code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to use Quandl with Python for Data Analysis
# This notebook demonstrates how to extract data from Quandl for data analysis. The example is based on United Kingdom Office of National Statistics:
# https://www.quandl.com/data/UKONS-United-Kingdom-Office-of-National-Statistics
# You must first register on Quandl website:
# https://www.quandl.com/
#
#
#
# Then find your own unique API key under the account setting.
# You must also install the quandl package for python:
#
# pip install quandl (check this out for further guidelines: https://docs.quandl.com/docs/python-installation)
import quandl
import pandas as pd
quandl.ApiConfig.api_key = 'type your unique API key here'
# This is an example of extracting a dataset:
quandl.get('UKONS/L5PA_A')
# We can also specify the date range.
quandl.get('UKONS/L5PA_A', start_date = '2010-01-01', end_date ='2020-06-30')
# So as you can see, we need to simply find the code of the dataset we are trying to get. Most datasets come with a metadata csv file, which include all the codes associated with the dataset.
# For UKONS, you can download it from here and save it into your local computer:
# https://www.quandl.com/data/UKONS-United-Kingdom-Office-of-National-Statistics/usage/export
# So we can read the metadata file using Pandas. There are 73502 datasets for UKONS.
codes = pd.read_csv('UKONS_metadata.csv', sep =',')
codes
# Let's select only the codes that are about Consumer Price Index. They are shown by 'CPI wts'.
CPI = codes[codes['name'].str.contains('CPI wts')]
CPI
# We need to add the string 'UKONS/' to each code:
CPI.code = 'UKONS/' + CPI.code
CPI
# You can rename the column if you want, though this is optional.
CPI = CPI.rename(columns={'name': 'category'})
# We are interested in two columns only.
CPI = CPI[['code','category']]
CPI
CPI.shape
# Now importing two more libraries. re is for spliting the text because we want to remove the string 'CPI wts'.
# pickle is also for saving the data.
import re
import pickle
# We can add the category column when extracting data from quandl. This is one example:
category ='CPI wts: Education, health and social protection SPECIAL AGGREGATES (Annual)'
df = quandl.get('UKONS/A9G7_A')
df['category'] = category
df
# This function, gets the data from quandl, then add the category column and also split the text based on ':'. It then dumps the extracted data into a pickle file.
def get_data(code,category):
df =quandl.get(code)
category = re.split(':',category)[1]
df['category'] = category
return df
with open('CPI_UKNONS.p', 'wb') as f:
pickle.dump(df, f)
# Here is one example. You can try it with other codes too. You will need to give the code and category to the function.
get_data('UKONS/A9G7_A','CPI wts: Education, health and social protection SPECIAL AGGREGATES (Annual)')
# Now we can concatenate all datasets using pd.concat method:
df_all = pd.concat(get_data(code, category) for code, category
in CPI.itertuples(index=False))
df_all
# Let's save the data into a pickle file:
with open('CPI_UKNONS.p', 'wb') as f:
pickle.dump(df_all, f)
with open('CPI_UKNONS.p', 'rb') as f:
CPI_Data = pickle.load(f)
CPI_Data
# The CPI dataset is now ready for analysis. Here is one simple example. Feel free to explore it further.
df_all.groupby('category').mean()
|
Quandl for DataVigo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Not completed.
import json
import requests
import csv
import pandas as pd
import os
import matplotlib.pylab as plt
import numpy as np
# %matplotlib inline
pd.options.mode.chained_assignment = None
from statsmodels.tsa.arima_model import ARIMA
import statsmodels.api as sm
import operator
from statsmodels.tsa.stattools import acf
from statsmodels.tsa.stattools import pacf
from pandas.tools.plotting import autocorrelation_plot
dateparse = lambda dates: pd.datetime.strptime(dates, '%Y-%m-%d')
indicator_data = pd.read_csv('P:\\ADS\\Final\\Indicators_Cleaned.csv',header=0,parse_dates=True,index_col='Year',date_parser=dateparse, low_memory=False)
indicator_data.head()
indicator_data.reset_index()
indicator_data.head()
argentina_df_ind = indicator_data[(indicator_data['IndicatorCode'].isin(['NY.GDP.MKTP.KD.ZG'])) & \
(indicator_data['CountryCode'] == 'AR')]
argentina_df_ind.index
argentina_df_ind
ts = argentina_df_ind['Value']
ts1 = argentina_df_ind[['Value']].copy()
ts1['Value']=ts1['Value']+20
ts1.head()
plt.plot(ts1)
from statsmodels.tsa.stattools import adfuller
def test_stationarity(timeseries):
#Determing rolling statistics
rolmean = pd.rolling_mean(timeseries, window=12)
rolstd = pd.rolling_std(timeseries, window=12)
#Plot rolling statistics:
orig = plt.plot(timeseries, color='blue',label='Original')
mean = plt.plot(rolmean, color='red', label='Rolling Mean')
std = plt.plot(rolstd, color='black', label = 'Rolling Std')
plt.legend(loc='best')
plt.title('Rolling Mean & Standard Deviation')
plt.show(block=False)
#Perform Dickey-Fuller test:
print ('Results of Dickey-Fuller Test:')
dftest = adfuller(timeseries, autolag='AIC')
dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','#Lags Used','Number of Observations Used'])
for key,value in dftest[4].items():
dfoutput['Critical Value (%s)'%key] = value
print (dfoutput)
test_stationarity(ts1.Value)
decomposition = sm.tsa.seasonal_decompose(ts1, model='additive')
fig = decomposition.plot()
plt.show()
def logTransform(df):
ts_log = np.log(df)
plt.plot(ts_log)
return ts_log
ts1_log = logTransform(ts1)
# +
#test_stationarity(ts1_log.Value)
# -
def logFirstDifference(ts1_log):
ts1_log_diff = ts1_log - ts1_log.shift()
ts1_log_diff.dropna(inplace=True)
return ts1_log_diff
ts1_log_diff = logFirstDifference(ts1_log)
test_stationarity(ts1_log_diff.Value)
def firstDifference(df):
ts_first_diff = df - df.shift()
ts_first_diff.dropna(inplace=True)
return ts_first_diff
ts1_first_diff = firstDifference(ts1)
test_stationarity(ts1_first_diff.Value)
lag_acf = acf(ts1_log_diff, nlags=10)
lag_pacf = pacf(ts1_log_diff, nlags=10, method='ols')
fig = plt.figure(figsize=(12,8))
ax1 = fig.add_subplot(211)
fig = sm.graphics.tsa.plot_acf(ts1_log_diff, lags=10, ax=ax1)
ax2 = fig.add_subplot(212)
fig = sm.graphics.tsa.plot_pacf(ts1_log_diff, lags=10, ax=ax2)
# - As seen from the graph above both ACF and PACF are geometric hence this is an ARMA model
autocorrelation_plot(ts1_log_diff)
plt.show()
plt.subplot(122)
plt.plot(lag_pacf)
plt.axhline(y=0,linestyle='--',color='gray')
plt.axhline(y=-1.96/np.sqrt(len(ts1_log_diff)),linestyle='--',color='gray')
plt.axhline(y=1.96/np.sqrt(len(ts1_log_diff)),linestyle='--',color='gray')
plt.title('Partial Autocorrelation Function')
plt.tight_layout()
aic_metric = pd.DataFrame({'Modelname':[],'AIC':[]})
aic_dict = {}
def cal_aic_metric(modelname,model):
global aic_metric
AIC = model.aic
aic_dict[modelname] = AIC
df_error = pd.DataFrame({'Modelname':[modelname],'AIC':[AIC]})
aic_metric = pd.concat([aic_metric,df_error])
return aic_metric
def AR_Model(ts):
model = ARIMA(ts, order=(1, 1, 0))
results_AR = model.fit(disp=0)
cal_aic_metric('ARIMA(ts, order=(1, 0, 0))',results_AR)
print('Lag: %s' % results_AR.k_ar)
print('Coefficients: %s' % results_AR.params)
#print(results_AR.summary())
predict_MA_HPI = np.exp(results_AR.predict(10, 10, dynamic=True))
print(predict_MA_HPI)
plt.plot(ts1_log)
plt.plot(results_AR.fittedvalues, color='red')
#print(np.exp(results_AR.fittedvalues))
print(results_AR.aic)
return results_AR
model_AR = AR_Model(ts1_log_diff)
def MA_Model(ts):
model = ARIMA(ts, order=(0,1, 1))
results_MA = model.fit(disp=0)
cal_aic_metric('ARIMA(ts, order=(2, 1, 2))',results_MA)
print('Lag: %s' % results_MA.k_ar)
print('Coefficients: %s' % results_MA.params)
print(results_MA.summary())
plt.plot(ts)
plt.plot(results_MA.fittedvalues, color='red')
return results_MA
model_MA = MA_Model(ts1_log_diff)
def Combined_Model(ts):
model = ARIMA(ts, order=(2, 1, 1))
results_ARIMA = model.fit(disp=0)
cal_aic_metric('ARIMA(ts, order=(2,1, 3))',results_ARIMA)
print('Lag: %s' % results_ARIMA.k_ar)
print('Coefficients: %s' % results_ARIMA.params)
print(results_ARIMA.summary())
plt.plot(ts)
plt.plot(results_ARIMA.fittedvalues, color='red')
return results_ARIMA
model_Combined = Combined_Model(ts1_log_diff)
best_model = min(aic_dict.items(),key=operator.itemgetter(1))[0]
print('Best Model is ', best_model)
aic_metric
#Forecast using Best Model
def forecast(model,numSteps):
#model.forecast(steps=numSteps)
output = model.forecast(steps=numSteps)[0]
output.tolist()
output = np.exp(output)
#print(output)
return normal(output)
# +
def forC(n):
output_forecast = forecast(model_Combined,57)
return output_forecast[:n]
forC(57)
# -
def FittedValues(model):
fittedVal=model.fittedvalues
PredictedVal=np.exp(fittedVal)
np.savetxt('PredictedValues.csv', PredictedVal, delimiter=",")
print('Predicted existing values are:')
return PredictedVal
# # Taking it to normal scale
def normal(predictions_ARIMA_diff):
#predictions_ARIMA_diff = pd.Series(results_ARIMA.fittedvalues, copy=True)
predictions_ARIMA_diff_cumsum = predictions_ARIMA_diff.cumsum()
predictions_ARIMA_log = pd.Series(ts1_log.ix[0], index=ts1_log.index)
#print(predictions_ARIMA_diff_cumsum.shape," ",predictions_ARIMA_log.shape)
predictions_ARIMA_log = predictions_ARIMA_log.add(predictions_ARIMA_diff_cumsum,fill_value=0)
#predictions_ARIMA = np.exp(predictions_ARIMA_log)
predictions_ARIMA_log = predictions_ARIMA_log -20
return predictions_ARIMA_log
|
Development Indicators Project/python notebooks/AR_GDP_Growth.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0 - Setup Notebook Pod
# ## 0.1 - Run in Jupyter Bash Terminal
# ```bash
# # create application-default credentials
# gcloud auth application-default login
# ```
# # 1 - Initialize SparkSession
# +
import pyspark
from pyspark.sql import SparkSession
# construct spark_jars list
spark_jars = ["https://storage.googleapis.com/hadoop-lib/gcs/gcs-connector-hadoop2-latest.jar"]
if pyspark.version.__version__[0] == "3":
spark_jars.append("https://storage.googleapis.com/spark-lib/bigquery/spark-bigquery-latest_2.12.jar")
else:
spark_jars.append("https://storage.googleapis.com/spark-lib/bigquery/spark-bigquery-latest_2.11.jar")
# create SparkSession
spark = SparkSession \
.builder \
.master("local[1]") \
.config("spark.driver.cores", "1") \
.config("spark.driver.memory", "4g") \
.config("spark.jars", ",".join(spark_jars)) \
.config("spark.sql.legacy.parquet.datetimeRebaseModeInWrite", "LEGACY") \
.config("spark.hadoop.fs.gs.impl", "com.google.cloud.hadoop.fs.gcs.GoogleHadoopFileSystem") \
.config("spark.hadoop.fs.AbstractFileSystem.gs.impl", "com.google.cloud.hadoop.fs.gcs.GoogleHadoopFS") \
.config("spark.hadoop.fs.gs.auth.service.account.enable", "true") \
.config("spark.hadoop.fs.gs.auth.service.account.json.keyfile", "/home/jovyan/.config/gcloud/application_default_credentials.json") \
.getOrCreate()
# -
# # 2 - SparkSQL
# ## 2.0 - Docs
# * https://spark.apache.org/docs/latest/sql-getting-started.html
# * https://spark.apache.org/docs/latest/api/python/pyspark.sql.html
# ## 2.1 - Write CSV
# +
# create a DataFrame
df = spark.createDataFrame(
[("aaa", 1, "!!!"),
("bbb", 2, "@@@"),
("ccc", 3, "###"),
("ddd", 4, "%%%")],
schema=["col1", "col2", "col3", ]
)
# write CSV
out_uri = f"gs://<<<MY_BUCKET>>>/example/spark_test.csv"
df.write \
.format("csv") \
.mode("overwrite") \
.option("header", "true") \
.save(out_uri)
# link to GUI
print("----------------")
print("View in GUI:")
print(f"https://console.cloud.google.com/storage/browser/${out_uri.lstrip('gs://')}/")
print("----------------")
# -
# ## 2.2 - Read CSV
# +
# read CSV
in_uri = f"gs://<<<MY_BUCKET>>>/example/spark_test.csv"
df2 = spark.read \
.format("csv") \
.option("mode", "FAILFAST") \
.option("inferSchema", "true") \
.option("header", "true") \
.load(in_uri)
# view DataFrame
df2.show()
# -
# # 3 - BigQuery
# ## 3.0 - Docs
# * https://github.com/GoogleCloudDataproc/spark-bigquery-connector
# ## 3.1 - Write to BigQuery
# +
# create a DataFrame
df3 = spark.createDataFrame(
[("aaa", 1, "!!!"),
("bbb", 2, "@@@"),
("ccc", 3, "###"),
("ddd", 4, "%%%")],
schema=["col1", "col2", "col3", ]
)
# write to BigQuery
out_project = "<<<MY_PROJECT>>>"
out_table = "<<<MY_DATABASE>>>.example__spark_notebook"
billing_project = "<<<MY_PROJECT>>>"
df3.write \
.format("bigquery") \
.mode("overwrite") \
.option("temporaryGcsBucket", "<<MY_BUCKET>>") \
.option("parentProject", billing_project) \
.option("project", out_project) \
.option("table", out_table) \
.save()
# link to GUI
print("----------------")
print("View in GUI:")
print(f"https://console.cloud.google.com/bigquery?project=${out_project}")
print("----------------")
# -
# ## 3.2 - Read from BigQuery
# +
# read from BigQuery
in_project = "<<<MY_PROJECT>>>"
in_table = "<<<MY_DATABASE>>>.example__spark_notebook"
billing_project = "<<<MY_PROJECT>>>"
df4 = spark.read \
.format("bigquery") \
.option("readDataFormat", "ARROW") \
.option("parentProject", billing_project) \
.option("project", in_project) \
.option("table", in_table) \
.load()
# view DataFrame
df4.show()
# -
# # 4 - Advanced Functions
# ## 4.1 - Write File (Hadoop Java API)
# +
def hadoop_write_file(spark: SparkSession,
fs_uri: str,
overwrite: bool,
file_data: str) -> str:
"""
Write a string as a file using the Hadoop Java API.
:param spark: a running SparkSession
:param fs_uri: the URI of the file
:param overwrite: if we should replace any existing file (error if False)
:param file_data: the string to write as the file data
:return the URI of the writen file
"""
# create py4j wrappers of java objects
hadoop = spark.sparkContext._jvm.org.apache.hadoop
java = spark.sparkContext._jvm.java
# create the FileSystem() object
conf = spark._jsc.hadoopConfiguration()
path = hadoop.fs.Path(java.net.URI(fs_uri))
fs = path.getFileSystem(conf)
# write the file
output_stream = fs.create(path, overwrite)
output_stream.writeBytes(file_data)
output_stream.close()
return fs_uri
# write file
out_uri = f"gs://<<<MY_BUCKET>>>/example/spark_test.txt"
file_data = "Hello World! " * 100
hadoop_write_file(spark=spark, fs_uri=out_uri, overwrite=True, file_data=file_data)
# link to GUI
print("----------------")
print("View in GUI:")
print(f"https://console.cloud.google.com/storage/browser/${out_project.lstrip('gs://')}")
print("----------------")
# -
# ## 4.2 - Read File (Hadoop Java API)
# +
def hadoop_read_file(spark: SparkSession,
fs_uri: str,
encoding: str = "utf-8") -> str:
"""
Read the content of a file as a string using the Hadoop Java API.
:param spark: a running SparkSession
:param fs_uri: the URI of the file
:param encoding: the file's encoding (defaults to utf-8)
:return: the content of the file (or None if the file is not present
"""
from py4j.protocol import Py4JJavaError
# create py4j wrappers of scala objects
commons = spark.sparkContext._jvm.org.apache.commons
hadoop = spark.sparkContext._jvm.org.apache.hadoop
java = spark.sparkContext._jvm.java
# create the FileSystem() object
conf = spark._jsc.hadoopConfiguration()
path = hadoop.fs.Path(java.net.URI(fs_uri))
fs = path.getFileSystem(conf)
# read file as string
try:
input_stream = fs.open(path)
file_data = commons.io.IOUtils.toString(input_stream, encoding)
input_stream.close()
return file_data
except Py4JJavaError as ex:
java_exception_class = ex.java_exception.getClass().getName()
if java_exception_class == "java.io.FileNotFoundException":
return None
else:
raise ex
# read file
in_uri = f"gs://<<<MY_BUCKET>>>/example/spark_test.txt"
file_data = hadoop_read_file(spark=spark, fs_uri=in_uri)
print("-------- File Content --------")
print(file_data)
print("------------------------------")
# -
|
jupyter/example_notebooks/.ipynb_checkpoints/spark_example_v0.2.1-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %%
import sys
import os
import time
import random
import re
import warnings
import glob
import pickle
import copy
from pathlib import Path
from tqdm import tqdm
from collections import OrderedDict
import functools
from enum import Enum, auto
import numpy as np
os.environ["CUDA_VISIBLE_DEVICES"] = '0'
os.environ["CUDA_DEVICE_ORDER"] = 'PCI_BUS_ID'
os.environ['MKL_THREADING_LAYER'] = 'GNU'
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.cuda.amp as amp
import torchvision
from torch.utils.data import Dataset, DataLoader
import nibabel as nib
import scipy
import wandb
import matplotlib.pyplot as plt
from IPython.display import display
from sklearn.model_selection import KFold
from deep_staple.metrics import dice3d, dice2d
from deep_staple.visualization import visualize_seg
from deep_staple.mindssc import mindssc
from deep_staple.CrossmodaHybridIdLoader import CrossmodaHybridIdLoader, get_crossmoda_data_load_closure
from deep_staple.MobileNet_LR_ASPP_3D import MobileNet_LRASPP_3D, MobileNet_ASPP_3D
from deep_staple.utils.torch_utils import get_batch_dice_per_class, get_batch_dice_over_all, get_2d_stack_batch_size, \
make_2d_stack_from_3d, make_3d_from_2d_stack, interpolate_sample, dilate_label_class, ensure_dense, get_module, set_module, save_model, reset_determinism
from deep_staple.utils.common_utils import DotDict, DataParamMode, LabelDisturbanceMode, in_notebook, get_script_dir
from deep_staple.utils.log_utils import get_global_idx, log_data_parameter_stats, log_class_dices
print(torch.__version__)
print(torch.backends.cudnn.version())
print(torch.cuda.get_device_name(0))
THIS_SCRIPT_DIR = get_script_dir(__file__)
print(f"Running in: {THIS_SCRIPT_DIR}")
# %%
config_dict = DotDict({
'num_folds': 3,
'only_first_fold': True,
# 'fold_override': 0,
# 'checkpoint_epx': 0,
'use_mind': False,
'epochs': 40,
'batch_size': 8,
'val_batch_size': 1,
'use_2d_normal_to': None,
'num_val_images': 20,
'atlas_count': 1,
'dataset': 'crossmoda',
'dataset_directory': Path(THIS_SCRIPT_DIR, "data/crossmoda_dataset"),
'reg_state': "acummulate_every_third_deeds_FT2_MT1",
'train_set_max_len': None,
'crop_3d_w_dim_range': (45, 95),
'crop_2d_slices_gt_num_threshold': 0,
'lr': 0.01,
'use_scheduling': True,
# Data parameter config
'data_param_mode': DataParamMode.INSTANCE_PARAMS, # DataParamMode.DISABLED
'init_inst_param': 0.0,
'lr_inst_param': 0.1,
'use_risk_regularization': True,
'use_fixed_weighting': True,
'use_ool_dp_loss': True,
# Extended config for loading pretrained data
'fixed_weight_file': None,
'fixed_weight_min_quantile': None,
'fixed_weight_min_value': None,
'override_embedding_weights': False,
'save_every': 200,
'mdl_save_prefix': 'data/models',
'debug': False,
'wandb_mode': 'disabled', # e.g. online, disabled
'do_sweep': False,
'checkpoint_name': None,
'fold_override': None,
'checkpoint_epx': None,
'do_plot': False,
'save_dp_figures': False,
'save_labels': False,
# Disturbance settings
'disturbance_mode': None, # LabelDisturbanceMode.FLIP_ROLL, LabelDisturbanceMode.AFFINE
'disturbance_strength': 0.,
'disturbed_percentage': 0.,
})
# %%
def prepare_data(config):
assert os.path.isdir(config.dataset_directory), "Dataset directory does not exist."
reset_determinism()
if config.reg_state:
print("Loading registered data.")
if config.reg_state == "mix_combined_best":
config.atlas_count = 1
domain = 'source'
label_data_left = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_left.pth"))
label_data_right = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_right.pth"))
loaded_identifier = label_data_left['valid_left_t1'] + label_data_right['valid_right_t1']
perm = np.random.permutation(len(loaded_identifier))
_clen = int(.5*len(loaded_identifier))
best_choice = perm[:_clen]
combined_choice = perm[_clen:]
best_label_data = torch.cat([label_data_left['best_all'].to_dense()[:44], label_data_right['best_all'].to_dense()[:63]], dim=0)[best_choice]
combined_label_data = torch.cat([label_data_left['combined_all'].to_dense()[:44], label_data_right['combined_all'].to_dense()[:63]], dim=0)[combined_choice]
label_data = torch.zeros([107,128,128,128])
label_data[best_choice] = best_label_data
label_data[combined_choice] = combined_label_data
var_identifier = ["mBST" if idx in best_choice else "mCMB" for idx in range(len(loaded_identifier))]
loaded_identifier = [f"{_id}:{var_id}" for _id, var_id in zip(loaded_identifier, var_identifier)]
elif config.reg_state == "acummulate_combined_best":
config.atlas_count = 2
domain = 'source'
label_data_left = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_left.pth"))
label_data_right = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_right.pth"))
loaded_identifier = label_data_left['valid_left_t1'] + label_data_right['valid_right_t1']
best_label_data = torch.cat([label_data_left['best_all'].to_dense()[:44], label_data_right['best_all'].to_dense()[:63]], dim=0)
combined_label_data = torch.cat([label_data_left['combined_all'].to_dense()[:44], label_data_right['combined_all'].to_dense()[:63]], dim=0)
label_data = torch.cat([best_label_data, combined_label_data])
loaded_identifier = [_id+':mBST' for _id in loaded_identifier] + [_id+':mCMB' for _id in loaded_identifier]
elif config.reg_state == "best":
config.atlas_count = 1
domain = 'source'
label_data_left = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_left.pth"))
label_data_right = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_right.pth"))
loaded_identifier = label_data_left['valid_left_t1'] + label_data_right['valid_right_t1']
label_data = torch.cat([label_data_left[config.reg_state+'_all'].to_dense()[:44], label_data_right[config.reg_state+'_all'].to_dense()[:63]], dim=0)
postfix = 'mBST'
loaded_identifier = [_id+':'+postfix for _id in loaded_identifier]
elif config.reg_state == "combined":
config.atlas_count = 1
domain = 'source'
label_data_left = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_left.pth"))
label_data_right = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220113_crossmoda_optimal/optimal_reg_right.pth"))
loaded_identifier = label_data_left['valid_left_t1'] + label_data_right['valid_right_t1']
label_data = torch.cat([label_data_left[config.reg_state+'_all'].to_dense()[:44], label_data_right[config.reg_state+'_all'].to_dense()[:63]], dim=0)
postfix = 'mCMB'
loaded_identifier = [_id+':'+postfix for _id in loaded_identifier]
elif config.reg_state == "acummulate_convex_adam_FT2_MT1":
config.atlas_count = 10
domain = 'target'
bare_data = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220318_crossmoda_convex_adam_lr/crossmoda_convex_registered_new_convex.pth"))
label_data = []
loaded_identifier = []
for fixed_id, moving_dict in bare_data.items():
sorted_moving_dict = OrderedDict(moving_dict)
for idx_mov, (moving_id, moving_sample) in enumerate(sorted_moving_dict.items()):
# Only use every third warped sample
if idx_mov % 3 == 0:
label_data.append(moving_sample['warped_label'].cpu())
loaded_identifier.append(f"{fixed_id}:m{moving_id}")
elif config.reg_state == "acummulate_every_third_deeds_FT2_MT1":
config.atlas_count = 10
domain = 'target'
bare_data = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220114_crossmoda_multiple_registrations/crossmoda_deeds_registered.pth"))
label_data = []
loaded_identifier = []
for fixed_id, moving_dict in bare_data.items():
sorted_moving_dict = OrderedDict(moving_dict)
for idx_mov, (moving_id, moving_sample) in enumerate(sorted_moving_dict.items()):
# Only use every third warped sample
if idx_mov % 3 == 0:
label_data.append(moving_sample['warped_label'].cpu())
loaded_identifier.append(f"{fixed_id}:m{moving_id}")
elif config.reg_state == "acummulate_every_deeds_FT2_MT1":
config.atlas_count = 30
domain = 'target'
bare_data = torch.load(Path(THIS_SCRIPT_DIR, "./data_artifacts/20220114_crossmoda_multiple_registrations/crossmoda_deeds_registered.pth"))
label_data = []
loaded_identifier = []
for fixed_id, moving_dict in bare_data.items():
sorted_moving_dict = OrderedDict(moving_dict)
for idx_mov, (moving_id, moving_sample) in enumerate(sorted_moving_dict.items()):
label_data.append(moving_sample['warped_label'].cpu())
loaded_identifier.append(f"{fixed_id}:m{moving_id}")
else:
raise ValueError()
modified_3d_label_override = {}
for idx, identifier in enumerate(loaded_identifier):
# Find sth. like 100r:mBST or 100r:m001l
nl_id, lr_id, m_id = re.findall(r'(\d{1,3})([lr]):m([A-Z0-9a-z]{3,4})$', identifier)[0]
nl_id = int(nl_id)
crossmoda_var_id = f"{nl_id:03d}{lr_id}:m{m_id}"
modified_3d_label_override[crossmoda_var_id] = label_data[idx]
prevent_disturbance = True
else:
domain = 'source'
modified_3d_label_override = None
prevent_disturbance = False
if config.dataset == 'crossmoda':
# Use double size in 2D prediction, normal size in 3D
pre_interpolation_factor = 2. if config.use_2d_normal_to is not None else 1.5
clsre = get_crossmoda_data_load_closure(
base_dir=str(config.dataset_directory),
domain=domain, state='l4', use_additional_data=False,
size=(128,128,128), resample=True, normalize=True, crop_3d_w_dim_range=config.crop_3d_w_dim_range,
ensure_labeled_pairs=True, modified_3d_label_override=modified_3d_label_override,
debug=config.debug
)
training_dataset = CrossmodaHybridIdLoader(
clsre,
size=(128,128,128), resample=True, normalize=True, crop_3d_w_dim_range=config.crop_3d_w_dim_range,
ensure_labeled_pairs=True,
max_load_3d_num=config.train_set_max_len,
prevent_disturbance=prevent_disturbance,
use_2d_normal_to=config.use_2d_normal_to,
crop_2d_slices_gt_num_threshold=config.crop_2d_slices_gt_num_threshold,
pre_interpolation_factor=pre_interpolation_factor,
fixed_weight_file=config.fixed_weight_file, fixed_weight_min_quantile=config.fixed_weight_min_quantile, fixed_weight_min_value=config.fixed_weight_min_value,
)
return training_dataset
# %%
if config_dict['do_plot'] and False:
# Plot label voxel W-dim distribution
training_dataset = prepare_data(config_dict)
_, all_labels, _ = training_dataset.get_data(use_2d_override=False)
print(all_labels.shape)
sum_over_w = torch.sum(all_labels, dim=(0,1,2))
plt.xlabel("W")
plt.ylabel("ground truth>0")
plt.plot(sum_over_w);
# %%
def save_parameter_figure(_path, title, text, parameters, reweighted_parameters, dices):
# Show weights and weights with compensation
fig, axs = plt.subplots(1,2, figsize=(12, 4), dpi=80)
sc1 = axs[0].scatter(
range(len(parameters)),
parameters.cpu().detach(), c=dices,s=1, cmap='plasma', vmin=0., vmax=1.)
sc2 = axs[1].scatter(
range(len(reweighted_parameters)),
reweighted_parameters.cpu().detach(), s=1,c=dices, cmap='plasma', vmin=0., vmax=1.)
fig.suptitle(title, fontsize=14)
fig.text(0, 0, text)
axs[0].set_title('Bare parameters')
axs[1].set_title('Reweighted parameters')
axs[0].set_ylim(-10, 10)
axs[1].set_ylim(-3, 1)
plt.colorbar(sc2)
plt.savefig(_path)
plt.clf()
plt.close()
def calc_inst_parameters_in_target_pos_ratio(dpm, disturbed_inst_idxs, target_pos='min'):
assert target_pos == 'min' or target_pos == 'max', "Value of target_pos must be 'min' or 'max'."
descending = False if target_pos == 'min' else True
target_len = len(disturbed_inst_idxs)
disturbed_params = dpm.get_parameter_list(inst_keys=disturbed_inst_idxs)
all_params = sorted(dpm.get_parameter_list(inst_keys='all'), reverse=descending)
target_param_ids = [id(param) for param in all_params[:target_len]]
ratio = [1. for param in disturbed_params if id(param) in target_param_ids]
ratio = sum(ratio)/target_len
return ratio
# %%
# %%
if config_dict['do_plot']:
training_dataset = prepare_data(config_dict)
# Print transformed 2D data
training_dataset.train(use_modified=True, augment=False)
# print(training_dataset.disturbed_idxs)
print("Displaying 2D training sample")
img_stack = []
label_stack = []
mod_label_stack = []
for sample in (training_dataset[idx] for idx in [500,590]):
print(sample['id'])
img_stack.append(sample['image'])
label_stack.append(sample['label'])
mod_label_stack.append(sample['modified_label'])
# Change label num == hue shift for display
img_stack = torch.stack(img_stack).unsqueeze(1)
label_stack = torch.stack(label_stack)
mod_label_stack = torch.stack(mod_label_stack)
mod_label_stack*=4
visualize_seg(in_type="batch_3D", reduce_dim="W",
img=img_stack,
# ground_truth=label_stack,
seg=(mod_label_stack-label_stack).abs(),
# crop_to_non_zero_gt=True,
crop_to_non_zero_seg=True,
alpha_seg = .5
)
def get_model(config, dataset_len, num_classes, THIS_SCRIPT_DIR, _path=None, device='cpu'):
_path = Path(THIS_SCRIPT_DIR).joinpath(_path).resolve()
if config.use_mind:
in_channels = 12
else:
in_channels = 1
if config.use_2d_normal_to is not None:
# Use vanilla torch model
lraspp = torchvision.models.segmentation.lraspp_mobilenet_v3_large(
pretrained=False, progress=True, num_classes=num_classes
)
set_module(lraspp, 'backbone.0.0',
torch.nn.Conv2d(in_channels, 16, kernel_size=(3, 3), stride=(2, 2),
padding=(1, 1), bias=False)
)
else:
# Use custom 3d model
lraspp = MobileNet_LRASPP_3D(
in_num=in_channels, num_classes=num_classes,
use_checkpointing=True
)
# lraspp.register_parameter('sigmoid_offset', nn.Parameter(torch.tensor([0.])))
lraspp.to(device)
print(f"Param count lraspp: {sum(p.numel() for p in lraspp.parameters())}")
optimizer = torch.optim.AdamW(lraspp.parameters(), lr=config.lr)
scaler = amp.GradScaler()
if config.use_2d_normal_to is not None:
scheduler = torch.optim.lr_scheduler.CosineAnnealingWarmRestarts(
optimizer, T_0=10, T_mult=2)
else:
# Use ExponentialLR in 3D
scheduler = torch.optim.lr_scheduler.ExponentialLR(optimizer, gamma=.99)
# Add data paramters embedding and optimizer
if config.data_param_mode == str(DataParamMode.INSTANCE_PARAMS):
embedding = nn.Embedding(dataset_len, 1, sparse=True)
embedding = embedding.to(device)
# Init embedding values
#
if config.override_embedding_weights:
fixed_weightdata = torch.load(config.fixed_weight_file)
fixed_weights = fixed_weightdata['data_parameters']
fixed_d_ids = fixed_weightdata['d_ids']
if config.use_2d_normal_to is not None:
corresp_dataset_idxs = [training_dataset.get_2d_ids().index(_id) for _id in fixed_d_ids]
else:
corresp_dataset_idxs = [training_dataset.get_3d_ids().index(_id) for _id in fixed_d_ids]
embedding_weight_tensor = torch.zeros_like(embedding.weight)
embedding_weight_tensor[corresp_dataset_idxs] = fixed_weights.view(-1,1).cuda()
embedding = nn.Embedding(len(training_dataset), 1, sparse=True, _weight=embedding_weight_tensor)
elif _path and _path.is_dir():
embedding.load_state_dict(torch.load(_path.joinpath('embedding.pth'), map_location=device))
else:
torch.nn.init.normal_(embedding.weight.data, mean=config.init_inst_param, std=0.00)
print(f"Param count embedding: {sum(p.numel() for p in embedding.parameters())}")
optimizer_dp = torch.optim.SparseAdam(
embedding.parameters(), lr=config.lr_inst_param,
betas=(0.9, 0.999), eps=1e-08)
scaler_dp = amp.GradScaler()
if _path and _path.is_dir():
print(f"Loading dp_optimizer and scaler_dp from {_path}")
optimizer_dp.load_state_dict(torch.load(_path.joinpath('optimizer_dp.pth'), map_location=device))
scaler_dp.load_state_dict(torch.load(_path.joinpath('scaler_dp.pth'), map_location=device))
else:
embedding = None
optimizer_dp = None
scaler_dp = None
if _path and _path.is_dir():
print(f"Loading lr-aspp model, optimizers and grad scalers from {_path}")
lraspp.load_state_dict(torch.load(_path.joinpath('lraspp.pth'), map_location=device))
optimizer.load_state_dict(torch.load(_path.joinpath('optimizer.pth'), map_location=device))
scheduler.load_state_dict(torch.load(_path.joinpath('scheduler.pth'), map_location=device))
scaler.load_state_dict(torch.load(_path.joinpath('scaler.pth'), map_location=device))
else:
print("Generating fresh lr-aspp model, optimizer and grad scaler.")
return (lraspp, optimizer, scheduler, optimizer_dp, embedding, scaler, scaler_dp)
# %%
def inference_wrap(lraspp, img, use_2d, use_mind):
with torch.inference_mode():
b_img = img.unsqueeze(0).unsqueeze(0).float()
if use_2d and use_mind:
# MIND 2D, in Bx1x1xHxW, out BxMINDxHxW
b_img = mindssc(b_img.unsqueeze(0)).squeeze(2)
elif not use_2d and use_mind:
# MIND 3D in Bx1xDxHxW out BxMINDxDxHxW
b_img = mindssc(b_img)
elif use_2d or not use_2d:
# 2D Bx1xHxW
# 3D out Bx1xDxHxW
pass
b_out = lraspp(b_img)['out']
b_out = b_out.argmax(1)
return b_out
def train_DL(run_name, config, training_dataset):
reset_determinism()
# Configure folds
kf = KFold(n_splits=config.num_folds)
# kf.get_n_splits(training_dataset.__len__(use_2d_override=False))
fold_iter = enumerate(kf.split(range(training_dataset.__len__(use_2d_override=False))))
if config.get('fold_override', None):
selected_fold = config.get('fold_override', 0)
fold_iter = list(fold_iter)[selected_fold:selected_fold+1]
elif config.only_first_fold:
fold_iter = list(fold_iter)[0:1]
if config.wandb_mode != 'disabled':
warnings.warn("Logging of dataset file paths is disabled.")
# # Log dataset info
# training_dataset.eval()
# dataset_info = [[smp['dataset_idx'], smp['id'], smp['image_path'], smp['label_path']] \
# for smp in training_dataset]
# wandb.log({'datasets/training_dataset':wandb.Table(columns=['dataset_idx', 'id', 'image', 'label'], data=dataset_info)}, step=0)
if config.use_2d_normal_to is not None:
n_dims = (-2,-1)
else:
n_dims = (-3,-2,-1)
fold_means_no_bg = []
for fold_idx, (train_idxs, val_idxs) in fold_iter:
train_idxs = torch.tensor(train_idxs)
val_idxs = torch.tensor(val_idxs)
all_3d_ids = training_dataset.get_3d_ids()
if config.debug:
num_val_images = 2
atlas_count = 1
else:
num_val_images = config.num_val_images
atlas_count = config.atlas_count
if config.use_2d_normal_to is not None:
# Override idxs
all_3d_ids = training_dataset.get_3d_ids()
val_3d_idxs = torch.tensor(list(range(0, num_val_images*atlas_count, atlas_count)))
val_3d_ids = training_dataset.switch_3d_identifiers(val_3d_idxs)
train_3d_idxs = list(range(num_val_images*atlas_count, len(all_3d_ids)))
# Get corresponding 2D idxs
train_2d_ids = []
dcts = training_dataset.get_id_dicts()
for id_dict in dcts:
_2d_id = id_dict['2d_id']
_3d_idx = id_dict['3d_dataset_idx']
if _2d_id in training_dataset.label_data_2d.keys() and _3d_idx in train_3d_idxs:
train_2d_ids.append(_2d_id)
train_2d_idxs = training_dataset.switch_2d_identifiers(train_2d_ids)
train_idxs = torch.tensor(train_2d_idxs)
else:
val_3d_idxs = torch.tensor(list(range(0, num_val_images*atlas_count, atlas_count)))
val_3d_ids = training_dataset.switch_3d_identifiers(val_3d_idxs)
train_3d_idxs = list(range(num_val_images*atlas_count, len(all_3d_ids)))
train_idxs = torch.tensor(train_3d_idxs)
print(f"Will run validation with these 3D samples (#{len(val_3d_ids)}):", sorted(val_3d_ids))
_, _, all_modified_segs = training_dataset.get_data()
if config.disturbed_percentage > 0.:
with torch.no_grad():
non_empty_train_idxs = [(all_modified_segs[train_idxs].sum(dim=n_dims) > 0)]
### Disturb dataset (only non-emtpy idxs)###
proposed_disturbed_idxs = np.random.choice(non_empty_train_idxs, size=int(len(non_empty_train_idxs)*config.disturbed_percentage), replace=False)
proposed_disturbed_idxs = torch.tensor(proposed_disturbed_idxs)
training_dataset.disturb_idxs(proposed_disturbed_idxs,
disturbance_mode=config.disturbance_mode,
disturbance_strength=config.disturbance_strength
)
disturbed_bool_vect = torch.zeros(len(training_dataset))
disturbed_bool_vect[training_dataset.disturbed_idxs] = 1.
else:
disturbed_bool_vect = torch.zeros(len(training_dataset))
clean_idxs = train_idxs[np.isin(train_idxs, training_dataset.disturbed_idxs, invert=True)]
print("Disturbed indexes:", sorted(training_dataset.disturbed_idxs))
if clean_idxs.numel() < 200:
print(f"Clean indexes: {sorted(clean_idxs.tolist())}")
wandb.log({f'datasets/disturbed_idxs_fold{fold_idx}':wandb.Table(columns=['train_idxs'], data=[[idx] for idx in training_dataset.disturbed_idxs])},
step=get_global_idx(fold_idx, 0, config.epochs))
### Configure MIND ###
if config.use_mind:
in_channels = 12
else:
in_channels = 1
### Add train sampler and dataloaders ##
train_subsampler = torch.utils.data.SubsetRandomSampler(train_idxs)
# val_subsampler = torch.utils.data.SubsetRandomSampler(val_idxs)
train_dataloader = DataLoader(training_dataset, batch_size=config.batch_size,
sampler=train_subsampler, pin_memory=False, drop_last=False,
# collate_fn=training_dataset.get_efficient_augmentation_collate_fn()
)
# training_dataset.set_augment_at_collate(True) # This function does not work as expected. Scores get worse.
### Get model, data parameters, optimizers for model and data parameters, as well as grad scaler ###
if 'checkpoint_epx' in config and config['checkpoint_epx'] is not None:
epx_start = config['checkpoint_epx']
else:
epx_start = 0
if config.checkpoint_name:
# Load from checkpoint
_path = f"{config.mdl_save_prefix}/{config.checkpoint_name}_fold{fold_idx}_epx{epx_start}"
else:
_path = f"{config.mdl_save_prefix}/{wandb.run.name}_fold{fold_idx}_epx{epx_start}"
(lraspp, optimizer, scheduler, optimizer_dp, embedding, scaler, scaler_dp) = get_model(config, len(training_dataset), len(training_dataset.label_tags),
THIS_SCRIPT_DIR=THIS_SCRIPT_DIR, _path=_path, device='cuda')
t_start = time.time()
dice_func = dice2d if config.use_2d_normal_to is not None else dice3d
bn_count = torch.zeros([len(training_dataset.label_tags)], device=all_modified_segs.device)
wise_dice = torch.zeros([len(training_dataset), len(training_dataset.label_tags)])
gt_num = torch.zeros([len(training_dataset)])
with torch.no_grad():
print("Fetching training metrics for samples.")
# _, wise_lbls, mod_lbls = training_dataset.get_data()
training_dataset.eval(use_modified=True)
for sample in tqdm((training_dataset[idx] for idx in train_idxs), desc="metric:", total=len(train_idxs)):
d_idxs = sample['dataset_idx']
wise_label, mod_label = sample['label'], sample['modified_label']
mod_label = mod_label.cuda()
wise_label = wise_label.cuda()
mod_label, _ = ensure_dense(mod_label)
dsc = dice_func(
torch.nn.functional.one_hot(wise_label.unsqueeze(0), len(training_dataset.label_tags)),
torch.nn.functional.one_hot(mod_label.unsqueeze(0), len(training_dataset.label_tags)),
one_hot_torch_style=True, nan_for_unlabeled_target=False
)
bn_count += torch.bincount(mod_label.reshape(-1).long(), minlength=len(training_dataset.label_tags)).cpu()
wise_dice[d_idxs] = dsc.cpu()
gt_num[d_idxs] = (mod_label > 0).sum(dim=n_dims).float().cpu()
class_weights = 1 / (bn_count).float().pow(.35)
class_weights /= class_weights.mean()
fixed_weighting = (gt_num+np.exp(1)).log()+np.exp(1)
class_weights = class_weights.cuda()
fixed_weighting = fixed_weighting.cuda()
for epx in range(epx_start, config.epochs):
global_idx = get_global_idx(fold_idx, epx, config.epochs)
lraspp.train()
### Disturb samples ###
training_dataset.train(use_modified=True)
epx_losses = []
dices = []
class_dices = []
# Load data
for batch_idx, batch in tqdm(enumerate(train_dataloader), desc="batch:", total=len(train_dataloader)):
optimizer.zero_grad()
if optimizer_dp:
optimizer_dp.zero_grad()
b_img = batch['image']
b_seg = batch['label']
b_seg_modified = batch['modified_label']
b_idxs_dataset = batch['dataset_idx']
b_img = b_img.float()
b_img = b_img.cuda()
b_seg_modified = b_seg_modified.cuda()
b_idxs_dataset = b_idxs_dataset.cuda()
b_seg = b_seg.cuda()
if training_dataset.use_2d() and config.use_mind:
# MIND 2D, in Bx1x1xHxW, out BxMINDxHxW
b_img = mindssc(b_img.unsqueeze(1).unsqueeze(1)).squeeze(2)
elif not training_dataset.use_2d() and config.use_mind:
# MIND 3D
b_img = mindssc(b_img.unsqueeze(1))
else:
b_img = b_img.unsqueeze(1)
### Forward pass ###
with amp.autocast(enabled=True):
assert b_img.dim() == len(n_dims)+2, \
f"Input image for model must be {len(n_dims)+2}D: BxCxSPATIAL but is {b_img.shape}"
for param in lraspp.parameters():
param.requires_grad = True
lraspp.use_checkpointing = True
logits = lraspp(b_img)['out']
### Calculate loss ###
assert logits.dim() == len(n_dims)+2, \
f"Input shape for loss must be BxNUM_CLASSESxSPATIAL but is {logits.shape}"
assert b_seg_modified.dim() == len(n_dims)+1, \
f"Target shape for loss must be BxSPATIAL but is {b_seg_modified.shape}"
ce_loss = nn.CrossEntropyLoss(class_weights)(logits, b_seg_modified)
if config.data_param_mode == str(DataParamMode.DISABLED) or config.use_ool_dp_loss:
scaler.scale(ce_loss).backward()
scaler.step(optimizer)
scaler.update()
if config.data_param_mode == str(DataParamMode.INSTANCE_PARAMS):
if config.use_ool_dp_loss:
# Run second consecutive forward pass
for param in lraspp.parameters():
param.requires_grad = False
lraspp.use_checkpointing = False
dp_logits = lraspp(b_img)['out']
else:
# Do not run a second forward pass
for param in lraspp.parameters():
param.requires_grad = True
lraspp.use_checkpointing = True
dp_logits = logits
dp_loss = nn.CrossEntropyLoss(reduction='none')(dp_logits, b_seg_modified)
dp_loss = dp_loss.mean(n_dims)
bare_weight = embedding(b_idxs_dataset).squeeze()
weight = torch.sigmoid(bare_weight)
weight = weight/weight.mean()
# This improves scores significantly: Reweight with log(gt_numel)
if config.use_fixed_weighting:
weight = weight/fixed_weighting[b_idxs_dataset]
if config.use_risk_regularization:
p_pred_num = (dp_logits.argmax(1) > 0).sum(dim=n_dims).detach()
if config.use_2d_normal_to is not None:
risk_regularization = -weight*p_pred_num/(dp_logits.shape[-2]*dp_logits.shape[-1])
else:
risk_regularization = -weight*p_pred_num/(dp_logits.shape[-3]*dp_logits.shape[-2]*dp_logits.shape[-1])
dp_loss = (dp_loss*weight).sum() + risk_regularization.sum()
else:
dp_loss = (dp_loss*weight).sum()
if str(config.data_param_mode) != str(DataParamMode.DISABLED):
scaler_dp.scale(dp_loss).backward()
if config.use_ool_dp_loss:
# LRASPP already stepped.
if not config.override_embedding_weights:
scaler_dp.step(optimizer_dp)
scaler_dp.update()
else:
scaler_dp.step(optimizer)
if not config.override_embedding_weights:
scaler_dp.step(optimizer_dp)
scaler_dp.update()
epx_losses.append(dp_loss.item())
else:
epx_losses.append(ce_loss.item())
logits_for_score = logits.argmax(1)
# Calculate dice score
b_dice = dice_func(
torch.nn.functional.one_hot(logits_for_score, len(training_dataset.label_tags)),
torch.nn.functional.one_hot(b_seg, len(training_dataset.label_tags)), # Calculate dice score with original segmentation (no disturbance)
one_hot_torch_style=True
)
dices.append(get_batch_dice_over_all(
b_dice, exclude_bg=True))
class_dices.append(get_batch_dice_per_class(
b_dice, training_dataset.label_tags, exclude_bg=True))
### Scheduler management ###
if config.use_scheduling and epx % atlas_count == 0:
scheduler.step()
if str(config.data_param_mode) != str(DataParamMode.DISABLED) and batch_idx % 10 == 0 and config.save_dp_figures:
# Output data parameter figure
train_params = embedding.weight[train_idxs].squeeze()
# order = np.argsort(train_params.cpu().detach()) # Order by DP value
order = torch.arange(len(train_params))
pearson_corr_coeff = np.corrcoef(train_params.cpu().detach(), wise_dice[train_idxs][:,1].cpu().detach())[0,1]
dp_figure_path = Path(f"data/output_figures/{wandb.run.name}_fold{fold_idx}/dp_figure_epx{epx:03d}_batch{batch_idx:03d}.png")
dp_figure_path.parent.mkdir(parents=True, exist_ok=True)
save_parameter_figure(dp_figure_path, wandb.run.name, f"corr. coeff. DP vs. dice(expert label, train gt): {pearson_corr_coeff:4f}",
train_params[order], train_params[order]/fixed_weighting[train_idxs][order], dices=wise_dice[train_idxs][:,1][order])
if config.debug:
break
### Logging ###
print(f"### Log epoch {epx} @ {time.time()-t_start:.2f}s")
print("### Training")
### Log wandb data ###
# Log the epoch idx per fold - so we can recover the diagram by setting
# ref_epoch_idx as x-axis in wandb interface
wandb.log({"ref_epoch_idx": epx}, step=global_idx)
mean_loss = torch.tensor(epx_losses).mean()
wandb.log({f'losses/loss_fold{fold_idx}': mean_loss}, step=global_idx)
mean_dice = np.nanmean(dices)
print(f'dice_mean_wo_bg_fold{fold_idx}', f"{mean_dice*100:.2f}%")
wandb.log({f'scores/dice_mean_wo_bg_fold{fold_idx}': mean_dice}, step=global_idx)
log_class_dices("scores/dice_mean_", f"_fold{fold_idx}", class_dices, global_idx)
# Log data parameters of disturbed samples
if str(config.data_param_mode) != str(DataParamMode.DISABLED):
# Calculate dice score corr coeff (unknown to network)
train_params = embedding.weight[train_idxs].squeeze()
order = np.argsort(train_params.cpu().detach())
pearson_corr_coeff = np.corrcoef(train_params[order].cpu().detach(), wise_dice[train_idxs][:,1][order].cpu().detach())[0,1]
spearman_corr_coeff, spearman_p = scipy.stats.spearmanr(train_params[order].cpu().detach(), wise_dice[train_idxs][:,1][order].cpu().detach())
wandb.log(
{f'data_parameters/pearson_corr_coeff_fold{fold_idx}': pearson_corr_coeff},
step=global_idx
)
wandb.log(
{f'data_parameters/spearman_corr_coeff_fold{fold_idx}': spearman_corr_coeff},
step=global_idx
)
wandb.log(
{f'data_parameters/spearman_p_fold{fold_idx}': spearman_p},
step=global_idx
)
print(f'data_parameters/pearson_corr_coeff_fold{fold_idx}', f"{pearson_corr_coeff:.2f}")
print(f'data_parameters/spearman_corr_coeff_fold{fold_idx}', f"{spearman_corr_coeff:.2f}")
print(f'data_parameters/spearman_p_fold{fold_idx}', f"{spearman_p:.5f}")
# Log stats of data parameters and figure
log_data_parameter_stats(f'data_parameters/iter_stats_fold{fold_idx}', global_idx, embedding.weight.data)
if (epx % config.save_every == 0 and epx != 0) \
or (epx+1 == config.epochs):
_path = f"{config.mdl_save_prefix}/{wandb.run.name}_fold{fold_idx}_epx{epx}"
save_model(
_path,
lraspp=lraspp,
optimizer=optimizer, optimizer_dp=optimizer_dp,
scheduler=scheduler,
embedding=embedding,
scaler=scaler,
scaler_dp=scaler_dp)
(lraspp, optimizer, optimizer_dp, embedding, scaler) = \
get_model(
config, len(training_dataset),
len(training_dataset.label_tags),
THIS_SCRIPT_DIR=THIS_SCRIPT_DIR,
_path=_path, device='cuda')
print()
print("### Validation")
lraspp.eval()
training_dataset.eval()
val_dices = []
val_class_dices = []
with amp.autocast(enabled=True):
with torch.no_grad():
for val_idx in val_3d_idxs:
val_sample = training_dataset.get_3d_item(val_idx)
stack_dim = training_dataset.use_2d_normal_to
# Create batch out of single val sample
b_val_img = val_sample['image'].unsqueeze(0)
b_val_seg = val_sample['label'].unsqueeze(0)
B = b_val_img.shape[0]
b_val_img = b_val_img.unsqueeze(1).float().cuda()
b_val_seg = b_val_seg.cuda()
if training_dataset.use_2d():
b_val_img_2d = make_2d_stack_from_3d(b_val_img, stack_dim=training_dataset.use_2d_normal_to)
if config.use_mind:
# MIND 2D model, in Bx1x1xHxW, out BxMINDxHxW
b_val_img_2d = mindssc(b_val_img_2d.unsqueeze(1)).squeeze(2)
output_val = lraspp(b_val_img_2d)['out']
val_logits_for_score = output_val.argmax(1)
# Prepare logits for scoring
# Scoring happens in 3D again - unstack batch tensor again to stack of 3D
val_logits_for_score = make_3d_from_2d_stack(
val_logits_for_score.unsqueeze(1), stack_dim, B
).squeeze(1)
else:
if config.use_mind:
# MIND 3D model shape BxMINDxDxHxW
b_val_img = mindssc(b_val_img)
else:
# 3D model shape Bx1xDxHxW
pass
output_val = lraspp(b_val_img)['out']
val_logits_for_score = output_val.argmax(1)
b_val_dice = dice3d(
torch.nn.functional.one_hot(val_logits_for_score, len(training_dataset.label_tags)),
torch.nn.functional.one_hot(b_val_seg, len(training_dataset.label_tags)),
one_hot_torch_style=True
)
# Get mean score over batch
val_dices.append(get_batch_dice_over_all(
b_val_dice, exclude_bg=True))
val_class_dices.append(get_batch_dice_per_class(
b_val_dice, training_dataset.label_tags, exclude_bg=True))
if config.do_plot:
print(f"Validation 3D image label/ground-truth {val_3d_idxs}")
print(get_batch_dice_over_all(
b_val_dice, exclude_bg=False))
# display_all_seg_slices(b_seg.unsqueeze(1), logits_for_score)
display_seg(in_type="single_3D",
reduce_dim="W",
img=val_sample['image'].unsqueeze(0).cpu(),
seg=val_logits_for_score_3d.squeeze(0).cpu(),
ground_truth=b_val_seg.squeeze(0).cpu(),
crop_to_non_zero_seg=True,
crop_to_non_zero_gt=True,
alpha_seg=.3,
alpha_gt=.0
)
mean_val_dice = np.nanmean(val_dices)
print(f'val_dice_mean_wo_bg_fold{fold_idx}', f"{mean_val_dice*100:.2f}%")
wandb.log({f'scores/val_dice_mean_wo_bg_fold{fold_idx}': mean_val_dice}, step=global_idx)
log_class_dices("scores/val_dice_mean_", f"_fold{fold_idx}", val_class_dices, global_idx)
print()
# End of training loop
if config.debug:
break
if str(config.data_param_mode) == str(DataParamMode.INSTANCE_PARAMS):
# Write sample data
save_dict = {}
training_dataset.eval(use_modified=True)
all_idxs = torch.tensor(range(len(training_dataset))).cuda()
train_label_snapshot_path = Path(THIS_SCRIPT_DIR).joinpath(f"data/output/{wandb.run.name}_fold{fold_idx}_epx{epx}/train_label_snapshot.pth")
seg_viz_out_path = Path(THIS_SCRIPT_DIR).joinpath(f"data/output/{wandb.run.name}_fold{fold_idx}_epx{epx}/data_parameter_weighted_samples.png")
train_label_snapshot_path.parent.mkdir(parents=True, exist_ok=True)
dp_weights = embedding(all_idxs)
save_data = []
data_generator = zip(
dp_weights[train_idxs], \
disturbed_bool_vect[train_idxs],
torch.utils.data.Subset(training_dataset, train_idxs)
)
for dp_weight, disturb_flg, sample in data_generator:
data_tuple = ( \
dp_weight,
bool(disturb_flg.item()),
sample['id'],
sample['dataset_idx'],
# sample['image'],
sample['label'].to_sparse(),
sample['modified_label'].to_sparse(),
inference_wrap(lraspp, sample['image'].cuda(), use_2d=training_dataset.use_2d(), use_mind=config.use_mind).to_sparse()
)
save_data.append(data_tuple)
save_data = sorted(save_data, key=lambda tpl: tpl[0])
(dp_weight, disturb_flags,
d_ids, dataset_idxs,
# _imgs,
_labels, _modified_labels, _predictions) = zip(*save_data)
dp_weight = torch.stack(dp_weight)
dataset_idxs = torch.stack(dataset_idxs)
save_dict.update(
{
'data_parameters': dp_weight.cpu(),
'disturb_flags': disturb_flags,
'd_ids': d_ids,
'dataset_idxs': dataset_idxs.cpu(),
}
)
if config.save_labels:
_labels = torch.stack(_labels)
_modified_labels = torch.stack(_modified_labels)
_predictions = torch.stack(_predictions)
save_dict.update(
{
'labels': _labels.cpu(),
'modified_labels': _modified_labels.cpu(),
'train_predictions': _predictions.cpu()
}
)
print(f"Writing data parameters output to '{train_label_snapshot_path}'")
torch.save(save_dict, train_label_snapshot_path)
if len(training_dataset.disturbed_idxs) > 0:
# Log histogram
separated_params = list(zip(dp_weights[clean_idxs], dp_weights[training_dataset.disturbed_idxs]))
s_table = wandb.Table(columns=['clean_idxs', 'disturbed_idxs'], data=separated_params)
fields = {"primary_bins": "clean_idxs", "secondary_bins": "disturbed_idxs", "title": "Data parameter composite histogram"}
composite_histogram = wandb.plot_table(vega_spec_name="rap1ide/composite_histogram", data_table=s_table, fields=fields)
wandb.log({f"data_parameters/separated_params_fold_{fold_idx}": composite_histogram})
# Write out data of modified and un-modified labels and an overview image
if training_dataset.use_2d():
reduce_dim = None
in_type = "batch_2D"
skip_writeout = len(training_dataset) > 3000 # Restrict dataset size to be visualized
else:
reduce_dim = "W"
in_type = "batch_3D"
skip_writeout = len(training_dataset) > 150 # Restrict dataset size to be visualized
skip_writeout = True
if not skip_writeout:
print("Writing train sample image.")
# overlay text example: d_idx=0, dp_i=1.00, dist? False
overlay_text_list = [f"id:{d_id} dp:{instance_p.item():.2f}" \
for d_id, instance_p, disturb_flg in zip(d_ids, dp_weight, disturb_flags)]
use_2d = training_dataset.use_2d()
scf = 1/training_dataset.pre_interpolation_factor
show_img = interpolate_sample(b_label=_labels.to_dense(), scale_factor=scf, use_2d=use_2d)[1].unsqueeze(1)
show_seg = interpolate_sample(b_label=_predictions.to_dense().squeeze(1), scale_factor=scf, use_2d=use_2d)[1]
show_gt = interpolate_sample(b_label=_modified_labels.to_dense(), scale_factor=scf, use_2d=use_2d)[1]
visualize_seg(in_type=in_type, reduce_dim=reduce_dim,
img=show_img, # Expert label in BW
seg=4*show_seg, # Prediction in blue
ground_truth=show_gt, # Modified label in red
crop_to_non_zero_seg=False,
alpha_seg = .5,
alpha_gt = .5,
n_per_row=70,
overlay_text=overlay_text_list,
annotate_color=(0,255,255),
frame_elements=disturb_flags,
file_path=seg_viz_out_path,
)
# End of fold loop
# %%
# Config overrides
# config_dict['wandb_mode'] = 'disabled'
# config_dict['debug'] = True
# Model loading
# config_dict['checkpoint_name'] = 'ethereal-serenity-1138'
# config_dict['fold_override'] = 0
# config_dict['checkpoint_epx'] = 39
# Define sweep override dict
sweep_config_dict = dict(
method='grid',
metric=dict(goal='maximize', name='scores/val_dice_mean_tumour_fold0'),
parameters=dict(
# disturbance_mode=dict(
# values=[
# 'LabelDisturbanceMode.AFFINE',
# ]
# ),
# disturbance_strength=dict(
# values=[0.1, 0.2, 0.5, 1.0, 2.0, 5.0]
# ),
# disturbed_percentage=dict(
# values=[0.3, 0.6]
# ),
# data_param_mode=dict(
# values=[
# DataParamMode.INSTANCE_PARAMS,
# DataParamMode.DISABLED,
# ]
# ),
use_risk_regularization=dict(
values=[False, True]
),
use_fixed_weighting=dict(
values=[False, True]
),
# fixed_weight_min_quantile=dict(
# values=[0.9, 0.8, 0.6, 0.4, 0.2, 0.0]
# ),
)
)
# %%
def normal_run():
with wandb.init(project="deep_staple", group="training", job_type="train",
config=config_dict, settings=wandb.Settings(start_method="thread"),
mode=config_dict['wandb_mode']
) as run:
run_name = run.name
print("Running", run_name)
training_dataset = prepare_data(config_dict)
config = wandb.config
train_DL(run_name, config, training_dataset)
def sweep_run():
with wandb.init() as run:
run = wandb.init(
settings=wandb.Settings(start_method="thread"),
mode=config_dict['wandb_mode']
)
run_name = run.name
print("Running", run_name)
training_dataset = prepare_data(config)
config = wandb.config
train_DL(run_name, config, training_dataset)
if config_dict['do_sweep']:
# Integrate all config_dict entries into sweep_dict.parameters -> sweep overrides config_dict
cp_config_dict = copy.deepcopy(config_dict)
# cp_config_dict.update(copy.deepcopy(sweep_config_dict['parameters']))
for del_key in sweep_config_dict['parameters'].keys():
if del_key in cp_config_dict:
del cp_config_dict[del_key]
merged_sweep_config_dict = copy.deepcopy(sweep_config_dict)
# merged_sweep_config_dict.update(cp_config_dict)
for key, value in cp_config_dict.items():
merged_sweep_config_dict['parameters'][key] = dict(value=value)
# Convert enum values in parameters to string. They will be identified by their numerical index otherwise
for key, param_dict in merged_sweep_config_dict['parameters'].items():
if 'value' in param_dict and isinstance(param_dict['value'], Enum):
param_dict['value'] = str(param_dict['value'])
if 'values' in param_dict:
param_dict['values'] = [str(elem) if isinstance(elem, Enum) else elem for elem in param_dict['values']]
merged_sweep_config_dict['parameters'][key] = param_dict
sweep_id = wandb.sweep(merged_sweep_config_dict, project="deep_staple")
wandb.agent(sweep_id, function=sweep_run)
else:
normal_run()
# %%
if not in_notebook():
sys.exit(0)
# %%
# Do any postprocessing / visualization in notebook here
|
main_deep_staple.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.1
# language: julia
# name: julia-0.6
# ---
# # Referências
# - Principles of Data Science - <NAME>
# - Practical Data Science — <NAME>
# - The Data Science Handbook - Field Cady
# - Introduction to Data Mining - <NAME>, <NAME>, <NAME>
# ### Dados
# Dados podem se encontrar de forma *organizada* onde linhas representam observações/entidades e colunas represetam características. Essa forma é representada por uma tabela ou matriz, e assim como uma matriz ela pode ser transposta onde as colunas representarão observações e linhas representarão as características.
#
# Note que utilizamos o conceito de matriz que pode ser estudada em álgebra linear. Em Data Science você utilizará muitos conceitos matemáticos relacionados à álgebra linear, estatística e cálculo por exemplo, mas este documento tem a intenção de ensiná-los (e também ao autor) todos os conceitos necessários e relacionados com a ciência de dados. Os conhecimentos básicos necessários como os de álgebra, estatística, cálculo e algoritmos estarão em capítulos simbolizados por letras ao invés de números, como por exemplo os capítulo A (ou A1, A2, etc) que corresponde ao capítulo dedicado ao ensino e aprendizagem de álgebra linear.
# ### Amostragem
# Amostragem é o processo de selecionar subconjuntos de dados para diminuir o volume para processamento. Se a amostra selecionada for representativa então ela funcionará tão bem quanto a utilização do conjunto inteiro de dados que poderia facilmente ultrapassar milhões de linhas em uma matriz ou tabela.
#
# A literatura traz diversas técnicas em relação à amostragem, definiremos aqui algumas e posteriormente introduziremos outras. *#(Ou este mesmo trecho/capítulo será editado)*
#
# #### Amostragem aleatória simples
# (sem substituição e com substituição)
#
# #### Amostragem estratificada
#
# #### Tamanho da amostra
#
# #### Amostragem progressiva
# Consiste em fazer várias amostragens de tamanho cada vez maior e escolher o mais apropriado.
#
# É preciso parar pois em determinado momento o aumento de tamanho já não implica em aumento de exatidão.
#
# #### Redução de dimensionalidade
# A redução da dimensionalidade pode eliminar características irrelevantes e reduzir ruídos.
#
# Isso pode ser feito distribuindo-se pesos para os atributos em grau de importância, também pode ser feito através da discretização que consiste em transformar atributos contínuos em discretos ou mesmo binarização que seria a transformação de atributos contínuos ou discretos em um ou mais atributos binários de acordo com o necessário.
# #### Discretização supervisionada
# *#(Necessário pesquisar melhor sobre discretiação supervisionada e entropia)*
#
# ##### Entropia do intervalo
#
# $$ e_i = \sum_{i = 1}^{k} p_{ij} log_2p_{ij} $$
#
# Onde $k$ = número de classes, $m_i$ = valores no intervalo i, $m_{ij}$ = valores da classe j no intervalo i e $i$ = índice. Seja $p_{ij} = \frac{m_{ij}}{m_i} $ a probabilidade da classe j no intervalo de índice i.
#
# ##### Entropia total
#
# $$ e = \sum_{i = 1}^{n} w_ie_i $$
#
# Onde $n$ = número de intervalos, $m$ = número de valores e $w_i = \frac{m_i}{m}$$ a fração dos valores no intervalo de índice 1.
|
1 - Dados.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import pickle
from tqdm import tqdm_notebook as tqdm
from joblib import Parallel
db = pickle.load(open( "db.pkl", "rb" ))
placement_hist = pickle.load(open("plot_hist.pkl","rb"))
db.num_physical_nodes + db.num_filler_nodes
# +
# first n nodes are physical. Last are filler.
# placement_hist is a list of [x:y] positions
# -
from draw_place import DrawPlace
d = DrawPlace(db)
d.forward(placement_hist[0],'test.png')
placement_hist[100]
id2name = {v: k for k, v in db.node_name2id_map.items()}
# +
# too slow
placement_hist_dfs = []
for i in tqdm(range(db.num_nodes)):
xpos = []
ypos = []
placement_df = pd.DataFrame(columns=['id','x','y'])
id_col = [id2name[i]]*len(placement_hist)
for placement in placement_hist:
xpos.append(placement[i])
ypos.append(placement[db.num_nodes + i])
placement_df['id'] = id_col
placement_df['x'] = xpos
placement_df['y'] = ypos
placement_hist_dfs.append(placement_df)
placement_hist_df = pd.concatenate(placement_hist_dfs)
# -
np.stack(placement_hist).shape
np.stack(placement_hist)[:,1]
# +
placement_mat = np.stack(placement_hist)
for i in tqdm(range(db.num_nodes)):
placement_df = pd.DataFrame(columns=['id','x','y'])
id_col = [id2name[i]]*len(placement_hist)
xpos = placement_mat[:,i]
ypos = placement_mat[:,db.num_nodes + i]
placement_df['id'] = id_col
placement_df['x'] = xpos
placement_df['y'] = ypos
placement_hist_dfs.append(placement_df)
placement_hist_df = pd.concatenate(placement_hist_dfs)
# -
|
.ipynb_checkpoints/cluster-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import datacube
dc = datacube.Datacube(config="/home/sharat910/.datacube.conf")
dc
dc.list_products()
dc.list_measurements()
la = dc.load(product='ls5_ledaps_albers', x=(79.0, 79.05), y=(30.0, 30.05))
la
la.data_vars
la.blue
a = la.blue.loc['1995-03']
a.shape
a.plot()
|
examples/notebooks/TestIpython.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating Financial Industry Word Dictionary
training_data_folder = "/Users/sudarshan/Google Drive/1. Academic Files/Semesters/Fall 2018/INDENG 135/Paradigm (Fall 18) - Team 2/Jupyter Notebooks/Sentiment Analysis/Sentiment Training Data/"
# +
import json
import pandas as pd
json_data = training_data_folder + "ntusd_data.json"
with open(json_data) as f:
json_df = pd.read_json(f)
print("Positive Words: {}".format(len(json_df[json_df["market_sentiment"] > 0])))
print("Neutral Words: {}".format(len(json_df[json_df["market_sentiment"] == 0])))
print("Negative Words: {}".format(len(json_df[json_df["market_sentiment"] < 0])))
min_val = json_df["market_sentiment"].min()
max_val = json_df["market_sentiment"].max()
for i, row in json_df.iterrows():
x = row["market_sentiment"]
if x > 0:
x = x / max_val
else:
x = -1 * (x / min_val)
json_df.at[i,'market_sentiment'] = x
json_df["market_sentiment"] = json_df["market_sentiment"] + 1
json_df["market_sentiment"] = json_df["market_sentiment"] / 2
print("\nNormalized Positive Words: {}".format(len(json_df[json_df["market_sentiment"] > 0.5])))
print("Normalized Neutral Words: {}".format(len(json_df[json_df["market_sentiment"] == 0.5])))
print("Normalized Negative Words: {}".format(len(json_df[json_df["market_sentiment"] < 0.5])))
json_df.to_csv("NTUSD_data.csv")
# -
# -----------------------------------------------
vocabulary = pd.read_csv(training_data_folder + "vocabulary.csv")
vocabulary["Word"] = vocabulary["Word"].str.lower()
# +
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
vocabulary_string = ""
for word in vocabulary["Word"]:
vocabulary_string += word + ","
tokenizer = Tokenizer(num_words=len(vocabulary["Word"]), split=",", char_level=False)
tokenizer.fit_on_texts(vocabulary["Word"].values)
X = tokenizer.texts_to_sequences(vocabulary["Word"].values)
temp = []
for i in X:
temp.append(len(i))
X = pad_sequences(X, maxlen = max(temp))
# -
|
past-team-code/Fall2018Team2/Sentiment Analysis/Old Iterations/WordDictionary.ipynb
|
# ---
# layout: post
# title: "Bitcoin Futures Arbitrage Part 4"
# categories:
# - Bitcoin Futures Series
# tags:
# - bitcoin
# - futures
# - perpetual future
# - deribit
# - python
# - arbitrage
# - data science
# - investments
# - monte carlo simulation
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This is the fourth of a series about bitcoin futures. We will use a Monte Carlo simulation to model the profitability of an arbitrage strategy.
#
# - [Part 1 - Getting the data]({% post_url 2019-05-11-bitcoin-futures-arbitrage-part-1 %})
# - [Part 2 - Were there arbitrage profits in the past?]({% post_url 2019-05-12-bitcoin-futures-arbitrage-part-2 %})
# - [Part 3 - Perpetual futures 101]({% post_url 2019-05-20-bitcoin-futures-arbitrage-part-3 %})
# - [Part 4 - Arbitrage Profit Analysis]({% post_url 2019-05-24-bitcoin-futures-arbitrage-part-4 %})
#
#
# Let's load our perpetual data again.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy
from matplotlib import style
import seaborn as sns
from IPython.core.pylabtools import figsize
import warnings
import glob
import statsmodels.api as sm
import datetime
# %matplotlib inline
plt.style.use('ggplot')
np.random.seed(17098234)
# -
df = pd.read_csv('/home/chris/code/deribit-future-arb/downloads/BTC-PERPETUAL.txt')
df['date'] = pd.to_datetime(df['timestamp'], unit='ms')
df.index = df['date']
df = df.drop(columns=['timestamp', 'date', 'instrument_name'])
df.columns = ['perpetual', 'index']
df['ratio'] = df['perpetual'] / df['index']
df['funding'] = np.maximum(0.0005, df['ratio'] - 1) + np.minimum(-0.0005, df['ratio'] - 1)
# df_D = df.resample('D').last()
# df_H = df.resample('H').last().interpolate()
df_5 = df.resample('5Min').last().interpolate()
# # Monte Carlo Simulation
#
# Let's calculate the profitability of arbitrage, given those prices from earlier that I sampled from a random moment in time.
# +
# Starting prices
start_future_price = 7270.13
start_perpetual_price = 7325.88
start_index_price = 7335.49
days = 46
# Initial Prices are the set of index, perpetual, and futures prices I recorded and used in the example earlier.
# This will simulate the profit considering you bought and sold for those prices.
# Enter in new prices to get a profitability analysis for those prices
# -
# I make a few assumptions:
#
# 1. The trades are done with 1 BTC of exposure.
# 2. The volatility of BTC is about 5% daily, based on a quick look [here](https://www.sk3w.co/options).
# 3. The BTC index price at the future's expiration date will be calculated by running forward random returns until expiration.
# 4. The perpetual price at the future's expiration date will equal the index price, times by a perpetual index ratio sampled from our perpetual index distribution.
# 5. The average funding rate over the investment period will be sampled from the distribution of past average funding rates over the same length period.
#
# Here's the full model specification as:
#
# $$\text{btc_daily_returns} \sim N(0, 0.05)$$
# $$\text{end_index_price} \sim f(\text{start_index_price}, \text{btc_daily_returns})$$
# $$\text{perpetual_index_ratio} \sim \text{real perpetual index data}$$
# $$\text{end_perpetual_price} = \text{end_index_price} * \text{perpetual_index_ratio}$$
# $$\text{average_funding_rate} \sim \text{real funding rate data}$$
# $$\text{total_funding} = f(\text{average_funding_rate}, \text{days})$$
# $$\text{profit} = f(\text{end_index_price}, \text{end_perpetual_price}, \text{total_funding})$$
#
# +
trials = 1000000
# Amount of btc trade
btc = 1
daily_btc_returns = np.random.normal(loc=0.00, scale=0.05, size=(trials, days))
total_btc_return = np.prod(1 + daily_btc_returns, axis=1)
end_index_price = total_btc_return * start_index_price
end_perpetual_ratio = np.random.choice(df_5['ratio'], size=trials)
end_perpetual_price = end_index_price * end_perpetual_ratio
# On deribit futures are restricted to contracts of $10
# That means your expoure might not be perfectly equal in long and short
future_amount = round(start_future_price * btc, -1)
perpetual_amount = round(start_perpetual_price * btc, -1)
# Payoffs from long future, short perpetual
long_future_payoff = future_amount * (1/start_future_price - 1/end_index_price) * end_index_price
short_perpetual_payoff = -perpetual_amount * (1/start_perpetual_price - 1/end_perpetual_price) * end_index_price
short_perpetual_profit = long_future_payoff + short_perpetual_payoff
# Payoffs from short future, long perpetual
short_future_payoff = -future_amount * (1/start_future_price - 1/end_index_price) * end_index_price
long_perpetual_payoff = perpetual_amount * (1/start_perpetual_price - 1/end_perpetual_price) * end_index_price
long_perpetual_profit = short_future_payoff + long_perpetual_payoff
# -
figsize(14, 3)
sns.kdeplot(short_perpetual_profit, label='Long Future Short Perpetual')
sns.kdeplot(long_perpetual_profit, label='Short Future Long Perpetual')
plt.xlabel('Profit ($)')
plt.ylabel('Relative Frequency')
plt.title('Distribution of Profits Before Fees and Funding Payments')
plt.xlim([-80, 80])
plt.show()
# ## Fees
#
# Our model has not included any fees. So we will do those calculations and update our profits.
#
# The fees for trading on Deribit can be found [here](https://www.deribit.com/pages/information/fees)
#
# They are listed below in a percentage of the underlying asset (negative numbers mean a rebate):
#
# Instrument | Maker Trade Fee | Taker Trade Fee | Delivery Fee
# --- | --- | --- | ---
# Future | 0.05% | -0.02% | 0.025%
# Perpetual | 0.075% | -0.025% | 0.075%
#
# It is possible to earn a profit on trades by only setting limit orders. Then our total fees would be:
#
# Trade | Fees
# --- | ---
# Buy Future | -0.02%
# Sell Perpetual | -0.025%
# Settle Future | 0.025%
# Buy Perpetual | -0.025%
#
# For a total rebate of 0.045% of the underlying asset.
#
# If we used all market orders the fees would be:
#
# Trade | Fees
# --- | ---
# Buy Future | 0.05%
# Sell Perpetual | 0.075%
# Settle Future | 0.025%
# Buy Perpetual | 0.075%
#
# For a total fee amount of 0.225%.
# +
taker_fees = start_index_price * 0.225 / 100
maker_rebate = start_index_price * 0.045 / 100
short_perpetual_taker = short_perpetual_profit - taker_fees
short_perpetual_maker = short_perpetual_profit + maker_rebate
long_perpetual_taker = long_perpetual_profit - taker_fees
long_perpetual_maker = long_perpetual_profit + maker_rebate
# -
figsize(14,3)
sns.kdeplot(short_perpetual_taker, label='Taker Short Perpetual')
sns.kdeplot(short_perpetual_maker, label='Maker Short Perpetual')
sns.kdeplot(long_perpetual_taker, label='Taker Long Perpetual')
sns.kdeplot(long_perpetual_maker, label='Maker Long Perpetual')
plt.xlabel('Profit ($)')
plt.ylabel('Relative Frequency')
plt.title('Distribution of Profits After Fees and Before Funding Payments')
plt.xlim([-100, 100])
plt.show()
# Going short on the perpetual is the most profitable strategy, as expected. So far we have a fairly narrow distribution around \\$60 profit, exactly what we expected from the difference in starting prices. We will limit our analysis to going short on the perpetual and paying maker fees.
#
# ## Funding
#
# We need to know what the expected funding rate is over the time period. So we will take average funding rates from a number of time periods the same length as our time horizon.
five_min_per_day = int(24 * 60 / 5)
samples = five_min_per_day * days
windows = np.convolve(df_5['funding'], np.ones((samples,))/samples, mode='valid')
funding_rate = np.random.choice(windows, size=trials)
funding_rate = np.random.normal(loc=windows.mean(), scale=windows.std(), size=trials)
figsize(8,3)
sns.kdeplot(funding_rate, label=f'8 Hour Funding Rate Averaged Over {days} days')
plt.xlabel('Rate')
plt.ylabel('Relative Frequency')
plt.title('Distribution of Average Funding Rates')
plt.xlim([-0.00075, 0.00075])
plt.show()
funding_payments = perpetual_amount * funding_rate * 3 * days
print(funding_payments.mean())
print(funding_payments.std())
figsize(6,3)
sns.kdeplot(funding_payments, label=f'Dollar Amount of Funding Payments over the {days} days')
plt.xlabel('Profit ($)')
plt.ylabel('Relative Frequency')
plt.title('Distribution of Funding Payments for our Perpetual Exposure')
plt.xlim([-750, 750])
plt.show()
# +
final_short_perpetual_taker = short_perpetual_taker + funding_payments
final_short_perpetual_maker = short_perpetual_maker + funding_payments
final_long_perpetual_taker = long_perpetual_taker - funding_payments
final_long_perpetual_maker = long_perpetual_maker - funding_payments
# -
figsize(6,3)
# sns.kdeplot(final_short_perpetual_taker, label='Taker Short Perpetual')
sns.kdeplot(final_short_perpetual_maker, label='Maker Short Perpetual')
# sns.kdeplot(final_long_perpetual_taker, label='Taker Long Perpetual')
# sns.kdeplot(final_long_perpetual_maker, label='Maker Long Perpetual')
plt.xlabel('Profit ($)')
plt.ylabel('Relative Frequency')
plt.title('Distribution of Final Profits')
plt.xlim([-750, 750])
plt.show()
# +
print(final_short_perpetual_maker.mean())
print(final_short_perpetual_maker.std())
# print(final_long_perpetual_maker.mean())
# print(final_long_perpetual_maker.std())
# -
# ## Rate of Return
#
# The model suggests that our strategy has positive expected profit (of approximately \\$60) with extremely high variance. And the benefit of this strategy, compared to traditional arbitrage, is that we earn that \\$60 using a much lower initial investment.
#
# Deribit futures allow up to 100x leverage. And Deribit has a portfolio margin system where your required margin is a function of your total portfolio risk due to bitcoin price fluctuations. The details are [here](https://www.deribit.com/pages/docs/portfoliomargin). As we are neutral on bitcoin, our required margin is only 1.2%. But we would still need margin to cover fluctuations in the funding rate.
#
# Below I will calculate the single largest day of funding.
single_day = np.max(np.abs(df_5.resample('D').mean()['funding']))
single_day
# This gives a good estimation of our required margin. It would be a good idea to have the minimum 1.2% plus enough to cover 5 days of the worst luck.
margin_amount = (single_day * 3 * perpetual_amount * 5) + (perpetual_amount * 0.012)
margin_amount
# This amount of upfront investment is much less than doing arbitrage the traditional way. That requires the purchase of 1 BTC or about \\$7000 of initial cash outflow.
#
# ## Margin Risk
#
# Depositing margin at Deribit means holding the margin in bitcoins while you wait for the future to expire. This exposes you to bitcoin risk as your margin will be worth more or less as the price changes. We can see that in the following graph.
# +
final_margin = margin_amount * (end_index_price/start_index_price)
margin_profit = final_margin - margin_amount
sns.kdeplot(margin_profit)
plt.title('Profit from Fluctuation of Margin Value')
plt.ylabel('Relative Frequency')
plt.xlabel('Profit ($)')
plt.xlim(-200,400)
plt.show()
# -
# Even holding a small amount of bitcoin is risky. A better solution is to short the perpetual for the amount we have in margin.
# +
margin_funding_payments = margin_amount * funding_rate * 3 * days
sns.kdeplot(margin_funding_payments)
plt.ylabel('Relative Frequency')
plt.xlabel('Amount ($)')
plt.title('Total Funding Payments due to Margin')
plt.xlim(-30,30)
plt.show()
# -
# ## Calculating Rate of Return
#
# We have calculated the total expected profit on making the arbitrage trade and holding the assets until expiration. To compare with other investments, let's calculate the annualized rate of return. The return will be our total profits over our initial required margin.
# +
profit = final_short_perpetual_maker + margin_funding_payments
# profit = final_long_perpetual_maker + margin_funding_payments
# Rate of Return (simple annualized) on the long perpetual strategy
RR = (profit / margin_amount) * (365 / days) * 100
# -
figsize(6,3)
sns.kdeplot(RR)
plt.ylabel('Relative Frequency')
plt.xlabel('Annual Return (%)')
plt.xlim(-1500,1500)
plt.show()
print(RR.mean())
print(RR.std())
# Our strategy has an extremely high rate of return with an insanely high variance. A 150% expected return with a 500% standard deviation is crazy high. But it makes sense with expecting to earn \\$60 on \\$300 of upfront capital in under 50 days.
#
# A globally diversified stock portfolio has something like a real expected rate of return of 6% with a standard deviation of 15%.
#
# ## Sensitivity Analysis
#
# Let's check which input variables have the biggest effect on profits.
#
# First we will see how the expected return varies as a function of our model parameters. We can do this by only looking at the modelled situations that happened to have a parameter fall within a specific range.
def linear_transform(x, xs):
max_xs = np.max(xs)
min_xs = np.min(xs)
return (x - min_xs) / (max_xs - min_xs)
# +
figsize(8,4)
btc_price_change = end_index_price / start_index_price - 1
labels = ['BTC Price Change', 'Final Perpetual Index Ratio', 'Funding Rate']
input_variables = [btc_price_change, end_perpetual_ratio, funding_rate]
# Sensitivity analysis
# expected value as each variable changes
for index, input_variable in enumerate(input_variables):
# split variable into groups
step = (input_variable.max() - input_variable.min())/10
ys = []
xs = np.linspace(input_variable.min(), input_variable.max(), 100)
for i in xs:
## values where input_variable is close to i
bool_array = np.logical_and(input_variable < i + step, input_variable > i - step)
ys.append(RR[bool_array].mean())
# transform xs to be on scale of 0 to 1 for the graph
linear_xs = []
for x in xs:
linear_xs.append(linear_transform(x, xs))
plt.plot(linear_xs, ys, label=labels[index])
plt.xlabel('Min Value to Max Value of Each Variable')
plt.ylabel('Expected Annualized RR (%)')
plt.title('Effects of Variable Changes on Expected RR')
plt.axhline(y=RR.mean(), color='g', linestyle='--', label='Total Average Expected Return')
plt.legend()
plt.show()
# -
# Most of these variables are normally distributed and thus there are fewer samples near the max and min values. This explains the variance of the RR in the bottom and top percentiles of parameter values.
#
# ## Profit Factors
#
# The single biggest factor in the profitability of this strategy is the funding rate we expect over the investment period.
#
# Otherwise we are approximately bitcoin neutral and slightly dependant on the final perpetual index ratio.
#
# The return seems to be only really strongly correlated with the average funding rate. But the funding rate appeared to be a random process. It's likely the funding rate isn't highly correlated with conventional asset classes such as stocks or bonds and thus this strategy is not correlated with the market.
#
# This strategy has poor risk-return characteristics if you were considering it as your only investment. But its low correlation with other assets makes it likely to be a part of an optimal mean-variance portfolio. In the next post I will perform a Markowitz portfolio optimization and see how this strategy fits as part of a portfolio.
#
# ## Was this situation unusual?
#
# Did I get lucky and find a particularly profitable moment in time that was quickly arbitraged away?
#
# Below I graph the ratio used in this analysis compared to the historical data.
# +
# Get a list of files with the historical data
files = glob.glob('/home/chris/code/deribit-future-arb/downloads/futures/*.txt')
# Combine the files into one large pandas df
data_list = []
for file in files:
data = pd.read_csv(file)
data_list.append(data)
df_futures = pd.concat(data_list)
df_futures['ratio'] = df_futures['price'] / df_futures['index_price']
instrument_names = ['BTC-27JUL18', 'BTC-31AUG18', 'BTC-28SEP18', 'BTC-28DEC18', 'BTC-29MAR19']
# -
figsize(10,4)
for i, instrument in enumerate(instrument_names):
temp = df_futures[df_futures['instrument_name'] == instrument]
sns.kdeplot(temp['ratio'], label=instrument)
plt.xlabel('Ratio')
plt.ylabel('Relative Frequency')
plt.title('Future Index Ratio Distribution')
plt.xlim(0.95, 1.05)
specific_ratio=start_future_price/start_index_price
plt.axvline(x=specific_ratio, color='g', linestyle='--', label='MC Model Ratio')
plt.legend()
plt.show()
# The prices used in this model are not particularly unusual. These situations happen frequently.
#
# ## Is the order book deep enough?
#
# In this post I assumed we just got one bitcoin of exposure, using about $300 of initial capital. That's not a large investment. And as you attempt to arbitrage the differences in prices, you will push the prices closer together and reduce your profits.
#
# So I took a quick look at the exchange. Right now as I type this, it would take about $100,000 to move the perpetual price by 0.05%.
#
# Our analysis was done based on a price difference of about 1% between the future and the perpetual. If you change the input prices it still has a large expected return at 0.95% difference.
#
# I predict that this strategy will be part of an optimal mean-variance portfolio and that you could put millions into this and still get a risk-return profile that should be included in an optimal portfolio. I will check this prediction in my next post.
#
# # Conclusion
#
# We created a model for the profitability of an arbitrage strategy given any future and perpetual prices. This model could be turned into a trading strategy.
#
# 1. Run this simulation on a variety of future/perpetual price ratios for different times to expiry. Cache the expected return and variance of each.
# 2. Calculate covariances between these strategies and other assets.
# 3. Get live data on ratios.
# 4. Calculate optimal portfolio using expected returns and covariances.
# 5. Invest
|
_notebooks/2019-05-24-bitcoin-futures-arbitrage-part-4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="3d979c7f"
# First we download and import all neccessary packages
# + id="5aVhK5fwUFZr" colab={"base_uri": "https://localhost:8080/"} outputId="782a9bcc-6aa4-4d8b-ba77-786a786db283"
# !git clone --depth=1 https://github.com/ntapiam/dagstuhl.git
# %cd dagstuhl
# + id="3f09593d"
import isstorch
from tqdm.notebook import trange
from utils import generate_examples, compute_signatures
import torch
import torch.nn as nn
from torch.optim import Adam
from torch.utils.data import DataLoader, TensorDataset
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="XUVWT0xoUa4e"
# Set the default `torch` device depending on whether there is a GPU available. We also run a random example to measure speed and check that everything is working OK.
# + id="0727924b" colab={"base_uri": "https://localhost:8080/", "height": 468} outputId="1f3b863f-6684-46a1-b0aa-fdec13e651a7"
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
x = torch.randn(1000, 100, device=device)
with torch.no_grad():
# %time isstorch.compute(x); torch.cuda.synchronize('cuda')
x = x.to('cpu')
with torch.no_grad():
# %time isstorch.compute(x);
# + [markdown] id="3f294f52"
# We build our model using by subclassing `torch.nn.Module`. We use a stack of dense layers with ReLU activation. To see all possible layers and activations, please see the `torch.nn` [documentation](https://pytorch.org/docs/stable/nn.html).
# + id="f2b3e163"
class DenseNet(nn.Module):
def __init__(self, hidden_layers=1, width=10, in_features=3):
super().__init__()
self.batch_norm = nn.BatchNorm1d(in_features)
input_layer = nn.Sequential(
nn.Linear(in_features=in_features, out_features=width),
nn.Tanh()
)
layers = [input_layer]
for k in range(hidden_layers):
layers.append(nn.Sequential(
nn.Linear(in_features=width, out_features=width),
nn.Tanh()
)
)
layers.append(
nn.Sequential(
nn.Linear(in_features=width, out_features=2),
nn.LogSoftmax(dim=1)
)
)
self.stack = nn.Sequential(*layers)
def forward(self, x):
if x.shape[0] > 1:
x = self.batch_norm(x)
return self.stack(x)
# + [markdown] id="31146f47"
# Next, we set up the training and testing loops
# + id="524e7150"
def train_loop(model, dataloader, loss_fn, optimizer, device):
for X, y in dataloader:
X, y = X.to(device), y.to(device)
pred = model(X)
loss = loss_fn(pred, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
def test_loop(model, dataloader, loss_fn, device):
total_samples, batches = len(dataloader.dataset), len(dataloader)
test_loss, correct = 0, 0
with torch.no_grad():
for X, y in dataloader:
X, y = X.to(device), y.to(device)
pred = model(X)
test_loss += loss_fn(pred, y).item()
correct += (pred.argmax(dim=1) == y).type(torch.long).sum().item()
return correct / total_samples, test_loss / batches
# + [markdown] id="73a55913"
# Let's look at some samples. The function `generate_samples` outputs a different set of data each time.
# + id="6637121d" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="c90f6960-1bd4-4e2c-fcc9-8f34c19ffa70"
X, labels = generate_examples()
plt.plot(range(100), X[0], label="class 0")
plt.plot(range(100), X[500], label="class 1")
plt.xlabel('t')
plt.legend()
plt.show()
# + [markdown] id="27d94aa5"
# Now, we compute the signature features using our `pytorch` implementation.
# + id="f9a2bd78" colab={"base_uri": "https://localhost:8080/"} outputId="1f53bbb5-ca20-4401-a1d8-40a01d4ff92c"
with torch.no_grad():
# %time sigs = compute_signatures(X, level=4)
print(sigs.shape)
in_features = sigs.shape[1]
# + [markdown] id="5f796922"
# In order to train and test our Neural Network, we need to wrap the data in a PyTorch `DataLoader`. We also load the data in batches.
# + id="0f11e74f"
train_sigs, test_sigs, train_labels, test_labels = train_test_split(sigs, labels, stratify=labels, train_size=0.7)
train_dataloader = DataLoader(TensorDataset(train_sigs, train_labels), batch_size=100)
test_dataloader = DataLoader(TensorDataset(test_sigs, test_labels), batch_size=100)
# + [markdown] id="7f7d5136"
# Setup our model, loss function and optimizer. Train the network and output the result.
# + id="eb4fe2ca" colab={"base_uri": "https://localhost:8080/", "height": 118, "referenced_widgets": ["cd59f6ede4ca4aa3a43bcc01c84ca1b9", "6fcb24ad3cc540f298804cd9b93ed96a", "<KEY>", "a3beeb83741644a0b6834fed914055c3", "3af1a4323243432c9e57f5c8da49414f", "9a4c8fe6e0e44ee688dd68e515d7ed18", "<KEY>", "ef4f725e2bc14e1f992db3452d18e1fd", "2caf5c249be34791bb8de2027946987c", "1eb29c0ecd1046d08900427a2edfa9ce", "b04e88ea69524eecbfcf6d3a97f62142"]} outputId="51290cef-33df-462e-d57e-f992a9102bbf"
# %%time
model = DenseNet(hidden_layers=1, width=in_features, in_features=in_features).to(device)
optimizer = Adam(model.parameters(), lr=1e-3)
loss_fn = nn.NLLLoss()
epochs = 200
losses = []
accuracies = []
with trange(epochs) as pbar:
for t in pbar:
train_loop(model, train_dataloader, loss_fn, optimizer, device)
acc, avg_loss = test_loop(model, test_dataloader, loss_fn, device)
losses.append(avg_loss)
accuracies.append(acc)
pbar.set_description(f"Training")
pbar.set_postfix(loss=avg_loss, accuracy=acc)
acc, avg_loss = test_loop(model, test_dataloader, loss_fn, device)
print(f"\nDone! Accuraccy: {acc:.2%}, Loss: {avg_loss:f}")
# + [markdown] id="0NEYm84TuEHp"
# Now, we can make a predicition on our test data (or even new inputs), which will be given as a probability distribution over class labels.
#
# Note that we apply the exponential mapping in order to obtain probabilities, as the network outputs negative log probas.
# + colab={"base_uri": "https://localhost:8080/"} id="EurkCtLfsE-F" outputId="afc06dd6-b017-47b2-d0b6-45747cd04096"
preds = model(test_sigs[:20].cuda()).exp().cuda()
preds
# + [markdown] id="IYdBTLwjuf8n"
# We check that they sum to 1 for each example.
# + colab={"base_uri": "https://localhost:8080/"} id="crgNwQUrunrK" outputId="3b354df6-6b86-4437-e123-93a19b08f0d8"
torch.allclose(preds.sum(dim=1), torch.ones(20, device='cuda'))
# + [markdown] id="HGv4o4ocwUWe"
# The predicted label is the one with the highest probability.
# + colab={"base_uri": "https://localhost:8080/"} id="Zda9btrxuxYT" outputId="f7b4281f-461d-4577-eab2-c9671d5e4b3d"
preds.argmax(dim=1), test_labels[:20]
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="xTL6ueOcWbIW" outputId="5c43172c-bf77-4d52-bbc1-0538769db14a"
sns.set_theme()
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.plot(range(epochs), losses, color='tab:red')
ax2.plot(range(epochs), accuracies, color='tab:blue')
ax1.set_xlabel('epochs')
ax1.set_ylabel('losses')
ax2.set_ylim(bottom=0, top=1)
ax2.set_ylabel('accuracy')
fig.tight_layout()
plt.plot()
# + id="2fecveXkdKOB"
train_data, test_data, train_labels, test_labels = train_test_split(X.flatten(start_dim=1), labels, stratify=labels, train_size=0.7)
train_dataloader = DataLoader(TensorDataset(train_data, train_labels), batch_size=100)
test_dataloader = DataLoader(TensorDataset(test_data, test_labels), batch_size=100)
# + id="MvwNfc0btD9j"
# + colab={"base_uri": "https://localhost:8080/", "height": 118, "referenced_widgets": ["447b7e71b46b4b6cbca2e8449c523d21", "8ecc77abab3540a580e6197e082b8e56", "<KEY>", "<KEY>", "dbe344e7b3a740cdb24467ec7b0eb7af", "79394bca56684c3e8c00fdca5f3bbc2e", "b62e6c7d1f9544d8ae482a3e52c2a579", "<KEY>", "<KEY>", "<KEY>", "9b8cd40ec09e4fd0a38d8092816498e3"]} id="ndZYJ0zfdaPm" outputId="50cd5229-544f-462d-9282-fabe482a0855"
# %%time
in_features=100
new_model = DenseNet(hidden_layers=1, width=in_features, in_features=in_features).to(device)
new_optimizer = Adam(new_model.parameters(), lr=1e-3)
with trange(epochs) as pbar:
for t in pbar:
train_loop(new_model, train_dataloader, loss_fn, new_optimizer, device)
acc, avg_loss = test_loop(new_model, test_dataloader, loss_fn, device)
pbar.set_description(f"Training")
pbar.set_postfix(loss=avg_loss, accuracy=acc)
acc, avg_loss = test_loop(new_model, test_dataloader, loss_fn, device)
print(f"\nDone! Accuraccy: {acc:.2%}, Loss: {avg_loss:f}")
|
train_toy_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mathematical model parameter fitting : a data science case study
# This Jupyter notebook is intended for new modelers to learn how to fit parameter values of their model to their data.
#
# However, if you're new to mathematical modeling this is the gist:
# # Introduction
# The type of mathematical modeling I'm referring to is a [system of ordinary differential equations](https://en.wikipedia.org/wiki/Ordinary_differential_equation). These models assess how a system (defined by non-linear interactions) changes over time. This type of modeling is used by the financial sector to predict [investment cycles](https://link.springer.com/article/10.1007/s10598-017-9371-4), by pharmaceutical companies [to predict if a preclinical drug will be effective](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5122624/), and by scientists trying to understand the susceptible-infection-recovery rates of [COVID-19](https://www.nature.com/articles/s41591-020-0883-7).
# ## Model definition
# For this notebook, we will be looking at a really simple model. My background is in biology, so this will be a very simple [gene regulatory network](https://en.wikipedia.org/wiki/Gene_regulatory_network) model:
# 
# where a signal activates ```Protein X``` with drives the transcription of ```Gene Y``` (shown by the black arrows). The gray dashed arrows indicate that both the protein and the gene are degraded over time. Note that there's a basal level of protein activation ($k_b$), and a [negative feedback loop](https://en.wikipedia.org/wiki/Negative_feedback) stemming from ```Gene Y``` supressing the ```Signal``` (at some rate $\beta$). The rates (hint, hint our parameters!) are defined next to each of the interactions. This model can be built from either the literature, your own hypotheses, or both!
# Now, we can write this model as a system of ordinary differential equations:
# $\frac{dX}{dt} = k_b + Signal - \beta * Y - d_1 * X$
#
#
# $\frac{dY}{dt} = s_1 * X - d_2*Y$
# Great! Now, let's say we recently collected some timeseries data for ```Gene Y``` that looks like this:
# 
# ## The problem: Can our model explain our data?
# Now we need to fit our model to the data, meaning that we need to find optimal values for our parameters within the model. Sometimes these parameter values are known from real world measurements (like the rate of transcription), but let's just say we don't know the values to any of these 5 parameters ($k_b, \beta, d_1, s_1, d_2$ - remember ```X``` and ```Y``` are our model species and we know our ```Signal```).
# Where do we start?
# If you tried plugging in random values it could take you a lifetime to get a good fit. And until you do get a good fit, you'll be wondering whether or not you can rule this model out.
# Let's use machine learning!
# ## Evolutionary Algorithm (EA):
# The EA is a genetic algorithm that hones in on an optimal values/conditions based off of an error function. In this way, it will plug in a handful of initial random guesses, simulate your model with those parameters, assess how well those simulations do in comparison to the data (determined by your own error function), and then select the best scoring parameter set.
#
# A diagram of of the EA works:
# 
# ## IO
# **Inputs**
# - Experimental time-series data
# - System of ordinary differential equations
# - Range for parameter search space
#
# **Output**
# - Model Simulations
# - Analysis
#
# This notebook creates synthetic data, defines a basic model, and has ranges for the parameters already. Feel free to play around with them yourself!
# # Housekeeping
# ## Import Packages
import matplotlib.pyplot as plt
import h5py
# ## Plotting settings
# +
# plt.style.use("seaborn-whitegrid")
plt.rcParams['figure.figsize'] = [6, 4]
plt.rcParams['axes.titlesize'] = 18
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['axes.grid'] = True
plt.rcParams['axes.grid.axis'] = 'y'
plt.rcParams['grid.linestyle'] = '-'
plt.rcParams['xtick.labelsize'] = 10
plt.rcParams['ytick.labelsize'] = 10
plt.rcParams['legend.fontsize'] = 10
green = '#228833'
blue = '#4477AA'
fig_folder = './Figs/'
# -
# ## Import modules
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.insert(1, './modules')
from EA import *
from model import *
import plotting
# -
# ## Model definition
# ### Model variables
# +
dt = 0.1
steps = 251
time = np.linspace(0, dt*steps, steps)
sig = 5
X = 0
Y = 0
inits = [X, Y]
B = 1
kb = 0.1
s1 = 2
d1 = .5
d2 = .5
params = [B, kb, s1, d1, d2]
# -
# ### Model definition
mod = model.Model(inits, time, model.m1)
sims = mod.run_simulation(params, sig)
plt.plot(time, sims[:,0], label = 'X')
plt.plot(time, sims[:,1], label = 'Y')
plt.xlabel('Time (min)')
plt.ylabel('Response')
plt.legend()
plt.show()
# ## Load experimental data
# ### Create synth data
# +
# synthetic data
def create_synth_data(species, interval, save_data):
synth_time = range(0,26,interval)
closest_idxs = [np.abs(time - t).argmin() for t in synth_time]
synth_data = np.zeros((species.shape[1], len(closest_idxs)))
for i, s in enumerate(species.T):
synth_data[i] = s[closest_idxs]
if save_data:
## TODO: add check if file exists
with h5py.File("./Data/"+save_data+".hdf5", "w") as f:
synth_data = f.create_dataset("response", data = synth_data)
synth_time = f.create_dataset("time", data = synth_time)
# return synth_time, synth_data
# -
create_synth_data(sims, 1, save_data='synth_1min')
create_synth_data(sims, 5, save_data='synth_5min')
# ### Load synthetic data
synth_data_1min = h5py.File('./Data/synth_1min.hdf5', 'r')
synth_data_5min = h5py.File('./Data/synth_5min.hdf5', 'r')
# Checking synthetic data
plt.plot(synth_data_1min['time'], synth_data_1min['response'][0], color=green, marker='o', linestyle="None", label='1 min')
plt.plot(synth_data_5min['time'], synth_data_5min['response'][0], color=blue, marker='^', linestyle="None", markersize=10, alpha=1, label='5 min')
plt.xlabel('Time (min)')
plt.ylabel('Response')
plt.legend()
plt.show()
# +
# plt.plot(synth_data_1min['time'], synth_data_1min['response'][0], color=green, marker='o', linestyle="None", label='1 min')
plt.plot(synth_data_5min['time'], synth_data_5min['response'][0], color=blue, marker='^', linestyle="None", markersize=10, alpha=1, label='5 min')
plt.xlabel('Time (min)')
plt.ylabel('Response')
plt.ylim(-.1,1.4)
plt.legend()
# plt.show()
plt.savefig('./img/synth_data.png', dpi=300,bbox_inches='tight')
# -
# # Parameter Fitting with the EA
# +
# Parameter ranges
minimums = [-2, -2, -2, -2, -2]
maximums = [2, 2, 2, 2, 2]
diff = np.asarray(maximums)-np.asarray(minimums) #for plotting
dims = (diff, minimums) #for plotting
labelnames = ['$B$','$k_b$','$s_1$','$d_1$','$d_2$']
species = [1]
sig = [5]
# -
# We'll use the synthetic data from ```Gene Y``` - 5 min data.
ea = EA(synth_data_5min, [1], [5], mod, minimums, maximums, 2000)
top_params, best_error = ea.run('./Simulation_data/')
synth_time = synth_data_5min['time']
synth_response = synth_data_5min['response']
# + [markdown] heading_collapsed=true
# ## Load in simulation data
# + hidden=true
ea_5min = plotting.get_sim_data('./Simulation_data', num_sims=5)
# -
# ## Check EA worked
# +
def sort_sims(top_params, best_error, gen=-1):
top_params_gen = np.array([top_params[i][gen] for i in range(len(top_params))])
best_error_gen = np.array([best_error[i][gen] for i in range(len(best_error))])
return top_params[np.argsort(best_error_gen)], best_error[np.argsort(best_error_gen)], top_params_gen[np.argsort(best_error_gen)], best_error_gen[np.argsort(best_error_gen)]
top_params_full, best_error_full, top_params_50g, best_error_50g = sort_sims(top_params, best_error)
# +
#plots the trajectory SSE over generations for the top 1% candidates
top1_error = best_error_full[:20]
for i in range(len(top1_error)):
mses = top1_error[i]
plt.plot(range(len(mses)), mses)
plt.xlabel('Generation')
plt.ylabel('SSE')
plt.show()
# -
# ## Check simulation runs!
# run simulations for the parameters sets in the last generation
sim_data = []
closest_idxs = [np.abs(time - t).argmin() for t in synth_time]
for params in top_params_50g:
sim_datum = mod.run_simulation(params, sig)
sim_data.append(sim_datum)
# +
fig, axs = plt.subplots(2, 5, figsize=(10, 8), sharey=True)
for datum, i in zip(sim_data[:5], range(5)):
axs[0,i].plot(time, datum[:,0], color=blue, label='X (simulated)')
axs[0,i].plot(time, datum[:,1], color=green, label='Y (simulated)')
axs[0,i].plot(synth_time, synth_response[0], color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
axs[0,i].plot(synth_time, synth_response[1], color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
for datum, i in zip(sim_data[5:10], range(5)):
axs[1,i].plot(time, datum[:,0], color=blue, label='X (simulated)')
axs[1,i].plot(time, datum[:,1], color=green, label='Y (simulated)')
axs[1,i].plot(synth_time, synth_response[0], color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
axs[1,i].plot(synth_time, synth_response[1], color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
plt.legend(loc='upper right', bbox_to_anchor=(1.25, -0.40),
fancybox=True, shadow=True, ncol=5)
plt.ylim(-2,5)
plt.show()
# +
#plot last mse (generation 50) vs. index
plt.plot(range(len(best_error_50g)), best_error_50g)
plt.xlabel('EA index')
plt.ylabel('SSE')
plt.show()
# +
# see how close you are to the actual parameter values
num_plt = 100
df_plt = pd.DataFrame(top_params_50g[:num_plt])
df_plt.columns = labelnames
df_plt_log = df_plt.apply(np.log10)
df_plt_log_M = df_plt_log.melt(var_name='param', value_name='vals')
synth_data_df = pd.DataFrame(var).T
synth_data_df.columns = labelnames
synth_data_df_log = synth_data_df.apply(np.log10)
plotting.plt_param_ranges(labelnames, dims, runs_sort, 100, synth_data=synth_data_df_log) #black dot is best param
# -
runs_sort[1][2]
# +
sse_params = []
for param in runs_sort[1]:
sse = [np.absolute(x-p) for x, p in zip(var,param)]
sse_params.append(sse)
# plt.plot(range(len(runs2)), sse_params)
plt.plot(range(len(runs))[:10], np.log(sse_params[:10]))
# plt.plot(range(len(runs))[:10], sse_params[:10])
plt.legend(labelnames)
# plt.plot(range(len(runs))[:5], np.log([(x-p)**2 for x, p in zip(var,var)]))
# -
# ## Runs to 5 min interval, both Y and X
runs4 = run([Y_data_synth], [X_data_synth],synth_time)
for i in range(len(runs)):
mses = runs4[i][0]
plt.plot(range(len(mses)), mses)
sim_data4 = []
closest_idxs = [np.abs(time - t).argmin() for t in synth_time]
runs_sort4 = sort_sims(runs4)
for param in runs_sort4[1]:
ss_inits = run_ss(model, inits, param)
sim_datum = run_experiment(model, ss_inits, time, param, sig)
# sim_datum2 = run_experiment(model, ss_inits, time, param, sig)
sim_data4.append(sim_datum)
plt.plot(range(len(runs)), runs_sort4[0])
fig, axs = plt.subplots(2, 5, figsize=(12, 6), sharey=True)
for datum, i in zip(sim_data4[:5], range(5)):
axs[0,i].plot(time, datum)
axs[0,i].plot(synth_time, Y_data_synth, color=green, marker='^', linestyle="None", alpha=.7)
axs[0,i].plot(synth_time, X_data_synth, color=blue, marker='o', linestyle="None", alpha=.3)
for datum, i in zip(sim_data4[5:], range(5)):
axs[1,i].plot(time, datum)
axs[1,i].plot(synth_time, Y_data_synth, color=green, marker='^', linestyle="None", alpha=.7)
axs[1,i].plot(synth_time, X_data_synth, color=blue, marker='o', linestyle="None", alpha=.3)
# +
num_plt = 100
df_plt = pd.DataFrame(runs_sort4[1][:num_plt])
df_plt.columns = labelnames
df_plt_log = df_plt.apply(np.log10)
df_plt_log_M = df_plt_log.melt(var_name='param', value_name='vals')
synth_data_df = pd.DataFrame(var).T
synth_data_df.columns = labelnames
synth_data_df_log = synth_data_df.apply(np.log10)
plt_param_ranges(labelnames, dims, runs_sort4, 10, synth_data=synth_data_df_log) #black dot is best param
# -
# ## Runs to 1 min interval, just Y
runs2 = run([Y_data_synth2], synth_time2)
# +
sim_data2 = []
runs_sort2 = sort_sims(runs2)
for param in runs_sort2[1]:
ss_inits = run_ss(model, inits, param)
sim_datum = run_experiment(model, ss_inits, time, param, sig)
sim_data2.append(sim_datum)
# -
plt.hist(runs_sort2[0])
for i in range(len(runs2)):
mses = runs2[i][0]
plt.plot(range(len(mses)), mses)
plt.savefig(fig_folder+'6B.jpg', dpi=300,bbox_inches='tight')
for i in range(len(runs2)):
mses = runs2[i][0]
if mses[-1] <= runs_sort2[0][10]:
plt.plot(range(len(mses)), mses)
plt.savefig(fig_folder+'6Bi.jpg', dpi=300,bbox_inches='tight')
# +
# use
# df_top_thetas_EA = pd.DataFrame(runs_sort2[1])
# df_top_thetas_EA.columns = labelnames
num_plt = 100
df_plt = pd.DataFrame(runs_sort2[1][:num_plt])
df_plt.columns = labelnames
df_plt_log = df_plt.apply(np.log10)
df_plt_log_M = df_plt_log.melt(var_name='param', value_name='vals')
synth_data_df = pd.DataFrame(var).T
synth_data_df.columns = labelnames
synth_data_df_log = synth_data_df.apply(np.log10)
plt_param_ranges(labelnames, dims, runs_sort2, 100, synth_data=synth_data_df_log, save_fig='6C.jpg') #black dot is best param
# +
num_plt = 10
df_plt = pd.DataFrame(runs_sort2[1][:num_plt])
df_plt.columns = labelnames
df_plt_log = df_plt.apply(np.log10)
df_plt_log_M = df_plt_log.melt(var_name='param', value_name='vals')
synth_data_df = pd.DataFrame(var).T
synth_data_df.columns = labelnames
synth_data_df_log = synth_data_df.apply(np.log10)
plt_param_ranges(labelnames, dims, runs_sort2, 10, synth_data=synth_data_df_log, save_fig='6Ci.jpg') #black dot is best param
# plt.savefig(fig_folder+'6Ci.jpg', dpi=300,bbox_inches='tight')
# -
plt.plot(range(len(runs2)), [runs2[i][0][-1] for i in range(len(runs2))])
plt.plot(range(len(runs2)), runs_sort2[0])
plt.plot(range(len(runs2[:10])), runs_sort2[0][:10])
for datum in sim_data2[11:15]:
plt.plot(time, datum[:,1])
plt.plot(synth_time2, Y_data_synth2, color='red', marker='^', linestyle="None", alpha=.3)
# pltplot(synth_time2, X_data_synth2, color='blue', marker='o', linestyle="None", alpha=.3)
# +
fig, axs = plt.subplots(2, 5, figsize=(8, 4), sharey=True)
for datum, i in zip(sim_data2[:5], range(5)):
axs[0,i].plot(time, datum[:,0], color=blue, label='X (simulated)')
axs[0,i].plot(time, datum[:,1], color=green, label='Y (simulated)')
axs[0,i].plot(synth_time2, Y_data_synth2, color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
axs[0,i].plot(synth_time2, X_data_synth2, color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
for datum, i in zip(sim_data2[5:10], range(5)):
axs[1,i].plot(time, datum[:,0], color=blue, label='X (simulated)')
axs[1,i].plot(time, datum[:,1], color=green, label='Y (simulated)')
axs[1,i].plot(synth_time2, X_data_synth2, color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
axs[1,i].plot(synth_time2, Y_data_synth2, color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
plt.legend(loc='upper right', bbox_to_anchor=(1.25, -0.40),
fancybox=True, shadow=True, ncol=5)
plt.savefig(fig_folder+'7B.jpg', dpi=300,bbox_inches='tight')
# +
# use
# df_top_thetas_EA = pd.DataFrame(runs_sort2[1])
# df_top_thetas_EA.columns = labelnames
num_plt = 100
df_plt = pd.DataFrame(runs_sort2[1][:num_plt])
df_plt.columns = labelnames
df_plt_log = df_plt.apply(np.log10)
df_plt_log_M = df_plt_log.melt(var_name='param', value_name='vals')
synth_data_df = pd.DataFrame(var).T
synth_data_df.columns = labelnames
synth_data_df_log = synth_data_df.apply(np.log10)
plt_param_ranges(labelnames, dims, runs_sort2, 10, synth_data=synth_data_df_log) #black dot is best param
# plt.savefig(fig_folder+'5B3.jpg', dpi=300,bbox_inches='tight')
# +
# use
sns.distplot(df_plt_log[labelnames[0]])
plt.xlim(-3,3)
# -
# use
sns.distplot(df_plt_log[labelnames[0]])
plt.xlim(-3,3)
# +
param_mses = []
for param in runs_sort2[1][:10]:
diff = np.sum(np.absolute(param-var))
param_mses.append(diff)
# 1/0
print(np.argmin(param_mses))
print(param_mses)
# +
# use
# x and y species' MSES
mses_YX = []
for param in runs_sort2[1]:
mses_YX.append(scorefxn2(param, [Y_data_synth2], [X_data_synth2], synth_time2, convert=False))
# plt.plot(range(len(runs2)), mses_YX)
plt.plot(range(len(runs2))[:10], mses_YX[:10])
print(np.argmin(mses_YX[:10]))
print(mses_YX[:10])
# +
# use
##sum of squared errors for each parameter set, log
sse_params = []
for param in runs_sort2[1]:
sse = [(x-p)**2 for x, p in zip(var,param)]
sse_params.append(sse)
# plt.plot(range(len(runs2)), sse_params)
plt.plot(range(len(runs2))[:10], np.log(sse_params))
# +
sse_params_arr = np.array(sse_params[:10])
N = 10
n1 = sse_params_arr[:,0]
n2 = sse_params_arr[:,1]
# menStd = (2, 3, 4, 1, 2)
# womenStd = (3, 5, 2, 3, 3)
ind = np.arange(N) # the x locations for the groups
width = 0.35 # the width of the bars: can also be len(x) sequence
p1 = plt.bar(ind, n1, width)
p2 = plt.bar(ind, n2, width,
bottom=n1)
plt.bar(ind, sse_params_arr[:,0], width)
for i in range(1,5):
n = sse_params_arr[:,i]
plt.bar(ind, n, width,
bottom=sse_params_arr[:,i-1])
plt.yscale('log')
# plt.ylabel('Scores')
# plt.title('Scores by group and gender')
# plt.xticks(ind, ('G1', 'G2', 'G3', 'G4', 'G5'))
# plt.yticks(np.arange(0, 81, 10))
# plt.legend((p1[0], p2[0]), ('Men', 'Women'))
plt.show()
# -
# ## Runs to 1 min interval, both Y and X
runs3 = run([Y_data_synth2], [X_data_synth2], synth_time2)
# +
sim_data3 = []
runs_sort3 = sort_sims(runs3)
for param in runs_sort3[1]:
ss_inits = run_ss(model, inits, param)
sim_datum = run_experiment(model, ss_inits, time, param, sig)
sim_data3.append(sim_datum)
# +
##sse / # datapoints <- incorrect
plt.plot(range(len(runs3)), runs_sort3[0]/len(Y_data_synth2)/2, color='green')
plt.plot(range(len(runs3)), runs_sort2[0]/len(Y_data_synth2), color='orange')
plt.plot(range(len(runs3)), runs_sort[0]/len(Y_data_synth), color='blue')
# -
fig, axs = plt.subplots(2, 5, figsize=(12, 6), sharey=True)
for datum, i in zip(sim_data3[:5], range(5)):
axs[0,i].plot(time, datum)
axs[0,i].plot(synth_time2, Y_data_synth2, color='red', marker='^', linestyle="None", alpha=.3)
axs[0,i].plot(synth_time2, X_data_synth2, color='blue', marker='o', linestyle="None", alpha=.3)
for datum, i in zip(sim_data3[5:10], range(5)):
axs[1,i].plot(time, datum)
axs[1,i].plot(synth_time2, Y_data_synth2, color='red', marker='^', linestyle="None", alpha=.3)
axs[1,i].plot(synth_time2, X_data_synth2, color='blue', marker='o', linestyle="None", alpha=.3)
# +
# use
# df_top_thetas_EA = pd.DataFrame(runs_sort2[1])
# df_top_thetas_EA.columns = labelnames
num_plt = 100
df_plt = pd.DataFrame(runs_sort3[1][:num_plt])
df_plt.columns = labelnames
df_plt_log = df_plt.apply(np.log10)
df_plt_log_M = df_plt_log.melt(var_name='param', value_name='vals')
synth_data_df = pd.DataFrame(var).T
synth_data_df.columns = labelnames
synth_data_df_log = synth_data_df.apply(np.log10)
plt_param_ranges(labelnames, dims, runs_sort3, 10, synth_data=synth_data_df_log) #black dot is best param
# +
error1 =[]
for param in runs_sort[1]:
error = scorefxn1(param, [Y_data_synth], synth_time)
error1.append(error)
error2 =[]
for param in runs_sort2[1]:
error = scorefxn1(param, [Y_data_synth2], synth_time2)
error2.append(error)
error3 =[]
for param in runs_sort3[1]:
error = scorefxn2(param, [Y_data_synth2],[X_data_synth2], synth_time2)
error3.append(error)
error4 =[]
for param in runs_sort4[1]:
error = scorefxn2(param, [Y_data_synth],[X_data_synth], synth_time)
error4.append(error)
# -
test = np.sort(error1)
print(test)
# +
#use
plt.plot(range(len(runs3)), np.log10(np.sort(error1)), color='#66ccee',label='5-min, Y') #5 min
plt.plot(range(len(runs3)), np.log10(np.sort(error4)/2), color='#ccbb44',label='5-min, XY') #10 min both YX
plt.plot(range(len(runs3)), np.log10(np.sort(error2)), color='#ee6677',label='1-min, Y')#1 min just Y
plt.plot(range(len(runs3)), np.log10(np.sort(error3)/2), color='#aa3377',label='1-min, XY') #1 min both YX
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.savefig(fig_folder+'8C.jpg', dpi=300,bbox_inches='tight')
# -
# # model no neg
def model2(inits, t, var, sig):
X, Y = inits
B, kb, s1, d1, d2 = var
dX = kb + sig - d1 * X
dY = s1 * X - d2*Y # * (Y_t-Y)
return dX, dY
runs_noneg = run([Y_data_synth], [X_data_synth],synth_time)
# +
sim_data_nn = []
runs_sort_nn = sort_sims(runs_noneg)
for param in runs_sort_nn[1]:
ss_inits = run_ss(model2, inits, param)
sim_datum = run_experiment(model2, ss_inits, time, param, sig)
sim_data_nn.append(sim_datum)
# -
plt.plot(range(len(runs_noneg)), runs_sort_nn[0], color='green')
plt.plot(range(len(runs_noneg))[:10], runs_sort_nn[0][:10], color='green')
# +
fig, axs = plt.subplots(2, 5, figsize=(8, 4), sharey=True)
for datum, i in zip(sim_data_nn[:5], range(5)):
axs[0,i].plot(time, datum[:,0], color=blue, label='X (simulated)')
axs[0,i].plot(time, datum[:,1], color=green, label='Y (simulated)')
axs[0,i].plot(synth_time, Y_data_synth, color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
axs[0,i].plot(synth_time, X_data_synth, color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
for datum, i in zip(sim_data_nn[5:10], range(5)):
axs[1,i].plot(time, datum[:,0], color=blue, label='X (simulated)')
axs[1,i].plot(time, datum[:,1], color=green, label='Y (simulated)')
axs[1,i].plot(synth_time, X_data_synth, color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
axs[1,i].plot(synth_time, Y_data_synth, color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
plt.legend(loc='upper right', bbox_to_anchor=(1.25, -0.40),
fancybox=True, shadow=True, ncol=5)
plt.savefig(fig_folder+'9.jpg', dpi=300,bbox_inches='tight')
# +
# use
# df_top_thetas_EA = pd.DataFrame(runs_sort2[1])
# df_top_thetas_EA.columns = labelnames
num_plt = 100
df_plt = pd.DataFrame(runs_sort_nn[1][:num_plt])
df_plt.columns = labelnames
df_plt_log = df_plt.apply(np.log10)
df_plt_log_M = df_plt_log.melt(var_name='param', value_name='vals')
synth_data_df = pd.DataFrame(var).T
synth_data_df.columns = labelnames
synth_data_df_log = synth_data_df.apply(np.log10)
plt_param_ranges(labelnames, dims, runs_sort_nn, 10, synth_data=synth_data_df_log) #black dot is best param
# +
fp = []
def find_fixed_points(r):
for x in range(r):
for y in range(r):
if ((f(x,y) == 0) and (g(x,y) == 0)):
fp.append((x,y))
print('The system has a fixed point in %s,%s' % (x,y))
return fp
find_fixed_points(5001)
# -
# # Model driven experimental design
# +
def model(inits, t, var, sig):
X, Y = inits
B, kb, s1, d1, d2 = var
d2 = 0
if t > 0:
sig = 5
if t > 10:
sig = 0
if t > 20:
sig = 5
dX = kb + sig - B * Y - d1 * X
dY = s1 * X - d2*Y # * (Y_t-Y)
return dX, dY
def model2(inits, t, var, sig):
X, Y = inits
B, kb, s1, d1, d2 = var
# d2 = 0
if t > 0:
sig = 5
if t > 10:
sig = 0
if t > 20:
sig = 5
dX = kb + sig - d1 * X
dY = s1 * X - d2*Y # * (Y_t-Y)
return dX, dY
# -
null_model_params = runs_sort_nn[1][0]
true_model_params = runs_sort4[1][0]
# +
ninits = run_ss(model, [0,0], param)
ndatum = run_experiment(model2, ninits, time, null_model_params, sig)
plt.plot(time, ndatum[:,0], color=blue, label='X')
plt.plot(time, ndatum[:,1], color=green, label='Y')
plt.plot(synth_time, X_data_synth, color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
plt.plot(synth_time, Y_data_synth, color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
plt.ylim(0, 6)
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
# plt.savefig(fig_folder+'101.jpg', dpi=300,bbox_inches='tight')
# +
trinits = run_ss(model, [0,0], param)
trdatum = run_experiment(model, trinits, time, true_model_params, sig)
plt.plot(time, trdatum[:,0], color=blue, label='X')
plt.plot(time, trdatum[:,1], color=green, label='Y')
plt.plot(synth_time, X_data_synth, color=blue, marker='o', linestyle="None", alpha=.3, label='X (synthetic data)')
plt.plot(synth_time, Y_data_synth, color=green, marker='^', linestyle="None", alpha=.3, label='Y (synthetic data)')
plt.ylim(0, 6)
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
# plt.savefig(fig_folder+'102.jpg', dpi=300,bbox_inches='tight')
# -
# ## Different inits
# +
tdatum = run_experiment(model, [10,0], time, true_model_params, sig)
# print(tdatum)
plt.plot(time, tdatum[:,0], color=blue, label='X')
plt.plot(time, tdatum[:,1], color=green, label='Y')
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
# plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# plt.savefig(fig_folder+'10ab.jpg', dpi=300,bbox_inches='tight')
# +
# tinits = run_ss(model2, [100,100], param)
tdatum = run_experiment(model2, [10,0], time, null_model_params, sig)
# print(tdatum)
plt.plot(time, tdatum[:,0], color=blue, label='X')
plt.plot(time, tdatum[:,1], color=green, label='Y')
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.savefig(fig_folder+'10a1.jpg', dpi=300,bbox_inches='tight')
# -
# ## Different input
# +
t2inits = run_ss(model, [0,0], param)
t2datum = run_experiment(model, t2inits, time, true_model_params, sig)
# print(tdatum)
plt.plot(time, t2datum[:,0], color=blue, label='X')
plt.plot(time, t2datum[:,1], color=green, label='Y')
# plt.ylim(-2, 6)
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
# plt.savefig(fig_folder+'10b1.jpg', dpi=300,bbox_inches='tight')
# +
t2inits = run_ss(model, [0,0], param)
t2datum = run_experiment(model2, t2inits, time, null_model_params, sig)
# print(tdatum)
plt.plot(time, t2datum[:,0], color=blue, label='X')
plt.plot(time, t2datum[:,1], color=green, label='Y')
# plt.ylim(-2, 6)
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
# plt.savefig(fig_folder+'10b2.jpg', dpi=300,bbox_inches='tight')
# -
# ## Different structures
# +
t2inits = run_ss(model, [0,0], param)
t2datum = run_experiment(model, t2inits, time, true_model_params, sig)
# print(tdatum)
plt.plot(time, t2datum[:,0], color=blue, label='X')
plt.plot(time, t2datum[:,1], color=green, label='Y')
# plt.ylim(-2, 6)
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
# plt.savefig(fig_folder+'10c1.jpg', dpi=300,bbox_inches='tight')
# +
t2inits = run_ss(model, [0,0], param)
t2datum = run_experiment(model2, t2inits, time, null_model_params, sig)
# print(tdatum)
plt.plot(time, t2datum[:,0], color=blue, label='X')
plt.plot(time, t2datum[:,1], color=green, label='Y')
# plt.ylim(-2, 6)
plt.grid(color='#606060', which='major', axis='y', linestyle='solid')
# plt.savefig(fig_folder+'10c2.jpg', dpi=300,bbox_inches='tight')
|
.ipynb_checkpoints/Insights Health Data Science Analysis Review-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with Numba
# This notebooks provides some examples of how to work with **Numba** and compare the speed-up with C++.
#
# From the **consav** package we will use the **runtools** module to control the behavior of **Numba**.
# **Links:**
#
# - [Supported Python features](https://numba.pydata.org/numba-doc/dev/reference/pysupported.html)
# - [Supported Numpy features](https://numba.pydata.org/numba-doc/dev/reference/numpysupported.html)
# # Decorating Python functions
# Imports and numba settings:
# +
import time
import numpy as np
from consav import runtools
runtools.write_numba_config(threads=8,threading_layer='tbb')
import numba as nb # must be imported after write_numba_config!
#nb.config.__dict__ # see all config options
# -
# ## Functions
# +
def test_standard(X,Y,Z,NX,NY):
# X is lenght NX
# Y is lenght NY
# Z is length NX
for i in range(NX):
for j in range(NY):
Z[i] += np.exp(np.log(X[i]*Y[j]))/(X[i]*Y[j])-1
@nb.njit(parallel=True)
def test(X,Y,Z,NX,NY):
for i in nb.prange(NX):
for j in range(NY):
Z[i] += np.exp(np.log(X[i]*Y[j]))/(X[i]*Y[j])-1
@nb.njit(parallel=True,fastmath=True)
def test_fast(X,Y,Z,NX,NY):
for i in nb.prange(NX):
for j in range(NY):
Z[i] += np.exp(np.log(X[i]*Y[j]))/(X[i]*Y[j])-1
# -
# ## Settings
# Choose settings and make random draws:
# +
# a. settings
NX = 100
NY = 20000
# b. random draws
np.random.seed(1998)
X = np.random.sample(NX)
Y = np.random.sample(NY)
Z = np.zeros(NX)
# -
# ## Examples
# +
tic = time.time()
test_standard(X,Y,Z,NX,NY)
toc = time.time()
print(f'numba {np.sum(Z):.8f} in {toc-tic:.1f} secs')
tic = time.time()
test(X,Y,Z,NX,NY)
toc = time.time()
print(f'numba {np.sum(Z):.8f} in {toc-tic:.1f} secs')
Z = np.zeros(NX)
tic = time.time()
test_fast(X,Y,Z,NX,NY)
toc = time.time()
print(f'numba (fastmath=true) {np.sum(Z):.8f} in {toc-tic:.1f} secs')
# -
# # Test parallization in Numba and C++
# Compile C++ function for comparison:
from consav import cpptools
#cpptools.compile('test_numba',compiler='vs',dllfilename='test_numba_vs')
cpptools.compile('cppfuncs/test_numba',compiler='intel',dllfilename='test_numba_intel')
# Run tests with different number of threads:
for threads in [8,4,2,1]:
print(f'threads = {threads}')
print(f' threading_layer = tbb')
runtools.write_numba_config(threads=threads,threading_layer='tbb')
# !python test_numba.py
print(f' threading_layer = omp')
runtools.write_numba_config(threads=threads,threading_layer='omp')
# !python test_numba.py
print('')
runtools.write_numba_config(threads=8,threading_layer='omp')
|
Numba and C++/Working with Numba.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Ic4_occAAiAT"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" id="ioaprt5q5US7"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" id="yCl0eTNH5RS3"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] id="ItXfxkxvosLH"
# # 电影评论文本分类
# + [markdown] id="hKY4XMc9o8iB"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://tensorflow.google.cn/tutorials/keras/text_classification"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />在 tensorFlow.google.cn 上查看</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/keras/text_classification.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />在 Google Colab 中运行</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/keras/text_classification.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />在 GitHub 上查看源代码</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/keras/text_classification.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载 notebook</a>
# </td>
# </table>
# + [markdown] id="GEe3i16tQPjo"
# Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
# [官方英文文档](https://tensorflow.google.cn/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
# [tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
# [<EMAIL> Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
# + [markdown] id="Eg62Pmz3o83v"
# 此笔记本(notebook)使用评论文本将影评分为*积极(positive)*或*消极(nagetive)*两类。这是一个*二元(binary)*或者二分类问题,一种重要且应用广泛的机器学习问题。
#
# 我们将使用来源于[网络电影数据库(Internet Movie Database)](https://www.imdb.com/)的 [IMDB 数据集(IMDB dataset)](https://tensorflow.google.cn/api_docs/python/tf/keras/datasets/imdb),其包含 50,000 条影评文本。从该数据集切割出的25,000条评论用作训练,另外 25,000 条用作测试。训练集与测试集是*平衡的(balanced)*,意味着它们包含相等数量的积极和消极评论。
#
# 此笔记本(notebook)使用了 [tf.keras](https://tensorflow.google.cn/guide/keras),它是一个 Tensorflow 中用于构建和训练模型的高级API。有关使用 `tf.keras` 进行文本分类的更高级教程,请参阅 [MLCC文本分类指南(MLCC Text Classification Guide)](https://developers.google.com/machine-learning/guides/text-classification/)。
# + id="2ew7HTbPpCJH"
import tensorflow as tf
from tensorflow import keras
import numpy as np
print(tf.__version__)
# + [markdown] id="iAsKG535pHep"
# ## 下载 IMDB 数据集
#
# IMDB 数据集已经打包在 Tensorflow 中。该数据集已经经过预处理,评论(单词序列)已经被转换为整数序列,其中每个整数表示字典中的特定单词。
#
# 以下代码将下载 IMDB 数据集到您的机器上(如果您已经下载过将从缓存中复制):
# + id="zXXx5Oc3pOmN"
imdb = keras.datasets.imdb
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
# + [markdown] id="odr-KlzO-lkL"
# 参数 `num_words=10000` 保留了训练数据中最常出现的 10,000 个单词。为了保持数据规模的可管理性,低频词将被丢弃。
#
# + [markdown] id="l50X3GfjpU4r"
# ## 探索数据
#
# 让我们花一点时间来了解数据格式。该数据集是经过预处理的:每个样本都是一个表示影评中词汇的整数数组。每个标签都是一个值为 0 或 1 的整数值,其中 0 代表消极评论,1 代表积极评论。
# + id="y8qCnve_-lkO"
print("Training entries: {}, labels: {}".format(len(train_data), len(train_labels)))
# + [markdown] id="RnKvHWW4-lkW"
# 评论文本被转换为整数值,其中每个整数代表词典中的一个单词。首条评论是这样的:
# + id="QtTS4kpEpjbi"
print(train_data[0])
# + [markdown] id="hIE4l_72x7DP"
# 电影评论可能具有不同的长度。以下代码显示了第一条和第二条评论的中单词数量。由于神经网络的输入必须是统一的长度,我们稍后需要解决这个问题。
# + id="X-6Ii9Pfx6Nr"
len(train_data[0]), len(train_data[1])
# + [markdown] id="4wJg2FiYpuoX"
# ### 将整数转换回单词
#
# 了解如何将整数转换回文本对您可能是有帮助的。这里我们将创建一个辅助函数来查询一个包含了整数到字符串映射的字典对象:
# + id="tr5s_1alpzop"
# 一个映射单词到整数索引的词典
word_index = imdb.get_word_index()
# 保留第一个索引
word_index = {k:(v+3) for k,v in word_index.items()}
word_index["<PAD>"] = 0
word_index["<START>"] = 1
word_index["<UNK>"] = 2 # unknown
word_index["<UNUSED>"] = 3
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
# + [markdown] id="U3CNRvEZVppl"
# 现在我们可以使用 `decode_review` 函数来显示首条评论的文本:
# + id="s_OqxmH6-lkn"
decode_review(train_data[0])
# + [markdown] id="lFP_XKVRp4_S"
# ## 准备数据
#
# 影评——即整数数组必须在输入神经网络之前转换为张量。这种转换可以通过以下两种方式来完成:
#
# * 将数组转换为表示单词出现与否的由 0 和 1 组成的向量,类似于 one-hot 编码。例如,序列[3, 5]将转换为一个 10,000 维的向量,该向量除了索引为 3 和 5 的位置是 1 以外,其他都为 0。然后,将其作为网络的首层——一个可以处理浮点型向量数据的稠密层。不过,这种方法需要大量的内存,需要一个大小为 `num_words * num_reviews` 的矩阵。
#
# * 或者,我们可以填充数组来保证输入数据具有相同的长度,然后创建一个大小为 `max_length * num_reviews` 的整型张量。我们可以使用能够处理此形状数据的嵌入层作为网络中的第一层。
#
# 在本教程中,我们将使用第二种方法。
#
# 由于电影评论长度必须相同,我们将使用 [pad_sequences](https://tensorflow.google.cn/api_docs/python/tf/keras/preprocessing/sequence/pad_sequences) 函数来使长度标准化:
# + id="2jQv-omsHurp"
train_data = keras.preprocessing.sequence.pad_sequences(train_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
test_data = keras.preprocessing.sequence.pad_sequences(test_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
# + [markdown] id="VO5MBpyQdipD"
# 现在让我们看下样本的长度:
# + id="USSSBnkE-lky"
len(train_data[0]), len(train_data[1])
# + [markdown] id="QJoxZGyfjT5V"
# 并检查一下首条评论(当前已经填充):
# + id="TG8X9cqi-lk9"
print(train_data[0])
# + [markdown] id="LLC02j2g-llC"
# ## 构建模型
#
# 神经网络由堆叠的层来构建,这需要从两个主要方面来进行体系结构决策:
#
# * 模型里有多少层?
# * 每个层里有多少*隐层单元(hidden units)*?
#
# 在此样本中,输入数据包含一个单词索引的数组。要预测的标签为 0 或 1。让我们来为该问题构建一个模型:
# + id="xpKOoWgu-llD"
# 输入形状是用于电影评论的词汇数目(10,000 词)
vocab_size = 10000
model = keras.Sequential()
model.add(keras.layers.Embedding(vocab_size, 16))
model.add(keras.layers.GlobalAveragePooling1D())
model.add(keras.layers.Dense(16, activation='relu'))
model.add(keras.layers.Dense(1, activation='sigmoid'))
model.summary()
# + [markdown] id="6PbKQ6mucuKL"
# 层按顺序堆叠以构建分类器:
#
# 1. 第一层是`嵌入(Embedding)`层。该层采用整数编码的词汇表,并查找每个词索引的嵌入向量(embedding vector)。这些向量是通过模型训练学习到的。向量向输出数组增加了一个维度。得到的维度为:`(batch, sequence, embedding)`。
# 2. 接下来,`GlobalAveragePooling1D` 将通过对序列维度求平均值来为每个样本返回一个定长输出向量。这允许模型以尽可能最简单的方式处理变长输入。
# 3. 该定长输出向量通过一个有 16 个隐层单元的全连接(`Dense`)层传输。
# 4. 最后一层与单个输出结点密集连接。使用 `Sigmoid` 激活函数,其函数值为介于 0 与 1 之间的浮点数,表示概率或置信度。
# + [markdown] id="0XMwnDOp-llH"
# ### 隐层单元
#
# 上述模型在输入输出之间有两个中间层或“隐藏层”。输出(单元,结点或神经元)的数量即为层表示空间的维度。换句话说,是学习内部表示时网络所允许的自由度。
#
# 如果模型具有更多的隐层单元(更高维度的表示空间)和/或更多层,则可以学习到更复杂的表示。但是,这会使网络的计算成本更高,并且可能导致学习到不需要的模式——一些能够在训练数据上而不是测试数据上改善性能的模式。这被称为*过拟合(overfitting)*,我们稍后会对此进行探究。
# + [markdown] id="L4EqVWg4-llM"
# ### 损失函数与优化器
#
# 一个模型需要损失函数和优化器来进行训练。由于这是一个二分类问题且模型输出概率值(一个使用 sigmoid 激活函数的单一单元层),我们将使用 `binary_crossentropy` 损失函数。
#
# 这不是损失函数的唯一选择,例如,您可以选择 `mean_squared_error` 。但是,一般来说 `binary_crossentropy` 更适合处理概率——它能够度量概率分布之间的“距离”,或者在我们的示例中,指的是度量 ground-truth 分布与预测值之间的“距离”。
#
# 稍后,当我们研究回归问题(例如,预测房价)时,我们将介绍如何使用另一种叫做均方误差的损失函数。
#
# 现在,配置模型来使用优化器和损失函数:
# + id="Mr0GP-cQ-llN"
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
# + [markdown] id="hCWYwkug-llQ"
# ## 创建一个验证集
#
# 在训练时,我们想要检查模型在未见过的数据上的准确率(accuracy)。通过从原始训练数据中分离 10,000 个样本来创建一个*验证集*。(为什么现在不使用测试集?我们的目标是只使用训练数据来开发和调整模型,然后只使用一次测试数据来评估准确率(accuracy))。
# + id="-NpcXY9--llS"
x_val = train_data[:10000]
partial_x_train = train_data[10000:]
y_val = train_labels[:10000]
partial_y_train = train_labels[10000:]
# + [markdown] id="35jv_fzP-llU"
# ## 训练模型
#
# 以 512 个样本的 mini-batch 大小迭代 40 个 epoch 来训练模型。这是指对 `x_train` 和 `y_train` 张量中所有样本的的 40 次迭代。在训练过程中,监测来自验证集的 10,000 个样本上的损失值(loss)和准确率(accuracy):
# + id="D6G9oqEV-Se-"
history = model.fit(partial_x_train,
partial_y_train,
epochs=40,
batch_size=512,
validation_data=(x_val, y_val),
verbose=1)
# + [markdown] id="9EEGuDVuzb5r"
# ## 评估模型
#
# 我们来看一下模型的性能如何。将返回两个值。损失值(loss)(一个表示误差的数字,值越低越好)与准确率(accuracy)。
# + id="zOMKywn4zReN"
results = model.evaluate(test_data, test_labels, verbose=2)
print(results)
# + [markdown] id="z1iEXVTR0Z2t"
# 这种十分朴素的方法得到了约 87% 的准确率(accuracy)。若采用更好的方法,模型的准确率应当接近 95%。
# + [markdown] id="5KggXVeL-llZ"
# ## 创建一个准确率(accuracy)和损失值(loss)随时间变化的图表
#
# `model.fit()` 返回一个 `History` 对象,该对象包含一个字典,其中包含训练阶段所发生的一切事件:
# + id="VcvSXvhp-llb"
history_dict = history.history
history_dict.keys()
# + [markdown] id="nRKsqL40-lle"
# 有四个条目:在训练和验证期间,每个条目对应一个监控指标。我们可以使用这些条目来绘制训练与验证过程的损失值(loss)和准确率(accuracy),以便进行比较。
# + id="nGoYf2Js-lle"
import matplotlib.pyplot as plt
acc = history_dict['accuracy']
val_acc = history_dict['val_accuracy']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
# “bo”代表 "蓝点"
plt.plot(epochs, loss, 'bo', label='Training loss')
# b代表“蓝色实线”
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + id="6hXx-xOv-llh"
plt.clf() # 清除数字
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# + [markdown] id="oFEmZ5zq-llk"
# 在该图中,点代表训练损失值(loss)与准确率(accuracy),实线代表验证损失值(loss)与准确率(accuracy)。
#
# 注意训练损失值随每一个 epoch *下降*而训练准确率(accuracy)随每一个 epoch *上升*。这在使用梯度下降优化时是可预期的——理应在每次迭代中最小化期望值。
#
# 验证过程的损失值(loss)与准确率(accuracy)的情况却并非如此——它们似乎在 20 个 epoch 后达到峰值。这是过拟合的一个实例:模型在训练数据上的表现比在以前从未见过的数据上的表现要更好。在此之后,模型过度优化并学习*特定*于训练数据的表示,而不能够*泛化*到测试数据。
#
# 对于这种特殊情况,我们可以通过在 20 个左右的 epoch 后停止训练来避免过拟合。稍后,您将看到如何通过回调自动执行此操作。
|
src/Basic_knowledge/.ipynb_checkpoints/keras_text_classification-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# load MDAnalysis library
# to deal with GRO/XTC files
import MDAnalysis as mda
# load nglview for visualization
# of molecular systems
import nglview as nv
# -
# Biomolecular system
# -------------------------
#
# In this exmaple we are going to analyze two MD simulations of **CLC**, a bacterial Cl$^-$/H$^+$ antiporter, embedded in a bilayer composed of **POPC** and **DLPC**. Simulations were run on *Anton 2* for about 15 $\mu$s each. The files included in the ``../datasets/trajectories/`` directory contain only protein and lipids heavy atoms, all other elements, like water and ions, were removed beforehand.
# +
# define path to structure files
PATHD = "../datasets/trajectories/"
FRAME = PATHD + "CLCec1_anton2_restrained_15us.gro"
# load just first frame for visualization
# you could add TRAJD to see the MD trajectory,
# but you should fit the trajectory right after
u = mda.Universe(FRAME)
# -
# In total each trajectory file containts 300 frames, which correspond to a stride of $\sim$50 ns.
print("The total number of atoms is %s," % u.atoms.n_atoms)
print("from which %s are part of protein," % u.select_atoms("protein").n_atoms)
print("and %s belong to lipids." % u.select_atoms("not protein").n_atoms)
# Visualizing the protein
# --------------------------
#
# Let's now check our protein. As you can see this is an integral membrane protein composed mainly of alpha-helixes. This protein is found mainly as a homo-dimer, even in very diluted conditions, 1 protein subunit per $\sim$10$^7$ lipids. The region shown in blue indicates the dimerization interface.
# +
# select atoms
protein = u.select_atoms("protein")
# nglview widget
view = nv.show_mdanalysis(protein)
# visualization options
view.clear_representations()
view.add_cartoon(selection="protein", colorScheme="sstruc")
view.add_cartoon(selection="372-410.CA or 160-202.CA")
view._remote_call("setSize", target="Widget", args=["1000px", "500px"])
view.camera = 'orthographic'
view
# -
# The whole package
# -----------------------
#
# The motivation for running these MD simulations was to study the preferential organization of lipids around the dimerization interface. In this tutorial we will investigate this relation. Here you can see a snippet of the protein-membrane system simulated using an all-atom potential.
# +
# select molecules: protein + membrane
protmemb = u.select_atoms('resname POPC DLPC',
'protein', updating=True)
# Jupyter widget
view = nv.show_mdanalysis(protmemb)
# visualization options
view.clear_representations()
view.add_licorice(selection="not protein")
view.add_cartoon(selection="protein", colorScheme="sstruc")
view._remote_call("setSize", target="Widget", args=["1000px", "600px"])
view.camera = 'orthographic'
view.center(selection='protein')
view
# -
|
examples/visualize_datasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Cheat Sheet
#
# Basic cheatsheet for Python mostly based on the book written by <NAME>, [Automate the Boring Stuff with Python](https://automatetheboringstuff.com/) under the [Creative Commons license](https://creativecommons.org/licenses/by-nc-sa/3.0/) and many other sources.
#
# ## Read It
#
# - [Website](https://www.pythoncheatsheet.org)
# - [Github](https://github.com/wilfredinni/python-cheatsheet)
# - [PDF](https://github.com/wilfredinni/Python-cheatsheet/raw/master/python_cheat_sheet.pdf)
# - [Jupyter Notebook](https://mybinder.org/v2/gh/wilfredinni/python-cheatsheet/master?filepath=jupyter_notebooks)
#
# ## Manipulating Strings
#
# ### Escape Characters
#
# | Escape character | Prints as |
# | ---------------- | -------------------- |
# | `\'` | Single quote |
# | `\"` | Double quote |
# | `\t` | Tab |
# | `\n` | Newline (line break) |
# | `\\` | Backslash |
#
# Example:
print("Hello there!\nHow are you?\nI\'m doing fine.")
Hello there!
How are you?
# ### Raw Strings
#
# A raw string completely ignores all escape characters and prints any backslash that appears in the string.
print(r'That is Carol\'s cat.')
# Note: mostly used for regular expression definition (see `re` package)
#
# ### Multiline Strings with Triple Quotes
print('''Dear Alice,
Eve's cat has been arrested for catnapping, cat burglary, and extortion.
Sincerely,
Bob''')
# To keep a nicer flow in your code, you can use the `dedent` function from the `textwrap` standard package.
# +
from textwrap import dedent
def my_function():
print('''
Dear Alice,
Eve's cat has been arrested for catnapping, cat burglary, and extortion.
Sincerely,
Bob
''').strip()
# -
# This generates the same string than before.
#
# ### Indexing and Slicing Strings
H e l l o w o r l d !
0 1 2 3 4 5 6 7 8 9 10 11
spam = 'Hello world!'
spam[0]
spam[4]
spam[-1]
# Slicing:
# +
spam[0:5]
# -
spam[:5]
spam[6:]
spam[6:-1]
spam[:-1]
spam[::-1]
spam = 'Hello world!'
fizz = spam[0:5]
fizz
# ### The in and not in Operators with Strings
'Hello' in 'Hello World'
'Hello' in 'Hello'
'HELLO' in 'Hello World'
'' in 'spam'
'cats' not in 'cats and dogs'
# ### The in and not in Operators with list
a = [1, 2, 3, 4]
5 in a
2 in a
# ### The upper, lower, isupper, and islower String Methods
#
# `upper()` and `lower()`:
spam = 'Hello world!'
spam = spam.upper()
spam
spam = spam.lower()
spam
# isupper() and islower():
spam = 'Hello world!'
spam.islower()
spam.isupper()
'HELLO'.isupper()
'abc12345'.islower()
'12345'.islower()
'12345'.isupper()
# ### The isX String Methods
#
# - **isalpha()** returns True if the string consists only of letters and is not blank.
# - **isalnum()** returns True if the string consists only of lettersand numbers and is not blank.
# - **isdecimal()** returns True if the string consists only ofnumeric characters and is not blank.
# - **isspace()** returns True if the string consists only of spaces,tabs, and new-lines and is not blank.
# - **istitle()** returns True if the string consists only of wordsthat begin with an uppercase letter followed by onlylowercase letters.
#
# ### The startswith and endswith String Methods
'Hello world!'.startswith('Hello')
'Hello world!'.endswith('world!')
'abc123'.startswith('abcdef')
'abc123'.endswith('12')
'Hello world!'.startswith('Hello world!')
'Hello world!'.endswith('Hello world!')
# ### The join and split String Methods
#
# join():
', '.join(['cats', 'rats', 'bats'])
' '.join(['My', 'name', 'is', 'Simon'])
'ABC'.join(['My', 'name', 'is', 'Simon'])
# split():
'My name is Simon'.split()
'MyABCnameABCisABCSimon'.split('ABC')
'My name is Simon'.split('m')
# ### Justifying Text with rjust, ljust, and center
#
# rjust() and ljust():
'Hello'.rjust(10)
'Hello'.rjust(20)
'Hello World'.rjust(20)
'Hello'.ljust(10)
# An optional second argument to rjust() and ljust() will specify a fill character other than a space character. Enter the following into the interactive shell:
'Hello'.rjust(20, '*')
'Hello'.ljust(20, '-')
# center():
'Hello'.center(20)
'Hello'.center(20, '=')
# ### Removing Whitespace with strip, rstrip, and lstrip
spam = ' Hello World '
spam.strip()
spam.lstrip()
spam.rstrip()
spam = 'SpamSpamBaconSpamEggsSpamSpam'
spam.strip('ampS')
# ### Copying and Pasting Strings with the pyperclip Module
#
# First, install `pypeerclip` with pip:
# + attributes={"classes": ["shell"], "id": ""}
pip install pyperclip
# +
import pyperclip
pyperclip.copy('Hello world!')
pyperclip.paste()
|
jupyter_notebooks/10_Manipulating_Strings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] delitable=false
# # Lab 2 (Part C) - Linear regression with multiple features
#
# <div class="alert alert-block alert-danger">
#
# __IMPORTANT__
# Please complete this Jupyter Notebook file and upload it to blackboard __before 05 February 2020__.
# </div>
#
# In this part of the lab, you will implement linear regression with multiple variables to predict the price of houses. Suppose you are selling your house and you want to know what a good market price would be. One way to do this is to first collect information on recent houses sold and make a model of housing prices.
#
# # 1. Loading the dataset
# The file `housing-dataset.csv` contains a training set of housing prices in Portland, Oregon. The first column is the size of the house (in square feet), the second column is the number of bedrooms, and the third column is the price of the house. The following Python code helps you load the dataset from the data file into the variables $X$ and $y$. Read the code and print a small subset of $X$ and $y$ to see what they look like.
# +
# %matplotlib inline
import numpy as np
filename = "datasets/housing-dataset.csv"
mydata = np.genfromtxt(filename, delimiter=",")
# We have n data-points (houses)
n = len(mydata)
# X is a matrix of two column, i.e. an array of n 2-dimensional data-points
X = mydata[:, :2].reshape(n, 2)
# y is the vector of outputs, i.e. an array of n scalar values
y = mydata[:, -1]
""" TODO:
You can print a small subset of X and y to see what it looks like.
"""
print(X[:10])
print(y[:10])
# -
# # 2. Data normalization
# By looking at the values, note that house sizes are about 1000 times the number of bedrooms. When features differ by orders of magnitude, first performing feature scaling can make gradient descent converge much more quickly. Your task here is to write the following code to:
# - Subtract the mean value of each feature from the dataset.
# - After subtracting the mean, additionally scale (divide) the feature values by their respective *standard deviations*.
#
# In Python, you can use the numpy function `np.mean(..)` to compute the mean. This function can directly be used on a $d$-dimensional dataset to compute a $d$-dimensional mean vector `mu` where each value `mu[j]` is the mean of the $j^{th}$ feature. This is done by setting the $2^{nd}$ argument `axis` of this function to `0`. For example, consider the following matrix `A` where each line corresponds to one data-point and each column corresponds to one feature:
#
# ```python
# A = [[ 100, 10],
# [ 30, 10],
# [ 230, 25]]
# ```
#
# In this case, `np.mean(A, axis=0)` will give `[120, 15]` where 120 is the mean of the 1st column (1st feature) and 15 is the mean of the 2nd column (2nd feature). Another function `np.std(..)` exists to compute the standard deviation. The standard deviation is a way of measuring how much variation there is in the range of values of a particular feature (usually, most data points will lie within the interval: mean $\pm$ 2 standard_deviation).
#
# Once the features are normalized, you can do a scatter plot of the original dataset `X` (size of the house vs. number of bedrooms) and a scatter plot of the normalized dataset `X_normalized`. You will notice that the normalized dataset still have the same shape as the original one; the difference is that the new feature values have a similar scale and are centred arround the origin.
#
# **Implementation Note**: When normalizing the features, it is important to store the values used for normalization (the mean and the standard deviation used for the computations). Indeed, after learning the parameters of a model, we often want to predict the prices of houses we have not seen before. Given a new $x$ value (living room area and number of bedrooms), we must first normalize $x$ using the mean and standard deviation that we had previously computed from the training set.
# +
import matplotlib.pylab as plt
""" TODO:
Complete the following code to compute a normalized version of X called: X_normalized
"""
# TODO: compute mu, the mean vector from X
mu = X.mean(axis=0)
# TODO: compute std, the standard deviation vector from X
std = X.std(axis=0)
# X_normalized = (X - mu) / std
X_normalized = (X-mu)/std
""" TODO:
- Do a scatter plot of the original dataset X
- Do a scatter plot of the normalized dataset X_normalized
"""
fig, ax = plt.subplots()
ax.set_xlabel('Size')
ax.set_ylabel('Rooms')
ax.scatter(X[:,0],X[:,1], color="red", marker='o', label='Data points')
fig, ax = plt.subplots()
ax.set_xlabel('Size')
ax.set_ylabel('Rooms')
ax.scatter(X_normalized[:,0],X_normalized[:,1], color="red", marker='x', label='Data points')
# -
# Similar to what you did in Lab2 Part B, you can simplify your implementation of linear regression by adding an additional first column to `X_normalized` with all the values of this column set to $1$. To do this you can re-use the function `add_all_ones_column(..)` defined in Lab2 Part B, which takes a matrix as argument and returns a new matrix with an additional first column (of ones).
# +
""" TODO:
Copy-past here the definition of the function add_all_ones_column(...) that
you have see in Lab 2 (Part B).
"""
# definition of the function add_all_ones_column() here ...
def add_all_ones_column(X):
n, d = X.shape # dimension of the matrix X (n lines, d columns)
XX = np.ones((n, d+1)) # new matrix of all ones with one additional column
XX[:, 1:] = X # set X starting from column 1 (keep only column 0 unchanged)
return XX
""" TODO:
Just uncomment the following lines to create a matrix
X_normalized_new with an additional first column (of ones).
"""
X_normalized_new = add_all_ones_column(X_normalized)
print("Subset of X_normalized_new")
print(X_normalized_new[:10])
# -
# You are now ready to implement the linear regression using gradient descent (with more than one feature). In this multivariate case, you can further simply your implementation by writing the cost function in the following vectorized form:
#
# $$E(\theta) = \frac{1}{2n} (X \theta - y)^T (X \theta - y)$$
#
# $$\text{where }\quad
# X = \begin{bmatrix}
# -- ~ {x^{(1)}}^T ~ -- \\
# -- ~ {x^{(2)}}^T ~ -- \\
# \vdots \\
# -- ~ {x^{(n)}}^T ~ --
# \end{bmatrix}
# \quad \quad \quad
# y = \begin{bmatrix}
# y^{(1)} \\
# y^{(2)} \\
# \vdots \\
# y^{(n)}
# \end{bmatrix}
# $$
#
# The vectorized form of the gradient of $E(\theta)$ is a vector denoted as $\nabla E(\theta)$ and defined follows:
#
# $$\nabla E(\theta) = \left ( \frac{\partial E}{\partial \theta_0}, \frac{\partial E}{\partial \theta_1}, \dots, \frac{\partial E}{\partial \theta_d} \right ) = \frac{1}{n} X^T (X \theta - y)$$
#
# this is a **vector** where each $j^{th}$ value corresponds to $\frac{\partial E}{\partial \theta_j}$ (the derivative of the function $E$ with respect to the parameter $\theta_j$)
#
# One your code is finished, you will get to try out different learning rates $\alpha$ for the dataset and find a learning rate that converges quickly. To do so, you can plot the history of the cost $E(\theta)$ with respect to the number of iterations at the end of your code.
#
# For example for alpha values of 0.01, 0.05 and 0.1, the plot should look like follows:
# <img src="imgs/costLab2C.png" width="400px" />
#
# If your learning rate is too large, $E(\theta)$ can diverge and *blow up*, resulting in values which are too large for computer calculations. In these situations, Python will tend to return `NaN` or `inf` (NaN stands for "*not a number*" and is often caused by undefined operations that involve $-\inf$ and $+\inf$). If your value of $E(\theta)$ increases or even blows up, adjust your learning rate and try again.
# +
""" TODO:
Write the cost function E using the vectorized form
"""
def E(theta, X, y):
return (1/(2*len(X)))*np.transpose(X@theta - y)@(X@theta - y)
""" TODO:
Define the function grad_E (the gradient of E) using the vectorized form.
This should return a vector of the same dimension as theta
"""
def grad_E(theta, X, y):
return 1/len(X)*np.transpose(X)@(X@theta-y)
""" TODO:
Complete the definition of the function LinearRegressionWithGD(...) below
Note: don't forget to call the functions E(..) and grad_E(..) with X_normalized_new instead of X
The arguments of LinearRegressionWithGD(..) are:
*** theta: vector of initial parameter values
*** alpha: the learning rate (used by gradient descent)
*** max_iterations: maximum number of iterations to perform
*** epsilon: to stop iterating if the cost decreases by less than epsilon
The function returns:
*** errs: a list corresponding to the historical cost values
*** theta: the final parameter values
"""
def LinearRegressionWithGD(theta, alpha, max_iterations, epsilon):
errs = []
cost_list = []
for itr in range(max_iterations):
mse = E(theta, X_normalized_new, y)
errs.append(mse)
# TODO: take a gradient descent step to adapt the vector of parameters theta
theta = theta - alpha*grad_E(theta, X_normalized_new,y) # Vectorized Gradient descent
# TODO: test if the cost decreases by less than epsilon (to stop iterating)
CONDITION = mse - E(theta, X_normalized_new, y) < epsilon
if CONDITION:
break
return errs, theta
""" TODO:
Here you will call LinearRegressionWithGD(..) in a loop with different values of alpha,
and plot the cost history (errs) returned by each call of LinearRegressionWithGD(..)
"""
fig, ax = plt.subplots()
ax.set_xlabel("Number of Iterations")
ax.set_ylabel(r"Cost $E(\theta)$")
theta_init = np.array([0, 0, 0])
max_iterations = 100
epsilon = 0.000000000001
for alpha in [0.01, 0.05, 0.1]:
# TODO: call LinearRegressionWithGD(...) using the current alpha, to get errs and theta
errs, theta = LinearRegressionWithGD(theta_init, alpha, max_iterations, epsilon)
print("alpha = {}, theta = {}".format(alpha, theta))
# plot the errs using ax.plot(..)
ax.plot(errs)
plt.legend()
fig.show()
# -
# Now, once you have found a good $\theta$ using gradient descent, use it to make a price prediction for a new house of 1650-square-foot with 3 bedrooms. **Note**: since the parameter vector $\theta$ was learned using the normalized dataset, you will need to normalize the new data-point corresponding to this new house before predicting its price.
# +
""" TODO:
Use theta to predict the price of a 1650-square-foot house with 3 bedrooms
Don't forget to normalize the feature values of this new house first.
"""
# Create a data-point x corresponding to the new house
x = (np.array([[1650,3]]))
# Normalize the feature values of x
x_normalized = (x-mu)/std
x_normalized = add_all_ones_column(x_normalized)
# Use the vector of parameters theta to predict the price of x
predict1 = x_normalized @ theta
print("Prediction", predict1)
"""
HINT: if you are not able to compute the dot product between x and theta, then
make sure that the arrays have the same size. Did you forget something?
"""
# -
# # Normal Equation: Linear regression without gradient descent
#
# As you know from the lecture, the MSE cost function $E(\theta)$ that we are trying to minimize is a convex function, and its derivative at the optimal $\theta$ (that minimizes $E(\theta)$) is equal to $0$. Therefore, to find the optimal $\theta$, one can simply compute the derivative of $E(\theta)$ with respect to $\theta$, set it equal to $0$, and solve for $\theta$.
#
# We have seen in the lecture that, by doing this, the closed-form solution is given as follows:
# $$\theta = (X^T X)^{-1} X^T y$$
#
# Using this formula does not require any feature scaling, and you will get an exact solution in one calculation: there is no "*loop until convergence*" like in gradient descent.
#
# You are asked to implement this equation to directly compute the best parameter vector $\theta$ for the linear regression. In Python, you can use the `inv` function from `numpy.linalg.inv` to compute the inverse of a function.
#
# Remember that while you don't need to scale your features, we still need to add a column of 1's to the $X$ matrix to have an intercept term ($\theta_0$).
# +
from numpy.linalg import inv
""" TODO:
Use the function add_all_ones_column(..) to add a column of 1's to X.
Let's call the returned dataset X_new.
"""
new_X = add_all_ones_column(X)
""" TODO:
Compute the optimal theta using new_X and y (without using gradient descent).
Use the normal equation shown above. You can use the function inv (imported above)
to compute the inverse of a matrix.
"""
theta = np.linalg.inv(np.transpose(new_X)@new_X)@np.transpose(new_X)@y
print("With the original (non-normalized) dataset: theta = {}".format(theta))
# -
# Now, once you have computed the optimal $\theta$, use it to make a price prediction for the new house of 1650-square-foot with 3 bedrooms. Remeber that $\theta$ was computed above based on the original dataset (without normalization); so, you do not need to normalize the feature values of the new house to make the prediction in this case.
""" TODO:
Use theta to predict the price of a 1650-square-foot house with 3 bedrooms
"""
x = add_all_ones_column(np.array([[1650,3]]))
prediction = x @ theta
print(prediction)
# Using the previous formula does not require any feature normalization or scaling. However, you can still compute again the optimal $\theta$ when using `X_normalized_new` instead of `new_X`.
#
# By doing this, you will be able to compare the $\theta$ that you compute here with the one you got previously when you used gradient descent. The two parameter vectors should be quite similar (but not necessarily exatly the same).
""" TODO:
Compute the optimal theta using X_normalized_new and y (without using gradient descent).
Use the normal equation (shown previously).
"""
theta = np.linalg.inv(np.transpose(X_normalized_new)@X_normalized_new)@np.transpose(X_normalized_new)@y
print("With the normalized dataset: theta = {}".format(theta))
# Again, now that you have computed the optimal $\theta$ based on `X_normalized_new`, use it to make a price prediction for the new house of 1650-square-foot with 3 bedrooms. Do you need to normalize the feature values of the new house here? Remeber that $\theta$ was computed here based on the normalized dataset.
#
# You should find that this predicted price similar to the price you predicted previsouly for the same house.
""" TODO:
Use theta to predict the price of a 1650-square-foot house with 3 bedrooms
"""
# Cretate a data-point x corresponding to the new house
x = np.array([[1650,3]])
# Normalize the feature values of x
x_normalized = (x-mu)/std
x_normalized = add_all_ones_column(x_normalized)
predict1 = x_normalized @ theta
# Use the vector of parameters theta to predict the price of x
print("prediction:", predict1)
|
Labs/Lab2_Regression/Lab 2 (Part C) - Linear regression with multiple features.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python3
import pandas as pd
import lz4.frame
import gzip
import io
import pyarrow.parquet as pq
import pyarrow as pa
import numpy as np
from glob import glob
from plumbum.cmd import rm
from keras.layers.core import Dense, Activation, Dropout
from keras.layers.recurrent import LSTM
from keras.layers import TimeDistributed
from keras.models import Sequential
from keras import regularizers
from keras.callbacks import EarlyStopping
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
import matplotlib.pyplot as plt
import os.path
import pickle
import datetime
import re
from keras.models import model_from_json
# +
def plotline(data):
plt.figure()
plt.plot(data)
plt.legend()
plt.show()
def event_count(time_series, data_name):
time_series = time_series[['Fill Price (USD)']].values
upevents = 0
downevents = 0
sameprice = 0
prev_obv = time_series[0]
for obv in time_series[1:]:
if obv > prev_obv:
upevents += 1
elif obv < prev_obv:
downevents += 1
elif obv == prev_obv:
sameprice += 1
prev_obv = obv
print('=== Event counts on %s ===' % data_name)
print('upevents')
print(upevents)
print('downevents')
print(downevents)
print('sameprice')
print(sameprice)
print()
def mse(time_series, data_name):
time_series = time_series[['Fill Price (USD)']].values
total_squared_error = 0
total_absolute_error = 0
prev_obv = time_series[0]
for obv in time_series[1:]:
total_squared_error += (obv - prev_obv)**2
total_absolute_error += abs(obv - prev_obv)
prev_obv = obv
num_predictions = len(time_series) - 1
mean_squared_error = total_squared_error / num_predictions
mean_absolute_error = total_absolute_error / num_predictions
root_mean_squared_error = np.sqrt(mean_squared_error)
print('=== baseline on %s ===' % data_name)
print('total squared error')
print(total_squared_error)
print('total absolute error')
print(total_absolute_error)
print('mean squared error')
print(mean_squared_error)
print('mean absolute error')
print(mean_absolute_error)
print('root mean squared error')
print(root_mean_squared_error)
print()
def show_summary_statistics():
#event_count(small_set, 'small')
train_set = df.iloc[0:num_samples_training]
dev_set = df.iloc[num_samples_training:num_samples_training+num_samples_dev]
test_set = df.iloc[num_samples_training+num_samples_dev:]
event_count(train_set, 'train')
event_count(dev_set, 'dev')
event_count(test_set, 'test')
mse(train_set, 'train')
mse(dev_set, 'dev')
mse(test_set, 'test')
#show_summary_statistics()
# -
def load_data(path, day_start=None, day_end=None):
# Concatenate dataframes
files = sorted(glob('%s/*.parquet' % path))
if day_start is not None:
start = day_start
else:
start = 0
if day_end is not None:
end = day_start
else:
end = len(files)
files = files[start:end]
all_dataframes = []
for file in files:
df = pq.read_table(file).to_pandas()
all_dataframes.append(df)
df = pd.concat(all_dataframes)
return df
def split(df, temporal_features, percent_train):
X_train, X_dev, X_test, train_prices, dev_prices, test_prices = split_X(df, temporal_features, percent_train)
Y_train, Y_dev, Y_test = split_Y(df, percent_train)
return X_train, X_dev, X_test, Y_train, Y_dev, Y_test, train_prices, dev_prices, test_prices
def split_X(df, temporal_features, percent_train):
n_all = df.shape[0]
n_train = round(n_all * percent_train)
n_dev = round(n_all * ((1 - percent_train)/2))
n_test = round(n_all * ((1 - percent_train)/2))
print('n_all: ', n_all)
print('n_train:', n_train)
print('n_dev: ', n_dev)
print('n_test: ', n_test)
if temporal_features:
end = 59
else:
end = 16
X_train = df.iloc[:n_train, 1:end].values.astype('float32')
X_dev = df.iloc[n_train:n_train+n_dev, 1:end].values.astype('float32')
X_test = df.iloc[n_train+n_dev:, 1:end].values.astype('float32')
print(X_train.shape)
print(X_dev.shape)
print(X_test.shape)
train_prices = df.iloc[:n_train, 0].values.astype('float32')
dev_prices = df.iloc[n_train:n_train+n_dev, 0].values.astype('float32')
test_prices = df.iloc[n_train+n_dev:, 0].values.astype('float32')
return X_train, X_dev, X_test, train_prices, dev_prices, test_prices
def split_Y(df, percent_train):
n_all = df.shape[0]
n_train = round(n_all * percent_train)
n_dev = round(n_all * ((1 - percent_train)/2))
n_test = round(n_all * ((1 - percent_train)/2))
Y_train = df.iloc[:n_train, -1:].values.astype('float32')
Y_dev = df.iloc[n_train:n_train+n_dev, -1:].values.astype('float32')
Y_test = df.iloc[n_train+n_dev:, -1:].values.astype('float32')
print(Y_train.shape)
print(Y_dev.shape)
print(Y_test.shape)
return Y_train, Y_dev, Y_test
def create_end_of_window_Y(Y, window, step):
# Returns label at the end of a given window
return np.array([Y[i] for i in range(window-1, len(Y), step)])
def create_all_XY(X_train, X_dev, X_test, Y_train, Y_dev, Y_test, window_size, step, predict_end_of_window):
X_train = create_sequenced_data(X_train, window=window_size, step=step)
X_dev = create_sequenced_data(X_dev, window=window_size, step=step)
X_test = create_sequenced_data(X_test, window=window_size, step=step)
if predict_end_of_window:
Y_train = create_end_of_window_Y(Y_train, window=window_size, step=step)
Y_dev = create_end_of_window_Y(Y_dev, window=window_size, step=step)
Y_test = create_end_of_window_Y(Y_test, window=window_size, step=step)
else:
Y_train = create_sequenced_data(Y_train, window=window_size, step=step)
Y_dev = create_sequenced_data(Y_dev, window=window_size, step=step)
Y_test = create_sequenced_data(Y_test, window=window_size, step=step)
print('Train, dev, test shapes:')
print(X_train.shape)
print(X_dev.shape)
print(X_test.shape)
print(Y_train.shape)
print(Y_dev.shape)
print(Y_test.shape)
return X_train, X_dev, X_test, Y_train, Y_dev, Y_test
def create_sequenced_data(data, window, step):
sequenced = []
for minute in range(0, len(data) - window + 1, step):
chunk = data[minute:minute+window]
sequenced.append(chunk)
sequenced = np.array(sequenced)
return sequenced
def df_to_parquet(df, outfile):
pq.write_table(pa.Table.from_pandas(df), outfile, compression='snappy')
def direction_prediction(y_true, y_pred, predict_end_of_window):
if predict_end_of_window:
prop_correct = np.sum(np.sign(y_pred) == np.sign(y_true)) / y_true.shape[0]
else:
prop_correct = np.sum(np.sign(y_pred) == np.sign(y_true)) / (y_true.shape[0] * y_true.shape[1])
return prop_correct
def price_diffs(prices, y_hats):
# Predict within $x of actual price on average
price_diffs = []
for i in range(len(prices) - 1):
current_price = prices[i]
percent_change_pred = y_hats[i]
next_pred_price = current_price + (current_price * percent_change_pred)
next_true_price = prices[i + 1]
price_diffs.append(np.abs(next_pred_price - next_true_price))
return price_diffs
def initialize_model(X_train, loss, optimizer, num_LSTMs, num_units, dropout, predict_end_of_window):
LSTM_input_shape = [X_train.shape[1], X_train.shape[2]]
# DEFINE MODEL
model = Sequential()
if num_LSTMs == 2:
model.add(LSTM(num_units[0], input_shape=LSTM_input_shape, return_sequences=True))
model.add(Dropout(dropout))
if predict_end_of_window:
model.add(LSTM(num_units[1], return_sequences=False))
else:
model.add(LSTM(num_units[1], return_sequences=True))
if num_LSTMs == 3:
model.add(LSTM(num_units[0], input_shape=LSTM_input_shape, return_sequences=True))
model.add(Dropout(dropout))
model.add(LSTM(num_units[1], return_sequences=True))
model.add(Dropout(dropout))
if predict_end_of_window:
model.add(LSTM(num_units[2], return_sequences=False))
else:
model.add(LSTM(num_units[2], return_sequences=True))
if predict_end_of_window:
model.add(Dense(1))
else:
model.add(TimeDistributed(Dense(1)))
model.add(Activation('linear'))
model.compile(loss=loss, optimizer=optimizer)
return model
def evaluate_model(model, history, prices, X_train, X_dev, X_test, Y_train, Y_dev, Y_test,
train_prices, dev_prices, test_prices, predict_end_of_window):
train_loss = history.history['loss'][-1]
dev_loss = history.history['val_loss'][-1]
print('Evaluating test loss...')
test_loss = model.evaluate(X_test, Y_test, verbose=0)
print('Predicting y_hat_train...')
y_hat_train = model.predict(X_train)
print('Predicting y_hat_dev...')
y_hat_dev = model.predict(X_dev)
print('Predicting y_hat_test...')
y_hat_test = model.predict(X_test)
train_mse, train_rmse = mse_rmse(Y_train, y_hat_train)
dev_mse, dev_rmse = mse_rmse(Y_dev, y_hat_dev)
test_mse, test_rmse = mse_rmse(Y_test, y_hat_test)
train_prop_correct = direction_prediction(Y_train, y_hat_train, predict_end_of_window)
dev_prop_correct = direction_prediction(Y_dev, y_hat_dev, predict_end_of_window)
test_prop_correct = direction_prediction(Y_test, y_hat_test, predict_end_of_window)
price_diffs_train = price_diffs(train_prices, y_hat_train)
price_diffs_dev = price_diffs(dev_prices, y_hat_dev)
price_diffs_test = price_diffs(test_prices, y_hat_test)
evaluation = {'train_loss': train_loss,
'dev_loss': dev_loss,
'test_loss': test_loss,
'train_mse': train_mse,
'train_rmse': train_rmse,
'dev_mse': dev_mse,
'dev_rmse': dev_rmse,
'test_mse': test_mse,
'test_rmse': test_rmse,
'train_prop_correct': train_prop_correct,
'dev_prop_correct': dev_prop_correct,
'test_prop_correct': test_prop_correct,
'y_hat_train': y_hat_train,
'y_hat_dev': y_hat_dev,
'y_hat_test': y_hat_test}
return evaluation
def save_model_history(model, history, model_path):
# serialize model to JSON
if os.path.exists(model_path):
suffix = ''.join(re.findall(r'\d+', str(datetime.datetime.now())))
model_path = model_path + '_' + suffix
os.makedirs(model_path)
model_json = model.to_json()
with open(model_path + '/model.json', 'w') as json_file:
json_file.write(model_json)
try:
model.save_weights(model_path + '/model.h5')
except:
print('WARNING: Could not save weights...')
with open(model_path + '/trainHistoryDict', 'wb') as file_pi:
pickle.dump(history.history, file_pi)
print("Saved model and history to:\n%s" % model_path)
def plot_price(df, X_train, X_dev, field):
X_train_stop = len(X_train)
X_dev_stop = X_train_stop + len(X_dev)
plt.figure(figsize=(20,4))
plt.plot(np.arange(0, X_train_stop), df.iloc[0:X_train_stop][field], 'k')
plt.plot(np.arange(X_train_stop, X_dev_stop), df.iloc[X_train_stop:X_dev_stop][field], 'r')
plt.plot(np.arange(X_dev_stop, len(df)), df.iloc[X_dev_stop:len(df)][field], 'g')
def plot_train_dev_losses(history):
train_loss = history.history['loss']
dev_loss = history.history['val_loss']
plt.figure(figsize=(20,4))
plt.plot(np.log(train_loss), 'k')
plt.figure(figsize=(20,4))
plt.plot(np.log(dev_loss), 'b')
plt.figure(figsize=(20,4))
plt.plot(train_loss, 'k')
plt.figure(figsize=(20,4))
plt.plot(dev_loss, 'b')
plt.show()
def plot_percent_change(y_pred, y_true, timestep_within_window, minute_start, minute_end, predict_end_of_window):
ys=[]
for i in range(len(y_pred)):
ys.append(y_pred[i][timestep_within_window])
original_ys=[]
for i in range(len(y_true)):
original_ys.append(y_true[i][timestep_within_window])
ys_orig = np.array(original_ys)
ys_pred = np.array(ys)
OldRange = (ys_pred.max() - ys_pred.min())
NewRange = (ys_orig.max() - ys_orig.min())
new_ys_pred = (((ys - ys_pred.min()) * NewRange) / OldRange) + ys_orig.min()
norm1 = ys_orig / np.linalg.norm(ys_orig)
norm2 = ys_pred / np.linalg.norm(ys_pred)
plt.figure(figsize=(20,10))
plt.plot(norm1[minute_start:minute_end], 'k', alpha=0.9)
plt.plot(norm2[minute_start:minute_end], 'r', alpha=0.9)
plt.figure(figsize=(20,10))
plt.plot(original_ys[minute_start:minute_end], 'k', alpha=0.9)
plt.plot(ys[minute_start:minute_end], 'r', alpha=0.9)
plt.figure(figsize=(20,10))
plt.plot(original_ys[minute_start:minute_end], 'k', alpha=0.9)
plt.plot(new_ys_pred[minute_start:minute_end], 'r', alpha=0.9)
try:
plt.figure(figsize=(20,10))
plt.plot(y_true[minute_start:minute_end], 'k', alpha=0.9)
plt.plot(y_pred[minute_start:minute_end], 'r', alpha=0.9)
except:
a = 1
def print_save_events_props(train, dev, test, evaluate, model_name, model_path):
train_event_counts = np.unique(np.sign(train), return_counts=True)
train_event_prop = train_event_counts[1] / len(train)
dev_event_counts = np.unique(np.sign(dev), return_counts=True)
dev_event_prop = dev_event_counts[1] / len(dev)
test_event_counts = np.unique(np.sign(test), return_counts=True)
test_event_prop = test_event_counts[1] / len(test)
print(model_name)
print('\n========== EVENT COUNTS AND PROPORTIONS ==========')
print('=== TRAIN ===')
print('Down, Same, Up:', train_event_counts[1])
print('Down, Same, Up:', train_event_prop)
print('\n=== DEV ===')
print('Down, Same, Up:', dev_event_counts[1])
print('Down, Same, Up:', dev_event_prop)
print('\n=== TEST ===')
print('Down, Same, Up:', test_event_counts[1])
print('Down, Same, Up:', test_event_prop)
print('\n========== CORRECTION DIRECTION PREDICTIONS ==========')
print("TRAIN: %f\nDEV: %f\nTEST: %f" % (evaluate['train_prop_correct'],
evaluate['dev_prop_correct'],
evaluate['test_prop_correct']))
print('\n========== FINAL LOSS ==========')
print("TRAIN: %s\nDEV: %s\nTEST: %s\n" % (evaluate['train_loss'], evaluate['dev_loss'], evaluate['test_loss']))
def plot_losses(history, field, title):
vals = np.log(restored_history[field])
#vals = restored_history[field]
new_df = pd.DataFrame(vals, columns=[field])
new_df.plot(y = field, figsize=(7,6), title=title, fontsize=14, legend=False, color='firebrick')
plt.xlabel('Epoch', fontsize=18)
plt.title(title, fontsize=15, fontweight='bold')
def mse(time_series):
total_squared_error = 0
total_absolute_error = 0
prev_obv = time_series[0]
for obv in time_series[1:]:
total_squared_error += (obv - prev_obv)**2
total_absolute_error += abs(obv - prev_obv)
prev_obv = obv
num_predictions = len(time_series) - 1
mean_squared_error = total_squared_error / num_predictions
mean_absolute_error = total_absolute_error / num_predictions
root_mean_squared_error = np.sqrt(mean_squared_error)
print('=== baseline ===')
print('total squared error')
print(total_squared_error)
print('total absolute error')
print(total_absolute_error)
print('mean squared error')
print(mean_squared_error)
print('mean absolute error')
print(mean_absolute_error)
print('root mean squared error')
print(root_mean_squared_error)
print()
def mse_rmse(Y_true, Y_pred, verbose=False, baseline_desc=None):
MSE = np.sum((Y_true - Y_pred) ** 2) / len(Y_true)
RMSE = np.sqrt(MSE)
if verbose:
print('\n%s' % baseline_desc)
print('MSE: %06f' % MSE)
print('RMSE: %06f' % RMSE)
return MSE, RMSE
def baselines(Y, data_name):
print('\n========= %s =========' % data_name)
Y_true = np.array(Y[1:])
# Predict no percent change
Y_pred = np.zeros((Y_true.shape))
mse_rmse(Y_true, Y_pred, verbose=True, baseline_desc='Predict NO Percent Change')
# Predict same percent change
Y_pred = np.array(Y[:-1])
mse_rmse(Y_true, Y_pred, verbose=True, baseline_desc='Predict SAME Percent Change')
# +
##### MAIN MODEL #####
# HYPERPARAMETERS
window_size = 60
step = 1
predict_end_of_window = False
temporal_features = False
batch_size = 2048 #8192
num_epochs = 15
#num_epochs = 30
verbose = 1
loss = 'mean_squared_error'
optimizer = 'adam'
num_LSTM = 3
num_units = [128, 256, 256]
#num_LSTM = 2
#num_units = [256, 256]
dropout = 0.1
path = 'cboe/parquet_preprocessed_BTCUSD_merged'
day_start = 401
day_end = None
percent_train = 0.9
num_units_string = '-'.join([str(u) for u in num_units])
model_name = 'window-%s_step-%s_predEndWindow-%s_temporalFeat-%s_batch-%s_epochs-%s_loss-%s_opt-%s_numLSTMs-%s_numUnits-%s_dropout-%s_dayStart-%s_dayEnd-%s' % (window_size, step, str(predict_end_of_window), str(temporal_features), batch_size, num_epochs, loss, optimizer, num_LSTM, num_units_string, dropout, str(day_start), str(day_end))
print(model_name+'\n')
model_path = 'models/%s' % model_name
min_delta = 0.1
patience = 15
early_stop = EarlyStopping(monitor='val_loss', min_delta=min_delta, patience=patience)
callbacks_list = [early_stop]
# LOAD DATA
df = load_data(path, day_start, day_end)
# CREATE XY DATA
X_train, X_dev, X_test, Y_train, Y_dev, Y_test, train_prices, dev_prices, test_prices = split(df, temporal_features, percent_train)
X_train, X_dev, X_test, Y_train, Y_dev, Y_test = create_all_XY(X_train, X_dev, X_test, Y_train, Y_dev, Y_test,
window_size, step, predict_end_of_window)
plot_price(df, X_train, X_dev, field='current_price')
plot_price(df, X_train, X_dev, field='percent_change')
# +
# INITIALIZE MODEL
print('Initializing model...')
model = initialize_model(X_train, loss, optimizer, num_LSTM, num_units, dropout,
predict_end_of_window=predict_end_of_window)
# TRAIN MODEL
print('Training model...')
history = model.fit(X_train, Y_train, batch_size=batch_size, epochs=num_epochs,
validation_data=(X_dev, Y_dev), callbacks=callbacks_list, verbose=verbose, shuffle=True)
# SAVE MODEL AND HISTORY
save_model_history(model, history, model_path)
# +
# EVALUATE MODEL
print('Evaluating model...')
prices = df['current_price']
evaluate = evaluate_model(model, history, prices, X_train, X_dev, X_test, Y_train, Y_dev, Y_test, predict_end_of_window)
with open(model_path + '/evaluate.pkl', 'wb') as f:
pickle.dump(evaluate, f, pickle.HIGHEST_PROTOCOL)
# +
# VISUALIZE
## Plot:
# Historical price, color coded with train, dev, test
# Historical percent change, color coded with train, dev, test
# Train Loss, Dev Loss
# Actual price vs predicted price (or percent change) for test set
# Example features time series for one day (NOTE: in the preprocessing_final notebook)
if predict_end_of_window:
timestep_within_window = 0
else:
timestep_within_window = window-1
minute_start = 0
print_save_events_props(Y_train.flatten(), Y_dev.flatten(), Y_test.flatten(), evaluate, model_name, model_path)
plot_price(df, X_train, X_dev, field='current_price')
plot_price(df, X_train, X_dev, field='percent_change')
plot_losses(restored_history, field='loss', title='Training Loss')
plot_losses(restored_history, field='val_loss', title='Dev Loss')
minute_end = len(Y_train)
plot_percent_change(evaluate['y_hat_train'], Y_train, timestep_within_window, minute_start, minute_end, predict_end_of_window)
minute_end = len(Y_dev)
plot_percent_change(evaluate['y_hat_dev'], Y_dev, timestep_within_window, minute_start, minute_end, predict_end_of_window)
minute_end = len(Y_test)
plot_percent_change(evaluate['y_hat_test'], Y_test, timestep_within_window, minute_start, minute_end, predict_end_of_window)
# -
print('================== BASELINES ==================')
baselines(Y_train, data_name='Training Data')
baselines(Y_dev, data_name='Dev Data')
baselines(Y_test, data_name='Test Data')
|
training_testing_FINAL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training at scale with AI Platform Training Service
# **Learning Objectives:**
# 1. Learn how to organize your training code into a Python package
# 1. Train your model using cloud infrastructure via Google Cloud AI Platform Training Service
# 1. (optional) Learn how to run your training package using Docker containers and push training Docker images on a Docker registry
#
# ## Introduction
#
# In this notebook we'll make the jump from training locally, to do training in the cloud. We'll take advantage of Google Cloud's [AI Platform Training Service](https://cloud.google.com/ai-platform/).
#
# AI Platform Training Service is a managed service that allows the training and deployment of ML models without having to provision or maintain servers. The infrastructure is handled seamlessly by the managed service for us.
#
# Each learning objective will correspond to a __#TODO__ in the [student lab notebook](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive2/building_production_ml_systems/labs/1_training_at_scale.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
# Specify your project name and bucket name in the cell below.
# !sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# +
import os
from google.cloud import bigquery
# -
# Change the following cell as necessary:
# +
# Change with your own bucket and project below:
BUCKET = "<BUCKET>"
PROJECT = "<PROJECT>"
REGION = "<YOUR REGION>"
OUTDIR = "gs://{bucket}/taxifare/data".format(bucket=BUCKET)
os.environ['BUCKET'] = BUCKET
os.environ['OUTDIR'] = OUTDIR
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = "2.1"
# + language="bash"
# gcloud config set project $PROJECT
# gcloud config set compute/region $REGION
# -
# ## Create BigQuery tables
# If you have not already created a BigQuery dataset for our data, run the following cell:
# +
bq = bigquery.Client(project = PROJECT)
dataset = bigquery.Dataset(bq.dataset("taxifare"))
try:
bq.create_dataset(dataset)
print("Dataset created")
except:
print("Dataset already exists")
# -
# Let's create a table with 1 million examples.
#
# Note that the order of columns is exactly what was in our CSV files.
# +
# %%bigquery
CREATE OR REPLACE TABLE taxifare.feateng_training_data AS
SELECT
(tolls_amount + fare_amount) AS fare_amount,
pickup_datetime,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers,
'unused' AS key
FROM `nyc-tlc.yellow.trips`
WHERE ABS(MOD(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING)), 1000)) = 1
AND
trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
# -
# Make the validation dataset be 1/10 the size of the training dataset.
# +
# %%bigquery
CREATE OR REPLACE TABLE taxifare.feateng_valid_data AS
SELECT
(tolls_amount + fare_amount) AS fare_amount,
pickup_datetime,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers,
'unused' AS key
FROM `nyc-tlc.yellow.trips`
WHERE ABS(MOD(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING)), 10000)) = 2
AND
trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
# -
# ## Export the tables as CSV files
# + language="bash"
#
# echo "Deleting current contents of $OUTDIR"
# gsutil -m -q rm -rf $OUTDIR
#
# echo "Extracting training data to $OUTDIR"
# bq --location=US extract \
# --destination_format CSV \
# --field_delimiter "," --noprint_header \
# taxifare.feateng_training_data \
# $OUTDIR/taxi-train-*.csv
#
# echo "Extracting validation data to $OUTDIR"
# bq --location=US extract \
# --destination_format CSV \
# --field_delimiter "," --noprint_header \
# taxifare.feateng_valid_data \
# $OUTDIR/taxi-valid-*.csv
#
# gsutil ls -l $OUTDIR
# -
# !gsutil cat gs://$BUCKET/taxifare/data/taxi-train-000000000000.csv | head -2
# ## Make code compatible with AI Platform Training Service
# In order to make our code compatible with AI Platform Training Service we need to make the following changes:
#
# 1. Upload data to Google Cloud Storage
# 2. Move code into a trainer Python package
# 4. Submit training job with `gcloud` to train on AI Platform
# ### Upload data to Google Cloud Storage (GCS)
#
# Cloud services don't have access to our local files, so we need to upload them to a location the Cloud servers can read from. In this case we'll use GCS.
# !gsutil ls gs://$BUCKET/taxifare/data
# ### Move code into a python package
#
#
# The first thing to do is to convert your training code snippets into a regular Python package that we will then `pip install` into the Docker container.
#
# A Python package is simply a collection of one or more `.py` files along with an `__init__.py` file to identify the containing directory as a package. The `__init__.py` sometimes contains initialization code but for our purposes an empty file suffices.
# #### Create the package directory
# Our package directory contains 3 files:
# ls ./taxifare/trainer/
# #### Paste existing code into model.py
#
# A Python package requires our code to be in a .py file, as opposed to notebook cells. So, we simply copy and paste our existing code for the previous notebook into a single file.
# In the cell below, we write the contents of the cell into `model.py` packaging the model we
# developed in the previous labs so that we can deploy it to AI Platform Training Service.
# +
# %%writefile ./taxifare/trainer/model.py
import datetime
import logging
import os
import shutil
import numpy as np
import tensorflow as tf
from tensorflow.keras import activations
from tensorflow.keras import callbacks
from tensorflow.keras import layers
from tensorflow.keras import models
from tensorflow import feature_column as fc
logging.info(tf.version.VERSION)
CSV_COLUMNS = [
'fare_amount',
'pickup_datetime',
'pickup_longitude',
'pickup_latitude',
'dropoff_longitude',
'dropoff_latitude',
'passenger_count',
'key',
]
LABEL_COLUMN = 'fare_amount'
DEFAULTS = [[0.0], ['na'], [0.0], [0.0], [0.0], [0.0], [0.0], ['na']]
DAYS = ['Sun', 'Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat']
def features_and_labels(row_data):
for unwanted_col in ['key']:
row_data.pop(unwanted_col)
label = row_data.pop(LABEL_COLUMN)
return row_data, label
def load_dataset(pattern, batch_size, num_repeat):
dataset = tf.data.experimental.make_csv_dataset(
file_pattern=pattern,
batch_size=batch_size,
column_names=CSV_COLUMNS,
column_defaults=DEFAULTS,
num_epochs=num_repeat,
)
return dataset.map(features_and_labels)
def create_train_dataset(pattern, batch_size):
dataset = load_dataset(pattern, batch_size, num_repeat=None)
return dataset.prefetch(1)
def create_eval_dataset(pattern, batch_size):
dataset = load_dataset(pattern, batch_size, num_repeat=1)
return dataset.prefetch(1)
def parse_datetime(s):
if type(s) is not str:
s = s.numpy().decode('utf-8')
return datetime.datetime.strptime(s, "%Y-%m-%d %H:%M:%S %Z")
def euclidean(params):
lon1, lat1, lon2, lat2 = params
londiff = lon2 - lon1
latdiff = lat2 - lat1
return tf.sqrt(londiff*londiff + latdiff*latdiff)
def get_dayofweek(s):
ts = parse_datetime(s)
return DAYS[ts.weekday()]
@tf.function
def dayofweek(ts_in):
return tf.map_fn(
lambda s: tf.py_function(get_dayofweek, inp=[s], Tout=tf.string),
ts_in
)
@tf.function
def fare_thresh(x):
return 60 * activations.relu(x)
def transform(inputs, NUMERIC_COLS, STRING_COLS, nbuckets):
# Pass-through columns
transformed = inputs.copy()
del transformed['pickup_datetime']
feature_columns = {
colname: fc.numeric_column(colname)
for colname in NUMERIC_COLS
}
# Scaling longitude from range [-70, -78] to [0, 1]
for lon_col in ['pickup_longitude', 'dropoff_longitude']:
transformed[lon_col] = layers.Lambda(
lambda x: (x + 78)/8.0,
name='scale_{}'.format(lon_col)
)(inputs[lon_col])
# Scaling latitude from range [37, 45] to [0, 1]
for lat_col in ['pickup_latitude', 'dropoff_latitude']:
transformed[lat_col] = layers.Lambda(
lambda x: (x - 37)/8.0,
name='scale_{}'.format(lat_col)
)(inputs[lat_col])
# Adding Euclidean dist (no need to be accurate: NN will calibrate it)
transformed['euclidean'] = layers.Lambda(euclidean, name='euclidean')([
inputs['pickup_longitude'],
inputs['pickup_latitude'],
inputs['dropoff_longitude'],
inputs['dropoff_latitude']
])
feature_columns['euclidean'] = fc.numeric_column('euclidean')
# hour of day from timestamp of form '2010-02-08 09:17:00+00:00'
transformed['hourofday'] = layers.Lambda(
lambda x: tf.strings.to_number(
tf.strings.substr(x, 11, 2), out_type=tf.dtypes.int32),
name='hourofday'
)(inputs['pickup_datetime'])
feature_columns['hourofday'] = fc.indicator_column(
fc.categorical_column_with_identity(
'hourofday', num_buckets=24))
latbuckets = np.linspace(0, 1, nbuckets).tolist()
lonbuckets = np.linspace(0, 1, nbuckets).tolist()
b_plat = fc.bucketized_column(
feature_columns['pickup_latitude'], latbuckets)
b_dlat = fc.bucketized_column(
feature_columns['dropoff_latitude'], latbuckets)
b_plon = fc.bucketized_column(
feature_columns['pickup_longitude'], lonbuckets)
b_dlon = fc.bucketized_column(
feature_columns['dropoff_longitude'], lonbuckets)
ploc = fc.crossed_column(
[b_plat, b_plon], nbuckets * nbuckets)
dloc = fc.crossed_column(
[b_dlat, b_dlon], nbuckets * nbuckets)
pd_pair = fc.crossed_column([ploc, dloc], nbuckets ** 4)
feature_columns['pickup_and_dropoff'] = fc.embedding_column(
pd_pair, 100)
return transformed, feature_columns
def rmse(y_true, y_pred):
return tf.sqrt(tf.reduce_mean(tf.square(y_pred - y_true)))
def build_dnn_model(nbuckets, nnsize, lr):
# input layer is all float except for pickup_datetime which is a string
STRING_COLS = ['pickup_datetime']
NUMERIC_COLS = (
set(CSV_COLUMNS) - set([LABEL_COLUMN, 'key']) - set(STRING_COLS)
)
inputs = {
colname: layers.Input(name=colname, shape=(), dtype='float32')
for colname in NUMERIC_COLS
}
inputs.update({
colname: layers.Input(name=colname, shape=(), dtype='string')
for colname in STRING_COLS
})
# transforms
transformed, feature_columns = transform(
inputs, NUMERIC_COLS, STRING_COLS, nbuckets=nbuckets)
dnn_inputs = layers.DenseFeatures(feature_columns.values())(transformed)
x = dnn_inputs
for layer, nodes in enumerate(nnsize):
x = layers.Dense(nodes, activation='relu', name='h{}'.format(layer))(x)
output = layers.Dense(1, name='fare')(x)
model = models.Model(inputs, output)
#TODO 1a
lr_optimizer = tf.keras.optimizers.Adam(learning_rate=lr)
model.compile(optimizer=lr_optimizer, loss='mse', metrics=[rmse, 'mse'])
return model
def train_and_evaluate(hparams):
#TODO 1b
batch_size = hparams['batch_size']
nbuckets = hparams['nbuckets']
lr = hparams['lr']
nnsize = hparams['nnsize']
eval_data_path = hparams['eval_data_path']
num_evals = hparams['num_evals']
num_examples_to_train_on = hparams['num_examples_to_train_on']
output_dir = hparams['output_dir']
train_data_path = hparams['train_data_path']
timestamp = datetime.datetime.now().strftime('%Y%m%d%H%M%S')
savedmodel_dir = os.path.join(output_dir, 'export/savedmodel')
model_export_path = os.path.join(savedmodel_dir, timestamp)
checkpoint_path = os.path.join(output_dir, 'checkpoints')
tensorboard_path = os.path.join(output_dir, 'tensorboard')
if tf.io.gfile.exists(output_dir):
tf.io.gfile.rmtree(output_dir)
model = build_dnn_model(nbuckets, nnsize, lr)
logging.info(model.summary())
trainds = create_train_dataset(train_data_path, batch_size)
evalds = create_eval_dataset(eval_data_path, batch_size)
steps_per_epoch = num_examples_to_train_on // (batch_size * num_evals)
checkpoint_cb = callbacks.ModelCheckpoint(
checkpoint_path,
save_weights_only=True,
verbose=1
)
tensorboard_cb = callbacks.TensorBoard(tensorboard_path)
history = model.fit(
trainds,
validation_data=evalds,
epochs=num_evals,
steps_per_epoch=max(1, steps_per_epoch),
verbose=2, # 0=silent, 1=progress bar, 2=one line per epoch
callbacks=[checkpoint_cb, tensorboard_cb]
)
# Exporting the model with default serving function.
tf.saved_model.save(model, model_export_path)
return history
# -
# ### Modify code to read data from and write checkpoint files to GCS
#
# If you look closely above, you'll notice a new function, `train_and_evaluate` that wraps the code that actually trains the model. This allows us to parametrize the training by passing a dictionary of parameters to this function (e.g, `batch_size`, `num_examples_to_train_on`, `train_data_path` etc.)
#
# This is useful because the output directory, data paths and number of train steps will be different depending on whether we're training locally or in the cloud. Parametrizing allows us to use the same code for both.
#
# We specify these parameters at run time via the command line. Which means we need to add code to parse command line parameters and invoke `train_and_evaluate()` with those params. This is the job of the `task.py` file.
# +
# %%writefile taxifare/trainer/task.py
import argparse
from trainer import model
if __name__ == '__main__':
parser = argparse.ArgumentParser()
parser.add_argument(
"--batch_size",
help="Batch size for training steps",
type=int,
default=32
)
parser.add_argument(
"--eval_data_path",
help="GCS location pattern of eval files",
required=True
)
parser.add_argument(
"--nnsize",
help="Hidden layer sizes (provide space-separated sizes)",
nargs="+",
type=int,
default=[32, 8]
)
parser.add_argument(
"--nbuckets",
help="Number of buckets to divide lat and lon with",
type=int,
default=10
)
parser.add_argument(
"--lr",
help = "learning rate for optimizer",
type = float,
default = 0.001
)
parser.add_argument(
"--num_evals",
help="Number of times to evaluate model on eval data training.",
type=int,
default=5
)
parser.add_argument(
"--num_examples_to_train_on",
help="Number of examples to train on.",
type=int,
default=100
)
parser.add_argument(
"--output_dir",
help="GCS location to write checkpoints and export models",
required=True
)
parser.add_argument(
"--train_data_path",
help="GCS location pattern of train files containing eval URLs",
required=True
)
parser.add_argument(
"--job-dir",
help="this model ignores this field, but it is required by gcloud",
default="junk"
)
args = parser.parse_args()
hparams = args.__dict__
hparams.pop("job-dir", None)
model.train_and_evaluate(hparams)
# -
# ### Run trainer module package locally
#
# Now we can test our training code locally as follows using the local test data. We'll run a very small training job over a single file with a small batch size and one eval step.
# + language="bash"
#
# EVAL_DATA_PATH=./taxifare/tests/data/taxi-valid*
# TRAIN_DATA_PATH=./taxifare/tests/data/taxi-train*
# OUTPUT_DIR=./taxifare-model
#
# test ${OUTPUT_DIR} && rm -rf ${OUTPUT_DIR}
# export PYTHONPATH=${PYTHONPATH}:${PWD}/taxifare
#
# python3 -m trainer.task \
# --eval_data_path $EVAL_DATA_PATH \
# --output_dir $OUTPUT_DIR \
# --train_data_path $TRAIN_DATA_PATH \
# --batch_size 5 \
# --num_examples_to_train_on 100 \
# --num_evals 1 \
# --nbuckets 10 \
# --lr 0.001 \
# --nnsize 32 8
# -
# ### Run your training package on Cloud AI Platform
#
# Once the code works in standalone mode locally, you can run it on Cloud AI Platform. To submit to the Cloud we use [`gcloud ai-platform jobs submit training [jobname]`](https://cloud.google.com/sdk/gcloud/reference/ml-engine/jobs/submit/training) and simply specify some additional parameters for AI Platform Training Service:
# - jobid: A unique identifier for the Cloud job. We usually append system time to ensure uniqueness
# - region: Cloud region to train in. See [here](https://cloud.google.com/ml-engine/docs/tensorflow/regions) for supported AI Platform Training Service regions
#
# The arguments before `-- \` are for AI Platform Training Service.
# The arguments after `-- \` are sent to our `task.py`.
#
# Because this is on the entire dataset, it will take a while. You can monitor the job from the GCP console in the Cloud AI Platform section.
# + language="bash"
#
# # Output directory and jobID
# OUTDIR=gs://${BUCKET}/taxifare/trained_model_$(date -u +%y%m%d_%H%M%S)
# JOBID=taxifare_$(date -u +%y%m%d_%H%M%S)
# echo ${OUTDIR} ${REGION} ${JOBID}
# gsutil -m rm -rf ${OUTDIR}
#
# # Model and training hyperparameters
# BATCH_SIZE=50
# NUM_EXAMPLES_TO_TRAIN_ON=100
# NUM_EVALS=100
# NBUCKETS=10
# LR=0.001
# NNSIZE="32 8"
#
# # GCS paths
# GCS_PROJECT_PATH=gs://$BUCKET/taxifare
# DATA_PATH=$GCS_PROJECT_PATH/data
# TRAIN_DATA_PATH=$DATA_PATH/taxi-train*
# EVAL_DATA_PATH=$DATA_PATH/taxi-valid*
#
# #TODO 2
# gcloud ai-platform jobs submit training $JOBID \
# --module-name=trainer.task \
# --package-path=taxifare/trainer \
# --staging-bucket=gs://${BUCKET} \
# --python-version=3.7 \
# --runtime-version=${TFVERSION} \
# --region=${REGION} \
# -- \
# --eval_data_path $EVAL_DATA_PATH \
# --output_dir $OUTDIR \
# --train_data_path $TRAIN_DATA_PATH \
# --batch_size $BATCH_SIZE \
# --num_examples_to_train_on $NUM_EXAMPLES_TO_TRAIN_ON \
# --num_evals $NUM_EVALS \
# --nbuckets $NBUCKETS \
# --lr $LR \
# --nnsize $NNSIZE
# -
# ### (Optional) Run your training package using Docker container
#
# AI Platform Training also supports training in custom containers, allowing users to bring their own Docker containers with any pre-installed ML framework or algorithm to run on AI Platform Training.
#
# In this last section, we'll see how to submit a Cloud training job using a customized Docker image.
# Containerizing our `./taxifare/trainer` package involves 3 steps:
#
# * Writing a Dockerfile in `./taxifare`
# * Building the Docker image
# * Pushing it to the Google Cloud container registry in our GCP project
# The `Dockerfile` specifies
# 1. How the container needs to be provisioned so that all the dependencies in our code are satisfied
# 2. Where to copy our trainer Package in the container and how to install it (`pip install /trainer`)
# 3. What command to run when the container is ran (the `ENTRYPOINT` line)
# +
# %%writefile ./taxifare/Dockerfile
FROM gcr.io/deeplearning-platform-release/tf2-cpu
# TODO 3
COPY . /code
WORKDIR /code
ENTRYPOINT ["python3", "-m", "trainer.task"]
# -
# !gcloud auth configure-docker
# + language="bash"
#
# PROJECT_DIR=$(cd ./taxifare && pwd)
# PROJECT_ID=$(gcloud config list project --format "value(core.project)")
# IMAGE_NAME=taxifare_training_container
# DOCKERFILE=$PROJECT_DIR/Dockerfile
# IMAGE_URI=gcr.io/$PROJECT_ID/$IMAGE_NAME
#
# docker build $PROJECT_DIR -f $DOCKERFILE -t $IMAGE_URI
#
# docker push $IMAGE_URI
# -
# **Remark:** If you prefer to build the container image from the command line, we have written a script for that `./taxifare/scripts/build.sh`. This script reads its configuration from the file `./taxifare/scripts/env.sh`. You can configure these arguments the way you want in that file. You can also simply type `make build` from within `./taxifare` to build the image (which will invoke the build script). Similarly, we wrote the script `./taxifare/scripts/push.sh` to push the Docker image, which you can also trigger by typing `make push` from within `./taxifare`.
# ### Train using a custom container on AI Platform
#
# To submit to the Cloud we use [`gcloud ai-platform jobs submit training [jobname]`](https://cloud.google.com/sdk/gcloud/reference/ml-engine/jobs/submit/training) and simply specify some additional parameters for AI Platform Training Service:
# - jobname: A unique identifier for the Cloud job. We usually append system time to ensure uniqueness
# - master-image-uri: The uri of the Docker image we pushed in the Google Cloud registry
# - region: Cloud region to train in. See [here](https://cloud.google.com/ml-engine/docs/tensorflow/regions) for supported AI Platform Training Service regions
#
#
# The arguments before `-- \` are for AI Platform Training Service.
# The arguments after `-- \` are sent to our `task.py`.
# You can track your job and view logs using [cloud console](https://console.cloud.google.com/mlengine/jobs).
# + language="bash"
#
# PROJECT_ID=$(gcloud config list project --format "value(core.project)")
# BUCKET=$PROJECT_ID
# REGION="us-central1"
#
# # Output directory and jobID
# OUTDIR=gs://${BUCKET}/taxifare/trained_model
# JOBID=taxifare_container_$(date -u +%y%m%d_%H%M%S)
# echo ${OUTDIR} ${REGION} ${JOBID}
# gsutil -m rm -rf ${OUTDIR}
#
# # Model and training hyperparameters
# BATCH_SIZE=50
# NUM_EXAMPLES_TO_TRAIN_ON=100
# NUM_EVALS=100
# NBUCKETS=10
# NNSIZE="32 8"
#
# # AI-Platform machines to use for training
# MACHINE_TYPE=n1-standard-4
# SCALE_TIER=CUSTOM
#
# # GCS paths.
# GCS_PROJECT_PATH=gs://$BUCKET/taxifare
# DATA_PATH=$GCS_PROJECT_PATH/data
# TRAIN_DATA_PATH=$DATA_PATH/taxi-train*
# EVAL_DATA_PATH=$DATA_PATH/taxi-valid*
#
# IMAGE_NAME=taxifare_training_container
# IMAGE_URI=gcr.io/$PROJECT_ID/$IMAGE_NAME
#
# gcloud beta ai-platform jobs submit training $JOBID \
# --staging-bucket=gs://$BUCKET \
# --region=$REGION \
# --master-image-uri=$IMAGE_URI \
# --master-machine-type=$MACHINE_TYPE \
# --scale-tier=$SCALE_TIER \
# -- \
# --eval_data_path $EVAL_DATA_PATH \
# --output_dir $OUTDIR \
# --train_data_path $TRAIN_DATA_PATH \
# --batch_size $BATCH_SIZE \
# --num_examples_to_train_on $NUM_EXAMPLES_TO_TRAIN_ON \
# --num_evals $NUM_EVALS \
# --nbuckets $NBUCKETS \
# --nnsize $NNSIZE
#
# -
# **Remark:** If you prefer submitting your jobs for training on the AI-platform using the command line, we have written the `./taxifare/scripts/submit.sh` for you (that you can also invoke using `make submit` from within `./taxifare`). As the other scripts, it reads it configuration variables from `./taxifare/scripts/env.sh`.
# Copyright 2020 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
|
courses/machine_learning/deepdive2/building_production_ml_systems/solutions/1_training_at_scale.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
from dolfin import *
from mshr import *
import numpy as np
from matplotlib import cm
import matplotlib.pyplot as plt
import matplotlib.tri as tri
from mpl_toolkits.mplot3d import Axes3D
import logging
logging.getLogger("FFC").setLevel(logging.WARNING)
set_log_active(False)
dolfin.parameters.reorder_dofs_serial = False
# +
def mesh2triang(mesh):
xy = mesh.coordinates()
return tri.Triangulation(xy[:, 0], xy[:, 1], mesh.cells())
def mplot_cellfunction(cellfn):
C = cellfn.array()
tri = mesh2triang(cellfn.mesh())
return plt.tripcolor(tri, facecolors=C)
def mplot_function(f):
mesh = f.function_space().mesh()
if (mesh.geometry().dim() != 2):
raise AttributeError('Mesh must be 2D')
# DG0 cellwise function
if f.vector().size() == mesh.num_cells():
C = f.vector().array()
return plt.tripcolor(mesh2triang(mesh), C)
# Scalar function, interpolated to vertices
elif f.value_rank() == 0:
C = f.compute_vertex_values(mesh)
return plt.tripcolor(mesh2triang(mesh), C, shading='gouraud')
# Vector function, interpolated to vertices
elif f.value_rank() == 1:
w0 = f.compute_vertex_values(mesh)
if (len(w0) != 2*mesh.num_vertices()):
raise AttributeError('Vector field must be 2D')
X = mesh.coordinates()[:, 0]
Y = mesh.coordinates()[:, 1]
U = w0[:mesh.num_vertices()]
V = w0[mesh.num_vertices():]
return plt.quiver(X,Y,U,V)
# Plot a generic dolfin object (if supported)
def plot(obj):
plt.gca().set_aspect('equal')
if isinstance(obj, Function):
return mplot_function(obj)
elif isinstance(obj, CellFunctionSizet):
return mplot_cellfunction(obj)
elif isinstance(obj, CellFunctionDouble):
return mplot_cellfunction(obj)
elif isinstance(obj, CellFunctionInt):
return mplot_cellfunction(obj)
elif isinstance(obj, Mesh):
if (obj.geometry().dim() != 2):
raise AttributeError('Mesh must be 2D')
return plt.triplot(mesh2triang(obj), color='#808080')
raise AttributeError('Failed to plot %s'%type(obj))
# end of commands for plotting
# +
s = 2
thetaS = 0.2
#THIS SECTION IS FOR MAKING THE MESH
r1 = 1 #radius of cylinder
r2 = 10 #radius of bounding circle
nr = 20 #number of radial divisions
nt = 50 #number of theta divisions
mesh = RectangleMesh(Point(r1, 0), Point(r2, 1), nr, nt, "crossed")
x = mesh.coordinates()[:,0]
y = mesh.coordinates()[:,1]
def denser(x,y):
return [r1 + (r2-r1)*((x-r1)/(r2-r1))**s, y]
x_bar, y_bar = denser(x, y)
xy_bar_coor = np.array([x_bar,y_bar]).transpose()
mesh.coordinates()[:] = xy_bar_coor
def cylinder(r, s, Theta):
return [r*np.cos(Theta*s), r*np.sin(Theta*s)]
x_hat, y_hat = cylinder(x_bar, y_bar,2*np.pi)
xy_hat_coor = np.array([x_hat,y_hat]).transpose()
mesh.coordinates()[:] = xy_hat_coor
# +
#THIS SECTION IS FOR MARKING BOUNDARIES OF THE MESH
class outer_boundary(SubDomain):
def inside(self, x, on_boundary):
tol = 1E-1
r = sqrt(x[0]*x[0] + x[1]*x[1])
return on_boundary and abs(r - r2) < tol
class inner_boundary(SubDomain):
def inside(self, x, on_boundary):
tol = 1E-1
r = sqrt(x[0]*x[0] + x[1]*x[1])
return on_boundary and abs(r - r1) < tol
class half_circleX(SubDomain):
def inside(self, x, on_boundary):
tol = 1E-1
return on_boundary and abs(x[1]) < tol
class half_circleY(SubDomain):
def inside(self, x, on_boundary):
tol = 1E-1
return on_boundary and abs(x[0]) < tol
outerradius = outer_boundary()
innerradius = inner_boundary()
half_circleX = half_circleX()
half_circleY = half_circleY()
boundaries = FacetFunction("size_t", mesh)
boundaries.set_all(0)
innerradius.mark(boundaries,1)
outerradius.mark(boundaries,2)
half_circleX.mark(boundaries,3)
half_circleY.mark(boundaries,4)
# -
#This section defines the finite element basis functions
V = FunctionSpace(mesh,'Lagrange',1)
nx = TrialFunction(V)
ny = TrialFunction(V)
v1 = TestFunction(V)
v2 = TestFunction(V)
# +
#Boundary Conditions
gx = Expression('-x[1]*sin(ThetaInner)/sqrt(x[0]*x[0] + x[1]*x[1])',ThetaInner = thetaS)
gy = Expression('x[0]*sin(ThetaInner)/sqrt(x[0]*x[0] + x[1]*x[1])',ThetaInner = thetaS)
#For nx
bcs1 = [DirichletBC(V, 0, boundaries, 2),
DirichletBC(V, gx, boundaries, 1),
DirichletBC(V, 0, boundaries, 3)]
#for ny
bcs2 = [DirichletBC(V, 0, boundaries, 2),
DirichletBC(V, gy, boundaries, 1),
DirichletBC(V, 0, boundaries, 4)]
# +
#Solving the problem
f = Constant(0.0)
a1 = inner(nabla_grad(nx), nabla_grad(v1))*dx
a2 = inner(nabla_grad(ny), nabla_grad(v2))*dx
L1 = f*v1*dx
L2 = f*v2*dx
nx = Function(V)
ny = Function(V)
solve(a1 == L1, nx, bcs1)
solve(a2 == L2, ny, bcs2)
V_d = VectorFunctionSpace(mesh,'Lagrange',1)
w1 = TrialFunction(V_d)
w2 = TrialFunction(V_d)
q1 = TestFunction(V_d)
q2 = TestFunction(V_d)
a3 = inner(w1,q1)*dx
a4 = inner(w2,q2)*dx
L3 = inner(grad(nx),q1)*dx
L4 = inner(grad(ny),q2)*dx
grad_nx = Function(V_d)
grad_ny = Function(V_d)
solve(a3 == L3, grad_nx)
solve(a4 == L4, grad_ny)
dxnx, dynx = grad_nx.split(deepcopy=True)
dxny, dyny = grad_ny.split(deepcopy=True)
# +
#Finding the energies
class int_radius(SubDomain):
def inside(self, x, on_boundary):
tol = 1E-4
r = sqrt(x[0]*x[0] + x[1]*x[1])
return ((r - r1)> tol)
int_radius = int_radius()
cell_markers = CellFunction("uint", mesh)
cell_markers.set_all(0)
int_radius.mark(cell_markers,5)
dx1 = Measure("dx", domain=mesh, subdomain_data=cell_markers)
splay = pow(dxnx+dyny,2)
twist = pow(dxny-dynx,2)
Esplay = assemble(splay*dx1(5))
Etwist = assemble(twist*dx1(5))
fel = splay+twist
f1 = project(fel,V)
Etot = Esplay+Etwist
print("Splay Energy %f" % Esplay)
print("Twist Energy %f" % Esplay)
print("Total Energy %f" % Etot)
# -
plt.figure(figsize=(10,10))
plot(project(curl(nx)/nper))
# +
nper = pow((pow(nx,2)+pow(ny,2)),1/2)
NPER = project(nper,V)
plt.figure(figsize=(10,10))
plot(project(curl(NPER)))
plt.colorbar(shrink=0.8);
# -
plt.figure(figsize=(10,10))
plot(f1)
plt.colorbar(shrink=0.8);
vec = as_vector([nx,ny])
plot(project(vec,V))
|
Nematic/SingleCylbadmesh.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get started
#
# <a href="https://mybinder.org/v2/gh/tinkoff-ai/etna/master?filepath=examples/get_started.ipynb">
# <img src="https://mybinder.org/badge_logo.svg" align='left'>
# </a>
# This notebook contains the simple examples of time series forecasting pipeline
# using ETNA library.
#
# **Table of Contents**
#
# * [Creating TSDataset](#chapter1)
# * [Plotting](#chapter2)
# * [Forecast single time series](#chapter3)
# * [Simple forecast](#section_3_1)
# * [Prophet](#section_3_2)
# * [Catboost](#section_3_3)
# * [Forecast multiple time series](#chapter4)
# * [Pipeline](#chapter5)
# ## 1. Creating TSDataset <a class="anchor" id="chapter1"></a>
# Let's load and look at the dataset
# + pycharm={"name": "#%%\n"}
import pandas as pd
# + pycharm={"name": "#%%\n"}
original_df = pd.read_csv("data/monthly-australian-wine-sales.csv")
original_df.head()
# + [markdown] pycharm={"name": "#%% md\n"}
# etna_ts is strict about data format:
# * column we want to predict should be called `target`
# * column with datatime data should be called `timestamp`
# * because etna is always ready to work with multiple time series, column `segment` is also compulsory
#
# Our library works with the special data structure TSDataset. So, before starting anything, we need to convert the classical DataFrame to TSDataset.
#
# Let's rename first
# + pycharm={"name": "#%%\n"}
original_df["timestamp"] = pd.to_datetime(original_df["month"])
original_df["target"] = original_df["sales"]
original_df.drop(columns=["month", "sales"], inplace=True)
original_df["segment"] = "main"
original_df.head()
# + [markdown] pycharm={"name": "#%% md\n"}
# Time to convert to TSDataset!
#
# To do this, we initially need to convert the classical DataFrame to the special format.
# + pycharm={"name": "#%%\n"}
from etna.datasets.tsdataset import TSDataset
# + pycharm={"name": "#%%\n"}
df = TSDataset.to_dataset(original_df)
df.head()
# -
# Now we can construct the TSDataset.
#
# Additionally to passing dataframe we should specify frequency of our data.
# In this case it is monthly data.
# + pycharm={"name": "#%%\n"}
ts = TSDataset(df, freq='1M')
# -
# Oups. Let's fix that
# + pycharm={"name": "#%%\n"}
ts = TSDataset(df, freq='MS')
# -
# ## 2. Plotting <a class="anchor" id="chapter2"></a>
#
# Let's take a look at the time series in the dataset
# + pycharm={"name": "#%%\n"}
ts.plot()
# -
# ## 3. Forecasting single time series <a class="anchor" id="chapter3"></a>
#
# Our library contains a wide range of different models for time series forecasting. Let's look at some of them.
# ### 3.1 Simple forecast<a class="anchor" id="section_3_1"></a>
# Let's predict the monthly values in 1994 in our dataset using the ```NaiveModel```
# + pycharm={"name": "#%%\n"}
train_ts, test_ts = ts.train_test_split(train_start='1980-01-01',
train_end='1993-12-01',
test_start='1994-01-01',
test_end='1994-08-01')
# + pycharm={"name": "#%%\n"}
HORIZON = 8
from etna.models import NaiveModel
#Fit the model
model = NaiveModel(lag=12)
model.fit(train_ts)
#Make the forecast
future_ts = train_ts.make_future(HORIZON)
forecast_ts = model.forecast(future_ts)
# + [markdown] pycharm={"name": "#%% md\n"}
# Now let's look at a metric and plot the prediction.
# All the methods already built-in in etna.
# + pycharm={"name": "#%%\n"}
from etna.metrics import SMAPE
# + pycharm={"name": "#%%\n"}
smape = SMAPE()
smape(y_true=test_ts, y_pred=forecast_ts)
# + pycharm={"name": "#%%\n"}
from etna.analysis import plot_forecast
# + pycharm={"name": "#%%\n"}
plot_forecast(forecast_ts, test_ts, train_ts, n_train_samples=10)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### 3.2 Prophet<a class="anchor" id="section_3_2"></a>
#
# Now try to improve the forecast and predict the values with the Facebook Prophet.
# + pycharm={"name": "#%%\n"}
from etna.models import ProphetModel
model = ProphetModel()
model.fit(train_ts)
#Make the forecast
future_ts = train_ts.make_future(HORIZON)
forecast_ts = model.forecast(future_ts)
# + pycharm={"name": "#%%\n"}
smape(y_true=test_ts, y_pred=forecast_ts)
# + pycharm={"name": "#%%\n"}
plot_forecast(forecast_ts, test_ts, train_ts, n_train_samples=10)
# -
# ### 3.2 Catboost<a class="anchor" id="section_3_3"></a>
# And finally let's try the Catboost model.
#
# Also etna has wide range of transforms you may apply to your data.
#
# Here how it is done:
# + pycharm={"name": "#%%\n"}
from etna.transforms.lags import LagTransform
lags = LagTransform(in_column="target", lags=list(range(8, 24, 1)))
train_ts.fit_transform([lags])
# + pycharm={"name": "#%%\n"}
from etna.models import CatBoostModelMultiSegment
model = CatBoostModelMultiSegment()
model.fit(train_ts)
future_ts = train_ts.make_future(HORIZON)
forecast_ts = model.forecast(future_ts)
# + pycharm={"name": "#%%\n"}
from etna.metrics import SMAPE
smape = SMAPE()
smape(y_true=test_ts, y_pred=forecast_ts)
# + pycharm={"name": "#%%\n"}
from etna.analysis import plot_forecast
train_ts.inverse_transform()
plot_forecast(forecast_ts, test_ts, train_ts, n_train_samples=10)
# -
# ## 4. Forecasting multiple time series <a class="anchor" id="chapter4"></a>
#
# In this section you may see example of how easily etna works
# with multiple time series and get acquainted with other transforms etna contains.
# + pycharm={"name": "#%%\n"}
original_df = pd.read_csv("data/example_dataset.csv")
original_df.head()
# + pycharm={"name": "#%%\n"}
df = TSDataset.to_dataset(original_df)
ts = TSDataset(df, freq='D')
ts.plot()
# + pycharm={"name": "#%%\n"}
import warnings
from etna.transforms import MeanTransform, LagTransform, LogTransform, \
SegmentEncoderTransform, DateFlagsTransform, LinearTrendTransform
warnings.filterwarnings("ignore")
log = LogTransform(in_column="target")
trend = LinearTrendTransform(in_column="target")
seg = SegmentEncoderTransform()
lags = LagTransform(in_column="target", lags=list(range(30, 96, 1)))
d_flags = DateFlagsTransform(day_number_in_week=True,
day_number_in_month=True,
week_number_in_month=True,
week_number_in_year=True,
month_number_in_year=True,
year_number=True,
special_days_in_week=[5, 6])
mean30 = MeanTransform(in_column="target", window=30)
# + pycharm={"name": "#%%\n"}
HORIZON = 31
train_ts, test_ts = ts.train_test_split(train_start='2019-01-01',
train_end='2019-11-30',
test_start='2019-12-01',
test_end='2019-12-31')
train_ts.fit_transform([log, trend, lags, d_flags, seg, mean30])
# + pycharm={"name": "#%%\n"}
from etna.models import CatBoostModelMultiSegment
model = CatBoostModelMultiSegment()
model.fit(train_ts)
future_ts = train_ts.make_future(HORIZON)
forecast_ts = model.forecast(future_ts)
# + pycharm={"name": "#%%\n"}
smape = SMAPE()
smape(y_true=test_ts, y_pred=forecast_ts)
# + pycharm={"name": "#%%\n"}
train_ts.inverse_transform()
plot_forecast(forecast_ts, test_ts, train_ts, n_train_samples=20)
# -
# ## 5. Pipeline <a class="anchor" id="chapter5"></a>
#
# Let's wrap everything into pipeline to create the end-to-end model from previous section.
from etna.pipeline import Pipeline
train_ts, test_ts = ts.train_test_split(train_start='2019-01-01',
train_end='2019-11-30',
test_start='2019-12-01',
test_end='2019-12-31')
# We put: **model**, **transforms** and **horizon** in a single object, which has the similar interface with the model(fit/forecast)
model = Pipeline(model=CatBoostModelMultiSegment(),
transforms=[log, trend, lags, d_flags, seg, mean30],
horizon=HORIZON)
model.fit(train_ts)
forecast_ts = model.forecast()
# As in the previous section, let's calculate the metrics and plot the forecast
smape = SMAPE()
smape(y_true=test_ts, y_pred=forecast_ts)
plot_forecast(forecast_ts, test_ts, train_ts, n_train_samples=20)
|
examples/get_started.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import px4tools
import pandas
import pylab as pl
# %matplotlib inline
pl.rcParams['figure.figsize'] = (15,5)
data = px4tools.process_data(
pandas.read_csv('logs/15-10-01-07_47_35-jgoppert-retune-flight.csv'))
px4tools.find_lpe_gains(data[10:100])
data.STAT_MainState.plot()
px4tools.plot_position_loops(data)
px4tools.plot_velocity_loops(data)
px4tools.pos_analysis(data[1:200]);
data.LPOS_VZ.plot()
|
15-10-01 jgoppert retune.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # Image classification - transfer learning demo
#
# 1. [Introduction](#Introduction)
# 2. [Prerequisites and Preprocessing](#Prequisites-and-Preprocessing)
# 3. [Fine-tuning the Image classification model](#Fine-tuning-the-Image-classification-model)
# 4. [Set up hosting for the model](#Set-up-hosting-for-the-model)
# 1. [Import model into hosting](#Import-model-into-hosting)
# 2. [Create endpoint configuration](#Create-endpoint-configuration)
# 3. [Create endpoint](#Create-endpoint)
# 5. [Perform Inference](#Perform-Inference)
#
# ## Introduction
#
# Welcome to our end-to-end example of distributed image classification algorithm in transfer learning mode. In this demo, we will use the Amazon sagemaker image classification algorithm in transfer learning mode to fine-tune a pre-trained model (trained on imagenet data) to learn to classify a new dataset. In particular, the pre-trained model will be fine-tuned using [CIFAR-10 dataset](https://www.cs.toronto.edu/~kriz/cifar.html).
#
# To get started, we need to set up the environment with a few prerequisite steps, for permissions, configurations, and so on.
# ## Prequisites and Preprocessing
#
# ### Permissions and environment variables
#
# Here we set up the linkage and authentication to AWS services. There are three parts to this:
#
# * The roles used to give learning and hosting access to your data. This will automatically be obtained from the role used to start the notebook
# * The S3 bucket that you want to use for training and model data
# * The Amazon sagemaker image classification docker image which need not be changed
# +
# %%time
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
bucket='jsimon-sagemaker-us' # customize to your bucket
containers = {'us-west-2': '433757028032.dkr.ecr.us-west-2.amazonaws.com/image-classification:latest',
'us-east-1': '811284229777.dkr.ecr.us-east-1.amazonaws.com/image-classification:latest',
'us-east-2': '825641698319.dkr.ecr.us-east-2.amazonaws.com/image-classification:latest',
'eu-west-1': '685385470294.dkr.ecr.eu-west-1.amazonaws.com/image-classification:latest'}
training_image = containers[boto3.Session().region_name]
print(training_image)
# -
# ## Fine-tuning the Image classification model
#
# The CIFAR-10 dataset consist of images from 10 categories and has 50,000 images with 5,000 images per category.
#
# The image classification algorithm can take two types of input formats. The first is a [recordio format](https://mxnet.incubator.apache.org/tutorials/basic/record_io.html) and the other is a [lst format](https://mxnet.incubator.apache.org/how_to/recordio.html?highlight=im2rec). Files for both these formats are available at http://data.mxnet.io/data/cifar10/. In this example, we will use the recordio format for training and use the training/validation split.
# +
import os
import urllib.request
import boto3
def download(url):
filename = url.split("/")[-1]
if not os.path.exists(filename):
urllib.request.urlretrieve(url, filename)
def upload_to_s3(channel, file):
s3 = boto3.resource('s3')
data = open(file, "rb")
key = channel + '/' + file
s3.Bucket(bucket).put_object(Key=key, Body=data)
# CIFAR-10
download('http://data.mxnet.io/data/cifar10/cifar10_train.rec')
download('http://data.mxnet.io/data/cifar10/cifar10_val.rec')
upload_to_s3('validation/cifar10', 'cifar10_val.rec')
upload_to_s3('train/cifar10', 'cifar10_train.rec')
# -
# Once we have the data available in the correct format for training, the next step is to actually train the model using the data. Before training the model, we need to setup the training parameters. The next section will explain the parameters in detail.
# ## Training parameters
# There are two kinds of parameters that need to be set for training. The first one are the parameters for the training job. These include:
#
# * **Input specification**: These are the training and validation channels that specify the path where training data is present. These are specified in the "InputDataConfig" section. The main parameters that need to be set is the "ContentType" which can be set to "application/x-recordio" or "application/x-image" based on the input data format and the S3Uri which specifies the bucket and the folder where the data is present.
# * **Output specification**: This is specified in the "OutputDataConfig" section. We just need to specify the path where the output can be stored after training
# * **Resource config**: This section specifies the type of instance on which to run the training and the number of hosts used for training. If "InstanceCount" is more than 1, then training can be run in a distributed manner.
#
# Apart from the above set of parameters, there are hyperparameters that are specific to the algorithm. These are:
#
# * **num_layers**: The number of layers (depth) for the network. We use 44 in this sample but other values can be used.
# * **num_training_samples**: This is the total number of training samples. It is set to 50000 for CIFAR-10 dataset with the current split
# * **num_classes**: This is the number of output classes for the new dataset. Imagenet was trained with 1000 output classes but the number of output classes can be changed for fine-tuning. For CIFAR-10, we use 10.
# * **epochs**: Number of training epochs
# * **learning_rate**: Learning rate for training
# * **mini_batch_size**: The number of training samples used for each mini batch. In distributed training, the number of training samples used per batch will be N * mini_batch_size where N is the number of hosts on which training is run
# After setting training parameters, we kick off training, and poll for status until training is completed, which in this example, takes between 10 to 12 minutes per epoch on a p2.xlarge machine. The network typically converges after 10 epochs.
# + isConfigCell=true
# The algorithm supports multiple network depth (number of layers). They are 18, 34, 50, 101, 152 and 200
# For this training, we will use 18 layers
num_layers = 50
# we need to specify the input image shape for the training data
image_shape = "3,28,28"
# we also need to specify the number of training samples in the training set
# for CIFAR-10 it is 50000
num_training_samples = 50000
# specify the number of output classes
num_classes = 10
# batch size for training
mini_batch_size = 128
# number of epochs
epochs = 10
# learning rate
learning_rate = 0.01
# Since we are using transfer learning, we set use_pretrained_model to 1 so that weights can be
# initialized with pre-trained weights
use_pretrained_model = 1
# -
# # Training
# Run the training using Amazon sagemaker CreateTrainingJob API
# +
# %%time
import time
import boto3
from time import gmtime, strftime
s3 = boto3.client('s3')
# create unique job name
job_name_prefix = 'sagemaker-imageclassification-cifar10'
timestamp = time.strftime('-%Y-%m-%d-%H-%M-%S', time.gmtime())
job_name = job_name_prefix + timestamp
training_params = \
{
# specify the training docker image
"AlgorithmSpecification": {
"TrainingImage": training_image,
"TrainingInputMode": "File"
},
"RoleArn": role,
"OutputDataConfig": {
"S3OutputPath": 's3://{}/{}/output'.format(bucket, job_name_prefix)
},
"ResourceConfig": {
"InstanceCount": 1,
"InstanceType": "ml.p2.8xlarge",
"VolumeSizeInGB": 50
},
"TrainingJobName": job_name,
"HyperParameters": {
"image_shape": image_shape,
"num_layers": str(num_layers),
"num_training_samples": str(num_training_samples),
"num_classes": str(num_classes),
"mini_batch_size": str(mini_batch_size),
"epochs": str(epochs),
"learning_rate": str(learning_rate),
"use_pretrained_model": str(use_pretrained_model)
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 360000
},
#Training data should be inside a subdirectory called "train"
#Validation data should be inside a subdirectory called "validation"
#The algorithm currently only supports fullyreplicated model (where data is copied onto each machine)
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": 's3://{}/train/cifar10'.format(bucket),
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "application/x-recordio",
"CompressionType": "None"
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": 's3://{}/validation/cifar10'.format(bucket),
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "application/x-recordio",
"CompressionType": "None"
}
]
}
print('Training job name: {}'.format(job_name))
print('\nInput Data Location: {}'.format(training_params['InputDataConfig'][0]['DataSource']['S3DataSource']))
# +
# create the Amazon SageMaker training job
sagemaker = boto3.client(service_name='sagemaker')
sagemaker.create_training_job(**training_params)
# confirm that the training job has started
status = sagemaker.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print('Training job current status: {}'.format(status))
try:
# wait for the job to finish and report the ending status
sagemaker.get_waiter('training_job_completed_or_stopped').wait(TrainingJobName=job_name)
training_info = sagemaker.describe_training_job(TrainingJobName=job_name)
status = training_info['TrainingJobStatus']
print("Training job ended with status: " + status)
except:
print('Training failed to start')
# if exception is raised, that means it has failed
message = sagemaker.describe_training_job(TrainingJobName=job_name)['FailureReason']
print('Training failed with the following error: {}'.format(message))
# -
training_info = sagemaker.describe_training_job(TrainingJobName=job_name)
status = training_info['TrainingJobStatus']
print("Training job ended with status: " + status)
# If you see the message,
#
# > `Training job ended with status: Completed`
#
# then that means training sucessfully completed and the output model was stored in the output path specified by `training_params['OutputDataConfig']`.
#
# You can also view information about and the status of a training job using the AWS SageMaker console. Just click on the "Jobs" tab.
# ## Plot training and validation accuracies
# +
import boto3
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
client = boto3.client('logs')
lgn='/aws/sagemaker/TrainingJobs'
lsn='sagemaker-imageclassification-cifar10-2018-01-16-11-05-28/algo-1-1516100993'
log=client.get_log_events(logGroupName=lgn, logStreamName=lsn)
trn_accs=[]
val_accs=[]
for e in log['events']:
msg=e['message']
if 'Validation-accuracy' in msg:
val = msg.split("=")
val = val[1]
val_accs.append(float(val))
if 'Train-accuracy' in msg:
trn = msg.split("=")
trn = trn[1]
trn_accs.append(float(trn))
print("Maximum validation accuracy: %f " % max(val_accs))
fig, ax = plt.subplots()
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
trn_plot, = ax.plot(range(epochs), trn_accs, label="Training accuracy")
val_plot, = ax.plot(range(epochs), val_accs, label="Validation accuracy")
plt.legend(handles=[trn_plot,val_plot])
ax.yaxis.set_ticks(np.arange(0.4, 1.05, 0.05))
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter('%0.2f'))
plt.show()
# -
# # Inference
#
# ***
#
# A trained model does nothing on its own. We now want to use the model to perform inference. For this example, that means predicting the topic mixture representing a given document.
#
# This section involves several steps,
#
# 1. [Create Model](#CreateModel) - Create model for the training output
# 1. [Create Endpoint Configuration](#CreateEndpointConfiguration) - Create a configuration defining an endpoint.
# 1. [Create Endpoint](#CreateEndpoint) - Use the configuration to create an inference endpoint.
# 1. [Perform Inference](#Perform Inference) - Perform inference on some input data using the endpoint.
# ## Create Model
#
# We now create a SageMaker Model from the training output. Using the model we can create an Endpoint Configuration.
# +
# %%time
import boto3
from time import gmtime, strftime
sage = boto3.Session().client(service_name='sagemaker')
model_name="image-classification-cifar-transfer"
print(model_name)
info = sage.describe_training_job(TrainingJobName=job_name)
model_data = info['ModelArtifacts']['S3ModelArtifacts']
print(model_data)
containers = {'us-west-2': '433757028032.dkr.ecr.us-west-2.amazonaws.com/image-classification:latest',
'us-east-1': '811284229777.dkr.ecr.us-east-1.amazonaws.com/image-classification:latest',
'us-east-2': '825641698319.dkr.ecr.us-east-2.amazonaws.com/image-classification:latest',
'eu-west-1': '685385470294.dkr.ecr.eu-west-1.amazonaws.com/image-classification:latest'}
hosting_image = containers[boto3.Session().region_name]
primary_container = {
'Image': hosting_image,
'ModelDataUrl': model_data,
}
create_model_response = sage.create_model(
ModelName = model_name,
ExecutionRoleArn = role,
PrimaryContainer = primary_container)
print(create_model_response['ModelArn'])
# -
# ### Create Endpoint Configuration
# At launch, we will support configuring REST endpoints in hosting with multiple models, e.g. for A/B testing purposes. In order to support this, customers create an endpoint configuration, that describes the distribution of traffic across the models, whether split, shadowed, or sampled in some way.
#
# In addition, the endpoint configuration describes the instance type required for model deployment, and at launch will describe the autoscaling configuration.
# +
from time import gmtime, strftime
timestamp = time.strftime('-%Y-%m-%d-%H-%M-%S', time.gmtime())
endpoint_config_name = job_name_prefix + '-epc-' + timestamp
endpoint_config_response = sage.create_endpoint_config(
EndpointConfigName = endpoint_config_name,
ProductionVariants=[{
'InstanceType':'ml.m4.xlarge',
'InitialInstanceCount':1,
'ModelName':model_name,
'VariantName':'AllTraffic'}])
print('Endpoint configuration name: {}'.format(endpoint_config_name))
print('Endpoint configuration arn: {}'.format(endpoint_config_response['EndpointConfigArn']))
# -
# ### Create Endpoint
# Lastly, the customer creates the endpoint that serves up the model, through specifying the name and configuration defined above. The end result is an endpoint that can be validated and incorporated into production applications. This takes 9-11 minutes to complete.
# +
# %%time
import time
timestamp = time.strftime('-%Y-%m-%d-%H-%M-%S', time.gmtime())
endpoint_name = job_name_prefix + '-ep-' + timestamp
print('Endpoint name: {}'.format(endpoint_name))
endpoint_params = {
'EndpointName': endpoint_name,
'EndpointConfigName': endpoint_config_name,
}
endpoint_response = sagemaker.create_endpoint(**endpoint_params)
print('EndpointArn = {}'.format(endpoint_response['EndpointArn']))
# -
# Finally, now the endpoint can be created. It may take sometime to create the endpoint...
# +
# get the status of the endpoint
response = sagemaker.describe_endpoint(EndpointName=endpoint_name)
status = response['EndpointStatus']
print('EndpointStatus = {}'.format(status))
# wait until the status has changed
sagemaker.get_waiter('endpoint_in_service').wait(EndpointName=endpoint_name)
# print the status of the endpoint
endpoint_response = sagemaker.describe_endpoint(EndpointName=endpoint_name)
status = endpoint_response['EndpointStatus']
print('Endpoint creation ended with EndpointStatus = {}'.format(status))
if status != 'InService':
raise Exception('Endpoint creation failed.')
# -
# If you see the message,
#
# > `Endpoint creation ended with EndpointStatus = InService`
#
# then congratulations! You now have a functioning inference endpoint. You can confirm the endpoint configuration and status by navigating to the "Endpoints" tab in the AWS SageMaker console.
#
# We will finally create a runtime object from which we can invoke the endpoint.
# ## Perform Inference
# Finally, the customer can now validate the model for use. They can obtain the endpoint from the client library using the result from previous operations, and generate classifications from the trained model using that endpoint.
#
import boto3
runtime = boto3.Session().client(service_name='runtime.sagemaker')
# ### Download test image
# Bird
# #!wget -O /tmp/test.jpg https://cdn.pixabay.com/photo/2015/12/19/10/54/bird-1099639_960_720.jpg
# Horse
# #!wget -O /tmp/test.jpg https://cdn.pixabay.com/photo/2016/02/15/13/26/horse-1201143_960_720.jpg
# Dog
# !wget -O /tmp/test.jpg https://cdn.pixabay.com/photo/2016/02/19/15/46/dog-1210559_960_720.jpg
# Truck
# #!wget -O /tmp/test.jpg https://cdn.pixabay.com/photo/2015/09/29/10/14/truck-truck-963637_960_720.jpg
file_name = '/tmp/test.jpg'
# test image
from IPython.display import Image
Image(file_name)
import json
import numpy as np
with open(file_name, 'rb') as f:
payload = f.read()
payload = bytearray(payload)
response = runtime.invoke_endpoint(EndpointName=endpoint_name,
ContentType='application/x-image',
Body=payload)
result = response['Body'].read()
# result will be in json format and convert it to ndarray
result = json.loads(result)
print(result)
# the result will output the probabilities for all classes
# find the class with maximum probability and print the class index
index = np.argmax(result)
object_categories = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
print("Result: label - " + object_categories[index] + ", probability - " + str(result[index]))
# ### Clean up
#
# When we're done with the endpoint, we can just delete it and the backing instances will be released. Run the following cell to delete the endpoint.
sage.delete_endpoint(EndpointName=endpoint_name)
|
40_AWS_SageMaker/transfer_learning/01-Image-classification-transfer-learning-cifar10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computational Assignment 1
# **Assigned Monday, 9-9-19.**, **Due Thursday, 9-12-19.**
# Most of the problems we encounter in computational chemistry are multidimensional. This means that we need to be able to work with vectors and matrices in our code. Even when we consider a 1-dimensional function, we still need to code all of the data points into a list.
#
# Additionally, when we need to analyze data, we don't want to reinvent the wheel. It can be useful to code your own math operation once to learn how it work, but most of the time you should be using existing libraries in your work. We will cover how to use math and science libraries to run calcualtions.
#
# This notebook will cover the following concepts:
#
# 1. Lists and arrays
# 1. Defining lists
# 1. Accessing list values
# 1. Changing lists
# 1. Counting, sorting, and looping methods
# ## 1. Lists and Arrays
# Computers regularly deal with multidimensional data. Even if you have one-dimensional data from some sort of function, $F(t)$, you would write the data points into a list of numbers. When talking about code, we call these lists or arrays.
my_list=[1,2,3,4,5,6,7]
# Python specifically has a few of ways to handle arrays. The most basic data object for handeling data is the **list**. The other two data objects are **tuples** and **dictionaries**.
#
# Lists are defined with brackets `[]`
# Tuples are defined with parenthessis `()`
# Dictionaries are defined with curly brackets `{}`
#
# We won't spend much time with tuples. They behave a lot like lists, but once make one, you can't change any of the values in the tuple. This can be useful if you want to make sure data values are not changed through a calculation. They also tend be faster to process, but you won't notice the speedup unless you are trying to process lots of data (Gb of data).
# ## 1.1 Defining Lists
# In python, we create **lists** by defining a set of numbers in a bracket
[1,2,3]
# You should also make sure to define your list as a variable so that you can use it later. Otherwise there is no way to call it. If you don't want to overwrite your variable, use a different name for the new varaiable
my_list2=[1,2,3]
print(my_list2)
# You can also define an empty list. This is sometimes usefull if you plan to add alements to the list (more on the later
empty=[]
# Lists also don't have to be numbers. They can be a collection of strings (text)
sonoran_plants=["saguaro","ocotillo","pitaya","creosote"]
# Your list can be a list of lists! Notice the line break. You can do that when there is a comma. This helps make your code readable, which is one of the underlying philosophies of python
sonoran_plants_animals=[["saguaro","ocotillo","pitaya","creosote"],
["coyote","kangaroo mouse","javalina","gila monster"]]
# Notice that the two nested lists are different sizes. Mathematically, these lists are not going to behave like vectors or matrices. Python will not add or multiply a list according to the rules of matrix algebra. This is fine. When we need matrices and vectors, it will make more sense to use arrays associated with the math library Numpy. And the Numpy library can usually understand most python lists. Most of the time, if you are using a list, you are mostly trying to organize data, not run a heavy calculation.
# ### Exercise 1.1:
# Make your own lists.
#
# 1. Make a one-dimensional list of numbers
# 1. A three dimensional list of numbers (a list of three lists)
# 1. Make a lists of strings
# 1. Make a two-dimensional list of strings
# 1. Print all of your lists
#
# Make sure to define your lists with variables. We will use your lists later on
# +
# Your code here
Even_numbers=[2,4,6]
Some_integers=[[1,2,3],[4,5,6],[7,8,9]]
Slang=["dope","swag","gucci","lit"]
Cool_words=[["yo","wassup","wasspoppin"],["peace","love","pixie_dust"]]
print(Even_numbers)
print(Some_integers)
print(Slang)
print(Cool_words)
# -
# ## 1.2 List indexing
# Once you've made a list, you need to know how to get values from the list. To get a value from list use square brackets next to the variable name. `my_list[index]`. The first thing to note is that list indeces start at 0
# Printing the list to remind us of the elements
print(my_list)
my_list[0]
# Your indexing can also be negative. The list indexing is cyclic, so if the first element is 0, the last element is -1, the second to last element is -2, etc.
my_list[-1]
my_list[-2]
# If your list is a list of lists, calling an index will give you the nested list. You need two indices to get individual items
# Printing the list to remind us of the elements
print(sonoran_plants_animals)
sonoran_plants_animals[0]
sonoran_plants_animals[0][-1]
# You can also make a new sublist by calling a range of indices
# Printing the list to remind us of the elements
print(my_list)
my_list[2:5]
# You can also make a range with negatice indices. Order matters here, more negative number must be first
my_list[-4:-1]
# ### Exercise 1.2
#
# Using the lists you made from up above do the following:
#
# 1. For each list you made before, print the first and last values
# 1. For each multi-dimensional list print the first and last entry of each nested list
# 1. For each one-dimensional list, use a range of indices to make a new sublist
# +
#Number 1
print("Number 1")
Even_numbers=[2,4,6]
print(Even_numbers[0],Even_numbers[-1])
print(Some_integers[0][0], Some_integers[2][-1])
Slang=["dope","swag","gucci","lit"]
print(Slang[0],Slang[-1])
print(Cool_words[0][0], Cool_words[1][-1])
print( )
#NUMBER 2
print("Number 2")
Some_integers=[[1,2,3],[4,5,6],[7,8,9]]
print(Some_integers[0][0], ",", Some_integers[0][-1])
print(Some_integers[1][0], ",", Some_integers[1][-1])
print(Some_integers[2][0], ",", Some_integers[2][-1])
Cool_words=[["yo","wassup","wasspoppin"],["peace","love","pixie_dust"]]
print(Cool_words[0][0], ",", Cool_words[0][-1])
print(Cool_words[1][0], ",", Cool_words[1][-1])
print( )
#NUMBER 3
print("Number 3")
print(Even_numbers[0:-1])
print(Slang[1:-1])
# -
# ## 1.3 Changing lists
# First, we can get the length of a list. Many times, our list is very long or read in from a file. We may need to knwo how long the list actuall is
# Printing the list to remind us of the elements
print(sonoran_plants)
len(sonoran_plants)
# We can change our lists after we make them. We can change individual values or we can add or remove values from a list. Note that tuples cannot be changed (they are called immutable)
# Printing the list to remind us of the elements
print(my_list)
# Individual values in a list can be changed
my_list[2] = -3
print(my_list)
# Values can be added to a list
my_list.append(8)
print(my_list)
# Values can be removed from a list
my_list.remove(-3)
print(my_list)
# A quick note about objects. Python is an object oriented language. It's underlying philosophy is that everything is an object. An object has atributes and methods. You can get information about the attributes and you can use the methods to change the properties of the object.
#
# In python you call the object attributes or methods using this format: `object_variable.attribute` For a list, you add values by changing the attribute `list.append(x)`, `list.remove(x)`.
# We can add the elements of a list to our list
my_list.extend([1,2,3,4])
print(my_list)
# We can insert values at a given index. When using insert, the first value is the index, the second value is the new list element
my_list.insert(0,15)
print(my_list)
# We can remove elements at a given index
my_list.pop(3)
print(my_list)
# ### Exercise 1.3
#
# For each one-dimensional list from above
#
# 1. Append a new element
# 1. Remove a previous element
# 1. Extend the lists with new lists of elements
# 1. Insert a value at the fourth index
# 1. Pop the last value
# +
Even_numbers=[2,4,6]
Slang=["dope","swag","gucci","lit"]
# Number 1
print("Number 1")
Even_numbers.append(8)
print(Even_numbers)
Slang.append("Z<NAME>")
print(Slang)
print( )
# Number 2
print("Number 2")
Even_numbers.remove(2)
print(Even_numbers)
Slang.remove("dope")
print(Slang)
print( )
# Number 3
print("Number 3")
Even_numbers.extend([12,14,16,18])
print(Even_numbers)
Slang.extend(["Dalia, fo'shizzle", "onomatopoeia"])
print(Slang)
print( )
# Number 4
print("Number 4")
Even_numbers.insert(3,10)
print(Even_numbers)
Slang.insert(3,"Yekaterina")
print(Slang)
print( )
# Number 5
print("Number 5")
Even_numbers.pop(-1)
print(Even_numbers)
Slang.pop(-1)
print(Slang)
# -
# ## 1.4 Counting, sorting, and looping methods
# There are a number of other list methods you can use to change your list. To demonstrate these methods, we will make a list of random integers using the append method
# +
# Build a list of random integers
import random # import random number generator library
rand_list=[] # Note this starts as an empty list
for i in range(0,100):
rand_num=random.randrange(0,10)
rand_list.append(rand_num)
print(rand_list)
# -
# We can count the number of times a value is found in our list. This can be really useful for analysis
rand_list.count(3)
# We can determine the index of the first instance of a value
rand_list.index(3)
# The first time that a 3 is found is at list index 0. If you want to keep finding values of 3, you can use a range index to get the other values
rand_list[1:-1].index(3)
rand_list[19:-1].index(3)
# The reason this gives you 1 is that the next instance of 3 is the $20^{\text{th}}$ element $(19+1)$
# We can also sort the list
rand_list.sort()
print(rand_list)
# We can reverse the list
rand_list.reverse()
print(rand_list)
# Lastly, any list can be looped over in python
for plants in sonoran_plants:
print(plants)
# Multidimensional lists can also be looped over. The first loop counts over the nested loops, the second loop counts over the list elements
for collections in sonoran_plants_animals:
for name in collections:
print(name)
# ### Excercise 1.4
# For this exercise, we will creat a list of random number
# +
rand1=[]
for i in range(0,100):
rand_num1=random.randrange(0,10)
rand1.append(rand_num1)
# -
print(rand1)
# Using the list above,
#
# 1. Count and print the nunber of instances of each integer (**Hint:** you can use a loop ranging from 0-9 to do this)
# 2. Loop over the elements of rand1 and make a new list that labels each element as even or odd. For example given the list `[1,6,9]`, you would have a list that looked like `["odd","even","odd"]`.
#
# Hint: You can use the modulo operator `a%b`. This operator give you the remainder when you divide a by b. If `a/b` has no remainder, then `a%b=0` See the examples below.
#
# Modulo example
print(4%2)
print(4%3)
# Module conditional
i=4
if (i%2==0):
print("even")
else:
print("odd")
# Remember that you can always make new cells. Use them to test parts of you code along the way and to seperate your code to make it readable
# +
# Number 1
print("Version #1 w/o loop")
print("Number of 0-")
print(rand.count(0))
print( )
print("Number of 1-")
print(rand.count(1))
print( )
print("Number of 2-")
print(rand.count(2))
print( )
print("Number of 3-")
print(rand.count(3))
print( )
print("Number of 4-")
print(rand.count(4))
print( )
print("Number of 5-")
print(rand.count(5))
print( )
print("Number of 6-")
print(rand.count(6))
print( )
print("Number of 7-")
print(rand.count(7))
print( )
print("Number of 8-")
print(rand.count(8))
print( )
print("Number of 9-")
print(rand.count(9))
print( )
print("Version #2 w/ loop")
for i in range(0,10):
print(rand.count(i))
# -
# Number 2
for i in rand:
print(i)
if (i%2==0):
print("even")
else:
print("odd")
|
comp_assignment-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R-4.0.3
# language: R
# name: ir403
# ---
# +
# Installation of dyno should refer to https://dynverse.org/users/1-installation/
# -
library(dyno)
library(tidyverse)
# # Bifurcation
# ## Run scAAnet (The following code block should be run in Python)
# +
import pandas as pd
import scanpy as sc
from scAAnet.api import scAAnet
data = pd.read_csv("../data/trajectory/bifurcating.csv", index_col=0)
adata = sc.AnnData(data)
sc.pp.normalize_total(adata, target_sum=1)
sc.pp.log1p(adata)
sc.pp.highly_variable_genes(adata, n_top_genes=2000)
K = 4
dist = 'ZIP'
methodmap = {"P": "poisson", "ZIP": "zipoisson", 'NB': 'nb', 'ZINB': 'zinb'}
rs = 1
preds= scAAnet(data.loc[:,adata.var.highly_variable], dispersion='gene-cell',
hidden_size=(128, K, 128), ae_type=methodmap[dist],
epochs=400, batch_size=32,
early_stop=100, reduce_lr=10, learning_rate=0.01,
warm_up=20, random_state=rs)
recon, usage, spectra = preds['recon'], preds['usage'], preds['spectra']
usage = pd.DataFrame(usage, columns = np.arange(1,(K+1)), index=data.index)
usage.to_csv('../data/trajectory/bifurcating.usage.wu20.%s.rs%d.k%s.csv'%(dist, rs, K))
spectra = pd.DataFrame(spectra.transpose(), columns = np.arange(1,(K+1)),
index=data.loc[:,adata.var.highly_variable].columns)
spectra.to_csv('../data/trajectory/bifurcating.program.wu20.%s.rs%d.k%s.csv'%(dist, rs, K))
recon = pd.DataFrame(recon, columns=data.loc[:,adata.var.highly_variable].columns, index=data.index)
recon.to_csv('../data/trajectory/bifurcating.recon.wu20.%s.rs%d.k%s.csv'%(dist, rs, K))
# -
# ## Back to R
bif <- readRDS("../data/trajectory/bifurcating.rds")
usage <- read.csv("../data/trajectory/bifurcating.usage.wu20.ZIP.rs1.k4.csv", row.names = 1)
bif <- bif %>% add_dimred(dyndimred::dimred_umap, expression_source = bif$expression)
for (i in 1:K){
print(plot_dimred(
bif, 'feature',
expression_source = as.matrix(usage),
feature_oi = paste0('X',i)
) + ggtitle(paste0('GEP ', i, ' usage')))
ggsave(sprintf("../figures/simulation/bifurcation3_gep%d.pdf", i))
}
# # Linear
# +
## Run scAAnet (The following code block should be run in Python)
# +
import pandas as pd
import scanpy as sc
from scAAnet.api import scAAnet
data = pd.read_csv("../data/trajectory/linear.csv", index_col=0)
adata = sc.AnnData(data)
sc.pp.normalize_total(adata, target_sum=1)
sc.pp.log1p(adata)
sc.pp.highly_variable_genes(adata, n_top_genes=2000)
K = 3
dist = 'ZIP'
methodmap = {"P": "poisson", "ZIP": "zipoisson", 'NB': 'nb', 'ZINB': 'zinb'}
rs = 1
preds= scAAnet(data.loc[:,adata.var.highly_variable], dispersion='gene-cell',
hidden_size=(128, K, 128), ae_type=methodmap[dist],
epochs=400, batch_size=32,
early_stop=100, reduce_lr=10, learning_rate=0.01,
warm_up=20, random_state=rs)
recon, usage, spectra = preds['recon'], preds['usage'], preds['spectra']
usage = pd.DataFrame(usage, columns = np.arange(1,(K+1)), index=data.index)
usage.to_csv('../data/trajectory/linear.usage.wu20.%s.rs%d.k%s.csv'%(dist, rs, K))
spectra = pd.DataFrame(spectra.transpose(), columns = np.arange(1,(K+1)),
index=data.loc[:,adata.var.highly_variable].columns)
spectra.to_csv('../data/trajectory/linear.program.wu20.%s.rs%d.k%s.csv'%(dist, rs, K))
recon = pd.DataFrame(recon, columns=data.loc[:,adata.var.highly_variable].columns, index=data.index)
recon.to_csv('../data/trajectory/linear.recon.wu20.%s.rs%d.k%s.csv'%(dist, rs, K))
# -
linear <- readRDS("../data/trajectory/linear.rds")
usage <- read.csv("../data/trajectory/linear.usage.wu20.ZIP.rs1.k3.csv", row.names = 1)
linear <- linear %>% add_dimred(dyndimred::dimred_umap, expression_source = linear$expression)
for (i in 1:K){
print(plot_dimred(
linear, 'feature',
expression_source = as.matrix(usage),
feature_oi = paste0('X',i)
) + ggtitle(paste0('GEP ', i, ' usage')))
ggsave(sprintf("../figures/simulation/linear1_gep%d.pdf", i))
}
|
scripts/simulation_trajectory_analysis.ipynb
|
# # Testing model acurracy
# +
import pandas
import matplotlib.pyplot
import numpy
# do ploting inline instead of in a separate window
# %matplotlib inline
#
df = pandas.read_csv("./data/pima-data.csv")
df.shape
# -
# ## Splitting the data
# 70/30 - train/test
# +
from sklearn.model_selection import train_test_split
feature_col_names = ['num_preg', 'glucose_conc', 'diastolic_bp', 'thickness', 'insulin','bmi', 'diab_pred', 'age']
predicted_class_names = ['diabetes']
X = df[feature_col_names].values # predictor feature columns (8 X m)
Y = df[predicted_class_names].values # predicted class (1=True, 0=False) column (1 X m)
split_test_size = 0.3
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=split_test_size, random_state=42)
# test_size = 0.3 is 30% 42 is the answer to everything
# -
# We check to ensure we hav the desired 70% train, 30% test split of the data
from __future__ import division
print "training set: \t {0:0.2f}%".format(len(X_train)/len(df.index) * 100)
print "testing set: \t {0:0.2f}%".format(len(X_test)/len(df.index) * 100)
# Verify the splited data prediction values correlation
print "Original True : {0} ({1:0.2f}%)".format(len(df.loc[df['diabetes'] == 1]), (len(df.loc[df['diabetes'] == 1])/len(df.index)) * 100.0)
print "Original False : {0} ({1:0.2f}%)".format(len(df.loc[df['diabetes'] == 0]), (len(df.loc[df['diabetes'] == 0])/len(df.index)) * 100.0)
print ""
print "Training True : {0} ({1:0.2f}%)".format(len(Y_train[Y_train[:] == 1]), (len(Y_train[Y_train[:] == 1])/len(Y_train) * 100.0))
print "Training False : {0} ({1:0.2f}%)".format(len(Y_train[Y_train[:] == 0]), (len(Y_train[Y_train[:] == 0])/len(Y_train) * 100.0))
print ""
print "Test True : {0} ({1:0.2f}%)".format(len(Y_test[Y_test[:] == 1]), (len(Y_test[Y_test[:] == 1])/len(Y_test) * 100.0))
print "Test False : {0} ({1:0.2f}%)".format(len(Y_test[Y_test[:] == 0]), (len(Y_test[Y_test[:] == 0])/len(Y_test) * 100.0))
# ### Post-split Data Preparation
#
#
# #### Hidding Missing Values
#
#
print "# rows in dataframe {0}".format(len(df))
print "# rows missing glucose_conc: {0}".format(len(df.loc[df['glucose_conc'] == 0]))
print "# rows missing diastolic_bp: {0}".format(len(df.loc[df['diastolic_bp'] == 0]))
print "# rows missing thickness: {0}".format(len(df.loc[df['thickness'] == 0]))
print "# rows missing insulin: {0}".format(len(df.loc[df['insulin'] == 0]))
print "# rows missing bmi: {0}".format(len(df.loc[df['bmi'] == 0]))
print "# rows missing diab_pred: {0}".format(len(df.loc[df['diab_pred'] == 0]))
print "# rows missing age: {0}".format(len(df.loc[df['age'] == 0]))
# Impute with the mean
# +
from sklearn.impute import SimpleImputer as Imputer
# Impute with mean all 0 readings
fill_0 = Imputer(missing_values=0, strategy="mean")
X_train = fill_0.fit(X_train)
X_test = fill_0.fit_transform(X_test)
# -
# ## Training Initial Algorithm - Naive Bayes
# +
from sklearn.naive_bayes import GaussianNB
# create Gaussian Naive Bayes model object and train it with the data
nb_model = GaussianNB()
nb_model.fit(X_train, Y_train.ravel())
# -
# ### Performance on Training Data
# +
# Predict values using the training data
nb_predict_train = nb_model.predict(X_train)
# Import the performance metrics library
from sklearn import metrics
# Accuracy
print "Accuracy: {0:.4f}".format(metrics.accuracy_score(Y_train, nb_predict_train))
print
# -
# ### Performance on Testing Data
# +
# Predict values using the testing data
nb_predict_test = nb_model.predict(X_test)
from sklearn import metrics
print "Accuracy: {0:.4f}".format(metrics.accuracy_score(Y_test, nb_predict_test))
# -
# #### Metrics
print "Confusion Matrix"
print "{}".format(metrics.confusion_matrix(Y_test, nb_predict_test))
print ""
print "[[True Negative] [False Positive]"
print "[False Negative] [True Positive]]"
print ""
print "Classification Report"
print metrics.classification_report(Y_test, nb_predict_test)
# ## Performance Improvement Options
#
# - Adjust current algorithm
# - Get more data or improve data
# - Improve training
# - Switch algorithms
#
# ### Random Forest
#
# - Ensemble Algorithm
# - Fits multiple trees with subsets of data
# - Averages tree results to improve performance and control overfitting
#
#
# +
from sklearn.ensemble import RandomForestClassifier
rf_model = RandomForestClassifier(random_state=42, n_estimators=10)
rf_model.fit(X_train, Y_train.ravel())
# Predict Training Data
rf_predict_train = rf_model.predict(X_train)
# Training metrics
print "Accuracy Training data: {0:.4f}".format(metrics.accuracy_score(Y_train, rf_predict_train))
rf_predict_test = rf_model.predict(X_test)
# Testing metrics
print "Accuracy Testing data: {0:.4f}".format(metrics.accuracy_score(Y_test, rf_predict_test))
# -
print metrics.confusion_matrix(Y_test, rf_predict_test)
print ""
print "Classification Report"
print metrics.classification_report(Y_test, rf_predict_test)
#
|
05.pima-predictions.training-performance.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ASTU Assignment
#
# ## BY-
# ### <NAME>- (170310007051)
#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# DATASET loading
data = pd.read_csv("PCA_practice_dataset.csv",header=None)
data.head()
# ### Covariance matrix of data
cov = np.cov(data)
cov.shape
# ### Eigenvalues and Eigenvectors of covariance matrix
e_val,e_vec = np.linalg.eig(cov)
e_val.shape
eig_vec_ls = []
for i in range(e_vec.shape[1]):
e = np.matrix(data).T@e_vec[:,i]
e = e/e_val[i]
eig_vec_ls.append(np.ravel(e))
# ## Principal Components
sort_index = np.argsort(e_val)
t_val = np.arange(0.9,0.98,0.01)
t_val = t_val.round(2)
sort_index = sort_index[::-1]
e_val_sum = np.sum(e_val)
temp_sum=0
prin_e_val=[]
prin_e_vec=[]
t=0.9
i=0
prin_comp=[]
while t<0.98:
while temp_sum<t*e_val_sum :
prin_e_val.append(e_val[sort_index[i]])
prin_e_vec.append(eig_vec_ls[sort_index[i]])
temp_sum += e_val[sort_index[i]]
i += 1
prin_comp.append(i)
t+=0.01
prin_comp
for i in range(len(t_val)):
print("Number of principal components at threshold {} is {}".format(t_val[i],prin_comp[i]))
# # Scree Plot
# Plotting number of principal components against threshold values
plt.plot(prin_comp,t_val)
plt.title("Scree Plot")
plt.xlabel("Number of principal components")
plt.ylabel("Threshold values")
plt.show()
|
Assignment 1 on pca/ASTU_assignment_on_pca.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# usage: from Portfolio.dataframes import *
# -
from Portfolio.paths import paths
from Portfolio.edit_frontmatter import *
doc = lambda function: print(function.__doc__.strip())
doc(write_frontmatter)
# +
## TOP-LEVEL FUNCTIONS:
def create_DataFrame(header_dict:dict):
"""
Trivially creates a pandas.DataFrame from a heading dictionary.
"""
return pd.DataFrame([header_dict])
def create_header_dict(post:frontmatter.Post, from_dict=None):
"""
create_header_dict(post:frontmatter.Post, from_dict=None)
Takes as inputs a post object, and optionally, a dictionary to being updated.
Returns a header dictionary from a Post YAML Frontmatter and the content.
Params:
post: frontmatter.Post.
from_dict: dict,bool. When it is specified, updates the from_dict with the new post.metadata values.
"""
if isinstance(from_dict, dict):
NEW_HEADER_DICT = from_dict.copy()
else:
NEW_HEADER_DICT = {}
NEW_HEADER_DICT.update({k:v for k,v in post.metadata.items()})
NEW_HEADER_DICT.update({
"metadata" : post.metadata,
"content" : post.content,
})
return NEW_HEADER_DICT
# -
## RUN:
if __name__ == "__main__":
posts_HEADER_TEMPLATE,_,_ = load_frontmatter(os.path.join(paths["postsDir"], "2020-07-03-lc0-guia.md"), False)
portfolio_HEADER_TEMPLATE,_,_ = load_frontmatter(os.path.join(paths["topicsDir"],"astrophysics.md"), False)
drafts_HEADER_TEMPLATE,_,_ = load_frontmatter(os.path.join(paths["draftsDir"], "2020-10-29-draft.md"), False)
posts_HEADER_DICT = create_header_dict(posts_HEADER_TEMPLATE, )
portfolio_HEADER_DICT = create_header_dict(portfolio_HEADER_TEMPLATE, )
drafts_HEADER_DICT = create_header_dict(drafts_HEADER_TEMPLATE, )
postsDF = create_DataFrame(posts_HEADER_DICT)
portfolioDF = create_DataFrame(portfolio_HEADER_DICT)
drafts_DF = create_DataFrame(drafts_HEADER_DICT)
# ## NOTES:
#
# ### FOREACH Approx:
#
# if __name__ == "__main__":
#
# glob_<collection> = glob.glob(os.path.join(paths["<collection>"],"*.md"))
#
# header_dict = {}
# <collection>DF = pd.DataFrame()
# for postfile in glob_<collection>:
# post,_,_ = load_frontmatter( postfile, _parsing=False )
# header_dict = create_header_dict(post, from_dict= header_dict)
# <collection>DF.append([header_dict])
#
#
|
notebooks/.ipynb_checkpoints/dataframes-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Edge detection
#
# <img align="right" width="30%" src="../images/edge_detection_app.png">
#
# With this application, users can create lineaments from gridded data using machine vision algorithms from the [Scikit-Image](https://scikit-image.org/) open-source package.
#
# New user? Visit the [Getting Started](https://geoapps.readthedocs.io/en/latest/content/installation.html) page.
#
# [**Online Documentation**](https://geoapps.readthedocs.io/en/latest/content/applications/edge_detection.html)
#
# [**Youtube demo**](https://youtu.be/Lpn3xA7xlBs)
#
#
# *Click on the cell below and press **Shift+Enter** to run the application*
# +
from geoapps.edge_detection import EdgeDetectionApp
app = EdgeDetectionApp()
app()
# -
# # Pro Tip
#
# Author: <NAME> (<EMAIL>)
#
# "Manual adjustement of each parameter is important to understand how the algorithm works, but finding interesting features manually can rapidly become a tedious process. The script below will let you automate this search by iterating over ranges of parameters. All results will be exported directly to Geoscience ANALYST for analysis.
#
# Enjoy."
#
# \- Phil
# +
x=1
sigmavalues = [4,5]
threshvalues = [1]
linelengthvalues =[5,10]
linegapvalues = [1]
windowsizevalues = [2500]
for sigma in sigmavalues:
for thresh in threshvalues:
for linelength in linelengthvalues:
for linegap in linegapvalues:
for windowsize in windowsizevalues:
print('computing ' + str(x) + ' of ' + str(len(sigmavalues) * len(threshvalues) * len(linelengthvalues) * len(linegapvalues) * len(windowsizevalues)))
x = x+1
app.sigma.value=sigma
app.line_length.value=linelength
app.line_gap.value = linegap
app.threshold.value = thresh
app.window_size.value = windowsize
app.compute.value=True
app.export_as.value=f"Sigma{sigma}_Thresh{thresh}_LineLength{linelength}_LineGap{linegap}_WindowSize{windowsize}"
app.trigger.click()
# -
# Need help? Contact us at <EMAIL>
#
|
geoapps/edge_detection/notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
from context import pypho
import matplotlib.pyplot as plt
# +
hc = 1.240e6
gamma_r = 3
gamma_nr = 0.3
wavelength_range = np.linspace(735, 765, 300)
wavelength0 = 750
order_num = 1
sim = pypho.simulation_1d.new()
sim.stack.define_material('vacuum', 1)
sim.stack.define_material('hBN', 4.7)
sim.stack.define_material('tmd', 1, two_dimensional=True)
sim.stack.define_material('gold', -19.9 + 1.23*1j)
sim.stack.define_material('graphene', 2.59 + 7.2*1j)
sim.stack.add_layers(['vacuum', 'hBN', 'tmd', 'hBN', 'graphene', 'vacuum', 'gold'],
[None, 23.5, None, 17, 3, 380, None])
r = np.zeros_like(wavelength_range)
for i, wavelength in enumerate(wavelength_range):
frequency = 2*np.pi/wavelength
detuning = hc*(1/wavelength - 1/wavelength0)
chi_2d = -gamma_r/frequency/(detuning + 1j*gamma_nr/2)
sim.stack.define_material('tmd', chi_2d)
sim.set_frequency(frequency)
sim.run()
r[i] = np.abs(sim.get_reflection(polarization_in=[1,0], polarization_out=[1,0]))**2
plt.plot(wavelength_range, r);
plt.ylim(0, 1);
plt.xlabel("wavelength (nm)");
plt.ylabel("$R$");
# +
hc = 1.240e6
# in meV
gamma_r = 3
gamma_nr = 0.3
detuning_range = np.linspace(-10, 20, 300)
# in nm
wavelength0 = 750
lattice_constant = 50
# number of Fourier orders
order_num = 50
sim = pypho.simulation_1d.new(order_num=order_num)
sim.stack.set_lattice_constant(lattice_constant)
sim.stack.define_material('vacuum', 1)
sim.stack.define_material('hBN', 4.7)
sim.stack.define_material('tmd', 1, two_dimensional=True)
sim.stack.define_material('silver', -26.1 + 0.60*1j)
sim.stack.define_pattern('grating', ['silver', 'vacuum'], [12.5, 37.5])
sim.stack.add_layers(['vacuum', 'hBN', 'tmd', 'hBN', 'grating', 'silver'], [None, 10, None, 10, 12.5, None])
rs = np.zeros_like(detuning_range)
rp = np.zeros_like(detuning_range)
for i, detuning in enumerate(detuning_range):
frequency = 2*np.pi*(1/wavelength0 + detuning/hc)
chi_2d = -gamma_r/frequency/(detuning + 1j*gamma_nr/2)
sim.stack.define_material('tmd', chi_2d)
sim.set_frequency(frequency)
sim.run()
rs[i] = np.abs(sim.get_reflection(polarization_in=[1,0], polarization_out=[1,0]))**2
rp[i] = np.abs(sim.get_reflection(polarization_in=[0,1], polarization_out=[0,1]))**2
plt.figure(1, figsize=(12, 4))
plt.subplot(121)
plt.plot(detuning_range, rp);
plt.xlabel("detuning (meV)")
plt.ylabel("$R$")
plt.title("x-polarized")
plt.ylim(0, 1.05);
plt.subplot(122)
plt.plot(detuning_range, rs);
plt.ylim(0, 1.05);
plt.xlabel("detuning (meV)");
plt.ylabel("$R$");
plt.title("y-polarized");
plt.show()
# -
|
examples/example_tmd.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Principal Component Regression (PCR)
#
# This notebook implements **Principal Component Regression (PCR)** from scratch and outlines the advantages of PCR over **Multi-variate Linear Regression**.
# ## What is PCR ?
#
# Going by the definition, **Principal Component Regression (PCR)** is a regression algorithm used for datasets having **multi-collinearity**.
#
# It constitutes the following steps:
#
# 1. Perform **Principal Component Analysis (PCA)** on the dataset to reduce the number of features and obtain the **Principal Components (PCs)**.
# 2. Perform **Linear Regression** on the obtained **Principal Components (PCs)**.
# ## When to apply PCR ?
#
# PCR is normally performed on multi-variate datasets having multi-collinearity.
#
# PCR is preferred over **OLS ( Ordinary Least Square ) Linear Regression** on these datasets.
# ## What are the problems with multi-variate Linear Regression?
#
# Multi-variate Linear Regression is Linear Regression with a large number of features.
#
# ### Problems:
#
# 1. In multi-variate Linear Regression, it may appear that **we do fit the model well** but there is normally a **high variance problem** on the test set.
#
# 2. The algorithm takes more time for training the features because of the larger number of input features.
# ## What are the advantages of PCR over OLS Linear Regression?
#
# We saw in the last part that multi-variate Linear Regression poses a problem of high variance and overfits the training set.
#
# PCR reduces the number of features on which regression is performed. It thus **adds a slight bias**. Hence, we now fit the model with a **slightly less training accuracy** but have **reduced the variance to a large extent**.
#
# The time required by the algorithm to train the features on the training set is also **reduced to a large extent**.
# ## Key idea behind PCR:
#
# The key idea behind **PCR** is to **apply PCA on the dataset before regression.**
# ## PCA - Feature Selection or Feature Reduction
#
# **PCA ia not a feature selection technique.** A feature selection technique would involve selecting a few features out of all of them, however we are combining features to create new PCs, which are different from the original features.
|
ml/Dimensionality Reduction Algorithms/Principal_Component_Regression(PCR)/Principal_Component_Regression(PCR).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Discriminant Analysis (LDA) [50 pts]
# In this part of the exercise, you will re-visit the problem of predicting whether a student gets admitted into a university. However, in this part, you will build a linear discriminant analysis (LDA) classifier for this problem.
#
# LDA is a generative model for classification that assumes the class covariances are equal. Given a training dataset of positive and negative features (x, y) with y $\in$ {0, 1} , LDA models the data x as generated from class-conditional Gaussians:
#
# $P(x, y) = P(x|y)P(y)$ where $P(y = 1) = \pi$ and $P(x|y) = N(x;\mu^y,\Sigma)$
#
# where means $\mu^y$ are class-dependent but the covariance matrix $\Sigma$ is class-independent (the same for all classes).
#
# A novel feature $x$ is classified as a positive if $P(y = 1|x) > P(y = 0|x)$, whichis equivalent to $a(x)\gt0$, where the linear classifier $a(x) = w^Tx+w_0$ has weights given by $w = \Sigma^{-1}(\mu^1-\mu^0)$.
#
# In practice, and in this assignment, we use $a(x)\gt$ some threshold, or equivalently, $w^Tx>T$ for some constant $T$.
#
# As we saw in lecture, LDA and logistic regression can be expressed in the same form
#
# $P(y=1|x) = \frac{1}{1+e^{-\theta^Tx}}.$
#
# However, they generally produce different solutions for the parameter theta.
# ## Implementation
#
# In this assignment, you can assume the prior probabilities for the two classes are the same (although the number of the positive and negative samples in the training data is not the same), and that the threshold $T$ is zero. As a bonus, you are encouraged to explore how the different prior probabilities shift the decision boundary.
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
datafile = 'data/ex2data1.txt'
# #!head $datafile
cols = np.loadtxt(datafile,delimiter=',',usecols=(0,1,2),unpack=True) #Read in comma separated data
##Form the usual "X" matrix and "y" vector
X = np.transpose(np.array(cols[:-1]))
y = np.transpose(np.array(cols[-1:]))
m = y.size # number of training examples
##Insert the usual column of 1's into the "X" matrix
X = np.insert(X,0,1,axis=1)
#Divide the sample into two: ones with positive classification, one with null classification
pos = np.array([X[i] for i in range(X.shape[0]) if y[i] == 1])
neg = np.array([X[i] for i in range(X.shape[0]) if y[i] == 0])
def plotData():
plt.figure(figsize=(10,6))
plt.plot(pos[:,1],pos[:,2],'k+',label='Admitted')
plt.plot(neg[:,1],neg[:,2],'yo',label='Not admitted')
plt.xlabel('Exam 1 score')
plt.ylabel('Exam 2 score')
plt.legend()
plt.grid(True)
plotData()
# -
# Implement the LDA classifier by completing the code here. As an implementation detail, you should first center the positive and negative data separately, so that each has a mean equal to 0, before computing the covariance, as this tends to give a more accurate estimate.
#
# You should center the whole training data set before applying the classifier. Namely, subtract the middle value of the two classes’ means ($\frac{1}{2}$(pos mean+neg mean)), which is on the separating plane when their prior probabilities are the same and becomes the ‘center’ of the data. [5 pts]
# +
# IMPLEMENT THIS
pos_mean = np.mean(pos,axis=0)
neg_mean = np.mean(neg,axis=0)
print(pos_mean,neg_mean)
pos_data = pos - 0.5*(pos_mean+neg_mean)
neg_data = neg - 0.5*(pos_mean+neg_mean)
plt.figure(figsize=(10,6))
plt.plot(pos_data[:,1],pos_data[:,2],'k+',label='Admitted')
plt.plot(neg_data[:,1],neg_data[:,2],'yo',label='Not admitted')
plt.xlabel('Exam 1 score')
plt.ylabel('Exam 2 score')
plt.legend()
plt.grid(True)
# -
# Implement the LDA algorithm here (Compute the covariance on all data): [10 pts each for getting cov_all, w and y_lda]
# +
# IMPLEMENT THIS
X_data = np.concatenate((neg_data,pos_data),axis=0)
label = np.sort(y,axis=None)
cov_all = np.cov(pos_data.T[1:]) + np.cov(neg_data.T[1:]) # SHAPE: (2,2)
print('cov_all shape:',cov_all.shape)
w = np.linalg.inv(cov_all)@((pos_mean-neg_mean)[1:]) # w=cov_all^(-1)(pos_mean-neg_mean)
print('w:',w)
y_lda = np.dot(X_data[:,1:],w) # SHAPE: (100,)
print('y_lda shape:',y_lda.shape)
# -
# Completing the code to compute the training set accuracy. You should get a training accuracy around 89%. [5 pts]
# +
# IMPLEMENT THIS
y_lda[y_lda<0]=0
y_lda[y_lda>0]=1
print(y_lda)
count = 0
for i in range(m):
if label[i] == y_lda[i]:
count+=1
accuracy = count/m
print(accuracy)
# -
# ## Written Problem [10 pts]
#
# Show that the log-odds decision function a(x) for LDA
#
# $a = \ln \frac{p(x|C_l)p(C_l)}{p(x|C_k)p(C_k)}$
#
# is linear in x, that is, we can express $a(x)=\theta^Tx$ for some $\theta$. Show all your steps. Hint: This is a binary problem.
# #### ANSWER:
#
# We notice that this is a binary problem. So that there are two classes: class l and k.
#
# By definition, the class-conditional density is multi-Gaussian:
# $$ p(x|C_k) = \frac{1}{(2\pi)^{p/2}|\Sigma_k|^{1/2}}e^{-\frac{1}{2}(x-\mu_k)^{T}\Sigma_k^{-1}(x-\mu_k)}$$
#
# And we assume $p(C_l)=p(C_k)=\frac{1}{2}$, $\Sigma=\Sigma_l=\Sigma_k$
#
# \begin{align*}
# a &=\log \frac{p(x|C_l)p(C_l)}{p(x|C_k)p(C_k)} \\
# &= \log p(x|C_l)-\log p(x|C_k)+\log \frac{p(C_l)}{p(C_k)} \\
# &= \log p(x|C_l)-\log p(x|C_k) \\
# &= -\log (2\pi)^{p/2}|\Sigma_l|^{1/2}-\frac{1}{2}(x-\mu_l)^{T}\Sigma_l^{-1}(x-\mu_l)+ \log (2\pi)^{p/2}|\Sigma_k|^{1/2}+\frac{1}{2}(x-\mu_k)^{T}\Sigma_k^{-1}(x-\mu_k)\\
# &= -\log (2\pi)^{p/2}|\Sigma|^{1/2}-\frac{1}{2}(x-\mu_l)^{T}\Sigma^{-1}(x-\mu_l)+ \log (2\pi)^{p/2}|\Sigma|^{1/2}+\frac{1}{2}(x-\mu_k)^{T}\Sigma^{-1}(x-\mu_k)\\
# &= -\frac{1}{2}(x-\mu_l)^{T}\Sigma^{-1}(x-\mu_l)+\frac{1}{2}(x-\mu_k)^{T}\Sigma^{-1}(x-\mu_k)\\
# &= -\frac{1}{2}(x^T\Sigma^{-1}x-2x^T\Sigma^{-1}\mu_l+\mu_l^T\Sigma^{-1}\mu_l)+\frac{1}{2}(x^T\Sigma^{-1}x-2x^T\Sigma^{-1}\mu_k+\mu_k^T\Sigma^{-1}\mu_k)\\
# &= -\frac{1}{2}(x^T\Sigma^{-1}x-2x^T\Sigma^{-1}\mu_l+\mu_l^T\Sigma^{-1}\mu_l-x^T\Sigma^{-1}x+2x^T\Sigma^{-1}\mu_k-\mu_k^T\Sigma^{-1}\mu_k)\\
# &= -\frac{1}{2}(-2x^T\Sigma^{-1}\mu_l+2x^T\Sigma^{-1}\mu_k+\mu_l^T\Sigma^{-1}\mu_l-\mu_k^T\Sigma^{-1}\mu_k) \\
# &= -\frac{1}{2}(-2x^T\Sigma^{-1}(\mu_l-\mu_k)+\mu_l^T\Sigma^{-1}\mu_l-\mu_k^T\Sigma^{-1}\mu_k) \\
# &= x^T\Sigma^{-1}(\mu_l-\mu_k) + const
# \end{align*}
#
# which is linear in x. So that we can express $a(x)=\theta^T x + \theta_0$, $\theta = \Sigma^{-1}(\mu_l-\mu_k)$, $\theta_0$ is the const.
#
# # CNN on MNIST using TensorFlow™ [50 pts]
# **Note 1**: The following has been verified to work with the current latest version of TensorFlow (1.11)
#
# \* Adapted from official TensorFlow™ tour guide.
#
# TensorFlow is a powerful library for doing large-scale numerical computation. One of the tasks at which it excels is implementing and training deep neural networks. In this assignment you will learn the basic building blocks of a TensorFlow model while constructing a deep convolutional MNIST classifier.
#
# What you are expected to implement in this tutorial:
#
# * Create a softmax regression function that is a model for recognizing MNIST digits, based on looking at every pixel in the image
#
# * Use Tensorflow to train the model to recognize digits by having it "look" at thousands of examples
#
# * Check the model's accuracy with MNIST test data
#
# * Build, train, and test a multilayer convolutional neural network to improve the results
#
# Here is a diagram, created with TensorBoard, of the model we will build:
#
# 
#
# ## Implement Utilities
#
# ### Weight Initialization
#
# To create this model, we're going to need to create a lot of weights and biases. One should generally initialize weights with a small amount of noise for symmetry breaking, and to prevent 0 gradients. Since we're using ReLU neurons, it is also good practice to initialize them with a slightly positive initial bias to avoid "dead neurons". Instead of doing this repeatedly while we build the model, let's create two handy functions to do it for us.
#
# +
import tempfile
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
tf.logging.set_verbosity(tf.logging.ERROR)
def weight_variable(shape):
"""weight_variable generates a weight variable of a given shape."""
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial)
def bias_variable(shape):
"""bias_variable generates a bias variable of a given shape."""
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial)
# -
# ### Convolution and Pooling [5 pts]
#
# Our convolutions uses a stride of one and are zero padded so that the output is the same size as the input. Our pooling is plain old max pooling over 2x2 blocks.
#
# NOTE: FOR ALL THE FOLLOWING CODES, DO NOT IMPLEMENT YOUR OWN VERSION. USE THE BUILT-IN METHODS FROM TENSORFLOW.
#
# Take a look at [TensorFlow API Docs](https://www.tensorflow.org/api_docs/python/).
# +
# IMPLEMENT THIS
def conv2d(x, W):
conv2d = tf.layers.conv2d(inputs=x,
filters=W,
kernel_size=[5,5],
padding="same",
)
return conv2d
def max_pool_2x2(x):
max_pool = tf.layers.max_pooling2d(inputs=x,
pool_size=[2,2],
strides=2)
return max_pool
# -
# ## Build the CNN
#
# ### First Convolutional Layer[10 pts]
#
# We can now implement our first layer. It will consist of convolution, followed by max pooling. The convolution will compute 32 features for each 5x5 patch. Its weight tensor will have a shape of [5, 5, 1, 32]. The first two dimensions are the patch size, the next is the number of input channels, and the last is the number of output channels. We will also have a bias vector with a component for each output channel.
#
# To apply the layer, we first reshape x to a 4d tensor, with the second and third dimensions corresponding to image width and height, and the final dimension corresponding to the number of color channels.
#
# We then convolve x_image with the weight tensor, add the bias, apply the ReLU function, and finally max pool. The max_pool_2x2 method will reduce the image size to 14x14.
#
# ### Second Convolutional Layer[5 pts]
#
# In order to build a deep network, we stack several layers of this type. The second layer will have 64 features for each 5x5 patch.
#
# ### Fully Connected Layer[10 pts]
#
# Now that the image size has been reduced to 7x7, we add a fully-connected layer with 1024 neurons to allow processing on the entire image. We reshape the tensor from the pooling layer into a batch of vectors, multiply by a weight matrix, add a bias, and apply a ReLU.
#
# ### SoftmaxLayer[5 pts]
#
# Finally, we add a layer of softmax regression.
def deepnn(x):
"""
deepnn builds the graph for a deep net for classifying digits.
Args:
x: an input tensor with the dimensions (N_examples, 784), where 784 is the
number of pixels in a standard MNIST image.
Returns:
A tuple (y, keep_prob). y is a tensor of shape (N_examples, 10), with values
equal to the logits of classifying the digit into one of 10 classes (the
digits 0-9). keep_prob is a scalar placeholder for the probability of
dropout.
"""
# Reshape to use within a convolutional neural net.
# Last dimension is for "features" - there is only one here, since images are
# grayscale -- it would be 3 for an RGB image, 4 for RGBA, etc.
with tf.name_scope('reshape'):
x_image = tf.reshape(x,[-1,28,28,1])
# First convolutional layer - maps one grayscale image to 32 feature maps.
with tf.name_scope('conv1'):
h_conv1 = tf.nn.relu(conv2d(x_image,32))
# Pooling layer - downsamples by 2X.
with tf.name_scope('pool1'):
h_pool1 = max_pool_2x2(h_conv1)
# Second convolutional layer -- maps 32 feature maps to 64.
with tf.name_scope('conv2'):
h_conv2 = tf.nn.relu(conv2d(h_pool1,64))
# Second pooling layer.
with tf.name_scope('pool2'):
h_pool2 = max_pool_2x2(h_conv2)
# Fully connected layer 1 -- after 2 round of downsampling, our 28x28 image
# is down to 7x7x64 feature maps -- maps this to 1024 features.
with tf.name_scope('fc1'):
h_pool2_flat = tf.contrib.layers.flatten(h_pool2,scope='pool2flat')
h_fc1 = tf.layers.dense(inputs=h_pool2_flat,units=1024,activation=tf.nn.relu)
# Map the 1024 features to 10 classes, one for each digit
with tf.name_scope('fc2'):
y_conv = tf.layers.dense(inputs=h_fc1,units=10)
return y_conv
# ## Complete the Graph[10 pts]
# We start building the computation graph by creating nodes for the input images and target output classes.
# +
# Import data
mnist = input_data.read_data_sets('/tmp/tensorflow/mnist/input_data', one_hot=True)
x = tf.placeholder(tf.float32, [None, 784])
y_ = tf.placeholder(tf.int32, [None, 10])
# Build the graph for the deep net
y_conv = deepnn(x)
# -
# We can specify a loss function just as easily. Loss indicates how bad the model's prediction was on a single example; we try to minimize that while training across all the examples. Here, our loss function is the cross-entropy between the target and the softmax activation function applied to the model's prediction. As in the beginners tutorial, we use the stable formulation:
# +
with tf.name_scope('loss'):
y_dict = dict(labels=y_,logits=y_conv)
losses = tf.nn.softmax_cross_entropy_with_logits(**y_dict)
cross_entropy = tf.reduce_mean(losses)
with tf.name_scope('adam_optimizer'):
trainer = tf.train.AdamOptimizer(learning_rate=0.001)
train_step = trainer.minimize(cross_entropy)
# -
# First we'll figure out where we predicted the correct label. tf.argmax is an extremely useful function which gives you the index of the highest entry in a tensor along some axis. For example, tf.argmax(y,1) is the label our model thinks is most likely for each input, while tf.argmax(y\_,1) is the true label. We can use tf.equal to check if our prediction matches the truth.
#
# That gives us a list of booleans. To determine what fraction are correct, we cast to floating point numbers and then take the mean. For example, [True, False, True, True] would become [1,0,1,1] which would become 0.75.
with tf.name_scope('accuracy'):
y_pred = tf.argmax(tf.nn.softmax(y_conv),axis=1)
y_true = tf.argmax(y_,axis=1)
correct_prediction = tf.equal(tf.cast(y_pred,tf.int64),y_true)
accuracy = tf.reduce_mean(tf.cast(correct_prediction,tf.float32))
# For saving the graph, DO NOT CHANGE.
graph_location = r'./graph'
print('Saving graph to: %s' % graph_location)
train_writer = tf.summary.FileWriter(graph_location)
train_writer.add_graph(tf.get_default_graph())
# ## Train and Evaluate the Model[5 pts]
#
# We will use a more sophisticated ADAM optimizer instead of a Gradient Descent Optimizer.
#
# We will add logging to every 100th iteration in the training process.
#
# Feel free to run this code. Be aware that it does 20,000 training iterations and may take a while (possibly up to half an hour), depending on your processor.
#
# The final test set accuracy after running this code should be approximately 99.2%.
#
# We have learned how to quickly and easily build, train, and evaluate a fairly sophisticated deep learning model using TensorFlow.
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for i in range(2000):
batch = mnist.train.next_batch(50)
if i % 100 == 0:
train_accuracy = accuracy.eval(feed_dict={x: batch[0], y_: batch[1]})
print('step %d, training accuracy %g' % (i, train_accuracy))
train_step.run(feed_dict={x: batch[0], y_: batch[1]})
print('test accuracy %g' % accuracy.eval(feed_dict={x: mnist.test.images, y_: mnist.test.labels}))
|
ps5/pset5_lda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Coopetition - Muon id classification
#
# <NAME>, <NAME>
#
#
# [x] Select features and find new features \
# [ ] Add Scaling to wide range variables \
# [ ] CatBoost \
# [x] Scale weight - NOT USED
#
# -> Need the module swifter with fsspec==0.3.3 , if the version is newer than this, the code might break
#
#
# ### Import part
#
# +
import os
import pandas as pd
import xgboost
import utils
import scoring
from sklearn.model_selection import train_test_split
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### Download dataset
# +
# The datasets are available in CoCalc in ~/share/data/I-coopetition-muon-id/
# Test
# # ! wget --content-disposition https://codalab.coresearch.club/my/datasets/download/dd6255a1-a14b-4276-9a2b-db7f360e01c7
# Train
# # ! wget --content-disposition https://codalab.coresearch.club/my/datasets/download/3a5e940c-2382-4716-9ff7-8fbc269b98ac
# -
# ### Step 1 Load Data
DATA_PATH = "~/share/data/I-coopetition-muon-id/"
columns = utils.SIMPLE_FEATURE_COLUMNS + ["id", "label", "weight", "sWeight", "kinWeight"]
train = utils.load_full_train_csv(DATA_PATH, 600000)
print(train.shape)
# ### Step 2 Data Preprocessing
# +
# Main Setting
weight_name = "weight"
num_sample = 10
train_vis = train.head(num_sample)
# +
'''
Utils functions
'''
def visualize(feature, target, weights, num_bins=100):
classes = np.unique(target)
bins = np.linspace(feature.min(), feature.max(), num_bins + 1)
# Plot all class
for c in classes:
selection = (target == c)
plt.hist(feature[selection], bins = bins, label = c, alpha = 0.5, weights = weights[selection])
plt.legend()
# -
print(train.columns)
# +
# Looking for high level parameters
# Get closest hits from utils
import swifter
def Get_closest_hits(data):
closest_hits_features = data.swifter.apply(utils.find_closest_hit_per_station, result_type="expand", axis=1)
closest_hits_features.columns = ["closest_{}".format(ind) for ind in range(len(closest_hits_features.columns))]
return closest_hits_features
close_hits = Get_closest_hits(train)
train_mod = pd.concat([train, close_hits], axis = 1)
# Save to files for backing up
close_hits.to_pickle('train_closest_hit.pkl')
# +
# Scale product of sWeight - considered NOT USED as we don't have an access in submission test
import pandas as pd
pd.options.mode.chained_assignment = None
def scale_weight(data, edge_weight = None):
if edge_weight != None:
data_mod = data[data['weight'].abs() < edge_weight]
data_mod['scale_weight'] = (data['weight'] + edge_weight)/(2 * edge_weight)
data_mod['scale_weight'] = data_mod['scale_weight'] * (data_mod['weight'].sum() / data_mod['scale_weight'].sum())
return data_mod
else:
data_mod = data
data_mod['scale_weight'] = (data_mod['weight'] + data_mod['weight'].min())
data_mod['scale_weight'] *= (data_mod['weight'].sum() / data_mod['scale_weight'].sum())
return data_mod
# Scale sweight
#train_mod = scale_weight(train_mod, edge_weight = None)
# +
# Add new features
def feat_PZ(data):
return np.sqrt(data['P'] ** 2 - data['PT'] ** 2)
def feat_eta(data):
P = data['P']
PZ = data['PZ']
return -0.5 * np.log((1.0 - ( PZ / P )) / (1.0 + (PZ / P)))
train_mod['PZ'] = feat_PZ(train_mod)
train_mod['eta'] = feat_eta(train_mod)
#train_mod['P']/train_mod['PZ']
# +
# Visualisation
for column in []:
plt.figure()
visualize(train[column][:num_sample], train['label'][:num_sample], train[weight_name][:num_sample])
plt.title(column)
# -
# ## Note:
# - Acc = 0.70, train_cols with close hit, add PZ, Model: XGBoost LR = 0.1 with scale weight
# - Acc = 0.72, train_cols with close hit, add PZ, Model: XGBoost LR = 0.1
# - Acc = 0.70, train_cols with close hit, add PZ, eta, Model: XGBoost LR = 0.1 with scale weight
# Def used columns
train_cols = ['ncl[0]', 'ncl[1]', 'ncl[2]', 'ncl[3]', 'avg_cs[0]', 'avg_cs[1]',
'avg_cs[2]', 'avg_cs[3]', 'ndof', 'MatchedHit_TYPE[0]',
'MatchedHit_TYPE[1]', 'MatchedHit_TYPE[2]', 'MatchedHit_TYPE[3]',
'MatchedHit_X[0]', 'MatchedHit_X[1]', 'MatchedHit_X[2]',
'MatchedHit_X[3]', 'MatchedHit_Y[0]', 'MatchedHit_Y[1]',
'MatchedHit_Y[2]', 'MatchedHit_Y[3]', 'MatchedHit_Z[0]',
'MatchedHit_Z[1]', 'MatchedHit_Z[2]', 'MatchedHit_Z[3]',
'MatchedHit_DX[0]', 'MatchedHit_DX[1]', 'MatchedHit_DX[2]',
'MatchedHit_DX[3]', 'MatchedHit_DY[0]', 'MatchedHit_DY[1]',
'MatchedHit_DY[2]', 'MatchedHit_DY[3]', 'MatchedHit_DZ[0]',
'MatchedHit_DZ[1]', 'MatchedHit_DZ[2]', 'MatchedHit_DZ[3]',
'MatchedHit_T[0]', 'MatchedHit_T[1]', 'MatchedHit_T[2]',
'MatchedHit_T[3]', 'MatchedHit_DT[0]', 'MatchedHit_DT[1]',
'MatchedHit_DT[2]', 'MatchedHit_DT[3]', 'NShared', 'Mextra_DX2[0]',
'Mextra_DX2[1]', 'Mextra_DX2[2]', 'Mextra_DX2[3]', 'Mextra_DY2[0]',
'Mextra_DY2[1]', 'Mextra_DY2[2]', 'Mextra_DY2[3]', 'FOI_hits_N',
'PT', 'PZ'] + close_hits.columns.tolist()
target_col = ['label', 'weight']
# +
import re
def prepare_data(data):
regex = re.compile(r"\[|\]|<", re.IGNORECASE)
data.columns = [regex.sub("_", col) if any(x in str(col) for x in set(('[', ']', '<'))) else col for col in data.columns.values]
# +
# Check Heading data
train.head()
X_mod, y_mod = train_mod[train_cols], train_mod[target_col]
# Rename - ignore []
prepare_data(X_mod)
# Splitting
X_train, X_val, y_train, y_val = train_test_split(X_mod, y_mod, test_size=0.25, shuffle=True, random_state=2342234)
# Test shift the weight on training
y_train = scale_weight(y_train) # Add ['scale_weight'] - shifting weight
# Scale
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
'''
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_val = scaler.transform(X_val)
pca = PCA()
X_train = pca.fit_transform(X_train)
X_val = pca.transform(X_val)
'''
# -
print(X_train)
# ### Step 3 Training part
# +
# Model Arch
import catboost
n_trees = 500
# +
from sklearn.metrics import accuracy_score
import joblib
# Trainning part
score_best = 0
model_best = None
lr_best = None
lrs = [1, 0.5, 0.1, 0.08]
for lr in lrs:
print("trainning with lr = {}".format(lr))
model = catboost.CatBoostClassifier(iterations=n_trees, verbose=False, thread_count=-1, learning_rate = lr)
model.fit(X_train.iloc[:, :],
y_train['label'].values,
#sample_weight=y_train.scale_weight.values,
#verbose=True,
)
'''
train with scaler and PCA
model.fit(X_train,
y_train['label'].values,
sample_weight=y_train.weight.values,
verbose=True,
)
'''
validation_predictions = model.predict_proba(X_val)[:, 1]
model_score = scoring.rejection90(y_val.label.values, validation_predictions, sample_weight = y_val.weight.values)
print("NN: {} , Test accuracy: {}".format(lr, model_score))
if model_score > score_best :
model_best = model
score_best = model_score
lr_best = lr
# Save Model
joblib.dump(model_best, 'Model_Best.pkl')
# -
print(score_best)
validation_predictions = model.predict_proba(X_val)[:, 1]
model_score = scoring.rejection90(y_val.label.values, validation_predictions, sample_weight = y_val.weight.values)
print(model_score)
# +
# Further test on 20% of original training
Eval_train = train.head(int(0.2 * train.shape[0]))
# Prep sample
close_hits = Get_closest_hits(Eval_train)
Eval_train = pd.concat([Eval_train, close_hits], axis = 1)
Eval_train['PZ'] = feat_PZ(Eval_train)
Eval_train['eta'] = feat_eta(Eval_train)
Eval_X = Eval_train[train_cols]
prepare_data(Eval_X)
# Predict and save file
Eval_true = Eval_train[['label']]
predictions = model_best.predict_proba(Eval_X)[:, 1]
model_score = scoring.rejection90(Eval_true.label.values, predictions, Eval_train.weight.values)
# -
print(model_score)
# ### Step 4 Predict on the whole test set and prepare submission
#
#
# +
# Read data
DATA_PATH = "~/share/data/I-coopetition-muon-id/"
test = utils.load_full_test_csv(DATA_PATH, None)
# Transform data
close_hits_test = Get_closest_hits(test)
test_mod = pd.concat([test, close_hits_test], axis = 1)
test_mod['PZ'] = feat_PZ(test_mod)
X_test = test_mod[train_cols]
prepare_data(X_test)
# Predict and save file
predictions = model_best.predict_proba(X_test)[:, 1]
compression_opts = dict(method='zip',
archive_name='submission.csv')
pd.DataFrame(data={"prediction": predictions}, index=test.index).to_csv(
"submission.zip", index_label=utils.ID_COLUMN, compression=compression_opts)
# -
|
model_muon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 04 : VGG architecture - exercise
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
from google.colab import drive
drive.mount('/content/gdrive')
file_name = 'vgg_exercise.ipynb'
import subprocess
path_to_file = subprocess.check_output('find . -type f -name ' + str(file_name), shell=True).decode("utf-8")
print(path_to_file)
path_to_file = path_to_file.replace(file_name,"").replace('\n',"")
os.chdir(path_to_file)
# !pwd
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from random import randint
import utils
import time
# ### With or without GPU?
#
# It is recommended to run this code on GPU:<br>
# * Time for 1 epoch on CPU : 841 sec (14.02 min)<br>
# * Time for 1 epoch on GPU : 9 sec w/ GeForce GTX 1080 Ti <br>
device= torch.device("cuda")
device= torch.device("cpu")
print(device)
# ### Download the CIFAR dataset
# +
from utils import check_cifar_dataset_exists
data_path=check_cifar_dataset_exists()
train_data=torch.load(data_path+'cifar/train_data.pt')
train_label=torch.load(data_path+'cifar/train_label.pt')
test_data=torch.load(data_path+'cifar/test_data.pt')
test_label=torch.load(data_path+'cifar/test_label.pt')
print(train_data.size())
print(test_data.size())
# -
# ### Compute mean pixel intensity over all training set and all channels
# +
mean= train_data.mean()
print(mean)
# -
# ### Compute standard deviation
# +
std= train_data.std()
print(std)
# -
# ### Make a VGG convnet class.
class VGG_convnet(nn.Module):
def __init__(self):
super(VGG_convnet, self).__init__()
# block 1: 3 x 32 x 32 --> 64 x 16 x 16
self.conv1a = nn.Conv2d(3, 64, kernel_size=3, padding=1 )
self.conv1b = nn.Conv2d(64, 64, kernel_size=3, padding=1 )
self.pool1 = nn.MaxPool2d(2,2)
# block 2: 64 x 16 x 16 --> 128 x 8 x 8
self.conv2a = # COMPLETE HERE
self.conv2b = # COMPLETE HERE
self.pool2 = # COMPLETE HERE
# block 3: 128 x 8 x 8 --> 256 x 4 x 4
self.conv3a = # COMPLETE HERE
self.conv3b = # COMPLETE HERE
self.pool3 = # COMPLETE HERE
#block 4: 256 x 4 x 4 --> 512 x 2 x 2
self.conv4a = # COMPLETE HERE
self.pool4 = # COMPLETE HERE
# linear layers: 512 x 2 x 2 --> 2048 --> 4096 --> 4096 --> 10
self.linear1 = # COMPLETE HERE
self.linear2 = # COMPLETE HERE
self.linear3 = # COMPLETE HERE
def forward(self, x):
# block 1: 3 x 32 x 32 --> 64 x 16 x 16
x = self.conv1a(x)
x = F.relu(x)
x = self.conv1b(x)
x = F.relu(x)
x = self.pool1(x)
# block 2: 64 x 16 x 16 --> 128 x 8 x 8
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
# block 3: 128 x 8 x 8 --> 256 x 4 x 4
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
#block 4: 256 x 4 x 4 --> 512 x 2 x 2
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
# linear layers: 512 x 2 x 2 --> 2048 --> 4096 --> 4096 --> 10
x = x.view(# COMPLETE HERE)
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
x = # COMPLETE HERE
return x
# ### Build the net. How many parameters in total? (the three layer net had 2 million parameters)
# +
net = # COMPLETE HERE
print(net)
utils.display_num_param(net)
# -
# ### Send the weights of the networks to the GPU (as well as the mean and std)
# +
net = # COMPLETE HERE
mean = mean.to(device)
std = std.to(device)
# -
# ### Choose the criterion, batch size, and initial learning rate. Select the following:
# * batch size =128
# * initial learning rate =0.25
#
criterion = nn.CrossEntropyLoss()
my_lr= # COMPLETE HERE
bs= # COMPLETE HERE
# ### Function to evaluate the network on the test set
def eval_on_test_set():
running_error=0
num_batches=0
for i in range(0,10000,bs):
minibatch_data = test_data[i:i+bs]
minibatch_label= test_label[i:i+bs]
minibatch_data=minibatch_data.to(device)
minibatch_label=minibatch_label.to(device)
inputs = (minibatch_data - mean)/std
scores=net( inputs )
error = utils.get_error( scores , minibatch_label)
running_error += error.item()
num_batches+=1
total_error = running_error/num_batches
print( 'error rate on test set =', total_error*100 ,'percent')
# ### Do 20 passes through the training set. Divide the learning rate by 2 at epoch 10, 14 and 18.
# +
start=time.time()
for epoch in range(1,20):
# divide the learning rate by 2 at epoch 10, 14 and 18
if # COMPLETE HERE
my_lr = # COMPLETE HERE
# create a new optimizer at the beginning of each epoch: give the current learning rate.
optimizer=torch.optim.SGD( net.parameters() , lr=my_lr )
# set the running quatities to zero at the beginning of the epoch
running_loss=0
running_error=0
num_batches=0
# set the order in which to visit the image from the training set
shuffled_indices=torch.randperm(50000)
for count in range(0,50000,bs):
# Set the gradients to zeros
optimizer.zero_grad()
# create a minibatch
indices=shuffled_indices[count:count+bs]
minibatch_data = train_data[indices]
minibatch_label= train_label[indices]
# send them to the gpu
minibatch_data=minibatch_data.to(device)
minibatch_label=minibatch_label.to(device)
# normalize the minibatch (this is the only difference compared to before!)
inputs = (minibatch_data - mean)/std
# tell Pytorch to start tracking all operations that will be done on "inputs"
inputs.requires_grad_()
# forward the minibatch through the net
scores=net( inputs )
# Compute the average of the losses of the data points in the minibatch
loss = criterion( scores , minibatch_label)
# backward pass to compute dL/dU, dL/dV and dL/dW
loss.backward()
# do one step of stochastic gradient descent: U=U-lr(dL/dU), V=V-lr(dL/dU), ...
optimizer.step()
# START COMPUTING STATS
# add the loss of this batch to the running loss
running_loss += loss.detach().item()
# compute the error made on this batch and add it to the running error
error = utils.get_error( scores.detach() , minibatch_label)
running_error += error.item()
num_batches+=1
# compute stats for the full training set
total_loss = running_loss/num_batches
total_error = running_error/num_batches
elapsed = (time.time()-start)/60
print('epoch=',epoch, '\t time=', elapsed,'min','\t lr=', my_lr ,'\t loss=', total_loss , '\t error=', total_error*100 ,'percent')
eval_on_test_set()
print(' ')
# -
# ### Choose image at random from the test set and see how good/bad are the predictions
# +
# choose a picture at random
idx=randint(0, 10000-1)
im=test_data[idx]
# diplay the picture
utils.show(im)
# send to device, rescale, and view as a batch of 1
im = im.to(device)
im= (im-mean) / std
im=im.view(1,3,32,32)
# feed it to the net and display the confidence scores
scores = net(im)
probs= F.softmax(scores, dim=1)
utils.show_prob_cifar(probs.cpu())
|
codes/labs_lecture08/lab04_vgg/vgg_exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import networkx as nx
import matplotlib.pyplot as plt
from graphein.construct_graphs import PPIGraph
# # Plot small PPI
# +
protein_list = ["CDC42", "CDK1", "KIF23", "PLK1", "RAC2", "RACGAP1", "RHOA", "RHOB"]
sources = ["STRING", "BIOGRID"]
kwargs = {"STRING_escore": 0.2, # Keeps STRING interactions with an experimental score >= 0.2
"BIOGRID_throughputTag": "high" # Keeps high throughput BIOGRID interactions
}
ppi_graph = PPIGraph(protein_list=protein_list,
sources=sources,
**kwargs)
# BIOGRID
df = ppi_graph._parse_BIOGRID(protein_list, 9606, **kwargs)
assert (df["THROUGHPUT"].str.contains("High Throughput")).all()
# STRING
df = ppi_graph._parse_STRING(protein_list=ppi_graph.protein_list,
ncbi_taxon_id=ppi_graph.ncbi_taxon_id)
df = ppi_graph._filter_STRING(df, **ppi_graph.kwargs)
assert (df["escore"] >= 0.2).all()
# Plot output network
nx_graph = ppi_graph.nx_graph()
nx_pos = nx.spring_layout(nx_graph)
nx.draw(nx_graph, nx_pos, with_labels=True, node_color="lightblue")
# -
# # Plot SARS-CoV-2 host-virus proteins (Gordon et al., 2020)
# List of human interacting proteins (`protein_list`) retrieved from:
#
# Gordon et al. (2020). A SARS-CoV-2-Human Protein-Protein Interaction Map Reveals Drug Targets and Potential Drug-Repurposing. [[paper](https://www.biorxiv.org/content/10.1101/2020.03.22.002386v2)]
# +
protein_list = ['AP3B1', 'BRD4', 'BRD2', 'CWC27', 'ZC3H18', 'SLC44A2', 'PMPCB',
'YIF1A', 'ATP1B1', 'ACADM', 'ETFA', 'STOM', 'GGCX', 'ATP6V1A',
'PSMD8', 'REEP5', 'PMPCA', 'ANO6', 'PITRM1', 'SLC30A9', 'FASTKD5',
'SLC30A7', 'TUBGCP3', 'COQ8B', 'SAAL1', 'REEP6', 'INTS4',
'SLC25A21', 'TUBGCP2', 'TARS2', 'RTN4', 'FAM8A1', 'AASS', 'AKAP8L',
'AAR2', 'BZW2', 'RRP9', 'PABPC1', 'CSNK2A2', 'CSNK2B', 'G3BP1',
'PABPC4', 'LARP1', 'FAM98A', 'SNIP1', 'UPF1', 'MOV10', 'G3BP2',
'DDX21', 'RBM28', 'RPL36', 'GOLGA7', 'ZDHHC5', 'POLA1', 'PRIM1',
'PRIM2', 'POLA2', 'COLGALT1', 'PKP2', 'AP2A2', 'GFER', 'ERGIC1',
'AP2M1', 'GRPEL1', 'TBCA', 'SBNO1', 'BCKDK', 'AKAP8', 'MYCBP2',
'SLU7', 'RIPK1', 'UBAP2L', 'TYSND1', 'PDZD11', 'PRRC2B', 'UBAP2',
'ZNF318', 'CRTC3', 'USP54', 'ZC3H7A', 'LARP4B', 'RBM41', 'TCF12',
'PPIL3', 'PLEKHA5', 'TBKBP1', 'CIT', 'HSBP1', 'PCNT', 'CEP43',
'PRKAR2A', 'PRKACA', 'PRKAR2B', 'RDX', 'CENPF', 'TLE1', 'TLE3',
'TLE5', 'GOLGA3', 'GOLGA2', 'GOLGB1', 'GRIPAP1', 'CEP350',
'PDE4DIP', 'CEP135', 'CEP68', 'CNTRL', 'ERC1', 'GCC2', 'CLIP4',
'NIN', 'CEP112', 'MIPOL1', 'USP13', 'GCC1', 'JAKMIP1', 'CDK5RAP2',
'AKAP9', 'GORASP1', 'FYCO1', 'C1orf50', 'CEP250', 'TBK1', 'HOOK1',
'NINL', 'GLA', 'IMPDH2', 'SIRT5', 'NUTF2', 'ARF6', 'RNF41',
'SLC27A2', 'EIF4E2', 'POR', 'RAP1GDS1', 'WASHC4', 'FKBP15',
'GIGYF2', 'IDE', 'TIMM10', 'ALG11', 'NUP210', 'TIMM29', 'DNAJC11',
'TIMM10B', 'TIMM9', 'HDAC2', 'GPX1', 'TRMT1', 'ATP5MG', 'ATP6AP1',
'SIGMAR1', 'ATP13A3', 'AGPS', 'CYB5B', 'ACSL3', 'CYB5R3', 'RALA',
'COMT', 'RAB5C', 'RAB7A', 'RAB8A', 'RAB2A', 'RAB10', 'RAB14',
'RHOA', 'RAB1A', 'GNB1', 'GNG5', 'LMAN2', 'MOGS', 'TOR1AIP1',
'MTARC1', 'QSOX2', 'HS2ST1', 'NDUFAF2', 'SCCPDH', 'SCARB1',
'NAT14', 'DCAKD', 'FAM162A', 'DNAJC19', 'SELENOS', 'PTGES2',
'RAB18', 'MPHOSPH10', 'SRP72', 'ATE1', 'NSD2', 'SRP19', 'SRP54',
'MRPS25', 'DDX10', 'LARP7', 'MEPCE', 'NGDN', 'EXOSC8', 'NARS2',
'NOL10', 'CCDC86', 'SEPSECS', 'EXOSC5', 'EXOSC3', 'AATF', 'HECTD1',
'MRPS2', 'MRPS5', 'EXOSC2', 'MRPS27', 'GTF2F2', 'FBN1', 'FBN2',
'NUP214', 'NUP62', 'DCAF7', 'EIF4H', 'NUP54', 'MIB1', 'SPART',
'NEK9', 'ZNF503', 'NUP88', 'NUP58', 'MAT2B', 'FBLN5', 'PPT1',
'CUL2', 'MAP7D1', 'THTPA', 'ZYG11B', 'TIMM8B', 'RBX1', 'ELOC',
'ELOB', 'HMOX1', 'TRIM59', 'ARL6IP6', 'VPS39', 'CLCC1', 'VPS11',
'SUN2', 'ALG5', 'STOML2', 'NUP98', 'RAE1', 'MTCH1', 'HEATR3',
'MDN1', 'PLOD2', 'TOR1A', 'STC2', 'PLAT', 'ITGB1', 'CISD3',
'COL6A1', 'PVR', 'DNMT1', 'LOX', 'PCSK6', 'INHBE', 'NPC2', 'MFGE8',
'OS9', 'NPTX1', 'POGLUT2', 'POGLUT3', 'ERO1B', 'PLD3', 'FOXRED2',
'CHPF', 'PUSL1', 'EMC1', 'GGH', 'ERLEC1', 'IL17RA', 'NGLY1',
'HS6ST2', 'SDF2', 'NEU1', 'GDF15', 'TM2D3', 'ERP44', 'EDEM3',
'SIL1', 'POFUT1', 'SMOC1', 'PLEKHF2', 'FBXL12', 'UGGT2', 'CHPF2',
'ADAMTS1', 'HYOU1', 'FKBP7', 'ADAM9', 'FKBP10', 'SLC9A3R1',
'CHMP2A', 'CSDE1', 'TOMM70', 'MARK3', 'MARK2', 'DPH5', 'DCTPP1',
'MARK1', 'PTBP2', 'BAG5', 'UBXN8', 'GPAA1', 'WFS1', 'ABCC1',
'F2RL1', 'SCAP', 'DPY19L1', 'TMEM97', 'SLC30A6', 'TAPT1', 'ERMP1',
'NLRX1', 'RETREG3', 'PIGO', 'FAR2', 'ECSIT', 'ALG8', 'TMEM39B',
'GHITM', 'ACAD9', 'NDFIP2', 'BCS1L', 'NDUFAF1', 'TMED5', 'NDUFB9',
'PIGS']
sources = ["STRING"]
kwargs = {"STRING_escore": 0.2, # Keeps STRING interactions with an experimental score >= 0.2
}
ppi_graph = PPIGraph(protein_list=protein_list,
sources=sources,
**kwargs)
# Plot output network
plt.figure(figsize=(16, 16))
nx_graph = ppi_graph.nx_graph()
nx_pos = nx.kamada_kawai_layout(nx_graph)
nx_pos = nx.spring_layout(nx_graph, pos=nx_pos, k=0.5, iterations=3)
nx.draw(nx_graph, nx_pos, with_labels=True, node_color="lightblue", node_size=500)
|
examples/ppi/ppi_graph.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# For a molecule in **solution**, the free energy of restraint can be calculated analytically as $\Delta G_{rest} = -kT \ln \big( \frac{2\pi}{V_0 \beta k} \big)^{3/2}$.
#
# For our system, with a box volume of about $V_0$ = 570.6 nm$^3$, this comes to $\Delta G_{rest}$ = +15.41 $kT$.
#
# WE NOTE however that we are NOT performing restrained simulations of ligand in solvent!
# +
import numpy as np
### The analytical estimate for adding a harmonic restraint
k = 800.0 # harmonic force constant, in kJ/nm^2
kB = 1.381e-23 * 6.022e23 / 1000.0 # Boltzmann constant in kJ/mol/K
temperature = 300. # in K
beta = 1.0 / (kB * temperature) # inverse temperature of simulations (in 1/(kJ/mol))
print('beta', beta)
box_line = " 8.29427 8.29427 8.29427"
box_dims = [float(s) for s in (box_line.strip()).split() ]
print('box_dims', box_dims, 'nm')
V0 = box_dims[0]*box_dims[1]*box_dims[2] # box volume in nm^3
print('box volume V_0 =', V0, 'nm^3')
f_rest = -(3./2.)*np.log( 2.*np.pi/(V0*beta*k)) # in units kT
print('f_rest (analytical)=', f_rest, 'kT')
# -
|
data-release-2020-05-10/Harmonic aside.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *Accompanying code examples of the book "Introduction to Artificial Neural Networks and Deep Learning: A Practical Guide with Applications in Python" by [<NAME>](https://sebastianraschka.com). All code examples are released under the [MIT license](https://github.com/rasbt/deep-learning-book/blob/master/LICENSE). If you find this content useful, please consider supporting the work by buying a [copy of the book](https://leanpub.com/ann-and-deeplearning).*
#
# Other code examples and content are available on [GitHub](https://github.com/rasbt/deep-learning-book). The PDF and ebook versions of the book are available through [Leanpub](https://leanpub.com/ann-and-deeplearning).
# %load_ext watermark
# %watermark -a '<NAME>' -v -p torch
# - Runs on CPU or GPU (if available)
# # Model Zoo -- Convolutional ResNet and Residual Blocks
# Please note that this example does not implement a really deep ResNet as described in literature but rather illustrates how the residual blocks described in He et al. [1] can be implemented in PyTorch.
#
# - [1] <NAME>, et al. "Deep residual learning for image recognition." *Proceedings of the IEEE conference on computer vision and pattern recognition*. 2016.
# ## Imports
# +
import time
import numpy as np
import torch
import torch.nn.functional as F
from torch.utils.data import DataLoader
from torchvision import datasets
from torchvision import transforms
if torch.cuda.is_available():
torch.backends.cudnn.deterministic = True
# -
# ## Settings and Dataset
# +
##########################
### SETTINGS
##########################
# Device
device = torch.device("cuda:1" if torch.cuda.is_available() else "cpu")
# Hyperparameters
random_seed = 123
learning_rate = 0.01
num_epochs = 10
batch_size = 128
# Architecture
num_classes = 10
##########################
### MNIST DATASET
##########################
# Note transforms.ToTensor() scales input images
# to 0-1 range
train_dataset = datasets.MNIST(root='data',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = datasets.MNIST(root='data',
train=False,
transform=transforms.ToTensor())
train_loader = DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# Checking the dataset
for images, labels in train_loader:
print('Image batch dimensions:', images.shape)
print('Image label dimensions:', labels.shape)
break
# -
# ## ResNet with identity blocks
# The following code implements the residual blocks with skip connections such that the input passed via the shortcut matches the dimensions of the main path's output, which allows the network to learn identity functions. Such a residual block is illustrated below:
#
# 
# +
##########################
### MODEL
##########################
class ConvNet(torch.nn.Module):
def __init__(self, num_classes):
super(ConvNet, self).__init__()
#########################
### 1st residual block
#########################
# 28x28x1 => 28x28x4
self.conv_1 = torch.nn.Conv2d(in_channels=1,
out_channels=4,
kernel_size=(1, 1),
stride=(1, 1),
padding=0)
self.conv_1_bn = torch.nn.BatchNorm2d(4)
# 28x28x4 => 28x28x1
self.conv_2 = torch.nn.Conv2d(in_channels=4,
out_channels=1,
kernel_size=(3, 3),
stride=(1, 1),
padding=1)
self.conv_2_bn = torch.nn.BatchNorm2d(1)
#########################
### 2nd residual block
#########################
# 28x28x1 => 28x28x4
self.conv_3 = torch.nn.Conv2d(in_channels=1,
out_channels=4,
kernel_size=(1, 1),
stride=(1, 1),
padding=0)
self.conv_3_bn = torch.nn.BatchNorm2d(4)
# 28x28x4 => 28x28x1
self.conv_4 = torch.nn.Conv2d(in_channels=4,
out_channels=1,
kernel_size=(3, 3),
stride=(1, 1),
padding=1)
self.conv_4_bn = torch.nn.BatchNorm2d(1)
#########################
### Fully connected
#########################
self.linear_1 = torch.nn.Linear(28*28*1, num_classes)
def forward(self, x):
#########################
### 1st residual block
#########################
shortcut = x
out = self.conv_1(x)
out = self.conv_1_bn(out)
out = F.relu(out)
out = self.conv_2(out)
out = self.conv_2_bn(out)
out += shortcut
out = F.relu(out)
#########################
### 2nd residual block
#########################
shortcut = out
out = self.conv_3(out)
out = self.conv_3_bn(out)
out = F.relu(out)
out = self.conv_4(out)
out = self.conv_4_bn(out)
out += shortcut
out = F.relu(out)
#########################
### Fully connected
#########################
logits = self.linear_1(out.view(-1, 28*28*1))
probas = F.softmax(logits, dim=1)
return logits, probas
torch.manual_seed(random_seed)
model = ConvNet(num_classes=num_classes)
model = model.to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# -
# ### Training
# +
def compute_accuracy(model, data_loader):
correct_pred, num_examples = 0, 0
for i, (features, targets) in enumerate(data_loader):
features = features.to(device)
targets = targets.to(device)
logits, probas = model(features)
_, predicted_labels = torch.max(probas, 1)
num_examples += targets.size(0)
correct_pred += (predicted_labels == targets).sum()
return correct_pred.float()/num_examples * 100
start_time = time.time()
for epoch in range(num_epochs):
model = model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
features = features.to(device)
targets = targets.to(device)
### FORWARD AND BACK PROP
logits, probas = model(features)
cost = F.cross_entropy(logits, targets)
optimizer.zero_grad()
cost.backward()
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 50:
print ('Epoch: %03d/%03d | Batch %03d/%03d | Cost: %.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_loader), cost))
model = model.eval() # eval mode to prevent upd. batchnorm params during inference
with torch.set_grad_enabled(False): # save memory during inference
print('Epoch: %03d/%03d training accuracy: %.2f%%' % (
epoch+1, num_epochs,
compute_accuracy(model, train_loader)))
print('Time elapsed: %.2f min' % ((time.time() - start_time)/60))
print('Total Training Time: %.2f min' % ((time.time() - start_time)/60))
# -
# ### Evaluation
print('Test accuracy: %.2f%%' % (compute_accuracy(model, test_loader)))
# ## ResNet with convolutional blocks for resizing
# The following code implements the residual blocks with skip connections such that the input passed via the shortcut matches is resized to dimensions of the main path's output. Such a residual block is illustrated below:
#
# 
# +
##########################
### MODEL
##########################
class ConvNet(torch.nn.Module):
def __init__(self, num_classes):
super(ConvNet, self).__init__()
#########################
### 1st residual block
#########################
# 28x28x1 => 14x14x4
self.conv_1 = torch.nn.Conv2d(in_channels=1,
out_channels=4,
kernel_size=(3, 3),
stride=(2, 2),
padding=1)
self.conv_1_bn = torch.nn.BatchNorm2d(4)
# 14x14x4 => 14x14x8
self.conv_2 = torch.nn.Conv2d(in_channels=4,
out_channels=8,
kernel_size=(1, 1),
stride=(1, 1),
padding=0)
self.conv_2_bn = torch.nn.BatchNorm2d(8)
# 28x28x1 => 14x14x8
self.conv_shortcut_1 = torch.nn.Conv2d(in_channels=1,
out_channels=8,
kernel_size=(1, 1),
stride=(2, 2),
padding=0)
self.conv_shortcut_1_bn = torch.nn.BatchNorm2d(8)
#########################
### 2nd residual block
#########################
# 14x14x8 => 7x7x16
self.conv_3 = torch.nn.Conv2d(in_channels=8,
out_channels=16,
kernel_size=(3, 3),
stride=(2, 2),
padding=1)
self.conv_3_bn = torch.nn.BatchNorm2d(16)
# 7x7x16 => 7x7x32
self.conv_4 = torch.nn.Conv2d(in_channels=16,
out_channels=32,
kernel_size=(1, 1),
stride=(1, 1),
padding=0)
self.conv_4_bn = torch.nn.BatchNorm2d(32)
# 14x14x8 => 7x7x32
self.conv_shortcut_2 = torch.nn.Conv2d(in_channels=8,
out_channels=32,
kernel_size=(1, 1),
stride=(2, 2),
padding=0)
self.conv_shortcut_2_bn = torch.nn.BatchNorm2d(32)
#########################
### Fully connected
#########################
self.linear_1 = torch.nn.Linear(7*7*32, num_classes)
def forward(self, x):
#########################
### 1st residual block
#########################
shortcut = x
out = self.conv_1(x) # 28x28x1 => 14x14x4
out = self.conv_1_bn(out)
out = F.relu(out)
out = self.conv_2(out) # 14x14x4 => 714x14x8
out = self.conv_2_bn(out)
# match up dimensions using a linear function (no relu)
shortcut = self.conv_shortcut_1(shortcut)
shortcut = self.conv_shortcut_1_bn(shortcut)
out += shortcut
out = F.relu(out)
#########################
### 2nd residual block
#########################
shortcut = out
out = self.conv_3(out) # 14x14x8 => 7x7x16
out = self.conv_3_bn(out)
out = F.relu(out)
out = self.conv_4(out) # 7x7x16 => 7x7x32
out = self.conv_4_bn(out)
# match up dimensions using a linear function (no relu)
shortcut = self.conv_shortcut_2(shortcut)
shortcut = self.conv_shortcut_2_bn(shortcut)
out += shortcut
out = F.relu(out)
#########################
### Fully connected
#########################
logits = self.linear_1(out.view(-1, 7*7*32))
probas = F.softmax(logits, dim=1)
return logits, probas
torch.manual_seed(random_seed)
model = ConvNet(num_classes=num_classes)
model = model.to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# -
# ### Training
# +
def compute_accuracy(model, data_loader):
correct_pred, num_examples = 0, 0
for i, (features, targets) in enumerate(data_loader):
features = features.to(device)
targets = targets.to(device)
logits, probas = model(features)
_, predicted_labels = torch.max(probas, 1)
num_examples += targets.size(0)
correct_pred += (predicted_labels == targets).sum()
return correct_pred.float()/num_examples * 100
for epoch in range(num_epochs):
model = model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
features = features.to(device)
targets = targets.to(device)
### FORWARD AND BACK PROP
logits, probas = model(features)
cost = F.cross_entropy(logits, targets)
optimizer.zero_grad()
cost.backward()
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 50:
print ('Epoch: %03d/%03d | Batch %03d/%03d | Cost: %.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_loader), cost))
model = model.eval() # eval mode to prevent upd. batchnorm params during inference
with torch.set_grad_enabled(False): # save memory during inference
print('Epoch: %03d/%03d training accuracy: %.2f%%' % (
epoch+1, num_epochs,
compute_accuracy(model, train_loader)))
# -
# ### Evaluation
print('Test accuracy: %.2f%%' % (compute_accuracy(model, test_loader)))
# ## ResNet with convolutional blocks for resizing (using a helper class)
# This is the same network as above but uses a `ResidualBlock` helper class.
class ResidualBlock(torch.nn.Module):
def __init__(self, channels):
super(ResidualBlock, self).__init__()
self.conv_1 = torch.nn.Conv2d(in_channels=channels[0],
out_channels=channels[1],
kernel_size=(3, 3),
stride=(2, 2),
padding=1)
self.conv_1_bn = torch.nn.BatchNorm2d(channels[1])
self.conv_2 = torch.nn.Conv2d(in_channels=channels[1],
out_channels=channels[2],
kernel_size=(1, 1),
stride=(1, 1),
padding=0)
self.conv_2_bn = torch.nn.BatchNorm2d(channels[2])
self.conv_shortcut_1 = torch.nn.Conv2d(in_channels=channels[0],
out_channels=channels[2],
kernel_size=(1, 1),
stride=(2, 2),
padding=0)
self.conv_shortcut_1_bn = torch.nn.BatchNorm2d(channels[2])
def forward(self, x):
shortcut = x
out = self.conv_1(x)
out = self.conv_1_bn(out)
out = F.relu(out)
out = self.conv_2(out)
out = self.conv_2_bn(out)
# match up dimensions using a linear function (no relu)
shortcut = self.conv_shortcut_1(shortcut)
shortcut = self.conv_shortcut_1_bn(shortcut)
out += shortcut
out = F.relu(out)
return out
# +
##########################
### MODEL
##########################
class ConvNet(torch.nn.Module):
def __init__(self, num_classes):
super(ConvNet, self).__init__()
self.residual_block_1 = ResidualBlock(channels=[1, 4, 8])
self.residual_block_2 = ResidualBlock(channels=[8, 16, 32])
self.linear_1 = torch.nn.Linear(7*7*32, num_classes)
def forward(self, x):
out = self.residual_block_1.forward(x)
out = self.residual_block_2.forward(out)
logits = self.linear_1(out.view(-1, 7*7*32))
probas = F.softmax(logits, dim=1)
return logits, probas
torch.manual_seed(random_seed)
model = ConvNet(num_classes=num_classes)
model.to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# -
# ### Training
# +
def compute_accuracy(model, data_loader):
correct_pred, num_examples = 0, 0
for i, (features, targets) in enumerate(data_loader):
features = features.to(device)
targets = targets.to(device)
logits, probas = model(features)
_, predicted_labels = torch.max(probas, 1)
num_examples += targets.size(0)
correct_pred += (predicted_labels == targets).sum()
return correct_pred.float()/num_examples * 100
for epoch in range(num_epochs):
model = model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
features = features.to(device)
targets = targets.to(device)
### FORWARD AND BACK PROP
logits, probas = model(features)
cost = F.cross_entropy(logits, targets)
optimizer.zero_grad()
cost.backward()
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 50:
print ('Epoch: %03d/%03d | Batch %03d/%03d | Cost: %.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_dataset)//batch_size, cost))
model = model.eval() # eval mode to prevent upd. batchnorm params during inference
with torch.set_grad_enabled(False): # save memory during inference
print('Epoch: %03d/%03d training accuracy: %.2f%%' % (
epoch+1, num_epochs,
compute_accuracy(model, train_loader)))
# -
# ### Evaluation
print('Test accuracy: %.2f%%' % (compute_accuracy(model, test_loader)))
# %watermark -iv
|
code/model_zoo/pytorch_ipynb/resnet-ex-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LeetCode
#
# ## Link
#
# 1. https://leetcode.com/
#
# 2. https://leetcode-cn.com/
#
# ## Demo
#
# 1. https://www.onlinegdb.com/
#
# ## Details
#
# LeetCode: 16, 17, 19
# # LeetCode 16. 3Sum Closest 整数反转
# Given an integer array nums of length n and an integer target, find three integers in nums such that the sum is closest to target.
#
# Return the sum of the three integers.
#
# You may assume that each input would have exactly one solution.
#
# 给你一个长度为 n 的整数数组 nums 和 一个目标值 target。请你从 nums 中选出三个整数,使它们的和与 target 最接近。
#
# 返回这三个数的和。
#
# 假定每组输入只存在恰好一个解。
#
#
# Example 1:
#
# ```
# Input: nums = [-1,2,1,-4], target = 1
# Output: 2
# Explanation: The sum that is closest to the target is 2. (-1 + 2 + 1 = 2).
# ```
#
# Example 2:
#
# ```
# Input: nums = [0,0,0], target = 1
# Output: 0
# ```
#
# Constraints:
#
# - 3 <= nums.length <= 1000
#
# - -1000 <= nums[i] <= 1000
#
# - -10^4 <= target <= 10^4
#
# ## 解题思路
#
# 这一题的解法是用两个指针夹逼的方法。先对数组进行排序,i 从头开始往后面扫。这里同样需要注意数组中存在多个重复数字的问题。具体处理方法很多,可以用 map 计数去重。这里笔者简单的处理,i 在循环的时候和前一个数进行比较,如果相等,i 继续往后移,直到移到下一个和前一个数字不同的位置。j,k 两个指针开始一前一后夹逼。j 为 i 的下一个数字,k 为数组最后一个数字,由于经过排序,所以 k 的数字最大。j 往后移动,k 往前移动,逐渐夹逼出最接近 target 的值。
#
# 这道题还可以用暴力解法,三层循环找到距离 target 最近的组合。
from typing import List
class Solution:
def threeSumClosest(self, nums: List[int], target: int) -> int:
n = len(nums)
nums.sort()
re_min = 0 #存储当前最小的差值
for i in range(n):
low = i+1
high = n-1
while low < high:
three_sum = nums[i] + nums[low] + nums[high]
x = target - three_sum #当前三数的差值
if re_min == 0:
re_min = abs(x)
sum_min = three_sum #sum_min为当前最接近的和
if abs(x) < re_min:
re_min = abs(x)
sum_min = three_sum
if three_sum == target:
return target
elif three_sum < target:
low += 1
else:
high -= 1
return sum_min
# # LeetCode 17. Letter Combinations of a Phone Number 电话号码的字母组合
# Given a string containing digits from 2-9 inclusive, return all possible letter combinations that the number could represent. Return the answer in any order.
#
# A mapping of digit to letters (just like on the telephone buttons) is given below. Note that 1 does not map to any letters.
#
# 给定一个仅包含数字 2-9 的字符串,返回所有它能表示的字母组合。答案可以按 任意顺序 返回。
#
# 给出数字到字母的映射如下(与电话按键相同)。注意 1 不对应任何字母。
#
# 
#
# Example 1:
#
# ```
# Input: digits = "23"
# Output: ["ad","ae","af","bd","be","bf","cd","ce","cf"]
# ```
#
# Example 2:
#
# ```
# Input: digits = ""
# Output: []
# ```
#
# Example 3:
#
# ```
# Input: digits = "2"
# Output: ["a","b","c"]
# ```
#
#
# Constraints:
#
# - 0 <= digits.length <= 4
#
# - digits[i] is a digit in the range ['2', '9'].(digits[i] 是范围 ['2', '9'] 的一个数字。)
#
#
#
# ## 解题思路
#
# DFS 递归深搜
#
# ## Reference
#
# 1. https://www.bilibili.com/video/BV1cy4y167mM/
#
# ```
# class Solution(object):
# def letterCombinations(self, digits):
# """
# 动态规划
# dp[i]: 前i个字母的所有组合
# 由于dp[i]只与dp[i-1]有关,可以使用变量代替列表存储降低空间复杂度
# :type digits: str
# :rtype: List[str]
# """
# if not digits:
# return []
# d = {'2': 'abc', '3': 'def', '4': 'ghi', '5': 'jkl',
# '6': 'mno', '7': 'pqrs', '8': 'tuv', '9': 'wxyz'}
# n = len(digits)
# dp = [[] for _ in range(n)]
# dp[0] = [x for x in d[digits[0]]]
# for i in range(1, n):
# dp[i] = [x + y for x in dp[i - 1] for y in d[digits[i]]]
# return dp[-1]
#
# def letterCombinations2(self, digits):
# """
# 使用变量代替上面的列表
# 降低空间复杂度
# :type digits: str
# :rtype: List[str]
# """
# if not digits:
# return []
# d = {'2': 'abc', '3': 'def', '4': 'ghi', '5': 'jkl',
# '6': 'mno', '7': 'pqrs', '8': 'tuv', '9': 'wxyz'}
# n = len(digits)
# res = ['']
# for i in range(n):
# res = [x + y for x in res for y in d[digits[i]]]
# return res
#
# def letterCombinations3(self, digits):
# """
# 递归
# :param digits:
# :return:
# """
# d = {'2': 'abc', '3': 'def', '4': 'ghi', '5': 'jkl',
# '6': 'mno', '7': 'pqrs', '8': 'tuv', '9': 'wxyz'}
# if not digits:
# return []
# if len(digits) == 1:
# return [x for x in d[digits[0]]]
# return [x + y for x in d[digits[0]] for y in self.letterCombinations3(digits[1:])]
# ```
class Solution(object):
def letterCombinations(self, digits):
if not digits:
return []
d = {'2': 'abc', '3': 'def', '4': 'ghi', '5': 'jkl',
'6': 'mno', '7': 'pqrs', '8': 'tuv', '9': 'wxyz'}
n = len(digits)
dp = [[] for _ in range(n)]
dp[0] = [x for x in d[digits[0]]]
for i in range(1, n):
dp[i] = [x + y for x in dp[i - 1] for y in d[digits[i]]]
return dp[-1]
# # LeetCode 19. Remove Nth Node From End of List 删除链表的倒数第 N 个结点
# Given the head of a linked list, remove the $n^{th}$ node from the end of the list and return its head.
#
# 给你一个链表,删除链表的倒数第 n 个结点,并且返回链表的头结点。
#
# 
#
# Example 1:
#
# ```
# Input: head = [1,2,3,4,5], n = 2
# Output: [1,2,3,5]
# ```
#
# Example 2:
#
# ```
# Input: head = [1], n = 1
# Output: []
# ```
#
# Example 3:
#
# ```
# Input: head = [1,2], n = 1
# Output: [1]
# ```
#
# Constraints:
#
# - The number of nodes in the list is sz.(链表中结点的数目为 sz)
#
# - 1 <= sz <= 30
#
# - 0 <= Node.val <= 100
#
# - 1 <= n <= sz
#
# Follow up: Could you do this in one pass?(你能尝试使用一趟扫描实现吗?)
#
# ## 解题思路
#
# - 先循环一次拿到链表的总长度,然后循环到要删除的结点的前一个结点开始删除操作。需要注意的一个特例是,有可能要删除头结点,要单独处理。
#
# - 这道题有一种特别简单的解法。设置 2 个指针,一个指针距离前一个指针 n 个距离。同时移动 2 个指针,2 个指针都移动相同的距离。当一个指针移动到了终点,那么前一个指针就是倒数第 n 个节点了。
#
# ## Reference
#
# https://stackoverflow.com/questions/61610160/remove-nth-node-from-end-of-listleetcode-python
"""
class Solution:
def removeNthFromEnd(self, head: ListNode, n: int) -> ListNode:
head_dummy = ListNode()
head_dummy.next = head
slow, fast = head_dummy, head_dummy
while(n!=0): # fast先往前走n步
fast = fast.next
n -= 1
while(fast.next!=None):
slow = slow.next
fast = fast.next
# fast 走到结尾后,slow 的下一个节点为倒数第N个节点
slow.next = slow.next.next # 删除
return head_dummy.next
"""
class Solution:
def removeNthFromEnd(self, head, n):
fast = slow = head
for _ in range(n):
fast = fast.next
if not fast:
return head.next
while fast.next:
fast = fast.next
slow = slow.next
slow.next = slow.next.next
return head
|
AATCC/lab-report/w3/practice-leetcode-labs-w3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install pyapacheatlas
from Authenticate_to_Purview_AML import *
ws,guid,client = authentitae_to_purview_AML()
from Create_ML_Lineage_Types import *
create_ml_lineage_types(client)
# %run Create_ML_Lineage_Functions
create_workspace_entities(ws)
create_datastore_entities(ws)
create_dataset_entities(ws)
create_experiment_entities(ws)
# +
## uncomment below code to link PowerBI Dataset and Report in lineage if you have access to a PBI workspace
# #The PowerBI entities will populate with more details if you set up a scan for PBI workspaces in Purview
# #We are just creating a placeholders and links for lineage below
# #get batch inference data entity name and exprimentname
# batchpred_data_ent_name = 'batchinfer_data.csv_CreditRiskPipeline'
# experimentname = "CreditRiskPipeline"
# #create PowerBI dataset entity and lineage
# pbi_workspace = '<YOUR PBIWORKSPACE URL>' #'https://xxx.powerbi.com/groups/7c555287-f9b8-45ff-be6c-9909afe9df40'
# pbi_datasetid = '<YOUR PBI Dataset ID>' #'c4a30c22-466d-4a30-a1ac-8736ed6567cc'
# pbidata_ent_name = 'creditriskpbidataset'
# create_powerbi_dataset_and_lineage(experimentname,pbi_workspace,pbi_datasetid,pbidata_ent_name,batchpred_data_ent_name,'custom_ml_dataset')
# #create PowerBI report entity and lineage
# pbi_reportid = '<YOUR PBI Report ID>' #'e495453d-6c0c-4fb9-bdc4-556319f6a57b'
# pbi_ent_name = 'creditriskpbireport'
# create_powerbi_report_and_lineage(experimentname,pbi_workspace,pbi_reportid,pbi_ent_name,pbi_datasetid)
|
AMLNotebooks/02_Create_CreditRisk_AML_Pipeline_Lineage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Step 1: Import required libraries and packages
# +
# import required libraries
import pandas as pd
import numpy as np
import datetime
from geopy.distance import distance
import time
pd.set_option('mode.chained_assignment', None)
# set directories
resource_directory = "00_Ntbk_Resources\\00_DataCollectionProcessingCleaning\\PMPML_BusRoutes_July2019\\"
result_directory = "00_Ntbk_Resources\\01_DataAnalysis\\00_ProcessedData\\PMPML_BusRoutes_July2019\\"
# -
# ## Step 2: Data Collection
# source: http://opendata.punecorporation.org/Citizen/CitizenDatasets/Index
#
# tags: PMPML
#
# title: PMPML Bus Routes - July 2019
#
# file type: .zip
#
# size: 8.81 MB
# ## Step 3: Data Processing and Cleaning
# import trip/service calendar information into the dataframe
df_TripCalendar = pd.read_csv(f"{resource_directory}calendar.txt")
df_TripCalendar
# import bus stop information into the dataframe
df_BusStops = pd.read_csv(f"{resource_directory}stops.txt")
df_BusStops = df_BusStops.drop(columns=["zone_id","stop_url","stop_desc","stop_code","location_type","parent_station","stop_timezone","wheelchair_boarding"])
df_BusStops
# check dataframe for NaN values
df_BusStops[df_BusStops.isna().any(axis=1)]
# import bus route information into the dataframe
df_BusRouteShapes = pd.read_csv(f"{resource_directory}shapes.txt")
df_BusRouteShapes
# +
# calculate the distance for each route and add it to separate column of dataframe
df_temp = df_BusRouteShapes.groupby("shape_id").transform(lambda x : x.shift(1))[["shape_pt_lat", "shape_pt_lon"]]
list_dist_traveled = []
for a,b,c,d in zip(df_BusRouteShapes["shape_pt_lat"], df_BusRouteShapes["shape_pt_lon"], df_temp["shape_pt_lat"], df_temp["shape_pt_lon"]):
if not np.isnan(c) :
list_dist_traveled.append(distance([a,b],[c,d]).km)
else :
list_dist_traveled.append(0)
df_BusRouteShapes ["shape_dist_traveled"] = list_dist_traveled
df_BusRouteShapes["trip_distance"] = df_BusRouteShapes.groupby("shape_id")["shape_dist_traveled"].transform(sum)
df_BusRouteShapes
# -
# import bus trip information into the dataframe and add trip distance information in a new column of dataframe
df_BusTrips = pd.read_csv(f"{resource_directory}trips.txt")
df_BusTrips = df_BusTrips.drop(columns=["trip_short_name","block_id","wheelchair_accessible","bikes_allowed","duty","duty_sequence_number","run_sequence_number"])
df_BusTrips = df_BusTrips.join(df_BusRouteShapes.set_index("shape_id").groupby("shape_id")[["trip_distance"]].max().reset_index("shape_id").set_index("shape_id"), on="shape_id")
df_BusTrips
# import bus stop times information into the dataframe
df_BusStopTimes = pd.read_csv(f"{resource_directory}stop_times.txt")
df_BusStopTimes = df_BusStopTimes.drop(columns=["stop_headsign","pickup_type","drop_off_type","shape_dist_traveled","timepoint"])
df_BusStopTimes
# information from this dataframe needs to verify because this dataframe has a lot of information and chances of human error are also high
df_BusStopTimes[df_BusStopTimes.isnull().any(axis=1)]
# To verify the information in dataframe we are going to run some filters and check whether wrong information is present or not.\
# We are also going to fix the wrong information with the average value of logically correct information, while doing this we also need to take care that already correct information is not getting changed.\
# 1) Check whether arrival_time or departure time are correctly entered i.e. they are in 24hr format.\
# 2) Check duration between arrival and departure time for each stop, it should be realistic\
# 3) Check duration between stop to stop arrival time, it should not be more than 2.5 to 3hrs considering traffic conditions
# check whether arrival_time or departure time is correctly entered i.e. they are in 24hr format.
df_temp1 = df_BusStopTimes[(df_BusStopTimes["arrival_time"] > '23:59:59') | (df_BusStopTimes["departure_time"] > '23:59:59')]
df_temp1["trip_id"].unique()
# +
# there are some entries which have arrival or departure time greater than 23:59:59
# the wrong information is found where bus trips are started late at night and end on next day early in the morning
# so time mentioned beyond 23:59:59 is nothing but early morning on the next day
# let's check and replace such entries with correct time values
index_bad1 = df_BusStopTimes[(df_BusStopTimes["trip_id"].str[-6:-2] >= str(2040)) & (df_BusStopTimes["arrival_time"].str.startswith("24:") | df_BusStopTimes["arrival_time"].str.startswith("25:"))].index
index_bad2 = df_BusStopTimes[(df_BusStopTimes["trip_id"].str[-6:-2] >= str(2040)) & (df_BusStopTimes["departure_time"].str.startswith("24:") | df_BusStopTimes["departure_time"].str.startswith("25:"))].index
df_BusStopTimes.loc[index_bad1,"arrival_time"] = df_BusStopTimes.iloc[index_bad1]["arrival_time"].str.replace('^24:','00:').str.replace('^25:','01:').to_list()
df_BusStopTimes.loc[index_bad2,"departure_time"] = df_BusStopTimes.iloc[index_bad2]["departure_time"].str.replace('^24:','00:').str.replace('^25:','01:').to_list()
# +
# for most of the trips from Swargate to Pune Station arrival and departure times are wrong which results in longer trip duration
# so lets filter out those bad trip ids and replace their arrival and departure times with average values of good trip_ids
df_temp1 = df_temp1[df_temp1["trip_id"].str.contains("NORMAL_5_Swargate To Pune Station_Up")]
list_BadTripIds = df_temp1["trip_id"].unique().tolist()
df_temp2 = df_BusStopTimes[df_BusStopTimes["trip_id"].str.contains("NORMAL_5_Swargate To Pune Station_Up")]
df_temp2.reset_index(inplace=True, drop =True)
df_temp3 = df_temp2[~ df_temp2["trip_id"].isin(list_BadTripIds)]
df_temp3.reset_index(inplace=True, drop =True)
df_temp3["deparr_time_diff"] = pd.to_timedelta(df_temp3["departure_time"]) - pd.to_timedelta(df_temp3["arrival_time"])
df_temp3["arr_time_diff"] = df_temp3.groupby("trip_id")["arrival_time"].transform(lambda x : pd.to_timedelta(x) - pd.to_timedelta(x.shift()))
df_temp3["avg_deparr_time_diff"] = df_temp3.groupby("stop_sequence")["deparr_time_diff"].transform(lambda x : pd.to_timedelta(x).sum()/ len(x))
df_temp3["avg_arr_time_diff"] = df_temp3.groupby("stop_sequence")["arr_time_diff"].transform(lambda x : pd.to_timedelta(x).sum()/ len(x))
df_temp3["avg_arr_time_diff"] = df_temp3.groupby("trip_id")["avg_arr_time_diff"].transform(lambda x : x.cumsum())
list_avg_deparr_time_diff = df_temp3[df_temp3["trip_id"] == "NORMAL_5_Swargate To Pune Station_Up-1010_0"]["avg_deparr_time_diff"].astype(str).str[7:15].tolist()
list_avg_arr_time_diff = df_temp3[df_temp3["trip_id"] == "NORMAL_5_Swargate To Pune Station_Up-1010_0"]["avg_arr_time_diff"].astype(str).str[7:15].tolist()
# -
# this function will replace the wrong arrival and departure times of trips considering average duration found in other trips with correct trip ids
def changeTimings(df) :
df_temp1 = pd.DataFrame()
df_temp1["avg_arr_time_diff"] = list_avg_arr_time_diff
list_arrival_time = (pd.to_timedelta(df["arrival_time"].tolist()[1]) + pd.to_timedelta(df_temp1["avg_arr_time_diff"])).astype(str).str[7:15].tolist()
df["arrival_time"][2:] = list_arrival_time[2:]
df_temp1["avg_deparr_time_diff"] = list_avg_deparr_time_diff
df["departure_time"] = ((pd.to_timedelta(df.reset_index(drop=True)["arrival_time"]) + pd.to_timedelta(df_temp1["avg_deparr_time_diff"])).astype(str).str[7:15].tolist())
return df
df_BusStopTimes[df_BusStopTimes["trip_id"].isin(list_BadTripIds)] = df_BusStopTimes[df_BusStopTimes["trip_id"].isin(list_BadTripIds)].groupby("trip_id").apply(lambda x: changeTimings(x))
df_BusStopTimes
# +
# still, some entries need to be corrected individually
df_temp1 = df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_181C_Na Ta Wadi To Kondhwa Bk_Up-1450_0"]
df_temp1["deparr_time_diff"] = pd.to_timedelta(df_temp1["departure_time"]) - pd.to_timedelta(df_temp1["arrival_time"])
df_temp1["arr_time_diff"] = pd.to_timedelta(df_temp1["arrival_time"]) - pd.to_timedelta(df_temp1["arrival_time"].shift())
df_temp1.iloc[0,6] = pd.to_timedelta("00:00:00")
df_temp1.iloc[1,6] = pd.to_timedelta("00:01:15")
df_temp1["arr_time_diff"] = df_temp1.groupby("trip_id")["arr_time_diff"].transform(lambda x: x.cumsum())
df_temp1["arrival_time"] = (pd.to_timedelta(df_temp1.iloc[0,1]) + pd.to_timedelta(df_temp1["arr_time_diff"])).astype(str).str[7:15].tolist()
df_temp1["departure_time"] = (pd.to_timedelta(df_temp1["arrival_time"]) + pd.to_timedelta(df_temp1["deparr_time_diff"])).astype(str).str[7:15].tolist()
df_temp1= df_temp1.drop(columns=["deparr_time_diff","arr_time_diff"])
df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_181C_Na Ta Wadi To Kondhwa Bk_Up-1450_0"] = df_temp1
print(df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_181C_Na Ta Wadi To Kondhwa Bk_Up-1450_0"])
df_temp1 = df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_181C_Na Ta Wadi To Kondhwa Bk_Up-1450_0"]
df_temp1["deparr_time_diff"] = pd.to_timedelta(df_temp1["departure_time"]) - pd.to_timedelta(df_temp1["arrival_time"])
df_temp1["arr_time_diff"] = pd.to_timedelta(df_temp1["arrival_time"]) - pd.to_timedelta(df_temp1["arrival_time"].shift())
df_temp1.iloc[0,6] = pd.to_timedelta("00:00:00")
df_temp1.iloc[1,6] = pd.to_timedelta("00:01:15")
df_temp1["arr_time_diff"] = df_temp1.groupby("trip_id")["arr_time_diff"].transform(lambda x: x.cumsum())
df_temp1["arrival_time"] = (pd.to_timedelta(df_temp1.iloc[0,1]) + pd.to_timedelta(df_temp1["arr_time_diff"])).astype(str).str[7:15].tolist()
df_temp1["departure_time"] = (pd.to_timedelta(df_temp1["arrival_time"]) + pd.to_timedelta(df_temp1["deparr_time_diff"])).astype(str).str[7:15].tolist()
df_temp1= df_temp1.drop(columns=["deparr_time_diff","arr_time_diff"])
df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_181C_Na Ta Wadi To Kondhwa Bk_Up-1450_0"] = df_temp1
print(df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_181C_Na Ta Wadi To Kondhwa Bk_Up-1450_0"])
df_temp1 = df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_256_Someshwarwadi To PMC_Down-1945_0"]
df_temp1["deparr_time_diff"] = pd.to_timedelta(df_temp1["departure_time"]) - pd.to_timedelta(df_temp1["arrival_time"])
df_temp1["arr_time_diff"] = pd.to_timedelta(df_temp1["arrival_time"]) - pd.to_timedelta(df_temp1["arrival_time"].shift())
df_temp1.iloc[0,6] = pd.to_timedelta("00:00:00")
df_temp1.iloc[1,6] = pd.to_timedelta("00:01:15")
df_temp1.iloc[2,6] = pd.to_timedelta("00:01:15")
df_temp1.iloc[1,5] = pd.to_timedelta("00:00:18")
df_temp1["arr_time_diff"] = df_temp1.groupby("trip_id")["arr_time_diff"].transform(lambda x: x.cumsum())
df_temp1["arrival_time"] = (pd.to_timedelta(df_temp1.iloc[0,1]) + pd.to_timedelta(df_temp1["arr_time_diff"])).astype(str).str[7:15].tolist()
df_temp1["departure_time"] = (pd.to_timedelta(df_temp1["arrival_time"]) + pd.to_timedelta(df_temp1["deparr_time_diff"])).astype(str).str[7:15].tolist()
df_temp1= df_temp1.drop(columns=["deparr_time_diff","arr_time_diff"])
df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_256_Someshwarwadi To PMC_Down-1945_0"] = df_temp1
print(df_BusStopTimes[df_BusStopTimes["trip_id"]=="NORMAL_256_Someshwarwadi To PMC_Down-1945_0"])
# -
# verify once again before proceeding to next filter
df_temp1 = df_BusStopTimes[(df_BusStopTimes["arrival_time"] > '23:59:59') | (df_BusStopTimes["departure_time"] > '23:59:59')]
df_temp1
# as arrival and departure values are within range we can change the data type of both columns to datetime
df_BusStopTimes["arrival_time"] = df_BusStopTimes["arrival_time"].astype('datetime64[ns]')
df_BusStopTimes["departure_time"] = df_BusStopTimes["departure_time"].astype('datetime64[ns]')
df_BusStopTimes
# this function will update the date of entry to next day if arrival time is less than its previous arrival time
def changeDate(df):
changeIndex = df[df["arrival_time"].shift(-1) < df["arrival_time"]].index + 1
if changeIndex.any() :
df.loc[changeIndex[0] : df.tail(1).index[0],"arrival_time"] = df.loc[changeIndex[0] : df.tail(1).index[0],"arrival_time"] + datetime.timedelta(days=1)
changeIndex = df[df["departure_time"].shift(-1) < df["departure_time"]].index + 1
if changeIndex.any() :
df.loc[changeIndex[0] : df.tail(1).index[0],"departure_time"] = df.loc[changeIndex[0] : df.tail(1).index[0],"departure_time"] + datetime.timedelta(days=1)
return df
df_BusStopTimes = df_BusStopTimes.groupby("trip_id").apply(lambda x: changeDate(x))
df_BusStopTimes
# let's add a column to the dataframe which will show arrival time difference between two consecutive stops
# also, add column showing duration between arrival and departure time for each stop
df_BusStopTimes["stp2stp_arrival_time"] = df_BusStopTimes.groupby("trip_id")["arrival_time"].transform(lambda x : x.shift(-1) - x)
df_BusStopTimes["deparr_time_diff"] = df_BusStopTimes["departure_time"] - df_BusStopTimes["arrival_time"]
df_BusStopTimes
# take a look at the maximum and 75% values of the stop to stop arrival duration as well as departure and arrival time difference
df_BusStopTimes.describe()
# +
# there are entries where the time difference between arrival and departure time is greater than 6hrs, i.e. not possible
# though 75% value is 30 seconds, for the safer side we'll assume duration as 3minutes and replace those with 75% value
# a similar technique can be applied for a stop to stop arrival time as well
df_BusStopTimes.loc[df_BusStopTimes[df_BusStopTimes["deparr_time_diff"] > datetime.timedelta(minutes = 3)].index,"deparr_time_diff"] = df_BusStopTimes.describe().loc["75%", "deparr_time_diff"]
df_BusStopTimes.loc[df_BusStopTimes[df_BusStopTimes["stp2stp_arrival_time"] > datetime.timedelta(minutes = 30)].index,"stp2stp_arrival_time"]= df_BusStopTimes.describe().loc["75%", "stp2stp_arrival_time"]
df_BusStopTimes
# -
# now we have to update arrival and departure times according to updated/modified durations
df_BusStopTimes["stp2stp_arrival_time"] = df_BusStopTimes.groupby("trip_id")["stp2stp_arrival_time"].transform(lambda x : x.cumsum().shift().fillna(datetime.timedelta(minutes = 0)))
df_BusStopTimes
# this function will update the arrival time with the help of first entry and stop to stop arrival time duration value
def updateArrivalTime(df):
df.loc[:,"arrival_time"] = df.iloc[0,1] + df["stp2stp_arrival_time"]
return df
df_BusStopTimes = df_BusStopTimes.groupby("trip_id").apply(lambda x: updateArrivalTime(x))
df_BusStopTimes
# similarly, update departure time as well
df_BusStopTimes.loc[:,"departure_time"] = df_BusStopTimes["arrival_time"] + df_BusStopTimes["deparr_time_diff"]
df_BusStopTimes
df_BusStopTimes["trip_bgn_time"] = df_BusStopTimes.groupby("trip_id")["arrival_time"].transform("first")
df_BusStopTimes["trip_end_time"] = df_BusStopTimes.groupby("trip_id")["arrival_time"].transform("last")
df_BusStopTimes["trip_duration"] = df_BusStopTimes["trip_end_time"] - df_BusStopTimes["trip_bgn_time"]
df_BusStopTimes.sort_values("trip_duration", ascending = True)
# let's drop this trip_id as it has only one stop_sequence
df_BusStopTimes = df_BusStopTimes.drop(df_BusStopTimes[df_BusStopTimes["trip_id"] == "NORMAL_107_Gharkul Vasahat Warje To Pimplegurav_Down-0530_0"].index[0])
df_BusStopTimes.reset_index(drop=True, inplace=True)
df_BusStopTimes
# let's verify the dataframe once again (check max and 75% values)
df_BusStopTimes["stp2stp_arrival_time"] = df_BusStopTimes.groupby("trip_id")["arrival_time"].transform(lambda x : x.shift(-1) - x)
df_BusStopTimes["deparr_time_diff"] = df_BusStopTimes["departure_time"] - df_BusStopTimes["arrival_time"]
df_BusStopTimes.describe()
# drop the duration columns as they are not required anymore
df_BusStopTimes.drop(["stp2stp_arrival_time", "deparr_time_diff", "trip_bgn_time", "trip_end_time", "trip_duration"], axis =1, inplace =True)
df_BusStopTimes
# now that we've repaired the dataframes we should save it to resources folder for further analysis
df_TripCalendar.to_csv(f"{result_directory}calendar.txt", index=False)
df_BusStops.to_csv(f"{result_directory}stops.txt", index=False)
df_BusRouteShapes.to_csv(f"{result_directory}shapes.txt", index=False)
df_BusTrips.to_csv(f"{result_directory}trips.txt", index=False)
df_BusStopTimes.to_csv(f"{result_directory}stop_times.txt", index=False)
|
00_Ntbk_PuneBusRoutes_DataCollection,Processing,Cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from carel import Carel
from carelgrid import CarelGrid
from gamecanvas import GameCanvas
class SmallCarl(Carel):
def __init__(self, canvas, grid):
super().__init__(canvas, grid)
self.beepers_count = 0
def drop_beeper(self):
if self.beepers_count != 0:
super().drop_beeper()
self.beepers_count -= 1
else:
print('Do not have beeper')
def collect_beeper(self):
super().collect_beeper()
self.beepers_count += 1
# +
field = [[0 for x in range(6)] for y in range(4)]
field[0][1] = 2
field[1][0] = 2
field[3][2] = 1
grid = CarelGrid(field)
canvas = GameCanvas(speed=1)
carel = SmallCarl(canvas, grid)
# -
carel.drop_beeper()
carel.move()
if carel.is_beeper():
carel.collect_beeper()
carel.move()
carel.drop_beeper()
carel.move()
carel.drop_beeper()
|
home_task_4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## VENRON-Electricity Dataset preprocessing for Fonduer
#
# This script is used to pre-process the spreadsheets in order to apply the cell annotations from the prediction json files or the corresponding manually labeled annotation range sheet.
import os
import pandas as pd
import json
# +
# First create xlsx output folder
# Run in terminal
# # ! libreoffice --headless --calc --convert-to xlsx --outdir src/data/gold/xlsx src/data/gold/spreadsheet/*
# -
# ## Cell annotations based on predictions
#
# We apply the json predictions to the gold data set of 114 spreadsheets (real annotations, not gold/manual labeled)
# +
from openpyxl import load_workbook
from openpyxl.cell.cell import Cell
from openpyxl.styles import Border, Side, PatternFill, Font, GradientFill, Alignment
from openpyxl.reader.drawings import find_images
xlsx_path = f"{os.getcwd()}/data/xlsx"
json_path = f"{os.getcwd()}/data/gold_pred_annotated/json_predictions"
pred_out_path = f"{os.getcwd()}/data/gold_pred_annotated/spreadsheet"
files_json = os.listdir(json_path)
pred_styles = {
"data": Font(color="000001"),
"derived": Font(color="000001"),
"header": Font(color="000002"),
"attributes": Font(color="000002"),
"metadata": Font(color="000003"),
"notes": Font(color="000004"),
}
def excel_wb(f):
wb = load_workbook(filename = f"{xlsx_path}/{f}")
images = find_images(f"{xlsx_path}/{f}")
return (wb, images)
(wb, images) = excel_wb("11_MeadMktplace.xlsx")
(wb, images)
# -
wb["MeadMktplace"]._images
# +
from openpyxl import load_workbook
from openpyxl.cell.cell import Cell
from openpyxl.styles import Border, Side, PatternFill, Font, GradientFill, Alignment
xlsx_path = f"{os.getcwd()}/data/gold/xlsx"
json_path = f"{os.getcwd()}/data/gold_pred_annotated/json_predictions"
pred_out_path = f"{os.getcwd()}/data/gold_pred_annotated/spreadsheet"
files_json = os.listdir(json_path)
pred_styles = {
"data": Font(color="000001"),
"derived": Font(color="000001"),
"header": Font(color="000002"),
"attributes": Font(color="000002"),
"metadata": Font(color="000003"),
"notes": Font(color="000004"),
}
def excel_wb(f):
wb = load_workbook(filename = f"{xlsx_path}/{f}")
return wb
# Assuming maximally ZZ column in excel
def int_to_char(i):
if (i <= 25):
return chr(65+j)
else:
return chr(90)+chr(65+ (i-25))
pred_styles = {
"data": Font(color="000001"),
"derived": Font(color="000001"),
"header": Font(color="000002"),
"attributes": Font(color="000002"),
"metadata": Font(color="000003"),
"notes": Font(color="000004"),
}
for json_file in files_json:
fname = json_file[0:-5]
f = open(f'./data/gold_pred_annotated/json_predictions/{json_file}', 'r')
data = f.read()
parsed = json.loads(data)
try:
wb = excel_wb(f"{fname}.xlsx")
# For each worksheet in the spreadsheet
for idx, (sheet_name, sheet_values) in enumerate(parsed.items()):
# load annotations
text = sheet_values['text']
labels = sheet_values['labels']
labels_probs = sheet_values['labels_probs']
# load sheet
ws = wb[sheet_name]
# Override the cell-style tags
for i, row in enumerate(text):
for j, cell in enumerate(row):
ws_index = f"{int_to_char(j)}{i+1}"
if (cell != "" and ws[ws_index].font.color is not None):
if (labels_probs[i][j] > 0.6):
ws[ws_index].font = pred_styles[labels[i][j]]
else:
ws[ws_index].font = None
# save the spreadsheet with annotated worksheets
wb.save(f"{pred_out_path}/{fname}.xlsx")
except:
# just copy the file if errors occur (openpyxl min value issue)
wb.save(f"{pred_out_path}/{fname}.xlsx")
print(f"FAILED to read {fname}.xlsx")
# -
# We also apply the json predictions to the full data set of 687 spreadsheets
# +
# First create xlsx output folder
# Run in terminal for each batch (batches are needed to avoid libreoffice failure)
# # ! libreoffice --headless --calc --convert-to xlsx --outdir src/data/full/xlsx src/data/full/spreadsheet/batch_X*
# +
from openpyxl import load_workbook
from openpyxl.cell.cell import Cell
from openpyxl.styles import Border, Side, PatternFill, Font, GradientFill, Alignment
xlsx_path = f"{os.getcwd()}/data/full/xlsx"
json_path = f"{os.getcwd()}/data/full_pred_annotated/json_predictions"
pred_out_path = f"{os.getcwd()}/data/full_pred_annotated/spreadsheet"
files_json = os.listdir(json_path)
pred_styles = {
"data": Font(color="000001"),
"derived": Font(color="000001"),
"header": Font(color="000002"),
"attributes": Font(color="000002"),
"metadata": Font(color="000003"),
"notes": Font(color="000004"),
}
def excel_wb(f):
wb = load_workbook(filename = f"{xlsx_path}/{f}")
return wb
# Assuming maximally ZZ column in excel
def int_to_char(i):
if (i <= 25):
return chr(65+j)
else:
return chr(90)+chr(65+ (i-25))
pred_styles = {
"data": Font(color="000001"),
"derived": Font(color="000001"),
"header": Font(color="000002"),
"attributes": Font(color="000002"),
"metadata": Font(color="000003"),
"notes": Font(color="000004"),
}
for json_file in files_json:
fname = json_file[0:-5]
f = open(f'./data/full_pred_annotated/json_predictions/{json_file}', 'r')
data = f.read()
parsed = json.loads(data)
try:
wb = excel_wb(f"{fname}.xlsx")
# For each worksheet in the spreadsheet
for idx, (sheet_name, sheet_values) in enumerate(parsed.items()):
# load annotations
text = sheet_values['text']
labels = sheet_values['labels']
labels_probs = sheet_values['labels_probs']
# load sheet
ws = wb[sheet_name]
# Override the cell-style tags
for i, row in enumerate(text):
for j, cell in enumerate(row):
ws_index = f"{int_to_char(j)}{i+1}"
if (cell != "" and ws[ws_index].font.color is not None):
if (labels_probs[i][j] > 0.6):
ws[ws_index].font = pred_styles[labels[i][j]]
else:
ws[ws_index].font = None
# save the spreadsheet with annotated worksheets
wb.save(f"{pred_out_path}/{fname}.xlsx")
except:
# just copy the file if errors occur (openpyxl min value issue)
wb.save(f"{pred_out_path}/{fname}.xlsx")
print(f"FAILED to read {fname}.xlsx")
# -
# ## HTML modifications
#
# Fonduer makes it difficult to deal with image names and document names similar to spans.
# In order to avoid rewriting all featurizers we simple construct the HTML files with new spans for the image and document name.
# +
import bs4
def extend_html_file(html_path, file_name):
# load the file
with open(f"{html_path}/{file_name}") as fin:
txt = fin.read()
soup = bs4.BeautifulSoup(txt)
# Insert document name
new_tag = soup.new_tag("div")
new_tag.string = f"Document name: {file_name}"
soup.body.insert(0, new_tag)
# Insert image urls
for i in soup.find_all("img"):
image_url = i["src"]
if (i.parent.name == "body"):
t = soup.new_tag("div")
t.string = image_url
i.parent.insert(-1, t)
else:
t = soup.new_tag("span")
t.string = image_url
i.parent.insert(0, t)
# save the file again
with open(fin.name, "w") as outf:
outf.write(str(soup))
# +
import os
from html.parser import HTMLParser
paths = ["gold", "full", "gold_pred_annotated", "full_pred_annotated"]
for p in paths:
html_path = f"{os.getcwd()}/data/{p}/html"
files_html = [x for x in os.listdir(html_path) if x[-4:] == "html"]
for file_html in files_html:
extend_html_file(html_path, file_html)
# -
# ### HTML Cell annotations
#
# Another approach to cell annotations is directly in the HTML.
# This is better for spreadsheets with a lot of formatting (colors, images, charts etc.), as openpyxl does modify the spreadsheet and formatting. (e.g. from $ 135.12 -> 135.124123123123 removes the rounding)
#
# Due to some pre-processign on both files the HTML (conversion via Libreoffice e.g. ignores collapsed cols/rows) and json-predictions (via cell annotation algorithm, e.g. has errors for multi-sheets) the annotations are far from perfect. However, we only take the ones we are certain and report incorrect/correct entries.
# In the future this should be done as one native process which converts an XLS file to HTML, makes predictiosn and annotates the files. However, e.g. openpyxl has also conversion issues at the moment, thus this is open for future work.
# +
import os
import bs4
import json
import math
import sys
html_path = f"{os.getcwd()}/data/gold/html"
json_path = f"{os.getcwd()}/data/gold_pred_annotated/json_predictions"
pred_out_path = f"{os.getcwd()}/data/gold_pred_annotated/html"
files_json = os.listdir(json_path)
def open_html(fname):
with open(f"{html_path}/{fname}.html") as fin:
txt = fin.read()
soup = bs4.BeautifulSoup(txt)
return soup
def save_html(fname, soup):
with open(f"{pred_out_path}/{fname}.html", "w") as outf:
outf.write(str(soup))
def get_sheet_name_from_table_soup(t):
# Previous sibling "\n", then sheet name
return list(t.previous_siblings)[1].em.string
def is_number(s):
try:
float(s)
return True
except ValueError:
return False
# Check if the cell content approx. matches (e.g. rounded data value)
def match_cells(cell, table_cell):
return (
cell == table_cell.string or
(
"sdval" in table_cell.attrs
and is_number(cell) and is_number(table_cell["sdval"])
and math.isclose(float(cell), float(table_cell["sdval"]), abs_tol=0.001)
)
)
# returns (row_diff, cell_diff) for json_predictions to HTML table
# Not guarantueed to be perfect, but for shifted tables (e.g. because of colapsed columns/rows) it works
def find_diffs(text, table_rows):
text_cords = (0,0)
for i, row in enumerate(text):
for j, cell in enumerate(row):
if (cell != ""):
text_cords = (i,j)
cell = text[text_cords[0]][text_cords[1]]
for i, table_row in enumerate(table_rows):
row_cells = table_row.find_all("td")
for j, table_cell in enumerate(row_cells):
if match_cells(cell, table_cell):
return (text_cords[0]-i, text_cords[1]-j)
return (0,0)
for json_file in files_json:
fname = json_file[0:-5]
f = open(f'./data/gold_pred_annotated/json_predictions/{json_file}', 'r')
data = f.read()
parsed = json.loads(data)
correct = 0
incorrect = 0
skipped_rows = 0
skipped_cells = 0
try:
html_soup = open_html(fname)
tables = html_soup.find_all("table")
# Has multiple sheets
if (len(tables) > 1):
table_dict = { get_sheet_name_from_table_soup(t):t for t in tables }
# For each worksheet in the spreadsheet
for idx, (sheet_name, sheet_values) in enumerate(parsed.items()):
# load annotations
text = sheet_values['text']
labels = sheet_values['labels']
labels_probs = sheet_values['labels_probs']
# load sheet
table = table_dict[sheet_name] if len(tables) > 1 else tables[0]
table_rows = table.find_all("tr")
(row_diff, col_diff) = find_diffs(text, table_rows)
# Override the cell-style tags
for i, row in enumerate(text):
i_t = i-row_diff
if (i_t >= len(table_rows) or i_t < 0):
skipped_rows += 1
continue
table_row = table_rows[i_t]
for j, cell in enumerate(row):
row_cells = table_row.find_all("td")
# Predictions for multi-sheet spreadsheets are partly corrupted. (Out of bounds)
j_t = j-col_diff
if (j_t >= len(row_cells) or j_t < 0):
skipped_cells += 1
continue
table_cell = row_cells[j_t]
if (cell != "" and labels_probs[i][j] > 0.6):
# print("Cell", cell, "vs. table", table_cell.string)
# Predictions for multi-sheet spreadsheets are partly corrupted.
# Thus explicit cell comparison for any predictions.
if match_cells(cell, table_cell):
table_cell["cellType"] = labels[i][j]
correct += 1
else:
incorrect += 1
# save the spreadsheet with annotated worksheets
print(f"{fname} with (row_diff={row_diff}, col_diff={col_diff}), " +
f"\ncorrect={correct}, \nincorrect={incorrect}, " +
f"\nskipped_row={skipped_rows}, \nskipped_cells={skipped_cells}\n")
save_html(fname, html_soup)
except KeyError as e:
# just copy the file if errors occur
save_html(fname, html_soup)
print(f"FAILED to read {fname}.html", sys.exc_info()[0])
print(e)
# -
# ## Playground
#
# Experimenting
|
src/preprocess.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # part1: tensorflow
#
# +
import tensorflow as tf
print('Tensorflow Version:{}'.format(tf.__version__))
print(tf.test.is_gpu_available())
# -
# # part2: opencv
import cv2 as cv
img1 = cv.imread("pic1.jpg")
img2 = cv.imread("pic2.jpg")
W = 3.752
H = 4.382
L = 8.342
W = round(W, 2) # round函数将W保留两位小数
H = round(H, 2)
L = round(L, 2)
text = "W:"+str(W)+" " + "H:"+str(H) + " " + "L:"+str(L)
cv.putText(img1, text, (40, 50), cv.FONT_HERSHEY_PLAIN, 2.0, (0, 0, 255), 2)
cv.imshow("image show",img1)
cv.waitKey(0)
cv.imwrite('lena.png',img1)
cv.destroyAllWindows()
# # part3: pytorch
# +
import torch
print(torch.cuda.is_available()) # 查看系统GPU是否可以使用,经常用来判断是否装好gpu版的pytorch
# -
print(torch.cuda.current_device())# 返回当前设备序号
print(torch.cuda.get_device_name(0))# 返回第0号设备的name
print(torch.cuda.device_count())# 返回可使用的gpu的数量
print(torch.cuda.memory_allocated(device="cuda:0"))#返回0号设备的当前GPU显存使用量(以字节为单位)
#判断是否安装了cuda
import torch
print(torch.cuda.is_available()) #返回True则说明已经安装了cuda
#判断是否安装了cuDNN
from torch.backends import cudnn
print(cudnn.is_available()) #返回True则说明已经安装了cuDNN
# # part4: CPU与GPU的计算能力对比
# +
# 屏蔽tensorflow输出的log信息
# 注意:代码在import tensorflow之前
import os
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "2"
import sys
print("python的版本信息:",sys.version)
#python的版本信息: 3.7.9 (default, Aug 31 2020, 17:10:11) [MSC v.1916 64 bit (AMD64)]
import tensorflow as tf
'''
验证GPU相对于CPU,在并行计算优势明显
'''
n=100000000 #1亿次 (2亿次 会发生内存分配OOM )
# -
# 创建在CPU环境上运算的 2 个矩阵
with tf.device('/cpu:0'):
cpu_a = tf.random.normal([1, n])
cpu_b = tf.random.normal([n, 1])
print(cpu_a.device, cpu_b.device)
cpu_b.device
cpu_a.device
# 创建使用 GPU环境运算的 2 个矩阵
with tf.device('/gpu:0'):
gpu_a = tf.random.normal([1, n])
gpu_b = tf.random.normal([n, 1])
print(gpu_a.device, gpu_b.device)
import timeit
def cpu_run(): # CPU 运算函数
with tf.device('/cpu:0'):
c = tf.matmul(cpu_a, cpu_b)
return c
def gpu_run():# GPU 运算函数
with tf.device('/gpu:0'):
c = tf.matmul(gpu_a, gpu_b)
return c
# 第一次计算需要热身,避免将初始化时间结算在内
cpu_time = timeit.timeit(cpu_run, number=10)
gpu_time = timeit.timeit(gpu_run, number=10)
print('首先计算10次(含热身环境)的平均时间,CPU计算消耗时间:%.3fms,GPU计算消耗时间:%.3fms!'%(cpu_time*1000, gpu_time*1000) )
#正式计算10次,取平均时间
cpu1_time = timeit.timeit(cpu_run, number=10)
gpu1_time = timeit.timeit(gpu_run, number=10)
print('正式计算10次的平均时间,CPU计算消耗时间:%.3fms,GPU计算消耗时间:%.3fms!'%(cpu1_time*1000, gpu1_time*1000))
print('正式计算10次的平均时间,CPU计算消耗时间:%.3fms,GPU计算消耗时间:%.3fms!'%(cpu1_time*1000, gpu1_time*1000))
# # Part5 : 获取CPU型号
# psutil是一个跨平台库(http://pythonhosted.org/psutil/)能够轻松实现获取系统运行的进程和系统利用率(包括CPU、内存、磁盘、网络等)信息。它主要用来做系统监控,性能分析,进程管理。它实现了同等命令行工具提供的功能,如ps、top、lsof、netstat、ifconfig、who、df、kill、free、nice、ionice、iostat、iotop、uptime、pidof、tty、taskset、pmap等。目前支持32位和64位的Linux、Windows、OS X、FreeBSD和Sun Solaris等操作系统.
# +
import psutil
def get_cpu_info():
cpu1 = psutil.cpu_count()
print("cpu逻辑个数:",cpu1)
def get_mem_info():
mem = psutil.virtual_memory()
mem1 = str(mem.total/1024/1024/1024)
mem2 = str(mem.free/1024/1024/1024)
print("内存总数为:",mem1[0:3],"G")
print("空闲内存总数:", mem2[0:3], "G")
get_cpu_info()
get_mem_info()
# -
psutil.cpu_times()
psutil.cpu_times().user
psutil.cpu_times().idle
psutil.cpu_count()
psutil.cpu_count(logical=False)
psutil.cpu_percent(11)
mem = psutil.virtual_memory()
mem
print(mem.used/1024/1024/1024)
psutil.disk_partitions()
psutil.disk_usage('/')
psutil.disk_io_counters()
psutil.disk_io_counters(perdisk=True)
psutil.net_io_counters()
psutil.net_io_counters(pernic=True)
psutil.boot_time()
import datetime
datetime.datetime.fromtimestamp(psutil.boot_time ()).strftime("%Y-%m-%d %H: %M: %S")
psutil.pids()
p = psutil.Process(22096)
p.name() #进程名
p.exe() #进程的bin路径
p.cwd() #进程的工作目录绝对路径
p.status() #进程状态
p.create_time() #进程创建时间
p.cpu_times() #进程的cpu时间信息,包括user,system两个cpu信息
p.cpu_affinity() #get进程cpu亲和度,如果要设置cpu亲和度,将cpu号作为参考就好
p.memory_percent() #进程内存利用率
p.memory_info() #进程内存rss,vms信息
p.io_counters() #进程的IO信息,包括读写IO数字及参数
p.num_threads() #进程开启的线程数
听过psutil的Popen方法启动应用程序,可以跟踪程序的相关信息
from subprocess import PIPE
p = psutil.Popen(["/usr/bin/python", "-c", "print('hello')"],stdout=PIPE)
p.name()
p.username()
import time
now_time = time.strftime('%Y-%m-%d-%H:%M:%S', time.localtime(time.time()))
print(now_time)
for pnum in psutil.pids():
p = psutil.Process(pnum)
print(u"进程名 %-20s 内存利用率 %-18s 进程状态 %-10s 创建时间 %-10s " \
% (p.name(), p.memory_percent(), p.status(), p.create_time()))
io = psutil.disk_partitions()
print("系统磁盘信息:" + str(io))
for i in io:
o = psutil.disk_usage(i.device)
print("总容量:" + str(int(o.total / (1024.0 * 1024.0 * 1024.0))) + "G")
print("已用容量:" + str(int(o.used / (1024.0 * 1024.0 * 1024.0))) + "G")
print("可用容量:" + str(int(o.free / (1024.0 * 1024.0 * 1024.0))) + "G")
net = psutil.net_io_counters()
bytes_sent = '{0:.2f} Mb'.format(net.bytes_recv / 1024 / 1024)
bytes_rcvd = '{0:.2f} Mb'.format(net.bytes_sent / 1024 / 1024)
print(u"网卡接收流量 %s 网卡发送流量 %s" % (bytes_rcvd, bytes_sent))
users_count = len(psutil.users())
users_list = ",".join([u.name for u in psutil.users()])
print(u"当前有%s个用户,分别是 %s" % (users_count, users_list))
free = str(round(psutil.virtual_memory().free / (1024.0 * 1024.0 * 1024.0), 2))
total = str(round(psutil.virtual_memory().total / (1024.0 * 1024.0 * 1024.0), 2))
memory = int(psutil.virtual_memory().total - psutil.virtual_memory().free) / float(psutil.virtual_memory().total)
print(u"物理内存: %s G" % total)
print(u"剩余物理内存: %s G" % free)
print(u"物理内存使用率: %s %%" % int(memory * 100))
# 系统启动时间
print(u"系统启动时间: %s" % datetime.datetime.fromtimestamp(psutil.boot_time()).strftime("%Y-%m-%d %H:%M:%S"))
# !pip install wmi
# !pip install pywin32
import wmi
c = wmi.WMI()
# # 硬盘序列号
for physical_disk in c.Win32_DiskDrive():
print(physical_disk.SerialNumber)
# CPU序列号
for cpu in c.Win32_Processor():
print(cpu.ProcessorId.strip())
# 主板序列号
for board_id in c.Win32_BaseBoard():
print(board_id.SerialNumber)
# mac地址
for mac in c.Win32_NetworkAdapter():
print(mac.MACAddress)
# bios序列号
for bios_id in c.Win32_BIOS():
print(bios_id.SerialNumber.strip())
# # PART6:获取NVIDIA GPU型号信息
#简单使用
from pynvml import *
nvmlInit() #初始化 ( 需要将 nvidia-smi.exe文件拷贝到C:\Program Files\NVIDIA Corporation\NVSMI, 否则 “NVML Shared Library Not Found“)
info1=nvmlSystemGetDriverVersion()
info1
print("Driver: ",info1) #显示驱动信息
def checkGPU():
import pynvml
pynvml.nvmlInit() #初始化
print("Driver: ",pynvml.nvmlSystemGetDriverVersion())#显示驱动信息
deviceCount = pynvml.nvmlDeviceGetCount()
for i in range(deviceCount):
handle = pynvml.nvmlDeviceGetHandleByIndex(i)
print("GPU", i, ":", pynvml.nvmlDeviceGetName(handle))
checkGPU()
# +
import pynvml #导包
NUM_EXPAND = 1024 * 1024
"""GPU消耗情况"""
pynvml.nvmlInit() #初始化
#print("Driver: ", pynvml.nvmlSystemGetDriverVersion()) #显示驱动信息
all_gpu_used = []#用于记录所有GPU内存使用信息的list
need_record_pid = 19289 #需要记录的进程PID
gpuDeviceCount = pynvml.nvmlDeviceGetCount()#获取Nvidia GPU块数
gpuDeviceCount
# -
for i in range(gpuDeviceCount):
handle = pynvml.nvmlDeviceGetHandleByIndex(i)#获取GPU i的handle,后续通过handle来处理
#info = pynvml.nvmlDeviceGetMemoryInfo(handle)#通过handle获取GPU i的信息
## gpu_memory_total = info.total #GPU i的总显存
#gpu_memory_used = info.used / NUM_EXPAND #转为MB单位
#all_gpu_used.append(gpu_memory_used) #添加进list
###还可以直接针对pid的gpu消耗进行统计
info_list = pynvml.nvmlDeviceGetComputeRunningProcesses(handle)#获取所有GPU上正在运行的进程信息
info_list_len = len(info_list)
gpu_memory_used = 0
if info_list_len > 0:#0表示没有正在运行的进程
for info_i in info_list:
if info_i.pid == need_record_pid:#如果与需要记录的pid一致
gpu_memory_used += info_i.usedGpuMemory / NUM_EXPAND #统计某pid使用的总显存
all_gpu_used.append(gpu_memory_used)
pynvml.nvmlShutdown() #最后关闭管理工具
|
gpu_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import seaborn as sns
if os.getcwd().endswith('notebook'):
os.chdir('..')
from rna_learn.load import load_dataset
from rna_learn.transform import (
sequence_embedding,
normalize, denormalize,
make_dataset_balanced,
one_hot_encode_classes,
split_train_test_set,
)
# -
sns.set(palette='colorblind', font_scale=1.3)
input_path = os.path.join(os.getcwd(), 'data/ncbi/dataset.csv')
alphabet = ['A', 'T', 'G', 'C']
classes = ['psychrophilic', 'mesophilic', 'thermophilic']
dataset_df = load_dataset(input_path, alphabet)
dataset_df.head()
# +
y_str, balanced_dataset_df = make_dataset_balanced(
dataset_df,
cat_name='temperature_range',
classes=classes,
)
y = one_hot_encode_classes(y_str, classes)
sequences = balanced_dataset_df['sequence'].values
x = sequence_embedding(sequences, alphabet)
x_train, y_train, x_test, y_test, train_idx, test_idx = split_train_test_set(
x, y, test_ratio=0.2, return_indices=True)
# -
output_train_path = os.path.join(os.getcwd(), 'data/dataset_train.csv')
output_test_path = os.path.join(os.getcwd(), 'data/dataset_test.csv')
unused_indices_path = os.path.join(os.getcwd(), 'data/dataset_unused_indices.csv')
# +
# balanced_dataset_df.iloc[train_idx].reset_index(drop=True).to_csv(output_train_path, index=False)
# balanced_dataset_df.iloc[test_idx].reset_index(drop=True).to_csv(output_test_path, index=False)
# -
idx_set = set(train_idx) | set(test_idx)
unused_indices = np.array([
idx for idx in range(len(dataset_df)) if idx not in idx_set
])[:,np.newaxis]
unused_indices.shape
# +
# pd.DataFrame(unused_indices, columns=['index']).to_csv(unused_indices_path, index=False)
# -
l = [len(sequence) for sequence in dataset_df['sequence'].values]
print('min', np.min(l))
print('max', np.max(l))
print('mean', np.mean(l))
print('std', np.std(l))
|
notebook/archive/Prepare training dataset.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PowerShell
# name: powershell
# ---
# + [markdown] azdata_cell_guid="a49ad7c3-e68e-4829-bcec-0bfc9115e476"
# # SQL Assessment API Quick Start
# ## Assess your SQL Server configuration for best practices in 2 simple steps
# + [markdown] azdata_cell_guid="2060c61e-6488-4b54-8aa8-8604a6159c0c"
# ### 1. Setup
# You need to install PowerShell SqlServer module using the following command. It is a good practice to run Import-Module at the beginning of your session as well. Get-Module will show you the version you have installed. The minimum version you want is 21.1.18206 — it is the version of SqlServer module containing SQL Assessment API GA.
# + azdata_cell_guid="d16282b0-a7fe-4f94-aa23-88b773dd0329"
# Uncomment and run Install-Module only the first time
# Install-Module -Name SqlServer -AllowClobber -Force
Import-Module -Name SqlServer
Get-Module
# + [markdown] azdata_cell_guid="69d60fd2-727d-4b30-b723-b217fa8ea128"
# ### 2. Invoke an assessment
# This command runs an assessment against your local SQL Server instance.
#
# + azdata_cell_guid="7468c134-77ff-4786-a795-a3767b55bd3b"
Get-SqlInstance -ServerInstance 'localhost' | Invoke-SqlAssessment
# + [markdown] azdata_cell_guid="546c7f04-c14f-449d-ba34-53bc52e545a5"
#
# You will see in the results that each rule has some properties (not the full list):
# - Severity (info, warning, critical)
# - Message property explains the recommendation but if you need more info, there is a HelpLink property that points at documentation on the subject.
# - Origin shows which ruleset and version the recommendation is coming from
#
# Visit SQL Assessment API GitHub page at http://aka.ms/sql-assessment-api for a full list of rules and properties.
#
# If you want to get recommendations for all databases on the local instance, you can run this command.
# + azdata_cell_guid="bbe2ef33-c5db-4a64-bb32-cd93b906c48d"
Get-SqlDatabase -ServerInstance 'localhost' | Invoke-SqlAssessment
# + [markdown] azdata_cell_guid="180b3a6d-1dab-4641-acd4-dc319e81ef1f"
# ### Learn more about SQL Assessment API
# To learn more about SQL Assesment API such as customizing and extending the rule set, saving the results in a table, etc., please visit:
# - Continue learning more about SQL Assessments with the ➡ [SQL Assessment API Tutorial notebook](https://github.com/microsoft/sql-server-samples/blob/master/samples/manage/sql-assessment-api/notebooks/SQLAssessmentAPITutorialNotebook.ipynb)
# - Docs online page: https://docs.microsoft.com/sql/sql-assessment-api/sql-assessment-api-overview
# - GitHub repo: http://aka.ms/sql-assessment-api
|
samples/manage/sql-assessment-api/notebooks/SQLAssessmentAPIQuickStartNotebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from thesis_initialise import *
# +
def testfunc(a, b,
c,
d = 4, # another comment
/,
e: int = 5,
# stuff
f = 6,
g = 7,
h = 8,
*args,
# morestuff
i,
j = 10,
# k
k0 = 11,
k1 = 110,
k2 = 1100,
l = 12,
m = 13,
# bonusstuff
n = 14,
# morebonusstuff
o = 15,
# _ignore
fee = 'fee', fie = 'fie', foe = 'foe',
fum = 'fum',
# subignore
boo = 'boo',
p = 16,
**kwargs,
):
print(a, b, c, d, e, f, g, h, args, i, j, k0, k1, k2, l, m, n, o, p, kwargs)
class A:
def testfunc(a, b,
c,
d = 4, # another comment
/,
e: int = 5,
# stuff
f = 6,
g = 7,
h = 8,
*args,
# morestuff
i,
j = 10,
# k
k0 = 11,
k1 = 110,
k2 = 1100,
l = 12,
m = 13,
# bonusstuff
n = 14,
# morebonusstuff
o = 15,
# _ignore
fee = 'fee', fie = 'fie', foe = 'foe',
fum = 'fum',
# subignore
boo = 'boo',
p = 16,
**kwargs,
):
print(a, b, c, d, e, f, g, h, args, i, j, k0, k1, k2, l, m, n, o, p, kwargs)
testfunc(1, 2, 3, i = 9)
# -
from everest.cascade.cascade import Cascade
from everest.cascade.signature import get_paramlevels, get_hierarchy, get_cascade, Signature
mylevs = get_paramlevels(testfunc)
mylevs
mylevs = get_paramlevels(A.testfunc)
mylevs
myhier = get_hierarchy(testfunc)
myhier
mycasc = get_cascade(testfunc)
mycasc
mysig = Signature(testfunc)
mysig
myskip = Signature(testfunc, skip = 2, skipkeys = {'e'})
myskip
mybnd = myskip.bind(1, i = 'bananas', newkwarg = 'mynewkwarg')
mybnd
mybnd.hashID
mybnd2 = myskip.bind(1, 4, i = 'bananas', newkwarg = 'mynewkwarg')
mybnd2
mybnd2.hashID
|
dev/dev_019_hierarchy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
def get_domains(examples):
d = [set() for i in examples[0]]
for x in examples:
for i, xi in enumerate(x):
d[i].add(xi)
return [list(sorted(x)) for x in d]
def more_general(h1, h2):
more_general_parts = []
for x, y in zip(h1, h2):
mg = x == "?" or (x != "0" and (x == y or y == "0"))
more_general_parts.append(mg)
return all(more_general_parts)
def fulfills(example, hypothesis):
# the implementation is the same as for hypotheses:
return more_general(hypothesis, example)
def min_generalizations(h, x):
h_new = list(h)
for i in range(len(h)):
if not fulfills(x[i:i+1], h[i:i+1]):
h_new[i] = '?' if h[i] != '0' else x[i]
return [tuple(h_new)]
def min_specializations(h, domains, x):
results = []
for i in range(len(h)):
if h[i] == "?":
for val in domains[i]:
if x[i] != val:
h_new = h[:i] + (val,) + h[i+1:]
results.append(h_new)
elif h[i] != "0":
h_new = h[:i] + ('0',) + h[i+1:]
results.append(h_new)
return results
def generalize_S(x, G, S):
S_prev = list(S)
for s in S_prev:
if s not in S:
continue
if not fulfills(x, s):
S.remove(s)
Splus = min_generalizations(s, x)
## keep only generalizations that have a counterpart in G
S.update([h for h in Splus if any([more_general(g,h)
for g in G])])
## remove hypotheses less specific than any other in S
S.difference_update([h for h in S if
any([more_general(h, h1)
for h1 in S if h != h1])])
return S
def specialize_G(x, domains, G, S):
G_prev = list(G)
for g in G_prev:
if g not in G:
continue
if fulfills(x, g):
G.remove(g)
Gminus = min_specializations(g, domains, x)
## keep only specializations that have a conuterpart in S
G.update([h for h in Gminus if any([more_general(h, s)
for s in S])])
## remove hypotheses less general than any other in G
G.difference_update([h for h in G if
any([more_general(g1, h)
for g1 in G if h != g1])])
return G
def candidate_elimination(examples):
domains = get_domains(examples)[:-1]
n = len(domains)
G = set([("?",)*n])
S = set([("0",)*n])
print("Maximally specific hypotheses - S ")
print("Maximally general hypotheses - G ")
i=0
print("\nS[0]:",str(S),"\nG[0]:",str(G))
for xcx in examples:
i=i+1
x, cx = xcx[:-1], xcx[-1] # Splitting data into attributes and decisions
if cx=='Y': # x is positive example
G = {g for g in G if fulfills(x, g)}
S = generalize_S(x, G, S)
else: # x is negative example
S = {s for s in S if not fulfills(x, s)}
G = specialize_G(x, domains, G, S)
print("\nS[{0}]:".format(i),S)
print("G[{0}]:".format(i),G)
return
with open('data22_sports.csv') as csvFile:
examples = [tuple(line) for line in csv.reader(csvFile)]
candidate_elimination(examples)
# -
|
Program 2 - Candidate Elim/.ipynb_checkpoints/pgm2_long-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import cloudpickle as pickle
from logging import getLogger
from walker_benchmark import walker_benchmark
from ipystate.impl.walker import Walker
from ipystate.impl.cython_walker import CythonWalker
from ipystate.impl.walker_master import WalkerMaster
# +
rows = {}
class PickleWalker:
def walk(self, ns):
return pickle.dumps(ns)
rows['PickleDumps'] = walker_benchmark(walker=PickleWalker())
# -
rows['CythonWalker'] = walker_benchmark(walker=CythonWalker(getLogger()))
rows['Walker'] = walker_benchmark(walker=Walker(getLogger()))
rows['WalkerMaster'] = walker_benchmark(walker=WalkerMaster(getLogger()))
# CythonWalker -- оптимизированный walker + компилляция Сython-ом
# Walker -- оптимизированный walker
# WalkerMaster -- текущая реализация walker из master
# PikleDumps -- вызов функции pickle.dumps()
df = pd.DataFrame.from_dict(rows)
df
|
src/main/python/ipystate/benchmarks/walker_benchmark.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Remote Query Examples
# This notebook demonstrates remote querying capabilities using `SPARQLWrapper`
# SPARQLWrapper is a Python wrapper around a SPARQL service that gives the capability to
# remotely execute queries on SPARQL endpoints. It simply invokes a SPARQLWrapper class
# that requires an endpoint which you can pass a query to. The following are a few
# examples that demonstate the capabilities:
# +
import ipywidgets as W
import pandas as pd
import traitlets as T
from rdflib import BNode, Graph, Literal, URIRef, namespace
from SPARQLWrapper import JSON, XML, SPARQLWrapper
from ipyradiant import (
CytoscapeVisualizer,
FileManager,
PathLoader,
QueryWidget,
service_patch_rdflib,
)
# -
try:
import requests
resp = requests.get("http://dbpedia.org/sparql/")
assert resp.ok
DBPEDIA_AVAILABLE = True
except AssertionError:
print("DBPedia not available (404).")
DBPEDIA_AVAILABLE = False
# ### Query a single remote endpoint using `SPARQLWrapper`
# This example creates a sparql class from the graph location: http://dbpedia.org/sparql.
# In the setQuery method, you can pass a string that describes the query you want. Here,
# we perform a SELECT query that returns the labels for the resource Asturias from
# dbpedia:
# +
sparql = SPARQLWrapper("http://dbpedia.org/sparql/")
sparql.setQuery(
"""
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
SELECT ?label
WHERE { <http://dbpedia.org/resource/Asturias> rdfs:label ?label }
"""
)
if DBPEDIA_AVAILABLE:
sparql.setReturnFormat(JSON)
results = sparql.query().convert()
results_df = pd.json_normalize(results["results"]["bindings"])
results_df[["label.value"]]
# -
# > Note: `SPARQLWrapper.setQuery` supports various query methods such as SELECT, ASK,
# > CONSTRUCT, and DESCRIBE
# ### SERVICE patch for rdflib
# Before we look at remote query examples with `SERVICE` calls, let's discuss an issue
# with rdflib:
# Currently, rdflib contains a bug where the SERVICE clause is not supported properly.
# ipyradiant detects when SERVICE is used for federated queries and converts the keyword
# into lower case for rdflib support. A warning is issued when SERVICE is detected. This
# patch is turned off for release>5.0.0
# Here is a working example of the query string conversion to a scheme that is supported
# by rdflib:
# +
query_str = """
SELECT DISTINCT ?s ?p ?o
WHERE
{
SERVICE <http://dbpedia.org/sparql>
{
SELECT ?s ?p ?o
WHERE {?s ?p ?o}
}
}
"""
if DBPEDIA_AVAILABLE:
query_str = service_patch_rdflib(query_str)
print(query_str)
# -
# ### Query Widget Example
# Here we show a working example of Remote Querying using the Query Widget available under
# ipyradiant. First we define a WidgetExample class and fill in the query form with query
# that contains a SERVICE call:
class WidgetExample(W.Tab):
query = T.Instance(QueryWidget)
log = W.Output()
def __init__(self, graph: Graph = None, *args, **kwargs):
super().__init__(*args, **kwargs)
self.children = [self.query]
@T.default("query")
def make_default_query_widget(self):
return QueryWidget()
# The following code snippet creates a Widget UI where you can see the query being passed
# in. Take a look at the UI panel on the right where you can see the query using the
# SPARQL endpoint from linkeddata. If you click on `Run Query`, you can immediate see the
# query string being modified to the correct form:
widget = WidgetExample()
widget.query.query_constructor.query_type = "SELECT DISTINCT"
widget.query.query_constructor.query_line = "*"
widget.query.query_constructor.query_body = """
{
SERVICE <http://dbpedia.org/sparql>
{
SELECT ?s ?p ?o
WHERE {?s ?p ?o}
LIMIT 10
}
}
"""
widget.query.query_constructor.formatted_query.value = """
SELECT DISTINCT *
WHERE {
SERVICE <http://dbpedia.org/sparql>
{
SELECT ?s ?p ?o
WHERE {?s ?p ?o}
LIMIT 10
}
}
"""
widget.query
# "click" the button
if DBPEDIA_AVAILABLE:
widget.query.run_button.click()
# ### Nested Query Example
# A known issue with rdflib is that it does not support nested service calls in its query
# body. An example of this can be seen in the
# <a href="examples/FederatedQuery_Example.ipynb">Federated Query Examples</a> notebook.
# We can utilize SPARQLWrapper to perform the same task. The following is a working
# example of utlizing two separate SERVICE calls and combining the results:
agent = "Chrome/86.0.4240.198" # default SPARQLWrapper agent does not work for wikidata
sparql = SPARQLWrapper("https://query.wikidata.org/sparql", agent=agent)
sparql.setQuery(
"""
PREFIX wd: <http://www.wikidata.org/entity/>
PREFIX wdt: <http://www.wikidata.org/prop/direct/>
PREFIX wikibase: <http://wikiba.se/ontology#>
PREFIX bd: <http://www.bigdata.com/rdf#>
SELECT ?p ?item
WHERE {
SELECT ?p ?item
WHERE
{
BIND(wikibase:label as ?p)
wd:Q28792126 wdt:P31 wd:Q146 .
service <https://query.wikidata.org/sparql>
{
SELECT ?item
WHERE {
?item wdt:P31 wd:Q146 .
}
LIMIT 5
}
}
}
"""
)
sparql.setReturnFormat(JSON)
if DBPEDIA_AVAILABLE:
results = sparql.query().convert()
results["results"]["bindings"]
results_df = pd.json_normalize(results["results"]["bindings"])
results_df
# ### Example using Wikidata and in-built service call
# In this example, we use SPARQLWrapper to query Wikidata via the Wikidata Query Service
# and organize the output response using pandas:
sparql = SPARQLWrapper("https://query.wikidata.org/sparql", agent=agent)
sparql.setQuery(
"""
PREFIX wd: <http://www.wikidata.org/entity/>
PREFIX wdt: <http://www.wikidata.org/prop/direct/>
PREFIX wikibase: <http://wikiba.se/ontology#>
PREFIX bd: <http://www.bigdata.com/rdf#>
SELECT ?item ?itemLabel
WHERE
{
?item wdt:P31 wd:Q146 .
service wikibase:label { bd:serviceParam wikibase:language "en" }
}
"""
)
sparql.setReturnFormat(JSON)
if DBPEDIA_AVAILABLE:
results = sparql.query().convert()
results_df = pd.json_normalize(results["results"]["bindings"])
results_df[["item.value"]].head()
# For more examples and usage of SPARQLWrapper, please visit the Github repo found here:
# https://github.com/RDFLib/sparqlwrapper
|
examples/RemoteQuery_Example.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala 2.10.4
// language: scala
// name: spark
// ---
// ### Getting started with SystemML
// Exercise objectives:
// - Load the SystemML library
// - Run basic SystemML code
// %AddJar https://sparktc.ibmcloud.com/repo/latest/SystemML.jar
import org.apache.sysml.api.MLContext
// Import the SQLContext to use some of its capabilites. This SQLContext library comes from the Apache Spark library.
import org.apache.spark.sql.SQLContext
val sqlCtx = new SQLContext(sc)
// Now do the same and create the MLContext variable from the SparkContext
val ml = new MLContext(sc)
val dml = """
X = rand(rows=100, cols=10)
sumX = sum(X)
outMatrix = matrix(sumX, rows=1, cols=1)
write(outMatrix, " ", format="csv")
"""
// Register the output variable.
ml.reset()
ml.registerOutput("outMatrix")
// Execute the script
val out = ml.executeScript(dml)
// Get the outputMatrix
val outMatrix = out.getDF(sqlCtx, "outMatrix")
// Print the matrix
outMatrix.show
|
Intro_Apache_SystemML.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pyEOF import *
import xarray as xr
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# create a function for visualization convenience
def visualization(da, pcs, eofs_da, evf):
fig = plt.figure(figsize = (6,12))
ax = fig.add_subplot(n+1,2,1)
da.mean(dim=["lat","lon"]).plot(ax=ax)
ax.set_title("average air temp")
ax = fig.add_subplot(n+1,2,2)
da.mean(dim="time").plot(ax=ax)
ax.set_title("average air temp")
for i in range(1,n+1):
pc_i = pcs["PC"+str(i)].to_xarray()
eof_i = eofs_da.sel(EOF=i)["air"]
frac = str(np.array(evf[i-1]*100).round(2))
ax = fig.add_subplot(n+1,2,i*2+1)
pc_i.plot(ax=ax)
ax.set_title("PC"+str(i)+" ("+frac+"%)")
ax = fig.add_subplot(n+1,2,i*2+2)
eof_i.plot(ax=ax,
vmin=-0.75, vmax=0.75, cmap="RdBu_r",
cbar_kwargs={'label': ""})
ax.set_title("EOF"+str(i)+" ("+frac+"%)")
plt.tight_layout()
plt.show()
# %matplotlib inline
# -
# ## load sample data
# note: if you got the complain of No module named 'pooch', please install "scikit-image"
# +
# load the DataArray
da = xr.tutorial.open_dataset('air_temperature')["air"]
print(da)
# create a mask
mask = da.sel(time=da.time[0])
mask = mask.where(mask<250).isnull().drop("time")
# get the DataArray with mask
da = da.where(mask)
da.sel(time=da.time[99]).plot()
plt.show()
# -
# convert DataArray to DataFrame
df = da.to_dataframe().reset_index() # get df from da
display(df.head(5))
print("DataFrame Shape:",df.shape)
# ## EOF Analysis
# ### reshape the dataframe to be [time, space]
df_data = get_time_space(df, time_dim = "time", lumped_space_dims = ["lat","lon"])
display(df_data.head(5))
print("DataFrame Shape:",df_data.shape)
# ### varimax rotated PCA analysis
# +
n = 4
pca = df_eof(df_data,pca_type="varimax",n_components=n)
eofs = pca.eofs(s=2, n=n) # get eofs
eofs_da = eofs.stack(["lat","lon"]).to_xarray() # make it convenient for visualization
pcs = pca.pcs(s=2, n=n) # get pcs
evfs = pca.evf(n=n) # get variance fraction
# plot
visualization(da, pcs, eofs_da, evfs)
# -
# ### unrotated EOFs analysis
# +
n = 4 # define the number of components
pca = df_eof(df_data) # implement EOF
eofs = pca.eofs(s=2, n=n) # get eofs
eofs_da = eofs.stack(["lat","lon"]).to_xarray() # make it convenient for visualization
pcs = pca.pcs(s=2, n=n) # get pcs
evfs = pca.evf(n=n) # get variance fraction
# plot
visualization(da, pcs, eofs_da, evfs)
# -
# ## compare with Eofs package (unrotated EOFs)
# link: https://ajdawson.github.io/eofs/latest/
# +
from eofs.standard import Eof
from sklearn.preprocessing import StandardScaler
solver = Eof(StandardScaler().fit_transform(df_data.values))
s_pcs = pd.DataFrame(data=solver.pcs(npcs=4, pcscaling=2),
columns = pcs.columns,
index = pcs.index)
s_eofs = pd.DataFrame(data = solver.eofs(neofs=4, eofscaling=2),
columns = eofs.columns,
index = eofs.index)
s_eofs_da = s_eofs.stack(["lat","lon"]).to_xarray() # make it convenient for visualization
s_evfs = solver.varianceFraction(neigs=4)
# plot
visualization(da, s_pcs, s_eofs_da, s_evfs)
|
docs/notebooks/basic_usage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import load_raw_data as lrd
from transforms3d import euler
from utils import nearest
import matplotlib.pyplot as plt
import seaborn as sns;sns.set(color_codes=True)
# %matplotlib notebook
def parse_dates(timestamp):
"""
parse_dates(timestampstring) takes a timestamp string formatted as Year-month-dayThour:min:sec.decimals+Timezone
and converts it into a datetime.datetime format, ignoring the timezone and the last decimal, keeping a microsecond
precision.
"""
return pd.datetime.strptime(timestamp[:26], '%Y-%m-%dT%H:%M:%S.%f')
filepath = './Data/Pilot/Jesse_FirstPilot/head.csv'
hd = lrd.load_head_data(filepath)
hd.head()
# determine the frequency of the signal
# first, get the minimum difference between timestamps:
np.diff(hd.index.values).min()
# Then, the frequency is 1 over that period
freq = 1e9 / np.diff(hd.index.values).min().astype(int)
np.diff(hd.index.values).min()
ed = lrd.load_event_data('./Data/Pilot/Jesse_FirstPilot/events.csv')
ed = ed.drop(['Value1','Value2'],axis=1)
ed.head()
trialstart_times = ed[ed['Name']=='DecisionStart'].index
ser = hd.loc[nearest(hd.index,trialstart_times[0])]
def series2mat4(head_pos):
return np.array([[head_pos.loc['Value.M11'],head_pos.loc['Value.M21'],head_pos.loc['Value.M31'],head_pos.loc['Value.M41']],
[head_pos.loc['Value.M12'],head_pos.loc['Value.M22'],head_pos.loc['Value.M32'],head_pos.loc['Value.M42']],
[head_pos.loc['Value.M13'],head_pos.loc['Value.M23'],head_pos.loc['Value.M33'],head_pos.loc['Value.M43']],
[head_pos.loc['Value.M14'],head_pos.loc['Value.M24'],head_pos.loc['Value.M34'],head_pos.loc['Value.M44']]])
head_position = series2mat4(ser)
## IMPORTANT: Watch out for gimbal lock.
euler_angles = euler.mat2euler(head_position)
np.degrees(euler_angles)
euler_angles = euler.mat2euler(head_position,'syzx')
np.degrees(euler_angles)
# Let's now find the end point location for the first trial
hitneutral_times = ed[ed['Name']=='Neutral'].index
ser = hd.loc[nearest(hd.index,hitneutral_times[0])]
head_position = series2mat4(ser)
euler_angles = euler.mat2euler(head_position)
np.degrees(euler_angles)
# +
# now get all of them in between
# -
trial1_trajectory = hd.loc[trialstart_times[0]:hitneutral_times[0]]
list_of_matrices = [series2mat4(hd.iloc[x]) for x in range(trial1_trajectory.shape[0])]
angles = np.array([np.degrees(euler.mat2euler(i,'syxz')) for i in list_of_matrices])
plt.plot(angles[:,1])
plt.title('Rotation trajectory around Y axis in first trial')
plt.show()
# ### Now let's extract the average for left and right
targetleft_times = ed[ed['Name']=='TargetLeft'].index
targetright_times = ed[ed['Name']=='TargetRight'].index
trial_numbers = np.argsort(targetleft_times.append(targetright_times))
trial_numbers
# get the indides (iloc in dataframe) of the end of each trial left and right
end_trial_indices_left = [ed.index.get_loc(trial)+1 for trial in targetleft_times]
end_trial_indices_right = [ed.index.get_loc(trial)+1 for trial in targetright_times]
# and now get the corresponding timestamps
end_trial_times_left = ed.iloc[end_trial_indices_left].index
end_trial_times_right = ed.iloc[end_trial_indices_right].index
# +
# let's do this differently. All at once, and then determine left and right after
start_trial_times = targetleft_times.append(targetright_times).sort_values()
end_trial_times = end_trial_times_left.append(end_trial_times_right).sort_values()
# -
# here, extract the list of left-right
target_sides = ed[ed.Name.str.get(0).isin(['T'])].reset_index()
# +
trajectories = []
counter = 0
# Left trials
for i, (start, end) in enumerate(zip(start_trial_times,end_trial_times)):
trial_trajectory = hd.loc[start:end]
trial_trajectory = trial_trajectory.resample('0.01S').pad()
trial_trajectory.loc[:,'Trial number'] = i
trial_trajectory.loc[:,'Target side'] = target_sides.iloc[i]['Name']
trial_trajectory['Trial time'] = trial_trajectory.index - trial_trajectory.index[0]
trajectories.append(trial_trajectory)
trajectories_df = pd.concat(trajectories).sort_index()
# -
# convert to matrices and then to angles
list_of_matrices = [series2mat4(trajectories_df.iloc[x]) for x in range(trajectories_df.shape[0])]
angles = np.array([np.degrees(euler.mat2euler(mat,'syzx')) for mat in list_of_matrices])
angles_df = pd.DataFrame(angles,index=trajectories_df.index,columns =['Y rotation','X rotation','Z rotation'])
trajectories_df = trajectories_df.join(angles_df)
trial_starts = trajectories_df[trajectories_df['Trial time']==trajectories_df.iloc[1]['Trial time']]
zeropoint = trial_starts['Y rotation'].mean()
trajectories_df['Y angle'] = trajectories_df['Y rotation'] - zeropoint
fig = plt.figure()
ax = sns.tsplot(data=trajectories_df, time="Trial time", value='Y angle', unit='Trial number',condition='Target side')
plt.title('Rotation Trajectory')
plt.xlabel('Time (seconds)')
#plt.savefig('./Figures/rotation_trajectory.png')
plt.show()
# ### Now, lets have a look at the distribution of movement endpoints in terms of Y angle and reaction times.
# TODO: fix trial numbering system so this works
RT=[]
for i in trajectories_df['Trial number'].unique():
idx = trajectories_df['Trial number'] == i
RT.append(trajectories_df[idx]['Trial time'].max())
trials = pd.DataFrame(index=trajectories_df['Trial number'].unique(),
columns=['RT'],
data=np.array(np.array(RT)))
trials.index.name = 'Trial'
# add the target side info to this dataframe
trials['Target side'] = target_sides['Name']
trials['Reaction time (ms)']= trials['RT'].apply(lambda x: x.microseconds/1000)
sns.distplot(trials['Reaction time (ms)'],rug=True)
# plot left and right separately
sns.distplot(trials.loc[trials['Target side']=='TargetRight','Reaction time (ms)'],
kde_kws={'label':'TargetRight'})
sns.distplot(trials.loc[trials['Target side']=='TargetLeft','Reaction time (ms)'],
kde_kws={'label':'TargetLeft'})
plt.title('Reaction time histograms with kernel density estimates')
# in order to fit a normal distribution instead: >>> from scipy.stats import norm, then fit=norm as argument
from scipy.stats import norm
# plot left and right separately
sns.distplot(trials.loc[trials['Target side']=='TargetRight','Reaction time (ms)'],
kde=False,
fit=norm,
fit_kws={'color':'b','label':'TargetRight'},
label='TargetRight')
sns.distplot(trials.loc[trials['Target side']=='TargetLeft','Reaction time (ms)'],
kde=False,
fit=norm,
fit_kws={'color':'g','label':'TargetLeft'})
plt.title('Reaction time histograms with Gaussian fit')
# ### Distribution of movement endpoints
startpoints=[]
endpoints=[]
for i in trajectories_df['Trial number'].unique():
idx = trajectories_df['Trial number'] == i
startpoints.append(trajectories_df[idx].iloc[1]['Y angle'] )
endpoints.append(trajectories_df[idx].iloc[-1]['Y angle'] )
trials['Starting points'] = startpoints
trials['Movement endpoints'] = endpoints
trials.head()
sns.distplot(trials.loc[trials['Target side']=='TargetRight','Movement endpoints'],
kde_kws={'label':'TargetRight'})
sns.distplot(trials.loc[trials['Target side']=='TargetLeft','Movement endpoints'],
kde_kws={'label':'TargetLeft'})
plt.xlim([-80,80])
plt.xlabel('Movement endpoint (degrees separation from midline)')
trial_results = ed[(ed['Name']=='Neutral') | (ed['Name']=='Missed') | (ed['Name']=='Hit') | (ed['Name']=='Penalty')]
trials['Outcome'] = np.array(trial_results['Name'])
trials.head()
mean_end_right = trials.loc[trials['Target side']=='TargetRight','Movement endpoints'].mean()
mean_end_left = trials.loc[trials['Target side']=='TargetLeft','Movement endpoints'].mean()
|
movement_endpoints.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="t9LpcoF-ATDz"
# # Yahoo Text Classification: ULMFiT vs BERT
#
# **NOTE: This notebook has been executed in Google Colab Pro Edition, reduce max_len in BERT model to avoid resource runout error.**
#
# ~ By <NAME>
#
# # Introduction
# Text classification is a classic Natural Language Processing (NLP) problem. The task is to assign predefined categories to a given text sequence.
#
# Tasks involved in this notebook:
# 1. General Classification
# 2. Language Modelling
# 3. Text Classification
#
# # Interesting Questions
# ## 1. Ask yourself why would they have selected this problem for the challenge? What are some gotchas in this domain I should know about?
#
# Ans: Text classification is the task of assigning a sentence or document to an appropriate category. The categories depend on the chosen dataset and can range from topics. Text Classification, both through supervised and unsupervised approach, finds application in various fields such as social media, marketing, customer experience management, digital media etc. This task is currently in SOTA, performaces are different for different complexities of data and the architecture used.
#
# Domains envisioned: NLP, supervised learning, Attention Mechanism, Transformer, Tokenization, RNN, LSTM
#
# ## 2. What is the highest level of accuracy that others have achieved with this dataset or similar problems / datasets ?
# Note: These results are obtained by using the whole dataset without dropping any rows.
#
# \# | Dataset | Model | Eval Metric | Value
# --- | --- | --- | --- | ---
# 01 | AG News | XLNet | Error | 4.8
# 02 | DBPedia | XLNet | Error | 0.62
# 03 | TREC-6 | USE + CNN | Error | 1.93
# 04 | 20NEWS | SSGC | Accuracy | 88.6
# 05 | Yahoo! Answers | BERT | Accuracy | 77.62
# 06 | YELP-5 | HAHNN | Accuracy | 73.28
#
# ## 3. What types of visualizations will help me grasp the nature of the problem / data?
# Ans:
# 1. Distributions to charts to check class balance
#
# 2. Distributions to charts to check where most nan values are present
#
# 3. Boxplots to visualize question lengths per category
#
# 4. Check class distribution for nan columns
#
# 5. Wordcloud visualization for most common words in the corpus
#
# 6. T-SNE visualization of target clusters using plotly (optional)
#
# ## 4. What feature engineering might help improve the signal?
#
# Ans: Combining all textual data to one column will help increase the corpus size and also increase relevant information for the target variable.
#
# ## 5. Which modeling techniques are good at capturing the types of relationships I see in this data?
#
# Ans: Language models, SOTA transformers like BERT, GPT-2, GPT-3, XLM, ROBERTa, XLNet and RNNs', LSTMs', GRUs'.
#
# ## 6. Now that I have a model, how can I be sure that I didn't introduce a bug in the code? If results are too good to be true, they probably are!
#
# Ans: We can perform hypothesis testing using custom examples and feed it the model to check the results with confidence score and target class.
#
# ## 7. What are some of the weaknesses of the model and and how can the model be improved with additional work
#
# Ans: I will be using ULMFiT and BERT models to compare my results. ULMFiT uses a language model and a classifier model to predict the classes. The ULMFiT langauge model is performing quite bad in terms of accuracy for this dataset (100000 rows only) but the accuracy model is doing great compared to current benchmarks on Yahoo! Answers dataset. The BERT model performs better than the ULMFiT model and is also the current best SOTA model for this dataset.
#
# ULMFiT is getting confused between few example, when I input the model with text as: **'homebrew is not working on macosx'** the ULTFiT model predicts the class as **'Education & Reference' with 35% confidence score** where the BERT model predicts **['Computers & Internet'] with 97% confidence**. In real-life scenario, the ULMFiT model prediction seems to correct. As homebrew and MacOSX strings may/may not come under **'Education & Reference'**. On the other hand, BERT seems to do a great job is predicting the classes accurately with a good confidence.
#
# ULMFiT can be improved if more dataset is passed for the language model. Fine-tuning the parameters using optuna can also slightly increase the accuracy. But overall, BERT is the clear winner for this task.
# + [markdown] id="JD4AMA3E94vJ"
# # Data Description
#
# The **Yahoo! Answers topic classification dataset** is constructed using 10 largest main categories. Each class contains 140,000 training samples and 6,000 testing samples. Therefore, the total number of training samples is 1,400,000 and testing samples 60,000 in this dataset. From all the answers and other meta-information, we only used the best answer content and the main category information.
#
# The file `classes.txt` contains a list of classes corresponding to each label.
#
# The files `train.csv` and `test.csv` contain all the training samples as comma-sparated values. There are 4 columns in them, corresponding to class index (1 to 10), question title, question content and best answer. The text fields are escaped using double quotes ("), and any internal double quote is escaped by 2 double quotes (""). New lines are escaped by a backslash followed with an "n" character, that is "\n".
#
# We'll start by mounting my google drive to colab, the dataset has been downloaded from [fastai](https://course.fast.ai/datasets#nlp).
# + colab={"base_uri": "https://localhost:8080/"} id="jo5OV8uX9tY0" outputId="0da5b9d8-6d01-4a92-d11f-9d0a0c5b3ed3"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="yoPytS4iB4w1"
# # EDA
#
# Let's explore the data using matplotlib and seaborn
# + id="GCbMLwPZ-c0S"
#@ Imports for data exploaration
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from wordcloud import WordCloud, STOPWORDS
stopwords = set(STOPWORDS)
# %matplotlib inline
# + id="B0pjTyksCEmo"
#@ Converting integers to strings to understand each label independently.
categories = {1: 'Society & Culture', 2: 'Science & Mathematics', 3: 'Health', 4: 'Education & Reference',
5: 'Computers & Internet', 6: 'Sports', 7: 'Business & Finance', 8: 'Entertainment & Music',
9: 'Family & Relationships', 10: 'Politics & Government'}
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="2E9mZk_ACXXe" outputId="d71f5186-e74b-4c61-8b68-427007080aee"
#@ Load raw training data using pandas
raw_data = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/train.csv',
names=['class', 'title', 'content', 'answer'])
#@ Mapping dict to target
raw_data['class'] = raw_data['class'].map(categories)
raw_data.head(3)
# + [markdown] id="DZ2buQ7DMyvZ"
# NaN values present?
# + colab={"base_uri": "https://localhost:8080/"} id="9QeoG0fKLrnd" outputId="6881afc8-643f-450e-e96e-74177b73ebbd"
raw_data.isna().sum()
# + [markdown] id="JG8qjcSBQXPl"
# We'll deal with this in preprocessing. Let's proceed with EDA
# + id="bvrSckjBQfzU"
#@ Add question length
raw_data['length'] = raw_data.loc[:, 'content'].astype(str).apply(len)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="8_Q5MA4iF3Y_" outputId="6960aa1d-7d90-4553-ba05-677acd7eeee6"
#@ Target distribution
plt.figure(figsize=(13, 7))
plt.style.use('ggplot')
sns.countplot(x = 'class', data = raw_data)
plt.title('Target distribution', fontweight='bold')
plt.xlabel('Target', fontweight='bold')
plt.xticks(rotation=45);
# + [markdown] id="hEb_qsiBdwXo"
# Insights from chart above:
# 1. No NaN values in target
# 2. All the classes are equally distributed
# 3. No need to use SMOTE since balanced
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="DfWbX4uKCs_D" outputId="f3dd3be6-095b-473c-c79d-5c1eb6ed145e"
#@ Exploring NaN values in dataset
fig = plt.figure(figsize=(15, 8))
plt.title("Distribution of NAN values")
plt.xlabel('Count')
plt.ylabel('Dataframe Columns')
raw_data.isna().sum().sort_values(ascending = True).plot(kind = 'barh')
plt.show();
# + [markdown] id="u8jvWxTqeB0f"
# Insights from chart above:
# 1. Almost 46% of the data is missing in content column
# 2. Around 0.017% of the data is missing in answers column
# 3. We'll drop nan values of content column in preprocessing, this will save us compute and remove irrelavant information
# + colab={"base_uri": "https://localhost:8080/", "height": 580} id="_qKaGK17IX-L" outputId="a0cd1b80-153c-4b9c-b620-b7e035569f4a"
#@ Boxplot of question lengths per category
plt.figure(figsize=(15, 8))
g = sns.boxplot(x='class', y='length', data=raw_data)
g.set_yscale('log')
plt.xticks(rotation=45)
plt.show();
# + [markdown] id="p7PtUqh4ehEo"
# Insights from chart above:
# 1. Most lengthy questions were from Family & Relationships category
# 2. Least from sports
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="d8nXn_8YCxTo" outputId="40127565-7655-4a80-bdfe-33ed7289ff29"
#@ Checking class distribution of NaN values as content
fig = plt.figure(figsize=(15, 8))
data_isnull = raw_data[raw_data['content'].isnull()]
index = pd.Index(data_isnull['class'])
index.value_counts().sort_values(ascending = True).plot(kind = 'barh')
plt.ylabel('Targets')
plt.xlabel('Count')
plt.show();
# + [markdown] id="iZm8iBYFgYdQ"
# Insights from chart above:
# 1. Most of the NaN values in content belongs to the Business & Finance class.
# 2. Least of the NaN values in content belongs to the Family & Relationships class.
# + id="vzVRjMP_HCtH"
#@ function to visualize word cloud for title - top occuring words in the corpus
def show_wordcloud(data, title = None):
wordcloud = WordCloud(
background_color='white',
stopwords=stopwords,
max_words=500,
max_font_size=40,
scale=3,
random_state=1
).generate(str(data))
fig = plt.figure(1, figsize=(12, 12))
plt.axis('off')
if title:
fig.suptitle(title, fontsize=20)
fig.subplots_adjust(top=2.3)
plt.imshow(wordcloud)
plt.show();
# + colab={"base_uri": "https://localhost:8080/", "height": 366} id="vmepYkChHCq1" outputId="7b3ae4d5-e46b-422f-f5f3-b3b4ad2e1f78"
#@ For title column
show_wordcloud(raw_data['title'])
# + colab={"base_uri": "https://localhost:8080/", "height": 366} id="26aiPInMHCob" outputId="06a09f0e-23ca-4a33-eb2b-00dfcfa4f729"
#@ For content column
show_wordcloud(raw_data['content'])
# + [markdown] id="RIGI06MPMYGR"
# # Preprocessing
# + colab={"base_uri": "https://localhost:8080/"} id="QKxrahbzHCko" outputId="bff4b360-b177-4a19-8f57-6ee5039e52f0"
#@ Imports for preprocessing data
import pandas as pd
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
import string
# + id="t4JCgxsxRbPD"
#@ Converting integers to strings to understand each label independently.
categories = {1: 'Society & Culture', 2: 'Science & Mathematics', 3: 'Health', 4: 'Education & Reference',
5: 'Computers & Internet', 6: 'Sports', 7: 'Business & Finance', 8: 'Entertainment & Music',
9: 'Family & Relationships', 10: 'Politics & Government'}
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="ysnHAf3ZRa6F" outputId="7e6253e2-1d0c-4a5e-c100-dbfe5de0ab4c"
#@ Load raw training data using pandas
raw_data = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/train.csv',
names=['class', 'title', 'content', 'answer'])
raw_data['class'] = raw_data['class'].map(categories)
raw_data.head(3)
# + colab={"base_uri": "https://localhost:8080/"} id="NoUy39PhECLN" outputId="1c842f88-7399-4b9c-f558-498f1386c366"
raw_data.shape
# + colab={"base_uri": "https://localhost:8080/"} id="paEOw-xWQpPK" outputId="67b405f9-fa42-46e2-80f6-3051d3f48fce"
#@ Remove all NaN values in content
data = raw_data[raw_data['content'].notnull()]
data.shape
# + id="u-ltENFVRSI6"
#@ Saving to csv
data.to_csv('yahoo_train_notnull.csv', index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="09vHSBueQreL" outputId="c270d47f-17fa-4704-85f1-ee0befa834c4"
#@ Remove capitalization, stopwords, and punctuation
def process(text) :
#@ Remove punctuation/captizalization
text = str(text)
text = text.lower()
nopunc = [char for char in text if char not in string.punctuation]
nopunc = ''.join(nopunc)
#@ Remove stopwords
procList = [word for word in nopunc.split() if word not in stopwords.words('english')]
return ' '.join(procList)
def cleanAndSave(source, dest) :
#@ Load data
df = pd.read_csv(source, names=['class', 'title', 'content', 'answer'])
X1, X2, X3, Y = df['title'], df['content'], df['answer'], df['class']
#@ Empty lists to append everything to one column
title, question, answer, clss = [], [], [], []
#@ Process text
for i in range(0, len(X1)) :
title.append(process(X1[i]))
question.append(process(X2[i]))
answer.append(process(X3[i]))
clss.append(Y[i])
#@ Merge processed text columns into dataframe and save
df = pd.DataFrame({"title" : title, "question" : question, "answer" : answer, "class" : clss})
df['text'] = df['title'].map(str) + ' ' + df['question'].map(str) + ' ' + df['answer'].map(str)
df_save = pd.DataFrame({'text': df['text'], "class" : df['class']})
df_save.to_csv(dest, index=False)
cleanAndSave('/content/yahoo_train_notnull.csv', '/content/yahoo_train_notnull_clean.csv')
print('done')
# + [markdown] id="UjjPB-2Thgcd"
# The above cell took around 4 hours to execute so I downloaded the preprocessed file from colab and saved in my drive.
# + [markdown] id="aj8gQ8KKaHeA"
# # Modeling: Using ULMFiT for Text Classification with FastAI
# ULMFiT, by <NAME> et. al. from fast.ai, gives us an incredibly powerful method to classify text using language modelling and transfer learning. ULMFiT stands for Universal Language Model Fine-tuning for Text Classification and is a transfer learning technique that involves creating a Language Model that is capable of predicting the next word in a sentence, based on unsupervised learning of the WikiText 103 corpus. The ULMFiT model uses multiple LSTM layers, with dropout applied to every layer (the secret sauce), developed by <NAME> (Salesforce) as the AWD-LSTM architecture.
# + colab={"base_uri": "https://localhost:8080/"} id="bHDvixTBcfcz" outputId="dc0087d4-08a9-4475-c074-3ba7e98b9880"
# !nvidia-smi
# + [markdown] id="3jd6PL2oodbx"
# ### 1. Load the preprocessed data
# + id="K7ghs28EazSW"
#@ Imports
from fastai.text import *
# + id="IMssjhfRQ8qz"
#@ Get cleaned dataset
path = '/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/yahoo_train_notnull_clean.csv'
cleaned_data = pd.read_csv(path)
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="5U0Jbx4uSOZt" outputId="5e03d697-1954-4cc5-9031-e6811a21532c"
#@ Fixing dataframe for ULMFiT
cleaned_data = cleaned_data.sample(frac=0.13).reset_index(drop=True)
cleaned_data = cleaned_data[['class', 'text']]
cleaned_data.head(2)
# + colab={"base_uri": "https://localhost:8080/"} id="Jh8NHQvIa-5u" outputId="5b7261f3-63ac-4f09-d6b7-057b361d796c"
cleaned_data.shape
# + [markdown] id="fuLgNHB4anGm"
# ### 2. Create train & validation datasets and FastAI data bunch
# + colab={"base_uri": "https://localhost:8080/"} id="yN3OBA0TSUWV" outputId="bc26b632-ca10-43e8-aa87-24d36d0d8ce3"
from sklearn.model_selection import train_test_split
df_trn, df_val = train_test_split(cleaned_data, stratify = cleaned_data['class'], test_size = 0.3)
df_trn.shape, df_val.shape
# + [markdown] id="5xlKtIuGTuYh"
# Next, we will setup our data in the format that FastAI requires it to be in. FastAI provides simple functions to create Language Model and Classification “data bunch”.
#
# Creating a data bunch automatically results in pre-processing of text, including vocabulary formation and tokenization.
#
# TextLMDataBunch creates a data bunch for language modelling. In this, labels are completely ignored. Instead, data is processed so that the RNN can learn what word comes next given a starting word. Read the documentation [here](https://docs.fast.ai/text.data.html#TextLMDataBunch).
#
# TextClasDataBunch sets up the data for classification. Labels play a key role here. We can also set the batch size for learning by changing the bs parameter. Read the documentation [here](https://docs.fast.ai/text.data.html#TextClasDataBunch).
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="zhcH6OGhauYB" outputId="e3a95d7a-25f3-4447-b9e4-cabf8daa6ee8"
# Language model data
data_lm = TextLMDataBunch.from_df(train_df = df_trn, valid_df = df_val, path = "")
# Classifier model data
data_clas = TextClasDataBunch.from_df(train_df = df_trn, valid_df = df_val, path = "", vocab=data_lm.train_ds.vocab, bs=32)
# + [markdown] id="aOXTx4FZT_HJ"
# You can print out a sample of the batch using this line:
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="Rq2zcqL9a1KB" outputId="fb9fa6e4-be4d-4993-9967-6cc7c9cc964a"
data_clas.show_batch()
# + [markdown] id="K3wHLuvjUFUZ"
# The xx___ tags represent the aspects of language in a way that the computer can understand. The xxbos tag marks the beginning of a sentence. The xxmaj tag is used to imply that the first letter of the next word is capitalized.
# With this in place, we are ready to create a language model and classify!
#
# + id="wyWuQzDIbL3Q"
#@ Saving language model in drive
data_lm.save('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/data_lm_export.pkl')
data_clas.save('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/data_clas_export.pkl')
# + id="8m-y8CNviAQN" colab={"base_uri": "https://localhost:8080/"} outputId="4ba67ccf-402d-4d18-fa99-95b8f6fb90ae"
#@ Loading language model from drive
data_lm = load_data('./', '/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/data_lm_export.pkl')
data_clas = load_data('./', '/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/data_clas_export.pkl',bs=64)
# + [markdown] id="C3yCaKDWipmC"
# ### 3. Create and Train the Language Model
#
# Creating a language model with the aforementioned AWD-LSTM model is done using the code below
# + id="ty9bJcLUiJ33" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9778a6f7-fbec-4c7a-f9ff-b733027a79af"
learn = language_model_learner(data_lm, AWD_LSTM, drop_mult=0.3)
# + [markdown] id="xbneTirXUWIZ"
# - data_lm is the language model data bunch
# - AWD-LSTM is the model architecture
# - drop_multi is the drop-out
#
# Next up, let’s find the optimal learning rate to train our language model
# + colab={"base_uri": "https://localhost:8080/", "height": 408} id="1OkLJ114irWp" outputId="49c37459-671b-4274-8de1-8491d459e5e5"
learn.lr_find()
learn.recorder.plot(suggestion=True)
min_grad_lr = learn.recorder.min_grad_lr
# + [markdown] id="V4oQuWDFUfha"
# lr_find() is a built in fast.ai function that runs a few epochs on the model to plot loss, and then calculate the minimum gradient.
#
# Now, let’s use this learning rate to train the language model:
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="5yoVzcsHitve" outputId="a6500683-46c6-41eb-cbbd-86129e2486c8"
learn.fit_one_cycle(2, min_grad_lr)
# + [markdown] id="2HNSWa1CUg34"
# We can do a few more epochs after unfreezing all the layers. This process will train the whole neural network rather than just the last few layers.
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="UCGN1vvNixuE" outputId="fa5f091f-f1f5-49d8-fb91-f2f906fabb3c"
learn.unfreeze()
learn.fit_one_cycle(2, 1e-3)
# + [markdown] id="HIBjwLucUnrE"
# Our language model only achieved around 12% accuracy, but that is okay. This accuracy represents how well the model does at predicting the next word, given one word. And 12% means that 1 out of 2 times, the model accurately predicts the next word. Pretty impressive!
#
# You can have some fun playing with the language model… here, we can ask the model to predict what comes after “How do”, till 10 words:
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="_kMDzMeki8M-" outputId="31bf23c0-41c9-42b2-8386-db7778d30330"
learn.predict("How do", n_words=10)
# + [markdown] id="S8ZOqGFRUvTV"
# Clearly, the sentence generated is not very meaningful, but it is grammatically accurate.
#
# Finally, let’s save the language model encoder so that we can load it later in our classifier
# + id="_m0M9TgQi4ad"
learn.save_encoder('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/ft_enc')
# + [markdown] id="Z621T1F1jOmc"
# ### 4. Using the Language Model to Train the Classifier
# + [markdown] id="TPFNeWslU40v"
# Creating and training the the text classifier is very similar to training the language model.
#
# Start by creating the text_classifier_learner with the data_clas DataBunch and the AWD_LSTM architecture. Then, you can load the language model encoder.
# + colab={"base_uri": "https://localhost:8080/"} id="TH4q2NNGi_2P" outputId="077e6896-a6cd-4b3e-a42f-88ec9f3746be"
learn = text_classifier_learner(data_clas, AWD_LSTM, drop_mult=0.5)
learn.load_encoder('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/ft_enc')
# + [markdown] id="q6T7usV4U8Tn"
# Let’s again find the optimal learning rate to start with
# + colab={"base_uri": "https://localhost:8080/", "height": 479} id="l0QdqHkxjS8Z" outputId="352b744a-077e-41ee-94a1-b0db91c3a7e6"
learn.lr_find()
learn.recorder.plot(suggestion=True)
min_grad_lr = learn.recorder.min_grad_lr
# + [markdown] id="gA05PlpxU-BN"
# To train the classifier, we will use a technique called gradual unfreezing. We can start by training the last few layers, then go backwards and unfreeze and train layers before. We can use the learner function learn.freeze_to(-2) to unfreeze the last 2 layers.
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="4LA1KzbcjWrN" outputId="9e1e3cf0-06bb-44df-e8ee-3a7db7e5421c"
learn.fit_one_cycle(2, min_grad_lr)
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="7V23LP3ojaIq" outputId="5e277ba8-e850-4189-da0e-636c1fa0e415"
learn.recorder.plot_losses()
# + colab={"base_uri": "https://localhost:8080/", "height": 397} id="_KMC2iBfjhZM" outputId="79b365e2-bfde-4d8d-a93e-fe9071de30ce"
learn.freeze_to(-2)
learn.fit_one_cycle(4, slice(5e-3, 2e-3), moms=(0.8,0.7))
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="yQJx27oQVGG5" outputId="2ef5a264-8b33-4592-9236-ad63ea8a5cfa"
learn.recorder.plot_losses()
# + [markdown] id="XcQaMUiZjib7"
# Finally, let us unfreeze all layers and train the model at a low learning rate.
# + colab={"base_uri": "https://localhost:8080/", "height": 397} id="PC7a1gm6VGEb" outputId="4d51b63c-66e5-4c8f-d5c7-83b784f02e9d"
learn.unfreeze()
learn.fit_one_cycle(4, slice(2e-3/100, 2e-3), moms=(0.8,0.7))
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="u9rz6tZ8Xy4i" outputId="3c5d9332-e142-45c6-a317-b713a1690479"
learn.recorder.plot_losses()
# + [markdown] id="Se-IQGwQ3soY"
# ### 5. Analyzing our results
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="CiYIhckD3ik1" outputId="ae4edace-74a5-49f1-a38e-498d7ee1ccc2"
preds,y,losses = learn.get_preds(with_loss=True)
interp = ClassificationInterpretation(learn, preds, y, losses)
interp.plot_confusion_matrix()
# + colab={"base_uri": "https://localhost:8080/"} id="64CsiRbNw21l" outputId="c43894e6-1747-4afa-97af-5d8966a744e7"
preds,tensor,probs=learn.predict("football is really nice")
probs
# + colab={"base_uri": "https://localhost:8080/"} id="O2bxiHs13iid" outputId="ddfcf958-2af4-4732-e7ad-95fb8eadb1a6"
preds,tensor,probs=learn.predict("football is really nice")
top_pred = probs.argsort(descending=True)[:1]
classes=learn.data.classes
labels = []
confidence = []
for i in top_pred:
x = classes[i]
p = probs[i]
labels.append(x)
print("The following sentence belongs to", labels, "with confidence as", p)
# + colab={"base_uri": "https://localhost:8080/"} id="ahnZdjgEd5Im" outputId="936d3e0c-0651-4f38-e4b5-ab0c8d0bd217"
preds,tensor,probs=learn.predict("homebrew is not working on macosx")
top_pred = probs.argsort(descending=True)[:1]
classes=learn.data.classes
labels = []
confidence = []
for i in top_pred:
x = classes[i]
p = probs[i]
labels.append(x)
print("The following sentence belongs to", labels, "with confidence as", p)
# + [markdown] id="tYfKG8jXgA4K"
# # Modeling: Using BERT for Text Classification with Ktrain Wrapper
# + colab={"base_uri": "https://localhost:8080/"} id="zB5QLjlGt41Y" outputId="0c342ba5-ceb4-4052-8eba-cbb486ef5a3a"
# !nvidia-smi
# + colab={"base_uri": "https://localhost:8080/"} id="YKBBcS6qgAvk" outputId="27f733d5-92d7-4713-cdfd-3dfbc28297e3"
#@ Necessary installations
# !pip install -q tensorflow==2.3.1
# !pip install -q ktrain==0.23.1
# + [markdown] id="V9TmTUjct8i-"
# ### 1. Load the preprocessed data
# + id="PBaQFxaIgAtX"
#@ Imports
import ktrain
from ktrain import text
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID";
os.environ["CUDA_VISIBLE_DEVICES"]="0";
# + id="gCIh_EqqgArC"
#@ Get clean data
path = '/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/yahoo_train_notnull_clean.csv'
cleaned_data = pd.read_csv(path)
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="UBpKxjsqgAol" outputId="ac08be80-caf0-4c99-ee10-d12452c5c94b"
#@ Fix data for BERT model
cleaned_data = cleaned_data.sample(frac=0.13).reset_index(drop=True)
cleaned_data = cleaned_data[['class', 'text']]
cleaned_data.head(2)
# + [markdown] id="VOwk1hRQt_3k"
# ### 2. Create train, validation & preprocessing datasets using texts_from_df in ktrain
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="Nt7YpLgxd84w" outputId="95b508ff-7fca-416f-c848-3b156bc18d7f"
(X_train, y_train), (X_test, y_test), preproc = text.texts_from_df(train_df=cleaned_data,
text_column='text',
label_columns='class',
maxlen=64,
preprocess_mode='bert')
# + [markdown] id="J6Hj3FVnuM2Y"
# ### 3. Create and Train the BERT Model
# + colab={"base_uri": "https://localhost:8080/"} id="6mYsAMg5h8ad" outputId="89be7b3b-1bc0-4618-b62e-be0340638e7c"
model = text.text_classifier(name='bert',
train_data=(X_train, y_train),
preproc=preproc)
# + id="O-aRNTV2iNC4"
#@ Setting up learner
learner = ktrain.get_learner(model=model,
train_data=(X_train, y_train),
val_data=(X_test, y_test),
batch_size=64)
# + colab={"base_uri": "https://localhost:8080/"} id="izppMLN8iOfQ" outputId="ab1a61d8-2843-4d80-d276-dc9dd1222e11"
#@ Fitting model using one-cycle poliy, lr=2e-5 from research paper
learner.fit_onecycle(lr=2e-5, epochs=3);
# + id="hwBgwlGUiPwS"
#@ Get predictor using preproc dataset
predictor = ktrain.get_predictor(learner.model, preproc)
# + colab={"base_uri": "https://localhost:8080/"} id="pxukNSV4sPMH" outputId="dd966dc5-28f2-4b0f-c2f3-c61553fd8486"
classes = predictor.get_classes()
classes
# + [markdown] id="-IVb_J9HzQ_T"
# ### 6. Experimenting results
# Example 1.
# + id="eMakacvcpIES"
data = ["football is really nice"]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="TwgxBmCUvSzi" outputId="807e894f-de97-4143-aab2-477ac83ac2b5"
predictor.predict(data)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="XV1w5XJqpICQ" outputId="fe2108a9-d5d3-4054-c554-0ec6be7b475f"
data = ['football is really nice']
classes=predictor.predict(data)
probs=predictor.predict(data, return_proba=True)
# probs = torch.Tensor(probs)
# top_pred = probs.argsort(descending=True)[:,0]
print("The following sentence belongs to", classes, "with confidence:", probs.max())
# + [markdown] id="PJ-DGEdEzWkU"
# Example 2.
# + id="bxSYFDQNpH_1"
data = ['homebrew is not working on macosx']
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="IqLY9ED_pH9i" outputId="5debec81-ac75-42d4-ccc8-1de954c984b7"
predictor.predict(data)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="Q6FWnoIQyf1r" outputId="ee03d120-a9e3-445c-f272-4084da9d08dd"
data = ['homebrew is not working on macosx, can someone solve this problem?']
classes=predictor.predict(data)
probs=predictor.predict(data, return_proba=True)
# probs = torch.Tensor(probs)
# top_pred = probs.argsort(descending=True)[:,0]
print("The following sentence belongs to", classes, "with confidence:", probs.max())
# + [markdown] id="TOkvFTQTzXx2"
# Example.3
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="It_peHAayhbM" outputId="86982ca7-36c9-4656-f1fc-649f9d9186be"
data = ['im bored got the ideas two say hey to a monkey']
classes=predictor.predict(data)
probs=predictor.predict(data, return_proba=True)
# probs = torch.Tensor(probs)
# top_pred = probs.argsort(descending=True)[:,0]
print("The following sentence belongs to", classes, "with confidence:", probs.max())
# + id="2JVqdjWDzk1p"
#@ Saved in h5 format by default
predictor.save('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/my_predictor')
# + id="Z1wb9eP0zrcs"
# reloaded_predictor = ktrain.load_predictor('/content/drive/MyDrive/Colab Notebooks/Internships/Fellowship.ai/yahoo_answers_csv/my_predictor')
|
Yahoo!_Answers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise: Many electron tunnelling systems
# Having read notebooks 1-4, you should be in a position to tackle this exercise. It is designed to test aspects from each of the topics you have learned so far. A worked solution to this exercise is provided, with a small amount of explanation of the underlying physics, but we highly encourage you to fully attempt this exercise first before consulting the solutions. There truly is no better way to check you're up to speed with everything you ought to be than by having a go yourself!
# The system of interest is going to be a double antisymmetric well, with the exact form of the potential being
#
# $$ -\frac{6}{5} exp \bigg(\frac{-1}{125}(x-10)^4\bigg) - \frac{9}{10} exp\bigg(\frac{-1}{10}(x+10)^2\bigg), $$
#
# or in python format
# +
# -(6.0/5.0)*math.e**(-(1.0/125.0)*(x-10)**4)-(9.0/10.0)*math.e**(-(1.0/10.0)*(x+10)**2)
# -
# Now we'd like you to ** run iDEA, with time dependence and reverse engineering switched on for the non-interacting, exact and LDA cases.** Be sure to adjust your system parameters to suitable values to ensure convergence and to include the whole of the potential. We also recommend running the time dependence for at least $t_{max} = 10 $ a.u. to ensure you are able to see the features we're interested in. The perturbation applied at $t = 0 $ is just the default linear electric field so there is no need to change this for the time being (but you're more than welcome to adjust this and see what its effect is).
#
# **After iDEA has finished running, plot your ground state densities for the each of the above cases and the external potential on the same graph commenting on any differences.**
# +
# Import iDEA and parameters
# +
# Adjust the parameters to answer the exercise
# NB - if your system has trouble running iDEA from the notebook, try setting pm.run.verbosity to 'low'
# +
# Change the external potential to that given above
# +
# Optional (but recommended) - check that parameters file has updated as you expected
# +
# Hopefully everything is good to go now, so go ahead and run iDEA
# -
# Hopefully everything has worked as expected so far. Getting to this point means you're comfortable with the basics of running the iDEA code. If you're having problems with this, go back and read the "Getting Started with iDEA" notebook.
# +
# Import matplotlib and plot each of the densities you've calculated thus far
# -
# Use this cell to comment on the differences you see in the graphs:
#
#
# That's the first part done! Now let's have a look at why DFT has done so well. **Plot the reverse engineered exact KS potential and the external potential on the same graph and comment on the major differences**
# +
# Plot the KS potential and the external potential on the same graph
# -
# **Use this cell to comment on the major differences between between the two potentials:**
#
#
# Now we're ready to move onto looking at the time dependence of the system. **Animate how the exact density changes over time.**
# +
# Import the relevant packages for animations
# +
# Store your time dependent solution as an array
# +
# Set the background for the animation
# +
# Define the init function
# +
# Define the animation function
# +
# Display the animation
# -
# **Use this cell to comment of the major features of the time-evolution. How can we tell it is a tunnelling system?**
#
#
|
05_tunneling/tunneling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
#making database using dictionary
food_eating = {
'FMenuRating' : [1.9,2.7,3.5,4.6,7.9,8.1,9,9.1,9.3,9.5],
'Wastage' : [7,6.4,5.4,4.7,3,1.4,1,0.6,0.4,0.2],
'FEating' :[2.4,3,4.1,4.9,7.4,8.3,9,9.3,9.4,9.6]
}
#FEating=(FMeanuRating+10-FWastage)/2
df = pd.DataFrame(food_eating,columns=['FMenuRating','Wastage','FEating'])
df.head()
plt.scatter(df['FMenuRating'], df['FEating'], color='blue')
plt.title('FMenuRating Vs FEating', fontsize=14)
plt.xlabel('FMenuRating', fontsize=14)
plt.ylabel('FEating', fontsize=14)
plt.grid(True)
plt.scatter(df['Wastage'], df['FEating'], color='blue')
plt.title('Wastage Vs FEating', fontsize=14)
plt.xlabel('Wastage', fontsize=14)
plt.ylabel('FEating', fontsize=14)
plt.grid(True)
X = df[['FMenuRating','Wastage']]
Y = df['FEating']
X.head()
Y.head()
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X, Y)
print('Intercept: \n', model.intercept_)
print('Coefficients: \n', model.coef_)
b1 = model.intercept_
m1 = model.coef_[0]
m2 = model.coef_[1]
print(b1)
print(m1)
print(m2)
# +
New_FRating = 9
New_Wastage = 1
print ('Should you go? I will tell you: \n', model.predict([[New_FRating,New_Wastage]]))
# -
if( model.predict([[New_FRating,New_Wastage]])>=5):
print('yes')
else:
print('no')
|
messgoingpreiction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import numpy as np
import sys
import matplotlib.pyplot as plt
import torch
from tqdm import tqdm
sys.path.append('..')
sys.path.append('../visualization')
import viz_2d as viz
import acd
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# get dataset
import pickle as pkl
from dsets.imagenet import dset
imnet_dict = pkl.load(open('../dsets/imagenet/imnet_dict.pkl', 'rb')) # contains 6 images (keys: 9, 10, 34, 20, 36, 32)
# get model
from torchvision import models
model = models.vgg16(pretrained=True).to(device).eval()
model_type='vgg' # alexnet, vgg
# -
# # # cd propagation fig
# +
def vgg_track(im_torch, model):
mods = list(model.modules())[2:]
scores = []
x = im_torch.clone()
for i in range(30):
x = mods[i](x)
if i in [0, 2, 5, 7, 10, 12, 14, 17, 19, 21, 24, 26, 28]: # all the conv 2d layers
scores.append(x.clone())
return scores
# calculate cd score
f = 16
im_num = 23
im_torch, im_orig, label_num = dset.get_im_and_label(im_num) # remember torch is H x W x C
# set up blob
blob = np.zeros((3, 224, 224))
blob[:, 60:80, 150:200] = 1
blob[:, 60:120, 180:200] = 1
rel1, irrel1, scores = acd.cd_track_vgg(blob, im_torch, model)
scores = [scores[i] for i in np.arange(0, len(scores), 2)] # pick every other
# calculate build_up score
im_torch2, im_orig, label_num = dset.get_im_and_label(im_num) # remember torch is H x W x C
im_torch2[0, :, :60, :] = 0
im_torch2[0, :, :, :150] = 0
im_torch2[0, :, 80:120, :180] = 0
im_torch2[0, :, :, 200:] = 0
im_torch2[0, :, 120:, :] = 0
scores2 = vgg_track(im_torch2, model)
scores2 = [scores2[i] for i in np.arange(0, len(scores2), 2)] # pick every other
# calculate occlusion score
im_torch, im_orig, label_num = dset.get_im_and_label(im_num) # remember torch is H x W x C
im_torch[0, :, 60:80, 150:200] = 0
im_torch[0, :, 60:120, 180:200] = 0
scores3 = vgg_track(im_torch, model)
scores3 = [scores3[i] for i in np.arange(0, len(scores3), 2)] # pick every other
plt.figure(figsize=(16, 8))
num_rows = 4
num_cols = len(scores) + 1
# show original ims
# plt.subplot2grid((num_rows, num_cols), (0, 0), rowspan=num_rows)
# plt.gcf().text(0.18, 0.85, 'Blob', fontsize=14)
# plt.gcf().text(0.16, 0.15, 'Non-blob', fontsize=14)
plt.subplot(num_rows, num_cols, 1)
plt.imshow(im_orig)
blob_show = np.copy(blob[0])
blob_show[blob_show == 0] = np.nan
plt.imshow(blob_show, alpha=0.6, cmap='Greens')
plt.ylabel('CD $\\beta$', fontsize=f)
plt.xticks([])
plt.yticks([])
plt.subplot(num_rows, num_cols, num_cols + 1)
plt.imshow(im_orig, cmap='Greens')
plt.imshow(blob_show, alpha=0.6, cmap='Greens')
plt.ylabel('CD $\\gamma$', fontsize=f)
plt.xticks([])
plt.yticks([])
plt.subplot(num_rows, num_cols, num_cols * 3 + 1)
im_blob = np.copy(im_orig)
blob_idxs = blob.astype(np.int).transpose((1, 2, 0))
im_blob[blob_idxs] = 0
im_blob[60:80, 150:200] = 0
im_blob[60:120, 180:200] = 0
plt.imshow(im_blob)
plt.ylabel('Occlusion', fontsize=f)
plt.xticks([])
plt.yticks([])
plt.subplot(num_rows, num_cols, num_cols * 2 + 1)
im_blob = np.copy(im_orig)
blob_idxs = blob.astype(np.int).transpose((1, 2, 0))
# im_blob[blob_idxs] = 0
# im_blob[60:80, 150:200] = 0
# im_blob[60:120, 180:200] = 0
im_blob[:60, :] = 0
im_blob[:, :150] = 0
im_blob[80:120, :180] = 0
im_blob[:, 200:] = 0
im_blob[120:, :] = 0
plt.imshow(im_blob)
plt.ylabel('Build up', fontsize=f)
plt.xticks([])
plt.yticks([])
# show propagating images
for i in range(len(scores)):
rel1, irrel1 = scores[i]
x = np.squeeze(rel1.data.cpu().numpy())
x = np.sum(np.abs(x), axis=0)
y = np.squeeze(irrel1.data.cpu().numpy())
y = np.sum(np.abs(y), axis=0)
rel2 = scores2[i]
z = np.squeeze(rel2.data.cpu().numpy())
z = np.sum(np.abs(z), axis=0)
rel3 = scores3[i]
zz = np.squeeze(rel3.data.cpu().numpy())
zz = np.sum(np.abs(zz), axis=0)
vmax1, vmin1 = max(np.max(x), np.max(z)), min(np.min(x), np.min(z))
vmax2, vmin2 = max(np.max(y), np.max(zz)), min(np.min(y), np.min(zz))
# top row
plt.subplot(num_rows, num_cols, i + 2)
plt.imshow(x, interpolation='None', vmin=vmin1, vmax=vmax1)
plt.axis('off')
plt.title('Conv ' + str(2*i+1), fontsize=f)
# plot 2
plt.subplot(num_rows, num_cols, num_cols + i + 2)
plt.imshow(y, interpolation='None', vmin=vmin2, vmax=vmax2)
plt.axis('off')
# plot 3
plt.subplot(num_rows, num_cols, num_cols * 2 + i + 2)
plt.imshow(z, interpolation='None', cmap='viridis', vmin=vmin1, vmax=vmax1)
plt.axis('off')
# plot 4
plt.subplot(num_rows, num_cols, num_cols * 3 + i + 2)
plt.imshow(zz, interpolation='None', cmap='viridis', vmin=vmin2, vmax=vmax2)
plt.axis('off')
plt.subplots_adjust(hspace=0, wspace=0)
plt.show()
# -
# # fig 3 - recreate hockey example
# +
# hyperparameters
num_iters = 5 # number of iterations to agglomerate before merging remaning blobs (fig uses 4)
percentile_include = 95 # values above this percentile will be added at each iteration (fig uses 95)
method = 'cd' # method to rank importance ('cd' works best, 'build_up' or 'occlusion' are simplest)
sweep_dim = 14 # importances are calculated by blocks of sweep_dim x sweep_dim (14 yields good results for imagenet)
im_torch, im_orig, lab_num_correct = imnet_dict[9] # the hockey example
lab_pred = np.argmax(dset.pred_ims(model, np.copy(im_orig)))
lists = acd.agg_2d.agglomerate(model, dset.pred_ims, percentile_include, method, sweep_dim, im_orig,
lab_pred, num_iters=num_iters, im_torch=im_torch, model_type=model_type)
# visualize
plt.figure(figsize=(12, 5), facecolor='white', dpi=100)
rows = 3
num_ims = len(lists['scores_list'])
# original plots
ind, labs = viz.visualize_original_preds(im_orig, lab_num_correct,
lists['comp_scores_raw_list'], lists['scores_orig_raw'],
subplot_rows=rows, dset=dset)
# comp plots
viz.visualize_ims_list(lists['comps_list'],
title='Chosen blobs',
subplot_row=1, subplot_rows=rows, colorbar=False, im_orig=im_orig, plot_overlay=True)
# dict plots
viz.visualize_dict_list_top(lists['comp_scores_raw_list'], method,
subplot_row=2, subplot_rows=rows, ind=ind, labs=labs, use_orig_top=True)
# -
# # fig s1 - compare different scores
# +
def get_diff_scores(im_torch, im_orig, label_num, model, preds, sweep_dim):
scores = []
# cd
method = 'cd'
tiles = acd.tiling_2d.gen_tiles(im_orig, fill=0, method=method, sweep_dim=sweep_dim)
scores_cd = acd.get_scores_2d(model, method=method, ims=tiles,
im_torch=im_torch, model_type=model_type, device=device)
scores.append(scores_cd)
for method in ['occlusion', 'build_up']: # 'build_up'
tiles_break = acd.tiling_2d.gen_tiles(im_orig, fill=0, method=method, sweep_dim=sweep_dim)
preds_break = acd.get_scores_2d(model, method=method, ims=tiles_break,
im_torch=im_torch, pred_ims=dset.pred_ims)
if method == 'occlusion':
preds_break += preds
scores.append(np.copy(preds_break))
# get integrated gradients scores
scores.append(acd.ig_scores_2d(model, im_torch, num_classes=1000,
im_size=224, sweep_dim=sweep_dim, ind=[label_num], device=device))
return scores
# pick an image + get scores
im_nums = [34, 20, 36, 32] # 34: screen, 20: snake, 36: trash can, 32: crane
sweep_dim = 14
# sweep_dim = 56
fig = plt.figure(figsize=(10, 8), facecolor='white')
for x, im_num in enumerate(im_nums):
im_torch, im_orig, label_num = imnet_dict[im_num] # remember torch is H x W x C
print('lab', dset.lab_dict[label_num])
# viz.visualize_ims_tiled(tiling.gen_tiles(im_orig, fill=np.nan))
preds = dset.pred_ims(model, im_orig).flatten()
ind = np.argpartition(preds, -8)[-8:] # top-scoring indexes
ind = ind[np.argsort(preds[ind])][::-1] # sort the indexes
scores = get_diff_scores(im_torch, im_orig, label_num, model, preds, sweep_dim)
# plot raw image
num_rows = len(im_nums)
num_cols = len(scores) + 1
plt.subplot(num_rows, num_cols, 1 + x * num_cols)
plt.imshow(im_orig)
# plt.axis('off')
plt.gca().xaxis.set_visible(False)
plt.yticks([])
if x == 0:
plt.title('Image', fontsize=16)
if x == 0:
plt.ylabel('CRT screen', fontsize=15)
elif x == 1:
plt.ylabel('Green mamba', fontsize=15)
elif x == 2:
plt.ylabel('Trash can', fontsize=15)
elif x == 3:
plt.ylabel('Crane', fontsize=15)
# plot scores
vmax = max([np.max(scores[i]) for i in range(len(scores))])
vmin = min([np.min(scores[i]) for i in range(len(scores))])
vabs = max(abs(vmax), abs(vmin))
for i, tit in enumerate(['CD', 'Occlusion', 'Build-Up', 'IG']):
plt.subplot(num_rows, num_cols, 2 + i + x * num_cols)
if i == 0:
plt.ylabel('pred: ' + dset.lab_dict[ind[0]][:16] + '...', fontsize=15)
if x == 0:
plt.title(tit, fontsize=16)
p = viz.visualize_preds(scores[i], num=label_num, cbar=False) #axis_off=False, vabs=vabs)
plt.xticks([])
plt.yticks([])
# divider = make_axes_locatable(plt.gca())
# cax = divider.append_axes("right", size="2%", pad=0.05)
# plt.colorbar(p, cax=cax)
plt.tight_layout()
plt.subplots_adjust(wspace=0, hspace=0)
plt.show()
# -
|
reproduce_figs/imagenet_fig3,s1,s2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import datetime as dt
music_0603 = pd.read_csv('music_data.csv')
music_0603
date = dt.date(2021, 6, 3)
music_0603['Date'] = date
music_clean_0603 = music_0603.drop(labels=[0,3], axis=0)
music_clean_0603
music_0609 = pd.read_csv('music_data0609.csv')
music_0609
date1 = dt.date(2021, 6, 9)
music_0609['Date'] = date1
music_0609
tiktok_growth = pd.DataFrame.append(music_clean_0603, music_0609).drop(columns='Unnamed: 0')
tiktok_growth
tiktok_growth.to_csv('tiktok_growth.csv')
|
notebooks/music_growth.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys # for automation and parallelisation
manual, scenario = (True, 'base') if 'ipykernel' in sys.argv[0] else (False, sys.argv[1])
if manual:
# %matplotlib inline
import numpy as np
import pandas as pd
import geopandas as gpd
from shapely import geometry
from quetzal.model import stepmodel
from quetzal.engine import engine, connectivity
from quetzal.io import excel
# # Preparation of the transport network.
# ## Saves access and egress links for road network
# ## Needs all networks.
#
input_path = '../input_static/'
output_path = '../output/'
model_path = '../model/'
# Load scenario parameters
params = excel.read_var(file='../input/parameters.xls', scenario=scenario)
sm = stepmodel.read_json(model_path + 'de_pt_network_agg')
road = stepmodel.read_json(input_path + 'de_road_network')
# Walking vs. driving to the train station
threshold = 500 # in m
speed_non_motorised_modes = 17 # in km/h
speed_car = 50 # in km/h
speed_footpaths = 5 # in km/h, assumed
# Max. distances for accessing the transport nodes
# in m
threshold_dict = {'road': 10000,
'road_pt': 500}
# ## Add access and egress links from zone centroids
# Compute controids
sm.preparation_ntlegs(
zone_to_transit=False,
zone_to_road=False)
# Compute road access and egress links
sm.zone_to_road = engine.ntlegs_from_centroids_and_nodes(
sm.centroids,
road.road_nodes,
short_leg_speed=params['pt_access']['speed_walk'],
long_leg_speed=params['pt_access']['speed_bicycle'],
threshold=params['pt_access']['catchment_radius_walk'],
n_neighbors=params['access-egress_links']['n_road'],
coordinates_unit=sm.coordinates_unit)
sm.zone_to_road = sm.zone_to_road.loc[
sm.zone_to_road['distance']<=params['access-egress_links']['road_max_dist']]
len(sm.zone_to_road.index)
# Compute road - PT links
sm.road_to_transit = engine.ntlegs_from_centroids_and_nodes(
sm.nodes,
road.road_nodes,
short_leg_speed=params['pt_access']['speed_walk'],
long_leg_speed=params['pt_access']['speed_bicycle'],
threshold=params['pt_access']['catchment_radius_walk'],
n_neighbors=params['access-egress_links']['n_road_pt'],
coordinates_unit=sm.coordinates_unit)
sm.zone_to_road = sm.zone_to_road.loc[
sm.zone_to_road['distance']<=params['access-egress_links']['road_pt_max_dist']]
len(sm.road_to_transit.index)
# ### Parametrise access and egress links
# Road - PT connectors
sm.road_to_transit['distance'] = 0
sm.road_to_transit['time'] = params['access-egress_links']['time_road_pt']
sm.road_to_transit.sample(n=2)
# Road - centroid connectors
sm.zone_to_road['distance'] = 0
sm.zone_to_road['time'] = params['access-egress_links']['time_road']
sm.zone_to_road.sample(n=2)
# ## Save model
# Drop unneccessary columns
cols = ['speed_factor', 'short_leg_speed', 'long_leg_speed', 'rank']
sm.zone_to_road.drop(cols, axis=1, inplace=True, errors='ignore')
sm.road_to_transit.drop(cols, axis=1, inplace=True, errors='ignore')
# Make tables lighter
cols = ['distance', 'speed', 'time']
sm.zone_to_road[cols] = sm.zone_to_road[cols].astype(int)
sm.road_to_transit[cols] = sm.road_to_transit[cols].astype(int)
sm.to_json(model_path + 'de_road_access_egress',
only_attributes=['centroids', 'zone_to_road', 'road_to_transit'],
encoding='utf-8')
|
notebooks/prep21_access_egress_road.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('MSCS-basic')
# language: python
# name: python3
# ---
# +
def fast_exp(base, exponent, modulus):
result = 1
while exponent != 0:
if exponent % 2 == 1:
result = (result * base) % modulus
base = base**2 % modulus
exponent = exponent // 2
return result
def extended_euclidean(m, n):
if n == 0:
return 1, 0, m
x, y, g = extended_euclidean(n, m % n)
return y, x - (m // n)*y, g
def inv_mod(a, modulus):
s, _, g = extended_euclidean(a, modulus)
assert g == 1, ValueError('the modular inverse does not exist')
return s % modulus
def crt(remainders, moduli):
assert len(remainders) == len(moduli), ValueError('the lists remainders and moduli are not the same length')
assert len(remainders) > 0, ValueError('the list lengths must be greater than zero')
first_modulus = moduli[0]
first_remainder = remainders[0]
if len(remainders) == 1:
return first_remainder % first_modulus, first_modulus
consecutive_remainder, consecutive_modulus = crt(remainders[1:], moduli[1:])
u, v, g = extended_euclidean(consecutive_modulus, first_modulus)
assert g == 1, ValueError('the moduli are not relatively prime')
return (first_remainder*consecutive_modulus*u + consecutive_remainder*first_modulus*v) % (first_modulus*consecutive_modulus), first_modulus*consecutive_modulus
def eulers_totient(p, q):
return (p-1)*(q-1)
def polynomial_congruence(e, m_to_the_e, totient_n, n):
d = inv_mod(e, totient_n)
m = fast_exp(m_to_the_e, d, n)
return m, d
def rsa_decrypt(e, m_to_the_e, p, q):
return polynomial_congruence(e, m_to_the_e, eulers_totient(p, q), p*q)
# -
# ### 1. Using the code developed in class, solve the following Chinese Remainder Theorem problem.
#
# $\begin{align*}
# x &\equiv 197 (\operatorname{mod} 1009) \\
# x &\equiv 917 (\operatorname{mod} 1013) \\
# x &\equiv 439 (\operatorname{mod} 1559) \\
# x &\equiv 777 (\operatorname{mod} 1439)
# \end{align*}$
# +
remainders = [197, 917, 439, 777]
moduli = [1009, 1013, 1559, 1439]
remainder, modulus = crt(remainders, moduli)
print(f'{remainder} (mod{modulus})')
# -
# ### 2. Using the code developed in class, compute the following exponentiation.
#
# $189723981723918723789^{8978234758972347892342789}(\operatorname{mod}999999999991)$
fast_exp(189723981723918723789, 8978234758972347892342789, 999999999991)
# ### 3. Use Fermat's Little Theorem to compute the following exponentiations. (This should be done by hand, make sure to show all work).
#
# $\begin{align*}
# &5^{117} (\operatorname{mod} 7) \\
# &7^{897123789} (\operatorname{mod} 11) \\
# &11^{200} (\operatorname{mod} 23)
# \end{align*}$
# $5^{117} = 5^{19*6 + 3} \approx 5^3 = 125 \approx 6 (\operatorname{mod} 7)$
# $7^{897123789} = 7^{89712378*10 + 9} \approx 7^{9} \approx 8 (\operatorname{mod} 11)$
#
# $7^{1} \approx 7$, $7^{2} \approx 5$, $7^{4} \approx 5^{2} \approx 3$, $7^{8} \approx 3^{2} \approx 9 (\operatorname{mod} 11)$
#
# $7^9 = 7^8*7 \approx 8 (\operatorname{mod} 11)$
# $11^{200} = 11^{9*22 + 2} \approx 11^{2} \approx 6 (\operatorname{mod} 23)$
# ### 4. Compute Euler's Totient Function for the following numbers.
#
# $\begin{align*}
# &\phi(2^{5}7^{3}) \\
# &\phi(11*23^{3}) \\
# &\phi(196)
# \end{align*}$
# $\phi(2^{5}7^{3}) = \phi(2^5)\phi(7^3) = (2^5-2^4)(7^3 - 7^2) = 4704$
# $\phi(11*23^3) = \phi(11)*\phi(23^3) = (11-1)(23^3-23^2) = 116380$
# $\phi(196) = \phi(2^{2}7^{2}) = \phi(2^2)\phi(7^2) = (2^2-2)(7^2-7) = 84$
# ### 5. The numbers $2459$ and $2503$ are prime with product $6154877$. Solve the following congruences for $x$.
#
#
# $\begin{align*}
# &x^{101} \approx 420539 (\operatorname{mod} 6154877) \\
# &x^{2393} \approx 4597153 (\operatorname{mod} 6154877) \\
# &x^{1373} \approx 2487125 (\operatorname{mod} 6154877)
# \end{align*}$
m, d = rsa_decrypt(101, 420539, 2459, 2503)
print(f'{m=}\n{d=}')
m, d = rsa_decrypt(2393, 4597153, 2459, 2503)
print(f'{m=}\n{d=}')
m, d = rsa_decrypt(1373, 2487125, 2459, 2503)
print(f'{m=}\n{d=}')
|
Homework/Homework 3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Power Spectrum Estimation
#
# This notebook is to understand the details of the power spectrum, its computation with FFT, carefully considering scaling issues, units and interpretation.
#
# Important lessons to be learned (non-windowed case)
#
# - Normalizing the FFT by sqrt(N)
# - squared magnitudes: __Energy Spectrum [V^2 s]__ - grows w/ N
# - Normalizing the FFT by N:
# - magnitudes are __RMS amplitudes [V]__ (for the given frequency bin)
# - squared magnitudes: __Power Spectrum [V^2]__
# - squared magnitudes normalized by the width of the bin: __Power Spectral Density [V^2/Hz]__
#
# __Power spectral density better suits wide-band (i.e. noise) signals. Power spectrum is better for interpreting narrow-band (i.e. single frequency) signals.__
#
# __Alternative view on DFT__: By looking at the definition of DFT, it can be interpreted as a mixer (complex exponential multipler) and a low-pass filer (box-car or simple average). The low-pass filter (hence the DFT bins) will gets narrower as you increase N.
#
# TODO: understand why we need to scale bins /2 (except at DC) - Hint: this is needed only for real (non-complex) signals
# Create a discrete sinusoid signal with some added noise. We assume that this is a voltage signal.
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
# constants
FS = 1e4 # sampling rate (Hz)
SIG_F = 1e3 # signal frequency (Hz)
SIG_DB = 0 # signal amplitude (dB)
NOISE_DB = -15 # noise amplitude (dB)
T = 1 # signal length (s)
dT = 1 / FS
t = np.arange(0, T, 1/FS)
sig = np.sin(2 * np.pi * SIG_F * t) * (10 ** (SIG_DB / 20))
noise = np.random.randn(sig.size) * (10 ** (NOISE_DB / 20))
samples = sig + noise
plt.plot(t[:100], samples[:100])
plt.xlabel('Time (s)')
plt.ylabel('Amplitude (V)')
# -
# Calculate the average power of the clean signal and of the noise from the time domain samples. Compute SNR.
#
# __Note__: the power of a sinusoid with unit amplitude is -3dB.
# +
P_sig_t = np.mean(sig ** 2) # same as np.sum((sig ** 2) * dT) / T
P_noise_t = np.mean(noise ** 2)
SNR_t = 10 * np.log10(P_sig_t / P_noise_t)
print('P(sig)= %.2f V^2, P(noise)= %.2f V^2, SNR= %.2f dB' % (P_sig_t, P_noise_t, SNR_t))
print('RMS(sig)= %.2f V, RMS(noise)= %.2f V' % (np.sqrt(P_sig_t), np.sqrt(P_noise_t)))
# -
# ## Power Spectrum
#
# Compute the DFT of the time domain samples using a fixed length (N).
#
# __Note__: the DFT results have to be scaled by 1/sqrt(N) to conserve energy (unitary operator). You can achieve the same results with `np.fft.fft(samples, norm='ortho')`. Also, see Parseval's Theorem.
# +
N = 1000 # must be even for these computations
X = np.fft.fft(samples, n=N) / np.sqrt(N)
f = np.fft.fftfreq(N, dT)
# Verify if time and frequency domain energies are the same
np.sum(np.abs(X) ** 2), np.sum(samples[:N] ** 2)
# -
# First important observation: the squared magnitude of the FFT values represent the __energy__ distribution across the frequency bins for the given signal length (N). Thus, the absolute bin values depend on N.
# +
Exx = np.abs(X) ** 2
plt.semilogy(np.fft.fftshift(f), np.fft.fftshift(Exx))
plt.title('Energy Spectrum')
plt.xlabel('Frequency (Hz)')
plt.ylabel('Energy in bin ($V^2s$)')
# -
# Let's convert the FFT values to power. In the time domain, we divided the sum energy by N. This is what we do in the frequency domain, too to get average power in each freq bin. If you followed carefully, we normalized the the FFT squared magnitudes by N to get energy and again by N to get power. __This is why people prefer to normalize the FFT values by N (so the squared magnitudes are in the power units)__.
Pxx = Exx / N
plt.semilogy(np.fft.fftshift(f), np.fft.fftshift(Exx))
plt.title('Power Spectrum')
plt.xlabel('Frequency (Hz)')
plt.ylabel('Power in bin ($V^2$)')
# Due to the real time-domain samples we have a symmetric spectrum (complex conjugate). Let's take and scale the positive half of it.
# +
Pxx = Pxx[:N // 2]
Pxx[1:-2] *= 2 # conserve energy
f = f[:N // 2]
plt.semilogy(f, Pxx)
plt.title('Power Spectrum')
plt.xlabel('Frequency (Hz)')
plt.ylabel('Power in bin ($V^2$)')
plt.ylim(1e-6, 1)
plt.grid()
# -
# Let's compare the result with the built-in periodogram function.
f2, Pxx2 = signal.periodogram(samples, FS, nfft=N, scaling='spectrum')
plt.semilogy(f2, Pxx2)
plt.title('Power Spectrum using scipy.signal.periodogram')
plt.xlabel('Frequency (Hz)')
plt.ylabel('Power in bin ($V^2$)')
plt.ylim(1e-6, 1)
plt.grid()
plt.show()
# Calculate SNR using the frequency domain (first peak is the signal assumption).
# +
f_sig_idx = np.argmax(Pxx)
SNR_f = 10 * np.log10(Pxx[f_sig_idx] / np.sum(np.delete(Pxx, f_sig_idx)))
print('SNR= %.2f dB (time domain SNR= %.2f dB)' % (SNR_f, SNR_t))
# -
# ## Power Spectrum Density
#
# Instead of ploting the (average) power in each frequency bin we can compute/plot the power density. This is a scaling of the power spectrum results by the width of the bin (in Hz). We also compare this to the built-in periodogram with density scaling.
# +
plt.semilogy(f, Pxx / (FS / N))
plt.title('PSD computed from DFT')
plt.xlabel('Frequency (Hz)')
plt.ylabel('PSD ($V^2/Hz$)')
plt.ylim(1e-7, 1)
plt.grid()
plt.show()
f2, Pxx2 = signal.periodogram(samples, FS, nfft=N, scaling='density')
plt.semilogy(f2, Pxx2)
plt.title('PSD using scipy.signal.periodogram')
plt.xlabel('Frequency (Hz)')
plt.ylabel('PSD ($V^2/Hz$)')
plt.ylim(1e-7, 1)
plt.grid()
# -
# __Observation__: the PSD figure is better for showing the noise level (it's height does not change with N), but is hard to interpret for the signal (it's height changes). The 'spectrum' scaling is better for the signal (does not change with N) but misleading for the noise level.
# +
f3, Pxx3 = signal.periodogram(samples, FS, nfft=512, scaling='density')
plt.semilogy(f3, Pxx3)
plt.title('PSD with N=1024')
plt.xlabel('Frequency (Hz)')
plt.ylabel('PSD ($V^2/Hz$)')
plt.ylim(1e-7, 1)
plt.grid()
plt.show()
f3, Pxx3 = signal.periodogram(samples, FS, nfft=8192, scaling='density')
plt.semilogy(f3, Pxx3)
plt.title('PSD with N=1024')
plt.xlabel('Frequency (Hz)')
plt.ylabel('PSD ($V^2/Hz$)')
plt.ylim(1e-7, 1)
plt.grid()
plt.show()
f3, Pxx3 = signal.periodogram(samples, FS, nfft=512, scaling='spectrum')
plt.semilogy(f3, Pxx3)
plt.title('Power Spectrum with N=1024')
plt.xlabel('Frequency (Hz)')
plt.ylabel('PSD ($V^2/Hz$)')
plt.ylim(1e-7, 1)
plt.grid()
plt.show()
f3, Pxx3 = signal.periodogram(samples, FS, nfft=8192, scaling='spectrum')
plt.semilogy(f3, Pxx3)
plt.title('Power Spectrum with N=1024')
plt.xlabel('Frequency (Hz)')
plt.ylabel('PSD ($V^2/Hz$)')
plt.ylim(1e-7, 1)
plt.grid()
plt.show()
# -
# ## TODO: Windowing
|
PSD.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 定义卷积神经网络(CNN)
#
# 查看正在使用的数据之后,了解图像与关键点的形状,接下来,就可以定义一个机器人可以从这些数据中 *学习*的卷积神经网络。
#
# 在这个notebook和`models.py`中,你的任务是:
# 1. 定义一个CNN,把图像作为输入,把关键点作为输出
# 2. 与以前一样,构造转换后的FaceKeypointsDataset
# 3. 使用训练数据训练这个CNN,并跟踪损失
# 4. 查看训练模型对测试数据的执行情况
# 5. 如有必要,请修改CNN结构并模拟超参数,使其*表现良好* **\***
#
# **\*** 什么是*表现良好*?
#
# “表现良好”意味着该模型的损失在训练期间有所降低,**而且**该模型应用于测试图像数据时,会产生与每个人脸的真实关键点紧密匹配的关键点。你会在这个notebook中看到这个例子。
#
# ---
#
#
# ## CNN架构
#
# 回想一下,CNN是由下列几种类型的层定义的:
# * 卷积层
# * 最大池化层
# * 全连接层
#
# 你需要使用上述层,而且我们建议你添加多个卷积层以及可能防止过度拟合的dropout层等。此外,你还可以查看一些有关关键点检测的文献,如 [这篇论文](https://arxiv.org/pdf/1710.00977.pdf),帮助你确定该网络的结构。
#
#
# ### TODO: 在`models.py`文件中定义你的模型
#
# 此文件大部分为空,但其中包含预期的名称和一些用于创建模型的TODO事项。
#
# ---
#
# ## PyTorch神经网络
#
# 要在PyTorch中定义神经网络,你可以在函数`__init__`中定义一个模型的各个层,并定义一个网络的前馈行为,该网络会在函数`forward`中使用这些初始化的层,而该函数会接收输入图像张量`x`。此Net类的结构如下所示,并由你来填充。
#
# 注意:在训练期间,PyTorch能够通过跟踪网络的前馈行为并使用autograd来计算该网络中权重的更新来执行反向传播。
#
# #### 在` __init__`中定义层
# 提醒一下,卷积层与池化层可以像这样来定义(在`__init__`中):
# +
# 1 input image channel (for grayscale images), 32 output channels/feature maps, 3x3 square convolution kernel
self.conv1 = nn.Conv2d(1, 32, 3)
# maxpool that uses a square window of kernel_size=2, stride=2
self.pool = nn.MaxPool2d(2, 2)
# -
# #### 引用`forward`中的层
# 然后在这样的`forward`函数中引用,其中卷积1层在应用最大池化之前应用了ReLu激活函数:
x = self.pool(F.relu(self.conv1(x)))
# 最佳做法是把权重将在训练过程中发生变化的任何层防治在`__init__`中,并在`forward`函数中引用它们。所有始终以相同方式运行的层或函数(例如预定义的激活函数)应*只* 出现在`forward` 函数中。
#
# #### 为什么要用models.py文件
#
# 你的任务是在`models.py`文件中定义该网络,便于在此项目目录中的不同notebook中按名称保存和加载你定义的任何模型。例如,通过在`models.py`中定义名为`Net`的CNN类,通过简单地导入该类并实例化模型,就可以在此notebook和其他notebook中创建相同的体系结构:
from models import Net
net = Net()
# load the data if you need to; if you have already loaded the data, you may comment this cell out
# -- DO NOT CHANGE THIS CELL -- #
# !mkdir /data
# !wget -P /data/ https://s3.amazonaws.com/video.udacity-data.com/topher/2018/May/5aea1b91_train-test-data/train-test-data.zip
# !unzip -n /data/train-test-data.zip -d /data
# <div class="alert alert-info">**注意:**工作区会在持续30分钟的不活动状态后,自动关闭连接,包括训练时出现不活动状态。使用下面的代码段可以在训练期间保持工作区的活动状态。下面导入了active_session上下文管理器。
# </div>
# +
from workspace_utils import active_session
with active_session():
train_model(num_epochs)
# +
# import the usual resources
import matplotlib.pyplot as plt
import numpy as np
# import utilities to keep workspaces alive during model training
from workspace_utils import active_session
# watch for any changes in model.py, if it changes, re-load it automatically
# %load_ext autoreload
# %autoreload 2
# +
## TODO: Define the Net in models.py
import torch
import torch.nn as nn
import torch.nn.functional as F
## TODO: Once you've define the network, you can instantiate it
# one example conv layer has been provided for you
from models import Net
net = Net()
print(net)
# -
# ## 转换数据集
#
# 为训练做准备,你还需要创建一个图像和关键点的转换数据集。
#
# ### TODO: 定义一个数据转换
#
# 在PyTorch中,卷积神经网络需要一个大小一致的torch图像作为输入。为了进行有效的训练,以及在训练过程中该模型的损失不会放大,我们还建议你对输入图像和关键点进行归一化。必要的转换已在`data_load.py`中定义,你无需再做修改。另外,你可以看一下这个文件,你会在该文件中看到Notebook 1中定义和应用的相同转换。
#
# 要定义下面的数据转换,请使用以下[组合](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html#compose-transforms) :
# 1. 重新缩放和/或裁剪数据,最终需要一个方形图像(建议大小为224x224px)
# 2. 归一化图像和关键点;将每个RGB图像转换为颜色范围为[0,1]的灰度图像,并将给定关键点转换为[-1,1]的范围
# 3. 将这些图像和关键点转换为张量
#
# 这些转换已在`data_load.py`中定义,但是否要在下面调用它们并创建一个`data_transform`,这都取决于你。**该转换将应用于训练数据,以及稍后的测试数据**。这样将改变显示这些图像和关键点的方式,但这些步骤对于高效训练来说非常重要。
#
# 需要说明的一点是,如果你想要执行数据增强(在此项目中是可选的),并随机旋转或移动这些图像,方形图像大小将会很有用,将224x224图像旋转90度就会产生相同的输出形状。
# +
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# the dataset we created in Notebook 1 is copied in the helper file `data_load.py`
from data_load import FacialKeypointsDataset
# the transforms we defined in Notebook 1 are in the helper file `data_load.py`
from data_load import Rescale, RandomCrop, Normalize, ToTensor
## TODO: define the data_transform using transforms.Compose([all tx's, . , .])
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(250),
RandomCrop(224),
Normalize(),
ToTensor()])
# testing that you've defined a transform
assert(data_transform is not None), 'Define a data_transform'
# +
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='/data/training_frames_keypoints.csv',
root_dir='/data/training/',
transform=data_transform)
print('Number of images: ', len(transformed_dataset))
# iterate through the transformed dataset and print some stats about the first few samples
for i in range(4):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
# -
# ## 批处理并加载数据
#
# 定义了转换数据集之后,接下来,我们可以使用PyTorch的DataLoader类来批量加载任意大小的训练数据,也可以对训练模型的数据进行置乱处理。你可以在 [本文档](http://pytorch.org/docs/master/data.html)中阅读有关DataLoader参数的更多信息。
#
# #### 批量大小
# 确定用于训练模型的最合适的批量是多少。小批量与大批量都要试一试,并注意在模型训练时损失会如何减少。批量过大可能会导致模型在训练时崩溃和/或内存不足。
#
# **Windows用户需要注意:**请将`num_workers`改为0,否则可能会遇到DataLoader失效的问题。
# +
# load training data in batches
batch_size = 32
train_loader = DataLoader(transformed_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=0)
# -
# ## 训练之前
#
# 看一下这个模型在训练之前的表现。你应该会看到,它预测的关键点从一个点开始,并且与人脸上的关键点根本不匹配!你可以把此行为可视化,并在训练后将其与模型进行比较,还可以查看该模型是如何改进的。
#
# #### 加载测试数据集
#
# 此模型之前*没有*见过这个测试数据集,这就是说,它没有使用这些图像进行过训练。在这里,我们将加载此测试数据,并在训练前后,查看你的模型在此数据集上的表现效果如何!
#
# 为了可视化这些测试数据,我们必须要做一些非转换步骤,将图像转换为张量的python图像,并将关键点重新转换回可识别的范围。
# +
# load in the test data, using the dataset class
# AND apply the data_transform you defined above
# create the test dataset
test_dataset = FacialKeypointsDataset(csv_file='/data/test_frames_keypoints.csv',
root_dir='/data/test/',
transform=data_transform)
# +
# load test data in batches
batch_size = 32
test_loader = DataLoader(test_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=0)
# -
# ## 将模型应用于测试样本
#
# 要在测试数据样本上测试模型,你必须执行以下步骤:
# 1. 从样本中提取图像和实际真值关键点
# 2. 将图像隐藏在变量中,便于你的网络将其作为输入处理,并跟踪图像在该网络中移动时发生的变化。
# 3. 确保图像是模型所需的FloatTensor。
# 4. 通过网络向前传递图像,获得预测的输出关键点。
#
# 此函数测试的是该网络在第一批测试数据上的执行情况。它会返回图像、转换图像、预测由模型产生的关键点以及实际真值关键点。
# +
# test the model on a batch of test images
def net_sample_output():
# iterate through the test dataset
for i, sample in enumerate(test_loader):
# get sample data: images and ground truth keypoints
images = sample['image']
key_pts = sample['keypoints']
# convert images to FloatTensors
images = images.type(torch.FloatTensor)
# forward pass to get net output
output_pts = net(images)
# reshape to batch_size x 68 x 2 pts
output_pts = output_pts.view(output_pts.size()[0], 68, -1)
# break after first image is tested
if i == 0:
return images, output_pts, key_pts
# -
# #### 调试技巧
#
# 如果此处出现尺寸或维度错误,请确保你的网络输出预期数量的关键点!或者,如果收到Tensor类型的错误,请考虑将数据转换为float类型的上述代码进行更改,float类型为:`images = images.type(torch.FloatTensor)`。
# +
# call the above function
# returns: test images, test predicted keypoints, test ground truth keypoints
test_images, test_outputs, gt_pts = net_sample_output()
# print out the dimensions of the data to see if they make sense
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
# -
# ## 将预测的关键点可视化
#
# 让模型生成一些预测的输出关键点之后,就可以用一种类似于我们之前显示这些数据的方式来显示这些点,只是这一次,要显示这些点,我们必须“取消转换”图像/关键点数据。
#
# 请注意,我已经定义了一个*新*函数`show_all_keypoints`,它会显示灰度图像、其预测的关键点以及其实际真值关键点(如果提供的话)。
def show_all_keypoints(image, predicted_key_pts, gt_pts=None):
"""Show image with predicted keypoints"""
# image is grayscale
plt.imshow(image, cmap='gray')
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
# plot ground truth points as green pts
if gt_pts is not None:
plt.scatter(gt_pts[:, 0], gt_pts[:, 1], s=20, marker='.', c='g')
# #### 非转换
#
# 接下来,你会看到一个辅助函数,即`visualize_output`,它会接收一批图像、预测关键点以及实际真值关键点,并显示一组图像及其真实/预测关键点。
#
# 此函数的主要作用是获取批量图像和关键点数据(CNN的输入和输出),并将它们转换为numpy图像和非归一化关键点(x,y),从而进行正常显示。非转换过程将关键点和图像转换为来自Tensors的numpy数组,*此外*, 它撤消了Normalize()转换中完成的关键点归一化。但前提是我们假设,你在载测试数据时应用了这些转换。
# +
# visualize the output
# by default this shows a batch of 10 images
def visualize_output(test_images, test_outputs, gt_pts=None, batch_size=10):
for i in range(batch_size):
plt.figure(figsize=(20,10))
ax = plt.subplot(1, batch_size, i+1)
# un-transform the image data
image = test_images[i].data # get the image from it's Variable wrapper
image = image.numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
# un-transform the predicted key_pts data
predicted_key_pts = test_outputs[i].data
predicted_key_pts = predicted_key_pts.numpy()
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50.0+100
# plot ground truth points for comparison, if they exist
ground_truth_pts = None
if gt_pts is not None:
ground_truth_pts = gt_pts[i]
ground_truth_pts = ground_truth_pts*50.0+100
# call show_all_keypoints
show_all_keypoints(np.squeeze(image), predicted_key_pts, ground_truth_pts)
plt.axis('off')
plt.show()
# call it
visualize_output(test_images, test_outputs, gt_pts)
# -
# ## 训练
#
# #### 损失函数
# 训练一个用于预测关键点的网络与训练一个用于预测类的网络不同。你可能希望选择适合回归的损失函数,而不是输出类的分布并使用交交叉熵损失函数,因为损失函数可以用于直接比较预测值和目标值。有关各种损失函数(如MSE或L1 / SmoothL1损失),请阅读 [本文档](http://pytorch.org/docs/master/_modules/torch/nn/modules/loss.html)中的内容。
#
# ### TODO: 定义损失与优化
#
# 接下来,你需要通过定义损失函数和优化程序来定义模型的训练方式。
#
# ---
# +
## TODO: Define the loss and optimization
import torch.optim as optim
criterion = nn.L1Loss()
# criterion = nn.SmoothL1Loss()
# criterion = nn.MSELoss()
optimizer = optim.Adam(net.parameters(), lr = 0.001)
# -
# ## 训练与初步观察
#
# 现在,你要使用大量epoch,从`train_loader`中训练你的批量训练数据。
#
# 为了快速观察你的模型是如何训练并决定是否应该修改它的结构或超参数,我们建议你最开始的时候使用一个或两个epoch。训练时,请注意观察模型的损失会如何随着时间的推移而变化:例如,它会先快速减少然后再减慢吗?或者起初会在一段时间后出现减少?如果更改了训练数据的批量大小或修改损失函数,会发生什么变化?
#
# 在使用多个epoch进行训练并创建最终模型之前,使用这些初始观察值对模型进行更改并确定一个最佳架构。
def train_net(n_epochs):
# prepare the net for training
net.train()
for epoch in range(n_epochs): # loop over the dataset multiple times
running_loss = 0.0
# train on batches of data, assumes you already have train_loader
for batch_i, data in enumerate(train_loader):
# get the input images and their corresponding labels
images = data['image']
key_pts = data['keypoints']
# flatten pts
key_pts = key_pts.view(key_pts.size(0), -1)
# convert variables to floats for regression loss
key_pts = key_pts.type(torch.FloatTensor)
images = images.type(torch.FloatTensor)
# forward pass to get outputs
output_pts = net(images)
# calculate the loss between predicted and target keypoints
loss = criterion(output_pts, key_pts)
# zero the parameter (weight) gradients
optimizer.zero_grad()
# backward pass to calculate the weight gradients
loss.backward()
# update the weights
optimizer.step()
# print loss statistics
running_loss += loss.item()
if batch_i % 50 == 49: # print every 10 batches
print('Epoch: {}, Batch: {}, Avg. Loss: {}'.format(epoch + 1, batch_i+1, running_loss/10))
running_loss = 0.0
print('Finished Training')
# +
# train your network
n_epochs = 35 # start small, and increase when you've decided on your model structure and hyperparams
# this is a Workspaces-specific context manager to keep the connection
# alive while training your model, not part of pytorch
with active_session():
train_net(n_epochs)
# -
# ## 测试数据
#
# 了解你的模型在之前未见过的测试数据上的表现如何。我们已经对测试数据进行加载与转换,这一点类似于与训练数据时的做法类似。接下来,在这些图像上运行已被训练的模型,查看其生成的关键点类型。你应该能够观察到你的模型是否拟合了它看到的每个新人脸,这些点是否是随机分布的,以及这些点实际上是否过度拟合了训练数据而没有进行归纳。
# +
# get a sample of test data again
test_images, test_outputs, gt_pts = net_sample_output()
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
# +
## TODO: visualize your test output
# you can use the same function as before, by un-commenting the line below:
visualize_output(test_images, test_outputs, gt_pts)
# -
# 找到了一个或两个表现良好的模型后,保存你的模型,这样你就可以加载它并在以后使用它了!
#
# 在这里,你需要保存模型,但请**在提交项目之前删除任何检查点和已保存的模型**,否则你的工作区可能会因为太大而无法提交。
# +
## TODO: change the name to something uniqe for each new model
model_dir = 'saved_models/'
model_name = 'keypoints_model_1.pt'
# after training, save your model parameters in the dir 'saved_models'
torch.save(net.state_dict(), model_dir+model_name)
# -
# 完成对一个表现良好的模型的训练后,请回答以下问题,以便我们对你的训练和架构选择过程有一些了解。要通过此项目,你需要回答下列所有问题。
#
# ### 问题1:你选择了哪些优化和损失函数?为什么会这样选择?
#
#
# **答案**: 优化器选择了Adam,因为它的学习速率是自适应的;损失函数选择了L1Loss,MSELoss对大的误差值最敏感,但在这里并没有特别的优势,所以最后选择了简洁直观的L1Loss
#
#
# ### 问题2:最开始,你的网络架构是什么样的?在尝试不同的架构时,又做了怎样的修改?为避免过度拟合数据,你是否决定添加了更多卷积层或其他层?
#
# **答案**: 最开始用了最基本的架构,只有一个卷积层;然后通过添加更多的卷积层和drop来调整网络结构;
# 为了避免过拟合,为每个卷积block添加了dropout层
#
# ### 问题3:你是如何决定训练模型的epoch数量和batch_size的?
#
# **答案**: batch_size不能太大,因为这会减慢loss,而且可能会出现内存不足,也不能太小,否则会训练速度太慢甚至不会收敛,所以最终选择了32作为batch_size;
# 通过多次尝试,epoch在35代之后loss减少的不明显了,所以epoch选择了35
#
#
# ## 特征可视化
#
# 有时,神经网络会被当做是一个黑盒子,给定一些输入,它就会学习产生一些输出。 事实上,CNN正在学习识别各种空间模式,你可以通过查看构成每个卷积核的权重并将这些一次性应用于样本图像来可视化每个卷积层已被训练识别的内容。这种技术称为特征可视化,它对于理解CNN的内部工作方式很有帮助。
#
# 在下面的单元格中,你可以看到如何从第一个卷积层中按索引提取单个滤波器。滤波器应显示为灰度网格。
# +
import torch
from models import Net
net = Net()
net.load_state_dict(torch.load('saved_models/keypoints_model_1.pt'))
net.eval()
# +
# Get the weights in the first conv layer, "conv1"
# if necessary, change this to reflect the name of your first conv layer
weights1 = net.conv1.weight.data
w = weights1.numpy()
filter_index = 3
print(w[filter_index][0])
print(w[filter_index][0].shape)
# display the filter weights
plt.imshow(w[filter_index][0], cmap='gray')
# -
# ## 特征映射
#
# 每个CNN至少包含一个由堆叠滤波器(也称为卷积核)组成的卷积层。CNN在进行训练时,它要学习在卷积内核中包含哪些权重,当这些内核应用于某些输入图像时,它们会产生一组**特征映射**。因此,特征映射只是过滤图像的集合,它们是通过将卷积核应用于输入图像而产生的图像。这些映射向我们展示了神经网络不同层学习提取的特征。例如,你可以想象一个卷积内核,它可以检测到脸部的垂直边缘,而另一个可以检测到眼角的边缘。通过将这些内核应用于图像,你可以看到每个内核检测到了哪些特征。具体请看以下示例,从它在图像中显示线条的方式,你可以将其表征为边缘检测滤波。
#
# <img src='images/feature_map_ex.png' width=50% height=50%/>
#
#
# 接下来,选择一个测试图像并使用已被训练的CNN中的一个卷积内核对其进行过滤。查看过滤后的输出,了解该内核检测到的内容。
#
# ### TODO: 过滤图像,查看卷积内核的效果
# ---
# +
import cv2
##TODO: load in and display any image from the transformed test dataset
img = test_dataset[20]
img = img["image"].numpy().reshape(224,224)
# plt.imshow(img,cmap='gray')
## TODO: Using cv's filter2D function,
## apply a specific set of filter weights (like the one displayed above) to the test image
dst = cv2.filter2D(img,-1,w[4][0])
# plt.imshow(dst,cmap='gray')
plt.figure(figsize=(12, 5))
plt.subplot(1, 2, 1)
plt.imshow(img,cmap='gray')
plt.subplot(1, 2, 2)
plt.imshow(dst,cmap='gray')
# -
# ### 问题4:从已被训练的CNN中选择一个滤波器并将其应用于测试图像。你认为它会起到什么作用?你认为它会检测到哪种特征?
#
#
# **答案**:从图中观察,图中的斜边更加明显,该滤波器的作用应该是检测斜边
#
# ---
# ## 继续加油吧!
#
# 现在,你已经定义并训练了模型,最终也保存了一个最佳模型。接下来,就是最后一个notebook,它会将人脸检测器与你保存的模型相结合,创建一个人脸关键点检测系统,用于预测一种图像中*任何一个* 人脸的关键点!
|
2. Define the Network Architecture-zh.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Making a static map of tephra fall on buildings
#
# Plot data from two different files with an open access basemap.
# +
# Import required libraries
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import contextily as ctx
# +
# Read in the shapefiles
fp = 'data/Rabaul_georef.shp'
fp2 = 'data/Tavurvur_iso.shp'
tephra = gpd.read_file(fp2)
data = gpd.read_file(fp)
tephra.head(1)
# +
# %matplotlib inline
# Check how the data look
data.plot('Rab_DI')
tephra.plot(facecolor = 'None', ec = 'k', linewidth=1)
# -
# Add a basemap using contextily
# Reproject data to web mercator (EPSG 3857)
tephra= tephra.to_crs(epsg=3857)
data = data.to_crs(epsg=3857)
data.crs
# +
# Create axes to plot figure into
fig, ax = plt.subplots(figsize=(10,8))
# Plot the data
data.plot(ax=ax, column = 'Rab_DI', cmap = 'RdYlBu_r', legend=True)
tephra.plot(ax=ax,facecolor = 'None', ec = 'k', linewidth=1, alpha = 0.5)
# Add a basemap using contextily
ctx.add_basemap(ax=ax, source=ctx.tile_providers.OSM_B)
ax.set_title('Building damage states from the 1994 Rabaul eruption')
# Use a tight layout to remove whitespace
plt.tight_layout()
# Save the figure
plt.savefig('Rabaul_94.png')
# -
|
.ipynb_checkpoints/static_map-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DNA Assemblies, gene expression, transcription, and translation
# The central Dogma - namely that DNA is transcribed to mRNA which is translated to proteins - is a key part of modelling many biochemical processes, especially in synthetic biology. Towards enabling easy modeling of transcription and translation in diverse context, BioCRNpyler has a number of Mixtures, Components and Mechanisms to produce models which include the central dogma.
# # Example 1: Creating a DNAassembly
# DNAassembly is a Component consisting of 2 sub Components: Promoter and RBS (ribosome binding site). DNAassembly automatically produces transcription and translation reactions from simple specifications: a DNAassembly X will produce 3 species dna_X, rna_X, and protein_X. These species can also be renamed manually if desired.
# +
# %matplotlib inline
from biocrnpyler import *
#promoter is the name of the promoter or a Promoter component
#RBS is the name of the RBS or an RBS Component
# RBS = None means there will be no translation
#By default (if transcript and protein are None), the transgript and protein will have the same name as an assmebly
# Users can also assign Species or String names to transcript and protein
G = DNAassembly("X", promoter = "P", rbs = "RBS", transcript = None, protein = None)
#create a transcription and translation Mechanisms.
mech_tx = SimpleTranscription()
mech_tl = SimpleTranslation()
#place that mechanism in a dictionary: {"transcription":mech_tx, "translation":mech_tl}
default_mechanisms = {mech_tx.mechanism_type:mech_tx, mech_tl.mechanism_type:mech_tl}
#Use default parameters for conviencience
default_parameters = {"kb":100, "ku":10, "ktx":.5, "ktl":2}
#Create a mixture.
M = Mixture("Catalysis Mixture", components = [G], parameters = default_parameters, default_mechanisms = default_mechanisms)
print("repr(Mixture) gives a printout of what is in a mixture and what it's Mechanisms are:\n", repr(M),"\n")
#Compile the CRN with Mixture.compile_crn
CRN = M.compile_crn()
print("Pretty_print representation of the CRN:\n",
CRN.pretty_print(show_rates = True, show_attributes = True, show_materials = True))
# -
# # Example 2: Placing a DNAassmebly in different Cell-like Mixtures for Transcription and Translation
# BioCRNpyler contains many pre-built Mixtures to model Transcription and Translation at different levels of detail in different contexts. Cell-like Mixtures are designed to more accurately model the internal environment of a cell.
#
# ## Cell-Like Mixtures:
# __ExpressionDilutionMixture__ uses the Mechanisms: OneStepGeneExpression
# * Genes express in a single step $G \to G + P_G$.
# * Proteins are degraded: $P_G \to \emptyset$.
#
# __SimpleTxTlDilutionMixture__ uses the Mechanisms: SimpleTranscription, SimpleTranslation, and Dilution (A global Mechanisms)
# * Genes transcribe via simple catalysis $G_X \to G_X + T_X$
# * mRNAs translate via simple catalyis $T_X \to T_X + P_X$.
# * Proteins and mRNA are degraded/diluted at different rates: $T_X \to \emptyset$, $P_X \to \emptyset$.
#
# __TxTlDilutionMixture__ uses the Mechanisms: Transcription_MM, Translation_MM, Degredation_mRNA_MM and Dilution (A global Mechanisms)
# * Genes transcribe via RNApolymerase (RNAP) $G_X + RNAP \rightleftarrows G_X:RNAP \to G_X + RNAP + T_X$
# * mRNAs translate via Ribosomes (Ribo) $T_X + Ribo \rightleftarrows T_X:Ribo \to T_X + Ribo + P_X$
# * mRNAs are degraded via endonucleases (Endo): $T_X + Endo \rightleftarrows T_X:Endo \to Endo$
# * Proteins & mRNA are Diluted: $P_X \to \emptyset$, $T_X \to \emptyset$
# * A Background DNAassembly $G_\textrm{cellular_processes}$ puts loading on all the cellular resources.
#
# In the following examples, the parameter file default_parameters.txt will be used to quickly produce more complex models with realistic parameters.
# +
#Changing the promoter and RBS name will result in different parameters
#Here are some parameters loaded into default_parameters.txt
#Promoter Names: strong, medium, weak correspond to bicistronic RBSs: BCD2, BCD8 and BCD12
#RBS Names: strong, medium, Weak correspond to Anderson Promoters: J23100, J23106, and J23103
G = DNAassembly("X", promoter = "strong", rbs = "weak", transcript = None, protein = None)
#Compare the Following Mixtures and Resulting CRNs
M1 = ExpressionDilutionMixture("ExpressionDilution", components = [G], parameter_file = "default_parameters.txt")
CRN1 = M1.compile_crn()
print(repr(M1),"\n", CRN1.pretty_print(show_attributes = True, show_material = True, show_rates = True),"\n\n")
M2 = SimpleTxTlDilutionMixture("SimpleTxTl", components = [G], parameter_file = "default_parameters.txt")
CRN2 = M2.compile_crn()
print(repr(M2),"\n", CRN2.pretty_print(show_attributes = True, show_material = True, show_rates = True),"\n\n")
M3 = TxTlDilutionMixture("e coli", components = [G], parameter_file = "default_parameters.txt")
CRN3 = M3.compile_crn()
print(repr(M3),"\n", CRN3.pretty_print(show_attributes = True, show_material = True, show_rates = True),"\n\n")
# -
# # Example 3: Placing a DNAassmebly in different Extract-like Mixtures for Transcription and Translation
# BioCRNpyler contains many pre-built Mixtures to model Transcription and Translation at different levels of detail in different contexts. Cell-like Mixtures are designed to more accurately model the internal environment of a cell.
#
# ## Extract-Like Mixtures:
# __ExpressionDilutionMixture__ uses the Mechanisms: OneStepGeneExpression
# * Genes express in a single step $G \to G + P_G$.
# __SimpleTxTlExtract__ uses the Mechanisms: SimpleTranscription, SimpleTranslation, and Dilution (A global Mechanisms)
# * Genes transcribe via simple catalysis $G_X \to G_X + T_X$
# * mRNAs translate via simple catalyis $T_X \to T_X + P_X$.
# * mRNA are degraded: $T_X \to \emptyset$
#
# __TxTlExtract__ uses the Mechanisms: Transcription_MM, Translation_MM, Degredation_mRNA_MM and Dilution (A global Mechanisms)
# * Genes transcribe via RNApolymerase (RNAP) $G_X + RNAP \rightleftarrows G_X:RNAP \to G_X + RNAP + T_X$
# * mRNAs translate via Ribosomes (Ribo) $T_X + Ribo \rightleftarrows T_X:Ribo \to T_X + Ribo + P_X$
# * mRNAs are degraded via endonucleases (Endo): $T_X + Endo \rightleftarrows T_X:Endo \to Endo$
#
# In the following examples, the parameter file default_parameters.txt will be used to quickly produce more complex models with realistic parameters.
# +
#Compare the Following Mixtures and Resulting CRNs
M1 = ExpressionExtract("ExpressionExtract", components = [G], parameter_file = "default_parameters.txt")
CRN1 = M1.compile_crn()
print(repr(M1),"\n", CRN1.pretty_print(show_attributes = True, show_material = True, show_rates = True),"\n\n")
M2 = SimpleTxTlExtract("SimpleTxTlExtract", components = [G], parameter_file = "default_parameters.txt")
CRN2 = M2.compile_crn()
print(repr(M2),"\n", CRN2.pretty_print(show_attributes = True, show_material = True, show_rates = True),"\n\n")
M3 = TxTlExtract("e coli extract", components = [G], parameter_file = "default_parameters.txt")
CRN3 = M3.compile_crn()
print(repr(M3),"\n", CRN3.pretty_print(show_attributes = True, show_material = True, show_rates = True),"\n\n")
# -
# ## Example 4: Retroactivity and Loading Using a Custom Promoter Object
# Most of the default Mixtures in BioCRNpyler include transcription, translation, and degredation machinery such as RNAP (RNA Polymerase), Ribosomes, and RNAases. In the following example, we will illustrate loading effects due to competition over this machinery. For this example, we will use the following model of constituitive transcription and translation:
#
# $G_i + \textrm{RNAP} \leftrightarrow G_i:\textrm{RNAP} \rightarrow G_i + \textrm{RNAP} + T_i$
#
# $T_i + \textrm{Ribosome} \leftrightarrow T_i:\textrm{Ribosome} \rightarrow T_i + \textrm{Ribosome} + P_i$
#
# $T_i + \textrm{RNAase} \leftrightarrow T_i:\textrm{RNAase} \rightarrow \textrm{RNAase}$
#
# Here $G_i$, $T_i$ and $P_i$ are gene, transcript, and protein $i$, respectively. In the example that follows, we will allow different RNA polymerases for different genes. Also, some genes may not be translated at all. By using orthogonal polymerases and/or loads without translation, we will see different kinds of loading effects.
#
# We will create 4 different DNA assemblies. The reference assembly, "ref", will have a RNAP promoter and an RBS. We will examine the output of this reporter as a function of the amount of various load assemblies.
# * The "Load" assembly will be identical to the "ref" assembly; this assembly will put load on all parts of transcription, RNA degredation, and translation for the ref assembly.
# * The "TxLoad" assembly will have an RNAP promoter, but no RBS, ensuring that only RNAP and RNAases experience loading, not ribosomes.
# * The "T7Load" assembly will have a T7 promoter and an RBS so there will be no loading on polymerases.
# * The "T7TxLoad" assmebly will have a T7 promoter and no RBS, so there will only be load on the RNAases.
#
# The creation of these assemblies highlights the flexible, object oriented nature of bioCRNpyler.
# +
from biocrnpyler import *
#Because we are lazy, all parameters will use the default "param_name"-->value key mapping.
#Parameter warnings will be later suppressed in the Mixture constructor
#For details on how parameter loading and defaulting works, see the Parameter notebook.
kb, ku, ktx, ktl, kdeg = 100, 10, 3.0, 5.0, 2
parameters = {"kb":kb, "ku":ku, "ktx":ktx, "ktl":ktl, "kdeg":kdeg}
#A constituitively expressed reporter
#By default the promoter 'P' will use the polymerase 'rnap'
reference_assembly = DNAassembly(name = "ref", promoter = "P", rbs = "BCD")
#A constiuitively expressed load (RNA and Protein)
full_load_assembly = DNAassembly(name = "Load", promoter = "P", rbs = "BCD")
#A constiutively transcribed (but not translated) load
#By putting rbs = None, DNAassembly automatically knows not to include translation
RNA_load_assembly = DNAassembly(name = "TxLoad", promoter = "P", rbs = None)
#Load genes on orthogonal polymerases
T7 = Protein("T7") #Create a new protein (polymerase) called 'T7'
#Create a custom promoter with a custom mechanism that uses T7 instead of RNAP
#instantiate a new mechanism Transcription_MM with its own name and overwrote the default parameter rnap
mechanism_txt7 = Transcription_MM(name = "T7_transcription_mm", rnap=T7)
#Create an instance of a promoter with this mechanism for transcription
T7P = Promoter("T7P", mechanisms={"transcription":mechanism_txt7})
#Create A load assembly with the custom T7 promoter
T7_load_assembly = DNAassembly(name = "T7Load", promoter = T7P, rbs = "BCD")
#Each new assembly requires its own promoter instance - so here I create another here
T7P = Promoter("T7P", mechanisms={
"transcription":Transcription_MM(name = "T7_transcription_mm", rnap=T7)})
#A load assembly with the custom T7 promoter and no RBS
T7RNA_load_assembly = DNAassembly(name = "T7TxLoad", promoter = T7P, rbs = None)
#Add all the assemblies to a mixture
components = [reference_assembly, full_load_assembly, T7_load_assembly, T7, RNA_load_assembly, T7RNA_load_assembly]
myMixture = TxTlExtract(name = "txtl", parameters = parameters, components = components, parameter_warnings = False)
#Print the CRN
myCRN = myMixture.compile_crn()
#The Species, Reaction, and CRN pretty_print functions return text which has been formated with a number of formatting options
print("\npretty_print gives a nicely formatted repesentation of the CRNS, reactions, and species. The names of species are formatted for clarity, but are not the actual species representations. Additionally a number of printing options are available.",
"\n", myCRN.pretty_print(show_material = True, show_rates = True, show_attributes = True))
#Simulate with BioSCRAPE if installed
try:
print("Simulating")
import numpy as np
import pylab as plt
timepoints = np.arange(0, 3, .01)
stochastic = False #Whether to use ODE models or Stochastic SSA models
plt.figure(figsize = (16, 8))
plt.subplot(221)
plt.title("Load on a RNAP Promoter")
loads = [0, 1.0, 5., 10., 50, 100, 500, 1000]
for dna_Load in loads:
#print("Simulating for dna_Load=", dna_Load)
x0_dict = {"protein_T7": 10., "protein_RNAP":10., "protein_RNAase":5.0, "protein_Ribo":50.,
'dna_ref':5., 'dna_Load':dna_Load}
results = myCRN.simulate_with_bioscrape(timepoints, x0_dict, stochastic = stochastic)
plt.plot(timepoints, results["protein_ref"], label = "Load = "+str(dna_Load))
plt.xlim(0, 5)
#plt.xlabel("time")
plt.ylabel("Reference Protein")
plt.legend()
plt.subplot(222)
plt.title("Load on a T7 Promotoer")
for dna_Load in loads:
#print("Simulating for dna_T7Load=", dna_Load)
x0_dict = {"protein_T7": 10., "protein_RNAP":10., "protein_RNAase":5.0, "protein_Ribo":50.,
'dna_ref':5., 'dna_T7Load':dna_Load}
results = myCRN.simulate_with_bioscrape(timepoints, x0_dict, stochastic = stochastic)
plt.plot(timepoints, results["protein_ref"], label="Load = " + str(dna_Load))
plt.xlim(0, 5)
#plt.xlabel("time")
plt.ylabel("Reference Protein")
plt.legend()
plt.subplot(223)
plt.title("Load on a RNAP Promotoer, No RBS")
for dna_Load in loads:
#print("Simulating for dna_TxLoad=", dna_Load)
x0_dict = {"protein_T7": 10., "protein_RNAP":10., "protein_RNAase":5.0, "protein_Ribo":50.,
'dna_ref':5., 'dna_TxLoad':dna_Load}
results = myCRN.simulate_with_bioscrape(timepoints, x0_dict, stochastic = stochastic)
plt.plot(timepoints, results["protein_ref"], label="Load = " + str(dna_Load))
plt.xlim(0, 5)
plt.xlabel("time")
plt.ylabel("Reference Protein")
plt.legend()
plt.subplot(224)
plt.title("Load on a T7 Promotoer, No RBS")
for dna_Load in loads:
#print("Simulating for dna_T7TxLoad=", dna_Load)
x0_dict = {"protein_T7": 10., "protein_RNAP":10., "protein_RNAase":5.0, "protein_Ribo":50.,
'dna_ref':5., 'dna_T7TxLoad':dna_Load}
results = myCRN.simulate_with_bioscrape(timepoints, x0_dict, stochastic = stochastic)
plt.plot(timepoints, results["protein_ref"], label="Load = " + str(dna_Load))
plt.xlim(0, 5)
plt.xlabel("time")
plt.ylabel("Reference Protein")
plt.legend()
plt.show()
except ModuleNotFoundError:
pass
# -
|
examples/3. DNA Assemblies, gene expression, transcription, and translation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
pd.set_option('max_rows', 7)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = 10, 8
import seaborn as sns
# -
movies = pd.read_csv("../data/Section6-Homework-Data.csv", encoding = "ISO-8859-1")
movies.head()
movies.describe()
movies.info()
selected_genres = ['action', 'comedy', 'adventure', 'animation',
'drama']
selected_studios = ['Buena Vista Studios', 'Sony', 'Universal',
'WB', 'Paramount Pictures', 'Fox']
filter = movies.Genre.isin(selected_genres) & movies.Studio.isin(selected_studios)
sns.set_style('whitegrid')
w = sns.boxplot(data=movies[filter],
x='Genre', y='Gross % US',
color='lightgrey')
w = sns.swarmplot(data=movies[filter],
x='Genre', y='Gross % US', hue='Studio')
w.set_xlabel('Genre', fontsize=20)
w.set_ylabel('Gross % US', fontsize=20)
w.set_title('Domestic Gross % by Genre', fontsize=40)
w.legend(loc='bottom left', bbox_to_anchor=(1, 1))
|
Section6/.ipynb_checkpoints/Homework-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import fsspec
import numpy as np
import imageio
# +
# %%time
url = 'cmgp-coastcam/cameras/caco-01/foo.txt'
fs = fsspec.filesystem('s3')
with fs.open(url) as f:
im = f.read()
print(im)
# +
with fs.open('cmgp-coastcam/cameras/caco-01/products/1600866001.c2.timex.jpg') as f:
im = imageio.read(f)
print(im)
# -
recent_list=fs.glob('cmgp-coastcam/cameras/caco-01/latest/*')
print(recent_list)
|
test_bucket_read.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="HMS4M6WQTOox" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592511766644, "user_tz": 420, "elapsed": 656, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
import os
import glob
import numpy as np
from tensorflow.keras import layers
from tensorflow import keras
import tensorflow as tf
# + id="teO33zkZTiLO" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592511769873, "user_tz": 420, "elapsed": 1401, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
categories_text = open('/content/drive/My Drive/category_mini.txt','r')
categories = categories_text.readlines()
categories_text.close()
# + id="YtLQivZ0T_Bc" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592511770977, "user_tz": 420, "elapsed": 586, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
categories = [c.replace('\n', '').replace(' ','_').replace("'", "").replace(',', '') for c in categories]
# + id="haoLTZKTUIF0" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592511773527, "user_tz": 420, "elapsed": 1659, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
# !mkdir npy
# + id="kC1flDuthmz3" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592511839068, "user_tz": 420, "elapsed": 603, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
cat_limit = categories[:25]
# + id="VsqvfHRBUJpY" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592511840001, "user_tz": 420, "elapsed": 519, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
import urllib.request
def download():
base = 'https://storage.googleapis.com/quickdraw_dataset/full/numpy_bitmap/'
counter = 1
for i in cat_limit:
space = i.replace('_', '%20')
path = base+space+'.npy'
print(f'{path} {counter}/{len(cat_limit)}')
counter += 1
urllib.request.urlretrieve(path, 'npy/'+i+'.npy')
# + id="jeBZS2KoUmCd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} executionInfo={"status": "ok", "timestamp": 1592511860225, "user_tz": 420, "elapsed": 19452, "user": {"displayName": "Brian Perry", "photoUrl": "", "userId": "10921880190671207311"}} outputId="e853cdce-e9e6-4b85-b2df-bc3141599660"
download()
# + id="-DD_wPWaUtX-" colab_type="code" colab={}
all_files = glob.glob(os.path.join('npy', '*.npy'))
# + id="D1ArXEhHE84s" colab_type="code" colab={}
X = np.empty([0, 784])
y = np.empty([0])
label_names = []
for idx, file in enumerate(all_files):
data = np.load(file)
data = data[0: 10000, :]
labels = np.full(data.shape[0], idx)
X = np.concatenate((X, data), axis=0)
y = np.append(y, labels)
label_name, extension = os.path.splitext(os.path.basename(file))
label_names.append(label_name)
# + colab_type="code" id="NR-ZLL8QriUO" colab={"base_uri": "https://localhost:8080/", "height": 298} executionInfo={"status": "ok", "timestamp": 1592452368337, "user_tz": 420, "elapsed": 64900, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="d5c11773-050e-4af7-a111-fc0c42cef24d"
import matplotlib.pyplot as plt
from random import randint
# %matplotlib inline
idx = randint(0, X.shape[0])
print(idx)
plt.imshow(X[idx].reshape(28,28), cmap='binary')
print(label_names[int(y[idx].item())])
# + id="SdVMH3bZFYqr" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.20, random_state=24)
# + id="JmlaONEAXemD" colab_type="code" colab={}
X_train, X_test = X_train / 255.0, X_test / 255.0
# + id="mqfAClCdvdXD" colab_type="code" colab={}
X_train = tf.reshape(X_train,[-1,28,28])
X_test = tf.reshape(X_test,[-1,28,28])
X_train = tf.expand_dims(X_train, 3)
X_test = tf.expand_dims(X_test, 3)
# + id="gH8aJjvWXtNH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1592452378856, "user_tz": 420, "elapsed": 75390, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="33f1fb8a-525e-4dc2-8a73-a3d46d39d5f3"
X_train.shape
# + id="wU0aEzgoGU9I" colab_type="code" colab={}
y_train = tf.keras.utils.to_categorical(y_train, num_classes=len(label_names))
y_test = tf.keras.utils.to_categorical(y_test, num_classes=len(label_names))
# y_train = tf.convert_to_tensor(y_train)
# y_test = tf.convert_to_tensor(y_test)
# + id="J3U4yuFtRQD4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} executionInfo={"status": "ok", "timestamp": 1592452378857, "user_tz": 420, "elapsed": 75377, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="85de06d1-fde4-4f26-d34a-50df9f837694"
print(f'{y_test.shape}.{y_train.shape}')
print(f'{X_test.shape},{X_train.shape}')
y_train.shape
# + id="NXi9UJ3WLwfO" colab_type="code" colab={}
# train_dataset = tf.data.Dataset.from_tensor_slices((X_train, y_train)) \
# .shuffle(2500).batch(32)
# test_dataset = tf.data.Dataset.from_tensor_slices((X_test, y_test)) \
# .batch(32)
# BATCH_SIZE = 64
# SHUFFLE_BUFFER_SIZE = 100
# train_dataset = train_dataset.shuffle(SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE)
# test_dataset = test_dataset.batch(BATCH_SIZE)
# print(train_dataset.element_spec)
# + id="LkOrSkyLIWuA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 538} executionInfo={"status": "ok", "timestamp": 1592452378859, "user_tz": 420, "elapsed": 75364, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="6631b63f-38ea-4b59-efab-c4c229681738"
model = keras.Sequential()
model.add(layers.Convolution2D(16, (3,3),
padding='same',
input_shape=X_train.shape[1:], activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(2,2)))
model.add(layers.Convolution2D(32, (3,3), padding='same', activation='relu'))
model.add(layers.Convolution2D(32, (3,3), padding='same', activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(2,2)))
model.add(layers.Convolution2D(64, (3,3), padding='same', activation='relu'))
model.add(layers.Convolution2D(64, (3,3), padding='same', activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(2,2)))
model.add(layers.Flatten())
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dense(len(cat_limit), activation='softmax'))
adam = tf.optimizers.Adam()
model.compile(loss='categorical_crossentropy',
optimizer=adam,
metrics=['accuracy'])
print(model.summary())
# + id="qTSuiEjzIYnG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"status": "ok", "timestamp": 1592452415509, "user_tz": 420, "elapsed": 112004, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="e4725df7-6fdc-4fca-e313-2226148d7d41"
model.fit(x=X_train, y=y_train, batch_size=256, validation_split=0.1, verbose=2, epochs=5)
# + [markdown] id="oz9adQjIPN_o" colab_type="text"
#
# + id="rU6lQVLhMAN8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} executionInfo={"status": "ok", "timestamp": 1592452419788, "user_tz": 420, "elapsed": 116275, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="b6ff07cd-6bb5-4c48-a75b-541c5bfcc54e"
score = model.evaluate(X_test, y_test, verbose=1)
print('Test accuracy: {:0.2f}%'.format(score[1] * 100))
# + colab_type="code" id="f-ZB6twccBIG" colab={"base_uri": "https://localhost:8080/", "height": 298} executionInfo={"status": "ok", "timestamp": 1592452419791, "user_tz": 420, "elapsed": 116271, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="c7139377-fcfb-42ab-ad23-bc03358690f0"
import matplotlib.pyplot as plt
from random import randint
# %matplotlib inline
idx = randint(0, X.shape[0])
print(idx)
plt.imshow(X[idx].reshape(28,28), cmap='binary')
print(label_names[int(y[idx].item())])
# + id="ai2ApJi9qr7m" colab_type="code" colab={}
# # !pip install tensorflowjs
# + id="uL6HUmCYlD_Q" colab_type="code" colab={}
# with open('class_names25.txt', 'w') as file_handler:
# for item in cat_limit:
# file_handler.write(f'{item}')
# + id="5Y947a_Ommsf" colab_type="code" colab={}
# model.save('keras25.h5')
# + id="f9-Ph5Ayuf_w" colab_type="code" colab={}
# # !mkdir model
# + id="_6qrDjYrmqGL" colab_type="code" colab={}
# # !tensorflowjs_converter --input_format keras keras25.h5 model/
# + id="5qQKsq01m22z" colab_type="code" colab={}
# # !cp class_names25.txt model/class_names25.txt
# + id="xFLmXinUnmQ8" colab_type="code" colab={}
# # !zip -r model25.zip model
# + id="My1A62VGnvyj" colab_type="code" colab={}
# from google.colab import files
# files.download('model25.zip')
# + id="kAFcpJTqn5WP" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592505948318, "user_tz": 420, "elapsed": 634, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
import matplotlib.pyplot as plt
import numpy as np
import random
# %matplotlib inline
# + colab_type="code" id="V63fBqz3nN-W" colab={"base_uri": "https://localhost:8080/", "height": 284} executionInfo={"status": "ok", "timestamp": 1592511746218, "user_tz": 420, "elapsed": 1475, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="a083041e-a3e1-41f6-a7cd-b1dc95b71359"
rand = np.floor(np.random.random((28,28)) + .5)
all_black = np.full((28,28, 3), 0)
all_white = np.ones((28,28))
diag = np.eye(28)
diag_low = np.eye(28, k=-15)
diag_high = np.eye(28, k=15)
plt.subplot(231)
plt.imshow(all_white, cmap='binary')
plt.subplot(232)
plt.imshow(rand, cmap='binary')
plt.subplot(233)
plt.imshow(all_black, cmap='binary')
plt.subplot(234)
plt.imshow(diag_low, cmap='binary')
plt.subplot(235)
plt.imshow(diag, cmap='binary')
plt.subplot(236)
plt.imshow(diag_high, cmap='binary')
# + id="YWQfcaLbnEx2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1592518459288, "user_tz": 420, "elapsed": 588, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="69b44c8b-2303-4555-9759-1a94d59e0545"
first6 = cat_limit[:6]
first6
# + id="OvsldMITufM6" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592518463113, "user_tz": 420, "elapsed": 830, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
dummy_files = glob.glob(os.path.join('npy', '*.npy'))
dummy_data = []
for index, file in enumerate(dummy_files):
test_data = np.load(file)
dummy_data.append(test_data[:])
# + [markdown] id="0wi-wUzbgqwj" colab_type="text"
# You can change which category of doodles by changing the index number of dummy_data below.
# + id="qDeTvPtIZpXm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1592518564757, "user_tz": 420, "elapsed": 1082, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="8f4e765a-8e13-42fd-8685-646463ee0aa9"
category_array = np.array(dummy_data[0])
category_array.shape
# + [markdown] id="nLmIog2IhEUq" colab_type="text"
# Set up to take select a random image from the corresponding category and reshapes it so it is plot-able.
#
# Feel free to select your own index if you prefer.
# + id="QzWDA_oqG36T" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592518445011, "user_tz": 420, "elapsed": 974, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
plot_me = category_array[random.randrange(len(category_array[0]))]
plot_me = plot_me.reshape(28,28)
# + id="krBW8PhmP1Ly" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} executionInfo={"status": "ok", "timestamp": 1592518446483, "user_tz": 420, "elapsed": 1102, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="c77dbda7-f68c-44e2-ceaf-8344ac09d120"
plt.imshow(plot_me, cmap='binary')
# + id="fQDro12NQskW" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1592518336189, "user_tz": 420, "elapsed": 586, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}}
butterfly = dummy_files[1]
# + id="XM1F7E7GY0Tz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1592518337514, "user_tz": 420, "elapsed": 660, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10921880190671207311"}} outputId="00741e18-be33-491f-f95f-86588fd30d90"
butterfly_data = np.load(butterfly)
butterfly_data.shape
# + id="r1oxonR_Y8h_" colab_type="code" colab={}
|
model/model25/doodle2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
base_dir = 'C:\\Users\\heineib\\Documents\\GitHub\\PlateReaderTools' #You will need to change this depending on which computer you are working on
os.chdir(base_dir)
from core import plate_reader_tools
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from datetime import datetime
from itertools import chain
data_dir = "C:\\Users\\heineib\\Google Drive\\Crick_LMS\\Wetlab\\"
# %load_ext autoreload
# %autoreload 2
# -
C:\Users\heineib\Google Drive\Crick_LMS\Wetlab\biospa_plate_reader\20201217_biospa_test_10strain_ox
# +
## In the output file it said the temp was 37 - is that correct?
# +
dirname = "C:\\Users\\heineib\\Google Drive\\Crick_LMS\\Wetlab\\biospa_plate_reader\\20201217_biospa_test_10strain_ox"
plate_data = pd.read_table(dirname + os.sep + 'P2.txt', skiprows=2, index_col = 0)
plate_data.drop(columns='Unnamed: 43', inplace=True)
plate_data.drop(index='Temp OD:595', inplace=True)
# +
#Build time vector
datetimes = [datetime.strptime(ind, '%H:%M:%S') for ind in plate_data.columns]
dt = np.array([(datetime_n-datetimes[0]).total_seconds() for datetime_n in datetimes])/3600
cycles = len(dt)
rows = 8
columns = 12
# +
#For tecan had this to filter over values
#data_filtered, OVER_list = plate_reader_tools.over_filter(data)
# -
#Plot all wells in an array similar to the plate.
letter_range = ['A','B','C','D','E','F','G','H']
number_range = range(1,13)
well_arr = plate_reader_tools.well_array_builder(letter_range,number_range)
fig = plate_reader_tools.subplot_array(plate_data, well_arr, dt)
# +
conds = ['cont', '0.07mM H2O2', '0.5mM H2O2', '1.0mM H2O2'] #each plate is a different condition so it is not part of the multi-index
layer_names = ['Strain','Dilution','Tech_Rep']
strains = ['WT','758','759', '762', '838', '837', '858', '869', '939', '1206']
#conds = ['SM','SM+HLUM']
dilutions = ['1x','0.5x']
tech_reps = ['TR1','TR2','TR3']
experiment_setup = [strains, dilutions, tech_reps]
data_inds = {}
for cond in conds:
data_inds[cond] = pd.MultiIndex.from_product(experiment_setup, names= layer_names)
#Print one data index
data_index = data_inds['cont']
print(data_index.levels)
print(data_index.names)
print(data_index)
# +
well_list = []
rows = 'BDFCEG'
col_groups = [[2,5,8],[3,6,9],[4,7,10], [5,8,11], [6,9,2], [7,10,3], [8,11,4],[9,2,5], [10,3,6],[11,4,7]]
for col_group in col_groups:
for row, col in zip(rows, col_group + col_group):
well_list.append(row + str(col))
# +
# Correcting for mistakes in experiment on second plate (had pipetter shifted over one)
#remove missing conditions from data
# Step 1: make index list for the coordinates of all the missing items based on pairs of missing conditions.
# # TR2 1x and TR1 0.5xx were shifted
data_index= data_inds['0.07mM H2O2']
data_index_df = data_index.to_frame()
data_index_df.drop(('WT','0.5x','TR1'), inplace=True)
data_index_df.drop(('762','1x','TR2'), inplace=True)
#data_index_adjusted, inds_to_remove = plate_reader_tools.delete_multiindex_for_missing_conditions(missing_items, data_index)
data_index_adjusted = pd.MultiIndex.from_frame(data_index_df)
data_index_adjusted
data_inds['0.07mM H2O2']=data_index_adjusted
well_list_adjusted = ['B2',
'D4',#'D5',
'F8',
# 'C2',
'E5',
'G8',
'B3',
'D5',# 'D6',
'F9',
'C2', #'C3',
'E6',
'G9',
'B4',
'D6', #'D7',
'F10',
'C3', #'C4',
'E7',
'G10',
'B5',
'D7', #'D8',
'F11',
'C4', #'C5',
'E8',
'G11',
'B6',
'D8', #'D9',
'F2',
'C5', #'C6',
'E9',
'G2',
'B7',
'D9', #'D10',
'F3',
'C6', #'C7',
'E10',
'G3',
'B8',
'D10', #'D11',
'F4',
'C7', #'C8',
'E11',
'G4',
'B9',
#'D2',
'F5',
'C8', #'C9',
'E2',
'G5',
'B10',
'D2', #'D3',
'F6',
'C9', #'C10',
'E3',
'G6',
'B11',
'D3', #'D4',
'F7',
'C10', #'C11',
'E4',
'G7']
# +
plate_dict = {'cont': 'P1', '0.07mM H2O2': 'P2', '0.5mM H2O2': 'P3', '1.0mM H2O2': 'P4'}
dirname = "C:\\Users\\heineib\\Google Drive\\Crick_LMS\\Wetlab\\biospa_plate_reader\\20201217_biospa_test_10strain_ox"
raw_data_all_plates = {}
growth_data_all_plates = {}
for cond in conds:
plate_data = pd.read_table(dirname + os.sep + plate_dict[cond] + '.txt', skiprows=2, index_col = 0)
plate_data.drop(columns='Unnamed: 43', inplace=True)
plate_data.drop(index='Temp OD:595', inplace=True)
raw_data_all_plates[cond] = plate_data
data=plate_data
blank = np.mean(data.loc['A1',:])
growth_data = []
if cond == '0.07mM H2O2':
growth_data.append([data.loc[well,:]-blank for well in well_list_adjusted])
else:
growth_data.append([data.loc[well,:]-blank for well in well_list])
growth_data = list(chain.from_iterable(growth_data)) # removes empty first dimension from growth_data
growth_data_df = pd.DataFrame(growth_data, index=data_inds[cond])
growth_data_all_plates[cond] = growth_data_df
# -
growth_data_all_plates['0.07mM H2O2']
int(np.floor(/5))
# +
fig, axarr = plt.subplots(2,5, figsize = (20,10), sharex=True, sharey = True)
colors = {'cont': 'black',
'0.07mM H2O2': 'orange',
'0.5mM H2O2': 'lightcoral',
'1.0mM H2O2': 'red'
}
linestyles = {'TR1': '-',
'TR2': '--',
'TR3': ':'
}
dilution = '0.5x'
for jj, strain in enumerate(strains):
axcol = np.mod(jj, 5)
axrow = int(np.floor(jj/5))
ax = axarr[axrow, axcol]
ax.set_title(strain)
for cond in conds:
color = colors[cond]
growth_data_df = growth_data_all_plates[cond]
for kk, tech_rep in enumerate(tech_reps):
try:
if kk==0:
ax.plot(dt,growth_data_df.loc[(strain, dilution, tech_rep)], color = color, linestyle = linestyles[tech_rep], label=cond )
else:
ax.plot(dt,growth_data_df.loc[(strain, dilution, tech_rep)], color = color, linestyle = linestyles[tech_rep] )
except KeyError:
print('missing data for {},{},{} in {}'.format(strain,dilution,tech_rep, cond))
if jj==0:
ax.legend()
# for jj, dil in enumerate(dilutions):
# ax = axarr[jj]
# for strain in strains:
# for cond in conds:
# color = colors[strain + '_' + cond]
# for kk, tech_rep in enumerate(tech_reps):
# if kk ==0:
# ax.plot(t,growth_data_df.loc[(strain, cond, dil, tech_rep)], color = color, label=strain + '_' + cond )
# else:
# ax.plot(t,growth_data_df.loc[(strain, cond, dil, tech_rep)], color = color )
# ax.set_title('Initial OD: ' + dil)
# if jj == 0:
# ax.legend()
|
scripts/Ben/20201217_10strain_biospa_growth.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Maximising classification accuracy via Ensemble Weight optimization
# ## Add the imports
# +
from models import wide_residual_net as WRN
import numpy as np
import sklearn.metrics as metrics
from scipy.optimize import minimize
from sklearn.metrics import log_loss
from tensorflow.keras.datasets import cifar100
from tensorflow.keras import backend as K
import tensorflow.keras.utils.np_utils as kutils
# -
# ## Some variables which we will use in some time
# +
NUM_TESTS = 25
n = 2 * 6 + 4
k = 4
models_filenames = [r"weights/WRN-CIFAR100-%d-%d-Best.h5" % (n, k),
r"weights/WRN-CIFAR100-%d-%d-1.h5" % (n, k),
r"weights/WRN-CIFAR100-%d-%d-2.h5" % (n, k),
r"weights/WRN-CIFAR100-%d-%d-3.h5" % (n, k),
r"weights/WRN-CIFAR100-%d-%d-4.h5" % (n, k),
r"weights/WRN-CIFAR100-%d-%d-5.h5" % (n, k)]
# -
# ## Load up the CIFAR 100 dataset and prepare for testing
# +
(trainX, trainY), (testX, testY) = cifar100.load_data()
nb_classes = len(np.unique(testY))
trainX = trainX.astype('float32')
trainX /= 255.0
testX = testX.astype('float32')
testX /= 255.0
trainY = kutils.to_categorical(trainY)
testY_cat = kutils.to_categorical(testY)
# -
# ## Create the Wide Residual Network (16-4)
# +
# Decide dim ordering for Theano and Tensorflow backends
if K.image_data_format() == "th":
init = (3, 32, 32)
else:
init = (32, 32, 3)
model = WRN.create_wide_residual_network(init, nb_classes=100, N=2, k=4, dropout=0.00)
model_prefix = 'WRN-CIFAR100-%d-%d' % (n, k)
# -
# ## Obtain predictions from each of the Ensemble models
preds = []
for fn in models_filenames:
model.load_weights(fn)
yPreds = model.predict(testX, batch_size=128)
preds.append(yPreds)
print("Obtained predictions from model with weights = %s" % (fn))
# ## Define helper function to calculate accuracy and error
def calculate_weighted_accuracy(prediction_weights):
weighted_predictions = np.zeros((testX.shape[0], nb_classes), dtype='float32')
for weight, prediction in zip(prediction_weights, preds):
weighted_predictions += weight * prediction
yPred = np.argmax(weighted_predictions, axis=1)
yTrue = testY
accuracy = metrics.accuracy_score(yTrue, yPred) * 100
error = 100 - accuracy
print("Accuracy : ", accuracy)
print("Error : ", error)
# ## Consider a Single Best Model prediction.
#
# We can load the weights of the single best model and make predictions
# +
# Load the weights of the best single model
model.load_weights(models_filenames[0])
# Get its predictions
yPreds = model.predict(testX, batch_size=128)
yPred = np.argmax(yPreds, axis=1)
yTrue = testY
# Calculate accuracy metric
accuracy = metrics.accuracy_score(yTrue, yPred) * 100
error = 100 - accuracy
print("Accuracy : ", accuracy)
print("Error : ", error)
# -
# ## Consider a non weighted ensemble prediction
#
# Here, each model has the same weight for predictions. However, this may not lead to optimal results.
#
# Notice that ensemble weighting is an improvement over the single best model, by a large margin for CIFAR100
# +
prediction_weights = [1. / len(models_filenames)] * len(models_filenames)
calculate_weighted_accuracy(prediction_weights)
# -
# ## Now we consider a weighted ensemble
#
# In a weighted ensemble, we try to optimize the weights of predictions of each model, so as to minimize the total log loss. This in turn improves the overall accuracy of the predictions
# Create the loss metric
def log_loss_func(weights):
''' scipy minimize will pass the weights as a numpy array '''
final_prediction = np.zeros((testX.shape[0], nb_classes), dtype='float32')
for weight, prediction in zip(weights, preds):
final_prediction += weight * prediction
return log_loss(testY_cat, final_prediction)
# +
best_acc = 0.0
best_weights = None
# Parameters for optimization
constraints = ({'type': 'eq', 'fun':lambda w: 1 - sum(w)})
bounds = [(0, 1)] * len(preds)
# Check for NUM_TESTS times
for iteration in range(NUM_TESTS):
# Random initialization of weights
prediction_weights = np.random.random(len(models_filenames))
# Minimise the loss
result = minimize(log_loss_func, prediction_weights, method='SLSQP', bounds=bounds, constraints=constraints)
print('Best Ensemble Weights: {weights}'.format(weights=result['x']))
weights = result['x']
weighted_predictions = np.zeros((testX.shape[0], nb_classes), dtype='float32')
# Calculate weighted predictions
for weight, prediction in zip(weights, preds):
weighted_predictions += weight * prediction
yPred = np.argmax(weighted_predictions, axis=1)
yTrue = testY
# Calculate weight prediction accuracy
accuracy = metrics.accuracy_score(yTrue, yPred) * 100
error = 100 - accuracy
print("Iteration %d: Accuracy : " % (iteration + 1), accuracy)
print("Iteration %d: Error : " % (iteration + 1), error)
# Save current best weights
if accuracy > best_acc:
best_acc = accuracy
best_weights = weights
print()
# -
# ## We can now compute the best accuracy ensemble model
print("Best Accuracy : ", best_acc)
print("Best Weights : ", best_weights)
calculate_weighted_accuracy(best_weights)
|
optimize_cifar100.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting data from web archives using Memento
#
# <p class="alert alert-info">New to Jupyter notebooks? Try <a href="getting-started/Using_Jupyter_notebooks.ipynb"><b>Using Jupyter notebooks</b></a> for a quick introduction.</p>
#
# Systems supporting the Memento protocol provide machine-readable information about web archive captures, even if other APIs are not available. In this notebook we'll look at the way the Memento protocol is supported across four web archive repositories – the UK Web Archive, the National Library of Australia, the National Library of New Zealand, and the Internet Archive. In particular we'll examine:
#
# * [Timegates](#Timegates) – request web page captures from (around) a particular date
# * [Timemaps](#Timemaps) – request a list of web archive captures from a particular url
# * [Mementos](#Mementos) – use url modifiers to change the way an archived web page is presented
#
# Notebooks using Timegates or Timemaps to access capture data include:
#
# * [Get the archived version of a page closest to a particular date](get_a_memento.ipynb)
# * [Find all the archived versions of a web page](find_all_captures.ipynb)
# * [Harvesting collections of text from archived web pages](getting_text_from_web_pages.ipynb)
# * [Compare two versions of an archived web page](show_diffs.ipynb)
# * [Create and compare full page screenshots from archived web pages](save_screenshot.ipynb)
# * [Using screenshots to visualise change in a page over time](screenshots_over_time_using_timemaps.ipynb)
# * [Display changes in the text of an archived web page over time](display-text-changes-from-timemap.ipynb)
# * [Find when a piece of text appears in an archived web page](find-text-in-page-from-timemap.ipynb)
#
# ## Useful tools and documentation
# * [Memento Protocol Specification](https://tools.ietf.org/html/rfc7089)
# * [Pywb Memento implementation](https://pywb.readthedocs.io/en/latest/manual/memento.html)
# * [Memento support in IA Wayback](https://ws-dl.blogspot.com/2013/07/2013-07-15-wayback-machine-upgrades.html)
# * [Time Travel APIs](https://timetravel.mementoweb.org/guide/api/)
# * [Memento Compliance Audit of PyWB](https://ws-dl.blogspot.com/2020/03/2020-03-26-memento-compliance-audit-of.html)
# * [Memento tools](http://mementoweb.org/tools/)
# * [Memento client](https://github.com/mementoweb/py-memento-client)
# * [Memgator](https://github.com/oduwsdl/MemGator) – Memento aggregator
# +
import requests
import arrow
import re
import json
# Alternatively use the python Memento client
# -
# These are the repositories we'll be using
TIMEGATES = {
'awa': 'https://web.archive.org.au/awa/',
'nzwa': 'https://ndhadeliver.natlib.govt.nz/webarchive/wayback/',
'ukwa': 'https://www.webarchive.org.uk/wayback/archive/',
'ia': 'https://web.archive.org/web/'
}
# ## Timegates
#
# Timegates let you query a web archive for the capture closest to a specific date. You do this by supplying your target date as the `Accept-Datetime` value in the headers of your request.
#
# For example, if you wanted to query the Australian Web Archive to find the version of `http://nla.gov.au/` that was captured as close as possible to 1 January 2001, you'd set the `Accept-Datetime` header to header to 'Fri, 01 Jan 2010 01:00:00 GMT' and request the url:
#
# ```
# https://web.archive.org.au/awa/http://nla.gov.au/
# ```
#
# A `get` request will return the captured page, but if all you want is the url of the archived page you can use a `head` request and extract the information you need from the response headers. Try this:
response = requests.head('https://web.archive.org.au/awa/http://nla.gov.au/', headers={'Accept-Datetime': 'Fri, 01 Jan 2010 01:00:00 GMT'})
response.headers
# The request above returns the following headers:
#
# ``` python
# {
# 'Server': 'nginx',
# 'Date': 'Wed, 06 May 2020 04:34:50 GMT',
# 'Content-Length': '0', 'Connection': 'keep-alive',
# 'Location': 'https://web.archive.org.au/awa/20100205144227/http://nla.gov.au/',
# 'Link': '<http://nla.gov.au/>; rel="original", <https://web.archive.org.au/awa/http://nla.gov.au/>; rel="timegate", <https://web.archive.org.au/awa/timemap/link/http://nla.gov.au/>; rel="timemap"; type="application/link-format", <https://web.archive.org.au/awa/20100205144227mp_/http://nla.gov.au/>; rel="memento"; datetime="Fri, 05 Feb 2010 14:42:27 GMT"',
# 'Vary': 'accept-datetime'
# }
# ```
#
# The `Link` parameter contains the Memento information. You can see that it's actually providing information on four types of link:
#
# * the `original` url (ie the url that was archived) – `<http://nla.gov.au/>`
# * the `timegate` for the harvested url (which us what we just used) – `<https://web.archive.org.au/awa/http://nla.gov.au/>`
# * the `timemap` for the harvested url (we'll look at this below) – `<https://web.archive.org.au/awa/timemap/link/http://nla.gov.au/>`
# * the `memento` – `<https://web.archive.org.au/awa/20100205144227mp_/http://nla.gov.au/>`
#
# The `memento` link is the capture closest in time to the date we requested. In this case there's only about a month's difference, but of course this will depend on how frequently a url is captured. Opening the link will display the capture in the web archive. As we'll see below, some systems provide additional links such as `first memento`, `last memento`, `prev memento`, and `next memento`.
# Here's some functions to query a timegate in one of the four systems we're exploring. We'll use them to compare the results we get from each.
# +
def format_date_for_headers(iso_date, tz):
'''
Convert an ISO date (YYYY-MM-DD) to a datetime at noon in the specified timezone.
Convert the datetime to UTC and format as required by Accet-Datetime headers:
eg Fri, 23 Mar 2007 01:00:00 GMT
'''
local = arrow.get(f'{iso_date} 12:00:00 {tz}', 'YYYY-MM-DD HH:mm:ss ZZZ')
gmt = local.to('utc')
return f'{gmt.format("ddd, DD MMM YYYY HH:mm:ss")} GMT'
def parse_links_from_headers(response):
'''
Extract original, timegate, timemap, and memento links from 'Link' header.
'''
links = response.links
return {k: v['url'] for k, v in links.items()}
def format_timestamp(timestamp, date_format='YYYY-MM-DD HH:mm:ss'):
return arrow.get(timestamp, 'YYYYMMDDHHmmss').format(date_format)
def query_timegate(timegate, url, date=None, tz='Australia/Canberra', request_type='head', allow_redirects=True):
headers = {}
if date:
formatted_date = format_date_for_headers(date, tz)
headers['Accept-Datetime'] = formatted_date
# Note that you don't get a timegate response if you leave off the trailing slash
tg_url = f'{TIMEGATES[timegate]}{url}/' if not url.endswith('/') else f'{TIMEGATES[timegate]}{url}'
print(tg_url)
if request_type == 'head':
response = requests.head(tg_url, headers=headers, allow_redirects=allow_redirects)
else:
response = requests.get(tg_url, headers=headers, allow_redirects=allow_redirects)
# print(response.headers)
return parse_links_from_headers(response)
# -
# ### Australian Web Archive
#
# A `HEAD` request that follows redirects returns no results
query_timegate('awa', 'http://www.nla.gov.au')
# ----
# A `HEAD` request that doesn't follow redirects returns results as expected
query_timegate('awa', 'http://www.nla.gov.au', allow_redirects=False)
# ----
# A query without an `Accept-Datetime` value returns a recent capture.
query_timegate('awa', 'http://www.nla.gov.au', allow_redirects=False)
# ----
#
# A query with an `Accept-Datetime` value of 1 January 2002 returns a capture from 20 January 2002.
query_timegate('awa', 'http://www.education.gov.au/', date='2002-01-01', allow_redirects=False)
# ----
#
# Using a `GET` rather than a `HEAD` request returns no Memento information when redirects are followed.
query_timegate('awa', 'http://www.education.gov.au/', date='2002-01-01', request_type='get')
# ----
#
# Using a `GET` rather than a `HEAD` request returns Memento information when redirects are not followed.
query_timegate('awa', 'http://www.education.gov.au/', date='2002-01-01', request_type='get', allow_redirects=False)
# ### New Zealand Web Archive
#
# Changing whether or not redirects are followed has no effect on any of these responses.
#
# A query without an `Accept-Datetime` value doesn't return a `memento`, but does include `first memento`, `last memento`, and `prev memento`.
query_timegate('nzwa', 'http://natlib.govt.nz')
# ----
#
# A query with an `Accept-Datetime` value of 1 January 2005 doesn't return a `memento`, even though there's a capture available from July 2004. I don't know why this is.
query_timegate('nzwa', 'http://natlib.govt.nz', date='2005-01-01')
# ----
#
# A query with an `Accept-Datetime` value of 1 January 2008 returns a `memento` from 25 February 2008, as well as `first memento`, `last memento`, `prev memento`, and `next memento`.
query_timegate('nzwa', 'http://natlib.govt.nz', date='2008-01-01')
# ----
#
# A `GET` request returns the same results as a `HEAD` request.
query_timegate('nzwa', 'http://natlib.govt.nz', date='2008-01-01', request_type='get')
# ### Internet Archive
#
# Using a `HEAD` request that follows redirects returns results as expected.
query_timegate('ia', 'http://discontents.com.au')
# ----
# Using a `HEAD` request returns no Memento information if redirects are not followed.
query_timegate('ia', 'http://discontents.com.au', allow_redirects=False)
# ----
#
# A query without an `Accept-Datetime` value returns a `memento` and also includes a `first memento`, `last memento`, `prev memento`, and `last memento`. It seems that the `memento` returned is the second last capture.
query_timegate('ia', 'http://discontents.com.au')
# ----
#
# A query with an `Accept-Datetime` value of 1 January 2010 returns a `memento` from 4 September 2010, even though the `prev memento` date, 30 October 2009, is closer.
query_timegate('ia', 'http://discontents.com.au', date='2010-01-01')
# ----
# `GET` requests return different results if redirects are not followed.
query_timegate('ia', 'http://discontents.com.au', date='2010-01-01', request_type='get')
query_timegate('ia', 'http://discontents.com.au', date='2010-01-01', request_type='get', allow_redirects=False)
# ### UK Web Archive
#
# Changing whether or not redirects are followed has no effect on any of these responses.
#
# A query without an `Accept-Datetime` value doesn't return a `memento`.
query_timegate('ukwa', 'http://bl.uk')
# ----
#
# A query with an `Accept-Datetime` value of 1 January 2006 returns a `memento` from 1 January 2006. However, this date doesn't seem to represent an actual capture. There seems to be a problem with the Timegate.
query_timegate('ukwa', 'http://bl.uk', date='2006-01-01')
# ----
#
# A `GET` request returns the same results as a `HEAD` request.
query_timegate('ukwa', 'http://bl.uk', date='2006-01-01', request_type='get')
# ### Summarising the differences
#
# As you can see above, there are a couple of significant differences in the way that Timegates behave across the four repositories.
#
# * The Wayback systems (IA and NZWA) provide more information than the Pywb systems (`first memento`, `last memento`, `prev memento`, and `last memento`)
# * The UKWA and NZWA don't return a `memento` unless you include a date in the `Accept-Datetime` header. The NLA and IA return a recently captured `memento` as a default. (Though no necessarily the *most* recent?)
# * You can use either `HEAD` or `GET` with UKWA and NZWA, but IA and AWA behave different depending on the type of request and whether redirects are followed. To get results from either a `HEAD` or `GET` request, AWA requests should not follow redirects. To get results from a `HEAD` requests, IA requests should follow redirects. `GET` requests to IA will return results whether or not redirects are allowed, however, those results differ.
# ### Normalising Timegate responses and queries
#
# Here's some code to smooth out the differences between systems, and return Memento data as a Python dictionary. Specifically it:
#
# * Inserts the current date into requests from the UKWA or NLNZ if no date is specified. This means they behave like the other repositories that return a recent Memento.
# * Follows redirects for requests to the IA.
# * If there is no `memento` value in the response (as sometimes happens with NLNZ), it looks for a `first`, `last`, `prev` or `next` value instead.
# +
def query_timegate(timegate, url, date=None, tz='Australia/Canberra'):
'''
Query the specified repository for a Memento.
'''
headers = {}
if date:
formatted_date = format_date_for_headers(date, tz)
headers['Accept-Datetime'] = formatted_date
# BL & NLNZ don't seem to default to latest date if no date supplied
elif not date and timegate in ['bl', 'nlnz']:
formatted_date = format_date_for_headers(arrow.utcnow().format('YYYY-MM-DD'), tz)
headers['Accept-Datetime'] = formatted_date
# Note that you don't get a timegate response if you leave off the trailing slash, but extras don't hurt!
tg_url = f'{TIMEGATES[timegate]}{url}/' if not url.endswith('/') else f'{TIMEGATES[timegate]}{url}'
# print(tg_url)
# IA only works if redirects are followed -- this defaults to False with HEAD requests...
if timegate == 'ia':
allow_redirects = True
else:
allow_redirects = False
response = requests.head(tg_url, headers=headers, allow_redirects=allow_redirects)
return parse_links_from_headers(response)
def get_memento(timegate, url, date=None, tz='Australia/Canberra'):
'''
If there's no memento in the results, look for an alternative.
'''
links = query_timegate(timegate, url, date, tz)
# NLNZ doesn't always seem to return a Memento, so we'll build in some fuzziness
if links:
if 'memento' in links:
memento = links['memento']
elif 'prev memento' in links:
memento = links['prev memento']
elif 'next memento' in links:
memento = links['next memento']
elif 'last memento' in links:
memento = links['last memento']
else:
memento = None
return memento
# -
# Now we can request a Memento from any of the four repositories and get back the results as a Python dictionary. You can see this code in action in the [Get full page screenshots from archived web pages](save_screenshot.ipynb) notebook.
query_timegate('ukwa', 'http://bl.uk', date='2015-01-01')
# Or if we just want to get the url for a Memento (and fallback to alternative values if `memento` is missing).
get_memento('nzwa', 'http://natlib.govt.nz')
# ----
#
# ## Timemaps
#
# Memento Timemaps provide machine-processable lists of web page captures from a particular archive. They are available from both OpenWayback and Pywb systems, though there are some differences. The [Pywb documentation](https://pywb.readthedocs.io/en/latest/manual/memento.html#timemap-api) notes that the following formats are available:
#
# * link – returns an application/link-format as required by the Memento spec
# * cdxj – returns a timemap in the native CDXJ format
# * json – returns the timemap as newline-delimited JSON lines (NDJSON) format
#
# Timemaps are requested using a url with the following format:
#
# ```
# http://[address.of.archive]/[collection]/timemap/[format]/[web page url]
# ```
#
# So if you wanted to query the Australian Web Archive to get a list of captures in JSON format from http://nla.gov.au/ you'd use this url:
#
# ```
# https://web.archive.org.au/awa/timemap/json/http://nla.gov.au/
# ```
#
# The examples below show how the format and behaviour of Timemaps vary slightly across the four respoitories we're interested in.
def get_timemap(timegate, url, format='json'):
'''
Basic function to get a Timemap for the supplied url.
'''
tg_url = f'{TIMEGATES[timegate]}timemap/{format}/{url}/'
response = requests.get(tg_url)
# Show the content-type
print(response.headers['content-type'])
return response.text
# ### National Library of Australia
#
# Request a Timemap in `link` format. Note that response headers include `content-type` of `application/link-format`.
timemap = get_timemap('awa', 'http://www.gov.au', 'link')
# Show the first 5 lines
print('\n'.join(timemap.splitlines()[:5]))
# ----
#
# Request a Timemap in `json` format. This returns `ndjson` (Newline Delineated JSON) – each capture is a JSON object, separated by a line break. Note that the response headers include `content-type` of `text/x-ndjson`.
timemap = get_timemap('awa', 'http://www.aph.gov.au/Senate/committee/eet_ctte/uni_finances/report/index.htm', 'json')
# Show the first line
print('\n'.join(timemap.splitlines()[:1]))
# ----
#
# Request a Timemap in `cdxj` format. Note that response headers include `content-type` of `text/x-cdxj`.
timemap = get_timemap('awa', 'http://www.aph.gov.au/Senate/committee/eet_ctte/uni_finances/report/index.htm', 'cdxj')
# Show the first line
print('\n'.join(timemap.splitlines()[:1]))
# ### UK Web Archive
#
# Request a Timemap in `link` format. Note that response headers include `content-type` of `application/link-format`.
timemap = get_timemap('ukwa', 'http://bl.uk', 'link')
print('\n'.join(timemap.splitlines()[:5]))
# ----
#
# Request a Timemap in `json` format. This returns `ndjson` (Newline Delineated JSON) – each capture is a JSON object, separated by a line break. Note that the response headers include `content-type` of `text/x-ndjson`.
timemap = get_timemap('ukwa', 'http://bl.uk', 'json')
print('\n'.join(timemap.splitlines()[:1]))
# ----
#
# Request a Timemap in `cdxj` format. Note that response headers include `content-type` of `text/x-cdxj`.
timemap = get_timemap('ukwa', 'http://bl.uk', 'cdxj')
print('\n'.join(timemap.splitlines()[:1]))
# ### National Library of New Zealand
#
# Request a Timemap in `link` format. Note that response headers include `content-type` of `application/link-format`.
timemap = get_timemap('nzwa', 'http://natlib.govt.nz', 'link')
print('\n'.join(timemap.splitlines()[:5]))
# ----
#
# A request for a Timemap in `json` returns results in `link` format. OpenWayback only supports the `link` format.
timemap = get_timemap('nzwa', 'http://natlib.govt.nz', 'json')
print('\n'.join(timemap.splitlines()[:5]))
# ### Internet Archive
#
# Request a Timemap in `link` format. Note that response headers include `content-type` of `application/link-format`.
timemap = get_timemap('ia', 'http://discontents.com.au', 'link')
print('\n'.join(timemap.splitlines()[:5]))
# ----
#
# Request for timemap in `json` format returns results in JSON as an array of arrays, where the first row provides the column headings. Response headers include `content-type` of `application/json`.
timemap = get_timemap('ia', 'http://discontents.com.au', 'json')
print('\n'.join(timemap.splitlines()[:5]))
# ----
#
# Request for timemap in `cdxj` returns results in plain text, with fields separated by spaces, and captures separated by line breaks. Response headers include `content-type` of `text/plain`.
timemap = get_timemap('ia', 'http://discontents.com.au', 'cdxj')
print('\n'.join(timemap.splitlines()[:5]))
# ### Differences in field labels
# If we compare the Pywb JSON output with the IA Wayback output, we see there are also some differences in the field labels. In particular `original` in IA Wayback is just `url` in Pywb, while `statuscode` and `mimetype` are shortened to `status` and `mime` in Pywb.
timemap = get_timemap('ia', 'http://bl.uk', 'json')
data = json.loads(timemap)
data[0]
timemap = get_timemap('ukwa', 'http://bl.uk', 'json')
data = [json.loads(line) for line in timemap.splitlines()]
list(data[0].keys())
# ### Summarising the differences
#
# The good news is that all repositories provide Timemaps in the standard `link` format as required by the Memento specification. However, there's more varation when it comes to other formats.
#
# * NLNZ only provides the `link` format.
# * IA's `json` format is different to the Pywb format from UKWA and NLA.
# * IA uses different labels for some values.
# ### Normalising Timemaps
#
# With the information above we can construct some functions to return normalised Timemap results as JSON. To do this we need to:
#
# * Convert the `link` format from NLNZ to JSON
# * Restructure the JSON output from IA to match the Pywb format
# * Change some of the column headings in the IA data to match the Pywb format
#
# Because the `link` format provides less information than the `json` format, we could also try to enrich the NLNZ data by requesting more information about individual Mementos.
# +
def convert_lists_to_dicts(results):
'''
Converts IA style timemap (a JSON array of arrays) to a list of dictionaries.
Renames keys to standardise IA with other Timemaps.
'''
if results:
keys = results[0]
results_as_dicts = [dict(zip(keys, v)) for v in results[1:]]
else:
results_as_dicts = results
for d in results_as_dicts:
d['status'] = d.pop('statuscode')
d['mime'] = d.pop('mimetype')
d['url'] = d.pop('original')
return results_as_dicts
def get_capture_data_from_memento(url, request_type='head'):
'''
For OpenWayback systems this can get some extra capture info to insert into Timemaps.
'''
if request_type == 'head':
response = requests.head(url)
else:
response = requests.get(url)
headers = response.headers
length = headers.get('x-archive-orig-content-length')
status = headers.get('x-archive-orig-status')
status = status.split(' ')[0] if status else None
mime = headers.get('x-archive-orig-content-type')
mime = mime.split(';')[0] if mime else None
return {'length': length, 'status': status, 'mime': mime}
def convert_link_to_json(results, enrich_data=False):
'''
Converts link formatted Timemap to JSON.
'''
data = []
for line in results.splitlines():
parts = line.split('; ')
if len(parts) > 1:
link_type = re.search(r'rel="(original|self|timegate|first memento|last memento|memento)"', parts[1]).group(1)
if link_type == 'memento':
link = parts[0].strip('<>')
timestamp, original = re.search(r'/(\d{14})/(.*)$', link).groups()
capture = {'timestamp': timestamp, 'url': original}
if enrich_data:
capture.update(get_capture_data_from_memento(link))
print(capture)
data.append(capture)
return data
def get_timemap_as_json(timegate, url):
'''
Get a Timemap then normalise results (if necessary) to return a list of dicts.
'''
tg_url = f'{TIMEGATES[timegate]}timemap/json/{url}/'
response = requests.get(tg_url)
response_type = response.headers['content-type']
print(response_type)
if response_type == 'text/x-ndjson':
data = [json.loads(line) for line in response.text.splitlines()]
elif response_type == 'application/json':
data = convert_lists_to_dicts(response.json())
elif response_type in ['application/link-format', 'text/html;charset=utf-8']:
data = convert_link_to_json(response.text)
return data
# -
#
# Now we can get information about captures in a standardised JSON format from all four repositories. Although, we can't rely on NLNZ data having anything more than `timestamp` and `url` for each capture. You can see this in action in the [Display changes in the text of an archived web page over time](display-text-changes-from-timemap.ipynb) notebook
timemap = get_timemap_as_json('ukwa', 'http://bl.uk')
timemap[0]
timemap = get_timemap_as_json('ia', 'http://bl.uk')
timemap[0]
# ----
#
# ## Mementos
#
# You can also modify the url of a Memento to change the way it's presented. In particular, adding `id_` after the timestamp will tell the server that you want the original harvested version of the webpage, without any rewriting of links, or web archive navigation features. For example:
#
# ```
# https://web.archive.org.au/awa/20200302223537id_/http://discontents.com.au/
# ```
#
# This works with all four repositories, however, note that for the Australian Web Archive you need to use the `web.archive.org.au` domain, not `webarchive.nla.gov.au`.
#
# In addition, NLNZ and IA both support the `if_` option, which provides a view of the archived page without web archive headers navigation inserted, but with links to CSS, JS, and images rewritten to point to archived versions. This is as close as you can get to looking at the original page, and I've used it in the [Get full page screenshots from archived web pages](save_screenshot.ipynb) notebook. Note that if you add `if_` to requests from the UKWA or the NLA you'll be redirected to the standard view with the original page framed by the web archive navigation.
#
# Pywb's page on [url rewriting](https://pywb.readthedocs.io/en/latest/manual/rewriter.html?highlight=id_#url-rewriting) has some useful information about this.
#
# ----
# Created by [<NAME>](https://timsherratt.org) for the [GLAM Workbench](https://glam-workbench.github.io). Support me by becoming a [GitHub sponsor](https://github.com/sponsors/wragge)!
#
# Work on this notebook was supported by the [IIPC Discretionary Funding Programme 2019-2020](http://netpreserve.org/projects/)
|
memento.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# default_exp modeling.core
# +
# all_slow
# -
#hide
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# # modeling.core
#
# > This module contains core custom models, loss functions, and a default layer group splitter for use in applying discriminiative learning rates to your Hugging Face models trained via fastai
# +
# export
import os, inspect
from typing import Any, Callable, Dict, List, Optional, Union, Type
from fastcore.all import *
from fastai.callback.all import *
from fastai.data.block import DataBlock, ColReader, CategoryBlock, MultiCategoryBlock, ColSplitter, RandomSplitter
from fastai.data.core import DataLoader, DataLoaders, TfmdDL
from fastai.imports import *
from fastai.learner import *
from fastai.losses import CrossEntropyLossFlat
from fastai.optimizer import Adam, OptimWrapper, params
from fastai.metrics import accuracy, F1Score, accuracy_multi, F1ScoreMulti
from fastai.torch_core import *
from fastai.torch_imports import *
from fastprogress.fastprogress import progress_bar, master_bar
from transformers import AutoModelForSequenceClassification, logging, PretrainedConfig, PreTrainedTokenizerBase, PreTrainedModel
from blurr.utils import BLURR, set_seed
from blurr.data.core import HF_TextBlock, HF_BaseInput, first_blurr_tfm
logging.set_verbosity_error()
# +
# hide_input
import pdb
from fastai.data.external import untar_data, URLs
from fastcore.test import *
from nbverbose.showdoc import show_doc
from blurr.utils import print_versions
from blurr.data.core import BlurrDataLoader
os.environ["TOKENIZERS_PARALLELISM"] = "false"
print("What we're running with at the time this documentation was generated:")
print_versions("torch fastai transformers")
# -
# hide
# cuda
torch.cuda.set_device(1)
print(f"Using GPU #{torch.cuda.current_device()}: {torch.cuda.get_device_name()}")
# ## Base splitter, model wrapper, and model callback
# export
def hf_splitter(m: Module):
"""Splits the Hugging Face model based on various model architecture conventions"""
model = m.hf_model if (hasattr(m, "hf_model")) else m
root_modules = list(model.named_children())
top_module_name, top_module = root_modules[0]
groups = L([m for m_name, m in list(top_module.named_children())])
groups += L([m for m_name, m in root_modules[1:]])
return groups.map(params).filter(lambda el: len(el) > 0)
show_doc(hf_splitter)
# +
# export
class HF_BaseModelWrapper(Module):
def __init__(
self,
# Your Hugging Face model
hf_model: PreTrainedModel,
# If True, hidden_states will be returned and accessed from Learner
output_hidden_states: bool = False,
# If True, attentions will be returned and accessed from Learner
output_attentions: bool = False,
# Any additional keyword arguments you want passed into your models forward method
hf_model_kwargs={},
):
super().__init__()
store_attr(self=self, names="output_hidden_states, output_attentions, hf_model_kwargs")
self.hf_model = hf_model.cuda() if torch.cuda.is_available() else hf_model
self.hf_model_fwd_args = list(inspect.signature(self.hf_model.forward).parameters.keys())
def forward(self, x):
for k in list(x):
if k not in self.hf_model_fwd_args:
del x[k]
return self.hf_model(
**x,
output_hidden_states=self.output_hidden_states,
output_attentions=self.output_attentions,
return_dict=True,
**self.hf_model_kwargs
)
# -
# Note that `HF_BaseModelWrapper` includes some nifty code for just passing in the things your model needs, as not all transformer architectures require/use the same information.
# +
# export
class HF_PreCalculatedLoss:
def __call__(self, inp, targ, **kwargs):
return tensor(0.0)
def decodes(self, x):
return x.argmax(dim=-1)
def activation(self, x):
return F.softmax(x, dim=-1)
# -
# If you want to let your Hugging Face model calculate the loss for you, make sure you include the `labels` argument in your inputs and use `HF_PreCalculatedLoss` as your loss function. Even though we don't really need a loss function per se, we have to provide a custom loss class/function for fastai to function properly (e.g. one with a `decodes` and `activation` methods). Why? Because these methods will get called in methods like `show_results` to get the actual predictions.
# export
class HF_BaseModelCallback(Callback):
def before_batch(self):
self.hf_loss = None
def after_pred(self):
model_outputs = self.pred
self.learn.blurr_model_outputs = {}
for k, v in model_outputs.items():
# if the "labels" are included, we are training with target labels in which case the loss is returned
if k == "loss" and isinstance(self.learn.loss_func, HF_PreCalculatedLoss):
self.hf_loss = to_float(v)
# the logits represent the prediction
elif k == "logits":
self.learn.pred = v
# add any other things included in model_outputs as blurr_{model_output_key}
else:
self.learn.blurr_model_outputs[k] = v
def after_loss(self):
# if we already have the loss from the model, update the Learner's loss to be it
if self.hf_loss is not None:
self.learn.loss_grad = self.hf_loss
self.learn.loss = self.learn.loss_grad.clone()
# We use a `Callback` for handling what is returned from the Hugging Face model. The return type is (`ModelOutput`)[https://huggingface.co/transformers/main_classes/output.html#transformers.file_utils.ModelOutput] which makes it easy to return all the goodies we asked for.
#
# Note that your `Learner`'s loss will be set for you only if the Hugging Face model returns one *and* you are using the `HF_PreCalculatedLoss` loss function.
#
# Also note that anything else you asked the model to return (for example, last hidden state, etc..) will be available for you via the `blurr_model_outputs` property attached to your `Learner`. For example, assuming you are using BERT for a classification task ... if you have told your `HF_BaseModelWrapper` instance to return attentions, you'd be able to access them via `learn.blurr_model_outputs['attentions']`.
# ## Sequence classification
#
# Below demonstrates how to setup your `blurr` pipeline for a sequence classification task (e.g., a model that requires a single text input) using the mid, high, and low-level API
# ### Using the mid-level API
# +
path = untar_data(URLs.IMDB_SAMPLE)
imdb_df = pd.read_csv(path / "texts.csv")
# -
imdb_df.head()
# +
# hide_output
model_cls = AutoModelForSequenceClassification
pretrained_model_name = "distilroberta-base" # "distilbert-base-uncased" "bert-base-uncased"
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(pretrained_model_name, model_cls=model_cls)
# +
# single input
set_seed()
blocks = (HF_TextBlock(hf_arch, hf_config, hf_tokenizer, hf_model), CategoryBlock)
dblock = DataBlock(blocks=blocks, get_x=ColReader("text"), get_y=ColReader("label"), splitter=RandomSplitter(seed=42))
# -
# hide
# dblock.summary(imdb_df)
dls = dblock.dataloaders(imdb_df, bs=4)
# dls.show_batch(dataloaders=dls, max_n=2, trunc_at=500)
# #### Training
#
# We'll also add in custom summary methods for blurr learners/models that work with dictionary inputs
# +
set_seed()
model = HF_BaseModelWrapper(hf_model)
learn = Learner(
dls,
model,
opt_func=partial(OptimWrapper, opt=torch.optim.Adam),
loss_func=CrossEntropyLossFlat(),
metrics=[accuracy],
cbs=[HF_BaseModelCallback],
splitter=hf_splitter,
)
learn.freeze()
# -
# `.to_fp16()` requires a GPU so had to remove for tests to run on github. Let's check that we can get predictions.
# hide_output
# learn.summary()
# print(len(learn.opt.param_groups))
# learn.lr_find(suggest_funcs=[minimum, steep, valley, slide])
set_seed()
learn.fit_one_cycle(1, lr_max=1e-3)
# epoch train_loss valid_loss accuracy time
# 0 0.324516 0.294210 0.885000 00:11
# #### Showing results
#
# And here we create a @typedispatched implementation of `Learner.show_results`.
# export
@typedispatch
def show_results(
# This typedispatched `show_results` will be called for `HF_BaseInput` typed inputs
x: HF_BaseInput,
# Your targets
y,
# Your raw inputs/targets
samples,
# The model's predictions
outs,
# Your `Learner`. This is required so as to get at the Hugging Face objects for decoding them into
# something understandable
learner,
# Your `show_results` context
ctxs=None,
# The maximum number of items to show
max_n=6,
# Any truncation your want applied to your decoded inputs
trunc_at=None,
# Any other keyword arguments you want applied to `show_results`
**kwargs,
):
# grab our tokenizer
tfm = first_blurr_tfm(learner.dls)
hf_tokenizer = tfm.hf_tokenizer
trg_labels = None
if hasattr(learner.dls, "label_names"):
trg_labels = learner.dls.label_names
res = L()
n_inp = learner.dls.n_inp
for idx, (input_ids, label, pred, sample) in enumerate(zip(x, y, outs, samples)):
if idx >= max_n:
break
# add in the input text
rets = [hf_tokenizer.decode(input_ids, skip_special_tokens=True)[:trunc_at]]
# add in the targets
for item in sample[n_inp:]:
if not torch.is_tensor(item):
trg = item
elif is_listy(item.tolist()):
trg = [trg_labels[idx] for idx, val in enumerate(label.numpy().tolist()) if (val == 1)] if (trg_labels) else label.item()
else:
trg = trg_labels[label.item()] if (trg_labels) else label.item()
rets.append(trg)
# add in the predictions
for item in pred:
if not torch.is_tensor(item):
p = item
elif is_listy(item.tolist()):
p = [trg_labels[idx] for idx, val in enumerate(item.numpy().tolist()) if (val == 1)] if (trg_labels) else item.item()
else:
p = trg_labels[item.item()] if (trg_labels) else item.item()
rets.append(p)
res.append(tuplify(rets))
cols = ["text"] + ["target" if (i == 0) else f"target_{i}" for i in range(len(res[0]) - n_inp * 2)]
cols += ["prediction" if (i == 0) else f"prediction_{i}" for i in range(len(res[0]) - n_inp * 2)]
display_df(pd.DataFrame(res, columns=cols)[:max_n])
return ctxs
learn.show_results(learner=learn, max_n=2, trunc_at=500)
# export
@patch
def blurr_predict(self: Learner, items, rm_type_tfms=None):
# grab our blurr tfm with the bits to properly decode/show our inputs/targets
tfm = first_blurr_tfm(self.dls)
is_split_str = tfm.is_split_into_words and isinstance(items[0], str)
is_df = isinstance(items, pd.DataFrame)
if not is_df and (is_split_str or not is_listy(items)):
items = [items]
dl = self.dls.test_dl(items, rm_type_tfms=rm_type_tfms, num_workers=0)
with self.no_bar():
probs, _, decoded_preds = self.get_preds(dl=dl, with_input=False, with_decoded=True)
trg_tfms = self.dls.tfms[self.dls.n_inp :]
outs = []
probs, decoded_preds = L(probs), L(decoded_preds)
for i in range(len(items)):
item_probs = probs.itemgot(i)
item_dec_preds = decoded_preds.itemgot(i)
item_dec_labels = tuplify([tfm.decode(item_dec_preds[tfm_idx]) for tfm_idx, tfm in enumerate(trg_tfms)])
outs.append((item_dec_labels, item_dec_preds, item_probs))
return outs
show_doc(Learner.blurr_predict)
# We need to replace fastai's `Learner.predict` method with the one above which is able to work with inputs that are represented by multiple tensors included in a dictionary.
# +
learn.blurr_predict("I really liked the movie")
# +
learn.blurr_predict(["I really liked the movie", "I really hated the movie"])
# -
# Though not useful in sequence classification, we will also add a `blurr_generate` method to `Learner` that uses Hugging Face's `PreTrainedModel.generate` for text generation tasks.
#
# For the full list of arguments you can pass in see [here](https://huggingface.co/transformers/main_classes/model.html#transformers.PreTrainedModel.generate). You can also check out their ["How To Generate"](https://github.com/huggingface/blog/blob/master/notebooks/02_how_to_generate.ipynb) notebook for more information about how it all works.
# export
@patch
def blurr_generate(self: Learner, inp, **kwargs):
"""Uses the built-in `generate` method to generate the text
(see [here](https://huggingface.co/transformers/main_classes/model.html#transformers.PreTrainedModel.generate)
for a list of arguments you can pass in)
"""
# grab our blurr tfm with the bits to properly decode/show our inputs/targets
tfm = first_blurr_tfm(self.dls)
# grab the Hugging Face tokenizer from the learner's dls.tfms
hf_config = tfm.hf_config
hf_tokenizer = tfm.hf_tokenizer
tok_kwargs = tfm.tok_kwargs
# grab the text generation kwargs
text_gen_kwargs = tfm.text_gen_kwargs if (len(kwargs) == 0) else kwargs
if isinstance(inp, str):
input_ids = hf_tokenizer.encode(inp, padding=True, truncation=True, return_tensors="pt", **tok_kwargs)
else:
# note (10/30/2020): as of pytorch 1.7, this has to be a plain ol tensor (not a subclass of TensorBase)
input_ids = inp.as_subclass(Tensor)
input_ids = input_ids.to(self.model.hf_model.device)
gen_texts = self.model.hf_model.generate(input_ids, **text_gen_kwargs)
outputs = [hf_tokenizer.decode(txt, skip_special_tokens=True, clean_up_tokenization_spaces=False) for txt in gen_texts]
if tfm.hf_arch == "pegasus":
outputs = [o.replace("<n>", " ") for o in outputs]
return outputs
show_doc(Learner.blurr_generate)
learn.unfreeze()
set_seed()
learn.fit_one_cycle(2, lr_max=slice(1e-7, 1e-4))
# epoch train_loss valid_loss accuracy time
# 0 0.263290 0.272322 0.895000 00:18
# 1 0.218568 0.263317 0.910000 00:18
learn.recorder.plot_loss()
learn.show_results(learner=learn, max_n=2, trunc_at=500)
learn.blurr_predict("This was a really good movie")
learn.blurr_predict("Acting was so bad it was almost funny.")
# #### Inference
# +
export_fname = "seq_class_learn_export"
# -
# #### Using fast.ai `Learner.export` and `load_learner`
learn.export(fname=f"{export_fname}.pkl")
inf_learn = load_learner(fname=f"{export_fname}.pkl")
inf_learn.blurr_predict("This movie should not be seen by anyone!!!!")
# hide
try:
del learn
del inf_learn
torch.cuda.empty_cache()
except:
pass
# ### Using the high-level API
# #### Blearner
# Instead of constructing our low-level `Learner`, we can use the `Blearner` class which provides sensible defaults for training
# +
# hide_output
model_cls = AutoModelForSequenceClassification
pretrained_model_name = "distilroberta-base" # "distilbert-base-uncased" "bert-base-uncased"
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(pretrained_model_name, model_cls=model_cls)
dls = dblock.dataloaders(imdb_df, bs=4)
# -
# export
@delegates(Learner.__init__)
class Blearner(Learner):
def __init__(
self,
# Your fast.ai DataLoaders
dls: DataLoaders,
# Your pretrained Hugging Face transformer
hf_model: PreTrainedModel,
# Your `HF_BaseModelCallback`
base_model_cb: HF_BaseModelCallback = HF_BaseModelCallback,
# Any kwargs you want to pass to your `BLearner`
**kwargs
):
model = kwargs.get("model", HF_BaseModelWrapper(hf_model))
loss_func = kwargs.pop("loss_func", dls.loss_func if hasattr(dls, "loss_func") else None)
splitter = kwargs.pop("splitter", hf_splitter)
super().__init__(dls, model=model, loss_func=loss_func, splitter=splitter, **kwargs)
self.add_cb(base_model_cb)
self.freeze()
learn = Blearner(dls, hf_model, metrics=[accuracy])
learn.fit_one_cycle(1, lr_max=1e-3)
learn.show_results(learner=learn, max_n=2, trunc_at=500)
learn.blurr_predict("This was a really good movie")
learn.export(fname=f"{export_fname}.pkl")
inf_learn = load_learner(fname=f"{export_fname}.pkl")
inf_learn.blurr_predict("This movie should not be seen by anyone!!!!")
# #### BlearnerForSequenceClassification
# We also introduce a task specific `Blearner` that get you your DataBlock, DataLoaders, and BLearner in one line of code!
# +
# hide
try:
del learn
del inf_learn
torch.cuda.empty_cache()
except:
pass
# +
# export
@delegates(Blearner.__init__)
class BlearnerForSequenceClassification(Blearner):
def __init__(self, dls: DataLoaders, hf_model: PreTrainedModel, **kwargs):
super().__init__(dls, hf_model, **kwargs)
@classmethod
def get_model_cls(self):
return AutoModelForSequenceClassification
@classmethod
def _get_x(cls, r, attr):
return r[attr] if (isinstance(attr, str)) else tuple(r[inp] for inp in attr)
@classmethod
def _get_y(cls, r, attr):
return r[attr] if (isinstance(attr, str)) else [r[inp] for inp in attr]
@classmethod
def _create_learner(
cls,
# Your raw dataset
data,
# The name or path of the pretrained model you want to fine-tune
pretrained_model_name_or_path: Optional[Union[str, os.PathLike]],
# A function to perform any preprocessing required for your Dataset
preprocess_func: Callable = None,
# The attribute in your dataset that contains your raw text
text_attr: str = "text",
# The attribute in your dataset that contains your labels/targets
label_attr: str = "label",
# The number of labels/classes your model should predict
n_labels: int = 2,
# A function that will split your Dataset into a training and validation set
# See [here](https://docs.fast.ai/data.transforms.html#Split) for a list of fast.ai splitters
dblock_splitter: Callable = RandomSplitter(),
# Any kwargs to pass to your `DataLoaders`
dl_kwargs={},
# Any kwargs to pass to your task specific `Blearner`
learner_kwargs={},
):
# get our hf objects
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(
pretrained_model_name_or_path, model_cls=cls.get_model_cls(), config_kwargs={"num_labels": n_labels}
)
# if we need to preprocess the raw data before creating our DataLoaders
if preprocess_func:
data = preprocess_func(data, hf_arch, hf_config, hf_tokenizer, hf_model, text_attr, label_attr)
# not all architectures include a native pad_token (e.g., gpt2, ctrl, etc...), so we add one here
if hf_tokenizer.pad_token is None:
hf_tokenizer.add_special_tokens({"pad_token": "<pad>"})
hf_config.pad_token_id = hf_tokenizer.get_vocab()["<pad>"]
hf_model.resize_token_embeddings(len(hf_tokenizer))
# defin our input/target getters
if isinstance(data, pd.DataFrame):
get_x = ColReader(text_attr)
get_y = ColReader(label_attr)
else:
get_x = partial(cls._get_x, attr=text_attr)
get_y = partial(cls._get_y, attr=label_attr)
# infer loss function and default metrics
if is_listy(label_attr):
trg_block = MultiCategoryBlock(encoded=True, vocab=label_attr)
learner_kwargs["metrics"] = learner_kwargs.get("metrics", [F1ScoreMulti(), accuracy_multi])
else:
trg_block = CategoryBlock
learner_kwargs["metrics"] = learner_kwargs.get("metrics", [F1Score(), accuracy])
# build our DataBlock and DataLoaders
blocks = (HF_TextBlock(hf_arch, hf_config, hf_tokenizer, hf_model), trg_block)
dblock = DataBlock(blocks=blocks, get_x=get_x, get_y=get_y, splitter=dblock_splitter)
dls = dblock.dataloaders(data, **dl_kwargs.copy())
# return BLearner instance
return cls(dls, hf_model, **learner_kwargs.copy())
@classmethod
def from_dataframe(
cls,
# Your pandas DataFrame
df: pd.DataFrame,
# The name or path of the pretrained model you want to fine-tune
pretrained_model_name_or_path: Optional[Union[str, os.PathLike]],
# A function to perform any preprocessing required for your Dataset
preprocess_func: Callable = None,
# The attribute in your dataset that contains your raw text
text_attr: str = "text",
# The attribute in your dataset that contains your labels/targets
label_attr: str = "label",
# The number of labels/classes your model should predict
n_labels: int = None,
# A function that will split your Dataset into a training and validation set
# See [here](https://docs.fast.ai/data.transforms.html#Split) for a list of fast.ai splitters
dblock_splitter: Callable = ColSplitter(),
# Any kwargs to pass to your `DataLoaders`
dl_kwargs={},
# Any kwargs to pass to your task specific `Blearner`
learner_kwargs={},
):
# we need to tell transformer how many labels/classes to expect
if n_labels is None:
n_labels = len(label_attr) if (is_listy(label_attr)) else len(df[label_attr].unique())
return cls._create_learner(
df, pretrained_model_name_or_path, preprocess_func, text_attr, label_attr, n_labels, dblock_splitter, dl_kwargs, learner_kwargs
)
@classmethod
def from_csv(
cls,
# The path to your csv file
csv_file: Union[Path, str],
# The name or path of the pretrained model you want to fine-tune
pretrained_model_name_or_path: Optional[Union[str, os.PathLike]],
# A function to perform any preprocessing required for your Dataset
preprocess_func: Callable = None,
# The attribute in your dataset that contains your raw text
text_attr: str = "text",
# The attribute in your dataset that contains your labels/targets
label_attr: str = "label",
# The number of labels/classes your model should predict
n_labels: int = None,
# A function that will split your Dataset into a training and validation set
# See [here](https://docs.fast.ai/data.transforms.html#Split) for a list of fast.ai splitters
dblock_splitter: Callable = ColSplitter(),
# Any kwargs to pass to your `DataLoaders`
dl_kwargs={},
# Any kwargs to pass to your task specific `Blearner`
learner_kwargs={},
):
df = pd.read_csv(csv_file)
return cls.from_dataframe(
df,
pretrained_model_name_or_path=pretrained_model_name_or_path,
preprocess_func=preprocess_func,
text_attr=text_attr,
label_attr=label_attr,
n_labels=n_labels,
dblock_splitter=dblock_splitter,
dl_kwargs=dl_kwargs,
learner_kwargs=learner_kwargs,
)
@classmethod
def from_dictionaries(
cls,
# A list of dictionaries
ds: List[Dict],
# The name or path of the pretrained model you want to fine-tune
pretrained_model_name_or_path: Optional[Union[str, os.PathLike]],
# A function to perform any preprocessing required for your Dataset
preprocess_func: Callable = None,
# The attribute in your dataset that contains your raw text
text_attr: str = "text",
# The attribute in your dataset that contains your labels/targets
label_attr: str = "label",
# The number of labels/classes your model should predict
n_labels: int = None,
# A function that will split your Dataset into a training and validation set
# See [here](https://docs.fast.ai/data.transforms.html#Split) for a list of fast.ai splitters
dblock_splitter: Callable = RandomSplitter(),
# Any kwargs to pass to your `DataLoaders`
dl_kwargs={},
# Any kwargs to pass to your task specific `Blearner`
learner_kwargs={},
):
# we need to tell transformer how many labels/classes to expect
if n_labels is None:
n_labels = len(label_attr) if (is_listy(label_attr)) else len(set([item[label_attr] for item in ds]))
return cls._create_learner(
ds, pretrained_model_name_or_path, preprocess_func, text_attr, label_attr, n_labels, dblock_splitter, dl_kwargs, learner_kwargs
)
# +
learn = BlearnerForSequenceClassification.from_dataframe(
imdb_df, "distilroberta-base", text_attr="text", label_attr="label", dl_kwargs={"bs": 4}
)
# -
learn.fit_one_cycle(1, lr_max=1e-3)
learn.show_results(learner=learn, max_n=2, trunc_at=500)
learn.blurr_predict("This was a really good movie")
learn.export(fname=f"{export_fname}.pkl")
inf_learn = load_learner(fname=f"{export_fname}.pkl")
inf_learn.blurr_predict("This movie should not be seen by anyone!!!!")
# ### Using the low-level API
# Thanks to the `BlurrDataLoader`, there isn't really anything you have to do to use plain ol' PyTorch or fast.ai `Dataset`s and `DataLoaders` with Blurr. Let's take a look at fine-tuning a model against Glue's MRPC dataset ...
# +
from datasets import load_dataset
from blurr.data.core import preproc_hf_dataset
raw_datasets = load_dataset("glue", "mrpc")
# +
def tokenize_function(example):
return hf_tokenizer(example["sentence1"], example["sentence2"], truncation=True)
tokenized_datasets = raw_datasets.map(tokenize_function, batched=True)
# +
label_names = raw_datasets["train"].features["label"].names
trn_dl = BlurrDataLoader(
tokenized_datasets["train"],
hf_arch=hf_arch,
hf_config=hf_config,
hf_tokenizer=hf_tokenizer,
hf_model=hf_model,
preproccesing_func=preproc_hf_dataset,
label_names=label_names,
shuffle=True,
batch_size=8,
)
val_dl = BlurrDataLoader(
tokenized_datasets["validation"],
hf_arch=hf_arch,
hf_config=hf_config,
hf_tokenizer=hf_tokenizer,
hf_model=hf_model,
preproccesing_func=preproc_hf_dataset,
label_names=label_names,
batch_size=16,
)
dls = DataLoaders(trn_dl, val_dl)
# -
# And with our fast.ai `DataLoaders` in hand, we can train our model's using the high or low-level Blurr API. The `BlurrDataLoader` class sets up everything so that we can use our objects just as if we built our `DataLoaders` with the mid-level `DataBlock` API. This means we get back methods like `one_batch`, `show_batch`, `show_results`, etc... with all levels of Blurr's API.
learn = BlearnerForSequenceClassification(dls, hf_model, loss_func=CrossEntropyLossFlat())
learn.lr_find()
learn.fit_one_cycle(1, lr_max=1e-5)
learn.unfreeze()
learn.fit_one_cycle(2, lr_max=slice(1e-8, 1e-6))
learn.show_results(learner=learn, max_n=2, trunc_at=500)
# ## Tests
#
# The tests below to ensure the core training code above works for **all** pretrained sequence classification models available in Hugging Face. These tests are excluded from the CI workflow because of how long they would take to run and the amount of data that would be required to download.
#
# **Note**: Feel free to modify the code below to test whatever pretrained classification models you are working with ... and if any of your pretrained sequence classification models fail, please submit a github issue *(or a PR if you'd like to fix it yourself)*
# +
# hide
try:
del learn
del inf_learn
torch.cuda.empty_cache()
except:
pass
# +
# hide
[model_type for model_type in BLURR.get_models(task="SequenceClassification") if (not model_type.startswith("TF"))]
# -
# hide
pretrained_model_names = [
"albert-base-v1",
"facebook/bart-base",
"bert-base-uncased",
"google/bigbird-roberta-base",
"sshleifer/tiny-ctrl",
"camembert-base",
"sarnikowski/convbert-medium-small-da-cased",
"microsoft/deberta-base",
"microsoft/deberta-v2-xlarge",
"distilbert-base-uncased",
"monologg/electra-small-finetuned-imdb",
"flaubert/flaubert_small_cased",
"huggingface/funnel-small-base",
"gpt2",
"kssteven/ibert-roberta-base",
"allenai/led-base-16384",
"microsoft/layoutlm-base-uncased",
"allenai/longformer-base-4096",
"sshleifer/tiny-mbart",
"microsoft/mpnet-base",
"google/mobilebert-uncased",
"openai-gpt",
#'reformer-enwik8', # (see model card; does not work with/require a tokenizer so no bueno here)
"roberta-base",
"squeezebert/squeezebert-uncased",
#'google/tapas-base', # (requires pip install torch-scatter)
"transfo-xl-wt103",
"xlm-mlm-en-2048",
"xlm-roberta-base",
"xlnet-base-cased",
]
# +
# hide
path = untar_data(URLs.IMDB_SAMPLE)
model_path = Path("models")
imdb_df = pd.read_csv(path / "texts.csv")
# +
# hide
from transformers import RobertaTokenizer
model_cls = AutoModelForSequenceClassification
bsz = 2
seq_sz = 32
test_results = []
for model_name in pretrained_model_names:
error = None
print(f"=== {model_name} ===\n")
# 1. get/configure our Hugging Face objects
tok_class = RobertaTokenizer if ("/ibert" in model_name) else None
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(
model_name, model_cls=model_cls, tokenizer_cls=tok_class, config_kwargs={"num_labels": 2}
)
print(f"architecture:\t{hf_arch}\ntokenizer:\t{type(hf_tokenizer).__name__}\nmodel:\t\t{type(hf_model).__name__}\n")
# not all architectures include a native pad_token (e.g., gpt2, ctrl, etc...), so we add one here
if hf_tokenizer.pad_token is None:
hf_tokenizer.add_special_tokens({"pad_token": "<pad>"})
hf_config.pad_token_id = hf_tokenizer.get_vocab()["<pad>"]
hf_model.resize_token_embeddings(len(hf_tokenizer))
# 2. get our DataLoaders
blocks = (HF_TextBlock(hf_arch, hf_config, hf_tokenizer, hf_model, max_length=seq_sz, padding="max_length"), CategoryBlock)
dblock = DataBlock(blocks=blocks, get_x=ColReader("text"), get_y=ColReader("label"), splitter=ColSplitter(col="is_valid"))
dls = dblock.dataloaders(imdb_df, bs=bsz)
# 3. configure our Learner
model = HF_BaseModelWrapper(hf_model)
learn = Learner(
dls,
model,
opt_func=partial(Adam),
loss_func=CrossEntropyLossFlat(),
metrics=[accuracy],
cbs=[HF_BaseModelCallback],
splitter=hf_splitter,
)
learn.freeze()
b = dls.one_batch()
# 4. train
try:
print("*** TESTING DataLoaders ***")
test_eq(len(b), bsz)
test_eq(len(b[0]["input_ids"]), bsz)
test_eq(b[0]["input_ids"].shape, torch.Size([bsz, seq_sz]))
test_eq(len(b[1]), bsz)
# print('*** TESTING One pass through the model ***')
# preds = learn.model(b[0])
# test_eq(len(preds[0]), bsz)
# test_eq(preds[0].shape, torch.Size([bsz, 2]))
print("*** TESTING Training/Results ***")
learn.fit_one_cycle(1, lr_max=1e-3, cbs=ShortEpochCallback(pct=0.2, short_valid=True))
test_results.append((hf_arch, type(hf_tokenizer).__name__, type(hf_model).__name__, "PASSED", ""))
learn.show_results(learner=learn, max_n=2, trunc_at=250)
except Exception as err:
test_results.append((hf_arch, type(hf_tokenizer).__name__, type(hf_model).__name__, "FAILED", err))
finally:
# cleanup
del learn
torch.cuda.empty_cache()
# -
# hide_input
test_results_df = pd.DataFrame(test_results, columns=["arch", "tokenizer", "model", "result", "error"])
display_df(test_results_df)
# ## Summary
# This module includes the fundamental building blocks for training using Blurr
# +
# hide
from nbdev.export import notebook2script
notebook2script()
# -
|
nbs/01_modeling-core.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
s = 1
d = 0
n_ex = 1
out_rows = 7
out_cols = 7
out_ch = 5
fr = 3
fc = 3
in_rows = out_rows + fr - 1
in_cols = out_cols + fc - 1
in_ch = 3
# simple test 1
# W = (np.array(range(fr*fc*in_ch*out_ch)) + 1).reshape(fr, fc, in_ch, out_ch)
# X = (np.array(range(n_ex*in_rows*in_cols*in_ch)) + 1).reshape(n_ex, in_rows, in_cols, in_ch)
# dZ = (np.array(range(n_ex*out_rows*out_cols*out_ch)) + 1).reshape(n_ex, out_rows, out_cols, out_ch)
# simple test 2
# W = np.ones(fr*fc*in_ch*out_ch).reshape(fr, fc, in_ch, out_ch)
# X = 2*np.ones(n_ex*in_rows*in_cols*in_ch).reshape(n_ex, in_rows, in_cols, in_ch)
# dZ = np.ones(n_ex*out_rows*out_cols*out_ch).reshape(n_ex, out_rows, out_cols, out_ch)
# random int test
W = np.random.randint(10, size=(fr*fc*in_ch*out_ch)).reshape(fr, fc, in_ch, out_ch)
X = np.random.randint(10, size=(n_ex*in_rows*in_cols*in_ch)).reshape(n_ex, in_rows, in_cols, in_ch)
dZ = np.random.randint(10, size=(n_ex*out_rows*out_cols*out_ch)).reshape(n_ex, out_rows, out_cols, out_ch)
# real test
# W = np.random.rand(fr*fc*in_ch*out_ch).reshape(fr, fc, in_ch, out_ch)
# X = np.random.rand(n_ex*in_rows*in_cols*in_ch).reshape(n_ex, in_rows, in_cols, in_ch)
# dZ = np.random.rand(n_ex*out_rows*out_cols*out_ch).reshape(n_ex, out_rows, out_cols, out_ch)
# -
# filter 0
print(W[:, :, 0, 0])
print(W[:, :, 1, 0])
# filter 1
print(W[:, :, 0, 1])
print(W[:, :, 1, 1])
# ifmap
print(X[0, :, :, 0])
print(X[0, :, :, 1])
# upstream gradients
print(dZ[0, :, :, 0])
print(dZ[0, :, :, 1])
# +
# ground truth placeholder
dW = np.zeros_like(W)
dX = np.zeros_like(X)
for m in range(n_ex):
for i in range(out_rows):
for j in range(out_cols):
for c in range(out_ch):
# compute window boundaries w. stride and dilation
i0, i1 = i * s, (i * s) + fr * (d + 1) - d
j0, j1 = j * s, (j * s) + fc * (d + 1) - d
wc = W[:, :, :, c]
kernel = dZ[m, i, j, c]
window = X[m, i0 : i1 : (d + 1), j0 : j1 : (d + 1), :]
# dB[:, :, :, c] += kernel
dW[:, :, :, c] += window * kernel
dX[m, i0 : i1 : (d + 1), j0 : j1 : (d + 1), :] += wc * kernel
# -
# reproducing with deep loop nest
N = n_ex
C = in_ch
K = out_ch
P = out_cols
Q = out_rows
R = fc
S = fr
(N, C, K, P, Q, R, S)
# +
# test placeholder
test_dW = np.zeros_like(W)
test_dX = np.zeros_like(X)
# gradient wrt filters
for n in range(N):
for c in range(C):
for k in range(K):
for p in range(P):
for q in range(Q):
for r in range(R):
for s in range(S):
test_dW[s, r, c, k] += X[n, q+s, p+r, c] * dZ[n, q, p, k]
# emmm numerical issue with FP...
dW - test_dW
# +
# gradient wrt to activations
# error gradient map need to be padded
# npad is a tuple of (n_before, n_after) for each dimension
npad = ((0, 0), (fr-1, fr-1), (fc-1, fc-1), (0, 0))
padded_dZ = np.pad(dZ, pad_width=npad, mode='constant', constant_values=0)
# print(padded_dZ[0, :, :, 0])
# print(padded_dZ[0, :, :, 1])
# generate spatially rotated (180 degree) weight
spat_W = np.zeros_like(W)
for i in range(in_ch):
for j in range(out_ch):
for s in range(fr):
for r in range(fc):
spat_W[fr-1-s, fc-1-r, i, j] = W[s, r, i, j]
print(W[:, :, 0, 0])
# print(W[:, :, 1, 0])
print(spat_W[:, :, 0, 0])
# print(spat_W[:, :, 1, 0])
# +
test_dX = np.zeros_like(X)
for n in range(N):
for c in range(C):
for k in range(K):
for pr in range(P+R-1):
for qs in range(Q+S-1):
for r in range(R):
for s in range(S):
# test_dX[n, qs, pr, c] += padded_dZ[n, qs+s, pr+r, k] * spat_W[s, r, c, k]
test_dX[n, qs, pr, c] += padded_dZ[n, qs+s, pr+r, k] * W[S-1-s, R-1-r, c, k]
# emmm numerical issue with FP...
dX - test_dX
# -
# gradient wrt filter 0
print(dW[:, :, 0, 0])
print(dW[:, :, 1, 0])
# gradient wrt filter 0
print(dW[:, :, 0, 1])
print(dW[:, :, 1, 1])
# gradient wrt ifmap
print(dX[0, :, :, 0])
print(dX[0, :, :, 1])
|
problem-shapes/conv_backprop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy.polynomial.polynomial as poly
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
import matplotlib
import seaborn as sns
matplotlib.rcParams['figure.figsize'] = [12.0, 8.0]
# ### 1. Show the first 5 rows of the dataset
df = pd.read_csv('./e-customers.csv')
df.head(5)
# ### 2. Show the description and the info on the dataset, using appropriate Pandas functions
df.info()
df.describe()
# +
# set up some manageable variables
asl = df['Avg. Session Length']
ta = df['Time on App']
tw = df['Time on Website']
lom = df['Length of Membership']
yas = df['Yearly Amount Spent']
evs = df[['Avg. Session Length', 'Time on App', 'Time on Website','Length of Membership']]
# -
# ### 3. Use seaborn jointplot function to see which fields correlate well with the "Yearly Amount Spent" column. Write your findings.
sns.jointplot(yas, lom)
# #### Length of Membership = Positive Correlation of 0.81, great fit
sns.jointplot(yas, ta)
# #### Time on App = Positive correlation of 0.5, good fit
sns.jointplot(yas, asl)
# #### Ave. Session Length = Positive correlation of 0.36, okay fit
sns.jointplot(yas, tw)
# #### Time on Website = Negative correlation of -.0026, bad fit
# ### 4. Make a summary plot of feature relationships using pairplot function of Seaborn
sns.pairplot(df)
# ### 5. Create two dataframes: one for the target variable ("Yearly Amount Spent"), the other - containing all the rest of numerical features
# +
# set up some manageable variables
asl = df['Avg. Session Length']
ta = df['Time on App']
tw = df['Time on Website']
lom = df['Length of Membership']
yas = df['Yearly Amount Spent']
evs = df[['Avg. Session Length', 'Time on App', 'Time on Website','Length of Membership']]
# -
# ### 6. Split the data into a training and test sets. Make a test set size 0.3 and random seed 123 (so the results are consistent for all students)
# +
X_full = yas
Y_full = evs
X_train, X_test, Y_train, Y_test = train_test_split(X_full, Y_full, test_size=0.3, random_state=123)
# -
# ### 7&8. Fit a regression model on the training set
# #### Print out the coefficients of the model
plt.plot(X_train, Y_train, ".", markersize=5)
plt.title("Train set")
# +
X_full = yas
Y_full = lom
X_train, X_test, Y_train, Y_test = train_test_split(X_full, Y_full, test_size=0.3, random_state=123)
# -
coefs_lin_lom = poly.polyfit(X_train, Y_train, 1)
coefs_sqr_lom = poly.polyfit(X_train, Y_train, 2)
coefs_cube_lom = poly.polyfit(X_train, Y_train, 3)
# +
X_line = np.linspace(min(X_train - 10), max(X_train + 10), 200, True)
ffit_lin = poly.polyval(X_line, coefs_lin_lom)
ffit_sqr = poly.polyval(X_line, coefs_sqr_lom)
ffit_cube = poly.polyval(X_line, coefs_cube_lom)
# +
plt.ylim(min(Y_train - 5), max(Y_train + 5))
plt.plot(X_line, ffit_lin, "g")
plt.plot(X_line, ffit_sqr, "k")
plt.plot(X_line, ffit_cube, "b")
plt.plot(X_train, Y_train, "r.", markersize=5)
plt.show()
print('Linear Coeff = ', coefs_lin_lom)
# +
X_full = yas
Y_full = ta
X_train, X_test, Y_train, Y_test = train_test_split(X_full, Y_full, test_size=0.3, random_state=123)
coefs_lin_ta = poly.polyfit(X_train, Y_train, 1)
coefs_sqr_ta = poly.polyfit(X_train, Y_train, 2)
coefs_cube_ta = poly.polyfit(X_train, Y_train, 3)
X_line = np.linspace(min(X_train - 10), max(X_train + 10), 200, True)
ffit_lin = poly.polyval(X_line, coefs_lin_ta)
ffit_sqr = poly.polyval(X_line, coefs_sqr_ta)
ffit_cube = poly.polyval(X_line, coefs_cube_ta)
plt.ylim(min(Y_train - 5), max(Y_train + 5))
plt.plot(X_line, ffit_lin, "g")
plt.plot(X_line, ffit_sqr, "k")
plt.plot(X_line, ffit_cube, "b")
plt.plot(X_train, Y_train, "r.", markersize=5)
plt.show()
print('Linear Coeff = ', coefs_lin_ta)
# +
X_full = yas
Y_full = asl
X_train, X_test, Y_train, Y_test = train_test_split(X_full, Y_full, test_size=0.3, random_state=123)
coefs_lin_asl = poly.polyfit(X_train, Y_train, 1)
coefs_sqr_asl = poly.polyfit(X_train, Y_train, 2)
coefs_cube_asl = poly.polyfit(X_train, Y_train, 3)
X_line = np.linspace(min(X_train - 10), max(X_train + 10), 200, True)
ffit_lin = poly.polyval(X_line, coefs_lin_asl)
ffit_sqr = poly.polyval(X_line, coefs_sqr_asl)
ffit_cube = poly.polyval(X_line, coefs_cube_asl)
plt.ylim(min(Y_train - 5), max(Y_train + 5))
plt.plot(X_line, ffit_lin, "g")
plt.plot(X_line, ffit_sqr, "k")
plt.plot(X_line, ffit_cube, "b")
plt.plot(X_train, Y_train, "r.", markersize=5)
plt.show()
print('Linear Coeff = ', coefs_lin_asl)
# +
X_full = yas
Y_full = tw
X_train, X_test, Y_train, Y_test = train_test_split(X_full, Y_full, test_size=0.3, random_state=123)
coefs_lin_tw = poly.polyfit(X_train, Y_train, 1)
coefs_sqr_tw = poly.polyfit(X_train, Y_train, 2)
coefs_cube_tw = poly.polyfit(X_train, Y_train, 3)
X_line = np.linspace(min(X_train - 10), max(X_train + 10), 200, True)
ffit_lin = poly.polyval(X_line, coefs_lin_tw)
ffit_sqr = poly.polyval(X_line, coefs_sqr_tw)
ffit_cube = poly.polyval(X_line, coefs_cube_tw)
plt.ylim(min(Y_train - 5), max(Y_train + 5))
plt.plot(X_line, ffit_lin, "g")
plt.plot(X_line, ffit_sqr, "k")
plt.plot(X_line, ffit_cube, "b")
plt.plot(X_train, Y_train, "r.", markersize=5)
plt.show()
print('Linear Coeff = ', coefs_lin_tw)
# -
# ### 9. Make a prediction of the target variable from features dataframe
# #### Target variable is Length of Membership, most strongly correlated to Yearly Amount Spent. Fostering loyalty in customers is key to the success of this business
# ### 10. Calculate the Mean Average Error (using sklearn.metrics module)
mse_lin_lom = mean_squared_error(Y_test, poly.polyval(X_test, coefs_lin_lom))
print("MSE for linear Length of Membership regressor: ", mse_lin_lom)
mse_lin_ta = mean_squared_error(Y_test, poly.polyval(X_test, coefs_lin_ta))
print("MSE for a linear Time On App regressor: ", mse_lin_ta)
mse_lin_asl = mean_squared_error(Y_test, poly.polyval(X_test, coefs_lin_asl))
print("MSE for a linear Avg. Session Length regressor: ", mse_lin_asl)
mse_lin_tw = mean_squared_error(Y_test, poly.polyval(X_test, coefs_lin_tw))
print("MSE for a linear Time On Website regressor: ", mse_lin_tw)
mse_sqr_lom = mean_squared_error(Y_test, poly.polyval(X_test, coefs_sqr_lom))
print("MSE for quadratic Length of Membership regressor: ", mse_sqr_lom)
mse_sqr_ta = mean_squared_error(Y_test, poly.polyval(X_test, coefs_sqr_ta))
print("MSE for a quadratic Time On App regressor: ", mse_sqr_ta)
mse_sqr_asl = mean_squared_error(Y_test, poly.polyval(X_test, coefs_sqr_asl))
print("MSE for a quadratic Avg. Session Length regressor: ", mse_sqr_asl)
mse_sqr_tw = mean_squared_error(Y_test, poly.polyval(X_test, coefs_sqr_tw))
print("MSE for a quadratic Time On Website regressor: ", mse_sqr_tw)
# +
# print(coefs_cube_tw)
mse_cube_lom = mean_squared_error(Y_test, poly.polyval(X_test, coefs_cube_lom))
print("MSE for cubed Length of Membership regressor: ", mse_cube_lom)
mse_cube_ta = mean_squared_error(Y_test, poly.polyval(X_test, coefs_cube_ta))
print("MSE for a cubed Time On App regressor: ", mse_cube_ta)
mse_cube_asl = mean_squared_error(Y_test, poly.polyval(X_test, coefs_cube_asl))
print("MSE for a cubed Avg. Session Length regressor: ", mse_cube_asl)
mse_cube_tw = mean_squared_error(Y_test, poly.polyval(X_test, coefs_cube_tw))
print("MSE for a cubed Time On Website regressor: ", mse_cube_tw)
# -
# ### 11. Using Seaborn distplot show the histogram of the residuals - differences between the target variable and predicted target variable
sns.distplot(yas)
# ### 12. Answer the main question: How should we allocate the engineering budget between website development and app development?
# #### Hint: look at the regression coefficients and contemplate their meaning
# #### Given the negative correlation of website use to sales, initial thoughts are engineering can probably allocate the majority of its resources to app development. However, it could be the app has already been optimized and the website was neglected. Average users are using the website for almost 3x use as the app, meaning important decisions are being formed but the user experience might not promote purchase behavior. I recommend a qualitative deep dive in to the underlying consumer sentiment causing the observed website vs. app behaviors. If forced into a decision, focus resources on improving website UX in an effort to improve conversion rates.
|
regression/Regression - E-commerce.csv - 28 Mar 18 - Jeremy Crawford.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fishscale
# language: python
# name: fishscale
# ---
# +
# pip install pandas
# pip install PyQt5
# pip install geopandas
# pip install matplotlib
# pip install networkx
# pip install scikit-image
# pip install scikit-learn
# pip install pint
# conda install numba
# pip install ripleyk
# python -m pip install "dask[distributed]" --upgrade
# pip install scanpy
# bone_fight from linnarsson github
# pip install torch
# pip install loompy
# pip install pyarrow
# pip install open3d
# -
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('/Users/simone.codeluppi/Github_code/')
from FISHscale.utils.dataset import Dataset, MultiDataset
DS = Dataset('/Users/simone.codeluppi/Downloads/220119_06_47_18_AMEXP20211026_EEL_MTG_data_summary_simple_plotting_cleaned_microscope_stitched.parquet',
x_label='r_px_microscope_stitched', y_label='c_px_microscope_stitched',gene_label='decoded_genes')
DS.visualize()
import pandas as pd
data = pd.read_parquet('/Users/simone.codeluppi/Downloads/220119_06_47_18_AMEXP20211026_EEL_MTG_data_summary_simple_plotting_cleaned_microscope_stitched.parquet')
data.loc[data.Gene.str.contains('Control'),:]
md = MultiDataset('/Users/simone.codeluppi/Downloads/dot_removed/',
x_label='c_px_global_stitched', y_label='r_px_global_stitched',
gene_label = 'decoded_genes', reparse=False, pixel_size='0.18 micrometer')
md.visualize()
import pandas as pd
data = pd.read_parquet('/Users/simone.codeluppi/Downloads/220119_06_47_18_AMEXP20211026_EEL_MTG_data_summary_simple_plotting_cleaned_microscope_stitched.parquet')
data.columns
data.shape
data.fov_num.unique()
|
notebooks/to_check/viz_fishscale.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Intorduction
# ## What is in this notebook?
# ## Inputs
# The following are the inputs which the model needs to run, please select one of the below for each input:
# +
# inputs go here
# -
# ## Magics & Versions
# The below table shows the version of libraries and packages used for running the model.
# +
# Inline matplotlib
# %matplotlib inline
# Interactive matplotlib plot()
# #%matplotlib notebook
# Autoreload packages before runs
# https://ipython.org/ipython-doc/dev/config/extensions/autoreload.html
# %load_ext autoreload
# %autoreload 2
# # %install_ext http://raw.github.com/jrjohansson/version_information/master/version_information.py
# ~/anaconda/bin/pip install version_information
# %load_ext version_information
# %version_information numpy, scipy, matplotlib, pandas
# -
# ## Standard imports
# +
# Standard library
import os
import sys
sys.path.append("../src/")
# Third party imports
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
# Date and time
import datetime
import time
# Ipython imports
from IPython.display import FileLink
# -
# ## Other imports
# +
# Other imports
# Stats models
import statsmodels.api as sm
from statsmodels.nonparametric.kde import KDEUnivariate
from statsmodels.nonparametric import smoothers_lowess
from patsy import dmatrices
from sklearn import datasets, svm
# Sk-learn
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble.gradient_boosting import GradientBoostingClassifier
from sklearn.feature_selection import SelectKBest
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import cross_val_score
from sklearn.feature_selection import SelectFromModel
from sklearn.linear_model import LogisticRegression, LogisticRegressionCV
# -
# ## Customization
# +
# Customizations
sns.set() # matplotlib defaults
# Any tweaks that normally go in .matplotlibrc, etc., should explicitly go here
plt.rcParams['figure.figsize'] = (12, 12)
# Silent mode
import warnings
warnings.filterwarnings('ignore')
warnings.filterwarnings('ignore', category=DeprecationWarning)
# -
# Find the notebook the saved figures came from
fig_prefix = "../figures/2019-04-25-mh-titanic"
# # Data preprocessing
# ## Reading the data
data = pd.read_csv('../data/training/train.csv')
data.shape
# We have:
# * 891 rows
# * 12 columns
# ## Exploring the data
data.head()
data.dtypes
# The Survived column is the target variable. If Suvival = 1 the passenger survived, otherwise he's dead. The is the variable we're going to predict.
#
# The other variables describe the passengers. They are the features.
# * PassengerId: and id given to each traveler on the boat
# * Pclass: the passenger class. It has three possible values: 1,2,3 (first, second and third class)
# * The Name of the passeger
# * The Sex
# * The Age
# * SibSp: number of siblings and spouses traveling with the passenger
# * Parch: number of parents and children traveling with the passenger
# * The ticket number
# * The ticket Fare
# * The cabin number
# * The embarkation. This describe three possible areas of the Titanic from which the people embark. Three possible values S,C,Q
# ### Features unique values
for col in data.columns.values:
print(col, ' :', data[col].nunique())
# Pclass, Sex, Embarked are categorical features.
data.describe()
data.info()
# Age seems to have 177 missing values. let's impute this using the median age.
# ## Missing values
data['Age'] = data['Age'].fillna(data['Age'].median())
data.describe()
# # Visualization
data['Died'] = 1 - data['Survived']
# ## Sex
# Survival count based on gender
data.groupby('Sex').agg('sum')[['Survived', 'Died']].plot(kind='Bar', figsize=(12, 7),
stacked=True, colors=['g', 'r'])
plt.savefig(fig_prefix + '-Sex', dpi=300)
# Survival ratio based on the gender
data.groupby('Sex').agg('mean')[['Survived', 'Died']].plot(kind='Bar', figsize=(12, 7),
stacked=True, colors=['g', 'r'])
plt.savefig(fig_prefix + '-Sex-ratio', dpi=300)
# ## Age
# Violin plots for correlating the survival with sex and age
fig = plt.figure(figsize=(12, 7))
sns.violinplot(x='Sex', y='Age',
hue='Survived', data=data,
split=True,
palette={0:'r', 1:'g'}
)
plt.savefig(fig_prefix + '-age', dpi=300)
# As we saw in the chart above and validate by the following:
# * Women survive more than men, as depicted by the larger female green histogram
#
# Now, we see that:
# * The age conditions the survival for male passengers:
# * Younger male tend to survive
# * A large number of passengers between 20 and 40 succumb
# * The age doesn't seem to have a direct impact on the female survival
# ## Ticket fare
figure = plt.figure(figsize=(25, 12))
plt.hist([data[data['Survived'] == 1]['Fare'], data[data['Survived'] == 0]['Fare']],
stacked=True, color=['g', 'r'],
bins=50, label = ['Survived', 'Dead'])
plt.xlabel('Fare')
plt.ylabel('Nubmer of passerngers')
plt.legend()
plt.savefig(fig_prefix + '-fare-dist', dpi=300)
# +
plt.figure(figsize=(25, 7))
ax = plt.subplot()
ax.scatter(data[data['Survived'] == 1]['Age'], data[data['Survived'] == 1]['Fare'],
c='g', s=data[data['Survived'] == 1]['Fare'])
ax.scatter(data[data['Survived'] == 0]['Age'], data[data['Survived'] == 0]['Fare'],
c='r', s=data[data['Survived'] == 0]['Fare'])
plt.savefig(fig_prefix + '-fare-scatter', dpi=300)
# -
# The size of the circles is proportional to the ticket fare.
#
# On the x-axis, we have the ages and the y-axis, we consider the ticket fare.
#
# We can observe different clusters:
# 1. Large green dots between x=20 and x=45: adults with the largest ticket fares
# 2. Small red dots between x=10 and x=45, adults from lower classes on the boat
# 3. Small greed dots between x=0 and x=7: these are the children that were saved
ax = plt.subplot()
ax.set_ylabel('Average fare')
(data.groupby('Pclass')['Fare'].mean()).plot(kind='bar', figsize=(25, 7), ax=ax)
plt.savefig(fig_prefix + '-fare-class', dpi=300)
# ## Embarked
fig = plt.figure(figsize=(25, 7))
sns.violinplot(x='Embarked', y='Fare', hue='Survived', data=data,
split=True, palette={0: 'r', 1: 'g'})
plt.savefig(fig_prefix + '-embared-fare', dpi=300)
# It seems that the embarkation C have a wider range of fare tickets and therefore the passengers who pay the highest prices are those who survive.
#
# We also see this happening in embarkation S and less in embarkation Q.
# # Feature engineering
# Function that asserts whether or not a feature has been processed.
def status(feature):
print('Processing', feature, ': ok')
# Function for combining the train and the test data
def get_combined_data():
# reading the train data
train = pd.read_csv('../data/training/train.csv')
# reading the test data
test = pd.read_csv('../data/training/test.csv')
# extracting and removing the target from the training data
targets = train.Survived
train.drop(['Survived'], 1, inplace=True)
# merging train data and test data for future feature engineering
# we'll also remove the PassengerID since this is not an informative feature
combined = train.append(test)
combined.reset_index(inplace=True)
combined.drop(['index', 'PassengerId'], inplace=True, axis=1)
return combined
combined = get_combined_data()
print(combined.shape)
# ## Passenger titles
titles = set()
for name in data['Name']:
titles.add(name.split(',')[1].split('.')[0].strip())
print(titles)
title_dic ={
"Capt": "Officer",
"Col": "Officer",
"Major": "Officer",
"Jonkheer": "Royalty",
"Don": "Royalty",
"Sir" : "Royalty",
"Dr": "Officer",
"Rev": "Officer",
"the Countess":"Royalty",
"Mme": "Mrs",
"Mlle": "Miss",
"Ms": "Mrs",
"Mr" : "Mr",
"Mrs" : "Mrs",
"Miss" : "Miss",
"Master" : "Master",
"Lady" : "Royalty"
}
def get_titles():
# We extract title from each name
combined['Title'] = combined['Name'].map(lambda name:name.split(',')[1].split('.')[0].strip())
# a map of more aggregated title
# we map each title
combined['Title'] = combined.Title.map(title_dic)
status('Title')
return combined
combined = get_titles()
combined.head()
# Checking
combined[combined.Title.isnull()]
# There is indeed a NaN value in the line 1305. In fact the corresponding name is <NAME>, <NAME>.
#
# This title was not encoutered in the train dataset.
# ## Passenger ages
# Number of missing ages in train set
print(combined.iloc[:891].Age.isnull().sum())
# Number of missing ages in test set
print(combined.iloc[891:].Age.isnull().sum())
# Grouping
grouped_train = combined.iloc[:891].groupby(['Sex', 'Pclass', 'Title'])
grouped_median_train = grouped_train.median()
grouped_median_train = grouped_median_train.reset_index()[['Sex', 'Pclass', 'Title', 'Age']]
grouped_median_train.head()
# +
def fill_age(row):
condition = (
(grouped_median_train['Sex'] == row['Sex']) &
(grouped_median_train['Title'] == row['Title']) &
(grouped_median_train['Pclass'] == row['Pclass'])
)
return grouped_median_train[condition]['Age'].values[0]
def process_age():
global combined
# a function that fills the missing values of the Age variable
combined['Age'] = combined.apply(lambda row: fill_age(row) if np.isnan(row['Age']) else row['Age'], axis=1)
status('age')
return combined
# -
combined = process_age()
combined.head()
# ## Names
def process_names():
global combined
# we clean the Name variable
combined.drop('Name', axis=1, inplace=True)
# encoding in dummy variable
titles_dummies = pd.get_dummies(combined['Title'], prefix='Title')
combined = pd.concat([combined, titles_dummies], axis=1)
# removing the title variable
combined.drop('Title', axis=1, inplace=True)
status('names')
return combined
combined = process_names()
combined.head()
# ## Fare
def process_fares():
global combined
# there's one missing fare value - replacing it with the mean.
combined.Fare.fillna(combined.iloc[:891].Fare.mean(), inplace=True)
status('fare')
return combined
combined = process_fares()
# ## Embarked
def process_embarked():
global combined
# two missing embarked values - filling them with the most frequent one in the train set(S)
combined.Embarked.fillna('S', inplace=True)
# dummy encoding
embarked_dummies = pd.get_dummies(combined['Embarked'], prefix='Embarked')
combined = pd.concat([combined, embarked_dummies], axis=1)
combined.drop('Embarked', axis=1, inplace=True)
status('embarked')
return combined
combined = process_embarked()
combined.head()
# ## Cabin
# +
train_cabin, test_cabin = set(), set()
for c in combined.iloc[:891]['Cabin']:
try:
train_cabin.add(c[0])
except:
train_cabin.add('U')
for c in combined.iloc[891:]['Cabin']:
try:
test_cabin.add(c[0])
except:
test_cabin.add('U')
# -
print(train_cabin)
print(test_cabin)
# We don't have any cabin letter in the test set that is not present in the train set.
def process_cabin():
global combined
# replacing missing cabins with U (for Uknown)
combined.Cabin.fillna('U', inplace=True)
# mapping each Cabin value with the cabin letter
combined['Cabin'] = combined['Cabin'].map(lambda c: c[0])
# dummy encoding ...
cabin_dummies = pd.get_dummies(combined['Cabin'], prefix='Cabin')
combined = pd.concat([combined, cabin_dummies], axis=1)
combined.drop('Cabin', axis=1, inplace=True)
status('cabin')
return combined
# This function replaces NaN values with U (for Unknow). It then maps each Cabin value to the first letter. Then it encodes the cabin values using dummy encoding again.
combined = process_cabin()
combined.head()
# ## Sex
def process_sex():
global combined
# mapping string values to numerical one
combined['Sex'] = combined['Sex'].map({'male':1, 'female':0})
status('Sex')
return combined
combined = process_sex()
# ## Pclass
def process_pclass():
global combined
# encoding into 3 categories:
pclass_dummies = pd.get_dummies(combined['Pclass'], prefix="Pclass")
# adding dummy variable
combined = pd.concat([combined, pclass_dummies],axis=1)
# removing "Pclass"
combined.drop('Pclass',axis=1,inplace=True)
status('Pclass')
return combined
combined = process_pclass()
# ## Ticket
def cleanTicket(ticket):
ticket = ticket.replace('.', '')
ticket = ticket.replace('/', '')
ticket = ticket.split()
ticket = map(lambda t : t.strip(), ticket)
ticket = list(filter(lambda t : not t.isdigit(), ticket))
if len(ticket) > 0:
return ticket[0]
else:
return 'XXX'
tickets = set()
for t in combined['Ticket']:
tickets.add(cleanTicket(t))
print(len(tickets))
def process_ticket():
global combined
# a function that extracts each prefix of the ticket, returns 'XXX' if no prefix (i.e the ticket is a digit)
def cleanTicket(ticket):
ticket = ticket.replace('.','')
ticket = ticket.replace('/','')
ticket = ticket.split()
ticket = map(lambda t : t.strip(), ticket)
ticket = list(filter(lambda t : not t.isdigit(), ticket))
if len(ticket) > 0:
return ticket[0]
else:
return 'XXX'
# Extracting dummy variables from tickets:
combined['Ticket'] = combined['Ticket'].map(cleanTicket)
tickets_dummies = pd.get_dummies(combined['Ticket'], prefix='Ticket')
combined = pd.concat([combined, tickets_dummies], axis=1)
combined.drop('Ticket', inplace=True, axis=1)
status('Ticket')
return combined
combined = process_ticket()
# ## Family
# This part includes creating new variables based on the size of the family (the size is by the way, another variable we create).
#
# This creation of new variables is done under a realistic assumption: Large families are grouped together, hence they are more likely to get rescued than people traveling alone.
def process_family():
global combined
# introducing a new feature : the size of families (including the passenger)
combined['FamilySize'] = combined['Parch'] + combined['SibSp'] + 1
# introducing other features based on the family size
combined['Singleton'] = combined['FamilySize'].map(lambda s: 1 if s == 1 else 0)
combined['SmallFamily'] = combined['FamilySize'].map(lambda s: 1 if 2 <= s <= 4 else 0)
combined['LargeFamily'] = combined['FamilySize'].map(lambda s: 1 if 5 <= s else 0)
status('family')
return combined
combined = process_family()
print(combined.shape)
combined.head()
|
develop/2019-04-25-mh-titanic.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
using DifferentialEquations
using Plots
# Problem definition
function competition(u0, p, t)
x, y = u0 # Initial populations sizes
dx = p.R*x - p.A*x*x - p.B*x*y # variation of the x population
dy = p.r*y - p.a*y*y - p.b*x*y # variation of the y population
return [dx, dy]
end
# +
# Expression of the zero growth isoclines
# Eq 1
x_hat_1 = 0
y_hat_1 = 0
# Eq 2
x_hat_2 = R/A
y_hat_2 = 0
# Eq 3
x_hat_3 = 0
y_hat_3 = r/a
#Eq 4
x_hat_4 = (r*B-R*a)/(B*b-A*a)
y_hat_4 = (R*b-r*A)/(B*b-A*a)
# +
# Finding when equilibrium with 2 species is possible
# Initial population sizes
u0 = [0.4, 0.6]
# t0 - t end
t = (0., 2000.)
function eq_2_sp(p)
#println(p)
w = p.B*p.b-p.A*p.a
x̂ = (p.r*p.B - p.R*p.a)/w
ŷ = (p.R*p.b - p.r*p.A)/w
return (x̂, ŷ)
end
function eq_1_sp_x(p)
x̂ = (p.R/p.A)
ŷ = 0
return (x̂, ŷ)
end
function eq_1_sp_y(p)
x̂ = 0
ŷ = (p.r/p.a)
return (x̂, ŷ)
end
"""
Salut
"""
eq_2_sp_exists(p) = (p.b/p.A < p.r/p.R < p.a/p.B)
eq_2_sp_exists2(p) = (p.a/p.B < p.r/p.R < p.b/p.A)
#eq_2_sp_exists(p) = all(eq_2_sp(p).>1e-3)
eq_1_sp_exists_x(p) = ((p.r < p.R) & (p.a == p.A == p.b == p.B) | (p.A < p.a) & (p.r == p.R == p.b == p.B) | (p.B < p.b) & (p.a == p.A == p.r == p.R)) & ~ (p.b/p.A < p.r/p.R < p.a/p.B) & ~ (p.a/p.B < p.r/p.R < p.b/p.A)
eq_1_sp_exists_y(p) = ((p.R < p.r) & (p.a == p.A == p.b == p.B) | (p.a < p.A) & (p.r == p.R == p.b == p.B) | (p.b < p.B) & (p.a == p.A == p.r == p.R)) & ~ (p.b/p.A < p.r/p.R < p.a/p.B) & ~ (p.a/p.B < p.r/p.R < p.b/p.A)
#eq_1_sp_exists_x(p) = (solve(ODEProblem(competition, u0, t, p_combin))).u[end][2] < 1e-3
#eq_1_sp_exists_y(p) = (solve(ODEProblem(competition, u0, t, p_combin))).u[end][1] < 1e-3
# Grid for the search
possible_values = 0.9:0.05:1.1
# Empty array for parameter values
p_combin = []
# Grid search
for r in possible_values, R in possible_values
for a in possible_values, A in possible_values
for b in possible_values, B in possible_values
p = (r=r, R=R, a=a, A=A, b=b, B=B)
push!(p_combin, p)
end
end
end
# Keep only parameters for which the 2sp equilibrium exists
p_eq = filter(eq_2_sp_exists, p_combin)
p_eq2 = filter(eq_2_sp_exists2, p_combin)
p_x = filter(eq_1_sp_exists_x, p_combin)
p_y = filter(eq_1_sp_exists_y, p_combin)
# +
# Definition of problem
rand_p_x = rand(p_x)
rand_p_y = rand(p_y)
rand_p_xy = rand(p_eq)
rand_p_xy2 = rand(p_eq2)
prob_x = ODEProblem(competition, u0, t, rand_p_x)
prob_y = ODEProblem(competition, u0, t, rand_p_y)
prob_xy = ODEProblem(competition, u0, t, rand_p_xy)
prob_xy2 = ODEProblem(competition, u0, t, rand_p_xy2)
# Resolution of problem
solution_x = solve(prob_x)
solution_y = solve(prob_y)
solution_xy = solve(prob_xy)
solution_xy2 = solve(prob_xy2)
p4 = plot(solution_xy2, title = "Equilibrium between both x and y populations \nTesting the second condition \n$(rand_p_xy)", xlab="Time", ylab="Population size")
plot(p4, lab=["x" "y"], ylim=(0,1.2), minorgrid=true)
#plot(p1, p2, p3, ylim=(0,1.2), legend = false)
#print(solution_xy.u[end][1])
# -
print("Expected value of (x̂,ŷ): $(eq_1_sp_x(rand_p_x))")
p1 = plot(solution_x, title = "Extinction of the y population \n$(rand_p_x)", xlab="Time", ylab="Population size")
plot(p1, lab=["x" "y"], ylim=(0,1.2), minorgrid=true)
print("Expected value of (x̂,ŷ):")
print(eq_1_sp_y(rand_p_y))
p2 = plot(solution_y, title = "Extinction of the x population \n$(rand_p_y)", xlab="Time", ylab="Population size")
plot(p2, lab=["x" "y"], ylim=(0,1.2), minorgrid=true)
print("Expected value of (x̂,ŷ):")
print(eq_2_sp(rand_p_xy))
p3 = plot(solution_xy, title = "Equilibrium between both x and y populations \nTesting the first condition\n$(rand_p_xy)", xlab="Time", ylab="Population size")
plot(p3, lab=["x" "y"], ylim=(0,1.2), minorgrid=true)
p4 = plot(solution_xy2, title = "Equilibrium between both x and y populations \nTesting the second condition \n$(rand_p_xy)", xlab="Time", ylab="Population size")
plot(p4, lab=["x" "y"], ylim=(0,1.2), minorgrid=true)
# +
# simulation of 4 combinaisons of parameters 1
solution1 = []
solution2 = []
solution3 = []
solution4 = []
for z in 1:100
# Parameters
x = rand(p_combin)
print(eq_2_sp(x))
if (eq_2_sp(x)[1] < 1e-1)
print("salut")
prob2 = ODEProblem(competition, u0, t, x)
# Resolution of problem
solution2 = solve(prob2)
end
if (eq_2_sp(x)[2] < 1e-1)
prob_eq_3 = ODEProblem(competition, u0, t, x)
# Resolution of problem
solution_eq_3 = solve(prob_eq_3)
end
if (eq_2_sp(x)[1] > 0.05 && eq_2_sp(x)[2] > 0.05)
# Definition of problem
prob4 = ODEProblem(competition, u0, t, x)
# Resolution of problem
solution4 = solve(prob4)
#push!(solution_graph, solution)
end
end
# Plot
#p1 = plot(solution_graph[1], title = "Parameters: a=")
#p2 = plot(solution_graph[2])
#p3 = plot(solution_graph[3])
#p4 = plot(solution_graph[4])
#plot(p1,p2,p3,p4,layout=(2,2), legend=false)
# Plot
p1 = plot(solution1, title = "Parameters: a=")
p2 = plot(solution2)
p3 = plot(solution3)
p4 = plot(solution4)
plot(p1,p2,p3,p4,layout=(2,2), legend=false)
#plot(solution)
#plot(solution, vars=(1,2), xlab="x", ylab="y", aspectratio=1, xlim=(0,1), ylim=(0,1))
#scatter!([u0[1]], [u0[2]], c=:black, lab="u0")
# -
# +
# simulation of 4 combinaisons of parameters 2
solution_graph = []
# Parameters
p_eq_2 = (r=1.0, R=1.0, a=1.01, A=0.99, b=1.0, B=1.0)
p_eq_3 = (r = 0.9, R = 0.9, a = 0.9, A = 0.95, b = 0.9, B = 0.9)
p_eq_4_1 = (r=1.0, R=1.0, a=0.9, A=0.9, b=1.1, B=1.1)
p_eq_4_2 = (r=1.0, R=1.0, a=1.1, A=1.1, b=0.9, B=0.9)
print(eq_2_sp(p_eq_4_1))
# Definition of problem
prob_2 = ODEProblem(competition, u0, t, p_eq_2)
prob_3 = ODEProblem(competition, u0, t, p_eq_3)
prob_4_1 = ODEProblem(competition, u0, t, p_eq_4_1)
prob_4_2 = ODEProblem(competition, u0, t, p_eq_4_2)
# Resolution of problem
solution_4_1 = solve(prob_4_1)
solution_4_2 = solve(prob_4_2)
solution_2 = solve(prob_2)
solution_3 = solve(prob_3)
p2 = plot(solution_2, label="a")
p3 = plot(solution_3, label="b")
p4_1 = plot(solution_4_1, label="c")
p4_2 = plot(solution_4_2, label="d")
plot(p2,p3,p4_1,p4_2)
#print(solution_2)
# +
p = rand(p_eq)
print(eq_2_sp(p))
print(eq_2_sp(p)[1])
print(eq_2_sp(p)[2])
if (eq_2_sp(p)[1] > 0.4 && eq_2_sp(p)[2] > 0.4)
print("PLUS GRAND")
else
print("PLUS PETIT")
end
# -
for i in 1:1000
p = rand(p_eq)
print(eq_2_sp(p))
end
|
homework/Devoir_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manifold
#
# **Manifold** is a tool for model-agnostic evaluation with visual support developed by Uber — you can find its repo [here](https://github.com/uber/manifold).
#
# In this quick demo, the [California housing dataset](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_california_housing.html) (regression) and part of the setup available in this [example](https://scikit-learn.org/stable/auto_examples/inspection/plot_partial_dependence.html) on the [scikit-learn](https://scikit-learn.org/stable/index.html) website are used.
#
# The main idea for this demo is to test **Manifold**'s [integration with Jupyter Notebook](https://eng.uber.com/manifold-open-source/) and the [Geo Feature View](https://github.com/uber/manifold#geo-feature-view) map.
# +
from sklearn.datasets import fetch_california_housing
import pandas as pd
import numpy as np
# Installation: https://github.com/uber/manifold/tree/master/bindings/jupyter
from mlvis import Manifold
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import QuantileTransformer
from sklearn.pipeline import make_pipeline
from sklearn.neural_network import MLPRegressor
# -
california_housing = fetch_california_housing()
california_housing.data.shape
california_housing.feature_names
california_housing.target.shape
california_housing.keys()
# +
X = pd.DataFrame(california_housing.data, columns=california_housing.feature_names)
y = california_housing.target
# +
# The Latitude feature must be called "lat".
# The Longitude feature must be called "lng".
X.rename(columns={'Latitude':'lat', 'Longitude': 'lng'}, inplace=True)
# -
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=0)
# +
est = make_pipeline(QuantileTransformer(),
MLPRegressor(hidden_layer_sizes=(50, 50),
learning_rate_init=0.01,
early_stopping=True))
est.fit(X_train, y_train)
print("Test R2 score: {:.2f}".format(est.score(X_test, y_test)))
# -
yPred = est.predict(X_test)
# +
# Mapbox access token: https://docs.mapbox.com/help/how-mapbox-works/access-tokens/
# It must be replaced with a valid token.
TOKEN = "<KEY>"
# +
# Props: https://github.com/uber/manifold/blob/master/bindings/jupyter-modules/jupyter-manifold/src/manifold.js
# Classification: https://github.com/uber/manifold/blob/master/bindings/jupyter/notebooks/manifold.ipynb
Manifold(props={'data': {
'x': X_test[['lat', 'lng']],
'yPred': [pd.DataFrame(yPred, columns=["Target"])], # Each element in this list contains the predictions for a model.
'yTrue': pd.DataFrame(y_test, columns=["Target"])
},
'mapboxAccessToken': TOKEN,
'width': 1000,
'height': 700
})
|
manifold-demo/Manifold demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bayesian HMM Model
#
# This notebook illustrate how to build and train a Bayesian Hidden Markov Model with the [beer framework](https://github.com/beer-asr/beer).
# +
# Add "beer" to the PYTHONPATH
import sys
sys.path.insert(0, '../')
import copy
import beer
import numpy as np
import torch
# For plotting.
from bokeh.io import show, output_notebook
from bokeh.plotting import figure, gridplot
from bokeh.models import LinearAxis, Range1d
output_notebook()
# Convenience functions for plotting.
import plotting
# %load_ext autoreload
# %autoreload 2
# +
def create_ali_trans_mat(tot_states):
'''Create align transition matrix for a sequence of units
Args:
tot_states (int): length of total number of states of the given
sequence.
'''
trans_mat = torch.diag(torch.ones(tot_states) * .5)
idx1 = torch.arange(0, tot_states-1, dtype=torch.long)
idx2 = torch.arange(1, tot_states, dtype=torch.long)
trans_mat[idx1, idx2] = .5
trans_mat[-1, -1] = 1.
return trans_mat
# Sequence: AB
seqs = ['A', 'B', 'A']
nsamples = 30
ndim = 2
units = ['A', 'B']
len_seqs = len(seqs)
num_unit_states = 3
tot_states = len(seqs) * num_unit_states
trans_mat = create_ali_trans_mat(tot_states)
means = [np.array([-1.5, 3]), np.array([-1.5, 4]), np.array([-1.5, 5]),
np.array([1, -3]), np.array([1, -2]), np.array([1, -1])]
covs = [np.array([[.75, -.5], [-.5, 2.]]), np.array([[.75, -.5], [-.5, 2.]]), np.array([[.75, -.5], [-.5, 2.]]),
np.array([[2, 1], [1, .75]]), np.array([[2, 1], [1, .75]]), np.array([[2, 1], [1, .75]])]
states_id = {'A':[0, 1, 2], 'B':[3, 4, 5]}
dict_seq_state = {}
seqs_id = []
for i, j in enumerate(seqs):
for u in range(num_unit_states):
dict_seq_state[num_unit_states * i + u] = states_id[j][u]
seqs_id.append(states_id[j][u])
normal_sets = list(zip(means,covs))
states = np.zeros(nsamples, dtype=np.int16)
data = np.zeros((nsamples, ndim))
states[0] = states_id['A'][0]
data[0] = np.random.multivariate_normal(means[0], covs[0], size=1)
colors = ['blue', 'blue', 'blue', 'red', 'red', 'red']
fig1 = figure(title='Samples', width=400, height=400)
fig1.circle(data[0, 0], data[0, 1], color=colors[states[0]])
for n in range(1, nsamples):
states[n] = np.random.choice(np.arange(tot_states), p=trans_mat[states[n-1]].numpy())
data[n] = np.random.multivariate_normal(means[dict_seq_state[states[n]]], covs[dict_seq_state[states[n]]], size=1)
fig1.circle(data[n, 0], data[n, 1], color=colors[dict_seq_state[states[n]]], line_width=1)
fig1.line(data[n-1:n+1, 0], data[n-1:n+1, 1], color='black', line_width=.5, alpha=.5)
states_id = [dict_seq_state[i] for i in states]
fig2 = figure(title='Emissions', width=400, height=400)
colors = ['darkblue', 'blue', 'skyblue', 'darkred','red', 'pink']
for i, n in enumerate(normal_sets):
plotting.plot_normal(fig2, n[0], n[1], alpha=.3, color=colors[i])
grid = gridplot([[fig1, fig2]])
show(grid)
print(states_id)
# -
# ## Model Creation
#
# We create several types of HMMs, each of them has the same transition matrix and initial / final state probability, and a specific type of emission density:
# * one Normal density per state with full covariance matrix
# * one Normal density per state with diagonal covariance matrix
# * one Normal density per state with full covariance matrix shared across states
# * one Normal density per state with diagonal covariance matrix shared across states.
graph = beer.graph.Graph()
s0 = graph.add_state()
s1 = graph.add_state(pdf_id=0)
s2 = graph.add_state(pdf_id=1)
s3 = graph.add_state(pdf_id=2)
s4 = graph.add_state()
graph.start_state = s0
graph.end_state = s4
graph.add_arc(s0, s1)
graph.add_arc(s1, s1)
graph.add_arc(s1, s2)
graph.add_arc(s2, s2)
graph.add_arc(s2, s3)
graph.add_arc(s3, s3)
graph.add_arc(s3, s1)
graph.add_arc(s3, s4)
graph.normalize()
graph
graph.normalize()
loop_graph = graph.compile()
graph = beer.graph.Graph()
s0 = graph.add_state()
s1 = graph.add_state(pdf_id=0)
s2 = graph.add_state(pdf_id=1)
s3 = graph.add_state(pdf_id=2)
s4 = graph.add_state(pdf_id=3)
s5 = graph.add_state(pdf_id=4)
s6 = graph.add_state(pdf_id=5)
s7 = graph.add_state(pdf_id=0)
s8 = graph.add_state(pdf_id=1)
s9 = graph.add_state(pdf_id=2)
s10 = graph.add_state()
graph.start_state = s0
graph.end_state = s10
graph.add_arc(s0, s1)
graph.add_arc(s1, s1)
graph.add_arc(s1, s2)
graph.add_arc(s2, s2)
graph.add_arc(s2, s3)
graph.add_arc(s3, s3)
graph.add_arc(s3, s4)
graph.add_arc(s4, s4)
graph.add_arc(s4, s5)
graph.add_arc(s5, s5)
graph.add_arc(s5, s6)
graph.add_arc(s6, s6)
graph.add_arc(s6, s7)
graph.add_arc(s7, s7)
graph.add_arc(s7, s8)
graph.add_arc(s8, s8)
graph.add_arc(s8, s9)
graph.add_arc(s9, s9)
graph.add_arc(s9, s10)
graph.normalize()
graph
ali_graph = graph.compile().double()
# +
# We use the global mean/cov. matrix of the data to initialize the mixture.
data_mean = torch.from_numpy(data.mean(axis=0)).float()
data_var = torch.from_numpy(np.cov(data.T)).float()
# HMM (diag cov).
modelset = beer.NormalSet.create(data_mean, data_var, size=loop_graph.n_states,
prior_strength=1., noise_std=1.,
cov_type='full')
hmm_diag_loop = beer.HMM.create(loop_graph, modelset)
modelset = beer.NormalSet.create(data_mean, data_var, size=ali_graph.n_states,
prior_strength=1., noise_std=1.,
cov_type='full')
hmm_diag_align = beer.HMM.create(ali_graph, modelset)
models = {
'hmm_diag_loop': hmm_diag_loop.double(),
'hmm_diag_align': hmm_diag_align.double()
}
# -
# ## Variational Bayes Training
# +
epochs = 100
lrate = 1.
X = torch.from_numpy(data).double()
optims = {
model_name: beer.VariationalBayesOptimizer(model.mean_field_factorization(), lrate)
for model_name, model in models.items()
}
elbos = {
model_name: []
for model_name in models
}
inf_graphs = {
'hmm_diag_loop': None,
'hmm_diag_align': ali_graph
}
for epoch in range(epochs):
for name, model in models.items():
optim = optims[name]
optim.init_step()
elbo = beer.evidence_lower_bound(model, X, datasize=len(X),
inference_graph=inf_graphs[name],
viterbi=True)
elbo.backward()
elbos[name].append(float(elbo) / len(X))
optim.step()
# +
colors = {
'hmm_diag_loop': 'green',
'hmm_diag_align': 'blue'
}
# Plot the ELBO.
fig = figure(title='ELBO', width=400, height=400, x_axis_label='step',
y_axis_label='ln p(X)')
for model_name, elbo in elbos.items():
fig.line(range(len(elbo)), elbo, legend=model_name, color=colors[model_name])
fig.legend.location = 'bottom_right'
show(fig)
# +
mean = data.mean(axis=0)
var = data.var(axis=0)
std_dev = np.sqrt(max(var))
x_range = (mean[0] - 2 * std_dev, mean[0] + 2 * std_dev)
y_range = (mean[1] - 2 * std_dev, mean[1] + 2 * std_dev)
global_range = (min(x_range[0], y_range[0]), max(x_range[1], y_range[1]))
fig1 = figure(title='HMM (diag) loop', x_range=global_range, y_range=global_range,
width=400, height=400)
fig1.circle(data[:, 0], data[:, 1], alpha=.5, color='blue')
plotting.plot_hmm(fig1, hmm_diag_loop, alpha=.1, color='blue')
fig2 = figure(title='HMM (diag) align', x_range=global_range, y_range=global_range,
width=400, height=400)
fig2.circle(data[:, 0], data[:, 1], alpha=.5, color='red')
plotting.plot_hmm(fig2, hmm_diag_align, alpha=.1, color='red')
grid = gridplot([[fig1, fig2]])
show(grid)
# -
# ### Plotting
# We are mixing bokeh and matplotlib >:-( ! .
import matplotlib.pyplot as plt
# %matplotlib inline
# +
posts1 = models['hmm_diag_loop'].posteriors(X).numpy().T
posts2 = models['hmm_diag_align'].posteriors(X, ali_graph).numpy().T
fig1, axarr = plt.subplots(2, 1)
axarr[0].imshow(posts1, origin='lower')
axarr[0].set_title('HMM loop (diag) lhs')
axarr[1].imshow(posts2, origin='lower')
axarr[1].set_title('HMM align (diag) lhs')
plt.tight_layout()
plt.show()
|
examples/HMM_align.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] colab_type="text" id="-2pzsVf_4e6E"
# # DL Indaba Practical 4
# # Gated Recurrent Models (GRUs and LSTMs)
#
# **Introduction**
#
# So far we have looked at feedforward models which learn to map a single input **x** to a label (prediction) **y**. However, a lot of real-world data comes in the form of sequences, for example the words in natural languages, phonemes in speech, and so forth. In this practical we move from feed-forward models to ***sequence models*** which are designed to specifically model the **dependencies** between inputs in sequences of data that change over time.
#
# We will again apply this model to predict the labels of the handwritten MNIST images (yes! images can be thought of as **sequences of pixels**!). Then we will show how a very small change turns our MNIST classifier into a **language model** that learns the distribution of words in language!
#
# **What is expected of you:**
# * **TODO: Stipulate desired outcomes (with time-limits).**
# + [markdown] colab_type="text" id="YZTO9IIiWQNR"
# # Recurrent Neural Networks (RNNs)
#
# **NOTE**: You can safely skip the first section below if you know RNNs. Skim the second and dip into the third, but be sure to resurface back at "Putting it all together"!
#
# ## The intuition
#
# RNNs generalize feedforward networks (FFNs) to be able to work with sequential data. FFNs take an input (e.g. an image) and immediately produce an output (e.g. a digit class). RNNs, on the other hand, consider the data sequentially and remembers what it has seen in the past in order to make new predictions about the future observations.
#
# To understand this distinction, consider the example where we want to label words as the part-of-speech categories that they belong to: E.g. for the input sentence 'I want a duck' and 'He had to duck', we want our model to predict that duck is a noun in the first sentence and a verb in the second. To do this successfully, the model needs to be aware of the surrounding context. However, if we feed a FFN model only one word at a time, how could it know the difference? If we want to feed it all the words at once, how do we deal with the fact that sentences are of different lengths? (We *could* try to find a trade-off by feeding it *windows of words*...)
#
# RNNs solve this issue by processing the sentence word-by-word, and maintaining an internal state summarizing what it has seen so far. This applies not only to words, but also to phonemes in speech, or even, as we will see, pixels of an image.
#
# ## The RNN API
#
# Feedforward neural networks operate on vectors of fixed size. As we have done before, we could think of the "API" of feedforward models as follows:
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="zBV_CSuvWPm5"
class FeedForwardModel():
# ...
def forward(self, x):
'''One example of a '''
# Compute activations on the hidden layer.
hidden_layer = np.tanh(np.dot(self.W_xh, x))
# Compute the (linear) output layer activations.
output = np.dot(self.W_ho, hidden_layer)
return output
# + [markdown] colab_type="text" id="oU95UPGzWXOR"
# Recurrent neural networks (RNNs) generalize this idea to operating on **sequences of vectors**. To process sequences, the model has an internal **state** which gets updated with each new observation. Computationally, one can think of this as recursively applying a function `recurrent_fn` to update the state of the model based on each new input in the sequence:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="FiNLIPNhWbiZ"
class RecurrentModel():
# ...
def recurrent_fn(self, x, prev_state):
'''Process the current input and the previous state and produce an output and a new state.'''
# Compute the new state based on the previous state and current input.
new_state = np.tanh(np.dot(self.W_hh, prev_state) + np.dot(self.W_xh, x))
# Compute the output vector.
y = np.dot(self.W_hy, new_state)
return new_state, y
def forward(self, data_sequence):
state = self.init_state()
all_states = [state]
last_output = None
for x in data_sequence:
new_state, last_output = recurrent_fn(x, state)
all_states.append(new_state)
state = new_state
return all_states, last_output
# + [markdown] colab_type="text" id="w-DG9N5tEdaD"
# ## Putting this together
#
# Look at the definition of `hidden_layer` in `FeedForwardModel.forward()` and `new_state` in `RecurrentModel.forward()`. If you're more comfortable with math, compare the expression for computing the hidden layer of a feedforward neural network:
#
# * $h = \sigma(W_{xh}x)$
#
# to the expression for computing the hidden layer at time step $t$ in an RNN:
#
# * $h_t = \sigma(W_{hh}h_{t-1} + W_{xh}x_t)$
#
# **QUESTIONS**:
# * How are they similar?
# * How are they different?
# * Why is $W_{hh}$ called "recurrent" weights?
#
# **NOTE**: Spend a few min to think about/discuss this before you move on.
#
#
# ## 'Unrolling' the network
#
# Imagine we are trying to classify sequences `X` into labels `y` (for now, let's keep it abstract). After running the `forward()` function of our RNN defined above on `X`, we would have a list of internal states of the model at each sequence position, and the final state of the network. This process is called **unrolling in time**, because you can think of it as unrolling the *computation graph* defined by the RNN `forward` function, over the inputs at each position of the sequence. RNNs are often used to model **time series data**, and therefore these positions are referred to as **time-steps**, hence, "unrolling over time".
#
# **TODO(sgouws)**: include graph to display this, e.g. https://r2rt.com/static/images/NH_StateLoop.png
#
# > **We can therefore think of an RNN as a composition of identical feedforward neural networks (with replicated/tied weights), one for each moment or step in time. **
#
# These feedforward functions (i.e. our `recurrent_fn` above) are typically referred to as **cells**, and the only restriction on its API is that the cell function needs to be a differentiable function that can map an input and a state vector to an output and a new state vector. What we have shown above is called the **vanilla RNN**, but there are many more possibilities. In this practical, we will build up to a family of **gated-recurrent cells**. One of the most popular variants is called the **Long short-term memory** cell. But we're getting ahead of ourselves.
# + [markdown] colab_type="text" id="kLweKkB7F4j4"
# ## Training RNNs: (Truncated) Back-prop through Time
#
# RNNs model sequential data, and are designed to capture how ***outputs*** at the current time step are influenced by the ***inputs*** that came before them. This is referred to as **long-range dependencies**. At a high level, this allows the model to `remember` what it has seen so far in order to better contextualize what it is seeing at the moment (think about how knowing the context of the sentence or conversation can sometimes help one to better figure out the intended meaning of a misheard word or ambiguous statement). It is what makes these models so powerful, but it is also what makes them so hard to train!
#
# ### BPTT: A quick theoretical overview
#
# The most well-known algorithm for training RNNs is called **back-propagation through time (BPTT)** (there are other algorithms). BPTT conceptually amounts to unrolling the computations of the RNN over time, computing the errors, and backpropagating the gradients through the unrolled graph structure. Ideally we want to unroll the graph up to the maximum sequence length, however in practice, since sequence lengths vary and memory is limited, we only end up unrolling sequences up to some length $T$. This is called **truncated BPTT**, and is the most used variant of BPTT.
#
# At a high level, there are two main issues when using (truncated) BPTT to train RNNs:
#
# * Shared / tied recurrent weights ($W_{hh}$) mean that **the gradient on these weights at some time step $t$ depends on all time steps up to time-step $T$**, the maximum length of the unrolled graph. This also leads to the **vanishing/exploding gradients** problem.
#
# * As alluded to above, **memory usage grows linearly with the total number of steps $T$ that we unroll for**, because we need to save/cache the activations at each time-step. This matters computationally, since memory is a limited resource. It also matters statistically, because it puts a limit on the types of dependencies the model is exposed to, and hence that it could learn.
#
# **NOTE**: Think about that last statement and make sure you understand those 2 points.
#
# BPTT is very similar to the standard back-propagation algorithm. Key to understanding the BPTT algorithm is to realize that gradients on the non-recurrent weights (weights of a per time-step classifier that tries to predict the next word in a sentence for example) and recurrent weights (that transform $h_{t-1}$ into $h_t$) are computed differently:
#
# * The gradients of **non-recurrent weights** ($W_{hy}$) depend only on the error at that time-step, $E_t$.
# * The gradients of **recurrent weights** ($W_{hh}$) depend on all time-steps up to maximum length $T$.
#
# The first point is fairly intuitive: predictions at time-step $t$ is related to the loss of that particular prediction.
#
# The second point will be explained in more detail in the lectures (see also [this great blog post](http://www.wildml.com/2015/10/recurrent-neural-networks-tutorial-part-3-backpropagation-through-time-and-vanishing-gradients/)), but briefly, this can be summarized in these equations:
#
# 1. The **current** state is a function of the **previous** state: $h_t = \sigma(W_{hh}h_{t-1} + W_{xh}x_t)$
# 2. The gradient of the loss $E_t$ at time $t$ on $W_{hh}$ is a function of the current hidden state and model predictions $\hat{y}_t$ at time t:
# $\frac{\partial E_t}{\partial W_{hh}} = \frac{\partial E_t}{\partial \hat{y}_t}\frac{\partial\hat{y}_t}{\partial h_t}\frac{\partial h_t}{\partial W_{hh}}$
# 3. Substituting (1) into (2) results in a **sum over all previous time-steps**:
# $\frac{\partial E_t}{\partial W_{hh}} = \sum\limits_{k=0}^{t} \frac{\partial E_t}{\partial \hat{y}_t}\frac{\partial\hat{y}_t}{\partial h_t}\frac{\partial h_t}{\partial h_k}\frac{\partial h_k}{\partial W_{hh}}$
#
# Because of this **repeated multiplicative interaction**, as the sequence length $t$ gets longer, the gradients themselves can get diminishingly small (**vanish**) or grow too large and result in numeric overflow (**explode**). This has been shown to be related to the norms of the recurrent weight matrices being less than or equal to 1. Intuitively, it works very similar to how multiplying a small number $v<1.0$ with itself repeatedly can quickly go to zero, or conversely, a large number $v>1.0$ could quickly go to infinity; only this is for matrices.
#
# + [markdown] colab_type="text" id="9KRWSWRbGzBJ"
# To implement this, we need three components:
#
# * The code to fprop one time-step though the cell,
# * the code to fprop through the unrolled RNN,
# * the code to backprop one time-step through the cell,
#
# And then we'll put all this together within the BPTT algorithm. Let's start.
#
# ### Forward Propagation
#
# Let's write the code to fprop one time-step though the cell. We'll need some helper functions:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="NINCPVs72WqW"
import numpy as np
## HELPER DEFINITIONS
## NOTE: WE KEEP THESE EXPLICIT BECAUSE WE WILL NEED THEIR DERIVATIVES BELOW.
def softmax(X):
eX = np.exp((X.T - np.max(X, axis=1)).T)
return (eX.T / eX.sum(axis=1)).T
def cross_entropy(y_pred, y_train):
m = y_pred.shape[0]
prob = softmax(y_pred)
log_like = -np.log(prob[range(m), y_train])
data_loss = np.sum(log_like) / m
return data_loss
def fc_forward(X, W, b):
'''A fully-connected feedforward layer.'''
out = np.dot(X, W) + b
cache = (W, X)
return out, cache
def tanh_forward(X):
out = np.tanh(X)
cache = out
return out, cache
# + [markdown] colab_type="text" id="kikiBXVjIwG8"
# Now we can implement the equation $h_t = \sigma(W_{hh}h_{t-1} + W_{xh}x_t)$ (where $\sigma$ is our non-linearity, tanh in this case) as follows:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Bw94oOd1I1Jj"
## Define the RNN Cell. We use a vanilla RNN.
def cell_fn_forward(X, h, model, train=True):
Wxh, Whh, Why = model['Wxh'], model['Whh'], model['Why']
bh, by = model['bh'], model['by']
hprev = h.copy()
h, h_cache = tanh_forward(np.dot(hprev, Whh) + np.dot(X, Wxh) + bh)
y, y_cache = fc_forward(h, Why, by)
cache = (X, Whh, h, hprev, y, h_cache, y_cache)
if not train:
# Compute per-time step outputs.
# NOTE: Here we build a classifer, but it could be anything else.
y = softmax(y)
return y, h, cache
# + [markdown] colab_type="text" id="TRFfMGOVTt-C"
# Put this together to do the RNN fprop over the entire sequence:
#
# **QUESTION**: Notice how we save all activations in `caches`. Why do we need to do this?
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="XQUTNuJYTxqM"
def rnn_forward(X_train, y_train, model, initial_state, verbose=True):
ys = []
caches = []
loss = 0.
h = initial_state
t = 0
for x, y in zip(X_train, y_train):
y_pred, h, cache = cell_fn_forward(x, h, model, train=True)
loss += cross_entropy(y_pred, y)
ys.append(y_pred)
caches.append(cache)
if verbose:
print "Time-step: ", t
print "x_t = ", x
print "cur_state = ", h
print "predicted y = ", y_pred
t += 1
# We return final hidden state, predictions, caches and final total loss.
return h, ys, caches, loss
# + [markdown] colab_type="text" id="WLVIHIz1Kvug"
# Let's test this on some dummy data.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 731, "output_extras": [{}]} colab_type="code" executionInfo={"elapsed": 536, "status": "ok", "timestamp": 1503667515621, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-6znVyM1oxdg/AAAAAAAAAAI/AAAAAAAAABI/vEPo2Ce7Rpc/s50-c-k-no/photo.jpg", "userId": "102606466886131565871"}, "user_tz": -60} id="NQJ7DH-gJxbq" outputId="ef70162d-0e2d-4316-bf85-e8efcdd55b5b"
# Create a helper function that calculates the relative error between two arrays
def relative_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
# We set the seed to make the results reproducible.
np.random.seed(1234)
def _initial_state(hidden_dim):
return np.zeros((1, hidden_dim))
def _init_model(input_dim, hidden_dim, output_dim):
D, H, C = input_dim, hidden_dim, output_dim # More compact.
model_params = dict(
Wxh=np.random.randn(D, H) / np.sqrt(D / 2.),
Whh=np.random.randn(H, H) / np.sqrt(H / 2.),
Why=np.random.randn(H, D) / np.sqrt(C / 2.),
bh=np.zeros((1, H)),
by=np.zeros((1, D)))
return model_params
# Initialize model.
input_dim=8
hidden_dim=4
output_dim=2 # num_classes
num_steps = 5
test_mdl = _init_model(input_dim, hidden_dim, output_dim)
# Create some dummy data (there's no batching, just num_steps input vectors)
X_test = np.split(np.random.randn(num_steps, input_dim), num_steps, axis=0)
y_test = np.random.randint(low=0, high=output_dim, size=num_steps).reshape(-1)
#y_onehot = np.eye(output_dim)[y_ids]
print "Created dummy input data X"
print "Created fake targets: \n", y_test
#print "Created fake onehot targets: \n", y_onehot
print "\nRunning FPROP on dummy data: "
initial_state = _initial_state(hidden_dim)
last_state, ys, caches, loss = rnn_forward(X_test, y_test, test_mdl, initial_state, verbose=True)
print "Final hidden state: ", last_state
correct_final_state = np.array([[ 0.19868001, 0.98286478, 0.76491549, -0.91578737]])
# Compare your output to the "correct" ones
# The difference should be around 2e-8 (or lower)
print "\n============================"
print 'Testing rnn_forward'
diff = relative_error(last_state, correct_final_state)
if diff <= 2e-8:
print 'PASSED'
else:
print 'The difference of %s is too high, try again' % diff
print "\n============================"
# + [markdown] colab_type="text" id="H3q0dGyzG6Ji"
# ### Computing the derivative: Truncated BPTT
#
# Let's start with computing the per time-step derivative of `recurrent_fn` wrt all model parameters. First, some helper derivative functions:
#
# **NOTE**: Make sure you understand how these were derived.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="rTCynUOYEg5i"
## HELPER DERIVATIVE FUNCTIONS
def fc_backward(dout, cache):
W, h = cache
dW = np.dot(h.T, dout)
db = np.sum(dout, axis=0)
dX = np.dot(dout, W.T)
return dX, dW, db
def tanh_backward(dout, cache):
dX = (1 - cache**2) * dout
return dX
def dcross_entropy(y_pred, y_train):
m = y_pred.shape[0]
grad_y = softmax(y_pred)
grad_y[range(m), y_train] -= 1.
grad_y /= m
return grad_y
# + [markdown] colab_type="text" id="evUka0sdM6ul"
# Let's put these together and write the code for $\frac{\partial E}{\partial \theta}$ for all parameters:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="wo2xTk99M76j"
## PERFORM PER-TIMESTEP BACKWARD STEP
def cell_fn_backward(y_pred, y_train, dh_next, cache):
X, Whh, h, hprev, y, h_cache, y_cache = cache
# Softmax gradient
dy = dcross_entropy(y_pred, y_train)
# Hidden to output gradient
dh, dWhy, dby = fc_backward(dy, y_cache)
dh += dh_next
dby = dby.reshape((1, -1))
# tanh
dh = tanh_backward(dh, h_cache)
# Hidden gradient
dbh = dh
dWhh = np.dot(hprev.T, dh)
dWxh = np.dot(X.T, dh)
dh_next = np.dot(dh, Whh.T)
grad = dict(Wxh=dWxh, Whh=dWhh, Why=dWhy, bh=dbh, by=dby)
return grad, dh_next
# + [markdown] colab_type="text" id="-6hdtyrPOCvH"
# Now let's put this together inside the BPTT algorithm:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="CHVwqUHvEmfa"
def bptt(model, X_train, y_train, initial_state):
# Forward
last_state, ys, caches, loss = rnn_forward(X_train, y_train, model, initial_state)
loss /= y_train.shape[0]
# Backward
dh_next = np.zeros((1, last_state.shape[0]))
grads = {k: np.zeros_like(v) for k, v in model.items()}
for t in reversed(range(len(X_train))):
grad, dh_next = cell_fn_backward(ys[t], y_train[t], dh_next, caches[t])
for k in grads.keys():
grads[k] += grad[k]
for k, v in grads.items():
grads[k] = np.clip(v, -5., 5.)
return grads, loss, last_state
# + [markdown] colab_type="text" id="55st2mH8ghcV"
# Finally, we can check our implementation using numerically-derived gradients:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="eNSXgrwvgtYY"
def rnn_gradient_check(model, x, y, init_state, h=0.001, error_threshold=0.01):
# Calculate the gradients using backpropagation. We want to checker if these are correct.
bptt_gradients, _, _ = bptt(model, x, y, init_state)
# List of all parameters we want to check.
model_parameters = ['Wxh', 'Whh', 'Why', 'bh', 'by']
# Gradient check for each parameter
for pidx, pname in enumerate(model_parameters):
# Get the actual parameter value from the model, e.g. model.W
parameter = model[pname]
print "Performing gradient check for parameter %s with size %d." % (pname, np.prod(parameter.shape))
# Iterate over each element of the parameter matrix, e.g. (0,0), (0,1), ...
it = np.nditer(parameter, flags=['multi_index'], op_flags=['readwrite'])
while not it.finished:
ix = it.multi_index
# Save the original value so we can reset it later
original_value = parameter[ix]
# Estimate the gradient using (f(x+h) - f(x-h))/(2*h)
parameter[ix] = original_value + h
# Compute the ENTIRE rnn_foward, evaluate cross entropy loss
_, _, _, gradplus = rnn_forward(x, y, model, init_state, verbose=False)
parameter[ix] = original_value - h
_, _, _, gradminus = rnn_forward(x, y, model, init_state, verbose=False)
estimated_gradient = (gradplus - gradminus)/(2*h)
# Reset parameter to original value
parameter[ix] = original_value
# The gradient for this parameter calculated using backpropagation
backprop_gradient = bptt_gradients[pname][ix]
# calculate The relative error: (|x - y|/(|x| + |y|))
relative_error = np.abs(backprop_gradient - estimated_gradient) / (np.abs(backprop_gradient) + np.abs(estimated_gradient))
# If the error is to large fail the gradient check
if relative_error > error_threshold:
print "Gradient Check ERROR: parameter=%s ix=%s" % (pname, ix)
print "+h Loss: %f" % gradplus
print "-h Loss: %f" % gradminus
print "Estimated_gradient: %f" % estimated_gradient
print "Backpropagation gradient: %f" % backprop_gradient
print "Relative Error: %f" % relative_error
return
it.iternext()
print "Gradient check for parameter %s: PASSED." % (pname)
# + [markdown] colab_type="text" id="CtGlXaAZkEpV"
# Aaaaand let's test it!
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 697, "output_extras": [{}]} colab_type="code" executionInfo={"elapsed": 410, "status": "ok", "timestamp": 1503668261812, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-6znVyM1oxdg/AAAAAAAAAAI/AAAAAAAAABI/vEPo2Ce7Rpc/s50-c-k-no/photo.jpg", "userId": "102606466886131565871"}, "user_tz": -60} id="yPt9o7pwkEVE" outputId="7137ac0e-93d0-4ea7-c9b7-96ccb1751379"
rnn_gradient_check(test_mdl, X_test, y_test, initial_state)
# + [markdown] colab_type="text" id="tYTNSQwQfpE7"
# # Moving on to MNIST data
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 85, "output_extras": [{}]} colab_type="code" executionInfo={"elapsed": 7545, "status": "ok", "timestamp": 1503667860083, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-6znVyM1oxdg/AAAAAAAAAAI/AAAAAAAAABI/vEPo2Ce7Rpc/s50-c-k-no/photo.jpg", "userId": "102606466886131565871"}, "user_tz": -60} id="IyIHQpUGgXyM" outputId="f77b737f-d410-4a1a-fb44-2fdb96ff8f7c"
import tensorflow as tf
import numpy as np
import functools
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
# + [markdown] colab_type="text" id="esdB5QtJFixP"
# # Gated Cells (GRUs and LSTMs)
#
# Vanilla RNNs are very powerful sequence models. However, they are difficult to train, in large part due to the difficulties with getting gradients to propagate through all the time-steps without vanishing or exploding. If the gradient explodes then backpropagation will not work because we will get NaN values for the gradient contributions from earlier layers. The simplest trick to overcome this is called **gradient clipping** (see [Pascanu et al., 2013](http://www.jmlr.org/proceedings/papers/v28/pascanu13.pdf)). One basically rescales the gradients once their norms exceed a certain threshold. Dealing with vanishing gradients is trickier. Proper weight initialization helps to overcome this at the start of training (e.g [orthogonal initialization](https://www.tensorflow.org/api_docs/python/tf/orthogonal_initializer)), and there are regularization tricks for encouraging constant backwards error flow, which works for some tasks, but is not theoretically well-motivated.
#
# Gated models (we will look at the GRU and LSTM) modify the architecture of the RNN cell to ensure constant gradient propagation. The problem with the vanilla RNN is that its entire state is multiplicatively updated (overwritten) at every time-step (notice the $W_{hh}h_{t-1}$ term):
#
# $h_t = \sigma(W_{hh}h_{t-1} ...)$
#
# Gated cells (GRUs/LSTMs) have two main new ideas:
#
# * Ensure **incremental state change** by updating state *additively*: $h_t = h_{t-1} + f(h_{t-1})$.
# * Control the update process by selectively **modulating** how much to keep/forget of the old, how much to read, and how much to write into the new state.
#
# These models modulate how much to throw away (forget) from the previous state when making a proposed new state, and then how much to read from the previous state and write to the output of the cell, by using **gates** (values between 0 and 1 which squash information flow to some extent; little neural networks, of course!). Gates are vectors of *per-dimension interpolation scalars*: When you multiply some vector by a gate vector, you essentially control how much of that vector you "let through". Below we show the generic equation for such a gate (they're all the same!):
#
# $g_t = \sigma(W_g h_{t-1} + U_g x_t + b_g)$
#
# where $\sigma(z) = 1 / (1+e^{-z})$ is the sigmoid function (i.e. $0 \leq \sigma(z) \leq 1.$).
#
# Think about this for a second:
#
# * What are the inputs of this gate (model)?
# * What are the parameters of this gate (model)?
# * What does this remind you of?
#
# **NOTE**: Gates are just (vectors of) simple, logistic regression models which take inputs from the previous hidden layer $h_{t-1}$ and the current input layer $x_t$) and produce outputs between 0 and 1.
#
# Now let's use them to modulate the flow of information. We'll start with the GRU and build up to the LSTM.
#
#
# ## The Gated Recurrent Unit (GRU)
#
# The GRU was introduced in 2014 by Cho et al. -- almost 2 decades after the LSTM -- but we'll start here because the GRU is simpler than the LSTM. It uses only two gates per cell. First, it uses a **reset** gate to control how much of the previous state is used in computing the new proposed state:
#
# $\tilde{h_t} = \phi(W(r_t \bigodot h_{t-1}) + Ux_t + b)$
#
# (where $\bigodot$ is element-wise multiplication). Then it ties "reading" and "writing" into an **update** gate (by bounding their sums to 1, i.e. the more you read the less you write) when the new state is calculated:
#
# $h_t = z_t \bigodot h_{t-1} + (1 - z_t)\bigodot \tilde{h_t}$
#
# Try to reconcile the equations with the following flow diagram of the same:
#
# **TODO**: Include the picture for this: https://r2rt.com/static/images/NH_GRUCell.png
#
# * What happens when the reset gate is high/low?
# * What happens when the update gate is high/low?
# * How do these two interact?
# * Why would this architecture be more powerful than a vanilla RNN?
#
# ## The Long Short-Term Memory unit (LSTM)
#
# The LSTM was introduced in 1997 by Hochreiter and Schmidhuber. There are several different architectual variations 'out there', but they all operate by maintaining a separate **memory vector** $c_t$ and a **state vector** $h_t$ (i.e. the model computes a tuple of vectors per time-step, not just a single vector). We'll just focus on the `BasicLSTM` version now. It uses three gates, the **input**, **output** and **forget** gates:
#
# \begin{aligned}
# i_t &= \sigma(W_i h_{t-1} + U_i x_t + b_i) \\
# o_t &= \sigma(W_o h_{t-1} + U_o x_t + b_o) \\
# f_t &= \sigma(W_f h_{t-1} + U_f x_t + b_f) \\
# \end{aligned}
#
# Don't be intimidated by these equations. We've seen them all above already in their generic form. They're all just doing the same thing (computationally, not functionally). Convince yourself of this, by answering the following:
#
# * What is the **same** between these equations?
# * What is **different** between them?
#
# 1. First, the `BasicLSTM` Cell uses no gating to create the proposed new hidden state (original notation uses g, but I use tilde h):
#
# $\tilde{h}_t = \phi(W h_{t-1}) + Ux_t + b)$
#
# 2. Then it updates its internal memory to be a combination of the previous memory $c_{t-1}$ (multiplied/modulated by the forget gates $f_t$) and the new proposed state $\tilde{h}_t$ (modulated by the input gates $i_t$):
#
# $c_t = f_t \bigodot c_{t-1} + i_t \bigodot \tilde{h}_t$
#
# **Intuitively**: the model could choose to ignore old memory completely (if $f_t$ is all 1s), or ignore the newly proposed state completely ($i_t$ all 0s), but more likely it would learn to do something in-between. Think about how and why this is true.
#
# 3. Finally, the *state* that is actually output by the cell is a gated version of the memory vector, squashed by a tanh (because not everything in the memory cell might be immediately useful to the surrounding network):
#
# $ h_t = o_t \bigodot \phi(c_t) $
#
# *Phewwww*. We know.. there is a lot going on here! The following flow-diagram might help a bit to make this more clear:
#
# **TODO** Add picture: https://r2rt.com/static/images/NH_BasicLSTMCell.png
#
# Look at the equations and at the flow-diagram, and then try to answer the following questions:
#
# * How is the LSTM similar to the GRU?
# * How is it different?
# * What is the function of the memory vector (think about edge cases, e.g. where the forget gate is set to all 1s)?
# * Is the LSTM theoretically more powerful than the GRU? If so, why?
# * What is the computational drawback to using LSTMs (think about the number of gates; these must be parameterized..)?
# + [markdown] colab_type="text" id="B_eGri3h8St8"
# # Implementing Recurrent Models in TensorFlow
#
# In TensorFlow, we implement recurrent models using two building blocks:
#
# 1. A (graph) definition for the cell (you can use [those provided](https://www.tensorflow.org/versions/r1.0/api_guides/python/contrib.rnn#Base_interface_for_all_RNN_Cells), or you can write your own); and
# 2. A method to unroll the graph over the sequence (dynamic vs unrolled).
#
# First, TensorFlow provides implementations for many of the standard RNN cells. Poke around in the docs.
#
# Second, TensorFlow provides two different ways to implement the recurrence operations: **dynamic or static**. Basically, static unrolling prebuilds the entire unrolled RNN over the maximum number of time-steps. Dynamic unrolling dynamically creates the graph at each time-step, and saves the activations during the forward phase for the backward phase.
#
# **QUESTION**: Why do activations need to be saved during the forward phase? (HINT: Look at our use of `cache` in the Numpy code above).
#
# We will use dynamic unrolling: it uses less memory, and (counterintuitively), oftentimes it turns out to be faster.
# + [markdown] colab_type="text" id="-wnPo8X1gp1e"
# # An RNN Image Labeler
#
# Let's reuse the BaseSoftmaxClassifier from the previous practicals:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="9rk6sOnPf286"
class BaseSoftmaxClassifier(object):
def __init__(self, input_size, output_size, l2_lambda):
# Define the input placeholders. The "None" dimension means that the
# placeholder can take any number of images as the batch size.
self.x = tf.placeholder(tf.float32, [None, input_size], name='x')
self.y = tf.placeholder(tf.float32, [None, output_size], name='y')
self.input_size = input_size
self.output_size = output_size
self.l2_lambda = l2_lambda
self._all_weights = [] # Used to compute L2 regularization in compute_loss().
# You should override these in your build_model() function.
self.logits = None
self.predictions = None
self.loss = None
self.build_model()
def get_logits(self):
return self.logits
def build_model(self):
# OVERRIDE THIS FOR YOUR PARTICULAR MODEL.
raise NotImplementedError("Subclasses should implement this function!")
def compute_loss(self):
"""All models share the same softmax cross-entropy loss."""
assert self.logits is not None # Ensure that logits has been created!
data_loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits=self.logits, labels=self.y))
reg_loss = 0.
for w in self._all_weights:
reg_loss += tf.nn.l2_loss(w)
return data_loss + self.l2_lambda * reg_loss
def accuracy(self):
# Calculate accuracy.
assert self.predictions is not None # Ensure that pred has been created!
correct_prediction = tf.equal(tf.argmax(self.predictions, 1), tf.argmax(self.y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
return accuracy
# + [markdown] colab_type="text" id="ZNEifoNh8H58"
# We override build_model to build the graph for the RNN classifier.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="TUZ5BZUiggm9"
class RecurrentClassifier(BaseSoftmaxClassifier):
def __init__(self, model_params):
self.config = model_params
super(RecurrentClassifier, self).__init__(model_params['input_size'],
model_params['output_size'],
model_params['l2_lambda'])
def build_model(self):
assert self.config['num_steps'] * self.config['pixels_per_step'] == self.config['input_size']
# We break up the input images into num_steps groups of pixels_per_step
# pixels each.
rnn_input = tf.reshape(self.x, [-1,
self.config['num_steps'],
self.config['pixels_per_step']])
# Define the main RNN 'cell', that will be applied to each timestep.
cell = self.config['cell_fn'](self.config['memory_units'])
# NOTE: This is how we apply Dropout to RNNs.
cell = tf.contrib.rnn.DropoutWrapper(
cell,
output_keep_prob = self.config['dropout_keep_prob'])
cell = tf.contrib.rnn.MultiRNNCell(cells=[cell] * self.config['num_layers'],
state_is_tuple=True)
outputs, state = tf.nn.dynamic_rnn(cell,
rnn_input,
dtype=tf.float32)
# Transpose the cell to get the output from the last timestep for each batch.
output = tf.transpose(outputs, [1, 0, 2])
last_hiddens = tf.gather(output, int(output.get_shape()[0]) - 1)
# Define weights and biases for output prediction.
out_weights = tf.Variable(tf.random_normal([self.config['memory_units'],
self.config['output_size']]))
self._all_weights.append(out_weights)
out_biases = tf.Variable(tf.random_normal([self.config['output_size']]))
self.logits = tf.matmul(last_hiddens, out_weights) + out_biases
self.predictions = tf.nn.softmax(self.logits)
self.loss = self.compute_loss()
def get_logits(self):
return self.logits
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="wkk33_lk7Z-I"
class MNISTFraction(object):
"""A helper class to extract only a fixed fraction of MNIST data."""
def __init__(self, mnist, fraction):
self.mnist = mnist
self.num_images = int(mnist.num_examples * fraction)
self.image_data, self.label_data = mnist.images[:self.num_images], mnist.labels[:self.num_images]
self.start = 0
def next_batch(self, batch_size):
start = self.start
end = min(start + batch_size, self.num_images)
self.start = 0 if end == self.num_images else end
return self.image_data[start:end], self.label_data[start:end]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Jid1Ejbj7PO8"
def train_tf_model(tf_model,
session, # The active session.
num_epochs, # Max epochs/iterations to train for.
batch_size=50, # Number of examples per batch.
keep_prob=1.0, # (1. - dropout) probability, none by default.
train_only_on_fraction=1., # Fraction of training data to use.
optimizer_fn=None, # TODO(sgouws): more correct to call this optimizer_obj
report_every=1, # Report training results every nr of epochs.
eval_every=1, # Evaluate on validation data every nr of epochs.
stop_early=True, # Use early stopping or not.
verbose=True):
# Get the (symbolic) model input, output, loss and accuracy.
x, y = tf_model.x, tf_model.y
loss = tf_model.loss
accuracy = tf_model.accuracy()
# Compute the gradient of the loss with respect to the model parameters
# and create an op that will perform one parameter update using the specific
# optimizer's update rule in the direction of the gradients.
if optimizer_fn is None:
optimizer_fn = tf.train.AdamOptimizer()
optimizer_step = optimizer_fn.minimize(loss)
# Get the op which, when executed, will initialize the variables.
init = tf.global_variables_initializer()
# Actually initialize the variables (run the op).
session.run(init)
# Save the training loss and accuracies on training and validation data.
train_costs = []
train_accs = []
val_costs = []
val_accs = []
if train_only_on_fraction < 1:
mnist_train_data = MNISTFraction(mnist.train, train_only_on_fraction)
else:
mnist_train_data = mnist.train
prev_c_eval = 1000000
# Main training cycle.
for epoch in range(num_epochs):
avg_cost = 0.
avg_acc = 0.
total_batch = int(train_only_on_fraction * mnist.train.num_examples / batch_size)
# Loop over all batches.
for i in range(total_batch):
batch_x, batch_y = mnist_train_data.next_batch(batch_size)
# Run optimization op (backprop) and cost op (to get loss value),
# and compute the accuracy of the model.
feed_dict = {x: batch_x, y: batch_y}
if keep_prob < 1.:
feed_dict["keep_prob:0"] = keep_prob
_, c, a = session.run(
[optimizer_step, loss, accuracy], feed_dict=feed_dict)
# Compute average loss/accuracy
avg_cost += c / total_batch
avg_acc += a / total_batch
train_costs.append((epoch, avg_cost))
train_accs.append((epoch, avg_acc))
# Display logs per epoch step
if epoch % report_every == 0 and verbose:
print "Epoch:", '%04d' % (epoch+1), "Training cost =", \
"{:.9f}".format(avg_cost)
if epoch % eval_every == 0:
val_x, val_y = mnist.validation.images, mnist.validation.labels
feed_dict = {x : val_x, y : val_y}
if keep_prob < 1.:
feed_dict['keep_prob:0'] = 1.0
c_eval, a_eval = session.run([loss, accuracy], feed_dict=feed_dict)
if verbose:
print "Epoch:", '%04d' % (epoch+1), "Validation acc=", \
"{:.9f}".format(a_eval)
if c_eval >= prev_c_eval and stop_early:
print "Validation loss stopped improving, stopping training early after %d epochs!" % (epoch + 1)
break
prev_c_eval = c_eval
val_costs.append((epoch, c_eval))
val_accs.append((epoch, a_eval))
print "Optimization Finished!"
return train_costs, train_accs, val_costs, val_accs
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="5Gnj1bC67dwT"
# Helper functions to plot training progress.
from matplotlib import pyplot as plt
def my_plot(list_of_tuples):
"""Take a list of (epoch, value) and split these into lists of
epoch-only and value-only. Pass these to plot to make sure we
line up the values at the correct time-steps.
"""
plt.plot(*zip(*list_of_tuples))
def plot_multi(values_lst, labels_lst, y_label, x_label='epoch'):
# Plot multiple curves.
assert len(values_lst) == len(labels_lst)
plt.subplot(2, 1, 2)
for v in values_lst:
my_plot(v)
plt.legend(labels_lst, loc='upper left')
plt.xlabel(x_label)
plt.ylabel(y_label)
plt.show()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 856, "output_extras": [{}, {}, {}, {}]} colab_type="code" executionInfo={"elapsed": 235339, "status": "ok", "timestamp": 1503339496820, "user": {"displayName": "<NAME>", "photoUrl": <KEY>", "userId": "102606466886131565871"}, "user_tz": -60} id="SPfywldo94TN" outputId="d78319c0-0d58-43fe-ef57-8fca5dabeed7"
# %%time
def build_train_eval_and_plot(model_params, train_params, verbose=True):
tf.reset_default_graph()
m = Recurrentlassifier(model_params)
with tf.Session() as sess:
# Train model on the MNIST dataset.
train_losses, train_accs, val_losses, val_accs = train_tf_model(
m,
sess,
verbose=verbose,
**train_params)
# Now evaluate it on the test set:
accuracy_op = m.accuracy() # Get the symbolic accuracy operation
# Calculate the accuracy using the test images and labels.
accuracy = accuracy_op.eval({m.x: mnist.test.images,
m.y: mnist.test.labels})
if verbose:
print "Accuracy on test set:", accuracy
# Plot losses and accuracies.
plot_multi([train_losses, val_losses], ['train', 'val'], 'loss', 'epoch')
plot_multi([train_accs, val_accs], ['train', 'val'], 'accuracy', 'epoch')
ret = {'train_losses': train_losses, 'train_accs' : train_accs,
'val_losses' : val_losses, 'val_accs' : val_accs,
'test_acc' : accuracy}
return m, ret
#################################CODE TEMPLATE##################################
# Specify the model hyperparameters:
model_params = {
'input_size' : 784,
'output_size' : 10,
'batch_size' : 100,
'num_steps' : 28,
'pixels_per_step' : 28, # NOTE: num_steps * pixels_per_step must = input_size
'cell_fn' : tf.contrib.rnn.BasicRNNCell,
'memory_units' : 128,
'num_layers' : 1,
'l2_lambda' : 1e-3,
'dropout_keep_prob': 1.
}
# Specify the training hyperparameters:
training_params = {
'num_epochs' : 100, # Max epochs/iterations to train for.
'batch_size' : 100, # Number of examples per batch, 100 default.
#'keep_prob' : 1.0, # (1. - dropout) probability, none by default.
'train_only_on_fraction' : 1., # Fraction of training data to use, 1. for everything.
'optimizer_fn' : None, # Optimizer, None for Adam.
'report_every' : 1, # Report training results every nr of epochs.
'eval_every' : 1, # Evaluate on validation data every nr of epochs.
'stop_early' : True, # Use early stopping or not.
}
# Build, train, evaluate and plot the results!
trained_model, training_results = build_train_eval_and_plot(
model_params,
training_params,
verbose=True # Modify as desired.
)
###############################END CODE TEMPLATE################################
# + [markdown] colab_type="text" id="H-GfLCrGUwsD"
# ## Exercise: Try out different cells!
#
# Replace the BasicLSTM cell with something else.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="HVLm6vflU5v8"
# Your code here...
# + [markdown] colab_type="text" id="hIAnbWvA-uDq"
# ## Known good settings
#
# We got **98.8%** with this model and hyperparams:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="btJIqQJ5-05c"
# Specify the model hyperparameters:
model_params = {
'input_size' : 784,
'output_size' : 10,
'batch_size' : 100,
'num_steps' : 28,
'pixels_per_step' : 28, # NOTE: num_steps * pixels_per_step must = input_size
'cell_fn' : tf.contrib.rnn.BasicLSTMCell,
'memory_units' : 128,
'num_layers' : 1,
'l2_lambda' : 1e-3,
'dropout_keep_prob': 1.
}
# Specify the training hyperparameters:
training_params = {
'num_epochs' : 100, # Max epochs/iterations to train for.
'batch_size' : 100, # Number of examples per batch, 100 default.
#'keep_prob' : 1.0, # (1. - dropout) probability, none by default.
'train_only_on_fraction' : 1., # Fraction of training data to use, 1. for everything.
'optimizer_fn' : None, # Optimizer, None for Adam.
'report_every' : 1, # Report training results every nr of epochs.
'eval_every' : 1, # Evaluate on validation data every nr of epochs.
'stop_early' : True, # Use early stopping or not.
}
# Build, train, evaluate and plot the results!
trained_model, training_results = build_train_eval_and_plot(
model_params,
training_params,
verbose=True # Modify as desired.
)
# + [markdown] colab_type="text" id="HEHT4Ne4b07g"
# # Resources
#
# * https://r2rt.com/written-memories-understanding-deriving-and-extending-the-lstm.html
# * HipsterNet Code: https://github.com/wiseodd/hipsternet/blob/master/hipsternet/neuralnet.py
# * http://peterroelants.github.io/posts/rnn_implementation_part01/
# * http://karpathy.github.io/2015/05/21/rnn-effectiveness/
#
|
practical4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tvm
from tvm import relay
import numpy as np
# +
dshape = [1, 32, 64, 64]
kshape = [32, 32, 3, 3]
dtype = 'int16'
x = relay.var("x", shape=dshape, dtype=dtype)
w = relay.var("w", dtype=dtype)
y = relay.nn.bitserial_conv2d(x, w, channels=32)
func = relay.Function([x, w], y)
data = np.random.uniform(-2, 2, size=dshape).astype(dtype)
kernel = np.random.uniform(-2, 2, size=kshape).astype(dtype)
# -
with relay.build_config(opt_level=3):
graph, lib, params = relay.build(func, 'llvm', params={'w': kernel})
intrp1 = relay.create_executor('graph', ctx=tvm.cpu(), target='llvm')
op_res1 = intrp1.evaluate(func)(data, kernel)
|
notebooks/Relay/bitserial_conv_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Tutorial-IllinoisGRMHD: reconstruct_set_of_prims_PPM.C
#
# ## Authors: <NAME> & <NAME>
#
# <font color='red'>**This module is currently under development**</font>
#
# ## In this tutorial module we explain the piecewise parabolic method (PPM) used to reconstruct the primitive variables within IllinoisGRMHD
#
# ### Required and recommended citations:
#
# * **(Required)** <NAME>., <NAME>., <NAME>., <NAME>., and <NAME>. IllinoisGRMHD: an open-source, user-friendly GRMHD code for dynamical spacetimes. Class. Quantum Grav. 32 (2015) 175009. ([arxiv:1501.07276](http://arxiv.org/abs/1501.07276)).
# * **(Required)** <NAME>., <NAME>., <NAME>., <NAME>. Primitive Variable Solvers for Conservative General Relativistic Magnetohydrodynamics. Astrophysical Journal, 641, 626 (2006) ([astro-ph/0512420](https://arxiv.org/abs/astro-ph/0512420)).
# * **(Recommended)** <NAME>., <NAME>., <NAME>. An efficient shock-capturing central-type scheme for multidimensional relativistic flows - II. Magnetohydrodynamics. A&A 400 (2) 397-413 (2003). DOI: 10.1051/0004-6361:20021641 ([astro-ph/0210618](https://arxiv.org/abs/astro-ph/0210618)).
#
# If using the version of `IllinoisGRMHD` with piecewise polytropic *or* tabulated (coming soon!) EOS support, then the following citation is also required:
#
# * **(Required)** <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., and <NAME>., *IllinoisGRMHD github repository* (2019). Source Code URL: https://github.com/zachetienne/nrpytutorial/tree/master/IllinoisGRMHD/.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This module is organized as follows
#
# 0. [Step 0](#src_dir): **Source directory creation**
# 1. [Step 1](#introduction): **Introduction**
# 1. [Step 1.a](#ppm): *The Piecewise Parabolic Method (PPM)*
# 1. [Step 1.b](#loop_defines_reconstruction): *The `loop_defines_reconstruction.h` header file*
# 1. [Step 1.c](#preamble_reconstruct_set_of_prims_ppm): *The preamble to the `reconstruct_set_of_prims_PPM.C` code file*
# 1. [Step 2](#reconstruct_set_of_prims_ppm): **The `reconstruct_set_of_prims_PPM()` function**
# 1. [Step 2.a](#reading_the_input_gfs): *Reading the input gridfunctions*
# 1. [Step 2.b](#computation_of_du): *Evaluation of $\delta U_{i}$*
# 1. [Step 2.c](#computation_of_ur_and_ul): *Computing $U_{r}$ and $U_{l}$*
# 1. [Step 2.d](#steepening_rhob): *Steepening $\rho_{b}$*
# 1. [Step 2.e](#flattening_and_monotonizing): *Flattening and monotonizing*
# 1. [Step 2.f](#shifting_ur_and_ul): *Shifting $U_{r}$ and $U_{l}$*
# 1. [Step 3](#slope_limit): **The `slope_limit()` function**
# 1. [Step 4](#steepen_rho): **The `steepen_rho()` function**
# 1. [Step 5](#monotonize): **The `monotonize()` function**
# 1. [Step 6](#compute_p_cold__Gamma_cold): **The `compute_P_cold__Gamma_cold()` function**
# 1. [Step 7](#ftilde_gf_compute): **The `ftilde_gf_compute()` function**
# 1. [Step 8](#ftilde_compute): **The `ftilde_compute()` function**
# 1. [Step 9](#code_validation): **Code validation**
# 1. [Step 9.a](#loop_defines_reconstruction__h_validation): *`loop_defines_reconstruction.h`*
# 1. [Step 9.b](#reconstruct_set_of_prims_ppm__c_validation): *`reconstruct_set_of_prims_PPM.C`*
# 1. [Step 10](#latex_pdf_output): **Output this notebook to $\LaTeX$-formatted PDF file**
# <a id='src_dir'></a>
#
# # Step 0: Source directory creation \[Back to [top](#toc)\]
# $$\label{src_dir}$$
#
# We will now use the [cmdline_helper.py NRPy+ module](Tutorial-Tutorial-cmdline_helper.ipynb) to create the source directory within the `IllinoisGRMHD` NRPy+ directory, if it does not exist yet.
# +
# Step 0: Creation of the IllinoisGRMHD source directory
# Step 0a: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import os,sys
nrpy_dir_path = os.path.join("..","..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
# Step 0b: Load up cmdline_helper and create the directory
import cmdline_helper as cmd
outdir = os.path.join("..","src")
cmd.mkdir(outdir)
# -
# <a id='introduction'></a>
#
# # Step 1: Introduction \[Back to [top](#toc)\]
# $$\label{introduction}$$
#
# In this tutorial module we will go through the implementation of the piecewise parabolic method (PPM), introduced by [Colella & Woodward (1984)](https://crd.lbl.gov/assets/pubs_presos/AMCS/ANAG/A141984.pdf) (which shall henceforth be our main reference), used by `IllinoisGRMHD`.
# <a id='ppm'></a>
#
# ## Step 1.a: The Piecewise Parabolic Method (PPM) \[Back to [top](#toc)\]
# $$\label{ppm}$$
#
# The piecewise parabolic method (PPM) is an algorithm used to construct the values of primitive variables, $U$, at cell interfaces. The interpolation procedure alone can lead to unstable evolutions. To remedy this, we introduce three different techniques:
#
# 1. Steepening
# 1. Flatenning
# 1. Monotonizing
#
# These algorithms are intended to also produce narrower profiles near the vicinity of a shock. These steps tend to reduce the third-order accuracy of the interpolation code, but only in cases where a third-order interpolation algorithm would produce *worse* results (e.g. at local extrema).
#
# The algorithmic flow of the code is as follows:
#
# 1. **Read the input**: determine which primitives are to be "reconstructed" (i.e. interpolated)
# 2. **Slope-limited gradient**: this must be computed for each of the primitives that will be reconstructed. We have:
#
# $$
# \boxed{\delta U^{\rm slope-lim} \equiv
# \left\{
# \begin{matrix}
# {\rm sign}\left(\delta_{m} U_{i}\right)\min\left(\left|\delta_{m} U_{i}\right|,c\left|\delta U_{i}\right|,c\left|\delta U_{i+1}\right|\right) & ,\ {\rm if}\ dU_{i}dU_{i+1} > 0\\
# 0 &,\ {\rm otherwise}
# \end{matrix}
# \right.}\ ,
# $$
#
# where $\delta U^{\rm slope-lim}$ is referred to as the slope-limited gradient of $U$ and
#
# \begin{align}
# \delta U_{i} &\equiv U_{i} - U_{i-1}\ ,\\
# \delta_{m} U_{i} &\equiv \frac{\delta U_{i} + \delta U_{i+1}}{2} = \frac{U_{i+1} - U_{i-1}}{2}\ .
# \end{align}
#
# 3. **Perform the interpolation**: We wish to determine $U_{r}$ and $U_{l}$, the values of $U$ at the cell interfaces. From the given set of known values $\left\{U_{i-2},U_{i-1},U_{i},U_{i+1},U_{i+2}\right\}$, one *interpolates* (with third-order accuracy) the values
#
# $$
# \begin{matrix}
# U_{i+1/2} \equiv U_{r,i} = U_{i+0} + \frac{1}{2}\left(U_{i+1} - U_{i+0}\right) + \frac{1}{6}\left(\delta U^{\rm slope-lim}_{i+0} - \delta U^{\rm slope-lim}_{i+1}\right)\\
# U_{i-1/2} \equiv U_{l,i} = U_{i-1} + \frac{1}{2}\left(U_{i+0} - U_{i-1}\right) + \frac{1}{6}\left(\delta U^{\rm slope-lim}_{i-1} - \delta U^{\rm slope-lim}_{i+0}\right)
# \end{matrix}
# $$
#
# 4. **Compute $P_{\rm cold}$ and $\Gamma_{\rm cold}$**: In order to decide whether or not to apply the steepening procedure, we must evaluate the contact discontinuity condition:
#
# $$
# \boxed{\Gamma_{\rm cold} K_{0}\frac{\left|\rho_{i+1}-\rho_{i-1}\right|}{\min\left(\rho_{i+1},\rho_{i-1}\right)} \geq \frac{\left|\left(P_{\rm cold}\right)_{i+1}-\left(P_{\rm cold}\right)_{i-1}\right|}{\min\left[\left(P_{\rm cold}\right)_{i+1},\left(P_{\rm cold}\right)_{i-1}\right]}}\ ,
# $$
#
# with $K_{0}$ a problem dependent constant.
#
# 5. **Steepening**: *if necessary*, apply the steepening proceedure *only* to $U = \rho_{b}^{\color{red}\ddagger}$. This involves performing the replacement
#
# $$
# \boxed{
# \begin{matrix}
# \rho_{r}\to \rho_{r}(1-\eta) + \rho^{\rm MC}_{r}\eta\\
# \rho_{l}\to \rho_{l}(1-\eta) + \rho^{\rm MC}_{l}\eta
# \end{matrix}
# }\ ,
# $$
#
# where
#
# \begin{align}
# \rho^{\rm MC}_{r,i+1} &= \rho_{i+1} - \frac{1}{2}\delta\rho^{\rm slope-lim}_{i+1}\ ,\\
# \rho^{\rm MC}_{l,i+0} &= \rho_{i-1} + \frac{1}{2}\delta\rho^{\rm slope-lim}_{i-1}\ ,\\
# \eta_{i} &= \max\left\{0,\min\left[\eta_{1}\left(\tilde\eta_{i}-\eta_{2}\right),1\right]\right\}\ ,
# \end{align}
#
# with $\eta_{1}$ and $\eta_{2}$ constants and
#
# $$
# \tilde\eta_{i} =
# \left\{
# \begin{matrix}
# 0&, \ {\rm if}\ \delta\rho_{i} = 0\ ,\\
# -\frac{1}{6}\left(\frac{\delta^{2}\rho_{i+1} - \delta^{2}\rho_{i-1}}{2\delta\rho_{i}}\right)&, \ {\rm otherwise}\ ,
# \end{matrix}
# \right.
# $$
#
# and finally
#
# \begin{align}
# \delta\rho_{i+0} &= \frac{\rho_{i+1} - \rho_{i-1}}{2}\ ,\\
# \delta^{2}\rho_{i-1} &= \rho_{i+0} - 2\rho_{i-1} + \rho_{i-2}\ ,\\
# \delta^{2}\rho_{i+1} &= \rho_{i+2} - 2\rho_{i+1} + \rho_{i+0}\ .
# \end{align}
#
# $^{\color{red}\ddagger}$: note that the common notation is $\rho_{0}$, but we will use $\rho_{b}$ to represent the baryonic matter density.
#
# 6. **Flattening**: *if necessary*, apply the flattening procedure to *all* primitives which are to be reconstructed. In the flattening procedure we modify either $U_{r}$, $U_{l}$, or both of them, according to
#
# $$
# \boxed{
# \begin{matrix}
# U_{r,i+0} = U_{i+0}\tilde{f} + U_{r,i+0}\left(1-\tilde{f}\right)\\
# U_{l,i+0} = U_{i+0}\tilde{f} + U_{l,i+0}\left(1-\tilde{f}\right)
# \end{matrix}
# }\ ,
# $$
#
# where
#
# \begin{align}
# \tilde{f} &= \min\left[1,w\max\left(0,q_{1}\right)\right]\ ,\\
# w &=
# \left\{
# \begin{matrix}
# 1\ , &\ {\rm if}\ q_{2} > \epsilon_{2}\ {\rm and}\ q_{2}\left(v^{\rm flux\ dirn}_{i-1}-v^{\rm flux\ dirn}_{i+1}\right)>0\ \left({\rm inside\ shock}\right)\ ,\\
# 0\ , &\ {\rm otherwise}\ \left({\rm outside\ shock}\right)\ ,
# \end{matrix}
# \right.
# \end{align}
#
# and
#
# \begin{align}
# q_{1} &= \left(\frac{\delta P_{1}}{\delta P_{2}}-\omega_{1}\right)\omega_{2}\ ,\\
# q_{2} &= \frac{\left|\delta P_{1}\right|}{\min\left(P_{i+1},P_{i-1}\right)}\ ,
# \end{align}
#
# with $\omega_{1}$ and $\omega_{2}$ constants and $\delta P_{n} \equiv P_{i+n} - P_{i-n}$.
#
# 7. **Monotonizing**: *if necessary*, apply the monotonizing procedure to *all* primitives which are to be reconstructed. We check three different cases, modifying $U_{r}$ and $U_{l}$ as follows:
#
# $$
# \boxed{
# \begin{matrix}
# \text{Case 1: if}\ \left(U_{r} - U\right)\left(U - U_{l}\right)\leq 0 \ \text{then:}\
# \left\{
# \begin{matrix}
# U_{r} \to U\ ,\\
# \ U_{l}\to U\ .
# \end{matrix}
# \right.\\
# \text{Case 2: if}\ \delta U\left(U - \delta_{m}U\right) > \frac{\left(\delta U\right)^{2}}{6} \ \text{then:}\
# \left\{
# \begin{matrix}
# U_{r} \to U_{r}\ ,\\
# \ U_{l}\to 3U-2U_{r}\ .
# \end{matrix}
# \right.\\
# \text{Case 3: if}\ \delta U\left(U - \delta_{m}U\right) < -\frac{\left(\delta U\right)^{2}}{6} \ \text{then:}\
# \left\{
# \begin{matrix}
# U_{r} \to 3U-2U_{l}\ ,\\
# \ U_{l}\to U_{l}\ .
# \end{matrix}
# \right.
# \end{matrix}
# }\ ,
# $$
#
# where
#
# \begin{align}
# \delta U &\equiv U_{r} - U_{l}\ ,\\
# \delta_{m}U &\equiv \frac{U_{r} + U_{l}}{2}\ .
# \end{align}
#
# 8. **Index shifting**: shift the indices of $U_{r}$ and $U_{l}$. We have, at this point
#
# \begin{align}
# U_{r,i} = U_{i+1/2}\ ,\\
# U_{l,i} = U_{i-1/2}\ .
# \end{align}
#
# We then perform the following shift
#
# $$
# \boxed{
# \begin{matrix}
# U_{i-1/2+\epsilon} = U_{l,i}^{\rm old} = U_{r,i}^{\rm new}\ ,\\
# U_{i-1/2-\epsilon} = U_{r,i-1}^{\rm old} = U_{l,i}^{\rm new}\ .\\
# \end{matrix}
# }
# $$
# <a id='loop_defines_reconstruction'></a>
#
# ## Step 1.b: The `loop_defines_reconstruction.h` header file \[Back to [top](#toc)\]
# $$\label{loop_defines_reconstruction}$$
#
# This header file defines useful quantities to be used throughout the `reconstruct_set_of_prims_PPM.C` code. They are:
#
# 1. LOOP_DEFINE $\rightarrow$ sets up a loop over the $x$, $y$, and $z$ directions, including the ghostzones
# 1. SET_INDEX_ARRAYS $\rightarrow$ for a given direction, $x^{j}$, and coordinate range, $i\in\left[i_\min,i_\max\right]$, finds the appropriate array indices
# 1. SET_INDEX_ARRAYS_3DBLOCK $\rightarrow$ finds the appropriate array indices in all directions of a given range
# +
# %%writefile $outdir/loop_defines_reconstruction.h
#ifndef LOOP_DEFINES_RECONSTRUCTION_H_
#define LOOP_DEFINES_RECONSTRUCTION_H_
#define LOOP_DEFINE(gz_shift_lo,gz_shift_hi, ext,flux_dirn, ijkgz_lo_hi,gz_lo,gz_hi) \
for(int rr=1;rr<=3;rr++) { \
ijkgz_lo_hi[rr][0]= gz_lo[rr]; \
ijkgz_lo_hi[rr][1]=ext[rr-1]-gz_hi[rr]; \
} \
ijkgz_lo_hi[flux_dirn][0] += gz_shift_lo; \
ijkgz_lo_hi[flux_dirn][1] -= gz_shift_hi; \
/* The following line is valid C99 */ \
_Pragma("omp parallel for private(U,dU,slope_lim_dU,Ur,Ul)") \
for(int k=ijkgz_lo_hi[3][0];k<ijkgz_lo_hi[3][1];k++) \
for(int j=ijkgz_lo_hi[2][0];j<ijkgz_lo_hi[2][1];j++) \
for(int i=ijkgz_lo_hi[1][0];i<ijkgz_lo_hi[1][1];i++)
// This define only sets indices.
// FIXME: benchmark with and without the if() statement.
// FIXME: try without index_arr being defined in all directions.
#define SET_INDEX_ARRAYS(IMIN,IMAX,flux_dirn) \
int max_shift=(MAXNUMINDICES/2); \
/* DEBUGGING ONLY: if(IMIN<-max_shift || IMAX>max_shift) CCTK_VError(VERR_DEF_PARAMS,"FIX MAXNUMINDICES!"); */ \
int index_arr[4][MAXNUMINDICES]; \
for(int idx=IMIN;idx<=IMAX;idx++) { \
index_arr[flux_dirn][idx+max_shift]= \
CCTK_GFINDEX3D(cctkGH, \
i+idx*kronecker_delta[flux_dirn][0], \
j+idx*kronecker_delta[flux_dirn][1], \
k+idx*kronecker_delta[flux_dirn][2]); \
}
#define SET_INDEX_ARRAYS_3DBLOCK(IJKLOHI) \
int max_shift=(MAXNUMINDICES/2); \
int index_arr_3DB[MAXNUMINDICES][MAXNUMINDICES][MAXNUMINDICES]; \
for(int idx_k=IJKLOHI[4];idx_k<=IJKLOHI[5];idx_k++) for(int idx_j=IJKLOHI[2];idx_j<=IJKLOHI[3];idx_j++) for(int idx_i=IJKLOHI[0];idx_i<=IJKLOHI[1];idx_i++) { \
index_arr_3DB[idx_k+max_shift][idx_j+max_shift][idx_i+max_shift]=CCTK_GFINDEX3D(cctkGH,i+idx_i,j+idx_j,k+idx_k); \
}
#endif /* LOOP_DEFINES_RECONSTRUCTION_H_ */
# -
# <a id='preamble_reconstruct_set_of_prims_ppm'></a>
#
# ## Step 1.c: The preamble to the `reconstruct_set_of_prims_PPM.C` code file \[Back to [top](#toc)\]
# $$\label{preamble_reconstruct_set_of_prims_ppm}$$
#
# We then initialize the `reconstruct_set_of_prims_PPM.C` code file with a basic preamble, some simple definitions, and function headers. Notice that these functions are defined in the other Steps of this tutorial notebook. See the [table of contents](#toc) above for more information.
# +
# %%writefile $outdir/reconstruct_set_of_prims_PPM.C
/*****************************************
* PPM Reconstruction Interface.
* <NAME> (2013)
*
* This version of PPM implements the standard
* Colella & Woodward PPM, though modified as in GRHydro
* to have 3 ghostzones instead of 4.
*****************************************/
#define MINUS2 0
#define MINUS1 1
#define PLUS0 2
#define PLUS1 3
#define PLUS2 4
#define MAXNUMINDICES 5
// ^^^^^^^^^^^^^ Be _sure_ to define MAXNUMINDICES appropriately!
// You'll find the #define's for LOOP_DEFINE and SET_INDEX_ARRAYS inside:
#include "loop_defines_reconstruction.h"
static inline CCTK_REAL ftilde_compute(const int flux_dirn,const CCTK_REAL rho_b_atm,CCTK_REAL U[MAXNUMVARS][MAXNUMINDICES]);
static inline CCTK_REAL slope_limit(CCTK_REAL dU,CCTK_REAL dUp1);
static inline void steepen_rho(CCTK_REAL U[MAXNUMVARS][MAXNUMINDICES],CCTK_REAL slope_lim_dU[MAXNUMVARS][MAXNUMINDICES],
CCTK_REAL Gamma_th,CCTK_REAL P_cold,CCTK_REAL Gamma_cold,
CCTK_REAL *rho_br_ppm,CCTK_REAL *rho_bl_ppm);
static inline void compute_P_cold__Gamma_cold(CCTK_REAL rho_b,eos_struct &eos, CCTK_REAL &P_cold,CCTK_REAL &Gamma_cold);
static inline void monotonize(CCTK_REAL U,CCTK_REAL &Ur,CCTK_REAL &Ul);
# -
# <a id='reconstruct_set_of_prims_ppm'></a>
#
# # Step 2: The `reconstruct_set_of_prims_PPM()` function \[Back to [top](#toc)\]
# $$\label{reconstruct_set_of_prims_ppm}$$
#
# The `reconstruct_set_of_prims_PPM()` function receives as input the direction in which to perform the reconstruction ($\rm flux\_dirn$), the number of primitives which are to be reconstructed ($\rm num\_prims\_to\_reconstruct$), which primitives are to be reconstructed ($\rm which\_prims\_to\_reconstruct$), the equation of state ($\rm eos$), and the array which stores the primitives ($\rm in\_prims$). The reconstructed primitive values are then stored in the output arrays $\rm out\_prims\_r$ and $\rm out\_prims\_l$. **Note**: you can find more information on the ETK specific parameters (such as ${\rm cctkGH}$ and $\rm cctk\_lsh$) by looking at the [official Einstein Toolkit documentation](http://cactuscode.org/documentation/referencemanual/ReferenceManualch2.html).
#
# Notice that we will start a loop that runs from 0 to $\rm num\_prims\_to\_reconstruct$, meaning that the discussion here will apply to *each* of the primitives that we wish to reconstruct.
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
static void reconstruct_set_of_prims_PPM(const cGH *cctkGH,const int *cctk_lsh,const int flux_dirn,const int num_prims_to_reconstruct,const int *which_prims_to_reconstruct,eos_struct &eos,
gf_and_gz_struct *IN_PRIMS,gf_and_gz_struct *OUT_PRIMS_R,gf_and_gz_struct *OUT_PRIMS_L,CCTK_REAL *ftilde_gf, CCTK_REAL *temporary) {
DECLARE_CCTK_PARAMETERS;
CCTK_REAL U[MAXNUMVARS][MAXNUMINDICES],dU[MAXNUMVARS][MAXNUMINDICES],slope_lim_dU[MAXNUMVARS][MAXNUMINDICES],
Ur[MAXNUMVARS][MAXNUMINDICES],Ul[MAXNUMVARS][MAXNUMINDICES];
int ijkgz_lo_hi[4][2];
for(int ww=0;ww<num_prims_to_reconstruct;ww++) {
int whichvar=which_prims_to_reconstruct[ww];
if(IN_PRIMS[whichvar].gz_lo[flux_dirn]!=0 || IN_PRIMS[whichvar].gz_hi[flux_dirn]!=0) {
CCTK_VError(VERR_DEF_PARAMS,"TOO MANY GZ'S! WHICHVAR=%d: %d %d %d : %d %d %d DIRECTION %d",whichvar,
IN_PRIMS[whichvar].gz_lo[1],IN_PRIMS[whichvar].gz_lo[2],IN_PRIMS[whichvar].gz_lo[3],
IN_PRIMS[whichvar].gz_hi[1],IN_PRIMS[whichvar].gz_hi[2],IN_PRIMS[whichvar].gz_hi[3],flux_dirn);
}
# -
# <a id='reading_the_input_gfs'></a>
#
# ## Step 2.a: Reading the input gridfunctions \[Back to [top](#toc)\]
# $$\label{reading_the_input_gfs}$$
#
# We start by reading in the input primitive gridfunction and storing them to a variable $U$. Notice that for a given direction and a given point $i$, we will know: $\left\{U_{i-2},U_{i-1},U_{i},U_{i+1},U_{i+2}\right\}$.
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// *** LOOP 1: Interpolate to Ur and Ul, which are face values ***
// You will find that Ur depends on U at MINUS1,PLUS0, PLUS1,PLUS2, and
// Ul depends on U at MINUS2,MINUS1,PLUS0,PLUS1.
// However, we define the below loop from MINUS2 to PLUS2. Why not split
// this up and get additional points? The reason is that later on,
// Ur and Ul depend on ftilde, which is defined from MINUS2 to PLUS2,
// so we would lose those points anyway.
LOOP_DEFINE(2,2, cctk_lsh,flux_dirn, ijkgz_lo_hi,IN_PRIMS[whichvar].gz_lo,IN_PRIMS[whichvar].gz_hi) {
SET_INDEX_ARRAYS(-2,2,flux_dirn);
/* *** LOOP 1a: READ INPUT *** */
// Read in a primitive at all gridpoints between m = MINUS2 & PLUS2, where m's direction is given by flux_dirn. Store to U.
for(int ii=MINUS2;ii<=PLUS2;ii++) U[whichvar][ii] = IN_PRIMS[whichvar].gf[index_arr[flux_dirn][ii]];
# <a id='computation_of_du'></a>
#
# ## Step 2.b: Evaluation of $\delta U_{i}$ \[Back to [top](#toc)\]
# $$\label{computation_of_du}$$
#
# We will need $\delta U_{i} \equiv U_{i} - U_{i-1}$ in order to compute $U_{r}$ and $U_{l}$. We will then compute (notice the notation change $i\to i+0$ so that it is easier to understand the C code that follows):
#
# \begin{align}
# \delta U_{i-1} &= U_{i-1} - U_{i-2}\ ,\\
# \delta U_{i+0} &= U_{i+0} - U_{i-1}\ ,\\
# \delta U_{i+1} &= U_{i+1} - U_{i+0} \ ,\\
# \delta U_{i+2} &= U_{i+2} - U_{i+1}\ .
# \end{align}
#
# After evaluating the differences $\delta U$, we compute the slope-limited $\delta U$, $\delta U^{\rm slope-lim}$, using the [`slope_limit()` function](#slope_limit).
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
/* *** LOOP 1b: DO COMPUTATION *** */
/* First, compute simple dU = U(i) - U(i-1), where direction of i
* is given by flux_dirn, and U is a primitive variable:
* {rho_b,P,vx,vy,vz,Bx,By,Bz}. */
// Note that for Ur and Ul at i, we must compute dU(i-1),dU(i),dU(i+1),
// and dU(i+2)
dU[whichvar][MINUS1] = U[whichvar][MINUS1]- U[whichvar][MINUS2];
dU[whichvar][PLUS0] = U[whichvar][PLUS0] - U[whichvar][MINUS1];
dU[whichvar][PLUS1] = U[whichvar][PLUS1] - U[whichvar][PLUS0];
dU[whichvar][PLUS2] = U[whichvar][PLUS2] - U[whichvar][PLUS1];
//static int iii=14, jjj=14, kkk=13;
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("UUUU0 %d | %e %e %e %e %e\n",whichvar,U[whichvar][MINUS2],U[whichvar][MINUS1],U[whichvar][PLUS0],U[whichvar][PLUS1],U[whichvar][PLUS2]);
// Then, compute slope-limited dU, using MC slope limiter:
slope_lim_dU[whichvar][MINUS1]=slope_limit(dU[whichvar][MINUS1],dU[whichvar][PLUS0]);
slope_lim_dU[whichvar][PLUS0] =slope_limit(dU[whichvar][PLUS0], dU[whichvar][PLUS1]);
slope_lim_dU[whichvar][PLUS1] =slope_limit(dU[whichvar][PLUS1], dU[whichvar][PLUS2]);
# <a id='computation_of_ur_and_ul'></a>
#
# ## Step 2.c: Computing $U_{r}$ and $U_{l}$ \[Back to [top](#toc)\]
# $$\label{computation_of_ur_and_ul}$$
#
# We now compute $U_{r}$ and $U_{l}$. Keep in mind that $U_{r,i} = U_{i+1/2}$, while $U_{l,i} = U_{i-1/2}$. The implemented equation follows eq. A1 in [Duez *et al.* (2005)](http://arxiv.org/pdf/astro-ph/0503420.pdf), but with the standardd PPM coefficient of $\frac{1}{6}$ (i.e. eq. A1 with $\frac{1}{8}\to\frac{1}{6}$). Keep in mind that we simplify the equation slightly before implementing it:
#
# \begin{align}
# U_{r,i+0} &= U_{i+0} + \frac{1}{2}\left(U_{i+1} - U_{i+0}\right) + \frac{1}{6}\left(\delta U^{\rm slope-lim}_{i+0} - \delta U^{\rm slope-lim}_{i+1}\right) \Rightarrow \boxed{U_{r,i+0} = \frac{1}{2}\left(U_{i+1} + U_{i+0}\right) + \frac{1}{6}\left(\delta U^{\rm slope-lim}_{i+0} - \delta U^{\rm slope-lim}_{i+1}\right)}\ ,\\
# U_{l,i+0} &= U_{i-1} + \frac{1}{2}\left(U_{i+0} - U_{i-1}\right) + \frac{1}{6}\left(\delta U^{\rm slope-lim}_{i-1} - \delta U^{\rm slope-lim}_{i+0}\right) \Rightarrow \boxed{U_{l,i+0} = \frac{1}{2}\left(U_{i+0} + U_{i-1}\right) + \frac{1}{6}\left(\delta U^{\rm slope-lim}_{i-1} - \delta U^{\rm slope-lim}_{i+0}\right)}\ .
# \end{align}
#
# After this step, the values of $U_{r,l,i+0}$ are stored as outputs.
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// Finally, compute face values Ur and Ul based on the PPM prescription
// (Eq. A1 in http://arxiv.org/pdf/astro-ph/0503420.pdf, but using standard 1/6=(1.0/6.0) coefficient)
// Ur[PLUS0] represents U(i+1/2)
// We applied a simplification to the following line: Ur=U+0.5*(U(i+1)-U) + ... = 0.5*(U(i+1)+U) + ...
Ur[whichvar][PLUS0] = 0.5*(U[whichvar][PLUS1] + U[whichvar][PLUS0] ) + (1.0/6.0)*(slope_lim_dU[whichvar][PLUS0] - slope_lim_dU[whichvar][PLUS1]);
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("UURR0 %d | %e\n",whichvar,Ur[whichvar][PLUS0]);
// Ul[PLUS0] represents U(i-1/2)
// We applied a simplification to the following line: Ul=U(i-1)+0.5*(U-U(i-1)) + ... = 0.5*(U+U(i-1)) + ...
Ul[whichvar][PLUS0] = 0.5*(U[whichvar][PLUS0] + U[whichvar][MINUS1]) + (1.0/6.0)*(slope_lim_dU[whichvar][MINUS1] - slope_lim_dU[whichvar][PLUS0]);
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("UULL0 %d | %e\n",whichvar,Ul[whichvar][PLUS0]);
/* *** LOOP 1c: WRITE OUTPUT *** */
// Store right face values to {rho_br,Pr,vxr,vyr,vzr,Bxr,Byr,Bzr},
// and left face values to {rho_bl,Pl,vxl,vyl,vzl,Bxl,Byl,Bzl}
OUT_PRIMS_R[whichvar].gf[index_arr[flux_dirn][PLUS0]] = Ur[whichvar][PLUS0];
OUT_PRIMS_L[whichvar].gf[index_arr[flux_dirn][PLUS0]] = Ul[whichvar][PLUS0];
}
# <a id='steepening_rhob'></a>
#
# ## Step 2.d: Steepening $\rho_{b}$ \[Back to [top](#toc)\]
# $$\label{steepening_rhob}$$
#
# Following the procedure described in [Step 4](#steepen_rho), we will now steepen $\rho_{b}$ using the [`steepen_rho()` function](#steepen_rho). Keep in mind that although we will loop over all primitives which are set for reconstruction, the steepening procedure is applied to $\rho_{b}$ *only*.
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// *** LOOP 2: STEEPEN RHOB ***
// Note that this loop applies ONLY to RHOB.
if(whichvar==RHOB) {
LOOP_DEFINE(2,2, cctk_lsh,flux_dirn, ijkgz_lo_hi,IN_PRIMS[whichvar].gz_lo,IN_PRIMS[whichvar].gz_hi) {
SET_INDEX_ARRAYS(-2,2,flux_dirn);
// Set rho and P separately, since within this loop,
// 1) steepen_rho() depends on RHOB(MINUS2,MINUS1,PLUS0,PLUS1,PLUS2)
// Read in all primitives between MINUS2 & PLUS2. Store to U.
for(int ii=MINUS2;ii<=PLUS2;ii++) U[RHOB][ii] = IN_PRIMS[RHOB ].gf[index_arr[flux_dirn][ii]];
for(int ii=MINUS1;ii<=PLUS1;ii++) U[PRESSURE][ii] = IN_PRIMS[PRESSURE].gf[index_arr[flux_dirn][ii]];
Ur[RHOB][PLUS0] = OUT_PRIMS_R[RHOB].gf[index_arr[flux_dirn][PLUS0]];
Ul[RHOB][PLUS0] = OUT_PRIMS_L[RHOB].gf[index_arr[flux_dirn][PLUS0]];
dU[whichvar][MINUS1] = U[whichvar][MINUS1]- U[whichvar][MINUS2];
dU[whichvar][PLUS0] = U[whichvar][PLUS0] - U[whichvar][MINUS1];
dU[whichvar][PLUS1] = U[whichvar][PLUS1] - U[whichvar][PLUS0];
dU[whichvar][PLUS2] = U[whichvar][PLUS2] - U[whichvar][PLUS1];
slope_lim_dU[whichvar][MINUS1]=slope_limit(dU[whichvar][MINUS1],dU[whichvar][PLUS0]);
//slope_lim_dU[whichvar][PLUS0] =slope_limit(dU[whichvar][PLUS0], dU[whichvar][PLUS1]);
slope_lim_dU[whichvar][PLUS1] =slope_limit(dU[whichvar][PLUS1], dU[whichvar][PLUS2]);
// Steepen rho
// DEPENDENCIES: RHOB face values, RHOB(MINUS2,MINUS1,PLUS0,PLUS1,PLUS2), P(MINUS1,PLUS0,PLUS1), and slope_lim_dU[RHOB](MINUS1,PLUS1)
CCTK_REAL P_cold,Gamma_cold;
compute_P_cold__Gamma_cold(U[RHOB][PLUS0],eos, P_cold,Gamma_cold);
steepen_rho(U,slope_lim_dU, Gamma_th,P_cold,Gamma_cold, Ur[RHOB],Ul[RHOB]);
// Output rho
OUT_PRIMS_R[RHOB].gf[index_arr[flux_dirn][PLUS0]] = Ur[RHOB][PLUS0];
OUT_PRIMS_L[RHOB].gf[index_arr[flux_dirn][PLUS0]] = Ul[RHOB][PLUS0];
}
}
}
# <a id='flattening_and_monotonizing'></a>
#
# ## Step 2.e: Flattening and monotonizing \[Back to [top](#toc)\]
# $$\label{flattening_and_monotonizing}$$
#
# The flattening procedure modifies $U_{r}$ and $U_{l}$ via
#
# $$
# \boxed{
# \begin{matrix}
# U_{r,i+0} = U_{i+0}\tilde{f} + U_{r,i+0}\left(1-\tilde{f}\right)\\
# U_{l,i+0} = U_{i+0}\tilde{f} + U_{l,i+0}\left(1-\tilde{f}\right)
# \end{matrix}
# }\ ,
# $$
#
# where $\tilde{f}$ is computed by the `ftilde_compute()` function, described in [Step 8](#ftilde_compute).
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
/* ORIGINAL PPM REQUIRES AT LEAST 4 GHOSTZONES, which can add
* significantly to the size of AMR ref. boundaries.
* To reduce to 3 ghostzones, we comment the following lines out:
* if ((P[indexp1] - P[indexm1]) <= 0.0) {
* f = MAX(ftilde,ftilde_p1);
* } else {
* f = MAX(ftilde,ftilde_m1);
* }
*/
// *** LOOP 3: FLATTEN BASED ON FTILDE AND MONOTONIZE ***
for(int ww=0;ww<num_prims_to_reconstruct;ww++) {
int whichvar=which_prims_to_reconstruct[ww];
// ftilde() depends on P(MINUS2,MINUS1,PLUS1,PLUS2)
LOOP_DEFINE(2,2, cctk_lsh,flux_dirn, ijkgz_lo_hi,IN_PRIMS[whichvar].gz_lo,IN_PRIMS[whichvar].gz_hi) {
SET_INDEX_ARRAYS(0,0,flux_dirn);
U[whichvar][PLUS0] = IN_PRIMS[whichvar].gf[index_arr[flux_dirn][PLUS0]];
Ur[whichvar][PLUS0] = OUT_PRIMS_R[whichvar].gf[index_arr[flux_dirn][PLUS0]];
Ul[whichvar][PLUS0] = OUT_PRIMS_L[whichvar].gf[index_arr[flux_dirn][PLUS0]];
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("READ %d | %e %e %e \n",whichvar,U[whichvar][PLUS0],Ur[whichvar][PLUS0],Ul[whichvar][PLUS0]);
// ftilde_gf was computed in the function compute_ftilde_gf(), called before this routine
CCTK_REAL ftilde = ftilde_gf[index_arr[flux_dirn][PLUS0]];
// ...and then flatten (local operation)
Ur[whichvar][PLUS0] = U[whichvar][PLUS0]*ftilde + Ur[whichvar][PLUS0]*(1.0-ftilde);
Ul[whichvar][PLUS0] = U[whichvar][PLUS0]*ftilde + Ul[whichvar][PLUS0]*(1.0-ftilde);
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("FLAT %d | %e %e %e %e \n",whichvar,U[whichvar][PLUS0],Ur[whichvar][PLUS0],Ul[whichvar][PLUS0],ftilde);
# Furthermore, we apply the `monotonize()` function, as described in [Step 5](#monotonize).
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// Then monotonize
monotonize(U[whichvar][PLUS0],Ur[whichvar][PLUS0],Ul[whichvar][PLUS0]);
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("MONO %d | %e %e \n",whichvar,Ur[whichvar][PLUS0],Ul[whichvar][PLUS0]);
OUT_PRIMS_R[whichvar].gf[index_arr[flux_dirn][PLUS0]] = Ur[whichvar][PLUS0];
OUT_PRIMS_L[whichvar].gf[index_arr[flux_dirn][PLUS0]] = Ul[whichvar][PLUS0];
}
# <a id='shifting_ur_and_ul'></a>
#
# ## Step 2.f: Shifting $U_{r}$ and $U_{l}$ \[Back to [top](#toc)\]
# $$\label{shifting_ur_and_ul}$$
#
# At this point, we have
#
# \begin{align}
# U_{r,i} &= U_{i+1/2}\ ,\\
# U_{l,i} &= U_{i-1/2}\ .
# \end{align}
#
# To keep things consistent, we shift indices to get
#
# $$
# \boxed{
# \begin{matrix}
# U_{i-1/2+\epsilon} = U_{l,i}^{\rm old} = U_{r,i}^{\rm new}\ ,\\
# U_{i-1/2-\epsilon} = U_{r,i-1}^{\rm old} = U_{l,i}^{\rm new}\ ,\\
# \end{matrix}
# }
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// Ur depends on ftilde, which depends on points of U between MINUS2 and PLUS2
OUT_PRIMS_R[whichvar].gz_lo[flux_dirn]+=2;
OUT_PRIMS_R[whichvar].gz_hi[flux_dirn]+=2;
// Ul depends on ftilde, which depends on points of U between MINUS2 and PLUS2
OUT_PRIMS_L[whichvar].gz_lo[flux_dirn]+=2;
OUT_PRIMS_L[whichvar].gz_hi[flux_dirn]+=2;
}
// *** LOOP 4: SHIFT Ur AND Ul ***
/* Currently face values are set so that
* a) Ur(i) represents U(i+1/2), and
* b) Ul(i) represents U(i-1/2)
* Here, we shift so that the indices are consistent:
* a) U(i-1/2+epsilon) = oldUl(i) = newUr(i)
* b) U(i-1/2-epsilon) = oldUr(i-1) = newUl(i)
* Note that this step is not strictly necessary if you keep
* track of indices when computing the flux. */
for(int ww=0;ww<num_prims_to_reconstruct;ww++) {
int whichvar=which_prims_to_reconstruct[ww];
LOOP_DEFINE(3,2, cctk_lsh,flux_dirn, ijkgz_lo_hi,IN_PRIMS[whichvar].gz_lo,IN_PRIMS[whichvar].gz_hi) {
SET_INDEX_ARRAYS(-1,0,flux_dirn);
temporary[index_arr[flux_dirn][PLUS0]] = OUT_PRIMS_R[whichvar].gf[index_arr[flux_dirn][MINUS1]];
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("TEMP %d || %d %d %d || %e\n",whichvar,index_arr[flux_dirn][PLUS0],index_arr[flux_dirn][MINUS1],CCTK_GFINDEX3D(cctkGH,14,14,13),temporary[index_arr[flux_dirn][PLUS0]]);
}
LOOP_DEFINE(3,2, cctk_lsh,flux_dirn, ijkgz_lo_hi,IN_PRIMS[whichvar].gz_lo,IN_PRIMS[whichvar].gz_hi) {
SET_INDEX_ARRAYS(0,0,flux_dirn);
// Then shift so that Ur represents the gridpoint at i-1/2+epsilon,
// and Ul represents the gridpoint at i-1/2-epsilon.
// Ur(i-1/2) = Ul(i-1/2) = U(i-1/2+epsilon)
// Ul(i-1/2) = Ur(i+1/2 - 1) = U(i-1/2-epsilon)
OUT_PRIMS_R[whichvar].gf[index_arr[flux_dirn][PLUS0]] = OUT_PRIMS_L[whichvar].gf[index_arr[flux_dirn][PLUS0]];
OUT_PRIMS_L[whichvar].gf[index_arr[flux_dirn][PLUS0]] = temporary[index_arr[flux_dirn][PLUS0]];
//if( (i==iii) && (j==jjj) && (k==kkk) ) printf("LAST %d | %d %d %d || %d | %e %e %e \n",whichvar,i,j,k,flux_dirn,OUT_PRIMS_R[whichvar].gf[index_arr[flux_dirn][PLUS0]],OUT_PRIMS_L[whichvar].gf[index_arr[flux_dirn][PLUS0]],temporary[index_arr[flux_dirn][PLUS0]]);
}
// Ul was just shifted, so we lost another ghostzone.
OUT_PRIMS_L[whichvar].gz_lo[flux_dirn]+=1;
OUT_PRIMS_L[whichvar].gz_hi[flux_dirn]+=0;
// As for Ur, we didn't need to get rid of another ghostzone,
// but we did ... seems wasteful!
OUT_PRIMS_R[whichvar].gz_lo[flux_dirn]+=1;
OUT_PRIMS_R[whichvar].gz_hi[flux_dirn]+=0;
}
}
# <a id='slope_limit'></a>
#
# # Step 3: The `slope_limit()` function \[Back to [top](#toc)\]
# $$\label{slope_limit}$$
#
# We will now show the definition of $\delta U^{\rm slope-lim}$. The reason why we introduce this slope-limited procedure is twofold. First, it leads to steeper representations of discontinuities, and second it guarantees that $U_{i+1/2}$ lies inside the range $\left[U_{i},U_{i+1}\right]$.
#
# We start by defining
#
# \begin{align}
# \delta U_{i} &\equiv U_{i} - U_{i-1}\ ,\\
# \delta_{m} U_{i} &\equiv \frac{\delta U_{i} + \delta U_{i+1}}{2} = \frac{U_{i+1} - U_{i-1}}{2}\ .
# \end{align}
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// Set SLOPE_LIMITER_COEFF = 2.0 for MC, 1 for minmod
#define SLOPE_LIMITER_COEFF 2.0
//Eq. 60 in JOURNAL OF COMPUTATIONAL PHYSICS 123, 1-14 (1996)
// [note the factor of 2 missing in the |a_{j+1} - a_{j}| term].
// Recall that dU = U_{i} - U_{i-1}.
static inline CCTK_REAL slope_limit(CCTK_REAL dU,CCTK_REAL dUp1) {
if(dU*dUp1 > 0.0) {
//delta_m_U=0.5 * [ (u_(i+1)-u_i) + (u_i-u_(i-1)) ] = (u_(i+1) - u_(i-1))/2 <-- first derivative, second-order; this should happen most of the time (smooth flows)
CCTK_REAL delta_m_U = 0.5*(dU + dUp1);
# -
# Next we implement the slope-limited $\delta U$ as
#
# $$
# \boxed{\delta U^{\rm slope-lim} \equiv
# \left\{
# \begin{matrix}
# {\rm sign}\left(\delta_{m} U_{i}\right)\min\left(\left|\delta_{m} U_{i}\right|,c\left|\delta U_{i}\right|,c\left|\delta U_{i+1}\right|\right) & ,\ {\rm if}\ dU_{i}dU_{i+1} > 0\\
# 0 &,\ {\rm otherwise}
# \end{matrix}
# \right.}\ .
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// EXPLANATION OF BELOW LINE OF CODE.
// In short, sign_delta_a_j = sign(delta_m_U) = (0.0 < delta_m_U) - (delta_m_U < 0.0).
// If delta_m_U>0, then (0.0 < delta_m_U)==1, and (delta_m_U < 0.0)==0, so sign_delta_a_j=+1
// If delta_m_U<0, then (0.0 < delta_m_U)==0, and (delta_m_U < 0.0)==1, so sign_delta_a_j=-1
// If delta_m_U==0,then (0.0 < delta_m_U)==0, and (delta_m_U < 0.0)==0, so sign_delta_a_j=0
int sign_delta_m_U = (0.0 < delta_m_U) - (delta_m_U < 0.0);
//Decide whether to use 2nd order derivative or first-order derivative, limiting slope.
return sign_delta_m_U*MIN(fabs(delta_m_U),MIN(SLOPE_LIMITER_COEFF*fabs(dUp1),SLOPE_LIMITER_COEFF*fabs(dU)));
}
return 0.0;
}
# <a id='steepen_rho'></a>
#
# # Step 4: The `steepen_rho()` function \[Back to [top](#toc)\]
# $$\label{steepen_rho}$$
#
# The steepening procedure withing the PPM algorithm is applied only to $\rho_{b}$. The idea here is to produce narrower profiles near the vicinity of a contact discontinuity.
#
# **A NOTE ON NOTATION**: in the discussion below we will refer to $\rho$ as $\rho$ to keep the notation a bit lighter. No confusion should arise from this since there is no other quantity $\rho$ involved.
#
# We start the algorithm by computing
#
# \begin{align}
# \delta\rho_{i+0} &= \frac{\rho_{i+1} - \rho_{i-1}}{2}\ ,\\
# \delta^{2}\rho_{i-1} &= \rho_{i+0} - 2\rho_{i-1} + \rho_{i-2}\ ,\\
# \delta^{2}\rho_{i+1} &= \rho_{i+2} - 2\rho_{i+1} + \rho_{i+0}\ .
# \end{align}
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// standard Colella-Woodward parameters:
// K0 = 0.1d0, eta1 = 20.0, eta2 = 0.05, epsilon = 0.01d0
#define K0 0.1
#define ETA1 20.0
#define ETA2 0.05
#define EPSILON 0.01
static inline void steepen_rho(CCTK_REAL U[MAXNUMVARS][MAXNUMINDICES],CCTK_REAL slope_lim_dU[MAXNUMVARS][MAXNUMINDICES],CCTK_REAL Gamma_th,CCTK_REAL P_cold,CCTK_REAL Gamma_cold,
CCTK_REAL *rho_br_ppm,CCTK_REAL *rho_bl_ppm) {
// Next compute centered differences d RHOB and d^2 RHOB
CCTK_REAL d1rho_b = 0.5*(U[RHOB][PLUS1] - U[RHOB][MINUS1]);
CCTK_REAL d2rho_b_m1 = U[RHOB][PLUS0] - 2.0*U[RHOB][MINUS1] + U[RHOB][MINUS2];
CCTK_REAL d2rho_b_p1 = U[RHOB][PLUS2] - 2.0*U[RHOB][PLUS1] + U[RHOB][PLUS0];
# -
# Then we evaluate
#
# $$
# \Gamma = \left.\left(\frac{\partial P}{\partial\rho}\right)\middle/\left(\frac{P}{\rho}\right)\right. = \Gamma_{\rm th} + \left(\Gamma_{\rm cold} - \Gamma_{\rm th}\right)\frac{P_{\rm cold}}{P}\ .
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// Compute effective Gamma = (partial P / partial rho0)_s /(P/rho0)
CCTK_REAL Gamma = Gamma_th + (Gamma_cold-Gamma_th)*P_cold/U[PRESSURE][PLUS0];
# Next the contact discontinuity condition, eq. (3.2) of [Colella & Woodward (1984)](https://crd.lbl.gov/assets/pubs_presos/AMCS/ANAG/A141984.pdf), is checked:
#
# $$
# \Gamma K_{0}\frac{\left|\rho_{i+1}-\rho_{i-1}\right|}{\min\left(\rho_{i+1},\rho_{i-1}\right)} \geq \frac{\left|P_{i+1}-P_{i-1}\right|}{\min\left(P_{i+1},P_{i-1}\right)}\ ,
# $$
#
# where $K_{0}$ is a problem dependent constant. Keep in mind that we implement the quantity
#
# $$
# \boxed{{\rm contact\_discontinuity\_check} \equiv \Gamma K_{0}\left|\rho_{i+1}-\rho_{i-1}\right|\min\left(P_{i+1},P_{i-1}\right) - \left|P_{i+1}-P_{i-1}\right|\min\left(\rho_{i+1},\rho_{i-1}\right)}\ ,
# $$
#
# and verify whether ${\rm contact\_discontinuity\_check} \geq 0$ to verify the discontinuity condition. We also define the quantities
#
# $$
# \boxed{{\rm second\_deriv\_check} \equiv - \delta^{2}\rho_{i-1}\delta^{2}\rho_{i+1}}\ ,
# $$
#
# and
#
# $$
# \boxed{{\rm relative\_change\_check} \equiv 2\left|\delta\rho\right| - \epsilon\min\left(\rho_{i+1},\rho_{i-1}\right)}\ ,
# $$
#
# where again $\epsilon$ is a constant. The contact discontinuity condition is then satisfied when all three quantities inside boxes above are non-negative. When that is the case, we evaluate
#
# $$
# \boxed{\eta_{i} = \max\left\{0,\min\left[\eta_{1}\left(\tilde\eta_{i}-\eta_{2}\right),1\right]\right\}}\ ,
# $$
#
# where $\eta_{1}$ and $\eta_{2}$ are constants and
#
# $$
# \boxed{\tilde\eta_{i} =
# \left\{
# \begin{matrix}
# 0&, \ {\rm if}\ \delta\rho_{i} = 0\ ,\\
# -\frac{1}{6}\left(\frac{\delta^{2}\rho_{i+1} - \delta^{2}\rho_{i-1}}{2\delta\rho_{i}}\right)&, \ {\rm otherwise}\ .
# \end{matrix}
# \right.}
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
CCTK_REAL contact_discontinuity_check = Gamma*K0*fabs(U[RHOB][PLUS1]-U[RHOB][MINUS1])*
MIN(U[PRESSURE][PLUS1],U[PRESSURE][MINUS1])
-fabs(U[PRESSURE][PLUS1]-U[PRESSURE][MINUS1])*MIN(U[RHOB][PLUS1],U[RHOB][MINUS1]);
CCTK_REAL second_deriv_check = -d2rho_b_p1*d2rho_b_m1;
CCTK_REAL relative_change_check = fabs(2.0*d1rho_b) - EPSILON*MIN(U[RHOB][PLUS1],U[RHOB][MINUS1]);
if(contact_discontinuity_check >= 0.0 && second_deriv_check >= 0.0
&& relative_change_check >= 0.0) {
CCTK_REAL eta_tilde=0.0;
if (fabs(d1rho_b) > 0.0) {
eta_tilde = -(1.0/6.0)*(d2rho_b_p1-d2rho_b_m1)/(2.0*d1rho_b);
}
CCTK_REAL eta = MAX(0.0,MIN(ETA1*(eta_tilde - ETA2),1.0));
# We then apply the monotonized central (MC) scheme of [van Leer](https://www.sciencedirect.com/science/article/pii/002199917790095X) (see also [Step 3](#slope_limit) for a discussion on the quantities $\delta\rho^{\rm slope-lim}_{i}$ below),
#
# \begin{align}
# \rho^{\rm MC}_{r,i+1} &= \rho_{i+1} - \frac{1}{2}\delta\rho^{\rm slope-lim}_{i+1}\ ,\\
# \rho^{\rm MC}_{l,i+0} &= \rho_{i-1} + \frac{1}{2}\delta\rho^{\rm slope-lim}_{i-1}\ ,
# \end{align}
#
# so that, finally, the steepening algorithm sets
#
# $$
# \begin{matrix}
# \rho_{r}\to \rho_{r}(1-\eta) + \rho^{\rm MC}_{r}\eta\ ,\\
# \rho_{l}\to \rho_{l}(1-\eta) + \rho^{\rm MC}_{l}\eta\ ,
# \end{matrix}
# $$
#
# or, as implemented below:
#
# $$
# \boxed{\begin{matrix}
# \rho_{r,i+0}\rightarrow \rho_{r,i+0}\left(1-\eta_{i+0}\right) + \rho^{\rm MC}_{r,i+1}\eta_{i+0}\ ,\\
# \rho_{l,i+0}\rightarrow \rho_{r,i+0}\left(1-\eta_{i+0}\right) + \rho^{\rm MC}_{l,i+0}\eta_{i+0}\ .
# \end{matrix}}
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// Next compute Urp1 and Ul for RHOB, using the MC prescription:
// Ur_p1 = U_p1 - 0.5*slope_lim_dU_p1
CCTK_REAL rho_br_mc_p1 = U[RHOB][PLUS1] - 0.5*slope_lim_dU[RHOB][PLUS1];
// Ul = U_m1 + 0.5*slope_lim_dU_m1
// Based on this line of code, Ur[index] = a_j - \delta_m a_j / 2. (cf. Eq. 65 in Marti & Muller's "PPM Method for 1D Relativistic Hydro." paper)
// So: Ur[indexp1] = a_{j+1} - \delta_m a_{j+1} / 2. This is why we have rho_br_mc[indexp1]
CCTK_REAL rho_bl_mc = U[RHOB][MINUS1] + 0.5*slope_lim_dU[RHOB][MINUS1];
rho_bl_ppm[PLUS0] = rho_bl_ppm[PLUS0]*(1.0-eta) + rho_bl_mc*eta;
rho_br_ppm[PLUS0] = rho_br_ppm[PLUS0]*(1.0-eta) + rho_br_mc_p1*eta;
}
}
# <a id='monotonize'></a>
#
# # Step 5: The `monotonize()` function \[Back to [top](#toc)\]
# $$\label{monotonize}$$
#
# The value $U_{i+1/2}$ will be assigned to $U_{l,i}$ and $U_{r,i-1}$ for most values of $i$, but in some cases this would lead to incorrect interpolation results. Near discontinuities, the value of either $U_{l}$, $U_{r}$, or both needs to be adjusted.
#
# Consider, then, the following quantities:
#
# \begin{align}
# \delta U &\equiv U_{r} - U_{l}\ ,\\
# \delta_{m}U &\equiv \frac{U_{r} + U_{l}}{2}\ .
# \end{align}
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
static inline void monotonize(CCTK_REAL U,CCTK_REAL &Ur,CCTK_REAL &Ul) {
CCTK_REAL dU = Ur - Ul;
CCTK_REAL mU = 0.5*(Ur+Ul);
# -
# Then, following eq. (1.10) of [Colella & Woodward (1984)](https://crd.lbl.gov/assets/pubs_presos/AMCS/ANAG/A141984.pdf), we will check the following three cases:
#
# $$
# \boxed{\text{Case 1: if}\ \left(U_{r} - U\right)\left(U - U_{l}\right)\leq 0 \ \text{then:}\
# \left\{
# \begin{matrix}
# U_{r} \to U\ ,\\
# \ U_{l}\to U\ .
# \end{matrix}
# \right.}
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
if ( (Ur-U)*(U-Ul) <= 0.0) {
Ur = U;
Ul = U;
return;
}
# $$
# \boxed{\text{Case 2: if}\ \delta U\left(U - \delta_{m}U\right) > \frac{\left(\delta U\right)^{2}}{6} \ \text{then:}\
# \left\{
# \begin{matrix}
# U_{r} \to U_{r}\ ,\\
# \ U_{l}\to 3U-2U_{r}\ .
# \end{matrix}
# \right.}
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
if ( dU*(U-mU) > (1.0/6.0)*SQR(dU)) {
Ul = 3.0*U - 2.0*Ur;
return;
}
# $$
# \boxed{\text{Case 3: if}\ \delta U\left(U - \delta_{m}U\right) < -\frac{\left(\delta U\right)^{2}}{6} \ \text{then:}\
# \left\{
# \begin{matrix}
# U_{r} \to 3U-2U_{l}\ ,\\
# \ U_{l}\to U_{l}\ .
# \end{matrix}
# \right.}
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
if ( dU*(U-mU) < -(1.0/6.0)*SQR(dU)) {
Ur = 3.0*U - 2.0*Ul;
return;
}
}
# <a id='compute_p_cold__Gamma_cold'></a>
#
# # Step 6: The `compute_P_cold__Gamma_cold()` function \[Back to [top](#toc)\]
# $$\label{compute_p_cold__Gamma_cold}$$
#
# This part of the code evaluates $P_{\rm cold}$ and $\Gamma_{\rm cold}$ for the equations of state (EOS) presented in eqs. 13-16 of [Stephens *et al.* (2008)](http://arxiv.org/pdf/0802.0200.pdf).
#
# First, if $\rho_{b} = 0$, then $P_{\rm cold} = 0$ and $\Gamma_{\rm cold}$ simply receives its tabulated value.
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
static inline void compute_P_cold__Gamma_cold(CCTK_REAL rho_b,eos_struct &eos, CCTK_REAL &P_cold,CCTK_REAL &Gamma_cold) {
// This code handles equations of state of the form defined
// in Eqs 13-16 in http://arxiv.org/pdf/0802.0200.pdf
// Default in case rho_b == 0.0
if(rho_b==0.0) { P_cold = 0.0; Gamma_cold = eos.Gamma_ppoly_tab[0]; return; }
# -
# Next we consider the case where the EOS is given by a single-polytrope
#
# $$
# \boxed{P_{\rm cold} = \kappa \rho_{b}^{\Gamma_{\rm cold}}}\ ,
# $$
#
# and also the piecewise polytrope EOS
#
# $$
# \boxed{
# P_{\rm cold} =
# \left\{
# \begin{matrix}
# K_{0}\rho^{\Gamma_{0}} & , & \rho \leq \rho_{0}\\
# K_{1}\rho^{\Gamma_{1}} & , & \rho_{0} \leq \rho \leq \rho_{1}\\
# \vdots & & \vdots\\
# K_{j}\rho^{\Gamma_{j}} & , & \rho_{j-1} \leq \rho \leq \rho_{j}\\
# \vdots & & \vdots\\
# K_{N-1}\rho^{\Gamma_{N-1}} & , & \rho_{N-2} \leq \rho \leq \rho_{N-1}\\
# K_{N}\rho^{\Gamma_{N}} & , & \rho \geq \rho_{N-1}
# \end{matrix}
# \right.
# }\ .
# $$
#
# Notice that we left the fact that $\Gamma_{i} \equiv \Gamma_{{\rm cold},i}$ implicit above.
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
/***********************************
* Piecewise Polytropic EOS Patch *
* Computing P_cold and Gamma_cold *
***********************************/
int polytropic_index = find_polytropic_K_and_Gamma_index(eos,rho_b);
Gamma_cold = eos.Gamma_ppoly_tab[polytropic_index];
P_cold = eos.K_ppoly_tab[polytropic_index]*pow(rho_b,Gamma_cold);
}
# -
# <a id='ftilde_gf_compute'></a>
#
# # Step 7: The `ftilde_gf_compute()` function \[Back to [top](#toc)\]
# $$\label{ftilde_gf_compute}$$
#
# This is the driver function of the `ftilde_compute()` function, setting up the dependencies needed to compute $\tilde{f}$. Please refer to the next step to see how $\tilde{f}$ is computed.
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
#define OMEGA1 0.75
#define OMEGA2 10.0
#define EPSILON2 0.33
static void ftilde_gf_compute(const cGH *cctkGH,const int *cctk_lsh,const int flux_dirn,gf_and_gz_struct *in_prims,CCTK_REAL *ftilde_gf) {
DECLARE_CCTK_PARAMETERS;
int ijkgz_lo_hi[4][2];
CCTK_REAL U[MAXNUMVARS][MAXNUMINDICES];
/*Remove gcc unused variable warning/error Re: Pragma statement in loop define:*/
CCTK_REAL dU,slope_lim_dU,Ur,Ul; dU=slope_lim_dU=Ur=Ul=0.0; dU*=0;
// Compute ftilde, which is used for flattening left and right face values
LOOP_DEFINE(2,2, cctk_lsh,flux_dirn, ijkgz_lo_hi,in_prims[VX+(flux_dirn-1)].gz_lo,in_prims[VX+(flux_dirn-1)].gz_hi) {
SET_INDEX_ARRAYS(-2,2,flux_dirn);
for(int ii=MINUS2;ii<=PLUS2;ii++) U[RHOB][ii] = in_prims[RHOB ].gf[index_arr[flux_dirn][ii]];
for(int ii=MINUS2;ii<=PLUS2;ii++) U[PRESSURE][ii] = in_prims[PRESSURE].gf[index_arr[flux_dirn][ii]];
U[VX+(flux_dirn-1)][MINUS1] = in_prims[VX+(flux_dirn-1)].gf[index_arr[flux_dirn][MINUS1]];
U[VX+(flux_dirn-1)][PLUS1] = in_prims[VX+(flux_dirn-1)].gf[index_arr[flux_dirn][PLUS1]];
// Compute ftilde, which is used for flattening left and right face values
// DEPENDENCIES: P(MINUS2,MINUS1,PLUS1,PLUS2) and v^m(MINUS1,PLUS1), where m=flux_dirn={1,2,3}={x,y,z}.
ftilde_gf[index_arr[flux_dirn][PLUS0]] = ftilde_compute(flux_dirn,rho_b_atm,U);
}
}
# -
# <a id='ftilde_compute'></a>
#
# # Step 8: The `ftilde_compute()` function \[Back to [top](#toc)\]
# $$\label{ftilde_compute}$$
#
# We start by evaluating
#
# \begin{align}
# \delta P_{1} &\equiv P_{i+1} - P_{i-1}\ ,\\
# \delta P_{2} &\equiv P_{i+2} - P_{i-2}\ .
# \end{align}
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
static inline CCTK_REAL ftilde_compute(const int flux_dirn,const CCTK_REAL rho_b_atm,CCTK_REAL U[MAXNUMVARS][MAXNUMINDICES]) {
CCTK_REAL dP1 = U[PRESSURE][PLUS1] - U[PRESSURE][MINUS1];
CCTK_REAL dP2 = U[PRESSURE][PLUS2] - U[PRESSURE][MINUS2];
# -
# Then we modify the standard PPM algorithm slightly by introducing the following conditions:
#
# \begin{align}
# {\rm if}\ \left|\frac{\delta P_{1}}{\delta_{m}P_{1}}\right| = 0\ {\rm or}\ \left|\left(\rho_{b}\right)_{i+1}+\left(\rho_{b}\right)_{i-1}\right| < 10\rho_{\rm atm}\ {\rm then\ set}\ \delta P_{1}=0\ ,\\
# {\rm if}\ \left|\frac{\delta P_{2}}{\delta_{m}P_{2}}\right| = 0\ {\rm or}\ \left|\left(\rho_{b}\right)_{i+1}+\left(\rho_{b}\right)_{i-1}\right| < 10\rho_{\rm atm}\ {\rm then\ set}\ \delta P_{2}=0\ ,
# \end{align}
#
# where
#
# \begin{align}
# \delta_{m} P_{1} &\equiv \frac{P_{i+1} + P_{i-1}}{2}\ ,\\
# \delta_{m} P_{2} &\equiv \frac{P_{i+2} + P_{i-2}}{2}\ .
# \end{align}
#
# Note that if the first condition above is satisfied then we are *not* inside a shock, while if the second condition is triggered *alone* there *may* be a shock.
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// MODIFICATION TO STANDARD PPM:
// Cure roundoff error issues when dP1==0 or dP2==0 to 15 or more significant digits.
CCTK_REAL avg1=0.5*(U[PRESSURE][PLUS1] + U[PRESSURE][MINUS1]);
CCTK_REAL avg2=0.5*(U[PRESSURE][PLUS2] + U[PRESSURE][MINUS2]);
if(fabs(dP1)/avg1<1e-15 || fabs(U[RHOB][PLUS1] + U[RHOB][MINUS1])<10.0*rho_b_atm ) dP1=0.0; /* If this is triggered, there is NO shock. Also ignore "shocks" that appear entirely in the atmosphere (or regions with densities ~ 10*rho_b_atm). */
if(fabs(dP2)/avg2<1e-15 || fabs(U[RHOB][PLUS2] + U[RHOB][MINUS2])<10.0*rho_b_atm ) dP2=0.0; /* If this is triggered alone, there may be a shock. Otherwise if triggered with above, NO shock. Also ignore "shocks" that appear entirely in the atmosphere (or regions with densities ~ 10*rho_b_atm). */
# Next we set
#
# $$
# {\rm dP1\_over\_dP2} =
# \left\{
# \begin{matrix}
# \frac{\delta P_{1}}{\delta P_{2}} &,\ {\rm if}\ \delta P_{2} \neq 0\\
# 1 &,\ {\rm otherwise}
# \end{matrix}
# \right.
# $$
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
CCTK_REAL dP1_over_dP2=1.0;
if (dP2 != 0.0) dP1_over_dP2 = dP1/dP2;
# We then construct
#
# \begin{align}
# q_{1} &= \left({\rm dP1\_over\_dP2}-\omega_{1}\right)\omega_{2}\ ,\\
# q_{2} &= \frac{\left|\delta P_{1}\right|}{\min\left(P_{i+1},P_{i-1}\right)}\ .
# \end{align}
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
CCTK_REAL q1 = (dP1_over_dP2-OMEGA1)*OMEGA2;
CCTK_REAL q2 = fabs(dP1)/MIN(U[PRESSURE][PLUS1],U[PRESSURE][MINUS1]);
# We then initialize a new variable, $w$, to 0, i.e. we assume there is no shock. In the old version of `IllinoisGRMHD`, we would perform the following check:
#
# $$
# \text{OLD LOGIC: }w =
# \left\{
# \begin{matrix}
# 1\ , &\ {\rm if}\ q_{2} > \epsilon_{2}\ {\rm and}\ q_{2}\left(v^{\rm flux\ dirn}_{i-1}-v^{\rm flux\ dirn}_{i+1}\right)>0, & \left({\rm inside\ shock}\right)\ ,\\
# 0\ , &\ {\rm otherwise}, & \left({\rm outside\ shock}\right)\ ,
# \end{matrix}
# \right.
# $$
#
# where $v^{\rm flux\ dirn}$ represents either $v^{x}$, $v^{y}$, or $v^{z}$, depending on the flux direction. This would, then, set $w=1$ when the shock condition above is satisfied. However, upon revision, a more robust logic is the following:
#
# * Compute $\delta v \equiv v^{\rm flux\ dirn}_{i-1} - v^{\rm flux\ dirn}_{i+1}$.
# * Compute $\delta_{m} v \equiv v^{\rm flux\ dirn}_{i-1} + v^{\rm flux\ dirn}_{i+1}$.
# * Compute $v_{\rm rel\ err} \equiv \left|\delta v\right|/\delta_{m} v$.
#
# Then, the shock condition becomes:
#
# $$
# {\rm if}\ v_{\rm rel\ err} > 10^{-15}\ {\rm and}\ q_{2}>\epsilon_{2}\ {\rm and}\ \delta v > 0 \implies \left({\rm inside\ shock}\right).
# $$
#
# This is a more robust test of whether we are in a shock or not, because we avoid problems in the atmosphere where the values of $v^{\rm flux\ dirn}_{i\pm1}$ can differ only by machine precision and cause an undesired trigger of the shock flag. We also take advantage of the fact that $q_{2}$ is necessarily positive, by definition, so it was not helping in the second Boolean comparison of the previous logic.
#
# Finally, $\tilde{f}$ is given by
#
# $$
# \boxed{\tilde{f} = \min\left[1,w\max\left(0,q_{1}\right)\right]}\ .
# $$
# +
# %%writefile -a $outdir/reconstruct_set_of_prims_PPM.C
// w==0 -> NOT inside a shock
CCTK_REAL w=0.0;
// w==1 -> inside a shock
// Original code: if (q2 > EPSILON2 && q2*( (U[VX+(flux_dirn-1)][MINUS1]) - (U[VX+(flux_dirn-1)][PLUS1]) ) > 0.0) w = 1.0;
// comments: 1. q2 = (positive number) / (positive number) is always positive, so it's useless in the second Boolean comparison
// comments: 2. It may be the case that (U[VX+(flux_dirn-1)][MINUS1]) and (U[VX+(flux_dirn-1)][PLUS1]) differ at 1 part in 1e16 or so. We don't want the point to be marked as a shock due to this imperceptible difference.
// New code addresses above comments by removing the "q2*" and computing the relative error between the velocities.
const CCTK_REAL v_fluxdirn_minus1 = U[VX+(flux_dirn-1)][MINUS1];
const CCTK_REAL v_fluxdirn_plus1 = U[VX+(flux_dirn-1)][PLUS1];
const CCTK_REAL dv = v_fluxdirn_minus1 - v_fluxdirn_plus1;
const CCTK_REAL avgabsv = 0.5*(fabs(v_fluxdirn_minus1) + fabs(v_fluxdirn_plus1));
CCTK_REAL v_rel_error = 0.0; //dv/avgabsv;
if(avgabsv != 0) v_rel_error = fabs(dv)/avgabsv; // We should never flag a point as a shock if the velocities agree to
if(v_rel_error > 1e-15 && q2 > EPSILON2 && dv > 0.0) w = 1.0;
return MIN(1.0, w*MAX(0.0,q1));
}
# -
# <a id='code_validation'></a>
#
# # Step 9: Code validation \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# <a id='loop_defines_reconstruction__h_validation'></a>
#
# ## Step 9.a: `loop_defines_reconstruction.h` \[Back to [top](#toc)\]
# $$\label{loop_defines_reconstruction__h_validation}$$
# +
# # Verify if the code generated by this tutorial module
# # matches the original IllinoisGRMHD source code
# # First download the original IllinoisGRMHD source code
# import urllib
# from os import path
# original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/src/loop_defines_reconstruction.h"
# original_IGM_file_name = "loop_defines_reconstruction-original.h"
# original_IGM_file_path = os.path.join(IGM_src_dir_path,original_IGM_file_name)
# # Then download the original IllinoisGRMHD source code
# # We try it here in a couple of ways in an attempt to keep
# # the code more portable
# try:
# original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# # Write down the file the original IllinoisGRMHD source code
# with open(original_IGM_file_path,"w") as file:
# file.write(original_IGM_file_code)
# except:
# try:
# original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# # Write down the file the original IllinoisGRMHD source code
# with open(original_IGM_file_path,"w") as file:
# file.write(original_IGM_file_code)
# except:
# # If all else fails, hope wget does the job
# # !wget -O $original_IGM_file_path $original_IGM_file_url
# # Perform validation
# # Validation__loop_defines_reconstruction__h = !diff $original_IGM_file_path $outfile_path__loop_defines_reconstruction__h
# if Validation__loop_defines_reconstruction__h == []:
# # If the validation passes, we do not need to store the original IGM source code file
# # !rm $original_IGM_file_path
# print("Validation test for loop_defines_reconstruction.h: PASSED!")
# else:
# # If the validation fails, we keep the original IGM source code file
# print("Validation test for loop_defines_reconstruction.h: FAILED!")
# # We also print out the difference between the code generated
# # in this tutorial module and the original IGM source code
# print("Diff:")
# for diff_line in Validation__loop_defines_reconstruction__h:
# print(diff_line)
# -
# <a id='reconstruct_set_of_prims_ppm__c_validation'></a>
#
# ## Step 9.b: `reconstruct_set_of_prims_PPM.C` \[Back to [top](#toc)\]
# $$\label{reconstruct_set_of_prims_ppm__c_validation}$$
# +
# # Verify if the code generated by this tutorial module
# # matches the original IllinoisGRMHD source code
# # First download the original IllinoisGRMHD source code
# import urllib
# from os import path
# original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/src/reconstruct_set_of_prims_PPM.C"
# original_IGM_file_name = "reconstruct_set_of_prims_PPM-original.C"
# original_IGM_file_path = os.path.join(IGM_src_dir_path,original_IGM_file_name)
# # Then download the original IllinoisGRMHD source code
# # We try it here in a couple of ways in an attempt to keep
# # the code more portable
# try:
# original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# # Write down the file the original IllinoisGRMHD source code
# with open(original_IGM_file_path,"w") as file:
# file.write(original_IGM_file_code)
# except:
# try:
# original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# # Write down the file the original IllinoisGRMHD source code
# with open(original_IGM_file_path,"w") as file:
# file.write(original_IGM_file_code)
# except:
# # If all else fails, hope wget does the job
# # !wget -O $original_IGM_file_path $original_IGM_file_url
# # Perform validation
# # Validation__reconstruct_set_of_prims_PPM__C = !diff $original_IGM_file_path $outfile_path__reconstruct_set_of_prims_PPM__C
# if Validation__reconstruct_set_of_prims_PPM__C == []:
# # If the validation passes, we do not need to store the original IGM source code file
# # !rm $original_IGM_file_path
# print("Validation test for reconstruct_set_of_prims_PPM.C: PASSED!")
# else:
# # If the validation fails, we keep the original IGM source code file
# print("Validation test for reconstruct_set_of_prims_PPM.C: FAILED!")
# # We also print out the difference between the code generated
# # in this tutorial module and the original IGM source code
# print("Diff:")
# for diff_line in Validation__reconstruct_set_of_prims_PPM__C:
# print(diff_line)
# -
# <a id='latex_pdf_output'></a>
#
# # Step 10: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-IllinoisGRMHD_reconstruct_set_of_prims_PPM.pdf](Tutorial-IllinoisGRMHD_reconstruct_set_of_prims_PPM.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means).
latex_nrpy_style_path = os.path.join(nrpy_dir_path,"latex_nrpy_style.tplx")
# #!jupyter nbconvert --to latex --template $latex_nrpy_style_path --log-level='WARN' Tutorial-IllinoisGRMHD__reconstruct_set_of_prims_PPM.ipynb
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD_reconstruct__set_of_prims_PPM.tex
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD_reconstruct__set_of_prims_PPM.tex
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD_reconstruct__set_of_prims_PPM.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
|
IllinoisGRMHD/doc/Tutorial-IllinoisGRMHD__reconstruct_set_of_prims_PPM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Evaluación de modelos, métricas de puntuación y manejo de conjuntos de datos no balanceados.
# En los cuadernos anteriores, hemos detallado como evaluar un modelo y como escoger el mejor modelo posible. Hasta ahora, hemos asumido que nos proporcionaban una medida de rendimiento, algo para medir la calidad del modelo. Sin embargo, no siempre está claro cuál debería ser la medida a utilizar.
# Por defecto, en scikit-learn, se utiliza el ``accuracy`` para clasificación, que es el ratio de patrones correctamente clasificados y el $R^2$ para regresión, que es el coeficiente de determinación.
# Estas medidas son razonables para muchos escenarios. Sin embargo, dependiendo de la tarea que estemos tratando, estas no tienen porque ser las mejores opciones (y a veces pueden ser opciones muy poco recomendables).
# Vamos a centrarnos en la tarea de clasificación, volviendo de nuevo al problema de clasificación de dígitos manuscritos. Scikit-learn tiene métodos muy útiles en el paquete ``sklearn.metrics`` para ayudarnos a entrenar un clasificador y luego evaluarlo de distintas formas:
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
np.set_printoptions(precision=2)
# +
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split
from sklearn.svm import LinearSVC
digits = load_digits()
X, y = digits.data, digits.target
X_train, X_test, y_train, y_test = train_test_split(X, y,
random_state=1,
stratify=y,
test_size=0.25)
classifier = LinearSVC(random_state=1).fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
print("CCR: %f"%(classifier.score(X_test, y_test)))
# -
# Vemos que hemos predicho alrededor de un 95% de patrones de forma correcta. Para problemas multi-clase, a veces es muy útil saber qué clases son más difíciles de predecir y cuáles más fáciles o incluso qué tipo de errores son los más comunes. Una forma de tener más información en este sentido es la **matriz de confusión**, que muestra para cada clase (filas) cuántas veces se predicen qué clases (columnas).
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, y_test_pred)
# A veces un gráfico es más fácil de leer:
plt.imshow(confusion_matrix(y_test, y_test_pred), cmap="Blues")
plt.colorbar(shrink=0.8)
plt.xticks(range(10))
plt.yticks(range(10))
plt.xlabel("Etiqueta predicha")
plt.ylabel("Etiqueta real");
# Podemos ver que la mayoría de valores están en la diagonal principal, lo que significa que predecimos casi todos los ejemplos correctamente. Las entradas que no están en la diagonal principal nos muestran que hay bastantes ochos clasificados como unos, y que los nueves son fácilmente confundibles con el resto de clases.
# Otra función muy útil es ``classification_report`` que nos proporciona los valores de precisión, recall, puntuación f y el soporte para todas las clases. La precisión nos dice cuantas de las predicciones de una clase, son realmente de esa clase. Sea TP, FP, TN, FN "true positive" (verdaderos positivos), "false positive", (falsos positivos),"true negative" (verdaderos negativos) y "false negative" (falsos negativos), respectivamente:
# Precision = TP / (TP + FP)
# El recall representa cuantos ejemplos de la clase fueron clasificados correctamente (accuracy considerando solo esa clase):
# Recall = TP / (TP + FN)
# El valor F1 es la media geométrica de la precisión y el recall:
#
# F1 = 2 x (precision x recall) / (precision + recall)
# Todas estas métricas están en el intervalo $[0,1]$, donde un 1 es una puntuación perfecta.
from sklearn.metrics import classification_report
print(classification_report(y_test, y_test_pred))
# Estas métricas son especialmente útiles en dos casos particulares:
# 1. Clasificación no balanceada, es decir, una o varias clases son mucho menos frecuentes (hay menos casos en el conjunto de entrenamiento) que el resto de clases.
# 2. Costes asimétricos, esto es, algunos tipos de errores son más "costosos" que el resto.
# Vamos a ver el primer caso. Imagina que tenemos un ratio de 1:9 para un problema de clasificación (lo cuál no es muy exagerado, piensa por ejemplo en la predicción de clicks sobre banners de publicidad, donde a lo mejor solo un 0.001% de los anunciados son visitados):
np.bincount(y) / y.shape[0]
# Para probar este escenario, vamos a clasificar el dígito 3 contra el resto (el problema de clasificación es un problema binario, ¿es este dígito un 3?):
X, y = digits.data, digits.target == 3
# Ahora vamos a aplicar validación cruzada con un clasificador para ver que tal funciona:
# +
from sklearn.model_selection import cross_val_score
from sklearn.svm import SVC
cross_val_score(SVC(), X, y)
# -
# Nuestro clasificador tienen un 90% de acierto siempre. ¿Es bueno o malo? Ten en cuenta que el 90% de los dígitos no son un 3. Vamos a ver que tal funciona un clasificador simple, que siempre predice la clase más frecuenta (ZeroR):
from sklearn.dummy import DummyClassifier
cross_val_score(DummyClassifier("most_frequent"), X, y)
# También un 90%, como esperábamos. Por tanto, podemos pensar que el clasificador SVC no es demasiado bueno, ya que funciona igual que una estrategia que ni si quiera mira los datos de entrada. De todas formas, esto sería sacar conclusiones muy rápido ya que, en general, el accuracy no es una buena medida de rendimiento para bases de datos no balanceadas.
np.bincount(y) / y.shape[0]
# Curvas ROC
# =======
#
# Una medida mucho mejor se puede obtener utilizando las llamadas curvas de operación características (ROC, *Receiver operating characteristics*). Una curva ROC trabaja con las medidas de incertidumbre de un clasificador, por ejemplo la función de decisión de un ``SVC``. En lugar de utilizar el cero como umbral para distinguir ejemplos negativos de ejemplos positivos, la curva ROC considera todos los posibles umbrales y almacena el ratio de ejemplos de la clase positiva que se predicen correctamente (TPR) y el ratio de fallos para la clase negativa (FPR).
#
# El siguiente gráfico compara la curva ROC de tres configuraciones distintas de nuestro clasificador para la tarea "tres vs el resto".
# +
from sklearn.metrics import roc_curve, roc_auc_score
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
for gamma in [.01, .095, 1]:
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate (recall)")
svm = SVC(gamma=gamma).fit(X_train, y_train)
decision_function = svm.decision_function(X_test)
fpr, tpr, _ = roc_curve(y_test, decision_function)
acc = svm.score(X_test, y_test)
auc = roc_auc_score(y_test, svm.decision_function(X_test))
plt.plot(fpr, tpr, label="gamma: %.2f (acc:%.2f auc:%.2f)" % (gamma, acc, auc), linewidth=3)
plt.legend(loc="best");
# -
# Si el valor de umbral es muy bajo, tendremos muchos falsos positivos y por tanto un TPR muy alto y un FPR muy alto (porque casi todo lo clasificamos como positivo). Si usamos un umbral muy alto, habrá muy pocos falsos positivos (casi todo se predice como negativo), y por tanto el TPR será bajo y el FPR también. Por lo que, en general, la curva va desde arriba a la derecha hasta abajo a la izquierda. Una línea diagonal indica que el rendimiento es aleatorio, mientras que el objetivo ideal sería que la curva se desplace arriba a la izquierda. Esto significa que el clasificador daría siempre valores más altos de la función de decisión a los ejemplos positivos que a los ejemplos negativos.
#
# En este sentido, esta curva solo considera el orden asignado a los ejemplos positivos y negativos según la función de decisión, pero no el valor asignado. Como puedes ver a partir de las curvas y de los valores de accuracy, aunque todos los clasificadores tengan el mismo accuracy, uno de ellos tiene una curva ROC perfecta, mientras que otro se comporta igual que un clasificador aleatorio.
#
# Para realizar búsqueda en rejilla y validación cruzada, nos gustaría que la evaluación se guiase por un único valor numérico. Una buena forma de hacer esto es considera el área bajo la curva ROC (*area under the curve*, AUC). Podemos usar esto en ``cross_val_score`` especificando ``scoring="roc_auc"``:
from sklearn.model_selection import cross_val_score
cross_val_score(SVC(), X, y, scoring="roc_auc")
# Compara el rendimiento con el DummyClassifier:
from sklearn.linear_model import LogisticRegression
cross_val_score(LogisticRegression(), X, y, scoring="roc_auc")
# Funciones de rendimiento por defecto y personalizadas
# =======================================
# Hay muchas medidas de rendimiento, que son útiles para problemas muy distintos. Puedes encontrarlas en el diccionario "SCORERS". La documentación explica todas ellas.
from sklearn.metrics.scorer import SCORERS
print(SCORERS.keys())
# También es posible escribir tu propia medida de rendimiento. En lugar de una cadena, puedes pasar un nombre de función como argumento ``scoring``, esto es, un objeto con un método ``__call__`` (o lo que es lo mismo, una función). Esa función debe recibir un modelo, un conjunto de características ``X_test`` y un conjutno de etiquetas ``y_test``, y devolver un valor real. Los valores más altos deberían indicar que el modelo es mejor.
#
# Para probarlo, vamos a reimplementar la medida estándar de accuracy:
# +
def my_accuracy_scoring(est, X, y):
return np.mean(est.predict(X) == y)
cross_val_score(SVC(), X, y, scoring=my_accuracy_scoring)
# -
# <div class="alert alert-success">
# <b>EJERCICIO</b>:
# <ul>
# <li>
# En las secciones anteriores, normalmente usábamos el accuracy para evaluar el rendimiento de nuestros clasificadores. Una medida relacionada de la cuál no hemos hablado aún es el accuracy medio por clase (average-per-class accuracy, APCA). Como recordarás, el accuracy se puede definir como:
#
# $$ACC = \frac{TP+TN}{n},$$
#
# donde *n* es el número total de ejemplos. Esto puede generalizarse para multiclase como:
#
# $$ACC = \frac{T}{n},$$
#
# donde *T* es el número total de predicciones correctas (diagonal principal).
# </li>
# </ul>
# 
# <li>
# Dados los siguientes arrays de etiquetas verdaderas y de etiquetas predichas, ¿puedes implementar una función que utilice la métrica accuracy para conseguir el APCA?
# </li>
# </div>
# +
y_true = np.array([0, 0, 0, 1, 1, 1, 1, 1, 2, 2])
y_pred = np.array([0, 1, 1, 0, 1, 1, 2, 2, 2, 2])
confusion_matrix(y_true, y_pred)
# -
|
talleres_inov_docente/3-03-metricas_rendimiento_evaluacion_modelos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ORF307 Precept 8
# # Duality motivation example
#
# We want to obtain a lower bound for the following LP
#
# \begin{array}{ll} \mbox{minimize} & x_1 + 3x_2 \\
# \mbox{subject to} & x_1 + x_2 \geq 2 \\
# & x_2 \geq 1 \\
# & x_1 - x_2 \geq 3
# \end{array}
#
# One idea: add the constraints
# \begin{array}{ll}
# & y_1(x_1 + x_2 \geq 2) \\
# & + y_2(x_2 \geq 1) \\
# & + y_3(x_1 - x_2 \geq 3)
# \end{array}
#
# and match the cost
# \begin{array}{ll}
# & y_1 + y_3 = 1 \\
# & y_1 + y_2 - y_3 = 3 \\
# & y_1, y_2, y_3 \geq 0
# \end{array}
#
# We have many options
# \begin{array}{ll}
# & y = (1, 2, 0) \text{ -> bound is 4} \\
# & y = (0, 4, 1) \text{ -> bound is 7}
# \end{array}
# # Weak Duality
# The objective value of the dual problem is less than or equal to the objective of primal problem, i.e. $d^* \leq p^*$
# # Duality for linear programs
# For most linear programs, we see than in fact the objective values of the two problems are equal, $d^* = p^*$. We say that strong duality holds when this happens. For linear programs, there only $4$ possibilities.
#
# 1: Primal and dual optimal values finite and equal
#
# 2: Primal is infeasible and dual is unbounded
#
# 3: Primal is unbounded and dual is infeasible
#
# 4: Primal infeasible and dual is infeasible
#
#
#
#
# # <NAME>
# Given $A$ and $b$ exactly one of the two following statements is true
#
#
# 1: there exists an $x$ such that $Ax = b$, $x \geq 0$
#
#
# 2: there exists a $y$ such that $A^T y \geq 0$, $b^T y < 0$
# # The dual of our standard form LP
#
#
# \begin{array}{ll} \mbox{minimize} & c^T x \\
# \mbox{subject to} & Ax = b \\
# & x \geq 0 \\
# \end{array}
#
# \begin{array}{ll} \mbox{maximize} & -b^T y \\
# \mbox{subject to} & A^T + c \geq 0 \\
# \end{array}
# # Taking the dual example
#
# Consider the following LP. Find the dual optimization problem.
#
# \begin{array}{ll} \mbox{minimize} & 3x_1 + 4x_2 \\
# \mbox{subject to} & x_1 + x_2 \geq 5 \\
# & 2x_1 + x_2 \geq 6 \\
# &x_1, x_2 \geq 0
# \end{array}
#
# # Dual of 1 norm problem
#
# Consider the following optimization problem in $x$
#
# \begin{array}{ll} \mbox{minimize} & c^T x \\
# \mbox{subject to} & \| Ax + b\|_1 \leq 1 \end{array}
#
# (a) Formulate this as an LP in inequality form
#
# (b) Derive the dual LP, and show that it is equivalent to the problem
#
# \begin{array}{ll} \mbox{maximize} & b^T z - \| z \|_{\infty} \\
# \mbox{subject to} & A^Tz + c = 0 \end{array}
# (c) Give a direct argument that whenever $x$ is primal feasible and $z$ is dual feasible, we have $c^T x \geq b^T z - \| z\|_{\infty}$
|
precepts/08_precept/08_precept.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
x = np.linspace (0, 5, 11)
y = x ** 2
plt.plot(x,y)
plt.xlabel("X Label")
plt.ylabel("Y Label")
plt.title("Title - X vs Y")
plt.show()
plt.subplot(1,2,1)
plt.plot(x,y,'r')
plt.subplot(1,2,2)
plt.plot(y,x,'b')
fig = plt.figure()
ax1 = fig.add_axes([0,0,1,1])
ax1.set_xlabel("X label")
ax1.set_ylabel("Y label")
ax1.set_title("Title X vs Y")
ax1.plot(x,y)
fig = plt.figure()
ax1 = fig.add_axes([0,0,0.4,1])
ax2 = fig.add_axes([0.5,0.0,0.4,1])
ax1.set_title("Left Plot")
ax2.set_title("Right Plot")
ax1.plot(x,y)
ax2.plot(y,x)
fig = plt.figure()
fig, axes = plt.subplots(nrows = 1, ncols = 2)
for ax in axes:
ax.plot(x,y, label = "Squared")
axes[0].plot(x,y)
axes[1].plot(y,x, label = "Roots")
axes[1].legend(loc=0)
plt.tight_layout()
fig, axes = plt.subplots(nrows = 2, ncols = 1, figsize = (8,2))
axes[0].plot(x,y)
axes[1].plot(x,y)
plt.tight_layout()
fig.savefig("uselessgraphs.png", dpi=200)
# +
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.set_title("Colored Graph")
ax.set_xlim ([0,6])
ax.set_ylim ([0,26])
ax.plot(x,y,color = "green", linewidth = 3, linestyle = "-.",
alpha = 0.85, marker = "x", markersize = 15, markerfacecolor = "yellow",
markeredgecolor = "green")
fig.savefig("anotheruselessgraph",dpi = 600)
# -
|
Python/Basics/py_R3/.ipynb_checkpoints/Untitled5-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The collaborative filter approach focuses on finding users who have given similar ratings to the same books, thus creating a link between users, to whom will be suggested books that were reviewed in a positive way. In this way, we look for associations between users, not between books.
import numpy as np
import pandas as pd
import tensorflow as tf
from sklearn.preprocessing import MinMaxScaler
rating = pd.read_csv('data/BX-Book-Ratings.csv', sep=';', error_bad_lines=False, encoding="latin-1")
user = pd.read_csv('data/BX-Users.csv', sep=';', error_bad_lines=False, encoding="latin-1")
book = pd.read_csv('data/BX-Books.csv', sep=';', error_bad_lines=False, encoding="latin-1")
book_rating = pd.merge(rating, book, on='ISBN')
cols = ['Year-Of-Publication', 'Publisher', 'Book-Author', 'Image-URL-S', 'Image-URL-M', 'Image-URL-L']
book_rating.drop(cols, axis=1, inplace=True)
book_rating.head()
book_rating.head(3)
rating_count = (book_rating.
groupby(by = ['Book-Title'])['Book-Rating'].
count().
reset_index().
rename(columns = {'Book-Rating': 'RatingCount_book'})
[['Book-Title', 'RatingCount_book']]
)
rating_count.head()
threshold = 25
rating_count = rating_count.query('RatingCount_book >= @threshold')
rating_count.head()
book_rating.head(3)
user_rating = pd.merge(rating_count, book_rating, left_on='Book-Title', right_on='Book-Title', how='left')
user_rating.head(3)
user_count = (user_rating.
groupby(by = ['User-ID'])['Book-Rating'].
count().
reset_index().
rename(columns = {'Book-Rating': 'RatingCount_user'})
[['User-ID', 'RatingCount_user']]
)
user_count.head()
threshold = 20
user_count = user_count.query('RatingCount_user >= @threshold')
user_count.head()
combined = user_rating.merge(user_count, left_on = 'User-ID', right_on = 'User-ID', how = 'inner')
combined.head(3)
combined.shape
print('Number of unique books: ', combined['Book-Title'].nunique())
print('Number of unique users: ', combined['User-ID'].nunique())
# Normalize the ratings.
scaler = MinMaxScaler()
combined['Book-Rating'] = combined['Book-Rating'].values.astype(float)
rating_scaled = pd.DataFrame(scaler.fit_transform(combined['Book-Rating'].values.reshape(-1,1)))
combined['Book-Rating'] = rating_scaled
# Abd build the user book matrix.
# +
combined = combined.drop_duplicates(['User-ID', 'Book-Title'])
user_book_matrix = combined.pivot(index='User-ID', columns='Book-Title', values='Book-Rating')
user_book_matrix.fillna(0, inplace=True)
users = user_book_matrix.index.tolist()
books = user_book_matrix.columns.tolist()
user_book_matrix = user_book_matrix.as_matrix()
# -
# tf.placeholder only available in v1, so we have to work around.
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
# We will initialize the TensorFlow placeholder. Then, weights and biases are randomly initialized, the following code are taken from the book: Python Machine Learning Cook Book - Second Edition
# +
num_input = combined['Book-Title'].nunique()
num_hidden_1 = 10
num_hidden_2 = 5
X = tf.placeholder(tf.float64, [None, num_input])
weights = {
'encoder_h1': tf.Variable(tf.random_normal([num_input, num_hidden_1], dtype=tf.float64)),
'encoder_h2': tf.Variable(tf.random_normal([num_hidden_1, num_hidden_2], dtype=tf.float64)),
'decoder_h1': tf.Variable(tf.random_normal([num_hidden_2, num_hidden_1], dtype=tf.float64)),
'decoder_h2': tf.Variable(tf.random_normal([num_hidden_1, num_input], dtype=tf.float64)),
}
biases = {
'encoder_b1': tf.Variable(tf.random_normal([num_hidden_1], dtype=tf.float64)),
'encoder_b2': tf.Variable(tf.random_normal([num_hidden_2], dtype=tf.float64)),
'decoder_b1': tf.Variable(tf.random_normal([num_hidden_1], dtype=tf.float64)),
'decoder_b2': tf.Variable(tf.random_normal([num_input], dtype=tf.float64)),
}
# -
# Now, we can build the encoder and decoder model, as follows:
# +
def encoder(x):
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(x, weights['encoder_h1']), biases['encoder_b1']))
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, weights['encoder_h2']), biases['encoder_b2']))
return layer_2
def decoder(x):
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(x, weights['decoder_h1']), biases['decoder_b1']))
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, weights['decoder_h2']), biases['decoder_b2']))
return layer_2
# -
# We will construct the model and the predictions
# +
encoder_op = encoder(X)
decoder_op = decoder(encoder_op)
y_pred = decoder_op
y_true = X
# -
# define loss function and optimizer, and minimize the squared error, and define the evaluation metrics
loss = tf.losses.mean_squared_error(y_true, y_pred)
optimizer = tf.train.RMSPropOptimizer(0.03).minimize(loss)
eval_x = tf.placeholder(tf.int32, )
eval_y = tf.placeholder(tf.int32, )
pre, pre_op = tf.metrics.precision(labels=eval_x, predictions=eval_y)
# Initialize the variables. Because TensorFlow uses computational graphs for its operations, placeholders and variables must be initialized.
init = tf.global_variables_initializer()
local_init = tf.local_variables_initializer()
pred_data = pd.DataFrame()
# We can finally start to train our model.
#
# We split training data into batches, and we feed the network with them.
#
# We train our model with vectors of user ratings, each vector represents a user and each column a book, and entries are ratings that the user gave to books.
#
# After a few trials, I discovered that training model for 5 epochs with a batch size of 10 would be consum enough memory. This means that the entire training set will feed our neural network 20 times, every time using 50 users.
with tf.Session() as session:
epochs = 100
batch_size = 35
session.run(init)
session.run(local_init)
num_batches = int(user_book_matrix.shape[0] / batch_size)
user_book_matrix = np.array_split(user_book_matrix, num_batches)
for i in range(epochs):
avg_cost = 0
for batch in user_book_matrix:
_, l = session.run([optimizer, loss], feed_dict={X: batch})
avg_cost += l
avg_cost /= num_batches
print("epoch: {} Loss: {}".format(i + 1, avg_cost))
user_book_matrix = np.concatenate(user_book_matrix, axis=0)
preds = session.run(decoder_op, feed_dict={X: user_book_matrix})
pred_data = pred_data.append(pd.DataFrame(preds))
pred_data = pred_data.stack().reset_index(name='Book-Rating')
pred_data.columns = ['User-ID', 'Book-Title', 'Book-Rating']
pred_data['User-ID'] = pred_data['User-ID'].map(lambda value: users[value])
pred_data['Book-Title'] = pred_data['Book-Title'].map(lambda value: books[value])
keys = ['User-ID', 'Book-Title']
index_1 = pred_data.set_index(keys).index
index_2 = combined.set_index(keys).index
top_ten_ranked = pred_data[~index_1.isin(index_2)]
top_ten_ranked = top_ten_ranked.sort_values(['User-ID', 'Book-Rating'], ascending=[True, False])
top_ten_ranked = top_ten_ranked.groupby('User-ID').head(10)
top_ten_ranked.loc[top_ten_ranked['User-ID'] == 278582]
book_rating.loc[book_rating['User-ID'] == 278582].sort_values(by=['Book-Rating'], ascending=False)
|
Collaborative Filtering Model with TensorFlow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: super-duper-fiesta_env
# language: python
# name: super-duper-fiesta_env
# ---
import matplotlib.pyplot as plt
import numpy as np
from scipy.optimize import curve_fit
atm_neutrino_fluxes = "./../data/atm_flux.txt"
cross_sections = "./../data/nu_xs.txt"
def read_flux_file(f):
d={}
log_e, nue, numu, nuebar, numubar = ([] for i in range (5))
File = open(f,"r")
lines = File.readlines()
for line in lines:
columns = line.split(' ')
log_e.append(float(columns[0]))
nue.append(float(columns[1]))
numu.append(float(columns[2]))
nuebar.append(float(columns[3]))
numubar.append(float(columns[4]))
d["log_E"]=np.array(log_e)
d["E"]=np.power(10, np.array(log_e))
d["nu_e"]=np.array(nue)
d["nu_mu"]=np.array(numu)
d["nu_e_bar"]=np.array(nuebar)
d["nu_mu_bar"]=np.array(numubar)
File.close()
return d
def read_xs_file(f):
d={}
log_e, cc_nu, nc_nu, cc_nu_bar, nc_nu_bar = ([] for i in range(5))
File = open(f,"r")
lines = File.readlines()
for line in lines:
columns = line.split(' ')
log_e.append(float(columns[0]))
cc_nu.append(float(columns[1]))
nc_nu.append(float(columns[2]))
cc_nu_bar.append(float(columns[3]))
nc_nu_bar.append(float(columns[4]))
d["log_E"]=np.array(log_e)
d["E"]=np.power(10, np.array(log_e))
d["cc_nu"]=np.array(cc_nu)
d["nc_nu"]=np.array(nc_nu)
d["cc_nu_bar"]=np.array(cc_nu_bar)
d["nc_nu_bar"]=np.array(nc_nu_bar)
File.close()
return d
flux_d = read_flux_file(atm_neutrino_fluxes)
# # Astrophysical neutrino flux
#
# We use the astrophysical flux detected by IceCube, as reported in this paper: https://arxiv.org/pdf/2001.09520.pdf
#
# There, the simplest parameterisation of the astrophysical neutrino flux is given by the following expression
#
# $\frac{\Phi _{astro} ^{\nu + \bar{\nu}}(E)}{C_0} = \phi _{astro}\cdot \left(\frac{E}{E_0}\right) ^{-\gamma}$
#
# where $C_0 = 3\cdot 10^{-18} \text{GeV} ^{-1} \cdot \text{cm} ^{-2}\cdot \text{s} ^{-1}\cdot \text{sr} ^{-1}$
#
# and $E_0=100 \text{TeV}$.
#
# The values of the spectral index and flux normalisation which best fit the icecube data are $\gamma = 2.53$ and $\phi _{astro} = 1.66$.
#
# In this fit, it is assumed that every neutrino flavor conttributes equally to the flux.
# +
# best fit 2020
C0 = 3e-18
E0 = 100e3
gamma = 2.53
phi = 1.66
# best fit 2016
# C0 = 1e-18
# E0 = 100e3
# gamma = 2.13
# phi = 0.9
Phi = lambda x : C0 * phi * np.power((x/E0), -gamma)
# -
plt.yscale("log")
plt.plot(flux_d["log_E"], Phi(flux_d["E"]))
xs_d = read_xs_file(cross_sections)
# +
plt.yscale("log")
factor = 1.e-4/(4*np.pi)
# plt.plot(flux_d["log_E"],flux_d["nu_mu"]*factor, label=r'$\nu_{\mu}$')
# plt.plot(flux_d["log_E"],flux_d["nu_e"]*factor, label=r'$\nu_{e}$')
# plt.plot(flux_d["log_E"],flux_d["nu_mu_bar"]*factor, label=r'$\bar{\nu}_{\mu}$')
# plt.plot(flux_d["log_E"],flux_d["nu_e_bar"]*factor, label=r'$\bar{\nu}_{e}$')
plt.plot(flux_d["log_E"],factor*(flux_d["nu_mu"] + flux_d["nu_e"] + flux_d["nu_mu_bar"] + flux_d["nu_e_bar"]), label = 'total atm')
plt.plot(flux_d["log_E"], Phi(flux_d["E"]), label='astro')
plt.legend()
#flux_d["nu_mu"]
# -
def Phi_atm(x, phi_atm, gamma_atm):
return phi_atm * np.power((x), -gamma_atm)
a,b = curve_fit(Phi_atm, flux_d["E"][800:], flux_d["nu_mu"][800:], maxfev=2000 )
plt.yscale("log")
plt.xscale("log")
plt.plot(flux_d["E"][800:], Phi_atm(flux_d["E"][800:], a[0], a[1]), label='Fitted function')
plt.plot(flux_d["E"],flux_d["nu_mu"], linewidth=3, alpha=0.5, label="data")
log_Emin = 2.0
log_Emax = 12.0
nPoints = 2000
step=(log_Emax-log_Emin)/nPoints
logE=np.arange(log_Emin, log_Emax, step)
plt.yscale("log")
plt.xscale("log")
factor = 1.e-4/(4*np.pi)
plt.plot(np.power(10,logE)[1000:], factor*Phi_atm(np.power(10,logE)[1000:], a[0], a[1]), label='Fitted function')
plt.plot(flux_d["E"],factor*flux_d["nu_mu"], linewidth=3, alpha=0.5, label="data")
plt.plot(np.power(10,logE), Phi(np.power(10,logE)), label='astro')
Phi_atm(np.power(10,12), a[0], a[1])
|
notebooks/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Predicting Product Success When Review Data Is Available
# _**Using XGBoost to Predict Whether Sales will Exceed the "Hit" Threshold**_
#
# ---
#
# ---
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Setup](#Setup)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Host](#Host)
# 1. [Evaluation](#Evaluation)
# 1. [Extensions](#Extensions)
#
#
# ## Background
#
# Word of mouth in the form of user reviews, critic reviews, social media comments, etc. often can provide insights about whether a product ultimately will be a success. In the video game industry in particular, reviews and ratings can have a large impact on a game's success. However, not all games with bad reviews fail, and not all games with good reviews turn out to be hits. To predict hit games, machine learning algorithms potentially can take advantage of various relevant data attributes in addition to reviews.
#
# For this notebook, we will work with the data set [Video Game Sales with Ratings](https://www.kaggle.com/rush4ratio/video-game-sales-with-ratings) from Kaggle. This [Metacritic](http://www.metacritic.com/browse/games/release-date/available) data includes attributes for user reviews as well as critic reviews, sales, ESRB ratings, among others. Both user reviews and critic reviews are in the form of ratings scores, on a scale of 0 to 10 or 0 to 100. Although this is convenient, a significant issue with the data set is that it is relatively small.
#
# Dealing with a small data set such as this one is a common problem in machine learning. This problem often is compounded by imbalances between the classes in the small data set. In such situations, using an ensemble learner can be a good choice. This notebook will focus on using XGBoost, a popular ensemble learner, to build a classifier to determine whether a game will be a hit.
#
# ## Setup
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the `get_execution_role()` call with the appropriate full IAM role arn string(s).
# + isConfigCell=true
bucket = '<your_s3_bucket_name_here>'
prefix = 'sagemaker/videogames_xgboost'
# Define IAM role
import sagemaker
role = sagemaker.get_execution_role()
# -
# Next we'll import the Python libraries we'll need.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import Image
from IPython.display import display
from sklearn.datasets import dump_svmlight_file
from time import gmtime, strftime
import sys
import math
import json
import boto3
# ---
# ## Data
#
# Before proceeding further, you'll need to sign in to Kaggle or create a Kaggle account if you don't have one. Then **upload the raw CSV data set from the above Kaggle link to the S3 bucket you specified above**. The raw_data_filename specified below is the name of the data file from Kaggle, but you should alter it if the name changes. Let's download the data from your S3 bucket to your notebook instance, where it will appear in the same directory as this notebook. Then we'll take an initial look at the data.
# +
raw_data_filename = 'Video_Games_Sales_as_at_22_Dec_2016.csv'
s3 = boto3.resource('s3')
s3.Bucket(bucket).download_file(raw_data_filename, 'raw_data.csv')
data = pd.read_csv('./raw_data.csv')
pd.set_option('display.max_rows', 20)
data
# -
# Before proceeding further, we need to decide upon a target to predict. Video game development budgets can run into the tens of millions of dollars, so it is critical for game publishers to publish "hit" games to recoup their costs and make a profit. As a proxy for what constitutes a "hit" game, we will set a target of greater than 1 million units in global sales.
data['y'] = (data['Global_Sales'] > 1)
# With our target now defined, let's take a look at the imbalance between the "hit" and "not a hit" classes:
plt.bar(['not a hit', 'hit'], data['y'].value_counts())
plt.show()
# Not surprisingly, only a small fraction of games can be considered "hits" under our metric. Next, we'll choose features that have predictive power for our target. We'll begin by plotting review scores versus global sales to check our hunch that such scores have an impact on sales. Logarithmic scale is used for clarity.
viz = data.filter(['User_Score','Critic_Score', 'Global_Sales'], axis=1)
viz['User_Score'] = pd.Series(viz['User_Score'].apply(pd.to_numeric, errors='coerce'))
viz['User_Score'] = viz['User_Score'].mask(np.isnan(viz["User_Score"]), viz['Critic_Score'] / 10.0)
viz.plot(kind='scatter', logx=True, logy=True, x='Critic_Score', y='Global_Sales')
viz.plot(kind='scatter', logx=True, logy=True, x='User_Score', y='Global_Sales')
plt.show()
# Our intuition about the relationship between review scores and sales seems justified. We also note in passing that other relevant features can be extracted from the data set. For example, the ESRB rating has an impact since games with an "E" for everyone rating typically reach a wider audience than games with an age-restricted "M" for mature rating, though depending on another feature, the genre (such as shooter or action), M-rated games also can be huge hits. Our model hopefully will learn these relationships and others.
#
# Next, looking at the columns of features of this data set, we can identify several that should be excluded. For example, there are five columns that specify sales numbers: these numbers are directly related to the target we're trying to predict, so these columns should be dropped. Other features may be irrelevant, such as the name of the game.
data = data.drop(['Name', 'Year_of_Release', 'NA_Sales', 'EU_Sales', 'JP_Sales', 'Other_Sales', 'Global_Sales', 'Critic_Count', 'User_Count', 'Developer'], axis=1)
# With the number of columns reduced, now is a good time to check how many columns are missing data:
data.isnull().sum()
# As noted in Kaggle's overview of this data set, many review ratings are missing. Unfortunately, since those are crucial features that we are relying on for our predictions, and there is no reliable way of imputing so many of them, we'll need to drop rows missing those features.
data = data.dropna()
# Now we need to resolve a problem we see in the User_Score column: it contains some 'tbd' string values, so it obviously is not numeric. User_Score is more properly a numeric rather than categorical feature, so we'll need to convert it from string type to numeric, and temporarily fill in NaNs for the tbds. Next, we must decide what to do with these new NaNs in the User_Score column. We've already thrown out a large number of rows, so if we can salvage these rows, we should. As a first approximation, we'll take the value in the Critic_Score column and divide by 10 since the user scores tend to track the critic scores (though on a scale of 0 to 10 instead of 0 to 100).
data['User_Score'] = data['User_Score'].apply(pd.to_numeric, errors='coerce')
data['User_Score'] = data['User_Score'].mask(np.isnan(data["User_Score"]), data['Critic_Score'] / 10.0)
# Let's do some final preprocessing of the data, including converting the categorical features into numeric using the one-hot encoding method.
data['y'] = data['y'].apply(lambda y: 'yes' if y == True else 'no')
model_data = pd.get_dummies(data)
# To help prevent overfitting the model, we'll randomly split the data into three groups. Specifically, the model will be trained on 70% of the data. It will then be evaluated on 20% of the data to give us an estimate of the accuracy we hope to have on "new" data. As a final testing dataset, the remaining 10% will be held out until the end.
train_data, validation_data, test_data = np.split(model_data.sample(frac=1, random_state=1729), [int(0.7 * len(model_data)), int(0.9 * len(model_data))])
# XGBoost operates on data in the libSVM data format, with features and the target variable provided as separate arguments. To avoid any misalignment issues due to random reordering, this split is done after the previous split in the above cell. As a last step before training, we'll copy the resulting files to S3 as input for SageMaker's managed training.
# +
dump_svmlight_file(X=train_data.drop(['y_no', 'y_yes'], axis=1), y=train_data['y_yes'], f='train.libsvm')
dump_svmlight_file(X=validation_data.drop(['y_no', 'y_yes'], axis=1), y=validation_data['y_yes'], f='validation.libsvm')
dump_svmlight_file(X=test_data.drop(['y_no', 'y_yes'], axis=1), y=test_data['y_yes'], f='test.libsvm')
boto3.Session().resource('s3').Bucket(bucket).Object(prefix + '/train/train.libsvm').upload_file('train.libsvm')
boto3.Session().resource('s3').Bucket(bucket).Object(prefix + '/validation/validation.libsvm').upload_file('validation.libsvm')
# -
# ---
# ## Train
#
# Our data is now ready to be used to train a XGBoost model. The XGBoost algorithm has many tunable hyperparameters. Some of these hyperparameters are listed below; initially we'll only use a few of them.
#
# - `max_depth`: Maximum depth of a tree. As a cautionary note, a value too small could underfit the data, while increasing it will make the model more complex and thus more likely to overfit the data (in other words, the classic bias-variance tradeoff).
# - `eta`: Step size shrinkage used in updates to prevent overfitting.
# - `eval_metric`: Evaluation metric(s) for validation data. For data sets such as this one with imbalanced classes, we'll use the AUC metric.
# - `scale_pos_weight`: Controls the balance of positive and negative weights, again useful for data sets having imbalanced classes.
#
# First we'll setup the parameters for a training job, then create a training job with those parameters and run it.
# +
job_name = 'videogames-xgboost-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("Training job", job_name)
containers = {
'us-west-2': '433757028032.dkr.ecr.us-west-2.amazonaws.com/xgboost:latest',
'us-east-1': '811284229777.dkr.ecr.us-east-1.amazonaws.com/xgboost:latest',
'us-east-2': '825641698319.dkr.ecr.us-east-2.amazonaws.com/xgboost:latest',
'eu-west-1': '685385470294.dkr.ecr.eu-west-1.amazonaws.com/xgboost:latest'
}
create_training_params = \
{
"RoleArn": role,
"TrainingJobName": job_name,
"AlgorithmSpecification": {
"TrainingImage": containers[boto3.Session().region_name],
"TrainingInputMode": "File"
},
"ResourceConfig": {
"InstanceCount": 1,
"InstanceType": "ml.c4.xlarge",
"VolumeSizeInGB": 10
},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/train".format(bucket, prefix),
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "libsvm",
"CompressionType": "None"
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/validation".format(bucket, prefix),
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "libsvm",
"CompressionType": "None"
}
],
"OutputDataConfig": {
"S3OutputPath": "s3://{}/{}/xgboost-video-games/output".format(bucket, prefix)
},
"HyperParameters": {
"max_depth":"3",
"eta":"0.1",
"eval_metric":"auc",
"scale_pos_weight":"2.0",
"subsample":"0.5",
"objective":"binary:logistic",
"num_class":"1",
"num_round":"100"
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 60 * 60
}
}
# +
# %%time
sm = boto3.client('sagemaker')
sm.create_training_job(**create_training_params)
status = sm.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print(status)
try:
sm.get_waiter('training_job_completed_or_stopped').wait(TrainingJobName=job_name)
finally:
status = sm.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print("Training job ended with status: " + status)
if status == 'Failed':
message = sm.describe_training_job(TrainingJobName=job_name)['FailureReason']
print('Training failed with the following error: {}'.format(message))
raise Exception('Training job failed')
# -
# ---
# ## Host
#
# Now that we've trained the XGBoost algorithm on our data, let's prepare the model for hosting on a SageMaker serverless endpoint. We will:
#
# 1. Point to the scoring container
# 1. Point to the model.tar.gz that came from training
# 1. Create the hosting model
# +
create_model_response = sm.create_model(
ModelName=job_name,
ExecutionRoleArn=role,
PrimaryContainer={
'Image': containers[boto3.Session().region_name],
'ModelDataUrl': sm.describe_training_job(TrainingJobName=job_name)['ModelArtifacts']['S3ModelArtifacts']})
print(create_model_response['ModelArn'])
# -
# Next, we'll configure our hosting endpoint. Here we specify:
#
# 1. EC2 instance type to use for hosting
# 1. The initial number of instances
# 1. Our hosting model name
#
# After the endpoint has been configured, we'll create the endpoint itself.
# +
xgboost_endpoint_config = 'videogames-xgboost-endpoint-config-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(xgboost_endpoint_config)
create_endpoint_config_response = sm.create_endpoint_config(
EndpointConfigName=xgboost_endpoint_config,
ProductionVariants=[{
'InstanceType': 'ml.t2.medium',
'InitialInstanceCount': 1,
'ModelName': job_name,
'VariantName': 'AllTraffic'}])
print("Endpoint Config Arn: " + create_endpoint_config_response['EndpointConfigArn'])
# +
# %%time
xgboost_endpoint = 'EXAMPLE-videogames-xgb-endpoint-' + strftime("%Y%m%d%H%M", gmtime())
print(xgboost_endpoint)
create_endpoint_response = sm.create_endpoint(
EndpointName=xgboost_endpoint,
EndpointConfigName=xgboost_endpoint_config)
print(create_endpoint_response['EndpointArn'])
resp = sm.describe_endpoint(EndpointName=xgboost_endpoint)
status = resp['EndpointStatus']
print("Status: " + status)
try:
sm.get_waiter('endpoint_in_service').wait(EndpointName=xgboost_endpoint)
finally:
resp = sm.describe_endpoint(EndpointName=xgboost_endpoint)
status = resp['EndpointStatus']
print("Arn: " + resp['EndpointArn'])
print("Status: " + status)
if status != 'InService':
message = sm.describe_endpoint(EndpointName=xgboost_endpoint)['FailureReason']
print('Endpoint creation failed with the following error: {}'.format(message))
raise Exception('Endpoint creation did not succeed')
# -
# ---
#
# ## Evaluation
#
# Now that we have our hosted endpoint, we can generate predictions from it. More specifically, let's generate predictions from our test data set to understand how well our model generalizes to data it has not seen yet.
#
# There are many ways to compare the performance of a machine learning model. We'll start simply by comparing actual to predicted values of whether the game was a "hit" (`1`) or not (`0`). Then we'll produce a confusion matrix, which shows how many test data points were predicted by the model in each category versus how many test data points actually belonged in each category.
runtime = boto3.client('runtime.sagemaker')
# +
def do_predict(data, endpoint_name, content_type):
payload = '\n'.join(data)
response = runtime.invoke_endpoint(EndpointName=endpoint_name,
ContentType=content_type,
Body=payload)
result = response['Body'].read()
result = result.decode("utf-8")
result = result.split(',')
preds = [float((num)) for num in result]
preds = [round(num) for num in preds]
return preds
def batch_predict(data, batch_size, endpoint_name, content_type):
items = len(data)
arrs = []
for offset in range(0, items, batch_size):
if offset+batch_size < items:
results = do_predict(data[offset:(offset+batch_size)], endpoint_name, content_type)
arrs.extend(results)
else:
arrs.extend(do_predict(data[offset:items], endpoint_name, content_type))
sys.stdout.write('.')
return(arrs)
# +
# %%time
import json
with open('test.libsvm', 'r') as f:
payload = f.read().strip()
labels = [int(line.split(' ')[0]) for line in payload.split('\n')]
test_data = [line for line in payload.split('\n')]
preds = batch_predict(test_data, 100, xgboost_endpoint, 'text/x-libsvm')
print ('\nerror rate=%f' % ( sum(1 for i in range(len(preds)) if preds[i]!=labels[i]) /float(len(preds))))
# -
pd.crosstab(index=np.array(labels), columns=np.array(preds))
# Of the 132 games in the test set that actually are "hits" by our metric, the model correctly identified 73, while the overall error rate is 13%. The amount of false negatives versus true positives can be shifted substantially in favor of true positives by increasing the hyperparameter scale_pos_weight. Of course, this increase comes at the expense of reduced accuracy/increased error rate and more false positives. How to make this trade-off ultimately is a business decision based on the relative costs of false positives, false negatives, etc.
# ---
# ## Extensions
#
# This XGBoost model is just the starting point for predicting whether a game will be a hit based on reviews and other features. There are several possible avenues for improving the model's performance. First, of course, would be to collect more data and, if possible, fill in the existing missing fields with actual information. Another possibility is further hyperparameter tuning, with Amazon SageMaker's Hyperparameter Optimization service. And, although ensemble learners often do well with imbalanced data sets, it could be worth exploring techniques for mitigating imbalances such as downsampling, synthetic data augmentation, and other approaches.
sm.delete_endpoint(EndpointName=xgboost_endpoint)
|
introduction_to_applying_machine_learning/video_game_sales/video-game-sales-xgboost.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.nn as nn
import pyro
from pyro.distributions import Bernoulli
from pyro.distributions import Delta
from pyro.distributions import Normal
from pyro.distributions import Uniform
from pyro.distributions import LogNormal
from pyro.infer import SVI
from pyro.infer import Trace_ELBO
from pyro.optim import Adam
import torch.distributions.constraints as constraints
from pyro.infer.autoguide import AutoDiagonalNormal
# initialize the autodiagonal with init_to_feasible instead of init_to_median
from pyro.infer.autoguide import init_to_feasible
# Data Loader
import sys
# insert at 1, 0 is the script path (or '' in REPL)
sys.path.insert(1, '../box_office/')
from data_loader import load_tensor_data
pyro.set_rng_seed(101)
# -
x_train_tensors, y_train_tensors, actors, full_data = load_tensor_data("../data/ohe_movies.csv")
# +
def f_z(params):
"""Samples from P(Z)"""
z_mean0 = params['z_mean0']
z_std0 = params['z_std0']
z = pyro.sample("z", Normal(loc = z_mean0, scale = z_std0))
return z
def f_x(z, params):
"""
Samples from P(X|Z)
P(X|Z) is a Bernoulli with E(X|Z) = logistic(Z * W),
where W is a parameter (matrix). In training the W is
hyperparameters of the W distribution are estimated such
that in P(X|Z), the elements of the vector of X are
conditionally independent of one another given Z.
"""
def sample_W():
"""
Sample the W matrix
W is a parameter of P(X|Z) that is sampled from a Normal
with location and scale hyperparameters w_mean0 and w_std0
"""
w_mean0 = params['w_mean0']
w_std0 = params['w_std0']
W = pyro.sample("W", Normal(loc = w_mean0, scale = w_std0))
return W
W = sample_W()
linear_exp = torch.matmul(z, W)
# sample x using the Bernoulli likelihood
x = pyro.sample("x", Bernoulli(logits = linear_exp))
return x
def f_y(x, z, params):
"""
Samples from P(Y|X, Z)
Y is sampled from a Gaussian where the mean is an
affine combination of X and Z. Bayesian linear
regression is used to estimate the parameters of
this affine transformation function. Use torch.nn.Module to create
the Bayesian linear regression component of the overall
model.
"""
predictors = torch.cat((x, z), 1)
w = pyro.sample('weight', Normal(params['weight_mean0'], params['weight_std0']))
b = pyro.sample('bias', Normal(params['bias_mean0'], params['bias_std0']))
y_hat = (w * predictors).sum(dim=1) + b
# variance of distribution centered around y
sigma = pyro.sample('sigma', Normal(params['sigma_mean0'], params['sigma_std0']))
with pyro.iarange('data', len(predictors)):
pyro.sample('y', LogNormal(y_hat, sigma))
return y_hat
# -
def model(params):
"""The full generative causal model"""
z = f_z(params)
x = f_x(z, params)
y = f_y(x, z, params)
return {'z': z, 'x': x, 'y': y}
def step_1_guide(params):
"""
Guide function for fitting P(Z) and P(X|Z) from data
"""
# Infer z hyperparams
qz_mean = pyro.param("qz_mean", params['z_mean0'])
qz_stddv = pyro.param("qz_stddv", params['z_std0'],
constraint=constraints.positive)
z = pyro.sample("z", Normal(loc = qz_mean, scale = qz_stddv))
# Infer w params
qw_mean = pyro.param("qw_mean", params["w_mean0"])
qw_stddv = pyro.param("qw_stddv", params["w_std0"],
constraint=constraints.positive)
w = pyro.sample("w", Normal(loc = qw_mean, scale = qw_stddv))
def step_2_guide(params):
# Z and W are just sampled using param values optimized in previous step
z = pyro.sample("z", Normal(loc = params['qz_mean'], scale = params['qz_stddv']))
w = pyro.sample("w", Normal(loc = params['qw_mean'], scale = params['qw_stddv']))
# Infer regression params
# parameters of (w : weight)
w_loc = pyro.param('w_loc', params['weight_mean0'])
w_scale = pyro.param('w_scale', params['weight_std0'])
# parameters of (b : bias)
b_loc = pyro.param('b_loc', params['bias_mean0'])
b_scale = pyro.param('b_scale', params['bias_std0'])
# parameters of (sigma)
sigma_loc = pyro.param('sigma_loc', params['sigma_mean0'])
sigma_scale = pyro.param('sigma_scale', params['sigma_std0'])
# sample (w, b, sigma)
w = pyro.sample('weight', Normal(w_loc, w_scale))
b = pyro.sample('bias', Normal(b_loc, b_scale))
sigma = pyro.sample('sigma', Normal(sigma_loc, sigma_scale))
def training_step_1(x_data, params):
adam_params = {"lr": 0.0005}
optimizer = Adam(adam_params)
conditioned_on_x = pyro.condition(model, data = {"x" : x_data})
svi = SVI(conditioned_on_x, step_1_guide, optimizer, loss=Trace_ELBO())
print("\n Training Z marginal and W parameter marginal...")
n_steps = 2000
pyro.set_rng_seed(101)
# do gradient steps
pyro.get_param_store().clear()
for step in range(n_steps):
loss = svi.step(params)
if step % 100 == 0:
print("[iteration %04d] loss: %.4f" % (step + 1, loss/len(x_data)))
# grab the learned variational parameters
updated_params = {k: v for k, v in params.items()}
for name, value in pyro.get_param_store().items():
print("Updating value of hypermeter{}".format(name))
updated_params[name] = value.detach()
return updated_params
def training_step_2(x_data, y_data, params):
print("Training Bayesian regression parameters...")
pyro.set_rng_seed(101)
num_iterations = 1000
pyro.clear_param_store()
# Create a regression model
optim = Adam({"lr": 0.003})
conditioned_on_x_and_y = pyro.condition(model, data = {
"x": x_data,
"y" : y_data
})
svi = SVI(conditioned_on_x_and_y, step_2_guide, optim, loss=Trace_ELBO(), num_samples=1000)
for step in range(num_iterations):
loss = svi.step(params)
if step % 100 == 0:
print("[iteration %04d] loss: %.4f" % (step + 1, loss/len(x_data)))
updated_params = {k: v for k, v in params.items()}
for name, value in pyro.get_param_store().items():
print("Updating value of hypermeter: {}".format(name))
updated_params[name] = value.detach()
print("Training complete.")
return updated_params
def train_model():
num_datapoints, data_dim = x_train_tensors.shape
latent_dim = 30 # can be changed
# print(torch.zeros(data_dim + latent_dim).shape)
params0 = {
'z_mean0': torch.zeros([num_datapoints, latent_dim]),
'z_std0' : torch.ones([num_datapoints, latent_dim]),
'w_mean0' : torch.zeros([latent_dim, data_dim]),
'w_std0' : torch.ones([latent_dim, data_dim]),
'weight_mean0': torch.zeros(data_dim + latent_dim),
'weight_std0': torch.ones(data_dim + latent_dim),
'bias_mean0': torch.tensor(0.),
'bias_std0': torch.tensor(1.),
'sigma_mean0' : torch.tensor(1.),
'sigma_std0' : torch.tensor(0.05)
}
params1 = training_step_1(x_train_tensors, params0)
params2 = training_step_2(x_train_tensors, y_train_tensors, params1)
return params1, params2
# trained_params = train_model()
p1, p2 = train_model()
# +
# Save all params to disk
import pickle
with open('params.pickle', 'wb') as handle:
pickle.dump(p2, handle, protocol=pickle.HIGHEST_PROTOCOL)
|
box_office/notebooks/new_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''Python39_venv'': venv)'
# name: python3
# ---
# # Задание 3.1 - Сверточные нейронные сети (Convolutional Neural Networks)
#
# Это последнее задание на numpy, вы до него дожили! Остался последний марш-бросок, дальше только PyTorch.
#
# В этом задании вы реализуете свою собственную сверточную нейронную сеть.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
from dataset import load_svhn, random_split_train_val
from gradient_check import check_layer_gradient, check_layer_param_gradient, check_model_gradient
from layers import FullyConnectedLayer, ReLULayer, ConvolutionalLayer, MaxPoolingLayer, Flattener
from model import ConvNet
from trainer import Trainer, Dataset
from optim import SGD, MomentumSGD
from metrics import multiclass_accuracy
# # Загружаем данные
#
# На этот раз мы не будем их преобразовывать в один вектор, а оставим размерности (num_samples, 32, 32, 3).
# +
def prepare_for_neural_network(train_X, test_X):
train_X = train_X.astype(float) / 255.0
test_X = test_X.astype(float) / 255.0
# Subtract mean
mean_image = np.mean(train_X, axis = 0)
train_X -= mean_image
test_X -= mean_image
return train_X, test_X
train_X, train_y, test_X, test_y = load_svhn("data", max_train=10000, max_test=1000)
train_X, test_X = prepare_for_neural_network(train_X, test_X)
# Split train into train and val
train_X, train_y, val_X, val_y = random_split_train_val(train_X, train_y, num_val = 1000)
# -
# # Реализуем новые слои!
#
# Сначала основной новый слой - сверточный (Convolutional layer).
# Для начала мы реализуем его для только одного канала, а потом для нескольких.
#
# Сверточный слой выполняет операцию свертки (convolution) с весами для каждого канала, а потом складывает результаты.
# Возможно, поможет пересмотреть Лекцию 6 или внимательно прочитать
# http://cs231n.github.io/convolutional-networks/
#
# Один из подходов к реализации сверточного слоя основан на том, что для конкретного "пикселя" выхода применение сверточного слоя эквивалентно обычному полносвязному.
# Рассмотрим один такой "пиксель":
# 
#
# Он получает на вход
# регион входа I размера `(batch_size, filter_size, filter_size, input_channels)`,
# применяет к нему веса W `(filter_size, filter_size, input_channels, output_channels`
# и выдает `(batch_size, output_channels)`.
#
# Если:
# - вход преобразовать в I' `(batch_size, filter_size*filter_size*input_channels)`,
# - веса в W' `(filter_size*filter_size*input_channels, output_channels)`,
# то выход "пикселе" будет эквивалентен полносвязному слою со входом I' и весами W'.
#
# Осталось выполнить его в цикле для каждого пикселя :)
# +
# TODO: Implement ConvolutionaLayer that supports only 1 output and input channel
# Note: now you're working with images, so X is 4-dimensional tensor of
# (batch_size, height, width, channels)
X = np.array([
[
[[1.0], [2.0]],
[[0.0], [-1.0]]
]
,
[
[[0.0], [1.0]],
[[-2.0], [-1.0]]
]
])
# Batch of 2 images of dimensions 2x2 with a single channel
print("Shape of X:", X.shape)
layer = ConvolutionalLayer(in_channels=1, out_channels=1, filter_size=2, padding=0)
print("Shape of W", layer.W.value.shape)
layer.W.value = np.zeros_like(layer.W.value)
layer.W.value[0, 0, 0, 0] = 1.0
layer.B.value = np.ones_like(layer.B.value)
result = layer.forward(X)
assert result.shape == (2, 1, 1, 1)
assert np.all(result == X[:, :1, :1, :1] +1), "result: %s, X: %s" % (result, X[:, :1, :1, :1])
# Now let's implement multiple output channels
layer = ConvolutionalLayer(in_channels=1, out_channels=2, filter_size=2, padding=0)
result = layer.forward(X)
assert result.shape == (2, 1, 1, 2)
# And now multple input channels!
X = np.array([
[
[[5.0, 0.0], [4.0, 1.0]],
[[0.0, -1.0], [-1.0, -2.0]]
]
,
[
[[0.0, 1.0], [1.0, -1.0]],
[[3.0, 2.0], [-1.0, 0.0]]
]
])
print("Shape of X:", X.shape)
layer = ConvolutionalLayer(in_channels=2, out_channels=2, filter_size=2, padding=0)
result = layer.forward(X)
assert result.shape == (2, 1, 1, 2)
# -
# ## А теперь имплементируем обратный проход
# Возможно, это самое сложное место в курсе. Дальше будет лучше.
#
# Раз выполнение сверточного слоя эквивалентно полносвязному слою для каждого "пикселя" выхода, то общий обратный проход эквивалентен обратному проходу каждого из таких "слоев".
# Градиенты от каждого из этих "слоев" в каждом пикселе надо сложить в соответствующие пиксели градиента по входу, а градиенты весов сложить все вместе.
# +
# First test - check the shape is right
layer = ConvolutionalLayer(in_channels=2, out_channels=2, filter_size=2, padding=0)
result = layer.forward(X)
# print(result)
d_input = layer.backward(np.ones_like(result))
assert d_input.shape == X.shape
# Actually test the backward pass
# As usual, you'll need to copy gradient check code from the previous assignment
layer = ConvolutionalLayer(in_channels=2, out_channels=2, filter_size=2, padding=0)
assert check_layer_gradient(layer, X)
layer = ConvolutionalLayer(in_channels=2, out_channels=2, filter_size=2, padding=0)
assert check_layer_param_gradient(layer, X, 'W')
layer = ConvolutionalLayer(in_channels=2, out_channels=2, filter_size=2, padding=0)
assert check_layer_param_gradient(layer, X, 'B')
# -
# Осталось реализовать дополнение нулями (padding).
# Достаточно дополнить входной тензор нулями по сторонам. Не забудьте учесть это при обратном проходе!
layer = ConvolutionalLayer(in_channels=2, out_channels=2, filter_size=3, padding=1)
result = layer.forward(X)
# Note this kind of layer produces the same dimensions as input
assert result.shape == X.shape,"Result shape: %s - Expected shape %s" % (result.shape, X.shape)
d_input = layer.backward(np.ones_like(result))
assert d_input.shape == X.shape
layer = ConvolutionalLayer(in_channels=2, out_channels=2, filter_size=3, padding=1)
assert check_layer_gradient(layer, X)
# ## После следующего слоя вам уже будет все ни по чем - max pooling
#
# Max Pooling - это слой, реализующий операцию максимума для каждого канала отдельно в окресности из `pool_size` "пикселей".
#
# 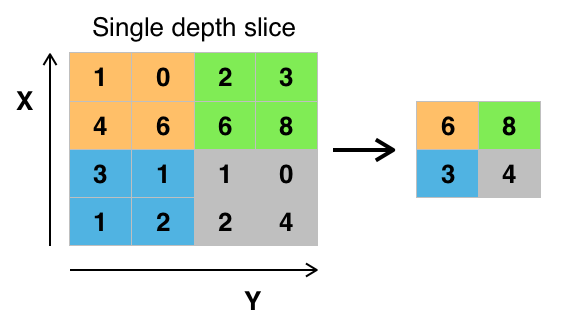
#
# И напомним что такое stride.
# Stride - это на сколько "пикселей" сдвигается окно на одном шаге.
# Вот здесь, например, stride = 2
#
# 
#
# На практике, для max pooling значение stride часто равно pool size.
# +
pool = MaxPoolingLayer(2, 2)
result = pool.forward(X)
assert result.shape == (2, 1, 1, 2)
assert check_layer_gradient(pool, X)
# -
# И на закуску - слой, преобразующий четырехмерные тензоры в двумерные.
#
# Этот слой понадобится нам, чтобы в конце сети перейти от сверточных слоев к полносвязным.
# +
flattener = Flattener()
result = flattener.forward(X)
assert result.shape == (2,8)
assert check_layer_gradient(flattener, X)
# -
# # Теперь есть все кирпичики, создаем модель
# +
# TODO: In model.py, implement missed functions function for ConvNet model
# No need to use L2 regularization
model = ConvNet(input_shape=(32,32,3), n_output_classes=10, conv1_channels=2, conv2_channels=2)
loss = model.compute_loss_and_gradients(train_X[:2], train_y[:2])
# TODO Now implement backward pass and aggregate all of the params
check_model_gradient(model, train_X[:2], train_y[:2])
# -
# # Оптимизатор и код для тренировки
# Должен заработать с кодом из прошлого задания без изменений!
# +
model = ConvNet(input_shape=(32,32,3), n_output_classes=10, conv1_channels=2, conv2_channels=2)
dataset = Dataset(train_X[:16], train_y[:16], val_X[:16], val_y[:16])
trainer = Trainer(model, dataset, SGD(), batch_size=16, learning_rate=1e-4)
loss_history, train_history, val_history = trainer.fit()
# -
plt.plot(train_history)
plt.plot(val_history)
# # Последнее упражнение
# В качестве последнего упражнения мы доведем точность на тренировочном наборе данных до 100% на небольшом наборе данных.
# Сверточные сети требуют большого количества вычислений и аккуратной эффективной реализации, поэтому настоящие модели мы будем тренировать уже на PyTorch в следующем задании.
# ## Итак, оверфитим маленький набор данных
#
# +
data_size = 128
model = ConvNet(input_shape=(32,32,3), n_output_classes=10, conv1_channels=2, conv2_channels=2)
dataset = Dataset(train_X[:data_size], train_y[:data_size], val_X[:data_size], val_y[:data_size])
# TODO: Change any hyperparamers or optimizators to reach 1.0 training accuracy in 50 epochs or less
# Hint: If you have hard time finding the right parameters manually, try grid search or random search!
trainer = Trainer(model, dataset, SGD(), learning_rate=1e-3, num_epochs=50, batch_size=32)
loss_history, train_history, val_history = trainer.fit()
# -
plt.plot(train_history)
plt.plot(val_history)
# Дальнейшие упражнения - уже на PyTorch, открывайте следующий notebook!
#
# 
|
assignments/assignment3/CNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (base)
# language: python
# name: base
# ---
from Bio import SeqIO
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import tqdm
import glob
import re
import requests
import io
#
pfamA_motors = pd.read_csv("../../data/pfamA_motors.csv")
pfamA_motors.head()
kinesin = pfamA_motors.loc[pfamA_motors["pfamA_acc"] == "PF00225",:]
kinesin.shape
tubulin = pfamA_motors.loc[pfamA_motors["pfamA_acc"] == "PF00091",:]
tubulin.shape
kinesin["uniprot_acc"] = kinesin["id"].apply(lambda s: s.split("/")[0])
tubulin["uniprot_acc"] = tubulin["id"].apply(lambda s: s.split("/")[0])
kinesin.head()
def get_uniprot_entry(acc):
url = "https://www.uniprot.org/uniprot/?query="+acc+"&sort=score&columns=id,entry name,reviewed,protein names,genes,organism,organism-id,lineage(all),sequence,length&format=tab"
try:
urlData = requests.get(url).content
rawData = pd.read_csv(io.StringIO(urlData.decode('utf-8')),sep = '\t')
except:
rawData = pd.DataFrame()
return rawData
a = get_uniprot_entry("A0A428QG75_9HYPO")
b = get_uniprot_entry("A0A2R6QVF3_ACTCH")
kinesin_uniprot = get_uniprot_entry(kinesin["uniprot_acc"].iloc[0])
for i in range(1,kinesin.shape[0]):
curr = get_uniprot_entry(kinesin["uniprot_acc"].iloc[i])
if not (curr.empty):
kinesin_uniprot = pd.concat([kinesin_uniprot,curr], ignore_index=True)
if i%500 ==0:
print(kinesin_uniprot.shape)
tubulin_uniprot = get_uniprot_entry(tubulin["uniprot_acc"].iloc[0])
for i in range(1,tubulin.shape[0]):
curr = get_uniprot_entry(kinesin["uniprot_acc"].iloc[i])
if not (curr.empty):
tubulin_uniprot = pd.concat([tubulin_uniprot,curr], ignore_index=True)
if i%500 ==0:
print(tubulin_uniprot.shape)
print('done')
tubulin_uniprot.shape
kinesin_uniprot.shape
tubulin_uniprot.to_csv("../../data/david_data/tubulin.csv",index = False)
kinesin_uniprot.to_csv("../../data/david_data/kinesin.csv",index = False)
print('done')
# ## Inspect whether the organisms of top interests are included in the dataset
#
# - Giardia lamblia: Giardia intestinalis
# - Paramecium: Paramecium
# - Diatom: Bacillariophyta
# - Dileptus sp.: Dileptus
# - Enchelyodon sp.: Enchelyodon
# - Opisthokonta: Opisthokonta
# - Amoebozoa: Amoebozoa
# - Hemimastigophora: Hemimastigophora
# - Euglenozoa: Euglenozoa
# - Archaeplastida: Rhodophyta,Rhodelphida,Glaucocystophyceae,Viridiplantae,Picozoa,Epimorpha
# - Carpediemonas
tubulin_uniprot = pd.read_csv("../../data/david_data/tubulin.csv")
tubulin_uniprot["Taxonomic lineage (all)"].head()
organism_ls = ["Giardia intestinalis","Paramecium","Bacillariophyta","Dileptus","Enchelyodon","Opisthokonta",\
"Amoebozoa","Hemimastigophora","Euglenozoa","Rhodophyta","Rhodelphida","Glaucocystophyceae","Viridiplantae",\
"Picozoa","Epimorpha"]
# +
lowered_description = tubulin_uniprot["Taxonomic lineage (all)"].str.lower()
tubulin_uniprot["og"] = "unlabeled"
for label in organism_ls:
print(label)
label = label.lower()
boo = lowered_description.str.contains( label,na=False)
tubulin_uniprot.loc[boo,"og"] = label
# -
tubulin_uniprot.groupby("og").count().iloc[:,0:1]
kinesin_uniprot = pd.read_csv("../../data/david_data/kinesin.csv")
# +
lowered_description = kinesin_uniprot["Taxonomic lineage (all)"].str.lower()
kinesin_uniprot["og"] = "unlabeled"
for label in organism_ls:
print(label)
label = label.lower()
boo = lowered_description.str.contains( label,na=False)
kinesin_uniprot.loc[boo,"og"] = label
# -
kinesin_uniprot.groupby("og").count().iloc[:,0:1]
kinesin_uniprot.head()
tubulin_uniprot.to_csv("../../data/david_data/tubulin.csv",index = False)
kinesin_uniprot.to_csv("../../data/david_data/kinesin.csv",index = False)
|
code/data_process/unicellular_motor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="JUgOrRxl0sKq"
# ## Google Colab
#
# Google provides a free cloud service based on Jupyter Notebooks that supports free CPU and GPU. It allows you to develop deep learning applications using popular libraries such as PyTorch, TensorFlow, Keras, and OpenCV (without installation). All these libraries are pre-installed on Google Colab along wilt Python.
# -
# ### 1. Notebook Creation
#
# Login with your account and got to [google colab](https://colab.research.google.com). You will be prompted to either create a new notebook or you can also upload your `.ipynb` notebook from your Github, Google Drive or your local machine.
#
# <img src="./images/colab_upload.png">
# Once you have created the notebook, you can rename it by clicking on notebook name in the upper right corner.
# <img src="./images/colab_rename.png">
#
#
# All your notebooks will be saved in your Google Drive inside the directory `Colab Notebooks`.
# ### 2. Dataset
#
# In upcoming assignments you would need data to train your model. The best way to use colab with your dataset is to upload your dataset to google drive and the mount your drive. You can do so with the following command
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="nyfGOBXU7cjL" outputId="f2c83b83-9633-403c-fe4e-5750acd9c33d"
from google.colab import drive
drive.mount("/content/gdrive", force_remount=True)
# + [markdown] colab_type="text" id="hAvnynvP0sKv"
# Now you should see your drive on the left-hand side of the screen!.(You may need to hit "refresh" if it doesn't occur immediately)
#
# <img src="./images/colab_mount.png">
# + [markdown] colab_type="text" id="YddOu3Bc0sKx"
# ### 3. Installing python libraries
#
# In general you would not need to install anything, but incase you have then you can do so with the following command.
# -
# !pip3 install torch torchvision
# <img src="./images/colab_install.png">
# ### 4. Download Notebooks
#
# Your notebooks are automatically saved in your google drive. But if you need to download them, you can do so by `File` -> `Download .ipynb`
|
exercise_0/3_colab_introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:caselaw]
# language: python
# name: conda-env-caselaw-py
# ---
# %matplotlib inline
import pandas as pd
import os
import matplotlib.pyplot as plt
from IPython.display import display
filepath = '/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/'
art_nodes = pd.read_csv(os.path.join(filepath, 'article_nodes_nodup_min5_bimodal_titles.csv'))
case_nodes = pd.read_csv(os.path.join(filepath, 'case_nodes_simple_bimodal_titles.csv'))
bimodal_links = pd.read_csv(os.path.join(filepath, 'case_to_article_title_links.csv'))
bimodal_clusters = pd.read_csv(os.path.join(filepath, 'bimodal_clusters_titles.csv'))
print(art_nodes.shape, case_nodes.shape)
bimodal_clusters.type.value_counts()
print("Number of communities: {}".format(bimodal_clusters['community'].nunique()))
print("Number of communities in largest cc: {}".format(bimodal_clusters[bimodal_clusters['cc']==0]['community'].nunique()))
comm_counts = bimodal_clusters.groupby(['community', 'type']).count()['name'].unstack()
# +
fig, axes = plt.subplots(3, 1, figsize=(8, 15))
n = 50
comm_counts[:n].plot(kind='bar', ax=axes[0], title='Cases and articles')
axes[0].set_ylabel('count')
comm_counts['article'][:n].plot(kind='bar', ax=axes[1], title='Articles')
axes[1].set_ylabel('count')
comm_counts['case'][:n].plot(kind='bar', ax=axes[2], title='Cases')
axes[2].set_ylabel('count')
# -
largest_coms = bimodal_clusters.community.value_counts().index
df_plot = case_nodes[case_nodes.community_bimodal.isin(largest_coms[:10])]
df_plot.groupby(['year','community_bimodal']).count()['ecli'].unstack().plot(colormap='nipy_spectral')
plt.gca().set_xlim(1990, 2017)
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
# ## compare with courts
courts = pd.read_csv(os.path.join(filepath, 'court_nodes_corrected.csv'))
case_nodes_courts = pd.merge(case_nodes, courts, how='left', left_on='court', right_on='id')
case_nodes_courts.Type.value_counts()
case_nodes_courts[case_nodes_courts.Type.isnull()].court.value_counts()
case_nodes_courts.loc[case_nodes_courts.Type.isnull(), 'Type'] = 'AndereGerechtelijkeInstantie'
case_nodes_courts.Type.value_counts().plot.bar()
df_plot = case_nodes_courts[case_nodes_courts.community_bimodal.isin(largest_coms[:20])]
ax = df_plot.groupby(['community_bimodal', 'Type']).count()['ecli'].unstack().plot.bar(stacked=True, colormap='nipy_spectral', figsize=(13,5))
ax.set_ylabel('count')
ax.set_xlabel('community')
plt.legend(bbox_to_anchor=(0, 1.1),loc=8, borderaxespad=0.)
plt.show()
for com in [0, 1, 3, 4]:
nodes_sub = art_nodes[art_nodes.community==com]
print('Community {}:'.format(com))
print(nodes_sub['id'].values)
for com in largest_coms[:20]:
nodes_sub = art_nodes[art_nodes.community==com]
nodes_sub.groupby(['book']).count()['id'].plot.barh(figsize=(3,3))
plt.title('Community {}, {} articles'.format(com, len(nodes_sub)))
plt.show()
#display(nodes_sub)
# ## summary statistics
import nwtools.communities
import networkx as nx
graph = nx.from_pandas_edgelist(bimodal_links, source='source', target='target')
partition = bimodal_clusters.set_index('name')['community'].to_dict()
# +
com_statistics = nwtools.communities.partition_statistics(partition, graph)
com_statistics_df = pd.DataFrame.from_dict(com_statistics, orient='index')
# -
com_statistics_df.head()
|
notebooks/Community_report_bimodal.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [Intermediate Machine Learning](https://www.kaggle.com/learn/intermediate-machine-learning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/xgboost).**
#
# ---
#
# In this exercise, you will use your new knowledge to train a model with **gradient boosting**.
#
# # Setup
#
# The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
# Set up code checking
import os
if not os.path.exists("../input/train.csv"):
os.symlink("../input/home-data-for-ml-course/train.csv", "../input/train.csv")
os.symlink("../input/home-data-for-ml-course/test.csv", "../input/test.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.ml_intermediate.ex6 import *
print("Setup Complete")
# You will work with the [Housing Prices Competition for Kaggle Learn Users](https://www.kaggle.com/c/home-data-for-ml-course) dataset from the previous exercise.
#
# 
#
# Run the next code cell without changes to load the training and validation sets in `X_train`, `X_valid`, `y_train`, and `y_valid`. The test set is loaded in `X_test`.
# +
import pandas as pd
from sklearn.model_selection import train_test_split
# Read the data
X = pd.read_csv('../input/train.csv', index_col='Id')
X_test_full = pd.read_csv('../input/test.csv', index_col='Id')
# Remove rows with missing target, separate target from predictors
X.dropna(axis=0, subset=['SalePrice'], inplace=True)
y = X.SalePrice
X.drop(['SalePrice'], axis=1, inplace=True)
# Break off validation set from training data
X_train_full, X_valid_full, y_train, y_valid = train_test_split(X, y, train_size=0.8, test_size=0.2,
random_state=0)
# "Cardinality" means the number of unique values in a column
# Select categorical columns with relatively low cardinality (convenient but arbitrary)
low_cardinality_cols = [cname for cname in X_train_full.columns if X_train_full[cname].nunique() < 10 and
X_train_full[cname].dtype == "object"]
# Select numeric columns
numeric_cols = [cname for cname in X_train_full.columns if X_train_full[cname].dtype in ['int64', 'float64']]
# Keep selected columns only
my_cols = low_cardinality_cols + numeric_cols
X_train = X_train_full[my_cols].copy()
X_valid = X_valid_full[my_cols].copy()
X_test = X_test_full[my_cols].copy()
# One-hot encode the data (to shorten the code, we use pandas)
X_train = pd.get_dummies(X_train)
X_valid = pd.get_dummies(X_valid)
X_test = pd.get_dummies(X_test)
X_train, X_valid = X_train.align(X_valid, join='left', axis=1)
X_train, X_test = X_train.align(X_test, join='left', axis=1)
# -
# # Step 1: Build model
#
# ### Part A
#
# In this step, you'll build and train your first model with gradient boosting.
#
# - Begin by setting `my_model_1` to an XGBoost model. Use the [XGBRegressor](https://xgboost.readthedocs.io/en/latest/python/python_api.html#xgboost.XGBRegressor) class, and set the random seed to 0 (`random_state=0`). **Leave all other parameters as default.**
# - Then, fit the model to the training data in `X_train` and `y_train`.
# +
from xgboost import XGBRegressor
# Define the model
my_model_1 = XGBRegressor(random_state=0)
# Fit the model
my_model_1.fit(X_train, y_train)
# Check your answer
step_1.a.check()
# +
# Lines below will give you a hint or solution code
#step_1.a.hint()
#step_1.a.solution()
# -
# ### Part B
#
# Set `predictions_1` to the model's predictions for the validation data. Recall that the validation features are stored in `X_valid`.
# +
from sklearn.metrics import mean_absolute_error
# Get predictions
predictions_1 = my_model_1.predict(X_valid) # Your code here
# Check your answer
step_1.b.check()
# +
# Lines below will give you a hint or solution code
#step_1.b.hint()
#step_1.b.solution()
# -
# ### Part C
#
# Finally, use the `mean_absolute_error()` function to calculate the mean absolute error (MAE) corresponding to the predictions for the validation set. Recall that the labels for the validation data are stored in `y_valid`.
# +
# Calculate MAE
mae_1 = mean_absolute_error(predictions_1, y_valid)
# Your code here
# Uncomment to print MAE
print("Mean Absolute Error:" , mae_1)
# Check your answer
step_1.c.check()
# +
# Lines below will give you a hint or solution code
#step_1.c.hint()
#step_1.c.solution()
# -
# # Step 2: Improve the model
#
# Now that you've trained a default model as baseline, it's time to tinker with the parameters, to see if you can get better performance!
# - Begin by setting `my_model_2` to an XGBoost model, using the [XGBRegressor](https://xgboost.readthedocs.io/en/latest/python/python_api.html#xgboost.XGBRegressor) class. Use what you learned in the previous tutorial to figure out how to change the default parameters (like `n_estimators` and `learning_rate`) to get better results.
# - Then, fit the model to the training data in `X_train` and `y_train`.
# - Set `predictions_2` to the model's predictions for the validation data. Recall that the validation features are stored in `X_valid`.
# - Finally, use the `mean_absolute_error()` function to calculate the mean absolute error (MAE) corresponding to the predictions on the validation set. Recall that the labels for the validation data are stored in `y_valid`.
#
# In order for this step to be marked correct, your model in `my_model_2` must attain lower MAE than the model in `my_model_1`.
# +
# Define the model
my_model_2 = XGBRegressor(n_estimators=1000, learning_rate=0.05)
# Fit the model
my_model_2.fit(X_train, y_train)
# Get predictions
predictions_2 = my_model_2.predict(X_valid)
# Calculate MAE
mae_2 = mean_absolute_error(predictions_2, y_valid)
# Uncomment to print MAE
print("Mean Absolute Error:" , mae_2)
# Check your answer
step_2.check()
# +
# Lines below will give you a hint or solution code
#step_2.hint()
#step_2.solution()
# -
# # Step 3: Break the model
#
# In this step, you will create a model that performs worse than the original model in Step 1. This will help you to develop your intuition for how to set parameters. You might even find that you accidentally get better performance, which is ultimately a nice problem to have and a valuable learning experience!
# - Begin by setting `my_model_3` to an XGBoost model, using the [XGBRegressor](https://xgboost.readthedocs.io/en/latest/python/python_api.html#xgboost.XGBRegressor) class. Use what you learned in the previous tutorial to figure out how to change the default parameters (like `n_estimators` and `learning_rate`) to design a model to get high MAE.
# - Then, fit the model to the training data in `X_train` and `y_train`.
# - Set `predictions_3` to the model's predictions for the validation data. Recall that the validation features are stored in `X_valid`.
# - Finally, use the `mean_absolute_error()` function to calculate the mean absolute error (MAE) corresponding to the predictions on the validation set. Recall that the labels for the validation data are stored in `y_valid`.
#
# In order for this step to be marked correct, your model in `my_model_3` must attain higher MAE than the model in `my_model_1`.
# +
# Define the model
my_model_3 = XGBRegressor(n_estimators=1)
# Fit the model
my_model_3.fit(X_train, y_train)
# Get predictions
predictions_3 = my_model_3.predict(X_valid)
# Calculate MAE
mae_3 = mean_absolute_error(predictions_3, y_valid)
# Uncomment to print MAE
print("Mean Absolute Error:" , mae_3)
# Check your answer
step_3.check()
# +
# Lines below will give you a hint or solution code
#step_3.hint()
#step_3.solution()
# -
# # Keep going
#
# Continue to learn about **[data leakage](https://www.kaggle.com/alexisbcook/data-leakage)**. This is an important issue for a data scientist to understand, and it has the potential to ruin your models in subtle and dangerous ways!
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161289) to chat with other Learners.*
|
Intermediate-Machine-Learning/exercise6-xgboost.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SSP-AHP based sustainability assessment
# This manual provides a simple explanation of the software provided in this repository based on an example of sustainability assessment from healthcare systems of 16 European countries considering their performance values of 25 criteria.
# Import the necessary libraries and modules
# +
import numpy as np
import pandas as pd
import copy
import os
from itertools import product
from PyPDF2 import PdfFileMerger
from normalizations import *
from visualizations import *
from weighting_methods import *
from ssp_ahp import AHP
from rank_preferences import *
# -
# Loading decision matrix containing performance values and criterion types (-1 denotes cost and 1 represents profit criterion) from CSV file
folder_name = './input'
# Name of file with input data
file_name = 'dataset.csv'
path_data = os.path.join(folder_name, file_name)
data = pd.read_csv(path_data, index_col = 'Country')
data
# Loading a hierarchical model of the investigated problem including main criteria, sub-criteria, and detailed criteria and matrix containing pairwise comparison criteria based on Saaty scale provided by the domain expert
# +
# Hierarchical model criteria tree
modules = [
[
['C1', 'C2'],
['C3', 'C4'],
['C5', 'C6', 'C7']
],
[
['C8', 'C9', 'C10', 'C11'],
['C12'],
['C13', 'C14']
],
[
['C15'],
['C16', 'C17']
],
[
['C18'],
['C19', 'C20']
],
[
['C21', 'C22'],
['C23', 'C24'],
['C25']
]
]
modules_indexes = [
[0, 1, 2, 3, 4, 5, 6],
[7, 8, 9, 10, 11, 12, 13],
[14, 15, 16],
[17, 18, 19],
[20, 21, 22, 23, 24]
]
# matrix with pairwise comparison criteria based on Saaty scale - provided by the domain expert
PCcriteria = np.array(([[1, 1, 5, 3, 9],
[1, 1, 3, 5, 7],
[1/5, 1/3, 1, 1, 9],
[1/3, 1/5, 1, 1, 7],
[1/9, 1/7, 1/9, 1/7, 1]
]))
# -
# Convert the criteria types and decision matrix from performance values to a NumPy array to simplify further calculations
# +
df_data = data.iloc[:len(data) - 1, :]
# criteria types
df_types = data.iloc[len(data) - 1, :]
types = df_types.to_numpy()
list_alt_names = [r'$A_{' + str(i + 1) + '}$' for i in range(0, len(df_data))]
# decision matrix with performance values
matrix = df_data.to_numpy()
# -
# The next step involves calculating the criteria weights using the AHP-based relative weighting method, using the eigenvector method weights. For this purpose, first, the consistency of the matrix with pairwise comparison criteria is checked. Matrices with result $\leq 0.1$ are acceptable
# calculation of main weights with AHP weighting method
ahp = AHP()
ahp._check_consistency(PCcriteria)
# The weights are then calculated using the eigenvector method. This procedure is easy to perform using the function provided by NumPy $\texttt{np.linalg.eig}$, which gives eigenvalues and eigenvectors as outputs. The first eigenvector is taken into account.
# eigenvector method
W = ahp._calculate_eigenvector(PCcriteria)
G = [r'$G_{' + str(i + 1) + '}$' for i in range(PCcriteria.shape[1])]
main_weights = copy.deepcopy(W)
# The resulting vector of weights for 5 main criteria:
main_weights
# Choose weighting method from three proposed: ahp critic entropy. ahp denotes subjective AHP-based relative weighting method and requires the matrix with pairwise comparison criteria constructed by the decision-maker. Other objective methods, namely Entropy weighting and CRITIC weighting, determine weights based on data in the decision matrix containing performance values
# +
# choose weighting method from three proposed: ahp critic entropy
weights_type = 'ahp'
if weights_type == 'ahp':
main_weights = copy.deepcopy(W)
weights, crit_list = structured_equal_weights(modules, main_weights)
# objective weights
elif weights_type == 'critic':
weights = critic_weighting(matrix)
crit_list = [r'$C_{' + str(i + 1) + '}$' for i in range(0, df_data.shape[1])]
elif weights_type == 'entropy':
weights = entropy_weighting(matrix)
crit_list = [r'$C_{' + str(i + 1) + '}$' for i in range(0, df_data.shape[1])]
# -
# The resulting vector of weights for all 25 criteria:
weights
# First, simple use of SSP-AHP without reducing the criteria compensation will be demonstrated. For this purpose, the parameter $mad$ denoting the mean average deviation of normalized values in the decision matrix is set to False, and there is no need to specify a vector with sustainability coefficients $s$ for the criteria. The SSP-AHP method includes normalization, multiplying the normalized matrix by weights and aggregation by summing the rows. The final utility values of alternatives are denoted by $pref$. They are ranked in descending order to generate a ranking of alternatives $rank$. Alternative with the highest utility value is the best variant.
df_ahp = pd.DataFrame(index = list_alt_names)
pref = ahp(matrix, weights, types, mad = False)
rank = rank_preferences(pref, reverse = True)
df_ahp['Utility value'] = pref
df_ahp['Rank'] = rank
df_ahp
# As can be easily observed in the table displayed above, the alternative $A_{11}$, which received the highest utility value, was the best evaluated.
# In the next step, the use of SSP-AHP with reduced compensation for all criteria will be demonstrated. For this purpose, the coefficient $s$ will be increased stepwise by 5% (0.05) from 0% to 100% (1.0). This time the $mad$ parameter is set to True, which means that the mean absolute deviation will be subtracted from the normalized values of the decision matrix. A vector with $s$ coefficients for the criteria is also given. The procedure is executed in a loop - for each value of the $s$ coefficient, the SSP-AHP procedure (ahp function) is run.
# SSP-AHP Simulation 1
# changes in the coefficient s in all criteria dimensions simultaneously
# The value 0 of s coefficient corresponds to the classical AHP method
df_sust = pd.DataFrame(index = list_alt_names)
df_sust_pref = copy.deepcopy(df_sust)
sust_coeffs = np.arange(0, 1.05, 0.05)
for s in sust_coeffs:
s_set = np.ones(matrix.shape[1]) * s
pref = ahp(matrix, weights, types, mad = True, s = s_set)
df_sust_pref["{:.2f}".format(s)] = pref
rank = rank_preferences(pref, reverse = True)
df_sust["{:.2f}".format(s)] = rank
df_sust_pref.to_csv('output/sust_utility_vals_' + weights_type + '.csv')
df_sust.to_csv('output/sust_rank_' + weights_type + '.csv')
pdf = plot_sustainability(sust_coeffs, df_sust, weights_type)
# The resulting chart displays the alternatives' rankings for each value of the $s$ coefficient (0-100%), which results in a progressive reduction of the criteria compensation. For example, it can be noticed that alternative $A_{11}$ with $s$ equal to 80% drops from the leading position to second place. On the other hand, alternative $A_{10}$ with increasing reduction of criteria compensation rises from the fourth place to first place, which indicates its high sustainability. The rankings for each $s$ value are displayed in the table below.
df_sust
# Analogous calculations can also be performed using the reduction of compensation of criteria only for their selected groups or for single criteria, which is demonstrated in the code in the file main.py (Simulation 2 and 3).
|
SSP-AHP Manual.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.8 64-bit
# name: python3
# ---
# # Regularization
#
# ## Ridge
#
# Add notes from missed class.
#
# The $\lambda I_n$ adds
#
# ## Lasso
#
#
# ## Shrinkage effect
#
# If each feature is independent, the design matrix (centerd and standardized) will have $x_j^T x_k = 1(j=k)$ because they are orthogonal.
#
# The least squares estimator will be reevalauted as
#
# $\hat{\beta}^{OLS} = (X^T X)^{-1} X^T y = X^T y$
#
# because $X^T X = I$. In other words, each predictor can be treated as a basis.
#
# In ridge regression, the $X^T X + \lambda I_p$ becomes a diagonal matrix of lambdas.
#
# $\hat{\beta}^R_\lambda = (X^T X + \lambda I_p)^{-1} X^T y = \frac{1}{1 + \lambda} X^T y = \frac{\beta^{OLS}}{1 + \lambda}$
#
# Therefore, the larger $\lambda$ is, the smaller the magnitude of each $\beta_j$. This is a linear function. Here, it is applied uniformally. However, in practice, features are not orthogonal. So, each coefficient is affected nonuniformally. It should shrink more on the smaller eigenvalues.
#
# This will shirnk the prediction by the same scale.
#
# $\hat{y}$ is scaled by ($1 + \lambda$)
#
# We know that the $\hat{\beta}^{OLS}$ is unbiased. $E \hat{\beta} = (X^T X)^{-1} X^T E(Y) = \beta$ because $E(Y) = X\beta$.
#
# However, the ridge regression is a biased estimator: $E \hat{\beta}_\lambda^R = \frac{\beta}{1 + \lambda} \neq \beta$
#
# Tradeoff between bias and variance. With collinearity, variance explodes (because we cannot take inverse bceause it is singular). Ridge regression will regularize the variance.
#
# For OLS, the $MSE(\hat{y}^{OLS}) = E(\hat{y}^{OLS} - f(x))^2 = \sigma^2_{OLS}$
#
# But, consider the ridge regression solution $\hat{y}_\lambda^R = E(\hat{y}^{OLS} / (1 + \lambda) - f(x))^2 = \frac{\sigma^2_{OLS}}{(1 + \lambda)^2} + \frac{\lambda^2}{(1 + \lambda)^2}f^2(x)$
#
# In some cases, the $\sigma_{OLS}$ can be very large due to singularity of $X^TX$. The shrinkage will increase bias but will reduce variance.
#
# Proof:
#
# $E(\frac{\hat{y}^{OLS}}{(1 + \lambda)} - f(x))$
#
# $= E[(\frac{\hat{y}}{1 + \lambda} - \frac{f(x)}{1 + \lambda}) - (\frac{\lambda}{1 + \lambda} f(x))]$
#
# $= \frac{\sigma^2_{OLS}}{1 + \lambda} + (\frac{\lambda}{1 + \lambda})^2 f^2(x)$
#
# ## PCA
#
# $X^T X = (V D U^T) (U D V^T) = U D V^T$
#
# $F = XV = UD$ where $f_j = X v_j$ is the projections to the PC directions and are the principal components.
#
# Note $f^T_j f_j = d^2_j$ and $f^T_i f_j = 0$ because orthogonal.
#
# So, with our PCs as $X$,
#
# $y = X\beta = y - UDV\beta = y - F \alpha = y - \sigma_{j=1}^p \alpha_j f_j$
#
# Regression after PCA agains all PCs
#
# $\alpha = V \beta$ and $||\alpha||_2 = ||\beta||_2$
#
# Therefore, the L2 norm of $\beta$ is the same as L2 norm of $\alpha$
#
# The ridge regression therefore
#
# $min_\beta ()...$
#
# is equivalent to
#
# $min_\alpha ()...$
#
# This allows $F = UD$ to satisfy the property $F^T F = DU^T UD = D^2$
#
# The ridge estimator
#
# $\hat{\alpha}_\lambda^R = (F^T F + \lambda I)^{-1} F^T y$
#
# $= diag(\frac{d_j}{d_j^2 + \lambda}) U^T y$
#
# $\hat{\alpha}_\lambda^R = (\frac{d_j^2}{d_j^2 + \lambda}) \hat{\alpha}_\lambda^{OLS} $
#
# The smaller the $d_j$, the more shrinkage.
#
# The prediction is $\hat{y}^R = F \alpha^R$
#
#
# ## Partial Least Squares
#
# Gradually capture the inform in $x$ corresponding to the information in $y$.
#
# $z_1 = \sum_{j=1}^p \phi_j x_j$
#
# where $\phi_j = \frac{<y, x_j>}{<x_j, x_j>} = <y, x_j>$
#
# We adjust the predictors by removing the effect containted on $Z_1$
#
# $e_j = X_j - Z_1 x_j$
#
# We do not want duplicate information. Info in $Z_1$ should not be in $Z_2$.
#
# ## Degrees of freedom
#
# We can evaluate the effective degree-of-freedom because
#
# In OLS, the trace of $H$ is $p$ because $\hat{y} = Hy = X(X^T X)^{-1}X^T y = (X^T X)^{-1}X^TX y = tr(I_p) = p$
#
# The ridge regerssion project $y$ to
#
# $\hat{\beta}^R_\lambda = (X^T X + \lambda I_p)^{-1} X^T y = S_\lambda y$
#
# We measure the effective degrees-of-freedom of ridge regression by
#
# $tr(S_\lambda) < p$
# ## Lasso
#
# Subset selection minimizes
#
# $(y - X\beta)^T (y - X\beta) + \lambda ||\beta||_1$
#
# where $1 = ||\beta||_1 = |\beta_1| + |\beta_2| = \sum_{j=1}^p |\beta_j|$. Therefore, a diamond (linear functions) is constructed.
#
# L2 norm encourages small $\beta_j$ but L1 will encourage sparse solutions.
#
#
#
|
HopML/Regularization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/FuriouStyles/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/module1-statistics-probability-and-inference/Stephen_P_LS_DS_131_Statistics_Probability_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="hml7gSN9WxG7" colab_type="text"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# ## *Data Science Unit 1 Sprint 3 Assignment 1*
#
# # Apply the t-test to real data
#
# Your assignment is to determine which issues have "statistically significant" differences between political parties in this [1980s congressional voting data](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records). The data consists of 435 instances (one for each congressperson), a class (democrat or republican), and 16 binary attributes (yes or no for voting for or against certain issues). Be aware - there are missing values!
#
# Your goals:
#
# 1. Load and clean the data (or determine the best method to drop observations when running tests)
# 2. Using hypothesis testing, find an issue that democrats support more than republicans with p < 0.01
# 3. Using hypothesis testing, find an issue that republicans support more than democrats with p < 0.01
# 4. Using hypothesis testing, find an issue where the difference between republicans and democrats has p > 0.1 (i.e. there may not be much of a difference)
#
# Note that this data will involve *2 sample* t-tests, because you're comparing averages across two groups (republicans and democrats) rather than a single group against a null hypothesis.
#
# Stretch goals:
#
# 1. Refactor your code into functions so it's easy to rerun with arbitrary variables
# 2. Apply hypothesis testing to your personal project data (for the purposes of this notebook you can type a summary of the hypothesis you formed and tested)
# + id="ousLF9T9X_GL" colab_type="code" colab={}
import pandas as pd
import numpy as np
from scipy.stats import ttest_ind, ttest_ind_from_stats, ttest_rel
# + id="psBYf9SfYoXx" colab_type="code" colab={}
df = pd.read_csv('house-votes-84.data', delimiter=',', header=None)
# + id="BI_jq799ZK6Z" colab_type="code" outputId="0293e0e8-53a6-49e8-8cc0-0c630fab8159" colab={"base_uri": "https://localhost:8080/", "height": 195}
df.head()
# + id="hhiAB8ioZQO6" colab_type="code" colab={}
cols = ['Class Name', 'handicapped-infants', 'water-project-cost-sharing', 'adoption-of-the-budget-resolution', 'physician-fee-freeze', 'el-salvador-aid', 'religious-groups-in-schools', 'anti-satellite-test-ban', 'aid-to-nicaraguan-contras', 'mx-missile', 'immigration', 'synfuels-corporation-cutback', 'education-spending', 'superfund-right-to-sue', 'crime', 'duty-free-exports', 'export-administration-act-south-africa']
df.columns = cols
# + id="0wrYXvgea0zY" colab_type="code" outputId="cf5d1bda-c2a7-4518-9e24-640e4f2770ed" colab={"base_uri": "https://localhost:8080/", "height": 266}
df = df.replace({'?': np.NaN, 'y': 1, 'n': 0})
df.head()
# + id="ZZrd4PC6durn" colab_type="code" colab={}
gop_votes = df[df['Class Name'] == 'republican']
dem_votes = df[df['Class Name'] == 'democrat']
# + id="fi0nQUcoQCe3" colab_type="code" colab={}
gop_crime_votes = gop_votes['crime']
dem_crime_votes = dem_votes['crime']
# + id="_McH4uu4XF5s" colab_type="code" outputId="4017963c-50bd-4e06-bb04-d1fac6c6cfbc" colab={"base_uri": "https://localhost:8080/", "height": 521}
df_grouped = df.groupby(df['Class Name']).mean()
df_grouped.T
# + [markdown] id="iUrCmuwVhLST" colab_type="text"
# ### Get ttest on the Crime Bill
# + id="na7cpT8PNA0f" colab_type="code" outputId="f65c6dd8-1db4-40e8-a06c-e3daf2e10d27" colab={"base_uri": "https://localhost:8080/", "height": 34}
ttest_ind(gop_crime_votes, dem_crime_votes, nan_policy='omit')
# + [markdown] id="tdb6A3qB4eFL" colab_type="text"
# With a p_value of 9.95 x 10^47, we can say that Republicans offer more support for the crime bill than Democrats in a statistically significant way
# + [markdown] id="Bvw__MGHhUTS" colab_type="text"
# ### Get ttest on the education-spending bill
# + id="27tXCIr8fjGq" colab_type="code" outputId="cc8562f0-8e8f-4341-9078-5f23bb195420" colab={"base_uri": "https://localhost:8080/", "height": 34}
gop_missile_votes = gop_votes['mx-missile']
dem_missile_votes = dem_votes['mx-missile']
ttest_ind(dem_missile_votes, gop_missile_votes, nan_policy='omit')
# + [markdown] id="MtHE5gzclE0p" colab_type="text"
# With p_value a 5.03 x 10^-47, we can reject the null hypothesis and confidently say that Democrats offer more support for the MX Missile bill than Republicans in a statistically significant way
# + [markdown] id="nQh-Uz3f5W9z" colab_type="text"
# ###Get ttest on the immigration bill
#
# + id="28yB6mjO5nNC" colab_type="code" outputId="a64874bd-f524-4a50-f4bd-bd43b18a9012" colab={"base_uri": "https://localhost:8080/", "height": 34}
gop_immigration_votes = gop_votes['immigration']
dem_immigration_votes = dem_votes['immigration']
ttest_ind(dem_immigration_votes, gop_immigration_votes, nan_policy='omit')
# + [markdown] id="o4N9RVH76OKb" colab_type="text"
# With a p_value of 0.08, we cannot confidently reject the null hypothesis, and cannot conclude that there is a signficant different that exists between Democrats and Republicans on this issue.
# + [markdown] id="6awBXqzndIne" colab_type="text"
# ##Stretch Goal - Refactor into a Function
#
#
#
# + id="-dI9XSvoNZQY" colab_type="code" colab={}
# We're going to hold on to gop_votes and dem_votes and use them here.
# -- I was made aware of a potential security flaw that by using gop_votes and dem_votes
# -- from the global scope I was introducing potential bugs and security vulerabilities.
# -- The below code should mitigate that, and it works as intended.
# x is the name of the bill in the column header in the dataframe
# The goal is to accept a dataframe, clean it, filter it, and successfully perform a ttest on it
def get_ttest(frame, x):
frame = frame.replace({'?': np.NaN, 'y': 1, 'n': 0})
gop_votes = frame[frame['Class Name'] == 'republican']
dem_votes = frame[frame['Class Name'] == 'democrat']
gop_bill_votes = gop_votes[x]
dem_bill_votes = dem_votes[x]
return ttest_ind(gop_bill_votes, dem_bill_votes, nan_policy='omit')
# + id="zUUuhRfKiyxh" colab_type="code" outputId="3d33b89e-100e-4115-cfc8-248027b38bab" colab={"base_uri": "https://localhost:8080/", "height": 34}
get_ttest(df, 'immigration')
|
module1-statistics-probability-and-inference/Stephen_P_LS_DS_131_Statistics_Probability_Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Blockmodel
#
#
# Example of creating a block model using the quotient_graph function in NX. Data
# used is the Hartford, CT drug users network::
#
# @article{weeks2002social,
# title={Social networks of drug users in high-risk sites: Finding the connections},
# url = {https://doi.org/10.1023/A:1015457400897},
# doi = {10.1023/A:1015457400897},
# author={Weeks, <NAME> and <NAME> and Borgatti, <NAME> and <NAME> and <NAME>},
# journal={{AIDS and Behavior}},
# volume={6},
# number={2},
# pages={193--206},
# year={2002},
# publisher={Springer}
# }
#
# +
# Authors: <NAME> <<EMAIL>>, <NAME> <<EMAIL>>
from collections import defaultdict
import matplotlib.pyplot as plt
import networkx as nx
import numpy
from scipy.cluster import hierarchy
from scipy.spatial import distance
def create_hc(G):
"""Creates hierarchical cluster of graph G from distance matrix"""
path_length = nx.all_pairs_shortest_path_length(G)
distances = numpy.zeros((len(G), len(G)))
for u, p in path_length:
for v, d in p.items():
distances[u][v] = d
# Create hierarchical cluster
Y = distance.squareform(distances)
Z = hierarchy.complete(Y) # Creates HC using farthest point linkage
# This partition selection is arbitrary, for illustrive purposes
membership = list(hierarchy.fcluster(Z, t=1.15))
# Create collection of lists for blockmodel
partition = defaultdict(list)
for n, p in zip(list(range(len(G))), membership):
partition[p].append(n)
return list(partition.values())
if __name__ == '__main__':
G = nx.read_edgelist("hartford_drug.edgelist")
# Extract largest connected component into graph H
H = G.subgraph(next(nx.connected_components(G)))
# Makes life easier to have consecutively labeled integer nodes
H = nx.convert_node_labels_to_integers(H)
# Create parititions with hierarchical clustering
partitions = create_hc(H)
# Build blockmodel graph
BM = nx.quotient_graph(H, partitions, relabel=True)
# Draw original graph
pos = nx.spring_layout(H, iterations=100)
plt.subplot(211)
nx.draw(H, pos, with_labels=False, node_size=10)
# Draw block model with weighted edges and nodes sized by number of internal nodes
node_size = [BM.nodes[x]['nnodes'] * 10 for x in BM.nodes()]
edge_width = [(2 * d['weight']) for (u, v, d) in BM.edges(data=True)]
# Set positions to mean of positions of internal nodes from original graph
posBM = {}
for n in BM:
xy = numpy.array([pos[u] for u in BM.nodes[n]['graph']])
posBM[n] = xy.mean(axis=0)
plt.subplot(212)
nx.draw(BM, posBM, node_size=node_size, width=edge_width, with_labels=False)
plt.axis('off')
plt.show()
|
NoSQL/NetworkX/plot_blockmodel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import time
Q1 = pd.read_csv('2007Q1_Charges.csv', index_col = 0)
Q1.head()
Q2 = pd.read_csv('2007Q2_Charges.csv', index_col = 0)
Q2.head()
Q3 = pd.read_csv('2007Q3_Charges.csv', index_col = 0)
Q3.head()
Q4 = pd.read_csv('2007Q4_Charges.csv', index_col = 0)
Q4.head()
Y2007 = pd.concat([Q1, Q2, Q3, Q4], sort = False)
Y2007.head()
Y2007.tail()
Y2007.shape
print(Q1.shape[0] + Q2.shape[0] + Q3.shape[0] + Q4.shape[0])
start = time.time()
Y2007.to_csv('Charges_07.csv')
end = time.time()
print(end - start)
|
2007/2007 Charges Merge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to implement a Proactive Chat widget
# This guide is intended to help Web developpers to implement a proactive chat widget on their website. This is the first section of a three part chapter:
#
# 1. How to implement a Proactive Chat widget
# 2. How to customize the Proactive chat widget
# 3. How to customize the Chat Window widget.
#
# At the end of this tutorial you will be able to create a webpage similar to the one below
# |  |
# |:--:|
# We will follow these steps:
#
#
# 1. Login to LH3
# 2. Creating and editing a Chat Snippet
# 3. Creating a test page in HTML
# 4. Test the result
#
# |  |
# |:--:|
# | *Proactive Chart Implementation FlowChart* |
# ### Login to LH3 Dashboard (Section 1/5)
# We will login to LH3 to create a Chat snipet.
# You need to have an admin account (or mini-admin account) in LH3. Plese can contact **Guinsly** the VR coordinator at SP to grant you an admin/create an admin account. ***guinsly [at] scholarsporta.info***
#
# action: login to [LibraryH3lp' deashboard](https://ca.libraryh3lp.com/dashboard/ "Google's Homepage")
#
# |  |
# |:--:|
# | *Login to LH3* |
# From the Navigation sidebar we will
# 1. click on **Chat widget**
# 2. click on **add**
#
# |  |
# |:--:|
# | *Login to LH3* |
# Type in the new name following this format.
#
# *. University [Language] (Button/Service type)
#
# example:
#
# 1. Western English (Proactive Chat)
# 2. UOIT (Chat tab)
# 3. Brock (Chat button)
#
from IPython.display import HTML, IFrame
HTML('<img src="./images/new_name.png" width="700px"/>')
# ### 2. Editing a Chat widget
#
# |  |
# |:--:|
# | *Proactive chat settings* |
# 1. This is where you want the new chat widget to point to. If you are a Home Team First school than you can select your school's queue. For the purpose of this tutorial we will choose **practice-webinars**.
# 2. Click on the **Proactive options** tab.
# 3. Select **Button**.
# 4. Select __where__ do you want to place the button.
# 5. Select a theme for your Proactive Chat. I selected the theme named **start** from the list.
# 7. Type the text that you want to display in the title bar. BTW My title is a bit too long. Therefore it will be truncated to fit the Chat widget
# 8. Type the invitation text or leave the default.
# 9. For a button (Step 3) we need to provide the choice to patrons to choose to accept or deny the chat request.
# 10. Type in the **Decline** text content.
#
# |  |
# |:--:|
# | *Proactive chat popup widget* |
# Delete the content found in the **Online appearance** tab.
#
# |  |
# |:--:|
# | *Delete content* |
# We will go down in the next section of the webpage
#
# 1. delete the content on **Offline appearance**
# 1. Click **SAVE**
#
# |  |
# |:--:|
# | *Saving the newly created Chat Snippet* |
# On the next page
# 1. click on the new Chat Snippet that you created
# * You can use **CTL+F** to search for your school name or the Chat Snippet number (i.e **899**)
# 2. Scroll to the top of the page and copy the Javascript snippet.
#
#
# + active=""
# <!-- Place this div in your web page where you want your chat widget to appear. -->
# <div class="needs-js">chat loading...</div>
#
# <!-- Place this script as near to the end of your BODY as possible. -->
# <script type="text/javascript">
# (function() {
# var x = document.createElement("script"); x.type = "text/javascript"; x.async = true;
# x.src = (document.location.protocol === "https:" ? "https://" : "http://") + "ca.libraryh3lp.com/js/libraryh3lp.js?899";
# var y = document.getElementsByTagName("script")[0]; y.parentNode.insertBefore(x, y);
# })();
# </script>
# -
# Make sure that your new Chat Snippet number (**899**) is include in the script.
# Starter Template **index.html**. We will use this starter template to test our newly created Chat Widget. Copy paste this content below in your text editor.
#
# + active=""
# <!doctype html>
# <html lang="en">
# <head>
# <!-- Required meta tags -->
# <meta charset="utf-8">
# <meta name="viewport" content="width=device-width, initial-scale=1, shrink-to-fit=no">
# <!-- Bootstrap CSS -->
# <link rel="stylesheet" href="https://maxcdn.bootstrapcdn.com/bootstrap/4.0.0/css/bootstrap.min.css" integrity="<KEY>" crossorigin="anonymous">
# <title>Hello, world!</title>
# </head>
# <body>
#
# <div class="jumbotron">
# <h1 class="display-4">Proactive Chat Example!</h1>
# <p class="lead">This is a simple proactive chat Example</p>
# <hr class="my-4">
# <p class="lead">
# <a class="btn btn-primary btn-lg" href="#" role="button" id="countdown">Learn more</a>
# </p>
# <div ></div>
# </div>
#
# <!-- Place this div in your web page where you want your chat widget to appear. -->
# <div class="needs-js">chat loading...</div>
#
# <!-- Place this script as near to the end of your BODY as possible. -->
# <script type="text/javascript">
# (function() {
# var x = document.createElement("script"); x.type = "text/javascript"; x.async = true;
# x.src = (document.location.protocol === "https:" ? "https://" : "http://") + "ca.libraryh3lp.com/js/libraryh3lp.js?899";
# var y = document.getElementsByTagName("script")[0]; y.parentNode.insertBefore(x, y);
# })();
#
#
# //JavaScript countdown
# var timeleft = 5;
# var downloadTimer = setInterval(function(){
# document.getElementById("countdown").innerHTML = timeleft + " seconds remaining";
# timeleft -= 1;
# if(timeleft <= 0){
# clearInterval(downloadTimer);
# document.getElementById("countdown").innerHTML = "Voila!"
# }
# }, 1000);
# </script>
# </body>
# </html>
# -
# Pay attention to the content.
# 1. This is the default message when the script is loading. It will not appear unless that we have a slow web connection
# 2. The chat widget number is in the script.
#
#
#
# |  |
# |:--:|
# | *Script* |
# # Voilà!!!
# Open that file (**index.html**) in your browser. Voilà!!!
#
# Make sure that the Service is on by login in the [LibraryH3lp' Weblient](https://ca.libraryh3lp.com/webclient/ "LH3 Webclient") (this is different from the Dashboard. If you are sending your Chats to "Pratice" Make sure that you are login to the practice webinar. The Proactive Chat widget will not pop up if the service is **off**
# |  |
# |:--:|
|
LH3_Proactive_Chat_Implementation_docs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#uberduck_ml_dev.exec.gather_dataset" data-toc-modified-id="uberduck_ml_dev.exec.gather_dataset-1"><span class="toc-item-num">1 </span>uberduck_ml_dev.exec.gather_dataset</a></span></li></ul></div>
# +
# default_exp exec.gather_dataset
# -
# # uberduck_ml_dev.exec.gather_dataset
# Create a zipped dataset from a filelist.
#
# Usage:
#
# ```
# python -m uberduck_ml_dev.exec.gather_dataset \
# --input filelist.txt \
# --ouput out.zip
# ```
#
# +
# export
import argparse
import os
from tempfile import NamedTemporaryFile
from typing import List
import sys
from zipfile import ZipFile
def _gather(filelist, output):
with open(filelist, "r") as f:
lines = f.readlines()
paths = []
for line in lines:
path, *_rest = line.split("|")
paths = [l.split("|")[0] for l in lines]
common_prefix = os.path.commonpath(paths)
archive_paths = []
archive_lines = []
for line in lines:
p, txn, *_rest = line.split("|")
relpath = os.path.relpath(p, common_prefix)
archive_paths.append(relpath)
archive_lines.append(f"{relpath}|{txn}|{''.join(_rest)}")
_, filelist_archive = os.path.split(filelist)
with NamedTemporaryFile("w") as tempfile:
for line in archive_lines:
tempfile.write(line)
tempfile.flush()
with ZipFile(output, "w") as zf:
zf.write(tempfile.name, filelist_archive)
for path, archive_path in zip(paths, archive_paths):
zf.write(path, archive_path)
def _parse_args(args: List[str]):
parser = argparse.ArgumentParser()
parser.add_argument("-i", "--input", help="Path to input filelist")
parser.add_argument(
"-o", "--output", help="Output zipfile", default="out.zip",
)
return parser.parse_args(args)
try:
from nbdev.imports import IN_NOTEBOOK
except:
IN_NOTEBOOK = False
if __name__ == "__main__" and not IN_NOTEBOOK:
args = _parse_args(sys.argv[1:])
_gather(args.input, args.output)
# -
# hide
_parse_args(["-i", "foo/bar.txt", "--output", "foo.zip"])
|
nbs/exec.gather_dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Conditional non-linear systems of equations
# Sometimes when performing modelling work in physical sciences we use different sets of equations to describe our system depending on conditions. Sometimes it is not known beforehand which of those formulations that will be applicable (only after having solved the system of equations can we reject or accept the answer). ``pyneqsys`` provides facilities to handle precisely this situation.
from __future__ import (absolute_import, division, print_function)
from functools import reduce
from operator import mul
import sympy as sp
import numpy as np
import matplotlib.pyplot as plt
from pyneqsys.symbolic import SymbolicSys, linear_exprs
sp.init_printing()
# Let's consider precipitation/dissolution of NaCl:
# $$
# \rm NaCl(s) \rightleftharpoons Na^+(aq) + Cl^-(aq)
# $$
init_concs = iNa_p, iCl_m, iNaCl = [sp.Symbol('i_'+str(i), real=True, negative=False) for i in range(3)]
c = Na_p, Cl_m, NaCl = [sp.Symbol('c_'+str(i), real=True, negative=False) for i in range(3)]
prod = lambda x: reduce(mul, x)
texnames = [r'\mathrm{%s}' % k for k in 'Na^+ Cl^- NaCl'.split()]
# if the solution is saturated, then the [solubility product](https://en.wikipedia.org/wiki/Solubility_equilibrium) will be constant:
#
# $$
# K_{\rm sp} = \mathrm{[Na^+][Cl^-]}
# $$
#
# in addition to this (conditial realtion) we can write equations for the preservation of atoms and charge:
# +
stoichs = [[1, 1, -1]]
Na = [1, 0, 1]
Cl = [0, 1, 1]
charge = [1, -1, 0]
preserv = [Na, Cl, charge]
eq_constants = [Ksp] = [sp.Symbol('K_{sp}', real=True, positive=True)]
def get_f(x, params, saturated):
init_concs = params[:3] if saturated else params[:2]
eq_constants = params[3:]
le = linear_exprs(preserv, x, linear_exprs(preserv, init_concs), rref=True)
return le + ([Na_p*Cl_m - Ksp] if saturated else [NaCl])
# -
# Our two sets of reactions are then:
get_f(c, init_concs + eq_constants, False)
f_true = get_f(c, init_concs + eq_constants, True)
f_false = get_f(c, init_concs + eq_constants, False)
f_true, f_false
# We have one condition (a boolean describing whether the solution is saturated or not). We provide two conditionals, one for going from non-saturated to saturated (forward) and one going from saturated to non-saturated (backward):
from pyneqsys.core import ConditionalNeqSys
cneqsys = ConditionalNeqSys(
[
(lambda x, p: (x[0] + x[2]) * (x[1] + x[2]) > p[3], # forward condition
lambda x, p: x[2] >= 0) # backward condition
],
lambda conds: SymbolicSys(
c, f_true if conds[0] else f_false, init_concs+eq_constants
),
latex_names=['[%s]' % n for n in texnames], latex_param_names=['[%s]_0' % n for n in texnames]
)
c0, K = [0.5, 0.5, 0], [1] # Ksp for NaCl(aq) isn't 1 in reality, but used here for illustration
params = c0 + K
# Solving for inital concentrations below the solubility product:
cneqsys.solve([0.5, 0.5, 0], params)
# no surprises there (it is of course trivial).
#
# In order to illustrate its usefulness, let us consider addition of a more soluable sodium salt (e.g. NaOH) to a chloride rich solution (e.g. HCl):
# %matplotlib inline
ax_out = plt.subplot(1, 2, 1)
ax_err = plt.subplot(1, 2, 2)
xres, sols = cneqsys.solve_and_plot_series(
c0, params, np.linspace(0, 3), 0, 'kinsol',
{'ax': ax_out}, {'ax': ax_err}, fnormtol=1e-14)
_ = ax_out.legend()
# note the (expected) discontinuity at $\mathrm{[Na^+]_0 = 2}$ at which point the solution became saturated
|
examples/conditional.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recreating k-sys Realtime R0 analysis
# #### This is a recreation of k-sys covid-19 repository where he uses Bettencourt and Ribeiro bayesian algorithm to estimate the most likely value of Rt and a credible interval.
#
# <details>
# <summary><strong>Goal</strong></summary>
# "The goal of this notebook is to ..."
# <ul>
# <li> measurable goals for this notebook </li>
# <li> Ex 1: Identify <i>variables of interest in dataset <strong>name.dta</strong></i> </li>
# <li> Ex 2: Create a <i>working dataset</i> from raw data.</li>
# </ul>
# </details>
#
# <details>
# <summary><strong>Context</strong></summary>
# Context at the moment (i.e. "We've downloaded raw data from <strong><i>website.com</i></strong> and are now in the process of creating a mastefile.")
# <li> What is the final goal of this project?</li>
# <li> What are we trying to recreate? Where are we in the process?</li>
# <li> Any links to documentation / figures.</li>
# </details>
# +
import pandas as pd
from zipfile import ZipFile
from pathlib import Path
from tools import tree
from datetime import datetime as dt
today = dt.today().strftime("%d-%b-%y")
today
# -
RAW_DATA = Path("../data/raw/")
INTERIM_DATA = Path("../data/interim/")
PROCESSED_DATA = Path("../data/processed/")
FINAL_DATA = Path("../data/final/")
tree(RAW_DATA)
data = pd.read_
|
notebooks/00_DataPrep.ipynb
|