pubs.year, author.researcher_id, COUNT(pubs.id) AS numpubs
FROM
`dimensions-ai.data_analytics.publications` AS pubs
CROSS JOIN
UNNEST(pubs.authors) AS author
WHERE
author.researcher_id IS NOT NULL
AND journal.id= "jour.1115214"
GROUP BY
author.researcher_id, pubs.year ),
authorfirst AS (
SELECT researcher_id, MIN(year) AS minyear
FROM
authoryear
GROUP BY
researcher_id ),
authorsummary AS (
SELECT ay.*,
IF
(ay.year=af.minyear,
TRUE,
FALSE) AS firstyear
FROM
authoryear ay
JOIN
authorfirst af
ON
af.researcher_id=ay.researcher_id
ORDER BY
ay.researcher_id, year ),
numauthors AS (
SELECT year, firstyear, COUNT(DISTINCT researcher_id) AS numresearchers
FROM
authorsummary
WHERE
year>2010
GROUP BY year, firstyear )
SELECT
year,
SUM(CASE
WHEN firstyear THEN numresearchers
ELSE
0
END
) AS num_first,
SUM(CASE
WHEN NOT firstyear THEN numresearchers
ELSE
0
END
) AS num_recurring
FROM numauthors
GROUP BY year
ORDER BY year
# + [markdown] Collapsed="false"
# ## 12. Funding by journal
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
WITH funding AS
(
SELECT funding.grid_id AS funders,
COUNT(id) AS pubs,
COUNT(funding.grant_id) AS grants
FROM `dimensions-ai.data_analytics.publications`,
UNNEST(funding_details) AS funding
WHERE journal.id = "jour.1113716" -- nature medicine
GROUP BY funders)
SELECT funding.*,
grid.NAME
FROM funding
JOIN `dimensions-ai.data_analytics.grid` grid
ON funding.funders = grid.id
ORDER BY pubs DESC,
grants DESC
LIMIT 10
# + [markdown] Collapsed="false"
# ## 13. Citations queries
# + [markdown] Collapsed="false"
# ### 13.1 Top N publications by citations percentile
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
WITH pubs AS (
SELECT
p.id as id,
p.title.preferred as title,
p.citations_count as citations,
FROM
`dimensions-ai.data_analytics.publications` p
WHERE year = 2020 AND "09" IN UNNEST(category_for.first_level.codes)
),
ranked_pubs AS (
SELECT
p.*,
PERCENT_RANK() OVER (ORDER BY p.citations DESC) citation_percentile
FROM
pubs p
)
SELECT * FROM ranked_pubs
WHERE citation_percentile <= 0.01
ORDER BY citation_percentile asc
# + [markdown] Collapsed="false"
# ### 13.2 Citations by journal, for a specific publisher
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
WITH publisher_pubs AS (
SELECT id FROM `dimensions-ai.data_analytics.publications`
WHERE publisher.id = "pblshr.1000340" AND type = "article"
)
SELECT
COUNT(p.id) as tot,
p.journal.title as journal
FROM `dimensions-ai.data_analytics.publications` p, UNNEST(p.reference_ids) r
WHERE
p.year = 2020 AND p.type = "article" -- restrict to articles with a published year of 2020
AND p.publisher.id <> "pblshr.1000340" -- where the publisher is not the same as the pusblisher above
AND r IN (SELECT * FROM publisher_pubs) -- the publication must reference a publishers publication
GROUP BY journal
ORDER BY tot DESC
LIMIT 10
# + [markdown] Collapsed="false"
# ### 13.3 One-degree citation network for a single publication
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
WITH level1 AS (
select "pub.1099396382" as citation_from, citations.id as citation_to, 1 as level, citations.year as citation_year
from `dimensions-ai.data_analytics.publications` p, unnest(citations) as citations
where p.id="pub.1099396382"
),
level2 AS (
select l.citation_to as citation_from, citations.id as citation_to, 2 as level, citations.year as citation_year
from `dimensions-ai.data_analytics.publications` p, unnest(citations) as citations, level1 l
where p.id = l.citation_to
)
SELECT * from level1
UNION ALL
SELECT * from level2
# + [markdown] Collapsed="false"
# ### 13.4 Incoming citations for a journal
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(DISTINCT id) AS totcount, year, type
FROM
`dimensions-ai.data_analytics.publications`
WHERE
id IN (
SELECT citing_pubs.id
FROM
`dimensions-ai.data_analytics.publications`,
UNNEST(citations) AS citing_pubs
WHERE journal.id = "jour.1115214" ) -- Nature Biotechnology
GROUP BY year, type
ORDER BY year, type
# + [markdown] Collapsed="false"
# ### 13.5 Outgoing citations to a journal
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
COUNT(DISTINCT id) AS totcount, year, type
FROM
`dimensions-ai.data_analytics.publications`
WHERE
id IN (
SELECT
DISTINCT reference_pubs
FROM
`dimensions-ai.data_analytics.publications`,
UNNEST(reference_ids) AS reference_pubs
WHERE
journal.id = "jour.1115214" ) -- Nature Biotechnology
GROUP BY year, type
ORDER BY year, type
# + [markdown] Collapsed="false"
# ## 14. Extracting complex publications records
#
# The query below combines various techniques presented in this notebook in order to extract full publication records that include both single-value metadata and unpacked lists.
#
# We use LEFT JOIN in order to ensure we obtain all records, not just the ones that have some value in the nested objects.
# + Collapsed="false"
# %%bigquery --project $MY_PROJECT_ID
SELECT
p.id,
p.title.preferred AS title,
p.doi,
p.year,
COALESCE(p.journal.title, p.proceedings_title.preferred, p.book_title.preferred, p.book_series_title.preferred) AS venue,
p.type,
p.date AS date_publication,
p.date_inserted,
p.altmetrics.score AS altmetrics_score,
p.metrics.times_cited,
grid.id AS gridid,
grid.name AS gridname,
grid.address.country AS gridcountry,
grid.address.city AS gridcity,
open_access_categories,
cat_for.name AS category_for,
FROM
`dimensions-ai.data_analytics.publications` p
LEFT JOIN
UNNEST(research_orgs) AS research_orgs_grids
LEFT JOIN
`dimensions-ai.data_analytics.grid` grid
ON
grid.id=research_orgs_grids
LEFT JOIN
UNNEST(p.open_access_categories) AS open_access_categories
LEFT JOIN
UNNEST(p.category_for.first_level.full) AS cat_for
WHERE
EXTRACT(YEAR
FROM
date_inserted) >= 2020
LIMIT 100
# + Collapsed="false"
|
archive/2-Publications-queries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install dna_features_viewer
# !pip install --upgrade forgi[all]
from Bio import Entrez
import matplotlib.pyplot as plt
from dna_features_viewer import *
Entrez.email = "<EMAIL>"
# access from NCBI RefSeq
handle = Entrez.efetch(db="nucleotide", id="NC_045512", retmode="xml")
features = Entrez.read(handle)[0]
features.keys()
seq = features['GBSeq_sequence']
len(seq)
features['GBSeq_comment']
features['GBSeq_definition']
[(f['GBFeature_key'], len(f['GBFeature_intervals'])) for f in features['GBSeq_feature-table']]
features['GBSeq_feature-table'][4]
len(seq)
# +
gfeatures = []
colors = {
"gene": "#ffd700",
"CDS": "#ffcccc",
"mat_peptide": "#cffccc",
"stem_loop": "#ccccff",
}
locations = set()
stem_loops = []
coverage = 0
elements = []
import subprocess
def rnafold(inseq, out=None, free_energy=False):
args = ['RNAfold', '-t4']
if free_energy:
args.append('-p')
try:
p = subprocess.run(args,
input=bytes(inseq, 'ascii'),
stdout=subprocess.PIPE, check=True,
stderr=subprocess.PIPE)
result = p.stdout.decode()
if out is not None:
subprocess.run(['mv', 'rna.ps', out+'.ps'], check=True)
return result
except subprocess.CalledProcessError as err:
print(err, err.output.decode(), p.stderr.decode())
for f in features['GBSeq_feature-table']:
key = f['GBFeature_key']
strand = +1
loc = f['GBFeature_location']
if loc in locations:
pprint.pprint(f)
continue
else:
locations.add(loc)
if key == 'CDS':
if len(f['GBFeature_intervals']) >= 2:
label = f['GBFeature_quals'][0]['GBQualifier_value'][:-len(f['GBFeature_intervals'])]
for i, interval in enumerate(f['GBFeature_intervals']):
gfeatures.append(GraphicFeature(
start=int(interval['GBInterval_from']),
end=int(interval['GBInterval_to']),
strand=strand,
label=label+chr(ord('a')+i),
color=colors.get(key, '#000080'),
))
continue
elif key == 'source':
continue
elif key == 'mat_peptide':
product = next(qual for qual in f['GBFeature_quals'] if qual['GBQualifier_name'] == 'product')['GBQualifier_value']
note = next(qual for qual in f['GBFeature_quals'] if qual['GBQualifier_name'] == 'note')['GBQualifier_value'].split('; ')
if len(note) >= 1:
label = note[0] if len(note[0]) < len(product) else product
else:
label = product
elif key == 'stem_loop':
f = f.copy()
f['label'] = next(qual for qual in f['GBFeature_quals'] if qual['GBQualifier_name'] == 'function')['GBQualifier_value']
stem_loops.append(f)
continue
elif 'GBFeature_quals' in f:
label = f['GBFeature_quals'][0]['GBQualifier_value'] + ' ' + key
else:
label = key
if key == 'gene' and not f['GBFeature_quals'][0]['GBQualifier_value'].startswith('ORF1a') or key == 'mat_peptide' or key.endswith('UTR'):
coverage += int(f['GBFeature_intervals'][-1]['GBInterval_to']) - int(f['GBFeature_intervals'][0]['GBInterval_from']) + 1
f = f.copy()
f['seq'] = seq[int(f['GBFeature_intervals'][0]['GBInterval_from'])-1:int(f['GBFeature_intervals'][-1]['GBInterval_to'])]
f['label'] = label
print(f)
f['qualifiers'] = dict((q['GBQualifier_name'], q.get('GBQualifier_value')) for q in f.get('GBFeature_quals', ()))
elements.append(f)
gfeatures.append(GraphicFeature(
start=int(f['GBFeature_intervals'][0]['GBInterval_from']),
end=int(f['GBFeature_intervals'][-1]['GBInterval_to']),
strand=strand,
label=label,
color=colors.get(key, '#000080'),
))
# -
len(seq) - coverage
# +
import tqdm
for f in tqdm.tqdm(elements):
lines = rnafold(f['seq'], f['label']).split('\n')
f['mfe_structures'] = lines
#f['mfe_structures'] = {
# 'brackets': [tuple(l.split(' ', maxsplit=1)) for l in lines[1:-2]],
# 'extra': [tuple(p.strip().rsplit(' ', 1)) for p in lines[-2].split('; ')],
#}
# -
elements[0]
entropy_tsv = np.genfromtxt('nextstrain_ncov_global_diversity_entropy.tsv', skip_header=1)
events_tsv = np.genfromtxt('nextstrain_ncov_global_diversity_events.tsv', skip_header=1)
# +
idx, ent = entropy_tsv.T
idx = idx.astype(int)
entropy = np.zeros(idx.max())
entropy[idx-1] = ent
idx, evt = events_tsv.T
idx = idx.astype(int)
events = np.zeros(idx.max())
events[idx-1] = evt
# +
#from dna_features_viewer import BiopythonTranslator
import numpy as np
fig, (ax1, ax2, ax3, ax4) = plt.subplots(
4, 1, figsize=(12, 6), sharex=True, gridspec_kw={"height_ratios": [4, 1, 1, 1]}
)
# PLOT THE RECORD MAP
record = GraphicRecord(sequence_length=len(seq), features=gfeatures)
record.plot(ax=ax1, with_ruler=False)
pad = 100
def interval_range(elems):
return int(elems[0]['GBFeature_intervals'][0]['GBInterval_from']), int(elems[-1]['GBFeature_intervals'][0]['GBInterval_to'])
lo1, hi1 = interval_range(stem_loops[:2])
lo2, hi2 = interval_range(stem_loops[2:])
ax1.fill_between((lo1-pad, hi1+pad), +1000, -1000, alpha=0.15)
ax1.fill_between((lo2-pad, hi2+pad), +1000, -1000, alpha=0.15)
# PLOT THE LOCAL GC CONTENT (we use 50bp windows)
window = 100
gc = lambda s: 100.0 * len([c for c in s if c in "gc"]) / window
xx = np.arange(len(seq) - window)
yy = [gc(seq[x : x + window]) for x in xx]
ax2.fill_between(xx + 25, yy, alpha=0.3)
ax2.set_ylim(bottom=0)
ax2.set_ylabel("GC(%)")
filter = np.ones(window)/window
def running_mean(x):
return np.convolve(x, filter, mode='full')
ax3.fill_between(np.arange(len(entropy)+window-1), running_mean(entropy), alpha=0.3)
ax3.set_ylim(bottom=0)
ax3.set_ylabel("entropy")
ax4.fill_between(np.arange(len(events)+window-1), running_mean(events), alpha=0.3)
ax4.set_ylim(bottom=0)
ax4.set_ylabel("events")
plt.show()
# +
import pandas as pd
rows = []
print(elements[-7])
for f in elements:
intervals = f['GBFeature_intervals']
lo, hi = int(intervals[0]['GBInterval_from']), int(intervals[-1]['GBInterval_to'])
span = hi - lo+1
rows.append((f['label'], f['GBFeature_key'], f['qualifiers'].get('locus_tag'), span, np.sum(events[lo-1:hi])/span, np.sum(entropy[lo-1:hi])/span, f['qualifiers'].get('note', '')))
pd.DataFrame(rows, columns=['label', 'type', 'locus', 'length', 'events/bp', 'entropy/bp', 'notes']).to_markdown()
# -
# !pip install tabulate
# +
from io import BytesIO
import matplotlib.image as img
def plot_stem_loops(sls, rng, n_lines=1, **plot_kwargs):
slfeatures = []
for f in sls:
slfeatures.append(GraphicFeature(
start=int(f['GBFeature_intervals'][0]['GBInterval_from']),
end=int(f['GBFeature_intervals'][-1]['GBInterval_to']),
strand=1,
label=str(f['label']),
color=colors.get(key, '#000080'),
))
rnafold(seq[rng[0]:rng[1]])
image = Image(filename='rna.ps')
factor = 100
image.resample(x_res=3*factor, y_res=factor)
blob = BytesIO(image.make_blob('png'))
record = GraphicRecord(sequence=seq, features=slfeatures)
fig, axs = record.crop(rng).plot_on_multiple_lines(plot_sequence=True, n_lines=n_lines, **plot_kwargs)
ax = fig.add_axes([.005, .5+.1*(n_lines-1), .99, 1.5])
ax.imshow(img.imread(blob), aspect='auto')
ax.set_axis_off()
plot_stem_loops(stem_loops[:2], (lo1-24, hi1+6), figure_width=7, max_label_length=60)
# -
from IPython.display import display
plot_stem_loops(stem_loops[2:], (lo2-12, hi2+6), n_lines=2, figure_width=8)
# +
import re
for m in re.finditer(r"(.)\1\1([au])\2\2[acu]", seq):
print(m.group(), m.start())
# +
import pprint
pprint.pprint(cdss)
# -
# !pip install viennarna
from wand.image import Image
Image(filename="5'UTR.ps")
Image(filename='dot.ps')
# +
import forgi
import forgi.graph.bulge_graph as fgb
import forgi.visual.mplotlib as fvm
fx = '\n'.join([elements[0]['seq'], elements[0]['mfe_structures'][1].split(' ', 1)[0]])
rna = fgb.BulgeGraph.from_fasta_text(fx)[0]
fig, (ax1) = plt.subplots(
1, 1, figsize=(12, 10)
)
fvm.plot_rna(rna, ax=ax1, lighten=.75)
plt.show()
# -
[f['GBFeature_quals'] for f in stem_loops]
# !pip install wand
print("| | label | type | locus | length | events/bp | entropy/bp | notes |\n|---:|:---------------------|:------------|:-----------|---------:|------------:|-------------:|:--------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|\n| 0 | 5'UTR | 5'UTR | | 265 | 0.479245 | 0.00320377 | |\n| 1 | nsp1 | mat_peptide | GU280_gp01 | 540 | 0.368519 | 0.00223148 | nsp1; produced by both pp1a and pp1ab |\n| 2 | nsp2 | mat_peptide | GU280_gp01 | 1914 | 0.429467 | 0.00215308 | produced by both pp1a and pp1ab |\n| 3 | nsp3 | mat_peptide | GU280_gp01 | 5835 | 0.320651 | 0.00158338 | former nsp1; conserved domains are: N-terminal acidic (Ac), predicted phosphoesterase, papain-like proteinase, Y-domain, transmembrane domain 1 (TM1), adenosine diphosphate-ribose 1''-phosphatase (ADRP); produced by both pp1a and pp1ab |\n| 4 | nsp4 | mat_peptide | GU280_gp01 | 1500 | 0.28 | 0.00140333 | nsp4B_TM; contains transmembrane domain 2 (TM2); produced by both pp1a and pp1ab |\n| 5 | 3C-like proteinase | mat_peptide | GU280_gp01 | 918 | 0.278867 | 0.00170915 | nsp5A_3CLpro and nsp5B_3CLpro; main proteinase (Mpro); mediates cleavages downstream of nsp4. 3D structure of the SARSr-CoV homolog has been determined (Yang et al., 2003); produced by both pp1a and pp1ab |\n| 6 | nsp6 | mat_peptide | GU280_gp01 | 870 | 0.375862 | 0.00172644 | nsp6_TM; putative transmembrane domain; produced by both pp1a and pp1ab |\n| 7 | nsp7 | mat_peptide | GU280_gp01 | 249 | 0.445783 | 0.00208032 | produced by both pp1a and pp1ab |\n| 8 | nsp8 | mat_peptide | GU280_gp01 | 594 | 0.26431 | 0.00117845 | produced by both pp1a and pp1ab |\n| 9 | nsp9 | mat_peptide | GU280_gp01 | 339 | 0.289086 | 0.00134218 | ssRNA-binding protein; produced by both pp1a and pp1ab |\n| 10 | nsp10 | mat_peptide | GU280_gp01 | 417 | 0.155875 | 0.000729017 | nsp10_CysHis; formerly known as growth-factor-like protein (GFL); produced by both pp1a and pp1ab |\n| 11 | nsp12 | mat_peptide | GU280_gp01 | 2795 | 0.26297 | 0.00148336 | nsp12; NiRAN and RdRp; produced by pp1ab only |\n| 12 | helicase | mat_peptide | GU280_gp01 | 1803 | 0.307266 | 0.00154631 | nsp13_ZBD, nsp13_TB, and nsp_HEL1core; zinc-binding domain (ZD), NTPase/helicase domain (HEL), RNA 5'-triphosphatase; produced by pp1ab only |\n| 13 | 3'-to-5' exonuclease | mat_peptide | GU280_gp01 | 1581 | 0.330803 | 0.00167426 | nsp14A2_ExoN and nsp14B_NMT; produced by pp1ab only |\n| 14 | endoRNAse | mat_peptide | GU280_gp01 | 1038 | 0.317919 | 0.00184971 | nsp15-A1 and nsp15B-NendoU; produced by pp1ab only |\n| 15 | nsp16_OMT | mat_peptide | GU280_gp01 | 894 | 0.293065 | 0.00156376 | nsp16_OMT; 2'-o-MT; produced by pp1ab only |\n| 16 | nsp11 | mat_peptide | GU280_gp01 | 39 | 0.230769 | 0.000666667 | produced by pp1a only |\n| 17 | S gene | gene | GU280_gp02 | 3822 | 0.360806 | 0.00189377 | |\n| 18 | ORF3a gene | gene | GU280_gp03 | 828 | 0.810386 | 0.00464614 | |\n| 19 | E gene | gene | GU280_gp04 | 228 | 0.342105 | 0.00153947 | |\n| 20 | M gene | gene | GU280_gp05 | 669 | 0.360239 | 0.00240359 | |\n| 21 | ORF6 gene | gene | GU280_gp06 | 186 | 0.510753 | 0.0021828 | |\n| 22 | ORF7a gene | gene | GU280_gp07 | 366 | 0.519126 | 0.00190437 | |\n| 23 | ORF7b gene | gene | GU280_gp08 | 132 | 0.477273 | 0.00218939 | |\n| 24 | ORF8 gene | gene | GU280_gp09 | 366 | 0.759563 | 0.00443989 | |\n| 25 | N gene | gene | GU280_gp10 | 1260 | 0.764286 | 0.00589048 | |\n| 26 | ORF10 gene | gene | GU280_gp11 | 117 | 0.615385 | 0.00362393 | |\n| 27 | 3'UTR | 3'UTR | | 229 | 1.12664 | 0.00556332 | |")
#
|
covid19/prelim.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1.2 引领浪潮的LaTeX
#
# ### 1.2.1 LaTeX的出现
#
# LaTeX是一款高质量的文档排版系统,LaTeX在读法上一般发作Lay-tek或者Lah-tek的音,而不是大家普遍认为的Lay-teks。LaTeX的历史可以追溯到1984年,在这一年里,兰波特博士作为早期开发者发布了LaTeX的最初版本。事实上,LaTeX完全是兰伯特博士的意外所得,他当年出于排版书籍的需要,在早先的文档排版系统TeX基础上新增了一些特定的宏包,为便于自己日后重复使用这些宏包,他将这些宏包构建成标准宏包。谁曾想,正是这些不经意间开发出来的宏包构成了LaTeX的雏形。
#
# <p align="center">
# <img align="middle" src="graphics/Leslie_Lamport.jpeg" width="200" />
# </p>
#
# <center><b>图1-2-1</b> 兰伯特博士,注:图片来源为兰伯特博士的维基百科网页。</center>
#
# 在很长一段时间里,LaTeX的版本其实没有多少大的更新,从技术层面来说,LaTeX实在没有什么可供更新的地方,它最初的面貌已趋近于完美且深入人心。LaTeX的最初版本是由兰伯特博士于上世纪80年代初开发出来的,目前,广泛使用的版本LaTeX2e是在1994年发布的,发布后一直没有大的更新,甚至发布后的首次更新出现在二十多年后的2020年。
#
# 尽管LaTeX2e的后续版本更新工作早在上世纪90年代初就已经开展,但时至今日,新版的LaTeX仍未进入人们的视野。从开发者兰伯特博士的视角来看,开发LaTeX的目的是为了降低TeX的使用门槛、发挥TeX强大的排版功能,提供一款高质量、解释性强的计算机程序语言,LaTeX最初定位的风格就是精简,这也是为什么LaTeX在日后可供提升的地方不是很多的原因。
# ### 1.2.2 LaTeX的特点
#
# 由于种种原因,时至今日,TeX慢慢淡出了人们的视线,不过我们依然能在LaTeX中看到TeX的身影:在使用LaTeX制作文档时,通常需要创建一个以`.tex`为拓展名的文件。通过LaTeX编译器对.tex文件进行编译,我们能得到一个PDF文档。
#
# 对于很多人来说,制作各类文档的首选可能是Word等软件,它简单好用、所写即所见,但当我们制作几十页甚至上百页的文档时,Word的劣势就会展露无疑,因为我们需要投入大量的时间和精力来对文档内容进行排版。反观LaTeX,它对文档的排版都是自动完成的,我们可在文档排版上节省大量的时间和精力,另外,使用LaTeX插入各种数学公式、表格、图形以及文献时,相应的索引出错的可能性也非常小,加之LaTeX对数学公式的强大排版能力,这些优点都是Word所无法比拟的。
#
# 在上个世纪80年代和90年代,LaTeX的用户群体非常庞大,然而,在世纪之交,随着微软推出的一系列Windows操作系统快速发展,相应的办公软件Office也以其便捷性吸引了人们的视线,致使大量LaTeX用户转而使用Office。即便如此,时至今日,LaTeX的用户群体依旧十分庞大,这主要得益于LaTeX强大的文档排版能力,虽然LaTeX复杂的语法结构与编译环境让很多初学者望而却步,但LaTeX能让用户更专注于内容创作,而非锦上添花的“排版”,这一特点也契合了人们对质量和效率的追求。在此基础上,具体来说,使得LaTeX历久弥新的关键可以归纳为以下五点:
#
# - 第一,LaTeX是专门用于制作文档的计算机程序语言。在众多计算机程序语言中,LaTeX可以制作排版质量极高的专业文档。
#
# - 第二,LaTeX拥有独特的创作方式。尽管LaTeX沿用了TeX排版系统的基本规则,但使用LaTeX制作文档时,内容创作和文档生成却是分开的,创作过程中也能随时预览创作文档。因此,在创作时,创作者不再像使用办公软件Word那样,既要关注创作内容,又要同步关注繁琐的排版和格式,使用LaTeX制作文档能在真正意义上让创作者专注于创作内容本身。值得一提的是,当文档篇幅较大时,使用LaTeX无疑会让我们在文档排版上节省大量的时间和精力。
#
# - 第三,LaTeX拥有简单的逻辑结构。使用LaTeX制作文档时,创作者可以通过一些非常简单的逻辑结构进行创作,如chapter(章)、section(节)、table(表格)。因此,LaTeX的使用门槛并不像常用的计算机程序语言那么高。
#
# - 第四,LaTeX对数学公式以及特殊符号具有极好的支持程度。众所周知,LaTeX在开发之初,是作为数学与计算机等相关领域研究人员的创作工具,这类群体喜欢使用LaTeX的原因无外乎是LaTeX可以通过一些简单的代码生成复杂的数学公式与特殊符号,编译后可呈现出高质量的排版效果。
#
# - 第五,LaTeX直接生成PDF文档。编译以`.tex`为拓展名的LaTeX文件后会得到一个PDF文档,PDF文档不存在跨平台、兼容性等问题,可以在各种操作系统上打开。
#
# 当然,除了上述五点,LaTeX制作文档的多元性这一特点也十分重要,LaTeX拥有众多封装好的文档类型,每一种文档类型对应着一类特定的文档结构及排版样式,从科技论文、技术报告、著作、学位论文、幻灯片甚至到科技绘图一应俱全,当然LaTeX也支持嵌入图片、绘制图形、设计表格、插入参考文献等。毋庸置疑,LaTeX在科技文档排版方面占有重要地位。
# 从LaTeX的出现到当下,它已经形成了一套非常高效的文档制作机制:
#
# - 文档类型 (document class)。文档类型是文档排版样式的基调,这些类型包括文章 (`article`)、报告 (`report`)、幻灯片 (`beamer`)等,申明文档类型往往是`.tex`文件的第一行代码,也是文档创作的第一步。
#
# - 宏包 (package)。它是LaTeX中的重要辅助工具,也可以把它理解为一般意义上的工具包。在使用时,调用宏包的基本命令为`\usepackage{}`,举例来说,包含颜色命令的宏包为`color`,其调用语句为`\usepackage{color}`。随着LaTeX的发展,越来越多的宏包被开发出来,这些宏包能满足特定的需求,如制表、插图、绘图,同时也能让LaTeX代码变得更加简洁。
#
# - 模板 (template)。LaTeX的发展催生了很多视觉和审美效果极好的模板,包括论文模板、幻灯片模板、报告模板甚至著作模板,这些模板在一定程度上能减少创作者在文档排版上的时间开销,也有很多学术刊物会给投稿作者提供相应的LaTeX模板。
#
# 通过对比LaTeX和Word,我们还会看到:
#
# - 第一,LaTeX的`.tex`源文件是无格式的,编译之后,根据特定的模板和指定的格式形成最终的PDF文档。因此,使用LaTeX制作文档能轻松切换文档类型、调整模板以及修改格式。
#
# - 第二,LaTeX对数学公式、图表以及文献索引的支持程度是Word所无法比拟的。尤为特殊的是,当文献数量达到上百篇时,在Word中修改参考文献可能是“牵一发而动全身”,费时耗力,而LaTeX根据已经整理好的`.bib`文件可自动完成文献引用与参考文献生成。
# ### 1.2.3 LaTeX编辑器
#
# 实际上,配置LaTeX环境包括两部分,即编译器和编辑器,对应的英文表达分别是complier和editor,两者不是一回事。LaTeX编译器又称为LaTeX编译工具,可根据系统属性与配置安装相应的编译工具:
#
# - Linux系统:可安装TeX Live,该编辑器拥有LaTeX编辑器;
# - Mac OS系统:可安装Mac TeX,该编译器拥有完整的TeX/LaTeX环境和LaTeX编辑器;
# - Windows系统:可安装MiKTeX或TeX Live,两者都拥有完整的TeX/LaTeX环境和LaTeX编辑器。
#
# 一般而言,LaTeX编辑器的界面大致由两部分组成,即LaTeX源码编译区域与PDF文档预览区域。以下几款LaTeX编辑器的使用体验较受人推崇:
#
# - TeXworks:这是TeX Live自带的一款轻量级编辑器。
# - TeXstudio:这款编辑器集代码编译与文档预览于一身。
# - WinEdt:这是CTeX自带的一款编辑器。
# - VS Code:这是微软推出的一款免费文本编辑器,功能包括文本编辑、日常开发等。
# - Atom:这是一款开源的跨平台编辑器(GitHub网址为[https://github.com/atom/atom](https://github.com/atom/atom)),支持多种程序语言。
# 【回放】[**1.1 横空出世的TeX**](https://nbviewer.jupyter.org/github/xinychen/latex-cookbook/blob/main/chapter-1/section1.ipynb)
#
# 【继续】[**1.3 应运而生的在线系统**](https://nbviewer.jupyter.org/github/xinychen/latex-cookbook/blob/main/chapter-1/section3.ipynb)
# ### License
#
# <div class="alert alert-block alert-danger">
# <b>This work is released under the MIT license.</b>
# </div>
|
chapter-1/section2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
iris=pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-database/iris/iris.data")
df=iris.head()
df.head(3)
df.columns=['sl','sw','pl','pw','flower_type']
print(df.shape)
print(df.dtypes)
df.describe()
df.sl
df["sl"]
df.isnull()
df.isnull().sum()
df.iloc[1:4,2:4]
df.head()
df.drop(0)
#drop by label
a=df.drop(0)
a.head()
df.drop(0,inplace=True)
#drop by position
df.drop(df.index[0])
df.index[0],df.index[3]
df.drop(df.index[0],inplace=True)
df.drop(df.index[[0,1]],inplace=True)
df.sl>3
df[df.sl>5]
df[df.flower_type=='Iris_setosa'].describe()
#add a row
df.iloc[0]
print(df.iloc[0])
print(df.loc[0])
df.loc[0]=[1,2,3,4,"Iris_setosa"]
df.tail()
df.reset_index()
df.reset_index(drop=True)
df.reset_index(drop=True,inplace=True)
df.drop('sl',axis=1,inplace=True)
del df['sw']
df=iris.copy()
df.columns=['sl','sw','pl','pw','flower_type']
df.describe()
df["diff_pl_pw"]=df["pl"]-df["pw"]
df.tail()
|
Lecture 5 Pandas/Manipulating Data in data frame-2/Manipulating Data in DataFrame -2-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Code for **"Inpainting"** figures $6$, $8$ and 7 (top) from the main paper.
# # Import libs
# +
from __future__ import print_function
import matplotlib.pyplot as plt
# %matplotlib inline
import os
# os.environ['CUDA_VISIBLE_DEVICES'] = '1'
import numpy as np
from models.resnet import ResNet
from models.unet import UNet
from models.skip import skip
import torch
import torch.optim
from torch.autograd import Variable
from utils.inpainting_utils import *
torch.backends.cudnn.enabled = True
torch.backends.cudnn.benchmark =True
dtype = torch.cuda.FloatTensor
PLOT = True
imsize=-1
dim_div_by = 64
dtype = torch.cuda.FloatTensor
# -
# # Choose figure
# +
## Fig 6
# img_path = 'data/inpainting/2.png'
# mask_path = 'data/inpainting/2_mask.png'
## Fig 8
# img_path = 'data/inpainting/1.png'
# mask_path = 'data/inpainting/1_mask.png'
## Fig 7 (top)
img_path = 'data/inpainting/lena.png'
mask_path = 'data/inpainting/lena_mask.png'
NET_TYPE = 'skip_depth6' # one of skip_depth4|skip_depth2|UNET|ResNet
# -
# # Load mask
img_pil, img_np = get_image(img_path, imsize)
img_mask_pil, img_mask_np = get_image(mask_path, imsize)
# ### Center crop
# +
img_mask_pil = crop_image(img_mask_pil, dim_div_by)
img_pil = crop_image(img_pil, dim_div_by)
img_np = pil_to_np(img_pil)
img_mask_np = pil_to_np(img_mask_pil)
# -
# ### Visualize
# +
img_mask_var = np_to_var(img_mask_np).type(dtype)
plot_image_grid([img_np, img_mask_np, img_mask_np*img_np], 3,11);
# -
# # Setup
pad = 'reflection' # 'zero'
OPT_OVER = 'net'
OPTIMIZER = 'adam'
# +
if '2.png' in img_path:
INPUT = 'meshgrid'
input_depth = 2
LR = 0.1
num_iter = 5001
param_noise = False
show_every = 100
figsize = 5
net = skip(input_depth, img_np.shape[0],
num_channels_down = [16, 32, 64, 128, 128],
num_channels_up = [16, 32, 64, 128, 128],
num_channels_skip = [0, 0, 0, 0, 0],
upsample_mode='nearest', filter_skip_size=1,
need_sigmoid=True, need_bias=True, pad=pad, act_fun='LeakyReLU').type(dtype)
elif 'lena.png' in img_path:
INPUT = 'noise'
input_depth = 32
LR = 0.01
num_iter = 3001
param_noise = False
show_every = 500
figsize = 5
net = skip(input_depth, img_np.shape[0],
num_channels_down = [16, 32, 64, 128, 128],
num_channels_up = [16, 32, 64, 128, 128],
num_channels_skip = [0, 0, 0, 0, 4],
filter_size_up = 7, filter_size_down = 7,
upsample_mode='nearest', filter_skip_size=1,
need_sigmoid=True, need_bias=True, pad=pad, act_fun='LeakyReLU').type(dtype)
elif '1.png' in img_path:
INPUT = 'noise'
input_depth = 1
num_iter = 3001
show_every = 250
figsize = 8
param_noise = True
if 'skip' in NET_TYPE:
depth = int(NET_TYPE[-1])
net = skip(input_depth, img_np.shape[0],
num_channels_down = [16, 32, 64, 128, 128, 128][:depth],
num_channels_up = [16, 32, 64, 128, 128, 128][:depth],
num_channels_skip = [0, 0, 0, 0, 0, 0][:depth],
filter_size_up = 3,filter_size_down = 5, filter_skip_size=1,
upsample_mode='nearest', # downsample_mode='avg',
need1x1_up=False,
need_sigmoid=True, need_bias=True, pad=pad, act_fun='LeakyReLU').type(dtype)
LR = 0.01
elif NET_TYPE == 'UNET':
net = UNet(num_input_channels=input_depth, num_output_channels=3,
feature_scale=8, more_layers=1,
concat_x=False, upsample_mode='deconv',
pad='zero', norm_layer=torch.nn.InstanceNorm2d, need_sigmoid=True, need_bias=True)
LR = 0.001
param_noise = False
elif NET_TYPE == 'ResNet':
net = ResNet(input_depth, img_np.shape[0], 8, 32, need_sigmoid=True, act_fun='LeakyReLU')
LR = 0.001
param_noise = False
else:
assert False
else:
assert False
net = net.type(dtype)
net_input = get_noise(input_depth, INPUT, img_np.shape[1:]).type(dtype)
# +
# Compute number of parameters
s = sum(np.prod(list(p.size())) for p in net.parameters())
print ('Number of params: %d' % s)
# Loss
mse = torch.nn.MSELoss().type(dtype)
img_var = np_to_var(img_np).type(dtype)
mask_var = np_to_var(img_mask_np).type(dtype)
# -
# # Main loop
# +
i = 0
def closure():
global i
if param_noise:
for n in [x for x in net.parameters() if len(x.size()) == 4]:
n.data += n.data.clone().normal_()*n.data.std()/50
out = net(net_input)
total_loss = mse(out * mask_var, img_var * mask_var)
total_loss.backward()
print ('Iteration %05d Loss %f' % (i, total_loss.data[0]), '\r', end='')
if PLOT and i % show_every == 0:
out_np = var_to_np(out)
plot_image_grid([np.clip(out_np, 0, 1)], factor=figsize, nrow=1)
i += 1
return total_loss
p = get_params(OPT_OVER, net, net_input)
optimize(OPTIMIZER, p, closure, LR, num_iter)
# -
out_np = var_to_np(net(net_input))
plot_image_grid([out_np], factor=5);
|
0-newbooks/deep-image-prior/inpainting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: p3.8
# language: python
# name: p3.8
# ---
# # Mathematical model of the curtain
# ## Model parameters
# - $a$: The height of the curtain, $\in [3,7]$m
# - $b$: The width of the curtain, $\in [4,8]$m
# - $c$: The thickness of the curtain, $\in [0.5, 2]$mm
# - $\rho = 1300$ kg/(m^3): Density of PVC
# - $r_r$: The radius of the core of the roll
# - $m_r$: The mass of the core, assumed to be a thin-walled tube.
# - $m_{w}$: Mass of the weights in the bottom of the curtain which keep it stretched
# ## Signals
# - $y$: Output signal. The height of the opening. $a \in [0, a)$ and $a-y$ is the amount of hanging curtain.
# - $\theta$: The angle of the roller axis. This has a static relationship with $y$ derived below.
# - $u$: Input signal. The torque acting on the roller axis.
# ## Kinematic relationships
# ### The radius of the roll as a function of the opening $y$
# We assume a circular shape of the cross-section. The area of this is $A=\pi r^2$, which must equal the sum of the area of the core $A_c = \pi r_r^2$ and the area of the rolled up curtain $A_c = cy$. So,
# $$r = \sqrt{r_r^2 + \frac{c}{\pi}y}, \quad \text{and}$$
# $$ y = \frac{\pi}{c}(r^2 - r_r^2).$$
# ### Relationship between $y$ and $\theta$
# Start with the relationship between the velocities
# $$ r(y)\frac{d\theta}{dt} = \frac{dy}{dt},$$
# which gives
# $$ \frac{dy}{r(y)} = d\theta$$
# $$ \frac{1}{\sqrt{r_r^2 + \frac{c}{\pi}y}} dy = d\theta.$$
# Integrate both sides to obtain
# $$ \frac{2\pi}{c}\sqrt{r_r^2 + \frac{c}{\pi} y} = \theta + K. $$
# We need an initial condition between $y$ and $\theta$ to determine the constant $K$.
# Let $y=0$ imply $\theta = 0$, hence
# $$ K = \frac{2\pi}{c}r_r.$$
# To find $y$ as a function of $\theta$:
# $$ r_r^2 + \frac{c}{\pi}y = (\frac{c}{2\pi} \theta + r_r)^2$$
# $$ r_r^2 + \frac{c}{\pi}y = (\frac{c}{2\pi})^2 \theta^2 + \frac{cr_r}{\pi}\theta + r_r^2$$
# $$ \frac{c}{\pi}y = (\frac{c}{2\pi})^2 \theta^2 + \frac{cr_r}{\pi}\theta$$
# $$ y = \frac{c}{4\pi}\theta^2 + r_r \theta = \theta(\frac{c}{4\pi}\theta + r_r).$$
# ## Inertial properties
# ### Mass of the hanging curtain and rolled-up curtain
# The hanging curtain has mass
# $$m_c = m_w + \rho b c (a-y),$$
# and the rolled up part
# $$m_{rc} = \rho b c y.$$
# ### Moment of inertia of the rolled-up and hanging curtain
# The moment of inertia of the core is
# $$I_r = m_rr_r^2, $$
# the moment of inertia of the rolled-up portion of the curtain is
# $$I_{rc}(y) = \frac{1}{2} m_{rc}(r^2 - r_r^2) = \frac{\rho b c^2}{\pi}y^2, $$ and the hanging mass, considering it to be a point mass at distance $r(y)$ from the axis of the roller is
# $$I_c(y) = m_c r(y)^2 = (m_w + \rho b c (a-y))(r_r^2 + \frac{c}{\pi}y). $$
# The complete moment of inertia with respect to the axis of the roller is
# $$I(y) = I_r + I_{cr}(y) + I_c(y).$$
# ## Friction
# We consider two types of friction. A friction in the bearings of the roller, and a friction between the curtain and the rails it travels in. The friction moment in the roller bearings is
# $$ F_{r} = f_r \frac{d \theta}{dt}.$$
# The friction in the curtain will be proportional to both the velocity $\frac{dy}{dt}$ and the length of hanging curtain $a-y$:
# $$F_c = f_c (a-y) \frac{d y}{dt}.$$
# ## ODE
# $$I(y) \ddot{\theta} = -m_c(y)r(y)g - f_r\dot{\theta}-f_c(a-y)\dot{y} + u.$$
import sympy as sy
sy.init_printing()
a,b,c,rr,rho,m_r,m_w = sy.symbols('a b c r_r rho m_r m_w')
y,u,theta,r = sy.symbols('y u theta r')
At = sy.pi*r**2
Ar = sy.pi*rr**2
Ac = y*c
eq1 = At - Ar -Ac
eq1
sy.solve(eq1, y)
sy.integrate(1/sy.sqrt(rr**2 + c/sy.pi*y), y)
|
challenges/curtain/notebooks/.ipynb_checkpoints/Mathematical model-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
df = pd.read_csv('movies_metadata.csv', low_memory=False)
df.head()
# -
c=df['vote_average'].mean()
c
m=df['vote_count'].quantile(0.9)
m
qm=df[(df['runtime']>150)&(df['revenue']>300000000)&(df['homepage'].notna())]
qm=qm[qm['vote_count']>=m]
qm.shape
def wr(x,m=m,c=c):
v=x['vote_count']
r=x['vote_average']
return (v/(v+m)*r)+(m/(m+v)*c)
qm['score']=qm.apply(wr,axis=1)
qm=qm.sort_value('score',ascedinding=False)
qm[['title','vote_count','vote_average','score']].head(10)
|
Section 2/Video 2.4/Video 2.4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Pima dataset for classification
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
import seaborn as sns
df=pd.read_csv('/Users/dantohe/Development/Projects/Springboard/datasets/Pima_Indians_Diabetes/diabetes.csv')
df.info()
df.shape
df.columns
# ### Pick a subset of features
feature_columns = ['Pregnancies','Insulin','BMI','Age']
x= df[feature_columns]
y = df.Outcome
x.info()
y.head
# ### Splitting in training and testing
# By default train_test_split will allocate 25% of the data set to testing.
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test = train_test_split(x,y,random_state=0)
x_train.shape
y_train.shape
x_test.shape
# ### Using logistic regression for learning.
from sklearn.linear_model import LogisticRegression
from sklearn import preprocessing
from sklearn.metrics import accuracy_score
#fitting
lr = LogisticRegression()
lr.fit(x_train, y_train)
#class prediction
y_pred_class = lr.predict(x_test)
#these are the predictions from the model
y_pred_class
# ### Calculate the classification accurcy
# How many were correctly classified.
print(accuracy_score(y_test, y_pred_class))
# ### The NULL ACCURACY
# The accuracy achieved by allways predicting the most frequent class.
y_test.count()
y_test.value_counts()
y.value_counts()
# +
#how many ones are there
# -
y_test.mean()
1-y_test.mean()
## null accuracy - for binary class
max(y_test.mean(),1-y_test.mean())
## null accuracy - for multi class
y_test.value_counts().head(1)/len(y_test)
# ### Comfusion Matrix
# ACTUAL, PREDICTED !!!!!
from sklearn import metrics
print(metrics.confusion_matrix(y_test,y_pred_class))
confusion_matrix = metrics.confusion_matrix(y_test, y_pred_class)
tp = confusion_matrix[1,1]
tn = confusion_matrix[0,0]
fn = confusion_matrix[1,0]
fp = confusion_matrix[0,1]
# ### Accuracy
accuracy = (tp+tn)*1.0/(tp+tn+fp+fn)
print(accuracy)
print(metrics.accuracy_score(y_test,y_pred_class))
# ### Classification error
classification_error = 1 - accuracy
print(1-metrics.accuracy_score(y_test,y_pred_class))
# ### Sensitivity - RECALL
# When the value is positive how many times is it predicted correctly.
sensitivity = tp*1.0/(tp+fn)
print(metrics.recall_score(y_test,y_pred_class))
# ### Speciicity
# When the value is negative how often is predicted corectly.
speciicity = tn*1.0/(tn+fp)
print(speciicity)
# ### Precision
# When te value is predicted how oten is it predicted correctly.
precision = tp *1.0/(tp+fp)
precision
print(metrics.precision_score(y_test,y_pred_class))
lr.predict(x_test)[0:10]
lr.predict_proba(x_test)[0:10]
lr.predict_proba(x_test)[0:10, 1]
y_pred_prob = lr.predict_proba(x_test)[0:10, 1]
import matplotlib.pyplot as plt
# plt.rcParams['font.size'=14]
plt.hist(y_pred_prob, bins =8)
plt.xlim(0,1)
# ### Decrese the treshhold - increase sensitivity
from sklearn.preprocessing import binarize
# ### AUC
# from sklearn.cross_validation
metrics.roc_auc_score(y_test, y_pred_class)
y_pred_prob = lr.predict_proba(x_test)[:, 1]
fpr, tpr, thresholds = metrics.roc_curve(y_test, y_pred_prob)
plt.plot(fpr, tpr)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title('ROC curve for diabetes classifier')
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.grid(True)
print(metrics.roc_auc_score(y_test, y_pred_prob))
# +
# calculate cross-validated AUC
from sklearn.model_selection import cross_val_score
cross_val_score(lr, x, y, cv=10, scoring='roc_auc').mean()
# -
|
classification/classification_pima_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TASK -2
# ## Exploring Supervised Machine Learning
#
# #### (we will predict the percentage of marks that a student is expected to score based upon the number of hours they studied.)
# + [markdown] colab_type="text" id="X6A8Hm86UUZ-"
# ### <NAME> ( _Data Science & Analytics Intern_ )
# **<EMAIL>**
# + colab={} colab_type="code" id="V9QN2ZxC38pB"
# Importing the liberaries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" executionInfo={"elapsed": 2534, "status": "ok", "timestamp": 1544113345787, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WI8p7JNWLic/AAAAAAAAAAI/AAAAAAAAAfs/vS8ElgH0p0c/s64/photo.jpg", "userId": "15341571102300750919"}, "user_tz": -480} id="LtU4YMEhqm9m" outputId="5b4b36af-1545-497e-a6dc-7658bab71dbc"
data = pd.read_csv("http://bit.ly/w-data")
print("Data from Link Imported here")
data.head()
# + [markdown] colab_type="text" id="RHsPneuM4NgB"
# #### Representing relationship graph of Hrs. vs %
# +
# Plotting the distribution of scores
data.plot(x='Hours', y='Scores', style='o')
#Changing tittle & label of graph
plt.title('Hours vs Percentage')
plt.xlabel('Hours Studied')
plt.ylabel('Percentage Score')
#Displaying Plot
plt.show()
# + [markdown] colab_type="text" id="fiQaULio4Rzr"
# **It's clear from the above graph that, Linear relation b/w hrs. & score_%**
# + [markdown] colab_type="text" id="WWtEr64M4jdz"
# ### **Data Preparation**
# + colab={} colab_type="code" id="LiJ5210e4tNX"
x = data.iloc[:, :-1].values
y = data.iloc[:, 1].values
# + colab={} colab_type="code" id="udFYso1M4BNw"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size=0.2, random_state=0)
# + [markdown] colab_type="text" id="a6WXptFU5CkC"
# ### **Training the Algorithm**
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 701, "status": "ok", "timestamp": 1544113358086, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WI8p7JNWLic/AAAAAAAAAAI/AAAAAAAAAfs/vS8ElgH0p0c/s64/photo.jpg", "userId": "15341571102300750919"}, "user_tz": -480} id="qddCuaS84fpK" outputId="befbd977-772c-4bd1-bb48-ee5dd6bae73c"
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
print("Training Done")
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" executionInfo={"elapsed": 985, "status": "ok", "timestamp": 1544113360867, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WI8p7JNWLic/AAAAAAAAAAI/AAAAAAAAAfs/vS8ElgH0p0c/s64/photo.jpg", "userId": "15341571102300750919"}, "user_tz": -480} id="J61NX2_2-px7" outputId="d20ec1fd-3e2d-4eae-84a2-a0df57d31009"
# Plotting the regression line
line = regressor.coef_*X+regressor.intercept_
# Plotting for the test data
plt.scatter(X, y)
plt.plot(X, line);
plt.show()
# + [markdown] colab_type="text" id="JCQn-g4m5OK2"
# ### **Making Predictions**
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 698, "status": "ok", "timestamp": 1544113363729, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WI8p7JNWLic/AAAAAAAAAAI/AAAAAAAAAfs/vS8ElgH0p0c/s64/photo.jpg", "userId": "15341571102300750919"}, "user_tz": -480} id="Tt-Fmzu55EGM" outputId="46f1acf8-91ac-4984-cfbe-e614aa9ea849"
# Testing data - In Hours
print(X_test)
# Predicting the scores
y_pred = regressor.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 753, "status": "ok", "timestamp": 1544113366918, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WI8p7JNWLic/AAAAAAAAAAI/AAAAAAAAAfs/vS8ElgH0p0c/s64/photo.jpg", "userId": "15341571102300750919"}, "user_tz": -480} id="6bmZUMZh5QLb" outputId="8ea11a9e-c1b7-4fab-ab62-4dcbd2c8607b"
# Comparing Actual vs Predicted
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
print(df)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 862, "status": "ok", "timestamp": 1544113370494, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WI8p7JNWLic/AAAAAAAAAAI/AAAAAAAAAfs/vS8ElgH0p0c/s64/photo.jpg", "userId": "15341571102300750919"}, "user_tz": -480} id="KAFO8zbx-AH1" outputId="fcb3830f-3cda-4dcb-f122-84b71f101fae"
# You can also test with your own data
hours = 9.25
pred = regressor.predict(np.array([9.25]).reshape(-1,1))
print("No of Hours = {}".format(hours))
print("Predicted Score = {}".format(pred[0]))
# + [markdown] colab_type="text" id="0AAsPVA_6KmK"
# ### **Evaluating the model**
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 834, "status": "ok", "timestamp": 1544113374919, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WI8p7JNWLic/AAAAAAAAAAI/AAAAAAAAAfs/vS8ElgH0p0c/s64/photo.jpg", "userId": "15341571102300750919"}, "user_tz": -480} id="r5UOrRH-5VCQ" outputId="7b9ddcf1-2848-408f-d81f-7a60652c381e"
from sklearn import metrics
print('Mean Absolute Error:',metrics.mean_absolute_error(y_test, y_pred))
# -
# # **Finally Submited the Task**
# #### Thank You **THE SPARKS FOUNDTION**
|
Task_1_Linear Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
path_raw = '../data/raw/original/'
filenames = ['Measurement_info.csv', 'Measurement_item_info.csv', 'Measurement_station_info.csv']
for i in range(0,3):
df = pd.read_csv(path_raw+filenames[i])
print(filenames[i], df.shape)
print(df.head(6))
print('____________________________\n')
# +
# Does instrument status vary within the same hour at the same station?
df = pd.read_csv(path_raw+filenames[0])
df_inst = df[['Measurement date', 'Station code', 'Instrument status']].copy()
stds = df_inst.groupby(['Measurement date', 'Station code']).std()
stds[(stds.select_dtypes(include=['number']) != 0).any(1)]
# +
# look at one datetime stamp at one station location to see how instrument status varies
df[(df.select_dtypes(include=['object']) == '2017-01-01 04:00').any(1)][(df.select_dtypes(include=['int']) == 112).any(1)]
# +
# Okay, let's go make some masks for the data
# -
|
notebooks/explore_data_original.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## pCMV-Target-AID
# QUEEN script for pCMV-Target-AID construction.
# 1. The N-terminus half of Target-AID was amplified from pcDNA3.1_pCMV-nCas-PmCDA1-ugi pH1-gRNA(HPRT) using the primer pair RS045/HM129.
# 2. The C-terminus half of Target-AID was amplified from pcDNA3.1_pCMV-nCas-PmCDA1-ugi pH1-gRNA(HPRT) using the primer pair HM128/RS046.
# 3. A backbone fragment was amplified from pCMV-ABE7.10 using the primer pair RS047/RS048.
# 4. The three fragments were assembled by Gibson Assembly.
# %matplotlib inline
import sys
from QUEEN.queen import *
set_namespace(globals())
if "output" not in os.listdir("./"):
os.mkdir("output")
QUEEN(record="https://benchling.com/s/seq-K4HkSd2E8WiTAulJUeBf", dbtype="benchling", product="pCMV_ABE") #Load pCMV-ABE plasmid object.
QUEEN(record="https://benchling.com/s/seq-cfnGDU0Mq8cUwn185LPF", dbtype="benchling", product="pcDNA31_Target_AID") #Load pCDNA3.1-Target-AID plasmid object.
processname1 = "PCR"
description1 = "1. The N-terminus half of Target-AID was amplified from pcDNA3.1_pCMV-nCas-PmCDA1-ugi pH1-gRNA(HPRT) using the primer pair RS045/HM129."
QUEEN("GAGAGCCGCCACCATGGCACCGAAGAAGAAGCG", product="RS045") #Create a QUEEN object for the forward primer.
QUEEN("CTGGGGCACGATATGATCCACGTCGTAGTCGGAGA", product="HM129") #Create a QUEEN object for the reverse primer.
pcDNA31_Target_AID.searchsequence(RS045.seq[-18:], product="FW1", pn=processname1, pd=description1) #Search for the 18-bp 3’-end sequences of the forward primer.
pcDNA31_Target_AID.searchsequence(HM129.seq[-18:], product="RV1", pn=processname1, pd=description1) #Search for the 18-bp 3’-end sequences of the reverse primer.
cropdna(pcDNA31_Target_AID, FW1[0].end, RV1[0].start, product="extract1", pn=processname1, pd=description1) #Crop the internal DNA sequence flanked by the primer annealing sites.
modifyends(extract1, RS045.seq, HM129.rcseq, product="fragment1", pn=processname1, pd=description1) #Add forward and reverse primer sequences to the both ends of the cropped fragment.
processname2 = "PCR"
description2 = "2. The C-terminus half of Target-AID was amplified from pcDNA3.1_pCMV-nCas-PmCDA1-ugi pH1-gRNA(HPRT) using the primer pair HM128/RS046."
QUEEN(seq="CTACGACGTGGATCATATCGTGCCCCAGTCTTTTC", product="HM128") #Create a QUEEN object for the forward primer.
QUEEN(seq="TTTAAACTCATTATAGCATCTTGATCTTGTTCTCTC", product="RS046") #Create a QUEEN object for the reverse primer.
pcDNA31_Target_AID.searchsequence(HM128.seq[-18:], product="FW2", pn=processname2, pd=description2) #Search for the 18-bp 3’-end sequences of the forward primer.
pcDNA31_Target_AID.searchsequence(RS046.seq[-18:], product="RV2", pn=processname2, pd=description2) #Search for the 18-bp 3’-end sequences of the reverse primer.
f2 = cropdna(pcDNA31_Target_AID, FW2[0].end, RV2[0].start, product="extract2", pn=processname2, pd=description2) #Crop the internal DNA sequence flanked by the primer annealing sites.
modifyends(extract2, HM128.seq, RS046.rcseq, product="fragment2", pn=processname2, pd=description2) #Add forward and reverse primer sequences to the both ends of the cropped fragment.
processname3 = "PCR"
description3 = "3. A backbone fragment was amplified from pCMV-ABE7.10 using the primer pair RS047/RS048."
QUEEN("ATCAAGATGCTATAATGAGTTTAAACCCGCTGATC", product="RS047") #Create a QUEEN object for the forward primer.
QUEEN("CTTCGGTGCCATGGTGGCGGCTCTCCCTATAG", product="RS048") #Create a QUEEN object for the reverse primer.
pCMV_ABE.searchsequence(RS047.seq[-18:], product="FW3", pn=processname3, pd=description3) #Search for the 18-bp 3’-end sequences of the forward primer.
pCMV_ABE.searchsequence(RS048.seq[-18:], product="RV3", pn=processname3, pd=description3) #Search for the 18-bp 3’-end sequences of the reverse primer.
f3 = cropdna(pCMV_ABE, FW3[0].end, RV3[0].start, product="extract3", pn=processname3, pd=description3) #Crop the internal DNA sequence flanked by the primer annealing sites.
modifyends(extract3, RS047.seq, RS048.rcseq, product="fragment3", pn=processname3, pd=description3) #Add forward and reverse primer sequences to the both ends of the cropped fragment.
processname4 = "Gibson Assembly"
description4 = "4. The three fragments were assembled by Gibson Assembly."
modifyends(fragment1, "*{25}/-{25}","-{28}/*{28}", product="fragment1_mod", pn=processname4, pd=description4) #Generate long sticky ends on the both sides of "fragment1".
modifyends(fragment2, "*{28}/-{28}","-{25}/*{25}", product="fragment2_mod", pn=processname4, pd=description4) #Generate long sticky ends on the both sides of "fragment2".
modifyends(fragment3, "*{25}/-{25}","-{25}/*{25}", product="fragment3_mod", pn=processname4, pd=description4) #Generate long sticky ends on the both sides of "fragment3".
joindna(fragment1_mod, fragment2_mod, fragment3_mod, topology="circular", product="pCMV_Target_AID", pn=processname4, pd=description4) #Join the fragments.
pCMV_Target_AID.printfeature()
# +
def add_fragment_annotation(dna, fragment, new_feature_id, color_set):
"""
Define seqeunce featurtes for "fragment" in "dna".
"""
f = dna.searchsequence(fragment.seq) #Search for "fragment" seqeunce in "dna"
editfeature(dna, source=f, target_attribute="feature_id", operation=createattribute(new_feature_id), new_copy=False) #Define a new feature whose feature_id becomes "new_feature_id".
editfeature(dna, key_attribute="feature_id", query=new_feature_id, target_attribute="qualifier:label", operation=createattribute(new_feature_id), new_copy=False) #Provide a qualifier:label to the new feature.
editfeature(dna, key_attribute="feature_id", query=new_feature_id, target_attribute="qualifier:edgecolor_queen", operation=createattribute(color_set[0]), new_copy=False) #Set a edge color for the new feature.
editfeature(dna, key_attribute="feature_id", query=new_feature_id, target_attribute="qualifier:facecolor_queen", operation=createattribute(color_set[1]), new_copy=False) #Set a face color for the new feature.
editfeature(dna, key_attribute="feature_id", query=new_feature_id, target_attribute="strand", operation=replaceattribute(0), new_copy=False) #Set a coding direction of the new feature
def visualization(dna):
"""
Set the start position of "dna" and visualize its circular sequence map.
"""
dna.searchfeature(key_attribute="qualifier:label", query="^Cas9", product="Cas9") #Search for the feature that represent the Cas9 gene.
plasmid = joindna(cutdna(dna, Cas9[0].start)[0], topology="circular") #Set the first nucleotide of the Cas9 gene as the start position of the plasmid.
fragments = plasmid.searchfeature(key_attribute="qualifier:label", query="fragment-[0-9]+") #Obtain the features whose feature_id are "fragment-[0-9]+".
fragments.sort(key=lambda x:x.qualifiers["label"][0]) #Sort the order of fragments according to qualifier.label.
features = plasmid.searchfeature(key_attribute="feature_type", query="CDS") + plasmid.searchfeature(key_attribute="feature_type", query="promoter") + plasmid.searchfeature(key_attribute="feature_type", query="rep_origin") + fragments #Select the sequence features to be visualized.
fig = visualizemap(plasmid, feature_list=features, map_view="circular", tick_interval=1000, title=dna.project) #Visualize the circular seqeunce map of the plasmid.
return fig, plasmid
#Paris of color codes for gene edge and face
color_sets = [('#E53935', '#ffcdd2'), ('#8E24AA', '#e1bee7'), ('#3949AB', '#c5cae9'), ('#1E88E5', '#bbdefb'), ('#00ACC1', '#b2ebf2'), ('#D81B60', '#f8bbd0'), ('#5E35B1', '#d1c4e9'),
('#43A047', '#c8e6c9'), ('#7CB342', '#dcedc8'), ('#FDD835', '#fff9c4'), ('#FB8C00', '#ffe0b2'), ('#6D4C41', '#d7ccc8'), ('#C0CA33', '#f0f4c3'), ('#546E7A', '#cfd8dc')]
add_fragment_annotation(pCMV_Target_AID, fragment1, "fragment-1", color_sets[0]) #Add "fragment1" to .dnafeatures of "pCMV_Target_AID" plasmid object.
add_fragment_annotation(pCMV_Target_AID, fragment2, "fragment-2", color_sets[1]) #Add "fragment2" to .dnafeatures of "pCMV_Target_AID" plasmid object.
add_fragment_annotation(pCMV_Target_AID, fragment3, "fragment-3", color_sets[2]) #Add "fragment3" to .dnafeatures of "pCMV_Target_AID" plasmid object.
fig, plasmid = visualization(pCMV_Target_AID) #Visualize the circular sequence map of pCMV-Target-AID.
# -
#Visualize the map of pCMV-Target-AID.
fig = visualizemap(plasmid, start=8688, end=5335, width_scale=0.40, tick_interval=1000, title="", fontsize=13, height_scale=1.05)
fig.savefig("output/fig2c.pdf", bbox_inches="tight")
#Visualize the operational process chart of pCMV-Target-AID.
flow = visualizeflow(pCMV_Target_AID, alias_dict={"fragment1":"fragment-1", "fragment2":"fragment-2", "fragment3":"fragment-3", "pcDNA31_Target_AID": "pcDNA31-Target-AID", "pCMV_ABE": "pCMV-ABE", "pCMV_Target_AID": "pCMV-Target-AID"})
flow.render("output/pCMV_Target_AID_construction")
flow
pCMV_Target_AID.outputgbk("output/pCMV-Target-AID.gbk")
quine(pCMV_Target_AID, execution=True)
quine(pCMV_Target_AID, process_description=True)
|
demo/sakata_et_al_2020/pCMV_Target_AID_construction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-3.4 Intro Python
# # The Power of List Iteration
# - for in: **`for`** loop using **`in`**
# - for range: **`for range(start,stop,step)`**
# - more list methods: **`.extend()`, `+, .reverse(), .sort()`**
# - **strings to lists,`.split()`, and list to strings, `.join()`**
# - **list cast & `print("hello", end='')`**
#
#
# -----
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - Iterate through Lists using **`for`** with **`in`**
# - Use **`for range()`** in looping operations
# - Use list methods **`.extend()`, `+, .reverse(), .sort()`**
# - **convert between lists and strings using `.split()` and `.join()`**
# - **cast strings to lists / direct multiple print outputs to a single line**
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Converting a string to a list with `.split()`
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/1a076a9c-842f-455f-91bf-48db837842e8/Unit2_Section3.4a-Split_on_Breaks.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/1a076a9c-842f-455f-91bf-48db837842e8/Unit2_Section3.4a-Split_on_Breaks.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### `.split()` by default, splits a string at spaces (" ") to create a list
# ```python
# tip = "Notebooks can be exported as .pdf"
# tip_words = tip.split()
#
# for word in tip_words:
# print(word)
# ```
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# +
# [ ] review and run example
tip = "Notebooks can be exported as .pdf"
tip_words = tip.split()
print("STRING:", tip)
print("LIST:", tip_words, "\n")
for word in tip_words:
print(word)
# +
# [ ] review and run example
rhyme = "London bridge is falling down"
rhyme_words = rhyme.split()
rhyme_words.reverse()
for word in rhyme_words:
print(word)
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
#
# ### using `.split()`
# +
# [ ] split the string(rhyme) into a list of words (rhyme_words)
# [ ] print each word on it's own line
rhyme = 'Jack and Jill went up the hill To fetch a pail of water'
# +
# [ ] split code_tip into a list and print the first and every other word
code_tip = "Python uses spaces for indentation"
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## `.split('-')`
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/55d37e65-fb49-4bf5-87f8-fb987d3ce7a4/Unit2_Section3.4b-Split_on_Strings.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/55d37e65-fb49-4bf5-87f8-fb987d3ce7a4/Unit2_Section3.4b-Split_on_Strings.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### to split on characters other than " " (space), provide `.split()` a string argument to use as break points
# ```python
# code_tip = "Python-uses-spaces-for-indentation"
# tip_words = code_tip.split('-')
# ```
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# ### `.split('-') : split with an argument
# +
# [ ] review and run example
code_tip = "Python-uses-spaces-for-indentation"
tip_words = code_tip.split('-')
print(tip_words)
# +
# [ ] review and run example - study the list print output
code_tip = "Python uses spaces for indentation"
# split on "a"
tip_words = code_tip.split('a')
print(code_tip)
print(tip_words)
# +
# [ ] review and run example
# triple quotes ''' ''' preserve formatting such as spaces and line breaks
big_quote = """Jack and Jill went up the hill
To fetch a pail of water
Jack fell down and broke his crown
And Jill came tumbling after"""
# split on line breaks (\n)
quote_lines = big_quote.split('\n')
print(quote_lines, '\n')
# print the list in reverse with index slicing
for line in quote_lines[::-1]:
print(line)
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
#
# ## `.split()`
# [ ] split poem into a list of phrases by splitting on "*" a
# [ ] print each phrase on a new line in title case
poem = "Write code frequently*Save code frequently*Comment code frequently*Study code frequently*"
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
#
# ## `.join()` build a string from a list
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/57decc97-801f-4f7e-8ab6-69a47cf1be7b/Unit2_Section3.4c-Build_using_Join_Sequence.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/57decc97-801f-4f7e-8ab6-69a47cf1be7b/Unit2_Section3.4c-Build_using_Join_Sequence.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### `.join()` is a method applied to a separator string and iterates through its argument
# ```python
# tip_words = ['Notebooks', 'can', 'be', 'exported', 'as', '.pdf']
#
# " ".join(tip_words)
# ```
# a space (" ") is the separator that gets injected between the objects in the argument (the list "tip_words")
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# ## `.join()`
# +
# [ ] review and run example
tip_words = ['Notebooks', 'can', 'be', 'exported', 'as', '.pdf']
# join tip_words objects with spaces
print(" ".join(tip_words))
# -
# [ ] review and run example
no_space = ""
letters = ["P", "y", "t", "h", "o", "n"]
print(no_space.join(letters))
# +
# [ ] review and run example - .join() iterates through sequences
dash = "-"
space = " "
word = "Iteration"
ellipises = "..."
dash_join = dash.join(word)
print(dash_join)
print(space.join(word))
print(ellipises.join(word))
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
# ## `.join()`
# +
# [ ] .join() letters list objects with an Asterisk: "*"
letters = ["A", "s", "t", "e", "r", "i", "s", "k"]
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 4</B></font>
# ## Program: Choose the separator
# - get user input on what to use to join words (" ", *, -, etc...) - store in variable: separator
# - join pharse_words with the separator and print
# +
# [ ] complete Choose the separator
phrase_words = ['Jack', 'and', 'Jill', 'went', 'up', 'the', 'hill', 'To', 'fetch', 'a', 'pail', 'of', 'water']
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concept</B></font>
# ## More Python string tools (tricks?)
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/aa6eaea3-a3cb-41d8-aee3-280d01a9f4f0/Unit2_Section3.4d-Useful_String_Tricks.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/aa6eaea3-a3cb-41d8-aee3-280d01a9f4f0/Unit2_Section3.4d-Useful_String_Tricks.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### Cast a string to a list of characters
# ```python
# hello_letters = list("Hello")
# ```
# ### print to the same line with multiple print statements (`end=`)
# or insert any character as an end in print("String", end="+")
# ```python
# print('Hello', end = '')
# print('world')
# ```
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# [ ] review and run example
hello_letters = list("Hello")
print(hello_letters)
# +
# [ ] review and run example
# cast sting to list
word_letters = list("concatenates")
# .join() concatenates the list
# print on same line setting the end character
print('~'.join(word_letters))
# -
# [ ] review and run example
print("Hello ", end = '')
print("world")
# [ ] review and run example
# This is the default print end
print("Hello World!", end="\n")
print('still something to learn about print()')
# [ ] review and run example
# end inserts any valid str character: A-z, 0-9,!,@,*,\n,\t or ''(empty string)...
for letter in "Concatenation":
print(letter, end='*')
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 5</B></font>
# ## `end=" " ` configuration in printing
# `print('The String', end='')`
# +
# [ ] use 3 print() statements to output text to one line
# [ ] separate the lines by using "- " (dash space)
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 6</B></font>
# ## cast: str to list
# `Msg_characters = list("Always test your code")`
# +
# [ ] create a string (fact) of 20 or more characters and cast to a list (fact_letters)
# [ ] iterate fact, printing each char on one line, except for spaces print a new line
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 7</B></font>
# ## Program: add the digits
# - create a 20 digit string, and cast to a list
# - then add all the digits as integers
# - print the equation and answer
#
# Hint: use cast to sum the digits, and .join() to create the equation (1+2+3+...)
# +
# [ ] create add the digits
# -
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
|
Python Fundamentals/Module_3_4_Python_Fundamentals.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Depth Map from Stereo Images
#
# _You can view [IPython Nootebook](README.ipynb) report._
#
# ----
#
# ## Contents
#
# - [GOAL](#GOAL)
# - [Basics](#Basics)
# - [Code](#Code)
# - [Exercises](#Exercises)
#
# ## GOAL
#
# In this session:
#
# - We will learn to create a depth map from stereo images.
#
# ## Basics
#
# In the last session, we saw basic concepts like epipolar constraints and other related terms. We also saw that if we have two images of same scene, we can get depth information from that in an intuitive way. Below is an image and some simple mathematical formulas which prove that intuition.
#
# 
#
# The above diagram contains equivalent triangles. Writing their equivalent equations will yield us following result:
#
# $$
# disparity = x - x' = \frac{Bf}{Z},
# $$
#
# $ x $ and $ x′ $ are the distance between points in image plane corresponding to the scene point 3D and their camera center. $ B $ is the distance between two cameras (which we know) and $ f $ is the focal length of camera (already known). So in short, the above equation says that the depth of a point in a scene is inversely proportional to the difference in distance of corresponding image points and their camera centers. So with this information, we can derive the depth of all pixels in an image.
#
# So it finds corresponding matches between two images. We have already seen how epiline constraint make this operation faster and accurate. Once it finds matches, it finds the disparity. Let's see how we can do it with OpenCV.
#
# ## Code
#
# Below code snippet shows a simple procedure to create a disparity map.
#
# ```python
# import cv2 as cv
# from matplotlib import pyplot as plt
#
# imgL = cv.imread("../../data/tsukuba_l.png", 0)
# imgR = cv.imread("../../data/tsukuba_r.png", 0)
#
# stereo = cv.StereoBM_create(numDisparities=16, blockSize=15)
# disparity = stereo.compute(imgL, imgR)
#
# # cv.imwrite("output-files/depth-map-res.png", disparity)
# plt.subplot(121), plt.imshow(imgL), plt.title("Image 1 - Original image")
# plt.xticks([]), plt.yticks([])
# plt.subplot(122), plt.imshow(disparity), plt.title("Image 2 - Disparity map")
# plt.xticks([]), plt.yticks([])
# plt.subplots_adjust(left=0.01, right=0.99, wspace=0.02)
# plt.show()
# ```
#
# Below image contains the original image (left) and its disparity map (right). As you can see, the result is contaminated with high degree of noise. By adjusting the values of numDisparities and blockSize, you can get a better result.
#
# 
#
# ### Note
#
# > More details to be added
#
# ## Exercises
#
# 1. OpenCV samples contain an example of generating disparity map and its 3D reconstruction. Check stereo_match.py in OpenCV-Python samples.
|
calibration-reconstruction/depth-map/README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="hQnUF4trhLQ4" colab_type="text"
# # Cliff World
#
# In this notebook, we will see the difference of using Q-learning and SARSA is solving the famous Cliff World problem.
#
# First, we have to make sure we are connected to the right **python 3 reutime and using the GPU**. (Click the 'Runtime' tab and choose 'Change runtime type'), then import the required package (all are already installed in Google Colab)
#
# The policy we're gonna use is epsilon-greedy policy, where agent takes optimal action with probability $(1-\epsilon)$, otherwise samples action at random. Note that agent __can__ occasionally sample optimal action during random sampling by pure chance.
# + id="iwGPAw-FgnB3" colab_type="code" colab={}
#Setting up the enviroment, ignore the warning
import os
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY"))==0:
# !bash ../xvfb start
# %env DISPLAY=:1
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from collections import defaultdict
import random, math
# + id="oscIYMvtgnB9" colab_type="code" colab={}
import gym, gym.envs.toy_text
env = gym.envs.toy_text.CliffWalkingEnv()
n_actions = env.action_space.n
print(env.__doc__)
# + id="mgdNdxFIgnB_" colab_type="code" colab={}
# Our cliffworld has one difference from what's on the image: there is no wall.
# Agent can choose to go as close to the cliff as it wishes. x:start, T:exit, C:cliff, o: flat ground
env.render()
# + [markdown] id="mLgCHNOEjNXb" colab_type="text"
# ## Task 1: Imprement Q-learning Agent
# + id="e7BUQS-yu9hD" colab_type="code" colab={}
class QLearningAgent:
def __init__(self, alpha, epsilon, discount, get_legal_actions):
"""
Q-Learning Agent
based on http://inst.eecs.berkeley.edu/~cs188/sp09/pacman.html
Instance variables you have access to
- self.epsilon (exploration prob)
- self.alpha (learning rate)
- self.discount (discount rate aka gamma)
Functions you should use
- self.get_legal_actions(state) {state, hashable -> list of actions, each is hashable}
which returns legal actions for a state
- self.get_qvalue(state,action)
which returns Q(state,action)
- self.set_qvalue(state,action,value)
which sets Q(state,action) := value
!!!Important!!!
Note: please avoid using self._qValues directly.
There's a special self.get_qvalue/set_qvalue for that.
"""
self.get_legal_actions = get_legal_actions
self._qvalues = defaultdict(lambda: defaultdict(lambda: 0))
self.alpha = alpha
self.epsilon = epsilon
self.discount = discount
def get_qvalue(self, state, action):
""" Returns Q(state,action) """
return self._qvalues[state][action]
def set_qvalue(self,state,action,value):
""" Sets the Qvalue for [state,action] to the given value """
self._qvalues[state][action] = value
#---------------------START OF YOUR CODE---------------------#
def get_value(self, state):
"""
Compute your agent's estimate of V(s) using current q-values
V(s) = max_over_action Q(state,action) over possible actions.
Note: please take into account that q-values can be negative.
"""
possible_actions = self.get_legal_actions(state)
#If there are no legal actions, return 0.0
<your_code>
return value
def update(self, state, action, reward, next_state):
"""
You should do your Q-Value update here:
Q(s,a) := (1 - alpha) * Q(s,a) + alpha * (r + gamma * V(s'))
"""
#agent parameters
gamma = self.discount
learning_rate = self.alpha
new_Q = <your_code>
self.set_qvalue(state, action, new_Q)
def get_best_action(self, state):
"""
Compute the best action to take in a state (using current q-values).
"""
possible_actions = self.get_legal_actions(state)
#If there are no legal actions, return None
<your_code>
return best_action
def get_action(self, state):
"""
Compute the action to take in the current state, including exploration.
With probability self.epsilon, we should take a random action.
otherwise - the best policy action (self.getPolicy).
Note: To pick randomly from a list, use random.choice(list).
To pick True or False with a given probablity, generate uniform number in [0, 1]
and compare it with your probability
"""
# Pick Action
<your_code>
#If there are no legal actions, return None
<your_code>
#agent parameters:
epsilon = self.epsilon
<your_code>
return chosen_action
# + [markdown] id="Lmi9Up-ljUWT" colab_type="text"
# ## Task 2: Imprement SARSA Agent
# + id="JT56U9OognB6" colab_type="code" colab={}
class EVSarsaAgent(QLearningAgent):
"""
An agent that changes some of q-learning functions to implement Expected Value SARSA.
Note: this demo assumes that your implementation of QLearningAgent.update uses get_value(next_state).
If it doesn't, please add
def update(self, state, action, reward, next_state):
and implement it for Expected Value SARSA's V(s')
"""
def get_value(self, state):
"""
Returns Vpi for current state under epsilon-greedy policy:
V_{pi}(s) = sum _{over a_i} {pi(a_i | s) * Q(s, a_i)}
Hint: all other methods from QLearningAgent are still accessible.
"""
epsilon = self.epsilon
possible_actions = self.get_legal_actions(state)
#If there are no legal actions, return 0.0
<your_code>
return state_value
# + [markdown] colab_type="text" id="4Okx7UNsi0Ft"
# ## Play and train and evaluate
#
# Let's now see how our algorithm compares against q-learning in case where we force agent to explore all the time.
#
# <img src=https://github.com/yandexdataschool/Practical_RL/raw/master/yet_another_week/_resource/cliffworld.png width=600>
# <center><i>image by cs188</i></center>
# + id="gOMwfhR9gnCC" colab_type="code" colab={}
def play_and_train(env,agent,t_max=10**4):
"""This function should
- run a full game, actions given by agent.getAction(s)
- train agent using agent.update(...) whenever possible
- return total reward"""
total_reward = 0.0
s = env.reset()
for t in range(t_max):
a = agent.get_action(s)
next_s,r,done,_ = env.step(a)
agent.update(s, a, r, next_s)
s = next_s
total_reward +=r
if done:break
return total_reward
# + id="wEvMCIeVgnCE" colab_type="code" colab={}
agent_sarsa = EVSarsaAgent(alpha=0.25, epsilon=0.2, discount=0.99,
get_legal_actions = lambda s: range(n_actions))
agent_ql = QLearningAgent(alpha=0.25, epsilon=0.2, discount=0.99,
get_legal_actions = lambda s: range(n_actions))
# + id="2hKgwv0ygnCG" colab_type="code" colab={}
from IPython.display import clear_output
from pandas import DataFrame
moving_average = lambda x, span=100: DataFrame({'x':np.asarray(x)}).x.ewm(span=span).mean().values
rewards_sarsa, rewards_ql = [], []
for i in range(5000):
rewards_sarsa.append(play_and_train(env, agent_sarsa))
rewards_ql.append(play_and_train(env, agent_ql))
#Note: agent.epsilon stays constant
if i %100 ==0:
clear_output(True)
print('EVSARSA mean reward =', np.mean(rewards_sarsa[-100:]))
print('QLEARNING mean reward =', np.mean(rewards_ql[-100:]))
plt.title("epsilon = %s" % agent_ql.epsilon)
plt.plot(moving_average(rewards_sarsa), label='ev_sarsa')
plt.plot(moving_average(rewards_ql), label='qlearning')
plt.grid()
plt.legend()
plt.ylim(-500, 0)
plt.show()
# + [markdown] id="qzx2xl0XgnCI" colab_type="text"
# Let's now see what did the algorithms learn by visualizing their actions at every state.
# + id="_v8wkDoagnCJ" colab_type="code" colab={}
def draw_policy(env, agent):
""" Prints CliffWalkingEnv policy with arrows. Hard-coded. """
n_rows, n_cols = env._cliff.shape
actions = '^>v<'
for yi in range(n_rows):
for xi in range(n_cols):
if env._cliff[yi, xi]:
print(" C ", end='')
elif (yi * n_cols + xi) == env.start_state_index:
print(" X ", end='')
elif (yi * n_cols + xi) == n_rows * n_cols - 1:
print(" T ", end='')
else:
print(" %s " % actions[agent.get_best_action(yi * n_cols + xi)], end='')
print()
# + id="K6w0605JgnCL" colab_type="code" colab={}
print("Q-Learning")
draw_policy(env, agent_ql)
print("SARSA")
draw_policy(env, agent_sarsa)
# + [markdown] id="VYn6MsaHkFoa" colab_type="text"
# As you can see, the SARSA agent will avoid the gutter and Q-learning will not.
# + [markdown] id="JjR_qJa0gnCR" colab_type="text"
# ## Wanting More?
#
# Here are some of the things you can do if you feel like it:
#
# * Play with epsilon. See learned how policies change if you set epsilon to higher/lower values (e.g. 0.75).
# * Expected Value SASRSA for softmax policy:
# $$ \pi(a_i|s) = softmax({Q(s,a_i) \over \tau}) = {e ^ {Q(s,a_i)/ \tau} \over {\sum_{a_j} e ^{Q(s,a_j) / \tau }}} $$
# * Implement N-step algorithms and TD($\lambda$): see [Sutton's book](http://incompleteideas.net/book/bookdraft2018jan1.pdf) chapter 7 and chapter 12.
# * Use those algorithms to train on CartPole in previous / next assignment for this week.
|
exercises/rl_workshop_cliff_world.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Markdown
#
# Per formattare il testo, Jupyter mette a disposizione un linguaggio chiamato Markdown. Perchè dovresti imparare Markdown? E' semplice, molto popolare ed è probabile ritrovarlo in molti posti (blog, sistemi di documentazione tecnica, etc).
#
# Qua riportiamo solo informazioni essenziali, per altre puoi consultare la [Guida di Jupyter (inglese)](http://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html)
#
# ## Celle Markdown
#
# Per dire a Jupyter che una cella è codice Markdown e non Python, dal menu seleziona `Cell->Cell type->Markdown`. Una shortcut veloce è premere `Esc` seguito poi da il tasto `m`
#
# ## Paragrafi
#
# Per suddividere paragrafi basta inserire una riga vuota:
#
# Per esempio, scrivendo così:
#
# ```
# Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua. Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris nisi ut aliquip ex ea commodo consequat.
#
# Duis aute irure dolor in reprehenderit in voluptate velit esse cillum dolore eu fugiat nulla pariatur. Excepteur sint occaecat cupidatat non proident, sunt in culpa qui officia deserunt mollit anim id est laborum.
# ```
#
# Si ottiene questo:
#
#
# Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua. Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris nisi ut aliquip ex ea commodo consequat.
#
# Duis aute irure dolor in reprehenderit in voluptate velit esse cillum dolore eu fugiat nulla pariatur. Excepteur sint occaecat cupidatat non proident, sunt in culpa qui officia deserunt mollit anim id est laborum.
#
# ## Titoli
# I titoli ( _headers_ ) si scrivono anteponendo dei cancelletti al titolo stesso:
#
# ```
#
# # Titolo pagina
#
# E' importante.
#
# ## Primo capitolo
#
# Ma perchè?
#
# ### Primo paragrafo
#
# Che ne so.
#
# ### Secondo paragrafo
#
# E spiegalo !
#
#
# ## Secondo capitolo
#
# Taci e scrivi.
# ```
#
# <h1> Titolo pagina </h1>
#
# E' importante.
#
# ## Primo capitolo
#
# Ma perchè?
#
# ### Primo paragrafo
#
# Che ne so.
#
# ### Secondo paragrafo
#
# E spiegalo !
#
#
# ## Secondo capitolo
#
# Taci e scrivi.
#
#
# ## Link
#
# Un link si scrive mettendo il testo visibile dall'utente tra parentesi quadre e l'indirizzo vero e proprio tra parentesi tonde:
#
# ```
# [Questo è il testo del link](http://www.google.com)
# ```
#
# Il risultato sarà il seguente:
#
# [Questo è il testo del link](http://www.google.com)
# ## Liste
#
# Le liste si scrivono anteponendo ad ogni elemento un asterisco:
#
# Per esempio, scrivendo questo:
#
# ```
# * Elemento 1
# * Elemento 2
# * Sotto elemento
# * Altro sotto elemento
# * Elemento 3
# ```
#
# Verrà visualizzato così:
#
# * Elemento 1
# * Elemento 2
# * Sotto elemento
# * Altro sotto elemento
# * Elemento 3
#
# Potete anche usare liste numerate anteponendo agli elementi un numero e un punto.
#
# Per esempio, questo:
#
# ```
# 1. Elemento 1
# 2. Elemento 2
# 1. Sotto elemento
# 2. Altro sotto elemento
# 3. Elemento 3
#
# ```
#
# Verrà visualizzato così:
#
# 1. Elemento 1
# 2. Elemento 2
# 1. Sotto elemento
# 2. Altro sotto elemento
# 3. Elemento 3
#
#
# Se la lista è lunga e dovete editarla spesso, invece di numeri espliciti conviene mettere sempre `1.` e Markdown automaticamente calcolerà i numeri corretti:
#
# ```
# 1. Elemento 1
# 1. Elemento 2
# 1. Sotto elemento
# 1. Altro sotto elemento
# 1. Elemento 3
# ```
#
# verrà visualizzato così:
#
# 1. Elemento 1
# 1. Elemento 2
# 1. Sotto elemento
# 1. Altro sotto elemento
# 1. Elemento 3
#
# ## Immagini
#
# Purtroppo Jupyter non supporta il copia e incolla di immagini, puoi solo inserire dei link alle immagini stesse da mettere nella stessa cartella del notebook oppure in una sottocartella.
#
# Una volta che la cella viene eseguita, se il percorso al file è corretto apparirà l'immagine. Per indicare il percorso, scrivi punto esclamativo, parentesi quadre aperta-chiusa, e poi tra parentesi tonde il percorso del file (possibilmente usando lo slash `'/'`), per es se l'immagine `notebook_icon.png` sta nella sotto-cartella `img`, scrivi così:
#
# ```
# 
# ```
#
# Eseguendo la cella, apparirà l'immagine:
#
# 
#
# Nota che puoi usare qualunque formato di immagine (`jpg`, `png`, `bmp`, `svg`, per gli altri prova a vedere se vengono visualizzati).
#
#
# ## Variabili e nomi tecnici
#
# Per visualizzare in evidenza variabili e nomi tecnici, come `x`, `faiQualcosa`, `percorso-di-file`, puoi includere il nome tra due cosiddetti backtick \`
#
# **NOTA**: il backtick \` NON è l'apice che usiamo di solito: `'`
#
# Per scrivere questo strano apice rovesciato, guarda qua, se non va fai copia e incolla !
#
# * Windows:
# - se hai il tastierino numerico: tenere premuto `Alt Gr`, scrivere `96` sul tastierino numerico, rilasciare `Alt Gr`
# - se non ce l'hai: prova a premere tasto windows + `\` (in alto a sinistra)
#
# * Mac: `alt+9`
# * Linux: `Alt-Gr` + carattere apice normale `'`
#
# ## Codice JSON / XML / Python
#
# Se in una cella Markdown vuoi visualizzare testo posizionato esattamente come lo scrivi, racchiudilo in un blocco delimitato da file di tre tre backtick \`\`\` :
#
# \`\`\`
# ```
# testo posizionato come
# voglio
# io
# ```
# \`\`\`
#
#
# Risultato:
#
# ```
#
# testo posizionato come
# voglio
# io
#
# ```
#
# Il codice python / json / xml e altri possono essere formattati automaticamente da Jupyter. Basta scriveterlo in blocchi da tre backtick come prima e in più specificare il linguaggio subito dopo i primi tre backtick, per esempio un json scritto così:
#
# \`\`\`json
# ```
# {
# "a" : ["b"],
# "c" : {
# "d":5
# }
#
# }
#
# ```
# \`\`\`
#
# Risulterà formattato in questo modo:
#
# ```json
# {
# "a" : ["b"],
# "c" : {
# "d":5
# }
#
# }
# ```
#
# ## Formule matematiche
#
# E' possibile scrivere formule in [Latex](http://www.lorenzopantieri.net/LaTeX_files/ArteLaTeX.pdf#chapter.5) (ma non è trattato in questo libro) mettendole tra segni di dollaro `$`. Per esempio `$\sum$` verrà visualizzato così:
#
# $\sum$
#
# Con due dollari ai lati `$$\sum$$` la formula viene centrata:
#
# $$\sum$$
#
# ## Tabelle
#
# Scrivere tabelle piccole in Markdown è ancora fattibile:
#
# Per esempio, scrivere questo:
#
# ```
# io |sono | una tabella
# ---|-----|------------
# 4|ciao|3
# 2|hello world|7
# ```
#
# risulta visualizzato così :
#
# io |sono | una tabella
# ---|-----|------------
# 4|ciao|3
# 2|hello world|7
#
#
# ma per tabelle grandi Markdown è terribile. Quindi si è più che giustificati a usare alternative, per esempio allegare fogli Excel. Si può anche prendere screenshot delle tabelle e includerli come immagini.
|
project/markdown.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Amazon Personalize Workshop Part 2 - View Campaign and Interactions
# > In the first part, you successfully built and deployed a recommendation model using deep learning with Amazon Personalize. This notebook will expand on that and will walk you through adding the ability to react to real time behavior of users. If their intent changes while browsing a movie, you will see revised recommendations based on that behavior. It will also showcase demo code for simulating user behavior selecting movies before the recommendations are returned.
# - toc: true
# - badges: true
# - comments: true
# - categories: [amazonpersonalize, movie, realtime]
# - image:
# Below we start with just importing libraries that we need to interact with Personalize
# Imports
import boto3
import json
import numpy as np
import pandas as pd
import time
import uuid
# The line below will retrieve your shared variables from the first notebook.
# %store -r
# +
# Setup and Config
# Recommendations from Event data
personalize = boto3.client('personalize')
personalize_runtime = boto3.client('personalize-runtime')
# Establish a connection to Personalize's Event Streaming
personalize_events = boto3.client(service_name='personalize-events')
# -
# ## Creating an Event Tracker
#
# Before your recommendation system can respond to real time events you will need an event tracker, the code below will generate one and can be used going forward with this lab. Feel free to name it something more clever.
response = personalize.create_event_tracker(
name='MovieClickTracker',
datasetGroupArn=dataset_group_arn
)
print(response['eventTrackerArn'])
print(response['trackingId'])
TRACKING_ID = response['trackingId']
event_tracker_arn = response['eventTrackerArn']
# ## Configuring Source Data
#
# Above you'll see your tracking ID and this has been assigned to a variable so no further action is needed by you. The lines below are going to setup the data used for recommendations so you can render the list of movies later.
# +
# First load items into memory
items = pd.read_csv('./ml-100k/u.item', sep='|', usecols=[0,1], encoding='latin-1', names=['ITEM_ID', 'TITLE'], index_col='ITEM_ID')
def get_movie_title(movie_id):
"""
Takes in an ID, returns a title
"""
movie_id = int(movie_id)-1
return items.loc[movie_id]['TITLE']
# -
# ## Getting Recommendations
#
# First we will render the recommendations again from the previous notebook:
recommendations_df
# ## Simulating User Behavior
#
# The lines below provide a code sample that simulates a user interacting with a particular item, you will then get recommendations that differ from those when you started.
session_dict = {}
def send_movie_click(USER_ID, ITEM_ID):
"""
Simulates a click as an event
to send an event to Amazon Personalize's Event Tracker
"""
# Configure Session
try:
session_ID = session_dict[USER_ID]
except:
session_dict[USER_ID] = str(uuid.uuid1())
session_ID = session_dict[USER_ID]
# Configure Properties:
event = {
"itemId": str(ITEM_ID),
}
event_json = json.dumps(event)
# Make Call
personalize_events.put_events(
trackingId = TRACKING_ID,
userId= USER_ID,
sessionId = session_ID,
eventList = [{
'sentAt': int(time.time()),
'eventType': 'EVENT_TYPE',
'properties': event_json
}]
)
# Immediately below this line will update the tracker as if the user has clicked a particular title.
#
#
# If the table generated by the cells below does not shift the recommendations simply try another random 3 digit number in the cell above and run both cells again. You'll see a third column generated of recommendations.
# Pick a movie, we will use ID 270 or Gattaca
movie_to_click = 270
movie_title_clicked = get_movie_title(movie_to_click)
send_movie_click(USER_ID=str(user_id), ITEM_ID=movie_to_click)
# After executing this block you will see the alterations in the recommendations now that you have event tracking enabled and that you have sent the events to the service.
# +
get_recommendations_response = personalize_runtime.get_recommendations(
campaignArn = campaign_arn,
userId = str(user_id),
)
print("Recommendations for user: ", user_id)
item_list = get_recommendations_response['itemList']
recommendation_list = []
for item in item_list:
title = get_movie_title(item['itemId'])
recommendation_list.append(title)
new_rec_DF = pd.DataFrame(recommendation_list, columns = [movie_title_clicked])
recommendations_df = recommendations_df.join(new_rec_DF)
recommendations_df
# -
# ## Conclusion
#
# You can see now that recommendations are altered by changing the movie that a user interacts with, this system can be modified to any application where users are interacting with a collection of items. These tools are available at any time to pull down and start exploring what is possible with the data you have.
#
# Execute the cell below to store values needed for the cleanup notebook.
#
# Finally when you are ready to remove the items from your account, open the `Cleanup.ipynb` notebook and execute the steps there.
#
# %store event_tracker_arn
|
getting_started/notebooks/2021-06-20-amazon-personalize-workshop-part-2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Graphs using Matplotlib
# Matplotlib is a plotting library used for data visulalization.It helps to create graphs ,pie charts etc for better understanding of the pattern that data is reflecting.
# We will discuss two of its types:
# Point graph
# Line graph
import matplotlib.pyplot as plt
x = [1,2,3,4]
y = [1,4,9,16]
plt.scatter(x,y)
plt.show()
plt.scatter(x,y)
plt.plot(x,y)
plt.show()
# we can also use range function instead of a list
a = [i for i in range(25)]
b = [j*j for j in range(25) ]
# scatter is used for points only
plt.scatter(a,b)
# plot is used to get the line in the graph that joins the points
plt.plot(a,b)
# show returns the graph
plt.show()
# "r" is used to get the red color and "g" is for green color
plt.plot(a,b,'r')
plt.show()
plt.plot(a,b,'g')
plt.show()
# +
# "r*" is used to get the red star instead of red line("r-")
# similarly we can use "r" with any special type of point you need
plt.plot(a,b,'r*')
plt.show()
# -
# to get the blue circles
plt.plot(a,b,'bo')
plt.show()
plt.plot(a,b,'g--')
plt.show()
plt.plot(a,b,'r+')
plt.show()
plt.plot(a,b,'gp')
plt.show()
plt.plot(a,b,'r^')
plt.show()
plt.plot(a,b,'r-.')
plt.show()
c = [i for i in range(15)]
d = [j*5 for j in range(15)]
plt.scatter(c,d)
plt.plot(c,d,'g')
plt.show()
import numpy as np
x = np.array([1,2,3,4])
y = x**3
plt.plot(x,y,'b-.')
plt.show()
a = np.array([0,5,0.5])
b = a**3
plt.scatter(a,b)
plt.plot(a,b)
plt.show()
x = np.array([1,2,3,4])
y = x **3
plt.scatter(x,y)
plt.show()
x = np.arange(0,10,0.5)
y = x**3
plt.scatter(x,y)
plt.show()
x = np.arange(0,10,0.3)
y = x**3
plt.scatter(x,y)
plt.plot(x,y,'g')
plt.show()
a = [0,1,2,3,4]
plt.plot(a) #Only One array has been Given. What It does is that it takes an array By defalut
plt.show() #The array by default is x = [0,1,2,....]
# +
# Customizing Graphs
#To make our Grpahs Look Pretty
x = [i for i in range(10)]
y = [j*j for j in range(10)]
plt.plot(x,y,color='red')
plt.show()
# -
x = [i for i in range(10)]
y = [j*j for j in range(10)]
#plt.scatter(x,y)
plt.plot(x,y,color='red',marker='*')
plt.show()
x = np.arange(0,5,0.1)
y = x**3
plt.plot(x,y,color = "black",marker = "o")
plt.scatter(x,y)
plt.show()
x = [i for i in range(10)]
y = [j*j for j in range(10)]
plt.plot(x,y,color='red',linewidth=3)
plt.show()
x = [i for i in range(10)]
y = [j*j for j in range(10)]
plt.plot(x,y,color='red',linewidth=3)
# to give the label to x and y axis
plt.xlabel(" X ")
plt.ylabel(" X**2 ")
plt.title("Graph For Y = x**2")
plt.show()
x = [i for i in range(10)]
y = [j*j for j in range(10)]
a = [j*2 for j in range(10)]
b = [j*3 for j in range(10)]
plt.plot(x,y,color='red',linewidth=3,label='x^2')
plt.plot(x,a,color='blue',linewidth=3,label='x*2')
plt.plot(x,b,color='green',linewidth=3,label='x*3')
# legend is used to get the grid in graph with labels
plt.legend()
plt.show()
x = [1,2,3]
y = [4,5,6]
x1 = [8,7,6]
y1 = [5,6,7]
print(plt.plot(x,y),"r^")
print(plt.plot(x1,y1),"rp")
print(plt.scatter(x,y),"rp")
# +
# Pie Chart
sizes = [3,4,5,6]
# assigning colors
# colors = ["blue","red","yellow","green"]
# assigning title
plt.title("Split among classes")
# assigning labels
labels = ["a","b","c","d"]
# explode is used to highlight the edge
explode = [0.1,0,0,0]
plt.pie(sizes,colors = colors,labels = labels,explode = explode)
# use autopct = "%.2f" for percentage/decimal digits
# use autopct = "%.2f%%" gives percentage sign after digits
plt.axis("equal")
# to make pie chart clockwise we use:
counterclockwise = False in plt.pie(a)
plt.show()
# +
sizes = [3,4,6,2]
plt.pie(sizes)
plt.axis('equal')
plt.show()
# -
# to give specific color and labels
sizes = [3,4,6,2]
labels = ["A","B","C","D"]
colors = ["blue","red",'yellow',"green"]
plt.pie(sizes,colors = colors,labels = labels)
plt.axis("equal")
plt.show()
|
algorithms/ml/Graphs(Matplotlib)/Graphs(Matplotlib).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torchvision
import torch
import torchvision.transforms as T
from torchvision.utils import make_grid
import torch.nn.functional as F
import torch.nn as nn
from torch.utils.data.dataloader import DataLoader
from torch.utils.data import random_split
from pathlib import Path
# %matplotlib inline
# -
train = Path(r'C:\Users\USER-PC\Documents\Data Science\data set\New Plant Diseases Dataset(Augmented)\New Plant Diseases Dataset(Augmented)\train')
val =Path(r'C:\Users\USER-PC\Documents\Data Science\data set\New Plant Diseases Dataset(Augmented)\New Plant Diseases Dataset(Augmented)\valid')
from torchvision.datasets import ImageFolder
image_size = 32
stats = ((0.5, 0.45, 0.425), (0.35, 0.5, 0.253))
# dataset = ImageFolder(train, transform = T.Compose([T.CenterCrop(image_size), T.Resize(image_size),
# T.ToTensor(), T.Normalize(*stats)]))
# validation_set = ImageFolder(val, transform = T.Compose([T.CenterCrop(image_size), T.Resize(image_size),
# T.ToTensor(), T.Normalize(*stats)]))
dataset = ImageFolder(train, transform = T.Compose([T.Resize(image_size), T.ToTensor()]))
validation_set = ImageFolder(val, transform = T.Compose([T.Resize(image_size), T.ToTensor()]))
print(len(dataset.classes))
dataset.classes
img, label = dataset[0]
print("label", dataset.classes[label], ':', label)
print(img.shape)
plt.imshow(img.permute(1, 2, 0))
random_seed = 20
torch.manual_seed(random_seed)
batch_size = 250
len(dataset), len(validation_set)
train_loader = DataLoader(dataset, batch_size, shuffle= True, collate_fn=None, num_workers=4, pin_memory = True)
val_loader = DataLoader(validation_set, batch_size=batch_size*2, num_workers=4, collate_fn=None, pin_memory=True)
def showbatch(data):
for image, label in data:
fig, ax = plt.subplots(figsize = (16, 8))
plt.xticks([]); plt.yticks([])
plt.imshow(make_grid(image, nrow=16).permute(1, 2, 0))
break
showbatch(train_loader)
def accuracy(outputs, labels):
_, preds = torch.max(outputs, dim =1)
return torch.tensor(torch.sum(preds == labels).item() / len(preds))
class ImageClassificationBase(nn.Module):
def training_step(self, batch):
images, labels = batch
out = self(images)
loss = F.cross_entropy(out, labels)
return loss
def validation_step(self, batch):
images, labels = batch
out = self(images)
acc = accuracy(out, labels)
loss = F.cross_entropy(out, labels)
return {'val_loss': loss.detach(), 'val_acc': acc}
def validation_epoch_end(self, outputs):
batch_losses = [x['val_loss'] for x in outputs]
batch_accuracies = [x['val_acc'] for x in outputs]
epoch_loss = torch.stack(batch_losses).mean()
epoch_accuracy = torch.stack(batch_accuracies).mean()
return {'val_loss': epoch_loss.item(), 'val_acc': epoch_accuracy.item()}
def epoch_end(self, epoch, result):
print("Epoch[{}], val_loss: {:.4f}, val_acc: {:.4f}".format(epoch, result['val_loss'], result['val_acc']))
# +
def evaluate(model, val_loader):
outputs = [model.validation_step(batch) for batch in val_loader]
return model.validation_epoch_end(outputs)
def fit(epochs, lr, model, train_loader, val_loader, opt_func = torch.optim.SGD):
history = []
optimizer = opt_func(model.parameters(), lr)
for epoch in range(epochs):
for batch in train_loader:
loss = model.training_step(batch)
loss = loss.backward()
optimizer.step()
optimizer.zero_grad()
result = evaluate(model, val_loader)
model.epoch_end(epoch, result)
history.append(result)
return history
# -
torch.cuda.is_available()
def get_default_device():
"""Pick GPU if available, else CPU"""
if torch.cuda.is_available():
return torch.device('cuda')
else:
return torch.device('cpu')
device = get_default_device()
device
# +
def to_device(data, device):
"""Move tensor(s) to chosen device"""
if isinstance(data, (list,tuple)):
return [to_device(x, device) for x in data]
return data.to(device, non_blocking=True)
class DeviceDataLoader():
"""Wrap a dataloader to move data to a device"""
def __init__(self, dl, device):
self.dl = dl
self.device = device
def __iter__(self):
"""Yield a batch of data after moving it to device"""
for b in self.dl:
yield to_device(b, self.device)
def __len__(self):
"""Number of batches"""
return len(self.dl)
# -
def plot_losses(history):
losses = [x['val_loss'] for x in history]
plt.plot(losses, '-x')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.title("Loss Vs. No of epoochs")
def plot_accuracies(history):
accuracies = [x['val_acc'] for x in history]
plt.plot(accuracies, '-x')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.title("Accuracy Vs. No of epochs")
train_loader = DeviceDataLoader(train_loader, device)
val_loader = DeviceDataLoader(val_loader, device)
input_size = 3*256*256
hidden_size1 = 1024
hidden_size2 = 512
hidden_size3 = 256
hidden_size4 = 128
hidden_size5 = 8
output_size = 38
batch_size = 190
class PlantDiseaseModel(ImageClassificationBase):
def __init__(self, input_size, hidden_size1, hidden_size2, hidden_size3, hidden_size4, output_size):
super().__init__()
self.linear1 = nn.Linear(input_size, hidden_size1)
self.linear2 = nn.Linear(hidden_size1, hidden_size2)
self.linear3 = nn.Linear(hidden_size2, hidden_size3)
self.linear4 = nn.Linear(hidden_size3, hidden_size4)
self.linear5 = nn.Linear(hidden_size4, output_size)
def forward(self, xb):
out = xb.view(xb.size(0), -1)
out = self.linear1(out)
out = F.relu(out)
out = self.linear2(out)
out = F.relu(out)
out = self.linear3(out)
out = F.relu(out)
out = self.linear4(out)
out = F.relu(out)
out = self.linear5(out)
return out
model = to_device(PlantDiseaseModel(input_size, hidden_size1, hidden_size2, hidden_size3, hidden_size4, output_size), device)
history = [evaluate(model, val_loader)]
history
history += fit(10, 0.1, model, train_loader, val_loader)
plot_losses(history)
plot_accuracies(history)
def predict_image(img, model):
xb = to_device(img.unsqueeze(0), device)
yb = model(xb)
_, preds = torch.max(yb, dim = 1)
return dataset.classes[preds[0].item()]
img, label = train[100]
plt.imshow(img.permute(1, 2, 0))
print("Label: ", dataset.classes[label], ", Predicted: ", predict_image(img, model))
img, label = dataset[38000]
plt.imshow(img.permute(1, 2, 0))
print("Label", dataset.classes[label], ", Predicted", predict_image(img, model))
val_loader = DataLoader(val, batch_size*3, shuffle = True, num_workers = 3, collate_fn = None, pin_memory = True)
result = evaluate(model, val_loader)
result
|
6.0 Computer Vision/2.0 Tensorflow/Plant Disease2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 给出矩阵 matrix 和目标值 target,返回元素总和等于目标值的非空子矩阵的数量。
#
# 子矩阵 x1, y1, x2, y2 是满足 x1 <= x <= x2 且 y1 <= y <= y2 的所有单元 matrix[x][y] 的集合。
#
# 如果 (x1, y1, x2, y2) 和 (x1', y1', x2', y2') 两个子矩阵中部分坐标不同(如:x1 != x1'),那么这两个子矩阵也不同。
#
# 示例 1:
# 输入:matrix = [[0,1,0],
# [1,1,1],
# [0,1,0]], target = 0
# 输出:4
# 解释:四个只含 0 的 1x1 子矩阵。
#
# 示例 2:
# 输入:matrix = [[1,-1],
# [-1,1]], target = 0
# 输出:5
# 解释:两个 1x2 子矩阵,加上两个 2x1 子矩阵,再加上一个 2x2 子矩阵。
#
# 提示:
# 1、1 <= matrix.length <= 300
# 2、1 <= matrix[0].length <= 300
# 3、-1000 <= matrix[i] <= 1000
# 4、-10^8 <= target <= 10^8
# -
# <img src='https://assets.leetcode.com/uploads/2020/09/02/mate1.jpg'>
# +
from collections import defaultdict
class Solution:
def numSubmatrixSumTarget(self, matrix, target: int) -> int:
rows, cols = len(matrix), len(matrix[0])
def calc(arr):
cnt = 0
presum = 0
record = defaultdict(int)
record[0] = 1
for i in range(cols):
presum += arr[i]
if presum - target in record:
cnt += record[presum - target]
record[presum] += 1
return cnt
count = 0
for i in range(rows):
temp = [0] * cols
for j in range(i, rows):
for k in range(cols):
temp[k] += matrix[j][k]
# 寻找sub_matrix == target的值
print(temp, i, j)
count += calc(temp)
return count
# -
solution = Solution()
solution.numSubmatrixSumTarget([[0,1,0],
[1,1,1],
[0,1,0]], 0)
# +
from collections import defaultdict
target = 0
cols = 4
def calc(arr):
cnt = 0
presum = 0
record = defaultdict(int)
record[0] = 1
for i in range(cols):
presum += arr[i]
if presum - target in record:
print(presum, target, record)
cnt += record[presum - target]
record[presum] += 1
print(record)
return cnt
# -
calc([0, 1, 2, 0])
|
Dynamic Programming/1104/1074. Number of Submatrices That Sum to Target.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from sqlalchemy import create_engine
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from scipy.sparse import csr_matrix
from sklearn.externals import joblib
import datetime as dt
import pickle
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
warnings.simplefilter('ignore')
# # Data Pre-Processing
engine = create_engine("sqlite:///../db/twitter_db.sqlite")
conn = engine.connect()
tweets_df = pd.read_sql("SELECT * FROM tweet_data", conn)
tweets_df.head(2)
len(tweets_df)
tweets_df["created_at_datetime"] = tweets_df["created_at_datetime"].apply(lambda x: dt.datetime.strptime(x,
"%Y-%m-%d %H:%M:%S.%f"))
tweets_df = tweets_df.loc[tweets_df["created_at_datetime"] < dt.datetime(2019,8,15), :].reset_index(drop=True)
len(tweets_df)
# +
# Below, the number of candidates was reduced to reduce mdoel file size (6 to 5)
# -
grouped_df = tweets_df.groupby(tweets_df['user_name']).median()
grouped_df = grouped_df[['retweet_count']].sort_values('retweet_count', ascending = False).iloc[:6]
# grouped_df.sort_values('retweet_count', ascendingh = False)
grouped_df
top_candidates =[]
for i, r in grouped_df.iterrows():
top_candidates.append(i)
# Remove <NAME> from list (dropped out)
top_candidates.pop(3)
top_candidates
with open('top_candidates.pkl', 'wb') as f:
pickle.dump(top_candidates, f)
tweets_df = tweets_df.loc[tweets_df['user_name'].isin(top_candidates), :].reset_index(drop=True)
len(tweets_df)
tweets_df["day"] = tweets_df["created_at_datetime"].apply(lambda x: dt.datetime.strftime(x, "%A"))
tweets_df["hour"] = tweets_df["created_at_datetime"].apply(lambda x: dt.datetime.strftime(x, "%H"))
tweets_df["month"] = tweets_df["created_at_datetime"].apply(lambda x: dt.datetime.strftime(x, "%B"))
tweets_df.drop(columns = ["created_at", "created_at_time", "created_at_date","created_at_datetime",
"tweet_id", "tweet_id_str", "in_reply_to_status_id",
"in_reply_to_status_id_str", "in_reply_to_user_id",
"in_reply_to_user_id_str", "in_reply_to_screen_name",
"user_id_str", "user_id", "user_screen_name", "id"], inplace = True)
tweets_df = tweets_df[["user_name", "month", "day", "hour", "retweet_count", "favorite_count", "full_text"]]
tweets_df.head(2)
len(tweets_df.groupby(tweets_df["user_name"]).count())
X_count_df = tweets_df[["full_text", "month", "day", "hour", "retweet_count", "favorite_count"]]
X_count_df = pd.get_dummies(X_count_df, columns = ["month", "day", "hour"])
X_count_df.head(2)
# +
import nltk
import re
import string
pd.set_option('display.max_colwidth', 100) # To extend column width
stopwords = nltk.corpus.stopwords.words('english')
wn = nltk.WordNetLemmatizer()
# -
def clean_text(text):
text = text.replace('&', '&')
text = text.replace('\n', ' ')
text = "".join([word.lower() for word in text if word not in string.punctuation])
tokens = re.split('\W+', text)
text = [wn.lemmatize(word) for word in tokens if word not in stopwords]
return text
# # Bag of Words
# # Count Vectorizer
# +
from sklearn.feature_extraction.text import CountVectorizer
# CountVectorizer
count_vect = CountVectorizer(analyzer=clean_text)
X_count_vect = count_vect.fit_transform(X_count_df['full_text'])
# -
X_count_df.drop(columns = ['full_text'], inplace = True)
X_count_df = pd.concat([X_count_df, pd.DataFrame(X_count_vect.toarray(), columns=count_vect.get_feature_names())], axis=1)
X_count_df.shape
rf_columns = X_count_df.columns
rf_columns
rf_columns_list = list(rf_columns)
len(rf_columns)
with open('rf_columns.pkl', 'wb') as f:
pickle.dump(rf_columns_list, f)
data_y = tweets_df.values
data_x = X_count_df.values
X_count = data_x[:, 0:]
y_count = data_y[:,0]
X_count
X_count.shape
X_count_sparse = csr_matrix(X_count)
X_count_sparse
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MaxAbsScaler
from sklearn.preprocessing import LabelEncoder
from matplotlib.legend_handler import HandlerLine2D
# +
# from keras.utils import to_categorical
# -
# ## Train-Test-Split/Label-Encoding (Grid Search)
X_train_cv, X_test_cv, y_train_cv, y_test_cv = train_test_split(X_count_sparse, y_count, random_state=42,
train_size = 0.9,
shuffle = True)
X_scaler_cv = MaxAbsScaler().fit(X_train_cv)
scaler_filename = "rf_scaler.save"
joblib.dump(X_scaler_cv, scaler_filename)
X_train_scaled_cv = X_scaler_cv.transform(X_train_cv)
X_test_scaled_cv = X_scaler_cv.transform(X_test_cv)
# +
# Step 1: Label-encode data set
label_encoder_cv = LabelEncoder()
label_encoder_cv.fit(y_train_cv)
encoded_y_train_cv = label_encoder_cv.transform(y_train_cv)
encoded_y_test_cv = label_encoder_cv.transform(y_test_cv)
# -
encoded_y_train_cv
encoded_y_train_cv = encoded_y_train_cv.reshape(-1, 1)
encoded_y_test_cv = encoded_y_test_cv.reshape(-1, 1)
encoded_y_train_cv
label_encoder_cv.classes_
np.save('rf_classes.npy', label_encoder_cv.classes_)
# ## Train-Test-Split/Label Encoding (In-Depth Parameter Tuning)
X_train, X_test, y_train, y_test = train_test_split(X_count_sparse, y_count, random_state=42)
X_scaler = MaxAbsScaler().fit(X_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
X_train_scaled.shape
# +
# Step 1: Label-encode data set
label_encoder = LabelEncoder()
label_encoder.fit(y_train)
encoded_y_train = label_encoder.transform(y_train)
encoded_y_test = label_encoder.transform(y_test)
# -
encoded_y_train = encoded_y_train.reshape(-1, 1)
encoded_y_test = encoded_y_test.reshape(-1, 1)
# ## N_estimators (Count Vectorizer)
# +
n_estimators = [1, 2, 4, 8, 16, 32, 64, 100, 150, 200, 300]
train_results = []
test_results = []
for estimator in n_estimators:
rf = RandomForestClassifier(n_estimators=estimator, n_jobs=-1)
rf.fit(X_train_scaled, encoded_y_train)
train_results.append(rf.score(X_train_scaled, encoded_y_train))
test_results.append(rf.score(X_test_scaled, encoded_y_test))
# +
line1, = plt.plot(n_estimators, train_results, 'b', label="Train")
line2, = plt.plot(n_estimators, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('n_estimators')
plt.show()
# +
# Best ~ 200 (50 - 250)
# -
# ## Max-Depth (Count Vectorizer)
# +
max_depths = list(np.linspace(1, 90, 90, endpoint=True))
train_results = []
test_results = []
for max_depth in max_depths:
rf = RandomForestClassifier(max_depth=max_depth, n_jobs=-1)
rf.fit(X_train_scaled, encoded_y_train)
train_results.append(rf.score(X_train_scaled, encoded_y_train))
test_results.append(rf.score(X_test_scaled, encoded_y_test))
# +
line1, = plt.plot(max_depths, train_results, 'b', label="Train")
line2, = plt.plot(max_depths, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('max_depth')
plt.show()
# +
# Best ~ 20 (5 - 15)
# -
# ## Min_samples_split (Count Vectorizer)
# +
min_samples_splits = list(np.linspace(0.1, 1.0, 10, endpoint=True))
train_results = []
test_results = []
for min_samples_split in min_samples_splits:
rf = RandomForestClassifier(min_samples_split=min_samples_split, n_jobs=-1)
rf.fit(X_train_scaled, encoded_y_train)
train_results.append(rf.score(X_train_scaled, encoded_y_train))
test_results.append(rf.score(X_test_scaled, encoded_y_test))
# +
line1, = plt.plot(min_samples_splits, train_results, 'b', label="Train")
line2, = plt.plot(min_samples_splits, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('min_samples_split')
plt.show()
# +
#Best ~ 2
# -
# ## Min_samples_leaf (Count Vectorizer)
# +
min_samples_leafs = list(np.linspace(.01, 0.5, 20, endpoint = True))
train_results = []
test_results = []
for min_samples_leaf in min_samples_leafs:
rf = RandomForestClassifier(min_samples_leaf=min_samples_leaf, n_jobs=-1)
rf.fit(X_train_scaled, encoded_y_train)
train_results.append(rf.score(X_train_scaled, encoded_y_train))
test_results.append(rf.score(X_test_scaled, encoded_y_test))
# +
line1, = plt.plot(min_samples_leafs, train_results, 'b', label="Train")
line2, = plt.plot(min_samples_leafs, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('min_samples_leaf')
plt.show()
# -
# ## Max_features (Count Vectorizer)
# +
max_features_list = list(range(1, 3000, 100))
train_results = []
test_results = []
for max_features in max_features_list:
rf = RandomForestClassifier(max_features=max_features, n_jobs=-1)
rf.fit(X_train_scaled, encoded_y_train)
train_results.append(rf.score(X_train_scaled, encoded_y_train))
test_results.append(rf.score(X_test_scaled, encoded_y_test))
# +
line1, = plt.plot(max_features_list, train_results, 'b', label="Train")
line2, = plt.plot(max_features_list, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('max_features')
plt.show()
# -
# ## In-Depth Parameter Tuning Scoring (Count Vectorizer)
# Create a random forest classifier
rf = RandomForestClassifier(n_estimators=200, max_depth=20, min_samples_split = 2,
min_samples_leaf = 1, max_features = 150)
rf = rf.fit(X_train_scaled, encoded_y_train)
rf.score(X_test_scaled, encoded_y_test)
# ## GridSearch (Count Vectorizer)
max_depth_gs = list(range(5, 16, 2))
max_depth_gs.append(None)
max_depth_gs
# +
rf = RandomForestClassifier()
param = {'n_estimators': list(range(10, 151, 14)),
'max_depth': max_depth_gs,
'max_features': list(range(100, 301, 20))
}
gs = GridSearchCV(rf, param, cv=5, n_jobs=-1, verbose = 3)# n_jobs=-1 for parallelizing search
gs_fit = gs.fit(X_train_scaled_cv, encoded_y_train_cv)
pd.DataFrame(gs_fit.cv_results_).sort_values('mean_test_score', ascending=False).head()
# -
# Create a random forest classifier
rf_model = RandomForestClassifier(n_estimators=136, max_depth=None, min_samples_split = 2,
min_samples_leaf = 1, max_features = 120)
rf_model = rf_model.fit(X_train_scaled_cv, encoded_y_train_cv)
rf_model.score(X_test_scaled_cv, encoded_y_test_cv)
filename = "rf_model.sav"
with open(filename, 'wb') as f:
pickle.dump(rf_model, f)
rf_model.feature_importances_
sorted(zip(rf_model.feature_importances_, rf_columns_list), reverse=True)
# # TF-IDF Vectorizer
X_tdidf_df = tweets_df[["full_text", "month", "day", "hour", "retweet_count", "favorite_count"]]
X_tdidf_df = pd.get_dummies(X_tdidf_df, columns = ["month", "day", "hour"])
# +
from sklearn.feature_extraction.text import TfidfVectorizer
# TF-IDF
tfidf_vect = TfidfVectorizer(analyzer=clean_text)
X_tfidf_vect = tfidf_vect.fit_transform(X_tdidf_df['full_text'])
# -
X_tdidf_df.drop(columns = ['full_text'], inplace = True)
X_tdidf_df = pd.concat([X_tdidf_df, pd.DataFrame(X_tfidf_vect.toarray(), columns=tfidf_vect.get_feature_names())], axis=1)
X_tdidf_df.shape
data_y_idf = tweets_df.values
data_x_idf = X_tdidf_df.values
X_count_idf = data_x_idf[:, 0:]
y_count_idf = data_y_idf[:,0]
X_count_idf
X_count_idf.shape
y_count_idf
X_count_idf_sparse = csr_matrix(X_count_idf)
# ## Train-Test-Split/Label-Encoding (Grid Search)
#
X_train_idf_cv, X_test_idf_cv, y_train_idf_cv, y_test_idf_cv = train_test_split(X_count_idf_sparse,
y_count_idf, random_state=42,
train_size = 0.9,
shuffle = True)
X_scaler_idf_cv = MaxAbsScaler().fit(X_train_idf_cv)
X_train_scaled_idf_cv = X_scaler_idf_cv.transform(X_train_idf_cv)
X_test_scaled_idf_cv = X_scaler_idf_cv.transform(X_test_idf_cv)
min_samples_leafs = list(np.linspace(.01, 0.5, 20, endpoint = True))
train_results = []
test_results = []
for min_samples_leaf in min_samples_leafs:
rf = RandomForestClassifier(min_samples_leaf=min_samples_leaf, n_jobs=-1)
rf.fit(X_train_scaled, encoded_y_train)
train_results.append(rf.score(X_train_scaled, encoded_y_train))
test_results.append(rf.score(X_test_scaled, encoded_y_test))
# +
# Step 1: Label-encode data set
label_encoder_idf_cv = LabelEncoder()
label_encoder_idf_cv.fit(y_train_idf_cv)
encoded_y_train_idf_cv = label_encoder_idf_cv.transform(y_train_idf_cv)
encoded_y_test_idf_cv = label_encoder_idf_cv.transform(y_test_idf_cv)
# -
encoded_y_train_idf_cv = encoded_y_train_idf_cv.reshape(-1, 1)
encoded_y_test_idf_cv = encoded_y_test_idf_cv.reshape(-1, 1)
# # TF-IDF Vectorizer
# ## Train-Test-Split/Label-Encoding (In-Depth Parameter Tuning)
X_train_idf, X_test_idf, y_train_idf, y_test_idf = train_test_split(X_count_idf_sparse, y_count_idf, random_state=42,
shuffle = True)
X_scaler_idf = MaxAbsScaler().fit(X_train_idf)
X_train_scaled_idf = X_scaler_idf.transform(X_train_idf)
X_test_scaled_idf = X_scaler_idf.transform(X_test_idf)
# +
# Step 1: Label-encode data set
label_encoder_idf = LabelEncoder()
label_encoder_idf.fit(y_train_idf)
encoded_y_train_idf = label_encoder_idf.transform(y_train_idf)
encoded_y_test_idf = label_encoder_idf.transform(y_test_idf)
# -
encoded_y_train_idf = encoded_y_train_idf.reshape(-1, 1)
encoded_y_test_idf = encoded_y_test_idf.reshape(-1, 1)
# ## In-Depth Parameter Tuning
# ## N-Estimators (TF-IDF Vectorizer)
# +
n_estimators = [1, 2, 4, 8, 16, 32, 64, 100, 150, 200, 300]
train_results = []
test_results = []
for estimator in n_estimators:
rf = RandomForestClassifier(n_estimators=estimator, n_jobs=-1)
rf.fit(X_train_scaled_idf, encoded_y_train_idf)
train_results.append(rf.score(X_train_scaled_idf, encoded_y_train_idf))
test_results.append(rf.score(X_test_scaled_idf, encoded_y_test_idf))
# +
line1, = plt.plot(n_estimators, train_results, 'b', label="Train")
line2, = plt.plot(n_estimators, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('n_estimators')
plt.show()
# +
# Best: n-estimators=150 (10 - 250)
# -
# ## Max-Depth (TF-IDF Vectorizer)
max_depths = list(np.linspace(1, 90, 90, endpoint=True))
train_results = []
test_results = []
for max_depth in max_depths:
rf = RandomForestClassifier(max_depth=max_depth, n_jobs=-1)
rf.fit(X_train_scaled_idf, encoded_y_train_idf)
train_results.append(rf.score(X_train_scaled_idf, encoded_y_train_idf))
test_results.append(rf.score(X_test_scaled_idf, encoded_y_test_idf))
# +
line1, = plt.plot(max_depths, train_results, 'b', label="Train")
line2, = plt.plot(max_depths, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('max_depths')
plt.show()
# +
#Best: max_depth= 8 (2-12)
# -
# ## Min_samples_split (TF-IDF Vectorizer)
min_samples_splits = list(np.linspace(0.1, 1.0, 10, endpoint=True))
train_results = []
test_results = []
for min_samples_split in min_samples_splits:
rf = RandomForestClassifier(min_samples_split=min_samples_split, n_jobs=-1)
rf.fit(X_train_scaled_idf, encoded_y_train_idf)
train_results.append(rf.score(X_train_scaled_idf, encoded_y_train_idf))
test_results.append(rf.score(X_test_scaled_idf, encoded_y_test_idf))
# +
line1, = plt.plot(min_samples_splits, train_results, 'b', label="Train")
line2, = plt.plot(min_samples_splits, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('min_samples_split')
plt.show()
# +
# Best: min_samples_split=2
# -
# ## Min_samples_leaf (TF-IDF Vectorizer)
min_samples_leafs = list(np.linspace(.01, 0.5, 20, endpoint = True))
train_results = []
test_results = []
for min_samples_leaf in min_samples_leafs:
rf = RandomForestClassifier(min_samples_leaf=min_samples_leaf, n_jobs=-1)
rf.fit(X_train_scaled_idf, encoded_y_train_idf)
train_results.append(rf.score(X_train_scaled_idf, encoded_y_train_idf))
test_results.append(rf.score(X_test_scaled_idf, encoded_y_test_idf))
# +
line1, = plt.plot(min_samples_leafs, train_results, 'b', label="Train")
line2, = plt.plot(min_samples_leafs, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('min_samples_leaf')
plt.show()
# +
#Best: min_samples_leaf = 1
# -
# ## Max_Features (TF-IDF Vectorizer)
max_features_list = list(range(1, 3000, 100))
train_results = []
test_results = []
for max_features in max_features_list:
rf = RandomForestClassifier(max_features=max_features, n_jobs=-1)
rf.fit(X_train_scaled_idf, encoded_y_train_idf)
train_results.append(rf.score(X_train_scaled_idf, encoded_y_train_idf))
test_results.append(rf.score(X_test_scaled_idf, encoded_y_test_idf))
# +
line1, = plt.plot(max_features_list, train_results, 'b', label="Train")
line2, = plt.plot(max_features_list, test_results, 'r', label="Test")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel('RF score')
plt.xlabel('max_features')
plt.show()
# +
#Best: max_features=200 (100-300)
# -
# ## In-Depth Parameter Tuning Scoring (TF-IDF Vectorizer)
# Create a random forest classifier
rf = RandomForestClassifier(n_estimators=150, max_depth=8, min_samples_split = 2,
min_samples_leaf = 1, max_features = 250)
rf = rf.fit(X_train_scaled_idf, encoded_y_train_idf)
rf.score(X_test_scaled_idf, encoded_y_test_idf)
# ## Grid Search (TF-IDF Vectorizer)
max_depth_gs = list(range(2, 20, 2))
max_depth_gs.append(None)
max_depth_gs
# +
rf = RandomForestClassifier()
param = {'n_estimators': list(range(10, 251, 24)),
'max_depth': max_depth_gs,
'max_features': list(range(100, 301, 20))
}
gs = GridSearchCV(rf, param, cv=5, n_jobs=-1, verbose = 2)# n_jobs=-1 for parallelizing search
gs_fit = gs.fit(X_train_scaled_idf_cv, encoded_y_train_idf_cv)
pd.DataFrame(gs_fit.cv_results_).sort_values('mean_test_score', ascending=False).head()
# -
# Create a random forest classifier
rf = RandomForestClassifier(n_estimators=250, max_depth=None, min_samples_split = 2,
min_samples_leaf = 1, max_features = 100)
rf = rf.fit(X_train_scaled_idf_cv, encoded_y_train_idf_cv)
rf.score(X_test_scaled_idf_cv, encoded_y_test_idf_cv)
|
jupyter_notebook_code/Random Forest Classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="YwMU3rCeoB-j" colab_type="text"
# # **nnU-Net : The no-new-UNet for automatic segmentation**
#
#
# + [markdown] id="EUwUYn48C62N" colab_type="text"
# ## **Introduction**
#
# nnU-Net is an open-source tool that can effectively be used out-of-the-box, rendering state of the art segmentation and catalyzing scientific progress as a framework for automated method design. It provides an end-to-end automated pipeline, which can be trained and inferred on any medical dataset for segmentation.
#
# nnU-Net has outperformed state-of-the-art architecture(s) in the [Medical Decathlon Challenge](http://medicaldecathlon.com/),comprising of 10 different datasets using an ensemble of the same U-Net architecture with an automated pipeline comprising of pre-processing, data augmentation and post-processing. It has set a new benchmark in the field of medical image segmentation, without the need to fine tuning a new architecture for every dataset individually. The pipeline itself takes care of the hyper-parameter tuning and requires no change in the network architecture to achieve start-of-the-art results.
# + [markdown] id="rC2pvOFvCtjS" colab_type="text"
# ## **Motivation**
#
#
# * Working through nnU-net has it's own hassles - manually renaming the files, putting the files in the right location, saving the data in the correct format. Each of these steps are time consuming and susceptible to human errors. Hence, this tutorial is written in a manner that will allow you to avoid the common errors.
#
# * The paper detailing nnU-net has a long supplementary material for reference (55 pages research paper). We want to save your worthy time by summarizing the internal working of the nnU-net to provide better understanding for first time users (having series of automation steps, it could take quite sometime to understand the internal workings.)
#
# * We have tried to provide a single notebook to run through all the necessary steps as part of getting started with nnU-net. Though nnU-net has 3 different documentations detailing all the steps, our tutorial gives brief description of the parts relevant to users who just want to get started.
#
#
#
#
#
#
#
#
# ## **Problem Statement**
#
#
# Though, nnU-Net takes care of most of the steps involved itself, sometimes getting it up and running for a custom dataset which is not in format mentioned [here](https://github.com/prateekgupta891/nnUNet/blob/master/documentation/setting_up_paths.md) is difficult. Training the system on a colab notebook is again a challenge, as it requires a lot of manual steps. This tutorial will run you through all the necessary steps required to train your first nnU-net model along with ways to ensure the correctness of the procedure.
#
# ## **Overview of this tutorial :**
#
# 1. Introduces the nnU-Net framework
# 2. One-stop tutorial to train and test with different datasets
# 3. Colab Notebook Based Tutorial (will work for local machines too).
# 4. Folder creation, command execution from particular folder location done by the code.
# 5. How to ingest the SCGM Challenge Dataset to the nnUNet pipeline.
# 6. Data Visualization for Train, Test and Prediction
# + [markdown] id="RIDI-C7sTbRZ" colab_type="text"
# ## **nnUNet Complete Work Flow**
# + [markdown] id="1wtzpPnjS7n5" colab_type="text"
# 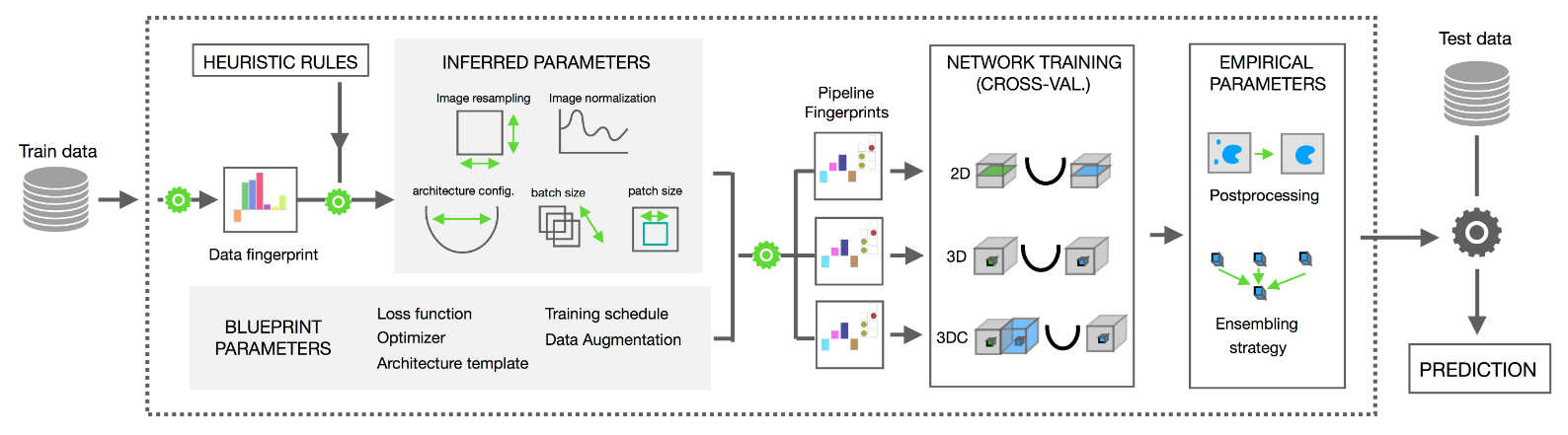
#
#
# The nnU-Net pipeline uses heuristic rule to determine the data-dependent hyperparamters, known as the **"data fingerprint"**, to ingest the training data. The **blueprint parameters** ( loss function, optimizer,architecture) and **inferred parameters** ( image resampling, normalization, batch and patch size) along with the data fingerprint generate pipeline fingerprints. **Pipeline fingerprints** produce network training for 2D, 3D and 3D-Cascade U-Net using the hyperparameters determined so far. The **ensemble** of different network configuration(s), along with post-processing determines the best average Dice coefficient for the training data. The **best configuration** will then be used to produce the **predictions** for the test data. Details about individual component are described below.
# + [markdown] id="ZwjffSPGofJb" colab_type="text"
# ### **1. Dataset Fingerprint**
#
# It contains a set of heuristic rules to infer data-dependent hyper-parameters of the pipeline.
#
#
# * Image size (i.e. number of voxels per spatial dimension) before and after cropping image
# * Image Spacing (i.e. physical size of the voxels)
# * Modalities (from metadata)
# * Number of classes for all images and total number of training cases
#
# + [markdown] id="QQcVo2YrZorQ" colab_type="text"
# ### **2. Blueprint Parameters**
#
# **a. Architecture Template**
#
# * nnU-Net architecture closely follows the original U-Net and has recently proposed variations such as residual connection, attention mechanisms, squeeze and excitation, and dilated convolutions.
#
# * Prefers large patch size rather than batch size.
#
# **b. Training Schedule**
#
# * 100 epochs; one epoch defined as iteration over 250 mini-batches.
#
# * Stochastic gradient descent with Nestrov momentum.
# * Loss is the combination of cross-entropy and dice loss.
# * Oversampling to handle class imbalance.
# * *Data Augmentation :* rotation, scaling, guassian noise, guassian blur, brightness, contrast, simulation of low resolution and gamma resolution.
#
# **c. Inference**
#
# * Images are predicted with a sliding window approach, where the window size equals the patch size used during training. Adjacent predictions overlap by half the size of a patch.
#
# * To suppress stitching artifacts and reduce the influence of positions close to the borders, a Gaussian importance weighing is applied, increasing the weight of the center voxels in the softmax aggregation.
#
# + [markdown] id="WJLB_aYXphHP" colab_type="text"
# ### **3. Inferred Parameters**
#
# **a. Intensity Normalization :**
# * Z-scoring for all modalities except CT.
# * For CT, it follows a global normalization scheme, which uses 0.5 and 99.5 percentiles of the foreground voxels for clipping.
#
# **b. Resampling :**
# * To cope up with the heterogeneity in medical domain,
# * Resamples all images to same target spacing using either third order spline, linear or nearest neighbor interpolation.
#
# **c. Adaptation of network topology, patch size and batch size :** The network topology for all U-Net configurations is chosen on the basis of the median image size after resampling as well as the target spacing the images were resampled to.
#
# **d. Initialization :** The patch size is initialized as the median image shape after resampling.
#
# **e. Architecture Topology:**
# * The architecture is configured by determining the number of down-sampling operations, performed until the feature map is reduced to 4 voxels or feature map space become anisotropic.
# * High resolution axes are down-sampled separately until their resolution is within factor 2 of the lower resolution axis.
# * Each axis is down-sampled individually, until the feature map constraints are triggered.
# * The default kernel size for convolutions is 3×3×3 and 3×3 for 3D U-Net and 2D U-Net, respectively.
#
# + [markdown] id="0q1Jhsqfxo_3" colab_type="text"
# ### **4. Empirical Parameters**
#
# **a. Ensembling and selection of U-Net configuration(s) :**
# * Automatically ensembles based on average foreground Dice coefficient computed via Cross-validation on training data to use for inference
# * **Configuration model(s):** Single models (2D, 3D fullres, 3D lowres or fullres U-Net of the cascade) or an ensemble of any two of these configurations.
# * Models are ensembled by averaging softmax probabilities.
#
# **b. Postprocessing:** Connected component-based postprocessing is used.
# All foreground classes are treated as one component, to improve the average foreground Dice coefficient and if it does not reduce the Dice coefficient for any of the classes, then nnU-Net builds on the outcome of this step and decides whether the same procedure should be performed for individual classes.
# + [markdown] id="qoBRVgXJjzn3" colab_type="text"
# ## **Different U-Net configurations**
#
# 1. **2D U-Net**
# 2. **3D U-Net Full Resolution**
# 3. **3D U-Net Cascaded :** The first U-Net operates on downsampled images and the second is trained to refine the segmentation maps created by the former at full resolution.
#
# + [markdown] id="mBJV4lJGTEex" colab_type="text"
# 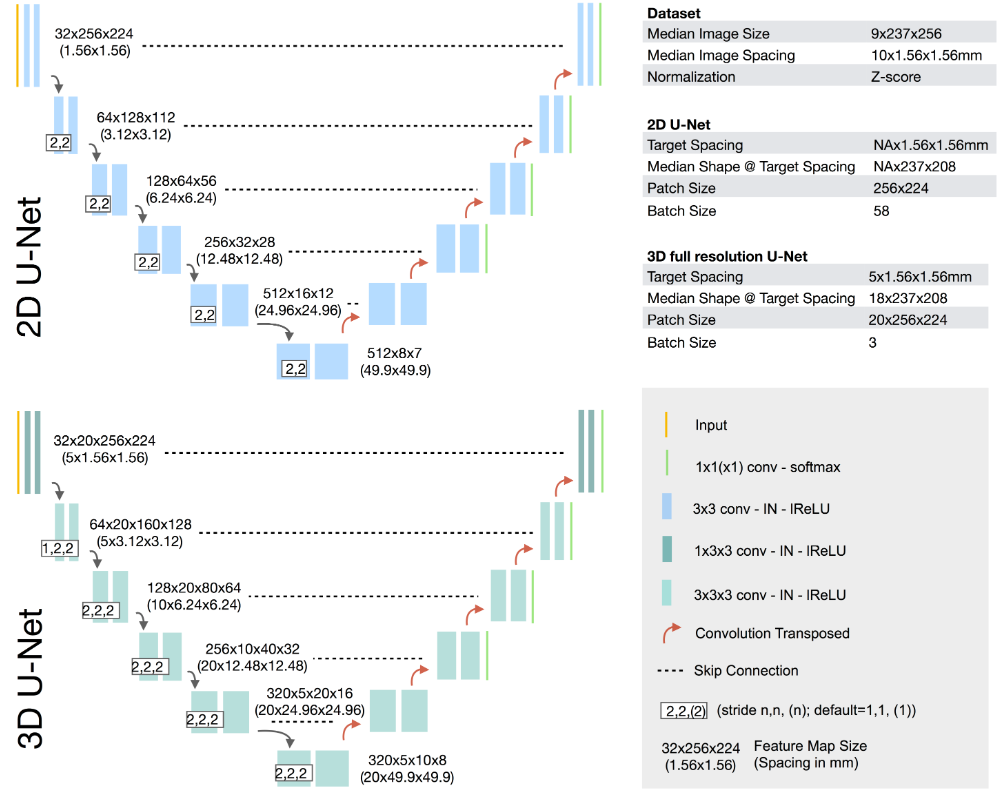
# **Figure :** for illustratation purposes. network architecture for ACDC dataset, from Medical Decathlon taken from Paper 1 mentioned below. Filters and no. of channels will remain same for any dataset.
# + [markdown] id="nK4WEPmCjGHZ" colab_type="text"
# ### **About the Spinal Cord Grey Matter Challenge**
#
# [SCGM Challenge](http://niftyweb.cs.ucl.ac.uk/program.php?p=CHALLENGE), has a collection of healthy spinal cord images collected at 4 different sites. As part of the challenge, you need to automatically or semi-automatically segment the anatomical MR Images into background class (as 0) and grey matter class (as 1). Algorithms will be evaluated against manual segmentations from four trained raters (one from each site) in terms of segmentation accuracy and precision.
# Predicted images must be saved in NIFIT format, with same space and resolution as the provided data.
#
# **Note:** Training images are labelled for Background class as (0), Grey Matter class as (1) and White Matter class as (2).
# + [markdown] id="XY2DYdEXbdyu" colab_type="text"
# ### **Major Sections:**
# Train a nnUNet model for the Spinal Cord Grey Matter Segmentation Dataset and infer results . Same approach can be applied on other datasets, with dataset specific modifications.
#
# + [markdown] id="FKwAI8FN4Iir" colab_type="text"
#
#
# 1. [Setup](#setup) *(Always,CPU, GPU)*
#
# 2. [Cloning the Repo's](#clone) *(One-time, CPU, GPU)*
#
# 3. [Install and import libraries](#libraries) *(Everytime, CPU, GPU)*
#
# 4. [Structure Dataset Folder](#folder_structure) *(Evertyime, CPU, GPU)*
#
# 5. [Environment Variables](#env_variables) *(Everytime, CPU, GPU)*
#
# 6. [Unzip, Rename, Train and Test Set](#dataset_setup) *(One-time, CPU, GPU)*
#
# 7. [Dataset verification](#dataset_verify) *(One-time, Optional, CPU, GPU)*
#
# 8. [Dataset Visualization](#data_visual) *(CPU,GPU)*
#
# 9. [Training Code](#train) *(GPU only)*
#
# 10. [Inference Code](#infer) *(GPU only)*
#
# 11. [Prediction Visualization](#predict_visual) *(CPU,GPU)*
#
# 12. [Submit Code for SCGM](#submit) *(Optional, CPU, GPU)*
#
#
# **Note:** For training and inference, you have to run through steps 1,3,4 everytime for colab users. For local machines, installing libraries and setting up environment variables again is not necessary.
#
# + [markdown] id="N5rkX_WLVM9D" colab_type="text"
# **Papers:**
#
# 1) nnU-Net: Self-adapting Framework for U-Net-Based Medical Image Segmentation [(arxiv)](https://arxiv.org/pdf/1809.10486.pdf)
#
# 2) Automated Design of Deep Learning Methods for Biomedical Image Segmentation [(arxiv)](https://arxiv.org/pdf/1904.08128.pdf)
#
# **Github repository:**
#
# 1) [Orginal Repository](https://github.com/MIC-DKFZ/nnUNet) - regularly maintained and changed by the authors.
#
# 2) [Forked Repository](https://github.com/prateekgupta891/nnUNet) - to ensure a version of nnUNet with which this tutorial works (as the original repository is constantly modified.)
# + [markdown] id="mP9BHLQ4AGTu" colab_type="text"
# (**Colab users** - Preferrably use GPU runtime, but you can change to GPU runtime afterwards, as and when required)
# + [markdown] id="HxUpC3-SLqd7" colab_type="text"
# ##**1. Setup Section**
# + id="oFAMjhgk91jX" colab_type="code" colab={}
#for colab users only - mounting the drive
from google.colab import drive
drive.mount('/content/drive',force_remount = False)
# + id="x4fqtH2k-OQn" colab_type="code" colab={}
#setup a base directory where everything will be installed - repo, dataset, libraries
#This .ipynb notebook needs to placed there as well.
import os
base_dir = '/content/drive/My Drive/Colab Notebooks'
os.chdir(base_dir)
# + [markdown] id="GrJJ3FWT_W7f" colab_type="text"
# ##**2. Clone the repository**
# + id="7Rt0PmGg_pbV" colab_type="code" colab={}
#comment after installed once
# #!git clone https://github.com/MIC-DKFZ/nnUNet.git
# !git clone https://github.com/prateekgupta891/nnUNet.git #my forked version
# !git clone https://github.com/NVIDIA/apex
# + [markdown] id="fSq91BNdPRsH" colab_type="text"
# ## **3. Install and Import Libraries**
# + id="sSNdbg2WDTna" colab_type="code" colab={}
#colab users - Do this everytime
#local machines - Once is enough
respository_dir = os.path.join(base_dir,'nnUNet')
os.chdir(respository_dir)
# !pip install -e .
#(optional installation)
# !pip install --upgrade git+https://github.com/nanohanno/hiddenlayer.git@bugfix/get_trace_graph#egg=hiddenlayer
os.chdir(base_dir)
# + [markdown] id="xj2vt5vkRqKv" colab_type="text"
# (**Colab Users:** You must restart your runtime after installing the libraries.)
# + id="bDA08Yo3PifU" colab_type="code" colab={}
#libraries
import shutil
from collections import OrderedDict
import json
import numpy as np
#visualization of the dataset
import matplotlib.pyplot as plt
import nibabel as nib
#for colab users only - keep the base directory same as above
import os
base_dir = "/content/drive/My Drive/Colab Notebooks"
repository_dir = os.path.join(base_dir,'nnUNet')
os.chdir(base_dir)
# + id="XdJ3WoUxRV4r" colab_type="code" colab={}
if os.getcwd()==base_dir:
print('We are in the correct directory')
else:
print("Run set base directory step again, then check to verify.")
# + [markdown] id="dFSfF07wXs1i" colab_type="text"
# ## **4. Dataset Folder Structure**
# + id="ARyAWPHFRbN0" colab_type="code" colab={}
def make_if_dont_exist(folder_path,overwrite=False):
"""
creates a folder if it does not exists
input:
folder_path : relative path of the folder which needs to be created
over_write :(default: False) if True overwrite the existing folder
"""
if os.path.exists(folder_path):
if not overwrite:
print(f'{folder_path} exists.')
else:
print(f"{folder_path} overwritten")
shutil.rmtree(folder_path)
os.makedirs(folder_path)
else:
os.makedirs(folder_path)
print(f"{folder_path} created!")
# + [markdown] id="vSX71vh73ugt" colab_type="text"
#
#
#
#
# **Custom Task Id starts at 101,** to ensure that there will be no conflicts with downloaded pretrained models.
#
#
#
# ```
# Task Naming Convention: Task[Task Id]_[Task Name] eg. Task101_SCGM
# ```
#
#
# + id="ib5XdSZIXcj4" colab_type="code" colab={}
task_name = 'Task101_SCGM' #change here for different task name
nnunet_dir = "nnUNet/nnunet/nnUNet_raw_data_base/nnUNet_raw_data"
task_folder_name = os.path.join(nnunet_dir,task_name)
train_image_dir = os.path.join(task_folder_name,'imagesTr')
train_label_dir = os.path.join(task_folder_name,'labelsTr')
test_dir = os.path.join(task_folder_name,'imagesTs')
main_dir = os.path.join(base_dir,'nnUNet/nnunet')
# + id="5pU7_h5JXeTH" colab_type="code" colab={}
make_if_dont_exist(task_folder_name,overwrite = False)
make_if_dont_exist(train_image_dir)
make_if_dont_exist(train_label_dir)
make_if_dont_exist(test_dir,overwrite= False)
make_if_dont_exist(os.path.join(main_dir,'nnunet_trained_models'))
# + [markdown] id="tYO2HVkegkkS" colab_type="text"
# ## **5.Environment Variables**
# + id="3TDrmYRLgjSt" colab_type="code" colab={}
os.environ['nnUNet_raw_data_base'] = os.path.join(main_dir,'nnUNet_raw_data_base')
os.environ['nnUNet_preprocessed'] = os.path.join(main_dir,'preprocessed')
os.environ['RESULTS_FOLDER'] = os.path.join(main_dir,'nnUNet_trained_models')
# + [markdown] id="1OSPNZos70Cz" colab_type="text"
# **Colab Users:** Everytime you re-run or restart your kernel always run until this point.
# + [markdown] id="1CfdGthC4KhY" colab_type="text"
# ## **6. Unzip, Rename, Train and Test Set**
#
#
# **(Manual Task)** Get the train and test data in zip form and place it
# in the folder: /nnUNet/nnunet/nnUNet_raw_data_base/nnUNet_raw_data/Task_folder_name .
#
# Apply to get Spinal Cord Grey Matter Challenge [Dataset](http://cmictig.cs.ucl.ac.uk/niftyweb/challenge/).
#
#
# + [markdown] id="pzD5Ya6wm5Xn" colab_type="text"
# Code will take care of the following:
#
# 1. Unzip train and test files
#
# 2. Rename the train images and labels to match and placing in respective directories.
#
# 3. Put testing data in the folder and remove text files
#
# 4. Add modality at the end of each file, as nnU-net can train on multiple modalities together
#
# 5. Create dataset.json
#
# **Note:** For a new dataset, you may need to do few changes for training. Also, nnUNet works with .nii.gz files only.
# + id="K6dqTN19kmET" colab_type="code" colab={}
def copy_and_rename(old_location,old_file_name,new_location,new_filename,delete_original = False):
shutil.copy(os.path.join(old_location,old_file_name),new_location)
os.rename(os.path.join(new_location,old_file_name),os.path.join(new_location,new_filename))
if delete_original:
os.remove(os.path.join(old_location,old_file_name))
# + id="5jhROK0ym6fO" colab_type="code" colab={}
os.chdir(task_folder_name)
if os.path.isfile('training-data-gm-sc-challenge-ismrm16-v20160302b.zip'):
print(f'Training file for exists')
else:
print('Training file for SCGM Challenge is not present in the directory')
if os.path.isfile('test-data-gm-sc-challenge-ismrm16-v20160401.zip'):
print('Testing file for SCGM Challenge exists')
else:
print('Testing file for SCGM Challenge is not present in the directory')
os.chdir(base_dir)
# + id="-O52YOFneGKR" colab_type="code" colab={}
#unzipping in nnUNet_raw folder the training data
os.chdir(task_folder_name)
# !unzip training-data-gm-sc-challenge-ismrm16-v20160302b.zip
os.chdir(base_dir)
# + [markdown] id="lsRlhvtonW4q" colab_type="text"
# ### *Rename and Relocate*
#
# We have 4 annotation of the same image, by different experts in the SCGM Challenge. *( Image , Ann1 )* and *( Image , Ann2 )* can be considered as a different image and label pairs. Hence, 4 copies of the training .nii.gz file is created with its mapping to the respective label name.
# + id="w89GuKbwnz8Q" colab_type="code" colab={}
#putting training images into folder
mask_count = 4 #change if more mask is available
for file in os.listdir(task_folder_name):
if file.endswith('.nii.gz'):
if file.find('mask')!=-1:
#putting mask
shutil.move(os.path.join(task_folder_name,file),train_label_dir)
else:
#making 4 copies
for mask in range(1,mask_count+1):
new_filename = file[:file.find('-image')] + '-mask-r' + str(mask) + '.nii.gz'
if mask==mask_count:
copy_and_rename(task_folder_name,file,train_image_dir,new_filename,delete_original = True)
else:
copy_and_rename(task_folder_name,file,train_image_dir,new_filename)
#removing all other files installed due to the unzip
elif file.endswith('.txt'):
os.remove(os.path.join(task_folder_name,file))
# + [markdown] id="jGR5w7n4oFCk" colab_type="text"
# ### *Verification*
# + id="B27xTmMcn1ca" colab_type="code" colab={}
train_files = os.listdir(train_image_dir)
label_files = os.listdir(train_label_dir)
print("train image files:",len(train_files))
print("train label files:",len(label_files))
print("Matches:",len(set(train_files).intersection(set(label_files))))
#should be equal to 160 for SCGM Challenge
# + id="bYuUAfjHoDgs" colab_type="code" colab={}
#unzip the testing files in nnUNet_raw folder
os.chdir(task_folder_name)
# !unzip test-data-gm-sc-challenge-ismrm16-v20160401.zip
os.chdir(base_dir)
# + id="jjHDon9Vn2_8" colab_type="code" colab={}
for file in os.listdir(task_folder_name):
if file.endswith('.nii.gz'):
#putting mask
shutil.move(os.path.join(task_folder_name,file),test_dir)
#removing all other files installed due to the unzip
elif file.endswith('.txt'):
os.remove(os.path.join(task_folder_name,file))
# + id="wn46w0FTn5-o" colab_type="code" colab={}
print("Testing files:",len(os.listdir(test_dir)))
print(test_dir)
#for spinal cord dataset testing files needs to be equal to 40.
# + id="4QUfzfcIqndD" colab_type="code" colab={}
#renaming to add the modality for SCGM there is only one modality
#images should be added with 0000
#can be skipped if modality is already mentioned
#re-write for multiple modalities
def check_modality(filename):
"""
check for the existence of modality
return False if modality is not found else True
"""
end = filename.find('.nii.gz')
modality = filename[end-4:end]
for mod in modality:
if not(ord(mod)>=48 and ord(mod)<=57): #if not in 0 to 9 digits
return False
return True
def rename_for_single_modality(directory):
for file in os.listdir(directory):
if check_modality(file)==False:
new_name = file[:file.find('.nii.gz')]+"_0000.nii.gz"
os.rename(os.path.join(directory,file),os.path.join(directory,new_name))
print(f"Renamed to {new_name}")
else:
print(f"Modality present: {file}")
rename_for_single_modality(train_image_dir)
rename_for_single_modality(test_dir)
# + [markdown] id="u-XfPdmpq79m" colab_type="text"
# ### Creating **dataset.json**
# + id="TmrSXf2srHjB" colab_type="code" colab={}
overwrite_json_file = True #make it True if you want to overwrite the dataset.json file in Task_folder
json_file_exist = False
if os.path.exists(os.path.join(task_folder_name,'dataset.json')):
print('dataset.json already exist!')
json_file_exist = True
if json_file_exist==False or overwrite_json_file:
json_dict = OrderedDict()
json_dict['name'] = task_name
json_dict['description'] = "Spinal Cord Grey Matter Segmenation Challenge"
json_dict['tensorImageSize'] = "3D"
json_dict['reference'] = "see challenge website"
json_dict['licence'] = "see challenge website"
json_dict['release'] = "0.0"
#you may mention more than one modality
json_dict['modality'] = {
"0": "MRI"
}
#labels+1 should be mentioned for all the labels in the dataset
json_dict['labels'] = {
"0": "background",
"1": "grey matter",
"2": "white matter"
}
train_ids = os.listdir(train_label_dir)
test_ids = os.listdir(test_dir)
json_dict['numTraining'] = len(train_ids)
json_dict['numTest'] = len(test_ids)
#no modality in train image and labels in dataset.json
json_dict['training'] = [{'image': "./imagesTr/%s" % i, "label": "./labelsTr/%s" % i} for i in train_ids]
#removing the modality from test image name to be saved in dataset.json
json_dict['test'] = ["./imagesTs/%s" % (i[:i.find("_0000")]+'.nii.gz') for i in test_ids]
with open(os.path.join(task_folder_name,"dataset.json"), 'w') as f:
json.dump(json_dict, f, indent=4, sort_keys=True)
if os.path.exists(os.path.join(task_folder_name,'dataset.json')):
if json_file_exist==False:
print('dataset.json created!')
else:
print('dataset.json overwritten!')
# + [markdown] id="gMPYPrUVslFi" colab_type="text"
# ##**7. Dataset Verification**
# + id="wjGLZk148MRb" colab_type="code" colab={}
#running it from the experiment_planning folder to verify the path settings
os.chdir(main_dir)
# !python experiment_planning/nnUNet_plan_and_preprocess.py -t 101 --verify_dataset_integrity
os.chdir(base_dir)
# + [markdown] id="I25_XU_Ie7TZ" colab_type="text"
# ## **8. Dataset Visualization**
# + [markdown] id="_EG_aX8qsrM6" colab_type="text"
# ### Train Data
# + id="bRCc6EPBwqNy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 499} outputId="d32c2028-b70c-4ea4-8e9d-43c9d0777730"
#visualizing some of the training images and labels
# (re-run to see random pick-ups)
# only maximum of first 5 slices are plotted
train_img_name = os.listdir(train_image_dir)[np.random.randint(0,160)]
train_img = np.array(nib.load(os.path.join(train_image_dir,train_img_name)).dataobj)[:,:,:5]
train_label_name = train_img_name[:train_img_name.find('_0000.nii.gz')]+'.nii.gz'
train_label = np.array(nib.load(os.path.join(train_label_dir,train_label_name)).dataobj)[:,:,:5]
print(train_img.shape,train_label.shape)
max_rows = 2
max_cols = train_img.shape[2]
fig, axes = plt.subplots(nrows=max_rows, ncols=max_cols, figsize=(20,8))
for idx in range(max_cols):
axes[0, idx].axis("off")
axes[0, idx].set_title('Train Image'+str(idx+1))
axes[0 ,idx].imshow(train_img[:,:,idx], cmap="gray")
for idx in range(max_cols):
axes[1, idx].axis("off")
axes[1, idx].set_title('Train Label'+str(idx+1))
axes[1, idx].imshow(train_label[:,:,idx])
plt.subplots_adjust(wspace=.1, hspace=.1)
plt.show()
# + [markdown] id="bE4ohbyTrVSk" colab_type="text"
# **Note:** In the label image, yellow color represents white Matter and green-ish color represents grey matter.
# + [markdown] id="3SwfcAeVfB94" colab_type="text"
# ### Test Data
# + id="rcDeru1kcLxQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 272} outputId="b8529720-dc90-44ff-e7bf-c5e133b12cf9"
#visualizing some of the test images
# (re-run to see random pick-ups)
# only maximum of first 5 slices are plotted
test_img_name = os.listdir(test_dir)[np.random.randint(0,40)]
test_img = np.array(nib.load(os.path.join(test_dir,test_img_name)).dataobj)[:,:,:5]
print(test_img.shape)
max_cols = test_img.shape[2]
max_rows = 1
fig, axes = plt.subplots(nrows=max_rows, ncols=max_cols, figsize=(20,20))
for idx in range(max_cols):
axes[ idx].axis("off")
axes[ idx].set_title('Test Image'+str(idx))
axes[ idx].imshow(test_img[:,:,idx], cmap="gray")
plt.subplots_adjust(wspace=.1, hspace=.1)
plt.show()
# + [markdown] id="DYdrPRfqVHl7" colab_type="text"
# ## **9. Training Code**
# + [markdown] id="UfapEc3NOmQb" colab_type="text"
# nnU-Net stores a checkpoint every 50 epochs. If you need to continue a previous training, just add a -c to the training command.
#
# **Generic Training Commands:**
#
# nnUNet_train CONFIGURATION TRAINER_CLASS_NAME TASK_NAME_OR_ID FOLD (additional options)
#
# + [markdown] id="OKAjmGzHvxUW" colab_type="text"
# **For 2D:** ``` nnUNet_train 2d nnUNetTrainerV2 TaskXXX_MYTASK FOLD```
#
# **For 3D Full resolution:** ``` nnUNet_train 3d_fullres nnUNetTrainerV2 TaskXXX_MYTASK FOLD```
#
# **For Cascaded 3D:**
#
# First Run lowres: ``` nnUNet_train 3d_lowres nnUNetTrainerV2 TaskXXX_MYTASK FOLD```
#
# Then Run fullres: ``` nnUNet_train 3d_cascade_fullres nnUNetTrainerV2CascadeFullRes TaskXXX_MYTASK FOLD ```
#
#
#
# + id="4OiB3r5sNguW" colab_type="code" colab={}
#colab users - mandatory
#local machine - once is sufficient
os.chdir('apex')
# !pip install -v --no-cache-dir --global-option="--cpp_ext" --global-option="--cuda_ext" ./
os.chdir(base_dir)
# + [markdown] id="KC614zRCRguc" colab_type="text"
# Training for 3D fullres with Trainer V2 and for Fold 0.
# + id="It1UZ04bQ62r" colab_type="code" colab={}
os.chdir(main_dir)
# !nnUNet_train 3d_fullres nnUNetTrainerV2 101 0
os.chdir(base_dir)
# + [markdown] id="6jvxL2xHQ4sm" colab_type="text"
# ##**10. Inference Code**
# ```nnUNet_find_best_configuration``` will print inference commands you need to use. The easiest way to run inference is to simply use these commands.
#
#
# For each of the desired configurations, run:
# ```
# nnUNet_predict -i INPUT_FOLDER -o OUTPUT_FOLDER -t TASK_NAME_OR_ID -m CONFIGURATION --save_npz
# ```
# Only specify --save_npz if you intend to use ensembling. --save_npz will make the command save the softmax probabilities alongside of the predicted segmentation masks requiring a lot of disk space.
#
# **Note:** Please select a separate OUTPUT_FOLDER for each configuration!
#
# + id="QSfJ8sQE3C37" colab_type="code" colab={}
#optional
os.chdir(repository_dir)
# !nnUNet_find_best_configuration -t 101
os.chdir(base_dir)
# + id="6bLhYZT8Poxf" colab_type="code" colab={}
result_dir = os.path.join(main_dir,'nnUNet_Prediction_Results',task_name)
make_if_dont_exist(result_dir)
team_name = 'prateek3' #make sure to change for your own team name
# + [markdown] id="UdGf_PNXhNiA" colab_type="text"
# **Note:** If you interrupted the training, then then rename **model_best.model.pkl** to **model_final_checkpoint.model.pkl** and **model_best.model** to **model_final_checkpoint.model** for the given fold.
# + id="4jaO3vKWQg0m" colab_type="code" colab={}
#location where you want save your results, will be created if dont exist
os.chdir(main_dir)
# !nnUNet_predict -i nnUNet_raw_data_base/nnUNet_raw_data/Task101_SCGM/imagesTs -o nnUNet_Prediction_Results/Task101_SCGM -t 101 -tr nnUNetTrainerV2 -m 3d_fullres --num_threads_preprocessing 1
os.chdir(base_dir)
# + [markdown] id="I1klVm1u3zm0" colab_type="text"
# If you wish to run ensembling, you can ensemble the predictions from several configurations with the following command:
# ```
# nnUNet_ensemble -f FOLDER1 FOLDER2 ... -o OUTPUT_FOLDER -pp POSTPROCESSING_FILE
# ```
# + [markdown] id="Fc1xlu6snBmc" colab_type="text"
# ## **11. Prediction Visualization**
# + id="rf4aAk_CfW7-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 518} outputId="b9040bdd-6ed4-4057-c614-1303c7c89df5"
#visualizing the predicted results
# (re-run to see random pick-ups)
# only maximum of first 5 slices are plotted
test_img_name = os.listdir(test_dir)[np.random.randint(0,40)]
test_img = np.array(nib.load(os.path.join(test_dir,test_img_name)).dataobj)[:,:,:5]
predicted_img_name = test_img_name[:test_img_name.find('_0000.nii.gz')]+'.nii.gz'
predicted_label = np.array(nib.load(os.path.join(result_dir,predicted_img_name)).dataobj)[:,:,:5]
print('Test Image Shape: ',test_img.shape)
print("Predicted Image Shape:",predicted_label.shape)
max_rows = 2
max_cols = test_img.shape[2]
fig, axes = plt.subplots(nrows=max_rows, ncols=max_cols, figsize=(20,8))
for idx in range(max_cols):
axes[0, idx].axis("off")
axes[0, idx].set_title('Test Image'+str(idx+1))
axes[0 ,idx].imshow(test_img[:,:,idx], cmap="gray")
for idx in range(max_cols):
axes[1, idx].axis("off")
axes[1, idx].set_title('Predicted Label'+str(idx+1))
axes[1, idx].imshow(predicted_label[:,:,idx])
plt.subplots_adjust(wspace=.1, hspace=.1)
plt.show()
# + [markdown] id="v_J4AHCePDLT" colab_type="text"
# ## **12. SCGM submission** (optional)
#
# While trianing, we trained the dataset to learn, **White Matter** as well as **Grey Matter**. But for the challenge we only need to predict *Gray matter* labelled as *1*, and *everything else is 0*. So we convert the labels.
#
# Test images naming convention: **[Original test filename]-[team name].nii.gz**
# + id="-LgCThfxQk94" colab_type="code" colab={}
#specific to the submission
submission_folder = os.path.join(result_dir,'submission_folder')
make_if_dont_exist(submission_folder)
for file in os.listdir(result_dir):
if file.endswith('.nii.gz'):
img = nib.load(os.path.join(result_dir,file))
img_np = np.array(img.dataobj)
img_np[img_np==2.0] = 0.0
img_nifti = nib.Nifti1Image(img_np,affine = np.eye(4))
new_file = file[:file.find('image')]+team_name+'.nii.gz'
nib.save(img_nifti, os.path.join(submission_folder,new_file))
print(new_file)
# + [markdown] id="tyKI8_5TPd5_" colab_type="text"
# **Test Results:**
# Download the the folder, and upload with at the following [link](http://niftyweb.cs.ucl.ac.uk/program.php?p=CHALLENGE).
# + [markdown] id="nnbYaA5_m2IU" colab_type="text"
#
# 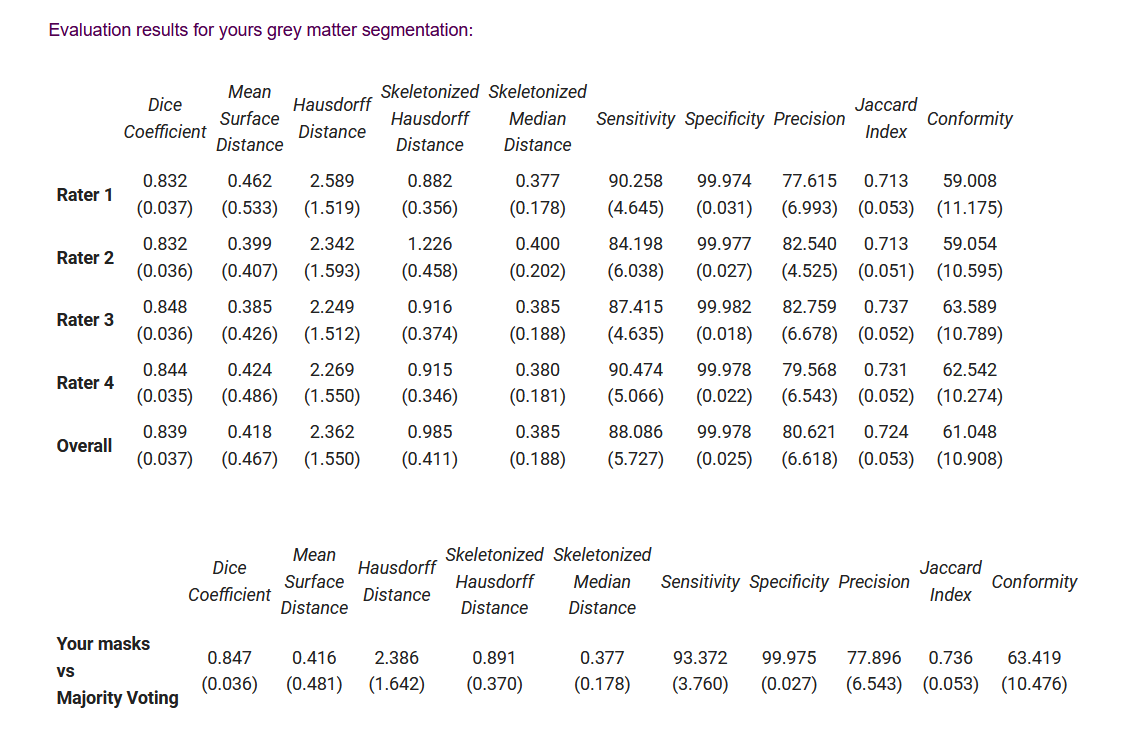
# + [markdown] id="mpb75yADwc4B" colab_type="text"
# **Note:** These results are achieved with only 10 epochs on 3D full resolution on a single fold without post-processing, and it is better than results, mentioned by SCGM Challenge website [here](http://niftyweb.cs.ucl.ac.uk/program.php?p=CHALLENGE).
# Dice Score is better by 0.04 from the best results.
|
nnunetmec2020.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import networkx as nx
G = nx.read_adjlist("data/tiny_network.adjlist", create_using=nx.DiGraph, delimiter=";")
list(G.nodes)
# +
graph = nx.read_adjlist("data/tiny_network.adjlist", create_using=nx.DiGraph, delimiter=";")
visited = []
queue = []
backtrace = {} # keep track of parent nodes
path = []
# add node with no connections
graph.add_node('pineapple')
def bfs(start, end=None):
"""
TODO: write a method that performs a breadth first traversal and pathfinding on graph G
* If there's no end node, just return a list with the order of traversal
* If there is an end node and a path exists, return a list of the shortest path
* If there is an end node and a path does not exist, return None
"""
# add start node to queue
queue.append(start)
# mark start node as visited
visited.append(start)
if start==end:
return visited
# while there is a queue, dequeue current node and loop through unvisited neighbors of current node
while queue:
just_popped = queue.pop(0) # first in first out
for neighbor in graph.neighbors(just_popped):
if neighbor not in visited:
queue.append(neighbor)
visited.append(neighbor)
backtrace[neighbor]=just_popped # set pointer from neighbor to current (parent) node
# If there is an end node and a path exists, return a list of the shortest path
if neighbor==end:
curr_node = end
path.append(curr_node)
while curr_node!=start:
parent = backtrace[curr_node] # identify parent node
path.append(parent) # add parent node to path
curr_node = parent # set current node to parent node
path.reverse() # reverse direction of path
return path
# If there's no end node, just return a list with the order of traversal
if end==None:
return visited
# If there is an end node and a path does not exist, return None
if end not in visited:
return None
bfs('<NAME>', '<NAME>')
# +
class Graph:
"""
Class to contain a graph and your bfs function
"""
def __init__(self, filename: str):
"""
Initialization of graph object which serves as a container for
methods to load data and
"""
self.graph = nx.read_adjlist(filename, create_using=nx.DiGraph, delimiter=";")
self.visited = []
self.queue = []
self.backtrace = {} # keep track of parent nodes
self.path = [] # shortest path
def bfs(self, start, end=None):
"""
TODO: write a method that performs a breadth first traversal and pathfinding on graph G
* If there's no end node, just return a list with the order of traversal
* If there is an end node and a path exists, return a list of the shortest path
* If there is an end node and a path does not exist, return None
"""
# add start node to queue
self.queue.append(start)
# mark start node as visited
self.visited.append(start)
if start==end:
return self.visited
# while there is a queue, dequeue current node and loop through unvisited neighbors of current node
while self.queue:
just_popped = self.queue.pop(0) # first in first out
for neighbor in self.graph.neighbors(just_popped):
if neighbor not in self.visited:
self.queue.append(neighbor)
self.visited.append(neighbor)
self.backtrace[neighbor]=just_popped # set pointer from neighbor to current (parent) node
# If there is an end node and a path exists, return a list of the shortest path
if neighbor==end:
curr_node = end
self.path.append(curr_node)
while curr_node!=start:
parent = backtrace[curr_node] # identify parent node
self.path.append(parent) # add parent node to path
curr_node = parent # set current node to parent node
path.reverse() # reverse direction of path
return path
# If there's no end node, just return a list with the order of traversal
if end==None:
return self.visited
# If there is an end node and a path does not exist, return None
if end not in self.visited:
return None
new_instance = Graph("data/tiny_network.adjlist")
print(new_instance.bfs('<NAME>'))
# -
|
testbed.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# !python setup.py build_ext --inplace
import os
import random
import math
import numpy as np
import pandas as pd
from PIL import Image, ImageDraw
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
import torchvision
from torchvision import datasets, models, transforms
from torchvision.models.detection.retinanet import RetinaNet
from torchvision.models.detection.faster_rcnn import FasterRCNN
import torchvision.transforms.functional as F
from typing import Any, Callable, Dict, List, Optional, Tuple, Union
import xml.etree.ElementTree as ET
import collections
from torchvision.datasets.voc import VisionDataset
from metrics import *
# +
from collections import defaultdict, deque
import time
import datetime
import torch.distributed as dist
import torchvision.transforms.functional as F
class SmoothedValue(object):
"""Track a series of values and provide access to smoothed values over a
window or the global series average.
"""
def __init__(self, window_size=20, fmt=None):
if fmt is None:
fmt = "{median:.4f} ({global_avg:.4f})"
self.deque = deque(maxlen=window_size)
self.total = 0.0
self.count = 0
self.fmt = fmt
def update(self, value, n=1):
self.deque.append(value)
self.count += n
self.total += value * n
def synchronize_between_processes(self):
"""
Warning: does not synchronize the deque!
"""
if not is_dist_avail_and_initialized():
return
t = torch.tensor([self.count, self.total], dtype=torch.float64, device='cuda')
dist.barrier()
dist.all_reduce(t)
t = t.tolist()
self.count = int(t[0])
self.total = t[1]
@property
def median(self):
d = torch.tensor(list(self.deque))
return d.median().item()
@property
def avg(self):
d = torch.tensor(list(self.deque), dtype=torch.float32)
return d.mean().item()
@property
def global_avg(self):
return self.total / self.count
@property
def max(self):
return max(self.deque)
@property
def value(self):
return self.deque[-1]
def __str__(self):
return self.fmt.format(
median=self.median,
avg=self.avg,
global_avg=self.global_avg,
max=self.max,
value=self.value)
class MetricLogger(object):
def __init__(self, delimiter="\t"):
self.meters = defaultdict(SmoothedValue)
self.delimiter = delimiter
def update(self, **kwargs):
for k, v in kwargs.items():
if isinstance(v, torch.Tensor):
v = v.item()
assert isinstance(v, (float, int))
self.meters[k].update(v)
def __getattr__(self, attr):
if attr in self.meters:
return self.meters[attr]
if attr in self.__dict__:
return self.__dict__[attr]
raise AttributeError("'{}' object has no attribute '{}'".format(
type(self).__name__, attr))
def __str__(self):
loss_str = []
for name, meter in self.meters.items():
loss_str.append(
"{}: {}".format(name, str(meter))
)
return self.delimiter.join(loss_str)
def synchronize_between_processes(self):
for meter in self.meters.values():
meter.synchronize_between_processes()
def add_meter(self, name, meter):
self.meters[name] = meter
def log_every(self, iterable, print_freq, header=None):
i = 0
if not header:
header = ''
start_time = time.time()
end = time.time()
iter_time = SmoothedValue(fmt='{avg:.4f}')
data_time = SmoothedValue(fmt='{avg:.4f}')
space_fmt = ':' + str(len(str(len(iterable)))) + 'd'
if torch.cuda.is_available():
log_msg = self.delimiter.join([
header,
'[{0' + space_fmt + '}/{1}]',
'eta: {eta}',
'{meters}',
'time: {time}',
'data: {data}',
'max mem: {memory:.0f}'
])
else:
log_msg = self.delimiter.join([
header,
'[{0' + space_fmt + '}/{1}]',
'eta: {eta}',
'{meters}',
'time: {time}',
'data: {data}'
])
MB = 1024.0 * 1024.0
for obj in iterable:
data_time.update(time.time() - end)
yield obj
iter_time.update(time.time() - end)
if i % print_freq == 0 or i == len(iterable) - 1:
eta_seconds = iter_time.global_avg * (len(iterable) - i)
eta_string = str(datetime.timedelta(seconds=int(eta_seconds)))
if torch.cuda.is_available():
print(log_msg.format(
i, len(iterable), eta=eta_string,
meters=str(self),
time=str(iter_time), data=str(data_time),
memory=torch.cuda.max_memory_allocated() / MB))
else:
print(log_msg.format(
i, len(iterable), eta=eta_string,
meters=str(self),
time=str(iter_time), data=str(data_time)))
i += 1
end = time.time()
total_time = time.time() - start_time
total_time_str = str(datetime.timedelta(seconds=int(total_time)))
print('{} Total time: {} ({:.4f} s / it)'.format(
header, total_time_str, total_time / len(iterable)))
def warmup_lr_scheduler(optimizer, warmup_iters, warmup_factor):
def f(x):
if x >= warmup_iters:
return 1
alpha = float(x) / warmup_iters
return warmup_factor * (1 - alpha) + alpha
return torch.optim.lr_scheduler.LambdaLR(optimizer, f)
def is_dist_avail_and_initialized():
if not dist.is_available():
return False
if not dist.is_initialized():
return False
return True
def get_world_size():
if not is_dist_avail_and_initialized():
return 1
return dist.get_world_size()
def reduce_dict(input_dict, average=True):
"""
Args:
input_dict (dict): all the values will be reduced
average (bool): whether to do average or sum
Reduce the values in the dictionary from all processes so that all processes
have the averaged results. Returns a dict with the same fields as
input_dict, after reduction.
"""
world_size = get_world_size()
if world_size < 2:
return input_dict
with torch.no_grad():
names = []
values = []
# sort the keys so that they are consistent across processes
for k in sorted(input_dict.keys()):
names.append(k)
values.append(input_dict[k])
values = torch.stack(values, dim=0)
dist.all_reduce(values)
if average:
values /= world_size
reduced_dict = {k: v for k, v in zip(names, values)}
return reduced_dict
def train_one_epoch(model, optimizer, data_loader, device, epoch, print_freq):
model.train()
metric_logger = MetricLogger(delimiter=" ")
metric_logger.add_meter('lr', SmoothedValue(window_size=1, fmt='{value:.6f}'))
header = 'Epoch: [{}]'.format(epoch)
lr_scheduler = None
if epoch == 0:
warmup_factor = 1. / 1000
warmup_iters = min(1000, len(data_loader) - 1)
lr_scheduler = warmup_lr_scheduler(optimizer, warmup_iters, warmup_factor)
for images, targets in metric_logger.log_every(data_loader, print_freq, header):
images = list(image.to(device) for image in images)
targets = [{k: v.to(device) for k, v in t.items()} for t in targets]
loss_dict = model(images, targets)
losses = sum(loss for loss in loss_dict.values())
# reduce losses over all GPUs for logging purposes
loss_dict_reduced = reduce_dict(loss_dict)
losses_reduced = sum(loss for loss in loss_dict_reduced.values())
loss_value = losses_reduced.item()
if not math.isfinite(loss_value):
print("Loss is {}, stopping training".format(loss_value))
print(loss_dict_reduced)
sys.exit(1)
optimizer.zero_grad()
losses.backward()
optimizer.step()
if lr_scheduler is not None:
lr_scheduler.step()
metric_logger.update(loss=losses_reduced, **loss_dict_reduced)
metric_logger.update(lr=optimizer.param_groups[0]["lr"])
return metric_logger
@torch.no_grad()
def evaluate(model, data_loader, device='cuda'):
n_threads = torch.get_num_threads()
torch.set_num_threads(1) # Does it nessesary? Who knows...
cpu_device = torch.device("cpu")
inference_res = []
model.eval()
for images, targets in data_loader:
images = list(img.to(device) for img in images)
if torch.cuda.is_available():
torch.cuda.synchronize()
outputs = model(images)
outputs = [{k: v.to(cpu_device) for k, v in t.items()} for t in outputs]
res = targets, outputs
inference_res.append(res)
torch.set_num_threads(n_threads)
return inference_res
def collate_fn(batch):
"""
This function helps when we have different number of object instances
in the batches in the dataset.
"""
return tuple(zip(*batch))
class ToTensor(object):
def __call__(self, image, target):
image = F.to_tensor(image)
return image, target
class Compose(object):
def __init__(self, transforms):
self.transforms = transforms
def __call__(self, image, target):
for t in self.transforms:
image, target = t(image, target)
return image, target
class RandomHorizontalFlip(object):
def __init__(self, prob):
self.prob = prob
def __call__(self, image, target):
if random.random() < self.prob:
height, width = image.shape[-2:]
image = image.flip(-1)
bbox = target["boxes"]
if bbox.shape[0]>0:
bbox[:, [0, 2]] = width - bbox[:, [2, 0]]
target["boxes"] = bbox
return image, target
class Resize(object):
def __init__(self, target_size):
self.target_size = target_size
def __call__(self, image, target):
old_height, old_width = image.shape[-2:]
image = F.resize(image, self.target_size, interpolation=F.InterpolationMode.BILINEAR)
new_height, new_width = image.shape[-2:]
bbox = target["boxes"]
if bbox.shape[0]>0:
bbox[:, [0, 2]] = bbox[:, [0, 2]] * new_width / old_width
bbox[:, [1, 3]] = bbox[:, [1, 3]] * new_height / old_height
target["boxes"] = bbox
return image, target
def get_transform(train, target_size):
transforms = []
# converts the image, a PIL image, into a PyTorch Tensor
transforms.append(ToTensor())
transforms.append(Resize(target_size))
if train:
# during training, randomly flip the training images
# and ground-truth for data augmentation
transforms.append(RandomHorizontalFlip(0.5))
return Compose(transforms)
# +
params = {}
params['target_size']=(1500,2000)
params['batch_size'] = 1
params['lr'] = 0.001
voc_root = '../data/TrainingData'
# +
# Reworked class from pytorch (see https://pytorch.org/vision/0.8/_modules/torchvision/datasets/voc.html#VOCDetection)
class LADDDataSET(torchvision.datasets.VisionDataset):
def __init__(
self,
root: str,
image_set: str,
transforms: Optional[Callable] = None):
super(LADDDataSET, self).__init__(root, transforms=transforms)
self.image_set = image_set
voc_root = root
image_dir = os.path.join(voc_root, 'JPEGImages')
annotation_dir = os.path.join(voc_root, 'Annotations')
if not os.path.isdir(voc_root):
raise RuntimeError('Dataset not found or corrupted.')
splits_dir = os.path.join(voc_root, 'ImageSets/Main')
split_f = os.path.join(splits_dir, image_set.rstrip('\n') + '.txt')
with open(os.path.join(split_f), "r") as f:
file_names = [x.strip() for x in f.readlines()]
self.images = [os.path.join(image_dir, x + ".jpg") for x in file_names]
self.annotations = [os.path.join(annotation_dir, x + ".xml") for x in file_names]
assert (len(self.images) == len(self.annotations))
def __getitem__(self, index: int) -> Tuple[Any, Any]:
"""
Args:
index (int): Index
Returns:
tuple: (image, target) where target is a dictionary of the XML tree.
"""
img = Image.open(self.images[index]).convert('RGB')
description = LADDDataSET.parse_voc_xml(
ET.parse(self.annotations[index]).getroot())
# get bounding box coordinates
num_objs = len(description['annotation']['object'])
boxes = []
for l in description['annotation']['object']:
bb = l['bndbox']
boxes.append([int(bb['xmin']), int(bb['ymin']), int(bb['xmax']), int(bb['ymax'])])
target = {}
target["boxes"] = torch.as_tensor(boxes, dtype=torch.float32) # there is only one class
target["labels"] = labels = torch.ones((num_objs,), dtype=torch.int64)
if self.transforms is not None:
img, target = self.transforms(img, target)
return img, target
def __len__(self) -> int:
return len(self.images)
@staticmethod
def parse_voc_xml(node: ET.Element) -> Dict[str, Any]:
voc_dict: Dict[str, Any] = {}
children = list(node)
if children:
def_dic: Dict[str, Any] = collections.defaultdict(list)
for dc in map(LADDDataSET.parse_voc_xml, children):
for ind, v in dc.items():
def_dic[ind].append(v)
if node.tag == 'annotation':
def_dic['object'] = [def_dic['object']]
voc_dict = {
node.tag:
{ind: v[0] if len(v) == 1 else v
for ind, v in def_dic.items()}
}
if node.text:
text = node.text.strip()
if not children:
voc_dict[node.tag] = text
return voc_dict
# +
# Pytorch implemenation of retinanet doesn't supports train on Images without any objects (which, probably need to be fixed)
# see https://github.com/pytorch/vision/blob/master/torchvision/models/detection/retinanet.py#L475
# As a temporary solution, yet, we just filtering out empty images
splits_dir = os.path.join(voc_root, 'ImageSets/Main')
annotation_dir = os.path.join(voc_root, 'Annotations')
with open(os.path.join(splits_dir,'train.txt'), "r") as f:
file_names = [x.strip() for x in f.readlines()]
non_empty = []
for a in file_names:
description = LADDDataSET.parse_voc_xml(
ET.parse(os.path.join(annotation_dir, a + ".xml")).getroot()
)
num_objs = len(description['annotation']['object'])
if num_objs > 0:
non_empty.append(a+'\n')
with open(os.path.join(splits_dir,'train_non_empty.txt'), "w") as f:
f.writelines(non_empty)
print('Total images '+str(len(file_names)), ' non empty: '+str(len(non_empty)))
# +
# test DS
im_idx = 99
dataset = LADDDataSET(voc_root,'test',get_transform(train=True,target_size=params['target_size']))
(image,target) = dataset[im_idx]
im = F.to_pil_image(image)
draw = ImageDraw.Draw(im)
for bb in target['boxes']:
draw.line([(bb[0], bb[1]), (bb[0], bb[3]), (bb[2], bb[3]),
(bb[2], bb[1]), (bb[0], bb[1])], width=4, fill=(255, 0, 0))
im.show()
# +
dataset_train = LADDDataSET(voc_root,'train_non_empty',get_transform(train=True,target_size=params['target_size']))
dataset_val = LADDDataSET(voc_root,'val',get_transform(train=False,target_size=params['target_size']))
# define training and validation data loaders
data_loader = torch.utils.data.DataLoader(
dataset_train, batch_size=params['batch_size'], shuffle=True, num_workers=4
,collate_fn=collate_fn
)
data_loader_val = torch.utils.data.DataLoader(
dataset_val, batch_size=1, shuffle=False, num_workers=4
,collate_fn=collate_fn
)
# -
model = torchvision.models.detection.fasterrcnn_resnet50_fpn(
pretrained=False,
num_classes=2,
pretrained_backbone=True,
min_size=params['target_size'][0],
max_size=params['target_size'][1],
trainable_backbone_layers = 0)
# +
device = torch.device('cuda')
model = model.to(device)
optimizer = torch.optim.SGD(
model.parameters(),
lr=params['lr'],
momentum=0.9,
weight_decay=0.0005)
lr_scheduler = torch.optim.lr_scheduler.StepLR(
optimizer,
step_size=3,
gamma=0.1)
# +
for epoch in range(2):
train_one_epoch(model, optimizer, data_loader, device, epoch, print_freq=100)
print ("Train done, evaluating.")
# update the learning rate
lr_scheduler.step()
# evaluate on the test dataset
inference_res = evaluate(model,data_loader_val)
print('Inference done, computing mAp : ')
print(evaluate_res(inference_res, iou_threshold = 0.5, score_threshold = 0.05))
print(evaluate_res(inference_res, iou_threshold = 0.6, score_threshold = 0.05))
print('Epoch Done')
torch.save(model.state_dict(), 'weights/resnet50_FRCNN_baseline.pth')
# +
dataset_test = LADDDataSET(voc_root,'test',get_transform(train=False,target_size=params['target_size']))
data_loader_test = torch.utils.data.DataLoader(
dataset_val, batch_size=1, shuffle=False, num_workers=1
,collate_fn=collate_fn
)
image_idx = 0
cpu_device = torch.device("cpu")
model.eval()
for images, targets in data_loader_test:
g_images = list(img.to(device) for img in images)
if torch.cuda.is_available():
torch.cuda.synchronize()
outputs = model(g_images)
outputs = [{k: v.to(cpu_device) for k, v in t.items()} for t in outputs]
res = targets, outputs
break
im = F.to_pil_image(images[image_idx])
targets
# im = to_pil_image(dataset[10][0])
draw = ImageDraw.Draw(im)
for idx in range(len(outputs[image_idx]['boxes'])):
width = math.ceil(outputs[image_idx]['scores'][idx]*10)
bb = outputs[0]['boxes'][idx]
draw.line([(bb[0], bb[1]), (bb[0], bb[3]), (bb[2], bb[3]),
(bb[2], bb[1]), (bb[0], bb[1])], width=width, fill=(255, 0, 0))
for bb in targets[image_idx]['boxes'][:10]:
draw.line([(bb[0], bb[1]), (bb[0], bb[3]), (bb[2], bb[3]),
(bb[2], bb[1]), (bb[0], bb[1])], width=4, fill=(0,255, 0))
im.show()
# -
|
cv-competition-1/pytorch_baseline/pytorch_baseline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Record Embedding
import pandas as pd
import numpy as np
import os
import calendar
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Embedding using fastText
# Details here: https://fasttext.cc/
# +
import warnings
warnings.filterwarnings('ignore')
import gensim
from gensim.models import FastText
# -
# Convert each tuple into a row
# +
#hospital dataset
#1. load hospital dataset and turthvalue.csv; mark error for unmatched values and cleanup using FastText; fill empty cells with word2vec
#finding errors and outliers -> discover FDs and try marking errors? or word2vec most_similar of val and if row does not contain the cell values??!
df_hospital = pd.read_csv('dirty_data_transformed.csv', dtype=object)
df_truth = pd.read_csv('truthvalue.csv', dtype=object)
provider = df_truth['ProviderNumber'].tolist()
hospital = df_truth['HospitalName'].tolist()
address = df_truth['Address1'].tolist()
city = df_truth['City'].tolist()
state = df_truth['State'].tolist()
zipcode = df_truth['ZipCode'].tolist()
county = df_truth['CountyName'].tolist()
phone = df_truth['PhoneNumber'].tolist()
hospType = df_truth['HospitalType'].tolist()
owner = df_truth['HospitalOwner'].tolist()
service = df_truth['EmergencyService'].tolist()
condition = df_truth['Condition'].tolist()
code = df_truth['MeasureCode'].tolist()
name = df_truth['MeasureName'].tolist()
score = df_truth['Score'].tolist()
sample = df_truth['Sample'].tolist()
stateavg = df_truth['Stateavg'].tolist()
combined_hosp = list(zip(provider, hospital, address, city, state, zipcode, county, phone, hospType, owner, service, condition, code, name, score, sample, stateavg))
# -
len(df_truth)
len(combined_hosp[1])
# +
#hospital dirty dataset; maintain dirty dataset separately
provider = df_hospital['ProviderNumber'].tolist()
hospital = df_hospital['HospitalName'].tolist()
address = df_hospital['Address1'].tolist()
city = df_hospital['City'].tolist()
state = df_hospital['State'].tolist()
zipcode = df_hospital['ZipCode'].tolist()
county = df_hospital['CountyName'].tolist()
phone = df_hospital['PhoneNumber'].tolist()
hospType = df_hospital['HospitalType'].tolist()
owner = df_hospital['HospitalOwner'].tolist()
service = df_hospital['EmergencyService'].tolist()
condition = df_hospital['Condition'].tolist()
code = df_hospital['MeasureCode'].tolist()
name = df_hospital['MeasureName'].tolist()
score = df_hospital['Score'].tolist()
sample = df_hospital['Sample'].tolist()
stateavg = df_hospital['Stateavg'].tolist()
combined_dirty = list(zip(provider, hospital, address, city, state, zipcode, county, phone, hospType, owner, service, condition, code, name, score, sample, stateavg))
len(combined_dirty)
# -
combined_dirty[1]
#Calculating error or dirty cells
dirty_list = []
truth_list = []
for i in range(len(combined_dirty)):
for j in range(len(combined_dirty[0])):
if combined_dirty[i][j] != combined_hosp[i][j]:
dirty_list.append(combined_dirty[i][j])
truth_list.append(combined_hosp[i][j])
combined_hosp[1]
#training the model with truth data.
model_hosp = FastText(combined_hosp, min_count=1, workers=8, iter=100)
#modelFast = load_model("amazonModelFastText.w2v")
model_hosp.most_similar("BIRMxNGHAM")
model_hosp.save("EmbeddingsFastText.txt")
model_hosp.save("EmbeddingsFastText.w2v")
# +
#module to predict the top value by the model.Some values are not broken into tokens.
#Values with keyerrors are stored in excluded_list
excluded_list = []
result = []
for i in range(len(dirty_list)):
if isinstance(dirty_list[i], str):
try:
result.append((model_hosp.most_similar(dirty_list[i])[0][0],dirty_list[i],truth_list[i]))
except KeyError:
excluded_list.append(dirty_list[i])
#print(dirty_list[i])
pass
# -
print (len(excluded_list))
print (len(result))
print (len(excluded_list) + len(result))
print (len(dirty_list))
excluded_list[0]
# +
#identify the true positives and false positives. Store them in the seperate list for further analysis.
true_pos = 0
false_pos = 0
correctly_predicted = []
falsely_predicted = []
for i in range(len(result)):
if result[i][0] == result[i][2]:
true_pos = true_pos + 1
correctly_predicted.append(result[i])
else:
false_pos = false_pos + 1
falsely_predicted.append(result[i])
# -
print(len(result))
print(len(correctly_predicted))
print(len(falsely_predicted))
#debugging wrong predictions
from gensim.models import KeyedVectors
model_hosp = KeyedVectors.load("EmbeddingsFastText.txt")
model_hosp.most_similar("3x640")
# +
correctly_predicted[8]
for i in falsely_predicted:
print (i)
# -
for i in correctly_predicted:
print (i)
falsely_predicted[1]
model_hosp.most_similar('3595x')
|
Embedding-FastText_hospital_results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import networkx as nx
import matplotlib.pyplot as plt
import numpy as np
import random
# +
def build_graph(edges,weights):
G = nx.Graph()
for i, e in enumerate(edges):
G.add_edge(*e, weight=weights[i])
return G
def generate_edges(G):
edges = []
for node in G:
for neighbour in G[node]:
edges.append((node, neighbour))
edges.append((neighbour, node))
return edges
def generate_weights(G):
weights = []
for node in G:
for neighbour in G[node]:
weights.append(G[node][neighbour]["weight"])
weights.append(G[node][neighbour]["weight"])
return weights
# -
def generate_tsp_path(G, origin):
tour = []
nodes_visited = []
actual_node = origin
while(len(nodes_visited) < G.number_of_nodes()):
min_distance = float('inf')
for n in G.neighbors(actual_node):
if G[actual_node][n]['weight'] < min_distance and n not in nodes_visited or len(nodes_visited) == G.number_of_nodes() - 1 and n == origin:
min_distance = G[actual_node][n]['weight']
next_node = n
next_edge = (actual_node,next_node)
nodes_visited.append(actual_node)
tour.append(next_edge)
actual_node = next_node
return tour
#return [(1,2),(2,3),(3,4),(4,5)] # example
def get_drawing_config(path, edges):
edges_config = [(4,'g') if n in path else (1,'b') for n in edges]
edges_config = np.array(edges_config)
edges_weights = list(edges_config[:,0])
edges_colors = list(edges_config[:,1])
edges_labels = nx.get_edge_attributes(G,'weight')
return (edges_weights, edges_colors, edges_labels)
# +
def random_colors(number_of_colors):
colors = ["#"+''.join([random.choice('0123456789ABCDEF') for j in range(6)])
for i in range(number_of_colors)]
return colors
if __name__ == "__main__":
# graph construction
#edges = [(1,2),(1,3),(1,4),(1,6),(2,3),(2,4),(2,5),(3,4),(3,5),(3,6),(4,5),(5,6),(4,6)] # example1
edges = [(1,2),(1,4),(1,6), (2,3), (2,4), (2,5), (3,4), (3,5)]
#weights = np.random.randint(low=1,high=12,size=len(edges))
#weights = [2,5, 3, 7, 5, 1, 5, 2,3, 7, 2, 2, 1] #example1
weights = [8, 3, 4, 1, 5, 9, 7, 2]
G = build_graph(edges,weights)
#edges1 = generate_edges(G)
#weights1 = generate_weights(G)
#print(edges1)
# tsp solving
path = generate_tsp_path(G,6)
print(path)
# drawing config
#edges1 = [(1,2),(2,1),(1,3),(3,1),(1,4),(4,1),(1,6),(6,1),(2,3),(2,3),(2,4),(4,2),(2,5),(5,2),(3,4),(4,3),(3,5),(5,3),(3,6),(6,3),(4,5),(5,4),(5,6),(6,5),(4,6),(6,4)]
#weights = [2, 2, 5, 5, 3, 3, 7, 7, 5, 5, 1, 1, 5, 5, 2, 2, 3, 3, 7, 7, 2, 2, 2, 2, 1, 1]
edge_weights, edge_colors, edge_labels = get_drawing_config(path,edges)
pos = nx.spring_layout(G, k=G.number_of_nodes()/len(edges))
node_mapping = dict(zip(G.nodes(),"abcdefg")) # example
node_colors = random_colors(G.number_of_nodes())
# graph plotting
plt.figure(1,figsize=(18,12))
nx.draw(G,pos, size=len(G.edges), labels=node_mapping, font_size=32, node_size=750,
node_color=node_colors, alpha=0.65, edge_color='b')
nx.draw_networkx_edge_labels(G,pos,font_size=21 , edge_labels=edge_labels)
plt.savefig('graph.png')
plt.close()
plt.figure(2,figsize=(18,12))
nx.draw(G,pos, size=len(G.edges), labels=node_mapping, font_size=32, node_size=750,
node_color=node_colors, alpha=0.65, width=edge_weights, edge_color=edge_colors)
nx.draw_networkx_edge_labels(G,pos,font_size=21 , edge_labels=edge_labels)
plt.savefig('solution.png')
plt.show()
# -
|
deprecated-tsp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Download** (right-click, save target as ...) this page as a jupyterlab notebook [Lab23](http://172.16.31.10/engr-1330-webroot/8-Labs/Lab23/Lab23_Class.ipynb)
#
# ___
# # <font color=darkred>Laboratory 23: KNN Classification </font>
#
# LAST NAME, FIRST NAME
#
# R00000000
#
# ENGR 1330 Laboratory 21 - In-Class and Homework
#
# ___
#
# ## Example: Iris Plants Classification <br>
#
# <img src="https://i.etsystatic.com/10589108/r/il/213b38/1876572420/il_570xN.1876572420_ikcm.jpg" width="200" align="left" style="padding-right: 20px">
#
# This is a well known problem and database to be found in the pattern recognition literature. Fisher's paper is a classic in the field and is referenced frequently to this day.
# The Iris Flower Dataset involves predicting the flower species given measurements of iris flowers.
#
# The Iris Data Set contains information on sepal length, sepal width, petal length, petal width all in cm, and class of iris plants.
# The data set contains 3 classes of 50 instances each, where each class refers to a type of iris plant.
# Hence, it is a multiclass classification problem and the number of observations for each class is balanced.
#
# <img src="https://miro.medium.com/max/1000/1*lFC_U5j_Y8IXF4Ga87KNVg.png" width="500"><br>
#
#
# Let's use a KNN model in Python and see if we can classifity iris plants based on the four given predictors.
#
# <hr>
#
# *__Acknowledgements__*
# 1. *Fisher,R.A. "The use of multiple measurements in taxonomic problems" Annual Eugenics, 7, Part II, 179-188 (1936); also in "Contributions to Mathematical Statistics" (John Wiley, NY, 1950).*
# 2. *Duda,R.O., & Hart,P.E. (1973) Pattern Classification and Scene Analysis. (Q327.D83) <NAME> & Sons. ISBN 0-471-22361-1. See page 218.*
# 3. *<NAME>. (1980) "Nosing Around the Neighborhood: A New System Structure and Classification Rule for Recognition in Partially Exposed Environments". IEEE Transactions on Pattern Analysis and Machine Intelligence, Vol. PAMI-2, No. 1, 67-71.*
# 4. *<NAME>. (1972) "The Reduced Nearest Neighbor Rule". IEEE Transactions on Information Theory, May 1972, 431-433.*
# 5. *See also: 1988 MLC Proceedings, 54-64. Cheeseman et al's AUTOCLASS II conceptual clustering system finds 3 classes in the data.*
# Load some libraries:
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import sklearn.metrics as metrics
import seaborn as sns
# %matplotlib inline
# Read the dataset and explore it using tools such as descriptive statistics:
# Read the remote directly from its url (Jupyter):
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data"
# Assign colum names to the dataset
names = ['sepal-length', 'sepal-width', 'petal-length', 'petal-width', 'Class']
# Read dataset to pandas dataframe
dataset = pd.read_csv(url, names=names)
dataset.tail(9)
dataset.describe()
# Split the predictors and target - similar to what we did for logisitc regression:
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, 4].values
# Then, the dataset should be split into training and testing. This way our algorithm is tested on un-seen data, as it would be in a real-world application. Let's go with a 80/20 split:
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
#This means that out of total 150 records:
#the training set will contain 120 records &
#the test set contains 30 of those records.
# -
# It is extremely straight forward to train the KNN algorithm and make predictions with it, especially when using Scikit-Learn. The first step is to import the `KNeighborsClassifier` class from the `sklearn.neighbors` library. In the second line, this class is initialized with one parameter, i.e. `n_neigbours`. This is basically the value for the K. There is no ideal value for K and it is selected after testing and evaluation, however to start out, 5 seems to be the most commonly used value for KNN algorithm.
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier(n_neighbors=5)
classifier.fit(X_train, y_train)
# The final step is to make predictions on our test data. To do so, execute the following script:
y_pred = classifier.predict(X_test)
# As it's time to evaluate our model, we will go to our rather new friends, confusion matrix, precision, recall and f1 score as the most commonly used discrete GOF metrics.
from sklearn.metrics import classification_report, confusion_matrix
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
cm = pd.DataFrame(confusion_matrix(y_test, y_pred))
sns.heatmap(cm, annot=True)
plt.title('Confusion matrix', y=1.1)
plt.ylabel('Predicted label')
plt.xlabel('Actual label')
# What if we had used a different value for K? What is the best value for K?
#
# One way to help you find the best value of K is to plot the graph of K value and the corresponding error rate for the dataset. In this section, we will plot the mean error for the predicted values of test set for all the K values between 1 and 50. To do so, let's first calculate the mean of error for all the predicted values where K ranges from 1 and 50:
# +
error = []
# Calculating error for K values between 1 and 50
# In each iteration the mean error for predicted values of test set is calculated and
# the result is appended to the error list.
for i in range(1, 50):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train, y_train)
pred_i = knn.predict(X_test)
error.append(np.mean(pred_i != y_test))
# -
# The next step is to plot the error values against K values:
plt.figure(figsize=(12, 6))
plt.plot(range(1, 50), error, color='red', linestyle='dashed', marker='o',
markerfacecolor='blue', markersize=10)
plt.title('Error Rate K Value')
plt.xlabel('K Value')
plt.ylabel('Mean Error')
# ## Exercise
# Everything above we did in class, but how about classifing a new measurement? Thats your lab exercise!
#
# A framework might be:
# - Use the database to train a classifier (done)
# - Explore the sturcture of `y_pred = classifier.predict(X_test)` This seems like the tool we want.
# - Once you know how to predict from the classifier, determine the classification of thge following sample
#
# |sepal-length|sepal-width|petal-length|petal-width|
# |---|---|---|---|
# |6.9|3.1|5.1|2.3|
|
8-Labs/Lab23/src-lessold/.ipynb_checkpoints/Lab23_Class-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sherkhan15/Random-Forest-Project-Lender-Score-/blob/master/Random_Forest_Project.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="aJc8ITW8qzAa" colab_type="text"
# # **Random Forest Project**
#
# ---
#
#
# For this project we will be exploring publicly available data from LendingClub.com. Lending Club connects people who need money (borrowers) with people who have money (investors). Hopefully, as an investor you would want to invest in people who showed a profile of having a high probability of paying you back. We will try to create a model that will help predict this.
#
#
# > We will use lending data from 2007-2010 and try to classify and predict whether or not the borrower paid back their loan in full.
#
#
#
#
#
# ***Here are what the columns represent:***
#
# ---
#
#
#
# *credit.policy: 1 if the customer meets the credit underwriting criteria of LendingClub.com, and 0 otherwise.
#
# *purpose: The purpose of the loan (takes values "credit_card", "debt_consolidation", "educational", "major_purchase", "small_business", and "all_other").
#
# *int.rate: The interest rate of the loan, as a proportion (a rate of 11% would be stored as 0.11).
#
# *Borrowers judged by LendingClub.com to be more risky are assigned higher interest rates.
#
# *installment: The monthly installments owed by the borrower if the loan is funded.
#
# *log.annual.inc: The natural log of the self-reported annual income of the borrower.
#
# *dti: The debt-to-income ratio of the borrower (amount of debt divided by annual income).
#
# *fico: The FICO credit score of the borrower.
#
# *days.with.cr.line: The number of days the borrower has had a credit line.
#
# *revol.bal: The borrower's revolving balance (amount unpaid at the end of the credit card billing cycle).
#
# *revol.util: The borrower's revolving line utilization rate (the amount of the credit line used
# *relative to total credit available).
#
# *inq.last.6mths: The borrower's number of inquiries by creditors in the last 6 months.
#
# *delinq.2yrs: The number of times the borrower had been 30+ days past due on a payment in the past 2 years.
#
# *pub.rec: The borrower's number of derogatory public records (bankruptcy filings, tax liens, or judgments).
#
#
#
#
#
#
#
#
# + [markdown] id="OY-Wcd37u0od" colab_type="text"
# # *Import Libraries*
# ***Import the usual libraries for pandas and plotting. You can import sklearn later on.***
# + id="oPkb1gRUt_vp" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from google.colab import files
uploaded = files.upload()
# + [markdown] id="gNhVTUi8u99c" colab_type="text"
#
# # Loading the Data
# + id="HSE6BGMclQ7W" colab_type="code" colab={}
loans=pd.read_csv('loan_data.csv')
# + [markdown] id="X2SQAdCZvAk9" colab_type="text"
#
# **Check out the info(), head(), and describe() methods on loans.**
# + id="GE4LYaa7mePg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 355} outputId="258617f1-633a-47c3-fa3f-8cca444c3c54"
loans.info()
# + id="DViZC1HnmfAI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 220} outputId="8950bdcc-f27e-4172-8d6d-32b207b1383d"
loans.head()
# + id="DkiD_i6MmfT1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 311} outputId="6178e308-e31f-4319-b5ff-0b8d9008c766"
loans.describe()
# + id="dpmvRpulmfm1" colab_type="code" colab={}
# + [markdown] id="XuLb0GZRvRCJ" colab_type="text"
# # Exploratory Data Analysis
# Let's do some data visualization! We'll use seaborn and pandas built-in plotting capabilities
#
# **Creating a histogram of two FICO distributions on top of each other, one for each credit.policy outcome.**
# + id="CoNO4AfFmf2q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="e403b42c-ded4-4083-a17e-87e07f49eb4a"
loans[loans['credit.policy']==1]['fico'].hist(alpha=0.5,color='blue',bins=30,label='Credit.Policy=1')
loans[loans['credit.policy']==0]['fico'].hist(alpha=0.5,color='red',bins=30,label='Credit.Policy=0')
plt.legend()
plt.xlabel('FICO')
# + [markdown] id="8NhH2O-hvgX5" colab_type="text"
# **Creating a histogram of two FICO distributions on top of each other, one for each "not fully.paid" column**
# + id="i2Ic6XRMvlq1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="62bbd513-942b-479c-aca8-f8e2d0e4e74f"
loans[loans['not.fully.paid']==1]['fico'].hist(alpha=0.5,color='blue',bins=30,label='not fully.paid=1')
loans[loans['not.fully.paid']==0]['fico'].hist(alpha=0.5,color='red',bins=30,label='not fully.paid=0')
plt.legend()
plt.xlabel('FICO')
# + [markdown] id="CTVQ2r1nv25Z" colab_type="text"
# **Creating a countplot using seaborn showing the counts of loans by purpose, with the color hue defined by not.fully.paid.**
# + id="gsZTNukNv8mu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 464} outputId="4148576d-ddd2-4fc5-f9ab-9348833551b3"
plt.figure(figsize=(11,7))
sns.countplot(x='purpose', hue='not.fully.paid', data=loans,palette='Set1')
# + id="4jMjMPz0wB2f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="fcb874c6-b409-4669-fc4a-757d6190489e"
sns.jointplot(x='fico', y='int.rate', data=loans, kind='scatter')
# + [markdown] id="Vnf1Jc2MwNKb" colab_type="text"
# Creating the following lmplots to see if the trend differed between not.fully.paid and credit.policy
# + id="8HSd-4qJwRaa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 387} outputId="64335277-4265-4b0d-fe69-acb8e99656d6"
sns.lmplot(x='fico',y='int.rate',data=loans,hue='credit.policy', col='not.fully.paid',palette='Set1')
# + id="qEbDeF_5xP68" colab_type="code" colab={}
loans['purpose'].unique()
# + id="T1C3leLbxUZp" colab_type="code" colab={}
change=['purpose']
# + id="UQEczZtZxW_9" colab_type="code" colab={}
#splitting column "purpose" into 6 distinctive columns
final_data=pd.get_dummies(loans,columns=change, drop_first=True )
# + id="7xTDeSdQxZh_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 444} outputId="c343bf43-d543-4715-94cc-cf494b5157b2"
final_data.info()
# + [markdown] id="5sTcsy2PxeNe" colab_type="text"
#
# **Train Test Split**
# + id="Ri3eTCtpxgk1" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
# + id="h_wWxcDJxokg" colab_type="code" colab={}
X = final_data.drop('not.fully.paid',axis=1)
y = final_data['not.fully.paid']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=101)
# + [markdown] id="bfT-ohFUxsIa" colab_type="text"
#
# **Training a Decision Tree Model**
# + id="NtJFk_IpyMwK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 220} outputId="f9a03640-bee3-424b-fed2-179a7ac27982"
from sklearn.tree import DecisionTreeClassifier
loans.head()
# + [markdown] id="UN0SSZyNyPxn" colab_type="text"
#
# **Creating an instance of DecisionTreeClassifier() called dtree and fitting it to the training data.**
# + id="02tJbOrLykz6" colab_type="code" colab={}
# + id="VgfP0vt-yUXy" colab_type="code" colab={}
dtree=DecisionTreeClassifier()
# + id="4ShlmkpkyZNE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 124} outputId="e20b6bca-8847-4c57-d323-fba287bfaa3d"
dtree.fit(X_train,y_train)
# + [markdown] id="m_6PxsDZyb1B" colab_type="text"
# **# Predictions and Evaluation of Decision Tree**
#
# **Creating predictions from the test set and creating a classification report and a confusion matrix.**
# + id="b-G28ElqylwR" colab_type="code" colab={}
predictions=dtree.predict(X_test)
# + id="hJoukrAFyqhX" colab_type="code" colab={}
from sklearn.metrics import classification_report,confusion_matrix
# + id="YTeuBUJyysi7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 177} outputId="be75f3d9-8796-444a-957e-2a511b999b62"
print(classification_report(y_test,predictions))
# + id="KRdOj4k8yvcW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="ed007702-94c0-4f79-d84b-ef29b1b7986c"
print(confusion_matrix(y_test,predictions) )
# + [markdown] id="9OieVJhgy9M6" colab_type="text"
# **Training the Random Forest model**
#
# *Creating an instance of the RandomForestClassifier class and fitting it to our training data from the previous step*
# + id="apoyYPldy6ZA" colab_type="code" colab={}
from sklearn.ensemble import RandomForestClassifier
# + id="VFu9vcUHy8ai" colab_type="code" colab={}
rfc=RandomForestClassifier(n_estimators=600)
# + id="9WlVeXTbzOeH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="2eac5862-c197-46e3-b5e2-eaeaa32b306f"
rfc.fit(X_train,y_train)
# + [markdown] id="F-FU-p7_zRWv" colab_type="text"
# # Predictions and Evaluation
# **Predicting y_test values**
# + id="9gof8IoHzW0m" colab_type="code" colab={}
predictions=rfc.predict(X_test)
# + id="Z5ozoqFZzaLY" colab_type="code" colab={}
from sklearn.metrics import classification_report,confusion_matrix
# + id="Ny7vg2lhzcW-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 177} outputId="159d197d-f1ae-4706-a594-8845bccb1f15"
print(classification_report(y_test,predictions))
# + id="pk-DMhsBzeu5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="8e91d46e-29bc-4927-9dc4-c3a4dc0248a6"
print(confusion_matrix(y_test,predictions))
|
Random_Forest_Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from GENIE3 import *
sizen = 200
# +
fname='/home/linaiqi/Lab/GPGene-TCBB/data/synthetic/synnet_and_data.txt'
data0 = loadtxt(fname,skiprows=1)
f = open(fname)
gene_names = f.readline()
f.close()
gene_names = gene_names.rstrip('\n').split('\t')
# -
data = data0[:sizen]
data.shape
# VIM = GENIE3(data)
# +
# # tree_method='RF'
# tree_method='ET'
# # Number of randomly chosen candidate regulators at each node of a tree
# K = 10
# # Number of trees per ensemble
# ntrees = 50
# # Run the method with these settings
# VIM3 = GENIE3(data,tree_method=tree_method,K=K,ntrees=ntrees)
# -
tree_method='RF'
# tree_method='ET'
# Number of randomly chosen candidate regulators at each node of a tree
K = 'sqrt'
# Number of trees per ensemble
ntrees = 1000
# Run the method with these settings
VIM3 = GENIE3(data,tree_method=tree_method,K=K,ntrees=ntrees)
# help(GENIE3)
# get_link_list(VIM)
res=get_link_list(VIM3,gene_names=gene_names,file_name='./tmp.txt')
# ! unset DISPLAY && java -jar ~/Lab/GNW/sandbox/gnw3-standalone.jar --evaluate --goldstandard /home/linaiqi/Lab/GPGene-TCBB/data/synthetic/synnet_and.tsv --prediction ./tmp.txt
|
eval/synnet_and_genie3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="_55E3-GVHuGx"
# # Get the data
# + colab={"base_uri": "https://localhost:8080/"} id="TNqqoz0-HFVC" outputId="235e6ea6-a57a-429c-d727-b51bdbe86adf"
# !gdown --id 1HVSazFk8m553VWPjFnZZ-YfJA_KecPea
# !unzip translated_data_updated.zip
# + id="OL-S8VmBy1tU"
# %%capture
# !pip install tensorflow_decision_forests
# + colab={"base_uri": "https://localhost:8080/"} id="6zhXcytVJq1w" outputId="b8878625-1d10-41bb-f2a2-f68974522cb3"
# imports
import pandas as pd
import seaborn as sns
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_decision_forests as tfdf
from tensorflow import keras
import sklearn
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
sns.set_theme(context='notebook', style='darkgrid')
mpl.rcParams['figure.figsize'] = (12, 10)
colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
# + id="66-dDHVzJ5NY"
# Important Note:
# Visits = browsing history in the training period. No test set available.
# Purchases = purchase history in the training period. No test set available.
df_users = pd.read_csv('data_translated/user_list.csv')
df_c_list_train = pd.read_csv('data_translated/coupon_list_train.csv')
df_c_list_test = pd.read_csv('data_translated/coupon_list_test.csv')
df_area_train = pd.read_csv('data_translated/coupon_area_train.csv')
df_area_test = pd.read_csv('data_translated/coupon_area_test.csv')
df_visit_train = pd.read_csv('data_translated/coupon_visit_train.csv')
df_purch_train = pd.read_csv('data_translated/coupon_detail_train.csv')
df_locations = pd.read_csv('data_translated/prefecture_locations.csv')
# + [markdown] id="yGhtMkAtJEQg"
# ## Feature Engineering
# Since TF Decision Forests can handle categorical variables just fine, we're not doing much preprocessing.
# + id="h3HNyFI7wmVf"
# rename SEX_ID column, change to categorical value (0 Male, 1 Female)
df_users['SEX'] = df_users['SEX_ID'].replace('f', 1)
df_users['SEX'] = df_users['SEX'].replace('m', 0)
# + id="Ekefadt28NG3"
# create a categorical variable for age group:
# 14-21, 22-35, 36-49, 50-65, 66-75, 76-90
def age_cat(age):
if age <= 21:
return 0
elif age <= 35:
return 1
elif age <= 49:
return 2
elif age <= 65:
return 3
elif age <= 75:
return 4
elif age <= 90:
return 5
else:
return 6
lbl_age_ranges = ['14-21', '22-35', '36-49', '50-65', '66-75', '76-90']
df_users['AGE_GROUP'] = [age_cat(a) for a in df_users['AGE']]
# + id="xYd_AMTSAVt7"
# Model Input Features
# For each user who purchased a coupon...
# Gender, Age, Prefecture, Coupon Genre, Coupon Prefecture, Price Rate, Catalog Price, Discount Rate, Ken Name
#############################
# BUILD DF_TRAIN DATAFRAME #
#############################
df_visit_train = df_visit_train.rename(columns={'VIEW_COUPON_ID_hash': 'COUPON_ID_hash'})
df_train = df_visit_train.join(df_users.set_index('USER_ID_hash'), on='USER_ID_hash', lsuffix='_v')
df_train = df_train.join(df_c_list_train.set_index('COUPON_ID_hash'), on='COUPON_ID_hash', rsuffix='_c')
# get a subset of the training columns and rename them
df_train = df_train[['AGE_GROUP', 'SEX', 'PREF_NAME_EN', 'KEN_NAME_EN', 'GENRE_NAME_EN', 'CAPSULE_TEXT_EN', 'PRICE_RATE', 'DISCOUNT_PRICE', 'PURCHASE_FLG']]
df_train.columns = ['age_group', 'sex', 'user_prefecture', 'coupon_prefecture', 'genre', 'capsule', 'discount_rate', 'discount_price', 'purchased']
# NaN preprocessing
# + colab={"base_uri": "https://localhost:8080/"} id="jahgKemcSyv3" outputId="0e2f5ca5-51c9-4647-ac08-f2f235cf6de6"
df_train.info()
# + colab={"base_uri": "https://localhost:8080/"} id="H2THoEJMxGAq" outputId="5b5ea913-2062-4322-973a-44dbf6550e1a"
# Train the model!
#df_train_set, df_test_set = train_test_split(df_train, test_size=0.2, stratify=df_train['purchased'])
ds_train_set = tfdf.keras.pd_dataframe_to_tf_dataset(df_train, label='purchased')
model = tfdf.keras.GradientBoostedTreesModel(num_trees=500,
growing_strategy='BEST_FIRST_GLOBAL',
max_depth=8, split_axis='SPARSE_OBLIQUE')
model.fit(ds_train_set)
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="u-_jHw8p_2at" outputId="5d39eb7c-afc5-42f2-c35b-9cc6a6838247"
# START HERE - run cells 108-114
tfdf.model_plotter.plot_model_in_colab(model, tree_idx=0)
# + [markdown] id="Vsx-v86C-J5e"
# #### Get User's Purchased Coupons
# + id="bpIDFsinMjic"
# preprocess the test set to make it a little faster
test_coupons = df_c_list_test
test_coupons = test_coupons[['PRICE_RATE', 'DISCOUNT_PRICE', 'COUPON_ID_hash', 'CAPSULE_TEXT_EN', 'GENRE_NAME_EN', 'KEN_NAME_EN']]
coupon_ids = test_coupons['COUPON_ID_hash']
def merge_user_with_test_coupons(user):
df = pd.DataFrame()
df['user_id'] = user['USER_ID_hash']
df['coupon_id'] = test_coupons['COUPON_ID_hash']
df['age_group'] = user['AGE_GROUP']
df['sex'] = user['SEX']
df['user_prefecture'] = np.array(user['PREF_NAME_EN']).astype(np.object)
df['coupon_prefecture'] = test_coupons['KEN_NAME_EN']
df['genre'] = test_coupons['GENRE_NAME_EN']
df['capsule'] = test_coupons['CAPSULE_TEXT_EN']
df['discount_rate'] = test_coupons['PRICE_RATE']
df['discount_price'] = test_coupons['DISCOUNT_PRICE']
df['sex'] = df['sex'].replace('m', 0)
df['sex'] = df['sex'].replace('f', 1)
return df
# + colab={"base_uri": "https://localhost:8080/", "height": 434} id="hMF4PzFiOsn_" outputId="4934fbc9-760a-44a0-9ff5-489d6ce3e7d1"
from tqdm import tqdm
all_predictions = []
for i, u in tqdm(df_users.iterrows(), total=len(df_users)):
user_coupons = merge_user_with_test_coupons(u)
ds_user_coupons = tfdf.keras.pd_dataframe_to_tf_dataset(user_coupons.drop(columns=['user_id', 'coupon_id']))
preds = model.predict(ds_user_coupons)
preds = preds.ravel()
df_pred = pd.DataFrame(data={'coupon_id': coupon_ids, 'likelihood': preds}, columns=['coupon_id', 'likelihood'])
top_coupons = df_pred.sort_values(by='likelihood', ascending=False)[:10]
coupon_string = ' '.join(top_coupons['coupon_id']).strip()
all_predictions.append({'USER_ID_hash': u['USER_ID_hash'], 'PURCHASED_COUPONS': coupon_string})
submission_df = pd.DataFrame.from_dict(all_predictions)
submission_df.to_csv('submission_decision_tree.csv', header=True, index=False)
submission_df
# + id="zgCasOIBA8Gi"
submission_df.to_csv('submission_gradient_boosted_hp.csv', header=True, index=False)
# + id="XSADJfv8BAy9"
|
notebooks/ipynb/BruteForce - Gradient Boosted Trees.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/martinfinis/HelloCodeSchoolProject/blob/master/covid_19_V2_Exploratory_Data_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="M-qejTAAaFiN"
# # Loading data
# + id="Rq5kGdgfTh45"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import glob
import time
# + id="c7ULD-yTD_0t" colab={"base_uri": "https://localhost:8080/"} outputId="3f69eeca-bb15-4afc-b84d-91ef4bc4b66d"
# mount google drive to access dataset food
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="Pa2Tr9QW626v"
# !rm -r curated_data
# !rm meta_data_cap.csv
# !rm meta_data_covid.csv
# !rm meta_data_normal.csv
# + id="naIEkdnjEVUF" colab={"base_uri": "https://localhost:8080/"} outputId="d3bae9bf-d4e8-4992-8720-1640ca65a0dc"
# %cd /content/
# !unzip '/content/drive/.shortcut-targets-by-id/1lp7vo6EG60jba1Gk_tuNpvDm7q5ZXlKX/Telecom_Specialization_ML3/Students/Peter_Christoph_Marko_Martin/data/archive.zip'
# + id="jYFH_-SOqDAy"
##### Inconsistency between file names and file names in meta data #####
## ==> rename files to the names used in meta_data_covid.csv
# !mv "/content/curated_data/curated_data/2COVID/17_Zhao_Recurrence-of-positive-SARS-CoV-2-RNA-in-C_2020_International-Journal-of-Inf-p1-21%0.png" "/content/curated_data/curated_data/2COVID/17_Zhao_Recurrence-of-COVID-19-SARS-CoV-2-RNA-in-C_2020_International-Journal-of-Inf-p1-21%0.png"
# !mv "/content/curated_data/curated_data/2COVID/17_Zhao_Recurrence-of-positive-SARS-CoV-2-RNA-in-C_2020_International-Journal-of-Inf-p1-21%1.png" "/content/curated_data/curated_data/2COVID/17_Zhao_Recurrence-of-COVID-19-SARS-CoV-2-RNA-in-C_2020_International-Journal-of-Inf-p1-21%1.png"
# + id="Psd9sPj7UKTT"
#17_Zhao_Recurrence-of-COVID-19-SARS-CoV-2-RNA-in-C_2020_International-Journal-of-Inf-p1-21%0
# + [markdown] id="xmtMMvbCVk1i"
# # Metadata-Analyse
# + id="-JoSWzwG4ZOX"
# Load data as dataframes
df_cap = pd.read_csv('meta_data_cap.csv')
df_covid = pd.read_csv('meta_data_covid.csv', encoding = "ISO-8859-1")
df_normal = pd.read_csv('meta_data_normal.csv')
# + [markdown] id="C9NekGUB4AiU"
# ### Analysis of df_cap
# + colab={"base_uri": "https://localhost:8080/", "height": 884} id="skcnpBBB4AiY" outputId="0e9756e4-7265-4dc1-93ab-f1c62fcfb9d2"
# Get some information about df_cap
print(df_cap.info())
display(df_cap.head())
# + colab={"base_uri": "https://localhost:8080/", "height": 120} id="f176KHMm4Aib" outputId="e8de10ad-a8e6-43b5-daab-d46bad6a7ea6"
df_cap.loc[df_cap.duplicated(),:] # get duplicates
# + colab={"base_uri": "https://localhost:8080/"} id="o8pUXcLU4Aid" outputId="09649e2b-0bb8-482e-9797-74aff7d324ac"
df_cap['Country'].unique() # unique countries
# + colab={"base_uri": "https://localhost:8080/"} id="IkfnNUjA4Aie" outputId="b6d423dd-e6e6-4ff7-95bf-9b959b42d69d"
df_cap['Patient ID'].nunique() # number of unique patients
# + colab={"base_uri": "https://localhost:8080/"} id="0oH3mI3J4Aif" outputId="210a7e51-b202-4a82-9799-abc8c8c954c9"
df_cap['Age'].unique() # unique ages
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="fhQSVdc_4Aig" outputId="c5d1d7c0-f0cc-4629-d5ce-a9404bb4fa4f"
# Display age distribution
df_cap.Age.plot(kind='hist')
plt.show()
# + [markdown] id="xJFKVdiny8z7"
# ##Analysis of df_normal
# + colab={"base_uri": "https://localhost:8080/", "height": 365} id="Lp17iuVaajPP" outputId="7bded768-1bab-4b73-e78e-ce21003e6da9"
#44 colums 21 one with NaN
df_normal = df_normal.dropna(axis=1, how='all')
df_normal.head(3)
# + colab={"base_uri": "https://localhost:8080/"} id="8d5oSo5mZnOu" outputId="18d527c7-c959-4f17-f7ad-92340d949542"
#after removen nan colums
df_normal.info()
# + [markdown] id="9EPXol9pels9"
# ## Analysis of df_covid
# + colab={"base_uri": "https://localhost:8080/", "height": 120} id="nHWNDoSEgwJx" outputId="5f8a815d-b204-4131-d347-9b43400a5b75"
df_covid.loc[df_covid.duplicated(),:] # get duplicates
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="WotgSpKGeozC" outputId="b5ad6c4c-e880-4243-f9df-7fcf26863f4e"
# Get some information about df_covid
print(df_covid.info())
display(df_covid.head())
# + colab={"base_uri": "https://localhost:8080/"} id="rY6beaAIgA1a" outputId="b6bf8761-5984-4e93-ddcf-67e0bcda7212"
df_covid['Country'].unique() # unique countries
# + colab={"base_uri": "https://localhost:8080/"} id="uTAwWL8mgIZp" outputId="26636f23-3749-4ce4-c7b2-367580d3f58e"
df_covid['Patient ID'].nunique() # number of unique patients
# + colab={"base_uri": "https://localhost:8080/"} id="HkMEJRS7IQ84" outputId="c8371333-629e-4274-b739-382c3373eaac"
# the Age column contains a - signa -> age unknown
df_covid['Age'].unique()
# + id="9KtYA069xmvU"
df = df_cap[['Patient ID','Age']]
# + [markdown] id="Dt33fQfcxQMZ"
# ### map '-' into 4711 as key for further plot taks
# + id="itZUSVBPIHYZ"
# clean '-' and add 4711 as key for further plot
df_covid['Age'] = df_covid['Age'].apply(lambda x: 4711 if x == '-' else x)
# + colab={"base_uri": "https://localhost:8080/"} id="_NAf8wUcIcXA" outputId="4a05d123-5d0e-4df7-bc8d-8dc10c269234"
df_covid['Age'].unique()
# + id="_x1dYUxyT62A"
df_covid['Age'] = df_covid['Age'].astype(int)
# + [markdown] id="BGvXdaCqv_Hq"
# #merge reports for statistic plot
# + colab={"base_uri": "https://localhost:8080/"} id="Dend3Kxhp18y" outputId="79b085ce-9bdb-4e0a-9e83-3634c6321c8f"
covid_name = df_covid.columns.tolist()
covid_name.sort()
print(covid_name)
# + colab={"base_uri": "https://localhost:8080/"} id="h5wwwO-Sm7wH" outputId="ea59ac9e-8db8-4bdb-b405-42c8c270419e"
normal_name = df_normal.columns.tolist()
normal_name.sort()
print(normal_name)
# + colab={"base_uri": "https://localhost:8080/"} id="sYT33HX1rPIT" outputId="cdc6f7f8-b45b-43c0-a945-986e0f3de29f"
cap_name = df_cap.columns.tolist()
cap_name.sort()
print(cap_name)
# + id="-PZUqMMfrsu7"
#the choosen columns
importend_columns = ['Patient ID','Diagnosis','Gender','Age','Country','File name']
# + colab={"base_uri": "https://localhost:8080/"} id="7MqGmhe_myR4" outputId="4c129b27-cb96-42f3-e027-a31ab546d932"
#check if all columns are present
set(importend_columns).issubset(cap_name)
# + colab={"base_uri": "https://localhost:8080/"} id="dri_x5EQtX77" outputId="fbef1af9-bef6-4e9a-ab4f-155527289d4e"
set(importend_columns).issubset(normal_name)
# + colab={"base_uri": "https://localhost:8080/"} id="_LhmM1oVtbrc" outputId="a14cdfca-6947-475c-d693-3c073043d8da"
set(importend_columns).issubset(covid_name)
# + id="V78h5N63tgHD"
df_all = df_cap[importend_columns]
# + id="K-XWVGRcuJ1y"
df_all = pd.concat([df_all, df_covid[importend_columns]], ignore_index=True)
# + id="r7XPP7rpvgub"
df_all = pd.concat([df_all, df_normal[importend_columns]], ignore_index=True)
# + colab={"base_uri": "https://localhost:8080/"} id="Ha-qrJIBvwoS" outputId="e202f72a-cec3-41a3-9f4e-603dad07748b"
df_all.info()
# + colab={"base_uri": "https://localhost:8080/"} id="hG0DY9zm2W61" outputId="2b972238-029d-4d48-a5ad-05d7f8c3d85a"
df_all['Diagnosis'].unique()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="4S0tgxYJ8XlB" outputId="80c3acfc-0ab4-4b34-92d4-301884fc0577"
#small methode for mapping the age into age groups
#90 -> between 81 - 90
age_group_lst =[10,20,30,40,50,60,70,80,90,120,4711]
age_group_name =['1-10','11-20','21-30','31-40','41-50','51-60','61-70','71-80','81-90','>91','unk']
def age_group(x):
for i,age in zip(range(len(age_group_lst)),age_group_lst):
if x <= age:
return age_group_name[i]
#result 55 -> between 51 - 60
#result 82 -> between 81 - 90
age_group(89)
# + id="JffwAb0c-g2T"
df_all['age_group'] = df_all['Age'].apply(age_group)
# + id="AI3wHBByPLG1"
#len(df_all['Patient ID'].unique())
# + id="l3qk3G6vPD9A"
# remove all duplicate patient information
df_all = df_all.groupby('Patient ID', as_index=False).last()
# + colab={"base_uri": "https://localhost:8080/", "height": 407} id="PN_kUoSUPsm-" outputId="5b17f81f-0577-4d43-be00-0ea75747fcab"
df_all
# + [markdown] id="btOzD319DG1L"
# ## preparing age plot
# + id="3DQpjqKILeCE"
#modify the given matplotlib example for our purpose
#https://matplotlib.org/stable/gallery/lines_bars_and_markers/barchart.html
# + id="d_SzT7BINRQi"
df_groupby_dia_age = df_all.groupby(['Diagnosis','age_group']).agg(count_per_age_group=('age_group','count'))
# + colab={"base_uri": "https://localhost:8080/", "height": 978} id="QzdkeWg7RmxR" outputId="26d61c54-49e5-4cb5-cdc5-733a70f31f58"
df_groupby_dia_age
# + id="fdEVrrmOQgCH"
df_for_plot = df_groupby_dia_age.reset_index()
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="C8j4iyNLf2ts" outputId="8b944e04-32c7-45ab-a773-c977432a7ca5"
df_for_plot.head()
# + colab={"base_uri": "https://localhost:8080/"} id="9RbYo1tyw1ZL" outputId="cc11da24-04b2-4087-f937-0b1fbe9974a0"
df_for_plot['count_per_age_group'].sum()
# + colab={"base_uri": "https://localhost:8080/"} id="2oYNOw-Nf_Lr" outputId="a2f648c4-478b-408f-f49d-126caa2d638b"
#mapping age groups for df in list
labels = ['1-10','11-20','21-30','31-40','41-50','51-60','61-70','71-80','81-90','>91','unk']
covid_n = [0] * len(labels)
cap_n = [0] * len(labels)
normal_n = [0] * len(labels)
for _, row in df_for_plot.iterrows():
i = labels.index(row['age_group'])
if row['Diagnosis'] == 'CAP':
cap_n[i] = row['count_per_age_group']
elif row['Diagnosis'] == 'COVID-19':
covid_n[i] = row['count_per_age_group']
elif row['Diagnosis'] == 'Normal':
normal_n[i] = row['count_per_age_group']
else:
print("should nerver happen")
covid_n,cap_n,normal_n
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="VulaY1FokPI8" outputId="f6d4e0a0-88cc-4a51-c9ee-2080c75692d7"
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
#modify the given matplotlib example for our purpose
#https://matplotlib.org/stable/gallery/lines_bars_and_markers/barchart.html
x = np.arange(len(labels)) # the label locations
width = 0.8 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(x - width/3, normal_n, width/3, label='normal_n')
rects2 = ax.bar(x , covid_n, width/3, label='covid_n')
rects2 = ax.bar(x + (width/3), cap_n, width/3, label='cap_n')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('count')
ax.set_title('age range of patients for different categories')
ax.set_xticks(x)
ax.set_xticklabels(labels)
ax.legend()
fig.tight_layout()
plt.show()
# + [markdown] id="dClcKChUDQIu"
# ## preparing country plot
# + colab={"base_uri": "https://localhost:8080/", "height": 138} id="Z2L7UkmBDTzd" outputId="60849b19-9b39-410f-cfdc-ff4b5ed847ca"
df_all.head(3)
# + id="YN9fgUoPDt8c"
df_groupby_dia_country = df_all.groupby(['Diagnosis','Country']).agg(count_per_county=('Country','count'))
# + colab={"base_uri": "https://localhost:8080/", "height": 588} id="dcW_5xTVDt8d" outputId="38474d39-fec0-4c1b-f61a-c2ec3f235a49"
df_groupby_dia_country
# + id="iI4EG9NnD2d-"
df_groupby_dia_country = df_groupby_dia_country.reset_index()
# + colab={"base_uri": "https://localhost:8080/", "height": 168} id="fk0QVgCnGYu0" outputId="1baec823-aef0-4039-f778-c1a002c6082e"
df_groupby_dia_country.head(4)
# + colab={"base_uri": "https://localhost:8080/"} id="i1Vxe6xiEwFu" outputId="02ee7c96-c685-4de3-c140-f2b6997ed721"
#reading labels for country plot out of df
labels = df_groupby_dia_country['Country'].unique()
labels.sort()
labels = labels.tolist()
#mapping age groups for df in list
covid_n = [0] * len(labels)
cap_n = [0] * len(labels)
normal_n = [0] * len(labels)
for _, row in df_groupby_dia_country.iterrows():
i = labels.index(row['Country'])
if row['Diagnosis'] == 'CAP':
cap_n[i] = row['count_per_county']
elif row['Diagnosis'] == 'COVID-19':
covid_n[i] = row['count_per_county']
elif row['Diagnosis'] == 'Normal':
normal_n[i] = row['count_per_county']
else:
print("should nerver happen")
covid_n,cap_n,normal_n
# + id="eI9N45zDIl-_"
#replace - with unk (unknown)
labels[0] ='unk'
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="dmEBX_DzG35b" outputId="1e4fcc09-9883-442f-9604-e1b9110f7d4e"
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
#modify the given matplotlib example for our purpose
#https://matplotlib.org/stable/gallery/lines_bars_and_markers/barchart.html
x = np.arange(len(labels)) # the label locations
width = 0.8 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(x - width/3, normal_n, width/3, label='normal_n')
rects2 = ax.bar(x , covid_n, width/3, label='covid_n')
rects2 = ax.bar(x + (width/3), cap_n, width/3, label='cap_n')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('count')
ax.set_title('number of patients for different countrys')
ax.set_xticks(x)
ax.set_xticklabels(labels,rotation = 45)
ax.legend()
fig.tight_layout()
plt.show()
|
covid_19_V2_Exploratory_Data_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 14.1 - 14.3
# Compute using $h=2^{-1}, 2^{-2}, \dots 2^{-5}$ and the forward, backward, and centered difference approximations the following derivatives.
# - $f(x) = \sqrt{x}$ at $x=0.5$. The answer is $f'(0.5) = 2^{-1/2} \approx 0.70710678118$.
# - $f(x) = \arctan(x^2 - 0.9 x + 2)$ at $x=0.5$. The answer is $f'(0.5) = \frac{5}{212}$.
# - $f(x) = J_0(x),$ at $x=1$, where $J_0(x)$ is a Bessel function of the first kind given by $$ J_\alpha(x) = \sum_{m=0}^\infty \frac{(-1)^m}{m! \, \Gamma(m+\alpha+1)} {\left(\frac{x}{2}\right)}^{2m+\alpha}. $$ The answer is $f'(1) \approx -0.4400505857449335.$
#
# ## Solution
# This problem is split up into the three given derivativesThis procedure is modeled to the one in found starting at page 239 of the lecture notes.
#
# We will make a function that can take the necessary inputs for each part and calculate the difference approximation for each h. It will also plot the absolute errors, which are not required for full credit.
#
# Here I also include $2^{-6}$ to $2^{-10}$
# +
import numpy as np
import matplotlib.pyplot as plt
import math
import cmath
# %matplotlib inline
def DiffApproximations(f,h,x,exact,loud=True,plot=True):
"""Determines the forward, backward, and centered difference
approximations for a given set of steps. Also prints the results
and plots the absolute errors if requested
Args:
f: function to approximate the derivative of
h: numpy array of step sizes
x: point at which to approximate
exact: the exact value at x for comparison purposes
loud: bool of whether to print a table of the results
plot: bool of whether to plot the errors
Returns:
forward: numpy array of the forward approximations
backward: numpy array of the backward approximations
centered: numpy array of the centered approximations"""
# Define variables to store approximations
forward = 0*h # forward difference
backward = 0*h # backward difference
center = 0*h # centered difference
compstep = 0*h # complex step
# Loop through each h
count = 0
for i in h:
forward[count] = (f(x+i) - f(x))/i
backward[count] = (f(x) - f(x-i))/i
center[count] = 0.5*(forward[count]+ backward[count])
compstep[count] = (f(x+i*1j)/i).imag
count += 1
# Print results
if(loud):
print('h\t forward\tbackward\tcentered\tcomplex step')
for i in range(count):
print("%.5f" % h[i]," %.11f" % forward[i],
" %.11f" % backward[i], " %.11f" % center[i], " %.11f" % compstep[i])
# Determine errors and plot
if(plot):
plt.loglog(h,np.fabs(forward-exact),'o-',label="Forward Difference")
plt.loglog(h,np.fabs(backward-exact),'o-',label="Backward Difference")
plt.loglog(h,np.fabs(center-exact),'o-',label="Central Difference")
plt.loglog(h,np.fabs(compstep-exact),'o-',label="Complex Step")
plt.legend(loc="best")
plt.title("Absolute Error on Log-Log Scale")
plt.xlabel("h")
plt.ylabel("Error")
plt.show()
return forward,backward,center
# Define step sizes
h = 2**np.linspace(-1,-10,10) #np.array([2**(-1),2**(-2),2**(-3),2**(-4),2**(-5)])
# -
# For the function
#
# $$f(x) = \sqrt{x},~\text{at}~x = 0.5.$$
# +
# Define knowns
f = lambda x: np.sqrt(x)
x = 0.5
exact = 0.70710678118
# Run function
forward,backward,center = DiffApproximations(f,h,x,exact)
# -
# For the function
#
# $$f(x) = \arctan(x^2 - 0.9 x + 2)~\text{at}~x=0.5$$
# +
# Define knowns
f = lambda x: np.arctan(x**2 - 0.9*x + 2)
x = 0.5
exact = 5/212
# Run function
forward,backward,center = DiffApproximations(f,h,x,exact)
# -
# For the function
#
# $$f(x) = J_0(x),~\text{at}~x = 1,~\text{where}~J_0(x)~\text{is a Bessel function of the first kind given by}$$
#
# $$J_\alpha(x) = \sum_{m=0}^\infty \frac{(-1)^m}{m! \, \Gamma(m+\alpha+1)} {\left(\frac{x}{2}\right)}^{2m+\alpha}.$$
# +
# Define knowns
def J_0(x, M = 100):
"""Order zero Bessel function of the first-kind
evaluated at x
Inputs:
alpha: value of alpha
x: point to evaluate Bessel function at
M: number of terms to include in sum
Returns:
J_0(x)
"""
total = 0.0
for m in range(M):
total += (-1)**m/(math.factorial(m)*math.gamma(m+1))*(0.5*x)**(2*m)
return total
x = 1
exact = -0.4400505857449335
# Run function
forward,backward,center = DiffApproximations(J_0,h,x,exact)
# -
# ## Comparison of Methods
# Consider the function
#
# $$f(x) = e^{-\frac{x^2}{\sigma^2}}.$$
#
# We will use finite differences to estimate derivatives of this function when $\sigma = 0.1$.
# - Using forward, backward, and centered differences evaluate the error in the function at 1000 points between $x=-1$ and $x=1$ ({\tt np.linspace} will be useful) using the following values of $h$:
# \[ h = 2^0, 2^{-1}, 2^{-2}, \dots, 2^{-7}.\]
# For each set of approximations compute the average absolute error over the one thousand points
# \[ \text{Average Absolute Error} = \frac{1}{N} \sum_{i=1}^{N} | f'(x_i) - f'_\mathrm{approx}(x_i)|,\]
# where $f'_\mathrm{approx}(x_i)$ is the value of an approximate derivative at $x_i$ and $N$ is the number of points the function derivative is evaluated at. You will need to find the exact value of the derivative to complete this estimate.
#
# Plot the value of the average absolute error error from each approximation on the same figure on a log-log scale. Discuss what you see. Is the highest-order method always the most accurate? Compute the order of accuracy you observe by computing the slope on the log-log plot.
#
# Next, compute the maximum absolute error for each value of $h$ as
# \[\text{Maximum Absolute Error} = \max_{i} | f'(x_i) - f'_\mathrm{approx}(x_i)|.\]
#
# Plot the value of the maximum absolute error error from each approximation on the same figure on a log-log scale. Discuss what you see. Is the highest-order method always the most accurate?
#
# - Repeat the previous part using the second-order version of the second-derivative approximation discussed above. You will only have one formula in this case.
# \item Now derive a formula for the fourth derivative and predict its order of accuracy. Then repeat the calculation and graphing of the average and maximum absolute errors and verify the order of accuracy.
#
# ## Solution
# We must know the exact first derivative, $f'(x)$, in order to determine the errors, therefore
#
# $$f'(x) = -\frac{2x}{\sigma^2}~e^{-\frac{x^2}{\sigma^2}} = -\frac{2x}{\sigma^2}~f(x).$$
#
# First, all of the constants, necessary functions and solution arrays are defined. The $\texttt{NumPy}$ function $\texttt{linspace}$ is used to define the evenly space values of $\texttt{x}$. Then, empty arrays are created that will fill all of the needed errors for each method (errors for each point, average errors for each step, and maximum errors for each step).
#
# $\br$A $\texttt{for}$ loop is used to loop through the index of each $h$, and then another loop is used to loop through the index of each $x$. Each approximation is then solved using the equations given in the Chapter 13 lecture notes. Next, the individual errors, average errors, and maximum errors are all calculated per the equations given in the problem statement. Last, the slopes for each method are determined using the approximations between $h = 2^{-6}$ and $h = 2^{-7}$, which approximate the order of error.
# +
import numpy as np
import matplotlib.pyplot as plt
# Define constants and functions
N = 1000
sigma = 0.1
f = lambda x: np.exp(-x**2/sigma**2)
fprime = lambda x: -2*x/sigma**2*f(x)
# Define step sizes
bases = 2*np.ones(8)
powers = np.array([0,-1,-2,-3,-4,-5,-6,-7])
h = np.power(bases,powers)
# Define values of x
x = np.linspace(-1,1,N)
# Evaluate derivative at each x
exact = fprime(x)
# Define arrays to fill with approximations
forward = np.zeros([h.size,x.size])
backward = np.zeros([h.size,x.size])
center = np.zeros([h.size,x.size])
comp1 = np.zeros([h.size,x.size])
comp2 = np.zeros([h.size,x.size])
# Define errors for each h
errorForward = np.zeros([h.size,x.size])
errorBackward = np.zeros([h.size,x.size])
errorCenter = np.zeros([h.size,x.size])
errorComp1 = np.zeros([h.size,x.size])
errorComp2 = np.zeros([h.size,x.size])
avgErrorForward = np.zeros(h.size)
avgErrorBackward = np.zeros(h.size)
avgErrorCenter = np.zeros(h.size)
avgErrorComp1 = np.zeros(h.size)
avgErrorComp2 = np.zeros(h.size)
maxErrorForward = np.zeros(h.size)
maxErrorBackward = np.zeros(h.size)
maxErrorCenter = np.zeros(h.size)
maxErrorComp1 = np.zeros(h.size)
maxErrorComp2 = np.zeros(h.size)
# Loop through indicies of h for h_i
for i in range(h.size):
# Loop through indicies x for x_j, solving for each x
for j in range(x.size):
forward[i,j] = (f(x[j]+h[i]) - f(x[j]))/h[i]
backward[i,j] = (f(x[j]) - f(x[j]-h[i]))/h[i]
center[i,j] = 0.5*(forward[i,j]+ backward[i,j])
comp1[i,j] = (f(x[j] +h[i]*1j)/h[i]).imag
comp2[i,j] = 8/3/h[i]*(f(x[j] +h[i]*1j/2)-1/8*f(x[j]+h[i]*1j)).imag
# Determine individual errors for h_i
errorForward[i,:] = np.fabs(exact-forward[i,:])
errorBackward[i,:] = np.fabs(exact-backward[i,:])
errorCenter[i,:] = np.fabs(exact-center[i,:])
errorComp1[i,:] = np.fabs(exact-comp1[i,:])
errorComp2[i,:] = np.fabs(exact-comp2[i,:])
# Determine average absolute error for h_i
avgErrorForward[i] = np.sum(errorForward[i,:])/N
avgErrorBackward[i] = np.sum(errorBackward[i,:])/N
avgErrorCenter[i] = np.sum(errorCenter[i,:])/N
avgErrorComp1[i] = np.sum(errorComp1[i,:])/N
avgErrorComp2[i] = np.sum(errorComp2[i,:])/N
# Determine max absolute error for h_i
maxErrorForward[i] = errorForward[i,:].max()
maxErrorBackward[i] = errorBackward[i,:].max()
maxErrorCenter[i] = errorCenter[i,:].max()
maxErrorComp1[i] = errorComp1[i,:].max()
maxErrorComp2[i] = errorComp2[i,:].max()
# Determine slope between last two approximations
slopeForward = (np.log(avgErrorForward[-1])-np.log(avgErrorForward[-2]))/(np.log(h[-1])-np.log(h[-2]))
slopeBackward = (np.log(avgErrorBackward[-1])-np.log(avgErrorBackward[-2]))/(np.log(h[-1])-np.log(h[-2]))
slopeCenter = (np.log(avgErrorCenter[-1])-np.log(avgErrorCenter[-2]))/(np.log(h[-1])-np.log(h[-2]))
slopeComp1 = (np.log(avgErrorComp1[-1])-np.log(avgErrorComp1[-2]))/(np.log(h[-1])-np.log(h[-2]))
slopeComp2 = (np.log(avgErrorComp2[-1])-np.log(avgErrorComp2[-2]))/(np.log(h[-1])-np.log(h[-2]))
# -
# The average error for each method is then plotted for each method, on a log-log scale.
# Plot average error
plt.loglog(h,avgErrorForward,'o-',label="Forward difference")
plt.loglog(h,avgErrorBackward,'o-',label="Backward difference")
plt.loglog(h,avgErrorCenter,'o-',label="Central difference")
plt.loglog(h,avgErrorComp1,'o-',label="Comp. Step 1")
plt.loglog(h,avgErrorComp2,'o-',label="Comp. Step 2")
plt.legend(loc="best")
plt.title('Average absolute error, log-log scale')
plt.xlabel('h')
plt.ylabel('Error')
plt.show()
# We see that the methods are rather similar in the terms of error up until $h = 2^{-2}$, and then the error of the central difference method diverges from the others. Throughout the domain of $h$, the forward and backward methods have errors of the same magnitude. The central difference method has the least error throughout the entire domain (note that this may not be the case for other functions). Of interest is that the error increases for all three methods up until $h = 2^{-2}$. This is due to the fact that this is the region in which $h^2 \approx h$, where the error then begins to decrease.
#
# $\br$The estimates for the order of accuracy are then printed.
# Print slopes for order accuracy
print('Order accuracies')
print('Forward difference\t',"%.5f" % slopeForward)
print('Backward difference\t',"%.5f" % slopeBackward)
print('Center difference\t',"%.5f" % slopeCenter)
print('Comp Step 1\t',"%.5f" % slopeComp1)
print('Comp Step 2\t',"%.5f" % slopeComp2)
# As expected, the forward and backward difference methods have the same order error. The divergence of the central difference method is also evident by the fact that it is of second-order error.
# Plot maximum error
plt.loglog(h,maxErrorForward,'o-',label="Forward difference")
plt.loglog(h,maxErrorBackward,'o-',label="Backward difference")
plt.loglog(h,maxErrorCenter,'o-',label="Central difference")
plt.loglog(h,maxErrorComp1,'o-',label="Comp Step 1")
plt.loglog(h,maxErrorComp2,'o-',label="Comp Step 1")
plt.legend(loc="best")
plt.title('Maximum absolute error, log-log scale')
plt.xlabel('h')
plt.ylabel('Error')
plt.show()
# For this example, the plot shows that the second-order method remains the most accurate in terms of maximum errors for all $h$. The increase in error for the first three step sizes of the forward and backward difference methods is more evident with the maximum error. Again, the orders of accuracy become more clear as $h \rightarrow 0$. Of interest is that the maximum errors are generally an order of magnitude higher than the average errors, meaning that for some values of $x$ the approximation is significantly less accurate.
#
# $\br$Next, we will estimate the second-derivative.
#
# $\br$It is necessary to determine the exact second derivative, $f''(x)$, in order to determine the errors, therefore
#
# $$f''(x) = \frac{4x^2 - 2\sigma^2}{\sigma^4}~e^{-\frac{x^2}{\sigma^2}} = \frac{4x^2 - 2\sigma^2}{\sigma^4}~f(x).$$
#
# $\br$The same constants are defined as were previously with the first-order approximation. In addition, the same set of $\texttt{for}$ loops is used to solve for the approximations for each $x$ and $h$. The errors are then calculated, the order of accuracy approximated, and plots are made for the average absolute error and the maximum absolute error.
# +
# Define array to fill with approximations
second = np.zeros([h.size,x.size])
# Define errors for each h
errorSecond = np.zeros([h.size,x.size])
avgErrorSecond = np.zeros(h.size)
maxErrorSecond = np.zeros(h.size)
# Define exact solution and evaluate at x
fprime2 = lambda x: (4*x**2-2*sigma**2)/sigma**4*f(x)
exact2 = fprime2(x)
# Loop through indicies of h for h_i
for i in range(h.size):
# Loop through indicies x for x_j, solving for each x
for j in range(x.size):
second[i,j] = (f(x[j]+h[i])-2*f(x[j])+f(x[j]-h[i]))/h[i]**2
# Determine individual errors for h_i
errorSecond[i,:] = np.fabs(exact2-second[i,:])
# Determine average absolute error for h_i
avgErrorSecond[i] = np.sum(errorSecond[i,:])/N
# Determine max absolute error for h_i
maxErrorSecond[i] = errorSecond[i,:].max()
# Determine slope between last two approximations
slopeSecond = (np.log(avgErrorSecond[-1])-np.log(avgErrorSecond[-2]))/(np.log(h[-1])-np.log(h[-2]))
# Plot average error
plt.loglog(h,avgErrorSecond,'o-')
plt.title('Average absolute error, log-log scale')
plt.xlabel('h')
plt.ylabel('Error')
plt.show()
# Print slope for order accuracy
print('Order accuracy')
print('Second-derivative approximation\t',"%.5f" % slopeSecond)
# Plot maximum error
plt.loglog(h,maxErrorSecond,'o-')
plt.title('Maximum absolute error, log-log scale')
plt.xlabel('h')
plt.ylabel('Error')
plt.show()
# -
# As seen, we have second-order accuracy that is evident in both of the plots above. In addition, it is important to take note of the magnitude of the maximum errors compared to the magnitude of the average errors. In this case again, the maximum errors are significantly larger.
# $\br$Next... we will venture into creating our own formula to approximate the fourth derivative. As usual, we must first know the exact solution of the fourth derivative, which is
#
# $$f^{(4)}(x) = \frac{4\Big(3\sigma^4 + 4x^4 - 12\sigma^2x^2\Big)}{\sigma^8}e^{-\frac{x^2}{\sigma^2}} = \frac{4\Big(3\sigma^4 + 4x^4 - 12\sigma^2x^2\Big)}{\sigma^8}~f(x).$$
#
# Here, we will use a central difference method to determine the fourth derivative. There are many finite difference approximations that can be made for the fourth-derivative: forward, backward, centered, etc. As long as the process made results in a viable method, credit will be awarded.
#
# $\br$First, we must start with the Taylor series expansion at $x+h$, $x-h$, $x+2h$, and $x-2h$:
#
# $$f(x+h) = f(x) + hf^{(1)}(x) + \frac{h^2}{2}f^{(2)}(x) + \frac{h^3}{6}f^{(3)}(x) + \frac{h^4}{24}f^{(4)}(x) + \frac{h^5}{120}f^{(5)}(x) + \frac{h^6}{720}f^{(6)}(x) + O(h^7),$$
#
# $$f(x-h) = f(x) - hf^{(1)}(x) + \frac{h^2}{2}f^{(2)}(x) - \frac{h^3}{6}f^{(3)}(x) + \frac{h^4}{24}f^{(4)}(x) - \frac{h^5}{120}f^{(5)}(x) + \frac{h^6}{720}f^{(6)}(x) + O(h^7),$$
#
# $$f(x+2h) = f(x) + 2hf^{(1)}(x) + 2h^2f^{(2)}(x) + \frac{4h^3}{3}f^{(3)}(x) + \frac{2h^4}{3}f^{(4)}(x) + \frac{4h^5}{15}f^{(5)}(x) + \frac{4h^6}{45}f^{(6)}(x) + O(h^7),$$
#
# and
#
# $$f(x-2h) = f(x) - 2hf^{(1)}(x) + 2h^2f^{(2)}(x) - \frac{4h^3}{3}f^{(3)}(x) + \frac{2h^4}{3}f^{(4)}(x) - \frac{4h^5}{15}f^{(5)}(x) + \frac{4h^6}{45}f^{(6)}(x) + O(h^7).$$
#
# Next, we will add the above four equations in a way such that the $h^2$ term is cancelled out. This will be done by adding $-2$ times the first two equations, and $1$ times the last two equations:
#
# $$-4f(x+h) - 4 f(x-h) + f(x+2h) + f(x-2h) = -6f(x) + h^4f^{(4)}(x) + \frac{h^6}{6}f^{(6)}(x).$$
#
# The equation is then solved for the fourth derivative:
#
# $$f^{(4)}(x) = \frac{f(x-2h) - 4f(x-h) + 6f(x) - 4f(x+h) + f(x+2h)}{h^4} - \frac{h^2}{6}f^{(6)}(x)$$
#
# Taking care of the last term, we can consider that the remaining error is on the order of $h^2$.
#
# $$f^{(4)}(x) = \frac{f(x-2h) - 4f(x-h) + 6f(x) - 4f(x+h) + f(x+2h)}{h^4} + O(h^2)$$
#
# Now, we have our centered finite difference approximation for the fourth derivative. Following the same process as done in the two parts above, we will evaluate its performance at varying values of $h$.
# +
# Define array to fill with approximations
fourth = np.zeros([h.size,x.size])
# Define errors for each h
errorFourth = np.zeros([h.size,x.size])
avgErrorFourth = np.zeros(h.size)
maxErrorFourth = np.zeros(h.size)
# Define exact solution and evaluate at x
fprime4 = lambda x: 4*f(x)*(3*sigma**4+4*x**4-12*sigma**2*x**2)/sigma**8
exact4 = fprime4(x)
# Loop through indicies of h for h_i
for i in range(h.size):
# Loop through indicies x for x_j, solving for each x
for j in range(x.size):
fourth[i,j] = (f(x[j]-2*h[i])-4*f(x[j]-h[i])+6*f(x[j])-4*f(x[j]+h[i])+f(x[j]+2*h[i]))/h[i]**4
# Determine individual errors for h_i
errorFourth[i,:] = np.fabs(exact4-fourth[i,:])
# Determine average absolute error for h_i
avgErrorFourth[i] = np.sum(errorFourth[i,:])/N
# Determine max absolute error for h_i
maxErrorFourth[i] = errorSecond[i,:].max()
# Determine slope between last two approximations
slopeFourth = (np.log(avgErrorFourth[-1])-np.log(avgErrorFourth[-2]))/(np.log(h[-1])-np.log(h[-2]))
# Plot average error
plt.loglog(h,avgErrorFourth,'o-')
plt.title('Average absolute error, log-log scale')
plt.xlabel('h')
plt.ylabel('Error')
plt.show()
# Print slope for order accuracy
print('Order accuracy')
print('Fourth-derivative approximation\t',"%.5f" % slopeFourth)
# Plot maximum error
plt.loglog(h,maxErrorFourth,'o-')
plt.title('Maximum absolute error, log-log scale')
plt.xlabel('h')
plt.ylabel('Error')
plt.show()
# -
# The calculation of the slope at the last two points leads to an order of accuracy of 2, as we expected in the formulation of our method above.
|
solution_chapter14.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="waM_Vx6Ej4DW" outputId="a4dbfe71-1c3f-469e-ff97-0d7837406cd7"
# Installation of PM4Py Python package
#Source: https://pm4py.fit.fraunhofer.de/install-page
# !pip install pm4py
# + id="7LnoPs-HexFk" colab={"base_uri": "https://localhost:8080/", "height": 647} outputId="728907f5-e919-41d8-cc9a-1081ac57a176"
from google.colab import drive
drive.mount('/content/drive')
# + id="TuNX1E8EkK0a"
# Import dependencies and set the filepath
import pm4py
import os
import pandas as pd
import numpy as np
import datetime as dt
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.express as px # needed for plotly express in general
import plotly.graph_objects as go # needed for plotly graph objects e.g. pie
# + id="ResKZ_cuqAoo"
# Set the file path
# Please upload the local file sent via email to colab before running this
file_path = '/content/Hospital Billing - Event Log.xes' # if using the original file
#file_path = '/content/log.xes' # if renamed
# + colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["820a25abca8c4d7eb7336314fd3d9a62", "82e57feef0474ad5a57c1e9b13278d35", "2126520666b14ad4946508a43a1047f1", "a99e491989f54e8e839d142fbfa24061", "0c1951cb6c0e4007a891d46b2d60091a", "4c2841b5df4c45fda31f9e27b0f5944f", "051a2c534c3145cdaa993ab60cffa9a3", "13dffe7bb3814b7aaa725f2342dd5ed0"]} id="SpV08HzYpiek" outputId="54185cd9-c2f4-46ac-bd22-5201be2f205f"
# Read the XES file and create the log
event_log = pm4py.read_xes(file_path)
# + id="7F0g5TIjmyIH" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="5bb90f7b-8986-4fd2-e0e2-0092ec22d5ac"
def import_xes(file_path):
event_log = pm4py.read_xes(file_path)
start_activities = pm4py.get_start_activities(event_log)
end_activities = pm4py.get_end_activities(event_log)
print("Start activities: {}\nEnd activities: {}".format(start_activities, end_activities))
# + id="IkcQ2bsyd9O7" colab={"base_uri": "https://localhost:8080/", "height": 232} outputId="ba267016-2bfe-4cc2-96f8-5e7f2a36037f"
# Obtaining a Process Model
# event_log = pm4py.read_xes('C:/Users/demo/Downloads/running-example.xes')
process_tree = pm4py.discover_process_tree_inductive(event_log) # discover_tree_inductive is deprecated
bpmn_model = pm4py.convert_to_bpmn(process_tree)
pm4py.view_bpmn(bpmn_model)
# + id="msikhvUYvPLU"
#obtain process map
dfg, start_activities, end_activities = pm4py.discover_dfg(event_log)
pm4py.view_dfg(dfg, start_activities, end_activities)
# + id="RfB93RYJvt3P"
#heuristics map
map = pm4py.discover_heuristics_net(event_log)
pm4py.view_heuristics_net(map)
# + id="v3f6FIPeuCuq"
# View Process Tree
process_tree = pm4py.discover_process_tree_inductive(event_log) # discover_tree_inductive is deprecated
pm4py.view_process_tree(process_tree)
# + colab={"base_uri": "https://localhost:8080/", "height": 164} id="CsnLkwlX76F5" outputId="f4c68efc-9c7e-405d-84aa-9a1beab5d79c"
print("Start activities: {}\nEnd activities: {}".format(start_activities, end_activities))
# + [markdown] id="wbtQto0x-Oew"
# ## Miner Algorithms
# Please refer to the documentation here:
# https://pm4py.fit.fraunhofer.de/documentation#item-3-1
# + id="dJYcOYCF-JC8"
# Alpha Miner implementation
from pm4py.algo.discovery.alpha import algorithm as alpha_miner
net, initial_marking, final_marking = alpha_miner.apply(event_log)
# Inductive Miner implementation
from pm4py.algo.discovery.inductive import algorithm as inductive_miner
net, initial_marking, final_marking = inductive_miner.apply(event_log)
# + [markdown] id="aNlOpRZs-Ifd"
# ## Visualisations
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="3Eh9q3t59PCR" outputId="a897a039-ca86-4c62-897b-9c8eb882f71e"
# Visualisation of IM
from pm4py.visualization.process_tree import visualizer as pt_visualizer
tree = inductive_miner.apply_tree(event_log)
gviz = pt_visualizer.apply(tree)
pt_visualizer.view(gviz)
# + id="zcjREevPuo7A"
# + colab={"base_uri": "https://localhost:8080/", "height": 37} id="p6H3fChVAmZz" outputId="a9b01993-7fa0-4bba-8989-bbe22893a299"
# Converting process tree in petri net
from pm4py.objects.conversion.process_tree import converter as pt_converter
net, initial_marking, final_marking = pt_converter.apply(tree, variant=pt_converter.Variants.TO_PETRI_NET)
gviz = pt_visualizer.apply(tree)
pt_visualizer.view(gviz)
# + id="h61RH4itDKWk"
# Dotted chart
pm4py.view_dotted_chart(event_log, format="svg")
# + [markdown] id="9s3yfBSorKat"
# Decision tree & Root Cause Analysis
#
# + colab={"base_uri": "https://localhost:8080/"} id="8oDtaz-TU8_E" outputId="3b9facc7-6d64-43a8-fdb5-9c024477821c"
from pm4py.objects.log.util import get_class_representation , get_log_representation
from sklearn import tree
data, feature_names = get_log_representation.get_default_representation(event_log)
#changing into pandas data frame
dataframe = pd.DataFrame(data, columns=feature_names)
#creating decision tree
target, classes = get_class_representation.get_class_representation_by_trace_duration(event_log, 11007360)
clf = tree.DecisionTreeClassifier()
clf.fit(data, target)
# + colab={"base_uri": "https://localhost:8080/", "height": 37} id="DpoF8J4XVKGl" outputId="5aff9e3c-b160-43a7-c50a-982aad4f7a61"
#visualizing decision tree
from pm4py.visualization.decisiontree import visualizer as dectree_visualizer
from pm4py.objects.log.util import get_class_representation
gviz = dectree_visualizer.apply(clf, feature_names, classes)
target, classes = get_class_representation.get_class_representation_by_trace_duration(event_log, 11007360)
dectree_visualizer.view(gviz)
|
notebooks/root-cause-analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="bwCxd7ugXMCu"
# # MNIST Dataseti
#
# 0'dan 10'a kadar yazılmış el yazısı rakamların fotoğraflarıdır. 60000 resim içerir. AI için aslında bir hello world'dür.[Datasetin rekoru](https://www.kaggle.com/cdeotte/mnist-perfect-100-using-knn) %100'dür.
# + id="ia0bmMWEydtD"
import matplotlib.pyplot as plt
from tensorflow import keras
import numpy as np
# %matplotlib inline
# + id="qOWccXdTypjb"
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
# + colab={"base_uri": "https://localhost:8080/"} id="S7ChF_9vytyF" outputId="9c3e2e8e-9e06-4c01-dca4-411bbed8f189"
x_train.shape, x_test.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="R5vEekaeyvxD" outputId="d99ae2a0-54f3-42d8-ff32-4296201f2be8"
plt.imshow(x_train[0])
y_train[0]
# + id="opmvJxMczI6X"
model = keras.Sequential([
keras.layers.Flatten(),
keras.layers.Dense(32),
keras.layers.Dense(10, activation='softmax')
])
# + id="aksCko82zU4B"
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="CQPnvhVjzTUc" outputId="99f12a02-e3f2-40bf-f329-62b3029c92e8"
model.fit(x_train, y_train, epochs=3)
# + id="1EX1u1es1QqW" colab={"base_uri": "https://localhost:8080/"} outputId="f46abf28-fe74-47a4-885c-26457838457e"
model.evaluate(x_test, y_test)
|
Models/MNIST/MNIST - Easy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# imports
import nibabel as nib
import numpy as np
import os
import random
import time
import warnings
warnings.filterwarnings('ignore')
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Function
from torch.autograd.functional import jacobian as J
from utils import *
# +
# settings
# data
cases = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44]
fold = 0
if fold == 0:
test_cases = [0, 5, 10, 15, 20, 25, 30, 35, 40]
elif fold == 1:
test_cases = [1, 6, 11, 16, 21, 26, 31, 36, 41]
elif fold == 2:
test_cases = [2, 7, 12, 17, 22, 27, 32, 37, 42]
elif fold == 3:
test_cases = [3, 8, 13, 18, 23, 28, 33, 38, 43]
elif fold == 4:
test_cases = [4, 9, 14, 19, 24, 29, 32, 39, 44]
train_cases = [i for i in cases if not i in test_cases]
# misc
device = 'cuda'
torch.backends.cudnn.enabled = True
torch.backends.cudnn.benchmark = True
model_dir = 'fold{}'.format(fold)
if not os.path.exists(os.path.join('./models/', model_dir)):
os.makedirs(os.path.join('./models/', model_dir))
data_dir = './data/preprocessed/resize_s'
# keypoints / graph
d = 5 # 3 (for refinement stage)
N_P = 2048 # 3072 (for refinement stage)
k = 15
# displacements
l_max = 14 # 8 (for refinement stage)
l_width = l_max * 2 + 1
q = 2 # 1 (for refinement stage)
# model
base = 4
sigma2 = 1
# training
num_epochs = 150
init_lr = 0.1
save_iter = 10
# +
# load data
imgs_fixed = {}
masks_fixed = {}
imgs_moving = {}
masks_moving = {}
for case in train_cases:
print('loading case {} ...'.format(case + 1), end=' ')
t0 = time.time()
input_img_fixed = os.path.join(data_dir, 'case{}_img_fixed.nii.gz'.format(case + 1))
input_mask_fixed = os.path.join(data_dir, 'case{}_mask_fixed.nii.gz'.format(case + 1))
input_img_moving = os.path.join(data_dir, 'case{}_img_moving.nii.gz'.format(case + 1))
input_mask_moving = os.path.join(data_dir, 'case{}_mask_moving.nii.gz'.format(case + 1))
img_fixed = (torch.from_numpy(nib.load(input_img_fixed).get_data()).unsqueeze(0).unsqueeze(0).float().clamp_(-1000, 1500) + 1000) / 2500
mask_fixed = torch.from_numpy(nib.load(input_mask_fixed).get_data()).unsqueeze(0).unsqueeze(0).bool()
img_moving = (torch.from_numpy(nib.load(input_img_moving).get_data()).unsqueeze(0).unsqueeze(0).float().clamp_(-1000, 1500) + 1000) / 2500
mask_moving = torch.from_numpy(nib.load(input_mask_moving).get_data()).unsqueeze(0).unsqueeze(0).bool()
imgs_fixed[case] = img_fixed
masks_fixed[case] = mask_fixed
imgs_moving[case] = img_moving
masks_moving[case] = mask_moving
t1 = time.time()
print('{:.2f} s'.format(t1-t0))
_, _, D, H, W = imgs_fixed[train_cases[0]].shape
# +
# displacement space
disp = torch.stack(torch.meshgrid(torch.arange(- q * l_max, q * l_max + 1, q * 2),
torch.arange(- q * l_max, q * l_max + 1, q * 2),
torch.arange(- q * l_max, q * l_max + 1, q * 2))).permute(1, 2, 3, 0).contiguous().view(1, -1, 3).float()
disp = (disp.flip(-1) * 2 / (torch.tensor([W, H, D]) - 1)).to(device)
# +
# graphregnet
class GaussianSmoothing(nn.Module):
def __init__(self, sigma):
super(GaussianSmoothing, self).__init__()
sigma = torch.tensor([sigma]).to(device)
N = torch.ceil(sigma * 3.0 / 2.0).long().item() * 2 + 1
weight = torch.exp(-torch.pow(torch.linspace(-(N // 2), N // 2, N).to(device), 2) / (2 * torch.pow(sigma, 2)))
weight /= weight.sum()
self.weight = weight
def forward(self, x):
x = filter1D(x, self.weight, 0)
x = filter1D(x, self.weight, 1)
x = filter1D(x, self.weight, 2)
return x
class Encoder(nn.Module):
def __init__(self, in_channels=1, base=4):
super(Encoder, self).__init__()
self.conv_in = nn.Sequential(nn.Conv3d(in_channels, base, 3, stride=2, padding=1, bias=False),
nn.InstanceNorm3d(base),
nn.LeakyReLU())
self.conv1 = nn.Sequential(nn.Conv3d(base, 2*base, 3, stride=2, padding=1, bias=False),
nn.InstanceNorm3d(2*base),
nn.LeakyReLU())
self.conv2 = nn.Sequential(nn.Conv3d(2*base, 4*base, 3, stride=2, padding=1, bias=False),
nn.InstanceNorm3d(4*base),
nn.LeakyReLU())
def forward(self, x):
x1 = self.conv_in(x)
x2 = self.conv1(x1)
x3 = self.conv2(x2)
return x1, x2, x3
class Decoder(nn.Module):
def __init__(self, out_channels=1, base=4):
super(Decoder, self).__init__()
self.conv1 = nn.Sequential(nn.Conv3d(4*base, 2*base, 3, stride=1, padding=1, bias=False),
nn.InstanceNorm3d(2*base),
nn.LeakyReLU())
self.conv1a = nn.Sequential(nn.Conv3d(4*base, 2*base, 3, stride=1, padding=1, bias=False),
nn.InstanceNorm3d(2*base),
nn.LeakyReLU())
self.conv2 = nn.Sequential(nn.Conv3d(2*base, base, 3, stride=1, padding=1, bias=False),
nn.InstanceNorm3d(base),
nn.LeakyReLU())
self.conv2a = nn.Sequential(nn.Conv3d(2*base, base, 3, stride=1, padding=1, bias=False),
nn.InstanceNorm3d(base),
nn.LeakyReLU())
self.conv_out = nn.Sequential(nn.Conv3d(base, 1, 3, padding=1))
def forward(self, x1, x2, x3):
x = F.interpolate(x3, size=x2.shape[-3:], mode='trilinear')
x = self.conv1(x)
x = self.conv1a(torch.cat([x, x2], dim=1))
x = F.interpolate(x, size=x1.shape[-3:], mode='trilinear')
x = self.conv2(x)
x = self.conv2a(torch.cat([x, x1], dim=1))
x = self.conv_out(x)
return x
class EdgeConv(nn.Module):
def __init__(self, in_channels, out_channels):
super(EdgeConv, self).__init__()
self.in_channels = in_channels
self.out_channels = out_channels
self.conv = nn.Sequential(
nn.Conv3d(self.in_channels * 2, self.out_channels, 1, bias=False),
nn.InstanceNorm3d(self.out_channels),
nn.LeakyReLU()
)
def forward(self, x, ind):
B, N, C, D, _, _ = x.shape
k = ind.shape[2]
y = x.view(B*N, C, D*D*D)[ind.view(B*N, k)].view(B, N, k, C, D*D*D)
x = x.view(B, N, C, D*D*D).unsqueeze(2).expand(-1, -1, k, -1, -1)
x = torch.cat([y - x, x], dim=3).permute(0, 3, 1, 2, 4)
x = self.conv(x)
x = x.mean(dim=3).permute(0, 2, 1, 3).view(B, N, -1, D, D, D)
return x
class GCN(nn.Module):
def __init__(self, base=4):
super(GCN, self).__init__()
self.base = base
self.conv1 = EdgeConv(4*self.base + 3, 4*self.base)
self.conv2 = EdgeConv(2*4*self.base + 3, 4*self.base)
self.conv3 = EdgeConv(3*4*self.base + 3, 4*self.base)
def forward(self, x1, x2, x3, kpts, ind):
expand = x3.shape[-1]
xa = self.conv1(torch.cat([x3, kpts.view(-1, 3, 1, 1, 1).expand(-1, -1, expand, expand, expand)], dim=1).unsqueeze(0), ind).squeeze(0)
xb = self.conv2(torch.cat([torch.cat([x3, kpts.view(-1, 3, 1, 1, 1).expand(-1, -1, expand, expand, expand)], dim=1), xa], dim=1).unsqueeze(0), ind).squeeze(0)
xc = self.conv3(torch.cat([torch.cat([x3, kpts.view(-1, 3, 1, 1, 1).expand(-1, -1, expand, expand, expand)], dim=1), xa, xb], dim=1).unsqueeze(0), ind).squeeze(0)
return x1, x2, xc
class GraphRegNet(nn.Module):
def __init__(self, base, smooth_sigma):
super(GraphRegNet, self).__init__()
self.base = base
self.smooth_sigma = smooth_sigma
self.pre_filter1 = GaussianSmoothing(self.smooth_sigma)
self.pre_filter2 = GaussianSmoothing(self.smooth_sigma)
self.encoder1 = Encoder(2, self.base)
self.gcn1 = GCN(self.base)
self.decoder1 = Decoder(1, self.base)
self.encoder2 = Encoder(4, self.base)
self.gcn2 = GCN(self.base)
self.decoder2 = Decoder(1, self.base)
def forward(self, x, kpts, kpts_knn):
x1 = self.encoder1(torch.cat([x, self.pre_filter1(x)], dim=1))
x1 = self.gcn1(*x1, kpts, kpts_knn)
x1 = self.decoder1(*x1)
x1 = F.interpolate(x1, size=x.shape[-3:], mode='trilinear')
x2 = self.encoder2(torch.cat([x, self.pre_filter1(x), x1, self.pre_filter2(x1)], dim=1))
x2 = self.gcn2(*x2, kpts, kpts_knn)
x2 = self.decoder2(*x2)
return x2
def init_weights(m):
if isinstance(m, nn.Conv3d):
nn.init.xavier_normal(m.weight)
if m.bias is not None:
nn.init.constant(m.bias, 0.0)
# +
# differentiable sparse-to-dense supervision
class InverseGridSample(Function):
@staticmethod
def forward(ctx, input, grid, shape, mode='bilinear', padding_mode='zeros', align_corners=None):
B, C, N = input.shape
D = grid.shape[-1]
device = input.device
dtype = input.dtype
ctx.save_for_backward(input, grid)
if D == 2:
input_view = [B, C, -1, 1]
grid_view = [B, -1, 1, 2]
elif D == 3:
input_view = [B, C, -1, 1, 1]
grid_view = [B, -1, 1, 1, 3]
ctx.grid_view = grid_view
ctx.mode = mode
ctx.padding_mode = padding_mode
ctx.align_corners = align_corners
with torch.enable_grad():
output = J(lambda x: InverseGridSample.sample(input.view(*input_view), grid.view(*grid_view), x, mode, padding_mode, align_corners), (torch.zeros(B, C, *shape).to(dtype).to(device)))
return output
@staticmethod
def backward(ctx, grad_output):
input, grid = ctx.saved_tensors
grid_view = ctx.grid_view
mode = ctx.mode
padding_mode = ctx.padding_mode
align_corners = ctx.align_corners
grad_input = F.grid_sample(grad_output, grid.view(*grid_view), mode, padding_mode, align_corners)
return grad_input.view(*input.shape), None, None, None, None, None
@staticmethod
def sample(input, grid, accu, mode='bilinear', padding_mode='zeros', align_corners=None):
sampled = F.grid_sample(accu, grid, mode, padding_mode, align_corners)
return -0.5 * ((input - sampled) ** 2).sum()
def inverse_grid_sample(input, grid, shape, mode='bilinear', padding_mode='zeros', align_corners=None):
return InverseGridSample.apply(input, grid, shape, mode, padding_mode, align_corners)
def densify(kpts, kpts_disp, shape, smooth_iter=3, kernel_size=5, eps=0.0001):
B, N, _ = kpts.shape
device = kpts.device
D, H, W = shape
grid = inverse_grid_sample(kpts_disp.permute(0, 2, 1), kpts, shape, padding_mode='border', align_corners=True)
grid_norm = inverse_grid_sample(torch.ones(B, 1, N).to(device), kpts, shape, padding_mode='border', align_corners=True)
avg_pool = nn.AvgPool3d(kernel_size, stride=1, padding=kernel_size // 2).to(device)
for i in range(smooth_iter):
grid = avg_pool(grid)
grid_norm = avg_pool(grid_norm)
grid = grid / (grid_norm + eps)
return grid
# +
# training
# model
graphregnet = GraphRegNet(base, sigma2).to(device)
graphregnet.apply(init_weights)
parameter_count(graphregnet)
# optimizer
optimizer = optim.Adam(graphregnet.parameters(), init_lr)
# criterion
def criterion(feat_fixed, feat_moving, disp, mask):
mse_loss = nn.MSELoss(reduction='none')
loss = (mse_loss(feat_fixed, warp_img(feat_moving, disp.permute(0, 2, 3, 4, 1))) * mask).sum() / mask.float().sum()
return loss
# statistics
losses = []
torch.cuda.synchronize()
t0 = time.time()
# for num_epochs epochs
for epoch in range(num_epochs):
# train mode
graphregnet.train()
# statistics
running_loss = 0.0
# shuffle training cases
train_cases_perm = random.sample(train_cases, len(train_cases))
# for all training cases
for case in train_cases_perm:
# zero out gradients
optimizer.zero_grad()
# load data
img_fixed = imgs_fixed[case].to(device)
mask_fixed = masks_fixed[case].to(device)
img_moving = imgs_moving[case].to(device)
mask_moving = masks_moving[case].to(device)
# extract kpts and generate knn graph
kpts_fixed = foerstner_kpts(img_fixed, mask_fixed, d=d, num_points=N_P)
kpts_fixed_knn = knn_graph(kpts_fixed, k, include_self=True)[0]
# extract mind features
mind_fixed = mindssc(img_fixed)
mind_moving = mindssc(img_moving)
# displacement cost computation
cost = ssd(kpts_fixed, mind_fixed, mind_moving, (D, H, W), l_max, q).view(-1, 1, l_width, l_width, l_width)
# forward
kpts_fixed_disp_pred = graphregnet(cost, kpts_fixed, kpts_fixed_knn)
# sparse to dense
disp_pred = densify(kpts_fixed, (disp.unsqueeze(1) * F.softmax(kpts_fixed_disp_pred.view(1, N_P, -1), 2).unsqueeze(3)).sum(2), (D//3, H//3, W//3))
disp_pred = F.interpolate(disp_pred, size=(D, H, W), mode='trilinear')
# loss
loss = criterion(mind_fixed, mind_moving, disp_pred, mask_moving)
# backward + optimize
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item()
running_loss /= (len(train_cases))
losses.append(running_loss)
if ((epoch + 1) % save_iter) == 0:
torch.cuda.synchronize()
t1 = time.time()
print('epoch: ', epoch + 1)
print('loss: {:.4f}'.format(running_loss))
print('time (epoch): {:.4f} s'.format((t1 - t0) / save_iter))
gpu_usage()
print('---')
torch.save(graphregnet.cpu().state_dict(), os.path.join('./models', model_dir, 'epoch{}.pth'.format(epoch)))
graphregnet.to(device)
torch.cuda.synchronize()
t0 = time.time()
torch.save(graph_reg_net.cpu().state_dict(), os.path.join('./models', model_dir, 'final.pth'))
# -
|
train.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # SELECT from Nobel
#
# ## `nobel` Nobel Laureates
#
# We continue practicing simple SQL queries on a single table.
#
# This tutorial is concerned with a table of Nobel prize winners:
#
# ```
# nobel(yr, subject, winner)
# ```
#
# Using the `SELECT` statement.
library(tidyverse)
library(DBI)
library(getPass)
drv <- switch(Sys.info()['sysname'],
Windows="PostgreSQL Unicode(x64)",
Darwin="/usr/local/lib/psqlodbcw.so",
Linux="PostgreSQL")
con <- dbConnect(
odbc::odbc(),
driver = drv,
Server = "localhost",
Database = "sqlzoo",
UID = "postgres",
PWD = <PASSWORD>("Password?"),
Port = 5432
)
options(repr.matrix.max.rows=20)
# ## 1. Winners from 1950
#
# Change the query shown so that it displays Nobel prizes for 1950.
nobel <- dbReadTable(con, 'nobel')
nobel %>%
filter(yr==1950)
# ## 2. 1962 Literature
#
# Show who won the 1962 prize for Literature.
nobel %>%
filter(yr==1962 & subject=='Literature') %>%
select(winner)
# ## 3. <NAME>
#
# Show the year and subject that won '<NAME>' his prize.
nobel %>%
filter(winner=='<NAME>') %>%
select(yr, subject)
# ## 4. Recent Peace Prizes
#
# Give the name of the 'Peace' winners since the year 2000, including 2000.
nobel %>%
filter(yr>=2000 & subject=='Peace') %>%
select(winner)
# ## 5. Literature in the 1980's
#
# Show all details **(yr, subject, winner)** of the Literature prize winners for 1980 to 1989 inclusive.
nobel %>%
filter(between(yr, 1980, 1989) &
subject=='Literature')
# ## 6. Only Presidents
#
# Show all details of the presidential winners:
#
# - <NAME>
# - <NAME>
# - <NAME>
# - <NAME>
nobel %>%
filter(winner %in% c(
'<NAME>', '<NAME>', '<NAME>', '<NAME>'))
# ## 7. John
#
# Show the winners with first name John
nobel %>%
filter(str_starts(winner, 'John')) %>%
select(winner)
# ## 8. Chemistry and Physics from different years
#
# **Show the year, subject, and name of Physics winners for 1980 together with the Chemistry winners for 1984.**
nobel %>%
filter(yr==1980 & subject=='Physics' |
yr==1984 & subject=='Chemistry') %>%
select(yr, subject, winner)
# ## 9. Exclude Chemists and Medics
#
# **Show the year, subject, and name of winners for 1980 excluding Chemistry and Medicine**
nobel %>%
filter(yr==1980 &
! subject %in% c('Chemistry', "Medicine")) %>%
select(yr, subject, winner)
# ## 10. Early Medicine, Late Literature
#
# Show year, subject, and name of people who won a 'Medicine' prize in an early year (before 1910, not including 1910) together with winners of a 'Literature' prize in a later year (after 2004, including 2004)
nobel %>%
filter((subject=='Medicine' & yr<1910) |
(subject=='Literature' & yr>=2004)) %>%
select(yr, subject, winner)
# ## 11. Umlaut
#
# Find all details of the prize won by <NAME>
#
# > _Non-ASCII characters_
# > The u in his name has an umlaut. You may find this link useful <https://en.wikipedia.org/wiki/%C3%9C#Keyboarding>
nobel %>%
filter(tolower(winner)=='<NAME>')
# ## 12. Apostrophe
#
# Find all details of the prize won by <NAME>
#
# > _Escaping single quotes_
# > You can't put a single quote in a quote string directly. You can use two single quotes within a quoted string.
nobel %>%
filter(tolower(winner)=="<NAME>")
# ## 13. Knights of the realm
#
# Knights in order
#
# **List the winners, year and subject where the winner starts with Sir. Show the the most recent first, then by name order.**
nobel %>%
filter(str_starts(tolower(winner), 'sir.')) %>%
select(winner, yr, subject) %>%
arrange(-yr, winner)
# ## 14. Chemistry and Physics last
#
# The expression **subject IN ('Chemistry','Physics')** can be used as a value - it will be 0 or 1.
#
# **Show the 1984 winners and subject ordered by subject and winner name; but list Chemistry and Physics last.**
nobel %>%
filter(yr==1984) %>%
arrange(subject %in% c('Chemistry', 'Physics'), subject, winner) %>%
select(winner, subject)
dbDisconnect(con)
|
R/03 SELECT from Nobel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from tensorflow.keras import models, layers, optimizers
from tensorflow.keras.utils import to_categorical
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import matplotlib.colors as colors
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib as mpl
from tensorflow.keras import regularizers
from tensorflow.keras.callbacks import LearningRateScheduler
import math
import matplotlib.colors as colors
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import tensorflow.keras.backend as K
plt.close()
df = pd.read_csv('facies_vectors.csv')
df.head()
df.isna().sum()
df = df.dropna()
df.isna().sum()
num_row, num_cols = df.shape
print(f"Number of datapoints : {num_row}")
print(f"Number of features : {num_cols}")
output_cols = [df.Facies.name]
print(f'Output feature are : {output_cols}')
input_cols = [cols for cols in df]
input_cols.remove(output_cols[0]) # input data = all data - output data.
input_cols = input_cols[3:] # start from Depth
del(input_cols[5])
print(f'Input layers are {input_cols}')
df[input_cols]
input_data = df[input_cols]
labels = df[output_cols]
print(input_data.head())
labels.head()
from sklearn.utils import shuffle
input_data, labels = shuffle(input_data, labels, random_state=42)
# +
### visualize out data
# 1=sandstone 2=c_siltstone 3=f_siltstone
# 4=marine_silt_shale 5=mudstone 6=wackestone 7=dolomite
# 8=packstone 9=bafflestone
facies_colors = ['#F4D03F', '#F5B041','#DC7633','#6E2C00',
'#1B4F72','#2E86C1', '#AED6F1', '#A569BD', '#196F3D']
facies_labels = ['SS', 'CSiS', 'FSiS', 'SiSh', 'MS',
'WS', 'D','PS', 'BS']
#facies_color_map is a dictionary that maps facies labels
#to their respective colors
facies_color_map = {}
for ind, label in enumerate(facies_labels):
facies_color_map[label] = facies_colors[ind]
def label_facies(row, labels):
return labels[ row['Facies'] -1]
df.loc[:,'FaciesLabels'] = df.apply(lambda row: label_facies(row, facies_labels), axis=1)
df['Facies_cnn'] = df['Facies']
df.describe()
# -
df.head()
df.isna().sum()
# +
def make_facies_log_plot(logs, facies_colors):
#make sure logs are sorted by depth
logs = logs.sort_values(by='Depth')
cmap_facies = colors.ListedColormap(
facies_colors[0:len(facies_colors)], 'indexed')
ztop=logs.Depth.min(); zbot=logs.Depth.max()
cluster=np.repeat(np.expand_dims(logs['Facies'].values,1), 100, 1)
f, ax = plt.subplots(nrows=1, ncols=6, figsize=(8, 12))
ax[0].plot(logs.GR, logs.Depth, '-g')
ax[1].plot(logs.ILD_log10, logs.Depth, '-')
ax[2].plot(logs.DeltaPHI, logs.Depth, '-', color='0.5')
ax[3].plot(logs.PHIND, logs.Depth, '-', color='r')
ax[4].plot(logs.PE, logs.Depth, '-', color='black')
im=ax[5].imshow(cluster, interpolation='none', aspect='auto',
cmap=cmap_facies,vmin=1,vmax=9)
divider = make_axes_locatable(ax[5])
cax = divider.append_axes("right", size="20%", pad=0.05)
cbar=plt.colorbar(im, cax=cax)
cbar.set_label((17*' ').join([' SS ', 'CSiS', 'FSiS',
'SiSh', ' MS ', ' WS ', ' D ',
' PS ', ' BS ']))
cbar.set_ticks(range(0,1)); cbar.set_ticklabels('')
for i in range(len(ax)-1):
ax[i].set_ylim(ztop,zbot)
ax[i].invert_yaxis()
ax[i].grid()
ax[i].locator_params(axis='x', nbins=3)
ax[0].set_xlabel("GR")
ax[0].set_xlim(logs.GR.min(),logs.GR.max())
ax[1].set_xlabel("ILD_log10")
ax[1].set_xlim(logs.ILD_log10.min(),logs.ILD_log10.max())
ax[2].set_xlabel("DeltaPHI")
ax[2].set_xlim(logs.DeltaPHI.min(),logs.DeltaPHI.max())
ax[3].set_xlabel("PHIND")
ax[3].set_xlim(logs.PHIND.min(),logs.PHIND.max())
ax[4].set_xlabel("PE")
ax[4].set_xlim(logs.PE.min(),logs.PE.max())
ax[5].set_xlabel('Facies')
ax[1].set_yticklabels([]); ax[2].set_yticklabels([]); ax[3].set_yticklabels([])
ax[4].set_yticklabels([]); ax[5].set_yticklabels([])
ax[5].set_xticklabels([])
f.suptitle('Well: %s'%logs.iloc[0]['Well Name'], fontsize=14,y=0.94)
make_facies_log_plot(
df[df['Well Name'] == 'SHRIMPLIN'],
facies_colors)
# +
#In addition to individual wells, we can look at how the various facies are represented by the entire training set.
#Let's plot a histgram of the number of training examples for each facies class
#count the number of unique entries for each facies, sort them by
#facies number (instead of by number of entries)
facies_counts = df['Facies'].value_counts().sort_index()
#use facies labels to index each count
facies_counts.index = facies_labels
facies_counts.plot(kind='bar',color=facies_colors,
title='Distribution of Training Data by Facies')
facies_counts
# -
input_data.isna().sum()
input_data = input_data.values
labels = labels.values
print(input_data.shape,labels.shape)
labels = labels.squeeze()
print(input_data.shape,labels.shape)
#Model_Seq
# normalize input data
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(input_data)
input_data = scaler.transform(input_data)
print(input_data[0:10])
labels[0:10]
np.isnan(input_data).sum()
val_data = input_data[0:640]
input_data = input_data[641:]
val_labels = labels[0:640]
labels = labels[641:]
input_data.shape
from CLR import CyclicLR
# +
## start model
from tensorflow.keras import regularizers
# Architecture
model_seq = models.Sequential()
model_seq.add(layers.Dense(512, activation='relu', input_shape = (input_data.shape[1],)))
model_seq.add(layers.Dropout(0.5))
model_seq.add(layers.Dense(256, activation='relu'))
model_seq.add(layers.Dropout(0.5))
model_seq.add(layers.Dense(128, activation='relu'))
model_seq.add(layers.Dropout(0.5))
model_seq.add(layers.Dense(128, activation='relu'))
model_seq.add(layers.Dropout(0.5))
model_seq.add(layers.Dense(64, activation='relu',kernel_regularizer = regularizers.l2()))
model_seq.add(layers.Dense(10, activation='softmax'))
model_seq.compile(
optimizer = optimizers.Adamax(),
loss = 'sparse_categorical_crossentropy',
metrics=['acc']
)
model_seq.summary()
# -
model = model_seq
from keras_lr_finder.lr_finder import LRFinder
lr_finder = LRFinder(model)
lr_finder.find(input_data, labels, start_lr=1e-6, end_lr=0.01,batch_size=128, epochs=80)
lr_finder.plot_loss(n_skip_beginning=10, n_skip_end=5)
lr_finder.plot_loss_change(sma=20, n_skip_beginning=10, n_skip_end=5, y_lim=(-0.001, 0.05))
clr = CyclicLR(base_lr=1e-6, max_lr=1e-2, step_size=8,mode='exp_range', gamma=0.99990)
K.set_value(model_seq.optimizer.lr, 0.01)
# +
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau, TensorBoard, TerminateOnNaN
callbacks = [
#ReduceLROnPlateau(monitor="val_loss",factor=0.2, patience=10, min_lr=5e-5, verbose=1),
clr,
ModelCheckpoint('model_seq.h5',monitor="val_loss", verbose=1, save_best_only=True, mode='auto'),
TerminateOnNaN()
]
history = model_seq.fit(
input_data,
labels,
epochs=1000,
validation_data=(val_data, val_labels),
verbose=2,
batch_size=64,
callbacks=callbacks
)
# +
def model_plots(history):
history_dict = history.history
# plot histories
epochs = history.epoch
epochs = epochs[1:]
epochs.append(len(epochs)+1)
## training loss and acc
acc = history_dict['acc']
loss = history_dict['loss']
## validation loss and acc
val_acc = history_dict['val_acc']
val_loss = history_dict['val_loss']
plt.figure("Losses")
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'r--', label='Validation loss')
plt.legend()
plt.title("Training and Validation loss")
plt.xlabel("Epochs")
plt.ylabel("Losses")
plt.show()
plt.figure("Accuracy")
plt.plot(epochs, acc, 'bo-', label='Training accuracy')
plt.plot(epochs, val_acc, 'r--', label='Validation accuracy')
plt.legend()
plt.title("Training and Validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.show()
model_plots(history)
# -
clr.history.keys()
# plot Learning rates
iterations = clr.history['iterations']
plt.figure("Learning curve")
plt.plot(iterations, clr.history["lr"])
plt.title("Cyclical Learning Rate (CLR)")
plt.xlabel("Training Iterations")
plt.ylabel("Learning Rate")
model_seq = models.load_model('model_seq.h5')
input_data.shape
cnn_data = input_data.reshape(input_data.shape[0], input_data.shape[1], 1)
cnnval_data = val_data.reshape(val_data.shape[0], val_data.shape[1], 1)
cnn_data.shape,cnnval_data.shape
# +
## start model
# Architecture
model_CNN = models.Sequential()
model_CNN.add(layers.Conv1D(64,3, activation='relu', input_shape = cnn_data.shape[1:],padding="same"))
model_CNN.add(layers.Conv1D(64,3, activation='relu', padding="same"))
model_CNN.add(layers.Dropout(0.5))
model_CNN.add(layers.Conv1D(64,3, activation='relu', padding="same"))
model_CNN.add(layers.Dropout(0.5))
model_CNN.add(layers.MaxPooling1D(2))
model_CNN.add(layers.Flatten())
# model.add(layers.Dense(16, activation='relu', input_shape = (input_data.shape[1],)))
# model.add(layers.Dense(32, activation='relu', kernel_regularizer=regularizers.l1_l2(l1=1e-4, l2=1e-4)))
# model.add(layers.Dense(64, activation='relu', kernel_regularizer=regularizers.l1_l2(l1=1e-4, l2=1e-4)))
# model.add(layers.Dense(128, activation='relu', kernel_regularizer=regularizers.l1_l2(l1=1e-4, l2=1e-4)))
# model.add(layers.Dense(256, activation='relu', kernel_regularizer=regularizers.l1_l2(l1=1e-4, l2=1e-4)))
# model.add(layers.Dense(512, activation='relu', kernel_regularizer=regularizers.l1_l2(l1=1e-4, l2=1e-4)))
# model.add(layers.Dense(1024, activation='relu', kernel_regularizer=regularizers.l1_l2(l1=1e-4, l2=1e-4)))
# model.add(layers.Dropout(0.4))
# model.add(layers.Dense(2048, activation='relu', kernel_regularizer=regularizers.l1_l2(l1=1e-4, l2=1e-4)))
#model.add(layers.Dense(1024, activation='relu',kernel_regularizer=regularizers.l2()))
model_CNN.add(layers.Dense(128, activation='relu',kernel_regularizer=regularizers.l2()))
model_CNN.add(layers.Dense(64, activation='relu',kernel_regularizer=regularizers.l2()))
model_CNN.add(layers.Dense(10, activation='softmax'))
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau, TensorBoard
model_CNN.compile(
optimizer = optimizers.Adam(lr=5e-5),
loss = 'sparse_categorical_crossentropy',
metrics=['acc']
)
model_CNN.summary()
callbacks = [
ReduceLROnPlateau(monitor="val_loss",factor=0.2, patience=10, min_lr=5e-6, verbose=1),
ModelCheckpoint('model_CNN.h5',monitor="val_loss", verbose=1, save_best_only=True, mode='auto')
]
history = model_CNN.fit(
cnn_data,
labels,
epochs=200,
validation_data=(cnnval_data, val_labels),
verbose=2,
batch_size=128,
callbacks=callbacks
)
def model_plots(history):
history_dict = history.history
# plot histories
epochs = history.epoch
epochs = epochs[1:]
epochs.append(len(epochs)+1)
## training loss and acc
acc = history_dict['acc']
loss = history_dict['loss']
## validation loss and acc
val_acc = history_dict['val_acc']
val_loss = history_dict['val_loss']
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.legend()
plt.title("Training and Validation loss")
plt.xlabel("Epochs")
plt.ylabel("Losses")
plt.show()
plt.figure()
plt.plot(epochs, acc, 'bo', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.legend()
plt.title("Training and Validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.show()
model_plots(history)
# +
#simple linear Model similar to SVM
import sys
input_data = df[input_cols]
input_stats = df[input_cols].describe()
means = input_stats.iloc[1]
stds = input_stats.iloc[2]
for col in input_cols:
input_data[col] = (input_data[col]-means[col])/stds[col]
print(input_data[0:10])
labels = df[output_cols]
(input_data, labels) = shuffle_datas(input_data, labels)
input_data = input_data.values
labels = labels.values
labels = to_categorical(labels)
print(f'Shape {input_data.shape},{labels.shape}')
val_data = input_data[0:800]
input_data = input_data[800:]
val_labels = labels[0:800]
labels = labels[800:]
# +
model_svm = models.Sequential()
model_svm.add(layers.Dense(64,activation='relu',input_shape = (input_data.shape[1],)))
model_svm.add(layers.Dense(10,activation='softmax'))
model_svm.compile(
optimizer = optimizers.Adam(2e-4),
loss = 'categorical_crossentropy',
metrics=['acc']
)
model_svm.summary()
# +
history = model_svm.fit(
input_data,
labels,
epochs=100,
validation_data=(val_data, val_labels),
verbose=2,
batch_size=1024
)
def model_plots(history):
history_dict = history.history
# plot histories
epochs = history.epoch
epochs = epochs[1:]
epochs.append(len(epochs)+1)
## training loss and acc
acc = history_dict['acc']
loss = history_dict['loss']
## validation loss and acc
val_acc = history_dict['val_acc']
val_loss = history_dict['val_loss']
plt.figure("Losses")
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.legend()
plt.title("Training and Validation loss")
plt.xlabel("Epochs")
plt.ylabel("Losses")
plt.show()
plt.figure("Accuracy")
plt.plot(epochs, acc, 'bo', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.legend()
plt.title("Training and Validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.show()
model_plots(history)
# +
#RNN
input_data = df[input_cols]
input_stats = df[input_cols].describe()
means = input_stats.iloc[1]
stds = input_stats.iloc[2]
for col in input_cols:
input_data[col] = (input_data[col]-means[col])/stds[col]
print(input_data[0:10])
labels = df[output_cols]
input_data = input_data.values
labels = labels.values
labels = to_categorical(labels)
print(f'Shape {input_data.shape},{labels.shape}')
val_data = input_data[0:500]
input_data = input_data[501:]
val_labels = labels[0:500]
labels = labels[501:]
input_data = np.expand_dims(input_data,axis=1)
val_data = np.expand_dims(val_data,axis=1)
print(f'Shape {input_data.shape},{labels.shape},{val_data.shape},{val_labels.shape}')
# -
input_data[0:10]
def expand_dim(x):
x = np.expand_dims(x,axis=0)
# +
## start model
# Architecture
model_RNN = models.Sequential()
model_RNN.add(layers.LSTM(512, activation='relu', input_shape = (input_data.shape[1], input_data.shape[2]), return_sequences=True))
model_RNN.add(layers.LSTM(256, return_sequences=True, activation='relu'))
model_RNN.add(layers.Dropout(0.5))
model_RNN.add(layers.LSTM(128, activation='relu', return_sequences=True))
model_RNN.add(layers.Dropout(0.5))
model_RNN.add(layers.LSTM(128, activation='relu', return_sequences=True))
model_RNN.add(layers.Dropout(0.5))
model_RNN.add(layers.LSTM(64, activation='relu', return_sequences=True))
model_RNN.add(layers.Dropout(0.5))
model_RNN.add(layers.LSTM(64, activation='relu', return_sequences=True))
model_RNN.add(layers.Dropout(0.5))
model_RNN.add(layers.LSTM(32, activation='relu', return_sequences=True))
model_RNN.add(layers.Dropout(0.5))
model_RNN.add(layers.LSTM(32, activation='relu', return_sequences=True))
model_RNN.add(layers.LSTM(10, activation='relu'))
model_RNN.add(layers.Dense(10,activation='softmax'))
# +
model_RNN.compile(
optimizer = optimizers.Adam(lr = 2e-4),
loss = 'categorical_crossentropy',
metrics=['acc']
)
model_RNN.summary()
# +
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau, TensorBoard
callbacks = [
EarlyStopping(monitor="val_loss",patience=30, verbose=1),
ReduceLROnPlateau(monitor="val_loss",factor=0.1, patience=5, min_lr=0.00004, verbose=1),
ModelCheckpoint('model_RNN.h5',monitor="val_loss", verbose=1, save_best_only=True, mode='auto')
]
history = model_RNN.fit(
input_data,
labels,
epochs=500,
validation_data=(val_data, val_labels),
verbose=2,
batch_size=256
#callbacks=callbacks
)
def model_plots(history):
history_dict = history.history
# plot histories
epochs = history.epoch
epochs = epochs[1:]
epochs.append(len(epochs)+1)
## training loss and acc
acc = history_dict['acc']
loss = history_dict['loss']
## validation loss and acc
val_acc = history_dict['val_acc']
val_loss = history_dict['val_loss']
plt.figure("Losses")
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.legend()
plt.title("Training and Validation loss")
plt.xlabel("Epochs")
plt.ylabel("Losses")
plt.show()
plt.figure("Accuracy")
plt.plot(epochs, acc, 'bo', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.legend()
plt.title("Training and Validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.show()
model_plots(history)
# -
import pandas as pd
well_data = pd.read_csv('validation_data_nofacies.csv')
well_data.head()
feature_vectors = df.drop(['Formation', 'Well Name','Facies','Depth','FaciesLabels'], axis=1)
well_data['Well Name'] = well_data['Well Name'].astype('category')
well_features = well_data.drop(['Formation', 'Well Name'], axis=1)
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(feature_vectors)
X_unknown = scaler.transform(well_features)
print(X_unknown.shape)
X_unknown = X_unknown[:,1:]
print(X_unknown.shape)
well_data.head()
#predict facies of unclassified data
y_unknown_seq = model_seq.predict_classes(X_unknown)
well_data['Facies_seq'] = y_unknown_seq
y_unknown_seq
X_unknown = X_unknown.reshape(830,7,1)
y_unknown_CNN = model_CNN.predict_classes(X_unknown)
well_data['Facies_CNN'] = y_unknown_CNN
y_unknown_CNN
X_unknown = X_unknown.reshape(-1,1,7)
y_unknown_RNN = model_RNN.predict_classes(X_unknown)
y_unknown_RNN.squeeze()
well_data['Facies_RNN'] = y_unknown_RNN
well_data['Well Name'].unique()
well_data
def predict_facies_log_plot(logs, facies_colors):
#make sure logs are sorted by depth
logs = logs.sort_values(by='Depth')
cmap_facies = colors.ListedColormap(
facies_colors[0:len(facies_colors)], 'indexed')
ztop=logs.Depth.min(); zbot=logs.Depth.max()
cluster_seq=np.repeat(np.expand_dims(logs['Facies_seq'].values,1), 100, 1)
cluster_CNN=np.repeat(np.expand_dims(logs['Facies_CNN'].values,1), 100, 1)
cluster_RNN=np.repeat(np.expand_dims(logs['Facies_RNN'].values,1), 100, 1) #1
f, ax = plt.subplots(nrows=1, ncols=8, figsize=(10, 14)) #2: ncols=6 to 7
ax[0].plot(logs.GR, logs.Depth, '-g')
ax[1].plot(logs.ILD_log10, logs.Depth, '-')
ax[2].plot(logs.DeltaPHI, logs.Depth, '-', color='0.5')
ax[3].plot(logs.PHIND, logs.Depth, '-', color='r')
ax[4].plot(logs.PE, logs.Depth, '-', color='black')
im_seq=ax[5].imshow(cluster_seq, interpolation='none', aspect='auto', #3
cmap=cmap_facies,vmin=1,vmax=9)
divider_seq = make_axes_locatable(ax[5])
cax_seq = divider_seq.append_axes("right", size="20%", pad=0.05)
cbar_seq=plt.colorbar(im_seq, cax=cax_seq)
cbar_seq.set_label((17*' ').join([' SS ', 'CSiS', 'FSiS',
'SiSh', ' MS ', ' WS ', ' D ',
' PS ', ' BS ']))
cbar_seq.set_ticks(range(0,1)); cbar_seq.set_ticklabels('')
im_CNN=ax[6].imshow(cluster_CNN, interpolation='none', aspect='auto', #3
cmap=cmap_facies,vmin=1,vmax=9)
divider_CNN = make_axes_locatable(ax[6])
cax_CNN = divider_CNN.append_axes("right", size="20%", pad=0.05)
cbar_CNN=plt.colorbar(im_CNN, cax=cax_CNN)
cbar_CNN.set_label((17*' ').join([' SS ', 'CSiS', 'FSiS',
'SiSh', ' MS ', ' WS ', ' D ',
' PS ', ' BS ']))
cbar_CNN.set_ticks(range(0,1)); cbar_CNN.set_ticklabels('')
im_RNN=ax[7].imshow(cluster_RNN, interpolation='none', aspect='auto', #3
cmap=cmap_facies,vmin=1,vmax=9)
divider_RNN = make_axes_locatable(ax[7])
cax_RNN = divider_RNN.append_axes("right", size="20%", pad=0.05)
cbar_RNN=plt.colorbar(im_RNN, cax=cax_RNN)
cbar_RNN.set_label((17*' ').join([' SS ', 'CSiS', 'FSiS',
'SiSh', ' MS ', ' WS ', ' D ',
' PS ', ' BS ']))
cbar_RNN.set_ticks(range(0,1)); cbar_RNN.set_ticklabels('')
num_facies_plot = 3
for i in range(len(ax)-num_facies_plot): #3 two is the problem
ax[i].set_ylim(ztop,zbot)
ax[i].invert_yaxis()
ax[i].grid()
ax[i].locator_params(axis='x', nbins=3)
ax[0].set_xlabel("GR")
ax[0].set_xlim(logs.GR.min(),logs.GR.max())
ax[1].set_xlabel("ILD_log10")
ax[1].set_xlim(logs.ILD_log10.min(),logs.ILD_log10.max())
ax[2].set_xlabel("DeltaPHI")
ax[2].set_xlim(logs.DeltaPHI.min(),logs.DeltaPHI.max())
ax[3].set_xlabel("PHIND")
ax[3].set_xlim(logs.PHIND.min(),logs.PHIND.max())
ax[4].set_xlabel("PE")
ax[4].set_xlim(logs.PE.min(),logs.PE.max())
ax[5].set_xlabel('Facies_seq')
ax[6].set_xlabel('Facies_CNN')
ax[7].set_xlabel('Facies_RNN') #4
ax[1].set_yticklabels([]); ax[2].set_yticklabels([]); ax[3].set_yticklabels([])
ax[4].set_yticklabels([]); ax[5].set_yticklabels([]); ax[6].set_yticklabels([]) ; ax[7].set_yticklabels([]) #5
ax[5].set_xticklabels([]); ax[6].set_xticklabels([]); ax[7].set_xticklabels([]) #6
f.suptitle('Well: %s'%logs.iloc[0]['Well Name'], fontsize=14,y=0.94)
facies = {'SS':1,'CSiS':2,'FSiS':3,'SiSh':4,'MS':5,'WS':6,'D':7,'PS':8,'BS':9} #7 for legends
hands = []
for k, col in zip(facies.keys(), facies_colors):
hands.append(mpatches.Patch(color=col, label=k))
plt.legend(handles=hands, loc=(2, 0.5), fontsize=18)
# +
predict_facies_log_plot(well_data[df['Well Name'] == 'SHRIMPLIN'], facies_colors)
labels = to_categorical(labels)
# -
from sklearn.metrics import f1_score_score
f1=f1_score(y_unknown_CNN)
model_1 = models.Sequential()
model_1.add(layers.Dense(100,input_shape=(2,1)))
for unit in [10,10,10]:
model_1.add(layers.Dense(unit,activation='relu'))
model_1.compile(optimizer='rmsprop',
loss = 'mse'
)
model_1.summary()
|
well_logs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py37
# language: python
# name: py37
# ---
import torch
from torch import nn
from torch.utils.data import DataLoader
from torchvision import datasets
from torchvision.transforms import ToTensor, Lambda, Compose
import matplotlib.pyplot as plt
# DataLoader wrap an iterable around Dataset which stores samples and corresponding labels.
# +
# Download training data from open datasets.
training_data = datasets.FashionMNIST(
root="data",
train=True,
download=True,
transform=ToTensor(),
)
# Download test data from open datasets.
test_data = datasets.FashionMNIST(
root="data",
train=False,
download=True,
transform=ToTensor(),
)
# +
batch_size = 64
# Create data loaders.
train_dataloader = DataLoader(training_data, batch_size=batch_size)
test_dataloader = DataLoader(test_data, batch_size=batch_size)
for X, y in test_dataloader:
print("Shape of X [N, C, H, W]: ", X.shape)
print("Shape of y: ", y.shape, y.dtype)
break
# +
# Get cpu or gpu device for training.
device = "cuda" if torch.cuda.is_available() else "cpu"
print("Using {} device".format(device))
# Define model
class NeuralNetwork(nn.Module):
def __init__(self):
super(NeuralNetwork, self).__init__()
self.flatten = nn.Flatten()
self.linear_relu_stack = nn.Sequential(
nn.Linear(28*28, 512),
nn.ReLU(),
nn.Linear(512, 512),
nn.ReLU(),
nn.Linear(512, 10)
)
def forward(self, x):
x = self.flatten(x)
logits = self.linear_relu_stack(x)
return logits
model = NeuralNetwork().to(device)
print(model)
# +
loss_fn = nn.CrossEntropyLoss()
# hold the current state and will update the parameters based on the computed gradients.
optimizer = torch.optim.SGD(model.parameters(), lr=1e-3)
# -
def train(dataloader, model, loss_fn, optimizer):
size = len(dataloader.dataset)
model.train()
for batch, (X, y) in enumerate(dataloader):
X, y = X.to(device), y.to(device)
# Compute prediction error
pred = model(X)
loss = loss_fn(pred, y)
# __Backpropagation__
# zero out accumlated gradients
optimizer.zero_grad()
# Computes the gradient of current tensor w.r.t. graph leaves.
# The graph is differentiated using the chain rule.
loss.backward()
optimizer.step()
if batch % 100 == 0:
loss, current = loss.item(), batch * len(X)
print(f"loss: {loss:>7f} [{current:>5d}/{size:>5d}]")
def test(dataloader, model, loss_fn):
size = len(dataloader.dataset)
num_batches = len(dataloader)
model.eval()
test_loss, correct = 0, 0
with torch.no_grad():
for X, y in dataloader:
X, y = X.to(device), y.to(device)
pred = model(X)
test_loss += loss_fn(pred, y).item()
correct += (pred.argmax(1) == y).type(torch.float).sum().item()
test_loss /= num_batches
correct /= size
print(f"Test Error: \n Accuracy: {(100*correct):>0.1f}%, Avg loss: {test_loss:>8f} \n")
epochs = 5
for t in range(epochs):
print(f"Epoch {t+1}\n-------------------------------")
train(train_dataloader, model, loss_fn, optimizer)
test(test_dataloader, model, loss_fn)
print("Done!")
torch.save(model.state_dict(), "model.pth")
print("Saved PyTorch Model State to model.pth")
model = NeuralNetwork()
model.load_state_dict(torch.load("model.pth"))
# Use loaded model to make predictions...
# +
classes = [
"T-shirt/top",
"Trouser",
"Pullover",
"Dress",
"Coat",
"Sandal",
"Shirt",
"Sneaker",
"Bag",
"Ankle boot",
]
model.eval()
x, y = test_data[0][0], test_data[0][1]
with torch.no_grad():
pred = model(x)
predicted, actual = classes[pred[0].argmax(0)], classes[y]
print(f'Predicted: "{predicted}", Actual: "{actual}"')
# -
|
modules/module_20/pytorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task 2: Create Bengali word embeddings
# ## Import libraries
# +
# Imports
import re
import string
import json
from datetime import datetime
from collections import defaultdict, Counter
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import tqdm
import torch
import torch.nn as nn
from torch.nn import Module
import torch.optim as optim
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from sklearn.model_selection import train_test_split
from nltk.corpus import stopwords
device = 'cuda'
import random
torch.manual_seed(123)
torch.cuda.manual_seed(234)
np.random.seed(345)
random.seed(456)
torch.manual_seed(567)
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
# -
# ## Load data
# +
ben_train_df = pd.read_csv('../hindi_bengali/save/bengali_hatespeech_sample_train_preprocessed.csv')
ben_test_df = pd.read_csv('../hindi_bengali/save/bengali_hatespeech_sample_test_preprocessed.csv')
display(ben_train_df.head())
# +
# train data:
# remove empty texts
ben_train_df = ben_train_df[ben_train_df.sentence.str.len() > 0]
# extract sentences and labels
train_sentences = [text.split() for text in ben_train_df['sentence']]
train_labels = ben_train_df['hate'].to_numpy()
# test data:
# remove empty texts
ben_test_df = ben_test_df[ben_test_df.sentence.str.len() > 0]
# extract sentences and labels
test_sentences = [text.split() for text in ben_test_df['sentence']]
test_labels = ben_test_df['hate'].to_numpy()
# -
# ### Print out data/statistics
print('Train data:')
print(train_sentences[:3])
print(train_labels)
print()
print('Test data:')
print(test_sentences[:3])
print(test_labels)
# ## Prepare vocab set
# +
# load mapping {word -> id} and {id -> word}
with open('../hindi_bengali/save/bengali_word_to_int_dict.json') as f:
word_to_int = json.load(f)
with open('../hindi_bengali/save/bengali_int_to_word_dict.json') as f:
int_to_word = json.load(f)
int_to_word = {int(k) : v for k, v in int_to_word.items()}
with open('../hindi_bengali/save/bengali_word_counter.json') as f:
word_counter = json.load(f)
# get vocab_size
vocab_size = len(word_to_int)
print(f'vocab_size: {vocab_size}')
# get total occurences
total_words = sum(word_counter.values())
print(f'total word occurences: {total_words}')
# -
# ## Define sub-sampling
def sampling_prob(word):
z = word_counter[word] / total_words
p_keep = ((z/0.000001)**0.5 + 1) * (0.000001/z)
return p_keep
# ## skip-gram
def get_target_context(sentence: list(str())):
for i, word in enumerate(sentence):
for j, context_word in enumerate(sentence[i-window_size:i+window_size+1]):
if j != i and random.random() < sampling_prob(context_word):
yield (torch.tensor(word_to_int[word], dtype=torch.long).unsqueeze(0),
torch.tensor(word_to_int[context_word], dtype=torch.long).unsqueeze(0))
# # Train word-embeddings
# ## hyper-parameters
window_size = 10
embedding_size = 300
learning_rate = 0.01
batch_size = 256
epochs = 100
# ## Model
# +
class Word2Vec(Module):
def __init__(self):
super(Word2Vec, self).__init__()
self.embed = nn.Embedding(vocab_size, embedding_size)
self.fc = nn.Linear(embedding_size, vocab_size)
def forward(self, word_id):
out = self.embed(word_id)
out = self.fc(out)
return out.squeeze(1)
def to_embed(self, word_id):
return self.embed(word_id)
word2vec = Word2Vec()
save_path = './save/bengali_word2vec.pt'
torch.save(word2vec.state_dict(), save_path)
display(word2vec.parameters)
# -
# ## Loss function and Optimizer
optimizer = optim.Adam(word2vec.parameters(), lr=learning_rate)
criterion = nn.CrossEntropyLoss()
# ## Dataset
class W2VDataset(Dataset):
def __init__(self, sentences):
self.data = []
for sentence in sentences:
for data_point in get_target_context(sentence):
self.data.append(data_point)
def __len__(self):
return len(self.data)
def __getitem__(self, index):
return self.data[index]
# ## Learning parameters
# +
# load initial weights
word2vec.load_state_dict(torch.load(save_path))
word2vec = word2vec.to(device)
# training
early_stop = 5
list_loss = []
for epoch in range(1, epochs+1):
train_dataset = W2VDataset(train_sentences)
train_loader = DataLoader(train_dataset, batch_size=batch_size, shuffle=True)
losses = 0.
cnt = 0
word2vec.train()
for words, context_words in tqdm(train_loader):
optimizer.zero_grad()
pred = word2vec(words.to(device))
loss = criterion(pred, context_words.squeeze(1).to(device))
loss.backward()
optimizer.step()
losses += loss.detach().item() * len(words)
cnt += len(words)
epoch_loss = losses / cnt
print(f'Epoch {epoch:2}: training loss: {epoch_loss:.4f} over {cnt} training points.')
if epoch % 10 == 0:
# save check-point embedding
embedding_weights = word2vec.embed.state_dict()
torch.save(embedding_weights, f'./save/embedding_checkpoints/embedding_weights_{epoch}_epoch_{embedding_size}_dim_{window_size}_wsize.pt')
list_loss.append(epoch_loss)
if len(list_loss) > early_stop and min(list_loss[-early_stop:]) > min(list_loss[:-early_stop]):
print('Training loss is not reducing anymore, terminate.')
break
print("Training finished")
# -
# save the word-embedding layer weights
embedding_weights = word2vec.embed.state_dict()
torch.save(embedding_weights, f'save/embedding_weights.pt')
|
Task_2/bengali_bengali/Task-2c.1_bengali_word_embeddings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center>
# <h1> ILI285 - Computación Científica I / INF285 - Computación Científica </h1>
# <h2> Finding 2 Chebyshev points graphycally </h2>
# <h2> <a href="#acknowledgements"> [S]cientific [C]omputing [T]eam </a> </h2>
# <h2> Version: 1.02</h2>
# </center>
# ## Table of Contents
# * [Finding 2 Chebyshev points](#cheb)
# * [Python Modules and Functions](#py)
# * [Acknowledgements](#acknowledgements)
import numpy as np
import matplotlib.pyplot as plt
from ipywidgets import interact
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import matplotlib as mpl
mpl.rcParams['font.size'] = 14
mpl.rcParams['axes.labelsize'] = 20
mpl.rcParams['xtick.labelsize'] = 14
mpl.rcParams['ytick.labelsize'] = 14
# %matplotlib inline
# <div id='cheb' />
#
# ## Finding 2 Chebyshev points
# We compute them so we can compare them later.
n=2
i=1
theta1=(2*i-1)*np.pi/(2*n)
i=2
theta2=(2*i-1)*np.pi/(2*n)
c1=np.cos(theta1)
c2=np.cos(theta2)
# Recall that the Chebyshev points are points that minimize the following expression:
# $$
# \displaystyle{\omega(x_1,x_2,\dots,x_n)=\max_{x} |(x-x_1)\,(x-x_2)\,\cdots\,(x-x_n)|}.
# $$
# This comes from the Interpolation Error Formula (I hope you remember it, otherwise see the textbook or the classnotes!).
# In this notebook, we will find the $\min$ for 2 points,
# this means:
# $$
# [x_1,x_2]= \displaystyle{\mathop{\mathrm{argmin}}_{x_1,x_2\in [-1,1]}} \,\omega(x_1,x_2)=\displaystyle{\mathop{\mathrm{argmin}}_{x_1,x_2\in [-1,1]}}\,
# \max_{x\in [-1,1]} |(x-x_1)\,(x-x_2)|.
# $$
# For doing this, we first need to build $\omega(x_1,x_2)$,
N=50
x=np.linspace(-1,1,N)
w = lambda x1,x2: np.max(np.abs((x-x1)*(x-x2)))
wv=np.vectorize(w)
# Now we need to evaluate $\omega(x_1,x_2)$ over the domain $\Omega=[0,1]^2$.
[X,Y]=np.meshgrid(x,x)
W=wv(X,Y)
# With this data, we can now plot the function $\omega(x_1,x_2)$ on $\Omega$.
# The minimun value of is shown by the color at the bottom of the colorbar.
# By visual inspection, we see that we have two mins.
# They are located at the bottom right and top left.
plt.figure(figsize=(8,8))
#plt.contourf(X, Y, W,100, cmap=cm.hsv, antialiased=False)
plt.contourf(X, Y, W,100, cmap=cm.nipy_spectral, antialiased=False)
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.colorbar()
plt.show()
# Finally, we have included the min in the plot and we see the agreement between the min of $\omega(x_1,x_2)$ and the Chebyshev points found.
plt.figure(figsize=(8,8))
plt.contourf(X, Y, W,100, cmap=cm.nipy_spectral, antialiased=False)
plt.plot(c1,c2,'k.',markersize=16)
plt.plot(c2,c1,'k.',markersize=16)
plt.colorbar()
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.show()
# <div id='py' />
#
# ## Python Modules and Functions
# An interesting module:
# https://docs.scipy.org/doc/numpy-1.13.0/reference/routines.polynomials.chebyshev.html
# <div id='acknowledgements' />
#
# # Acknowledgements
# * _Material created by professor <NAME>_ (`<EMAIL>`). DI UTFSM. _May 2018._
# * _Update June 2020 - v1.02 - C.Torres_ : Fixing formatting issues.
|
SC1/Bonus_Finding_2_Chebyshev_Points_Graphically.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from requests import get
from bs4 import BeautifulSoup
from html.parser import HTMLParser
import pandas as pd
import numpy as np
import pickle
# +
club_power_index = get('https://projects.fivethirtyeight.com/global-club-soccer-rankings/', 'lxml')
''' Grabbing the country names, scores and ranks '''
club_power_index = BeautifulSoup(club_power_index.text, 'html.parser')
team_list=[p.text for p in club_power_index.findAll('div', attrs={'class':'name'})]
score_list=[p.text for p in club_power_index.findAll('td', attrs={'class':'num'}) if len(p.text)>3]
league_list=[p.text.strip() for p in club_power_index.findAll('td', attrs={'class':'league drop-5'}) ]
country_list=[p.text.strip() for p in club_power_index.findAll('td', attrs={'class':'country drop-1'}) ]
# -
team_list=team_list[:600]
score_list=score_list[:600]
league_list=league_list[:600]
country_list=country_list[:600]
club_score_df=pd.DataFrame(list(zip(score_list, league_list, country_list)),
index=team_list, columns=["score", "league", "country"])
club_score_df.head(30)
# +
# keep only teams in 7 major european leagues
# we find that Austrian top soccer league is also named Bundasliga and Russian's league named
# Premier League. To avoid ambiguity, we use country as a selection criteria
mask1=club_score_df['league'].isin(
["Premier League", "Serie A", "La Liga", "Bundesliga", "Ligue 1","Primeira Liga","Eredivisie"])
club_score_df=club_score_df[mask1]
mask2=club_score_df['country'].isin(['England', 'Germany', 'France', 'Spain', 'Italy', 'Netherlands',
'Portugal'])
club_score_df=club_score_df[mask2]
# -
club_score_df.reset_index(level=0, inplace=True)
club_score_df.rename(columns={'index':'club'}, inplace=True)
club_score_df
# +
def team_name_unify(team):
if team=='Man. City':
return 'Manchester City'
if team=='Man. United':
return 'Manchester United'
if team=='Tottenham':
return 'Tottenham Hotspur'
if team== 'West Ham':
return 'West Ham United'
if team=='Leicester':
return 'Leicester City'
if team=='Newcastle':
return 'Newcastle United'
if team =='PSG':
return 'Paris Saint-Germain'
if team == 'Dortmund':
return 'Borussia Dortmund'
# if team =='Hoffenheim':
# return 'TSG 1899 Hoffenheim'
if team =='Eintracht':
return 'Eintracht Frankfurt'
# if team == 'Leverkusen':
# return 'Bayer 04 Leverkusen'
if team == 'Norwich':
return 'Norwich City'
if team =='Sheffield Utd':
return 'Sheffield United'
if team == 'Brighton':
return 'Brighton & Hovelbion'
if team =='Bordeaux':
return 'Girondins Bordeaux'
if team == 'St Étienne':
return 'Saint-Étienne'
if team == 'Twente':
return 'Twente Enschede'
else:
return team
club_score_df.club=club_score_df.club.apply(team_name_unify)
# -
club_score_df['score']=pd.to_numeric(club_score_df['score'])
pickle.dump(club_score_df[['club','score']], open('./temporary_pkl/club_score_df.pkl', 'wb'))
avrg_sc_by_league_dict = club_score_df.groupby(
by='league')['score'].mean().round(3).to_dict()
avrg_sc_by_league_dict
pickle.dump(avrg_sc_by_league_dict, open('./temporary_pkl/avrg_sc_by_league_dict.pkl', 'wb'))
list_short=club_score_df.club.to_list()
pkl_file = open('./temporary_pkl/list.pkl','rb')
list_long = pickle.load(pkl_file)
pkl_file.close()
pickle.dump(club_score_df, open('./temporary_pkl/club_score_df.pkl', 'wb'))
list_long
club_score_df
|
.ipynb_checkpoints/scrape+clean data-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import urllib
from bs4 import BeautifulSoup
import requests
import numpy
import os
import re
import json
import unicodedata
import datetime
url_general = 'https://www.factcheck.org/the-factcheck-wire/'
html_general = requests.get(url_general).text
soup_general = BeautifulSoup(html_general, 'html.parser')
# +
num_pags = int(soup_general.find_all('span', attrs={'class' : 'sr-only'})[1].text)
print(num_pags)
# -
def replace_characters(text):
text_aux = ''.join(c for c in unicodedata.normalize('NFD', text.strip())
if unicodedata.category(c) != 'Mn')
return re.sub(r'[^\w\s]', '', text_aux.lower().replace(u'\xa0', u' ').replace(u'\n', u' ')).split(" ")
def replace_characters_title(text):
return ''.join(c for c in unicodedata.normalize('NFD', text.strip())
if unicodedata.category(c) != 'Mn')
# +
bulos = {}
cont = 0
for i in range(1, num_pags+1):
url = 'https://www.factcheck.org/the-factcheck-wire/page/' + str(i)
html = requests.get(url).text
soup = BeautifulSoup(html, 'html.parser')
for h3 in soup.find_all('h3',attrs={'class' : 'entry-title'}):
a = h3('a')[0]
title = a.text
link = a.get('href', None)
words = dict()
html_new = requests.get(link, allow_redirects=False).text
soup_new = BeautifulSoup(html_new, 'html.parser')
title_list = re.sub(r'[^\w\s]', '', title.lower()).split(" ")
for w in title_list:
if w in words:
words[w] += 1
else:
words[w] = 1
new_text = soup_new.find('div',attrs={'class' : "entry-content"})
# Obtener la fecha
date = soup_new.find(class_="entry-date published updated")
for d in date:
new_date = datetime.datetime.strptime(str(d), "%B %d, %Y")
formated_date = new_date.strftime('%Y/%m/%d')
for text in new_text('p'):
for w in replace_characters(text.get_text()):
if w in words:
words[w] += 1
else:
words[w] = 1
bulos[cont] = {"titulo" : replace_characters_title(title), "link" : link, "date" : formated_date, "words_count" : words}
cont += 1
# +
#bulos
# -
len(bulos)
with open('fakenewsFactCheck.json', 'w') as fp:
json.dump(bulos, fp)
|
WebScraping/FactCheckWebScraping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# ## Contents
#
# - [Abstract](index.html)
# - [Project Motivation](motivation.html)
# - [Biological & Theoretical Background](background.html)
# - [Model Structure](structure.html)
# - [Usage](usage.html)
# - [First Steps: Pyro](pyro.html)
# - [Model Reconstruction](model.html)
# - [Performance Comparison](performance.html)
# - [Conclusions](conclusions.html)
#
# # Model Structure
#
# ## Data Structure
#
# Detailed descriptions of how data was collected and processed can be found in the Marks lab's papers describing their [EVmutation](https://www.nature.com/articles/nbt.3769) and [DeepSequence](https://www.nature.com/articles/s41592-018-0138-4) models. For ease of model testing and comparison, datasets prepared by the lab were used exclusively for this project. This preparation involved the following process: For a particular protein of interest (the focus sequence of the data set), a multiple-sequence alignment was performed with five iterations of the HMM homology search tool `jackhmmer` against the UniRef database of nonredundant protein sequences. The threshold for inclusion is 80% coverage, as this was found to be a good value for ensuring that sufficient sequence alignments were generated but the sequences aligned were still sufficiently similar.
#
# The biomolecule sequences found in databases have biases due to human sampling (as some phylogeny have been sequenced more frequently than other) and due to evolutionary sampling (as some proteins are over-represented in some groups of species). As such, before applying data to the model, a sequence re-weighting schedule is performed to reduce these biases. Each sequence weight is calculated as the reciprocal of the number of sequences within a given Hamming distance cutoff. The sample size, N is thus changed to be the sum of of these sequence weights. The cutoff for this re-weighting schedule has been found to work well as 0.2 (80% sequence identity) for non-viral biomolecules.
#
# As will be discussed below, loading of prepared data sets and performance of this re-weighting scheme is done by the data helper class.
#
# * * *
#
# ## Code Structure
#
# ### pt_helper.py
#
# Dependencies: numpy, torch
#
# This module contains important classes and functions for data loading and mutation effect prediction. This module is largely unchanged from the original version - the only changes made are to remove Theano dependence and facilitate Torch/GPU compatibility.
#
# #### DataHelper Class
#
# The main component of this module, responsible for loading data from datasets, calculating sequence weight. Configures the data set, performs one-hot encoding of amino acids, generates alignment and contains the functions required to perform ELBO calculations for log-ratio calculations used in mutation effect prediction.
#
# #### gen_simple_job_string
#
# Generates a basic job string specifying the key parameters of the model - used during model performance evaluation. Job string used for output and model file names
#
# #### gen_job_string
#
# Generates a detailed job string specifying model parameters, used for output and model file names.
#
# ----
#
# ### pt_model.py
#
# Dependencies: numpy, scipy, torch
#
# The module for construction of the model. All model classes inherit torch's nn.Module class and call `super(cls_name, self).__init__()` to add itself to this class. In doing so, all model classes can access each other, creating an easy-to-use modular structure for building models. In doing this, the Encoder class, for example, can be shared easily by both the MLE and SVI versions of the VAE.
#
# #### set_tensor_environment
#
# Function used by all model classes to set correct torch tensor type with respect to float-size and cuda/GPU capabilities (both of which specified in training.)
#
# #### Encoder Class
#
# sub-class used to define the encoder model for both SVI and MLE forms of the VAE model. Receives arguments from full VAE class specifying parameters.
#
# In `__init__` the encoder architecture is specified, with the option for convolving the input layer, non-linear functions are set up and the weights of the layers are initialised in accordance with the original model (weights are glorot-normal initialised, biases are set to 0.1 apart from the output log-sigma bias, which is set to -5). `forward()` defines the forward pass, accepting data batch $x$ and passing the data through the architecture to return `z_mu` and `z_logsig`.
#
# #### DecoderMLE Class
#
# sub-class for decoder where parameters are not subject to variational approximation (parameters determined through maximum likelihood estimation rather than stochastic variational inference).
#
# `__init__` creates architecture, initialises weights as per original model, sets up non-linear functions and accepts arguments passed from the full VAE model. In setting up the architecture, if so specified, will create sparsity parameters, a final temperature parameter, a convolutional layer for the output and a dropout layer. `forward()` accepts two arguments; the original input `x` which is used in calculations for the ELBO, and `z`, which is passed through the decoder architecture. If sparsity is used, `forward` takes sparsity parameters and tiles them to correct dimension. If passed sparsity argument is `'logit'` then the sigmoid of these parameters is applied to the final weights, otherwise the exponential is applied. If temperature or convolution are used, they are also applied here. Once the final output of the decoder is created, log(P(x|z)) is calculated. The forward pass returns the reconstruction of `x`, this log-probability value and the values of the final output.
#
# #### DecoderSVI Class
#
# sub-class for decoder that performs variational approximation on parameters.
#
# `__init__` accepts arguments from full VAE class, sets up parameters and architecture, initialises variables and non-linear functions to be used in forward pass. For each weight, mu and sigma weights and biases are specified rather than just weights themselves, and these are used to sample the used weights. Since the weights and biases used in the forward pass of the decoder are actually sampled from these learnt weights, the layers specified here serve only to act as containers for the weights that are learnt, as opposed to being the actual objects applied in the forward pass. Another key difference here is that two containers are constructed; first a list called `variational_param_identifiers` which stores an identifier key for all decoder parameters subject to variational parameters other than the sparsity parameters, for which loss is calculated separately. Second, `variational_param_name_to_sigma`, a dictionary mapping each parameter to a prior sigma value for loss calculations - the default for all prior sigmas is 1, this dictionary is created to allow flexibility in changing this.
#
# The `sampler` function inherits the GPU-enabled torch random number generator constructed in the VAE class, and applies this in for the decoder's own reparameterisation trick function for sampling from a gaussian distribution.
#
# `forward()` accepts `x` and `z` as arguments, passes `z` through the model, tiling and applying sparsity parameters if specified, as well as convolutions and a final temperature parameter. The key difference is that for each step, the `sampler` function is used to create weights and biases sampled from the model's parameters, and these sampled values are what is applied to the data. Once the reconstruction of x has been created, log(P(x|z)) is calculated and the forward pass returns the reconstruction of `x`, this log-probability value and the values of the final output.
#
# #### VAE_MLE Class
#
# Class that constructs the full bayesian VAE model.
#
# `__init__` adds the class to nn.Module, accepts parameters passed by user in training, sets up the correct torch tensor environment, and creates the encoder and decoder as per these parameters.
#
# `sampler` is a function that, through the reparameterization trick, applies a torch random number generator to perform gaussian sampling.
#
# `_anneal` is a function used to perform annealing if specified. This is where the KLD loss is down-scaled in early updates to allow the model to 'learn' the data first, increasing stability. Default is to not use annealing, and annealing is not used in performance comparisons.
#
# `update` takes in the log(P(x|z)) value from the decoder, the z-mu and z-logsigma values returned by the encoder, the update number (for annealing should it be activated) and the effective sequence number. Calculates the latent-space loss. If sparsity is used, will calculate the sparsity loss according to the specific sparsity prior selected. Calculates the l2-regularisation penalty term. Applies annealing to the KLD loss if used, applies scaling as well should this be used (should the user want the model to focus on reconstruction rather than KLD loss - by default this is not applied). Finally, calculates the log(P(x)) value approximation based on ELBO. returns this ELBO, the log(P(x|z)) value, the regularization loss and the latent space loss.
#
# `likelihoods` takes in a particular `x` and returns the log(P(x)) estimation for that `x`, used in mutation effect prediction. This and subsequent functions are used in prediction and testing rather than training.
#
# `all_likelihood_components` takes in a particular `x` and returns the log(P(x)) estimation, the latent space KLD loss and the log(P(x|z)) value.
#
# `recognize` takes in a particular `x` and returns the mu and log-sigma of the latent variable outputted by the model's encoder.
#
# `get_pattern_activations` takes in a particular `x` and returns the output of the decoder's final layer from that x.
#
# #### VAE_SVI Class
#
# Class that constructs the full bayesian VAE model.
#
# `__init__` adds the class to nn.Module, accepts parameters passed by user in training, sets up the correct torch tensor environment, and creates the encoder and decoder as per these parameters.
#
# `KLD_diag_gaussians` is a function to calculate the KL divergence between two diagonal gaussians, used throughout the variational approximation loss calculations.
#
# `sampler` is a function that, through the reparameterization trick, applies a torch random number generator to perform gaussian sampling.
#
# `_anneal` is a function used to perform annealing if specified. This is where the KLD loss is down-scaled in early updates to allow the model to 'learn' the data first, increasing stability. Default is to not use annealing, and annealing is not used in performance comparisons.
#
# `gen_KLD_params` iterates through all parameters identified by the `variational_param_identifiers` and, by accessing the mu and logsigma values, calculates the KL divergence loss, summing across all the variational parameters and returning this sum.
#
# `gen_KLD_sparsity` calculates the same loss for the sparsity parameters, which must be calculated differently due to the sparsity constraints. Here, loss can be calculated differently depending on which sparsity prior is chosen - the continuous relaxation of a spike and slab prior is the default that is used for performance comparisons.
#
# `update` takes in the log(P(x|z)) value from the decoder, the z-mu and z-logsigma values returned by the encoder, the update number (for annealing should it be activated) and the effective sequence number. applies the KLD loss of variational decoder parameters, along with the KLD loss of the latent space and the log(P(x|z)) value to determine the full ELBO loss value, which is returned, along with values for log(P(x|z)), the variational parameter loss and the latent variable loss.
#
# `likelihoods` takes in a particular `x` and returns the log(P(x)) estimation for that `x`, used in mutation effect prediction. This and subsequent functions are used in prediction and testing rather than training.
#
# `all_likelihood_components` takes in a particular `x` and returns the log(P(x)) estimation, the latent space KLD loss and the log(P(x|z)) value.
#
# `recognize` takes in a particular `x` and returns the mu and log-sigma of the latent variable outputted by the model's encoder.
#
# `get_pattern_activations` takes in a particular `x` and returns the output of the decoder's final layer from that x.
#
# ----
#
# ### pt_train.py
#
# Module containing the main training function, as well as loading and saving functions.
#
# #### save
#
# accepts a model and filepath amongst arguments, and saves the model.
#
# #### load
#
# Loads weights from a given path to a given model.
#
# #### train
#
# Takes in an instance of the DataHelper and VAE model, along with training parameters. Sets up saving file path, embeddings (if the use of embeddings is chosen in the DataHelper) and sets up the Adam solver with the model's parameters and a learning rate of 0.001). Generates training loop, whereby for each loop a mini-batch is selected (with calculated sequence weights used to select the weights to recalibrate against biases) and then, if required, the model and data are transferred to GPU with cuda. The Adam solver's gradients are reset (as is required with Torch, due to accumulating gradients), the forward pass is performed, loss is found, backpropagation is performed and the solver updates parameters. Updated values are saved, printed or stored as specified and this process repeats for each epoch.
#
# ### run_ptmle.py / run_ptsvi.py
#
# These are the scripts that should be called to run the model. In each, the above modules are imported; data, model and training parameters are specified in dictionaries, the DataHelper and relevant model are constructed. Parameters are printed to output, and then the model is trained - at the end, the model is saved. Due to the different model architectures, different scripts are used for the SVI and MLE versions of the model.
#
# ### SVI_mutation_analysis.py / MLE_mutation_analysis
#
# Example scripts for performing mutation effect prediction - largely based off code from the original project. Model is constructed and loaded with saved parameters, and a function is defined and called to calculate the Spearman R value for comparison of the model's predictions with experimental data. Greater agreement between the model's predictions and the experimental data will lead to larger values of Spearman R. Although the code for this analysis was largely written before this project, the code will be supplied for reference in [model performance](performance.html)
#
# ## Key Changes
#
# Here, we briefly highlight some of the main differences and developments that are present in the new model.
#
# - Increased Modularity. Through the use of torch's nn.Module class, the encoder, decoder and VAE elements of the models have been defined as separate classes. This makes the model cleaner, more readable and more adjustable - for example, both the SVI and MLE versions of the model inherit the Encoder class, meaning that adjustments are instantly shared between models and do not need to be repeated.
# - Increased concision. Through use of torch's machine learning functions and modules, the code is more concise - the full model takes up about half the number of lines of code. Again, this improves readability and manageability of the model which makes usage easier for new users
# - Less hard-coding. The original model was almost entirely defined in standard Python code, depending on Theano largely just for GPU compatibility. The new model takes advantage of PyTorch's many built in functionalities to define the model. Not only does this make the model more interpretable, but it makes hyper-parameter exploration and adjustment significantly easier. For example, to change the original model's optimizer from Adam to Adadelta, for example, would require adjustments to a minimum of 25 lines of code - now, this can be done by changing only one line.
# - Increased documentation. It is important for the development of this model that new members of the lab can easily learn how it is constructed and how it is used. To this end, the degree of commenting and documentation in the model has been increased so that throughout the model it is more clear what is being done.
# - More Pythonic structure. Without needing to use compiled code or symbolic tensors, model functions can be defined by standard Python functions rather than with the use of the compiled function call `theano.function()`.
# - A data-loading bug that was present in the original DataHelper script was fixed to ensure proper functionality. This bug prevented the loading of pre-prepared datasets.
|
notebooks/structure.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# # Detecting rooftop available surface for installing PV modules in aerial images using Machine Learning
# In this notebook we will present the entire pipeline to train a Unet model from a desired data set, evaluate the results and visualize the predictions. We present multiple ways to initialize and train a Unet. All the methods are availble under the section *Training Methods*. Note that in our results *Adaptative Training* provided the best results.
# +
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch.autograd import Variable
from torchvision.transforms.functional import normalize
from torch.utils.data import DataLoader
from torch.utils.data import DataLoader, ConcatDataset
from train.train import *
from tempfile import TemporaryFile
from process_data.normalize import *
from model.unet import *
from loss.loss import *
from process_data.data_loader import *
from hyperparameters.select_param import *
from process_data.import_test import *
from plots.plots import *
# %load_ext autoreload
# %autoreload 2
# -
device=torch.device('cuda' if torch.cuda.is_available() else 'cpu')
seed_torch() # For reproducibility we set the seed with a seed_torch() method that set the seed in numpy and pytorch
# # Loading the Data Set
# First we load the data set that we will use for training. Each sample is an image with its mask (label). An image is represented as a 3x250x250 array with each of the 3 color chanel being 250x250 pixels. The asssociated mask is a 250x250 array. Note that we already split the images in train/val/test 80/10/10 in advance to make our reproducibility as clear as possible.
#
# We perform data augmentation and transformation on the training set to counter the low amount of images in our data set. However in the validation set and test set, we only perform transformatio and no augmentation. Again this is to make reproducibility easier.
# +
folder_path_train_image = 'data/train/images'
folder_path_train_masks = 'data/train/labels'
folder_path_test_image = 'data/test/images'
folder_path_test_masks = 'data/test/labels'
folder_path_val_image = 'data/val/images'
folder_path_val_masks = 'data/val/labels'
folder_path_custom_image = 'data/custom/images'
# Load dataset
train_set = DataLoaderSegmentation(folder_path_train_image,folder_path_train_masks) # 80%
test_set = DataLoaderSegmentation(folder_path_test_image,folder_path_test_masks,augment=False)# 10%, no augmentation
val_set = DataLoaderSegmentation(folder_path_val_image,folder_path_val_masks,augment=False) # 10%, no augmentation
custom_set = DataLoaderSegmentation(folder_path_custom_image, augment=False)
# Init data loader
train_loader = DataLoader(train_set,batch_size=2, shuffle=True ,num_workers=0)
val_loader = DataLoader(val_set,batch_size=2, shuffle=True ,num_workers=0)
test_loader = DataLoader(test_set,batch_size=2 , shuffle=True ,num_workers=0)
custom_loader = DataLoader(custom_set,batch_size=2 , shuffle=True ,num_workers=0)
print(len(train_set),len(test_set),len(val_set),len(custom_set))
# +
model = UNet(3,1,False).to(device)
# + tags=[]
# We may compute the mean and standard deviation of the train loader. This is used either to check if the data loader is normalized, or to compute the mean and std for the normalizer in data_loader.
mean_std(train_loader)
# -
# # Training Methods
# We now present a certain number of training methods, they all initialize a new Unet model from scratch and train it. Note that this methods should not be used simultaneously. Instead you should choose one of the methods, run it and evaluate its performance.
#
# ## Regular training
# This is a simple training loop. We can tune the num_epochs, the learning rate and the parameter of the loss function.
# + tags=[]
# Init training parameters
num_epochs = 50
model = UNet(3,1,False).to(device)
loss_function = torch.nn.BCEWithLogitsLoss(weight=torch.FloatTensor([4]))#.cuda())
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
# Train model
history_train_loss, history_val_loss, history_train_iou, history_val_iou = training_model(train_loader,loss_function,optimizer,model,num_epochs)
# -
np.random.uniform(-0.1,0.1)
# Visualize the evolution of the loss and the IoU, either on the train or validation set.
plot_train_val(history_train_loss,history_val_loss,period=25, al_param=False, metric='loss')
plot_train_val(history_train_iou,history_val_iou,period=25, al_param=False, metric='IoU')
# ## Training with adaptative learning rate
# Training with adaptative learning rate is a regular training with an added learning rate scheduler. The task of the scheduler is to change the learning rate depending of the number of epochs. In our testing, the linear learning rate scheduler provided the best results.
#
#
# +
# Init training parameters
num_epochs = (150)
loss_function = torch.nn.BCEWithLogitsLoss(weight=torch.FloatTensor([15]).cuda())
optimizer = torch.optim.Adam(model.parameters(), lr=0.00008)
# We opted for the linear scheduler. For example, every 60 epochs the learning rate is multiplied by 0.8.
al_param=60
scheduler = torch.optim.lr_scheduler.StepLR(optimizer, al_param, gamma=0.8, last_epoch=-1, verbose=False)
# Train model
history_train_loss, history_val_loss, history_train_iou, history_val_iou = training_model(train_loader,loss_function,optimizer,model,num_epochs,scheduler,val_loader)
# -
# Visualize the evolution of the loss and the IoU, either on the train or validation set.
plot_train_val(history_train_loss,history_val_loss,period=25, al_param=al_param, metric='loss')
plot_train_val(history_train_iou,history_val_iou,period=25, al_param=al_param, metric='IoU')
# ## Adaptative Learning
# Performs a training on a model over a training data set by doing the following: we first fix the learing rate, then we split the training set into two folds, the model is trained on the first fold then on the second fold. After this has been done, we move on the next learning rate.
#
# Note that this method as no theoritical basis and came up from a flawed cross validation method we did not implemented correctly. However since it was providing good results we decided to clean it and keep it as a training method.
# +
# Init training parameters
lr_candidates = np.logspace(-1,-2,num=5)
num_epochs = 5
loss_function = torch.nn.BCEWithLogitsLoss(pos_weight=torch.FloatTensor([6]).cuda())
model = UNet(3,1,False).to(device)
# Train model
best_iou, history_iou = adptative_learning(train_set,val_loader,loss_function,best_model,num_epochs,lr_candidates)
# -
# # Miscellaneous methods
# We may try to identify the best learning rate for our task, either by using a regular grid search algorithm on the learning rate or performing cross validation to have a good estimate of the IoU we may get.
#
# ## Find Best Learning Rate
# We perform a grid search algorithm on the learning rate, by using a predefined range of learning rates lr and keeping the model that has the best IoU score.
# +
loss_function = torch.nn.BCEWithLogitsLoss(pos_weight=torch.FloatTensor([25]).cuda())
n_splits = 2
num_epochs = 300
lr = np.logspace(-1,-2,num=5)
best_lr, best_iou = select_hyper_param(train_loader,n_splits,loss_function,num_epochs,lr)
# -
# ## Cross Validation
# We can perform K-fold cross validation on our training set, to have an estimate of the mean iou and accuracy we may hope to achieve. Note that K-Fold CV is not considered as appropriate for CNN parameters tuning, since it is very costly.
# +
loss_function = torch.nn.BCEWithLogitsLoss(pos_weight=torch.FloatTensor([6]).cuda())
n_splits = 2
num_epochs = 10
lr = 0.01
iou_, acc_ = cross_validation(train_dataset=train_set,n_splits, loss_function, num_epochs, lr)
# -
# # Export or import a model
#
# ## Export a model
# To save a model trained with one of the aforementionned methods, one could use the following cell. The model is save in the *model* folder.
torch.save(model.state_dict(), 'model/'+input('Name of the model file:')+".pt")
# ## Import a model
# To import a model, one could use the following cell. The model must be located in the *model* folder. In the command prompt you can for example write *all/batch5loss4_200*
# path = 'models/all/'+input('Name of the model file:')+".pt"
path = 'models/all/batch5_loss4_1000.pt'
model = UNet(3,1,False).to(device)
# model.load_state_dict(torch.load(path))
model.load_state_dict(torch.load(path, map_location=torch.device('cpu')))
# # Evaluation of the model
# We can evaluate the model to have the mean (IoU, Accuracy) on every data set, and print the number of parameters of the Unet.
print('Train:', test_model(train_loader,model))
print('Val:', test_model(val_loader,model))
print('Test:', test_model(test_loader,model))
sum(p.numel() for p in model.parameters())
# # Visualization of the model
# With the model we trained or imported, we can display from the test_loader examples of its prediction.
# + tags=[]
# Get the input, transformed input and prediction made by the model
model.eval()
index_random_sample = int(np.random.random()*len(test_loader.dataset))
(x,y,z) = test_loader.dataset.__getitem__(index_random_sample,show_og=True)
ypred = torch.squeeze(model.predict(torch.unsqueeze(x,0))).cpu().detach().numpy()
# Display all 4 images
fig = plt.figure()
fig.set_size_inches(12, 7, forward=True)
ax1 = fig.add_subplot(1,4,1)
ax1.title.set_text('Input Image')
ax2 = fig.add_subplot(1,4,2)
ax2.title.set_text('Transformed Input Image')
ax3 = fig.add_subplot(1,4,3)
ax3.title.set_text('Expected Mask')
ax4 = fig.add_subplot(1,4,4)
ax4.title.set_text('Predicted Mask')
ax1.imshow(z)
ax2.imshow(np.transpose(x.numpy(),(1,2,0)))
ax3.imshow(y)
ax4.imshow(np.around((ypred)))
# Compute IoU and accuracy on prediction and mask
predict_iou = np.around(iou(np.around(ypred),y.numpy()),4)
predict_acc = accuracy(np.around(ypred),y.numpy())
print('Iou:',predict_iou)
print('Accuracy:', predict_acc)
# +
# Get the input, transformed input and prediction made by the model
model.eval()
index_random_sample = int(np.random.random()*len(custom_loader.dataset))
(x,y,z) = custom_loader.dataset.__getitem__(index_random_sample,show_og=True)
ypred = torch.squeeze(model.predict(torch.unsqueeze(x,0))).cpu().detach().numpy()
# Display all 4 images
fig = plt.figure()
fig.set_size_inches(12, 7, forward=True)
ax1 = fig.add_subplot(1,4,1)
ax1.title.set_text('Input Image')
ax2 = fig.add_subplot(1,4,2)
ax2.title.set_text('Transformed Input Image')
ax3 = fig.add_subplot(1,4,3)
ax3.title.set_text('Expected Mask')
ax4 = fig.add_subplot(1,4,4)
ax4.title.set_text('Predicted Mask')
ax1.imshow(z)
ax2.imshow(np.transpose(x.numpy(),(1,2,0)))
ax3.imshow(y)
ax4.imshow(np.around((ypred)))
# Compute IoU and accuracy on prediction and mask
predict_iou = np.around(iou(np.around(ypred),y.numpy()),4)
predict_acc = accuracy(np.around(ypred),y.numpy())
print('Iou:',predict_iou)
print('Accuracy:', predict_acc)
# -
|
rooftop-detection/main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Space Flight Mice Kidney Gene Analysis
# ## network_bio_toolkit all 3 pipelines
#
# ----------------------
#
# Author: <NAME> (<EMAIL>)
#
# Date: 21st June, 2018
#
# ----------------------
#
# <a id='toc'></a>
# ## Table of Contents
# 1. [Background](#background)
# 2. [Import packages](#import)
# 3. [Upstream Regulator Analysis](#ura)
# 1. [Load Networks](#loadura)
# 2. [Localization](#local)
# 3. [TF Enrichment](#enrich)
# 4. [TF Activation State Prediction](#active)
# 5. [Display Our results](#display)
# 4. [Heat Propagation and Clustering Analysis](#hpc)
# 1. [Load Networks](#loadhpc)
# 2. [Run Heat Propagation](#heat)
# 3. [Clustering](#cluster)
# 5. [Gene Set Enrichment Analysis](#gsea)
# 1. [User Preferences for Metafile Filtering](#meta)
# 2. [Filter Expression File](#expression)
# 3. [GSEA call](#gseacall)
# 4. [Visualization Functions](#vis)
# ## Background
# <a id='background'></a>
#
# Network biology is quickly becoming an indispensable tool for elucidating dysregulated biological pathways and furthering understanding in disease. However, network analysis is intimidating to many researchers. We present an integrated network analysis toolkit, network_bio_toolkit, built upon our previous interactive network visualization tool, visJS2jupyter, to provide three common network analysis workflows for RNAseq data, from start to finish, requiring only an expression file, or a list of differentially expressed genes (DEGs). The tool is designed for use in Jupyter notebooks, to promote reproducible research.
#
# This package includes a set of start-to-finish functions for three common network analysis pipelines: upstream regulator analysis, network propagation and clustering, and gene set enrichment analysis. The upstream regulator analysis pipeline includes localization, transcription factor (TF) enrichment, and TF activation state prediction methods. The network propagation and clustering pipeline includes network propagation, clustering, and annotation functionality. The gene set enrichment analysis includes easy-to-use data filtering methods, that help the user prep their data as input for GSEA’s enrichment calculation function. Each pipeline includes functions to easily load and process a variety of genome networks, such as STRING and GIANT, from NDEXbio, eliminating the need for downloading large and unwieldy network files. Each pipeline also has data visualization functions, so interpreting results from our pipelines is intuitive. Many of the analysis components of each pipeline are modular; the user can pick and choose which functions are relevant to their data and leave out unnecessary steps.
#
# In this notebook, we demonstrate the functionality of network_bio_toolkit by analyzing a set of differentially expressed genes derived from NASA's [Rodent Research Project on the ISS](https://genelab-data.ndc.nasa.gov/genelab/accession/GLDS-102). The tissues sampled for this experiment were taken from the kidneys of mice who experienced space flight. The data and files used in this analysis are the results after RNAseq alignment and quantification using STAR+RSEM, and differential expression analysis using Limma-voom, on the NASA kidney data.
#
# This
# ## Import packages
# <a id='import'></a>
# +
# import upstream regulator module
import sys
code_path = '../../network_bio_toolkit'
sys.path.append(code_path)
import Upstream
reload(Upstream)
import Heat2
reload(Heat2)
import PrepGsea
reload(PrepGsea)
# -
# ## Upstream Regulator Analysis
# <a id='ura'></a>
#
# The inspiration for these modules comes from Ingenuity System's [Ingenuity Upstream Regulator Analysis in IPA®](http://pages.ingenuity.com/rs/ingenuity/images/0812%20upstream_regulator_analysis_whitepaper.pdf).
# +
# User preferences
symbol = 'symbol'
entrez = 'entrez'
human = 'human'
mouse = 'mouse'
ura = Upstream.Upstream(gene_type = symbol, species = mouse)
# -
# ### Load Networks
# <a id='loadura'></a>
#
# 1. List of all **Transcription Factors** (TF's): [slowkow](https://github.com/slowkow/tftargets) and [jaspar](http://jaspar.genereg.net/)
# 2. **Background Network**: [STRING human protein interactions network](https://string-db.org/cgi/download.pl?)
# 3. User-supplied list of **Differentially Expressed Genes** (DEG's)
#
# Note: We need to use the String protein ACTIONS file because it has activating and inhibiting interaction information.
# transcription factors
ura.easy_load_TF_list('../../TF_databases/TF_database_URA.csv')
TF_list = ura.get('TF_list')
print "Number of TF's: " + str(len(TF_list))
# +
# background network
filename = "../../background_networks/10090.protein.actions.v10.5.txt"
confidence_filter = 400
ura.load_STRING_to_digraph(filename, confidence_filter)
DG_TF = ura.get('DG_TF')
print "\nNumber of interactions: " + str(len(list(DG_TF.edges())))
# +
# differentially expressed genes
DEG_filename = "../../DEG_databases/DE_SFvsGC_12_Kidney.csv"
ura.create_DEG_list(DEG_filename, p_value_filter = 0.05, sep = ',')
DEG_list = ura.get('DEG_list')
print "Number of DEG's: " + str(len(DEG_list))
# -
# ### Localization
# <a id='local'></a>
#
# Below we see that the set of DEGs is visibly more localized when compared to randomly selected gene sets with similar degree distributions. In the first 2 figures, the blue distributions are formed from randomly sampling 80% of the genes from the set of DEGs. In the third figure, the red line is the absolute size of the largest connected component of the DEGs. The blue distributions and red line are visibly distinct from the randomly sampled sets.
numedges_list, numedges_rand, LCC_list, LCC_rand = ura.localization(num_reps = 100,
sample_frac = 0.8,
method = 'both',
plot = True)
# The p-value of zero indicates extreme significance.
# test out plotting parameters with a smaller num_reps first.
numedges_list, numedges_rand, LCC_list, LCC_rand = ura.localization_full(num_reps = 100,
method = 'LCC',
label = 'focal genes',
line_height = 1,
legend_loc = 'upper left')
# ## TF Enrichment
# <a id='enrich'></a>
# How relevant is a TF to its DEG targets? Are they connected by chance, or is their connection statistically significant? Are the TFs themselves enriched? We calculate the -log(p-value) using [scipy.stats.hypergeom.logsf](https://docs.scipy.org/doc/scipy-0.19.1/reference/generated/scipy.stats.hypergeom.html) for every TF-plus-downstream-targets network, and then for the group of TFs themselves.
# - high value = significant connection between this TF and its DEG targets
# - low value = TF is randomly associated with its DEG targets
# - zero = None of this TF's targets were DEG's
# - inf = original p-value was so small that its log is inf. Very high significance.
#
# Below, we see that the TF Zbtb4 and its downstream targets are significnantly enriched, however the set of TFs themselves are not.
# Enrichment of every TF with respect to its targets
ura.tf_target_enrichment_calc()
ura.get('tf_target_enrichment').head()
# Enrichment of TF's themselves
ura.tf_enrichment_calc()
ura.get('tf_enrichment')
# ## TF Activation State Prediction
# <a id='active'></a>
#
# Is a given TF activating or inhibiting to its downstream targets? We quantify the activation state of the TF using a z-score calculation. A positive z-score indicates activating while a negative score indicates inhibiting. The significance of the z-score is a vote for how confident the algorithm is of that TF's state prediction.
#
# Below we display the top 5 most significant activating and top 5 inhibiting genes.
ura.tf_zscore()
display(ura.top_values(act = True, top = 5))
display(ura.top_values(act = False, top = 5))
# ### Display Our results
# <a id='display'></a>
# Here we plot those 10 genes along the full distribution of activation state z-scores. They are visibly the most significant TFs in the distribution.
genes_to_rank = ['Ikbkb', 'Btrc', 'Fbxw11', 'Bhlhe41','Per2', 'Per1', 'Akt1', 'Myc', 'Anapc2']
ura.compare_genes(genes_to_rank, font_size = 14)
# Per2 was predicted to be inhibiting to its downstream targets. The network below shows that Per2 has 3 DEG targets, where each of those targets is significantly up-regulated (bright red node color indicates up regulation). Per2's interaction with each of those up-regulated DEGs is inhibiting (as indicated by the blue edges between Per2 and the 3 red nodes). The following chart is represents our activation state prediction algorithm:
#
# (interaction type/regulation direction = predicted state)
# - activating/up = activating
# - activating/down = inhibiting
# - **inhibiting/up = inhibiting**
# - inhibiting/down = activating
#
# According to our algorithm, Per2 should be strongly inhibiting because all of the DEGs uninanimously agree (by being up-regulated and having an inhibiting interaction with Per2) that it is.
ura.vis_tf_network('Per2',
directed_edges = True,
color_non_DEGs = False,
node_spacing = 1100,
graph_id = 1, # needs to be different for every graph in the notebook
tf_size_amplifier = 5)
# ## Heat Propagation and Clustering Analysis
# <a id='hpc'></a>
#
# The Heat2 module provides tools to conduct an integrated network analysis of a set of differentially expressed genes (DEGs).
heat = Heat2.Heat(gene_type = symbol, species = mouse)
# ### Load Networks
# <a id='loadhpc'></a>
# +
filename = '../../background_networks/10090.protein.links.v10.5.txt'
heat.load_STRING_links(filename, confidence_filter = 700)
print('\nNumber of interactions: ' + str(len(list(heat.DG_universe.edges()))))
# +
# using the same DEG file as with URA
heat.create_DEG_list(DEG_filename, p_value_filter = 0.05, sep = ',')
print('Number of DEG\'s: ' + str(len(heat.DEG_list)))
# -
# ### Run Heat Propagation
# <a id='heat'></a>
#
# Below is a visualization of our heat propagation. Here we are displaying roughly the hottest 200 nodes, where triangle nodes are DEGs. The redder the node, the hotter it is.
heat.normalized_adj_matrix()
heat.draw_heat_prop(num_nodes = 200,
random_walk = True,
edge_width = 2,
edge_smooth_enabled = True,
edge_smooth_type = 'bezier',
node_size_multiplier = 5,
hover = False,
hover_connected_edges = False,
largest_connected_component = True,
physics_enabled = True,
node_font_size = 40,
graph_id = 2, # needs to be different for every graph in the notebook
node_shadow_x = 6)
# ### Clustering
# <a id='cluster'></a>
# Now we take those same 200 nodes displayed above, and we run a cluster analysis on them. Each cluster has been assigned a different color. If you hover your mouse over a node, you will see its assigned cluster id, or you can check out the cluster legend below our network.
heat.draw_clustering(rad_positions = False,
k = None,
largest_connected_component = True,
num_top_genes = 200,
cluster_size_cut_off = 0,
remove_stray_nodes = True,
node_spacing = 700,
node_size_multiplier = 10,
physics_enabled = True,
node_font_size = 40,
graph_id = 3, # needs to be different for every graph in the notebook
edge_width = 2,
edge_smooth_enabled = True,
edge_smooth_type = 'bezier',
hover = False,
hover_connected_edges = False
)
heat.cluster_legend()
# Now we spacially separate the clusters, so you can see them better.
heat.draw_clustering(rad_positions = True,
k = None,
r = 0.5,
largest_connected_component = True,
num_top_genes = 200,
cluster_size_cut_off = 0,
remove_stray_nodes = True,
node_spacing = 1000,
node_size_multiplier = 5,
physics_enabled = False,
node_font_size = 20,
graph_id = 4, # needs to be different for every graph in the notebook
edge_width = 1,
edge_smooth_enabled = True,
edge_smooth_type = 'bezier',
hover = False,
hover_connected_edges = False
)
# Lastly, rather than color by cluster, we color by log fold change. Blue nodes have negative log fold change values and red nodes have positive ones.
heat.draw_clustering(rad_positions = True,
k = None,
r = 0.5,
num_top_genes = 200,
cluster_size_cut_off = 0,
remove_stray_nodes = True,
node_spacing = 1000,
node_size_multiplier = 5,
physics_enabled = False,
node_font_size = 20,
graph_id = 5, # needs to be different for every graph in the notebook
edge_width = 1,
edge_smooth_enabled = True,
edge_smooth_type = 'bezier',
hover = False,
hover_connected_edges = False,
color_lfc = True,
largest_connected_component = True,
vmin = -1,
vmax = 1
)
min_lfc = min(zip(*heat.get('node_to_lfc').items())[1])
max_lfc = max(zip(*heat.get('node_to_lfc').items())[1])
heat.draw_legend(label = 'log fold change')
# ## Gene Set Enrichment Analysis
# <a id='gsea'></a>
gsea_prepper = PrepGsea.PrepGsea(gmt_file = '../../gseapy_databases/h.all.v6.1.symbols.gmt',
expression_file = '../../gseapy_databases/cpm_12_Kidney_symbol.tsv',
meta_file = '../../gseapy_databases/NASA_RR1kidney_metadata.csv',
output_dir = '../../gsea_output')
# ### User Preferences for Metafile Filtering
# <a id='meta'></a>
gsea_prepper.remove_extra_columns()
gsea_prepper.choose_comparison_col('Sample_name')
gsea_prepper.choose_two_classes()
gsea_prepper.choose_controls()
gsea_prepper.filter_metafile()
# ### Filter Expression File
# <a id='expression'></a>
gsea_prepper.filter_expression_file()
# ### GSEA call
# <a id='gseacall'></a>
gsea_prepper.call_gsea(method = 'log2_ratio_of_classes',
processes = 4,
format = 'png',
permutation_num = 10,
weighted_score_type = 1)
# ### Visualization Functions
# <a id='vis'></a>
gsea_prepper.plot_gsea(style_content = 'ggplot', top = 20, y = 'fdr', x = 'Term', fontsize = 8)
# We are currently working on fixing the text location in the visualization.
gsea_prepper.plot_individual_pathway_heatmap('HALLMARK_G2M_CHECKPOINT')
|
notebooks/general_notebooks/all_pipelines_mice_kidneys.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Tutorial 7: Pancan data access
#
# This tutorial shows how to use the `cptac.pancan` submodule to access data from the harmonized pipelines for all cancer types.
#
# Before the harmonized pipelines, the team working on each cancer type had their own pipeline for each data type. So for example, the ccRCC team ran the ccRCC data through their own transcriptomics pipeline, and the HNSCC team ran the HNSCC data through a different transcriptomics pipeline. However, this made it hard to study trends across multiple cancer types, since each cancer type's data had been processed differently.
#
# To fix this problem, all data for all cancer types was run through the same pipelines for each data type. These are the harmonized pipelines. Now, for example, you can get transcriptomics data for both ccRCC and HNSCC (and all other cancer types) that came from the same pipeline.
#
# For some data types, multiple harmonized pipelines were available. In this cases, all cancers were run through each pipeline, and you can choose which one to use. For example, you can get transcriptomics data from either the BCM pipeline, the Broad pipeline, or the WashU pipeline. But whichever pipeline you choose, you can get transcriptomics data for all cancer types through that one pipeline.
#
# First, we'll import the package.
import cptac.pancan as pc
# We can list which cancers we have data for.
pc.list_datasets()
# ## Download
#
# Authentication through your Box account is required when you download files. Pass the name of the dataset you want, as listed by `list_datasets`. Capitalization does not matter.
#
# See the end of this tutorial for how to download files on a remote computer that doesn't have a web browser for logging into Box.
pc.download("pancanbrca")
# ## Load the BRCA dataset
br = pc.PancanBrca()
# We can list which data types are available from which sources.
br.list_data_sources()
# Let's get some data tables.
br.get_clinical(source="mssm")
br.get_somatic_mutation(source="washu")
br.get_proteomics(source="umich")
# ## Box authentication for remote downloads
#
# Normally, when you download the `cptac.pancan` data files you're required to log into your Box account, as these files are not released publicly. However, there may be situations where the computer you're running your analysis on doesn't have a web browser you can use to log in to Box. For example, you may be running your code in a remotely hosted notebook (e.g. Google Colabs), or on a computer cluster that you access using ssh.
#
# In these situations, follow these steps to take care of Box authenication:
# 1. On a computer where you do have access to a web browser to log in to Box, load the `cptac.pancan` module.
# 2. Call the `cptac.pancan.get_box_token` function. This will return a temporary access token that gives permission to download files from Box with your credentials. The token expires 1 hour after it's created.
# 3. On the remote computer, when you call the `cptac.pancan.download` function, copy and paste the access token you generated on your local machine into the `box_token` parameter of the function. The program will then be able to download the data files.
#
# Below is all the code you would need to call for this process on each machine. For security, we will not actually run it in this notebook.
#
# On your local machine:
# ```
# import cptac.pancan as pc
# pc.get_box_token()
# ```
#
# On the remote machine:
# ```
# import cptac.pancan as pc
# pc.download("pancanbrca", box_token=[INSERT TOKEN HERE])
# ```
|
notebooks/tutorial07_pancan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="3o8Qof7Cy165"
# # LAB 2b: Prepare babyweight dataset.
#
# **Learning Objectives**
#
# 1. Setup up the environment
# 1. Preprocess natality dataset
# 1. Augment natality dataset
# 1. Create the train and eval tables in BigQuery
# 1. Export data from BigQuery to GCS in CSV format
#
#
# ## Introduction
# In this notebook, we will prepare the babyweight dataset for model development and training to predict the weight of a baby before it is born. We will use BigQuery to perform data augmentation and preprocessing which will be used for AutoML Tables, BigQuery ML, and Keras models trained on Cloud AI Platform.
#
# In this lab, we will set up the environment, create the project dataset, preprocess and augment natality dataset, create the train and eval tables in BigQuery, and export data from BigQuery to GCS in CSV format.
#
# Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/1b_prepare_data_babyweight.ipynb).
# + [markdown] colab_type="text" id="hJ7ByvoXzpVI"
# ## Set up environment variables and load necessary libraries
# + [markdown] colab_type="text" id="mC9K9Dpx1ztf"
# Check that the Google BigQuery library is installed and if not, install it.
# + colab={"base_uri": "https://localhost:8080/", "height": 609} colab_type="code" id="RZUQtASG10xO" outputId="5612d6b0-9730-476a-a28f-8fdc14f4ecde" language="bash"
# pip freeze | grep google-cloud-bigquery==1.6.1 || \
# pip install google-cloud-bigquery==1.6.1
# -
# Import necessary libraries.
import os
from google.cloud import bigquery
# ## Lab Task #1: Set environment variables.
#
# Set environment variables so that we can use them throughout the entire lab. We will be using our project name for our bucket, so you only need to change your project and region.
# + language="bash"
# export PROJECT=$(gcloud config list project --format "value(core.project)")
# echo "Your current GCP Project Name is: "$PROJECT
# +
# TODO: Change environment variables
PROJECT = "cloud-training-demos" # REPLACE WITH YOUR PROJECT NAME
REGION = "us-central1" # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
# Do not change these
os.environ["BUCKET"] = PROJECT # DEFAULT BUCKET WILL BE PROJECT ID
os.environ["REGION"] = REGION
if PROJECT == "cloud-training-demos":
print("Don't forget to update your PROJECT name! Currently:", PROJECT)
# + [markdown] colab_type="text" id="L0-vOB4y2BJM"
# ## The source dataset
#
# Our dataset is hosted in [BigQuery](https://cloud.google.com/bigquery/). The CDC's Natality data has details on US births from 1969 to 2008 and is a publically available dataset, meaning anyone with a GCP account has access. Click [here](https://console.cloud.google.com/bigquery?project=bigquery-public-data&p=publicdata&d=samples&t=natality&page=table) to access the dataset.
#
# The natality dataset is relatively large at almost 138 million rows and 31 columns, but simple to understand. `weight_pounds` is the target, the continuous value we’ll train a model to predict.
# -
# ## Create a BigQuery Dataset and Google Cloud Storage Bucket
#
# A BigQuery dataset is a container for tables, views, and models built with BigQuery ML. Let's create one called __babyweight__ if we have not already done so in an earlier lab. We'll do the same for a GCS bucket for our project too.
# + language="bash"
#
# ## Create a BigQuery dataset for babyweight if it doesn't exist
# datasetexists=$(bq ls -d | grep -w # TODO: Add dataset name)
#
# if [ -n "$datasetexists" ]; then
# echo -e "BigQuery dataset already exists, let's not recreate it."
#
# else
# echo "Creating BigQuery dataset titled: babyweight"
#
# bq --location=US mk --dataset \
# --description "Babyweight" \
# $PROJECT:# TODO: Add dataset name
# echo "Here are your current datasets:"
# bq ls
# fi
#
# ## Create GCS bucket if it doesn't exist already...
# exists=$(gsutil ls -d | grep -w gs://${BUCKET}/)
#
# if [ -n "$exists" ]; then
# echo -e "Bucket exists, let's not recreate it."
#
# else
# echo "Creating a new GCS bucket."
# gsutil mb -l ${REGION} gs://${BUCKET}
# echo "Here are your current buckets:"
# gsutil ls
# fi
# + [markdown] colab_type="text" id="b2TuS1s9vREL"
# ## Create the training and evaluation data tables
#
# Since there is already a publicly available dataset, we can simply create the training and evaluation data tables using this raw input data. First we are going to create a subset of the data limiting our columns to `weight_pounds`, `is_male`, `mother_age`, `plurality`, and `gestation_weeks` as well as some simple filtering and a column to hash on for repeatable splitting.
#
# * Note: The dataset in the create table code below is the one created previously, e.g. "babyweight".
# -
# ### Lab Task #2: Preprocess and filter dataset
#
# We have some preprocessing and filtering we would like to do to get our data in the right format for training.
#
# Preprocessing:
# * Cast `is_male` from `BOOL` to `STRING`
# * Cast `plurality` from `INTEGER` to `STRING` where `[1, 2, 3, 4, 5]` becomes `["Single(1)", "Twins(2)", "Triplets(3)", "Quadruplets(4)", "Quintuplets(5)"]`
# * Add `hashcolumn` hashing on `year` and `month`
#
# Filtering:
# * Only want data for years later than `2000`
# * Only want baby weights greater than `0`
# * Only want mothers whose age is greater than `0`
# * Only want plurality to be greater than `0`
# * Only want the number of weeks of gestation to be greater than `0`
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_data AS
SELECT
# TODO: Add selected raw features and preprocessed features
FROM
publicdata.samples.natality
WHERE
# TODO: Add filters
# ### Lab Task #3: Augment dataset to simulate missing data
#
# Now we want to augment our dataset with our simulated babyweight data by setting all gender information to `Unknown` and setting plurality of all non-single births to `Multiple(2+)`.
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_augmented_data AS
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks,
hashmonth
FROM
babyweight.babyweight_data
UNION ALL
SELECT
# TODO: Replace is_male and plurality as indicated above
FROM
babyweight.babyweight_data
# ### Lab Task #4: Split augmented dataset into train and eval sets
#
# Using `hashmonth`, apply a modulo to get approximately a 75/25 train/eval split.
# ### Split augmented dataset into train dataset
# #### **Exercise**: **RUN** the query to create the training data table.
# + colab={} colab_type="code" id="CMNRractvREL"
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_data_train AS
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks
FROM
babyweight.babyweight_augmented_data
WHERE
# TODO: Modulo hashmonth to be approximately 75% of the data
# -
# ### Split augmented dataset into eval dataset
# #### **Exercise**: **RUN** the query to create the evaluation data table.
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_data_eval AS
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks
FROM
babyweight.babyweight_augmented_data
WHERE
# TODO: Modulo hashmonth to be approximately 25% of the data
# + [markdown] colab_type="text" id="clnaaqQsXkwC"
# ## Verify table creation
#
# Verify that you created the dataset and training data table.
#
# -
# %%bigquery
-- LIMIT 0 is a free query; this allows us to check that the table exists.
SELECT * FROM babyweight.babyweight_data_train
LIMIT 0
# %%bigquery
-- LIMIT 0 is a free query; this allows us to check that the table exists.
SELECT * FROM babyweight.babyweight_data_eval
LIMIT 0
# ## Lab Task #5: Export from BigQuery to CSVs in GCS
#
# Use BigQuery Python API to export our train and eval tables to Google Cloud Storage in the CSV format to be used later for TensorFlow/Keras training. We'll want to use the dataset we've been using above as well as repeat the process for both training and evaluation data.
# +
# Construct a BigQuery client object.
client = bigquery.Client()
dataset_name = # TODO: Add dataset name
# Create dataset reference object
dataset_ref = client.dataset(
dataset_id=dataset_name, project=client.project)
# Export both train and eval tables
for step in [# TODO: Loop over train and eval]:
destination_uri = os.path.join(
"gs://", BUCKET, dataset_name, "data", "{}*.csv".format(step))
table_name = "babyweight_data_{}".format(step)
table_ref = dataset_ref.table(table_name)
extract_job = client.extract_table(
table_ref,
destination_uri,
# Location must match that of the source table.
location="US",
) # API request
extract_job.result() # Waits for job to complete.
print("Exported {}:{}.{} to {}".format(
client.project, dataset_name, table_name, destination_uri))
# -
# ## Verify CSV creation
#
# Verify that we correctly created the CSV files in our bucket.
# + language="bash"
# gsutil ls gs://${BUCKET}/babyweight/data/*.csv
# + language="bash"
# gsutil cat gs://${BUCKET}/babyweight/data/train000000000000.csv | head -5
# + language="bash"
# gsutil cat gs://${BUCKET}/babyweight/data/eval000000000000.csv | head -5
# -
# ## Lab Summary:
# In this lab, we setup our environment, created a BigQuery dataset, preprocessed and augmented the natality dataset, created train and eval tables in BigQuery, and exported data from BigQuery to GCS in CSV format.
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
|
courses/machine_learning/deepdive2/structured/labs/1b_prepare_data_babyweight.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from simulation import inverse_kinematics_objective, draw_solutions
from idea import IDEA
# +
rectangles = [((0.4, 2.), (5., 4.)), ((-5., 2.), (-0.8, 4.)), ((-0.5, 2.), (0.1, 4.)),
((-5., 5.), (-0.3, 6.)), ((-0.1, 5.), (5., 6.))]
target_x = 1.
target_y = 8.
S = [0.7] * 10 + [0.3] * 10
d = len(S)
x_min = -np.pi
x_max = np.pi
objective = inverse_kinematics_objective(S, (target_x, target_y), rectangles)
n_constraints = len(rectangles)
n = 600
alpha_inf = 0.7
eta_c = 3.
eta_m = 20.
p_c = 0.9
p_m = 0.05
num_iterations = 300
populations, scores = IDEA(objective, n_constraints, x_min, x_max, d, n, alpha_inf, eta_c, eta_m, p_c, p_m, num_iterations, log_interval=20)
# -
mask = scores[-1][:, 1] == 0.
feasible_population = populations[-1][mask, :]
draw_solutions(feasible_population, S, (target_x, target_y), rectangles, figsize=(10, 8))
infeasible_population = populations[-1][~mask, :]
draw_solutions(infeasible_population, S, (target_x, target_y), rectangles, figsize=(10, 8))
|
First_experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + init_cell=true
# %logstop
# %logstart -rtq ~/.logs/ML_Natural_Language_Processing.py append
# %matplotlib inline
import matplotlib
import seaborn as sns
sns.set()
matplotlib.rcParams['figure.dpi'] = 144
# -
# # Natural Language Processing
#
# Natural language processing (NLP) is the field devoted to methods and algorithms for processing human (natural) languages for computers. NLP is a vast discipline that is actively being researched. For this notebook, we will be concerned with NLP tools and techniques we can use for machine learning applications. Some examples of machine learning applications using NLP include sentiment analysis, topic modeling, and language translation. In NLP, the following terms have specific meanings:
#
# * **Corpus**: The body/collection of text being investigated.
# * **Document**: The unit of analysis, what is considered a single observation.
#
# Examples of corpora include a collection of reviews and tweets, the text of the _Iliad_, and Wikipedia articles. Documents can be whatever you decided, it is what your model will consider an observation. For the example when the corpus is a collection of reviews or tweets, it is logical to make the document a single review or tweet. For the example of the text of the _Iliad_, we can set the document size to a sentence or a paragraph. The choice of document size will be influenced by the size of our corpus. If it is large, it may make sense to call each paragraph a document. As is usually the case, some design choices that need to be made.
#
# For this notebook, we will build a classifier to discern homonyms, words that are spelled the same but that have different meanings. The exact use case we will explore is to discern if the word "python" refers to the programming language or the animal.
# ## NLP with spaCy
#
# spaCy is a Python package that bills itself as "industrial-strength" natural language processing. We will use the tools spaCy provides in conjunction with `scikit-learn`. Let's explore some of spaCy's capabilities; we will introduce more functionality when needed. More about spaCy can be found [here](https://spacy.io/).
# +
import spacy
# load text processing pipeline
nlp = spacy.load('en')
# nlp accepts a string
doc = nlp("Let's try out spacy. We can easily divide our text into sentences! I've run out of ideas.")
# iterate through each sentence
for sent in doc.sents:
print(sent)
# index words
print(doc[0])
print(doc[6])
# -
# Another nice feature from spaCy is part-of-speech tagging, the process of identifying whether a word is a noun, adjective, adverb, etc. A processed word has the attribute `pos_` and `tag_`; the former identifies the simple part of speech (e.g., noun) wile the latter identifies the more detailed part of speech (e.g., proper noun). The meaning of the resulting abbreviations of the `tag_` are listed [here](https://www.ling.upenn.edu/courses/Fall_2003/ling001/penn_treebank_pos.html) or can be revealed by running `spacy.explain` function.
# +
doc = nlp("The quick brown fox jumped over the lazy dog. Mr. Peanut wears a top hat.")
tags = set()
# reveal part of speech
for word in doc:
tags.add(word.tag_)
print((word.text, word.pos_, word.tag_))
# revealing meaning of tags
print()
for tag in tags:
print(tag, spacy.explain(tag))
# -
# ## Obtaining a corpus
#
# Before we can move on with our analysis, we need to obtain a corpus. For our intended classifier, we need documents pertaining to python the animal and Python the programming language. Let's use Wikipedia articles to form our corpus. Luckily, there's a Python package called `wikipedia` that makes it easy to fetch articles. We will create documents based on the sentences in the articles. The function allows us to pass multiples pages in constructing the documents, allowing us to prevent one class of documents from dominating the corpus.
# +
import wikipedia
def pages_to_sentences(*pages):
"""Return a list of sentences in Wikipedia articles."""
sentences = []
for page in pages:
p = wikipedia.page(page)
doc = nlp(p.content)
sentences += [sent.text for sent in doc.sents]
return sentences
animal_sents = pages_to_sentences("Reticulated python", "Ball Python")
language_sents = pages_to_sentences("Python (programming language)")
documents = animal_sents + language_sents
print(language_sents[:5])
print()
print(animal_sents[:5])
# -
# **Question**
# * Given the example documents, what patterns should our word usage classifier learn?
# * We chose to create documents from sentences. What are other options? What are some pros and cons?
# ## Bag of words model
#
# Machine learning models needs to ingest data in a structured form, a matrix where the rows represents observations and the columns are features/attributes. When working with text data, we need a method to convert this unstructured data into a form that the machine learning model can work with. Let's consider our motivating example to create a classifier to discern the usage of "python" in a document. We understand that documents referring to the programming language will use words such as "integer", "byte", and "error" at higher frequency than documents that refer to python the animal. The reverse is true for words such as "bite", "snake", and "pet". One technique to _transform_ text data into a matrix is to count the number of appearances of each word in each document. This technique is called the **bag of words** model. The model gets its name because each document is viewed as a bag holding all the words, disregarding word order, context, and grammar. After applying the bag of words model to a corpus, the resulting matrix will exhibit patterns that a machine learning model can exploit. See the example below for the result of applying the bag of words model to a corpus of two documents.
# Document 0: "The python is a large snake, although the snake is not venomous." <br>
# Document 1: "Python is an interpreted programming language for general purpose programming." <br>
# <br>
#
# | although | an | for | general | interpreted | is | language | large | not | programming | purpose | python | snake | the | venomous |
# |:--------:|----|-----|---------|-------------|----|----------|-------|-----|-------------|---------|--------|-------|-----|----------|
# | 1 | 0 | 0 | 0 | 0 | 2 | 0 | 1 | 1 | 0 | 0 | 1 | 2 | 2 | 1 |
# | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 2 | 1 | 1 | 0 | 0 | 0 |
#
# ### The `CountVectorizer` transformer
#
# The bag of words model is found in `scikit-learn` with the `CountVectorizer` transformer. Note, `scikit-learn` uses the word `Vectorizer` to refer to transformers that convert a data structure (like a dictionary) into a NumPy array. Since it is a transformer, we need to first fit the object and _then_ call `transform`.
# +
from sklearn.feature_extraction.text import CountVectorizer
bag_of_words = CountVectorizer()
bag_of_words.fit(documents)
word_counts = bag_of_words.transform(documents)
print(word_counts)
word_counts
# -
# The `transform` method returns a sparse matrix. A sparse matrix is a more efficient manner of storing a matrix. If a matrix has mostly zero entries, it is better to just store the non-zero entries and their occurrence, their row and column. Sparse matrices have the method `toarray()` that returns a full matrix **but** doing so may result in memory issues. Some key hyperparameters of the `CountVectorizer` are shown below:
#
# * `min_df`: only counts words that appear in a minimum number of documents.
# * `max_df`: only counts words that do not appear more than a maximum number of documents.
# * `max_features`: limits the number of generated features, based on the frequency.
#
# After fitting a `CountVectorizer` object, the following method and attribute help with determining which index belongs to which word.
#
# * `get_feature_names()`: Returns a list of words used as features. The index of the word corresponds to the column index.
# * `vocabulary_`: A dictionary mapping a word to its corresponding feature index.
#
# Let's use `vocabulary_` to determine how many times "programming" occurs in the documents for Python the programming language and python the animal. Do the results make sense?
# +
# get word counts
counts_animal = bag_of_words.transform(animal_sents)
counts_language = bag_of_words.transform(language_sents)
# index for "programming"
ind_programming = bag_of_words.vocabulary_['programming']
# total counts across all documents
print(counts_animal.sum(axis=0)[0, ind_programming])
print(counts_language.sum(axis=0)[0, ind_programming])
# -
# ### The `HashingVectorizer` transformer
#
# The `CountVectorizer` requires that we hold the mapping of words to features in memory. In addition, document processing cannot be parallelized because each worker needs to have the same mapping of word to column index. `CountVectorizer` objects are said to have _state_, they retain information of previous interactions and usage. A trick to improve the `CountVectorizer` is to use a hash function to convert the words into numbers. A hash function is a function that converts an input into a _deterministic_ value. In our context, we will use a hash function to convert a word into a number. The resulting number determines which feature column the word is mapped to. Python has a built-in hash function, seen below.
print(hash("hi!"))
print(hash("python"))
print(hash("Pyton"))
print(hash("hi!"))
# Notice how the function returns different values for different words. Also notice, the hash values of "apple" and "apples" are significantly different. Ideally no two inputs result in the same hash value, but this is impossible to avoid; when different inputs generate the same hash, it is referred to as a "hash collision".
#
# The `HashingVectorizer` class is similar to the `CountVectorizer` but it uses a hash function to render it *stateless*. The stateless nature of `HashingVectorizer` objects allows it to parallelize the counting process. There are two main disadvantages of `HashingVectorizer`:
#
# * Hash collisions are possible but in practice are often inconsequential.
# * Because the transformer is stateless, there is no mapping between word to feature index.
# +
from sklearn.feature_extraction.text import HashingVectorizer
hashing_bag_of_words = HashingVectorizer(norm=None) # by default, it normalizes the vectors
hashing_bag_of_words.fit(documents)
hashing_bag_of_words.transform(documents)
# -
# See how the feature matrix has over a million columns? This is in contrast from the result of the count vectorizer. The discrepancy is from the `HashingVectorizer` using, by default, $2^{20}=1048576$ different hash values to construct the count matrix. A vast majority of those indices will have no counts across all documents, and since we represent our feature matrix using a sparse matrix, we pay no cost for empty features!
# +
import time
t_0 = time.time()
CountVectorizer().fit_transform(documents)
t_elapsed = time.time() - t_0
print("Fitting time for CountVectorizer: {}".format(t_elapsed))
t_0 = time.time()
HashingVectorizer(norm=None).fit_transform(documents)
t_elapsed = time.time() - t_0
print("Fitting time for HashingVectorizer: {}".format(t_elapsed))
# -
# ## Term frequency-inverse document frequency
#
# Both the `CountVectorizer` and `HashingVectorizer` creates a feature matrix of raw counts. Using raw counts has two problems, documents vary widely in length and the counts will be large for common words such as "the" and "is". We need to use a weighting scheme that considers the aforementioned attributes. The term frequency-inverse document frequency, **tf-idf** for short, is a popular weighting scheme to improve the simple count based data from the bag of words model. It is the product of two values, the term frequency and the inverse document frequency. There are several variants but the most popular is defined below.
#
# * **Term Frequency:**
# $$ \mathrm{tf}(t, d) = \frac{\mathrm{counts}(t, d)}{\sqrt{\sum_{t \in d} \mathrm{counts}(t, d)^2}}, $$
# where $\mathrm{counts}(t, d)$ is the raw count of term $t$ in document $d$ and $t \in d$ are the terms in document $d$. The normalization results in a vector of unit length.
#
# * **Inverse Document Frequency:**
# $$ \mathrm{idf}(t, D) = \ln\left(\frac{\text{number of documents in corpus } D}{1 + \text{number of documents with term } t}\right). $$
# Every counted term $t$ in the corpus will have its own idf weight. The $1+$ in the denominator is to ensure no division by zero if a term does not appear in the corpus. The idf weight is simply the log of the inverse of a term's document frequency.
#
# With both $\mathrm{tf}(t, d)$ and $\mathrm{idf}(t, D)$ calculated, the tf-idf weight is
#
# $$ \mathrm{tfidf}(t, d, D) = \mathrm{tf}(t, d) \mathrm{idf}(t, D).$$
#
# With the idf weighting, words that are very common throughout the documents get weighted down. The reverse is true; the count of rare words get weighted up. With the tf-idf weighting scheme, a machine learning model will have an easier time to learn patterns to properly predict labels.
# There are two ways to apply the tf-idf weighting in `scikit-learn`, differing in what input they work on. `TfidfVectorizer` works on an array of documents (e.g., list of sentences) while the `TfidfTransformer` works on a count matrix, like the outputs of `HashingVectorizer` and `CountVectorizer`. `TfidfVectorizer` encapsulates the `CountVectorizer` and `TfidfTransformer` into one class. Since we have already calculated the word counts, we will demonstrate the `TfidfTransformer`.
# +
from sklearn.feature_extraction.text import TfidfTransformer
tfidf = TfidfTransformer()
tfidf_weights = tfidf.fit_transform(word_counts)
print(tfidf_weights)
# -
# We no longer have raw counts in our feature matrix. Let's use the `idf_` attribute of the fitted tf-idf transformer to inspect the top idf weights and their corresponding terms.
# +
top_idf_indices = tfidf.idf_.argsort()[:-20:-1]
ind_to_word = bag_of_words.get_feature_names()
for ind in top_idf_indices:
print(tfidf.idf_[ind], ind_to_word[ind])
# -
# Using tf-idf weighting renders the process as _stateful_; to apply the idf weight, we need to know the frequency of each word across all documents. While we may initially use `HasingVectorizer` to have a stateless transformer, coupling it with `TfidfTransformer` will create a stateful process.
# ## Improving signal
#
# So far, we have discussed how using tf-idf rather than raw counts will improve the performance of our machine learning model. There are several other approaches that can boost performance; we will discuss techniques that improve the signal in our data set. Note, the following techniques may marginally increase model performance. It may be best to create a baseline model and measure the increased performance with the new model additions.
# ### Stop words
#
# Words such as "the", "a", and "or" are so common throughout our corpus that they do not contribute any signal to our data set. Further, omitting these words will reduce our already high dimensional data set. It is best to not have these words as features and not be counted in the analysis. The set of words that will not factor into our analysis are called **stop words**.
#
# spaCy provides a `set` of around 300 commonly used English words. When using stop words, it is best to examine the entries in case there are certain words you want to be included or not included. Since the words are provided as a Python `set`, we can use methods available to `set` objects to modify entries of the `set` object.
# +
from spacy.lang.en import STOP_WORDS
print(type(STOP_WORDS))
STOP_WORDS_python = STOP_WORDS.union({"python"})
STOP_WORDS_python
# -
# ### Stemming and lemmatization
#
# In our current analysis, words like "python" and "pythons" will be counted as separate words. We understand that they represent the same concept and want them to be treated as the same word. The same applies to other words like "run", "runs", "ran", and "running", they all represent the same meaning. **Stemming** is the process of reducing a word to its stem. Note, the stemming process is not 100% effective and sometimes the resulting stem is not an actual word. For example, the popular Porter stemming algorithm applied to "argues" and "arguing" returns `"argu"`.
#
# **Lemmatization** is the process of reducing a word to its lemma, or the dictionary form of the word. It is a more sophisticated process than stemming as it considers context and part of speech. Further, the resulting lemma is an actual word. spaCy does not have a stemming algorithm but does offer lemmatization. Each word analyzed by spaCy has the attribute `lemma_` which returns the lemma of the word.
print([word.lemma_ for word in nlp('run runs ran running')])
print([word.lemma_ for word in nlp('buy buys buying bought')])
print([word.lemma_ for word in nlp('see saw seen seeing')])
# **Note**: As of version 2.0.16 of spaCy, there is the bug with the English lemmatization and will fail in instances it should not. However, the bug has been fixed and a patch will be included in a future update, version 2.1.x.
# To apply lemmatization in `scikit-learn`, you need to pass a function to the keyword `tokenizer` of whatever text vectorizer you are deploying. See the example below were we apply lemmatization for a `TfidfVectorizer` transformer.
# +
from sklearn.feature_extraction.text import TfidfVectorizer
def lemmatizer(text):
return [word.lemma_ for word in nlp(text)]
# we need to generate the lemmas of the stop words
stop_words_str = " ".join(STOP_WORDS) # nlp function needs a string
stop_words_lemma = set(word.lemma_ for word in nlp(stop_words_str))
tfidf_lemma = TfidfVectorizer(max_features=100,
stop_words=stop_words_lemma.union({"python"}),
tokenizer=lemmatizer)
tfidf_lemma.fit(documents)
print(tfidf_lemma.get_feature_names())
# -
# ### Tokenization and n-grams
#
# Tokenization refers to dividing up a document into pieces to be counted. In our analysis so far, we are only counting words. However, it may be useful to count a sequence of words such as "natural environment" and "virtual environment". Counting these **bigrams** for our word usage analyzer may boost performance. More generally, an n-gram refers to the n sequence of words. In `scikit-learn`, n-grams can be included by setting `ngram_range=(min_n, max_n)` for the vectorizer, where `min_n` and `max_n` are the lower and upper bound of the range of n-grams to include. For example, `ngram_range=(1, 2)` will include words and bigrams while `ngram_range=(2, 2)` will only count bigrams. Let's see what are the most frequent bigrams in our corpus.
# +
bigram_counter=CountVectorizer(max_features=20, ngram_range=(2,2), stop_words=STOP_WORDS.union({"python"}))
bigram_counter.fit(documents)
bigram_counter.get_feature_names()
# -
# **Questions**
# * Is using stop words more important when using `CountVectorizer`/`HashingVectorizer` or when using the `TfidfVectorizer`?
# * Is it practical to use a large n-gram range, for example, count 3-grams?
# ## Document similarity
#
# After we have transformed our corpus into a matrix, we can interpret our data set as representing a set of vectors in a $p$-dimensional space, where each document is its own vector. One common analysis is to find similar documents. The cosine similarity is a metric that measure how well aligned in space are two vectors, equal to the cosine of the angle in between the two vectors. If the vectors are perfectly aligned, they point in the same direction, the angle they form is 0 and the similarity score is 1. If the vectors are orthogonal, forming an angle of 90 degrees, the similarity metric is 0. Mathematically, the cosine similarity metric is equal to the dot product of two vectors, normalized,
#
# $$ \frac{v_1 \cdot v_2}{\|v_1 \|\|v_2 \|}, $$
#
# where $v_1$ and $v_2$ are two document vectors and $\| v_1 \|$ and $\| v_2 \|$ are their lengths.
# ## Word usage classifier
#
# Let's build a word usage classifier with all the techniques we have seen. The model will include:
#
# * tf-idf weighting
# * stop words
# * words and bigrams
# * lemmatization
#
# Applying the above techniques should result in a data set with enough signal that a machine learning model can learn from. For this exercise, we will use the naive Bayes model; a probabilistic model that calculates conditional probabilities using Bayes theorem. The term naive is applied because it assumes the features are conditionally independent from each other. You can think of a naive Bayes classifier working by determining what class should a document be assigned based upon the frequencies of words in the different classes in the training set. Naive Bayes is often used as benchmark model for NLP as it is quick to train. More about the model in general can be found [here](https://en.wikipedia.org/wiki/Naive_Bayes_classifier) and details of the `scikit-learn` implementation is found [here](https://scikit-learn.org/stable/modules/naive_bayes.html). After training our model, we will see how well it performs for a chosen set of sentences.
# +
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
# create data set and labels
documents = animal_sents + language_sents
labels = ["animal"]*len(animal_sents) + ["language"]*len(language_sents)
# lemma of stop words
stop_words_str = " ".join(STOP_WORDS)
stop_words_lemma = set(word.lemma_ for word in nlp(stop_words_str))
# create and train pipeline
tfidf = TfidfVectorizer(stop_words=stop_words_lemma, tokenizer=lemmatizer, ngram_range=(1, 2))
pipe = Pipeline([('vectorizer', tfidf), ('classifier', MultinomialNB())])
pipe.fit(documents, labels)
print("Training accuracy: {}".format(pipe.score(documents, labels)))
# +
test_docs = ["My Python program is only 100 bytes long.",
"A python's bite is not venomous but still hurts.",
"I can't find the error in the python code.",
"Where is my pet python; I can't find her!",
"I use for and while loops when writing Python.",
"The python will loop and wrap itself onto me.",
"I use snake case for naming my variables.",
"My python has grown to over 10 ft long!",
"I use virtual environments to manage package versions.",
"Pythons are the largest snakes in the environment."]
class_labels = ["animal", "language"]
y_proba = pipe.predict_proba(test_docs)
predicted_indices = (y_proba[:, 1] > 0.5).astype(int)
for i, index in enumerate(predicted_indices):
print(test_docs[i], "--> {} at {:g}%".format(class_labels[index], 100*y_proba[i, index]))
# -
# ## Exercises
#
# 1. Encapsulate the entire process of gathering a corpus, constructing, and training a model into a function. Afterwards, deploy the model to other sets of homonyms.
# 1. Measure the model's improvements by stripping out things such as the use of stop words and lemmatization. Perhaps you can incorporate model additions as parameters to the previously mentioned function. What model additions increases the performance the most?
# 1. Consider another source of data and see how well the model performs with the new corpus.
# 1. Naive Bayes classifier calculates conditional probabilities from the training set. In other words, it determines values like $P(\text{snake | }Y = \text{animal})$, the probability a document has the word "snake" given if the document belongs to those of python the animal. These values are stored in `coef_` attribute of a trained naive Bayes model. Can you use these coefficients to determine the most discriminative features? In other words, what terms when found in a document really help classify the document.
# *Copyright © 2020 The Data Incubator. All rights reserved.*
|
9_ML_Natural_Language_Processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import can
import udsoncan
# +
bus = can.Bus(interface='socketcan',
channel='vcan0',
receive_own_messages=True)
# send a message
message = can.Message(arbitration_id=123, is_extended_id=True,
data=[0x11, 0x22, 0x33])
bus.send(message, timeout=0.2)
# iterate over received messages
for msg in bus:
print("{:X}: {}".format(msg.arbitration_id, msg.data))
# -
# or use an asynchronous notifier
notifier = can.Notifier(bus, [can.Logger("recorded.log"), can.Printer()])
|
notebooks/00_Bootstrap.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### XGBoosting Easy version
# +
import numpy as np # array, vector, matrix calculations
import pandas as pd # DataFrame handling
#import shap # for consistent, signed variable importance measurements
import xgboost as xgb # gradient boosting machines (GBMs)
import math
import matplotlib.pyplot as plt # plotting
pd.options.display.max_columns = 999 # enable display of all columns in notebook
# enables display of plots in notebook
# %matplotlib inline
np.random.seed(42) # set random seed for reproducibility
# +
# import XLS file
path = ".\\credit_cards_dataset.csv"
#data = pd.read_excel(path, skiprows=1) # skip the first row of the spreadsheet
#path = 'C:\\Users\\User\\Desktop\\data\\original_data.csv'
#data = pd.read_csv(path, skiprows=1) # skip the first row of the spreadsheet
data = pd.read_csv(path) # skip the first row of the spreadsheet
# remove spaces from target column name
data = data.rename(columns={'default payment next month': 'DEFAULT_NEXT_MONTH'})
# -
# assign target and inputs for GBM
#y = 'DEFAULT_NEXT_MONTH'
y='default.payment.next.month'
X = [name for name in data.columns if name not in [y, 'ID', 'Y_Value']]
print('y =', y)
print('X =', X)
# +
split_ratio=0.7
# execute split
split = np.random.rand(len(data)) < split_ratio
train=data[split]
test=data[~split]
print('Train data rows = %d. columns = %d' % (train.shape[0], train.shape[1]))
print('Test data rows = %d. columns = %d' % (test.shape[0], test.shape[1]))
# +
# XGBoost uses SVMLight data structure, not Numpy arrays or Pandas DataFrames
mod = xgb.XGBRegressor(
gamma=1,
learning_rate=0.01,
max_depth=3,
n_estimators=10000,
subsample=0.8,
random_state=42,
verbosity=1
)
# -
mod.fit(train[X], train[y])
predictions = mod.predict(test[X])
from sklearn.metrics import mean_squared_error
test[y]
rmse = math.sqrt(mean_squared_error(test[y], predictions))
print(rmse)
#print("score: {0:,.0f}".format(rmse))
predictions=np.rint(predictions)
predictions
# +
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn import metrics
from sklearn.model_selection import GridSearchCV
accuracy = accuracy_score(test[y], predictions)
cm = confusion_matrix(test[y], predictions)
precision = precision_score(test[y], predictions)
recall = recall_score(test[y], predictions)
# -
print(accuracy)
print(cm)
print(precision)
print(recall)
# +
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
import itertools
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
# -
plt.figure()
plot_confusion_matrix(cm, classes=['Non_Default','Default'], normalize=False,
title='Non Normalized confusion matrix')
|
Big_Data_Analysis/feature_selection/12-1xgb(easy).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.5
# language: julia
# name: julia-1.0
# ---
# Objectives :
# Re-itirate and understand
#
#
# Setting up training and test data sets
#
#
# Data pre-processing
#
#
# model performance Classification
#
#
# Cross-validation
using RDatasets
using Distributions
using MLBase
iris =dataset("datasets","iris")
features=iris[:,1:4]
labels=convert( Array , iris[:,5])
nrow(iris)
# +
#just loop revision . don't notice me . hehe xd
#train= Int64[]
#for i in 1:nrow(iris)
# push!(train, 1)
#end
# -
train=rand(Bernoulli(0.75),nrow(iris)) #random array of trues and false to feed to the model as training data
show(train)
typeof(features)
print(features[train,:])
labels[train,:]
#convert(UInt8, x)
test=Bool[]
for i in 1:nrow(iris)
push!(test,!train[i])
end
show(test)
length(test)
features[test,:]
labels[test,:]
species=["setosa","versicolor","virginica"]
lm=labelmap(species) #code the species to 1 2 3
labelencode(lm,labels[train,:]) #replace the label with the encoded value
MLBase.groupindices(lm,labels[train,:]) #show the indices of each encoded value in the training set
# # Basic teamplate of how the modeling phase goes in ML :
#
# model=build_model(labels[train,:],features[train,:])
#
# model=prune_model(model,0.7)
#
# print_tree(model)
#
# predictions=apply_model(model,features[test,:])
#
# #gridtune method is also useful for adjusting the model parameters
#
# #performance Checks
#
# gt=["setosa","versicolor","virginica","setosa","virginica"]
#
# pred=["setosa","versicolor","setosa","setosa","virginica" ]
#
# gtl=labelencode(lm,gt)
#
# predl=labelencode(lm,pred)
#
# correctrate(gtl,predl)
#
# errorrate(gtl,predl)
#
# confusmat(3,gtl,predl)
# +
#Receiver Operating Characteristics ROC
#r=roc(gtl,predl)
# +
#true_positive_rate(r)
#true_negative_rate(r)
#precision(r)
#recall(r)
# -
# Classification using Decision Trees and Rules
#
#
# Objectives :
#
#
# Understand the usage of Decision Trees
#
#
# Pruned Tree classifer
#
#
# Random Forest Classifer
#
#
# adaptive Boosted Decision Stump Classifier
iris[:,1:4]
iris[:,1:4][2,3]
using DecisionTree #warning Decision Trees have a habit of overfitting the model to the dataset
iris =dataset("datasets","iris")
features=convert(Array , features)
labels=convert( Array , iris[:,5])
model=build_tree(labels,features)
model=prune_tree(model,0.9)
print_tree(model,5) #5level depth
apply_tree(model,[5.9,3.0,.1,1.9])
# +
#cross-validation
accuracy=nfoldCV_tree(labels,features, 0.9,3 ) #90% purity #3folds
#this function has not been maintained in Julia 1.0.5 but after chatting with support and
#community members on discord they confirmed that the DecisionTree package will overhauled and the cross validation function
#will drop in the upcomming version
# -
using Base.Test
@test mean(accuracy)
#improve performance by using Adaptive_boosted Decisions Stumps Classifier
model,coeffs=build_adaboost_stumps(labels,features,10)
accuracy=nfoldCV_stumps(labels,features, 10 ,3 ) #3folds
#in this specific dataset the boosting technique gives us no improvement probably due to outliers or other problems
# Using a Random Forest Classifier
#
# to create multiple trees and ouputting a class and it's the mean prediction of the trees' ouputs
model=build_forest(labels,features,2,10,0.5)
#2 random features #10 trees #0.5 of samples per tree
#accuracy rising
|
Machine Learning/Julia/Basics of Machine Learning with Julia.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import necessary packages
import os
import glob
import pandas as pd
import numpy as np
from scipy import stats
import scikit_posthocs
import iqplot
import bokeh.io
bokeh.io.output_notebook()
# # Import data from directory of measurement tables, collected from Fiji
# +
# Define path to directory with measurements
path = os.path.abspath('2_Puncta/')
df_summary = pd.DataFrame()
list_summary = []
# For loop to bring in files and concatenate them into a single dataframe
for file_ in glob.glob(path + "/*Summary.csv"):
df = pd.read_csv(file_)
# Determine Image name from file name, then parse experiment details from Image name
df['Image'] = os.path.splitext(os.path.basename(file_))[0]
# (df['Date'], df['CellLine'], df['Dose'], df['Time'], df['Treatment'],
# df['Channels'], df['Mag'], df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
(df['Date'], df['CellLine'], df['Treatment'], df['Rep'], df['Channels'],
df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
# (df['Date'], df['Treatment'], df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
# Drop unnecessary columns for tidiness
df = df.drop(['Total Area', 'Average Size', '%Area', 'Mean', 'IntDen', 'Image', 'Channels'], axis = 1)
# Compile data
list_summary.append(df)
df_summary = pd.concat(list_summary, sort=False)
# Preview dataframe to confirm import successful
df_summary.head()
# -
# # Parse dataframe by desired 'cell line' and 'treatment' combinations, then plot results
# +
# Generate and display list of cell lines and treatments present in this dataset
cell_list = df_summary['CellLine'].unique().tolist()
treatment_list = df_summary['Treatment'].unique().tolist()
print('Cells lines: ' + str(cell_list))
print('Treatments: ' + str(treatment_list))
# +
# Prepare for parsing data:
# To populate this "comment on/off" code block, copy the results of th cell lists above
cells = [
'U2OS',
# 'DF1',
]
treatments = [
'2aRFP',
'Dyn1K44A',
'SMPD3',
'SMPD3N130A',
# 'SGMS2',
# 'ControlgRNA',
# 'SMPD3gRNA',
]
# Copy dataset to not disrupt raw data
df_subset = df_summary
# Pull out only cells and treaments of interest
df_subset = df_subset.loc[df_subset['CellLine'].isin(cells)]
df_subset = df_subset.loc[df_subset['Treatment'].isin(treatments)]
# df_subset = df_subset['Count'].dropna()
# Make ECDF plot using iqplot
data_ecdf = iqplot.ecdf(
data=df_subset, q='Count', cats='Treatment'
,title=str(cells) + ' cells treated with ' + str(treatments), style='staircase'
# ,conf_int=True, n_bs_reps=1000, ptiles=[2.5, 97.5] # ptiles values equate to 95% CIs
,conf_int=True, n_bs_reps=1000, ptiles=[16, 84] # ptiles values equate to 68% CIs (SEM)
,show_legend=True
# Other customization parameters
,frame_height = 300, frame_width = 500
,order = ['2aRFP', 'Dyn1K44A', 'SMPD3', 'SMPD3N130A']
# ,palette = ['#E8754F', '#2066AC', '#68ADCC']
# ,x_axis_label='Transferrin-633 Puncta', y_axis_label='ECDF'
)
# Other customization parameters
data_ecdf.title.text_font_size = '18px'
data_ecdf.legend.title='Treatment'
# data_ecdf.legend.title_text_font_style = 'bold'
data_ecdf.legend.title_text_font_size = '16px'
data_ecdf.legend.label_text_font_size = '14px'
data_ecdf.axis.axis_label_text_font_size = '16px'
data_ecdf.axis.axis_label_text_font_style = 'bold'
data_ecdf.axis.major_label_text_font_size = '14px'
bokeh.io.show(data_ecdf)
# -
# # Kruskal-Wallis Test with Dunn's Multiple Comparisons Correction
# Useful for comparing multiple datasets
# Reminder of treatments to compare
treatment_list
# +
### Kruskal-Wallis test
# Define samples to compare
sample1 = '2aRFP'
sample2 = 'Dyn1K44A'
sample3 = 'SMPD3'
sample4 = 'SMPD3N130A'
# sample5 = 'SGMS2'
# sample6 = 'nSMase2gRNA1'
# Run Kruskal-Wallis test
kw_result = stats.kruskal(
df_subset.loc[df_subset['Treatment']==sample1]['Count']
,df_subset.loc[df_subset['Treatment']==sample2]['Count']
,df_subset.loc[df_subset['Treatment']==sample3]['Count']
,df_subset.loc[df_subset['Treatment']==sample4]['Count']
# ,df_subset.loc[df_subset['Treatment']==sample5]['Count']
# ,df_subset.loc[df_subset['Treatment']==sample6]['Count']
)
# Dunn's Posthoc for Multiple Comparisons
mult_compar = scikit_posthocs.posthoc_dunn(df_subset, val_col='Count', group_col='Treatment'
, sort=False, p_adjust='bonferroni').round(6)
# Display test results
print('Kruskal-Wallis test results: \n\t\t\t\t statistic=' + str(kw_result[0]) +
'\n\t\t\t\t p-value=' + str(kw_result[1]))
print("\nDunn's posthoc multiple comparison result: \n" + str(mult_compar))
# mult_compar.to_csv("Results of Dunn's Posthoc.csv")
# -
# # Kolmogorov-Smirnov Test
# Useful for comparing two datasets, not multiple comparisons
# +
### Kolmogorov-Smirnov test - NO MULTIPLE COMPARISONS
# Define samples to compare
sample1 = '2aRFP'
sample2 = 'SMPD3'
# Run 2-sample Kolmogorov-Smirnov Test
ks_result = stats.ks_2samp(df_subset.loc[df_subset['Treatment']==sample1]['Count']
,df_subset.loc[df_subset['Treatment']==sample2]['Count']
)
# Display results of Kolmogorov-Smirnov test
print('Two-sample Kolmogorov-Smirnov test results for ' + sample1 + ' vs ' + sample2 + ': \n\t\t\t\t statistic=' + str(ks_result[0]) +
'\n\t\t\t\t p-value=' + str(ks_result[1]))
# -
# ## Examine data using other plots from iqplot
# +
# Make the variety of plots
data_box =iqplot.box(data=df_subset, q='Count', cats='Treatment', q_axis='y',
# order=['RFP', 'Dyn1KA'],
whisker_caps=True, frame_height = 300, frame_width = 200)
data_strip =iqplot.strip(data=df_subset, q='Count', cats='Treatment', q_axis='y',
# order=['RFP', 'Dyn1KA'],
jitter=True, frame_height=300, frame_width=200)
data_histo = iqplot.histogram(data=df_subset, q='Count', cats='Treatment', density=True, frame_width=550)
# Display plots
bokeh.io.show(bokeh.layouts.gridplot([data_box, data_strip, data_histo], ncols=3))
# -
data_strip =iqplot.strip(data=df_subset, q='Count', cats='Treatment', q_axis='y',
# order=['RFP', 'Dyn1KA'],
jitter=True, frame_height=300, frame_width=200)
bokeh.io.show(data_strip)
# +
# Define path to directory with measurements
path = os.path.abspath('2_Puncta/')
df_summary = pd.DataFrame()
list_summary = []
df_details = pd.DataFrame()
list_details = []
# For loop to bring in files and concatenate them into a single dataframe
for file_ in glob.glob(path + "/*Summary.csv"):
df = pd.read_csv(file_)
# Determine Image name from file name, then parse experiment details from Image name
df['Image'] = os.path.splitext(os.path.basename(file_))[0]
(df['Date'], df['CellLine'], df['Treatment'], df['Rep'], df['Channels'],
df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
# Drop unnecessary columns for tidiness
df = df.drop(['Total Area', 'Average Size', '%Area', 'Mean', 'IntDen', 'Image', 'Channels'], axis = 1)
# Compile data
list_summary.append(df)
df_summary = pd.concat(list_summary, sort=False)
# For loop to bring in files and concatenate them into a single dataframe
for file_ in glob.glob(path + "/*Details.csv"):
df = pd.read_csv(file_)
# Determine Image name from file name, then parse experiment details from Image name
df['Image'] = os.path.splitext(os.path.basename(file_))[0]
(df['Date'], df['CellLine'], df['Treatment'], df['Rep'], df['Channels'],
df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
(df['ROI'], df['?unknown?']) = zip(*df['Label'].map(lambda x:x.split(':')))
df = df.rename({" ": "?Vesicle?"}, axis=1)
# Drop unnecessary columns for tidiness
df = df.drop(['Mean', 'IntDen', 'Image', 'Channels', 'Mean', 'IntDen', 'RawIntDen'], axis = 1)
# Compile data
list_details.append(df)
df_details = pd.concat(list_details, sort=False)
full_df = pd.merge(df_summary, df_details, on=['Date', 'CellLine', 'Treatment', 'Rep', 'FOV'])
# full_df = full_df.dropna()
# Preview dataframe to confirm import successful
# full_df
|
Endocytosis_Analysis_Confocal/20200823_Tf633PunctaCounts_CatData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # xspec Documentation (v1.3.2)
#
# This ipython Notebook is intended to provide documentation for the linetools GUI named XSpecGUI.
#
# Enjoy and feel free to suggest edits/additions, etc.
#
# Here is a screenshot of the XSpecGUI in action:
from IPython.display import Image
Image(filename="images/xspec_example.png")
# The example spectrum file used below is part of the linetools package.
import imp
lt_path = imp.find_module('linetools')[1]
spec_fil = lt_path+'/spectra/tests/files/PH957_f.fits'
# ## Before Launching the GUI
#
# If you are a Mac user, we **highly** recommend that you set your
# matplotlib backend from MacOSX to TkAgg (or another option, see
# [backends](http://matplotlib.org/faq/usage_faq.html#what-is-a-backend)).
#
# ## Launching the GUI
# ### From the command line (recommended)
# We recommend you use the script provided with linetools.
#
# Then it is as simple as:
#
# > lt_xspec filename
#
# Here are the current command-line options:
# profx.ucolick.org> lt_xspec -h
# usage: lt_xspec [-h] [-z ZSYS] [--norm] [--air] [--exten EXTEN]
# [--splice SPLICE]
# file
#
# Parse for XSpec; Extra arguments are passed to read_spec
#
# positional arguments:
# file Spectral file; specify extension by appending #exten#
#
# optional arguments:
# -h, --help show this help message and exit
# -z ZSYS, --zsys ZSYS System Redshift
# --norm Show spectrum continuum normalized (if continuum is
# provided)
# --air Convert input spectrum wavelengths from air to vacuum
# --exten EXTEN FITS extension
# --splice SPLICE Splice with the input file; extension convention
# applies
# ### From within ipython or equivalent
# from linetools.guis import xspecgui as ltxsg
#
# import imp; imp.reload(ltxsg)
# ltxsg.main(spec_fil)
# ### File inputs
#
# You may specify an extension for multi-extension binary tables by appending the number within # signs, e.g.
# lt_xspec file.fits#1#
# ### Splice
# Use --splice=filename.fits to splice a second spectrum to the first
# ---
# ## Navigating - These key strokes help you explore the spectrum (be sure to click in the spectrum panel first!)
#
# ### Setting the window edges (mouse+keystroke)
# * l -- Set left edge of window
# * r -- Set right edge of window
# * t -- Set top edge of window
# * b -- Set bottom edge of window
# * Z -- Set bottom edge to 0.
# * W -- View full spectrum
# * s,s -- Set a zoom-in window at 2 mouse positions
#
# ### Zoom in/out Wavelength
# * i -- Zoom in on cursor
# * I -- Zoom in extra fast
# * o -- Zoom out
# * O -- Zoom out extra fast
#
# ### Best estimate Flux limits
# * y -- Make a guess for the flux plotting limits
#
# ### Zoom out Flux
# * Y -- Zoom out
#
# ### Pan
# * [ -- Pan left
# * { -- Pan left extra
# * ] -- Pan right
# * } -- Pan right extra
#
#
#
# ---
# ## Overlaying Line Lists
# You can overlay a series of vertical lines at standard spectral
# lines at any given redshift.
#
# ### Setting the Line List
# You must choose a line-list by clicking one.
#
# ### Setting the redshift
# * Type one in
# * RMB on a spectral feature (Ctrl-click on Emulated 3-button on Macs)
# * Choose the rest wavelength
#
# ### Marking Doublets
# * C -- CIV
# * M -- MgII
# * X -- OVI
# * 4 -- SiIV
# * 8 -- NeVIII
# * B -- Lyb/Lya
#
# ### Velocity plot
# Once **both** a line list and redshift are set, type 'v'
# to launch a Velocity Plot GUI.
# ---
# ## Simple Analysis
#
#
# ### Gaussian Fit
#
# You can fit a Gaussian to any single feature in the spectrum
# as follows:
# 1. Click "G" at the continuum at one edge of the feature
# 1. And then another "G" at the other edge (also at the continuum)
# 1. A simple Gaussian is
# fit and reported.
#
# ### Equivalent Width
# You can measure the rest EW of a spectral feature as follows:
# 1. Click "E" at the continuum at one edge of the feature
# 1. And then another "E" at the other edge (also at the continuum)
# 1. A simple boxcar integration is performed and reported.
#
# ### Apparent Column Density
# You can measure the apparent column via AODM as follows:
# 1. Click "N" at the continuum at one edge of the feature
# 1. And then another "EN" at the other edge (also at the continuum)
# 1. A simple AODM integration is performed and reported.
#
# ### Ly$\alpha$ Lines
# * "D" - Plot a DLA with $N_{\rm HI} = 10^{20.3} \rm cm^{-2}$
# * "R" - Plot a SLLS with $N_{\rm HI} = 10^{19} \rm cm^{-2}$
|
docs/examples/xspecgui.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### <NAME> - Udacity / SDCND - May./2019.
#
#
# # Self-Driving Car Engineer Nanodegree
#
# ## Project3: ** Traffic Sign Recognition**
#
# The goals / steps of this project are the following according to the [rubric points](https://review.udacity.com/#!/rubrics/481/view)
#
# 1. **Dataset Exploration**<br/>
# 1.1 Dataset summary.<br/>
# 1.2 Exploratory Visualization.<br/>
#
# 2. **Design and Test a Model Architecture**<br/>
# 2.1 Preprocessing.<br/>
# 2.2 Model Architecture.<br/>
# 2.3 Model Training.<br/>
# 2.4 Solution Approach.<br/>
#
# 3. **Test a Model on New Images**<br/>
# 3.1 Acquiring New Images.<br/>
# 3.2 Performance on New Images.<br/>
# 3.3 Model Certainty - Softmax Probabilities.<br/>
#
#
# **Here is a link to [code file](./Advanced_Lane_Finding.ipynb)**
#
# [//]: # (Image References)
#
# [image1]: ./output_images/dataset_expl_qty.jpg
# [image2]: ./output_images/dataset_expl_percent.jpg
# [image3]: ./output_images/dataset_German_traffic_signs.jpg
#
# [image4]: ./output_images/same_labes_examples1.jpg
# [image5]: ./output_images/same_labes_examples2.jpg
# [image6]: ./output_images/same_labes_examples3.jpg
# [image7]: ./output_images/same_labes_examples4.jpg
# [image8]: ./output_images/same_labes_examples5.jpg
# [image9]: ./output_images/same_labes_examples6.jpg
#
#
# [image10]: ./output_images/preprocessed_examples1.jpg
# [image11]: ./output_images/preprocessed_examples2.jpg
# [image12]: ./output_images/preprocessed_examples3.jpg
#
# [image13]: ./output_images/lenet.jpg
#
# [image14]: ./output_images/Training_process.jpg
#
# [image15]: ./output_images/New_images.jpg
#
# [image16]: ./output_images/New_images_preprocessed.jpg
#
# [image17]: ./output_images/5_New_images_selecteds.jpg
#
# [image18]: ./output_images/5_New_images_predictions.jpg
#
#
#
# [image19]: ./output_images/softmax_probabilities1.jpg
# [image20]: ./output_images/softmax_probabilities2.jpg
# [image21]: ./output_images/softmax_probabilities3.jpg
# [image22]: ./output_images/softmax_probabilities4.jpg
# [image23]: ./output_images/softmax_probabilities5.jpg
#
#
#
#
# _____
#
# ## 1 - Load the data set.
#
# ### 1.1 - Dataset sumary.
#
# The dataset was analysed using dictionary function from python. The table below shows the summary.
#
# | Features | quantity |
# |:-----------------:|:------------------:|
# | training | 34799 |
# | testing | 12630 |
# | Validation | 4410 |
#
# The image shape for the features is: 32x32x3
#
# The number of unique classes (Labels) is: 43
#
# The next histograms show the distribution for the labels for each set. (Train,Validation and Test).
#
# ![alt text][image1]
#
#
# Regarding the label distribution for the datasets, it is possible identify that we have the approximately the same distributions from the percentiles of the labels comparing with the total for the each Dataset. The next histogram show the distribution:
#
# ![alt text][image2]
#
# ---
#
# ### 1.2 - Exploratory Visualization.
#
# The Next picture shows the 43 Labels used as traffic signs dataset and the image of them.
#
# ![alt text][image3]
#
# From the histogram, below it is possible see 7 differents images from the same Label. The Labels 2, 1 and 13 are the labels with more number of examples and the Labels 0, 19 and 37 are the labels with less number of examples.
#
# ![alt text][image4]
# ![alt text][image5]
# ![alt text][image6]
# ![alt text][image7]
# ![alt text][image8]
# ![alt text][image9]
#
# _____
#
#
# ## 2 -Design and Test a Model Architecture.
#
# ### 2.1 - Preprocessing.
#
# For the Pre- process was applied 3 steps to the image data.<br/>
# $1^{st}$ - Converted to gray scale.<br/>
# $2^{nd}$ - Normalized the image data using the formula (pixel - 128)/ 128<br/>
# $3^{rd}$ - reshape image to add channel and reach the shape: 32x32x1<br/>
#
# These techniques were chosen to normalize the images, so that the data has mean zero and equal variance and ensure a shape to be correctly in the next steps. ( Tensorflow functions).
#
# The final results are shown below in 7 pictures as example for the labels 2,1 and 13:
#
# ![alt text][image10]
# ![alt text][image11]
# ![alt text][image12]
#
#
# ### 2.2 - Model Architecture.
#
# The Convolutional Neural networks used have the same architecture of the example for Lenet-5. Used in the Udacity class, but the difference is that the last Linear Layer was chane the value from 10 to 50.
# The architeture is: Two convolutional layers followed by one flatten layer, drop out layer, and Three fully connected linear layers.As the follow picture.
#
# ![alt text][image13]
# Source: Yan LeCun / edited to adjust the new value for the last layer.
#
# Details about the architeture:
#
# **Input:** Image in GrayScale with the size: 32x32x1
#
# 1. convolution 1: 32x32x1 -> 28x28x6 -> relu -> 14x14x6 (pooling)
# 2. convolution 2: 14x14x6 -> 10x10x16 -> relu -> 5x5x16 (pooling)
# 3. flatten: 5x5x16 -> 400
# 4. drop out: 400 -> 120
# 5. linear: 120 -> 84
# 6. linear: 84 -> 50
#
# **Output:** Return the result of the 2nd fully connected layer. Size = 50
#
# It was canceled the last layer to check the accuracy for the validation Set, the results is very similar , but the results to predict the label for the images from the internet it was not good. For this reason was kept the architeture as the LeNet-5 example.
#
#
# ### 2.3 - Model Training.
#
# Here working with a lower value for 'Batch_size', I have realized an opportunity to increase the accuracy over than 0.98 in the validation data set, but the results accuracy with the Dowloaded images is very bad.So it was necessary work with a high value for batch size. The Number of EPOCHS greather than 10 did not show advantages to justify use values higher than 10.
# the Learning rate have contributed working with values less than $10^{-3}$. Below the parameter values:
#
# EPOCHS = 10<br/>
# BATCH_SIZE = 200<br/>
# learning rate 0.001<br/>
#
# The training process was executed using the Tensor flow library. and could be checked from cells 20 to 26 in the [code file](./Advanced_Lane_Finding.ipynb)
#
#
#
# ### 2.4 - Solution Approach.
#
# The picture below shows the accuracy variation over each EPOCH:
#
# ![alt text][image14].
#
# The Validation accuracy have accomplished the value above the target of the project 0.93.<br/>
# The parameter and the architeture were checked in order to reached a high accuracy with the new images from the internet. The results will be shown on the next part of this write up.
#
# ____
#
# ## 3 - Test a Model on New Images.
#
# ### 3.1 - Acquiring New Images.
#
# The New images from the internet for German traffic signs could be see in this [link](./German_traffic_Signs_Download/original_images).
#
# It was neccesary crop the images to isolate the traffic signs and this step could be see in this [link](./German_traffic_Signs_Download/Manual_crop_images).
#
#
# To use the pictures to check the accuracy using the model trained and already explained previously, it was necessary load the data , add margins do adapt the traffic signs image to similar of the data set used ,it was necessary also apply the `blur function` from the cv2 library ,before create the data set with the new images.
#
# Below it is possible to see that it was capture 15 pictures from the internet.
#
# ![alt text][image15].
#
# It was applied the same preprocessing function to match the new images to the others available on the data-set reference.
#
# ![alt text][image16].
#
#
# Using the `random.sample` function , it was selected 5 picures to check the accuracy.
# Below the pictures selected.<br/>
#
# ![alt text][image17].
#
# ### 3.2 - Performance on New Images.
#
# Below we have the same five images and the OK title where the prediction have reached success.
#
# ![alt text][image18].
#
# It is possible to see that our model under a small data set with only 5 images reach an accuracy value equal 1 (100%).
#
#
# ### 3.3 - Model Certainty - Softmax Probabilities.
#
# In our example, using the Tensor flow function `tf.nn.top_k` and K value equal to 5. it is possible see 5 probabilities for each image.
#
# From the cells 45 to 49 were applied this function and the final result is an image showing in the left side the original image and the others 5 images in the same row are the probability to the model reach the success to predict the traffic sign. It is possible have a look that in the 5 rows number the model hit the correct label. It is the same result shown in the performance on New Images (item 3.2) confirming the accuracy = 1.
#
#
# ![alt text][image19].
# ![alt text][image20].
# ![alt text][image21].
# ![alt text][image22].
# ![alt text][image23].
#
#
#
#
|
writeup_Eduardo_Campos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Datasheet for "Where it pays to go to college"
# ## *Motivation for Data Creation*
#
# #### Why was the dataset created? (e.g., were there specific tasks in mind, or a specific gap that needed to be filled?)
#
# The salary report was created by PayScale to provide its users with a detailed compensation report that compares their compensation to others like them. “Where it Pays to Attend College” was created by the WSJ by extracting some of the data from PayScale’s report to provide information and compare the incomes of certain college graduates (mainly Ivy League) depending on what (major) and where (university) they studied during undergrad.
#
# #### What (other) tasks could the dataset be used for? Are there obvious tasks for which it should not be used?
# Nothing too obvious.
#
# #### Has the dataset been used for any tasks already? If so, where are the results so others can compare (e.g., links to published papers)?
# The dataset was somewhat analyzed by the WSJ in Ivy Leaguers’ Big Edge: Starting Pay: https://www.wsj.com/articles/SB121746658635199271
# Also, some of Kaggle users did their own analysis: https://www.kaggle.com/wsj/college-salaries/kernels
#
# #### Who funded the creation of the dataset? If there is an associated grant, provide the grant number.
# There is no mentioning of any funding for this dataset both in the WSJ, PayScale and Kaggle.
#
# ## *Data Composition*
#
# #### What are the instances? (that is, examples; e.g., documents, images, people, countries) Are there multiple types of instances? (e.g., movies, users, ratings; people, interactions between them; nodes, edges)
#
# There are 3 subsets to the dataset. In total, there are multiple types of instances such as different salaries (starting, mid-career percentiles), colleges (name and type), major, and region.
#
# #### Are relationships between instances made explicit in the data?
# There are no known relationships between instances except for the fact that graduate degree holders and self-employed/contract-based employees were excluded from the dataset.
#
# #### How many instances of each type are there?
# There are 50 majors and corresponding salaries; 269 universities and types and corresponding salaries; and 320 universities and region and corresponding salaries.
#
# #### What data does each instance consist of? “Raw” data (e.g., unprocessed text or images)? Features/attributes? Is there a label/target associated with instances? If the instances are related to people, are subpopulations identified (e.g., by age, gender, etc.) and what is their distribution?
# Each instance for the major subset contains associated starting, mid-career, and percentiles mid-career salaries. Same goes for university/type and region instances. The subpopulation data was not available in the WSJ except for the fact that there are no graduate degree holders/self-employed/contract workers.
#
# #### Is everything included or does the data rely on external resources? (e.g., websites, tweets, datasets) If external resources, a) are there guarantees that they will exist, and remain constant, over time; b) is there an official archival version. Are there licenses, fees or rights associated with any of the data?
# Some of the salaries are missing, and there are some duplicates.
#
# #### Are there recommended data splits or evaluation measures? (e.g., training, development, testing; accuracy/AUC)
# The data comes with specified splits (3 subsets) for major, college/type and college/region. The recommended measures are unknown.
#
# #### What experiments were initially run on this dataset? Have a summary of those results and, if available, provide the link to a paper with more information here.
# The WSJ’s article can be found here: https://www.wsj.com/articles/SB121746658635199271 and it mainly suggests the following:
# * The median starting salary for Ivy Leaguers is 32% higher than that of liberal-arts college graduates
# * The incomes of graduates from all types of schools grow almost at the same rate
# * Salary increase percentage in 10 years is highest for liberal art schools (95%) but lowest for engineering (76%)
# * It is more about the career path one chooses rather than the major one studied
#
#
# ## *Dataset Collection Process*
#
# #### How was the data collected? (e.g., hardware apparatus/sensor, manual human curation, software pro- gram, software interface/API; how were these con- structs/measures/methods validated?)
# The data used in PayScale’s College Salary Report was collected through online compensation survey, where people self-report data about their jobs, compensation, employer, demographics and educational background. However, it is unknown whether the WSJ copied the full dataset or extracted some part of it.
#
# #### Who was involved in the data collection process?(e.g., students, crowdworkers) How were they compensated? (e.g., how much were crowdworkers paid?)
# Anyone who wanted to get PayScale’s compensation report. They were “compensated” with a detailed compensation report that compares their compensation to others like them.
#
# #### Over what time-frame was the data collected? Does the collection time-frame match the creation time-frame?
# For this dataset unknown. However, PayScale’s survey is ongoing.
#
# #### How was the data associated with each instance acquired? Was the data directly observable (e.g., raw text, movie ratings), reported by subjects (e.g., survey responses), or indirectly inferred/derived from other data (e.g., part of speech tags; model-based guesses for age or language)? If the latter two, were they validated/verified and if so how?
# The data was reported by individuals (through a survey.) PayScale claims that the data is verified2, but it is unknown how.
#
# #### Does the dataset contain all possible instances? Or is it, for instance, a sample (not necessarily random) from a larger set of instances?
# It is a sample from a larger set, aka the dataset does not contain all possible instances.
#
# #### If the dataset is a sample, then what is the population? What was the sampling strategy (e.g., deterministic, probabilistic with specific sampling probabilities)? Is the sample representative of the larger set (e.g., geographic coverage)? If not, why not (e.g., to cover a more diverse range of in- stances)? How does this affect possible uses?
# PayScale states: “the sample size is 3.2 M. The sample size for each school included ranges from 28 profiles to over 25,000, depending largely upon the size of the school.”2 However, it is unclear what was the sampling strategy used for the WSJ.
#
# #### Is there information missing from the dataset and why? (this does not include intentionally dropped instances; it might include, e.g., redacted text, withheld documents) Is this data missing because it was unavailable?
# Unknown. (the WSJ does not provide any explanation)
#
# #### Are there any known errors, sources of noise, or redundancies in the data?
# Missing data
#
# ## *Data Preprocessing*
#
# #### What preprocessing/cleaning was done? (e.g., discretization or bucketing, tokenization, part-of-speech tagging, SIFT feature extraction, removal of instances, processing of missing values, etc.)
# Unknown. (neither the WSJ nor PayScale does not provide any information)
#
# #### Was the “raw” data saved in addition to the preprocessed/cleaned data? (e.g., to support unanticipated future uses)
# Unknown.
#
# #### Is the preprocessing software available?
# Unknown.
#
# #### Does this dataset collection/processing procedure achieve the motivation for creating the dataset stated in the first section of this datasheet?
# Unknown.
#
# #### Any other comments?
# PayScale’s core business is building software that utilizes our data and compensation algorithm. The “Where it Pays to Attend College” dataset by the WSJ is based on PayScale data.
#
# ## *Dataset Distribution*
#
# #### How is the dataset distributed? (e.g., website, API, etc.; does the data have a DOI; is it archived redundantly?)
# The dataset can be downloaded from: http://online.wsj.com/public/resources/documents/info-Salaries_for_Colleges_by_Type-sort.html
# and from: https://www.kaggle.com/wsj/college-salaries
#
# #### When will the dataset be released/first distributed? (Is there a canonical paper/reference for this dataset?)
# The dataset was released on July 31, 2008 on the WSJ and was updated 2 years ago on Kaggle.
# #### What license (if any) is it distributed under? Are there any copyrights on the data?
# Copyrights on the data belong to PayScale.
#
# #### Are there any fees or access/export restrictions?
# None.
#
# ## *Data Maintenance*
#
# #### Who is supporting/hosting/maintaining the dataset? How does one contact the owner/curator/manager of the dataset (e.g. email address, or other contact info)?
# The dataset is hosted at the WSJ and all comments can be sent to <EMAIL>
#
# #### Will the dataset be updated? How often and by whom? How will updates/revisions be documented and communicated (e.g., mailing list, GitHub)? Is there an erratum?
# There is a newer dataset available on the PayScale’s website with fewer salary instances, but with the new ones like STEM variables and High Meaning. PayScale seems to produce new college report every year.
#
# #### If the dataset becomes obsolete how will this be communicated?
# Since PayScale’s whole business is built around this report they probably won’t ever claim it as obsolete. However, if more verified similar dataset will be available, this one will become useless.
#
# #### Is there a repository to link to any/all papers/systems that use this dataset?
# Unknown. (I could not find one)
#
# #### If others want to extend/augment/build on this dataset, is there a mechanism for them to do so? If so, is there a process for tracking/assessing the quality of those contributions. What is the process for communicating/distributing these contributions to users?
# Since the original collected data is not public and there is no way of accessing those contributions, the only way to build on this dataset is to use external data.
#
# ## *Legal and Ethical Considerations*
#
# #### If the dataset relates to people (e.g., their attributes) or was generated by people, were they informed about the data collection? (e.g., datasets that collect writing, photos, interactions, transactions, etc.)
# According to PayScale, people willingly self-reported the data.
# #### If it relates to other ethically protected subjects, have appropriate obligations been met? (e.g., medical data might include information collected from animals)
# Not applicable
# #### If it relates to people, were there any ethical review applications/reviews/approvals? (e.g. Institutional Review Board applications)
# Unknown
# #### If it relates to people, were they told what the dataset would be used for and did they consent? What community norms exist for data collected from human communications? If consent was obtained, how? Were the people provided with any mechanism to revoke their consent in the future or for certain uses?
# Since the data was self-reported it implies that people consent.
# #### If it relates to people, could this dataset expose people to harm or legal action? (e.g., financial social or otherwise) What was done to mitigate or reduce the potential for harm?
# There is a minimal risk because people provided consent when provided the data.
# #### If it relates to people, does it unfairly advantage or dis- advantage a particular social group? In what ways? How was this mitigated?
# Unknown
# #### If it relates to people, were they provided with privacy guarantees? If so, what guarantees and how are these ensured?
# Unknown
# #### Does the dataset comply with the EU General Data Protection Regulation (GDPR)? Does it comply with any other standards, such as the US Equal Employment Opportunity Act?
# Probably, https://www.payscale.com/compensation-today/2018/05/gdpr-commitments
# #### Does the dataset contain information that might be considered sensitive or confidential? (e.g., personally identifying information)
# Possibly yes.
# #### Does the dataset contain information that might be considered inappropriate or offensive?
# Probably no.
#
|
botasakhi/Datasheet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="RWPN3PCP-IwW" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204} outputId="3a6060e4-b54b-432d-90ef-1fd7438d17c9" executionInfo={"status": "ok", "timestamp": 1530084292943, "user_tz": -330, "elapsed": 4054, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
# !wget https://play.minio.io:9000/rao/creditcard.csv -P /tmp
# + id="gbG-0hh0AEtT" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
#Importing librairies
import pandas as pd
import numpy as np
# Scikit-learn library: For SVM
from sklearn import preprocessing
from sklearn.metrics import confusion_matrix
from sklearn import svm
import itertools
# Matplotlib library to plot the charts
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
# Library for the statistic data vizualisation
import seaborn
# %matplotlib inline
# + id="ppYEsnOvBWPu" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 346} outputId="c7445f0e-90c8-4691-e053-2ba30db708bc" executionInfo={"status": "ok", "timestamp": 1530084299544, "user_tz": -330, "elapsed": 4649, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
data = pd.read_csv('/tmp/creditcard.csv') # Reading the file .csv
df = pd.DataFrame(data) # Converting data to Panda DataFrame
df.describe()
# + id="zutF9XH0CNdh" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 648} outputId="1652a666-90cf-49b3-910f-3313032b2eab" executionInfo={"status": "ok", "timestamp": 1530084300764, "user_tz": -330, "elapsed": 1078, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
df_fraud = df[df['Class'] == 1] # Recovery of fraud data
plt.figure(figsize=(15,10))
plt.scatter(df_fraud['Time'], df_fraud['Amount']) # Display fraud amounts according to their time
plt.title('Scratter plot amount fraud')
plt.xlabel('Time')
plt.ylabel('Amount')
plt.xlim([0,175000])
plt.ylim([0,2500])
plt.show()
# + id="ygBc2E6-Cd6k" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="e91ccdc7-cd28-4f2b-cae1-d182300373c7" executionInfo={"status": "ok", "timestamp": 1530084302099, "user_tz": -330, "elapsed": 1221, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
nb_big_fraud = df_fraud[df_fraud['Amount'] > 1000].shape[0] # Recovery of frauds over 1000
print('There are only '+ str(nb_big_fraud) + ' frauds where the amount was bigger than 1000 over ' + str(df_fraud.shape[0]) + ' frauds')
# + id="jVk9305QCig6" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="fb6586e3-ed09-4073-936f-6ac3cafb394a" executionInfo={"status": "ok", "timestamp": 1530084303166, "user_tz": -330, "elapsed": 951, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
number_fraud = len(data[data.Class == 1])
number_no_fraud = len(data[data.Class == 0])
print('There are only '+ str(number_fraud) + ' frauds in the original dataset, even though there are ' + str(number_no_fraud) +' no frauds in the dataset.')
# + id="IKDqygwLDubk" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="d19a4105-daf0-43c3-91f5-d5a51487fe85" executionInfo={"status": "ok", "timestamp": 1530084304392, "user_tz": -330, "elapsed": 1093, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print("The accuracy of the classifier then would be : "+ str((284315-492)/284315)+ " which is the number of good classification over the number of tuple to classify")
# + id="yE5H0K3QDzjV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
df_corr = df.corr() # Calculation of the correlation coefficients in pairs, with the default method:
# Pearson, Standard Correlation Coefficient
# + id="fnyJ2QOqD4J7" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 673} outputId="0fbff38d-4362-429d-af40-ab51ed195034" executionInfo={"status": "ok", "timestamp": 1530084307628, "user_tz": -330, "elapsed": 1365, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
plt.figure(figsize=(15,10))
seaborn.heatmap(df_corr, cmap="YlGnBu") # Displaying the Heatmap
seaborn.set(font_scale=2,style='white')
plt.title('Heatmap correlation')
plt.show()
# + id="GMSFhwAHD9VY" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
rank = df_corr['Class'] # Retrieving the correlation coefficients per feature in relation to the feature class
df_rank = pd.DataFrame(rank)
df_rank = np.abs(df_rank).sort_values(by='Class',ascending=False) # Ranking the absolute values of the coefficients
# in descending order
df_rank.dropna(inplace=True) # Removing Missing Data (not a number)
# + id="ZkzvMuCAECWV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="6dfac083-0175-41c9-94ec-1eb75a88dc64" executionInfo={"status": "ok", "timestamp": 1530084309759, "user_tz": -330, "elapsed": 1046, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
# We seperate ours data in two groups : a train dataset and a test dataset
# First we build our train dataset
df_train_all = df[0:150000] # We cut in two the original dataset
df_train_1 = df_train_all[df_train_all['Class'] == 1] # We seperate the data which are the frauds and the no frauds
df_train_0 = df_train_all[df_train_all['Class'] == 0]
print('In this dataset, we have ' + str(len(df_train_1)) +" frauds so we need to take a similar number of non-fraud")
df_sample=df_train_0.sample(300)
df_train = df_train_1.append(df_sample) # We gather the frauds with the no frauds.
df_train = df_train.sample(frac=1) # Then we mix our dataset
# + id="_gonODlLEFtm" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
X_train = df_train.drop(['Time', 'Class'],axis=1) # We drop the features Time (useless), and the Class (label)
y_train = df_train['Class'] # We create our label
X_train = np.asarray(X_train)
y_train = np.asarray(y_train)
# + id="87HfN1W5EI_H" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
############################## with all the test dataset to see if the model learn correctly ##################
df_test_all = df[150000:]
X_test_all = df_test_all.drop(['Time', 'Class'],axis=1)
y_test_all = df_test_all['Class']
X_test_all = np.asarray(X_test_all)
y_test_all = np.asarray(y_test_all)
# + id="u4hGWTqYEMeX" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
X_train_rank = df_train[df_rank.index[1:11]] # We take the first ten ranked features
X_train_rank = np.asarray(X_train_rank)
# + id="CIMmKVoNEPco" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
############################## with all the test dataset to see if the model learn correctly ##################
X_test_all_rank = df_test_all[df_rank.index[1:11]]
X_test_all_rank = np.asarray(X_test_all_rank)
y_test_all = np.asarray(y_test_all)
# + id="XdwH6jleETAx" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
class_names=np.array(['0','1']) # Binary label, Class = 1 (fraud) and Class = 0 (no fraud)
# + id="5J7iSZt1EzeN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Function to plot the confusion Matrix
def plot_confusion_matrix(cm, classes,
title='Confusion matrix',
cmap=plt.cm.Blues):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# + id="gv62cf5tE7KV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
classifier = svm.SVC(kernel='linear') # We set a SVM classifier, the default SVM Classifier (Kernel = Radial Basis Function)
# + id="W_4Ljuc1E-lZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} outputId="32d1caec-fb0f-4d2b-f38b-36ac65bef07d" executionInfo={"status": "ok", "timestamp": 1530084323008, "user_tz": -330, "elapsed": 4591, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
classifier.fit(X_train, y_train) # Then we train our model, with our balanced data train.
# + id="xzPUwVNPFK35" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
prediction_SVM_all = classifier.predict(X_test_all) #And finally, we predict our data test.
# + id="wWrPvjaDFOhe" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 400} outputId="44a640d6-44a1-4030-9d9d-eb845cb165ce" executionInfo={"status": "ok", "timestamp": 1530084325745, "user_tz": -330, "elapsed": 1381, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
cm = confusion_matrix(y_test_all, prediction_SVM_all)
plot_confusion_matrix(cm,class_names)
# + id="qB8JYgaFFSQc" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="38e87e6b-ad3b-4455-c270-b1fb6724a88d" executionInfo={"status": "ok", "timestamp": 1530084327253, "user_tz": -330, "elapsed": 1396, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('Our criterion give a result of '
+ str( ( (cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1])) + 4 * cm[1][1]/(cm[1][0]+cm[1][1])) / 5))
# + id="Y_UV4WH1FVr-" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} outputId="f47a5d2f-5628-4539-8f0a-016d18755343" executionInfo={"status": "ok", "timestamp": 1530084328562, "user_tz": -330, "elapsed": 1194, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('We have detected ' + str(cm[1][1]) + ' frauds / ' + str(cm[1][1]+cm[1][0]) + ' total frauds.')
print('\nSo, the probability to detect a fraud is ' + str(cm[1][1]/(cm[1][1]+cm[1][0])))
print("the accuracy is : "+str((cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1]))))
# + id="YJ1XQahSFZGX" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
classifier.fit(X_train_rank, y_train) # Then we train our model, with our balanced data train.
prediction_SVM = classifier.predict(X_test_all_rank) #And finally, we predict our data test.
# + id="cQC7qxglFcmE" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 400} outputId="e3885dea-e183-46d1-9fd9-69e4fd8026bf" executionInfo={"status": "ok", "timestamp": 1530084331675, "user_tz": -330, "elapsed": 1907, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
cm = confusion_matrix(y_test_all, prediction_SVM)
plot_confusion_matrix(cm,class_names)
# + id="OJiwHNV8Ff4_" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="f7004f95-ec27-46c3-b410-5ee4f6ae8843" executionInfo={"status": "ok", "timestamp": 1530084332433, "user_tz": -330, "elapsed": 645, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('Our criterion give a result of '
+ str( ( (cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1])) + 4 * cm[1][1]/(cm[1][0]+cm[1][1])) / 5))
# + id="lfb_akz7FjCQ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} outputId="bfc2bf65-6287-42d4-fa96-083f9c4cf9b2" executionInfo={"status": "ok", "timestamp": 1530084334006, "user_tz": -330, "elapsed": 1426, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('We have detected ' + str(cm[1][1]) + ' frauds / ' + str(cm[1][1]+cm[1][0]) + ' total frauds.')
print('\nSo, the probability to detect a fraud is ' + str(cm[1][1]/(cm[1][1]+cm[1][0])))
print("the accuracy is : "+str((cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1]))))
# + id="mx5WudfCFmbm" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
classifier_b = svm.SVC(kernel='linear',class_weight={0:0.60, 1:0.40})
# + id="knUidEq4F51A" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} outputId="2fd94ca2-8a0d-4514-8697-dc1b4b863080" executionInfo={"status": "ok", "timestamp": 1530084340261, "user_tz": -330, "elapsed": 4743, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
classifier_b.fit(X_train, y_train) # Then we train our model, with our balanced data train.
# + id="XG8eSEFfGBnD" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
prediction_SVM_b_all = classifier_b.predict(X_test_all) #We predict all the data set.
# + id="BMfvXbw8GE5K" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 400} outputId="456c71eb-b5d1-4e27-da10-4343822bdc50" executionInfo={"status": "ok", "timestamp": 1530084342646, "user_tz": -330, "elapsed": 1173, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
cm = confusion_matrix(y_test_all, prediction_SVM_b_all)
plot_confusion_matrix(cm,class_names)
# + id="ZuBUnLH5GINT" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="b7149807-9ace-4c1c-bfc6-391a27c068b2" executionInfo={"status": "ok", "timestamp": 1530084343700, "user_tz": -330, "elapsed": 913, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('Our criterion give a result of '
+ str( ( (cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1])) + 4 * cm[1][1]/(cm[1][0]+cm[1][1])) / 5))
# + id="Nj1HGCWTGMTw" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} outputId="f5a16c7d-d4bf-4432-bbff-05b9f5cf48de" executionInfo={"status": "ok", "timestamp": 1530084344998, "user_tz": -330, "elapsed": 1188, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('We have detected ' + str(cm[1][1]) + ' frauds / ' + str(cm[1][1]+cm[1][0]) + ' total frauds.')
print('\nSo, the probability to detect a fraud is ' + str(cm[1][1]/(cm[1][1]+cm[1][0])))
print("the accuracy is : "+str((cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1]))))
# + id="UI2qjvRQGPmE" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
classifier_b.fit(X_train_rank, y_train) # Then we train our model, with our balanced data train.
prediction_SVM = classifier_b.predict(X_test_all_rank) #And finally, we predict our data test.
# + id="O70X3SVQGS9h" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 400} outputId="fabc9585-49a2-495f-e8a1-e28ae2c27b83" executionInfo={"status": "ok", "timestamp": 1530084347664, "user_tz": -330, "elapsed": 1501, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
cm = confusion_matrix(y_test_all, prediction_SVM)
plot_confusion_matrix(cm,class_names)
# + id="vdHHhadOGWZO" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="b7e607d7-3deb-4e8c-9225-36d3133a3a7b" executionInfo={"status": "ok", "timestamp": 1530084348853, "user_tz": -330, "elapsed": 1075, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('Our criterion give a result of '
+ str( ( (cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1])) + 4 * cm[1][1]/(cm[1][0]+cm[1][1])) / 5))
# + id="8n6XxmvUGZsN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} outputId="21b221ff-a9b3-4080-8b12-92432d40f8b5" executionInfo={"status": "ok", "timestamp": 1530084350216, "user_tz": -330, "elapsed": 1252, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-ocaYyjrX3Z4/AAAAAAAAAAI/AAAAAAAAABQ/cPl8U19wmMg/s50-c-k-no/photo.jpg", "userId": "115904709554351828058"}}
print('We have detected ' + str(cm[1][1]) + ' frauds / ' + str(cm[1][1]+cm[1][0]) + ' total frauds.')
print('\nSo, the probability to detect a fraud is ' + str(cm[1][1]/(cm[1][1]+cm[1][0])))
print("the accuracy is : "+str((cm[0][0]+cm[1][1]) / (sum(cm[0]) + sum(cm[1]))))
|
Machine_Learning/SVM/Credit_Card_Fraud_Detection/Credit_Card_Fraud_Detection_Using_SVM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mass Detection training
#
# +
import os
import sys
import itertools
import math
import logging
import json
import re
import random
import time
import concurrent.futures
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.lines as lines
from matplotlib.patches import Polygon
import imgaug
from imgaug import augmenters as iaa
# Root directory of the project
ROOT_DIR = os.getcwd()
print(ROOT_DIR)
if ROOT_DIR.endswith("mammography"):
# Go up one level to the repo root
ROOT_DIR = os.path.dirname(ROOT_DIR)
print(ROOT_DIR)
# Import Mask RCNN
sys.path.append(ROOT_DIR)
from mrcnn import utils
from mrcnn import visualize
from mrcnn.visualize import display_images
from mrcnn import model as modellib
from mrcnn.model import log
import mammo_baseline
# %matplotlib inline
# -
# Comment out to reload imported modules if they change
# %load_ext autoreload
# %autoreload 2
# ## Configurations
# +
# Dataset directory
DATASET_DIR = os.path.join(ROOT_DIR, "datasets/mammo")
# Load dataset
subset = "mass_train_3x"
if "3x" in subset:
augmented=True
else:
augmented=False
dataset_train = mammo_baseline.MammoDataset()
dataset_train.load_mammo(DATASET_DIR, subset=subset, augmented=augmented)
# Must call before using the dataset
dataset_train.prepare()
print("Image Count: {}".format(len(dataset_train.image_ids)))
print("Class Count: {}".format(dataset_train.num_classes))
for i, info in enumerate(dataset_train.class_info):
print("{:3}. {:50}".format(i, info['name']))
# Load validation dataset
dataset_val = mammo_baseline.MammoDataset()
dataset_val.load_mammo(DATASET_DIR, "val", augmented=augmented)
dataset_val.prepare()
print("Images: {}\nClasses: {}".format(len(dataset_val.image_ids), dataset_val.class_names))
# -
# ## Notebook Preferences
def get_ax(rows=1, cols=1, size=16):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Adjust the size attribute to control how big to render images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
# +
#######################################
# Training with higher resolution #
# images. Max_dim = 1408 #
# Training set size = 4x #
# Augmentation = FlipLR and FlipUD #
#######################################
# Configurations
# Use configuation from mammo.py, but override
# image resizing so we see the real sizes here
class NoResizeConfig(mammo.MammoConfig):
BACKBONE = "resnet101"
# Adjust depending on your GPU memory
IMAGES_PER_GPU = 1
IMAGE_MAX_DIM = 1408
# Number of classes (including background)
NUM_CLASSES = 1 + 1 # Background + mass
# Number of training and validation steps per epoch
STEPS_PER_EPOCH = (len(dataset_train.image_ids)) // IMAGES_PER_GPU
VALIDATION_STEPS = max(1, len(dataset_val.image_ids) // IMAGES_PER_GPU)
MINI_MASK_SHAPE = (56, 56) # (height, width) of the mini-mask
RPN_ANCHOR_SCALES = (32, 64, 128, 256, 512)
RPN_TRAIN_ANCHORS_PER_IMAGE = 512
RPN_ANCHOR_RATIOS = [0.5, 1, 2]
# ROIs kept after non-maximum supression (training and inference)
POST_NMS_ROIS_TRAINING = 2000
POST_NMS_ROIS_INFERENCE = 1000
# Non-max suppression threshold to filter RPN proposals.
# You can increase this during training to generate more propsals.
RPN_NMS_THRESHOLD = 0.7
config = NoResizeConfig()
config.display()
MODEL_DIR = 'checkpoints'
# Create model
model = modellib.MaskRCNN(mode="training", config=config,
model_dir=MODEL_DIR)
# Select weights file to load
weights_path = model.get_imagenet_weights()
# Load weights
print("Loading weights ", weights_path)
model.load_weights(weights_path, by_name=True)
model.train(dataset_train, dataset_val, config.LEARNING_RATE, epochs=9, layers='all')
|
mammography/Mass_detection_Mask_RCNN_training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from collections import defaultdict, deque
# ## algorithm
def knowledge_base(formulas):
rules, variable, dependency = [], defaultdict(bool), defaultdict(list)
def _clause(formula):
# A, B, C => P
neg, pos = formula.replace(' ', '').split('=>')
neg, pos = set(neg.split('&')) - {''}, pos or None
# add rule
rules.append((neg, pos))
# set variable and track dependencies
for i in neg:
dependency[i].append((neg, pos))
# parse formulas and build knowledge base
deque((_clause(i) for i in formulas.split('\n') if i), 0)
return rules, variable, dependency
def resolution(rules, variable, dependency):
# initial variables that have to be satisfied
to_satisfy = [(neg, pos) for neg, pos in rules if not neg]
while to_satisfy:
neg, pos = to_satisfy.pop()
# contradiction: true => false
if not pos:
return False
# satisfy variable
variable[pos] = True
# update dependent rules
for d_neg, d_pos in dependency[pos]:
d_neg.remove(pos)
# next variable to be satisfied
if not d_neg and d_pos not in variable:
to_satisfy.append((d_neg, d_pos))
return True
def hornsat(formulas):
rules, variable, dependency = knowledge_base(formulas)
satisfiable = resolution(rules, variable, dependency)
print(['CONTRADICTION', 'SATISFIABLE'][satisfiable])
print(', '.join('%s=%s' % i for i in variable.items()))
# ## run
hornsat("""
X => Y
Y => Z
=> X
""")
hornsat("""
X => Y
Y => X
=> X
Y =>
""")
hornsat("""
P & Q & R & S => X
P & Q => R
R => S
X =>
=> P
=> R
""")
hornsat("""
P & Q & R & S => X
P & Q => R
R => S
X =>
=> P
=> Q
""")
|
100days/day 78 - horn-satifiability.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Generate points for the cases tab
#
# This notebook generates the points GeoJSON for the points on the Cases tab of the dashboard. It pulls the point data from JHU confirmed cases. It also produces some intermediate outputs, such as mappings to the feature IDs, for use in the case data processing notebook that runs as part of the data update pipeline.
#
# ### Papermill
# + tags=["parameters"]
# parameters
data_dir = '/opt/src/data'
# -
# For papermill execution, the pameters are:
# - data_dir: That data directory to read data from and publish data to.
# +
import json
import io
import os
from datetime import datetime
import requests
import numpy as np
import pandas as pd
import geopandas as gpd
from shapely.geometry import Point, shape, mapping
from slugify import slugify
# +
def get_code(admin0, admin1=None, admin2=None):
slug_txt = admin0
if admin1 is not None:
slug_txt = "{} {}".format(admin1, slug_txt)
if admin2 is not None:
slug_txt = "{} {}".format(admin2, slug_txt)
return slugify(slug_txt)
def fetch_df(url):
"""Fetches a Pandas DataFrame from a remote source"""
r = requests.get(url)
return pd.read_csv(io.BytesIO(r.content))
# -
cases_df = fetch_df('https://github.com/CSSEGISandData/COVID-19/raw/master/'
'csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv')
# +
us_territories = [
'American Samoa',
'Guam',
'Northern Mariana Islands',
'Puerto Rico',
'Virgin Islands'
]
def filter_us(df):
# Filter out counties that have 0 latest data.
filtered_df = df[df.iloc[:,-1] != 0]
filtered_df = filtered_df[
(filtered_df['Province_State'].isin(us_territories)) |
(
(~filtered_df['Lat'].isnull()) &
(filtered_df['Lat'] != 0.0) &
(~filtered_df['FIPS'].isnull())
)
]
return filtered_df
us_cases_df = fetch_df('https://github.com/CSSEGISandData/COVID-19/raw/master/'
'csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_US.csv')
us_cases_df = filter_us(us_cases_df)
# -
countries_gdf = gpd.read_file(os.path.join(data_dir, 'published/countries.geojson'))
# +
country_data = {}
case_features = []
codes_to_id = {}
codes_to_alpha2 = {}
current_id = 0
def add_country_data(country):
bounds = country['geometry'].bounds
bounds = [[bounds[0], bounds[1]], [bounds[2], bounds[3]]]
if country['ADM0_A3'] is None or type(country['ADM0_A3']) is float:
raise Exception('ADM0_A3 is None or nan for {}'.format(country['ADMIN']))
if country['ISO_A2'] is None or type(country['ISO_A2']) is float:
raise Exception('ISO_A2 is None or nan for {}'.format(country['ADMIN']))
country_data[code] = {
'name': country['ADMIN'],
'alpha3': country['ADM0_A3'],
'alpha2': country['ISO_A2'],
'bounds': bounds
}
for _, row in cases_df.sort_values(by=['Country/Region', 'Province/State']).iterrows():
name = row['Country/Region']
region_name = None if type(row['Province/State']) is float else row['Province/State']
if name == 'Congo (Brazzaville)':
# Fix the lat/lng of Congo, which is the same location
# the Democratic Republic of Congo in the JHU data.
lat, lon = -1.402385, 15.405892
else:
lat, lon = row['Lat'], row['Long']
pt = Point(lon, lat)
if (lon, lat) == (0, 0):
print('Skipping {}'.format(get_code(name, region_name)))
else:
matching_countries = countries_gdf[countries_gdf['geometry'].contains(pt)]
if len(matching_countries) < 1:
if name == 'Saint Vincent and the Grenadines':
matching_countries = countries_gdf[
countries_gdf['NAME'] == 'St. Vin. and Gren.'
]
else:
matching_countries = countries_gdf[
countries_gdf['NAME'] == name
]
if len(matching_countries) < 1:
print(row['Country/Region'])
print(' Not found: {}'.format(pt))
else:
country = matching_countries.iloc[0]
code = get_code(name, region_name)
point_id = current_id
current_id += 1
if region_name is not None:
display_name = '{}, {}'.format(region_name, country['ADMIN'])
else:
display_name = country['ADMIN']
case_features.append({
'id': point_id,
'type': 'Feature',
'geometry': mapping(pt),
'properties': {
'displayName': display_name,
'code': code,
'id': point_id
}
})
codes_to_id[code] = point_id
codes_to_alpha2[code] = country['ISO_A2']
# Process countries
if region_name is None:
add_country_data(country)
SKIP_COUNTIES = True
for _, row in us_cases_df.sort_values(by=['Country_Region', 'Province_State', 'Admin2']).iterrows():
region_name = row['Province_State']
county_name = None if type(row['Admin2']) is float else row['Admin2']
if county_name is not None and (
county_name.startswith('Out of') or
county_name == 'Unassigned'):
print('Skipping {}, {}'.format(county_name, region_name))
continue
lat, lon = row['Lat'], row['Long_']
pt = Point(lon, lat)
if (lon, lat) == (0, 0):
print('Skipping {}'.format(get_code('US', region_name, county_name)))
else:
code = get_code('US', admin1=region_name, admin2=county_name)
point_id = current_id
current_id += 1
display_name = '{}, US'
if county_name is not None:
display_name = '{}, {}, US'.format(county_name, region_name)
else:
display_name = '{}, US'.format(region_name)
if SKIP_COUNTIES and county_name is not None:
pass
else:
case_features.append({
'id': point_id,
'type': 'Feature',
'geometry': mapping(pt),
'properties': {
'displayName': display_name,
'code': code,
'id': point_id
}
})
codes_to_id[code] = point_id
codes_to_alpha2[code] = 'US'
# -
# Handle adding country information for countries that only have regions in the JHU data.
for _, row in countries_gdf[countries_gdf['ADMIN'].isin(['Australia', 'Canada', 'China'])].iterrows():
add_country_data(row)
codes_to_alpha2[get_code(row['ADMIN'])] = row['ISO_A2']
with open(os.path.join(data_dir, 'published/case-points.geojson'), 'w') as f:
f.write(json.dumps({
'type': 'FeatureCollection',
'features': case_features
}, sort_keys=True))
with open(os.path.join(data_dir, 'case-codes-to-ids-intermidiate.json'), 'w') as f:
f.write(json.dumps(codes_to_id, sort_keys=True))
with open(os.path.join(data_dir, 'case-country-config.json'), 'w') as f:
f.write(json.dumps(country_data, sort_keys=True))
with open(os.path.join(data_dir, 'case-codes-to-alpha2.json'), 'w') as f:
f.write(json.dumps(codes_to_alpha2, sort_keys=True))
|
data-processing/notebooks/generate_points_from_JHU.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task 3: preprocessing
# +
# Imports
import re
import string
import json
from datetime import datetime
from collections import defaultdict, Counter
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import tqdm
import torch
import torch.nn as nn
from torch.nn import Module
import torch.optim as optim
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from torch.nn.utils.rnn import pad_sequence, pack_padded_sequence, pad_packed_sequence
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score
from nltk.corpus import stopwords
device = 'cpu'
import random
random.seed(26)
np.random.seed(62)
torch.manual_seed(2021)
torch.cuda.manual_seed(123)
# -
# # Load Bengali datasets
# save sample Bengali datasets
bengali_train_df = pd.read_csv('../../Task_2/hindi_bengali/save/bengali_hatespeech_sample_train.csv')
bengali_test_df = pd.read_csv('../../Task_2/hindi_bengali/save/bengali_hatespeech_sample_test.csv')
bengali_other_df = pd.read_csv('../../Task_2/hindi_bengali/save/bengali_hatespeech_other.csv')
# # Preprocessing
train_sentences = bengali_train_df['sentence']
test_sentences = bengali_test_df['sentence']
other_sentences = bengali_other_df['sentence']
# +
# remove user taggings
user_tag_pattern = re.compile(r'\@\w*')
def remove_tagging(sentence):
return re.sub(user_tag_pattern, ' ', sentence)
# remove punctuations and urls
http_re = re.compile('http://[^ ]*')
https_re = re.compile('https://[^ ]*')
punctuation = string.punctuation[:2] + string.punctuation[3:]
translator = str.maketrans(punctuation, ' '*len(punctuation))
def remove_punc_and_urls(s):
s = re.sub(http_re, ' ', s)
s = re.sub(https_re, ' ', s)
s = s.translate(translator)
return s
# substitute numbers
# when there is a number in the string:
# if that number is 0 or 1 or 2, then there is no change.
# else, that number is substituted by a word describing how many digits it has.
def substitute_number(x):
x = x.group(0)
if x in {'0', '1', '2'}:
return x
return '{}_digits_number'.format(len(x))
# stopwords BENGALI (source: https://www.ranks.nl/stopwords/bengali)
stopwords = ['অবশ্য', 'অনেক', 'অনেকে', 'অনেকেই', 'অন্তত', 'অথবা', 'অথচ', 'অর্থাত', 'অন্য', 'আজ', 'আছে', 'আপনার',
'আপনি', 'আবার', 'আমরা', 'আমাকে', 'আমাদের', 'আমার', 'আমি', 'আরও', 'আর', 'আগে', 'আগেই', 'আই',
'অতএব', 'আগামী', 'অবধি', 'অনুযায়ী', 'আদ্যভাগে', 'এই', 'একই', 'একে', 'একটি', 'এখন', 'এখনও', 'এখানে',
'এখানেই', 'এটি', 'এটা', 'এটাই', 'এতটাই', 'এবং', 'একবার', 'এবার', 'এদের', 'এঁদের', 'এমন', 'এমনকী', 'এল',
'এর', 'এরা', 'এঁরা', 'এস', 'এত', 'এতে', 'এসে', 'একে', 'এ', 'ঐ', ' ই', 'ইহা', 'ইত্যাদি', 'উনি', 'উপর',
'উপরে', 'উচিত', 'ও', 'ওই', 'ওর', 'ওরা', 'ওঁর', 'ওঁরা', 'ওকে', 'ওদের', 'ওঁদের', 'ওখানে', 'কত', 'কবে',
'করতে', 'কয়েক', 'কয়েকটি', 'করবে', 'করলেন', 'করার', 'কারও', 'করা', 'করি', 'করিয়ে', 'করার', 'করাই',
'করলে', 'করলেন', 'করিতে', 'করিয়া', 'করেছিলেন', 'করছে', 'করছেন', 'করেছেন', 'করেছে', 'করেন', 'করবেন',
'করায়', 'করে', 'করেই', 'কাছ', 'কাছে', 'কাজে', 'কারণ', 'কিছু', 'কিছুই', 'কিন্তু', 'কিংবা', 'কি', 'কী', 'কেউ',
'কেউই', 'কাউকে', 'কেন', 'কে', 'কোনও', 'কোনো', 'কোন', 'কখনও', 'ক্ষেত্রে', 'খুব গুলি', 'গিয়ে', 'গিয়েছে',
'গেছে', 'গেল', 'গেলে', 'গোটা', 'চলে', 'ছাড়া', 'ছাড়াও', 'ছিলেন', 'ছিল', 'জন্য', 'জানা', 'ঠিক', 'তিনি',
'তিনঐ', 'তিনিও', 'তখন', 'তবে', 'তবু', 'তাঁদের', 'তাঁাহারা', 'তাঁরা', 'তাঁর', 'তাঁকে', 'তাই', 'তেমন', 'তাকে',
'তাহা', 'তাহাতে', 'তাহার', 'তাদের', 'তারপর', 'তারা', 'তারৈ', 'তার', 'তাহলে', 'তিনি', 'তা', 'তাও', 'তাতে',
'তো', 'তত', 'তুমি', 'তোমার', 'তথা', 'থাকে', 'থাকা', 'থাকায়', 'থেকে', 'থেকেও', 'থাকবে', 'থাকেন', 'থাকবেন',
'থেকেই', 'দিকে', 'দিতে', 'দিয়ে', 'দিয়েছে', 'দিয়েছেন', 'দিলেন', 'দু', 'দুটি', 'দুটো', 'দেয়', 'দেওয়া', 'দেওয়ার',
'দেখা', 'দেখে', 'দেখতে', 'দ্বারা', 'ধরে', 'ধরা', 'নয়', 'নানা', 'না', 'নাকি', 'নাগাদ', 'নিতে', 'নিজে', 'নিজেই',
'নিজের', 'নিজেদের', 'নিয়ে', 'নেওয়া', 'নেওয়ার', 'নেই', 'নাই', 'পক্ষে', 'পর্যন্ত', 'পাওয়া', 'পারেন', 'পারি', 'পারে',
'পরে', 'পরেই', 'পরেও', 'পর', 'পেয়ে', 'প্রতি', 'প্রভৃতি', 'প্রায়', 'ফের', 'ফলে', 'ফিরে', 'ব্যবহার', 'বলতে',
'বললেন', 'বলেছেন', 'বলল', 'বলা', 'বলেন', 'বলে', 'বহু', 'বসে', 'বার', 'বা', 'বিনা', 'বরং', 'বদলে', 'বাদে',
'বার', 'বিশেষ', 'বিভিন্ন বিষয়টি', 'ব্যবহার', 'ব্যাপারে', 'ভাবে', 'ভাবেই', 'মধ্যে', 'মধ্যেই', 'মধ্যেও', 'মধ্যভাগে',
'মাধ্যমে', 'মাত্র', 'মতো', 'মতোই', 'মোটেই', 'যখন', 'যদি', 'যদিও', 'যাবে', 'যায়', 'যাকে', 'যাওয়া', 'যাওয়ার',
'যত', 'যতটা', 'যা', 'যার', 'যারা', 'যাঁর', 'যাঁরা', 'যাদের', 'যান', 'যাচ্ছে', 'যেতে', 'যাতে', 'যেন', 'যেমন',
'যেখানে', 'যিনি', 'যে', 'রেখে', 'রাখা', 'রয়েছে', 'রকম', 'শুধু', 'সঙ্গে', 'সঙ্গেও', 'সমস্ত', 'সব', 'সবার', 'সহ',
'সুতরাং', 'সহিত', 'সেই', 'সেটা', 'সেটি', 'সেটাই', 'সেটাও', 'সম্প্রতি', 'সেখান', 'সেখানে', 'সে', 'স্পষ্ট', 'স্বয়ং',
'হইতে', 'হইবে', 'হৈলে', 'হইয়া', 'হচ্ছে', 'হত', 'হতে', 'হতেই', 'হবে', 'হবেন', 'হয়েছিল', 'হয়েছে', 'হয়েছেন', 'হয়ে',
'হয়নি', 'হয়', 'হয়েই', 'হয়তো', 'হল', 'হলে', 'হলেই', 'হলেও', 'হলো', 'হিসাবে', 'হওয়া', 'হওয়ার', 'হওয়ায়', 'হন',
'হোক', 'জন', 'জনকে', 'জনের', 'জানতে', 'জানায়', 'জানিয়ে', 'জানানো', 'জানিয়েছে', 'জন্য', 'জন্যওজে', 'জে',
'বেশ', 'দেন', 'তুলে', 'ছিলেন', 'চান', 'চায়', 'চেয়ে', 'মোট', 'যথেষ্ট', 'টি']
# -
def clean_texts(sentences):
# tags
sentences = [remove_tagging(sentence) for sentence in sentences]
# lower case
sentences = [sentence.lower() for sentence in sentences]
# remove punctuations and urls
sentences = [remove_punc_and_urls(sentence) for sentence in sentences]
# substitute numbers
sentences = [re.sub('\\b[0-9]+\\b', substitute_number, sentence) for sentence in sentences]
# remove stopwords
sentences = [[word for word in sentence.split() if word not in stopwords] for sentence in sentences]
return sentences
# +
# perform cleaning
train_sentences = clean_texts(train_sentences)
train_texts = [' '.join(l) for l in train_sentences]
bengali_train_df['sentence'] = train_texts
test_sentences = clean_texts(test_sentences)
test_texts = [' '.join(l) for l in test_sentences]
bengali_test_df['sentence'] = test_texts
other_sentences = clean_texts(other_sentences)
other_texts = [' '.join(l) for l in other_sentences]
bengali_other_df['sentence'] = other_texts
# -
print('train:')
display(bengali_train_df.head())
print('test:')
display(bengali_test_df.head())
print('other:')
display(bengali_other_df.head())
# ## Vocab and Word <-> int transformation
embed_train_df = pd.concat([bengali_train_df, bengali_other_df])
# +
train_sentences = [sentence.split() for sentence in bengali_train_df['sentence']]
test_sentences = [sentence.split() for sentence in bengali_test_df['sentence']]
embed_train_sentences = [sentence.split() for sentence in embed_train_df['sentence']]
flattened_words = [word for sentence in embed_train_sentences for word in sentence]
V = sorted(list(set(flattened_words)))
vocab_size = len(V)
print(f'vocab_size: {vocab_size}')
word_to_int = {}
int_to_word = {}
for i, word in enumerate(V):
word_to_int[word] = i
int_to_word[i] = word
# save dicts for transformation word <-> int
with open('save/word_to_int_dict.json', 'w') as f:
json.dump(word_to_int, f)
with open('save/int_to_word_dict.json', 'w') as f:
json.dump(int_to_word, f)
# save word-counter for sampling
word_counter = Counter(flattened_words)
with open('save/word_counter.json', 'w') as f:
json.dump(word_counter, f)
# -
# ## Save
bengali_train_df.to_csv('save/bengali_hatespeech_sample_train_preprocessed.csv', index=False)
bengali_test_df.to_csv('save/bengali_hatespeech_sample_test_preprocessed.csv', index=False)
bengali_other_df.to_csv('save/bengali_hatespeech_other_preprocessed.csv', index=False)
embed_train_df.to_csv('save/bengali_hatespeech_embed_train_preprocessed.csv', index=False)
|
Task_3/bengali_bengali/Task-3c.1_bengali_preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import re
import string
import nltk
from nltk.stem import PorterStemmer, WordNetLemmatizer
from nltk.corpus import stopwords
from nltk.tokenize import sent_tokenize, word_tokenize
from sklearn.feature_extraction.text import CountVectorizer
from fuzzywuzzy import fuzz
import matplotlib.pyplot as plt
import scipy.stats as st
nltk.download('punkt')
nltk.download('stopwords')
nltk.download('wordnet')
# -
# A. Using the **McDonalds Yelp Review CSV file**, **process the reviews**.
# This means you should think briefly about:
# * what stopwords to remove (should you add any custom stopwords to the set? Remove any stopwords?)
# * what regex cleaning you may need to perform (for example, are there different ways of saying `hamburger` that you need to account for?)
# * stemming/lemmatization (explain in your notebook why you used stemming versus lemmatization).
#
# Next, **count-vectorize the dataset**. Use the **`sklearn.feature_extraction.text.CountVectorizer`** examples from `Linear Algebra, Distance and Similarity (Completed).ipynb` and `Text Preprocessing Techniques (Completed).ipynb` (read the last section, `Vectorization Techniques`).
#
# I do not want redundant features - for instance, I do not want `hamburgers` and `hamburger` to be two distinct columns in your document-term matrix. Therefore, I'll be taking a look to make sure you've properly performed your cleaning, stopword removal, etc. to reduce the number of dimensions in your dataset.
stopword_list = stopwords.words('english')
negative_review_df = pd.read_csv('mcdonalds-yelp-negative-reviews.csv', encoding = 'latin-1')
negative_review_df['lower_case'] = negative_review_df['review'].str.lower()
negative_review_df['timestamp'] = negative_review_df['lower_case'].str.replace(
r'(?:[0-1][0-9]:[0-5][0-9])|(?:[0-1]?[0-9]?:?[0-5]?[0-9](?:ish)?\s?(?:am|pm))','TIMESTAMP_TOKEN')
negative_review_df['stopword'] = negative_review_df['timestamp'].str.replace(
r'\b('+'|'.join(stopword_list)+r')\b','')
negative_review_df['word_list'] = negative_review_df['stopword'].apply(word_tokenize)
punctuation_list = set(negative_review_df['stopword'].str.findall(r'['+string.punctuation+r']+').explode())
stemmer = PorterStemmer()
negative_review_df['stem'] = negative_review_df['word_list'].apply(lambda x: [stemmer.stem(word) for word in x if word not in punctuation_list])
negative_review_df['join'] = negative_review_df['stem'].apply(lambda x: ' '.join(x))
negative_review_df
vectorizer = CountVectorizer(min_df=2)
X = vectorizer.fit_transform(negative_review_df['join'])
X = X.toarray()
corpus_df = pd.DataFrame(X, columns=vectorizer.get_feature_names())
corpus_df
# **Answer:**
#
# 1. Read the data and lowercase
# 2. Replace timestamp value by Regular Expression
# 3. Stopword removal
# 4. Stemming because I want to lower the number of features more.
# 5. Vectorization
# B. **Stopwords, Stemming, Lemmatization Practice**
#
# Using the `tale-of-two-cities.txt` file from Week 1:
# * Count-vectorize the corpus. Treat each sentence as a document.
#
# How many features (dimensions) do you get when you:
# * Perform **stemming and then count-vectorization
# * Perform **lemmatization** and then **count-vectorization**.
# * Perform **lemmatization**, remove **stopwords**, and then perform **count-vectorization**?
text = open('tale-of-two-cities.txt', "r", encoding='utf8').read().replace('\n',' ')
sent_text = nltk.sent_tokenize(text) # this gives us a list of sentences
word_list_sent = [word_tokenize(sent) for sent in sent_text]
stemmer = PorterStemmer()
stem_only = []
for sent in word_list_sent:
stem_only.append([stemmer.stem(word) for word in sent])
lemmatizer = WordNetLemmatizer()
lemma_only = []
for sent in word_list_sent:
lemma_only.append([lemmatizer.lemmatize(word) for word in sent])
stopword_list = stopwords.words('english')
lemma_stop = []
for sent in word_list_sent:
lemma_stop.append([lemmatizer.lemmatize(word) for word in sent if word not in stopword_list])
def get_num_features(word_list_sent):
vectorizer = CountVectorizer()
X = vectorizer.fit_transform([' '.join(sent) for sent in word_list_sent])
return len(vectorizer.get_feature_names())
get_num_features(stem_only)
get_num_features(lemma_only)
get_num_features(lemma_stop)
# **Answer: As we can see here, stemming have the lowest number of features, while lemmatization have much more. Also removing stopword will decrease the number a little bit**
|
week2/Yixin_Ouang_HW2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/AI4Finance-LLC/FinRL-Library/blob/master/FinRL_multiple_stock_trading.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="gXaoZs2lh1hi"
# # Deep Reinforcement Learning for Stock Trading from Scratch: Multiple Stock Trading
#
# Tutorials to use OpenAI DRL to trade multiple stocks in one Jupyter Notebook | Presented at NeurIPS 2020: Deep RL Workshop
#
# * This blog is based on our paper: FinRL: A Deep Reinforcement Learning Library for Automated Stock Trading in Quantitative Finance, presented at NeurIPS 2020: Deep RL Workshop.
# * Check out medium blog for detailed explanations: https://towardsdatascience.com/finrl-for-quantitative-finance-tutorial-for-multiple-stock-trading-7b00763b7530
# * Please report any issues to our Github: https://github.com/AI4Finance-LLC/FinRL-Library/issues
# * **Pytorch Version**
#
#
# + [markdown] id="lGunVt8oLCVS"
# # Content
# + [markdown] id="HOzAKQ-SLGX6"
# * [1. Problem Definition](#0)
# * [2. Getting Started - Load Python packages](#1)
# * [2.1. Install Packages](#1.1)
# * [2.2. Check Additional Packages](#1.2)
# * [2.3. Import Packages](#1.3)
# * [2.4. Create Folders](#1.4)
# * [3. Download Data](#2)
# * [4. Preprocess Data](#3)
# * [4.1. Technical Indicators](#3.1)
# * [4.2. Perform Feature Engineering](#3.2)
# * [5.Build Environment](#4)
# * [5.1. Training & Trade Data Split](#4.1)
# * [5.2. User-defined Environment](#4.2)
# * [5.3. Initialize Environment](#4.3)
# * [6.Implement DRL Algorithms](#5)
# * [7.Backtesting Performance](#6)
# * [7.1. BackTestStats](#6.1)
# * [7.2. BackTestPlot](#6.2)
# * [7.3. Baseline Stats](#6.3)
# * [7.3. Compare to Stock Market Index](#6.4)
# + [markdown] id="sApkDlD9LIZv"
# <a id='0'></a>
# # Part 1. Problem Definition
# + [markdown] id="HjLD2TZSLKZ-"
# This problem is to design an automated trading solution for single stock trading. We model the stock trading process as a Markov Decision Process (MDP). We then formulate our trading goal as a maximization problem.
#
# The algorithm is trained using Deep Reinforcement Learning (DRL) algorithms and the components of the reinforcement learning environment are:
#
#
# * Action: The action space describes the allowed actions that the agent interacts with the
# environment. Normally, a ∈ A includes three actions: a ∈ {−1, 0, 1}, where −1, 0, 1 represent
# selling, holding, and buying one stock. Also, an action can be carried upon multiple shares. We use
# an action space {−k, ..., −1, 0, 1, ..., k}, where k denotes the number of shares. For example, "Buy
# 10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or −10, respectively
#
# * Reward function: r(s, a, s′) is the incentive mechanism for an agent to learn a better action. The change of the portfolio value when action a is taken at state s and arriving at new state s', i.e., r(s, a, s′) = v′ − v, where v′ and v represent the portfolio
# values at state s′ and s, respectively
#
# * State: The state space describes the observations that the agent receives from the environment. Just as a human trader needs to analyze various information before executing a trade, so
# our trading agent observes many different features to better learn in an interactive environment.
#
# * Environment: Dow 30 consituents
#
#
# The data of the single stock that we will be using for this case study is obtained from Yahoo Finance API. The data contains Open-High-Low-Close price and volume.
#
# + [markdown] id="Ffsre789LY08"
# <a id='1'></a>
# # Part 2. Getting Started- ASSUMES USING DOCKER, see readme for instructions
# + [markdown] id="Uy5_PTmOh1hj"
# <a id='1.1'></a>
# ## 2.1. Add FinRL to your path. You can of course install it as a pipy package, but this is for development purposes.
#
# + colab={"base_uri": "https://localhost:8080/"} id="mPT0ipYE28wL" outputId="802ae0b5-d88e-46ba-8082-9eb5890f9cba"
import sys
sys.path.append("..")
# -
import pandas as pd
print(pd.__version__)
# + [markdown] id="osBHhVysOEzi"
#
# <a id='1.2'></a>
# ## 2.2. Check if the additional packages needed are present, if not install them.
# * Yahoo Finance API
# * pandas
# * numpy
# * matplotlib
# * stockstats
# * OpenAI gym
# * stable-baselines
# * tensorflow
# * pyfolio
# + [markdown] id="nGv01K8Sh1hn"
# <a id='1.3'></a>
# ## 2.3. Import Packages
# + colab={"base_uri": "https://localhost:8080/"} id="lPqeTTwoh1hn" outputId="c437c266-2780-4c50-af8b-6868e7fdaa1f"
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# matplotlib.use('Agg')
import datetime
# %matplotlib inline
from finrl.config import config
from finrl.marketdata.yahoodownloader import YahooDownloader
from finrl.preprocessing.preprocessors import FeatureEngineer
from finrl.preprocessing.data import data_split
from finrl.env.env_stocktrading_v2 import StockTradingEnvV2
from finrl.model.models import DRLAgent
from finrl.trade.backtest import BackTestStats, BaselineStats, BackTestPlot
from pprint import pprint
# + [markdown] id="T2owTj985RW4"
# <a id='1.4'></a>
# ## 2.4. Create Folders
# + id="w9A8CN5R5PuZ"
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR):
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
# + [markdown] id="A289rQWMh1hq"
# <a id='2'></a>
# # Part 3. Download Data
# Yahoo Finance is a website that provides stock data, financial news, financial reports, etc. All the data provided by Yahoo Finance is free.
# * FinRL uses a class **YahooDownloader** to fetch data from Yahoo Finance API
# * Call Limit: Using the Public API (without authentication), you are limited to 2,000 requests per hour per IP (or up to a total of 48,000 requests a day).
#
# + [markdown] id="NPeQ7iS-LoMm"
#
#
# -----
# class YahooDownloader:
# Provides methods for retrieving daily stock data from
# Yahoo Finance API
#
# Attributes
# ----------
# start_date : str
# start date of the data (modified from config.py)
# end_date : str
# end date of the data (modified from config.py)
# ticker_list : list
# a list of stock tickers (modified from config.py)
#
# Methods
# -------
# fetch_data()
# Fetches data from yahoo API
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="h3XJnvrbLp-C" outputId="87dea23f-469d-4e9d-de91-0f8a74929de2"
# from config.py start_date is a string
config.START_DATE
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="FUnY8WEfLq3C" outputId="c635ae69-a13e-408f-d932-9d386d1d6dcf"
# from config.py end_date is a string
config.END_DATE
# + colab={"base_uri": "https://localhost:8080/"} id="JzqRRTOX6aFu" outputId="d3baf63f-948a-49f9-f6f2-b7241971b8ea"
print(config.DOW_30_TICKER)
# + colab={"base_uri": "https://localhost:8080/"} id="yCKm4om-s9kE" outputId="932583d8-f98b-4243-c02d-375f7272db1a"
df = YahooDownloader(start_date = '2009-01-01',
end_date = '2021-01-01',
ticker_list = config.DOW_30_TICKER).fetch_data()
# + colab={"base_uri": "https://localhost:8080/"} id="CV3HrZHLh1hy" outputId="b7b78172-8c8a-41c9-c8a6-0167edb9bd11"
df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="4hYkeaPiICHS" outputId="ce9d7463-a74c-4917-c96d-848a1e8ad493"
df.sort_values(['date','tic'],ignore_index=True).head()
# + [markdown] id="uqC6c40Zh1iH"
# # Part 4: Preprocess Data
# Data preprocessing is a crucial step for training a high quality machine learning model. We need to check for missing data and do feature engineering in order to convert the data into a model-ready state.
# * Add technical indicators. In practical trading, various information needs to be taken into account, for example the historical stock prices, current holding shares, technical indicators, etc. In this article, we demonstrate two trend-following technical indicators: MACD and RSI.
# * Add turbulence index. Risk-aversion reflects whether an investor will choose to preserve the capital. It also influences one's trading strategy when facing different market volatility level. To control the risk in a worst-case scenario, such as financial crisis of 2007–2008, FinRL employs the financial turbulence index that measures extreme asset price fluctuation.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
fe = FeatureEngineer(
use_technical_indicator=True,
tech_indicator_list = config.TECHNICAL_INDICATORS_LIST,
use_turbulence=True,
user_defined_feature = False)
processed = fe.preprocess_data(df)
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="grvhGJJII3Xn" outputId="91d09c37-b0e9-4c5c-d532-967e40d11f41"
processed.sort_values(['date','tic'],ignore_index=True).head(10)
# + [markdown] id="-QsYaY0Dh1iw"
# <a id='4'></a>
# # Part 5. Design Environment
# Considering the stochastic and interactive nature of the automated stock trading tasks, a financial task is modeled as a **Markov Decision Process (MDP)** problem. The training process involves observing stock price change, taking an action and reward's calculation to have the agent adjusting its strategy accordingly. By interacting with the environment, the trading agent will derive a trading strategy with the maximized rewards as time proceeds.
#
# Our trading environments, based on OpenAI Gym framework, simulate live stock markets with real market data according to the principle of time-driven simulation.
#
# The action space describes the allowed actions that the agent interacts with the environment. Normally, action a includes three actions: {-1, 0, 1}, where -1, 0, 1 represent selling, holding, and buying one share. Also, an action can be carried upon multiple shares. We use an action space {-k,…,-1, 0, 1, …, k}, where k denotes the number of shares to buy and -k denotes the number of shares to sell. For example, "Buy 10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or -10, respectively. The continuous action space needs to be normalized to [-1, 1], since the policy is defined on a Gaussian distribution, which needs to be normalized and symmetric.
# + [markdown] id="5TOhcryx44bb"
# ## Training data split: 2009-01-01 to 2018-12-31
# ## Trade data split: 2019-01-01 to 2020-09-30
# + colab={"base_uri": "https://localhost:8080/"} id="W0qaVGjLtgbI" outputId="c98aeb90-84e3-4b83-9671-d679f3fe148f"
train = data_split(processed, '2009-01-01','2019-01-01')
trade = data_split(processed, '2019-01-01','2021-01-01')
print(len(train))
print(len(trade))
# -
import time
milliseconds = int(round(time.time() * 1000))
print(milliseconds)
# +
import numpy as np
import pandas as pd
from gym.utils import seeding
import gym
from gym import spaces
import matplotlib
from copy import deepcopy
matplotlib.use("Agg")
import matplotlib.pyplot as plt
import pickle
from stable_baselines3.common.vec_env import DummyVecEnv
import numpy as np
import pandas as pd
from gym.utils import seeding
import gym
from gym import spaces
import matplotlib
import random
matplotlib.use("Agg")
import matplotlib.pyplot as plt
import pickle
from stable_baselines3.common.vec_env import DummyVecEnv, SubprocVecEnv
from stable_baselines3.common import logger
import time
class StockTradingEnvV2(gym.Env):
"""
A stock trading environment for OpenAI gym
Parameters:
state space: {start_cash, <owned_shares>, for s in stocks{<stock.values>}, }
df (pandas.DataFrame): Dataframe containing data
transaction_cost (float): cost for buying or selling shares
hmax (int): max number of share purchases allowed per asset
turbulence_threshold (float): Maximum turbulence allowed in market for purchases to occur. If exceeded, positions are liquidated
print_verbosity(int): When iterating (step), how often to print stats about state of env
reward_scaling (float): Scaling value to multiply reward by at each step.
initial_amount: (int, float): Amount of cash initially available
daily_information_columns (list(str)): Columns to use when building state space from the dataframe.
out_of_cash_penalty (int, float): Penalty to apply if the algorithm runs out of cash
action space: <share_dollar_purchases>
TODO:
property for date index - starting point
tests:
after reset, static strategy should result in same metrics
buy zero should result in no costs, no assets purchased
given no change in prices, no change in asset values
"""
metadata = {"render.modes": ["human"]}
def __init__(
self,
df,
transaction_cost_pct=3e-3,
date_col_name="date",
hmax=10,
turbulence_threshold=None,
print_verbosity=10,
reward_scaling=1e-4,
initial_amount=1e6,
daily_information_cols=["open", "close", "high", "low", "volume"],
out_of_cash_penalty=None,
cache_indicator_data = True,
daily_reward = None,
cash_penalty_proportion = 0.1,
random_start = True
):
self.df = df
self.stock_col = "tic"
self.assets = df[self.stock_col].unique()
self.dates = df[date_col_name].sort_values().unique()
self.random_start = random_start
self.df = self.df.set_index(date_col_name)
self.hmax = hmax
self.initial_amount = initial_amount
if out_of_cash_penalty is None:
out_of_cash_penalty=-initial_amount*0.5
self.out_of_cash_penalty = out_of_cash_penalty
self.print_verbosity = print_verbosity
self.transaction_cost_pct = transaction_cost_pct
self.reward_scaling = reward_scaling
self.daily_information_cols = daily_information_cols
self.close_index = self.daily_information_cols.index("close")
self.state_space = (
1 + len(self.assets) + len(self.assets) * len(self.daily_information_cols)
)
self.action_space = spaces.Box(low=-1, high=1, shape=(len(self.assets),))
self.observation_space = spaces.Box(
low=-np.inf, high=np.inf, shape=(self.state_space,)
)
self.episode = -1 # initialize so we can call reset
self.episode_history = []
self.printed_header = False
self.daily_reward = daily_reward
self.cache_indicator_data = cache_indicator_data
self.cached_data = None
self.cash_penalty_proportion = cash_penalty_proportion
if self.cache_indicator_data:
print("caching data")
self.cached_data = [self.get_date_vector(i) for i, _ in enumerate(self.dates)]
print("data cached!")
def seed(self, seed=None):
if seed is None:
seed = int(round(time.time() * 1000))
random.seed(seed)
@property
def current_step(self):
return self.date_index-self.starting_point
def reset(self):
self.seed()
self.sum_trades = 0
if self.random_start:
starting_point = random.choice(range(int(len(self.dates)*0.5)))
self.starting_point = starting_point
else:
self.starting_point = 0
self.date_index = self.starting_point
self.episode += 1
self.actions_memory = []
self.transaction_memory = []
self.state_memory = []
self.account_information = {
"cash": [],
"asset_value": [],
"total_assets": [],
'reward': []
}
init_state = np.array(
[self.initial_amount]
+ [0] * len(self.assets)
+ self.get_date_vector(self.date_index)
)
self.state_memory.append(init_state)
return init_state
def get_date_vector(self, date, cols=None):
if (cols is None) and (self.cached_data is not None):
return self.cached_data[date]
else:
date = self.dates[date]
if cols is None:
cols = self.daily_information_cols
trunc_df = self.df.loc[date]
v = []
for a in self.assets:
subset = trunc_df[trunc_df[self.stock_col] == a]
v += subset.loc[date, cols].tolist()
assert len(v) == len(self.assets) * len(cols)
return v
def return_terminal(self, reason='Last Date', reward=0):
state = self.state_memory[-1]
self.log_step(reason = reason, terminal_reward= reward)
reward = reward*self.reward_scaling
# Add outputs to logger interface
reward_pct = self.account_information['total_assets'][-1]/self.initial_amount
logger.record("environment/total_reward_pct", (reward_pct-1)*100)
logger.record("environment/daily_trades", self.sum_trades/(self.current_step)/len(self.assets))
logger.record("environment/completed_steps", self.current_step)
logger.record("environment/sum_rewards", np.sum(self.account_information['reward']))
logger.record("environment/cash_proportion", self.account_information['cash'][-1]/self.account_information['total_assets'][-1])
return state, reward, True, {}
def log_step(self, reason, terminal_reward=None):
if terminal_reward is None:
terminal_reward = self.account_information['reward'][-1]
cash_pct = self.account_information['cash'][-1]/self.account_information['total_assets'][-1]
rec = [self.episode, self.date_index-self.starting_point, reason, f"${int(self.account_information['total_assets'][-1])}",f"${terminal_reward:0.2f}", f"{cash_pct*100:0.2f}%"]
self.episode_history.append(rec)
print(self.template.format(*rec))
def step(self, actions):
#let's just log what we're doing in terms of max actions at each step.
self.sum_trades += np.sum(np.abs(actions))
#print header only first time
if self.printed_header is False:
self.template = "{0:4}|{1:4}|{2:15}|{3:10}|{4:10}|{5:10}" # column widths: 8, 10, 15, 7, 10
print(self.template.format("EPISODE", "STEPS", "TERMINAL_REASON", "TOT_ASSETS", "TERMINAL_REWARD_unsc", "CASH_PCT"))
self.printed_header = True
# define terminal function in scope so we can do something about the cycle being over
# print if it's time.
if (self.current_step+ 1) % self.print_verbosity == 0:
self.log_step(reason = 'update')
#if we're at the end
if self.date_index == len(self.dates) - 1:
#if we hit the end, set reward to total gains (or losses)
terminal_reward = self.account_information['total_assets'][-1]-self.initial_amount
return self.return_terminal(reward = terminal_reward)
else:
#compute value of cash + assets
begin_cash = self.state_memory[-1][0]
holdings = self.state_memory[-1][1 : len(self.assets) + 1]
assert (min(holdings)>=0)
closings = np.array(self.get_date_vector(self.date_index, cols=["close"]))
asset_value = np.dot(holdings, closings)
# reward is (cash + assets) - (cash_last_step + assets_last_step)
if self.current_step==0:
reward = 0
else:
#stepwise reward
reward = (
begin_cash + asset_value - self.account_information["total_assets"][-1]
)
#
if self.daily_reward is not None:
reward+=(self.daily_reward*self.current_step)
if self.cash_penalty_proportion is not None:
cash_penalty = min(0, begin_cash-(self.cash_penalty_proportion*self.initial_amount))
reward += cash_penalty
# log the values of cash, assets, and total assets
self.account_information["cash"].append(begin_cash)
self.account_information["asset_value"].append(asset_value)
self.account_information["total_assets"].append(begin_cash + asset_value)
self.account_information['reward'].append(reward)
# multiply action values by our scalar multiplier and save
actions = actions * self.hmax
self.actions_memory.append(actions)
#scale cash purchases to asset # changes
actions = actions/closings
self.transaction_memory.append(actions)
# clip actions so we can't sell more assets than we hold
actions = np.maximum(actions, -np.array(holdings))
# compute our proceeds from sales, and add to cash
sells = -np.clip(actions, -np.inf, 0)
proceeds = np.dot(sells, closings)
costs = proceeds * self.transaction_cost_pct
coh = begin_cash + proceeds
# compute the cost of our buys
buys = np.clip(actions, 0, np.inf)
spend = np.dot(buys, closings)
costs += spend * self.transaction_cost_pct
# if we run out of cash, end the cycle and penalize
if (spend + costs) > coh:
return self.return_terminal(reason = 'CASH SHORTAGE',
reward=self.out_of_cash_penalty,
)
# verify we didn't do anything impossible here
assert (spend + costs) <= coh
# update our holdings
coh = coh - spend - costs
holdings_updated = holdings + actions
self.date_index += 1
state = (
[coh] + list(holdings_updated) + self.get_date_vector(self.date_index)
)
self.state_memory.append(state)
reward = reward * self.reward_scaling
return state, reward, False, {}
def get_sb_env(self):
def get_self():
return deepcopy(self)
e = DummyVecEnv([get_self])
obs = e.reset()
return e, obs
def get_multiproc_env(self, n = 10):
def get_self():
return deepcopy(self)
e = SubprocVecEnv([get_self for _ in range(n)], start_method = 'fork')
obs = e.reset()
return e, obs
def save_asset_memory(self):
if self.current_step==0:
return None
else:
self.account_information["date"] = self.dates[- len(self.account_information['cash']):]
return pd.DataFrame(self.account_information)
def save_action_memory(self):
if self.current_step==0:
return None
else:
return pd.DataFrame(
{"date": self.dates[-len(self.account_information['cash']): ],
"actions": self.actions_memory,
"transactions": self.transaction_memory}
)
# -
print(StockTradingEnvV2.__doc__)
# + [markdown] colab={"base_uri": "https://localhost:8080/"} id="Q2zqII8rMIqn" outputId="8a2c943b-1be4-4b8d-b64f-666e0852b7e6"
# #### state space
# The state space of the observation is as follows
#
# `start_cash, <owned_shares_of_n_assets>, <<indicator_i_for_asset_j> for j in assets>`
#
# indicators are any daily measurement you can achieve. Common ones are 'volume', 'open' 'close' 'high', 'low'.
# However, you can add these as needed,
# The feature engineer adds indicators, and you can add your own as well.
#
# + id="AWyp84Ltto19"
information_cols = ['open', 'high', 'low', 'close', 'volume', 'day', 'macd', 'rsi_30', 'cci_30', 'dx_30', 'turbulence']
env_kwargs = {
"hmax": 5000,
"daily_reward": 5,
"out_of_cash_penalty": 0,
"cash_penalty_proportion": 0.1,
"daily_information_cols": information_cols,
"print_verbosity": 500,
}
e_train_gym = StockTradingEnvV2(df = train, cache_indicator_data=True,
**env_kwargs)
# e_train_obs = StockTradingEnvV2(df = train, cache_indicator_data=False, **env_kwargs)
# e_trade_gym = StockTradingEnvV2(df = train,**env_kwargs)
# env_trade, obs_trade = e_trade_gym.get_sb_env()
# + [markdown] id="64EoqOrQjiVf"
# ## Environment for Training
# There are two available environments. The multiprocessing and the single processing env.
# Some models won't work with multiprocessing.
#
# ```python
# # single processing
# env_train, _ = e_train_gym.get_sb_env()
#
#
# #multiprocessing
# env_train, _ = e_train_gym.get_multiproc_env(n = <n_cores>)
# ```
#
# + colab={"base_uri": "https://localhost:8080/"} id="xwSvvPjutpqS" jupyter={"outputs_hidden": true} outputId="406e5ec3-28ba-4a72-9b22-0d031f7bf9a6"
# for this example, let's do multiprocessing with n_cores-2
import multiprocessing
n_cores = multiprocessing.cpu_count() - 2
n_cores = 24
print(f"using {n_cores} cores")
#
e_train_gym.print_verbosity = 500
env_train, _ = e_train_gym.get_multiproc_env(n = n_cores)
# env_train, _ = e_train_gym.get_sb_env()
env_train_obs, _ = e_train_gym.get_sb_env()
# + [markdown] id="HMNR5nHjh1iz"
# <a id='5'></a>
# # Part 6: Implement DRL Algorithms
# * The implementation of the DRL algorithms are based on **OpenAI Baselines** and **Stable Baselines**. Stable Baselines is a fork of OpenAI Baselines, with a major structural refactoring, and code cleanups.
# * FinRL library includes fine-tuned standard DRL algorithms, such as DQN, DDPG,
# Multi-Agent DDPG, PPO, SAC, A2C and TD3. We also allow users to
# design their own DRL algorithms by adapting these DRL algorithms.
# + id="364PsqckttcQ"
agent = DRLAgent(env = env_train)
# -
print(config.PPO_PARAMS)
# + [markdown] id="YDmqOyF9h1iz"
# ### Model Training: 5 models, A2C DDPG, PPO, TD3, SAC
#
# + [markdown] id="uijiWgkuh1jB"
# ### Model 1: A2C
#
# +
from torch.nn import Softsign, ReLU
ppo_params ={'n_steps': 256,
'ent_coef': 0.01,
'learning_rate': 0.00001,
'batch_size': 256,
'gamma': 0.99}
policy_kwargs = {
# "activation_fn": ReLU,
"net_arch": [1024, 1024, 1024, 1024],
# "squash_output": True
}
model = agent.get_model("ppo", model_kwargs = ppo_params, policy_kwargs = policy_kwargs, verbose = 0)
# -
model.learn(total_timesteps = 10000000,
eval_env = env_train_obs,
eval_freq = 1000,
log_interval = 1,
tb_log_name = 'cashbuffer_1_16_longrun',
n_eval_episodes = 1,
reset_num_timesteps = True)
model.save("quicksave_ppo_dow_1_17.model")
# + id="efwBi84ch1jE"
data_turbulence = processed[(processed.date<'2019-01-01') & (processed.date>='2009-01-01')]
insample_turbulence = data_turbulence.drop_duplicates(subset=['date'])
# + colab={"base_uri": "https://localhost:8080/"} id="VHZMBpSqh1jG" outputId="f750f515-9f4f-4adb-846e-ea0bdf15ea6b"
insample_turbulence.turbulence.describe()
# + id="yuwDPkV9h1jL"
turbulence_threshold = np.quantile(insample_turbulence.turbulence.values,1)
# + colab={"base_uri": "https://localhost:8080/"} id="wwoz_7VSh1jO" outputId="37894e93-d22e-4e3f-f23a-d3ca08bf8342"
turbulence_threshold
# + [markdown] id="U5mmgQF_h1jQ"
# ### Trade
#
# DRL model needs to update periodically in order to take full advantage of the data, ideally we need to retrain our model yearly, quarterly, or monthly. We also need to tune the parameters along the way, in this notebook I only use the in-sample data from 2009-01 to 2018-12 to tune the parameters once, so there is some alpha decay here as the length of trade date extends.
#
# Numerous hyperparameters – e.g. the learning rate, the total number of samples to train on – influence the learning process and are usually determined by testing some variations.
# + id="eLOnL5eYh1jR"
trade = data_split(processed, '2019-01-01','2021-01-01')
env_kwargs = {
"hmax": 5000,
"daily_reward": 5,
"out_of_cash_penalty": 0,
"cash_penalty_proportion": 0.1,
"daily_information_cols": information_cols,
"print_verbosity": 50,
"random_start": False,
"cache_indicator_data": False
}
e_trade_gym = StockTradingEnvV2(df = trade,**env_kwargs)
env_trade, obs_trade = e_trade_gym.get_sb_env()
# -
print(len(e_trade_gym.dates))
df_account_value, df_actions = DRLAgent.DRL_prediction(model=model,
test_data = trade,
test_env = env_trade,
test_obs = obs_trade)
# + colab={"base_uri": "https://localhost:8080/"} id="ERxw3KqLkcP4" outputId="cbb465c9-38dc-4d88-e79a-6ae29025164b"
df_account_value.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="2yRkNguY5yvp" outputId="53ec139f-88e7-4291-cf11-8e6766184265"
df_account_value.head(50)
# + [markdown] id="W6vvNSC6h1jZ"
# <a id='6'></a>
# # Part 7: Backtest Our Strategy
# Backtesting plays a key role in evaluating the performance of a trading strategy. Automated backtesting tool is preferred because it reduces the human error. We usually use the Quantopian pyfolio package to backtest our trading strategies. It is easy to use and consists of various individual plots that provide a comprehensive image of the performance of a trading strategy.
# + [markdown] id="Lr2zX7ZxNyFQ"
# <a id='6.1'></a>
# ## 7.1 BackTestStats
# pass in df_account_value, this information is stored in env class
#
# + colab={"base_uri": "https://localhost:8080/"} id="Nzkr9yv-AdV_" outputId="1053083a-d74c-48b0-a623-de33282e2fff"
print("==============Get Backtest Results===========")
now = datetime.datetime.now().strftime('%Y%m%d-%Hh%M')
perf_stats_all = BackTestStats(account_value=df_account_value, value_col_name = 'total_assets')
perf_stats_all = pd.DataFrame(perf_stats_all)
perf_stats_all.to_csv("./"+config.RESULTS_DIR+"/perf_stats_all_"+now+'.csv')
# + [markdown] id="9U6Suru3h1jc"
# <a id='6.2'></a>
# ## 7.2 BackTestPlot
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="lKRGftSS7pNM" outputId="4f77cef2-3934-444a-cacc-4ed<PASSWORD>"
print("==============Compare to DJIA===========")
# %matplotlib inline
# S&P 500: ^GSPC
# Dow Jones Index: ^DJI
# NASDAQ 100: ^NDX
BackTestPlot(df_account_value,
baseline_ticker = '^DJI',
baseline_start = '2019-01-01',
baseline_end = '2021-01-01', value_col_name = 'total_assets')
# + [markdown] id="SlLT9_5WN478"
# <a id='6.3'></a>
# ## 7.3 Baseline Stats
# + colab={"base_uri": "https://localhost:8080/"} id="YktexHcqh1jc" outputId="38566531-a3a0-4705-db30-d437e8f8fc73"
print("==============Get Baseline Stats===========")
baesline_perf_stats=BaselineStats('^DJI',
baseline_start = '2019-01-01',
baseline_end = '2021-01-01')
# + id="A6W2J57ch1j9"
|
notebooks/multistock_variant_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Корректность проверена на Python 3.7:**
# + pandas 0.23.0
# + numpy 1.14.5
# + scipy 1.1.0
# + statsmodels 0.9.0
# # <NAME>
# +
import numpy as np
import pandas as pd
import scipy
from statsmodels.stats.weightstats import *
# -
import scipy
import statsmodels
print(np.__version__)
print(pd.__version__)
print(scipy.__version__)
print(statsmodels.__version__)
# %pylab inline
# ## Treatment effects of methylphenidate
# В рамках исследования эффективности препарата метилфенидат 24 пациента с синдромом дефицита внимания и гиперактивности в течение недели принимали либо метилфенидат, либо плацебо. В конце недели каждый пациент проходили тест на способность к подавлению импульсивных поведенческих реакций. На втором этапе плацебо и препарат менялись, и после недельного курса каждый испытуемые проходили второй тест.
#
# Требуется оценить эффект применения препарата.
#
# <NAME>, <NAME>., <NAME>., et al. (2004). Treatment effects of methylphenidate on cognitive functioning in children with mental retardation and ADHD. Journal of the American Academy of Child and Adolescent Psychiatry, 43(6), 677–685.
data = pd.read_csv('ADHD.txt', sep = ' ', header = 0)
data.columns = ['Placebo', 'Methylphenidate']
data.plot.scatter('Placebo', 'Methylphenidate', c = 'r', s = 30)
pylab.grid()
pylab.plot(range(100), c = 'black')
pylab.xlim((20, 80))
pylab.ylim((20, 80))
pylab.show()
data.plot.hist()
pylab.show()
# ## Одновыборочный критерий Стьюдента
# Исходя из того, что способность к подавлению испульсивных поведенческих реакций измеряется по шкале [0, 100], можно предположить, что при хорошей калибровке теста средняя способоность к подавлению реакций в популяции составляет 50. Тогда для того, чтобы проверить гипотезу о том, что пациенты в выборке действительно в среднем хуже справляются с подавлением импульсивных реакций (нуждаются в лечении), давайте проверим, что их способность к подавлению реакций отличается от средней (не равна 50).
# $H_0\colon$ среднее значение способности к подавлению испульсивных поведенческих реакций равно 50.
#
# $H_1\colon$ не равно.
stats.ttest_1samp(data.Placebo, 50.0)
print("95%% confidence interval: [%f, %f]" % zconfint(data.Placebo))
# ## Двухвыборочный критерий Стьюдента (независимые выборки)
# Для того, чтобы использовать двухвыборочный критерий Стьюдента, убедимся, что распределения в выборках существенно не отличаются от нормальных.
pylab.figure(figsize=(12,8))
pylab.subplot(2,2,1)
stats.probplot(data.Placebo, dist="norm", plot=pylab)
pylab.subplot(2,2,2)
stats.probplot(data.Methylphenidate, dist="norm", plot=pylab)
pylab.show()
# Критерий Шапиро-Уилка:
#
# $H_0\colon$ способности к подавлению импульсивных реакций распредлены нормально
#
# $H_1\colon$ не нормально.
print("Shapiro-Wilk normality test, W-statistic: %f, p-value: %f" % stats.shapiro(data.Placebo))
print("Shapiro-Wilk normality test, W-statistic: %f, p-value: %f" % stats.shapiro(data.Methylphenidate))
# С помощью критерия Стьюдента проверим гипотезу о развенстве средних двух выборок.
# Критерий Стьюдента:
#
# $H_0\colon$ средние значения способности к подавлению испульсивных поведенческих реакций одинаковы для пациентов, принимавших препарат, и для пациентов, принимавших плацебо.
#
# $H_0\colon$ не одинаковы.
scipy.stats.ttest_ind(data.Placebo, data.Methylphenidate, equal_var = False)
cm = CompareMeans(DescrStatsW(data.Methylphenidate), DescrStatsW(data.Placebo))
print("95%% confidence interval: [%f, %f]" % cm.tconfint_diff(usevar='unequal'))
# ## Двухвыборочный критерий Стьюдента (зависмые выборки)
# Для того, чтобы использовать критерй Стьюдента для связанных выборок, давайте проверим, что распределение попарных разностей существенно не отличается от нормального.
stats.probplot(data.Placebo - data.Methylphenidate, dist = "norm", plot = pylab)
pylab.show()
# <NAME>:
#
# $H_0\colon$ попарные разности распределены нормально.
#
# $H_1\colon$ не нормально.
print("Shapiro-Wilk normality test, W-statistic: %f, p-value: %f" % stats.shapiro(data.Methylphenidate - data.Placebo))
# Критерий Стьюдента:
#
# $H_0\colon$ средние значения способности к подавлению испульсивных поведенческих реакций одинаковы для пациентов, принимавших препарат, и для пациентов, принимавших плацебо.
#
# $H_1\colon$ не одинаковы.
stats.ttest_rel(data.Methylphenidate, data.Placebo)
print("95%% confidence interval: [%f, %f]" % DescrStatsW(data.Methylphenidate - data.Placebo).tconfint_mean())
|
Yandex data science/4/Week 2/.ipynb_checkpoints/stat.student_tests-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 3
import numpy as np
from matplotlib import pyplot as plt
path="https://raw.githubusercontent.com/Anasuya-Sahoo/DMDW-Lab/main/Toyota.csv"
import pandas as pd
data=pd.read_csv(path)
type(data)
data.shape
data.info()
data.index
data.columns
data.head()
data.tail()
data.head(5)
data[['Price',"Age"]].head(10)
data.isnull().sum()
data.dropna(inplace=True)
data.isnull().sum()
data.shape
data.head(10)
data['MetColor'].mean()
data['MetColor'].head()
data.head(10)
data['CC'].mean()
data['CC'].head()
data[['Age',"KM"]].head(20)
|
18cse002-Assignment 3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 練習時間
# 資料的操作有很多,接下來的馬拉松中我們會介紹常被使用到的操作,參加者不妨先自行想像一下,第一次看到資料,我們一般會想知道什麼訊息?
#
# #### Ex: 如何知道資料的 row 數以及 column 數、有什麼欄位、多少欄位、如何截取部分的資料等等
#
# 有了對資料的好奇之後,我們又怎麼通過程式碼來達成我們的目的呢?
#
# #### 可參考該[基礎教材](https://bookdata.readthedocs.io/en/latest/base/01_pandas.html#DataFrame-%E5%85%A5%E9%97%A8)或自行 google
# # [作業目標]
# - 熟悉更多的 Python 資料操作
# # [作業重點]
# - 列出資料的大小 (In[4], Hint : shape)
# - 列出所有欄位 (In[5], 有多種寫法)
# - 擷取部分資料 (In[6], Hint : loc 或 iloc)
import os
import numpy as np
import pandas as pd
# 設定 data_path
dir_data = 'D:\\GitHub\\3rd-ML100Days\\HomeWork\\Data'
crime_csv = os.path.join(dir_data, 'crime.csv')
print('Path of read in data: %s' % (crime_csv))
crime_csv_train = pd.read_csv(crime_csv)
# ### 如果沒有想法,可以先嘗試找出剛剛例子中提到的問題的答案
# #### 資料的 row 數以及 column 數
print ('row : ', crime_csv_train.shape[0] ,', column : ', crime_csv_train.shape[1])
# #### 列出所有欄位
#crime_csv_train_DF = pd.DataFrame(crime_csv_train)
crime_csv_train_DF = pd.DataFrame(crime_csv_train)
print ('Each columns : ', list(crime_csv_train_DF.columns.values))
# #### 截取部分資料
crime_csv_train_DF.tail(1)
# #### 還有各種數之不盡的資料操作,重點還是取決於實務中遇到的狀況和你想問的問題,在馬拉松中我們也會陸續提到更多例子
search = crime_csv_train_DF["IS_CRIME"] >= 1
print(crime_csv_train_DF[search])
|
HomeWork/Day_004_HW.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="JgBwwu4ZAHKK"
# # Interactive sensitivity analysis with GP regression model
#
# In this notebook we create an interactive version of the sensitivity analysis we developed the previous step of the tutorial.
# You can use this to explore the predictions and sensitivities when varying the inputs to the model.
# In particular, you can see how the behaviour of certain important inputs is affected when changing the values of other important inputs.
#
# This notebook contains a lot of code, however, most of it is for creating the interactive plots. The code used for the sensitivity analysis is the same as we have seen earlier in the tutorial.
# + [markdown] id="s01R_4uPAP9k"
# ## Dependencies
#
# As in the previous notebooks, we start by importing all dependencies.
#
# If you are in Colab, you need to install the [pyro](https://pyro.ai/) package by uncommenting and running the line `!pip3 install pyro-ppl` below before proceeding.
# + executionInfo={"elapsed": 315, "status": "ok", "timestamp": 1642407257748, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="_Q3hnowbBhdQ"
# install dependencies
# # !pip3 install pyro-ppl
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1104, "status": "ok", "timestamp": 1642407259070, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="qQqUp2oV_7S5" outputId="abd34b68-c633-4002-a0df-2509214804ec"
# imports
from collections import defaultdict
from pathlib import Path
import matplotlib.pyplot as plt
from matplotlib.ticker import FormatStrFormatter
import numpy as np
import pandas as pd
import torch
import pyro
import pyro.contrib.gp as gp
import ipywidgets as widgets
from matplotlib.ticker import FormatStrFormatter
pyro.set_rng_seed(0)
print(f"torch version: {torch.__version__}")
print(f"pyro version: {pyro.__version__}")
# + [markdown] id="eDiPq5VOArDQ"
# ## Load the dataset and model parameters
#
# We can load the dataset directly from the GitHub URL.
# Alternatively, the dataset can be loaded from a local file.
# + executionInfo={"elapsed": 419, "status": "ok", "timestamp": 1642407259486, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="9K_6MiokAuVc"
# load dataset
dataset_path = "https://raw.githubusercontent.com/BIG-MAP/sensitivity_analysis_tutorial/main/data/p2d_sei_10k.csv"
# dataset_path = "data/p2d_sei_10k.csv" # local
df = pd.read_csv(dataset_path, index_col=0)
# store the names of the features and the name of the target variable
features = df.columns[:15].tolist() # use input parameters as features
target = "SEI_thickness(m)" # primary target
# target = "Capacity loss (%)" # secondary target
# + [markdown] id="GNGMKLjfAt0x"
# We also need to load the trained model parameters that we saved previously.
#
# If you are running this notebook in Colab, you need to make the parameter file available in the working directory by uploading it to the Files section to the left.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 17, "status": "ok", "timestamp": 1642407259487, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="1-iz-4W8A1oP" outputId="8e97e40a-ad33-4ff0-8054-3fc3d280f267"
pyro.clear_param_store()
if target == "SEI_thickness(m)":
pyro.get_param_store().load("sgpr_params_sei.p")
if target == "Capacity loss (%)":
pyro.get_param_store().load("sgpr_params_cap.p")
params = pyro.get_param_store()
params.keys()
# + [markdown] id="0soo-23HA7KJ"
# ## Setup model
#
# Setup the model with the trained parameters.
# + executionInfo={"elapsed": 12, "status": "ok", "timestamp": 1642407259487, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="JlLeifboA_NF"
kernel = gp.kernels.RBF(input_dim=params["data.x_train"].shape[1], variance=params["kernel.variance"], lengthscale=params["kernel.lengthscale"])
model = gp.models.SparseGPRegression(params["data.x_train"], params["data.y_train"], kernel, Xu=params["Xu"], noise=params["noise"])
# + [markdown] id="X2BhUVtgBJ14"
# ## Interactive sensitivity analysis
#
#
# + executionInfo={"elapsed": 11, "status": "ok", "timestamp": 1642407259487, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="ttPKxfUuBMlp"
def sa_autograd(model, X, reduce=None):
"""Sensitivity analysis of GP regression model with automatic differentiation.
Args:
model: Gaussian process regression model
X (tensor): Input data (design matrix)
reduce (string): method used to reduce the sensitivity result: sum, mean, none.
"""
X.requires_grad = True
# compute gradient of the mean prediction
model.zero_grad()
mean, _ = model(X, full_cov=False, noiseless=False)
gmean = torch.autograd.grad(mean.sum(), X)[0]
# compute gradient of the variance prediction
model.zero_grad()
_, var = model(X, full_cov=False, noiseless=False)
gvar = torch.autograd.grad(var.sum(), X)[0]
X.requires_grad = False
if reduce == "sum":
return mean, var, torch.sqrt(torch.sum(gmean**2, dim=0)), torch.sqrt(torch.sum(gvar**2, dim=0))
elif reduce == "mean":
return mean, var, torch.sqrt(torch.mean(gmean**2, dim=0)), torch.sqrt(torch.mean(gvar**2, dim=0))
else:
return mean, var, torch.sqrt(gmean**2), torch.sqrt(gvar**2)
# + executionInfo={"elapsed": 12, "status": "ok", "timestamp": 1642407259488, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="mzUX9Z8bBUL-"
def predict_sa(x, reduce=None):
return sa_autograd(model, x, reduce=reduce)
# + [markdown] id="ficqxHv5Cgm6"
# ### Interactive sensitivity analysis: global
#
# Here you can experiment with how the average sensitivities are affected by range we consider for each input.
# + colab={"base_uri": "https://localhost:8080/", "height": 497, "referenced_widgets": ["bddade887dbe4e7b920aae3267df35b5", "9760b524ee8c4cf78be2d3a21654ff6d", "13f3be5ea2a04d2f80566d8d33a18ddf", "<KEY>", "<KEY>", "<KEY>", "c92c519a2d8542a4ae8c587818ceed65", "c5ed03be7f08411893e080395a2480fa", "daa8d48eba374c2e85bb08f092062db8", "dad5f206e25d492a81e30d0d42f05373", "07ede0d40fdc42c3ac1fd8b6864c53c3", "<KEY>", "<KEY>", "5aa4975fd3c74c7aa5c88eb4dfbe96ca", "8c26af306c5844128e6ac386c87bac77", "641e26ed2e594dd3aafb86877c61a3d9", "c7df870edabc42e3905f7c4db06bdf0f", "b57302cb8a3441a28e3e0f85c79c239e", "<KEY>", "fb3525aca9c74523848ceb9128e62aab", "<KEY>", "8cc1ce6ac67a45029f7df7e5e35d4d95", "f6264eada7bd42fcb62f912492fe6c82", "ce744a59c50048ca85cdce1c2ac7a3aa", "<KEY>", "<KEY>", "8e9ed535f7b645bda09c5ac30238fb50", "d91a56c269974e7997e46a81364b93a5", "<KEY>", "838e2fb5efa14705987429a1f9af45f6", "1e44ed3891614587819b489a9fee57b6", "<KEY>", "72f655d5ab2f44fba0e1aff3f0d7defb", "6e71344920694d4ca3bdce8564a1d9cb", "2138d79545084d3f8e7cb650eb20a695", "<KEY>", "<KEY>", "d0d348644b21453c863ecd1f7262a51d", "<KEY>", "2ca827738d4840058ca37f7d61b88783", "<KEY>", "<KEY>", "<KEY>", "16c26dbed83d4695b1b46a8f77fb19c1", "<KEY>", "ecec07a3c89c4b5ea8bac85214c9c34d", "be6885744edc4b168e3794096e56edeb", "<KEY>", "e9f66cc818114f4ebb139305e532712a", "75e357ab40ac48189176a134d5c65fa9", "cd1f01ff7ac04eafbd9220d99319c6c0"]} executionInfo={"elapsed": 942, "status": "ok", "timestamp": 1642407260421, "user": {"displayName": "jonas busk", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="efxzCcKfBr3Y" outputId="fdb85fdb-6fef-4e32-823f-15232ce9d61b"
def create_predict_and_plot_global(predict_sa, features, n_sample=1000, normalise=False, figsize=(12,6)):
def predict_and_plot(**x_dict):
x_min = torch.tensor([x_dict[f][0] for f in features])
x_max = torch.tensor([x_dict[f][1] for f in features])
# create inputs
X = torch.distributions.Uniform(x_min, x_max).sample((n_sample,))
# predict
mean, var, s_mean, s_var = predict_sa(X, reduce="mean")
mean, var, s_mean, s_var = mean.detach(), var.detach(), s_mean.detach(), s_var.detach()
std = var.sqrt().detach()
# normalise
if normalise:
s_mean = s_mean / s_mean.sum()
s_var = s_var / s_var.sum()
# plot
plt.figure(figsize=figsize)
# plot sensitivity of mean prediction
plt.subplot(121)
plt.bar(range(len(features)), s_mean, label="s_mean")
#plt.bar(range(len(features)), s_var, alpha=0.75, label="s_var")
if normalise:
plt.ylim((0,.8))
plt.xticks(range(len(features)), [f"x{i}: {f}" for i,f in enumerate(features)], rotation=90)
plt.xlabel("Input feature"); plt.ylabel("Sensitivity")
plt.legend()
plt.grid(axis='y')
# plot sensitivity of var prediction
plt.subplot(122)
#plt.bar(range(len(features)), s_mean, label="s_mean")
plt.bar(range(len(features)), s_var, color="C1", label="s_var")
if normalise:
plt.ylim((0,.8))
plt.xticks(range(len(features)), [f"x{i}: {f}" for i,f in enumerate(features)], rotation=90)
plt.xlabel("Input feature"); plt.ylabel("Sensitivity")
plt.legend()
plt.grid(axis='y')
plt.tight_layout()
plt.show()
return predict_and_plot
def interactive_global(on_change, features, x_min, x_max, n_steps=20):
sliders = {}
#sliders["d"] = widgets.IntSlider(value=0, min=0, max=len(features)-1, description="dim")
for i, f in enumerate(features):
sliders[f] = widgets.FloatRangeSlider(
value=[x_min[i], x_max[i]],
min=x_min[i],
max=x_max[i],
step=(x_max[i] - x_min[i]) / n_steps,
description=f"x{i}: {f}",
readout_format=".1f",
)
# setup ui
out = widgets.interactive_output(on_change, sliders)
controls = widgets.VBox(list(sliders.values()))
ui = widgets.HBox([controls, out])
# display ui
display(ui)
# on change function
on_change_global = create_predict_and_plot_global(
predict_sa,
features,
n_sample=5000,
normalise=True,
)
# setup ui
interactive_global(
on_change_global,
features,
x_min=params["data.x_min"].detach().numpy(),
x_max=params["data.x_max"].detach().numpy(),
)
# + [markdown] id="M_0jwmPhCsnn"
# ### Interactive sensitivity analysis: 1D
#
# Here you can see how the behaviour of certain important inputs along their range of variation is affected when changing the values of other important inputs.
# + colab={"base_uri": "https://localhost:8080/", "height": 529, "referenced_widgets": ["ada9d2f832714cddbe29e171f10cf9c2", "63d70e40c3a745a2bd02944a747732b6", "74b1947906504e9b94ecc2f79bb7ea7f", "418ac64bd49248af975cab76a2fc9e52", "df68e9150716411caaa7f83723a04133", "<KEY>", "<KEY>", "1791cca6824f408d958c209854be5012", "<KEY>", "<KEY>", "<KEY>", "288d6506a7084906b345a00455f31620", "<KEY>", "5b030b62296d434aa3aa0b5c66d301df", "5b25afc1702245ca98a888a08ebeee6a", "603ff4e41fee4a91889948ad50718334", "5153fb93a7a94532879ee3e81dd32edd", "368fe401fc8941b9a592f7e0faff10e3", "66f60aa5b9564d14b468ec8491a83149", "ffbe629cece7427a84d78201fff1fc52", "5fc89c44ade0445593a9534b6e61601a", "60de0696c0964b5ab62f8e97d0967ca8", "<KEY>", "<KEY>", "11ee799f5ac24327a87015e945bde102", "b29c06a34cf44ac781a4725c2806c89a", "bde2c7666c71459a8eed8345fa951c46", "9501e0df5ad54ee2b08a1e33cf91737c", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "1c198650cafd4d18a26033de2340841d", "<KEY>", "e9f4377761e840ecb9669b5b682271be", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8283be0f4e2f439ba9f3c7ced93f4d72", "<KEY>", "<KEY>", "0d2e6a2df58e4d3ca3746f7360cb5979", "2f94cad5c81344b986ef1b5c6ef52a25", "07480d171e4446c98721f5683925e880", "381f11e8fff24484839afe63ca954aac", "5af80034ada149b7a63dd1461de40379", "<KEY>", "<KEY>", "ef817541b52944c992913ef3de3306c5", "<KEY>", "<KEY>", "dd3b65fe4c58487e91d83f2d8b48f29f"]} executionInfo={"elapsed": 1506, "status": "ok", "timestamp": 1642407261923, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="GoU4oIdkB_82" outputId="33cdf956-383a-489e-ab2f-f3d9b6d133f6"
def create_predict_and_plot_1d(predict_sa, features, target, x_min, x_max, y_lim=None, n_points=100, figsize=(12,7)):
def predict_and_plot(d, **x_dict):
x_list = [x_dict[f] for f in features]
# create inputs
x = torch.tensor(x_list)
X = x.repeat(n_points, 1)
xd = torch.linspace(x_min[d], x_max[d], n_points)
X[:,d] = xd
# predict point
mean0, var0, s_mean0, s_var0 = predict_sa(x.unsqueeze(0))
mean0, var0, s_mean0, s_var0 = mean0.detach(), var0.detach(), s_mean0.detach(), s_var0.detach()
std0 = var0.sqrt()
# predict grid
mean, var, s_mean, s_var = predict_sa(X)
mean, var, s_mean, s_var = mean.detach(), var.detach(), s_mean.detach(), s_var.detach()
std = var.sqrt().detach()
plt.figure(figsize=figsize)
# plot mean prediction with uncertainty
plt.subplot(221)
plt.title("mean prediction with uncertainty (2*std)")
plt.plot(xd.numpy(), mean.numpy())
plt.fill_between(xd.numpy(), (mean.numpy() - 2.0 * std.numpy()), (mean.numpy() + 2.0 * std.numpy()), color='C0', alpha=0.3)
plt.axvline(x[d].numpy(), color="k", linewidth=1, label=f"{mean0.item():.4f} ({std0.item():.4f})")
plt.xlim((x_min[d], x_max[d]))
if y_lim is not None:
plt.ylim(y_lim)
plt.xlabel(f"x{d}: {features[d]}")
plt.ylabel(f"log y: {target}")
plt.grid()
plt.legend(loc=4)
# plot uncertainty
plt.subplot(222)
plt.title("uncertainty prediction (2*std)")
plt.plot(xd.numpy(), 2*std.numpy())
plt.axvline(x[d].numpy(), color="k", linewidth=1, label=f"{2 * std0.item():.4f}")
plt.xlim((x_min[d], x_max[d]))
plt.ylim((0,1))
plt.xlabel(f"x{d}: {features[d]}")
plt.ylabel("uncertainty")
plt.grid()
plt.legend(loc=4)
# plot sensitivity of mean
plt.subplot(223)
plt.title("sensitivity of mean prediction")
plt.plot(xd.numpy(), s_mean[:, d].numpy())
plt.axvline(x[d].numpy(), color="k", linewidth=1, label=f"{s_mean0[:,d].item():.4f}")
plt.xlim((x_min[d], x_max[d]))
plt.ylim((0,5))
plt.xlabel(f"x{d}: {features[d]}")
plt.ylabel("sensitivity")
plt.grid()
plt.legend(loc=4)
# plot sensitivity of var
plt.subplot(224)
plt.title("sensitivity of uncertainty prediction")
plt.plot(xd.numpy(), s_var[:, d].numpy())
plt.axvline(x[d].numpy(), color="k", linewidth=1, label=f"{s_var0[:,d].item():.4f}")
plt.xlim((x_min[d], x_max[d]))
plt.ylim((0,0.3))
plt.xlabel(f"x{d}: {features[d]}")
plt.ylabel("sensitivity")
plt.grid()
plt.legend(loc=4)
plt.tight_layout()
plt.show()
return predict_and_plot
def interactive_1d(on_change, features, x_min, x_max, x_init, n_steps=20):
sliders = {}
sliders["d"] = widgets.IntSlider(value=0, min=0, max=len(features)-1, description="dim")
for i, f in enumerate(features):
sliders[f] = widgets.FloatSlider(
value=x_init[i],
min=x_min[i],
max=x_max[i],
step=(x_max[i] - x_min[i]) / n_steps,
description=f"x{i}: {f}",
readout_format=".1f",
)
# setup ui
out = widgets.interactive_output(on_change, sliders)
controls = widgets.VBox(list(sliders.values()))
ui = widgets.HBox([controls, out])
# display ui
display(ui)
# on change function
on_change_1d = create_predict_and_plot_1d(
predict_sa,
features,
target,
x_min=params["data.x_min"].detach().numpy(),
x_max=params["data.x_max"].detach().numpy(),
y_lim=(params["data.y_min"].item(), params["data.y_max"].item()),
)
# setup ui
interactive_1d(
on_change_1d,
features,
x_min=params["data.x_min"].detach().numpy(),
x_max=params["data.x_max"].detach().numpy(),
x_init=params["data.x_train"][0],
)
# + [markdown] id="yIXu_UauC7f_"
# ### Interactive sensitivity analysis: 2D
#
# This lets you plot two inputs at a time to see how their combined behaviour is affected when changing the values of other important inputs.
# + colab={"base_uri": "https://localhost:8080/", "height": 764, "referenced_widgets": ["da32ff4d3f4b4c809e3cfa0920cb5f9b", "760f800cca694275b690384fc4647a82", "37a0493a72c840f7929b11fb24f1f34a", "30b40c02f0f645a6918505dc447483a0", "3c6146903c68431788b167cf071e3896", "757baf992d0f43f098e89185e49423f5", "a8a5141f73c5499883f54d002864d643", "f59ac41310604efca4f657695d5bb2e5", "a4399524d4c645ec854725025a43c74f", "fb99ea5333ce4c82a5f4edd440212fe3", "144eed98cf3b4e5297c886d4a1f95509", "68492df211d84b30877fe36d89b0ceb3", "9fe6680c56a4422482238ba608fc6b2b", "<KEY>", "34b1b214d7514bf3a88be9de4e8c6903", "<KEY>", "4c077e3b0c1a4b25863e100d8e2c06aa", "23adc59a3a764547bc1050ef4ce30128", "fbba1d1411e244a3a79658d3db5dccba", "<KEY>", "3d60d74d9e4542de837e0594011d8f3d", "511199631f974fa3b70bd411f536648f", "80b707edd98745b0991e87a61b0f7aae", "<KEY>", "86233be6f94542ad8018a2e3175e4c64", "<KEY>", "5591bec3966e40e1bfe1a421961eb3db", "<KEY>", "cbc8b5291ed547079fe2ade0864360c0", "<KEY>", "<KEY>", "2562b3ede2044b07891ffc337aa98f3e", "63ed26e3759a4997b1c807fea252a03b", "7eb448e8de4b42cd86dbe6cb5195725e", "3e3b2e66228d4c43a4c7274e51a55588", "<KEY>", "<KEY>", "957efdf74e054483ab982e2418ae9227", "5448248eb1654270a6203e7eedbab191", "<KEY>", "d8c37c99b87e458e83ab42213733978b", "<KEY>", "76d4fbdacff6428fb2cee8261145137d", "<KEY>", "2d5643f147dc498a955b972a34a84321", "189c9db8c3d3463c8ccedb308160e891", "<KEY>", "11272c0e272f450e91f9a56d03f66d67", "c96fc4ca39ab44dd980eb98ce99c3e26", "29f949f4925348f49e6cfd44e6130f08", "4c3ee3e0ec254a7e9c45d35859cd40f9", "69bf43856f464f8caeeb7bc5ec27ef56", "<KEY>", "e9797f6768c442748258efcef7bb686c", "27ed7fdd50d6435ba3f31d2a38373606", "f8a6733dbe8d4f5184305dd0fd779f8c", "f4a502411dda465b8199549a827b008a"]} executionInfo={"elapsed": 2083, "status": "ok", "timestamp": 1642407263999, "user": {"displayName": "jonas busk", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj7-1gAF7PppPBq1jWtOrRLj_kiVnCZpQDWsCTO4g=s64", "userId": "13756499934799797810"}, "user_tz": -60} id="sptR-_4DC8x1" outputId="5ae9428b-8170-4da3-ec27-01e493b78cc1"
# setup on_change function
def create_predict_and_plot_2d(predict_sa, features, target, x_min, x_max, y_lim=None, n_points=100, n_levels=21, figsize=(12,10)):
def predict_and_plot(d0, d1, **x_dict):
x_list = [x_dict[f] for f in features]
# create inputs
x = torch.tensor(x_list)
X = x.repeat(n_points**2, 1)
# setup grid
xd0 = torch.linspace(x_min[d0], x_max[d0], n_points)
xd1 = torch.linspace(x_min[d1], x_max[d1], n_points)
grid_xd0, grid_xd1 = torch.meshgrid(xd0, xd1)
X[:,d0] = grid_xd0.reshape(len(X))
X[:,d1] = grid_xd1.reshape(len(X))
# predict point
mean0, var0, s_mean0, s_var0 = predict_sa(x.unsqueeze(0))
mean0, var0, s_mean0, s_var0 = mean0.detach(), var0.detach(), s_mean0.detach(), s_var0.detach()
std0 = var0.sqrt()
# predict grid
mean, var, s_mean, s_var = predict_sa(X)
mean, var, s_mean, s_var = mean.detach(), var.detach(), s_mean.detach(), s_var.detach()
std = var.sqrt()
s_mean0_d = (s_mean0[:, d0] + s_mean0[:, d1]).item()
s_var0_d = (s_var0[:, d0] + s_var0[:, d1]).item()
s_mean_d = (s_mean[:, d0] + s_mean[:, d1]).reshape(n_points, n_points)
s_var_d = (s_var[:, d0] + s_var[:, d1]).reshape(n_points, n_points)
# plot
plt.figure(figsize=figsize)
# plot mean prediction
ax = plt.subplot(221)
plt.title("mean prediction of log y")
if y_lim is None:
levels = torch.linspace(mean.min().item(), mean.max().item(), n_levels).numpy()
else:
levels = torch.linspace(y_lim[0], y_lim[1], n_levels).numpy()
plt.contourf(grid_xd0.numpy(), grid_xd1.numpy(), mean.reshape(n_points, n_points).numpy(), levels=levels, cmap="plasma")
plt.axvline(x[d0].numpy(), color="k", linewidth=1, label=f"{mean0.item():.4f} ({std0.item():.4f})")
plt.axhline(x[d1].numpy(), color="k", linewidth=1)
plt.xlabel(f"x{d0}: {features[d0]}"); plt.ylabel(f"x{d1}: {features[d1]}")
plt.colorbar(shrink=0.9)
ax.yaxis.set_major_formatter(FormatStrFormatter('%6.2f'))
plt.legend(loc=4)
# plot uncertainty
ax = plt.subplot(222)
plt.title("uncertainty (2*std)")
levels = torch.linspace(0, 1.0, 21).numpy()
plt.contourf(grid_xd0.numpy(), grid_xd1.numpy(), 2*std.reshape(n_points, n_points).numpy(), levels=levels, cmap="plasma")
plt.axvline(x[d0].numpy(), color="k", linewidth=1, label=f"{std0.item()*2:.4f}")
plt.axhline(x[d1].numpy(), color="k", linewidth=1)
plt.xlabel(f"x{d0}: {features[d0]}"); plt.ylabel(f"x{d1}: {features[d1]}")
plt.colorbar(shrink=0.9)
ax.yaxis.set_major_formatter(FormatStrFormatter('%6.2f'))
plt.legend(loc=4)
# plot sensitivity of mean prediction
ax = plt.subplot(223)
plt.title("sensitivity of mean prediction")
levels = torch.linspace(0, 5.0, 21).numpy()
plt.contourf(grid_xd0.numpy(), grid_xd1.numpy(), s_mean_d.numpy(), levels=levels, cmap="plasma")
plt.axvline(x[d0].numpy(), color="k", linewidth=1, label=f"{s_mean0_d:.4f}")
plt.axhline(x[d1].numpy(), color="k", linewidth=1)
plt.xlabel(f"x{d0}: {features[d0]}"); plt.ylabel(f"x{d1}: {features[d1]}")
plt.colorbar(shrink=0.9)
ax.yaxis.set_major_formatter(FormatStrFormatter('%6.2f'))
plt.legend(loc=4)
# plot sensitivity of uncertainty prediction
ax = plt.subplot(224)
plt.title("sensitivity of uncertainty prediction")
levels = torch.linspace(0, 0.25, 21).numpy()
plt.contourf(grid_xd0.numpy(), grid_xd1.numpy(), s_var_d.numpy(), levels=levels, cmap="plasma")
plt.axvline(x[d0].numpy(), color="k", linewidth=1, label=f"{s_var0_d:.4f}")
plt.axhline(x[d1].numpy(), color="k", linewidth=1)
plt.xlabel(f"x{d0}: {features[d0]}"); plt.ylabel(f"x{d1}: {features[d1]}")
plt.colorbar(shrink=0.9)
ax.yaxis.set_major_formatter(FormatStrFormatter('%6.2f'))
plt.legend(loc=4)
plt.tight_layout()
plt.show()
return predict_and_plot
def interactive_2d(on_change, features, x_min, x_max, x_init, n_steps=20):
sliders = {}
sliders["d0"] = widgets.IntSlider(value=0, min=0, max=len(features)-1, description="dim 0")
sliders["d1"] = widgets.IntSlider(value=1, min=0, max=len(features)-1, description="dim 1")
for i, f in enumerate(features):
sliders[f] = widgets.FloatSlider(
value=x_init[i],
min=x_min[i],
max=x_max[i],
step=(x_max[i] - x_min[i]) / n_steps,
description=f"x{i}: {features[i]}",
readout_format=".1f",
)
# setup ui
out = widgets.interactive_output(on_change, sliders)
controls = widgets.VBox(list(sliders.values()))
ui = widgets.HBox([controls, out])
# display ui
display(ui)
# on change function
on_change_2d = create_predict_and_plot_2d(
predict_sa,
features,
target,
x_min=params["data.x_min"].detach().numpy(),
x_max=params["data.x_max"].detach().numpy(),
y_lim=(params["data.y_min"].item(), params["data.y_max"].item()),
)
# setup ui
interactive_2d(
on_change_2d,
features,
x_min=params["data.x_min"].detach().numpy(),
x_max=params["data.x_max"].detach().numpy(),
x_init=params["data.x_train"][0],
)
|
notebooks/7_interactive_sensitivity_analysis_of_gpr.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Active Learning on Logistic Regression
# ## Configuration
# +
# Config
"""
Set to a number to generate same split
Number of Iteration: number of iteraton of active learning
Number of Query: number of queries for to extract samples near to decision boundary
"""
random_state=0
number_of_iteration=7
number_of_query = 20
N_THRESHOLD = 1
# -
# ## Active Learning Helper
# ### least_confident
#
# 1. Get probability of test data by calling `predict_proba`, return 2d vectors [0_probability, 1_probability]
# 2. For each vector, get max value of 2 probability values and negate the number to generate score. Higher score means higher uncertainty
# 3. Get top N data.
#
# +
# Active Learning Helper
# From https://github.com/davefernig/alp/
from __future__ import unicode_literals, division
from scipy.sparse import csc_matrix, vstack
from scipy.stats import entropy
from collections import Counter
import numpy as np
class ActiveLearner(object):
_uncertainty_sampling_frameworks = [
'entropy',
'max_margin',
'least_confident',
]
_query_by_committee_frameworks = [
'vote_entropy',
'average_kl_divergence',
]
def __init__(self, strategy='least_confident'):
self.strategy = strategy
def rank(self, clf, X_unlabeled, num_queries=None):
if num_queries == None:
num_queries = X_unlabeled.shape[0]
elif type(num_queries) == float:
num_queries = int(num_queries * X_unlabeled.shape[0])
if self.strategy in self._uncertainty_sampling_frameworks:
scores = self.__uncertainty_sampling(clf, X_unlabeled)
elif self.strategy in self._query_by_committee_frameworks:
scores = self.__query_by_committee(clf, X_unlabeled)
else:
raise ValueError(
"No input strategy"
)
rankings = np.argsort(-scores)[:num_queries]
return rankings
def __uncertainty_sampling(self, clf, X_unlabeled):
probs = clf.predict_proba(X_unlabeled)
if self.strategy == 'least_confident':
return 1 - np.amax(probs, axis=1)
elif self.strategy == 'max_margin':
margin = np.partition(-probs, 1, axis=1)
return -np.abs(margin[:,0] - margin[:, 1])
elif self.strategy == 'entropy':
return np.apply_along_axis(entropy, 1, probs)
def __query_by_committee(self, clf, X_unlabeled):
num_classes = len(clf[0].classes_)
C = len(clf)
preds = []
if self.strategy == 'vote_entropy':
for model in clf:
y_out = map(int, model.predict(X_unlabeled))
preds.append(np.eye(num_classes)[y_out])
votes = np.apply_along_axis(np.sum, 0, np.stack(preds)) / C
return np.apply_along_axis(entropy, 1, votes)
elif self.strategy == 'average_kl_divergence':
for model in clf:
preds.append(model.predict_proba(X_unlabeled))
consensus = np.mean(np.stack(preds), axis=0)
divergence = []
for y_out in preds:
divergence.append(entropy(consensus.T, y_out.T))
return np.apply_along_axis(np.mean, 0, np.stack(divergence))
class RandomLearner(object):
"""
Random return idx
for control group
"""
def __init__(self, strategy=''):
self.strategy = strategy
def rank(self, clf, X_unlabeled, num_queries=None):
if num_queries == None:
num_queries = X_unlabeled.shape[0]
elif type(num_queries) == float:
num_queries = int(num_queries * X_unlabeled.shape[0])
idx = np.arange(len(X_unlabeled))
np.random.shuffle(idx)
return idx[:num_queries]
# -
def process(X, y, learner, model, config):
# Initialization
precisions = []
recalls = []
predict_proba = []
parameter = [
"test_size", # test/(train + test)
"random_state",
"init_size", #
"query_each_iteration",
]
default_config = {
"test_size": 0.4,
"random_state": 0,
"init_size": 0.5,
"query_each_iteration": 5,
}
cfg = {}
# load config
for k in parameter:
cfg[k] = config[k] if config.get(k) else default_config[k]
# Split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=cfg["test_size"], random_state=cfg["random_state"])
init_size = int(np.ceil(X_train.shape[0] * cfg["init_size"]))
pool_size = X_train.shape[0] - init_size
X_init = X_train[:init_size]
y_init = y_train[:init_size]
X_pool = X_train[init_size:]
y_pool = y_train[init_size:]
X_train = X_init
y_train = y_init
number_of_iteration = int(np.ceil(pool_size/cfg["query_each_iteration"]))
query_each_iteration = cfg["query_each_iteration"]
print("Initial training set size:", init_size, "(", cfg["init_size"], ")")
print("Query each iteration:", cfg["query_each_iteration"])
# init state
model.fit(X_train, y_train)
y_predict = model.predict(X_test)
precision, recall, thresholds = precision_recall_curve(y_test, y_predict)
precisions.append(precision[N_THRESHOLD])
recalls.append(recall[N_THRESHOLD])
print("init")
print("@P:", precision)
print("@R:", recall)
print(thresholds)
print()
# Active Learning
for i in range(number_of_iteration):
idx = learner.rank(model, X_pool, query_each_iteration)
t = model.predict_proba(X_pool[idx[0]:])
min_certain = np.amax(model.predict_proba(X_pool[idx[0:1]]))
max_certain = np.amax(model.predict_proba(X_pool[idx[-1:]]))
predict_proba.append([min_certain, max_certain])
X_train = np.concatenate([X_train, X_pool[idx, :]])
y_train = np.concatenate([y_train, y_pool[idx]])
X_pool = np.delete(X_pool, idx, axis=0)
y_pool = np.delete(y_pool, idx, axis=0)
model.fit(X_train, y_train)
# metrics
y_predict = model.predict(X_test)
precision, recall, thresholds = precision_recall_curve(y_test, y_predict)
precisions.append(precision[N_THRESHOLD])
recalls.append(recall[N_THRESHOLD])
print("Round: ", i+1)
print("Proba:", min_certain, max_certain)
print("Train: ", len(X_train))
print("Test: ", len(X_test))
print("@P:", precision)
print("@R:", recall)
print(thresholds)
debug = {
"X_train": X_train
}
info = {
"predict_proba": [predict_proba[0]] + predict_proba
}
return precisions, recalls, debug, info
import numpy as np
import sklearn
from sklearn import datasets
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
# metrics
from sklearn.metrics import precision_recall_curve
# plot
import matplotlib.pyplot as plt
# %matplotlib inline
config = {
"test_size":0.4,
"random_state": random_state,
"init_size": 0.3,
"query_each_iteration": 5
}
# +
# Active Learning
#Load Data
path = "data/"
y = np.array([])
T1 = np.loadtxt(path+"hurricane.vec")
T = T1
y = np.append(y, np.ones(len(T)))
F1 = np.loadtxt(path+"lasvegas.vec")
F2 = np.loadtxt(path+"false.vec")
F = np.append(F1, F2, axis=0)
y = np.append(y, np.zeros(len(F)))
embedding = np.append(T,F,axis=0)
X = sklearn.preprocessing.normalize(embedding)
model = sklearn.linear_model.LogisticRegressionCV()
active_learner = ActiveLearner("least_confident")
active_learning_precision, active_learning_recall, debug1, info1 = process(X, y, active_learner, model, config)
# +
# Control Group
#Load Data
path = "data/"
y = np.array([])
T1 = np.loadtxt(path+"hurricane.vec")
T = T1
y = np.append(y, np.ones(len(T)))
F1 = np.loadtxt(path+"lasvegas.vec")
F2 = np.loadtxt(path+"false.vec")
F = np.append(F1, F2, axis=0)
y = np.append(y, np.zeros(len(F)))
embedding = np.append(T,F,axis=0)
X = sklearn.preprocessing.normalize(embedding)
model = sklearn.linear_model.LogisticRegressionCV()
random_learner = RandomLearner()
control_group_precision, control_group_recall, debug2, info2 = process(X, y, random_learner, model, config)
# -
x = range(len(active_learning_precision))
plt.figure(figsize=(18, 18))
plt.xlabel("number of iteration")
proba, = plt.plot(x, [n[0] for n in info1["predict_proba"]],"g--" , label="proba")
p1, = plt.plot(x, active_learning_precision, "r-", label="AL @P")
r1, = plt.plot(x, active_learning_recall, "r--", label="AL @R")
p2, = plt.plot(x, control_group_precision, "b-", label="Control @P")
r2, = plt.plot(x, control_group_recall, "b--", label="Control @R")
plt.legend(handles=[p1, r1, p2, r2, proba])
# Debug Utils
def compare_X_train():
xt1 = debug1["X_train"]
xt2 = debug2["X_train"]
a = set([tuple(l) for l in xt1])
b = set([tuple(l) for l in xt2])
if a - b:
print("invalid X train: should be same X train")
compare_X_train()
|
Active Learning Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/timeseriesAI/tsai/blob/master/tutorial_nbs/00c_Time_Series_data_preparation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# created by <NAME> - email: <EMAIL>
# ## Import libraries 📚
# Since some of you have been asking questions as to how to prepare your data to be able to use timeserisAI, I've prepared a short tutorial to address this.
#
# There are endless options in terms of how your source data may be stored, so I'll cover a few of the most frequent ones I've seen. I may be expanding this in the future if needed.
# +
# # **************** UNCOMMENT AND RUN THIS CELL IF YOU NEED TO INSTALL/ UPGRADE TSAI ****************
# stable = True # Set to True for latest pip version or False for main branch in GitHub
# # !pip install {"tsai -U" if stable else "git+https://github.com/timeseriesAI/tsai.git"} >> /dev/null
# -
from tsai.all import *
computer_setup()
# ## Required input shape 🔶
# To be able to use timeseriesAI your data needs to have 3 dimensions:
#
# * **number of samples**
# * **number of features** (aka variables, dimensions, channels)
# * **number of steps** (or length, time steps, sequence steps)
#
# There are a few convenience functions that you may want to use to prepare your data.
#
# We are going to see how you could prepare your data in a few scenarios.
# **Note: I've recently modified timeseriesAI so that you can also use 2d input data in the case of univariate time series (they'll be converted to 3d internally), although you can still pass univariate time series as 3d or pass them if you prefer. You'll get the same result.**
# ## UCR time series data ⏳
# The easiest case if if you want to use some of the data already preprocessed in timeseriesAI (all UCR datasets have been included). In this case, the only thing you need to do is:
#
# * select a univariate or multivariate dataset from the list
# * use the get_UCR_data function
print('univariate datasets: ', get_UCR_univariate_list())
print('multivariate datasets: ', get_UCR_multivariate_list())
ds_name = 'NATOPS'
X, y, splits = get_UCR_data(ds_name, return_split=False)
X.shape, y.shape, splits
tfms = [None, [Categorize()]]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits, inplace=True)
dsets
# As you can see, X has 3 dimensions:
#
# * 360 samples
# * 24 features
# * 51 time steps
#
# Let's visualize of the samples:
plt.plot(X[0].T);
# ## 2d or 3d np.ndarray/ torch.Tensor ⌗
# Another option is that you have your data as an array or a tensor.
# In this case, the only thing you'll need to do is to transform your data to 3d (if not already done), and generate your splits.
# We are going to simulate this scenario generating 2d data for a univariate dataset:
ds_name = 'OliveOil'
X, y, _ = get_UCR_data(ds_name, return_split=False)
X_2d = X[:, 0]
X_2d.shape, y.shape
# To make data 3d you use `to3d`:
X_3d = to3d(X_2d)
X_3d.shape
# To generate your splits, you would use `get_splits`. Here you need to indicate:
# * valid_size=0.2
# * test_size (optional)
# * stratify=True if you want stratified splits
# * random_state=seed or None (random)
splits = get_splits(y, valid_size=.2, stratify=True, random_state=23, shuffle=True)
splits
X_3d.shape, y.shape, splits
tfms = [None, [Categorize()]]
dsets = TSDatasets(X_3d, y, tfms=tfms, splits=splits, inplace=True)
dsets
# In fastai I've modified TS datasets so that you can pass univariate time series as a 2d or 3d arrays.
tfms = [None, [Categorize()]]
dsets = TSDatasets(X_2d, y, tfms=tfms, splits=splits, inplace=True)
dsets
# ### Pre-split 2d or 3d np.ndarray/ torch.Tensor
# If your data is already split into Train and Valid/ Test, you may the use `get_predefined_split` to generate the splits:
ds_name = 'OliveOil'
X_train, y_train, X_valid, y_valid = get_UCR_data(ds_name, return_split=True)
X, y, splits = combine_split_data([X_train, X_valid], [y_train, y_valid])
tfms = [None, [Categorize()]]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits, inplace=True)
dsets
# ## Pandas dataframe with samples as rows 🐼
# ### Univariate
ds_name = 'OliveOil'
X, y, _ = get_UCR_data(ds_name, return_split=False)
X = X[:, 0]
y = y.reshape(-1, 1)
data = np.concatenate((X, y), axis=-1)
df = pd.DataFrame(data)
df = df.rename(columns={570: 'target'})
df.head()
X, y = df2xy(df, target_col='target')
test_eq(X.shape, (60, 1, 570))
test_eq(y.shape, (60, ))
splits = get_splits(y, valid_size=.2, stratify=True, random_state=23, shuffle=True)
splits
tfms = [None, [Categorize()]]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits, inplace=True)
dsets
# ### Multivariate
ds_name = 'OliveOil'
X, y, _ = get_UCR_data(ds_name, return_split=False)
X = X[:, 0]
y = y.reshape(-1, 1)
data = np.concatenate((X, y), axis=-1)
df = pd.DataFrame(data).astype(float)
df = df.rename(columns={570: 'target'})
df1 = pd.concat([df, df + 10, df + 100], axis=0).reset_index(drop=False)
df2 = pd.DataFrame(np.array([1] * 60 + [2] * 60 + [3] * 60), columns=['feature'])
df = pd.merge(df2, df1, left_index=True, right_index=True)
df
X, y = df2xy(df, sample_col='index', feat_col='feature', target_col='target', data_cols=None)
test_eq(X.shape, (60, 3, 570))
test_eq(y.shape, (60, 3))
splits = get_splits(y, valid_size=.2, stratify=True, random_state=23, shuffle=True)
splits
tfms = [None, TSRegression()]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits, inplace=True)
dsets
# ## Single, long time series 🤥
# Sometimes, instead of having the data already split into samples, you only have a single (univariate or multivariate) time series that you need to split.
# The recommended way to do this is to use a sliding window. In `timeseriesAI`there is a function called `SlidingWindow`that performs this task in a flexible way.
#
# This function applies a sliding window to a 1d or 2d input (np.ndarray, torch.Tensor or pd.DataFrame).
#
# * Args:
# * window_length = length of lookback window
# * stride = n datapoints the window is moved ahead along the sequence. Default: 1. If None, stride=window_length (no overlap)
# * horizon = number of future datapoints to predict. 0 for last step in the selected window. > 0 for future steps. List for several steps.
# * get_x = indices of columns that contain the independent variable (xs). If get_x=None, all data will be used as x
# * get_y = indices of columns that contain the target (ys). If y_idx is None, no y will be returned
# * seq_first = True if input shape (seq_len, n_vars), False if input shape (n_vars, seq_len)
# * random_start = determines the step where the first window is applied: 0 (default), a given step (int), or random within the 1st stride (None).
#
# * Input:
# * shape: (seq_len, ) or (seq_len, n_vars) if seq_first=True else (n_vars, seq_len)
# ### Univariate
# You may use it just without a target
window_length = 5
t = np.arange(100)
print('input shape:', t.shape)
X, y = SlidingWindow(window_length)(t)
test_eq(X.shape, ((95, 1, 5)))
# If the target is the next step in the univariate time series set `horizon=1`:
# +
window_length = 5
horizon = 1
t = np.arange(100)
print('input shape:', t.shape)
X, y = SlidingWindow(window_length, horizon=horizon)(t)
test_eq(X.shape, ((95, 1, 5)))
test_eq(y.shape, ((95,)))
# -
# Horizon may be > 1 to select multiple steps in the future:
# +
window_length = 5
horizon = 2
t = np.arange(100)
print('input shape:', t.shape)
X, y = SlidingWindow(window_length, horizon=horizon)(t)
test_eq(X.shape, ((94, 1, 5)))
test_eq(y.shape, ((94, 2)))
# -
# To have non-overlapping samples, we need to set `stride=None`:
window_length = 5
stride = None
horizon = 1
t = np.arange(100)
print('input shape:', t.shape)
X, y = SlidingWindow(window_length, stride=stride, horizon=horizon)(t)
test_eq(X.shape, ((19, 1, 5)))
test_eq(y.shape, ((19, )))
window_length = 5
stride = 3
horizon = 1
t = np.arange(100)
print('input shape:', t.shape)
X, y = SlidingWindow(window_length, stride=stride, horizon=horizon)(t)
test_eq(X.shape, ((32, 1, 5)))
test_eq(y.shape, ((32, )))
# We can also decide where to start the sliding window using `start`:
window_length = 5
stride = None
horizon = 1
t = np.arange(100)
print('input shape:', t.shape)
X, y = SlidingWindow(window_length, stride=stride, start=20, horizon=horizon)(t)
test_eq(X.shape, ((15, 1, 5)))
test_eq(y.shape, ((15, )))
# If the time series is of shape (1, seq_len) we need to set `seq_first=False`
window_length = 5
stride = 3
horizon = 1
t = np.arange(100).reshape(1, -1)
print('input shape:', t.shape)
X, y = SlidingWindow(window_length, stride=stride, horizon=horizon, seq_first=False)(t)
test_eq(X.shape, ((32, 1, 5)))
test_eq(y.shape, ((32, )))
# Your univariate time series may be in a pandas DataFrame:
# +
window_length = 5
stride = None
horizon=1
t = np.arange(20)
df = pd.DataFrame(t, columns=['var'])
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length, stride=stride, horizon=horizon)(df)
test_eq(X.shape, ((3, 1, 5)))
test_eq(y.shape, ((3, )))
# +
window_length = 5
stride = None
horizon=1
t = np.arange(20)
df = pd.DataFrame(t, columns=['var']).T
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length, stride=stride, horizon=horizon, seq_first=False)(df)
test_eq(X.shape, ((3, 1, 5)))
test_eq(y.shape, ((3, )))
# -
# ### Multivariate
# When using multivariate data, all parameters shown before work in the same way, but you always need to indicate how to get the X data and the y data (as there are multiple features). To do that, we'll use get_x and get_y.
#
# By default get_x is set to None, which means that all features will be used.
# By default get_y is set to None, which means that all features will be used as long as horizon > 0 (to avoid leakage).
#
# If you get the time series in a np.ndarray or a torch.Tensor, you should use integers, a list or slice as get_x/ get_y.
# +
window_length = 5
stride = None
n_vars = 3
t = (np.random.rand(1000, n_vars) - .5).cumsum(0)
print(t.shape)
plt.plot(t)
plt.show()
X, y = SlidingWindow(window_length, stride=stride, get_x=[0,1], get_y=2)(t)
test_eq(X.shape, ((199, 2, 5)))
test_eq(y.shape, ((199, )))
# +
window_length = 5
n_vars = 3
t = (torch.stack(n_vars * [torch.arange(10)]).T * tensor([1, 10, 100]))
df = pd.DataFrame(t, columns=[f'var_{i}' for i in range(n_vars)])
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length)(df)
test_eq(X.shape, ((5, 3, 5)))
test_eq(y.shape, ((5, 3)))
# +
window_length = 5
n_vars = 3
horizon = 1
t = (torch.stack(n_vars * [torch.arange(10)]).T * tensor([1, 10, 100]))
df = pd.DataFrame(t, columns=[f'var_{i}' for i in range(n_vars)])
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length, horizon=horizon)(df)
test_eq(X.shape, ((5, 3, 5)))
test_eq(y.shape, ((5, 3)))
# -
# You may also get the target from a different column:
# +
window_length = 5
n_vars = 3
t = (torch.stack(n_vars * [torch.arange(10)]).T * tensor([1, 10, 100]))
columns=[f'var_{i}' for i in range(n_vars-1)]+['target']
df = pd.DataFrame(t, columns=columns)
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length, get_x=columns[:-1], get_y='target')(df)
test_eq(X.shape, ((5, 2, 5)))
test_eq(y.shape, ((5, )))
# +
window_length = 5
n_vars = 5
horizon = 1
t = (torch.stack(n_vars * [torch.arange(10)]).T * tensor([10**i for i in range(n_vars)]))
columns=[f'var_{i}' for i in range(n_vars-1)]+['target']
df = pd.DataFrame(t, columns=columns)
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length, horizon=horizon, get_x=columns[:-1], get_y='target')(df)
test_eq(X.shape, ((5, 4, 5)))
test_eq(y.shape, ((5, )))
# +
window_length = 4
n_vars = 5
seq_len = 100
horizon = 1
t1 = (np.random.rand(seq_len, n_vars-1) - .5).cumsum(0)
t2 = np.random.randint(0, 10, (seq_len,1))
t = np.concatenate((t1, t2), axis=-1)
columns=[f'var_{i}' for i in range(n_vars-1)]+['target']
df = pd.DataFrame(t, columns=columns)
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length, horizon=horizon, get_x=columns[:-1], get_y='target')(df)
splits = get_splits(y, valid_size=.2, stratify=True, random_state=23, shuffle=False)
tfms = [None, [Categorize()]]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits, inplace=True)
dsets
# -
dsets[0][0].data, dsets[0][1].data
# +
window_length = 4
start = 3
n_vars = 5
seq_len = 100
horizon = 0
t1 = (np.random.rand(seq_len, n_vars-1) - .5).cumsum(0)
t2 = np.random.randint(0, 10, (seq_len,1))
t = np.concatenate((t1, t2), axis=-1)
columns=[f'var_{i}' for i in range(n_vars-1)]+['target']
df = pd.DataFrame(t, columns=columns).T
print('input shape:', df.shape)
display(df)
X, y = SlidingWindow(window_length, start=start, horizon=horizon, get_x=columns[:-1], get_y='target', seq_first=False)(df)
splits = get_splits(y, valid_size=.2, stratify=True, random_state=23, shuffle=False)
tfms = [None, [Categorize()]]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits, inplace=True)
dsets
# -
dsets[0][0].data, dsets[0][1].data
# ## End-to-end example 🎬
# ### Data split by sample
# This is a example using real data where the dataframe already the data split by sample. Let's first simulate how you could get the pandas df.
#
# In this case, you only need to convert the df format to X and y using `df2xy`as we have seen before.
ds_name = 'NATOPS'
X, y, splits = get_UCR_data(ds_name, return_split=False)
data = np.concatenate((np.arange(len(X)).repeat(X.shape[1]).reshape(-1,1), np.tile(np.arange(X.shape[1]), len(X)).reshape(-1,1)), axis=1)
df1 = pd.DataFrame(data, columns=['sample', 'feature'])
df2 = pd.DataFrame(X.reshape(-1, 51))
df3 = pd.DataFrame(np.repeat(y, X.shape[1]), columns=['target'])
df = df1.merge(df2, left_index=True, right_index=True)
df = df.merge(df3, left_index=True, right_index=True)
df
# In this case, we can shuffle the data as the individual time series are independent from the rest.
def y_func(o): return o[0]
X, y = df2xy(df, sample_col='sample', feat_col='feature', target_col='target', data_cols=df.columns[2:-1], y_func=y_func)
splits = get_splits(y, valid_size=.2, stratify=True, random_state=23, shuffle=True)
tfms = [None, TSClassification()]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits)
dsets
# ### Single multivariate time series
# In this second scenario, we have a single time series, and we'll need to decide how to create the individual samples using the `SlidingWindow`function.
#
# This is how you could get the dataframe, with many columns for each feature, and a target.
ds_name = 'NATOPS'
X, y, splits = get_UCR_data(ds_name, return_split=False)
data = X.transpose(1,0,2).reshape(X.shape[1], -1)
print(X.shape, data.shape)
df = pd.DataFrame(data).T
df2 = pd.DataFrame(np.repeat(y, X.shape[2]), columns=['target'])
df = df.merge(df2, left_index=True, right_index=True)
df
# In this case, you'll need to set the following parameters:
#
# * window_length
# * stride
# * start
# * horizon
# * get_x
# * get_y
# * seq_first
#
# You also need to bear in mind that you sould set shuffle=False when using splits since the individual time series are correlated with rest.
# +
window_length = X.shape[-1] # window_length is usually selected based on prior domain knowledge or by trial and error
stride = None # None for non-overlapping (stride = window_length) (default = 1). This depends on how often you want to predict once the model is trained
start = 0 # use all data since the first time stamp (default = 0)
get_x = df.columns[:-1] # Indicates which are the columns that contain the x data.
get_y = 'target' # In multivariate time series, you must indicate which is/are the y columns
horizon = 0 # 0 means y is taken from the last time stamp of the time sequence (default = 0)
seq_first = True
X, y = SlidingWindow(window_length, stride=stride, start=start, get_x=get_x, get_y=get_y, horizon=horizon, seq_first=seq_first)(df)
splits = get_splits(y, valid_size=.2, stratify=True, random_state=23, shuffle=False)
tfms = [None, [Categorize()]]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits)
dsets
# -
beep()
|
tutorial_nbs/00c_Time_Series_data_preparation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="woaQzCU4smj8"
# # Image Classification
# In this project, you'll classify images from the [CIFAR-10 dataset](https://www.cs.toronto.edu/~kriz/cifar.html). The dataset consists of airplanes, dogs, cats, and other objects. You'll preprocess the images, then train a convolutional neural network on all the samples. The images need to be normalized and the labels need to be one-hot encoded. You'll get to apply what you learned and build a convolutional, max pooling, dropout, and fully connected layers. At the end, you'll get to see your neural network's predictions on the sample images.
# ## Get the Data
# Run the following cell to download the [CIFAR-10 dataset for python](https://www.cs.toronto.edu/~kriz/cifar-10-python.tar.gz).
# + id="iythrj3CtJ-5"
import os
import numpy as np
import tensorflow as tf
import random
from unittest.mock import MagicMock
def _print_success_message():
print('Tests Passed')
def test_folder_path(cifar10_dataset_folder_path):
assert cifar10_dataset_folder_path is not None,\
'Cifar-10 data folder not set.'
assert cifar10_dataset_folder_path[-1] != '/',\
'The "/" shouldn\'t be added to the end of the path.'
assert os.path.exists(cifar10_dataset_folder_path),\
'Path not found.'
assert os.path.isdir(cifar10_dataset_folder_path),\
'{} is not a folder.'.format(os.path.basename(cifar10_dataset_folder_path))
train_files = [cifar10_dataset_folder_path + '/data_batch_' + str(batch_id) for batch_id in range(1, 6)]
other_files = [cifar10_dataset_folder_path + '/batches.meta', cifar10_dataset_folder_path + '/test_batch']
missing_files = [path for path in train_files + other_files if not os.path.exists(path)]
assert not missing_files,\
'Missing files in directory: {}'.format(missing_files)
print('All files found!')
def test_normalize(normalize):
test_shape = (np.random.choice(range(1000)), 32, 32, 3)
test_numbers = np.random.choice(range(256), test_shape)
normalize_out = normalize(test_numbers)
assert type(normalize_out).__module__ == np.__name__,\
'Not Numpy Object'
assert normalize_out.shape == test_shape,\
'Incorrect Shape. {} shape found'.format(normalize_out.shape)
assert normalize_out.max() <= 1 and normalize_out.min() >= 0,\
'Incorect Range. {} to {} found'.format(normalize_out.min(), normalize_out.max())
_print_success_message()
def test_one_hot_encode(one_hot_encode):
test_shape = np.random.choice(range(1000))
test_numbers = np.random.choice(range(10), test_shape)
one_hot_out = one_hot_encode(test_numbers)
assert type(one_hot_out).__module__ == np.__name__,\
'Not Numpy Object'
assert one_hot_out.shape == (test_shape, 10),\
'Incorrect Shape. {} shape found'.format(one_hot_out.shape)
n_encode_tests = 5
test_pairs = list(zip(test_numbers, one_hot_out))
test_indices = np.random.choice(len(test_numbers), n_encode_tests)
labels = [test_pairs[test_i][0] for test_i in test_indices]
enc_labels = np.array([test_pairs[test_i][1] for test_i in test_indices])
new_enc_labels = one_hot_encode(labels)
assert np.array_equal(enc_labels, new_enc_labels),\
'Encodings returned different results for the same numbers.\n' \
'For the first call it returned:\n' \
'{}\n' \
'For the second call it returned\n' \
'{}\n' \
'Make sure you save the map of labels to encodings outside of the function.'.format(enc_labels, new_enc_labels)
for one_hot in new_enc_labels:
assert (one_hot==1).sum() == 1,\
'Each one-hot-encoded value should include the number 1 exactly once.\n' \
'Found {}\n'.format(one_hot)
assert (one_hot==0).sum() == len(one_hot)-1,\
'Each one-hot-encoded value should include zeros in all but one position.\n' \
'Found {}\n'.format(one_hot)
_print_success_message()
def test_nn_image_inputs(neural_net_image_input):
image_shape = (32, 32, 3)
nn_inputs_out_x = neural_net_image_input(image_shape)
assert nn_inputs_out_x.get_shape().as_list() == [None, image_shape[0], image_shape[1], image_shape[2]],\
'Incorrect Image Shape. Found {} shape'.format(nn_inputs_out_x.get_shape().as_list())
assert nn_inputs_out_x.op.type == 'Placeholder',\
'Incorrect Image Type. Found {} type'.format(nn_inputs_out_x.op.type)
assert nn_inputs_out_x.name == 'x:0', \
'Incorrect Name. Found {}'.format(nn_inputs_out_x.name)
print('Image Input Tests Passed.')
def test_nn_label_inputs(neural_net_label_input):
n_classes = 10
nn_inputs_out_y = neural_net_label_input(n_classes)
assert nn_inputs_out_y.get_shape().as_list() == [None, n_classes],\
'Incorrect Label Shape. Found {} shape'.format(nn_inputs_out_y.get_shape().as_list())
assert nn_inputs_out_y.op.type == 'Placeholder',\
'Incorrect Label Type. Found {} type'.format(nn_inputs_out_y.op.type)
assert nn_inputs_out_y.name == 'y:0', \
'Incorrect Name. Found {}'.format(nn_inputs_out_y.name)
print('Label Input Tests Passed.')
def test_nn_keep_prob_inputs(neural_net_keep_prob_input):
nn_inputs_out_k = neural_net_keep_prob_input()
assert nn_inputs_out_k.get_shape().ndims is None,\
'Too many dimensions found for keep prob. Found {} dimensions. It should be a scalar (0-Dimension Tensor).'.format(nn_inputs_out_k.get_shape().ndims)
assert nn_inputs_out_k.op.type == 'Placeholder',\
'Incorrect keep prob Type. Found {} type'.format(nn_inputs_out_k.op.type)
assert nn_inputs_out_k.name == 'keep_prob:0', \
'Incorrect Name. Found {}'.format(nn_inputs_out_k.name)
print('Keep Prob Tests Passed.')
def test_con_pool(conv2d_maxpool):
test_x = tf.placeholder(tf.float32, [None, 32, 32, 5])
test_num_outputs = 10
test_con_k = (2, 2)
test_con_s = (4, 4)
test_pool_k = (2, 2)
test_pool_s = (2, 2)
conv2d_maxpool_out = conv2d_maxpool(test_x, test_num_outputs, test_con_k, test_con_s, test_pool_k, test_pool_s)
assert conv2d_maxpool_out.get_shape().as_list() == [None, 4, 4, 10],\
'Incorrect Shape. Found {} shape'.format(conv2d_maxpool_out.get_shape().as_list())
_print_success_message()
def test_flatten(flatten):
test_x = tf.placeholder(tf.float32, [None, 10, 30, 6])
flat_out = flatten(test_x)
assert flat_out.get_shape().as_list() == [None, 10*30*6],\
'Incorrect Shape. Found {} shape'.format(flat_out.get_shape().as_list())
_print_success_message()
def test_fully_conn(fully_conn):
test_x = tf.placeholder(tf.float32, [None, 128])
test_num_outputs = 40
fc_out = fully_conn(test_x, test_num_outputs)
assert fc_out.get_shape().as_list() == [None, 40],\
'Incorrect Shape. Found {} shape'.format(fc_out.get_shape().as_list())
_print_success_message()
def test_output(output):
test_x = tf.placeholder(tf.float32, [None, 128])
test_num_outputs = 40
output_out = output(test_x, test_num_outputs)
assert output_out.get_shape().as_list() == [None, 40],\
'Incorrect Shape. Found {} shape'.format(output_out.get_shape().as_list())
_print_success_message()
def test_conv_net(conv_net):
test_x = tf.placeholder(tf.float32, [None, 32, 32, 3])
test_k = tf.placeholder(tf.float32)
logits_out = conv_net(test_x, test_k)
assert logits_out.get_shape().as_list() == [None, 10],\
'Incorrect Model Output. Found {}'.format(logits_out.get_shape().as_list())
print('Neural Network Built!')
def test_train_nn(train_neural_network):
mock_session = tf.Session()
test_x = np.random.rand(128, 32, 32, 3).astype(np.float32)
test_y = np.random.rand(128, 10)
test_k = np.random.rand(1)
test_optimizer = tf.train.AdamOptimizer()
mock_session.run = MagicMock()
train_neural_network(mock_session, test_optimizer, test_k, test_x, test_y)
assert mock_session.run.called, 'Session not used'
_print_success_message()
# + id="U67oPzAAsmkk" colab={"base_uri": "https://localhost:8080/"} outputId="609502e3-b410-4e45-d06b-e3bf74b21445"
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
from urllib.request import urlretrieve
from os.path import isfile, isdir
from tqdm import tqdm
#import problem_unittests as tests
import tarfile
cifar10_dataset_folder_path = 'cifar-10-batches-py'
# Use Floyd's cifar-10 dataset if present
floyd_cifar10_location = '/cifar/cifar-10-python.tar.gz'
if isfile(floyd_cifar10_location):
tar_gz_path = floyd_cifar10_location
else:
tar_gz_path = 'cifar-10-python.tar.gz'
class DLProgress(tqdm):
last_block = 0
def hook(self, block_num=1, block_size=1, total_size=None):
self.total = total_size
self.update((block_num - self.last_block) * block_size)
self.last_block = block_num
if not isfile(tar_gz_path):
with DLProgress(unit='B', unit_scale=True, miniters=1, desc='CIFAR-10 Dataset') as pbar:
urlretrieve(
'https://www.cs.toronto.edu/~kriz/cifar-10-python.tar.gz',
tar_gz_path,
pbar.hook)
if not isdir(cifar10_dataset_folder_path):
with tarfile.open(tar_gz_path) as tar:
tar.extractall()
tar.close()
test_folder_path(cifar10_dataset_folder_path)
# + [markdown] id="KYD7cudjsmkq"
# ## Explore the Data
# The dataset is broken into batches to prevent your machine from running out of memory. The CIFAR-10 dataset consists of 5 batches, named `data_batch_1`, `data_batch_2`, etc.. Each batch contains the labels and images that are one of the following:
# * airplane
# * automobile
# * bird
# * cat
# * deer
# * dog
# * frog
# * horse
# * ship
# * truck
#
# Understanding a dataset is part of making predictions on the data. Play around with the code cell below by changing the `batch_id` and `sample_id`. The `batch_id` is the id for a batch (1-5). The `sample_id` is the id for a image and label pair in the batch.
#
# Ask yourself "What are all possible labels?", "What is the range of values for the image data?", "Are the labels in order or random?". Answers to questions like these will help you preprocess the data and end up with better predictions.
# + id="FvCU-93Zu-lw"
import pickle
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelBinarizer
def _load_label_names():
"""
Load the label names from file
"""
return ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
def load_cfar10_batch(cifar10_dataset_folder_path, batch_id):
"""
Load a batch of the dataset
"""
with open(cifar10_dataset_folder_path + '/data_batch_' + str(batch_id), mode='rb') as file:
batch = pickle.load(file, encoding='latin1')
display(batch['data'].shape)
features = batch['data'].reshape((len(batch['data']), 3, 32, 32)).transpose(0, 2, 3, 1)
labels = batch['labels']
return features, labels
def display_stats(cifar10_dataset_folder_path, batch_id, sample_id):
"""
Display Stats of the the dataset
"""
batch_ids = list(range(1, 6))
if batch_id not in batch_ids:
print('Batch Id out of Range. Possible Batch Ids: {}'.format(batch_ids))
return None
features, labels = load_cfar10_batch(cifar10_dataset_folder_path, batch_id)
if not (0 <= sample_id < len(features)):
print('{} samples in batch {}. {} is out of range.'.format(len(features), batch_id, sample_id))
return None
print('\nStats of batch {}:'.format(batch_id))
print('Samples: {}'.format(len(features)))
print('Label Counts: {}'.format(dict(zip(*np.unique(labels, return_counts=True)))))
print('First 20 Labels: {}'.format(labels[:20]))
sample_image = features[sample_id]
sample_label = labels[sample_id]
label_names = _load_label_names()
print('\nExample of Image {}:'.format(sample_id))
print('Image - Min Value: {} Max Value: {}'.format(sample_image.min(), sample_image.max()))
print('Image - Shape: {}'.format(sample_image.shape))
print('Label - Label Id: {} Name: {}'.format(sample_label, label_names[sample_label]))
plt.axis('off')
plt.imshow(sample_image)
def _preprocess_and_save(normalize, one_hot_encode, features, labels, filename):
"""
Preprocess data and save it to file
"""
features = normalize(features)
labels = one_hot_encode(labels)
pickle.dump((features, labels), open(filename, 'wb'))
def preprocess_and_save_data(cifar10_dataset_folder_path, normalize, one_hot_encode):
"""
Preprocess Training and Validation Data
"""
n_batches = 5
valid_features = []
valid_labels = []
for batch_i in range(1, n_batches + 1):
features, labels = load_cfar10_batch(cifar10_dataset_folder_path, batch_i)
validation_count = int(len(features) * 0.1)
# Prprocess and save a batch of training data
_preprocess_and_save(
normalize,
one_hot_encode,
features[:-validation_count],
labels[:-validation_count],
'preprocess_batch_' + str(batch_i) + '.p')
# Use a portion of training batch for validation
valid_features.extend(features[-validation_count:])
valid_labels.extend(labels[-validation_count:])
# Preprocess and Save all validation data
_preprocess_and_save(
normalize,
one_hot_encode,
np.array(valid_features),
np.array(valid_labels),
'preprocess_validation.p')
with open(cifar10_dataset_folder_path + '/test_batch', mode='rb') as file:
batch = pickle.load(file, encoding='latin1')
# load the test data
test_features = batch['data'].reshape((len(batch['data']), 3, 32, 32)).transpose(0, 2, 3, 1)
test_labels = batch['labels']
# Preprocess and Save all test data
_preprocess_and_save(
normalize,
one_hot_encode,
np.array(test_features),
np.array(test_labels),
'preprocess_test.p')
def batch_features_labels(features, labels, batch_size):
"""
Split features and labels into batches
"""
for start in range(0, len(features), batch_size):
end = min(start + batch_size, len(features))
yield features[start:end], labels[start:end]
def load_preprocess_training_batch(batch_id, batch_size):
"""
Load the Preprocessed Training data and return them in batches of <batch_size> or less
"""
filename = 'preprocess_batch_' + str(batch_id) + '.p'
features, labels = pickle.load(open(filename, mode='rb'))
# Return the training data in batches of size <batch_size> or less
return batch_features_labels(features, labels, batch_size)
def display_image_predictions(features, labels, predictions):
n_classes = 10
label_names = _load_label_names()
label_binarizer = LabelBinarizer()
label_binarizer.fit(range(n_classes))
label_ids = label_binarizer.inverse_transform(np.array(labels))
fig, axies = plt.subplots(nrows=4, ncols=2)
fig.tight_layout()
fig.suptitle('Softmax Predictions', fontsize=20, y=1.1)
n_predictions = 3
margin = 0.05
ind = np.arange(n_predictions)
width = (1. - 2. * margin) / n_predictions
for image_i, (feature, label_id, pred_indicies, pred_values) in enumerate(zip(features, label_ids, predictions.indices, predictions.values)):
pred_names = [label_names[pred_i] for pred_i in pred_indicies]
correct_name = label_names[label_id]
axies[image_i][0].imshow(feature)
axies[image_i][0].set_title(correct_name)
axies[image_i][0].set_axis_off()
axies[image_i][1].barh(ind + margin, pred_values[::-1], width)
axies[image_i][1].set_yticks(ind + margin)
axies[image_i][1].set_yticklabels(pred_names[::-1])
axies[image_i][1].set_xticks([0, 0.5, 1.0])
# + id="2MpvMJ4Osmkt" colab={"base_uri": "https://localhost:8080/", "height": 435} outputId="89797ea4-6866-450d-817b-8fffd5625655"
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
#import helper
import numpy as np
# Explore the dataset
batch_id = 1
sample_id = 5
display_stats(cifar10_dataset_folder_path, batch_id, sample_id)
# + [markdown] id="otWjejd5smkw"
# ## Implement Preprocess Functions
# ### Normalize
# In the cell below, implement the `normalize` function to take in image data, `x`, and return it as a normalized Numpy array. The values should be in the range of 0 to 1, inclusive. The return object should be the same shape as `x`.
# + id="zkGPqn1Csmky" colab={"base_uri": "https://localhost:8080/"} outputId="f4d36a9d-d3f0-401d-81f0-f366314b33a0"
def normalize(x):
"""
Normalize a list of sample image data in the range of 0 to 1
: x: List of image data. The image shape is (32, 32, 3)
: return: Numpy array of normalize data
"""
# TODO: Implement Function
x_min = x.min(axis=(0, 1), keepdims=True)
x_max = x.max(axis=(0, 1), keepdims=True)
norm_x = (x-x_min)/(x_max-x_min)
return norm_x
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
test_normalize(normalize)
# + [markdown] id="0o61J7cPsmk1"
# ### One-hot encode
# Just like the previous code cell, you'll be implementing a function for preprocessing. This time, you'll implement the `one_hot_encode` function. The input, `x`, are a list of labels. Implement the function to return the list of labels as One-Hot encoded Numpy array. The possible values for labels are 0 to 9. The one-hot encoding function should return the same encoding for each value between each call to `one_hot_encode`. Make sure to save the map of encodings outside the function.
#
# Hint: Don't reinvent the wheel.
# + id="CSMy2fHWsmk3" colab={"base_uri": "https://localhost:8080/"} outputId="770e666e-796a-4981-8f02-66f0fb4900d1"
from sklearn.preprocessing import OneHotEncoder
def one_hot_encode(x):
"""
One hot encode a list of sample labels. Return a one-hot encoded vector for each label.
: x: List of sample Labels
: return: Numpy array of one-hot encoded labels
"""
#display(x)
onehot_encoder = np.eye(10)[x]
# TODO: Implement Function
return onehot_encoder
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
test_one_hot_encode(one_hot_encode)
# + [markdown] id="szxlQP5osmk5"
# ### Randomize Data
# As you saw from exploring the data above, the order of the samples are randomized. It doesn't hurt to randomize it again, but you don't need to for this dataset.
# + [markdown] id="FnvI7ogWsmk8"
# ## Preprocess all the data and save it
# Running the code cell below will preprocess all the CIFAR-10 data and save it to file. The code below also uses 10% of the training data for validation.
# + id="F6CTd9zhsmk-" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="4d63d05a-96a4-460f-a650-8d9d2e412df8"
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# Preprocess Training, Validation, and Testing Data
preprocess_and_save_data(cifar10_dataset_folder_path, normalize, one_hot_encode)
# + [markdown] id="BhPmB0OosmlA"
# # Check Point
# This is your first checkpoint. If you ever decide to come back to this notebook or have to restart the notebook, you can start from here. The preprocessed data has been saved to disk.
# + id="VTYFo-D0smlC"
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import pickle
#import problem_unittests as tests
import helper
# Load the Preprocessed Validation data
valid_features, valid_labels = pickle.load(open('preprocess_validation.p', mode='rb'))
# + [markdown] id="U5WNCTBysmlE"
# ## Build the network
# For the neural network, you'll build each layer into a function. Most of the code you've seen has been outside of functions. To test your code more thoroughly, we require that you put each layer in a function. This allows us to give you better feedback and test for simple mistakes using our unittests before you submit your project.
#
# >**Note:** If you're finding it hard to dedicate enough time for this course each week, we've provided a small shortcut to this part of the project. In the next couple of problems, you'll have the option to use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages to build each layer, except the layers you build in the "Convolutional and Max Pooling Layer" section. TF Layers is similar to Keras's and TFLearn's abstraction to layers, so it's easy to pickup.
#
# >However, if you would like to get the most out of this course, try to solve all the problems _without_ using anything from the TF Layers packages. You **can** still use classes from other packages that happen to have the same name as ones you find in TF Layers! For example, instead of using the TF Layers version of the `conv2d` class, [tf.layers.conv2d](https://www.tensorflow.org/api_docs/python/tf/layers/conv2d), you would want to use the TF Neural Network version of `conv2d`, [tf.nn.conv2d](https://www.tensorflow.org/api_docs/python/tf/nn/conv2d).
#
# Let's begin!
#
# ### Input
# The neural network needs to read the image data, one-hot encoded labels, and dropout keep probability. Implement the following functions
# * Implement `neural_net_image_input`
# * Return a [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder)
# * Set the shape using `image_shape` with batch size set to `None`.
# * Name the TensorFlow placeholder "x" using the TensorFlow `name` parameter in the [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder).
# * Implement `neural_net_label_input`
# * Return a [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder)
# * Set the shape using `n_classes` with batch size set to `None`.
# * Name the TensorFlow placeholder "y" using the TensorFlow `name` parameter in the [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder).
# * Implement `neural_net_keep_prob_input`
# * Return a [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder) for dropout keep probability.
# * Name the TensorFlow placeholder "keep_prob" using the TensorFlow `name` parameter in the [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder).
#
# These names will be used at the end of the project to load your saved model.
#
# Note: `None` for shapes in TensorFlow allow for a dynamic size.
# + id="i6ui94FksmlH" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="355ea248-7fbe-49e0-cf2d-aedbfaacbc84"
#import tensorflow as tf
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
def neural_net_image_input(image_shape):
"""
Return a Tensor for a batch of image input
: image_shape: Shape of the images
: return: Tensor for image input.
"""
x = tf.compat.v1.placeholder(tf.float32, shape=[None,image_shape[0],image_shape[1],image_shape[2]], name='x')
# TODO: Implement Function
return x
def neural_net_label_input(n_classes):
"""
Return a Tensor for a batch of label input
: n_classes: Number of classes
: return: Tensor for label input.
"""
# TODO: Implement Function
y = tf.compat.v1.placeholder(tf.int32, [None,n_classes], name='y')
return y
def neural_net_keep_prob_input():
"""
Return a Tensor for keep probability
: return: Tensor for keep probability.
"""
# TODO: Implement Function
keep_prob = tf.compat.v1.placeholder(tf.int32, name='keep_prob')
return keep_prob
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
#tf.reset_default_graph()
#test_nn_image_inputs(neural_net_image_input)
#test_nn_label_inputs(neural_net_label_input)
#test_nn_keep_prob_inputs(neural_net_keep_prob_input)
# + [markdown] id="K0QECKCLsmlK"
# ### Convolution and Max Pooling Layer
# Convolution layers have a lot of success with images. For this code cell, you should implement the function `conv2d_maxpool` to apply convolution then max pooling:
# * Create the weight and bias using `conv_ksize`, `conv_num_outputs` and the shape of `x_tensor`.
# * Apply a convolution to `x_tensor` using weight and `conv_strides`.
# * We recommend you use same padding, but you're welcome to use any padding.
# * Add bias
# * Add a nonlinear activation to the convolution.
# * Apply Max Pooling using `pool_ksize` and `pool_strides`.
# * We recommend you use same padding, but you're welcome to use any padding.
#
# **Note:** You **can't** use [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) for **this** layer, but you can still use TensorFlow's [Neural Network](https://www.tensorflow.org/api_docs/python/tf/nn) package. You may still use the shortcut option for all the **other** layers.
# + id="YjMzIwjHsmlM" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e58cdda5-bb08-4523-d9af-00cc6e3439fa"
#tf.compat.v1.reset_default_graph()
def conv2d_maxpool(x_tensor, conv_num_outputs, conv_ksize, conv_strides, pool_ksize, pool_strides,W,b):
"""
Apply convolution then max pooling to x_tensor
:param x_tensor: TensorFlow Tensor
:param conv_num_outputs: Number of outputs for the convolutional layer
:param conv_ksize: kernal size 2-D Tuple for the convolutional layer
:param conv_strides: Stride 2-D Tuple for convolution
:param pool_ksize: kernal size 2-D Tuple for pool
:param pool_strides: Stride 2-D Tuple for pool
: return: A tensor that represents convolution and max pooling of x_tensor
"""
# TODO: Implement Function
#display(x_tensor.shape[0])
x = tf.nn.conv2d(x_tensor, W, strides=[1, conv_strides[0], conv_strides[1], 3], padding='SAME')
x = tf.nn.bias_add(x, b)
x = tf.nn.relu(x)
x= tf.nn.max_pool(x, ksize=[1, pool_ksize[0], pool_ksize[1], 3], strides=[1,pool_strides[0] , pool_strides[1], 1],padding='SAME')
return x
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
#tf.compat.v1.reset_default_graph()
#test_con_pool(conv2d_maxpool)
# + [markdown] id="uzSt5KaXsmlO"
# ### Flatten Layer
# Implement the `flatten` function to change the dimension of `x_tensor` from a 4-D tensor to a 2-D tensor. The output should be the shape (*Batch Size*, *Flattened Image Size*). Shortcut option: you can use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages for this layer. For more of a challenge, only use other TensorFlow packages.
# + id="klWi-wCRsmlP" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5c3c811f-4b18-4d1d-9f10-4e8d1609ac72"
def flatten(x_tensor):
"""
Flatten x_tensor to (Batch Size, Flattened Image Size)
: x_tensor: A tensor of size (Batch Size, ...), where ... are the image dimensions.
: return: A tensor of size (Batch Size, Flattened Image Size).
"""
# TODO: Implement Function
fc1 = tf.reshape(x_tensor, [-1,128*4*4])
return fc1
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
#test_flatten(flatten)
# + [markdown] id="fAkpKF4ismlR"
# ### Fully-Connected Layer
# Implement the `fully_conn` function to apply a fully connected layer to `x_tensor` with the shape (*Batch Size*, *num_outputs*). Shortcut option: you can use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages for this layer. For more of a challenge, only use other TensorFlow packages.
# + id="X2_gt_65smlT" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="78a169b0-f8c4-4c39-ce1f-cb81ebd54f76"
def fully_conn(x_tensor, num_outputs,W,b):
"""
Apply a fully connected layer to x_tensor using weight and bias
: x_tensor: A 2-D tensor where the first dimension is batch size.
: num_outputs: The number of output that the new tensor should be.
: return: A 2-D tensor where the second dimension is num_outputs.
"""
# TODO: Implement Function
out = tf.add(tf.matmul(x_tensor, W), b)
return out
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
#tf.compat.v1.reset_default_graph()
#test_fully_conn(fully_conn)
# + [markdown] id="PScNi2kssmlV"
# ### Output Layer
# Implement the `output` function to apply a fully connected layer to `x_tensor` with the shape (*Batch Size*, *num_outputs*). Shortcut option: you can use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages for this layer. For more of a challenge, only use other TensorFlow packages.
#
# **Note:** Activation, softmax, or cross entropy should **not** be applied to this.
# + id="AdIKE8m1smlX" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b09f2b10-f92f-48e3-c5d8-dfd86692c8bd"
def output(x_tensor, num_outputs,W,b):
"""
Apply an output layer to x_tensor using weight and bias
: x_tensor: A 2-D tensor where the first dimension is batch size.
: num_outputs: The number of output that the new tensor should be.
: return: A 2-D tensor where the second dimension is num_outputs.
"""
# TODO: Implement Function
# TODO: Implement Function
out = tf.add(tf.matmul(x_tensor, W), b)
return out
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
#test_output(output)
# + [markdown] id="Chy2AVXHsmlY"
# ### Create Convolutional Model
# Implement the function `conv_net` to create a convolutional neural network model. The function takes in a batch of images, `x`, and outputs logits. Use the layers you created above to create this model:
#
# * Apply 1, 2, or 3 Convolution and Max Pool layers
# * Apply a Flatten Layer
# * Apply 1, 2, or 3 Fully Connected Layers
# * Apply an Output Layer
# * Return the output
# * Apply [TensorFlow's Dropout](https://www.tensorflow.org/api_docs/python/tf/nn/dropout) to one or more layers in the model using `keep_prob`.
# + id="8m-3Hk9Nsmla" colab={"base_uri": "https://localhost:8080/", "height": 497} outputId="b2929da6-be9a-45b7-e5ea-171a887b3833"
def conv_net(x, keep_prob):
"""
Create a convolutional neural network model
: x: Placeholder tensor that holds image data.
: keep_prob: Placeholder tensor that hold dropout keep probability.
: return: Tensor that represents logits
"""
# TODO: Apply 1, 2, or 3 Convolution and Max Pool layers
# Play around with different number of outputs, kernel size and stride
# Function Definition from Above:
display('weights shape')
with tf.variable_scope("other", reuse=tf.AUTO_REUSE) as scope:
W0 = tf.get_variable('W0', shape=(3,3,3,32), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
display(W0.shape)
B0 = tf.get_variable('B0', shape=(32), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
W1 = tf.get_variable('W1', shape=(3,3,32,64), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
display(W1.shape)
B1 = tf.get_variable('B1', shape=(64), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
W2 = tf.get_variable('W2', shape=(3,3,64,128), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
display(W2.shape)
B2 = tf.get_variable('B2', shape=(128), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
W3 = tf.get_variable('W3', shape=(128*4*4,128), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
display(W3.shape)
B3 = tf.get_variable('B3', shape=(128), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
W4 = tf.get_variable('W4', shape=(128,10), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
display(W4.shape)
B4 = tf.get_variable('B4', shape=(10), initializer=tf.compat.v1.keras.initializers.glorot_uniform())
display('Convolution shape')
conv1 = conv2d_maxpool(x, 10, [3,3], [1,1], [2,2], [1,1],W0,B0)
display(conv1.shape)
conv2 = conv2d_maxpool(conv1, 10, [3,3], [1,1], [2,2], [1,1],W1,B1)
display(conv2.shape)
conv3 = conv2d_maxpool(conv2, 10, [3,3], [1,1], [2,2], [1,1],W2,B2)
display(conv3.shape)
# TODO: Apply a Flatten Layer
# Function Definition from Above:
# flatten(x_tensor)
flat1 = flatten(conv3)
display(flat1.shape)
# TODO: Apply 1, 2, or 3 Fully Connected Layers
# Play around with different number of outputs
# Function Definition from Above:
# fully_conn(x_tensor, num_outputs)
full1 = fully_conn(flat1,10,W3,B3)
#full2 = fully_conn(full1,10,weights['wd1'],biases['bd1'])
display(full1.shape)
# TODO: Apply an Output Layer
# Set this to the number of classes
# Function Definition from Above:
# output(x_tensor, num_outputs)
out = output(full1,10,W4,B4)
display(out.shape)
# TODO: return output
return out
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
##############################
## Build the Neural Network ##
##############################
# Remove previous weights, bias, inputs, etc..
tf.reset_default_graph()
tf.compat.v1.disable_eager_execution()
# Inputs
x = neural_net_image_input((32, 32, 3))
y = neural_net_label_input(10)
keep_prob = neural_net_keep_prob_input()
display(x.shape)
# Model
logits = conv_net(x, keep_prob)
# Name logits Tensor, so that is can be loaded from disk after training
logits = tf.identity(logits, name='logits')
# Loss and Optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=y))
optimizer = tf.train.AdamOptimizer().minimize(cost)
# Accuracy
correct_pred = tf.equal(tf.argmax(logits, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32), name='accuracy')
test_conv_net(conv_net)
# + [markdown] id="O6t5YeGHsmlc"
# ## Train the Neural Network
# ### Single Optimization
# Implement the function `train_neural_network` to do a single optimization. The optimization should use `optimizer` to optimize in `session` with a `feed_dict` of the following:
# * `x` for image input
# * `y` for labels
# * `keep_prob` for keep probability for dropout
#
# This function will be called for each batch, so `tf.global_variables_initializer()` has already been called.
#
# Note: Nothing needs to be returned. This function is only optimizing the neural network.
# + id="r0xSboqnsmld" colab={"base_uri": "https://localhost:8080/", "height": 563} outputId="258a2551-629f-498b-9871-01b1e14a8119"
def train_neural_network(session, optimizer, keep_probability, feature_batch, label_batch):
"""
Optimize the session on a batch of images and labels
: session: Current TensorFlow session
: optimizer: TensorFlow optimizer function
: keep_probability: keep probability
: feature_batch: Batch of Numpy image data
: label_batch: Batch of Numpy label data
"""
# TODO: Implement Function
pred = conv_net(feature_batch,keep_probability)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=pred, labels=y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
pass
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
test_train_nn(train_neural_network)
# + [markdown] id="s637f5CQsmle"
# ### Show Stats
# Implement the function `print_stats` to print loss and validation accuracy. Use the global variables `valid_features` and `valid_labels` to calculate validation accuracy. Use a keep probability of `1.0` to calculate the loss and validation accuracy.
# + id="tXT0Ql90smlf"
def print_stats(session, feature_batch, label_batch, cost, accuracy):
"""
Print information about loss and validation accuracy
: session: Current TensorFlow session
: feature_batch: Batch of Numpy image data
: label_batch: Batch of Numpy label data
: cost: TensorFlow cost function
: accuracy: TensorFlow accuracy function
"""
# TODO: Implement Function
pass
# + [markdown] id="CFwE7Tjesmlg"
# ### Hyperparameters
# Tune the following parameters:
# * Set `epochs` to the number of iterations until the network stops learning or start overfitting
# * Set `batch_size` to the highest number that your machine has memory for. Most people set them to common sizes of memory:
# * 64
# * 128
# * 256
# * ...
# * Set `keep_probability` to the probability of keeping a node using dropout
# + id="-yY82kbSsmlh"
# TODO: Tune Parameters
epochs = None
batch_size = None
keep_probability = None
# + [markdown] id="lLFhlge-smlj"
# ### Train on a Single CIFAR-10 Batch
# Instead of training the neural network on all the CIFAR-10 batches of data, let's use a single batch. This should save time while you iterate on the model to get a better accuracy. Once the final validation accuracy is 50% or greater, run the model on all the data in the next section.
# + id="o4gOXncSsmll"
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
print('Checking the Training on a Single Batch...')
with tf.Session() as sess:
# Initializing the variables
sess.run(tf.global_variables_initializer())
# Training cycle
for epoch in range(epochs):
batch_i = 1
for batch_features, batch_labels in helper.load_preprocess_training_batch(batch_i, batch_size):
train_neural_network(sess, optimizer, keep_probability, batch_features, batch_labels)
print('Epoch {:>2}, CIFAR-10 Batch {}: '.format(epoch + 1, batch_i), end='')
print_stats(sess, batch_features, batch_labels, cost, accuracy)
# + [markdown] id="oCwiRovxsmlm"
# ### Fully Train the Model
# Now that you got a good accuracy with a single CIFAR-10 batch, try it with all five batches.
# + id="3VVqFZINsmln"
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
save_model_path = './image_classification'
print('Training...')
with tf.Session() as sess:
# Initializing the variables
sess.run(tf.global_variables_initializer())
# Training cycle
for epoch in range(epochs):
# Loop over all batches
n_batches = 5
for batch_i in range(1, n_batches + 1):
for batch_features, batch_labels in helper.load_preprocess_training_batch(batch_i, batch_size):
train_neural_network(sess, optimizer, keep_probability, batch_features, batch_labels)
print('Epoch {:>2}, CIFAR-10 Batch {}: '.format(epoch + 1, batch_i), end='')
print_stats(sess, batch_features, batch_labels, cost, accuracy)
# Save Model
saver = tf.train.Saver()
save_path = saver.save(sess, save_model_path)
# + [markdown] id="vRhaQaXVsmlo"
# # Checkpoint
# The model has been saved to disk.
# ## Test Model
# Test your model against the test dataset. This will be your final accuracy. You should have an accuracy greater than 50%. If you don't, keep tweaking the model architecture and parameters.
# + id="srLp7Fvlsmlp"
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import tensorflow as tf
import pickle
import helper
import random
# Set batch size if not already set
try:
if batch_size:
pass
except NameError:
batch_size = 64
save_model_path = './image_classification'
n_samples = 4
top_n_predictions = 3
def test_model():
"""
Test the saved model against the test dataset
"""
test_features, test_labels = pickle.load(open('preprocess_test.p', mode='rb'))
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
# Load model
loader = tf.train.import_meta_graph(save_model_path + '.meta')
loader.restore(sess, save_model_path)
# Get Tensors from loaded model
loaded_x = loaded_graph.get_tensor_by_name('x:0')
loaded_y = loaded_graph.get_tensor_by_name('y:0')
loaded_keep_prob = loaded_graph.get_tensor_by_name('keep_prob:0')
loaded_logits = loaded_graph.get_tensor_by_name('logits:0')
loaded_acc = loaded_graph.get_tensor_by_name('accuracy:0')
# Get accuracy in batches for memory limitations
test_batch_acc_total = 0
test_batch_count = 0
for test_feature_batch, test_label_batch in helper.batch_features_labels(test_features, test_labels, batch_size):
test_batch_acc_total += sess.run(
loaded_acc,
feed_dict={loaded_x: test_feature_batch, loaded_y: test_label_batch, loaded_keep_prob: 1.0})
test_batch_count += 1
print('Testing Accuracy: {}\n'.format(test_batch_acc_total/test_batch_count))
# Print Random Samples
random_test_features, random_test_labels = tuple(zip(*random.sample(list(zip(test_features, test_labels)), n_samples)))
random_test_predictions = sess.run(
tf.nn.top_k(tf.nn.softmax(loaded_logits), top_n_predictions),
feed_dict={loaded_x: random_test_features, loaded_y: random_test_labels, loaded_keep_prob: 1.0})
helper.display_image_predictions(random_test_features, random_test_labels, random_test_predictions)
test_model()
# + [markdown] id="TU0wflGIsmlr"
# ## Why 50-80% Accuracy?
# You might be wondering why you can't get an accuracy any higher. First things first, 50% isn't bad for a simple CNN. Pure guessing would get you 10% accuracy. However, you might notice people are getting scores [well above 80%](http://rodrigob.github.io/are_we_there_yet/build/classification_datasets_results.html#43494641522d3130). That's because we haven't taught you all there is to know about neural networks. We still need to cover a few more techniques.
# ## Submitting This Project
# When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_image_classification.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "helper.py" and "problem_unittests.py" files in your submission.
|
image-classification/dlnd_image_classification_copy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.8 64-bit
# language: python
# name: python3
# ---
# # Assignment - Index splitting (take home)
#
# 1. Create an array of shape (100,100,3) of unsigned integer 8 bits filled with random values between 0 to 256.
# 2. Find the values that are less than 30, 50, 100, 250.
# 3. Split the array into 4 parts vertically and horizontally
# 4. Copy array from Question 1 and change the values that are les than 110 to 0 and others to 255
# 5. Hint: Use random seed of 12345
import numpy as np
np.random.seed(12345)
ar100x100x3 = np.random.randint(0,255,(100,100,3),dtype=np.uint8)
a30 = ar100x100x3<30
ar100x100x3[a30]
# print(a30)
a50 = ar100x100x3 < 50
print(ar100x100x3[a30])
# print(a30)
a100 = ar100x100x3 < 100
print(ar100x100x3[a100])
a250 = ar100x100x3 < 250
print(ar100x100x3[a250])
#print(a50)
a_4h = np.hsplit(ar100x100x3,4)
print(len(a_4h))
a_4v = np.vsplit(ar100x100x3,4)
print(len(a_4v))
a_5h = np.hsplit(ar100x100x3,4)
print(len(a_4h))
aryNew = ar100x100x3.copy()
aryNew = np.where(aryNew<110,0,255)
print(aryNew)
|
advance/cv-master/Assignments/a07.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import
import sys
import os
try:
from dotenv import find_dotenv, load_dotenv
except:
pass
import argparse
try:
sys.path.append(os.path.join(os.path.dirname(__file__), '../src'))
except:
sys.path.append(os.path.join(os.getcwd(), '../src'))
import pandas as pd
import numpy as np
import torch
import torch.nn as nn
from torchcontrib.optim import SWA
from torch.optim import Adam, SGD
from torch.optim.lr_scheduler import CosineAnnealingLR, ReduceLROnPlateau, CyclicLR, \
CosineAnnealingWarmRestarts
from consNLP.data import load_data, data_utils, fetch_dataset
from consNLP.models import transformer_models, activations, layers, losses, scorers
from consNLP.visualization import visualize
from consNLP.trainer.trainer import BasicTrainer, PLTrainer, test_pl_trainer
from consNLP.trainer.trainer_utils import set_seed, _has_apex, _torch_lightning_available, _has_wandb, _torch_gpu_available, _num_gpus, _torch_tpu_available
from consNLP.preprocessing.custom_tokenizer import BERTweetTokenizer
if _has_apex:
#from torch.cuda import amp
from apex import amp
if _torch_tpu_available:
import torch_xla
import torch_xla.core.xla_model as xm
import torch_xla.distributed.xla_multiprocessing as xmp
if _has_wandb:
import wandb
try:
load_dotenv(find_dotenv())
wandb.login(key=os.environ['WANDB_API_KEY'])
except:
_has_wandb = False
if _torch_lightning_available:
import pytorch_lightning as pl
from pytorch_lightning import Trainer, seed_everything
from pytorch_lightning.loggers import WandbLogger
from pytorch_lightning.metrics.metric import NumpyMetric
from pytorch_lightning.callbacks import ModelCheckpoint, EarlyStopping, Callback
import tokenizers
from transformers import AutoModel, AutoTokenizer, AdamW, get_linear_schedule_with_warmup, AutoConfig
# -
load_dotenv(find_dotenv())
fetch_dataset(project_dir='../',download_from_kaggle=True,\
kaggle_dataset='lakshmi25npathi/imdb-dataset-of-50k-movie-reviews')
# +
parser = argparse.ArgumentParser(prog='Torch trainer function',conflict_handler='resolve')
parser.add_argument('--train_data', type=str, default='../data/raw/IMDB Dataset.csv', required=False,
help='train data')
parser.add_argument('--val_data', type=str, default='', required=False,
help='validation data')
parser.add_argument('--test_data', type=str, default=None, required=False,
help='test data')
parser.add_argument('--task_type', type=str, default='binary_sequence_classification', required=False,
help='type of task')
parser.add_argument('--transformer_model_pretrained_path', type=str, default='roberta-base', required=False,
help='transformer model pretrained path or huggingface model name')
parser.add_argument('--transformer_config_path', type=str, default='roberta-base', required=False,
help='transformer config file path or huggingface model name')
parser.add_argument('--transformer_tokenizer_path', type=str, default='roberta-base', required=False,
help='transformer tokenizer file path or huggingface model name')
parser.add_argument('--bpe_vocab_path', type=str, default='', required=False,
help='bytepairencoding vocab file path')
parser.add_argument('--bpe_merges_path', type=str, default='', required=False,
help='bytepairencoding merges file path')
parser.add_argument('--berttweettokenizer_path', type=str, default='', required=False,
help='BERTweet tokenizer path')
parser.add_argument('--max_text_len', type=int, default=100, required=False,
help='maximum length of text')
parser.add_argument('--epochs', type=int, default=5, required=False,
help='number of epochs')
parser.add_argument('--lr', type=float, default=.00003, required=False,
help='learning rate')
parser.add_argument('--loss_function', type=str, default='bcelogit', required=False,
help='loss function')
parser.add_argument('--metric', type=str, default='f1', required=False,
help='scorer metric')
parser.add_argument('--use_lightning_trainer', type=bool, default=False, required=False,
help='if lightning trainer needs to be used')
parser.add_argument('--use_torch_trainer', type=bool, default=True, required=False,
help='if custom torch trainer needs to be used')
parser.add_argument('--use_apex', type=bool, default=False, required=False,
help='if apex needs to be used')
parser.add_argument('--use_gpu', type=bool, default=False, required=False,
help='GPU mode')
parser.add_argument('--use_TPU', type=bool, default=False, required=False,
help='TPU mode')
parser.add_argument('--num_gpus', type=int, default=0, required=False,
help='Number of GPUs')
parser.add_argument('--num_tpus', type=int, default=0, required=False,
help='Number of TPUs')
parser.add_argument('--train_batch_size', type=int, default=16, required=False,
help='train batch size')
parser.add_argument('--eval_batch_size', type=int, default=16, required=False,
help='eval batch size')
parser.add_argument('--model_save_path', type=str, default='../models/sentiment_classification/', required=False,
help='seed')
parser.add_argument('--wandb_logging', type=bool, default=False, required=False,
help='wandb logging needed')
parser.add_argument('--seed', type=int, default=42, required=False,
help='seed')
args, _ = parser.parse_known_args()
print ("Wandb Logging: {}, GPU: {}, Pytorch Lightning: {}, TPU: {}, Apex: {}".format(\
_has_wandb and args.wandb_logging, _torch_gpu_available,\
_torch_lightning_available and args.use_lightning_trainer, _torch_tpu_available, _has_apex))
# +
reshape = False
final_activation = None
convert_output = None
if args.task_type == 'binary_sequence_classification':
if args.metric != 'roc_auc_score':
convert_output = 'round'
if args.loss_function == 'bcelogit':
final_activation = 'sigmoid'
elif args.task_type == 'multiclass_sequence_classification':
convert_output = 'max'
elif args.task_type == 'binary_token_classification':
reshape = True
if args.metric != 'roc_auc_score':
convert_output = 'round'
if args.loss_function == 'bcelogit':
final_activation = 'sigmoid'
elif args.task_type == 'multiclass_token_classification':
reshape = True
convert_output = 'max'
# -
df = load_data.load_pandas_df(args.train_data,sep=',')
df = df.iloc[:1000]
df.head(5)
model_save_dir = args.model_save_path
try:
os.makedirs(model_save_dir)
except OSError:
pass
df.sentiment, label2idx = data_utils.convert_categorical_label_to_int(df.sentiment, \
save_path=os.path.join(model_save_dir,'label2idx.pkl'))
df.head(5)
# +
from sklearn.model_selection import KFold
kf = KFold(5)
for train_index, val_index in kf.split(df.review, df.sentiment):
break
train_df = df.iloc[train_index].reset_index(drop=True)
val_df = df.iloc[val_index].reset_index(drop=True)
# -
train_df.shape, val_df.shape
# +
if args.berttweettokenizer_path:
tokenizer = BERTweetTokenizer(args.berttweettokenizer_path)
else:
tokenizer = AutoTokenizer.from_pretrained(args.transformer_model_pretrained_path)
if not args.berttweettokenizer_path:
try:
bpetokenizer = tokenizers.ByteLevelBPETokenizer(args.bpe_vocab_path, \
args.bpe_merges_path)
except:
bpetokenizer = None
else:
bpetokenizer = None
# +
train_dataset = data_utils.TransformerDataset(train_df.review, bpetokenizer=bpetokenizer, tokenizer=tokenizer, MAX_LEN=args.max_text_len, \
target_label=train_df.sentiment, sequence_target=False, target_text=None, conditional_label=None, conditional_all_labels=None)
val_dataset = data_utils.TransformerDataset(val_df.review, bpetokenizer=bpetokenizer, tokenizer=tokenizer, MAX_LEN=args.max_text_len, \
target_label=val_df.sentiment, sequence_target=False, target_text=None, conditional_label=None, conditional_all_labels=None)
# -
config = AutoConfig.from_pretrained(args.transformer_config_path, output_hidden_states=True, output_attentions=True)
basemodel = AutoModel.from_pretrained(args.transformer_model_pretrained_path,config=config)
model = transformer_models.TransformerWithCLS(basemodel)
# +
if _torch_tpu_available and args.use_TPU:
train_sampler = torch.utils.data.distributed.DistributedSampler(
train_dataset,
num_replicas=xm.xrt_world_size(),
rank=xm.get_ordinal(),
shuffle=True
)
val_sampler = torch.utils.data.distributed.DistributedSampler(
val_dataset,
num_replicas=xm.xrt_world_size(),
rank=xm.get_ordinal(),
shuffle=False
)
if _torch_tpu_available and args.use_TPU:
train_data_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=args.train_batch_size, sampler=train_sampler,
drop_last=True,num_workers=2)
val_data_loader = torch.utils.data.DataLoader(
val_dataset, batch_size=args.eval_batch_size, sampler=val_sampler,
drop_last=False,num_workers=1)
else:
train_data_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=args.train_batch_size)
val_data_loader = torch.utils.data.DataLoader(
val_dataset, batch_size=args.eval_batch_size)
# -
# ### Run with Pytorch Trainer
# +
if args.use_torch_trainer:
device = torch.device("cuda" if _torch_gpu_available and args.use_gpu else "cpu")
if _torch_tpu_available and args.use_TPU:
device=xm.xla_device()
print ("Device: {}".format(device))
if args.use_TPU and _torch_tpu_available and args.num_tpus > 1:
train_data_loader = torch_xla.distributed.parallel_loader.ParallelLoader(train_data_loader, [device])
train_data_loader = train_data_loader.per_device_loader(device)
trainer = BasicTrainer(model, train_data_loader, val_data_loader, device, args.transformer_model_pretrained_path, \
final_activation=final_activation, \
test_data_loader=val_data_loader)
param_optimizer = list(trainer.model.named_parameters())
no_decay = ["bias", "LayerNorm.bias", "LayerNorm.weight"]
optimizer_parameters = [
{
"params": [
p for n, p in param_optimizer if not any(nd in n for nd in no_decay)
],
"weight_decay": 0.001,
},
{
"params": [
p for n, p in param_optimizer if any(nd in n for nd in no_decay)
],
"weight_decay": 0.0,
},
]
num_train_steps = int(len(train_data_loader) * args.epochs)
if _torch_tpu_available and args.use_TPU:
optimizer = AdamW(optimizer_parameters, lr=args.lr*xm.xrt_world_size())
else:
optimizer = AdamW(optimizer_parameters, lr=args.lr)
if args.use_apex and _has_apex:
model, optimizer = amp.initialize(model, optimizer, opt_level="O1")
scheduler = get_linear_schedule_with_warmup(optimizer, num_warmup_steps=0, num_training_steps=num_train_steps)
loss = losses.get_loss(args.loss_function)
scorer = scorers.SKMetric(args.metric, convert=convert_output, reshape=reshape)
def _mp_fn(rank, flags, trainer, epochs, lr, metric, loss_function, optimizer, scheduler, model_save_path, num_gpus, num_tpus, \
max_grad_norm, early_stopping_rounds, snapshot_ensemble, is_amp, use_wandb, seed):
torch.set_default_tensor_type('torch.FloatTensor')
a = trainer.train(epochs, lr, metric, loss_function, optimizer, scheduler, model_save_path, num_gpus, num_tpus, \
max_grad_norm, early_stopping_rounds, snapshot_ensemble, is_amp, use_wandb, seed)
FLAGS = {}
if _torch_tpu_available and args.use_TPU:
xmp.spawn(_mp_fn, args=(FLAGS, trainer, args.epochs, args.lr, scorer, loss, optimizer, scheduler, args.model_save_path, args.num_gpus, args.num_tpus, \
1, 3, False, args.use_apex, False, args.seed), nprocs=8, start_method='fork')
else:
use_wandb = _has_wandb and args.wandb_logging
trainer.train(args.epochs, args.lr, scorer, loss, optimizer, scheduler, args.model_save_path, args.num_gpus, args.num_tpus, \
max_grad_norm=1, early_stopping_rounds=3, snapshot_ensemble=False, is_amp=args.use_apex, use_wandb=use_wandb, seed=args.seed)
elif args.use_lightning_trainer and _torch_lightning_available:
from pytorch_lightning import Trainer, seed_everything
seed_everything(args.seed)
loss = losses.get_loss(args.loss_function)
scorer = scorers.PLMetric(args.metric, convert=convert_output, reshape=reshape)
log_args = {'description': args.transformer_model_pretrained_path, 'loss': loss.__class__.__name__, 'epochs': args.epochs, 'learning_rate': args.lr}
if _has_wandb and not _torch_tpu_available and args.wandb_logging:
wandb.init(project="Project",config=log_args)
wandb_logger = WandbLogger()
checkpoint_callback = ModelCheckpoint(
filepath=args.model_save_path,
save_top_k=1,
verbose=True,
monitor='val_loss',
mode='min'
)
earlystop = EarlyStopping(
monitor='val_loss',
patience=3,
verbose=False,
mode='min'
)
if args.use_gpu and _torch_gpu_available:
print ("using GPU")
if args.wandb_logging:
if _has_apex:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, logger=wandb_logger, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, logger=wandb_logger, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
if _has_apex:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
elif args.use_TPU and _torch_tpu_available:
print ("using TPU")
if _has_apex:
trainer = Trainer(num_tpu_cores=args.num_tpus, max_epochs=args.epochs, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(num_tpu_cores=args.num_tpus, max_epochs=args.epochs, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
print ("using CPU")
if args.wandb_logging:
if _has_apex:
trainer = Trainer(max_epochs=args.epochs, logger=wandb_logger, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(max_epochs=args.epochs, logger=wandb_logger, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
if _has_apex:
trainer = Trainer(max_epochs=args.epochs, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(max_epochs=args.epochs, checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
num_train_steps = int(len(train_data_loader) * args.epochs)
pltrainer = PLTrainer(num_train_steps, model, scorer, loss, args.lr, \
final_activation=final_activation, seed=42)
#try:
# print ("Loaded model from previous checkpoint")
# pltrainer = PLTrainer.load_from_checkpoint(args.model_save_path)
#except:
# pass
trainer.fit(pltrainer, train_data_loader, val_data_loader)
# -
test_output1 = trainer.test_output
# ### Run with Pytorch Lightning Trainer
# +
parser = argparse.ArgumentParser(prog='Torch trainer function',conflict_handler='resolve')
parser.add_argument('--train_data', type=str, default='../data/raw/IMDB Dataset.csv', required=False,
help='train data')
parser.add_argument('--val_data', type=str, default='', required=False,
help='validation data')
parser.add_argument('--test_data', type=str, default=None, required=False,
help='test data')
parser.add_argument('--transformer_model_pretrained_path', type=str, default='roberta-base', required=False,
help='transformer model pretrained path or huggingface model name')
parser.add_argument('--transformer_config_path', type=str, default='roberta-base', required=False,
help='transformer config file path or huggingface model name')
parser.add_argument('--transformer_tokenizer_path', type=str, default='roberta-base', required=False,
help='transformer tokenizer file path or huggingface model name')
parser.add_argument('--bpe_vocab_path', type=str, default='', required=False,
help='bytepairencoding vocab file path')
parser.add_argument('--bpe_merges_path', type=str, default='', required=False,
help='bytepairencoding merges file path')
parser.add_argument('--berttweettokenizer_path', type=str, default='', required=False,
help='BERTweet tokenizer path')
parser.add_argument('--max_text_len', type=int, default=100, required=False,
help='maximum length of text')
parser.add_argument('--epochs', type=int, default=5, required=False,
help='number of epochs')
parser.add_argument('--lr', type=float, default=.00003, required=False,
help='learning rate')
parser.add_argument('--loss_function', type=str, default='bcelogit', required=False,
help='loss function')
parser.add_argument('--metric', type=str, default='f1', required=False,
help='scorer metric')
parser.add_argument('--use_lightning_trainer', type=bool, default=True, required=False,
help='if lightning trainer needs to be used')
parser.add_argument('--use_torch_trainer', type=bool, default=False, required=False,
help='if custom torch trainer needs to be used')
parser.add_argument('--use_apex', type=bool, default=False, required=False,
help='if apex needs to be used')
parser.add_argument('--use_gpu', type=bool, default=False, required=False,
help='GPU mode')
parser.add_argument('--use_TPU', type=bool, default=False, required=False,
help='TPU mode')
parser.add_argument('--num_gpus', type=int, default=0, required=False,
help='Number of GPUs')
parser.add_argument('--num_tpus', type=int, default=0, required=False,
help='Number of TPUs')
parser.add_argument('--train_batch_size', type=int, default=16, required=False,
help='train batch size')
parser.add_argument('--eval_batch_size', type=int, default=16, required=False,
help='eval batch size')
parser.add_argument('--model_save_path', type=str, default='../models/sentiment_classification/', required=False,
help='seed')
parser.add_argument('--wandb_logging', type=bool, default=False, required=False,
help='wandb logging needed')
parser.add_argument('--seed', type=int, default=42, required=False,
help='seed')
args, _ = parser.parse_known_args()
print ("Wandb Logging: {}, GPU: {}, Pytorch Lightning: {}, TPU: {}, Apex: {}".format(\
_has_wandb and args.wandb_logging, _torch_gpu_available,\
_torch_lightning_available and args.use_lightning_trainer, _torch_tpu_available, _has_apex))
# +
if args.use_torch_trainer:
device = torch.device("cuda" if _torch_gpu_available and args.use_gpu else "cpu")
if _torch_tpu_available and args.use_TPU:
device=xm.xla_device()
print ("Device: {}".format(device))
if args.use_TPU and _torch_tpu_available and args.num_tpus > 1:
train_data_loader = torch_xla.distributed.parallel_loader.ParallelLoader(train_data_loader, [device])
train_data_loader = train_data_loader.per_device_loader(device)
trainer = BasicTrainer(model, train_data_loader, val_data_loader, device, args.transformer_model_pretrained_path, \
final_activation=final_activation, \
test_data_loader=val_data_loader)
param_optimizer = list(trainer.model.named_parameters())
no_decay = ["bias", "LayerNorm.bias", "LayerNorm.weight"]
optimizer_parameters = [
{
"params": [
p for n, p in param_optimizer if not any(nd in n for nd in no_decay)
],
"weight_decay": 0.001,
},
{
"params": [
p for n, p in param_optimizer if any(nd in n for nd in no_decay)
],
"weight_decay": 0.0,
},
]
num_train_steps = int(len(train_data_loader) * args.epochs)
if _torch_tpu_available and args.use_TPU:
optimizer = AdamW(optimizer_parameters, lr=args.lr*xm.xrt_world_size())
else:
optimizer = AdamW(optimizer_parameters, lr=args.lr)
if args.use_apex and _has_apex:
model, optimizer = amp.initialize(model, optimizer, opt_level="O1")
scheduler = get_linear_schedule_with_warmup(optimizer, num_warmup_steps=0, num_training_steps=num_train_steps)
loss = losses.get_loss(args.loss_function)
scorer = scorers.SKMetric(args.metric, convert=convert_output, reshape=reshape)
def _mp_fn(rank, flags, trainer, epochs, lr, metric, loss_function, optimizer, scheduler, model_save_path, num_gpus, num_tpus, \
max_grad_norm, early_stopping_rounds, snapshot_ensemble, is_amp, use_wandb, seed):
torch.set_default_tensor_type('torch.FloatTensor')
a = trainer.train(epochs, lr, metric, loss_function, optimizer, scheduler, model_save_path, num_gpus, num_tpus, \
max_grad_norm, early_stopping_rounds, snapshot_ensemble, is_amp, use_wandb, seed)
FLAGS = {}
if _torch_tpu_available and args.use_TPU:
xmp.spawn(_mp_fn, args=(FLAGS, trainer, args.epochs, args.lr, scorer, loss, optimizer, scheduler, args.model_save_path, args.num_gpus, args.num_tpus, \
1, 3, False, args.use_apex, False, args.seed), nprocs=8, start_method='fork')
else:
use_wandb = _has_wandb and args.wandb_logging
trainer.train(args.epochs, args.lr, scorer, loss, optimizer, scheduler, args.model_save_path, args.num_gpus, args.num_tpus, \
max_grad_norm=1, early_stopping_rounds=3, snapshot_ensemble=False, is_amp=args.use_apex, use_wandb=use_wandb, seed=args.seed)
elif args.use_lightning_trainer and _torch_lightning_available:
from pytorch_lightning import Trainer, seed_everything
seed_everything(args.seed)
loss = losses.get_loss(args.loss_function)
scorer = scorers.PLMetric(args.metric, convert=convert_output, reshape=reshape)
log_args = {'description': args.transformer_model_pretrained_path, 'loss': loss.__class__.__name__, 'epochs': args.epochs, 'learning_rate': args.lr}
if _has_wandb and not _torch_tpu_available and args.wandb_logging:
wandb.init(project="Project",config=log_args)
wandb_logger = WandbLogger()
checkpoint_callback = ModelCheckpoint(
filepath=args.model_save_path,
save_top_k=1,
verbose=True,
monitor='val_loss',
mode='min'
)
earlystop = EarlyStopping(
monitor='val_loss',
patience=3,
verbose=False,
mode='min'
)
if args.use_gpu and _torch_gpu_available:
print ("using GPU")
if args.wandb_logging:
if _has_apex:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, logger=wandb_logger, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, logger=wandb_logger, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
if _has_apex:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(gpus=args.num_gpus, max_epochs=args.epochs, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
elif args.use_TPU and _torch_tpu_available:
print ("using TPU")
if _has_apex:
trainer = Trainer(num_tpu_cores=args.num_tpus, max_epochs=args.epochs, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(num_tpu_cores=args.num_tpus, max_epochs=args.epochs, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
print ("using CPU")
if args.wandb_logging:
if _has_apex:
trainer = Trainer(max_epochs=args.epochs, logger=wandb_logger, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(max_epochs=args.epochs, logger=wandb_logger, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
if _has_apex:
trainer = Trainer(max_epochs=args.epochs, precision=16, \
checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
else:
trainer = Trainer(max_epochs=args.epochs, checkpoint_callback=checkpoint_callback, callbacks=[earlystop])
num_train_steps = int(len(train_data_loader) * args.epochs)
pltrainer = PLTrainer(num_train_steps, model, scorer, loss, args.lr, \
final_activation=final_activation, seed=42)
#try:
# print ("Loaded model from previous checkpoint")
# pltrainer = PLTrainer.load_from_checkpoint(args.model_save_path)
#except:
# pass
trainer.fit(pltrainer, train_data_loader, val_data_loader)
# +
from tqdm import tqdm
test_output2 = []
for val_batch in tqdm(val_data_loader):
out = torch.sigmoid(pltrainer(val_batch)).detach().cpu().numpy()
test_output2.extend(out[:,0].tolist())
#test_output2 = np.concatenate(test_output2)
# -
test_output1 = np.array(test_output1)[:,0]
test_output2 = np.array(test_output2)
np.corrcoef(test_output1,test_output2)
|
notebooks/imdb_binary_sentiment_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Built-in libraries
from datetime import datetime, timedelta
# NumPy, SciPy and Pandas
import pandas as pd
import numpy as np
# -
def hourly_dataset(name):
"""
Constants for time period with maximum number of buildings measured simultaneously in the BDG dataset.
For more details, go to old_files/RawFeatures_BDG.ipynb
"""
BDG_STARTDATE = datetime.strptime('01/01/15 00:00', '%d/%m/%y %H:%M')
BDG_ENDDATE = datetime.strptime('30/11/15 23:00','%d/%m/%y %H:%M')
# Building Data Genome dataset
if name == 'BDG':
df = pd.read_csv('../data/raw/temp_open_utc_complete.csv', parse_dates=True,
infer_datetime_format=True, index_col=0)
# truncate the dataframe based on a pre-calculated time period, if needed
startDate = BDG_STARTDATE
endDate = BDG_ENDDATE
df = df[(df.index >= startDate) & (df.index <= endDate)]
# Washington D.C. dataset
elif name == 'DGS':
df = pd.read_csv('../data/raw/DGS_322_Buildings-15m-By_Building-DST-gap-filled-3-2-18-508pm.csv',
parse_dates=[['Building ID', 'Unnamed: 1']], infer_datetime_format=True)
# get rid of temperature column
del df['Unnamed: 2']
# update column names to match the row of building names
new_column_names = df.iloc[0,:]
df.columns = new_column_names
# get rid of rows with metadata and update index
df = df.drop([0,1,2], axis=0)
df = df.rename(columns = {'Building nan':'timestamp'})
df.index = df['timestamp'].astype('datetime64[ns]')
del df['timestamp']
df = df.astype(float)
# since the dataset is made from 15min interval readings, resample to 1 hr
df = df.resample('1H').sum()
else:
print("Please choose a valid dataset")
exit()
# save the file to csv before exit
df.to_csv('../data/processed/{}_dataset.csv'.format(name))
return df
# +
from collections import Counter
def resampleDGS():
df = pd.read_csv("../data/processed/DGS_dataset.csv", parse_dates=True, infer_datetime_format=True, index_col=0)
og_index = df.index.values
df = df.T
df_meta = pd.read_csv('../data/raw/dgs_metadata.csv')
df_aux = pd.read_csv("../data/raw/DGS_322_Buildings-15m-By_Building-DST-gap-filled-3-2-18-508pm.csv")
# get labels for all buildings
df_aux = df_aux.T
df_aux_og = df_aux.copy()
df_label = df_aux[df_aux.iloc[:, 0].isin(df.index.values)] # get id based on names
df_label = df_meta[df_meta['id'].isin(df_label.index.values)] # get label based on id
# print(c.value_counts())
cnt = Counter(df_label['espm_type_name'
])
for i in df_label['espm_type_name']:
print(cnt[i])
df_label = df_label[(df_label['espm_type_name'] == 'K-12 School') |
(df_label['espm_type_name'] == 'Other - Recreation') |
(df_label['espm_type_name'] == 'Fire Station') |
(df_label['espm_type_name'] == 'Office') |
(df_label['espm_type_name'] == 'Library') |
(df_label['espm_type_name'] == 'Other - Public Services') |
(df_label['espm_type_name'] == 'Police Station')]
# print(df_label['espm_type_name'].value_counts())
df_aux_og = df_aux_og.drop(df_aux_og.index[0:3])
df_aux_og.index = list(map(int, df_aux_og.index.values))
df_bdg_name = df_aux_og[df_aux_og.index.isin(df_label['id'])]
df = df[df.index.isin(df_bdg_name.iloc[:, 0])]
df = df.T
df.index = og_index
# df.to_csv('../data/processed/DGS_dataset.csv')
# +
# load building gnome dataset (BDG)
df_BDG = hourly_dataset('BDG')
# load dc building dataset (DC)
df_DGS = hourly_dataset('DGS')
# -
resampleDGS()
|
Preprocessing/preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %matplotlib inline
# %autoreload 2
# +
import numpy as np
import os
import time
import h5py
import keras
import pandas as pd
import math
import joblib
import json
import matplotlib.pyplot as plt
from fuel.datasets.svhn import SVHN
from IPython.display import display
from keras.layers import (Input, Dense, Lambda, Flatten, Reshape, BatchNormalization,
Activation, Dropout, Conv2D, Conv2DTranspose,
Concatenate, Add, Multiply)
from keras.regularizers import l2
from keras.initializers import RandomUniform
from keras.optimizers import RMSprop, Adam, SGD
from keras.models import Model
from keras import metrics
from keras import backend as K
from keras_tqdm import TQDMNotebookCallback
from keras.datasets import mnist
from keras.preprocessing.image import ImageDataGenerator
from resnet import *
# +
img_rows, img_cols, img_chns = 32, 32, 3
original_img_size = (img_rows, img_cols, img_chns)
num_classes = 10
learning_rate = float(os.environ.get('LEARNING_RATE', 0.001))
decay = float(os.environ.get('DECAY', 0.0))
batch_size = int(os.environ.get('BATCH_SIZE', 250))
epochs = int(os.environ.get('EPOCHS', 100))
run_num = int(os.environ.get('RUN_NUM', 0))
use_preprocessing = int(os.environ.get('USE_PREPROCESSING', 1))
file_prefix = 'results_preproc_%d_run_%d_' % (use_preprocessing, run_num)
# -
# # Load Data
# +
def get_svhn(split):
f = SVHN(which_format=2, which_sets=(split,), load_in_memory=True)
f.load()
X_dataset, y_dataset = f.data_sources
X_dataset, y_dataset = np.moveaxis(X_dataset, 1, 3), y_dataset
X_dataset = X_dataset / 255.
y_dataset = keras.utils.to_categorical(y_dataset, 10)
print ("%s - DType X=%s, y=%s" % (split, X_dataset.dtype, y_dataset.dtype))
print ("%s - Shape X=%s, y=%s" % (split, X_dataset.shape, y_dataset.shape))
return X_dataset, y_dataset
X_train_raw, y_train_raw = get_svhn('train')
validation_index = int(len(X_train_raw) * 0.85)
X_validation, y_validation = X_train_raw[validation_index:], y_train_raw[validation_index:]
X_train, y_train = X_train_raw[:validation_index], y_train_raw[:validation_index]
X_test, y_test = get_svhn('test')
print("raw", len(X_train_raw), len(y_train_raw))
print("validation", len(X_validation), len(y_validation))
print("train", len(X_train), len(y_train))
print("test", len(X_test), len(y_test))
# -
# Augment data
if use_preprocessing:
datagen = ImageDataGenerator(
zoom_range=0.10,
width_shift_range=0.1,
height_shift_range=0.1,
fill_mode='constant',
rotation_range=10)
datagen.fit(X_train)
X_aug = [X_train]
y_aug = [y_train]
for i in range(2):
X_gen, y_gen = next(datagen.flow(X_train, y_train, batch_size=len(X_train)))
X_aug.append(X_gen)
y_aug.append(y_gen)
X_train = np.concatenate(tuple(X_aug))
y_train = np.concatenate(tuple(y_aug))
del X_aug, y_aug, X_gen, y_gen
print(X_train.shape, X_train.shape)
# +
def display_grid(dataset, digit_size=32, grid_size=5, seed=None):
# Display some digits to figure out what's going on
figure = np.zeros((digit_size * grid_size, digit_size * grid_size, 3))
if seed is not None:
np.random.seed(seed)
for i in range(grid_size):
for j in range(grid_size):
digit = dataset[np.random.randint(len(dataset))]
d_x, d_y = i * digit_size, j * digit_size
figure[d_x:d_x + digit_size, d_y:d_y + digit_size, :] = digit.astype(float)
plt.figure(figsize=(5, 5))
plt.imshow(figure)
plt.show()
display_grid(X_train, seed=0)
display_grid(X_test, seed=0)
# -
def make_model():
x_input = Input(batch_shape=(None,) + original_img_size)
resnet_model = ResNet50(weights=None, pooling='avg', input_shape=original_img_size, include_top=False)
model_out = resnet_model(x_input)
out = Dense(num_classes, activation='softmax', name='fc10')(model_out)
model = Model(x_input, out, name='myresent50')
return model
def train_model(model, y_train_vals):
optimizer = Adam(lr=learning_rate, decay=decay)
model.compile(optimizer=optimizer,
loss='categorical_crossentropy',
metrics=['accuracy'])
start = time.time()
early_stopping = keras.callbacks.EarlyStopping('val_acc', min_delta=0.1, patience=20)
reduce_lr = keras.callbacks.ReduceLROnPlateau(monitor='val_acc', factor=0.1, patience=10, min_lr=0.01 * learning_rate)
callbacks=[early_stopping, reduce_lr]
if 'CMDLINE' not in os.environ:
callbacks += [TQDMNotebookCallback()]
history = model.fit(
X_train, y_train_vals,
batch_size=batch_size,
epochs=epochs,
callbacks=callbacks,
validation_data=(X_validation, y_validation),
verbose=0
)
done = time.time()
elapsed = done - start
print("Elapsed: ", elapsed)
return model, history
model = make_model()
model.summary()
# +
# Testing
# epochs=1
# model, history = train_model(model, y_train)
# history.history
# -
y_train_predict = y_train
for i in range(3):
print("Iteration", i)
model = make_model()
model.summary()
model, history = train_model(model, y_train_predict)
df = pd.DataFrame(history.history)
display(df.describe(percentiles=[0.25 * i for i in range(4)] + [0.95, 0.99]))
df.plot(figsize=(8, 6))
df.to_csv(file_prefix + ('history_iter%d' % i) + '.csv', index=False)
with open(file_prefix + 'vars.txt', 'w') as f:
f.write(str(locals()))
y_train_predict = model.predict(X_train)
y_train_predict
test_results = model.evaluate(X_test, y_test)
print(test_results)
with open('allresults.csv', 'a') as f:
line = ','.join([str(use_preprocessing), str(run_num), str(i)] + [str(x) for x in test_results])
f.write(line + '\n')
locals()
|
notebooks/label_refinery/output-1535206418/label_refinery-use_pre1-run_num4.nbconvert.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Divorce rates and their relationship with Marriage rate and Median Age Marriage
# load data and copy
library(rethinking)
options(mc.cores = parallel::detectCores())
data(WaffleDivorce)
d <- WaffleDivorce
# standardize variables
d$A <- scale( d$MedianAgeMarriage )
d$D <- scale( d$Divorce )
sd( d$MedianAgeMarriage )
m5.1 <- quap(
alist(
D ~ dnorm( mu , sigma ) ,
mu <- a + bA * A ,
a ~ dnorm( 0 , 0.2 ) ,
bA ~ dnorm( 0 , 0.5 ) ,
sigma ~ dexp( 1 )
) , data = d )
# ## Prior Predictive Simulation
set.seed(10)
prior <- extract.prior( m5.1 )
mu <- link( m5.1 , post=prior , data=list( A=c(-2,2) ) )
plot( NULL , xlim=c(-2,2) , ylim=c(-2,2) )
for ( i in 1:50 ) lines( c(-2,2) , mu[i,] , col=col.alpha("black",0.4) )
# ## Divorce vs Median Age Marriage
# compute percentile interval of mean
A_seq <- seq( from=-3 , to=3.2 , length.out=30 )
mu <- link( m5.1 , data=list(A=A_seq) )
mu.mean <- apply( mu , 2, mean )
mu.PI <- apply( mu , 2 , PI )
# plot it all
plot( D ~ A , data=d , col=rangi2 )
lines( A_seq , mu.mean , lwd=2 )
shade( mu.PI , A_seq )
d$M <- scale( d$Marriage )
m5.2 <- quap(
alist(
D ~ dnorm( mu , sigma ) ,
mu <- a + bM * M ,
a ~ dnorm( 0 , 0.2 ) ,
bM ~ dnorm( 0 , 0.5 ) ,
sigma ~ dexp( 1 )
) , data = d )
# compute percentile interval of mean
A_seq <- seq( from=-3 , to=3.2 , length.out=30 )
mu <- link( m5.2 , data=list(M=A_seq) )
mu.mean <- apply( mu , 2, mean )
mu.PI <- apply( mu , 2 , PI )
# plot it all
plot( D ~ M , data=d , col=rangi2 )
lines( A_seq , mu.mean , lwd=2 )
shade( mu.PI , A_seq )
# # Directed Acyclic Graph
library(dagitty)
dag5.1 <- dagitty( "dag {
A -> D
A -> M
M -> D
}")
coordinates(dag5.1) <- list( x=c(A=0,D=1,M=2) , y=c(A=0,D=1,M=0) )
drawdag( dag5.1 )
# ## Another DAG with a different causal implication
DMA_dag2 <- dagitty('dag{ D <- A -> M }')
coordinates(DMA_dag2) <- list( x=c(A=0,D=1,M=2) , y=c(A=0,D=1,M=0) )
drawdag(DMA_dag2)
impliedConditionalIndependencies( DMA_dag2 )
# ### The above relation translates to D is independent of A conditional on A
# ## The following is a multiple regression :-
m5.3 <- quap(
alist(
D ~ dnorm( mu , sigma ) ,
mu <- a + bM*M + bA*A ,
a ~ dnorm( 0 , 0.2 ) ,
bM ~ dnorm( 0 , 0.5 ) ,
bA ~ dnorm( 0 , 0.5 ) ,
sigma ~ dexp( 1 )
) , data = d )
precis( m5.3 )
plot( coeftab(m5.1,m5.2,m5.3), par=c("bA","bM") )
# call link without specifying new data
# so it uses original data
mu <- link( m5.3 )
# summarize samples across cases
mu_mean <- apply( mu , 2 , mean )
mu_PI <- apply( mu , 2 , PI )
mu_mean
mu_PI
D_sim <- sim( m5.3 , n=1e4 )
plot( mu_mean ~ d$D , col=rangi2 , ylim=range(mu_PI) ,
xlab="Observed divorce" , ylab="Predicted divorce" )
abline( a=0 , b=1 , lty=2 )
for ( i in 1:nrow(d) ) lines( rep(d$D[i],2) , mu_PI[,i] , col=rangi2 )
identify( x=d$D , y=mu_mean , labels=d$Loc )
N <- 100 # number of cases
x_real <- rnorm( N ) # x_real as Gaussian with mean 0 and stddev 1
x_spur <- rnorm( N , x_real ) # x_spur as Gaussian with mean=x_real
y <- rnorm( N , x_real ) # y as Gaussian with mean=x_real
d <- data.frame(y,x_real,x_spur) # bind all together in data frame
x_spur
x_real
d
pairs(d)
# # Counterfactual Plots
# ## Considering the below DAG we try to simulate the effect of A on D while also including the effect A has on M
drawdag( dag5.1 )
data(WaffleDivorce)
d <- list()
d$A <- standardize( WaffleDivorce$MedianAgeMarriage )
d$D <- standardize( WaffleDivorce$Divorce )
d$M <- standardize( WaffleDivorce$Marriage )
m5.3_A <- quap(
alist(
## A -> D <- M
D ~ dnorm( mu , sigma ) ,
mu <- a + bM*M + bA*A ,
a ~ dnorm( 0 , 0.2 ) ,
bM ~ dnorm( 0 , 0.5 ) ,
bA ~ dnorm( 0 , 0.5 ) ,
sigma ~ dexp( 1 ),
## A -> M
M ~ dnorm( mu_M , sigma_M ),
mu_M <- aM + bAM*A,
aM ~ dnorm( 0 , 0.2 ),
bAM ~ dnorm( 0 , 0.5 ),
sigma_M ~ dexp( 1 )
) , data = d )
A_seq <- seq( from=-2 , to=2 , length.out=30 )
# prep data
sim_dat <- data.frame( A=A_seq )
# ## Given the data(A_seq) we simulate M first and later use this A_seq,M to generate D
# simulate M and then D, using A_seq
s <- sim( m5.3_A , data=sim_dat , vars=c("M","D") )
# display counterfactual predictions
plot( sim_dat$A , colMeans(s$D) , ylim=c(-2,2) , type="l" ,
xlab="manipulated A" , ylab="counterfactual D" )
shade( apply(s$D,2,PI) , sim_dat$A )
mtext( "Total counterfactual effect of A on D" )
plot( sim_dat$A , colMeans(s$M) , ylim=c(-2,2) , type="l" ,
xlab="manipulated A" , ylab="counterfactual M" )
shade( apply(s$M,2,PI) , sim_dat$A )
mtext( "Total counterfactual effect of A on M" )
# ## The above counterfactual plots show that A has a direct influence on both A and M as assumed by our model
# ## What happends when we try to manipulate A without involving the causal change in M ?
# 1. Implies that we use A -> D and M -> D
# 2. The code uses simulating **only D** assuming A = 0 (average State in US )
# simulate D, using A_seq
sim_dat <- data.frame( M=seq(from=-2,to=2,length.out=30) , A=0 )
s <- sim( m5.3_A , data=sim_dat , vars="D")
plot( sim_dat$M , colMeans(s) , ylim=c(-2,2) , type="l" ,
xlab="manipulated M" , ylab="counterfactual D" )
shade( apply(s,2,PI) , sim_dat$M )
mtext( "Total counterfactual effect of M on D" )
# ### The above plot shows that M doesnt have a significant affect on D
# # Spurious Waffles
library(rethinking)
data(milk)
d <- milk
str(d)
d$K <- scale( d$kcal.per.g )
d$N <- scale( d$neocortex.perc )
d$M <- scale( log(d$mass) )
m5.5_draft <- quap(
alist(
K ~ dnorm( mu , sigma ) ,
mu <- a + bN*N ,
a ~ dnorm( 0 , 1 ) ,
bN ~ dnorm( 0 , 1 ) ,
sigma ~ dexp( 1 )
) , data=d )
d$neocortex.perc
dcc <- d[ complete.cases(d$K,d$N,d$M) , ]
m5.5_draft <- quap(
alist(
K ~ dnorm( mu , sigma ) ,
mu <- a + bN*N ,
a ~ dnorm( 0 , 1 ) ,
bN ~ dnorm( 0 , 1 ) ,
sigma ~ dexp( 1 )
) , data=dcc )
prior <- extract.prior( m5.5_draft )
xseq <- c(-2,2)
mu <- link( m5.5_draft , post=prior , data=list(N=xseq) )
plot( NULL , xlim=xseq , ylim=xseq )
for ( i in 1:50 ) lines( xseq , mu[i,] , col=col.alpha("black",0.3) )
# ## Impossible priors
# ## Tightening the priors so that we have reasonable values
m5.5 <- quap(
alist(
K ~ dnorm( mu , sigma ) ,
mu <- a + bN*N ,
a ~ dnorm( 0 , 0.2 ) ,
bN ~ dnorm( 0 , 0.5 ) ,
sigma ~ dexp( 1 )
) , data=dcc )
prior <- extract.prior( m5.5 )
xseq <- c(-2,2)
mu <- link( m5.5_draft , post=prior , data=list(N=xseq) )
plot( NULL , xlim=xseq , ylim=xseq )
for ( i in 1:50 ) lines( xseq , mu[i,] , col=col.alpha("black",0.3) )
precis( m5.5 )
xseq <- seq( from=min(dcc$N)-0.15 , to=max(dcc$N)+0.15 , length.out=30 )
mu <- link( m5.5 , data=list(N=xseq) )
mu_mean <- apply(mu,2,mean)
mu_PI <- apply(mu,2,PI)
plot( K ~ N , data=dcc )
lines( xseq , mu_mean , lwd=2 )
shade( mu_PI , xseq )
m5.6 <- quap(
alist(
K ~ dnorm( mu , sigma ) ,
mu <- a + bM*M ,
a ~ dnorm( 0 , 0.2 ) ,
bM ~ dnorm( 0 , 0.5 ) ,
sigma ~ dexp( 1 )
) , data=dcc )
precis(m5.6)
xseq <- seq( from=min(dcc$M)-0.15 , to=max(dcc$M)+0.15 , length.out=30 )
mu <- link( m5.6 , data=list(M=xseq) )
mu_mean <- apply(mu,2,mean)
mu_PI <- apply(mu,2,PI)
plot( K ~ M , data=dcc )
lines( xseq , mu_mean , lwd=2 )
shade( mu_PI , xseq )
m5.7 <- quap(
alist(
K ~ dnorm( mu , sigma ) ,
mu <- a + bN*N + bM*M ,
a ~ dnorm( 0 , 0.2 ) ,
bN ~ dnorm( 0 , 0.5 ) ,
bM ~ dnorm( 0 , 0.5 ) ,
sigma ~ dexp( 1 )
) , data=dcc )
precis(m5.7)
plot( coeftab( m5.5 , m5.6 , m5.7 ) , pars=c("bM","bN") )
# ## Using both M and N have increased their influence on the outcome in a more farther direction
pairs( ~K + M + N ,
dcc )
# ## Counterfactual with N = 0
xseq <- seq( from=min(dcc$M)-0.15 , to=max(dcc$M)+0.15 , length.out=30 )
mu <- link( m5.7 , data=data.frame( M=xseq , N=0 ) )
mu_mean <- apply(mu,2,mean)
mu_PI <- apply(mu,2,PI)
plot( NULL , xlim=range(dcc$M) , ylim=range(dcc$K) )
lines( xseq , mu_mean , lwd=2 )
shade( mu_PI , xseq )
# ## Counterfactual with M = 0
xseq <- seq( from=min(dcc$N)-0.15 , to=max(dcc$N)+0.15 , length.out=30 )
mu <- link( m5.7 , data=data.frame( N=xseq , M=0 ) )
mu_mean <- apply(mu,2,mean)
mu_PI <- apply(mu,2,PI)
plot( NULL , xlim=range(dcc$N) , ylim=range(dcc$K) )
lines( xseq , mu_mean , lwd=2 )
shade( mu_PI , xseq )
# ## So it is clear that M and N when applied together lead to stronger effect
# # Index Variables
# ### Using dummy variables for categorical data doesnot help with the prior so we try to use Index variables
data(Howell1)
d <- Howell1
str(d)
mu_female <- rnorm(1e4,178,20)
mu_male <- rnorm(1e4,178,20) + rnorm(1e4,0,10)
precis( data.frame( mu_female , mu_male ) )
# ### Prior predictive simulation clearly shows that male has more deviation in its values even though there is no prior reasoning to that
#
# \begin{align}
# h_i = Normal(\mu_i,\sigma)\\
# \mu_i = \alpha_i + \beta_m*m_i
# \end{align}
#
# since $\alpha_i$ represents the avg female height when $m_i$ = 0 and if it is 1 there are 2 parameters that determine male height causing more uncertainity even when there is no prior evidence
d$sex <- ifelse( d$male==1 , 2 , 1 )
str( d$sex )
# +
m5.8 <- quap(
alist(
height ~ dnorm( mu , sigma ) ,
mu <- a[sex] ,
a[sex] ~ dnorm( 178 , 20 ) ,
sigma ~ dunif( 0 , 50 )
) , data=d )
precis( m5.8 , depth=2 )
# -
post <- extract.samples(m5.8)
post$diff_fm <- post$a[,1] - post$a[,2]
precis( post , depth=2 )
# ### So this variable diff_fm is called the Contrast
data(milk)
d <- milk
unique(d$clade)
d$clade_id <- as.integer( d$clade )
d$clade_id
d$K <- scale( d$kcal.per.g )
m5.9 <- quap(
alist(
K ~ dnorm( mu , sigma ),
mu <- a[clade_id],
a[clade_id] ~ dnorm( 0 , 0.5 ),
sigma ~ dexp( 1 )
) , data=d )
labels <- paste( "a[" , 1:4 , "]:" , levels(d$clade) , sep="" )
plot( precis( m5.9 , depth=2 , pars="a" ) , labels=labels ,
xlab="expected kcal (std)" )
# ### The above example shows how this idea of Index Variables scales up
# And when you want to find the difference u need to simulate from the posterior distribution
|
The Many Variables & The Spurious Waffles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # word2vec IMDB data
#
# Training word2vec embeddings on the IMDB database and experimenting.
#
# Referência: Tutorial Kagggle ["Bag of Words meets Bags of Popcorn"](https://www.kaggle.com/c/word2vec-nlp-tutorial#part-2-word-vectors)
from bs4 import BeautifulSoup
import re
from nltk.corpus import stopwords
import nltk.data
import pandas as pd
import gensim
train = pd.read_csv( "labeledTrainData.tsv", header=0,
delimiter="\t", quoting=3 )
test = pd.read_csv( "testData.tsv", header=0, delimiter="\t", quoting=3 )
unlabeled_train = pd.read_csv( "unlabeledTrainData.tsv", header=0,
delimiter="\t", quoting=3 )
train
def review_to_wordlist( review, remove_stopwords=False ):
# Function to convert a document to a sequence of words,
# optionally removing stop words. Returns a list of words.
#
# 1. Remove HTML
review_text = BeautifulSoup(review).get_text()
#
# 2. Remove non-letters
review_text = re.sub("[^a-zA-Z]"," ", review_text)
#
# 3. Convert words to lower case and split them
words = review_text.lower().split()
#
# 4. Optionally remove stop words (false by default)
if remove_stopwords:
stops = set(stopwords.words("english"))
words = [w for w in words if not w in stops]
#
# 5. Return a list of words
return(words)
# +
# Load the punkt tokenizer
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
# Define a function to split a review into parsed sentences
def review_to_sentences( review, tokenizer, remove_stopwords=False ):
# Function to split a review into parsed sentences. Returns a
# list of sentences, where each sentence is a list of words
#
# 1. Use the NLTK tokenizer to split the paragraph into sentences
raw_sentences = tokenizer.tokenize(review.decode('utf-8').strip())
#
# 2. Loop over each sentence
sentences = []
for raw_sentence in raw_sentences:
# If a sentence is empty, skip it
if len(raw_sentence) > 0:
# Otherwise, call review_to_wordlist to get a list of words
sentences.append( review_to_wordlist( raw_sentence, \
remove_stopwords ))
#
# Return the list of sentences (each sentence is a list of words,
# so this returns a list of lists
return sentences
# +
sentences = [] # Initialize an empty list of sentences
print("Parsing sentences from training set")
for review in train["review"]:
sentences += review_to_sentences(review, tokenizer)
print("Parsing sentences from unlabeled set")
for review in unlabeled_train["review"]:
sentences += review_to_sentences(review, tokenizer)
# -
y = train["sentiment"]
X = []
for review in train["review"]:
sentences = review_to_sentences(review, tokenizer)
words = []
for sentence in sentences:
words += sentence
X.append(sentence)
print len(train["sentiment"]), len(train["review"]), len(X), X[0]
# print review_to_sentences(train[0], tokenizer)
model = gensim.models.Word2Vec(sentences, min_count=1)
def vectorize(sentence):
return [model[word] for word in sentence]
print(model.wv.most_similar(positive=['he', 'her'], negative=['she']))
acc = model.accuracy('questions-words.txt')
[(d.keys()[1], d[d.keys()[1]]) for d in acc]
for i in range(0, len(acc)):
print(acc[i][acc[i].keys()[1]], len(acc[i]['correct']), len(acc[i]['incorrect']))#, len(acc[i]['correct']/len(acc[i]['incorrect']))
model.wv.syn0.shape
# ## Keras model
# +
from keras.preprocessing import sequence
from keras.optimizers import SGD, RMSprop, Adagrad
from keras.utils import np_utils
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation
from keras.layers.embeddings import Embedding
from keras.layers.recurrent import LSTM, GRU
from keras.datasets import imdb
from keras import backend as K
from theano import function
# +
max_features = 20000
max_len = 500
# weights = model.wv.syn0
weights = np.vstack([np.zeros(100), model.wv.syn0])
lstm_model = Sequential()
lstm_model.add(Embedding(input_dim=weights.shape[0], output_dim=weights.shape[1], weights=[weights]))
# lstm_model.add(Embedding(max_features, 128, input_length = max_len))
lstm_model.add(LSTM(100))
lstm_model.add(Dropout(0.5))
lstm_model.add(Dense(1))
lstm_model.add(Activation('sigmoid'))
print(lstm_model.summary())
# -
max_features = 5000
(X_train, y_train), (X_test, y_test) = imdb.load_data(nb_words = max_features)
lstm_model.compile(loss='binary_crossentropy',
optimizer = 'adam',
metrics=["accuracy"])
print("Train..")
batch_size = 30
score = lstm_model.fit(X_train, y_train, batch_size = batch_size,
nb_epoch = 4, validation_data = (X_test, y_test))
|
models/word2vec-imdb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../myimageclassifier')
import engine as eng
import urllib.request
from PIL import Image
import numpy as np
import joblib
url = "https://raw.githubusercontent.com/wesleybeckner/myImageClassifier/main/myimageclassifier/data/test/drone_10T094634322772.bmp"
path = '../myimageclassifier/data/models/image_classifier_pipe.pkl'
urllib.request.urlretrieve(url, 'image.bmp')
im = Image.open("image.bmp")
img = np.array(im)
loaded_model = joblib.load(open(path, 'rb'))
loaded_model.predict(np.reshape(img,(1,240,320,3)))
#
|
examples/example_api.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import plotly.express as px
import numpy
import matplotlib.pyplot as plt
import seaborn as sns
data = pd.read_csv('Titanic Data.csv')
data.head()
fig = px.histogram(data, x='Age', nbins=26, marginal='box')
fig.show()
fig = px.histogram(data, x='Age',y="Sex", nbins=26, marginal='box')
fig.show()
correlations = data.corr()
correlations
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(correlations, vmin=-1, vmax=1)
fig.colorbar(cax)
ticks = numpy.arange(0,9,1)
plt.show()
pd.pivot_table(data, index='Survived', columns='Pclass', values='Fare')
table = pd.pivot_table(data,index=['Sex','Pclass'])
table
table.plot(kind='box');
pivot=pd.pivot_table(data, index='Survived', columns='Pclass', values='Fare', aggfunc='count')
pivot
boxplot = pivot.boxplot(column=Pclass)
pivot.boxplot(by ='Survived', column =['], grid = False)
ax = data.plot.Scatter(figsize=(16,4), title='Fare Average by Class and Sex')
ax.set_ylabel('Average Fare')
# +
def scatter_plot_class(pclass):
g = sns.FacetGrid(data[data['Pclass'] == pclass],
col='Sex',
col_order=['male', 'female'],
hue='Survived',
hue_kws=dict(marker=['v', '^']),
height=6)
g = (g.map(plt.scatter, 'Age', 'Fare', edgecolor='w', alpha=0.7, s=80).add_legend())
plt.subplots_adjust(top=0.9)
g.fig.suptitle('CLASS {}'.format(pclass))
# plotted separately because the fare scale for the first class makes it difficult to visualize second and third class charts
scatter_plot_class(1)
scatter_plot_class(2)
scatter_plot_class(3)
# -
plt.scatter(x=data.Age,
y=data.Fare
)
plt.xlabel("Age",
fontweight ='bold',
size=14)
plt.ylabel("Fare",
fontweight ='bold',
size=14)
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 815, "status": "ok", "timestamp": 1544418391131, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="iJwLMwEBOXd-" outputId="2ac02fac-77f0-4984-edec-441d364cd143"
# !pip install gensim
# + colab={} colab_type="code" id="H9AV9hKQdo5R"
import os
import re
import json
import pandas as pd
import numpy as np
from keras.models import Sequential
from keras.layers import LSTM, Dense, Embedding, Bidirectional, Dropout
from keras.layers import SpatialDropout1D, Conv1D, MaxPooling1D
from gensim.models import Word2Vec
from keras.utils import np_utils
from keras.preprocessing import sequence
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.model_selection import train_test_split
from keras.regularizers import l2
import seaborn as sns
from gensim.models import KeyedVectors
from sklearn.feature_extraction.text import TfidfVectorizer
from keras.models import model_from_json
pd.options.display.max_rows = 10
pd.set_option('display.max_columns', None)
from google.colab import files
import pickle
from keras.models import Model
from keras.layers import Input, LSTM, Dense, Bidirectional, Concatenate
from keras.utils import to_categorical
from keras.models import load_model
# + colab={"base_uri": "https://localhost:8080/", "height": 120} colab_type="code" executionInfo={"elapsed": 33040, "status": "ok", "timestamp": 1544418428960, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="ZNbsIlbFN5xn" outputId="8e762ab6-b76a-4bf1-eea1-b1328aac6c47"
from google.colab import drive
drive.mount('/content/gdrive')
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 582, "status": "ok", "timestamp": 1544418439031, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="VPa-6NDckouB" outputId="07e51cdd-d3fb-4de4-a5ac-8bdaf226b6fe"
try:
device_name = os.environ['COLAB_TPU_ADDR']
TPU_ADDRESS = 'grpc://' + device_name
print('Found TPU at: {}'.format(TPU_ADDRESS))
except KeyError:
print('TPU not found')
# + colab={} colab_type="code" id="u7BXxoRhN6Y7"
w2v_path = os.path.join("/content/gdrive/My Drive/", "Colab Notebooks/ruwikiruscorpora-nobigrams_upos_skipgram_300_5_2018.vec.gz")
hh_data_path = os.path.join("/content/gdrive/My Drive/", "Colab Notebooks/hh_dataset.csv")
jooble_data_path = os.path.join("/content/gdrive/My Drive/", "Colab Notebooks/by_jobs.csv")
weights_dir = os.path.join("/content/gdrive/My Drive/", "Colab Notebooks/weights")
# + colab={"base_uri": "https://localhost:8080/", "height": 552} colab_type="code" executionInfo={"elapsed": 9059, "status": "ok", "timestamp": 1544418452711, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="PBUAImzpN8eX" outputId="e199e318-5458-4a8f-bed8-0e28e77632e4"
dataset_hh = pd.read_csv(hh_data_path, sep="\t")
dataset_hh = dataset_hh.loc[dataset_hh["lang_text"] == "russian"]
#dataset_hh.dropna(inplace=True)
#dataset_hh = dataset_hh.loc[isinstance(dataset_hh["responsibility"], str)]
#dataset_hh.drop(dataset_hh.index[dataset_hh["lang_text"] == "english"], inplace=True)
dataset_hh.reset_index(drop=True, inplace=True)
print(dataset_hh.info())
#dataset_hh.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 552} colab_type="code" executionInfo={"elapsed": 627, "status": "ok", "timestamp": 1544439105005, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="GuGS9XMbk6Z4" outputId="710550ef-ff1c-41f9-dcd3-61baf69955c9"
dataset_hh.drop_duplicates(["text"], inplace=True)
print("Final size of dataset =", len(dataset_hh))
dataset_hh.reset_index(drop=True, inplace=True)
dataset_hh.info()
# + colab={} colab_type="code" id="w-zbfd7Flemd"
#dataset_hh.to_csv(os.path.join(weights_dir, "hh_dataset_all_uniq_text.csv"), sep='\t', header=True, index=None)
# + colab={} colab_type="code" id="w2BQAAwllytC"
#files.download("hh_dataset_all_uniq_text.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 120542, "status": "ok", "timestamp": 1544418600988, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="tts29-15OY9T" outputId="18e8b5c7-15a6-41cb-c1b3-9196a3df6a0a"
# %%time
vectorizer = KeyedVectors.load_word2vec_format(w2v_path, binary=False)
# + [markdown] colab_type="text" id="iW-wqlnMPww6"
# ### Prepare dataset
# + colab={} colab_type="code" id="4hzMgC-ZObl9"
def get_mask(text, req):
text = text.split(" ")
req = req.split(" ")
mask = np.zeros(len(text))
i = 0
while i <= (len(text)):
if (req == text[i: i + len(req)]):
for j in range(len(req)):
mask[i + j] = 1
i += 1
break
i += 1
return mask
# + colab={} colab_type="code" id="WBRiJSO_P42Z"
def get_training_sample(model, text):
"""
Gets 1 sample of training data
:param model text:
:return str text:
"""
tagged_list = text.split(" ")
vec_list = []
#converting word2vec
for word in tagged_list:
try:
vec_list.append(model[word])
except:
#print("Word " + word + " isn't in vocab. Embeding as zeros")
vec_list.append(np.zeros(300))
return vec_list
# + colab={"base_uri": "https://localhost:8080/", "height": 67} colab_type="code" executionInfo={"elapsed": 116111, "status": "ok", "timestamp": 1544418602647, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="ADcED57Ii3_a" outputId="51a45b25-f197-4d27-a020-85fe359f65fb"
lens = []
total_len = 0
for i, raw in dataset_hh.iterrows():
doc_len = len(raw["text_lemmas"].split(" "))
total_len += doc_len
lens.append(doc_len)
mean_len = total_len / len( dataset_hh)
print("Mean text len =", mean_len)
print(np.mean(lens))
print(np.std(lens))
# + colab={} colab_type="code" id="OJTf9l9dRZhR"
#text = dataset_hh.loc[0, "text_lemmas_tags"]
#print(text)
#sub_text = dataset_hh.loc[0, "requirement_lemmas_tags"]
#print(sub_text)
# + colab={} colab_type="code" id="OjnQMUH4UmZd"
#index = 0
#text = dataset_hh.loc[index, "text_lemmas"]
#sub_text = dataset_hh.loc[index, "requirement_lemmas"]
#sub_text2 = dataset_hh.loc[index, "responsibility_lemmas"]
#
#y1 = get_mask(text, sub_text)
#y2 = get_mask(text, sub_text2)
#y = np.logical_or(y1, y2).astype(int)
#
#x = get_training_sample(vectorizer, text)
# + colab={} colab_type="code" id="i6u1stoRRBDN"
#print(len(x))
#print(len(y))
#print(y)
#x = sequence.pad_sequences([x], maxlen=100, dtype='float', padding="post", truncating="post")
#y = sequence.pad_sequences([list(y)], maxlen=100, dtype='float', padding="post", truncating="post")
#print()
#print(y)
#print()
#print(len(x[0]))
#print(len(y[0]))
# + colab={} colab_type="code" id="_oFZwjHXjXKU"
#x[0][99]
# + colab={} colab_type="code" id="OoYsBV_dh_jw"
#print(sub_text)
#print(sub_text2)
#print(text)
# + colab={} colab_type="code" id="swdTHYfBhxAT"
def get_one_sample(row, max_len=200):
if not isinstance(row["requirement_norm"], str) or not isinstance(row["responsibility_norm"], str) :
print("Try another document")
return None
text = row["text_lemmas_tags"]
x_sample = get_training_sample(vectorizer, text)
text = row["text_lemmas"]
sub_text = row["requirement_lemmas"]
sub_text2 = row["responsibility_lemmas"]
y1 = get_mask(text, sub_text)
y2 = get_mask(text, sub_text2)
y_sample = np.logical_or(y1, y2).astype(int)
assert(len(x_sample) == len(y_sample))
x_sample = sequence.pad_sequences([x_sample], maxlen=max_len, dtype='float', padding="post", truncating="post")
y_sample = sequence.pad_sequences([list(y_sample)], maxlen=max_len, dtype='float', padding="post", truncating="post")
y_sample = np.array(y_sample)
y_sample = np.reshape(y_sample, (1, -1, 1))
x_sample = np.array(x_sample)
y_new = np.zeros((y_sample.shape[0], y_sample.shape[1], 2))
for j in range(y_sample.shape[1]):
y_new[0, j] = to_categorical(y_sample[0, j, 0], num_classes=2)
y_sample = y_new
#print("x shape =", x_sample.shape, " y shape =", y_sample.shape)
return x_sample, y_sample
# + colab={} colab_type="code" id="aWupQ_T9ldpI"
x, y = get_one_sample(dataset_hh.loc[0], max_len=200)
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" executionInfo={"elapsed": 118475, "status": "ok", "timestamp": 1544418617687, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="cSr4oPdtSD4Y" outputId="adb824e2-5fe4-48ca-a066-4b62aa1ba939"
# %%time
X = []
Y = []
max_len_real = 0
max_len = 200
n_samples = 10_709
# 10_000: 12_000 - for text
for i, row in dataset_hh.loc[: n_samples].iterrows():
if not isinstance(row["requirement_norm"], str) or not isinstance(row["responsibility_norm"], str) :
continue
text = row["text_lemmas_tags"]
x_sample = get_training_sample(vectorizer, text)
text = row["text_lemmas"]
sub_text = row["requirement_lemmas"]
sub_text2 = row["responsibility_lemmas"]
y1 = get_mask(text, sub_text)
y2 = get_mask(text, sub_text2)
y_sample = np.logical_or(y1, y2).astype(int)
Y.append(y_sample)
X.append(x_sample)
if (len(x_sample) != len(y_sample)):
print(len(x_sample))
print(len(y_sample))
print(len(row["text_normalized"].split(" ")))
print(y_sample)
print(i)
assert(len(x_sample) == len(y_sample))
if len(y_sample) > max_len_real:
max_len_real = len(y_sample)
X = sequence.pad_sequences(X, maxlen=max_len, dtype='float', padding="post", truncating="post")
Y = sequence.pad_sequences(Y, maxlen=max_len, dtype='float', padding="post", truncating="post")
Y = np.array(Y)
Y = np.reshape(Y, (Y.shape[0], Y.shape[1], -1))
X = np.array(X)
print("Max len real=", max_len_real)
print("Max len to reshape =", max_len)
# + colab={} colab_type="code" id="7cDHzKDgshna"
#print(Y[index])
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 114206, "status": "ok", "timestamp": 1544418617691, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="ln9xVkHiYcLU" outputId="bc27b65d-e3d6-4418-efd9-8a2164dc5080"
print(X.shape)
print(Y.shape)
# + [markdown] colab_type="text" id="wF1JdoOPeGbh"
# ### seq2seq model
# + colab={"base_uri": "https://localhost:8080/", "height": 67} colab_type="code" executionInfo={"elapsed": 18417, "status": "ok", "timestamp": 1544418845873, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="MqQGyqzumLIF" outputId="f0a99d47-e8be-4c93-8f80-1fcbecfb2195"
# %%time
Y_new = np.zeros((Y.shape[0], Y.shape[1], 2))
for i in range(Y.shape[0]):
for j in range(Y.shape[1]):
Y_new[i, j] = to_categorical(Y[i, j, 0], num_classes=2)
Y = Y_new
print(Y.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 16308, "status": "ok", "timestamp": 1544418847653, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="1fpSLvO0i-4T" outputId="d303a909-1901-44dd-8334-dc1d5cbe810c"
# %%time
encoder_input_data = X
decoder_input_data = Y
decoder_target_data = np.zeros(Y.shape)
for i in range(Y.shape[0]):
for j in range(0, Y.shape[1]-1):
decoder_target_data[i, j] = Y[i, j + 1]
# + colab={} colab_type="code" id="cZquRHwAjhIe"
#for i in range(0, 30):
# print('Y:', np.argmax(Y[index, i]), 'Target', np.argmax(decoder_target_data[index, i]))
# + colab={} colab_type="code" id="DW834k1rfVNI"
max_len = 200
num_encoder_tokens = 300
latent_dim = 400
num_decoder_tokens = 2
max_decoder_seq_length = max_len
batch_size = 64
epochs = 40
# + colab={} colab_type="code" id="KKSoC2ofo16y"
from keras.callbacks import ModelCheckpoint, EarlyStopping
# + colab={} colab_type="code" id="_ZCgcfc2qbIe"
callbacks = [ModelCheckpoint(os.path.join(weights_dir, 'seq2seq_2.{epoch:02d}-{acc:.2f}.hdf5'))]
# + colab={} colab_type="code" id="ncp8J_eMZXSZ"
# Define an input sequence and process it.
encoder_inputs = Input(shape=(None, num_encoder_tokens))
encoder = Bidirectional(LSTM(latent_dim // 2, return_state=True))
#a = encoder(encoder_inputs)
#print(len(a))
#encoder_outputs, state_h, state_c = encoder(encoder_inputs)
encoder_outputs, forward_h, forward_c, backward_h, backward_c = encoder(encoder_inputs)
state_h = Concatenate(axis=1)([forward_h, backward_h])
state_c = Concatenate(axis=1)([forward_c, backward_c])
# We discard `encoder_outputs` and only keep the states.
encoder_states = [state_h, state_c]
# Set up the decoder, using `encoder_states` as initial state.
decoder_inputs = Input(shape=(None, num_decoder_tokens))
# We set up our decoder to return full output sequences,
# and to return internal states as well. We don't use the
# return states in the training model, but we will use them in inference.
#decoder_lstm = LSTM(latent_dim, return_sequences=True, return_state=True)
decoder_lstm = LSTM(latent_dim, return_sequences=True, return_state=True)
decoder_outputs, _, _ = decoder_lstm(decoder_inputs, initial_state=encoder_states)
decoder_dense = Dense(num_decoder_tokens, activation='softmax')
decoder_outputs = decoder_dense(decoder_outputs)
# Define the model that will turn
# `encoder_input_data` & `decoder_input_data` into `decoder_target_data`
model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
# + colab={} colab_type="code" id="v1owkEuch2hT"
#model = load_model(os.path.join(weights_dir, 'seq2seq_2.33-0.99.hdf5'))
# + colab={} colab_type="code" id="qM5qUw-imzgs"
#model.save_weights(os.path.join(weights_dir, 'seq2seq_2.33-0.99_weights.h5'))
# + colab={} colab_type="code" id="cN8_bkbfnGlM"
model.load_weights(os.path.join(weights_dir, 'seq2seq_2.33-0.99_weights.h5'))
# + colab={} colab_type="code" id="IdN2gd8WhnFW"
# Run training
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', loss_weights=[[0.1, 1000_000]], metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/", "height": 435} colab_type="code" executionInfo={"elapsed": 506, "status": "ok", "timestamp": 1544420252770, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="ggeOuyn4Y5JN" outputId="6779f4b8-81b0-4ceb-d86b-bcff25f11f98"
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1442} colab_type="code" executionInfo={"elapsed": 8694908, "status": "ok", "timestamp": 1544429079212, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="EZ0oj78OhpHO" outputId="2ecc1acd-1950-4c75-9af3-cb68c17aee62"
# %%time
history = model.fit([encoder_input_data, decoder_input_data], decoder_target_data,
batch_size=batch_size,
epochs=epochs,
#validation_split=0.1,
callbacks=callbacks,
verbose=1)
# class_weight={0: 1.0, 1: 1000}
# + colab={"base_uri": "https://localhost:8080/", "height": 134} colab_type="code" executionInfo={"elapsed": 507, "status": "ok", "timestamp": 1544420273636, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="zHp-52ETi9u9" outputId="e6896567-33af-4951-8b8f-da7a85e54ae4"
model.layers
# + colab={} colab_type="code" id="NWxOZ9NEjgYs"
#encoder_inputs = model.layers[0]
#encoder_states = model.layers[3: 5]
#decoder_inputs = model.layers[2]
#decoder_lstm = LSTM(latent_dim, return_sequences=True, return_state=True)
# + colab={} colab_type="code" id="GUdRsBRZp18w"
encoder_model = Model(encoder_inputs, encoder_states)
decoder_state_input_h = Input(shape=(latent_dim,))
decoder_state_input_c = Input(shape=(latent_dim,))
decoder_states_inputs = [decoder_state_input_h, decoder_state_input_c]
decoder_outputs, state_h, state_c = decoder_lstm(
decoder_inputs, initial_state=decoder_states_inputs)
decoder_states = [state_h, state_c]
decoder_outputs = decoder_dense(decoder_outputs)
decoder_model = Model(
[decoder_inputs] + decoder_states_inputs,
[decoder_outputs] + decoder_states)
# + colab={} colab_type="code" id="YqbUFhdyeMFT"
def decode_sequence(input_seq):
# Encode the input as state vectors.
states_value = encoder_model.predict(input_seq)
# Generate empty target sequence of length 1.
target_seq = np.zeros((1, 1, num_decoder_tokens))
# Populate the first character of target sequence with the start character.
target_seq[0, 0, 0] = 1.
# Sampling loop for a batch of sequences
# (to simplify, here we assume a batch of size 1).
stop_condition = False
decoded_sentence = []
while not stop_condition:
output_tokens, h, c = decoder_model.predict(
[target_seq] + states_value)
# Sample a token
sampled_token_index = np.argmax(output_tokens[0, -1, :])
"""sampled_char = reverse_target_char_index[sampled_token_index]"""
decoded_sentence.append(sampled_token_index)
# Exit condition: either hit max length
# or find stop character.
if (len(decoded_sentence) == max_decoder_seq_length) or \
(len(decoded_sentence) > max_decoder_seq_length):
stop_condition = True
# Update the target sequence (of length 1).
target_seq = np.zeros((1, 1, num_decoder_tokens))
target_seq[0, 0, sampled_token_index] = 1.
# Update states
states_value = [h, c]
return decoded_sentence
# + [markdown] colab_type="text" id="FiuuXtj3nrtZ"
# ### Save model
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" executionInfo={"elapsed": 8637456, "status": "ok", "timestamp": 1544429079925, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="aJOshA2zpi4Y" outputId="db171045-02ac-4f5e-a3f9-ad1f80502e64"
with open(os.path.join(weights_dir, 'encoder_model.json'), 'w', encoding='utf8') as f:
f.write(encoder_model.to_json())
encoder_model.save_weights(os.path.join(weights_dir,'encoder_model_weights.h5'))
with open(os.path.join(weights_dir,'decoder_model.json'), 'w', encoding='utf8') as f:
f.write(decoder_model.to_json())
decoder_model.save_weights(os.path.join(weights_dir, 'decoder_model_weights.h5'))
# + colab={} colab_type="code" id="UU_MhzMtuJFb"
del X
del Y
# + colab={} colab_type="code" id="CgrPZm7Ia_w2"
files.download("encoder_model.json")
files.download("decoder_model.json")
files.download("encoder_model_weights.h5")
files.download("decoder_model_weights.h5")
# + [markdown] colab_type="text" id="RG_Tzx37cBoO"
# ### Load model from file to test
# + colab={"base_uri": "https://localhost:8080/", "height": 56, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>WUsCn07Cn0pKHNlbGYpOwo=", "headers": [["content-type", "application/javascript"]], "ok": true, "status": 200, "status_text": ""}}} colab_type="code" executionInfo={"elapsed": 104684, "status": "ok", "timestamp": 1544390514744, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="FUglLAfkbEGp" outputId="e502a583-0358-4bb5-9407-4ec5931e35cc"
#files.upload()
# + colab={} colab_type="code" id="6w_juSU-qkJ3"
#def load_model(model_filename, model_weights_filename):
# with open(model_filename, 'r', encoding='utf8') as f:
# model = model_from_json(f.read())
# model.load_weights(model_weights_filename)
# return model
#
#encoder_model = load_model('encoder_model.json', 'encoder_model_weights.h5')
#decoder_model = load_model('decoder_model.json', 'decoder_model_weights.h5')
# + colab={} colab_type="code" id="DEzrv0IAbgiO"
def decode_sequence(input_seq):
# Encode the input as state vectors.
states_value = encoder_model.predict(input_seq)
# Generate empty target sequence of length 1.
target_seq = np.zeros((1, 1, num_decoder_tokens))
# Populate the first character of target sequence with the start character.
target_seq[0, 0, 0] = 1.
# Sampling loop for a batch of sequences
# (to simplify, here we assume a batch of size 1).
stop_condition = False
decoded_sentence = []
while not stop_condition:
output_tokens, h, c = decoder_model.predict(
[target_seq] + states_value)
# Sample a token
sampled_token_index = np.argmax(output_tokens[0, -1, :])
"""sampled_char = reverse_target_char_index[sampled_token_index]"""
decoded_sentence.append(sampled_token_index)
# Exit condition: either hit max length
# or find stop character.
if (len(decoded_sentence) == max_decoder_seq_length) or \
(len(decoded_sentence) > max_decoder_seq_length):
stop_condition = True
# Update the target sequence (of length 1).
target_seq = np.zeros((1, 1, num_decoder_tokens))
target_seq[0, 0, sampled_token_index] = 1.
# Update states
states_value = [h, c]
return decoded_sentence
# + colab={} colab_type="code" id="0n1KS7xndJcZ"
index = 12001
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 872, "status": "ok", "timestamp": 1544440722628, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="0GGvfTBRmqql" outputId="9375aa5e-70f7-45b2-9195-4c8747bc1881"
# %%time
x, y = get_one_sample(dataset_hh.loc[index])
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 1367, "status": "ok", "timestamp": 1544440723512, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="lYc539OZcUsV" outputId="60725673-7493-4690-e611-5b2fddff4332"
# %%time
predict = decode_sequence(x)
# + colab={"base_uri": "https://localhost:8080/", "height": 120} colab_type="code" executionInfo={"elapsed": 895, "status": "ok", "timestamp": 1544440723514, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="OJgEG4VKciIV" outputId="3203fdba-d47a-4244-c7d5-55494cf9bd05"
print(dataset_hh.loc[index, "title"])
print()
print(dataset_hh.loc[index, "requirement_lemmas"])
print(dataset_hh.loc[index, "responsibility_lemmas"])
print(dataset_hh.loc[index, "text_lemmas"])
# + colab={"base_uri": "https://localhost:8080/", "height": 1689} colab_type="code" executionInfo={"elapsed": 786, "status": "ok", "timestamp": 1544440725290, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="O-NcSbMacXwn" outputId="1128ca27-1868-44d4-ae7d-f65ccef65d8d"
words = dataset_hh.loc[index, "text_lemmas"].split(" ")
for i in range(0, 100):
if len(words) > i:
word = words[i]
else:
word = "END"
print(i, 'Expected:', np.argmax(y[0, i]), 'Predicted', predict[i], word)
# + [markdown] colab_type="text" id="fESe3tXxpvHx"
# ### Text on jooble data
# + colab={"base_uri": "https://localhost:8080/", "height": 351} colab_type="code" executionInfo={"elapsed": 1255, "status": "ok", "timestamp": 1544440745693, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="2EobS2jMp6II" outputId="98eb6047-f7f3-4af1-bfe9-24ea3c447e03"
dataset_jooble = pd.read_csv(jooble_data_path, sep="\t")
dataset_jooble = dataset_jooble.loc[dataset_jooble["lang_text"] == "russian"]
#dataset_hh.dropna(inplace=True)
#dataset_hh = dataset_hh.loc[isinstance(dataset_hh["responsibility"], str)]
#dataset_hh.drop(dataset_hh.index[dataset_hh["lang_text"] == "english"], inplace=True)
dataset_jooble.reset_index(drop=True, inplace=True)
print(dataset_jooble.info())
#dataset_jooble.head()
# + colab={} colab_type="code" id="SgTTv962puoB"
maxlen = 200
def get_sample_nonlabeled(row):
text = row["text_lemmas_tags"]
x_sample = get_training_sample(vectorizer, text)
x_sample = sequence.pad_sequences([x_sample], maxlen=max_len, dtype='float', padding="post", truncating="post")
x_sample = np.array(x_sample)
#print("x shape =", x_sample.shape)
return x_sample
# + colab={"base_uri": "https://localhost:8080/", "height": 134} colab_type="code" executionInfo={"elapsed": 80757, "status": "ok", "timestamp": 1544440851732, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="s2c4UKYFtrJD" outputId="38750703-087f-4597-9391-b5ab7eb93899"
# %%time
indexes_labeled = []
n_test = 100
for i, row in dataset_jooble.loc[: n_test].iterrows():
x = get_sample_nonlabeled(dataset_jooble.loc[i])
predict = decode_sequence(x)
if np.sum(predict) > 0:
indexes_labeled.append(i)
if i % 100 == 0:
print(i)
print()
print("Total:")
print(len(indexes_labeled))
# + [markdown] colab_type="text" id="3_kFxa7G1cb_"
# 80 epochs
# Total:
# 15
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 450, "status": "ok", "timestamp": 1544441059174, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="D-2AUnH9ukun" outputId="9c292819-e22c-4d3e-8ad2-7149d6a1d8d7"
print(indexes_labeled)
# + [markdown] colab_type="text" id="UWlQC2b91XZj"
# 80 epoch
# [8, 9, 18, 27, 36, 57, 61, 63, 68, 70, 71, 85, 87, 97, 100]
#
# new(best):
# [8, 9, 18, 27, 36, 41, 57, 61, 65, 68, 69, 85, 97]
# + colab={"base_uri": "https://localhost:8080/", "height": 418} colab_type="code" executionInfo={"elapsed": 1268, "status": "ok", "timestamp": 1544441473516, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="vAFL4yxZq8yi" outputId="dfc7555e-92fe-4c6d-f085-dfe7d5e2c4e2"
# %%time
index = 65
x = get_sample_nonlabeled(dataset_jooble.loc[index])
predict = decode_sequence(x)
print(np.sum(predict))
print(dataset_jooble.loc[index, "title"])
print()
print(dataset_jooble.loc[index, "text"])
# + colab={"base_uri": "https://localhost:8080/", "height": 870} colab_type="code" executionInfo={"elapsed": 490, "status": "ok", "timestamp": 1544441477997, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="HkZZK2DwrdFF" outputId="ba6eb071-06d8-4cbc-bd65-1193f8c5aeee"
print(np.sum(predict))
words = dataset_jooble.loc[index, "text_lemmas"].split(" ")
for i in range(0, 50):
if len(words) > i:
word = words[i]
else:
word = "END"
print(i, 'Predicted', predict[i], " - ", word)
# + [markdown] colab_type="text" id="6FDWq1s2uTaO"
# ### Test
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" executionInfo={"elapsed": 8860661, "status": "ok", "timestamp": 1544429475262, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="y7ZST_wCwwy-" outputId="b2bb9a04-96ad-401a-8a1c-70935fe1b835"
# %%time
X = []
Y = []
max_len_real = 0
max_len = 200
n_samples = 10_709
# 10_000: 12_000 - for text
for i, row in dataset_hh.loc[10_000: 13_000].iterrows():
if not isinstance(row["requirement_norm"], str) or not isinstance(row["responsibility_norm"], str) :
continue
text = row["text_lemmas_tags"]
x_sample = get_training_sample(vectorizer, text)
text = row["text_lemmas"]
sub_text = row["requirement_lemmas"]
sub_text2 = row["responsibility_lemmas"]
y1 = get_mask(text, sub_text)
y2 = get_mask(text, sub_text2)
y_sample = np.logical_or(y1, y2).astype(int)
Y.append(y_sample)
X.append(x_sample)
if (len(x_sample) != len(y_sample)):
print(len(x_sample))
print(len(y_sample))
print(len(row["text_normalized"].split(" ")))
print(y_sample)
print(i)
assert(len(x_sample) == len(y_sample))
if len(y_sample) > max_len_real:
max_len_real = len(y_sample)
X = sequence.pad_sequences(X, maxlen=max_len, dtype='float', padding="post", truncating="post")
Y = sequence.pad_sequences(Y, maxlen=max_len, dtype='float', padding="post", truncating="post")
Y = np.array(Y)
Y = np.reshape(Y, (Y.shape[0], Y.shape[1], -1))
X = np.array(X)
print("Max len real=", max_len_real)
print("Max len to reshape =", max_len)
# + colab={"base_uri": "https://localhost:8080/", "height": 518} colab_type="code" executionInfo={"elapsed": 10951884, "status": "ok", "timestamp": 1544431573563, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="QWMRsUYwuDdO" outputId="14caa6e6-e2c6-4d6c-829f-cd2b73dafa5e"
# %%time
#X.shape[0]
Y_predict = []
for i in range(X.shape[0]):
predict = decode_sequence(X[i].reshape((1, max_len, 300)))
Y_predict.append(predict)
if i % 100 == 0:
print(i)
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 10948313, "status": "ok", "timestamp": 1544431573564, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="tJWfmi9RuiTK" outputId="11a988db-49f3-42a8-e302-e9541aadcb7b"
Y_predict = np.array(Y_predict)
Y_predict.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 10943411, "status": "ok", "timestamp": 1544431573564, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="cqTLqPhAthG-" outputId="ca3d8fb5-871a-4dc0-95ee-3c2c9ddf83ff"
Y.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 10855708, "status": "ok", "timestamp": 1544431573568, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="xnXCxx4m00Nk" outputId="bc9c76e3-0ebf-448c-85e6-9bbefaaddc1c"
np.sum(Y[:, :, 0])
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 10854753, "status": "ok", "timestamp": 1544431573570, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="w0qQ5cn40veQ" outputId="afc74a06-ea75-4eba-8664-6fd6bde43804"
np.sum(Y_predict)
# + colab={} colab_type="code" id="xVwPYQWUp9n3"
from sklearn import metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 10853117, "status": "ok", "timestamp": 1544431574611, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="2zGybev0n1HP" outputId="70846d04-37de-4f13-c5e5-af16bf0af20b"
tp = 0
for i in range(Y.shape[0]):
for j in range(Y.shape[1]):
if Y_predict[i, j] == Y[i, j]:
tp += 1
accuracy = tp / (Y.shape[0] * Y.shape[1]) * 100
print("Accuracy =", accuracy)
# + [markdown] colab_type="text" id="h9RUG7qy3Aqh"
# 60 epochs
# Accuracy = 97.1189866939611
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 10846377, "status": "ok", "timestamp": 1544431574814, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="5pKSglYUs1oL" outputId="3db70e2b-4f38-4214-8dc4-790d6f3e2c1c"
f1_score = metrics.f1_score(Y.reshape(-1), Y_predict.reshape(-1))
print("F1-score =", f1_score)
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" executionInfo={"elapsed": 10837850, "status": "ok", "timestamp": 1544431575086, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="Nv3PcgM-7yXl" outputId="554b1577-01a7-4215-9751-40c8c9c3904f"
f1_score = metrics.f1_score(Y.reshape(-1), Y_predict.reshape(-1), average='weighted')
print("F1-score =", f1_score)
# + [markdown] colab_type="text" id="PSfxowdS0kWN"
# F1-score = 0.8140268578318826
# + colab={"base_uri": "https://localhost:8080/", "height": 184} colab_type="code" executionInfo={"elapsed": 10833372, "status": "ok", "timestamp": 1544431576863, "user": {"displayName": "\u0412\u0438\u043a\u0442\u043e\u0440 \u041e\u0432\u0435\u0440\u043a\u043e", "photoUrl": "https://lh6.googleusercontent.com/-_I9YbAF9z5o/AAAAAAAAAAI/AAAAAAAAABA/NUb59sonytE/s64/photo.jpg", "userId": "04713510988786792129"}, "user_tz": -120} id="_Kt1FBV9ygFo" outputId="5939614d-f88b-42d7-eed0-ff824b7a2448"
print(metrics.classification_report(Y.reshape(-1), Y_predict.reshape(-1)))
print("Confussion matrix: \n", metrics.confusion_matrix(Y.reshape(-1), Y_predict.reshape(-1), labels=[0, 1]))
# + [markdown] colab_type="text" id="EGeNeX4V0nLx"
# 80 epochs
#
# precision recall f1-score support
#
# 0.0 0.98 0.99 0.98 359547
# 1.0 0.84 0.79 0.81 31253
#
# avg / total 0.97 0.97 0.97 390800
#
# Confussion matrix:
#
# [[354900 4647]
#
# [ 6612 24641]]
# + colab={} colab_type="code" id="U-Y9dHf_v7HS"
#count = 0
#for i in range(Y_predict.shape[0]):
# if np.sum(Y_predict[i, :]) != 0:
# print(" I =", i, "____________")
# print(Y_predict[i])
# count += 1
#print("total =", count)
# + colab={} colab_type="code" id="Bmo7DteQv9iN"
files.download("encoder_model.json")
files.download("decoder_model.json")
files.download("encoder_model_weights.h5")
files.download("decoder_model_weights.h5")
# + colab={} colab_type="code" id="iyNar7YXwGRN"
|
data/drafts/seq2seq.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CarND Object Detection Lab
#
# Let's get started!
# +
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
from PIL import ImageDraw
from PIL import ImageColor
import time
from scipy.stats import norm
# %matplotlib inline
plt.style.use('ggplot')
# -
# ## MobileNets
#
# [*MobileNets*](https://arxiv.org/abs/1704.04861), as the name suggests, are neural networks constructed for the purpose of running very efficiently (high FPS, low memory footprint) on mobile and embedded devices. *MobileNets* achieve this with 3 techniques:
#
# 1. Perform a depthwise convolution followed by a 1x1 convolution rather than a standard convolution. The 1x1 convolution is called a pointwise convolution if it's following a depthwise convolution. The combination of a depthwise convolution followed by a pointwise convolution is sometimes called a separable depthwise convolution.
# 2. Use a "width multiplier" - reduces the size of the input/output channels, set to a value between 0 and 1.
# 3. Use a "resolution multiplier" - reduces the size of the original input, set to a value between 0 and 1.
#
# These 3 techniques reduce the size of cummulative parameters and therefore the computation required. Of course, generally models with more paramters achieve a higher accuracy. *MobileNets* are no silver bullet, while they perform very well larger models will outperform them. ** *MobileNets* are designed for mobile devices, NOT cloud GPUs**. The reason we're using them in this lab is automotive hardware is closer to mobile or embedded devices than beefy cloud GPUs.
# ### Convolutions
#
# #### Vanilla Convolution
#
# Before we get into the *MobileNet* convolution block let's take a step back and recall the computational cost of a vanilla convolution. There are $N$ kernels of size $D_k * D_k$. Each of these kernels goes over the entire input which is a $D_f * D_f * M$ sized feature map or tensor (if that makes more sense). The computational cost is:
#
# $$
# D_g * D_g * M * N * D_k * D_k
# $$
#
# Let $D_g * D_g$ be the size of the output feature map. Then a standard convolution takes in a $D_f * D_f * M$ input feature map and returns a $D_g * D_g * N$ feature map as output.
#
# (*Note*: In the MobileNets paper, you may notice the above equation for computational cost uses $D_f$ instead of $D_g$. In the paper, they assume the output and input are the same spatial dimensions due to stride of 1 and padding, so doing so does not make a difference, but this would want $D_g$ for different dimensions of input and output.)
#
# 
#
#
#
# #### Depthwise Convolution
#
# A depthwise convolution acts on each input channel separately with a different kernel. $M$ input channels implies there are $M$ $D_k * D_k$ kernels. Also notice this results in $N$ being set to 1. If this doesn't make sense, think about the shape a kernel would have to be to act upon an individual channel.
#
# Computation cost:
#
# $$
# D_g * D_g * M * D_k * D_k
# $$
#
#
# 
#
#
# #### Pointwise Convolution
#
# A pointwise convolution performs a 1x1 convolution, it's the same as a vanilla convolution except the kernel size is $1 * 1$.
#
# Computation cost:
#
# $$
# D_k * D_k * D_g * D_g * M * N =
# 1 * 1 * D_g * D_g * M * N =
# D_g * D_g * M * N
# $$
#
# 
#
#
#
# Thus the total computation cost is for separable depthwise convolution:
#
# $$
# D_g * D_g * M * D_k * D_k + D_g * D_g * M * N
# $$
#
# which results in $\frac{1}{N} + \frac{1}{D_k^2}$ reduction in computation:
#
# $$
# \frac {D_g * D_g * M * D_k * D_k + D_g * D_g * M * N} {D_g * D_g * M * N * D_k * D_k} =
# \frac {D_k^2 + N} {D_k^2*N} =
# \frac {1}{N} + \frac{1}{D_k^2}
# $$
#
# *MobileNets* use a 3x3 kernel, so assuming a large enough $N$, separable depthwise convnets are ~9x more computationally efficient than vanilla convolutions!
# ### Width Multiplier
#
# The 2nd technique for reducing the computational cost is the "width multiplier" which is a hyperparameter inhabiting the range [0, 1] denoted here as $\alpha$. $\alpha$ reduces the number of input and output channels proportionally:
#
# $$
# D_f * D_f * \alpha M * D_k * D_k + D_f * D_f * \alpha M * \alpha N
# $$
# ### Resolution Multiplier
#
# The 3rd technique for reducing the computational cost is the "resolution multiplier" which is a hyperparameter inhabiting the range [0, 1] denoted here as $\rho$. $\rho$ reduces the size of the input feature map:
#
# $$
# \rho D_f * \rho D_f * M * D_k * D_k + \rho D_f * \rho D_f * M * N
# $$
# Combining the width and resolution multipliers results in a computational cost of:
#
# $$
# \rho D_f * \rho D_f * a M * D_k * D_k + \rho D_f * \rho D_f * a M * a N
# $$
#
# Training *MobileNets* with different values of $\alpha$ and $\rho$ will result in different speed vs. accuracy tradeoffs. The folks at Google have run these experiments, the result are shown in the graphic below:
#
# 
# MACs (M) represents the number of multiplication-add operations in the millions.
# ### Exercise 1 - Implement Separable Depthwise Convolution
#
# In this exercise you'll implement a separable depthwise convolution block and compare the number of parameters to a standard convolution block. For this exercise we'll assume the width and resolution multipliers are set to 1.
#
# Docs:
#
# * [depthwise convolution](https://www.tensorflow.org/api_docs/python/tf/nn/depthwise_conv2d)
# +
def vanilla_conv_block(x, kernel_size, output_channels):
"""
Vanilla Conv -> Batch Norm -> ReLU
"""
x = tf.layers.conv2d(
x, output_channels, kernel_size, (2, 2), padding='SAME')
x = tf.layers.batch_normalization(x)
return tf.nn.relu(x)
# TODO: implement MobileNet conv block
def mobilenet_conv_block(x, kernel_size, output_channels):
"""
Depthwise Conv -> Batch Norm -> ReLU -> Pointwise Conv -> Batch Norm -> ReLU
"""
# assumes BHWC format
input_channel_dim = x.get_shape().as_list()[-1]
W = tf.Variable(tf.truncated_normal((kernel_size, kernel_size, input_channel_dim, 1)))
# depthwise conv
x = tf.nn.depthwise_conv2d(x, W, (1, 2, 2, 1), padding='SAME')
x = tf.layers.batch_normalization(x)
x = tf.nn.relu(x)
# pointwise conv
x = tf.layers.conv2d(x, output_channels, (1, 1), padding='SAME')
x = tf.layers.batch_normalization(x)
return tf.nn.relu(x)
# -
# **[Sample solution](./exercise-solutions/e1.py)**
#
# Let's compare the number of parameters in each block.
# +
# constants but you can change them so I guess they're not so constant :)
INPUT_CHANNELS = 32
OUTPUT_CHANNELS = 512
KERNEL_SIZE = 3
IMG_HEIGHT = 256
IMG_WIDTH = 256
with tf.Session(graph=tf.Graph()) as sess:
# input
x = tf.constant(np.random.randn(1, IMG_HEIGHT, IMG_WIDTH, INPUT_CHANNELS), dtype=tf.float32)
with tf.variable_scope('vanilla'):
vanilla_conv = vanilla_conv_block(x, KERNEL_SIZE, OUTPUT_CHANNELS)
with tf.variable_scope('mobile'):
mobilenet_conv = mobilenet_conv_block(x, KERNEL_SIZE, OUTPUT_CHANNELS)
vanilla_params = [
(v.name, np.prod(v.get_shape().as_list()))
for v in tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, 'vanilla')
]
mobile_params = [
(v.name, np.prod(v.get_shape().as_list()))
for v in tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, 'mobile')
]
print("VANILLA CONV BLOCK")
total_vanilla_params = sum([p[1] for p in vanilla_params])
for p in vanilla_params:
print("Variable {0}: number of params = {1}".format(p[0], p[1]))
print("Total number of params =", total_vanilla_params)
print()
print("MOBILENET CONV BLOCK")
total_mobile_params = sum([p[1] for p in mobile_params])
for p in mobile_params:
print("Variable {0}: number of params = {1}".format(p[0], p[1]))
print("Total number of params =", total_mobile_params)
print()
print("{0:.3f}x parameter reduction".format(total_vanilla_params /
total_mobile_params))
# -
# Your solution should show the majority of the parameters in *MobileNet* block stem from the pointwise convolution.
# ## *MobileNet* SSD
#
# In this section you'll use a pretrained *MobileNet* [SSD](https://arxiv.org/abs/1512.02325) model to perform object detection. You can download the *MobileNet* SSD and other models from the [TensorFlow detection model zoo](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/detection_model_zoo.md) (*note*: we'll provide links to specific models further below). [Paper](https://arxiv.org/abs/1611.10012) describing comparing several object detection models.
#
# Alright, let's get into SSD!
# ### Single Shot Detection (SSD)
#
# Many previous works in object detection involve more than one training phase. For example, the [Faster-RCNN](https://arxiv.org/abs/1506.01497) architecture first trains a Region Proposal Network (RPN) which decides which regions of the image are worth drawing a box around. RPN is then merged with a pretrained model for classification (classifies the regions). The image below is an RPN:
#
# 
# The SSD architecture is a single convolutional network which learns to predict bounding box locations and classify the locations in one pass. Put differently, SSD can be trained end to end while Faster-RCNN cannot. The SSD architecture consists of a base network followed by several convolutional layers:
#
# 
#
# **NOTE:** In this lab the base network is a MobileNet (instead of VGG16.)
#
# #### Detecting Boxes
#
# SSD operates on feature maps to predict bounding box locations. Recall a feature map is of size $D_f * D_f * M$. For each feature map location $k$ bounding boxes are predicted. Each bounding box carries with it the following information:
#
# * 4 corner bounding box **offset** locations $(cx, cy, w, h)$
# * $C$ class probabilities $(c_1, c_2, ..., c_p)$
#
# SSD **does not** predict the shape of the box, rather just where the box is. The $k$ bounding boxes each have a predetermined shape. This is illustrated in the figure below:
#
# 
#
# The shapes are set prior to actual training. For example, In figure (c) in the above picture there are 4 boxes, meaning $k$ = 4.
# ### Exercise 2 - SSD Feature Maps
#
# It would be a good exercise to read the SSD paper prior to a answering the following questions.
#
# ***Q: Why does SSD use several differently sized feature maps to predict detections?***
# A: Differently sized feature maps allow for the network to learn to detect objects at different
# resolutions. This is illustrated in the figure with the 8x8 and 4x4 feature maps. This may remind you
# of skip connections in fully convolutional networks.
#
# **[Sample answer](./exercise-solutions/e2.md)**
# The current approach leaves us with thousands of bounding box candidates, clearly the vast majority of them are nonsensical.
#
# ### Exercise 3 - Filtering Bounding Boxes
#
# ***Q: What are some ways which we can filter nonsensical bounding boxes?***
# A: You may have come up with different answers. The SSD paper does 2 things:
#
# 1. Filters boxes based on IoU metric. For example, if a box has an IoU score
# less than 0.5 on all ground truth boxes it's removed.
#
# 2. *Hard negative mining*. This is a fancy way of saying "search for negatives examples
# the highest confidence". For example, a box that misclassifies a dog as a cat with 80% confidence.
# The authors of the SSD paper limit the positive to hard negative ratio to 3:1 at most. The actual positive to negative ratio is typically much higher and the number of boxes are typically reduced substantially.
#
# **[Sample answer](./exercise-solutions/e3.md)**
# #### Loss
#
# With the final set of matched boxes we can compute the loss:
#
# $$
# L = \frac {1} {N} * ( L_{class} + L_{box})
# $$
#
# where $N$ is the total number of matched boxes, $L_{class}$ is a softmax loss for classification, and $L_{box}$ is a L1 smooth loss representing the error of the matched boxes with the ground truth boxes. L1 smooth loss is a modification of L1 loss which is more robust to outliers. In the event $N$ is 0 the loss is set 0.
#
#
# ### SSD Summary
#
# * Starts from a base model pretrained on ImageNet.
# * The base model is extended by several convolutional layers.
# * Each feature map is used to predict bounding boxes. Diversity in feature map size allows object detection at different resolutions.
# * Boxes are filtered by IoU metrics and hard negative mining.
# * Loss is a combination of classification (softmax) and dectection (smooth L1)
# * Model can be trained end to end.
# ## Object Detection Inference
#
# In this part of the lab you'll detect objects using pretrained object detection models. You can download the latest pretrained models from the [model zoo](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/detection_model_zoo.md), although do note that you may need a newer version of TensorFlow (such as v1.8) in order to use the newest models.
#
# We are providing the download links for the below noted files to ensure compatibility between the included environment file and the models.
#
# [SSD_Mobilenet 11.6.17 version](http://download.tensorflow.org/models/object_detection/ssd_mobilenet_v1_coco_11_06_2017.tar.gz)
#
# [RFCN_ResNet101 11.6.17 version](http://download.tensorflow.org/models/object_detection/rfcn_resnet101_coco_11_06_2017.tar.gz)
#
# [Faster_RCNN_Inception_ResNet 11.6.17 version](http://download.tensorflow.org/models/object_detection/faster_rcnn_inception_resnet_v2_atrous_coco_11_06_2017.tar.gz)
#
# Make sure to extract these files prior to continuing!
# Frozen inference graph files. NOTE: change the path to where you saved the models.
SSD_GRAPH_FILE = 'ssd_mobilenet_v1_coco_11_06_2017/frozen_inference_graph.pb'
RFCN_GRAPH_FILE = 'rfcn_resnet101_coco_11_06_2017/frozen_inference_graph.pb'
FASTER_RCNN_GRAPH_FILE = 'faster_rcnn_inception_resnet_v2_atrous_coco_11_06_2017/frozen_inference_graph.pb'
# Below are utility functions. The main purpose of these is to draw the bounding boxes back onto the original image.
# +
# Colors (one for each class)
cmap = ImageColor.colormap
print("Number of colors =", len(cmap))
COLOR_LIST = sorted([c for c in cmap.keys()])
#
# Utility funcs
#
def filter_boxes(min_score, boxes, scores, classes):
"""Return boxes with a confidence >= `min_score`"""
n = len(classes)
idxs = []
for i in range(n):
if scores[i] >= min_score:
idxs.append(i)
filtered_boxes = boxes[idxs, ...]
filtered_scores = scores[idxs, ...]
filtered_classes = classes[idxs, ...]
return filtered_boxes, filtered_scores, filtered_classes
def to_image_coords(boxes, height, width):
"""
The original box coordinate output is normalized, i.e [0, 1].
This converts it back to the original coordinate based on the image
size.
"""
box_coords = np.zeros_like(boxes)
box_coords[:, 0] = boxes[:, 0] * height
box_coords[:, 1] = boxes[:, 1] * width
box_coords[:, 2] = boxes[:, 2] * height
box_coords[:, 3] = boxes[:, 3] * width
return box_coords
def draw_boxes(image, boxes, classes, thickness=4):
"""Draw bounding boxes on the image"""
draw = ImageDraw.Draw(image)
for i in range(len(boxes)):
bot, left, top, right = boxes[i, ...]
class_id = int(classes[i])
color = COLOR_LIST[class_id]
draw.line([(left, top), (left, bot), (right, bot), (right, top), (left, top)], width=thickness, fill=color)
def load_graph(graph_file):
"""Loads a frozen inference graph"""
graph = tf.Graph()
with graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(graph_file, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
return graph
# -
# Below we load the graph and extract the relevant tensors using [`get_tensor_by_name`](https://www.tensorflow.org/api_docs/python/tf/Graph#get_tensor_by_name). These tensors reflect the input and outputs of the graph, or least the ones we care about for detecting objects.
# +
detection_graph = load_graph(SSD_GRAPH_FILE)
# detection_graph = load_graph(RFCN_GRAPH_FILE)
# detection_graph = load_graph(FASTER_RCNN_GRAPH_FILE)
# The input placeholder for the image.
# `get_tensor_by_name` returns the Tensor with the associated name in the Graph.
image_tensor = detection_graph.get_tensor_by_name('image_tensor:0')
# Each box represents a part of the image where a particular object was detected.
detection_boxes = detection_graph.get_tensor_by_name('detection_boxes:0')
# Each score represent how level of confidence for each of the objects.
# Score is shown on the result image, together with the class label.
detection_scores = detection_graph.get_tensor_by_name('detection_scores:0')
# The classification of the object (integer id).
detection_classes = detection_graph.get_tensor_by_name('detection_classes:0')
# -
# Run detection and classification on a sample image.
# +
# Load a sample image.
image = Image.open('./assets/sample1.jpg')
image_np = np.expand_dims(np.asarray(image, dtype=np.uint8), 0)
#config = tf.ConfigProto()
#config.gpu_options.allow_growth = True
gpu_options = tf.GPUOptions(allow_growth=True)
session_config = tf.ConfigProto(allow_soft_placement=True, log_device_placement=False, gpu_options=gpu_options)
with tf.Session(config=session_config, graph=detection_graph) as sess:
# Actual detection.
(boxes, scores, classes) = sess.run([detection_boxes, detection_scores, detection_classes],
feed_dict={image_tensor: image_np})
# Remove unnecessary dimensions
boxes = np.squeeze(boxes)
scores = np.squeeze(scores)
classes = np.squeeze(classes)
confidence_cutoff = 0.8
# Filter boxes with a confidence score less than `confidence_cutoff`
boxes, scores, classes = filter_boxes(confidence_cutoff, boxes, scores, classes)
# The current box coordinates are normalized to a range between 0 and 1.
# This converts the coordinates actual location on the image.
width, height = image.size
box_coords = to_image_coords(boxes, height, width)
# Each class with be represented by a differently colored box
draw_boxes(image, box_coords, classes)
plt.figure(figsize=(12, 8))
plt.imshow(image)
# -
# ## Timing Detection
#
# The model zoo comes with a variety of models, each its benefits and costs. Below you'll time some of these models. The general tradeoff being sacrificing model accuracy for seconds per frame (SPF).
def time_detection(sess, img_height, img_width, runs=10):
image_tensor = sess.graph.get_tensor_by_name('image_tensor:0')
detection_boxes = sess.graph.get_tensor_by_name('detection_boxes:0')
detection_scores = sess.graph.get_tensor_by_name('detection_scores:0')
detection_classes = sess.graph.get_tensor_by_name('detection_classes:0')
# warmup
gen_image = np.uint8(np.random.randn(1, img_height, img_width, 3))
sess.run([detection_boxes, detection_scores, detection_classes], feed_dict={image_tensor: gen_image})
times = np.zeros(runs)
for i in range(runs):
t0 = time.time()
sess.run([detection_boxes, detection_scores, detection_classes], feed_dict={image_tensor: image_np})
t1 = time.time()
times[i] = (t1 - t0) * 1000
return times
with tf.Session(graph=detection_graph) as sess:
times = time_detection(sess, 600, 1000, runs=10)
# +
# Create a figure instance
fig = plt.figure(1, figsize=(9, 6))
# Create an axes instance
ax = fig.add_subplot(111)
plt.title("Object Detection Timings")
plt.ylabel("Time (ms)")
# Create the boxplot
plt.style.use('fivethirtyeight')
bp = ax.boxplot(times)
# -
# ### Exercise 4 - Model Tradeoffs
#
# Download a few models from the [model zoo](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/detection_model_zoo.md) and compare the timings.
graphlist = []
graphlist.append(load_graph(SSD_GRAPH_FILE))
graphlist.append(load_graph(RFCN_GRAPH_FILE))
graphlist.append(load_graph(FASTER_RCNN_GRAPH_FILE))
timeslist = []
for detection_graph in graphlist:
gpu_options = tf.GPUOptions(allow_growth=True)
session_config = tf.ConfigProto(allow_soft_placement=True, log_device_placement=False, gpu_options=gpu_options)
with tf.Session(config=session_config, graph=detection_graph) as sess:
timeslist.append(time_detection(sess, 600, 1000, runs=10))
def set_box_color(bp, color):
plt.setp(bp['boxes'], color=color)
plt.setp(bp['whiskers'], color=color)
plt.setp(bp['caps'], color=color)
plt.setp(bp['medians'], color=color)
# +
# Create a figure instance
fig = plt.figure(1, figsize=(9, 6))
# Create an axes instance
ax = fig.add_subplot(111)
plt.title("Object Detection Timings")
plt.ylabel("Time (ms)")
# Create the boxplot
plt.style.use('fivethirtyeight')
bp1 = ax.boxplot(timeslist[0])
bp2 = ax.boxplot(timeslist[1])
bp3 = ax.boxplot(timeslist[2])
set_box_color(bp1, '#D7191C') # colors are from http://colorbrewer2.org/
set_box_color(bp2, '#2C7BB6')
set_box_color(bp3, '#2CA25F')
# draw temporary red and blue lines and use them to create a legend
plt.plot([], c='#D7191C', label='SSD')
plt.plot([], c='#2C7BB6', label='RFCN')
plt.plot([], c='#2CA25F', label='RCNN')
plt.legend()
# -
# ## Detection on a Video
#
# Finally run your pipeline on [this short video](https://s3-us-west-1.amazonaws.com/udacity-selfdrivingcar/advanced_deep_learning/driving.mp4).
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
HTML("""
<video width="960" height="600" controls>
<source src="{0}" type="video/mp4">
</video>
""".format('driving.mp4'))
# ### Exercise 5 - Object Detection on a Video
#
# Run an object detection pipeline on the above clip.
clip = VideoFileClip('driving.mp4')
# TODO: Complete this function.
# The input is an NumPy array.
# The output should also be a NumPy array.
def pipeline(img):
draw_img = Image.fromarray(img)
boxes, scores, classes = sess.run([detection_boxes, detection_scores, detection_classes], feed_dict={image_tensor: np.expand_dims(img, 0)})
# Remove unnecessary dimensions
boxes = np.squeeze(boxes)
scores = np.squeeze(scores)
classes = np.squeeze(classes)
confidence_cutoff = 0.8
# Filter boxes with a confidence score less than `confidence_cutoff`
boxes, scores, classes = filter_boxes(confidence_cutoff, boxes, scores, classes)
# The current box coordinates are normalized to a range between 0 and 1.
# This converts the coordinates actual location on the image.
width, height = draw_img.size
box_coords = to_image_coords(boxes, height, width)
# Each class with be represented by a differently colored box
draw_boxes(draw_img, box_coords, classes)
return np.array(draw_img)
# **[Sample solution](./exercise-solutions/e5.py)**
# +
gpu_options = tf.GPUOptions(allow_growth=True)
session_config = tf.ConfigProto(allow_soft_placement=True, log_device_placement=False, gpu_options=gpu_options)
with tf.Session(config=session_config, graph=detection_graph) as sess:
image_tensor = sess.graph.get_tensor_by_name('image_tensor:0')
detection_boxes = sess.graph.get_tensor_by_name('detection_boxes:0')
detection_scores = sess.graph.get_tensor_by_name('detection_scores:0')
detection_classes = sess.graph.get_tensor_by_name('detection_classes:0')
new_clip = clip.fl_image(pipeline)
# write to file
new_clip.write_videofile('result.mp4')
# -
HTML("""
<video width="960" height="600" controls>
<source src="{0}" type="video/mp4">
</video>
""".format('result.mp4'))
# ## Further Exploration
#
# Some ideas to take things further:
#
# * Finetune the model on a new dataset more relevant to autonomous vehicles. Instead of loading the frozen inference graph you'll load the checkpoint.
# * Optimize the model and get the FPS as low as possible.
# * Build your own detector. There are several base model pretrained on ImageNet you can choose from. [Keras](https://keras.io/applications/) is probably the quickest way to get setup in this regard.
#
|
CarND-Object-Detection-Lab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ltoosaint24/DS-Unit-3-Sprint-2-SQL-and-Databases/blob/master/northwind_py.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="BsppG0G9krBo"
# + [markdown] id="wevrWlEaSdCX"
# Part II- Northwid Database
# + id="RggUd-ybMINv"
import sqlite3
import numpy as numy
import pandas as pd
import random
# + id="CcscDG0rSu9d"
xon = sqlite3.connect('/content/northwind_small.sqlite3')
# + id="bTI-X_0yXcbw"
xo = xon.cursor()
# + colab={"base_uri": "https://localhost:8080/"} id="xwpyLqttSv28" outputId="cab67373-0e45-4c17-8345-48ebae944d5b"
DF = pd.read_sql_query('SELECT * FROM OrderDetail GROUP BY UnitPrice ORDER BY UnitPrice DESC LIMIT 10', xon)
print(DF)
expensive_items =DF.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="KfXq7iCeSzTs" outputId="ee919f2b-9352-42e4-aacb-9006958294f1"
emp =pd.read_sql_query('SELECT Avg(HireDate-Birthdate) FROM Employee',xon)
print(emp.columns)
print(emp)
avg_hire_age = emp
print(avg_hire_age)
# + colab={"base_uri": "https://localhost:8080/"} id="qEimP2sES2Sv" outputId="325d7d1a-f242-4ec3-a488-531905b6f3a6"
er = pd.read_sql_query('Select city, AVG(HireDate -Birthdate) from Employee GROUP BY city',xon)
avg_age_by_city = print(er)
# + colab={"base_uri": "https://localhost:8080/", "height": 317} id="uBDM3-zlfigR" outputId="94fb5570-eeff-46e7-f068-12f6bb0f3992"
pd.read_sql_query('Select EmployeeId, Count(TerritoryId) from EmployeeTerritory GROUP BY EmployeeId ORDER BY Count(TerritoryId) DESC',xon)
# + id="aiFwXbu2llF4"
# + colab={"base_uri": "https://localhost:8080/"} id="cU-T2gMzTQ_f" outputId="f038fc8e-2169-4c5b-cb0c-bc58910e07f3"
dff = pd.read_sql_query('SELECT Supplier.CompanyName, OrderDetail.Id, OrderDetail.OrderId, OrderDetail.ProductId, OrderDetail.UnitPrice FROM OrderDetail, Supplier WHERE OrderDetail.Id = Supplier.Id GROUP BY UnitPrice ORDER BY OrderDetail.UnitPrice DESC LIMIT 10', xon)
print(dff)
expensive_items =dff.head(10)
ten_most_expensive = print(dff)
# + colab={"base_uri": "https://localhost:8080/"} id="ztjo6Al9TT5o" outputId="065ced62-2e0d-439d-8386-d8a8303269cb"
largest_category=pd.read_sql_query('Select CategoryName, MAX(Description) from Category',xon)
print(largest_category)
# + colab={"base_uri": "https://localhost:8080/"} id="g40HKqRETWJa" outputId="fd71437c-7e08-4175-b6c3-2da0d456dda1"
most_territories = pd.read_sql_query('Select EmployeeId, Count(TerritoryId) from EmployeeTerritory GROUP BY EmployeeId ORDER BY Count(TerritoryId) DESC',xon)
print(most_territories)
# + [markdown] id="U8OtAXyHTc1G"
# Part IV - Questions
#
# 1. The Territory table can be considered to be another class table, while it may contain subclass items, such as TerritoryId can be utilized as connecting variables in subclass tables within Employee, such as EmployeeTerritory tables.
#
# 2.MongoDB is appropriate within maintianing large data sets, or Big Data. It may not be suitable for the database that require direct access.
#
# 3.NewSQL is a way of reformating how database are stored. This usually intends to create various environment that supports the storage of databased when dealing with big data sets. They are associated within new SQL database platforms.
|
northwind_py.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Autoencoder
#
# Sticking with the MNIST dataset, let's improve our autoencoder's performance using convolutional layers. Again, loading modules and the data.
# +
# %matplotlib inline
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# -
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets('MNIST_data', validation_size=0)
img = mnist.train.images[2]
plt.imshow(img.reshape((28, 28)), cmap='Greys_r')
# ## Network Architecture
#
# The encoder part of the network will be a typical convolutional pyramid. Each convolutional layer will be followed by a max-pooling layer to reduce the dimensions of the layers. The decoder though might be something new to you. The decoder needs to convert from a narrow representation to a wide reconstructed image. For example, the representation could be a 4x4x8 max-pool layer. This is the output of the encoder, but also the input to the decoder. We want to get a 28x28x1 image out from the decoder so we need to work our way back up from the narrow decoder input layer. A schematic of the network is shown below.
#
# <img src='assets/convolutional_autoencoder.png' width=500px>
#
# Here our final encoder layer has size 4x4x8 = 128. The original images have size 28x28 = 784, so the encoded vector is roughly 16% the size of the original image. These are just suggested sizes for each of the layers. Feel free to change the depths and sizes, but remember our goal here is to find a small representation of the input data.
#
# ### What's going on with the decoder
#
# Okay, so the decoder has these "Upsample" layers that you might not have seen before. First off, I'll discuss a bit what these layers *aren't*. Usually, you'll see **transposed convolution** layers used to increase the width and height of the layers. They work almost exactly the same as convolutional layers, but in reverse. A stride in the input layer results in a larger stride in the transposed convolution layer. For example, if you have a 3x3 kernel, a 3x3 patch in the input layer will be reduced to one unit in a convolutional layer. Comparatively, one unit in the input layer will be expanded to a 3x3 path in a transposed convolution layer. The TensorFlow API provides us with an easy way to create the layers, [`tf.nn.conv2d_transpose`](https://www.tensorflow.org/api_docs/python/tf/nn/conv2d_transpose).
#
# However, transposed convolution layers can lead to artifacts in the final images, such as checkerboard patterns. This is due to overlap in the kernels which can be avoided by setting the stride and kernel size equal. In [this Distill article](http://distill.pub/2016/deconv-checkerboard/) from <NAME>, *et al*, the authors show that these checkerboard artifacts can be avoided by resizing the layers using nearest neighbor or bilinear interpolation (upsampling) followed by a convolutional layer. In TensorFlow, this is easily done with [`tf.image.resize_images`](https://www.tensorflow.org/versions/r1.1/api_docs/python/tf/image/resize_images), followed by a convolution. Be sure to read the Distill article to get a better understanding of deconvolutional layers and why we're using upsampling.
#
# > **Exercise:** Build the network shown above. Remember that a convolutional layer with strides of 1 and 'same' padding won't reduce the height and width. That is, if the input is 28x28 and the convolution layer has stride = 1 and 'same' padding, the convolutional layer will also be 28x28. The max-pool layers are used the reduce the width and height. A stride of 2 will reduce the size by a factor of 2. Odena *et al* claim that nearest neighbor interpolation works best for the upsampling, so make sure to include that as a parameter in `tf.image.resize_images` or use [`tf.image.resize_nearest_neighbor`]( `https://www.tensorflow.org/api_docs/python/tf/image/resize_nearest_neighbor). For convolutional layers, use [`tf.layers.conv2d`](https://www.tensorflow.org/api_docs/python/tf/layers/conv2d). For example, you would write `conv1 = tf.layers.conv2d(inputs, 32, (5,5), padding='same', activation=tf.nn.relu)` for a layer with a depth of 32, a 5x5 kernel, stride of (1,1), padding is 'same', and a ReLU activation. Similarly, for the max-pool layers, use [`tf.layers.max_pooling2d`](https://www.tensorflow.org/api_docs/python/tf/layers/max_pooling2d).
# +
learning_rate = 0.001
# Input and target placeholders
inputs_ = tf.placeholder(tf.float32, shape=[None, 28, 28, 1], name='inputs')
targets_ = tf.placeholder(tf.float32, shape=[None, 28, 28, 1], name='targets')
### Encoder
conv1 = tf.layers.conv2d(inputs_, 16, kernel_size=(5,5), padding='same', activation=tf.nn.relu)
# Now 28x28x16
maxpool1 = tf.layers.max_pooling2d(conv1, pool_size=(2,2), strides=(2,2), padding='same')
# Now 14x14x16
conv2 = tf.layers.conv2d(maxpool1, 8, kernel_size=(5,5), padding='same', activation=tf.nn.relu)
# Now 14x14x8
maxpool2 = tf.layers.max_pooling2d(conv2, pool_size=(2,2), strides=(2,2), padding='same')
# Now 7x7x8
conv3 = tf.layers.conv2d(maxpool2, 8, kernel_size=(5,5), padding='same', activation=tf.nn.relu)
# Now 7x7x8
encoded = tf.layers.max_pooling2d(conv3, pool_size=(2,2), strides=(2,2), padding='same')
# Now 4x4x8
### Decoder
upsample1 = tf.image.resize_nearest_neighbor(encoded, size=(7,7))
# Now 7x7x8
conv4 = tf.layers.conv2d(upsample1, 8, kernel_size=(5,5), padding='same', activation=tf.nn.relu)
# Now 7x7x8
upsample2 = tf.image.resize_nearest_neighbor(conv4, size=(14,14))
# Now 14x14x8
conv5 = tf.layers.conv2d(upsample2, 8, kernel_size=(5,5), padding='same', activation=tf.nn.relu)
# Now 14x14x8
upsample3 = tf.image.resize_nearest_neighbor(conv5, size=(28,28))
# Now 28x28x8
conv6 = tf.layers.conv2d(upsample3, 16, kernel_size=(5,5), padding='same', activation=tf.nn.relu)
# Now 28x28x16
logits = tf.layers.conv2d(conv6, 1, kernel_size=(5,5), padding='same', activation = None)
#Now 28x28x1
# Pass logits through sigmoid to get reconstructed image
decoded = tf.sigmoid(logits)
# Pass logits through sigmoid and calculate the cross-entropy loss
loss = tf.nn.sigmoid_cross_entropy_with_logits(labels = targets_,
logits = logits)
# Get cost and define the optimizer
cost = tf.reduce_mean(loss)
opt = tf.train.AdamOptimizer(learning_rate).minimize(cost)
# -
# ## Training
#
# As before, here we'll train the network. Instead of flattening the images though, we can pass them in as 28x28x1 arrays.
sess = tf.Session()
epochs = 20
batch_size = 200
sess.run(tf.global_variables_initializer())
for e in range(epochs):
for ii in range(mnist.train.num_examples//batch_size):
batch = mnist.train.next_batch(batch_size)
imgs = batch[0].reshape((-1, 28, 28, 1))
batch_cost, _ = sess.run([cost, opt], feed_dict={inputs_: imgs,
targets_: imgs})
print("Epoch: {}/{}...".format(e+1, epochs),
"Training loss: {:.4f}".format(batch_cost))
# +
fig, axes = plt.subplots(nrows=2, ncols=10, sharex=True, sharey=True, figsize=(20,4))
in_imgs = mnist.test.images[:10]
reconstructed = sess.run(decoded, feed_dict={inputs_: in_imgs.reshape((10, 28, 28, 1))})
for images, row in zip([in_imgs, reconstructed], axes):
for img, ax in zip(images, row):
ax.imshow(img.reshape((28, 28)), cmap='Greys_r')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
fig.tight_layout(pad=0.1)
# -
sess.close()
# ## Denoising
#
# As I've mentioned before, autoencoders like the ones you've built so far aren't too useful in practive. However, they can be used to denoise images quite successfully just by training the network on noisy images. We can create the noisy images ourselves by adding Gaussian noise to the training images, then clipping the values to be between 0 and 1. We'll use noisy images as input and the original, clean images as targets. Here's an example of the noisy images I generated and the denoised images.
#
# 
#
#
# Since this is a harder problem for the network, we'll want to use deeper convolutional layers here, more feature maps. I suggest something like 32-32-16 for the depths of the convolutional layers in the encoder, and the same depths going backward through the decoder. Otherwise the architecture is the same as before.
#
# > **Exercise:** Build the network for the denoising autoencoder. It's the same as before, but with deeper layers. I suggest 32-32-16 for the depths, but you can play with these numbers, or add more layers.
# +
learning_rate = 0.001
inputs_ = tf.placeholder(tf.float32, (None, 28, 28, 1), name='inputs')
targets_ = tf.placeholder(tf.float32, (None, 28, 28, 1), name='targets')
### Encoder
conv1 = tf.layers.conv2d(inputs_, 32, kernel_size=(3,3), padding='same', activation=tf.nn.relu)
# Now 28x28x32
maxpool1 = tf.layers.max_pooling2d(conv1, pool_size=(2,2), strides=(2,2), padding='same')
# Now 14x14x32
conv2 = tf.layers.conv2d(maxpool1, 32, kernel_size=(3,3), padding='same', activation=tf.nn.relu)
# Now 14x14x32
maxpool2 = tf.layers.max_pooling2d(conv2, pool_size=(2,2), strides=(2,2), padding='same')
# Now 7x7x32
conv3 = tf.layers.conv2d(maxpool2, 16, kernel_size=(3,3), padding='same', activation=tf.nn.relu)
# Now 7x7x16
encoded = tf.layers.max_pooling2d(conv3, pool_size=(2,2), strides=(2,2), padding='same')
# Now 4x4x16
### Decoder
upsample1 = tf.image.resize_nearest_neighbor(encoded, size=(7,7))
# Now 7x7x16
conv4 = tf.layers.conv2d(upsample1, 16, kernel_size=(3,3), padding='same', activation=tf.nn.relu)
# Now 7x7x16
upsample2 = tf.image.resize_nearest_neighbor(conv4, size=(14,14))
# Now 14x14x16
conv5 = tf.layers.conv2d(upsample2, 32, kernel_size=(3,3), padding='same', activation=tf.nn.relu)
# Now 14x14x32
upsample3 = tf.image.resize_nearest_neighbor(conv5, size=(28,28))
# Now 28x28x32
conv6 = tf.layers.conv2d(upsample3, 32, kernel_size=(3,3), padding='same', activation=tf.nn.relu)
# Now 28x28x32
logits = tf.layers.conv2d(conv6, 1, kernel_size=(3,3), padding='same', activation = None)
#Now 28x28x1
# Pass logits through sigmoid to get reconstructed image
decoded = tf.sigmoid(logits)
# Pass logits through sigmoid and calculate the cross-entropy loss
loss = tf.nn.sigmoid_cross_entropy_with_logits(labels = targets_,
logits = logits)
# Get cost and define the optimizer
cost = tf.reduce_mean(loss)
opt = tf.train.AdamOptimizer(learning_rate).minimize(cost)
# -
sess = tf.Session()
epochs = 100
batch_size = 1000
# Set's how much noise we're adding to the MNIST images
noise_factor = 0.5
sess.run(tf.global_variables_initializer())
for e in range(epochs):
for ii in range(mnist.train.num_examples//batch_size):
batch = mnist.train.next_batch(batch_size)
# Get images from the batch
imgs = batch[0].reshape((-1, 28, 28, 1))
# Add random noise to the input images
noisy_imgs = imgs + noise_factor * np.random.randn(*imgs.shape)
# Clip the images to be between 0 and 1
noisy_imgs = np.clip(noisy_imgs, 0., 1.)
# Noisy images as inputs, original images as targets
batch_cost, _ = sess.run([cost, opt], feed_dict={inputs_: noisy_imgs,
targets_: imgs})
print("Epoch: {}/{}...".format(e+1, epochs),
"Training loss: {:.4f}".format(batch_cost))
# ## Checking out the performance
#
# Here I'm adding noise to the test images and passing them through the autoencoder. It does a suprisingly great job of removing the noise, even though it's sometimes difficult to tell what the original number is.
# +
fig, axes = plt.subplots(nrows=2, ncols=10, sharex=True, sharey=True, figsize=(20,4))
in_imgs = mnist.test.images[:10]
noise_factor = 0.5
noisy_imgs = in_imgs + noise_factor * np.random.randn(*in_imgs.shape)
noisy_imgs = np.clip(noisy_imgs, 0., 1.)
reconstructed = sess.run(decoded, feed_dict={inputs_: noisy_imgs.reshape((10, 28, 28, 1))})
for images, row in zip([noisy_imgs, reconstructed], axes):
for img, ax in zip(images, row):
ax.imshow(img.reshape((28, 28)), cmap='Greys_r')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
fig.tight_layout(pad=0.1)
|
autoencoder/Convolutional_Autoencoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:machine_learning_cookbook]
# language: python
# name: conda-env-machine_learning_cookbook-py
# ---
# ## Chapter 16
# ---
# # Logistic Regression
#
# Despire being called a regression, logistic regression is actually a widely used supervised classification technique.
# Allows us to predict the probability that an observation is of a certain class
#
# ## 16.1 Training a Binary Classifier
# +
from sklearn.linear_model import LogisticRegression
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
iris = datasets.load_iris()
features = iris.data[:100,:]
target = iris.target[:100]
scaler = StandardScaler()
features_standardized = scaler.fit_transform(features)
logistic_regression = LogisticRegression(random_state=0)
model = logistic_regression.fit(features_standardized, target)
new_observation = [[.5, .5, .5, .5]]
print("model.predict: {}".format(model.predict(new_observation)))
print("model.predict_proba: {}".format(model.predict_proba(new_observation)))
# -
# ### Discussion
# Dispire having "regression" in its name, a logistic regression is actually a widely used binary lassifier (i.e. the target vector can only take two values). In a logistic regression, a linear model (e.g. $\beta_0 + \beta_i x$) is included in a logistic (also called sigmoid) function, $\frac{1}{1+e^{-z }}$, such that:
# $$
# P(y_i = 1 | X) = \frac{1}{1+e^{-(\beta_0 + \beta_1x)}}
# $$
# where $P(y_i = 1 | X)$ is the probability of the ith obsevation's target, $y_i$ being class 1, X is the training data, $\beta_0$ and $\beta_1$ are the parameters to be learned, and e is Euler's number. The effect of the logistic function is to constrain the value of the function's output to between 0 and 1 so that i can be interpreted as a probability. If $P(y_i = 1 | X)$ is greater than 0.5, class 1 is predicted; otherwise class 0 is predicted
#
# ## 16.2 Training a Multiclass Classifier
# +
from sklearn.linear_model import LogisticRegression
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
iris = datasets.load_iris()
features = iris.data
target = iris.target
scaler = StandardScaler()
features_standardized = scaler.fit_transform(features)
logistic_regression = LogisticRegression(random_state=0, multi_class="ovr")
#logistic_regression_MNL = LogisticRegression(random_state=0, multi_class="multinomial")
model = logistic_regression.fit(features_standardized, target)
# -
# ### Discussion
# On their own, logistic regressions are only binary classifiers, meaning they cannot handle target vectors with more than two classes. However, two clever extensions to logistic regression do just that. First, in one-vs-rest logistic regression (OVR) a separate model is trained for each class predicted whether an observation is that class or not (thus making it a binary classification problem). It assumes that each observation problem (e.g. class 0 or not) is independent
#
# Alternatively in multinomial logistic regression (MLR) the logistic function we saw in Recipe 15.1 is replaced with a softmax function:
# $$
# P(y_I = k | X) = \frac{e^{\beta_k x_i}}{\sum_{j=1}^{K}{e^{\beta_j x_i}}}
# $$
# where $P(y_i = k | X)$ is the probability of the ith observation's target value, $y_i$, is class k, and K is the total number of classes. One practical advantage of the MLR is that its predicted probabilities using `predict_proba` method are more reliable
#
# We can switch to an MNL by setting `multi_class='multinomial'`
#
# ## 16.3 Reducing Variance Through Regularization
# +
from sklearn.linear_model import LogisticRegressionCV
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
iris = datasets.load_iris()
features = iris.data
target = iris.target
scaler = StandardScaler()
features_standardized = scaler.fit_transform(features)
logistic_regression = LogisticRegressionCV(
penalty='l2', Cs=10, random_state=0, n_jobs=-1)
model = logistic_regression.fit(features_standardized, target)
# -
# ### Discussion
# Regularization is a method of penalizing complex models to reduce their variance. Specifically, a penalty term is added to the loss function we are trying to minimize typically the L1 and L2 penalties
#
# In the L1 penalty:
# $$
# \alpha \sum_{j=1}^{p}{|\hat\beta_j|}
# $$
# where $\hat\beta_j$ is the parameters of the jth of p features being learned and $\alpha$ is a hyperparameter denoting the regularization strength.
#
# With the L2 penalty:
# $$
# \alpha \sum_{j=1}^{p}{\hat\beta_j^2}
# $$
# higher values of $\alpha$ increase the penalty for larger parameter values(i.e. more complex models). scikit-learn follows the common method of using C instead of $\alpha$ where C is the inverse of the regularization strength: $C = \frac{1}{\alpha}$. To reduce variance while using logistic regression, we can treat C as a hyperparameter to be tuned to find thevalue of C that creates the best model. In scikit-learn we can use the `LogisticRegressionCV` class to efficiently tune C.
#
# ## 16.4 Training a Classifier on Very Large Data
# +
from sklearn.linear_model import LogisticRegression
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
iris = datasets.load_iris()
features = iris.data
target = iris.target
scaler = StandardScaler()
features_standardized = scaler.fit_transform(features)
logistic_regression = LogisticRegression(random_state=0, solver="sag") # stochastic average gradient (SAG) solver
model = logistic_regression.fit(features_standardized, target)
# -
# ### Discussion
# scikit-learn's `LogisticRegression` offers a number of techniques for training a logistic regression, called solvers. Most of the time scikit-learn will select the best solver automatically for us or warn us we cannot do something with that solver.
#
# Stochastic averge gradient descent allows us to train a model much faster than other solvers when our data is very large. However, it is also very sensitive to feature scaling, so standardizing our features is particularly important
#
# ### See Also
# * Minimizing Finite Sums with the Stochastic Average Gradient Algorithm, <NAME> (http://www.birs.ca/workshops/2014/14w5003/files/schmidt.pdf)
#
# ## 16.5 Handling Imbalanced Classes
# +
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
iris = datasets.load_iris()
features = iris.data[40:, :]
target = iris.target[40:]
target = np.where((target == 0), 0, 1)
scaler = StandardScaler()
features_standardized = scaler.fit_transform(features)
logistic_regression = LogisticRegression(random_state=0, class_weight="balanced")
model = logistic_regression.fit(features_standardized, target)
# -
# ### Discussion
# `LogisticRegression` comes with a built in method of handling imbalanced classes.
# `class_weight="balanced"` will automatically weigh classes inversely proportional to their frequency:
# $$
# w_j = \frac{n}{kn_j}
# $$
# where $w_j$ is the weight to class j, n is the number of observations, $n_j$ is the number of observations in class j, and k is the total number of classes
|
Chapter 16 - Logistic Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introdução
# Este arquivo se refere aos dados obtidos pela turma de Química Geral I do curso de Processos
# Químicos do IFRJ _campus_ Rio de Janeiro, no 1º semestre de 2019, em uma prática de laboratório cujo objetivo era ensinar
# aos alunos como ler o volume escoado de um dado líquido em buretas e analisar os resultados da turma como um todo, entendendo os motivos que levam a ter leituras distintas.
#
# Ao se repetir várias vezes uma medição são encontrados vários resultados diferentes,
# embora alguns possam coincidir. Ou seja, ao medir várias vezes a mesma grandeza, usando
# o mesmo instrumento e o mesmo observador é possı́vel que as medidas encontradas sejam
# diferentes. A isto se chama *dispersão* das medidas.
#
# No caso em questão, foram realizadas diversas medidas por diferentes pessoas. As medidas
# foram de volume escoado em buretas. O líquido da primeira bureta era água e o da segunda
# era uma solução de permanganato de potássio, que possui uma coloração roxa intensa.
#
# O objetivo é demonstrar que, realizando muitas medidas, boa parte irá se concentrar ao redor
# de uma faixa de valores. Eventualmente, podem surgir valores discrepantes (*outliers*) que
# podem ser devido a erros de leitura. A forma correta de ler uma escala em uma vidraria
# como a bureta é manter a escala na altura dos olhos. Em soluções aquosas, a leitura do volume
# é feita pela parte de baixo do menisco formado pelo líquido. No entanto, quando o líquido
# é muito escuro, como no caso da solução de permanganto, a visualização do menisco fica dificultada.
#
# O erro devido a uma leitura feita acima ou abaixo da linha do menisco do líquido é chamado de
# **erro de paralaxe** e é exemplificado na figura abaixo. Esse erro é uma possível fonte de
# *outliers* por ser um erro grosseiro.
#
# <img src='images/parallax.png' width=200 height=200>
#
# A dispersão nos valores, desconsiderando erros grosseiros, podem ter diversas origens.
# Como as medidas são feitas por pessoas distintas, a própria acuidade visual da pessoa
# pode afetar, além de efeitos de iluminação do local, reflexos no caso de usuários de óculos,
# dentre diversos outros fatores.
#
# Tópicos que podem ser discutidos com base nesse experimento:
#
# - conceitos de *precisão* e *exatidão*
# - conceitos de *repetibilidade* e *reprodutibilidade*
# - tipos de erros (grosseiros, sistemáticos, randômicos)
# - conceitos básicos de estatística (dispersão, média, mediana, algarismos significativos)
# - construção e interpretação de histogramas e boxplots
# - influência da largura de classes (*bins*) na interpretação de um histograma
# # Setup
# Importe as seguintes bibliotecas para que os exemplos e tratamentos dos dados do notebook funcionem.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# A biblioteca a seguir foi criada especialmente para esse trabalho, permitindo uma rápida análise dos dados e geração de gráficos.
import statistics
# # Importação dos dados
df = pd.read_csv('dados_brutos/volumes2019_1.csv')
df.head()
# Breve detalhamento dos dados com o método `describe` do `pandas`.
desc_table = df.describe()
desc_table
# ## Tabela com IQR
#
# Vamos criar uma linha para o intervalo interquartil (*interquartile range* - IQR em inglês) na tabela de descrição de dados. O IQR é o intervalo onde se concentra parte significativa dos dados e sua largura dá uma indicação da dispersão desses valores centrais.
desc_table.loc['IQR'] = desc_table.loc['75%'] - desc_table.loc['25%']
desc_table
# Há um maior *range* de valores para as leituras de permanganato de potássio, talvez *bins* mais largos no histograma sejam melhores para visualizar. Isso será testado a seguir.
# ## Tabelas de frequências
# Vamos criar tabelas de fequências para ter uma ideia melhor de qual a largura de _bins_ mais adequada para cada histograma.
# ### Água
# Vamos obter os valores mínimos e máximos de leitura.
min_water = df['agua / mL'].min()
max_water = df['agua / mL'].max()
# Testando intervalo de *bins* de 0.05
bins_tab = statistics.freq_table(df['agua / mL'], min_water, max_water + 0.03, 0.05)
bins_tab
# Testando intervalo de *bins* de 0.02
bins_tab = statistics.freq_table(df['agua / mL'], min_water, max_water+0.02, 0.02)
bins_tab
# Um intervalo de _bins_ menor permite um maior detalhamento dos dados, mas talvez não gere um histograma que seja adequado para visualização pois há intervalos sem dados. Mais abaixo serão construídos os dois histogramas.
#
# Mais detalhes sobre _bins_ e suas larguras podem ser lidos [aqui](https://en.wikipedia.org/wiki/Histogram#Number_of_bins_and_width).
# ### $KMnO_4$
# Vamos obter os valores mínimos e máximos de leitura.
min_perm = df['KMnO4 / mL'].min()
max_perm = df['KMnO4 / mL'].max()
# Testando intervalo de *bins* de 0.05
bins_tab = statistics.freq_table(df['KMnO4 / mL'], min_perm, max_perm + 0.04, 0.05)
bins_tab
# Testando intervalo de *bins* de 0.02
bins_tab = statistics.freq_table(df['KMnO4 / mL'], min_perm, max_perm+0.02, 0.02)
bins_tab
# O ideal é que a escala, nesse caso, seja de múltiplos de 2. Vamos modificar os valores de início e fim:
bins_tab = statistics.freq_table(df['KMnO4 / mL'], min_perm-0.01, max_perm+0.03, 0.02)
bins_tab
# Repare que há muitos intervalos sem dados até chegar ao valor da última leitura. Provavelmente há um outlier nesses dados, mas precisamos confirmar com ferramentas mais adequadas. Veremos ao plotar o boxplot.
# # Plots
# Para os gráficos, inicialmente criamos uma figura com dois sistemas de eixo, um acima do outro e compartilhando o mesmo eixo horizontal. Assim, passamos esses eixos para a função de histograma e de boxplot e os gráficos irão compartilhar os mesmos valores e serão apresentados um acima do outro.
# ## Água
# Testando plot com intervalo de bins 0.02.
# + code_folding=[]
fig1, (ax2, ax1) = plt.subplots(figsize=(12, 8),
nrows=2,
sharex=True,
facecolor=(1, 1, 1),
gridspec_kw={
"height_ratios": (.15, .85),
'hspace': 0.02
})
statistics.plot_hist(df['agua / mL'], min_water, max_water + 0.03, 0.02, ax=ax1, outlier=True)
statistics.plot_boxplot(df['agua / mL'], ax=ax2)
fig1.subplots_adjust(top=0.90)
fig1.suptitle('Análise estatística - Água', fontsize=20)
plt.show()
# -
# Testando plot com intervalo de bins 0.05.
# +
fig2, (ax2, ax1) = plt.subplots(figsize=(12, 8),
nrows=2,
sharex=True,
facecolor=(1, 1, 1),
gridspec_kw={
"height_ratios": (.15, .85),
'hspace': 0.02
})
statistics.plot_hist(df['agua / mL'], min_water, max_water + 0.03, 0.05, ax=ax1, outlier=True)
statistics.plot_boxplot(df['agua / mL'], ax=ax2)
fig2.subplots_adjust(top=0.90)
fig2.suptitle('Análise estatística - Água', fontsize=20)
plt.show()
# -
# Os histogramas ficam bem distintos, e podemos aproveitar e discutir o efeito da escolha do intervalo de *bins* na interpretação dos dados.
#
# Muito embora um menor intervalo permita analisar mais detalhadamente os dados, ele pode dar uma falsa sensação de descontinuidade. No histograma com menor intervalo, percebemos onde os dados se concentram, o que não é possível de perceber no de maior intervalo de forma independente, apenas com auxílio do boxplot. No entanto, o de maior intervalo apresenta continuidade entre os _bins_ contribuindo para a visualização de que se trata de uma distribuição.
#
# Por fim, o de menor intervalo serve também para destacar melhor os outliers que só são percebidos no histograma de maior intervalo com auxílio do boxplot na parte superior.
# ## $KMnO_4$
# Testando plot com intervalo de bins 0.02.
# +
fig3, (ax2, ax1) = plt.subplots(figsize=(12, 8),
nrows=2,
sharex=True,
facecolor=(1, 1, 1),
gridspec_kw={
"height_ratios": (.15, .85),
'hspace': 0.02
})
statistics.plot_hist(df['KMnO4 / mL'], min_perm - 0.01, max_perm + 0.03, 0.02, ax=ax1, outlier=False)
statistics.plot_boxplot(df['KMnO4 / mL'], ax=ax2)
fig3.subplots_adjust(top=0.90)
fig3.suptitle('Análise estatística - $KMnO_4$', fontsize=20)
plt.show()
# -
# Testando plot com intervalo de bins 0.05.
# +
fig4, (ax2, ax1) = plt.subplots(figsize=(12, 8),
nrows=2,
sharex=True,
facecolor=(1, 1, 1),
gridspec_kw={
"height_ratios": (.15, .85),
'hspace': 0.02
})
statistics.plot_hist(df['KMnO4 / mL'], min_perm, max_perm + 0.01, 0.05, ax=ax1, outlier=False)
statistics.plot_boxplot(df['KMnO4 / mL'], ax=ax2)
fig4.subplots_adjust(top=0.90)
fig4.suptitle('Análise estatística - $KMnO_4$', fontsize=20)
plt.show()
# -
# Repare que o histograma com maior intervalo não mostra o comportamento quase bimodal que se mostra no histograma de menor intervalo.
#
# Além disso, como há um grande IQR nesses dados, a leitura entre 15.20 e 15.22 não é considerada um outlier como se mostra no boxplot. Assim, vemos a importância de utilizarmos as ferramentas corretas. A análise apenas do histograma poderia levar um leitor a achar que tal leitura seria um outlier, mas o boxplot mostra que não.
#
# Por fim, o de menor intervalo serve também para mostrar que indicadores não necessariamente são valores que realmente foram medidos. Repare que a média se encontra numa região onde não foram obtidos valores por medida.
#
# # Informação de versão dos packages utilizados
# %load_ext version_information
# %version_information pandas, numpy, matplotlib
|
volumes_2019_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Notebook to play with WOD data using the wodpy package
#
# WOD data
# * http://www.nodc.noaa.gov/OC5/WOD/datageo.html
#
# wodpy package
# * https://github.com/BillMills/wodpy
#
# Playing with new pandas supprot
# +
from wodpy import wod
from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import netCDF4 as nc
from salishsea_tools import viz_tools, tidetools
import numpy as np
import glob
import os
import datetime
from salishsea_tools.nowcast import analyze
# %matplotlib inline
# -
#make plots pretty
sns.set_style('darkgrid')
# #Load Model Grid
f=nc.Dataset('/data/nsoontie/MEOPAR/NEMO-forcing/grid/bathy_meter_SalishSea2.nc')
bathy=f.variables['Bathymetry'][:]
X=f.variables['nav_lon'][:]
Y=f.variables['nav_lat'][:]
# #Load WOD observations
# Experiment with functions to manipulate the data.
def extract_data(filename, lat_min, lat_max, lon_min, lon_max, start_date, end_date ):
"""Reads a WOD file (filename).
Returns a list of dataframes with profiles in the region defined by lat_min, lat_max, lon_min, lon_max
and between start_date, end_date (datetime objects)
"""
file = open(filename)
#empty list for gatherthing profiles.
list_data=[]
#loop through profiles
profile = wod.WodProfile(file)
while not profile.is_last_profile_in_file(file):
df=profile.df()
profile_date = datetime.datetime(df.year, df.month, df.day)
#isolate
if profile_date >= start_date and profile_date <= end_date:
if df.latitude >= lat_min and df.latitude <= lat_max:
if df.longitude >= lon_min and df.longitude <= lon_max:
list_data.append(df)
profile = wod.WodProfile(file)
#again for last profile
df=profile.df()
profile_date = datetime.datetime(df.year, df.month, df.day)
#isolate
if profile_date >= start_date and profile_date <= end_date:
if df.latitude >= lat_min and df.latitude <= lat_max:
if df.longitude >= lon_min and df.longitude <= lon_max:
list_data.append(df)
return list_data
# +
#define region
lon_min=-123.5; lat_min=49;
lon_max=-123.2; lat_max=49.2;
#define time period
sdt = datetime.datetime(2000,1,1)
edt = datetime.datetime(2015,12,31)
data = extract_data('/ocean/nsoontie/MEOPAR/WOD/CTDS7412', lat_min, lat_max, lon_min, lon_max, sdt, edt)
# -
data[0].latitude, data[0].longitude, data[0].year, data[0].month, data[0].day, data[0].uid
data[0]
# Examine data by plotting
fig,axm = plt.subplots(1,figsize=(5,5))
for df in data:
axm.plot(df.longitude, df.latitude, 'o')
viz_tools.plot_coastline(axm,f,coords='map')
axm.set_xlim([-124.1,-122.5])
axm.set_ylim([48.4,49.5])
|
Nancy/strat/comparisons/WOD Data-pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.9 64-bit (''venv'': venv)'
# language: python
# name: python3
# ---
# +
from sklearn.tree import DecisionTreeClassifier
from sklearn.datasets import make_blobs
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
RANDOM_SEED = 139
train_data, train_labels = make_blobs(n_samples=200, centers=[(0,1),(-3,-3),(4,2)],
n_features=2, random_state=RANDOM_SEED,
cluster_std=(1.2,1.5,1,))
# Let’s write an auxiliary function that will return grid for further visualization.
def get_grid(data):
x_min, x_max = data[:, 0].min() - 1, data[:, 0].max() + 1
y_min, y_max = data[:, 1].min() - 1, data[:, 1].max() + 1
return np.meshgrid(np.arange(x_min, x_max, 0.01), np.arange(y_min, y_max, 0.01))
clf_tree = DecisionTreeClassifier(criterion='entropy', max_depth=3,
random_state=RANDOM_SEED)
# training the tree
clf_tree.fit(train_data, train_labels)
# some code to depict separating surface
xx, yy = get_grid(train_data)
predicted = clf_tree.predict(np.c_[xx.ravel(),
yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, predicted, cmap='coolwarm')
plt.scatter(train_data[:, 0], train_data[:, 1], c=train_labels, s=100,
cmap='coolwarm', edgecolors='black', linewidth=1.5);
|
unit_6/ml_5.2.1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: predict_pv_yield
# language: python
# name: predict_pv_yield
# ---
# +
# Python core
from typing import Optional, Callable, TypedDict, Union
from dataclasses import dataclass
# Scientific python
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import xarray as xr
import numcodecs
# Cloud compute
import gcsfs
# PyTorch
import torch
from torch import nn
import torch.nn.functional as F
from torchvision import transforms
import pytorch_lightning as pl
# -
# ## Consts & config
ZARR = 'solar-pv-nowcasting-data/satellite/EUMETSAT/SEVIRI_RSS/OSGB36/all_zarr_int16'
plt.rcParams['figure.figsize'] = (10, 10)
plt.rcParams['image.interpolation'] = 'none'
torch.cuda.is_available()
# The [Zarr docs](https://zarr.readthedocs.io/en/stable/tutorial.html#configuring-blosc) say we should tell the Blosc compression library to not use threads when reading from a Zarr store using multiple processes:
numcodecs.blosc.use_threads = False
# # Load satellite data
def get_sat_data(filename: str=ZARR) -> xr.DataArray:
"""Lazily opens the Zarr store on Google Cloud Storage (GCS).
Selects the High Resolution Visible (HRV) satellite channel.
"""
gcs = gcsfs.GCSFileSystem()
store = gcsfs.GCSMap(root=filename, gcs=gcs)
dataset = xr.open_zarr(store, consolidated=True)
return dataset['stacked_eumetsat_data'].sel(variable='HRV')
# %%time
sat_data = get_sat_data()
# Caution: Wierdly, plotting `sat_data` at this point causes the code to hang (with no errors messages) when it gets to `enumerate(dataloader)`. The code hangs even if we first do `sat_data.close(); del sat_data`
sat_data
# ## Simple PyTorch Dataset
#
# We have several TB of satellite data. To keep the GPU fed with data during training, we need to read chunks of data quickly from the Zarr store; and we also want to load data asynchronously. That is, while the GPU is training on the current batch, the data loader should simultaneously load the _next_ batch from disk.
#
# PyTorch makes this easy! PyTorch's `DataLoader` spawns multiple worker processes when constructed with `num_workers` set to more than 1. Each worker process receives a copy of the `SatelliteDataset` object.
#
# There is a small challenge: The code hangs when it gets to `enumerate(dataloader)` if we open the `xarray.DataArray` in the main process and copy that opened `DataArray` to the child processes. Our solution is to delay the creation of the `DataArray` until _after_ the worker processes have been created. PyTorch makes this easy by allowing us to pass a `worker_init_fn` to `DataLoader`. `worker_init_fn` is called on each worker process. Our `worker_init_fn` just has one job: to call `SatelliteDataset.per_worker_init()` which, in turn, opens the `DataArray`.
#
# This approach achieves read speeds of 600 MB/s from Google Cloud Storage to a single GCP VM with 12 vCPUs (as measured by `nethogs`).
#
# We use `IterableDataset` instead of `Dataset` so `SatelliteDataset` can pre-load the next example from disk and then block (on the `yield`) waiting for PyTorch to read that data. This allows the worker processes to load the next batch from disk while the main process is training the current batch on the GPU.
#
# We can't pin the memory in each worker process because pinned memory can't be shared across processes. Instead we ask `DataLoader` to pin the collated batch so that pytorch-lightning can asynchronously load the next batch from pinned CPU memory into GPU memory.
#
# The satellite data is stored on disk as `int16`. We keep the data as `int16` until it gets to the
# ### Timestep numbering:
#
# * t<sub>0</sub> is 'now': It's the most recent observation.
# * t<sub>1</sub> is the first forecast.
# +
Array = Union[np.ndarray, xr.DataArray]
IMAGE_ATTR_NAMES = ('historical_sat_images', 'target_sat_images')
class Sample(TypedDict):
"""Simple class for structuring data for the ML model.
Using typing.TypedDict gives us several advantages:
1. Single 'source of truth' for the type and documentation of each example.
2. A static type checker can check the types are correct.
Instead of TypedDict,, we could use typing.NamedTuple,
which would provide runtime checks, but, crucially, Tuples are immutable
so we cannot change the values in the transforms.
"""
# IMAGES
# Shape: batch_size, seq_length, width, height
historical_sat_images: Array
target_sat_images: Array
class BadData(Exception):
pass
@dataclass
class RandomSquareCrop():
size: int = 128 #: Size of the cropped image.
def __call__(self, sample: Sample) -> Sample:
crop_params = None
for attr_name in IMAGE_ATTR_NAMES:
image = sample[attr_name]
# TODO: Random crop!
cropped_image = image[..., :self.size, :self.size]
sample[attr_name] = cropped_image
return sample
class CheckForBadData():
def __call__(self, sample: Sample) -> Sample:
for attr_name in IMAGE_ATTR_NAMES:
image = sample[attr_name]
if np.any(image < 0):
raise BadData(f'{attr_name} has negative values at {image.time}!')
return sample
# TODO: Delete Normalize if we continue normalising on the GPU instead of in the dataset.
class Normalize():
HRV_MEAN = 93.23458
HRV_STD = 115.34247
def __call__(self, sample: Sample) -> Sample:
for attr_name in IMAGE_ATTR_NAMES:
image = sample[attr_name].astype(np.float32)
image -= Normalize.HRV_MEAN
image /= Normalize.HRV_STD
sample[attr_name] = image
return sample
class ToTensor():
def __call__(self, sample: Sample) -> Sample:
for key, value in sample.items():
if isinstance(value, xr.DataArray):
value = value.values
sample[key] = torch.from_numpy(value)
return sample
# +
@dataclass
class SatelliteDataset(torch.utils.data.IterableDataset):
total_dataset_len: int #: The total number of timesteps in the entire dataset.
history_len: int = 1 #: The number of timesteps of 'history' to load.
forecast_len: int = 1 #: The number of timesteps of 'forecast' to load.
transform: Optional[Callable] = None
n_samples_per_epoch_total: int = 1_000_000
def __post_init__(self):
#: Total sequence length of each sample.
self.total_seq_len = self.history_len + self.forecast_len
#: Effective length of entire dataset.
self.effective_dataset_len = self.total_dataset_len - self.total_seq_len
def per_worker_init(self, n_workers: int=1) -> None:
"""Called by worker_init_fn on each copy of SatelliteDataset after the worker process has been spawned."""
self.data_array = get_sat_data()
self.n_samples_per_epoch_per_worker = self.n_samples_per_epoch_total // n_workers
# Each worker must have a different seed for its random number generator.
# Otherwise all the workers will output exactly the same data!
seed = torch.initial_seed()
self.rng = np.random.default_rng(seed=seed)
def __iter__(self):
for _ in range(self.n_samples_per_epoch_per_worker):
# TODO: Ensure images are contiguous in time.
start_idx = self.rng.integers(low=0, high=self.effective_dataset_len, dtype=np.uint32)
end_idx = start_idx + self.total_seq_len
sat_images = self.data_array.isel(time=slice(start_idx, end_idx))
sample = Sample(
historical_sat_images=sat_images[:self.history_len],
target_sat_images=sat_images[self.history_len:])
if self.transform:
try:
sample = self.transform(sample)
except BadData as e:
print(e)
continue
yield sample
def worker_init_fn(worker_id):
"""Configures each dataset worker process.
Just has one job! To call SatelliteDataset.per_worker_init().
"""
# get_worker_info returns information specific to each worker process.
worker_info = torch.utils.data.get_worker_info()
if worker_info is None:
print('worker_info is None!')
else:
dataset_obj = worker_info.dataset # The Dataset copy in this worker process.
dataset_obj.per_worker_init(n_workers=worker_info.num_workers)
torch.manual_seed(42)
dataset = SatelliteDataset(
total_dataset_len=len(sat_data),
transform=transforms.Compose([
RandomSquareCrop(),
#CheckForBadData(),
ToTensor(),
]),
)
# -
dataloader = torch.utils.data.DataLoader(
dataset,
batch_size=8,
num_workers=10, # timings: 4=13.8s; 8=11.6; 10=11.3s; 11=11.5s; 12=12.6s. 10=3it/s
worker_init_fn=worker_init_fn,
pin_memory=True
)
# %%time
for i, batch in enumerate(dataloader):
print(i, batch['historical_sat_images'].shape)
if i > 2:
break
batch['historical_sat_images'].shape
batch['target_sat_images'].shape
batch['historical_sat_images'].dtype
plt.imshow(batch['historical_sat_images'][0, 0])
# # Simple ML model
def normalise_images_in_model(images, device):
SAT_IMAGE_MEAN = torch.tensor(93.23458, dtype=torch.float, device=device)
SAT_IMAGE_STD = torch.tensor(115.34247, dtype=torch.float, device=device)
images = images.float()
images -= SAT_IMAGE_MEAN
images /= SAT_IMAGE_STD
return images
# +
CHANNELS = 32
KERNEL = 3
class LitAutoEncoder(pl.LightningModule):
def __init__(self):
super().__init__()
self.encoder_conv1 = nn.Conv2d(in_channels=1, out_channels=CHANNELS//2, kernel_size=KERNEL)
self.encoder_conv2 = nn.Conv2d(in_channels=CHANNELS//2, out_channels=CHANNELS, kernel_size=KERNEL)
self.encoder_conv3 = nn.Conv2d(in_channels=CHANNELS, out_channels=CHANNELS, kernel_size=KERNEL)
self.encoder_conv4 = nn.Conv2d(in_channels=CHANNELS, out_channels=CHANNELS, kernel_size=KERNEL)
self.maxpool = nn.MaxPool2d(kernel_size=KERNEL)
self.decoder_conv1 = nn.ConvTranspose2d(in_channels=CHANNELS, out_channels=CHANNELS, kernel_size=KERNEL)
self.decoder_conv2 = nn.ConvTranspose2d(in_channels=CHANNELS, out_channels=CHANNELS//2, kernel_size=KERNEL)
self.decoder_conv3 = nn.ConvTranspose2d(in_channels=CHANNELS//2, out_channels=CHANNELS//2, kernel_size=KERNEL)
self.decoder_conv4 = nn.ConvTranspose2d(in_channels=CHANNELS//2, out_channels=1, kernel_size=KERNEL)
def forward(self, x):
images = x['historical_sat_images']
images = normalise_images_in_model(images, self.device)
# Pass data through the network :)
# ENCODER
out = F.relu(self.encoder_conv1(images))
out = F.relu(self.encoder_conv2(out))
out = F.relu(self.encoder_conv3(out))
out = F.relu(self.encoder_conv4(out))
out = self.maxpool(out)
# DECODER
out = F.relu(self.decoder_conv1(out))
out = F.relu(self.decoder_conv2(out))
out = F.relu(self.decoder_conv3(out))
out = self.decoder_conv4(out)
return out
def _training_or_validation_step(self, batch, is_train_step):
y_hat = self(batch)
y = batch['target_sat_images']
y = normalise_images_in_model(y, self.device)
y = y[..., 40:-40, 40:-40] # Due to the CNN stride, the output image is 48 x 48
loss = F.mse_loss(y_hat, y)
tag = "Loss/Train" if is_train_step else "Loss/Validation"
self.log_dict({tag: loss}, on_step=is_train_step, on_epoch=True)
return loss
def training_step(self, batch, batch_idx):
return self._training_or_validation_step(batch, is_train_step=True)
def validation_step(self, batch, batch_idx):
return self._training_or_validation_step(batch, is_train_step=False)
def configure_optimizers(self):
optimizer = torch.optim.Adam(self.parameters(), lr=0.001)
return optimizer
# -
model = LitAutoEncoder()
trainer = pl.Trainer(gpus=1, max_epochs=400, terminate_on_nan=False)
# %%time
trainer.fit(model, train_dataloader=dataloader)
|
notebooks/20.0_simplify_data_loading.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import ee
import geodatatool
Map = geodatatool.Map()
Map
geodatatool.ee_initialize()
# +
dem = ee.Image('USGS/SRTMGL1_003')
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# -
ee_layer = geodatatool.ee_tile_layer(dem, vis_params, "DEM")
Map.add_layer(ee_layer)
# +
# alternate name can be used for Map.add_layer, conventional to JavaScript
# Map.addLayer(dem, vis_params, "DEM")
# -
import ipywidgets as widgets
slider = widgets.FloatSlider(min=0, max=1, step=0.01)
slider
widgets.jslink((ee_layer, "opacity"), (slider, "value"))
from ipyleaflet import WidgetControl
control = WidgetControl(widget=slider, position="bottomright")
Map.add_control(control)
|
docs/notebooks/earthengine_intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basics tutorial
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# ## Exercise 1: Matrix manipulations and numerics warm-up
# +
filepath = '/Users/RuoyanWang/Documents/LEIDEN/SPRING2019/MAS/Modern-Astrostatistics/AllExercises/DataFiles/'
sn_covmat = pd.read_csv(filepath+'/SN_covmat.txt', sep='\s+', header=None)
sn_covmat
# -
np.sqrt(len(sn_covmat.values))
matrix = np.reshape(sn_covmat.values, (31,31))
cov_matrix = np.cov(matrix)
plt.figure(figsize=(8,8))
plt.imshow(cov_matrix)
plt.colorbar()
corr_matrix = np.corrcoef(matrix)
plt.figure(figsize=(8,8))
plt.imshow(corr_matrix)
plt.colorbar()
np.where((corr_matrix <= 0.99)*(corr_matrix >= 0.6))
corr_matrix[np.where((corr_matrix <= 0.99)*(corr_matrix >= 0.6))]
np.where((corr_matrix >= -0.99)*(corr_matrix <= -0.7))
corr_matrix[np.where((corr_matrix >= -0.99)*(corr_matrix <= -0.7))]
np.where((corr_matrix <= 0.99)*(corr_matrix >= 0.45))
corr_matrix[np.where((corr_matrix <= 0.99)*(corr_matrix >= 0.45))]
np.where((corr_matrix >= -0.99)*(corr_matrix <= -0.5148))
corr_matrix[np.where((corr_matrix >= -0.99)*(corr_matrix <= -0.5148))]
plt.figure(figsize=(8,6))
plt.errorbar(sn_covmat.index, sn_covmat.values)
#plt.xlim(30,35)
np.linalg.det(cov_matrix)
inv_cov_matrix = np.linalg.inv(cov_matrix)
plt.figure(figsize=(8,8))
plt.imshow(inv_cov_matrix)
plt.colorbar()
# ## Exercise 3: Bargaining for funding.
# ### $\sigma_m = \frac{\sigma}{\sqrt{N}}$
#
# ### $ f(x)=\begin{cases}
# \frac {1}{2 \sigma \sqrt{3}} & \mbox{for }-\sigma\sqrt{3} \le x-\mu \le \sigma\sqrt{3} \\
# 0 & \text{otherwise}
# \end{cases}$
|
Basics_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W0D4_Calculus/W0D4_Tutorial2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Neuromatch Academy: Week 0, Day 4, Tutorial 2
# # Differential Equations
#
# __Content creators:__ <NAME>, <NAME>
#
# __Content reviewers:__
#
# __Production editors:__
# ---
# # Tutorial Objectives
#
# A great deal of neuroscience can be modelled using differential equations, from gating channels to single neurons to a network of neurons to blood flow, to behaviour. A simple way to think about differential equations is they are equations that describe how something changes.
#
# The most famous of these in neuroscience is the Nobel Prize winning Hodgkin Huxley equation, which describes a neuron by modelling the gating of each axon. But we will not start there; we will start a few steps back.
#
# Differential Equations are mathematical equations that describe how something like population or a neuron changes over time. The reason why differential equations are so useful is they can generalise a process such that one equation can be used to describe many different outcomes.
# The general form of a first order differential equation is:
#
# \begin{align*}
# \frac{d}{dt}y(t)&=f(t,y(t))\\
# \end{align*}
#
# which can be read as "the change in a process $y$ over time $t$ is a function $f$ of time $t$ and itself $y$". This might initially seem like a paradox as you are using a process $y$ you want to know about to describe itself, a bit like the MC Escher drawing of two hands painting [each other](https://en.wikipedia.org/wiki/Drawing_Hands). But that is the beauty of mathematics - this can be solved some of time, and when it cannot be solved exactly we can use numerical methods to estimate the answer (see next tutorial).
#
#
# In this tutorial, we will see how __differential equations are motivated by observations of physical responses.__ We will break down the population differential equation, then the integrate and fire model, which leads nicely into raster plots and frequency-current curves to rate models.
#
# **Steps:**
# - Get an intuitive understanding of a linear population differential equation (humans, not neurons)
# - Visualize the relationship between the change in population and the population
# - Breakdown the Leaky Integrate and Fire (LIF) differential equation
# - Code the exact solution of an LIF for a constant input
# - Visualize and listen to the response of the LIF for different inputs
#
# + cellView="form"
#@markdown Tutorial Slides
# you should link the slides for all tutorial videos here (we will store pdfs on osf)
from IPython.display import HTML
HTML('<iframe src="" frameborder="0" width="960" height="569" allowfullscreen="true" mozallowfullscreen="true" webkitallowfullscreen="true"></iframe>')
# +
# Imports
import numpy as np
import matplotlib.pyplot as plt
# + cellView="form"
# @title Figure Settings
import IPython.display as ipd
from matplotlib import gridspec
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
# use NMA plot style
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
my_layout = widgets.Layout()
# + cellView="form"
# @title Plotting Functions
def plot_dPdt(alpha=.3):
""" Plots change in population over time
Args:
alpha: Birth Rate
Returns:
A figure two panel figure
left panel: change in population as a function of population
right panel: membrane potential as a function of time
"""
with plt.xkcd():
time=np.arange(0, 10 ,0.01)
fig = plt.figure(figsize=(12,4))
gs = gridspec.GridSpec(1, 2)
## dpdt as a fucntion of p
plt.subplot(gs[0])
plt.plot(np.exp(alpha*time), alpha*np.exp(alpha*time))
plt.xlabel(r'Population $p(t)$ (millions)')
plt.ylabel(r'$\frac{d}{dt}p(t)=\alpha p(t)$')
## p exact solution
plt.subplot(gs[1])
plt.plot(time, np.exp(alpha*time))
plt.ylabel(r'Population $p(t)$ (millions)')
plt.xlabel('time (years)')
plt.show()
def plot_V_no_input(V_reset=-75):
"""
Args:
V_reset: Reset Potential
Returns:
A figure two panel figure
left panel: change in membrane potential as a function of membrane potential
right panel: membrane potential as a function of time
"""
E_L=-75
tau_m=10
t=np.arange(0,100,0.01)
V= E_L+(V_reset-E_L)*np.exp(-(t)/tau_m)
V_range=np.arange(-90,0,1)
dVdt=-(V_range-E_L)/tau_m
with plt.xkcd():
time=np.arange(0, 10, 0.01)
fig = plt.figure(figsize=(12, 4))
gs = gridspec.GridSpec(1, 2)
plt.subplot(gs[0])
plt.plot(V_range,dVdt)
plt.hlines(0,min(V_range),max(V_range), colors='black', linestyles='dashed')
plt.vlines(-75, min(dVdt), max(dVdt), colors='black', linestyles='dashed')
plt.plot(V_reset,-(V_reset - E_L)/tau_m, 'o', label=r'$V_{reset}$')
plt.text(-50, 1, 'Positive')
plt.text(-50, -2, 'Negative')
plt.text(E_L - 1, max(dVdt), r'$E_L$')
plt.legend()
plt.xlabel('Membrane Potential V (mV)')
plt.ylabel(r'$\frac{dV}{dt}=\frac{-(V(t)-E_L)}{\tau_m}$')
plt.subplot(gs[1])
plt.plot(t,V)
plt.plot(t[0],V_reset,'o')
plt.ylabel(r'Membrane Potential $V(t)$ (mV)')
plt.xlabel('time (ms)')
plt.ylim([-95, -60])
plt.show()
## LIF PLOT
def plot_IF(t, V,I,Spike_time):
"""
Args:
t : time
V : membrane Voltage
I : Input
Spike_time : Spike_times
Returns:
figure with three panels
top panel: Input as a function of time
middle panel: membrane potential as a function of time
bottom panel: Raster plot
"""
with plt.xkcd():
fig = plt.figure(figsize=(12, 4))
gs = gridspec.GridSpec(3, 1, height_ratios=[1, 4, 1])
# PLOT OF INPUT
plt.subplot(gs[0])
plt.ylabel(r'$I_e(nA)$')
plt.yticks(rotation=45)
plt.hlines(I,min(t),max(t),'g')
plt.ylim((2, 4))
plt.xlim((-50, 1000))
# PLOT OF ACTIVITY
plt.subplot(gs[1])
plt.plot(t,V)
plt.xlim((-50, 1000))
plt.ylabel(r'$V(t)$(mV)')
# PLOT OF SPIKES
plt.subplot(gs[2])
plt.ylabel(r'Spike')
plt.yticks([])
plt.scatter(Spike_time, 1 * np.ones(len(Spike_time)), color="grey", marker=".")
plt.xlim((-50, 1000))
plt.xlabel('time(ms)')
plt.show()
## Plotting the differential Equation
def plot_dVdt(I=0):
"""
Args:
I : Input Current
Returns:
figure of change in membrane potential as a function of membrane potential
"""
with plt.xkcd():
E_L = -75
tau_m = 10
V = np.arange(-85, 0, 1)
g_L = 10.
fig = plt.figure(figsize=(6, 4))
plt.plot(V,(-(V-E_L) + I*10) / tau_m)
plt.hlines(0, min(V), max(V), colors='black', linestyles='dashed')
plt.xlabel('V (mV)')
plt.ylabel(r'$\frac{dV}{dt}$')
plt.show()
# + cellView="form"
# @title Helper Functions
## EXACT SOLUTION OF LIF
def Exact_Integrate_and_Fire(I,t):
"""
Args:
I : Input Current
t : time
Returns:
Spike : Spike Count
Spike_time : Spike time
V_exact : Exact membrane potential
"""
Spike = 0
tau_m = 10
R = 10
t_isi = 0
V_reset = E_L = -75
V_exact = V_reset * np.ones(len(t))
V_th = -50
Spike_time = []
for i in range(0, len(t)):
V_exact[i] = E_L + R*I + (V_reset - E_L - R*I) * np.exp(-(t[i]-t_isi)/tau_m)
# Threshold Reset
if V_exact[i] > V_th:
V_exact[i-1] = 0
V_exact[i] = V_reset
t_isi = t[i]
Spike = Spike+1
Spike_time = np.append(Spike_time, t[i])
return Spike, Spike_time, V_exact
# -
# ---
# # Section 1: Population Differential Equation
# + cellView="form"
#@title Video 1: Differential Equations Introduction
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="nWKgnouNzGY", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
# -
#
#
# To get an intuitive feel of a differential equations, we will start with a population differential equation, which models the change in population [1], that is human population not neurons, we will get to neurons later. Mathematically it is written like:
# \begin{align*}
# \\
# \frac{d}{dt}\,p(t) &= \alpha p(t),\\
# \end{align*}
#
# where $p(t)$ is the population of the world and $\alpha$ is a parameter representing birth rate.
#
# Another way of thinking about the models is that the equation
# \begin{align*}
# \\
# \frac{d}{dt}\,p(t) &= \alpha p(t),\\
# \text{can be written as:}\\
# \text{"Change in Population"} &= \text{ "Birth rate times Current population."}
# \end{align*}
#
# The equation is saying something reasonable maybe not the perfect model but a good start.
# + cellView="form"
#@title Video 2: The Population Equation
video = YouTubeVideo(id="yaQdEayD9Nw", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
# -
# ## Coding Exercise 1: Plot the Change in Population
#
# Let's investigate what the rate of change of population looks like as a function of the current population.
#
# Write Python code to calculate the change in population `dpdt` with an `alpha=0.3` as a function of population `p`. Uncomment the plot line and interpret the plot.
# +
p = np.arange(0, 100, 0.1)
########################################################################
## TODO for students: calculate the dp/dt
## Complete line of code
raise NotImplementedError("Calculate the dp/dt")
########################################################################
dpdt = ...
## Uncomment to plot answer
fig = plt.figure(figsize=(6, 4))
#plt.plot(p, dpdt)
plt.xlabel(r'Population $p(t)$ (millions)')
plt.ylabel(r'$\frac{d}{dt}p(t)=\alpha p(t)$')
plt.show()
# +
# to_remove solution
p = np.arange(0, 100, 0.1)
with plt.xkcd():
dpdt = 0.3*p
fig = plt.figure(figsize=(6, 4))
plt.plot(p, dpdt)
plt.xlabel(r'Population $p(t)$ (millions)')
plt.ylabel(r'$\frac{d}{dt}p(t)=\alpha p(t)$')
plt.show()
# -
# ## Think! 1: Linear Differential Equations and the Population Equation
#
# 1. Why is the population differential equation is known as a linear differential equation?
#
# 2. How does population size affect the rate of change of the population?
#
# +
# to_remove explanation
"""
1. The plot of $\frac{dp}{dt}$ is a line, which is why the differential
equation is known as a linear differential equation.
2. As the population increases, the change of population increases. A
population of 20 has a change of 6 while a population of 100 has a change of
30. This makes sense - the larger the population the larger the change.
"""
# -
# ## Section 1.1: Exact Solution of the Population Equation
# ### Section 1.1.1: Initial Condition
# The linear population differential equation is known as an initial value differential equation because we need an initial population value to solve it, so here we will set our initial population to:
#
# \begin{align*}
# &p(0)=1.\\
# \end{align*}
#
# Different initial conditions will lead to different answers, but they will not change the differential equation. This is one of the strengths of a differential equation.
# ### Section 1.1.2: Exact Solution
# To calculate the exact solution of a differential equation, we must integrate both sides and use some mathematical tricks - the harder the equation the more obscure the trick.
#
# Given the linear population equation
# \begin{align*}
# \frac{d}{dt}\,p(t) &= \alpha p(t),\\\\
# p(0)=1,\\
# \end{align*}
# has the exact solution:
# \begin{align*}
# p(t)&=e^{\alpha t}.\\
# \end{align*}
#
# The exact solution written in words is:
#
# \begin{align*}
# \text{"Population"}&=\text{"grows/declines exponentially as a function time and birth rate"}.\\
# \end{align*}
#
# Most differential equations do not have a known exact solution, so in the next tutorial on numerical methods we will show how the solution can be estimated.
#
# A small aside: a good deal of progress in mathematics was due to mathematicians writing taunting letters to each other saying they had a trick that could solve something better than everyone else. So do not worry too much about the tricks.
# #### Coding Exercise 1.1.2: Exact Solution of the Population Equation
# Let's consider the population differential equation with a birth rate $\alpha=0.3$:
#
# \begin{align*}
# \frac{d}{dt}\,p(t) = 0.3 p(t),\\
# \text{with the initial condition}\\
# p(0)=1.\\
# \end{align*}
#
# It has an exact solution
# \begin{align*}
# \\
# p(t)=e^{0.3 t}.
# \end{align*}
#
# Code and plot the exact solution of the population differential equation $p(t)$ for $\alpha=0.3$ for the time period $t=0$ to $t=10$.
#
# Use the function `np.exp()` for the exponential.
# +
t = np.arange(0, 10, 0.1) # Time from 0 to 10 years in 0.1 steps
########################################################################
## TODO for students: calculate the exact solution of p
# Fill out function and remove
raise NotImplementedError("Calculate the exact solution of p")
########################################################################
p = ...
## Uncomment to plot answer
fig = plt.figure(figsize=(6, 4))
#plt.plot(t, p)
plt.ylabel('Population (millions)')
plt.xlabel('time (years)')
plt.show()
# +
# to_remove solution
t = np.arange(0, 10, 0.1) # Time from 0 to 10 years in 0.1 steps
with plt.xkcd():
p = np.exp(0.3 * t)
fig = plt.figure(figsize=(6, 4))
plt.plot(t, p)
plt.ylabel('Population (millions)')
plt.xlabel('time (years)')
plt.show()
# -
# ## Section 1.2: Parameters (Birth Rate $\alpha$)
# One of the goals when design a differential equation is to make it generalisable. For example, so that the differential equation will give reasonable solutions for different countries with different birth rates $\alpha$.
#
#
# ## Interactive Demo 1.2: Interactive Parameter Change
# Play with the widget to see the relationship between $\alpha$ and the population differential equation as a function of population (left-hand side), and the population solution as a function of time(right-hand side). Pay close attention to the transision point from positive to negative.
# + cellView="form"
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
alpha=widgets.FloatSlider(.3, min=-1., max=1., step=.1, layout=my_layout)
)
def Pop_widget(alpha):
plot_dPdt(alpha=alpha)
plt.show()
# -
# ### Think! 1.2: How do changing parameters of the Population Equation affect the outcome?
#
# 1. What happens when $\alpha < 0$?
# 2. What happens when $\alpha > 0$?
# 3. What happens when $\alpha = 0$?
#
# +
#to_remove explanation
"""
1. Negative values of alpha result in an exponential decrease.
2. Positive Values of alpha in an exponential increase.
3. Alpha equal to 0 is a unique point known as an equilibrium point when the
dp/dt=0 and there is no change in population. This is known as a stable point.
"""
# -
# ## Secton 1.3: Population Differential Equation Summary
#
# The population differential equation is an over-simplification and has some very obvious limitations:
# 1. Population growth is not exponential as there are limited number of resources so the population will level out at some point.
# 2. It does not include any external factors on the populations like weather, predators and preys.
#
# These kind of limitations can be addressed by extending the model.
#
#
# While it might not seem that the population equation has direct relevance to neuroscience a similar equation is used to describe the accumulation of evidence for decision making known as the Drift Diffusion Model which you will see in more detail in the Linear System day in Neuromatch (W2D2).
#
#
# Another differential equation that is similar to the population equation is the Leaky Integrate and Fire Model which we will explore below.
# ---
# # Section 2: The Leaky Integrate and Fire Model
#
# + cellView="form"
#@title Video 3: Leaky Integrate and Fire Model
video = YouTubeVideo(id="htYJ84aVK3s", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
# -
# The Leaky Integrate and Fire Model is a linear differential equation that describes the membrane potential ($V$) of a single neuron which was proposed by <NAME> in 1907 [2].
#
# The subthreshold membrane potential dynamics of a LIF neuron is described by
# \begin{align}
# \tau_m\frac{dV}{dt} = -(V-E_L) + R_mI\,
# \end{align}
#
#
# where $\tau_m$ is the time constant, $V$ is the membrane potential, $E_L$ is the resting potential, $R_m$ is membrance resistance, and $I$ is the external input current.
#
# In the next few sections, we will break down the equation and then build it back up to get an intuitive feel of the different facets of the differential equation.
# ## Section 2.1: LIF without Input
#
# First, we will simplify the equation by removing the input, which gives the equation
#
# \begin{align}
# \tau_m\frac{dV}{dt} &= -V+E_L,\\
# \end{align}
#
# which can be written in words as:
#
# \begin{align}
# \begin{matrix}\text{"Time constant multiplied by the} \\ \text{change in membrane potential"}\end{matrix}&=\begin{matrix}\text{"Minus Current} \\ \text{membrane potential"} \end{matrix}+
# \begin{matrix}\text{"resting potential"}\end{matrix}.\\
# \end{align}
#
#
# The equation can be re-arranged to look even more like the population equation:
#
# \begin{align}
# \frac{dV}{dt} &= \frac{-(V-E_L)}{\tau_m}.\\
# \end{align}
#
# ### Coding Exercise 2.1: LIF without Input
# Code the right hand side `dVdt` of the differential equation,
# \begin{align}
# \frac{dV}{dt} &= \frac{-(V-E_L)}{\tau_m},\\
# \end{align}
# with the parameters set as:
# * `E_L = -75`
# * `V_reset = -50`
# * `tau_m = 10`
#
# and uncommment the code to plot for 'dVdt' as a function of $V$ between -90 and 0.
#
# Take note of where the curve of `dVdt` cuts the dashed line indicating `dVdt = 0`.
#
# +
# Parameter definition
E_L = -75
tau_m = 10
# Range of Values of V
V = np.arange(-90, 0, 1)
########################################################################
## TODO for students: Complete line of code for the dVdt
# Fill out function and remove
raise NotImplementedError("Complete line of code for the dVdt")
########################################################################
dVdt = ...
# Uncomment code below to plot
fig = plt.figure(figsize=(6, 4))
# plt.plot(V, dVdt)
plt.vlines(-75, min(dVdt), max(dVdt), colors='black', linestyles='dashed')
plt.text(E_L, max(dVdt) + 1, r'$E_L$')
plt.hlines(0, min(V), max(V), colors='black', linestyles='dashed')
plt.text(-50, 1, 'Positive')
plt.text(-50, -2, 'Negative')
plt.xlabel(r'$V(t)$ (mV)')
plt.ylabel(r'$\frac{dV}{dt}=\frac{-(V-E_L)}{\tau_m}$')
plt.ylim(-8, 2)
plt.show()
# +
# to_remove solution
# Parameter definition
E_L = -75
tau_m = 10
# Range of Values of V
V = np.arange(-90, 0, 1)
dVdt = -(V - E_L) / tau_m
with plt.xkcd():
fig = plt.figure(figsize=(6, 4))
plt.plot(V, dVdt)
plt.hlines(0, min(V), max(V), colors='black', linestyles='dashed')
plt.vlines(-75, min(dVdt), max(dVdt), colors='black', linestyles='dashed')
plt.text(-50, 1, 'Positive')
plt.text(-50, -2, 'Negative')
plt.text(E_L, max(dVdt) + 1, r'$E_L$')
plt.xlabel(r'$V(t)$ (mV)')
plt.ylabel(r'$\frac{dV}{dt}=\frac{-(V-E_L)}{\tau_m}$')
plt.ylim(-8, 2)
plt.show()
# -
# ### Think! 2.1: Effect on Membrane Potential $V$ on the LIF Model
#
# 1. What is the effect on $\frac{dV}{dt}$ when $V>-75$ mV?
# 2. What is the effect on $\frac{dV}{dt}$ when $V<-75$ mV
# 3. What is the effect on $\frac{dV}{dt}$ when $V=-75$ mV?
# +
#to_remove explanation
"""
1. For $V>-75$ mV, the derivative is negative.
2. For $V<-75$ mV, the derivative is positive.
3. For $V=-75$ mV, the derivative is equal to $0$ is and a stable point.
"""
# -
# ### Section 2.1.1: Exact Solution of the LIF model without Input
#
# Similar to the population equation, we need an initial membrane potential at time $0$ to solve the LIF model.
#
# With this equation
# \begin{align}
# \frac{dV}{dt} &= \frac{-(V-E_L)}{\tau_m}\,\\
# V(0)&=V_{reset},
# \end{align}
# where is $V_{reset}$ is called the reset potential.
#
# The LIF model has the exact solution:
# \begin{align*}
# V(t)=&\ E_L+(V_{reset}-E_L)e^{\frac{-t}{\tau_m}}\\
# \text{ which can be written as: }\\
# \begin{matrix}\text{"Current membrane} \\ \text{potential}"\end{matrix}=&\text{"Resting potential"}+\begin{matrix}\text{"Reset potential minus resting potential} \\ \text{times exponential with rate one over time constant."}\end{matrix}\\
# \end{align*}
# #### Interactive Demo 2.1.1: Initial Condition $V_{reset}$
# This exercise is to get an intuitive feel of how the different initial conditions $V_{reset}$ impact the differential equation of the LIF and the exact soluation for the equation:
#
# \begin{align}
# \frac{dV}{dt} &= \frac{-(V-E_L)}{\tau_m}\,\\
# \end{align}
# with the parameters set as:
# * `E_L = -75,`
# * `tau_m = 10.`
#
# The panel on the left-hand side plot the change in membrane potential $\frac{dV}{dt}$ is a function of membrane potential $V$ and right-hand side panel plots the exact solution $V$ as a function of time $t,$ the green dot in both panels is the reset potential $V_{reset}$.
#
# Pay close attention to when $V_{reset}=E_L=-75$mV.
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
V_reset=widgets.FloatSlider(-77., min=-91., max=-61., step=2,
layout=my_layout)
)
def V_reset_widget(V_reset):
plot_V_no_input(V_reset)
# -
# #### Think! 2.1.1: Effect of $V_{reset}$ on the solution
#
#
# 1. How does the solution look with initial values of $V_{reset} < -75$?
# 2. How does the solution look with initial values of $V_{reset} > -75$?
# 3. How does the solution look with initial values of $V_{reset} = -75$?
#
#
# +
#to_remove explanation
"""
1. Initial Values of $V_{reset} < -75$ result in the solution increasing to
-75mV because $\frac{dV}{dt} > 0$.
2. Initial Values of $V_{reset} > -75$ result in the solution decreasing to
-75mV because $\frac{dV}{dt} < 0$.
3. Initial Values of $V_{reset} = -75$ result in a constant $V = -75$ mV
because $\frac{dV}{dt} = 0$ (Stable point).
"""
# -
# ## Section 2.2: LIF with Input
# We will re-introduce the input $I$ and membrane resistance $R_m$ giving the original equation:
#
# \begin{align}
# \tau_m\frac{dV}{dt} = -(V-E_L) + \color{blue}{R_mI}\,
# \end{align}
#
# the input can be other neurons or sensory information.
# ### Interactive Demo 2.2: The Impact of Input
# The interactive plot below mainpulates $I$ in the differential equation.
# With increasing input what would you expect to happen to the solution.
# + cellView="form"
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
I=widgets.FloatSlider(3., min=0., max=20., step=2,
layout=my_layout)
)
def Pop_widget(I):
plot_dVdt(I=I)
plt.show()
# -
# ### Think! 2.2: Effect of increasing Input
#
# 1. As $I$ increases, less and less of $\frac{dV}{dt}$ is below 0. How would this impact the solution?
#
#
# +
#to_remove explanation
"""
"""
# -
# ### Section 2.2.1: LIF Exact Solution
#
# The LIF with a constant input is a linear differential equation has a known exact solution:
# \begin{align*}
# V(t)=&\ E_L+R_mI+(V_{reset}-E_L-R_mI)e^{\frac{-t}{\tau_m}}\\
# \text{which is written as:}\\
# \begin{matrix}\text{"Current membrane} \\ \text{potential"}\end{matrix}=&\text{"Resting potential"}+\begin{matrix}\text{"Reset potential minus resting potential} \\ \text{times exponential with rate one over time constant." }\end{matrix}\\
# \end{align*}
# #### Coding Exercise 2.2.1: LIF Exact Solution
# Code the exact solution `V`:
# $$
# V(t)= E_L+RI+(V_{reset}-E_L-RI)e^{\frac{-t}{\tau_m}},\\
# $$
# with the parameters set as:
# * `V_reset = -75,`
# * `E_L = -75,`
# * `tau_m = 10,`
# * `R_m = 10,`
# * `I = 10,`
# of the LIF model and uncomment the code to plot the result.
#
# Ask yourself, does the result make biological sense?
# If not, what would you change?
# +
dt = 0.5
t_rest = 0
t = np.arange(0, 1000, dt)
tau_m = 10
R_m = 10
V_reset = E_L = -75
I = 10
########################################################################
## TODO for students: Complete line of code for the exact Solution V
# Fill out function and remove
raise NotImplementedError("Complete line of code for the exact Solution V")
########################################################################
V = ...
# Uncomment below to plot the result
fig = plt.figure(figsize=(6, 4))
#plt.plot(t,V)
plt.ylabel('V (mV)')
plt.xlabel('time (ms)')
plt.show()
# +
# to_remove solution
dt = 0.5
t_rest = 0
t = np.arange(0, 1000, dt)
tau_m = 10
R_m = 10
V_reset = E_L = -75
I = 10
V = E_L + R_m*I + (V_reset - E_L - R_m*I) * np.exp(-(t)/tau_m)
with plt.xkcd():
fig = plt.figure(figsize=(6, 4))
plt.plot(t,V)
plt.ylabel('V (mV)')
plt.xlabel('time (ms)')
plt.show()
# -
# ## Section 2.3: Maths is one thing, but biology matters
# While the mathematics of the exact solution is exact, it is not biologically valid as neurons spike and defintely do not plateau at a very positive value.
#
# To model the firing of a spike, we must have a threshold voltage $V_{th}$ such that if the voltage $V(t)$ goes above it, the neuron spikes
# $$V(t)>V_{th}.$$
# We must record the time of spike $t_{isi}$ and count the number of spikes
# $$t_{isi}=t, $$
# $$𝑆𝑝𝑖𝑘𝑒=𝑆𝑝𝑖𝑘𝑒+1.$$
# Then reset the membrane voltage $V(t)$
# $$V(t_{isi} )=V_{Reset}.$$
# To take into account the spike the exact solution becomes:
# \begin{align*}
# V(t)=&\ E_L+R_mI+(V_{reset}-E_L-R_mI)e^{\frac{-(t-t_{isi})}{\tau_m}},&\qquad V(t)<V_{th} \\
# V(t)=&V_{reset},&\qquad V(t)>V_{th}\\
# Spike=&Spike+1,&\\
# t_{isi}=&t,\\
# \end{align*}
# while this does make the neuron spike, it introduces a discontinuity which is not as elegant mathematically as it could be, but it gets results so that is good.
# ### Interactive Demo 2.3.1: Listen to the Spikes
# This exercise show the relationship between firing rate and the Input for exact solution `V` of the LIF:
# $$
# V(t)=\ E_L+R_mI+(V_{reset}-E_L-R_mI)e^{\frac{-(t-t_{isi})}{\tau_m}},
# $$
# with the parameters set as:
# * `V_reset = -75,`
# * `E_L = -75,`
# * `tau_m = 10,`
# * `R_m = 10.`
#
#
# Below is a figure with with three panels;
# * the top panel is the input, $I,$
# * the middle panel is the membrane potential $V(t)$ to illustrative the spike $V(t)$ is set to $0$ and then reset to $-75$ mV,
# * the bottom panel is the raster plot with each dot indicating a spike.
#
# As electrophysiologist normally listen to spikes when conducting experiments, so listen to the music of different firing rates by changing the input value $I.$
# + cellView="form"
# @markdown Make sure you execute this cell to be able to hear the neuron
I = 3
t = np.arange(0, 1000, dt)
Spike, Spike_time, V = Exact_Integrate_and_Fire(I, t)
plot_IF(t, V, I, Spike_time)
ipd.Audio(V, rate=len(V))
# -
# ### Interactive Demo 2.3.2: Input on Spikes
# Manipulate the input into the LIF to see the impact of input on the firing pattern (rate).
# + cellView="form"
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
I=widgets.FloatSlider(3, min=2.0, max=4., step=.1,
layout=my_layout)
)
def Pop_widget(I):
Spike, Spike_time, V = Exact_Integrate_and_Fire(I, t)
plot_IF(t, V, I, Spike_time)
# -
# ### Think! 2.3: Is the model biologically valid?
#
# 1. What is the effect of $I$ on spiking?
# 2. Is this biologically valid?
# +
#to_remove explanation
"""
1. As $I$ increases, the number of spikes increases.
2.
"""
# -
# ## Section 2.4 Firing Rate as a function of Input
#
# The firing frequency of a neuron plotted as a fucntion of current is called an input-output curve (F–I curve). It is also known as a transfer function, which you came accross in the previous tutorial. This function is one of the starting points for the rate model, which extends from modelling single neurons to collections of neurons.
#
# By fitting this to a function, we can start to generalise the firing pattern of many neurons, which can be used to build rate models which is discussed later in Neuromatch.
# + cellView="form"
# @markdown *Execture this cell to visualize the FI curve*
I_range = np.arange(2.0, 4.0, 0.1)
Spike_rate = np.ones(len(I_range))
for i, I in enumerate(I_range):
Spike_rate[i], _, _ = Exact_Integrate_and_Fire(I, t)
with plt.xkcd():
fig = plt.figure(figsize=(6, 4))
plt.plot(I_range,Spike_rate)
plt.xlabel('Input Current (nA)')
plt.ylabel('Spikes per Second (Hz)')
plt.show()
# -
# ---
# ## Summary of LIF model
#
# The LIF model is a very nice differential equation to start with in computational neuroscience as it has been used as a building block for many papers that simulate neuronal response.
#
# __Strenghts of LIF model:__
# + Has an exact solution;
# + Easy to interpret;
# + Great to build network of neurons.
#
# __Weaknesses of the LIF model:__
# - Spiking is a discontinuity;
# - Abstraction from biology;
# - Cannot generate different spiking patterns.
#
#
# ---
# # Take Home Message
# + cellView="form"
#@title Video 4: Take Home Message and Outro
video = YouTubeVideo(id="TcKWdBD3jcA", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
# -
# In this tutorial, we have have seen two differential equations, the population differential equations and the leaky integrate and fire model.
#
#
# We learned about:
# * The motivation for differential equations.
# * An intuitive relationship between the solution and the form of the differential equation.
# * How different parameters of the differential equation impact the solution.
# * The strengths and limitations of the simple differential equations.
#
# ---
# # Links to Neuromatch Days
#
# Differential equations turn up in a number of different Neuromatch days:
# * The LIF model is discussed in more details in Model Types (Week 1 Day 1) and Real Neurons (Week 2 Day 3).
# * Drift Diffusion model which is a differential equation for decision making is discussed in Linear Systems (Week 2 Day 2).
# * Systems of differential equations are discussed in Linear Systems (Week 2 Day 2) and Dynamic Networks (Week 2 Day 4).
#
#
# ---
# # References
# 1. <NAME>, (1920) Analytical note on certain rhythmic relations inorganic systems.Proceedings of the National Academy of Sciences,6(7):410–415,1920.
#
# 2. <NAME>, <NAME>. Lapicque's 1907 paper: from frogs to integrate-and-fire. Biol Cybern. 2007 Dec;97(5-6):337-9. doi: 10.1007/s00422-007-0190-0. Epub 2007 Oct 30. PMID: 17968583.
#
#
#
# # Bibliography
# 1. <NAME>., & <NAME>. (2001). Theoretical neuroscience: computational and mathematical modeling of neural systems. Computational Neuroscience Series.
# 2. <NAME>. Nonlinear dynamics and chaos: with applications to physics, biology, chemistry, and engineering (studies in nonlinearity), Westview Press; 2 edition (29 July 2014)
#
# ## Supplemental Popular Reading List
# 1. <NAME>. (2021). Models of the Mind: How Physics, Engineering and Mathematics Have Shaped Our Understanding of the Brain. Bloomsbury Publishing.
# 2. <NAME>. (2004). Sync: The emerging science of spontaneous order. Penguin UK.
#
# ## Popular Podcast
# 1. <NAME>. (Host). (2020-), Joy of X https://www.quantamagazine.org/tag/the-joy-of-x/ Quanta Magazine
#
|
tutorials/W0D4_Calculus/W0D4_Tutorial2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Landsat 8 NDVI Analysis on the Cloud
#
# This notebook demonstrates a "Cloud Native" analysis of [Normalized Difference Vegetation Index (NDVI)](https://en.wikipedia.org/wiki/Normalized_difference_vegetation_index) using [Landsat 8 satellite imagery](https://landsat.usgs.gov/landsat-8).
#
# **To run, simply hit Shift+Enter to run each code block**
#
# By "Cloud Native" we mean that images are not downloaded to your local machine. Instead, calculations are performed efficiently in parallel across many distributed machines on Google Cloud ([where the imagery is stored](https://cloud.google.com/storage/docs/public-datasets/landsat)).
#
# This workflow is possible because the Landsat 8 data is stored in [Cloud-Optimized Geotiff](http://www.cogeo.org) format, which can be accessed remotely via [xarray](http://xarray.pydata.org/en/stable/) and [rasterio](https://rasterio.readthedocs.io/en/latest/) Python libraries. Interactive, dynamically updating visualization is done with [Holoviews](http://holoviews.org). Distributed computing is enabled through a [Pangeo](http://pangeo.io) JupyterHub deployment with [Dask Kubernetes](https://github.com/dask/dask-kubernetes).
#
# Created on August 30, 2018 by:
#
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>
# +
# Import required libraries
import os
import pandas as pd
import rasterio
import xarray as xr
import requests
import geoviews as gv
import holoviews as hv
import hvplot.xarray
import hvplot.pandas
import shapely
#import numpy as np
import dask
from dask_kubernetes import KubeCluster
from dask.distributed import Client
from dask.distributed import wait, progress
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Print package versions
print('Xarray version: ', xr.__version__)
print('Rasterio version: ', rasterio.__version__)
print('dask version: ', dask.__version__)
print('hvplot version: ', hvplot.__version__)
# +
# Set environment variables for cloud-optimized-geotiffs efficiency
#os.environ['GDAL_DISABLE_READDIR_ON_OPEN']='YES'
#os.environ['CPL_VSIL_CURL_ALLOWED_EXTENSIONS']='TIF'
env = rasterio.Env(GDAL_DISABLE_READDIR_ON_OPEN='EMPTY_DIR',
CPL_VSIL_CURL_USE_HEAD=False,
CPL_VSIL_CURL_ALLOWED_EXTENSIONS='TIF',
#CPL_DEBUG=True,
#CPL_CURL_VERBOSE=True,
#VSI_CACHE=True, #The cache size defaults to 25 MB, but can be modified by setting the configuration option VSI_CACHE_SIZE (in bytes). Content in that cache is discarded when the file handle is closed.
)
# -
# ## Use NASA Common Metadata Repository (CMR) to get Landsat 8 images
#
# [NASA CMR](https://earthdata.nasa.gov/about/science-system-description/eosdis-components/common-metadata-repository) is a new unified way to search for remote sensing assests across many archive centers. If you prefer a graphical user interface, NASA [Earthdata Search](https://search.earthdata.nasa.gov/search) is built on top of CMR. CMR returns download links through the USGS (https://earthexplorer.usgs.gov), but the same archive is mirrored as a (Google Public Dataset)[https://cloud.google.com/storage/docs/public-datasets/landsat], so we'll make a function that queries CMR and returns URLs to the imagery stored on Google Cloud.
def query_cmr_landsat(collection='Landsat_8_OLI_TIRS_C1',tier='T1', path=47, row=27):
"""Query NASA CMR for Collection1, Tier1 Landsat scenes from a specific path and row."""
data = [f'short_name={collection}',
f'page_size=2000',
f'attribute[]=string,CollectionCategory,{tier}',
f'attribute[]=int,WRSPath,{path}',
f'attribute[]=int,WRSRow,{row}',
]
query = 'https://cmr.earthdata.nasa.gov/search/granules.json?' + '&'.join(data)
r = requests.get(query, timeout=100)
print(r.url)
df = pd.DataFrame(r.json()['feed']['entry'])
# Save results to a file
#print('Saved results to cmr-result.json')
#with open('cmr-result.json', 'w') as j:
# j.write(r.text)
return df
def make_google_archive(pids, bands):
"""Turn list of product_ids into pandas dataframe for NDVI analysis."""
path = pids[0].split('_')[2][1:3]
row = pids[0].split('_')[2][-2:]
baseurl = f'https://storage.googleapis.com/gcp-public-data-landsat/LC08/01/0{path}/0{row}'
dates = [pd.to_datetime(x.split('_')[3]) for x in pids]
df = pd.DataFrame(dict(product_id=pids, date=dates))
for band in bands:
df[band] = [f'{baseurl}/{x}/{x}_{band}.TIF' for x in pids]
return df
# Landsat scenes over a specific area are categorized by Path and Row
df = query_cmr_landsat(path=47, row=27)
# +
# Holoviews maps are interactive, you can zoom in with the magnifying glass tool
coordlist = df.polygons.iloc[0]
lats = coordlist[0][0].split(' ')[::2]
lons = coordlist[0][0].split(' ')[1::2]
coords = [(float(lon),float(lat)) for lon, lat in zip(lons,lats)]
poly = shapely.geometry.Polygon(coords)
buffer = 1 #degrees
left, bottom, right, top = poly.bounds
footprint = gv.Shape(poly, label=df.title.iloc[0]).options(alpha=0.5)
tiles = gv.tile_sources.CartoEco.options(width=700, height=500).redim.range(Latitude=(bottom-1, top+1), Longitude=(left-1,right+1))
labels = gv.tile_sources.StamenLabels.options(level='annotation')
tiles * footprint * labels
# -
# Get all scenes for a given path and row, bands 4 and 5
pids = df.title.tolist()
# Don't use the most recent date since there can be a lag in data being on Google Storage
df = make_google_archive(pids[:-1], ['B4', 'B5'])
#df.head()
df.tail()
# ## Launch Dask Kubernetes Cluster
#
# This will allow us to distribute our analysis across many machines. In the default configuration for Pangeo Binder, each worker has 2 vCPUs and 7Gb of RAM. It may take several minutes to initialize these workers and make them available to Dask.
# Select 10 'workers' under 'manual scaling' menu below and click 'Scale'
# Click on the 'Dashboard link' to monitor calculation progress
cluster = KubeCluster(n_workers=10)
cluster
# Attach Dask to the cluster
client = Client(cluster)
# ## Examine a single band Landsat image
#
# The *rasterio* library allows us to read Geotiffs on the web without downloading the entire image. *Xarray* has a built-in load_rasterio() function that allows us to open the file as a DataArray. Xarray also uses Dask for lazy reading, so we want to make sure the native block tiling of the image matches the dask "chunk size". These dask chunks are automatically distributed among all our workers when a computation is requested, so ideally they will fit in the worker memory. A chunk size of 2048x2048 with a float32 datatype implies a 16Mb array.
#
# Load with rasterio
image_url = df.iloc[5]['B4']
with env:
with rasterio.open(image_url) as src:
width = src.width
blockx = src.profile['blockxsize']
blocky = src.profile['blockysize']
print(src.profile)
# Note that the blocksize of the image is 256 by 256, so we want xarray to use some multiple of that
xchunk = int(width/blockx)*blockx
ychunk = blocky
with env:
da = xr.open_rasterio(image_url, chunks={'band': 1, 'x': xchunk, 'y': ychunk})
da
# ### holoviews visualization
# If we request to compute something or plot these arrays, the necessary data chunks will be accessed on cloud storage. Watch the KubeCluster dashboard to see the worker activity when this command is run. Note that no data is stored on the disk here, it's all in memory
#
# Use the magnifying glass button on the right to interactively zoom in. The image resolution automatically updates based on zoom level. The cursor gives you UTM coordinates and the image value at that point!
band1 = da.sel(band=1).persist()
img = band1.hvplot(rasterize=True, dynamic=True, width=700, height=500, cmap='magma')
img
# ## Load all Landsat bands into an xarray dataset
#
# Often we want to analyze a time series of satellite imagery, but we are constrained by computational resources. So we either download all the images, extract a small subset and then do our analysis. Or, we coarsen the resolution of all our images so that the entire set fits into our computer RAM. Because this notebook is running on Google Cloud with access to many resources in our Kube Cluster, we no longer have to worry about the computational constraints, and can conduct our analysis at full resoution!
#
# First we need to construct an xarray dataset object (which has data variables 'band4' and 'band5' in a n-dimensional array with x-coordinates representing UTM easting, y-coordinates representing UTM northing, and a time coordinate representing the image acquisition date).
#
# There are different ways to go about this, but we will load our images with a timestamp index since each image is taken on a different date. Typically, this is a chore if our images are not on the same grid to begin with, but xarray knows how to automatically align images based on their georeferenced coordinates.
# Note that these landsat images are not necessarily the same shape or on the same grid:
for image_url in df.B4[:5]:
with env:
with rasterio.open(image_url) as src:
print(src.shape, src.bounds)
def create_multiband_dataset(row, bands=['B4','B5'], chunks={'band': 1, 'x': xchunk, 'y': ychunk}):
'''A function to load multiple landsat bands into an xarray dataset
'''
# Each image is a dataset containing both band4 and band5
datasets = []
for band in bands:
url = row[band]
with env:
da = xr.open_rasterio(url, chunks=chunks)
da = da.squeeze().drop(labels='band')
ds = da.to_dataset(name=band)
datasets.append(ds)
DS = xr.merge(datasets)
return DS
# Merge all acquisitions into a single large Dataset, this will take a minute
datasets = []
for i,row in df.iterrows():
try:
print('loading...', row.date)
ds = create_multiband_dataset(row)
datasets.append(ds)
except Exception as e:
print('ERROR loading, skipping acquistion!')
print(e)
# Create an xarray dataset
# This takes a while to expand dimensions
DS = xr.concat(datasets, dim=pd.DatetimeIndex(df.date.tolist(), name='time'))
print('Dataset size (Gb): ', DS.nbytes/1e9)
DS
# Set chunks now
#xchunk = int(DS.dims['x']/blockx)*blockx
xchunk = DS.dims['x']
chunks = {'x': xchunk, 'y': ychunk}
DS = DS.chunk(chunks)
DS
# ### Note that xarray has automatically expanded the dimensions to include the maximum extents of all the images, also the chunksize has been automatically adjusted.
#
# There is definitely some room for improvement here from a computational efficiency standpoint - in particular the dask chunks are no longer aligned with the image tiles. This is because each image starts at different coordinates and has different shapes, but xarray uses a single chunk size for the entire datasets. There will also be many zeros in this dataset, so future work could take advantage of sparse arrays.
#
# These points aside, our KubeCluster will automatically parallelize our computations for us, so we can not worry too much about optimal efficiency and just go ahead and run our analysis!
# Again, only metadata is retrieved at this point, wich is why it's so quick!
da = DS.sel(time='2013-04-21')['B4']
print('Image size (Gb): ', da.nbytes/1e9)
da
# ## Distributed NDVI computations
#
# Set up our NDVI dataset. Note that NDVI is not actually computed until we call the Dask compute(), persist(), or call other functions such as plot() that require actually operate on the data!
#
# Because we now have a timeseries of images Holoviews automatically will add a time slider to our visualization! We add a 'SingleTap' features as well to keep track of interesting coordinates (by clicking on the image a white dot will appear and the coordinates are stored in the 'taps' list)
NDVI = (DS['B5'] - DS['B4']) / (DS['B5'] + DS['B4'])
NDVI
taps = []
def record_coords(x, y):
if None not in [x,y]:
taps.append([x, y])
return hv.Points(taps)
# +
# NOTE: this will take a minute to load and is best viewed on a wide monitor
# the time slider can get hidden on small screens
img = NDVI.hvplot('x', 'y', groupby='time', dynamic=True, rasterize=True, width=700, height=500, cmap='magma')
tap = hv.streams.SingleTap(transient=True, source=img)
clicked_points = hv.DynamicMap(record_coords, streams=[tap])
img * clicked_points.options(size=10, color='w')
# +
# Points clicked are stored in the 'taps list'
if len(taps) == 0:
taps = [(562370, 5312519)]
print('Selected points:')
taps
# -
# ### Extract time series for region around point selected from map
#
# This uses a buffer around a selected point and does monthly resampling and will probably take a minute or so to pull the necessary data and run computations
xcen,ycen = taps[0]
buf = 5000 # look at point +/- 5km
ds = NDVI.sel(x=slice(xcen-buf,xcen+buf), y=slice(ycen-buf,ycen+buf))
timeseries = ds.resample(time='1MS').mean().persist()
# Store as pandas series
s = timeseries.to_series()
# +
# Holoviews is also great for interative 2D plots
line = s.hvplot(width=700, height=300, legend=False)
points = s.hvplot.scatter(width=700, height=300, legend=False)
label = f'Mean NDVI: easting={xcen:g} , northing={ycen:g}'
(line * points).relabel(label)
# -
# ### Plot subset of selected region at full resolution
ds.sel(time=slice('2015-01-01', '2015-06-15')).plot.imshow('x', 'y', col='time', col_wrap=4, cmap='magma', vmin=0, vmax=1)
# ## In conclusion
#
# * This notebook demonstrates the power of storing data publically in the Cloud as optimized geotiffs - scientists can conduct scalable analysis without downloading the data to a local machine. Only derived subsets and figures need to be downloaded!
# * We used a crude NDVI calculation, designed to demonstrate the syntax and tools - a proper analysis should take into account cloud masks and other corrections
|
notebooks/0-landsat8-on-gcs.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Arc
// language: javascript
// name: arc
// ---
// # Analysing JSON Data
//
// This example demonstrates how to download and process [JSON](https://en.wikipedia.org/wiki/JSON) datasets using the open source data transformation tool: [Arc](https://arc.tripl.ai/). In this example the data is downloaded directly from the source using an Arc [HTTPExtract](https://arc.tripl.ai/extract/#httpextract) stage but the data could equally have been downloaded prior to the job.
//
// This example aims to show how to:
//
// 1. source data using the [HTTPExtract](https://arc.tripl.ai/extract/#httpextract) stage.
// 1. parse a [JSON](https://en.wikipedia.org/wiki/JSON) response using the [JSONExtract](https://arc.tripl.ai/extract/#jsonextract) stage.
// 1. process the nested result into a standard tabular representation using the [SQLTransform](https://arc.tripl.ai/transform/#sqltransform) stage with the inline `%sql` 'magic' functionality
// 1. ensure data quality and load assurance needs are met using the [SQLValidate](https://arc.tripl.ai/validate/#sqlvalidate) stage.
// 1. write out the data so it can be consumed by other people or jobs.
//
// This example is possible due to data provided by the excellent [Australian Bureau of Meteorology](http://www.bom.gov.au).
// ### Define variables
//
// To make this process reusable we first define a variable which we can use to dynamically replace the weather station identifier at execution time with a different station. This makes this job reusable for all [Australian Bureau of Meteorology](http://www.bom.gov.au) weather stations.
//
// To set a variable to use when developing logic it can be set with the `%env` magic:
// %env
WMO_STATION_ID=94768
// ### Download the weather data
//
// This step uses the [HTTPExtract](https://arc.tripl.ai/extract/#httpextract) stage to directly download the data at `http://www.bom.gov.au/fwo/IDN60901/IDN60901.94768.json` and makes that dataset available with the alias `weather_raw` (defined by `outputView`). The reponse is the raw response data to allow you to choose how to process the data.
//
// Here we are using the `WMO_STATION_ID` variable and [string interpolation](https://en.wikipedia.org/wiki/String_interpolation) to call the endpoint based on the provided station identifier.
// %arc truncate=100
{
"type": "HTTPExtract",
"name": "download weather data",
"environments": [
"production",
"test"
],
"inputURI": "http://www.bom.gov.au/fwo/IDN60901/IDN60901."${WMO_STATION_ID}".json",
"outputView": "weather_raw",
"persist": true
}
// ### Parse the response
//
// This step uses the [JSONExtract](https://arc.tripl.ai/extract/#jsonextract) stage to parse data in the `body` field (defined by `inputField`) of the incoming `weather_raw` dataset (defined by `inputView`) and makes that dataset available with the alias `weather_nested` (defined by `outputView`).
//
// If the `IDN60901.94768.json` file had been downloaded prior to this job (instead of being directly downloaded using the [HTTPExtract](https://arc.tripl.ai/extract/#httpextract) stage) then `inputView` and `inputField` would be replaced with an `inputURI` pointing to that file or multiple files.
{
"type": "JSONExtract",
"name": "parse weather data http response",
"environments": [
"production",
"test"
],
"inputView": "weather_raw",
"inputField": "body",
"outputView": "weather_nested"
}
// ### View the schema
//
// Arc runs on top of [Apache Spark](https://spark.apache.org/) which supports advanced data types such as nested objects inside arrays like returned in the `IDN60901.94768.json` [JSON](https://en.wikipedia.org/wiki/JSON) dataset in addition to the standard `string`, `float` and `date` data types.
//
// To see the schema and help write queries to extract the data the `%printschema` magic can be used to show the layout of the data within the parsed [JSON](https://en.wikipedia.org/wiki/JSON) response.
// %printschema
weather_nested
// ### Explode the data
//
// [Arc](https://arc.tripl.ai/) aims to help business users safely and independently build their own data processing jobs by removing the multiple translations traditionally required when having to hand off the work to developers or other 'go-between' roles.
//
// To do this we need a way of those users expressing business intent for which we employ [SQL](https://en.wikipedia.org/wiki/SQL) as a dialect (which is why we say that [Arc](https://arc.tripl.ai/) is 'SQL First'). We have found that SQL supports most standard data transformation requirements, is relatively easy to learn and easy to hire for.
//
// The statement below demonstrates three key operations for processing the `IDN60901.94768.json` nested dataset:
//
// 1. use of the [POSEXPLODE](https://spark.apache.org/docs/latest/api/sql/index.html#posexplode) SQL function to separates the elements of the `observations.data` into multiple rows. [POSEXPLODE](https://spark.apache.org/docs/latest/api/sql/index.html#posexplode) also returns the `position` of the data (i.e. the index) in the array which can be useful if the position in the array is important. There is also a most simplistic [EXPLODE](https://spark.apache.org/docs/latest/api/sql/index.html#explode) function which does not return the position.
// 1. use of a subquery to turn the data returned by [POSEXPLODE](https://spark.apache.org/docs/latest/api/sql/index.html#posexplode) into a normal tabular representation by selecting the required fields. If desired the use of `observation.*` would also work instead of selecting individual fields.
// 1. parsing the `aifstime_utc` field into a [UTC](https://en.wikipedia.org/wiki/Coordinated_Universal_Time) timestamp which can then be safely used to order the data.
// +
// %sql name="calculate weather" outputView=weather environments=production,test persist=true
SELECT
-- convert aifstime_utc to a timestamp object
TIMESTAMP(
CONCAT(
-- date
SUBSTR(observation.aifstime_utc, 0, 4),'-', SUBSTR(observation.aifstime_utc, 5, 2),'-', SUBSTR(observation.aifstime_utc, 7, 2)
,' ',
-- time
SUBSTR(observation.aifstime_utc, 9, 2),':', SUBSTR(observation.aifstime_utc, 11, 2),':', SUBSTR(observation.aifstime_utc, 13, 2)
)
) AS timestamp
,observation.air_temp
,observation.apparent_t
,observation.delta_t
,observation.dewpt
,observation.press
,observation.wind_spd_kmh
,observation.history_product
,observation.wmo
,header.refresh_message
,_index
FROM (
SELECT
POSEXPLODE(observations.data) AS (_index, observation)
FROM weather_nested
) observations
CROSS JOIN
(
SELECT
EXPLODE(observations.header) AS header
FROM weather_nested
) header
// -
// ### Validate the data / load assurance
//
// Before using the data it is a good idea to ensure that certain data quality guarantees can be provided to downstream consumers of the data. To do this we once again use an [Arc](https://arc.tripl.ai/) 'SQL First' approach to define data quality rules using a [SQLValidate](https://arc.tripl.ai/validate/#sqlvalidate) stage. When building [Arc](https://arc.tripl.ai/) we could have tried to impose a set of standardised business rules but inevitably someone would have a case which was not covered but can be met using a SQL statement.
//
// The statement below demonstrates three key operations for our processed the `IDN60901.94768.json` flattened dataset:
//
// 1. to apply individual rules a good method is to use case statements. For example if we want to ensure that all the timestamp values are populated we can write a statement to find any missing values like: `CASE WHEN timestamp IS NULL THEN 1 ELSE 0 END AS timestamp_null`. `CASE` statements allow very detailed business rules to be defined. Many of these rules can be quickly added for different conditions you care about.
// 1. once the individual rules have been applied we need to define what conditions need to be met for the data to be considered to have met data quality guarantees (and return a `TRUE`/`FALSE` response). In this case we are asserting that the record set must meet these condtions for this stage to be successful:
// - has 144 rows (3 days of 30 minute interval data) - this is Load Assurance to ensure all records have been received and processed.
// - AND the `SUM` of the `timestamp_null` rule must equal `0` - this is a Data Quality 'completentess' guarantee that can be provided to consumers of the data
// - AND the `SUM` of the `air_temp_null` rule must also equal `0` - this is a Data Quality 'completentess' guarantee that can be provided to consumers of the data
//
// 1. additionally we can return a message that is added to the logs. in this case we are returning a [JSON](https://en.wikipedia.org/wiki/JSON) formatted string which looks like `{"count":144,"timestamp_null":0,"air_temp_null":0}`. This is very useful when monitoring this job in when it is operational as we can track metrics or set up alerts when certain conditions occur.
// %sqlvalidate name="validate dataset" environments=production,test
SELECT
COUNT(*) = 144 AND SUM(timestamp_null) = 0 AND SUM(air_temp_null) = 0 AS valid
,TO_JSON(
NAMED_STRUCT(
'count', COUNT(*),
'timestamp_null', SUM(timestamp_null),
'air_temp_null', SUM(air_temp_null)
)
) AS message
FROM (
SELECT
CASE
WHEN timestamp IS NULL THEN 1
ELSE 0
END AS timestamp_null
,CASE
WHEN air_temp IS NULL THEN 1
ELSE 0
END AS air_temp_null
FROM weather
) input_table
// ### Use the data
//
// At this point we can write arbitrary SQL against the `weather` dataset as a standard [SQL](https://en.wikipedia.org/wiki/SQL) table using the `%sql` command:
// %sql
-- find records where the difference between apparent temperature (feels like) and air temperature correlates with wind speed
SELECT
air_temp
,apparent_t
,air_temp - apparent_t AS difference_t
,wind_spd_kmh
FROM weather
WHERE air_temp > apparent_t
// ### Export the data
//
// As we are confident our data is of good quality (due to passing the [SQLValidate](https://arc.tripl.ai/validate/#sqlvalidate) stage) we can export the data so it can be safely consumed by other people and jobs.
//
// To do so we can use the [ParquetLoad](https://arc.tripl.ai/load/#parquetload) stage to write the data out to a [Parquet](https://parquet.apache.org/) format which can then can be easily reloaded without losing any data precision using a [ParquetExtract](https://arc.tripl.ai/extract/#parquetextract) stage.
{
"type": "ParquetLoad",
"name": "write out flattened weather dataset",
"environments": ["production", "test"],
"inputView": "weather",
"outputURI": "/home/jovyan/examples/weather/output/"${WMO_STATION_ID}"/weather.parquet",
"saveMode": "Append"
}
// ### Execute the Job
//
// Now that a job has been built in this notebook it is possible to execute it using the [Arc](https://arc.tripl.ai/) Docker image. Notice that we have defined the `WMO_STATION_ID` parameter using an environment variable which can be easily changed to a different station. This could be scheduled to run periodically to retrieve this data.
//
// ```bash
// docker run \
// --rm \
// -v $(pwd)/examples:/home/jovyan/examples:Z \
// -e "ETL_CONF_ENV=production" \
// -e "WMO_STATION_ID=94768" \
// -p 4040:4040 \
// triplai/arc:arc_2.8.0_spark_2.4.5_scala_2.12_hadoop_2.9.2_1.0.0 \
// bin/spark-submit \
// --master local[*] \
// --driver-memory 4g \
// --class ai.tripl.arc.ARC \
// /opt/spark/jars/arc.jar \
// --etl.config.uri=file:///home/jovyan/examples/weather/ProcessJSON.ipynb
// ```
|
examples/weather/ProcessJSON.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd # data package
import matplotlib.pyplot as plt # graphics
import datetime as dt
import numpy as np
from census import Census # This is new...
import requests, io # internet and input tools
import zipfile as zf # zip file tools
import os
#import weightedcalcs as wc
#import numpy as np
import pyarrow as pa
import pyarrow.parquet as pq
# +
#https://www.ons.gov.uk/economy/nationalaccounts/balanceofpayments/datasets/uktradecountrybycommodityimports
url = "https://www.ons.gov.uk/file?uri=%2feconomy%2fnationalaccounts%2fbalanceofpayments%2fdatasets%2fuktradecountrybycommodityimports%2fcurrent/cxcimportshistoricaldata.zip"
# +
r = requests.get(url)
brexit_FEB21 = zf.ZipFile(io.BytesIO(r.content))
brexit_FEB21.extractall(".\\data")
filename = brexit_FEB21.namelist()
# -
df = pd.read_excel(".\\data" + "\\" + filename[0], sheet_name= "Country by Commodity Import")
df.head()
# +
df = df.melt(id_vars = ["COMMODITY", "COUNTRY", "DIRECTION"])
df.drop("DIRECTION", axis = 1, inplace = True)
df.variable= pd.to_datetime(df.variable, format="%Y%b")
df.rename({"variable":"time", "value": "imports"},inplace = True, axis = 1)
df.COUNTRY = df.COUNTRY.str[3:]
df.imports = df.imports*1000000
# +
grp = df[df.COMMODITY == 'T Total'].groupby("COUNTRY")
test = grp.agg({"imports": "sum"})
# -
topcntry = test.sort_values(by = "imports", ascending = False).iloc[0:23].index.to_list()
df = df[df.COUNTRY.isin(topcntry)]
df.COMMODITY.replace({"T Total": "Total"}, inplace = True)
df.COUNTRY.replace({"United States inc Puerto Rico": "United States"}, inplace = True)
df.COUNTRY.replace({"Extra EU 28 (Rest of World)": "Excl. EU 28 (Rest of World)"}, inplace = True)
grp = df.groupby(["COUNTRY"])
def ex_metal_total(df):
if not (df[df["COMMODITY"] == "9 Unspecified goods"].empty):
foo = pd.DataFrame([])
fooall = df[df["COMMODITY"] == "Total"].set_index(["time"])["imports"]
foobar = df[df["COMMODITY"] == "9 Unspecified goods"].set_index(["time"])["imports"]
foo["imports"] = fooall - foobar
# Grab the total and then sbutract off the unspecified goods
foo["COUNTRY"] = df.COUNTRY.unique()[0]
#print(df.COUNTRY.unique())
foo["COMMODITY"] = "Total (ex. metals)"
foo.reset_index(inplace = True)
foo[df.columns]
df = df.append(foo)
return df
df = grp.apply(ex_metal_total)
# +
df.drop(["COUNTRY"], axis = 1, inplace = True)
df.reset_index(inplace = True)
df.drop(["level_1"], axis = 1, inplace = True)
# +
out_file = ".\\data"+ "\\UK-imports-1997-2017.parquet"
pq.write_table(pa.Table.from_pandas(df), out_file)
# -
df.tail()
|
brexit-data-1997-2017.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Modelling lithium plating in PyBaMM
#
# This notebook shows how PyBaMM [7] can be used to model both reversible and irreversible lithium plating.
# %pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import os
import numpy as np
import matplotlib.pyplot as plt
os.chdir(pybamm.__path__[0]+'/..')
# The Doyle-Fuller-Newman model [3] is upgraded with two different lithium plating models. Model 1 contains the reversible lithium plating model of O'Kane et al. [5], while model 2 contains the same model but with the lithium stripping capability removed, making the plating irreversible. The parameters are taken from Chen et al.'s investigation [2] of an LG M50 cell.
# +
# choose models
model1 = pybamm.lithium_ion.DFN(options={"lithium plating": "reversible"})
model2 = pybamm.lithium_ion.DFN(options={"lithium plating": "irreversible"})
# pick parameters
parameter_values = pybamm.ParameterValues("Chen2020_plating")
#parameter_values.update({"Reference temperature [K]": 268.15})
parameter_values.update({"Ambient temperature [K]": 268.15})
#parameter_values.update({"Initial temperature [K]": 268.15})
parameter_values.update({"Upper voltage cut-off [V]": 4.21})
parameter_values.update({"Lithium plating kinetic rate constant [m.s-1]": 1E-9})
# -
# A series of simple fast charging experiments based on those of Ren et al. [6] is defined here. We first initialise the model at 0% SoC by performing a C/20 discharge (see more details on how to initialise a model from a simulation in [this notebook](./initialise-model-with-solution.ipynb)).
# +
# specify experiments
pybamm.citations.register("Ren2018")
experiment_discharge = pybamm.Experiment(
[
("Discharge at C/20 until 2.5 V (10 minute period)",
"Rest for 1 hour (3 minute period)")
]
)
sim_discharge1 = pybamm.Simulation(model1, parameter_values=parameter_values, experiment=experiment_discharge)
sol_discharge1 = sim_discharge1.solve()
model1.set_initial_conditions_from(sol_discharge1, inplace=True)
sim_discharge2 = pybamm.Simulation(model2, parameter_values=parameter_values, experiment=experiment_discharge)
sol_discharge2 = sim_discharge2.solve()
model2.set_initial_conditions_from(sol_discharge2, inplace=True)
# -
# And we can now define the different experiments to charge at different C-rates.
experiment_2C = pybamm.Experiment(
[
("Charge at 2C until 4.2 V",
"Hold at 4.2 V until C/20",
"Rest for 1 hour")
]
)
experiment_1C = pybamm.Experiment(
[
("Charge at 1C until 4.2 V",
"Hold at 4.2 V until C/20",
"Rest for 1 hour")
]
)
experiment_Cover2 = pybamm.Experiment(
[
("Charge at C/2 until 4.2 V",
"Hold at 4.2 V until C/20",
"Rest for 1 hour")
]
)
experiment_Cover4 = pybamm.Experiment(
[
("Charge at C/4 until 4.2 V",
"Hold at 4.2 V until C/20",
"Rest for 1 hour")
]
)
experiment_Cover8 = pybamm.Experiment(
[
("Charge at C/8 until 4.2 V",
"Hold at 4.2 V until C/20",
"Rest for 1 hour")
]
)
# Solve the reversible plating model first. The default CasADi [1] solver is used here.
sim1_2C = pybamm.Simulation(model1, experiment=experiment_2C, parameter_values=parameter_values)
sim1_2C.solve()
sim1_1C = pybamm.Simulation(model1, experiment=experiment_1C, parameter_values=parameter_values)
sim1_1C.solve()
sim1_Cover2 = pybamm.Simulation(model1, experiment=experiment_Cover2, parameter_values=parameter_values)
sim1_Cover2.solve()
sim1_Cover4 = pybamm.Simulation(model1, experiment=experiment_Cover4, parameter_values=parameter_values)
sim1_Cover4.solve()
sim1_Cover8 = pybamm.Simulation(model1, experiment=experiment_Cover8, parameter_values=parameter_values)
sim1_Cover8.solve()
A = parameter_values.evaluate(model1.param.L_y * model1.param.L_z)
F = parameter_values.evaluate(model1.param.F)
L_n = parameter_values.evaluate(model1.param.L_n)
# Isolate final equilibration phase
sol_2C_rest = sim1_2C.solution.cycles[0].steps[2]
sol_1C_rest = sim1_1C.solution.cycles[0].steps[2]
sol_Cover2_rest = sim1_Cover2.solution.cycles[0].steps[2]
sol_Cover4_rest = sim1_Cover4.solution.cycles[0].steps[2]
sol_Cover8_rest = sim1_Cover8.solution.cycles[0].steps[2]
# Time (hours)
t_2C = sol_2C_rest["Time [s]"].entries-sol_2C_rest["Time [s]"].entries[0]
t_1C = sol_1C_rest["Time [s]"].entries-sol_1C_rest["Time [s]"].entries[0]
t_Cover2 = sol_Cover2_rest["Time [s]"].entries-sol_Cover2_rest["Time [s]"].entries[0]
t_Cover4 = sol_Cover4_rest["Time [s]"].entries-sol_Cover4_rest["Time [s]"].entries[0]
t_Cover8 = sol_Cover8_rest["Time [s]"].entries-sol_Cover8_rest["Time [s]"].entries[0]
# Intercalated capacity (Ah)
Q_main_2C = sol_2C_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_1C = sol_1C_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_Cover2 = sol_Cover2_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_Cover4 = sol_Cover4_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_Cover8 = sol_Cover8_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
# Plated capacity (Ah)
Q_Li_2C = sol_2C_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_1C = sol_1C_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_Cover2 = sol_Cover2_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_Cover4 = sol_Cover4_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_Cover8 = sol_Cover8_rest["Loss of capacity to lithium plating [A.h]"].entries
# Voltage (V)
V_2C = sol_2C_rest["Terminal voltage [V]"].entries
V_1C = sol_1C_rest["Terminal voltage [V]"].entries
V_Cover2 = sol_Cover2_rest["Terminal voltage [V]"].entries
V_Cover4 = sol_Cover4_rest["Terminal voltage [V]"].entries
V_Cover8 = sol_Cover8_rest["Terminal voltage [V]"].entries
# Dimensionless current components
j_2C = sol_2C_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_2C = sol_2C_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_2C = sol_2C_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_1C = sol_1C_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_1C = sol_1C_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_1C = sol_1C_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_Cover2 = sol_Cover2_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_Cover2 = sol_Cover2_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_Cover2 = sol_Cover2_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_Cover4 = sol_Cover4_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_Cover4 = sol_Cover4_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_Cover4 = sol_Cover4_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_Cover8 = sol_Cover8_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_Cover8 = sol_Cover8_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_Cover8 = sol_Cover8_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
fig, axs = plt.subplots(2, 2, figsize=(13,9))
axs[0,0].plot(t_2C/60, V_2C, color='tab:purple', linestyle='solid')
axs[0,0].plot(t_1C/60, V_1C, color='tab:cyan', linestyle='solid')
axs[0,0].plot(t_Cover2/60, V_Cover2, color='tab:red', linestyle='solid')
axs[0,0].plot(t_Cover4/60, V_Cover4, color='tab:green', linestyle='solid')
axs[0,0].plot(t_Cover8/60, V_Cover8, color='tab:blue', linestyle='solid')
axs[0,0].set_xlabel("Time [minutes]")
axs[0,0].set_ylabel("Voltage [V]")
axs[0,0].legend(('2C','1C','C/2','C/4','C/8'))
axs[0,1].plot(t_2C/60, j_2C, color='tab:purple', linestyle='dashed')
axs[0,1].plot(t_2C/60, j_sr_2C, color='tab:purple', linestyle='dotted')
axs[0,1].plot(t_2C/60, j_sum_2C, color='tab:purple', linestyle='solid')
axs[0,1].plot(t_1C/60, j_1C, color='tab:cyan', linestyle='dashed')
axs[0,1].plot(t_1C/60, j_sr_1C, color='tab:cyan', linestyle='dotted')
axs[0,1].plot(t_1C/60, j_sum_1C, color='tab:cyan', linestyle='solid')
axs[0,1].plot(t_Cover2/60, j_Cover2, color='tab:red', linestyle='dashed')
axs[0,1].plot(t_Cover2/60, j_sr_Cover2, color='tab:red', linestyle='dotted')
axs[0,1].plot(t_Cover2/60, j_sum_Cover2, color='tab:red', linestyle='solid')
axs[0,1].plot(t_Cover4/60, j_Cover4, color='tab:green', linestyle='dashed')
axs[0,1].plot(t_Cover4/60, j_sr_Cover4, color='tab:green', linestyle='dotted')
axs[0,1].plot(t_Cover4/60, j_sum_Cover4, color='tab:green', linestyle='solid')
axs[0,1].plot(t_Cover8/60, j_Cover8, color='tab:blue', linestyle='dashed')
axs[0,1].plot(t_Cover8/60, j_sr_Cover8, color='tab:blue', linestyle='dotted')
axs[0,1].plot(t_Cover8/60, j_sum_Cover8, color='tab:blue', linestyle='solid')
axs[0,1].set_xlabel("Time [minutes]")
axs[0,1].set_ylabel("Interfacial current density [dimensionless]")
axs[0,1].legend(('Deintercalation current','Stripping current','Total current'))
axs[1,0].plot(t_2C/60, Q_Li_2C, color='tab:purple', linestyle='solid')
axs[1,0].plot(t_1C/60, Q_Li_1C, color='tab:cyan', linestyle='solid')
axs[1,0].plot(t_Cover2/60, Q_Li_Cover2, color='tab:red', linestyle='solid')
axs[1,0].plot(t_Cover4/60, Q_Li_Cover4, color='tab:green', linestyle='solid')
axs[1,0].plot(t_Cover8/60, Q_Li_Cover8, color='tab:blue', linestyle='solid')
axs[1,0].set_xlabel("Time [minutes]")
axs[1,0].set_ylabel("Plated lithium capacity [Ah]")
axs[1,1].plot(t_2C/60, Q_main_2C, color='tab:purple', linestyle='solid')
axs[1,1].plot(t_1C/60, Q_main_1C, color='tab:cyan', linestyle='solid')
axs[1,1].plot(t_Cover2/60, Q_main_Cover2, color='tab:red', linestyle='solid')
axs[1,1].plot(t_Cover4/60, Q_main_Cover4, color='tab:green', linestyle='solid')
axs[1,1].plot(t_Cover8/60, Q_main_Cover8, color='tab:blue', linestyle='solid')
axs[1,1].set_xlabel("Time [minutes]")
axs[1,1].set_ylabel("Intercalated lithium capacity [Ah]")
plt.tight_layout()
plt.show()
# The results show both similarities and differences with those of Ren et al. [6]. Notably, unlike Ren et al., this model uses equations [5] that result in a small but finite amount of plated lithium being present in the steady state.
# Now solve the irreversible plating model and see how it compares.
sim2_2C = pybamm.Simulation(model2, experiment=experiment_2C, parameter_values=parameter_values)
sim2_2C.solve()
sim2_1C = pybamm.Simulation(model2, experiment=experiment_1C, parameter_values=parameter_values)
sim2_1C.solve()
sim2_Cover2 = pybamm.Simulation(model2, experiment=experiment_Cover2, parameter_values=parameter_values)
sim2_Cover2.solve()
sim2_Cover4 = pybamm.Simulation(model2, experiment=experiment_Cover4, parameter_values=parameter_values)
sim2_Cover4.solve()
sim2_Cover8 = pybamm.Simulation(model2, experiment=experiment_Cover8, parameter_values=parameter_values)
sim2_Cover8.solve()
# WARNING: RUNNING THIS CELL WILL OVERWRITE PREVIOUS VARIABLES (but not the raw solution data)
A = parameter_values.evaluate(model2.param.L_y * model2.param.L_z)
F = parameter_values.evaluate(model2.param.F)
L_n = parameter_values.evaluate(model2.param.L_n)
# Isolate final equilibration phase
sol_2C_rest = sim2_2C.solution.cycles[0].steps[2]
sol_1C_rest = sim2_1C.solution.cycles[0].steps[2]
sol_Cover2_rest = sim2_Cover2.solution.cycles[0].steps[2]
sol_Cover4_rest = sim2_Cover4.solution.cycles[0].steps[2]
sol_Cover8_rest = sim2_Cover8.solution.cycles[0].steps[2]
# Time (hours)
t_2C = sol_2C_rest["Time [s]"].entries-sol_2C_rest["Time [s]"].entries[0]
t_1C = sol_1C_rest["Time [s]"].entries-sol_1C_rest["Time [s]"].entries[0]
t_Cover2 = sol_Cover2_rest["Time [s]"].entries-sol_Cover2_rest["Time [s]"].entries[0]
t_Cover4 = sol_Cover4_rest["Time [s]"].entries-sol_Cover4_rest["Time [s]"].entries[0]
t_Cover8 = sol_Cover8_rest["Time [s]"].entries-sol_Cover8_rest["Time [s]"].entries[0]
# Intercalated capacity (Ah)
Q_main_2C = sol_2C_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_1C = sol_1C_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_Cover2 = sol_Cover2_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_Cover4 = sol_Cover4_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
Q_main_Cover8 = sol_Cover8_rest["Negative electrode volume-averaged concentration [mol.m-3]"].entries * F * A * L_n / 3600
# Plated capacity (Ah)
Q_Li_2C = sol_2C_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_1C = sol_1C_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_Cover2 = sol_Cover2_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_Cover4 = sol_Cover4_rest["Loss of capacity to lithium plating [A.h]"].entries
Q_Li_Cover8 = sol_Cover8_rest["Loss of capacity to lithium plating [A.h]"].entries
# Voltage (V)
V_2C = sol_2C_rest["Terminal voltage [V]"].entries
V_1C = sol_1C_rest["Terminal voltage [V]"].entries
V_Cover2 = sol_Cover2_rest["Terminal voltage [V]"].entries
V_Cover4 = sol_Cover4_rest["Terminal voltage [V]"].entries
V_Cover8 = sol_Cover8_rest["Terminal voltage [V]"].entries
# Dimensionless current components
j_2C = sol_2C_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_2C = sol_2C_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_2C = sol_2C_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_1C = sol_1C_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_1C = sol_1C_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_1C = sol_1C_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_Cover2 = sol_Cover2_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_Cover2 = sol_Cover2_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_Cover2 = sol_Cover2_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_Cover4 = sol_Cover4_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_Cover4 = sol_Cover4_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_Cover4 = sol_Cover4_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
j_Cover8 = sol_Cover8_rest["X-averaged negative electrode interfacial current density"].entries
j_sr_Cover8 = sol_Cover8_rest["X-averaged lithium plating interfacial current density"].entries
j_sum_Cover8 = sol_Cover8_rest["Sum of x-averaged negative electrode interfacial current densities"].entries
fig, axs = plt.subplots(2, 2, figsize=(13,9))
axs[0,0].plot(t_2C/60, V_2C, color='tab:purple', linestyle='solid')
axs[0,0].plot(t_1C/60, V_1C, color='tab:cyan', linestyle='solid')
axs[0,0].plot(t_Cover2/60, V_Cover2, color='tab:red', linestyle='solid')
axs[0,0].plot(t_Cover4/60, V_Cover4, color='tab:green', linestyle='solid')
axs[0,0].plot(t_Cover8/60, V_Cover8, color='tab:blue', linestyle='solid')
axs[0,0].set_xlabel("Time [minutes]")
axs[0,0].set_ylabel("Voltage [V]")
axs[0,0].legend(('2C','1C','C/2','C/4','C/8'))
axs[0,1].plot(t_2C/60, j_2C, color='tab:purple', linestyle='dashed')
axs[0,1].plot(t_2C/60, j_sr_2C, color='tab:purple', linestyle='dotted')
axs[0,1].plot(t_2C/60, j_sum_2C, color='tab:purple', linestyle='solid')
axs[0,1].plot(t_1C/60, j_1C, color='tab:cyan', linestyle='dashed')
axs[0,1].plot(t_1C/60, j_sr_1C, color='tab:cyan', linestyle='dotted')
axs[0,1].plot(t_1C/60, j_sum_1C, color='tab:cyan', linestyle='solid')
axs[0,1].plot(t_Cover2/60, j_Cover2, color='tab:red', linestyle='dashed')
axs[0,1].plot(t_Cover2/60, j_sr_Cover2, color='tab:red', linestyle='dotted')
axs[0,1].plot(t_Cover2/60, j_sum_Cover2, color='tab:red', linestyle='solid')
axs[0,1].plot(t_Cover4/60, j_Cover4, color='tab:green', linestyle='dashed')
axs[0,1].plot(t_Cover4/60, j_sr_Cover4, color='tab:green', linestyle='dotted')
axs[0,1].plot(t_Cover4/60, j_sum_Cover4, color='tab:green', linestyle='solid')
axs[0,1].plot(t_Cover8/60, j_Cover8, color='tab:blue', linestyle='dashed')
axs[0,1].plot(t_Cover8/60, j_sr_Cover8, color='tab:blue', linestyle='dotted')
axs[0,1].plot(t_Cover8/60, j_sum_Cover8, color='tab:blue', linestyle='solid')
axs[0,1].set_xlabel("Time [minutes]")
axs[0,1].set_ylabel("Interfacial current density [dimensionless]")
axs[0,1].legend(('Deintercalation current','Stripping current','Total current'))
axs[1,0].plot(t_2C/60, Q_Li_2C, color='tab:purple', linestyle='solid')
axs[1,0].plot(t_1C/60, Q_Li_1C, color='tab:cyan', linestyle='solid')
axs[1,0].plot(t_Cover2/60, Q_Li_Cover2, color='tab:red', linestyle='solid')
axs[1,0].plot(t_Cover4/60, Q_Li_Cover4, color='tab:green', linestyle='solid')
axs[1,0].plot(t_Cover8/60, Q_Li_Cover8, color='tab:blue', linestyle='solid')
axs[1,0].set_xlabel("Time [minutes]")
axs[1,0].set_ylabel("Plated lithium capacity [Ah]")
axs[1,1].plot(t_2C/60, Q_main_2C, color='tab:purple', linestyle='solid')
axs[1,1].plot(t_1C/60, Q_main_1C, color='tab:cyan', linestyle='solid')
axs[1,1].plot(t_Cover2/60, Q_main_Cover2, color='tab:red', linestyle='solid')
axs[1,1].plot(t_Cover4/60, Q_main_Cover4, color='tab:green', linestyle='solid')
axs[1,1].plot(t_Cover8/60, Q_main_Cover8, color='tab:blue', linestyle='solid')
axs[1,1].set_xlabel("Time [minutes]")
axs[1,1].set_ylabel("Intercalated lithium capacity [Ah]")
plt.tight_layout()
plt.show()
# Unlike in the reversible case, there is no steady state and the capacity degrades over time. Perhaps not leaving the battery at 100% charge would reduce the rate of degradation...
# # References
pybamm.print_citations()
|
examples/notebooks/models/lithium-plating.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pylab import *
import numpy as np
import matplotlib, pylab
from dcll.npamlib import plotLIF, sigmoid, spikes_to_evlist
from dcll.pytorch_libdcll import *
from dcll.load_dvsgestures_sparse import *
matplotlib.rcParams['text.usetex']=False
matplotlib.rcParams['savefig.dpi']=400.
matplotlib.rcParams['font.size']=14.0
matplotlib.rcParams['figure.figsize']=(5.0,3.5)
matplotlib.rcParams['axes.formatter.limits']=[-10,10]
matplotlib.rcParams['axes.labelsize']= 14.
matplotlib.rcParams['figure.subplot.bottom'] = .2
matplotlib.rcParams['figure.subplot.left'] = .2
# -
Nlayers = 3
directory = "../Paper_results/synthetic/"
do_ = [np.load(directory+'do1.npy'), np.load(directory+'do2.npy'), np.load(directory+'do3.npy')]
loss = np.load(directory+'loss.npy')
dwt = np.array(np.load(directory+'dwt.npy'))
deps = np.array(np.load(directory+'deps.npy')).squeeze()
target = np.load(directory+'target.npy')
layer=2
nrn=2
innrn=3
# +
sinput = np.array([d[0] for d in do_[layer-1]]).squeeze()
soutput = np.array([d[0] for d in do_[layer]]).squeeze()
softvoutput = np.array([d[2] for d in do_[layer]]).squeeze()
pvoutput= np.array([d[1] for d in do_[layer]]).squeeze()
pvoutput0= np.array([d[1] for d in do_[0]]).squeeze()
pvoutput1= np.array([d[1] for d in do_[1]]).squeeze()
pvoutput2= np.array([d[1] for d in do_[2]]).squeeze()
voutput = np.array([d[3] for d in do_[layer]]).squeeze()
loss_iter = np.array([d[3] for d in do_[layer]]).squeeze()
target = np.array([d.squeeze() for d in target])
# -
figure(figsize=(5,20))
for nrn_ in range(10):
subplot(10,1,nrn_+1)
plot(torch.nn.Sigmoid()(torch.Tensor(voutput[:,nrn_])).numpy(), linewidth=2, alpha=.8)
plot(softvoutput[:,nrn_], linewidth=2, alpha=.8)
xticks([])
for t in np.where(soutput[:, nrn]==1)[0]:
axvline(t, color='k', alpha=.1)
# +
fig = plt.figure(figsize=(8,9))
k=1
xmin=50
xmax=500
subplot(6,1,k)
t,n = spikes_to_evlist(sinput[:,innrn+1:innrn+10])
plot(t, n+1, 'k|', markersize=6)
t,n = spikes_to_evlist(sinput[:,innrn:innrn+1])
plot(t, n, 'r|', markersize=6)
xlim(xmin,xmax)
ylim([-.5,10])
yticks([])
ylabel('$s_{in}(t)$')
k+=1
subplot(6,1,k)
for tt in t:
axvline(tt, color='r', alpha=.5)
plot(deps[:,innrn], linewidth=2, alpha=.8)
xlim(xmin,xmax)
yticks([])
ylabel('$\epsilon \\ast s_{in}(t)$')
k+=1
subplot(6,1,k)
plot(voutput[:,nrn], linewidth=2, alpha=.8)
for t in np.where(soutput[:, nrn]==1)[0]:
axvline(t, color='k', alpha=.1)
xlim(xmin,xmax)
yticks([])
ylabel('$u(t)$')
k+=1
subplot(6,1,k)
x = voutput[:, nrn]
plot(sigmoid(x)*(1-sigmoid(x)))
xlim(xmin,xmax)
ylim(0,.5)
#yticks([0,.5],[0,.5])
ylabel('$\sigma\'(u)$')
yticks([])
k+=1
ax1=subplot(6,1,k)
#Get rid of zeros due to dropout
ax1.plot(pvoutput2, color='C0')
ax1.plot(target[2], color='C0', linewidth = 3, alpha=.5)
# ax1.plot(pvoutput1, color='C1')
# ax1.plot(target[1], color='C1', linewidth = 3, alpha=.5)
# ax1.plot(pvoutput2, color='C2')
# ax1.plot(target[2], color='C2', linewidth = 3, alpha=.5)
yticks([])
ylabel('$Output$')
xlim(xmin,xmax)
k+=1
subplot(6,1,k)
plot(dwt[:,nrn,innrn])
xlim(xmin,xmax)
yticks([])
ylabel('$\Delta w$')
k+=1
xlabel('Time [au]')
tight_layout()
savefig(directory+'/snapshot.png', dpi=200)
# +
fig = plt.figure(figsize=(4,6))
ax1 = subplot(3,1,1)
ax1.plot(pvoutput2, color='C0')
ax1t = ax1.twinx()
ax1t.plot(target[2], color='C0', linewidth = 3, alpha=.5)
ax1.set_xticks([])
ax1.set_xlim(xmin,xmax)
ax1.set_ylabel('$y_3$', color='C0')
ax1.set_title('Layer 3')
ax1.set_yticks([])
ax1t.set_yticks([])
ax1t.set_ylabel('$\hat{y}_3$', color='C0', alpha=.6)
ax2 = subplot(3,1,2)
ax2.plot(pvoutput1, color='C1')
ax2t = ax2.twinx()
ax2t.plot(target[1], color='C1', linewidth = 3, alpha=.5)
ax2.set_xticks([])
ax2.set_xlim(xmin,xmax)
ax2.set_ylabel('$y_2$', color='C1')
ax2.set_yticks([])
ax2t.set_yticks([])
ax2t.set_ylabel('$\hat{y}_2$', color='C1', alpha=.6)
ax2.set_title('Layer 2')
ax3 = subplot(3,1,3)
ax3.plot(pvoutput0, color='C2')
ax3t = ax3.twinx()
ax3t.plot(target[0], color='C2', linewidth = 3, alpha=.5)
title('Layer 1')
xlim(xmin,xmax)
ax3.set_ylabel('$y_1$', color='C2')
ax3.set_yticks([])
ax3t.set_yticks([])
ax3t.set_ylabel('$\hat{y}_1$', color='C2', alpha=.6)
ax3.set_xlabel('Time [au]')
# Plot 1:
# just align the last column of axes:
fig.align_ylabels()
plt.tight_layout()
savefig(directory+'/layer_outs.png', dpi=200)
# -
|
notebooks/plot_synthetic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#自作moduleのパスを追加。
import sys
sys.path.append('../my-python-module/MyModule')
#自作moduleのインポート
import DataAnalysis
import importlib
importlib.reload(DataAnalysis)
#Html作成用
from jinja2 import Environment, FileSystemLoader
#データベースを読み込む
filepath='./input/FEI_PREF_210502090318.csv'
df=DataAnalysis.readcsv(filepath,6)
#print(df)
#テンプレートファイルを読み込む
env = Environment(loader=FileSystemLoader('./input/', encoding='utf8'))
tmpl = env.get_template('sample_template.html')
#ここで shop に入る文字を指定している
data = {'name_title': '部署の問題', 'item_a': 'Python','value_a':'3.9.4','PdfFilePath':'./pdffiles/sample.pdf'}
html = tmpl.render(data)
with open('./output/jinja2_test.html',mode='w', encoding='utf-8') as f:
f.write(str(html))
print(html)
# -
|
CreateHtmlFile/CreateHtml.ipynb
|