code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1-2.2 Intro Python
# ## Strings: input, testing, formatting
# - input() - gathering user input
# - **print() formatting**
# - Quotes inside strings
# - Boolean string tests methods
# - String formatting methods
# - Formatting string input()
# - Boolean `in` keyword
#
# -----
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - gather, store and use string `input()`
# - **format `print()` output**
# - test string characteristics
# - format string output
# - search for a string in a string
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## comma print formatting
# ### print() comma separated strings
# Python provides several methods of formatting strings in the **`print()`** function beyond **string addition**
#
# **`print()`** provides using **commas** to combine stings for output
# by comma separating strings **`print()`** will output each separated by a space by default
#
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/ac97f001-e639-494e-aa15-e420efb5a7a8/Unit1_Section2-2-Print-Comma_Format.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/ac97f001-e639-494e-aa15-e420efb5a7a8/Unit1_Section2-2-Print-Comma_Format.vtt","srclang":"en","kind":"subtitles","label":"english"}])
#
# #### comma formatted `print()`
# - **[ ]** print 3 strings on the same line using commas inside the `print()` function
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# +
# review and run code
name = "Collette"
# string addition
print("Hello " + name + "!")
# comma separation formatting
print("Hello to",name,"who is from the city")
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
# **print 3 strings on the same line using commas inside the print() function**
# +
# [ ] print 3 strings on the same line using commas inside the print() function
print ("Hi,","it's been a long", "day!")
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## using commas in `print()` with strings and numbers together
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/46b436d7-31ed-4e9a-a4c9-55f9eaacfb84/Unit1_Section2-2-Print-String_Number.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/46b436d7-31ed-4e9a-a4c9-55f9eaacfb84/Unit1_Section2-2-Print-String_Number.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# - **`print()`** function formatting with comma separation works different than with string addition.
# - **`print()`** using comma separation can mix numbers (int & float) and strings without a TypeError
#
# #### `print()` with numbers and strings together using commas
# - **[ ]** use a **`print()`** function with comma separation to combine 2 numbers and 2 strings
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# review and run code
print("I will pick you up @",6,"for the party")
# review and run code
number_errors = 0
print("An Integer of", 14, "combined with strings causes",number_errors,"TypeErrors in comma formatted print!")
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
# **use a print() function with comma separation to combine 2 numbers and 2 strings**
# +
# [ ] use a print() function with comma separation to combine 2 numbers and 2 strings
print ("my wife turned", 50, "this past Sunday, Spet.", 6)
# -
# <font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
# **print() comma separated mixing of strings and variables**
# by comma separating strings and/or string variables **`print()`** will output each separated by a space by default
#
# **display text describing an address, made from stings and variables of different types**
# - initialize variables with input()
# - street
# - st_number
# - Display a message about the street and street number using comma separation formatting
# +
# [ ] get user input for a street name in the variable, street
street = input ("enter street name")
# [ ] get user input for a street number in the variable, st_number
st_number = input ("enter street number")
# [ ] display a message about the street and st_number
print (st_number, street)
# -
# <font size="6" color="#B24C00" face="verdana"> <B>Task 4</B></font>
# **`print()` number, strings, variables from input**
# - [ ] display text made from combining a variable, a literal string and a number
# +
# [ ] define a variable with a string or numeric value
first_house = 322
st_name = "T Street"
second_house = 326
# [ ] display a message combining the variable, 1 or more literal strings and a number
print ("my fraternity was founded at", first_house, st_name, "then moved to", second_house, st_name)
# -
# <font size="6" color="#B24C00" face="verdana"> <B>Task 5</B></font>
# ## Program: How many for the training?
# Create a program that prints out a reservation for a training class. Gather the name of the party, the number attending and the time.
# >**example** of what input/output might look like:
# ```
# enter name for contact person for training group: <NAME>
# enter the total number attending the course: 7
# enter the training time selected: 3:25 PM
# ------------------------------
# Reminder: training is schedule at 3:25 PM for the <NAME> group of 7 attendees
# Please arrive 10 minutes early for the first class
# ```
#
# Design and Create your own reminder style
# - **[ ]** get user input for variables:
# - **owner**: name of person the reservation is for
# - **num_people**: how many are attending
# - **training_time**: class time
# - **[ ]** create an integer variable **min_early**: number of minutes early the party should arrive
# - **[ ]** using comma separation, print reminder text
# - use all of the variables in the text
# - use additional strings as needed
# - use multiple print statements to format message on multiple lines (optional)
# +
# [ ] get input for variables: owner, num_people, training_time - use descriptive prompt text
owner = input ("enter name of person or group")
num_people = input ("enter number of people")
training_time = input ("enter training time")
# [ ] create a integer variable min_early and "hard code" the integer value (e.g. - 5, 10 or 15)
min_early = "10 minutes"
# [ ] print reminder text using all variables & add additional strings - use comma separated print formatting
print ('Reminder: training is scheduled at', training_time, "for the", owner, "group of", num_people, "attendees.",
"Please arrive", min_early, "early for the first class")
# -
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
|
Python Absolute Beginner/Module_1_2.2_Absolute_Beginner.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Veri Görselleştirme Kütüphanesi - Matplotlib
#
# MATLAB-tarzı grafikler çizdirmek için Python'a eklenmiş bir kütüphane olan matplotlib ile kendi grafiklerimizi çizmeye çalışacağız.
# +
# gerekli kütüphaneler
import matplotlib.pyplot as plt # Grafik çizdirme kütüphanesi
import pandas as pd
import numpy as np
veriler = pd.read_csv("heart_disease.csv")
# -
plt.boxplot(x=veriler['max_heart_rate'])
veriler.info()
# "max_heart_rate" nümerik olarak tanımlanmadığı için grafiğini çizemedik.
# Önce '?' olarak girilen değerleri değiştirmemiz lazım
veriler["max_heart_rate"].unique()
veriler["max_heart_rate"].replace(to_replace="?",value=0, inplace=True) # inplace = True yazmazsak değiştirmez
veriler["max_heart_rate"].unique()
# şimdi dönüşümü yapalım
veriler["max_heart_rate"]=veriler["max_heart_rate"].astype(np.int64)
veriler["max_heart_rate"].describe()
# boxplot figure
plt.boxplot(x=veriler["max_heart_rate"]);
# görünümü değiştirme
plt.style.use('ggplot')
# plt.style.use('classic')
plt.boxplot(x=veriler["max_heart_rate"])
plt.ylabel("Max Heart Rate");
# histogram
plt.hist(x=veriler["max_heart_rate"]);
# çizgi grafik 0-100 değerleri için
plt.plot(veriler["max_heart_rate"][0:100])
plt.xlabel("Heart Rate");
# +
# Sihirli Fonksiyonlar
# # %matplotlib notebook: İnteraktif grafikler çizdirmek için
# # %matplotlib inline: Statik görüntüler çizdirmek için
# -
# %matplotlib notebook
plt.plot(veriler["max_heart_rate"][0:100])
# %matplotlib inline
# ## Resimleri kaydetmek
x = np.linspace(0, 10, 100)
fig = plt.figure()
plt.plot(x, np.sin(x), '-', label='sinüs')
plt.plot(x, np.cos(x), '--', label='kosinüs')
plt.legend()
plt.xlabel("x")
plt.ylabel("y")
plt.title("Sinüs ve Kosinüs")
fig.savefig("figur.png")
# Scatter Plot
x = np.linspace(0, 10, 30)
y = np.sin(x)
plt.plot(x, y, 'o', color='black')
# +
# histogram
data = np.random.randn(1000)
plt.hist(data)
# -
plt.hist(data, bins=30, normed=True, alpha=0.5,
histtype='stepfilled', color='steelblue',
edgecolor='none')
# Çoklu grafik çizdirmek için
plt.subplot(2,1,1)
plt.plot([1,2,3])
plt.subplot(2,1,2)
plt.plot([1,2,5])
# Diğer yöntem
x = np.arange(10)
y1 = np.random.rand(10)
y2 = np.random.rand(10)
fig = plt.figure()
ax1 = fig.add_subplot(221)
ax1.plot(x,y1)
ax2 = fig.add_subplot(222)
ax2.scatter(x,y2)
ax3 = fig.add_subplot(223)
ax3.hist(y1)
ax4 = fig.add_subplot(224)
ax4.barh(x,y2)
# Online Kaynaklar
# 1. [Matplotlib Sitesi](matplotlib.org)
# 2. [Kitap](https://www.packtpub.com/application-development/interactive-applications-using-matplotlib)
|
uygulama_dersleri/veri_gorsellestirme.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This notebook tests the functionality of position.py
import os
os.chdir('../../source')
import position as pos
# ### Test StagePosition()
# +
stage = pos.StagePosition(x=0)
stage2 = pos.StagePosition(x=3, y=4, z=0)
stage3 = pos.StagePosition(x=0, y=0, z=0)
stage4 = pos.StagePosition(x=9, y=9, z=0, theta=90)
stage5 = pos.StagePosition(x=0, y=0, z=0)
stagelist = [stage, stage2, stage3, stage4, stage5]
stagelist2 = [stage2, stage]
# test __str__()
print (stage)
print (stage2)
print (stage3)
# -
# Test __eq__() function
assert (stage==stage2) is False, '== error'
assert (stage==stage3) is False, '== error'
assert (stage3==stage5) is True, '== error'
# Test dist()
assert stage3.dist(stage5) == 0, 'dist error'
assert stage2.dist(stage3) == 5, 'dist error'
# ### Test PositionList()
# +
# Test __init__(), append()
posit = pos.PositionList(stage)
posit.append(stage2)
posit2 = pos.PositionList(stage)
posit2.append(stage2)
posit3 = pos.PositionList(positions=stagelist)
# -
# Test __len__()
assert len(posit3) == 5
assert len(posit) == 2
assert len(posit2) == 2
# Test __add__()
p = posit + posit3
assert len(p) == 7
# Test __iter__()
for position in p:
print (position)
# +
# Test __getitem__(), __setitem__(), __delitem__()
assert p[1].x == 3
assert p[4].z == 0
p[1].x = 4
assert p[1].x == 4
del p[1]
for position in p:
print (position)
# -
# Test visualize()
del p[0]
del p[0]
del p[2]
p.visualize()
# Test save()
for i in p:
print (i)
p.save('test', './')
# Test load()
loaded = pos.load('test', './')
for i in loaded:
print (i)
# +
# Test current()
import MMCorePy
mmc = MMCorePy.CMMCore()
mmc.loadSystemConfiguration("../../config/MMConfig_YellenLab_ubuntu.cfg")
mmc.setFocusDevice("FocusDrive")
curr = pos.current(mmc)
print (curr)
# -
# Test set_pos()
pos.set_pos(mmc, x=curr[0], y=curr[1]+10)
print (pos.current(mmc))
|
SmartScope/smartscope/notebook/testing/.ipynb_checkpoints/position_test-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/algoritmosdenegociacion/modulo4/blob/main/M4_L2_Modelos_de_ML_para_Trading.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HDMUuXGmB_q1"
# # Algoritmos de Negociación basados en Machine Learning - Módulo 4
# - <NAME>, Ph.D., Universidad de los Andes
# - <NAME>, T.A., Universidad de los Andes
#
# https://github.com/algoritmosdenegociacion/
# + [markdown] id="CSr9zTNECaPv"
# ## 1. Carga de librerías, funciones y APIs necesarias.
#
# + [markdown] id="mF4dwMlYCb8D"
# #### 1.1. Instalan las librerías que no incluye Google Colab
# + id="aKnoONgqf_EC" colab={"base_uri": "https://localhost:8080/"} outputId="d29065e7-27ad-45a0-d489-6ad6cab1376d"
pip install yfinance
# + id="7wCoftrqrreP" colab={"base_uri": "https://localhost:8080/"} outputId="8fb2e511-a8f4-46e9-bad4-e4f7fb659c62"
pip install ta
# + [markdown] id="6zaoGOtoCjPq"
# #### 1.2. Se cargan las librerías necesarias
# + id="cVO4GoZZCgc8"
# Funciones numéricas adicionales
import numpy as np
# Lectura de datos y manejo de Data-sets
import pandas as pd
# Datos
import yfinance as yfin
# Gráficos
import matplotlib.pyplot as plt
# Análisis Técnico
import ta
# Funciones
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# Medidas de desempeño
from sklearn.metrics import accuracy_score, confusion_matrix, plot_roc_curve
# Support Vector Machine (SVM) / Support Vector Classification (SVC)
from sklearn.svm import SVC
# Decision Tree Classifier
from sklearn.tree import DecisionTreeClassifier
# Random Forest Classifier
from sklearn.ensemble import RandomForestClassifier
# Redes Neuronales / Multi-layer Perceptron (MLP)
from sklearn.neural_network import MLPClassifier
# + [markdown] id="0Dvota99Cn_O"
# ## 2. Obtención de datos históricos
#
# + [markdown] id="R_4CbC3JCq--"
# https://finance.yahoo.com/
# + id="7z9AKmNdCr7v" colab={"base_uri": "https://localhost:8080/", "height": 472} outputId="0916bdc2-ff27-4b18-f7cd-12c2dcb50ed4"
# Descargamos datos de 1 años de la acción de Tesla
df = yfin.download('BTC-USD',start='2016-01-01', end='2021-01-01')
df
# + [markdown] id="hJAMd1vGtaJM"
# ## 3. Análisis Técnico
# + colab={"base_uri": "https://localhost:8080/", "height": 649} id="Fgd3zwkadz3W" outputId="d0400fc6-b6f1-40b5-b954-1f569ae34a8f"
# Incluir todo el anáñisis técnico a la base de datos
# Medias móviles exponenciales
df['EMA_5'] = ta.trend.ema_indicator(close=df["Close"], window=5, fillna=True)/df["Close"]
df['EMA_20'] = ta.trend.ema_indicator(close=df["Close"], window=20, fillna=True)/df["Close"]
df['EMA_50'] = ta.trend.ema_indicator(close=df["Close"], window=50, fillna=True)/df["Close"]
df['EMA_100'] = ta.trend.ema_indicator(close=df["Close"], window=100, fillna=True)/df["Close"]
# Índice de fuerza relativa
df['RSI'] = ta.momentum.rsi(close=df["Close"], fillna=True)
# Rango medio verdadero
df['ATR'] = ta.volatility.average_true_range(high=df["High"], low=df["Low"], close=df["Close"], fillna=True)
# Rango de porcentaje de wiliams
df['WR'] = ta.momentum.williams_r(high=df["High"], low=df["Low"], close=df["Close"], fillna=True)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 649} id="Aq5LU3L0ynyO" outputId="50327317-2f3e-4dd6-8614-c13a342dc6d7"
# Crea el Target (1 si el precio sube el día siguiente, -1 si baja)
cl = np.array(df['Close'])
target = np.where(cl[1:] > cl[:-1], 1, -1)
# Elimina el ultimo día para el que no tenemos info del precio el día siguiente.
df.drop(df.tail(1).index, inplace=True)
# Creamos la columna target
df['Target'] = target
# Elimina los primeros 29 días dónde los indicadores técnicos no tienen suficiente info
df.drop(df.head(29).index, inplace=True)
df
# + [markdown] id="L0uCcDmyJTFM"
# ## 4. Conjunto de entrenamiento y prueba
# + colab={"base_uri": "https://localhost:8080/"} id="E1xXhH-mr7Iz" outputId="1fc2b690-4db9-4025-a16d-74b5d032c840"
# Divida el conjunto de datos en una característica o un conjunto de datos independiente (X) y un destino o conjunto de datos dependiente (Y)
X = np.array(df.iloc[:, 6:-1])
Y = np.array(df['Target'])
print(X.shape)
print(Y.shape)
# + id="SHrAH-ZsI2s1" colab={"base_uri": "https://localhost:8080/"} outputId="a65b29ff-ecfd-46d0-e7bb-9622b06f9107"
# Vuelva a dividir los datos, pero esta vez en 90% de entrenamiento y 10% de prueba.
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.1, shuffle=False)
print(X_train.shape)
print(X_test.shape)
# + id="cAXksFBUsbzO"
#Standardization
ss = StandardScaler()
X_train = ss.fit_transform(X_train)
X_test = ss.transform(X_test)
# + [markdown] id="o01VaYhRB4CU"
# ## 5. Machine Learning - Support Vector Classifier (SVC)
# + id="ITp-5OPyCWVF"
# Crear el modelo clasificador de Support Vector Machine.
# Kernel radial
svc = SVC(kernel='rbf')
# Entrenar el modelo de SVM
svc = svc.fit(X_train, Y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="a0NXNN1sCb6I" outputId="47422215-89d6-4e54-c530-5440b2f30c64"
# Compruebe cómo le fue al modelo en el conjunto de datos de prueba
svm_pred = svc.predict(X_test)
print("Accuracy: {:.4f}\n".format( accuracy_score(Y_test, svm_pred) ))
print("Confusion Matrix:")
print( confusion_matrix(Y_test, svm_pred) )
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="9kClbWh0Dgso" outputId="b8b47c4b-211b-4216-efe5-688e55333be7"
# Curva ROC y AUC.
plot_roc_curve(svc, X_test, Y_test)
# + [markdown] id="UunnZSSZe72G"
# ## 6. Machine Learning - Decision Tree Classifer
# + id="rcSj0_7p0EcX"
# Crear el modelo clasificador de Árbol de Decisión.
dtc = DecisionTreeClassifier()
# Entrenar el Árbol de Decisión
dtc = dtc.fit(X_train, Y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="Kw-GX3rT0o2h" outputId="74b08ba0-0705-4919-98f9-66175070c1a0"
# Compruebe cómo le fue al modelo en el conjunto de datos de prueba
dtc_pred = dtc.predict(X_test)
print("Accuracy: {:.4f}\n".format( accuracy_score(Y_test, dtc_pred) ))
print("Confusion Matrix:")
print( confusion_matrix(Y_test, dtc_pred) )
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="MpsH35pW1K7N" outputId="698236ab-4aee-4e64-a60e-9602c247ff9a"
# Curva ROC y AUC.
plot_roc_curve(dtc, X_test, Y_test)
plt.show()
# + [markdown] id="romG8tcxuud7"
# ## 7. Machine Learning - Random Forest Classifier
# + id="0nHxvMmi2U96"
# Crear el modelo clasificador de Random Forest.
rfc = RandomForestClassifier()
# Entrenar el modelo de Random Forest
rfc = rfc.fit(X_train, Y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="G8XinU042eMj" outputId="9eea3032-89d0-4a54-eecd-e3833b8c2a09"
# Compruebe cómo le fue al modelo en el conjunto de datos de prueba
rfc_pred = rfc.predict(X_test)
print("Accuracy: {:.4f}\n".format( accuracy_score(Y_test, rfc_pred) ))
print("Confusion Matrix:")
print( confusion_matrix(Y_test, rfc_pred) )
# + id="6gEIamrY2lGm" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="ae47defb-8d56-4fc3-8ec2-9786f15d4a0d"
# Curva ROC y AUC.
plot_roc_curve(rfc, X_test, Y_test)
plt.show()
# + [markdown] id="4hgJYDZzDyBt"
# ## 8. Machine Learning - Redes neuronales / Multilayer Perceptron (MLP)
# + id="9Ob_BqWVe0ZD"
# Redes Neuronales / Multi-layer Perceptron (MLP)
from sklearn.neural_network import MLPClassifier
# + id="proAGOujEG6i"
# Crear el modelo clasificador de Redes Neuronales.
mlp = MLPClassifier(hidden_layer_sizes=(64,32), max_iter=1000)
# Entrenar el modelo de Redes Neuronales
mlp = mlp.fit(X_train, Y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="z2mCFeN6EO1s" outputId="4bb2c7f0-5853-4dcf-dbcf-42bd899b5d7c"
# Compruebe cómo le fue al modelo en el conjunto de datos de prueba
mlp_pred = mlp.predict(X_test)
print("Accuracy: {:.4f}\n".format( accuracy_score(Y_test, mlp_pred) ))
print("Confusion Matrix:")
print( confusion_matrix(Y_test, mlp_pred) )
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="aAvh_JvSEcOT" outputId="f6b7b890-ca8a-46db-d8e2-160d5703f85f"
# Get the model metrics
plot_roc_curve(mlp, X_test, Y_test)
plt.show()
# + [markdown] id="f6zGffMEhstm"
# ## 9. Backtesting - Random Forest
# + [markdown] id="zZCmMnZ4Zv9p"
# #### 9.1. Cálculo de capital a lo largo de la venta de tiempo
# + colab={"base_uri": "https://localhost:8080/", "height": 753} id="-Z8uBrBviUEQ" outputId="20892cea-9fa3-411e-a51f-a43b1d2ed249"
N = len(rfc_pred)
df_test = df.tail(N)
df_test['Signals'] = rfc_pred
df_test
# + id="On59u_WhGYY_" colab={"base_uri": "https://localhost:8080/"} outputId="2dd79679-6f96-4dc6-b8b9-900dc93ee4b8"
# Crea una lista para almacenar el capital
# Emezamos con 100 USD
equity = [100]
# Take Profit al 3% y Stop Loss al 1%
TP = 0.03
SL = 0.01
pos = 0
price = -1
# Recorre la ventana de tiempo a partir del día 1
for i in range(1, N):
equity.append( equity[i-1] )
if pos == 1:
if df_test['Close'][i] >= price*(1 + TP):
equity[i] *= 1 + TP
pos = 0
elif df_test['Close'][i] <= price*(1 - SL):
equity[i] *= 1 - SL
pos = 0
elif pos == -1:
if df_test['Close'][i] <= price*(1 - TP):
equity[i] *= 1 + TP
pos = 0
elif df_test['Close'][i] >= price*(1 + SL):
equity[i] *= 1 - SL
pos = 0
else:
if df_test['Signals'][i] != 0:
pos = df_test['Signals'][i]
price = df_test['Close'][i]
df_test['Equity'] = equity
# + [markdown] id="x4KTqWfiZ4Tx"
# #### 9.2. Gráfico del capital
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="77mvALXgVWJ9" outputId="6c26edcc-e16f-4702-eee1-f1f7ecbb189b"
# Mostrar visualmente el equity a lo largo de la ventana de tiempo
plt.figure(figsize=(16,8))
plt.plot(df_test['Equity'])
plt.title('Capital - Random Forest')
plt.xlabel('Fecha')
plt.ylabel('USD ($)')
plt.show()
# + [markdown] id="BOW_qn-eaCAc"
# #### 9.3. Comparación con un portafolio de mercado
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="BFfct0RpaLif" outputId="563a8a75-27de-4cf3-e634-b2f12210a348"
mkt = yfin.download('^GSPC', start='2020-07-02', end='2020-12-31')
mkt
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="Eu10Ha0eaLgY" outputId="1fc58226-7458-4155-c0be-60fd60dba648"
mkt['Equity'] = (100/mkt['Close'][0])*mkt['Close']
mkt
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="QcPVCBSUbZCj" outputId="647f7b4a-555b-4fbc-9518-c08e50a5a804"
# Comparación de la estrategia con el portafolio de mercado
plt.figure(figsize=(16,8))
plt.plot(df_test['Equity'], color = 'green', label = 'Estrategia de trading (ML-Random Forest)')
plt.plot(mkt['Equity'], color = 'red', label = 'Portafolio de mercado')
plt.title('Comparación de la estrategia de ML con el portafolio de mercado')
plt.xlabel('Fecha')
plt.ylabel('Index (100)')
plt.legend(loc='lower right')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="IOPBWPCGc6aL" outputId="c6963a17-4b9c-4cd2-853b-05b27bf0a3cd"
# Desempeño de la estrategia planteada
ret = 252*np.log(df_test['Equity']).diff().mean()
print('Retorno esperado anulizado: {:.4f}'.format(ret))
vol = np.sqrt(252)*np.log(df_test['Equity']).diff().std()
print('Volatilidad anulizada: {:.4f}'.format(vol))
sharpe_ratio = (ret - 0.01)/vol
print('Sharpe Ratio: {:.4f}'.format(sharpe_ratio))
# + colab={"base_uri": "https://localhost:8080/"} id="taXd72Ghc6Pt" outputId="01656c28-92df-421b-dc70-a28b002e21f4"
# Desempeño del portafolio de mercado
ret = 252*np.log(mkt['Equity']).diff().mean()
print('Retorno esperado anulizado: {:.4f}'.format(ret))
vol = np.sqrt(252)*np.log(mkt['Equity']).diff().std()
print('Volatilidad anulizada: {:.4f}'.format(vol))
sharpe_ratio = (ret - 0.01)/vol
print('Sharpe Ratio: {:.4f}'.format(sharpe_ratio))
# + id="tGeF-3Okm7Qi"
|
M4_L2_Modelos_de_ML_para_Trading.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import random
# +
def remove_space(string, threshold = 0.3):
string = [s for s in string if not (s == ' ' and random.random() >= threshold)]
return ''.join(string)
def package(string, repeat = 2):
result = [(string, string)]
result.append((string.lower(), string.lower()))
for _ in range(repeat):
result.append((remove_space(string), string))
result.append((remove_space(string.lower()), string.lower()))
return result
from tqdm import tqdm
def loop(strings):
results = []
for i in tqdm(range(len(strings))):
p = package(strings[i])
results.extend(p)
return results
def slide(strings, n = 5):
result = []
for i in range(0, len(strings), len(strings) - (n - 1)):
result.append(strings[i: i + n])
return result
# -
files = ['/home/husein/pure-text/filtered-dumping-wiki.txt',
'/home/husein/pure-text/dumping-cleaned-news.txt',
'/home/husein/pure-text/dumping-iium.txt']
# +
with open(files[0]) as fopen:
data = fopen.read().split('\n')
results, result = [], []
for i in data:
if len(i) and i[-1] != '.':
i = i + '.'
if not len(i) and len(result):
results.append(result)
result = []
else:
if len(i):
result.append(i)
if len(result):
results.append(result)
len(results)
# +
from tqdm import tqdm
def loop(strings):
results = []
for i in tqdm(range(len(strings))):
try:
slided = slide(strings[i])
slided = [s for s in slided if len(s) > 1]
for s in slided:
s = ' '.join(s)
p = package(s)
results.extend(p)
except:
pass
return results
# +
import cleaning
results1 = cleaning.multiprocessing(random.sample(results, 1000), loop)
# -
testset = []
testset.extend(results1)
# +
with open(files[1]) as fopen:
data = fopen.read().split('\n')
len(data)
# +
results, result = [], []
for i in data:
if len(i) and i[-1] != '.':
i = i + '.'
if not len(i) and len(result):
results.append(result)
result = []
else:
if len(i):
result.append(i)
if len(result):
results.append(result)
# -
results = random.sample(results, 1000)
results1 = cleaning.multiprocessing(results, loop)
testset.extend(results1)
# +
with open(files[2]) as fopen:
data = fopen.read().split('\n')
results, result = [], []
for i in data:
if len(i) and i[-1] != '.':
i = i + '.'
if not len(i) and len(result):
results.append(result)
result = []
else:
if len(i):
result.append(i)
if len(result):
results.append(result)
# -
results = random.sample(results, 1000)
results1 = cleaning.multiprocessing(results, loop)
testset.extend(results1)
def generate_short(string):
splitted = string.split()
random_length = random.randint(2, min(len(splitted), 10))
end = random.randint(0 + random_length, len(splitted))
return ' '.join(splitted[end - random_length: end])
# +
with open(files[0]) as fopen:
data = list(filter(None, fopen.read().split('\n')))
data = [i for i in data if len(i) >= 2]
len(data)
# -
data = random.sample(data, 2000)
def loop(strings):
results = []
for i in tqdm(range(len(strings))):
try:
p = package(generate_short(strings[i]))
results.extend(p)
except:
pass
return results
results1 = cleaning.multiprocessing(data, loop)
results1[:10]
testset.extend(results1)
# +
with open(files[1]) as fopen:
data = list(filter(None, fopen.read().split('\n')))
data = random.sample(data, 2000)
# -
results1 = cleaning.multiprocessing(data, loop)
testset.extend(results1)
# +
with open(files[2]) as fopen:
data = list(filter(None, fopen.read().split('\n')))
len(data)
# -
data = random.sample(data, 2000)
results1 = cleaning.multiprocessing(data, loop)
testset.extend(results1)
def loop(strings):
results = []
for i in tqdm(range(len(strings))):
p = package(strings[i])
results.extend(p)
return results
# +
with open(files[0]) as fopen:
data = list(filter(None, fopen.read().split('\n')))
data = [i for i in data if len(i) >= 2]
# -
data = random.sample(data, 2000)
results1 = cleaning.multiprocessing(data, loop)
testset.extend(results1)
# +
with open(files[1]) as fopen:
data = list(filter(None, fopen.read().split('\n')))
data = random.sample(data, 2000)
results1 = cleaning.multiprocessing(data, loop)
# -
results1
testset.extend(results1)
# +
with open(files[2]) as fopen:
data = list(filter(None, fopen.read().split('\n')))
len(data)
# -
data = random.sample(data, 2000)
results1 = cleaning.multiprocessing(data, loop)
testset.extend(results1)
len(testset)
# +
import os
b2_application_key_id = os.environ['b2_application_key_id']
b2_application_key = os.environ['b2_application_key']
# -
from b2sdk.v1 import *
info = InMemoryAccountInfo()
b2_api = B2Api(info)
application_key_id = b2_application_key_id
application_key = b2_application_key
b2_api.authorize_account("production", application_key_id, application_key)
file_info = {'how': 'good-file'}
b2_bucket = b2_api.get_bucket_by_name('malay-dataset')
# +
import json
with open('test-set-segmentation.json', 'w') as fopen:
json.dump(testset, fopen)
# -
b2_bucket.upload_local_file(
local_file='test-set-segmentation.json',
file_name='segmentation/test-set-segmentation.json',
file_infos=file_info,
)
|
session/segmentation/t5/test-set-segmentation-augmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Advanced Python Programming
# ... or, how to avoid repeating yourself.
# ## Avoid Boiler-Plate
# Code can often be annoyingly full of "boiler-plate" code: characters you don't really want to have to type.
#
# Not only is this tedious, it's also time-consuming and dangerous: unnecessary code is an unnecessary potential place for mistakes.
#
# There are two important phrases in software design that we've spoken of before in this context:
#
# > Once And Only Once
#
# > Don't Repeat Yourself (DRY)
#
# All concepts, ideas, or instructions should be in the program in just one place.
# Every line in the program should say something useful and important.
#
# We refer to code that respects this principle as DRY code.
#
# In this chapter, we'll look at some techniques that can enable us to refactor away repetitive code.
#
# Since in many of these places, the techniques will involve working with
# functions as if they were variables, we'll learn some **functional**
# programming. We'll also learn more about the innards of how Python implements
# classes.
#
# We'll also think about how to write programs that *generate* the more verbose, repetitive program we could otherwise write.
# We call this **metaprogramming**.
#
|
ch07dry/01intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf15
# language: python
# name: tf15
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
params = {
"legend.fontsize": "x-large",
"axes.labelsize": "x-large",
"axes.titlesize": "x-large",
"xtick.labelsize": "x-large",
"ytick.labelsize": "x-large",
"figure.facecolor": "w",
"xtick.top": True,
"ytick.right": True,
"xtick.direction": "in",
"ytick.direction": "in",
"font.family": "serif",
"mathtext.fontset": "dejavuserif",
}
plt.rcParams.update(params)
data = np.load("./z_phot/all_paper1_regression_80perc_0_100.npz", allow_pickle=True)
cat_test = data["cat_test"]
y_caps_all = data["y_caps_all_test"]
y_prob = data["y_prob_test"]
morpho = np.argmax(y_prob, axis =-1)
caps = y_caps_all[range(len(y_caps_all)),morpho,:]
dim_names = [str(i+1) for i in range(16)]
caps = pd.DataFrame(caps, columns=dim_names)
# caps["Caps Length"] = np.max(y_prob, axis=-1)
dim_names = list(caps.columns)
fig, ax = plt.subplots(1,1)
cbar = ax.matshow(caps.corr(), cmap ="coolwarm", vmin=-1, vmax=1)
fig.colorbar(cbar)
xaxis = np.arange(len(dim_names))
ax.set_xticks(xaxis)
ax.set_yticks(xaxis)
ax.set_xticklabels(dim_names, rotation="vertical")
ax.set_yticklabels(dim_names)
plt.show()
extra_cat = np.load(
"/data/bid13/photoZ/data/pasquet2019/sdss_vagc.npz", allow_pickle=True
)["labels"]
extra_cat = pd.DataFrame(
{
"specObjID": extra_cat["specObjID"],
"sersic_R50_r": extra_cat["sersic_R50_r"],
"sersic_R90_r": extra_cat["sersic_R90_r"],
"sersic_R0_r": extra_cat["sersic_R0_r"],
# "sersicN_r": extra_cat["sersicN_r"],
}
)
# plt.scatter(extra_cat["sersicN_r"],extra_cat["sersic_R50_r"],marker=".")
# plt.xlabel("n")
# plt.ylabel("R")
cat_test = pd.DataFrame(cat_test)
cat_test = cat_test.merge(extra_cat, how="left", on="specObjID")
# +
cat = pd.DataFrame()
# cat["EBV"] = cat_test["EBV"]
cat["u"] = cat_test["cModelMag_u"]- cat_test["extinction_u"]
cat["g"] = cat_test["cModelMag_g"]- cat_test["extinction_g"]
cat["r"] = cat_test["cModelMag_r"]- cat_test["extinction_r"]
cat["i"] = cat_test["cModelMag_i"]- cat_test["extinction_i"]
cat["z"] = cat_test["cModelMag_z"]- cat_test["extinction_z"]
cat["u-g"] = (cat_test["modelMag_u"] - cat_test["extinction_u"]) - (
cat_test["modelMag_g"] - cat_test["extinction_g"]
)
cat["u-r"] = (cat_test["modelMag_u"] - cat_test["extinction_u"]) - (
cat_test["modelMag_r"] - cat_test["extinction_r"]
)
cat["u-i"] = (cat_test["modelMag_u"] - cat_test["extinction_u"]) - (
cat_test["modelMag_i"] - cat_test["extinction_i"]
)
cat["u-z"] = (cat_test["modelMag_u"] - cat_test["extinction_u"]) - (
cat_test["modelMag_z"] - cat_test["extinction_z"]
)
cat["g-r"] = (cat_test["modelMag_g"] - cat_test["extinction_g"]) - (
cat_test["modelMag_r"] - cat_test["extinction_r"]
)
cat["g-i"] = (cat_test["modelMag_g"] - cat_test["extinction_g"]) - (
cat_test["modelMag_i"] - cat_test["extinction_i"]
)
cat["g-z"] = (cat_test["modelMag_g"] - cat_test["extinction_g"]) - (
cat_test["modelMag_z"] - cat_test["extinction_z"]
)
cat["r-i"] = (cat_test["modelMag_r"] - cat_test["extinction_r"]) - (
cat_test["modelMag_i"] - cat_test["extinction_i"]
)
cat["r-z"] = (cat_test["modelMag_r"] - cat_test["extinction_r"]) - (
cat_test["modelMag_z"] - cat_test["extinction_z"]
)
cat["i-z"] = (cat_test["modelMag_i"] - cat_test["extinction_i"]) - (
cat_test["modelMag_z"] - cat_test["extinction_z"]
)
cat["sersicN_r"] = cat_test["sersicN_r"]
# cat["deVRad_r"] = cat_test["deVRad_r"]
# cat["sersic_R50_r"] = cat_test["sersic_R50_r"]
cat["sersic_R90_r"] = cat_test["sersic_R90_r"]
# cat["sersic_R0_r"] = cat_test["sersic_R0_r"]
cat["z_spec"] = cat_test["z"]
cat["absMag_u"] = cat_test["absMag_u"]
cat["absMag_g"] = cat_test["absMag_g"]
cat["absMag_r"] = cat_test["absMag_r"]
cat["absMag_i"] = cat_test["absMag_i"]
cat["absMag_z"] = cat_test["absMag_z"]
cat["lgm_tot_p50"] = cat_test["lgm_tot_p50"]
cat["sfr_tot_p50"] = cat_test["sfr_tot_p50"]
cat["specsfr_tot_p50"] = cat_test["specsfr_tot_p50"]
cat["v_disp"] = cat_test["v_disp"]
# cat["bptclass"] = cat_test["bptclass"]
# cat["age_mean"] = cat_test["age_mean"]
# cat["ssfr_mean"] = cat_test["ssfr_mean"]
# cat["logMass_median"] = cat_test["logMass_median"]
# cat["sersicN_u"] = cat_test["sersicN_u"]
# cat["sersicN_g"] = cat_test["sersicN_g"]
# cat["sersicN_i"] = cat_test["sersicN_i"]
# cat["sersicN_z"] = cat_test["sersicN_z"]
# cat["fracDev_r"] = cat_test["fracDev_r"]
# cat["deVAB_r"] = cat_test["deVAB_r"]
# cat["expAB_r"] = cat_test["expAB_r"]
# cat["petroR90_r"] = cat_test["petroR90_r"]
# cat["P_disk"] = cat_test["P_disk"]
# cat["P_edge_on"] = cat_test["P_edge_on"]
# cat["modelMag_u"] = cat_test["modelMag_u"]
# cat["modelMag_g"] = cat_test["modelMag_g"]
# cat["modelMag_r"] = cat_test["modelMag_r"]
# cat["modelMag_i"] = cat_test["modelMag_i"]
mask = np.all(np.isfinite(cat), axis =1)
cat_corr = np.array(cat)
caps_corr= np.array(caps)
# -
# # Distance Correlation
import dcor
nmad_threshold = 5
caps_dim = caps_corr.shape[1]
num_features = cat_corr.shape[1]
dcorr_mat = np.zeros((num_features, caps_dim))
for i in range(num_features):
x = caps_corr.T
y = cat_corr[:,i]
finite_mask = np.isfinite(y)
y = y[finite_mask]
x = x[:,finite_mask]
median = np.median(y)
nmad = np.abs(stats.median_abs_deviation(y, scale="normal"))
mad_mask = (y>= (median - nmad_threshold*nmad)) & (y<= (median + nmad_threshold*nmad))
y = y[mad_mask]
x = x[:,mad_mask]
y = np.repeat(y[np.newaxis,:], x.shape[0], 0)
dcorr_mat[i] = dcor.rowwise(dcor.distance_correlation, x, y, compile_mode=dcor.CompileMode.COMPILE_PARALLEL)
print(f"{cat.columns.to_list()[i]} percent rejected: {(~mad_mask).sum()*100/len(mad_mask)}")
# + tags=[]
y_labels_phot = [
r"$u$",
r"$g$",
r"$r$",
r"$i$",
r"$z$",
r"$u-g$",
r"$u-r$",
r"$u-i$",
r"$u-z$",
r"$g-r$",
r"$g-i$",
r"$g-z$",
r"$r-i$",
r"$r-z$",
r"$i-z$",
r"$n_{r}$",
r"$R_{90, r}$",
]
y_labels_spec = [
r"$z_{spec}$",
r"$\mathrm{M}_{u}$",
r"$\mathrm{M}_{g}$",
r"$\mathrm{M}_{r}$",
r"$\mathrm{M}_{i}$",
r"$\mathrm{M}_{z}$",
"log(M$_{\star}$)",
"log(SFR)",
"log(sSFR)",
r"$\sigma_{v}$",
]
fig, ax = plt.subplots(2, 1, figsize=(25, 20), sharex=True, gridspec_kw={'height_ratios': [1.7, 1]})
_ = sns.heatmap(
dcorr_mat[:17],
linewidths=0.2,
annot=True,
cmap="rocket",
cbar=False,
xticklabels=dim_names,
yticklabels=y_labels_phot,
# yticklabels=cat.columns.to_list(),
ax=ax[0],
robust=True,
annot_kws={"fontsize": 20},
vmin=0,
vmax=1,
)
_ = sns.heatmap(
dcorr_mat[17:],
linewidths=0.2,
annot=True,
cmap="rocket",
cbar=False,
xticklabels=dim_names,
yticklabels=y_labels_spec,
# yticklabels=cat.columns.to_list(),
ax=ax[1],
robust=True,
annot_kws={"fontsize": 20},
vmin=0,
vmax=1,
)
fig.subplots_adjust(hspace=0.05)
cbar = fig.colorbar(ax[0].collections[0], ax=ax)
cbar.ax.tick_params(axis="both", which="major", labelsize=25)
cbar.ax.set_ylabel("Distance Correlation", fontsize=40, labelpad=30)
ax[0].tick_params(axis="both", which="major", labelsize=25, labeltop=True, bottom=False, top=True, left=True, right=False)
ax[0].tick_params(axis="both", which="minor", labelsize=25)
ax[1].tick_params(axis="both", which="major", labelsize=25, labeltop=False, bottom=True, top=False, left=True, right=False)
ax[1].tick_params(axis="both", which="minor", labelsize=25)
ax[1].set_xlabel("Capsule Dimension", size=40)
fig.text(0.05,0.4,"Galaxy Property", size=40, rotation=90 )
fig.savefig("./figs/correlations.pdf", dpi=300, bbox_inches="tight")
# -
# plt.scatter(cat["sersicN_r"],cat["sersic_R50_r"],marker=".")
# plt.xlabel("n")
# plt.ylabel("R")
# ### Correlations among capsule dims
caps_dim = caps_corr.shape[1]
dcorr_caps_mat = np.zeros((caps_dim, caps_dim))
for i in range(caps_dim):
x = caps_corr.T
y = caps_corr[:,i]
y = np.repeat(y[np.newaxis,:], x.shape[0], 0)
dcorr_caps_mat[i] = dcor.rowwise(dcor.distance_correlation, x, y, compile_mode=dcor.CompileMode.COMPILE_PARALLEL)
fig, ax = plt.subplots(1, 1, figsize=(25, 20))
ax = sns.heatmap(
dcorr_caps_mat,
linewidths=0.2,
annot=True,
cmap="rocket",
xticklabels=dim_names,
yticklabels=dim_names,
# yticklabels=cat.columns.to_list(),
ax=ax,
robust=True,
annot_kws={"fontsize": 20},
vmin=0,
vmax=1,
)
cbar = ax.collections[0].colorbar
cbar.ax.tick_params(axis="both", which="major", labelsize=25)
cbar.ax.set_ylabel("Distance Correlation", fontsize=40)
ax.tick_params(axis="both", which="major", labelsize=25, labeltop=True)
ax.tick_params(axis="both", which="minor", labelsize=25)
ax.set_xlabel("Capsule Dimension", size=40)
ax.set_ylabel("Capsule Dimension", size=40)
fig.savefig("./figs/correlations.pdf", dpi=300, bbox_inches="tight")
from scipy.cluster import hierarchy
from scipy.spatial.distance import squareform
corr_linkage = hierarchy.linkage(dcorr_caps_mat, method="ward")#, optimal_ordering=True)
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(30,15))
dendro = hierarchy.dendrogram(corr_linkage, labels=dim_names, leaf_rotation=90, ax=ax1,)
dendro_idx = np.arange(0, len(dendro['ivl']))
ax2 = sns.heatmap(
dcorr_caps_mat[dendro['leaves'], :][:, dendro['leaves']],
linewidths=0.2,
annot=True,
cmap="rocket",
xticklabels=dendro['ivl'],
yticklabels=dendro['ivl'],
# yticklabels=cat.columns.to_list(),
ax=ax2,
robust=True,
annot_kws={"fontsize": 20},
vmin=0,
vmax=1,
)
cbar = ax2.collections[0].colorbar
cbar.ax.tick_params(axis="both", which="major", labelsize=25)
cbar.ax.set_ylabel("Distance Correlation", fontsize=40)
ax2.tick_params(axis="both", which="major", labelsize=25, labeltop=True)
ax2.tick_params(axis="both", which="minor", labelsize=25)
ax2.set_xlabel("Capsule Dimension", size=40)
ax2.set_ylabel("Capsule Dimension", size=40)
fig.tight_layout()
plt.show()
# -
from collections import defaultdict
cluster_ids = hierarchy.fcluster(corr_linkage, 0.5, criterion='distance')
cluster_id_to_feature_ids = defaultdict(list)
for idx, cluster_id in enumerate(cluster_ids):
cluster_id_to_feature_ids[cluster_id].append(idx)
selected_features = [v[0]+1 for v in cluster_id_to_feature_ids.values()]
selected_features
# # Spearman's correlation (not for paper)
from scipy.stats import spearmanr
spearman_corr = spearmanr(cat_corr[mask],caps_corr[mask],)[0]
spearman_corr = spearman_corr[:cat.shape[1],cat.shape[1]:]
plt.figure(figsize=(20,15))
sns.heatmap(spearman_corr, annot=True, cmap="icefire", xticklabels=dim_names, yticklabels=cat.columns.to_list())
|
notebooks/plot_correlations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] nbpresent={"id": "ccaf80c3-1220-4001-80ed-1519ddf82a05"} slideshow={"slide_type": "slide"}
# using hydrochemistry and simple visualization to differentiate groundwater samples
# ===
#
# presented in 2018 International ITB Geothermal Workshop
# 21 - 22 Maret 2018
#
# authors:
#
# - <NAME> [ORCID](https://orcid.org/0000-0002-1526-0863) [Google Scholar]()
# - <NAME> [Google Scholar](https://scholar.google.co.id/citations?hl=en&user=zymmkxUAAAAJ&view_op=list_works&sortby=pubdate)
# - <NAME> [Google Scholar](https://scholar.google.co.id/citations?user=G3uiPMoAAAAJ&hl=en)
# - <NAME> [Google Scholar](https://scholar.google.co.id/citations?user=t7CtT5MAAAAJ&hl=en&oi=ao)
# - <NAME> [Google Scholar](https://scholar.google.co.id/citations?user=-Z9rgsQAAAAJ&hl=en&oi=ao)
#
# 
#
# **Faculty of Earth Sciences and Technology**
#
# **Institut Teknologi Bandung**
# + [markdown] nbpresent={"id": "18903a2f-38e8-47d8-9310-aa6619abf376"} slideshow={"slide_type": "slide"}
# # before you continue
#
# this talk **is not** about:
#
# - **geology**, but how to differentiate the geology.
# - **geothermal**, but you may look at your data differently.
# - **results**, but methods.
#
# 
# + [markdown] nbpresent={"id": "1b872498-e595-4dce-bc7a-03287ef8d1fb"} slideshow={"slide_type": "slide"}
# # introduction
#
# - we have lots of data.
# - we know they are different but don't how to visualize the difference.
# - if we know how to do it, we don't have the skills and tools.
# - here we are proposing a new way to look at your data using **free tools** with a bit of programming skills.
#
# 
# + [markdown] nbpresent={"id": "35268107-19df-41af-b24e-30bdec734ee8"} slideshow={"slide_type": "slide"}
# ## but we do have spreadsheet
#
# yes, but with some limitations:
#
# - it's cell-based, you have to scroll you way to get what you want.
# - it's limited ability to visualize your data.
# - it has reproducibility issues: versioning, _point and click_, copy and paste to your show the results.
#
# 
# + [markdown] nbpresent={"id": "94554f0e-4aa5-4077-af52-c031fdc43c79"} slideshow={"slide_type": "slide"}
# ## I like it but I don't have programming skills
#
# - it's not that hard.
# - many **good people** share their work (including codes).
# - the **difficulties** are not comparable to the **coolness**. :-)
#
# 
# + [markdown] nbpresent={"id": "4daed158-5a99-4a46-9e05-4d36632a908f"} slideshow={"slide_type": "slide"}
# ## why codes?
#
# - **it's reproducible**: people can get the same results from the same code and data, with no copy-pasting.
# - **it's not only about the results**: but also the process. you can learn the process step by step.
# - it's about **pretty-informative** visualization
#
# 
# + [markdown] nbpresent={"id": "e9ed126e-f317-46a5-8f4d-301232b46827"} slideshow={"slide_type": "slide"}
# # what do we need?
#
# you may choose on or both:
#
# - `python` installation [Anaconda installation instruction](https://conda.io/docs/installation.html) [on Youtube](https://www.youtube.com/watch?v=YJC6ldI3hWk) or
# - `R`installation [instructions](a-little-book-of-r-for-time-series.readthedocs.io/en/latest/src/installr.html) or [on Youtube](https://www.youtube.com/watch?v=cX532N_XLIs)
# - in this case we use `python` with its `pandas` package
#
# 
# 
# 
# 
#
# + [markdown] nbpresent={"id": "8bf405c9-b409-466a-9f6e-7060ba853493"} slideshow={"slide_type": "notes"}
# Kami menggunakan `Python-Pandas` karena `Pandas` adalah `python library` yang lengkap, mudah digunakan, dan seperti halnya `R`, `Pandas` memiliki basis pengguna yang sangat banyak dan berkomitmen untuk membagikan ilmu dan pengetahuannya. Karena ini, Anda dapat menemukan dengan mudah berbagai tutorial berbasis teks atau video di internet. Sebelum menggunakannya, maka Anda akan perlu mengunduh dan menginstalasi `Python` dan `Pandas` yang kami tulis dalam tutorial terpisah.
#
# Tutorial ini akan terkait dengan:
#
# - artikel kami berjudul" `Using hydrochemistry and simple visualisation to differentiate groundwater samples`
# - penulis: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
# - event: ITB International Geothermal Workshop ITB
# - organizer: Faculty of Mining and Petroleum Engineering
# - references:
#
# - [Codebasics Youtube Channel](https://www.youtube.com/channel/UCh9nVJoWXmFb7sLApWGcLPQ)
# - [Pandas 0.22.0 documentation](https://pandas.pydata.org/pandas-docs/stable/)
# - [A little book of Python for multivariate analysis](http://python-for-multivariate-analysis.readthedocs.io/)
# - [<NAME>'s PCA tutorial](http://sebastianraschka.com/Articles/2015_pca_in_3_steps.html)
# - [<NAME>'s MachineLearningMastery Blog](https://machinelearningmastery.com/visualize-machine-learning-data-python-pandas/)
# - [Jupyter Notebook documentation](http://jupyter-notebook.readthedocs.io/)
# + [markdown] nbpresent={"id": "04e35d15-6168-4c91-8607-449a41861d1a"} slideshow={"slide_type": "slide"}
# # loading libraries
#
# we will use the following libraries:
# - `pandas` and `numpy` for numerical calculation,
# - `matplotlib`and `seaborn` for plotting, and
# - `scikitlearn` for the PCA and other machine learning techniques.
# + nbpresent={"id": "6a0db77a-3cff-4d9e-9a03-731084f2d0ba"} slideshow={"slide_type": "subslide"}
import pandas as pd # loading Pandas on to memory
from pandas.tools.plotting import scatter_matrix
import numpy as np # loading Numpy library on to memory
import matplotlib.pyplot as plt # loading plotting library on to memory
# %matplotlib inline
import seaborn as sns # loading seaborn library
# loading some functions from sklearn
from sklearn.preprocessing import scale
from sklearn.decomposition import PCA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from scipy import stats
# + [markdown] nbpresent={"id": "1b9556c2-6414-420d-beb8-98e54bc67061"} slideshow={"slide_type": "slide"}
# # data description
#
# we use:
#
# - `describe()`
# - `boxplot`
# - `scatter plot matrix`
# + [markdown] nbpresent={"id": "717c140a-46ac-48ae-b735-3012d7a12049"} slideshow={"slide_type": "notes"}
# # Deskripsi data
#
# Kami menggunakan fungsi describe() dan boxplot untuk mengevaluasi data. Selain itu scatter plot matrix juga digunakan untuk mengetahui korelasi antar parameter.
# + nbpresent={"id": "0520781b-79c4-45be-893d-895182f60a6e"} slideshow={"slide_type": "subslide"}
df = pd.read_csv('data_arifs_2.csv') # loading data
# + [markdown] nbpresent={"id": "1004ea42-1e91-4d1e-b562-17babcd198ef"} slideshow={"slide_type": "subslide"}
# ## head and tail
#
# - we could see the first 5 lines using `foo.head()` and the last 5 lines using `foo.tail()`.
# - we need to see the type of data using `type(foo)` command.
# - checking the dimension or shape of the data (sum of rows and columns) using `foo.shape` command.
# - Change `foo` with your own data frame.
# + nbpresent={"id": "f4672e50-4a8c-42ad-8ea1-12c8340daf41"} slideshow={"slide_type": "subslide"}
df.head() # showing first 5 rows
# + nbpresent={"id": "d0fa239a-ad7c-4570-98bd-74345718f57b"} slideshow={"slide_type": "subslide"}
df.tail() # showing last 5 rows
# + nbpresent={"id": "340c4fd9-b5a1-46ee-964b-19ceffca21ce"} slideshow={"slide_type": "skip"}
type(df)
# + nbpresent={"id": "b6ea4cb0-f333-41ad-ba64-2a66c53f7cc7"} slideshow={"slide_type": "skip"}
df.shape # table size showing number of (rows, columns)
# + nbpresent={"id": "00d528bb-12f8-4041-8dba-c4c9c9499a96"} slideshow={"slide_type": "subslide"}
df.describe() # selecting columns in number and describe basics stats
# + nbpresent={"id": "1b82ffd6-a57e-4472-b97a-a3cf7a513d7b"} slideshow={"slide_type": "subslide"}
list(df)
# + [markdown] nbpresent={"id": "cd9b4ef4-22c8-4af9-b2c6-1e577a15a391"} slideshow={"slide_type": "slide"}
# # creating boxplot
#
# - here we create a boxplot to visualize the distribution of dataset.
# - we're going to make two kinds of layout.
# + nbpresent={"id": "40afa030-2f8c-4a11-b15b-010972e9398e"} slideshow={"slide_type": "subslide"}
df.boxplot(figsize=[20,10]) # creating boxplot
plt.savefig('box.png')
# + nbpresent={"id": "90b7e561-ca4b-47f2-bf4d-7b8cab2a392e"} slideshow={"slide_type": "subslide"}
df.boxplot(by=['litho'], figsize=[30,15]) # creating boxplot grouped by lithology
plt.savefig('panel_box.png')
# + [markdown] nbpresent={"id": "0670d0c7-a3bb-4796-8777-a858935c4db8"} slideshow={"slide_type": "slide"}
# # Correlation matrix
#
# ## omitting some non-numeric columns
#
# In the PCA process, we will not be using non-numerical columns: `sample`, `litho`, `turb`, `col`, and `source`. Also, `li` (Lithium) column contains zeros. We will drop them. First we're going to see the correlation matrix. Here we build the matrix in table form and in plot form.
# + nbpresent={"id": "2e5c108c-6014-45b2-b9d6-dc5ad5964a28"} slideshow={"slide_type": "subslide"}
df_cor = df.drop(['sample', 'litho', 'turb', 'col', 'source', 'li'], axis=1)
df_cor
list(df_cor)
# + nbpresent={"id": "7c1ec84b-7701-494c-906b-a56872026623"} slideshow={"slide_type": "subslide"}
df_cor
# + nbpresent={"id": "4322b618-0891-4ebb-8cd1-23f866cd9123"} slideshow={"slide_type": "subslide"}
corr = df_cor.corr()
corr
# + [markdown] nbpresent={"id": "671a38be-2fab-4b2c-b6e1-d1f80aac7dcd"} slideshow={"slide_type": "slide"}
# ## scatter plot matrix
#
# Then we visualize the correlation matrix in form of scatter plot matrix. We're going to see to types of scatter plot matrix. The first one builds on `pandas` function. The function automatically produces a separate window to contain the plot. The second plot, we define a custom-made function.
# + nbpresent={"id": "6446cc79-1d1a-495d-8c6c-c2cda733ab8a"} slideshow={"slide_type": "subslide"}
scatter_matrix(df_cor, figsize=(8,8))
plt.savefig('scatmat1.png')
# + nbpresent={"id": "3688b5dd-9c8d-49e5-89e3-6587bd306928"} slideshow={"slide_type": "subslide"}
def plot_corr(df_cor, size=10):
'''function to plot a graphical correlation matrix input: df: pandas DataFrame, size: vertical and horizontal size of the plot'''
fig, ax = plt.subplots(figsize = (size, size))
ax.matshow(corr)
plt.xticks(range(len(corr.columns)), corr.columns);
plt.yticks(range(len(corr.columns)), corr.columns)
plot_corr(df_cor, size = 10)
# + nbpresent={"id": "53136348-6e31-4779-8797-1ac9384ccd69"} slideshow={"slide_type": "notes"}
plt.savefig('scatmat2.png') # use this line only if you want to save the plot
# + [markdown] nbpresent={"id": "644ca130-617d-44a4-8b2e-540ff6800626"} slideshow={"slide_type": "subslide"}
# # we find the following correlations
#
# - TDS-DHL/EC with: K, HCO3, Cl, SO4, CO2, dan NO3
# - K with HCO3 and Cl
# - NH4 with Cl, SO4, NO2, dan NO3
# - Cl with SO4, NO2 dan NO3
# - NO2 with NO3
# + [markdown] nbpresent={"id": "dcc319c0-21b6-4039-ba7f-757dde4b10a5"} slideshow={"slide_type": "slide"}
# # multivariate analysis
#
# we will use principal component analysis (PCA) and later on cluster analysis (CA) to separate water quality samples.
# + [markdown] nbpresent={"id": "21c802fc-5c87-4ca7-a509-da4932631cbe"} slideshow={"slide_type": "subslide"}
# # steps
#
# - scale or normalize of dataset using `scale()` function
# - creating PCA model using `PCA()`
# - evaluating PCA
# - visualize PCA
# + [markdown] nbpresent={"id": "a590eacd-ec8c-476e-bc62-d3832dcf6247"} slideshow={"slide_type": "notes"}
# # Multivariate analysis
#
# Di sini kita mencoba menggunakan dua analisis multivariabel, Principal Component Analysis dan Cluster Analysis, untuk memisahkan sampel air berdasarkan karakter sifat fisik dan kimianya. Kami menggunakan library Scikit-Learn untuk melakukan ini.
#
# ## Principal component analysis (PCA)
# Dalam tahap ini kami menggunakan fungsi `PCA` dari Pandas. Sebelumnya proses standardisasi atau normalisasi perlu dilakukan dengan fungsi `scale`. Hasil dari fungsi `PCA` adalah nilai per variabel terhadap komponen 1 dan komponen 2. Jadi nantinya 18 variabel yang diukur akan dimasukkan ke dalam dua komponen utama (PC1 dan PC2). Dengan demikian akan muncul dua komponen yang merupakan transformasi dari 18 variabel awal. Berbekal dua komponen besar dan bukan 18 variabel terpisah, akan memudahkan kami untuk melakukan interpretasi lebih lanjut. Karena sebab inilah, maka PCA adalah salah satu teknik pengurangan dimensi atau _dimension reduction_.
# + [markdown] nbpresent={"id": "3353c68a-b3e2-4726-a16d-e8e020f0427a"} slideshow={"slide_type": "notes"}
# ### Creating PCA model and fitting
# Tahap pertama adalah melakukan normalisasi dengan `scale()` dan kemudian menjalankan proses PCA dengan `pca()`. Dalam proses PCA, data yang tadinya terdiri dari 18 variabel (atau sumbu atau dimensi), ditransformasi menjadi beberapa komponen saja. Biasanya fungsi `pca()` akan mengajukan empat komponen untuk dipilih, tapi pengguna bisa saja menentukan berapa jumlah komponen yang dihasilkan sejak awal, misalnya 2 komponen.
# + nbpresent={"id": "f5742379-ea00-4f03-883b-b33436a38ad3"} slideshow={"slide_type": "subslide"}
# scaling the dataset
standardisedX = scale(df_cor) # scale() from sklearn
standardisedX = pd.DataFrame(standardisedX, index=df_cor.index, columns=df_cor.columns)
# + nbpresent={"id": "4bf894e7-8d5c-481d-8fb9-3c6c9682b07c"} slideshow={"slide_type": "subslide"}
from sklearn.decomposition import PCA
pca = PCA(n_components=2, svd_solver='full')
pca.fit(df_cor)
existing_2d = pca.transform(df_cor)
existing_df_2d = pd.DataFrame(existing_2d)
existing_df_2d.index = df_cor.index
existing_df_2d.columns = ['PC1','PC2']
existing_df_2d
existing_df_2d.to_csv('us_pc.csv')
# + nbpresent={"id": "289efd73-d11c-4c13-9fcc-0d4259a26c19"} slideshow={"slide_type": "subslide"}
print(pca.explained_variance_ratio_)
# + [markdown] nbpresent={"id": "d4db8c79-a674-4727-a1cf-75bf7ea2563c"} slideshow={"slide_type": "notes"}
# ### Evaluating PCA fit
# Di sini kami mengevaluasi model PCA yang telah dihasilkan, yaitu dengan menghitung dan memplot jumlah komponen yang mampu menangkap variansi kumulatif terbesar dari data yang mampu dijelaskan oleh model (_cumulative explained variance_).
# + nbpresent={"id": "71fa3c09-1e6b-4a98-aba8-36f1e2ed6aeb"} slideshow={"slide_type": "subslide"}
cumsum = plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance');
# + [markdown] nbpresent={"id": "94c63f24-7def-458f-b1e5-43663dfee159"} slideshow={"slide_type": "subslide"}
# ## calculating `eigenvalue`
#
# This function is borrowed from this [source](http://python-for-multivariate-analysis.readthedocs.io/a_little_book_of_python_for_multivariate_analysis.html#loadings-for-the-principal-components) for calculating `eigenvalue`.
# + nbpresent={"id": "3dce15dd-59c5-4994-abd2-a9a69bd9ef6f"} slideshow={"slide_type": "subslide"}
pca = PCA(n_components=2, svd_solver='full').fit(standardisedX)
pca.fit = pca.fit(df_cor)
# + nbpresent={"id": "562ee0b1-f9fe-4db7-a490-0348e9633219"} slideshow={"slide_type": "subslide"}
X_transformed = pca.fit_transform(df_cor)
# We center the data and compute the sample covariance matrix.
df_cor_centered = df_cor - np.mean(df_cor, axis=0)
cov_matrix = np.dot(df_cor_centered.T, df_cor_centered) / 20
eigenvalues = pca.explained_variance_
for eigenvalue, eigenvector in zip(eigenvalues, pca.components_):
print(np.dot(eigenvector.T, np.dot(cov_matrix, eigenvector)))
print(eigenvalue)
# + nbpresent={"id": "09d8876e-94f9-4e19-b00c-b99cc82336bc"} slideshow={"slide_type": "subslide"}
X_transformed
# + nbpresent={"id": "0555b237-cf1e-4357-bdf5-13929f0a56cb"} slideshow={"slide_type": "subslide"}
type(eigenvalue)
eigval = pd.Series({2: 521036.29562818405, 1: 548459.2585559834, 4: 24341.049177525907, 3: 25622.157028974627})
eigval.plot.bar(figsize=(16,8))
# + [markdown] nbpresent={"id": "033dea76-1b81-4b9d-9eb8-b0e52cd9fcb1"} slideshow={"slide_type": "subslide"}
# ## ploting loading/vector
#
# Here we plot loadings (R's term) or vectors (python's term) of the PCA model.
# + nbpresent={"id": "af793dda-c41d-46b0-b3d7-fe4a7324dcd0"} slideshow={"slide_type": "subslide"}
pcdf = pd.DataFrame(data = X_transformed, columns = ['PC1', 'PC2'])
fig, ax = plt.subplots()
ax.scatter(x=pcdf["PC1"], y=pcdf["PC2"])
# + nbpresent={"id": "b2f83761-fffb-40ea-aaf2-fd4051fcc584"} slideshow={"slide_type": "subslide"}
pcdf
# + nbpresent={"id": "adaad656-789f-40b2-8a69-1a927f4eb9f3"} slideshow={"slide_type": "subslide"}
df_cor
# + nbpresent={"id": "3a5bdaf7-dbfd-46c0-98ae-bb7724fe02fa"} slideshow={"slide_type": "subslide"}
df_cor.columns
# + nbpresent={"id": "67b9417e-4c22-4f47-bbfe-9485cf17c4f8"} slideshow={"slide_type": "subslide"}
pcdf
# + nbpresent={"id": "90f2aa39-1fa2-433f-b0f4-100d8890fc11"} slideshow={"slide_type": "subslide"}
varid = pd.DataFrame(df_cor.columns)
# + nbpresent={"id": "01cadbeb-0aa5-482d-ae5a-3dca8c8e7141"} slideshow={"slide_type": "subslide"}
pcdf = varid.join(pcdf) # adding variable id to pcdf
pcdf
# + nbpresent={"id": "3c891fc2-0300-4eb4-ab90-cec4ffdf4bff"} slideshow={"slide_type": "subslide"}
def biplot(score, coeff, labels=None):
xs = score[:,0]
ys = score[:,1]
n = coeff.shape[0]
scalex = 1.0/(xs.max() - xs.min())
scaley = 1.0/(ys.max() - ys.min())
plt.scatter(xs * scalex, ys * scaley)
for i in range(n):
plt.arrow(0, 0, coeff[i,0], coeff[i,1],color = 'r',alpha = 0.5)
if labels is None:
plt.text(coeff[i,0]* 1.15, coeff[i,1] * 1.15, "Var"+str(i+1), color = 'g', ha = 'center', va = 'center')
else:
plt.text(coeff[i,0]* 1.15, coeff[i,1] * 1.15, labels[i], color = 'g', ha = 'center', va = 'center')
plt.xlim(-1,1)
plt.ylim(-1,1)
plt.xlabel("PC{}".format(1))
plt.ylabel("PC{}".format(2))
plt.grid()
#Call the function. Use only the 2 PCs.
biplot(X_transformed[:,0:2], np.transpose(pca.components_[0:2, :]), labels=pcdf[0])
# + [markdown] nbpresent={"id": "2df9f1bf-7c65-495b-895d-a41ee20a25fb"} slideshow={"slide_type": "notes"}
# PCA pada dasarnya adalah alat untuk mengurangi dimensi (baca: variabel) atau *dimension reduction*. Bila semula kami punya 18 variabel, kemudian ditransformasi oleh PCA menjadi dua variabel, berupa `komponen principal (PC)` (*principal component*), seperti dapat dilihat pada diagram balok di atas. Sekarang kita lihat variabel apa saja yang berperan membentuk PC 1 dan PC2.
# + [markdown] nbpresent={"id": "ed042510-3279-4e06-bd45-a3660283039e"} slideshow={"slide_type": "notes"}
# Plot di atas memperlihatkan lompatan nilai eigen value yang sangat besar antara PC2 dan PC3. Atas dasar itu, kami memilih analisis dilanjutkan pada PC1 dan PC2 untuk menangkap variansi terbesar dari data yang ada.
# + [markdown] nbpresent={"id": "f84f937b-dbac-4c59-af5b-0f362351f996"} slideshow={"slide_type": "notes"}
# ### Visualizing PCA fit
# Di sini kami membuat beberapa visualisasi model PCA menggunakan _scatter plot_ sederhana.
# + nbpresent={"id": "50b7fae9-bc2f-4d80-8b35-bd6f165db8d8"} slideshow={"slide_type": "subslide"}
ax = existing_df_2d.plot(kind='scatter', x='PC2', y='PC1', figsize=(16,8))
for i, sample in enumerate(existing_df_2d.index):
ax.annotate(sample, (existing_df_2d.iloc[i].PC2, existing_df_2d.iloc[i].PC1))
# + [markdown] nbpresent={"id": "f33dc61c-95a4-48e2-a563-f49eee07b1ee"} slideshow={"slide_type": "notes"}
# Perhatikan plot di atas. Index data menggunakan no urut. Kami ingin menambahkan identitas pada tiap titik data. Untuk itu kami tambahkan kolom `litho` dan `sampleid` ke dalam `existing_df_2d` (dataframe hasil fit PCA). Kemudian kami set kolom `sampleid` sebagai index.
# + nbpresent={"id": "39a97fc8-0701-4bc0-95da-e2b19be65af7"} slideshow={"slide_type": "skip"}
lithoid = pd.DataFrame(df['litho'])
type(lithoid)
sampleid = pd.DataFrame(df['sample'])
type(sampleid)
existing_df_2d = lithoid.join(existing_df_2d)
# + nbpresent={"id": "77295454-cb5b-4b33-9d0b-b223bc32f3e4"} slideshow={"slide_type": "subslide"}
existing_df_2d
# + nbpresent={"id": "246c6a25-2b0e-41ad-be24-a0b7dd3e7e34"} slideshow={"slide_type": "subslide"}
existing_df_2d = pd.concat([sampleid, existing_df_2d], axis=1)
existing_df_2d
# + nbpresent={"id": "d4bf1280-d2a8-4af2-9d54-bba79f9406db"} slideshow={"slide_type": "subslide"}
existing_df_2d.set_index('sample', inplace=True)
ax = existing_df_2d.plot(kind='scatter', x='PC2', y='PC1', figsize=(16,8))
for i, sample in enumerate(existing_df_2d.index):
ax.annotate(sample, (existing_df_2d.iloc[i].PC2, existing_df_2d.iloc[i].PC1))
# + [markdown] nbpresent={"id": "dfe35934-8adf-4f56-a5bb-1d13a7212c7b"} slideshow={"slide_type": "slide"}
# # results and discussions
#
# we should see separation between samples:
#
# - group 1 strong `Cl`
# - group 2 strong `CO3`
# - group 3 strong contaminations of NO2, NO3, NH4?
# - group 4 strong `SO4`
# + [markdown] nbpresent={"id": "629f933d-7d1a-4e1b-8f33-c22a1d25da9f"} slideshow={"slide_type": "notes"}
# # Hasil dan diskusi
#
# Dari plot di atas dapat dilihat bahwa sampel dari kawasan pantai Indramayu (Indra1 - Indra5) dan sampel dari Padalarang (Pad1 - Pad4) terpisah dari sampel endapan gunung api (Bdg1 - Bdg8) dan Pangalengan (Pang1 dan Pang2). Kemungkinan besar dari nilai Cl nya untuk sampel dari daerah pesisi Indramayu dan karena tingginya nilai CO3 atau HCO3 untuk sampal-sampel dari Padalarang. Namun demikian, model ini akan jadi berbeda bila ada data air panas jenis klorida yang masuk ke dalam plot. Demikian pula untuk sampel air dari akuifer endapan volcanik Bdg7 dan Bdg8 memisahkan dari sampel dari endapan gunung api lainnya berkode Bdg1-Bdg6 dan Pang1-Pang2. Ini menarik bila dilihat bahwa sampel Bdg7 dan Bdg8 terletak lebih mendekati Kota Bandung dibanding sampel-sampel lainnya. Kedua sampel tersebut telah mendapatkan pengaruh dari komponen NH4, NO2 dan NO3, yang menjadi tolok ukur adanya aktivitas manusia. Apakah ini berarti mata air tersebut telah mengalami pencampuran dari resapan limbah domestik atau pertanian dari permukaan? Memerlukan data dan observasi lebih rinci untuk menjawabnya.
# + [markdown] nbpresent={"id": "43b2406e-7fd9-4cc0-9c55-12b57648b430"} slideshow={"slide_type": "slide"}
# # conclusion
#
# we can divide the `toy samples` in to 3 groups:
#
# - **group 1** samples from coastal area (eg: Indramayu)
# - **group 2** samples from limestone area (eg: Padalarang)
# - **group 3** samples from `inner city-lowland` volcanic area (eg: Bandung)
# - **group 4** samples from `outer city-highland` volcanic area (eg: Pangalengan)
# + [markdown] nbpresent={"id": "d5c692ce-21dc-4ef2-9dd1-1ba1a6e06a89"} slideshow={"slide_type": "notes"}
# # Kesimpulan
#
# Dari proses ini dapat dilihat bahwa kami berhasil menguraikan sampel-sampel kualitas air yang dimiliki menjadi beberapa bagian, yakni __Kelompok 1__: sampel dari pesisir Indramayu; __Kelompok 2__: sampel dari kawasan batugamping Padalarang; serta __Kelompok 3__: sampel dari endapan gunungapi Bandung dan Pangalengan. Kelompok 3 masih dapat dibagi lagi menjadi sampel yang berada di bagian hulu yang relatif belum mendapatkan pengaruh aktivitas manusia, dan sampel dari bagian hilir (mungkin dekat dengan kawasan pemukiman) yang mungkin telah menerima pengaruh dari aktivitas manusia. Harapan kami, metode ini dapat diaplikasikan untuk analisis sampel data kualitas air hipertermal guna mengidentifikasi proses yang terjadi, untuk membedakannya dari air dingin (mesotermal), atau mengidentifikasi ada atau tidaknya pengaruh sistem geotermal kepada sistem air tanah yang digunakan oleh penduduk sekitarnya.
# + [markdown] nbpresent={"id": "9d7a6086-b519-490b-bdf3-fe44a374bb9e"} slideshow={"slide_type": "slide"}
# ## take home message
#
# - sometimes data might behave beyond our visual recognition.
# - this multivariable technique might give you more insight from your data.
# - we hope this method can assist you to foreseen the unseen in your data.
# - all resources are accessible via [GitHub repository](https://github.com/dasaptaerwin/iigw2018).
# + [markdown] nbpresent={"id": "d5d428e5-ecc5-4530-b576-67598c8565ce"} slideshow={"slide_type": "slide"}
# # should anyone be interested in this `mess`, here's my contact:
#
# - email: `dasaptaerwin at gmail`
# - twitter handle: `@dasaptaerwin`
|
Old-projects/Jupyter/IIGW2018-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Install all requirements for the project
# +
# # !pip install numpy
# # !pip install pandas
# # !pip install matplotlib
# # !pip install seaborn
# -
# #### Importing Libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# Path
DOCS_PATH = 'docs/'
DATA_PATH = 'data/'
# -
# #### Importing our Datasets
commune_df = pd.read_excel((DATA_PATH + "commune.xlsx"))
enroll_df = pd.read_csv(DATA_PATH + "enroll.csv")
industry_df = pd.read_csv(DATA_PATH + "industry.csv")
ord_df = pd.read_csv(DATA_PATH + "ord.csv")
quest_df = pd.read_csv(DATA_PATH + "quest.csv")
studyd_df = pd.read_csv(DATA_PATH + "study_domain.csv")
tech_df = pd.read_csv(DATA_PATH + "technology.csv")
transaction_df = pd.read_csv(DATA_PATH + "transaction.csv")
# # Ayiti Analytics Data Processing Bootcamp
# Ayiti Analytics Data wants to expand its training centers throughout all the communes of the country. Your role as a data analyst is to help them realize this dream.
#
# Its objective is to know which three communes of the country will be the most likely to expand its training centers.
#
# Knowing that each cohort must have 30 students
#
# * How many applications must be made to select 25% women for each on average
#
# * What are the most effective communication channels (Alumni, Facebook, WhatsApp, Friend ...) that will allow a student to be susceptible to selection
#
# * What is the average number of university students who should participate in this program
# * What will be the average number of applications per week that we could have
# * How many weeks should we extend the application process to select 60 students per commune?
# * If we were to do all the bootcamp online, who would be the best communes and how many applications would we need to select 30 student and what percentage of students would have a laptop, an internet connection, both at the same time
# * What are the most effective communication channels (Alumni, Facebook, WhatsApp, Friend ...) that will allow a women to be susceptible to selection
#
# ### NB
# Use the same framework of the BA project to complete this project
# >## Commune dataset
# #### Display the Dataframe
commune_df.head(5)
# #### Data overview
# Check the dimensions of the DataFrame
row, col = commune_df.shape
print("This datasset have",row,"rows","and",col,'columns')
# Data Types
commune_df.dtypes
# Data info
commune_df.info()
# Describe the dataset
commune_df.describe()
# #### Data cleaning
# Make sure our columns don't have duplicate values.
commune_df.duplicated().sum()
# Describe the dataset
commune_df.describe()
# dealing with missing data
commune_df.isna().sum()
# Drop all columns we will not use in this dataset
to_drop = []
clean_commune_df = commune_df.drop(['Commune_FR'], axis = 1)
# display ou new dataframe
clean_commune_df.head(3)
# **Conclusion:** just checking that there's no missing and duplicated values in the commune dataset...
# >## Enroll Dataset
# #### Import Dataset
# Affiche done enroll yo
enroll_df.head(2)
# #### Data overview
enroll_df.shape
# #### Data cleaning
# Finding duplicated rows
print(enroll_df.duplicated().sum(),"Duplicated rows")
# Count how many missing values we have in this dataset
enroll_df.isnull().sum()
# Total missing values we have
print("We have", enroll_df.isnull().sum().sum(), "missing values in this dataset")
# Dealing with our null values
clean_enroll_df = enroll_df.fillna(value = 0)
print("We now have", clean_enroll_df.isnull().sum().sum(), "missing value")
# Our cleaned Dataframe
clean_enroll_df.head(2)
# **Conclusion:** All the missing values in the Enroll dataset have been filled with 0 as new value.
# Also we will be using our new dataframe called **clean_enroll_df**
# >## Industry Dataset
# Affiche done industry yo
industry_df.head(3)
# >## Ord Dataset
# Affiche done ord yo
ord_df.head(3)
# #### Data overview
ord_df.info()
ord_df.shape
ord_df.describe()
# #### Data cleaning
# Duplicated rows
print(ord_df.duplicated().sum(), "Duplicated rows")
# Show how many missing values we have in this dataset
ord_df.isnull().sum()
# Count all of our missing values
print(ord_df.isnull().sum().sum(), "Null values")
# Dealing with our missing values
clean_ord_df = ord_df.fillna(value=0)
print("Now we have", clean_ord_df.isnull().sum().sum(), "missing value Now")
# Our new dataframe
clean_ord_df.head(3)
# **Conclusion:** All the missing values in the Ord dataset have been filled with 0 as new value.
# Also we will be using our new dataframe called **clean_ord_df**
# >## Questions dataset
# Affiche done quest yo
quest_df.head(2)
# #### Data overview
quest_df.shape
quest_df.describe()
quest_df.info()
# #### Data cleaning
# Looking for duplicated rows
quest_df.duplicated().sum()
# Missing values
quest_df.isnull().sum()
# Total missing values
print('We have',quest_df.isnull().sum().sum(),'missing values in our dataset')
# Dealing with our missing values
clean_quest_df = quest_df.fillna({'dob':'not specified', 'department':'no info'}).fillna(0)
print("Now all this dataframe is clean we have", clean_quest_df.isnull().sum().sum(), "missing value")
# Rename the column commune to Commune_Id
# I do this because i need this dataframe to be able to merge easily with the commune dataset
clean_quest_df = clean_quest_df.rename(columns={'commune':'Commune_Id'})
# Add all our "commune" values on uppercase to match with our commune ID
clean_quest_df["Commune_Id"] = clean_quest_df["Commune_Id"].str.upper()
# As we can see in our dataframe we have Date let's convert the date to DateTime type
clean_quest_df['created_at'] = pd.to_datetime(clean_quest_df['created_at'])
clean_quest_df['created_at'].dtype
# Display our clean dataframe
clean_quest_df.head(2)
# >## Study domain Dataset
# Affiche done Study domain yo
studyd_df.head(2)
studyd_df.isnull().sum().sum()
studyd_df.duplicated().sum()
# >## Technology Dataset
# Affiche done technology yo
tech_df.head(3)
# Duplicated rows in our dataset
print(tech_df.duplicated().sum(),'Dupicated rows')
# Missing values
print(tech_df.isnull().sum().sum(),"Null values")
# Affiche done industry yo
transaction_df.head(3)
print(transaction_df.duplicated().sum(),'Duplicated rows')
print(transaction_df.isnull().sum().sum(),'Missing values')
# ## Looking for the top three communes of the country that will be the most likely to expand its training centers.
# Review our commune dataframe columns
clean_commune_df.columns
# Review our Quest dataframe columns
clean_quest_df.columns
# Merge our Commune DF and ou Quest DF to see where most students are most interesting to the Bootcamp
location_merge_df = pd.merge(clean_commune_df[['Commune_Id','Commune_en']],clean_quest_df[['Commune_Id','quest_id']], on="Commune_Id")
location_merge_df.columns
# Creating a new DF
location_df = location_merge_df[['Commune_en']]
location_df.shape
# Top three communes of the country that will be the most likely to expand its training centers.
result_location_df = location_df.value_counts()
result_location_df.head(3)
# **Conclusion:** The top three communes of the country that will be the most likely to expand its training centers are **Delmas, Port-au-Prince, Petion-Ville** because we have more interessed students in these communes
# #### 2. Data vizualisation
result_location_df.plot.bar()
plt.show()
# ## Question 1. How many applications must be made to select 25% women for each on average
# First let's see the percentages we have for our top three communes
for_delmas = result_location_df.Delmas.sum()
for_paup = result_location_df['Port-au-Prince'].sum()
for_pv = result_location_df['Petion-Ville'].sum()
# ## Question 2. What are the most effective communication channels (Alumni, Facebook, WhatsApp, Friend ...) that will allow a student to be susceptible to selection
# #### Data overview
# Verify all the communication channel based on our Question dataframe
communication_channels = clean_quest_df["hear_AA_1"]
communication_channels.shape
final_channels = communication_channels.value_counts()
final_channels.head(5)
# #### Data visualization
final_channels.plot.bar()
# **Conclusion:** Based on all communications the most effective communication channels are: Friend, Whatsapp, Facebook, ESIH, LinkedIn
# ## Question 3. What is the average number of university students who should participate in this program
clean_quest_df.columns
# Let's see how many students we have in our dataframe
uni_students_df = clean_quest_df['education_level']
uni_students_df
count_uni_students = uni_students_df.value_counts()
count_uni_students
# Display only the ones with a university Status
univ_status = uni_students_df.loc[(uni_students_df =='Bachelors (bacc +4)' ) | (uni_students_df=='Masters') | (uni_students_df =='Doctorate (PhD, MD, JD)') ]
univ_status.value_counts()
# Display the total students with university status
result_univ_status = univ_status.value_counts().sum()
print(result_univ_status,"students with university status")
# Display the average
result_univ_status.mean()
# ## Question 4. What will be the average number of applications per week that we could have
clean_quest_df.columns
# Let's create a new DF with date
app_date_df = clean_quest_df['created_at']
app_date_df.head()
# Total number of applications per week we have
per_week = clean_quest_df.groupby(pd.Grouper(freq='W', key='created_at'))['quest_id'].count()
per_week
# Average per week
per_week.mean()
# ## Question 5. How many weeks should we extend the application process to select 60 students per commune?
# ## Question 6. If we were to do all the bootcamp online, who would be the best communes and how many applications would we need to select 30 student and what percentage of students would have a laptop, an internet connection, both at the same time
# > To reply to this question, First we are going to merge our commune dataframe with our question's dataframe and we are going to look in our new data frame to see where all our canditates located.
# #### 2. Merge/join the dataframe
# Review our commun columns
clean_commune_df.columns
# Review our questions columns
clean_quest_df.columns
merge_df = pd.merge(clean_commune_df,clean_quest_df, on="Commune_Id")
new_df = merge_df[["Commune_en","Commune_Id","Departement","have_computer_home","internet_at_home"]]
new_df.head(3)
# #### If all the bootcamp were online, who would be the best communes
# Display all students with a laptop, an internet connection, both at the same time
both_yes = new_df.loc[(new_df['have_computer_home'] == "Yes") & (new_df['internet_at_home'] == "Yes")]
both_yes.head()
best_communes_df = both_yes[['Commune_en','have_computer_home','internet_at_home']]
# Now let's check for the best communes
best_communes = best_communes_df.value_counts()
best_communes.head(5)
# **Conclusion:** If all the bootcamp were online, the best communes would be: **Delmas, Port-au-Prince, Petion-Ville, Carrefour, Tabarre.**
#
# POV: Based on all Minimum technical and software requirements for online courses we have selected all our students with **Computer at home** and **Internet at home** and we look for their location to what communes would be the best if the bootcamp were online
# #### Data visualization
best_communes.plot.bar()
# #### How many applications would we need to select 30 student
# #### Percentage of students would have a laptop, an internet connection, both at the same time
# Select specifie
df_both_yes = both_yes[["have_computer_home","internet_at_home"]]
df_both_yes.head(3)
# Control the some for both
result_both_yes = df_both_yes.value_counts()
result_both_yes
# Sum of total students
total_result_both_yes = result_both_yes.sum()
print("We have",total_result_both_yes,"students with a laptop, and an internet connection")
'''
Now let's give the results in %
'''
percents_both_yes = (total_result_both_yes / 250 * 100).round(2).astype(str) + '%'
print(percents_both_yes,"of students with a laptop, and an internet connection")
# ## Question 7. What are the most effective communication channels (Alumni, Facebook, WhatsApp, Friend ...) that will allow a women to be susceptible to selection
# Let's check our questions dataframe to see what columns we are going to work on
clean_quest_df.head(3)
# Let's make a new dataframe with only our needed columns
new_df_channels = clean_quest_df[["gender","hear_AA_1"]]
# Display our new DF
new_df_channels.head(2)
# Display only Females to see their most effective communication channels
only_females_df_channels = new_df_channels.loc[(new_df_channels["gender"] == "female")]
only_females_df_channels.head(3)
# Overview our new dataframe
row,col = only_females_df_channels.shape
print("We have", row, "Females")
# Now let's check for their most effective communication channels
final_channels = only_females_df_channels.value_counts()
final_channels.head(5)
# #### Data visualization
final_channels.plot.bar()
# **Conclusion:** As we can see thier Top communication channels are **Friends, Whatsapp, Bootcamp Alumni.**
|
data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="hXLFbvLVoi6L"
# # Load the needed data from Foundry.
#
#
# + id="w3h7efGhr_pl" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623804472992, "user_tz": 300, "elapsed": 6612, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="64d200bd-dcc3-4ab8-c0ca-7221be42fc45"
# install the foundry and pymatgen
# !pip install foundry_ml
# !pip install pymatgen
# + id="Iq3tNnQoD3XT" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623804543640, "user_tz": 300, "elapsed": 49625, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="895352e4-5cfa-4da6-ec6c-db2691df19ea"
# initiate the foundry
from foundry import Foundry
f = Foundry(no_browser=True, no_local_server=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 700} id="VDadSOwuSx79" executionInfo={"status": "ok", "timestamp": 1623804559142, "user_tz": 300, "elapsed": 1465, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="d490bda2-f253-402d-ed82-3544d9e86125"
f.list()
# + [markdown] id="Wfs-hbpKElgl"
# Load and clean the experimental data. For clean method details, please refer to the paper
# + colab={"base_uri": "https://localhost:8080/"} id="ls3RjBR1EnGr" executionInfo={"status": "ok", "timestamp": 1622731497941, "user_tz": 300, "elapsed": 46315, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="40f45e30-131f-4ea8-e52b-e2bd03bce254"
import numpy as np
from pymatgen.core import Structure
import pandas as pd
f = f.load('_test_exp_bandgap_v1.1', globus=False)
df_X, df_Y = f.load_data()
N = len(df_Y)
drop_index = []
for i in range(N):
if type(df_Y.loc[i,'bandgap value (eV)']) == list:
new = [float(k) for k in df_Y.loc[i,'bandgap value (eV)']]
mean = np.mean(new)
std = np.std(new)
if abs(mean) > 1.e-6 and std/mean > 0.1:
drop_index.append(i)
else:
df_Y.loc[i,'bandgap value (eV)'] = mean
df1_exp_Y = df_Y.drop(drop_index)
df1_exp_X = df_X.drop(drop_index)
# select 300K, optical, smallest bandgap data
composition = []
sp = []
N1 = len(df1_exp_Y)
for i in range(N1):
str_temp = Structure.from_dict(df1_exp_X.iloc[i]['structure'])
composition.append(str_temp.composition.reduced_formula)
sp.append(str_temp.get_space_group_info()[1])
# composition.append(i['Composition'])
df_300_smallest_X = []
df_300_smallest_Y = []
for com in list({value:"" for value in composition}):
temp = []
for index in range(N1):
if composition[index] == com:
temp.append(index)
spacegroup = []
for index in temp:
spacegroup.append(sp[index])
# print(spacegroup, list(set(spacegroup)))
for sp1 in list({value:"" for value in spacegroup}):
temp1 = []
kinds = []
for index in temp:
if sp[index] == sp1:
temp1.append(index)
string = df1_exp_X.iloc[index]['temp (K)'] + df1_exp_X.iloc[index]['bandgap type'] + df1_exp_X.iloc[index]['exp method']
kinds.append(string)
# print('temp1',temp1)
for case in ['300IO','300I','IO','I','300O','300','O','','300DO','300D','DO','D']:
if case in kinds:
for index in temp1:
string = df1_exp_X.iloc[index]['temp (K)'] + df1_exp_X.iloc[index]['bandgap type'] + df1_exp_X.iloc[index]['exp method']
if string == case:
df_300_smallest_X.append(df1_exp_X.iloc[index])
df_300_smallest_Y.append(df1_exp_Y.iloc[index])
break
else:
continue
print(len(df_300_smallest_X))
df_exp_X = pd.DataFrame(df_300_smallest_X)
df_exp_Y = pd.DataFrame(df_300_smallest_Y)
df_exp_X, df_exp_Y
# + [markdown] id="SUuDo35wET6f"
# Load the materials project PBE data
# + colab={"base_uri": "https://localhost:8080/"} id="RBEFxdtwEWKC" executionInfo={"status": "ok", "timestamp": 1622731566044, "user_tz": 300, "elapsed": 57960, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="90fc3486-956b-4486-ddca-7bbc48abc561"
f = f.load('_test_mp_bandgap_v1.1', globus=False)
df_pbe_X, df_pbe_Y = f.load_data()
# + [markdown] id="0OCl-Ik6HBTN"
# Load other computational data
# + colab={"base_uri": "https://localhost:8080/"} id="EwkrX4gjHVTq" executionInfo={"status": "ok", "timestamp": 1622731615722, "user_tz": 300, "elapsed": 49688, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="1a2c8d2b-f681-4bd2-d98b-87746d4e2a97"
f = f.load('_test_comp_bandgap_v1.1',globus=False)
df_X, df_Y = f.load_data()
# + [markdown] id="Kk2HaiElHaO_"
# Obtain the Jarvis PBE data with OPTB88 (vdW)
# + id="KycEghVCICuV"
df_pbe1_X = df_X[(df_X['comp method'] == 'optb88') ]
df_pbe1_Y = df_Y[(df_X['comp method'] == 'optb88') ]
# + [markdown] id="Vq3po71IIo6l"
# Obtain the data for mbj
# + colab={"base_uri": "https://localhost:8080/"} id="y73hLhQmI5LL" executionInfo={"status": "ok", "timestamp": 1622731824820, "user_tz": 300, "elapsed": 86191, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="52913e9a-e43f-48be-8123-927bd5b79288"
df_temp1_X = df_X[(df_X['comp method'] == 'MBJ') | (df_X['comp method'] == 'mbj')]
df_temp1_Y = df_Y[(df_X['comp method'] == 'MBJ') | (df_X['comp method'] == 'mbj')]
df_temp_X = df_temp1_X[df_temp1_X['bandgap type'] != 'D']
df_temp_Y = df_temp1_Y[df_temp1_X['bandgap type'] != 'D']
df_temp2_X = df_temp_X.copy()
df_temp2_Y = df_temp_Y.copy()
# remove repeated case
composition = []
sp = []
N = len(df_temp_X)
for i in range(N):
str_temp = Structure.from_dict(df_temp_X.iloc[i]['structure'])
composition.append(str_temp.composition.reduced_formula)
sp.append(str_temp.get_space_group_info()[1])
df_no_repeat_X = []
df_no_repeat_Y = []
for com in list({value:"" for value in composition}):
temp = []
for index in range(N):
if composition[index] == com:
temp.append(index)
if len(temp) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp[0]])
else:
spacegroup = []
for index in temp:
spacegroup.append(sp[index])
for sp1 in list({value:"" for value in spacegroup}):
temp1 = []
for index in temp:
if sp[index] == sp1:
temp1.append(index)
if len(temp1) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
else:
values = []
for index in temp1:
values.append(float(df_temp_Y.iloc[index]['bandgap value (eV)']))
mean = np.mean(values)
std = np.std(values)
if abs(mean) < 1.e-10 or abs(std) < 1.e-10:
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
elif std/mean < 0.1:
df_temp2_Y.iloc[temp1[0],df_temp2_Y.columns.get_loc('bandgap value (eV)')] = mean
df_no_repeat_Y.append(df_temp2_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp2_X.iloc[temp1[0]])
df_mbj_X = pd.DataFrame(df_no_repeat_X)
df_mbj_Y = pd.DataFrame(df_no_repeat_Y)
# + [markdown] id="_7aKDky0JRs1"
# Obtain the data for GLLB-SC
# + id="nEFziglAJWS2"
df_temp1_X = df_X[(df_X['comp method'] == 'GLLB_SC') | (df_X['comp method'] == 'GLLB-SC')]
df_temp1_Y = df_Y[(df_X['comp method'] == 'GLLB_SC') | (df_X['comp method'] == 'GLLB-SC')]
df_temp_X = df_temp1_X[df_temp1_X['bandgap type'] != 'D']
df_temp_Y = df_temp1_Y[df_temp1_X['bandgap type'] != 'D']
df_temp2_X = df_temp_X.copy()
df_temp2_Y = df_temp_Y.copy()
# remove repeated case
composition = []
sp = []
N = len(df_temp_X)
for i in range(N):
str_temp = Structure.from_dict(df_temp_X.iloc[i]['structure'])
composition.append(str_temp.composition.reduced_formula)
sp.append(str_temp.get_space_group_info()[1])
df_no_repeat_X = []
df_no_repeat_Y = []
for com in list({value:"" for value in composition}):
temp = []
for index in range(N):
if composition[index] == com:
temp.append(index)
if len(temp) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp[0]])
else:
spacegroup = []
for index in temp:
spacegroup.append(sp[index])
for sp1 in list({value:"" for value in spacegroup}):
temp1 = []
for index in temp:
if sp[index] == sp1:
temp1.append(index)
if len(temp1) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
else:
values = []
for index in temp1:
values.append(float(df_temp_Y.iloc[index]['bandgap value (eV)']))
mean = np.mean(values)
std = np.std(values)
if abs(mean) < 1.e-10 or abs(std) < 1.e-10:
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
elif std/mean < 0.1:
df_temp2_Y.iloc[temp1[0],df_temp2_Y.columns.get_loc('bandgap value (eV)')] = mean
df_no_repeat_Y.append(df_temp2_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp2_X.iloc[temp1[0]])
df_gllb_X = pd.DataFrame(df_no_repeat_X)
df_gllb_Y = pd.DataFrame(df_no_repeat_Y)
# + [markdown] id="-TU8GnkvJb3q"
# Obtain the data for HSE
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="pvvrZFBWJjT4" executionInfo={"status": "ok", "timestamp": 1622732012511, "user_tz": 300, "elapsed": 50855, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="ec1c879d-229a-4ba2-ec31-6c56357e5ed0"
df_temp1_X = df_X[(df_X['comp method'] == 'HSE') | (df_X['comp method'] == 'hse')]
df_temp1_Y = df_Y[(df_X['comp method'] == 'HSE') | (df_X['comp method'] == 'hse')]
df_temp_X = df_temp1_X[df_temp1_X['bandgap type'] != 'D']
df_temp_Y = df_temp1_Y[df_temp1_X['bandgap type'] != 'D']
df_temp2_X = df_temp_X.copy()
df_temp2_Y = df_temp_Y.copy()
# remove repeated case
composition = []
sp = []
N = len(df_temp_X)
for i in range(N):
str_temp = Structure.from_dict(df_temp_X.iloc[i]['structure'])
composition.append(str_temp.composition.reduced_formula)
sp.append(str_temp.get_space_group_info()[1])
df_no_repeat_X = []
df_no_repeat_Y = []
for com in list({value:"" for value in composition}):
temp = []
for index in range(N):
if composition[index] == com:
temp.append(index)
if len(temp) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp[0]])
else:
spacegroup = []
for index in temp:
spacegroup.append(sp[index])
for sp1 in list({value:"" for value in spacegroup}):
temp1 = []
for index in temp:
if sp[index] == sp1:
temp1.append(index)
if len(temp1) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
else:
values = []
for index in temp1:
values.append(float(df_temp_Y.iloc[index]['bandgap value (eV)']))
mean = np.mean(values)
std = np.std(values)
if abs(mean) < 1.e-10 or abs(std) < 1.e-10:
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
elif std/mean < 0.1:
df_temp2_Y.iloc[temp1[0],df_temp2_Y.columns.get_loc('bandgap value (eV)')] = mean
df_no_repeat_Y.append(df_temp2_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp2_X.iloc[temp1[0]])
df_hse_X = pd.DataFrame(df_no_repeat_X)
df_hse_Y = pd.DataFrame(df_no_repeat_Y)
# + [markdown] id="MVlZn4yHJpiC"
# Obtain the data for GW
# + id="KFTfSTQ1JvSn"
df_temp1_X = df_X[(df_X['comp method'] == 'GW') | (df_X['comp method'] == 'GWVD')]
df_temp1_Y = df_Y[(df_X['comp method'] == 'GW') | (df_X['comp method'] == 'GWVD')]
df_temp_X = df_temp1_X[df_temp1_X['bandgap type'] != 'D']
df_temp_Y = df_temp1_Y[df_temp1_X['bandgap type'] != 'D']
df_temp2_X = df_temp_X.copy()
df_temp2_Y = df_temp_Y.copy()
# remove repeated case
composition = []
sp = []
N = len(df_temp_X)
for i in range(N):
str_temp = Structure.from_dict(df_temp_X.iloc[i]['structure'])
composition.append(str_temp.composition.reduced_formula)
sp.append(str_temp.get_space_group_info()[1])
df_no_repeat_X = []
df_no_repeat_Y = []
for com in list({value:"" for value in composition}):
temp = []
for index in range(N):
if composition[index] == com:
temp.append(index)
if len(temp) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp[0]])
else:
spacegroup = []
for index in temp:
spacegroup.append(sp[index])
for sp1 in list({value:"" for value in spacegroup}):
temp1 = []
for index in temp:
if sp[index] == sp1:
temp1.append(index)
if len(temp1) == 1:
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
else:
values = []
for index in temp1:
values.append(float(df_temp_Y.iloc[index]['bandgap value (eV)']))
mean = np.mean(values)
std = np.std(values)
if abs(mean) < 1.e-10 or abs(std) < 1.e-10:
df_no_repeat_Y.append(df_temp_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp_X.iloc[temp1[0]])
elif std/mean < 0.1:
df_temp2_Y.iloc[temp1[0],df_temp2_Y.columns.get_loc('bandgap value (eV)')] = mean
df_no_repeat_Y.append(df_temp2_Y.iloc[temp1[0]])
df_no_repeat_X.append(df_temp2_X.iloc[temp1[0]])
df_gw_X = pd.DataFrame(df_no_repeat_X)
df_gw_Y = pd.DataFrame(df_no_repeat_Y)
# + [markdown] id="8mVapx7VKB9S"
# # Set and initiate the MEGNet model
# + colab={"base_uri": "https://localhost:8080/"} id="1vcSiVvsoykP" executionInfo={"status": "ok", "timestamp": 1622740956655, "user_tz": 300, "elapsed": 6193, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="2bcb4a5d-28fc-4512-a36a-7d791cebadd0"
# install megnet
# !pip install megnet
# !pip install jsonlines
# + [markdown] id="vL9yjN2fqCgj"
# Define data fidelity
# + id="ePVLK6ZNpVe7"
ALL_FIDELITIES = ['pbe','pbe1','mbj', 'gllb-sc', 'hse','gw','exp']
TRAIN_FIDELITIES= ['pbe','pbe1','mbj', 'gllb-sc', 'hse', 'gw','exp']
VAL_FIDELITIES = ['mbj','gllb-sc', 'hse', 'gw','exp']
TEST_FIDELITIES= ['pbe','pbe1','mbj', 'gllb-sc', 'hse', 'gw','exp']
# + [markdown] id="v8I4ZwkYqLtP"
# Import needed modules
# + id="J819vrIvqH6k"
from megnet.callbacks import ReduceLRUponNan, ManualStop
from megnet.data.crystal import CrystalGraph
from megnet.data.graph import GraphBatchDistanceConvert, GaussianDistance
from megnet.models import MEGNetModel
# import jsonlines
import tensorflow as tf
tf.compat.v1.disable_eager_execution()
# + [markdown] id="vI0YEbk4qvd-"
# Initiate the megnet model
# + id="sUXmOIpyql-f" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1622732025447, "user_tz": 300, "elapsed": 3248, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="5b18daa4-c0ea-418a-815b-4647c9d9de6b"
## Set the number of maximum epochs to 1500
EPOCHS = 1500
## Random seed
SEED = 11742
crystal_graph = CrystalGraph(
bond_converter=GaussianDistance(centers=np.linspace(0, 6, 100),
width=0.5),
cutoff=5.0)
model = MEGNetModel(nfeat_edge=100, nfeat_global=None, ngvocal=len(TRAIN_FIDELITIES),
global_embedding_dim=16, nblocks=3, nvocal=95,
npass=2, graph_converter=crystal_graph, lr=1e-3)
# + [markdown] id="GH5_6yJKdhO_"
# Set the structure feature(graph) and target (bandgap value).
# + id="O9CsSyzgdmmG"
graphs = []
targets = []
indexs = []
for fidelity_id, fidelity in enumerate(ALL_FIDELITIES):
if fidelity == 'hse':
N = len(df_hse_X)
df_temp_X = df_hse_X.copy()
df_temp_Y = df_hse_Y.copy()
elif fidelity == 'gw':
N = len(df_gw_X)
df_temp_X = df_gw_X.copy()
df_temp_Y = df_gw_Y.copy()
elif fidelity == 'gllb-sc':
N = len(df_gllb_X)
df_temp_X = df_gllb_X.copy()
df_temp_Y = df_gllb_Y.copy()
elif fidelity == 'exp':
N = len(df_exp_X)
df_temp_X = df_exp_X.copy()
df_temp_Y = df_exp_Y.copy()
elif fidelity == 'pbe':
N = len(df_pbe_X)
df_temp_X = df_pbe_X.copy()
df_temp_Y = df_pbe_Y.copy()
elif fidelity == 'mbj':
N = len(df_mbj_X)
df_temp_X = df_mbj_X.copy()
df_temp_Y = df_mbj_Y.copy()
for index in range(N):
try:
graph = crystal_graph.convert(Structure.from_dict(df_temp_X.iloc[index]['structure']))
graph['state'] = [fidelity_id]
graphs.append(graph)
bd_value = float(df_temp_Y.iloc[index]['bandgap value (eV)'])
targets.append(bd_value)
indexs.append('%s_%s' % (str(index), fidelity))
except:
pass
final_graphs = {i:j for i, j in zip(indexs, graphs)}
final_targets = {i:j for i, j in zip(indexs, targets)}
# + [markdown] id="RoEE16PseJH7"
# Data split
# + id="ic-Wol8MeLEH" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1622732479938, "user_tz": 300, "elapsed": 639, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14643087371112424475"}} outputId="a2ee179b-3958-438a-a7c1-d770461b6c01"
from sklearn.model_selection import train_test_split
## train:val:test = 8:1:1
fidelity_list = [i.split('_')[1] for i in indexs]
train_val_ids, test_ids = train_test_split(indexs, stratify=fidelity_list,
test_size=0.1, random_state=SEED)
fidelity_list = [i.split('_')[1] for i in train_val_ids]
train_ids, val_ids = train_test_split(train_val_ids, stratify=fidelity_list,
test_size=0.1/0.9, random_state=SEED)
## remove pbe from validation
val_ids = [i for i in val_ids if not i.endswith('pbe')]
print("Train, val and test data sizes are ", len(train_ids), len(val_ids), len(test_ids))
# + id="2gY6mAHHeOXS"
## Get the train, val and test graph-target pairs
def get_graphs_targets(ids):
"""
Get graphs and targets list from the ids
Args:
ids (List): list of ids
Returns:
list of graphs and list of target values
"""
ids = [i for i in ids if i in final_graphs]
return [final_graphs[i] for i in ids], [final_targets[i] for i in ids]
train_graphs, train_targets = get_graphs_targets(train_ids)
val_graphs, val_targets = get_graphs_targets(val_ids)
# + [markdown] id="Kah789ezeT4w"
# Model training
# + id="MIc3XkJbeST9" colab={"base_uri": "https://localhost:8080/"} outputId="3c137955-09b5-4cc1-8be0-a4b27104bf37"
callbacks = [ReduceLRUponNan(patience=500), ManualStop()]
model.train_from_graphs(train_graphs, train_targets, val_graphs, val_targets,
epochs=EPOCHS, verbose=2, initial_epoch=0, callbacks=callbacks)
|
Notebook/Bandgap_model_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Code Samples From Large Group
#
# +
import pandas as pd
dataset = "/home/jovyan/datasets/ist256/UCDP-Georeferenced-Event-Dataset/v21.1/ged211.csv"
df = pd.read_csv(dataset)
# -
confs = df[ ["year","code_status","conflict_name" ] ]
confs[ confs.year == 1989 ]
# +
stocks = pd.read_json("https://raw.githubusercontent.com/mafudge/advanced-databases/main/datasets/json-samples/stocks.json")
stocks
# -
bigstocks = stocks[ stocks['price'] > 1000 ]
bigstocks
for s in bigstocks.to_records():
print(f"{s['symbol']} is a big stock!")
stocks
students = [
{'name' : 'mike', 'gpa' : 2.1},
{'name' : 'dan', 'gpa' : 2.7},
{'name' : 'deb', 'gpa' : 3.4}
]
student_df = pd.DataFrame(students)
student_df
people = [
{'name' : 'mike', 'age' : 100},
{'name' : 'dan', 'age' : 50},
{'name' : 'deb', 'credit_score' : 880}
]
people_df = pd.DataFrame(people)
people_df
people_df.columns
df.columns
df.columns
df[ ["year","country","latitude", "longitude"] ]
student_df
student_df['gpa'] < 3.0
student_df[ student_df['gpa'] < 3.0 ]
# +
from IPython.display import display, HTML
from ipywidgets import interact_manual
import pandas as pd
dataset = "/home/jovyan/datasets/ist256/UCDP-Georeferenced-Event-Dataset/v21.1/ged211.csv"
df = pd.read_csv(dataset)
countries = list(df["country"].unique())
years = list(df['year'].unique())
@interact_manual( country = countries, year = years )
def onclick(country,year):
filtered_df = df[ (df["country"] == country) & (df['year'] == year) ]
cols_df = filtered_df[["year","country","latitude", "longitude"]]
display(cols_df)
# -
DATA FRAME OPERATORS
& AND
| OR
~ NOT
|
lessons/12-pandas/LargeGroup.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/djvaroli/samsung_oct/blob/Janhavi-Colab-Notebooks/50per_train_data_JG_oct__2_model_SimCLR.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="IOHjctPzkGDD" outputId="83e55589-c1ac-4f01-b94b-3b7a8f3ef9b6"
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
import os
os.chdir('/content/drive/MyDrive/Colab Notebooks/Samsung-OCT-Project-Work/')
pwd = os.getcwd()
print(pwd)
# + colab={"base_uri": "https://localhost:8080/"} id="wPisAhxbuGpo" outputId="66e7063f-c21c-45f7-b21b-35928aae4905"
os.chdir('/content/drive/MyDrive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/')
# %mkdir train_data_50per
# %cd train_data_50per
# %mkdir 0
# %mkdir 1
# %mkdir 2
# %mkdir 3
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="3qqLOUZUugnK" outputId="6cb31b29-99a5-40b5-b28e-669a057bc815"
# %pwd
# + id="atFDJpWjt6vp"
import pandas as pd
import numpy as np
def load_data(folder):
X=[]
Y=[]
Z=[]
for folderName in os.listdir(folder):
if not folderName.startswith('.'):
if folderName == '0':
label = 0
elif folderName == '1':
label = 1
elif folderName == '2':
label = 2
elif folderName == '3':
label = 3
else:
label = 4
for file in os.listdir(folder+'/'+folderName):
if not file.startswith('.'):
X.append(file)
Y.append(label)
Z.append(folder+'/'+folderName+'/'+file)
return pd.DataFrame(np.hstack((np.asarray(X).reshape(-1,1),np.asarray(Z).reshape(-1,1), np.asarray(Y).reshape(-1,1))),columns=['filename','path','Class']).sort_values('Class')
###############################
# + id="mEgEGLU8xLy3"
train_data = load_data('/content/drive/My Drive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/train/')
# + id="o-PKUYkuwo3p"
# Creating reduced train dataset
# Selecting smaller sample size
def sample_data(data, sample_size):
if sample_size == data.shape[0]:
return data
class_weights = dict(data["Class"].value_counts()/data.shape[0])
print("Before sub-sampling=", class_weights)
df0 = data[data["Class"]=='0'].sample(n=min(int(round(sample_size * class_weights['0'])),len(data[data["Class"]=='0'])))
df1 = data[data["Class"]=='1'].sample(n=min(int(round(sample_size * class_weights['1'])),len(data[data["Class"]=='1'])))
df2 = data[data["Class"]=='2'].sample(n=min(int(round(sample_size * class_weights['2'])),len(data[data["Class"]=='2'])))
df3 = data[data["Class"]=='3'].sample(n=min(int(round(sample_size * class_weights['3'])),len(data[data["Class"]=='3'])))
return pd.concat([df0,df1,df2,df3])
# + colab={"base_uri": "https://localhost:8080/"} id="0J7-AAfVws6v" outputId="79423208-6485-4d2c-8284-662c7932fcf0"
train_50per = int((1/50)*(train_data.shape[0]))
print("50% reduced train data sample size=",train_50per)
train_data_red = sample_data(train_data,train_50per)
class_weights = dict(train_data_red["Class"].value_counts()/train_data_red.shape[0])
print("Sub-sampled train data=", class_weights)
print(train_data.shape, train_data_red.shape)
# + id="o7ZEco1at7so"
# Copying to 50 per train data
import shutil
train_files = train_data_red.path.to_list()
for filenum in range(len(train_files)):
x = train_files[filenum]
x_split = x.split('/')
filename = x_split[-1]
separator = '/'
x_path = separator.join(x_split[:-1])
#print(x_path)
os.chdir(x_path)
if '0' in x_path:
#print(x_path)
shutil.copy(filename, '../../train_data_50per/0/')
elif '1' in x_path:
shutil.copy(filename, '../../train_data_50per/1/')
elif '2' in x_path:
shutil.copy(filename, '../../train_data_50per/2/')
else:
shutil.copy(filename, '../../train_data_50per/3/')
# + colab={"base_uri": "https://localhost:8080/"} id="lcGWHBLhxBf-" outputId="915796e0-dab2-4194-e443-861462635c73"
os.chdir('/content/drive/My Drive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/train_data_50per/')
# !ls ./*/* | wc -l
os.chdir('/content/drive/My Drive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/test/')
# !ls ./*/* | wc -l
os.chdir('/content/drive/My Drive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/val/')
# !ls ./*/* | wc -l
# + [markdown] id="HbpMmt6uMl0Z"
# # Load Dataframe
# + [markdown] id="G-yBcycIcNYK"
# #JG: Start execution
# + id="Ri8myIeI4l3T"
os.chdir('/content/drive/MyDrive/Colab Notebooks/simclr_tf_keras_07022021/oct_run/50per_train_data')
# + colab={"base_uri": "https://localhost:8080/"} id="q0dvE0YZANZm" outputId="073feece-233f-444f-fcb1-ccb6d9e4b4a2"
# !pip install keras==2.3.1
# !pip install tensorflow==2.1.0
# !pip install opencv-python==4.2.0.32
# !pip install scikit-learn==0.23.1
# !pip install scipy==1.4.1
# !pip install DateTime==4.3
# + id="hsi65mRTMl0G"
# %ls
# %cd SimCLRv1-keras-tensorflow/
# #!pip install -r requirements.txt
# + colab={"base_uri": "https://localhost:8080/", "height": 69} id="IcnnSU90iSFK" outputId="231af54f-c2ae-4e49-fe84-2a34f8a00671"
# %ls
# %cd SimCLRv1-keras-tensorflow/
# %pwd
# + id="YqzIOfEZMl0R"
import numpy as np
import pickle
import pandas as pd
from tensorflow import keras
from sklearn.model_selection import train_test_split
from tensorflow.keras.applications.vgg16 import VGG16
from evaluate_features import get_features, linear_classifier, tSNE_vis
# + colab={"base_uri": "https://localhost:8080/"} id="PVbgtOvpd5py" outputId="ecb7ad32-1d00-4630-acd7-0cc17e777e65"
train_data = pd.read_pickle('/content/drive/My Drive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/train_data_50per_resize/train_data_50per_df.pickle')
train_data.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="CkeNPZ-xfVB1" outputId="a60943a0-bddd-4485-de51-62a658762657"
train_data.loc[train_data['class_label']=='3']
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="aEdGXl1weQC6" outputId="b13a9e2a-6185-4145-de6f-ddb0ccbe1580"
test_data = pd.read_pickle("/content/drive/My Drive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/test_resize/test_df.pickle")
test_data.loc[test_data['class_label']=='2']
# + id="rnx-GJqHIU2K"
#NOT needed
'''for i in range(len(test_data.class_label)):
if test_data.class_label[i] == '0':
#print(i,test_data.class_label[i])
test_data.class_one_hot[i] = [1,0,0,0]
#print(i,test_data.class_one_hot[i])
if test_data.class_label[i] == '1':
#print(i,test_data.class_label[i])
test_data.class_one_hot[i] = [0,1,0,0]
#print(i,test_data.class_one_hot[i])
if test_data.class_label[i] == '2':
#print(i,test_data.class_label[i])
test_data.class_one_hot[i] = [0,0,1,0]
#print(i,test_data.class_one_hot[i])
if test_data.class_label[i] == '3':
#print(i,test_data.class_label[i])
test_data.class_one_hot[i] = [0,0,0,1]
#print(i,test_data.class_one_hot[i]) '''
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="QgDhTPgPf3yu" outputId="7c713aa6-9317-4530-eca2-212ac220911e"
val_data = pd.read_pickle("/content/drive/My Drive/Colab Notebooks/Samsung-OCT-Project-Work/processed_data/data/val_resize/val_df.pickle")
val_data.loc[val_data['class_label']=='3']
# + colab={"base_uri": "https://localhost:8080/"} id="c-YZpNjEMl0g" outputId="40dc0f2c-d332-4090-91c7-10ba584e86e3"
class_labels = ["0", "1", "2", "3"]
num_classes = len(train_data['class_one_hot'][0])
print("# of training instances:", len(train_data.index), "\n")
for label in class_labels:
print(f"# of '{label}' training instances: {(train_data.class_label == label).sum()}")
# + colab={"base_uri": "https://localhost:8080/"} id="Ew6ItQW9aYKS" outputId="199c18aa-2b49-4a8f-8e2f-13b5e92b471c"
df_train = train_data
df_val = val_data
df_test = test_data
print(df_train.shape,df_test.shape,df_val.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="d2qU1NriMl0n" outputId="02701283-4742-41f6-b33c-e7cda92de1a7"
#Not executing these 2 lines since we already have data splitted in train/val/test sets.
#Testing with train data only
df_train, df_val_test = train_test_split(df_train, test_size=0.30, random_state=42, shuffle=True)
df_val, df_test = train_test_split(df_val_test, test_size=0.50, random_state=42, shuffle=True)
print("# of training instances:", len(df_train.index), "\n")
for label in class_labels:
print(f"# of '{label}' training instances: {(df_train.class_label == label).sum()}")
print()
print("# of validation instances:", len(df_val.index), "\n")
for label in class_labels:
print(f"# of '{label}' training instances: {(df_val.class_label == label).sum()}")
print()
print("# of test instances:", len(df_test.index), "\n")
for label in class_labels:
print(f"# of '{label}' training instances: {(df_test.class_label == label).sum()}")
dfs = {
"train": df_train,
"val": df_val,
"test": df_test
}
# + id="N9GgCItGMl0u"
# Img size
size = 80
height_img = size
width_img = size
input_shape = (height_img, width_img, 3)
# + [markdown] id="yYrLvS3iMl0y"
# # Load pretrained VGG16 & Feature evaluation
# + colab={"base_uri": "https://localhost:8080/"} id="4z587NS5Ml00" outputId="9311b12c-a920-4aee-a6ab-a7c1bc2b018c"
params_vgg16 = {'weights': "imagenet",
'include_top': False,
'input_shape': input_shape,
'pooling': None}
# Design model
base_model = VGG16(**params_vgg16)
base_model.summary()
# + id="XcdE2JU5Ml04"
feat_dim = 2 * 2* 512
# + colab={"base_uri": "https://localhost:8080/"} id="ngpgI6FsE4tM" outputId="8065086f-751c-48c5-d5e7-40c804cb231a"
# !pip install keras==2.3.1
# + id="SSYpY2bmFfvR"
import tensorflow
from tensorflow.keras.applications.vgg16 import preprocess_input
# + id="nqlPYBJKjiGD"
from tensorflow import keras
from DataGeneratorSimCLR import DataGeneratorSimCLR as DataGenerator
# + [markdown] id="75j95YbIMl09"
# # Build SimCLR-Model
# + id="pNq93i59Ml0-"
#from DataGeneratorSimCLR import DataGeneratorSimCLR as DataGenerator
from SimCLR import SimCLR
#if it errors out, execute cell # 28 : pip install keras==2.3.1
# + [markdown] id="b0_nz95NMl1D"
# ### Properties
# + id="EqkkdfRPMl1E"
batch_size =32
# Projection_head
num_layers_ph = 2
feat_dims_ph = [2048, 128]
num_of_unfrozen_layers = 1 #Note: with 1, all weights of the base_model are still frozen (last layer is max_pool)
save_path = 'models/trashnet'
# + id="X1dGZztKMl1K" colab={"base_uri": "https://localhost:8080/"} outputId="8764cd83-a40b-462b-f1ca-79aea28a4537"
SimCLR = SimCLR(
base_model = base_model,
input_shape = input_shape,
batch_size = batch_size,
feat_dim = feat_dim,
feat_dims_ph = feat_dims_ph,
num_of_unfrozen_layers = num_of_unfrozen_layers,
save_path = save_path
)
# + colab={"base_uri": "https://localhost:8080/"} id="fGizWCnA_TkF" outputId="ed023cd5-5a9b-47ac-df5c-8d9d91616bbf"
print(SimCLR)
# + id="bvdVxJTiMl1W"
params_generator = {'batch_size': batch_size,
'shuffle' : True,
'width':width_img,
'height': height_img,
'VGG': True
}
# Generators
data_train = DataGenerator(df_train.reset_index(drop=True), **params_generator)
data_val = DataGenerator(df_val.reset_index(drop=True), subset = "val", **params_generator) #val keeps the unity values on the same random places ~42
data_test = DataGenerator(df_test.reset_index(drop=True), subset = "test", **params_generator) #test keeps the unity values on the diagonal
# + colab={"base_uri": "https://localhost:8080/"} id="L4uLn_lcLdvV" outputId="376122fa-58c1-49dc-cc06-c0ff5b00d106"
data_train.height
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="BQhOPioiuw8Y" outputId="b4a15e64-5aa7-4518-b08b-a5f3369ea11c"
data_train.df
# + id="-ipobvp7Ml1a"
y_predict_test_before = SimCLR.predict(data_test)
# + [markdown] id="3XrbFJGsMl1g"
# # SimCLR - Round 1: Only Projection head
# + [markdown] id="pssY8CS6Ml1g"
# ## Training SimCLR
# + colab={"base_uri": "https://localhost:8080/"} id="JLMsqpJcMl1h" outputId="f354f218-6427-422f-f4de-fd3e2e31176d"
SimCLR.train(data_train, data_val, epochs = 5)
# + id="YfWiM4gfMl1n"
y_predict_test_after = SimCLR.predict(data_test)
# + [markdown] id="MITBpHs6Ml1r"
# ## SimCLR-output check
# + colab={"base_uri": "https://localhost:8080/"} id="M92VNCQHMl1s" outputId="f043a5dd-96df-4888-a8c6-1ce612861288"
print(f"Random guess accuracy: {round(1/(2*batch_size),4)}")
print(f"accuracy - test - before: {np.round(np.sum(data_test[0][1] * y_predict_test_before[:batch_size])/(2*batch_size),2)}")
print(f"accuracy - test - after: {np.round(np.sum(data_test[0][1] * y_predict_test_after[:batch_size])/(2*batch_size),2)}")
# + colab={"base_uri": "https://localhost:8080/"} id="K7NxSEoyMl1w" outputId="78960273-f35f-44e8-cae1-79780221021f"
print("y_predict_test_before")
for i in range(min(batch_size, 15)):
print(np.round(y_predict_test_before[i][i],2), end=" | ")
print("\n")
print("y_predict_test_after")
for i in range(min(batch_size, 15)):
print(np.round(y_predict_test_after[i][i],2), end=" | ")
print("\n")
print("y_predict_test_after - Second diagonal")
for i in range(min(batch_size, 15)):
print(np.round(y_predict_test_after[i + 2 * batch_size][i],2), end=" | ")
print("\n")
# + [markdown] id="K_k39St9Ml1z"
# ## Feature Evaluation
#
# Note that this evaluation corresponds with the unaltered pretrained weights
# + id="IOXEi_YVMl10"
fractions = [1.0, 0.2, 0.05]
# + [markdown] id="_36LmVuuMl15"
# ### Logistic regression evaluation
# + colab={"base_uri": "https://localhost:8080/"} id="yDgFplaQMl16" outputId="06efdfd2-c177-4004-cf12-53bb9bc9f732"
features_train, y_train, feats = get_features(base_model, df_train, class_labels)
features_test, y_test, feats = get_features(base_model, df_test, class_labels)
np.count_nonzero(features_train[0])
# + colab={"base_uri": "https://localhost:8080/"} id="lRR_M3xGMl1-" outputId="bcd82286-0faf-46b0-80e4-448111c02838"
# Training logistic regression classifier on 3 fractions of the data
# Optimal regularization is determined from a 5-fold cross-validation
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
linear_classifier(features_train, y_train, features_test, y_test, class_labels, fraction = fraction)
# + [markdown] id="3BT4MD5bMl2G"
# ### Fine tuned model
# + id="bWJlfGi5Ml2G"
batch_size_classifier = 32
params_generator_classifier = {'max_width':width_img,
'max_height': height_img,
'num_classes': num_classes,
'VGG': True
}
params_training_classifier = {'1.0':{
"reg_dense" : 0.005,
"reg_out" : 0.005,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 5e-5],
"epochs" : [5, 5, 15, 10]
},
'0.2':{
"reg_dense" : 0.075,
"reg_out" : 0.01,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 5e-5],
"epochs" : [5, 5, 20, 15]
},
'0.05':{
"reg_dense" : 0.01,
"reg_out" : 0.02,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 1e-5],
"epochs" : [5, 5, 20, 15]
}
}
# + colab={"base_uri": "https://localhost:8080/"} id="Z_IONDPzMl2J" outputId="0cf3ab1a-3427-41eb-b5da-b182f5d6f930"
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
SimCLR.train_NL_and_evaluate(dfs = dfs,
batch_size = batch_size_classifier,
params_generator = params_generator_classifier,
fraction = fraction,
class_labels = class_labels,
reg_dense = params_training_classifier[str(fraction)]["reg_dense"],
reg_out = params_training_classifier[str(fraction)]["reg_out"],
nums_of_unfrozen_layers = params_training_classifier[str(fraction)]["nums_of_unfrozen_layers"],
lrs = params_training_classifier[str(fraction)]["lrs"],
epochs = params_training_classifier[str(fraction)]["epochs"],
verbose_epoch = 0,
verbose_cycle = 0
)
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="YOD7ahLaMl2Q" outputId="fb8009a0-cb24-4e16-8bf3-aaf984363ebd"
tSNE_vis(df_train, features_train, class_labels)
# + [markdown] id="7uyPJiScMl2U"
# # SimCLR - Round 2: Unfreeze last convolutional layer
# + [markdown] id="CNKDhiABMl2V"
# ## Training SimCLR
# + colab={"base_uri": "https://localhost:8080/"} id="r54NABsoMl2V" outputId="ed6a137c-dd40-4643-9135-7f84d7fe6ee3"
#Unfreeze
SimCLR.unfreeze_and_train(data_train, data_val, num_of_unfrozen_layers = 2, r = 2, lr = 1e-5, epochs = 5)
# + [markdown] id="W0ZSWi5EMl2Y"
# ## Feature Evaluation
# + [markdown] id="1OXphSXTMl2Y"
# ### Logisitic regression
# + id="CtO5h6u2Ml2Z"
base_model = SimCLR.base_model
fractions = [1.0, 0.2, 0.05]
# + colab={"base_uri": "https://localhost:8080/"} id="48mY8eOgMl2c" outputId="45899161-1823-4a9c-ab2b-ea3ddb49b5e6"
features_train, y_train, feats = get_features(base_model, df_train, class_labels)
features_test, y_test, feats = get_features(base_model, df_test, class_labels)
np.count_nonzero(features_train[0])
# + colab={"base_uri": "https://localhost:8080/"} id="0UnlErH6Ml2f" outputId="692bfb50-51eb-4cf2-c6d6-7710b131b12d"
# Training logistic regression classifier on 3 fractions of the data
# Optimal regularization is determined from a 5-fold cross-validation
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
linear_classifier(features_train, y_train, features_test, y_test, class_labels, fraction = fraction)
# + [markdown] id="AMwQBHz1Ml2k"
# ### Fine tuned model
# + id="pyxa7SJZMl2k"
batch_size_classifier = 32
params_generator_classifier = {'max_width':width_img,
'max_height': height_img,
'num_classes': num_classes,
'VGG': True
}
params_training_classifier = {'1.0':{
"reg_dense" : 0.005,
"reg_out" : 0.005,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 5e-5],
"epochs" : [5, 5, 15, 10]
},
'0.2':{
"reg_dense" : 0.075,
"reg_out" : 0.01,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 5e-5],
"epochs" : [5, 5, 20, 15]
},
'0.05':{
"reg_dense" : 0.01,
"reg_out" : 0.02,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 1e-5],
"epochs" : [5, 5, 20, 15]
}
}
# + colab={"base_uri": "https://localhost:8080/"} id="de23l-B6Ml2m" outputId="a0348a1b-89ea-4985-8be0-2f8009544bad"
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
SimCLR.train_NL_and_evaluate(dfs = dfs,
batch_size = batch_size_classifier,
params_generator = params_generator_classifier,
fraction = fraction,
class_labels = class_labels,
reg_dense = params_training_classifier[str(fraction)]["reg_dense"],
reg_out = params_training_classifier[str(fraction)]["reg_out"],
nums_of_unfrozen_layers = params_training_classifier[str(fraction)]["nums_of_unfrozen_layers"],
lrs = params_training_classifier[str(fraction)]["lrs"],
epochs = params_training_classifier[str(fraction)]["epochs"],
verbose_epoch = 0,
verbose_cycle = 0
)
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="Z9VQvF9yMl2s" outputId="b061fff2-72c5-43bc-bfbe-4bebbc38884e"
tSNE_vis(df_train, features_train, class_labels)
# + [markdown] id="_Bkh50_zMl2v"
# # SimCLR - Round 3: Unfreeze 2 last convolutional layer
# + [markdown] id="KgjG_EyBMl2w"
# ## Training SimCLR
# + colab={"base_uri": "https://localhost:8080/"} id="8srBtciWMl2y" outputId="b168e6e7-5ea4-4fc4-aa11-f2a66b1bcc1b"
#Unfreeze
SimCLR.unfreeze_and_train(data_train, data_val, num_of_unfrozen_layers = 3, r = 3, lr = 5e-6, epochs = 5)
# + id="TIvT7J9lMl21"
y_predict_test_after = SimCLR.predict(data_test)
# + [markdown] id="fR_iD6kjMl23"
# ## Feature Evaluation
# + id="HfCtfu-FMl24"
base_model = SimCLR.base_model
# + colab={"base_uri": "https://localhost:8080/"} id="QCVop5DtMl26" outputId="1c876439-3f09-4190-d214-4fac8fcc4810"
features_train, y_train, feats = get_features(base_model, df_train, class_labels)
features_test, y_test, feats = get_features(base_model, df_test, class_labels)
np.count_nonzero(features_train[0])
# + colab={"base_uri": "https://localhost:8080/"} id="t8ufdXeRMl28" outputId="9c38ca5b-84ec-43a4-f751-954fd4394059"
# Training logistic regression classifier on 3 fractions of the data
# Optimal regularization is determined from a 5-fold cross-validation
fractions = [1.0, 0.2, 0.05]
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
linear_classifier(features_train, y_train, features_test, y_test, class_labels, fraction = fraction)
# + [markdown] id="5AVOuKXlMl2-"
# ### Fine tuned model
# + id="ABGWyLgSMl2_"
batch_size_classifier = 32
params_generator_classifier = {'max_width':width_img,
'max_height': height_img,
'num_classes': num_classes,
'VGG': True
}
# + colab={"base_uri": "https://localhost:8080/"} id="_sHcT6tiMl3B" outputId="4d1b7da3-c805-4287-8c9e-0bd31f147b02"
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
SimCLR.train_NL_and_evaluate(dfs = dfs,
batch_size = batch_size_classifier,
params_generator = params_generator_classifier,
fraction = fraction,
class_labels = class_labels,
reg_dense = 0.005,
reg_out = 0.003,
nums_of_unfrozen_layers = [5, 5, 6, 7],
lrs = [1e-3, 1e-4, 5e-5, 1e-5],
epochs = [5, 5, 15, 10],
verbose_epoch = 0,
verbose_cycle = 0
)
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="TfrOqkyIMl3E" outputId="b398ed44-35b1-4dd6-9c8b-9e017cf80f9b"
tSNE_vis(df_train, features_train, class_labels)
# + [markdown] id="wgd8co1ZMl3H"
# # SimCLR - Round 4: Unfreeze 3 last convolutional layer
# + [markdown] id="_T2CPs44L7Wj"
# #JG: Achieved quite good classification as seen above. Step not needed ; it leads to NaN features.
# + id="H85sA9-jMl3H"
y_predict_test_before = y_predict_test_after
# + [markdown] id="TbTY6SdtMl3K"
# ## Training SimCLR
# + colab={"base_uri": "https://localhost:8080/"} id="dbj61XF1Ml3K" outputId="e08224e3-2e80-4ea3-aa72-cda145d1e6cc"
#Unfreeze
SimCLR.unfreeze_and_train(data_train,
data_val,
num_of_unfrozen_layers = 4,
r = 4,
lr = 1e-6,
epochs = 5)
# + id="DWApei37Ml3O"
y_predict_test_after = SimCLR.predict(data_test)
# + [markdown] id="IMdwX9kXMl3R"
# ## Feature Evaluation
# + [markdown] id="pO4paMl9Ml3R"
# ### Logistic Regression
# + id="P-IHH1xVMl3S"
base_model = SimCLR.base_model
# + colab={"base_uri": "https://localhost:8080/"} id="cnECT4QJMl3T" outputId="46cb84fc-b87c-4f20-fcce-11a9fdf30547"
features_train, y_train, feats = get_features(base_model, df_train, class_labels)
features_test, y_test, feats = get_features(base_model, df_test, class_labels)
np.count_nonzero(features_train[0])
# + colab={"base_uri": "https://localhost:8080/"} id="L9jr7LcF6d79" outputId="32bb606b-f044-449a-cbb5-334b4bedc4d5"
features_train_nzero = (features_train[np.nonzero(features_train)]).reshape([1168,2048])
features_test_nzero = features_test[np.nonzero(features_test)]
y_train_nzero = y_train[np.nonzero(y_train)]
y_test_nzero = y_test[np.nonzero(y_test)]
print(y_train_nzero.shape, y_train.shape, y_test_nzero.shape, y_test.shape, '\n')
print(features_train.shape)
print(features_test.shape)
print(features_test_nzero.shape)
print(features_train_nzero.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 374} id="TU8NPJBkMl3X" outputId="7dd98137-47f1-4562-a6c6-a91348fdc84f"
# Training logistic regression classifier on 3 fractions of the data
# Optimal regularization is determined from a 5-fold cross-validation
fractions = [1.0, 0.2, 0.05]
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
linear_classifier(features_train, y_train, features_test, y_test, class_labels, fraction = fraction)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="ubK-UPEB8a1S" outputId="bc74e80e-310d-48d5-9439-507657278626"
# Training logistic regression classifier on 3 fractions of the data
# Optimal regularization is determined from a 5-fold cross-validation
fractions = [1.0, 0.2, 0.05]
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
linear_classifier(features_train, y_train_nzero, features_test, y_test_nzero, class_labels, fraction = fraction)
# + [markdown] id="XlVebWE7Ml3Z"
# ### Fine tuned model
# + id="vhnYkrhZMl3a"
batch_size_classifier = 32
params_generator_classifier = {'max_width':width_img,
'max_height': height_img,
'num_classes': num_classes,
'VGG': True
}
params_training_classifier = {'1.0':{
"reg_dense" : 0.005,
"reg_out" : 0.005,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 5e-5],
"epochs" : [5, 5, 15, 10]
},
'0.2':{
"reg_dense" : 0.075,
"reg_out" : 0.01,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 5e-5],
"epochs" : [5, 5, 20, 15]
},
'0.05':{
"reg_dense" : 0.01,
"reg_out" : 0.02,
"nums_of_unfrozen_layers" : [5, 5, 6, 7],
"lrs" : [1e-3, 1e-4, 5e-5, 1e-5],
"epochs" : [5, 5, 20, 15]
}
}
# + colab={"base_uri": "https://localhost:8080/", "height": 425} id="QtoW3bL4Ml3g" outputId="768a9b6c-060c-4c6a-e925-957e0a5b430b"
for fraction in fractions:
print(f" ==== {fraction * 100}% of the training data used ==== \n")
SimCLR.train_NL_and_evaluate(dfs = dfs,
batch_size = batch_size_classifier,
params_generator = params_generator_classifier,
fraction = fraction,
class_labels = class_labels,
reg_dense = params_training_classifier[str(fraction)]["reg_dense"],
reg_out = params_training_classifier[str(fraction)]["reg_out"],
nums_of_unfrozen_layers = params_training_classifier[str(fraction)]["nums_of_unfrozen_layers"],
lrs = params_training_classifier[str(fraction)]["lrs"],
epochs = params_training_classifier[str(fraction)]["epochs"],
verbose_epoch = 0,
verbose_cycle = 0
)
# + id="PAyCxLZXMl3j"
tSNE_vis(df_train, features_train, class_labels)
# + id="RItzibZvMl3l" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="288e9d00-f3b7-4fc5-8bee-7cd9c77e7a35"
os.getcwd()
# + [markdown] id="wrDAbEICODF3"
# # JG: Load pre-trained best performaing SimCLR based model trained with 1168 images only and make predictions
# + id="mAN4YBLCN2Du"
import tensorflow as tf
import json
import os
from tensorflow import keras
from tensorflow.keras.models import load_model
# + colab={"base_uri": "https://localhost:8080/"} id="nYSfHh6GPkQy" outputId="3b4d4282-f458-4cb7-d4fb-d10732301c4a"
os.chdir('/content/drive/My Drive/Colab Notebooks/simclr_tf_keras_07022021/oct_run/50per_train_data/SimCLRv1-keras-tensorflow/models/trashnet/SimCLR')
# %ls
# + colab={"base_uri": "https://localhost:8080/", "height": 459} id="vPtCLYZCQYfl" outputId="50f21cd2-2347-4941-ec58-8c4a8208fd9d"
# !pip install 'h5py==2.10.0' --force-reinstall
# + id="wNvo4rS3RJUk"
os.chdir('/content/drive/My Drive/Colab Notebooks/simclr_tf_keras_07022021/oct_run/50per_train_data/SimCLRv1-keras-tensorflow')
# + colab={"base_uri": "https://localhost:8080/"} id="VJLda2OhRzoo" outputId="2e4ae6a1-49fe-411e-bbee-c562c1a3264e"
# !pip install keras==2.3.1
# + id="kqenISG7RYt1"
from SimCLR import SimCLR
# + colab={"base_uri": "https://localhost:8080/"} id="qjlvK809Of-m" outputId="8afc121c-deb1-454f-ae32-4d49325a8de6"
#model = load_model('/content/drive/My Drive/Colab Notebooks/simclr_tf_keras_07022021/oct_run/50per_train_data/SimCLRv1-keras-tensorflow/models/trashnet/SimCLR/SimCLR_07_04_22h_42.h5')
model = load_model('/content/drive/My Drive/Colab Notebooks/simclr_tf_keras_07022021/oct_run/50per_train_data/SimCLRv1-keras-tensorflow/models/trashnet/base_model/base_model_round_3.h5')
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="ebAkNmkfUMqc" outputId="0cadd6f8-051f-4985-c9d1-c5ec7e476146"
os.getcwd()
# + id="Ka2YwlxDXLeH"
#from swish import Swish
#from SoftmaxCosineSim import SoftmaxCosineSim
# + colab={"base_uri": "https://localhost:8080/"} id="Qh7-389YSbqJ" outputId="1ee513cb-22a5-4acf-c06e-7637cb11cfed"
model2 = load_model('/content/drive/My Drive/Colab Notebooks/simclr_tf_keras_07022021/oct_run/50per_train_data/SimCLRv1-keras-tensorflow/models/trashnet/SimCLR/SimCLR_07_04_22h_42.h5', custom_objects={'SoftmaxCosineSim':SoftmaxCosineSim,'Swish':Swish})
model2.summary()
# + id="KcqmJym0b2jT"
|
50per_train_data_JG_oct__2_model_SimCLR.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Structured data prediction using Vertex AI Platform
#
#
# ## Learning Objectives
#
# 1. Create a BigQuery Dataset and Google Cloud Storage Bucket
# 2. Export from BigQuery to CSVs in GCS
# 3. Training on Cloud AI Platform
# 4. Deploy trained model
#
# ## Introduction
#
# In this notebook, you train, evaluate, and deploy a machine learning model to predict a baby's weight.
#
#
# + colab={} colab_type="code" id="Nny3m465gKkY"
# !sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# -
# !pip install --user google-cloud-bigquery==2.26.0
# **Note**: Restart your kernel to use updated packages.
# Kindly ignore the deprecation warnings and incompatibility errors related to google-cloud-storage.
# + [markdown] colab_type="text" id="hJ7ByvoXzpVI"
# ## Set up environment variables and load necessary libraries
# -
# Set environment variables so that we can use them throughout the entire notebook. We will be using our project name for our bucket, so you only need to change your project and region.
# + deletable=true editable=true
# change these to try this notebook out
BUCKET = 'qwiklabs-gcp-02-2372fbdc4b9d' # Replace with the your bucket name
PROJECT = 'qwiklabs-gcp-02-2372fbdc4b9d' # Replace with your project-id
REGION = 'us-central1'
# +
import os
from google.cloud import bigquery
# -
os.environ["PROJECT"] = PROJECT
os.environ["BUCKET"] = BUCKET
os.environ["REGION"] = REGION
os.environ["TFVERSION"] = "2.6"
os.environ["PYTHONVERSION"] = "3.7"
# + language="bash"
# export PROJECT=$(gcloud config list project --format "value(core.project)")
# echo "Your current GCP Project Name is: "$PROJECT
# + [markdown] colab_type="text" id="L0-vOB4y2BJM"
# ## The source dataset
#
# Our dataset is hosted in [BigQuery](https://cloud.google.com/bigquery/). The CDC's Natality data has details on US births from 1969 to 2008 and is a publically available dataset, meaning anyone with a GCP account has access. Click [here](https://console.cloud.google.com/bigquery?project=bigquery-public-data&p=publicdata&d=samples&t=natality&page=table) to access the dataset.
#
# The natality dataset is relatively large at almost 138 million rows and 31 columns, but simple to understand. `weight_pounds` is the target, the continuous value we’ll train a model to predict.
# -
# ## Create a BigQuery Dataset and Google Cloud Storage Bucket
#
# A BigQuery dataset is a container for tables, views, and models built with BigQuery ML. Let's create one called __babyweight__. We'll do the same for a GCS bucket for our project too.
# + language="bash"
#
# # Create a BigQuery dataset for babyweight if it doesn't exist
# datasetexists=$(bq ls -d | grep -w babyweight)
#
# if [ -n "$datasetexists" ]; then
# echo -e "BigQuery dataset already exists, let's not recreate it."
#
# else
# echo "Creating BigQuery dataset titled: babyweight"
#
# bq --location=US mk --dataset \
# --description "Babyweight" \
# $PROJECT:babyweight
# echo "Here are your current datasets:"
# bq ls
# fi
#
# ## Create GCS bucket if it doesn't exist already...
# exists=$(gsutil ls -d | grep -w gs://${BUCKET}/)
#
# if [ -n "$exists" ]; then
# echo -e "Bucket exists, let's not recreate it."
#
# else
# echo "Creating a new GCS bucket."
# gsutil mb -l ${REGION} gs://${BUCKET}
# echo "Here are your current buckets:"
# gsutil ls
# fi
# + [markdown] colab_type="text" id="b2TuS1s9vREL"
# ## Create the training and evaluation data tables
#
# Since there is already a publicly available dataset, we can simply create the training and evaluation data tables using this raw input data. First we are going to create a subset of the data limiting our columns to `weight_pounds`, `is_male`, `mother_age`, `plurality`, and `gestation_weeks` as well as some simple filtering and a column to hash on for repeatable splitting.
#
# * Note: The dataset in the create table code below is the one created previously, e.g. "babyweight".
# -
# ### Preprocess and filter dataset
#
# We have some preprocessing and filtering we would like to do to get our data in the right format for training.
#
# Preprocessing:
# * Cast `is_male` from `BOOL` to `STRING`
# * Cast `plurality` from `INTEGER` to `STRING` where `[1, 2, 3, 4, 5]` becomes `["Single(1)", "Twins(2)", "Triplets(3)", "Quadruplets(4)", "Quintuplets(5)"]`
# * Add `hashcolumn` hashing on `year` and `month`
#
# Filtering:
# * Only want data for years later than `2000`
# * Only want baby weights greater than `0`
# * Only want mothers whose age is greater than `0`
# * Only want plurality to be greater than `0`
# * Only want the number of weeks of gestation to be greater than `0`
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_data AS
SELECT
weight_pounds,
CAST(is_male AS STRING) AS is_male,
mother_age,
CASE
WHEN plurality = 1 THEN "Single(1)"
WHEN plurality = 2 THEN "Twins(2)"
WHEN plurality = 3 THEN "Triplets(3)"
WHEN plurality = 4 THEN "Quadruplets(4)"
WHEN plurality = 5 THEN "Quintuplets(5)"
END AS plurality,
gestation_weeks,
FARM_FINGERPRINT(
CONCAT(
CAST(year AS STRING),
CAST(month AS STRING)
)
) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND weight_pounds > 0
AND mother_age > 0
AND plurality > 0
AND gestation_weeks > 0
# ### Augment dataset to simulate missing data
#
# Now we want to augment our dataset with our simulated babyweight data by setting all gender information to `Unknown` and setting plurality of all non-single births to `Multiple(2+)`.
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_augmented_data AS
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks,
hashmonth
FROM
babyweight.babyweight_data
UNION ALL
SELECT
weight_pounds,
"Unknown" AS is_male,
mother_age,
CASE
WHEN plurality = "Single(1)" THEN plurality
ELSE "Multiple(2+)"
END AS plurality,
gestation_weeks,
hashmonth
FROM
babyweight.babyweight_data
# ### Split augmented dataset into train and eval sets
#
# Using `hashmonth`, apply a module to get approximately a 75/25 train-eval split.
# #### Split augmented dataset into train dataset
# + colab={} colab_type="code" id="CMNRractvREL"
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_data_train AS
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks
FROM
babyweight.babyweight_augmented_data
WHERE
ABS(MOD(hashmonth, 4)) < 3
# -
# #### Split augmented dataset into eval dataset
# %%bigquery
CREATE OR REPLACE TABLE
babyweight.babyweight_data_eval AS
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks
FROM
babyweight.babyweight_augmented_data
WHERE
ABS(MOD(hashmonth, 4)) = 3
# + [markdown] colab_type="text" id="clnaaqQsXkwC"
# ## Verify table creation
#
# Verify that you created the dataset and training data table.
# -
# %%bigquery
-- LIMIT 0 is a free query; this allows us to check that the table exists.
SELECT * FROM babyweight.babyweight_data_train
LIMIT 0
# %%bigquery
-- LIMIT 0 is a free query; this allows us to check that the table exists.
SELECT * FROM babyweight.babyweight_data_eval
LIMIT 0
# ## Export from BigQuery to CSVs in GCS
#
# Use BigQuery Python API to export our train and eval tables to Google Cloud Storage in the CSV format to be used later for TensorFlow/Keras training. We'll want to use the dataset we've been using above as well as repeat the process for both training and evaluation data.
# +
# Construct a BigQuery client object.
client = bigquery.Client()
dataset_name = "babyweight"
# Create dataset reference object
dataset_ref = client.dataset(
dataset_id=dataset_name, project=client.project)
# Export both train and eval tables
for step in ["train", "eval"]:
destination_uri = os.path.join(
"gs://", BUCKET, dataset_name, "data", "{}*.csv".format(step))
table_name = "babyweight_data_{}".format(step)
table_ref = dataset_ref.table(table_name)
extract_job = client.extract_table(
table_ref,
destination_uri,
# Location must match that of the source table.
location="US",
) # API request
extract_job.result() # Waits for job to complete.
print("Exported {}:{}.{} to {}".format(
client.project, dataset_name, table_name, destination_uri))
# -
# ## Verify CSV creation
#
# Verify that we correctly created the CSV files in our bucket.
# + language="bash"
# gsutil ls gs://${BUCKET}/babyweight/data/*.csv
# -
# ## Check data exists
#
# Verify that you previously created CSV files we'll be using for training and evaluation.
# + language="bash"
# gsutil ls gs://${BUCKET}/babyweight/data/*000000000000.csv
# -
# ## Training on Cloud AI Platform
#
# Now that we see everything is working locally, it's time to train on the cloud!
# To submit to the Cloud we use [`gcloud ai-platform jobs submit training [jobname]`](https://cloud.google.com/sdk/gcloud/reference/ml-engine/jobs/submit/training) and simply specify some additional parameters for AI Platform Training Service:
# - jobname: A unique identifier for the Cloud job. We usually append system time to ensure uniqueness
# - job-dir: A GCS location to upload the Python package to
# - runtime-version: Version of TF to use.
# - python-version: Version of Python to use.
# - region: Cloud region to train in. See [here](https://cloud.google.com/ml-engine/docs/tensorflow/regions) for supported AI Platform Training Service regions
#
# Below the `-- \` we add in the arguments for our `task.py` file.
# + language="bash"
#
# OUTDIR=gs://${BUCKET}/babyweight/trained_model
# JOBID=babyweight_$(date -u +%y%m%d_%H%M%S)
#
# gcloud ai-platform jobs submit training ${JOBID} \
# --region=${REGION} \
# --module-name=trainer.task \
# --package-path=$(pwd)/babyweight/trainer \
# --job-dir=${OUTDIR} \
# --staging-bucket=gs://${BUCKET} \
# --master-machine-type=n1-standard-8 \
# --scale-tier=CUSTOM \
# --runtime-version=${TFVERSION} \
# --python-version=${PYTHONVERSION} \
# -- \
# --train_data_path=gs://${BUCKET}/babyweight/data/train*.csv \
# --eval_data_path=gs://${BUCKET}/babyweight/data/eval*.csv \
# --output_dir=${OUTDIR} \
# --num_epochs=10 \
# --train_examples=10000 \
# --eval_steps=100 \
# --batch_size=32 \
# --nembeds=8
# -
# The training job should complete within 15 to 20 minutes. You do not need to wait for this training job to finish before moving forward in the notebook, but will need a trained model.
# ## Check our trained model files
#
# Let's check the directory structure of our outputs of our trained model in folder we exported. We'll want to deploy the saved_model.pb within the timestamped directory as well as the variable values in the variables folder. Therefore, we need the path of the timestamped directory so that everything within it can be found by Cloud AI Platform's model deployment service.
# + language="bash"
# gsutil ls gs://${BUCKET}/babyweight/trained_model
# + language="bash"
# MODEL_LOCATION=$(gsutil ls -ld -- gs://${BUCKET}/babyweight/trained_model/2* \
# | tail -1)
# gsutil ls ${MODEL_LOCATION}
# -
# ## Deploy trained model
#
# Deploying the trained model to act as a REST web service is a simple gcloud call.
# + language="bash"
# gcloud config set ai_platform/region global
#
# + language="bash"
# MODEL_NAME="babyweight"
# MODEL_VERSION="ml_on_gcp"
# MODEL_LOCATION=$(gsutil ls -ld -- gs://${BUCKET}/babyweight/trained_model/2* \
# | tail -1 | tr -d '[:space:]')
# echo "Deleting and deploying $MODEL_NAME $MODEL_VERSION from $MODEL_LOCATION"
# # gcloud ai-platform versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
# # gcloud ai-platform models delete ${MODEL_NAME}
# gcloud ai-platform models create ${MODEL_NAME} --regions ${REGION}
# gcloud ai-platform versions create ${MODEL_VERSION} \
# --model=${MODEL_NAME} \
# --origin=${MODEL_LOCATION} \
# --runtime-version=2.6 \
# --python-version=3.7
# -
# Copyright 2021 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
courses/machine_learning/deepdive2/production_ml/babyweight/train_deploy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # "Transfer Learning with TensorFlow : Feature Extraction"
# > "Notebook demonstrates Transfer Learning using Feature Extraction in TensorFlow"
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [Deep Learning, NeuralNetworks, TensorFlow, Transfer-Learning, Feature-Extraction]
# - image: images/efficientnet.png
# + [markdown] id="Zds_xieR_GXe"
# ## Transfer Learning with TensorFlow : Feature Extraction
# > This Notebook is an account of my working for the Udemy course : [TensorFlow Developer Certificate in 2022: Zero to Mastery](https://www.udemy.com/course/tensorflow-developer-certificate-machine-learning-zero-to-mastery/).
#
# Transfer learning is a machine learning method where a model developed for a task is reused as the starting point for a model on a second task.
#
# It is a popular approach in deep learning where pre-trained models are used as the starting point on computer vision and natural language processing tasks given the vast compute and time resources required to develop neural network models on these problems and from the huge jumps in skill that they provide on related problems.
# + [markdown] id="2hpdfZcxK643"
# Concepts covered in this Notebook:
# * Introduce transfer learning with TensorFlow
# * Using a small dataset to experiment faster(10% of training samples)
# * Building a transfer learning feature extraction model with TensorFlow Hub.
# * Use TensorBoard to track modelling experiments and results
# + [markdown] id="2MnCCap9HUsl"
# **Why use transfer learning?**
#
# * Can leverage an existing neural network architecture **proven to work** on problems similar to our own
# * Can leverage a working network architecture which has **already learned patterns** on similar data to our own (often results in great ML products with less data)
#
# + [markdown] id="eELe13wLHUvv"
# Example of tranfer learning use cases:
# * Computer Vision (using ImageNet for our Image classification problems)
# * Natural Language Processing (detecting spam mails - spam filter)
# + [markdown] id="em6PbYNrHUni"
# ### Downloading and getting familiar with data
#
# + colab={"base_uri": "https://localhost:8080/"} id="5M2Q8-YUHUet" outputId="d7ec14cc-d74b-464e-c578-215befe14617"
# Get data (10% of 10 food classes from food101 dataset from kaggle)
import zipfile
# Download the data
# !wget https://storage.googleapis.com/ztm_tf_course/food_vision/10_food_classes_10_percent.zip
# Unzip the downloaded file
zip_ref = zipfile.ZipFile("10_food_classes_10_percent.zip")
zip_ref.extractall()
zip_ref.close()
# + id="g9dnRKR5HUTR"
# How many images in each folder
import os
# Walk through 10 percent data dir and list number of files
for dirpath, dirnames, filenames in os.walk("10_food_classes_10_percent)"):
print(f"There are {len(dirnames)} directories and {len(filenames)} images in '{dirpath}'.")
# + [markdown] id="hsPC9AG9HUP8"
# ### Creating data loader (preparing the data)
#
# We'll use the `ImageDataGenerator` class to load in our images in batches.
#
# + colab={"base_uri": "https://localhost:8080/"} id="5MTnMPNdHUK6" outputId="71ef886d-d094-42b4-a5a0-cdd52ecbd886"
# Setup data inputs
from tensorflow.keras.preprocessing.image import ImageDataGenerator
IMAGE_SHAPE = (224,224)
BATCH_SIZE = 32
train_dir = "10_food_classes_10_percent/train"
test_dir = "10_food_classes_10_percent/test"
train_datagen = ImageDataGenerator(rescale=1/255.)
test_datagen = ImageDataGenerator(rescale = 1/255.)
print("Training images:")
train_data_10_percent = train_datagen.flow_from_directory(train_dir,
target_size = IMAGE_SHAPE,
batch_size = BATCH_SIZE,
class_mode = "categorical")
print("Testing Images:")
test_data = test_datagen.flow_from_directory(test_dir,
target_size = IMAGE_SHAPE,
batch_size = 32,
class_mode ="categorical")
# + [markdown] id="BMu8qiDoQx54"
# ### Setting up callbacks (things to run whilst our model trains)
#
# Callbacks are extra functionality you can add to your models to be performed during or after training. Some of the most popular callbacks:
#
# * Tracking experiments with the `TensorBoard` callback
# * Model checkpoint with the `ModelCheckpoint` callback
# * Stopping a model from training (before it trains too long and overfits) with the `EarlyStopping` callback
#
# Some popular callbacks include:
#
# | Callback name | Use case | Code |
# | --- | --- | --- |
# | TensorBoard | Log the performance of multiple models and then view and compare these models in a visual way on TensorBoard (a dashboard for inspecting neural network parameters). Helpful to compare the results of different models on your data | `tf.keras.callbacks.TensorBoard()` |
# | Model checkpointing | Save your model as it trains so you can stop training if needed and come back to continue off where you left. Helpful if training takes a long time and can't be done in one sitting | `tf.keras.callbacks.ModelCheckpoint()` |
# | Early Stopping | Leave your model training for an arbitary amount of time and have it stop training automatically when it ceases to improve. Helpful when you've got a large dataset and don't know how long training will take | `tf.keras.callbacks.EarlyStopping()`
# + id="E077l4rGXUPT"
# Create TensorBoard callback (functionized because we need to create a new one for each model)
import datetime
import tensorflow as tf
def create_tensorboard_callback(dir_name, experiment_name):
log_dir = dir_name + "/" + experiment_name + "/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir = log_dir)
print(f"Saving TensorBoard log files to : {log_dir}")
return tensorboard_callback
# + [markdown] id="ImRmpsbLZ0e8"
# > **Note:** You can customize the directory where your TensorBoard logs (model training metrics) get saved to whatever you like. The `log_dir` parameter we've created above is only one option.
# + [markdown] id="W_mFpBK0amsy"
# ### Creating models using Tensorflow Hub
#
# TensorFlow Hub is the repository of pre-trained machine learning models created with tensorflow.
#
# We are going to do a similar process, except the majority of our model's layers are going to come from TensorFlow Hub.
#
# [TensorFlow Hub](https://www.tensorflow.org/hub)
# + id="fstRn16QatVY"
# Let's compare the following two models
efficientnet_url = "https://tfhub.dev/google/efficientnet/b0/classification/1"
resnet_url = "https://tfhub.dev/google/imagenet/resnet_v2_50/feature_vector/5"
# + id="LaIZDn6datSZ"
# Import dependencies
import tensorflow as tf
import tensorflow_hub as hub
from tensorflow.keras import layers
# + id="EaWQzoKoasKV"
# Let's make a create_model() function to create a model from a URL
def create_model(model_url, num_classes = 10):
"""
Takes a TensorFlow Hub URL and creates a Keras Sequential model with it
Args:
model_url(str): A TensorFlow Hub features extraction URL.
num_classes(int) : Number of output neurons in the output layer,
should be equal to number of target classes, default 10.
Returns:
An uncompiled Keras Sequential model with model_url as feature extractor
layers and Dense output layers with num_classes output neurons.
"""
# Download the pretrained model and save it
feature_extractor_layer = hub.KerasLayer(model_url,
trainable =False,
name = "feature_extraction_layer",
input_shape = IMAGE_SHAPE+(3,)) # Freeze the already learned parameters
# Create our model
model = tf.keras.Sequential([
feature_extractor_layer,
layers.Dense(num_classes, activation = "softmax", name = "output_layer")
])
return model
# + [markdown] id="6xAjYjIqasHt"
# ### Creating and testing ResNet TensorFlow Hub Feature Extraction model
# + id="HC8DDXO-asFO"
# Create Resnet model
resnet_model = create_model(resnet_url,
num_classes=train_data_10_percent.num_classes)
# + id="QAFNpqr3g3ud"
resnet_model.compile(loss = "categorical_crossentropy",
optimizer = tf.keras.optimizers.Adam(),
metrics = ["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="92WvUoW4asC_" outputId="38257b44-1012-4b96-c4ae-92f503e81c8b"
resnet_model.summary()
# + [markdown] id="Ir9JgJbcgoWl"
# whoaa..the number of parameters are ~23 Million. But only ~20,000 are trainable parameters other are from the freeze model we uploaded from the tensorflow hub. Here's look at the Resnet50 architecture:
#
# 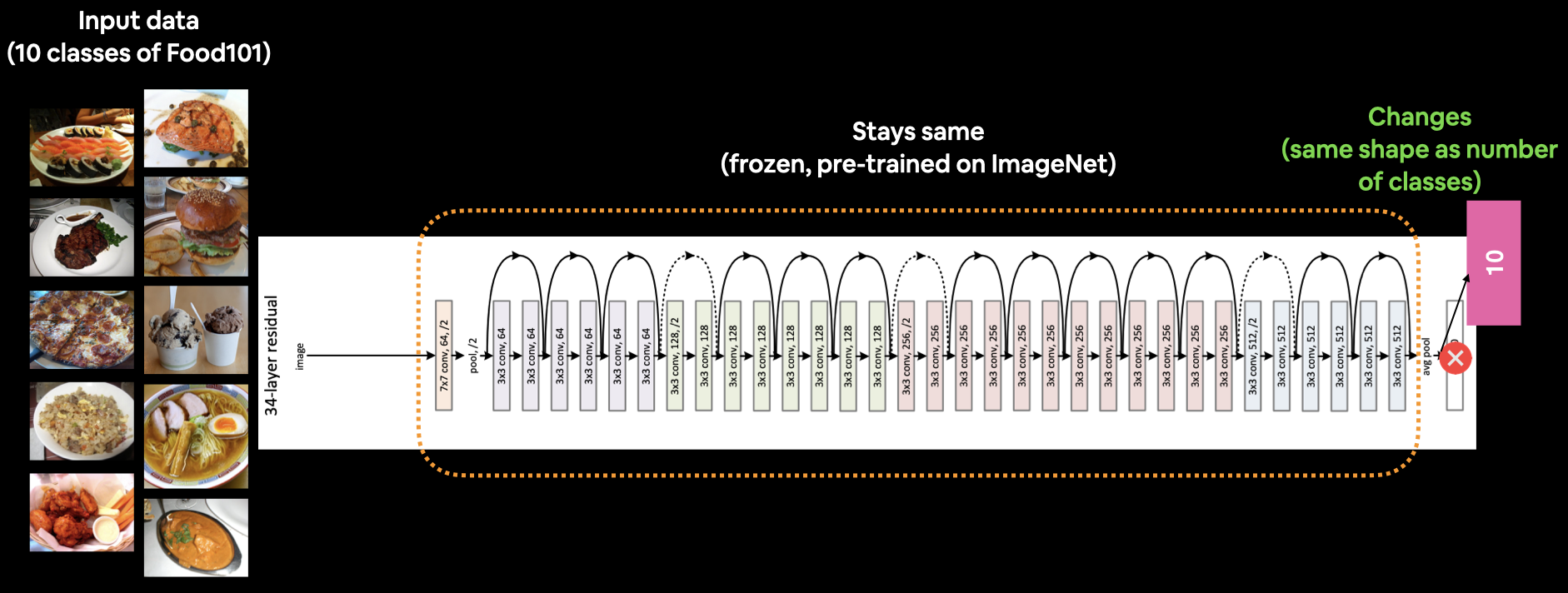
# *What our current model looks like. A ResNet50V2 backbone with a custom dense layer on top (10 classes instead of 1000 ImageNet classes). **Note:** The Image shows ResNet34 instead of ResNet50. **Image source:** https://arxiv.org/abs/1512.03385.*
#
# + colab={"base_uri": "https://localhost:8080/"} id="jPrWoWN1hD21" outputId="b35e84a0-718a-4bf6-feec-5c8645d85d8f"
# Fit the resnet model to our 10% of the data
resnet_history = resnet_model.fit(train_data_10_percent,
epochs = 5,
steps_per_epoch =len(train_data_10_percent),
validation_data = test_data,
validation_steps = len(test_data),
callbacks = [create_tensorboard_callback(dir_name="tensorflow_hub",
experiment_name ="resnet50V2")])
# + [markdown] id="m0O4naTjrliM"
# Our transfer learning feature extractor model outperformed all of the previous models we built by hand. We have achieved around ~91% accuracy!! with only 10% of the data. we have a validation accuracy ~77%
# + id="c1fTVUvPasAI"
# let's create a function to plot our loss curves
# we can put a function like this into a script called "helper.py and import "
import matplotlib.pyplot as plt
plt.style.use('dark_background') # set dark background for plots
def plot_loss_curves(history):
"""
Returns training and validation metrics
Args:
history : TensorFlow History object.
Returns:
plots of training/validation loss and accuracy metrics
"""
loss = history.history["loss"]
val_loss = history.history["val_loss"]
accuracy = history.history["accuracy"]
val_accuracy = history.history["val_accuracy"]
epochs = range(len(history.history["loss"]))
# Plot loss
plt.plot(epochs, loss, label = "training_loss")
plt.plot(epochs, val_loss, label = "validation_loss")
plt.title("Loss")
plt.xlabel("Epochs")
plt.legend()
# Plot Accuracy
plt.figure()
plt.plot(epochs, accuracy, label ="training_accuracy")
plt.plot(epochs, val_accuracy, label = "validation_accuracy")
plt.title("Accuracy")
plt.xlabel("Accuracy")
plt.legend();
# + colab={"base_uri": "https://localhost:8080/", "height": 573} id="JYepR4ynar9I" outputId="46f0f329-e93b-413a-cb24-e339876472b7"
plot_loss_curves(resnet_history)
# + [markdown] id="mstf-FFlar6q"
# ### Creating and testing EfficientNetB0 TensorFlow Hub Feature Extraction model
# + colab={"base_uri": "https://localhost:8080/"} id="VK8mkElCar4O" outputId="d8e19724-2478-4274-fc43-a1a5d184a977"
# Create Efficient model
efficientnet_model = create_model(model_url=efficientnet_url,
num_classes=train_data_10_percent.num_classes)
# Compile the EfficientNet model
efficientnet_model.compile(loss = "categorical_crossentropy",
optimizer = tf.keras.optimizers.Adam(),
metrics = ["accuracy"])
# Fit EfficientNet model to 10% of training data
efficientnet_history = efficientnet_model.fit(train_data_10_percent,
epochs =5,
steps_per_epoch =len(train_data_10_percent),
validation_data = test_data,
validation_steps = len(test_data),
callbacks = [create_tensorboard_callback(dir_name ="tensorflow_hub",
experiment_name="efficientnetb0")])
# + [markdown] id="iI-n7IZD27-A"
# We have done really well with EfficientNetb0 model. we have training accuracy ~91% and validation accuracy ~84%.
# + colab={"base_uri": "https://localhost:8080/", "height": 573} id="_L2uqfRtar1Q" outputId="797cdee2-f566-444e-cb54-070584dc48cc"
# Let's plot the loss curves
plot_loss_curves(efficientnet_history)
# + colab={"base_uri": "https://localhost:8080/"} id="bCs-CXDA2mzz" outputId="3fcb12e7-1e4a-4c92-b69c-9eab1c28fb9e"
efficientnet_model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="ZlthUDkF3N5r" outputId="cd4fca46-8dcf-4e08-b499-7933c1efc876"
resnet_model.summary()
# + [markdown] id="y5ZmBHru3TO5"
# The EfficientNet architecture performs really good even though the number of parameters in the model are far less than ResNet model.
# + colab={"base_uri": "https://localhost:8080/"} id="PunZSzsg5XnI" outputId="9a47f406-a791-4e4b-d8ea-07c172ab4951"
len(efficientnet_model.layers[0].weights)
# + [markdown] id="3wherogl5O-u"
# our Neural network learns these weights/parameters for extracting and generalizing features for better prediction on new data
# + [markdown] id="NA1pe2f83fVu"
# ### Different types of transfer learning
#
# * **"As is" transfer learning** - using an existing model with no changes(eg. using ImageNet model on 1000 ImageNet classes, none of your own)
# * **"Feature extraction"** transfer learning - use the pre-learned patterns of an existing model (eg. EffiientNetB0 trained on ImageNet) and adjust the output layer for your own problem(eg. 1000 classes -> 10 classes of food)
# * **"Fine tuning" transfer learning** - use the prelearned patterns of an existing model and "fine-tune" many or all of the underlying layers(including new output layers)
#
# 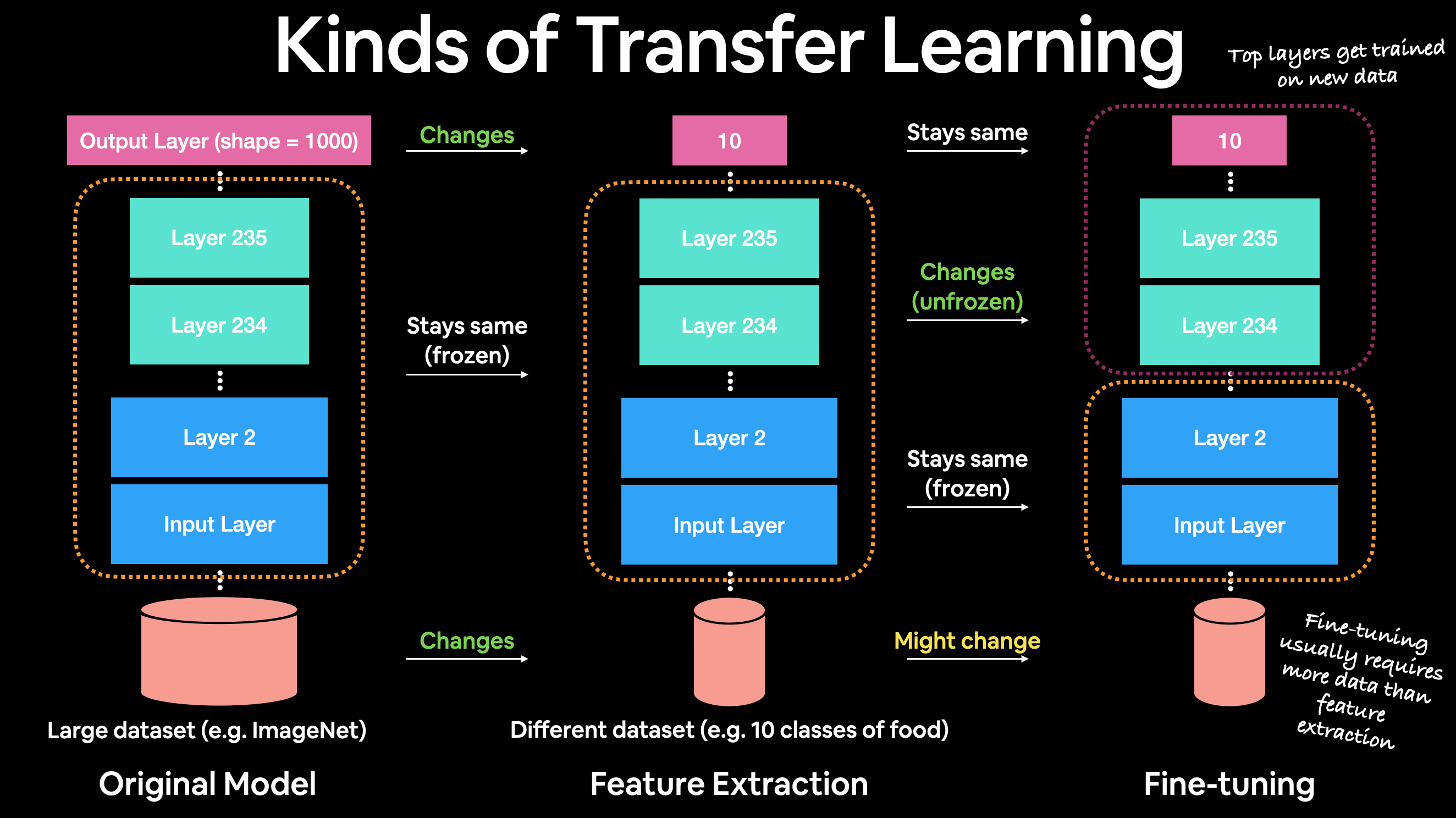
# *The different kinds of transfer learning. An original model, a feature extraction model (only top 2-3 layers change) and a fine-tuning model (many or all of original model get changed).*
# + [markdown] id="YhX0eNTH3fTE"
# ### Comparing our model's results using TensorBoard
#
# **What is TensorBoard?**
# * A way to visually explore your machine learning models performance and internals.
# * Host, track and share your machine learning experiments on [TensorBoard.dev](https://tensorboard.dev/)
#
# > **Note:** When you upload things to TensorBoard.dev, your experiments are public. So, if you're running private experiments (things you don't want other to see) do not upload them to TensorBoard.dev
# + id="B0JnLPpA3fOk"
# Run thie code cell for uploading the experiment to TensorBoard
# Upload TensorBoard dev records
# !tensorboard dev upload --logdir ./tensorflow_hub/ \
# --name "EfficientNetB0 vs. ResNet50V2" \
# --description "Comparing two different TF Hub Feature extraction model architectures using 10% of the training data" \
# --one_shot
# + colab={"base_uri": "https://localhost:8080/"} id="As7cGJDY3fMC" outputId="24ffb9da-085a-4aa7-d859-5f5963296af3"
#Check out what TensorBoard experiments you have
# !tensorboard dev list
# + id="HRoFtMGC3fKI"
# Delete an experiment
# # !tensorboard dev delete --experiment_id [copy your id here]
# + [markdown] id="Yw-BS-TN3SaZ"
# Weights&Biases also integreates with TensorFlow so we can use that as visualization tool.
# + [markdown] id="KfBwE_UK_wqy"
# ## References:
#
# * [weights&bisases](https://wandb.ai/site)
# * [TensorBoard experiment](https://tensorboard.dev/experiment/1OBT56EnQ7GbJSOdsOKjZg/)
# * [TensorFlow Developer Certificate in 2022: Zero to Mastery](https://www.udemy.com/course/tensorflow-developer-certificate-machine-learning-zero-to-mastery/)
|
_notebooks/2022-02-18-Transfer-Learning-in-TensorFlow-Feature-Extraction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 ('ml_Test')
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df=pd.read_csv(r"C:\Users\HP\OneDrive\Desktop\ml\modular\ml-class\Data\wine_fraud.csv")
df.head()
df["quality"].unique()
df["type"].unique()
sns.countplot(x="quality",data=df)
# +
#showing that it is highly imbalanced dataset
# -
sns.countplot(x="type",data=df,hue="quality")
reds=df[df["type"]=="red"]
whites=df[df["type"]=="white"]
# +
len(whites[whites['quality'] == 'Fraud'])/len(whites)*100
# -
len(reds[reds["quality"]=="Fraud"])/len(reds)*100
df["Fraud"]=df["quality"].map({"Legit":0,"Fraud":1})
df.head()
df.corr()["Fraud"].sort_values()
plt.figure(dpi =150)
df.corr()['Fraud'][: -1].sort_values().plot(kind = 'bar')
plt.figure(figsize=(11,8),dpi =150)
sns.heatmap(df.corr(), cmap = 'viridis', annot = True)
df=df.drop("Fraud",axis=1)
df.head()
df['type'] = pd.get_dummies(df['type'], drop_first = True) #Encoding
x = df.drop('quality', axis =1)
y = df['quality']
# +
# TRAIN TEST SPLIT
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.1, random_state=101)
# SCALE DATA
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled_x_train = scaler.fit_transform(X_train)
scaled_x_test = scaler.transform(X_test)
# -
from sklearn.svm import SVC
# +
svc = SVC(class_weight = 'balanced')
# -
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [0.001, 0.01, 0.1, 0.5,1], 'kernel': ['linear', 'rbf', 'sigmoid', 'poly']}
# +
grid_model =GridSearchCV(svc, param_grid)
# +
grid_model.fit(scaled_x_train, y_train)
# -
grid_model.best_params_
# +
from sklearn.metrics import confusion_matrix, classification_report, plot_confusion_matrix
# +
grid_preds = grid_model.predict(scaled_x_test)
# +
confusion_matrix(y_test, grid_preds)
# +
plot_confusion_matrix(grid_model, scaled_x_test, y_test)
# +
print(classification_report(y_test, grid_preds))
|
SVM_Project/SVM_wine.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
# example text for model training (SMS messages)
simple_train = ['call you tonight', 'Call me a cab', 'please call me... PLEASE!']
from sklearn.feature_extraction.text import CountVectorizer
vect = CountVectorizer()
# learn the 'vocabulary' of the training data (occurs in-place)
vect.fit(simple_train)
# examine the fitted vocabulary
vect.get_feature_names()
# +
# transform training data into a 'document-term matrix'
# row = document , Column = terms
# here we have 3 documents & 6 terms, so makes it a
# 3*6 matrics# convert sparse matrix to a dense matrix
simple_train_dtm = vect.transform(simple_train)
simple_train_dtm
# -
# convert sparse matrix to a dense matrix
# in "sparse matrix" all of the value initially conserder
# as zero, so when we get a value it stores only the
# co-ordinate of non-zero values but in "dense matrix"
# zeros & non-zeros both have the presence thus occupy
# more memory
simple_train_dtm.toarray()
pd.DataFrame(simple_train_dtm.toarray(), columns=vect.get_feature_names())
# check the type of the document-term matrix
type(simple_train_dtm)
# examine the sparse matrix contents
print(simple_train_dtm)
# example text for model tesing
simple_test = [" please don't call me "]
vect.transform(simple_test)
# transform testing data into a document-term matrix (using existing vocabulary)
simple_test_dtm = vect.transform(simple_test)
simple_test_dtm.toarray()
# examine the vocabulary and document-term matrix together
pd.DataFrame(simple_test_dtm.toarray(), columns=vect.get_feature_names())
# +
#==================================================
#==================================================
#==================================================
# -
# alternative: read file into pandas from a URL
url = 'https://raw.githubusercontent.com/justmarkham/pycon-2016-tutorial/master/data/sms.tsv'
sms = pd.read_table(url, header=None, names=['label', 'message'])
# examine the shape
sms.shape
# examine the first 10 rows
sms.head()
# examine the class distribution
sms.label.value_counts()
# convert label to a numerical variable
sms['label_num'] = sms.label.map({'ham':0, 'spam':1})
sms.head(10)
# how to define X and y (from the SMS data) for use with COUNTVECTORIZER
X = sms.message
y = sms.label_num
print(X.shape)
print(y.shape)
# split X & y into training & testing sets
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,random_state=1)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print( y_test.shape )
# instantiate the vectorizer
vect = CountVectorizer()
# learn training data vocabulary, then use it to creat "document term matrix"'
vect.fit(X_train)
X_train_dtm = vect.transform(X_train)
# examine the dtm matrix
X_train_dtm
# transform testing data (using fitter vocabulary) into dtm
X_test_dtm = vect.transform(X_test)
X_test_dtm
# import & instantiate a Multinomial Naive Bayes Model
from sklearn.naive_bayes import MultinomialNB
nb = MultinomialNB()
# %time nb.fit(X_train_dtm, y_train)
# we are using X_train_dtm as it's mathmatical
# transformation of X_train
y_pred_class = nb.predict(X_test_dtm)
from sklearn import metrics
metrics.accuracy_score(y_test, y_pred_class)
metrics.confusion_matrix(y_test,y_pred_class)
# +
# [ true_neg false_pos
# false_neg true_pos ]
# -
# print message text for false positive (ham incorrectly specified)
X_test[ y_pred_class > y_test ]
#find false negative
X_test[ y_pred_class < y_test ]
# example of false negative
X_test[3132]
nb.predict_proba(X_test_dtm)
# array([ [ probability of being 1 probability of being 0]])
y_pred_prob = nb.predict_proba(X_test_dtm)[:,1]
y_pred_prob
# calculate AUC
metrics.roc_auc_score(y_test, y_pred_prob)
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression()
# %time logreg.fit(X_train_dtm, y_train)
y_pred_class = logreg.predict(X_test_dtm)
y_pred_prob = logreg.predict_proba(X_test_dtm)[:,1]
y_pred_prob
metrics.accuracy_score(y_test,y_pred_class)
metrics.roc_auc_score(y_test,y_pred_prob)
|
Misc/Spam Filtering/Spam_Filtering_of_SMS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: vinyl
# language: python
# name: vinyl
# ---
# ## Python Imports
# +
import librosa
import spotipy
import os, requests, time, random, json
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.layers.recurrent import LSTM
from keras.layers.convolutional import Conv3D
from keras.layers.convolutional_recurrent import ConvLSTM2D
from keras.layers.normalization import BatchNormalization
from keras.optimizers import Adam
# %matplotlib inline
import matplotlib.pyplot as plt
import librosa.display
import IPython.display as ipd
# +
from src.obtain.spotify_metadata import generate_token, download_playlist_metadata
from src.vinyl.build_datasets import extract_features
from src.vinyl.build_datasets import build_dataset
import src.vinyl.db_manager as crates
# -
# ## [Globals](https://www.geeksforgeeks.org/global-local-variables-python/)
# +
# globals
spotify_username = 'djconxn'
user_id = "spotify:user:djconxn"
zoukables_uri = "spotify:playlist:79QPn32wwghlJfTImywNgV"
zouk_features_path = "data/zoukable_spectral.npy"
# -
# ## Model Config
# ### Features Set
features_dict = {
librosa.feature.mfcc : {'n_mfcc':12},
librosa.feature.spectral_centroid : {},
librosa.feature.chroma_stft : {'n_chroma':12},
librosa.feature.spectral_contrast : {'n_bands':6},
#librosa.feature.tempogram : {'win_length':192}
}
# ### Model Architecture
# #### TODO: Design a schema for configuring Keras models to build
# +
# seq.add(ConvLSTM2D(filters=40, kernel_size=(3, 3),
# input_shape=input_shape,
# padding='same', return_sequences=True))
# seq.add(BatchNormalization())
# seq.add(ConvLSTM2D(filters=40, kernel_size=(3, 3),
# padding='same', return_sequences=True))
# seq.add(BatchNormalization())
# seq.add(ConvLSTM2D(filters=40, kernel_size=(3, 3),
# padding='same', return_sequences=True))
# seq.add(BatchNormalization())
# seq.add(ConvLSTM2D(filters=40, kernel_size=(3, 3),
# padding='same', return_sequences=True))
# seq.add(BatchNormalization())
# seq.add(Conv3D(filters=1, kernel_size=(3, 3, 3),
# activation='sigmoid',
# padding='same', data_format='channels_last'))
# seq.compile(loss='binary_crossentropy', optimizer='adadelta')
# Keras optimizer defaults:
# Adam : lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-8, decay=0.
# RMSprop: lr=0.001, rho=0.9, epsilon=1e-8, decay=0.
# SGD : lr=0.01, momentum=0., decay=0.
# -
# # Obtain Data
#
# Set up the Spotify client, download metadata from a Zouk playlist and a non-Zouk playlist.
#
# Download song mp3 samples.
# ## Authenticate Spotify Client
token=generate_token(username=spotify_username)
sp = spotipy.Spotify(auth=token)
# ## Download Zouk Playlist Metadata
zouk_songs = crates.download_playlist_songs(sp, user_id, "zoukables", zoukables_uri)
# zouk_metadata = download_playlist_metadata(user_id, zoukables_uri, "pname", sp)
# ## Download Zouk Playlist Sample mp3's
# zouk_songs = crates.get_playlist_songs('zoukables')
for song_id in zouk_songs:
crates.get_preview_mp3(song_id)
# ## Sample Non-Zouk Songs
# #### TODO: Remove songs in `zoukables` list
non_zouk_songs = crates.sample_other_songs(n_songs=len(zouk_songs), skip_genres=["zoukables"])
# # Calculate Audio Features for Songs
#
#
# Sample 10 other genres. Add the songs from their playlists to one list. Sample `n_zouk_songs` from that list. Use these as negative cases for training our zouk classifier. Train to convergence, then repeat with another sample of non-zouk songs.
# ## Build Audio Features
#
# #### TODO: save features to `/data/librosa_features`
# Saving the feature array to a numpy file is a terrible caching practice.
#
# New workflow for `build_dataset`:
# - Download preview mp3's, extract features and save to `/data/librosa_features`
# - Return list of (unique) mp3's successfully downloaded + extracted
# - Add new songs if needed for balanced training sets
# - Build training dataset from already-extracted features
zouk_data = build_dataset(zouk_songs, features_dict)
non_zouk_data = build_dataset(non_zouk_songs, features_dict)
# ## Build Targets
target = np.array([1] * len(zouk_songs) + [0] * len(non_zouk_songs))
# ## Train Test Split
print(zouk_data.shape)
print(non_zouk_data.shape)
# +
X = np.concatenate((zouk_data, non_zouk_data))
train_idx, test_idx, y_train, y_test = train_test_split(
range(X.shape[0]), target, test_size=0.33, random_state=42, stratify=target)
X_train = X[train_idx,:,:]
X_test = X[test_idx,:,:]
# -
# # Generating Sequences for an LSTM Classifier
# ## Build Model
#
# #### TODO: Study Convolutional LSTMs
# I think this would make the model robust to handling similar songs in different keys
# +
input_shape = (X_train.shape[1], X_train.shape[2])
print("Build LSTM model ...")
model = Sequential()
model.add(LSTM(units=128, dropout=0.05, recurrent_dropout=0.35, return_sequences=True, input_shape=input_shape))
model.add(LSTM(units=64, dropout=0.05, recurrent_dropout=0.35, return_sequences=True))
model.add(LSTM(units=32, dropout=0.05, recurrent_dropout=0.35, return_sequences=False))
model.add(Dense(units=1, activation="sigmoid"))
print("Compiling ...")
opt = Adam()
model.compile(loss="binary_crossentropy", optimizer=opt, metrics=["accuracy"])
model.summary()
# -
# ## Train Model
# #### TODO: log the training reports to keep track of learning rates and training times.
print("Training ...")
batch_size = 35 # num of training examples per minibatch
num_epochs = 400
model.fit(
X_train,
y_train,
batch_size=batch_size,
epochs=num_epochs,
validation_split=.25,
verbose=1,
callbacks=[
keras.callbacks.EarlyStopping(patience=8, verbose=1, restore_best_weights=True),
keras.callbacks.ReduceLROnPlateau(factor=.5, patience=3, verbose=1),
]
)
# ## Evaluate Model
print("\nTesting ...")
score, accuracy = model.evaluate(
X_test, y_test, batch_size=batch_size, verbose=1
)
print("Test loss: ", score)
print("Test accuracy: ", accuracy)
# ## Save Model
model.save("models/zouk_classifier_spectral_LSTM3.h5")
# # Is It Any Good?
#
# Do some explanatory analysis to see what songs are being misclassified. I know that the "labels" are sketchy, so I'll need to do some data cleaning and re-training. How bad is it?
# ## Get Predictions From Training Set
# +
all_songs = pd.DataFrame({'song_id':zouk_songs + non_zouk_songs,
'target':target})
trainers = all_songs.iloc[train_idx,:].reset_index()
sample0 = trainers[trainers.target==0].sample(10).index
sample1 = trainers[trainers.target==1].sample(10).index
sample_idx = sample0.append(sample1)
samples = trainers.loc[sample_idx]
# -
# ## Print Classification Report
y_pred = model.predict(X_train[sample_idx,:])
y_pred_bool = y_pred > 0.75
samples['prediction'] = y_pred_bool.astype(int)
print(classification_report(samples.target, y_pred_bool))
# #### TODO: Add False Positives, False Negatives to Spotify playlists
# +
candidates_uri = 'spotify:playlist:69K5ogTF87NeSFvU9ePI3x'
suspects_uri = 'spotify:playlist:3M1IBVChmAYh7srqwK0CDt'
def update_screening_playlists(false_positives, false_negatives):
global user_id
global candidates_uri
global suspects_uri
sp.user_playlist_add_tracks(user_id, candidates_uri, false_positives)
sp.user_playlist_add_tracks(user_id, suspects_uri, false_negatives)
# -
# ## Sample False Positives and False Negatives
# +
fp_index = samples[(samples.target==0) & (samples.prediction==1)].index
fn_index = samples[(samples.target==1) & (samples.prediction==0)].index
print("False Positives:")
for i in fp_index:
song_id = samples['song_id'][i]
filepath = crates.get_preview_mp3(song_id)
print(crates.load_song_metadata(song_id)['title'])
ipd.display(ipd.Audio(filepath))
print("~" * 32)
print("False Negatives:")
for i in fn_index:
song_id = samples['song_id'][i]
filepath = crates.get_preview_mp3(song_id)
print(crates.load_song_metadata(song_id)['title'])
ipd.display(ipd.Audio(filepath))
# -
# # Ship It!
#
# Create a new notebook and copy over the code it needs to run the app from scratch.
#
# Copy over the functions that return the output, and then iterate running the function and copying over the imports and function definitions that are needed to get it to execute without crashing.
#
# (MVP for this should probably run on a single song, not all the songs on a playlist... downloading and extracting the features for many songs is going to take a long time.)
# # References
#
# - [Every Noise At Once](http://everynoise.com/)
# - [Keras docs](https://keras.io/)
# - [Librosa docs](https://librosa.github.io/librosa/index.html)
# - [Spotipy docs](https://spotipy.readthedocs.io)
# - [ruohoruotsi: LSTM Music Genre Classification on GitHub](https://github.com/ruohoruotsi/LSTM-Music-Genre-Classification)
# - [Music Genre classification using a hierarchical Long Short Term Memory (LSTM) Model](http://www.cs.cuhk.hk/~khwong/p186_acm_00_main_lstm_music_rev5.pdf)
# - [Using CNNs and RNNs for Music Genre Recognition](https://towardsdatascience.com/using-cnns-and-rnns-for-music-genre-recognition-2435fb2ed6af) [(GitHub)](https://github.com/priya-dwivedi/Music_Genre_Classification)
# - [The dummy’s guide to MFCC](https://medium.com/prathena/the-dummys-guide-to-mfcc-aceab2450fd)
# - [Convolutional LSTM Network: A Machine Learning Approach for Precipitation Nowcasting](https://arxiv.org/abs/1506.04214v1)
# - [An introduction to ConvLSTM](https://medium.com/neuronio/an-introduction-to-convlstm-55c9025563a7)
# # Storage Space Requirements
#
# .model files = 1 - 6 MB
#
# features = 250MB (spectral), 1.7GB(tempo)
#
# mp3 previews = 365 kB ea
#
# librosa features = 420 kB ea
# # Action Plan
#
# This process should train a decent classifier for songs from this playlist, but I really need to find a much larger list of positive cases. My plan is to maintain three playlists on Spotify:
# - the Zoukables list, which I've curated
# - a False Positives list, non-zouk songs which have been classified as zoukable and may very well be zoukable (since in our workflow, "negative" just means "has not been tagged positive"), which I can then screen and possibly add to the Zoukables list
# - a False Negatives list, zouk songs which have been classified as not zoukable and may not actually belong in the Zoukables list
#
# Once I set up these playlists and connect them to my pipeline, I can run and re-run the training pipeline, and listen and screen the Spotify playlists to curate my training set.
#
# And then the next step would be to engineer a system where other users can vote on songs to add to the Zoukables list, and automatically add songs with a threshold of votes and a high enough percentage of Yes votes.
#
# ## Spotify Playlist Updates
#
# - Refresh Zoukables list when training models
# - Update FP/FN screening playlists on Spotify
# - Update GitHub
#
# ## Re-implement EveryNoise Scraper
#
# (I think this is working)
#
# - Download EveryNoise playlist URLs
# - Download Spotify playlist metadata
# - Download preview mp3s (during model training)
# - Update GitHub
#
# ## Mongo DB: Songs Database
#
# (I've got this working in flat files)
#
# - Song IDs
# - Spotify metadata
# - Librosa Features
# - Genre Labels
# - Python API (1.4.0.1/2/3, 2.1.0.1)
# - Update GitHub
#
# ## Mongo DB: Models Database
#
# - Keras schema (0.3.2.1)
# - Feature sets
# - Training reports
# - Python API (3.1.0.1, 3.2.0.1)
# - Update GitHub
#
# ## Python Package
#
# - Keras model API
# - Organize modules
# - Write docstrings
# - Conda environment
# - Update GitHub
#
# ## Deployment
# - Reproduce pipeline on other machines
# - Reproduce pipeline for other genres
# - Deploy to AWS
#
|
Is This Zoukable (Dev).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import higra as hg
from functools import partial
from scipy.cluster.hierarchy import fcluster
from ultrametric.optimization import UltrametricFitting
from ultrametric.data import load_datasets, show_datasets
from ultrametric.graph import build_graph, show_graphs
from ultrametric.utils import Experiments
from ultrametric.evaluation import eval_clustering
# The following line requires that a C++14 compiler be installed
# On Windows, you should probably run
# c:\Program Files (x86)\Microsoft Visual Studio\2017\Enterprise\VC\Auxiliary\Build\vcvars64.bat
# to properly setup all environment variables
from ultrametric.loss import loss_closest, loss_closest_and_cluster_size, make_triplets, loss_closest_and_triplet, loss_dasgupta
# -
# ## Toy data
sets = load_datasets(n_samples=200, n_labeled=20)
exp = Experiments(sets)
show_datasets(sets, show_labeled=True)
show_graphs(sets, "knn-mst")
# ## Test function
def run_test(exp, method_name, method):
for set_name in exp.sets:
X, y, n_clusters, labeled = exp.get_data(set_name)
A = build_graph(X, 'knn-mst', mst_weight=1)
graph, edge_weights = hg.adjacency_matrix_2_undirected_graph(A)
hierarchy = method(X, labeled, y[labeled], graph, edge_weights)
Z = hg.binary_hierarchy_to_scipy_linkage_matrix(*hierarchy)
y_prediction = fcluster(Z, n_clusters, criterion='maxclust') - 1
exp.add_results(method_name, set_name, y=y_prediction, linkage=Z)
scores = eval_clustering( y, y_prediction)
d_purity = hg.dendrogram_purity(hierarchy[0], y)
print('{:10s} - {:7s} - {:.4f} - {:.4f} - {:.4f} - {:.4f} - {:.4f}'.format(method_name, set_name, *scores, d_purity))
# # Agglomerative Clustering
# +
methods = {
'average': lambda X, labeled, y_labeled, graph, edge_weights: hg.binary_partition_tree_average_linkage(graph, edge_weights),
'ward': lambda X, labeled, y_labeled, graph, edge_weights: hg.binary_partition_tree_ward_linkage(graph, X)
}
for method_name, method in methods.items():
print('{:10s} - {:7s} - {:6s} - {:6s} - {:6s} - {:6s} - {:6s}'.format("method", "set", "acc", "pur", "nmi", "randi", "dendpur"))
run_test(exp, method_name, method)
exp.show(method_name, ["clustering", "dendrogram"])
# -
# # Closest ultrametric fitting
# +
def closest_ultrametric(X, labeled, y_labeled, graph, edge_weights):
optim = UltrametricFitting(500, 0.1, loss_closest)
ultrametric = optim.fit(graph, edge_weights)
return hg.bpt_canonical(graph, ultrametric)
print('{:10s} - {:7s} - {:6s} - {:6s} - {:6s} - {:6s} - {:6s}'.format("method", "set", "acc", "pur", "nmi", "randi", "dendpur"))
run_test(exp, "closest", closest_ultrametric)
exp.show("closest", ["clustering", "dendrogram"])
# -
# # Closest ultrametric fitting + Cluster size regularization
# +
def closest_and_cluster_size_ultrametric(X, labeled, y_labeled, graph, edge_weights):
loss = partial(loss_closest_and_cluster_size, top_nodes=10)
optim = UltrametricFitting(500, 0.1, loss)
ultrametric = optim.fit(graph, edge_weights)
return hg.bpt_canonical(graph, ultrametric)
print('{:12s} - {:7s} - {:6s} - {:6s} - {:6s} - {:6s} - {:6s}'.format("method", "set", "acc", "pur", "nmi", "randi", "dendpur"))
run_test(exp, "closest+size", closest_and_cluster_size_ultrametric)
exp.show("closest+size", ["clustering", "dendrogram"])
# -
# # Closest ultrametric fitting + Triplet regularization
# +
def closest_and_triplet(X, labeled, y_labeled, graph, edge_weights):
triplets = make_triplets(y_labeled, labeled)
loss = partial(loss_closest_and_triplet, triplets=triplets, margin=1)
optim = UltrametricFitting(500, 0.1, loss)
ultrametric = optim.fit(graph, edge_weights)
return hg.bpt_canonical(graph, ultrametric)
print('{:10s} - {:7s} - {:6s} - {:6s} - {:6s} - {:6s} - {:6s}'.format("method", "set", "acc", "pur", "nmi", "randi", "dendpur"))
run_test(exp, "closest+triplet", closest_and_triplet)
exp.show("closest+triplet", ["clustering", "dendrogram"])
# -
# # Dasgupta ultrametric fitting
# +
def dasgupta(X, labeled, y_labeled, graph, edge_weights):
optim = UltrametricFitting(500, 0.1, partial(loss_dasgupta, sigmoid_param=50))
edge_weights = edge_weights / np.max(edge_weights)
ultrametric = optim.fit(graph, edge_weights)
return hg.bpt_canonical(graph, ultrametric)
print('{:10s} - {:7s} - {:6s} - {:6s} - {:6s} - {:6s} - {:6s}'.format("method", "set", "acc", "pur", "nmi", "randi", "dendpur"))
run_test(exp, "dasgupta", dasgupta)
exp.show("dasgupta", ["clustering", "dendrogram"])
# -
|
Clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The question is how well the Balmer-decrement-derived mean extinction, can correct line-flux ratios at other wavelength combinations; after discussion with LVM team
# +
import numpy as np
import matplotlib
from astropy.io import fits as fits
from astropy.table import Table
from matplotlib.colors import LogNorm
import scipy.stats as stats
# Set up matplotlib
import matplotlib.pyplot as plt
#reddening curves KK
from dust_extinction.parameter_averages import CCM89, F99
import astropy.units as u
# -
#define a few extinction-related quanbtities (REF)
def kl(lam): # get the extinction at wavelength lambda [in microns], to be multiplied by E(B-V)
if (lam <0.6):
return -5.726 + 4.004/lam - 0.525/lam**2 +0.029/lam**3 + 2.505
else:
return -2.672 - 0.010/lam + 1.532/lam**2 - 0.412/lam**3 +2.505
#define a few extinction-related quanbtities (REF) ##KK edited
def k_dust(lam): # get the extinction at wavelength lambda [in microns], to be multiplied by E(B-V)
lam2=(lam*u.micron)
#ext_model = CCM89(Rv=3.1)
#ext_model = F99(Rv=3.1)
#return ext_model(lam2)
return F99.evaluate(F99,lam2,Rv=3.1)*3.1
#return CCM89.evaluate(lam2,3.1)*3.1
print(kl(0.3727), kl(0.4868), kl(0.5007), kl(0.6564)) # just testing
print(k_dust(0.3727), k_dust(0.4868), k_dust(0.5007), k_dust(0.6564)) # just testing KK
# #### Now define the true change in the line ratio (at $\lambda_1$ vs $\lambda_2$), caused by patchy dust-extinction with E(B-V), except for a clear (area) fraction of $\epsilon$. And define the estimated E(B-V) from the observed Balmer decrement (or any other line ratio, assuming homogeneity)
# +
def line_ratio_reddening(lam1,lam2,EBV,eps):
exp_alam1 = np.exp(-k_dust(lam1)*EBV) #KK
exp_alam2 = np.exp(-k_dust(lam2)*EBV) #KK
return (eps*(1-exp_alam1)+exp_alam1) / (eps*(1-exp_alam2)+exp_alam2)
def estimate_EBV(lam1,lam2,line_ratio_change): # "line_ratio_change" is the ratio by
#which the observed line ratio differs from the expected (unreddened) one; e.g. 2.86 for the Balmer decrement
if (line_ratio_change>1.):
print('wrong line ratio regime')
else:
return -np.log((line_ratio_change))/(k_dust(lam1)-k_dust(lam2)) #KK dust
def sys_err(lam1,lam2,EBV,eps): # systematic error in dereddening line ratios at lam1 and lam2,
# using the Balmer decrement, when E(B-V) and epsilon
BD_obs = line_ratio_reddening(0.4868,0.6564,EBV,eps) # true amount by which the Balmer decretent is altered
EBV_estimated = estimate_EBV(0.4868,0.6564,BD_obs) # actually estimated B.d. to be from the observed line ratios
line_ratio_obs = line_ratio_reddening(lam1,lam2,EBV,eps)
line_ratio_after_inferred_correction = line_ratio_reddening(lam1,lam2,EBV_estimated,0.)
return line_ratio_obs/line_ratio_after_inferred_correction
def sys_err_array(lam1,lam2,X,Y): # get the previous function for a 2D array
Z = 0*X
for i in range(len(X[0,:])):
for j in range(len(Y[:,0])):
Z[i,j] = np.log10( np.abs( sys_err(lam1,lam2,X[i,j],Y[i,j]) ) ) #log to log10
return Z
# -
# Now assume there is a certain foreground absorption of E(B-V), that covers all but $\epsilon$ of the spaxel (where the flux emerges unattenuated).
# Let's make a 2D plot of the systematic error incurred when using the Balmer decrement to de-reddene [OII]/[OIII], as a function of E(B-V) and $\epsilon$
# +
x = np.linspace(0.05, 1.1, 50)
y = np.linspace(0.01, 0.3, 50)
X, Y = np.meshgrid(x, y)
Z = sys_err_array(0.3727,0.5007,X,Y) # this is specific to 3727 / 5007
#plt.contourf(X, Y, Z, 20, cmap='RdGy');
#plt.contourf(X, Y, Z, 20, cmap='nipy_spectral'); #orig
plt.contourf(X, Y, Z, 20, cmap='nipy_spectral',vmin=0,vmax=0.2); #KK
#plt.colorbar();
plt.xlabel('E(B-V)',fontsize=18,labelpad=0)
plt.tick_params(labelsize=14)
plt.ylabel('$\epsilon_{unobscured}$',fontsize=18,labelpad=0)
cbar = plt.colorbar()
cbar.set_label('log(sys. [OII]/[OIII] error)', rotation=270,fontsize=16,labelpad=23)
plt.savefig('systematic_dereddening_error_F99.pdf')
# -
x[26]
tmparr=Z[:,26]
print(tmparr)
np.median(tmparr)
# The following uses HWs original code:
# +
##original from HW
def line_ratio_reddening_orig(lam1,lam2,EBV,eps):
exp_alam1 = np.exp(-kl(lam1)*EBV)
exp_alam2 = np.exp(-kl(lam2)*EBV)
return (eps*(1-exp_alam1)+exp_alam1) / (eps*(1-exp_alam2)+exp_alam2)
def estimate_EBV_orig(lam1,lam2,line_ratio_change): # "line_ratio_change" is the ratio by
#which the observed line ratio differs from the expected (unreddened) one; e.g. 3.86 for the Balmer decrement
if (line_ratio_change>1.):
print('wrong line ration regime')
else:
return -np.log(line_ratio_change)/(kl(lam1)-kl(lam2))
def sys_err_orig(lam1,lam2,EBV,eps): # systematic error in dereddening line ratios at lam1 and lam2,
# using the Balmer decrement, when E(B-V) and epsilon
BD_obs = line_ratio_reddening_orig(0.4868,0.6564,EBV,eps) # true amount by which the Balmer decretent is altered
EBV_estimated = estimate_EBV_orig(0.4868,0.6564,BD_obs) # actually estimated B.d. to be from the observed line ratios
line_ratio_obs = line_ratio_reddening_orig(lam1,lam2,EBV,eps)
line_ratio_after_inferred_correction = line_ratio_reddening_orig(lam1,lam2,EBV_estimated,0.)
return line_ratio_obs/line_ratio_after_inferred_correction
def sys_err_array_orig(lam1,lam2,X,Y): # get the previous function for a 2D array
Z = 0*X
for i in range(len(X[0,:])):
for j in range(len(Y[:,0])):
Z[i,j] = np.log10( np.abs( sys_err_orig(lam1,lam2,X[i,j],Y[i,j]) ) ) #log to log10
return Z
# +
x = np.linspace(0.05, 1.1, 50)
y = np.linspace(0.01, 0.3, 50)
X, Y = np.meshgrid(x, y)
Z = sys_err_array_orig(0.3727,0.5007,X,Y) # this is specific to 3727 / 5007
#plt.contourf(X, Y, Z, 20, cmap='RdGy');
#plt.contourf(X, Y, Z, 20, cmap='nipy_spectral'); #orig
plt.contourf(X, Y, Z, 20, cmap='nipy_spectral',vmin=0,vmax=0.2); #KK
#plt.colorbar();
plt.xlabel('E(B-V)',fontsize=18,labelpad=10)
plt.tick_params(labelsize=14)
plt.ylabel('$\epsilon_{unobscured}$',fontsize=18,labelpad=10)
plt.clim([0,.2])#KK
cbar = plt.colorbar()
cbar.set_label('log(sys. [OII]/[OIII] error)', rotation=270,fontsize=16,labelpad=23)
plt.savefig('systematic_dereddening_error_orig.pdf')
# -
# Now assume there is a certain foreground absorption of E(B-V), that covers all but $\epsilon$ of the spaxel (where the flux emerges unattenuated).
# Let's make a 2D plot of the systematic error incurred when using the Balmer decrement to de-reddene [SII]/[SIII], as a function of E(B-V) and $\epsilon$
# +
x = np.linspace(0.05, 1.1, 50)
y = np.linspace(0.01, 0.3, 50)
X, Y = np.meshgrid(x, y)
Z = sys_err_array(0.9069,0.6716,X,Y) # this is specific to 3727 / 5007
#plt.contourf(X, Y, Z, 20, cmap='RdGy');
#plt.contourf(X, Y, Z, 20, cmap='nipy_spectral'); #orig
plt.contourf(X, Y, Z, 20, cmap='nipy_spectral',vmin=0,vmax=0.2); #KK
#plt.colorbar();
plt.xlabel('E(B-V)',fontsize=18,labelpad=0)
plt.tick_params(labelsize=14)
plt.ylabel('$\epsilon_{unobscured}$',fontsize=18,labelpad=0)
cbar = plt.colorbar()
cbar.set_label('log(sys. [SIII]/[SII] error)', rotation=270,fontsize=16,labelpad=23)
plt.savefig('systematic_dereddening_error_F99_Sulphur.pdf')
# -
tmparr=Z[:,26]
np.median(tmparr)
|
notebooks/Dereddening_under_PatchyExtinction_KK.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="G9kMNtqz8vxy"
# ## 1. urllib 활용
# + id="YRWcMwablErt"
import requests as r
import urllib.request as ur
import os
import pandas as pd
import time
# + id="Gtb05lJ-lKy0"
# 해당 디렉토리가 없으면 만드는 함수
def createFolder(dir_name):
try:
if not os.path.exists(dir_name):
os.makedirs(dir_name)
except OSError:
print ('Error: Creating directory. ' + dir_name)
# + colab={"base_uri": "https://localhost:8080/"} id="VftMTBPPlh5Y" outputId="46af9a21-652d-4f3f-b58a-d1baa26eea36"
# 쿼리를 요청할 검색어 리스트 만들거나 불러오기
name_list = ['<NAME>','<NAME>' ,'<NAME>']
# + id="CPP5ppjQ5sf1"
key1 = "" # Your bing API key
# + id="Wvv3_-ib8ZQo"
def make_img_list(lst, apikey):
# 시작 시간 저장
start = time.time()
# 에러 로그 저장을 위한 리스트 만들기
error_log = []
# 주어진 리스트에 있는 모든 이름 검색
for item in lst:
# 폴더 만들고 경로 이동하기
os.chdir("./bing_img") # your designated directory
createFolder(item)
os.chdir(f"./{item}")
url = "https://api.bing.microsoft.com/v7.0/images/search"
headers = {
"Ocp-Apim-Subscription-Key" : key1
}
param = {"q" : item,
"count" : 10,
"imageType" : "Photo",
"imageContent" : "Face",
"minHeight" : 256,
"safeSearch" : "strict",
"color" : "ColorOnly"
}
s = r.Session()
test = s.get(url,headers=headers, params=param).json()
# 검색결과 링크에서 이미지 저장하기
for i in range(0,len(test.get("value"))):
print(item,'-',i)
print(test.get('value')[i].get('contentUrl'))
file_name = f"{item}_{i}.{test.get('value')[i].get('encodingFormat')}"
try:
ur.urlretrieve(test.get('value')[i].get('contentUrl'),file_name)
except Exception as e:
print(e)
error_log.append(f"{file_name}: {e}")
pass
os.chdir("../..")
time_taken = time.time()-start
return time_taken, error_log
# + colab={"base_uri": "https://localhost:8080/"} id="2AxHwiqj8yj5" outputId="e8238267-fd96-4252-d55a-5238b85208b1"
make_img_list(name_list, key1)
# -
|
bing_img_search_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
# %matplotlib inline
import argparse
import os
import pprint
import shutil
import pickle
import cv2
from collections import defaultdict
import matplotlib.pyplot as plt
import numpy as np
import random
import torch
import torch.nn.parallel
import torch.backends.cudnn as cudnn
import torch.optim
import torch.utils.data
import torch.utils.data.distributed
import torchvision.transforms as transforms
#from tensorboardX import SummaryWriter
import _init_paths
from config import cfg
from config import update_config
from core.loss import JointsMSELoss
from core.function import train
from core.function import validate
from utils.utils import get_optimizer
from utils.utils import save_checkpoint
from utils.utils import create_logger
from utils.utils import get_model_summary
import dataset
import models
# +
with open('/u/snaha/v6/dataset/AWA/Animals_with_Attributes2/quadruped_keypoints/coco_format/dataset.pickle', 'rb') as handle:
dataset = pickle.load(handle)
dataset.keys()
# -
dataset['images'][0]
# +
path = '/u/snaha/v6/dataset/AWA/Animals_with_Attributes2/quadruped_keypoints/Annotations/'
list_subfolders_with_paths = [f.path for f in os.scandir(path) if f.is_dir()]
animal_list = []
for animal in list_subfolders_with_paths:
animal_folder = animal#os.path.join(path, animal)
animal_list.append(animal_folder.split('/')[-1])
animal_list
# +
def generate_split(animal_list, dataset):
rand_inds = np.arange(len(animal_list)).tolist()
random.shuffle(rand_inds)
anim_list = []
for ind in rand_inds:
anim_list.append(animal_list[ind])
dataset_train = dict()
dataset_test = dict()
annotations_train = []
annotations_test = []
images_train = []
images_test = []
train_animals = anim_list[:-4]
test_animal = anim_list[-4:]
for anno in dataset['annotations']:
if anno['animal'] in test_animal:
annotations_test.append(anno)
else:
annotations_train.append(anno)
for anno in dataset['images']:
if anno['id'].split('_')[0] in test_animal:
images_test.append(anno)
else:
images_train.append(anno)
dataset_train['annotations'] = annotations_train
dataset_test['annotations'] = annotations_test
dataset_train['images'] = images_train
dataset_test['images'] = images_test
dataset_train['categories'] = dataset['categories']
dataset_test['categories'] = dataset['categories']
dataset_train['info'] = dataset['info']
dataset_test['info'] = dataset['info']
return train_animals, test_animal, dataset_train, dataset_test
for i in range(5):
train_animals, test_animals, dataset_train, dataset_test = generate_split(animal_list, dataset)
#test_animals
print('Number of train instances = ', len(dataset_train['images']), len(dataset_train['annotations']))
print('Number of test instances = ', len(dataset_test['images']), len(dataset_test['annotations']))
with open('/u/snaha/v6/dataset/AWA/Animals_with_Attributes2/quadruped_keypoints/coco_format/dataset_train_' + str(i+1) + '.pickle', 'wb') as handle:
pickle.dump(dataset_train, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('/u/snaha/v6/dataset/AWA/Animals_with_Attributes2/quadruped_keypoints/coco_format/dataset_val_' + str(i+1) + '.pickle', 'wb') as handle:
pickle.dump(dataset_test, handle, protocol=pickle.HIGHEST_PROTOCOL)
# -
|
code/deep-high-resolution-net.pytorch/tools/.ipynb_checkpoints/awa_dataset_split-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Theory and Practice of Visualization Exercise 1
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
from IPython.display import Image
# + [markdown] nbgrader={}
# ## Graphical excellence and integrity
# + [markdown] nbgrader={}
# Find a data-focused visualization on one of the following websites that is a *positive* example of the principles that Tufte describes in *The Visual Display of Quantitative Information*.
#
# * [Vox](http://www.vox.com/)
# * [Upshot](http://www.nytimes.com/upshot/)
# * [538](http://fivethirtyeight.com/)
# * [BuzzFeed](http://www.buzzfeed.com/)
#
# Upload the image for the visualization to this directory and display the image inline in this notebook.
# + deletable=false nbgrader={"checksum": "9c86bcce96065a2133bab497403e3291", "grade": true, "grade_id": "theorypracticeex01a", "points": 2}
# Add your filename and uncomment the following line:
Image(filename='graphie.JPG')
# + [markdown] nbgrader={}
# Describe in detail the ways in which the visualization exhibits graphical *integrity* and *excellence*:
# + [markdown] deletable=false nbgrader={"checksum": "80145499593a6a8f756ab550388d18ea", "grade": true, "grade_id": "theorypracticeex01b", "points": 8, "solution": true}
# This is an okay example of Tufte's principles. It clearly shows the data trend. Upon first glance the viewer can see the how different "home prices" relate to one another in terms of "equivalent rent". The data contains a large domain, making it a large data set, but the data does not seem overwhelming or disconnected. This is an interactive graph, so for what ever "home price" the viewer selects, the only color on the graph is the "equivalent rent". This is efficient and precise and shows the data at mmore levels. All of the data is conveyed with consistent axes that do not skew or distort the data. One detail that could be improved is the labeling of the x-axis. In this graph, the axis label is also used as a graph title. It would be more effective to have the axis labeled near the axis to efficiently convey data.
# -
|
assignments/assignment04/TheoryAndPracticeEx01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# The code in this notebok reproduces the figures in **Examining the Evolution of Legal Precedent through Citation Network Analysis**.
#
# The code that produces the results can be found at https://github.com/idc9/law-net.
#
# The very last cell contains code to test pairwise comparisions between two metrics.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import igraph as ig
from scipy.stats import linregress
from scipy.stats import ttest_rel
repo_directory = '/Users/iaincarmichael/Dropbox/Research/law/law-net/'
data_dir = '/Users/iaincarmichael/data/courtlistener/'
network_name = 'scotus'
raw_dir = data_dir + 'raw/'
subnet_dir = data_dir + network_name + '/'
text_dir = subnet_dir + 'textfiles/'
results_dir = subnet_dir + 'results/'
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
name = '1_16_17'
sort_path = results_dir + 'sort/%s/rankloss_sort.p' % name
rankloss_sort = pd.read_pickle(open(sort_path, "rb"))
rankloss = {'sort': rankloss_sort,
'match': rankloss_match}#,
# -
G = ig.Graph.Read_GraphML(subnet_dir + network_name +'_network.graphml')
exper = 'sort'
metric = 'MRS'
# # Helper functions
def plot_scores(results, exper='', metric='', network_name=''):
"""
plots the results
"""
# compute mean and std of data
data = pd.DataFrame(index=results.columns, columns=['score', 'error'])
data['score'] = results.median(axis=0)
data.sort_values(by='score', inplace=True)
# label locations
pos = np.arange(data.shape[0])
plt.barh(pos,
data['score'],
color='grey')
plt.xlim([0, 1.2 * data['score'].max()])
axis_font = {'fontname': 'Arial', 'size': '12'}
plt.yticks(pos, data.index, **axis_font)
plt.xlabel('mean rank score')
plt.gca().spines.values()[1].set_visible(False)
plt.gca().spines.values()[3].set_visible(False)
# # Sort Experiment Results
# ## Figure E: compare in-degree driven metrics
# +
metrics_to_show = ['indegree', 'd_pagerank', 'authorities', 'd_betweenness']
plt.figure(figsize=[8, 8])
plot_scores(rankloss[exper][metric][metrics_to_show], exper=exper, metric=metric, network_name=network_name)
# -
# ## Figure F: include out-degree
# +
metrics_to_show = ['indegree', 'd_pagerank', 'authorities', 'd_betweenness', 'outdegree']
plt.figure(figsize=[8, 8])
plot_scores(rankloss[exper][metric][metrics_to_show], exper=exper, metric=metric, network_name=network_name)
# -
# ## figure H: num words vs. out-degree
num_words = np.array(G.vs['num_words'])
outdegrees = np.array(G.outdegree())
indegrees = G.indegree()
years = G.vs['year']
# +
# remove some outliers
out_deg_upper = np.percentile(outdegrees, 99)
out_deg_lower = np.percentile(outdegrees, 0)
num_words_upper = np.percentile(num_words, 99)
num_words_lower = np.percentile(num_words, 0)
od_to_keep = (out_deg_lower <= outdegrees) & (outdegrees <= out_deg_upper)
nw_to_keep = (num_words_lower <= num_words) & (num_words <= num_words_upper)
to_keep = od_to_keep & nw_to_keep
nw = num_words[to_keep]
od = outdegrees[to_keep]
# -
# remove cases that have zero out-degree
slope, intercept, r_value, p_value, std_err = linregress(nw, od)
# +
plt.figure(figsize=[8, 8])
plt.scatter(nw, od, color='grey', s=10)
plt.xlabel('number of words')
plt.ylabel('out-degre')
# kill top and right axes
plt.gca().spines.values()[1].set_visible(False)
plt.gca().spines.values()[3].set_visible(False)
plt.xlim([0, max(nw)*1.1])
plt.ylim([0, max(od)*1.1])
xvals = np.array([0, max(nw)])
line = slope * xvals + intercept
plt.plot(xvals, line, color='red', linewidth=5.0)
plt.title('opinion text length vs. out-degree')
# -
# # Figure I
# +
metrics_to_show = ['indegree', 'd_pagerank', 'authorities', 'd_betweenness', 'outdegree', 'num_words']
plt.figure(figsize=[8, 8])
plot_scores(rankloss[exper][metric][metrics_to_show], exper=exper, metric=metric, network_name=network_name)
# -
# # Figure J: citation ages
diffs = [G.vs[e[0]]['year'] - G.vs[e[1]]['year'] for e in G.get_edgelist()]
# +
plt.figure(figsize=[8, 8])
bins = np.linspace(-40, 300, 100)
plt.hist(diffs, bins=bins, color='grey')
plt.xlim(0, 300)
plt.xlabel('citation age')
plt.gca().spines.values()[1].set_visible(False)
plt.gca().spines.values()[3].set_visible(False)
plt.title('distribution of SCOTUS citation ages')
# -
# # Figure K: time aware
# +
metrics_to_show = [ 'd_pagerank','citerank_50',
'indegree', 'd_betweenness',
'authorities', 'recentcite_2',
'outdegree', 'recentcite_5',
'recentcite_20', 'citerank_10',
'recentcite_10', 'citerank_5',
'age', 'citerank_2']
plt.figure(figsize=[8, 8])
plot_scores(rankloss[exper][metric][metrics_to_show], exper=exper, metric=metric, network_name=network_name)
# -
# # Figure L: Federal
# +
rankloss_sort_federal = pd.read_pickle('/Users/iaincarmichael/data/courtlistener/federal/results/sort/federal_test/rankloss_sort.p')
rankloss_federal = {'sort': rankloss_sort_federal}
# +
metrics_to_show = ['hubs', 'd_pagerank', 'authorities', 'outdegree', 'indegree']
plt.figure(figsize=[8, 8])
plot_scores(rankloss_federal[exper][metric][metrics_to_show], exper=exper, metric=metric, network_name=network_name)
# -
# # Figure M: warren court
# +
def get_year_aggregate(years, x, fcn):
by_year = {y: [] for y in set(years)}
for i in range(len(years)):
by_year[years[i]].append(x[i])
year_agg_dict = {y: fcn(by_year[y]) for y in by_year.keys()}
return pd.Series(year_agg_dict)
in_year_median = get_year_aggregate(years, indegrees, np.median)
nw_year_median = get_year_aggregate(years, num_words, np.median)
od_year_median = get_year_aggregate(years, outdegrees, np.median)
# +
# Text length
plt.figure(figsize=[6, 9])
plt.subplot(3,1,1)
plt.plot(nw_year_median.index, nw_year_median/1000,
color='black', marker='.', linestyle=':')
plt.axvline(1953, color='black', alpha=.5)
plt.axvline(1969, color='black', alpha=.5)
plt.ylabel('median text length')
plt.xlim([1800, 2017])
plt.ylim([0, 30])
plt.title('citation and case length statistics by year')
plt.annotate('<NAME> \n (1953-1969)', xy=(1952, 15), xytext=(1890, 20),
arrowprops=dict(fc='grey', ec='grey', shrink=0.01, width=1, headwidth=10))
plt.gca().spines.values()[1].set_visible(False)
plt.gca().spines.values()[3].set_visible(False)
# out degree
plt.subplot(3,1,2)
plt.plot(od_year_median.index, od_year_median,
color='black', marker='.', linestyle=':')
plt.axvline(1953, color='black', alpha=.5)
plt.axvline(1969, color='black', alpha=.5)
plt.ylabel('median outdegree')
plt.xlim([1800, 2017])
plt.ylim([0, 30])
plt.gca().spines.values()[1].set_visible(False)
plt.gca().spines.values()[3].set_visible(False)
# in degree
plt.subplot(3,1,3)
plt.plot(in_year_median.index, in_year_median,
color='black', marker='.', linestyle=':')
plt.axvline(1953, color='black', alpha=.5)
plt.axvline(1969, color='black', alpha=.5)
plt.ylabel('median indegree')
plt.xlabel('year')
plt.xlim([1800, 2017])
plt.ylim([0, 30])
plt.gca().spines.values()[1].set_visible(False)
plt.gca().spines.values()[3].set_visible(False)
# -
# # Figure O: page rank bias
years = np.array(G.vs['year'])
pr = np.array(G.pagerank())
# +
plt.figure(figsize=[8, 8])
plt.scatter(years, pr, color='grey', s=15)
plt.xlabel('year')
plt.ylabel('PageRank')
plt.xlim([1800, 2017])
plt.ylim([0, 1.2 *max(pr)])
plt.title('PageRank of each Supreme Court case')
plt.gca().spines.values()[1].set_visible(False)
plt.gca().spines.values()[3].set_visible(False)
# -
# # Figure P
# +
metrics_to_show = ['d_pagerank', 'indegree', 'd_betweenness', 'u_betweenness',
'authorities', 'u_pagerank', 'outdegree', 'degree', 'u_eigen']
plt.figure(figsize=[8, 8])
plot_scores(rankloss[exper][metric][metrics_to_show], exper=exper, metric=metric, network_name=network_name)
# -
# # Statsitical significance
# +
# to_compare = ['outdegree', 'hubs']
# to_compare = ['recentcite_10', 'citerank_2']
to_compare = ['num_words', 'indegree']
exper = 'sort'
metric = 'MRS'
data = rankloss[exper][metric][to_compare]
print '%s vs. %s' % ( to_compare[0], to_compare[1])
print '%s experiment, %s' % (exper,metric)
print 'two sided t-test for equal means'
print
print 'dependent paired samples'
print ttest_rel(data[to_compare[0]], data[to_compare[1]])
# -
|
examining_evolution_code/figures_for_paper.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZHHTFMsziMya" colab_type="text"
# ## PageRank
# + [markdown] id="PdFfDePRnGkR" colab_type="text"
# #### Ways to think about SVD
#
# - Data compression
# - SVD trades a large number of features for a smaller set of better features
# - All matrices are diagonal (if you use change of bases on the domain and range)
#
# **Relationship between SVD and Eigen Decomposition**: the left-singular vectors of A are the eigenvectors of $AA^T$. The right-singular vectors of A are the eigenvectors of $A^T A$. The non-zero singular values of A are the square roots of the eigenvalues of $A^T A$ (and $A A^T$).
#
# SVD is a generalization of eigen decomposition. Not all matrices have eigen values, but ALL matrices have singular values.
#
# A **Hermitian** matrix is one that is equal to it's own conjugate transpose. In the case of real-valued matrices (which is all we are considering in this course), **Hermitian** means the same as **Symmetric**.
#
# **Relevant Theorems:**
# - If A is symmetric, then eigenvalues of A are real and $A = Q \Lambda Q^T$
# - If A is triangular, then its eigenvalues are equal to its diagonal entries
#
# * 确定图中顶点的相对重要性的经典方法是计算邻接矩阵的主特征向量,以便将每个顶点的第一特征向量的分量值分配为中心性分数
#
# 1. [维基百科主要特征向量 - scikit-learn 0.19.1文档](http://scikit-learn.org/stable/auto_examples/applications/wikipedia_principal_eigenvector.html#sphx-glr-auto-examples-applications-wikipedia-principal-eigenvector-py)
# 2.[Eigenvector centrality - Wikipedia](https://en.wikipedia.org/wiki/Eigenvector_centrality)
# 3. [Power iteration - Wikipedia](https://en.wikipedia.org/wiki/Power_iteration)
# 4. [Katz centrality - Wikipedia](https://en.wikipedia.org/wiki/Katz_centrality)
# 5. [PageRank - Wikipedia](https://en.wikipedia.org/wiki/PageRank)
# 6. [PageRank算法--从原理到实现 - CSDN博客](http://blog.csdn.net/rubinorth/article/details/52215036)
#
# + id="v4kdcgiUnU4-" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}} cellView="code"
#@title Power iteration
import numpy as np
def power_iteration(A, num_simulations):
# Ideally choose a random vector
# To decrease the chance that our vector
# Is orthogonal to the eigenvector
b_k = np.random.rand(A.shape[0])
for _ in range(num_simulations):
# calculate the matrix-by-vector product Ab
b_k1 = np.dot(A, b_k)
# calculate the norm
b_k1_norm = np.linalg.norm(b_k1)
# re normalize the vector
b_k = b_k1 / b_k1_norm
return b_k
power_iteration(np.array([[0.5, 0.5], [0.2, 0.8]]), 100)
# + id="wOc2pjYjG-fD" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# 稀疏矩阵
import numpy as np
from scipy import sparse
def power_method(A, max_iter=100):
n = A.shape[1]
A = np.copy(A)
A.data /= np.take(A.sum(axis=0).A1, A.indices)
scores = np.ones(n, dtype=np.float32) * np.sqrt(A.sum()/(n*n)) # initial guess
for i in range(max_iter):
scores = A @ scores
nrm = np.linalg.norm(scores)
scores /= nrm
print(nrm)
return scores
# + id="Fdxpp3gQHtH0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
x = np.matrix(np.arange(12).reshape((3,4)))
a = sparse.csr_matrix(x, dtype=np.float32)
power_method(a, max_iter=10)
# + id="h2X3Wz0PKoRh" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# + id="V5ebU342tWKf" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
np.random.randn(2, 4)
# + [markdown] id="KtFUdeEqDGJs" colab_type="text"
# * numpy.matrix.A1
#
# Return self as a flattened ndarray. Equivalent to np.asarray(x).ravel()
#
# 1. [numpy.matrix.A1 — NumPy v1.14 Manual](https://docs.scipy.org/doc/numpy/reference/generated/numpy.matrix.A1.html#numpy.matrix.A1)
# + id="9iOwLyd8DJX0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
x = np.matrix(np.arange(12).reshape((3,4)))
# + id="aThhf8vXDDAS" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
x
# + id="Tnq8WfUWDDrp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
x.A1
# + id="ArdaqmJRDZSo" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
x.ravel()
# + id="RRSBa24cDc4Q" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
x.A1.shape, x.ravel().shape
# + [markdown] id="PSu46NULDu90" colab_type="text"
# ### How to normalize a sparse matrix
# + id="LNOdCEnFD16d" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
from scipy import sparse
S = sparse.csr_matrix(np.array([[1,2],[3,4]]))
S
# + id="7vlslNChD9yp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
Sr = S.sum(axis=0).A1
Sr
# + id="tOMJy8vcENgi" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
S.indices
# + id="NOhh5nPeEX3p" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
S.data
# + id="T_cC4G0HEzxO" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
S.data / np.take(Sr, S.indices)
# + id="FbI_7s17FXKH" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
np.take(Sr, S.indices)
# + [markdown] id="lGRkIORMNFCk" colab_type="text"
#
# + [markdown] id="Nd0jxdmFNFWT" colab_type="text"
# ### QR 分解
# + id="0j7hgwGgNJeQ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
from numba import jit
@jit()
def pure_qr(A, max_iter=50000):
Ak = np.copy(A)
n = A.shape[0]
QQ = np.eye(n)
for k in range(max_iter):
Q, R = np.linalg.qr(Ak)
Ak = R @ Q
QQ = QQ @ Q
if k % 100 == 0:
print(Ak)
print("\n")
return Ak, QQ
# + id="JVjXZwL2NLO9" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
n = 6
A = np.random.rand(n,n)
AT = A @ A.T
# + id="TvyVKxjsNQNN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
Ak, Q = pure_qr(A)
# + id="KqrvfGXNNVrC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# 特征值
np.linalg.eigvals(A)
# + id="woadPsUqNywg" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Q 是正交的
np.allclose(np.eye(n), Q @ Q.T), np.allclose(np.eye(n), Q.T @ Q)
# + id="TKtUXwEZN7SC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# + [markdown] id="P3rDIOXsO_Dv" colab_type="text"
# The Arnoldi Iteration is two things:
# 1. the basis of many of the iterative algorithms of numerical linear algebra
# 2. a technique for finding eigenvalues of nonhermitian matrices
# (Trefethen, page 257)
#
# **How Arnoldi Locates Eigenvalues**
#
# 1. Carry out Arnoldi iteration
# 2. Periodically calculate the eigenvalues (called *Arnoldi estimates* or *Ritz values*) of the Hessenberg H, using the QR algorithm
# 3. Check at whether these values are converging. If they are, they're probably eigenvalues of A.
# + id="aEOnd5QYPCDz" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Decompose square matrix A @ Q ~= Q @ H
def arnoldi(A):
m, n = A.shape
assert(n <= m)
# Hessenberg matrix
H = np.zeros([n+1,n]) #, dtype=np.float64)
# Orthonormal columns
Q = np.zeros([m,n+1]) #, dtype=np.float64)
# 1st col of Q is a random column with unit norm
b = np.random.rand(m)
Q[:,0] = b / np.linalg.norm(b)
for j in range(n):
v = A @ Q[:,j]
for i in range(j+1):
#This comes from the formula for projection of v onto q.
#Since columns q are orthonormal, q dot q = 1
H[i,j] = np.dot(Q[:,i], v)
v = v - (H[i,j] * Q[:,i])
H[j+1,j] = np.linalg.norm(v)
Q[:,j+1] = v / H[j+1,j]
# printing this to see convergence, would be slow to use in practice
print(np.linalg.norm(A @ Q[:,:-1] - Q @ H))
return Q[:,:-1], H[:-1,:]
# + id="93dvMIoGPKgg" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
Q, H = arnoldi(A)
# + id="niJA8uocPMSv" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
H
# + id="l9L48xiAPNtk" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
Q
# + id="lm0sK6KRPelf" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
n = 10
A0 = np.random.rand(n,n)
A = A0 @ A0.T
# + id="iY_w4Cq3PjwZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
np.linalg.eigvals(A)
|
fastai_notes/LinearAlgebra/speech04.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
import pandas as pd
import yfinance as yf
def get_balancesheet(ticker):
stock_obj = yf.Ticker(ticker)
return stock_obj.balancesheet
def income_statement(ticker):
stock_obj = yf.Ticker(ticker)
return stock_obj.financials
def get_info(ticker):
df = {}
ticker_info = yf.Ticker(ticker).info
df['regularMarketPrice'] = ticker_info['regularMarketPrice']
df['trailingEps'] = ticker_info['trailingEps']
df['trailingAnnualDividendRate'] = ticker_info['trailingAnnualDividendRate']
df['enterpriseValue'] = ticker_info['enterpriseValue']
return df
def magic_formula(tickers):
df = {}
for ticker in tickers:
print(ticker)
balance_sheet = get_balancesheet(ticker).transpose()
curr_assets = balance_sheet['Total Current Assets'].iloc[0]
curr_liab = balance_sheet['Total Current Liabilities'].iloc[0]
tang_assets = balance_sheet['Net Tangible Assets'].iloc[0]
working_cap = curr_assets - curr_liab
income_sheet = income_statement(ticker).transpose()
EBIT = income_sheet['Ebit'].iloc[0]
stats = get_info(ticker)
if stats['trailingEps'] != None and stats['regularMarketPrice'] != None and stats['enterpriseValue'] != None:
ROIC = EBIT / (working_cap + tang_assets)
EY_perc = (stats['trailingEps'] / stats['regularMarketPrice']) * 100
EY = EBIT / stats['enterpriseValue']
DivdendRate = stats['trailingAnnualDividendRate']
df[ticker] = [EY_perc, EY, ROIC, DivdendRate, EBIT]
df = pd.DataFrame(df).transpose()
df.columns = ["EarningsYield%", "EarningsYield", "ReturnOnInvestedCapital", 'trailingAnnualDividendRate', "EBIT"]
df["CombineRanking"] = df["EarningsYield"].rank(ascending=False,na_option='bottom') + df["ReturnOnInvestedCapital"].rank(ascending=False,na_option='bottom')
df["Magic"] = df["CombineRanking"].rank(method='first')
return df.sort_values("Magic")
tickers = pd.read_csv('s_&_p_500_tickers.csv')['Ticker']
data = magic_formula(tickers)
data
|
value_investing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Image-to-Image Translation with Conditional Adversarial Networks
# * BAIR, UC Berkeley
# ### Background
#
# ## Video-to-Video Synthesis
# * Nvidia, MIT, 2018
#
# ### Background
# * Image-to-Image Translation
# * Without modeling temporal dynamics, directly applying existing image synthesis approaches to an input video often results in temporally incoherent videos of low visual quality.
#
# ### Main Strategy
# * with a spatio-temporal adversarial objective
# * Through carefully-designed generator and discriminator architectures, coupled with a spatio-temporal adversarial objective, we achieve high-resolution, photorealistic, temporally coher- ent video results on a diverse set of input formats including segmentation masks, sketches, and poses.
# * future video prediction
# * Finally, we apply our approach to future video prediction, outperforming several state-of-the-art competing systems.
|
papers/Generative_Models/Video_Generative.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Interact Exercise 4
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# + nbgrader={}
from IPython.html.widgets import interact, interactive, fixed
from IPython.display import display
# + [markdown] nbgrader={}
# ## Line with Gaussian noise
# + [markdown] nbgrader={}
# Write a function named `random_line` that creates `x` and `y` data for a line with y direction random noise that has a normal distribution $N(0,\sigma^2)$:
#
# $$
# y = m x + b + N(0,\sigma^2)
# $$
#
# Be careful about the `sigma=0.0` case.
# + nbgrader={"checksum": "f1fccd14526477d1457886a737404055", "solution": true}
def random_line(m, b, sigma, size=10):
"""Create a line y = m*x + b + N(0,sigma**2) between x=[-1.0,1.0]
Parameters
----------
m : float
The slope of the line.
b : float
The y-intercept of the line.
sigma : float
The standard deviation of the y direction normal distribution noise.
size : int
The number of points to create for the line.
Returns
-------
x : array of floats
The array of x values for the line with `size` points.
y : array of floats
The array of y values for the lines with `size` points.
"""
x=np.linspace(-1.0,1.0,size)
N=np.empty(size)
if sigma==0.0: #I received some help from classmates here
y=m*x+b
else:
for i in range(size):
N[i]=np.random.normal(0,sigma**2)
y=m*x+b+N
return(x,y)
# + deletable=false nbgrader={"checksum": "085b717fea11f553f5549a88b1090e24", "grade": true, "grade_id": "interactex04a", "points": 2}
m = 0.0; b = 1.0; sigma=0.0; size=3
x, y = random_line(m, b, sigma, size)
assert len(x)==len(y)==size
assert list(x)==[-1.0,0.0,1.0]
assert list(y)==[1.0,1.0,1.0]
sigma = 1.0
m = 0.0; b = 0.0
size = 500
x, y = random_line(m, b, sigma, size)
assert np.allclose(np.mean(y-m*x-b), 0.0, rtol=0.1, atol=0.1)
assert np.allclose(np.std(y-m*x-b), sigma, rtol=0.1, atol=0.1)
# + [markdown] nbgrader={}
# Write a function named `plot_random_line` that takes the same arguments as `random_line` and creates a random line using `random_line` and then plots the `x` and `y` points using Matplotlib's `scatter` function:
#
# * Make the marker color settable through a `color` keyword argument with a default of `red`.
# * Display the range $x=[-1.1,1.1]$ and $y=[-10.0,10.0]$.
# * Customize your plot to make it effective and beautiful.
# + nbgrader={}
def ticks_out(ax):
"""Move the ticks to the outside of the box."""
ax.get_xaxis().set_tick_params(direction='out', width=1, which='both')
ax.get_yaxis().set_tick_params(direction='out', width=1, which='both')
# + nbgrader={"checksum": "701a9529400e32449715b0090b912d11", "solution": true}
def plot_random_line(m, b, sigma, size=10, color='red'):
"""Plot a random line with slope m, intercept b and size points."""
x=np.linspace(-1.0,1.0,size)
N=np.empty(size)
if sigma==0.0:
y=m*x+b
else:
for i in range(size):
N[i]=np.random.normal(0,sigma**2)
y=m*x+b+N
plt.figure(figsize=(9,6))
plt.scatter(x,y,color=color)
plt.xlim(-1.1,1.1)
plt.ylim(-10.0,10.0)
plt.box(False)
plt.grid(True)
# + nbgrader={"solution": false}
plot_random_line(5.0, -1.0, 2.0, 50)
# + deletable=false nbgrader={"checksum": "b079fa9a413c8bc761692d3bfd9eb813", "grade": true, "grade_id": "interactex04b", "points": 4}
assert True # use this cell to grade the plot_random_line function
# + [markdown] nbgrader={}
# Use `interact` to explore the `plot_random_line` function using:
#
# * `m`: a float valued slider from `-10.0` to `10.0` with steps of `0.1`.
# * `b`: a float valued slider from `-5.0` to `5.0` with steps of `0.1`.
# * `sigma`: a float valued slider from `0.0` to `5.0` with steps of `0.01`.
# * `size`: an int valued slider from `10` to `100` with steps of `10`.
# * `color`: a dropdown with options for `red`, `green` and `blue`.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
interact(plot_random_line,m=(-10.0,10.0,0.1), b=(-5.0,5.0,0.1), sigma=(0.0,5.0,0.01),size=(10,100,10),color=('r','b','g'))
# + deletable=false nbgrader={"checksum": "49bbb321697a88612357059cba486cd3", "grade": true, "grade_id": "interactex04c", "points": 4}
#### assert True # use this cell to grade the plot_random_line interact
# -
|
assignments/assignment05/InteractEx04.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# Extract segmentation features
# =============================
#
# This example shows how to extract segmentation features from the tissue
# image.
#
# Features extracted from a nucleus segmentation range from the number of
# nuclei per image, over nuclei shapes and sizes, to the intensity of the
# input channels within the segmented objects. They are very interpretable
# features and provide valuable additional information. Use
# `features='segmentation'` to calculate the features.
#
# In addition to `feature_name` and `channels` we can specify the
# following `features_kwargs`:
#
# - `label_layer` - name of label image layer in `img`.
# - `props` - segmentation features that are calculated. See properties
# in skimage.measure.regionprops\_table.
#
#
# +
import scanpy as sc
import squidpy as sq
import matplotlib.pyplot as plt
# -
# Lets load a fluorescence Visium dataset.
#
img = sq.datasets.visium_fluo_image_crop()
adata = sq.datasets.visium_fluo_adata_crop()
# Before calculating segmentation features, we need to first calculate a
# segmentation using squidpy.im.segment.
#
sq.im.segment(img=img, layer="image", layer_added="segmented_watershed", method="watershed", channel=0)
# Now we can calculate segmentation features. Here, we will calculate the
# following features:
#
# - number of nuclei (`label`).
# - mean area of nuclei (`area`).
# - mean intensity of channels 1 (anti-NEUN) and 2 (anti-GFAP) within
# nuclei (`mean_intensity`).
#
# We use `mask_cicle = True` to ensure that we are only extracting
# features from the tissue underneath each Visium spot. For more details
# on the image cropping, see
# sphx\_glr\_auto\_examples\_image\_compute\_crops.py.
#
sq.im.calculate_image_features(
adata,
img,
layer="image",
features="segmentation",
key_added="segmentation_features",
features_kwargs={
"segmentation": {
"label_layer": "segmented_watershed",
"props": ["label", "area", "mean_intensity"],
"channels": [1, 2],
}
},
mask_circle=True,
)
# The result is stored in `adata.obsm['segmentation_features']`.
#
adata.obsm["segmentation_features"].head()
# Use squidpy.pl.extract to plot the texture features on the tissue image
# or have a look at [our interactive visualisation
# tutorial](../../external_tutorials/tutorial_napari.html) to learn how to
# use our interactive napari plugin. Here, we show all calculated
# segmentation features.
#
# +
# show all channels (using low-res image contained in adata to save memory)
fig, axes = plt.subplots(1, 3, figsize=(8, 4))
for i, ax in enumerate(axes):
ax.imshow(adata.uns["spatial"]["V1_Adult_Mouse_Brain_Coronal_Section_2"]["images"]["hires"][:, :, i])
ax.set_title(f"ch{i}")
# plot segmentation features
sc.pl.spatial(
sq.pl.extract(adata, "segmentation_features"),
color=[
"segmentation_label",
"segmentation_area_mean",
"segmentation_ch-1_mean_intensity_mean",
"segmentation_ch-2_mean_intensity_mean",
],
bw=True,
ncols=2,
vmin="p1",
vmax="p99",
)
# -
# segmentation\_label shows the number of nuclei per spot and
# segmentation\_area\_mean the mean are of nuclei per spot. The remaining
# two plots show the mean intensity of channels 1 and 2 per spot. As the
# stains for channels 1 and 2 are specific to Neurons and Glial cells,
# respectively, these features show us Neuron and Glial cell dense areas.
#
|
docs/source/auto_examples/image/compute_segmentation_features.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="x8Q7Un821X1A"
# ##### Copyright 2018 The TensorFlow Hub Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + id="1W4rIAFt1Ui3"
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] id="cDq0CIKc1vO_"
# # 팽창된 3D CNN을 사용한 행동 인식
#
# + [markdown] id="MfBg1C5NB3X0"
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/hub/tutorials/action_recognition_with_tf_hub"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">}TensorFlow.org에서 보기</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/hub/tutorials/action_recognition_with_tf_hub.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab에서 실행하기</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/hub/tutorials/action_recognition_with_tf_hub.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub에서 소스 보기</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/hub/tutorials/action_recognition_with_tf_hub.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드하기</a></td>
# </table>
# + [markdown] id="h6W3FhoP3TxC"
# 이 Colab에서는 [tfhub.dev/deepmind/i3d-kinetics-400/1](https://tfhub.dev/deepmind/i3d-kinetics-400/1) 모듈을 사용하여 비디오 데이터에서 행동 인식을 사용하는 방법을 보여줍니다.
#
# 기본 모델은 <NAME>와 <NAME>의 "[Quo Vadis, Action Recognition? A New Model and the Kinetics Dataset](https://arxiv.org/abs/1705.07750)" 논문에 설명되어 있습니다. 이 논문은 2017년 5월 arXiv에 게시되었으며 CVPR 2017 컨퍼런스 논문으로 발표되었습니다. 소스 코드는 [github](https://github.com/deepmind/kinetics-i3d)에서 공개적으로 사용할 수 있습니다.
#
# "<NAME>"는 비디오 분류를 위한 새로운 아키텍처인 Inflated 3D Convnet 또는 I3D를 도입했습니다. 이 아키텍처는 이러한 모델을 미세 조정하여 UCF101 및 HMDB51 데이터세트에서 최고의 결과를 얻었습니다. Kinetics에 사전 훈련된 I3D 모델도 CVPR 2017 [Charades 챌린지](http://vuchallenge.org/charades.html)에서 1위를 차지했습니다.
#
# 원래 모듈은 [kinetics-400 데이터세트](https://deepmind.com/research/open-source/open-source-datasets/kinetics/)에서 훈련되었으며 약 400가지의 행동을 인식합니다. 이러한 행동의 레이블은 [레이블 맵 파일](https://github.com/deepmind/kinetics-i3d/blob/master/data/label_map.txt)에서 찾을 수 있습니다.
#
# 이 Colab에서는 UCF101 데이터세트에서 비디오 활동을 인식하는 데 이 모듈을 사용합니다.
# + [markdown] id="R_0xc2jyNGRp"
# ## 설정
# + id="mOHMWsFnITdi"
# !pip install -q imageio
# !pip install -q opencv-python
# !pip install -q git+https://github.com/tensorflow/docs
# + cellView="both" id="USf0UvkYIlKo"
#@title Import the necessary modules
# TensorFlow and TF-Hub modules.
from absl import logging
import tensorflow as tf
import tensorflow_hub as hub
from tensorflow_docs.vis import embed
logging.set_verbosity(logging.ERROR)
# Some modules to help with reading the UCF101 dataset.
import random
import re
import os
import tempfile
import cv2
import numpy as np
# Some modules to display an animation using imageio.
import imageio
from IPython import display
from urllib import request # requires python3
# + cellView="both" id="IuMMS3TGdws7"
#@title Helper functions for the UCF101 dataset
# Utilities to fetch videos from UCF101 dataset
UCF_ROOT = "https://www.crcv.ucf.edu/THUMOS14/UCF101/UCF101/"
_VIDEO_LIST = None
_CACHE_DIR = tempfile.mkdtemp()
# As of July 2020, crcv.ucf.edu doesn't use a certificate accepted by the
# default Colab environment anymore.
unverified_context = ssl._create_unverified_context()
def list_ucf_videos():
"""Lists videos available in UCF101 dataset."""
global _VIDEO_LIST
if not _VIDEO_LIST:
index = request.urlopen(UCF_ROOT, context=unverified_context).read().decode("utf-8")
videos = re.findall("(v_[\w_]+\.avi)", index)
_VIDEO_LIST = sorted(set(videos))
return list(_VIDEO_LIST)
def fetch_ucf_video(video):
"""Fetchs a video and cache into local filesystem."""
cache_path = os.path.join(_CACHE_DIR, video)
if not os.path.exists(cache_path):
urlpath = request.urljoin(UCF_ROOT, video)
print("Fetching %s => %s" % (urlpath, cache_path))
data = request.urlopen(urlpath, context=unverified_context).read()
open(cache_path, "wb").write(data)
return cache_path
# Utilities to open video files using CV2
def crop_center_square(frame):
y, x = frame.shape[0:2]
min_dim = min(y, x)
start_x = (x // 2) - (min_dim // 2)
start_y = (y // 2) - (min_dim // 2)
return frame[start_y:start_y+min_dim,start_x:start_x+min_dim]
def load_video(path, max_frames=0, resize=(224, 224)):
cap = cv2.VideoCapture(path)
frames = []
try:
while True:
ret, frame = cap.read()
if not ret:
break
frame = crop_center_square(frame)
frame = cv2.resize(frame, resize)
frame = frame[:, :, [2, 1, 0]]
frames.append(frame)
if len(frames) == max_frames:
break
finally:
cap.release()
return np.array(frames) / 255.0
def to_gif(images):
converted_images = np.clip(images * 255, 0, 255).astype(np.uint8)
imageio.mimsave('./animation.gif', converted_images, fps=25)
return embed.embed_file('./animation.gif')
# + cellView="form" id="pIKTs-KneUfz"
#@title Get the kinetics-400 labels
# Get the kinetics-400 action labels from the GitHub repository.
KINETICS_URL = "https://raw.githubusercontent.com/deepmind/kinetics-i3d/master/data/label_map.txt"
with request.urlopen(KINETICS_URL) as obj:
labels = [line.decode("utf-8").strip() for line in obj.readlines()]
print("Found %d labels." % len(labels))
# + [markdown] id="GBvmjVICIp3W"
# # UCF101 데이터세트 사용하기
# + id="V-QcxdhLIfi2"
# Get the list of videos in the dataset.
ucf_videos = list_ucf_videos()
categories = {}
for video in ucf_videos:
category = video[2:-12]
if category not in categories:
categories[category] = []
categories[category].append(video)
print("Found %d videos in %d categories." % (len(ucf_videos), len(categories)))
for category, sequences in categories.items():
summary = ", ".join(sequences[:2])
print("%-20s %4d videos (%s, ...)" % (category, len(sequences), summary))
# + id="c0ZvVDruN2nU"
# Get a sample cricket video.
video_path = fetch_ucf_video("v_CricketShot_g04_c02.avi")
sample_video = load_video(video_path)
# + id="hASLA90YFPTO"
sample_video.shape
# + id="POf5XgffvXlD"
i3d = hub.load("https://tfhub.dev/deepmind/i3d-kinetics-400/1").signatures['default']
# + [markdown] id="mDXgaOD1zhMP"
# id3 모델을 실행하고 상위 5개의 행동 예측값을 출력합니다.
# + id="3mTbqA5JGYUx"
def predict(sample_video):
# Add a batch axis to the to the sample video.
model_input = tf.constant(sample_video, dtype=tf.float32)[tf.newaxis, ...]
logits = i3d(model_input)['default'][0]
probabilities = tf.nn.softmax(logits)
print("Top 5 actions:")
for i in np.argsort(probabilities)[::-1][:5]:
print(f" {labels[i]:22}: {probabilities[i] * 100:5.2f}%")
# + id="ykaXQcGRvK4E"
predict(sample_video)
# + [markdown] id="PHsq0lHXCsD4"
# 이제 https://commons.wikimedia.org/wiki/Category:Videos_of_sports에서 새 비디오를 시도해 봅니다.
#
# <NAME>의 [이 비디오](https://commons.wikimedia.org/wiki/File:End_of_a_jam.ogv)는 어떻습니까?
# + id="p-mZ9fFPCoNq"
# !curl -O https://upload.wikimedia.org/wikipedia/commons/8/86/End_of_a_jam.ogv
# + id="lpLmE8rjEbAF"
video_path = "End_of_a_jam.ogv"
# + id="CHZJ9qTLErhV"
sample_video = load_video(video_path)[:100]
sample_video.shape
# + id="2ZNLkEZ9Er-c"
to_gif(sample_video)
# + id="yskHIRbxEtjS"
predict(sample_video)
|
site/ko/hub/tutorials/action_recognition_with_tf_hub.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Make nice COS spectra plots
# +
import numpy as np
import matplotlib.pyplot as plt
import astropy.io.fits as fits
import os
import glob
from astropy.table import Table
from astropy.io import ascii
import astropy.units as u
import astropy.constants as const
from astropy.convolution import convolve, Box1DKernel
from scipy.optimize import leastsq
from scipy.interpolate import interp1d
from astropy.modeling import models, fitting
from astropy.io.votable import parse
from dust_extinction.parameter_averages import F99
#matplotlib set up
# %matplotlib inline
from matplotlib import rcParams
rcParams["figure.figsize"] = (14, 5)
rcParams["font.size"] = 20
#from cycler import cycler
#plt.rcParams['axes.prop_cycle'] = cycler(color=[plt.cm.plasma(r)])
# rcParams['image.cmap']='plasma'
# -
# path = '/media/david/5tb_storage1/cc_cet/hst/data/'
path = '/media/david/1tb_storage1/emergency_data/cc_cet/hst/data/'
x1ds = glob.glob(path+'*x1dsum.fits')
x1ds
smooth = 5
for i, x in enumerate(x1ds):
data = fits.getdata(x,1)
for dt in data:
w, f, e, dq = dt['WAVELENGTH'], dt['FLUX'], dt['ERROR'], dt['DQ']
mask = (f>0) & (dq == 0) & (w < 1214) | (w > 1217) & (f>0) & (dq == 0)
w, f, e = w[mask], f[mask], e[mask]
f = convolve(f,Box1DKernel(smooth))
e = convolve(e,Box1DKernel(smooth))/(smooth**0.5)
plt.plot(w, f+0.5*np.mean(f)*i)
plt.xlabel('Wavelength (\AA)')
plt.ylabel('Flux (erg s$^{-1}$ cm$^{-2}$ \AA$^{-1}$)')
# Just show one spectrum and a bunch of lines?
ism = Table.read('../../ism_lines.csv')
print(ism.dtype.names)
ism = ism[ism['line'] != 'SiIV']
# +
si2 = [1264.738]
si3 = [1294.545,1296.726,1298.892,1301.149,1303.323,1312.591]
si4 = [1393.775,1402.770]
plt.figure(figsize=(12, 6))
smooth = 5
x = x1ds[0]
data = fits.getdata(x,1)
w0, w1 = 10000, 0
for dt in data:
w, f, e, dq = dt['WAVELENGTH'], dt['FLUX'], dt['ERROR'], dt['DQ']
mask = (f>0) & (dq == 0) & (w < 1214) | (w > 1217) & (f>0) & (dq == 0)
w, f, e = w[mask], f[mask], e[mask]
if w[0] < w0:
w0= w[0]
if w[-1] > w1:
w1 = w[-1]
f = convolve(f,Box1DKernel(smooth))
e = convolve(e,Box1DKernel(smooth))/(smooth**0.5)
plt.plot(w, f, c='C0')
plt.xlabel('Wavelength (\AA)')
plt.ylabel('Flux (erg s$^{-1}$ cm$^{-2}$ \AA$^{-1}$)')
[plt.axvline(line, ls='--', c='C1', alpha=0.5) for line in ism['rest_lambda']]
plt.xlim(w0, w1)
plt.ylim(0.1e-14, 4.09e-13)
names = ['Si\,{\sc ii}', 'Si\,{\sc iii}', 'Si\,{\sc iv}']
lines = [si2, si3, si4]
for name, si in zip(names, lines):
[plt.annotate('',(line, 3.e-13), xytext=(line, 3.5e-13),arrowprops=dict(arrowstyle='-'), horizontalalignment='center') for line in si]
plt.annotate(name,(np.mean(si), 3.e-13), xytext=(np.mean(si), 3.6e-13), horizontalalignment='center')
#[plt.annotate('',(line, 3.5e-13), xytext=(line, 4e-13),arrowprops=dict(arrowstyle='-'), horizontalalignment='center') for line in si3]
#plt.annotate('Si\,{\sc iii}',(np.mean(si3), 3.5e-13), xytext=(np.mean(si3), 4.1e-13), horizontalalignment='center')
#[plt.annotate('',(line, 3.5e-13), xytext=(line, 4e-13),arrowprops=dict(arrowstyle='-'), horizontalalignment='center') for line in si4]
#plt.annotate('Si\,{\sc iv}',(np.mean(si4), 3.5e-13), xytext=(np.mean(si4), 4.1e-13), horizontalalignment='center')
plt.tight_layout()
# plt.savefig('plots/cc_cet_cos.pdf')
# -
# Looking for variation in the split spectra (try S iv lines)
# +
# smooth=10
# npath = '/media/david/5tb_storage1/cc_cet/hst/newx1ds/'
# nx1ds = glob.glob(npath+'*100*x1d.fits')
# data = fits.getdata(nx1ds[9], 1)[0]
# w, f, e = data['WAVELENGTH'], data['FLUX'], data['ERROR']
# f = convolve(f,Box1DKernel(smooth))
# e = convolve(e,Box1DKernel(smooth))/(smooth**0.5)
# plt.plot(w, f)
# plt.plot(w,e)
# plt.xlim(1380, 1420)
# plt.ylim(0, 3e-13)
# +
# times = []
# for x in nx1ds:
# hdr = fits.getheader(x,1)
# ti = (hdr['EXPSTART'] + hdr['EXPEND'])/2
# times.append(ti)
# args = np.argsort(np.array(times))
# nx1ds = np.array(nx1ds)[args]
# times = np.array(times)[args]
# -
"""from matplotlib.animation import FuncAnimation
smooth=50
fig, ax = plt.subplots(figsize=(5,5))
fig.set_tight_layout(True)
#ax[0].plot(t, f_lc)
#ax[0].set_xlabel('Time (s)')
#ax[0].set_ylabel('Flux (erg s$^{-1}$ cm$^{-2}$)')
#ax[0].set_ylim(0.4, 1.2)
ax.set_xlim(1380.1, 1414.9)
ax.set_ylim(1.11e-13, 2.09e-13)
#line, = ax[0].plot([0,0], [-0.1e-12,1.3e-12], 'C1--', linewidth=2)
ax.set_ylabel('Flux (erg s$^{-1}$ cm$^{-2}$ \AA$^{-1}$)')
ax.set_xlabel('Wavelength (\AA)')
ax.axvline(1393.775, ls='--', c='C1', alpha=0.5)
ax.axvline(1402.770, ls='--', c='C1', alpha=0.5)
#[ax[1].axvline(line, ls='--', c='r') for line in [8498.02,8542.09,8662.14]]
#ext = hdul[1::][0]
#dt = ext.data[0]
#w, f = dt['WAVELENGTH'], dt['FLUX']
w, f, e = np.array([], dtype=float), np.array([], dtype=float), np.array([], dtype=float)
#w, f, e = np.loadtxt(csv_files[0], unpack=True, delimiter=',')
line1, = ax.step(w,f, where='mid')
t0 = fits.getheader(nx1ds[0],1)['EXPSTART']
an = ax.annotate('', (0.75, 0.1), xycoords ='axes fraction')
obs = 1
def update(i):
#time = t[i]
#line.set_xdata([t[i], t[i]])
#ext = hdul[1::][i]
hdr = fits.getheader(nx1ds[i],1)
ti = (hdr['EXPSTART'] + hdr['EXPEND'])/2
if ti > 58152:
obs=2
else:
obs = 1
data = fits.getdata(nx1ds[i],1)[0]
w, f = data['WAVELENGTH'], data['FLUX']
f = convolve(f,Box1DKernel(smooth))
line1.set_xdata(w)
line1.set_ydata(f)
## if ti > t0+2:
# t0 = ti
# obs =
an.set_text('Ob {0}'.format(obs))
# print(ti)
return ax, line1, an
#ax.legend()
anim = FuncAnimation(fig, update, frames=np.arange(len(nx1ds)), interval=300)
anim.save('hst.gif', dpi=80, writer='imagemagick')
plt.show()
"""
# +
#gaia
p = 8.23807235942898e-3
pe = 0.07578241768233003e-3
d = 1/p
de = pe/p**2
print(d, de)
print(pe/p)
# +
#model
mw, mf = np.loadtxt('models/ldlc01010.dk', unpack=True, skiprows=34)
#plt.plot(mw, mf)
r = (0.0179*u.Rsun).to(u.m).value
dm = (d*u.pc).to(u.m).value
scale = (np.pi)*((r/dm)**2)*1e-8
print(scale)
plt.plot(mw, mf*scale)
# +
si2 = [1264.738]
si3 = [1294.545,1296.726,1298.892,1298.946, 1301.149,1303.323,1312.591]
si4 = [1393.775,1402.770]
c3 = [1174.935, 1175.265, 1175.592, 1175.713, 1175.713, 1175.989, 1176.372]
plt.figure(figsize=(12, 6))
smooth = 5
x = x1ds[0]
data = fits.getdata(x,1)
wb = np.array([], dtype=float)
fb = np.array([], dtype=float)
eb = np.array([], dtype=float)
for dt in data[::-1]:
w, f, e, dq = dt['WAVELENGTH'], dt['FLUX'], dt['ERROR'], dt['DQ']
mask = (f>0) & (dq == 0) & (w < 1214) | (w > 1217) & (f>0) & (dq == 0)
w, f, e = w[mask], f[mask], e[mask]
wb = np.concatenate((wb, w))
fb = np.concatenate((fb, f))
eb = np.concatenate((eb, e))
f = convolve(f,Box1DKernel(smooth))
e = convolve(e,Box1DKernel(smooth))/(smooth**0.5)
plt.plot(w, f, c='C0')
plt.xlabel('Wavelength (\AA)')
plt.ylabel('Flux (erg s$^{-1}$ cm$^{-2}$ \AA$^{-1}$)')
[plt.axvline(line, ls='--', c='C2', alpha=0.5) for line in ism['rest_lambda']]
plt.xlim(wb[0], wb[-1])
plt.ylim(0.1e-14, 3.89e-13)
names = ['Si\,{\sc ii}', 'Si\,{\sc iii}', 'Si\,{\sc iv}', 'C\,{\sc iii}']
lines = [si2, si3, si4, [np.mean(c3)]]
for name, si in zip(names, lines):
[plt.annotate('',(line, 2.7e-13), xytext=(line, 3.2e-13),arrowprops=dict(arrowstyle='-'), horizontalalignment='center') for line in si]
plt.annotate(name,(np.mean(si), 3.e-13), xytext=(np.mean(si), 3.3e-13), horizontalalignment='center', bbox=dict(facecolor='white', edgecolor='none'))
def residuals(scale, f, mf):
return f - mf/scale
mmask = (mw > wb[0]) & (mw < wb[-1])
mw1, mf1 = mw[mmask], mf[mmask]
mf1 = interp1d(mw1, mf1, fill_value='extrapolate')(wb)
alllines = np.hstack((si2, si3, si4, c3, ism['rest_lambda']))
C = np.zeros_like(wb,dtype='bool')
for a in alllines:
C |= (wb> a-0.5)&(wb <a+0.5)
mask = ~C
normfac = leastsq(residuals, 1., args=(fb[mask], mf1[mask]))[0]
print(normfac)
# define the model
ext = F99(Rv=3.1)
p = 8.23807235942898e-3
r = (0.0179*u.Rsun).to(u.m)
dm = ((1/p)*u.pc).to(u.m)
red = ext.extinguish(wb*u.AA, Ebv=0.021)
normfac1 = (1e8)/(np.pi*((r/dm)**2)*red)
print(normfac1)
normfac2 = leastsq(residuals, 1., args=(fb[mask], (mf1/normfac1)[mask]))[0]
print(normfac2)
plt.plot(wb, mf1/(normfac1*normfac2), c='C1', lw=2)
plt.tight_layout()
# plt.savefig('plots/cc_cet_cos.pdf', dpi=300)
# plt.savefig('plots/cc_cet_cos.png',dip=150, facecolor='white')
# -
"""x = x1ds[1]
data = fits.getdata(x,1)
for dt in data[::-1]:
w, f, e, dq = dt['WAVELENGTH'], dt['FLUX'], dt['ERROR'], dt['DQ']
plt.plot(w, f)
w2, f2, e2, dq2 = np.loadtxt('CC-Cet_ldlc51010.dat', unpack=True)
plt.plot(w2, f2)
plt.xlim(1300, 1350)
plt.ylim(0, 0.5e-12)"""
"""x = '/media/david/5tb_storage1/pceb_data/ldlc04010_x1dsum.fits'
data = fits.getdata(x,1)
rootname = fits.getheader(x, 0)['ASN_ID']
wb = np.array([], dtype=float)
fb = np.array([], dtype=float)
eb = np.array([], dtype=float)
dqb = np.array([], dtype=int)
for dt in data[::-1]:
w, f, e, dq = dt['WAVELENGTH'], dt['FLUX'], dt['ERROR'], dt['DQ']
# mask = (f>0) & (dq == 0) & (w < 1214) | (w > 1217) & (f>0) & (dq == 0)
#w, f, e = w[mask], f[mask], e[mask]
wb = np.concatenate((wb, w))
fb = np.concatenate((fb, f))
eb = np.concatenate((eb, e))
dqb = np.concatenate((dqb, dq))
savdat = Table([wb, fb, eb, dqb], names=['#WAVELENGTH', 'FLUX', 'ERROR', 'DQ'])
ascii.write(savdat, 'LM-COM_'+rootname.lower()+'.dat', format='basic', overwrite=True)"""
# Making plots with the magnetic models
# +
mods = glob.glob('magnetic_models/*1400*.dat')
mods.sort()
print(mods)
# -
# Adding another spectrum to compare with
"""def make_plot_spec(w, f, e, mask1, mask2): #cuts spectrum down to the bit to plot
fitter = fitting.LinearLSQFitter()
#mask = (w > 8450) & (w < 8480) | (w > 8520) & (w <8540) | (w > 8560) & (w< 8660) | (w > 8680) & (w < 8700) #mask out emmission lines
w1, f1 = w[mask1], f[mask1]
n_init = models.Polynomial1D(3)
n_fit = fitter(n_init, w1, f1)
#mask = (w > 8450) & (w < 8700)
w1, f1, e1 = w[mask2], f[mask2], e[mask2]
nf = f1/n_fit(w1)
ne = e1/n_fit(w1)
smooth = 5
nf = convolve(nf,Box1DKernel(smooth))
ne = convolve(ne,Box1DKernel(smooth))/smooth**0.5
return w1,nf, ne
wc, fc, ec, dqc = np.loadtxt('LM-COM_ldlc04010.dat', unpack=True) #picking lm com for now, might change!
mask2 = (wc > 1390) & (wc < 1410)
mask1 = (wc > 1390) & (wc < 1392) | (wc > 1395) & (wc < 1401) | (wc > 1405) & (wc < 1410)
wn, fn, en = make_plot_spec(wc, fc, ec, mask1, mask2)
plt.plot(wn, fn)"""
# +
mods = ['magnetic_models/lmcom-1400-0kG-plot.dat', 'magnetic_models/cccet-1400-B710kG-40kms-02-plot.dat', 'magnetic_models/cccet-1400-B630kG-40kms-01-plot.dat']
si4 = [1393.775,1402.770]
dates = ['LM Com \n 2017~December~17','CC\,Cet \n 2018~July~22', 'CC\,Cet \n 2018~February~01']
Bs = [100, 710, 630]
plt.figure(figsize = (9, 12))
for i, mod in enumerate(mods):
w, f, m = np.loadtxt(mod, unpack=True)
f = convolve(f,Box1DKernel(5))
if i == 0:
mask = (w < 1393.280) | (w > 1393.310)
w, f, m = w[mask], f[mask], m[mask]
plt.plot(w,f+0.5*i, c='C0')
plt.plot(w, m+0.5*i, lw=2, c='C1')
#if i == 0:
#[plt.annotate('',(line, 2.7e-13), xytext=(line, 3.2e-13),arrowprops=dict(arrowstyle='-'), horizontalalignment='center') for line in si4]
# plt.xticks(visible=False)
plt.xlim(1390.1, 1408.9)
plt.ylabel('Normalised Flux')
if i == 1:
plt.xlabel('Wavelength (\AA)')
plt.annotate(dates[i], (0.3, 0.75+(0.5*i)), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
if i > 0:
plt.annotate(r'$\langle \vert B \vert \rangle = {}$\,kG'.format(Bs[i]), (0.75, 0.75+(0.5*i)), xycoords = ('axes fraction', 'data'))
else:
plt.annotate(r'$\langle \vert B \vert \rangle <$ {}\,kG'.format(Bs[i]), (0.75, 0.75+(0.5*i)), xycoords = ('axes fraction', 'data'))
plt.ylim(0.45, 2.19)
#plt.plot(wn, fn+1)
[plt.axvline(line, ls='--', c='C2', alpha=0.5) for line in si4]
[plt.annotate('Si\,{\sc iv}',(line, 1), xytext=(line, 2.1), horizontalalignment='center', bbox=dict(facecolor='white', edgecolor='none')) for line in si4]
#plt.annotate('LM Com', (0.75, 1.8), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
plt.tight_layout()
plt.subplots_adjust(hspace=0.02)
plt.savefig('plots/siiv_lines.pdf', dpi=300)
plt.savefig('plots/siiv_lines.png', dpi=150, facecolor='white')
#plt.show()
# -
#
mods = glob.glob('magnetic_models/cccet*1300*.dat')
#mods.sort()
mods = mods[::-1]
print(mods)
# +
dates = ['2018~July~22', '2018~February~01']
Bs = [710, 630]
si3 = [1294.545,1296.726,1298.892,1298.946,1301.149,1303.323]#,1312.591]
plt.figure(figsize = (10, 10))
for i, mod in enumerate(mods):
w, f, m = np.loadtxt(mod, unpack=True)
f = convolve(f,Box1DKernel(5))
plt.plot(w,f+0.5*i, c='C0')
plt.plot(w, m+0.5*i, lw=2, c='C1')
plt.xlim(1292.1, 1307.9)
plt.ylabel('Normalised Flux')
if i == 1:
plt.xlabel('Wavelength (\AA)')
plt.annotate(dates[i], (0.02, 0.67+(0.55*i)), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
plt.annotate(r'$\langle \vert B \vert \rangle <$ {}\,kG'.format(Bs[i]), (0.77, 0.67+(0.55*i)), xycoords = ('axes fraction', 'data'))
plt.ylim(0.61, 1.69)
#plt.plot(wn, fn+1)
[plt.axvline(line, ls='--', c='C2', alpha=0.5) for line in si3]
[plt.annotate('Si\,{\sc iii}',(line, 1), xytext=(line, 1.6), horizontalalignment='center', bbox=dict(facecolor='white', edgecolor='none')) for line in si3]
#plt.annotate('LM Com', (0.75, 1.8), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
plt.tight_layout()
plt.subplots_adjust(hspace=0.02)
plt.savefig('plots/nolm_siiii_lines.pdf')
# -
# See what it looks like with LM com as well
# +
mods = ['magnetic_models/lmcom-1300-0kG-plot.dat','magnetic_models/cccet-1300-B710kG-40kms-02-plot.dat', 'magnetic_models/cccet-1300-B630kG-40kms-01-plot.dat']
dates = ['LM Com \n 2017~December~17','CC\,Cet \n 2018~July~22', 'CC\,Cet \n 2018~February~01']
Bs = [100, 710, 630]
plt.figure(figsize = (9, 12))
for i, mod in enumerate(mods):
w, f, m = np.loadtxt(mod, unpack=True)
f = convolve(f,Box1DKernel(5))
if i == 0:
mask = (w < 1393.280) | (w > 1393.310)
w, f, m = w[mask], f[mask], m[mask]
plt.plot(w,f+0.5*i, c='C0')
plt.plot(w, m+0.5*i, lw=2, c='C1')
#if i == 0:
#[plt.annotate('',(line, 2.7e-13), xytext=(line, 3.2e-13),arrowprops=dict(arrowstyle='-'), horizontalalignment='center') for line in si4]
# plt.xticks(visible=False)
plt.xlim(1292.1, 1307.9)
plt.ylabel('Normalised Flux')
if i == 1:
plt.xlabel('Wavelength (\AA)')
if i > 0:
plt.annotate(r'$\langle \vert B \vert \rangle = {}$\,kG'.format(Bs[i]), (0.75, 0.7+(0.5*i)), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
plt.annotate(dates[i], (0.02, 0.65+(0.5*i)), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
else:
plt.annotate(r'$\langle \vert B \vert \rangle <$ {}\,kG'.format(Bs[i]), (0.75, 0.65+(0.5*i)), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
plt.annotate(dates[i], (0.02, 0.6+(0.5*i)), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
plt.ylim(0.45, 2.19)
#plt.plot(wn, fn+1)
[plt.axvline(line, ls='--', c='C2', alpha=0.5) for line in si3]
[plt.annotate('Si\,{\sc iii}',(line, 1), xytext=(line, 2.1), horizontalalignment='center', bbox=dict(facecolor='white', edgecolor='none')) for line in si3]
#plt.annotate('LM Com', (0.75, 1.8), xycoords = ('axes fraction', 'data'), bbox=dict(facecolor='white', edgecolor='none'))
plt.tight_layout()
#plt.subplots_adjust(hspace=0.02)
plt.savefig('plots/siiii_lines.pdf', dpi=300)
plt.savefig('plots/siiii_lines.png', dpi=150, facecolor='white')
# -
# Saving a scaled model for use in the COS etc.
mmask = (mw > 950) & (mw < 1600) #safe side to overlap g130m
mws, mfs = mw[mmask], mf[mmask]/normfac
plt.plot(mws, mfs)
savdat = Table([mws, mfs], names=['#WAVELENGTH', 'FLUX'])
ascii.write(savdat, 'models/CC_CET_scaled_fuv_model.dat', format='basic', overwrite=True)
# 20210119 what are the lines around 1140?
x = x1ds[0]
data = fits.getdata(x,1)
wb = np.array([], dtype=float)
fb = np.array([], dtype=float)
eb = np.array([], dtype=float)
for dt in data[::-1]:
w, f, e, dq = dt['WAVELENGTH'], dt['FLUX'], dt['ERROR'], dt['DQ']
mask = (f>0) & (dq == 0) & (w < 1214) | (w > 1217) & (f>0) & (dq == 0)
w, f, e = w[mask], f[mask], e[mask]
wb = np.concatenate((wb, w))
fb = np.concatenate((fb, f))
eb = np.concatenate((eb, e))
f = convolve(f,Box1DKernel(smooth))
e = convolve(e,Box1DKernel(smooth))/(smooth**0.5)
plt.plot(w, f, c='C0')
plt.xlim(1130, 1150)
# Can the M4 emmmison lines poke through? Compare with GJ1214 (M5V from MUSCLES).
muspath = '/media/david/5tb_storage1/mast_muscles/gj1214/'
spec = 'hlsp_muscles_multi_multi_gj1214_broadband_v22_adapt-var-res-sed.fits'
gjdat = fits.getdata(muspath+spec)
gjw, gjf = gjdat['WAVELENGTH'], gjdat['FLUX']
plt.plot(gjw, gjf)
# +
gjmask = (gjw > w[0]) & (gjw < w[-1])
gjw1, gjf1 = gjw[gjmask], gjf[gjmask]
gj_d = 14.65*u.pc
cc_d = 121.4*u.pc
scale = (gj_d/cc_d)**2
scale
# -
plt.plot(w,f)
plt.plot(gjw1, gjf1*scale)
# Making an SED. First scale a template spectrum.
# +
tphot = 'cc_cet_vizier_votable.vot'
c = 2.998e8*u.m/u.s
votable = parse(tphot)
table = votable.get_first_table()
data = table.array
mask = ~data['sed_eflux'].mask
masked_data = data[mask].data
filters = np.unique(masked_data['sed_filter'].data)
wp = []
fp = []
ep = []
#print(filters)
# filters = [b'2MASS:H', b'2MASS:J', b'2MASS:Ks', b'GALEX:FUV',
# b'GALEX:NUV', b'Gaia:G', b'Johnson:B', b'Johnson:H', b'Johnson:J',
# b'Johnson:K', b'Johnson:V', b'PAN-STARRS/PS1:g', b'PAN-STARRS/PS1:i',
# b'PAN-STARRS/PS1:r', b'PAN-STARRS/PS1:y', b'PAN-STARRS/PS1:z', b"SDSS:g'",
# b"SDSS:r'", b'WISE:W1', b'WISE:W2'] #picking my own
# filters = [b'2MASS:H', b'2MASS:J', b'2MASS:Ks', b'GALEX:FUV',
# b'GALEX:NUV', b'PAN-STARRS/PS1:g', b'PAN-STARRS/PS1:i',
# b'PAN-STARRS/PS1:r', b'PAN-STARRS/PS1:y', b'PAN-STARRS/PS1:z']#, b'WISE:W1', b'WISE:W2'] #picking my own
filters = [b'GALEX:FUV',
b'GALEX:NUV', b'PAN-STARRS/PS1:g', b'PAN-STARRS/PS1:i',
b'PAN-STARRS/PS1:r', b'PAN-STARRS/PS1:y', b'PAN-STARRS/PS1:z']
#filters = [b'GALEX:NUV']
for flt in filters:
w1 = (np.mean(masked_data['sed_freq'][masked_data['sed_filter']==flt])*u.GHz).to(u.AA, equivalencies=u.spectral())
fj1 = masked_data['sed_flux'][masked_data['sed_filter']==flt]
e1 = masked_data['sed_eflux'][masked_data['sed_filter']==flt]
if len(fj1) >1:
fj_av = np.average(fj1, weights = (1/(e1**2)))
e1_av = abs(np.average((fj1-fj_av), weights = (1/(e1**2))))**0.5
e1_av = 1 / np.sum(1/(e1**2), axis=0)**0.5
else:
fj_av, e1_av = fj1[0], e1[0]
f1 = (fj_av*u.Jy).to(u.erg / u.cm**2 / u.s / u.AA, equivalencies=u.spectral_density(w1))
wp.append(w1.value)
fp.append(f1.value)
e1 = ((e1_av*f1)/fj_av).value
ep.append(e1)
wp, fp, ep = np.array(wp), np.array(fp), np.array(ep)
# +
temp_path = '/media/david/5tb_storage1/pyhammer/PyHammer-2.0.0/resources/templates/'
specs = glob.glob('{}M4_+0.0_Dwarf.fits'.format(temp_path))
data = fits.getdata(specs[0])
tempscale= 3e-16
wt, ft = 10**data['Loglam'], data['Flux']*tempscale
mwt, mft = mw[mw >wt[0]], mf[mw > wt[0]]/normfac
plt.plot(wt, ft*tempscale)
plt.plot(mwt, mft)
ftm = interp1d(wt, ft, fill_value='extrapolate')(mwt)
plt.plot(mwt, ftm)
plt.errorbar(wp[ep>0], fp[ep>0], yerr=ep[ep>0], marker='o', ls='none', c='C0')
plt.xlim(mwt[0], mwt[-1])
plt.ylim(0, 0.6e-14)
com_f = mft+ftm
plt.plot(mwt, com_f)
specs = glob.glob('{}M5_+0.0_Dwarf.fits'.format(temp_path))
data = fits.getdata(specs[0])
tempscale= 3e-16
wt, ft = 10**data['Loglam'], data['Flux']*tempscale
mwt, mft = mw[mw >wt[0]], mf[mw > wt[0]]/normfac
plt.plot(wt, ft*tempscale)
ftm = interp1d(wt, ft, fill_value='extrapolate')(mwt)
plt.plot(mwt, ftm)
com_f = mft+ftm
plt.plot(mwt, com_f)
# +
plt.figure(figsize=(12,6))
plt.plot(mw, mf/normfac, c='C1', zorder=10)
# plt.plot(w,f)
x = x1ds[0]
data = fits.getdata(x,1)
wb = np.array([], dtype=float)
fb = np.array([], dtype=float)
eb = np.array([], dtype=float)
for dt in data[::-1]:
w, f, e, dq = dt['WAVELENGTH'], dt['FLUX'], dt['ERROR'], dt['DQ']
mask = (f>0) & (dq == 0) & (w < 1214) | (w > 1217) & (f>0) & (dq == 0)
w, f, e = w[mask], f[mask], e[mask]
wb = np.concatenate((wb, w))
fb = np.concatenate((fb, f))
eb = np.concatenate((eb, e))
f = convolve(f,Box1DKernel(smooth))
e = convolve(e,Box1DKernel(smooth))/(smooth**0.5)
plt.plot(w, f, c='C0')
# plt.plot(gjw, gjf)
plt.yscale('log')
plt.xscale('log')
plt.xlim(1000, 50000)
plt.errorbar(wp[ep>0], fp[ep>0], yerr=ep[ep>0], marker='o', ls='none', c='C3')
plt.xlim(1051, 9999)
plt.ylim(1e-16)
uves_path = '/media/david/5tb_storage1/cc_cet/uves/'
dats = glob.glob('{}/*.dat'.format(uves_path))
dat = dats[0]
w, f, e = np.loadtxt(dat, unpack=True)
f = convolve(f,Box1DKernel(20))
plt.plot(w[5:-6], f[5:-6])
temp_path = '/media/david/5tb_storage1/pyhammer/PyHammer-2.0.0/resources/templates/'
specs = glob.glob('{}M5_+0.0_Dwarf.fits'.format(temp_path))
data = fits.getdata(specs[0])
tempscale= 1e-15
plt.plot(10**data['Loglam'], data['Flux']*tempscale,c='C3')
#plt.yscale('log')
#plt.xscale('log')
plt.ylabel('Flux (erg s$^{-1}$ cm$^{-2}$ \AA$^{-1}$)')
plt.xlabel('Wavelength (\AA)')
#plt.plot(mw[mw>1400] , mf[mw>1400]/normfac)
#plt.xlim(1200, 60000)
#plt.xticks((2000, 10000, 40000), ('2000', '10000', '40000'))
plt.tight_layout()
#plt.axvline(2311)
#plt.savefig('plots/cc_cet_phot.png', dpi=150, facecolor='white')
plt.plot(mwt, com_f)
# -
# Huh. Plot of the Si 1264 line
# +
def make_plot_model(w, f, e, mask1, mask2): #cuts spectrum down to the bit to plot
fitter = fitting.LinearLSQFitter()
w1, f1, m1 = w[mask1], f[mask1], m[mask1]
n_init = models.Polynomial1D(3)
n_fit = fitter(n_init, w1, f1)
n_fit2 = fitter(n_init, w1, m1)
#mask = (w > 8450) & (w < 8700)
w1, f1, m1 = w[mask2], f[mask2], m[mask2]
nf = f1/n_fit(w1)
nm = m1/n_fit2(w1)
return w1,nf,nm
mods = ['magnetic_models/plot-obs-model-si2-1265.dat']
# dates = ['2018~July~22', '2018~February~01']
# Bs = [710, 630]
# si3 = [1294.545,1296.726,1298.892,1301.149,1303.323]#,1312.591]
def picomps(l0, B, z=1.43):
"""
Returns the pi components of a zeeman-split line l0 for magnetic field B
"""
dl = 4.67e-13 * z* l0**2 * B
return [l0-dl, l0+dl]
si2 = [1264.73]
pis = picomps(si2[0], 450e3)
print(pis)
fig, ax = plt.subplots(figsize = (10, 7))
for i, mod in enumerate(mods):
w, f, m = np.loadtxt(mod, unpack=True)
mask1 = (w > 1262) & (w < 1263.5) | (w > 1266) & (w < 1267)
mask2 = (w> 1261) & (w < 1269)
w, f, m = make_plot_model(w, f,m , mask1, mask2)
f = convolve(f,Box1DKernel(5))
plt.plot(w,f+0.5*i, c='C0')
plt.plot(w, m+0.5*i, lw=2, c='C1')
#pidispaly = ax.transData.transform(pis[1]-pis[0])
#print(pidisplay)
#[plt.axvline(line, ls='--', c='C2', alpha=0.5) for line in si2]
#[plt.axvline(line, ls='--', c='C2', alpha=0.5) for line in pis]
plt.annotate('Si\,{\sc ii}\n$\pi$', (si2[0], 1.09),xytext=(si2[0], 1.13), arrowprops=dict(arrowstyle='-[,widthB=2.6,lengthB=1.2') , ha='center')
plt.annotate('$\sigma-$', (pis[0], 1.05), ha='center')
plt.annotate('$\sigma+$', (pis[1], 1.05), ha='center')
plt.xlim(1262.1, 1267.9)
plt.ylim(0.76, 1.19)
plt.ylabel('Normalised Flux')
plt.xlabel('Wavelength (\AA)')
plt.tight_layout()
plt.savefig('plots/siii_lines.pdf')
# -
t1 = 25245.
t1e=18.5
t2=25162.
t2e=19.5
paper = 25203
paper_e = 42
wm = ((t1/t1e) + (t2/t2e))/((1/t1e)+(1/t2e))
print('value in paper', '{}+/-{}'.format(paper, paper_e))
print('weighted mean',wm)
print('mean', np.mean([t1,t2]))
print('std', np.std([t1, t2]))
|
cc_cet/cc_cet_hst.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
# %matplotlib inline
pd.options.display.float_format = '{:.2f}'.format
#Read excel dataset
df = pd.read_excel('timeseries_covid19_us_confirmed.xlsx',index_col='date',parse_dates=True)
#since US is irrelevant in this dataframe
df = df.drop(columns=['country'])
df.head()
# -
df.tail()
train_data = df.iloc[:242] # Goes up to but not including 242
#train_data = train_data.astype('double')
test_data = df.iloc[242:]
#test_data = test_data.astype('double')
test_data.index
# +
from statsmodels.tsa.holtwinters import ExponentialSmoothing
import warnings
warnings.filterwarnings("ignore")
fitted_model = ExponentialSmoothing(train_data['total'],trend='mul',seasonal='mul',seasonal_periods=7).fit()
# -
test_predictions = fitted_model.forecast(60).rename('Case Forecast')
test_predictions
train_data['total'].plot(legend=True,label='TRAIN')
test_data['total'].plot(legend=True,label='TEST',figsize=(15,10))
test_predictions.plot(legend=True,label='PREDICTION');
train_data['total'].plot(legend=True,label='TRAIN')
test_data['total'].plot(legend=True,label='TEST',figsize=(15,10))
#let's zoom into the period we were predicting for
test_predictions.plot(legend=True,label='PREDICTION',xlim=['9/17/2020','11/20/2020']);
from sklearn.metrics import mean_squared_error,mean_absolute_error
mean_absolute_error(test_data,test_predictions)
mean_squared_error(test_data,test_predictions)
np.sqrt(mean_squared_error(test_data,test_predictions))
test_data.describe() #we want to compare MSE with Avergae of Test Data, or RMSE with STD of overall True Data
# ## Forecasting into Future
final_model = ExponentialSmoothing(df['total'],trend='mul',seasonal='mul',seasonal_periods=7).fit()
forecast_predictions = final_model.forecast(43)#forecast to December 31,2020
forecast_predictions
df['total'].plot(figsize=(12,8))
forecast_predictions.plot(legend=True,label='PREDICTION');
|
Forecasting The Future.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Module 1 Required Coding Activity
#
# Introduction to Python (Unit 2) Fundamentals
#
# | Important Assignment Requirements |
# |:-------------------------------|
# | **NOTE:** This program **requires** **`print`** output and using code syntax used in module 1 such as keywords **`for`**/**`in`** (iteration), **`input`**, **`if`**, **`else`**, **`.isalpha()`** method, **`.lower()`** or **`.upper()`** method |
#
#
# ## Program: Words after "G"/"g"
# Create a program inputs a phrase (like a famous quotation) and prints all of the words that start with h-z
#
# Sample input:
# `enter a 1 sentence quote, non-alpha separate words:` **`Wheresoever you go, go with all your heart`**
#
# Sample output:
# ```
# WHERESOEVER
# YOU
# WITH
# YOUR
# HEART
# ```
# 
#
# - split the words by building a placeholder variable: **`word`**
# - loop each character in the input string
# - check if character is a letter
# - add a letter to **`word`** each loop until a non-alpha char is encountered
#
# - **if** character is alpha
# - add character to **`word`**
# - non-alpha detected (space, punctuation, digit,...) defines the end of a word and goes to **`else`**
# - **`else`**
# - check **`if`** word is greater than "g" alphabetically
# - print word
# - set word = empty string
# - or **else**
# - set word = empty string and build the next word
#
# Hint: use `.lower()`
#
# Consider how you will print the last word if it doesn't end with a non-alpha character like a space or punctuation?
#
#
# +
# [] create words after "G" following the Assignment requirements use of functions, menhods and kwyowrds
# sample quote "Wheresoever you go, go with all your heart" ~ Confucius (551 BC - 479 BC)
# [] copy and paste in edX assignment page
# -
# Submit this by creating a python file (.py) and submitting it in D2L. Be sure to test that it works.
|
Python Fundamentals/Module_1_Required_Code_Python_Fundamentals.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/joshuajhchoi/Real-Time-Voice-Cloning/blob/master/WGAN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_4ZcUUel91ps"
# # Wasserstein GAN
# + [markdown] id="VEIeFkRj91pt"
# [WGAN](https://arxiv.org/abs/1701.07875) is an important version of GANs. It's based on the Earthmover distance distance which is an optimization problem that solve transportation problem.
#
# GAN duty is to learn a probability distribution, but it's job become harder if we are dealing with distributions supported by low dimensional manifolds. It is then unlikely that the model manifold and the true distribution’s support have a non-negligible intersection.
#
# The solution is to treat each point in the noise as a discret probability distribution and then calculate the Euclidean distance between the fake point and real point in the real data distribution. The objective is to minimize the cost of the sum of the distance between points in real and fake data distribution. With Earthmover distance, we get point-by-point details about how each data point affects the output. This kind of attention to detail is very important in our situation.
#
#
# 
# 
#
# The diffrence between GAN and WGAN is that the discriminator predicts the probability of a generated image being “real” and the critic model scores the “realness” of a given image.
# + [markdown] id="e-Y3rNA191pt"
# [click for more ressources](https://jeremykun.com/2018/03/05/earthmover-distance/)
# + [markdown] id="MYZMQu3e91pu"
# ### Learning Objectives
# + [markdown] id="cmjHLR8291pv"
# - Learn how to train WGAN
# - Learning the implementation details for WGAN
# - Knowing the diffrence between a discriminator predicting a probability and a critic predicting a score
# + [markdown] id="F_XR9o0TA8L_"
# ### Practical steps we will follow
#
# 1. Explore data and prepare the dataset for training
# 3. Define and Implement the Generator and the discriminator Network Architecture
# 4. Work on the training/ evaluation loop and generate new realistic samples using WGAN
#
#
# + [markdown] id="Kjb-uNtPQZJL"
# ### Steps to run this notebook from Colaboratory
#
# This colab will run much faster on GPU. To use a Google Cloud
# GPU:
#
# 1. Go to `Runtime > Change runtime type`.
# 1. Click `Hardware accelerator`.
# 1. Select `GPU` and click `Save`.
# 1. Click `Connect` in the upper right corner and select `Connect to hosted runtime`.
# + [markdown] id="hQ3wCzCA91pv"
# ### Imports (RUN ME!)
# + [markdown] id="9HTCfyuw91pw"
# Make sure to run the imports cell above, otherwise the rest of the cells will fail when you try to run them. (To run a cell press `shift` + `enter` with your mouse cursor in the cell or press the play button in the top right of the cell.)
# + id="KDvyG-5091px"
# %load_ext tensorboard
import os
from glob import glob
import time
import random
import IPython.display as display
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import PIL
from PIL import Image
import imageio
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras.initializers import RandomNormal
# %matplotlib inline
# + [markdown] id="d2GpQofxEpLP"
# ### Mount Drive
# + id="_e7u2b2AEuln" outputId="a60bc2b1-1160-415b-cebe-c197017cf113" colab={"base_uri": "https://localhost:8080/", "height": 33}
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="dd11WdlH91p1"
# ### Configs
# + id="U92L25aQ91p1"
# Experiment paths
# Save the model for further use
EXPERIMENT_ID = "train_wgan"
MODEL_SAVE_PATH = os.path.join("/content/drive/My Drive/lecture hands on lab/wgan/results", EXPERIMENT_ID)
if not os.path.exists(MODEL_SAVE_PATH):
os.makedirs(MODEL_SAVE_PATH)
CHECKPOINT_DIR = os.path.join(MODEL_SAVE_PATH, 'training_checkpoints')
# Data path
DATA_PATH = "/content/drive/My Drive/lecture hands on lab/datasets/cars/cars_images/"
# Model parameter
BATCH_SIZE = 64
EPOCHS = 9000
LATENT_DEPTH = 100
IMAGE_SHAPE = [100,100]
NB_CHANNELS = 3
LR = 1e-4
BETA = 0.5
NOISE = tf.random.normal([1, LATENT_DEPTH])
# Parameters for THE CRITIC
N_CRITIC = 5
CLIPPING_WEIGHT = 0.01
# A fixed random seed is a common "trick" used in ML that allows us to recreate
# the same data when there is a random element involved.
seed = random.seed(30)
# + [markdown] id="drVU_E4Z91p4"
# ### Data understanding and exploration
# + [markdown] id="bPQfQvA391p5"
# We will be using the same dataset [cars dataset](https://www.kaggle.com/prondeau/the-car-connection-picture-dataset) as DCGAN to see the diffrence more clearly.
# + id="0Hj-OWUABigv" outputId="862fb719-5ed9-4488-bcd6-83530cd52307" colab={"base_uri": "https://localhost:8080/", "height": 34}
image_count = len(list(glob(str( DATA_PATH + '*.jpg'))))
image_count
# + id="n24GGNgK91p6" outputId="e0f17517-08df-43da-c270-b27f2c0ee9d5" colab={"base_uri": "https://localhost:8080/", "height": 737}
cars_images_path = list(glob(str(DATA_PATH + '*.jpg')))
for image_path in cars_images_path[:3]:
display.display(Image.open(str(image_path)))
# + id="uhYIMU7vBfR_"
# Our goal to get images filenames
images_name = [i.split(DATA_PATH) for i in cars_images_path]
images_name = [x[:][1] for x in images_name]
cars_model = [i.split('_')[0] for i in images_name]
# + id="dyQ6xEvqBm2y"
# Extract cars models
def unique(list1):
list_set = set(list1)
unique_list = (list(list_set))
return unique_list
# + id="uBRSzqexBoRZ" outputId="ec3836eb-1174-4ef2-ce65-641e477c554e" colab={"base_uri": "https://localhost:8080/", "height": 357}
unique_cars = unique(cars_model)
unique_cars
# + id="X6v_qgGBBpnB" outputId="b22a1555-624b-4b5d-c99e-e10b53615950" colab={"base_uri": "https://localhost:8080/", "height": 606}
plt.figure(figsize=(20,10))
plt.hist(cars_model, color = "blue", lw=0, alpha=0.7)
plt.ylabel('images number')
plt.xlabel('car model')
plt.show()
# + id="BX8K1NB7Brgl"
image_size = []
for filename in cars_images_path:
im=Image.open(filename)
im =im.size
image_size.append(im)
print(max(image_size))
print(min(image_size))
# + id="nTS_R-EiBtIA"
# Read in the image
image = mpimg.imread(cars_images_path[20])
plt.axis("off")
plt.imshow(image)
# + id="rCF31vNvBulQ"
# Isolate RGB channels
r = image[:,:,0]
g = image[:,:,1]
b = image[:,:,2]
# Visualize the individual color channels
f, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,10))
ax1.set_title('R channel')
ax1.imshow(r, cmap='gray')
ax2.set_title('G channel')
ax2.imshow(g, cmap='gray')
ax3.set_title('B channel')
ax3.imshow(b, cmap='gray')
# + [markdown] id="TK9rOsRa91p6"
# The dataset contains 8960 images:
# * Three-channel each one.
# * Diverse styles
# * the minimum shape is (320, 124)
# * the maximum shape is (320, 360)
# + [markdown] id="ZCidwOSA91p-"
# ### Data Loader
# + [markdown] id="xGEDsmng91p_"
# For the Data pipeline, we will be using Data API provide it by TensorFlow. What we have to do is create a dataset object, tell it where to get the data, then transform it in any way we want, and TensorFlow takes care of all the implementation details, such as multithreading, queuing, batching, prefetching, and so on.\
# (1) Create dataset entirely in RAM using tf.data.Dataset.from_tensor_slices() \
# (2) Call shuffle to ensure that the training set are independent and identically distributed. \
# NB: the buffer size must be specified, and it is important to make it large enough or else shuffling will not be very efficient.\
# (3) Do the necessary transformation by calling the map() method. \
# (4) Call the batch() method. It will group the items of the previous dataset in batches of n items. \
# (5) Using prefetch will let the dataset work on parallel with training algorithm to get the next batch ready.
# 
#
#
# We must scale the pixel values from the range of unsigned integers in [0,255] to the normalized range of [-1,1] or [0,1]. But you should know that if you're using [-1,1] you need to choose tanh as an activation function and if you are normalizing to [0,1] you use sigmoid activation function.
# + [markdown] id="ytZ-dgAGB_yY"
# ### Data Loader using TF API
# + id="2RGpWU7S91qB"
@tf.function
def preprocessing_data(path):
image = tf.io.read_file(path)
image = tf.image.decode_jpeg(image, channels=3)
image = tf.image.resize(image, [IMAGE_SHAPE[0],IMAGE_SHAPE[1]])
image = image / 255.0
return image
# + id="s2vWihAh91qE"
def dataloader(paths):
dataset = tf.data.Dataset.from_tensor_slices(paths)
dataset = dataset.shuffle(buffer_size=len(paths))
dataset = dataset.map(preprocessing_data)
dataset = dataset.batch(10* BATCH_SIZE)
dataset = dataset.prefetch(1)
return dataset
# + id="6PCX7jz791qH"
dataset = dataloader(cars_images_path)
for batch in dataset.take(1):
for img in batch:
img_np = img.numpy()
plt.figure()
plt.axis('off')
plt.imshow((img_np-img_np.min())/(img_np.max()-img_np.min()))
# + [markdown] id="RPeLxdC-91qL"
# # Modeling
# + [markdown] id="2pbAfHvm91qM"
# 
#
# The differences in implementation for the WGAN and DCGAN are as follows:
#
# 1. Adding a linear activation function in the output layer of the critic model or remove sigmoid function if you're using it.
#
# 2. Use Wasserstein loss to train the critic and generator models that promote larger difference between scores for real and generated images.
# 3. Adding a constraint to limit the weight range after each mini batch update (e.g. [-0.01,0.01] like the paper).
# 4. Update the critic model more times than the generator each iteration (e.g. 5 like the paper).
# 5. Use the RMSProp version of gradient descent with small learning rate and no momentum (e.g. 0.00005 like the paper).
#
# Recommandation: Use always the paper hyperparameter to get a baseline model and then built on it the changes
# + [markdown] id="BUUEF-7191qM"
# ### Generator
# + [markdown] id="Hmau8JZf91qN"
# The generator model is responsible for creating new, fake, but plausible small photographs of objects.
# It does this by taking a point from the latent space as input and outputting a square color image and this will not change in WGAN.
# 
# + id="Kl1CzCFH91qO"
def make_generator_model():
#weight initialization
#init = RandomNormal(stddev=0.02)
model = tf.keras.Sequential()
model.add(layers.Dense(25*25*128, use_bias=False, input_shape=(100,))) # add kernel_initializer=init in case of weight initialization
model.add(layers.BatchNormalization())
model.add(layers.ReLU())
model.add(layers.Reshape((25, 25, 128)))
assert model.output_shape == (None, 25, 25, 128) # Note: None is the batch size
model.add(layers.Conv2DTranspose(128, (5, 5), strides=(1, 1), padding='same', use_bias=False))# add kernel_initializer=init in case of weight initialization
assert model.output_shape == (None, 25, 25, 128)
model.add(layers.BatchNormalization())
model.add(layers.ReLU())
model.add(layers.Conv2DTranspose(64, (5, 5), strides=(2, 2), padding='same', use_bias=False))# add kernel_initializer=init in case of weight initialization
assert model.output_shape == (None, 50, 50, 64)
model.add(layers.BatchNormalization())
model.add(layers.ReLU())
model.add(layers.Conv2DTranspose(3, (5, 5), strides=(2, 2), padding='same', use_bias=False, activation='sigmoid'))# add kernel_initializer=init in case of weight initialization
assert model.output_shape == (None, 100, 100, 3)
model.summary()
return model
# + id="kWUa_aPZ91qQ" outputId="5cd6899a-84eb-41b1-9e73-1df69fe3fd7d" colab={"base_uri": "https://localhost:8080/", "height": 795}
generator = make_generator_model()
noise = tf.random.normal([1, 100])
generated_image = generator(noise, training=True)
plt.imshow(generated_image[0, :, :, :], cmap='gray')
# + [markdown] id="1N-sZGs491qT"
# ### Critic
# + [markdown] id="658kwxq491qU"
# The model must take a sample image from our dataset as input and ~~output a classification prediction as to whether the sample is real or fake. This is a binary classification problem.~~ output scores for real and generated images.
#
# - Inputs: Image with three color channel and 100×100 pixels in size.
# - Outputs: ~~Binary classification, likelihood the sample is real (or fake).~~
# - Outputs: scores for real and generated images.
#
# 
# + id="3w8RsUgs91qU"
def make_critic_model():
model = tf.keras.Sequential()
model.add(layers.Conv2D(64, (5, 5), strides=(2, 2), padding='same',
input_shape=[100, 100, 3]))
model.add(layers.ReLU(0.2))
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(128, (5, 5), strides=(2, 2), padding='same'))
model.add(layers.ReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Flatten())
model.add(layers.Dense(1)) #activation='linear'
model.summary()
return model
# + id="BTYWrIv791qX" outputId="a8cd9369-fa28-4d14-fe44-cdb747629fe3"
critic = make_critic_model()
decision = critic(generated_image)
print (decision)
# + [markdown] id="VHt1Eaah91qa"
# ## Loss and Optimization
# + [markdown] id="rt9QH1tu91qb"
# ~~cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)~~
# + [markdown] id="-BipjOyN91qa"
# The Wasserstein GAN uses the 1-Wasserstein distance, rather than the JS-Divergence, to measure the difference between the model and target distributions.
# + id="2uLOrOwN91qb"
def critic_loss(r_logit, f_logit):
real_loss = - tf.reduce_mean(r_logit)
fake_loss = tf.reduce_mean(f_logit)
return real_loss, fake_loss
def generator_loss(f_logit):
fake_loss = - tf.reduce_mean(f_logit)
return fake_loss
# + id="k3wjWGcp91qf"
generator_optimizer = tf.keras.optimizers.Adam(learning_rate= LR) #tf.keras.optimizers.RMSprop(learning_rate= LR)
critic_optimizer = tf.keras.optimizers.Adam(learning_rate= LR) #tf.keras.optimizers.RMSprop(learning_rate= LR)
# + [markdown] id="Ri8gMQIM91qh"
# ## Experiment utils (RUN ME!)
# + id="4d7Rjrvu91qi"
def summary(name_data_dict,
step=None,
types=['mean', 'std', 'max', 'min', 'sparsity', 'histogram', 'image'],
historgram_buckets=None,
name='summary'):
"""Summary.
Examples
--------
>>> summary({'a': data_a, 'b': data_b})
"""
def _summary(name, data):
if data.shape == ():
tf.summary.scalar(name, data, step=step)
else:
if 'mean' in types:
tf.summary.scalar(name + '-mean', tf.math.reduce_mean(data), step=step)
if 'std' in types:
tf.summary.scalar(name + '-std', tf.math.reduce_std(data), step=step)
if 'max' in types:
tf.summary.scalar(name + '-max', tf.math.reduce_max(data), step=step)
if 'min' in types:
tf.summary.scalar(name + '-min', tf.math.reduce_min(data), step=step)
if 'sparsity' in types:
tf.summary.scalar(name + '-sparsity', tf.math.zero_fraction(data), step=step)
if 'histogram' in types:
tf.summary.histogram(name, data, step=step, buckets=historgram_buckets)
if 'image' in types:
tf.summary.image(name, data, step=step)
with tf.name_scope(name):
for name, data in name_data_dict.items():
_summary(name, data)
# + id="jeB-uZ_091ql"
train_summary_writer = tf.summary.create_file_writer(os.path.join(MODEL_SAVE_PATH, 'summaries', 'train'))
checkpoint_prefix = os.path.join(CHECKPOINT_DIR, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=critic_optimizer,
generator=generator,
discriminator=critic)
# + id="AwEDq0T991qn"
def generate_and_save_images(model, epoch, noise):
plt.figure(figsize=(15,10))
for i in range(4):
images = model(noise, training=False)
image = images[0, :, :, :]
image = np.reshape(image, [100, 100, 3])
plt.subplot(1, 4, i+1)
plt.imshow(np.uint8(image), cmap='gray')
plt.axis('off')
plt.title("Randomly Generated Images")
plt.tight_layout()
plt.savefig(os.path.join(MODEL_SAVE_PATH,'image_at_epoch_{:02d}.png'.format(epoch)))
plt.show()
# + [markdown] id="rSw2b9aC91qq"
# ## Training Process
# + id="S8CbAmbN91qq"
@tf.function
def train_generator(noise):
with tf.GradientTape() as tape:
# create fake image
generated_images = generator(noise, training=True)
fake_logit = critic(generated_images, training=True)
# calculate generator loss
g_loss = generator_loss(fake_logit)
gradients = tape.gradient(g_loss, generator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients, generator.trainable_variables))
return {'Generator loss': g_loss}
# + id="6wVBnVs491qt"
@tf.function
def train_Critic(noise, real_img):
with tf.GradientTape() as t:
fake_img = generator(noise, training=True)
real_logit = critic(real_img, training=True)
fake_logit = critic(fake_img, training=True)
real_loss, fake_loss = critic_loss(real_logit, fake_logit)
d_loss = (real_loss + fake_loss)
D_grad = t.gradient(d_loss, critic.trainable_variables)
critic_optimizer.apply_gradients(zip(D_grad, critic.trainable_variables))
for w in critic.trainable_variables:
w.assign(tf.clip_by_value(w, -CLIPPING_WEIGHT, CLIPPING_WEIGHT))
return {'Critic loss': real_loss + fake_loss}
# + id="HIllLMT591qv"
def train(dataset, epochs):
with train_summary_writer.as_default():
with tf.summary.record_if(True):
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
C_loss_dict = train_Critic(NOISE, image_batch)
summary(C_loss_dict, step=critic_optimizer.iterations, name='critic_losses')
if critic_optimizer.iterations.numpy() % N_CRITIC == 0:
G_loss_dict = train_generator(NOISE)
summary(G_loss_dict, step=generator_optimizer.iterations, name='generator_losses')
display.clear_output(wait=True)
generate_and_save_images(generator,
epoch + 1, NOISE)
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time for epoch {} is {} sec'.format(epoch + 1, time.time()-start))
display.clear_output(wait=True)
generate_and_save_images(generator, epochs, NOISE)
# + id="m0vRGsIl91qx"
train(dataset, EPOCHS)
# + id="V7WlzymG91q0"
# %tensorboard --logdir='/content/drive/My Drive/lecture hands on lab/wgan/results/summaries'
# + [markdown] id="OK1YYrAJ91q3"
# ### Generate Gif with the whole generated images during training
# + id="K8EXPgSY91q3"
anim_file = 'dcgan.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
import IPython
if IPython.version_info > (6,2,0,''):
display.Image(filename=anim_file)
# + [markdown] id="nk72PQzh91q6"
# ### Congratulations!
# you have built your first WGAN
# + [markdown] id="Ut7Q2B5891q6"
# ### What's expected from you
#
# * Play arround with optimizer
# * Tuning the model
# * Observe the critic and generator loss failure and image generation failure using tensorboard
# * Add L2 regularization
# + [markdown] id="iQxXsoDX91q7"
# ### Extra Reading :
# * https://paper.dropbox.com/doc/Wasserstein-GAN-GvU0p2V9ThzdwY3BbhoP7
# * https://arxiv.org/pdf/1803.00567.pdf
# * https://www.youtube.com/watch?v=SZHumKEhgtA
# * https://mindcodec.ai/2018/09/19/an-intuitive-guide-to-optimal-transport-part-i-formulating-the-problem/
# * https://vincentherrmann.github.io/blog/wasserstein/
# * https://mindcodec.ai/2018/09/23/an-intuitive-guide-to-optimal-transport-part-ii-the-wasserstein-gan-made-easy/
|
WGAN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solar Reflex Motion
#
# This notebook is aimed at investigating the calculation of solar reflex motion -- the vector correction for the motion of the sun around the Galaxy.
#
# Developed based on <NAME>'s Gala example [here](https://github.com/adrn/gala/blob/master/docs/coordinates/index.rst)
import astropy
import astropy.coordinates as coord
import astropy.units as u
print('astropy v.%s'%astropy.__version__)
# Our first task is to define a 6D phase space coordinate. This is defined in the heliocentric reference frame for both position (ra, dec, distance) and velocity (proper motion and radial velocity). We define a set of keyword arguments first, and then create the `SkyCoord` itself.
# +
# From Simon 2018 (https://arxiv.org/abs/1804.10230)
kwargs = dict(ra = 54 * u.deg,
dec = -54 * u.deg,
distance = 31e3 * u.pc,
pm_ra_cosdec=2.393*u.mas/u.yr,
pm_dec=-1.300*u.mas/u.yr,
radial_velocity=62.8*u.km/u.s
)
c = coord.SkyCoord(**kwargs)
# -
# Following the Price-Whelan example, we can now transform to a Galactocentric reference frame. Note that this is **both** the Galactic Standard of Rest (GSR) in velocity space **and** the Galactocentric spatial coordinate reference frame.
print(c.transform_to(coord.Galactocentric))
# In our case, we do not know the heliocentric radial velocity. To examine the impact of this uncertaintly, we loop through several values of the radial velocity and re-calculate the Galactocentric Cartesian velocities.
for v in [10, 30, 62.8, 90, 120]:
kwargs.update(radial_velocity = v * u.km/u.s)
c = coord.SkyCoord(**kwargs)
print("Heliocentric radial velocity: %.1f km/s"%v)
print("Galactocentric Cartesian velocity:")
print(c.transform_to(coord.Galactocentric).velocity)
print()
|
notebooks/SolarReflexMotion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] nbsphinx="hidden"
# This notebook is part of the $\omega radlib$ documentation: http://wradlib.org/wradlib-docs.
#
# Copyright (c) 2016, $\omega radlib$ developers.
# Distributed under the MIT License. See LICENSE.txt for more info.
# -
# # For Developers
# This section provides a collection of code snippets we use in $\omega radlib$ to achieve certain features.
#
# + [markdown] nbsphinx-toctree={"maxdepth": 2}
# ## Examples List
# - [Automate Image Generation](../tutorial_autoimages.rst)
# - [Apichange Function Decorators](develop/wradlib_api_change.ipynb)
|
notebooks/develop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# Google Trends gives us an estimate of search volume. Let's explore if search popularity relates to other kinds of data. Perhaps there are patterns in Google's search volume and the price of Bitcoin or a hot stock like Tesla. Perhaps search volume for the term "Unemployment Benefits" can tell us something about the actual unemployment rate?
#
# Data Sources: <br>
# <ul>
# <li> <a href="https://fred.stlouisfed.org/series/UNRATE/">Unemployment Rate from FRED</a></li>
# <li> <a href="https://trends.google.com/trends/explore">Google Trends</a> </li>
# <li> <a href="https://finance.yahoo.com/quote/TSLA/history?p=TSLA">Yahoo Finance for Tesla Stock Price</a> </li>
# <li> <a href="https://finance.yahoo.com/quote/BTC-USD/history?p=BTC-USD">Yahoo Finance for Bitcoin Stock Price</a> </li>
# </ul>
# # Import Statements
import pandas as pd
import matplotlib.pyplot as plt
# # Read the Data
#
# Download and add the .csv files to the same folder as your notebook.
df_tesla = pd.read_csv('TESLA Search Trend vs Price.csv')
df_btc_search = pd.read_csv('Bitcoin Search Trend.csv')
df_btc_price = pd.read_csv('Daily Bitcoin Price.csv')
df_unemployment = pd.read_csv('UE Benefits Search vs UE Rate 2004-19.csv')
# # Data Exploration
# ### Tesla
# **Challenge**: <br>
# <ul>
# <li>What are the shapes of the dataframes? </li>
# <li>How many rows and columns? </li>
# <li>What are the column names? </li>
# <li>Complete the f-string to show the largest/smallest number in the search data column</li>
# <li>Try the <code>.describe()</code> function to see some useful descriptive statistics</li>
# <li>What is the periodicity of the time series data (daily, weekly, monthly)? </li>
# <li>What does a value of 100 in the Google Trend search popularity actually mean?</li>
# </ul>
print(df_tesla.shape)
for col in df_tesla.columns:
print(col)
print(df_tesla.head(2))
print(f'Largest value for Tesla in Web Search: {df_tesla["TSLA_WEB_SEARCH"].min()}')
print(f'Smallest value for Tesla in Web Search: {df_tesla["TSLA_WEB_SEARCH"].max()}')
print(df_tesla.describe())
# ### Unemployment Data
print('Largest value for "Unemployemnt Benefits" '
f'in Web Search: ')
# ### Bitcoin
print(f'largest BTC News Search: ')
# # Data Cleaning
# ### Check for Missing Values
# **Challenge**: Are there any missing values in any of the dataframes? If so, which row/rows have missing values? How many missing values are there?
print(f'Missing values for Tesla?: {df_tesla.isna().values.any()}')
print(f'Missing values for U/E?: {df_unemployment.isna().values.any()}')
print(f'Missing values for BTC Search?: {df_btc_search.isna().values.any()}')
print(f'Missing values for BTC price?: {df_btc_price.isna().values.any()}')
print(f'Number of missing values: {df_btc_price.isna().values.sum()}')
# **Challenge**: Remove any missing values that you found.
df_btc_price.dropna(inplace=True)
# ### Convert Strings to DateTime Objects
# **Challenge**: Check the data type of the entries in the DataFrame MONTH or DATE columns. Convert any strings in to Datetime objects. Do this for all 4 DataFrames. Double check if your type conversion was successful.
df_tesla.MONTH = pd.to_datetime(df_tesla.MONTH)
df_btc_search.MONTH = pd.to_datetime(df_btc_search.MONTH)
df_unemployment.MONTH = pd.to_datetime(df_unemployment.MONTH)
df_btc_price.DATE = pd.to_datetime(df_btc_price.DATE)
df_tesla["MONTH"].head()
# ### Converting from Daily to Monthly Data
#
# [Pandas .resample() documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.resample.html) <br>
df_btc_monthly = df_btc_price.resample('M', on='DATE').last()
df_btc_monthly.head()
# # Data Visualisation
# ### Notebook Formatting & Style Helpers
# +
# Create locators for ticks on the time axis
# +
# Register date converters to avoid warning messages
# -
# ### Tesla Stock Price v.s. Search Volume
# **Challenge:** Plot the Tesla stock price against the Tesla search volume using a line chart and two different axes. Label one axis 'TSLA Stock Price' and the other 'Search Trend'.
# +
ax1 = plt.gca() # get current axis
ax2 = ax1.twinx()
ax1.set_ylabel('TSLA Stock Price')
ax2.set_ylabel('Search Trend')
ax1.plot(df_tesla.MONTH, df_tesla.TSLA_USD_CLOSE)
ax2.plot(df_tesla.MONTH, df_tesla.TSLA_WEB_SEARCH)
# -
# **Challenge**: Add colours to style the chart. This will help differentiate the two lines and the axis labels. Try using one of the blue [colour names](https://matplotlib.org/3.1.1/gallery/color/named_colors.html) for the search volume and a HEX code for a red colour for the stock price.
# <br>
# <br>
# Hint: you can colour both the [axis labels](https://matplotlib.org/3.3.2/api/text_api.html#matplotlib.text.Text) and the [lines](https://matplotlib.org/3.2.1/api/_as_gen/matplotlib.lines.Line2D.html#matplotlib.lines.Line2D) on the chart using keyword arguments (kwargs).
# +
ax1 = plt.gca()
ax2 = ax1.twinx()
ax1.set_ylabel('TSLA Stock Price', color='#DF3E1B') # can use a HEX code
ax2.set_ylabel('Search Trend') # or a named colour
ax1.plot(df_tesla.MONTH, df_tesla.TSLA_USD_CLOSE, color='#DF3E1B')
ax2.plot(df_tesla.MONTH, df_tesla.TSLA_WEB_SEARCH)
# -
# **Challenge**: Make the chart larger and easier to read.
# 1. Increase the figure size (e.g., to 14 by 8).
# 2. Increase the font sizes for the labels and the ticks on the x-axis to 14.
# 3. Rotate the text on the x-axis by 45 degrees.
# 4. Make the lines on the chart thicker.
# 5. Add a title that reads 'Tesla Web Search vs Price'
# 6. Keep the chart looking sharp by changing the dots-per-inch or [DPI value](https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.pyplot.figure.html).
# 7. Set minimum and maximum values for the y and x axis. Hint: check out methods like [set_xlim()](https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.axes.Axes.set_xlim.html).
# 8. Finally use [plt.show()](https://matplotlib.org/3.2.1/api/_as_gen/matplotlib.pyplot.show.html) to display the chart below the cell instead of relying on the automatic notebook output.
# +
plt.figure(figsize=(14,8), dpi=120)
plt.title('Tesla Web Search vs Price', fontsize=18)
ax1 = plt.gca()
ax2 = ax1.twinx()
# Also, increase fontsize and linewidth for larger charts
ax1.set_ylabel('TSLA Stock Price', color='#E6232E', fontsize=14)
ax2.set_ylabel('Search Trend', color='skyblue', fontsize=14)
ax1.plot(df_tesla.MONTH, df_tesla.TSLA_USD_CLOSE, color='#E6232E', linewidth=3)
ax2.plot(df_tesla.MONTH, df_tesla.TSLA_WEB_SEARCH, color='skyblue', linewidth=3)
# Displays chart explicitly
plt.show()
# -
# How to add tick formatting for dates on the x-axis.
# ### Bitcoin (BTC) Price v.s. Search Volume
# **Challenge**: Create the same chart for the Bitcoin Prices vs. Search volumes. <br>
# 1. Modify the chart title to read 'Bitcoin News Search vs Resampled Price' <br>
# 2. Change the y-axis label to 'BTC Price' <br>
# 3. Change the y- and x-axis limits to improve the appearance <br>
# 4. Investigate the [linestyles](https://matplotlib.org/3.2.1/api/_as_gen/matplotlib.pyplot.plot.html ) to make the BTC price a dashed line <br>
# 5. Investigate the [marker types](https://matplotlib.org/3.2.1/api/markers_api.html) to make the search datapoints little circles <br>
# 6. Were big increases in searches for Bitcoin accompanied by big increases in the price?
# ### Unemployement Benefits Search vs. Actual Unemployment in the U.S.
# **Challenge** Plot the search for "unemployment benefits" against the unemployment rate.
# 1. Change the title to: Monthly Search of "Unemployment Benefits" in the U.S. vs the U/E Rate <br>
# 2. Change the y-axis label to: FRED U/E Rate <br>
# 3. Change the axis limits <br>
# 4. Add a grey [grid](https://matplotlib.org/3.2.1/api/_as_gen/matplotlib.pyplot.grid.html) to the chart to better see the years and the U/E rate values. Use dashes for the line style<br>
# 5. Can you discern any seasonality in the searches? Is there a pattern?
# **Challenge**: Calculate the 3-month or 6-month rolling average for the web searches. Plot the 6-month rolling average search data against the actual unemployment. What do you see in the chart? Which line moves first?
#
# ### Including 2020 in Unemployment Charts
# **Challenge**: Read the data in the 'UE Benefits Search vs UE Rate 2004-20.csv' into a DataFrame. Convert the MONTH column to Pandas Datetime objects and then plot the chart. What do you see?
|
day074/Google Trends and Data Visualisation (start).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import fastf1 as ff1
from matplotlib import pyplot as plt
from fastf1 import plotting
plotting.setup_mpl()
# -
# # Variables
# +
directory = '../Telemetry/2021/TUR/Alpine/Race/'
year = 2021
raceNumber = 16
driver1_name = 'OCO'
driver1_fullname = 'Ocon'
driver2_name = 'ALO'
driver2_fullname = 'Alonso'
graphLinediWidth = 0.3
color1 = plotting.TEAM_COLORS['Alpine']
color2 = plotting.TEAM_COLORS['McLaren']
# -
# # Loads Session DATA
# +
ff1.Cache.enable_cache('cache')
race = ff1.get_session(year, raceNumber, 'R')
laps = race.load_laps()
lapsTelemetry = race.load_laps(with_telemetry=True)
# -
# # Initialization
# +
driver1 = lapsTelemetry.pick_driver(driver1_name).pick_fastest()
driver2 = lapsTelemetry.pick_driver(driver2_name).pick_fastest()
driver1LapsWithoutBox = laps.pick_driver(driver1_name).pick_wo_box()
driver2LapsWithoutBox = laps.pick_driver(driver2_name).pick_wo_box()
driver1Laps = laps.pick_driver(driver1_name)
driver2Laps = laps.pick_driver(driver2_name)
driver1Data = driver1.telemetry
driver2Data = driver2.telemetry
name1 = driver1_fullname
name2 = driver2_fullname
# -
# # Telemetry charts
# ## Fastest Lap
# +
fig, ax = plt.subplots()
ax.plot(driver1Data['Distance'], driver1Data['Speed'], color=color1, linewidth=graphLinediWidth, label=name1)
ax.plot(driver2Data['Distance'], driver2Data['Speed'], color=color2, linewidth=graphLinediWidth, label=name2)
ax.set_title("Fastest race lap")
ax.set_xlabel("Distance (m)")
ax.set_ylabel("Speed (kph)")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.text(-0.05, 1.13, driver1['LapTime'].total_seconds(),
horizontalalignment='center',
verticalalignment='center',
transform = ax.transAxes)
plt.text(-0.05, 1.06,driver2['LapTime'].total_seconds(),
horizontalalignment='center',
verticalalignment='center',
transform = ax.transAxes)
plt.savefig(directory + 'telemetrySpeed.png', dpi=1200)
plt.show()
# +
fig, ax = plt.subplots()
ax.plot(driver1Data['Distance'], driver1Data['RPM'], color=color1, linewidth=graphLinediWidth, label=name1)
ax.plot(driver2Data['Distance'], driver2Data['RPM'], color=color2, linewidth=graphLinediWidth, label=name2)
ax.set_title("Fastest race lap")
ax.set_xlabel("Distance (m)")
ax.set_ylabel("RPM")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory +'telemetryRPM.png', dpi=1200)
plt.show()
# +
fig, ax = plt.subplots()
ax.plot(driver1Data['Distance'], driver1Data['nGear'], color=color1, linewidth=graphLinediWidth, label=name1)
ax.plot(driver2Data['Distance'], driver2Data['nGear'], color=color2, linewidth=graphLinediWidth, label=name2)
ax.set_title("Fastest race lap")
ax.set_xlabel("Distance (m)")
ax.set_ylabel("Gear)")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory + 'telemetryGear.png', dpi=1200)
plt.show()
# +
fig, ax = plt.subplots()
ax.plot(driver1Data['Distance'], driver1Data['Throttle'], color=color1, linewidth=graphLinediWidth, label=name1)
ax.plot(driver2Data['Distance'], driver2Data['Throttle'], color=color2, linewidth=graphLinediWidth, label=name2)
ax.set_title("Fastest race lap")
ax.set_xlabel("Distance (m)")
ax.set_ylabel("Throttle pedal pressure 0-100")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory + 'telemetryThrottle2.png', dpi=1200)
plt.show()
# +
fig, ax = plt.subplots()
ax.plot(driver1Data['Distance'], driver1Data['Brake'], color=color1, linewidth=graphLinediWidth, label=name1)
ax.plot(driver2Data['Distance'], driver2Data['Brake'], color=color2, linewidth=graphLinediWidth, label=name2)
ax.set_title("Fastest race lap")
ax.set_xlabel("Distance (m)")
ax.set_ylabel("Brake pedal pressure 0-100")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory + 'telemetryBrake.png', dpi=1200)
plt.show()
# +
fig, ax = plt.subplots()
ax.plot(driver1Data['Distance'], driver1Data['DRS'], color=color1, linewidth=graphLinediWidth, label=name1)
ax.plot(driver2Data['Distance'], driver2Data['DRS'], color=color2, linewidth=graphLinediWidth, label=name2)
ax.set_title("Fastest race lap")
ax.set_xlabel("Distance (m)")
ax.set_ylabel("DRS indicator")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory + 'telemetryDRS.png', dpi=1200)
plt.show()
# -
# ## Race pace
# +
plotting.setup_mpl()
fig, ax = plt.subplots()
plt.ylim(top=100, bottom=93)
ax.plot(driver1LapsWithoutBox['LapNumber'], driver1LapsWithoutBox['LapTime'].dt.total_seconds(), color=color1, linewidth=graphLinediWidth, label=name1)
ax.plot(driver2LapsWithoutBox['LapNumber'], driver2LapsWithoutBox['LapTime'].dt.total_seconds(), color=color2, linewidth=graphLinediWidth, label=name2)
ax.set_title("Race pace")
ax.set_xlabel("Lap Number")
ax.set_ylabel("Lap Time")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory + 'lapsTrimmed.png', dpi=1200)
plt.show()
# +
import numpy as np
plotting.setup_mpl()
fig, ax = plt.subplots()
plt.ylim(top=2, bottom=-2)
o = driver1Laps['LapTime'].dt.total_seconds().to_numpy()
a = driver2Laps['LapTime'].dt.total_seconds().to_numpy()
#o = np.delete(o, -1, 0) # remove last lap for the non-lapped drivers
diff = np.subtract(o, a)
ax.plot(driver1Laps['LapNumber'], diff, color='green', linewidth=1)
ax.set_title("Race pace")
ax.set_xlabel("Lap Number")
ax.set_ylabel("Delta (Ocon to Alonso)")
#ax.legend(bbox_to_anchor=(0,1),loc=3)
ma = np.ma.MaskedArray(diff, mask=np.isnan(diff))
avg = "average " + str(round(np.ma.average(ma), 3))
med = "median " + str(round(np.ma.median(ma), 3))
plt.text(-0.05, 1.13,avg,
horizontalalignment='center',
verticalalignment='center',
transform = ax.transAxes)
plt.text(-0.05, 1.06,med,
horizontalalignment='center',
verticalalignment='center',
transform = ax.transAxes)
plt.savefig(directory + 'diff.png', dpi=1200)
plt.show()
# -
# ## Experimental
# ### Race pace with tyre compounds
# +
plotting.setup_mpl()
fig, ax = plt.subplots()
driver1LapsHard = driver1Laps.pick_tyre('HARD')
driver2LapsHard = driver2Laps.pick_tyre('HARD')
driver1LapsMedium = driver1Laps.pick_tyre('MEDIUM')
driver2LapsMedium = driver2Laps.pick_tyre('MEDIUM')
colorHard = 'white'
colorMedium = 'yellow'
ax.plot(driver1LapsHard['LapNumber'], driver1LapsHard['LapTime'].dt.total_seconds(), color=colorHard, linewidth=graphLinediWidth, label=name1, linestyle='-')
ax.plot(driver2LapsHard['LapNumber'], driver2LapsHard['LapTime'].dt.total_seconds(), color=colorHard, linewidth=graphLinediWidth, label=name2, linestyle='--')
ax.plot(driver1LapsMedium['LapNumber'], driver1LapsMedium['LapTime'].dt.total_seconds(), color=colorMedium, linewidth=graphLinediWidth, label=name1, linestyle='-')
ax.plot(driver2LapsMedium['LapNumber'], driver2LapsMedium['LapTime'].dt.total_seconds(), color=colorMedium, linewidth=graphLinediWidth, label=name2, linestyle='--')
ax.set_title("Race pace")
ax.set_xlabel("Lap Number")
ax.set_ylabel("Lap Time")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory + 'laps.png', dpi=1200)
plt.show()
# +
plotting.setup_mpl()
fig, ax = plt.subplots()
plt.ylim(top=100, bottom=90)
driver1LapsHard = driver1Laps.pick_tyre('HARD')
driver2LapsHard = driver2Laps.pick_tyre('HARD')
driver1LapsMedium = driver1Laps.pick_tyre('MEDIUM')
driver2LapsMedium = driver2Laps.pick_tyre('MEDIUM')
ax.plot(driver1LapsHard['LapNumber'], driver1LapsHard['LapTime'].dt.total_seconds(), color=colorHard, linewidth=graphLinediWidth, label=name1, linestyle='-')
ax.plot(driver2LapsHard['LapNumber'], driver2LapsHard['LapTime'].dt.total_seconds(), color=colorHard, linewidth=graphLinediWidth, label=name2, linestyle='--')
ax.plot(driver1LapsMedium['LapNumber'], driver1LapsMedium['LapTime'].dt.total_seconds(), color=colorMedium, linewidth=graphLinediWidth, label=name1, linestyle='-')
ax.plot(driver2LapsMedium['LapNumber'], driver2LapsMedium['LapTime'].dt.total_seconds(), color=colorMedium, linewidth=graphLinediWidth, label=name2, linestyle='--')
ax.set_title("Race pace")
ax.set_xlabel("Lap Number")
ax.set_ylabel("Lap Time")
ax.legend(bbox_to_anchor=(0,1),loc=3)
plt.savefig(directory + 'lapsTrimmedWithCompounds.png', dpi=1200)
plt.show()
|
Notebooks/.ipynb_checkpoints/Analysis Race-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 객담도말 결핵진단 딥러닝 모델
#
# CNN 기반의 객담도말 결핵진단 딥러닝 모델 소스코드입니다.
# +
import os
from __future__ import print_function
import numpy as np
np.random.seed(1337) # for reproducibility
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution2D, MaxPooling2D
from keras.utils import np_utils
# -
# 딥러닝 모델관련 환경설정입니다.
BATCH_SIZE = 128 # 한 epoch에서 실행시키는 단위(배치)크기
NUM_CLASSES = 2 # 클래스 수
NUM_EPOCHS = 1 # epoch 수
NUM_FILTERS = 32 # convolution 필터 수
NUM_POOL = 2 # max plling을 위한 pooling 영역 크기
NUM_CONV = 3 # convolution 커널 크기
# 데이터셋 관련 환경설정입니다.
# +
IMG_CHANNELS = 1
IMG_ROWS = 64
IMG_COLS = 64
TRAIN_DATA_COUNT = 447648
train_img_filename = './datasets/train_image_64x64_gray_447648.bin'
train_label_filename = './datasets/train_label_64x64_gray_447648.bin'
TEST_DATA_COUNT = 15873
test_img_filename = './datasets/test_image_64x64_gray_15873.bin'
test_label_filename = './datasets/test_label_64x64_gray_15873.bin'
VALIDATION_DATA_COUNT = int(TRAIN_DATA_COUNT * 1.0/4.0)
# -
MODEL_SAVE_FILE_PATH = './seq_model_cnn.h5'
PREDICT_FILE_PATH = './predict.txt'
# img 자료 로딩 함수 입니다.
def load_img(filename, count, channel, row, col):
print('Loading data from', filename)
print('file size : ', os.path.getsize(filename))
print('calc size : ', count * channel * row * col)
fp = open(filename, 'rb')
buf = fp.read(count * channel * row * col)
data = np.frombuffer(buf, dtype=np.uint8)
data = data.reshape(count, channel, row, col)
print('loaded shape : ', data.shape)
data = data.astype('float32')
data /= 255
return data
# label 자료 로딩함수입니다.
def load_label(filename, count, classes):
print('Loading labels from ', filename)
print('file size : ', os.path.getsize(filename))
print('calc size : ', count)
fp = open(filename, 'r')
buf = fp.read(count)
data_bin = []
for i in buf:
data_bin.append(i)
data = np.asarray(data_bin, dtype=np.uint8, order='C')
print('loaded shape : ', data.shape)
label_hist = np.histogram(data, bins=range(NUM_CLASSES+1))
print(label_hist)
# convert class vectors to binary class matrices
data = np_utils.to_categorical(data, classes)
return data
# +
# the data, shuffled and split between train and test sets
train_img = load_img(train_img_filename, TRAIN_DATA_COUNT, IMG_CHANNELS, IMG_ROWS, IMG_COLS)
test_img = load_img(test_img_filename, TEST_DATA_COUNT, IMG_CHANNELS, IMG_ROWS, IMG_COLS)
#validation_img = load_img(validation_img_filename, VALIDATION_DATA_COUNT, IMG_CHANNELS, IMG_ROWS, IMG_COLS)
train_label = load_label(train_label_filename, TRAIN_DATA_COUNT, NUM_CLASSES)
test_label = load_label(test_label_filename, TEST_DATA_COUNT, NUM_CLASSES)
#validation_label = load_label(validation_label_filename, VALIDATION_DATA_COUNT, NUM_CLASSES)
# -
# 훈련셋의 일부로부터 검증셋을 생성합니다.
# +
validation_img = train_img[:VALIDATION_DATA_COUNT, ...]
validation_label = train_label[:VALIDATION_DATA_COUNT, ...]
train_img = train_img[VALIDATION_DATA_COUNT:, ...]
train_label = train_label[VALIDATION_DATA_COUNT:, ...]
print('train count : ' + str(len(train_img)))
print('validation count : ' + str(len(validation_img)))
# -
# 딥리닝 모델을 구축합니다.
# +
model = Sequential()
model.add(Convolution2D(NUM_FILTERS, NUM_CONV, NUM_CONV,
border_mode='valid',
input_shape=(IMG_CHANNELS, IMG_ROWS, IMG_COLS)))
model.add(Activation('relu'))
model.add(Convolution2D(NUM_FILTERS, NUM_CONV, NUM_CONV))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(NUM_POOL, NUM_POOL)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(NUM_CLASSES))
model.add(Activation('softmax'))
# np_utils.visualize_util.plot(model, to_file='model.png')
# -
# 딥러닝 모델을 구축합니다.
# +
model.compile(loss='categorical_crossentropy',
optimizer='adadelta',
metrics=['accuracy'])
model.fit(train_img,
train_label,
batch_size=BATCH_SIZE,
nb_epoch=NUM_EPOCHS,
verbose=1,
validation_data=(validation_img, validation_label))
# -
# 딥러닝 모델 테스트를 수행합니다.
score = model.evaluate(test_img, test_label, verbose=1)
print('Test score:', score[0])
print('Test accuracy:', score[1])
classes = model.predict_classes(test_img, batch_size=32)
np.savetxt(PREDICT_FILE_PATH, classes, fmt='%d')
model.summary()
model.save_weights(MODEL_SAVE_FILE_PATH)
|
_writing/tb/tb_seq_cnn.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// In this GettingStarted article, we will build a robot for answering questions in IQ test with the help of [DeepLearning.scala](http://deeplearning.thoughtworks.school/).
// ## Background
// Suppose we are building a robot for answering questions in IQ test like this:
//
// > What is the next number in sequence:
// >> 3, 6, 9, ?
// >
// > The answer is 12.
// We prepared some questions and corresponding answers as [INDArray](https://oss.sonatype.org/service/local/repositories/public/archive/org/nd4j/nd4j-api/0.8.0/nd4j-api-0.8.0-javadoc.jar/!/org/nd4j/linalg/api/ndarray/INDArray.html)s:
// +
import $ivy.`org.nd4j::nd4s:0.8.0`
import $ivy.`org.nd4j:nd4j-native-platform:0.8.0`
import org.nd4j.linalg.api.ndarray.INDArray
val TrainingQuestions: INDArray = {
import org.nd4s.Implicits._
Array(
Array(0, 1, 2),
Array(4, 7, 10),
Array(13, 15, 17)
).toNDArray
}
val ExpectedAnswers: INDArray = {
import org.nd4s.Implicits._
Array(
Array(3),
Array(13),
Array(19)
).toNDArray
}
// -
// These samples will be used to train the robot.
// In the rest of this article, we will build the robot in the following steps:
//
// 1. Install DeepLearning.scala, which is the framework that helps us build the robot.
// 1. Setup configuration (also known as hyperparameters) of the robot.
// 1. Build an untrained neural network of the robot.
// 1. Train the neural network using the above samples.
// 1. Test the robot seeing if the robot have been learnt how to answer these kind of questions.
// ## Install DeepLearning.scala
// DeepLearning.scala is hosted on Maven Central repository.
//
// You can use magic imports in [jupyter-scala](https://github.com/alexarchambault/jupyter-scala) or [Ammonite-REPL](http://www.lihaoyi.com/Ammonite/#Ammonite-REPL) to download DeepLearning.scala and its dependencies.
import $ivy.`com.thoughtworks.deeplearning::plugins-builtins:2.0.0-RC1`
// If you use [sbt](http://www.scala-sbt.org), please add the following settings into your `build.sbt`:
//
// ``` scala
// libraryDependencies += "com.thoughtworks.deeplearning" %% "plugins-builtins" % "latest.release"
//
// libraryDependencies += "org.nd4j" %% "nd4j-native-platform" % "0.8.0"
//
// fork := true
//
// scalaVersion := "2.11.11"
// ```
//
// Note that this example must run on Scala 2.11.11 because [nd4s](http://nd4j.org/scala) does not support Scala 2.12. Make sure there is not a setting like `scalaVersion := "2.12.x"` in your `build.sbt`.
//
// See [Scaladex](https://index.scala-lang.org/thoughtworksinc/deeplearning.scala) to install DeepLearning.scala in other build tools!
// ## Setup hyperparameters
// Hyperparameters are global configurations for a neural network.
//
// For this robot, we want to set its learning rate, which determines how fast the robot change its inner weights.
//
// In DeepLearning.scala, hyperparameters can be introduced by plugins, which is a small piece of code loaded from a URL.
val INDArrayLearningRatePluginUrl = "https://gist.githubusercontent.com/Atry/1fb0608c655e3233e68b27ba99515f16/raw/27c7d00dd37785335b6acfe1f1c5614843bc6d9f/INDArrayLearningRate.sc"
interp.load(scala.io.Source.fromURL(new java.net.URL(INDArrayLearningRatePluginUrl)).mkString)
// By loading the hyperparameter plugin `INDArrayLearningRate`, we are able to create the context of neural network with `learningRate` parameter.
import com.thoughtworks.deeplearning.plugins.Builtins
// All DeepLearning.scala built-in features are also provided by plugins. [Builtins](https://javadoc.io/page/com.thoughtworks.deeplearning/plugins-builtins_2.11/latest/com/thoughtworks/deeplearning/plugins/Builtins.html) is the plugin that contains all other DeepLearning.scala built-in plugins.
// Now we create the context and setup learning rate to `0.001`.
// `interp.load` is a workaround for https://github.com/lihaoyi/Ammonite/issues/649 and https://github.com/scala/bug/issues/10390
interp.load("""
import scala.concurrent.ExecutionContext.Implicits.global
import com.thoughtworks.feature.Factory
val hyperparameters = Factory[Builtins with INDArrayLearningRate].newInstance(learningRate = 0.001)
""")
// See [Factory](https://javadoc.io/page/com.thoughtworks.feature/factory_2.11/latest/com/thoughtworks/feature/Factory.html) if you are wondering how those plugins are composed together.
// The `Builtins` plugin contains some implicit values and views, which should be imported as following:
import hyperparameters.implicits._
// ## Build an untrained neural network of the robot
// In DeepLearning.scala, a neural network is simply a function that references some **weights**, which are mutable variables being changed automatically according to some goals during training.
//
// For example, given `x0`, `x1` and `x2` are the input sequence passed to the robot, we can build a function that returns the answer as `robotWeight0 * x0 + robotWeight1 * x1 + robotWeight2 * x2`, by adjusting those weights during training, the result should become close to the expected answer.
// In DeepLearning.scala, weights can be created as following:
// +
def initialValueOfRobotWeight: INDArray = {
import org.nd4j.linalg.factory.Nd4j
import org.nd4s.Implicits._
Nd4j.randn(3, 1)
}
import hyperparameters.INDArrayWeight
val robotWeight = INDArrayWeight(initialValueOfRobotWeight)
// -
// In the above code, `robotWeight` is a weight of n-dimensional array, say, [INDArrayWeight], initialized from random values. Therefore, the formula `robotWeight0 * x0 + robotWeight1 * x1 + robotWeight2 * x2` can be equivalent to a matrix multipication, written as a `dot` method call:
//
// [INDArrayWeight]: https://javadoc.io/page/com.thoughtworks.deeplearning/plugins-builtins_2.11/latest/com/thoughtworks/deeplearning/plugins/INDArrayWeights$INDArrayWeight.html
import hyperparameters.INDArrayLayer
def iqTestRobot(questions: INDArray): INDArrayLayer = {
questions dot robotWeight
}
// Note that the `dot` method is a differentiable function provided by DeepLearning.scala.
// You can find other [n-dimensional array differentiable methods in Scaladoc](https://javadoc.io/page/com.thoughtworks.deeplearning/plugins-builtins_2.11/latest/com/thoughtworks/deeplearning/plugins/RawINDArrayLayers$ImplicitsApi$INDArrayLayerOps.html)
//
// Unlike the functions in nd4s, all those differentiable functions accepts either an `INDArray`, `INDArrayWeight`
// or [INDArrayLayer], and returns one [Layer] of neural network, which can be composed into another differentiable function call.
//
// [INDArrayLayer]: https://javadoc.io/page/com.thoughtworks.deeplearning/plugins-builtins_2.11/latest/com/thoughtworks/deeplearning/plugins/RawINDArrayLayers$INDArrayLayer.html
//
// [Layer]: https://javadoc.io/page/com.thoughtworks.deeplearning/plugins-builtins_2.11/latest/com/thoughtworks/deeplearning/plugins/Layers$Layer.html
// ## Training the network
// ### Loss function
// In DeepLearning.scala, when we train a neural network, our goal should always be minimizing the return value.
//
// For example, if `iqTestRobot(TrainingQuestions).train` get called repeatedly,
// the neural network would try to minimize `input dot robotWeight`.
// `robotWeight` would become smaller and smaller in order to make `input dot robotWeight` smaller,
// and `iqTestRobot(TrainingQuestions).predict` would return an `INDArray` of small numbers.
//
// What if you expect `iqTestRobot(TrainingQuestions).predict` to return `ExpectedAnswers`?
//
// You can create another neural network that evaluates how far between the result of `myNeuralNetwork` and your expectation. The new neural network is usually called **loss function**.
//
// In this article we will use square loss as the loss function:
import hyperparameters.DoubleLayer
def squareLoss(questions: INDArray, expectAnswer: INDArray): DoubleLayer = {
val difference = iqTestRobot(questions) - expectAnswer
(difference * difference).mean
}
// When the `lossFunction` get trained continuously, its return value will be close to zero, and the result of `myNeuralNetwork` must be close to the expected result at the same time.
//
// Note the `lossFunction` accepts a `questions` and `expectAnswer` as its parameter.
// The first parameter is the input data used to train the neural network, and the second array is the expected output.
//
// The `squareLoss` function itself is a neural network, internally using the layer returned by `iqTestRobot` method.
// ### Run the training task
// As I mentioned before, there is a [train] method for `DoubleLayer`. It is a [Task] that performs one iteration of training.
//
// Since we want to repeatedly train the neural network of the robot, we need to create another `Task` that performs many iterations of training.
//
// In this article, we use [ThoughtWorks Each] to build such a `Task`:
//
// [train]: https://javadoc.io/page/com.thoughtworks.deeplearning/plugins-builtins_2.11/latest/com/thoughtworks/deeplearning/DeepLearning$$Ops.html#train(implicitmonoid:spire.algebra.MultiplicativeMonoid[Ops.this.typeClassInstance.Delta]):scalaz.concurrent.Task[Ops.this.typeClassInstance.Data]
//
// [Task]: https://javadoc.io/page/org.scalaz/scalaz-concurrent_2.11/latest/scalaz/concurrent/Task.html
//
// [ThoughtWorks Each]: https://github.com/ThoughtWorksInc/each
// +
import $ivy.`com.thoughtworks.each::each:3.3.1`
import $plugin.$ivy.`org.scalamacros:paradise_2.11.11:2.1.0`
import com.thoughtworks.each.Monadic._
import scalaz.concurrent.Task
import scalaz.std.stream._
// +
val TotalIterations = 500
@monadic[Task]
def train: Task[Stream[Double]] = {
for (iteration <- (0 until TotalIterations).toStream) yield {
squareLoss(TrainingQuestions, ExpectedAnswers).train.each
}
}
// -
// Then we can run the task to train the robot.
val lossByTime: Stream[Double] = train.unsafePerformSync
// Then we create a plot to show how the loss changed during iterations.
// +
import $ivy.`org.plotly-scala::plotly-jupyter-scala:0.3.2`
import plotly._
import plotly.element._
import plotly.layout._
import plotly.JupyterScala._
plotly.JupyterScala.init()
// -
Scatter(lossByTime.indices, lossByTime).plot(title = "loss by time")
// After these iterations, the loss should be close to zero.
// ## Test the trained robot
val TestQuestions: INDArray = {
import org.nd4s.Implicits._
Array(Array(3, 6, 9)).toNDArray
}
iqTestRobot(TestQuestions).predict.unsafePerformSync
// The result should be close to `12`.
// You may also see the value of weights in the trained neural network:
val weightData: INDArray = robotWeight.data
// ## Conclusion
// In this article, we have created a IQ test robot with the help of DeepLearning.scala.
//
// The model of robot is linear regression with a square loss, which consists of some `INDArryWeight`s and `INDArrayLayer`s.
//
// After many iterations of `train`ing, the robot finally learnt the pattern of arithmetic progression.
|
demo/2.0.0-Preview/GettingStarted.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.colors import LogNorm
import copy
import random
import time
def initialise_state(N): #N is the grid dimension (in the above example, N=4)
'''
Author: <NAME>
~Function Description~
'''
grid = np.ones((N,N,2),dtype=int)
return np.array(grid)
# +
def plot_vector(p1,p2):
'''
Author: <NAME>
~Function Description~
'''
p1 = np.array(p1)
p2 = np.array(p2)
dp = p2-p1
plt.quiver(p1[0], p1[1], dp[0], dp[1],angles='xy', scale_units='xy', scale=1, headwidth = 5, headlength = 7)
def get_coord_list(arr):
'''
Author: <NAME>
~Function Description~
'''
coord_list=[]
num = len(arr)
for i in range(num):
temp_coord = []
for j in range(num):
current_elems = arr[i][j]
xpt = (num-1)-i
ypt = j
temp_coord.append((xpt,ypt))
coord_list.append(temp_coord)
return coord_list
def visualise_2d_model(arr,savefig=False,savename=".temp"):
'''
Author: <NAME>
~Function Description~
'''
num = len(arr)
plt.axes().set_aspect('equal')
coord_list = get_coord_list(arr)
for i in range(num):
for j in range(num):
current_up_state = arr[i][j][0]
current_right_state = arr[i][j][1]
x_current = coord_list[i][j][1]
y_current = coord_list[i][j][0]
lower_neighbour_up_state = arr[(i+1)%num][j][0]
x_up = coord_list[(i+1)%num][j][1]
y_up = coord_list[(i+1)%num][j][0]
left_neighbour_right_state = arr[i][j-1][1]
x_left = coord_list[i][j-1][1]
y_left = coord_list[i][j-1][0]
current_down_state = -(lower_neighbour_up_state)
current_left_state = -(left_neighbour_right_state)
# plt.plot(x_current,y_current,'ob')
plt.plot(x_current,y_current,
marker="o", markersize=9, markeredgecolor="k",
markerfacecolor="red",
zorder=1)
if current_up_state == 1:
plot_vector([x_current,y_current],[x_current,y_current+1])
elif current_up_state == -1:
plot_vector([x_current,y_current+1],[x_current,y_current])
if current_right_state == 1:
plot_vector([x_current,y_current],[x_current+1,y_current])
elif current_right_state == -1:
plot_vector([x_current+1,y_current],[x_current,y_current])
if current_down_state == 1:
plot_vector([x_current,y_current],[x_current,y_current-1])
elif current_down_state == -1:
plot_vector([x_current,y_current-1],[x_current,y_current])
if current_left_state == 1:
plot_vector([x_current,y_current],[x_current-1,y_current])
elif current_left_state == -1:
plot_vector([x_current-1,y_current],[x_current,y_current])
plt.xlim(-1,num+1)
plt.ylim(-1,num+1)
plt.axis('off')
if savefig:
plt.savefig(f"{savename}.png",dpi=300)
plt.show()
plt.close()
# -
def check_config(arr):
'''
Author: <NAME>
~Function Description~
'''
flag=True
N=len(arr)
for i in range(len(arr)):
for j in range(len(arr)):
current_up_state = arr[i][j][0]
current_right_state = arr[i][j][1]
lower_neighbour_up_state = arr[(i+1)%N][j][0]
left_neighbour_right_state = arr[i][j-1][1]
current_left_state = -(left_neighbour_right_state)
current_down_state = -(lower_neighbour_up_state)
if (current_up_state + current_right_state + current_left_state + current_down_state) != 0:
flag=False
break
return flag
# # LONG LOOP
def long_loop(arr2, verbose=False):
'''
Author: Team ℏ
~Function Description~
'''
arr = copy.deepcopy(arr2)
N=len(arr)
iters=0
n1 = np.random.randint(low=0, high=N)
n2 = np.random.randint(low=0, high=N)
inital_pt =(n1,n2)
prev_choice=None
while True:
iters+=1
if n1==inital_pt[0] and n2==inital_pt[1] and iters!=1:
if verbose:
print(f"Completed in {iters} iterations.")
# assert(check_config(arr))
break
current_up_state = arr[n1][n2][0]
current_right_state = arr[n1][n2][1]
lower_neighbour_up_state = arr[(n1+1)%N][n2][0]
left_neighbour_right_state = arr[n1][n2-1][1]
current_down_state = -(lower_neighbour_up_state)
current_left_state = -(left_neighbour_right_state)
current_states_dict = {"up":current_up_state,"right":current_right_state,"down":current_down_state,"left":current_left_state}
outgoing_state_dict={}
incoming_state_dict={}
for key in current_states_dict.keys():
if current_states_dict[key]==1: #current state is outgoing
outgoing_state_dict[key]=current_states_dict[key]
else:
incoming_state_dict[key]=current_states_dict[key]
if prev_choice =="right":
forbidden_choice="left"
if prev_choice =="up":
forbidden_choice="down"
if prev_choice =="left":
forbidden_choice="right"
if prev_choice =="down":
forbidden_choice="up"
else:
forbidden_choice=None
while True:
out_choice = np.random.choice(list(outgoing_state_dict.keys()))
if out_choice !=forbidden_choice:
break
prev_choice=out_choice
if out_choice == "up":
arr[n1][n2][0]= - (arr[n1][n2][0])
n1=(n1-1)%N
n2=n2
continue
if out_choice == "right":
arr[n1][n2][1]= - (arr[n1][n2][1])
n1=n1
n2=(n2+1)%N
continue
if out_choice == "down":
arr[(n1+1)%N][n2][0]= - (arr[(n1+1)%N][n2][0])
n1=(n1+1)%N
n2=n2
continue
if out_choice == "left":
arr[n1][(n2-1)%N][1]= - (arr[n1][(n2-1)%N][1])
n1=n1
n2=(n2-1)%N
continue
return arr
def count_states(num,error_threshold,return_dict = False,verbose=False):
'''
Author: Team ℏ
~Function Description~
'''
if not (error_threshold<=100 and error_threshold>0):
print("Error! Please input error_threshold as a value between 0 and 100")
assert (error_threshold<=100 and error_threshold>0)
state_dict={}
oldarr = long_loop(initialise_state(num), verbose=False)
good_iterations = 0 #Iterations that gave us a new state, so good.
bad_iterations = 0 #Iterations that gave us an already found state,so a waste and hence bad.
while True:
newarr = long_loop(oldarr,verbose=False)
name =arr_to_string(newarr)
if name not in state_dict:
count_repetitions=0
state_dict[name]=1
good_iterations+=1
else:
bad_iterations+=1
count_repetitions+=1
state_dict[name]+=1
percent_approx_err=good_iterations*100/(good_iterations+bad_iterations)
if verbose:
print(f"Good iterations = {good_iterations} and bad iterations = {bad_iterations} and Error % = {percent_approx_err}", end="\r",flush=True)
if percent_approx_err < error_threshold:
break
oldarr=newarr
if return_dict:
return len(state_dict),state_dict
else:
return len(state_dict)
def state2to4(arr):
'''
Author: <NAME>
Examine once.
~Function Description~
'''
fourstatearr=np.zeros((arr.shape[0],arr.shape[1],4))
N=len(arr)
for i in range(len(arr)):
for j in range(len(arr)):
current_up_state = arr[i][j][0]
current_right_state = arr[i][j][1]
lower_neighbour_up_state = arr[(i+1)%N][j][0]
left_neighbour_right_state = arr[i][j-1][1]
current_left_state = -(left_neighbour_right_state)
current_down_state = -(lower_neighbour_up_state)
fourstatearr[i][j][0] = current_up_state
fourstatearr[i][j][1] = current_right_state
fourstatearr[i][j][2] = current_down_state
fourstatearr[i][j][3] = current_left_state
return fourstatearr
# +
#Rot 90 anticlock
#Up becomes left, left becomes down, down becomes right, right becomes up
def rot90_anticlock(arr2):
'''
Author: <NAME>
~Function Description~
'''
fourstatearr = state2to4(arr2)
fourstatearr = np.rot90(fourstatearr,1)
arr=np.zeros((fourstatearr.shape[0],fourstatearr.shape[1],2))
N=len(arr)
for i in range(len(arr)):
for j in range(len(arr)):
current_up_state = fourstatearr[i][j][0]
current_right_state = fourstatearr[i][j][1]
current_down_state = fourstatearr[i][j][2]
current_left_state = fourstatearr[i][j][3]
new_up_state = current_right_state
new_right_state = current_down_state
arr[i][j][0]=new_up_state
arr[i][j][1]=new_right_state
return arr.astype(int)
#Rot 180 anticlock
#Up becomes down, left becomes right, down becomes up, right becomes left
def rot180_anticlock(arr2):
'''
Author: <NAME>
~Function Description~
'''
fourstatearr = state2to4(arr2)
fourstatearr = np.rot90(fourstatearr,2)
arr=np.zeros((fourstatearr.shape[0],fourstatearr.shape[1],2))
N=len(arr)
for i in range(len(arr)):
for j in range(len(arr)):
current_up_state = fourstatearr[i][j][0]
current_right_state = fourstatearr[i][j][1]
current_down_state = fourstatearr[i][j][2]
current_left_state = fourstatearr[i][j][3]
new_up_state = current_down_state
new_right_state = current_left_state
arr[i][j][0]=new_up_state
arr[i][j][1]=new_right_state
return arr.astype(int)
#Rot 270 anticlock
#Up becomes right, left becomes up, down becomes left, right becomes down
def rot270_anticlock(arr2):
'''
Author: <NAME>
~Function Description~
'''
fourstatearr = state2to4(arr2)
fourstatearr = np.rot90(fourstatearr,3)
arr=np.zeros((fourstatearr.shape[0],fourstatearr.shape[1],2))
N=len(arr)
for i in range(len(arr)):
for j in range(len(arr)):
current_up_state = fourstatearr[i][j][0]
current_right_state = fourstatearr[i][j][1]
current_down_state = fourstatearr[i][j][2]
current_left_state = fourstatearr[i][j][3]
new_up_state = current_left_state
new_right_state = current_up_state
arr[i][j][0]=new_up_state
arr[i][j][1]=new_right_state
return arr.astype(int)
# +
#Flip horizontally
#Up becomes right, left becomes up, down becomes left, right becomes down
def hor_flip(arr2):
'''
Author: <NAME> and <NAME>
~Function Description~
'''
arr = np.flip(arr2,1)
proper_arr=np.zeros_like(arr2)
num = len(arr)
for i in range(num):
for j in range(num):
current_up_state = arr[i][j][0]
current_left_state = arr[i][j][1]
right_neighbour_left_state = arr[i][(j+1)%num][1]
current_right_state = - (right_neighbour_left_state)
proper_arr[i][j][0]=current_up_state
proper_arr[i][j][1]=current_right_state
return proper_arr.astype(int)
#Flip vertically
#Up becomes right, left becomes up, down becomes left, right becomes down
def ver_flip(arr2):
'''
Author: <NAME> and <NAME>
~Function Description~
'''
arr = np.flip(arr2,0)
proper_arr=np.zeros_like(arr2)
num = len(arr)
for i in range(num):
for j in range(num):
current_down_state = arr[i][j][0]
current_right_state = arr[i][j][1]
upper_neighbour_down_state = arr[i-1][j][0]
current_up_state = - (upper_neighbour_down_state)
proper_arr[i][j][0]=current_up_state
proper_arr[i][j][1]=current_right_state
return proper_arr.astype(int)
# +
def flip_secondary_diag(arr2):
'''
Author: <NAME>
~Function Description~
'''
arr = copy.deepcopy(arr2)
N = len(arr)
for i in range(N):
for j in range(N):
if (i+j)<=N-1:
dist = N-(i+j+1)
arr[i][j][0], arr[i+dist][j+dist][0], arr[i][j][1], arr[i+dist][j+dist][1] = arr[i+dist][j+dist][1], arr[i][j][1], arr[i+dist][j+dist][0], arr[i][j][0]
return arr.astype(int)
def flip_primary_diag(arr2):
'''
Author: <NAME>
~Function Description~
'''
arr = copy.deepcopy(arr2)
N = len(arr)
arr = rot90_anticlock(flip_secondary_diag(rot270_anticlock(arr)))
return arr.astype(int)
# -
def get_all_column_translations(arr):
'''
Author: <NAME>
~Function Description~
'''
result_arr_list=[]
N=len(arr)
for i in range(1,N):
a1 = arr[:,0:i].reshape(N,-1,2)
a2 = arr[:,i:].reshape(N,-1,2)
res = np.hstack([a2,a1])
result_arr_list.append(res)
return result_arr_list
def get_all_row_translations(arr):
'''
Author: <NAME>
~Function Description~
'''
result_arr_list=[]
N=len(arr)
for i in range(1,N):
a1 = arr[0:i,:].reshape(-1,N,2)
a2 = arr[i:,:].reshape(-1,N,2)
res = np.vstack([a2,a1])
result_arr_list.append(res)
return result_arr_list
def arr_to_string(arr2):
'''
Author: <NAME>
~Function Description~
'''
arr = copy.deepcopy(arr2)
name = ' '.join(map(str, arr.flatten())).replace(' ','')
return name
def string_to_arr(s):
'''
Author: <NAME>
~Function Description~
'''
replaced_str = s.replace("-1","0")
arr=[]
for i in replaced_str:
if i=='1':
arr.append(1)
elif i=="0":
arr.append(-1)
else:
print("ERROR")
assert(1==0)
arr = np.array(arr)
arr = arr.reshape(int(np.sqrt(len(arr)/2)),int(np.sqrt(len(arr)/2)),2)
return arr
def remove_symmetries(all_names, verbose=False):
'''
Author: <NAME>
~Function Description~
'''
assert type(all_names)==list
for i,given_name in enumerate(all_names):
if verbose and i%100 ==0:
print(f"Loading... {i}/{len(all_names)} done. Percent Completed = {100*i/len(all_names)}.", end="\r",flush=True)
# print("*******************************")
# print(f"Original Name = {given_name}")
arr = string_to_arr(given_name)
#Column Translation symmetries
templist=get_all_column_translations(arr)
for newarr in templist:
name = arr_to_string(newarr)
# print(f"Col Trans Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Row Translation symmetries
templist=get_all_row_translations(arr)
for newarr in templist:
name = arr_to_string(newarr)
# print(f"Row Trans Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Check 90 degree rotation symmetry
name = arr_to_string(rot90_anticlock(arr))
# print(f"Rot 90 Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Check 180 degree rotation symmetry
name = arr_to_string(rot180_anticlock(arr))
# print(f"Rot 180 Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Check 270 degree rotation symmetry
name = arr_to_string(rot270_anticlock(arr))
# print(f"Rot 270 Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Check horizontal flip symmetry
name = arr_to_string(hor_flip(arr))
# print(f"Flip Hor Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Check vertical flip symmetry
name = arr_to_string(ver_flip(arr))
# print(f"Flip Ver Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Check secondary diagonal flip symmetry
name = arr_to_string(flip_secondary_diag(arr))
# print(f"Sec Diag Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
#Check primary diagonal flip symmetry
name = arr_to_string(flip_primary_diag(arr))
# print(f"Prim Diag Name = {name}")
if name in all_names[i+1:]:
idx = all_names[i+1:].index(name) + i+1
del all_names[idx]
i = all_names.index(given_name)
return all_names
# # ENERGETIC: F1
arr = [[[-1,1],[-1,1],[1,-1],[-1,1]],
[[-1,-1],[-1,-1],[-1,1],[1,-1]],
[[-1,1],[-1,1],[1,1],[-1,1]],
[[-1,1],[-1,1],[1,1],[-1,1]]]
arr = np.array(arr)
def calculate_energy(arr,eps=1):
'''
Author: <NAME>
~Function Description~
'''
N = len(arr)
s = 0
for i in range(len(arr)):
for j in range(len(arr)):
current_up_state = arr[i][j][0]
current_right_state = arr[i][j][1]
lower_neighbour_up_state = arr[(i+1)%N][j][0]
left_neighbour_right_state = arr[i][j-1][1]
if current_up_state ==1 and lower_neighbour_up_state == -1 :
s+= 1
elif current_right_state == 1 and left_neighbour_right_state == -1:
s+= 1
return (-s*eps)
def calculate_atom_config(arr):
'''
Author: <NAME>
~Function Description~
'''
tupdict = {
(1,1,-1,-1):1,
(-1,-1,1,1):2,
(1,-1,-1,1):3,
(-1,1,1,-1):4,
(-1,1,-1,1):5,
(1,-1,1,-1):6
}
num=len(arr)
typearr = np.zeros((arr.shape[0],arr.shape[1]))
for i in range(len(arr)):
for j in range(len(arr)):
current_up_state = arr[i][j][0]
current_right_state = arr[i][j][1]
lower_neighbour_up_state = arr[(i+1)%num][j][0]
left_neighbour_right_state = arr[i][j-1][1]
current_down_state = -(lower_neighbour_up_state)
current_left_state = -(left_neighbour_right_state)
tup = (current_up_state,current_right_state,current_down_state,current_left_state)
atomtype = tupdict[tup]
typearr[i][j] = atomtype
return typearr.astype(int)
def calculate_energy_using_atom_config(arr, eps):
'''
Author: <NAME>
~Function Description~
'''
atom_config_arr = calculate_atom_config(arr).flatten()
energy = len(np.where(atom_config_arr==5)[0])+len(np.where(atom_config_arr==6)[0])
energy = energy * eps
return energy
def visualise_atom_config_with_bonds(arr,savefig=False,savename=".temp"):
'''
Author: <NAME>
~Function Description~
'''
num = len(arr)
plt.axes().set_aspect('equal')
coord_list = get_coord_list(arr)
atom_config_arr = calculate_atom_config(arr)
for i in range(num):
for j in range(num):
current_up_state = arr[i][j][0]
current_right_state = arr[i][j][1]
x_current = coord_list[i][j][1]
y_current = coord_list[i][j][0]
lower_neighbour_up_state = arr[(i+1)%num][j][0]
x_up = coord_list[(i+1)%num][j][1]
y_up = coord_list[(i+1)%num][j][0]
left_neighbour_right_state = arr[i][j-1][1]
x_left = coord_list[i][j-1][1]
y_left = coord_list[i][j-1][0]
current_down_state = -(lower_neighbour_up_state)
current_left_state = -(left_neighbour_right_state)
if atom_config_arr[i][j]==5:
plt.plot(x_current,y_current,
marker="o", markersize=9, markeredgecolor="k",
markerfacecolor="cyan",
zorder=1)
elif atom_config_arr[i][j]==6:
plt.plot(x_current,y_current,
marker="o", markersize=9, markeredgecolor="k",
markerfacecolor="magenta",
zorder=1)
else:
plt.plot(x_current,y_current,
marker="o", markersize=9, markeredgecolor="k",
markerfacecolor="gray",
zorder=1)
if current_up_state == 1:
plot_vector([x_current,y_current],[x_current,y_current+1])
elif current_up_state == -1:
plot_vector([x_current,y_current+1],[x_current,y_current])
if current_right_state == 1:
plot_vector([x_current,y_current],[x_current+1,y_current])
elif current_right_state == -1:
plot_vector([x_current+1,y_current],[x_current,y_current])
if current_down_state == 1:
plot_vector([x_current,y_current],[x_current,y_current-1])
elif current_down_state == -1:
plot_vector([x_current,y_current-1],[x_current,y_current])
if current_left_state == 1:
plot_vector([x_current,y_current],[x_current-1,y_current])
elif current_left_state == -1:
plot_vector([x_current-1,y_current],[x_current,y_current])
plt.xlim(-1,num+1)
plt.ylim(-1,num+1)
plt.axis('off')
if savefig:
plt.savefig(f"{savename}.png",dpi=300)
plt.show()
plt.close()
def visualise_final_state(arr,savefig=False,savename=".temp"):
'''
Author: <NAME> and <NAME>
~Function Description~
'''
plt.axes().set_aspect('equal')
atom_config_arr = calculate_atom_config(arr)
colordict = {1:0,2:0,3:0,4:0,5:-1,6:1}
atom_config_arr_copy = np.copy(atom_config_arr)
for key, val in colordict.items(): atom_config_arr_copy[atom_config_arr==key] = val
flipped_arr = np.flip(atom_config_arr_copy,0)
plt.pcolormesh(flipped_arr,cmap=plt.cm.coolwarm)
if savefig:
plt.savefig(f"{savename}.png",dpi=300)
plt.show()
plt.close()
def metropolis_move(arr,temp,eps=1,verbose = False):
'''
Author: <NAME>
~Function Description~
Dependencies: Uses energy(arr) function
and long_loop(arr) function.
'''
new = long_loop(arr, verbose = verbose)
old_energy = calculate_energy(arr,eps)
new_energy = calculate_energy(new,eps)
delta_E = new_energy - old_energy
if temp >0:
beta = 1/(temp)
if delta_E <0:
arr = new
elif np.random.uniform() < np.exp(-beta*delta_E):
arr = new
elif temp==0:
if delta_E <0:
arr = new
return arr
def calculate_polarization(arr):
'''
Author: <NAME>
~Function Description~
'''
vert = 0
hor = 0
N = len(arr)**2
for i in arr:
for j in i:
vert+=j[0]
hor+=j[1]
polarization= ((1/((2**0.5)*N))*hor,(1/((2**0.5)*N))*vert)
polval = np.sqrt(polarization[0]**2 + polarization[1]**2)
return polval
# +
#Temp array, Energy @that temp array
# -
def equilibrate(n,temp,breaking_iters=3000,eps=1,verbose=False):
'''
Author: Team ℏ
~Function Description~
'''
assert temp>=0
arr = initialise_state(n)
polar_list=[]
iter_list=[]
polval = calculate_polarization(arr)
for i in range(breaking_iters):
if verbose:
print(f"Pol = {polval} and iter = {i}",end="\r",flush=True)
arr=metropolis_move(arr,temp,eps=eps)
newpolval = calculate_polarization(arr)
polar_list.append(polval)
polval=newpolval
return polar_list,arr
np.array([1,2,3]).mean()
def get_sp_heat_and_energy(n,temp,breaking_iters=3000):
polar_list, arr = equilibrate(n,temp,breaking_iters=breaking_iters)
energy_list=[]
for i in range(100):
arr=metropolis_move(arr,temp)
en=calculate_energy(arr,eps=1)
energy_list.append(en)
energy_list = np.array(energy_list)
energy = energy_list.mean()
beta = 1/temp
sp_heat = beta**2 * ((energy_list**2).mean() - (energy_list.mean())**2)/n**2
return sp_heat, energy
temp_arr=np.linspace(0.1,4.5,20)
sp_heat=[]
energy_list=[]
for temp in temp_arr:
print(f"******* TEMP = {temp} *******")
sp,en = get_sp_heat_and_energy(n=25,temp=temp,breaking_iters=4000)
sp_heat.append(sp)
energy_list.append(en)
sp_heat
plt.plot(temp_arr,sp_heat)
plt.xlabel("Temperature")
plt.ylabel("Specific Heat")
plt.title("T vs C")
plt.show()
plt.plot(temp_arr,energy_list)
plt.xlabel("Temperature")
plt.ylabel("Energy")
plt.title("E vs C")
plt.show()
# +
# def get_spec_heat(arr,temperature,energy,eps=1):#eps is energy for config 5
# '''
# Author: <NAME>
# ~Function Description~
# '''
# # kb=1.38064 *10**(-23)
# kb=1
# diversion=np.zeros(n)
# energy_div=np.zeros(n)
# spec_heat=np.zeros(n)
# n=len(temperature)
# N=len(arr)
# for i in range(n):
# if i==0:
# spec_heat[i]=abs(energy[i+1]-energy[i])/h #O(h) differentiation at the end points
# elif i==n-1:
# spec_heat[i]=abs(energy[i]-energy[i-1])/h #O(h) differentiation at the end points
# else:
# spec_heat[i]=abs((energy[i+1]-energy[i-1])/2*h) #O(h^2) differentiation
# diversion[i]=abs((spec_heat[i]/N**2)-kb*((ln2)**2)*(28/45))
# energy_div[i]=abs((energy[i]/N**2)-(eps/3))
# minpos = diversion.index(min(diversion))
# print("The critical temp found is at",temp_plot[minpos]) #ye minpos +1 kar
# print("The error in Specific Heat per vertex is",min(diversion))
# plt.scatter(temperature, spec_heat)
# plt.plot(temperature, spec_heat)
# plt.xlabel("Temperature")
# plt.ylabel("Specific Heat")
# plt.title("T vs C")
# plt.show()
# return spec_heat
# -
# ## ERROR MODELLING: TRUE AND APPROX ERROR COMPARISONS FOR N=1,2,3,4,5
true_val_dict = {1:4,2:18,3:148,4:2940,5:142815}
def count_states_err(num,error_threshold,return_dict = False,verbose=False):
true_err_list = []
approx_err_list = []
if not (error_threshold<=100 and error_threshold>0):
print("Error! Please input error_threshold as a value between 0 and 100")
assert (error_threshold<=100 and error_threshold>0)
state_dict={}
oldarr = long_loop(initialise_state(num), verbose=False)
good_iterations = 0 #Iterations that gave us a new state, so good.
bad_iterations = 0 #Iterations that gave us an already found state,so a waste and hence bad.
while True:
newarr = long_loop(oldarr,verbose=False)
name =arr_to_string(newarr)
if name not in state_dict:
count_repetitions=0
state_dict[name]=1
good_iterations+=1
else:
bad_iterations+=1
count_repetitions+=1
state_dict[name]+=1
percent_approx_err=good_iterations*100/(good_iterations+bad_iterations)
if verbose:
print(f"Good iterations = {good_iterations} and bad iterations = {bad_iterations} and Error % = {percent_approx_err}", end="\r",flush=True)
true_err_list.append((true_val_dict[num] - good_iterations)*100/true_val_dict[num])
approx_err_list.append(percent_approx_err)
if percent_approx_err < error_threshold:
break
oldarr=newarr
if return_dict:
return len(state_dict),state_dict,true_err_list,approx_err_list
else:
return len(state_dict),true_err_list,approx_err_list
count_states_err(4,5,verbose=True)
errorlistdict={}
for n in true_val_dict.keys():
t1=time.process_time()
print(f"************ N = {n} ************")
cnt, true_err_list,approx_err_list = count_states_err(n,5)
errorlistdict[n]=[true_err_list,approx_err_list]
t2=time.process_time()
print(f"Completed in {t2-t1} seconds.")
for n in errorlistdict.keys():
true=errorlistdict[n][0]
approx=errorlistdict[n][1]
plt.plot(true, label='True Error')
plt.plot(approx, label='Approx Error')
plt.legend()
plt.title(f"For {n}x{n} grid")
plt.xlabel("No. of iterations")
plt.ylabel("% Error")
plt.show()
|
Archived IPYNBs/ColdAsIcev8.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os
from sklearn import datasets
from sklearn.cluster import KMeans
import sklearn.metrics as sm
import sklearn.preprocessing as pre
import numpy as np
import pandas as pd
import os
import matplotlib as mpl
import matplotlib.pyplot as plt
# +
#Show the available images
#rootdir = "/data10/shared/jlogan/wensi20170523/luad"
rootdir = "/data01/shared/tcga_analysis/seer_data/results"
allfiles = os.listdir(rootdir)
imgdirs = [f for f in allfiles if not f.endswith(".svs") ]
zip (range(len(imgdirs)), imgdirs) #show image names with indices
# -
#Get a list of all of the csv files across all images, picking the first set of results for each
allcsv = []
for imgdir in imgdirs:
imgroot = "%s/%s"%(rootdir,imgdir)
first_seg = "%s/%s"%(imgroot,os.listdir(imgroot)[0])
#allcsv = os.listdir(first_seg)
csvfiles = ["%s/%s"%(first_seg,f) for f in os.listdir(first_seg) if f.endswith("features.csv")]
allcsv.extend (csvfiles)
allcsv
#Perform a query by loading all data frames and filtering desired results
million = 1000000
results = []
for csvfile in allcsv:
df = pd.read_csv(csvfile, index_col=None, header=0)
results.extend(df.loc[df['AreaInPixels'] > 1*million])
# +
frame = pd.DataFrame()
flist = []
for f in csvfiles:
#Use pandas to read csv...
df = pd.read_csv(f, index_col=None, header=0, usecols=range(93))
#df = pd.read_csv(f, index_col=None, header=0, usecols=[0,1,2,3])
#df = pd.read_csv(f, index_col=None, header=0)
df['file'] = f
flist.append(df)
frame = pd.concat(flist, ignore_index=True)
frame
# -
|
explore/seer_explore.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# # MNIST BASICS
from fastai.vision.all import *
#from fastbook import *
path = untar_data(URLs.MNIST_SAMPLE)
path.ls()
(path/'train').ls()
threes = (path/'train'/'3').ls().sorted()
sevens = (path/'train'/'7').ls().sorted()
threes
img3_path = threes[0]
img3 = Image.open(img3_path)
img3
type(img3)
# ## IMAGE AS ARRAY / TENSOR
array(img3).shape
array(img3)[4:24,6:17]
img3_t = tensor(img3)
img3_t[4:24,6:17]
# +
import pandas as pd
df = pd.DataFrame(img3_t)
df.style.set_properties(**{'font-size':'6pt'}).background_gradient('Greys')
|
Courses/course.fast.ai/Notebooks/04_01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature lab
import pandas as pd
import numpy as np
import seaborn as sns
import sklearn as sk
import matplotlib.pyplot as plt
from tqdm import tqdm
dataset_path = "../datasets/1.0v/"
# !ls ../datasets/1.0v
# ## Loading our datasets...
infos = pd.read_csv(dataset_path + 'infos.csv', sep='|')
items = pd.read_csv(dataset_path + 'items.csv', sep='|')
orders = pd.read_csv(dataset_path + 'orders.csv', sep='|')
# ## FIle heads
orders.head()
items.head()
infos.head()
# ## Basic stats
# **Conclusion**: Our items have a high ```simulationPrice``` standard deviation, which affects directly our approach.
infos.describe().round(1)
items.describe().round(1)
orders.describe().round(1)
# ### Extracting the dates of promotion
# This may be useful later on the competition. For now, let's just understand we have access to this data.
promotions = infos['promotion'].value_counts()
infos['promotion'].unique()
# ### Removing the items that were never sold
# Taking the ids of the items that were sold at least once...
sold_items_id = orders['itemID'].unique()
sold_items_id
# Items that were sold at least once will be stored in 'sold_items'
sold_items = items[items.itemID.isin(sold_items_id)]
sold_items
# Just making a sanity check to be sure if we really have kept only the items that had been sold at least once...
fact_check = np.array(sold_items.itemID)
fact_check == sorted(sold_items_id)
# We'll extract only the infos about the items that were sold at least once...
sold_infos = infos[infos.itemID.isin(sold_items_id)]
sold_infos
# Another sanity check...
fact_check_2 = np.array(sold_infos.itemID)
fact_check_2 == sorted(sold_items_id)
# ### <span style="color:red">Most revenue items</span>
# It's very strange that some items are sold below the 'recommended retail price'
# Concatenating the itemID column with the revenue column...
sold_items_revenue = pd.DataFrame(pd.concat([sold_items.itemID, sold_infos.simulationPrice - sold_items.recommendedRetailPrice], axis=1, names=['itemID', 'revenue']))
items_revenue = pd.concat([items.itemID, infos.simulationPrice - items.recommendedRetailPrice], axis=1, names=['itemID', 'revenue'])
items_revenue
# ### Demystifying the categories
# Ordering the items by how much they were sold and displaying it's categories (the first position has the id of the most sold item, and so goes on).
category = items[['itemID','category1','category2','category3']]
category_and_orders = pd.merge(orders, category, on='itemID')
category_and_orders_stats = category_and_orders.groupby('itemID').aggregate({'order' : ['sum','count'],'category1': ['mean'], 'category2': ['mean'], 'category3': ['mean']})
category_and_orders_stats = category_and_orders_stats.sort_values(('order', 'sum'), ascending=False)
category_and_orders_stats
# #### How well distribuited are the categories in the orders?
cat1_order_amount = category_and_orders.groupby('category1').aggregate({'order' : ['sum']})
cat1_order_amount.sort_values(('order', 'sum'), ascending=False)
plt.plot(cat1_order_amount)
cat2_order_amount = category_and_orders.groupby('category2').aggregate({'order' : ['sum']})
cat2_order_amount.sort_values(('order', 'sum'), ascending=False)
plt.plot(cat2_order_amount)
cat3_order_amount = category_and_orders.groupby('category3').aggregate({'order' : ['sum']})
cat3_order_amount.sort_values(('order', 'sum'), ascending=False)
plt.plot(cat3_order_amount)
# ### Customer rating evaluation
# Almost 70% of our customer ratings consist of zeros, <span style="color:red">so this feature might not be as useful as I expected at first glance</span>
sold_items.customerRating.hist()
print(f'{len(sold_items.customerRating[sold_items.customerRating == 0])/len(sold_items) * 100}%')
|
dora/pre-processing-features/feature_lab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.8 64-bit (''base'': conda)'
# name: python3
# ---
# # Adaptive partitioning algorithm for mutual information estimation
#
# Python implementation of mutual information estimation, where adaptive partitioning strategy is applied.
#
# ## Reference
# - <NAME>., & <NAME>. (1999). Estimation of the information by an adaptive partitioning of the observation space. IEEE Transactions on Information Theory, 45(4), 1315-1321.
# +
import numpy as np
from minfo.mi_float import mutual_info as mi_cy
from minfo.mi_float import tdmi as TDMI_cy
from minfo.mi_float import tdmi_omp as TDMI_cy_omp
from mutual_info import mutual_info as mi_py
import time
def TDMI_py(dat, n):
"""Time-delay mutual information estimator. (Pure Python Version)
Parameters:
dat (np.ndarray) : 2D array of time series with 2 column.
each column is a variable, and each row is a sample.
n (int) : number of delays, including zero time-lag case.
Returns:
np.ndarray : 1d array of delayed mutual information series.
"""
tdmi = np.zeros(n)
N = dat.shape[0]
for i in range(n):
dat_buffer = np.zeros((N-i, 2))
dat_buffer[:,0] = dat[:N-i,0]
dat_buffer[:,1] = dat[i:,1]
tdmi[i] = mi_py(dat_buffer)
return tdmi
# -
# load sample time series
dat = np.load('sample.npy', allow_pickle=True)
dat.shape
print('[INFO]: Testing mi (Python) ...')
# %timeit mi_py(dat)
print('[INFO]: Testing mi (Cython) ...')
# %timeit mi_cy(dat[:,0], dat[:,1])
n_delay = 100
print('[INFO]: Testing tdmi (Python) ...')
# %timeit TDMI_py(dat, n_delay)
print('[INFO]: Testing tdmi (Cython) ...')
# %timeit TDMI_cy(dat[:,0], dat[:,1], n_delay)
print('[INFO]: Testing tdmi (Cython/OpenMP) ...')
# %timeit TDMI_cy_omp(dat[:,0], dat[:,1], n_delay)
|
example/example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# +
import torch
import torch.nn as nn
import numpy as np
import matplotlib.pyplot as plt
import random
# GPU if available
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
BOARD_WIDTH = 10
BOARD_HEIGHT = 20
BLANK = 0
TEMPLATE_WIDTH = 5
TEMPLATE_HEIGHT = 5
S_SHAPE_TEMPLATE = [['.....',
'.....',
'..OO.',
'.OO..',
'.....'],
['.....',
'..O..',
'..OO.',
'...O.',
'.....']]
Z_SHAPE_TEMPLATE = [['.....',
'.....',
'.OO..',
'..OO.',
'.....'],
['.....',
'..O..',
'.OO..',
'.O...',
'.....']]
I_SHAPE_TEMPLATE = [['..O..',
'..O..',
'..O..',
'..O..',
'.....'],
['.....',
'.....',
'OOOO.',
'.....',
'.....']]
O_SHAPE_TEMPLATE = [['.....',
'.....',
'.OO..',
'.OO..',
'.....']]
J_SHAPE_TEMPLATE = [['.....',
'.O...',
'.OOO.',
'.....',
'.....'],
['.....',
'..OO.',
'..O..',
'..O..',
'.....'],
['.....',
'.....',
'.OOO.',
'...O.',
'.....'],
['.....',
'..O..',
'..O..',
'.OO..',
'.....']]
L_SHAPE_TEMPLATE = [['.....',
'...O.',
'.OOO.',
'.....',
'.....'],
['.....',
'..O..',
'..O..',
'..OO.',
'.....'],
['.....',
'.....',
'.OOO.',
'.O...',
'.....'],
['.....',
'.OO..',
'..O..',
'..O..',
'.....']]
T_SHAPE_TEMPLATE = [['.....',
'..O..',
'.OOO.',
'.....',
'.....'],
['.....',
'..O..',
'..OO.',
'..O..',
'.....'],
['.....',
'.....',
'.OOO.',
'..O..',
'.....'],
['.....',
'..O..',
'.OO..',
'..O..',
'.....']]
PIECES = {'S': S_SHAPE_TEMPLATE,
'Z': Z_SHAPE_TEMPLATE,
'J': J_SHAPE_TEMPLATE,
'L': L_SHAPE_TEMPLATE,
'I': I_SHAPE_TEMPLATE,
'O': O_SHAPE_TEMPLATE,
'T': T_SHAPE_TEMPLATE}
PIECES_IND = {'S': 0,
'Z': 1,
'J': 2,
'L': 3,
'I': 4,
'O': 5,
'T': 6}
PIECES_MARGINS = {'S': [[1,1],[0,1]],
'Z': [[1,1],[1,0]],
'J': [[1,1],[0,1],[1,1],[1,0]],
'L': [[1,1],[0,1],[1,1],[1,0]],
'I': [[0,0],[2,1]],
'O': [[1,0]],
'T': [[1,1],[0,1],[1,1],[1,0]]}
class Tetris:
def __init__(self):
self.board = self.getBlankBoard()
self.current_piece = self.getNewPiece()
def reset(self):
"""
Restarts the game with a blank board and new piece.
Returns: torch tensor
A tensor representing the state.
"""
self.board = self.getBlankBoard()
self.current_piece = self.getNewPiece()
return self.convertToFeatures()
def isOnBoard(self, x, y):
"""
Checks if the position (x,y) is on the board.
Args:
x: int
The x position
y: int
The y position
Returns: Boolean
If (x,y) is on the board.
"""
return 0<=x<BOARD_WIDTH and 0<=y<BOARD_HEIGHT
def getBlankBoard(self):
return np.zeros((BOARD_WIDTH, BOARD_HEIGHT))
def isValidPosition(self, x, y, rotation):
"""
Checks if a piece has a valid position on the board.
Args:
shape: str
The shape of the tetris piece.
x: int
The x position of the piece.
y: int
The y position of the piece.
rotation: int
The rotation of the piece.
Returns: Boolean
If the piece has a valid position on the board.
"""
shape = self.current_piece
for dx in range(TEMPLATE_WIDTH):
for dy in range(TEMPLATE_HEIGHT):
template = PIECES[shape][rotation % len(PIECES[shape])]
if template[dy][dx] == 'O':
board_x_pos, board_y_pos = x + dx - 2, y + dy - 2
if not self.isOnBoard(board_x_pos, board_y_pos) or self.board[board_x_pos][board_y_pos]:
return False
return True
def getNewPiece(self):
return random.choice(list(PIECES.keys()))
def getNextState(self, action):
"""
Returns the next state given the current action.
Args:
action: int
An integer representing the action chosen.
In total, there are BOARD_WIDTH x 4 actions, representing
choices in the x coordinate and rotation of the piece.
For a chosen x and rotation r, the action is 4 * x + r.
Returns: tuple
A tuple (reward, next_state, done) representing the reward, next state,
and if the game has finished.
"""
rotation = action % 4
left_margin, right_margin = PIECES_MARGINS[self.current_piece][rotation % len(PIECES_MARGINS[self.current_piece])]
x = max(left_margin, min(action // 4, BOARD_WIDTH - right_margin - 1))
for y in range(BOARD_HEIGHT):
if self.isValidPosition(x, y, rotation):
self.placeOnBoard(x, y, rotation)
lines_cleared = self.clearLines()
#delta_r, delta_c = self.countHoles()
#reward = lines_cleared**2/16
#- delta_r/(BOARD_HEIGHT*BOARD_WIDTH) - delta_c/(BOARD_HEIGHT*BOARD_WIDTH))/2
reward = lines_cleared**2/16
self.current_piece = self.getNewPiece()
next_state = self.convertToFeatures()
return reward, next_state, False, lines_cleared
return -1, None, True, 0
def placeOnBoard(self, x, y, rotation):
"""
Places the current piece on the board. Assumes that the piece
is in a valid position.
Args:
x: int
The x position of the piece.
y: int
The y position of the piece.
rotation: int
The rotation of the piece.
Returns: None
"""
template = PIECES[self.current_piece][rotation % len(PIECES[self.current_piece])]
for dx in range(TEMPLATE_WIDTH):
for dy in range(TEMPLATE_HEIGHT):
if template[dy][dx] == 'O':
board_x_pos, board_y_pos = x + dx - 2, y + dy - 2
self.board[board_x_pos][board_y_pos] = 1
def clearLines(self):
"""
Removes completed lines from the board.
Returns: int
The number of lines removed.
"""
lines_removed = 0
y = 0 # start y at the bottom of the board
while y < BOARD_HEIGHT:
if self.isCompleteLine(y):
# Remove the line and pull boxes down by one line.
for pull_down_Y in range(y, BOARD_HEIGHT-1):
for x in range(BOARD_WIDTH):
self.board[x][pull_down_Y] = self.board[x][pull_down_Y + 1]
# Set very top line to blank.
for x in range(BOARD_WIDTH):
self.board[x][BOARD_HEIGHT-1] = BLANK
lines_removed += 1
# Note on the next iteration of the loop, y is the same.
# This is so that if the line that was pulled down is also
# complete, it will be removed.
else:
y += 1 # move on to check next row up
return lines_removed
def isCompleteLine(self, y):
"""
Checks if the line at height y is complete.
Args:
y: int
The height of the row to check.
Returns: Boolean
True if the row is complete.
"""
for x in range(BOARD_WIDTH):
if not self.board[x][y]: return False
return True
def convertToFeatures(self):
"""
Converts the current board position and falling piece to a
list of features.
The features consist of:
- 7 entries representing a 1 hot vector for the current piece.
- BOARD_WIDTH entries representing the maximum height for each column.
- BOARD_WIDTH - 1 entries representing the difference in heights between successive columns.
Returns: torch tensor
Torch tensor of the features described above. Values normalized to be between -1 and 1.
"""
features = torch.zeros(len(PIECES) + 2 * BOARD_WIDTH - 1)
# One hot vector for the current piece
features[PIECES_IND[self.current_piece]] = 1.0
# Maximum heights of each column
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT-1, -1, -1):
if self.board[x][y]: break
features[len(PIECES) + x] = y/BOARD_HEIGHT
# Differences in heights between each column
for x in range(BOARD_WIDTH-1):
features[len(PIECES) + BOARD_WIDTH + x] = (features[len(PIECES) + x + 1] - features[len(PIECES) + x])/BOARD_HEIGHT
return features.to(device)
def countHoles(self):
"""
Counts the number of transitions from filled to empty or vice
versa in the rows and columns.
Returns: tuple[int]
A tuple (delta_r, delta_c) representing the number of transitions
from filled to empty squares or vice versa across rows and columns respectively.
"""
# Across rows:
delta_r = 0
for y in range(BOARD_HEIGHT):
for x in range(BOARD_WIDTH-1):
if self.board[x][y] != self.board[x+1][y]:
delta_r += 1
# Across columns:
delta_c = 0
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT-1):
if self.board[x][y] != self.board[x][y+1]:
delta_c += 1
return delta_r, delta_c
class Agent:
""" Agent object that uses the actor-critic network to find the
optimal policy.
"""
def __init__(self, env, NN_value, NN_pi):
""" Initializes the agent.
@type env: Tetris
The Tetris environment.
@type NN_value: NeuralNet
Neural network for computing the state values.
@type NN_pi: NeuralNet
Neural network for computing the policy.
"""
self.env = env
self.NN_value = NN_value
self.NN_pi = NN_pi
def chooseAction(self, state):
""" Chooses action according to the current policy.
@type state: torch tensor
A torch tensor representing the current state.
@rtype: int
An integer representing the action.
"""
with torch.no_grad():
action_probs = self.NN_pi(state).detach().cpu().numpy()
action_probs = action_probs.astype('float64') # More precision for normalization
action_probs /= action_probs.sum() # Normalize probabilities so it sums closer to 1
return np.random.choice(range(len(action_probs)), p=action_probs)
def initializeActorTrace(self):
"""
Initializes the eligibility trace for the policy network.
@rtype: dict
A dictionary that maps the layers of the policy network
to their eligibility traces.
"""
z_theta = {}
with torch.no_grad():
for p in self.NN_pi.parameters():
z_theta[p] = torch.zeros(size=p.data.size()).to(device)
return z_theta
def initializeCriticTrace(self):
"""
Initializes the eligibility trace for the value network.
@rtype: dict
A dictionary that maps the layers of the value network
to their eligibility traces.
"""
z_w = {}
with torch.no_grad():
for p in self.NN_value.parameters():
z_w[p] = torch.zeros(size=p.data.size()).to(device)
return z_w
def oneStepActor(self, state, action, delta, gamma, I, z_theta, alpha_theta, lmbda_theta):
""" Performs one training step on observed transition.
@type state: torch tensor
A tensor denoting the position.
@type action: int
An integer representing the action.
@type delta: float
The TD error.
@type gamma: float
The discount factor.
@type I: float
A number that multiples the gradient in policy gradient updates.
@type z_theta: dict
A dictionary that maps actor model parameters to their
eligibility traces.
@type alpha_theta: float
The step size.
@type lmbda_theta: float
Trace decay parameter.
@rtype: tuple
A tuple (dict, I) denoting the updated eligibility trace dictionary
and the new I value.
"""
pred = self.NN_pi(state)
self.NN_pi.zero_grad()
pred[action].backward()
# Update eligibility trace and parameters
with torch.no_grad():
prob = pred[action]
for p in self.NN_pi.parameters():
z_theta[p] = gamma * lmbda_theta * z_theta[p] + I * p.grad / prob
p.copy_(p + alpha_theta * delta * z_theta[p])
return z_theta, gamma * I
def oneStepCritic(self, state, delta, gamma, z_w, alpha_w, lmbda_w):
""" Performs one training step on observed transition.
@type state: torch tensor
A torch tensor denoting the position.
@type delta: float
The TD error.
@type gamma: float
The discount factor.
@type z_w: dict
A dictionary that maps value model parameters to their
eligibility traces.
@type alpha_w: float
The step size.
@type lmbda_w: float
Trace decay parameter.
@rtype: dict
A dictionary denoting the updated eligibility trace dictionary.
"""
pred = self.NN_value(state)
self.NN_value.zero_grad()
pred[0].backward()
# Update eligibility trace and parameters
with torch.no_grad():
for p in self.NN_value.parameters():
z_w[p] = gamma * lmbda_w * z_w[p] + p.grad
p.copy_(p + alpha_w * delta * z_w[p])
return z_w
def train(self, episodes, gamma, alpha_w, alpha_theta, lmbda_w, lmbda_theta):
""" Trains the agent using the actor-critic method with eligibility traces.
@type episodes: int
The number of episodes to train.
@type gamma: float
The discount factor.
@type alpha_w: float
Step size for the value network (critic).
@type alpha_theta: float
Step size for the policy network (actor).
@type lmbda_w: float
Trace decay parameter for the value network (critic).
@type lmbda_theta: float
Trace decay parameter for the policy network (actor).
"""
tot_steps = 0
LC = 0
for episode in range(episodes):
if (episode + 1) % 10 == 0:
print(f'Episode {episode + 1}/{episodes} completed!')
torch.save(self.NN_value.state_dict(), 'tetris_NN_value_model')
torch.save(self.NN_pi.state_dict(), 'tetris_NN_pi_model')
print(f'Average steps per episode: {tot_steps/10}')
print(f'Average lines cleared per episode: {LC/10}')
tot_steps = 0
LC = 0
state, done = self.env.reset(), False
# Initialize eligibility traces
z_w = self.initializeCriticTrace()
z_theta = self.initializeActorTrace()
I = 1.0
while not done:
tot_steps += 1
action = self.chooseAction(state)
reward, next_state, done, lines_cleared = self.env.getNextState(action)
LC += lines_cleared
# TD error
target = reward + gamma * self.NN_value(next_state) if not done else reward
delta = target - self.NN_value(state)
z_w = self.oneStepCritic(state, delta, gamma, z_w, alpha_w, lmbda_w)
z_theta, I = self.oneStepActor(state, action, delta, gamma, I, z_theta, alpha_theta, lmbda_theta)
state = next_state
class QNetwork(nn.Module):
def __init__(self, input_size, hidden_size1, hidden_size2):
super(QNetwork, self).__init__()
self.l1 = nn.Linear(input_size, hidden_size1)
self.l2 = nn.Linear(hidden_size1, hidden_size2)
self.l3 = nn.Linear(hidden_size2, 1)
def forward(self, x):
x = torch.tanh(self.l1(x))
x = torch.tanh(self.l2(x))
x = self.l3(x)
return x
class piNetwork(nn.Module):
def __init__(self, input_size, hidden_size1, hidden_size2, action_size):
super(piNetwork, self).__init__()
self.l1 = nn.Linear(input_size, hidden_size1)
self.l2 = nn.Linear(hidden_size1, hidden_size2)
self.l3 = nn.Linear(hidden_size2, action_size)
def forward(self, x):
x = torch.tanh(self.l1(x))
x = torch.tanh(self.l2(x))
x = torch.softmax(self.l3(x), dim=-1)
return x
if __name__ == "__main__":
# Network parameters
input_size = len(PIECES) + 2 * BOARD_WIDTH - 1
action_size = BOARD_WIDTH * 4
hidden_size1 = 20
hidden_size2 = 20
# Training parameters
episodes = 1000000
gamma = 1.0
alpha_w = 1e-3
alpha_theta = 1e-3
lmbda_w = 0.85
lmbda_theta = 0.85
env = Tetris()
model_value = QNetwork(input_size, hidden_size1, hidden_size2).to(device)
model_pi = piNetwork(input_size, hidden_size1, hidden_size2, action_size).to(device)
#model_value.load_state_dict(torch.load('tetris_NN_value_model'))
#model_pi.load_state_dict(torch.load('tetris_NN_pi_model'))
tetris_agent = Agent(env, model_value, model_pi)
tetris_agent.train(episodes, gamma, alpha_w, alpha_theta, lmbda_w, lmbda_theta)
# +
def animate(env, actions):
# %matplotlib
fig = plt.gcf()
fig.show()
fig.canvas.draw()
plt.grid()
for action in actions:
time.sleep(0.2)
state, reward, done = env.one_step(action)
if done:
break
falling_piece_shape = state[1]['shape']
next_piece_shape = state[2]['shape']
plt.title('Action: ' + str(action) + ', Cur: ' + falling_piece_shape + ', Next: ' + next_piece_shape)
board = state[0]
print(np.shape(board))
print(type(board))
plt.imshow(np.transpose(board), cmap=plt.cm.binary, interpolation='none')
width = len(board)
height = len(board[0])
plt.xlim(-0.5, width-0.5)
plt.ylim(height-0.5, 0.5)
ax = plt.gca()
ax.set_xticks(np.arange(-0.5, width-0.5, 1))
ax.set_yticks(np.arange(0.5, height-0.5, 1))
fig.canvas.draw()
if __name__ == "__main__":
random.seed(2)
env = Env()
action_length = 100
actions = [random.randint(0, 5) for _ in range(action_length)]
animate(env, actions)
plt.show()
# +
BOARD_WIDTH = 10
BOARD_HEIGHT = 20
board1 = np.random.randint(2, size = (BOARD_WIDTH, 10))
board2 = board = np.random.randint(1, size = (BOARD_WIDTH, 10))
board = np.concatenate((board2, board1), axis=1)
def is_valid_position(board, shape,px,py,rot, adjX, adjY):
"""Return whether the falling piece is within the board and not colliding,
after adding (adjX, adjY) to the current coordinates (x, y) of the falling piece.
Args:
adjX (int): move x-coordinate by adjX
adjY (int): move y-coordinate by adjY
Returns:
Boolean value. True if the resulting coordinate is in valid position. False otherwise.
"""
for x in range(TEMPLATE_WIDTH):
for y in range(TEMPLATE_HEIGHT):
is_above_board = y + py + adjY < 0
if is_above_board or PIECES[shape][rot][y][x] == '.':
continue
if not is_on_board(x + px + adjX, y + py + adjY):
return False
if board[x + px + adjX][y + py + adjY] != BLANK:
return False
return True
def compute_metric(board, shape, px, py, rot):
H = []
PA = PIECES[shape][rot]
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT):
if board[x][y] or (0<=y-py<5 and 0<=x-px<5 and PA[y-py][x-px]=='O'): break
H.append(BOARD_HEIGHT-y)
height = sum(H)
bump = sum([abs(H[i]-H[i-1]) for i in range(1, len(H))])
lines = 0
for y in range(BOARD_HEIGHT):
b = False
for x in range(BOARD_WIDTH):
if not (board[x][y] or (0<=y-py<5 and 0<=x-px<5 and PA[y-py][x-px]=='O')):
b = True
break
if not b: lines += 1
holes = 0
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT):
if board[x][y] or (0<=y-py<5 and 0<=x-px<5 and PA[y-py][x-px]=='O'):
for t in range(y, BOARD_HEIGHT):
if not (board[x][t] or (0<=t-py<5 and 0<=x-px<5 and PA[t-py][x-px]=='O')): holes += 1
break
return -0.51*height+0.76*lines-0.186*bump-0.35*holes
def find_best(board, shape):
baseline_metric = -float('inf')
bestx,besty,bestr = 0,0,0
for rot in range(len(PIECES[shape])):
for x in range(-3, BOARD_WIDTH+3):
if not is_valid_position(board, shape, x, 0, rot,0, 0): continue
for y in range(BOARD_HEIGHT+3):
if not is_valid_position(board, shape, x, y, rot,0, 0):
y -= 1
m = compute_metric(board, shape, x,y,rot)
if m>=baseline_metric:
baseline_metric = m
bestx,besty,bestr = x,y,rot
break
return bestx,besty,bestr
def is_valid_position(board, shape,px,py,rot, adjX, adjY):
"""Return whether the falling piece is within the board and not colliding,
after adding (adjX, adjY) to the current coordinates (x, y) of the falling piece.
Args:
adjX (int): move x-coordinate by adjX
adjY (int): move y-coordinate by adjY
Returns:
Boolean value. True if the resulting coordinate is in valid position. False otherwise.
"""
for x in range(TEMPLATE_WIDTH):
for y in range(TEMPLATE_HEIGHT):
is_above_board = y + py + adjY < 0
if is_above_board or PIECES[shape][rot][y][x] == '.':
continue
if not is_on_board(x + px + adjX, y + py + adjY):
return False
if board[x + px + adjX][y + py + adjY] != BLANK:
return False
return True
def is_on_board(x, y):
"""Return whether the position (x, y) is on the board.
Args:
x (int): x-coordinate
y (int): y-coordinate
Returns:
Boolean value. True if (x, y) is on the board. False otherwise.
"""
return x >= 0 and x < BOARD_WIDTH and 0<= y < BOARD_HEIGHT
t1 = time.time()
px,py,r = find_best(board, 'Z')
print('Best', px,py,r)
print('Increased Metric', compute_metric(board, 'Z', px, py, r))
for y in range(BOARD_HEIGHT):
for x in range(BOARD_WIDTH):
if 0<=y-py<5 and 0<=x-px<5 and PIECES['Z'][r][y-py][x-px]=='O':
board[x][y]+=2
t2 = time.time()
print(t2-t1)
plt.figure()
plt.imshow(np.transpose(board),cmap=plt.cm.binary, interpolation='none')
# Try using the "right" policy
# penalize holding time more
# -
model = Sequential()
model.add(Conv2D(32,3,activation='relu',padding='same', input_shape=(10,20, 1)))#120
model.add(Conv2D(64,1,activation = 'relu', padding = 'same'))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dense(5, activation='linear'))
model.compile(loss='mse', optimizer = RMSprop(learning_rate=0.01, momentum = 0.95, rho = 0.95, epsilon = 0.01))
# +
fig = plt.figure()
ax1 = fig.add_subplot(121)
ax2 = fig.add_subplot(122)
plt.ion()
ax1.grid()
ax2.grid()
width = 10
height = 20
ax1.set_xlim(-0.5, width-0.5)
ax1.set_ylim(height-0.5, 0.5)
ax1.set_xticks(np.arange(-0.5, width-0.5, 1))
ax1.set_yticks(np.arange(0.5, height-0.5, 1))
ax2.set_xlim(-0.5, width-0.5)
ax2.set_ylim(height-0.5, 0.5)
ax2.set_xticks(np.arange(-0.5, width-0.5, 1))
ax2.set_yticks(np.arange(0.5, height-0.5, 1))
def compute_metric(board, shape, px, py, rot):
H = []
PA = PIECES[shape][rot]
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT):
if board[x][y] or (0<=y-py<5 and 0<=x-px<5 and PA[y-py][x-px]=='O'): break
H.append(BOARD_HEIGHT-y)
height = sum(H)
bump = sum([abs(H[i]-H[i-1]) for i in range(1, len(H))])
lines = 0
for y in range(BOARD_HEIGHT):
b = False
for x in range(BOARD_WIDTH):
if not (board[x][y] or (0<=y-py<5 and 0<=x-px<5 and PA[y-py][x-px]=='O')):
b = True
break
if not b: lines += 1
holes = 0
for x in range(BOARD_WIDTH):
for y in range(BOARD_HEIGHT):
if board[x][y] or (0<=y-py<5 and 0<=x-px<5 and PA[y-py][x-px]=='O'):
for t in range(y, BOARD_HEIGHT):
if not (board[x][t] or (0<=t-py<5 and 0<=x-px<5 and PA[t-py][x-px]=='O')): holes += 1
break
return -0.51*height+0.76*lines-0.35*holes-0.18*bump
def is_valid_position(board, shape,px,py,rot, adjX, adjY):
for x in range(TEMPLATE_WIDTH):
for y in range(TEMPLATE_HEIGHT):
is_above_board = y + py + adjY < 0
if is_above_board or PIECES[shape][rot][y][x] == '.':
continue
if not is_on_board(x + px + adjX, y + py + adjY):
return False
if board[x + px + adjX][y + py + adjY] != BLANK:
return False
return True
def find_best(board, shape):
baseline_metric = -float('inf')
bestx,besty,bestr = 0,0,0
for rot in range(len(PIECES[shape])):
for x in range(-3, BOARD_WIDTH+3):
if not is_valid_position(board, shape, x, 0, rot,0, 0): continue
for y in range(BOARD_HEIGHT+3):
if not is_valid_position(board, shape, x, y, rot,0, 0):
y -= 1
m = compute_metric(board, shape, x,y,rot)
if m>=baseline_metric:
baseline_metric = m
bestx,besty,bestr = x,y,rot
break
return bestx,besty,bestr
t = 30
randBoard = np.random.normal(size = (1,10,20,1))
print('Move: ', memory[t][1])
print('Reward: ', memory[t][2])
print('Piece: ', memory[t][-1])
p = memory[t][-1]
print(model.predict(memory[t][0]))
print(model.predict(memory[t][3]))
print(model.predict(randBoard))
grid = deepcopy(memory[t][0][0,:,:,0])
px,py,r = find_best(grid, p)
for y in range(BOARD_HEIGHT):
for x in range(BOARD_WIDTH):
if 0<=y-py<5 and 0<=x-px<5 and PIECES[p][r][y-py][x-px]=='O':
grid[x][y]+=3
ax1.imshow(np.transpose(grid), cmap=plt.cm.binary, interpolation='none')
grid = deepcopy(memory[t][3][0,:,:,0])
for y in range(BOARD_HEIGHT):
for x in range(BOARD_WIDTH):
if 0<=y-py<5 and 0<=x-px<5 and PIECES[p][r][y-py][x-px]=='O':
grid[x][y]+=3
ax2.imshow(np.transpose(grid), cmap=plt.cm.binary, interpolation='none')
fig.canvas.draw()
# +
import numpy as np
import tensorflow as tf
from tensorflow.keras import backend as K
from tensorflow.keras.layers import Input,Conv2D, Activation
from tensorflow.keras.models import Sequential
from tensorflow.keras.models import clone_model
from tensorflow.keras.layers import Dense, Conv2D, Flatten
from tensorflow.keras.optimizers import RMSprop
from collections import deque
from copy import deepcopy
tf.keras.backend.clear_session()
import gc
matrixSide = 128 #define a big enough matrix to give memory issues
model = Sequential()
model.add(Conv2D(32,3,activation='relu',padding='same', input_shape=(matrixSide, matrixSide, 12)))#120
model.add(Conv2D(64,1,activation = 'relu', padding = 'same'))
model.add(Conv2D(64,3,activation = 'relu', padding = 'same'))
model.add(Conv2D(1,1,activation = 'relu', padding = 'same'))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.compile(loss='mse', optimizer = RMSprop(learning_rate=0.01, momentum = 0.95, rho = 0.95, epsilon = 0.01))
#run predictions
for i in range (30000):
inImm = np.zeros((64,matrixSide,matrixSide,12))
outImm = model.predict(inImm)
# +
# MEMORY LEAK TESTING
import matplotlib.pyplot as plt
import numpy as np
import random
import time
import os
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.models import clone_model
from tensorflow.keras.layers import Dense, Conv2D, Flatten
from tensorflow.keras.optimizers import RMSprop
from collections import deque
from copy import deepcopy
tf.keras.backend.clear_session()
model = Sequential()
model.add(Conv2D(32, kernel_size=4, input_shape=(10,20,1)))
model.add(Conv2D(64, kernel_size=3, activation='relu'))
model.add(Conv2D(64, kernel_size=3, activation='relu'))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dense(8, activation='linear'))
model.compile(loss='mse', optimizer = RMSprop(learning_rate=0.001, momentum = 0.95, rho = 0.95, epsilon = 0.01))
times = deque(maxlen = 500)
for i in range(200000):
tempx = np.zeros((256,10,20,1))
tempy = np.zeros((256,8))
t0 = time.time()
act_values = model.fit(tempx, tempy, epochs = 1, verbose=0, batch_size = 256)
times.append(time.time()-t0)
if i % 50 == 0: print(sum(times)/len(times))
# +
episodes = 30000
updatefreq = 420
# %matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
import random
import time
import os
import tensorflow as tf
tf.compat.v1.disable_eager_execution()
from tensorflow.keras.models import Sequential
from tensorflow.keras.models import clone_model
from tensorflow.keras.layers import Dense, Conv2D, Flatten
from tensorflow.keras.optimizers import RMSprop
from collections import deque
from copy import deepcopy
fig = plt.figure()
ax1 = fig.add_subplot(131)
ax2 = fig.add_subplot(132)
ax3 = fig.add_subplot(133)
plt.ion()
ax1.grid()
width = 10
height = 20
ax1.set_xlim(-0.5, width-0.5)
ax1.set_ylim(height-0.5, 0.5)
ax1.set_xticks(np.arange(-0.5, width-0.5, 1))
ax1.set_yticks(np.arange(0.5, height-0.5, 1))
# initialize gym environment and the agent
totalR = deque(maxlen = 50)
pieces = deque(maxlen = 50)
pieces.append(0)
totalR.append(0)
env = Env()
### AGENT #########################
def build_model():
# Neural Net for Deep-Q learning Model
model = Sequential()
model.add(Conv2D(32, kernel_size=4, activation='relu', input_shape=(10,20,1)))
model.add(Conv2D(64, kernel_size=3, activation='relu'))
model.add(Conv2D(64, kernel_size=3, activation='relu'))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dense(action_size, activation='linear'))
model.compile(loss='mse', optimizer = RMSprop(learning_rate=learning_rate, momentum = 0.95, rho = 0.95, epsilon = 0.01))
return model
def remember(state, action, reward, next_state, done,piece):
# Places the state in memory
memory.append((state, action, reward, next_state, done, piece))
def act(state):
# Selects an action based on epsilon-greedy algorithm
if np.random.rand() <= epsilon:
x = np.random.rand()
if x < 3.0/7: return random.randrange(2)
elif x < 5.0/7: return 3
elif x < 6.0/7: return 2
else: return 4
act_values = model.predict(state)
return np.argmax(act_values[0]) # returns action
def update():
lag_model.set_weights(model.get_weights())
def replay(batch_size):
# Memory replay
minibatch = random.sample(memory, min(batch_size, len(memory)))
States,R,Ns,D,A = [],[],[], [],[]
for state, action, reward, next_state, done, ps in minibatch:
R.append(reward)
D.append(not done)
Ns.append(next_state[0])
A.append(action)
States.append(state[0])
Ns, States = np.array(Ns), np.array(States)
#print(np.shape(Ns[0]))
R,D,A = np.array(R), np.array(D), np.array(A)
#x = lag_model.predict(Ns)
#del States, R, Ns, D, A
#R = R + D*gamma*np.amax(lag_model.predict(Ns), axis = 1)
#model = tf.keras.models.load_model('mymodel')
#targetF = model.predict(States)
#for i in range(len(targetF)):
#targetF[i][A[i]] = targetF[i][A[i]]+max(-1, min(1, R[i]-targetF[i][A[i]]))
#States = deepcopy(States)
#targetF = deepcopy(targetF)
#x = model.fit(States, targetF, epochs=1, verbose=0, batch_size=batch_size).history['loss']
#x = model.train_on_batch(States, targetF)
#model.save('mymodel')
#Errs.append(x)
state_size, action_size = (10,20), 5
memory = deque(maxlen=1000)
gamma = 0.99 # discount rate
epsilon = 0 # exploration rate
epsilon_min = 0.1
learning_rate = 0.001
model = build_model()
#model.save('mymodel')
Errs = deque(maxlen = 100)
Errs.append(0)
lag_model = build_model()
# Iterate the game
j = 0
it = 0
for e in range(episodes):
if e % 10 == 0 and e!=0:
#model.save_weights('my_checkpoint_LargerNetwork_overnight')
ax2.plot(e,sum(totalR)/len(totalR), marker = 'o', markersize = 2, color = 'k')
ax3.plot(e,sum(Errs)/len(Errs), marker = 'o', markersize = 2, color = 'k')
fig.canvas.draw()
print("episode: {}/{}, Avg score: {}, Errs: {}, epsilon: {}"
.format(e, episodes, sum(totalR)/len(totalR), sum(Errs)/len(Errs), epsilon))
nums = 0
# reset state in the beginning of each game
state, reward, done = env.reset()
state = np.reshape(state, [-1,10, 20,1])
# time_t represents each frame of the game
# Our goal is to keep the pole upright as long as possible until score of 500
# the more time_t the more score
total_reward = reward
while not done:
it += 1
# Decide action
#bestx, besty, bestr = env.bestx, env.besty, env.bestr
#if env.falling_piece['rotation']!=bestr: action = 3
#elif env.falling_piece['x'] > bestx: action = 0
#elif env.falling_piece['x']<bestx: action = 1
#else: action = 4
#action = 4
action = act(state)
# Advance the game to the next frame based on the action.
next_state, reward, done,lines, piece = env.one_step(action)
next_state = np.reshape(next_state, [-1,10, 20, 1])
total_reward += reward
# Remember the previous state, action, reward, and done
remember(state, action, reward, next_state, done, piece)
# make next_state the new current state for the next frame.
state = next_state
# done becomes True when the game ends
#if it % 1000 == 0:
#ax2.plot(it/1000.0,sum(agent.Errs)/len(agent.Errs), marker = 'o', markersize = 2, color = 'k')
#fig.canvas.draw()
if done:
# print the score and break out of the loop
#print("episode: {}/{}, score: {}, lines cleared: {}, Avg Time per cycle: {}, epsilon: {}"
#.format(e, episodes, total_reward, lines, totalt/nums, agent.epsilon))
totalR.append(total_reward)
pieces.append(env.pieces)
#ax2.plot(e,total_reward, marker = 'o', markersize = 2, color = 'k')
#ax3.plot(e,agent.epsilon, marker = 'o', markersize = 2, color = 'k')
#fig.canvas.draw()
break
# train the agent with the experience of the episode
if it>2:
t0 = time.time()
replay(256)
#if epsilon>epsilon_min: epsilon -= 0.00001
nums += 1
j += 1
if j >= updatefreq:
#update()
j = 0
#if it % 5 == 0:
#grid = state[0,:,:,0]
#ax1.imshow(np.transpose(grid), cmap=plt.cm.binary, interpolation='none')
#fig.canvas.draw()
# -
model.save_weights('my_checkpoint_LargerNetwork_overnight_lag')
|
.ipynb_checkpoints/Testing-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Healthcare & Medical Analytics
# ### Individual Assignment - <NAME>
# ##### "The relationship between gender and seasonal allergies; influence by socioeconomic factors"
# First we load the necessary libraries.
import string
import pyreadr
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from statsmodels.graphics.mosaicplot import mosaic
# Then we load and visually inspect our data set.
data_orig = pyreadr.read_r('21600-0022-Data.rda')
data_orig = data_orig['da21600.0022']
data_orig
# We then choose the variables that we are interested in.
data_new = data_orig[['H4ID9F','BIO_SEX4','H4ID5I','H4CJ1','H4SE32','H4GH8','H4EC7','H4IR4']]
# After that we give our variables appropriate names.
data_new = data_new.rename(columns ={'H4ID9F':'allergies','BIO_SEX4':'gender','H4ID5I':'stress','H4CJ1':'arrested','H4SE32':'rape','H4GH8':'nutrition','H4EC7':'assets','H4IR4':'ethnicity'})
# Then we clean our data by removing any special characters from the columns needed.
# +
columns = ['allergies','gender','stress','arrested','rape','ethnicity']
for i in columns:
for j in string.punctuation:
data_new[i] = data_new[i].str.lstrip(' ')
data_new[i] = data_new[i].str.replace('\d+','')
data_new[i] = data_new[i].astype(str).str.replace(j,'')
# -
# Afterwards, we prepare our data. First, we pick the variables that we need to use in our model and drop "NaN" values.
data = data_new[['allergies','gender','stress','arrested','rape','nutrition','assets','ethnicity']]
data = data.dropna()
# Let's now reduce the number of categories of some of our categorical variables. We reduce the dimension of the fast-food consumption variable from 8 to 3, and classify the nutrition as "healthy", "normal" and "unhealthy". We also reduce the dimension of total assets besides homes variable from 9 to 3, "poor", "normal" and "wealthy". Moreover, we rename our categories of our ethnicity variable so as to have shorter names.
# +
for i in range(len(data['nutrition'])):
if data['nutrition'].iloc[i] == 0:
data['nutrition'].iloc[i] = 'healthy'
elif data['nutrition'].iloc[i] == 1 or data['nutrition'].iloc[i] == 2:
data['nutrition'].iloc[i] = 'normal'
else:
data['nutrition'].iloc[i] = 'unhealthy'
mapping = {'(1) (1) Less than $5,000': 'poor', '(2) (2) $5,000 to $9,999': 'poor', '(3) (3) $10,000 to $24,999': 'normal', '(4) (4) $25,000 to $49,999': 'normal', '(5) (5) $50,000 to $99,999': 'wealthy','(6) (6) $100,000 to $249,999': 'wealthy','(7) (7) $250,000 to $499,999': 'wealthy','(8) (8) $500,000 to $999,999': 'wealthy','(9) (9) $1,000,000 or more': 'wealthy'}
data['assets'] = data.assets.map(mapping)
mapping2 = {'White': 'white', 'Black or African American': 'black', 'Asian or Pacific Islander': 'asian', 'American Indian or Alaska Native': 'indian'}
data['ethnicity'] = data.ethnicity.map(mapping2)
# -
# Now, in order to deal with perfect multicolinearity issues, after we create dummy variables for each of the categories of our categorical variables, we drop one column.
# +
# remove allergies from our list so as not to create a dummy for it
columns = list(data.columns)
columns.pop(0)
# deal with perfect multicolinearity
model = pd.get_dummies(data, columns = columns, drop_first = True)
# -
# We remove "NaN" dummy columns and we visually inspect our final data for our model.
model = model.drop(['arrested_nan'], axis = 1)
model = model.drop(['rape_nan'], axis = 1)
model
# We store the final data for our model in a csv file for future purposes.
model.to_csv('data_model.csv')
# Lastly, we visually inspect our final data for our descriptive statistics and store them in a csv file for future analysis.
data.to_csv('data_eda.csv')
data
# Let's now draw some descriptive statistics.
# We choose the Mosaic plot from statsmodels, since it gives us statistical highlighting for the variances. In other words, it is a graphical method for visualizing data from two or more qualitative variables. It is the multidimensional extension of spineplots, which graphically display the same information for only one variable. It gives an overview of the data and makes it possible to recognize relationships between different variables. For example, independence is shown when the boxes across categories all have the same areas.
#
# We now draw mosaic plots for several variables.
data.gender.value_counts()
plt.rcParams['font.size'] = 16.0
pd.crosstab(data.allergies, data.gender)
ct = pd.crosstab(data.allergies, data.gender)
mosaic(ct.unstack(), gap = 0.006675, title = 'Allergies per Gender')
plt.show()
plt.rcParams['font.size'] = 16.0
pd.crosstab(data.gender, data.nutrition)
ct = pd.crosstab(data.gender, data.nutrition)
mosaic(ct.unstack(), title = 'Nutrition type per Gender')
plt.show()
# Hence, since the area for unhealthy nutrition for men is larger than that of women and vice versa, we can conclude that men have on average an unhealthier nutrition in comparison with that of women.
# We can see that female are more prone to seasonal allergies than men.
plt.rcParams['font.size'] = 16.0
pd.crosstab(data.gender, data.assets)
ct = pd.crosstab(data.gender, data.assets)
mosaic(ct.unstack(), gap = 0.006675, title = 'Assets per Gender')
plt.show()
# Lastly, we can clearly see that females have on average a lower number of total assets (bank accounts, retirement plans and stocks) in comparison with men, who own more.
|
health_individual.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Apache Ambari - API Fundamentals & Examples
#
# 
#
#
# ## Introduction
#
#
# This living document will give an introduction & ongoing examples for integrating with Ambari's RESTful API.
#
# ### Knowledge Requirements:
#
# - An understanding of RESTful APIs. This will help: [Learn REST: A RESTful Tutorial](http://www.restapitutorial.com/)
# - Basic programming experience. This document uses Python, but the methods can be translated to other languages.
# - Basic understanding of Ambari's functions.
#
# ### How to use this notebook
#
# - a) Browse a read-only verison of the notebook [here](http://nbviewer.ipython.org/github/HortonworksUniversity/Ops_Labs/1.1.0/build/security/ambari-bootstrap/blob/master/api-examples/ambari-api-examples.ipynb).
# - b) or, download the notebook and use [your own ipython Notebook](http://ipython.org/install.html).
# - c) or, have some inception by running ipython notebook from within Ambari!: https://github.com/randerzander/ipython-stack
#
#
# ## Questions & Help
#
# * [Ambari Wiki](https://ambari.apache.org)
# * [Ambari Mailing Lists](https://ambari.apache.org/mail-lists.html)
# * [Github Issues for this Repo](https://github.com/HortonworksUniversity/Ops_Labs/1.1.0/build/security/ambari-bootstrap/issues)
#
#
# ## The Apache Ambari Ecosystem
# ----
#
# * Interfaces:
# * Ambari API
# * Ambari Web UI (Web interface to the API)
#
# * Functions:
# * Management, Metrics, Monitoring, Operations
# * Blueprints
# * Stacks
# * Views
#
# * Backend:
# * Ambari Agent & Server
#
# ## Ambari Web is an API Client
#
# Ambari Web is a graphical front-end to the API.
#
# Note the output on the right showing the requests which my browser is making:
# 
# ## API Examples
#
# Below we will cover:
#
# * Authentication/Sessions
# * Change Password
# * List Clusters
# * List Hosts
# * Upload blueprint
# * Create cluster
# * Export of Cluster's blueprint
# * Show Cluster details
# * List Cluster's hosts
# * Change configuration _(example with `hive.execution.engine`)_
#
# Todo:
#
# * Restart service
# ### API Examples: Authentication
# +
### Authenticate to Ambari
#### Python requirements
import difflib
import getpass
import json
import requests
import sys
import time
#### Change these to fit your Ambari configuration
ambari_protocol = 'http'
ambari_server = '172.28.128.3'
ambari_port = 8080
ambari_user = 'admin'
#cluster = 'Sandbox'
#### Above input gives us http://user:pass@hostname:port/api/v1/
api_url = ambari_protocol + '://' + ambari_server + ':' + str(ambari_port)
#### Prompt for password & build the HTTP session
ambari_pass = getpass.getpass()
s = requests.Session()
s.auth = (ambari_user, ambari_pass)
s.headers.update({'X-Requested-By':'seanorama'})
#### Authenticate & verify authentication
r = s.get(api_url + '/api/v1/clusters')
assert r.status_code == 200
print("You are authenticated to Ambari!")
# -
# ### API Examples: Users
# +
#### Change password of admin user
old_pass = getpass.getpass()
new_pass = getpass.getpass()
body = {
"Users": {
"user_name": "admin",
"old_password": <PASSWORD>,
"password": <PASSWORD>
}}
r = s.put(api_url + '/api/v1/users/' + ambari_user, data=json.dumps(body))
print(r.url)
assert r.status_code == 200
print("Password changed successfully!")
# +
#### Add a user
new_pass = getpass.getpass()
body = {
"Users/user_name": "sean",
"Users/password": <PASSWORD>,
"Users/active": 'true',
"Users/admin": 'false'
}
r = s.post(api_url + '/api/v1/users', data=json.dumps(body))
print(r.url)
assert r.status_code == 201
print("User created successfully!")
# +
### Grant privileges
#### allowed permission_names: CLUSTER.OPERATE, CLUSTER.READ
body = [{
"PrivilegeInfo": {
"permission_name":"CLUSTER.READ",
"principal_name":"sean",
"principal_type":"USER"
}}]
r = s.post(api_url + '/api/v1/clusters/Sandbox/privileges', data=json.dumps(body))
print(r.url)
assert r.status_code == 201
print("Privilege granted!")
# -
# ### API Examples: Clusters & Hosts
# +
### List Clusters
r = s.get(api_url + '/api/v1/clusters')
print(r.url)
print(json.dumps(r.json(), indent=2))
# +
### Set cluster based on existing cluster
cluster = r.json()['items'][0]['Clusters']['cluster_name']
cluster
# +
#### List registered hosts
r = s.get(api_url + '/api/v1/hosts')
print(r.url)
#print(json.dumps(r.json(), indent=2))
for host in [item["Hosts"]["host_name"] for item in r.json()["items"]]:
print(host)
# +
### Cluster details
r = s.get(api_url + '/api/v1/clusters/' + cluster)
print(r.url)
print(json.dumps(r.json(), indent=2))
# +
#### List hosts in cluster
r = s.get(api_url + '/api/v1/clusters/' + cluster + '/hosts')
print(r.url)
print(json.dumps(r.json(), indent=2))
# -
# ### API Examples: Ambari Stacks
# +
#### List Stacks
r = s.get(api_url + '/api/v1/stacks')
print(json.dumps(r.json(), indent=2))
# +
#### List Stack services
r = s.get(api_url + '/api/v1/stacks/HDP/versions/2.2')
#print(json.dumps(r.json(), indent=2))
print("\nServices which Ambari 1.7.0 can deploy for HDP 2.2:")
#for services in r.json()['services']:
# print('\t',services['StackServices']['service_name'])
for service in [service['StackServices']['service_name'] for service in r.json()['services']]:
print('\t',service)
# -
# ### API Examples: Ambari Blueprints
# +
#### List Blueprints
r = s.get(api_url + '/api/v1/blueprints')
print(r.url)
print(json.dumps(r.json(), indent=2))
# +
#### Blueprint: Load from file
blueprint = json.loads(open('blueprint_hdp22-single-node-simple.json').read())
# +
#### Blueprint: Upload to Ambari
body = blueprint
r = s.post(api_url + '/api/v1/blueprints/myblueprint', data=json.dumps(body))
assert r.status_code == 201
print("Blueprint uploaded successfully!")
# +
#### Show Blueprint
r = s.get(api_url + '/api/v1/blueprints/myblueprint')
print(r.url)
print(json.dumps(r.json(), indent=2))
# +
#### Create Cluster from Blueprint
body = {
"blueprint": "myblueprint",
"default_password": "<PASSWORD>",
"configurations": [
{ "hive-site": { "properties": { "javax.jdo.option.ConnectionPassword": "<PASSWORD>" } } }
],
"host_groups": [
{
"hosts": [ { "fqdn": "hdp" } ],
"name": "host-group-1"
}]}
r = s.post(api_url + '/api/v1/clusters/mycluster', data=json.dumps(body))
print(r.url)
print(r.status_code)
print(json.dumps(r.json(), indent=2))
status_url = r.json()['href']
# +
#### Check Cluster creation status
r = s.get(status_url)
print(r.url)
print(json.dumps(r.json()['Requests'], indent=2))
# +
#### Blueprint: Export Blueprint from existing Cluster
r = s.get(api_url + '/api/v1/clusters/' + cluster + '?format=blueprint')
print(r.url)
print(json.dumps(r.json(), indent=2))
# +
#### Blueprint: Export Blueprint from existing Cluster
r = s.get(api_url + '/api/v1/clusters/' + cluster + '?format=blueprint')
print(r.url)
#print(json.dumps(r.json(), indent=2))
print(r.json()['configurations']['hdfs-site'])
# -
# ### API Examples: Ambari Views
# +
#### List Views
r = s.get(api_url + '/api/v1/views')
#print(json.dumps(r.json(), indent=2))
for view in [item['ViewInfo']['view_name'] for item in r.json()['items']]:
print(view)
# -
# ### API Examples: Change Configuration
#
# As part of the Stinger project, Tez brings many performance improvements to Hive. But as of HDP 2.2.0 they are not turned on my default.
#
# The below will make the required changes using the API.
#
# See the blog for more details: http://hortonworks.com/hadoop-tutorial/supercharging-interactive-queries-hive-tez/
#
# #### Process to change configuration from API:
#
# 1. Get current configuration tag
# 2. Get current configuration as JSON
# 3. Update the configuration JSON with your changes
# 4. Modify the configuration JSON to Ambari's required format, including setting new tag
# 5. Submit the new configuration JSON to Ambari
# 6. Restart services
#
# +
#### Get current configuration tag
r = s.get(api_url + '/api/v1/clusters/' + cluster + '?fields=Clusters/desired_configs/hive-site')
print(r.url)
print(json.dumps(r.json(), indent=2))
tag = r.json()['Clusters']['desired_configs']['hive-site']['tag']
# +
### Get current configuration
r = s.get(api_url + '/api/v1/clusters/' + cluster + '/configurations?type=hive-site&tag=' + tag)
print(r.url)
print(json.dumps(r.json(), indent=2))
# +
#### Change the configuration
config_old = r.json()['items'][0]
config_new = r.json()['items'][0]
#### The configurations you want to change
config_new['properties']['hive.execution.engine'] = 'tez'
# +
#### Show the differences
a = json.dumps(config_old, indent=2).splitlines(1)
b = json.dumps(config_new, indent=2).splitlines(1)
for line in difflib.unified_diff(a, b):
sys.stdout.write(line)
# +
#### Manipulate the document to match the format Ambari expects
#### Adds new configuration tag, deletes fields, and wraps in appropriate json
config_new['tag'] = 'version' + str(int(round(time.time() * 1000000000)))
del config_new['Config']
del config_new['href']
del config_new['version']
config_new = {"Clusters": {"desired_config": config_new}}
print(json.dumps(config_new, indent=2))
# +
body = config_new
r = s.put(api_url + '/api/v1/clusters/' + cluster, data=json.dumps(body))
print(r.url)
print(r.status_code)
assert r.status_code == 200
print("Configuration changed successfully!")
print(json.dumps(r.json(), indent=2))
# -
# #### What you'll see from the Ambari UI:
#
# 
# ### API Examples: Managing Services & Components
# +
#### Create Cluster from Blueprint
service = 'GANGLIA'
body = {
"ServiceInfo": {
"state" : "INSTALLED”
}
}
r = s.post(api_url + '/api/v1/clusters/ + cluster + 'services/' + service , data=json.dumps(body))
print(r.url)
print(r.status_code)
print(json.dumps(r.json(), indent=2))
status_url = r.json()['href']
# +
PUT /clusters/c1/services/HDFS
{
"ServiceInfo": {
"state" : "INSTALLED”
}
}
PUT /api/v1/clusters/c1/services?ServiceInfo/state=INSTALLED
{
"ServiceInfo": {
"state" : "STARTED”
}
}
|
build/security/ambari-bootstrap-master/api-examples/ambari-api-examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
# Numpy import for array processing, python doesn’t have built in array support. The feature of working with native arrays can be used in python with the help of numpy library.
#
# Pandas is a library of python used for working with tables, on importing the data, mostly data will be of table format, for ease manipulation of tables pandas library is imported
#
# Matplotlib is a library of python used to plot graphs, for the purpose of visualizing the results we would be plotting the results with the help of matplotlib library.
#
# Math import is just used to square the numerical values
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math
# # Reading the dataset from data
# In this line of code using the read_excel method of pandas library, the dataset has been imported from data folder and stored in dataset variable.
dataset = pd.read_csv(r'..\\data\\auto_insurance.csv')
# On viewing the dataset, it contains of two columns X and Y where X is dependent variable and Y is Independent Variable.
dataset.head()
# # Creating Dependent and Independent variables
# The X Column from the dataset is extracted into an X variable of type numpy, similarly the y variable
# X is an independent variable
# Y is dependent variable Inference
X = dataset['X'].values
y = dataset['Y'].values
# On execution of first line would result in a pandas Series Object
# On using values attribute it would result in an numpy array
print(type(dataset['X']))
print(type(dataset['X'].values))
# # Visualizing the data
# The step is to just see how the dataset is
# On visualization the data would appear something like this
# The X and Y attributes would vary based on dataset.
# Each point on the plot is a data point showing the respective Number of Claims on x-axis and Total Payment on y-axis
title='Linear Regression on Auto Insurance in Sweden Dataset'
x_axis_label = 'Number of Claims'
y_axis_label = 'Total Payment'
plt.scatter(X,y)
plt.title(title)
plt.xlabel(x_axis_label)
plt.ylabel(y_axis_label)
plt.show()
# # Splitting the data into training set and test set
# We are splitting the whole dataset into training and test set where training set is used for fitting the line to data and test set is used to check how good the line if for the data.
# >This splitting can be dont with scikit learns test train split or manully by below code
X_train,X_test = np.split(X,indices_or_sections = [int(len(X)*0.8)])
y_train,y_test = np.split(y,indices_or_sections = [int(len(X)*0.8)])
# # Reshaping the numpy arrays since the scikit learn model expects 2-D array in further code
# In further the scikit learn model would be expecting a 2-D array of shape (length,1).
X_train_fw = np.reshape(X_train,newshape = (-1,1))
y_train_fw = np.reshape(y_train,newshape = (-1,1))
X_test_fw = np.reshape(X_test,newshape = (-1,1))
y_test_fw = np.reshape(y_test,newshape = (-1,1))
# The code was just to convert a single dimensional array into a 2-D array where each element is an array.
print('Before Reshaping',np.shape(X))
print('After Reshaping',np.shape(X_train))
# # Computing the values of sigma
# As per the derivation formula we are computing the values of sigma x sigme x^2 sigma y simg x*y
# n is the number of terms in the dataset
sigma_X = sum(X_train)
sigma_y = sum(y_train)
sigma_xy = sum(np.multiply(X_train,y_train))
sigma_X_square = sum(np.square(X_train))
n = len(X_train)
# # Computing the values of slope and intercept
# As our linear regression line requires a slope and intercept
# we are computing their values using statistical formulas
# +
m_numerator = (n*sigma_xy)-(sigma_X*sigma_y)
m_denominator = n*sigma_X_square - math.pow(sigma_X,2)
m = m_numerator/m_denominator
c_numerator = (sigma_y*sigma_X_square)-(sigma_xy*sigma_X)
c_denominator = (n*sigma_X_square) - math.pow(sigma_X,2)
c = c_numerator/c_denominator
# -
# # Importing the linear model from sklearn framework
# From scikit learn Library LinearRegression is imported. Lr is an object of LinearRegression.
# The process of training is done in the fit method, our dependent and independent variable are fed into to the fit method in which it would try to fit a line to the data provided.
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X = X_train_fw, y = y_train_fw)
# # Predicting the Results
# `Line 1` By the knowing the slope and intercept values of linear regression model we are trying to predict the values of test data. Y_pred variable contains all the predicted y-values of the test x-values.
#
# `Line 2` By the trained linear regression model we are trying to predict the values of test data. Y_pred variable contains all the predicted y-values of the test x-values.
y_pred_stat = X_test*m + c
y_pred_fw = lr.predict(X_test_fw)
# # Visualizing the Results
# As we have predicted the y-values for a set of x-values we are visualizing the results to check how good did our line fit for our predictions.
# # Plotting each result individually
# The plot shows the blue points are the data points are actual values where the cyan and Green lines is the predictions
plt.scatter(X_test,y_test,c='red')
plt.plot(X_test,y_pred_fw,c='cyan',label='framework')
plt.scatter(X,y)
plt.title(title)
plt.xlabel(x_axis_label)
plt.ylabel(y_axis_label)
plt.show()
plt.scatter(X_test,y_test,c='red')
plt.plot(X_test,y_pred_stat,c='green',label='statistical formula')
plt.scatter(X,y)
plt.title(title)
plt.legend()
plt.xlabel(x_axis_label)
plt.ylabel(y_axis_label)
plt.show()
# # Combining the results
# The plot shows the blue points are the data points are actual values where the lines is the predictions
# > Note: In the below graph one is was hidden over the other
plt.scatter(X_test,y_test,c='red')
plt.plot(X_test,y_pred_fw,c='cyan',label='framework')
plt.plot(X_test,y_pred_stat,c='green',label='statistical formula')
plt.scatter(X,y)
plt.title(title)
plt.legend()
plt.xlabel(x_axis_label)
plt.ylabel(y_axis_label)
plt.show()
|
src/linear_regression_comparison_framework_vs_scratch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import numpy as np
import cv2
from matplotlib import pyplot as plt
from sklearn.model_selection import train_test_split
from keras.utils import np_utils
from keras import utils, losses, optimizers
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation, Flatten, Lambda
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.layers import Conv2D, MaxPooling2D
SEED = 2017
# +
# Specify data directory and extract all file names for both classes
DATA_DIR = 'Data/PetImages/'
cats = glob.glob(DATA_DIR + "Cat/*.jpg")
dogs = glob.glob(DATA_DIR + "Dog/*.jpg")
print('#Cats: {}, #Dogs: {}'.format(len(cats), len(dogs)))
# #Cats: 12500, #Dogs: 12500
# -
dogs_train, dogs_val, cats_train, cats_val = train_test_split(dogs, cats, test_size=0.2, random_state=SEED)
def batchgen(cats, dogs, batch_size, img_size=50):
# Create empty numpy arrays
batch_images = np.zeros((batch_size, img_size, img_size, 3))
batch_label = np.zeros(batch_size)
# Custom batch generator
while 1:
n = 0
while n < batch_size:
# Randomly pick a dog or cat image
if np.random.randint(2) == 1:
i = np.random.randint(len(dogs))
img = cv2.imread(dogs[i])
if img is None:
break
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# The images have different dimensions, we resize all to 100x100
img = cv2.resize(img, (img_size, img_size), interpolation = cv2.INTER_AREA)
y = 1
else:
i = np.random.randint(len(cats))
img = cv2.imread(cats[i])
if img is None:
break
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = cv2.resize(img, (img_size, img_size), interpolation = cv2.INTER_AREA)
y = 0
batch_images[n] = img
batch_label[n] = y
n+=1
yield batch_images, batch_label
def create_model(init_type='xavier', img_size=100):
# Define architecture
model = Sequential()
model.add(Lambda(lambda x: (x / 255.) - 0.5, input_shape=(img_size, img_size, 3)))
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer=init_type))
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer=init_type))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.5))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer=init_type))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer=init_type))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
sgd = optimizers.Adam()
model.compile(loss='binary_crossentropy', optimizer=sgd, metrics=['binary_accuracy'])
return model
models = []
for init_type in ['random_uniform', 'glorot_normal', 'glorot_uniform', 'lecun_uniform', 'he_uniform']:
model = create_model(init_type, img_size=50)
models.append(dict({'setting': '{}'.format(init_type),
'model': model
}))
callbacks = [EarlyStopping(monitor='val_binary_accuracy', patience=3)]
# +
batch_size = 512
n_epochs = 500
steps_per_epoch = 100
validation_steps = round((len(dogs_val)+len(cats_val))/batch_size)
train_generator = batchgen(dogs_train, cats_train, batch_size)
val_generator = batchgen(dogs_val, cats_val, batch_size)
history = []
for i in range(len(models)):
print(models[i])
history.append(
models[i]['model'].
fit_generator(train_generator, steps_per_epoch=steps_per_epoch, epochs=n_epochs, validation_data=val_generator, validation_steps=validation_steps, callbacks=callbacks)
)
# -
for i in range(len(models)):
plt.plot(range(len(history[i].history['val_binary_accuracy'])), history[i].history['val_binary_accuracy'], label=models[i]['setting'])
print('Max accuracy model {}: {}'.format(models[i]['setting'], max(history[i].history['val_binary_accuracy'])))
plt.title('Accuracy on the validation set')
plt.xlabel('epochs')
plt.ylabel('accuracy')
plt.legend()
plt.show()
|
Chapter03/Chapter 3 - Experimenting with different types of initialization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 数据操作
# :label:`sec_ndarray`
#
# 为了能够完成各种数据操作,我们需要某种方法来存储和操作数据。
# 通常,我们需要做两件重要的事:(1)获取数据;(2)将数据读入计算机后对其进行处理。
# 如果没有某种方法来存储数据,那么获取数据是没有意义的。
#
# 首先,我们介绍$n$维数组,也称为*张量*(tensor)。
# 使用过Python中NumPy计算包的读者会对本部分很熟悉。
# 无论使用哪个深度学习框架,它的*张量类*(在MXNet中为`ndarray`,
# 在PyTorch和TensorFlow中为`Tensor`)都与Numpy的`ndarray`类似。
# 但深度学习框架又比Numpy的`ndarray`多一些重要功能:
# 首先,GPU很好地支持加速计算,而NumPy仅支持CPU计算;
# 其次,张量类支持自动微分。
# 这些功能使得张量类更适合深度学习。
# 如果没有特殊说明,本书中所说的张量均指的是张量类的实例。
#
# ## 入门
#
# 本节的目标是帮助读者了解并运行一些在阅读本书的过程中会用到的基本数值计算工具。
# 如果你很难理解一些数学概念或库函数,请不要担心。
# 后面的章节将通过一些实际的例子来回顾这些内容。
# 如果你已经具有相关经验,想要深入学习数学内容,可以跳过本节。
#
# + [markdown] origin_pos=3 tab=["tensorflow"]
# 首先,我们导入`tensorflow`。
# 由于`tensorflow`名称有点长,我们经常在导入它后使用短别名`tf`。
#
# + origin_pos=6 tab=["tensorflow"]
import tensorflow as tf
# + [markdown] origin_pos=7
# [**张量表示由一个数值组成的数组,这个数组可能有多个维度**]。
# 具有一个轴的张量对应数学上的*向量*(vector);
# 具有两个轴的张量对应数学上的*矩阵*(matrix);
# 具有两个轴以上的张量没有特殊的数学名称。
#
# 首先,可以使用`arange`创建一个行向量`x`。
# 这个行向量包含从0开始的前12个整数,它们被默认创建为浮点数。
# 张量中的每个值都称为张量的*元素*(element)。
# 例如,张量`x`中有12个元素。
# 除非额外指定,新的张量默认将存储在内存中,并采用基于CPU的计算。
#
# + origin_pos=10 tab=["tensorflow"]
x = tf.range(12)
x
# + [markdown] origin_pos=11
# [**可以通过张量的`shape`属性来访问张量(沿每个轴的长度)的*形状***]
# (~~和张量中元素的总数~~)。
#
# + origin_pos=12 tab=["tensorflow"]
x.shape
# + [markdown] origin_pos=13
# 如果只想知道张量中元素的总数,即形状的所有元素乘积,可以检查它的大小(size)。
# 因为这里在处理的是一个向量,所以它的`shape`与它的`size`相同。
#
# + origin_pos=16 tab=["tensorflow"]
tf.size(x)
# + [markdown] origin_pos=17
# [**要想改变一个张量的形状而不改变元素数量和元素值,可以调用`reshape`函数。**]
# 例如,可以把张量`x`从形状为(12,)的行向量转换为形状为(3,4)的矩阵。
# 这个新的张量包含与转换前相同的值,但是它被看成一个3行4列的矩阵。
# 要重点说明一下,虽然张量的形状发生了改变,但其元素值并没有变。
# 注意,通过改变张量的形状,张量的大小不会改变。
#
# + origin_pos=19 tab=["tensorflow"]
X = tf.reshape(x, (3, 4))
X
# + [markdown] origin_pos=20
# 我们不需要通过手动指定每个维度来改变形状。
# 也就是说,如果我们的目标形状是(高度,宽度),
# 那么在知道宽度后,高度会被自动计算得出,不必我们自己做除法。
# 在上面的例子中,为了获得一个3行的矩阵,我们手动指定了它有3行和4列。
# 幸运的是,我们可以通过`-1`来调用此自动计算出维度的功能。
# 即我们可以用`x.reshape(-1,4)`或`x.reshape(3,-1)`来取代`x.reshape(3,4)`。
#
# 有时,我们希望[**使用全0、全1、其他常量,或者从特定分布中随机采样的数字**]来初始化矩阵。
# 我们可以创建一个形状为(2,3,4)的张量,其中所有元素都设置为0。代码如下:
#
# + origin_pos=23 tab=["tensorflow"]
tf.zeros((2, 3, 4))
# + [markdown] origin_pos=24
# 同样,我们可以创建一个形状为`(2,3,4)`的张量,其中所有元素都设置为1。代码如下:
#
# + origin_pos=27 tab=["tensorflow"]
tf.ones((2, 3, 4))
# + [markdown] origin_pos=28
# 有时我们想通过从某个特定的概率分布中随机采样来得到张量中每个元素的值。
# 例如,当我们构造数组来作为神经网络中的参数时,我们通常会随机初始化参数的值。
# 以下代码创建一个形状为(3,4)的张量。
# 其中的每个元素都从均值为0、标准差为1的标准高斯分布(正态分布)中随机采样。
#
# + origin_pos=31 tab=["tensorflow"]
tf.random.normal(shape=[3, 4])
# + [markdown] origin_pos=32
# 我们还可以[**通过提供包含数值的Python列表(或嵌套列表),来为所需张量中的每个元素赋予确定值**]。
# 在这里,最外层的列表对应于轴0,内层的列表对应于轴1。
#
# + origin_pos=35 tab=["tensorflow"]
tf.constant([[2, 1, 4, 3], [1, 2, 3, 4], [4, 3, 2, 1]])
# + [markdown] origin_pos=36
# ## 运算符
#
# 我们的兴趣不仅限于读取数据和写入数据。
# 我们想在这些数据上执行数学运算,其中最简单且最有用的操作是*按元素*(elementwise)运算。
# 它们将标准标量运算符应用于数组的每个元素。
# 对于将两个数组作为输入的函数,按元素运算将二元运算符应用于两个数组中的每对位置对应的元素。
# 我们可以基于任何从标量到标量的函数来创建按元素函数。
#
# 在数学表示法中,我们将通过符号$f: \mathbb{R} \rightarrow \mathbb{R}$
# 来表示*一元*标量运算符(只接收一个输入)。
# 这意味着该函数从任何实数($\mathbb{R}$)映射到另一个实数。
# 同样,我们通过符号$f: \mathbb{R}, \mathbb{R} \rightarrow \mathbb{R}$
# 表示*二元*标量运算符,这意味着该函数接收两个输入,并产生一个输出。
# 给定同一形状的任意两个向量$\mathbf{u}$和$\mathbf{v}$和二元运算符$f$,
# 我们可以得到向量$\mathbf{c} = F(\mathbf{u},\mathbf{v})$。
# 具体计算方法是$c_i \gets f(u_i, v_i)$,
# 其中$c_i$、$u_i$和$v_i$分别是向量$\mathbf{c}$、$\mathbf{u}$和$\mathbf{v}$中的元素。
# 在这里,我们通过将标量函数升级为按元素向量运算来生成向量值
# $F: \mathbb{R}^d, \mathbb{R}^d \rightarrow \mathbb{R}^d$。
#
# 对于任意具有相同形状的张量,
# [**常见的标准算术运算符(`+`、`-`、`*`、`/`和`**`)都可以被升级为按元素运算**]。
# 我们可以在同一形状的任意两个张量上调用按元素操作。
# 在下面的例子中,我们使用逗号来表示一个具有5个元素的元组,其中每个元素都是按元素操作的结果。
#
# + origin_pos=39 tab=["tensorflow"]
x = tf.constant([1.0, 2, 4, 8])
y = tf.constant([2.0, 2, 2, 2])
x + y, x - y, x * y, x / y, x ** y # **运算符是求幂运算
# + [markdown] origin_pos=40
# (**“按元素”方式可以应用更多的计算**),包括像求幂这样的一元运算符。
#
# + origin_pos=43 tab=["tensorflow"]
tf.exp(x)
# + [markdown] origin_pos=44
# 除了按元素计算外,我们还可以执行线性代数运算,包括向量点积和矩阵乘法。
# 我们将在 :numref:`sec_linear-algebra`中解释线性代数的重点内容。
#
# [**我们也可以把多个张量*连结*(concatenate)在一起**],
# 把它们端对端地叠起来形成一个更大的张量。
# 我们只需要提供张量列表,并给出沿哪个轴连结。
# 下面的例子分别演示了当我们沿行(轴-0,形状的第一个元素)
# 和按列(轴-1,形状的第二个元素)连结两个矩阵时,会发生什么情况。
# 我们可以看到,第一个输出张量的轴-0长度($6$)是两个输入张量轴-0长度的总和($3 + 3$);
# 第二个输出张量的轴-1长度($8$)是两个输入张量轴-1长度的总和($4 + 4$)。
#
# + origin_pos=47 tab=["tensorflow"]
X = tf.reshape(tf.range(12, dtype=tf.float32), (3, 4))
Y = tf.constant([[2.0, 1, 4, 3], [1, 2, 3, 4], [4, 3, 2, 1]])
tf.concat([X, Y], axis=0), tf.concat([X, Y], axis=1)
# + [markdown] origin_pos=48
# 有时,我们想[**通过*逻辑运算符*构建二元张量**]。
# 以`X == Y`为例:
# 对于每个位置,如果`X`和`Y`在该位置相等,则新张量中相应项的值为1。
# 这意味着逻辑语句`X == Y`在该位置处为真,否则该位置为0。
#
# + origin_pos=49 tab=["tensorflow"]
X == Y
# + [markdown] origin_pos=50
# [**对张量中的所有元素进行求和,会产生一个单元素张量。**]
#
# + origin_pos=52 tab=["tensorflow"]
tf.reduce_sum(X)
# + [markdown] origin_pos=53
# ## 广播机制
# :label:`subsec_broadcasting`
#
# 在上面的部分中,我们看到了如何在相同形状的两个张量上执行按元素操作。
# 在某些情况下,[**即使形状不同,我们仍然可以通过调用
# *广播机制*(broadcasting mechanism)来执行按元素操作**]。
# 这种机制的工作方式如下:首先,通过适当复制元素来扩展一个或两个数组,
# 以便在转换之后,两个张量具有相同的形状。
# 其次,对生成的数组执行按元素操作。
#
# 在大多数情况下,我们将沿着数组中长度为1的轴进行广播,如下例子:
#
# + origin_pos=56 tab=["tensorflow"]
a = tf.reshape(tf.range(3), (3, 1))
b = tf.reshape(tf.range(2), (1, 2))
a, b
# + [markdown] origin_pos=57
# 由于`a`和`b`分别是$3\times1$和$1\times2$矩阵,如果让它们相加,它们的形状不匹配。
# 我们将两个矩阵*广播*为一个更大的$3\times2$矩阵,如下所示:矩阵`a`将复制列,
# 矩阵`b`将复制行,然后再按元素相加。
#
# + origin_pos=58 tab=["tensorflow"]
a + b
# + [markdown] origin_pos=59
# ## 索引和切片
#
# 就像在任何其他Python数组中一样,张量中的元素可以通过索引访问。
# 与任何Python数组一样:第一个元素的索引是0,最后一个元素索引是-1;
# 可以指定范围以包含第一个元素和最后一个之前的元素。
#
# 如下所示,我们[**可以用`[-1]`选择最后一个元素,可以用`[1:3]`选择第二个和第三个元素**]:
#
# + origin_pos=60 tab=["tensorflow"]
X[-1], X[1:3]
# + [markdown] origin_pos=62 tab=["tensorflow"]
# TensorFlow中的`Tensors`是不可变的,也不能被赋值。
# TensorFlow中的`Variables`是支持赋值的可变容器。
# 请记住,TensorFlow中的梯度不会通过`Variable`反向传播。
#
# 除了为整个`Variable`分配一个值之外,我们还可以通过索引来写入`Variable`的元素。
#
# + origin_pos=64 tab=["tensorflow"]
X_var = tf.Variable(X)
X_var[1, 2].assign(9)
X_var
# + [markdown] origin_pos=65
# 如果我们想[**为多个元素赋值相同的值,我们只需要索引所有元素,然后为它们赋值。**]
# 例如,`[0:2, :]`访问第1行和第2行,其中“:”代表沿轴1(列)的所有元素。
# 虽然我们讨论的是矩阵的索引,但这也适用于向量和超过2个维度的张量。
#
# + origin_pos=67 tab=["tensorflow"]
X_var = tf.Variable(X)
X_var[0:2, :].assign(tf.ones(X_var[0:2,:].shape, dtype = tf.float32) * 12)
X_var
# + [markdown] origin_pos=68
# ## 节省内存
#
# [**运行一些操作可能会导致为新结果分配内存**]。
# 例如,如果我们用`Y = X + Y`,我们将取消引用`Y`指向的张量,而是指向新分配的内存处的张量。
#
# 在下面的例子中,我们用Python的`id()`函数演示了这一点,
# 它给我们提供了内存中引用对象的确切地址。
# 运行`Y = Y + X`后,我们会发现`id(Y)`指向另一个位置。
# 这是因为Python首先计算`Y + X`,为结果分配新的内存,然后使`Y`指向内存中的这个新位置。
#
# + origin_pos=69 tab=["tensorflow"]
before = id(Y)
Y = Y + X
id(Y) == before
# + [markdown] origin_pos=70
# 这可能是不可取的,原因有两个:首先,我们不想总是不必要地分配内存。
# 在机器学习中,我们可能有数百兆的参数,并且在一秒内多次更新所有参数。
# 通常情况下,我们希望原地执行这些更新。
# 其次,如果我们不原地更新,其他引用仍然会指向旧的内存位置,
# 这样我们的某些代码可能会无意中引用旧的参数。
#
# + [markdown] origin_pos=72 tab=["tensorflow"]
# `Variables`是TensorFlow中的可变容器,它们提供了一种存储模型参数的方法。
# 我们可以通过`assign`将一个操作的结果分配给一个`Variable`。
# 为了说明这一点,我们创建了一个与另一个张量`Y`相同的形状的`Z`,
# 使用`zeros_like`来分配一个全$0$的块。
#
# + origin_pos=75 tab=["tensorflow"]
Z = tf.Variable(tf.zeros_like(Y))
print('id(Z):', id(Z))
Z.assign(X + Y)
print('id(Z):', id(Z))
# + [markdown] origin_pos=77 tab=["tensorflow"]
# 即使你将状态持久存储在`Variable`中,
# 你也可能希望避免为不是模型参数的张量过度分配内存,从而进一步减少内存使用量。
#
# 由于TensorFlow的`Tensors`是不可变的,而且梯度不会通过`Variable`流动,
# 因此TensorFlow没有提供一种明确的方式来原地运行单个操作。
#
# 但是,TensorFlow提供了`tf.function`修饰符,
# 将计算封装在TensorFlow图中,该图在运行前经过编译和优化。
# 这允许TensorFlow删除未使用的值,并复用先前分配的且不再需要的值。
# 这样可以最大限度地减少TensorFlow计算的内存开销。
#
# + origin_pos=79 tab=["tensorflow"]
@tf.function
def computation(X, Y):
Z = tf.zeros_like(Y) # 这个未使用的值将被删除
A = X + Y # 当不再需要时,分配将被复用
B = A + Y
C = B + Y
return C + Y
computation(X, Y)
# + [markdown] origin_pos=80
# ## 转换为其他Python对象
#
# + [markdown] origin_pos=81 tab=["tensorflow"]
# 将深度学习框架定义的张量[**转换为NumPy张量(`ndarray`)**]很容易,反之也同样容易。
# 转换后的结果不共享内存。
# 这个小的不便实际上是非常重要的:当你在CPU或GPU上执行操作的时候,
# 如果Python的NumPy包也希望使用相同的内存块执行其他操作,你不希望停下计算来等它。
#
# + origin_pos=85 tab=["tensorflow"]
A = X.numpy()
B = tf.constant(A)
type(A), type(B)
# + [markdown] origin_pos=86
# 要(**将大小为1的张量转换为Python标量**),我们可以调用`item`函数或Python的内置函数。
#
# + origin_pos=89 tab=["tensorflow"]
a = tf.constant([3.5]).numpy()
a, a.item(), float(a), int(a)
# + [markdown] origin_pos=90
# ## 小结
#
# * 深度学习存储和操作数据的主要接口是张量($n$维数组)。它提供了各种功能,包括基本数学运算、广播、索引、切片、内存节省和转换其他Python对象。
#
# ## 练习
#
# 1. 运行本节中的代码。将本节中的条件语句`X == Y`更改为`X < Y`或`X > Y`,然后看看你可以得到什么样的张量。
# 1. 用其他形状(例如三维张量)替换广播机制中按元素操作的两个张量。结果是否与预期相同?
#
# + [markdown] origin_pos=93 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/1746)
#
|
d2l/tensorflow/chapter_preliminaries/ndarray.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ROI Pooling Demo
#
# The ROI pooling layer is a specialized version of the max pooling layer commonly used in convolutional neural networks. A max pooling layer is typically used to spatially downsample a 2d feature map by choosing the maximum value from a small region corresponding to a particular output pixel. The ROI pooling layer does the same thing, but also takes as input a set of ROIs (regions of interest) over which to pool. This is essentially equivalent to max pooling the input image croppped to each ROI, and computing the stride of the pooling such that each output feature map has the same size no matter the ROI size.
#
# The ROI pooling layer is used in algorithms such as Fast-RCNN, which use it to pool over the output of the last convolutional layer of a CNN for a number of input ROIs. This allows the inspection of various regions of an image, but without having to run all the convoulational layers on each individual crop of the image.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import PIL
from PIL import Image
import tensorflow as tf
from tensorflow.python.ops import array_ops
from tensorflow.python.framework import ops
import os
home = os.getenv("HOME")
# +
# Since we've added custom operations, we need to import them. Tensorflow does not automatically add custom ops.
# Adjust the paths below to your tensorflow source folder.
# Import the forward op
roi_pooling_module = tf.load_op_library(
home + "/packages/tensorflow/bazel-bin/tensorflow/core/user_ops/roi_pooling_op.so")
roi_pooling_op = roi_pooling_module.roi_pooling
# Import the gradient op
roi_pooling_module_grad = tf.load_op_library(
home + "/packages/tensorflow/bazel-bin/tensorflow/core/user_ops/roi_pooling_op_grad.so")
roi_pooling_op_grad = roi_pooling_module_grad.roi_pooling_grad
# -
# Open a demo image
im = Image.open("cat.png").convert("L")
# Show the image
im
# Image dimensions - HW ordering
np.asarray(im).shape
# +
# Prepare demo inputs
num_batches = 1
# Test ROIs
rois = [(0, 0, 300, 400), (150, 150, 50, 200)]
# Data ordering: num batches, num ROIs, (x, y, height, width)
roi_array = np.asarray(rois).astype(int).reshape(len(rois), 4)
roi_array = np.asarray([roi_array for i in range(num_batches)])
# Input image
# Crop dimensions (for easy testing of various input sizes)
image_width = 403
image_height = 304
input_image = Image.open("cat.png").convert("L").crop((0, 0, image_width, image_height))
# Should be in NCHW format (num batches, num channels, height, width)
input_array = np.asarray(input_image).reshape(1, image_height, image_width)
input_array = np.asarray([input_array for i in range(num_batches)])
# Output should be a tensor of this shape
# height, width
output_shape = np.asarray((50, 80)).astype(np.int32)
# Prepare an array of random numbers to test gradient backpropagation
# Should be same size as output of ROI pooling layer
grad_test = np.random.random((1, 1, len(roi_array), output_shape[0], output_shape[1]))
# -
# Here we register our gradient op as the gradient function for our ROI pooling op.
@ops.RegisterGradient("RoiPooling")
def _roi_pooling_grad(op, grad0, grad1):
# The input gradients are the gradients with respect to the outputs of the pooling layer
input_grad = grad0
# We need the argmax data to compute the gradient connections
argmax = op.outputs[1]
# Grab the shape of the inputs to the ROI pooling layer
input_shape = array_ops.shape(op.inputs[0])
# Compute the gradient
backprop_grad = roi_pooling_op_grad(input_grad, argmax, input_shape)
# Return the gradient for the feature map, but not for the other inputs
return [backprop_grad, None, None]
# +
# Set up the TensorFlow operations to test
# Placeholders are placeholders for the input data. Passed in for each computation.
data = tf.placeholder(tf.float32)
rois = tf.placeholder(tf.int32)
grad = tf.placeholder(tf.float32)
output_shape_tensor = tf.placeholder(tf.int32)
# Get the shape of the input feature map
input_shape = array_ops.shape(data)
# We can use either the CPU or GPU. Here we'll use the gpu. Other examples test both implementations.
with tf.device("/gpu:0"):
# Compute the forward pass - returns the feature map and the argmax data for backprop
result, argmax = roi_pooling_op(data, rois, output_shape_tensor)
# Compute the backwards pass - the gradient between result and data.
# Calls the registered gradient operation
gradient, = tf.gradients(result, data)
# -
# Set up a Tensorflow session
sess = tf.InteractiveSession()
# +
# Run the forward pass. We ask for result and argmax to be computed, and pass
# in a feed dictionary to fill in the placeholders.
# result_out is the computed pooled feature map.
# argmax_out contains the location the max pixels were found
result_out, argmax_out = \
sess.run([result, argmax], \
feed_dict={data:input_array, rois:roi_array, grad:grad_test, output_shape_tensor:output_shape})
# -
# Run the backward pass.
gradient_out, = \
sess.run([gradient], \
feed_dict={data:input_array, rois:roi_array, grad:grad_test, output_shape_tensor:output_shape})
# Display the result of pooling the first ROI
plt.imshow(result_out[0, 0, 0, :, :].reshape(output_shape))
# Display the result of pooling the second ROI
plt.imshow(result_out[0, 0, 1, :, :].reshape(output_shape))
# Print a sample of argmax_out. Contains the locations that the max pooling
# value for each output pixel was found. For example, pixel (0, 0) in the first
# ROI pooling output was found at index 3 (indices in the flattened version of the
# input feature map).
argmax_out
# Shows the first 100 elements of the back-propagated gradient.
# The gradient computation connects all pixels from the input that were chosen
# during max pooling to the upstream gradient. Thus, all pixels that were not
# chosen will have a computed gradient of 0.
gradient_out.flatten()[:100]
|
tensorflow/examples/roi_pooling/demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Building your own container as Algorithm / Model Package
#
# With Amazon SageMaker, you can package your own algorithms that can than be trained and deployed in the SageMaker environment. This notebook will guide you through an example that shows you how to build a Docker container for SageMaker and use it for training and inference.
#
# This is an extension of the [scikit-bring-your-own notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/advanced_functionality/scikit_bring_your_own/scikit_bring_your_own.ipynb). We append specific steps that help you create a new Algorithm / Model Package SageMaker entities, which can be sold on AWS Marketplace
#
# By packaging an algorithm in a container, you can bring almost any code to the Amazon SageMaker environment, regardless of programming language, environment, framework, or dependencies.
#
# 1. [Building your own algorithm container](#Building-your-own-algorithm-container)
# 1. [When should I build my own algorithm container?](#When-should-I-build-my-own-algorithm-container?)
# 1. [Permissions](#Permissions)
# 1. [The example](#The-example)
# 1. [The presentation](#The-presentation)
# 1. [Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker](#Part-1:-Packaging-and-Uploading-your-Algorithm-for-use-with-Amazon-SageMaker)
# 1. [An overview of Docker](#An-overview-of-Docker)
# 1. [How Amazon SageMaker runs your Docker container](#How-Amazon-SageMaker-runs-your-Docker-container)
# 1. [Running your container during training](#Running-your-container-during-training)
# 1. [The input](#The-input)
# 1. [The output](#The-output)
# 1. [Running your container during hosting](#Running-your-container-during-hosting)
# 1. [The parts of the sample container](#The-parts-of-the-sample-container)
# 1. [The Dockerfile](#The-Dockerfile)
# 1. [Building and registering the container](#Building-and-registering-the-container)
# 1. [Testing your algorithm on your local machine or on an Amazon SageMaker notebook instance](#Testing-your-algorithm-on-your-local-machine-or-on-an-Amazon-SageMaker-notebook-instance)
# 1. [Part 2: Training and Hosting your Algorithm in Amazon SageMaker](#Part-2:-Training-and-Hosting-your-Algorithm-in-Amazon-SageMaker)
# 1. [Set up the environment](#Set-up-the-environment)
# 1. [Create the session](#Create-the-session)
# 1. [Upload the data for training](#Upload-the-data-for-training)
# 1. [Create an estimator and fit the model](#Create-an-estimator-and-fit-the-model)
# 1. [Run a Batch Transform Job](#Batch-Transform-Job)
# 1. [Deploy the model](#Deploy-the-model)
# 1. [Optional cleanup](#Cleanup-Endpoint)
# 1. [Part 3: Package your resources as an Amazon SageMaker Algorithm](#Part-3---Package-your-resources-as-an-Amazon-SageMaker-Algorithm)
# 1. [Algorithm Definition](#Algorithm-Definition)
# 1. [Part 4: Package your resources as an Amazon SageMaker ModelPackage](#Part-4---Package-your-resources-as-an-Amazon-SageMaker-ModelPackage)
# 1. [Model Package Definition](#Model-Package-Definition)
# 1. [Debugging Creation Issues](#Debugging-Creation-Issues)
# 1. [List on AWS Marketplace](#List-on-AWS-Marketplace)
#
# ## When should I build my own algorithm container?
#
# You may not need to create a container to bring your own code to Amazon SageMaker. When you are using a framework (such as Apache MXNet or TensorFlow) that has direct support in SageMaker, you can simply supply the Python code that implements your algorithm using the SDK entry points for that framework. This set of frameworks is continually expanding, so we recommend that you check the current list if your algorithm is written in a common machine learning environment.
#
# Even if there is direct SDK support for your environment or framework, you may find it more effective to build your own container. If the code that implements your algorithm is quite complex on its own or you need special additions to the framework, building your own container may be the right choice.
#
# If there isn't direct SDK support for your environment, don't worry. You'll see in this walk-through that building your own container is quite straightforward.
#
# ## Permissions
#
# Running this notebook requires permissions in addition to the normal `SageMakerFullAccess` permissions. This is because we'll creating new repositories in Amazon ECR. The easiest way to add these permissions is simply to add the managed policy `AmazonEC2ContainerRegistryFullAccess` to the role that you used to start your notebook instance. There's no need to restart your notebook instance when you do this, the new permissions will be available immediately.
#
# ## The example
#
# Here, we'll show how to package a simple Python example which showcases the [decision tree][] algorithm from the widely used [scikit-learn][] machine learning package. The example is purposefully fairly trivial since the point is to show the surrounding structure that you'll want to add to your own code so you can train and host it in Amazon SageMaker.
#
# The ideas shown here will work in any language or environment. You'll need to choose the right tools for your environment to serve HTTP requests for inference, but good HTTP environments are available in every language these days.
#
# In this example, we use a single image to support training and hosting. This is easy because it means that we only need to manage one image and we can set it up to do everything. Sometimes you'll want separate images for training and hosting because they have different requirements. Just separate the parts discussed below into separate Dockerfiles and build two images. Choosing whether to have a single image or two images is really a matter of which is more convenient for you to develop and manage.
#
# If you're only using Amazon SageMaker for training or hosting, but not both, there is no need to build the unused functionality into your container.
#
# [scikit-learn]: http://scikit-learn.org/stable/
# [decision tree]: http://scikit-learn.org/stable/modules/tree.html
#
# ## The presentation
#
# This presentation is divided into two parts: _building_ the container and _using_ the container.
# # Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker
#
# ### An overview of Docker
#
# If you're familiar with Docker already, you can skip ahead to the next section.
#
# For many data scientists, Docker containers are a new concept, but they are not difficult, as you'll see here.
#
# Docker provides a simple way to package arbitrary code into an _image_ that is totally self-contained. Once you have an image, you can use Docker to run a _container_ based on that image. Running a container is just like running a program on the machine except that the container creates a fully self-contained environment for the program to run. Containers are isolated from each other and from the host environment, so the way you set up your program is the way it runs, no matter where you run it.
#
# Docker is more powerful than environment managers like conda or virtualenv because (a) it is completely language independent and (b) it comprises your whole operating environment, including startup commands, environment variable, etc.
#
# In some ways, a Docker container is like a virtual machine, but it is much lighter weight. For example, a program running in a container can start in less than a second and many containers can run on the same physical machine or virtual machine instance.
#
# Docker uses a simple file called a `Dockerfile` to specify how the image is assembled. We'll see an example of that below. You can build your Docker images based on Docker images built by yourself or others, which can simplify things quite a bit.
#
# Docker has become very popular in the programming and devops communities for its flexibility and well-defined specification of the code to be run. It is the underpinning of many services built in the past few years, such as [Amazon ECS].
#
# Amazon SageMaker uses Docker to allow users to train and deploy arbitrary algorithms.
#
# In Amazon SageMaker, Docker containers are invoked in a certain way for training and a slightly different way for hosting. The following sections outline how to build containers for the SageMaker environment.
#
# Some helpful links:
#
# * [Docker home page](http://www.docker.com)
# * [Getting started with Docker](https://docs.docker.com/get-started/)
# * [Dockerfile reference](https://docs.docker.com/engine/reference/builder/)
# * [`docker run` reference](https://docs.docker.com/engine/reference/run/)
#
# [Amazon ECS]: https://aws.amazon.com/ecs/
#
# ### How Amazon SageMaker runs your Docker container
#
# Because you can run the same image in training or hosting, Amazon SageMaker runs your container with the argument `train` or `serve`. How your container processes this argument depends on the container:
#
# * In the example here, we don't define an `ENTRYPOINT` in the Dockerfile so Docker will run the command `train` at training time and `serve` at serving time. In this example, we define these as executable Python scripts, but they could be any program that we want to start in that environment.
# * If you specify a program as an `ENTRYPOINT` in the Dockerfile, that program will be run at startup and its first argument will be `train` or `serve`. The program can then look at that argument and decide what to do.
# * If you are building separate containers for training and hosting (or building only for one or the other), you can define a program as an `ENTRYPOINT` in the Dockerfile and ignore (or verify) the first argument passed in.
#
# #### Running your container during training
#
# When Amazon SageMaker runs training, your `train` script is run just like a regular Python program. A number of files are laid out for your use, under the `/opt/ml` directory:
#
# /opt/ml
# |-- input
# | |-- config
# | | |-- hyperparameters.json
# | | `-- resourceConfig.json
# | `-- data
# | `-- <channel_name>
# | `-- <input data>
# |-- model
# | `-- <model files>
# `-- output
# `-- failure
#
# ##### The input
#
# * `/opt/ml/input/config` contains information to control how your program runs. `hyperparameters.json` is a JSON-formatted dictionary of hyperparameter names to values. These values will always be strings, so you may need to convert them. `resourceConfig.json` is a JSON-formatted file that describes the network layout used for distributed training. Since scikit-learn doesn't support distributed training, we'll ignore it here.
# * `/opt/ml/input/data/<channel_name>/` (for File mode) contains the input data for that channel. The channels are created based on the call to CreateTrainingJob but it's generally important that channels match what the algorithm expects. The files for each channel will be copied from S3 to this directory, preserving the tree structure indicated by the S3 key structure.
# * `/opt/ml/input/data/<channel_name>_<epoch_number>` (for Pipe mode) is the pipe for a given epoch. Epochs start at zero and go up by one each time you read them. There is no limit to the number of epochs that you can run, but you must close each pipe before reading the next epoch.
#
# ##### The output
#
# * `/opt/ml/model/` is the directory where you write the model that your algorithm generates. Your model can be in any format that you want. It can be a single file or a whole directory tree. SageMaker will package any files in this directory into a compressed tar archive file. This file will be available at the S3 location returned in the `DescribeTrainingJob` result.
# * `/opt/ml/output` is a directory where the algorithm can write a file `failure` that describes why the job failed. The contents of this file will be returned in the `FailureReason` field of the `DescribeTrainingJob` result. For jobs that succeed, there is no reason to write this file as it will be ignored.
#
# #### Running your container during hosting
#
# Hosting has a very different model than training because hosting is reponding to inference requests that come in via HTTP. In this example, we use our recommended Python serving stack to provide robust and scalable serving of inference requests:
#
# 
#
# This stack is implemented in the sample code here and you can mostly just leave it alone.
#
# Amazon SageMaker uses two URLs in the container:
#
# * `/ping` will receive `GET` requests from the infrastructure. Your program returns 200 if the container is up and accepting requests.
# * `/invocations` is the endpoint that receives client inference `POST` requests. The format of the request and the response is up to the algorithm. If the client supplied `ContentType` and `Accept` headers, these will be passed in as well.
#
# The container will have the model files in the same place they were written during training:
#
# /opt/ml
# `-- model
# `-- <model files>
#
#
# ### The parts of the sample container
#
# In the `container` directory are all the components you need to package the sample algorithm for Amazon SageMager:
#
# .
# |-- Dockerfile
# |-- build_and_push.sh
# `-- decision_trees
# |-- nginx.conf
# |-- predictor.py
# |-- serve
# |-- train
# `-- wsgi.py
#
# Let's discuss each of these in turn:
#
# * __`Dockerfile`__ describes how to build your Docker container image. More details below.
# * __`build_and_push.sh`__ is a script that uses the Dockerfile to build your container images and then pushes it to ECR. We'll invoke the commands directly later in this notebook, but you can just copy and run the script for your own algorithms.
# * __`decision_trees`__ is the directory which contains the files that will be installed in the container.
# * __`local_test`__ is a directory that shows how to test your new container on any computer that can run Docker, including an Amazon SageMaker notebook instance. Using this method, you can quickly iterate using small datasets to eliminate any structural bugs before you use the container with Amazon SageMaker. We'll walk through local testing later in this notebook.
#
# In this simple application, we only install five files in the container. You may only need that many or, if you have many supporting routines, you may wish to install more. These five show the standard structure of our Python containers, although you are free to choose a different toolset and therefore could have a different layout. If you're writing in a different programming language, you'll certainly have a different layout depending on the frameworks and tools you choose.
#
# The files that we'll put in the container are:
#
# * __`nginx.conf`__ is the configuration file for the nginx front-end. Generally, you should be able to take this file as-is.
# * __`predictor.py`__ is the program that actually implements the Flask web server and the decision tree predictions for this app. You'll want to customize the actual prediction parts to your application. Since this algorithm is simple, we do all the processing here in this file, but you may choose to have separate files for implementing your custom logic.
# * __`serve`__ is the program started when the container is started for hosting. It simply launches the gunicorn server which runs multiple instances of the Flask app defined in `predictor.py`. You should be able to take this file as-is.
# * __`train`__ is the program that is invoked when the container is run for training. You will modify this program to implement your training algorithm.
# * __`wsgi.py`__ is a small wrapper used to invoke the Flask app. You should be able to take this file as-is.
#
# In summary, the two files you will probably want to change for your application are `train` and `predictor.py`.
# ### The Dockerfile
#
# The Dockerfile describes the image that we want to build. You can think of it as describing the complete operating system installation of the system that you want to run. A Docker container running is quite a bit lighter than a full operating system, however, because it takes advantage of Linux on the host machine for the basic operations.
#
# For the Python science stack, we will start from a standard Ubuntu installation and run the normal tools to install the things needed by scikit-learn. Finally, we add the code that implements our specific algorithm to the container and set up the right environment to run under.
#
# Along the way, we clean up extra space. This makes the container smaller and faster to start.
#
# Let's look at the Dockerfile for the example:
# !cat container/Dockerfile
# ### Building and registering the container
#
# The following shell code shows how to build the container image using `docker build` and push the container image to ECR using `docker push`. This code is also available as the shell script `container/build-and-push.sh`, which you can run as `build-and-push.sh decision_trees_sample` to build the image `decision_trees_sample`.
#
# This code looks for an ECR repository in the account you're using and the current default region (if you're using an Amazon SageMaker notebook instance, this will be the region where the notebook instance was created). If the repository doesn't exist, the script will create it.
# + language="sh"
#
# # The name of our algorithm
# algorithm_name=decision-trees-sample
#
# cd container
#
# chmod +x decision_trees/train
# chmod +x decision_trees/serve
#
# account=$(aws sts get-caller-identity --query Account --output text)
#
# # Get the region defined in the current configuration (default to us-west-2 if none defined)
# region=$(aws configure get region)
#
# fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
#
# # If the repository doesn't exist in ECR, create it.
#
# aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
#
# if [ $? -ne 0 ]
# then
# aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
# fi
#
# # Get the login command from ECR and execute it directly
# $(aws ecr get-login --region ${region} --no-include-email)
#
# # Build the docker image locally with the image name and then push it to ECR
# # with the full name.
#
# docker build -t ${algorithm_name} .
# docker tag ${algorithm_name} ${fullname}
#
# docker push ${fullname}
# -
# ## Testing your algorithm on your local machine or on an Amazon SageMaker notebook instance
#
# While you're first packaging an algorithm use with Amazon SageMaker, you probably want to test it yourself to make sure it's working right. In the directory `container/local_test`, there is a framework for doing this. It includes three shell scripts for running and using the container and a directory structure that mimics the one outlined above.
#
# The scripts are:
#
# * `train_local.sh`: Run this with the name of the image and it will run training on the local tree. You'll want to modify the directory `test_dir/input/data/...` to be set up with the correct channels and data for your algorithm. Also, you'll want to modify the file `input/config/hyperparameters.json` to have the hyperparameter settings that you want to test (as strings).
# * `serve_local.sh`: Run this with the name of the image once you've trained the model and it should serve the model. It will run and wait for requests. Simply use the keyboard interrupt to stop it.
# * `predict.sh`: Run this with the name of a payload file and (optionally) the HTTP content type you want. The content type will default to `text/csv`. For example, you can run `$ ./predict.sh payload.csv text/csv`.
#
# The directories as shipped are set up to test the decision trees sample algorithm presented here.
# # Part 2: Training, Batch Inference and Hosting your Algorithm in Amazon SageMaker
#
# Once you have your container packaged, you can use it to train and serve models. Let's do that with the algorithm we made above.
#
# ## Set up the environment
#
# Here we specify a bucket to use and the role that will be used for working with Amazon SageMaker.
# +
# S3 prefix
common_prefix = "DEMO-scikit-byo-iris"
training_input_prefix = common_prefix + "/training-input-data"
batch_inference_input_prefix = common_prefix + "/batch-inference-input-data"
import os
from sagemaker import get_execution_role
role = get_execution_role()
# -
# ## Create the session
#
# The session remembers our connection parameters to Amazon SageMaker. We'll use it to perform all of our SageMaker operations.
# +
import sagemaker as sage
sess = sage.Session()
# -
# ## Upload the data for training
#
# When training large models with huge amounts of data, you'll typically use big data tools, like Amazon Athena, AWS Glue, or Amazon EMR, to create your data in S3. For the purposes of this example, we're using some the classic [Iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set), which we have included.
#
# We can use use the tools provided by the Amazon SageMaker Python SDK to upload the data to a default bucket.
# +
TRAINING_WORKDIR = "data/training"
training_input = sess.upload_data(TRAINING_WORKDIR, key_prefix=training_input_prefix)
print ("Training Data Location " + training_input)
# -
# ## Create an estimator and fit the model
#
# In order to use Amazon SageMaker to fit our algorithm, we'll create an `Estimator` that defines how to use the container to train. This includes the configuration we need to invoke SageMaker training:
#
# * The __container name__. This is constructed as in the shell commands above.
# * The __role__. As defined above.
# * The __instance count__ which is the number of machines to use for training.
# * The __instance type__ which is the type of machine to use for training.
# * The __output path__ determines where the model artifact will be written.
# * The __session__ is the SageMaker session object that we defined above.
#
# Then we use fit() on the estimator to train against the data that we uploaded above.
account = sess.boto_session.client('sts').get_caller_identity()['Account']
region = sess.boto_session.region_name
image = '{}.dkr.ecr.{}.amazonaws.com/decision-trees-sample:latest'.format(account, region)
tree = sage.estimator.Estimator(image,
role, 1, 'ml.c4.2xlarge',
output_path="s3://{}/output".format(sess.default_bucket()),
sagemaker_session=sess)
tree.fit(training_input)
# ## Batch Transform Job
#
# Now let's use the model built to run a batch inference job and verify it works.
#
# ### Batch Transform Input Preparation
#
# The snippet below is removing the "label" column (column indexed at 0) and retaining the rest to be batch transform's input.
#
# NOTE: This is the same training data, which is a no-no from a statistical/ML science perspective. But the aim of this notebook is to demonstrate how things work end-to-end.
# +
import pandas as pd
## Remove first column that contains the label
shape=pd.read_csv(TRAINING_WORKDIR + "/iris.csv", header=None).drop([0], axis=1)
TRANSFORM_WORKDIR = "data/transform"
shape.to_csv(TRANSFORM_WORKDIR + "/batchtransform_test.csv", index=False, header=False)
transform_input = sess.upload_data(TRANSFORM_WORKDIR, key_prefix=batch_inference_input_prefix) + "/batchtransform_test.csv"
print("Transform input uploaded to " + transform_input)
# -
# ### Run Batch Transform
#
# Now that our batch transform input is setup, we run the transformation job next
# +
transformer = tree.transformer(instance_count=1, instance_type='ml.m4.xlarge')
transformer.transform(transform_input, content_type='text/csv')
transformer.wait()
print("Batch Transform output saved to " + transformer.output_path)
# -
# #### Inspect the Batch Transform Output in S3
# +
from urllib.parse import urlparse
parsed_url = urlparse(transformer.output_path)
bucket_name = parsed_url.netloc
file_key = '{}/{}.out'.format(parsed_url.path[1:], "batchtransform_test.csv")
s3_client = sess.boto_session.client('s3')
response = s3_client.get_object(Bucket = sess.default_bucket(), Key = file_key)
response_bytes = response['Body'].read().decode('utf-8')
print(response_bytes)
# -
# ## Deploy the model
#
# Deploying the model to Amazon SageMaker hosting just requires a `deploy` call on the fitted model. This call takes an instance count, instance type, and optionally serializer and deserializer functions. These are used when the resulting predictor is created on the endpoint.
# +
from sagemaker.predictor import csv_serializer
model = tree.create_model()
predictor = tree.deploy(1, 'ml.m4.xlarge', serializer=csv_serializer)
# -
# ### Choose some data and use it for a prediction
#
# In order to do some predictions, we'll extract some of the data we used for training and do predictions against it. This is, of course, bad statistical practice, but a good way to see how the mechanism works.
# +
shape=pd.read_csv(TRAINING_WORKDIR + "/iris.csv", header=None)
import itertools
a = [50*i for i in range(3)]
b = [40+i for i in range(10)]
indices = [i+j for i,j in itertools.product(a,b)]
test_data=shape.iloc[indices[:-1]]
test_X=test_data.iloc[:,1:]
test_y=test_data.iloc[:,0]
# -
# Prediction is as easy as calling predict with the predictor we got back from deploy and the data we want to do predictions with. The serializers take care of doing the data conversions for us.
print(predictor.predict(test_X.values).decode('utf-8'))
# ### Cleanup Endpoint
#
# When you're done with the endpoint, you'll want to clean it up.
sess.delete_endpoint(predictor.endpoint)
# # Part 3 - Package your resources as an Amazon SageMaker Algorithm
# (If you looking to sell a pretrained model (ModelPackage), please skip to Part 4.)
#
# Now that you have verified that the algorithm code works for training, live inference and batch inference in the above sections, you can start packaging it up as an Amazon SageMaker Algorithm.
# +
import boto3
smmp = boto3.client('sagemaker')
# -
# ## Algorithm Definition
#
# SageMaker Algorithm is comprised of 2 parts:
#
# 1. A training image
# 2. An inference image (optional)
#
# The key requirement is that the training and inference images (if provided) remain compatible with each other. Specifically, the model artifacts generated by the code in training image should be readable and compatible with the code in inference image.
#
# You can reuse the same image to perform both training and inference or you can choose to separate them.
#
#
# This sample notebook has already created a single algorithm image that perform both training and inference. This image has also been pushed to your ECR registry at {{image}}. You need to provide the following details as part of this algorithm specification:
#
# #### Training Specification
#
# You specify details pertinent to your training algorithm in this section.
#
# #### Supported Hyper-parameters
#
# This section captures the hyper-parameters your algorithm supports, their names, types, if they are required, default values, valid ranges etc. This serves both as documentation for buyers and is used by Amazon SageMaker to perform validations of buyer requests in the synchronous request path.
#
# Please Note: While this section is optional, we strongly recommend you provide comprehensive information here to leverage our validations and serve as documentation. Additionally, without this being specified, customers cannot leverage your algorithm for Hyper-parameter tuning.
#
# *** NOTE: The code below has hyper-parameters hard-coded in the json present in src/training_specification.py. Until we have better functionality to customize it, please update the json in that file appropriately***
#
#
# +
from src.training_specification import TrainingSpecification
from src.training_channels import TrainingChannels
from src.metric_definitions import MetricDefinitions
from src.tuning_objectives import TuningObjectives
import json
training_specification = TrainingSpecification().get_training_specification_dict(
ecr_image=image,
supports_gpu=True,
supported_channels=[
TrainingChannels("training", description="Input channel that provides training data", supported_content_types=["text/csv"])],
supported_metrics=[MetricDefinitions("validation:accuracy", "validation-accuracy: (\\S+)")],
supported_tuning_job_objective_metrics=[TuningObjectives("Maximize", "validation:accuracy")]
)
print(json.dumps(training_specification, indent=2, sort_keys=True))
# -
# #### Inference Specification
#
# You specify details pertinent to your inference code in this section.
#
# +
from src.inference_specification import InferenceSpecification
import json
inference_specification = InferenceSpecification().get_inference_specification_dict(
ecr_image=image,
supports_gpu=True,
supported_content_types=["text/csv"],
supported_mime_types=["text/csv"])
print(json.dumps(inference_specification, indent=4, sort_keys=True))
# -
# #### Validation Specification
#
# In order to provide confidence to the sellers (and buyers) that the products work in Amazon SageMaker before listing them on AWS Marketplace, SageMaker needs to perform basic validations. The product can be listed in AWS Marketplace only if this validation process succeeds. This validation process uses the validation profile and sample data provided by you to run the following validations:
#
# 1. Create a training job in your account to verify your training image works with SageMaker.
# 2. Once the training job completes successfully, create a Model in your account using the algorithm's inference image and the model artifacts produced as part of the training job we ran.
# 3. Create a transform job in your account using the above Model to verify your inference image works with SageMaker
# +
from src.algorithm_validation_specification import AlgorithmValidationSpecification
import json
validation_specification = AlgorithmValidationSpecification().get_algo_validation_specification_dict(
validation_role = role,
training_channel_name = "training",
training_input = training_input,
batch_transform_input = transform_input,
content_type = "text/csv",
instance_type = "ml.c4.xlarge",
output_s3_location = 's3://{}/{}'.format(sess.default_bucket(), common_prefix))
print(json.dumps(validation_specification, indent=4, sort_keys=True))
# -
# ## Putting it all together
#
# Now we put all the pieces together in the next cell and create an Amazon SageMaker Algorithm
# +
import json
import time
algorithm_name = "scikit-decision-trees-" + str(round(time.time()))
create_algorithm_input_dict = {
"AlgorithmName" : algorithm_name,
"AlgorithmDescription" : "Decision trees using Scikit",
"CertifyForMarketplace" : True
}
create_algorithm_input_dict.update(training_specification)
create_algorithm_input_dict.update(inference_specification)
create_algorithm_input_dict.update(validation_specification)
print(json.dumps(create_algorithm_input_dict, indent=4, sort_keys=True))
print ("Now creating an algorithm in SageMaker")
smmp.create_algorithm(**create_algorithm_input_dict)
# -
# ### Describe the algorithm
#
# The next cell describes the Algorithm and waits until it reaches a terminal state (Completed or Failed)
# +
import time
import json
while True:
response = smmp.describe_algorithm(AlgorithmName=algorithm_name)
status = response["AlgorithmStatus"]
print (status)
if (status == "Completed" or status == "Failed"):
print (response["AlgorithmStatusDetails"])
break
time.sleep(5)
# -
# # Part 4 - Package your resources as an Amazon SageMaker ModelPackage
#
# In this section, we will see how you can package your artifacts (ECR image and the trained artifact from your previous training job) into a ModelPackage. Once you complete this, you can list your product as a pretrained model in the AWS Marketplace.
#
# ## Model Package Definition
# A Model Package is a reusable model artifacts abstraction that packages all ingredients necessary for inference. It consists of an inference specification that defines the inference image to use along with an optional model weights location.
#
smmp = boto3.client('sagemaker')
# #### Inference Specification
#
# You specify details pertinent to your inference code in this section.
#
# +
from src.inference_specification import InferenceSpecification
import json
modelpackage_inference_specification = InferenceSpecification().get_inference_specification_dict(
ecr_image=image,
supports_gpu=True,
supported_content_types=["text/csv"],
supported_mime_types=["text/csv"])
# Specify the model data resulting from the previously completed training job
modelpackage_inference_specification["InferenceSpecification"]["Containers"][0]["ModelDataUrl"]=tree.model_data
print(json.dumps(modelpackage_inference_specification, indent=4, sort_keys=True))
# -
# #### Validation Specification
#
# In order to provide confidence to the sellers (and buyers) that the products work in Amazon SageMaker before listing them on AWS Marketplace, SageMaker needs to perform basic validations. The product can be listed in the AWS Marketplace only if this validation process succeeds. This validation process uses the validation profile and sample data provided by you to run the following validations:
#
# * Create a transform job in your account using the above Model to verify your inference image works with SageMaker.
#
# +
from src.modelpackage_validation_specification import ModelPackageValidationSpecification
import json
modelpackage_validation_specification = ModelPackageValidationSpecification().get_validation_specification_dict(
validation_role = role,
batch_transform_input = transform_input,
content_type = "text/csv",
instance_type = "ml.c4.xlarge",
output_s3_location = 's3://{}/{}'.format(sess.default_bucket(), common_prefix))
print(json.dumps(modelpackage_validation_specification, indent=4, sort_keys=True))
# -
# ## Putting it all together
#
# Now we put all the pieces together in the next cell and create an Amazon SageMaker Model Package.
# +
import json
import time
model_package_name = "scikit-iris-detector-" + str(round(time.time()))
create_model_package_input_dict = {
"ModelPackageName" : model_package_name,
"ModelPackageDescription" : "Model to detect 3 different types of irises (Setosa, Versicolour, and Virginica)",
"CertifyForMarketplace" : True
}
create_model_package_input_dict.update(modelpackage_inference_specification)
create_model_package_input_dict.update(modelpackage_validation_specification)
print(json.dumps(create_model_package_input_dict, indent=4, sort_keys=True))
smmp.create_model_package(**create_model_package_input_dict)
# -
# #### Describe the ModelPackage
#
# The next cell describes the ModelPackage and waits until it reaches a terminal state (Completed or Failed)
# +
import time
import json
while True:
response = smmp.describe_model_package(ModelPackageName=model_package_name)
status = response["ModelPackageStatus"]
print (status)
if (status == "Completed" or status == "Failed"):
print (response["ModelPackageStatusDetails"])
break
time.sleep(5)
# -
# ## Debugging Creation Issues
#
# Entity creation typically never fails in the synchronous path. However, the validation process can fail for many reasons. If the above Algorithm creation fails, you can investigate the cause for the failure by looking at the "AlgorithmStatusDetails" field in the Algorithm object or "ModelPackageStatusDetails" field in the ModelPackage object. You can also look for the Training Jobs / Transform Jobs created in your account as part of our validation and inspect their logs for more hints on what went wrong.
#
# If all else fails, please contact AWS Customer Support for assistance!
#
# ## List on AWS Marketplace
#
# Next, please go back to the Amazon SageMaker console, click on "Algorithms" (or "Model Packages") and you'll find the entity you created above. If it was successfully created and validated, you should be able to select the entity and "Publish new ML Marketplace listing" from SageMaker console.
# <img src="images/publish-to-marketplace-action.png"/>
|
aws_marketplace/creating_marketplace_products/Bring_Your_Own-Creating_Algorithm_and_Model_Package.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from itertools import zip_longest
def return_dict_from_csv_line(header, line):
# Zip them
zipped_line = zip_longest(header, line, fillvalue=None)
# Use dict comprehension to generate the final dict
ret_dict = {kv[0]: kv[1] for kv in zipped_line}
return ret_dict
open("sales_record.csv", "r") as fd
first_line = fd.readline()
header = first_line.replace("\n", "").split(",")
for i, line in enumerate(fd):
line = line.replace("\n", "").split(",")
d = return_dict_from_csv_line(header, line)
print(d)
if i > 10:
break
|
Chapter02/.ipynb_checkpoints/Activity 2.02-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="vfxqky4PCUnh"
# # How JAX primitives work
#
# [](https://colab.research.google.com/github/google/jax/blob/main/docs/notebooks/How_JAX_primitives_work.ipynb)
#
# *<EMAIL>*, October 2019.
#
# JAX implements certain transformations of Python functions, e.g., `jit`, `grad`,
# `vmap`, or `pmap`. The Python functions to be transformed must be JAX-traceable,
# which means that as the Python function executes
# the only operations it applies to the data are either inspections of data
# attributes such as shape or type, or special operations called JAX primitives.
# In particular, a JAX-traceable function is sometimes invoked by JAX with
# abstract arguments. An example of a JAX abstract value is `ShapedArray(float32[2,2])`,
# which captures the type and the shape of values, but not the concrete data values.
# JAX primitives know how to operate on both concrete data
# values and on the JAX abstract values.
#
#
# The JAX-transformed functions must themselves be JAX-traceable functions,
# to ensure that these transformations
# can be composed, e.g., `jit(jacfwd(grad(f)))`.
#
# There are pre-defined JAX primitives corresponding to most XLA operations,
# e.g., add, matmul, sin, cos, indexing.
# JAX comes with an implementation of numpy functions in terms of JAX primitives, which means that Python programs
# using JAX’s implementation of numpy are JAX-traceable and therefore transformable.
# Other libraries can be made JAX-traceable by implementing them in terms of JAX primitives.
#
# The set of JAX primitives is extensible. Instead of reimplementing a function in terms of pre-defined JAX primitives,
# one can define a new primitive that encapsulates the behavior of the function.
#
# **The goal of this document is to explain the interface that a JAX primitive must support in order to allow JAX to perform all its transformations.**
#
# Consider that we want to add to JAX support for a multiply-add function with three arguments, defined mathematically
# as "multiply_add(x, y, z) = x * y + z".
# This function operates on 3 identically-shaped tensors of floating point
# values and performs the operations pointwise.
# + [markdown] id="HIJYIHNTD1yI"
# ## Using existing primitives
#
# The easiest way to define new functions is to write them in terms of JAX primitives, or in terms of other
# functions that are themselves written using JAX primitives, e.g., those
# defined in the `jax.lax` module:
# + id="tbOF0LB0EMne" outputId="3fb1c8a7-7a4c-4a3a-f7ff-37b7dc740528"
from jax import lax
from jax._src import api
def multiply_add_lax(x, y, z):
"""Implementation of multiply-add using the jax.lax primitives."""
return lax.add(lax.mul(x, y), z)
def square_add_lax(a, b):
"""A square-add function using the newly defined multiply-add."""
return multiply_add_lax(a, a, b)
print("square_add_lax = ", square_add_lax(2., 10.))
# Differentiate w.r.t. the first argument
print("grad(square_add_lax) = ", api.grad(square_add_lax, argnums=0)(2.0, 10.))
# + [markdown] id="Cgv60Wm3E_D5"
# In order to understand how JAX is internally using the primitives,
# we add some helpers for tracing function calls.
# + cellView="form" id="mQRQGEGiE53K"
#@title Helper functions (execute this cell)
import functools
import traceback
_indentation = 0
def _trace(msg=None):
"""Print a message at current indentation."""
if msg is not None:
print(" " * _indentation + msg)
def _trace_indent(msg=None):
"""Print a message and then indent the rest."""
global _indentation
_trace(msg)
_indentation = 1 + _indentation
def _trace_unindent(msg=None):
"""Unindent then print a message."""
global _indentation
_indentation = _indentation - 1
_trace(msg)
def trace(name):
"""A decorator for functions to trace arguments and results."""
def trace_func(func): # pylint: disable=missing-docstring
def pp(v):
"""Print certain values more succinctly"""
vtype = str(type(v))
if "jax.lib.xla_bridge._JaxComputationBuilder" in vtype:
return "<JaxComputationBuilder>"
elif "jaxlib.xla_extension.XlaOp" in vtype:
return "<XlaOp at 0x{:x}>".format(id(v))
elif ("partial_eval.JaxprTracer" in vtype or
"batching.BatchTracer" in vtype or
"ad.JVPTracer" in vtype):
return "Traced<{}>".format(v.aval)
elif isinstance(v, tuple):
return "({})".format(pp_values(v))
else:
return str(v)
def pp_values(args):
return ", ".join([pp(arg) for arg in args])
@functools.wraps(func)
def func_wrapper(*args):
_trace_indent("call {}({})".format(name, pp_values(args)))
res = func(*args)
_trace_unindent("|<- {} = {}".format(name, pp(res)))
return res
return func_wrapper
return trace_func
class expectNotImplementedError(object):
"""Context manager to check for NotImplementedError."""
def __enter__(self): pass
def __exit__(self, type, value, tb):
global _indentation
_indentation = 0
if type is NotImplementedError:
print("\nFound expected exception:")
traceback.print_exc(limit=3)
return True
elif type is None: # No exception
assert False, "Expected NotImplementedError"
else:
return False
# + [markdown] id="Qf4eLrLCFYDl"
# Instead of using `jax.lax` primitives directly, we can use other functions
# that are already written in terms of those primitives, such as those in `jax.numpy`:
# + id="QhKorz6cFRJb" outputId="aba3cef3-6bcc-4eb3-c7b3-34e405f2f82a"
import jax.numpy as jnp
import numpy as np
@trace("multiply_add_numpy")
def multiply_add_numpy(x, y, z):
return jnp.add(jnp.multiply(x, y), z)
@trace("square_add_numpy")
def square_add_numpy(a, b):
return multiply_add_numpy(a, a, b)
print("\nNormal evaluation:")
print("square_add_numpy = ", square_add_numpy(2., 10.))
print("\nGradient evaluation:")
print("grad(square_add_numpy) = ", api.grad(square_add_numpy)(2.0, 10.))
# + [markdown] id="Sg-D8EdeFn4a"
# Notice that in the process of computing `grad`, JAX invokes `square_add_numpy` and
# `multiply_add_numpy` with special arguments `ConcreteArray(...)` (described further
# below in this colab).
# It is important to remember that a JAX-traceable function must be able to
# operate not only on concrete arguments but also on special abstract arguments
# that JAX may use to abstract the function execution.
#
# The JAX traceability property is satisfied as long as the function is written
# in terms of JAX primitives.
# + [markdown] id="WxrQO7-XGLcg"
# ## Defining new JAX primitives
#
# The right way to add support for multiply-add is in terms of existing
# JAX primitives, as shown above. However, in order to demonstrate how JAX
# primitives work let us pretend that we want to add a new primitive to
# JAX for the multiply-add functionality.
# + id="cPqAH1XOGTN4"
from jax import core
multiply_add_p = core.Primitive("multiply_add") # Create the primitive
@trace("multiply_add_prim")
def multiply_add_prim(x, y, z):
"""The JAX-traceable way to use the JAX primitive.
Note that the traced arguments must be passed as positional arguments
to `bind`.
"""
return multiply_add_p.bind(x, y, z)
@trace("square_add_prim")
def square_add_prim(a, b):
"""A square-add function implemented using the new JAX-primitive."""
return multiply_add_prim(a, a, b)
# + [markdown] id="LMzs5PAKGr-4"
# If we try to call the newly defined functions we get an error, because
# we have not yet told JAX anything about the semantics of the new primitive.
# + id="_X3PAYxhGpWd" outputId="90ea2c6a-9ef3-40ea-e9a3-3ab1cfc59fc8"
with expectNotImplementedError():
square_add_prim(2., 10.)
# + [markdown] id="elha0FdgHSEF"
# ### Primal evaluation rules
# + id="FT34FFAGHARU" outputId="4c54f1c2-8a50-4788-90e1-06aee412c43b"
@trace("multiply_add_impl")
def multiply_add_impl(x, y, z):
"""Concrete implementation of the primitive.
This function does not need to be JAX traceable.
Args:
x, y, z: the concrete arguments of the primitive. Will only be called with
concrete values.
Returns:
the concrete result of the primitive.
"""
# Note that we can use the original numpy, which is not JAX traceable
return np.add(np.multiply(x, y), z)
# Now we register the primal implementation with JAX
multiply_add_p.def_impl(multiply_add_impl)
# + id="G5bstKaeNAVV" outputId="deb94d5b-dfea-4e6f-9ec2-70b416c996c5"
assert square_add_prim(2., 10.) == 14.
# + [markdown] id="upBf-uAuHhPJ"
# ### JIT
#
# If we now try to use `jit` we get a `NotImplementedError`:
# + id="QG-LULjiHk4b" outputId="d4ef4406-8dae-4c96-97ca-b662340474ee"
with expectNotImplementedError():
api.jit(square_add_prim)(2., 10.)
# + [markdown] id="rHS1bAGHH44E"
# #### Abstract evaluation rules
# In order to JIT the function, and for other transformations as well,
# JAX first evaluates it abstractly using only the
# shape and type of the arguments. This abstract evaluation serves multiple
# purposes:
#
# * Gets the sequence of JAX primitives that are used in the computation. This
# sequence will be compiled.
# * Computes the shape and type of all vectors and operations used in the computation.
#
#
# For example, the abstraction of a vector with 3 elements may be `ShapedArray(float32[3])`, or `ConcreteArray([1., 2., 3.])`.
# In the latter case, JAX uses the actual concrete value wrapped as an abstract value.
# + id="ctQmEeckIbdo" outputId="e751d0cc-460e-4ffd-df2e-fdabf9cffdc2"
from jax._src import abstract_arrays
@trace("multiply_add_abstract_eval")
def multiply_add_abstract_eval(xs, ys, zs):
"""Abstract evaluation of the primitive.
This function does not need to be JAX traceable. It will be invoked with
abstractions of the actual arguments.
Args:
xs, ys, zs: abstractions of the arguments.
Result:
a ShapedArray for the result of the primitive.
"""
assert xs.shape == ys.shape
assert xs.shape == zs.shape
return abstract_arrays.ShapedArray(xs.shape, xs.dtype)
# Now we register the abstract evaluation with JAX
multiply_add_p.def_abstract_eval(multiply_add_abstract_eval)
# + [markdown] id="RPN88X6YI43A"
# If we re-attempt to JIT, we see how the abstract evaluation proceeds, but
# we get another error, about missing the actual XLA compilation rule:
# + id="eOcNR92SI2h-" outputId="356ef229-3703-4696-cc3d-7c05de405fb0"
with expectNotImplementedError():
api.jit(square_add_prim)(2., 10.)
# + [markdown] id="9IOV1R-fJMHp"
# #### XLA Compilation rules
#
# JAX compilation works by compiling each primitive into a graph of XLA operations.
#
# This is the biggest hurdle to adding new functionality to JAX, because the
# set of XLA operations is limited, and JAX already has pre-defined primitives
# for most of them. However, XLA includes a `CustomCall` operation that can be used to encapsulate arbitrary functionality defined using C++.
# + id="FYQWSSjKJaWP"
from jax.lib import xla_client
@trace("multiply_add_xla_translation")
def multiply_add_xla_translation(c, xc, yc, zc):
"""The compilation to XLA of the primitive.
Given an XlaBuilder and XlaOps for each argument, return the XlaOp for the
result of the function.
Does not need to be a JAX-traceable function.
"""
return xla_client.ops.Add(xla_client.ops.Mul(xc, yc), zc)
# Now we register the XLA compilation rule with JAX
# TODO: for GPU? and TPU?
from jax.interpreters import xla
xla.backend_specific_translations['cpu'][multiply_add_p] = multiply_add_xla_translation
# + [markdown] id="K98LX-VaJkFu"
# Now we succeed to JIT. Notice below that JAX first evaluates the function
# abstractly, which triggers the `multiply_add_abstract_eval` function, and
# then compiles the set of primitives it has encountered, including `multiply_add`.
# At this point JAX invokes `multiply_add_xla_translation`.
# + id="rj3TLsolJgEc" outputId="e384bee4-1e9c-4344-f49c-d3b5ec08eb32"
assert api.jit(lambda x, y: square_add_prim(x, y))(2., 10.) == 14.
# + [markdown] id="Omrez-2_KFfo"
# Below is another use of `jit` where we compile only
# with respect to the first argument. Notice how the second argument to `square_add_prim` is concrete, which leads
# in the third argument to `multiply_add_abstract_eval` being
# `ConcreteArray`. We see that `multiply_add_abstract_eval` may be used with
# both `ShapedArray` and `ConcreteArray`.
# + id="mPfTwIBoKOEK" outputId="b293b9b6-a2f9-48f5-f7eb-d4f99c3d905b"
assert api.jit(lambda x, y: square_add_prim(x, y),
static_argnums=1)(2., 10.) == 14.
# + [markdown] id="_Ya3B5l4J1VA"
# ### Forward differentiation
#
# JAX implements forward differentiation in the form of
# a Jacobian-vector product (see the [JAX autodiff cookbook](https://jax.readthedocs.io/en/latest/notebooks/autodiff_cookbook.html#Jacobian-Matrix-and-Matrix-Jacobian-products)).
#
# If we attempt now to compute the `jvp` function we get an
# error because we have not yet told JAX how to differentiate
# the `multiply_add` primitive.
# + id="OxDx6NQnKwMI" outputId="ce659ef3-c03c-4856-f252-49ec4b6eb964"
# The second argument `(2., 10.)` are the argument values
# where we evaluate the Jacobian, and the third `(1., 1.)`
# are the values of the tangents for the arguments.
with expectNotImplementedError():
api.jvp(square_add_prim, (2., 10.), (1., 1.))
# + id="zxG24C1JMIMM"
from jax.interpreters import ad
@trace("multiply_add_value_and_jvp")
def multiply_add_value_and_jvp(arg_values, arg_tangents):
"""Evaluates the primal output and the tangents (Jacobian-vector product).
Given values of the arguments and perturbation of the arguments (tangents),
compute the output of the primitive and the perturbation of the output.
This method must be JAX-traceable. JAX may invoke it with abstract values
for the arguments and tangents.
Args:
arg_values: a tuple of arguments
arg_tangents: a tuple with the tangents of the arguments. The tuple has
the same length as the arg_values. Some of the tangents may also be the
special value ad.Zero to specify a zero tangent.
Returns:
a pair of the primal output and the tangent.
"""
x, y, z = arg_values
xt, yt, zt = arg_tangents
_trace("Primal evaluation:")
# Now we have a JAX-traceable computation of the output.
# Normally, we can use the ma primtive itself to compute the primal output.
primal_out = multiply_add_prim(x, y, z)
_trace("Tangent evaluation:")
# We must use a JAX-traceable way to compute the tangent. It turns out that
# the output tangent can be computed as (xt * y + x * yt + zt),
# which we can implement in a JAX-traceable way using the same "multiply_add_prim" primitive.
# We do need to deal specially with Zero. Here we just turn it into a
# proper tensor of 0s (of the same shape as 'x').
# An alternative would be to check for Zero and perform algebraic
# simplification of the output tangent computation.
def make_zero(tan):
return lax.zeros_like_array(x) if type(tan) is ad.Zero else tan
output_tangent = multiply_add_prim(make_zero(xt), y, multiply_add_prim(x, make_zero(yt), make_zero(zt)))
return (primal_out, output_tangent)
# Register the forward differentiation rule with JAX
ad.primitive_jvps[multiply_add_p] = multiply_add_value_and_jvp
# + id="ma3KBkiAMfW1" outputId="f34cbbc6-20d9-48ca-9a9a-b5d91a972cdd"
# Tangent is: xt*y + x*yt + zt = 1.*2. + 2.*1. + 1. = 5.
assert api.jvp(square_add_prim, (2., 10.), (1., 1.)) == (14., 5.)
# + [markdown] id="69QsEcu-lP4u"
# TO EXPLAIN:
#
# * Why is JAX using ConcreteArray in square_add_prim? There is no abstract evaluation going on here.
# * Not sure how to explain that multiply_add_prim is invoked with ConcreteValue, yet
# we do not call the multiply_add_abstract_eval.
# * I think it would be useful to show the jaxpr here
# + [markdown] id="Sb6e3ZAHOPHv"
# #### JIT of forward differentiation
#
# We can apply JIT to the forward differentiation function:
# + id="hg-hzVu-N-hv" outputId="38d32067-e152-4046-ad80-7f95a31ba628"
assert api.jit(lambda arg_values, arg_tangents:
api.jvp(square_add_prim, arg_values, arg_tangents))(
(2., 10.), (1., 1.)) == (14., 5.)
# + [markdown] id="jlZt1_v2mU88"
# Notice that first we evaluate `multiply_add_value_and_jvp` abstractly, which in turn
# evaluates abstractly both the primal and the tangent evaluation (a total of
# 3 invocations of the `ma` primitive). Then we compile the 3 occurrences
# of the primitive.
# + [markdown] id="555yt6ZIOePB"
# ### Reverse differentiation
#
# If we attempt now to use reverse differentiation we
# see that JAX starts by using the `multiply_add_value_and_jvp` to
# compute the forward differentiation for abstract values, but then runs
# into a `NotImplementedError`.
#
# When computing the reverse differentiation JAX first does abstract evaluation
# of the forward differentiation code `multiply_add_value_and_jvp` to obtain a
# trace of primitives that compute the output tangent.
# Observe that JAX performs this abstract evaluation with concrete values
# for the differentiation point, and abstract values for the tangents.
# Observe also that JAX uses the special abstract tangent value `Zero` for
# the tangent corresponding to the 3rd argument of `ma`. This reflects the
# fact that we do not differentiate w.r.t. the 2nd argument to `square_add_prim`,
# which flows to the 3rd argument to `multiply_add_prim`.
#
# Observe also that during the abstract evaluation of the tangent we pass the
# value 0.0 as the tangent for the 3rd argument. This is due to the use
# of the `make_zero` function in the definition of `multiply_add_value_and_jvp`.
# + id="8eAVnexaOjBn" outputId="e4ee89cf-ab4a-4505-9817-fa978a2865ab"
# This is reverse differentiation w.r.t. the first argument of square_add_prim
with expectNotImplementedError():
api.grad(square_add_prim)(2., 10.)
# + [markdown] id="fSHLUMDN26AY"
# The above error is because there is a missing piece for JAX to be able
# to use the forward differentiation code to compute reverse differentiation.
# + [markdown] id="3ibDbGF-PjK9"
# #### Transposition
#
#
# As explained above, when computing reverse differentiation JAX obtains
# a trace of primitives that compute the tangent using forward differentiation.
# Then, **JAX interprets this trace abstractly backwards** and for each
# primitive it applies a **transposition** rule.
#
# To understand what is going on, consider for now a simpler example of the function "f(x, y) = x * y + y". Assume we need to differentiate at the point `(2., 4.)`. JAX will produce the following JVP tangent calculation of `ft` from the tangents of the input `xt` and `yt`:
# ```
# a = xt * 4.
# b = 2. * yt
# c = a + b
# ft = c + yt
# ```
#
# By construction, the tangent calculation is always linear in the input tangents.
# The only non-linear operator that may arise in the tangent calculation is multiplication,
# but then one of the operands is constant.
#
# JAX will produce the reverse differentiation computation by processing the
# JVP computation backwards. For each operation in the tangent computation,
# it accumulates the cotangents
# of the variables used by the operation, using the cotangent of the result
# of the operation:
# ```
# # Initialize cotangents of inputs and intermediate vars
# xct = yct = act = bct = cct = 0.
# # Initialize cotangent of the output
# fct = 1.
# # Process "ft = c + yt"
# cct += fct
# yct += fct
# # Process "c = a + b"
# act += cct
# bct += cct
# # Process "b = 2. * yt"
# yct += 2. * bct
# # Process "a = xt * 4."
# xct += act * 4.
# ```
#
# One can verify that this computation produces `xct = 4.` and `yct = 3.`, which
# are the partial derivatives of the function `f`.
#
# JAX knows for each primitive that may appear in a JVP calculation how to transpose it. Conceptually, if the primitive `p(x, y, z)` is linear in the arguments `y` and `z` for a constant value of `x`, e.g., `p(x, y, z) = y*cy + z*cz`, then the transposition of the primitive is:
# ```
# p_transpose(out_ct, x, _, _) = (None, out_ct*cy, out_ct*cz)
# ```
#
# Notice that `p_transpose` takes the cotangent of the output of the primitive and a value corresponding to each argument of the primitive. For the linear arguments, the transposition gets an undefined `_` value, and for the other
# arguments it gets the actual constants. The transposition returns a cotangent value for each argument of the primitive, with the value `None` returned
# for the constant arguments.
#
# In particular,
# ```
# add_transpose(out_ct, _, _) = (out_ct, out_ct)
# mult_transpose(out_ct, x, _) = (None, x * out_ct)
# mult_transpose(out_ct, _, y) = (out_ct * y, None)
# ```
# + id="JaHxFdkRO42r"
@trace("multiply_add_transpose")
def multiply_add_transpose(ct, x, y, z):
"""Evaluates the transpose of a linear primitive.
This method is only used when computing the backward gradient following
value_and_jvp, and is only needed for primitives that are used in the JVP
calculation for some other primitive. We need transposition for multiply_add_prim,
because we have used multiply_add_prim in the computation of the output_tangent in
multiply_add_value_and_jvp.
In our case, multiply_add is not a linear primitive. However, it is used linearly
w.r.t. tangents in multiply_add_value_and_jvp:
output_tangent(xt, yt, zt) = multiply_add_prim(xt, y, multiply_add_prim(x, yt, zt))
Always one of the first two multiplicative arguments is a constant.
Args:
ct: the cotangent of the output of the primitive.
x, y, z: values of the arguments. The arguments that are used linearly
get an ad.UndefinedPrimal value. The other arguments get a constant
value.
Returns:
a tuple with the cotangent of the inputs, with the value None
corresponding to the constant arguments.
"""
if not ad.is_undefined_primal(x):
# This use of multiply_add is with a constant "x"
assert ad.is_undefined_primal(y)
ct_y = ad.Zero(y.aval) if type(ct) is ad.Zero else multiply_add_prim(x, ct, lax.zeros_like_array(x))
res = None, ct_y, ct
else:
# This use of multiply_add is with a constant "y"
assert ad.is_undefined_primal(x)
ct_x = ad.Zero(x.aval) if type(ct) is ad.Zero else multiply_add_prim(ct, y, lax.zeros_like_array(y))
res = ct_x, None, ct
return res
ad.primitive_transposes[multiply_add_p] = multiply_add_transpose
# + [markdown] id="PpChox-Jp7wb"
# Now we can complete the run of the `grad`:
# + id="PogPKS4MPevd" outputId="d33328d4-3e87-45b5-9b31-21ad624b67af"
assert api.grad(square_add_prim)(2., 10.) == 4.
# + [markdown] id="8M1xLCXW4fK7"
# Notice the two calls to `multiply_add_transpose`. They correspond to the two
# uses of `multiply_add_prim` in the computation of the `output_tangent` in `multiply_add_value_and_jvp`. The first call to transpose corresponds to the
# last use of `multiply_add_prim`: `multiply_add_prim(xt, y, ...)` where `y` is the constant 2.0.
# + [markdown] id="EIJs6FYmPg6c"
# #### JIT of reverse differentiation
#
# Notice that the abstract evaluation of the `multiply_add_value_and_jvp` is using only
# abstract values, while in the absence of JIT we used `ConcreteArray`.
# + id="FZ-JGbWZPq2-" outputId="e42b5222-9c3e-4853-e13a-874f6605d178"
assert api.jit(api.grad(square_add_prim))(2., 10.) == 4.
# + [markdown] id="-3lqPkdQPvl5"
# ### Batching
#
# The batching transformation takes a point-wise computation and turns it
# into a computation on vectors. If we try it right now, we get a `NotImplementedError`:
# + id="hFvBR3I9Pzh3" outputId="434608bc-281f-4d3b-83bd-eaaf3b51b1cd"
# The arguments are two vectors instead of two scalars
with expectNotImplementedError():
api.vmap(square_add_prim, in_axes=0, out_axes=0)(np.array([2., 3.]),
np.array([10., 20.]))
# + [markdown] id="gILasMiP6elR"
# We need to tell JAX how to evaluate the batched version of the primitive. In this particular case, the `multiply_add_prim` already operates pointwise for any dimension of input vectors. So the batched version can use the same `multiply_add_prim` implementation.
# + id="KQfeqRIrP7zg"
from jax.interpreters import batching
@trace("multiply_add_batch")
def multiply_add_batch(vector_arg_values, batch_axes):
"""Computes the batched version of the primitive.
This must be a JAX-traceable function.
Since the multiply_add primitive already operates pointwise on arbitrary
dimension tensors, to batch it we can use the primitive itself. This works as
long as both the inputs have the same dimensions and are batched along the
same axes. The result is batched along the axis that the inputs are batched.
Args:
vector_arg_values: a tuple of two arguments, each being a tensor of matching
shape.
batch_axes: the axes that are being batched. See vmap documentation.
Returns:
a tuple of the result, and the result axis that was batched.
"""
assert batch_axes[0] == batch_axes[1]
assert batch_axes[0] == batch_axes[2]
_trace("Using multiply_add to compute the batch:")
res = multiply_add_prim(*vector_arg_values)
return res, batch_axes[0]
batching.primitive_batchers[multiply_add_p] = multiply_add_batch
# + id="VwxNk869P_YG" outputId="9d22c921-5803-4d33-9e88-b6e439ba9738"
assert np.allclose(api.vmap(square_add_prim, in_axes=0, out_axes=0)(
np.array([2., 3.]),
np.array([10., 20.])),
[14., 29.])
# + [markdown] id="NmqLlV1TQDCC"
# #### JIT of batching
# + id="xqEdXVUgQCTt" outputId="9c22fd9c-919c-491d-bbeb-32c241b808fa"
assert np.allclose(api.jit(api.vmap(square_add_prim, in_axes=0, out_axes=0))
(np.array([2., 3.]),
np.array([10., 20.])),
[14., 29.])
|
docs/notebooks/How_JAX_primitives_work.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="border: 1.5px solid #ccc;
# padding: 8px 12px;
# color:#56BFCB;"
# >
# <center> <br/>
# Lista de Exercícios 5a <br/>
# <span style="font-size:18px;"> <NAME> </span>
# </center>
# </h1>
# ---
# <b>
# <center>
# Imports
# </center>
# </b>
# +
import numpy as np
import pandas as pd
from scipy import optimize as opt
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('seaborn-poster')
import sympy as sp
sp.init_printing()
# -
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>Exercicio 1:</b> Implemente os algoritmos da iteração de ponto fixo, Newton-Raphson e secante usando somente a biblioteca Numpy.
# </div>
# **Funções de teste:**
# +
g_aula = lambda x: np.exp(-x)
g_aula_x0 = 0
fn_youtube = lambda x: np.cos(x) - np.sin(x)
fD_youtube = lambda x: -np.sin(x) - np.cos(x)
er_youtube = 0.01
x0_youtube = 0
x1_youtube = np.pi/2
f_aula = lambda x: np.sin(x) - x
f_aula_x1 = 0.7
f_aula_x0 = 0.8
# -
#
# <div class="alert alert-block alert-info" style="color:Blue;">
# Método da Iteração de Ponto Fixo
# </div>
# + tags=[]
def ponto_fixo(f, x0, maxiter=1000, xtol=1e-10, verbose=True, r_tb=False, p_tb=False):
''' Opcional... '''
if r_tb or p_tb: tb = []
i = 0 # contador de iterações
while i < maxiter:
# enquanto tanto o erro quanto o número de iterações não extrapolarem os limites estabelecidos...
y = f(x0) # calcula f(x0) e atribui a y
# calcula o erro abs
erro = np.abs(x0 - y)
''' Opcional... '''
if r_tb or p_tb: tb.append([]); tb[i].append(x0); tb[i].append(y); tb[i].append(erro); tb[i].append(xtol);
if erro < xtol:
msg = f'Raiz encontrada: {x0} | Em {i} iterações'
if verbose:
print('-'*len(msg))
print(msg)
print('-'*len(msg))
''' Opcional... '''
if p_tb:
print(pd.DataFrame(tb, columns=["x0", "f(x0)", "Erro", "xtol"]))
print('-'*len(msg))
''' ----------- '''
# retorna a raiz e o número de iterações
if r_tb:
return x0, i, tb
else:
return x0, i
x0 = y # atualiza o valor de x0 com o atual y
i += 1 # incrementa contador
if verbose: print("Número de iterações extrapolou o limite sem encontrar a raíz!")
if r_tb: return False, False, tb
return None # saiu do while sem encontrar uma solução
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
x, y, tb = ponto_fixo(g_aula, g_aula_x0, xtol=1e-1, r_tb=True)
pd.DataFrame(tb, columns=["x0", "f(x0)", "Erro", "xtol"])
# -
#
# <div class="alert alert-block alert-info" style="color:Blue;">
# Método Newton-Raphson
# </div>
def newton_raphson(f, fD, x, tol=1e-10, maxiter=500, verbose=False, r_tb=False, p_tb=False):
Xk = x # o x da primeira iteração recebe o x de entrada
k = 0 # contador de iterações
''' Opcional... '''
if r_tb or p_tb: tb = []
while k < maxiter:
# enquanto o número de iterações não extrapolarem o limite estabelecido...
''' Opcional... '''
if r_tb or p_tb: tb.append([]);
f_Xk = f(Xk) # calcula f(x) do x atual (dessa k-ézima iteração)
fD_Xk = fD(Xk) # calcula a derivada f'(x) do x atual (dessa k-ézima iteração)
''' Opcional... '''
if r_tb or p_tb: tb[k].append(Xk); tb[k].append(f_Xk); tb[k].append(fD_Xk);
# se a derivada for 0, não há o que fazer
if fD_Xk == 0:
if verbose: print("Derivada é == 0. Divisão por zero. Sem soluções possíveis!")
return None
# atualiza o valor de Xk+1 (x da próxima k-ézima iteração)
newton_div = f_Xk / fD_Xk
Xk1 = Xk - newton_div
''' Opcional... '''
if r_tb or p_tb: tb[k].append(newton_div); tb[k].append(Xk1);
erro = np.abs( Xk1 - Xk ) # calcula o erro relativo
''' Opcional... '''
if r_tb or p_tb: tb[k].append(erro);
# se o erro for menor ou igual a tolerância, retorna o resultado
if erro <= tol:
msg = f'Raiz encontrada: {Xk1} | Em {k} iterações'
if verbose:
print('-'*len(msg))
print(msg)
print('-'*len(msg))
''' Opcional... '''
if p_tb:
print(pd.DataFrame(tb, columns=["Xk", "f(Xk)", "f(Xk)", "f(Xk)/f'(Xk)", "Xk+1", "Erro"]))
print('-'*len(msg))
''' ----------- '''
# retorna a raiz e o número de iterações
if r_tb:
return Xk1, k, tb
else:
return Xk1, k, False
Xk = Xk1 # atualiza o valor do x para a próxima iteração
k += 1 # incrementa o contador
if verbose: print("Número de iterações extrapolou o limite sem encontrar a raíz!")
if r_tb: return False, False, tb
return None # saiu do while sem encontrar uma solução
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
x, k, tb = newton_raphson(fn_youtube, fD_youtube, x0_youtube, er_youtube, verbose=True, r_tb=True)
pd.DataFrame(tb, columns=["Xk", "f(Xk)", "f(Xk)", "f(Xk)/f'(Xk)", "Xk+1", "Erro"])
# -
#
# <div class="alert alert-block alert-info" style="color:Blue;">
# Método da Secante
# </div>
def secante(f, x0, x1, tol=1e-10, maxiter=500, verbose=True, r_tb=False, p_tb=False):
if f(x0) * f(x1) >= 0:
if verbose: print("Incapaz de prosseguir!")
return None
''' Opcional... '''
if r_tb or p_tb: tb = []
erro = None
k = 0
while k < maxiter:
''' Opcional... '''
# if r_tb or p_tb: tb.append([]);
fX1 = f(x1)
fX0 = f(x0)
# calcula o valor intermediário
Xk = (x0 * fX1 - x1 * fX0) / (fX1 - fX0)
# atualiza valor de x
x = fX0 * f(Xk);
# verifica se x é a raiz da equação, em caso positivo, retorna Xk e o número de iterações
if (x == 0):
if verbose: print(f"Encontrou a raiz {Xk} em {k} iterações!")
return Xk, k
else:
# atualiza os valores do intervalo
x0 = x1;
x1 = Xk;
Xk1 = Xk
erro = abs(Xk1 - Xk)
if erro <= tol:
if verbose: print(f"Encontrou a raiz {Xk} em {k} iterações!")
return Xk1, k # retorna a raiz e o número de iterações
# update number of iteration
k += 1
if verbose: print(f"Número de iterações extrapolou o limite sem encontrar a raíz! Valor final foi {Xk1}")
return None # saiu do while sem encontrar uma soluçãoint("Número de iterações extrapolou o limite sem encontrar a raíz!")
secante(fn_youtube, x0_youtube, x1_youtube, 0.01, verbose=True)
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>Exercicio 2:</b> Use a iteração de ponto fixo simples para localizar a raiz de $f(x)=2 sin(\sqrt{x})−x$, tendo $x_0 = 0,5$ e adotando como critério de parada o erro $e_a ≤ 0,001\%$.
# </div>
ex2_f = lambda x: ( 2 * np.sin(np.sqrt(x)) - x )
ex2_x0 = 0.5
ex2_tol = 0.001
ponto_fixo(ex2_f, ex2_x0, xtol=ex2_tol)
opt.fixed_point(ex2_f, ex2_x0, xtol=ex2_tol, maxiter=3)
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>Exercicio 3:</b> Determine a maior raiz real de $f(x)=2x^3 − 11.7x^2 + 17.7x − 5$.
# </div>
# +
# Definindo f(x)
ex3_f = lambda x: 2 * x**3 - 11.7 * x**2 + 17.7 * x - 5
sp.var('x')
sp.Lambda(x, ex3_f(x)) # 'printando' a função de forma simbólica
# +
# ex3_fD = lambda x: 2*x**3 - 11.7*x**2 + 17.7*x - 5
sp.var('x')
# Calculando f'(x)
ex3_fD_sym = lambda x: eval(sp.ccode(sp.diff(ex3_f(x), x)))
sp.Lambda(x, ex3_fD_sym(x)) # 'printando' a função de forma simbólica
# -
ex3_fD = lambda x: 6*x**2 - 23.4*x + 17.7
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>a)</b> Graficamente;
# </div>
# +
x = np.linspace(0, 4, 50)
y = ex3_f(x)
raiz1 = opt.root(ex3_f, 0)
raiz2 = opt.root(ex3_f, 2)
raiz3 = opt.root(ex3_f, 4)
raizes = np.array([raiz1.x, raiz2.x, raiz3.x])
fig = plt.figure(figsize=(10, 8))
ax = fig.add_subplot(111)
plt.vlines(x=raiz1.x, ymin=ex3_f(0), ymax=0, colors='gray', ls=':', lw=2)
plt.vlines(x=raiz2.x, ymin=ex3_f(0), ymax=0, colors='gray', ls=':', lw=2)
plt.vlines(x=raiz3.x, ymin=ex3_f(0), ymax=0, colors='gray', ls=':', lw=2)
ax.axhline(0, color='b')
ax.plot(x, ex3_f(x), 'r', label="$f(x)$")
ax.plot(raizes, ex3_f(raizes), 'kv', label="$Raizes$")
ax.legend(loc='best')
ax.set_xlabel('$x$')
ax.set_ylabel('$f(x)$')
plt.show()
# -
# Analisando o gráfico plotado, podemos notar que a maior raíz real está entre 3.5 e 4.0.
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>b)</b> Pelo método da iteração de ponto fixo (três iterações, $x_0=3$) (certifique-se de desenvolver uma solução que convirja para a raiz);
# </div>
ponto_fixo(ex3_f, 3, 3)
try: opt.fixed_point(ex3_f, 3, maxiter=3)
except RuntimeError as re: print(str(re))
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>c)</b> Pelo método de Newton-Raphson (três iterações, $x_0=3$);
# </div>
opt.root_scalar(ex3_f, fprime=ex3_fD, x0=3, maxiter=3, method='newton')
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>d)</b> Pelo método da secante (três iterações, $x_{−1}=3$, $x_0=4$).
# </div>
secante(ex3_f, 3, 4, 3)
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>Exercicio 4:</b> Compare os métodos da bisseção, falsa posição, do ponto fixo, de Newton-Raphson e da secante, localizando a raiz das seguintes equações:
# </div>
# <div class="alert alert-block alert-warning">
# <p><b>Para as avaliações, deve-se considerar:</b></p>
# <ul>
# <li>o número máximo de iterações de todos os métodos testados não pode ultrapassar 200;</li>
# <li>a tolerância deve ser de $10^{-10}$;</li>
# <li>para os métodos abertos, escolha os limites do intervalo, respectivamente como $x_{?1}$ e $x_0$.</li>
# </ul>
# <p><b>Para cada método, estamos interessados em comparar:</b></p>
# <ul>
# <li>raiz;</li>
# <li>número de iterações até o critério de parada;</li>
# <li>se houve erro de convergência;</li>
# <li>tempo de cálculo (procure como calcular tempo de execução usando jupyter notebooks, como %timeit).</li>
# </ul>
# </div>
# Constantes
ex4_maxit = 200
ex4_tol = 1e-10
# ---
# Método da Falsa Posição:
# + tags=[]
def regula_falsi(f, xl, xu, tol=1e-10, maxit=10000):
if (f(xl) * f(xu) >= 0):
return -1
i = 0
x = xl
erro, x_ant = 1, x
while erro > tol and i < maxit:
x = xu - ( ( f(xu)*(xl-xu) ) / (f(xl)-f(xu)) )
if f(x) * f(xl) < 0:
xu = x
else:
xl = x
erro = np.abs((x - x_ant) / np.abs(x))
x_ant = x
i += 1
return ( x, i )
# -
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>a)</b> $f_1(x) = 2x^4 + 4x^3 + 3x^2 - 10x - 15$, com $x^* \in [0, 3]$
# </div>
# **Definindo $f_1(x)$**
sp.var('x')
f1 = lambda x: 2*x**4 + 4*x**3 + 3*x**2 - 10*x - 15
sp.Lambda(x, f1(x)) # 'printando' a função simbolicamente
# **Calculando $f_1'(x)$**
sp.var('x')
f1D = lambda x: eval(sp.ccode(sp.diff(f1(x), x)))
sp.Lambda(x, f1D(x)) # 'printando' a função simbolicamente
f1D_2 = lambda x: 8*x**3 + 12*x**2 + 6*x - 10
# **Refatorando $f_1'(x)$**
sp.var('x')
f1_ref = lambda x: 15 / (2*(x**3) + 4*(x**2) + 3*(x) - 10)
sp.Lambda(x, f1_ref(x)) # 'printando' a função simbolicamente
# **Limites do Intervalo**
f1_x0 = 0
f1_x1 = 3
#
# **$f_1(x)$ - Bisseção**
# %timeit opt.root_scalar(f1, method= 'bisect', bracket=[f1_x0, f1_x1], rtol=ex4_tol)
opt.root_scalar(f1, method= 'bisect', bracket=[f1_x0, f1_x1], rtol=ex4_tol)
#
# <p style="font-weight: bold">$f_1(x)$ - Falsa Posição</p>
# %timeit regula_falsi(f1, f1_x0, f1_x1, ex4_tol, ex4_maxit)
regula_falsi(f1, f1_x0, f1_x1, ex4_tol, ex4_maxit)
#
# <p><b>$f_1(x)$ - Ponto Fixo</b></p>
# + tags=[]
# %timeit opt.fixed_point(f1_ref, 1.5, xtol=ex4_tol, maxiter=ex4_maxit)
opt.fixed_point(f1_ref, 1.5, xtol=ex4_tol, maxiter=ex4_maxit)
# -
#
# <p><b>$f_1(x)$ - Newton-Raphson</b></p>
# + tags=[]
# %timeit opt.root_scalar(f1, fprime=f1D_2, x0=1, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
opt.root_scalar(f1, fprime=f1D_2, x0=1, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
# -
#
# <p><b>$f_1(x)$ - Secante</b></p>
# %timeit opt.root_scalar(f1, x0=1, x1=f1_x1, method='secant')
opt.root_scalar(f1, x0=1, x1=f1_x1, method='secant')
#
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>b)</b> $f_2(x) = (x + 3)(x + 1)(x - 2)^3$, com $x^* \in [0,5]$
# </div>
# **Definindo $f_2(x)$**
sp.var('x')
f2 = lambda x: (x + 3)*(x + 1)*(x - 2)**3
sp.Lambda(x, f2(x)) # 'printando' a função simbolicamente
# **Refatorando $f_2(x)$**
sp.var('x')
f2R = lambda x: 24 / ( (x**4) - 2*(x**3) - 9*(x**2) + 22*x + 4 )
sp.Lambda(x, f2R(x)) # vendo o resultado simbólico
# **Calculando $f_2'(x)$**
sp.var('x')
f2D_sym = lambda x: eval(sp.ccode(sp.diff(f2(x), x)))
sp.Lambda(x, f2D_sym(x)) # 'printando' a função simbolicamente
f2D = lambda x: (x - 2)**3 * (x + 1) + (x - 2)**3 * (x + 3) + 3*(x - 2)**2 * (x + 1) * (x + 3)
# **Limites do Intervalo**
f2_x0 = 0
f2_x1 = 5
#
# **$f_2(x)$ - Bisseção**
# %timeit opt.root_scalar(f2, method= 'bisect', bracket=[f2_x0, f2_x1], rtol=ex4_tol)
opt.root_scalar(f2, method= 'bisect', bracket=[f2_x0, f2_x1], rtol=ex4_tol)
#
# **$f_2(x)$ - Falsa Posição**
# %timeit regula_falsi(f2, f2_x0, f2_x1, ex4_tol, ex4_maxit)
regula_falsi(f2, f2_x0, f2_x1, ex4_tol, ex4_maxit)
#
# **$f_2(x)$ - Ponto Fixo**
# + tags=[]
# %timeit opt.fixed_point(f2R, x0=1, xtol=ex4_tol, maxiter=ex4_maxit)
opt.fixed_point(f2R, x0=1, xtol=ex4_tol, maxiter=ex4_maxit)
# -
#
# **$f_2(x)$ - Newton-Raphson**
# + tags=[]
# %timeit opt.root_scalar(f2, fprime=f2D, x0=1, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
opt.root_scalar(f2, fprime=f2D, x0=1, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
# -
#
# **$f_2(x)$ - Secante**
# %timeit opt.root_scalar(f2, x0=f2_x0, x1=f2_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
opt.root_scalar(f2, x0=f2_x0, x1=f2_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
#
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>c)</b> $f_3(x) = 5x^3 + x^2 - e^{1-2x} + cos(x) + 20$, com $x^* \in [-5, 5]$
# </div>
# **Definindo $f_3(x)$**
# +
sp.var('x')
def f3(x, t=None):
if t == 'sp':
return (5*x**3 + x**2 - sp.exp(1-2*x) + sp.cos(x) + 20)
return (5*x**3 + x**2 - np.exp(1-2*x) + np.cos(x) + 20)
sp.Lambda(x, f3(x, 'sp')) # 'printando' a função simbolicamente
# -
# **Refatorando $f_3(x)$**
# +
sp.var('x')
def f3R(x, t=None):
if t == 'sp':
return ( 5*x**3 + x**2 - sp.exp(1-2*x) + sp.cos(x) + 20 + x )
return ( 5*x**3 + x**2 - np.exp(1-2*x) + np.cos(x) + 20 + x )
sp.Lambda(x, f3R(x, 'sp')) # vendo o resultado simbólico
# -
# **Calculando $f_3'(x)$**
# +
sp.var('x')
f3D_sym = lambda x: sp.diff(f3(x, 'sp'), x) # calcula a derivada
sp.Lambda(x, f3D_sym(x)) # vendo o resultado simbólico
# +
# aplicando o resultado
def f3D(x, t=None):
if t == 'sp':
return (15*x**2 + 2*x + 2*sp.exp(1-2*x) - sp.sin(x))
return (15*x**2 + 2*x + 2*np.exp(1-2*x) - np.sin(x))
sp.Lambda(x, f3D(x, 'sp')) # 'printando' a função simbolicamente
# -
# **Limites do Intervalo**
f3_x0 = -5
f3_x1 = 5
#
# **$f_3(x)$ - Bisseção**
# %timeit opt.root_scalar(f3, method= 'bisect', bracket=[f3_x0, f3_x1], rtol=ex4_tol)
opt.root_scalar(f3, method= 'bisect', bracket=[f3_x0, f3_x1], rtol=ex4_tol)
#
# **$f_3(x)$ - Falsa Posição**
# %timeit regula_falsi(f3, f3_x0, f3_x1, ex4_tol, ex4_maxit)
regula_falsi(f3, f3_x0, f3_x1, ex4_tol, ex4_maxit)
#
# **$f_3(x)$ - Ponto Fixo**
# + tags=[]
# %timeit opt.fixed_point(f3R, x0=1, xtol=ex4_tol, maxiter=ex4_maxit)
opt.fixed_point(f3R, x0=1, xtol=ex4_tol, maxiter=ex4_maxit)
# -
#
# **$f_3(x)$ - Newton-Raphson**
# + tags=[]
# %timeit opt.root_scalar(f3, fprime=f3D, x0=1, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
opt.root_scalar(f3, fprime=f3D, x0=1, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
# -
#
# **$f_3(x)$ - Secante**
# %timeit opt.root_scalar(f3, x0=f3_x0, x1=f3_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
opt.root_scalar(f3, x0=f3_x0, x1=f3_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
#
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>d)</b> $f_4(x) = sin(x)x + 4$, com $x^* \in [1, 5]$
# </div>
# **Definindo $f_4(x)$**
# +
sp.var('x')
def f4(x, t=None):
if t == 'sp':
return (sp.sin(x)*x + 4)
return (np.sin(x)*x + 4)
sp.Lambda(x, f4(x, 'sp')) # 'printando' a função simbolicamente
# -
# **Refatorando $f_4(x)$**
# +
sp.var('x')
def f4R(x, t=None):
if t == 'sp':
return ( (-4) / sp.sin(x) )
return ( (-4) / np.sin(x) )
sp.Lambda(x, f4R(x, 'sp')) # vendo o resultado simbólico
# -
# **Calculando $f_4'(x)$**
# +
sp.var('x')
f4D_sym = lambda x: sp.diff(f4(x, 'sp'), x) # calcula a derivada
sp.Lambda(x, f4D_sym(x)) # vendo o resultado simbólico
# +
# aplicando o resultado
def f4D(x, t=None):
if t == 'sp':
return (x * sp.cos(x) + sp.sin(x))
return (x * np.cos(x) + np.sin(x))
sp.Lambda(x, f4D(x, 'sp')) # 'printando' a função simbolicamente
# -
# **Limites do Intervalo**
f4_x1 = 1
f4_x0 = 5
#
# **$f_4(x)$ - Bisseção**
# %timeit opt.root_scalar(f4, method= 'bisect', bracket=[f4_x0, f4_x1], rtol=ex4_tol)
opt.root_scalar(f4, method= 'bisect', bracket=[f4_x0, f4_x1], rtol=ex4_tol)
#
# **$f_4(x)$ - Falsa Posição**
# %timeit regula_falsi(f4, f4_x0, f4_x1, ex4_tol, ex4_maxit)
regula_falsi(f4, f4_x0, f4_x1, ex4_tol, ex4_maxit)
#
# **$f_4(x)$ - Ponto Fixo**
# + tags=[]
# %timeit opt.fixed_point(f4R, x0=f4_x0, xtol=ex4_tol, maxiter=ex4_maxit)
opt.fixed_point(f4R, x0=f4_x0, xtol=ex4_tol, maxiter=ex4_maxit)
# -
#
# **$f_4(x)$ - Newton-Raphson**
# + tags=[]
# %timeit opt.root_scalar(f4, fprime=f4D, x0=4, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
opt.root_scalar(f4, fprime=f4D, x0=4, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
# -
#
# **$f_4(x)$ - Secante**
# %timeit opt.root_scalar(f4, x0=f4_x0, x1=f4_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
opt.root_scalar(f4, x0=f4_x0, x1=f4_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
#
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>e)</b> $f_5(x) = (x - 3)^5 ln(x)$, com $x^* \in [2, 5]$
# </div>
# **Definindo $f_5(x)$**
# +
sp.var('x')
def f5(x, t=None):
if t == 'sp':
return ( (x - 3)**5 * sp.ln(x) )
return ( (x - 3)**5 * np.log(x) )
sp.Lambda(x, f5(x, 'sp')) # 'printando' a função simbolicamente
# -
# **Refatorando $f_5(x)$**
# +
sp.var('x')
def f5R(x, t=None):
if t == 'sp':
return ( (x-3)**5 * sp.log(x) + x )
return ( (x-3)**5 * np.log(x) + x )
sp.Lambda(x, f5R(x, 'sp')) # vendo o resultado simbólico
# -
# **Calculando $f_5'(x)$**
# +
sp.var('x')
f5D_sym = lambda x: sp.diff(f5(x, 'sp'), x) # calcula a derivada
sp.Lambda(x, f5D_sym(x)) # vendo o resultado simbólico
# +
# aplicando o resultado
def f5D(x, t=None):
if t == 'sp':
return ( 5*(x - 3)**4 * sp.log(x) + sp.Pow((x-3),5) / x )
return ( 5*(x - 3)**4 * np.log(x) + ((x-3)**5 / x) )
sp.Lambda(x, f5D(x, 'sp')) # 'printando' a função simbolicamente
# -
# **Limites do Intervalo**
f5_x0 = 2
f5_x1 = 5
#
# **$f_5(x)$ - Bisseção**
# %timeit opt.root_scalar(f5, method= 'bisect', bracket=[f5_x0, f5_x1], rtol=ex4_tol)
opt.root_scalar(f5, method= 'bisect', bracket=[f5_x0, f5_x1], rtol=ex4_tol)
#
# **$f_5(x)$ - Falsa Posição**
# %timeit regula_falsi(f5, f5_x0, f5_x1, ex4_tol, ex4_maxit)
regula_falsi(f5, f5_x0, f5_x1, ex4_tol, ex4_maxit)
#
# **$f_5(x)$ - Ponto Fixo**
# + tags=[]
# %timeit opt.fixed_point(f5R, x0=2, xtol=10**-9.9, maxiter=ex4_maxit)
opt.fixed_point(f5R, x0=2, xtol=10**-9.9, maxiter=ex4_maxit)
# -
# **$f_5(x)$ - Newton-Raphson**
# + tags=[]
# %timeit opt.root_scalar(f5, fprime=f5D, x0=f5_x0, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
opt.root_scalar(f5, fprime=f5D, x0=f5_x0, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
# -
#
# **$f_5(x)$ - Secante**
# %timeit opt.root_scalar(f5, x0=f5_x0, x1=f5_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
opt.root_scalar(f5, x0=f5_x0, x1=f5_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
#
# ---
# <div class="alert alert-block alert-info" style="color:#20484d;">
# <b>f)</b> $f_6(x) = x^{10} - 1$, com $x^* \in [0.8, 1.2]$
# </div>
# **Definindo $f_6(x)$**
# +
sp.var('x')
f6 = lambda x: x**10 - 1
sp.Lambda(x, f6(x)) # 'printando' a função simbolicamente
# -
# **Refatorando $f_6(x)$**
# +
sp.var('x')
f6R = lambda x: 1/(x)**9
sp.Lambda(x, f6R(x)) # vendo o resultado simbólico
# -
# **Calculando $f_6'(x)$**
# +
sp.var('x')
f6D_sym = lambda x: eval(sp.ccode(sp.diff(f6(x), x)))
sp.Lambda(x, f6D_sym(x)) # 'printando' a função simbolicamente
# -
f6D = lambda x: 10 * x**9
# **Limites do Intervalo**
f6_x0 = 0.8
f6_x1 = 1.2
#
# **$f_6(x)$ - Bisseção**
# %timeit opt.root_scalar(f6, method= 'bisect', bracket=[f6_x0, f6_x1], rtol=ex4_tol)
opt.root_scalar(f6, method= 'bisect', bracket=[f6_x0, f6_x1], rtol=ex4_tol)
#
# **$f_6(x)$ - Falsa Posição**
# %timeit regula_falsi(f6, f6_x0, f6_x1, ex4_tol, ex4_maxit)
regula_falsi(f6, f6_x0, f6_x1, ex4_tol, ex4_maxit)
#
# **$f_6(x)$ - Ponto Fixo**
# + tags=[]
# %timeit opt.fixed_point(f6R, x0=1, xtol=ex4_tol, maxiter=ex4_maxit)
opt.fixed_point(f6R, x0=1, xtol=ex4_tol, maxiter=ex4_maxit)
# -
#
# **$f_6(x)$ - Newton-Raphson**
# + tags=[]
# %timeit opt.root_scalar(f6, fprime=f6D, x0=f6_x0, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
opt.root_scalar(f6, fprime=f6D, x0=f6_x0, xtol=ex4_tol, maxiter=ex4_maxit, method='newton')
# -
#
# **$f_6(x)$ - Secante**
# %timeit opt.root_scalar(f6, x0=f6_x0, x1=f6_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
opt.root_scalar(f6, x0=f6_x0, x1=f6_x1, maxiter=ex4_maxit, xtol=ex4_tol, method='secant')
#
# ---
|
Class 05/Lista_5a - Guilherme Esdras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Figure S7A
# +
# Preliminaries to work with the data.
# %matplotlib inline
# %run __init__.py
from utils import loading, scoring, prog
from gerkin import dream,params,fit2
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import matplotlib as mpl
mpl.rcParams.update({'font.size':14})
# -
# Load the data
descriptors = loading.get_descriptors(format='True')
all_CIDs = loading.get_CIDs(['training','leaderboard','testset'])
testset_CIDs = loading.get_CIDs(['testset'])
all_CID_dilutions = loading.get_CID_dilutions(['training','leaderboard','testset'])
#mdx_full = dream.get_molecular_data(['dragon','episuite','morgan','nspdk','gramian'],all_CIDs)
features = loading.get_molecular_data(['dragon','morgan'],all_CIDs)
# Create the feature and descriptor arrays
X,_,_,_,_,_ = dream.make_X(features,all_CID_dilutions)
X_train = X.drop(testset_CIDs)
X_test = X.drop(X_train.index)
# Load and split perceptual data
Y_train = loading.load_perceptual_data(['training','leaderboard'])
Y_train = Y_train.groupby(level=['Descriptor','CID','Dilution']).mean() # Average over replicates
Y_test = loading.load_perceptual_data('testset')
# ### Load or compute the random forest model
# +
from sklearn.model_selection import ShuffleSplit
from sklearn.ensemble import RandomForestRegressor
n_subjects = 49
n_splits = 25
trans_params = params.get_trans_params(Y_train, descriptors, plot=False)
use_et, max_features, max_depth, min_samples_leaf, trans_weight, regularize, use_mask = params.get_other_params()
ss = ShuffleSplit(n_splits=n_splits,test_size=(24./49),random_state=0)
rs_in = pd.DataFrame(index=range(n_splits),columns=descriptors) # Same subject, different molecules correlations.
rs_out = pd.DataFrame(index=range(n_splits),columns=descriptors) # Different subject, different molecule correlations.
for d,descriptor in enumerate(descriptors):
print("%d) %s" % (d,descriptor))
rfc = RandomForestRegressor(n_estimators=30, max_features='auto', random_state=0)
for i,(train,test) in enumerate(ss.split(range(n_subjects))):
prog(i,n_splits)
train+=1; test+=1; # Subjects are 1-indexed.
rfc.fit(X_train,Y_train['Subject'][train].mean(axis=1).loc[descriptor])
Y_test_in = Y_test['Subject'][train].mean(axis=1).loc[descriptor]
Y_test_out = Y_test['Subject'][test].mean(axis=1).loc[descriptor]
Y_predicted = rfc.predict(X_test)
rs_in.loc[i,descriptor] = np.corrcoef(Y_predicted,Y_test_in)[0,1]
rs_out.loc[i,descriptor] = np.corrcoef(Y_predicted,Y_test_out)[0,1]
# +
# 25 x 30
fig,axes = plt.subplots(2,2,figsize=(10,10))
ax = axes.flat
ax[0].errorbar(range(len(descriptors)),rs_in.mean(),yerr=rs_in.sem(),
color='k',fmt='o-',label='Same %d subjects' % 25)
ax[0].errorbar(range(len(descriptors)),rs_out.mean(),yerr=rs_out.sem(),
color='r',fmt='o-',label='Different %d subjects' % 24)
order = rs_in.mean().sort_values()[::-1].index
ax[1].errorbar(range(len(descriptors)),rs_in.mean()[order],yerr=rs_in.sem()[order],
color='k',fmt='o-',label='Same %d subjects' % 25)
ax[1].errorbar(range(len(descriptors)),rs_out.mean()[order],yerr=rs_out.sem()[order],
color='r',fmt='o-',label='Different %d subjects' % 24)
for i in [0,1]:
ax[i].set_xlim(-0.5,len(descriptors)-0.5)
ax[i].set_ylim(0,0.82)
ax[i].set_xticklabels(order,rotation=90);
ax[i].set_ylabel('Correlation')
ax[i].legend(fontsize=10)
ax[2].errorbar(rs_in.mean(),rs_out.mean(),
xerr=rs_in.sem(),yerr=rs_out.sem(),
color='k',fmt='o')
ax[2].plot([0,1],[0,1],'--')
ax[2].set_xlim(0,0.82)
ax[2].set_xlabel('Correlation\n(Same 25 subjects)')
ax[2].set_ylabel('Correlation\n(Different 25 subjects)')
order = (rs_in-rs_out).mean().sort_values()[::-1].index
ax[3].errorbar(range(len(descriptors)),(rs_in-rs_out).mean()[order],
yerr=(rs_in-rs_out).sem()[order],
color='k',fmt='o-')
ax[3].plot([0,len(descriptors)],[0,0],'--')
ax[3].set_xlim(-0.5,len(descriptors)-0.5)
ax[3].set_ylim(-0.05,0.1)
ax[3].set_xticklabels(order,rotation=90);
ax[3].set_ylabel('Correlation Difference')
plt.tight_layout()
plt.savefig('../../figures/subject-splits.eps',format='eps')
# -
print('%.3f +/- %.3f, with maximum value %.3f' % \
((rs_in-rs_out).mean().mean(),(rs_in-rs_out).mean().std(),(rs_in-rs_out).mean().max()))
from scipy.stats import ttest_rel,chi2
# No FDR correction
chi2_ = 0
for d,descriptor in enumerate(descriptors):
p = ttest_rel(rs_in[descriptor],rs_out[descriptor])[1]
chi2_ += -2*np.log(p)
print('%s%.3f' % ((descriptor+':').ljust(15),p))
p_pooled = 1-chi2.cdf(chi2_,2*len(descriptors))
print("Pooled p-value = %.3g" % p_pooled)
|
opc_python/paper/subject-splits.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torchgeometry as tgm
import cv2
# +
# read the image with OpenCV
image = cv2.imread('./data/bruce.png')[..., (2,1,0)]
print(image.shape)
img = tgm.image_to_tensor(image)
img = torch.unsqueeze(img.float(), dim=0) # BxCxHxW
# +
# the source points are the region to crop corners
points_src = torch.FloatTensor([[
[125, 150], [562, 40], [562, 282], [54, 328],
]])
# the destination points are the image vertexes
h, w = 64, 128 # destination size
points_dst = torch.FloatTensor([[
[0, 0], [w - 1, 0], [w - 1, h - 1], [0, h - 1],
]])
# compute perspective transform
M = tgm.get_perspective_transform(points_src, points_dst)
# warp the original image by the found transform
img_warp = tgm.warp_perspective(img, M, dsize=(h, w))
# convert back to numpy
image_warp = tgm.tensor_to_image(img_warp.byte())
# draw points into original image
for i in range(4):
center = tuple(points_src[0, i].long().numpy())
image = cv2.circle(image.copy(), center, 5, (0, 255, 0), -1)
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# create the plot
fig, axs = plt.subplots(1, 2, figsize=(16, 10))
axs = axs.ravel()
axs[0].axis('off')
axs[0].set_title('image source')
axs[0].imshow(image)
axs[1].axis('off')
axs[1].set_title('image destination')
axs[1].imshow(image_warp)
|
examples/warp_perspective.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from subprocess import call
from glob import glob
from nltk.corpus import stopwords
import os, struct
from tensorflow.core.example import example_pb2
import pyrouge
import shutil
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
from nltk.stem.porter import *
# +
ratio = 1
duc_num = 6
max_len = 250
#cmd = '/root/miniconda2/bin/python ../pointer-generator-master/run_summarization.py --mode=decode --single_pass=1 --coverage=True --vocab_path=finished_files/vocab --log_root=log --exp_name=myexperiment --data_path=test/temp_file'
#cmd = '/root/miniconda2/bin/python run_summarization.py --mode=decode --single_pass=1 --coverage=True --vocab_path=finished_files/vocab --log_root=log --exp_name=myexperiment --data_path=test/temp_file --max_enc_steps=4000'
#cmd = cmd.split()
#generated_path = '/gttp/pointer-generator-master/log/myexperiment/decode_test_4000maxenc_4beam_35mindec_120maxdec_ckpt-238410/'
#generated_path = '/gttp/pointer-generator-tal/log/myexperiment/decode_test_4000maxenc_4beam_35mindec_100maxdec_ckpt-238410/'
vocab_path = '../data/DMQA/finished_files/vocab'
log_root = 'log'
exp_name = 'myexperiment'
data_path= 'test/temp_file'
max_enc_steps = 4000
cmd = ['python',
'run_summarization.py',
'--mode=decode',
'--single_pass=1',
'--coverage=True',
'--vocab_path=' + vocab_path,
'--log_root=' + log_root,
'--exp_name=' + exp_name,
'--data_path=' + data_path,
'--max_enc_steps=' + str(max_enc_steps)]
generated_path = 'log/myexperiment/decode_test_4000maxenc_4beam_35mindec_100maxdec_ckpt-238410/'
stopwords = set(stopwords.words('english'))
stemmer = PorterStemmer()
# +
def pp(string):
return ' '.join([stemmer.stem(word.decode('utf8')) for word in string.lower().split() if not word in stopwords])
def write_to_file(article, abstract, rel, writer):
abstract = '<s> '+' '.join(abstract)+' </s>'
#abstract = abstract.encode('utf8', 'ignore')
#rel = rel.encode('utf8', 'ignore')
#article = article.encode('utf8', 'ignore')
tf_example = example_pb2.Example()
tf_example.features.feature['abstract'].bytes_list.value.extend([bytes(abstract)])
tf_example.features.feature['relevancy'].bytes_list.value.extend([bytes(rel)])
tf_example.features.feature['article'].bytes_list.value.extend([bytes(article)])
tf_example_str = tf_example.SerializeToString()
str_len = len(tf_example_str)
writer.write(struct.pack('q', str_len))
writer.write(struct.pack('%ds' % str_len, tf_example_str))
def duck_iterator(i):
duc_folder = 'duc0' + str(i) + 'tokenized/'
for topic in os.listdir(duc_folder + 'testdata/docs/'):
topic_folder = duc_folder + 'testdata/docs/' + topic
if not os.path.isdir(topic_folder):
continue
query = ' '.join(open(duc_folder + 'queries/' + topic).readlines())
model_files = glob(duc_folder + 'models/' + topic[:-1].upper() + '.*')
topic_texts = [' '.join(open(topic_folder + '/' + file).readlines()).replace('\n', '') for file in
os.listdir(topic_folder)]
abstracts = [' '.join(open(f).readlines()) for f in model_files]
yield topic_texts, abstracts, query
def ones(sent, ref): return 1.
def count_score(sent, ref):
ref = pp(ref).split()
sent = ' '.join(pp(w) for w in sent.lower().split() if not w in stopwords)
return sum([1. if w in ref else 0. for w in sent.split()])
def get_w2v_score_func(magic = 10):
import gensim
google = gensim.models.KeyedVectors.load_word2vec_format(
'GoogleNews-vectors-negative300.bin', binary=True)
def w2v_score(sent, ref):
ref = ref.lower()
sent = sent.lower()
sent = [w for w in sent.split() if w in google]
ref = [w for w in ref.split() if w in google]
try:
score = google.n_similarity(sent, ref)
except:
score = 0.
return score * magic
return w2v_score
def get_tfidf_score_func_glob(magic = 1):
corpus = []
for i in range(5, 8):
for topic_texts, _, _ in duck_iterator(i):
corpus += [pp(t) for t in topic_texts]
vectorizer = TfidfVectorizer()
vectorizer.fit_transform(corpus)
def tfidf_score_func(sent, ref):
#ref = [pp(s) for s in ref.split(' . ')]
sent = pp(sent)
v1 = vectorizer.transform([sent])
#v2s = [vectorizer.transform([r]) for r in ref]
#return max([cosine_similarity(v1, v2)[0][0] for v2 in v2s])
v2 = vectorizer.transform([ref])
return cosine_similarity(v1, v2)[0][0]
return tfidf_score_func
# +
def get_tfidf_score_func(magic = 10):
corpus = []
for i in range(5, 8):
for topic_texts, _, _ in duck_iterator(i):
corpus += [t.lower() for t in topic_texts]
vectorizer = TfidfVectorizer()
vectorizer.fit_transform(corpus)
def tfidf_score_func(sent, ref):
ref = ref.lower()
sent = sent.lower()
v1 = vectorizer.transform([sent])
v2 = vectorizer.transform([ref])
return cosine_similarity(v1, v2)[0][0]*magic
return tfidf_score_func
def just_relevant(text, query):
text = text.split(' . ')
score_per_sent = [count_score(sent, query) for sent in text]
sents_gold = list(zip(*sorted(zip(score_per_sent, text), reverse=True)))[1]
sents_gold = sents_gold[:int(len(sents_gold)*ratio)]
filtered_sents = []
for s in text:
if not s: continue
if s in sents_gold: filtered_sents.append(s)
return ' . '.join(filtered_sents)
class Summary:
def __init__(self, texts, abstracts, query):
#texts = sorted([(tfidf_score(query, text), text) for text in texts], reverse=True)
#texts = sorted([(tfidf_score(text, ' '.join(abstracts)), text) for text in texts], reverse=True)
#texts = [text[1] for text in texts]
self.texts = texts
self.abstracts = abstracts
self.query = query
self.summary = []
self.words = set()
self.length = 0
def add_sum(self, summ):
for sent in summ:
self.summary.append(sent)
def get(self):
text = max([(len(t.split()), t) for t in self.texts])[1]
#text = texts[0]
if ratio < 1: text = just_relevant(text, self.query)
sents = text.split(' . ')
score_per_sent = [(score_func(sent, self.query), sent) for sent in sents]
#score_per_sent = [(count_score(sent, ' '.join(self.abstracts)), sent) for sent in sents]
scores = []
for score, sent in score_per_sent:
scores += [score] * (len(sent.split()) + 1)
scores = str(scores[:-1])
return text, 'a', scores
def get_summaries(path):
path = path+'decoded/'
out = {}
for file_name in os.listdir(path):
index = int(file_name.split('_')[0])
out[index] = open(path+file_name).readlines()
return out
def rouge_eval(ref_dir, dec_dir):
"""Evaluate the files in ref_dir and dec_dir with pyrouge, returning results_dict"""
r = pyrouge.Rouge155()
r.model_filename_pattern = '#ID#_reference_(\d+).txt'
r.system_filename_pattern = '(\d+)_decoded.txt'
r.model_dir = ref_dir
r.system_dir = dec_dir
return r.convert_and_evaluate()
def evaluate(summaries):
for path in ['eval/ref', 'eval/dec']:
if os.path.exists(path): shutil.rmtree(path, True)
os.mkdir(path)
for i, summ in enumerate(summaries):
for j,abs in enumerate(summ.abstracts):
with open('eval/ref/'+str(i)+'_reference_'+str(j)+'.txt', 'w') as f:
f.write(abs)
with open('eval/dec/'+str(i)+'_decoded.txt', 'w') as f:
f.write(' '.join(summ.summary))
print rouge_eval('eval/ref/', 'eval/dec/')
# +
score_func = get_w2v_score_func()
summaries = [Summary(texts, abstracts, query) for texts, abstracts, query in duck_iterator(duc_num)]
with open('test/temp_file', 'wb') as writer:
for summ in summaries:
article, abstract, scores = summ.get()
write_to_file(article, abstracts, scores, writer)
call(['rm', '-r', generated_path])
call(cmd)
generated_summaries = get_summaries(generated_path)
for i in range(len(summaries)):
summaries[i].add_sum(generated_summaries[i])
# -
evaluate(summaries)
print duc_num
print score_func
|
notebooks/RSA-word2vec-DUC2006.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Whatsapp Messages Processing</h1>
import numpy as np
import pandas as pd
from datetime import date
from stop_words import get_stop_words # python -m pip install stop-words
import string
from nltk.probability import FreqDist
from nltk.tokenize import word_tokenize
import seaborn as sns
import matplotlib.pyplot as plt
# +
# %%time
# Runs in a few (3-4) seconds for a messages file of 44k lines
def get_word_frequencies(file_url, start_date):
# Example message:
# '22-05-19 15:22 - Neal: Hahaha'
# Determine stop words (words to be removed because they are not interesting)
stop_words_nltk = get_stop_words('nl')
stop_words_custom = ['\n','<Media weggelaten>','en']
stop_words = set(stop_words_nltk) | set(stop_words_custom)
messages = [] # This list will contain messages as lists of words
start_to_capture = False # This boolean will determine whether we are past the
# start_date and can start to capture messages.
# Read file line by line
with open(file_url) as file:
for line in file:
# Some messages are just showing that a user has sent an image file; skip these
if ('<Media weggelaten>' in line):
continue
# Extract message contents (split by ':' results in the last elt containing the message)
message = line.split(':')
# Try to extract the message date (quick and dirty)
try:
# Extract message date
message_date = np.array(message[0].split(' ')[0].split('-')).astype(int)
message_date = date(message_date[2]+2000, message_date[1], message_date[0])
except:
continue
# Determine whether we can start to capture the messages
# (test whether we are past the 'start' date)
if (start_to_capture == False and message_date >= start_date):
start_to_capture = True
# Process the contents of the message if past the start date
if (start_to_capture):
# Remove some stuff and split into single words
message_text = ":".join(message[2:]).replace('\n','').replace(',','').split(' ')
# Add the message as a list of words
message_text = [word.lower() for word in message_text if word.lower() not in stop_words and len(word) > 0]
# If the message is still interesting, add it to our list of messages
if (len(message) > 0):
messages.append(message_text)
# Return a dictionary-like object containing words and their frequencies
# Warning: not sorted. Use the method .most_common() to obtain a sorted of the data (see code below))
return FreqDist(word for message in messages for word in message)
# -
# <h3>Obtain frequencies</h3>
# +
file_url = '/Users/jan/Downloads/WhatsApp-chat met Ægir.txt'
start_date = date(2020, 3, 12) # Format: Y,M,D
frequencies_postlockdown = get_word_frequencies(file_url, start_date)
start_date = date(1970,1,1)
frequencies_overall = get_word_frequencies(file_url, start_date)
# -
# <h3>Plot top 10 in lockdown period</h3>
# +
# Post Corona (set filter date in cell above)
top_k = 10
frequencies_common = frequencies_postlockdown.most_common()
frequencies_common = {'words': [word for (word,_) in frequencies_common[:top_k]],
'freqs': [freq for (_,freq) in frequencies_common[:top_k]]}
fig = sns.barplot(x=frequencies_common['words'],y=frequencies_common['freqs'],zorder=4)
fig.set_title('Top 10 meest gebruikte woorden vanaf begin lockdown')
plt.ylabel('count')
plt.xticks(rotation=25)
plt.grid(linestyle=":")
plt.savefig('aegir/post_lockdown.pdf')
# -
# <h4>Save words that were used only once in this period to disk</h4>
# Words that are used only once
once_words = np.array([str(word) for (word,freq) in frequencies_postcorona.most_common() if freq==1])
pd.DataFrame(once_words).to_csv('aegir/once_words_post_lockdown.txt', header=None, index=None)
# <h3>Plot top 10 overall</h3>
# +
# Over all time (set filter date in cell above)
top_k = 10
frequencies_common = frequencies_overall.most_common()
frequencies_common = {'words': [word for (word,_) in frequencies_common[:top_k]],
'freqs': [freq for (_,freq) in frequencies_common[:top_k]]}
fig = sns.barplot(x=frequencies_common['words'],y=frequencies_common['freqs'],zorder=4)
fig.set_title('Top 10 meest gebruikte woorden vanaf begin dispuutswhatsappgroep')
plt.ylabel('count')
plt.xticks(rotation=25)
plt.grid(linestyle=":")
plt.savefig('aegir/overall.pdf')
# -
# <h4>Show words that occur within a certain frequency window</h4>
# Show words that occur within a certain frequency window
min_freq = 1
max_freq = 12
words = np.array([(word,freq) for (word,freq) in frequencies_postcorona.most_common() if freq>=min_freq and freq<=max_freq])
for w,f in words:
print(w,f)
|
assets/docs/whatsapp_export_processor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + dc={"key": "3"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 1. Credit card applications
# <p>Commercial banks receive <em>a lot</em> of applications for credit cards. Many of them get rejected for many reasons, like high loan balances, low income levels, or too many inquiries on an individual's credit report, for example. Manually analyzing these applications is mundane, error-prone, and time-consuming (and time is money!). Luckily, this task can be automated with the power of machine learning and pretty much every commercial bank does so nowadays. In this notebook, we will build an automatic credit card approval predictor using machine learning techniques, just like the real banks do.</p>
# <p><img src="https://assets.datacamp.com/production/project_558/img/credit_card.jpg" alt="Credit card being held in hand"></p>
# <p>We'll use the <a href="http://archive.ics.uci.edu/ml/datasets/credit+approval">Credit Card Approval dataset</a> from the UCI Machine Learning Repository. The structure of this notebook is as follows:</p>
# <ul>
# <li>First, we will start off by loading and viewing the dataset.</li>
# <li>We will see that the dataset has a mixture of both numerical and non-numerical features, that it contains values from different ranges, plus that it contains a number of missing entries.</li>
# <li>We will have to preprocess the dataset to ensure the machine learning model we choose can make good predictions.</li>
# <li>After our data is in good shape, we will do some exploratory data analysis to build our intuitions.</li>
# <li>Finally, we will build a machine learning model that can predict if an individual's application for a credit card will be accepted.</li>
# </ul>
# <p>First, loading and viewing the dataset. We find that since this data is confidential, the contributor of the dataset has anonymized the feature names.</p>
# + dc={"key": "3"} tags=["sample_code"]
# Import pandas
import pandas as pd
# Load dataset
cc_apps = pd.read_csv("datasets/cc_approvals.data", header=None)
# Inspect data
cc_apps.head()
# + dc={"key": "10"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 2. Inspecting the applications
# <p>The output may appear a bit confusing at its first sight, but let's try to figure out the most important features of a credit card application. The features of this dataset have been anonymized to protect the privacy, but <a href="http://rstudio-pubs-static.s3.amazonaws.com/73039_9946de135c0a49daa7a0a9eda4a67a72.html">this blog</a> gives us a pretty good overview of the probable features. The probable features in a typical credit card application are <code>Gender</code>, <code>Age</code>, <code>Debt</code>, <code>Married</code>, <code>BankCustomer</code>, <code>EducationLevel</code>, <code>Ethnicity</code>, <code>YearsEmployed</code>, <code>PriorDefault</code>, <code>Employed</code>, <code>CreditScore</code>, <code>DriversLicense</code>, <code>Citizen</code>, <code>ZipCode</code>, <code>Income</code> and finally the <code>ApprovalStatus</code>. This gives us a pretty good starting point, and we can map these features with respect to the columns in the output. </p>
# <p>As we can see from our first glance at the data, the dataset has a mixture of numerical and non-numerical features. This can be fixed with some preprocessing, but before we do that, let's learn about the dataset a bit more to see if there are other dataset issues that need to be fixed.</p>
# + dc={"key": "10"} tags=["sample_code"]
# Print summary statistics
cc_apps_description = cc_apps.describe()
print(cc_apps_description)
print("\n")
# Print DataFrame information
cc_apps_info = cc_apps.info()
print(cc_apps_info)
print("\n")
# Inspect missing values in the dataset
cc_apps.tail(17)
# + dc={"key": "17"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 3. Handling the missing values (part i)
# <p>We've uncovered some issues that will affect the performance of our machine learning model(s) if they go unchanged:</p>
# <ul>
# <li>Our dataset contains both numeric and non-numeric data (specifically data that are of <code>float64</code>, <code>int64</code> and <code>object</code> types). Specifically, the features 2, 7, 10 and 14 contain numeric values (of types float64, float64, int64 and int64 respectively) and all the other features contain non-numeric values.</li>
# <li>The dataset also contains values from several ranges. Some features have a value range of 0 - 28, some have a range of 2 - 67, and some have a range of 1017 - 100000. Apart from these, we can get useful statistical information (like <code>mean</code>, <code>max</code>, and <code>min</code>) about the features that have numerical values. </li>
# <li>Finally, the dataset has missing values, which we'll take care of in this task. The missing values in the dataset are labeled with '?', which can be seen in the last cell's output.</li>
# </ul>
# <p>Now, let's temporarily replace these missing value question marks with NaN.</p>
# + dc={"key": "17"} tags=["sample_code"]
# Import numpy
import numpy as np
# Inspect missing values in the dataset
print(cc_apps.isnull().values.sum())
# Replace the '?'s with NaN
cc_apps = cc_apps.replace('?', np.nan)
# Inspect the missing values again
cc_apps.tail(17)
# + dc={"key": "24"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 4. Handling the missing values (part ii)
# <p>We replaced all the question marks with NaNs. This is going to help us in the next missing value treatment that we are going to perform.</p>
# <p>An important question that gets raised here is <em>why are we giving so much importance to missing values</em>? Can't they be just ignored? Ignoring missing values can affect the performance of a machine learning model heavily. While ignoring the missing values our machine learning model may miss out on information about the dataset that may be useful for its training. Then, there are many models which cannot handle missing values implicitly such as LDA. </p>
# <p>So, to avoid this problem, we are going to impute the missing values with a strategy called mean imputation.</p>
# + dc={"key": "24"} tags=["sample_code"]
# Impute the missing values with mean imputation
cc_apps.fillna(cc_apps.mean(), inplace=True)
# Count the number of NaNs in the dataset to verify
print(cc_apps.isnull().values.sum())
# + dc={"key": "31"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 5. Handling the missing values (part iii)
# <p>We have successfully taken care of the missing values present in the numeric columns. There are still some missing values to be imputed for columns 0, 1, 3, 4, 5, 6 and 13. All of these columns contain non-numeric data and this why the mean imputation strategy would not work here. This needs a different treatment. </p>
# <p>We are going to impute these missing values with the most frequent values as present in the respective columns. This is <a href="https://www.datacamp.com/community/tutorials/categorical-data">good practice</a> when it comes to imputing missing values for categorical data in general.</p>
# + dc={"key": "31"} tags=["sample_code"]
# Iterate over each column of cc_apps
for col in cc_apps.columns:
# Check if the column is of object type
if cc_apps[col].dtypes == 'object':
# Impute with the most frequent value
cc_apps = cc_apps.fillna(cc_apps[col].value_counts().index[0])
# Count the number of NaNs in the dataset and print the counts to verify
print(cc_apps.isnull().values.sum())
# + dc={"key": "38"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 6. Preprocessing the data (part i)
# <p>The missing values are now successfully handled.</p>
# <p>There is still some minor but essential data preprocessing needed before we proceed towards building our machine learning model. We are going to divide these remaining preprocessing steps into three main tasks:</p>
# <ol>
# <li>Convert the non-numeric data into numeric.</li>
# <li>Split the data into train and test sets. </li>
# <li>Scale the feature values to a uniform range.</li>
# </ol>
# <p>First, we will be converting all the non-numeric values into numeric ones. We do this because not only it results in a faster computation but also many machine learning models (like XGBoost) (and especially the ones developed using scikit-learn) require the data to be in a strictly numeric format. We will do this by using a technique called <a href="http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html">label encoding</a>.</p>
# + dc={"key": "38"} tags=["sample_code"]
# Import LabelEncoder
from sklearn.preprocessing import LabelEncoder
# Instantiate LabelEncoder
le = LabelEncoder()
# Iterate over all the values of each column and extract their dtypes
for col in cc_apps.columns:
# Compare if the dtype is object
if cc_apps[col].dtypes=='object':
# Use LabelEncoder to do the numeric transformation
cc_apps[col]=le.fit_transform(cc_apps[col])
# As we can see all features are of numeric type now
cc_apps.info()
# + dc={"key": "45"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 7. Splitting the dataset into train and test sets
# <p>We have successfully converted all the non-numeric values to numeric ones.</p>
# <p>Now, we will split our data into train set and test set to prepare our data for two different phases of machine learning modeling: training and testing. Ideally, no information from the test data should be used to scale the training data or should be used to direct the training process of a machine learning model. Hence, we first split the data and then apply the scaling.</p>
# <p>Also, features like <code>DriversLicense</code> and <code>ZipCode</code> are not as important as the other features in the dataset for predicting credit card approvals. We should drop them to design our machine learning model with the best set of features. In Data Science literature, this is often referred to as <em>feature selection</em>. </p>
# + dc={"key": "45"} tags=["sample_code"]
# Import train_test_split
from sklearn.model_selection import train_test_split
# Drop the features 11 and 13 and convert the DataFrame to a NumPy array
cc_apps = cc_apps.drop([cc_apps.columns[11], cc_apps.columns[13]], axis=1)
cc_apps = cc_apps.values
# Segregate features and labels into separate variables
X,y = cc_apps[:,0:13] , cc_apps[:,13]
# Split into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.33,
random_state=42)
# + dc={"key": "52"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 8. Preprocessing the data (part ii)
# <p>The data is now split into two separate sets - train and test sets respectively. We are only left with one final preprocessing step of scaling before we can fit a machine learning model to the data. </p>
# <p>Now, let's try to understand what these scaled values mean in the real world. Let's use <code>CreditScore</code> as an example. The credit score of a person is their creditworthiness based on their credit history. The higher this number, the more financially trustworthy a person is considered to be. So, a <code>CreditScore</code> of 1 is the highest since we're rescaling all the values to the range of 0-1.</p>
# + dc={"key": "52"} tags=["sample_code"]
# Import MinMaxScaler
from sklearn.preprocessing import MinMaxScaler
# Instantiate MinMaxScaler and use it to rescale X_train and X_test
scaler = MinMaxScaler(feature_range=(0, 1))
rescaledX_train = scaler.fit_transform(X_train)
rescaledX_test = scaler.fit_transform(X_test)
# + dc={"key": "59"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 9. Fitting a logistic regression model to the train set
# <p>Essentially, predicting if a credit card application will be approved or not is a <a href="https://en.wikipedia.org/wiki/Statistical_classification">classification</a> task. <a href="http://archive.ics.uci.edu/ml/machine-learning-databases/credit-screening/crx.names">According to UCI</a>, our dataset contains more instances that correspond to "Denied" status than instances corresponding to "Approved" status. Specifically, out of 690 instances, there are 383 (55.5%) applications that got denied and 307 (44.5%) applications that got approved. </p>
# <p>This gives us a benchmark. A good machine learning model should be able to accurately predict the status of the applications with respect to these statistics.</p>
# <p>Which model should we pick? A question to ask is: <em>are the features that affect the credit card approval decision process correlated with each other?</em> Although we can measure correlation, that is outside the scope of this notebook, so we'll rely on our intuition that they indeed are correlated for now. Because of this correlation, we'll take advantage of the fact that generalized linear models perform well in these cases. Let's start our machine learning modeling with a Logistic Regression model (a generalized linear model).</p>
# + dc={"key": "59"} tags=["sample_code"]
# Import LogisticRegression
from sklearn.linear_model import LogisticRegression
# Instantiate a LogisticRegression classifier with default parameter values
logreg = LogisticRegression()
# Fit logreg to the train set
logreg.fit(rescaledX_train, y_train)
# + dc={"key": "66"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 10. Making predictions and evaluating performance
# <p>But how well does our model perform? </p>
# <p>We will now evaluate our model on the test set with respect to <a href="https://developers.google.com/machine-learning/crash-course/classification/accuracy">classification accuracy</a>. But we will also take a look the model's <a href="http://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/">confusion matrix</a>. In the case of predicting credit card applications, it is equally important to see if our machine learning model is able to predict the approval status of the applications as denied that originally got denied. If our model is not performing well in this aspect, then it might end up approving the application that should have been approved. The confusion matrix helps us to view our model's performance from these aspects. </p>
# + dc={"key": "66"} tags=["sample_code"]
# Import confusion_matrix
from sklearn.metrics import confusion_matrix
# Use logreg to predict instances from the test set and store it
y_pred = logreg.predict(rescaledX_test)
# Get the accuracy score of logreg model and print it
print("Accuracy of logistic regression classifier: ", logreg.score(rescaledX_test, y_test))
# Print the confusion matrix of the logreg model
confusion_matrix(y_pred, y_test)
# + dc={"key": "73"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 11. Grid searching and making the model perform better
# <p>Our model was pretty good! It was able to yield an accuracy score of almost 84%.</p>
# <p>For the confusion matrix, the first element of the of the first row of the confusion matrix denotes the true negatives meaning the number of negative instances (denied applications) predicted by the model correctly. And the last element of the second row of the confusion matrix denotes the true positives meaning the number of positive instances (approved applications) predicted by the model correctly.</p>
# <p>Let's see if we can do better. We can perform a <a href="https://machinelearningmastery.com/how-to-tune-algorithm-parameters-with-scikit-learn/">grid search</a> of the model parameters to improve the model's ability to predict credit card approvals.</p>
# <p><a href="http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html">scikit-learn's implementation of logistic regression</a> consists of different hyperparameters but we will grid search over the following two:</p>
# <ul>
# <li>tol</li>
# <li>max_iter</li>
# </ul>
# + dc={"key": "73"} tags=["sample_code"]
# Import GridSearchCV
from sklearn.model_selection import GridSearchCV
# Define the grid of values for tol and max_iter
tol = [0.01, 0.001, 0.0001]
max_iter = [100, 150, 200]
# Create a dictionary where tol and max_iter are keys and the lists of their values are corresponding values
param_grid = dict(tol=tol, max_iter=max_iter)
print(param_grid)
# + dc={"key": "80"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 12. Finding the best performing model
# <p>We have defined the grid of hyperparameter values and converted them into a single dictionary format which <code>GridSearchCV()</code> expects as one of its parameters. Now, we will begin the grid search to see which values perform best.</p>
# <p>We will instantiate <code>GridSearchCV()</code> with our earlier <code>logreg</code> model with all the data we have. Instead of passing train and test sets separately, we will supply <code>X</code> (scaled version) and <code>y</code>. We will also instruct <code>GridSearchCV()</code> to perform a <a href="https://www.dataschool.io/machine-learning-with-scikit-learn/">cross-validation</a> of five folds.</p>
# <p>We'll end the notebook by storing the best-achieved score and the respective best parameters.</p>
# <p>While building this credit card predictor, we tackled some of the most widely-known preprocessing steps such as <strong>scaling</strong>, <strong>label encoding</strong>, and <strong>missing value imputation</strong>. We finished with some <strong>machine learning</strong> to predict if a person's application for a credit card would get approved or not given some information about that person.</p>
# + dc={"key": "80"} tags=["sample_code"]
# Instantiate GridSearchCV with the required parameters
grid_model = GridSearchCV(estimator=logreg, param_grid=param_grid, cv=5)
# Use scaler to rescale X and assign it to rescaledX
rescaledX = scaler.fit_transform(X)
# Fit data to grid_model
grid_model_result = grid_model.fit(rescaledX, y)
# Summarize results
best_score, best_params = grid_model_result.best_score_, grid_model_result.best_params_
print("Best: %f using %s" % (best_score, best_params))
|
6 - Predicting Credit Card Approvals/Predicting Credit Card Approvals.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.5.6 64-bit (''cv3'': conda)'
# language: python
# name: python35664bitcv3conda56b31b492c17456d86703f6408b0e697
# ---
import os
import sys
import time
import base64
import uuid
import pandas as pd
from zipfile import ZipFile
from lxml import etree
import xml.etree.ElementTree as ET
import codecs
import json
from itertools import groupby
import difflib
# +
input_filepath = '/Users/kd/Workspace/python/DOCX/document-formatting/data/input/long_paragraph.docx'
output_dir = '/Users/kd/Workspace/python/DOCX/document-formatting/data/output'
fetch_content_filepath = '/Users/kd/Workspace/python/DOCX/document-formatting/data/input/long_paragraph.json'
filename = os.path.splitext(os.path.basename(input_filepath))[0]
translated_filename = filename + '_translated' + '.docx'
# +
def get_string_xmltree(xml):
return etree.tostring(xml)
def get_xml_tree(xml_string):
return etree.fromstring(xml_string)
def get_xmltree(filepath, parse='xml'):
if parse == 'html':
parser = etree.HTMLParser()
tree = etree.parse(open(filepath, mode='r', encoding='utf-8'), parser)
return tree
else:
with open(filepath,'r') as file:
xml_string = file.read()
return etree.fromstring(bytes(xml_string, encoding='utf-8'))
return None
def check_element_is(element, type_char):
word_schema1 = "http://schemas.openxmlformats.org/wordprocessingml/2006/main"
word_schema2 = 'http://purl.oclc.org/ooxml/wordprocessingml/main'
return (element.tag == '{%s}%s' % (word_schema1, type_char)) or (element.tag == '{%s}%s' % (word_schema2, type_char))
def get_specific_tags(node, type_char):
nodes = []
for elem in node.iter():
if check_element_is(elem, type_char):
nodes.append(elem)
return nodes
def add_identifier(node):
node.attrib['id'] = str(uuid.uuid4())
def is_run_superscript(run):
attrib = {}
vertAlign = get_specific_tags(run, 'vertAlign')
if len(vertAlign) > 0:
for key in vertAlign[0].attrib.keys():
attrib['vertAlign_' + key.split('}')[-1]] = vertAlign[0].attrib[key]
if 'vertAlign_val' in attrib:
if attrib['vertAlign_val'] == 'superscript':
return True
return False
def update_run_text(r1, r2):
t1s = get_specific_tags(r1, 't')
t2s = get_specific_tags(r2, 't')
# print('r1 text [%s], r2 text [%s]'% (t1s[0].text, t2s[0].text))
t1s[0].text = t1s[0].text + t2s[0].text
t2s[0].text = ''
def get_run_properties(run):
attrib = {}
rFonts = get_specific_tags(run, 'rFonts')
sz = get_specific_tags(run, 'sz')
szCs = get_specific_tags(run, 'szCs')
if len(rFonts) > 0:
for key in rFonts[0].attrib.keys():
attrib['rFonts_' + key.split('}')[-1]] = rFonts[0].attrib[key]
if len(sz) > 0:
for key in sz[0].attrib.keys():
attrib['sz_' + key.split('}')[-1]] = sz[0].attrib[key]
if len(szCs) > 0:
for key in szCs[0].attrib.keys():
attrib['szCs_' + key.split('}')[-1]] = szCs[0].attrib[key]
return attrib
def update_font_property(p, reduce=4):
szs = get_specific_tags(p, 'sz')
szCss = get_specific_tags(p, 'szCs')
value = '{%s}%s' % ("http://schemas.openxmlformats.org/wordprocessingml/2006/main", 'val')
for szCs in szCss:
size = szCs.attrib[value]
szCs.set(value, str(int(size) - reduce))
for sz in szs:
size = sz.attrib[value]
sz.set(value, str(int(size) - reduce))
def compare_run_properties(run1, run2):
attrib1 = get_run_properties(run1)
attrib2 = get_run_properties(run2)
if all (k in attrib1 for k in ('rFonts_ascii', 'sz_val', 'szCs_val')):
if all (k in attrib2 for k in ('rFonts_ascii', 'sz_val', 'szCs_val')):
if (attrib1['rFonts_ascii'] == attrib2['rFonts_ascii']) and \
(attrib1['szCs_val'] == attrib2['szCs_val']) and \
(attrib1['sz_val'] == attrib2['sz_val']) :
return True
return False
def get_line_connections(p):
runs = get_specific_tags(p, 'r')
text_runs = []
for run in runs:
if is_run_superscript(run) == False:
text_runs.append(run)
line_connections = []
for index in range(len(text_runs) - 1):
if (compare_run_properties(text_runs[index], text_runs[index+1])):
line_connections.append((index, index+1, 'CONNECTED'))
else:
line_connections.append((index, index+1, 'NOT_CONNECTED'))
return line_connections
def arrange_grouped_line_indices(line_connections, debug=False):
lines = [list(i) for j, i in groupby(line_connections, lambda a: a[2])]
if debug:
print('arrange_grouped_line_indices: %s \n---------\n' % (str(lines)))
arranged_lines = []
for line_items in lines:
indices = []
for line_item in line_items:
indices.append(line_item[0])
indices.append(line_item[1])
indices = sorted(list(set(indices)))
arranged_lines.append([indices, line_items[0][2]])
if debug:
print('arrange_grouped_line_indices,arranged_lines : %s \n---------\n' % (str(arranged_lines)))
final_arranged_lines = []
if len(arranged_lines) == 1:
final_arranged_lines.append([arranged_lines[0][0], arranged_lines[0][1]])
else:
for index, line_item in enumerate(arranged_lines):
if index == 0 and line_item[1] == 'NOT_CONNECTED':
del line_item[0][-1]
if index > 0 and index < (len(arranged_lines) - 1) and line_item[1] == 'NOT_CONNECTED':
del line_item[0][0]
del line_item[0][-1]
if index == (len(arranged_lines) - 1) and line_item[1] == 'NOT_CONNECTED':
del line_item[0][0]
final_arranged_lines.append([line_item[0], line_item[1]])
if debug:
print('final_arrange_grouped_line_indices,arranged_lines : %s \n---------\n' % (str(final_arranged_lines)))
return final_arranged_lines
def merge_runs(node, grouped_runs, debug=False):
runs = get_specific_tags(node, 'r')
text_runs = []
for run in runs:
if is_run_superscript(run) == False:
text_runs.append(run)
for element in grouped_runs:
if (element[1] == 'CONNECTED'):
for index, run_index in enumerate(element[0]):
if (index > 0):
if (debug):
print('merge index %d with %d' % ( run_index, 0))
update_run_text(text_runs[0], text_runs[run_index])
text_runs[run_index].getparent().remove(text_runs[run_index])
def update_document_runs(document):
'''
the function iterates through the p tags and merges run that have exactly same
visual property.
'''
tag_name = 'p'
tags = get_specific_tags(document, tag_name)
for p in tags:
grouped_runs = arrange_grouped_line_indices(get_line_connections(p))
merge_runs(p, grouped_runs, debug=False)
return document
def get_text_tags(document):
tags = []
runs = get_specific_tags(document, 'r')
for run in runs:
if is_run_superscript(run) == False:
texts = get_specific_tags(run, 't')
for text in texts:
if text.text and len(text.text.strip()) > 0:
add_identifier(text)
tags.append(text)
return tags
# +
def extract_docx(filepath, working_dir):
filename = os.path.splitext(os.path.basename(filepath))[0]
extract_dir = os.path.join(working_dir, filename)
with ZipFile(filepath, 'r') as file:
file.extractall(path=extract_dir)
filenames = file.namelist()
return extract_dir, filenames
def save_docx(extracted_dir, filenames, output_filename):
with ZipFile(output_filename, 'w') as docx:
for filename in filenames:
docx.write(os.path.join(extracted_dir, filename), filename)
def save_document_xml(extracted_dir, xml):
with open(os.path.join(extracted_dir,'word/document.xml'), 'wb') as f:
xmlstr = get_string_xmltree(xml)
f.write(xmlstr)
# -
def get_tokenized_sentences(filepath):
from jsonpath_rw import jsonpath, parse
json_data = json.load(codecs.open(fetch_content_filepath, 'r', 'utf-8-sig'))
jsonpath_expr = parse('$..tokenized_sentences[*]')
matches = jsonpath_expr.find(json_data)
tokenized_sentences = []
for match in matches:
tokenized_sentences.append(match.value)
return tokenized_sentences
# +
def count_occurrences(string, substring):
count = 0
start = 0
while start < len(string):
pos = string.find(substring, start)
if pos != -1:
start = pos + 1
count += 1
else:
break
return count
def check_string_status(doc_tag, tokenized):
doc_text = doc_tag.text.replace(" ", "")
tokenized_text = tokenized['src'].replace(" ", "")
if len(doc_text) < 2 or len(tokenized_text) < 2:
if doc_text.isdigit() == False or tokenized_text.isdigit() == False:
return (False, False)
'''
perfect match
'''
if doc_text == tokenized_text:
return (True, 0)
count = 0
if len(doc_text) > len(tokenized_text):
count = count_occurrences(doc_text, tokenized_text)
if count != 0:
return (True, -1)
else:
count = count_occurrences(tokenized_text, doc_text)
if count != 0:
return (True, 1)
return (False, False)
def string_overlap(str1, str2):
str1_list = [x for x in str1.split(' ') if x]
str1_set = set(str1_list)
str2_list = [x for x in str2.split(' ') if x]
str2_set = set(str2_list)
common_set = str1_set.intersection(str2_set)
diff_set = str1_set.difference(str2_set)
overlap_list = []
if len(str1_list) > len(str2_list):
for word in str2_list:
if word in list(common_set):
overlap_list.append(word)
else:
for word in str1_list:
if word in list(common_set):
overlap_list.append(word)
return ' '.join(overlap_list)
def check_string_status_v1(doc_tag, tokenized, overlap_threshold=4):
doc_text = doc_tag.text.replace(" ", "")
tokenized_text = tokenized['src'].replace(" ", "")
if len(doc_text) < 2 or len(tokenized_text) < 2:
if doc_text.isdigit() == False or tokenized_text.isdigit() == False:
return (False, False)
'''
perfect match
'''
if doc_text == tokenized_text:
return (True, 0)
doc_text = doc_tag.text
tokenized_text = tokenized['src']
overlap, start, end, percentage, smaller = get_overlap(doc_text, tokenized_text)
# overlap_str_list = [x for x in overlap_str.split(' ') if x]
# doc_text_list = [x for x in doc_text.split(' ') if x]
# tokenized_text_list = [x for x in tokenized_text.split(' ') if x]
# if len(overlap_str) > 0:
# if (len(doc_text_list) <= len(tokenized_text_list)):
# if (abs(len(doc_text_list) - len(overlap_str_list)) <= overlap_threshold):
# return (True, 1)
# else:
# if (abs(len(tokenized_text_list) - len(overlap_str_list)) <= overlap_threshold):
# return (True, -1)
# else:
# '''
# when sentence overlap is not found, trying overlap at character level
# '''
# count = 0
# if len(doc_text) > len(tokenized_text):
# count = count_occurrences(doc_text, tokenized_text)
# if count != 0:
# return (True, -1)
# else:
# count = count_occurrences(tokenized_text, doc_text)
# if count != 0:
# return (True, 1)
# return (False, False)
# -
def get_as_df(tags, tokenized_sentences):
doc_texts = []
doc_ids = []
for tag in tags:
doc_texts.append(tag.text)
doc_ids.append(tag.attrib['id'])
tokenized_src_texts = []
tokenized_tgt_texts = []
for tokenized_sentence in tokenized_sentences:
tokenized_src_texts.append(tokenized_sentence['src'])
tokenized_tgt_texts.append(tokenized_sentence['tgt'])
if len(doc_texts) > len(tokenized_src_texts):
empty = [''] * (len(doc_texts) - len(tokenized_src_texts))
tokenized_src_texts.extend(empty)
tokenized_tgt_texts.extend(empty)
else:
empty = [''] * (len(tokenized_src_texts) - len(doc_texts))
doc_texts.extend(empty)
doc_ids.extend(empty)
df = pd.DataFrame(list(zip(doc_texts, doc_ids, tokenized_src_texts, tokenized_tgt_texts)),
columns =['doc_texts', 'doc_ids', 'tokenized_src_texts', 'tokenized_tgt_texts'])
return df
# +
def get_updated_tokenized_text_matched(df):
'''
return df, df1, creates merged string of tokenized sentences that are equivalent to document text
'''
df1 = []
for doc_id in df['doc_id'].unique():
temp_df = df[df['doc_id'] == doc_id]
src_text = ''
tgt_text = ''
for index, row in temp_df.iterrows():
src_text = src_text + row['src'] + ' '
tgt_text = tgt_text + row['tgt'] + ' '
temp_df_first = temp_df[0:1].reset_index(drop=True)
temp_df_first.at[0, 'src'] = src_text
temp_df_first.at[0, 'tgt'] = tgt_text
df1.append(temp_df_first)
return pd.concat(df1)
def get_updated_document_text_matched(df):
'''
returns two dfs, df3: contains text that are actually substring of tokenized sentence
df4: are multiple occurance text that have same spelling.
'''
df1 = []
df2 = []
for s_id in df['s_id'].unique():
sid_df = df[df['s_id'] == s_id].reset_index(drop=True)
doc_text = ''
for index, row in sid_df.iterrows():
doc_text = doc_text + row['doc_text'] + ' '
doc_text_list = [x for x in doc_text.split(' ') if x]
sid_src_text_list = [x for x in sid_df.iloc[0]['src'].split(' ') if x]
if len(doc_text_list) <= len(sid_src_text_list):
for index, row in sid_df.iterrows():
if index > 0:
sid_df.at[index, 'tgt'] = ''
df1.append(sid_df)
else:
df2.append(sid_df)
df3 = pd.DataFrame()
df4 = pd.DataFrame()
if len(df1) > 0:
df3 = pd.concat(df1).reset_index(drop=True)
if len(df2) > 0:
df4 = pd.concat(df2).drop_duplicates(subset='doc_id', keep='first').reset_index(drop=True)
return df3, df4
def replace_translated_df(df, texts):
for index, row in df.iterrows():
for text in texts:
if 'id' in text.attrib:
if text.attrib['id'] == row['doc_id']:
text.text = row['tgt']
def get_matched_dfs(tokenized_sentences, texts):
matched_ids = []
matched_sids = []
founds = []
is_substrings = []
doc_texts = []
srcs = []
tgts = []
ids = []
s_ids = []
for sent_index in range(len(tokenized_sentences)):
for text_index in range(len(texts)):
if (texts[text_index].attrib['id'] in matched_ids) or \
(tokenized_sentences[sent_index]['s_id'] in matched_sids):
continue
is_found, is_substring = check_string_status_v1(texts[text_index], tokenized_sentences[sent_index])
if is_found and is_substring == 0:
matched_ids.append(texts[text_index].attrib['id'])
matched_sids.append(tokenized_sentences[sent_index]['s_id'])
founds.append(is_found)
is_substrings.append(is_substring)
doc_texts.append(texts[text_index].text)
ids.append(texts[text_index].attrib['id'])
s_ids.append(tokenized_sentences[sent_index]['s_id'])
srcs.append(tokenized_sentences[sent_index]['src'])
tgts.append(tokenized_sentences[sent_index]['tgt'])
df = pd.DataFrame(list(zip(founds, is_substrings, doc_texts, ids, s_ids, srcs, tgts)),
columns =['found', 'substr', 'doc_text', 'doc_id', 's_id', 'src', 'tgt'])
return df.loc[(df['substr'] == 0) & (df['found'] == True)].reset_index(drop=True), \
df.loc[(df['substr'] == -1) & (df['found'] == True)].reset_index(drop=True), \
df.loc[(df['substr'] == 1) & (df['found'] == True)].reset_index(drop=True), \
df.loc[(df['found'] == False)].reset_index(drop=True)
# +
def get_overlap(s1, s2):
if len(s1) < len(s2):
s = difflib.SequenceMatcher(lambda x: x == '.', s1, s2)
pos_a, pos_b, size = s.find_longest_match(0, len(s1), 0, len(s2))
return s1[pos_a:pos_a+size], pos_a, (pos_a+size), size/len(s1), s.ratio()
s = difflib.SequenceMatcher(None, s2, s1)
pos_a, pos_b, size = s.find_longest_match(0, len(s2), 0, len(s1))
return s2[pos_a:pos_a+size], pos_a, (pos_a+size), size/len(s2), s.ratio()
def get_filtered_dfs(tokenized_sentences, texts):
percent_threshold = 0.3
ratio_threshold = 0.5
overlaps = []
percents = []
ratios = []
docs = []
srcs = []
tgts = []
ids = []
s_ids = []
for sent_index in range(len(tokenized_sentences)):
for text_index in range(len(texts)):
doc_text = texts[text_index].text
tokenized_text = tokenized_sentences[sent_index]['src']
overlap, start, end, percent, ratio = get_overlap(doc_text, tokenized_text)
overlaps.append(overlap)
percents.append(percent)
ratios.append(ratio)
docs.append(texts[text_index].text)
ids.append(texts[text_index].attrib['id'])
srcs.append(tokenized_sentences[sent_index]['src'])
tgts.append(tokenized_sentences[sent_index]['tgt'])
s_ids.append(tokenized_sentences[sent_index]['s_id'])
df = pd.DataFrame(list(zip(percents, ratios, overlaps, docs, srcs, tgts, s_ids, ids)),
columns =['percent', 'ratio', 'overlap', 'doc_text', 'src', 'tgt', 's_id', 'doc_id'])
filtered_df = df.loc[(df['percent'] >= percent_threshold ) & (df['ratio'] >= ratio_threshold)]
pm_df = filtered_df.loc[(filtered_df['percent'] == 1.0 ) & (filtered_df['ratio'] == 1.0)]
pm_df1 = filtered_df[~filtered_df.index.isin(pm_df.index)]
return pm_df, pm_df1, df
# -
extracted_dir, filenames = extract_docx(input_filepath, output_dir)
# +
document_xml = get_xmltree(os.path.join(extracted_dir, 'word', 'document.xml'))
document_xml = update_document_runs(document_xml)
texts = get_text_tags(document_xml)
tokenized_sentences = get_tokenized_sentences(fetch_content_filepath)
print('document has (%d) text tags, tokenized sentences (%d)' % (len(texts), len(tokenized_sentences)))
pm_df, filtered_df, df = get_filtered_dfs(tokenized_sentences, texts)
tm_df = get_updated_tokenized_text_matched(filtered_df)
substring_df, multiple_df = get_updated_document_text_matched(filtered_df)
df.to_csv('df.csv')
filtered_df.to_csv('filtered_df.csv')
pm_df.to_csv('pm_df.csv')
tm_df.to_csv('tm_df.csv')
substring_df.to_csv('substring_df.csv')
multiple_df.to_csv('multiple_df.csv')
print('filtered match (%d)' % (len(filtered_df)))
print('perfect match (%d), tokenized match (%d) substring match (%d) multiple match (%d)' \
% (len(pm_df), len(tm_df), len(substring_df), len(multiple_df)))
# pm_df, tm_df, dtm_df, df = get_matched_dfs(tokenized_sentences, texts)
# print('perfect match (%d), tokenized sentences matched (%d), document text matched (%d)' % (len(pm_df), len(tm_df), len(dtm_df)))
# +
'''
replace and create translated sentence
'''
replace_translated_df(pm_df, texts)
# replace_translated_df(tm_df, texts)
# replace_translated_df(substring_df, texts)
# replace_translated_df(multiple_df, texts)
# replace_translated_df(get_updated_tokenized_text_matched(tm_df), texts)
# substring_df, multiple_df = get_updated_document_text_matched(dtm_df)
# if substring_df.empty == False:
# replace_translated_df(substring_df, texts)
# if multiple_df.empty != False:
# replace_translated_df(multiple_df, texts)
# -
ps = get_specific_tags(document_xml, 'p')
for p in ps:
update_font_property(p, 5)
save_document_xml(extracted_dir, document_xml)
save_docx(extracted_dir, filenames, os.path.join(output_dir, translated_filename))
# +
# doc_text = 'recruited prior to change of policy as (ii) aforesaid. The Permanent Commission shall be offered to them after completion of five years. They would also be entitled to all consequential benefits such as promotion and other financial benefits. However, the aforesaid benefits are to be made available only to women officers in service or who have approached this Court by filing these petitions and have retired during the course of pendency of the petitions.'
# tokenized_text = 'This benefit would be conferred to women officers recruited prior to change of policy as (ii) aforesaid.'
tokenized_text = 'The original ToE provided for a contractual period of five years after which the officers were to be released from service. The officers who were granted commission under the Army instruction were not entitled to PC or to any extension beyond five years of commissioned service.'
doc_text = '8. The original ToE provided for a contractual period of five years after which the officers were to be released from service.'
# +
def get_overlap_v1(s1, s2):
if len(s1) > len(s2):
return get_diff(s1, s2)
else:
return get_diff(s2, s1)
def get_diff(s1, s2):
s = difflib.SequenceMatcher(lambda x: x == '.', s1, s2)
pos_s1, pos_s2, match_size = s.find_longest_match(0, len(s1), 0, len(s2))
return s1[pos_s1:pos_s1+match_size], pos_s1, pos_s2, \
len(s1[pos_s1:pos_s1+match_size])/len(s1), \
len(s2[pos_s2:pos_s2+match_size])/len(s2), \
s.ratio()
# -
print(get_overlap_v1('1. 1999', '1999'))
print(get_overlap_v1('1999', '1. 1999'))
|
notebooks/doc_construct_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Art Style Transfer
#
# This notebook is an implementation of the algorithm described in "A Neural Algorithm of Artistic Style" (http://arxiv.org/abs/1508.06576) by Gatys, Ecker and Bethge. Additional details of their method are available at http://arxiv.org/abs/1505.07376 and http://bethgelab.org/deepneuralart/.
#
# An image is generated which combines the content of a photograph with the "style" of a painting. This is accomplished by jointly minimizing the squared difference between feature activation maps of the photo and generated image, and the squared difference of feature correlation between painting and generated image. A total variation penalty is also applied to reduce high frequency noise.
#
# This notebook was originally sourced from [Lasagne Recipes](https://github.com/Lasagne/Recipes/tree/master/examples/styletransfer), but has been modified to use a GoogLeNet network (pre-trained and pre-loaded), and given some features to make it easier to experiment with.
# +
import theano
import theano.tensor as T
import lasagne
from lasagne.utils import floatX
import numpy as np
import scipy
import matplotlib.pyplot as plt
# %matplotlib inline
import os # for directory listings
import pickle
import time
AS_PATH='./images/art-style'
# +
from model import googlenet
net = googlenet.build_model()
net_input_var = net['input'].input_var
net_output_layer = net['prob']
# -
# Load the pretrained weights into the network :
# +
params = pickle.load(open('./data/googlenet/blvc_googlenet.pkl', 'rb'), encoding='iso-8859-1')
model_param_values = params['param values']
#classes = params['synset words']
lasagne.layers.set_all_param_values(net_output_layer, model_param_values)
IMAGE_W=224
print("Loaded Model parameters")
# -
# ### Choose the Photo to be *Enhanced*
#
photos = [ '%s/photos/%s' % (AS_PATH, f) for f in os.listdir('%s/photos/' % AS_PATH) if not f.startswith('.')]
photo_i=-1 # will be incremented in next cell (i.e. to start at [0])
# Executing the cell below will iterate through the images in the ```./images/art-style/photos``` directory, so you can choose the one you want
photo_i += 1
photo = plt.imread(photos[photo_i % len(photos)])
photo_rawim, photo = googlenet.prep_image(photo)
plt.imshow(photo_rawim)
# ### Choose the photo with the required 'Style'
styles = [ '%s/styles/%s' % (AS_PATH, f) for f in os.listdir('%s/styles/' % AS_PATH) if not f.startswith('.')]
style_i=-1 # will be incremented in next cell (i.e. to start at [0])
# Executing the cell below will iterate through the images in the ```./images/art-style/styles``` directory, so you can choose the one you want
style_i += 1
art = plt.imread(styles[style_i % len(styles)])
art_rawim, art = googlenet.prep_image(art)
plt.imshow(art_rawim)
# This defines various measures of difference that we'll use to compare the current output image with the original sources.
def plot_layout(combined):
def no_axes():
plt.gca().xaxis.set_visible(False)
plt.gca().yaxis.set_visible(False)
plt.figure(figsize=(9,6))
plt.subplot2grid( (2,3), (0,0) )
no_axes()
plt.imshow(photo_rawim)
plt.subplot2grid( (2,3), (1,0) )
no_axes()
plt.imshow(art_rawim)
plt.subplot2grid( (2,3), (0,1), colspan=2, rowspan=2 )
no_axes()
plt.imshow(combined, interpolation='nearest')
plt.tight_layout()
# +
def gram_matrix(x):
x = x.flatten(ndim=3)
g = T.tensordot(x, x, axes=([2], [2]))
return g
def content_loss(P, X, layer):
p = P[layer]
x = X[layer]
loss = 1./2 * ((x - p)**2).sum()
return loss
def style_loss(A, X, layer):
a = A[layer]
x = X[layer]
A = gram_matrix(a)
G = gram_matrix(x)
N = a.shape[1]
M = a.shape[2] * a.shape[3]
loss = 1./(4 * N**2 * M**2) * ((G - A)**2).sum()
return loss
def total_variation_loss(x):
return (((x[:,:,:-1,:-1] - x[:,:,1:,:-1])**2 + (x[:,:,:-1,:-1] - x[:,:,:-1,1:])**2)**1.25).sum()
# -
# Here are the GoogLeNet layers that we're going to pay attention to :
layers = [
# used for 'content' in photo - a mid-tier convolutional layer
'inception_4b/output',
# used for 'style' - conv layers throughout model (not same as content one)
'conv1/7x7_s2', 'conv2/3x3', 'inception_3b/output', 'inception_4d/output',
]
#layers = [
# # used for 'content' in photo - a mid-tier convolutional layer
# 'pool4/3x3_s2',
#
# # used for 'style' - conv layers throughout model (not same as content one)
# 'conv1/7x7_s2', 'conv2/3x3', 'pool3/3x3_s2', 'inception_5b/output',
#]
layers = {k: net[k] for k in layers}
# ### Precompute layer activations for photo and artwork
# This takes ~ 20 seconds
# +
input_im_theano = T.tensor4()
outputs = lasagne.layers.get_output(layers.values(), input_im_theano)
photo_features = {k: theano.shared(output.eval({input_im_theano: photo}))
for k, output in zip(layers.keys(), outputs)}
art_features = {k: theano.shared(output.eval({input_im_theano: art}))
for k, output in zip(layers.keys(), outputs)}
# +
# Get expressions for layer activations for generated image
generated_image = theano.shared(floatX(np.random.uniform(-128, 128, (1, 3, IMAGE_W, IMAGE_W))))
gen_features = lasagne.layers.get_output(layers.values(), generated_image)
gen_features = {k: v for k, v in zip(layers.keys(), gen_features)}
# -
# ### Define the overall loss / badness function
# +
losses = []
# content loss
cl = 10 /1000.
losses.append(cl * content_loss(photo_features, gen_features, 'inception_4b/output'))
# style loss
sl = 20 *1000.
losses.append(sl * style_loss(art_features, gen_features, 'conv1/7x7_s2'))
losses.append(sl * style_loss(art_features, gen_features, 'conv2/3x3'))
losses.append(sl * style_loss(art_features, gen_features, 'inception_3b/output'))
losses.append(sl * style_loss(art_features, gen_features, 'inception_4d/output'))
#losses.append(sl * style_loss(art_features, gen_features, 'inception_5b/output'))
# total variation penalty
vp = 0.01 /1000. /1000.
losses.append(vp * total_variation_loss(generated_image))
total_loss = sum(losses)
# -
# ### The *Famous* Symbolic Gradient operation
grad = T.grad(total_loss, generated_image)
# ### Get Ready for Optimisation by SciPy
# +
# Theano functions to evaluate loss and gradient - takes around 1 minute (!)
f_loss = theano.function([], total_loss)
f_grad = theano.function([], grad)
# Helper functions to interface with scipy.optimize
def eval_loss(x0):
x0 = floatX(x0.reshape((1, 3, IMAGE_W, IMAGE_W)))
generated_image.set_value(x0)
return f_loss().astype('float64')
def eval_grad(x0):
x0 = floatX(x0.reshape((1, 3, IMAGE_W, IMAGE_W)))
generated_image.set_value(x0)
return np.array(f_grad()).flatten().astype('float64')
# -
# Initialize with the original ```photo```, since going from noise (the code that's commented out) takes many more iterations.
# +
generated_image.set_value(photo)
#generated_image.set_value(floatX(np.random.uniform(-128, 128, (1, 3, IMAGE_W, IMAGE_W))))
x0 = generated_image.get_value().astype('float64')
iteration=0
# -
# ### Optimize all those losses, and show the image
#
# To refine the result, just keep hitting 'run' on this cell (each iteration is about 60 seconds) :
# +
t0 = time.time()
scipy.optimize.fmin_l_bfgs_b(eval_loss, x0.flatten(), fprime=eval_grad, maxfun=40)
x0 = generated_image.get_value().astype('float64')
iteration += 1
if False:
plt.figure(figsize=(8,8))
plt.imshow(googlenet.deprocess(x0), interpolation='nearest')
plt.axis('off')
plt.text(270, 25, '# {} in {:.1f}sec'.format(iteration, (float(time.time() - t0))), fontsize=14)
else:
plot_layout(googlenet.deprocess(x0))
print('Iteration {}, ran in {:.1f}sec'.format(iteration, float(time.time() - t0)))
# -
|
notebooks/2-CNN/6-StyleTransfer/2-Art-Style-Transfer-googlenet_theano.ipynb
|
# +
import numpy as np
from numpy.linalg import cholesky
from matplotlib import pyplot as plt
from scipy.stats import multivariate_normal
from numpy.linalg import inv
import probml_utils as pml
np.random.seed(10)
def gaussSample(mu, sigma, n):
A = cholesky(sigma)
Z = np.random.normal(loc=0, scale=1, size=(len(mu), n))
return np.dot(A, Z).T + mu
mtrue = {}
prior = {}
muTrue = np.array([0.5, 0.5])
Ctrue = 0.1 * np.array([[2, 1], [1, 1]])
mtrue["mu"] = muTrue
mtrue["Sigma"] = Ctrue
xyrange = np.array([[-1, 1], [-1, 1]])
ns = [10]
X = gaussSample(mtrue["mu"], mtrue["Sigma"], ns[-1])
# fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(24, 8))
# fig.suptitle('gauss2dUpdateData')
fig, ax1 = plt.subplots()
ax1.plot(X[:, 0], X[:, 1], "o", markersize=8, markerfacecolor="b")
ax1.set_ylim([-1, 1])
ax1.set_xlim([-1, 1])
ax1.set_title("data")
ax1.plot(muTrue[0], muTrue[1], "x", linewidth=5, markersize=20, color="k")
pml.savefig("gauss_2d_update_data.pdf")
prior["mu"] = np.array([0, 0])
prior["Sigma"] = 0.1 * np.eye(2)
npoints = 100j
out = np.mgrid[xyrange[0, 0] : xyrange[0, 1] : npoints, xyrange[1, 0] : xyrange[1, 1] : npoints]
X1, X2 = out[0], out[1]
nr = X1.shape[0]
nc = X2.shape[0]
points = np.vstack([np.ravel(X1), np.ravel(X2)]).T
p = multivariate_normal.pdf(points, mean=prior["mu"], cov=prior["Sigma"]).reshape(nr, nc)
fig, ax2 = plt.subplots()
ax2.contour(X1, X2, p)
ax2.set_ylim([-1, 1])
ax2.set_xlim([-1, 1])
ax2.set_title("prior")
pml.savefig("gauss_2d_update_prior.pdf")
post = {}
data = X[: ns[0], :]
n = ns[0]
S0 = prior["Sigma"]
S0inv = inv(S0)
S = Ctrue
Sinv = inv(S)
Sn = inv(S0inv + n * Sinv)
mu0 = prior["mu"]
xbar = np.mean(data, 0)
muN = np.dot(Sn, (np.dot(n, np.dot(Sinv, xbar)) + np.dot(S0inv, mu0)))
post["mu"] = muN
post["Sigma"] = Sn
p = multivariate_normal.pdf(points, mean=post["mu"], cov=post["Sigma"]).reshape(nr, nc)
fig, ax3 = plt.subplots()
ax3.contour(X1, X2, p)
ax3.set_ylim([-1, 1])
ax3.set_xlim([-1, 1])
ax3.set_title("post after 10 observation")
pml.savefig("gauss_2d_update_post.pdf")
# fig.savefig("gauss2dUpdatePostSubplot.pdf")
|
notebooks/book1/03/gauss_infer_2d.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# dataset is taken from Kaggle website (https://www.kaggle.com/saurabh00007/diabetescsv)
df = pd.read_csv('./datasets/diabetes.csv')
df.head()
listOfUniqueAge = df['Age'].unique()
listOfUniqueAge
|
08-Getting-all-unique-attribute-values-of-a-column.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function
import os
import argparse
import time
import numpy as np
import mxnet as mx
import h5py
from mxnet import nd,gluon,autograd
from mxnet.gluon import nn,data as gdata
import pandas as pd
from d2l import mxnet as d2l
from visualization.util_vtk import visualization
from dataset import ShapeNet3D
from model import BlockOuterNet,Conv3DNet
from criterion import BatchIoU
from misc import decode_multiple_block, execute_shape_program
from interpreter import Interpreter
from programs.loop_gen import translate, rotate, end
from options import options_guided_adaptation,options_train_generator,options_train_executor
import socket
# +
def parse_argument():
parser = argparse.ArgumentParser(description="testing the program generator")
parser.add_argument('--model', type=str, default='model/generator of GA on shapenet {}',
help='path to the testing model')
parser.add_argument('--data', type=str, default='./data/{}_testing.h5',
help='path to the testing data')
parser.add_argument('--save_path', type=str, default='./output/{}/',
help='path to save the output results')
parser.add_argument('--batch_size', type=int, default=32, help='batch size')
parser.add_argument('--num_workers', type=int, default=4, help='number of workers')
parser.add_argument('--info_interval', type=int, default=10, help='freq for printing info')
parser.add_argument('--save_prog', action='store_true', help='save programs to text file')
parser.add_argument('--save_img', action='store_true', help='render reconstructed shapes to images')
parser.add_argument('--num_render', type=int, default=10, help='how many samples to be rendered')
#parser = get_parser()
opt,unknown = parser.parse_known_args()
opt.prog_save_path = os.path.join(opt.save_path, 'programs')
opt.imgs_save_path = os.path.join(opt.save_path, 'images')
return opt
def test_on_shapenet_data(epoch, test_loader, model,model_wo, opt, ctx, gen_shape=False):
generated_shapes = []
generated_shapes_wo = []
original_shapes = []
gen_pgms = []
gen_params = []
for idx, data in enumerate(test_loader):
start = time.time()
shapes = data
shape = nd.expand_dims(data,axis = 1).as_in_context(ctx)
with autograd.train_mode():
out = model.decode(shape)
out_wo = model_wo.decode(shape)
end = time.time()
out[0] = nd.round(out[0]).astype('int64')
out[1] = nd.round(out[1]).astype('int64')
out_wo[0] = nd.round(out_wo[0]).astype('int64')
out_wo[1] = nd.round(out_wo[1]).astype('int64')
if gen_shape:
generated_shapes.append(decode_multiple_block(out[0], out[1]))
generated_shapes_wo.append(decode_multiple_block(out_wo[0], out_wo[1]))
original_shapes.append(data.asnumpy())
save_pgms = nd.argmax(out[0], axis=3).asnumpy()
save_params = out[1].asnumpy()
gen_pgms.append(save_pgms)
gen_params.append(save_params)
if idx % opt.info_interval == 0:
print("Test: epoch {} batch {}/{}, time={:.3f}".format(epoch, idx, len(test_loader), end - start))
if gen_shape:
generated_shapes = np.concatenate(generated_shapes, axis=0)
generated_shapes_wo = np.concatenate(generated_shapes_wo, axis=0)
original_shapes = np.concatenate(original_shapes, axis=0)
gen_pgms = np.concatenate(gen_pgms, axis=0)
gen_params = np.concatenate(gen_params, axis=0)
return original_shapes, generated_shapes,generated_shapes_wo,gen_pgms, gen_params
# +
opt = parse_argument()
print('========= arguments =========')
for key, val in vars(opt).items():
print("{:20} {}".format(key, val))
print('========= arguments =========')
def visual(path,gen_shapes,file_name,nums_samples):
data = gen_shapes.transpose((0, 3, 2, 1))
data = np.flip(data, axis=2)
num_shapes = data.shape[0]
for i in range(min(nums_samples,num_shapes)):
voxels = data[i]
save_name = os.path.join(path, file_name.format(i))
visualization(voxels,
threshold=0.1,
save_name=save_name,
uniform_size=0.9)
def test_on_shape_net(tp,model_path, data_path,save_path,num_img):
# data loader
test_set = ShapeNet3D(data_path)
test_loader = gdata.DataLoader(
dataset=test_set,
batch_size=opt.batch_size,
shuffle=False,
num_workers=opt.num_workers,
)
# model
ctx = d2l.try_gpu()
opt_gen = options_train_generator.parse()
model = BlockOuterNet(opt_gen)
model_wo = BlockOuterNet(opt_gen)
model.init_blocks(ctx)
model_wo.init_blocks(ctx)
model_wo.load_parameters("model/model of blockouternet")
model.load_parameters(model_path)
# test the model and evaluate the IoU
ori_shapes, gen_shapes,gen_shapes_wo ,pgms, params = test_on_shapenet_data(epoch=0,
test_loader=test_loader,
model=model,
model_wo = model_wo,
opt=opt,ctx=ctx,
gen_shape=True)
# Visualization
visual(save_path,gen_shapes,'GA {}.png',num_img)
visual(save_path,ori_shapes,'GT {}.png',num_img)
visual(save_path,gen_shapes_wo,'Before GA {}.png',num_img)
gen_shapes = nd.from_numpy(gen_shapes).astype('float32')
ori_shapes = nd.from_numpy(ori_shapes).astype('float32')
gen_shapes_wo = nd.from_numpy(gen_shapes_wo).astype('float32')
IoU1 = BatchIoU(ori_shapes.copy(), gen_shapes)
IoU2 = BatchIoU(ori_shapes.copy(), gen_shapes_wo)
# execute the generated program to generate the reconstructed shapes
# for double-check purpose, can be disabled
'''
num_shapes = gen_shapes.shape[0]
res = []
for i in range(num_shapes):
data = execute_shape_program(pgms[i], params[i])
res.append(data.reshape((1, 32, 32, 32)))
res = nd.from_numpy(np.concatenate(res, axis=0)).astype('float32')
IoU_2 = BatchIoU(ori_shapes.copy(), res)
print(IoU1.mean() - IoU_2.mean())
assert abs(IoU1.mean() - IoU_2.mean()) < 0.1, 'IoUs are not matched'
# save results
save_file = os.path.join(opt.save_path.format(tp), 'shapes.h5')
f = h5py.File(save_file, 'w')
f['data'] = gen_shapes
f['pgms'] = pgms
f['params'] = params
f.close()
'''
# Interpreting programs to understandable program strings
interpreter = Interpreter(translate, rotate, end)
num_programs = gen_shapes.shape[0]
for i in range(min(num_programs, opt.num_render)):
program = interpreter.interpret(pgms[i], params[i])
save_file = os.path.join(opt.prog_save_path.format(tp), '{}.txt'.format(i))
print(save_file)
with open(save_file, 'w') as out:
out.write(program)
return IoU1.mean(),IoU2.mean()
# -
tp_ls = ["chair","table","bed","sofa","cabinet","bench"]
results = pd.DataFrame(columns=["chair","table","bed","sofa","cabinet","bench"],
index=["shape2prog w/o GA","shape2progs"])
for tp in tp_ls:
if not os.path.isdir(opt.prog_save_path.format(tp)):
os.makedirs(opt.prog_save_path.format(tp))
if not os.path.isdir(opt.imgs_save_path.format(tp)):
os.makedirs(opt.imgs_save_path.format(tp))
model_path = opt.model.format(tp);
data_path = opt.data.format(tp)
opt.save_path = opt.save_path.format(tp)
save_path = opt.imgs_save_path.format(tp)
IoU1 = test_on_shape_net(tp,model_path,data_path,save_path,12)
results[tp][0] = IoU1[1]
results[tp][1]= IoU1[0]
results
|
test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Checking data recovery matrices: modal DOF (`cb.cbtf`)
#
# This and other notebooks are available here: https://github.com/twmacro/pyyeti/tree/master/docs/tutorials.
# As with the `cbcheck` demo, we'll use superelement 102. The data recovery matrices were formed in the test directory: "tests/nastran_drm12".
#
# The `cb.cbtf` routine aides in checking the modal DOF. This function performs a base-drive analysis and returns the boundary and modal responses. These are then used by the analyst to plot frequency response curves as a sanity check.
#
# Notes:
#
# * This model uses units of kg, mm, s
# * It's a very light-weight truss: mass = 1.755 kg
#
# .. image:: se102.png
# <img src="./se102.png" />
# ---
# First, do some imports:
import numpy as np
import matplotlib.pyplot as plt
from pyyeti import cb, nastran
np.set_printoptions(precision=3, linewidth=130, suppress=True)
# Some settings specifically for the jupyter notebook.
# %matplotlib inline
plt.rcParams['figure.figsize'] = [6.4, 4.8]
plt.rcParams['figure.dpi'] = 150.
# Need path to data files:
import os
import inspect
pth = os.path.dirname(inspect.getfile(cb))
pth = os.path.join(pth, '../tests')
pth = os.path.join(pth, 'nastran_drm12')
# #### Load data recovery matrices
# We'll use the function `procdrm12` from the `nastran.op2` module. (This gets imported into the `nastran` namespace automatically.)
otm = nastran.procdrm12(os.path.join(pth, 'drm12'))
sorted(otm.keys())
# #### Load the mass and stiffness from the "nas2cam" output
#
# Use the `rdnas2cam` routine (imported from `nastran.op2`) to read data from the output of the "nas2cam" DMAP. This loads the data into a dict:
nas = nastran.rdnas2cam(os.path.join(pth, 'inboard_nas2cam'))
nas.keys()
maa = nas['maa'][102]
kaa = nas['kaa'][102]
# #### Get the USET table for the b-set DOF
uset = nas['uset'][102]
b = nastran.mksetpv(uset, 'p', 'b')
usetb = uset[b]
# show the coordinates (which are in basic):
usetb.loc[(slice(None), 1), :]
# #### Form b-set partition vector into a-set
# In this case, we already know the b-set are first but, since we have the nas2cam output, we can use `n2p.mksetpv` to be more general. We'll also get the q-set partition vector for later use.
b = nastran.mksetpv(uset, 'a', 'b')
q = ~b
b = np.nonzero(b)[0]
q = np.nonzero(q)[0]
print('b =', b)
print('q =', q)
# #### Form the damping matrix
# We'll use 2.5% critical damping.
baa = 2*.025*np.sqrt(np.diag(kaa))
baa[b] = 0
baa = np.diag(baa)
# ---
# #### Form rigid-body modes
# These are used to define the acceleration(s) of the boundary DOF. Each rigid-body mode defines a consistent acceleration field which is needed for a base-drive (which is really what `cbtf` does).
#
# Note the the second boundary grid is in a different coordinate system.
rbg = nastran.rbgeom_uset(usetb, [600, 150, 150])
rbg
# Do a check of the mass:
bb = np.ix_(b, b)
rbg.T @ maa[bb] @ rbg
# #### Define analysis frequency vector and run `cbtf`
# The ``save`` option is useful for speeding up loops 2 to 6:
freq = np.arange(0.1, 200., .1)
save = {}
sol = {}
for i in range(6):
sol[i] = cb.cbtf(maa, baa, kaa, rbg[:, i], freq, b, save)
# Each solution (eg, ``sol[0]``) has:
#
# * The boundary and modal accelerations, velocities and displacements (``.a, .v, .d``)
# * The boundary force (``.frc``)
# * The analysis frequency vector (``.freq``)
[i for i in dir(sol[0]) if i[0] != '_']
# Just to check the solution, we'll first look at the boundary responses. The acceleration should be the same as the input (0 or 1), and velocity & displacement should be large approaching zero, but approach zero as frequency increases. (They should equal 1 where $2\pi f$ is 1, or $f \approx 0.16$.) Off-axis values should be zero.
h = plt.plot(freq, abs(sol[0].a[b]).T, 'b',
freq, abs(sol[0].v[b]).T, 'r',
freq, abs(sol[0].d[b]).T, 'g')
plt.title('Boundary Responses')
plt.legend(h[::len(b)], ('Acce', 'Velo', 'Disp'), loc='best')
plt.ylim(-.1, 3)
plt.xlim(0, 5)
# The modal part has dynamic content as we'll see next. Note: for the x-direction, the modes of interest are above 50 Hz. The other directions have modal content much lower in frequency.
plt.figure(figsize=(8, 8))
plt.subplot(311); plt.plot(freq, abs(sol[0].a[q]).T); plt.title('Modal Acce')
plt.subplot(312); plt.plot(freq, abs(sol[0].v[q]).T); plt.title('Modal Velo')
plt.subplot(313); plt.plot(freq, abs(sol[0].d[q]).T); plt.title('Modal Disp')
plt.tight_layout()
# We can plot ``sol.frc`` to see the boundary forces needed to run the base-drive. Here, we'll use the rigid-body modes to sum the forces to the center point and plot that. The starting value for the x-direction should be 1.755 to match the mass.
plt.plot(freq, abs(rbg.T @ sol[0].frc).T)
plt.title('Boundary Forces');
# ---
# Finally, let's get to checking the data recovery matrices.
#
# The first one we'll check is the ``SPCF`` recovery. Since that was defined to recovery the boundary forces, the components should match the b-set parts of the mass and stiffness. (Note that ``SPCFD`` loses some precision through the DMAP as compared to the original stiffness.)
assert np.allclose(otm['SPCFA'], maa[b])
assert np.allclose(otm['SPCFD'], kaa[bb])
# For the ``ATM``, there should be some lines that start at 1.0. Other lines, should start at zero. These curves make sense.
plt.semilogy(freq, abs(otm['ATM'] @ sol[0].a).T)
plt.title('ATM')
plt.ylim(.001, 10)
# The ``LTMA`` curves should all start with zero slope. ``LTMD`` curves should be numerically zero since rigid-body displacement should not cause any loads. These look reasonable.
plt.subplot(211)
plt.semilogy(freq, abs(otm['LTMA'] @ sol[0].a).T)
plt.title('LTMA')
plt.subplot(212)
plt.semilogy(freq, abs(otm['LTMD'] @ sol[0].d[b]).T)
plt.title('LTMD')
plt.tight_layout()
# The ``DTMA`` curves should also start with zero slope, with values much less than 1.0. Some of the ``DTMD`` curves (the ones in the 'x' direction) should start at high values then quickly drop off as frequency increases.
plt.subplot(211)
plt.semilogy(freq, abs(otm['DTMA'] @ sol[0].a).T)
plt.title('DTMA')
plt.subplot(212)
plt.semilogy(freq, abs(otm['DTMD'] @ sol[0].d[b]).T)
plt.title('DTMD')
plt.tight_layout()
|
docs/tutorials/cbtf.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: p38C
# language: python
# name: p38c
# ---
import glob
import os
import json
import jsonlines
from transformers import BertTokenizer
import torch.nn as nn
import torch
import os
# HuggingFace
from transformers import BertTokenizer, BertModel, BertConfig
data=[json.loads(l) for l in open("data_sets/proc/moviescope/train_sentences.jsonl")]
class BertClf(nn.Module):
def __init__(self, args):
super(BertClf, self).__init__()
self.args = args
bert=BertModel.from_pretrained("bert-base-uncased")
self.bert_last_layer=bert.encoder.layer[11]
self.bert = BertModel.from_pretrained(args.bert_model)
self.dropout= nn.Dropout(args.dropout)
self.clf = nn.Linear(args.hidden_sz, args.n_classes)
def forward(self, txt1, mask1, segment1,txt2, mask2, segment2):
bsz= txt1.shape[0]
_, cls_1 = self.bert(
input_ids=txt1,
token_type_ids=segment1,
attention_mask=mask1,
return_dict=False,
)
cls_1=cls_1.reshape(bsz,1,self.args.hidden_sz)
embedding_t2=self.bert.embeddings(input_ids=txt2,token_type_ids=segment2)
embedinng_t2=torch.cat([cls_1,embedding_t2],dim=1)
outputs=self.bert(inputs_embeds=embedinng_t2,attention_mask=mask2)
cls_2 = outputs[1]
cls_2=cls_2.reshape(bsz,1,self.args.hidden_sz)
mask_id = torch.LongTensor([103])#.cuda()
mask_id = mask_id.unsqueeze(0).expand(bsz, 1)
mask_token_embeds = self.bert.embeddings(mask_id)
print(mask_token_embeds.shape,cls_1.shape,cls_2.shape)
x=torch.cat([mask_token_embeds,cls_1,cls_2],dim=1)
x= self.bert_last_layer(x)
return self.clf(x)
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
index=0
input1=tokenizer(data[index]["sentences"][0], return_tensors="pt", padding='max_length',truncation=True)
sentence1=input1.input_ids
segment1=input1.token_type_ids
attmask1=input1.attention_mask
# +
if len(data[index]["sentences"])<2:
data[index]["sentences"].append(" ")
input2=tokenizer(data[index]["sentences"][1], return_tensors="pt", padding='max_length',truncation=True)
sentence2=input2.input_ids[:,1:]
segment2=input2.token_type_ids[:,1:]
attmask2=input2.attention_mask
# -
import argparse
args = argparse.Namespace(
bert_model="bert-base-uncased",
model="mmbt",
batch_sz=8,
task_type="multilabel",
max_seq_len=512,
num_image_embeds=3,
n_workers=2,
dropout=0.1,
hidden_sz=768,
gradient_accumulation_steps=32,
max_epochs=40,
lr=1e-4,
patience=5,
lr_patience=2,
warmup=0.1,
freeze_img=3,
freeze_txt=6,
img_hidden_sz=2048,
data_path="",
task="",
img_embed_pool_type="avg",
n_classes=13,
savedir="checkpoints2",
name="mmbt"
)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
device
bert2=BertClf(args)
bert2.to(device)
bert2(sentence1,attmask1,segment1,sentence2,attmask2,segment2)
bert=BertModel.from_pretrained("bert-base-uncased")
bert_last_layer=bert.encoder.layer[11]
bert_last_layer
|
notebooks/.ipynb_checkpoints/bert3-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yukinaga/twitter_bot/blob/master/section_5/01_input_padding.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-9xgHp5jqQQO"
# # 入力のパディング
# 入力となる文章の長さは様々なのですが、バッチ内ではデータの時系列の長さを揃える必要があります。
# 今回は、短い文章に対して「パディング」を行い、バッチ内の全ての文章の長さを揃えます。
# + [markdown] id="Yar4MmG9ufdW"
# ## テンソルのパディング
# `nn.utils.rnn.pad_sequence`でパディングを行うことができます。
# + [markdown] id="Z9VjHFyXupy-"
# サイズが異なる3つのテンソルを用意します。
# + id="g2hYapMVuScF"
import torch
import torch.nn as nn
a = torch.ones(3, 5)
print(a)
b = torch.ones(2, 5) * 2
print(b)
c = torch.ones(1, 5) * 3
print(c)
# + [markdown] id="sCX-sQ3PvKwi"
# `nn.utils.rnn.pad_sequence`によりパディングを行います。
# + id="uD2oTL-vvPmk"
padded = nn.utils.rnn.pad_sequence([a, b, c], batch_first=True)
print(padded)
# + [markdown] id="-rHKMusSvjSN"
# ## Tensorのパッキング
# このままだと0の入力が多数になってしまうので、RNNで適切に扱うためには「パッキング」を行う必要があります。
# `rnn.pack_padded_sequence`により、0を除いたPackedSequence型のデータを作成できます。
# PackedSequence型のデータは、RNNに入力することが可能です。
# + id="Gkpk0jOwvjl0"
packed = nn.utils.rnn.pack_padded_sequence(padded, [3, 2, 1], batch_first=True, enforce_sorted=False)
print(packed)
# + [markdown] id="PKPZvXSFyC0O"
# ## RNNへ入力
# パッキングされたTensorをRNNへ入力します。
# 出力は`PackedSequence`型のデータに、隠れ層の状態hは通常のTensorになります。
# + id="GL3tsCWVyDC1"
rnn = nn.RNN(
input_size=5, # 入力サイズ
hidden_size=2, # ニューロン数
batch_first=True, # 入力を (バッチサイズ, 時系列の数, 入力の数) にする
)
y, h = rnn(packed)
print(y)
print(h)
# + [markdown] id="vT1Hodyf2uDW"
# ## PackedSequence型をTensorに戻す
# `pad_packed_sequence`により、RNNの出力をパディングされたTensorに戻します。
# + id="k-k1yfuo4XI1"
y_unpacked = nn.utils.rnn.pad_packed_sequence(y, batch_first=True)
print(y_unpacked)
|
section_5/01_input_padding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
buy = int(input("lotto 구매 개수를 입력 : "))
print("----------------------")
for x in range(1, buy+1):
lucky = [0, 0, 0, 0, 0, 0]
lucky[0] = random.randrange(1, 46, 1)
lucky[1] = lucky[0]
lucky[2] = lucky[0]
lucky[3] = lucky[0]
lucky[4] = lucky[0]
lucky[5] = lucky[0]
while (lucky[0] == lucky[1]):
lucky[1] = random.randrange(1, 46, 1)
while (lucky[0] == lucky[2] or lucky[1] == lucky[2]):
lucky[2] = random.randrange(1, 46, 1)
while (lucky[0] == lucky[3] or lucky[1] == lucky[3] or lucky[2] == lucky[3]):
lucky[3] = random.randrange(1, 46, 1)
while (lucky[0] == lucky[4] or lucky[1] == lucky[4] or lucky[2] == lucky[4] or lucky[3] == lucky[4]):
lucky[4] = random.randrange(1, 46, 1)
while (lucky[0] == lucky[5] or lucky[1] == lucky[5] or lucky[2] == lucky[5] or lucky[3] == lucky[5] or lucky[4] == lucky[5]):
lucky[5] = random.randrange(1, 46, 1)
lucky.sort()
print(lucky)
# -
|
Untitled1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collect ICU Stay and Caregiver Data
# ## NOTE: This is the first notebook of a 3 notebook series.
# +
# Data processing libraries
import pandas as pd
import numpy as np
#Util
import itertools
import datetime
# Database libraries
import psycopg2
# Stats libraries
from tableone import TableOne
import statsmodels.api as sm
import statsmodels.formula.api as smf
import scipy.stats
# Image libraries
# https://jakevdp.github.io/pdvega/
# jupyter nbextension enable vega3 --py --sys-prefix
import matplotlib.pyplot as plt
import pdvega
# %matplotlib inline
# -
# Create a database connection
# Replace user and password with credentials
user = 'xxx'
password = '<PASSWORD>'
host = 'hst953.csail.mit.edu'
dbname = 'mimic'
schema = 'mimiciii'
# Connect to the database
con = psycopg2.connect(dbname=dbname, user=user, host=host,
password=password)
cur = con.cursor()
cur.execute('SET search_path to {}'.format(schema))
# ## Querying the Data
# Here we query the database and extract information about a specific ICU stay. To increase performance we subset the data by age range.
# +
# Run query and assign the results to a Pandas DataFrame
# Requires the icustay_detail view from:
# https://github.com/MIT-LCP/mimic-code/tree/master/concepts/demographics
# And the OASIS score from:
# https://github.com/MIT-LCP/mimic-code/tree/master/concepts/severityscores
query = \
"""
WITH first_icu AS (
SELECT i.subject_id, i.hadm_id, i.icustay_id, i.gender, i.admittime admittime_hospital,
i.dischtime dischtime_hospital, i.los_hospital, i.age, i.admission_type,
i.hospital_expire_flag, i.intime intime_icu, i.outtime outtime_icu, i.los_icu, i.hospstay_seq, i.icustay_seq,
s.first_careunit,s.last_careunit,s.first_wardid, s.last_wardid
FROM icustay_detail i
LEFT JOIN icustays s
ON i.icustay_id = s.icustay_id
WHERE i.age > 50 AND i.age <= 60
)
SELECT f.*, o.icustay_expire_flag, o.oasis, o.oasis_prob
FROM first_icu f
LEFT JOIN oasis o
ON f.icustay_id = o.icustay_id;
"""
data = pd.read_sql_query(query,con)
# -
# After the data is loaded, we can take a look at it
data.columns
data
# We are interested in all of the rows related to a certain subject. We could do this with a database query, but to save network overhead we will do this here in memory
subj_rows = []
for i,subj_id in enumerate(data['subject_id']):
if subj_id == 13033:
subj_rows.append(i)
# Some subjects have multiple ICU stays, we would like to analyze the last stay because if the patient dies within a stay it will be their last.
# +
#This tuple is row, value
m_icu_id = (0,0)
#We want to find the last ICU stay so we find the maximum
for row_i in subj_rows:
d = data['icustay_seq'][row_i]
if d > m_icu_id[1]:
m_icu_id = (row_i,d)
m_icu_id
# -
# One off code is great and all, but we would prefer to create maintainable code that can later be extracted from the notebook, so the above code is merged into a function which creates a dictionary. This dictionary contains all of the tuples we generate keyed by subject
def create_icu_table():
icu_table = {}
sub_m = {}
#Find the rows related to each subject
for i,subj_id in enumerate(data['subject_id']):
if subj_id not in sub_m:
sub_m[subj_id] = []
sub_m[subj_id].append(i)
# For each row across the subject we find the last ICU stay
for subj,subj_rows in sub_m.items():
for row_i in subj_rows:
d = data['icustay_seq'][row_i]
if d > icu_table.get(subj,(0,0))[1]:
icu_table[subj]=(row_i,d)
return icu_table
it = create_icu_table()
# Now that we have all the relavant rows, we can subset our initial data set
target_rows = []
for row_i, _ in it.values():
target_rows.append(row_i)
data.iloc[target_rows]
# Just to be safe, we check that the table length is the same as the unique subjects
len(data['subject_id'].unique())
# +
#optional write out to spreadsheet
#writer = pd.ExcelWriter('max_icu_stay.xlsx')
#data.iloc[target_rows].to_excel(writer,'Sheet1')
#writer.save()
# -
# ## Getting caregiver data
# Test a query against the database for the caregivers associated with a specific chart
# +
item = 228232
query = \
"""
SELECT c.subject_id, c.hadm_id, c.icustay_id, c.charttime,
c.cgid,g.label
FROM chartevents c
LEFT JOIN caregivers g
ON c.cgid = g.cgid
WHERE c.icustay_id = """+str(item)+"""
"""
data_cg = pd.read_sql_query(query,con)
# -
# We see here that there a multiple caregivers which monitored this paitent; however, we do not know the role of each caregiver
data_cg['cgid'].value_counts()
# To find the caregiver label we check another row
def get_cgid_label(df, cgid):
return df.loc[df['cgid'] == cgid]['label'].values[0]
#test functionality
get_cgid_label(data_cg,18765)
# List comprehensions are always 100% easy to understand :P
#
# This list comprehension finds the associated label for each caregiver
[get_cgid_label(data_cg,idxx) for idxx in data_cg['cgid'].value_counts().index]
# Our previous query was a little too broad. Let's try looking at just some common labels
# +
query = \
"""
SELECT g.label
FROM caregivers g
WHERE g.label = 'RN' OR g.label = 'MD' OR g.label = 'Res' OR g.label = 'RO' OR g.label = 'MDs'
"""
data_cglabel = pd.read_sql_query(query,con)
# -
data_cglabel['label'].value_counts()
# Functions are useful, and in this case we would like to quickly count the number of labels from each group given a certain ICU stay
def get_measure_info(subj_icustay_id):
#Check type for safety
if type(subj_icustay_id)!= int:
raise TypeError
#TODO: Params for query
query = \
"""
SELECT c.icustay_id,c.cgid,g.label
FROM chartevents c
LEFT JOIN caregivers g
ON c.cgid = g.cgid
WHERE c.icustay_id = """+str(subj_icustay_id)+"""
"""
data_cg = pd.read_sql_query(query,con)
#The same list comprehension we saw above
mea_list = [(get_cgid_label(data_cg,idxx),v) for idxx, v in data_cg['cgid'].value_counts().items()]
#clinic_types -> ['RO','MD','Res','RN','MDs']
counts = {"RO":[0,0],"MDs":[0,0],"RN":[0,0],"OTH":[0,0]}
# We will count the total measurements
total_meas = 0
# Iterate over the measurements and count for each label group
for m_inst, m_visit_count in mea_list:
total_meas = total_meas + m_visit_count
if (m_inst == None):
counts["OTH"][0] = counts["OTH"][0] + 1
counts["OTH"][1] = counts["OTH"][1] + m_visit_count
else:
cmp = m_inst.upper()
if (cmp == "RO"):
counts["RO"][0] = counts["RO"][0] + 1
counts["RO"][1] = counts["RO"][1] + m_visit_count
elif (cmp == "MDS"):
counts["MDs"][0] = counts["MDs"][0] + 1
counts["MDs"][1] = counts["MDs"][1] + m_visit_count
elif (cmp == "MD"):
counts["MDs"][0] = counts["MDs"][0] + 1
counts["MDs"][1] = counts["MDs"][1] + m_visit_count
elif (cmp == "RES"):
counts["MDs"][0] = counts["MDs"][0] + 1
counts["MDs"][1] = counts["MDs"][1] + m_visit_count
elif (cmp == "RN"):
counts["RN"][0] = counts["RN"][0] + 1
counts["RN"][1] = counts["RN"][1] + m_visit_count
else:
counts["OTH"][0] = counts["OTH"][0] + 1
counts["OTH"][1] = counts["OTH"][1] + m_visit_count
# Returns a dictionary and int
return (counts,total_meas)
get_measure_info(228232)
#subset data to only the rows that contain the last visit
data_mro = data.iloc[target_rows]
# Produce measurement info for every row in the dataset. We slice to keep the queries small and the internet in the conference space was not great and caused large network delays
data_slices = []
cur_b = 0
width = 29
while cur_b < len(data_mro):
s = datetime.datetime.now()
d_info = data_mro['icustay_id'][cur_b:cur_b + width].apply(get_measure_info)
data_slices.append(d_info)
e = datetime.datetime.now()
print((e-s).total_seconds(),cur_b)
cur_b = cur_b + width + 1
data_slices
# We can look at the age distribution with a histogram
plt.hist(data_mro['age'])
# ## Save the data
pickle.dump(data_slices,open("d_slices_5060_g.p", "wb" ))
pickle.dump(data_mro,open("d_mro_5060_g.p","wb"))
|
FLOW1_GetICUStayAndCaregiverData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Tce3stUlHN0L"
# ##### Copyright 2018 The TensorFlow Authors.
#
#
# + colab={} colab_type="code" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="MfBg1C5NB3X0"
# # Using GPUs
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/alpha/guide/using_gpu"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/guide/using_gpu.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/guide/using_gpu.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="xHxb-dlhMIzW"
# TensorFlow supports running computations on a variety of types of devices, including CPU and GPU. They are represented with string identifiers for example:
#
# * `"/device:CPU:0"`: The CPU of your machine.
# * `"/GPU:0"`: Short-hand notation for the first GPU of your machine that is visible to TensorFlow
# * `"/job:localhost/replica:0/task:0/device:GPU:1"`: Fully qualified name of the second GPU of your machine that is visible to TensorFlow.
#
# If a TensorFlow operation has both CPU and GPU implementations, by default the GPU devices will be given priority when the operation is assigned to a device. For example, `tf.matmul` has both CPU and GPU kernels. On a system with devices `CPU:0` and `GPU:0`, the `GPU:0` device will be selected to run `tf.matmul` unless you explicitly request running it on another device.
# + [markdown] colab_type="text" id="MUXex9ctTuDB"
# ## Setup
#
# Ensure you have the latest TensorFlow release installed.
# + colab={} colab_type="code" id="IqR2PQG4ZaZ0"
from __future__ import absolute_import, division, print_function, unicode_literals
# !pip install tf-nightly-gpu-2.0-preview
import tensorflow as tf
# + [markdown] colab_type="text" id="UhNtHfuxCGVy"
# ## Logging device placement
#
# To find out which devices your operations and tensors are assigned to, put
# `tf.debugging.set_log_device_placement(True)` as the first statement of your
# program. Enabling device placement logging causes any Tensor allocations or operations to be printed.
# + colab={"height": 85} colab_type="code" id="2Dbw0tpEirCd" outputId="2ae5cf66-2cdf-49b2-d8d0-87492e97e137"
tf.debugging.set_log_device_placement(True)
# Create some tensors
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
# + [markdown] colab_type="text" id="kKhmFeraTdEI"
# The above code will print an indication the `MatMul` op was executed on `GPU:0`.
# + [markdown] colab_type="text" id="U88FspwGjB7W"
# ## Manual device placement
#
# If you would like a particular operation to run on a device of your choice
# instead of what's automatically selected for you, you can use `with tf.device`
# to create a device context, and all the operations within that context will
# run on the same designated device.
# + colab={"height": 85} colab_type="code" id="8wqaQfEhjHit" outputId="f96e5676-0dbc-462d-fde9-7c9831bfd01a"
tf.debugging.set_log_device_placement(True)
# Place tensors on the CPU
with tf.device('/CPU:0'):
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
# + [markdown] colab_type="text" id="8ixO89gRjJUu"
# You will see that now `a` and `b` are assigned to `CPU:0`. Since a device was
# not explicitly specified for the `MatMul` operation, the TensorFlow runtime will
# choose one based on the operation and available devices (`GPU:0` in this
# example) and automatically copy tensors between devices if required.
# + [markdown] colab_type="text" id="ARrRhwqijPzN"
# ## Limiting GPU memory growth
#
# By default, TensorFlow maps nearly all of the GPU memory of all GPUs (subject to
# [`CUDA_VISIBLE_DEVICES`](https://docs.nvidia.com/cuda/cuda-c-programming-guide/index.html#env-vars)) visible to the process. This is done to more efficiently use the relatively precious GPU memory resources on the devices by reducing memory fragmentation. To limit TensorFlow to a specific set of GPUs we use the `tf.config.experimental.set_visible_devices` method.
# + colab={} colab_type="code" id="hPI--n_jhZhv"
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Restrict TensorFlow to only use the first GPU
try:
tf.config.experimental.set_visible_devices(gpus[0], 'GPU')
except RuntimeError as e:
# Visible devices must be set at program startup
print(e)
# + [markdown] colab_type="text" id="N3x4M55DhYk9"
# In some cases it is desirable for the process to only allocate a subset of the available memory, or to only grow the memory usage as is needed by the process. TensorFlow provides two methods to control this.
#
# The first option is to turn on memory growth by calling `tf.config.experimental.set_memory_growth`, which attempts to allocate only as much GPU memory in needed for the runtime allocations: it starts out allocating very little memory, and as the program gets run and more GPU memory is needed, we extend the GPU memory region allocated to the TensorFlow process. Note we do not release memory, since it can lead to memory fragmentation. To turn on memory growth for a specific GPU, use the following code prior to allocating any tensors or executing any ops.
# + colab={} colab_type="code" id="jr3Kf1boFnCO"
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
tf.config.experimental.set_memory_growth(gpus[0], True)
except RuntimeError as e:
# Memory growth must be set at program startup
print(e)
# + [markdown] colab_type="text" id="I1o8t51QFnmv"
# Another way to enable this option is to set the environmental variable `TF_FORCE_GPU_ALLOW_GROWTH` to `true`. This configuration is platform specific.
#
# The second method is to configure a virtual GPU device with `tf.config.experimental.set_virtual_device_configuration` and set a hard limit on the total memory to allocate on the GPU.
# + colab={} colab_type="code" id="2qO2cS9QFn42"
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Restrict TensorFlow to only allocate 1GB of memory on the first GPU
try:
tf.config.experimental.set_virtual_device_configuration(
gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit=1024)])
except RuntimeError as e:
# Virtual devices must be set at program startup
print(e)
# + [markdown] colab_type="text" id="Bsg1iLuHFoLW"
# This is useful if you want to truly bound the amount of GPU memory available to the TensorFlow process. This is common practise for local development when the GPU is shared with other applications such as a workstation GUI.
# + [markdown] colab_type="text" id="B27_-1gyjf-t"
# ## Using a single GPU on a multi-GPU system
#
# If you have more than one GPU in your system, the GPU with the lowest ID will be
# selected by default. If you would like to run on a different GPU, you will need
# to specify the preference explicitly:
# + colab={"height": 34} colab_type="code" id="wep4iteljjG1" outputId="3f86b9f7-5c82-420e-a7c1-e34d3c115454"
tf.debugging.set_log_device_placement(True)
try:
# Specify an invalid GPU device
with tf.device('/device:GPU:2'):
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
except RuntimeError as e:
print(e)
# + [markdown] colab_type="text" id="jy-4cCO_jn4G"
# If the device you have specified does not exist, you will get a `RuntimeError`:
#
# If you would like TensorFlow to automatically choose an existing and supported device to run the operations in case the specified one doesn't exist, you can call `tf.config.set_soft_device_placement(True)`.
# + colab={"height": 85} colab_type="code" id="sut_UHlkjvWd" outputId="d01be27d-cadd-4e7d-a827-89d10e830762"
tf.config.set_soft_device_placement(True)
tf.debugging.set_log_device_placement(True)
# Creates some tensors
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
# + [markdown] colab_type="text" id="sYTYPrQZj2d9"
# ## Using multiple GPUs
# + [markdown] colab_type="text" id="IDZmEGq4j6kG"
# #### With `tf.distribute.Strategy`
#
# The best practice for using multiple GPUs is to use `tf.distribute.Strategy`.
# Here is a simple example:
# + colab={"height": 255} colab_type="code" id="1KgzY8V2AvRv" outputId="a74aca5d-bd8a-49a1-d345-ef9bbf933ddb"
strategy = tf.distribute.MirroredStrategy()
with strategy.scope():
inputs = tf.keras.layers.Input(shape=(1,))
predictions = tf.keras.layers.Dense(1)(inputs)
model = tf.keras.models.Model(inputs=inputs, outputs=predictions)
model.compile(loss='mse',
optimizer=tf.keras.optimizers.SGD(learning_rate=0.2))
# + [markdown] colab_type="text" id="Dy7nxlKsAxkK"
# This program will run a copy of your model on each GPU, splitting the input data
# between them, also known as "[data parallelism](https://en.wikipedia.org/wiki/Data_parallelism)".
#
# For more information about distribution strategies, check out the guide [here](./distribute_strategy.ipynb).
# + [markdown] colab_type="text" id="8phxM5TVkAY_"
# #### Without `tf.distribute.Strategy`
#
# `tf.distribute.Strategy` works under the hood by replicating computation across devices. You can manually implement replication by constructing your model on each GPU. For example:
# + colab={"height": 119} colab_type="code" id="AqPo9ltUA_EY" outputId="83a9a1fa-cf3f-4f08-f121-f70649831aad"
tf.debugging.set_log_device_placement(True)
gpus = tf.config.experimental.list_logical_devices('GPU')
if gpus:
# Replicate your computation on multiple GPUs
c = []
for gpu in gpus:
with tf.device(gpu.name):
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c.append(tf.matmul(a, b))
with tf.device('/CPU:0'):
matmul_sum = tf.add_n(c)
print(matmul_sum)
|
site/en/r2/guide/using_gpu.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="aD3O3_zBXRNA" outputId="8b4eceb6-416b-444f-d577-52eacfb0621f"
# !pip install pyspark
# + id="A6QpiLo_Yp9Z" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="02154bac-9918-48be-8864-b826fbfec495"
from pyspark import SparkContext
sc = SparkContext()
sc
# + colab={"base_uri": "https://localhost:8080/"} id="Vgip0aeKXuVc" outputId="0f456c05-6afa-4cc7-d1c9-ca3bfa2c9532"
# flatMap
rdd = sc.parallelize([2, 3, 4])
newRDD = rdd.flatMap(lambda x: range(1, x*2))
newRDD.collect()
# + colab={"base_uri": "https://localhost:8080/"} id="5PWek5HIYpPn" outputId="aacf32a7-9980-4304-98fb-32e14cb6297d"
# mapValues
rdd = sc.parallelize([("a", ["apple", "banana", "lemon"]), ("b", ["grapes"])])
rdd.mapValues(lambda x: len(x)).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="zo4X1NYNLgK9" outputId="3521a80c-6d7f-4124-f01c-21244aa5e448"
# mapValues
rdd = sc.parallelize([("a", ["apple", "banana", "lemon"]), ("b", ["grapes"])])
rdd.mapValues(len).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="0L5MobtMZ7IO" outputId="da02f9fc-a940-4b17-ddd4-478ad2000c67"
# map
rdd.map(lambda x: (x[0], len(x[1]))).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="snzPiPmfbNwg" outputId="b33a863d-de40-4836-cf24-d2c8cb75be90"
# filter
rdd = sc.parallelize([1, 2, 3, 4, 5])
rdd.filter(lambda x: x%2 == 0).collect()
# + [markdown] id="uoQhct0yX9Fh"
# Reduce
# + colab={"base_uri": "https://localhost:8080/"} id="20E2P47JbNzR" outputId="df934b66-3575-4ce5-bda0-60c5f656486a"
# reduce
sc.parallelize([1, 2, 3, 4, 5]).reduce(lambda x, y: x+y)
# + colab={"base_uri": "https://localhost:8080/"} id="qynkIX3CbN2Q" outputId="7d02b8ef-7a95-4831-c5b2-d3bb1c71c2ce"
# reduceByKey
rdd = sc.parallelize([("to", 1), ("be", 1), ("or", 1), ("not", 1), ("to", 1),("be", 1)])
rdd.reduceByKey(lambda x, y: x+y).collect()
# + id="Yg11sOGifchl" colab={"base_uri": "https://localhost:8080/"} outputId="2d36be55-21f2-4817-ea36-778652c79b29"
# union
rdd = sc.parallelize([1, 1, 2, 3])
rdd.union(rdd).collect()
# + id="ExKoVVy8fckm" colab={"base_uri": "https://localhost:8080/"} outputId="dd665bfe-78bd-46c6-c3cd-03fd32c3f038"
# distinct
sc.parallelize([1, 1, 2, 3]).distinct().collect()
# + id="Rf-cDI86fcny" colab={"base_uri": "https://localhost:8080/"} outputId="96b748fc-9347-4dda-d4dc-ffe924eeee1e"
# join
rdd1 = sc.parallelize([(1, 'a'), (1, 'b'), (5, 'c'), (2, 'd'), (3, 'e')])
rdd2 = sc.parallelize([(1, 'AA'), (5, 'BB'), (5, 'CC'), (6, 'DD')])
rdd1.join(rdd2).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="XQb9zReajP6k" outputId="73b03202-cc2c-4c5b-fae2-dc6dd202c0f0"
# leftOuterJoin
rdd1 = sc.parallelize([(1, 'a'), (1, 'b'), (5, 'c'), (2, 'd'), (3, 'e')])
rdd2 = sc.parallelize([(1, 'AA'), (5, 'BB'), (5, 'CC'), (6, 'DD')])
rdd1.leftOuterJoin(rdd2).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="47VmGPRhjfDs" outputId="b23d8495-fdf4-4fc7-a5fe-6de40b795daa"
# rightOuterJoin
rdd1 = sc.parallelize([(1, 'a'), (1, 'b'), (5, 'c'), (2, 'd'), (3, 'e')])
rdd2 = sc.parallelize([(1, 'AA'), (5, 'BB'), (5, 'CC'), (6, 'DD')])
rdd1.rightOuterJoin(rdd2).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="bmBGjKjyjkHl" outputId="198e1215-155d-4d90-d306-0dd7f9df8445"
# fullOuterJoin
rdd1 = sc.parallelize([(1, 'a'), (1, 'b'), (5, 'c'), (2, 'd'), (3, 'e')])
rdd2 = sc.parallelize([(1, 'AA'), (5, 'BB'), (5, 'CC'), (6, 'DD')])
rdd1.fullOuterJoin(rdd2).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="seE7YNEZjph2" outputId="60581da9-3189-4dc4-9d37-49eff79db4d3"
# cartesian
rdd = sc.parallelize([1,2,3])
rdd.cartesian(rdd).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="jjOi5nrCk3MX" outputId="22115faa-e5ed-4b86-d53b-c4a71330d71b"
# groupByKey
rdd = sc.parallelize([("a", 1), ("b", 1), ("a", 1)])
rdd.groupByKey().collect()
# + colab={"base_uri": "https://localhost:8080/"} id="W7lwOt6ulWdq" outputId="6530313d-caff-467a-d07d-04052a17f58b"
# groupByKey
rdd = sc.parallelize([("a", 1), ("b", 1), ("a", 1), ("a", 2), ("b", 3)])
rdd.groupByKey().mapValues(list).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="VqBTcy8qllhw" outputId="84b1872a-1d27-4caa-a039-55856042a81a"
# sortByKey
rdd = sc.parallelize([("c", 1), ("a", 1), ("b", 1)])
rdd.sortByKey().collect()
rdd.collect()
# + colab={"base_uri": "https://localhost:8080/"} id="VTckjADYoiwB" outputId="e732db87-6fc8-494e-cfe2-7f040d3da085"
# Aggregate the above RDD
def mySequenceFunction(x, y):
x.add(y)
return x
def myCombinerFunction(x, y):
x.update(y)
return x
rdd = sc.parallelize([("c1", "p1"), ("c2", "p1"), ("c1", "p1"), ("c2", "p2"), ("c2", "p3")])
rdd.aggregateByKey(set(), mySequenceFunction, myCombinerFunction).collect()
# + id="NdloaTSAolsu"
#rdd.treeAggregate(0, mySequenceFunction, myCombinerFunction).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="kZr0cNrjp_Y8" outputId="11f5d106-5833-4652-ba63-2b96c5563cf1"
# zip
x = sc.parallelize(range(0, 5))
y = sc.parallelize(range(1000, 1005))
x.zip(y).collect()
# + colab={"base_uri": "https://localhost:8080/"} id="VF-TJ0JL-KVW" outputId="1533920b-8c9b-4b71-829b-8b443f10f38d"
# zipWithIndex
y.zipWithIndex().collect()
# + colab={"base_uri": "https://localhost:8080/"} id="1XRBYzEp-Vq2" outputId="1612f10b-a6c0-4305-b349-08718f3cd38f"
sc.parallelize(["a", "b", "c", "d"], 3).zipWithIndex().collect()
# + [markdown] id="bSyWo1E5VxXJ"
# # Glom Operation
# The glom() operation returns an RDD created by coalescing all elements within each partition into a list. Glom is a highly useful operation when you want to access batches of an RDD. Listing 1.34 is a simple example of how we can access data batches of an RDD
# + colab={"base_uri": "https://localhost:8080/"} id="B7H0kyJR-bwW" outputId="1bc31476-17e0-4da9-9ba1-84ace7e6ac6f"
# glom
rdd = sc.parallelize([1, 2, 3, 4], 3)
sorted(rdd.glom().collect())
# + colab={"base_uri": "https://localhost:8080/"} id="-2qCthVZ-of6" outputId="f9634781-ac44-4477-fc51-4038dae16177"
# glom
rdd = sc.parallelize([1,2,3,4,5,6,7,8], 4)
rdd.glom().collect()
# + colab={"base_uri": "https://localhost:8080/"} id="07shqJTl-_xB" outputId="5304a178-7b73-4192-d996-c741507b0a6b"
# repartition
rdd = rdd.repartition(2)
rdd.glom().collect()
# + colab={"base_uri": "https://localhost:8080/"} id="MT458FNE_DF8" outputId="20ca5eb5-8551-4ac3-e385-42ec22b2c01d"
# coalesce: This is optimized or improved version of repartition()
# where the movement of the data across the partitions is lower using coalesce.
rdd = sc.parallelize([1,2,3,4,5,6,7,8], 4)
rdd.coalesce(2).glom().collect()
# + colab={"base_uri": "https://localhost:8080/"} id="qLq91TgXACBJ" outputId="0af578a3-6bb6-404a-9bc8-0f1063e85f86"
# cache
rdd = sc.parallelize([1, 2, 3, 4])
rdd.cache()
# + colab={"base_uri": "https://localhost:8080/"} id="CzfMsIC9C-jv" outputId="ab82e9e1-3ffe-4b97-820b-981aae75255d"
# countByValue
sc.parallelize([1, 2, 1, 2, 2], 2).countByValue()
# + colab={"base_uri": "https://localhost:8080/"} id="9kj2l3_DDNzp" outputId="2fe1eaa4-b6dd-44d9-f091-750917f6285a"
# first
sc.parallelize([(4, 2), (1, 2), (3, 2)]).first()
# + colab={"base_uri": "https://localhost:8080/"} id="S6s63tqKDndd" outputId="7252cca7-e8ed-43b2-9592-216b22364be2"
# take
sc.parallelize([2, 3, 4, 5, 6]).take(2)
# + id="meGSi_rrDntW" colab={"base_uri": "https://localhost:8080/"} outputId="ca517af9-1b1e-49cb-d26a-accd593344a9"
# groupByKey
x = sc.parallelize([('B',5),('B',4),('A',3),('A',2),('A',1)])
y = x.groupByKey()
print(x.collect())
print([(j[0],[i for i in j[1]]) for j in y.collect()])
# + id="e50cHHGGDnoh" colab={"base_uri": "https://localhost:8080/"} outputId="e005b85c-3fc2-498f-857b-e10bb8b6d525"
# aggregateByKey
x = sc.parallelize([('B',1),('B',2),('A',3),('A',4),('A',5)])
zeroValue = [] # empty list is 'zero value' for append operation
mergeVal = (lambda aggregated, el: aggregated + [(el,el**2)])
mergeComb = (lambda agg1,agg2: agg1 + agg2 )
y = x.aggregateByKey(zeroValue,mergeVal,mergeComb)
print(x.collect())
print(y.collect())
# + id="PtOw2W0ZVwjx"
|
Starter pack/RDD_Operations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="_wIWPxBVc3_O"
# # Getting Started: Voice swap application
# This notebook shows how to use NVIDIA NeMo (https://github.com/NVIDIA/NeMo) to construct a toy demo which will swap a voice in the audio fragment with a computer generated one.
#
# At its core the demo does:
#
# * Automatic speech recognition of what is said in the file. E.g. converting audio to text
# * Adding punctuation and capitalization to the text
# * Generating spectrogram from resulting text
# * Generating waveform audio from the spectrogram.
# + [markdown] colab_type="text" id="gzcsqceVdtj3"
# ## Installation
# NeMo can be installed via simple pip command.
# + colab={} colab_type="code" id="I9eIxAyKHREB"
BRANCH = 'main'
# !python -m pip install git+https://github.com/NVIDIA/NeMo.git@$BRANCH#egg=nemo_toolkit[all]
# + colab={} colab_type="code" id="-X2OyAxreGfl"
# Ignore pre-production warnings
import warnings
warnings.filterwarnings('ignore')
import nemo
# Import Speech Recognition collection
import nemo.collections.asr as nemo_asr
# Import Natural Language Processing colleciton
import nemo.collections.nlp as nemo_nlp
# Import Speech Synthesis collection
import nemo.collections.tts as nemo_tts
# We'll use this to listen to audio
import IPython
# + colab={} colab_type="code" id="1vC2DHawIGt8"
# Download audio sample which we'll try
# This is a sample from LibriSpeech Dev Clean dataset - the model hasn't seen it before
Audio_sample = '2086-149220-0033.wav'
# !wget https://dldata-public.s3.us-east-2.amazonaws.com/2086-149220-0033.wav
# Listen to it
IPython.display.Audio(Audio_sample)
# + [markdown] colab_type="text" id="zodyzdyTVXas"
# ## Instantiate pre-trained NeMo models which we'll use
# ``from_pretrained(...)`` API downloads and initialized model directly from the cloud.
#
# We will load audio_sample and convert it to text with QuartzNet ASR model (an action called transcribe).
# To convert text back to audio, we actually need to generate spectrogram with FastPitch first and then convert it to actual audio signal using the HiFiGAN vocoder.
# + colab={} colab_type="code" id="f_J9cuU1H6Bn"
# Speech Recognition model - QuartzNet
quartznet = nemo_asr.models.EncDecCTCModel.from_pretrained(model_name="stt_en_quartznet15x5").cuda()
# Punctuation and capitalization model
punctuation = nemo_nlp.models.PunctuationCapitalizationModel.from_pretrained(model_name='punctuation_en_distilbert').cuda()
# Spectrogram generator which takes text as an input and produces spectrogram
spectrogram_generator = nemo_tts.models.FastPitchModel.from_pretrained(model_name="tts_en_fastpitch").cuda()
# Vocoder model which takes spectrogram and produces actual audio
vocoder = nemo_tts.models.HifiGanModel.from_pretrained(model_name="tts_hifigan").cuda()
# + [markdown] colab_type="text" id="jQSj-IhEhrtI"
# ## Using the models
# + colab={} colab_type="code" id="s0ERrXIzKpwu"
# Convert our audio sample to text
files = [Audio_sample]
raw_text = ''
text = ''
for fname, transcription in zip(files, quartznet.transcribe(paths2audio_files=files)):
raw_text = transcription
# Add capitalization and punctuation
res = punctuation.add_punctuation_capitalization(queries=[raw_text])
text = res[0]
print(f'\nRaw recognized text: {raw_text}. \nText with capitalization and punctuation: {text}')
# + colab={} colab_type="code" id="-0Sk0C9-LmAR"
# A helper function which combines TTS models to go directly from
# text to audio
def text_to_audio(text):
parsed = spectrogram_generator.parse(text)
spectrogram = spectrogram_generator.generate_spectrogram(tokens=parsed)
audio = vocoder.convert_spectrogram_to_audio(spec=spectrogram)
return audio.to('cpu').detach().numpy()
# + [markdown] colab_type="text" id="Q8Jvwe4Ahncx"
# ## Results
# + colab={} colab_type="code" id="-im5TDF-MP2N"
# This is our original audio sample
IPython.display.Audio(Audio_sample)
# + colab={} colab_type="code" id="SNOMquwviEEQ"
# This is what was recognized by the ASR model
print(raw_text)
# + colab={} colab_type="code" id="6qRpDPfNiLOU"
# This is how punctuation model changed it
print(text)
# + [markdown] colab_type="text" id="di2IzMsdiiWq"
# Compare how the synthesized audio sounds when using text with and without punctuation.
# + colab={} colab_type="code" id="EIh8wTVs5uH7"
# Without punctuation
IPython.display.Audio(text_to_audio(raw_text), rate=22050)
# + colab={} colab_type="code" id="_qgKa9L954bJ"
# Final result - with punctuation
IPython.display.Audio(text_to_audio(text), rate=22050)
# + [markdown] colab_type="text" id="JOEFYywbctbJ"
# ## Next steps
# A demo like this is great for prototyping and experimentation. However, for real production deployment, you would want to use a service like [NVIDIA Riva](https://developer.nvidia.com/riva).
#
# **NeMo is built for training.** You can fine-tune, or train from scratch on your data all models used in this example. We recommend you checkout the following, more in-depth, tutorials next:
#
# * [NeMo fundamentals](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/00_NeMo_Primer.ipynb)
# * [NeMo models](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/01_NeMo_Models.ipynb)
# * [Speech Recognition](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/asr/ASR_with_NeMo.ipynb)
# * [Punctuation and Capitalization](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/nlp/Punctuation_and_Capitalization.ipynb)
# * [Speech Synthesis](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/tts/Inference_ModelSelect.ipynb)
#
#
# You can find scripts for training and fine-tuning ASR, NLP and TTS models [here](https://github.com/NVIDIA/NeMo/tree/main/examples).
# + [markdown] colab_type="text" id="ahRh2Y0Lc0G1"
# That's it folks! Head over to NeMo GitHub for more examples: https://github.com/NVIDIA/NeMo
|
tutorials/VoiceSwapSample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
ageinc_df = pd.read_csv('ageinc.csv')
ageinc_df['z_income'] = (ageinc_df['income'] - ageinc_df['income'].mean())/ageinc_df['income'].std()
ageinc_df['z_age'] = (ageinc_df['age'] - ageinc_df['age'].mean())/ageinc_df['age'].std()
# +
from sklearn import cluster
model = cluster.KMeans(n_clusters=2, random_state=10)
X = ageinc_df[['z_income','z_age']].as_matrix()
cluster_assignments = model.fit_predict(X)
centers = model.cluster_centers_
# +
import numpy as np
print(np.sum((X - centers[cluster_assignments]) ** 2))
# +
import matplotlib.pyplot as plt
# %matplotlib inline
ss = []
krange = list(range(2,11))
X = ageinc_df[['z_income','z_age']].values
for n in krange:
model = cluster.KMeans(n_clusters=n, random_state=10)
model.fit_predict(X)
cluster_assignments = model.labels_
centers = model.cluster_centers_
ss.append(np.sum((X - centers[cluster_assignments]) ** 2))
plt.plot(krange, ss)
plt.xlabel("$K$")
plt.ylabel("Sum of Squares")
plt.show()
# -
|
Lesson04/Exercise 15.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Day 8 - Part One
from aocd import get_data
raw_data = get_data(day=8, year=2021)
data = [x for x in raw_data.strip().split('\n')]
sep_parts = [x.split('|') for x in data]
signals = [x[0].strip() for x in sep_parts]
outputs = [x[1].strip() for x in sep_parts]
digit_counts = {}
for output in outputs:
for part in output.split():
if len(part) not in digit_counts:
digit_counts[len(part)] = 0
digit_counts[len(part)] += 1
print(sum([digit_counts[2], digit_counts[4], digit_counts[3], digit_counts[7]]))
# # Day 8 - Part Two
#
# This one was really difficult and required a "creative" solution ( :/ ).
# +
LENGTH_TO_DIGIT = {
2: (1, ),
3: (7, ),
4: (4, ),
5: (2, 3, 5),
6: (0, 6, 9),
7: (8, )
}
sep_parts = [x.split('|') for x in data]
signals = [x[0].strip() for x in sep_parts]
outputs = [x[1].strip() for x in sep_parts]
# +
output_total = 0
for (signal, output) in zip(signals, outputs):
candidates = [[] for _ in range(10)]
digit_letters = {key: set() for key in range(10)}
for part in signal.split():
for digit in LENGTH_TO_DIGIT[len(part)]:
candidates[digit].append(set(part))
# 1, 4, 7, and 8 have unique numbers of segments
digit_letters[1] = candidates[1][0]
digit_letters[4] = candidates[4][0]
digit_letters[7] = candidates[3][0]
digit_letters[8] = candidates[7][0]
# 2, 5, 6 must not contain both segments of 1
for i in (2, 5, 6):
candidates[i] = [c for c in candidates[i] if not digit_letters[1].issubset((c))]
# The intersection of 5 and 4 must have length 3
candidates[5] = [c for c in candidates[5] if len(c & candidates[4][0]) == 3]
# Since 2 now contains either 2 or 5, remove the intersection between 2 and 5
candidates[2] = [c for c in candidates[2] if c & candidates[5][0] != c]
# Take 4 chunk out of 9 and check if length is 2
candidates[9] = [c for c in candidates[9] if len(c - candidates[4][0]) == 2]
# Take 9 chunk out of 3 and check if empty; also remove 5 since we know it
candidates[3] = [c for c in candidates[3] if not(c - candidates[9][0]) and not(c == candidates[5][0])]
# Remove candidates for 6 and 9 from 0
candidates[0] = [c for c in candidates[0] if not(c == candidates[6][0]) and not(c == candidates[9][0])]
digits_list = []
for output_part in output.split():
output_part = set(output_part)
for d in range(len(candidates)):
if output_part == candidates[d][0]:
digits_list.append(d)
break
this_sum = 0
for digit, weight in zip(digits_list, iter([1000, 100, 10, 1])):
this_sum += digit * weight
output_total += this_sum
# -
for i, j in zip(['a', 'b', 'c', 'd'], iter([1000, 100, 10, 1])):
print(i, j)
print(signal, '|', output)
print(output_total)
|
2021/Day08.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## IMPORT LIBRARIES
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import altair as alt
import seaborn as sns
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_val_score
from sklearn import preprocessing
from sklearn.feature_selection import RFE
# Model Building
from sklearn.neighbors import KNeighborsClassifier
# Model Validation
from sklearn.model_selection import KFold
from sklearn.model_selection import LeaveOneOut
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
df = pd.read_csv('E:\ExcelR Assignment\Assignment 13 - KNN\Zoo.csv')
df.head()
df.info()
df.isnull().sum()
df.describe()
df.shape
df.columns
df['type'].value_counts().plot(kind='bar')
# ## 1. Data Analysis & Data Visualization
df.head()
df1 = df.copy()
df1 = df1.drop(['animal name'],axis=1)
df1.head()
# ## Splitting the variables
X = df1.iloc[0:91,0:-1]
Y = df1.iloc[0:91:,-1]
# ## Up-sampling as the Target Variable is not balanced
import smote_variants as sv
oversampler= sv.MulticlassOversampling(sv.distance_SMOTE())
X_samp, y_samp= oversampler.sample(X, Y)
# ## Train-Test Split Model Validation Technique
X_train, X_test, y_train, y_test = train_test_split(X,Y,test_size=0.2,random_state=42,stratify=Y)
# ## Transforming Variables
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.fit_transform(X_test)
from yellowbrick.features import Rank1D
visualizer = Rank1D(algorithm='shapiro')
visualizer.fit(X_train, y_train) # Fit the data to the visualizer
visualizer.transform(X_train) # Transform the data
visualizer.show()
# ## Build KNN model
kmodel = KNeighborsClassifier()
param_grid = [{'n_neighbors':range(2,20)}]
gsv = GridSearchCV(kmodel,param_grid)
gsv.fit(X_train,y_train)
gsv.best_params_,gsv.best_score_
result1 = []
result2 = []
for n in range(2,20):
model = KNeighborsClassifier(n_neighbors=n,metric='euclidean')
model.fit(X_train,y_train)
result1.append(model.score(X_train,y_train))
result2.append(model.score(X_test,y_test))
frame = pd.DataFrame({'n_neighbors':range(2,20),'Train Accuracy':result1,'Test Accuracy':result2})
frame
plt.plot(frame['n_neighbors'],frame['Train Accuracy'],marker='o',linestyle='dashed')
plt.plot(frame['n_neighbors'],frame['Test Accuracy'],marker='o')
model = KNeighborsClassifier(n_neighbors=13,metric='euclidean')
model.fit(X_train,y_train)
result1 = model.score(X_train,y_train)
result2 = model.score(X_test,y_test)
result1,result2
x_val = df.iloc[-10:,1:-1]
y_val = df.iloc[-10:,-1]
model.score(x_val,y_val)
|
KNN/Assignment 13 Part 2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PODPAC Introduction
#
# *Author: Creare* <br>
# *Date: April 01 2020* <br>
#
# **Keywords**: podpac
# ## Overview
#
# This notebook provides a high level overview of the PODPAC library.
# ### Prerequisites
#
# - Python 2.7 or above
# - [`podpac`](https://podpac.org/install.html#install)
# - *Review the [README.md](../../README.md) and [jupyter-tutorial.ipynb](../jupyter-tutorial.ipynb) for additional info on using jupyter notebooks*
# ### See Also
#
# - [python/basic-python.ipynb](../python/basic-reference.ipynb): Basic introduction to Python language features
# - [python/matlab.ipynb](../python/matlab.ipynb): Introduction to Python for MATLAB users
# - [xarray](xarray.ipynb): Short reference for the core [`xarray`](https://xarray.pydata.org/en/stable/) module.
# + [markdown] slideshow={"slide_type": "slide"}
# # Importing modules
#
# PODPAC has multiple modules, which can be imported all at once, or individually:
# -
import podpac # Import PODPAC with the namespace 'podpac'
import podpac as pc # Import PODPAC with the namespace 'pc'
from podpac import Coordinates # Import Coordinates from PODPAC into the main namespace
# + [markdown] slideshow={"slide_type": "slide"}
# # PODPAC library structure
# PODPAC is composed out of multiple sub-modules/sub-libraries. The major ones, from a user's perspective are shown below.
# <img src='../../images/podpac-user-api.png' style='width:80%; margin-left:auto;margin-right:auto;' />
#
#
# We can examine what's in the PODPAC library by using the `dir` function
# + slideshow={"slide_type": "subslide"}
dir(podpac)
# + [markdown] slideshow={"slide_type": "subslide"}
# In PODPAC, the top-level classes and functions are frequently used and include:
#
# * `Coordinates`: class for defining coordinates
# * `Node`: Base class for defining PODPAC compute Pipeline
# * `NodeException`: The error type thrown by Nodes
# * `clinspace`: A helper function used to create uniformly spaced coordinates based on the number of points
# * `crange`: Another helper function used to create uniformly spaced coordinates based on step size
# * `settings`: A module with various settings that define caching behavior, login credentials, etc.
# * `version_info`: Python dictionary giving the version of the PODPAC library
# + [markdown] slideshow={"slide_type": "subslide"}
# The top-level modules or sub-packages (or sub libraries) include:
# * `algorithm`: here you can find generic `Algorithm` nodes to do different types of computations
# * `authentication`: this contains utilities to help authenticate users to download data
# * `compositor`: here you can find nodes that help to combine multiple data sources into a single node
# * `coordinates`: this module contains additional utilities related to creating coordinates
# * `core`: this is where the core library is implemented, and follows the directory structure of the code
# * `data`: here you can find generic `DataSource` nodes for reading and interpreting data sources
# * `datalib`: here you can find domain-specific `DataSource` nodes for reading data from specific instruments, studies, and programs
# * `interpolators`: this contains classes for dealing with automatic interpolation
# * `pipeline`: this contains generic `Pipeline` nodes which can be used to share and re-create PODPAC processing routines
#
# Diving into specifically what's available in some of these submodules
# + slideshow={"slide_type": "subslide"}
# Generic Algorithm nodes
dir(podpac.algorithm)
# + slideshow={"slide_type": "subslide"}
# Generic DataSource nodes
dir(podpac.data)
# + slideshow={"slide_type": "subslide"}
# Specific data libraries built into podpac
import podpac.datalib # not loaded by default
dir(podpac.datalib)
# + slideshow={"slide_type": "skip"}
# Algorithms to compute climatology -- used for computing beta distributions of soil moisture for the drought-monitor application
import podpac.alglib
dir(podpac.alglib)
|
notebooks/0-concepts/introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ungraded Lab: Coding a Wide and Deep Model
#
# In this lab, we'll show how you can implement a wide and deep model. We'll first look at how to build it with the Functional API then show how to encapsulate this into a class. Let's get started!
# ## Imports
# + colab={} colab_type="code" id="CmI9MQA6Z72_"
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow.keras.layers import Dense
from tensorflow.keras import Model
from tensorflow.keras.layers import concatenate
from tensorflow.keras.layers import Input
from tensorflow.keras.utils import plot_model
# + [markdown] colab={} colab_type="code" id="8RKbMogoaHvc"
# ## Build the Model
#
# Let's implement the wide and deep model as shown in class. As shown below, the Functional API is very flexible in implementing complex models.
# - You will specify the previous layer when you define a new layer.
# - When you define the `Model`, you will specify the inputs and output.
# + colab={} colab_type="code" id="Uz4pA6uEucZ8"
# define inputs
input_a = Input(shape=[1], name="Wide_Input")
input_b = Input(shape=[1], name="Deep_Input")
# define deep path
hidden_1 = Dense(30, activation="relu")(input_b)
hidden_2 = Dense(30, activation="relu")(hidden_1)
# define merged path
concat = concatenate([input_a, hidden_2])
output = Dense(1, name="Output")(concat)
# define another output for the deep path
aux_output = Dense(1,name="aux_Output")(hidden_2)
# build the model
model = Model(inputs=[input_a, input_b], outputs=[output, aux_output])
# visualize the architecture
plot_model(model)
# -
# ## Implement as a Class
#
# Alternatively, you can also implement this same model as a class.
# - For that, you define a class that inherits from the [Model](https://keras.io/api/models/model/) class.
# - Inheriting from the existing `Model` class lets you use the Model methods such as `compile()`, `fit()`, `evaluate()`.
#
# When inheriting from `Model`, you will want to define at least two functions:
# - `__init__()`: you will initialize the instance attributes.
# - `call()`: you will build the network and return the output layers.
#
# If you compare the two methods, the structure is very similar, except when using the class, you'll define all the layers in one function, `init`, and connect the layers together in another function, `call`.
# + colab={} colab_type="code" id="NwyCp57qqdXS"
# inherit from the Model base class
class WideAndDeepModel(Model):
def __init__(self, units=30, activation='relu', **kwargs):
'''initializes the instance attributes'''
super().__init__(**kwargs)
self.hidden1 = Dense(units, activation=activation)
self.hidden2 = Dense(units, activation=activation)
self.main_output = Dense(1)
self.aux_output = Dense(1)
def call(self, inputs):
'''defines the network architecture'''
input_A, input_B = inputs
hidden1 = self.hidden1(input_B)
hidden2 = self.hidden2(hidden1)
concat = concatenate([input_A, hidden2])
main_output = self.main_output(concat)
aux_output = self.aux_output(hidden2)
return main_output, aux_output
# + colab={} colab_type="code" id="KVOkjlgwuD_9"
# create an instance of the model
model = WideAndDeepModel()
|
Custom Models, Layers, and Loss Functions with TensorFlow/Week 4 Custom Models/Lab_1_basic-model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualization
# PySwarms implements tools for visualizing the behavior of your swarm. These are built on top of `matplotlib`, thus rendering charts that are easy to use and highly-customizable. However, it must be noted that in order to use the animation capability in PySwarms (and in `matplotlib` for that matter), at least one writer tool must be installed. Some available tools include:
# * ffmpeg
# * ImageMagick
# * MovieWriter (base)
#
# In the following demonstration, the `ffmpeg` tool is used. For Linux and Windows users, it can be installed via:
# ```shell
# $ conda install -c conda-forge ffmpeg
# ```
import sys
sys.path.append('../')
# First, we need to import the `pyswarms.utils.environments.PlotEnvironment` class. This enables us to use various methods to create animations or plot costs.
# +
# Import modules
import matplotlib.pyplot as plt
import numpy as np
from matplotlib import animation, rc
from IPython.display import HTML
# Import PySwarms
import pyswarms as ps
from pyswarms.utils.functions import single_obj as fx
from pyswarms.utils.environments import PlotEnvironment
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# -
# The first step is to create an optimizer. Here, we're going to use Global-best PSO to find the minima of a sphere function. As usual, we simply create an instance of its class `pyswarms.single.GlobalBestPSO` by passing the required parameters that we will use.
options = {'c1':0.5, 'c2':0.3, 'w':0.9}
optimizer = ps.single.GlobalBestPSO(n_particles=10, dimensions=3, options=options)
# ## Initializing the `PlotEnvironment`
#
# Think of the `PlotEnvironment` as a container in which various plotting methods can be called. In order to create an instance of this class, we need to pass the optimizer object, the objective function, and the number of iterations needed. The `PlotEnvironment` will then simulate these parameters so as to build the plots.
plt_env = PlotEnvironment(optimizer, fx.sphere_func, 1000)
# ## Plotting the cost
#
# To plot the cost, we simply need to call the `plot_cost()` function. There are pre-set defaults in this method already, but we can customize by passing various arguments into it such as figure size, title, x- and y-labels and etc. Furthermore, this method also accepts a keyword argument `**kwargs` similar to `matplotlib`. This enables us to further customize various artists and elements in the plot.
#
# For now, let's stick with the default one. We'll just call the `plot_cost()` and `show()` it.
plt_env.plot_cost(figsize=(8,6));
plt.show()
# ## Animating swarms
# The `PlotEnvironment()` offers two methods to perform animation, `plot_particles2D()` and `plot_particles3D()`. As its name suggests, these methods plot the particles in a 2-D or 3-D space. You can choose which dimensions will be plotted using the `index` argument, but the default takes the first 2 (or first three in 3D) indices of your swarm dimension.
#
# Each animation method returns a `matplotlib.animation.Animation` class that still needs to be animated by a `Writer` class (thus necessitating the installation of a writer module). For the proceeding examples, we will convert the animations into an HTML5 video. In such case, we need to invoke some extra methods to do just that.
# equivalent to rcParams['animation.html'] = 'html5'
# See http://louistiao.me/posts/notebooks/save-matplotlib-animations-as-gifs/
rc('animation', html='html5')
# ### Plotting in 2-D space
#
HTML(plt_env.plot_particles2D(limits=((-1.2,1.2),(-1.2,1.2))).to_html5_video())
# ### Plotting in 3-D space
HTML(plt_env.plot_particles3D(limits=((-1.2,1.2),(-1.2,1.2),(-1.2,1.2))).to_html5_video())
|
examples/visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: saturn (Python 3)
# language: python
# name: python3
# ---
# # GPU random forest with data from Snowflake
# <table>
# <tr>
# <td>
# <img src="https://saturn-public-assets.s3.us-east-2.amazonaws.com/example-resources/rapids.png" width="300">
# </td>
# <td>
# <img src="https://saturn-public-assets.s3.us-east-2.amazonaws.com/example-resources/snowflake.png" width="300">
# </td>
# </tr>
# </table>
# This notebook describes a machine learning training workflow using the famous [NYC Taxi Dataset](https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page). That dataset contains information on taxi trips in New York City.
#
# In this exercise, you'll use `cudf` to load a subset of the data from Snowflake and `cuml` to answer this classification question:
#
# > based on characteristics that can be known at the beginning of a trip, will this trip result in a high tip?
# ## Use RAPIDS libraries
#
# RAPIDS is a collection of libraries which enable you to take advantage of NVIDIA GPUs to accelerate machine learning workflows. This exercise uses the following RAPIDS packages:
#
# * [`cudf`](https://github.com/rapidsai/cudf): data frame manipulation, similar to `pandas` and `numpy`
# * [`cuml`](https://github.com/rapidsai/cuml): machine learning training and evaluation, similar to `scikit-learn`
#
# For more information on RAPIDS, see ["Getting Started"](https://rapids.ai/start.html) in the RAPIDS docs.
#
# ### Monitor Resource Usage
#
# This tutorial aims to teach you how to take advantage of the GPU for data science workflows. To prove to yourself that RAPIDS is utilizing the GPU, it's important to understand how to monitor that utilization while your code is running. If you already know how to do that, skip to the next section.
#
# <details><summary>(click here to learn how to monitor resource utilization)</summary>
#
# <br>
#
# **Monitoring CPU and Main Memory**
#
# CPUs and GPUs are two different types of processors, and the GPU has its own dedicated memory. Many data science libraries that claim to offer GPU acceleration accomplish their tasks with a mix of CPU and GPU use, so it's important to monitor both to see what that code is doing.
#
# To monitor CPU utilization and the amount of free main memory (memory available to the CPU), you can use `htop`.
#
# Open a new terminal and run `htop`. That will keep an auto-updating dashboard up that shows the CPU utilization and memory usage.
#
# **Monitoring GPU and GPU memory**
#
# Open a new terminal and run the following command.
#
# ```shell
# watch -n 5 nvidia-smi
# ```
#
# <br>
# This command will update the output in the terminal every 5 seconds. It shows some information like:
#
# * current CUDA version
# * NVIDIA driver version
# * internal temperature
# * current utilization of GPU memory
# * list of processes (if any) currently running on the GPU, and how much GPU memory they're consuming
#
# If you'd prefer a simpler view, you can also consider `gpustat`, which simply tracks temperature, GPU utilization, and memory. This is not available by default in the Saturn GPU images, but you can install it from PyPi.
#
# ```shell
# pip install gpustat
# ```
#
# <br>
#
# And then run it
#
# ```shell
# gpustat -cp --watch
# ```
#
# <br>
#
# Whichever option you choose, leave these terminals with the monitoring process running while you work, so you can see how the code below uses the available resources.
#
# </details>
# <hr>
#
# ## Connect to Snowflake
#
# This example uses data stored in a Snowflake data warehouse that is managed by the team at Saturn Cloud. We've set up a read-only user for use in these examples. If you would like to access data stored in your own Snowflake account, you should set up [Credentials](https://saturncloud.io/docs/concepts/credentials/) for your account, user, and password then set the other connection information accordingly. For more details on Snowflake connection information, see ["Connecting to Snowflake"](https://docs.snowflake.com/en/user-guide/python-connector-example.html#connecting-to-snowflake) in the `snowflake-connector-python` docs.
#
# Note that in order to update environment variables your Jupyter server will need to be stopped.
# +
import os
import cudf
import pandas as pd
import snowflake.connector
conn_info = {
"account": os.environ["EXAMPLE_SNOWFLAKE_ACCOUNT"],
"user": os.environ["EXAMPLE_SNOWFLAKE_USER"],
"password": <PASSWORD>["<PASSWORD>"],
"database": os.environ["TAXI_DATABASE"],
}
conn = snowflake.connector.connect(**conn_info)
# -
# > Don't worry if you see a warning about an incompatible version of pyarrow installed. This is because `snowflake.connector` relies on a older version of pyarrow for certain methods. We won't use those methods here so it's not a problem!
#
# ## Load data
#
# This example is designed to run quickly with small resources. So let's just load a single month of taxi data for training.
#
# The code below loads the data into a `cudf` data frame. This is similar to a `pandas` dataframe, but it lives in GPU memory and most operations on it are done on the GPU.
#
# This example uses Snowflake to handle the hard work of creating new features, then creates a `cudf` data frame with the result.
query = """
SELECT
pickup_taxizone_id,
dropoff_taxizone_id,
passenger_count,
DIV0(tip_amount, fare_amount) > 0.2 AS high_tip,
DAYOFWEEKISO(pickup_datetime) - 1 AS pickup_weekday,
WEEKOFYEAR(pickup_datetime) AS pickup_weekofyear,
HOUR(pickup_datetime) AS pickup_hour,
(pickup_weekday * 24) + pickup_hour AS pickup_week_hour,
MINUTE(pickup_datetime) AS pickup_minute
FROM taxi_yellow
WHERE
DATE_TRUNC('MONTH', pickup_datetime) = '{day}'
"""
taxi = pd.read_sql(query.format(day="2019-01-01"), conn)
taxi.columns = taxi.columns.str.lower()
taxi = cudf.from_pandas(taxi)
# +
numeric_feat = [
"pickup_weekday",
"pickup_weekofyear",
"pickup_hour",
"pickup_week_hour",
"pickup_minute",
"passenger_count",
]
categorical_feat = [
"pickup_taxizone_id",
"dropoff_taxizone_id",
]
features = numeric_feat + categorical_feat
y_col = "high_tip"
taxi_train = taxi[features + [y_col]]
taxi_train[features] = taxi_train[features].astype("float32").fillna(-1)
taxi_train[y_col] = taxi_train[y_col].astype("int32").fillna(-1)
# -
# The code below computes the size of this dataset in memory.
print(f"Num rows: {len(taxi_train)}, Size: {taxi_train.memory_usage(deep=True).sum() / 1e6} MB")
# You can examine the structure of the data with `cudf` commands:
#
# `.head()` = view the first few rows
taxi_train.head()
# `.dtypes` = list all the columns and the type of data in them
taxi_train.dtypes
# <hr>
#
# ## Train a Model
#
# Now that the data have been prepped, it's time to build a model!
#
# For this task, we'll use the `RandomForestClassifier` from `cuml`. If you've never used a random forest or need a refresher, consult ["Forests of randomized trees"](https://scikit-learn.org/stable/modules/ensemble.html#forest) in the `sciki-learn` documentation.
#
# The code below initializes a random forest classifier with the following parameter values.
#
# * `n_estimators=100` = create a 100-tree forest
# * `max_depth=10` = stop growing a tree once it contains a leaf node that is 10 levels below the root
# * `n_streams=4` = create 4 decision trees at a time
# - setting this to a value higher than 1 can reduce training time, but setting it too high can increase training time
# - increasing this parameter's value increases the memory requirements for training
#
# All other parameters use the defaults from `RandomForestClassifier`.
#
# <details><summary>(click here to learn why data scientists do this)</summary>
#
# **Setting max_depth**
#
# Tree-based models split the training data into smaller and smaller groups, to try to group together records with similar values of the target. A tree can be thought of as a collection of rules like `pickup_hour greater than 11` and `pickup_minute less than 31.0`. As you add more rules, those groups (called "leaf nodes") get smaller. In an extreme example, a model could create a tree with enough rules to place each record in the training data into its own group. That would probably take a lot of rules, and would be referred to as a "deep" tree.
#
# Deep trees are problematic because their descriptions of the world are too specific to be useful on new data. Imagine training a classification model to predict whether or not visitors to a theme park will ride a particular rollercoaster. You could measure the time down to the millisecond that every guest's ticket is scanned at the entrance, and a model might learn a rule like *"if the guest has been to the park before and if the guest is older than 40 and younger than 41, and if the guest is staying at Hotel A and if the guest enters the park after 1:00:17.456 and if the guest enters the park earlier than 1:00:17.995, they will ride the rollercoaster"*. This is very very unlikely to ever match any future visitors, and if it does it's unlikely that this prediction will be very good unless you have some reason to believe that a visitor arriving at 1:00:18 instead of 1:00:17 really changes the probability that they'll ride that rollercoaster.
#
# To prevent this situation (called "overfitting"), most tree-based machine learning algorithms accept parameters that control how deep the trees can get. `max_depth` is common, and says "don't create a rule more complex than this". In the example above, that rule has a depth of 7.
#
# 1. visiting the park
# 2. has been to the park before?
# 3. older than 40?
# 4. younger than 41?
# 5. staying at Hotel A?
# 6. entered the park after 1:00:17.456?
# 7. entered the park before 1:00:17.995?
#
# Setting `max_depth = 5` would have prevented those weirdly-specific timing rules from ever being generated.
#
# Choosing good values for this parameter is part art, part science, and is outside the scope of this tutorial.
#
# </details>
# +
from cuml.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=100, max_depth=10, n_streams=4)
# -
# With the classifier created, fit it to some data! The code below uses `%%time` to print out a timing, so you can see how long it takes to train. This can be used to compare `cuml` to methods explored in other notebooks, or to test how changing some parameters to `RandomForestClassifier` changes the runtime for training.
# %%time
_ = rfc.fit(taxi_train[features], taxi_train[y_col])
# <hr>
#
# ## Save model
#
# Once you've trained a model, save it in a file to use later for scoring or for comparison with other models.
#
# There are several ways to do this, but `cloudpickle` is likely to give you the best experience. It handles some common drawbacks of the built-in `pickle` library.
#
# `cloudpickle` can be used to write a Python object to bytes, and to create a Python object from that binary representation.
# +
import cloudpickle
import os
MODEL_PATH = "models"
if not os.path.exists(MODEL_PATH):
os.makedirs(MODEL_PATH)
with open(f"{MODEL_PATH}/random_forest_rapids.pkl", "wb") as f:
cloudpickle.dump(rfc, f)
# -
# <hr>
#
# ## Calculate metrics on test set
#
# Machine learning training tries to create a model which can produce useful results on new data that it didn't see during training. To test how well we've accomplished that in this example, read in another month of taxi data from Snowflake.
# +
taxi_test = pd.read_sql(query.format(day="2019-02-01"), conn)
taxi_test.columns = taxi_test.columns.str.lower()
taxi_test = cudf.from_pandas(taxi_test)
taxi_test[features] = taxi_test[features].astype("float32").fillna(-1)
taxi_test[y_col] = taxi_test[y_col].astype("int32").fillna(-1)
# -
# `cuml` comes with many functions for calculating metrics that describe how well a model's predictions match the actual values. For a complete list, see [the cuml API docs](https://docs.rapids.ai/api/cuml/stable/api.html#metrics-regression-classification-and-distance) or run the code below.
# +
import cuml.metrics
[m for m in dir(cuml.metrics) if not m.startswith("_")]
# -
# This tutorial uses the `roc_auc_score` to evaluate the model. This metric measures the area under the [receiver operating characteristic](https://en.wikipedia.org/wiki/Receiver_operating_characteristic) curve. Values closer to 1.0 are desirable.
# +
from cuml.metrics import roc_auc_score
preds = rfc.predict_proba(taxi_test[features])[1]
roc_auc_score(taxi_test[y_col], preds)
# -
# <hr>
#
# ## Next Steps
#
# In this tutorial, you learned how to train a model for a binary classification task, using `cuml`, based on data in Snowflake. Training took around 5 seconds for a dataset that was 0.31 GB in memory, a huge improvement on the almost 9 minutes it took to [train the same model using `scikit-learn`](./rf-scikit.ipynb)!
#
# If you wanted to train a much larger model (think `max_depth=16, num_iterations=10000`) or use a much larger dataset or both, it might not be possible on a single machine. Try [this dask-cudf notebook](./rf-rapids-dask.ipynb) to learn how to use Dask to take advantage of multiple-machine, multi-GPU training.
#
# <hr>
|
examples/snowflake/advanced/rf-rapids.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Hello, and welcome to this tutorial on simulating circuit noise in Tequila! In this tutorial, we will briefly detail what quantum noise is, its mathematical modeling, and how specific popular simulation packages handle noisy simulation, before diving in to building Tequila `NoiseModel`s and applying them to sampling different circuits.
# # <center> What is noise?</center>
#
# In case you need a quick refresher: Real quantum systems undergo the effects of noise, a catch-all term for 'anything the user didn't ask the computer to do'. Such noise can be caused by a number of physical processes, including but not limited to:
#
# - **Thermal fluctuations**
# - **Interaction with the environment**
# - **Uncontrolled interaction between qubits (cross-talk)**
# - **Imperfections in gate implementation**
#
#
#
# # <center> How is noise represented mathematically?</center>
#
#
# Commonly, the effects of noise on quantum systems are treated as the evolution of the system's density matrix under Krauss maps. Krauss maps are mappings of the form $ A: \rho \rightarrow \rho' = \sum_{i} A_i \rho A_{i}^{\dagger}$, where $\sum_{i} A_i A_{i}^{\dagger} = I$. These Krauss maps are parametrized, in general by probabilities.
#
# For example, bit flip noise -- which takes qubits from the 0 to the 1 state and vice versa-- is a krauss map with two operators, and a single probabilistic parameter, p. The operators are:
# $$A_0 = \sqrt{1-p} I, A_1 = \sqrt{p} X$$
# **Note that the square root is present, so that bit flip map is:**
# $$ A_{bf}(p): \rho \rightarrow (1-p) * I\rho I + p * X\rho X$$
# other noise operations may be defined similarly.
#
# **Note that such krauss operators may only affect subsystems of the system;** one can have a single qubit undergo bit-flip noise in an 8 qubit state. In such cases, the krauss maps are merely the 1-qubit maps tensored with the identity on all other qubits. Multi-qubit krauss operators will involve tensor products of single qubit krauss operators.
#
# For example, the 2-qubit bit flip krauss map has 4 operators:
# $$A_{00}=(1-p)I\otimes I, A_{01}=\sqrt{p-p^2}I\otimes X,A_{10}=\sqrt{p-p^2}X \otimes I,A_{11}=pX\otimes X$$
# Which are just all the tensor products of $A_{0}$ and $A_{1}$.
# # <center> 3. How is noise simulated? </center>
#
# Different simulation packages handle noise in radically different ways.
#
# *Cirq* and *Qulacs*, for example, use noise channels, parametrized operations which are inserted into circuits the same way regular, unitary gates are.
#
# *Pyquil* asks its users to define noisy gate operations, and then instantiate those.
#
# *Qiskit*, meanwhile, simply takes a dictionary-like object as an argument to its simulator, and applies the noise on the user-chosen gates.
#
# In tequila, we try to hew toward making users write as few lines of code as possible. We therefore implement a simple framework for the application of noise, meant to be compatible with all our supported platforms. To do this, we make a few assumptions:
#
# 1. If noise is present, any gate may be affected by noise.
# 2. The noise that affects 1..k..n-qubit gates is independent of the noise on 1...k-1,k+1...n qubit gates.
# 3. Noise probabilities are independent of position in the circuit.
# 4. The number of qubits involved in a gate, not the operation performed, dictates what noises may occur.
#
#
# # <center> Noise in Tequila: Overview </center>
#
# noise in Tequila is centered upon the `NoiseModel` class, itself used to store and combine `QuantumNoise` objects.
# Each `QuantumNoise` internally designates what operation it will perform, with what probability (or probabilities), and on how many qubits. Only at the time of translation to a backend -- or in the case of *Qiskit*, at time of simulation -- do`NoiseModel`s and simulateables -- circuits, ExpectationValues, Objectives -- interact.
#
# Tequila at present supports six common quantum noise operations, all of which can at present be employed by all the noise-supporting simulation backgrounds. These six operations are:
#
# 1. Bit flips, a probabilistic application of pauli X;
# 2. Phase flips, a probablistic application of pauli Z;
# 3. Amplitude damps, which take qubits in state |1> to |0>;
# 4. Phase damps, which are a different formalization of the phase flip;
# 5. Phase-Amplitude damps, which simultaneously perform said operations;
# 6. (Symmetric) depolarizing, which (equi)probabilistically performs pauli X, Y, and Z.
#
# in Tequila, custom members of the `QuantumNoise` class are not possible, and so they should be initialized by the constructor function for each supported channel, which creates a `NoiseModel` containing one operation. all six of which are shown in the import statement below.
#
# `NoiseModel`s combine with eachother through addition, creating a new `NoiseModel` with all the operations of the two summands. Note that in those simulators which employ noise channels, the order of the noises in the noise model will dictate the order of application in the circuit; users should be mindful of this.
#
# To use a `NoiseModel` to apply noise, one may provide a noise model to the *tq.compile*, *tq.simulate*, and optimization calls like *tq.minimize*, through the keyword `noise=my_noise_model`. Noise is only supported when sampling; if in the above functions the keyword *samples* is `None` (defaullt), noise cannot function.
#
# additionally, Tequila supports the use of device-noise-emulation for those backends which allow the emulation of specific real devices. If in compilation, simulation, or optimization, emulated backends have been selected (such as 'fake_vigo', for IBMQ), the known noise of this device may be employed. In these cases, the keyword assignment *noise='device'* should be used, if these known noise models are desired.
#
#
### first, we import tequila!
import tequila as tq
from tequila.circuit.noise import BitFlip,PhaseFlip,AmplitudeDamp,PhaseDamp,PhaseAmplitudeDamp,DepolarizingError
# We will first examine bit flip noise on a simple circuit with a simple Hamiltonian.
# +
H=tq.paulis.Qm(1) ### this hamiltonian is 0 for a qubit that is 0, and 1 for a qubit that is 1.
U=tq.gates.X(0)+tq.gates.CNOT(0,1)
O1=tq.ExpectationValue(U=U,H=H)
print('simulating: ',H)
print('acting on: ')
tq.draw(U)
# -
# Say that we wanted a noise model where 1-qubit gates and 2-qubit gates undergo bit flips, but with different probabilities.
bf_1=BitFlip(p=0.1,level=1)
bf_2=BitFlip(p=0.3,level=2)
# `NoiseModel` objects, like those initialized above, can be combined into new `NoiseModel`s by simple addition.
my_nm=bf_1+bf_2
print('applying:',my_nm)
# we will now sample our `Objective` O1, both with and without noise.
E=tq.simulate(O1)
### noise models are fed to tequila functions with the noise keyword.
E_noisy=tq.simulate(O1,samples=5000,noise=my_nm)
print('Without noise, E =',E)
print('With noise, E =',E_noisy)
# **Because noise is stochastic, results may vary wildly if the number of samples is low.**
for i in range(1,11):
print('round',i,'sampling with 5 samples, E = ', tq.simulate(O1,samples=5,noise=my_nm))
# Note that the *BitFlip* functions returned applicable `NoiseModel`s in their own right:
#
E_1_only=tq.simulate(O1,samples=1000,noise=bf_1)
print('With 1-qubit noise only, E =',E_1_only)
E_2_only=tq.simulate(O1,samples=1000,noise=bf_2)
print('With 2-qubit noise only, E =',E_2_only)
# Below, we demonstrate the effects of the ordering of the noise operations applied.
#
# +
amp=AmplitudeDamp(0.3,1)
bit=BitFlip(0.4,1)
forward=amp+bit
backward=bit+amp
H = tq.paulis.Z(0)
U = tq.gates.X(target=0)
O = tq.ExpectationValue(U=U, H=H)
E_1 = tq.simulate(O,samples=100000,noise=forward)
E_2 = tq.simulate(O,samples=100000,noise=backward)
print('amplitude damping before bit flip leads to E = ',E_1)
print('amplitude damping after bit flip leads to E = ',E_2)
# -
# Tequila will *always* attempt to apply noise to the circuit *in the order each noise was added to the noise model*. Some backends have behavior which is harder to control than others, but in general, this order will be preserved.
# Below, we will optimize a noisy circuit.
# +
import numpy as np
# -
# Consider the 1-qubit expectation value, $<0|U^{\dagger}\hat{Y}U|0>$, with $U=H Rz(\theta) H $. In the absence of noise, this expectation value just yields $Sin(\theta)$. This circuit therefore has a minimum at $\theta = -\pi$. We can minimize this circuit under phase flip noise -- which is probabilistic application of pauli Z -- and see what happens!
# +
U=tq.gates.H(0) +tq.gates.Rz('a',0)+tq.gates.H(0)
H=tq.paulis.Y(0)
O=tq.ExpectationValue(U=U,H=H)
### we pick a random, small probability to apply noise
p=np.random.uniform(0,.1)
NM=PhaseFlip(p,1)
print('optimizing expectation value with phase flip probability {}.'.format(str(p)))
result=tq.minimize(objective=O,lr=0.5,maxiter=60,initial_values={'a':np.pi},method='adam',samples=5000,noise=NM,silent=True)
result.history.plot()
# -
# The final energy is not -1.0, because the application of noise leads the expected output to be $(-1+2*p)^{3} Sin(\theta)$. One sees that this is approximately the value reached by minimizing $\theta$. Because the number of samples is not infinite, the 'expected' best energy may be exceeded:
out=result.energy
best=((-1+2*p)**3)*np.sin(np.pi/2)
print('best energy: ',out)
print('expected best ',best)
# ## This concludes our brief tutorial on Noise. Stay tuned (and up to date) for more exciting noise features in the future!
|
tutorials/Noise_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np
import os
import cv2
import matplotlib.pyplot as plt
from keras.models import load_model
import numpy as np
import pandas as pd
import os
import glob
import cv2
import random
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.optimizers import RMSprop
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Activation, Dropout, Flatten, Dense
from keras import regularizers
from keras.callbacks import CSVLogger
#from livelossplot import PlotLossesKeras
import os
import numpy as np
#from imgaug import augmenters as iaa
#import cv2
from keras.layers.normalization import BatchNormalization
#import seaborn as sns
import pandas as pd
from keras import initializers
from keras import optimizers
import keras.backend as K
import tensorflow as tf
import operator
# -
val_all_path = '../input/cmncthesis/C-NMC_test_final_phase_data'
val_all_list = os.listdir(val_all_path)
val_all_list.sort()
print('validation: ', len(val_all_list))
val_all_batch = np.zeros((len(val_all_list), 150, 150, 3), dtype=np.uint8)
print(val_all_batch.shape)
def Read_n_Crop(list_data, batch, path):
i=0
for x in list_data:
image = cv2.imread(os.path.join(path, x))
image = cv2.cvtColor(image,cv2.COLOR_BGR2RGB)
image = crop_center(image, (150,150,3))
batch[i] = image
i+=1
print(type(batch), batch.shape, batch.dtype, batch[0].shape, batch[0].dtype)
return batch
def crop_center(img, bounding):
start = tuple(map(lambda a, da: a//2-da//2, img.shape, bounding))
end = tuple(map(operator.add, start, bounding))
slices = tuple(map(slice, start, end))
return img[slices]
images=Read_n_Crop(val_all_list, val_all_batch, val_all_path)
images = images/255.0
model= None
# load the trained model
model = load_model('../input/effnetb0-5xaug-test8/EffnetB0_5xAug_test8.h5', compile=False)
model.summary()
predictions = model.predict(images, verbose = 10)
y_pred_flat = []
for pred in predictions:
if pred > 0.3:
y_pred_flat.append(0)
else:
y_pred_flat.append(1)
y_pred_flat = np.array(y_pred_flat)
print(len(y_pred_flat))
with open('isbi_valid.predict', 'w') as f:
for item in y_pred_flat:
f.write("%s\n" %item)
|
submission-5x.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SanchitaMahajan/ELP.Sem2_2021_20467876/blob/main/ELP_Ques2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="OK8U629C040h"
# # **Extended Learning Portfolio Ques 2:-**
#
# ### ***By <NAME>***
#
# ### ***20467876***
# + [markdown] id="Ydhd2LaaC9AZ"
# # **This is a function which is used to get the URL (search_term) :-**
# ***1. The User has to input an item in place of search_term..***
#
# ***2. The function will then return the Webpage of the search_term for the item provided by the user.***
#
#
#
#
# ---
#
#
#
#
# # **WORKING:-**
#
# Let's understand the following function using the Amazon URL example...
#
# **url** = 'https://www.amazon.com.au'
#
#
# **template(ultrawide monitor)** = 'https://www.amazon.com.au/s?k=ultrawide+monitor&ref=nb_sb_noss_2'
#
#
#
#
#
#
# ## ***After careful analysis we realised that, when we create a function to get url for Search_term:-***
#
# ---
#
#
#
# We get, **url** = 'https://www.amazon.com.au/s?k=ultrawide monitor&ref=nb_sb_noss_2'
#
#
#
# ---
#
#
# ***This is not a valid url,In the actual URL, We need '+' between ultrawide and monitor***
#
# `#replace spaces with a '+' sign`
#
# ## **So, we will replace ' ' (space) with a '+' sign !**
#
#
#
#
# ---
#
#
#
#
# ###***So,the result URL will be***:-
# 'https://www.amazon.com.au/s?k=ultrawide+monitor&ref=nb_sb_noss_2'
#
#
#
# + id="Yh8skwYs0pHf"
''' This function is used to get_url for
search_term & template '''
def get_url(search_term, template):
#replace spaces with a (' ', '+') sign
search_term = search_term.replace(' ', '+')
url = template.format(search_term)
# This will append all the pages of url and add the page code to get query parameter
url += '&page={}'
return url
# + id="LA3M3BpEBN0X"
|
ELP_Ques2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load data
#
# <https://www.kaggle.com/c/bike-sharing-demand>
import sage
import numpy as np
from sklearn.model_selection import train_test_split
# Load data
df = sage.datasets.bike()
feature_names = df.columns.tolist()[:-3]
# Split data, with total count serving as regression target
train, test = train_test_split(
df.values, test_size=int(0.1 * len(df.values)), random_state=123)
train, val = train_test_split(
train, test_size=int(0.1 * len(df.values)), random_state=123)
Y_train = train[:, -1].copy()
Y_val = val[:, -1].copy()
Y_test = test[:, -1].copy()
train = train[:, :-3].copy()
val = val[:, :-3].copy()
test = test[:, :-3].copy()
# # Train model
import xgboost as xgb
# +
# Set up data
dtrain = xgb.DMatrix(train, label=Y_train)
dval = xgb.DMatrix(val, label=Y_val)
# Parameters
param = {
'max_depth' : 10,
'objective': 'reg:squarederror',
'nthread': 4
}
evallist = [(dtrain, 'train'), (dval, 'val')]
num_round = 50
# Train
model = xgb.train(param, dtrain, num_round, evallist, verbose_eval=False)
# +
# Calculate performance
mean = np.mean(Y_train)
base_mse = np.mean((mean - Y_test) ** 2)
mse = np.mean((model.predict(xgb.DMatrix(test)) - Y_test) ** 2)
print('Base rate MSE = {:.2f}'.format(base_mse))
print('Model MSE = {:.2f}'.format(mse))
# -
# # Feature importance (SAGE)
# Setup and calculate
imputer = sage.MarginalImputer(model, test[:512])
estimator = sage.PermutationEstimator(imputer, 'mse')
sage_values = estimator(test, Y_test)
# Plot results
sage_values.plot(feature_names)
# # Model sensitivity (Shapley Effects)
# Setup and calculate
imputer = sage.MarginalImputer(model, test[:512])
estimator = sage.PermutationEstimator(imputer, 'mse')
sensitivity = estimator(test)
# Plot results
sensitivity.plot(feature_names, title='Model Sensitivity')
|
notebooks/bike.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sbi_prinz
# language: python
# name: sbi_prinz
# ---
# ## Running simulations with `pyloric`
# +
# # %load ../pyloric/utils/pandas_setup.py
import pandas as pd
pd.set_option("display.max_rows", 10)
pd.set_option("display.max_columns", 100)
pd.set_option("display.width", 2000)
pd.set_option("display.float_format", "{:,.4f}".format)
# -
from pyloric import create_prior, simulate, stats
from pyloric.utils import show_traces
import torch
# ### Obtain parameter sets
_ = torch.manual_seed(0)
prior = create_prior()
p = prior.sample((2,))
p
# ### Simulate them and plot the traces
sim_outputs = [simulate(param_set, seed=0) for param_set in p.to_numpy()]
for sim_ in sim_outputs:
_ = show_traces(sim_)
# ### Compute summary statistics
summstats = pd.concat([stats(sim_) for sim_ in sim_outputs], ignore_index=True)
summstats
|
tutorials/01_run_pyloric.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding=utf-8
# Copyright 2019 The Google Research Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.optim import lr_scheduler
from torchvision import datasets, transforms, utils
import numpy as np
import pdb
import argparse
import time
import os
import sys
import foolbox
import wideresnet
from collections import OrderedDict
from utils import *
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
def remove_module_state_dict(state_dict):
new_state_dict = OrderedDict()
for key in state_dict.keys():
new_key = '.'.join(key.split('.')[1:])
new_state_dict[new_key] = state_dict[key]
return new_state_dict
# +
# #!pip3 install foolbox
# -
# Setup parameters
class eval_args():
def __init__(self, param_dict):
# training
self.dataset = 'cifar' #parser.add_argument('--dataset', type=str, default='cifar')
self.batch_size = 50 #parser.add_argument('--batch_size', type=int, default=50)
self.norm = None #parser.add_argument("--norm", type=str, default=None, choices=[None, "norm", "batch", "instance", "layer", "act"])
# EBM specific
self.n_steps = 100 #parser.add_argument("--n_steps", type=int, default=100)
self.width = 10 #parser.add_argument("--width", type=int, default=10)
self.depth = 28 #parser.add_argument("--depth", type=int, default=28)
#
self.n_steps_refine = 0 #parser.add_argument('--n_steps_refine', type=int, default=0)
self.n_classes = 10 #parser.add_argument('--n_classes',type=int,default=10)
self.init_batch_size = 128 #parser.add_argument('--init_batch_size', type=int, default=128)
self.softmax_ce = False #parser.add_argument('--softmax_ce', action='store_true')
# attack
self.attack_conf = False #parser.add_argument('--attack_conf', action='store_true')
self.random_init = False #parser.add_argument('--random_init', action='store_true')
self.threshold = .7 #parser.add_argument('--threshold', type=float, default=.7)
self.debug = False #parser.add_argument('--debug', action='store_true')
self.no_random_start = False #parser.add_argument('--no_random_start', action='store_true')
self.load_path = None #parser.add_argument("--load_path", type=str, default=None)
self.distance = 'Linf' #parser.add_argument("--distance", type=str, default='Linf')
self.n_steps_pgd_attack = 40 #parser.add_argument("--n_steps_pgd_attack", type=int, default=40)
self.start_batch = -1 #parser.add_argument("--start_batch", type=int, default=-1)
self.end_batch = 2 #parser.add_argument("--end_batch", type=int, default=2)
self.sgld_sigma = 1e-2 #parser.add_argument("--sgld_sigma", type=float, default=1e-2)
self.n_dup_chains = 5 #parser.add_argument("--n_dup_chains", type=int, default=5)
self.sigma = .03 #parser.add_argument("--sigma", type=float, default=.03)
self.base_dir = './experiment_attacks' #parser.add_argument("--base_dir", type=str, default='./adv_results')
# added
self.exp_name = 'exp' #parser.add_argument('--exp_name', type=str, default='exp', help='saves everything in ?r/exp_name/')
# set from inline dict
for key in param_dict:
#print(key, '->', param_dict[key])
setattr(self, key, param_dict[key])
# +
# instantiate
# --start_batch 0 --end_batch 6 --load_path /cloud_storage/BEST_EBM.pt
#--exp_name rerun_ebm_1_step_5_dup_l2_no_sigma_REDO --n_steps_refine 1 --distance L2
#--random_init --n_dup_chains 5 --sigma 0.0 --base_dir /cloud_storage/adv_results &
args = eval_args({"load_path": "./experiment_weights/energy_with_aug_L2_dlogpx_dx_3/last_ckpt.pt", "distance": 'L2', \
"start_batch": 0, "end_batch": 1, "n_dup_chains": 5, "sigma": 0.0, "random_init": True, \
"batch_size": 50})
device = torch.device('cuda')
# locations
base_dir = args.base_dir
save_dir = os.path.join(base_dir, args.exp_name, 'saved_model')
last_dir = os.path.join(save_dir,'last')
best_dir = os.path.join(save_dir,'best')
data_dir = os.path.join(base_dir,'data')
# -
class gradient_attack_wrapper(nn.Module):
def __init__(self, model):
super(gradient_attack_wrapper, self).__init__()
self.model = model.eval()
def forward(self, x):
x = x - 0.5
x = x / 0.5
x.requires_grad_()
out = self.model.module.refined_logits(x)
return out
def eval(self):
return self.model.eval()
class F(nn.Module):
def __init__(self, depth=28, width=2, norm=None):
super(F, self).__init__()
self.f = wideresnet.Wide_ResNet(depth, width, norm=norm)
self.energy_output = nn.Linear(self.f.last_dim, 1)
self.class_output = nn.Linear(self.f.last_dim, 10)
def forward(self, x, y=None):
penult_z = self.f(x)
return self.energy_output(penult_z).squeeze()
def classify(self, x):
penult_z = self.f(x)
return self.class_output(penult_z)
class CCF(F):
def __init__(self, depth=28, width=2, norm=None):
super(CCF, self).__init__(depth, width, norm=norm)
def forward(self, x, y=None):
logits = self.classify(x)
if y is None:
return logits.logsumexp(1)
else:
return torch.gather(logits, 1, y[:, None])
# wrapper class to provide utilities for what you need
class DummyModel(nn.Module):
def __init__(self, f):
super(DummyModel, self).__init__()
self.f = f
def logits(self, x):
return self.f.classify(x)
def refined_logits(self, x, n_steps=args.n_steps_refine):
xs = x.size()
dup_x = x.view(xs[0], 1, xs[1], xs[2], xs[3]).repeat(1, args.n_dup_chains, 1, 1, 1)
dup_x = dup_x.view(xs[0] * args.n_dup_chains, xs[1], xs[2], xs[3])
dup_x = dup_x + torch.randn_like(dup_x) * args.sigma
refined = self.refine(dup_x, n_steps=n_steps, detach=False)
logits = self.logits(refined)
logits = logits.view(x.size(0), args.n_dup_chains, logits.size(1))
logits = logits.mean(1)
return logits
def classify(self, x):
logits = self.logits(x)
pred = logits.max(1)[1]
return pred
def logpx_score(self, x):
# unnormalized logprob, unconditional on class
return self.f(x)
def refine(self, x, n_steps=args.n_steps_refine, detach=True):
# runs a markov chain seeded at x, use n_steps=10
x_k = torch.autograd.Variable(x, requires_grad=True) if detach else x
# sgld
for k in range(n_steps):
f_prime = torch.autograd.grad(self.f(x_k).sum(), [x_k], retain_graph=True)[0]
x_k.data += f_prime + args.sgld_sigma * torch.randn_like(x_k)
final_samples = x_k.detach() if detach else x_k
return final_samples
def grad_norm(self, x):
x_k = torch.autograd.Variable(x, requires_grad=True)
f_prime = torch.autograd.grad(self.f(x_k).sum(), [x_k], retain_graph=True)[0]
grad = f_prime.view(x.size(0), -1)
return grad.norm(p=2, dim=1)
def logpx_delta_score(self, x, n_steps=args.n_steps_refine):
# difference in logprobs from input x and samples from a markov chain seeded at x
#
init_scores = self.f(x)
x_r = self.refine(x, n_steps=n_steps)
final_scores = self.f(x_r)
# for real data final_score is only slightly higher than init_score
return init_scores - final_scores
def logp_grad_score(self, x):
return -self.grad_norm(x)
# +
model_attack_wrapper =gradient_attack_wrapper
transformer_train = transforms.Compose([transforms.ToTensor()])
transformer_test = transforms.Compose([transforms.ToTensor()])
data_loader = torch.utils.data.DataLoader(datasets.CIFAR10(data_dir, train=False,
transform=transformer_test, download=True),
batch_size=args.batch_size, shuffle=False, num_workers=10)
init_loader = torch.utils.data.DataLoader(datasets.CIFAR10(data_dir, train=True,
download=True, transform=transformer_train),
batch_size=args.init_batch_size, shuffle=True, num_workers=1)
# -
# construct model and ship to GPU
f = CCF(args.depth, args.width, args.norm)
print(args.load_path)
print("loading model from {args.load_path}")
ckpt_dict = torch.load(args.load_path)
if "model_state_dict" in ckpt_dict:
# loading from a new checkpoint
f.load_state_dict(ckpt_dict["model_state_dict"])
else:
# loading from an old checkpoint
f.load_state_dict(ckpt_dict)
# +
f = DummyModel(f)
model = f.to(device)
model = nn.DataParallel(model).to(device)
model.eval()
## Define criterion
criterion = foolbox.criteria.Misclassification()
# +
## Initiate attack and wrap model
model_wrapped = model_attack_wrapper(model)
fmodel = foolbox.models.PyTorchModel(model_wrapped, bounds=(0.,1.), num_classes=10, device=device)
if args.distance == 'L2':
distance = foolbox.distances.MeanSquaredDistance
attack = foolbox.attacks.L2BasicIterativeAttack(model=fmodel, criterion=criterion, distance=distance)
else:
distance = foolbox.distances.Linfinity
attack = foolbox.attacks.RandomStartProjectedGradientDescentAttack(model=fmodel, criterion=criterion, distance=distance)
print('Starting...')
# Grab 50 images at a time (k index), go though batches start_batch to end_batch (i index)
for i, (img, label) in enumerate(data_loader):
print("i... ", i)
adversaries = []
if i < args.start_batch:
continue
if i >= args.end_batch:
break
# get images and their logits from classification with the model
img = img.data.cpu().numpy()
logits = model_wrapped(torch.from_numpy(img[:, :, :, :]).to(device))
# Get the k largest elements of the logits along dimention 1
_, top = torch.topk(logits,k=2,dim=1)
top = top.data.cpu().numpy()
# prediction is the first element
pred = top[:,0]
# loop through the 50 images
#print("pred",pred,len(label))
for k in range(len(label)):
print("k... ", k)
# get image and its label
im = img[k,:,:,:]
orig_label = label[k].data.cpu().numpy()
# check its croorectly classified else we cant use for mis-classification
if pred[k] != orig_label:
print("Error in prediction")
continue
# setup for adversarial with lowest distance
best_adv = None
# try 20 times for best
for ii in range(1): #20):
try:
# create adversarial
adversarial = attack(im, label=orig_label, unpack=False, random_start=True, iterations=args.n_steps_pgd_attack)
print("adv", adversarial.distance)
# save if its the lowest distance
# I guess it iterates to failure and we want the smallest of these
if ii == 0 or best_adv.distance > adversarial.distance:
best_adv = adversarial
except Exception as e:
print("Failed Adv.", e)
continue
try:
# save our adversarial
adversaries.append((im, orig_label, adversarial.image, adversarial.adversarial_class))
except:
continue
# save our adversarials
adv_save_dir = os.path.join(base_dir, args.exp_name)
save_file = 'adversarials_batch_'+str(i)
if not os.path.exists(os.path.join(adv_save_dir,save_file)):
os.makedirs(os.path.join(adv_save_dir,save_file))
np.save(os.path.join(adv_save_dir,save_file),adversaries)
# +
# save and load our 7 images
#np.save('./experiment_attacks/exp/adversarials_batch_test2.npy', adversaries)
#adversaries_rl = np.load("./experiment_attacks/exp/adversarials_batch_test2.npy", allow_pickle=True)
# -
# plot data
def find_epsilons(adversaries_rl):
adv_size = adversaries_rl.shape[0]
eps = np.zeros(adv_size + 1)
acc = np.zeros(adv_size + 1)
eps[0] = 0.
acc[0] = 1.
for i in range(adv_size):
perturbation = adversaries_rl[i,2] - adversaries_rl[i,0]
#print('L2 norm of perturbation: {}'.format(np.linalg.norm(perturbation.flatten()*255, 2)))
eps[i+1] = np.linalg.norm(perturbation.flatten()*255, 2)
acc[i+1] = 1. - (i+1.)/adv_size
eps = np.sort(eps)
#print(eps[0],acc[0],eps[6],acc[6],eps[7],acc[7])
return eps, acc
#print("image label", adversaries_rl[i,1], "peturb label", adversaries_rl[i,3])
# +
# plot results
adversaries_rl_ewa = np.load("./experiment_attacks/exp/adversarials_batch_0_full_with-aug_max-ent.npy", allow_pickle=True)
eps_ewa, acc_ewa = find_epsilons(adversaries_rl_ewa)
#eps_ewa
adversaries_rl_2 = np.load("./experiment_attacks/exp/adversarials_batch_0_energy_with_aug.npy", allow_pickle=True) #_energy_with_aug
eps_2, acc_2 = find_epsilons(adversaries_rl_2)
adversaries_rl_2a = np.load("./experiment_attacks/exp/adversarials_batch_0.npy", allow_pickle=True) #_energy_with_aug
eps_2a, acc_2a = find_epsilons(adversaries_rl_2a)
adversaries_rl_b = np.load("./experiment_attacks/exp/adversarials_batch_0_with_aug.npy", allow_pickle=True)
eps_b, acc_b = find_epsilons(adversaries_rl_b)
import matplotlib.pyplot as plt
plt.plot(eps_b, acc_b * 100, label="Baseline")
plt.plot(eps_ewa, acc_ewa * 100, label="JEM")
plt.plot(eps_2, acc_2 * 100, label="JEM-CRS")
plt.plot(eps_2a, acc_2a * 100, label="JEM-CRS-abs-dpx")
plt.legend(loc="upper right")
plt.xlim([0,300])
plt.xlabel("epsilon")
plt.ylabel("accuracy")
plt.title("Adversarial attack")
plt.savefig('experiment_output/images/adversarials.png')
plt.show()
# -
# %matplotlib inline
from matplotlib import pyplot as plt
img = adversaries_rl[1,0]
img = img.swapaxes(0,1)
img = img.swapaxes(1,2)
plt.imshow(img, interpolation='nearest')
plt.show()
# %matplotlib inline
from matplotlib import pyplot as plt
img = adversaries_rl[1,2]
img = img.swapaxes(0,1)
img = img.swapaxes(1,2)
plt.imshow(img, interpolation='nearest')
plt.show()
# %matplotlib inline
from matplotlib import pyplot as plt
img = adversaries_rl[1,2] - adversaries_rl[1,0]
img = img.swapaxes(0,1)
img = img.swapaxes(1,2)
plt.imshow(img, interpolation='nearest')
plt.show()
adversaries_rl[1,2] - adversaries_rl[1,0]
import matplotlib.pyplot as plt
plt.plot(eps_ewa[:40], acc_ewa[:40])
plt.show()
|
attack_wrn_ebm_nb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %config InlineBackend.figure_format = True
import matplotlib.pyplot as plt
import numpy as np
from astropy.time import Time
import astropy.units as u
from rms import STSP, LightCurve, Star, Spot
# +
spot_contrast = 0.7 # 0 -> perfectly dark; 1 -> same as photosphere
rotation_period = 30 # days
inc_stellar = 90 # Tilt of stellar rotation axis away from observer
star = Star(rotation_period, inc_stellar, spot_contrast)
start_time = Time('2009-01-01 00:00')
end_time = start_time + rotation_period * u.day
duration = end_time - start_time
time_steps = 30*u.min
times = start_time + np.arange(0, duration.to(u.day).value,
time_steps.to(u.day).value) * u.day
# -
n_spots = 30
random_radii = 0.1 * np.random.rand(n_spots)**3
spots = [Spot.at_random_position(radius=r) for r in random_radii]
with STSP(times, star, spots) as stsp:
lc = stsp.generate_lightcurve(n_ld_rings=5)
# +
fig = plt.figure(figsize=(12, 4))
ax_star_map = fig.add_subplot(121, projection='hammer')
ax_lightcurve = fig.add_subplot(122)
for spot in spots:
spot.plot(ax=ax_star_map)
ax_star_map.grid()
lc.plot(star, ax=ax_lightcurve)
# -
inverse_transform
# +
def sunspot_distribution(latitude):
return np.exp(-0.5 * (abs(latitude) - 15)**2 / 6**2)
def sunspot_inverse_transform(x):
lats = np.linspace(-60, 60, 1000)
prob = np.cumsum(sunspot_distribution(lats))
prob /= np.max(prob)
return np.interp(x, prob, lats)
def draw_sunspots(n):
return sunspot_inverse_transform(np.random.rand(n))
plt.hist(inverse_transform(np.random.rand(1000)))
# +
lat =
lon = 2*np.pi * np.random.rand(len(lat))
plt.hist()
|
notebooks/random_draws.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
import copy
from collections import OrderedDict
import argparse
import json
import pickle
from datetime import datetime
from sklearn.metrics import classification_report, roc_curve, precision_recall_curve,roc_auc_score, f1_score, confusion_matrix
from sklearn.model_selection import GridSearchCV
from sklearn.neighbors import KNeighborsRegressor
import xgboost as xgb
import shap
from imblearn.over_sampling import SMOTE
import utils
# %matplotlib inline
# -
# Recall from previous notebook the top feature importances:
#
# 
#
#
# We now wish to take what we learned to create a smaller version of the model that can take in user input and convert it to useful features.
# # 1.0 Load Data
#get directory
df_train_path = os.path.join('data','df_train_scaled.csv')
df_test_path = os.path.join('data','df_test_scaled.csv')
df_train = pd.read_csv(df_train_path, compression='zip',index_col=0)
df_train.head()
df_train.shape
df_test = pd.read_csv(df_test_path, compression='zip', index_col=0)
df_test.head()
df_test.shape
# # 2.0 FICO to Grade: An Example
df_fico_grade = pd.read_csv('data/grade_to_fico.csv')
df_fico_grade.head()
plt.figure(figsize=(10,5))
plt.plot(df_fico_grade['value'], df_fico_grade['score'], marker='x')
plt.xticks(range(df_fico_grade.shape[0]) , df_fico_grade['sub_grade'][::-1], rotation='vertical')
plt.xlabel('Sub Grade')
plt.ylabel("FICO Score")
plt.title("FICO Versus Sub Grade Per SEC Filing")
plt.savefig(os.path.join('plots','fico_grade_line.png'))
plt.show()
#instantiate regressor
knn = KNeighborsRegressor(n_neighbors=1)
#fit to data
knn.fit(np.reshape(df_fico_grade['score'].values, (-1, 1)),
df_fico_grade['value'])
#make sample prediction
knn.predict(np.reshape([750], (1,-1)))[0]
# ### Save KNN Regression
#save the model
with open(os.path.join('models', 'knn_regression.pkl'), 'wb') as handle:
pickle.dump(knn,
handle)
# # 3.0 FICO to Interest Rate
df_fico_apr = pd.read_csv('data/grade_to_apr.csv')
df_fico_apr.head()
df_fico_apr[df_fico_apr['grade_num']==30]['36_mo']
# # 4.0 Fit Model
# ## 4.1 Prepare Data
# +
#target variable
target_col = 'loan_status'
#training variables
X_train = df_train.drop(columns=[target_col])
y_train = df_train[target_col]
#instantiate smote
sm = SMOTE(random_state=42)
#apply oversampling to training data
X_train, y_train = sm.fit_resample(X_train, y_train)
#test variables
X_test = df_test.drop(columns=[target_col])
y_test = df_test[target_col]
# -
# ## 4.2 Training
#instantiate model
clf = xgb.XGBClassifier(colsample_bytree=0.9, eta=0.3, max_depth=3)
#fit to data
clf.fit(X_train, y_train)
# ## 4.3 Predictions
#make predictions
y_pred = clf.predict(X_test.values)
confusion_matrix(y_test, y_pred)
fpr, tpr, thresholds = roc_curve(y_test,
clf.predict_proba(X_test.values)[:,1],
pos_label=1)
precision, recall, thresholds = precision_recall_curve(y_test,
clf.predict_proba(X_test.values)[:,1],
pos_label=1)
plt.plot(fpr, tpr, label='XGB')
plt.plot([0,1], [0,1], label='No Discrimination', linestyle='-', dashes=(5, 5))
plt.show()
xgb_auc = roc_auc_score(y_test, clf.predict_proba(X_test.values)[:,1])
print("xgb_auc: {}".format(xgb_auc))
# +
f1_xgb = f1_score(y_test, y_pred)
print("f1_xgb (binary): {}\n".format(f1_xgb))
print(classification_report(y_test, y_pred))
# -
# ## 4.4 Save Model
# +
#define location to save trained model
save_model_dir = os.path.join('models','xgb_cv_compact.pkl')
print("Saving model at: {}".format(save_model_dir))
#save the model
with open(save_model_dir, 'wb') as handle:
pickle.dump(clf,
handle)
# -
# # 5.0 Case by Case Timing
# +
predictions_list = []
for x, y in zip(X_test.values, y_test):
start = datetime.now()
pred = clf.predict(x.reshape(1, -1))[0]
delta = (datetime.now() - start).microseconds
pred_dict = {'y_true': y, 'y_pred': pred, 'pred_time': delta}
predictions_list.append(pred_dict)
pred_df = pd.DataFrame(predictions_list)
pred_df.head()
# -
pred_df.describe()
plt.hist(pred_df['pred_time'], bins=np.arange(100, 255, 5))
plt.xlabel("Prediction Time (microseconds)")
plt.show()
# ## 6.0 Custom Input
df_train.columns
# Ask user for:
# - FICO score
# - loan_amount
# - term
# - dti
# - home_ownership
# - mort_acc: number of mortage accounts
# - annual_inc
# - open_acc: number of open credit lines
# - employment
# - verification_status: loan verified or not
# - revol_bal
# - revol_util: Revolving linΩ utilization rate, or the amount of credit the borrower is using relative to all available revolving credit
# - pub_rec
# - emp_length
# - purpose
#
#
# calculate:
# - sub_grade
# - grade
# - installment
# - int_rate
#
# +
def emp_title_to_dict(e_title):
#force make string if not and make lower
title_lower = str(e_title).lower()
#list of employment types to consider
emp_list = ['e_manager', 'e_educ', 'e_self',
'e_health', 'e_exec', 'e_driver',
'e_law', 'e_admin', 'e_fin', 'e_other']
#instantiate title dict
title_dict = dict(zip(emp_list, len(emp_list)*[0]))
#check and fill out dict
if any(job in title_lower for job in ['manag', 'superv']):
title_dict['e_manager'] = 1
elif 'teacher' in title_lower:
title_dict['e_educ'] = 1
elif 'owner' in title_lower:
title_dict['e_self'] = 1
elif any(job in title_lower for job in ['rn', 'registered nurse', 'nurse',
'doctor', 'pharm', 'medic']):
title_dict['e_health'] = 1
elif any(job in title_lower for job in ['vice president', 'president', 'director',
'exec', 'chief']):
title_dict['e_exec'] = 1
elif any(job in title_lower for job in ['driver', 'trucker']):
title_dict['e_driver'] = 1
elif any(job in title_lower for job in ['lawyer', 'legal', 'judg']):
title_dict['e_law'] = 1
elif 'admin' in title_lower:
title_dict['e_admin'] = 1
elif any(job in title_lower for job in ['analyst', 'financ', 'sales']):
title_dict['e_fin'] = 1
else:
title_dict['e_other'] = 1
return title_dict
def purp_to_int(title):
#force make string if not and make lower
title_lower = str(title).lower()
#list of employment types to consider
title_list = ['purp_car', 'purp_credit_card', 'purp_debt_consolidation',
'purp_educational', 'purp_home_improvement',
'purp_house', 'purp_major_purchase', 'purp_medical',
'purp_moving', 'purp_other', 'purp_renewable_energy',
'purp_small_business', 'purp_vacation', 'purp_wedding']
#instantiate title dict
title_dict = dict(zip(title_list,
len(title_list)*[0]))
#check if any shared items
for key in title_dict:
if bool(set(title_lower.split()) & set(key.split('_'))):
title_dict[key] = 1
return title_dict
home_to_int = {'MORTGAGE': 4,
'RENT': 3,
'OWN': 5,
'ANY': 2,
'OTHER': 1,
'NONE':0 }
# +
#get input
fico = 750
loan_amnt = 6000
term = 36
dti = 20
home_ownership = 'rent'
mort_acc = 5
annual_inc = 50_000
open_acc = 5
employment = 'nurse'
verification_status = 1
revol_bal = 14_000
revol_util = 60
pub_rec = 0
emp_length = 10
application_type = 0
emp_title = 'sales'
purpose = 'wedding'
pub_rec_bankruptcies = 0
#calculate grade from FICO
sub_grade = knn.predict(np.reshape([fico], (1,-1)))[0]
#calculate grade
grade = round(sub_grade/5) + 1
#get purpose of loan
title_dict = emp_title_to_dict(emp_title)
#get purpose dict
purp_dict = purp_to_int(purpose)
#get interest rate
apr_row = df_fico_apr[df_fico_apr['grade_num']==sub_grade]
if term<=36:
int_rate = apr_row['36_mo'].values[0]
installment = float(loan_amnt)/36
else:
int_rate = apr_row['60'].values[0]
installment = float(loan_amnt)/60
# -
apr_row['36_mo'].values[0]
# +
title_df = pd.DataFrame({k: [v] for k, v in title_dict.items()})
purp_df = pd.DataFrame({k: [v] for k, v in purp_dict.items()})
temp =pd.concat([title_df, purp_df], axis=1)
#temp['fico'] = fico
temp['loan_amnt'] = loan_amnt
temp['term'] = term
temp['dti'] = dti
temp['home_ownership'] = home_to_int[home_ownership.upper()]
temp['mort_acc'] = mort_acc
temp['annual_inc'] = annual_inc
temp['open_acc'] = open_acc
temp['verification_status'] = verification_status
temp['revol_bal'] = revol_bal
temp['revol_util'] = revol_util
temp['pub_rec'] = pub_rec
temp['emp_length'] = emp_length
temp['application_type'] = application_type
temp['int_rate'] = int_rate
temp['installment'] = installment
temp['grade'] = grade
temp['sub_grade'] = sub_grade
temp['time_delta'] = 20
temp['pub_rec_bankruptcies'] = pub_rec_bankruptcies
temp['total_acc'] = 1
temp = temp[df_train.drop(columns=['loan_status']).columns]
temp.shape
# +
temp = temp[['pub_rec', 'grade', 'purp_renewable_energy', 'revol_bal', 'open_acc',
'mort_acc', 'purp_credit_card', 'purp_car', 'e_exec',
'verification_status', 'purp_other', 'e_other', 'home_ownership',
'e_educ', 'emp_length', 'e_driver', 'revol_util',
'purp_moving', 'loan_amnt', 'time_delta', 'e_law', 'e_health', 'e_fin',
'purp_small_business', 'sub_grade', 'application_type', 'dti',
'e_manager', 'purp_major_purchase', 'pub_rec_bankruptcies',
'purp_house', 'term', 'installment', 'total_acc', 'e_admin',
'purp_medical', 'purp_vacation', 'e_self', 'purp_debt_consolidation',
'int_rate', 'annual_inc', 'purp_home_improvement']]
temp.head()
# -
X_train.shape
# +
df_macro_mean = pd.read_csv('data/df_macro_mean.csv', index_col=0)
df_macro_std = pd.read_csv('data/df_macro_std.csv', index_col=0)
# -
df_macro_mean.head()
# +
scale = temp.copy()
code=23
for feat in df_macro_mean.columns:
scale[feat] = (scale[feat] - df_macro_mean.loc[code,feat]) / df_macro_std.loc[code,feat]
scale.head()
# -
clf.predict(scale.values)[0]
temp.to_dict(orient='record')[0]
clf.save_model('models/clf.pkl')
import sklearn
sklearn.__version__
|
notebooks/05_CompactModel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/zerotodeeplearning/ztdl-masterclasses/blob/master/solutions_do_not_open/Data_Augmentation_solution.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="2bwH96hViwS7"
# #### Copyright 2020 Catalit LLC.
# + colab={} colab_type="code" id="bFidPKNdkVPg"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="DvoukA2tkGV4"
# # Data Augmentation
# + colab={} colab_type="code" id="d85dZiUHNQsQ"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
import os
from tensorflow.keras.preprocessing import image
# + colab={} colab_type="code" id="W9rpfDisNSs9"
# sports_images_path = tf.keras.utils.get_file(
# 'sports_images',
# 'https://archive.org/download/ztdl_sports_images/sports_images.tgz',
# untar=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="WeovACo4OyAM" outputId="97bf6652-1b99-4108-ef1e-a0b8f249c691"
![[ ! -f sports_images.tar.gz ]] && gsutil cp gs://ztdl-datasets/sports_images.tar.gz .
![[ ! -d sports_images ]] && echo "Extracting images..." && tar zxf sports_images.tar.gz
sports_images_path = './sports_images'
# + colab={} colab_type="code" id="FOjyb1QXV7Vz"
train_path = os.path.join(sports_images_path, 'train')
test_path = os.path.join(sports_images_path, 'test')
# + colab={} colab_type="code" id="Z_pFvoKoWCG9"
batch_size = 16
img_size = 224
# + colab={} colab_type="code" id="Rdrw2B16Nbvk"
datagen = image.ImageDataGenerator(
rescale=1./255.,
rotation_range=15,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=5,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
# + colab={"base_uri": "https://localhost:8080/", "height": 241} colab_type="code" id="EynSA1dQYXmp" outputId="138ccfdf-22ce-43cf-f8f2-ab4ca1786961"
input_path = os.path.join(train_path, 'Beach volleyball/1e9ce0e76695de2e5d1f6964ab8c538.jpg')
img = image.load_img(input_path, target_size=(img_size, img_size))
img
# + colab={} colab_type="code" id="L3L_2T8u8U-S"
img_array = image.img_to_array(img)
# + colab={} colab_type="code" id="qVX0AA7J8U6r"
img_tensor = np.expand_dims(img_array, axis=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 717} colab_type="code" id="ohkU-lku8U2R" outputId="c837cc3a-f688-4eab-94be-e487bcdeec37"
plt.figure(figsize=(10, 10))
i = 0
for im in datagen.flow(img_tensor, batch_size=1):
i += 1
if i > 16:
break
plt.subplot(4, 4, i)
plt.imshow(im[0])
plt.axis('off')
plt.tight_layout()
# + [markdown] colab_type="text" id="ppplZ34g8UyI"
# ### Exercise 1
#
# - Use the `flow_from_directory` method of the data generator to produce a batch of images of sports flowing from the training directory `train_path`.
# - display the images with their labels
#
# Your code should look like:
#
# ```python
# train_datagen = datagen.flow_from_directory(
# # YOUR CODE HERE
# )
#
# batch, labels = train_datagen.next()
#
# plt.figure(figsize=(10, 10))
# for i in range(len(batch)):
# # YOUR CODE HERE
# ```
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="U7UUzGIW8Uuw" outputId="aaee65ee-8ee9-4c17-c0ac-75902dab5ba8" tags=["solution", "empty"]
train_datagen = datagen.flow_from_directory(
train_path,
target_size=(img_size, img_size),
batch_size=batch_size,
class_mode = 'sparse',
shuffle=True)
classes_dict = train_datagen.class_indices
classes = list(classes_dict.keys())
# + colab={} colab_type="code" id="gCPiSFzK8UrZ" tags=["solution"]
batch, labels = train_datagen.next()
# + colab={"base_uri": "https://localhost:8080/", "height": 729} colab_type="code" id="Sy7HcmUa8UoL" outputId="517f2e27-8d38-42da-cfb7-aa0d23542ad0" tags=["solution"]
plt.figure(figsize=(10, 10))
for i in range(len(batch)):
plt.subplot(4, 4, i+1)
plt.imshow(batch[i])
plt.title(classes[int(labels[i])])
plt.axis('off')
plt.tight_layout()
# + [markdown] colab_type="text" id="wCq_t4tVC0rE"
# ### Tensorflow Data Generators
# + colab={} colab_type="code" id="6K5TN9MV-_G8"
from imutils import paths
# + colab={} colab_type="code" id="tmUpG2Nziik7"
def parse_images(im_path):
im = tf.io.read_file(im_path)
im = tf.image.decode_jpeg(im, channels=3)
im = tf.image.convert_image_dtype(im, tf.float32)
im = tf.image.resize(im, [img_size, img_size])
label = tf.strings.split(im_path, os.path.sep)[-2]
return (im, label)
# + colab={} colab_type="code" id="lvu-1nO8ny3k"
im_paths = list(paths.list_images(train_path))
path_ds = tf.data.Dataset.from_tensor_slices((im_paths))
# + colab={} colab_type="code" id="Idc45mpUoDJW"
AUTO = tf.data.experimental.AUTOTUNE
# + colab={} colab_type="code" id="2xC7x6F0oD9e"
train_ds = (
path_ds
.map(parse_images, num_parallel_calls=AUTO)
.shuffle(10000)
.batch(batch_size)
.prefetch(AUTO)
)
# + colab={} colab_type="code" id="IWITasIDoqiV"
batch, labels = next(iter(train_ds))
# + colab={"base_uri": "https://localhost:8080/", "height": 729} colab_type="code" id="WilMQSo5pruo" outputId="5cb0fd38-9dd7-4c8f-f195-009964c07e34"
plt.figure(figsize=(10, 10))
for i in range(len(batch)):
plt.subplot(4, 4, i+1)
plt.imshow(batch[i])
plt.title(labels[i].numpy().decode('utf-8'))
plt.axis('off')
plt.tight_layout()
# + [markdown] colab_type="text" id="JqEEhAzWC4dX"
# ### Exercise 2: Data augmentation with Keras layers
#
# Keras provides a few experimental layers to include data augmentation in the model.
#
# - Define a data augmentation model using a `Sequential` model with a few layers from the `tensorflow.keras.layers.experimental.preprocessing` submodule.
# - Apply this model on the batch using the flag `training=True` to ensure data augmentation is applied
# - Visualize the augmented images as above
# - What are the advantages of including data augmentation in the model?
# + colab={} colab_type="code" id="KmU4ki5CASIF" tags=["solution", "empty"]
from tensorflow.keras.layers.experimental.preprocessing import RandomFlip, RandomRotation, RandomTranslation
# + colab={} colab_type="code" id="rgk3wkHFoSvO" tags=["solution"]
data_aug_layer = tf.keras.Sequential([
RandomFlip('horizontal'),
RandomRotation(0.3),
RandomTranslation(height_factor=0.2, width_factor=0.2)
])
# + colab={} colab_type="code" id="WtlFwN0XoXC0" tags=["solution"]
aug_batch = data_aug_layer(batch, training=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 729} colab_type="code" id="M01r4q3aAu5U" outputId="da8a71c7-78da-4035-f6e4-31b533c76372" tags=["solution"]
plt.figure(figsize=(10, 10))
for i in range(len(batch)):
plt.subplot(4, 4, i+1)
plt.imshow(aug_batch[i])
plt.title(labels[i].numpy().decode('utf-8'))
plt.axis('off')
plt.tight_layout()
|
solutions_do_not_open/Data_Augmentation_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="TymKjsOlbojL" outputId="a176f22c-b064-4607-ed31-2d1088cbb2d3"
# ! pip install kaggle
# + id="EYtCb-jsZZ2k"
# ! mkdir ~/.kaggle
# + id="aclV4CpBZl82"
# ! cp kaggle.json ~/.kaggle/
# ! chmod 600 ~/.kaggle/kaggle.json
# + colab={"base_uri": "https://localhost:8080/"} id="2TZYv6L9bBBY" outputId="9d816413-60eb-42f1-915a-77f7cebaa2d4"
# ! kaggle competitions download -c fake-news
# + colab={"base_uri": "https://localhost:8080/"} id="GGvdBBBjbL2h" outputId="b2814138-03fb-4efe-b0bb-c402dfe76a34"
# ! unzip train.csv.zip
# + colab={"base_uri": "https://localhost:8080/"} id="-jCKM8MTclaQ" outputId="a6187283-0d5b-46cb-c8e2-e22fe24d6c78"
# ! unzip test.csv.zip
# + colab={"base_uri": "https://localhost:8080/"} id="Jx9Mtj2QVsJa" outputId="11114e1a-3b1e-437f-9469-b243e25a5add"
#dependencies
import warnings
warnings.filterwarnings("ignore")
# !pip install contractions
# !pip install kaggle_datasets
import nltk
nltk.download('stopwords')
nltk.download('punkt')
nltk.download('words')
nltk.download('wordnet')
import re
import pickle
import numpy as np
import pandas as pd
import contractions
import seaborn as sns
import matplotlib.pyplot as plt
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Embedding, LSTM, Conv1D, MaxPooling1D, Dropout, BatchNormalization
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, accuracy_score
import gensim
from sklearn.metrics import confusion_matrix
import tensorflow as tf
from tensorflow.keras.models import Model, load_model
#from tensorflow.keras.callbacks import ReduceLROnPlateau, LearningRateScheduler, EarlyStopping, ModelCheckpoint
#from kaggle_datasets import KaggleDatasets
#import transformers
#from transformers import TFAutoModel, AutoTokenizer
#from tqdm.notebook import tqdm
#from tokenizers import Tokenizer
from tensorflow.keras.layers import Input, Dense
from tensorflow.keras.optimizers import Adam
import tensorflow.keras
# + id="Sj9A3IY5VsJe"
df_2 = pd.read_csv("/content/train.csv", header=0, index_col=0)
df_t = pd.read_csv("/content/test.csv", header=0, index_col=0)
df_2 = df_2.drop(['title','author'], axis = 1)
df_t = df_t.drop(['title','author'], axis = 1)
df_2.dropna(inplace = True)
df_t.fillna('',inplace = True)
#print(df_2.isnull().sum(axis = 0))
# + id="6K4MfGXwVsJg"
def clean_text(text_col ):
text_col = text_col.apply(lambda x: [contractions.fix(word, slang=False).lower() for word in x.split()])
text_col = text_col.apply(lambda x: [re.sub(r'[^\w\s]','', word) for word in x])
stop_words = set(stopwords.words('english'))
text_col = text_col.apply(lambda x: [word for word in x if word not in stop_words])
text_col = text_col.apply(lambda x: [word for word in x if re.search("[@_!#$%^&*()<>?/|}{~:0-9]", word) == None])
return text_col
df_2["text"] = clean_text(df_2["text"])
df_t["text"] = clean_text(df_t["text"])
df_2['label'] = df_2['label'].apply(lambda x: int(x))
y = df_2['label']
# + id="rYcLecLfVsJi" colab={"base_uri": "https://localhost:8080/"} outputId="5149d49a-a929-40cc-e876-6ea11d29ed13"
y = df_2["label"]
type(y)
# + id="nX_4aEyvVsJk"
#lemmatizing
wordnet_lemmatizer = WordNetLemmatizer()
x = []
x_test = []
english_words = set(nltk.corpus.words.words())
for words in df_2['text']:
tmp = []
fil_wor = [wordnet_lemmatizer.lemmatize(word, 'n') for word in words if word in english_words]
tmp.extend(fil_wor)
x.append(tmp)
for words in df_t['text']:
tmp = []
fil_wor = [wordnet_lemmatizer.lemmatize(word, 'n') for word in words if word in english_words]
tmp.extend(fil_wor)
x_test.append(tmp)
df_2["text"] = x
df_t["text"] = x_test
# + colab={"base_uri": "https://localhost:8080/"} id="wk9aMgC4VsJm" outputId="3eb7edfc-02cd-49cf-a9a8-fabb4ef0a620"
#creating word embedding
x_all = x.copy()
for z in range(len(x_test)):
x_all.append(x_test[z])
#n of vectors we are generating
EMBEDDING_DIM = 100
#Creating Word Vectors by Word2Vec Method (takes time...)
w2v_model = gensim.models.Word2Vec(sentences=x_all,size = EMBEDDING_DIM, window=5, min_count=1)
#print(len(w2v_model.wv.vocab))
#testing a word embedding
print(w2v_model.wv["liberty"])
#similarity between words
word = 'people'
w2v_model.wv.most_similar(word)
# + id="PvhjJrkOVsJp" colab={"base_uri": "https://localhost:8080/"} outputId="0010d887-d77e-44f0-95ef-1868fac5d698"
#tokenizing
tokenizer = Tokenizer()
tokenizer.fit_on_texts(x)
x = tokenizer.texts_to_sequences(x)
x_test = tokenizer.texts_to_sequences(x_test)
print(x[0][:10])
word_index = tokenizer.word_index
for word, num in word_index.items():
print(f"{word} -> {num}")
if num == 10:
break
# + id="s5cpUbpUVsJu"
#padding
maxlen = 700
x = pad_sequences(x, maxlen=maxlen)
x_test = pad_sequences(x_test, maxlen=maxlen)
# + id="6de49bfjVsJx"
# Function to create weight matrix from word2vec gensim model
def get_weight_matrix(model, vocab):
# total vocabulary size plus 0 for unknown words
vocab_size = len(vocab) + 1
# define weight matrix dimensions with all 0
weight_matrix = np.zeros((vocab_size, EMBEDDING_DIM))
# step vocab, store vectors using the Tokenizer's integer mapping
for word, i in vocab.items():
weight_matrix[i] = model[word]
return weight_matrix, vocab_size
#Getting embedding vectors from word2vec and usings it as weights of non-trainable keras embedding layer
embedding_vectors, vocab_size = get_weight_matrix(w2v_model.wv, word_index)
# + id="cNYlQ686VsJy" colab={"base_uri": "https://localhost:8080/"} outputId="0ac71178-04f8-4823-e886-7d953820eaf7"
#Defining Neural Network
model = Sequential()
#Non-trainable embeddidng layer
model.add(Embedding(vocab_size, output_dim=EMBEDDING_DIM, weights=[embedding_vectors], input_length=maxlen, trainable=False))
#LSTM
model.add(Dropout(0.2))
#model.add(Conv1D(filters=32, kernel_size=5, padding='same', activation='relu'))
#model.add(MaxPooling1D(pool_size=2))
model.add(Conv1D(filters=64, kernel_size=3, padding='same', activation='relu'))
model.add(MaxPooling1D(pool_size=2))
model.add(LSTM(units=128,dropout=0.2, return_sequences=True))
model.add(BatchNormalization())
model.add(LSTM(units=128,dropout=0.2))
model.add(BatchNormalization())
#model.add(Dropout(0.2))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['acc'])
model.summary()
# + id="_A6yCCmOVsJz" colab={"base_uri": "https://localhost:8080/"} outputId="5f25a859-b5a6-48f4-8dfd-a4779898da48"
X_train, X_test, y_train, y_test = train_test_split(x, y)
model.fit(X_train, y_train, validation_data= (X_test,y_test), epochs=50)
# + id="Di5wjb8kVsJ0" colab={"base_uri": "https://localhost:8080/", "height": 633} outputId="74b345ba-d2be-41fe-9311-0154568b5d04"
#validation_data_performance evaluation
y_pred = (model.predict(X_test) >= 0.5).astype("int")
print(accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
cm = confusion_matrix(y_test, y_pred)
plt.figure(figsize=(10,7))
sns.heatmap(cm, annot=True)
plt.xlabel('Predicted')
plt.ylabel('Truth')
# + id="3WYgcs3rVsJ1"
#test_data_for_scoring_on_kaggle
y_t = (model.predict(x_test) >= 0.5).astype("int")
result = pd.DataFrame({"id" :df_t.index, "label":y_t.squeeze() }, index = None )
result.to_csv("result_rnn.csv",index = False)
# + id="dpPU1UcgVsJ1"
#let's include an attention layer in our model
class Attention(tf.keras.Model):
def __init__(self, units):
super(Attention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, features, hidden):
hidden_with_time_axis = tf.expand_dims(hidden, 1)
score = tf.nn.tanh(self.W1(features)+ self.W2(hidden_with_time_axis))
attention_weights = tf.nn.softmax(self.V(score),axis = 1)
context_vector = attention_weights * features
context_vector = tf.reduce_sum(context_vector, axis = 1)
return context_vector, attention_weights
# + id="qx-8Zd2NVsJ2"
'''#dd attention layer to the deep learning network
class Attention(Layer):
def __init__(self,**kwargs):
super(Attention,self).__init__(**kwargs)
def build(self,input_shape):
self.W=self.add_weight(name='attention_weight', #shape=(input_shape[-1],1),
initializer='random_normal', trainable=True)
self.b=self.add_weight(name='attention_bias', #shape=(input_shape[1],1),
initializer='zeros', trainable=True)
super(Attention, self).build(input_shape)
def call(self,x):
# Alignment scores. Pass them through tanh function
e = K.tanh(K.dot(x,self.W)+self.b)
# Remove dimension of size 1
e = K.squeeze(e, axis=-1)
# Compute the weights
alpha = K.softmax(e)
# Reshape to tensorFlow format
alpha = K.expand_dims(alpha, axis=-1)
# Compute the context vector
context = x * alpha
context = K.sum(context, axis=1)
return context'''
# + id="UgU2d0BqVsJ3" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="de610116-73bd-4ea5-80bd-7dfa5a8520bc"
#RNN with Attention model
sequence_input = Input(shape = (maxlen,), dtype = "int32")
embedding = Embedding(vocab_size, output_dim=EMBEDDING_DIM, weights=[embedding_vectors], input_length=maxlen, trainable=False)(sequence_input)
dropout = Dropout(0.2)(embedding)
conv1 = Conv1D(filters=64, kernel_size=3, padding='same', activation='relu')(dropout)
maxp = MaxPooling1D(pool_size=2)(conv1)
#(lstm, state_h, state_c) = LSTM(units=128,return_sequences=True,dropout=0.2, return_state= True)(maxp)
#bn1 = BatchNormalization()((lstm, state_h, state_c))
(lstm, state_h, state_c) = LSTM(units=128,dropout=0.2, return_sequences=True, return_state= True)(maxp)
context_vector, attention_weights = Attention(10)(lstm, state_h)
densee = Dense(20, activation='relu')(context_vector)
#bn = BatchNormalization()(densee)
dropout2 = Dropout(0.2)(densee)
densef = Dense(1, activation='sigmoid')(dropout2)
model = tensorflow.keras.Model(inputs = sequence_input, outputs = densef)
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['acc'])
display(model.summary())
history = model.fit(X_train, y_train, validation_data= (X_test,y_test), epochs=50)
# + id="8E8qApwtVsJ3" colab={"base_uri": "https://localhost:8080/", "height": 633} outputId="8b6bbd43-289c-4b9f-cfe8-8f292f5d64ee"
#checking accuracy on validation data
y_pred = (model.predict(X_test) >= 0.5).astype("int")
print(accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
cm = confusion_matrix(y_test, y_pred)
plt.figure(figsize=(10,7))
sns.heatmap(cm, annot=True)
plt.xlabel('Predicted')
plt.ylabel('Truth')
# + colab={"base_uri": "https://localhost:8080/"} id="Bzn7vHP0NEAq" outputId="e88900f1-c072-4a19-9507-b5416afa9173"
history.history.keys()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="hIfcmBpi8aMf" outputId="04e9c289-8945-4b93-d6ff-c09b34dd70a3"
# summarize history for accuracy
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="QuFvB8zk8bLd" outputId="0ddfcd4e-5755-458d-8f05-80899415b9ce"
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# + id="SYxvZc7BVsJ4"
y_t = (model.predict(x_test) >= 0.5).astype("int")
result = pd.DataFrame({"id" :df_t.index, "label":y_t.squeeze() }, index = None )
result.to_csv("result_rnnattenion.csv",index = False)
# + id="7wxtEutLVsJ4"
|
Notebooks/fake_news_rnn_variants_.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''base'': conda)'
# name: python3
# ---
# # Week 3 handin
# ## 01 Assignment
# Here is the text for the assignment linked to
# ## 02 Status
# Here is a status on the handin. How far you got. What is implemented and what is not
# ## 03 Solution part 1
# ## Ex1 Use data from Danmarks Statistik - Databanken
# 1. Go to https://www.dst.dk/da/Statistik/brug-statistikken/muligheder-i-statistikbanken/api#testkonsol
# 2. Open 'Konsol' and click 'Start Konsol'
# 3. In the console at pt 1: choose 'Retrieve tables', pt 2: choose get request and json format and pt 3: execute:
# 1. check the result
# 2. in the code below this same get request is used to get information about all available data tables in 'databanken'.
# 4. Change pt. 1 in the console to 'Retrieve data', pt 2: 'get request' and Table id: 'FOLK1A', format: csv, delimiter: semicolon and click: 'Variable and value codes' and choose some sub categories (Hint: hover over the codes to see their meaning). Finally execute and see what data you get.
# 5. With data aggregation and data visualization answer the following questions:
# 1. What is the change in pct of divorced danes from 2008 to 2020?
# 2. Which of the 5 biggest cities has the highest percentage of 'Never Married' in 2020?
# 3. Show a bar chart of changes in marrital status in Copenhagen from 2008 till now
# 4. Show 2 plots in same figure: 'Married' and 'Never Married' for all ages in DK in 2020 (Hint: x axis is age from 0-125, y axis is how many people in the 2 categories). Add lengend to show names on graphs
#
# ## Ex2 Use another table (extra)
# Choose any of the other tables in 'databanken' to find interesting data.
# 1. Collect the data
# 2. Pose 5 or more interesing questions to the data
# 3. Answer the questions by aggregating the data
# 4. Illustrate the answers with visual plots
import pandas as pd
# +
# 8. Change in pct of divorced danes from 2008 to 2020
url = "https://api.statbank.dk/v1/data/FOLK1A/CSV?delimiter=Semicolon&CIVILSTAND=F&Tid=*"
divorced = pd.read_csv(url, sep=";", encoding="utf-8")
def get_percent():
first = divorced.iloc[0, 2]
last = divorced.iloc[-1, 2]
return (last - first) / first * 100
print(f"change in percent: {round(get_percent(), 2)}%")
# +
# 9. Which of the 5 biggest cities has the highest percentage of 'Never Married' in 2020?
url = "https://api.statbank.dk/v1/data/FOLK1A/CSV?delimiter=Semicolon&Tid=2020K1%2C2020K2%2C2020K3%2C2020K4&OMR%C3%85DE=*&CIVILSTAND=U%2CG%2CE%2CF"
data = pd.read_csv(url, sep=";", encoding="utf-8")
no_all_cities = data[data["OMRÅDE"] != "Hele landet"]
def get_five_biggest():
biggest_cities = no_all_cities.sort_values("INDHOLD", ascending=False)
return biggest_cities[:20]
five_biggest = get_five_biggest()
print(five_biggest[five_biggest.CIVILSTAND == "Ugift"].iloc[0])
# +
import matplotlib.pyplot as plt
# 3. Show a bar chart of changes in marrital status in Copenhagen from 2008 till now
url = "https://api.statbank.dk/v1/data/FOLK1A/CSV?delimiter=Semicolon&OMR%C3%85DE=101&CIVILSTAND=G&Tid=*"
data = pd.read_csv(url, sep=";", encoding="utf-8")
def get_calculated_years():
currentYear = ""
currentTime = 0
time = dict()
for idx, year in enumerate(data["TID"]):
year = year[:4]
if currentYear == "":
currentYear = year
if currentYear == year:
currentTime = currentTime + data.iloc[idx]["INDHOLD"]
else:
time.setdefault(year, currentTime)
currentYear = year
currentTime = 0
return time
years = get_calculated_years()
print(years)
plt.plot(list(years.keys()), list(years.values()))
# +
# 4. Show 2 plots in same figure: 'Married' and 'Never Married' for all ages in DK in 2020
# (Hint: x axis is age from 0-125, y axis is how many people in the 2 categories). Add lengend to show names on graphs
url = "https://api.statbank.dk/v1/data/FOLK1A/CSV?delimiter=Semicolon&CIVILSTAND=G%2CU&ALDER=*&Tid=2020K1%2C2020K2%2C2020K3%2C2020K4"
data = pd.read_csv(url, sep=";", encoding="utf-8")
data = data[data["ALDER"] != "I alt"]
plt.figure(figsize= (20, 8))
def get_plot(civil_stand: str):
values = data[data["CIVILSTAND"] == civil_stand]
x_values = [x.split(" ")[0] for x in values["ALDER"]]
plt.plot(x_values, values["INDHOLD"])
get_plot("Gift/separeret")
get_plot("Ugift")
plt.legend()
# -
# ## 04 Solution part 2
|
Week_5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import style
style.use("ggplot")
from sklearn.cluster import KMeans
from sklearn.metrics import accuracy_score
from sklearn.cross_validation import train_test_split
import warnings
warnings.simplefilter('ignore')
df=pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/water-treatment/water-treatment.data')
df.shape
# 1 Q-E (input flow to plant)
# 2 ZN-E (input Zinc to plant)
# 3 PH-E (input pH to plant)
# 4 DBO-E (input Biological demand of oxygen to plant)
# 5 DQO-E (input chemical demand of oxygen to plant)
# 6 SS-E (input suspended solids to plant)
# 7 SSV-E (input volatile supended solids to plant)
# 8 SED-E (input sediments to plant)
# 9 COND-E (input conductivity to plant)
# 10 PH-P (input pH to primary settler)
# 11 DBO-P (input Biological demand of oxygen to primary settler)
# 12 SS-P (input suspended solids to primary settler)
# 13 SSV-P (input volatile supended solids to primary settler)
# 14 SED-P (input sediments to primary settler)
# 15 COND-P (input conductivity to primary settler)
# 16 PH-D (input pH to secondary settler)
# 17 DBO-D (input Biological demand of oxygen to secondary settler)
# 18 DQO-D (input chemical demand of oxygen to secondary settler)
# 19 SS-D (input suspended solids to secondary settler)
# 20 SSV-D (input volatile supended solids to secondary settler)
# 21 SED-D (input sediments to secondary settler)
# 22 COND-D (input conductivity to secondary settler)
# 23 PH-S (output pH)
# 24 DBO-S (output Biological demand of oxygen)
# 25 DQO-S (output chemical demand of oxygen)
# 26 SS-S (output suspended solids)
# 27 SSV-S (output volatile supended solids)
# 28 SED-S (output sediments)
# 29 COND-S (output conductivity)
# 30 RD-DBO-P (performance input Biological demand of oxygen in primary settler)
# 31 RD-SS-P (performance input suspended solids to primary settler)
# 32 RD-SED-P (performance input sediments to primary settler)
# 33 RD-DBO-S (performance input Biological demand of oxygen to secondary settler)
# 34 RD-DQO-S (performance input chemical demand of oxygen to secondary settler)
# 35 RD-DBO-G (global performance input Biological demand of oxygen)
# 36 RD-DQO-G (global performance input chemical demand of oxygen)
# 37 RD-SS-G (global performance input suspended solids)
# 38 RD-SED-G (global performance input sediments)
#
#
headers= [x for x in range(0,39)]
df.columns=headers
df.sample()
df=df.replace('?', df.replace(['?'], [None]))
# +
# df.isnull().sum()
# -
# converting to float from string
for i in range(1,39):
df[i]=df[i].astype(float)
for column in range(1,39):
df[column]=df[column].fillna(int(df[column].mean()))
df=df.drop(0,axis=1)
train,test=train_test_split(df,test_size=0.4)
train.shape
test.shape
colors = ["g.","r.","c.","y.","k.",'-c.','r.','g.']
# +
kmeans6 = KMeans(n_clusters=6)
kmeans6.fit(train)
centroids = kmeans6.cluster_centers_
labels = kmeans6.labels_
# -
centroids[0]
labels
df=pd.read_csv("https://dl.dropboxusercontent.com/u/75194/stats/data/01_heights_weights_genders.csv")
df.sample(5)
df.drop("Gender",axis=1,inplace=True)
df.sample(5)
train,test=train_test_split(df,test_size=0.4)
# +
kmeans6 = KMeans(n_clusters=2)
kmeans6.fit(train)
centroids = kmeans6.cluster_centers_
labels = kmeans6.labels_
# -
centroids
labels
def toList(df):
idx =df.index.tolist()
columns=df.columns
l=[]
for i in idx:
temp=[]
for j in columns:
a=df.get_value(i,j)
temp.append(a)
l.append(temp)
return l
lis=toList(train)
# +
for i in range(len(lis)):
# print("coordinate:",lis[i], "label:", labels[i])
plt.plot(lis[i][0], lis[i][1], colors[labels[i]], markersize = 10)
# plt.show()
plt.scatter(centroids[:, 0],centroids[:, 1], marker = "x", s=150, linewidths = 5, zorder = 10)
plt.show()
# -
|
Clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 12: Discrete vs Continuous Distributions
#
#
# ## Stat 110, Prof. <NAME>, Harvard University
#
# ----
# ## Discrete vs Continuous Random Variables
#
# So, we've completed our whirlwind introduction of discrete random variables, covering the following:
#
# 1. Bernoulli
# 1. Binomial
# 1. Hypergeometric
# 1. Geometric
# 1. Negative Binomial
# 1. Poisson
#
# As we now move into continuous random variables, let's create a table to compare/contrast important random variable properties and concepts.
#
# | Discrete | Continuous |
# | ------------- |---------------|
# | $X$ | $X$ |
# | PMF $P(X=x)$ | PDF $f_x(x) = F^\prime(x)$<br/>note $P(X=x)=0$ |
# | CDF $F_x(x)=P(X \le x)$| CDF $F_x(x) = P(X \le x)$ |
# | $\mathbb{E}(X) = \sum_{x} x P(X=x)$ | $\mathbb{E}(X) = \int_{-\infty}^{\infty} x f(x)dx$ |
# | $Var(X) = \mathbb{E}X^2 - \mathbb{E}(X)^2$ | $Var(X) = \mathbb{E}X^2 - \mathbb{E}(X)^2$ |
# | $SD(X) = \sqrt{Var(X)}$ | $SD(X) = \sqrt{Var(X)}$ |
# | LOTUS $\mathbb{E}(g(x)) = \sum_{x} g(x) P(X=x)$ | LOTUS $\mathbb{E}( g(x) ) = \int_{-\infty}^{\infty} g(x) f(x)dx$ |
#
# ----
# ## Probability Density Function
#
# In the _discrete_ case, we could calculate probability by __summing__ (counting) the discrete elements, since each element represents a bit of mass. The probability would be the total of all elements concerned, divided by total mass.
#
# But we cannot count discrete elements in the _continuous_ case. Instead, we __integrate__ the density function over a range to get probability mass per _area_.
#
# #### Definition: random variable
#
# > A __random variable__ $X$ has PDF $f(x)$ if for all $a$ and $b$
# >
# > \\begin{align}
# > & P(a \le x \le b) = \int_a^b f(x) dx \\\\
# > \\end{align}
#
# ### Test for validity
#
# Note that to be a valid PDF,
#
# 1. $f(x) \ge 0$
# 1. $\int_{-\infty}^{\infty} f(x) = 1$
#
#
# ### Probability at a _point_ is 0
#
# - $a = b \Rightarrow \int_{a}^{a} f(x)dx = 0$
#
#
# ### Density $\times$ Length
#
# But for some point $x_0$ and some _very, very small value_ $\epsilon$, we can derive $f(x_0) \epsilon \approx P(X \in (x_0-\frac{\epsilon}{2}, x_0+\frac{\epsilon}{2})$
#
#
#
# ----
# ## Cumulative Distribution Function
#
# ### Deriving CDF from PDF
#
# If continuous r.v. $X$ has PDF $f$, the CDF is
#
# \begin{align}
# F(x) &= P(X \le x) \\
# &= \int_{-\infty}^{x} f(t) dt \\
# \\
# \Rightarrow P(a \le x \le b) &= \int_{a}^{b} f(x)dx \\
# &= F(b) - F(a) \\
# \end{align}
#
# ### Deriving PDF from CDF
#
# If continuous r.v. $X$ has CDF $F$ (and $X$ is continuous), the PDF is
#
# \begin{align}
# f(x) &= F^\prime(x) & &\text{ by the Fundamental Theorem of Calculus} \\
# \end{align}
#
# ----
# ## Variance
#
# Mean only tells you where the average is. Another useful statistic is the _variance_ of a random variable, which tells you how the random variable is spread out around the mean.
#
# In other words, _variance_ answers the question _How far is $X$ from its mean, on average?_
#
# #### Definition: variance
# > Variance is a measure of how a random variable is spread about its mean.
# >
# > \\begin{align}
# > \operatorname{Var}(X) &= \mathbb{E}(X - \mathbb{E}X)^2 & \quad \text{or alternatively} \\\\
# > \\\\
# > &= \mathbb{E}X^2 - 2X(\mathbb{E}X) + \mathbb{E}(X^2) & \quad \text{by Linearity}\\\\
# > &= \boxed{\mathbb{E}X^2 - \mathbb{E}(X)^2}
# > \\end{align}
#
# Sometimes the second form of variance is easier to use.
#
# Note that the formula variance is the same for both discrete and continuous r.v.
#
# But you might be wondering right now how to calculate $\mathbb{E}X^2$. We will get to that in a bit...
#
# ----
# ## Standard Deviation
#
# But note that variance is expressed in terms of units __squared__. _Standard deviation_ is sometimes easier to use than variance, as it is given in the original units.
#
# #### Definition: standard deviation
# > The _standard deviation_ the square root of the variance.
# >
# > \\begin{align}
# > SD(X) &= \sqrt{\operatorname{Var}(X)}
# > \\end{align}
#
# Note that like variance, the formula for standard deviation is the same for both discrete and continuous r.v.
#
# ----
# ## Uniform Distribution
#
# ### Description
#
# The simplest and perhaps the most famous continuous distribution. Given starting point $a$ and ending point $b$, probability $\propto$ length.
#
# 
#
# ### Notation
#
# $X \sim \operatorname{Unif}(a,b)$
#
# ### Parameters
#
# - $a$ start of the segment, $a < b$
# - $b$ end of the segment, $b > a$
#
# ### Probability density function
#
# \begin{align}
# f(x) &=
# \begin{cases}
# c & \quad \text{ if } a \le x \le b \\
# 0 & \quad \text{ otherwise }
# \end{cases} \\
# \\
# \\
# 1 &= \int_{a}^{b} c dx \\
# \Rightarrow c &= \boxed{\frac{1}{b-a}}
# \end{align}
#
# ### Cumulative distribution function
#
# \begin{align}
# F(x) &= \int_{-\infty}^{x} f(t)dt \\
# &= \int_{a}^{x} f(t)dt \\
# &=
# \begin{cases}
# 0 & \quad \text{if } x \lt a \\
# \frac{x-a}{b-a} & \quad \text{if } a \lt x \lt b \\
# 1 & \quad \text{if } x \gt b
# \end{cases} \\
# \end{align}
#
# So this means that as $X$ increase, its probability increase likewise in a _linear_ fashion.
#
# ### Expected value
#
# For continuous r.v.
#
# \begin{align}
# \mathbb{E}(X) &= \int_{a}^{b} x \frac{1}{b-a} dx \\
# &=\left. \frac{x^2}{2(b-a)} ~~ \right\vert_{a}^{b} \\
# &= \frac{(b^2-a^2)}{2(b-a)} \\
# &= \boxed{\frac{b+a}{2}}
# \end{align}
#
# ### Variance
#
# Remember that lingering doubt about $\mathbb{E}X^2$?
#
# Let random variable $Y = X^2$.
#
# \begin{align}
# \mathbb{E}X^2 &= \mathbb{E}(Y) \\
# &\stackrel{?}{=} \int_{-\infty}^{\infty} x^2 f(x) dx & &\text{since we need the PDF of Y..?}
# \end{align}
#
# ----
# ## Law of the Unconscious Statistician (LOTUS)
#
# Actually, that last bit of wishful thinking is correct and will work in both the discrete and continuous cases.
#
# In general for continuous r.v.
#
# \begin{align}
# \mathbb{E}( g(x) ) = \int_{-\infty}^{\infty} g(x) f(x)dx
# \end{align}
#
# And likewise for discrete r.v.
#
# \begin{align}
# \mathbb{E}(g(x)) = \sum_{x} g(x) P(X=x)
# \end{align}
#
# ### Variance of $U \sim \operatorname{Unif}(0,1)$
#
# \begin{align}
# \mathbb{E}(U) &= \frac{1}{b-a} \\
# &= \frac{1}{2} \\
# \\
# \\
# \mathbb{E}U^2 &= \int_{0}^{1} u^2 \underbrace{f(u) du}_{1} \\
# &= \left.\frac{u^3}{3} ~~ \right\vert_{0}^{1} \\
# &= \frac{1}{3} \\
# \\
# \\
# \Rightarrow Var(U) &= \mathbb{E}U^2 - \mathbb{E}(U)^2 \\
# &= \frac{1}{3} - \left(\frac{1}{2}\right)^2 \\
# &= \frac{1}{12}
# \end{align}
#
# ----
# ## Universality of the Uniform
#
# Given an arbitrary CDF $F$ and the uniform $\operatorname{U} \sim \operatorname{Unif}(0,1)$, it is possible to simulate a draw from the continuous r.v. of the CDF $F$.
#
# Assume:
#
# 1. $F$ is strictly increasing
# 1. $F$ is continuous as a function
#
# If we define $X = F^{-1}(U)$. Then $X \sim F$.
#
# \begin{align}
# P(X \le x) &= P(F^{-1}(U) \le x) \\
# &= P(U \le F(x)) \\
# &= F(x) & \quad \text{ since } P(U \le u) \propto 1~~ \blacksquare
# \end{align}
#
# 
#
# ----
# View [Lecture 12: Discrete vs. Continuous, the Uniform | Statistics 110](http://bit.ly/2wex5yh) on YouTube.
|
Lecture_12.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import astropy as ap
import matplotlib.pyplot as plt
data = np.genfromtxt('data/obj5.data')
print(data)
plt.scatter(data[:,1], data[:,2], c = data[:,3])
plt.colorbar()
plt.show()
plt.hist(data[:,3], bins = 30)
# +
wantedx = []
for i in range(len(data[:,1])):
if data[i,3]<= 0.01:
wantedx.append(i)
wantedx = np.array(wantedx)
newdatax = data[wantedx,:]
plt.scatter(newdatax[:,1], newdatax[:,2], c = newdatax[:,3])
plt.xlabel('Dec')
plt.ylabel('RA')
plt.colorbar()
plt.show()
# +
wanted = []
for i in range(len(data[:,1])):
if data[i,3]<= 0.01 and data[i,1]<=10:
wanted.append(i)
wanted = np.array(wanted)
newdata = data[wanted,:]
plt.scatter(newdata[:,1], newdata[:,2], c = newdata[:,3])
plt.xlabel('Dec')
plt.ylabel('RA')
plt.colorbar()
plt.show()
plt.hist(newdata[:,3]*1000, bins=30)
plt.xlabel('Z*1000')
plt.show()
fig = plt.figure()
ax = fig.add_subplot(111, projection='polar')
c = ax.scatter(newdata[:,2], newdata[:,1], c=newdata[:,3])
fig.colorbar(c)
plt.show()
# -
avg_redshift= np.average(newdata[:,3])
print(avg_redshift)
std_dev = np.std(newdata[:,3])
print(std_dev)
|
day4/Object 5 .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
# ### In this kernel, I will walk you through some extra text cleaning methods and how they work on sample comments.
# <center><img src="https://i.imgur.com/CtyQ8Ag.png" width="250px"></center>
# ## Acknowledgements
# I have borrowed the text cleaning functions from Dimitrios in [this great kernel](https://www.kaggle.com/deffro/text-pre-processing-techniques) from the Quora Insincere Questions Classification competition.
# ### Import necessary libraries
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import os
import gc
import re
import numpy as np
import pandas as pd
import nltk
from nltk.corpus import wordnet, stopwords
from nltk.stem import WordNetLemmatizer
from nltk.stem.porter import PorterStemmer
from colorama import Fore, Back, Style
# -
# ### Download data
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
train_df = pd.read_csv('../input/train.csv')
# -
# ### See first few rows of dataframe
train_df.head()
# ### Extract comments from data
comments = train_df['comment_text']
# ### Create function for visualizing the effect of text cleaning function
def example_cleaning_results(function):
select_comments = []
for i, comment in enumerate(comments):
if comment != function(comment):
select_comments.append(comment)
if len(select_comments) == 5:
break
print(" " +\
f'{Style.DIM}'+\
"EXAMPLE WORKING OF TEXT CLEANING FUNCTION"+\
f'{Style.RESET_ALL}')
print(" " +\
f'{Style.DIM}'+\
"-------------------------------------------"+\
f'{Style.RESET_ALL}')
print("")
for comment in select_comments:
print(f'{Fore.YELLOW}{Style.DIM}' + comment + f'{Style.RESET_ALL}' +\
'\n\n' + " "+\
'CHANGES TO' + '\n\n' +\
f'{Fore.CYAN}{Style.DIM}' + function(comment) + f'{Style.RESET_ALL}')
print("")
print(f'{Fore.WHITE}{Style.DIM}' +\
"-------------------------"+\
"-------------------------"+\
"-------------------------"+\
"------------------" +\
f'{Style.RESET_ALL}')
# ### Remove the numbers
# This function removes all the numbers in the comment
#
# Eg. I'm 25 years old. --> I'm years old.
def remove_numbers(text):
""" Removes integers """
text = ''.join([i for i in text if not i.isdigit()])
return text
example_cleaning_results(remove_numbers)
# ### Remove the exclamation, question and full stop marks
# This function removes the exclamation, question and full stop marks from the comment.
#
# Eg. This is awesome ! --> This is awesome
# +
def replace_multi_exclamation_mark(text):
""" Replaces repetitions of exlamation marks """
text = re.sub(r"(\!)\1+", ' multiExclamation ', text)
return text
def replace_multi_question_mark(text):
""" Replaces repetitions of question marks """
text = re.sub(r"(\?)\1+", ' multiQuestion ', text)
return text
def replace_multi_stop_mark(text):
""" Replaces repetitions of stop marks """
text = re.sub(r"(\.)\1+", ' multiStop ', text)
return text
# -
example_cleaning_results(lambda x: replace_multi_exclamation_mark(replace_multi_question_mark(replace_multi_stop_mark(x))))
# ### Remove the exclamation, question and full stop marks
# This function removes the exclamation, question and full stop marks from the comment.
#
# Eg. You love cats !? I prefer dogs. --> You love cats I prefer dogs
# +
contraction_patterns = [(r'won\'t', 'will not'), (r'can\'t', 'cannot'), (r'i\'m', 'i am'),\
(r'ain\'t', 'is not'), (r'(\w+)\'ll', '\g<1> will'),\
(r'(\w+)n\'t', '\g<1> not'),\
(r'(\w+)\'ve', '\g<1> have'), (r'(\w+)\'s', '\g<1> is'),\
(r'(\w+)\'re', '\g<1> are'), (r'(\w+)\'d', '\g<1> would'),\
(r'&', 'and'), (r'dammit', 'damn it'), (r'dont', 'do not'),\
(r'wont', 'will not')]
def replace_contraction(text):
patterns = [(re.compile(regex), repl) for (regex, repl) in contraction_patterns]
for (pattern, repl) in patterns:
(text, count) = re.subn(pattern, repl, text)
return text
# -
example_cleaning_results(replace_contraction)
# ### Replace the negations with antonyms
# This function rplaces negations with their respective antonyms.
#
# Eg. I am not happy. --> I am unhappy.
# +
def replace(word, pos=None):
""" Creates a set of all antonyms for the word and if there is only one antonym, it returns it """
antonyms = set()
for syn in wordnet.synsets(word, pos=pos):
for lemma in syn.lemmas():
for antonym in lemma.antonyms():
antonyms.add(antonym.name())
if len(antonyms) == 1:
return antonyms.pop()
else:
return None
def replace_negations(text):
""" Finds "not" and antonym for the next word and if found, replaces not and the next word with the antonym """
i, l = 0, len(text)
words = []
while i < l:
word = text[i]
if word == 'not' and i+1 < l:
ant = replace(text[i+1])
if ant:
words.append(ant)
i += 2
continue
words.append(word)
i += 1
return words
def tokenize_and_replace_negations(text):
tokens = nltk.word_tokenize(text)
tokens = replace_negations(tokens)
text = " ".join(tokens)
return text
# -
example_cleaning_results(tokenize_and_replace_negations)
# ### Remove stopwords
# This function removes the most common words used in English (stop words) like 'a', 'is', 'are' etc.
#
# Eg. He is a very humorous person. --> He very humorous person.
# +
stoplist = stopwords.words('english')
def remove_stop_words(text):
finalTokens = []
tokens = nltk.word_tokenize(text)
for w in tokens:
if (w not in stoplist):
finalTokens.append(w)
text = " ".join(finalTokens)
return text
# -
example_cleaning_results(remove_stop_words)
# ### Replace elongated words with the basic form
# This function replaces elongated words with its basic form.
#
# Eg. I eat little food --> I eat litle food
# +
def replace_elongated(word):
""" Replaces an elongated word with its basic form, unless the word exists in the lexicon """
repeat_regexp = re.compile(r'(\w*)(\w)\2(\w*)')
repl = r'\1\2\3'
if wordnet.synsets(word):
return word
repl_word = repeat_regexp.sub(repl, word)
if repl_word != word:
return replace_elongated(repl_word)
else:
return repl_word
def replace_elongated_words(text):
finalTokens = []
tokens = nltk.word_tokenize(text)
for w in tokens:
finalTokens.append(replace_elongated(w))
text = " ".join(finalTokens)
return text
# -
example_cleaning_results(replace_elongated_words)
# ### Stem words
# This function "stems" the words in the comments. It only keeps the stem of the word, which need not be an actual word.
#
# Eg. I love swimming and driving happily --> I love swimm and driv happi
# +
stemmer = PorterStemmer()
def stem_words(text):
finalTokens = []
tokens = nltk.word_tokenize(text)
for w in tokens:
finalTokens.append(stemmer.stem(w))
text = " ".join(finalTokens)
return text
# -
example_cleaning_results(stem_words)
# ### Lemmatize words
# The function lemmatizes the words in the comments. It only keeps the lemma of the actual words, which needs to be an actual word.
#
# Eg. I love swimming and driving happily --> I love swim and drive happy
# +
lemmatizer = WordNetLemmatizer()
def lemmatize_words(text):
finalTokens = []
tokens = nltk.word_tokenize(text)
for w in tokens:
finalTokens.append(lemmatizer.lemmatize(w))
text = " ".join(finalTokens)
return text
# -
example_cleaning_results(lemmatize_words)
# ### That's it ! Thanks for reading my kernel and I hope you found it useful. Your upvote will be appreciated :)
|
4 jigsaw/jigsaw-competition-more-text-cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Chapter: Decision Trees and Ensemble Learning
#
#
# # Topic: Robust fitting via Bagging: Illustration
# import
import numpy as np
# generate training samples
from sklearn import datasets
noisy_moons = datasets.make_moons(n_samples=200, noise=0.3, random_state=10)
X, y = noisy_moons
# fit DT model
from sklearn import tree
DT_model = tree.DecisionTreeClassifier().fit(X, y)
# +
# plot decision surface of the decision tree
import matplotlib.pyplot as plt
plt.figure()
cmap = plt.cm.RdBu
# Plot the decision boundary
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02), np.arange(y_min, y_max, 0.02))
# create grid to evaluate model
Z = DT_model.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=cmap, alpha=0.2)
# plot the training data-points
plt.scatter(X[:, 0], X[:, 1], marker="o", c=y, cmap=cmap, s=25, edgecolor="k")
plt.xlabel('x1'), plt.ylabel('x2')
plt.show()
# -
# fit bagging model
from sklearn.ensemble import BaggingClassifier
Bagging_model = BaggingClassifier(n_estimators=500, max_samples=50, random_state=100).fit(X, y)
# +
# plot decision surface of the decision tree
plt.figure()
# Plot the decision boundary
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02), np.arange(y_min, y_max, 0.02))
# create grid to evaluate model
Z = Bagging_model.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=cmap, alpha=0.2)
# plot the trainign data-points
plt.scatter(X[:, 0], X[:, 1], marker="o", c=y, cmap=cmap, s=25, edgecolor="k")
plt.xlabel('x1'), plt.ylabel('x2')
plt.show()
|
Chapter_DecisionTrees_EnsembleLearning/Bagging_illustration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Tz-zgucvuT2m" executionInfo={"status": "ok", "timestamp": 1624545402919, "user_tz": -60, "elapsed": 2665, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}}
import io
import math
import keras
import pandas as pd
import numpy as np
import requests
import matplotlib.pyplot as plt
from keras.models import Sequential, model_from_json
from keras.layers import Dense, LSTM, Dropout
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from google.colab import files
plt.style.use('default')
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="ues3ln6iOawo" executionInfo={"status": "ok", "timestamp": 1624545403602, "user_tz": -60, "elapsed": 686, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="75a62e97-20d9-4fd2-f8de-e403eee1c840"
# https://finance.yahoo.com/quote/AAPL/history?period1=345427200&period2=1624233600&interval=1d&filter=history&frequency=1d&includeAdjustedClose=true (Max Historical Apple Daily)
resp = requests.get(f'https://query1.finance.yahoo.com/v7/finance/download/AAPL?period1=345427200&period2=1624233600&interval=1d&events=history&includeAdjustedClose=true') # download link
csv_data = io.StringIO(resp.content.decode('utf8').replace("'", '"'))
df = pd.read_csv(csv_data, sep=',')
df
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="8w3CQB0chlLJ" executionInfo={"status": "ok", "timestamp": 1624545403603, "user_tz": -60, "elapsed": 16, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="5361c020-ced1-4a41-b5ed-5d295d3a8532"
df['EMA12'] = df['Adj Close'].ewm(span=12, adjust=False).mean()
df['EMA26'] = df['Adj Close'].ewm(span=26, adjust=False).mean()
df['MACD'] = df['EMA12'] - df['EMA26']
df['Signal'] = df['MACD'].ewm(span=9, adjust=False).mean()
df['OBV'] = np.where(df['Adj Close'] == df['Adj Close'].shift(1), 0, np.where(df['Adj Close'] > df['Adj Close'].shift(1), df['Volume'],
np.where(df['Adj Close'] < df['Adj Close'].shift(1), -df['Volume'], df.iloc[0]['Volume']))).cumsum() # on-balance volume
df['OBV Percent'] = df['OBV'].pct_change() * 100
df['OBV Percent'] = round(df['OBV Percent'].fillna(0), 2) # on-balance volume as a percentage
df['Adj Close Delta'] = df['Adj Close'].diff().fillna(0) # delta between adj close
df['Next Adj Close'] = df['Adj Close'].shift(-1)
df.dropna(inplace=True)
df
# + colab={"base_uri": "https://localhost:8080/"} id="vqYRlYZWdufJ" executionInfo={"status": "ok", "timestamp": 1624545485248, "user_tz": -60, "elapsed": 187, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="4a2a9133-2c27-4701-f2be-e4e480e967e3"
df_X = df[['Open','High','Low','Close','Volume','EMA12','EMA26','MACD','Signal','OBV Percent','Adj Close Delta']]
df_y = df[['Next Adj Close']]
X_scaler = MinMaxScaler(feature_range=(0, 1))
X_scaled = X_scaler.fit_transform(df_X)
df_X = pd.DataFrame(X_scaled, columns=[df_X.columns])
y_scaler = MinMaxScaler(feature_range=(0, 1))
y_scaled = y_scaler.fit_transform(df_y)
df_y = pd.DataFrame(y_scaled, columns=[df_y.columns])
X_train, X_test, y_train, y_test = train_test_split(X_scaled, y_scaled, shuffle=False, stratify=None)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# + id="GHGKVTvXaDKK" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1624545531642, "user_tz": -60, "elapsed": 184, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="1edd856f-913a-4348-9638-34c7132a9b9b"
window = 1 # 1 day
X_train_window=[]
y_train_window=[]
for i in range(window, len(X_train)):
X_train_window.append(X_scaled[i-window:i, :])
y_train_window.append(X_scaled[i, 0])
X_train_window, y_train_window = np.array(X_train_window), np.array(y_train_window)
X_train_window = np.reshape(X_train_window, (X_train_window.shape[0], X_train_window.shape[1], X_train.shape[1]))
print ('X_train_window.shape', X_train_window.shape)
print ('y_train_window.shape', y_train_window.shape)
# + id="oML26KP1aEIh" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1624545601987, "user_tz": -60, "elapsed": 49933, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="b67e4063-dce5-4257-93f9-581f3dbab7cf"
def create_model():
model = Sequential()
model.add(LSTM(units=50, return_sequences=True, input_shape=(X_train_window.shape[1], X_train.shape[1]))) # layer 1 lstm
model.add(Dropout(0.2)) # layer 1 dropout regularisation
model.add(LSTM(units=50, return_sequences=True)) # layer 2 lstm
model.add(Dropout(0.2)) # layer 2 dropout regularisation
model.add(LSTM(units=50, return_sequences=True)) # layer 3 lstm
model.add(Dropout(0.2)) # layer 3 dropout regularisation
model.add(LSTM(units=50)) # layer 4 lstm
model.add(Dropout(0.2)) # layer 4 dropout regularisation
model.add(Dense(units=1)) # output layer
model.compile(optimizer='adam', loss='mse', metrics=['mae']) # compile the rnn
return model
model = create_model()
history = model.fit(X_train_window, y_train_window, epochs=300, batch_size=1500, verbose=2)
model.summary()
for metric in history.history:
plt.title(metric)
plt.plot(history.history[metric])
plt.show()
# + id="_Pwg5momQELY" colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"status": "ok", "timestamp": 1624281332400, "user_tz": -60, "elapsed": 263, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="9fe06eb2-15bd-4267-d2fb-469616970539"
'''Optionally save model, model.json and weights.h5'''
model_filename = f'model_apple_daily_{len(X_train.columns)}-inputs.json'
weights_filename = f'weights_apple_daily_{len(X_train.columns)}-inputs.h5'
# !ls /content
# save structure to json
model_json = model.to_json()
with open(model_filename, 'w') as json_file:
json_file.write(model_json)
# save weights to hdf5
model.save_weights(weights_filename)
files.download(f'/content/{model_filename}')
files.download(f'/content/{weights_filename}')
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 157} id="Zs50-wzE1fZE" executionInfo={"status": "ok", "timestamp": 1624278452191, "user_tz": -60, "elapsed": 21020, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="6dec0ae3-43ca-40d1-c4e8-3d851ee935bb"
'''Optionally load model, model.json and weights.h5'''
model_filename = f'model_apple_daily_{len(X_train.columns)}-inputs.json'
weights_filename = f'weights_apple_daily_{len(X_train.columns)}-inputs.h5'
try:
files.upload()
# !ls /content
# read structure from json
model = open(model_filename, 'r')
json = model.read()
model.close()
model = model_from_json(json)
# read weights from hdf5
model.load_weights(f'/content/{weights_filename}')
except Exception as e:
print (e)
# + id="LQP2ZA0txrhZ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1624545930185, "user_tz": -60, "elapsed": 2671, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="c8c0accf-5ef1-45aa-ab33-b236b8880723"
X_test_window=[]
y_test_window=[]
for i in range(window, len(X_test)):
X_test_window.append(X_scaled[i-window:i, :])
y_test_window.append(X_scaled[i, 0])
X_test_window, y_test_window = np.array(X_test_window), np.array(y_test_window)
X_test_window = np.reshape(X_test_window, (X_test_window.shape[0], X_test_window.shape[1], X_test.shape[1]))
print ('X_test_window.shape:', X_test_window.shape)
print ('y_test_window.shape:', y_test_window.shape)
y_pred = model.predict(X_test_window)
y_pred = y_scaler.inverse_transform(y_pred)
print ('y_pred.shape:', y_pred.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="lA6Dqqllf3Cv" executionInfo={"status": "ok", "timestamp": 1624546188026, "user_tz": -60, "elapsed": 368, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="5d6b8010-24e9-402a-af67-95f93131b10e"
y_pred.min(), y_pred.max()
# + id="xgkK5E9K5Dn5" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1624545930186, "user_tz": -60, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="408b2815-0f8a-443a-82b9-93f8a42694ce"
print ('ae:', mean_absolute_error(df['Adj Close'][-len(y_pred):].values, y_pred))
print ('mse:', mean_squared_error(df['Adj Close'][-len(y_pred):].values, y_pred, squared=False))
print ('rmse:', mean_squared_error(df['Adj Close'][-len(y_pred):].values, y_pred))
# + id="0ci9sG1zcLle" colab={"base_uri": "https://localhost:8080/", "height": 670} executionInfo={"status": "ok", "timestamp": 1624545996593, "user_tz": -60, "elapsed": 6153, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="c322fb9c-b9af-4f9c-d2a4-1743d193a869"
plt.figure(figsize=(30,10))
plt.plot(df['Date'].tail(len(y_pred)), df['Adj Close'][-len(y_pred):].values, color='red', label=f'Actual Apple Daily {df_y.columns.values[0][0].title()}')
plt.plot(df['Date'].tail(len(y_pred)), y_pred, color='blue', label=f'Predicted Apple Daily {df_y.columns.values[0][0].title()}')
plt.title(f'Apple Daily {df_y.columns.values[0][0].title()} Prediction')
plt.xlabel('Time')
plt.ylabel(f'{df_y.columns.values[0][0].title()}')
plt.xticks(rotation=90)
ax = plt.gca()
for index, label in enumerate(ax.xaxis.get_ticklabels()):
if index % 92 != 0:
label.set_visible(False)
plt.legend()
plt.show()
# + id="XZlO5cxit3oR" colab={"base_uri": "https://localhost:8080/", "height": 419} executionInfo={"status": "ok", "timestamp": 1624402166883, "user_tz": -60, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gih6jGZ_ZDRChRiZZo2DwHoBerm3MdEG4NWtpSGYg=s64", "userId": "16245925105622995030"}} outputId="df0a6b31-7d12-40e8-da1b-b651545c6cad"
pd.set_option('mode.chained_assignment', None)
df_test = df.tail(len(y_pred))
df_test.loc[:, 'Predicted Next Adj Close'] = y_pred
df_test[['Open','High','Low','Close','Volume','EMA12','EMA26','MACD','Signal','OBV Percent','Adj Close Delta','Next Adj Close','Next Adj Close']]
df_test.rename(columns={'Next Adj Close': 'Actual Next Adj Close'}, errors='raise', inplace=True)
df_test
# + id="UstG4kwolrrU"
|
TradingAI_LSTM_AppleDaily_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="UB0JdwYh4TXP" colab_type="text"
# # Ridge Regression
# + id="C7YFRZD74TXQ" colab_type="code" colab={}
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression, Ridge
from sklearn.metrics import mean_squared_error
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import MinMaxScaler, PolynomialFeatures
# + id="o_LETRC34TXW" colab_type="code" colab={}
_df = pd.read_csv('https://raw.githubusercontent.com/PacktWorkshops/The-Data-Science-Workshop/master/Chapter07/Dataset/ccpp.csv')
# + id="AJruh2jv4TXb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="c992073a-0b55-4818-f789-5fbb0b2c66a1"
_df.info()
# + id="ncMDnCFH4TXi" colab_type="code" colab={}
X = _df.drop(['PE'], axis=1).values
y = _df['PE'].values
# + id="MWeCBwia4TXs" colab_type="code" colab={}
train_X, eval_X, train_y, eval_y = train_test_split(X, y, train_size=0.8, random_state=0)
# + [markdown] id="tM3c9EIN4TXw" colab_type="text"
# # Implement a LinearRegression model
# + id="4T2HmvOL4TXx" colab_type="code" colab={}
lr_model_1 = LinearRegression()
# + id="jtxCFguq4TX0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6baaa6a0-1ee2-4081-b807-943ef0dacf4a"
lr_model_1.fit(train_X, train_y)
# + id="DmjxSQFx4TX4" colab_type="code" colab={}
lr_model_1_preds = lr_model_1.predict(eval_X)
# + id="2tjzBNFY4TX-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="af7ceef5-e293-4c03-9da9-3ca1d8449793"
print('lr_model_1 R2 Score: {}'.format(lr_model_1.score(eval_X, eval_y)))
# + id="7s2xf7ZJ4TYG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f51ebf29-81bf-4950-faf4-d145a83bb85a"
print('lr_model_1 MSE: {}'.format(mean_squared_error(eval_y, lr_model_1_preds)))
# + [markdown] id="KdDWbtcH4TYM" colab_type="text"
# # Engineer cubic features
# + id="NgKhTb974TYN" colab_type="code" colab={}
steps = [
('scaler', MinMaxScaler()),
('poly', PolynomialFeatures(degree=3)),
('lr', LinearRegression())
]
# + id="qXAuIVa_4TYS" colab_type="code" colab={}
lr_model_2 = Pipeline(steps)
# + id="a8es2V-O4TYX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 177} outputId="8be3b435-f59d-405c-f63e-2dd78dbb87c2"
lr_model_2.fit(train_X, train_y)
# + id="ctQ5S3hC4TYb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5740cb06-5003-4466-f2d7-eb5e9f55b728"
print('lr_model_2 R2 Score: {}'.format(lr_model_2.score(eval_X, eval_y)))
# + id="SuXPIB-k4TYj" colab_type="code" colab={}
lr_model_2_preds = lr_model_2.predict(eval_X)
# + id="ITfCmnHP4TYm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="65ca8ff8-08da-4d7d-c4a1-7a85fe5ddda5"
print('lr_model_2 MSE: {}'.format(mean_squared_error(eval_y, lr_model_2_preds)))
# + id="hbbvnP8H4TYp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 177} outputId="8f23907b-0bf0-484d-907c-39c88d117645"
print(lr_model_2[-1].coef_)
# + id="0HLfk7zh4TYu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="86414a27-a260-4d7e-dc99-710d60ebd800"
print(len(lr_model_2[-1].coef_))
# + [markdown] id="IurzrMU74TYz" colab_type="text"
# # Engineer polynomial features
# + id="UPaMX0Os4TY0" colab_type="code" colab={}
steps = [
('scaler', MinMaxScaler()),
('poly', PolynomialFeatures(degree=10)),
('lr', LinearRegression())
]
# + id="gNa1_R4F4TY5" colab_type="code" colab={}
lr_model_3 = Pipeline(steps)
# + id="PEvqJh5V4TY9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 177} outputId="08bd757d-1293-4d03-ee73-791a7ac114b9"
lr_model_3.fit(train_X, train_y)
# + id="Ux-fhkRp4TZE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="744336a7-c29c-48b2-9ca8-a77b7a2644e5"
print('lr_model_3 R2 Score: {}'.format(lr_model_3.score(eval_X, eval_y)))
# + id="6TCmu8ON4TZH" colab_type="code" colab={}
lr_model_3_preds = lr_model_3.predict(eval_X)
# + id="Z2AVT7xh4TZK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0a0b22cc-a49f-4b3a-a4a0-ee48a2891ca7"
print('lr_model_3 MSE: {}'.format(mean_squared_error(eval_y, lr_model_3_preds)))
# + id="nioc06jt4TZO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="525ac3aa-b84c-4fe6-f6f9-6c2bb325cd0e"
print(len(lr_model_3[-1].coef_))
# + id="QkgaxPCR4TZT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 177} outputId="37d463b6-187b-490c-b65f-e1f180d7acdd"
print(lr_model_3[-1].coef_[:35])
# + [markdown] id="mW6S5WTh4TZY" colab_type="text"
# # Implement Ridge on the same pipeline
# + id="__LdiSB84TZY" colab_type="code" colab={}
steps = [
('scaler', MinMaxScaler()),
('poly', PolynomialFeatures(degree=10)),
('lr', Ridge(alpha=0.9))
]
# + id="p0a-4cxH4TZd" colab_type="code" colab={}
ridge_model = Pipeline(steps)
# + id="ENQhe2T84TZi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="45d16816-b8fd-4d90-b7d1-53990669b268"
ridge_model.fit(train_X, train_y)
# + id="8mIo0z5-4TZl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1800a63e-8a8d-4393-bc2f-0b5431d4c8e1"
print('ridge_model R2 Score: {}'.format(ridge_model.score(eval_X, eval_y)))
# + id="bRz7syxm4TZr" colab_type="code" colab={}
ridge_model_preds = ridge_model.predict(eval_X)
# + id="Ho4NgDe_4TZu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d3c54a15-ff04-4db6-add5-be5c354ec894"
print('ridge_model MSE: {}'.format(mean_squared_error(eval_y, ridge_model_preds)))
# + id="HosEKfHd4TZy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="821e57b2-929f-4284-85bc-cd4b911aa0b2"
print(len(ridge_model[-1].coef_))
# + id="52c0ZM1a4TZ2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="2c9d310b-6dab-4357-fda1-f388fa4f6881"
print(ridge_model[-1].coef_[:35])
# + id="ier6eRNz4TZ5" colab_type="code" colab={}
|
Chapter07/Exercise7.10/Exercise7_10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Issue #4: understand data
# +
import pandas as pd
import re
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set(style='white', context='notebook', palette='deep')
# -
# ### Profiles DF
df_profiles = pd.read_csv("../data/raw/data_set_phase1/profiles.csv")
# There are no null values
df_profiles.isnull().sum().sum()
df_profiles.sample(5)
df_profiles.shape
# 63k profiles
# ### Queries DF
df_queries = pd.read_csv("../data/raw/data_set_phase1/train_queries.csv")
df_queries.shape
# 163k queries are done by anonymous users
df_queries.isnull().sum()
df_queries.sample(5)
df_queries.shape
df_queries.dtypes
df_queries.describe()
# req_time, o and d are considered objets. Let's turn them into floats
df_queries[['ox','oy']] = df_queries.o.str.split(",",expand=True,).astype(float)
df_queries[['dx','dy']] = df_queries.d.str.split(",",expand=True,).astype(float)
df_queries = df_queries.drop('o', 1)
df_queries = df_queries.drop('d', 1)
df_queries.sort_values('req_time')
df_queries['req_time'] = pd.to_datetime(df_queries['req_time'])
df_queries['day_of_week'] = df_queries['req_time'].dt.day_name()
df_queries['req_date'] = df_queries['req_time'].dt.strftime('%m-%d')
df_queries['req_hour'] = df_queries['req_time'].dt.hour
df_queries['req_minute'] = df_queries['req_time'].dt.minute
df_queries['if_holiday'] = df_queries['req_date'].astype(int)
reg_day = re.compile("-(\d\d) ")
reg_month = re.compile("-(\d\d)-")
def preprocess_time(row):
day = int(reg_day.search(row.req_time).group(1))
month = int(reg_month.search(row.req_time).group(1))
if month > firstmonth:
day += 30*(month - firstmonth)
return day
firstmonth = 10
df_queries['req_time'] = df_queries.apply(preprocess_time, axis=1)
df_sorted = df_queries.sort_values('req_time')
df_sorted.sample(5)
sns.countplot(df_queries.req_time)
# #### Queries per day of the week
#
# Right now we have queries per day, let's get some number for average queries every day of the week
queries_day = df_queries.groupby(['req_time']).size()
# +
queries_weekday = [0]*7
num_queries_weekday = [0]*7
for i, v in queries_day.items():
queries_weekday[(i-1) % 7] = v
num_queries_weekday[(i-1) % 7] += 1
for i in range(7):
queries_weekday[i] //= num_queries_weekday[i]
queries_weekday
# -
plt.plot(queries_weekday)
# #### Statistics of df_queries
#
# More info in `reports/EDA.md`
df_queries.describe()
# #### User analysis
#
# Analyze queries made by each user
df_most_pop_users = df_queries['pid'].value_counts()
# there are 46k users
df_most_pop_users.shape
df_most_pop_users.describe()
df_most_pop_users.head(50).plot(kind='bar',x='user IDs',y='# queries',color='red')
plt.show()
# number of profiles with one or two queries
df_most_pop_users[df_most_pop_users < 5].count()
# ### Clicks DF
df_clicks = pd.read_csv("../data/raw/data_set_phase1/train_clicks.csv")
df_clicks.isnull().sum()
df_clicks.shape
df_clicks.sample(5)
sns.countplot(df_clicks.click_mode)
# ### Plans DF
df_plans = pd.read_csv("../data/raw/data_set_phase1/train_plans.csv")
df_plans.isnull().sum()
df_plans.sample(5)
df_plans.shape
|
notebooks/.ipynb_checkpoints/1_AB_EDA-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# # Choropleth Maps Exercise
#
# Welcome to the Choropleth Maps Exercise! In this exercise we will give you some simple datasets and ask you to create Choropleth Maps from them. Due to the Nature of Plotly we can't show you examples
#
# [Full Documentation Reference](https://plot.ly/python/reference/#choropleth)
#
# ## Plotly Imports
import plotly.graph_objs as go
from plotly.offline import init_notebook_mode,iplot
init_notebook_mode(connected=True)
import pandas as pd
# ** Import pandas and read the csv file: 2014_World_Power_Consumption**
df = pd.read_csv('./2014_World_Power_Consumption')
df.head()
# ** Referencing the lecture notes, create a Choropleth Plot of the Power Consumption for Countries using the data and layout dictionary. **
# +
data = {'type':'choropleth', 'locations':df['Country'],'locationmode':'country names',
'z':df['Power Consumption KWH'], 'text':df['Text']}
layout={'title':'World power consumption',
'geo':{'showframe':True, 'projection':{'type':'Mercator'}}}
# -
choromap = go.Figure(data = [data],layout = layout)
iplot(choromap,validate=False)
# ## USA Choropleth
#
# ** Import the 2012_Election_Data csv file using pandas. **
df2 = pd.read_csv('./2012_Election_Data')
# ** Check the head of the DataFrame. **
df2.head()
# ** Now create a plot that displays the Voting-Age Population (VAP) per state. If you later want to play around with other columns, make sure you consider their data type. VAP has already been transformed to a float for you. **
data = {'type':'choropleth', 'locations':df2['State Abv'],'locationmode':'USA-states',
'z':df2['% Non-citizen'], 'text':df2['% Non-citizen']}
layout={'geo':{'scope':'usa'}}
choromap = go.Figure(data = [data],layout = layout)
iplot(choromap,validate=False)
# # Great Job!
|
udemy_ml_bootcamp/Python-for-Data-Visualization/Geographical Plotting/Choropleth Maps Exercise .ipynb
|