code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import ipyrad
import ipyrad.analysis as ipa
import ipcoal
import matplotlib.pyplot as plt
import msprime
import numpy as np
import toytree
import toyplot
print(ipyrad.__version__)
# +
# Many loci
def simloci(sample_size_pop1=10,
sample_size_pop2=10,
get_pis=False,
TMRCA=220e3,
Ne=100000,
Ne_ratio=1,
mig=3e-3,
sequence_length=100,
num_replicates=10,
debug=False):
T_MRCA = TMRCA
m_a_u = mig
m_b_u = mig
migmat = [[0, mig], [mig, 0]]
sample_size_a=sample_size_pop1
sample_size_b=sample_size_pop2
ancpopa = msprime.PopulationConfiguration(sample_size=sample_size_a, initial_size=Ne)
ancpopb = msprime.PopulationConfiguration(sample_size=sample_size_b, initial_size=Ne/Ne_ratio)
## All lineages coalesce at some point in the past
atou_event = msprime.MassMigration(time=T_MRCA, source=0, destination=1, proportion=1.0)
# au_change = msprime.MigrationRateChange(time=T_Extinction, rate=m_a_u, matrix_index=(0,1))
# ua_change = msprime.MigrationRateChange(time=T_Extinction, rate=m_a_u, matrix_index=(1,0))
demographic_events = [atou_event]
dp = msprime.DemographyDebugger(
Ne=Ne,
population_configurations=[ancpopa, ancpopb],
demographic_events=demographic_events)
if debug:
dp.print_history()
ts = msprime.simulate(length=sequence_length,
Ne=Ne,
mutation_rate=1e-8,
migration_matrix=migmat,
population_configurations=[ancpopa, ancpopb],
demographic_events=demographic_events,
num_replicates=num_replicates)
if debug:
tree = next(next(ts).trees())
f = "/tmp/watt.svg"
tree.draw(f, height=600, width=600)
show_svg(f)
pis0 = []
pis2 = []
divs = []
sfss = []
for tre in ts:
pop0 = tre.get_samples(0)
pop2 = tre.get_samples(1)
#pis0.append(tre.get_pairwise_diversity(pop0)/sequence_length)
#pis2.append(tre.get_pairwise_diversity(pop2)/sequence_length)
#divs.append(tre.divergence(sample_sets=[pop0, pop2]))
sfss.append(tre.allele_frequency_spectrum([pop0, pop2], polarised=False, span_normalise=False))
#return pis0, pis2, divs, sfss
return np.mean(pis0), np.mean(pis2), np.mean(divs), (np.sum(sfss, axis=0)/np.sum(sfss)).flatten()
_, _, _, sfs = simloci(num_replicates=100)
sfs
# -
plt.imshow(np.reshape(sfs, (11, 11)), cmap="gray_r")
# +
# generate a balance tree
tree = toytree.rtree.baltree(ntips=2, treeheight=1e5)
# draw the tree w/ an admixture edge
tree.draw(ts='p');
# -
|
jupyter-notebooks/easySFS_simdata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="zZavc8oCeKil"
# By working for the company, we have access to each customer’s app behaviour data. This data allows us to see the date and time of app installation,as well as the features the users engaged with within the app.App behaviour is characterized as the list of app screens the user looked at, and whether the user looked at,and where the user played the financial mini-games available.The app usage data is only from the user’s first day in the app. This limitation exists because users can enjoy a 24-hour free trial of the premium features,and the company wants to target them with new offers shortly after the trial is over.
#
# + colab={} colab_type="code" id="oCMDjev_6o4W"
#### Importing Libraries ####
import pandas as pd
from dateutil import parser
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="Cl0BgtKh6vej" outputId="f2a8f96b-5428-4658-838f-94fb2ccc1b83"
#Running or Importing .py Files with Google Colab
from google.colab import drive
drive.mount('/content/drive/')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="NP5efe3P6yyh" outputId="fdaf99cf-179f-4572-f8da-a38c94ec5114"
dataset = pd.read_csv("/content/drive/My Drive/app/appdata10.csv")
dataset
# + [markdown] colab_type="text" id="xgxbPhmzPp-b"
# #### Exploratory Data Analysis(EDA)
# + colab={"base_uri": "https://localhost:8080/", "height": 294} colab_type="code" id="sMoX59US7Xyk" outputId="85a8ada4-c33f-43c7-c755-d3fcd2ab498f"
dataset.head(10) # Viewing the Data
dataset.describe() # Distribution of Numerical Variables
# First set of Feature cleaning
dataset["hour"] = dataset.hour.str.slice(1, 3).astype(int)
### Plotting
dataset2 = dataset.copy().drop(columns = ['user', 'screen_list', 'enrolled_date',
'first_open', 'enrolled'])
dataset2.head()
## Histograms
plt.suptitle('Histograms of Numerical Columns', fontsize=20)
for i in range(1, dataset2.shape[1] + 1):
plt.subplot(3, 3, i)
f = plt.gca()
# f.axes.get_yaxis().set_visible(False)
f.set_title(dataset2.columns.values[i - 1])
vals = np.size(dataset2.iloc[:, i - 1].unique())
plt.hist(dataset2.iloc[:, i - 1], bins=vals, color='#3F5D7D')
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
#plt.savefig('app_data_hist.jpg')
# + colab={"base_uri": "https://localhost:8080/", "height": 732} colab_type="code" id="rPmE_ZhE7ebM" outputId="acffa56a-6cff-4691-e848-462a7e743970"
## Correlation with Response Variable
dataset2.corrwith(dataset.enrolled).plot.bar(figsize=(20,10),
title = 'Correlation with Reposnse variable',
fontsize = 15, rot = 45,
grid = True)
## Correlation Matrix
sns.set(style="white", font_scale=2)
# Compute the correlation matrix
corr = dataset2.corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="j9gJ6Oj47vgG" outputId="65abddef-61c7-4fa3-d391-17f62989163f"
#Correlation Matrix
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(18, 15))
f.suptitle("Correlation Matrix", fontsize = 40)
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# + [markdown] colab_type="text" id="LAZCn0b9P87g"
# #### Feature Engineering ####
# + colab={"base_uri": "https://localhost:8080/", "height": 589} colab_type="code" id="Q4SJZpxP7lvv" outputId="4b053f04-7ba9-47be-b6c8-281d404b2700"
# Formatting Date Columns
dataset.dtypes
dataset["first_open"] = [parser.parse(row_date) for row_date in dataset["first_open"]]
dataset["enrolled_date"] = [parser.parse(row_date) if isinstance(row_date, str) else row_date for row_date in dataset["enrolled_date"]]
dataset.dtypes
# Selecting Time For Response
dataset["difference"] = (dataset.enrolled_date-dataset.first_open).astype('timedelta64[h]')
response_hist = plt.hist(dataset["difference"].dropna(), color='#3F5D7D')
plt.title('Distribution of Time-Since-Screen-Reached')
plt.show()
plt.hist(dataset["difference"].dropna(), color='#3F5D7D', range = [0, 100])
plt.title('Distribution of Time-Since-Screen-Reached')
plt.show()
dataset.loc[dataset.difference > 48, 'enrolled'] = 0
dataset = dataset.drop(columns=['enrolled_date', 'difference', 'first_open'])
# + colab={} colab_type="code" id="VGy2afnq8YYJ"
## Formatting the screen_list Field
# Load Top Screens
top_screens = pd.read_csv("/content/drive/My Drive/app/top_screens.csv").top_screens.values
top_screens
# Mapping Screens to Fields
dataset["screen_list"] = dataset.screen_list.astype(str) + ','
for sc in top_screens:
dataset[sc] = dataset.screen_list.str.contains(sc).astype(int)
dataset['screen_list'] = dataset.screen_list.str.replace(sc+",", "")
dataset['Other'] = dataset.screen_list.str.count(",")
dataset = dataset.drop(columns=['screen_list'])
# Funnels
savings_screens = ["Saving1",
"Saving2",
"Saving2Amount",
"Saving4",
"Saving5",
"Saving6",
"Saving7",
"Saving8",
"Saving9",
"Saving10"]
dataset["SavingCount"] = dataset[savings_screens].sum(axis=1)
dataset = dataset.drop(columns=savings_screens)
cm_screens = ["Credit1",
"Credit2",
"Credit3",
"Credit3Container",
"Credit3Dashboard"]
dataset["CMCount"] = dataset[cm_screens].sum(axis=1)
dataset = dataset.drop(columns=cm_screens)
cc_screens = ["CC1",
"CC1Category",
"CC3"]
dataset["CCCount"] = dataset[cc_screens].sum(axis=1)
dataset = dataset.drop(columns=cc_screens)
loan_screens = ["Loan",
"Loan2",
"Loan3",
"Loan4"]
dataset["LoansCount"] = dataset[loan_screens].sum(axis=1)
dataset = dataset.drop(columns=loan_screens)
#### Saving Results ####
dataset.head()
dataset.describe()
dataset.columns
dataset.to_csv('new_appdata10.csv', index = False)
# + colab={} colab_type="code" id="XnLL4dZo81f7"
#### Data Pre-Processing ####
# Splitting Independent and Response Variables
response = dataset["enrolled"]
dataset = dataset.drop(columns="enrolled")
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(dataset, response,
test_size = 0.2,
random_state = 0)
# + colab={} colab_type="code" id="gqbGSE0JPJRm"
#Balancing the Training Set
import random
y_train.value_counts()
pos_index = y_train[y_train.values == 1].index
neg_index = y_train[y_train.values == 0].index
if len(pos_index) > len(neg_index):
higher = pos_index
lower = neg_index
else:
higher = neg_index
lower = pos_index
random.seed(0)
higher = np.random.choice(higher, size=len(lower))
lower = np.asarray(lower)
new_indexes = np.concatenate((lower, higher))
# + colab={} colab_type="code" id="i2iAcXw-PTC-"
X_train = X_train.loc[new_indexes,]
y_train = y_train[new_indexes]
# + colab={} colab_type="code" id="dq1ooPOE9g-A"
# Removing Identifiers
train_identity = X_train['user']
X_train = X_train.drop(columns = ['user'])
test_identity = X_test['user']
X_test = X_test.drop(columns = ['user'])
# + colab={} colab_type="code" id="7TdbLLba9ji6"
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_train2 = pd.DataFrame(sc_X.fit_transform(X_train))
X_test2 = pd.DataFrame(sc_X.transform(X_test))
X_train2.columns = X_train.columns.values
X_test2.columns = X_test.columns.values
X_train2.index = X_train.index.values
X_test2.index = X_test.index.values
X_train = X_train2
X_test = X_test2
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="R45S1WEF9MZX" outputId="66b11c3a-22ff-4027-d6a7-61960714dc86"
#### Model Building ####
# Fitting Model to the Training Set
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state = 0, penalty = 'l1')
classifier.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Ja4n15t_9QPZ" outputId="541f9680-9a90-4329-988f-0e6a5fbc41fe"
# Predicting Test Set
y_pred = classifier.predict(X_test)
y_pred
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="w2QbXdFh9Thv" outputId="6e5fdebe-0608-4632-f064-7630319dadec"
# Evaluating Results
from sklearn.metrics import confusion_matrix, accuracy_score, f1_score, precision_score, recall_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
print(accuracy_score(y_test, y_pred))
print(precision_score(y_test, y_pred)) # tp / (tp + fp)
print(recall_score(y_test, y_pred)) # tp / (tp + fn)
print(f1_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/", "height": 459} colab_type="code" id="Rtg3D5ndQOFP" outputId="84eac30c-4934-4c95-bc5e-f0edc4e5bcb9"
df_cm = pd.DataFrame(cm, index = (0, 1), columns = (0, 1))
plt.figure(figsize = (10,7))
sns.set(font_scale=1.4)
sns.heatmap(df_cm, annot=True, fmt='g')
print("Test Data Accuracy: %0.4f" % accuracy_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/", "height": 374} colab_type="code" id="dUnKWpPO9qvK" outputId="2e2f7b44-e175-49cb-d26b-76b9fc2ebe5f"
# Applying k-Fold Cross Validation
from sklearn.model_selection import cross_val_score
accuracies = cross_val_score(estimator = classifier, X = X_train, y = y_train, cv = 10)
print("SVM Accuracy: %0.3f (+/- %0.3f)" % (accuracies.mean(), accuracies.std() * 2))
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="8N69mUCA9ver" outputId="cb676f01-1979-4199-e4e9-30514c7773ac"
# Analyzing Coefficients
pd.concat([pd.DataFrame(dataset.drop(columns = 'user').columns, columns = ["features"]),
pd.DataFrame(np.transpose(classifier.coef_), columns = ["coef"])
],axis = 1)
# + [markdown] colab_type="text" id="62PtyghGOcpM"
# #### Model Tuning ####
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="j822lIkv8_TV" outputId="a28f5e01-d73b-4a5b-e79f-a4c95c67d879"
## Grid Search (Round 1)
import time
from sklearn.model_selection import GridSearchCV
# Select Regularization Method
penalty = ['l1', 'l2']
# Create regularization hyperparameter space
C = [0.001, 0.01, 0.1, 1, 10, 100, 1000]
# Combine Parameters
parameters = dict(C=C, penalty=penalty)
grid_search = GridSearchCV(estimator = classifier,param_grid = parameters,scoring = "accuracy",cv = 10,n_jobs = -1)
t0 = time.time()
grid_search = grid_search.fit(X_train, y_train)
t1 = time.time()
print("Took %0.2f seconds" % (t1 - t0))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="1mi9cPYJOd6s" outputId="be1ae4cb-b80c-4b28-a358-71d41de30995"
rf_best_accuracy = grid_search.best_score_
rf_best_parameters = grid_search.best_params_
rf_best_accuracy, rf_best_parameters
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="NVHIcV7_9CTf" outputId="7c4789c1-0a14-40f9-a685-c707cb477943"
## Grid Search (Round 2)
# Select Regularization Method
penalty = ['l1', 'l2']
# Create regularization hyperparameter space
C = [0.1, 0.5, 0.9, 1, 2, 5]
# Combine Parameters
parameters = dict(C=C, penalty=penalty)
grid_search = GridSearchCV(estimator = classifier,param_grid = parameters,scoring = "accuracy",cv = 10,n_jobs = -1)
t0 = time.time()
grid_search = grid_search.fit(X_train, y_train)
t1 = time.time()
print("Took %0.2f seconds" % (t1 - t0))
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="57E_TYKzO7F_" outputId="755b4378-f924-42fb-d08f-9c62cb522031"
rf_best_accuracy = grid_search.best_score_
rf_best_parameters = grid_search.best_params_
print(rf_best_accuracy, rf_best_parameters)
print(grid_search.best_score_)
# + [markdown] colab_type="text" id="OlLfu5rSO9g2"
# #### End of Model ####
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="j7X1iGyD9E_2" outputId="ce6435d1-b967-4b38-92a4-4032d4272109"
# Formatting Final Results
final_results = pd.concat([y_test, test_identity], axis = 1).dropna()
final_results['predicted_reach'] = y_pred
final_results = final_results[['user', 'enrolled', 'predicted_reach']].reset_index(drop=True)
final_results
|
Data-Science-Portfolio-master/Python/Directing_Customers_to_Subscription_Through_App_Behavior_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scraping using BeautifulSoup, Pandas, and Requests/Splinter
# +
## Dependencies
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all'
import requests
from splinter import Browser
from bs4 import BeautifulSoup as bs
import time
import pandas as pd
import re
from urls_list import * #where all urls and paths are saved
# -
# ## NASA Mars News
#
# * **Scrape the NASA Mars News Site and collect the latest News Title and Paragraph Text. Assign the text to variables that we can reference later.**
def scrape_latest_news():
#########################################################################################
#Scrape the latest news
#Returns news_title, news_p
#########################################################################################
news_title, news_p = None, None
#Configure Browser
browser = Browser(browser_choice, executable_path=executable_path, headless=True)
try:
#Visit url
browser.visit(nasa_mars_news)
#bs object with lxml parser
time.sleep(4)#This is very important
soup = bs(browser.html, 'lxml')
news = soup.find('li', class_='slide').div.find(class_='list_text').find_all('div')
news_title, news_p = news[1].text, news[2].text
except Exception as e:
print(e)
#Close browser to avoid resource issue
browser.quit()
return (news_title, news_p)
news_title, news_p = scrape_latest_news()
news_title
news_p
# ## JPL Mars Space Images - Featured Image
#
# * **Find the image url for the current Featured Mars Image and assign the url string to a variable called featured_image_url**
def scrape_featured_image_url():
#########################################################################################
#Scrape the featured image url from nasa jpl site
#Returns featured_image_url
#########################################################################################
featured_image_url = None
#Configure Browser
browser = Browser(browser_choice, executable_path=executable_path, headless=True)
try:
#Visit url
browser.visit(nasa_jpl)
#bs object with lxml parser
time.sleep(4)#This is very important
soup = bs(browser.html, 'lxml')
#Click a button "FULL IMAGE"
browser.find_by_id('full_image', wait_time=1).click()
#Click more info button
browser.find_by_css('[id="fancybox-lock"]')[0].find_by_css('div[class="buttons"] a:nth-child(2)')[0].click()
time.sleep(1)
#Take the image link (largesize)
featured_image_url = browser.find_by_css('figure[class="lede"] a')['href']
except Exception as e:
print(e)
#Close browser to avoid resource issue
browser.quit()
return featured_image_url
# +
featured_image_url = scrape_featured_image_url()
# -
featured_image_url
# ## Mars Weather - from twitter page
#
# * **Visit the Mars Weather twitter account and scrape the latest Mars weather tweet from the page. Save the tweet text for the weather report as a variable called mars_weather.**
#
# * **Note: Be sure you are not signed in to twitter, or scraping may become more difficult.**
#
# * **Note: Twitter frequently changes how information is presented on their website.**
def scrape_mars_weather():
#########################################################################################
#Scrape the latest Mars weather tweet from the twitter page
#Returns mars_weather
#########################################################################################
mars_weather = None
#Configure Browser
browser = Browser(browser_choice, executable_path=executable_path, headless=True)
try:
#Visit url
browser.visit(mars_twitter_page)
#bs object with lxml parser
time.sleep(4)#This is very important
soup = bs(browser.html, 'lxml')
#Extract the weather info using soup css selector
mars_weather = soup.find('div', attrs={'data-testid':'tweet'}).select('div:nth-of-type(2) > div:nth-of-type(2) > div:nth-of-type(1) > div:nth-of-type(1) > span')[0].text
except Exception as e:
print(e)
#Close browser to avoid resource issue
browser.quit()
return mars_weather
scrape_mars_weather()
# ## Mars Facts
#
# * **Visit the Mars Facts webpage here and use Pandas to scrape the table containing facts about the planet including Diameter, Mass, etc.**
#
# * **Use Pandas to convert the data to a HTML table string.**
#
DF = pd.read_html(mars_facts, attrs={'id':'tablepress-p-mars'})[0]
DF.rename(columns={0:'attributes', 1:'value'}, inplace=True)
DF
# ## Mars Hemispheres - from USGS Astrogeology site
#
# * **Visit the USGS Astrogeology site to obtain high resolution images for each of Mar's hemispheres.**
#
# * **Save both the image url string for the full resolution hemisphere image, and the Hemisphere title containing the hemisphere name. Use a Python dictionary to store the data using the keys img_url and title.**
#
# * **Append the dictionary with the image url string and the hemisphere title to a list. This list will contain one dictionary for each hemisphere.**
def scrape_hemispheres():
#########################################################################################
#Scrape high resolution images for each of Mar's hemispheres from USGS Astrogeology site
#Returns list of dictionaries with title and image urls
#########################################################################################
hemisphere_image_urls = []
#Configure Browser
browser = Browser(browser_choice, executable_path=executable_path, headless=True)
try:
#Visit url
browser.visit(usgs_search)
#bs object with lxml parser
time.sleep(10)#This is very important (site loads super slow)
soup = bs(browser.html, 'lxml')
hs_links = soup.find(id='product-section').find_all('a',class_="itemLink product-item")
for index,link in enumerate(hs_links):
if link.img is None:
title = re.sub(' Enhanced', '', link.text)
else:
browser.visit(usgs_base+link['href'])
time.sleep(1)
img_url = browser.find_by_css('img[class="wide-image"]')['src']
if index%2:#Image and title come together
hemisphere_image_urls.append({'title':title,'img_url':img_url})
except Exception as e:
print(e)
#Close browser to avoid resource issue
browser.quit()
return hemisphere_image_urls
hemisphere_image_urls = scrape_hemispheres()
hemisphere_image_urls
|
Analysis_Code/mission_to_mars.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
import cufflinks as cf
# -
fn=r'/media/Data1/pyWorkDir/Bigdata/Pyspark/data/COVID19/OntarioTorontOttawaCOVID-19ConfirmedCases.csv'
df = pd.read_csv(fn)
df=df.set_index('Date_health_region')
#print(df.dtypes)
df.head(1)
@interact
def plot_Ontario():
df.iplot(xTitle='Date', yTitle='Convirmed Cases',title='Ontario COVID-19 Confirmed Cases',theme='solar')
|
codes/A05_OntarioTorontoOttawaCovid19_run05.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:NLP_DEMO]
# language: python
# name: conda-env-NLP_DEMO-py
# ---
# +
import spacy
nlp = spacy.load("zh_core_web_md")
doc = nlp("今天天气不错,挺风和日丽的,我们下午没有课。")
for token in doc:
print(token.text, token.lemma_, token.pos_, token.tag_, token.dep_,
token.shape_, token.is_alpha, token.is_stop)
# +
from spacy import displacy
html = displacy.render(doc, style="dep",options = {'distance':105})
# -
|
src/NLP_visualization/Dependency Parsing 依存句法可视化.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.metrics import mean_absolute_error as mae
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# !wget https://raw.githubusercontent.com/dataworkshop/5dwchallenge_2019/master/challenge5/input/airmiles.csv
# !ls
df = pd.read_csv("airmiles.csv")
df.head()
plt.figure(figsize=(15, 5))
plt.plot(df.time, df.airmiles, 'o-');
df = pd.read_csv("airmiles.csv", parse_dates=["time"], index_col="time")
df[['airmiles']].plot();
# +
def linear_func(airmiles, k=None, b=0):
mean_value = np.mean(airmiles)
if k is None:
return [mean_value] * len(airmiles)
return [idx*k + b for idx,_ in enumerate(airmiles)]
plt.figure(figsize=(15, 5))
plt.plot(df.index, df.airmiles, 'o-', label='original');
plt.plot(df.index, linear_func(df.airmiles), 'x-', label='mean');
plt.plot(df.index, linear_func(df.airmiles, 1300, -3000), 'x-', label='linear');
plt.legend();
# -
# ## Metryka sukcesu
print('mean', mae(df.airmiles, linear_func(df.airmiles)))
print('mean', mae(df.airmiles, linear_func(df.airmiles, 1300, -3000)))
# +
best_k = 1300
best_b = -3000
best_mae = mae(df.airmiles, linear_func(df.airmiles, best_k, -best_b))
for k in range(1000, 1400, 50):
for b in (-4000, -2000, 50):
actual = mae(df.airmiles, linear_func(df.airmiles, k, b))
if actual < best_mae:
best_mae = actual
best_k = k
best_b = b
print("Best mea={} for k={} and b={}".format(best_mae, best_k, best_b))
|
week5/day2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Best Time to Buy and Sell Stock II
# + active=""
# Say you have an array for which the ith element is the price of a given stock on day i.
# Design an algorithm to find the maximum profit. You may complete as many transactions as you like
# (i.e., buy one and sell one share of the stock multiple times).
# Note: You may not engage in multiple transactions at the same time (i.e., you must sell the stock before you buy again).
#
# Example 1: # 能多 n 次交易 找到最佳利潤~
# Input: [7,1,5,3,6,4]
# Output: 7
# Explanation: Buy on day 2 (price = 1) and sell on day 3 (price = 5), profit = 5-1 = 4.
# Then buy on day 4 (price = 3) and sell on day 5 (price = 6), profit = 6-3 = 3.
# Example 2:
# Input: [1,2,3,4,5]
# Output: 4
# Explanation: Buy on day 1 (price = 1) and sell on day 5 (price = 5), profit = 5-1 = 4.
# Note that you cannot buy on day 1, buy on day 2 and sell them later, as you are
# engaging multiple transactions at the same time. You must sell before buying again.
# Example 3:
# Input: [7,6,4,3,1]
# Output: 0
# Explanation: In this case, no transaction is done, i.e. max profit = 0.
# +
class Solution:
def maxProfit(self, prices): # 62.97%
"""
:type prices: List[int]
:rtype: int
"""
if prices == []: # 沒資料的狀況
return 0
profit = 0
for ind in range(1, len(prices)):
profit += max(0, (prices[ind] - prices[ind-1])) # 抓到每次的差價 <0 的算 0, >0 加入profit
return profit
###############################################################
def maxProfit2(self, prices): # 45 %
if prices == []: # 沒資料的狀況
return 0
yester = prices[0]
profit = 0
all_profit = 0
for p in prices[1:]:
if p > yester:
profit += (p-yester)
elif p < yester: # 往下就
all_profit += profit
profit = 0
yester = p
all_profit += profit
return all_profit
# -
prices = [7,1,5,3,6,4,5,7,1,5]
ans = Solution()
ans.maxProfit(prices)
|
122. Best Time to Buy and Sell Stock II.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2
# language: sage
# name: sagemath
# ---
# +
q = 16798108731015832284940804142231733909889187121439069848933715426072753864723
F = GF(q)
K2.<x> = PolynomialRing(F)
K.<a> = GF(q**2, name='a', modulus=x ^ 2 + 1)
F2.<u> = F.extension(x ^ 2 + 1)
K6.<y> = PolynomialRing(F2)
F6.<v> = F2.extension(y ^ 3 - (u + 1))
# towering is extermely slow in sage, be careful
# -
(u + 1) ^ ((q - 1) / 3)
(u + 1) ^ ((q^2 - 1) / 3)
(u + 1) ^ ((q^3 - 1) / 3)
(u + 1) ^ ((q^4 - 1) / 3)
(u + 1) ^ ((q^5 - 1) / 3)
(u + 1) ^ ((2*q - 2) / 3)
(u + 1) ^ ((2*(q^2) - 2) / 3)
(u + 1) ^ ((2*(q^3) - 2) / 3)
(u + 1) ^ ((2*(q^4) - 2) / 3)
(u + 1) ^ ((2*(q^5) - 2) / 3)
|
src/curve_bn254/sage_scripts/Fq6.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (Lab 1)
# language: python
# name: pycharm-8b40bb0f
# ---
# + [markdown] id="s3iu_wRpS8xc"
#
# # **ESS 314: Lab 1**
#
# This is the first lab of ESS 314. The lab was designed by <NAME>, <NAME>, and modified by <NAME> (<EMAIL>) and <NAME> (<EMAIL>)
#
# + [markdown] id="Eds_9NmfSziJ"
# ## **Lab 1A: Setting up Your Workspace**
#
#
# This lab focuses on helping students set up their workspace so that they can access the relevant
# software packages necessary for future lab work. *You do not need to turn anything in for portion 1A of the lab.*
#
#
#
# ### **Computer Resources**
#
# 1. ***UW-JupyterHub***
#
# UW-IT set up a jupyterhub for our course. You can start one with the following link in your browser:
#
# https://rttl.axdd.s.uw.edu/2021-autumn-ess-314-a
#
# You will need UW ID credentials to get started. You can log in from the ESS Lab computers and from your laptop.
#
# You will be prompted to a Jupyter Notebook in Python. We thrive to use open-source python codes that should be available through our python environment or easy to install using simple commands.
#
#
#
# **Now, we will go through an example on how to get the hub ready**.
#
# * Log in to the server
# * To add files, you can click the upload button on the top and upload individual files and folters.
# * To add the git repos with the scripts and lab notebook, open a terminal window and type ``git clone "https://github.com/UW-geophysics-edu/ESS314-fall21.git"``. The entire updated github repository will be transfered to your environments.
# * The files that you add to the environment will be saved and left as is for future times.
# * To update the github repository. Open a terminal, move into the directory using the command ``cd ~/ESS314-fall21``, and type ``git pull``. This will update your directory with the updated version of the remote class github.
#
#
# 2. ***Google Colab***
#
# Google Colab is a free python notebook environment. It comes in with a built-in python environment. Once you open a notebook in Colab, click ``Connect`` on the top right, select ``Connect to a hosted runtime`` to launch your instance on the free Google Cloud Platform.
#
# To add packages, just use the following command everytime you log in, for instance:
#
# ``!pip install mypackage``
#
#
#
# 3. ***Your own Local Python***
#
# We created an conda environment in the class Github Page: https://github.com/UW-geophysics-edu/ess314-2021-image.
# Similar to the hub, open a terminal window. Move to the directory of choice and type:
#
# * ``git clone "https://github.com/UW-geophysics-edu/ESS314-fall21.git"``.
#
# * Move into the directory: ``cd ESS314-fall21``
# * Create the conda environment: ``conda env create -f environment.yml``
# * start the conda environment: ``conda activate ess314``
#
# Now you are ready to do some pythoning!
#
#
# + [markdown] id="vh4X66FzoBMh"
# ### **Getting Familiar with Shell scripting**
#
# Shell scripting allows you to navigate through your computer directories, files, and perform basic manipulation of text files. It can also allows you to run big simulations on large computers. It is extremely flexible!
#
# Start by opening a **Terminal** window. The basic commands you will need are:
#
# * ``pwd`` to see what directory you are in.
# * ``cd FOLDER`` to move to another directory
# * ``cd ../OTHER_FOLDER`` to move to another directory that is ahead of your current directory. This provides a *relative* path.
# * ``cd /home/user/whatever/your/path/is`` is you know the *absolute* path of the directory you want to go to.
# * ``ls `` it will list all of the files in the current directory
# * ``ls ./*``, exact same as above
# * ``ls ../*``, will list the files in the directory ahead of your current directory
# * ``mkdir NEW_DIRECTORY`` wil create a new directory
# * ``cp file to_new_directory`` will copy ``file`` to another directory
# * ``cp -r this_directory to_this_directory``, will copy a folder and everything within it into another directory.
#
#
# We recommend to take the SoftwareCarpentry classes if you want to learn more. They have sometimes workshop through UW/eScience, or you can follow their lessons on Shell scripting here:
# https://swcarpentry.github.io/shell-novice/
#
# + [markdown] id="3QuW6Vtl0yHI"
# ### **Getting Familiar with Git**
#
#
# Github is becoming a fundamental tool in STEM education and research. It allows you to keep track of code versions, without having to save a file per version. It also allows you to share your work, share your code, and work as a team very efficiently.
#
# Github has a web-platform: https://github.com/
#
# Our class Github is here: https://github.com/UW-geophysics-edu/ESS314-fall21
#
#
# Github toolbox is installed on the JupyterHub. You may already have it installed on your local machine (lapto/desktop), or you may have to install it. Follow the instructions here: https://github.com/git-guides/install-git
#
# A cheat sheet of commands is added on Canvas and on the github website.
#
# You can just download all courses labs and scripts through the Github website as the repository is public. If you wish to create your own repository for this class and upload your own codes:
# * create a Github account (I advise using your uwnet ID or an EASY identifier).
# * Follow guidelines from the cheat sheet to use command line to navigate through git.
#
#
# Further master your Git Skills by taking the basic lesson from the Software Carpentry: https://swcarpentry.github.io/git-novice/.
# + [markdown] id="HBb5NmMAVVTB"
# ## **Lab 1B: Introduction to Python**
#
# Python is an increasingly common scientific tool which we will be utilizing in this course. Lab 1B focuses on gettnig acquainted with Python basics.
#
# *Questions/Items for submission are shown in italics. For all questions, show the command you used to arrive at the answer.*
# + [markdown] id="mrapRBHdv_GC"
# **Setting Variables**
# + [markdown] id="7OJqYXMAw6aO"
#
# 2. Many basic commands are available in Python, including arithmetic functions. For example, type 2+2 into a code cell and press SHIFT+ENTER.
# + colab={"base_uri": "https://localhost:8080/"} id="phT_aoQzwNkJ" outputId="993f4bb8-e06a-4685-d3bb-0f49d268ea48"
2+2
# + [markdown] id="uQaUmjmrx2IE"
# 3. Arithmetic operators are + for addition, - for subtraction, * for multiplication, and / for division. Raising a value to a power is accomplished by using ** (e.g. 3**2).
# + [markdown] id="AykQtz0RyGIM"
# 4. Type a=2 into the next cell. This will assign the value 2 to the variable a, which will hold this value until it is reassigned. To reassign a, type a=3 in the next cell.
# + id="7iWOCVWAwP7O"
a = 2 # Make sure to add comments to your code as you go to ensure clarity
# + id="f_Fgzb4OwRgg"
a = 3 # Here you could remark on what the variable a represents in your code
# + [markdown] id="_EbXyR2_ya7J"
# 5. Assign a value to b. Now type c = a+b and Python will calculate the value of c. Note that Python will not print out the result of c automatically unless it is told to do so. To see the value of c, type print(c).
# + colab={"base_uri": "https://localhost:8080/"} id="J-RfeqiHwTXb" outputId="48f5dbff-31ce-4407-bb42-3acb8a011cd0"
b = 4
c = a + b
print(c)
# + [markdown] id="8PGTOjSdzBPL"
# 6. There are several kinds of variables in Python. Numbers can be integers, like a, whole numbers without a decimal point. Numbers designated as float have a decimal value. For computational purposes, float numbers are more useful. Python also allows the designation of text, or 'strings' as variables. Type d = 7.2 and e = 'hello!' to utilize float and string variables.
# + id="nscC4qLQwdh4"
d = 7.2
e = 'hello!'
# + [markdown] id="kqX093s90UTO"
# 7. To see all variables, their type, and their value, use the command %whos. Deleting a single variable can be accomplished using the command del following by the desired variable (e.g., del a). To delete all defined variables, use %reset. Check if the deletions were successful using %whos.
# + colab={"base_uri": "https://localhost:8080/"} id="XP40ACd1wrmw" outputId="d0db6b3a-783c-47ad-bb1c-12723175fb5c"
del a
# %whos
# + [markdown] id="RuVf09Cj1gkD"
# 8. It is important to note that variables in Python can consist of numbers, upper and lowercase letters, and underscores. A variable name cannot start with a number. Variable names are generally more useful when they are more descriptive (e.g., gravity_anomaly rather than ga).
# + [markdown] id="cIsn4c__1ZqA"
# ### **Question 1:**
#
#
#
# > ***a.*** What command would be used to clear just the b variable?*
#
#
#
#
# + [markdown] id="HQ8ujqVaqdz7"
# answer below:
#
# del b
# + [markdown] id="ykKOVC4BqgBu"
#
# > ***b.*** What command would be used to clear all variables?*
#
# + [markdown] id="rzMrLJKGqk5v"
# answer below:
# + [markdown] id="xsO0ToInqmRv"
#
# > ***c.*** Which of the following are valid variable names?*
# * *_n*
# * *n6*
# * *n_*
# * *6n*
# * *nnnnnnnnnnnnnnnnnnnnn*
#
# + [markdown] id="s8evhTwNqqxM"
# answer below:
#
# All except 6n
# + [markdown] id="MAR9vX9_qr8F"
# > ***d.*** *What error message appears when you try to use an invalid variable name?*
# + [markdown] id="Wwv6C0euqubg"
# answer below:
#
# "End of statement expected"
# + [markdown] id="-ZSLNN2h3Mhd"
# **Basic Commands and NumPy**
#
#
# + [markdown] id="uNeHDtu93Urm"
#
#
# 1. In order to perform more advanced calculations, certain Python packages must be imported. For example, numerical python is a key package. To import this, use the command
#
#
#
#
#
# + id="lLOD2oHM00jR"
import numpy as np
# + [markdown] id="yHQ2EtqO1GFd"
# 2. To use numpy, commands must begin with np. For example, type np.pi and you will see the result printed to the screen. To round this number to a certain number of decimal places, assign the result to a variable, then use the function round, for example:
#
# + colab={"base_uri": "https://localhost:8080/"} id="T3Fxxmbd1JE4" outputId="96aa5766-0f23-4351-bc56-fd3701de61ac"
pi = np.pi
round(pi, 2)
# + [markdown] id="05gLzG876W8U"
# 3. In numpy, log(x) calculates the natural log of a number. log10(x) will return the log base 10 and log2(x) will return the log base 2.
#
# + colab={"base_uri": "https://localhost:8080/"} id="Wm_KPly5ad90" outputId="f6c1b3c7-db46-49de-bbfc-9d097dfdbde6"
np.log10(10)
# + [markdown] id="S8BsnlP5aeIO"
#
# 4. The command np.sqrt() can be used to calculate the square root of positive numbers.
#
# + colab={"base_uri": "https://localhost:8080/"} id="iKBPQfXsafqb" outputId="4b589236-62d1-4ec2-db27-3fe51edc14cc"
np.sqrt(3)
# + [markdown] id="MzOKRTDpaf-s"
#
# 5. Trigonometric functions in NumPy take angles in radians, so make sure your input angles are not in degrees. You can also use the built-in NumPy conversion between degrees and radians.
# + colab={"base_uri": "https://localhost:8080/"} id="gXf_zX9hxSaB" outputId="918fac52-b79c-4d8e-f50b-23914e96a146"
print(np.sin(np.pi/2))
print(np.sin(np.deg2rad(90))) # You can also use np.rad2deg() for other calculations
# + [markdown] id="Gcjex5HpzD9f"
# 6. Inverse trigonometric functions can be calculated using np.arcsin(), np.arccos(), and np.arctan().
# + colab={"base_uri": "https://localhost:8080/"} id="oD2w9svEax9H" outputId="a9724add-43b7-43b2-b24f-b007a0677cc1"
print(np.arcsin(np.sin(np.pi/3)))
print(np.pi/3)
# + [markdown] id="fU8WEGBq698H"
# ### **Question 2:**
#
# > ***a.*** *What is the command to return the value of e (Euler’s number)?*
#
# + id="IqTkZWzlbRzp"
# answer below
np.e
# + [markdown] id="j5T3TeZfbT9R"
# > ***b.*** *What is the sin of 270 degrees?*
# + id="x6VuZxzhbV-J"
# answer below
np.sin(np.deg2rad(270))
# + [markdown] id="XMAssY4DbXO0"
# > ***c.*** *What is arccos(0.5) in degrees?*
# + id="AiJZOTCzbZO8"
#answer below
np.rad2deg(np.arccos(0.5))
# + [markdown] id="p_BIO5XQbaea"
# > ***d.*** *What is arccos(0.5) in radians?*
# + id="XrAqWIzhbcDf"
# answer below
np.arccos(0.5)
# + [markdown] id="E_xrFhN0qow-"
# **Vectors and Matrices**
#
# NumPy can be utilized to create "arrays," which function as vectors or matrices. The size of a matrix with r rows and c columns is 𝑟 × 𝑐. Think of a row vector as a matrix with 𝑟 = 1, and a column vector as a matrix with 𝑐 = 1.
#
#
#
# 1. Use the command np.array() to create row and column vectors as follows:
#
# + colab={"base_uri": "https://localhost:8080/"} id="2yr-0FFSznKH" outputId="49d45dfc-2854-4c77-c6b2-0bb800c43d38"
row_vector = np.array([[1, 2, 3]])
print(row_vector)
col_vector = np.array([[1], [2], [3]])
print(col_vector)
# You can check that these vectors are indeed the dimensions you seek by using the following command:
print(row_vector.shape) # Should be (1, 3) for (r, c)
print(col_vector.shape) # Should be (3, 1) for (r, c)
# + [markdown] id="HUZFU-PU0q4D"
# 2. Selecting specific values in a vector or matrix is known as indexing. Start by creating a 3 x 4 matrix as follows.
# + colab={"base_uri": "https://localhost:8080/"} id="cXSXVtWV0wzM" outputId="ef31d8c4-237a-4034-8dbc-b7ed662ba004"
matrix = np.array([[1, 2, 3, 4], [5, 6, 7, 8], [9, 10, 11, 12]])
print(matrix)
# + [markdown] id="8UusThFZ1EAu"
# To select elements within the matrix, specify their location within the rows and columns, making sure to begin counting at 0. If you are used to MATLAB, it is important to remember that Python indexing starts at 0, not 1. Here are some examples of indexing:
# + colab={"base_uri": "https://localhost:8080/"} id="I8HWL4DD0_E5" outputId="93f55769-3f2c-4d80-c9c6-3826ec39e390"
print(matrix[1,2]) # The number 7
print(matrix[2,0]) # The number 9
print(matrix[0, :]) # The first row
column1 = matrix[:, 0] # The first column - but note it only has one dimension
print(column1)
column1.shape # but note it only has one dimension and is not technically a 'row' or 'column'
matrixT = np.transpose(matrix) # Transpose a matrix using this command
print(matrixT)
# + [markdown] id="QtdzVeAE4Gia"
# 3. There are other ways to create more specialized kinds of arrays, such as those made out of zeroes, ones, or a linearly spaced array.
# + colab={"base_uri": "https://localhost:8080/"} id="W9QoCrSE1fUS" outputId="2fbc5961-4f31-46bb-d7be-d262abae944a"
print(np.zeros([3,1]))
print(np.ones([4,1]))
print(np.linspace(0,10,5)) # Specify start, stop, and the number of elements for a linearly spaced array
print(np.arange(0, 10, 2)) # Specify start, stop, and spacing
# Note the difference between the following:
print(np.linspace(0,10,10))
print(np.linspace(0,10,11))
# + [markdown] id="RkXjNvK-5ngd"
# 4. You can concatenate or truncate vectors once they are created. The two arrays must have the same shape, except in the direction of the axis.
# + colab={"base_uri": "https://localhost:8080/"} id="nOUQVnkL4glx" outputId="086d4d2e-e073-4ec9-ddb9-8daf1021caf1"
g = np.ones([4,3])
print(g)
h = np.zeros([2,3])
print(h)
i = np.concatenate((g,h)) # axis is assumed 0 unless specified
print(i)
j = np.ones([1,2])
print(j)
k = np.zeros([1, 3])
print(k)
l = np.concatenate((j, k), axis=1)
print(l)
m = np.concatenate((k, j), axis=1) # reverses the order from above
print(m)
# + id="t-zJqYl56xVc"
# Use these for Question 3
v = np.array([[1, 2, 3]])
w = np.array([[1], [2], [3]])
print(v)
print(w)
# + [markdown] id="K_hYsu6j8_zq"
# ### **Question 3:**
#
# > ***a.*** *What is the difference between v and w?*
#
#
# + [markdown] id="tcYZ2S7ptaBf"
# answer here
#
# V is a row matrix whereas W is a column matrix
# + [markdown] id="870vNDbStbP0"
#
# > ***b.*** *Explain the function of np.transpose().*
#
# + [markdown] id="DkfeagvNtc6n"
# answer here
#
# Turns the columns of the matrix into rows and vice versa
# + [markdown] id="_6Sqa18atfpc"
# > ***c.*** *Create a matrix with three columns, each of which is a copy of vector w*
# + id="4opEzIWftxsI"
#answer below
z = np.array([w, w, w]).transpose()
print(z)
# + [markdown] id="5mHMwC8_tizc"
# > ***d.*** *Select the entire third column of the matrix you just created?*
# + id="MI05b3K0t7Am"
#answer below
print(z[0, :, 2])
# + [markdown] id="-1Mn7wUf6Z_o"
# **Operations with Vectors and Matrices**
#
#
# + id="s2AFFYiy8ZUS"
v = np.array([[1, 2, 3]])
w = np.array([[1], [2], [3]])
p = ([[3, 4, 5, 6]])
M = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
N = np.concatenate((v, v, v))
# + [markdown] id="3ctDjZvEY1xv"
# 1. Adding a scalar to a vector or matrix is the same as adding two scalar values. Similarly,
# multiplying a vector or matrix by a scalar value looks just like multiplying two scalars.
# a. To add 2 to all elements of your vector v, type v+2.
# b. To multiply each element of matrix M by 3, type M*3.
# + colab={"base_uri": "https://localhost:8080/"} id="_VfcutaRYyRJ" outputId="3a0961f6-23dc-4911-e8b4-e2f508a8d3da"
print(v+2)
print(M*3)
# + [markdown] id="-r9KqlgJZoic"
# 2. Adding two matrices may work differently than you expect. Try adding v+N and p+N.
# + colab={"base_uri": "https://localhost:8080/"} id="hwXNNEamY8Xr" outputId="7676a703-9cab-4634-ff67-5ff9cad5c1a4"
v+N
# + colab={"base_uri": "https://localhost:8080/", "height": 164} id="d8KgYjbfZgob" outputId="97f5a497-97d0-4dc2-ae1a-8c77372eed94"
p+N
# + [markdown] id="J26dQxJcZ2ay"
# 3. To perform matrix multiplication, just multiply the two matrices by each other. Recall
# that to multiply two matrices together, the number of columns in the first must equal
# the number of rows in the second.
# a. Try multiplying vector v by vector w. Make sure and show that they have the complementary dimensions.
# + colab={"base_uri": "https://localhost:8080/"} id="JazWdY7hZlzT" outputId="e54557f0-e5f3-4ddf-c756-adde6b8da7f7"
print(v.shape)
print(w.shape)
v*w
# + [markdown] id="yMMTtfu-Z-Y7"
# 4. For square matrices, you can also raise the matrix to a power using the same function as
# you did with scalar values. Square the matrix M using the command M**2, then review
# the output.
# + colab={"base_uri": "https://localhost:8080/"} id="wRMWS2LKZ4TY" outputId="0ff2146b-b90a-4c05-86c5-2c4a2c6cfd90"
M**2
# + [markdown] id="xylu7k0maHnS"
# 5. Operations that apply to each element of the matrix individually are sometimes called array operations or element-wise operations. Many functions, such as trig functions, will operate on a matrix element-by-element. For example, try np.sin(N).
# + colab={"base_uri": "https://localhost:8080/"} id="s7ioHsu0aAA6" outputId="1ed04c41-9929-40ac-8191-3057dbc46132"
np.sin(N)
print(v)
print(M)
print(p)
print(v+M)
# print(p+M)
# + [markdown] id="o0n9c1GIao4o"
# *Question 4:*
#
#
#
# > ***a.*** *Why do you get an error message when you multiply v*v *but not when you multiply v*w?
#
#
# + [markdown] id="SLnya6I5MaQm"
#
# + [markdown] id="ekgAka6wMayh"
# > ***b.*** *What does Python do when you ask it to add v+M? What about p+M? Describe as best you can from your own experiments the logic that Python uses to add two matrices together. Include any relevant errors that might occur.*
#
# Python will add the values in V (1 row, 3 columns) to every row in V (3 rows, 3 columns). But because P has 4 columns,
# it is impossible for Python to add every value in p to M.
# + [markdown] id="v21pOH8Gbbqx"
# **Plotting**
#
#
#
# 1. Plotting in Python requires importing another package, known as matplotlib. You can find tutorials and self-teach yourself about plotting in Python on the Matplotlib tutorial page: https://matplotlib.org/stable/tutorials/index.html
#
#
# + id="P54mcCaIbdTs"
import matplotlib.pyplot as plt # You will have to run this import cell each time you open the notebook
# + [markdown] id="gtPwc_o3KS0I"
# 2. Create a row vector x with the numbers 1 through 50. Then, use the sine function to vector y where each element is the sine of the corresponding value in x.
# + id="b0zSVjFTKbY8"
x = np.linspace(1,50,50)
y = np.sin(x)
# + [markdown] id="nM_F0KXlL0tj"
# 3. Create a simple plot using matplotlib.
# + id="6yQseom3MZPy"
fig = plt.plot(x,y, 'g:') # 'g:' defines the color (g) and the pattern (:) of the lines here.
# You can search for color legends and more line patterns. - is a line, o is dots.
plt.xlabel('x label')
plt.ylabel('y label')
plt.title("Example Plot 1")
plt.show()
# + [markdown] id="kxxDgtI9SjmW"
# 4. Create a figure with 2 subplots.
# + id="NSJBr9BbKjU7"
z = np.cos(x)
# + id="Z-HI1ZlvLzrc" colab={"base_uri": "https://localhost:8080/", "height": 232} outputId="4861d135-0f4e-4b6b-c210-809e835b02ef"
fig, ax = plt.subplots(2,1) # control how many rows and columns here
ax[0].plot(x,y, 'r-', label='sin(x)')
ax[0].set_title('Example Plot 2a')
ax[0].set_xlabel('This is x (radians)')
ax[0].set_ylabel('Sine of x')
ax[0].legend()
ax[1].plot(x, z, 'bo', label='cos(x)')
ax[1].set_title('Example Plot 2b')
ax[1].set_xlabel('This is x (radians)')
ax[1].set_ylabel('Cosine of x')
ax[1].legend()
plt.tight_layout() # This will add space between your subplots (try commenting out this line!)
plt.show()
# Use the following command to save the image
plt.savefig('example2.png')
# + [markdown] id="ltDWL7hgSm1h"
# 5. Redefine x with twice as many points (i.e., define x as a row vector from 1 to 50, but include points at every half integer). Try to plot x and y again. What's wrong?
# + id="o9Tfi7F3N1fG"
x = np.linspace(1,50,99)
fig = plt.plot(x, y, 'b-')
# + [markdown] id="mOTW4yeLUiKK"
# ### **Question 5:**
#
# > ***a.*** *Redefine y as appropriate. Keep iterating on the resolution of x and y until you're satisfied that the graph looks like a sine wave. Add a title and axis labels. Export the plot image as a png named fig5. Open it from a file management app (Finder on Mac, Explorer on Windows) so you can be aware of the idiosyncrasies of exporting an
# image. Include the fig5.png export file with your assignment submission when you turn in this lab on Canvas.*
#
# + id="GQxfdxKfgY0H"
y=np.sin(x)
fig = plt.plot(x, y, 'b-')
plt.savefig('fig5.png')
# + [markdown] id="vWeTK7KsgZLy"
# > ***b.*** *Create a row vector from 0 – 100, counting by 5s (i.e. [0 5 10 15 20 … 100])?*
#
# + id="IKgwH8TvEsBx"
row_vector = np.linspace(0,100,21)
print(row_vector)
# + [markdown] id="kI1hV-wyEsZk"
# > ***c.*** *Plot x and y with maroon upward-facing triangles?*
# + id="HEZrgk_VJhqw"
fig10 = plt.plot(x, y, 'm^')
# + [markdown] id="vQeFKx1zG03P"
#
# > ***d.*** *What was the command you used to define x for your last plot?*
#
# x = np.linspace(1,50,99)
# + [markdown] id="9UfQf1OuUWrk"
# ### **Question 6:**
#
#
# > ***a***. *Plot two functions on the same plot. Define a new row vector x where the values range from 1 to 40.*
#
# + id="KwdubgqsLG-T"
x1 = np.linspace(1, 40, 40)
print(x1)
fig11 = plt.plot(x, y, 'm^')
fig11 = plt.plot(x1, x1, 'g-')
# + [markdown] id="PRzgEP3wLHYC"
# > ***b.*** *Define a vector y that is equal to x^2 + 50cos(x + 4). Please express the cosine in radians.*
#
# + id="POx3siCKLdot"
y1 = np.square(x1) + 50 * np.cos(x1 + 4)
print(y1)
# + [markdown] id="jx6Iih7gLdyK"
# > ***c.*** *Define a vector z that is equal to 1.5(tan(y)). Please express the tangent in radians.*
#
# + id="q_ZqQYg8LmS0"
z1 = 1.5 * np.tan(y1)
print(z1)
# + [markdown] id="uB0hZkjILmct"
# > ***d.*** *Plot y vs. x and z vs. x on the same graph. First, open a figure. Plot the y function as a blue solid line, and the z function as a red dotted line.*
# + id="KSaBion6Lp4i"
fig12 = plt.plot(y1, x1, 'b-')
fig12 = plt.plot(z1, x1, 'r.')
# + [markdown] id="ZAFQOtkdLqB5"
# > ***e.*** *Add a title, and label the x and y axes ‘x’ and 'y(x) and z(y(x))’, respectively.*
#
# + id="ZrttxIrJLsfD"
fig12 = plt.plot(y1, x1, 'b-')
fig12 = plt.plot(z1, x1, 'r.')
plt.title('Graph of y vs x and z vs x')
plt.xlabel('x')
plt.ylabel('y(x) and z(y(x))')
plt.savefig('fig6')
# + [markdown] id="h0wk59cgLsof"
# > ***f.*** *Plot more than one function on the same graph.*
# + id="WZ9y1sPDLuLv"
# + [markdown] id="rQ8t-qGwLuVO"
# > ***g.*** *Save the plot image as a png named fig6. You'll need to include this file in your submission for this lab.*
# + id="4TBLx2ctLue6"
|
LABS/LAB1/Lab1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.8 - AzureML
# language: python
# name: python38-azureml
# ---
# # AutoML: Train "the best" classifier model for the UCI Bank Marketing dataset.
#
# **Requirements** - In order to benefit from this tutorial, you will need:
# - A basic understanding of Machine Learning
# - An Azure account with an active subscription. [Create an account for free](https://azure.microsoft.com/free/?WT.mc_id=A261C142F)
# - An Azure ML workspace. [Check this notebook for creating a workspace](../../../resources/workspace/workspace.ipynb)
# - A Compute Cluster. [Check this notebook to create a compute cluster](../../../resources/compute/compute.ipynb)
# - A python environment
# - Installed Azure Machine Learning Python SDK v2 - [install instructions](../../../README.md) - check the getting started section
# - The latest MLFlow packages:
#
# pip install azureml-mlflow
#
# pip install mlflow
#
# **Learning Objectives** - By the end of this tutorial, you should be able to:
# - Connect to your AML workspace from the Python SDK
# - Create an `AutoML classification Job` with the 'classification()' factory-fuction.
# - Submit and run the AutoML classification job
# - Obtaing the model and score predictions with it
#
# **Motivations** - This notebook explains how to setup and run an AutoML classification job. This is one of the nine ML-tasks supported by AutoML. Other ML-tasks are 'regression', 'time-series forecasting', 'image classification', 'image object detection', 'nlp text classification', etc.
#
# In this example we use the UCI Bank Marketing dataset to showcase how you can use AutoML for a classification problem. The classification goal is to predict if the client will subscribe to a term deposit with the bank.
# # 1. Connect to Azure Machine Learning Workspace
#
# The [workspace](https://docs.microsoft.com/en-us/azure/machine-learning/concept-workspace) is the top-level resource for Azure Machine Learning, providing a centralized place to work with all the artifacts you create when you use Azure Machine Learning. In this section we will connect to the workspace in which the job will be run.
#
# ## 1.1. Import the required libraries
# + name="automl-import" gather={"logged": 1634852261599}
# Import required libraries
from azure.identity import DefaultAzureCredential
from azure.ai.ml import MLClient
from azure.ai.ml.constants import AssetTypes
from azure.ai.ml import automl
from azure.ai.ml import Input
# -
# ## 1.2. Workspace details
#
# To connect to a workspace, we need identifier parameters - a subscription, resource group and workspace name. We will use these details in the `MLClient` from `azure.ai.ml` to get a handle to the required Azure Machine Learning workspace. We use the default [default azure authentication](https://docs.microsoft.com/en-us/python/api/azure-identity/azure.identity.defaultazurecredential?view=azure-python) for this tutorial. Check the [configuration notebook](../../configuration.ipynb) for more details on how to configure credentials and connect to a workspace.
#
# By default, we try to use the by default workspace configuration (available out-of-the-box in Compute Instances) or from any Config.json file you might have copied into the folders structure.
# If no Config.json is found, then you need to manually introduce the subscription_id, resource_group and workspace when creating MLClient .
# + name="mlclient-setup" gather={"logged": 1634852261744} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
credential = DefaultAzureCredential()
ml_client = None
try:
ml_client = MLClient.from_config(credential)
except Exception as ex:
print(ex)
# Enter details of your AML workspace
subscription_id = "<SUBSCRIPTION_ID>"
resource_group = "<RESOURCE_GROUP>"
workspace = "<AML_WORKSPACE_NAME>"
ml_client = MLClient(credential, subscription_id, resource_group, workspace)
# -
# ### Show Azure ML Workspace information
# +
workspace = ml_client.workspaces.get(name=ml_client.workspace_name)
output = {}
output["Workspace"] = ml_client.workspace_name
output["Subscription ID"] = ml_client.connections._subscription_id
output["Resource Group"] = workspace.resource_group
output["Location"] = workspace.location
output
# -
# # 2. Configure and run the AutoML classification job
# In this section we will configure and run the AutoML classification job.
#
# ## 2.1 Configure the job through the classification() factory function
#
# ### classification() parameters:
#
# The `classification()` factory function allows user to configure AutoML for the classification task for the most common scenarios with the following properties.
#
# - `target_column_name` - The name of the column to target for predictions. It must always be specified. This parameter is applicable to 'training_data', 'validation_data' and 'test_data'.
# - `primary_metric` - The metric that AutoML will optimize for Classification model selection.
# - `training_data` - The data to be used for training. It should contain both training feature columns and a target column. Optionally, this data can be split for segregating a validation or test dataset.
# You can use a registered MLTable in the workspace using the format '<mltable_name>:<version>' OR you can use a local file or folder as a MLTable. For e.g Input(mltable='my_mltable:1') OR Input(mltable=MLTable(local_path="./data"))
# The parameter 'training_data' must always be provided.
# - `compute` - The compute on which the AutoML job will run. In this example we are using a compute called 'cpu-cluster' present in the workspace. You can replace it any other compute in the workspace.
# - `name` - The name of the Job/Run. This is an optional property. If not specified, a random name will be generated.
# - `experiment_name` - The name of the Experiment. An Experiment is like a folder with multiple runs in Azure ML Workspace that should be related to the same logical machine learning experiment.
#
# ### set_limits() parameters:
# This is an optional configuration method to configure limits parameters such as timeouts.
#
# - timeout_minutes - Maximum amount of time in minutes that the whole AutoML job can take before the job terminates. This timeout includes setup, featurization and training runs but does not include the ensembling and model explainability runs at the end of the process since those actions need to happen once all the trials (children jobs) are done. If not specified, the default job's total timeout is 6 days (8,640 minutes). To specify a timeout less than or equal to 1 hour (60 minutes), make sure your dataset's size is not greater than 10,000,000 (rows times column) or an error results.
#
# - trial_timeout_minutes - Maximum time in minutes that each trial (child job) can run for before it terminates. If not specified, a value of 1 month or 43200 minutes is used.
#
# - max_trials - The maximum number of trials/runs each with a different combination of algorithm and hyperparameters to try during an AutoML job. If not specified, the default is 1000 trials. If using 'enable_early_termination' the number of trials used can be smaller.
#
# - max_concurrent_trials - Represents the maximum number of trials (children jobs) that would be executed in parallel. It's a good practice to match this number with the number of nodes your cluster.
#
# - enable_early_termination - Whether to enable early termination if the score is not improving in the short term.
#
# + name="data-load"
# Create MLTables for training dataset
my_training_data_input = Input(
type=AssetTypes.MLTABLE, path="./data/training-mltable-folder"
)
# Remote MLTable definition
# my_training_data_input = Input(type=AssetTypes.MLTABLE, path="azureml://datastores/workspaceblobstore/paths/Classification/Train")
# + tags=["parameters"]
# General job parameters
compute_name = "cpu-cluster"
max_trials = 5
exp_name = "dpv2-classifier-experiment"
# + name="classification-configuration" gather={"logged": 1634852262026} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Create the AutoML classification job with the related factory-function.
classification_job = automl.classification(
compute=compute_name,
experiment_name=exp_name,
training_data=my_training_data_input,
target_column_name="y",
primary_metric="accuracy",
n_cross_validations=5,
enable_model_explainability=True,
tags={"my_custom_tag": "My custom value"},
)
# Limits are all optional
classification_job.set_limits(
timeout_minutes=600,
trial_timeout_minutes=20,
max_trials=max_trials,
# max_concurrent_trials = 4,
# max_cores_per_trial: -1,
enable_early_termination=True,
)
# Training properties are optional
classification_job.set_training(
blocked_training_algorithms=["LogisticRegression"],
enable_onnx_compatible_models=True,
)
# -
# ## 2.2 Run the Command
# Using the `MLClient` created earlier, we will now run this Command in the workspace.
# + name="job-submit" gather={"logged": 1634852267930} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Submit the AutoML job (CDLTLL: Is it ml_client.create_or_update(classification_job))
returned_job = ml_client.jobs.create_or_update(
classification_job
) # submit the job to the backend
print(f"Created job: {returned_job}")
# Get a URL for the status of the job
# returned_job.services["Studio"].endpoint
# -
# ### Wait until the AutoML job is finished
# ml_client.jobs.stream(returned_job.name) waits until the specified job is finished
ml_client.jobs.stream(returned_job.name)
print(returned_job.name)
# # Next Steps
# You can see further examples of other AutoML tasks such as Image-Classification, Image-Object-Detection, NLP-Text-Classification, Time-Series-Forcasting, etc.
#
|
sdk/jobs/automl-standalone-jobs/automl-classification-task-bankmarketing/automl-classification-task-bankmarketing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Usage
#
# This tutorial walks through the basic usage of the `pynrc` package to calculate sensitivities and saturation limits for NIRCam in a variety of modes.
# +
# Makes print and division act like Python 3
from __future__ import print_function, division
# Import the usual libraries
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# Enable inline plotting at lower left
# %matplotlib inline
from IPython.display import display, Latex, clear_output
# -
# ## Getting Started
#
# We assume you have already installed `pynrc` as outlined in the documentation.
# +
# import main module
import pynrc
from pynrc import nrc_utils
# import pysynphot instance
#from pynrc.nrc_utils import S
# -
# Log messages for `pynrc` follow the same the logging functionality included in `webbpsf`. Logging levels include `DEBUG`, `INFO`, `WARN`, and `ERROR`.
pynrc.setup_logging()
# If you get tired of the `INFO` level messages, simply type:
# ```python
# pynrc.setup_logging('WARN', verbose=False)
# ```
# ## First NIRCam Observation
#
# The basic NIRCam object consists of all the instrument settings one would specify for a JWST observation, including filter, pupil, and coronagraphic mask selections along with detector subarray settings and ramp sampling cadence (i.e., `MULTIACCUM`).
#
# The NIRCam class makes use of high order polynomial coefficient maps to quickly generate large numbers of monochromatic PSFs that can be convolved with arbitrary spectra and collapsed into a final broadband PSF (or dispersed with NIRCam's slitless grisms). The PSF coefficients are calculated from a series of WebbPSF monochromatic PSFs and saved to disk. These polynomial coefficients are further modifed based on focal plane position and drift in the wavefront error relative to nominal OPD mao.
#
# There are a multitude of posssible keywords one can pass upon initialization, including certain detector settings and PSF generation parameters. If not passed initially, then defaults are assumed. The user can update these parameters at any time by either setting attributes directly (e.g., `filter`, `mask`, `pupil`, etc.) along with using the `update_detectors()` and `update_psf_coeff()` methods.
#
# For instance,
# ```python
# nrc = pynrc.NIRCam('F210M')
# nrc.module = 'B'
# nrc.update_detectors(read_mode='DEEP8', nint=10, ngroup=5)
# ```
# is the same as:
# ```python
# nrc = pynrc.NIRCam('F210M', module='B', read_mode='DEEP8', nint=10, ngroup=5)
# ```
#
# To start, we'll set up a simple observation using the `F430M` filter. Defaults will be populated for unspecified attributes such as `module`, `pupil`, `mask`, etc.
#
# **Check the function docstrings for more detailed information**
nrc = pynrc.NIRCam(filter='F430M')
print('Filter: {}; Pupil: {}; Mask: {}; Module: {}'\
.format(nrc.filter, nrc.pupil, nrc.mask, nrc.module))
# Keyword information for detector and PSF settings are stored in the `det_info` and `psf_info` dictionaries. These cannot be modified directly, but instead are updated via the `update_detectors()` and `update_psf_coeff()` methods.
print('Detector Info Keywords:')
print(nrc.det_info)
print('')
print('PSF Info Keywords:')
print(nrc.psf_info)
# PSF coefficient information is stored in the `psf_coeff` attribute. This data is accessed by many of the NIRCam class functions to generate PSFs with arbitrary wavelength weights, such as the `gen_psf()` function.
# +
# Demonstrate the color difference of the PSF for different spectral types, same magnitude
sp_M0V = pynrc.stellar_spectrum('M0V', 10, 'vegamag', nrc.bandpass)
sp_A0V = pynrc.stellar_spectrum('A0V', 10, 'vegamag', nrc.bandpass)
# Generate oversampled PSFs (counts/sec)
_, psf_M0V = nrc.gen_psf(sp_M0V, return_oversample=True)
_, psf_A0V = nrc.gen_psf(sp_A0V, return_oversample=True)
fig, axes = plt.subplots(1,3, figsize=(12,4))
axes[0].imshow(psf_M0V**0.5)
axes[0].set_title('M0V PSF ({})'.format(nrc.filter))
axes[1].imshow(psf_A0V**0.5)
axes[1].set_title('A0V PSF ({})'.format(nrc.filter))
diff = psf_M0V - psf_A0V
minmax = np.abs(diff).max() / 2
axes[2].imshow(diff, cmap='RdBu', vmin=-minmax, vmax=minmax)
axes[2].set_title('Difference')
fig.tight_layout()
# -
# Bandpass information is stored in the `bandpass` attribute and can be plotted with the convenience function `plot_bandpass()`.
nrc.plot_bandpass()
# ##1. Saturation Limits
#
# One of the most basic functions is to determine the saturation limit of a CDS observation, so let's try this for the current filter selection. Generally, saturation is considered to be 80% of the full well.
# +
# Turn off those pesky informational texts
pynrc.setup_logging('WARN', verbose=False)
# Configure the observation for CDS frames (ngroup=2)
# Print out frame and ramp information using verbose=True
nrc.update_detectors(ngroup=2, verbose=True)
# -
# The `sat_limits()` function returns a dictionary of results. There's the option in include a Pysynphot spectrum, but if none is specificed the it defaults to a G2V star.
# +
# Set verbose=True to print results in a user-friendly manner
sat_lims = nrc.sat_limits(verbose=True)
# Dictionary information
print("\nDictionary Info:", sat_lims)
# -
# By default, the function `sat_limits()` uses a G2V stellar spectrum, but any arbritrary spectrum can be passed via the `sp` keyword. In addition, using the `bp_lim` keyword, you can use spectral information to determine the brightness in some other bandpass that saturates the source within the NIRCam filter.
# +
# Spectrum of an M0V star (not normalized)
sp_M0V = pynrc.stellar_spectrum('M0V')
# 2MASS Ks Bandpass
bp_k = pynrc.bp_2mass('K')
sat_lims = nrc.sat_limits(sp=sp_M0V, bp_lim=bp_k, verbose=True)
# -
# Now, let's get the same saturation limit assuming a 128x128 subarray.
nrc.update_detectors(wind_mode='WINDOW', xpix=128, ypix=128)
sat_lims = nrc.sat_limits(sp=sp_M0V, bp_lim=bp_k, verbose=True)
# You can also use the `saturation_levels()` function to generate an image of a point source indicating the fractional well fill level.
# +
# Spectum of A0V star with Ks = 8 mag
sp = pynrc.stellar_spectrum('M0V', 8, 'vegamag', bp_k)
sat_levels = nrc.saturation_levels(sp, full_size=False, ngroup=nrc.det_info['ngroup'])
print('Max Well Fraction: {:.2f}'.format(sat_levels.max()))
# +
# Plot the well fill levels for each pixel
fig, ax = plt.subplots(1,1)
extent = 0.5*nrc.psf_info['fov_pix'] * np.array([-1,1,-1,1])
cax = ax.imshow(sat_levels, extent=extent, vmin=0, vmax=1)
ax.set_xlabel('Pixels')
ax.set_ylabel('Pixels')
ax.set_title('Well Fraction in {} of $K_s = 5$ M0V star'.format(nrc.filter))
cbar = fig.colorbar(cax)
cbar.set_label('Well Fill Fraction')
ax.tick_params(axis='both', color='white', which='both')
for k in ax.spines.keys():
ax.spines[k].set_color('white')
# -
# Information for slitless grism observations show wavelength-dependent results.
nrc = pynrc.NIRCam('F444W', pupil='GRISM0', ngroup=2, wind_mode='STRIPE', ypix=128)
sat_lims = nrc.sat_limits(sp=sp_M0V, bp_lim=bp_k, verbose=True)
# ## 2. Sensitivity Limits
#
# Similarly, we can determine sensitivity limits of point sources (and extended sources) for the defined instrument configuration. By default, the `sensitivity()` function uses a flat spectrum. In this case, let's find the sensitivities NIRCam can reach in a single ~1000sec integration with the F430M filter. Noise values will depend on the exact `MULTIACCUM` settings.
# +
nrc = pynrc.NIRCam('F430M')
nrc.update_detectors(read_mode='MEDIUM8', ngroup=10)
# The multiaccum_times attribute describes the various timing information
print(nrc.multiaccum_times)
# -
sens = nrc.sensitivity(nsig=5, units='vegamag', verbose=True)
# The sensitivity function also includes a keyword `forwardSNR`, which allows the user to pass a normalized spectrum and estimate the SNR For some extraction aperture.
sp = pynrc.stellar_spectrum('M0V', 20, 'vegamag', nrc.bandpass)
snr = nrc.sensitivity(sp=sp, forwardSNR=True, units='vegamag', verbose=True)
# ## 3. Ramp Optimization
#
# Armed with these two basic functions, we can attempt to determine the best instrument settings to optimize for SNR and efficiency. In these types of optimizations, we must consider observational constraints such as saturation levels, SNR requirements, and limits on acquisition time.
#
# **Note**: The reported acquisition times do not include obsevatory and instrument-level overheads, such as slew times, filter changes, script compilations, etc. It only includes detector readout times (including reset frames and Fast Row Resets).
#
# For instance, we want to observe an M-Dwarf (K=18 mag) in the F430M filter. What is the most efficient configuration to obtain an SNR of 100?
# +
# Setup observation
nrc = pynrc.NIRCam('F430M', wind_mode='WINDOW', xpix=160, ypix=160)
# Spectrum of an M2V star
bp_k = pynrc.bp_2mass('K')
sp_M0V = pynrc.stellar_spectrum('M2V', 18, 'vegamag', bp_k)
# -
# Run optimizer. Result is a ranked list sorted by efficiency.
tbl = nrc.ramp_optimize(sp_M0V, snr_goal=100, ng_min=5, nint_min=10, verbose=True)
# For a slightly more complicated scenario, consider an additional foreground source. In this scenario, the F0V star will saturate much more quickly compared to the fainter M2V, so it limits which ramp settings we may want to use (assuming we want unsaturated frames, which isn't always necessarily true).
sp_F0V = pynrc.stellar_spectrum('F0V', 10, 'vegamag', bp_k)
tbl = nrc.ramp_optimize(sp_M0V, sp_bright=sp_F0V,
snr_goal=100, ng_min=5, nint_min=10, verbose=True)
# If there are no objections to saturating the bright source, then we can set the `well_frac_max` parameter to something like 5 times the hard saturation limit. This allows for more efficient exposure settings.
tbl = nrc.ramp_optimize(sp_M0V, sp_bright=sp_F0V, well_frac_max=5,
snr_goal=100, ng_min=5, nint_min=10, verbose=True)
|
notebooks/Basic_Usage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: forallpurposes
# language: python
# name: forallpurposes
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import copy
from altaipony.ffd import FFD
import time
tstamp = time.strftime("%d_%m_%Y_%H_%M", time.localtime())
for att in ['axes.labelsize', 'axes.titlesize', 'legend.fontsize',
'legend.fontsize', 'xtick.labelsize', 'ytick.labelsize']:
plt.rcParams[att] = 13
import logging
logger = logging.getLogger()
logger.setLevel(logging.CRITICAL)
# -
ccol = dict(zip(['pleiades',"hyades","praesepe","ngc6774","ngc2682"],
[["Pleiades","brown","x",135,25,25],
["Hyades","orangered","*",690,160,100],
["Praesepe","orange","d",750,3,7],
["Rup 147", "green",r"$m$",2650,380,380],
["M67","blue",">",3639,17,17]]))
# +
df = pd.read_csv("../k2scoc/results/tables/full_table.csv")
hasflares = (df.real==1) & (df.todrop.isnull())
wassearched = (df.real==0) & (df.todrop.isnull())
teffrange = (df.Teff_median<=6000) & (df.Teff_median>=2500)
df = df[(hasflares | wassearched) & teffrange]
df.shape
# +
df["LC_time_years"] = df.dtp_tot / 2. / 24. / 365.25
df = df[(df.Teff_median<=6000) & (df.Teff_median>=2500)]
cepic = df[["LC_time_years","EPIC","C", "Teff_median", "cluster"]].drop_duplicates()
tobs = cepic.LC_time_years.sum()
print(f"Total observing time: {tobs:.2f} years!")
labels = ["2500-3000","3000-3250","3250-3500",
"3500-3750","3750-4000","4000-5000",
"5000-6000"]
Mmid = [0.097, .19, .36, .47, .55, .71, .94]
masses = pd.DataFrame({"Teff_median" : labels,
"M_Msun" : Mmid})
params = pd.read_csv("cluster_parameters_merged.csv")
bins_T = pd.cut(cepic["Teff_median"],
bins=[2500,3000,3250,3500,3750,4000,5000,6000],
labels=labels)
T = cepic.groupby([bins_T, "cluster"]).LC_time_years.sum().reset_index()
T = T.merge(masses, how="left", on="Teff_median")
T = T.merge(params[["cluster","age (Myr)"]], how="left", on="cluster")
assert T.LC_time_years.sum() - tobs < 1e-10
T.shape
# +
def gyroflaremodel(t, m):
a1, a2, a3 = -.07, .79, -1.06
b1, b2, b3 = 2.01, -25.15, 33.99
alpha = a1 * np.log10(t) + a2 * m + a3
beta = b1 * np.log10(t) + b2 * m + b3
return np.array([alpha, beta, t, m])
s = T.apply(lambda x: gyroflaremodel(x["age (Myr)"], x["M_Msun"]), axis=1)
s = pd.DataFrame(s.tolist(), columns=['alpha', 'beta', "age (Myr)", "M_Msun"])
T = T.merge(s, how="left", on=[ "age (Myr)", "M_Msun"])
T.shape
# -
gyroflaremodel(135, 0.5)
# +
fig, ax = plt.subplots(1, figsize=(8,6))
df2 = copy.deepcopy(df)
df2["ed_rec"] = df.ed_rec * df.Lum_Kepler
ffd = FFD(f=df2[hasflares], tot_obs_time=tobs, ID="EPIC")
#---------- No correction ----------------
ed, freq, counts = ffd.ed_and_freq(energy_correction=False,
recovery_probability_correction=False,
multiple_stars=False)
ax.scatter(ed, freq, c="orange", marker="*", label="no correction")
#---------- multiple stars
ed, freq, counts = ffd.ed_and_freq(energy_correction=False,
recovery_probability_correction=False,
multiple_stars=True)
#ax.scatter(ffd.ed, ffd.freq, marker="^", c="c", label="correcting for multiple stars in sample")
cts = pd.Series(counts).value_counts()
thresh = cts[cts==1].sort_index().index.min()
ed, freq, counts = ffd.ed_and_freq(energy_correction=False,
recovery_probability_correction=False,
multiple_stars=False)
ffd.count_ed = ffd.count_ed[np.where(ffd.count_ed>thresh)]
ffd.ed = ed[np.where(ed>thresh)]
ffd.freq = freq[np.where(ed>thresh)]
alpha_en, alpha_en_err = ffd.fit_powerlaw()
print(f"Fitted power law alpha={ffd.alpha}")
#ffd.alpha, ffd.alpha_err = 2., .2
betas, beta, beta_err = ffd.fit_beta_to_powerlaw(mode="energy")
print(f"Fitted power law beta={ffd.beta}")
ispowerlaw = ffd.is_powerlaw()
truncated = ffd.is_powerlaw_truncated()
ax.scatter(ffd.ed, ffd.freq, marker="^", c="grey", label="all flaring stars contribute = above detection threshold")
a, x, y = ffd.plot_powerlaw(ax, c="k", label=fr"$\alpha=-${ffd.alpha:.2f}({ffd.alpha_err:.2f}), $\beta=${ffd.beta:.2e}")
# resfullsample["alpha_en"] = alpha_en
# resfullsample["alpha_en_err"] = alpha_en_err
# resfullsample["beta_en"] = beta
# resfullsample["beta_en_err"] = beta_err
# resfullsample["nflares_en_tot"] = len(ed)
# resfullsample["nflares_en_for_fit"] = len(ffd.ed)
# resfullsample["ispowerlaw_en"] = ispowerlaw
# resfullsample["istruncated_en"] = truncated
plt.xscale("log")
plt.yscale("log")
plt.legend(frameon=False)
plt.xlabel(r"$E_\mathrm{Kp}$ [erg]")
plt.ylabel("cumulative number of flares per year")
#plt.savefig(f"plots/{tstamp}_full_sample_ffd_energy.png", dpi=300)
# -
# ## Teff bins
#
# +
resen ={}
bins_T = pd.cut(cepic["Teff_median"],
bins=[2500,3000,3250,3500,3750,4000,5000,6000],
labels=labels)
df2 = copy.deepcopy(df)
df2["ed_rec"] = df.ed_rec * df.Lum_Kepler
df2 = df2[hasflares]
bins_T = pd.cut(df2["Teff_median"],
bins=[2500,3000,3250,3500,3750,4000,5000,6000],
labels=labels)
print(df2.shape)
fig, axes = plt.subplots(nrows=4,ncols=2,figsize=(12.5,18.5),sharey=True, squeeze=True)
k = 0
axes = [l for x in axes for l in x]
for i, g in df2.groupby(bins_T):
resen[i]={}
g.loc[:,"cluster"] = pd.Categorical(g.cluster,
categories=["pleiades","hyades","praesepe","ngc6774", "ngc2682"],
ordered=True)
for j, h in g.groupby("cluster"):
# print(i,j)
# print(h.shape)
tobs = T[(T.Teff_median == i) & (T.cluster == j)].LC_time_years.values[0]
h = h[(h.real == 1)]
if h.shape[0] == 0:
continue
else:
ffd = FFD(f=h, tot_obs_time=tobs, ID="EPIC")
#---------- get thershold
ed, freq, counts = ffd.ed_and_freq(energy_correction=False,
recovery_probability_correction=False,
multiple_stars=True)
cts = pd.Series(counts).value_counts()
thresh = cts[cts==1].sort_index().index.min()
# ----------- use threshold
ed, freq, counts = ffd.ed_and_freq(energy_correction=False,
recovery_probability_correction=False,
multiple_stars=False)
ffd.count_ed = ffd.count_ed[np.where(ffd.count_ed>=thresh)]
ffd.ed = ed[np.where(ed>=thresh)]
ffd.freq = freq[np.where(ed>=thresh)]
ffd.alpha, ffd.alpha_err = alpha_en, alpha_en_err
betas, beta, beta_err = ffd.fit_beta_to_powerlaw(mode="energy")
#print(f"Fitted power law beta={ffd.beta}")
if j == "ngc2682":
E, y = ffd.ed[0], ffd.freq[0]
ffd.beta = y * np.power(E, ffd.alpha-1)
a, x, y = ffd.plot_powerlaw(axes[k], c="k",)
else:
a, x, y = ffd.plot_powerlaw(axes[k], c="k",)
ab = T[(T.cluster==j) & (T.Teff_median==i)]
A = 1. - ab.alpha.iloc[0]
B = np.power(10, ab.beta.iloc[0]) * (1 - A) * 365.25
ffdd = FFD(alpha=A, beta=B)
ffdd.ed = ffd.ed
a, x, y = ffdd.plot_powerlaw(axes[k], c=ccol[j][1], linewidth=3)#, label=f"{A}, {B}")
resen[i][ccol[j][0]]={"beta_en":ffd.beta,
"beta_en_err":ffd.beta_err,
"nflares_en":len(ed),
"age":ccol[j][3],
"age_uperr":ccol[j][4],
"age_lowerr":ccol[j][5],
"ispowerlaw_en":ffd.is_powerlaw(),
"istruncated_en": ffd.is_powerlaw_truncated()
}
ed, freq, counts = ffd.ed_and_freq(energy_correction=False,
recovery_probability_correction=False,
multiple_stars=False)
axes[k].scatter(ed, freq, c=ccol[j][1], marker=ccol[j][2], s=60,
label=fr"{ccol[j][0]}: $\beta=${ffd.beta:.2e}")
axes[k].set_xscale('log')
axes[k].set_yscale('log')
axes[k].set_title(f"{i} K")
axes[k].set_xlim(8e30, 5e35)
axes[k].set_xlabel(r"$E_\mathrm{Kp}$ [erg]")
axes[k].legend()
k += 1
axes[-1].axis('off')
for i in [0,2,4,6]:
axes[i].set_ylabel(r"cumulative flare frequency [yr$^{-1}$]")
plt.tight_layout()
#plt.savefig(f"plots/{tstamp}_SpT_wise_sample_ffd_energy.png", dpi=300)
# +
reform = {(outerKey, innerKey): values for outerKey, innerDict in resed.items() for innerKey, values in innerDict.items()}
betaed_ = pd.DataFrame(reform).T
betaed = betaed_.reset_index().rename(index=str, columns={"level_0":"Teff",
"level_1":"cluster"})
reform = {(outerKey, innerKey): values for outerKey, innerDict in resen.items() for innerKey, values in innerDict.items()}
betaen_ = pd.DataFrame(reform).T
betaen = betaen_.reset_index().rename(index=str, columns={"level_0":"Teff",
"level_1":"cluster"})
beta = betaed.merge(betaen)
beta["Tmin"] = beta.Teff.apply(lambda x: int(x.split("-")[0]))
beta["Tmax"] = beta.Teff.apply(lambda x: int(x.split("-")[1]))
beta["Tmid"] = ((beta.Tmax + beta.Tmin) / 2).astype(int)
beta["Tmid_err"] = ((beta.Tmax - beta.Tmin) / 2).astype(int)
beta.sort_values(by="nflares")
# +
fig, ax = plt.subplots(figsize=(10,5))
Tcols = {"2750":["^","maroon"],
"3125":[">","orangered"],
"3375":["<","orange"],
"3625":["*","green"],
"3875":["d","lime"],
"4500":["x","c"],
"5500":["o","blue"],}
for label, g in beta.groupby("Tmid"):
g.loc[:,"cluster"] = pd.Categorical(g.cluster,
categories=["Pleiades","Hyades","Praesepe","Rup 147", "M67"],
ordered=True)
g = g.sort_values(by="cluster")
g.plot(x="age", xerr="age_uperr",y="beta",yerr="beta_err",
label=f"{g.Tmin.iloc[0]}-{g.Tmax.iloc[0]} K", ax=ax, kind="line",c=Tcols[str(label)][1],
marker=Tcols[str(label)][0],markersize=10)#s=120,)#
ax.set_xlabel("age [Myr]")
ax.set_ylabel(r"$\beta_\mathrm{s}$ [yr$^{-1}$]")
ax.set_xscale("log")
ax.set_yscale("log")
ax.set_xticks(beta.age.astype(float).unique())
ax.set_xticklabels(labels=beta.age.unique(), rotation=90)
ax.legend(loc=(1.03,.5))
plt.tight_layout()
plt.savefig(f"plots/{tstamp}_beta_T_age_ED.png", dpi=300)
# +
beta_s = betaen_.merge(betaed_, left_index=True,right_index=True)
for b in ["beta","beta_err","beta_en","beta_en_err"]:
beta_s.loc[:,b] = beta_s[b].astype(float)
for out, beta, beta_err in [(r"$\beta_\mathrm{s}$","beta","beta_err"),
(r"$\beta_\mathrm{erg}$","beta_en","beta_en_err")
]:
beta_s["perr"] = (np.rint(np.log10(beta_s[beta]/beta_s[beta_err]))+1).astype(int)
beta_s["b"] = (np.rint(np.log10(beta_s[beta]))).astype(int)
beta_s["berr"] = (np.rint(np.log10(beta_s[beta_err]))).astype(int)
beta_s["B"] = beta_s.apply(lambda x: np.round(x[beta]/10**x.b, x.perr).astype(str), axis=1)
beta_s["Berr"] = beta_s.apply(lambda x: np.round(x[beta_err]/10**x.b, x.perr), axis=1)
beta_s[out] = beta_s.apply(lambda x: f"${x.B}\left({x.Berr}\right)\cdot 10^{x.b}$", axis=1)
beta_s[out] = beta_s[out].apply(lambda x: "$" + x.replace("^","^{").replace("_","}_{").replace("$","}")[1:] + "$")
rename = {"nflares":r"$n_\mathrm{s}$",
"nflares_en":r"$n_\mathrm{erg}$",
"istruncated":r"$tr_\mathrm{s}$",
"istruncated_en":r"$tr_\mathrm{erg}$",
"ispowerlaw":r"$pl_\mathrm{s}$",
"ispowerlaw_en":r"$pl_\mathrm{erg}$",}
beta_s = beta_s[[r"$\beta_\mathrm{s}$","nflares","istruncated","ispowerlaw",
r"$\beta_\mathrm{erg}$","nflares_en","istruncated_en","ispowerlaw_en",]].rename(index=str, columns=rename)
beta_s = beta_s.sort_index()
nc = 'c' * (beta_s.shape[1])
stri = beta_s.to_latex(index=True,escape=False, column_format=f"l{nc}r")
stri = stri.replace("\\toprule","\hline")
stri = stri.replace("\\midrule","\hline")
stri = stri.replace("\\bottomrule","\hline\n" )
with open(f"/home/ekaterina/Documents/002_writing/flares-in-clusters-ii-draft/Flares_in_Clusters_with_K2_2/tables/powerlaw_SpT.tex", "w") as f:
f.write(stri)
# -
print(stri)
# # Notes on very old clusters
# ## Rup 147
#
# - 219426848 spectroscopic binary with double line (Curtis 2013)
# - **219591752** unremarkable Gaia target [simbad](http://simbad.u-strasbg.fr/simbad/sim-coo?Coord=19+14+11.962+%09-16+21+38.93+%09&CooFrame=FK5&CooEpoch=2000&CooEqui=2000&CooDefinedFrames=none&Radius=2&Radius.unit=arcsec&submit=submit+query&CoordList=) the cool star! 3315 K
# - 219341906 - nothing special from Curtis 2013
# - **219601739** - nothing special from Curtis 2013, CWW79
# - **219610232** - unremarkable Gaia target [simbad](http://simbad.u-strasbg.fr/simbad/sim-coo?Coord=19+13+31.091+%09-16+18+40.14+%09&CooFrame=FK5&CooEpoch=2000&CooEqui=2000&CooDefinedFrames=none&Radius=2&Radius.unit=arcsec&submit=submit+query&CoordList=), Cantat-Gaudin+ 2018: 100% member
# +
df[(df.cluster=="ngc6774") & (df.Teff_median > 3000.)].dropna(subset=["real"])[["dtp_tot", "EPIC","C","Teff_median", "ed_rec","note", "todrop"]]
# -
# ## M67
#
# - 211406144 - RG
# - 211409376 - RS CVn
# - **211434440** - ? 5000-6000 (no obvious info on [simbad](http://simbad.u-strasbg.fr/simbad/sim-coo?Coord=08+48+55.901+%09%2B12+09+42.93+%09&CooFrame=FK5&CooEpoch=2000&CooEqui=2000&CooDefinedFrames=none&Radius=2&Radius.unit=arcsec&submit=submit+query&CoordList=))
# - Huber+ 2016:
# ```
# lo [F
# Tef logg ([c Fe/H] ( Rad Mass E(B-V F RA DE
# EPIC f (K) m/s2]) [Sun]) (Rsun) (Msun) rho (Sun) Dist (pc) ) (mag) lag J2000 (deg) J2000 (deg) A15 2M
# --------- ----- ------ ------ ------- ------ --------- --------- ------- --- ----------- ----------- --- --
# 211434440 5236 4.545 -0.210 0.823 0.870 1.53e+00 6.20e+02 0.0338 rpm 132.2329270 +12.1619250 0 2M
# ```
# If the flare is real: 6.3e+33 erg energy released!
# - 211412587 - ? 5000-6000 spectroscopic binary [simbad](http://simbad.u-strasbg.fr/simbad/sim-coo?Coord=08+51+23.778+%09%2B11+49+49.38+%09&CooFrame=FK5&CooEpoch=2000&CooEqui=2000&CooDefinedFrames=none&Radius=2&Radius.unit=arcsec&submit=submit+query&CoordList=)
# - 211417284 - ? 5000-6000 cataclysmic variable star [simbad](http://simbad.u-strasbg.fr/simbad/sim-coo?Coord=08+50+36.996+%09%2B11+54+04.67+%09&CooFrame=FK5&CooEpoch=2000&CooEqui=2000&CooDefinedFrames=none&Radius=2&Radius.unit=arcsec&submit=submit+query&CoordList=), (0.64 day period, star 01610 in Nardiello+ 2016), [Mooley and Singh 2015](https://academic.oup.com/mnras/article/452/4/3394/1055661) :
# ```
# _Belloni et al. (1998)useB−V=0.38 from Sanders (1977) and hardness of the X-rayspectrum to claim that this could be an accreting white dwarf system. Orbital parameters areunknown. However, EIS hasB−V=0.79, and our hardness ratios are HR1=0.66±0.20,HR2=−0.48±0.30. The position in the HR diagram and absence of soft and hard X-raycomponents therefore argue against the CV hypothesis for this source_
# ```
# - 200233344 - ? 4000-5000 resolved to EPIC 211414300 2MASS J08514235+1151230 RGB star
# - 200233429 - ? 5000-6000 2MASS J08504609+1143082 spectroscopic binary
# - 211390071 - It's an SSO! 12.6 period rotationally var. 4000-5000
# If the flare is real 5.9e34 erg
# - 211396661 - It's an SSO! 21.23 period rotationally var 3750-4000, "no clear variation" (Gonzalez 2016)
# If the flare is real 4.6e34 erg
# - 211409644 - spectrocscopic binary
#
# The poster flare was in the Nardiello sample.
df2 = df[(df.real != -1) & (df.real != 0) & (df.note.isnull())]
df2[(df2.cluster=="ngc2682") & (df2.Teff_median < 6000.)].dropna(subset=["real"]).Teff_median
# # Compare Shibayama 2013 Kepler superflares to our flares
ens = np.linspace(5e33,5e34,200000)
#ens = np.linspace(5e34,5e35,200000)
1 / np.sum(np.power(ens[:-1]*.72,-1.85) * 2.05e27 / .85 *np.diff(ens)) #M67
365.25 / np.sum(np.power(ens[:-1]*.72,-1.85) * 5.2e28 / .85 *np.diff(ens)) # Pleiades
1/800, 1/5000
800/18, 5000/128
np.power(1e34,-1.85) * 2.05e27 / .85
18-130
|
VettingFlares/FFDs_vs_Davenport.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3.7
# language: python
# name: py3.7
# ---
# # Activity 7.01
# Import the Libraries
from keras.models import Sequential
from keras.layers import Conv2D
from keras.layers import MaxPool2D
from keras.layers import Flatten
from keras.layers import Dense
import numpy as np
from tensorflow import random
# Set the seed and initialize the CNN model
# +
seed = 1
np.random.seed(seed)
random.set_seed(seed)
# Initialising the CNN
classifier = Sequential()
# -
# Add the convolutional layers
# +
# Convolution
classifier.add(Conv2D(32, (3, 3), input_shape = (64, 64, 3), activation = 'relu'))
classifier.add(Conv2D(32, (3, 3), activation = 'relu'))
classifier.add(Conv2D(32, (3, 3), activation = 'relu'))
# Pooling
classifier.add(MaxPool2D(pool_size = (2, 2)))
# Flattening
classifier.add(Flatten())
# -
# Add the dense layers to the network
# Full ANN Connection
classifier.add(Dense(units = 128, activation = 'relu'))
classifier.add(Dense(128,activation='relu'))
classifier.add(Dense(128,activation='relu'))
classifier.add(Dense(128,activation='relu'))
classifier.add(Dense(units = 1, activation = 'softmax'))
# Compiling the CNN
classifier.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# Create the training and test data generators
# +
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
test_datagen = ImageDataGenerator(rescale = 1./255)
# -
# Create training and test datasets
# +
training_set = train_datagen.flow_from_directory('../dataset/training_set',
target_size = (64, 64),
batch_size = 32,
class_mode = 'binary')
test_set = test_datagen.flow_from_directory('../dataset/test_set',
target_size = (64, 64),
batch_size = 32,
class_mode = 'binary')
# -
# Fit the model to the training data
classifier.fit_generator(training_set,
steps_per_epoch = 10000,
epochs = 2,
validation_data = test_set,
validation_steps = 2500,
shuffle=False)
|
Chapter07/Activity7.01/Activity7_01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Tensorflow Multi-GPU VAE-GAN implementation
# - This is an implementation of the VAE-GAN based on the implementation described in *<a href="http://arxiv.org/abs/1512.09300">Autoencoding beyond pixels using a learned similarity metric</a>*
# - I implement a few useful things like
# - Visualizing Movement Through Z-Space
# - Latent Space Algebra
# - Spike Triggered Average Style Receptive Fields
# ### How does a VAE-GAN work?
# - We have three networks, an <font color="#38761d"><strong>Encoder</strong></font>,
# a <font color="#1155cc"><strong>Generator</strong></font>, and a <font color="#ff0000"><strong>Discriminator</strong></font>.
# - The <font color="#38761d"><strong>Encoder</strong></font> learns to map input x onto z space (latent space)
# - The <font color="#1155cc"><strong>Generator</strong></font> learns to generate x from z space
# - The <font color="#ff0000"><strong>Discriminator</strong></font> learns to discriminate whether the image being put in is real, or generated
#
# ### Diagram of basic network input and output
# 
# ### `l_x_tilde` and `l_x` here become layers of *high level features* that the discriminator learns.
# - we train the network to minimize the difference between the high level features of `x` and `x_tilde`
# - This is basically an autoencoder that works on *high level features* rather than pixels
# - Adding this *autoencoder* to a GAN helps to stabilize the GAN
# ### Training
# Train <font color="#38761d"><strong>Encoder</strong></font> on minimization of:
# - `kullback_leibler_loss(z_x, gaussian)`
# - `mean_squared_error(l_x_tilde_, l_x)`
#
# Train <font color="#1155cc"><strong>Generator</strong></font> on minimization of:
# - `kullback_leibler_loss(z_x, gaussian)`
# - `mean_squared_error(l_x_tilde_, l_x)`
# - `-1*log(d_x_p)`
#
# Train <font color="#ff0000"><strong>Discriminator</strong></font> on minimization of:
# - `-1*log(d_x) + log(1 - d_x_p)`
#
# +
# Import all of our packages
import os
import numpy as np
import prettytensor as pt
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
import matplotlib.pyplot as plt
from deconv import deconv2d
import IPython.display
import math
import tqdm # making loops prettier
import h5py # for reading our dataset
import ipywidgets as widgets
from ipywidgets import interact, interactive, fixed
# %matplotlib inline
# -
# ## Parameters
#
dim1 = 64 # first dimension of input data
dim2 = 64 # second dimension of input data
dim3 = 3 # third dimension of input data (colors)
batch_size = 32 # size of batches to use (per GPU)
hidden_size = 2048 # size of hidden (z) layer to use
num_examples = 60000 # how many examples are in your training set
num_epochs = 10000 # number of epochs to run
### we can train our different networks with different learning rates if we want to
e_learning_rate = 1e-3
g_learning_rate = 1e-3
d_learning_rate = 1e-3
# ### Which GPUs are we using?
# - Set `gpus` to a list of the GPUs you're using. The network will then split up the work between those gpus
gpus = [] # Here I set CUDA to only see one GPU
os.environ["CUDA_VISIBLE_DEVICES"]=','.join([str(i) for i in gpus])
num_gpus = len(gpus) # number of GPUs to use
cpus = [0,1,2,3]
num_cpus = len(cpus)
# ### Reading the dataset from HDF5 format
# - open `makedataset.ipynb' for instructions on how to build the dataset
with h5py.File(''.join(['datasets/faces_dataset_new.h5']), 'r') as hf:
faces = hf['images'].value
headers = hf['headers'].value
labels = hf['label_input'].value
# Normalize the dataset between 0 and 1
faces = (faces/255.)
# Just taking a look and making sure everything works
plt.imshow(np.reshape(faces[1], (64,64,3)), interpolation='nearest')
# grab the faces back out after we've flattened them
def create_image(im):
return np.reshape(im,(dim1,dim2,dim3))
# Lets just take a look at our channels
cm = plt.cm.hot
test_face = faces[0].reshape(dim1,dim2,dim3)
fig, ax = plt.subplots(nrows=1,ncols=4, figsize=(20,8))
ax[0].imshow(create_image(test_face), interpolation='nearest')
ax[1].imshow(create_image(test_face)[:,:,0], interpolation='nearest', cmap=cm)
ax[2].imshow(create_image(test_face)[:,:,1], interpolation='nearest', cmap=cm)
ax[3].imshow(create_image(test_face)[:,:,2], interpolation='nearest', cmap=cm)
# ### A data iterator for batching (drawn up by <NAME>)
# - https://indico.io/blog/tensorflow-data-inputs-part1-placeholders-protobufs-queues/
#
# +
def data_iterator():
""" A simple data iterator """
batch_idx = 0
while True:
idxs = np.arange(0, len(faces))
np.random.shuffle(idxs)
for batch_idx in range(0, len(faces), batch_size*num_cpus):
cur_idxs = idxs[batch_idx:batch_idx+batch_size*num_cpus]
images_batch = faces[cur_idxs]
#images_batch = images_batch.astype("float32")
labels_batch = labels[cur_idxs]
yield images_batch, labels_batch
iter_ = data_iterator()
# -
iter_ = data_iterator()
# +
#face_batch, label_batch
# -
# ### <NAME>
fig, ax = plt.subplots(nrows=1,ncols=4, figsize=(20,8))
ax[0].imshow(create_image(faces[labels[:,4] == 1][0]), interpolation='nearest')
ax[1].imshow(create_image(faces[labels[:,4] == 1][1]), interpolation='nearest')
ax[2].imshow(create_image(faces[labels[:,4] == 1][2]), interpolation='nearest')
ax[3].imshow(create_image(faces[labels[:,4] == 1][3]), interpolation='nearest')
# ### Draw out the architecture of our network
# - Each of these functions represent the <font color="#38761d"><strong>Encoder</strong></font>,
# <font color="#1155cc"><strong>Generator</strong></font>, and <font color="#ff0000"><strong>Discriminator</strong></font> described above.
# - It would be interesting to try and implement the inception architecture to do the same thing, next time around:
# <br /><br />
# 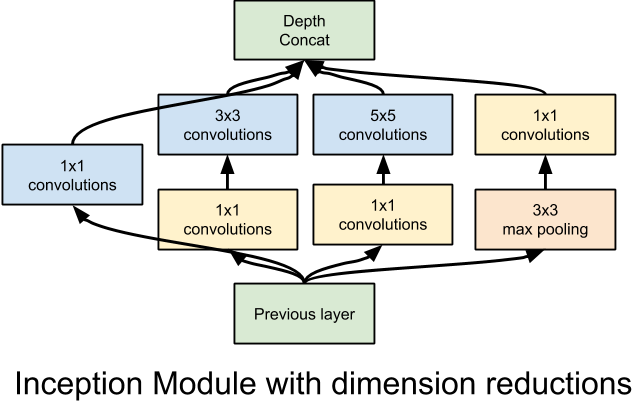
# - They describe how to implement inception, in prettytensor, here: https://github.com/google/prettytensor
# +
def encoder(X):
'''Create encoder network.
Args:
x: a batch of flattened images [batch_size, 28*28]
Returns:
A tensor that expresses the encoder network
# The transformation is parametrized and can be learned.
# returns network output, mean, setd
'''
lay_end = (pt.wrap(X).
reshape([batch_size, dim1, dim2, dim3]).
conv2d(5, 64, stride=2).
conv2d(5, 128, stride=2).
conv2d(5, 256, stride=2).
flatten())
z_mean = lay_end.fully_connected(hidden_size, activation_fn=None)
z_log_sigma_sq = lay_end.fully_connected(hidden_size, activation_fn=None)
return z_mean, z_log_sigma_sq
def generator(Z):
'''Create generator network.
If input tensor is provided then decodes it, otherwise samples from
a sampled vector.
Args:
x: a batch of vectors to decode
Returns:
A tensor that expresses the generator network
'''
return (pt.wrap(Z).
fully_connected(8*8*256).reshape([batch_size, 8, 8, 256]). #(128, 4 4, 256)
deconv2d(5, 256, stride=2).
deconv2d(5, 128, stride=2).
deconv2d(5, 32, stride=2).
deconv2d(1, dim3, stride=1, activation_fn=tf.sigmoid).
flatten()
)
def discriminator(D_I):
''' A encodes
Create a network that discriminates between images from a dataset and
generated ones.
Args:
input: a batch of real images [batch, height, width, channels]
Returns:
A tensor that represents the network
'''
descrim_conv = (pt.wrap(D_I). # This is what we're descriminating
reshape([batch_size, dim1, dim2, dim3]).
conv2d(5, 32, stride=1).
conv2d(5, 128, stride=2).
conv2d(5, 256, stride=2).
conv2d(5, 256, stride=2).
flatten()
)
lth_layer= descrim_conv.fully_connected(1024, activation_fn=tf.nn.elu)# this is the lth layer
D =lth_layer.fully_connected(1, activation_fn=tf.nn.sigmoid) # this is the actual discrimination
return D, lth_layer
# -
# ### Defining the forward pass through the network
# - This function is based upon the inference function from tensorflows cifar tutorials
# - https://github.com/tensorflow/tensorflow/blob/r0.10/tensorflow/models/image/cifar10/cifar10.py
# - Notice I use `with tf.variable_scope("enc")`. This way, we can reuse these variables using `reuse=True`. We can also specify which variables to train using which error functions based upon the label `enc`
def inference(x):
"""
Run the models. Called inference because it does the same thing as tensorflow's cifar tutorial
"""
z_p = tf.random_normal((batch_size, hidden_size), 0, 1) # normal dist for GAN
eps = tf.random_normal((batch_size, hidden_size), 0, 1) # normal dist for VAE
with pt.defaults_scope(activation_fn=tf.nn.elu,
batch_normalize=True,
learned_moments_update_rate=0.0003,
variance_epsilon=0.001,
scale_after_normalization=True):
with tf.variable_scope("enc"):
z_x_mean, z_x_log_sigma_sq = encoder(x) # get z from the input
with tf.variable_scope("gen"):
z_x = tf.add(z_x_mean,
tf.multiply(tf.sqrt(tf.exp(z_x_log_sigma_sq)), eps)) # grab our actual z
x_tilde = generator(z_x)
with tf.variable_scope("dis"):
_, l_x_tilde = discriminator(x_tilde)
with tf.variable_scope("gen", reuse=True):
x_p = generator(z_p)
with tf.variable_scope("dis", reuse=True):
d_x, l_x = discriminator(x) # positive examples
with tf.variable_scope("dis", reuse=True):
d_x_p, _ = discriminator(x_p)
return z_x_mean, z_x_log_sigma_sq, z_x, x_tilde, l_x_tilde, x_p, d_x, l_x, d_x_p, z_p
# ### Loss - define our various loss functions
# - **SSE** - we don't actually use this loss (also its the MSE), its just to see how close x is to x_tilde
# - **KL Loss** - our VAE gaussian distribution loss.
# - See https://arxiv.org/abs/1312.6114
# - **D_loss** - Our descriminator loss, how good the discriminator is at telling if something is real
# - **G_loss** - essentially the opposite of the D_loss, how good the generator is a tricking the discriminator
# - ***notice we clip our values to make sure learning rates don't explode***
def loss(x, x_tilde, z_x_log_sigma_sq, z_x_mean, d_x, d_x_p, l_x, l_x_tilde, dim1, dim2, dim3):
"""
Loss functions for SSE, KL divergence, Discrim, Generator, Lth Layer Similarity
"""
### We don't actually use SSE (MSE) loss for anything (but maybe pretraining)
SSE_loss = tf.reduce_mean(tf.square(x - x_tilde)) # This is what a normal VAE uses
# We clip gradients of KL divergence to prevent NANs
KL_loss = tf.reduce_sum(-0.5 * tf.reduce_sum(1 + tf.clip_by_value(z_x_log_sigma_sq, -10.0, 10.0)
- tf.square(tf.clip_by_value(z_x_mean, -10.0, 10.0) )
- tf.exp(tf.clip_by_value(z_x_log_sigma_sq, -10.0, 10.0) ), 1))/dim1/dim2/dim3
# Discriminator Loss
D_loss = tf.reduce_mean(-1.*(tf.log(tf.clip_by_value(d_x,1e-5,1.0)) +
tf.log(tf.clip_by_value(1.0 - d_x_p,1e-5,1.0))))
# Generator Loss
G_loss = tf.reduce_mean(-1.*(tf.log(tf.clip_by_value(d_x_p,1e-5,1.0))))# +
#tf.log(tf.clip_by_value(1.0 - d_x,1e-5,1.0))))
# Lth Layer Loss - the 'learned similarity measure'
LL_loss = tf.reduce_sum(tf.square(l_x - l_x_tilde))/dim1/dim2/dim3
return SSE_loss, KL_loss, D_loss, G_loss, LL_loss
# ### Average the gradients between towers
# - This function is taken directly from
# - https://github.com/tensorflow/tensorflow/blob/r0.10/tensorflow/models/image/cifar10/cifar10_multi_gpu_train.py
# - Basically we're taking a list of gradients from each tower, and averaging them together
def average_gradients(tower_grads):
"""Calculate the average gradient for each shared variable across all towers.
Note that this function provides a synchronization point across all towers.
Args:
tower_grads: List of lists of (gradient, variable) tuples. The outer list
is over individual gradients. The inner list is over the gradient
calculation for each tower.
Returns:
List of pairs of (gradient, variable) where the gradient has been averaged
across all towers.
"""
average_grads = []
for grad_and_vars in zip(*tower_grads):
# Note that each grad_and_vars looks like the following:
# ((grad0_gpu0, var0_gpu0), ... , (grad0_gpuN, var0_gpuN))
grads = []
for g, _ in grad_and_vars:
# Add 0 dimension to the gradients to represent the tower.
expanded_g = tf.expand_dims(g, 0)
# Append on a 'tower' dimension which we will average over below.
grads.append(expanded_g)
# Average over the 'tower' dimension.
grad = tf.concat(grads, 0)
grad = tf.reduce_mean(grad, 0)
# Keep in mind that the Variables are redundant because they are shared
# across towers. So .. we will just return the first tower's pointer to
# the Variable.
v = grad_and_vars[0][1]
grad_and_var = (grad, v)
average_grads.append(grad_and_var)
return average_grads
# ### Plot network output
# - This is just my ugly function to regularly plot the output of my network - tensorboard would probably be a better option for this
def plot_network_output():
""" Just plots the output of the network, error, reconstructions, etc
"""
random_x, recon_z, all_d= sess.run((x_p, z_x_mean, d_x_p), {all_input: example_data})
top_d = np.argsort(np.squeeze(all_d))
recon_x = sess.run((x_tilde), {z_x: recon_z})
examples = 8
random_x = np.squeeze(random_x)
recon_x = np.squeeze(recon_x)
random_x = random_x[top_d]
fig, ax = plt.subplots(nrows=3,ncols=examples, figsize=(18,6))
for i in xrange(examples):
ax[(0,i)].imshow(create_image(random_x[i]), cmap=plt.cm.gray, interpolation='nearest')
ax[(1,i)].imshow(create_image(recon_x[i]), cmap=plt.cm.gray, interpolation='nearest')
ax[(2,i)].imshow(create_image(example_data[i + (num_cpus-1)*batch_size]), cmap=plt.cm.gray, interpolation='nearest')
ax[(0,i)].axis('off')
ax[(1,i)].axis('off')
ax[(2,i)].axis('off')
fig.suptitle('Top: random points in z space | Bottom: inputs | Middle: reconstructions')
plt.show()
#fig.savefig(''.join(['imgs/test_',str(epoch).zfill(4),'.png']),dpi=100)
fig, ax = plt.subplots(nrows=1,ncols=1, figsize=(20,10), linewidth = 4)
KL_plt, = plt.semilogy((KL_loss_list), linewidth = 4, ls='-', color='r', alpha = .5, label='KL')
D_plt, = plt.semilogy((D_loss_list),linewidth = 4, ls='-', color='b',alpha = .5, label='D')
G_plt, = plt.semilogy((G_loss_list),linewidth = 4, ls='-', color='k',alpha = .5, label='G')
SSE_plt, = plt.semilogy((SSE_loss_list),linewidth = 4,ls='-', color='g',alpha = .5, label='SSE')
LL_plt, = plt.semilogy((LL_loss_list),linewidth = 4,ls='-', color='m',alpha = .5, label='LL')
axes = plt.gca()
leg = plt.legend(handles=[KL_plt, D_plt, G_plt, SSE_plt, LL_plt], fontsize=20)
leg.get_frame().set_alpha(0.5)
plt.show()
# Make lists to save the losses to
# You should probably just be using tensorboard to do any visualization(or just use tensorboard...)
G_loss_list = []
D_loss_list = []
SSE_loss_list = []
KL_loss_list = []
LL_loss_list = []
dxp_list = []
dx_list = []
tf.reset_default_graph()
# ### With your graph, define what a step is (needed for multi-gpu), and what your optimizers are for each of your networks
#with tf.Graph().as_default(), tf.device('/cpu:0'):
# Create a variable to count number of train calls
global_step = tf.get_variable(
'global_step', [],
initializer=tf.constant_initializer(0), trainable=False)
# different optimizers are needed for different learning rates (using the same learning rate seems to work fine though)
lr_D = tf.placeholder(tf.float32, shape=[])
lr_G = tf.placeholder(tf.float32, shape=[])
lr_E = tf.placeholder(tf.float32, shape=[])
with tf.variable_scope(tf.get_variable_scope(),reuse=None):
opt_D = tf.train.AdamOptimizer(lr_D, epsilon=1.0)
opt_G = tf.train.AdamOptimizer(lr_G, epsilon=1.0)
opt_E = tf.train.AdamOptimizer(lr_E, epsilon=1.0)
# ### Run all of the functions we defined above
# - `tower_grads_e` defines the list of gradients for the encoder for each tower
# - For each GPU we grab parameters corresponding to each network, we then calculate the gradients, and add them to the twoers to be averaged
#
# +
# These are the lists of gradients for each tower
tower_grads_e = []
tower_grads_g = []
tower_grads_d = []
all_input = tf.placeholder(tf.float32, [batch_size*num_cpus, dim1*dim2*dim3])
KL_param = tf.placeholder(tf.float32)
LL_param = tf.placeholder(tf.float32)
G_param = tf.placeholder(tf.float32)
# Define the network for each GPU
for i in xrange(num_cpus):
with tf.device('/cpu:%d' % i):
with tf.variable_scope('Tower' , reuse=(i>0)):
#with tf.name_scope('Tower_%d' % (i)) as scope:
# grab this portion of the input
next_batch = all_input[i*batch_size:(i+1)*batch_size,:]
# Construct the model
z_x_mean, z_x_log_sigma_sq, z_x, x_tilde, l_x_tilde, x_p, d_x, l_x, d_x_p, z_p = inference(next_batch)
# Calculate the loss for this tower
SSE_loss, KL_loss, D_loss, G_loss, LL_loss = loss(next_batch, x_tilde, z_x_log_sigma_sq, z_x_mean, d_x, d_x_p, l_x, l_x_tilde, dim1, dim2, dim3)
# specify loss to parameters
params = tf.trainable_variables()
E_params = [i for i in params if 'enc' in i.name]
G_params = [i for i in params if 'gen' in i.name]
D_params = [i for i in params if 'dis' in i.name]
# Calculate the losses specific to encoder, generator, decoder
L_e = tf.clip_by_value(KL_loss*KL_param + LL_loss, -100, 100)
L_g = tf.clip_by_value(LL_loss*LL_param+G_loss*G_param, -100, 100)
L_d = tf.clip_by_value(D_loss, -100, 100)
# Reuse variables for the next tower.
# Calculate the gradients for the batch of data on this CIFAR tower.
grads_e = opt_E.compute_gradients(L_e, var_list = E_params)
grads_g = opt_G.compute_gradients(L_g, var_list = G_params)
grads_d = opt_D.compute_gradients(L_d, var_list = D_params)
# Keep track of the gradients across all towers.
tower_grads_e.append(grads_e)
tower_grads_g.append(grads_g)
tower_grads_d.append(grads_d)
# -
# ### Now lets average, and apply those gradients
# +
# Average the gradients
grads_e = average_gradients(tower_grads_e)
grads_g = average_gradients(tower_grads_g)
grads_d = average_gradients(tower_grads_d)
# apply the gradients with our optimizers
#with tf.variable_scope(tf.get_variable_scope(),reuse=False):
train_E = opt_E.apply_gradients(grads_e, global_step=global_step)
train_G = opt_G.apply_gradients(grads_g, global_step=global_step)
train_D = opt_D.apply_gradients(grads_d, global_step=global_step)
# -
# ### Now lets actually run our session
# +
# Start the Session
init = tf.global_variables_initializer()
saver = tf.train.Saver() # initialize network saver
sess = tf.InteractiveSession(config=tf.ConfigProto(allow_soft_placement=True, log_device_placement=True))
sess.run(init)
# -
# ### Get some example data to do visualizations with
example_data, _ = iter_.next()
np.shape(example_data)
# ### Initialize our epoch number, and restore a saved network by uncommening `#tf.train...`
epoch = 0
tf.train.Saver.restore(saver, sess, 'models/faces_multiGPU_64_0000.tfmod')
# ### Now we actually run the network
# - Importantly, notice how we define the learning rates
# - `e_current_lr = e_learning_rate*sigmoid(np.mean(d_real),-.5,10)`
# - we calculate the sigmoid of how the network has been performing, and squash the learning rate using a sigmoid based on that. So if the discriminator has been winning, it's learning rate will be low, and if the generator is winning, it's learning rate will be lower on the next batch.
def sigmoid(x,shift,mult):
"""
Using this sigmoid to discourage one network overpowering the other
"""
return 1 / (1 + math.exp(-(x+shift)*mult))
fig, ax = plt.subplots(nrows=1,ncols=1, figsize=(18,4))
plt.plot(np.arange(0,1,.01), [sigmoid(i/100.,-.5,10) for i in range(100)])
ax.set_xlabel('Mean of Discriminator(Real) or Discriminator(Fake)')
ax.set_ylabel('Multiplier for learning rate')
plt.title('Squashing the Learning Rate to balance Discrim/Gen network performance')
# +
total_batch = int(np.floor(num_examples / batch_size*num_cpus)) # how many batches are in an epoch
# We balance of generator and discriminators learning rate by using a sigmoid function,
# encouraging the generator and discriminator be about equal
d_real = 0
d_fake = 0
while epoch < num_epochs:
for i in tqdm.tqdm(range(total_batch)):
iter_ = data_iterator()
# balence gen and descrim
e_current_lr = e_learning_rate*sigmoid(np.mean(d_real),-.5,15)
g_current_lr = g_learning_rate*sigmoid(np.mean(d_real),-.5,15)
d_current_lr = d_learning_rate*sigmoid(np.mean(d_fake),-.5,15)
next_batches, _ = iter_.next()
_, _, _, D_err, G_err, KL_err, SSE_err, LL_err, d_fake,d_real = sess.run([
train_E, train_G, train_D,
D_loss, G_loss, KL_loss, SSE_loss, LL_loss,
d_x_p, d_x,
],
{
lr_E: e_current_lr,
lr_G: g_current_lr,
lr_D: d_current_lr,
all_input: next_batches,
KL_param: 1,
G_param: 1,
LL_param: 1
}
)
#KL_err= SSE_err= LL_err = 1
# Save our lists
dxp_list.append(d_fake)
dx_list.append(d_real)
G_loss_list.append(G_err)
D_loss_list.append(D_err)
KL_loss_list.append(KL_err)
SSE_loss_list.append(SSE_err)
LL_loss_list.append(LL_err)
if i%300 == 0:
# print display network output
IPython.display.clear_output()
print('Epoch: '+str(epoch))
plot_network_output()
# save network
saver.save(sess,''.join(['models/faces_multiGPU_64_',str(epoch).zfill(4),'.tfmod']))
epoch +=1
# -
# ### This is how we save our network
# - Just uncomment, and name it.
total_batch
#saver.save(sess,''.join(['models/faces_multiGPU_64_',str(epoch).zfill(4),'.tfmod']))
# ### Visualize movement through z-space
# - we're using jupyter widgets to slide through z-space from one point to another
# +
n_steps = 20
examples = 10
all_x_recon = np.zeros((batch_size, dim1*dim2*dim3,n_steps))
z_point_a= np.random.normal(0,1,(batch_size,hidden_size))
z_point_b= np.random.normal(0,1,(batch_size,hidden_size))
recon_z_step = (z_point_b - z_point_a)/n_steps
for i in range(n_steps):
z_point_a += recon_z_step
all_x_recon[:,:,i] = sess.run((x_tilde), {z_x: z_point_a})
canvas = np.zeros((dim1,dim2*examples,dim3, n_steps))
print np.shape(canvas)
for f in range(n_steps):
for i in range(examples):
canvas[:,dim2*i:dim2*(i+1),:,f] = create_image(all_x_recon[i,:,f])
# -
def plt_random_faces(f):
fig, ax = plt.subplots(nrows=1,ncols=1, figsize=(18,12))
plt.imshow(canvas[:,:,:,f],interpolation='nearest')
plt.title('This slider won\.t work in Github')
plt.show()
interact(plt_random_faces, f = (0,n_steps-1,1))
# ### 'Spike Triggered Average' style receptive fields.
# - We take a look at what makes a neuron respond, by taking a bunch of images, and averaging them based on how much the neuron was activated.
def norm(x):
return (x - np.min(x)) / np.max(x - np.min(x))
# get a bunch of images and their corresponding z points in the network
recon_z = np.random.normal(0,1,(batch_size,hidden_size))
recon_x, recon_l = sess.run((x_tilde, l_x_tilde), {z_x: recon_z})
for i in range(100):
rz = np.random.normal(0,1,(batch_size,hidden_size))
rx, rl = sess.run((x_tilde, l_x_tilde), {z_x: rz})
recon_z= np.concatenate((recon_z,rz),axis = 0)
recon_l = np.concatenate((recon_l,rl),axis = 0)
recon_x = np.concatenate((recon_x,rx),axis = 0)
# #### Z-Neurons
# +
num_neurons = 25
neuron = 0
fig, ax = plt.subplots(nrows=int(np.sqrt(num_neurons)),ncols=int(np.sqrt(num_neurons)), figsize=(18,12))
for a in range(int(np.sqrt(num_neurons))):
for b in range(int(np.sqrt(num_neurons))):
proportions = (recon_z[:,neuron] - min(recon_z[:,neuron])) / max((recon_z[:,neuron] - min(recon_z[:,neuron])))
receptive_field = norm(np.sum(([proportions[i] * recon_x[i,:] for i in range(len(proportions))]),axis = 0)/np.sum(proportions)- np.mean(recon_x,axis = 0))
ax[(a,b)].imshow(create_image(receptive_field), cmap=plt.cm.gray, interpolation='nearest')
ax[(a,b)].axis('off')
neuron+=1
# -
# #### Deep Descriminator Neurons
# +
num_neurons = 25
neuron = 0
fig, ax = plt.subplots(nrows=int(np.sqrt(num_neurons)),ncols=int(np.sqrt(num_neurons)), figsize=(18,12))
for a in range(int(np.sqrt(num_neurons))):
for b in range(int(np.sqrt(num_neurons))):
proportions = (recon_l[:,neuron] - min(recon_l[:,neuron])) / max((recon_l[:,neuron] - min(recon_l[:,neuron])))
receptive_field = norm(np.sum(([proportions[i] * recon_x[i,:] for i in range(len(proportions))]),axis = 0)/np.sum(proportions)- np.mean(recon_x,axis = 0))
#test = norm(test/np.mean(test_list, axis = 0))
ax[(a,b)].imshow(create_image(receptive_field), cmap=plt.cm.gray, interpolation='nearest')
ax[(a,b)].axis('off')
neuron+=1
# -
# ### Now lets try some latent space algebra
# Here are the attribute types
print [str(i) + ': ' + headers[i] for i in range(len(headers))]
# +
# Go through a bunch of inputs, get their z values and their attributes
iter_ = data_iterator()
all_batch, all_attrib = iter_.next()
all_z = sess.run((z_x_mean), {all_input: all_batch})
all_recon_x = sess.run((x_tilde), {z_x: all_z})
for i in range(200):
next_batch, next_attrib = iter_.next()
recon_z = sess.run((z_x_mean), {all_input: next_batch})
recon_x = sess.run((x_tilde), {z_x: recon_z})
all_z = np.concatenate((all_z,recon_z),axis = 0)
all_batch = np.concatenate((all_batch,next_batch),axis = 0)
all_recon_x = np.concatenate((all_recon_x,recon_x),axis = 0)
all_attrib = np.concatenate((all_attrib,next_attrib),axis = 0)
# +
# for each attribute type, get the difference between the mean z-vector of faces with
# the attribute, and without the attribute
attr_vector_list = []
avg_attr_list = []
avg_not_attr_list = []
for i in range(np.shape(all_attrib)[1]):
has_attribute = all_attrib[:,i] == 1
average_attribute = np.mean(all_z[has_attribute], axis=0)
average_not_attribute = np.mean(all_z[has_attribute == False], axis=0)
avg_attr_list.append(average_attribute)
avg_not_attr_list.append(average_not_attribute)
attr_vector_list.append(average_attribute - average_not_attribute)
# -
feature_to_look_at = 9 # specify the attribute we want to look at
# #### Look at some blonde people (bottom), and their reconstructions (top)
# +
# show some faces which have this attribute
recon_faces = all_recon_x[all_attrib[:,feature_to_look_at] == 1,:]
new_faces = all_batch[all_attrib[:,feature_to_look_at] == 1,:]
examples = 4
canvas = np.zeros((dim1*2,dim2*examples,dim3))
for i in range(examples):
canvas[0:dim1,dim2*i:dim2*(i+1),:] = create_image(recon_faces[i])
canvas[dim1:,dim2*i:dim2*(i+1),:] = create_image(new_faces[i])
fig, ax = plt.subplots(nrows=1,ncols=1, figsize=(18,6))
ax.imshow(canvas)
ax.axis('off')
# -
# #### Take random z-points, and add the blonde vector
recon_z = np.random.normal(0,1,(batch_size,hidden_size))
recon_x = sess.run((x_tilde), {z_x: recon_z})
recon_z_with_attribute = [recon_z[i] + attr_vector_list[feature_to_look_at] for i in range(len(recon_z))]
recon_x_with_attribute = sess.run((x_tilde), {z_x: recon_z_with_attribute})
examples = 12
canvas = np.zeros((dim1*2,dim2*examples,dim3))
for i in range(examples):
canvas[:dim1,dim2*i:dim2*(i+1),:] = create_image(recon_x[i])
canvas[dim1:,dim2*i:dim2*(i+1),:] = create_image(recon_x_with_attribute[i])
fig, ax = plt.subplots(nrows=1,ncols=1, figsize=(18,6))
ax.imshow(canvas)
ax.axis('off')
plt.title('Top: random points in z space | Bottom: random points + blonde vector')
# #### Look at the average blonde person, the average not blonde person, and their difference
# +
recon_z = np.random.normal(0,1,(batch_size,hidden_size))
recon_z[0] = avg_attr_list[feature_to_look_at]
recon_z[1] = avg_not_attr_list[feature_to_look_at]
recon_z[2] = attr_vector_list[feature_to_look_at]
recon_x = sess.run((x_tilde), {z_x: recon_z})
# -
examples = 3
canvas = np.zeros((dim1,dim2*examples,dim3))
for i in range(examples):
canvas[:,dim2*i:dim2*(i+1),:] = create_image(recon_x[i])
fig, ax = plt.subplots(nrows=1,ncols=1, figsize=(18,6))
ax.imshow(canvas)
ax.axis('off')
plt.title('Average Blonde Person | Average Not Blonde Person | ABP-ANBP')
# ### This implementation is based on a few other things:
# - [Autoencoding beyond pixels](http://arxiv.org/abs/1512.09300) [*(Github)*](https://github.com/andersbll/autoencoding_beyond_pixels)
# - [VAE and GAN implementations in prettytensor/tensorflow (*Github*)](https://github.com/ikostrikov/TensorFlow-VAE-GAN-DRAW)
# - [Tensorflow VAE tutorial](https://jmetzen.github.io/2015-11-27/vae.html)
# - [DCGAN](https://arxiv.org/abs/1511.06434) [*(Github)*](https://github.com/Newmu/dcgan_code)
# - [Torch GAN tutorial](http://torch.ch/blog/2015/11/13/gan.html) [*(Github)*](https://github.com/skaae/torch-gan)
# - [Open AI improving GANS](https://openai.com/blog/generative-models/) [*(Github)*](https://github.com/openai/improved-gan)
# - Other things which I am probably forgetting...
#
|
VAE-GAN-multi-gpu-celebA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align=center>Naive Bayes</h1>
#
# <p><code>Naive Bayes</code> methods are a set of supervised learning algorithms
# based on applying Bayes’ theorem with the “naive” assumption of
# conditional independence between every pair of features given the
# value of the class variable. Bayes’ theorem states the following
# relationship, given class variable $y$ and dependent feature vector $x_1$ through $x_n$</p>
#
#
# $P(y \mid x_1, \dots, x_n) = \frac{P(y) P(x_1, \dots x_n \mid y)}
# {P(x_1, \dots, x_n)}$
# Using the naive conditional independence assumption that
#
# $P(x_i | y, x_1, \dots, x_{i-1}, x_{i+1}, \dots, x_n) = P(x_i | y)$
#
# for all , this relationship is simplified to
#
# $P(y \mid x_1, \dots, x_n) = \frac{P(y) \prod_{i=1}^{n} P(x_i \mid y)}
# {P(x_1, \dots, x_n)}$
#
# Since $P(x_1, \dots, x_n)$ is constant given the input, we can use the following classification rule:
#
# $ \begin{align}\begin{aligned}P(y \mid x_1, \dots, x_n) \propto P(y) \prod_{i=1}^{n} P(x_i \mid y)\\\Downarrow\\\hat{y} = \arg\max_y P(y) \prod_{i=1}^{n} P(x_i \mid y),\end{aligned}\end{align} $
#
# and we can use Maximum A Posteriori (MAP) estimation to estimate $P(y)$ and $P(x_i \mid y)$ the former is then the relative frequency of class $y$ in the training set.
#
#
# The different naive Bayes classifiers differ mainly by the assumptions they make regarding the distribution of $P(x_i \mid y)$
#
# In spite of their apparently over-simplified assumptions, naive Bayes classifiers have worked quite well in many real-world situations, famously document classification and spam filtering. They require a small amount of training data to estimate the necessary parameters. (For theoretical reasons why naive Bayes works well, and on which types of data it does, see the references below.)
#
# Naive Bayes learners and classifiers can be extremely fast compared to more sophisticated methods. The decoupling of the class conditional feature distributions means that each distribution can be independently estimated as a one dimensional distribution. This in turn helps to alleviate problems stemming from the curse of dimensionality.
# ### Gaussian Naive Bayes
# GaussianNB implements the Gaussian Naive Bayes algorithm for classification. The likelihood of the features is assumed to be Gaussian:
#
# $P(x_i \mid y) = \frac{1}{\sqrt{2\pi\sigma^2_y}} \exp\left(-\frac{(x_i - \mu_y)^2}{2\sigma^2_y}\right)$
#
#
# The parameters $\sigma_y$
# and $\mu_y$
# are estimated using `maximum likelihood`.
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.naive_bayes import GaussianNB
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
Data = load_iris()
Data.data
Data.target
df = pd.DataFrame(Data.data,columns=Data.feature_names)
df
df['Target'] = Data.target
df.head()
df.tail()
df.isna().sum()
sns.pairplot(df)
x = df.drop('Target',axis=1)
y = df['Target']
sns.pairplot(x)
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.35, random_state=0)
X_train.shape
X_test.shape
y_train.shape
y_test.shape
gnb = GaussianNB()
gnb.fit(X_train,y_train)
New_preddiction = gnb.predict(X_test)
New_preddiction
print("Number of mislabeled points out of a total %d points : %d"
% (X_test.shape[0], (y_test != New_preddiction).sum()))
gnb_Train_score = gnb.score(X_train,y_train)*100
gnb_Train_score
gnb_Test_score= gnb.score(X_test,y_test)*100
gnb_Test_score
from sklearn.metrics import classification_report,confusion_matrix
cm = confusion_matrix(y_test,New_preddiction)
cm
sns.heatmap(cm,annot=True)
print(classification_report(y_test,New_preddiction))
# ### Bernoulli Naive Bayes
# BernoulliNB implements the naive Bayes training and classification algorithms for data that is distributed according to multivariate Bernoulli distributions; i.e., there may be multiple features but each one is assumed to be a `binary-valued (Bernoulli, boolean) variable`. Therefore, this class requires samples to be represented as `binary-valued` feature vectors; if handed any other kind of data, a BernoulliNB instance may binarize its input (depending on the binarize parameter).
#
# The decision rule for Bernoulli naive Bayes is based on
#
# $P(x_i \mid y) = P(i \mid y) x_i + (1 - P(i \mid y)) (1 - x_i)$
#
# which differs from multinomial NB’s rule in that it explicitly penalizes the non-occurrence of a feature $i$ that is an indicator for class $y$, where the multinomial variant would simply ignore a non-occurring feature.
#
# In the case of text classification, word occurrence vectors (rather than word count vectors) may be used to train and use this classifier. BernoulliNB might perform better on some datasets, especially those with shorter documents. It is advisable to evaluate both models, if time permits.
from sklearn.naive_bayes import BernoulliNB
clf = BernoulliNB()
clf.fit(X_train,y_train)
y_pred = clf.predict(X_test)
pd.DataFrame({'Actual_data':y_test,
"New_Predication":y_pred})
print("Number of mislabeled points out of a total %d points : %d"
% (X_test.shape[0], (y_test != y_pred).sum()))
Clf_Train = clf.score(X_train,y_train)
Clf_Train
Clf_Test = clf.score(X_test,y_test)
Clf_Test
# ### Multinomial Naive Bayes
#
# MultinomialNB implements the naive Bayes algorithm for multinomially distributed data, and is one of the two classic naive Bayes variants used in text classification (where the data are typically represented as word vector counts, although tf-idf vectors are also known to work well in practice). The distribution is parametrized by vectors $\theta_y = (\theta_{y1},\ldots,\theta_{yn})$ for each class $y$,where $n$ is the number of features (in text classification, the size of the vocabulary) and $\theta_{yi}$ is the probability $P(x_i \mid y)$ of feature $i$ appearing in a sample belonging to class $y$.
# The parameters $\theta_y$ is estimated by a smoothed version of maximum likelihood, i.e. relative frequency counting:
#
# $\hat{\theta}_{yi} = \frac{ N_{yi} + \alpha}{N_y + \alpha n}$
#
# where
#
# $N_{yi} = \sum_{x \in T} x_i$ s the number of times feature $i$ appears in a sample of class $y$ in the training set $T$ and $N_{y} = \sum_{i=1}^{n} N_{yi}$ the total count of all features for class $y$
#
#
#
# The smoothing priors $\alpha \ge 0$ accounts for features not present in the learning samples and prevents zero probabilities in further computations. Setting $\alpha = 1$ is called Laplace smoothing, while $\alpha < 1$ is called Lidstone smoothing.
from sklearn.naive_bayes import MultinomialNB
mnb = MultinomialNB()
mnb.fit(X_train,y_train)
mnb.coef_
y_pred = mnb.predict(X_test)
pd.DataFrame({"Actual":y_test,
"predicated":y_pred})
print("Number of mislabeled points out of a total %d points : %d"
% (X_test.shape[0], (y_test != y_pred).sum()))
mnb_Train = mnb.score(X_train,y_train)
mnb_Train
mnb_Test = mnb.score(X_test,y_test)
mnb_Test
cm = confusion_matrix(y_test,y_pred)
cm
sns.heatmap(cm,annot=True)
print(classification_report(y_test,y_pred))
# ### Complement Naive Bayes
# ComplementNB implements the complement naive Bayes (CNB) algorithm. CNB is an adaptation of the standard multinomial naive Bayes (MNB) algorithm that is particularly suited for `imbalanced data sets`. Specifically, CNB uses statistics from the complement of each class to compute the model’s weights. The inventors of CNB show empirically that the parameter estimates for CNB are more stable than those for MNB. Further, CNB regularly outperforms MNB (often by a considerable margin) on text classification tasks. The procedure for calculating the weights is as follows:
#
# $ \begin{align}\begin{aligned}\hat{\theta}_{ci} = \frac{\alpha_i + \sum_{j:y_j \neq c} d_{ij}}
# {\alpha + \sum_{j:y_j \neq c} \sum_{k} d_{kj}}\\w_{ci} = \log \hat{\theta}_{ci}\\w_{ci} = \frac{w_{ci}}{\sum_{j} |w_{cj}|}\end{aligned}\end{align} $
#
#
# where the summations are over all documents $j$ not in class $c$, $d_{ij}$ is either the count or `tf-idf` value of term $i$ in document $j$ $\alpha_i$ is a smoothing hyperparameter like that found in MNB, and $\alpha = \sum_{i} \alpha_i$ The second normalization addresses the tendency for longer documents to dominate parameter estimates in MNB. The classification rule is
#
#
#
# $\hat{c} = \arg\min_c \sum_{i} t_i w_{ci}$
#
from sklearn.naive_bayes import ComplementNB
CNB = ComplementNB()
CNB
CNB.fit(X_train,y_train)
New_predi = CNB.predict(X_test)
pd.DataFrame({'Actual_Data':y_test,
"New_Data":New_predi})
print("Number of mislabeled points out of a total %d points : %d"
% (X_test.shape[0], (y_test != New_predi).sum()))
CNB_Train_score =CNB.score(X_train,y_train)
CNB_Train_score
CNB_Test_score= CNB.score(X_test,y_test)
CNB_Test_score
cm = confusion_matrix(y_test,New_predi)
cm
sns.heatmap(cm,annot=True)
# ### Categorical Naive Bayes
#
# CategoricalNB implements the categorical naive Bayes algorithm for `categorically distributed data`. It assumes that each feature, which is described by the index $i$, has its own categorical distribution.
#
# For each feature $i$ in the training set $X$, CategoricalNB estimates a categorical distribution for each feature i of X conditioned on the class y. The index set of the samples is defined as $J = \{ 1, \dots, m \}$ , with $m$ as the number of samples.
#
#
# The probability of category $t$ in feature $i$ given class $c$ is estimated as:
#
# $P(x_i = t \mid y = c \: ;\, \alpha) = \frac{ N_{tic} + \alpha}{N_{c} +
# \alpha n_i}$
#
# where $N_{tic} = |\{j \in J \mid x_{ij} = t, y_j = c\}|$ is the number of times category $t$ appears in the samples $x_i$ which belong to class $c$, $N_{c} = |\{ j \in J\mid y_j = c\}$ is the number of samples with class c,$\alpha$ is a smoothing parameter and $n_i$ is the number of available categories of feature $i$.
#
# CategoricalNB assumes that the sample matrix $X$ is encoded (for instance with the help of OrdinalEncoder) such that all categories for each feature $i$ are represented with numbers $0, ..., n_i - 1$ where $n_i$ is the number of available categories of feature $i$.
from sklearn.naive_bayes import CategoricalNB
CatNB = CategoricalNB()
CatNB.fit(X_train,y_train)
CatNB_train_score = CatNB.score(X_train,y_train)
CatNB_train_score
CatNB_Test_Scoer = CatNB.score(X_test,y_test)
CatNB_Test_Scoer
# + jupyter={"outputs_hidden": true}
New_pred = CatNB.predict(X_test)
pd.DataFrame({'Actual_Data':y_test,
"New_predict":New_pred})
# -
print("Number of mislabeled points out of a total %d points : %d"
% (X_test.shape[0], (y_test != New_pred).sum()))
cm = confusion_matrix(y_test,New_pred)
sns.heatmap(cm,annot=True)
print(classification_report(y_test,New_pred))
from sklearn.metrics import accuracy_score
accuracy_score(y_test,New_pred)
di = {'Multinomial_Train_score':mnb_Train,
'Multinomial_Test_score':mnb_Test,
'Gaussian_Train_score':gnb_Train_score,
'Gaussian_Test_score':gnb_Test_score,
'Bernoulli_Train_score':Clf_Train,
'Bernoulli_Test_Score':Clf_Test,
'CategoricalNB_Train_Score':CatNB_train_score,
'CategoricalNB_Test_score':CatNB_Test_Scoer,
'ComplementNB_Train_score':CNB_Train_score,
'ComplementNB_Test_score':CNB_Test_score
}
di
index=['Multinomial','Gaussian','Bernoulli','CategoricalNB','ComplementNB']
# |Model|Data type|Difference|
# |------|--------|----------|
# |Multinomial|Discrete eg=count||
# |Gaussian|Continuous|
# |Bernoulli|Binary(T/F,Y/N,0/1)|
|
Classification/Naive Bayes/ML0101EN-Naive Bayes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon SageMaker
# # <a name="0">Sample Note Book: Feature Engineering, Gradient Boosting Model, and Hyperparameter Tuning</a>
#
#
# ## Random Forest Tree Model
#
# In this notebook, we build, train, and tune by [__GridSearchCV__](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html) a [Gradient Boosting Classifier__](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingClassifier.html) to predict the __target_label__ field (default or no default) of Credit Cards from UCI dataset.
#
# 1. <a href="#1">Read the dataset</a>
# 2. <a href="#2">Exploratory Data Analysis</a>
# 3. <a href="#3">Select features to build the model</a>
# 4. <a href="#4">Resample Dataset</a>
# 4. <a href="#5">Training and test datasets</a>
# 5. <a href="#6">Data processing with Pipeline and ColumnTransformer</a>
# 6. <a href="#7">Train and tune a classifier</a>
# 7. <a href="#8">Test the classifier</a>
#
#
# ## 1. <a name="1">Read the dataset</a>
# (<a href="#0">Go to top</a>)
#
# Let's read the dataset into a dataframe, using Pandas.
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import Image
from IPython.display import display
from sklearn.datasets import dump_svmlight_file
from time import gmtime, strftime
import sys
import math
import json
# +
import pandas as pd
import warnings
warnings.filterwarnings("ignore")
df = pd.read_csv('../input/dataset/default of credit card clients.csv', error_bad_lines=False)
print('The shape of the training dataset is:', df.shape)
# -
# ## 2. <a name="2">Exploratory Data Analysis</a>
# (<a href="#0">Go to top</a>)
#
# We look at number of rows, columns and some simple statistics of the dataset.
# Print the first five rows
# NaN means missing data
df.head(10)
# Let's see the data types and non-null values for each column
df.info()
# This will print basic statistics for numerical columns
df.describe()
# #### Target distribution
#
# Let's check our target distribution.
# We can see from the target plot above that we are dealing with an imbalanced dataset. This means one result type is dominating the other one(s). But this is a regression model, and no issue for this data set
# Number of unique elements in dataset
df.nunique()
# What columns are in dataset?
df.columns
# +
# Display correlation after removing correlated features
import matplotlib.pyplot as plt
import seaborn as sns
f, ax = plt.subplots(figsize=(32, 26))
corr = df.corr()
mp = sns.heatmap(corr, mask=np.zeros_like(corr, dtype=np.bool), cmap=sns.diverging_palette(220, 10, as_cmap=True),
square=True, ax=ax, annot = True)
mp.set_title(label='dataset correlation', fontsize=20)
# +
text_categorical_features = ['SEX', 'EDUCATION', 'MARRIAGE']
numeric_categorical_features = [ 'PAY_0', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']
numeric_features = ['LIMIT_BAL', 'AGE', 'BILL_AMT1', 'BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6', 'PAY_AMT1', 'PAY_AMT2', 'PAY_AMT3', 'PAY_AMT4', 'PAY_AMT5', 'PAY_AMT6']
non_useful_features = ['ID']
label_feature = ['default payment next month']
# -
text_categorical_features = [col for col in df.select_dtypes(include = 'object').columns]
print([col for col in df.select_dtypes(include = 'object').columns])
df[numeric_features].head()
df[text_categorical_features].nunique()
df[numeric_categorical_features].nunique()
df[numeric_features].nunique()
#remove duplicates
df.drop_duplicates(inplace=True)
# +
#drop non necessary columns
#dfc.drop(features_drop, axis=1, inplace=True)
#dfc_t.drop(features_drop, axis=1, inplace=True)
# -
# ## 3. <a name="3">Select features to build the model</a>
# (<a href="#0">Go to top</a>)
#
# We build a model using selected features (except __ID, Name, Links__). That is, we build a regressor including __numerical, and categorical__ features.
label = 'default payment next month'
all_features = numeric_features + text_categorical_features + numeric_categorical_features + label_feature #+ date_features
print('Numerical categorial columns:', df[numeric_categorical_features].select_dtypes(include=np.number).columns)
print('Numerical columns:', df[numeric_features].select_dtypes(include=np.number).columns)
print('Text Categorical columns:',df[text_categorical_features].select_dtypes(include='object').columns)
# +
# Display correlation after removing correlated features
import matplotlib.pyplot as plt
import seaborn as sns
plt.subplots(figsize=(32, 26))
# corr = df[numeric_features].corr()
sns.heatmap(df[numeric_features].corr(), annot = True)
# mp.set_title(label='dataset correlation', fontsize=20)
# -
# #### checking for numerical features outliers
for c in numeric_features:
print(c)
print(df[c].value_counts(bins=10, sort=False))
df = df[df["AGE"] < 100]
for c in numeric_features:
print(c)
print(df[c].value_counts(bins=10, sort=False))
# Let's check missing values for these numerical features.
print(df[numeric_features].isna().sum())
# As a quick fix, we will apply mean imputation. This will replace the missing values with the mean value of the corresponding column.
#
# __Note__: The statistically correct way to perform mean/mode imputation before training an ML model is to compute the column-wise means on the training data only, and then use these values to impute missing data in both the train and test sets. So, we'll need to split our dataset first. Same goes for any other transformations we would like to apply to these numerical features, such as scaling.
# #### Cleaning categorical features
#
# Let's also examine the categorical features.
for c in text_categorical_features:
print(c)
print(df[c].unique()) #value_counts())
df[text_categorical_features] = df[text_categorical_features].astype('str')
df[numeric_categorical_features] = df[numeric_categorical_features].astype('str')
df[numeric_features] = df[numeric_features].astype('float32')
df[label_feature] = df[label_feature].astype('int')
# __Note on boolean type__: Most categories are strings, except the __nan__s, and the booleans __False__ and __True__. The booleans will raise errors when trying to encode the categoricals with sklearn encoders, none of which accept boolean types. If using pandas get_dummies to one-hot encode the categoricals, there's no need to convert the booleans. However, get_dummies is trickier to use with sklearn's Pipeline and GridSearch.
#
# One way to deal with the booleans is to convert them to strings, by using a mask and a map changing only the booleans.
#
# The current data set doesn't have booleans, but here is a sample code
#
# mask = df.applymap(type) != bool
#
# do = {True: 'TRUE', False: 'FALSE'}
#
# df_masked = df.where(mask, df.replace(do))
# Another way to handle the booleans is to convert them to strings by changing the type of all categoricals to 'str'. This will also affect the nans, basically performing imputation of the nans with a 'nans' placeholder value!
#
# Applying the type conversion to both categoricals and text features, takes care of the nans in the text fields as well. In case other imputations are planned for the categoricals and/or test fields, notice that the masking shown above leaves the nans unchanged.
#
# df[boolean_features] = df[boolean_features].astype('str')
# Converting categoricals into useful numerical features, will also have to wait until after the train/test split.
# ## 4. <a name="4">Re-sample DataSet</a>
# (<a href="#0">Go to top</a>)
#
# We will use sklearn's __resample()__ method for both upsampling and downsampling. Here, we will upsample negative scored data to match the positive scored number of records
#
# __Important note:__ For upsampling, we have to use __replace=True__
# +
from sklearn.utils import shuffle, resample
# Let's seperate our data into two based on the Score (True of False).
df_negative= df[df['default payment next month'] == 0]
df_positive = df[df['default payment next month'] == 1]
print("Number of records before upsampling: ")
print("1:", len(df_positive), "0:", len(df_negative))
# +
# Let's use the resample function for upsampling.
df_positive = resample(df_positive, replace=True, n_samples=len(df_negative))
# Let's put the separated data frames together.
df = pd.concat([df_negative, df_positive], axis=0)
# Let's shuffle the data
df = shuffle(df)
print("Number of records after upsampling: ")
print("Positive:", len(df[df['default payment next month'] == 1]), "Negative:", len(df[df['default payment next month'] == 0]))
# +
import matplotlib.pyplot as plt
import seaborn as sb
plt.subplots(figsize = (15,15))
sb.countplot(x=df['default payment next month'], y = None, palette = "Reds")
plt.title('Ratings after upsampling the negative class')
plt.yticks([0,5000,10000,15000,20000,25000])
# -
# ## 5. <a name="5">Training and test datasets</a>
# (<a href="#0">Go to top</a>)
#
# We split our dataset into training (90%) and test (10%) subsets using sklearn's [__train_test_split()__](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function.
# +
from sklearn.model_selection import train_test_split
train_data, test_data = train_test_split(df[all_features], test_size=0.2, shuffle=True, random_state=23)
# -
# ## 6. <a name="6">Data processing with Pipeline and ColumnTransformer</a>
# (<a href="#0">Go to top</a>)
#
# Let's build a more complex pipeline today. We first build separate pipelines to handle the numerical, categorical, and text features, and then combine them into a composite pipeline along with an estimator, a [Decision Tree Classifier](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html) here.
#
# * For the numerical features pipeline, the __numerical_processor__ below, we impute missing values with the mean using sklearn's SimpleImputer, followed by a MinMaxScaler (don't have to scale features when using Decision Trees, but it's a good idea to see how to use more data transforms). If different processing is desired for different numerical features, different pipelines should be built - just like shown below for the two text features.
#
#
# * In the categoricals pipeline, the __categorical_processor__ below, we impute with a placeholder value (no effect here as we already encoded the 'nan's), and encode with sklearn's OneHotEncoder. If computing memory is an issue, it is a good idea to check categoricals' unique values, to get an estimate of many dummy features will be created by one-hot encoding. Note the __handle_unknown__ parameter that tells the encoder to ignore (rather than throw an error for) any unique value that might show in the validation/and or test set that was not present in the initial training set.
#
#
# The selective preparations of the dataset features are then put together into a collective ColumnTransformer, to be finally used in a Pipeline along with an estimator. This ensures that the transforms are performed automatically on the raw data when fitting the model and when making predictions, such as when evaluating the model on a validation dataset via cross-validation or making predictions on a test dataset in the future.
# +
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LinearRegression
### COLUMN_TRANSFORMER ###
##########################
# Preprocess the numerical features
numerical_processor = Pipeline([
('num_imputer', SimpleImputer(strategy='mean')),
('num_scaler', MinMaxScaler()) # Shown in case is needed, not a must with Decision Trees
])
# Preprocess the categorical features
categorical_processor = Pipeline([
('cat_imputer', SimpleImputer(strategy='constant', fill_value='0')), # Shown in case is needed, no effect here as we already imputed with 'nan' strings
('cat_encoder', OneHotEncoder(handle_unknown='ignore')) # handle_unknown tells it to ignore (rather than throw an error for) any value that was not present in the initial training set.
])
# Combine all data preprocessors from above (add more, if you choose to define more!)
# For each processor/step specify: a name, the actual process, and finally the features to be processed
data_preprocessor = ColumnTransformer([
('numerical_pre', numerical_processor, numeric_features),
('categorical_pre', categorical_processor, (text_categorical_features + numeric_categorical_features)),
])
### PIPELINE ###
################
# Pipeline desired all data transformers, along with an estimator at the end
# Later you can set/reach the parameters using the names issued - for hyperparameter tuning, for example
pipeline = Pipeline([
('data_preprocessing', data_preprocessor),
('dt', GradientBoostingClassifier(n_estimators=100))
])
# Visualize the pipeline
# This will come in handy especially when building more complex pipelines, stringing together multiple preprocessing steps
from sklearn import set_config
set_config(display='diagram')
pipeline
# -
# ## 6. <a name="6">Train and tune a classifier</a>
# (<a href="#0">Go to top</a>)
#
# Let's first train and test the above composite pipeline on the train and the test sets.
# +
y_train = train_data[label]
y_train.head(10)
# +
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
# Get train data to train the pipeline
X_train = train_data[numeric_features + text_categorical_features + numeric_categorical_features]
y_train = train_data[label]
# Fit the Pipeline to training data
pipeline.fit(X_train, y_train)
# Use the fitted pipeline to make predictions on the train dataset
train_predictions = pipeline.predict(X_train)
#this is for binary classification
print(confusion_matrix(y_train, train_predictions))
print(classification_report(y_train, train_predictions))
print("Accuracy (training):", accuracy_score(y_train, train_predictions))
# Get test data to test the pipeline
X_test = test_data[numeric_features + text_categorical_features + numeric_categorical_features]
y_test = test_data[label]
# Use the fitted pipeline to make predictions on the test dataset
test_predictions = pipeline.predict(X_test)
print(confusion_matrix(y_test, test_predictions))
print(classification_report(y_test, test_predictions))
print("Accuracy (test):", accuracy_score(y_test, test_predictions))
# -
# #### Hyperparameter Tuning
#
# We next use sklearn's [GridSearchCV](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html) to look for hyperparameter combinations to improve the accuracy on the test set (and reduce the generalization gap). As GridSearchCV does cross-validation train-validation split internally,
# our data transformers inside the Pipeline context will force the correct behavior of learning data transformations on the training set, and applying the transformations to the validation set when cross-validating, as well as on the test set later when running test predictions.
#
# Also, Pipeline's steps names give easy access to hyperparameters for hyperparameter tuning while cross-validating. Parameters of the estimators in the pipeline can be accessed using the __estimator__ __ __parameter__ syntax. Note the __double underscores__ connecting the __estimator__ and __parameter__!
# +
from sklearn.model_selection import train_test_split
train_data, test_data = train_test_split(df[all_features], test_size=0.2, shuffle=True, random_state=23)
# +
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
### PIPELINE GRID_SEARCH ###
############################
# Get train data to train the pipeline
X_train = train_data[numeric_features + text_categorical_features + numeric_categorical_features]
y_train = train_data[label]
# Get test data to test the pipeline
X_test = test_data[numeric_features + text_categorical_features + numeric_categorical_features]
y_test = test_data[label]
# Parameter grid for GridSearch
parameters = {
#"dt__loss":["deviance"],
"dt__learning_rate": [0.5,0.6,0.8,1],
# " #"dt__min_samples_split": np.linspace(0.1, 0.5, 12),
#"dt__min_samples_leaf": np.linspace(0.1, 0.5, 12),
"dt__max_depth":[8,10,12],
#"dt__max_features":["sqrt"],
#"dt__criterion": ["friedman_mse", "mae"],
#"dt__subsample":[0.5, 0.618, 0.8, 0.85, 0.9, 0.95, 1.0],
"dt__n_estimators":[30,50,70,100]
}
grid_search = GridSearchCV(pipeline, param_grid=parameters, scoring='accuracy',n_jobs=4,iid=False, cv=5)
# Fit the GridSearch to our training data
grid_search.fit(X_train, y_train)
# -
print(grid_search.best_params_)
print(grid_search.best_score_)
# +
# Get the best model out of GridSearchCV
classifier = grid_search.best_estimator_
# Fit the best model to the train data once more
classifier.fit(X_train, y_train)
# -
# ### <a name="8">Test the classifier</a>
# (<a href="#0">Go to top</a>)
#
# Now we test the best model with the best parameters on "unseen" data (our test data).
#
# Before that, let's first see how the model works on the training dataset.
# +
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
# Use the fitted model to make predictions on the train dataset
train_predictions = classifier.predict(X_train)
print('Model performance on the train set:')
print(confusion_matrix(y_train, train_predictions))
print(classification_report(y_train, train_predictions))
print("Train accuracy:", accuracy_score(y_train, train_predictions))
# -
# And now, let's evaluate the performance of the classifier on the test set.
|
classification/credit_card_clients classifier/credit-card-clients-classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="jJPIig7DM95P"
# from google.colab import drive
# drive.mount('/content/drive')
# + id="yTZ5-cnpNVTZ"
# # !pip install unrar
# # !unrar x "/content/drive/MyDrive/Mask Dataset/dataset.rar"
# + id="lDcZO9KnGBE-"
# importing libraries
import tensorflow
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.applications import MobileNetV2
from tensorflow.keras.layers import AveragePooling2D
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras import callbacks
from tensorflow.keras.applications.mobilenet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import img_to_array
from sklearn.model_selection import train_test_split
from tensorflow.keras.preprocessing.image import load_img
from tensorflow.keras.utils import to_categorical
from sklearn.metrics import classification_report
import matplotlib.pyplot as plt
import numpy as np
import argparse
import pandas as pd
import glob
import os
import cv2
import seaborn as sns
import random as rand
from sklearn.utils import shuffle
# + id="USF1gMUgMUkd"
# Defining path to subdirectories in dataset
mask_path = "/content/dataset/with_mask"
no_mask_path = "/content/dataset/without_mask"
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="Y_VGTy6i05fW" outputId="fd456673-5c00-44f7-bc4f-dab1ff7859e2"
# Creating a dataframe containing labels and image paths
image_mask = []
target_mask = []
for i in os.listdir(mask_path):
pic = os.path.join(mask_path + "/", i)
image_mask.append(pic)
target_mask.append("mask")
image_no_mask = []
target_no_mask = []
for i in os.listdir(no_mask_path):
pic = os.path.join(no_mask_path + "/", i)
image_no_mask.append(pic)
target_no_mask.append("without_mask")
mask = pd.DataFrame()
mask["image"] = image_mask
mask["target"] = target_mask
no_mask = pd.DataFrame()
no_mask["image"] = image_no_mask
no_mask["target"] = target_no_mask
data = pd.concat([mask, no_mask], axis = 0, ignore_index = True)
data = shuffle(data)
data
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="WSmr6wLq09Qp" outputId="650feb4c-9cad-468e-e671-215da4a1eede"
# Plot to check dataset balance
sns.countplot(data["target"])
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="XTT5K7Gw1IlX" outputId="ddd38948-8e7c-4226-d890-616bbb2af2d4"
# Image with mask
plt.figure(figsize=(12,8))
img = load_img(mask["image"][8])
plt.imshow(img)
plt.title("With Mask", color = "green", size = 14)
plt.grid(color='#999999', linestyle='-')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="cx3lrv1s1OVf" outputId="38a959da-09bb-4cd5-84ba-3b06abe3718a"
# Image without mask
plt.figure(figsize=(12,8))
img = load_img(no_mask["image"][750])
plt.imshow(img)
plt.title("Without Mask", color = "green", size = 14)
plt.grid(color='#999999', linestyle='-')
plt.show()
# + id="PBnSA1gq1SKK"
# making train, validation and test split
train, val, test = np.split(data.sample(frac=1, random_state=42), [int(.6*len(data)), int(.8*len(data))])
# + colab={"base_uri": "https://localhost:8080/"} id="VRAOcH0P1hH7" outputId="bfd1d0c4-0f63-406f-b24f-eb097e8928f6"
train["target"].value_counts() #checking the variation of the labels
# + colab={"base_uri": "https://localhost:8080/"} id="bBTd_Jj41o9h" outputId="8bb4a4d8-11ea-4475-cfad-5d8bff15bed3"
val["target"].value_counts() #checking the variation of the labels
# + colab={"base_uri": "https://localhost:8080/"} id="Hf8JpWAH95mG" outputId="890a960a-bef6-4934-f8ac-f85effaed69c"
test["target"].value_counts() #checking the variation of the labels
# + colab={"base_uri": "https://localhost:8080/"} id="CpemruUA1xMR" outputId="451cdc44-e159-43cc-f055-92fcecd63a65"
# making generators for train, validation and test data
train_datagen = ImageDataGenerator(rescale = 1./255,
rotation_range = 40,
width_shift_range = 0.2,
height_shift_range = 0.2,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True,
fill_mode = "nearest")
train_generator = train_datagen.flow_from_dataframe(dataframe = train,
x_col = "image",
y_col = "target",
target_size = (150, 150),
batch_size = 32,
class_mode = "binary")
val_datagen = ImageDataGenerator(rescale = 1./255)
val_generator = val_datagen.flow_from_dataframe(dataframe = val,
x_col = "image",
y_col = "target",
target_size = (150, 150),
batch_size = 32,
class_mode = "binary")
test_datagen = ImageDataGenerator(rescale = 1./255)
test_generator = test_datagen.flow_from_dataframe(dataframe = test,
x_col = "image",
y_col = "target",
target_size = (150, 150),
class_mode = "binary",
batch_size = 32)
# + colab={"base_uri": "https://localhost:8080/"} id="vTuONPsP12pu" outputId="dc5a9900-c7cb-443e-85f7-fa922647de10"
# using VGG19 architecture and its wieghts to train
from keras import layers
from keras import models
from keras import optimizers
from keras.applications.vgg19 import VGG19
from keras.applications.vgg19 import preprocess_input
vgg19 = VGG19(weights='imagenet',include_top=False,input_shape=(128,128,3))
for layer in vgg19.layers:
layer.trainable = False
model = models.Sequential()
model.add(vgg19)
model.add(layers.Flatten())
model.add(layers.Dense(1,activation='sigmoid'))
model.summary()
# + id="I-aEolxl1-Rb"
# Compiling the model
model.compile(loss = "binary_crossentropy",
optimizer = optimizers.Adam(lr = 1e-4),
metrics = ["acc"])
# + id="9fJ79suY9U-Q"
# Storing the model with least validation loss using callbacks
filepath = '/content/callbacks'
callback = callbacks.ModelCheckpoint(filepath=filepath,monitor='val_loss',
mode='min',save_best_only=True)
# + colab={"base_uri": "https://localhost:8080/"} id="wvXwL6MH2BKx" outputId="26766937-30c8-4adb-e1c5-92b7cb52e00a"
# training the model on 50 epochs
history = model.fit(train_generator,
steps_per_epoch=len(train_generator)//32,
epochs=50,
validation_data=val_generator,
validation_steps=len(val_generator)//32,
callbacks=[callback])
# + id="pqRcm_vt2DFM"
# saving the model
tensorflow.keras.models.save_model(model, '/content/callbacks/model.h5')
# + id="cu8A9ORe_kSQ"
# loading the saved model
mask_model = tensorflow.keras.models.load_model('/content/callbacks/model.h5')
# + colab={"base_uri": "https://localhost:8080/", "height": 731} id="eGNM0KtmAd_i" outputId="e7cc3aad-7e88-4e68-8c8b-d5afaba3518b"
# visualising the losses
acc = history.history["acc"]
val_acc = history.history["val_acc"]
loss = history.history["loss"]
val_loss = history.history["val_loss"]
epochs = range(1, len(acc) + 1)
plt.figure(figsize = (15, 6))
plt.plot(epochs, acc, "bo", label = "Train Accuracy")
plt.plot(epochs, val_acc, "b", label = "Validation Accuracy")
plt.legend()
plt.figure(figsize = (15, 6))
plt.plot(epochs, loss, "bo", label = "Train Loss")
plt.plot(epochs, val_loss, "b", label = "Validation Loss")
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="JtKyEFgMAlcJ" outputId="e6d6cbe3-a6d5-4d63-aace-8e467558e8d2"
# testing the model
test_loss, test_acc = mask_model.evaluate(test_generator, steps = 9)
print("Test ACC: ", round(test_acc, 2))
# + id="l_buktSvCzGX"
# Test accuracy comes out to be 90%
# + colab={"base_uri": "https://localhost:8080/"} id="d1uouxhjkKo-" outputId="83de2e3f-b5db-4b36-a6fe-47ae1cf27fa2"
prediction = mask_model.predict(test_generator)
prediction
# + id="EXEuy_o9kW_x"
classes = []
for x in prediction:
if x > 0.5:
classes.append(1)
else:
classes.append(0)
# + colab={"base_uri": "https://localhost:8080/"} id="Qxx42SL9nct9" outputId="03f1e722-287e-434d-f1b5-514af22c580b"
sum(classes)
# + colab={"base_uri": "https://localhost:8080/", "height": 335} id="4F7qGOmVnwQf" outputId="c5f4ff3e-5a33-4896-d536-ae42131b56fc"
sns.countplot(classes)
# + [markdown] id="2V8_BF6KHcPB"
# Checking model on new images
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="g-IR93boCZNP" outputId="ae9fc7a5-5db1-4481-adde-55cc26c5d871"
from tensorflow.keras.preprocessing import image
test_image = image.load_img('/content/dataset/with_mask/0with_mask.jpg', target_size=(128, 128))
img = test_image
test_image = image.img_to_array(test_image)
test_image = test_image.reshape(128, 128, 3)
test_image = np.expand_dims(test_image, axis=0)
#test_image.shape
result = mask_model.predict(test_image, batch_size=1)
if result[0][0] < 0.5:
# with mask
plt.figure(figsize=(12,8))
plt.imshow(img)
plt.title("With Mask", color = "red", size = 14)
plt.grid(color='#999999', linestyle='-')
plt.show()
if result[0][0] > 0.5:
# without mask
plt.figure(figsize=(12,8))
plt.imshow(img)
plt.title("Without Mask", color = "red", size = 14)
plt.grid(color='#999999', linestyle='-')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="rMd56G3rC-Al" outputId="59a138c3-af05-49d4-8fc7-36f51614ddae"
from tensorflow.keras.preprocessing import image
test_image = image.load_img('/content/dataset/without_mask/1282 without_mask.jpg', target_size=(128, 128))
img = test_image
test_image = image.img_to_array(test_image)
test_image = test_image.reshape(128, 128, 3)
test_image = np.expand_dims(test_image, axis=0)
#test_image.shape
result = mask_model.predict(test_image, batch_size=1)
if result[0][0] < 0.5:
# with mask
plt.figure(figsize=(12,8))
plt.imshow(img)
plt.title("With Mask", color = "red", size = 14)
plt.grid(color='#999999', linestyle='-')
plt.show()
if result[0][0] > 0.5:
# without mask
plt.figure(figsize=(12,8))
plt.imshow(img)
plt.title("Without Mask", color = "red", size = 14)
plt.grid(color='#999999', linestyle='-')
plt.show()
# + id="jSu5y--RIctx"
# Thank you for Watching...
|
Mask_or_No_Mask.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# # Text Classification Project
# Now we're at the point where we should be able to:
# * Read in a collection of documents - a *corpus*
# * Transform text into numerical vector data using a pipeline
# * Create a classifier
# * Fit/train the classifier
# * Test the classifier on new data
# * Evaluate performance
#
# For this project we'll use the Cornell University Movie Review polarity dataset v2.0 obtained from http://www.cs.cornell.edu/people/pabo/movie-review-data/
#
# In this exercise we'll try to develop a classification model as we did for the SMSSpamCollection dataset - that is, we'll try to predict the Positive/Negative labels based on text content alone. In an upcoming section we'll apply *Sentiment Analysis* to train models that have a deeper understanding of each review.
# ## Perform imports and load the dataset
# The dataset contains the text of 2000 movie reviews. 1000 are positive, 1000 are negative, and the text has been preprocessed as a tab-delimited file.
# +
import numpy as np
import pandas as pd
df = pd.read_csv('../TextFiles/moviereviews.tsv', sep='\t')
df.head()
# -
len(df)
# ### Take a look at a typical review. This one is labeled "negative":
from IPython.display import Markdown, display
display(Markdown('> '+df['review'][0]))
# ## Check for missing values:
# We have intentionally included records with missing data. Some have NaN values, others have short strings composed of only spaces. This might happen if a reviewer declined to provide a comment with their review. We will show two ways using pandas to identify and remove records containing empty data.
# * NaN records are efficiently handled with [.isnull()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.isnull.html) and [.dropna()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.dropna.html)
# * Strings that contain only whitespace can be handled with [.isspace()](https://docs.python.org/3/library/stdtypes.html#str.isspace), [.itertuples()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.itertuples.html), and [.drop()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.drop.html)
#
# ### Detect & remove NaN values:
# Check for the existence of NaN values in a cell:
df.isnull().sum()
# 35 records show **NaN** (this stands for "not a number" and is equivalent to *None*). These are easily removed using the `.dropna()` pandas function.
# <div class="alert alert-info" style="margin: 20px">CAUTION: By setting inplace=True, we permanently affect the DataFrame currently in memory, and this can't be undone. However, it does *not* affect the original source data. If we needed to, we could always load the original DataFrame from scratch.</div>
# +
df.dropna(inplace=True)
len(df)
# -
# ### Detect & remove empty strings
# Technically, we're dealing with "whitespace only" strings. If the original .tsv file had contained empty strings, pandas **.read_csv()** would have assigned NaN values to those cells by default.
#
# In order to detect these strings we need to iterate over each row in the DataFrame. The **.itertuples()** pandas method is a good tool for this as it provides access to every field. For brevity we'll assign the names `i`, `lb` and `rv` to the `index`, `label` and `review` columns.
# +
blanks = [] # start with an empty list
for i,lb,rv in df.itertuples(): # iterate over the DataFrame
if type(rv)==str: # avoid NaN values
if rv.isspace(): # test 'review' for whitespace
blanks.append(i) # add matching index numbers to the list
print(len(blanks), 'blanks: ', blanks)
# -
# Next we'll pass our list of index numbers to the **.drop()** method, and set `inplace=True` to make the change permanent.
# +
df.drop(blanks, inplace=True)
len(df)
# -
# Great! We dropped 62 records from the original 2000. Let's continue with the analysis.
# ## Take a quick look at the `label` column:
df['label'].value_counts()
# ## Split the data into train & test sets:
# +
from sklearn.model_selection import train_test_split
X = df['review']
y = df['label']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# -
# ## Build pipelines to vectorize the data, then train and fit a model
# Now that we have sets to train and test, we'll develop a selection of pipelines, each with a different model.
# +
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import LinearSVC
# Naïve Bayes:
text_clf_nb = Pipeline([('tfidf', TfidfVectorizer()),
('clf', MultinomialNB()),
])
# Linear SVC:
text_clf_lsvc = Pipeline([('tfidf', TfidfVectorizer()),
('clf', LinearSVC()),
])
# -
# ## Feed the training data through the first pipeline
# We'll run naïve Bayes first
text_clf_nb.fit(X_train, y_train)
# ## Run predictions and analyze the results (naïve Bayes)
# Form a prediction set
predictions = text_clf_nb.predict(X_test)
# Report the confusion matrix
from sklearn import metrics
print(metrics.confusion_matrix(y_test,predictions))
# Print a classification report
print(metrics.classification_report(y_test,predictions))
# Print the overall accuracy
print(metrics.accuracy_score(y_test,predictions))
# Naïve Bayes gave us better-than-average results at 76.4% for classifying reviews as positive or negative based on text alone. Let's see if we can do better.
# ## Feed the training data through the second pipeline
# Next we'll run Linear SVC
text_clf_lsvc.fit(X_train, y_train)
# ## Run predictions and analyze the results (Linear SVC)
# Form a prediction set
predictions = text_clf_lsvc.predict(X_test)
# Report the confusion matrix
from sklearn import metrics
print(metrics.confusion_matrix(y_test,predictions))
# Print a classification report
print(metrics.classification_report(y_test,predictions))
# Print the overall accuracy
print(metrics.accuracy_score(y_test,predictions))
# Not bad! Based on text alone we correctly classified reviews as positive or negative **84.7%** of the time. In an upcoming section we'll try to improve this score even further by performing *sentiment analysis* on the reviews.
# ## Advanced Topic - Adding Stopwords to CountVectorizer
# By default, **CountVectorizer** and **TfidfVectorizer** do *not* filter stopwords. However, they offer some optional settings, including passing in your own stopword list.
# <div class="alert alert-info" style="margin: 20px">CAUTION: There are some [known issues](http://aclweb.org/anthology/W18-2502) using Scikit-learn's built-in stopwords list. Some words that are filtered may in fact aid in classification. In this section we'll pass in our own stopword list, so that we know exactly what's being filtered.</div>
# The [CountVectorizer](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) class accepts the following arguments:
# > *CountVectorizer(input='content', encoding='utf-8', decode_error='strict', strip_accents=None, lowercase=True, preprocessor=None, tokenizer=None, **stop_words=None**, token_pattern='(?u)\b\w\w+\b', ngram_range=(1, 1), analyzer='word', max_df=1.0, min_df=1, max_features=None, vocabulary=None, binary=False, dtype=<class 'numpy.int64'>)*
#
# [TfidVectorizer](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html) supports the same arguments and more. Under *stop_words* we have the following options:
# > stop_words : *string {'english'}, list, or None (default)*
#
# That is, we can run `TfidVectorizer(stop_words='english')` to accept scikit-learn's built-in list,<br>
# or `TfidVectorizer(stop_words=[a, and, the])` to filter these three words. In practice we would assign our list to a variable and pass that in instead.
# Scikit-learn's built-in list contains 318 stopwords:
# > <pre>from sklearn.feature_extraction import text
# > print(text.ENGLISH_STOP_WORDS)</pre>
# ['a', 'about', 'above', 'across', 'after', 'afterwards', 'again', 'against', 'all', 'almost', 'alone', 'along', 'already', 'also', 'although', 'always', 'am', 'among', 'amongst', 'amoungst', 'amount', 'an', 'and', 'another', 'any', 'anyhow', 'anyone', 'anything', 'anyway', 'anywhere', 'are', 'around', 'as', 'at', 'back', 'be', 'became', 'because', 'become', 'becomes', 'becoming', 'been', 'before', 'beforehand', 'behind', 'being', 'below', 'beside', 'besides', 'between', 'beyond', 'bill', 'both', 'bottom', 'but', 'by', 'call', 'can', 'cannot', 'cant', 'co', 'con', 'could', 'couldnt', 'cry', 'de', 'describe', 'detail', 'do', 'done', 'down', 'due', 'during', 'each', 'eg', 'eight', 'either', 'eleven', 'else', 'elsewhere', 'empty', 'enough', 'etc', 'even', 'ever', 'every', 'everyone', 'everything', 'everywhere', 'except', 'few', 'fifteen', 'fifty', 'fill', 'find', 'fire', 'first', 'five', 'for', 'former', 'formerly', 'forty', 'found', 'four', 'from', 'front', 'full', 'further', 'get', 'give', 'go', 'had', 'has', 'hasnt', 'have', 'he', 'hence', 'her', 'here', 'hereafter', 'hereby', 'herein', 'hereupon', 'hers', 'herself', 'him', 'himself', 'his', 'how', 'however', 'hundred', 'i', 'ie', 'if', 'in', 'inc', 'indeed', 'interest', 'into', 'is', 'it', 'its', 'itself', 'keep', 'last', 'latter', 'latterly', 'least', 'less', 'ltd', 'made', 'many', 'may', 'me', 'meanwhile', 'might', 'mill', 'mine', 'more', 'moreover', 'most', 'mostly', 'move', 'much', 'must', 'my', 'myself', 'name', 'namely', 'neither', 'never', 'nevertheless', 'next', 'nine', 'no', 'nobody', 'none', 'noone', 'nor', 'not', 'nothing', 'now', 'nowhere', 'of', 'off', 'often', 'on', 'once', 'one', 'only', 'onto', 'or', 'other', 'others', 'otherwise', 'our', 'ours', 'ourselves', 'out', 'over', 'own', 'part', 'per', 'perhaps', 'please', 'put', 'rather', 're', 'same', 'see', 'seem', 'seemed', 'seeming', 'seems', 'serious', 'several', 'she', 'should', 'show', 'side', 'since', 'sincere', 'six', 'sixty', 'so', 'some', 'somehow', 'someone', 'something', 'sometime', 'sometimes', 'somewhere', 'still', 'such', 'system', 'take', 'ten', 'than', 'that', 'the', 'their', 'them', 'themselves', 'then', 'thence', 'there', 'thereafter', 'thereby', 'therefore', 'therein', 'thereupon', 'these', 'they', 'thick', 'thin', 'third', 'this', 'those', 'though', 'three', 'through', 'throughout', 'thru', 'thus', 'to', 'together', 'too', 'top', 'toward', 'towards', 'twelve', 'twenty', 'two', 'un', 'under', 'until', 'up', 'upon', 'us', 'very', 'via', 'was', 'we', 'well', 'were', 'what', 'whatever', 'when', 'whence', 'whenever', 'where', 'whereafter', 'whereas', 'whereby', 'wherein', 'whereupon', 'wherever', 'whether', 'which', 'while', 'whither', 'who', 'whoever', 'whole', 'whom', 'whose', 'why', 'will', 'with', 'within', 'without', 'would', 'yet', 'you', 'your', 'yours', 'yourself', 'yourselves']
#
# However, there are words in this list that may influence a classification of movie reviews. With this in mind, let's trim the list to just 60 words:
stopwords = ['a', 'about', 'an', 'and', 'are', 'as', 'at', 'be', 'been', 'but', 'by', 'can', \
'even', 'ever', 'for', 'from', 'get', 'had', 'has', 'have', 'he', 'her', 'hers', 'his', \
'how', 'i', 'if', 'in', 'into', 'is', 'it', 'its', 'just', 'me', 'my', 'of', 'on', 'or', \
'see', 'seen', 'she', 'so', 'than', 'that', 'the', 'their', 'there', 'they', 'this', \
'to', 'was', 'we', 'were', 'what', 'when', 'which', 'who', 'will', 'with', 'you']
# Now let's repeat the process above and see if the removal of stopwords improves or impairs our score.
# +
# YOU DO NOT NEED TO RUN THIS CELL UNLESS YOU HAVE
# RECENTLY OPENED THIS NOTEBOOK OR RESTARTED THE KERNEL:
import numpy as np
import pandas as pd
df = pd.read_csv('../TextFiles/moviereviews.tsv', sep='\t')
df.dropna(inplace=True)
blanks = []
for i,lb,rv in df.itertuples():
if type(rv)==str:
if rv.isspace():
blanks.append(i)
df.drop(blanks, inplace=True)
from sklearn.model_selection import train_test_split
X = df['review']
y = df['label']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import LinearSVC
from sklearn import metrics
# -
# RUN THIS CELL TO ADD STOPWORDS TO THE LINEAR SVC PIPELINE:
text_clf_lsvc2 = Pipeline([('tfidf', TfidfVectorizer(stop_words=stopwords)),
('clf', LinearSVC()),
])
text_clf_lsvc2.fit(X_train, y_train)
predictions = text_clf_lsvc2.predict(X_test)
print(metrics.confusion_matrix(y_test,predictions))
print(metrics.classification_report(y_test,predictions))
print(metrics.accuracy_score(y_test,predictions))
# Our score didn't change that much. We went from 84.7% without filtering stopwords to 84.4% after adding a stopword filter to our pipeline. Keep in mind that 2000 movie reviews is a relatively small dataset. The real gain from stripping stopwords is improved processing speed; depending on the size of the corpus, it might save hours.
# ## Feed new data into a trained model
# Once we've developed a fairly accurate model, it's time to feed new data through it. In this last section we'll write our own review, and see how accurately our model assigns a "positive" or "negative" label to it.
# ### First, train the model
# +
# YOU DO NOT NEED TO RUN THIS CELL UNLESS YOU HAVE
# RECENTLY OPENED THIS NOTEBOOK OR RESTARTED THE KERNEL:
import numpy as np
import pandas as pd
df = pd.read_csv('../TextFiles/moviereviews.tsv', sep='\t')
df.dropna(inplace=True)
blanks = []
for i,lb,rv in df.itertuples():
if type(rv)==str:
if rv.isspace():
blanks.append(i)
df.drop(blanks, inplace=True)
from sklearn.model_selection import train_test_split
X = df['review']
y = df['label']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import LinearSVC
from sklearn import metrics
# Naïve Bayes Model:
text_clf_nb = Pipeline([('tfidf', TfidfVectorizer()),
('clf', MultinomialNB()),
])
# Linear SVC Model:
text_clf_lsvc = Pipeline([('tfidf', TfidfVectorizer()),
('clf', LinearSVC()),
])
# Train both models on the moviereviews.tsv training set:
text_clf_nb.fit(X_train, y_train)
text_clf_lsvc.fit(X_train, y_train)
# -
# ### Next, feed new data to the model's `predict()` method
myreview = "A movie I really wanted to love was terrible. \
I'm sure the producers had the best intentions, but the execution was lacking."
# +
# Use this space to write your own review. Experiment with different lengths and writing styles.
myreview =
# -
print(text_clf_nb.predict([myreview])) # be sure to put "myreview" inside square brackets
print(text_clf_lsvc.predict([myreview]))
# Great! Now you should be able to build text classification pipelines in scikit-learn, apply a variety of algorithms like naïve Bayes and Linear SVC, handle stopwords, and test a fitted model on new data.
# ## Up next: Text Classification Assessment
|
02-Text-Classification-Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
# !{sys.executable} -m pip install --user wordcloud
# +
import pandas as pd
import nltk
from nltk.tokenize import word_tokenize
from nltk.corpus import stopwords
import string
from wordcloud import WordCloud
import matplotlib.pyplot as plt
regular_punct = list(string.punctuation)
regular_punct.append("'")
df = pd.read_csv('Data/CrisisLogger/crisislogger.csv')
stopwords = nltk.corpus.stopwords.words('english')
stopwords.append("'m")
stopwords.append("n't")
stopwords.append("'s")
stopwords.append("'re")
data = df.transcriptions.str.cat(sep=' ')
tokens = word_tokenize(data)
vocabulary = set(tokens)
tokens = [w.lower() for w in tokens if not w.lower() in stopwords and not w in regular_punct]
frequency_dist = nltk.FreqDist(tokens)
wordcloud = WordCloud()
wordcloud.generate_from_frequencies(frequency_dist)
plt.imshow(wordcloud)
plt.axis("off")
plt.show(block=True)
# -
|
WordCloud.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Data Quality Analysis
# This notebook is created to found issues about data quality in 'Transactions','CustomerDemographic','NewCustomerList' and 'CustomerAdress' datasets given by Sprocket Central Pty Ltd Data Set
# +
#Importing libraries
#-------------------------------
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
pd.plotting.register_matplotlib_converters()
# %matplotlib inline
import seaborn as sns
print("setup complete!")
# +
#Opening the File.xlsx
#-------------------------------
file_path = "KPMG_VI_New_raw_data_update_final.xlsx"
# +
#Function to open the sheets as dataframes
#-------------------------------
def analysing_data(file_path, sheet, col):
data = pd.read_excel(file_path, sheet_name=sheet, index_col=0, skiprows=1, usecols=col)
return data
# +
#Sheet_1 - Transactions
sheet_1 = analysing_data(file_path,'Transactions','A:M')
#Sheet_2 - CustomerDemographic
sheet_2 = analysing_data(file_path,'CustomerDemographic','A:M')
#Sheet_3 - NewCustomersList
sheet_3 = analysing_data(file_path,'NewCustomerList','A:W')
#Sheet_4 - CustomerAdress
sheet_4 = analysing_data(file_path,'CustomerAddress','A:F')
# -
#Sheet_1 - Transactions
sheet_1.info()
#Sheet_2 - CustomerDemographic
sheet_2.info()
#Sheet_3 - NewCustomersList
sheet_3.info()
#Sheet_4 - CustomerAdress
sheet_4.info()
# +
#Function to Accuracy
#-------------------------------
def Accuracy(file):
value = file.columns.to_series().groupby(file.dtypes).groups
return value
# +
#Sheet_1 - Transactions
print('---------|Sheet(1)-Transactions:')
print(Accuracy(sheet_1))
print('\n')
#Sheet_2 - CustomerDemographic
print('---------|Sheet(2)-CustomerDemographic:')
print(Accuracy(sheet_2))
print('\n')
#Sheet_3 - NewCustomersList
print('---------|Sheet(3)-NewCustomerList:')
print(Accuracy(sheet_3))
print('\n')
#Sheet_4 - CustomerAdress
print('---------|Sheet(4)-CustomerAdress:')
print(Accuracy(sheet_4))
print('\n')
# -
#Function to Completeness
#-------------------------------
def Completeness (file):
missing_values_count = file.isnull().sum()
total_cells = np.product(file.shape)
total_missing = missing_values_count.sum()
#percent of data missed
percent = (total_missing / total_cells) * 100
complete = (total_cells - total_missing)
print(f'Data Empty:', round(percent,2))
print(f'Data Complete:', round(complete, 2))
#Creating a dataframe to stackedbar
data = pd.DataFrame({'Total_Row': file.count(axis=0), 'Missing': file.isnull().sum()})
# create stacked bar chart for monthly temperatures
data.plot.bar(stacked=True, figsize=(12,6))
plt.ylabel('Number of Records')
plt.title(f'Total Value Missed per Column')
#Sheet_1 - Transactions
print('---------|Sheet(1)-Transactions:')
print(Completeness(sheet_1))
print('\n')
sheet_1['online_order']
#Sheet_2 - CustomerDemographic
print('---------|Sheet(2)-CustomerDemographic:')
print(Completeness(sheet_2))
print('\n')
#Sheet_3 - NewCustomersList
print('---------|Sheet(3)-NewCustomerList:')
print(Completeness(sheet_3))
print('\n')
#Sheet_4 - CustomerAdress
print('---------|Sheet(4)-CustomerAdress:')
print(Completeness(sheet_4))
print('\n')
# +
#Function to Consistency
def Consistency(file_list):
neo = list()
dic = dict()
for x in file_list:
for w in x.columns:
neo.append(w)
for item in neo:
dic[item] = dic.get(item,0)+1
return dic
# -
file_list = [sheet_1, sheet_2, sheet_3, sheet_4]
Consistency(file_list)
#Comments:
#4 columns without description?
sheet_3.head()
# +
#Function to Currency - datetime64[ns]
#-------------------------------
def Currency(file, path):
print("Max date: {}".format(file[path].max()))
print('------------------')
print("Min date: {}".format(file[path].min()))
# -
Currency(sheet_1, 'transaction_date')
Currency(sheet_2, 'DOB')
sheet_2.query("DOB == '1843-12-21'")
Currency(sheet_3, 'DOB')
#'product_first_sold_date' with a wrong type data - it should be date not float and cannot transform it...
sheet_1['product_first_sold_date']
pd.to_datetime(sheet_1['product_first_sold_date'][[1]], format='%m%d%y')
# +
#Validity -> Data containing allowable values
def Validity_id(file):
for x in file:
if type(x) != int:
print(x)
else:
pass
print('All validated!')
def Validity_gender(file):
dic = dict()
for x in file:
if x not in dic:
dic[x] = 1
else:
dic[x] = dic[x] + 1
return dic
# +
print('Sheet 1 - Transactions')
print(Validity_id(sheet_1['product_id']))
print('\n')
print('Sheet 1 - Transactions')
print(Validity_id(sheet_1['customer_id']))
print('\n')
# +
print('Sheet 2 - Customer Demographic')
print(Validity_gender(sheet_2['gender']))
print('\n')
print('Sheet 3 - New Customer List')
print(Validity_gender(sheet_3['gender']))
print('\n')
# -
print('Sheet 4 - Customer Address')
print(Validity_gender(sheet_4['state']))
print('\n')
#Relevancy -> Data Items with Value Meta-Data
sheet_2['default'].head()
#Uniqueness -> Values that are duplicated
def Unique(file):
return file[file.duplicated()]
Unique(sheet_1)
sheet_3['Rank']
|
Task_1/KPMG_VI_New_raw_data_update_final.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/HSE-LAMBDA/MLatMIPS-2020/blob/master/Introduction/02-Libraries.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="Aw1cuRi7hhzz"
# # Titanic: Machine Learning from Disaster
# + [markdown] colab_type="text" id="w3muhqoaJXVL"
# 
#
# This notebook's gonna teach you to use the basic data science stack for python: jupyter, numpy, pandas and matplotlib.
# + [markdown] colab_type="text" id="I_J_iM9Hjwc3"
# ## Part I: Jupyter notebooks recap
# + [markdown] colab_type="text" id="De2BsSxGJXVV"
# **The most important feature** of jupyter notebooks for this course:
# * if you're typing something, press `Tab` to see automatic suggestions, use arrow keys + enter to pick one.
# * if you move your cursor inside some function and press `Tab`, you'll get a help window.
# + colab={} colab_type="code" id="5oQdc9MIJXVW"
# run this first
import math
# + colab={} colab_type="code" id="4FZxJodmJXVZ"
# Place your cursor at the end of the unfinished line below and press tab to
# find a function that computes arctangent from two parameters (should
# have 2 in it's name).
# Once you chose it, put an opening bracket character and press tab to
# see the docs.
math.atan2(1,2) # <---
# + [markdown] colab_type="text" id="iD6NjAugj_3y"
# ## Part II: Loading data with Pandas
# + [markdown] colab_type="text" id="K0OYaO-6JXVb"
# Pandas is a library that helps you load the data, prepare it and perform some lightweight analysis. The god object here is the `pandas.DataFrame` - a 2d table with batteries included.
#
# In the cell below we use it to read the data on the infamous titanic shipwreck.
#
# __please keep running all the code cells as you read__
# + colab={} colab_type="code" id="rMpClKdKzasG"
# !wget https://github.com/HSE-LAMBDA/MLatMIPS-2020/raw/master/Introduction/train.csv
# + colab={} colab_type="code" id="G36oVo3RJXVc"
import pandas as pd
data = pd.read_csv("train.csv", index_col='PassengerId') # this yields a pandas.DataFrame
# + colab={} colab_type="code" id="gEvLO4nvJXVf"
# Selecting rows
head = data[:10]
head #if you leave an expression at the end of a cell, jupyter will "display" it automatically
# + [markdown] colab_type="text" id="EDHw3sHgJXVj"
# #### About the data
# Here's some of the columns
# * Name - a string with person's full name
# * Survived - 1 if a person survived the shipwreck, 0 otherwise.
# * Pclass - passenger class. Pclass == 3 is cheap'n'cheerful, Pclass == 1 is for moneybags.
# * Sex - a person's gender
# * Age - age in years, if available
# * Sibsp - number of siblings on a ship
# * Parch - number of parents on a ship
# * Fare - ticket cost
# * Embarked - port where the passenger embarked
# * C = Cherbourg; Q = Queenstown; S = Southampton
# + colab={} colab_type="code" id="PWmRBFP6JXVl"
# table dimensions
print("len(data) = ", len(data))
print("data.shape = ", data.shape)
# + colab={} colab_type="code" id="dt6itIHBJXVn"
# select a single row
print(data.loc[4])
# + colab={} colab_type="code" id="wdpiKMr7JXVq"
# select a single column.
ages = data["Age"]
print(ages[:10]) # alternatively: data.Age
# + colab={} colab_type="code" id="-ao3OhDtJXVu"
# select several columns and rows at once
data.loc[5:10, ("Fare", "Pclass")] # alternatively: data[["Fare","Pclass"]].loc[5:10]
# + [markdown] colab_type="text" id="h7jqzyjLp9qe"
# ### `loc` vs `iloc`
# + [markdown] colab_type="text" id="qR-MSiiiqDB4"
# There are two ways of indexing the rows in pandas:
# * by index column values (`PassengerId` in our case) – use `data.loc` for that
# * by positional index - use `data.iloc` for that
# + [markdown] colab_type="text" id="uXCOFn0Fqr1M"
# Note that index column starts from 1, so positional index 0 will correspond to index column value 1, positional 1 to index column value 2, and so on:
# + colab={} colab_type="code" id="PWD9vfcIqjAH"
print(data.index)
print('------')
print("data.iloc[0]:")
print(data.iloc[0])
print('------')
print("data.loc[1]:")
print(data.loc[1])
# + [markdown] colab_type="text" id="ln3hUck0JXVx"
# ### Your turn:
#
# + colab={} colab_type="code" id="r0surSILJXVy"
# select passengers number 13 and 666 - did they survive?
data.loc[13]
# + colab={} colab_type="code" id="WfL_YYOVJXV3"
# compute the overall survival rate (what fraction of passengers survived the shipwreck)
# + [markdown] colab_type="text" id="NaDF00ADJXV7"
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
#
# + [markdown] colab_type="text" id="dZAezJgDJXV8"
# Pandas also has some basic data analysis tools. For one, you can quickly display statistical aggregates for each column using `.describe()`
# + colab={} colab_type="code" id="yHtn-GlbJXV8"
data.describe()
# + [markdown] colab_type="text" id="hp-OG6bOJXWA"
# Some columns contain __NaN__ values - this means that there is no data there. For example, passenger `#5` has unknown age. To simplify the future data analysis, we'll replace NaN values by using pandas `fillna` function.
#
# _Note: we do this so easily because it's a tutorial. In general, you think twice before you modify data like this._
# + colab={} colab_type="code" id="h62nzR9vJXWB"
data.iloc[5]
# + colab={} colab_type="code" id="EHEegWTOJXWK"
data['Age'] = data['Age'].fillna(value=data['Age'].mean())
data['Fare'] = data['Fare'].fillna(value=data['Fare'].mean())
# + colab={} colab_type="code" id="jSIS7m9RJXWO"
data.iloc[5]
# + [markdown] colab_type="text" id="ASsnUCRMJXWS"
# More pandas:
# * A neat [tutorial](http://pandas.pydata.org/) from pydata
# * Official [tutorials](https://pandas.pydata.org/pandas-docs/stable/tutorials.html), including this [10 minutes to pandas](https://pandas.pydata.org/pandas-docs/stable/10min.html#min)
# * Bunch of cheat sheets awaits just one google query away from you (e.g. [basics](http://blog.yhat.com/static/img/datacamp-cheat.png), [combining datasets](https://pbs.twimg.com/media/C65MaMpVwAA3v0A.jpg) and so on).
# + [markdown] colab_type="text" id="-cY1PEyznRYw"
# ## Part III: Numpy and vectorized computing
# + [markdown] colab_type="text" id="4luG6vWXJXWU"
# Almost any machine learning model requires some computational heavy lifting usually involving linear algebra problems. Unfortunately, raw python is terrible at this because each operation is interpreted at runtime.
#
# So instead, we'll use `numpy` - a library that lets you run blazing fast computation with vectors, matrices and other tensors. Again, the god oject here is `numpy.ndarray`:
# + colab={} colab_type="code" id="gcTokNL-JXWV"
import numpy as np
a = np.array([1,2,3,4,5])
b = np.array([5,4,3,2,1])
print("a = ", a)
print("b = ", b)
# math and boolean operations can applied to each element of an array
print("a + 1 =", a + 1)
print("a * 2 =", a * 2)
print("a == 2", a == 2)
# ... or corresponding elements of two (or more) arrays
print("a + b =", a + b)
print("a * b =", a * b)
# + colab={} colab_type="code" id="40slF_H1JXWY"
# Your turn: compute half-products of a and b elements (halves of products)
a * b / 2
# + colab={} colab_type="code" id="EnNXgtaaJXWa"
# compute elementwise quotient between squared a and (b plus 1)
a * a / (b +1)
# + [markdown] colab_type="text" id="KSkrSyKTnJBp"
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
# + [markdown] colab_type="text" id="uHV6HlQgoL-r"
# There's a number of functions to create arrays of zeros, ones, ascending/descending numbers etc.:
# + colab={} colab_type="code" id="dCqkfcCWoi1I"
np.zeros(shape=(3, 4))
# + colab={} colab_type="code" id="NAee3VdtorX8"
np.ones(shape=(2, 5), dtype=np.bool)
# + colab={} colab_type="code" id="ymW4lMqpozFn"
np.arange(3, 15, 2) # start, stop, step
# + colab={} colab_type="code" id="KQi-FGqto_ty"
np.linspace(0, 10, 11) # divide [0, 10] interval into 11 points
# + colab={} colab_type="code" id="7d7MlvuqpJ0S"
np.logspace(1, 10, 10, base=2, dtype=np.int64)
# + [markdown] colab_type="text" id="C4hb3shSpdBf"
# You can easily reshape arrays:
# + colab={} colab_type="code" id="83xNeecTphm4"
np.arange(24).reshape(2, 3, 4)
# + [markdown] colab_type="text" id="F_evgKCtpzpk"
# or add dimensions of size 1:
# + colab={} colab_type="code" id="It_JTbIiqCTH"
print(np.arange(3)[:, np.newaxis])
print('---')
print(np.arange(3)[np.newaxis, :])
# + [markdown] colab_type="text" id="4eE5Uq8GqO75"
# Such dimensions are automatically broadcast when doing mathematical operations:
# + colab={} colab_type="code" id="VmO2UtjtqcEU"
np.arange(3)[:, np.newaxis] + np.arange(3)[np.newaxis, :]
# + [markdown] colab_type="text" id="pfmFlVopqzId"
# There is also a number of ways to stack arrays together:
# + colab={} colab_type="code" id="WzfKzymGq3wX"
matrix1 = np.arange(50).reshape(10, 5)
matrix2 = -np.arange(20).reshape(10, 2)
np.concatenate([matrix1, matrix2], axis=1)
# + colab={} colab_type="code" id="EqJnzNehrYYP"
np.stack([matrix1[:,0], matrix2[:,0]], axis=1)
# + [markdown] colab_type="text" id="MulZJhPBr6ie"
# Any matrix can be transposed easily:
# + colab={} colab_type="code" id="KIeOCQ-wr_Q6"
matrix2.T
# + colab={} colab_type="code" id="WnajGvSrsP-t"
# Your turn: make a (7 x 5) matrix with e_ij = i
# (i - row number, j - column number)
<YOUR CODE>
# + [markdown] colab_type="text" id="56X724n8nNK9"
# ### How fast is it, harry?
# + [markdown] colab_type="text" id="J7ugCw1wJXWc"
# 
#
# Let's compare computation time for python and numpy
# * Two arrays of 10^6 elements
# * first - from 0 to 1 000 000
# * second - from 99 to 1 000 099
#
# * Computing:
# * elemwise sum
# * elemwise product
# * square root of first array
# * sum of all elements in the first array
#
# + colab={} colab_type="code" id="hkaH4AkUJXWd"
# %%time
# ^-- this "magic" measures and prints cell computation time
# Option I: pure python
arr_1 = range(1000000)
arr_2 = range(99,1000099)
a_sum = []
a_prod = []
sqrt_a1 = []
for i in range(len(arr_1)):
a_sum.append(arr_1[i]+arr_2[i])
a_prod.append(arr_1[i]*arr_2[i])
a_sum.append(arr_1[i]**0.5)
arr_1_sum = sum(arr_1)
# + colab={} colab_type="code" id="BasUnM6uJXWf"
# %%time
# Option II: start from python, convert to numpy
arr_1 = range(1000000)
arr_2 = range(99,1000099)
arr_1, arr_2 = np.array(arr_1) , np.array(arr_2)
a_sum = arr_1 + arr_2
a_prod = arr_1 * arr_2
sqrt_a1 = arr_1 ** .5
arr_1_sum = arr_1.sum()
# + colab={} colab_type="code" id="whU6BvJ6JXWi"
# %%time
# Option III: pure numpy
arr_1 = np.arange(1000000)
arr_2 = np.arange(99,1000099)
a_sum = arr_1 + arr_2
a_prod = arr_1 * arr_2
sqrt_a1 = arr_1 ** .5
arr_1_sum = arr_1.sum()
# + [markdown] colab_type="text" id="7Jc5_Z5wJXWl"
# If you want more serious benchmarks, take a look at [this](http://brilliantlywrong.blogspot.ru/2015/01/benchmarks-of-speed-numpy-vs-all.html).
# + [markdown] colab_type="text" id="H2cYqf_UJXWm"
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
# + [markdown] colab_type="text" id="OPtaPKy3uI6A"
# ### Other numpy functions and features
# + [markdown] colab_type="text" id="S-8KaYRmuM4m"
# There's also a bunch of pre-implemented operations including logarithms, trigonometry, vector/matrix products and aggregations.
# + colab={} colab_type="code" id="RklGSklbJXWn"
a = np.array([1,2,3,4,5])
b = np.array([5,4,3,2,1])
print("numpy.sum(a) = ", np.sum(a))
print("numpy.mean(a) = ", np.mean(a))
print("numpy.min(a) = ", np.min(a))
print("numpy.argmin(b) = ", np.argmin(b)) # index of minimal element
print("numpy.dot(a,b) = ", np.dot(a, b)) # dot product. Also used for matrix/tensor multiplication
print("numpy.unique(['male','male','female','female','male']) = ", np.unique(['male','male','female','female','male']))
# and tons of other stuff. see http://bit.ly/2u5q430 .
# + [markdown] colab_type="text" id="Wk8fA8HBJXWs"
# The important part: all this functionality works with dataframes, as you can get their numpy representation with `.values` (most numpy functions will even work on pure pandas objects):
# + colab={} colab_type="code" id="uE53osRgJXWs"
# calling np.max on a pure pandas column:
print("Max ticket price: ", np.max(data["Fare"]))
# calling np.argmax on a numpy representation of a pandas column
# to get its positional index:
print("\nThe guy who paid the most:\n", data.iloc[np.argmax(data["Fare"].values)])
# + colab={} colab_type="code" id="p7O68NpyJXWw"
# your code: compute mean passenger age and the oldest guy on the ship
np.max(data["Age"])
data.iloc[np.argmax(data["Age"].values)]
# + colab={} colab_type="code" id="Jkq07GT5JXW0"
print("Boolean operations")
print('a = ', a)
print('b = ', b)
print("a > 2", a > 2)
print("numpy.logical_not(a>2) = ", np.logical_not(a>2))
print("numpy.logical_and(a>2,b>2) = ", np.logical_and(a > 2,b > 2))
print("numpy.logical_or(a>2,b<3) = ", np.logical_or(a > 2, b < 3))
print("\n shortcuts")
print("~(a > 2) = ", ~(a > 2)) #logical_not(a > 2)
print("(a > 2) & (b > 2) = ", (a > 2) & (b > 2)) #logical_and
print("(a > 2) | (b < 3) = ", (a > 2) | (b < 3)) #logical_or
# + [markdown] colab_type="text" id="VBkDTTOLJXW3"
# Another numpy feature we'll need is indexing: selecting elements from an array.
# Aside from python indexes and slices (e.g. a[1:4]), numpy also allows you to select several elements at once.
# + colab={} colab_type="code" id="MhXHB6fgJXW4"
a = np.array([0, 1, 4, 9, 16, 25])
ix = np.array([1,2,5])
print("a = ", a)
print("Select by element index")
print("a[[1,2,5]] = ", a[ix])
print("\nSelect by boolean mask")
print("a[a > 5] = ", a[a > 5]) # select all elements in a that are greater than 5
print("(a % 2 == 0) =", a % 2 == 0) # True for even, False for odd
print("a[a % 2 == 0] =", a[a % 2 == 0]) # select all elements in a that are even
print("data[(data['Age'] < 18) & (data['Sex'] == 'male')] = (below)") # select male children
data[(data['Age'] < 18) & (data['Sex'] == 'male')]
# + [markdown] colab_type="text" id="T_xqJK6UJXW6"
# ### Your turn
#
# Use numpy and pandas to answer a few questions about data
# + colab={} colab_type="code" id="h6eXINxLJXW6"
# who on average paid more for their ticket, men or women?
mean_fare_men = np.mean(data[data["Sex"] == "male"]["Fare"])
mean_fare_women = np.mean(data[data["Sex"] == "female"]["Fare"])
print(mean_fare_men, mean_fare_women)
# + colab={} colab_type="code" id="EyDEaifrJXW8"
# who is more likely to survive: a child (<18 yo) or an adult?
child_survival_rate = np.sum(data[data["Age"] < 18]["Survived"])/np.shape(data[data["Age"] < 18])[0]
adult_survival_rate = np.sum(data[data["Age"] >= 18]["Survived"])/np.shape(data[data["Age"] >= 18])[0]
print(child_survival_rate, adult_survival_rate)
# + [markdown] colab_type="text" id="yt0lgtQox2e1"
# ## Part IV: plots and matplotlib
# + [markdown] colab_type="text" id="Odx24QWTJXW-"
# Using python to visualize the data is covered by yet another library: `matplotlib`.
#
# Just like python itself, matplotlib has an awesome tendency of keeping simple things simple while still allowing you to write complicated stuff with convenience (e.g. super-detailed plots or custom animations).
# + colab={} colab_type="code" id="7QxD0DoLJXW-"
import matplotlib.pyplot as plt
# %matplotlib inline
# ^-- this "magic" selects specific matplotlib backend suitable for
# jupyter notebooks. For more info see:
# https://ipython.readthedocs.io/en/stable/interactive/plotting.html#id1
# line plot
plt.plot([0,1,2,3,4,5],[0,1,4,9,16,25]);
# + colab={} colab_type="code" id="huybuiQkJXXB"
#scatter-plot
x = np.arange(5)
print("x =", x)
print("x**2 =", x**2)
print("plotting x**2 vs x:")
plt.scatter(x, x**2)
plt.show() # show the first plot and begin drawing next one
plt.plot(x, x**2);
# + colab={} colab_type="code" id="Q4Bj57P_JXXF"
# draw a scatter plot with custom markers and colors
plt.scatter([1, 1, 2, 3, 4, 4.5], [3, 2, 2, 5, 15, 24],
c=["red", "blue", "orange", "green", "cyan", "gray"],
marker="x")
# without plt.show(), several plots will be drawn on top of one another
plt.plot([0, 1, 2, 3, 4, 5], [0, 1, 4, 9, 16, 25], c="black")
# adding more sugar
plt.title("Conspiracy theory proven!!!")
plt.xlabel("Per capita alcohol consumption")
plt.ylabel("# Layers in state of the art image classifier");
# fun with correlations: http://bit.ly/1FcNnWF
# + colab={} colab_type="code" id="fz5WDA4YJXXI"
# histogram - showing data density
plt.hist([0,1,1,1,2,2,3,3,3,3,3,4,4,5,5,5,6,7,7,8,9,10])
plt.show()
plt.hist([0,1,1,1,2,2,3,3,3,3,3,4,4,5,5,5,6,7,7,8,9,10], bins=5);
# + colab={} colab_type="code" id="XmapamrGJXXM"
# plot a histogram of age and a histogram of ticket fares on separate plots
plt.subplot(211)
plt.hist(data["Age"]);
plt.subplot(212)
plt.hist(data["Fare"]);
#bonus: use tab to see if there is a way to draw a 2D histogram of age vs fare.
# + colab={} colab_type="code" id="QIzNFjffJXXP"
# make a scatter plot of passenger age vs ticket fare
m_data = data[data["Sex"] == "male"]
f_data = data[data["Sex"] == "female"]
plt.scatter(m_data["Age"], m_data["Fare"], c='r')
plt.scatter(f_data["Age"], f_data["Fare"], c='g');
# kudos if you add separate colors for men and women
# + [markdown] colab_type="text" id="bHDcDWFKJXXS"
# * Extended [tutorial](https://matplotlib.org/2.0.2/users/pyplot_tutorial.html)
# * A [cheat sheet](http://bit.ly/2koHxNF)
# * Other libraries for more sophisticated stuff: [Seaborn](https://seaborn.pydata.org/), [Plotly](https://plot.ly/python/), and [Bokeh](https://bokeh.pydata.org/en/latest/)
|
Introduction/02-Libraries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# # Selección de parámetros, validación y test
# Muchos algoritmos tienen asociados algunos parámetros que influyen en la complejidad del modelo que pueden aprender. Recuerda cuando usamos `KNeighborsRegressor`. Si cambiamos el número de vecinos a considerar, obtenemos progresivamente predicciones más y más *suavizadas*:
# <img src="figures/plot_kneigbors_regularization.png" width="100%">
# En la figura anterior, podemos ver ajustes con tres valores diferentes para ``n_neighbors``. Con ``n_neighbors=2``, los datos se sobreajustan, el modelo es muy flexible y ajusta demasiado bien el ruido que hay presente en el dataset. Para ``n_neighbors=20``, el modelo no es suficientemente flexible y no puede ajustar la variación en los datos.
#
# En la subfigura intermedia, hemos encontrado un buen punto intermedio, ``n_neighbors = 5``. Ajusta los datos bastante bien y no sufre ni de sobre-aprendizaje ni de infra-aprendizaje. Nos gustaría disponer de un método cuantitativo para identificar tanto el sobre-entrenamiento como el infra-entrenamiento y optimizar los hiperparámetros (en este caso, el número de vecinos) para llegar a los mejores resultados.
#
# Intentamos obtener un equilibrio entre recordar particularidades (y ruido) de los datos de entrenamiento y modelar la suficiente variabilidad de los mismos. Este equilibrio necesita obtenerse para cualquier algoritmo de aprendizaje automático y es un concepto central, denominado equilibrio bias-varianza o "sobre-ajuste Vs. infra-ajuste"
# <img src="figures/overfitting_underfitting_cartoon.svg" width="100%">
#
# ## Hiperparámetros, sobre-ajuste e infra-ajuste
#
# Desafortunadamente, no hay un regla general para conseguir llegar a este punto óptimo y, por ello, el usuario debe encontrar el mejor equilibrio posible entre complejidad del modelo y generalización, probando distintas opciones para los hiper-parámetros. Los hiper-parámetros son aquellos parámetros que podemos ajustar sobre un algoritmos de aprendizaje automático (algoritmo que, a su vez, ajusta los parámetros del modelo en función de los datos de entrenamiento, de ahí el "hiper"). El número de vecinos $k$ del algoritmo kNN es un hiper-parámetro.
#
# A menudo este ajuste de hiper-parámetros se hace mediante una búsqueda por fuerza bruta, por ejemplo usando varios valores de ``n_neighbors``:
# +
from sklearn.model_selection import cross_val_score, KFold
from sklearn.neighbors import KNeighborsRegressor
# Generamos un dataset sintético:
x = np.linspace(-3, 3, 100)
rng = np.random.RandomState(42)
y = np.sin(4 * x) + x + rng.normal(size=len(x))
X = x[:, np.newaxis]
cv = KFold(shuffle=True)
# Para cada parámetro, repetimos una validación cruzada
for n_neighbors in [1, 3, 5, 10, 20]:
scores = cross_val_score(KNeighborsRegressor(n_neighbors=n_neighbors), X, y, cv=cv)
print("n_neighbors: %d, rendimiento medio: %f" % (n_neighbors, np.mean(scores)))
# -
# Hay una función en scikit-learn, llamada ``validation_plot``, que produce una figura similar a la que vimos previamente. Representa un parámetro, como el número de vecinos, enfrentado a los errores de entrenamiento y validación (utilizando validación cruzada):
from sklearn.model_selection import validation_curve
n_neighbors = [1, 3, 5, 10, 20, 50]
train_scores, test_scores = validation_curve(KNeighborsRegressor(), X, y, param_name="n_neighbors",
param_range=n_neighbors, cv=cv)
plt.plot(n_neighbors, train_scores.mean(axis=1), 'b', label="precisión de entrenamiento")
plt.plot(n_neighbors, test_scores.mean(axis=1), 'g', label="precisión de test")
plt.ylabel('Precisión')
plt.xlabel('Número de vecinos')
plt.xlim([50, 0])
plt.legend(loc="best");
# <div class="alert alert-warning">
# Observa que muchos vecinos resultan en un modelo suavizado o más simple, por lo que el eje X se ha dibujado invertido.
# </div>
# Si más de un parámetro es importante, como los parámetros ``C`` y ``gamma`` de una máquina de vectores soporte (SVM) (de las cuales hablaremos después), se intentan todas las posibles combinaciones de parámetros:
# +
from sklearn.model_selection import cross_val_score, KFold
from sklearn.svm import SVR
# Hacer validación cruzada para cada combinación de parámetros:
for C in [0.001, 0.01, 0.1, 1, 10]:
for gamma in [0.001, 0.01, 0.1, 1]:
scores = cross_val_score(SVR(C=C, gamma=gamma), X, y, cv=cv)
print("C: %f, gamma: %f, valor medio de R^2: %f" % (C, gamma, np.mean(scores)))
# -
# Como esto es algo que se hace frecuentemente en aprendizaje automático, hay una clase ya implementada en scikit-learn, ``GridSearchCV``. ``GridSearchCV`` utiliza un diccionario que describe los parámetros que deberían probarse y un modelo que entrenar.
#
# La rejilla de parámetros se define como un diccionario, donde las claves son los parámetros y los valores son las cantidades a probar.
# +
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10], 'gamma': [0.001, 0.01, 0.1, 1]}
grid = GridSearchCV(SVR(), param_grid=param_grid, cv=cv, verbose=3)
# -
# Una de las cosas interesantes de GridSearchCV es que es un *meta-estimador*. Utiliza un estimador como SVR y crea un nuevo estimador que se comporta exactamente igual que SVR, por lo que podemos llamar a ``fit`` para entrenarlo:
grid.fit(X, y)
# ``GridSearchCV`` aplica un proceso algo más complejo que el visto anteriormente. Primero, ejecuta el mismo bucle de validación cruzada para encontrar la mejor combinación de parámetros. Una vez tiene la mejor combinación, ejecuta el método ``fit`` de nuevo sobre todos los datos que se le pasan (sin validación cruzada), para construir un nuevo modelo con los parámetros óptimos obtenidos anteriormente.
# Después, utilizando los métodos ``predict`` o ``score`` podemos realizar una nueva predicción:
#
grid.predict(X)
# Puedes observar los mejores parámetros obtenidos por ``GridSearchCV`` en su atributo ``best_params_`` y la puntuación correspondiente en su atributo ``best_score_``:
print(grid.best_score_)
print(grid.best_params_)
# Pero puedes investigar más a fondo el rendimiento y algunas cosas más sobre cada una de las combinaciones de parámetros accediendo al atributo `cv_results_`. `cv_results_` es un diccionario donde cada clave es una cadena y cada valor un array. Se puede por tanto usar para crear un ``DataFrame`` de pandas.
type(grid.cv_results_)
print(grid.cv_results_.keys())
# +
import pandas as pd
cv_results = pd.DataFrame(grid.cv_results_)
cv_results.head()
# -
cv_results_tiny = cv_results[['param_C', 'param_gamma', 'mean_test_score']]
cv_results_tiny.sort_values(by='mean_test_score', ascending=False).head()
# Sin embargo, hay un problema en la utilización de este rendimiento para la evaluación. Puedes estar incurriendo en lo que se denomina un error de probar varias hipótesis. Si tienes muchas combinaciones de parámetros, algunas de ellas puede ser que funcionen mejor solo por aleatoriedad y que el rendimiento que estás obteniendo no sea el mismo cuando tengamos nuevos datos. Por tanto, es en general buena idea realizar una separación en entrenamiento y test previa a la búsqueda *grid*. Este patrón se suele denominar partición de entrenamiento, test y validación, y es bastante común en aprendizaje automático:
# <img src="figures/grid_search_cross_validation.svg" width="100%">
# Podemos emular este proceso fácilmente dividiendo primero los datos con ``train_test_split``, aplicando ``GridSearchCV`` al conjunto de entrenamiento, y calculando el ``score`` correspondiente solo con el conjunto de test:
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10], 'gamma': [0.001, 0.01, 0.1, 1]}
cv = KFold(n_splits=10, shuffle=True)
grid = GridSearchCV(SVR(), param_grid=param_grid, cv=cv)
grid.fit(X_train, y_train)
grid.score(X_test, y_test)
# -
# Podemos comprobar de nuevo los parámetros obtenidos con:
grid.best_params_
# A veces se utiliza un esquema más simple, que parte los datos en tres subconjuntos entrenamiento, validación y test. Esto es una alternativa si tu conjunto de datos es muy grande o si es imposible entrenar muchos modelos mediante validación cruzada, porque entrenar cada modelo es muy costoso computacionalmente. Para hacer este tipo de partición tendríamos que hacer una partición con ``train_test_split`` y después aplicar ``GridSearchCV`` con un ``ShuffleSplit`` y una sola iteración:
#
# <img src="figures/train_validation_test2.svg" width="100%">
# +
from sklearn.model_selection import train_test_split, ShuffleSplit
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10], 'gamma': [0.001, 0.01, 0.1, 1]}
single_split_cv = ShuffleSplit(n_splits=1)
grid = GridSearchCV(SVR(), param_grid=param_grid, cv=single_split_cv, verbose=3)
grid.fit(X_train, y_train)
grid.score(X_test, y_test)
# -
# Esto es mucho más rápido pero puede resultar en valores peores de los hiper-parámetros y, por tanto, peores resultados.
clf = GridSearchCV(SVR(), param_grid=param_grid)
clf.fit(X_train, y_train)
clf.score(X_test, y_test)
# <div class="alert alert-success">
# <b>EJERCICIO</b>:
# <ul>
# <li>
# Aplica una búsqueda *grid* para encontrar el mejor valor del parámetro número de vecinos para el ``KNeighborsClassifier`` para el dataset de ``digits``.
# </li>
# </ul>
# </div>
|
notebooks-spanish/14-complejidad_modelos_busqueda_grid.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using the `inferelator_view` module
#
# The `jp_doodle.inferelator_view` module provides interfaces
# for presenting information generated by the inferelator network
# inference techniques.
#
# The "user friendly" interfacese of the module read data from
# a TAB seperated values file as generated by the inferelator
# and either create a Jupyter widget view of the data requested
# or a stand alone HTML file containing an interactive visualization
# of the data requested.
#
# For example below we create two Jupyter widget viewe of a a network.
# +
file_name = "network.tsv"
from jp_doodle import inferelator_view
# The subnetwork related to Akna and Jax2
nodes = "<NAME>".split()
inferelator_view.relatedness_view(file_name, nodes)
# -
# The subnetwork containing the 100 most positive and 100 most negative edges
inferelator_view.head_and_tail_view(file_name, 100, 100)
# # Command line interface
#
# The "script" command line interface to the module can be used to create stand alone HTML files.
# !python -m jp_doodle.inferelator_view -h
# +
# Create a network with edges related to Ahr and Sox4 in "N.html"
# !python -m jp_doodle.inferelator_view network.tsv Ahr Sox4 --out N.html
# +
# Create a network with the 100 most positive and most negative edges in N2.html
# !python -m jp_doodle.inferelator_view network.tsv --negative_limit 100 --positive_limit 100 --out N2.html
# -
|
notebooks/misc/inferelator/Using the inferelator view module.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # General Implementation of Grover Over Limited Search Space Using W states (Oracle 1)
# ---
import networkx as nx
import numpy as np
import math
import itertools as it
import matplotlib.pyplot as plt
import matplotlib.axes as axes
import warnings
from random import choice, randint
import os
import re
import itertools
from IPython.display import IFrame
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, Aer, execute
from qiskit.tools.visualization import plot_histogram
import csv
# %matplotlib inline
warnings.filterwarnings("ignore", category=UserWarning)
S_simulator = Aer.backends(name='statevector_simulator')[0]
M_simulator = Aer.backends(name='qasm_simulator')[0]
#A function to display results in a ket format (as descripted in arXiv:1903.04359v1)
def Measurement(quantumcircuit, *args, **kwargs):
#Displays the measurement results of a quantum circuit
p_M = True
S = 1
ref = False
NL = False
if 'shots' in kwargs:
S = int(kwargs['shots'])
if 'return_M' in kwargs:
ret = kwargs['return_M']
if 'print_M' in kwargs:
p_M = kwargs['print_M']
if 'column' in kwargs:
NL = kwargs['column']
M1 = execute(quantumcircuit, M_simulator, shots=S).result().get_counts(quantumcircuit)
M2 = {}
k1 = list(M1.keys())
v1 = list(M1.values())
for k in np.arange(len(k1)):
key_list = list(k1[k])
new_key = ''
for j in np.arange(len(key_list)):
new_key = new_key+key_list[len(key_list)-(j+1)]
M2[new_key] = v1[k]
if(p_M):
k2 = list(M2.keys())
v2 = list(M2.values())
measurements = ''
for i in np.arange(len(k2)):
m_str = str(v2[i])+'|'
for j in np.arange(len(k2[i])):
if(k2[i][j] == '0'):
m_str = m_str + '0'
if(k2[i][j] == '1'):
m_str = m_str + '1'
if( k2[i][j] == ' ' ):
m_str = m_str +'>|'
m_str = m_str + '> '
if(NL):
m_str = m_str + '\n'
measurements = measurements + m_str
#print(measurements)
return measurements
if(ref):
return M2
# # Get graph from list of edges
# ---
#Produce a graph from a list of edges
#The edges list must be ordered as in, all edges from node 0 then 1 then 2 and so on...
G = nx.Graph()
default_axes = plt.axes(frameon=True)
#[(0, 1), (0, 3), (1, 3), (1, 2)] Orignal
#[(0, 1), (2, 3), (1, 3), (1, 2)] Test
#edges = [(4,1),(4,2),(4,3),(1,2),(1,3),(1,0),(2,3),(2,0)]
#edges = [(0, 1), (2, 3), (0, 2), (1, 2)]#The graph we used so far
#edges = [(0,1),(0,3),(1,2),(1,3),(2,4),(2,5),(3,4),(4,5)] #two non-overlapping triangles.
edges =[(0, 1), (2, 3), (0, 2), (1, 2)]
#1- Find number of nodes based on a given list of edges
n_nodes = len(set(itertools.chain.from_iterable(edges)))
#2- Draw graph based on number of nodes and edges
G.add_edges_from(edges)
G.add_nodes_from(range(n_nodes))
G.add_edges_from(edges)
colors = ['r' for node in G.nodes()]
pos = nx.shell_layout(G)
nx.draw_networkx(G, node_color=colors, node_size=600, alpha=.8, ax=default_axes, pos=pos)
# # Define the Wn gate
# ---
# +
#Implementation of W state as described in arXiv:1807.05572v1
def cg (qcir,cQbit,tQbit,theta):
theta_dash = math.asin(math.cos(math.radians(theta/2)))
qcir.u3(theta_dash,0,0,tQbit)
qcir.cx(cQbit,tQbit)
qcir.u3(-theta_dash,0,0,tQbit)
return qcir
def wn (qcir,qbits):
for i in range(len(qbits)):
if i == 0:
qcir.x(qbits[0])
qcir.barrier()
else:
p = 1/(len(qbits)-(i-1))
theta = math.degrees(math.acos(math.sqrt(p)))
theta = 2* theta
qcir = cg(qcir,qbits[i-1],qbits[i],theta)
qcir.cx(qbits[i],qbits[i-1])
qcir.barrier()
#print(p)
return qcir,qbits
# -
# # State Preparation
# ---
k = 3 #Size of the clique, in this case a triangle
sub_qbits = QuantumRegister(n_nodes)
sub_cir = QuantumCircuit(sub_qbits, name="state")
sub_cir, sub_qbits = wn(sub_cir, sub_qbits)
sub_cir.x(sub_qbits)
stat_prep = sub_cir.to_instruction()
inv_stat_prep = sub_cir.inverse().to_instruction()
print(sub_cir.decompose().size())
state_prep_size = sub_cir.decompose().size()-1
print("total number of operations in the circuit.")
print(sub_cir.size())
print("depth of circuit (number of ops on the critical path)")
print(sub_cir.depth())
print("a breakdown of operations by type")
print(sub_cir.count_ops())
# # Define cz gate
# ---
def cnz(qc, num_control, node, anc):
"""
num_control : number of control qubit of cnz gate
node : node qubit
anc : ancillaly qubit
"""
if num_control>2:
qc.ccx(node[0], node[1], anc[0])
for i in range(num_control-2):
qc.ccx(node[i+2], anc[i], anc[i+1])
qc.cz(anc[num_control-2], node[num_control])
for i in range(num_control-2)[::-1]:
qc.ccx(node[i+2], anc[i], anc[i+1])
qc.ccx(node[0], node[1], anc[0])
if num_control==2:
qc.h(node[2])
qc.ccx(node[0], node[1], node[2])
qc.h(node[2])
if num_control==1:
qc.cz(node[0], node[1])
# # The Diff operator
# ---
def grover_diff(qc, nodes_qubits,edge_anc,ancilla,stat_prep,inv_stat_prep):
qc.append(inv_stat_prep,qargs=nodes_qubits)
qc.x(nodes_qubits)
#====================================================
#3 control qubits Z gate
cnz(qc,len(nodes_qubits)-1,nodes_qubits[::-1],ancilla)
#====================================================
qc.x(nodes_qubits)
qc.append(stat_prep,qargs=nodes_qubits)
# # The Oracle
# ---
def edge_counter(qc,qubits,anc,flag_qubit,k):
bin_k = bin(k)[2:][::-1]
l = []
for i in range(len(bin_k)):
if int(bin_k[i]) == 1:
l.append(qubits[i])
qc.mct(l,flag_qubit,[anc])
def oracle(n_nodes, edges, qc, nodes_qubits, edge_anc, ancilla, neg_base):
#1- edge counter
#forward circuit
qc.barrier()
qc.ccx(nodes_qubits[edges[0][0]],nodes_qubits[edges[0][1]],edge_anc[0])
for i in range(1,len(edges)):
qc.mct([nodes_qubits[edges[i][0]],nodes_qubits[edges[i][1]],edge_anc[0]],edge_anc[1],[ancilla[0]])
qc.ccx(nodes_qubits[edges[i][0]],nodes_qubits[edges[i][1]],edge_anc[0])
#----------------------------------------------------------------------------------------------------------
#Edges check Qubit
#print(k)
edg_k = int((k/2)*(k-1))
edge_counter(qc,edge_anc,ancilla[0],neg_base[0],edg_k)
#----------------------------------------------------------------------------------------------------------
#4- Reverse edge count
for i in range(len(edges)-1,0,-1):
qc.ccx(nodes_qubits[edges[i][0]],nodes_qubits[edges[i][1]],edge_anc[0])
qc.mct([nodes_qubits[edges[i][0]],nodes_qubits[edges[i][1]],edge_anc[0]],edge_anc[1],[ancilla[0]])
qc.ccx(nodes_qubits[edges[0][0]],nodes_qubits[edges[0][1]],edge_anc[0])
qc.barrier()
#return nodes_qubits, edge_anc,node_anc, ancilla, neg_base, class_bits, tri_flag
# # The Algorithm
# ---
# Grover algo function
def grover(x,n_nodes,stat_prep,inv_stat_prep): # X >> Number of iterations
N = 2**n_nodes # for optimal iterations count
nodes_qubits = QuantumRegister(n_nodes, name='nodes')
edge_anc = QuantumRegister(2, name='edge_anc')
#node_anc = QuantumRegister(2, name='node_anc')
ancilla = QuantumRegister(n_nodes-2, name = 'cccx_diff_anc')
neg_base = QuantumRegister(1, name='check_qubits')
class_bits = ClassicalRegister(n_nodes, name='class_reg')
tri_flag = ClassicalRegister(3, name='tri_flag')
qc = QuantumCircuit(nodes_qubits, edge_anc, ancilla, neg_base, class_bits, tri_flag)
# Initialize qunatum flag qubits in |-> state
qc.x(neg_base[0])
qc.h(neg_base[0])
# Initializing i/p qubits in superposition
qc.append(stat_prep,qargs=nodes_qubits)
qc.barrier()
# Calculate iteration count
iterations = round(math.pi/4*math.sqrt(N))
# Calculate iteration count
for i in np.arange(x):
qc.barrier()
oracle(n_nodes, edges, qc, nodes_qubits, edge_anc, ancilla, neg_base)
qc.barrier()
grover_diff(qc, nodes_qubits,edge_anc,ancilla,stat_prep,inv_stat_prep)
return qc, nodes_qubits, edge_anc, ancilla, neg_base, class_bits, tri_flag
# # For different iterations
# ---
l = [] #a list to store different number of iterations results
N = 2**n_nodes # for iterations count
data =[]
n_ans = 2
iterations = round(math.pi/4*math.sqrt(N/n_ans))
for i in range(1,2): #Grover for different iterations (optimal is 1)
qc, nodes_qubits, edge_anc, ancilla, neg_base, class_bits, tri_flag = grover(i,n_nodes,stat_prep,inv_stat_prep)
qc.measure(nodes_qubits,class_bits)
#Executing circuit and show results
ex = execute(qc, M_simulator, shots = 5000)
res = ex.result()
M = res.get_counts(qc)
#printing measurements results
s = Measurement(qc,shots=5000)
pattern = "\d+\|\d{4}\>"
res = re.findall(pattern,s)
dct = {}
for item in res:
if item[item.index("|"):] not in dct.keys():
dct[item[item.index("|"):]] = int(item[:item.index("|")])
else:
dct[item[item.index("|"):]] += int(item[:item.index("|")])
l.append(dct)
#print(dct)
print("===========================")
print("at "+str(i)+" Iterations")
print(dct)
print("total number of operations in the circuit.")
print(qc.size())
print("depth of circuit (number of ops on the critical path)")
print(qc.depth())
print("number of qubits in the circuit")
print(len(qc.qubits))
print("a breakdown of operations by type")
print(qc.decompose().count_ops())
data.append([str(i),str(qc.size()),str(qc.depth()),str(len(qc.qubits)),qc.count_ops()])
# # Adding Noise
# ---
# ## Needed Imports
from qiskit.quantum_info import Kraus, SuperOp
from qiskit.providers.aer import QasmSimulator
# Import from Qiskit Aer noise module
from qiskit.providers.aer.noise import NoiseModel
from qiskit.providers.aer.noise import QuantumError, ReadoutError
from qiskit.providers.aer.noise import pauli_error
from qiskit.providers.aer.noise import depolarizing_error
from qiskit.providers.aer.noise import thermal_relaxation_error
simulator = QasmSimulator()
# ## Error Probability
# ### Bit-flip noise
# +
# Example error probabilities
p_reset = 0.3
p_meas = 0.3
p_gate1 = 0.5
# QuantumError objects
error_reset = pauli_error([('X', p_reset), ('I', 1 - p_reset)])
error_meas = pauli_error([('X',p_meas), ('I', 1 - p_meas)])
error_gate1 = pauli_error([('X',p_gate1), ('I', 1 - p_gate1)])
error_gate2 = error_gate1.tensor(error_gate1)
# Add errors to noise model
noise_bit_flip = NoiseModel()
noise_bit_flip.add_all_qubit_quantum_error(error_reset, "reset")
noise_bit_flip.add_all_qubit_quantum_error(error_meas, "measure")
noise_bit_flip.add_all_qubit_quantum_error(error_gate1, ["u1", "u2", "u3"])
noise_bit_flip.add_all_qubit_quantum_error(error_gate2, ["cx"])
print(noise_bit_flip)
# -
# ### Thermal noise
def noise_func(n,T1in,T2in):
# T1 and T2 values for qubits 0-3
T1s = np.random.normal(T1in, 10e3, n) # Sampled from normal distribution mean 50 microsec
T2s = np.random.normal(T2in, 10e3, n) # Sampled from normal distribution mean 50 microsec
# Truncate random T2s <= T1s
T2s = np.array([min(T2s[j], 2 * T1s[j]) for j in range(n)])
# Instruction times (in nanoseconds)
time_u1 = 0 # virtual gate
time_u2 = 50 # (single X90 pulse)
time_u3 = 100 # (two X90 pulses)
time_cx = 300
time_reset = 1000 # 1 microsecond
time_measure = 1000 # 1 microsecond
# QuantumError objects
errors_reset = [thermal_relaxation_error(t1, t2, time_reset)
for t1, t2 in zip(T1s, T2s)]
errors_measure = [thermal_relaxation_error(t1, t2, time_measure)
for t1, t2 in zip(T1s, T2s)]
errors_u1 = [thermal_relaxation_error(t1, t2, time_u1)
for t1, t2 in zip(T1s, T2s)]
errors_u2 = [thermal_relaxation_error(t1, t2, time_u2)
for t1, t2 in zip(T1s, T2s)]
errors_u3 = [thermal_relaxation_error(t1, t2, time_u3)
for t1, t2 in zip(T1s, T2s)]
errors_cx = [[thermal_relaxation_error(t1a, t2a, time_cx).expand(
thermal_relaxation_error(t1b, t2b, time_cx))
for t1a, t2a in zip(T1s, T2s)]
for t1b, t2b in zip(T1s, T2s)]
# Add errors to noise model
noise_thermal = NoiseModel()
for j in range(n):
noise_thermal.add_quantum_error(errors_reset[j], "reset", [j])
noise_thermal.add_quantum_error(errors_measure[j], "measure", [j])
noise_thermal.add_quantum_error(errors_u1[j], "u1", [j])
noise_thermal.add_quantum_error(errors_u2[j], "u2", [j])
noise_thermal.add_quantum_error(errors_u3[j], "u3", [j])
for k in range(n):
noise_thermal.add_quantum_error(errors_cx[j][k], "cx", [j, k])
#print(noise_thermal)
return noise_thermal
# ## Simulation
#
k = 3
sub_qbits = QuantumRegister(n_nodes)
sub_cir = QuantumCircuit(sub_qbits, name="state")
sub_cir, sub_qbits = wn(sub_cir, sub_qbits)
sub_cir.x(sub_qbits)
stat_prep = sub_cir.to_instruction()
inv_stat_prep = sub_cir.inverse().to_instruction()
l_noise = [] #a list to store different number of iterations results
ss = []
N = 2**n_nodes # for iterations count
data_noise =[]
iterations = round(math.pi/4*math.sqrt(N))
for i in range(1,6):
qc, nodes_qubits, edge_anc, ancilla, neg_base, class_bits, tri_flag = grover(i,n_nodes,stat_prep,inv_stat_prep)
qc.measure(nodes_qubits,class_bits)
#Executing circuit and show results
T1 = 500e3 #rochester
T2 = 500e3
noise_thermal = noise_func(len(qc.qubits),T1,T2)
ex = execute(qc, simulator, basis_gates=noise_thermal.basis_gates, noise_model=noise_thermal)
#ex = execute(qc, simulator,basis_gates=noise_bit_flip.basis_gates,noise_model=noise_bit_flip)
res = ex.result()
M = res.get_counts(qc)
print(M)
ss.append(M)
#printing measurements results
s = oq.Measurement(qc,shots=5000)
print(s)
print('============================================')
pattern = "\d+\|\d+\>"
res = re.findall(pattern,s)
dct_noise = {}
for item in res:
if item[item.index("|"):] not in dct_noise.keys():
dct_noise[item[item.index("|"):]] = int(item[:item.index("|")])
else:
dct_noise[item[item.index("|"):]] += int(item[:item.index("|")])
l_noise.append(dct_noise)
h = []
for item in ss:
test_dict = {}
for key in item.keys():
k = '|'+key[4:][::-1]+'>'
test_dict[k]= item[key]
h.append(test_dict)
print(h)
|
Grover with W state GitHub.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Logistic Regression with L2 Regularization
import numpy as np
import pandas as pd
# +
from sklearn.datasets import load_iris
dataset = load_iris()
X = dataset.data
y = dataset.target
target_names = list(dataset.target_names)
print(target_names)
# -
# Change to binary class
y = (y > 0).astype(int)
y
class LogReg:
"""
This implementation of Logistic Regression uses mini-batch gradient descent with l2-regularization.
"""
def __init__(self,
epochs=100,
tolerance = 1e-10,
alpha=0.001,
lambd=0,
threshold=0.5,
verbose=False,
minibatch_size=30,
):
self.epochs = epochs
self.alpha = alpha # Learning rate
self.lambd = lambd # Regularization parameter
self.tolerance = tolerance
self.threshold = threshold
self.verbose = verbose
self.minibatch_size = minibatch_size
def add_ones(self, X):
return np.concatenate((np.ones((len(X),1)), X), axis = 1)
def sigmoid(self, X, theta):
return 1/(1 + np.exp(X@theta))
def cost(self, X, y_true):
m = X.shape[0]
y_hat = self.sigmoid(X, self.theta)
temp_theta = self.theta[:, 1:].copy()
Cost = np.sum(-1*y_true*np.log(y_hat)-(1-y_true)*np.log(1-y_hat)) + self.lambd * np.sum(temp_theta**2)
return Cost
def get_minibatch(self, X, y, minibatch):
X_mb = X[minibatch*self.minibatch_size: (minibatch+1)*self.minibatch_size]
y_mb = y[minibatch*self.minibatch_size: (minibatch+1)*self.minibatch_size]
return X_mb, y_mb
def fit(self, X, y):
X = X.copy()
X = self.add_ones(X)
n, d = X.shape
self.classes = np.unique(y)
self.no_classes = len(self.classes)
# Turn y into one-hot-labels if number of classes is greater than 2
if self.no_classes > 2:
y_encode = np.zeros((n, self.no_classes))
y_encode[range(n), y] = 1 #numpy advanced indexing
y = y_encode
else:
y = y.reshape(-1, 1)
if self.no_classes > 2:
self.theta = np.zeros((d, self.no_classes))
else:
self.theta = np.zeros((d, 1))
current_epoch = 1
norm = 1
no_of_minibatch = int(n/self.minibatch_size)
while (norm >= self.tolerance and current_epoch < self.epochs):
# Shuffle X for minibatch gradient descent
shuffled_index = np.random.permutation(n)
X_shuffled = X[shuffled_index]
y_shuffled = y[shuffled_index]
old_theta = self.theta.copy()
theta_wo_bias = self.theta[:, 1:].copy()
for mb in range(no_of_minibatch):
X_mb, y_mb = self.get_minibatch(X, y, mb)
grad = X_mb.T@(y_mb - self.sigmoid(X_mb, self.theta)) + self.lambd * np.sum(theta_wo_bias)
if self.no_classes <= 2:
grad= grad.reshape(-1, 1)
self.theta = self.theta - self.alpha*grad
if self.verbose and (current_epoch%100 == 0):
print(f'cost for {current_epoch} epoch : {self.cost(X, y)}')
norm = np.linalg.norm(old_theta - self.theta)
current_epoch += 1
return self.theta
def evaluate(self, X, y):
"""
Returns mse loss for a dataset evaluated on the hypothesis
"""
X = self.add_ones(X)
return self.cost(X, y)
def predict(self, X):
proba = self.predict_proba(X)
if self.no_classes > 2:
# Multiclass classification
y_hat = np.argmax(proba, axis=1)
elif self.no_classes == 2:
# Binary classification
y_hat = (proba >= self.threshold).astype(int)
return y_hat
def predict_proba(self, X):
"""
Returns probability of predictions.
"""
X = self.add_ones(X)
return self.sigmoid(X, self.theta)
logreg = LogReg(verbose=True)
logreg.fit(X, y)
predictions = logreg.predict(X)
predictions = predictions.squeeze()
np.sum(y == predictions) / len(y)
predictions
|
Logistic Regression L2Regularization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from os import system
import graphviz #pip install graphviz
from sklearn.cross_validation import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn import tree
carmpg = pd.read_csv("car-mpg.csv")
carmpg.head(5)
columns = carmpg.columns
mask = np.ones(columns.shape, dtype=bool)
i = 0
mask[i] = 0
mask[7] = 0
X = carmpg[columns[mask]]
Y = carmpg["mpg"]
X_train, X_test, y_train, y_test = train_test_split( X, Y, test_size = 0.3, random_state = 100)
clf_gini = tree.DecisionTreeClassifier(criterion = "gini", random_state = 100, max_depth=3, min_samples_leaf=5)
clf_gini.fit(X_train, y_train)
|
Section 10/Decision trees using python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bus
#
# Este bus tiene un sistema de control de subida y bajada de pasajeros para monitorizar el número de ocupantes que lleva y así detectar cuando hay un aforo demasiado alto.
#
# En cada parada la subida y bajada de pasajeros se representa por una tupla compuesta por dos números enteros.
# ```
# bus_stop = (in, out)
# ```
# La sucesión de paradas se representa con una lista estas tuplas.
# ```
# stops = [(in1, out1), (in2, out2), (in3, out3), (in4, out4)]
# ```
#
# ## Objetivos:
# * listas, tuplas
# * bucles while/for
# * mínimo, máximo, longitud
# * media, desviación estandard
#
# ## Tareas
# 1. Calcula el número de paradas.
# 2. Asigna a una variable una lista cuyos elementos sean el número de pasajeros en cada parada (in-out),
# 3. Halla el máximo de ocupación del autobús.
# 4. Calcula la media de la ocupación. Y la desviación estandard.
#
# +
stops = [(10, 0), (4, 1), (3, 5), (3, 4), (5, 1), (1, 5), (5, 8), (4, 6), (2, 3)]
# 1. Calcula el número de paradas.
# 2. Asigna a una variable una lista cuyos elementos sean el número de pasajeros en
# cada parada: Cada elemento depende del elemento previo en la lista + in - out.
# 3. Halla el máximo de ocupación del autobús.
# 4. Calcula la media de la ocupación. Y la desviación estandard.
# -
len(stops)
|
01-intro-101/python/practices/03-bus/bus.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Centerloss
# Centerloss is a loss that can minimize the intra-class distance
# I am proposing the use of centerloss in the 2-layer-network.
#
# Where the Hidden Layer Unit is set to be 3, so that we can visualize it.
# In this extra task, I am trying to compare the difference between softmax loss and centerloss
#
# We fixed these params to compare the loss.
# input_size,hidden_size,num_classes,num_iters,batch_size
# +
import numpy as np
import matplotlib.pyplot as plt
import time
from cs231n.classifiers.neural_net import TwoLayerNet
from cs231n.classifiers import LinearClassifier
# %matplotlib inline
#plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
#plt.rcParams['image.interpolation'] = 'nearest'
#plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
# -
# Load Data
# +
from cs231n.data_utils import load_CIFAR10
def get_CIFAR10_data(num_training=49000, num_validation=1000, num_test=1000):
"""
Load the CIFAR-10 dataset from disk and perform preprocessing to prepare
it for the two-layer neural net classifier. These are the same steps as
we used for the SVM, but condensed to a single function.
"""
# Load the raw CIFAR-10 data
cifar10_dir = 'cs231n/datasets/cifar-10-batches-py'
X_train, y_train, X_test, y_test = load_CIFAR10(cifar10_dir)
# Subsample the data
mask = list(range(num_training, num_training + num_validation))
X_val = X_train[mask]
y_val = y_train[mask]
mask = list(range(num_training))
X_train = X_train[mask]
y_train = y_train[mask]
mask = list(range(num_test))
X_test = X_test[mask]
y_test = y_test[mask]
# Normalize the data: subtract the mean image
mean_image = np.mean(X_train, axis=0)
X_train -= mean_image
X_val -= mean_image
X_test -= mean_image
# Reshape data to rows
X_train = X_train.reshape(num_training, -1)
X_val = X_val.reshape(num_validation, -1)
X_test = X_test.reshape(num_test, -1)
return X_train, y_train, X_val, y_val, X_test, y_test
# Invoke the above function to get our data.
X_train, y_train, X_val, y_val, X_test, y_test = get_CIFAR10_data()
print('Train data shape: ', X_train.shape)
print('Train labels shape: ', y_train.shape)
print('Validation data shape: ', X_val.shape)
print('Validation labels shape: ', y_val.shape)
print('Test data shape: ', X_test.shape)
print('Test labels shape: ', y_test.shape)
# +
from mpl_toolkits.mplot3d import Axes3D
import matplotlib
def show_features(X_test,y_test,net):
# color_map = np.array([[0,0,0],
# [255,0,0],
# [0,255,0],
# [0,0,255],
# [255,255,0],
# [0,255,255],
# [255,0,255],
# [128,128,128],
# [128,0,128],
# [0,128,0]])
color_map = ["#000000","#FF0000","#00FF00","#0000FF","#FFFF00","#00FFFF","#FF00FF","#808080","#008000","#000080"]
# get the first layer output ( after reLu)
W1 = net.params['W1']
b1 = net.params['b1']
layer1 = X_val.dot(W1) + b1.reshape(1, -1)
out = np.maximum(0, layer1)
# out shpae (N,3)
fig = plt.figure(figsize=(18, 16), dpi= 80)
# ax = fig.add_subplot(111, projection='3d')
ax = fig.add_subplot(111)
for i in range(10):
class_i_out = out[y_val==i,:]
# ax.scatter(class_i_out[:,0],class_i_out[:,1],class_i_out[:,2],c=color_map[i],marker='.')
ax.scatter(class_i_out[:,0],class_i_out[:,1],c=color_map[i],marker='.')
plt.show()
return
# -
# Using Softmaxloss
# +
input_size = 32 * 32 * 3
hidden_size = 2
num_classes = 10
num_iters=1000
batch_size=400
learning_rate = 1e-3
reg = 0.3
net = TwoLayerNet(input_size, hidden_size, num_classes)
# Train the network
start_time = time.time()
stats = net.train(X_train, y_train, X_val, y_val,
num_iters=num_iters, batch_size=batch_size,
learning_rate=learning_rate, learning_rate_decay=0.95,
reg=reg, verbose=True)
print('Elasped Time :{:.3f}'.format(time.time()-start_time))
y_test_pred = net.predict(X_test)
acc = np.mean(y_test == y_test_pred)
print("accuracy =",acc)
# -
# show features
show_features(X_test,y_test,net)
# Centerloss
# +
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
from past.builtins import xrange
class Center_loss_net(object):
def __init__(self, input_size, hidden_size, output_size, std=1e-4):
self.params = {}
self.params['W1'] = std * np.random.randn(input_size, hidden_size)
self.params['b1'] = np.zeros(hidden_size)
self.params['W2'] = std * np.random.randn(hidden_size, output_size)
self.params['b2'] = np.zeros(output_size)
def loss(self, X, y=None, reg=0.0, ratio=0.5):
# Unpack variables from the params dictionary
W1, b1 = self.params['W1'], self.params['b1']
W2, b2 = self.params['W2'], self.params['b2']
N, D = X.shape
C = W2.shape[1]
H = W1.shape[1]
scores = None
# X: (N, D)
# W1: (D, H)
# b1: (H,)
# W2: (H, C)
# b2: (C,)
layer1 = X.dot(W1) + b1.reshape(1, -1)
reLu = np.maximum(0, layer1)
layer2 = reLu.dot(W2) + b2
# layer1: (N,H), relu: (N,H), layer2: (N,C)
scores = layer2
if y is None:
return scores
# Compute the loss
loss = None
# score: (N,C), softmax: (N,C)
exp_scores = np.exp(scores)
softmax = exp_scores[np.arange(N), y] / np.sum(exp_scores, axis=1)
loss = np.average(-np.log(softmax)) + reg * (np.sum(W1 * W1) + np.sum(W2 * W2))
### Center loss here
# cal center, shape = (C,H)
center_loss = 0
centers = np.zeros((C,H))
diff = np.zeros((N,H))
Centers = np.zeros((N,H))
for i in range(C):
reLu_i = reLu[y==i,:]
centers[i] = np.mean(reLu_i,axis = 0)
for i in range(N):
diff[i] = reLu[i]-centers[y[i]]
# print(diff[i])
# center_loss +=(np.sum(diff[i])) # L1 distance
center_loss = np.sum(diff)/(N)
# print(center_loss)
loss += ratio * center_loss
# Backward pass: compute gradients
grads = {}
#############################################################################
# TODO: Compute the backward pass, computing the derivatives of the weights #
# and biases. Store the results in the grads dictionary. For example, #
# grads['W1'] should store the gradient on W1, and be a matrix of same size #
#############################################################################
# D_softmax wrt S, shape = (N,C)
D_center = np.ones_like(diff)
D_center[np.where(reLu <= 0)] = 0
DCDW = np.zeros((C,D))
for i in range(C):
a = X
a[np.where(reLu <= 0)] = 0
a=a[y==i]
DCDW[i] = 1/N * np.sum(a,axis=0)
# bb = np.ones((N,))
# bb[np.where(reLu <= 0)] = 0
DXDW_ = np.zeros_like(X)
for i in range(N):
DXDW_[i] = DCDW[y[i]]
D_center_W1 = np.dot(X.T-DXDW_.T,D_center)
D_center_W1 = D_center_W1*ratio / N
D_center_b1 = np.dot(np.ones((N,)),D_center)
D_center_b1 = D_center_b1*ratio/N
D_softmax = exp_scores / np.sum(exp_scores, axis=1).reshape(-1, 1)
D_softmax[np.arange(N), y] -= 1
D_W2 = np.dot(reLu.T, D_softmax)
grads['W2'] = D_W2 / N + 2 * reg * W2
D_b2 = np.dot(np.ones(N).T, D_softmax)
grads['b2'] = D_b2 / N
# D_ReLu wrt ReLu, shape = (N,H)
D_reLu = np.dot(D_softmax, W2.T)
D_reLu[np.where(reLu <= 0)] = 0
D_W1 = X.T.dot(D_reLu)
grads['W1'] = D_W1 / N + 2 * reg * W1 + D_center_W1
D_b1 = np.dot(np.ones(N), D_reLu)
grads['b1'] = D_b1 / N + D_center_b1
#############################################################################
# END OF YOUR CODE #
#############################################################################
return loss, grads
def train(self, X, y, X_val, y_val,
learning_rate=1e-3, learning_rate_decay=0.95,
reg=5e-6, num_iters=100,
batch_size=200, verbose=False):
"""
"""
num_train = X.shape[0]
iterations_per_epoch = max(num_train / batch_size, 1)
# Use SGD to optimize the parameters in self.model
loss_history = []
train_acc_history = []
val_acc_history = []
for it in xrange(num_iters):
X_batch = None
y_batch = None
#########################################################################
# TODO: Create a random minibatch of training data and labels, storing #
# them in X_batch and y_batch respectively. #
#########################################################################
indices = np.random.choice(X.shape[0], batch_size, replace=True)
X_batch = X[indices, :]
y_batch = y[indices]
#########################################################################
# END OF YOUR CODE #
#########################################################################
# Compute loss and gradients using the current minibatch
loss, grads = self.loss(X_batch, y=y_batch, reg=reg)
loss_history.append(loss)
#########################################################################
# TODO: Use the gradients in the grads dictionary to update the #
# parameters of the network (stored in the dictionary self.params) #
# using stochastic gradient descent. You'll need to use the gradients #
# stored in the grads dictionary defined above. #
#########################################################################
for k in self.params:
self.params[k] += - learning_rate * grads[k]
#########################################################################
# END OF YOUR CODE #
#########################################################################
if verbose and it % 100 == 0:
print('iteration %d / %d: loss %f' % (it, num_iters, loss))
# Every epoch, check train and val accuracy and decay learning rate.
if it % iterations_per_epoch == 0:
# Check accuracy
train_acc = (self.predict(X_batch) == y_batch).mean()
val_acc = (self.predict(X_val) == y_val).mean()
train_acc_history.append(train_acc)
val_acc_history.append(val_acc)
# Decay learning rate
learning_rate *= learning_rate_decay
return {
'loss_history': loss_history,
'train_acc_history': train_acc_history,
'val_acc_history': val_acc_history,
}
def predict(self, X):
"""
Use the trained weights of this two-layer network to predict labels for
data points. For each data point we predict scores for each of the C
classes, and assign each data point to the class with the highest score.
Inputs:
- X: A numpy array of shape (N, D) giving N D-dimensional data points to
classify.
Returns:
- y_pred: A numpy array of shape (N,) giving predicted labels for each of
the elements of X. For all i, y_pred[i] = c means that X[i] is predicted
to have class c, where 0 <= c < C.
"""
y_pred = None
###########################################################################
# TODO: Implement this function; it should be VERY simple! #
###########################################################################
scores = self.loss(X)
y_pred = np.argmax(scores,axis=1)
###########################################################################
# END OF YOUR CODE #
###########################################################################
return y_pred
# +
input_size = 32 * 32 * 3
hidden_size = 2
num_classes = 10
num_iters=1000
batch_size=200
learning_rate = 1e-5
reg = 3000
clnet = Center_loss_net(input_size, hidden_size, num_classes)
# Train the network
start_time = time.time()
stats = clnet.train(X_train, y_train, X_val, y_val,
num_iters=num_iters, batch_size=batch_size,
learning_rate=learning_rate, learning_rate_decay=0.95,
reg=reg, verbose=True)
print('Elasped Time :{:.3f}'.format(time.time()-start_time))
y_test_pred = net.predict(X_test)
acc = np.mean(y_test == y_test_pred)
print("accuracy =",acc)
# -
# show features
show_features(X_test,y_test,net)
|
assignment1/Extra.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="V8JDmA21FffQ" colab_type="code" outputId="978af748-2c21-4e6e-c480-1411dcbdcf22" executionInfo={"status": "ok", "timestamp": 1591926776186, "user_tz": 300, "elapsed": 20939, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GinuX1vGa2YjjYRYk0K3BaUnB028-gIIiyo76XGFw=s64", "userId": "05082376392541668854"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
message_to_encrypted = bytes(input(), 'utf-8')
# + id="it_zhoTO-c3L" colab_type="code" outputId="134ea522-3080-4016-cce7-6cb6d4eda674" executionInfo={"status": "ok", "timestamp": 1591926812003, "user_tz": 300, "elapsed": 32255, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GinuX1vGa2YjjYRYk0K3BaUnB028-gIIiyo76XGFw=s64", "userId": "05082376392541668854"}} colab={"base_uri": "https://localhost:8080/", "height": 593}
pip install eciespy
# + id="O74ORzFgAFNL" colab_type="code" colab={}
from ecies import encrypt
# + id="K0RLI3HgAK43" colab_type="code" outputId="d4596865-1c59-4cbb-9e3b-ba1ca769a765" executionInfo={"status": "ok", "timestamp": 1591926838954, "user_tz": 300, "elapsed": 21144, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GinuX1vGa2YjjYRYk0K3BaUnB028-gIIiyo76XGFw=s64", "userId": "05082376392541668854"}} colab={"base_uri": "https://localhost:8080/", "height": 124}
from google.colab import drive
drive.mount('/content/drive')
# + id="h9NyKHoMAK8X" colab_type="code" colab={}
with open("/content/drive/My Drive/Colab Notebooks/ECC/public_key/pub", "r") as public_key:
public = public_key.read()
# + id="2970LiUQALDL" colab_type="code" colab={}
encripted_msg = encrypt(public, message_to_encrypted)
# + id="_1JXLZEQAzIM" colab_type="code" colab={}
with open("/content/drive/My Drive/Colab Notebooks/ECC/responses/response", "wb") as response:
response.write(encripted_msg)
# + id="4pcA_BMsBGgJ" colab_type="code" colab={}
|
Elliptic Curve Cryptography August 2020/ECC Reponse Example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Instructions
#
# To start, go to Kernal -> 'Restart and Run All' -> 'Restart and Run All Cells'
#
# replace the wallet address in the cell below with the address you want to analyse
# Bearwhale wallet v2 : <KEY>
# v1 : 9hyDXH72HoNTiG2pvxFQwxAhWBU8CrbvwtJDtnYoa4jfpaSk1d3
target = '<KEY>'
# # Imports
# #!pip install pandas
import pandas as pd
import requests
pd.set_option('display.float_format', lambda x: '%.3f' % x)
# # Pull
#
# API Request to pull the data and then place it into a pandas dataframe. You can analyse the json directly by uncommenting these lines below.
# +
# API Request
# Docs : https://api.ergoplatform.com/api/v1/docs/
import requests
for p in range(1,50):
url = 'https://api.ergoplatform.com/api/v1/boxes/byAddress/<KEY>'+'?page='+str(p)
r = requests.get(url)
json = r.json()
# +
# Query the json directly
#json.keys()
#json['items']
json['total']
#json
# -
# load json into a dataframe
df = pd.DataFrame(json['items'])
df
# # Dataframe
#
# Pandas allows us to view the information easier, we drop the columns we don't need to here to clear up clutter.
# +
df.info()
# boxId
# transactionId - tx id displayed on explorer
# blockId
# value - unspent
# index - ?
# creationHeight - time
# ergoTree - 0008cd023c166f1ba9de6fdd5ed90a6f15e037b0a9247a...
# address - wallet address
# assets - tokens
# additionalRegisters - sigma? R4
# spentTxId
# mainChain - main-net - True
#df.sort_values(by=['BalanceChange (ERG)'])
df
# +
# Drop unneeded columns
df.drop(columns=['mainChain','ergoTree','transactionId','boxId',
'blockId','address','index',
'additionalRegisters','spentTransactionId'], inplace=True)
# -
# # Tidy
#
# The UTXO model tracks the total balance on each transaction in nanoErgs. Some small fixes to make it more readable.
# Finding difference
df['BalanceChange (ERG)'] = df.value.diff()
df['HeightChange (h)'] = df.creationHeight.diff()
# nanoErg -> Erg
df['value'] = df['value'] / 1000000000
df['BalanceChange (ERG)'] = df['BalanceChange (ERG)'] / 1000000000
# Track this from geneis block ?
df['HeightChange (h)'] = df['HeightChange (h)'] * 2
df['HeightChange (h)'] = df['HeightChange (h)'] / 60
# Reversing polarity
df['BalanceChange (ERG)'] = df['BalanceChange (ERG)'] * -1
df['HeightChange (h)'] = df['HeightChange (h)'] * -1
df['BalanceChange (ERG)'] = df['BalanceChange (ERG)'].shift(-1)
df['HeightChange (h)'] = df['HeightChange (h)'].shift(-1)
# # Tokens
# +
# Pulling SigmaUSD out asset tokens
df3 = df['assets'].apply(pd.Series)
df4 = df3[0].apply(pd.Series)
df4.drop(columns=['tokenId','index','decimals','type',
], inplace=True)
# Merging back into original frame and tidying
df = pd.concat([df, df4], axis=1)
df.drop(columns=['assets'], inplace=True)
df = df.rename(columns={'value': 'Final Balance', 'amount': 'Sigma', 'name': 'type'})
# -
pd.set_option('display.max_rows', 30)
df.sort_values(by=['BalanceChange (ERG)'], ascending=False)
df
df['BalanceChange (ERG)'].sum()
import numpy as np
df['Final Balance'].sum()
df['Sigma'] = df['Sigma'].replace(np.nan, 0)
df
# +
df['Sigma Minted'] = df['Sigma'].diff(periods=-1)
df
# -
df['Price Paid'] = df['Sigma Minted'] / (df['BalanceChange (ERG)'] * -1)
df['Price Paid'] = df['Price Paid'] / 100
df
|
wallet-query.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## A model build using TweetBERT to identify cause-effect pairs in sentences of tweets
# The cause-effect pair sentence prediction model will be trained on the dataset which got augmented in an active learning approach in 5 steps.
#
# +
import pandas as pd
import numpy as np
import spacy
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, precision_recall_fscore_support, matthews_corrcoef
from transformers import BertForSequenceClassification, AutoTokenizer
from transformers import AdamW, get_linear_schedule_with_warmup
from tqdm import tqdm, trange
import random
import os
import torch.nn.functional as F
import torch
from torch.nn import CrossEntropyLoss
from torch.utils.data import DataLoader
import transformers
from tqdm import tqdm, trange
from utils import normalizeTweet, split_into_sentences, bio_tagging, EarlyStopping
import matplotlib.pyplot as plt
########################### Check if cuda available ############################
print("Cuda available: ", torch.cuda.is_available())
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
########################### DATA FILE ###################################
# dataPath = "/home/adrian/workspace/causality/Causal-associations-diabetes-twitter/data/Causality_tweets_data.xlsx"
dataPath = "data/Causality_tweets_data.xlsx"
#dataPath = "Causality_tweets_data.xlsx"
########################### MODEL PARAMETERS ############################
lr = 1e-3
adam_eps = 1e-8
epochs = 35
num_warmup_steps = 0
early_patience = 5 # how long to wait after last time validation loss improved
train_batch_size = 16
val_batch_size = 16
test_batch_size = 32
train_to_test_ratio = 0.9 # 10% test and 90% train
val_to_train_ratio = 0.2
metrics_average = "binary" # this will give measure for class_1,i.e., causal class
saveModelName = "./model-causal-model/NER_model_1_finetuned-{}-epochs-lr_{}.pth".format(epochs, lr) # it should be epoch so that the name shows at what epoch teh mdel ws saved
# +
##### DATA TO LOAD ######
data_round0 = pd.read_excel(dataPath, sheet_name="round0")
data_round0 = data_round0[data_round0["Causal association"].notnull()] # some tweets at the end are not labeled yet
data_round0 = data_round0[["full_text", "Intent", "Cause", "Effect", "Causal association"]]
print("Data round 0 (tweets!):")
print(data_round0["Causal association"].value_counts())
print("-----"*5)
##### additional data labeled through active learning strategy - round 1 ########
data_round1 = pd.read_excel(dataPath, sheet_name="round1")
data_round1 = data_round1[data_round1["Causal association"].notnull()]
data_round1 = data_round1[["sentence", "Intent", "Cause", "Effect", "Causal association"]]
data_round1.rename(columns ={"sentence":"full_text"}, inplace=True) # rename for merge
print("Sentences round 1:")
print(data_round1["Causal association"].value_counts())
print("-----"*5)
##### additional data labeled through active learning strategy - round 2 ########
data_round2 = pd.read_excel(dataPath, sheet_name="round2")
data_round2 = data_round2[data_round2["Causal association"].notnull()]
data_round2 = data_round2[["sentence", "Intent", "Cause", "Effect", "Causal association"]]
data_round2.rename(columns ={"sentence":"full_text"}, inplace=True) # rename for merge
print("sentences round 2:")
print(data_round2["Causal association"].value_counts())
print("-----"*5)
##### additional data labeled through active learning strategy - round 3 ########
data_round3 = pd.read_excel(dataPath, sheet_name="round3")
data_round3 = data_round3[data_round3["Causal association"].notnull()]
data_round3 = data_round3[["sentence", "Intent", "Cause", "Effect", "Causal association"]]
data_round3.rename(columns ={"sentence":"full_text"}, inplace=True) # rename for merge
print("sentences round 3:")
print(data_round3["Causal association"].value_counts())
print("-----"*5)
##### additional data labeled through active learning strategy - round 4 ########
data_round4 = pd.read_excel(dataPath, sheet_name="round4")
data_round4 = data_round4[data_round4["Causal association"].notnull()]
data_round4 = data_round4[["sentence", "Intent", "Cause", "Effect", "Causal association"]]
data_round4.rename(columns ={"sentence":"full_text"}, inplace=True) # rename for merge
print("sentences round 3:")
print(data_round3["Causal association"].value_counts())
#### merge both datasets ######
data = data_round0.append(data_round1).append(data_round2).append(data_round3).append(data_round4)
print("\nAfter merge old data:")
print(data["Causal association"].value_counts())
data.head()
# -
# ## Preprocessing
# +
################## add BIO tags and tokenized tweets ################
data["tokenized"] = data["full_text"].map(lambda tweet: normalizeTweet(tweet).split(" "))
data["bio_tags"] = data.apply(lambda row: bio_tagging(row["full_text"],row["Cause"], row["Effect"]), axis=1)
data.head(n=20)
# +
##### TEST to see on a random subsample which tokens get associated to which bio tag #######
#for i, row in data[data["Causal association"] == 1].sample(n=100).iterrows():
# print()
# print(row["Causal association"], "\tCause:",row["Cause"], "\tEffect:", row["Effect"])
# for tok, tag in zip(row["tokenized"], row["bio_tags"]):
# print(tok,tag)
# +
def get_start_end_index_of_sentence_in_tweet(tweet, sentence):
"""
The sentence tokens are included in the tweet tokens.
Return the start end end indices of the sentence tokens in the tweet tokens
"""
sentence_start_word = sentence[0]
start_indices = [i for i, x in enumerate(tweet) if x == sentence_start_word] # find all indices of the start word of the sentence
try:
for start_index in start_indices:
isTrueStartIndex = all([tweet[start_index+i] == sentence[i] for i in range(len(sentence))])
#print("start_index:", start_index, "isTrueStartIndex:", isTrueStartIndex)
if isTrueStartIndex:
return start_index, start_index + len(sentence)
except:
print("ERROR: StartIndex should have been found for sentence:")
print("tweet:")
print(tweet)
print("sentence:")
print(sentence)
return -1, -2 # should not be returned
def split_tweets_to_sentences(data):
"""
Splits tweets into sentences and associates the appropriate intent, causes, effects and causal association
to each sentence.
Parameters:
- min_words_in_sentences: Minimal number of words in a sentence such that the sentence is kept.
Assumption: A sentence with too few words does not have enough information
Ex.:
full_text | Intent | Cause | Effect | Causal association | ...
--------------------------------------------------------------------------------------------
what? type 1 causes insulin dependence | q;msS | type 1|insulin dependence | 1 | ...
New dataframe returned:
full_text | Intent | Cause | Effect | Causal association | ...
--------------------------------------------------------------------------------------------
what? | q | | | 0 | ...
type 1 causes insulin dependence | | type 1| insulin dependence | 1 | ...
"""
newDF = pd.DataFrame(columns=["sentence", "Intent", "Cause", "Effect", "Causal association", "tokenized", "bio_tags"])
for i,row in data.iterrows():
causes = row["Cause"]
effects = row["Effect"]
sentences = split_into_sentences(normalizeTweet(row["full_text"]))
# single sentence in tweet
if len(sentences) == 1:
singleSentenceIntent = ""
if isinstance(row["Intent"], str):
if len(row["Intent"].split(";")) > 1:
singleSentenceIntent = row["Intent"].strip().replace(";msS", "").replace("msS;", "").replace(";mS", "").replace("mS;", "")
else:
if row["Intent"] == "mS" or row["Intent"] == "msS":
singleSentenceIntent = ""
else:
singleSentenceIntent = row["Intent"].strip()
newDF=newDF.append(pd.Series({"sentence": sentences[0] # only one sentence
, "Intent": singleSentenceIntent
, "Cause" : row["Cause"]
, "Effect": row["Effect"]
, "Causal association" : row["Causal association"]
, "tokenized": row["tokenized"]
, "bio_tags": row["bio_tags"]}), ignore_index=True)
# tweet has several sentences
else:
intents = str(row["Intent"]).strip().split(";")
for sentence in sentences:
sent_tokenized = sentence.split(" ")
causeInSentence = np.nan if not isinstance(causes, str) or not any([cause in sentence for cause in causes.split(";")]) else ";".join([cause for cause in causes.split(";") if cause in sentence])
effectInSentence = np.nan if not isinstance(effects, str) or not any([effect in sentence for effect in effects.split(";")]) else ";".join([effect for effect in effects.split(";") if effect in sentence])
causalAssociationInSentence = 1 if isinstance(causeInSentence, str) and isinstance(effectInSentence, str) else 0
startIndex, endIndex = get_start_end_index_of_sentence_in_tweet(row["tokenized"], sent_tokenized)
sentence_tokenized = row["tokenized"][startIndex:endIndex]
sentence_bio_tags = row["bio_tags"][startIndex:endIndex]
if "q" in intents and sentence[-1] == "?": # if current sentence is question
newDF=newDF.append(pd.Series({"sentence": sentence, "Intent": "q", "Cause" : causeInSentence
, "Effect": effectInSentence, "Causal association" : causalAssociationInSentence
, "tokenized": sentence_tokenized, "bio_tags": sentence_bio_tags}), ignore_index=True)
elif "joke" in intents: # all sentences with "joke" in tweet keep the intent "joke"
newDF=newDF.append(pd.Series({"sentence": sentence, "Intent": "joke", "Cause" : causeInSentence
, "Effect": effectInSentence, "Causal association" : causalAssociationInSentence
, "tokenized": sentence_tokenized, "bio_tags": sentence_bio_tags}), ignore_index=True)
elif "neg" in intents: # all sentences with "neg" in tweet keep intent "neg"
newDF=newDF.append(pd.Series({"sentence": sentence, "Intent": "neg", "Cause" : causeInSentence
, "Effect": effectInSentence, "Causal association" : causalAssociationInSentence
, "tokenized": sentence_tokenized, "bio_tags": sentence_bio_tags}), ignore_index=True)
elif isinstance(causeInSentence, str) and isinstance(effectInSentence, str): # cause effect sentence
causalIntent = ""
if len(causeInSentence.split(";")) > 1:
causalIntent = "mC"
if len(effectInSentence.split(";")) > 1:
causalIntent = "mC;mE"
elif len(effectInSentence.split(";")) > 1:
causalIntent = "mE"
newDF=newDF.append(pd.Series({"sentence": sentence, "Intent": causalIntent, "Cause" : causeInSentence
, "Effect": effectInSentence, "Causal association" : causalAssociationInSentence
, "tokenized": sentence_tokenized, "bio_tags": sentence_bio_tags}), ignore_index=True)
else:
nonCausalIntent = ""
if isinstance(causeInSentence, str): # only cause is given
if len(causeInSentence.split(";")) > 1:
nonCausalIntent = "mC"
elif isinstance(effectInSentence, str): # only effect is given
if len(effectInSentence.split(";")) > 1:
nonCausalIntent = "mE"
newDF=newDF.append(pd.Series({"sentence": sentence, "Intent": nonCausalIntent, "Cause" : causeInSentence
, "Effect": effectInSentence, "Causal association" : causalAssociationInSentence
, "tokenized": sentence_tokenized, "bio_tags": sentence_bio_tags}), ignore_index=True)
return newDF
# +
### Split tweets into sentences (train classifier on sentence level) ####
print("N tweets:", data.shape[0])
dataSentences = split_tweets_to_sentences(data)
print("N sentences:", dataSentences.shape[0])
dataSentences.head()
# +
########## Remove sentences with joke, question, negation and keep only sentences with more than 3 tokens #####
print("N sentences before filtering: ", dataSentences.shape[0])
dataSentFiltered = dataSentences[~dataSentences["Intent"].str.contains("neg|joke|q")] # remove sentences with joke, q, neg
dataSentFiltered = dataSentFiltered[dataSentFiltered["tokenized"].map(len) >= 3] # only keep sentences with at least 3 words
print("N sentences after filtering: ", dataSentFiltered.shape[0])
dataSentFiltered.head()
# +
############ Choose causal tweets to train on ##################
# choose sentences with cause and effect
#trainingData = dataSentFiltered[dataSentFiltered["Causal association"] == 1]
# choose sentences with cause or effect
trainingData = dataSentFiltered[(dataSentFiltered["Cause"].notnull()) | (dataSentFiltered["Effect"].notnull())]
trainingData.shape
# -
# ### Training
trainingDataSample = trainingData#.sample(n=200) # VIVEK: DELETE TAKING SAMPLE. THIS WAS ONLY FOR TESTING
train = trainingDataSample.sample(frac=train_to_test_ratio, random_state=0)
test = trainingDataSample.drop(train.index)
validate = train.sample(frac=val_to_train_ratio, random_state=0)
train = train.drop(validate.index)
print("Train:", train.shape)
print("Validate:", validate.shape)
print("Test:", test.shape)
# +
# Transform labels + encodings into Pytorch DataSet object (including __len__, __getitem__)
class TweetDataSet(torch.utils.data.Dataset):
def __init__(self, text, labels, bio_tags, tokenizer):
self.text = text
self.labels = labels
self.tokenizer = tokenizer
self.bio_tags = bio_tags
self.tag2id = {label: idx for idx, label in enumerate(["O", "B-C", "I-C", "B-E", "I-E"])}
self.tag2id[-100] = -100
self.id2tag = {id:tag for tag,id in self.tag2id.items()}
def __getitem__(self, idx):
inputs = self.tokenizer(self.text, padding=True, truncation=True, return_token_type_ids=True)
ids = inputs["input_ids"]
mask = inputs["attention_mask"]
token_type_ids = inputs["token_type_ids"]
bio_tags_extended = self.extend_tags(self.text[idx], self.bio_tags[idx], ids[idx])
assert(len(ids[idx]) == len(bio_tags_extended), "token ids and BIO tags lengths do not match!")
return {
"input_ids" : torch.tensor(ids[idx], dtype=torch.long)
, "attention_mask" : torch.tensor(mask[idx], dtype=torch.long)
, "token_type_ids" : torch.tensor(token_type_ids[idx], dtype=torch.long)
, "labels" : torch.tensor(self.labels[idx], dtype=torch.long)
, "bio_tags" : torch.tensor(list(map(lambda bioTags: self.tag2id[bioTags], bio_tags_extended))
, dtype=torch.long)
}
def __len__(self):
return len(self.labels)
def extend_tags(self, tokens_old, tags_old, ids_tokenized_padded):
"""
Each token has a BIO tag label.
However BERT's tokenization splits tokens into subwords. How to label those subwords?
Option 1:
---------
add the same label to each subword than the first subword. Only replace "B" by "I"
Ex.
#lowbloodsugar => '#low@@', 'blood@@', 'sugar@@'
"B-C" => "B-C" , "I-C" , "I-C"
Option 2 (implemented):
---------
From : https://huggingface.co/transformers/custom_datasets.html#token-classification-with-w-nut-emerging-entities
A common obstacle with using pre-trained models for token-level classification: many of the tokens in
the W-NUT corpus are not in DistilBert’s vocabulary. Bert and many models like it use a method called
WordPiece Tokenization, meaning that single words are split into multiple tokens such that each token
is likely to be in the vocabulary. For example, DistilBert’s tokenizer would split the Twitter
handle @huggingface into the tokens ['@', 'hugging', '##face']. This is a problem for us because we
have exactly one tag per token. If the tokenizer splits a token into multiple sub-tokens, then we will
end up with a mismatch between our tokens and our labels.
One way to handle this is to only train on the tag labels for the first subtoken of a split token.
We can do this in 🤗 Transformers by setting the labels we wish to ignore to -100.
In the example above, if the label for @HuggingFace is 3 (indexing B-corporation), we would set
the labels of ['@', 'hugging', '##face'] to [3, -100, -100].
"""
tags = [-100] # add for start token <CLS>
for token_old, tag in zip(tokens_old.split(" "), tags_old):
# print(F"\ntoken_old: {token_old}; tag: {tag}")
for i, sub_token in enumerate(self.tokenizer.tokenize(token_old)):
if (i == 0):
tags.append(tag)
else:
tags.append(-100)
tags.append(-100) # 0 for end of sentence token
# append -100 for all padded elements
padded_elements = ids_tokenized_padded.count(1) # id 1 is <PAD> ; Alternative: where attention_mask == 0 add -100
tags.extend([-100]*padded_elements)
return tags
tokenizer = AutoTokenizer.from_pretrained("vinai/bertweet-base")
train_dataset = TweetDataSet(train["sentence"].values.tolist()
, train["Causal association"].values.tolist()
, train["bio_tags"].values.tolist()
, tokenizer)
val_dataset = TweetDataSet(validate["sentence"].values.tolist()
, validate["Causal association"].values.tolist()
, validate["bio_tags"].values.tolist()
, tokenizer)
test_dataset = TweetDataSet(test["sentence"].values.tolist()
, test["Causal association"].values.tolist()
, test["bio_tags"].values.tolist()
, tokenizer)
print(len(train_dataset))
print(len(val_dataset))
print(len(test_dataset))
# put data to batches
train_loader = DataLoader(train_dataset, batch_size=16, shuffle=True)
validation_loader = DataLoader(val_dataset, batch_size=8, shuffle=True)
test_loader = DataLoader(test_dataset, batch_size=8, shuffle=True)
# +
############ class weights #####################
# Since -100 will anyway be ignored by loss func, drop it
#train_bio_tags = np.hstack([t["bio_tags"][t["bio_tags"]!=-100].numpy() for t in train_dataset])
#print("train_bio_tags:", train_bio_tags)
#train_bio_tags_count_info = (pd.Series(train_bio_tags).value_counts(normalize=True))
#print("train_bio_tags_count_info:", train_bio_tags_count_info)
# for class-imbalanced dataset, the class weight for a ith class
# to be specified for balancing in the loss function is given by:
# weight[i] = num_samples / (num_classes * num_samples[i])
# since train_bio_tags_count_info obtained above has fraction of
# samples for ith class, hence the corresponding weight calculation is:
#class_weight = (1/train_bio_tags_count_info)/len(train_bio_tags_count_info)
#class_weight = class_weight[sorted(class_weight.index)]
#class_weight
# -
# ### Evaluation Metrics
# +
from sklearn.metrics import accuracy_score, precision_recall_fscore_support
def compute_metrics(pred, labels):
"""
Dataset is unbalanced -> measure weighted metrics
Calculate metrics for each label, and find their average wieghted by support (Number of true instances for each label)
This alters 'macro' to account for label imbalance;
it can result in an F-Score taht is not between precision and recall
"""
precision, recall, f1, _ = precision_recall_fscore_support(labels, pred, average='macro') # TODO: check weightin
acc = accuracy_score(labels, pred)
return {
'accuracy': acc,
'f1': f1,
'precision': precision,
'recall': recall
}
# -
# ### Model definition
class CausalNER(torch.nn.Module):
""" Model Bert"""
def __init__(self):
super(CausalNER, self).__init__()
self.num_labels = 5 # B-C, I-C, B-E, I-E, O
self.bert = transformers.BertModel.from_pretrained("vinai/bertweet-base")
self.dropout = torch.nn.Dropout(0.3)
self.linear1 = torch.nn.Linear(768, 256)
self.linear2 = torch.nn.Linear(256, self.num_labels)
self.softmax = torch.nn.Softmax(-1)
def forward(self, input_ids, attention_mask, token_type_ids):
# _, output_1 = self.bert(input_ids, attention_mask = attention_mask, token_type_ids=token_type_ids, return_dict=False) # if output 1 is our cls token
output_seq, _ = self.bert(input_ids, attention_mask = attention_mask, token_type_ids=token_type_ids, return_dict=False) # if output 1 is our cls token
output_2 = self.dropout(output_seq)
output_3 = self.linear1(output_2)
output_4 = self.dropout(output_3)
output_5 = self.linear2(output_4)
return output_5 # 5
# +
model = CausalNER()
model.to(device)
# fine-tune only the task-specific parameters
for param in model.bert.parameters():
param.requires_grad = False
# param[:-1].requires_grad = False
# param[-1].requires_grad = True
num_training_steps = np.ceil(len(train_dataset)/train_batch_size)*epochs
optim = AdamW(model.parameters(), lr=lr, eps=adam_eps)
# scheduler with a linearly decreasing learning rate from the initial lr set in the optimizer to 0;
# after a warmup period during which it increases linearly from to the initial lr set in the optimizer
scheduler = get_linear_schedule_with_warmup(optim, num_warmup_steps=num_warmup_steps, num_training_steps=num_training_steps)
## adding weight to the loss function
#loss_fn = CrossEntropyLoss(
# weight=torch.tensor(class_weight.to_list()).to(device),
# ignore_index=-100 # ignore subwords/tokens with label -100
#)
loss_fn = CrossEntropyLoss(ignore_index=-100)
# + tags=[]
################ TRAINING ######################
# initialise the early_stopping object
early_stopping = EarlyStopping(patience=early_patience, path=saveModelName, verbose=True)
train_avg_loss = [] # avg training loss per epoch
val_avg_loss = [] # avg validation loss per epoch
train_avg_acc = [] # avg training accuracy per epoch
val_avg_acc = [] # avg val accuracy per epoch
n_trained_epochs = 0
N_bio_tags = 5 # "O", "B-C", "I-C", "B-E", "I-C"
for epoch in trange(1, epochs+1, desc='Epoch'):
print("<" + "="*22 + F" Epoch {epoch} "+ "="*22 + ">")
############ training eval metrics ######################
train_loss = []
train_acc = []
train_prec = []
train_rec = []
train_f1 = []
#########################################################
for batch in tqdm(train_loader):
optim.zero_grad() # gradients get accumulated by default -> clear previous accumulated gradients
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
token_type_ids = batch["token_type_ids"].to(device)
labels = batch['labels'].to(device)
bio_tags = batch['bio_tags'].to(device)
################################################
model.train() # set model to training mode
logits = model(**{"input_ids":input_ids, "attention_mask":attention_mask, "token_type_ids":token_type_ids}) # forward pass
################################################
# similar to the class RobertaForToken classification in transformers: https://github.com/huggingface/transformers/blob/master/src/transformers/models/roberta/modeling_roberta.py
active_loss = attention_mask.view(-1) == 1 # either based on attention_mask (includes <CLS>, <SEP> token)
active_logits = logits.view(-1, N_bio_tags)[active_loss] # N_bio_tags=5
active_tags = bio_tags.view(-1)[active_loss]
loss = loss_fn(active_logits, active_tags)
print("loss:", loss) ## TODO VIVEK: check loss function calculation
loss.backward() # backward pass
optim.step() # update parameters and take a steup using the computed gradient
scheduler.step()# update learning rate scheduler
train_loss.append(loss.item())
################## Training Performance Measures ##########
logits = logits.detach().to('cpu').numpy()
tags_ids = bio_tags.to('cpu').numpy()
# calculate performance measures only on tokens and not subwords or special tokens
tags_mask = tags_ids != -100 # only get token labels and not labels from subwords or special tokens
pred = np.argmax(logits, axis=2)[tags_mask] #.flatten() # convert logits to list of predicted labels
tags = tags_ids[tags_mask]
metrics = compute_metrics(pred, tags)
train_acc.append(metrics["accuracy"])
train_prec.append(metrics["precision"])
train_rec.append(metrics["recall"])
train_f1.append(metrics["f1"])
train_avg_loss.append(np.mean(train_loss))
train_avg_acc.append(np.mean(train_acc))
print(F'\n\tTraining Loss: {np.mean(train_loss)}')
print(F'\n\tTraining acc: {np.mean(train_acc)}')
print(F'\n\tTraining prec: {np.mean(train_prec)}')
print(F'\n\tTraining rec: {np.mean(train_rec)}')
print(F'\n\tTraining f1: {np.mean(train_f1)}')
n_trained_epochs += 1
###################################################################################
## ---- Validation ------
val_accuracy = []
val_loss = []
val_acc = []
val_prec = []
val_rec = []
val_f1 = []
# Evaluate data for one epoch
for batch in tqdm(validation_loader):
batch = tuple(batch[t].to(device) for t in batch) # batch to GPU
v_input_ids, v_input_mask, v_token_type_ids, v_labels, v_bio_tags = batch # unpack inputs from dataloader
with torch.no_grad(): # tell model not to compute or store gradients -> saves memory + speeds up validation
model.eval() # put model in evaluation mode for validation set
logits = model(**{"input_ids":v_input_ids, "attention_mask":v_input_mask, "token_type_ids":v_token_type_ids}) # forward pass, calculates logit predictions
######################################################
# similar to the class RobertaForToken classification in transformers: https://github.com/huggingface/transformers/blob/master/src/transformers/models/roberta/modeling_roberta.py
v_active_loss = v_input_mask.view(-1) == 1 # either based on attention_mask (includes <CLS>, <SEP> token)
v_active_logits = logits.view(-1, N_bio_tags)[v_active_loss] # 5
v_active_tags = v_bio_tags.view(-1)[v_active_loss]
v_loss = loss_fn(v_active_logits, v_active_tags)
val_loss.append(v_loss.item())
#########################################################
logits = logits.detach().to('cpu').numpy()
tags_ids = v_bio_tags.to('cpu').numpy()
# calculate performance measures only on tokens and not subwords or special tokens
tags_mask = tags_ids != -100 # only get token labels and not labels from subwords or special tokens
pred = np.argmax(logits, axis=2)[tags_mask] #.flatten() # convert logits to list of predicted labels
tags = tags_ids[tags_mask]#.flatten()
metrics = compute_metrics(pred, tags)
val_acc.append(metrics["accuracy"])
val_prec.append(metrics["precision"])
val_rec.append(metrics["recall"])
val_f1.append(metrics["f1"])
val_avg_loss.append(np.mean(val_loss))
val_avg_acc.append(np.mean(val_acc))
print(F'\n\tValidation Loss: {np.mean(val_loss)}')
print(F'\n\tValidation acc: {np.mean(val_acc)}')
print(F'\n\tValidation prec: {np.mean(val_prec)}')
print(F'\n\tValidation rec: {np.mean(val_rec)}')
print(F'\n\tValidation f1: {np.mean(val_f1)}')
# early_stopping needs the validation loss to check if it has decreased,
# and if it has, it will make a checkpoint of the current model
early_stopping(np.average(val_loss), model)
if early_stopping.early_stop:
print("Early stopping")
break
# -
print(n_trained_epochs)
print(train_avg_loss)
print(val_avg_loss)
plt.plot(range(1, n_trained_epochs+1), train_avg_loss, label="train loss")
plt.plot(range(1, n_trained_epochs+1), val_avg_loss, label="val loss")
plt.title("Training Curve (lr={})".format(lr))
plt.xlabel("epochs")
plt.ylabel("Train Loss")
plt.legend(loc="best")
plt.show()
plt.plot(range(1, n_trained_epochs+1), train_avg_acc, label="train acc")
plt.plot(range(1, n_trained_epochs+1), val_avg_acc, label="val acc")
plt.title("Training Curve (lr={})".format(lr))
plt.xlabel("epochs")
plt.ylabel("Train/val Accuracy")
plt.legend(loc="best")
plt.show()
# +
########### save model ###########
# torch.save(model.state_dict(), "finetuned-35-epochs-1e3-lr-with-weighted-loss.pth") # early stopping saves model
# -
# ### Evaluation on the test dataset
# +
############ test eval metrics ######################
test_true_labels = []
test_predict_labels = []
test_loss = []
test_acc = []
test_prec = []
test_rec = []
test_f1 = []
########################################################
for batch in tqdm(test_loader):
batch = tuple(batch[t].to(device) for t in batch) # batch to GPU
t_input_ids, t_input_mask, t_token_type_ids, t_labels, t_bio_tags = batch # unpack inputs from dataloader
with torch.no_grad(): # tell model not to compute or store gradients -> saves memory + speeds up validation
model.eval() # put model in evaluation mode for validation set
logits = model(**{"input_ids":t_input_ids, "attention_mask":t_input_mask, "token_type_ids":t_token_type_ids}) # forward pass, calculates logit predictions
######################################################
# similar to the class RobertaForToken classification in transformers: https://github.com/huggingface/transformers/blob/master/src/transformers/models/roberta/modeling_roberta.py
t_active_loss = t_input_mask.view(-1) == 1 # either based on attention_mask (includes <CLS>, <SEP> token)
t_active_logits = logits.view(-1, N_bio_tags)[t_active_loss] # 5
t_active_tags = t_bio_tags.view(-1)[t_active_loss]
t_loss = loss_fn(t_active_logits, t_active_tags)
test_loss.append(t_loss.item())
#########################################################
logits = logits.detach().to('cpu').numpy()
tags_ids = t_bio_tags.to('cpu').numpy()
# calculate performance measures only on tokens and not subwords or special tokens
tags_mask = tags_ids != -100 # only get token labels and not labels from subwords or special tokens
pred = np.argmax(logits, axis=2)[tags_mask] #.flatten() # convert logits to list of predicted labels
tags = tags_ids[tags_mask]#.flatten()
test_true_labels.append(tags) # appends true labels for batch
test_predict_labels.append(pred) # # appends predicted labels for batch
metrics = compute_metrics(pred, tags)
test_acc.append(metrics["accuracy"])
test_prec.append(metrics["precision"])
test_rec.append(metrics["recall"])
test_f1.append(metrics["f1"])
print(F'\n\tTest Loss: {np.mean(test_loss)}')
print(F'\n\tTest acc: {np.mean(test_acc)}')
print(F'\n\tTest prec: {np.mean(test_prec)}')
print(F'\n\tTest rec: {np.mean(test_rec)}')
print(F'\n\tTest f1: {np.mean(test_f1)}')
# -
# ### Classification reports
from sklearn.metrics import classification_report
tag2id = {label: idx for idx, label in enumerate(["O", "B-C", "I-C", "B-E", "I-E"])}
tag2id[-100] = -100
id2tag = {id:tag for tag,id in tag2id.items()}
test_true_tag = [id2tag[ID] for ID in np.concatenate(test_true_labels)]#
test_predict_tag = [id2tag[ID] for ID in np.concatenate(test_predict_labels)]
print(classification_report(test_true_tag, test_predict_tag))
# +
from seqeval.metrics import classification_report as classification_report_seqeval
#The metrics we are seeing in this report are designed specifically for NLP tasks such as NER and POS tagging,
#in which all words of an entity need to be predicted correctly to be counted as one correct prediction.
#Therefore, the metrics in this classification report are much lower than in scikit-learn's classification report.
test_true_tag = [[id2tag[ID] for ID in IDS] for IDS in test_true_labels]
test_predict_tag = [[id2tag[ID] for ID in IDS] for IDS in test_predict_labels]
print(classification_report_seqeval(test_true_tag, test_predict_tag))
# -
# ### bio tags back to tokens
# +
# take last batch of test set:
t_input_ids, t_input_mask, t_token_type_ids, t_labels, t_bio_tags = batch
for i in range(len(batch)):
tags_mask = t_bio_tags[i].to("cpu").numpy() != -100 # only get token labels and not labels from subwords or special tokens
pred = np.argmax(logits[i], axis=1)[tags_mask]
true_tags = t_bio_tags[i][tags_mask].to("cpu").numpy()
tokens = tokenizer.convert_ids_to_tokens(t_input_ids[i])
print("\n\nPadded Sentence:")
print(tokens)
print("true labels:")
print(t_bio_tags[i])
for token, true_label, pred in zip(np.array(tokens)[tags_mask], true_tags, pred):
print(token, "\t\ttrue:", true_label, " pred:", pred)
break
# -
# ### Save model
torch.save(model.state_dict(), "weighted-loss-moreData.pth")
# ### Load model locally
# + tags=[]
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = CausalNER()
model.load_state_dict(torch.load("weighted-loss-moreData.pth"))
model.to(device)
model.eval()
# +
# dir(TweetDataSet)
# -
from transformers import BertTokenizer
# +
tokenizer1 = AutoTokenizer.from_pretrained("vinai/bertweet-base")
tokenizer2 = AutoTokenizer.from_pretrained("roberta-base")
tokenizer3 = AutoTokenizer.from_pretrained("bert-base-cased")
tokenizer4 = BertTokenizer.from_pretrained("bert-base-uncased")
#bert = transformers.BertModel.from_pretrained("vinai/bertweet-base")
seq = normalizeTweet("#diabetes is a shitty disease")
print(tokenizer1.tokenize(seq))
print(tokenizer2.tokenize(seq))
print(tokenizer3.tokenize(seq))
print(tokenizer4.tokenize(seq))
# -
for i, row in data[1:10].iterrows():
print(tokenizer1.tokenize(row["full_text"]))
print(tokenizer2.tokenize(row["full_text"]))
print(tokenizer4.tokenize(row["full_text"]))
print()
|
Cause-effect-identification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#
# +
import numpy as np
import pandas as pd
import uproot
import matplotlib.pyplot as plt
from tensorflow import keras
import tensorflow.keras.backend as K
from sificc_lib import AI, utils, Event, Simulation, root_files, DataModel
pd.options.display.max_columns = 100
pd.options.display.max_rows = 100
pd.options.display.float_format = '{:,.3f}'.format
# %matplotlib inline
# +
model_name = 'model-2a-lsf'
shuffle_clusters = False
data = DataModel('data-mtx-enough-top-6.npz',
batch_size = 64, validation_percent = .05, test_percent = .1)
data.append_dim = True
ai = AI(data, model_name)
np.random.seed(888)
ai.data.shuffle(only_train=False)
if shuffle_clusters:
ai.data.shuffle_training_clusters()
ai.data.weight_non_compton = .5
ai.weight_type = .05
ai.weight_e_cluster = .8
ai.weight_p_cluster = .8
ai.weight_pos_x = 12
ai.weight_pos_y = 2
ai.weight_pos_z = 8
ai.weight_energy = 5
# -
ai.create_model(conv_layers=[128, 64], classifier_layers=[32], type_layers=[8],
pos_layers=[64], energy_layers=[32], base_l2=.000, limbs_l2=.000)
ai.compile_model(learning_rate=0.0003)
# %%time
ai.train(epochs=100, shuffle=True, shuffle_clusters=shuffle_clusters, verbose=0)
ai.model.evaluate(ai.data.train_x, ai.data.train_y, verbose=1)
print()
ai.plot_training_loss(smooth=True)
ai.evaluate()
ai.save(file_name=model_name)
|
shallow-nn/model-2a-lsf.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="9yj_fpKmg-6P"
# This notebook contains data preparation pipeline for the [Book-Crossing Dataset](http://www2.informatik.uni-freiburg.de/~cziegler/BX/) that was originaly collected in paper [Improving Recommendation Lists Through
# Topic Diversification](http://www2.informatik.uni-freiburg.de/~cziegler/BX/WWW-2005-Preprint.pdf). Authors crawled [BookCrossing](https://www.bookcrossing.com/) website and collected data on 278 858 members and 1 157 112 ratings, both implicit and explicit, referring to 271 379 distinct ISBNs. Invalid ISBNs
# were excluded from the outset.
# + [markdown] id="pWzEuF-r0Wqp"
# # Setup
#
#
# + [markdown] id="zaznfJEj2dVy"
# ## Packages
#
# Installing and importing packages. We will work with this nice python package called [isbnlib](https://github.com/xlcnd/isbnlib) that can be used to validate, clean, transform, hyphenate and get metadata for ISBN strings.
# + colab={"base_uri": "https://localhost:8080/"} id="DGO2kBFa2t8Q" outputId="5cbe77c3-b8b1-40bf-963c-b6c2e98b67d3"
# !pip install isbnlib
# + id="UP4ASdA6559G"
import pandas as pd
import isbnlib
from tqdm import tqdm
import matplotlib.pyplot as plt
# + [markdown] id="<KEY>"
# ## Data
#
# We download Book-Crossing dataset in CSV format, unzip it and load it into pandas DataFrame. During data loading, we need to set `encoding` parameter, because data are in `ISO-8859-1` encoding, not default `UTF-8`. Also, one book title contains quote, so we need to escape it, because quote is also used to encapsulate fileds in CSV file. Troubling title looks like this:
#
# > `Peterman Rides Again: Adventures Continue with the Real \"J. Peterman\" Through Life & the Catalog Business`
#
# We need to explicitly set backslash (`\`) as escape chareacter.
# + id="NoWyhb2GU3bH" colab={"base_uri": "https://localhost:8080/"} outputId="5c59fdb1-e380-440e-c8e9-41b119366995"
# !wget http://www2.informatik.uni-freiburg.de/~cziegler/BX/BX-CSV-Dump.zip
# + colab={"base_uri": "https://localhost:8080/"} id="5IJqWy-Y1MoZ" outputId="49eb6a14-0e34-49bb-95d0-7f39ae7e1596"
# !unzip BX-CSV-Dump.zip
# + id="LSDwKzvq1O_b"
users = pd.read_csv('BX-Users.csv', sep=';', encoding = "ISO-8859-1")
books = pd.read_csv('BX-Books.csv', sep=';', encoding = "ISO-8859-1", escapechar = "\\")
ratings = pd.read_csv('BX-Book-Ratings.csv', sep=';', encoding = "ISO-8859-1")
# + [markdown] id="0EpyvWldDks9"
# # Data cleaning
#
#
# + [markdown] id="QVqGPzNRGKls"
# ## Useless columns
#
# Let's look at columns in `books` table if there are some useful information.
# + id="CUC1fmAJGTnG" colab={"base_uri": "https://localhost:8080/", "height": 434} outputId="ba437c73-d0fd-4f5a-8516-a5ef1c6396ba"
books.head()
# + [markdown] id="gyMgV3pYGbVk"
# We might use image in final application to show a preview of sugested books to a user. Or we could use it in multi-modal model as one of input for computing similarity between books. But for now, we will not need it, since we will do just simple proof-of-concept recommendation system.
# + id="7_TTo9vnGb9G"
books = books.drop(columns=['Image-URL-S', 'Image-URL-M', 'Image-URL-L'])
# + id="7ABLuwiCGff0" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="7cf2e9a4-a3ef-4cdf-cd66-fbdb638b161c"
books.head()
# + [markdown] id="X1M9-zaPGjhw"
# ## Renaming columns
#
# What we could do to make our life easier is to rename columns.
# + id="URFrOVeDGxN4" colab={"base_uri": "https://localhost:8080/"} outputId="5137f6fe-78c1-4d40-daf0-02cf3b84d1d7"
books.columns
# + id="B-pSK9AlG0Uk"
books.columns = ['ISBN', 'Title', 'Author', 'Year', 'Publisher']
# + id="zNS7SnWjHBaH" colab={"base_uri": "https://localhost:8080/"} outputId="b22794dc-15d5-42bb-b16b-75b535e7a7a0"
books.columns
# + id="2FsEhHUoHC_S" colab={"base_uri": "https://localhost:8080/"} outputId="a32a35a8-bee1-4088-a1c5-53b0a55dc416"
ratings.columns
# + id="YjDRLawBHHbc"
ratings.columns = ['User-ID', 'ISBN', 'Rating']
# + id="DtHjd5b2HMAh" colab={"base_uri": "https://localhost:8080/"} outputId="407a8e1a-5db8-4004-acb4-f2fafca433ec"
ratings.columns
# + id="F0aaeJC1HOZS" colab={"base_uri": "https://localhost:8080/"} outputId="90c6da4f-0163-4f9e-9df9-0479a528b05c"
users.columns
# + [markdown] id="RosfMfiBHQl2"
# Columns in `users` table are okey, we will keep them.
# + [markdown] id="krdq4lDiSiFk"
# ## Transforming ISBN to canonical form
#
# Reasons for bothering with ISBN numbers:
#
# - transform ISBN numbers into standard form to prevent duplicate entries
# - use ISBN as unique and valid identificator of a book, so we could connect it with other resources where books are identified by ISBN number.
#
# Data cleaning on ISBN in `books` table is connected to `ratings` table, because ISBN is used as primary key, so we need to work with both tables.
# + [markdown] id="V5v6lRfPezI4"
# First, we create a helper function to print some statistics about number of books and number of books with unique ISBNs to see if it is changing during our ISBN transformation.
# + id="Vw8IrL2GbJxk"
def book_stats():
# this is not the first run of this function
if hasattr(book_stats, "books_count"):
print("Removed books: ", book_stats.books_count - len(books))
print("Removed unique books: ", book_stats.unique_books - len(books['ISBN'].unique()))
print("Removed books in ratings: ", book_stats.ratings_count - len(ratings))
print("Removed unique books in ratings: ", book_stats.unique_ratings - len(ratings['ISBN'].unique()))
print()
# update count in each run
book_stats.books_count = len(books)
book_stats.ratings_count = len(ratings)
book_stats.unique_books = len(books['ISBN'].unique())
book_stats.unique_ratings = len(ratings['ISBN'].unique())
print("Current number of all books in books: ", book_stats.books_count)
print("Current number of entries in books with unique ISBN: ", book_stats.unique_books)
print("Current number of all books in ratings: ", book_stats.ratings_count)
print("Current number of entries in ratings with unique ISBN: ", book_stats.unique_ratings)
# + id="H3Byd7SCfW1o" outputId="73812a04-a5ed-4b41-cdaf-cecb26feb161" colab={"base_uri": "https://localhost:8080/"}
book_stats()
# + id="OUFq3OfafeZj"
books['ISBN'] = books['ISBN'].apply(lambda x: isbnlib.canonical(isbnlib.clean(x)))
ratings['ISBN'] = ratings['ISBN'].apply(lambda x: isbnlib.canonical(isbnlib.clean(x)))
# + id="g_HEw2A3fkDh" outputId="09e4e33a-3456-46dc-cdcb-160baefe6b04" colab={"base_uri": "https://localhost:8080/"}
book_stats()
# + [markdown] id="b_si9BwfgKmX"
# As we can see, we still have the same number of entries in `books` and `ratings` tables, but we have less entries with unique ISBN. Multiple ISBNs were transformed into the same ISBN number.
|
BookRecommendations/data_preparation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # gapminder
# ## Author Information
#
# **Name**: <NAME>
# **Institution**: UCLA
# **Email**: <EMAIL>
#
# **Name**: <NAME>
# **Institution**: UCLA
# **Email**: <EMAIL>
# ## Data & File Overview
#
# Excerpt from the Gapminder data for the purpose of instruction and demonstrating computational reproducibility in Dataverse. This data is an excerpt of data found in specific spreadsheets on [Gapminder.org](https://www.gapminder.org/) circa 2010. It is not a definitive source of socioeconomic data will not be updated. Use other data sources if it’s important to have the current best estimate of these statistics.
#
# The gapminder data include six variables, ([Gapminder.org documentation page](https://www.gapminder.org/data/documentation/)):
# | variable | meaning |
# |:---------|:--------|
# | country | |
# | continent | |
# | year | |
# | lifeExp | life expectancy at birth |
# | pop | total population |
# | gdpPercap | per-capita GDP |
# Per-capita GDP (Gross domestic product) is given in units of international dollars, “a hypothetical unit of currency that has the same purchasing power parity that the U.S. dollar had in the United States at a given point in time” – 2005, in this case.
# ## Geographic location of data collection
#
# * 195 states recognised by the UN which include 193 member states and 2 observing states
# * 4 regions: The Americas,
# ## File List
# |Filename | Description |
# |:--------|:------------|
# |IS262B_Daniels_Final_Project_WIP.ipynb | Jupyter Notebook demonstrating use of `gapminder data` |
# |IS262B_Daniels_Final_Project_WIP.pdf | PDF output of Jupyter Notebook above |
# |README.ipynb | Jupyter Notebook Readme file of contents of repository |
# |README.md | Jupyter Notebook converted to markdown |
# |my_figure.png | Figure saved from notebook |
# |requirements.txt | A list of Python modules used in Jupyter notebook & used by Mybinder to build Python environment|
# |data/gapminder_all_data.tab | |
# |data/gapminder_gdp_africa.tab | Subset of gapminder_all_data.tab by African countries |
# |data/gapminder_gdp_asia.tab | Subset of gapminder_all_data.tab by Asian countries |
# |data/gapminder_gdp_europe.tab | Subset of gapminder_all_data.tab by European countries |
# |data/gapminder_gdp_oceania.tab | Subset of gapminder_all_data.tab by Oceania countries |
# |data/gapminder_wide.tab | Rawer version of gapminder_all_data.tab with indicators and years in columns |
#
# ## Sharing/Acess Information
#
# Data are available freely under [CC BY 4.0 LICENCE](https://creativecommons.org/licenses/by/4.0/)
|
README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem 2
x = int(input("Value is "))
y = int(input("Value is "))
z = x + y
if z%2 == 0:
print("Even wins")
else:
print("Odd wins")
# # Problem 3
R_D_Spend = [165349.2, 162597.7, 153441.51, 144372.41, 142107.34, 131876.9, 134615.46, 130298.13, 120542.52, 123334.88]
Administration = [45654, 6475, 65785, 765856,54375, 76585, 7586, 7595, 8967876, 876976]
Marketing_Spend = [6585,769696, 758658, 758756, 5378596, 765485, 5348967098, 675433567, 1234, 12345678]
State = ['New York', 'California', 'Florida', 'New York', 'Florida', 'New York', 'Çalifornia', 'Florida', 'New York', 'California']
Profit = [234, 34567, 237, 6854, 674874, 65364, 123457898, 6383, 82633, 7636840]
tab = {'R_D_Spend': R_D_Spend, 'Administration': Administration, 'Marketing_Spend': Marketing_Spend, 'State': State, 'Profit': Profit}
tab
import pandas
dir(pandas)
data = pandas.DataFrame(tab)
data
data.to_excel('problems.xlsx', index = False)
# # Problem 4
x = int(input("Distance is "))
y = int(input("Time is "))
z = x/y
print ("Speed is ", z,"mil/hr")
# # Problem 5
|
Wednesday lab practice-Problems.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # V-Type Three-Level: 0.5π Sech Pulse, 1.5π Coupling — Simulton Propagation
# +
import numpy as np
sech_fwhm_conv = 1./2.6339157938
t_width = 1.0*sech_fwhm_conv # [τ]
print('t_width', t_width)
n = 0.5 # For a pulse area of nπ
ampl = n/t_width/(2*np.pi) # Pulse amplitude [2π Γ]
print('ampl', ampl)
n = 1.5 # For a pulse area of nπ
ampl_2 = n/t_width/(2*np.pi) # Pulse amplitude [2π Γ]
print('ampl_2', ampl_2)
# +
a = 0.5 #np.sqrt(2)
b = 1.5 #np.sqrt(2)
np.sqrt(a**2 + b**2)
# -
mb_solve_json = """
{
"atom": {
"fields": [
{
"coupled_levels": [[0, 1]],
"detuning": 0.0,
"detuning_positive": true,
"label": "probe",
"rabi_freq": 0.20960035913554168,
"rabi_freq_t_args":
{
"ampl": 1.0,
"centre": 0.0,
"width": 0.3796628587572578
},
"rabi_freq_t_func": "sech"
},
{
"coupled_levels": [[0, 2]],
"detuning": 0.0,
"detuning_positive": true,
"label": "coupling",
"rabi_freq": 0.628801077406625,
"rabi_freq_t_args":
{
"ampl": 1.0,
"centre": 0.0,
"width": 0.3796628587572578
},
"rabi_freq_t_func": "sech"
}
],
"num_states": 3
},
"t_min": -2.0,
"t_max": 10.0,
"t_steps": 60,
"z_min": -0.2,
"z_max": 1.2,
"z_steps": 70,
"z_steps_inner": 2,
"interaction_strengths": [10.0, 10.0],
"savefile": "mb-solve-vee-sech-0.5pi-1.5pi"
}
"""
# +
from maxwellbloch import mb_solve
mb_solve_00 = mb_solve.MBSolve().from_json_str(mb_solve_json)
# %time Omegas_zt, states_zt = mb_solve_00.mbsolve(recalc=False)
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_style('darkgrid')
fig = plt.figure(1, figsize=(16, 12))
# Probe
ax = fig.add_subplot(211)
cmap_range = np.linspace(0.0, 0.8, 11)
cf = ax.contourf(mb_solve_00.tlist, mb_solve_00.zlist,
np.abs(mb_solve_00.Omegas_zt[0]/(2*np.pi)),
cmap_range, cmap=plt.cm.Blues)
ax.set_title('Rabi Frequency ($\Gamma / 2\pi $)')
ax.set_ylabel('Distance ($L$)')
ax.text(0.02, 0.95, 'Probe',
verticalalignment='top', horizontalalignment='left',
transform=ax.transAxes, color='grey', fontsize=16)
plt.colorbar(cf)
# Coupling
ax = fig.add_subplot(212)
cmap_range = np.linspace(0.0, 0.8, 11)
cf = ax.contourf(mb_solve_00.tlist, mb_solve_00.zlist,
np.abs(mb_solve_00.Omegas_zt[1]/(2*np.pi)),
cmap_range, cmap=plt.cm.Greens)
ax.set_xlabel('Time ($1/\Gamma$)')
ax.set_ylabel('Distance ($L$)')
ax.text(0.02, 0.95, 'Coupling',
verticalalignment='top', horizontalalignment='left',
transform=ax.transAxes, color='grey', fontsize=16)
plt.colorbar(cf)
# Both
for ax in fig.axes:
for y in [0.0, 1.0]:
ax.axhline(y, c='grey', lw=1.0, ls='dotted')
plt.tight_layout()
# +
total_area = np.sqrt(mb_solve_00.fields_area()[0]**2 + mb_solve_00.fields_area()[1]**2)
fig, ax = plt.subplots(figsize=(16, 4))
ax.plot(mb_solve_00.zlist, mb_solve_00.fields_area()[0]/np.pi, label='Probe', clip_on=False)
ax.plot(mb_solve_00.zlist, mb_solve_00.fields_area()[1]/np.pi, label='Coupling', clip_on=False)
ax.plot(mb_solve_00.zlist, total_area/np.pi, label='Total', ls='dashed', clip_on=False)
ax.legend()
ax.set_ylim([0.0, 2.0])
ax.set_xlabel('Distance ($L$)')
ax.set_ylabel('Pulse Area ($\pi$)');
|
docs/examples/mbs-vee-sech-0.5pi-1.5pi.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# name: python38164bited0cfbe7c3504f05a4d155e3328c3124
# ---
# # Duplicate Zeros
#
# Given a fixed length array arr of integers, duplicate each occurrence of zero, shifting the remaining elements to the right.
#
# Note that elements beyond the length of the original array are not written.
#
# Do the above modifications to the input array in place, do not return anything from your function.
# ## 解析
# 本题非常简单,注意在复制0之后,0不能当作有效数字,否则数组会变成`[x,x,x,0,0,0,0,0]`的情况
def duplicateZeros(arr):
"""
Do not return anything, modify arr in-place instead.
"""
idx = 0
while idx < len(arr):
if arr[idx] == 0:
for move_idx in reversed(range(idx + 1,len(arr))):
arr[move_idx] = arr[move_idx - 1]
idx += 2
else:
idx += 1
return arr
# + tags=[]
print(duplicateZeros([1,0,2,3,0,4,5,0]))
|
Duplicate-Zeros.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bitcoin Battle Royale - Getting started (Python 2 notebook)
# Welcome to Bitcoin Battle Royale, a contest where you have the predict the probability of bitcoin moving more than 0.25% over the next 30min.
#
# This notebook aims to provide a few code snippets you can use to download data from s3, and also a few ways to interact with the feature set.
#
# You don't have to use any of this code, this should only be considered as a quick start guide...
#
# If you have any questions, ask Eben or Mike :)
# ## How to download data from S3
# This notebook references a config.ini file in the config folder. You should enter your AWS access key and secret key in the config.ini file. If you wanted to use ConfigParser to load your credentials, this is what it would look like:
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import statsmodels.graphics.tsaplots as tsa
# +
import configparser, os
Config = configparser.ConfigParser(allow_no_value=True)
Config.read(os.path.join(os.path.dirname(os.getcwd()), 'config', 'config.ini'))
aws_access_key_id = Config.get("aws_credentials", "aws_access_key_id")
aws_secret_access_key = Config.get("aws_credentials", "aws_secret_access_key")
bucket_name_value = Config.get("aws_credentials", "bucket_name_value")
# -
# To download data, we're going to create a download folder on our local machine...
download_folder = os.path.join(os.path.dirname(os.getcwd()), 'data')
if not os.path.exists(download_folder):
os.makedirs(download_folder)
print("Created download folder", download_folder)
else:
print("Download folder already exists", download_folder)
# Great! Now we're ready to download data from s3.
# The following function makes it easy to list all the bitcoin files available for download in our s3 bucket...
# +
import boto
from boto.s3.key import Key
from boto.s3.connection import OrdinaryCallingFormat
def download_most_recent_tick_data_files(download_destination=download_folder,
number_of_files_to_download=5):
# establish a connection using our access keys...
conn = boto.connect_s3(aws_access_key_id,
aws_secret_access_key,
calling_format=OrdinaryCallingFormat())
# connect to our bucket...
bucket = conn.get_bucket(bucket_name_value)
# connect to the folder that contains all the bitcoin trading data...
most_recent_files = sorted([k.key for k in boto.s3.bucketlistresultset.bucket_lister(bucket,
prefix="featuredata/GDAX/")],
reverse=True)[0:int(number_of_files_to_download)]
for most_recent_file_key in most_recent_files:
k = Key(bucket, most_recent_file_key)
k.get_contents_to_filename(os.path.join(download_destination,
most_recent_file_key.split("/")[-1]))
print("Downloaded to local...", most_recent_file_key)
# call the function...
download_most_recent_tick_data_files()
# -
# ## Quick description of features...
# +
import pandas as pd
# pick the most recent file for analysis...
most_recent_tick_data_file = os.path.join(download_folder,
sorted(os.listdir(download_folder), reverse=True)[0])
data_df = pd.read_csv(most_recent_tick_data_file)
print(most_recent_tick_data_file, "df loaded, shape", data_df.shape)
# -
# Plot of prices
# convert times to datetimes
data_df['time'] = pd.to_datetime(data_df['time'])
# +
from datetime import timedelta
def hhmax(row):
ser = data_df['price'][(data_df['time'] > row) &
(data_df['time'] <= row + timedelta(minutes=30))]
return ser.max()
data_df['MaxY'] = data_df['time'].apply(hhmax)
# +
from datetime import timedelta
import numpy as np
def hhmax(row):
ser = data_df['price'][(data_df['time'] > row) &
(data_df['time'] <= row + timedelta(minutes=30))]
minY = np.min(ser)
maxY = np.max(ser)
currentY = ser[0]
thresh_low = currentY * (1 - 0.0025)
thres_hi = currentY * (1 + 0.0025)
exceeds_low == minY <= thresh_low
exceeds_hi == maxY >= thresh_hi
if (not exceeds_low) and (not exceeds_hi):
target = 0
elif (exceeds_low) and (not exceeds_hi):
target = 1
elif (not exceeds_low) and (exceeds_hi):
target = 2
else:
target = 3
return target
data_df['MinY'] = data_df['time'].apply(hhmax)
# -
if 1==0:
a = 1
elif 1 == 1:
a = 2
# +
# define plotting function
def tsplot2(y, title, lags=None, figsize=(12, 8)):
"""
Credit to <NAME>, PyData NYC 2017, "Time Series Forecasting using Statistical and Machine Learning Models"
Examine the patterns of ACF and PACF, along with the time series plot and histogram.
:param y:
:param title:
:param lags:
:param figsize:
:return:
"""
fig = plt.figure(figsize=figsize)
layout = (2,2)
ts_ax = plt.subplot2grid(layout, (0,0))
hist_ax = plt.subplot2grid(layout, (0, 1))
acf_ax = plt.subplot2grid(layout, (1, 0))
pacf_ax = plt.subplot2grid(layout, (1, 1))
y.plot(ax=ts_ax)
ts_ax.set_title(title, fontsize=14, fontweight='bold')
y.plot(ax=hist_ax, kind='hist', bins=25)
hist_ax.set_title('Histogram')
tsa.plot_acf(y, lags=lags, ax=acf_ax)
tsa.plot_pacf(y, lags=lags, ax=pacf_ax)
[ax.set_xlim(0) for ax in [acf_ax, pacf_ax]]
sns.despine()
plt.tight_layout()
return ts_ax, acf_ax, pacf_ax
# -
data = pd.read_csv('target_test.csv')
# +
layout = (1,2)
price_ax = plt.subplot2grid(layout, (0,0))
target_ax = plt.subplot2grid(layout, (0,1))
data['price'][:10000].plot(ax=price_ax)
data['target'][:10000].plot(ax=target_ax)
# -
# for each series, plot a time series, histogram, autocorr, and partial autocorr
tsplot2(data_df['target'].dropna(), title='MaxY', lags=100)
# we could just dump all the feature names in alphabetical order...
for feature in sorted(data_df.columns.values):
print(feature)
# Key points:
#
# 1. Whenever you see a _rar value, remember that it refers to rolling average return by time, e.g. 10m = 10 minutes, 1m = 1 minute, 60m = an hour...
# 2. "larg_order" features refer to the orderbook on GDAX, largest orders, already normalized
# 3. "last_size" refers to the size of the most recent trade in BTC
# 4. "max_price", "min_price", "std_price", "max_volume", "mean_price" etc. all refers to statistical properties of recent price and volume trends
# 5. "side" is a variable that shows if the recent trade was a buy order or a sell order...
# 6. "time" is the UTC timestamp of the trade
# 7. "volume_over_price" and "volume_under_price" - ?
# 8. "vwp" refers to volume weighted price
# ## Display plots for feature interaction
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = [10, 10]
feature_categories_to_plot = ["_larg_order_10_rar", "_price_10_rar", "_volume_10_rar"]
for feature_category in feature_categories_to_plot:
features_to_scan = sorted([h for h in data_df.columns.values if h.find(feature_category) != -1])[0:4]
data_df[features_to_scan].hist()
# -
# ...notice how very few of these features look like they have a "normal" distribution...this is likely due to the impact of outliers...
#
# One quick and easy way to filter for outliers is to simply remove the bottom x% and the top x% of a feature set...
# +
import numpy as np
import warnings
warnings.filterwarnings('ignore')
feature_categories_to_plot = ["_larg_order_10_rar", "_price_10_rar", "_volume_10_rar"]
for feature_category in feature_categories_to_plot:
features_to_scan = sorted([h for h in data_df.columns.values if h.find(feature_category) != -1])[0:4]
outliers_removed_df = data_df[features_to_scan].copy()
outliers_removed_df.dropna(inplace=True)
for feature_to_scan in features_to_scan:
lower_bound = np.percentile(outliers_removed_df[feature_to_scan], 10)
upper_bound = np.percentile(outliers_removed_df[feature_to_scan], 90)
print(feature_to_scan, "lower_bound", lower_bound, "upper_bound", upper_bound)
outliers_removed_df[feature_to_scan] = outliers_removed_df[feature_to_scan].map(lambda x: lower_bound if x < lower_bound
else upper_bound if x > upper_bound
else x)
outliers_removed_df[features_to_scan].hist()
# +
# some of the feature sets have very high correlations with other feature sets...
# this is a function that takes a feature set as a string tag and generates correlation matrices
import seaborn as sns
def generate_correlation_plot(feature_category_to_plot):
correlation_df = data_df.copy()
headers_to_plot = sorted([h for h in correlation_df.columns.values if h.find(feature_category_to_plot) != -1])
correlation_df = correlation_df[headers_to_plot]
correlation_df.dropna(inplace=True)
corr = correlation_df.corr()
sns.heatmap(corr,
xticklabels=corr.columns.values,
yticklabels=corr.columns.values)
# -
generate_correlation_plot("_volume_")
generate_correlation_plot("_price_")
generate_correlation_plot("_larg_order_10_")
|
notebooks/feature_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MARIO GAME
# <img src='./img/ejercicio1.png'>
# Implemente un programa que imprima una media pirámide de una altura específica, como se indica a continuación.
# <code>$ ./mario
# Height: 4
# #
# ##
# ###
# ####</code>
# ## Especificaciones
# - Cree un archivo llamado <code>mario.py</code>,el cual es un programa que recrea una media pirámide usando los hash (#) para los bloques.
# - Para hacer las cosas más interesantes, primero solicite al usuario <code>input</code> la altura de la media pirámide, el cual debe ser un número entero positivo entre <code>1</code> y <code>8</code>, inclusive.
# - Si el usuario no proporciona un número entero positivo no mayor que <code>8</code>, debe volver a solicitar el mismo.
# - Luego, genere (con la ayuda de <code>print</code> uno o más bucles) la media pirámide deseada.
# - Tenga cuidado de alinear la esquina inferior izquierda de su media pirámide con el borde izquierdo de la ventana de su terminal.
# ## Uso
# Su programa debería comportarse según el ejemplo siguiente.
# <code>$ ./mario
# Height: 4
# #
# ##
# ###
# ####</code>
# ## Pruebas
# - Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. Escribe <code>-1</code> y presiona enter. Su programa debe rechazar esta entrada como inválida, por ejemplo, volviendo a solicitar al usuario que escriba otro número.
# - Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. Escribe <code>0</code> y presiona enter. Su programa debe rechazar esta entrada como inválida, por ejemplo, volviendo a solicitar al usuario que escriba otro número.
# - Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. Escribe <code>1</code> y presiona enter. Su programa debería generar la siguiente salida. Asegúrese de que la pirámide esté alineada con la esquina inferior izquierda de su terminal y de que no haya espacios adicionales al final de cada línea.
# <code>#</code>
# Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. Escribe <code>2</code> y presiona enter. Su programa debería generar la siguiente salida. Asegúrese de que la pirámide esté alineada con la esquina inferior izquierda de su terminal y de que no haya espacios adicionales al final de cada línea.
#
# <code> #
# ##</code>
# Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. Escribe <code>8</code> y presiona enter. Su programa debería generar la siguiente salida. Asegúrese de que la pirámide esté alineada con la esquina inferior izquierda de su terminal y de que no haya espacios adicionales al final de cada línea.
#
# <code> #
# ##
# ###
# ####
# #####
# ######
# #######
# ########</code>
# Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. Escribe <code>9</code> y presiona enter. Su programa debe rechazar esta entrada como inválida, por ejemplo, volviendo a solicitar al usuario que escriba otro número. Luego, escribe <code>2</code> y presiona enter. Su programa debería generar la siguiente salida. Asegúrese de que la pirámide esté alineada con la esquina inferior izquierda de su terminal y de que no haya espacios adicionales al final de cada línea.
#
# <code> #
# ##</code>
#
#
# - Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. Escribe fooy presiona enter. Su programa debe rechazar esta entrada como inválida, por ejemplo, volviendo a solicitar al usuario que escriba otro número.
# - Ejecute su programa como <code>python mario.py</code> y espere una solicitud de entrada. No escriba nada y presione enter. Su programa debe rechazar esta entrada como inválida, por ejemplo, volviendo a solicitar al usuario que escriba otro número.
|
Modulo2/Ejercicios/Problema1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Activity 5.1: Storefront location associations
import numpy as np, pandas as pd
import matplotlib.pyplot as plt, seaborn as sns
# %matplotlib inline
df = pd.read_csv('location_rev.csv')
df.head()
df.plot.scatter("median_income", 'revenue', figsize=[5,5])
plt.show()
# +
import seaborn as sns
# %matplotlib inline
sns.pairplot(df)
plt.show()
# -
sns.pairplot(df,y_vars="revenue")
plt.show()
df.corr()
# ## Activity 5.2
import pandas as pd
df = pd.read_csv('location_rev.csv')
df.head()
X = df[['num_competitors',
'median_income',
'num_loyalty_members',
'population_density',
'location_age'
]]
y = df['revenue']
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 100)
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X_train,y_train)
#Print out the model coefficients:
model.coef_
# -
model.intercept_
# +
single_location = pd.DataFrame({
'num_competitors': [3],
'median_income': [30000],
'num_loyalty_members': [1200],
'population_density': [2000],
'location_age': [10]
})
model.predict(single_location)
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.scatter(model.predict(X_test),y_test)
plt.xlabel('Model Predictions')
plt.ylabel('True Value')
plt.plot([0, 100000], [0, 100000], 'k-', color = 'r')
plt.show()
# +
from scipy.stats.stats import pearsonr
pearsonr(model.predict(X_test),y_test)
# -
|
Chapter05/Ch05 Activities v0.1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Implementing a Priority Queue ###
#
# #### Problem: Implement a queue that sorts items by a given priority and always returns the item with the highest priority on each pop operation.
#
# #### Solution:
# - Using the `heapq` module to implement a simple priority queue.
# +
import heapq
class PriorityQueue(object):
def __init__(self):
self._queue = []
self._index = 0
def push(self, item, priority):
heapq.heappush(self._queue, (-priority, self._index, item))
self._index += 1
def pop(self):
return heapq.heappop(self._queue)[-1]
# +
#Example of using the queue
class Item(object):
def __init__(self, name):
self.name = name
def __repr__(self):
return 'Item([!r])'.format(self.name)
q = PriorityQueue()
q.push(Item('foo'), 1)
q.push(Item('bar'), 5)
q.push(Item('spam'), 4)
q.push(Item('grok'), 1)
q.pop()
q.pop()
q.pop()
q.pop()
# -
# - The first `pop()` operation returns the item with the highest priority.
# - And, the iems, foo and grok, having the same priority are returned in the same order in which they are inserted into the queue.
#
# #### Discussion
# The core concept is the use of `heapq` module.
# - the functions `heapq.heappush()` and the `heapq.headppop()` insert and remove itesm from the list `_queue` in a way that the first item is always the smallest priority.
# - Example: instances of `item` that can't be ordered.
a = Item('foo')
b = Item('bar')
a < b
|
data structures & algorithms/5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# metadata:
# interpreter:
# hash: 7d6993cb2f9ce9a59d5d7380609d9cb5192a9dedd2735a011418ad9e827eb538
# name: python3
# ---
# + dotnet_interactive={"language": "python"}
# Assignment One
# Course: MIS 5150
# Professor: Dr. <NAME>
# + dotnet_interactive={"language": "python"}
import tensorflow as tf
import numpy as np
# + dotnet_interactive={"language": "python"}
# Question 1:
#
import tensorflow as tf
import numpy as np
# a.
# Creating dataset
data_set = tf.data.Dataset.from_tensor_slices([[0, 1, 9, 8, 7, 3], [2, 9, 4, 0, 2, 3], [7, 3, 3, 2, 2, 1], [0, 0, 1, 2, 2, 5]])
# b.
# Iterating data
# method 1
for elements in data_set:
print(elements)
print('')
# method 2
for elements in data_set:
print(elements.numpy())
print('')
# method 3
iterator = iter(data_set)
print(iterator.get_next())
print(iterator.get_next())
print(iterator.get_next())
print(iterator.get_next())
print('')
# method 4
for data in data_set:
print (data.numpy())
print('')
# c.
# display 3rd element from 2nd tensor
# method 1
ls = []
for item in data_set:
e = item.numpy()
ls.append(e)
print('3rd element from 2nd tensor:',ls[1][2])
print('')
# method 2
np_arr = np.asarray(ls, dtype=np.float32)
print('3rd element from 2nd tensor:',int(np_arr[1][2]))
print('')
# method 3
display_element = np.array([[0, 1, 9, 8, 7, 3], [2, 9, 4, 0, 2, 3], [7, 3, 3, 2, 2, 1], [0, 0, 1, 2, 2, 5]])
print('3rd element from 2nd tensor:', display_element[1][2])
# + dotnet_interactive={"language": "python"}
# Question 2:
#
import tensorflow as tf
# GPU Enable
tf.__version__, tf.test.gpu_device_name()
# output ('2.4.1', '/device:GPU:0')
# + dotnet_interactive={"language": "python"}
# Question 3:
#
import tensorflow as tf
import pandas as pd
from tensorflow import keras
ds = 'flag.data'
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/flags/flag.data'
dataset_path = tf.keras.utils.get_file(ds, url)
# downloaded the data using keras
dataset_path
# cols from the flag.names file
cols = ['name','landmass','zone','area','population','language','religion','bars','stripes','colours','red','green','blue','gold','white','black','orange','mainhue','circles','crosses','saltires','quarters','sunstars','crescent','triangle','icon','animate','text','topleft','botright']
# raw dataset using pandas
raw_dataset = pd.read_csv(dataset_path, names=cols)
# displaying dataset
raw_dataset.head()
# remove non numerical values
raw_dataset.pop('name')
raw_dataset.pop('mainhue')
raw_dataset.pop('topleft')
raw_dataset.pop('botright')
# print to check removal of non-numerical values
raw_dataset.head()
# step 2
dataset = tf.data.Dataset.from_tensor_slices(raw_dataset.values)
# range for dataset
dataset = tf.data.Dataset.range(10)
# using take function
for i in dataset.take(3):
print(i)
print('')
# + dotnet_interactive={"language": "python"}
# Question 4:
#
import tensorflow as tf
# a. create a tensor
dataset = tf.data.Dataset.range(10)
# b. display the tendor
for row in dataset:
print(row)
print('')
# c. repeat and batch
data_batch = dataset.repeat(5).batch(4)
# d. display batched sensor
for row in data_batch:
print(row)
print('')
# e. map to cube the dataset
data_map = data_batch.map(lambda x:x ** 3)
# f. batched tensor display
for item in data_map.take(2):
print(item)
print('')
# + dotnet_interactive={"language": "python"}
# Question 5:
#
import tensorflow as tf
# a. create dataset
dataset = tf.data.Dataset.range(10).repeat(5)
print ('dataset has', len(list(dataset)), 'elements')
print('')
# b. dataset buffer and display
ds = dataset.shuffle(buffer_size=5).batch(7)
for item in ds:
print(item)
print('')
|
assignment/assignment1/assignment_one.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Part 6 - Utility functions for the map widget
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Demonstrate-callback-features-with-on_click,-and-on_draw_end" data-toc-modified-id="Demonstrate-callback-features-with-on_click,-and-on_draw_end-1">Demonstrate callback features with <code>on_click</code>, and <code>on_draw_end</code></a></span></li><li><span><a href="#Show-message-in-map-widget-via-display_message" data-toc-modified-id="Show-message-in-map-widget-via-display_message-2">Show message in map widget via <code>display_message</code></a></span></li><li><span><a href="#Hide-the-Map/Scene-Switch-button-using-hide_mode_switch" data-toc-modified-id="Hide-the-Map/Scene-Switch-button-using-hide_mode_switch-3">Hide the Map/Scene Switch button using <code>hide_mode_switch</code></a></span></li><li><span><a href="#Configure-the-map-widget-with-the-Javascript-API-using-set_js_cdn" data-toc-modified-id="Configure-the-map-widget-with-the-Javascript-API-using-set_js_cdn-4">Configure the map widget with the Javascript API using <code>set_js_cdn</code></a></span></li><li><span><a href="#Conclusion" data-toc-modified-id="Conclusion-5">Conclusion</a></span></li></ul></div>
# -
# In this notebook, we will explore using utility functions to build interactivity. This can usually be split into two procedures: `call back` and `display message`. The `call back` function can be used to collect `gis` data on screen through digitization, while the `display message` function can be used to prompt users to perform a desired action.
#
# To do this, we will need to setup an asynchronous `callback` function using the `on_click()` or `on_draw_end()` methods to create dynamic, interactive 'apps'. We will need to create a `callback` function, like `function_name(map_inst, geometry)`, with map_inst being the `MapView` instance and geometry being the geometry instance that the user is clicking.
# ## Demonstrate callback features with `on_click`, and `on_draw_end`
#
# Our first example will take a point that a user clicks on the map, reverse geocode it from the geometry, and print out the resulting location.
#
# Note, you can either create the `GIS` connection using an existing profile, or you can simply enter your username and password, e.g. `gis = GIS("https://www.arcgis.com", "username", "password")`.
# +
from arcgis.gis import GIS
import arcgis.geocoding as geocoding
gis = GIS('home')
callback_map = gis.map('San Diego convention center, San Diego, CA', 16)
# -
def find_addr(callback_map, g):
try:
callback_map.draw(g)
geocoded = geocoding.reverse_geocode(g)
print(geocoded['address']['Match_addr'])
except:
print("Couldn't match address. Try another place...")
callback_map
callback_map.on_click(find_addr)
# Next, we want a user to draw a freehand `polyline` to indicate the paths they take for their runs. When the drawing operation ends, we use the `GIS`'s Geometry service to compute the length of the drawn path. We can do this by adding an event listener to the map widget that gets called when drawing is completed (i.e. `on_draw_end`). The event listener then computes the geodesic length of the drawn geometry using the geometry service and prints it out:
drawend_map = gis.map('San Diego convention center, San Diego, CA', 16)
# +
from arcgis.geometry import lengths
# Define the callback function that computes the length.
def calc_dist(drawend_map, g):
print("Computing length of drawn polyline...")
length = lengths(g['spatialReference'], [g], "", "geodesic")
print("Length: " + str(length[0]) + " mile(s).")
# -
# Set calc_dist as the callback function to be invoked when a polyline is drawn on the map
drawend_map.on_draw_end(calc_dist)
drawend_map
drawend_map.draw("polyline")
# ## Show message in map widget via `display_message`
#
# The `display_message` method displays a message on the upper-right corner of the map widget.
#
# <div class="alert alert-info">
# <b>Note:</b> Only one message can be sent at a time, and multiple messages will not show up. Let's experiment with the first example by changing the print method to display_message so the desired text output can be shown inside the widget.
# </div>
callback_map
def find_addr2(callback_map, g):
try:
callback_map.draw(g)
geocoded = geocoding.reverse_geocode(g)
callback_map.display_message(geocoded['address']['Match_addr'])
except:
callback_map.display_message("Couldn't match address. Try another place...")
callback_map.on_click(find_addr2)
# ## Hide the Map/Scene Switch button using `hide_mode_switch`
#
# When `hide_mode_switch` is set to `True`, the 2D/3D switch button will be hidden from the widget.
#
# <div class="alert alert-info">
# <b>Note:</b> Once the button is hidden, it cannot be made visible again. A new `MapView` instance must be created to see the button.
# </div>
callback_map.hide_mode_switch = False
# ## Configure the map widget with the Javascript API using `set_js_cdn`
#
# The `set_js_cdn` function is called before the creation of any `MapView` object, and each instantiated object will use the specified js_cdn parameter as the ArcGIS API for JavaScript CDN URL, rather than the default `http://js.arcgis.com/4.X/`. This functionality is necessary in disconnected environments if the enterprise you are connecting to does not ship with the minimum necessary JavaScript API version.
#
# <div class="alert alert-info">
# <b>Note:</b> You may not need to call this function to view the widget in disconnected environments. For instance, if your computer cannot reach `js.arcgis.com` and you have a `GIS` connection to a portal, the widget will automatically attempt to use that enterprise’s JS API that it ships with.
# </div>
#
# For example, if `gis.map()` does not return a map, run the following code to configure the map widget with the Javascript API shipped with the enterprise:
if callback_map.set_js_cdn("https://pythonapi.playground.esri.com/portal/home/10.8.1/js/jsapi") :
print("True")
# ## Conclusion
#
# In Part 6 of this guide series, we discussed the utility functions of the `MapView` object, particularly in regard to using the `callback` and `display_message` mechanisms in creating interactivity for users.
#
# <a href="#Part-6---Utility-functions-for-the-map-widget">Back to Top</a>
|
guide/10-mapping-and-visualization/part6_utility_functions_for_the_map_widget.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # DLVC 2017
# # Tutorial 4 : PyTorch for Deep Neural Networks
#
# ## Follow instructions given in the PyTorch [website](http://pytorch.org/) for installation
# ### Tensor initialization and slicing
# Importing the library
import torch
# Defining tensors
x = torch.Tensor(2,2) # Uninitialized (garbage value present in memory)
y = torch.rand(2,2) # Random initialization
print x,y
# Obtaining size of tensors
print x.size()
# Arithmetic operations
z = torch.rand(2,2)
print y,z,y+z
print z,2*z
# Indexing is similar to numpy indexing
print y
print y[1,1]
print y[:,0]
# Converting to numpy
y_np = y.numpy()
print y
print y_np
import numpy as np
# Converting from numpy to tensor
x_np = np.ones((3,3))
x_py = torch.from_numpy(x_np)
print x_np,x_py
# Improving computational time with GPU acceleration
import time
use_gpu = torch.cuda.is_available()
x = torch.randn(10000,10000)
if use_gpu:
x = torch.randn(100,100)
cpuStart = time.time()
y = x*x
cpuEnd = time.time()-cpuStart
x = x.cuda()
gpuStart = time.time()
y = x*x
gpuEnd = time.time()-gpuStart
print('CPU computation completed in {:.6f}s, GPU computation completed in {:.6f}s'\
.format(cpuEnd,gpuEnd))
# ### Autograd
# This package provides automatic differentiation for all operations on Tensors
from torch.autograd import Variable
a = torch.ones(2,2)
a_var = Variable(a,requires_grad=True)
print a
print(a_var)
b = a_var+2
print b
print(b.grad_fn)
c = b*b*3
d = c.mean()
print c
print d
d.backward()
# Gradients
print a_var.grad
# ### Torchvision datasets
# %matplotlib inline
import torch
from torchvision import datasets,transforms
import matplotlib.pyplot as plt
apply_transform = transforms.Compose([transforms.ToTensor()])
trainDset = datasets.MNIST('./MNIST',train=True, download=True, transform= apply_transform)
testDset = datasets.MNIST('./MNIST',train=False, download=True, transform= apply_transform)
# Number of samples
print(len(trainDset),len(testDset))
# Displaying sample image from the dataset
img = trainDset[0][0].numpy().transpose(1,2,0).squeeze(2)
plt.imshow(img,'gray')
print('Label: '+str(trainDset[0][1]))
# Creating dataloader for loading data in batches
trainLoader = torch.utils.data.DataLoader(trainDset, batch_size=10, shuffle=True, num_workers=1, pin_memory=False)
testLoader = torch.utils.data.DataLoader(testDset, batch_size=10, shuffle=True, num_workers=1, pin_memory=False)
# ### Defining a multi-layer perceptron
import torch.nn.functional as F
import torch.nn as nn
class MLP(nn.Module):
def __init__(self):
super(MLP, self).__init__()
self.fc1 = nn.Linear(28*28,100)
self.fc2 = nn.Linear(100,10)
def forward(self, x):
x = self.fc1(x)
x = F.sigmoid(x)
x = self.fc2(x)
x = F.log_softmax(x)
return x
net = MLP()
print(net)
params = list(net.parameters())
print('No. of parameters :'+str(len(params)))
print('Dimensions of first parameter: '+str(params[0].size())) # Weights of fc1
print('Dimensions of second parameter: '+str(params[1].size())) # Biases of fc1
inp = trainLoader.dataset[0][0]
label = trainLoader.dataset[0][1]
from torch.autograd import Variable
# Feed-forward data through network
out = net(Variable(inp.view(-1,28*28)))
print(inp.size())
print(out.size())
# Backpropagating gradients
net.zero_grad()
out.backward(torch.randn(1, 10)) # Using random gradients
out = net(Variable(inp.view(-1,28*28)))
# Defining loss function
criterion = nn.NLLLoss() # Negative log-likelihood loss
label = label*torch.ones(1) # Converting to tensor
loss = criterion(out,Variable(label.long())) # NLLLoss() expects the labels to be of dtype 'long'
print(loss)
# +
# Backprogattion
net.zero_grad() # zeroes the gradient buffers of all parameters
print('Bias gradient of fc1 before backward')
print(net.fc1.bias.grad[:10])
loss.backward()
print('Bias gradient of fc1 after backward')
print(net.fc1.bias.grad[:10])
# +
import copy
# Updataing weights of the network
learning_rate = 1
init_params = copy.deepcopy(net.fc2.weight.data) # Copying initial parameters
for f in net.parameters():
f.data.sub_(f.grad.data * learning_rate)
updated_params = net.fc2.weight.data
print(init_params[0,:5])
print(updated_params[0,:5])
# -
|
Notebooks/4_PyTorch for Deep Neural Networks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# launch XVFB if you run on a server
import os
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY"))==0:
# !bash ../xvfb start
# %env DISPLAY=:1
# ### Let's make a TRPO!
#
# In this notebook we will write the code of the one Trust Region Policy Optimization.
# As usually, it contains a few different parts which we are going to reproduce.
#
#
import numpy as np
import tensorflow as tf
# +
import gym
env = gym.make("Acrobot-v1")
env.reset()
observation_shape = env.observation_space.shape
n_actions = env.action_space.n
print("Observation Space", env.observation_space)
print("Action Space", env.action_space)
# -
import matplotlib.pyplot as plt
# %matplotlib inline
plt.imshow(env.render('rgb_array'))
# ### Step 1: Defining a network
#
# With all it's complexity, at it's core TRPO is yet another policy gradient method.
#
# This essentially means we're actually training a stochastic policy $ \pi_\theta(a|s) $.
#
# And yes, it's gonna be a neural network. So let's start by defining one.
# +
#input tensors
observations_ph = tf.placeholder(shape=(None, observation_shape[0]), dtype=tf.float32)
# Actions that we made
actions_ph = tf.placeholder(shape=(None,), dtype=tf.int32)
# "G = r + gamma*r' + gamma^2*r'' + ..."
cummulative_returns_ph = tf.placeholder(shape=(None,), dtype=tf.float32)
# Action probabilities from previous iteration
old_probs_ph = tf.placeholder(shape=(None, n_actions), dtype=tf.float32)
all_inputs = [observations_ph,actions_ph,cummulative_returns_ph,old_probs_ph]
# +
def denselayer(name, x, out_dim, nonlinearity=None):
with tf.variable_scope(name):
if nonlinearity is None:
nonlinearity = tf.identity
x_shape = x.get_shape().as_list()
w = tf.get_variable('w', shape=[x_shape[1], out_dim])
b = tf.get_variable('b', shape=[out_dim], initializer=tf.constant_initializer(0))
o = nonlinearity(tf.matmul(x, w) + b)
return o
sess = tf.InteractiveSession()
nn = observations_ph
#<your network here>
dense1 = denselayer('hidden1',nn, 128)
dense2 = denselayer('hidden2',dense1, 32)
policy_out = denselayer('policy',dense2, n_actions, nonlinearity = tf.log) #<layer that predicts action log-probabilities>
probs_out = tf.exp(policy_out)
weights = tf.trainable_variables()
sess.run(tf.global_variables_initializer())
# -
# ### Step 2: Actions and rollouts
#
# In this section, we'll define functions that take actions $ a \sim \pi_\theta(a|s) $ and rollouts $ <s_0,a_0,s_1,a_1,s_2,a_2,...s_n,a_n> $.
# +
#compile function
def act(obs, sample=True):
"""
Samples action from policy distribution (sample = True) or takes most likely action (sample = False)
:param: obs - single observation vector
:param sample: if True, samples from \pi, otherwise takes most likely action
:returns: action (single integer) and probabilities for all actions
"""
probs = sess.run(probs_out, feed_dict = {observations_ph:obs.reshape((1, -1))})[0]
if sample:
action = int(np.random.choice(n_actions,p=probs))
else:
action = int(np.argmax(probs))
return action, probs
# -
#demo
print ("sampled:", [act(env.reset()) for _ in range(5)])
print ("greedy:", [act(env.reset(),sample=False) for _ in range(5)])
# Compute cummulative reward just like you did in vanilla REINFORCE
import scipy.signal
def get_cummulative_returns(r, gamma=1):
"""
Computes cummulative discounted rewards given immediate rewards
G_i = r_i + gamma*r_{i+1} + gamma^2*r_{i+2} + ...
Also known as R(s,a).
"""
r = np.array(r)
assert r.ndim >= 1
return scipy.signal.lfilter([1], [1, -gamma], r[::-1], axis=0)[::-1]
#simple demo on rewards [0,0,1,0,0,1]
get_cummulative_returns([0,0,1,0,0,1],gamma=0.9)
# **Rollout**
def rollout(env, act, max_pathlength=2500, n_timesteps=50000):
"""
Generate rollouts for training.
:param: env - environment in which we will make actions to generate rollouts.
:param: act - the function that can return policy and action given observation.
:param: max_pathlength - maximum size of one path that we generate.
:param: n_timesteps - total sum of sizes of all pathes we generate.
"""
paths = []
total_timesteps = 0
while total_timesteps < n_timesteps:
obervations, actions, rewards, action_probs = [], [], [], []
obervation = env.reset()
for _ in range(max_pathlength):
action, policy = act(obervation)
obervations.append(obervation)
actions.append(action)
action_probs.append(policy)
obervation, reward, done, _ = env.step(action)
rewards.append(reward)
total_timesteps += 1
if done or total_timesteps==n_timesteps:
path = {"observations": np.array(obervations),
"policy": np.array(action_probs),
"actions": np.array(actions),
"rewards": np.array(rewards),
"cumulative_returns":get_cummulative_returns(rewards),
}
paths.append(path)
break
return paths
paths = rollout(env,act,max_pathlength=5,n_timesteps=100)
print (paths[-1])
assert (paths[0]['policy'].shape==(5, n_actions))
assert (paths[0]['cumulative_returns'].shape==(5,))
assert (paths[0]['rewards'].shape==(5,))
assert (paths[0]['observations'].shape==(5,)+observation_shape)
assert (paths[0]['actions'].shape==(5,))
print ('It\'s ok')
# ### Step 3: loss functions
#
# Now let's define the loss functions and constraints for actual TRPO training.
# The surrogate reward should be
# $$J_{surr}= {1 \over N} \sum\limits_{i=0}^N \frac{\pi_{\theta}(s_i, a_i)}{\pi_{\theta_{old}}(s_i, a_i)}A_{\theta_{old}(s_i, a_i)}$$
#
# For simplicity, let's use cummulative returns instead of advantage for now:
# $$J'_{surr}= {1 \over N} \sum\limits_{i=0}^N \frac{\pi_{\theta}(s_i, a_i)}{\pi_{\theta_{old}}(s_i, a_i)}G_{\theta_{old}(s_i, a_i)}$$
#
# Or alternatively, minimize the surrogate loss:
# $$ L_{surr} = - J'_{surr} $$
#select probabilities of chosen actions
batch_size = tf.shape(observations_ph)[0]
probs_all = tf.reshape(probs_out, [-1])
probs_for_actions = tf.gather(probs_all, tf.range(0, batch_size) * n_actions + actions_ph)
old_probs_all = tf.reshape(old_probs_ph, [-1])
old_probs_for_actions = tf.gather(old_probs_all, tf.range(0, batch_size) * n_actions + actions_ph)
# +
# Compute surrogate loss: negative importance-sampled policy gradient
J_surr = tf.reduce_mean(((probs_for_actions/ old_probs_for_actions) * cummulative_returns_ph),axis = 0, keepdims = True)
L_surr = - J_surr #<compute surrogate loss, aka _negative_ importance-sampled policy gradient>
# +
#compute and return surrogate policy gradient
def var_shape(x):
return [k.value for k in x.get_shape()]
def numel(x):
return np.prod(var_shape(x))
def flatgrad(loss, var_list):
grads = tf.gradients(loss, var_list)
return tf.concat([tf.reshape(grad, [numel(v)])
for (v, grad) in zip(var_list, grads)], 0)
flat_grad_surr = flatgrad(L_surr, weights)
# -
# We can ascend these gradients as long as our $pi_\theta(a|s)$ satisfies the constraint
# $$E_{s,\pi_{\Theta_{t}}}\Big[KL(\pi(\Theta_{t}, s) \:||\:\pi(\Theta_{t+1}, s))\Big]< \alpha$$
#
#
# where
#
# $$KL(p||q) = E _p log({p \over q})$$
# +
# Compute Kullback-Leibler divergence (see formula above)
# Note: you need to sum KL and entropy over all actions, not just the ones agent took
old_log_probs = tf.log(old_probs_ph+1e-10)
kl = tf.reduce_mean(old_probs_all * tf.log(old_probs_all/probs_all+1e-10)) #<compute kullback-leibler as per formula above>
#Compute policy entropy
entropy = tf.reduce_mean(-probs_all * tf.log(probs_all+1e-10), axis = 0, keepdims = True) #<compute policy entropy. Don't forget the sign!>
losses = [L_surr, kl, entropy]
# -
# **Linear search**
#
# TRPO in its core involves ascending surrogate policy gradient constrained by KL divergence.
#
# In order to enforce this constraint, we're gonna use linesearch. You can find out more about it [here](https://en.wikipedia.org/wiki/Linear_search)
def linesearch(f, x, fullstep, max_kl):
"""
Linesearch finds the best parameters of neural networks in the direction of fullstep contrainted by KL divergence.
:param: f - function that returns loss, kl and arbitrary third component.
:param: x - old parameters of neural network.
:param: fullstep - direction in which we make search.
:param: max_kl - constraint of KL divergence.
:returns:
"""
max_backtracks = 10
loss, _, _ = f(x)
for stepfrac in .5**np.arange(max_backtracks):
xnew = x + stepfrac * fullstep
new_loss, kl, _ = f(xnew)
actual_improve = new_loss - loss
if kl<=max_kl and actual_improve < 0:
x = xnew
loss = new_loss
return x
# ### Step 4: training
# In this section we construct rest parts of our computational graph
def slice_vector(vector, shapes):
"""
Slices symbolic vector into several symbolic tensors of given shapes.
Auxilary function used to un-flatten gradients, tangents etc.
:param vector: 1-dimensional symbolic vector
:param shapes: list or tuple of shapes (list, tuple or symbolic)
:returns: list of symbolic tensors of given shapes
"""
assert len(vector.get_shape())==1,"vector must be 1-dimensional"
start = 0
tensors = []
for shape in shapes:
size = np.prod(shape)
tensor = tf.reshape(vector[start:(start + size)],shape)
tensors.append(tensor)
start += size
return tensors
# +
#intermediate grad in conjugate_gradient
conjugate_grad_intermediate_vector = tf.placeholder(dtype=tf.float32, shape=(None,))
#slice flat_tangent into chunks for each weight
weight_shapes = [sess.run(var).shape for var in weights]
tangents = slice_vector(conjugate_grad_intermediate_vector,weight_shapes)
# KL divergence where first arg is fixed
kl_firstfixed = tf.reduce_sum((tf.stop_gradient(probs_out) * (tf.stop_gradient(tf.log(probs_out)) - tf.log(probs_out))))/ tf.cast(batch_size, tf.float32)
#compute fisher information matrix (used for conjugate gradients and to estimate KL)
gradients = tf.gradients(kl_firstfixed, weights)
gradient_vector_product = [tf.reduce_sum(g[0] * t) for (g, t) in zip(gradients, tangents)]
fisher_vec_prod = flatgrad(gradient_vector_product, weights)
# -
# ### TRPO helpers
#
# Here we define a few helper functions used in the main TRPO loop
# **Conjugate gradients**
#
# Since TRPO includes contrainted optimization, we will need to solve Ax=b using conjugate gradients.
#
# In general, CG is an algorithm that solves Ax=b where A is positive-defined. A is Hessian matrix so A is positive-defined. You can find out more about them [here](https://en.wikipedia.org/wiki/Conjugate_gradient_method)
from numpy.linalg import inv
def conjugate_gradient(f_Ax, b, cg_iters=10, residual_tol=1e-10):
"""
This method solves system of equation Ax=b using iterative method called conjugate gradients
:f_Ax: function that returns Ax
:b: targets for Ax
:cg_iters: how many iterations this method should do
:residual_tol: epsilon for stability
"""
p = b.copy()
r = b.copy()
x = np.zeros_like(b)
rdotr = r.dot(r)
for i in range(cg_iters):
z = f_Ax(p)
v = rdotr / (p.dot(z) + 1e-8)
x += v * p
r -= v * z
newrdotr = r.dot(r)
mu = newrdotr / (rdotr + 1e-8)
p = r + mu * p
rdotr = newrdotr
if rdotr < residual_tol:
break
return x
# +
#This code validates conjugate gradients
A = np.random.rand(8, 8)
A = np.matmul(np.transpose(A), A)
def f_Ax(x):
return np.matmul(A, x.reshape(-1, 1)).reshape(-1)
b = np.random.rand(8)
w = np.matmul(np.matmul(inv(np.matmul(np.transpose(A), A)), np.transpose(A)), b.reshape((-1, 1))).reshape(-1)
print (w)
print (conjugate_gradient(f_Ax, b))
# +
#Compile a function that exports network weights as a vector
flat_weights = tf.concat([tf.reshape(var, [-1]) for var in weights], axis=0)
#... and another function that imports vector back into network weights
flat_weights_placeholder = tf.placeholder(tf.float32, shape=(None,))
assigns = slice_vector(flat_weights_placeholder, weight_shapes)
load_flat_weights = [w.assign(ph) for w, ph in zip(weights, assigns)]
# -
# ##### Step 5: Main TRPO loop
#
# Here we will train our network!
# +
import time
from itertools import count
from collections import OrderedDict
max_kl=0.01 #this is hyperparameter of TRPO. It controls how big KL divergence may be between old and new policy every step.
cg_damping=0.1 #This parameters regularize addition to
numeptotal = 0 #this is number of episodes that we played.
start_time = time.time()
for i in count(1):
print ("\n********** Iteration %i ************" % i)
# Generating paths.
print("Rollout")
paths = rollout(env,act)
print ("Made rollout")
# Updating policy.
observations = np.concatenate([path["observations"] for path in paths])
actions = np.concatenate([path["actions"] for path in paths])
returns = np.concatenate([path["cumulative_returns"] for path in paths])
old_probs = np.concatenate([path["policy"] for path in paths])
inputs_batch=[observations,actions,returns,old_probs]
feed_dict = {observations_ph:observations,
actions_ph:actions,
old_probs_ph:old_probs,
cummulative_returns_ph:returns,
}
old_weights = sess.run(flat_weights)
def fisher_vector_product(p):
"""gets intermediate grads (p) and computes fisher*vector """
feed_dict[conjugate_grad_intermediate_vector] = p
return sess.run(fisher_vec_prod, feed_dict) + cg_damping * p
flat_grad = sess.run(flat_grad_surr, feed_dict)
stepdir = conjugate_gradient(fisher_vector_product, -flat_grad)
shs = .5 * stepdir.dot(fisher_vector_product(stepdir))
lm = np.sqrt(shs / max_kl)
fullstep = stepdir / lm
#Compute new weights with linesearch in the direction we found with CG
def losses_f(flat_weights):
feed_dict[flat_weights_placeholder] = flat_weights
sess.run(load_flat_weights, feed_dict)
return sess.run(losses, feed_dict)
new_weights = linesearch(losses_f, old_weights, fullstep, max_kl)
feed_dict[flat_weights_placeholder] = new_weights
sess.run(load_flat_weights, feed_dict)
#Report current progress
L_surr, kl, entropy = sess.run(losses, feed_dict)
episode_rewards = np.array([path["rewards"].sum() for path in paths])
stats = OrderedDict()
numeptotal += len(episode_rewards)
stats["Total number of episodes"] = numeptotal
stats["Average sum of rewards per episode"] = episode_rewards.mean()
stats["Std of rewards per episode"] = episode_rewards.std()
stats["Entropy"] = entropy
stats["Time elapsed"] = "%.2f mins" % ((time.time() - start_time)/60.)
stats["KL between old and new distribution"] = kl
stats["Surrogate loss"] = L_surr
for k, v in stats.items():
print(k + ": " + " " * (40 - len(k)) + str(v))
i += 1
# -
# # Homework option I: better sampling (10+pts)
#
# In this section, you're invited to implement a better rollout strategy called _vine_.
#
# 
#
# In most gym environments, you can actually backtrack by using states. You can find a wrapper that saves/loads states in [the mcts seminar](https://github.com/yandexdataschool/Practical_RL/blob/master/yet_another_week/seminar_MCTS.ipynb).
#
# You can read more about in the [TRPO article](https://arxiv.org/abs/1502.05477) in section 5.2.
#
# The goal here is to implement such rollout policy (we recommend using tree data structure like in the seminar above).
# Then you can assign cummulative rewards similar to `get_cummulative_rewards`, but for a tree.
#
# __bonus task__ - parallelize samples using multiple cores
# # Homework option II (10+pts)
#
# Let's use TRPO to train evil robots! (pick any of two)
# * [MuJoCo robots](https://gym.openai.com/envs#mujoco)
# * [Box2d robot](https://gym.openai.com/envs/BipedalWalker-v2)
#
# The catch here is that those environments have continuous action spaces.
#
# Luckily, TRPO is a policy gradient method, so it's gonna work for any parametric $\pi_\theta(a|s)$. We recommend starting with gaussian policy:
#
# $$\pi_\theta(a|s) = N(\mu_\theta(s),\sigma^2_\theta(s)) = {1 \over \sqrt { 2 \pi {\sigma^2}_\theta(s) } } e^{ (a -
# \mu_\theta(s))^2 \over 2 {\sigma^2}_\theta(s) } $$
#
# In the $\sqrt { 2 \pi {\sigma^2}_\theta(s) }$ clause, $\pi$ means ~3.1415926, not agent's policy.
#
# This essentially means that you will need two output layers:
# * $\mu_\theta(s)$, a dense layer with linear activation
# * ${\sigma^2}_\theta(s)$, a dense layer with activation tf.exp (to make it positive; like rho from bandits)
#
# For multidimensional actions, you can use fully factorized gaussian (basically a vector of gaussians).
#
# __bonus task__: compare performance of continuous action space method to action space discretization
|
week9_policy_II/seminar_TRPO_tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import os
sns.set_style('white')
sns.set_context('poster')
# -
# # Solutions: Model Validation and Optimization
# For some introduction, please see notebook 1. This notebook assumes some acquintance with the scikit-learn API. Much of the material here is adapted from [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook) by <NAME> and a [workshop at SciPY 2016](https://github.com/amueller/scipy-2016-sklearn) by <NAME> and <NAME>.
#
# We have been using train_test_split already to divide data between a training set to train the data and a test set to have an independent check of the trained model. In practice, you could imagine that sometimes, by pure chance, your crazy outliers will be in the test set, bringing your test score down, while in another random realization of the train_test_split this is not the case. You could wrongfully conclude you're overfitting.
#
# In order to keep all this all under control, machine learners often use k-fold cross validation. In the k folds, a fraction 1/k of the train data is used for testing and (1-1/k) for training. That is repeated k times. All data points are in the "test" set (in this case, it's called the validation set) exactly once, and k values of the test score (on the 'real' test data).
#
# Mostly to get some insight into the methods, we will simply create some cross-validation by hand. For now, forget about splitting training and testing first and using validation, but just do the cross validation using the whole iris data set as training set. Load the data and train a kNN classifier using 5-fold cross-validation, without making use of the sklearn.model_selection stuff. What are the scores, and their mean and standard deviation?
# Now that you know what is done, let's introduce sklearn's function for doing cross-validation:
from sklearn.model_selection import cross_val_score
# Try it! How many folds did it use? You can control that. Try reproducing your own result.
# There are a few more objects to import for cross-validation:
from sklearn.model_selection import KFold, StratifiedKFold, ShuffleSplit, StratifiedShuffleSplit
# All these are made to make your life easier. The function cross_val_score uses StratifiedKFold, which ensures that the target values are present in the same proportions in all subsamples. Please compare KFold and StratifiedKFold on the un-shuffled iris data set. Would there ever be a reason to use non-stratified?
# ## Model optimization: taming the hyperparameters
#
# Many, in fact almost all, machine learning models come with a suite of hyperparameters that control the behavior of the algorithm. Sometimes just a few are important, sometimes more. Sometimes one particular set of parameters, sometimes another. At some point you might develop some sort of intuition because you know the model well, and you understand your data, and you might naturally pick a decent value. Before that time (and to be sure, also at that time), you might want to search parameter space for the optimal set of hyperparameters for the current combination of algorithm and data.
#
#
# ### Validation and learning curves
#
# As an example of how to use validation and learning curves, we will create a data set and do polynomial regression. Polynomial regression is not a model in sklearn, but it turns out to be easy to do, using one of the data preprocessing methods (it will become linear regression to polynomial combinations of the features):
#
# +
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import make_pipeline
def PolynomialRegression(degree=2, **kwargs):
return make_pipeline(PolynomialFeatures(degree),
LinearRegression(**kwargs))
# +
def make_data(N, err=1.0, rseed=1):
# randomly sample the data
rng = np.random.RandomState(rseed)
X = rng.rand(N, 1) ** 2
y = 10 - 1. / (X.ravel() + 0.1)
if err > 0:
y += err * rng.randn(N)
return X, y
X, y = make_data(40)
# -
# Here is what it looks like, together with three polynomial fits:
# +
X_test = np.linspace(-0.1, 1.1, 500)[:, None]
plt.figure(figsize=(12,8))
plt.scatter(X.ravel(), y, color='black')
axis = plt.axis()
for degree in [1, 3, 8]:
y_test = PolynomialRegression(degree).fit(X, y).predict(X_test)
plt.plot(X_test.ravel(), y_test, label='degree={0}, training score={1:.2f}'.format(
degree, PolynomialRegression(degree).fit(X, y).score(X, y)))
plt.xlim(-0.1, 1.0)
plt.ylim(-2, 12)
plt.legend(loc='lower right');
# -
# Use sklearn.learning_curve.validation_curve to create the validation curves for this model, where the complexity parameter is the degree of polynomial.
#
# The validation curve needs to pass the polynomial degree to PolynomialFeatures. The pipeline automatically generates the name of this parameter: 'polynomialfeatures\_\_degree'.
#
# What is the optimal degree for fitting and how do you know?
#
# Is this a high bias or a high variance model?
#
# How does a nearest neighbor regressor compare?
# While validation curves look at the scores and their dependence on model complexity, learning curves investigate the dependence of both scores on the training set size.
#
# Why would need that is illustrated here. Use the same data set generator to make a sample that has 5 times more data points than the previous. While the model before started overfitting at roughly a twelfth-order polynomial, what is that like for this bigger data set? Use sklearn.model_selection.learning_curve to compare learning curves for a low and a high order polynomial, in the original, 40-point data set. Can you understand the results?
# With the same code as above run on X2, y2, you should find that overfitting occurs at much higher order polynomials. In the analysis below, also look at the variation with the number of folds!
# ### Grid search
#
# In practice, you will want to search a multi-dimensional grid of (hyper-)parameters to find your best model. Obviously, sklearn provides you with an easy interface to do just that: sklearn.model_selection.GridSearchCV. Us that, and its documentation to perform a grid search optimization of the iris data classifier that we started this notebook with. For that one, the number of neihgbors would be a reasonable parameter to play with. Try at least that, and if you can think of more, please go ahead. Cross-validate.
#
# What are the parameters of the best model? What is it's score? How can you make a script that after a grid search uses the best model of teh script to evaluate new data?
# ## Bonus exercise: text analytics and spam detection
#
# If you have some time left, are looking for something more fun or challenging or if you cheated your way through the solutions to get here at high speed, you may find this extra exercise amusing. It has no solutions, so you will have to do stuff yourself.
#
# There is a dataset included that contains a bunch of emails that are hand-classified to be either spam, or ham. Here is an outline of what needs to be done, with some hints, to get to a spam filter for this data set.
#
# - Read the data set. This is done for you, so as to avoid confusion.
# - Labels are fine. Words as features are not yet in great shape to run a classifier on. They will need to be transformed into numeric features. This comes down to counting words. Look for CountVectorizer and TfidfVectorizer to do so. What does this result in? What's the dimensionality of your problem? What does that imply for models?
# - Train a classifier of your choice. Investigate a few. Go through hyperparmeters of the classifier, but also of the vectorizers (mostly min_df and ngram_range).
# - Experiment with feature engineering. For example (but you should definitely feel free to come up with something yourself, coming up with things is aided by visualisation): length of the email.
#
# How pure can you get the detection (on data the classifier hasn't seen during training)? Can you identify words that are crucial in spam detection?
#
# Enjoy!
# +
with open(os.path.join("data", "SMSSpamCollection")) as f:
lines = [line.strip().split("\t") for line in f.readlines()]
text = [x[1] for x in lines]
y = [int(x[0] == "spam") for x in lines]
print("Labels are 0 for ham, 1 for spam!")
print()
print("Example, the first message is", lines[0][0], ":", text[0])
print("Example, the third message is", lines[2][0], ":", text[2])
# -
|
3_Model_Validation_Optimization_exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# <img src="logo.png" width="200">
#
# <center> <h1>Linear Regression with Gradient Descent Using a Made Up Example</h1> </center>
#
# + [markdown] deletable=true editable=true
# ## Gradient descent algorithm
# From our [video](https://youtu.be/fkS3FkVAPWU) on linear regression, we derived the equation to update the linear model parameters as:
#
#
# \begin{equation}
# \theta^{+} = \theta^{-} + \frac{\alpha}{m} (y_{i} - h(x_{i}) )\bar{x}
# \end{equation}
#
# This minimizes the following cost function
#
# \begin{equation}
# J(x, \theta, y) = \frac{1}{2m}\sum_{i=1}^{m}(h(x_i) - y_i)^2
# \end{equation}
#
# where
# \begin{equation}
# h(x_i) = \theta^T \bar{x}
# \end{equation}
# + [markdown] deletable=true editable=true
# ### Batch gradient descent
# ```FOR j FROM 0 -> max_iteration:
# FOR i FROM 0 -> m:
# theta += (alpha / m) * (y[i] - h(x[i])) * x_bar
# ENDLOOP
# ENDLOOP
# ```
# + [markdown] deletable=true editable=true
# ### Stochastic gradient descent
# ```shuffle(x, y)
# FOR i FROM 0 -> m:
# theta += (alpha / m) * (y[i] - h(x[i])) * x_bar
# ENDLOOP
# ```
# + deletable=true editable=true
import numpy as np
import matplotlib.pyplot as plt
# + deletable=true editable=true
"""Generate data"""
true_slope = 10.889
true_intercept = 3.456
input_var = np.arange(0.0,100.0)
output_var = true_slope * input_var + true_intercept + 500.0 * np.random.rand(len(input_var))
# + deletable=true editable=true
# %matplotlib notebook
plt.figure()
plt.scatter(input_var, output_var)
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# + deletable=true editable=true
def compute_cost(input_var, output_var, params):
"Compute linear regression cost"
num_samples = len(input_var)
cost_sum = 0.0
for x,y in zip(input_var, output_var):
y_hat = np.dot(params, np.array([1.0, x]))
cost_sum += (y_hat - y) ** 2
cost = cost_sum / (num_samples * 2.0)
return cost
# + deletable=true editable=true
def lin_reg_batch_gradient_descent(input_var, output_var, params, alpha, max_iter):
"""Compute the params for linear regression using batch gradient descent"""
iteration = 0
num_samples = len(input_var)
cost = np.zeros(max_iter)
params_store = np.zeros([2, max_iter])
while iteration < max_iter:
cost[iteration] = compute_cost(input_var, output_var, params)
params_store[:, iteration] = params
print('--------------------------')
print(f'iteration: {iteration}')
print(f'cost: {cost[iteration]}')
for x,y in zip(input_var, output_var):
y_hat = np.dot(params, np.array([1.0, x]))
gradient = np.array([1.0, x]) * (y - y_hat)
params += alpha * gradient/num_samples
iteration += 1
return params, cost, params_store
# + deletable=true editable=true
"""Train the model"""
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(input_var, output_var, test_size=0.20)
params_0 = np.array([20.0, 80.0])
alpha_batch = 1e-3
max_iter = 500
params_hat_batch, cost_batch, params_store_batch =\
lin_reg_batch_gradient_descent(x_train, y_train, params_0, alpha_batch, max_iter)
# + deletable=true editable=true
def lin_reg_stoch_gradient_descent(input_var, output_var, params, alpha):
"""Compute the params for linear regression using stochastic gradient descent"""
num_samples = len(input_var)
cost = np.zeros(num_samples)
params_store = np.zeros([2, num_samples])
i = 0
for x,y in zip(input_var, output_var):
cost[i] = compute_cost(input_var, output_var, params)
params_store[:, i] = params
print('--------------------------')
print(f'iteration: {i}')
print(f'cost: {cost[i]}')
y_hat = np.dot(params, np.array([1.0, x]))
gradient = np.array([1.0, x]) * (y - y_hat)
params += alpha * gradient/num_samples
i += 1
return params, cost, params_store
# + deletable=true editable=true
alpha = 1e-3
params_0 = np.array([20.0, 80.0])
params_hat, cost, params_store =\
lin_reg_stoch_gradient_descent(x_train, y_train, params_0, alpha)
# + deletable=true editable=true
plt.figure()
plt.scatter(x_test, y_test)
plt.plot(x_test, params_hat_batch[0] + params_hat_batch[1]*x_test, 'g', label='batch')
plt.plot(x_test, params_hat[0] + params_hat[1]*x_test, '-r', label='stochastic')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plt.show()
print(f'batch T0, T1: {params_hat_batch[0]}, {params_hat_batch[1]}')
print(f'stochastic T0, T1: {params_hat[0]}, {params_hat[1]}')
rms_batch = np.sqrt(np.mean(np.square(params_hat_batch[0] + params_hat_batch[1]*x_test - y_test)))
rms_stochastic = np.sqrt(np.mean(np.square(params_hat[0] + params_hat[1]*x_test - y_test)))
print(f'batch rms: {rms_batch}')
print(f'stochastic rms: {rms_stochastic}')
# + deletable=true editable=true
plt.figure()
plt.plot(np.arange(max_iter), cost_batch, 'r', label='batch')
plt.plot(np.arange(len(cost)), cost, 'g', label='stochastic')
plt.xlabel('iteration')
plt.ylabel('normalized cost')
plt.legend()
plt.show()
print(f'min cost with BGD: {np.min(cost_batch)}')
print(f'min cost with SGD: {np.min(cost)}')
# + deletable=true editable=true
|
notebooks/linear-regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Handwritten dataset
# +
# %run "../config/local.ipynb"
# %run "../utils/functions.ipynb"
# %config IPCompleter.greedy=True
import numpy as np
import pandas as pd
import sklearn
from sklearn.model_selection import StratifiedKFold
from src.mrcnn.utils import Dataset
# -
# ## Dataset class
class HandwrittenDataset(Dataset):
def __init__(self,
images_path,
labels_file_path,
classes=None,
stage: str = "train",
img_ids: np.array = None,
transforms=None,
size=(512,512)):
super().__init__()
# Add classes
self.init_classes(classes)
# input files paths
self.images_dir_path = images_path
self.labels_file_path = labels_file_path
# images ids
self.img_ids = img_ids
# data transformation function
self.transforms = transforms
# load referenced masks
self.load_masks_references()
# load the images
self.load_images()
def init_classes(self, classes):
""" add classes """
for item in classes:
self.add_class(item['source'], int(item['num']), item['name'])
def load_images(self):
""" Generate the images list
"""
# Add images
for idx in range(len(self.img_ids)):
self.add_image("pages", image_id=idx, path=os.path.join(self.images_dir_path, self.img_ids[idx]), name=self.img_ids[idx])
def load_masks_references(self):
# load all masks
all_masks = pd.read_csv(self.labels_file_path)
all_masks = all_masks.set_index(['name', 'num'])
# filter the masks by images ids
self.masks_references = all_masks.loc[self.img_ids].index.get_level_values(['name', 'num'])
print("finish load_masks_references")
def load_mask(self, image_id):
"""Generate instance masks for shapes of the given image ID.
"""
img = self.image_info[image_id]
# image name
name = img['name']
# check if the image is referenced
if not name in self.masks_references.index:
return None, []
# list of labels
labels = self.masks_references.loc[name]['label']
if type(labels) is str:
labels = [labels]
else:
labels = list(labels)
masks = np.array(list(refs.loc[name]['mask'].apply(lambda v: rle_to_mask(v, size[0], size[1]))))
# map the labels to the class indexes
class_ids = np.array([self.class_names.index(label) for label in labels])
# create the masks tensor
masks_tensor = np.array([mask.astype(np.bool) for mask in masks])
return np.stack(masks_tensor,axis=-1), class_ids.astype(np.int32)
def get_random_image(self, min_label_count=0):
""" return a filename and a (widht, height, RGB) array of a randmly selected image
with a label !
"""
# idx = random.randint(0,len(self.image_info)-1)
# img = self.image_info[idx]
# pick an image id from the masks references
# refs_idx = random.randint(0,len(self.masks_references.index)-1)
# img_id = self.masks_references.index[refs_idx]
# list of multi-label image
df = self.masks_references.reset_index()
df_a = df.groupby('name').count() >= min_label_count
multi_labels = list(df_a[df_a['label']].index)
# random selection of an image
ref_idx = random.randint(0,len(multi_labels))
img_id = multi_labels[ref_idx]
idx = [info['id'] for info in self.image_info if info['name'] == img_id][0]
return idx, self.load_image(idx)
def get_transformation(self, image_id):
# load the image
img = self.load_image(image_id)
# img = img.astype(np.uint8)
# get the mask
masks, class_ids = self.load_mask(image_id)
# image name
img_infos = self.image_info[image_id]
img_name = img_infos['name']
# img = img.astype(np.uint8)
data = {"image": img}
# add each mask for the transformation
for i in range(len(class_ids)):
idx = 'mask{}'.format(i)
mask = masks[:,:,i].astype(np.uint8)
data[idx] = mask
# m = masks[:,:,0].astype(np.uint8)
# apply augementation
transformer = self.__get_training_transformer(data)
augmented = transformer(**data)
# augmented image
img = augmented['image']
# list of augmented masks
tranformed_masks = [augmented[key].astype(bool) for key in augmented.keys() if not key == 'image']
return img, np.stack(tranformed_masks,axis=-1)
def __get_training_transformer(self, data):
""" Return the albumentation transformation function with respect to the number of masks
"""
train_transform = [
albu.HorizontalFlip(p=0.5),
albu.VerticalFlip(p=0.5),
albu.Blur(p=0.5),
albu.ShiftScaleRotate(
scale_limit=0.5,
rotate_limit=0,
shift_limit=0.1,
p=0.5,
border_mode=0
),
albu.GridDistortion(p=0.5),
]
target = {}
for i in range(len(data.keys()) -1):
target['mask' + str(i)] = 'mask'
return albu.Compose(train_transform, additional_targets=target)
# ## Train test split
def train_valid_split(masks_path, fold_num=0, n_folds=4, seed=42):
"""
return train and validation ids from the label file
parameters:
model_no: the model number if you train multiples models with the same data
n_folds: number of folds to create
"""
df_masks = pd.read_csv(masks_path)
# get count of label by file
refs = df_masks.groupby('name').count().reset_index()
# define the kfold generator
skfolds = StratifiedKFold(n_splits=n_folds,random_state=seed)
# create the folds
folds = [[train_idxs, valid_idxs] for train_idxs, valid_idxs in skfolds.split(refs['name'], refs['label'])]
sampled_train_ids = list(refs.iloc[folds[fold_num][0]]['name'])
sampled_valid_ids = list(refs.iloc[folds[fold_num][1]]['name'])
return sampled_train_ids, sampled_valid_ids
# ## Check dataset
train_ids, valid_ids = train_valid_split(MASKS_FILE)
print("train_ids: {} items, valid_ids: {} items".format(len(train_ids), len(valid_ids)))
# ## Build train and valid datasets
def build_train_valid_dataset(images_path, masks_path, classes, seed=42):
# get train and validation ids
train_ids, valid_ids = train_valid_split(masks_path=masks_path, seed=seed)
# Training dataset
dataset_train = HandwrittenDataset(images_path=images_path,
labels_file_path=masks_path,
classes=classes,
img_ids=train_ids,
transforms=get_training_augmentation())
dataset_train.prepare()
# Validation dataset
dataset_val = HandwrittenDataset(images_path=images_path,
labels_file_path=masks_path,
classes=classes,
img_ids=valid_ids,
transforms=get_training_augmentation())
dataset_val.prepare()
return dataset_train, dataset_val
# ## Check datasets
classes = [{'source':'clouds', 'num':1, 'name':'Fish'}]
train_ds, val_ds = build_train_valid_dataset(RESIZED_512x512_FEATURES_DIR, MASKS_FILE, classes)
train_ds.get_random_image()
|
notebooks/datasets/handwritten_dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Groupby and Pivot Tables in Python
import pandas as pd
# +
import pandas as pd
url = 'https://raw.githubusercontent.com/davidrkearney/Kearney_Data_Science/master/_notebooks/df_panel_fix.csv'
df = pd.read_csv(url, error_bad_lines=False)
#df
df.pivot_table(index='province', columns='year', values='it', aggfunc='mean').round(-1).style.highlight_max(color='blue').highlight_max(axis=1, color='green')
# -
df_subset = df[["year", "reg", "province", "gdp", "fdi", 'it',"specific"]]
df_subset.columns = ["year", "region", "province", "gdp", "fdi", 'it',"specific"]
df=df_subset
df
# +
# Add distributions by region
import matplotlib.pyplot as plt
#fig, axes = plt.subplots(nrows=3, ncols=3)
test_cells = ['East China', 'North China']
metrics = ['gdp', 'fdi', 'it']
for test_cell in test_cells:
for metric in metrics:
df.loc[df["region"] == test_cell].hist(column=[metric], bins=60)
print(test_cell)
print(metric)
# -
df.hist(column=['fdi'], bins=60)
# ## Distributions of Dependant Variables
# ### Right skew
df.hist(column=['fdi'], bins=60)
sns.histplot(df['fdi'])
sns.displot(df['gdp'])
sns.displot(df['fdi'])
sns.displot(df['it'])
sns.displot(df['specific'].dropna())
df.hist(column=['fdi'], bins=60)
# ## Removal of GDP value outliers more than 3 standard deviations away from the mean
# ## outlier removal of rows with GDP values that are > 3 standard deviations away form the mean
import scipy.stats as stats
df['gdp_zscore'] = stats.zscore(df['gdp'])
# ## these are the observations more then > 3 SDs away from the mean of gdp that will be dropped
df[abs(df['gdp_zscore'])>3].hist(column = ['gdp'])
df_no_gdp_outliers=df[abs(df['gdp_zscore'])<3]
df_no_gdp_outliers
df_no_gdp_outliers.hist(column=['gdp'], bins=60)
counts_fiscal=df.groupby('region').count()
counts_fiscal
counts_fiscal=df.groupby('province').count()
counts_fiscal
# +
#df_no_gdp_outliers.pivot_table(index='grouping column 1', columns='grouping column 2', values='aggregating column', aggfunc='sum')
# +
#pd.crosstab(df_no_gdp_outliers, 'year')
# -
df_no_gdp_outliers_subset = df_no_gdp_outliers[['region', 'gdp', 'fdi', 'it']]
df_no_gdp_outliers_subset
def aggregate_and_ttest(dataset, groupby_feature='province', alpha=.05, test_cells = [0, 1]):
#Imports
from tqdm import tqdm
from scipy.stats import ttest_ind_from_stats
metrics = ['gdp', 'fdi', 'it']
feature_size = 'size'
feature_mean = 'mean'
feature_std = 'std'
for metric in tqdm(metrics):
#print(metric)
crosstab = dataset.groupby(groupby_feature, as_index=False)[metric].agg(['size', 'mean', 'std'])
print(crosstab)
treatment = crosstab.index[test_cells[0]]
control = crosstab.index[test_cells[1]]
counts_control = crosstab.loc[control, feature_size]
counts_treatment = crosstab.loc[treatment, feature_size]
mean_control = crosstab.loc[control, feature_mean]
mean_treatment = crosstab.loc[treatment, feature_mean]
standard_deviation_control = crosstab.loc[control, feature_std]
standard_deviation_treatment = crosstab.loc[treatment, feature_std]
t_statistic, p_value = ttest_ind_from_stats(mean1=mean_treatment, std1=standard_deviation_treatment, nobs1=counts_treatment,mean2=mean_control,std2=standard_deviation_control,nobs2=counts_control)
#fstring to print the p value and t statistic
print(f"The t statistic of the comparison of the treatment test cell of {treatment} compared to the control test cell of {control} for the metric of {metric} is {t_statistic} and the p value is {p_value}.")
#f string to say of the comparison is significant at a given alpha level
if p_value < alpha:
print(f'The comparison between {treatment} and {control} is statistically significant at the threshold of {alpha}')
else:
print(f'The comparison between {treatment} and {control} is not statistically significant at the threshold of {alpha}')
aggregate_and_ttest(df_no_gdp_outliers, test_cells = [0,2])
EastvNorth=pd.DataFrame()
EastvNorth= aggregate_and_ttest(df_no_gdp_outliers_subset, test_cells = [0,1])
EastvNorth
# +
import numpy as np
import bootstrapped.bootstrap as bs
import bootstrapped.stats_functions as bs_stats
test_1=df_no_gdp_outliers[df_no_gdp_outliers['province']=='Beijing']
test=test_1['gdp'].to_numpy()
test
control_1=df_no_gdp_outliers[df_no_gdp_outliers['province']=='Shanxi']
control=control_1['gdp'].to_numpy()
control
# +
bins = np.linspace(0, 40, 20)
plt.hist(control, label='Control')
plt.hist(test, label='Test', color='orange')
plt.title('Test/Ctrl Data')
plt.legend()
# -
bs.bootstrap_ab(test, control, stat_func=bs_stats.sum, compare_func=bs_compare.percent_change)
# +
# run an a/b test simulation considering the lengths of the series (sum)
# consider the full 'volume' of values that are passed in
print(bs_compare.percent_change(test.sum(), control.sum()))
print(bs.bootstrap_ab(
test,
control,
stat_func=bs_stats.sum,
compare_func=bs_compare.percent_change
))
# +
# run an a/b test simulation ignoring the lengths of the series (average)
# just what is the 'typical' value
# use percent change to compare test and control
print(bs_compare.difference(test.mean(), control.mean()))
# -
print(bs.bootstrap_ab(test, control, bs_stats.mean, bs_compare.difference))
|
_notebooks/2021-05-18-groupby_pivot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import numpy as np
from IPython.core.interactiveshell import InteractiveShell
import random
InteractiveShell.ast_node_interactivity = "all"
import os
from datetime import datetime
import plotly.express as px
import glob
from tqdm import tqdm
# ## NASA FIRMS - Fire Information for Resource Management System
#
# 
#
# FIRMS distributes Near Real-Time (NRT) active fire data within 3 hours of satellite observation from the Moderate Resolution Imaging Spectroradiometer ([MODIS](https://modis.gsfc.nasa.gov/)) aboard the Aqua and Terra satellites, and the Visible Infrared Imaging Radiometer Suite ([VIIRS](https://www.jpss.noaa.gov/viirs.html)) aboard S-NPP and NOAA 20.
#
# ### Acknowledgement & Disclaimer
#
# We acknowledge the use of data and/or imagery from NASA's FIRMS (https://earthdata.nasa.gov/firms), part of NASA's Earth Observing System Data and Information System (EOSDIS).
#
# * Do not use for the preservation of life or property. Satellite-derived active fire / thermal anomalies have limited accuracy.
# * Active fire/thermal anomalies may be from fire, hot smoke, agriculture or other sources.
# * Cloud cover may obscure active fire detections.
#
# Please see the [official page](https://earthdata.nasa.gov/earth-observation-data/near-real-time/citation#ed-lance-disclaimer) for further details.
# ### Read and check the chunks
#
# The archive fire/hotspot datasets could be requested at
# https://firms.modaps.eosdis.nasa.gov/download/ in yearly chunks for each instrument.
#
# * MODIS Collection 6.1: Temporal Coverage: 11 November 2000 - present
# * VIIRS S-NPP 375m: Temporal Coverage: 20 January 2012 - present
# * VIIRS NOAA-20 375m: Temporal Coverage: 1 January 2020 - present
#
# Since NOAA-20 has less than 2 years data let's focus ont the other instruments.
t0 = datetime.now()
filenames = glob.glob('data/*/*.csv')
filenames
len(filenames)
# ### File stats
# +
rows = []
for f in tqdm(filenames):
df = pd.read_csv(f, parse_dates=['acq_time'], low_memory=False #, nrows=1000
)
csv_name = f.split('/')[-1]
row = [
f, csv_name, df.shape[0], df.shape[1], df.acq_date.min(), df.acq_date.max(),
df.satellite.max(), df.instrument.max(), df.version.max(),
df.latitude.nunique(), df.longitude.nunique(),
df.confidence.nunique(), df.satellite.nunique(), df.acq_date.nunique()
]
rows.append(row)
cols = [
'path', 'csv', 'rows', 'cols', 'start', 'end',
'satellite', 'instrument', 'version',
'lats', 'lons', 'confs', 'sats', 'days'
]
filestats = pd.DataFrame(rows, columns=cols)
filestats.sort_values(by=['start', 'instrument'])
# -
# ### Raw fire readings
#
# The satellite takes a ‘snapshot’ of events as it passes over the earth. Each hotspot/active fire detection represents the center of a pixel flagged as containing one or more fires, or other thermal anomalies (such as volcanoes). For MODIS the pixel is approximately 1km and for VIIRS the pixel is approximately 375m. The “location” is the center point of the pixel (not necessarily the coordinates of the actual fire).
df.head()
# ### Confidence
#
# The raw dataset has more detailed sensor measurements
#
# * **brightness**: Channel 21/22 brightness temperature of the fire pixel measured in Kelvin.
# * **bright_t31**: Channel 31 brightness temperature of the fire pixel measured in Kelvin.
# * **frp**: Fire Radiative Power depicts the pixel-integrated fire radiative power in MW (megawatts).
# * **type** Inferred hot spot type (0 = presumed vegetation fire, 1 = active volcano, 2 = other static land source, 3 = offshore)
# * **confidence** This value is based on a collection of intermediate algorithm quantities used in the detection process. It is intended to help users gauge the quality of individual hotspot/fire pixels. Confidence estimates range between 0 and 100% and are assigned one of the three fire classes (low-confidence fire, nominal-confidence fire, or high-confidence fire).
#
# For the baseline model we only keep the provided confidence values to filter less confident fire detection records.
dfs = []
for f in tqdm(filenames):
c = pd.read_csv(f, usecols=['confidence'], low_memory=False)
csv_name = f.split('/')[-1]
cnt = c.groupby('confidence').size().reset_index()
cnt['csv'] = csv_name
dfs.append(cnt)
confidences = pd.concat(dfs)
# # Process each chunk
#
# We removed fire readings with low or less than 50 confidence. For simplicity the coordinates are rounded to two decimal degrees. That is roughly 1.1 km at the Equator. For better spatial resolution the original VIIRS records could be used.
#
chunks = []
cols_to_read = ['latitude', 'longitude', 'acq_date', 'satellite', 'instrument', 'confidence']
for f in tqdm(filenames):
fire = pd.read_csv(f, usecols=cols_to_read, parse_dates=['acq_date'], low_memory=False)
if fire.satellite.loc[0] in ['Terra', 'Aqua', 'N']:
fire.latitude = fire.latitude.round(2)
fire.longitude = fire.longitude.round(2)
fire.confidence = fire.confidence.replace({'l': 0, 'n': 50, 'h': 100})
daily_fires = fire.groupby(
['latitude', 'longitude', 'acq_date', 'satellite', 'instrument']).confidence.max().reset_index()
daily_fires = daily_fires[daily_fires.confidence >= 50] # Remove low confidence records
instrument = fire.instrument.loc[0]
start = fire.acq_date.min()
print(instrument, start, fire.shape[0], daily_fires.shape[0])
daily_fires.to_csv(f'{instrument}_{start.strftime("%Y%m%d")}.csv', index=False)
chunks.append(daily_fires)
else:
'skip', f
daily_fires
full_dataset = pd.concat(chunks)
full_dataset.shape
full_dataset.head()
# full_dataset.to_csv('firms_fire_daily.csv.gz', index=False, compression='gzip')
AUS_LAT_RANGE = (-40, -9)
AUS_LON_RANGE = (112, 155)
aus = full_dataset[
(full_dataset.latitude > AUS_LAT_RANGE[0]) & (full_dataset.latitude < AUS_LAT_RANGE[1])]
aus = aus[
(aus.longitude > AUS_LON_RANGE[0]) & (aus.longitude < AUS_LON_RANGE[1])]
aus.shape
aus.to_csv('australia_fire_daily.csv.gz', index=False, compression='gzip')
end = datetime.now()
end.strftime('%Y-%m-%d %H:%M:%S')
f'Total time {(end - t0).seconds} (s)'
|
notebook/DataPreparation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright <NAME>/Attribution-NonCommercial 4.0 International (CC BY-NC 4.0)
# # MySQL Exercise 2: Using WHERE to select specific data
#
# When you are querying a business-related data set, you are usually doing so to answer a question about a subset of the data. In this lesson you will learn how to select subsets of rows of data that meet criteria you specify, to help you prepare for these types of business questions. The mechanism within a SQL query that allows you specify which subset of data you want to retrieve is the WHERE clause.
#
# Before we begin, let's load the SQL library and Dognition database, and make the Dognition database our default database. As a reminder, these are the lines of code you should input:
#
# ```python
# # %load_ext sql
# # %sql mysql://studentuser:studentpw@localhost/dognitiondb
# # %sql USE dognitiondb
# ```
# Recall the general syntax structure we learned from the "Introduction to Query Syntax" video at the beginning of the week:
#
# <img src="https://duke.box.com/shared/static/vnnubyrx8r46me7fmw1ayhcd8wn16wf8.jpg" width=400 alt="SELECT FROM WHERE" />
#
# This guide indicates that whenever the data we select need to meet certain criteria (specified using a "WHERE" clause), we specify those criteria after we have specified where the data come from.
#
# Let's say we want to know which Dognition customers received access to Dognition's first four tests for free. These customers have a 1 in the "free_start_user" column of the users table. The syntax you would use to select the data for these customers would be:
#
# ```mySQL
# SELECT user_guid
# FROM users
# WHERE free_start_user=1;
# ```
# (Note: user_guid is the field that specifies the unique User ID number of each customer in the users table)
#
# If you wanted to double-check that the outputted data indeed met the criteria you specified, you could include a second column in your output that would give you the value in the free_start_user field for each row of the output:
#
# ```mySQL
# SELECT user_guid, free_start_user
# FROM users
# WHERE free_start_user=1;
# ```
#
# Try this on your own below. Remember to use %%sql to indicate that your query will span multiple lines, and consider whether you would like to limit the number of results you ouput using the syntax we learned last lesson. If you do use a LIMIT statement, remember that it has to be the last item in your query, so this time you will place it after your WHERE statement instead of after your FROM statement.
# **Question 1: How would you select the Dog IDs for the dogs in the Dognition data set that were DNA tested (these should have a 1 in the dna_tested field of the dogs table)? Try it below (if you do not limit your output, your query should output data from 1433 dogs):**
# The SELECT statement can be used to interact with all data types, and there are many operators and functions that allow you to interact with the data in different ways. Here are some resources that describe these operators and functions:
#
# https://dev.mysql.com/doc/refman/5.7/en/sql-function-reference.html
# http://www.w3resource.com/mysql/mysql-functions-and-operators.php
#
#
# Some of the most common operators include: =,<,>,<=, and >=. If you want to select something that is NOT a specific value, use != or <>. You can also use logical operators, such as AND and OR.
#
# Let's start by examining how operators can be used with numerical data.
#
# If you wanted to examine the Dog IDs of dogs who weighed between 10 and 50 pounds, you could query:
#
# ```mySQL
# SELECT dog_guid, weight
# FROM dogs
# WHERE weight BETWEEN 10 AND 50;
# ```
#
# The above query provided an example of how to use the BETWEEN operator (described in the links provided above), as well as an example of how AND can be used to specify multiple criteria. If you wanted to examine the Dog IDs of dogs who were "fixed" (neutered) OR DNA tested, you could use OR in the following query:
#
# ```mySQL
# SELECT dog_guid, dog_fixed, dna_tested
# FROM dogs
# WHERE dog_fixed=1 OR dna_tested=1;
# ```
#
# If you wanted to examine the Dog IDs of dogs who were fixed but NOT DNA tested, you could query:
# ```mySQL
# SELECT dog_guid, dog_fixed, dna_tested
# FROM dogs
# WHERE dog_fixed=1 AND dna_tested!=1;
# ```
#
# **Question 2: How would you query the User IDs of customers who bought annual subscriptions, indicated by a "2" in the membership_type field of the users table? (If you do not limit the output of this query, your output should contain 4919 rows.)**
#
# Now let's try using the WHERE statement to interact with text data (called "strings").
#
# Strings need to be surrounded by quotation marks in SQL. MySQL accepts both double and single quotation marks, but some database systems only accept single quotation marks. Whenever a string contains an SQL keyword, the string must be enclosed in backticks instead of quotation marks.
#
# >` 'the marks that surrounds this phrase are single quotation marks' `
# >` "the marks that surrounds this phrase are double quotation marks" `
# >`` `the marks that surround this phrase are backticks` ``
#
# Strings enclosed in quotation or backticks can be used with many of the same operators as numerical data. For example, imagine that you only wanted to look at data from dogs of the breed "Golden Retrievers." You could query (note that double quotation marks could have been used in this example is well):
#
# ```mySQL
# SELECT dog_guid, breed
# FROM dogs
# WHERE breed='golden retriever';
# ```
#
# The IN operator allows you to specify multiple values in a WHERE clause. Each of these values must be separated by a comma from the other values, and the entire list of values should be enclosed in parentheses. If you wanted to look at all the data from Golden Retrievers and Poodles, you could certainly use the OR operator, but the IN operator would be even more efficient (note that single quotation marks could have been used in this example, too):
#
# ```mySQL
# SELECT dog_guid, breed
# FROM dogs
# WHERE breed IN ("golden retriever","poodle");
# ```
#
# The LIKE operator allows you to specify a pattern that the textual data you query has to match. For example, if you wanted to look at all the data from breeds whose names started with "s", you could query:
#
# ```mySQL
# SELECT dog_guid, breed
# FROM dogs
# WHERE breed LIKE ("s%");
# ```
#
# In this syntax, the percent sign indicates a wild card. Wild cards represent unlimited numbers of missing letters. This is how the placement of the percent sign would affect the results of the query:
#
# + WHERE breed LIKE ("s%") = the breed must start with "s", but can have any number of letters after the "s"
# + WHERE breed LIKE ("%s") = the breed must end with "s", but can have any number of letters before the "s"
# + WHERE breed LIKE ("%s%") = the breed must contain an "s" somewhere in its name, but can have any number of letters before or after the "s"
#
# **Question 3: How would you query all the data from customers located in the state of North Carolina (abbreviated "NC") or New York (abbreviated "NY")? If you do not limit the output of this query, your output should contain 1333 rows. **
#
#
# Next, let's try using the WHERE statement to interact with datetime data. Time-related data is a little more complicated to work with than other types of data, because it must have a very specific format. MySQL comes with the following data types for storing a date or a date/time value in the database:
#
# DATE - format YYYY-MM-DD
# DATETIME - format: YYYY-MM-DD HH:MI:SS
# TIMESTAMP - format: YYYY-MM-DD HH:MI:SS
# YEAR - format YYYY or YY
#
# One of the interesting things about time-related data is that SQL has commands to break the data into different "time parts" or "date parts" as described here:
#
# http://www.tutorialspoint.com/mysql/mysql-date-time-functions.htm
#
#
# A time stamp stored in one row of data might look like this:
#
# ```
# 2013-02-07 02:50:52
# ```
# The year part of that entry would be 2013, the month part would be "02" or "February" (depending on the requested format), the seconds part would be "52", and so on. SQL functions easily allow you to convert those parts into formats you might need for specific analyses. For example, imagine you wanted to know how many tests Dognition customers complete on different days of the week. To complete this analysis, you would need to convert the time stamps of each completed test to a variable that outputted the correct day of the week for that date. DAYNAME is a function that will do this for you. You can combine DAYNAME with WHERE to select data from only a single day of the week:
#
# ```mySQL
# SELECT dog_guid, created_at
# FROM complete_tests
# WHERE DAYNAME(created_at)="Tuesday"
# ```
#
# You can also use common operators like =,<,>,<=,>=,!=, or <> with dates just like you would with other types of data, but whether you refer to the date as a number or text will depend on whether you are selecting individual date parts or treating the date/time entry as a single clause. For example, you could select all the Dog IDs and time stamps of tests completed after the 15 of every month with this command that extracts the "DAY" date part out of each time stamp:
#
# ```mySQL
# SELECT dog_guid, created_at
# FROM complete_tests
# WHERE DAY(created_at) > 15
# ```
#
# You could also select all the Dog IDs and time stamps of completed tests from after February 4, 2014 by treating date entries as text clauses with the following query:
#
# ```mySQL
# SELECT dog_guid, created_at
# FROM complete_tests
# WHERE created_at > '2014-02-04'
# ```
#
# Note that you have to use a different set of functions than you would use for regular numerical data to add or subtract time from any values in these datetime formats. For example, instead of using a minus sign to find the difference in time between two time stamps or dates, you would use the TIMEDIFF or DATEDIFF function. See the references provided above for a list of these functions.
#
# **Question 4: Now that you have seen how datetime data can be used to impose criteria on the data you select, how would you select all the Dog IDs and time stamps of Dognition tests completed before October 15, 2015 (your output should have 193,246 rows)?**
# Last, let's use the WHERE statement in combination with two very important operators: IS NULL and IS NOT NULL. IS NULL will indicate rows of data that have null values. IS NOT NULL will indicate rows that do not have null values. We saw in previous exercises that many of the entries in the free_start_user field of the user table in the Dognition data set had NULL values. To select only the rows that have non-null data you could query:
#
# ```mySQL
# SELECT user_guid
# FROM users
# WHERE free_start_user IS NOT NULL;
# ```
#
# To select only the rows that have null data so that you can examine if these rows share something else in common, you could query:
#
# ```mySQL
# SELECT user_guid
# FROM users
# WHERE free_start_user IS NULL;
# ```
#
# **Question 5: How would you select all the User IDs of customers who do not have null values in the State field of their demographic information (if you do not limit the output, you should get 17,985 from this query -- there are a lot of null values in the state field!)?**
# ## Practice writing your own SELECT and WHERE statements!
#
# ### These queries will combine what you've learned in the past two lessons.
#
# **Question 6: How would you retrieve the Dog ID, subcategory_name, and test_name fields, in that order, of the first 10 reviews entered in the Reviews table to be submitted in 2014?**
#
# ** Question 7: How would you select all of the User IDs of customers who have female dogs whose breed includes the word "terrier" somewhere in its name (if you don't limit your output, you should have 1771 rows in your output)? **
# **Question 8: How would you select the Dog ID, test name, and subcategory associated with each completed test for the first 100 tests entered in October, 2014?**
# There are many more operators you can use in your WHERE clauses to restrict the data you select as well. We do not have the space to go over each one individually in this lesson, but I encourage you to explore them on your own. <mark>*This is a great area to practice being fearless and bold in your desire to learn new things! The more you try, the more you will learn.*</mark>
#
# **Feel free to practice any other functions or operators you discover in the space below:**
|
MySQL_exercises/.ipynb_checkpoints/MySQL_Exercise_02_Selecting_Data_Subsets_using_WHERE-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import holoviews as hv
hv.extension('bokeh')
import numpy as np
import scipy.signal
# # Bayesian modeling
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Introduction
#
# In this lecture we return to parametric modeling but using the bayesian approach.
#
# A summary of the bayesian premise
#
# - Inference is made by producing probability density functions (pdf): **posterior**
# - We model the uncertainty of the data, experiment, parameters, etc. as a **joint pdf**
# - The parameter vector $\theta$ is a R.V., *i.e.* it follows a distribution: **prior**
#
# The Bayes theorem and the law of total probability tell us
#
# $$
# p(\theta| \{x\}) = \frac{p(\{x\}, \theta)}{p(\{x\})}= \frac{p(\{x\}|\theta) p(\theta)}{\int p(\{x\}|\theta) p(\theta) d\theta} \propto p(\{x\}|\theta) p(\theta),
# $$
#
#
#
# -
# :::{note}
#
# The posterior is build from the **likelihood**, **prior** and **evidence** (marginal data likelihood), *i.e.* the posterior can be small if either the likelihood or the prior are small
#
# :::
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# **Why/When should I use the Bayesian formalism?**
#
# In many cases bayesian inference will not differ much from frequentist techniques. Also, in general, bayesian inference is harder to compute and requires more sophisticated methods
#
# But bayesian modeling gives us some key advantages:
#
# - We know the uncertainty of our parameters/predictions, i.e. and we can take more informed decisions
# - It gives a principled way of injecting prior knowledge (regularization)
# - We can integrate unknown or missing (nuisance) parameters
#
# + [markdown] slideshow={"slide_type": "subslide"}
# The following is a summary of the Bayesian inference procedure
#
# 1. Formulate your problem: likelihood and prior
# 1. Build a joint distribution (relation of all parameters)
# 1. Determine the posterior using Bayes Theorem. Find MAP and credible regions
# 1. Test your hypothesis
# 1. **Criticize:** Evaluate how appropriate the model is and suggest improvements
#
# We will review these steps in this lesson
# + [markdown] slideshow={"slide_type": "slide"}
# ## Maximum *a posteriori* (MAP) estimation
#
# In the Bayesian setting the best "point estimate" of the parameters of the model is given by the MAP
#
# $$
# \hat \theta = \text{arg} \max_\theta p(\theta|\{x\}) = \text{arg} \max_\theta p(\{x\}| \theta) p(\theta),
# $$
#
# where we "omit" the evidence (denominator in Bayes rule) because it does not depend on $\theta$
#
# Applying the logarithm (monotonic) we can decouple the likelihood from the prior
#
# $$
# \hat \theta = \text{arg} \max_\theta \log p(\{x\}| \theta) + \log p(\theta),
# $$
#
# :::{note}
#
# MAP is still a point estimate: poor's man Bayes
#
# :::
#
# The main difference to what we saw in previous lessons is **the prior**
# + [markdown] slideshow={"slide_type": "subslide"}
# ### What can I do with priors?
#
# Priors are distributions that summarize what we know about the parameters before-hand, for example
#
# - a parameter is continuous and has no bounds: Normal
# - a parameter is continuous and positive: Lognormal, Inverse gamma, Half-normal, etc
# - a parameter is positive-semidefinite: Inverse Wishart, LKJ, etc
# - a parameter is in the simplex: Dirichlet
#
# Priors can be described as
#
# - Informative: $\mathcal{N}(\theta|\mu=5.4, \sigma^2=0.1)$
# - Weakly informative: $\mathcal{N}(\theta|\mu=0, \sigma^2=100.)$
# - Uninformative (or objective): My parameter is positive
#
# Of course these notions depend on the problem at hand.
#
# We should select priors that
#
# - add a positive weight on values that may occur
# - put zero weight to impossible values
# - help regularize the solution
#
# Later we will see the case of **conjugate prior**, which are very convenient from a computational point of view
#
# I suggest reading the [practical principles for choosing priors](https://github.com/stan-dev/stan/wiki/Prior-Choice-Recommendations) in the Stan repository
#
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# ### Example: MAP estimate of the mean of a Gaussian distribution
#
# Assuming $N$ i.i.d samples and a Gaussian likelihood with known variance we can write
#
# $$
# \log p(\{x\}|\theta) = \log L (\mu) = - \frac{N}{2} \log 2\pi\sigma^2 - \frac{1}{2\sigma^{2}} \sum_{i=1}^N (x_i-\mu)^2,
# $$
#
# In this particular example we will select a Gaussian prior with parameters $\mu_0$ and $\sigma_0$ for $\mu$
#
# $$
# \log p(\theta) = -\frac{1}{2} \log 2 \pi \sigma^2_0 - \frac{1}{2 \sigma^2_0} (\mu - \mu_0)^2,
# $$
#
# Adding the log likelihood and log prior and taking the derivative
#
# $$
# \frac{d}{d\mu} \log p(\{x\}|\theta) + \log p(\theta) = \frac{1}{\sigma^{2}} \sum_{i=1}^N (x_i-\mu) - \frac{1}{ \sigma^2_0} (\mu - \mu_0),
# $$
#
# then setting the derivative equal to zero gives us the MAP estimate
#
# $$
# \hat \mu_{\text{map}} = \left(\frac{N}{\sigma^2} + \frac{1}{\sigma^2_0} \right)^{-1} \left(\frac{N}{\sigma^2} \bar x + \frac{1}{\sigma^2_0} \mu_0 \right),
# $$
#
# where $\bar x = \frac{1}{N} \sum_{i=1}^N x_i$.
#
# :::{important}
#
# Do not confuse $\sigma^2$ (the likelihood/noise variance) and $\sigma^2_0$ (prior variance)
#
# :::
#
# (Using a bit of algebra) we can write the MAP expression as
#
# $$
# \begin{align}
# \hat \mu_{\text{map}} &= \left(\frac{N}{\sigma^2} + \frac{1}{\sigma^2_0} \right)^{-1} \left(\frac{N\bar x}{\sigma^2} + \frac{\mu_0}{\sigma^2_0} \right) \nonumber \\
# &= \frac{N \bar x \sigma^2_0 + \mu_0 \sigma^2}{N\sigma^2_0+ \sigma^2} \nonumber \\
# &= \frac{\bar x + \mu_0 \frac{\sigma^2}{\sigma^2_0 N}}{1 + \frac{\sigma^2}{\sigma^2_0 N}} \nonumber \\
# &= w \bar x + (1-w) \mu_0, \qquad \text{where} \quad w = \frac{1}{1 + \frac{\sigma^2}{\sigma^2_0 N}} \nonumber
# \end{align}
# $$
#
# > The MAP estimate of $\mu$ is a weighted average between $\mu_0$ (prior) and $\bar x$ (the MLE solution)
#
# :::{note}
#
# In the last expression:
#
# - if either $\sigma^2_0 \to \infty$ or $N \to \infty$ then $w\to1$, i.e. the MAP converges to the MLE solution
# - the prior is more relevant if have a few sample (small $N$) or a noisy samples (large $\sigma^2$)
#
# :::
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Extra: MAP intepretation as a penalized MLE/regularized LS
#
# We can rewrite the MAP optimization problem for a Gaussian likelihood with known variance and a zero-mean Gaussian prior as
#
# $$
# \begin{align}
# \hat \mu_{\text{map}} &= \text{arg} \max_\mu \log p(\{x\}| \mu, \sigma^2) + \log p(\mu) \nonumber \\
# &= \text{arg} \max_\mu - \frac{N}{2} \log 2\pi\sigma^2 - \frac{1}{2\sigma^{2}} \sum_{i=1}^N (x_i-\mu)^2 - \frac{1}{2\sigma_0^2} \mu^2 \nonumber \\
# &= \text{arg} \min_\mu \frac{1}{2\sigma^{2}} \sum_{i=1}^N (x_i-\mu)^2 + \frac{1}{2\sigma_0^2} \mu^2 \nonumber \\
# &= \text{arg} \min_\mu \|x-\mu\|^2 + \lambda \|\mu \|^2, \nonumber
# \end{align}
# $$
#
# where $\lambda = \frac{\sigma^2}{\sigma_0^2}$.
#
# We can recognize the last equation as a regularized least squares problem. In this case a using a Gaussian priors is equivalent to using a L2 norm regularizar on the parameters (this is known as ridge regression). A Laplacian prior yields a L1 regularizer (LASSO) [^lasso]
#
# [^lasso]: Hastie, Tibshirani, Friedman, chapter 3.4 (Shrinkage methods), page 61.
#
# We will review ridge regression in a future lecture
# -
# ## Analytical posterior with conjugate priors
#
# Remember that the MAP is only a point estimate. In a fully-bayesian setting want we are interested in is the posterior of the parameter
#
# In the particular case of a Gaussian likelihod and a Gaussian prior we can rearrange the terms to show that
#
# $$
# \begin{align}
# p(\theta |\{x\}) &\propto p(\{x\} |\theta ) p(\theta ) \nonumber \\
# &\propto \exp \left ( \frac{1}{2\sigma^2} \sum_{i=1}^N (x_i - \mu)^2 \right) \exp \left ( \frac{1}{2\sigma_0^2} (\mu - \mu_0)^2 \right) \nonumber \\
# &\propto \exp \left ( -\frac{1}{2 \hat \sigma^2} (\mu - \hat \mu_{\text{map}} )^2 \right), \nonumber
# \end{align}
# $$
#
# where
#
# $$
# \hat \sigma^2 = \left(\frac{N}{\sigma^2} + \frac{1}{\sigma^2_0} \right)^{-1},
# $$
#
# i.e. the posterior has a closed analytical form and is also Gaussian [^gaussmult]
#
# [^gaussmult]: Another way to show that the posterior is Gaussian is to use the [property of Gaussian pdf multiplication](http://www.tina-vision.net/docs/memos/2003-003.pdf)
#
# > When the resulting posterior has the same distribution as the specified prior we say that the prior is a **conjugate prior for the specified likelihood**
#
# In this particular case the Gaussian distribution is conjugate with itself
#
# Other examples are:
#
# ```{list-table} Conjugacy table
# :header-rows: 1
#
# * - Likelihood
# - Conjugate prior
# * - Bernoulli
# - Beta
# * - Poisson
# - Gamma
# * - Multinomial or categorial
# - Dirichlet
# * - Exponential
# - Gamma
# * - Normal with unknown variance
# - Normal-inverse gamma (NIG)
# * - Multivariate normal with unknown covariance
# - Normal-inverse Wishart
# ```
#
#
# ### Interactive example
#
# We generate Gaussian distributed data with $\mu=2$ and $\sigma=1$ and plot the asymptotic distribution of the MLE (yellow) and the analytical posterior (red) and the prior (blue)
# +
from scipy.stats import norm
def mle_mu(xi: np.array) -> float:
return np.mean(xi)
def asymptotic_mle(x: np.array, xi: np.array, s2: float) -> np.array:
N = len(xi)
return norm(loc=mle_mu(xi), scale=np.sqrt(s2/N)).pdf(x)
def map_mu(xi: np.array, mu0: float, s20: float, s2: float):
N = len(xi)
w = (N*s20)/(N*s20 + s2)
return mle_mu(xi)*w + mu0*(1. - w)
def prior_mu(x: np.array, mu0: float, s20: float) -> np.array:
return norm(loc=mu0, scale=np.sqrt(s20)).pdf(x)
def posterior_mu(x: np.array, xi: np.array, mu0: float, s20: float, s2: float) -> np.array:
N = len(xi)
s2_pos = s2*s20/(N*s20 + s2)
mu_pos = map_mu(xi, mu0, s20, s2)
return norm(loc=mu_pos, scale=np.sqrt(s2_pos)).pdf(x)
# -
# Explore
#
# - What happens with $N$ grows?
# - What happens when $\sigma_0$ grows?
# +
mu_real, s2_real = 2., 1.
x_plot = np.linspace(-5, 5, num=1000)
true_value = hv.VLine(mu_real).opts(color='k', line_width=2, alpha=0.5)
hmap = hv.HoloMap(kdims=['N', 'mu0', 's20'])
for N in [1, 5, 10, 50, 100, 500]:
for mu0 in np.linspace(-3, 3, num=5):
for s20 in np.logspace(-1, 1, num=3):
data = norm(loc=mu_real, scale=np.sqrt(s2_real)).rvs(N, random_state=1234)
plot_prior = hv.Curve((x_plot, prior_mu(x_plot, mu0, s20)), 'x', 'density', label='prior')
plot_mle = hv.Curve((x_plot, asymptotic_mle(x_plot, data, s2_real)), label='MLE')
plot_post = hv.Curve((x_plot, posterior_mu(x_plot, data, mu0, s20, s2_real)), label='posterior')
hmap[(N, mu0, s20)] = (plot_prior * plot_post * plot_mle * true_value).opts(hv.opts.Curve(width=500))
hmap
# -
# ### Conjugate prior for Gaussian likelihood when $\sigma^2$ is unknown
#
# Before we assumed that $\sigma^2$ was a known quantity and we focused on estimating $\mu$
#
# If we now assume that the mean $\mu$ is known and the variance is unknown then the conjugate prior for the variance is an inverse-Gamma distribution
#
# $$
# p(\sigma^2) = \text{IG}(\sigma^2| \alpha_0, \beta_0) = \frac{\beta_0^{\alpha_0}}{\Gamma(\alpha_0)} x^{-\alpha_0-1} e^{-\frac{\beta_0}{x}}
# $$
#
# With which the resulting posterior is also
#
# $$
# \text{IG}\left(\sigma^2| \alpha_N , \beta_N \right),
# $$
#
# where
#
# - $ \alpha_N = \alpha_0 + N/2$
# - $\beta_N = \beta_0 + \frac{1}{2} \sum_{i=1}^N (x_i - \mu)^2$
#
# As both $\alpha$ and $\beta$ encode the strength of the prior the following parameterization is broadly used
#
# $$
# p(\sigma^2) = \text{IG}(\sigma^2| \alpha, \beta) = \text{IG}\left(\sigma^2 \bigg| \frac{\nu}{2}, \frac{\nu \sigma_0^2}{2}\right)
# $$
#
# where $\sigma_0^2$ controls the value of the prior and $\nu$ the strength. Note that this is also closely related to the [inverse chi-square distribution](https://en.wikipedia.org/wiki/Inverse-chi-squared_distribution)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Conjugate prior for Gaussian likelihood when both $\mu$ and $\sigma^2$ are unknown
#
# Multiplying the normal prior and the IG prior does not yield a conjugate prior (assumes independence of $\mu$ and $\sigma$). In this case the conjugate prior is hierarchical
#
# $$
# \begin{align}
# p(x_i|\mu, \sigma^2) &= \mathcal{N}(\mu, \sigma^2) \nonumber \\
# p(\mu|\sigma^2) &= \mathcal{N}(\mu_0, \sigma^2/\lambda_0) \nonumber \\
# p(\sigma^2) &= \text{IG}(\alpha, \beta) \nonumber
# \end{align}
# $$
#
# which is called **normal-inverse-gamma (NIG)**, a four parameter distribution
#
# The NIG prior is
#
# $$
# p(\mu, \sigma^2) = \text{NIG}(\mu_0, \lambda_0, \alpha_0, \beta_0) = \mathcal{N}(\mu|\mu_0 , \sigma^2/\lambda_0) \text{IG}(\sigma^2|\alpha_0, \beta_0)
# $$
#
# An the posterior is also NIG
#
# $$
# p(\mu, \sigma^2|\{x\}) = \text{NIG}(\mu_n, \lambda_n, \alpha_n, \beta_n)
# $$
#
# where
#
# - $\lambda_n = \lambda_0 + N$
# - $\mu_n = \lambda_n^{-1} \left ( \lambda_0 \mu_0 + N \bar x \right)$
# - $\alpha_n = \alpha_0 + N/2$
# - $\beta_n = \beta_0 + 0.5\mu_0^2\lambda_0 + 0.5\sum_i x_i^2 - 0.5\lambda_n \mu_n^2$
# -
# ## Describing the posterior using Credible Interval (CI) and the High Posterior Density (HPD) regions
#
# One way to summarize the posterior is to measure its **width**
#
# The $100(1-\alpha)$ % CI of $\theta$ is a contiguous region $[\theta_{l}, \theta_{u}]$ such that
#
# $$
# P(\theta_{l}< \theta < \theta_{u}) = 1 - \alpha
# $$
#
# We have to either know the functional form of the posterior (analytical) or have a posterior from which we can sample from (this is the case if we are using MCMC)
#
# The HPD is an alternative to CI that is better when we have multiple modes. The HPD depends not only on the width but also on the height of the posterior. The following figure shows the difference between them
#
#
# <img src="img/HPD.png">
#
# ### Example
#
# The 95% CI for the previous example for a given combination of $\mu_0$, $\sigma_0^2$ and $N$ is
# +
mu0, s20, N = 0., 10., 100
data = norm(loc=mu_real, scale=np.sqrt(s2_real)).rvs(N, random_state=12345)
N = len(data)
s2_pos = s2_real*s20/(N*s20 + s2_real)
mu_pos = map_mu(data, mu0, s20, s2_real)
dist = norm(loc=mu_pos, scale=np.sqrt(s2_pos))
display(f'95 % CI for mu: [{dist.ppf(0.025):0.4f}, {dist.ppf(0.975):0.4f}]')
# + [markdown] slideshow={"slide_type": "skip"}
# ### Extra: Mean of the posterior
#
# Other point estimate that can be used to characterize the posterior is
#
# $$
# \hat \theta = \mathbb{E}[\theta|\{x\}] = \int \theta p(\theta| \{x\}) d\theta,
# $$
#
# *i.e.* the mean or expected value of the posterior
# + [markdown] slideshow={"slide_type": "slide"}
# ## Help: My posterior does not have an analytical form
#
# In this case we resort to either variational inference (VI) or Markov Chain Monte Carlo (MCMC) methods
#
# We will learn how to use MCMC to sample from intractable posterior distributions in a future lesson
# -
|
lectures/2_statistical_modeling/part3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def allInOne(path):
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
def filterArraysToShortest(arr1, arr1Pos, arr2, arr2Pos):
both_pos = np.logical_and(arr1Pos, arr2Pos)
arr1Filt = arr1[both_pos]
arr2Filt = arr2[both_pos]
return arr1Filt, arr2Filt
def filterArraysToShortest2(arr1, arr1Pos, arr2, arr2Pos):
if np.count_nonzero(arr1Pos) < np.count_nonzero(arr2Pos):
arr1Filt = arr1[arr1Pos]
arr2Filt = arr2[arr1Pos]
else:
arr1Filt = arr1[arr2Pos]
arr2Filt = arr2[arr2Pos]
return arr1Filt, arr2Filt
df = pd.read_csv(path)
display(df.head(5))
reps = df['rep'].to_numpy()
hopsRanked = df['hopsRanked'].to_numpy()
hopsUnranked = df['hopsUnranked'].to_numpy()
hopsRankedOne = df['hopsRankedOne'].to_numpy()
hopsPath = df['hopsPath'].to_numpy()
hopsRankedPositive = hopsRanked > -1
hopsUnrankedPositive = hopsUnranked > -1
hopsRankedOnePositive = hopsRankedOne > -1
hopsPathPositive = hopsPath > -1
total = len(df.index)
successesRanked = np.count_nonzero(hopsRanked > -1)
successesRankedOne = np.count_nonzero(hopsRankedOne > -1)
successesUnranked = np.count_nonzero(hopsUnranked > -1)
successesPath = np.count_nonzero(hopsPath > -1)
print("Resilience Paths: {}%".format((successesPath / total) * 100))
print("Resilience One Tree: {}%".format((successesRankedOne / total) * 100))
print("Resilience Trees: {}%".format((successesRanked / total) * 100))
#print("Resilience Unranked: {}%".format((successesUnranked / total) * 100))
hopsPathFiltered,hopsRankedFiltered = filterArraysToShortest(hopsPath, hopsPathPositive, hopsRanked, hopsRankedPositive)
diff = np.subtract(hopsPathFiltered,hopsRankedFiltered)
plt.scatter(np.arange(len(hopsPathFiltered)), diff, color="blue", s=1.5)
plt.show()
higherZero = np.count_nonzero(diff > 0)
lowerZero = np.count_nonzero(diff < 0)
equalZero = np.count_nonzero(diff == 0)
print("Trees beat paths in {} runs".format(higherZero))
print("Paths beat trees in {} runs".format(lowerZero))
print("Both performed equally good in {} runs".format(equalZero))
hopsPathFiltered,hopsRankedOneFiltered = filterArraysToShortest(hopsPath, hopsPathPositive, hopsRankedOne, hopsRankedOnePositive)
diff = np.subtract(hopsPathFiltered,hopsRankedOneFiltered)
plt.scatter(np.arange(len(hopsPathFiltered)), diff, color="blue", s=1.5);
plt.show()
higherZero = np.count_nonzero(diff > 0)
lowerZero = np.count_nonzero(diff < 0)
equalZero = np.count_nonzero(diff == 0)
print("One Tree beat paths in {} runs".format(higherZero))
print("Paths beat one tree in {} runs".format(lowerZero))
print("Both performed equally good in {} runs".format(equalZero))
hopsRankedOneFiltered,hopsRankedFiltered = filterArraysToShortest(hopsRankedOne, hopsRankedOnePositive, hopsRanked, hopsRankedPositive)
diff = np.subtract(hopsRankedOneFiltered,hopsRankedFiltered)
plt.scatter(np.arange(len(hopsRankedFiltered)), diff, color="blue", s=1.5);
plt.show()
higherZero = np.count_nonzero(diff > 0)
lowerZero = np.count_nonzero(diff < 0)
equalZero = np.count_nonzero(diff == 0)
print("Multiple Trees beat One Tree in {} runs".format(higherZero))
print("One Tree beat multiple trees in {} runs".format(lowerZero))
print("Both performed equally good in {} runs".format(equalZero))
hopsUnrankedFiltered,hopsRankedFiltered = filterArraysToShortest(hopsUnranked, hopsUnrankedPositive, hopsRanked, hopsRankedPositive)
diff = np.subtract(hopsUnrankedFiltered,hopsRankedFiltered)
plt.scatter(np.arange(len(hopsRankedFiltered)), diff, color="blue", s=1.5);
plt.show()
higherZero = np.count_nonzero(diff > 0)
lowerZero = np.count_nonzero(diff < 0)
equalZero = np.count_nonzero(diff == 0)
print("Ranked Trees beat Unranked Tree in {} runs".format(higherZero))
print("Unranked Tree beat ranked trees in {} runs".format(lowerZero))
print("Both performed equally good in {} runs".format(equalZero))
# ## CLUSTERED, 60%, -30% per hop
allInOne("CSVs/trees-2021-08-02_10-54-26-n25.csv")
# ## CLUSTERED, 50%, -15% per hop
allInOne('CSVs/trees-2021-07-25_14-24-23-p0.05-n100.csv')
# ## Adversarial failures around D (varying failure rate)
allInOne('CSVs/trees-2021-07-24_19-48-33-p0.05-n100.csv')
# ## Fixed adversarial failure rate 80%
allInOne('CSVs/trees-2021-07-25_14-37-35-p0.05-n100.csv')
# ## AFTER BUGFIX
allInOne('CSVs/trees-2021-08-08_13-05-29-n25.csv')
allInOne('CSVs/trees-2021-08-08_13-09-49-n25.csv')
allInOne('CSVs/trees-2021-08-08_13-33-06-n25.csv')
# ### Reverse=False
# +
allInOne('CSVs/trees-2021-08-08_14-32-46-n25.csv')
# -
allInOne('CSVs/trees-2021-08-08_14-44-09-n197.csv')
allInOne('CSVs/trees-2021-08-08_14-45-37-n113.csv')
# +
allInOne('CSVs/trees-2021-08-08_14-47-03-n149.csv')
# +
allInOne('CSVs/trees-2021-08-08_14-48-35-n110.csv')
# -
allInOne('CSVs/trees-2021-08-08_14-50-24-n93.csv')
#adversarial failures 80%
allInOne('CSVs/trees-2021-08-08_14-52-48-n113.csv')
# +
#adversarial failures 95%
allInOne('CSVs/trees-2021-08-08_14-57-45-n113.csv')
# -
|
evaluation/notebooks/hops/p2-allInOne-topologyZoo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Table of Content <a id='toc'></a>
#
#
# [1. correlation](#0)
#
# [1.1. Pearson's (linear) correlation](#1)
#
# [1.2. Spearman's (rank) correlation coefficient](#2)
#
# [1.3. Significance of Pearson and Spearman correlation coefficient.](#3)
#
# [1.4 Kendall tau correlation coefficient (for fun)](#4)
#
# [Exercise 01](#5)
#
# [1.5 Correlation and causation](#6)
#
# [2.Linear regression](#7)
#
# [2.1.Presentation](#8)
#
# [2.2.Underlying hypothesis](#9)
#
# [2.3. Goodness of fit](#10)
#
# [2.4. Confidence interval and test statistics](#11)
#
# [2.5. Maximum Likelihood](#12)
#
# [2.6. Model choosing](#13)
#
# [2.7. What to do when some hypothesis about OLS are not true](#14)
#
# [Exercise 02](#15)
# +
import matplotlib.pyplot as plt
from IPython.display import Image
import seaborn as sns
import scipy.stats as stats
import pandas as pd
import numpy as np
from matplotlib import collections as mc
from operator import itemgetter
from mpl_toolkits.mplot3d import Axes3D
# -
#
# So far we have seen how to evaluate the relationship between
# * 2 categorical variables (fisher's exact test, chi-square)
# * 1 quantitative and a categorical variable (t-test, anova)
#
# Now we are going to see how to relate 2 quantitative variables together.
#
# In this notebook we will use the folloming dataset:
df=pd.read_csv('data/etubiol.csv')
df.rename(columns={i:i.replace('.','_') for i in df.columns},inplace=True)
df.eye_colour = df.eye_colour.astype(str) # making eye colour a non-numerical variable
df.diet = df.diet.astype(str) # making diet a non-numerical variable
df
# From which we would like to model the height of individuals given the following informations :
for s in set(df.columns)-set(['height']):
print(s)
#
# [back to the toc](#toc)
#
# <br>
#
# # 1. correlation <a id='0'></a>
#
# Correlation is a measure of the amount of relatedness between two measured variables.
#
# A correlation measure typically goes from -1 (anti-correlation) to 1 (correlation), where 0 is the absence of correlation (independence).
#
# When two variables show a very large correlation, one can be said to be a **predictor** of the other, in the sense that knowing the value of one of the two variable allows us to make a reasonnable guess about the value of the second variable.
# Another way of looking at this relationship is to see those variable as **redundant**, in the sense that they carry the same information : knowing the value of both variable does not bring much more insight compared to knowing the value of only one.
#
#
#
# [back to the toc](#toc)
#
# <br>
#
# ## 1.1. Pearson's (linear) correlation <a id='1'></a>
#
#
# Given 1 sample where each individual $i$ has 2 measures $x_i$ and $y_i$, Pearson's correlation coefficient between $x$ and $y$ is :
#
# $$r_{x,y} = \frac{\sum(x_i - \bar{x})(y_i - \bar{y})}{\sqrt{\sum(x_i - \bar{x})^2}\sqrt{\sum(y_i - \bar{y})^2}}$$
#
# A way to look at this formula is that $r_{x,y}$ tends to move away from zeros when points for which $x_i$ is very different from its mean corresponds to points for which $y_i$ is also very different from its mean. Thus, we are looking for an association in the variation of the variables, which is why Pearson correlation coefficient is also defined as a **standardized covariance** of the two variables.
#
# Pearson's correlation coefficient measures the **linear correlation** between variables, which means that its value is only relevant for the evaluation of a linear relationship. In other words, **two variables can have a strong relationship (i.e. be correlated) but display a Pearson's coefficient of correlation equal to 0**.
#
# 
# > Image by DenisBoigelot, released into the public domain (CC0 license)
#
# This emphasises the danger of relying on a single number for representing sometimes complex notions and the
# importance of always representing visually the variables you want to describe.
#
# Another (fun) example is the [datasaurus dozen](https://www.autodeskresearch.com/publications/samestats) :
#
# 
#
# [back to the toc](#toc)
#
# <br>
#
# ## 1.2. Spearman's (rank) correlation coefficient <a id='2'></a>
#
# Spearman's correlation coefficient corresponds to Pearson's correlation coefficient, but on the **ranks** of observations rather than their values.
#
# Spearman correlation coefficient is used to describe the correlation between two variables when their relation is *monotonic* (i.e. it goes in a single direction: if it is increasing it is always increasing, it never goes down) but non linear (e.g. an exponential relationship)
#
# The formula to calculate the Spearman's rank correlation coefficients between two random variables **X** and **Y** associated to n individual drawns is:
#
# $$\rho=1-6\frac{\sum d_{i}^{2}}{n^{3}-n}$$
#
# Where i is the individual number, n the number of individuals and $d_i$ defined as follow :
#
# $d_i=rank(x_i)-rank(y_i)$
#
# Where of course $x_i$ and $y_i$ are the realization of **X** and **Y** for individual i.
# It is easy to calculate pearson and spearman coeffcient using the scipy.stats library (in the following code abbreviating by stats) followed by either `pearsonr` or `spearmanr`. Both take two lists or arrays as a input and return an array comprised of the coefficient and the p-values.
# +
sigma=1./5
linear=[[u,(u)/100+sigma*np.random.randn()] for u in range(10,500)]
monotonic=[[u,50*(0.8**(u/10))+sigma*np.random.randn()] for u in range(10,500)]
non_monotonic=[[u,(u)**3+3*u**2+sigma*np.random.randn()] for u in np.arange(-1,1,1./250)]
together=[linear,monotonic,non_monotonic]
plt.subplots(133,figsize=(15,5))
for i in range(3):
plt.subplot(1,3,i+1)
x=[u[0] for u in together[i]]
y=[u[1] for u in together[i]]
plt.scatter(x,y)
plt.title('Pearson: {0:.3f}, Spearman: {1:.3f}'.format(
stats.pearsonr(x,y)[0],##just like that
stats.spearmanr(x,y)[0]))
plt.tight_layout()
plt.show()
# -
# Going back to our real dataset
x=df['shoe_size']
y=df['height']
plt.scatter(x,y)
plt.title('Pearson: {0:.3f}, Spearman: {1:.3f}'.format(
stats.pearsonr(x,y)[0],##just like that
stats.spearmanr(x,y)[0]))
plt.xlabel('Shoe size')
plt.ylabel('Height')
plt.tight_layout()
plt.show()
#
# [back to the toc](#toc)
#
# <br>
#
# ## 1.3. Significance of Pearson and Spearman correlation coefficient. <a id='3'></a>
#
# There are 3 main ways to evaluate the significance of $\rho$ (the coefficient of correlation) compared to zero. The most straighforward rely on a t-test to evaluate if $\rho$ is significantly different from 0.
#
# Following a permutation argument (so looking at a null model where you break data correlation/class by creating all the possible arrangements of your data), you can rationalize the usage of the following test statistic :
#
# $$t=\rho\sqrt{\frac{n-2}{1-\rho^2}}$$
#
# which follow a Student's t-distribution under the null hypothesis that $\rho=0$.
#
#
# The other two ways to evaluate the significance of $\rho$ are :
#
# - Do the permutation test yourself
# - Transform the data using Fisher transformation ($F(\rho)=arctanh(\rho)$) to calculate a z variable (instead of a t-student variable), which is normally distributed under $H_0$ : $z=\sqrt{\frac{n-3}{1.06}}F(\rho)$
#
# [back to the toc](#toc)
#
# <br>
#
# ## 1.4 Kendall tau correlation coefficient (for fun) <a id='4'></a>
#
# You have already seen that you could define a correlation coefficient in multiple manners using multiple metrics : multiplicative distances to the means for Pearson, ranking distances between the two random variables for Spearman. For completness let's rapidly present another way to measure correlation : Kendall tau.
#
# Kendal tau is based on the concept of concordant or discordant pairs. A concordant pair is a pair of individual i and j, $i<j$, for which the order relation between the two random variables stands : either $x_i>x_j$ and $y_i>y_j$ or $x_i<x_j$ and $y_i<y_j$. Discordant pairs have opposite signs between x and y.
#
# The coefficient is defined as follow:
#
# $$\tau=\frac{\text{number of concordant pairs}-\text{number of discordant pairs}}{\frac{n(n-1)}{2}}$$
#
# Obviously this coefficient is comprised between -1 and 1, 0 meaning no correlation (indeed there is a total of $\frac{n(n-1)}{2}$ unique pairs).
#
# A rank version also exists (which is mathematically equivalent to the expression above):
#
# $$\tau_{rank}=\frac{2}{n(n-1)}\sum_{i<j} sgn(x_i-x_j)sgn(y_i-y_j)$$
#
# Both the Spearman and and Pearson correlation coefficent are relying on distances, which means they are sensitive to the intensity of the error term and to outliers. Kendall tau is way less influenced by the scale of the variability since it relies only on an order relation.
#
#
#
#
#
#
plt.subplots(133,figsize=(15,5))
for i in range(3):
plt.subplot(1,3,i+1)
x=[u[0] for u in together[i]]
y=[u[1] for u in together[i]]
plt.scatter(x,y)
plt.title('Kendall_tau: {0:.3f}, Kendall_tau_rank: {1:.3f}'.format(
stats.kendalltau(x,y)[0],
stats.mstats.kendalltau(x,y)[0]))
plt.tight_layout()
plt.show()
#
# [back to the toc](#toc)
#
# <br>
#
# # Exercise 01 <a id='5'></a>
#
# Given the *etubiol* dataset, rank the best correlators of *height*.
#
# Among these, which seem redundant to you (because of a high correlation)?
#
# > If you work with a panda dataframe you can use (or not) the functionality `DataFrame.corr()` to calculate the pearson correlation between all the variable of the dataframe
# +
# # %load solutions/solution_04_corr.py
# +
# # %load solutions/solution_04_more.py
# -
#
# [back to the toc](#toc)
#
# <br>
#
# ## 1.5 Correlation and causation <a id='6'></a>
#
# "Correlation does not equal causation" is one of these [often](https://xkcd.com/552/) [repeated](https://www.explainxkcd.com/wiki/index.php/925:_Cell_Phones) [sentence](https://en.wikipedia.org/wiki/Correlation_does_not_imply_causation), but it is still true nonetheless.
#
# Observing that A and B are correlated (linearly or otherwise) gives you **no information** about whether A causes B or B causes A. One may not cause the other at all (they might both be caused by another [unidentified process](https://xkcd.com/1138/)!).
#
# Furthermore, even if a coefficient of correlation if very high, it might still be completely [spurious](https://www.tylervigen.com/spurious-correlations).
#
# 
# It is thus important to always shed a critical eye on correlations and the conclusion we could be tempted to draw from them.
#
# [back to the toc](#toc)
#
# <br>
#
# # 2.Linear regression <a id='7'></a>
#
# Now that we have defined correlation, we can see further ways to characterise the relationship between two variables.
#
# The problem we are interested in is the following:
#
# We measured a bunch of variables per individual, for many individuals. We are interested in the relationship between one of this variable that we will call the *response variable* ($Y$) and the other variables that we will call *covariables* ($X$).
# Of course our measurments are not perfect so there is some noise associated to it ($\epsilon$). In mathematical term we are interested in a class of problem that we can write as :
#
# $\pmb{Y}=f(\pmb{X})+\epsilon$
#
# The function $f$ is called the regression function, and today we will be interested in looking at a particular form of those function: **linear combination**.
#
# A particular case of linear combination would be a single covariable with an intercept like :
#
# $y_i=\beta x_i+c$
#
# 
# A more general case would have more covariables and would be written like:
#
# $$f(\textbf{X}_i,\pmb{\beta})=\sum_{p} \beta_p x_{i,p}= \textbf{X}_{i}^{T}\pmb{\beta}$$
#
# Where *$X_i$* is a vector of p covariables associated to point individual i.
#
# Note that for now nothing is said about the nature of the $x_{i,p}$, for example some could be constant instead of being a variable and thus you could go back to a more specific affine function (like $\beta x+c$).
#
# So of course now the game become to best choose the vector of parameters $\pmb{\beta}$. For that there are two main methods (sorry Bayesian people...):
# - Least Square fit
# - Maximum Likelihood
#
# We will discuss both those methods. Least square fit is the most intuitive and easy to get a hold on, so hopefully you will leave this class with a rather good understanding of it. Maximum likelihood is a bit more advanced in terms of the concepts it utilizes, but being introduce to it will allow you to manipulate cool concepts that you will need by the end of this notebook and if you keep learning about statistics in general.
#
# Underlying those different methods, there are different models:
#
# - Linear models
# - Generalized linear models
#
# The way we wrote the function linking $Y$ to $X$ above, have the noise term $\epsilon$ outside of the function. So one would say that this function only try to represent the mean of the response variable $Y$ along the curve, and as importantly, it does it looking at linear function.
# . This is what we actually do in the framework of Linear models : we only aim to fit the mean response using linear funcitons.
#
# Generalized linear model, in another hand, are more flexible : they allow us to transform the mean response and to fit that transformed response with a linear model. It is very powerfull, as now we could better modeled response variable with broader properties (count data, catgorical data etc....), but significantly more complicated and so we will not talk about those methods here.
#
# # Least square
#
# [back to the toc](#toc)
#
# <br>
#
# ## 2.1.Presentation <a id='8'></a>
#
# For clarity let's define once for all some variables : we have a sample of size n, for each individual on this sample there are p+1 measurments, p covariables and one response variable.
#
# In the least square method we are interested in making the smallest overall distance error between our model and the response variable.
# Typically we want to find the $\beta$ that minimizes:
#
# $S(\pmb\beta)=\sum_i (y_i-f(\textbf{X},\pmb{\beta}))^2=\sum_i \epsilon_i^2$
#
# in mathematical terms you are looking for
#
# $\hat{\pmb\beta}=\text{arg min}_{\pmb\beta}S(\pmb\beta)$
#
# Here the sum is over i, which counts the number of individuals.
#
# > The hat $\hat{.}$, is a notation we use to denote our estimate of the true value of something. So in that sense $\hat{\pmb\beta}$ is the estimate of the "real" coefficient values, and $\hat{Y}$ is the estimation of $Y$ given by our model (also called the model predictions).
#
# Let's try to represent this:
# First a real case scenario just to show case the outcome of what we are trying to do ultimately
# Basically we want to end up with thefollowing graph and outputs.
# +
x=df['shoe_size']
y=df['height']
plt.scatter(x,y,label='data')
slope , intercept , r , pval , stderr = stats.linregress(x,y)#linear regression explaining disease progression thanks
#to bmi
print(r"slope also called beta in our notation= ",slope)
print("intercept also called c in our notation= ",intercept)
yPredict = x * slope + intercept #now that we have the outcome iof the regression which is in this case a slope and
#an intercept we can calulate what the model will predict as a diseas progression given a bmi
plt.plot( x , yPredict , color = 'red',label='model')#the outcome of the regression is this red line
plt.legend(loc='best')
plt.xlabel('shoe_size')
plt.ylabel('height')
# -
# What does Least square method do to end up with this line? Well let's see it on some simple mock data
# +
from matplotlib import collections as mc
## let's create some data to plot
slopeReal = 3
noise = 3
x = np.arange(10)
y = slopeReal * x + noise * np.random.randn(len(x)) # y = beta * x + some noise (no intercept here)
## alternatively we could have:
# x= df['shoe_size']
# y= df['height']
## although note that in that case we also need an intercept. you can try 70
# The challenge of least square regression is to find the slope that minimizes the squared error
# let's try two possible values for the slope
estimatedSlopes = [1,2.5]
fig, ax = plt.subplots(ncols=len(estimatedSlopes) , figsize = (14,7))
for i,slopeEstimate in enumerate(estimatedSlopes):
yPredicted = slopeEstimate * x # prediction of y given the estimated slope and values of x
# error of the prediction
predictionSquaredError = sum( ( yPredicted - y )**2 )
ax[i].plot(x,y, 'o')
ax[i].plot(x,yPredicted, color='orange' , linewidth=2)
# now, let's represent the fitting error as segments between real and estimated values
Real = [i for i in zip(x,y)]
Predicted = [i for i in zip(x,yPredicted)]
lc = mc.LineCollection(zip(Real,Predicted) , colors='black')
ax[i].add_collection(lc)
ax[i].set_title('slope : {} - squared error : {:.2f}'.format(slopeEstimate,predictionSquaredError) )
# +
# to find the best value for the slope, we could try a lot of them :
possibleSlopes = np.linspace(0,6,101)
print('all the slopes tested',possibleSlopes)
errors = []
for sl in possibleSlopes: # we compute the sum of squared error for each slopes
yPred = sl*x
errors.append( sum( yPred - y )**2 )
plt.plot(possibleSlopes , errors )
plt.xlabel('estimated slope')
plt.ylabel('sum of squared errors')
print( 'slope estimate with the smallest error : ', possibleSlopes[np.argmin(errors)] )
# -
#
# While we could use various optimization algorithms to find the best value for $\beta$,
# when the system is overdetermined (*i.e.*, you have more points than coefficients $\beta_i$) an analytical solution exists. It is of the form:
#
# $$\hat{\pmb\beta}=(\pmb X^T \pmb X)^{-1}\pmb X^T \pmb Y$$
#
#
#
#
#
#
# [back to the toc](#toc)
#
# <br>
#
# ## 2.2.Underlying hypothesis <a id='9'></a>
#
# There are a couple of important hypothesis behind this method:
#
# - **Correct specification** : have a good incentive for the function you use
# - **Strict exogeneity** : the errors are centered around the true value of y
# - **No linear dependance** : you can not reconstruct one of your covariable by summing a subset of your covariables with some set of constant weights
# - **Spherical errors**:
# - Homoscedasticity : the spread of the error is the same along the curve (for example not true for counts data).
# - No autocorrelation : error are not correlated along the curve.
#
# The linear dependance part has to do with the part of the exercise where I clustered highly correlated covariables together. If you want to produce a good model for prediction then be carefull about that point. You can have a feeling of what the problem is by imagining that 2 covariables are actually copy pasta of each other : there is not unique way to associate a weight to them... Also then you have 1 variable which bring nothing new to the modeling... so kind of worthless. This is why later on I will ask you to work on a restricted part of the covariables.
#
# If your goal is not really to produce a predictive model but more to infer the size effect of some covariables on your target variable, then it is not too crucial. Just remember that if this is what you want there are other steps to take, that are far beyond the scope of this course, and which are related to the field of causal inference
#
# Normality is not strictly needed for Least Square fitting, neither for the variables nor for their errors.
# However you may need that hypothesis downstream in your analysis, for instance when using a test statistic.
#
# If you errors are normally distributed, then Least Square fitting and Maximum Likelihood are equivalent, showing that your method for choosing $\pmb\beta$ is efficient and sound.
#
# We will quickly present the Maximum Likelihood equivalent as it is both a very useful technic and helps broadening linear models to Generalized Linear Models.
#
# Finally, within that set of constraints and even if the method is called Linear Models, it is possible to fit polynomials of a degree bigger than 1. To do so you just have to precompute the monomials and add them to your set of covariables.
#
# For example :
#
# $y=\beta x +c$ is a linear combination of x
#
# $y=\beta_{1}x+\beta_{2}x^{2}+\beta_{3}x^{3}$ is still a linear combination of features (covariables) x, $x^{2}$ and $x^{3}$, and **X** becomes {$x,x^2,x^3$\}
#
#
# [back to the toc](#toc)
#
# <br>
#
# ## 2.3. Goodness of fit <a id='10'></a>
#
# To have an idea of how good your fit is, you can either directly use the Mean Squared Error (MSE) or the adjusted coefficient of determination $\pmb R^2_a$.
#
# The MSE is defined as follow:
#
# <br>
#
# $$MSE=\frac{\sum (y_i-\hat{y_i})^2}{n-2}$$
# and accounts for what your model is missing.
# That could be the simple inherent variance induced by the noise term or the noise term and a missing term that your model doesn't take into account. By its nature, this metric makes it hard to compare between different hypothetical fitting models or different dataset.
#
# A better normalized metric is the **adjusted coefficient of determination $\pmb R^2_a$**.
# The adjusted part is very necessary when we work in the context of multiple linear regression (more than one covariable).
#
# Let's start by defining the coefficient of determination $\pmb R^2$.
# This coefficient partitions the variance present in your data between what is taken into account by your model and what is not.
#
# $$R^2=1-\frac{SSE}{SST}$$, where SSE is the sum of squared errors ($\sum_i (y_i-\hat{y_i})^2$) and SST in the sum of squares total ($\sum_i (y_i-\bar{y})^2$)
#
# For the adjusted coefficient of determination you have to take into account that SSE and SST don't have the same degree of freedom and you should adjust for that.
#
# $$R^2_a=1-\frac{n-1}{n-p}(1-R^2)$$, with $p$ the number of covariables and $n$ the number of individuals.
#
# > Note : you can see that when there is only one covariable then $R^2_a = R^2$
#
#
#
# ### Examples: some linear regression examples
# #### Affine
# This is typically the case where we would like to describe height = $\beta$ shoe_size + c
#
# Here we look at a model y=1+3*x
# +
import statsmodels
import statsmodels.api as sm
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import PolynomialFeatures
X=np.array(np.arange(-1,1,10**-2))
fig,ax=plt.subplots(2,3,figsize=(15,10))
for k,epsilon in enumerate([0,1,10]):
y = 1+3*X + epsilon* np.random.randn( len(X) )
## creating a dataframe with the data
X1 = sm.add_constant(X)##adding the intercept
df_=pd.DataFrame(X1,columns=['c','x'])
model = sm.OLS( y , df_[['c','x']])##defining an Ordinary Least Square variable
results = model.fit()##fitting it
y_predict=results.predict(X1)# predict back what your target variable would be in that model
R2=r2_score(y,y_predict)#evaluate R2
MSE=mean_squared_error(y,y_predict)#evaluate MSE
## plotting the data and model
ax[0,k].plot(X,y,'ko',label='Data',linewidth=10,alpha=0.5)
ax[0,k].plot(X,y_predict,'r-.',label='Predicted')
ax[0,k].legend(loc='best',fontsize=10)
ax[0,k].set_title('R2={0:.2f}, MSE={1:.2f}, noise={2}'.format(R2,MSE,epsilon))
ax[0,k].set_xlabel('X')
ax[0,k].set_ylabel('y')
## plotting predicted value versus real value is a good way to visualize a fit
ax[1,k].plot(y,y_predict,'ko')
ax[1,k].set_xlabel('true y')
ax[1,k].set_ylabel('predicted y')
print('epsilon',epsilon)
print('fit param c= {0:.3f} beta= {1:.3f}'.format(results.params['c'],results.params['x']))
print('true param c= {0:.3f} beta= {1:.3f}'.format(1,3))
print()
plt.tight_layout()
plt.show()
# -
# Let's see what happened if we miss specify the polynomial degree :
# +
X=np.array(np.arange(-1,1,10**-2))
fig,ax=plt.subplots(2,3,figsize=(15,10))
for k,epsilon in enumerate([0,1,10]):
y = 1+3*X + epsilon* np.random.randn( len(X) )
X1 = np.column_stack((X, X**2,X**3))
X1 = sm.add_constant(X1)##adding the intercept
df_=pd.DataFrame(X1,columns=['c','x','x²','x³'])
model = sm.OLS( y , df_[['c','x','x²','x³']])##defining an Ordinary Least Square variable
results = model.fit()##fitting it
y_predict=results.predict(X1)# predict back what your target variable would be in that model
R2=r2_score(y,y_predict)#evaluate R2
MSE=mean_squared_error(y,y_predict)#evaluate MSE
ax[0,k].plot(X,y,'ko',label='Data',linewidth=10,alpha=0.5)
ax[0,k].plot(X,y_predict,'r-.',label='Predicted')
ax[0,k].legend(loc='best',fontsize=10)
ax[0,k].set_title('R2={0:.2f}, MSE={1:.2f}, noise={2}'.format(R2,MSE,epsilon))
ax[0,k].set_xlabel('X')
ax[0,k].set_ylabel('y')
ax[1,k].plot(y,y_predict,'ko')
ax[1,k].set_xlabel('true y')
ax[1,k].set_ylabel('predicted y')
print('epsilon',epsilon)
print('fit param c= {0:.3f} beta1= {1:.3f} beta2= {2:.3f} beta3= {3:.3f}'.format(results.params['c'],results.params['x'],results.params['x²'],results.params['x³']))
print('true param c= {0:.3f} beta= {1:.3f}'.format(1,3))
print()
plt.tight_layout()
plt.show()
# -
# On the height datset
# +
import statsmodels.formula.api as smf
model = smf.ols(formula='height ~ shoe_size', data=df)
results = model.fit()#we do the actual fit
y_predict=results.predict(df)# predict back what your target variable would be in that model
R2=r2_score(df['height'],y_predict)
MSE=mean_squared_error(df['height'],y_predict)
plt.plot(df['shoe_size'],df['height'],'ko',label='Data',linewidth=10,alpha=0.5)
plt.plot(df['shoe_size'],y_predict,'r-.',label='Predicted')
plt.legend(loc='best',fontsize=10)
plt.title('R2={0:.2f}, MSE={1:.2f}'.format(R2,MSE))
plt.xlabel('shoe_size')
plt.ylabel('height')
plt.show()
plt.plot(df['height'],y_predict,'ko')
plt.xlabel('true height')
plt.ylabel('predicted height')
plt.show()
print('fit param for shoe_size ',results.params['Intercept'])
print('fit intercept ', results.params['shoe_size'])
print()
# -
# ##### Unidimensional, multiple covariables
# This is typically the case where we would like to describe $\text{height}$ = $\beta_1$ $shoesize$ +$\beta_2$ $shoesize^2$ +$\beta_3$ $shoesize^3$+c
#
#
# Here we look at a model y=1-3*x+6*x^3
# +
X=np.array(np.arange(-1,1,10**-2))
fig,ax=plt.subplots(2,3,figsize=(15,10))
for k,epsilon in enumerate([0,1,10]):
y = 1-3*X+6*X**3 +epsilon*np.random.randn(len(X))
X1 = np.column_stack((X, X**2,X**3))
X1 = sm.add_constant(X1)##adding the intercept
df_=pd.DataFrame(X1,columns=['c','x','x²','x³'])
model = sm.OLS( y , df_[['c','x','x²','x³']])##defining an Ordinary Least Square variable
results = model.fit()##fitting it
y_predict=results.predict(X1)# predict back what your target variable would be in that model
R2=r2_score(y,y_predict)#evaluate R2
MSE=mean_squared_error(y,y_predict)#evaluate MSE
ax[0,k].plot(X,y,'ko',label='Data',linewidth=10,alpha=0.5)
ax[0,k].plot(X,y_predict,'r-.',label='Predicted')
ax[0,k].legend(loc='best',fontsize=10)
ax[0,k].set_title('R2={0:.2f}, MSE={1:.2f}, noise={2}'.format(R2,MSE,epsilon))
ax[0,k].set_xlabel('X')
ax[0,k].set_ylabel('y')
ax[1,k].plot(y,y_predict,'ko')
ax[1,k].set_xlabel('true y')
ax[1,k].set_ylabel('predicted y')
print('epsilon',epsilon)
print('fit param c= {0:.3f} beta1= {1:.3f} beta2= {2:.3f} beta3= {3:.3f}'.format(results.params['c'],results.params['x'],results.params['x²'],results.params['x³']))
print('true param c= {0:.3f} beta1= {1:.3f} beta2= {2:.3f} beta3= {3:.3f}'.format(1,-3,0,6))
print()
plt.tight_layout()
plt.show()
# -
# On the real data, eventhough you probably have no reason to model it with something else than a degree 1 polynomial
# +
df_extended=df.copy()
df_extended['shoe_size2']=df['shoe_size']**2
df_extended['shoe_size3']=df['shoe_size']**3
model = smf.ols(formula='height ~ shoe_size + shoe_size2 + shoe_size3', data=df_extended)
results = model.fit()#we do the actual fit
y_predict=results.predict(df_extended)# predict back what your target variable would be in that model
R2=r2_score(df_extended['height'],y_predict)
MSE=mean_squared_error(df_extended['height'],y_predict)
plt.plot(df_extended['shoe_size'] , df_extended['height'] ,'ko',label='Data',linewidth=10,alpha=0.5)
plt.plot(df_extended['shoe_size'] , y_predict ,'ro',label='Predicted')
plt.legend(loc='best',fontsize=10)
plt.title('R2={0:.2f}, MSE={1:.2f}'.format(R2,MSE))
plt.xlabel('shoe_size')
plt.ylabel('height')
plt.show()
plt.plot( df_extended['height'], y_predict,'ko')
plt.xlabel('true height')
plt.ylabel('predicted height')
plt.show()
print('fit param for shoe_size ',results.params['Intercept'])
print('fit intercept ', results.params['shoe_size'])
print()
# -
# #### Multidimensional
# Let see what that look like with our dataset
# +
model = smf.ols(formula='height ~ shoe_size+height_M', data=df)
results = model.fit()#we do the actual fit
y_predict=results.predict(df)# predict back what your target variable would be in that model
R2=r2_score(df['height'],y_predict)
MSE=mean_squared_error(df['height'],y_predict)
fig = plt.figure(figsize=(14,10))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(df['shoe_size'], df['height_M'], df['height'], s=20, c='k',label='Data', depthshade=True)
ax.scatter(df['shoe_size'], df['height_M'],y_predict,label='Predicted' ,color='m')
plt.legend(loc='best',fontsize=10)
plt.title('R2={0:.2f}, MSE={1:.2f}'.format(R2,MSE))
plt.xlabel('shoe_size')
plt.ylabel('height_M')
ax.set_zlabel('height')
plt.show()
plt.plot(df['height'],y_predict,'ko')
plt.xlabel('true height')
plt.ylabel('predicted height')
plt.show()
print('fit param for shoe_size ',results.params['shoe_size'])
print('fit param for height_M ',results.params['height_M'])
print('fit intercept ',results.params['Intercept'])
print()
# -
# Stats model gives you way more info than just a predicition and your fitted parameters. But to really use those info that we will see later on, we need to introduce some other stuff first.
#
# [back to the toc](#toc)
#
# <br>
#
# ## 2.4. Confidence interval and test statistics <a id='11'></a>
#
# After your fitting, you would probably like to know the confidence interval for each of your estimated $\beta$, as well as if they are truly necessary (significantly different from zero).
# For both **you can't truly do anything without making an hypothesis about the statistic of the noise** : here comes the part where assuming your noise to be normally distributed ($N(0,\sigma^2)$) becomes important, but potentially wrong too.
#
# For the confidence interval, if you have an infinite amount of data, and your noise distribution is not heavytailed, you can show that the estimators are well described by a normal statistic (there is convergence in the distribution so that $(\hat{\pmb\beta}-\pmb\beta)\rightarrow N(0,\sigma^2 (\pmb X^T \pmb X)^{-1})$).
# So for big amount of points relative to the number of estimated parameters, you are not making a big mistake by writting:
#
# $$\beta_p \in [\hat{\beta_p} \pm z_{1-\frac{\alpha}{2}}\sqrt{\hat{\sigma}^2 [(\pmb X^T \pmb X)^{-1}]_{p,p}}]$$
#
# If you don't have a huge amount of data you need to show that you have an incentive about your noise statistic to use these kind of confidence intervals (some libraries that we are going to use can do that for you!).
#
#
# For the significance of the coefficients, **if you know that your noise is normally distributed then you can use a t-test**.
#
#
# [back to the toc](#toc)
#
# <br>
#
# ## 2.5. Maximum Likelihood <a id='12'></a>
#
# Maximum Likelihood is a method that is used to estimate parameters of a probablililty distribution, and is usefull for model choosing. It is done by maximizing the likelihood function. In the case that we are interested in (i.e. independant identically distributed) this likelihood function is simply the product of a density function values over the entire sample. It is a parametric method since it needs to have an a priory about the density function for it to work. Since it is a product, most of the time we would rather work with the log likelihood function which transforms this product into a sum.
#
# So we would like to maximize $l$, the loglikelihood function, by choosing a set of parameters $\Theta$.
# Where $l$ is of the form:
#
# $l(\Theta;Y)=\sum_i ln(p(y_i|\Theta))$
#
# Where $Y$ is a random variable and $p()$ is the density function associated to $Y$.So you want to find the following estimation for $\pmb\Theta$
#
# $$\hat{\pmb\Theta}=\text{arg max}_{\pmb\Theta}l(\pmb\Theta;Y)$$
#
#
#
#
# ### What are we looking at?
#
# Let's take the example of a gaussian where you would like to estimate the $\sigma$ and the $\mu$, given your data. As they are simulated data we chose that $\mu=2$ and $\sigma=0.5$
def gaussian_dist(x,mu,sigma):
""" returns the probability of observing x in a normal distribution of mean mu and standard deviation sigma """
return 1./(sigma*np.sqrt(2*np.pi))*np.exp(-1./(2*sigma**2)*(x-mu)**2)
# note : this is equivalent to stats.norm.pdf( x , mu , sigma )
# +
X_small=np.random.randn(10)*0.5+2 # this is our observed data, with ( mean=2 , sd=0.5 )
m=[2,0.5] # we will try 2 possible combinations of paramters ( mean=2 , sd=0.5 ) and ( mean=0.5 , sd=0.5 )
s=[0.5,0.5]
fig, ax = plt.subplots(ncols=len(m) , figsize = (14,7))
X_small_=[[v,0] for v in X_small]
x=np.arange(-2,4,0.005) # we will plot between -2 and 4
print('the data that we observed',[v[0] for v in X_small_])
for q in range(len(m)): # for each of the parameter combinations we want to try
ax[q].plot(X_small,[0]*len(X_small),'k+') # we plot the observed data as crosses
ax[q].plot( x , stats.norm.pdf( x , loc = m[q] , scale = s[q] ),'k') # we plot the distribution we are testing
Predicted = stats.norm.pdf( X_small , loc = m[q] , scale = s[q] )
Predicted_= [i for i in zip(X_small,Predicted)] # this is to plot segments
lc = mc.LineCollection(zip(X_small_,Predicted_) , colors='red',linewidths=5,alpha=0.7,label='Predicted likelihood')
ax[q].add_collection(lc)
ax[q].legend(loc='best',fontsize=10)
# the log likelihood of this set of parameters is the sum of the log of the probability densities of the sample
sum_like=sum(np.log(Predicted))
ax[q].set_title('$\mu$ : {} - $\sigma$: {:.2f} - log likelihood : {:.2f}'.format(m[q],s[q],sum_like) ,fontsize=13)
ax[q].set_xlabel('X')
ax[q].set_ylabel('Likelihood')
plt.tight_layout()
# -
# Multiplying those red bars is exactly what the maximum likelihood does.
#
# Basically, you shift your theoritical distribution to the right or the left (trying different means), and you narrow it or widen it (trying different variances).
#
# For each of those try you multiply those red bars together, and the combination of parameters giving highest result is the one maximizing the likelihood of your data being produced by that distribution with those parameters.
#
#
# It is important to point out here that **even when our data are actually coming from a certain distribution, there will (almost) always be a difference between the theoretical distribution and the recovered one**, as to have perfect match you would need an infinite number of data points.
X=np.random.randn(800)*0.5+2
fig = plt.figure(figsize = (10,7))
sns.kdeplot(X,label='data probability\ndensity function')
x=np.arange(0,4,0.005)
plt.plot(X,[0]*len(X) ,'k+',label='data')
plt.plot(x, stats.norm.pdf( x , loc = 2 , scale = 0.5 ) ,'r',label='generative probability\ndensity function')
plt.ylabel('Likelihood')
plt.xlabel('X')
plt.legend(loc='best',fontsize=10)
# Let's test many combinations of possible means and standard deviations to see where our maximum of likelihood lies.
# +
import math
mu=np.arange(0,4,0.1) # from 0 to 4 by increments of 0.1
sigma=np.arange(0.1,2.1,0.1) # from 0.1 to 2.1 by increments of 0.1
mu,sigma=np.meshgrid(mu,sigma) # this useful function combines all possibles values for mu and sigma
def loglike_func(X,mu,sigma):
"""returns a list of the loglikelihoods of mus and sigmas given data X"""
ll = []
for i in range(len(mu)):
ll.append( sum(np.log(stats.norm.pdf(X,mu[i],sigma[i]))) )
if math.isnan(ll[-1]) or ll[-1] < -10000:
ll[-1] = -10000 # we verify that no numerical error gave us an NaN or very small log value
return ll
# we compute the log-likelihood for all tested parameters values
zs=np.array(
loglike_func(X,np.ravel(mu),np.ravel(sigma))
)
loglike=zs.reshape(mu.shape)
bestMu = np.ravel(mu)[np.argmax(zs)]
bestSigma = np.ravel(sigma)[np.argmax(zs)]
# make a 3D figure of our loglikelihood landscape
from mpl_toolkits.mplot3d import Axes3D
print(r'Highest likelihood is for \mu and \sigma :',bestMu,bestSigma)
fig = plt.figure(figsize=(14,8))
ax = Axes3D(fig)
ax.plot_surface(mu,sigma,loglike,cmap='plasma')
ax.scatter(bestMu,bestSigma,max(zs),s=200,c='r') # put a dot at the ML value
ax.set_xlabel('$\mu$')
ax.set_ylabel('$\sigma$')
ax.set_zlabel('Loglike')
plt.title("Loglikelihood landscape")
plt.show()
# -
# ### What is the link between OLS and maximum likelihood (optional but a good gateway to understand GLM)
#
# Let's now imagine that we try to fit the average of a Y, $\bar{Y}$, along the curve $\bar{Y}=\beta X+c$ for which the noise around those averages is gaussian. Since we didn't put the noise in this equality, thus it really represents a fit of the average of Y. The equation representing the fitting of Y would be $Y=\beta X+c+\epsilon$. We could thus consider that we can switch to the following problem of distribution fitting, defined by the density function:
#
# $$p(y_i|\bar{y_i},\sigma)=\frac{1}{\sqrt{2\pi\sigma^2}}*\exp(-\frac{1}{2}\frac{(y_i-\bar{y_i})^2}{\sigma^2})$$
#
# Note that the parameters you want to estimate are $\bar{y_i}$ and $\sigma$.
#
# By definition of the likelihood function over $n$ individuals in a sample is:
#
# $$\Pi_i \frac{1}{\sqrt{2\pi\sigma^2}}*\exp(-\frac{1}{2}\frac{(y_i-\bar{y_i})^2}{\sigma^2})$$
#
# which transformed into the loglikelihood function:
#
# $$l(\bar{y_i},\sigma;Y) = \sum_i -\frac{1}{2}\frac{(y_i-\bar{y_i})^2}{\sigma^2} + constant$$
#
# Now let's rewrite $\bar{y_i}=\beta x_i+c=f(x_i,\beta)$. So now the game is to find $\beta$ and $c$.
#
# You see now that maximizing $\sum_i -(y_i-f(x_i,\beta))^2$ over $\beta$ is the same than minimizing $\sum_i (y_i-f(x_i,\beta))^2$ over $\beta$(which is what we wrote for Ordinary Least Square)
#
#
#
# [back to the toc](#toc)
#
# <br>
#
# ## 2.6. Model choosing <a id='13'></a>
#
# Most of the time you are not sure of the model you want to fit. You might have a broad idea of the different forms of the function but you don't really know for example what would be the best degree for your poynomial or if all the covariables are actually necessary. Of course you could say "I am keeping the model that fit the best in term of $R^2$". But the question really is : is that bunch of extra parameters that are complexifying my model worth the increase in $R^2$?
#
# We touched that question in 1.4 by asking about the significance of parameters values. Again if you are confident on the noise distribution you are dealing with (let's say it is normally distributed), and you have a function in mind but you don't know if you should include 1,2 or $p$ covariables then the problem is easy: you can use a log-likelihood ratio test.
#
#
# ### Likelihood ratio test (LRT)
#
#
# To perform a likelihood ratio test you just have to calculate the difference between the maximised log-likelihood of the two models you are comparing. You can estimate the significance of that difference either by using a test statistic (approximate method) or by simulation.
#
# LRT are to be used in the case of nested function comparison. Nested functions are functions that have the same form but differ from the number of parameters used : for example comparing $y=\beta_1 x_1 +c$ and $y=\beta_1 x_1 +\beta_2 x_2 +c$. In this course this will always be the case (but just remember that outside of this course you might want to do other comparison, so be carefull).
#
# Quickly :
#
# You want to compare model $M_0$ and $M_1$, respectively having $\{\beta_{1,0}\}$ and $\{\beta_{1,2},\beta_{2,2}\}$ as parameters. You want to see if adding this extra parameter $\beta_{2,2}$ is worth it.
#
# The LRT statistics is :
#
# $2*(l(Y;\hat{\beta}_{1,2},\hat{\beta}_{2,2},X)-l(Y;\hat{\beta}_{1,0},X))$
#
# Where the hat represents the maximum likelihood estimates. The LRT statistic asymptoptically, for your sample size going to infinity, follows a **chi-square distribution with a number of degree of freedom equal to the difference between the number of degrees of freedom in your models**. You have thus access to a P-value which will help you to decide if complexifying your model is worth it.
#
# To calulate this P-value you can use 1-scipy.stats.chi2.cdf(LRT,$df_{M_1}-df_{M_0}$), where $df$ is the number of degree of freedom of the models.
#
#
# ### Regularization (for the culture)
#
# If you don't have access to the noise properties (*i.e.* you have no good reason to say it is normally distributed), you can always use a technic called regularization which is going to penalize covariables that are not really important to your fit. This is more on the machine learning side, and so a lot should be said about how to properly use this technic (splitting your dataset between train, validation and test set, *etc.*).
# But let's just check what the principle behind it is and I will give an additionnal example on it later on.
#
# The only thing that this method does is to add a penalization term to the least square minimization method seen before.
# This penalization is based on the size of the parameters estimated.
# The rational is that some time, parameters estimated will be inflated to compensate the fact that the covariable is not really important to fit the data, but is rather important to understand the noise. So regularization minimizes square error while balancing the overall size of the parameters.
#
# Broadly, it can looks like that:
#
# * $S(\pmb{\beta}) + \frac{1}{C}\Sigma^{n}_{i=1}|\beta_{i}|$ , l1 regularization (Lasso) C being the inverse of the weight that you put on that regularization
#
# * $S(\pmb{\beta}) + \frac{1}{C}\Sigma^{n}_{i=1}\beta_{i}^{2}$ , l2 regularization (Ridge)
#
# * $S(\pmb{\beta}) + \frac{1}{C}\Sigma^{n}_{i=1}(\alpha|\beta_{i}|+(1-\alpha)\beta_{i}^{2})$ , elasticnet
#
# How to choose this C, or sometime $\alpha$, is related to the field of machine learning and has to do with splitting your data set into train, validation and test sets. We will not go deeper than that but statsmodels has it implemented `statsmodels.regression.linear_model.OLS.fit_regularized` and scikitlearn, a python library specialized in machine learning has even more option.
#
# This is really just for culture, there are many more things to learn before applying those technics rigorously.
# ### Examples with Stats model
# On a mock dataset for which we know the ground truth : y=1-3x+6x^3
# +
## Statsmodel scale your variable to unit lenght automatically so no need for scaling here.
import statsmodels
import statsmodels.api as sm
###making toy data
nsample = 200
x = np.linspace(0, 10, nsample)
X = np.column_stack((x, x**3))
beta = np.array([1, -3, 6])
e = 1000*np.random.normal(size=nsample)#for now noise is 1000, but play with it
X = sm.add_constant(X)##adding the intercept
y = np.dot(X, beta) + e## making y=1-3x+6x^3 +noise
y_true=np.dot(X, beta)
df_=pd.DataFrame(X,columns=['c','x','x³'])
df_['y']=y
model = sm.OLS(df_['y'], df_[['c','x','x³']])##defining an Ordinary Least Square variable
results = model.fit()##fitting it
res=results.summary()
print(res)
# -
# The first panel gives you an overview of the fit quality:
# * You recognize the good old $R^2$ and $R_a^2$
# * The F-statistic and its associated P-value test the hypothesis that all the coefficients are 0 (normality assumption)
# * You should also recognize the log-likelihood (normality assumption)
# * AIC and BIC respectively Aikike Information Criterion and Bayesian Information Criterion are equivalent of likelihood but that you can use to compare non nested models.
#
# The second panel is quite self explanatory, just be careful with this t-test which again makes the assumption that errors are normally distributed, same for the standard error and the 95% confidence interval.
#
# The third panel is a summary of a few statistical tests that will give you a sense of how all of the hypothesis needed for OLS are plausible:
# * Omnibus and Prob(omnibus): this is a test for normality of residuals. Low P-values means that your linear model is not adapted
# * Durbin-Watson : tests autocorrelation in the error terms (2 is no autocorrelation, less than 1 is bad)
# * Jarque-Bera: tests if the skewness and kurtosis of your errors are looking like a normal distribution. If the Pvalue is high then they look normal.
# * Condition Number : sensibility to noise of the fit.Skewness and kurtosis of your noise (both 0 for normally distributed noise).
#
# +
###Plotting the fit
#for some noise: scale=1000
from statsmodels.sandbox.regression.predstd import wls_prediction_std
prstd, iv_l, iv_u = wls_prediction_std(results)
fig, ax = plt.subplots(figsize=(8,6))
ax.plot(x, y, 'o', label="data")
ax.plot(x, y_true, 'b-', label="True")
ax.plot(x, results.fittedvalues, 'r--.', label="OLS")
ax.plot(x, iv_u, 'r--')
ax.plot(x, iv_l, 'r--')
ax.legend(loc='best');
# +
#### a little bit of gymnastic to get this summary saved and usable.
results_as_html = res.tables[0].as_html()
result_general_df2=pd.read_html(results_as_html, header=0, index_col=0)[0]
list1=["Dep. Variable:"]+list(result_general_df2.index)+[result_general_df2.columns[1]]+list(result_general_df2[result_general_df2.columns[1]])
list2=[result_general_df2.columns[0]]+list(result_general_df2[result_general_df2.columns[0]])+[result_general_df2.columns[2]]+list(result_general_df2[result_general_df2.columns[2]])
dico_i={s:v for s,v in zip(list1,list2)}
result_general_df=pd.DataFrame([[dico_i[v]] for v in list1],index=list1,columns=['Value']).transpose()
results_as_html = res.tables[1].as_html()
result_fit_df=pd.read_html(results_as_html, header=0, index_col=0)[0]
#print(result_general_df)
#print(result_fit_df)
# +
# I have put the overall recap of the fit in a dataframe format so you can use it later
result_general_df
# -
# I have put the outcome of the fit in a dataframe format so you can use it later
result_fit_df
# #### Stats model on etubiol dataset
#
# let's see how that work on real data: let's say we want to predict height using the height of the mother (`height_M`) and and shoe size.
# +
import statsmodels
import statsmodels.api as sm
import statsmodels.formula.api as smf
model1 = smf.ols(formula='height ~ height_M + shoe_size', data=df)
results_model1 = model1.fit()#we do the actual fit
res=results_model1.summary()#we print the summary
print(res)
# -
# let's check with only shoe_size
# +
model2 = smf.ols(formula='height ~ shoe_size', data=df)
results_model2 = model2.fit()#we do the actual fit
res=results_model2.summary()#we print the summary
print(res)
# -
# let's add one more covariable
print('loglikelihood model shoe_size:', results_model2.llf )
print('loglikelihood model shoe_size + mother height:',results_model1.llf)
#print('loglikelihood model shoe_size + mother height + number of siblings:',result_general_df_3['Log-Likelihood:']['Value'])
# We can already rule out number of siblings as it didn't change the loglikelihood. Adding weight did increase the loglikelihood, is it significant enought for us to keep it for modelling?
LRT=2*(results_model1.llf - results_model2.llf)
print('The LRT statistics is ',LRT)
print('The associated pvalue to that difference of Log likelihood is', 1-stats.chi2.cdf(LRT,2-1))
#
# [back to the toc](#toc)
#
# <br>
#
# ## 2.7. What to do when some hypothesis about OLS are not true <a id='14'></a>
#
# When the homoscedasticity of your data is not true you have a few possibilities:
# - you can transform your data so your data become homoscedastic (for example you could use variance stabilizing transformation, or a simple log transform or other...)
# - you can change your loss function that we previously called $S(\beta)$ to reweight the different members of that equation by taking into account the discrepancy in terms of variance. That only works if there is no correlation between the error terms. In that case the method is called Weighted Least Square and it simply transformed to $S(\pmb\beta)=\sum_i \frac{1}{\sigma_i^2} (y_i-f(\textbf{X},\pmb{\beta}))^2$.
# - if there is a correlation between the different error terms then it becomes more complicated, but technics exist such as Generalized Least Square model
#
# Finally if you know what statistics your measurement follow, you can bypass all of those problems (and encounter others :-)) by using a maximum likelihood estimation rather than an LS method. By doing so you will have to put yourself in the framework of Generalized Linear Models, which is outside of the scope of this course.
#
# [back to the toc](#toc)
#
# <br>
#
# # Exercise 02 <a id='15'></a>
#
# Using the following subset of covariables ['shoe_size','height_M','nb_siblings_F'] find the best model to predict height.
#
# +
# # %load solutions/solution_04_reg.py
# -
# Final remark : to actually have the best model I invite you to follow the practice of machine learning that is based in splitting your dataset, cross validation etc... What you have learn today is still an introduction. You are more ready than ever to do modelisation but be aware that many things still need to be done to have you derive a model following state of the art methodology.
|
Intro_to_stats_in_life_sciences/04_correlation_and_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ローカルにあるデータをフィーチャ レイヤーとして公開する
#
# ArcGIS Developersから作成した無償アカウント、または既にお持ちの組織サイトのアカウントの両方で実践できます。
#
# ### ここでは次の 3 つのファイルを ArcGIS Online へ公開します。
# * CSV
# * シェープ ファイル(*.shp)
# * ファイルジオデータベース(*.gdb)
#
# ### おまけ1
# * PDFファイル(*.pdf)
# * Microsoft Wordファイル(*.docx)
#
# ※これら2つは、ArcGIS Onlineに作成可能なフォルダに格納します。
#
# そのほか ArcGIS API for Python を使ってArcGIS Online へ公開できるファイルは[こちら](https://developers.arcgis.com/rest/users-groups-and-items/items-and-item-types.htm)
#
# ### おまけ2
# * 作成したフィーチャサービスを全体公開にする
#
#
# #### ユーザでログインする
# ArcGIS for Developers ログイン情報
develoersUser = ''
develoersPass = ''
# +
from arcgis.gis import GIS
gis = GIS("http://"+ develoersUser +".maps.arcgis.com/",develoersUser,develoersPass)
user = gis.users.get(develoersUser)
user
# -
# ## CSVファイルを公開する
# ※緯度経度またはXYなど位置情報が含まれることが条件です
# +
# csvファイルの読み込み
import pandas as pd
from IPython.display import display
csvfile = 'data/kanazawaParking.csv'
# データ内容の確認
dataflame = pd.read_csv(csvfile)
display(dataflame)
# -
# CSV ファイルのアップロード
csvItem = gis.content.add({},csvfile)
# FeatureService として公開
csvService = csvItem.publish()
csvService
# ## シェープ ファイルを公開する
#
# シェープ ファイルのアップロード
shpfile = 'data/yokohamaPolygonShp.zip'
ShpItem = gis.content.add({}, shpfile)
# FeatureService として公開
shpService = ShpItem.publish()
shpService
# ## ファイルジオデータベースを公開する
# ファイルジオデータベースのアップロード
# ファイルタイプ(この場合必要)と公開名称(任意)とタグ(任意)を指定します。
fgdb = 'data/kawasakiEscapeFGDB.zip'
fgdbItem = gis.content.add({ 'type': 'File Geodatabase' , 'title':'川崎避難所ファイルジオ' , 'tags':'geodev_jp'}, fgdb )
# FeatureService として公開
fgdbItem.publish()
# ### おまけ1
#
# * PDFファイル(*.pdf)
# * Microsoft Wordファイル(*.docx)
# #### PDF ファイルをアップロードする
#
# * フォルダを作成してみよう
# * PDFファイルをアップロードしてみよう
# 日付フォルダを作成する
from datetime import datetime as dt
tdatetime = dt.now()
today = tdatetime.strftime('%Y/%m/%d')
FolderName = 'geodev_'+ today
# ArcGIS Onlineへ フォルダを作成する
newFolder = gis.content.create_folder(FolderName)
newFolder
pdffile = 'data/ESRIジャパンPDFファイル.pdf'
pdffileItem = gis.content.add({'type': 'PDF','tags':'geodev_jp' }, pdffile)
pdffileItem
# フォルダ移動
pdffileItem.move(newFolder)
# #### Microsoft Word ファイルをアップロードする
#
# * フォルダ指定でWordファイルをアップロードしてみよう
wordfile = 'data/ESRIジャパンWordファイル.docx'
wordfileItem = gis.content.add({'type': 'Microsoft Word','tags':'geodev_jp' }, wordfile , folder=newFolder.get('title'))
wordfileItem
# +
# ArcGIS Onlineへログインして、コンテンツとフォルダを見てみましょう
# -
# ### おまけ2
#
# * 作成したフィーチャサービスを全体公開にする
#
# CSVファイルから作成したフィーチャ サービスを全体公開にします。
csvService.share(everyone=True, org=True, allow_members_to_edit=False)
|
samples/1.Create published Service from my local.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Original Demo: https://github.com/keras-team/keras/blob/master/examples/mnist_mlp.py
from __future__ import print_function
import syft
import syft.nn as nn
import syft.optim as optim
from syft.controller import models, tensors
import imp
imp.reload(syft)
imp.reload(syft.nn)
import numpy as np
from syft import FloatTensor
import syft.interfaces.keras as keras
from syft.interfaces.keras.datasets import mnist
batch_size = 128
num_classes = 10
epochs = 20
# the data, shuffled and split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, 784)
x_test = x_test.reshape(10000, 784)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# +
model = nn.Sequential([
nn.Linear(784,10)
])
for p in model.parameters():
p *= 0
model.summary()
optimizer = optim.SGD(model.parameters(),0.00001)
criterion = syft.nn.CrossEntropyLoss()
input = FloatTensor(x_train,autograd=True)
target = FloatTensor(y_train,autograd=True)
# -
final_loss = model.fit(input=input,
target=target,
batch_size=100,
criterion=criterion,
optim=optimizer,
iters=3,log_interval=1)
final_loss
|
notebooks/demos/MNIST MLP Using Optimizer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Two qubits
# +
import numpy as np
import pandas as pd
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, execute
from qiskit.tools import visualization
from qiskit import BasicAer as Aer
import matplotlib.pyplot as plt
# %matplotlib auto
from IPython.display import Latex
from IPython.display import Math
import ancillary_functions as anf
# -
# ## Preliminary task
# ### Create two-qubit circuit and perform X gate on one of them.
# * Try to use it only on first qubit, later only on second qubit.
# * Test how results are numbered. You may change mapping to classical register when you measure.
# * Check what is the convention for numbering qubits in state vector.
# * This may seem a little silly, but it's actually quite easy to get lost in naming conventions if you won't try it yourself.
# +
# specify variables
qrs =
crs =
circuit_name =
nos =
backend_name =
backend = Aer.get_backend(backend_name)
circuit, qreg, creg = anf.create_circuit_draft(qrs=qrs, crs=crs, circuit_name=circuit_name)
# add things to circuit
# ....
# perform experiments
job = execute(circuit, backend=backend, shots=nos)
results = job.result();
counts = results.get_counts()
print(counts)
# -
# ## Two-qubit gates
# ### CNOT
# * The most important two-qubit gate is controlled-NOT (CNOT) gate.
#
# * One qubit is called CONTROL, the other one is called TARGET.
#
# * The gate simply applies NOT gate to the TARGET, if CONTROL is in |1> state.
# * If CONTROL is in |0> state, it does nothing.
# +
qrs = 2
crs = 2
circuit_name = 'cnot'
nos = 8192
backend_name = 'qasm_simulator'
backend = Aer.get_backend(backend_name)
circuit, qreg, creg = anf.create_circuit_draft(qrs=qrs, crs=crs, circuit_name=circuit_name)
# CONTROL QUBIT, TARGET QUBIT
circuit.cx(qreg[0], qreg[1])
# Measure
circuit.measure(qreg[0], creg[0])
circuit.measure(qreg[1], creg[1])
# perform experiments
job = execute(circuit, backend=backend, shots=nos)
results = job.result();
counts = results.get_counts()
print(counts)
# -
# * Since our control qubit is in |0> state, our gate does nothing.
print(circuit)
# ### Tasks
# * a) Check how CNOT works for the other basis states |01>, |10>, |11>
# * b) Check what happens if you put Hadamard gates on BOTH qubits BEFORE and AFTER cnot gate.
# ## SWAP gate
# * Next very important two-qubit gate is called SWAP. It swaps two states with each other (in terms of tensor product).
# ### Task
# * c) Implement two-qubit circuit, which starts with one qubit in |0> state and other in |1> state. (it was done previously)
# * d) Swap the qubits using circuit.swap(qreg[q0],qreg[q1]) and look how they swap in statevector_simulator.
# * e) Try this for some superposition states.
# ## Bell states
# * Most famous two-qubit entangled states are so-called Bell states.
# * For Bell states, (in ideal scenario) we have always perfect correlations between measurement outcomes.
# ### Task
# * e) Try to figure out how to create any of the Bell states using standard quantum gates.
# (hint: you will need X, H and CNOT).
# * f) Create a quantum circuit which implements chosen Bell state. Implement it. You may wish to use 'visualization.plot_histogram(counts)' for histogram of results.
|
exercises/CW_0_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:env]
# language: python
# name: conda-env-env-py
# ---
import numpy as np
# ### definitions
# **underflow:** numbers near zero are rounded to zero
#
# **overflow:** large-magnitude numbers are approximated as $\infty$ or $-\infty$
# **softmax:**
#
# $$\text{softmax}(\boldsymbol{x})_{i} = \frac{\text{exp}(x_{i})}{\sum_{j=1}^{n} \text{exp}(x_{j})}$$
#
# Softmax must be stabilized against underflow and overflow.
def softmax(x):
print("\t(denominator =", np.sum(np.exp(x)), ")")
return np.exp(x)/np.sum(np.exp(x))
# non-problematic example
x = [4.0, 6.0, 8.0]
print("x =", x, "\n")
print("softmax(x) =", softmax(x), "\n")
print("sum(softmax(x)) =", np.sum(softmax(x)), "\n")
# underflow example: denominator will become 0, so result is undefined
x = [-4e100, -6e100, -8e100]
print("x =", x, "\n")
print("softmax(x) =", softmax(x), "\n")
print("sum(softmax(x)) =", np.sum(softmax(x)), "\n")
# overflow example: denominator will become inf, so result is undefined
x = [4e100, 6e100, 8e100]
print("x =", x, "\n")
print("softmax(x) =", softmax(x), "\n")
print("sum(softmax(x)) =", np.sum(softmax(x)), "\n")
# **improved softmax**
#
# $$\text{softmax}(\boldsymbol{z})_{i},\quad \boldsymbol{z} = \boldsymbol{x} - \text{max}_{i}x_{i}$$
def improved_softmax(x):
print("\t(max =", np.max(x), ")")
z = x - np.max(x)
print("\t(z =", z, ")")
print("\t(denominator =", np.sum(np.exp(z)), ")")
return np.exp(z)/np.sum(np.exp(z))
# non-problematic example
x = [4.0, 6.0, 8.0]
print("x =", x, "\n")
print("improved_softmax(x) =", improved_softmax(x), "\n")
print("sum(improved_softmax(x)) =", np.sum(improved_softmax(x)), "\n")
# underflow example: denominator will become 0, so result is undefined
x = [-4e100, -6e100, -8e100]
print("x =", x, "\n")
print("improved_softmax(x) =", improved_softmax(x), "\n")
print("sum(improved_softmax(x)) =", np.sum(improved_softmax(x)), "\n")
# overflow example: denominator will become inf, so result is undefined
x = [4e100, 6e100, 8e100]
print("x =", x, "\n")
print("improved_softmax(x) =", improved_softmax(x), "\n")
print("sum(improved_softmax(x)) =", np.sum(improved_softmax(x)), "\n")
# ---
#
# Obviously, improving **softmax** doesn't resolve all possible future underflow/overflow issues.
#
# For example, $\text{log}(\text{softmax}(x))$ could lead to some values returning as $-\infty$.
# underflow example again, but taking log after
x = [-4e100, -6e100, -8e100]
print("x =", x, "\n")
print("improved_softmax(x) =", improved_softmax(x), "\n")
print("sum(improved_softmax(x)) =", np.sum(improved_softmax(x)), "\n")
print("log(improved_softmax(x)) =", np.log(improved_softmax(x)), "\n")
|
notebooks/Softmax.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Transpiler Passes and Pass Manager
# ## Introduction
# A central component of Qiskit Terra is the transpiler, which is designed for modularity and extensibility. The goal is to be able to easily write new circuit transformations (known as transpiler **passes**), and combine them with other existing passes. Which passes are chained together and in which order has a major effect on the final outcome. This pipeline is determined by a **pass manager**, which schedules the passes and also allows passes to communicate with each other by providing a shared space. In this way, the transpiler opens up the door for research into aggressive optimization of quantum circuits.
#
# In this notebook, we look at the built-in passes, how to use the pass manager, and develop a simple custom transpiler pass. In order to do the latter, we first need to introduce the internal representation of quantum circuits in Qiskit, in the form of a Directed Acyclic Graph, or **DAG**. Then, we illustrate a simple swap mapper pass, which transforms an input circuit to be compatible with a limited-connectivity quantum device.
#
# ***Before you start***: You may need to install the `pydot` library and the `graphviz` library for the DAG plotting routines. If you are using Anaconda Python, you can install both with the `conda` command. If you use your system's native Python interpreter, install `pydot` using the `pip` command, and install `graphviz` using your system's native package manager (e.g. `yum`, `apt`, `dnf`, `brew`, etc.).
from qiskit import QuantumCircuit
from qiskit.compiler import transpile
from qiskit.transpiler import PassManager
# ## PassManager object
#
# Lets you specify the set of passes you want.
circ = QuantumCircuit(3)
circ.ccx(0, 1, 2)
circ.draw(output='mpl')
from qiskit.transpiler.passes import Unroller
pass_ = Unroller(['u1', 'u2', 'u3', 'cx'])
pm = PassManager(pass_)
new_circ = pm.run(circ)
new_circ.draw(output='mpl')
# All of Qiskit's transpiler passes are accessible from ``qiskit.transpiler.passes``.
from qiskit.transpiler import passes
[pass_ for pass_ in dir(passes) if pass_[0].isupper()]
# ## Different Variants of the Same Pass
#
# There can be passes that do the same job, but in different ways. For example, the ``TrivialLayout``, ``DenseLayout`` and ``NoiseAdaptiveLayout`` all choose a layout (binding of virtual qubits to physical qubits), but use different algorithms and objectives. Similarly, the ``BasicSwap``, ``LookaheadSwap`` and ``StochasticSwap`` all insert swaps to make the circuit compatible with the coupling map. The modularity of the transpiler allows plug-and-play replacements for each pass.
#
# Below, we show the swapper passes all applied to the same circuit, to transform it to match a linear chain topology. You can see differences in performance, where the ``StochasticSwap`` is clearly the best. However, this can vary depending on the input circuit.
# +
from qiskit.transpiler import CouplingMap, Layout
from qiskit.transpiler.passes import BasicSwap, LookaheadSwap, StochasticSwap
coupling = [[0, 1], [1, 2], [2, 3], [3, 4], [4, 5], [5, 6]]
circuit = QuantumCircuit(7)
circuit.h(3)
circuit.cx(0, 6)
circuit.cx(6, 0)
circuit.cx(0, 1)
circuit.cx(3, 1)
circuit.cx(3, 0)
coupling_map = CouplingMap(couplinglist=coupling)
bs = BasicSwap(coupling_map=coupling_map)
pass_manager = PassManager(bs)
basic_circ = pass_manager.run(circuit)
ls = LookaheadSwap(coupling_map=coupling_map)
pass_manager = PassManager(ls)
lookahead_circ = pass_manager.run(circuit)
ss = StochasticSwap(coupling_map=coupling_map)
pass_manager = PassManager(ss)
stochastic_circ = pass_manager.run(circuit)
# -
circuit.draw(output='mpl')
basic_circ.draw(output='mpl')
lookahead_circ.draw(output='mpl')
stochastic_circ.draw(output='mpl')
# ## Preset Pass Managers
#
# Qiskit comes with several pre-defined pass managers, corresponding to various levels of optimization achieved through different pipelines of passes. Currently ``optimization_level`` 0 through 3 are supported; the higher the number, the more optimized it is, at the expense of more time. Choosing a good pass manager may take trial and error, as it depends heavily on the circuit being transpiled and the backend being targeted.
#
# Here we illustrate the different levels by looking at a state synthesis circuit. We initialize four qubits to an arbitrary state, and then try to optimize the circuit that achieves this.
#
# - ``optimization_level=0``: just maps the circuit to the backend, with no explicit optimization (except whatever optimizations the mapper does).
#
# - ``optimization_level=1``: maps the circuit, but also does light-weight optimizations by collapsing adjacent gates.
#
# - ``optimization_level=2``: medium-weight optimization, including a noise-adaptive layout and a gate-cancellation procedure based on gate commutation relationships.
#
# - ``optimization_level=3``: heavy-weight optimization, which in addition to previous steps, does resynthesis of two-qubit blocks of gates in the circuit.
# +
import math
from qiskit.test.mock import FakeTokyo
backend = FakeTokyo() # mimics the tokyo device in terms of coupling map and basis gates
# +
qc = QuantumCircuit(10)
random_state = [
1 / math.sqrt(4) * complex(0, 1),
1 / math.sqrt(8) * complex(1, 0),
0,
0,
0,
0,
0,
0,
1 / math.sqrt(8) * complex(1, 0),
1 / math.sqrt(8) * complex(0, 1),
0,
0,
0,
0,
1 / math.sqrt(4) * complex(1, 0),
1 / math.sqrt(8) * complex(1, 0)]
qc.initialize(random_state, range(4))
qc.draw()
# -
# Now map this to the 20-qubit Tokyo device, with different optimization levels:
optimized_0 = transpile(qc, backend=backend, seed_transpiler=11, optimization_level=0)
print('gates = ', optimized_0.count_ops())
print('depth = ', optimized_0.depth())
optimized_1 = transpile(qc, backend=backend, seed_transpiler=11, optimization_level=1)
print('gates = ', optimized_1.count_ops())
print('depth = ', optimized_1.depth())
optimized_2 = transpile(qc, backend=backend, seed_transpiler=11, optimization_level=2)
print('gates = ', optimized_2.count_ops())
print('depth = ', optimized_2.depth())
optimized_3 = transpile(qc, backend=backend, seed_transpiler=11, optimization_level=3)
print('gates = ', optimized_3.count_ops())
print('depth = ', optimized_3.depth())
# ## Introducing the DAG
# In Qiskit, we represent circuits internally using a Directed Acyclic Graph (DAG). The advantage of this representation over a pure list of gates (i.e., *netlist*) is that the flow of information between operations are explicit, making it easier for passes to make transformation decisions without changing the semantics of the circuit.
#
# Let's start by building a simple circuit, and examining its DAG.
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit
from qiskit.dagcircuit import DAGCircuit
q = QuantumRegister(3, 'q')
c = ClassicalRegister(3, 'c')
circ = QuantumCircuit(q, c)
circ.h(q[0])
circ.cx(q[0], q[1])
circ.measure(q[0], c[0])
circ.rz(0.5, q[1]).c_if(c, 2)
circ.draw(output='mpl')
# In the DAG, there are three kinds of graph nodes: qubit/clbit input nodes (green), operation nodes (blue), and output nodes (red). Each edge indicates data flow (or dependency) between two nodes.
from qiskit.converters import circuit_to_dag
from qiskit.tools.visualization import dag_drawer
dag = circuit_to_dag(circ)
dag_drawer(dag)
# Therefore, writing a transpiler pass means using Qiskit's DAGCircuit API to analyze or transform the circuit. Let's see some examples of this.
# **a. Get all op nodes in the DAG:**
dag.op_nodes()
# Each node is an instance of the ``DAGNode`` class. Let's examine the information stored in the second op node.
node = dag.op_nodes()[3]
print("node name: ", node.name)
print("node op: ", node.op)
print("node qargs: ", node.qargs)
print("node cargs: ", node.cargs)
print("node condition: ", node.condition)
# **b. Add an operation to the back:**
# + tags=["nbsphinx-thumbnail"]
from qiskit.circuit.library import HGate
dag.apply_operation_back(HGate(), qargs=[q[0]])
dag_drawer(dag)
# -
# **c. Add an operation to the front:**
from qiskit.circuit.library import CCXGate
dag.apply_operation_front(CCXGate(), qargs=[q[0], q[1], q[2]], cargs=[])
dag_drawer(dag)
# **d. Substitute a node with a subcircuit:**
# +
from qiskit.circuit.library import CHGate, U2Gate, CXGate
mini_dag = DAGCircuit()
p = QuantumRegister(2, "p")
mini_dag.add_qreg(p)
mini_dag.apply_operation_back(CHGate(), qargs=[p[1], p[0]])
mini_dag.apply_operation_back(U2Gate(0.1, 0.2), qargs=[p[1]])
# substitute the cx node with the above mini-dag
cx_node = dag.op_nodes(op=CXGate).pop()
dag.substitute_node_with_dag(node=cx_node, input_dag=mini_dag, wires=[p[0], p[1]])
dag_drawer(dag)
# -
# Finally, after all transformations are complete, we can convert back to a regular QuantumCircuit object.
# This is what the transpiler does! It takes a circuit, operates on it in DAG form, and outputs a transformed circuit.
from qiskit.converters import dag_to_circuit
circuit = dag_to_circuit(dag)
circuit.draw(output='mpl')
# ## Implementing a BasicMapper Pass
# Now that we are familiar with the DAG, let's use it to write a transpiler pass. Here we will implement a basic pass for mapping an arbitrary circuit to a device with limited qubit connectivity. We call this the BasicMapper. This pass is included in Qiskit Terra as well.
#
# The first thing to do when writing a transpiler pass is to decide whether the pass class derives from a ``TransformationPass`` or ``AnalysisPass``. Transformation passes modify the circuit, while analysis passes only collect information about a circuit (to be used by other passes). Then, the ``run(dag)`` method is implemented, which does the main task. Finally, the pass is registered inside the ``qiskit.transpiler.passes`` module.
#
# This pass functions as follows: it traverses the DAG layer-by-layer (each layer is a group of operations that does not act on independent qubits, so in theory all operations in a layer can be done independently). For each operation, if it does not already meet the coupling map constraints, the pass identifies a swap path and inserts swaps to bring the two qubits close to each other.
#
# Follow the comments in the code for more details.
# +
from copy import copy
from qiskit.transpiler.basepasses import TransformationPass
from qiskit.transpiler import Layout
from qiskit.circuit.library import SwapGate
class BasicSwap(TransformationPass):
"""Maps (with minimum effort) a DAGCircuit onto a `coupling_map` adding swap gates."""
def __init__(self,
coupling_map,
initial_layout=None):
"""Maps a DAGCircuit onto a `coupling_map` using swap gates.
Args:
coupling_map (CouplingMap): Directed graph represented a coupling map.
initial_layout (Layout): initial layout of qubits in mapping
"""
super().__init__()
self.coupling_map = coupling_map
self.initial_layout = initial_layout
def run(self, dag):
"""Runs the BasicSwap pass on `dag`.
Args:
dag (DAGCircuit): DAG to map.
Returns:
DAGCircuit: A mapped DAG.
Raises:
TranspilerError: if the coupling map or the layout are not
compatible with the DAG.
"""
new_dag = DAGCircuit()
for qreg in dag.qregs.values():
new_dag.add_qreg(qreg)
for creg in dag.cregs.values():
new_dag.add_creg(creg)
if self.initial_layout is None:
if self.property_set["layout"]:
self.initial_layout = self.property_set["layout"]
else:
self.initial_layout = Layout.generate_trivial_layout(*dag.qregs.values())
if len(dag.qubits) != len(self.initial_layout):
raise TranspilerError('The layout does not match the amount of qubits in the DAG')
if len(self.coupling_map.physical_qubits) != len(self.initial_layout):
raise TranspilerError(
"Mappers require to have the layout to be the same size as the coupling map")
canonical_register = dag.qregs['q']
trivial_layout = Layout.generate_trivial_layout(canonical_register)
current_layout = trivial_layout.copy()
for layer in dag.serial_layers():
subdag = layer['graph']
for gate in subdag.two_qubit_ops():
physical_q0 = current_layout[gate.qargs[0]]
physical_q1 = current_layout[gate.qargs[1]]
if self.coupling_map.distance(physical_q0, physical_q1) != 1:
# Insert a new layer with the SWAP(s).
swap_layer = DAGCircuit()
swap_layer.add_qreg(canonical_register)
path = self.coupling_map.shortest_undirected_path(physical_q0, physical_q1)
for swap in range(len(path) - 2):
connected_wire_1 = path[swap]
connected_wire_2 = path[swap + 1]
qubit_1 = current_layout[connected_wire_1]
qubit_2 = current_layout[connected_wire_2]
# create the swap operation
swap_layer.apply_operation_back(SwapGate(),
qargs=[qubit_1, qubit_2],
cargs=[])
# layer insertion
order = current_layout.reorder_bits(new_dag.qubits)
new_dag.compose(swap_layer, qubits=order)
# update current_layout
for swap in range(len(path) - 2):
current_layout.swap(path[swap], path[swap + 1])
order = current_layout.reorder_bits(new_dag.qubits)
new_dag.compose(subdag, qubits=order)
return new_dag
# -
# Let's test this pass on a small example circuit.
q = QuantumRegister(7, 'q')
in_circ = QuantumCircuit(q)
in_circ.h(q[0])
in_circ.cx(q[0], q[4])
in_circ.cx(q[2], q[3])
in_circ.cx(q[6], q[1])
in_circ.cx(q[5], q[0])
in_circ.rz(0.1, q[2])
in_circ.cx(q[5], q[0])
# Now we construct a pass manager that contains our new pass. We pass the example circuit above to this pass manager, and obtain a new, transformed circuit.
# +
from qiskit.transpiler import PassManager
from qiskit.transpiler import CouplingMap
from qiskit import BasicAer
pm = PassManager()
coupling = [[0, 1], [1, 2], [2, 3], [3, 4], [4, 5], [5, 6]]
coupling_map = CouplingMap(couplinglist=coupling)
pm.append([BasicSwap(coupling_map)])
out_circ = pm.run(in_circ)
# -
in_circ.draw(output='mpl')
out_circ.draw(output='mpl')
# Note that this pass only inserts the swaps necessary to make every two-qubit interaction conform to the device coupling map. It does not, for example, care about the direction of interactions, or the native gate set supported by the device. This is a design philosophy of Qiskit's transpiler: every pass performs a small, well-defined action, and the aggressive circuit optimization is achieved by the pass manager through combining multiple passes.
# ## Transpiler Logging <a name='logging'></a>
#
# Due to the complexity of the internal operations that the transpiler is performing it's likely that you'll end up in a situation where you'd like to debug an issue or just understand more of what is happening inside the transpiler when you call it. To facilitate this the transpiler emits log messages as part of its normal operation. This logging uses the Python standard library `logging` module to emit the log messages. Python's standard logging was used because it allows Qiskit-Terra's logging to integrate in a standard way with other applications and libraries.
#
# For a more thorough introduction to Python logging refer to the [official documentation](https://docs.python.org/3/library/logging.html) and the tutorials and cookbook linked off of there.
# <div class="alert alert-block alert-success">
# <b>Note:</b> Most of the <code>logging</code> module functions used in this section adjust global settings. If you run commands in this section it might effect the output from other cells if they are run in a different order.
# </div>
# ### Configuring Python Standard Library Logging
#
# By default Python Standard Logging only prints log messages at the `WARNING`, `ERROR`, or `CRITICAL` log levels.
# Since none of the logs emitted by the transpiler use these log levels (they're all informative) you need to configure logging.
#
# The simplest way to do this is to just run:
# +
import logging
logging.basicConfig(level='DEBUG')
# -
# The `basicConfig()` function (see the docs here: https://docs.python.org/3/library/logging.html#logging.basicConfig) configures a root handler and formatter. We also specify the [log level](https://docs.python.org/3/library/logging.html#levels) to display with the `level` kwarg. Setting it to a level will also include and higher levels. For example, if you set it to `'INFO'` in addition to the `INFO` level this will also include the `WARNING`, `ERROR`, and `CRITICAL` log levels.
#
# Now the python environment in this notebook is configured to emit log messages to stderr when you run the transpiler. For example:
# <div class="alert alert-block alert-success">
# <b>Note:</b> <code>basicConfig()</code> will only work when called the first time it's called. It detects if a root handler and formatter have already been setup (either by using an earlier <code>basicConfig()</code> call or otherwise) and does nothing if they have. Further adjustments will have to by interacting with the handler directly.
# </div>
# +
from qiskit.test.mock import FakeTenerife
log_circ = QuantumCircuit(2, 2)
log_circ.h(0)
log_circ.h(1)
log_circ.h(1)
log_circ.x(1)
log_circ.cx(0, 1)
log_circ.measure([0,1], [0,1])
backend = FakeTenerife()
transpile(log_circ, backend);
# -
# As you can clearly see here when calling `transpile()` it now prints 2 types of log messages. The first is at the `INFO` log level and come from the pass manager. These indicate each pass that was executed and how long that took. The second are at the `DEBUG` level and come from the StochasticSwap pass and describes the internal operation of that pass. It's useful for debugging issues in the pass's operation.
# ### Adjusting the log level for the transpiler
#
# The qiskit transpiler uses a single namespace ``qiskit.transpiler``, as used by ``logging.getLogger('qiskit.transpiler')``. This makes it very easy to adjust the log level for just the transpiler. For example if you only wish to see log messages at the INFO level or above you can run:
logging.getLogger('qiskit.transpiler').setLevel('INFO')
transpile(log_circ, backend);
# ### Setting up logging to deal with parallel execution
#
# When running the transpiler with multiple circuits by default these circuits are transpiled in parallel. If you want to do this with logging enabled and be able to understand the output some additional steps are required.
#
# If you were just to enable logging as above and then pass `transpile()` multiple circuits you'll get results that are difficult to decipher. For example:
# Change log level back to DEBUG
logging.getLogger('qiskit.transpiler').setLevel('DEBUG')
# Transpile multiple circuits
circuits = [log_circ, log_circ]
transpile(circuits, backend);
# As you can see here we get log messages from all 3 circuits being transpiled together. There is no way to know which pass is part of which circuit's transpilation. Luckily Python logging provides tools to deal with this. The simplest one is to just change the [log formatter](https://docs.python.org/3/library/logging.html#logging.Formatter) so that includes additional information so we can associate a log message with the process it came from.
formatter = logging.Formatter('%(name)s - %(processName)-10s - %(levelname)s: %(message)s')
handler = logging.getLogger().handlers[0]
handler.setFormatter(formatter)
# Then rerun the `transpile()` call and see the new log formatter.
transpile(circuits, backend);
# Now the format for the log messages has been changed and it includes a process name for each of the transpilation processes so it's at least clear which log messages go together.
#
# There are many different options for how you can configure, this example is pretty limited. Refer to the documentation for more examples and options to build more sophisticated use cases that suit your specific use case or preferences.
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
|
tutorials/circuits_advanced/04_transpiler_passes_and_passmanager.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#pip install seaborn
# -
# # Import Libraries
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# # Read the CSV and Perform Basic Data Cleaning
# Raw dataset drop NA
df = pd.read_csv("../resources/train_predict.csv")
# Drop the null columns where all values are null
df1 = df.dropna(axis='columns', how='all')
df1.head()
#Reviewing the % of null values
100*df1.isnull().sum()/df.shape[0]
# Drop the null rows data cleaning, making all column headers lowercase
loan_df = df.dropna()
loan_df.columns=df.columns.str.lower()
loan_df.head()
#Update column names
loan_df.columns=['loan_id', 'gender', 'married', 'dependents', 'education','self_employed'
, 'income', 'co_income'
, 'loan_amount', 'loan_term', 'credit_history', 'property_area', 'loan_status']
#Test data_df after drop NAN
loan_df.dtypes
loan_df.shape
#Reviewing data
loan_df['dependents'].unique()
#Reviewing data
loan_df['self_employed'].unique()
#Reviewing data
loan_df['loan_term'].unique()
#Reviewing data
loan_df['credit_history'].unique()
loan_df.describe()
# # Select your features (columns)
# Set features. This will also be used as your x values. Removed 'loan_id', 'property_area'
loan_features_df = loan_df[['gender', 'married', 'dependents', 'education','self_employed'
, 'income', 'co_income'
, 'loan_amount', 'loan_term', 'credit_history', 'loan_status']]
loan_features_df.head()
sns.countplot(y='loan_term', hue ='loan_status',data =loan_features_df)
sns.countplot(y='married', hue ='loan_status',data =loan_features_df)
sns.countplot(y='gender', hue ='loan_status',data =loan_features_df)
sns.countplot(y='credit_history', hue ='loan_status',data =loan_features_df)
# # Create a Train Test Split
#
# Use `loan_status` for the y values
y = loan_features_df[["loan_status"]]
X = loan_features_df.drop(columns=["loan_status"])
print(X.shape, y.shape)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1, stratify=y)
#code to numberic Hold-> ‘Urban’: 3, ‘Semiurban’: 2,’Rural’: 1,
code_numeric = {'Female': 1, 'Male': 2,'Yes': 1, 'No': 2,
'Graduate': 1, 'Not Graduate': 2, 'Y': 1, 'N': 0, '3+': 3}
#code to numberic
loan_features_df = loan_features_df.applymap(lambda s: code_numeric.get(s) if s in code_numeric else s)
loan_features_df.info()
# # Pre-processing
#
# Scale the data and perform some feature selection
# Create a StandardScater model and fit it to the training data
from sklearn.preprocessing import StandardScaler
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
X_scaler = StandardScaler().fit(X_train)
# +
# Preprocessing
from tensorflow.keras.utils import to_categorical
y_train_categorical = to_categorical(y_train)
y_test_categorical = to_categorical(y_test)
# -
# # Train the Model
#
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
model = Sequential()
model.add(Dense(units=500, activation='relu', input_dim=10))
# model.add(Dense(units=100, activation='relu'))
model.add(Dense(units=2, activation='softmax'))
# -
model.summary()
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
# Fit the model to the training data
model.fit(
X_train_scaled,
y_train_categorical,
epochs=100,
shuffle=True,
verbose=2
)
# +
from sklearn.svm import SVC
model = SVC(kernel='linear')
model.fit(X_train_scaled, y_train.values.ravel())
print(f"Training Data Score: {model.score(X_train_scaled, y_train)}")
print(f"Testing Data Score: {model.score(X_test_scaled, y_test)}")
# -
from sklearn.metrics import classification_report
predictions = model.predict(X_test)
print(classification_report(y_test, predictions))
# # Hyperparameter Tuning
#
# Use `GridSearchCV` to tune the model's parameters
# Create the GridSearchCV model
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [1, 2, 10, 50],
'gamma': [0.0001, 0.0005, 0.001, 0.005]}
grid = GridSearchCV(model, param_grid, verbose=3)
# Train the model with GridSearch
grid.fit(X_train, y_train.values.ravel())
#print params, scores
print(grid.best_params_)
print(grid.best_score_)
# # Save the Model
import joblib
# +
# save your model by updating "your_name" with your name
# and "your_model" with your model variable
# be sure to turn this in to BCS
# if joblib fails to import, try running the command to install in terminal/git-bash
filename = 'Final_loan_model1.sav'
joblib.dump(model, filename)
# +
#To be done later, load the model from disk
# loaded_model = joblib.load(filename)
# result = loaded_model.score(X_test, y_test_categorical)
# print(result)
# -
|
Dawn/Loan-Model-old-ML.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/alfmorais/estrutura_de_dados_em_python/blob/main/secao_5/aula_59.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qYVeK_fTYoDM"
# Vetores Ordenados
#
# 1. Ordem crescente
# 2. Vantagem: agiliza o tempo de pesquisa
#
# Operações
#
# - Inserção:
# 1. Pesquisa uma média de N/2 elementos (pesquisa linear) - Pior caso: N
# 2. Mover elementos restantes (N/2 passos) - Pior caso: N
# 3. Big-O-O(2n) = O(n)
#
# - Pesquisa Linear:
# 1. A pesquisa termina quando o primeiro item maior que o valor da pesquisa é atingido.
# 2. Como o vetor está ordenado, o algoritmo sabe que não há necessidade de procurar mais.
# 3. Pior caso: se o elemento não estiver no vetor ou na última posição.
# 4. Big-O-O(n)
# 5. Visualização Online: https://www.cs.usfca.edu/~galles/visualization/Search.html
#
# - Exclusão:
# 1. O algoritmo pode terminar na metade do caminho se não encontrar o item
# 2. Pesquisa uma média de N/2 elementos (pesquisa linear)
# 3. Pior caso: N
# 4. Mover os elementos restantes (N/2 passos)
# 5. Pior caso: N
# 6. Big-O-O(2n) = O(n)
# + id="L4SLFk_4YksW"
|
secao_5/aula_58.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''SNN'': conda)'
# language: python
# name: python38364bitsnnconda9a8bd72d9c7f40db9654d23148b18415
# ---
# %pylab inline
# +
import torch
import torch.nn as nn
import torchvision.transforms as transforms
import torchvision.datasets as dsets
import torch.nn.functional as F
import numpy as np
from torch.autograd import Variable
from torch.optim.lr_scheduler import StepLR,MultiStepLR
import math
import keras
from torch.utils import data
import matplotlib.pyplot as plt
from datetime import datetime
def load_dataset(task='smnist'):
if task == 'smnist':
(X_train, y_train), (X_test, y_test) = keras.datasets.mnist.load_data()
# elif task == 'psmnist':
# X_train = np.load('./ps_data/ps_X_train.npy')
# X_test = np.load('./ps_data/ps_X_test.npy')
# y_train = np.load('./ps_data/Y_train.npy')
# y_test = np.load('./ps_data/Y_test.npy')
else:
print('only two task, -- smnist and psmnist')
return 0
X_train = torch.from_numpy(X_train).float()
X_test = torch.from_numpy(X_test).float()
y_train = torch.from_numpy(y_train).long()
y_test = torch.from_numpy(y_test).long()
train_dataset = data.TensorDataset(X_train,y_train) # create train datset
test_dataset = data.TensorDataset(X_test,y_test) # create test datset
return train_dataset,test_dataset
'''
STEP 3a_v2: CREATE Adaptative spike MODEL CLASS
'''
b_j0 = 0.01 # neural threshold baseline
tau_m = 20 # ms membrane potential constant
R_m = 1 # membrane resistance
dt = 1 #
gamma = .5 # gradient scale
lens = 0.5
def gaussian(x, mu=0., sigma=.5):
return torch.exp(-((x - mu) ** 2) / (2 * sigma ** 2)) / torch.sqrt(2 * torch.tensor(math.pi)) / sigma
class ActFun_adp(torch.autograd.Function):
@staticmethod
def forward(ctx, input): # input = membrane potential- threshold
ctx.save_for_backward(input)
return input.gt(0).float() # is firing ???
@staticmethod
def backward(ctx, grad_output): # approximate the gradients
input, = ctx.saved_tensors
grad_input = grad_output.clone()
# temp = abs(input) < lens
scale = 6.0
hight = .15
#temp = torch.exp(-(input**2)/(2*lens**2))/torch.sqrt(2*torch.tensor(math.pi))/lens
temp = gaussian(input, mu=0., sigma=lens) * (1. + hight) \
- gaussian(input, mu=lens, sigma=scale * lens) * hight \
- gaussian(input, mu=-lens, sigma=scale * lens) * hight
# temp = gaussian(input, mu=0., sigma=lens)
return grad_input * temp.float() * gamma
act_fun_adp = ActFun_adp.apply
def mem_update_adp(inputs, mem, spike, tau_adp,tau_m, b, dt=1, isAdapt=1):
# tau_adp = torch.FloatTensor([tau_adp])
alpha = torch.exp(-1. * dt / tau_m).cuda()
ro = torch.exp(-1. * dt / tau_adp).cuda()
# tau_adp is tau_adaptative which is learnable # add requiregredients
if isAdapt:
beta = 1.8
else:
beta = 0.
b = ro * b + (1 - ro) * spike
B = b_j0 + beta * b
mem = mem * alpha + (1 - alpha) * R_m * inputs - B * spike * dt
inputs_ = mem - B
spike = act_fun_adp(inputs_) # act_fun : approximation firing function
return mem, spike, B, b
def output_Neuron(inputs, mem, tau_m, dt=1):
"""
The read out neuron is leaky integrator without spike
"""
# alpha = torch.exp(-1. * dt / torch.FloatTensor([30.])).cuda()
alpha = torch.exp(-1. * dt / tau_m).cuda()
mem = mem * alpha + (1. - alpha) * R_m * inputs
return mem
class RNN_custom(nn.Module):
def __init__(self, input_size, stride, hidden_dims, output_size, DC_f='mem'):
super(RNN_custom, self).__init__()
self.DC_f = DC_f
self.stride = stride
self.input_size = input_size
self.output_size = output_size
self.r1_dim = hidden_dims[0]
self.r2_dim = hidden_dims[1]
self.d1_dim = hidden_dims[2]
self.i2h = nn.Linear(input_size, self.r1_dim)
self.h2h = nn.Linear(self.r1_dim, self.r1_dim)
self.h2d = nn.Linear(self.r1_dim, self.r2_dim)
self.d2d = nn.Linear(self.r2_dim, self.r2_dim)
self.dense1 = nn.Linear(self.r2_dim, self.d1_dim)
self.d2o = nn.Linear(self.d1_dim, self.output_size)
self.tau_adp_r1 = nn.Parameter(torch.Tensor(self.r1_dim))
self.tau_adp_r2 = nn.Parameter(torch.Tensor(self.r2_dim))
self.tau_adp_d1 = nn.Parameter(torch.Tensor(self.d1_dim))
self.tau_adp_o = nn.Parameter(torch.Tensor(self.output_size))
self.tau_m_r1 = nn.Parameter(torch.Tensor(self.r1_dim))
self.tau_m_r2 = nn.Parameter(torch.Tensor(self.r2_dim))
self.tau_m_d1 = nn.Parameter(torch.Tensor(self.d1_dim))
self.tau_m_o = nn.Parameter(torch.Tensor(self.output_size))
nn.init.orthogonal_(self.h2h.weight)
nn.init.xavier_uniform_(self.i2h.weight)
nn.init.xavier_uniform_(self.h2d.weight)
nn.init.xavier_uniform_(self.d2d.weight)
nn.init.xavier_uniform_(self.dense1.weight)
nn.init.xavier_uniform_(self.d2o.weight)
nn.init.constant_(self.i2h.bias, 0)
nn.init.constant_(self.h2h.bias, 0)
nn.init.constant_(self.h2d.bias, 0)
nn.init.constant_(self.d2d.bias, 0)
nn.init.constant_(self.dense1.bias, 0)
nn.init.constant_(self.d2o.bias, 0)
nn.init.normal_(self.tau_adp_r1, 700,25)
nn.init.normal_(self.tau_adp_r2, 700,25)
nn.init.normal_(self.tau_adp_o, 700,25)
nn.init.normal_(self.tau_adp_d1, 700,25)
nn.init.normal_(self.tau_m_r1, 20,5)
nn.init.normal_(self.tau_m_r2, 20,5)
nn.init.normal_(self.tau_m_o, 20,5)
nn.init.normal_(self.tau_m_d1, 20,5)
self.b_r1 =self.b_r2 = self.b_o = self.b_d1 = 0
def compute_input_steps(self,seq_num):
return int(seq_num/self.stride)
def forward(self, input):
batch_size, seq_num, input_dim = input.shape
self.b_r1 =self.b_r2 = self.b_o = self.b_d1 = b_j0
r1_mem = r1_spike = torch.rand(batch_size, self.r1_dim).cuda()
r2_mem = r2_spike = torch.rand(batch_size, self.r2_dim).cuda()
d1_mem = d1_spike = torch.rand(batch_size, self.d1_dim).cuda()
d2o_spike = output_sumspike = d2o_mem = torch.rand(batch_size, output_dim).cuda()
input = input/255.
input_steps = self.compute_input_steps(seq_num)
r1_spikes = []
r2_spikes = []
d1_spikes = []
d2_spikes = []
for i in range(input_steps):
start_idx = i*self.stride
if start_idx < (seq_num - self.input_size):
input_x = input[:, start_idx:start_idx+self.input_size, :].reshape(-1,self.input_size)
else:
input_x = input[:, -self.input_size:, :].reshape(-1,self.input_size)
#print(input_x.shape)
h_input = self.i2h(input_x.float()) + self.h2h(r1_spike)
r1_mem, r1_spike, theta_r1, self.b_r1 = mem_update_adp(h_input,r1_mem, r1_spike, self.tau_adp_r1, self.tau_m_r1,self.b_r1)
d_input = self.h2d(r1_spike) + self.d2d(r2_spike)
r2_mem, r2_spike, theta_r2, self.b_r2 = mem_update_adp(d_input, r2_mem, r2_spike, self.tau_adp_r2,self.tau_m_r2, self.b_r2)
d1_mem, d1_spike, theta_d1, self.b_d1 = mem_update_adp(self.dense1(r2_spike), d1_mem, d1_spike, self.tau_adp_d1,self.tau_m_d1, self.b_d1)
if self.DC_f[:3]=='adp':
d2o_mem, d2o_spike, theta_o, self.b_o = mem_update_adp(self.d2o(d1_spike),d2o_mem, d2o_spike, self.tau_adp_o, self.tau_m_o, self.b_o)
elif self.DC_f == 'integrator':
d2o_mem = output_Neuron(self.d2o(d1_spike),d2o_mem, self.tau_m_o)
if i >= 0:
if self.DC_f == 'adp-mem':
output_sumspike = output_sumspike + F.softmax(d2o_mem,dim=1)
elif self.DC_f =='adp-spike':
output_sumspike = output_sumspike + d2o_spike
elif self.DC_f =='integrator':
output_sumspike =output_sumspike+ F.softmax(d2o_mem,dim=1)
r1_spikes.append(r1_spike.detach().cpu().numpy())
r2_spikes.append(r2_spike.detach().cpu().numpy())
d1_spikes.append(d1_spike.detach().cpu().numpy())
d2_spikes.append(d2o_spike.detach().cpu().numpy())
return output_sumspike, [r1_spikes,r2_spikes,d1_spikes,d2_spikes]
# -
def test(model, dataloader):
correct = 0
total = 0
# Iterate through test dataset
for images, labels in dataloader:
images = images.view(-1, seq_dim, input_dim).to(device)
outputs, _ = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
if torch.cuda.is_available():
correct += (predicted.cpu() == labels.long().cpu()).sum()
else:
correct += (predicted == labels).sum()
accuracy = 100. * correct.numpy() / total
return accuracy
# +
batch_size = 200
task = 'smnist'
DC_f = 'adp-spike'
train_dataset,test_dataset = load_dataset(task)
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,batch_size=batch_size,shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,batch_size=batch_size,shuffle=False)
input_dim = 1
input_size=8
stride = 1
hidden_dims = [64,256,256]
output_dim = 10
seq_dim = int(784 / input_dim) # Number of steps to unroll
model = RNN_custom(input_size, stride,hidden_dims, output_dim,DC_f=DC_f)
model = torch.load('./model/model_98.21_Task-smnist||Time-30-06-2020 16:33:05||EC_f--rbf||DC_f--adp-spike||multiinput-multi_input.pth')
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print("device:",device)
model.to(device)
# -
accuracy = test(model,test_loader)
print('test Accuracy: ', accuracy)
i = 0
for images, labels in test_loader:
if i == 0:
i+=1
images = images.view(-1, seq_dim, input_dim).to(device)
outputs, states = model(images)
else:
break
r1_spike_np = np.array(states[0])
r2_spike_np = np.array(states[1])
d1_spike_np = np.array(states[2])
d2_spike_np = np.array(states[3])
r1_spike_np.shape,r2_spike_np.shape,d1_spike_np.shape,d2_spike_np.shape
b = r1_spike_np.shape[1]
spikes = np.zeros((784,b,64+256+256+10))
spikes[:,:,:64] = r1_spike_np
spikes[:,:,64:64+256]= r2_spike_np
spikes[:,:,64+256:64+256+256] = d1_spike_np
spikes[:,:,64+256+256:] = d2_spike_np
np.mean(r1_spike_np),np.mean(r2_spike_np),np.mean(d1_spike_np),np.mean(d2_spike_np),np.mean(spikes)
plt.imshow(spikes[:,1,:].T)
spike_count = {'total':[],'fr':[],'per step':[]}
for images, labels in test_loader:
images = images.view(-1, seq_dim, input_dim).to(device)
outputs, states = model(images)
r1_spike_np = np.array(states[0])
r2_spike_np = np.array(states[1])
d1_spike_np = np.array(states[2])
d2_spike_np = np.array(states[3])
b = r1_spike_np.shape[1]
spikes = np.zeros((784,b,64+256+256+10))
spikes[:,:,:64] = r1_spike_np
spikes[:,:,64:64+256]= r2_spike_np
spikes[:,:,64+256:64+256+256] = d1_spike_np
spikes[:,:,64+256+256:] = d2_spike_np
spike_count['total'].append([np.mean(np.sum(spikes,axis=(0,2))),np.max(np.sum(spikes,axis=(0,2))),np.min(np.sum(spikes,axis=(0,2)))])
spike_count['per step'].append([np.mean(np.sum(spikes,axis=(2))),np.max(np.sum(spikes,axis=(2))),np.min(np.sum(spikes,axis=(2)))])
spike_count['fr'].append(np.mean(spikes))
spike_total = np.array(spike_count['total'])
np.mean(spike_total[0]),np.max(spike_total[1]),np.min(spike_total[2])
spike_per = np.array(spike_count['per step'])
np.mean(spike_per[0]),np.max(spike_per[1]),np.min(spike_per[2])
spike_fr = np.array(spike_count['fr'])
np.mean(spike_fr[0])
|
NMI_vis/smnist/smnist_vis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
# filenamesjpgs.txt contains a list of JPG files in
# the 'images' directory (without the JPG extension at the end).
# To generate filenamesjpgs.txt using Terminal:
# 1. Navigate to 'images' directory.
# 2. Run the following command: ls -1 | sed -e 's/\.jpg$//' > ../filenamesjpgs.txt
jpgs = [line.rstrip() for line in open('filenamesjpgs.txt')]
# filenameslogs.txt contains a list of TXT files in
# the 'logs' directory (without the TXT extension at the end).
# To generate file using Terminal:
# 1. Navigate to 'logs' directory.
# 2. Run the following command: ls -1 | sed -e 's/\.txt$//' > ../filenameslogs.txt
logs = [line.rstrip() for line in open('filenameslogs.txt')]
for i in range(len(logs)):
logs[i] = logs[i][:-4] # Deletes '_log' suffix for each filename
# filenamesresults.txt contains a list of CSV files in
# the 'results' directory (without the CSV extension at the end).
# To generate file using Terminal:
# 1. Navigate to 'results' directory.
# 2. Run the following command: ls -1 | sed -e 's/\.csv$//' > ../filenamesresults.txt
results = [line.rstrip() for line in open('filenamesresults.txt')]
for i in range(len(results)):
results[i] = results[i][:-8] # Deletes '_results' suffix for each filename
filenames = []
error_files = []
# This part checks if for every image there exists a
# corresponding log and result file
for f in jpgs:
if f in logs and f in results:
filenames.append(f)
elif f not in logs:
error_files.append(f + " missing log file.")
elif f not in results:
error_files.append(f + " missing results file.")
else:
error_files.append(f + " invalid.")
# Save an output file filenames.txt with valid filenames.
if os.path.isfile('./filenames.txt'):
print("Delete filenames.txt and rerun script.")
else:
output_file = open('filenames.txt', "w+")
for f in filenames:
print(f, file=output_file)
output_file.close()
# Save an output file invalid_filenames.txt with invalid filenames.
if os.path.isfile('./invalid_filenames.txt'):
print("Delete invalid_filenames.txt and rerun script.")
else:
output_file = open('invalid_filenames.txt', "w+")
for f in error_files:
print(f, file=output_file)
output_file.close()
# -
|
Archive/TF_VERSION_MASKRCNN/datasets/ellipse/defectextraction/Data3TypesYminXminYmaxXmax5/mislabeled_files_remover.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# language: python
# name: python3
# ---
# Import Package
import math
import numpy as np
import tabulate
import matplotlib.pyplot as plt
# Data Entry
# +
fr=int(input('The number of Frequency: '))
fr_list=list()
for i in range(fr):
fr_list.append(float(input()))
fr_high=max(fr_list)
fr_low=min(fr_list)
fr_list
# -
# Number of classes and class interval
k = math.ceil((1 + 3.322 * math.log10(fr)))
i = math.ceil((fr_high-fr_low)/k)
k,i
# Arranginging the classes
main_list = list()
sub_list = list()
con=np.arange(fr_low,fr_high,i)
con=list(con)
con.append(fr_high)
con
# Arranging the Frequency
j=0
while j<k:
sub_list.append(con[j])
j=j+1
sub_list.append(con[j])
get=0
for key in fr_list:
if key >= con[j-1] and key < con[j] :
get+=1
elif j==k and key==con[k]:
get+=1
sub_list.append(get)
temp=sub_list.copy()
main_list.append(temp)
sub_list.clear()
main_list
# Distribution Table
# +
import tabulate
head = list()
tbl_main_list = list()
tbl_sub_list = list()
fr_get=0
j=0
while j<k:
tbl_sub_list.append(str(main_list[j][0])+' - '+str(main_list[j][1]))
tbl_sub_list.append(main_list[j][2])
if j==0:
fr_get+=main_list[j][2]
tbl_sub_list.append(fr_get)
else:
fr_get+=main_list[j][2]
tbl_sub_list.append(fr_get)
temp1 = tbl_sub_list.copy()
tbl_main_list.append(temp1)
tbl_sub_list.clear()
j+=1
head.append('Class')
head.append('Frequency')
head.append('Cumulative Frequency')
print(tabulate.tabulate(tbl_main_list,headers=head,tablefmt='fancy_grid'))
# -
# Histogram Plots for Frequency and Cumulative Frequency.
# +
import matplotlib.pyplot as plt
plt.style.use('seaborn')
fig,ax= plt.subplots(1,2)
fig.suptitle('Frequency Plot',color='blueviolet',size=14)
fig.tight_layout(pad=3)
ax[0].set_title('Frequency Plot',color='deepskyblue',size=12)
ax[0].hist(fr_list,con,color='deepskyblue',edgecolor='b',label='Frequency')
ax[0].set_xlabel('Class',color='deepskyblue',size=10)
ax[0].set_ylabel('Frequency',color='deepskyblue',size=10)
ax[0].legend()
ax[1].set_title('Cumulative Frequency Plot',color='lime',size=12)
ax[1].hist(fr_list,con,cumulative='True',color='lime',edgecolor='b',label='Cumulative Frequency')
ax[1].set_xlabel('Class',color='lime',size=10)
ax[1].set_ylabel('Cumulative Frequency',color='lime',size=10)
ax[1].legend()
plt.show()
# -
# The Project is completed.
# Thank You !
|
Main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
#
# By <NAME> and <NAME>
#
# This Jupyter Notebook will introduce to you to how to make a Linear Regression model using the Sci-kit Learn (aka `sklearn`) Python library.
#
# You can see basic example here:
# > http://scikit-learn.org/stable/modules/linear_model.html#ordinary-least-squares
#
# and full documentation of the sklearn linear_model module here:
# > http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html
#
# # Instructions
#
# 0. Make sure you've read and learned a bit about the Linear Regression model. [Click here for course notes.](https://jennselby.github.io/MachineLearningCourseNotes/#linear-regression)
# 1. Read through the instructions and code behind the following sections:
#
# * [Setup](#Setup)
# * [Fake Data Generation](#Fake-Data-Generation)
# * [Training](#Training)
# * [Results and Visualization](#Results-and-Visualization)
# 2. Then, pick and complete at least one of the set of exercises (Standard or Advanced) and write code that answers each set of questions.
# * [Option 1 - Standard Difficulty](#Exercise-Option-#1---Standard-Difficulty)
# * [Option 2 - Standard Difficulty](#Exercise-Option-#2---Standard-Difficulty)
# * [Option 3 - Advanced Difficulty](#Exercise-Option-#3---Advanced-Difficulty)
# * [Option 4 - Advanced Difficulty](#Exercise-Option-#4---Advanced-Difficulty)
# ## Setup
#
# First, make sure you have installed all of the necessary Python libraries, following [the instructions here](https://jennselby.github.io/MachineLearningCourseNotes/#setting-up-python3).
#
# You should have `sklearn`, `numpy`, `matplotlib` and `pandas` installed.
#
# If you haven't installed them, use `pip install <library here>` to install them in your Terminal.
#
# Next, we want to make sure we can display our graphs in this notebook and import all of the libraries we'll need into the notebook.
# We're going to be doing some plotting, and we want to be able to see these plots.
# To display graphs in this notebook, run this cell.
# %matplotlib inline
# +
# We're now going to import some important libraries
import numpy.random # for generating a noisy data set
from sklearn import linear_model # for training a linear model
import matplotlib.pyplot # for plotting in general
from mpl_toolkits.mplot3d import Axes3D # for 3D plotting
import pandas as pd
# -
# ## Fake Data Generation
#
# We're going to generate some fake data to test out our ideas about linear regression. These constant variables decide some of the characteristics of our data: the `x` range (which will also be used to set the size of the graph later) and how many inputs we should generate.
# +
# Setting the limits and number of our first, X, variable
MIN_X = -10
MAX_X = 10
NUM_INPUTS = 50
# -
# ### Fake Dataset 1 - Single x Variable
#
# Our first dataset has just one input feature. We are going to pick out 50 random real numbers between our min and max. Then, we will generate one output for each of these inputs following the function $y = 0.3x + 1$.
# +
# randomly pick numbers for x
x_one_x = numpy.random.uniform(low=MIN_X, high=MAX_X, size=(NUM_INPUTS, 1))
print(x_one_x)
# -
# Let's store this data into a `pandas` `DataFrame` object and name the column `'x'`.
data_one_x = pd.DataFrame(data=x_one_x, columns=['x'])
data_one_x.head()
# Cool. Now we have some fake `x` data.
#
# Let's make the fake `y` data now.
#
# Let's try to make data that follows the equation: $y = 0.3x + 1$.
data_one_x['y'] = 0.3 * data_one_x['x'] + 1
data_one_x.plot.scatter(x='x', y='y')
# Okay. That looks *too* perfect.
#
# Most data in the real world look less linear than that.
#
# So let's add a little bit of noise. Noise are random pertubations to your data that happens naturally in the real world. We will simulate some noise.
#
# Otherwise our linear model will be too easy.
#
# **Note:** We can generate some noise by picking numbers in a [normal distribution (also called bell curve)](http://www.statisticshowto.com/probability-and-statistics/normal-distributions/) around zero.
# +
# First, let's create some noise to make our data a little bit more spread out.
# generate some normally distributed noise
noise_one_x = numpy.random.normal(size=NUM_INPUTS)
# -
# Now let's create the 'y' variable
# It turns out you can make a new column in pandas just by doing the below.
# It's so simple!
data_one_x['y'] = data_one_x['y'] + noise_one_x
data_one_x.plot.scatter(x='x', y='y')
# Great!
#
# This looks more like real data now.
# ## Training
#
# Now that we have our data, we can train our model to find the best fit line. We will use the linear model module from the scikit-learn library to do this.
#
# Note: you may get a warning about LAPACK. According to [this discussion on the scikit-learn github page](https://github.com/scipy/scipy/issues/5998), this is safe to ignore.
# +
# This creates an "empty" linear model
model_one_x = linear_model.LinearRegression()
# -
# First, we need to reshape our data.
#
# Currently, our data looks like the following:
#
# ```python
# # data_one_x['x'] looks like
# [-3.44342026, 9.60082542, 4.99683803, 7.11339915, 9.69287893, ...]
#
# ```
#
# In other words, it's just a list.
#
# However, this isn't sufficient.
#
# That's because later on, we will use a command called `.fit()` and this command expects our data to look like a list of lists.
#
# For example:
#
# ```python
# [[-3.44342026],
# [ 9.60082542],
# [ 4.99683803],
# [ 7.11339915],
# [ 9.69287893],
# [-5.1383316 ],
# [ 8.96638209],
# ...
# [-9.12492363]]
# ```
#
# We will use a the command `.reshape()`.
# Run this code
x_one_x = data_one_x['x'].values.reshape(-1, 1)
y_one_x = data_one_x['y'].values.reshape(-1, 1)
# There we go. Now we can "fit" the data.
#
# "Fitting" the data means to give the "empty model" real data and ask it to find the "best parameters" that "best fits" the data.
#
# Using the amazing `sklearn` library, it's as easy as running the `.fit()` command.
#
# Note: you may get a warning about LAPACK. According to [this discussion on the scikit-learn github page](https://github.com/scipy/scipy/issues/5998), this is safe to ignore.
# Run this code
model_one_x.fit(X=x_one_x, y=y_one_x)
# ## Results and Visualization
#
# Now, let's see what our model learned. We can look at the results numerically:
def print_model_fit(model):
# Print out the parameters for the best fit line
print('Intercept: {i} Coefficients: {c}'.format(i=model.intercept_, c=model.coef_))
print_model_fit(model_one_x)
# +
## How would this model make predictions?
# Let's make some new data that have the following values and see how to predict their corresponding 'y' values.
# Print out the model's guesses for some values of x
new_x_values = [ [-1.23], [0.66], [1.98] ]
predictions = model_one_x.predict(new_x_values)
print(predictions)
# -
# Let's print them a little bit nicer
for datapoint, prediction in zip(new_x_values, predictions):
print('Model prediction for {}: {}'.format(datapoint[0], prediction))
# We can also look at them graphically.
def plot_best_fit_line(model, x, y):
# create the figure
fig = matplotlib.pyplot.figure(1)
fig.suptitle('Data and Best-Fit Line')
matplotlib.pyplot.xlabel('x values')
matplotlib.pyplot.ylabel('y values')
# put the generated dataset points on the graph
matplotlib.pyplot.scatter(x, y)
# Now we actually want to plot the best-fit line.
# To simulate that, we'll simply generate all the
# inputs on the graph and plot that.
# predict for inputs along the graph to find the best-fit line
X = numpy.linspace(MIN_X, MAX_X) # generates all the possible values of x
Y = model.predict(list(zip(X)))
matplotlib.pyplot.plot(X, Y)
plot_best_fit_line(model_one_x, x_one_x, y_one_x)
# # Exercise Option #1 - Standard Difficulty
#
# Answer the following questions about dataset 1:
# 1. Take a look at the output of the `print_model_fit()` function in the "Results and Visualization" section above. What numbers did you expect to see printed if the linear regression code was working, and why?
# 1. What numbers did you expect the model to predict when we gave it our new x values, -1.23, 0.66, and 1.98, and why?
# 1. What did you expect to see on the graph if the linear regression code was working, and why?
# 1. Pick some lines of code that you could change to continue testing that the linear regression worked properly. What lines did you choose and how did you change them? How did the output change, and why does that tell you that the code is working correctly?
# +
# -
# ### Fake Dataset 2 - Two x Values
#
# Let's look at a dataset has two inputs, like [the tree example in our notes](https://jennselby.github.io/MachineLearningCourseNotes/#linear-regression).
#
# **NOTE**: This will make it a littler harder to visualize, particularly because you cannot rotate the graph interactively in the Jupyter notebook. If you are interested in looking more closely at this graph, you can copy the code below in the next several cells into a file and run it through Python normally. This will open a graph window that will allow you to drag to rotate the graph.
# +
# generate some normally distributed noise
noise_two_x = numpy.random.normal(size=NUM_INPUTS)
# randomly pick pairs of numbers for x
x1_two_x = numpy.random.uniform(low=MIN_X, high=MAX_X, size=NUM_INPUTS)
x2_two_x = numpy.random.uniform(low=MIN_X, high=MAX_X, size=NUM_INPUTS)
y_two_x = 0.5 * x1_two_x - 2.7 * x2_two_x - 2 + noise_two_x
# -
data_two_x = pd.DataFrame(data=x1_two_x, columns = ['x1'])
data_two_x['x2'] = x2_two_x
data_two_x['y'] = y_two_x
data_two_x.head()
# +
# use scikit-learn's linear regression model and fit to our data
model_two_x = linear_model.LinearRegression()
model_two_x.fit(data_two_x[['x1', 'x2']], data_two_x['y'])
# Print out the parameters for the best fit plane
print_model_fit(model_two_x)
# +
## Now create a function that can plot in 3D
def plot_3d(model, x1, x2, y):
# 3D Plot
# create the figure
fig = matplotlib.pyplot.figure(1)
fig.suptitle('3D Data and Best-Fit Plane')
# get the current axes, and tell them to do a 3D projection
axes = fig.gca(projection='3d')
axes.set_xlabel('x1')
axes.set_ylabel('x2')
axes.set_zlabel('y')
# put the generated points on the graph
axes.scatter(x1, x2, y)
# predict for input points across the graph to find the best-fit plane
# and arrange them into a grid for matplotlib
X1 = X2 = numpy.arange(MIN_X, MAX_X, 0.05)
X1, X2 = numpy.meshgrid(X1, X2)
Y = numpy.array(model.predict(list(zip(X1.flatten(), X2.flatten())))).reshape(X1.shape)
# put the predicted plane on the graph
axes.plot_surface(X1, X2, Y, alpha=0.1)
# show the plots
matplotlib.pyplot.show()
# -
# Now let's use the function
plot_3d(model_two_x, x1_two_x, x2_two_x, y_two_x)
# # Exercise Option #2 - Standard Difficulty
#
# Now, answer the following questions about [Fake Dataset 2](#Fake-Dataset-2---Two-x-Values):
# 1. Take a look at the output of the `print_model_fit()` function for this above dataset. What output did you expect to see printed if the linear regression code was working, and why?
# 1. What did you expect to see on the graph if the linear regression code was working, and why?
# 1. Pick some lines of code that you could change to continue testing that the linear regression worked properly. What lines did you choose and how did you change them? How did the output change, and why does that tell you that the code is working correctly?
# 1. Explain any differences you noticed between working with dataset 1 and dataset 2.
# +
# -
# ### Fake Dataset 3 - Quadratic
#
# The new equation we'll try to model is $y = 0.7x^2 - 0.4x + 1.5$.
#
#
# This dataset still just has one input, so the code is very similar to our first one. However, now the generating function is quadratic, so this one will be trickier to deal with.
#
# Again, we'll go through dataset generation, training, and visualization.
# +
# randomly pick numbers for x
x_quadratic = numpy.random.uniform(low=MIN_X, high=MAX_X, size=(NUM_INPUTS, 1))
data_quadratic = pd.DataFrame(data=x_quadratic, columns=['x'])
# -
# Let's create some noise to make our data a little bit more spread out.
# generate some normally distributed noise
noise_quadratic = numpy.random.normal(size=NUM_INPUTS)
# Let's generate the y values
# Our equation:
# y = 0.7x^2 - 0.4x + 1.5
data_quadratic['y'] = 0.7 * data_quadratic['x'] * data_quadratic['x'] - 0.4 * data_quadratic['x'] + 1.5 + noise_quadratic
# +
# get a 1D array of the input data
x_quadratic = data_quadratic['x'].values.reshape(-1, 1)
y_quadratic = data_quadratic['y'].values.reshape(-1, 1)
# Let's try use scikit-learn's linear regression model and fit to our data
model_quadratic = linear_model.LinearRegression()
model_quadratic.fit(x_quadratic, y_quadratic)
# show results
print_model_fit(model_quadratic)
plot_best_fit_line(model_quadratic, x_quadratic, y_quadratic)
# -
# # Exercise Option #3 - Advanced Difficulty
#
# First, look over and understand the data for [Fake Dataset 3](#Fake-Dataset-3---Quadratic).
#
# There are some issues here. Clearly the linear model that we have isn't working great.
#
# Your challenge is to write some new code that will better fit a linear model to this data. There are a couple different ways to do this, but all of them will involve some new code. If you have ideas but just aren't sure how to translate them into code, please ask for help!
# +
### Your code here
# -
# # Exercise Option #4 - Advanced Difficulty
#
# Try adding some [regularization](https://jennselby.github.io/MachineLearningCourseNotes/#regularization-ridge-lasso-and-elastic-net) to your linear regression model. This will get you some practice in using the sci-kit learn documentation to find new functions and figure out how to use them.
#
#
# +
### Your code here
|
Examples/.ipynb_checkpoints/Linear Regression Example-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Counterfactuals guided by prototypes on Boston housing dataset
# This notebook goes through an example of [prototypical counterfactuals](../methods/CFProto.ipynb) using [k-d trees](https://en.wikipedia.org/wiki/K-d_tree) to build the prototypes. Please check out [this notebook](./cfproto_mnist.ipynb) for a more in-depth application of the method on MNIST using (auto-)encoders and trust scores.
#
# In this example, we will train a simple neural net to predict whether house prices in the Boston area are above the median value or not. We can then find a counterfactual to see which variables need to be changed to increase or decrease a house price above or below the median value.
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.ERROR) # suppress deprecation messages
from tensorflow.keras import backend as K
from tensorflow.keras.layers import Dense, Input
from tensorflow.keras.models import Model, load_model
from tensorflow.keras.utils import to_categorical
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import os
from sklearn.datasets import load_boston
from alibi.explainers import CounterFactualProto
# ## Load and prepare Boston housing dataset
boston = load_boston()
data = boston.data
target = boston.target
feature_names = boston.feature_names
# Transform into classification task: target becomes whether house price is above the overall median or not
y = np.zeros((target.shape[0],))
y[np.where(target > np.median(target))[0]] = 1
# Remove categorical feature
data = np.delete(data, 3, 1)
feature_names = np.delete(feature_names, 3)
# Explanation of remaining features:
#
# - CRIM: per capita crime rate by town
# - ZN: proportion of residential land zoned for lots over 25,000 sq.ft.
# - INDUS: proportion of non-retail business acres per town
# - RM: average number of rooms per dwelling
# - AGE: proportion of owner-occupied units built prior to 1940
# - DIS: weighted distances to five Boston employment centres
# - RAD: index of accessibility to radial highways
# - TAX: full-value property-tax rate per USD10,000
# - PTRATIO: pupil-teacher ratio by town
# - B: 1000(Bk - 0.63)^2 where Bk is the proportion of blacks by town
# - LSTAT: % lower status of the population
# Standardize data
mu = data.mean(axis=0)
sigma = data.std(axis=0)
data = (data - mu) / sigma
# Define train and test set
idx = 475
x_train,y_train = data[:idx,:], y[:idx]
x_test, y_test = data[idx:,:], y[idx:]
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# ## Train model
np.random.seed(0)
tf.set_random_seed(0)
def nn_model():
x_in = Input(shape=(12,))
x = Dense(40, activation='relu')(x_in)
x = Dense(40, activation='relu')(x)
x_out = Dense(2, activation='softmax')(x)
nn = Model(inputs=x_in, outputs=x_out)
nn.compile(loss='categorical_crossentropy', optimizer='sgd', metrics=['accuracy'])
return nn
nn = nn_model()
nn.summary()
nn.fit(x_train, y_train, batch_size=64, epochs=500, verbose=0)
nn.save('nn_boston.h5', save_format='h5')
score = nn.evaluate(x_test, y_test, verbose=0)
print('Test accuracy: ', score[1])
# ## Generate counterfactual guided by the nearest class prototype
# Original instance:
X = x_test[1].reshape((1,) + x_test[1].shape)
shape = X.shape
# Run counterfactual:
# +
# define model
nn = load_model('nn_boston.h5')
# initialize explainer, fit and generate counterfactual
cf = CounterFactualProto(nn, shape, use_kdtree=True, theta=10., max_iterations=1000,
feature_range=(x_train.min(axis=0), x_train.max(axis=0)),
c_init=1., c_steps=10)
cf.fit(x_train)
explanation = cf.explain(X)
# -
# The prediction flipped from 0 (value below the median) to 1 (above the median):
print('Original prediction: {}'.format(explanation['orig_class']))
print('Counterfactual prediction: {}'.format(explanation['cf']['class']))
# Let's take a look at the counterfactual. To make the results more interpretable, we will first undo the pre-processing step and then check where the counterfactual differs from the original instance:
orig = X * sigma + mu
counterfactual = explanation['cf']['X'] * sigma + mu
delta = counterfactual - orig
for i, f in enumerate(feature_names):
if np.abs(delta[0][i]) > 1e-4:
print('{}: {}'.format(f, delta[0][i]))
# So in order to increase the house price, the proportion of owner-occupied units built prior to 1940 should decrease by ~11-12%. This is not surprising since the proportion for the observation is very high at 93.6%. Furthermore, the % of the population with "lower status" should decrease by ~5%.
print('% owner-occupied units built prior to 1940: {}'.format(orig[0][5]))
print('% lower status of the population: {}'.format(orig[0][11]))
# Clean up:
os.remove('nn_boston.h5')
|
examples/cfproto_housing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import packages
# +
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import os
import sys
import dill
import yaml
import numpy as np
import pandas as pd
import ast
import seaborn as sns
from collections import Iterable
from collections import Counter
sns.set(style='ticks')
# -
# ### Utility functions
def flatten(x):
result = []
for el in x:
if hasattr(el, "__iter__") and not isinstance(el, str):
result.extend(flatten(el))
else:
result.append(el)
return result
def create_skills_list(df_user):
skills_one_nested_list = df_user['skills'].tolist()
skills_one_list = flatten(skills_one_nested_list)
return list(skills_one_list)
def create_skills_frequencies_dictionary(skills_list):
skills_dict_counter = {}
for skill in skills_list:
if skill not in skills_dict_counter:
skills_dict_counter[skill] = 0
skills_dict_counter[skill] += 1
return skills_dict_counter
def convert_skills_to_frequencies(skills_list):
skills_dict_counter = create_skills_frequencies_dictionary(skills_list)
skills_occurrences = list(skills_dict_counter.values())
return skills_occurrences
def set_style():
# This sets reasonable defaults for font size for a paper
sns.set_context("paper")
# Set the font to be serif
sns.set(font='serif')#, rc={'text.usetex' : True})
# Make the background white, and specify the specific font family
sns.set_style("white", {
"font.family": "serif",
"font.serif": ["Times", "Palatino", "serif"]
})
# Set tick size for axes
sns.set_style("ticks", {"xtick.major.size": 6, "ytick.major.size": 6})
def set_size(fig, width=12, height=9):
fig.set_size_inches(width, height)
plt.tight_layout()
def save_fig(fig, filename):
fig.savefig(os.path.join(VIZ_DIR, filename), dpi=600, format='pdf', bbox_inches='tight')
def plot_skills_histogram(alist):
bins = 100
title = "Distribution of skill occurrences in users."
width = 0.8
xmin = min(alist) - 1; xmax = max(alist) + 1
fig = plt.gcf()
(n, bins, patches) = plt.hist(alist, bins=bins, color="#3F5D7D", normed=False,\
histtype='bar')
plt.xticks(fontsize = 16)
plt.yticks(fontsize = 16)
plt.xlabel('Frequency',fontsize=16)
plt.ylabel('Skills',fontsize=16)
plt.xlim([xmin-10, xmax+10])
plt.yscale('log')
plt.title(title)
return fig
# ### Pre-processing of skill occurrences
generic = lambda x: ast.literal_eval(x)
df_user = pd.read_csv("/Users/smnikolakaki/Desktop/research/data-x/submodular-optimization/guru/guru_user_df.csv",
header=0,
index_col=False,
converters={'skills': generic})
skills_user_list = create_skills_list(df_user)
print('Total number of skills:',len(skills_user_list))
skills_set = list(set(skills_user_list))
print('Number of distinct skills in users:',len(skills_set))
df_skills = pd.read_csv("/Users/smnikolakaki/Desktop/research/data-x/submodular-optimization/guru/guru_skill_df.csv",
header=0,
index_col=False)
skills_list = df_skills['skill'].tolist()
print('Number of distinct skills in skills:',len(skills_list))
skill_dict = {}
for skill in skills_list:
if skill not in skill_dict:
skill_dict[skill] = 0
skill_dict[skill] += 1
skills_list = [x for x in skills_user_list if x in skill_dict]
skills_occurrences = convert_skills_to_frequencies(skills_list)
# ### Plot histogram
set_style()
fig = plot_skills_histogram(skills_occurrences)
set_size(fig, 10, 8)
# ### Plot histogram of sampled users
def sample_users(num_users,fraction=1.0):
if fraction < 1.0:
num_sampled_users = int(fraction * num_users)
sampled_users = np.random.choice(num_users, size=num_sampled_users, replace=False)
E = set(sampled_users)
else:
E = set(np.arange(num_users))
return E
def convert_sampled_skills_to_frequencies(num_users,fraction=1.0):
E = sample_users(num_users,users_sample_fraction)
list_of_available_skills = [user_skills_dict[x] for x in E]
list_of_available_skills = [y for x in list_of_available_skills for y in x]
skills_occurrences = convert_skills_to_frequencies(list_of_available_skills)
return skills_occurrences
num_users = len(df_user)
skills_nested_list = df_user['skills'].tolist()
users_list = df_user['user_id'].tolist()
user_skills_dict = {}
for i, user in enumerate(users_list):
user_skills_dict[user] = skills_nested_list[i]
users_sample_fraction = 0.05
skill_occurrences = convert_sampled_skills_to_frequencies(num_users,users_sample_fraction)
skill_occurrences_sorted = sorted(skill_occurrences)
print('Total number of skills:',len(skill_occurrences))
total_num_of_skills = len(skill_occurrences)
num_of_skills_with_less_than_3_occurrences = len([x for x in skill_occurrences_sorted if x < 3])
num_of_skills_with_more_than_200_occurrences = len([x for x in skill_occurrences_sorted if x > 50])
print('Num of skills with 2 or less occurrences:',num_of_skills_with_less_than_3_occurrences)
print('% of skills with 2 or less occurrences:',(num_of_skills_with_less_than_3_occurrences/total_num_of_skills)*100)
print('Num of skills with 200 or more occurrences:',num_of_skills_with_more_than_200_occurrences)
print('% of skills with 200 or more occurrences:',(num_of_skills_with_more_than_200_occurrences/total_num_of_skills)*100)
set_style()
fig = plot_skills_histogram(skill_occurrences)
set_size(fig, 10, 8)
users_sample_fraction = 0.2
skill_occurrences = convert_sampled_skills_to_frequencies(num_users,users_sample_fraction)
skill_occurrences_sorted = sorted(skill_occurrences)
print('Total number of skills:',len(skill_occurrences))
total_num_of_skills = len(skill_occurrences)
num_of_skills_with_less_than_3_occurrences = len([x for x in skill_occurrences_sorted if x < 3])
num_of_skills_with_more_than_200_occurrences = len([x for x in skill_occurrences_sorted if x > 50])
print('Num of skills with 2 or less occurrences:',num_of_skills_with_less_than_3_occurrences)
print('% of skills with 2 or less occurrences:',(num_of_skills_with_less_than_3_occurrences/total_num_of_skills)*100)
print('Num of skills with 200 or more occurrences:',num_of_skills_with_more_than_200_occurrences)
print('% of skills with 200 or more occurrences:',(num_of_skills_with_more_than_200_occurrences/total_num_of_skills)*100)
set_style()
fig = plot_skills_histogram(skill_occurrences)
set_size(fig, 10, 8)
users_sample_fraction = 0.1
skill_occurrences = convert_sampled_skills_to_frequencies(num_users,users_sample_fraction)
skill_occurrences_sorted = sorted(skill_occurrences)
print('Total number of skills:',len(skill_occurrences))
total_num_of_skills = len(skill_occurrences)
num_of_skills_with_less_than_3_occurrences = len([x for x in skill_occurrences_sorted if x < 3])
num_of_skills_with_more_than_200_occurrences = len([x for x in skill_occurrences_sorted if x > 50])
print('Num of skills with 2 or less occurrences:',num_of_skills_with_less_than_3_occurrences)
print('% of skills with 2 or less occurrences:',(num_of_skills_with_less_than_3_occurrences/total_num_of_skills)*100)
print('Num of skills with 200 or more occurrences:',num_of_skills_with_more_than_200_occurrences)
print('% of skills with 200 or more occurrences:',(num_of_skills_with_more_than_200_occurrences/total_num_of_skills)*100)
set_style()
fig = plot_skills_histogram(skill_occurrences)
set_size(fig, 10, 8)
|
jupyter/Skills_histogram.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # MEGSIM experimental and simulation datasets
#
#
# The MEGSIM consists of experimental and simulated MEG data
# which can be useful for reproducing research results.
#
# The MEGSIM files will be dowloaded automatically.
#
# The datasets are documented in:
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>,
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
# (2012) MEG-SIM: A Web Portal for Testing MEG Analysis Methods using
# Realistic Simulated and Empirical Data. Neuroinformatics 10:141-158
#
#
# +
import mne
from mne import find_events, Epochs, pick_types, read_evokeds
from mne.datasets.megsim import load_data
print(__doc__)
condition = 'visual' # or 'auditory' or 'somatosensory'
# Load experimental RAW files for the visual condition
raw_fnames = load_data(condition=condition, data_format='raw',
data_type='experimental', verbose=True)
# Load simulation evoked files for the visual condition
evoked_fnames = load_data(condition=condition, data_format='evoked',
data_type='simulation', verbose=True)
raw = mne.io.read_raw_fif(raw_fnames[0], verbose='error') # Bad naming
events = find_events(raw, stim_channel="STI 014", shortest_event=1)
# Visualize raw file
raw.plot()
# Make an evoked file from the experimental data
picks = pick_types(raw.info, meg=True, eog=True, exclude='bads')
# Read epochs
event_id, tmin, tmax = 9, -0.2, 0.5
epochs = Epochs(raw, events, event_id, tmin, tmax, baseline=(None, 0),
picks=picks, reject=dict(grad=4000e-13, mag=4e-12, eog=150e-6))
evoked = epochs.average() # average epochs and get an Evoked dataset.
evoked.plot(time_unit='s')
# Compare to the simulated data (use verbose='error' b/c of naming)
evoked_sim = read_evokeds(evoked_fnames[0], condition=0, verbose='error')
evoked_sim.plot(time_unit='s')
|
0.16/_downloads/plot_megsim_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Py3 research env
# language: python
# name: py3_research
# ---
# + [markdown] colab_type="text" id="eulvfJWl7ueY"
# # Lab 1
#
#
# ## Part 1: Bilingual dictionary induction and unsupervised embedding-based MT (30%)
# *Note: this homework is based on materials from yandexdataschool [NLP course](https://github.com/yandexdataschool/nlp_course/). Feel free to check this awesome course if you wish to dig deeper.*
#
# *Refined by [<NAME>](https://www.linkedin.com/in/nikolay-karpachev-b0146a104/)*
# + [markdown] colab_type="text" id="fV4rIjxa7uei"
# **In this homework** **<font color='red'>YOU</font>** will make machine translation system without using parallel corpora, alignment, attention, 100500 depth super-cool recurrent neural network and all that kind superstuff.
#
# But even without parallel corpora this system can be good enough (hopefully), in particular for similar languages, e.g. Ukrainian and Russian.
# + [markdown] colab_type="text" id="idSYq2GU7uew"
# ### Frament of the Swadesh list for some slavic languages
#
# The Swadesh list is a lexicostatistical stuff. It's named after American linguist Morris Swadesh and contains basic lexis. This list are used to define subgroupings of languages, its relatedness.
#
# So we can see some kind of word invariance for different Slavic languages.
#
#
# | Russian | Belorussian | Ukrainian | Polish | Czech | Bulgarian |
# |-----------------|--------------------------|-------------------------|--------------------|-------------------------------|-----------------------|
# | женщина | жанчына, кабета, баба | жінка | kobieta | žena | жена |
# | мужчина | мужчына | чоловік, мужчина | mężczyzna | muž | мъж |
# | человек | чалавек | людина, чоловік | człowiek | člověk | човек |
# | ребёнок, дитя | дзіця, дзіцёнак, немаўля | дитина, дитя | dziecko | dítě | дете |
# | жена | жонка | дружина, жінка | żona | žena, manželka, choť | съпруга, жена |
# | муж | муж, гаспадар | чоловiк, муж | mąż | muž, manžel, choť | съпруг, мъж |
# | мать, мама | маці, матка | мати, матір, неня, мама | matka | matka, máma, 'стар.' mateř | майка |
# | отец, тятя | бацька, тата | батько, тато, татусь | ojciec | otec | баща, татко |
# | много | шмат, багата | багато | wiele | mnoho, hodně | много |
# | несколько | некалькі, колькі | декілька, кілька | kilka | několik, pár, trocha | няколко |
# | другой, иной | іншы | інший | inny | druhý, jiný | друг |
# | зверь, животное | жывёла, звер, істота | тварина, звір | zwierzę | zvíře | животно |
# | рыба | рыба | риба | ryba | ryba | риба |
# | птица | птушка | птах, птиця | ptak | pták | птица |
# | собака, пёс | сабака | собака, пес | pies | pes | куче, пес |
# | вошь | вош | воша | wesz | veš | въшка |
# | змея, гад | змяя | змія, гад | wąż | had | змия |
# | червь, червяк | чарвяк | хробак, черв'як | robak | červ | червей |
# | дерево | дрэва | дерево | drzewo | strom, dřevo | дърво |
# | лес | лес | ліс | las | les | гора, лес |
# | палка | кій, палка | палиця | patyk, pręt, pałka | hůl, klacek, prut, kůl, pálka | палка, пръчка, бастун |
# + [markdown] colab_type="text" id="cNM3_fjr7ue2"
# But the context distribution of these languages demonstrates even more invariance. And we can use this fact for our for our purposes.
# + [markdown] colab_type="text" id="YLppwa527ue6"
# ## Data
# + colab={} colab_type="code" id="lYBGKAUn7ue_"
import gensim
import numpy as np
from gensim.models import KeyedVectors
# + [markdown] colab_type="text" id="MwGoVhRA7ufP"
# In this notebook we're going to use pretrained word vectors - FastText (original paper - https://arxiv.org/abs/1607.04606).
#
# You can download them from the official [website](https://fasttext.cc/docs/en/crawl-vectors.html). We're going to need embeddings for Russian and Ukrainian languages. Please use word2vec-compatible format (.text).
# + colab={} colab_type="code" id="u1JjQv_97ufT"
uk_emb = KeyedVectors.load_word2vec_format("cc.uk.300.vec")
# + colab={} colab_type="code" id="ffzuept_7ufd"
ru_emb = KeyedVectors.load_word2vec_format("cc.ru.300.vec")
# + colab={} colab_type="code" id="nTkXfT0W7ufk"
ru_emb.most_similar([ru_emb["август"]], topn=10)
# + colab={} colab_type="code" id="vdBA8lcg7ufs"
uk_emb.most_similar([uk_emb["серпень"]])
# + colab={} colab_type="code" id="_yJvcKXO7uf0"
ru_emb.most_similar([uk_emb["серпень"]])
# + [markdown] colab_type="text" id="pNdYAR1q7uf6"
# Load small dictionaries for correspoinding words pairs as trainset and testset.
# + colab={} colab_type="code" id="35d_DAK67uf8"
def load_word_pairs(filename):
uk_ru_pairs = []
uk_vectors = []
ru_vectors = []
with open(filename, "r") as inpf:
for line in inpf:
uk, ru = line.rstrip().split("\t")
if uk not in uk_emb or ru not in ru_emb:
continue
uk_ru_pairs.append((uk, ru))
uk_vectors.append(uk_emb[uk])
ru_vectors.append(ru_emb[ru])
return uk_ru_pairs, np.array(uk_vectors), np.array(ru_vectors)
# + colab={} colab_type="code" id="wkNL602WHJyO"
# !wget -O ukr_rus.train.txt http://tiny.cc/jfgecz
# + colab={} colab_type="code" id="uoclU6JcHCcn"
# !wget -O ukr_rus.test.txt http://tiny.cc/6zoeez
# + colab={} colab_type="code" id="05BqsdSK7ugD"
uk_ru_train, X_train, Y_train = load_word_pairs("ukr_rus.train.txt")
# + colab={} colab_type="code" id="zQOZw51r7ugL"
uk_ru_test, X_test, Y_test = load_word_pairs("ukr_rus.test.txt")
# + [markdown] colab_type="text" id="-ZBBNvpz7ugQ"
# ## Embedding space mapping (0.3 pts)
# + [markdown] colab_type="text" id="x_Dhk5gL7ugS"
# Let $x_i \in \mathrm{R}^d$ be the distributed representation of word $i$ in the source language, and $y_i \in \mathrm{R}^d$ is the vector representation of its translation. Our purpose is to learn such linear transform $W$ that minimizes euclidian distance between $Wx_i$ and $y_i$ for some subset of word embeddings. Thus we can formulate so-called Procrustes problem:
#
# $$W^*= \arg\min_W \sum_{i=1}^n||Wx_i - y_i||_2$$
# or
# $$W^*= \arg\min_W ||WX - Y||_F$$
#
# where $||*||_F$ - Frobenius norm.
# + [markdown] colab_type="text" id="acOjDdtL7ugY"
# $W^*= \arg\min_W \sum_{i=1}^n||Wx_i - y_i||_2$ looks like simple multiple linear regression (without intercept fit). So let's code.
# + colab={} colab_type="code" id="Lb-KN1be7uga"
from sklearn.linear_model import LinearRegression
# YOUR CODE HERE
# mapping = ...
# -------
# + [markdown] colab_type="text" id="X7tqJwoY7ugf"
# Let's take a look at neigbours of the vector of word _"серпень"_ (_"август"_ in Russian) after linear transform.
# + colab={} colab_type="code" id="31SrFSbn7ugi"
august = mapping.predict(uk_emb["серпень"].reshape(1, -1))
ru_emb.most_similar(august)
# + [markdown] colab_type="text" id="okSkjk597ugo"
# We can see that neighbourhood of this embedding cosists of different months, but right variant is on the ninth place.
# + [markdown] colab_type="text" id="o2uY6Y9B7ugt"
# As quality measure we will use precision top-1, top-5 and top-10 (for each transformed Ukrainian embedding we count how many right target pairs are found in top N nearest neighbours in Russian embedding space).
# + colab={} colab_type="code" id="zptuho8LAfIE"
def precision(pairs, mapped_vectors, topn=1):
"""
:args:
pairs = list of right word pairs [(uk_word_0, ru_word_0), ...]
mapped_vectors = list of embeddings after mapping from source embedding space to destination embedding space
topn = the number of nearest neighbours in destination embedding space to choose from
:returns:
precision_val, float number, total number of words for those we can find right translation at top K.
"""
assert len(pairs) == len(mapped_vectors)
num_matches = 0
for i, (_, ru) in enumerate(pairs):
# YOUR CODE HERE
precision_val = num_matches / len(pairs)
return precision_val
# + colab={} colab_type="code" id="duhj9hpv7ugy"
assert precision([("серпень", "август")], august, topn=5) == 0.0
assert precision([("серпень", "август")], august, topn=9) == 1.0
assert precision([("серпень", "август")], august, topn=10) == 1.0
# + colab={} colab_type="code" id="0-iyd5gP7ug5"
assert precision(uk_ru_test, X_test) == 0.0
assert precision(uk_ru_test, Y_test) == 1.0
# + colab={} colab_type="code" id="U-ssEJ3x7uhA"
precision_top1 = precision(uk_ru_test, mapping.predict(X_test), 1)
precision_top5 = precision(uk_ru_test, mapping.predict(X_test), 5)
# + colab={} colab_type="code" id="7K-hy7a6Ksn2"
print(precision_top1)
print(precision_top5)
# + [markdown] colab_type="text" id="hf6Ou8bx7uhH"
# ## Making it better (orthogonal Procrustean problem) (0.3 pts)
# + [markdown] colab_type="text" id="4oLs-drN7uhK"
# It can be shown (see original paper) that a self-consistent linear mapping between semantic spaces should be orthogonal.
# We can restrict transform $W$ to be orthogonal. Then we will solve next problem:
#
# $$W^*= \arg\min_W ||WX - Y||_F \text{, where: } W^TW = I$$
#
# $$I \text{- identity matrix}$$
#
# Instead of making yet another regression problem we can find optimal orthogonal transformation using singular value decomposition. It turns out that optimal transformation $W^*$ can be expressed via SVD components:
# $$X^TY=U\Sigma V^T\text{, singular value decompostion}$$
# $$W^*=UV^T$$
# + colab={} colab_type="code" id="_KSaRJFGMFiJ"
import numpy as np
# + colab={} colab_type="code" id="DdFQ7qti7uhL"
def learn_transform(X_train, Y_train):
"""
:returns: W* : float matrix[emb_dim x emb_dim] as defined in formulae above
"""
# YOUR CODE GOES HERE
# compute orthogonal embedding space mapping
# mapping = ...
return mapping
# + colab={} colab_type="code" id="7X7QfYDd7uhQ"
W = learn_transform(X_train, Y_train)
# + colab={} colab_type="code" id="OVOFYYa37uhX"
ru_emb.most_similar([np.matmul(uk_emb["серпень"], W)])
# + colab={} colab_type="code" id="r297sYP37uhb"
print(precision(uk_ru_test, np.matmul(X_test, W)))
print(precision(uk_ru_test, np.matmul(X_test, W), 5))
# + [markdown] colab_type="text" id="hvUZ72U5AfJg"
# ## Unsupervised embedding-based MT (0.4 pts)
# + [markdown] colab_type="text" id="LLyuVfHBLrJn"
# Now, let's build our word embeddings-based translator!
# + [markdown] colab_type="text" id="tPAURW1CMuP7"
# Firstly, download OPUS Tatoeba corpus.
# + colab={} colab_type="code" id="F80kUKzQMsDu"
# !wget https://object.pouta.csc.fi/OPUS-Tatoeba/v20190709/mono/uk.txt.gz
# + colab={} colab_type="code" id="0CGFZoxCUVf1"
# !gzip -d ./uk.txt.gz
# + colab={} colab_type="code" id="2MV3VvoVUX5U"
with open('./uk.txt', 'r') as f:
uk_corpus = f.readlines()
# + colab={} colab_type="code" id="tU7nPVf0UhbI"
# To save your time and CPU, feel free to use first 1000 sentences of the corpus
uk_corpus = uk_corpus[:1000]
# + colab={} colab_type="code" id="FLN8dBOXAfJ1"
# Any necessary preprocessing if needed
# YOUR CODE HERE
# + colab={} colab_type="code" id="FGksC7l_NMi9"
def translate(sentence):
"""
:args:
sentence - sentence in Ukrainian (str)
:returns:
translation - sentence in Russian (str)
* find ukrainian embedding for each word in sentence
* transform ukrainian embedding vector
* find nearest russian word and replace
"""
# YOUR CODE GOES HERE
return " ".join(translated)
# + colab={} colab_type="code" id="4hbbMy-tNxlf"
assert translate(".") == "."
assert translate("1 , 3") == "1 , 3"
assert translate("кіт зловив мишу") == "кот поймал мышку"
# + [markdown] colab_type="text" id="ia6I2ce7O_HI"
# Now you can play with your model and try to get as accurate translations as possible. **Note**: one big issue is out-of-vocabulary words. Try to think of various ways of handling it (you can start with translating each of them to a special **UNK** token and then move to more sophisticated approaches). Good luck!
# + colab={} colab_type="code" id="ap1W7ZCeOAVU"
for sent in uk_corpus[::10]:
print(translate(sent))
# -
# Great!
# See second notebook for the Neural Machine Translation assignment.
|
homeworks_advanced/Lab1_NLP/Lab1_NLP_part1_Embedding_based_MT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ginttone/test_visuallization/blob/master/1_autompg_pands_info.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8pWfzTFyQQnp"
# 정보(데이터)단계
#
# + id="9FrYULO-PBw2"
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/"} id="deC-NpK9Ql5D" outputId="ef83f9cc-7620-4077-c374-cc2de602c6f3"
# !pwd #현재 어느 디랙토리에 있니?
# + colab={"base_uri": "https://localhost:8080/"} id="jmpg5AyWRvcd" outputId="c60313b0-6718-4d8d-fb59-06387628fd85"
# !ls -l ./auto-mpg.csv #자세히보는 -l 현재 있느냐 ./
# + id="0-Khd8EFQltI"
df=pd.read_csv('./auto-mpg.csv', header=None)
# + [markdown] id="lXj0uYcpT3Bp"
# * ------------info구성
# * Range index : 행의 사이즈
# * column : 각각의 구성
# * null의 count: non-null은 널이없다 표현
# * dataset
# + colab={"base_uri": "https://localhost:8080/"} id="k94yMEWBQmRD" outputId="ef4a46c2-d09a-4de6-b3a3-b00a5a31e7e9"
df.info() #df의 모든것 보기
# + colab={"base_uri": "https://localhost:8080/", "height": 315} id="NCSyhb1GRuqm" outputId="9cc6ffa3-1cfa-44f6-9e24-bf16aa385f53"
df.describe()
# + [markdown] id="QlqRm25_WKP4"
# <데이터 구성의 분포 확인>
#
# count 398 전체<br>
# unique 94 유니크 카테고리<br>
# top 150.0 카테고리가 150번반복<br>
# freq 22<br>
# Name: 3, dtype: object<br>
#
# + colab={"base_uri": "https://localhost:8080/"} id="TIdHOQtiT0dK" outputId="a95a19fb-0a15-4dc7-f64f-660ee9eb38be"
#위 데이터중 없는 컬럼 3 확인
df[3].describe()
# + [markdown] id="gVduf9kWaoLM"
# <데이터 편중여부를 수치로 알수있다>
# * top은 가장 많이 반복된 최고 높은 카테고리를 말하는데
# * 전체(행) 398개- 탑150 = 나머지 248개
# * 데이터의 구성이 93개의 카테고리로 분할 되어있다는 것을 확인.
#
# + colab={"base_uri": "https://localhost:8080/"} id="PswDhpxrUNiV" outputId="e21525f5-924e-4aa0-c30c-ac598ac44dc2"
398 - 150.
# + colab={"base_uri": "https://localhost:8080/"} id="QqfoD6wuW3Vx" outputId="dd7de13b-647a-4ae5-a39c-c44d5cde7a68"
df[8].describe()
# + colab={"base_uri": "https://localhost:8080/"} id="UJWls-uMa4hR" outputId="bd7b39c7-b2ec-4411-ecab-47ec13e28c82"
#특정 위치 뽑아오기
#예)0컬럼 mean값만 뽑아오고 싶을때
df[0].mean()
# + colab={"base_uri": "https://localhost:8080/"} id="lrpXvYp5a4t9" outputId="ef8adb72-59f2-45bc-d951-4ea8abe84835"
#예)0컬럼 std값만 뽑아오고 싶을때
df[0].std()
# + colab={"base_uri": "https://localhost:8080/"} id="Oh1w_K3SbooT" outputId="34617667-5341-4726-f92a-c3380dc9731f"
#예)5컬럼 count값만 뽑아오고 싶을때
df[5].count()
# + colab={"base_uri": "https://localhost:8080/"} id="BJo1iSp0bxhh" outputId="7da50459-ada0-4568-fbe2-73992c5bcfef"
#예)5컬럼 min값만 뽑아오고 싶을때
df[5].min()
# + colab={"base_uri": "https://localhost:8080/"} id="0ieTAudub1iO" outputId="add7273f-d2f1-486b-ddff-a57ec4fb76a9"
#예)5컬럼 max값만 뽑아오고 싶을때
df[5].max()
# + id="pRK0mlkfJXSN"
#컬럼명 지정
df.columns = ['mpg','cylinders','displacement','horsepower','weight',
'acceleration','model year','origin','name']
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="JnXhKbjFJgZ3" outputId="18d259a7-24e0-4acd-99d2-184b6d71811f"
df.head()
# + [markdown] id="ceGZCNc7N2uJ"
# * 시각화 하는 이유는 데이터의 분포를 보고 어떻게 처리할지 판단의 기준을 세우기 위해 한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="t7B9shMZJvFR" outputId="9b23e949-c990-44f8-cc86-978826fe704f"
df.plot(x='weight',y='mpg',kind='scatter')
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="QzdbYRPvKxL8" outputId="5f62c66d-fb44-4113-d011-0427e05ca412"
#데이터 모여있어 중앙에 박스화 되었다.min과max의 정도가 박스 위아래 표시
df.plot(kind='box')
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="8PC7swirKxdq" outputId="de130181-42d0-4084-eb75-c31be8896797"
df[['mpg','acceleration']].plot(kind='box')
# + id="82Clv2FFKxuE"
# + [markdown] id="XIpN-Kd0QVwM"
# 교육단계
#
# + id="A9cXsmwmQX4U"
# + [markdown] id="MzBdGhbOQYQg"
# 서비스 단계
# + id="UnDLrdjIQbEY"
|
1_autompg_pands_info.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rome, Italy
# **Source of original dataset:** https://dati.comune.roma.it/catalog/it/dataset?q=incidenti\&sort=score+desc\%2C+dataset_last_update+desc
#
# **Location of accidents:** Latitude, Longitude
#
# **Date of accidents:** Date
#
# **Outcome of accidents:** Fatality, Injury, PDO
import pandas as pd
pd.set_option('max_columns', None)
pd.set_option('display.max_colwidth', -1)
import numpy as np
from plotly import graph_objects as go
import plotly.express as px
from itertools import chain
import matplotlib.pyplot as plt
import glob
# Setup input files
# +
data_dir = "../data/rome/"
files = glob.glob(data_dir+'*.csv')
# -
# Read original data
data = []
for x in files:
aux = pd.read_csv(x, sep=';', encoding = "ISO-8859-1")
data.append(aux)
for x in range(len(data)):
try:
data[x] = data[x].drop(['Unnamed: 37'], axis=1, errors='ignore')
except e:
pass
# Setup Longitude & Latitude columns
for i in range(len(data)):
name1 = ['Latitudine',
'Longitudine',
]
name2 = ['Latitude',
'Longitude',
]
for key1, key2 in zip(name1, name2):
try:
data_aux[i] = data_aux[i].rename(columns={key1: key2})
except:
pass
# Join the different files
list_of_dfs = data
list_of_dicts = [cur_df.T.to_dict().values() for cur_df in list_of_dfs]
data = pd.DataFrame(list(chain(*list_of_dicts)))
data
# Create Datetime column
data['Date'] = pd.to_datetime(data['DataOraIncidente'])
# Setup bicycles filter
data_bicycles = data[(data['TipoVeicolo'] == 'Velocipede')]
# Setup Longitude & Latitude columns
data_bicycles['Longitude'] = data_bicycles['Longitude'].str.replace(',','.').astype(float)
data_bicycles['Latitude'] = data_bicycles['Latitude'].str.replace(',','.').astype(float)
# Some key statistics
# +
print('Accidents between '+str(data['Date'].min())+' and '+str(data['Date'].max()))
total_accidents = data.shape[0]
print("There are a total of "+str(total_accidents)+" accidents.")
fatalities = data_bicycles["NUM_MORTI"].sum()
print("There are a total of bicyle "+str(fatalities)+" fatalities.")
injuries = data_bicycles["NUM_FERITI"].astype(float).sum()
print("There are a total of bicyle "+str(injuries)+" major injured.")
bicycles = data_bicycles.shape[0]
print("There are a total of "+str(bicycles)+" bicycles involved in all the accidents.")
# -
# Slice all bicycle accidents
data_bicycles.head()
# Save to file
print(data_bicycles.shape)
data_bicycles.to_csv('cycling_safety_rome.csv')
print('Wrote file to: cycling_safety_rome.csv')
|
dataset_curation/rome.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import random
def welcome():
print('Welcome to the game.')
def win():
print('Congratulation !you win!')
def lose():
print('Sorry you lose. ')
def gameover():
print('GAMEOVER, Goodbye!')
def get_idioms (filename):
fh=open(filename)
text=fh.read()
fh.close()
idioms=text.split()
return idioms
def get_chtable(filename):
fh=open(filename)
text=fh.read()
text.replace('\n','')
ch_table=[]
for ch in text :
if ch not in ch_table:
ch_table.append(ch)
return ch_table
def get_gusstable(idiom,ch_table):
guss_table=[ch for ch in idiom]
while len(guss_table)<6:
ch=random.choice(ch_table)
if ch not in guss_table:
guss_table.append(ch)
random.shuffle(guss_table)
for i in range(0,6,2):
print(guss_table[i],guss_table[i+1],sep=' ')
def play_gussidiom():
filename=r'd:/temp/idioms_correct.txt'
idioms=get_idioms(filename)
ch_table=get_chtable(filename)
score=10
welcome()
print('你的初始分数是:',score,'分')
while score>0:
idiom=random.choice(idioms)
get_gusstable(idiom,ch_table)
your_guss=input('请输入你猜的成语: ')
if your_guss==idiom :
score+=10
if score==100:
print('你当前的分数是:',score,'分')
win()
break
print('你猜对了!你当前的分数是:',score,'分')
elif your_guss=='':
break
else:
score-=10
print('你猜错了!你当前的分数是:',score,'分')
if score==0:
lose()
print('正确的成语是:',idiom)
gameover()
play_gussidiom()
# +
import random
filename=r'd:/temp/words.txt'
def get_words(filename):
with open(filename) as fh:
text=fh.read()
text.replace(' ','')
words=text.split()
return words
def get_key(words):
key=[]
for word in words:
n=len(word)
m=random.randrange(10**8,10**9)
key.append(str(n)+str(m))
return key
def change_word(xkey,xword):
i=0
word=[]
for ch in xword :
n=xkey[i]
if ch>'Z':
m=ord(ch)+int(n)
if m>122:
m=m%122+97
else:
m=ord(ch)+int(n)
if m>90:
m=m%90+65
word.append(chr(m))
i+=1
while len(word)<10:
num=random.randrange(97,112)
word.append(chr(num))
return word
filename=r'd:/temp/words.txt'
words=get_words(filename)
key=get_key(words)
new_words=[]
i=0
for xword in words:
xkey=key[i]
new_word=change_word(xkey,xword)
new_words.append(new_word)
i+=1
fh=open(r'd:/temp/jiami.txt','w')
for word in new_words:
fh.writelines(word)
fh.write(' ')
fh.close()
def recover_word(xword,xkey):
n=int(xkey[0])
origin_word=[]
for i in range(n):
m=ord(xword[i])
k=int(xkey[i])
if xword[i]<'Z':
if m-k<65:
m=m+25-k
else:
m-=k
else:
if m-k<97:
m=m+25-k
else:
m-=k
origin_word.append(chr(m))
return origin_word
filename=r'd:/temp/jiami.txt'
words=get_words(filename)
key=get_key(words)
origin_words=[]
i=0
for xword in words:
xkey=key[i]
origin_word=recover_word(xword,xkey)
print(origin_word)
origin_words.append(origin_word)
i+=1
fh=open(r'd:/temp/jiemi.txt','w')
for word in origin_words:
fh.writelines(word)
fh.write(' ')
fh.close()
|
chapter2/homework/computer/5-10/201611680672_test8.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cord
# language: python
# name: cord
# ---
# # Downsample Document Vectors
#
#
from cord.core import DOCUMENT_VECTOR_PATH, DOCUMENT_VECTOR_LENGTH, JSON_CATALOGS, cord_support_dir
from cord.jsonpaper import load_json_cache
from cord import ResearchPapers
import pandas as pd
from pathlib import Path, PurePath
import numpy as np
# ## 1. Load Document Vectors
document_vectors = pd.read_parquet(DOCUMENT_VECTOR_PATH)
document_vectors = document_vectors[['sha', 'pmcid','document_vector']]
document_vectors
# ## 2. Create Downsampled Vectors
# +
docvector_arr = np.stack(document_vectors.document_vector.values)
RANDOM_STATE = 42
def kmean_labels(docvectors, n_clusters=6, random_state=RANDOM_STATE):
print('Setting cluster labels')
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=n_clusters,
random_state=random_state).fit(docvectors)
return kmeans.labels_, kmeans
def tsne_embeddings(docvectors, dimensions=2):
print(f'Creating {dimensions}D embeddings')
from sklearn.manifold import TSNE
tsne = TSNE(verbose=1,
perplexity=15,
early_exaggeration=24,
n_components=dimensions,
n_jobs=8,
random_state=RANDOM_STATE,
learning_rate=600)
embeddings = tsne.fit_transform(docvectors)
return embeddings, tsne
# %time document_vector_2d, tsne2d = tsne_embeddings(docvector_arr, 2)
# %time document_vector_1d, tsne1d = tsne_embeddings(docvector_arr, 1)
# %time cluster_id, kmeans = kmean_labels(docvector_arr, 7)
# -
document_vectors['document_vector_2d'] = document_vector_2d.tolist()
document_vectors['document_vector_1d'] = document_vector_1d
document_vectors['cluster_id'] = cluster_id
# ## 3. Save Document Vectors
docvector_savepath = Path(cord_support_dir()) / f'DocumentVectors_{DOCUMENT_VECTOR_LENGTH}.pq'
document_vectors.to_parquet(docvector_savepath)
# ## 4. Save TSNE
# +
import pickle
with Path(cord_support_dir()/ 'TSNE2d.pickle').open('wb') as f:
pickle.dump(tsne2d, f)
with Path(cord_support_dir()/ 'TSNE1d.pickle').open('wb') as f:
pickle.dump(tsne1d, f)
with Path(cord_support_dir()/ 'KMeans.pickle').open('wb') as f:
pickle.dump(kmeans, f)
# -
pd.read_parquet(DOCUMENT_VECTOR_PATH)
|
job-notebooks/04-downsample-document-vectors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import json
import datetime
import time
import os
import numpy as np
import lightgbm as lgb
from config import DATA_BIM_RAW_FOLDER
from lgbm_imputer import imputer
import matplotlib
from daily_enthalpy_gradient_module import daily_enthalpy_gradients
epw_file_raw_path = os.path.join(DATA_BIM_RAW_FOLDER, 'Weather_template_epw.epw')
epw_file_raw_df = pd.read_csv(epw_file_raw_path)
|
model/3_1_weather_data_to_BIM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
using SciPy
using PyPlot
# # polynomial
x = collect(0.0:0.1:5.0)
y = sin.(x)
poly_model = odr.polynomial(3) # using third order polynomial model
data = odr.Data(x, y)
odr_obj = odr.ODR(data, poly_model)
output = odr_obj.run() # running ODR fitting
a = output.beta
y_odr = a[4].*x.^3+a[3].*x.^2+a[2].*x.^1 .+ a[1]
println(y_odr)
plt.plot(x, y, label="input data")
plt.plot(x, y_odr, label="polynomial ODR")#
plt.legend()
|
examples/odr.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
def square(number):
return number * number
# **`__map__`**
# **`__filter__`**
def is_positive(number):
if number > 0:
return True
return False
# **`all`**
# **`any`**
class Foo(object):
def __init__(self):
self.value_1 = 5
def test_method(self):
return 10
foo = Foo()
# **`getter`**
# **`hasattr`**
# **`setattr`**
# **`decorator`**
# +
>>> def memoize(f):
... pass
>>> @memoize
... def factorial(n):
... if n == 1:
... return 1
... else:
... return n * factorial(n-1)
# -
# **`generator`**
def generator():
pass
# **`coroutine`**
def coroutine():
pass
# **`@staticmethod`** and **`@classmethod`**
class Fraction(object):
def __str__(self):
return 'Please complete'
x = Fraction()
print(x)
# **`Diamond Problem`**
# +
>>> class A:
... def m(self):
... print("m of A called")
>>> class B(A):
... def m(self):
... print("m of B called")
>>> class C(A):
... def m(self):
... print("m of C called")
>>> class D(B,C):
... pass
# -
# **`MRO`**
# +
>>> class A:
... def m(self):
... print("m of A called")
>>> class B(A):
... def m(self):
... print("m of B called")
>>> class C(A):
... def m(self):
... print("m of C called")
>>> class D(B,C):
... pass
# -
# **`singleton`**
# +
>>> class Singleton(type):
... pass
>>> class SingletonClass(metaclass=Singleton):
... pass
>>> class RegularClass():
... pass
# -
# **`namedtuple`**
from collections import namedtuple
import csv
# **`Signal Handler`**
# +
import signal
import time
def receive_alarm(signum, stack):
print('Alarm :', time.ctime())
# -
# **Recursion**
# How to sum the Integers from `1` to `N`?
def sum():
pass
def tail_sum():
pass
# **Time for exercise on Binary Tree**
# - Return the count of nodes of a binary tree.
# - Return the maximum of data fields of nodes.
# - Return the height of the tree.
# - Return the number of leaves in the tree.
class Node(object):
def __init__(self, data, left=None, right=None):
self.data = data
self.left = left
self.right = right
|
00-PythonLearning/01-Tutorials/python_examples/exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch.nn.functional as F
# %load_ext autoreload
# %autoreload 2
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from dvrl.training.train import run_mnist_dvrl, run_fashion_mnist_dvrl, run_cifar_dvrl
# -
prediction_hparams = {'activation_fn': F.relu,
'predictor_lr': 1e-3,
'num_classes': 10,
'inner_batch_size': 256,
'num_inner_iterations': 15
}
dvrl_hparams = {'dve_lr': 1e-3,
'epsilon': 1e-8,
'T': 20,
'num_classes':10,
'outer_batch_size': 2000,
'exploration_threshold':0.9
}
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
mnist_pred_model, mnist_dve_model = run_mnist_dvrl(prediction_hparams=prediction_hparams, dvrl_hparams=dvrl_hparams)
# -
fasion_mnist_pred_model, fashion_mnist_dve_model = run_fashion_mnist_dvrl(prediction_hparams=prediction_hparams, dvrl_hparams=dvrl_hparams)
cifar_pred_model, cifar_dve_model = run_cifar_dvrl(prediction_hparams=prediction_hparams, dvrl_hparams=dvrl_hparams)
|
notebooks/run_dvrl.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Solscan Llama Python API Tutorial
# This tutorial aims to be a quick guide to get you started using the Solscan API integrated into messari's python library.
from messari.solscan import Solscan
ss = Solscan()
# ## get_last_block
# returns info for last num_blocks (default is 1, limit is 20)
last_block = ss.get_last_block(num_blocks=5)
last_block
# ## get_block_transactions
# get last num_transactions (default is 10) transactions of given block numbers
blocks = ['109452586', '109452587']
block_transactions = ss.get_block_transactions(blocks)
block_transactions.head()
# ## get_block
#
blocks = ['109452586', '109452587']
blocks_info = ss.get_block(blocks)
blocks_info
# ## get_last_transaction
# return last num_transactions (default 10, limit 20) transactions
last_transactions = ss.get_last_transaction(num_transactions=5)
last_transactions
# ## get_transaction
#
transactions=['T4ipYTjKUqHQpfuA8ZM5E4iJag9kX9nGhjbY974oq2ucyYRL6eWhqTjtmk3cqfqTSu8Qdce33vzKQd7bWEX3H21']
transactions_info = ss.get_transaction(transactions)
transactions_info
# ## get_account_tokens
#
accounts = ['<KEY>', '<KEY>']
account_tokens = ss.get_account_tokens(accounts)
account_tokens.head()
# ## get_account_transactions
#
accounts = ['<KEY>', '<KEY>']
account_transactions = ss.get_account_transactions(accounts)
account_transactions
# ## get_account_stake
#
accounts = ['<KEY>', '<KEY>']
account_stake = ss.get_account_stake(accounts)
# ## get_account_spl_transactions
#
accounts = ['<KEY>', '<KEY>']
account_spl_transactions = ss.get_account_spl_transactions(accounts)
# ## get_account_sol_transactions
#
accounts = ['<KEY>', '<KEY>']
account_sol_transactions = ss.get_account_sol_transactions(accounts)
# ## get_account_export_transactions
#
accounts = ['<KEY>', '<KEY>']
account_export_transactions = ss.get_account_export_transactions(accounts)
# ## get_account
#
accounts = ['<KEY>', '<KEY>']
account = ss.get_account(accounts)
# ## get_token_holders
#
token_holders = ss.get_token_holders()
# ## get_token_meta
#
tokens = ['<KEY>', '<KEY>']
token_meta = ss.get_token_meta(tokens)
# ## get_token_list
#
token_list = ss.get_token_list()
# ## get_market_info
#
market_info = ss.get_market_info()
# ## get_chain_info
#
chain_info = ss.get_chain_info()
chain_info
|
examples/notebooks/.ipynb_checkpoints/Solscan API Tutorial-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="x68NLyOxMw88"
from collections import Counter, OrderedDict
import ipywidgets as widgets
import itertools
import json
import pandas as pd
from urllib.request import urlopen
import numpy as np
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="J9l2tTp2M0hN" outputId="4afaaf97-d309-4c8f-c848-f45c9eb54e95"
union_df = pd.read_csv("master_catmodel_dataset_union_decoded.csv")
#majority_df = pd.read_csv("master_catmodel_dataset_majority_decoded.csv")
df = union_df
df['category'] = df['category'].apply(lambda x: eval(x))
df.head()
# + id="dWTRe2WLNEbW"
import nltk
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
import re
# + colab={"base_uri": "https://localhost:8080/"} id="G5e2k30QAVZS" outputId="c9f5cd15-05e7-4b59-83d4-044e83bf4a22"
nltk.download('stopwords')
STOPWORDS = stopwords.words('english')
porter = PorterStemmer()
# + id="gfGtO00vNHQx"
def preprocess(text, lower=True, stem=False,
filters="[!\"'#$%&()*\+,-./:;<=>?@\\\[\]^_`{|}~]",
stopwords=STOPWORDS):
"""Conditional preprocessing on our text unique to our task."""
# Lower
if lower:
text = text.lower()
# Remove stopwords
pattern = re.compile(r'\b(' + r'|'.join(stopwords) + r')\b\s*')
text = pattern.sub('', text)
# Spacing and filters
text = re.sub(r"([-;;.,!?<=>])", r" \1 ", text)
text = re.sub(filters, r"", text)
text = re.sub('[^A-Za-z0-9]+', ' ', text) # remove non alphanumeric chars
text = re.sub(' +', ' ', text) # remove multiple spaces
text = text.strip()
# Remove links
text = re.sub(r'http\S+', '', text)
# Stemming
if stem:
text = " ".join([porter.stem(word) for word in text.split(' ')])
return text
# + colab={"base_uri": "https://localhost:8080/"} id="M0sdZME2NiXb" outputId="3cfdfbbb-8810-4807-f531-242c84d5a17e"
# Apply to dataframe
original_df = df.copy()
df.segment_text = df.segment_text.apply(preprocess, lower=True, stem=False)
print (f"{original_df.segment_text.values[0]}\n{df.segment_text.values[0]}")
# + id="iJkbZgaONkpv"
import numpy as np
import random
# + id="SoxCIBCeNmi4"
# Set seeds for reproducibility
seed = 42
np.random.seed(seed)
random.seed(seed)
# + id="qhZDpRpHNoDg"
# Get data
X = df.segment_text.to_numpy()
y = df.category
# + id="U2oWIV-3NplY"
class LabelEncoder(object):
"""Label encoder for tag labels."""
def __init__(self, class_to_index={}):
self.class_to_index = class_to_index
self.index_to_class = {v: k for k, v in self.class_to_index.items()}
self.classes = list(self.class_to_index.keys())
def __len__(self):
return len(self.class_to_index)
def __str__(self):
return f"<LabelEncoder(num_classes={len(self)})>"
def fit(self, y):
classes = np.unique(list(itertools.chain.from_iterable(y)))
for i, class_ in enumerate(classes):
self.class_to_index[class_] = i
self.index_to_class = {v: k for k, v in self.class_to_index.items()}
self.classes = list(self.class_to_index.keys())
return self
def encode(self, y):
y_one_hot = np.zeros((len(y), len(self.class_to_index)), dtype=int)
for i, item in enumerate(y):
for class_ in item:
y_one_hot[i][self.class_to_index[class_]] = 1
return y_one_hot
def decode(self, y):
classes = []
for i, item in enumerate(y):
indices = np.where(item == 1)[0]
classes.append([self.index_to_class[index] for index in indices])
return classes
def save(self, fp):
with open(fp, 'w') as fp:
contents = {'class_to_index': self.class_to_index}
json.dump(contents, fp, indent=4, sort_keys=False)
@classmethod
def load(cls, fp):
with open(fp, 'r') as fp:
kwargs = json.load(fp=fp)
return cls(**kwargs)
# + id="4M4jJgk5Nq_i"
# Encode
label_encoder = LabelEncoder()
label_encoder.fit(y)
num_classes = len(label_encoder)
# + colab={"base_uri": "https://localhost:8080/"} id="ycQkr9IdNtFW" outputId="e282d34e-befe-454b-e3dd-99275ac5e457"
label_encoder.class_to_index
# + colab={"base_uri": "https://localhost:8080/"} id="K2N83CuyNukw" outputId="20f9e64e-1c98-4b83-b113-87beee8f8a2f"
# Sample
label_encoder.encode([["Data Retention", "International and Specific Audiences"]])
# + colab={"base_uri": "https://localhost:8080/"} id="OxH8B4A1NxTY" outputId="c8006f64-d2f2-42e2-bc58-7e802c146228"
# Encode all our labels
y = label_encoder.encode(y)
print (y.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="e_TPQjDqN0CY" outputId="d2b4f356-98e2-4bee-e133-5d334cbbaf59"
# !pip install scikit-multilearn==0.2.0 -q
# + id="psKDNtKGN1c_"
from sklearn.model_selection import train_test_split
from skmultilearn.model_selection.measures import get_combination_wise_output_matrix
# + id="5I_sp0DjN3sA"
# Split sizes
train_size = 0.7
val_size = 0.15
test_size = 0.15
# + id="MiJ5cxvfN5Ck"
# Split (train)
X_train, X_, y_train, y_ = train_test_split(X, y, train_size=train_size)
# + colab={"base_uri": "https://localhost:8080/"} id="LjXOOHtrN6aT" outputId="0e7746b6-e25c-4a50-c221-0254f3b8b10e"
print (f"train: {len(X_train)} ({(len(X_train) / len(X)):.2f})\n"
f"remaining: {len(X_)} ({(len(X_) / len(X)):.2f})")
# + id="oXQDbk17N745"
# Split (test)
X_val, X_test, y_val, y_test = train_test_split(
X_, y_, train_size=0.5)
# + colab={"base_uri": "https://localhost:8080/"} id="cDobAxBKN9ia" outputId="34ed65b5-a4a2-4450-8b0c-548add1a082e"
print(f"train: {len(X_train)} ({len(X_train)/len(X):.2f})\n"
f"val: {len(X_val)} ({len(X_val)/len(X):.2f})\n"
f"test: {len(X_test)} ({len(X_test)/len(X):.2f})")
# + id="9rZr-qniN_BB"
# Get counts for each class
counts = {}
counts['train_counts'] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_train, order=1) for combination in row)
counts['val_counts'] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_val, order=1) for combination in row)
counts['test_counts'] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_test, order=1) for combination in row)
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="TdAY-tQ5OAiD" outputId="31c34a1f-b311-46f7-b5ba-4b21a7bf2ef4"
# View distributions
pd.DataFrame({
"Train": counts["train_counts"],
"Val": counts["val_counts"],
"Test": counts["test_counts"]
}).T.fillna(0)
# + id="pQoyT8atOCF4"
# Adjust counts across splits
for k in counts["val_counts"].keys():
counts["val_counts"][k] = int(counts["val_counts"][k] * \
(train_size/val_size))
for k in counts["test_counts"].keys():
counts["test_counts"][k] = int(counts["test_counts"][k] * \
(train_size/test_size))
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="PR5k39BlODwn" outputId="ec8ece7f-1501-410d-9762-3c039a94b83f"
dist_df = pd.DataFrame({
"Train": counts["train_counts"],
"Val": counts["val_counts"],
"Test": counts["test_counts"]
}).T.fillna(0)
dist_df
# + colab={"base_uri": "https://localhost:8080/"} id="6GscNRa9OFbQ" outputId="e402d4c4-b99d-4786-8f44-7018001e5675"
# Standard deviation
np.mean(np.std(dist_df.to_numpy(), axis=0))
# + id="qm2RMCNLOHNL"
from skmultilearn.model_selection import IterativeStratification
# + id="aae3ivotOI3e"
def iterative_train_test_split(X, y, train_size):
"""Custom iterative train test split which
'maintains balanced representation with respect
to order-th label combinations.'
"""
stratifier = IterativeStratification(
n_splits=2, order=1, sample_distribution_per_fold=[1.0-train_size, train_size, ])
train_indices, test_indices = next(stratifier.split(X, y))
X_train, y_train = X[train_indices], y[train_indices]
X_test, y_test = X[test_indices], y[test_indices]
return X_train, X_test, y_train, y_test
# + id="zf5lppoMOKQA"
# Get data
X = df.segment_text.to_numpy()
y = df.category
# + id="gFo0GnUVOL3f"
# Binarize y
label_encoder = LabelEncoder()
label_encoder.fit(y)
y = label_encoder.encode(y)
# + id="GVfn1xihONUD"
# Split
X_train, X_, y_train, y_ = iterative_train_test_split(
X, y, train_size=train_size)
X_val, X_test, y_val, y_test = iterative_train_test_split(
X_, y_, train_size=0.5)
# + colab={"base_uri": "https://localhost:8080/"} id="Wxm9hfnaOOkg" outputId="0be23ca5-537e-4c24-ec4e-6548bc6911e7"
print(f"train: {len(X_train)} ({len(X_train)/len(X):.2f})\n"
f"val: {len(X_val)} ({len(X_val)/len(X):.2f})\n"
f"test: {len(X_test)} ({len(X_test)/len(X):.2f})")
# + id="gZzrMbeKOP3b"
# Get counts for each class
counts = {}
counts["train_counts"] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_train, order=1) for combination in row)
counts["val_counts"] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_val, order=1) for combination in row)
counts["test_counts"] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_test, order=1) for combination in row)
# + id="moRa_ilXORml"
# Adjust counts across splits
for k in counts["val_counts"].keys():
counts["val_counts"][k] = int(counts["val_counts"][k] * \
(train_size/val_size))
for k in counts["test_counts"].keys():
counts["test_counts"][k] = int(counts["test_counts"][k] * \
(train_size/test_size))
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="tDvfzv6TOTM3" outputId="3affba79-78fd-47f5-e286-298391ab4c71"
# View distributions
pd.DataFrame({
"Train": counts["train_counts"],
"Val": counts["val_counts"],
"Test": counts["test_counts"]
}).T.fillna(0)
# + id="dwOR9Iq1OUyu"
dist_df = pd.DataFrame({
'train': counts['train_counts'],
'val': counts['val_counts'],
'test': counts['test_counts']
}).T.fillna(0)
# + colab={"base_uri": "https://localhost:8080/"} id="vci6Rj22OWkv" outputId="30e8bc6f-f6e5-40ec-fec3-19235bc38d04"
# Standard deviation
np.mean(np.std(dist_df.to_numpy(), axis=0))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="mzKLeV-ZOaPH" outputId="fd6a2b3a-4c0e-4807-e1e0-495d8471795d"
# Split DataFrames
train_df = pd.DataFrame({"segment_text": X_train, "category": label_encoder.decode(y_train)})
val_df = pd.DataFrame({"segment_text": X_val, "category": label_encoder.decode(y_val)})
test_df = pd.DataFrame({"segment_text": X_test, "category": label_encoder.decode(y_test)})
train_df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="OUw6Lxn6Obm2" outputId="aab48a87-5b32-4dc4-ebb9-8068e90e1ea7"
# !pip install nlpaug==1.1.0 transformers==3.0.2 -q
# !pip install snorkel==0.9.7 -q
# + id="dKqh4oe1BpGC"
# + id="XD5cMQ9NBpDk"
# + id="3cZOZTfoBpAp"
# + id="Yff4ZdDdBo9k"
# + id="bxrf6Cp4Bo7o"
# + id="R_f6dIAGBo4X"
# + id="HAtUU13EBo1O"
# + id="qU-O26QCOdIS"
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.multiclass import OneVsRestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import LinearSVC
# + id="sR1TwHavOhvT"
from sklearn import metrics
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.metrics import precision_recall_curve
from sklearn.preprocessing import MultiLabelBinarizer
# + id="LfljLd8ePF_-"
from sklearn.metrics import precision_recall_fscore_support
import torch
# + id="F2boqTcyO3Ky"
def set_seeds(seed=1234):
"""Set seeds for reproducibility."""
np.random.seed(seed)
random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.cuda.manual_seed_all(seed) # multi-GPU
# + id="wB0FWqMBO6HM"
def get_data_splits(df, train_size=0.7):
""""""
# Get data
X = df.segment_text.to_numpy()
y = df.category
# Binarize y
label_encoder = LabelEncoder()
label_encoder.fit(y)
y = label_encoder.encode(y)
# Split
X_train, X_, y_train, y_ = iterative_train_test_split(
X, y, train_size=train_size)
X_val, X_test, y_val, y_test = iterative_train_test_split(
X_, y_, train_size=0.5)
return X_train, X_val, X_test, y_train, y_val, y_test, label_encoder
# + id="OaG9wvlEO6CK"
class Trainer(object):
def __init__(self, model, device, loss_fn=None, optimizer=None, scheduler=None):
# Set params
self.model = model
self.device = device
self.loss_fn = loss_fn
self.optimizer = optimizer
self.scheduler = scheduler
def train_step(self, dataloader):
"""Train step."""
# Set model to train mode
self.model.train()
loss = 0.0
# Iterate over train batches
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, targets = batch[:-1], batch[-1]
self.optimizer.zero_grad() # Reset gradients
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, targets) # Define loss
J.backward() # Backward pass
self.optimizer.step() # Update weights
# Cumulative Metrics
loss += (J.detach().item() - loss) / (i + 1)
return loss
def eval_step(self, dataloader):
"""Validation or test step."""
# Set model to eval mode
self.model.eval()
loss = 0.0
y_trues, y_probs = [], []
# Iterate over val batches
with torch.inference_mode():
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, y_true = batch[:-1], batch[-1]
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, y_true).item()
# Cumulative Metrics
loss += (J - loss) / (i + 1)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
y_trues.extend(y_true.cpu().numpy())
return loss, np.vstack(y_trues), np.vstack(y_probs)
def predict_step(self, dataloader):
"""Prediction step."""
# Set model to eval mode
self.model.eval()
y_probs = []
# Iterate over val batches
with torch.inference_mode():
for i, batch in enumerate(dataloader):
# Forward pass w/ inputs
inputs, targets = batch[:-1], batch[-1]
z = self.model(inputs)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
return np.vstack(y_probs)
def train(self, num_epochs, patience, train_dataloader, val_dataloader,
tolerance=1e-5):
best_val_loss = np.inf
for epoch in range(num_epochs):
# Steps
train_loss = self.train_step(dataloader=train_dataloader)
val_loss, _, _ = self.eval_step(dataloader=val_dataloader)
self.scheduler.step(val_loss)
# Early stopping
if val_loss < best_val_loss - tolerance:
best_val_loss = val_loss
best_model = self.model
_patience = patience # reset _patience
else:
_patience -= 1
if not _patience: # 0
print("Stopping early!")
break
# Logging
print(
f"Epoch: {epoch+1} | "
f"train_loss: {train_loss:.5f}, "
f"val_loss: {val_loss:.5f}, "
f"lr: {self.optimizer.param_groups[0]['lr']:.2E}, "
f"_patience: {_patience}"
)
return best_model
# + id="CyCPG2l_OjGR"
# Set seeds
set_seeds()
# + id="twyOEIycOnt1"
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.segment_text = preprocessed_df.segment_text.apply(preprocess, lower=True, stem=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
# + colab={"base_uri": "https://localhost:8080/"} id="oAtv-kKHOo7J" outputId="744dfb2b-8b31-4b55-da1a-f398447e596f"
# Tf-idf
vectorizer = TfidfVectorizer()
print (X_train[0])
X_train = vectorizer.fit_transform(X_train)
X_val = vectorizer.transform(X_val)
X_test = vectorizer.transform(X_test)
print (X_train.shape)
print (X_train[0]) # scipy.sparse.csr_matrix
# + id="k9n1HfgUOrDs"
def fit_and_evaluate(model):
"""Fit and evaluate each model."""
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
return {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
# + colab={"base_uri": "https://localhost:8080/"} id="j93Vq4mKOsvQ" outputId="3f0f4670-dd8c-49fc-84ed-afcb3e8ad067"
# Models
performance = {}
performance["logistic-regression"] = fit_and_evaluate(OneVsRestClassifier(
LogisticRegression(), n_jobs=1))
performance["k-nearest-neighbors"] = fit_and_evaluate(
KNeighborsClassifier())
performance["random-forest"] = fit_and_evaluate(
RandomForestClassifier(n_jobs=-1))
performance["gradient-boosting-machine"] = fit_and_evaluate(OneVsRestClassifier(
GradientBoostingClassifier()))
performance["support-vector-machine"] = fit_and_evaluate(OneVsRestClassifier(
LinearSVC(), n_jobs=-1))
print (json.dumps(performance, indent=2))
# + [markdown] id="VYTHG_7pPM_t"
# # CNN
# + id="85KFu7u1OuIQ"
import math
import torch
import torch.nn as nn
import torch.nn.functional as F
# + id="kdHZBSbKPP9x"
# Set seeds
set_seeds()
# + id="UH2eV8hyPRLb"
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.segment_text = preprocessed_df.segment_text.apply(preprocess, lower=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
X_test_raw = X_test # use for later
# + id="uW5TVwa-PSPZ"
# Split DataFrames
train_df = pd.DataFrame({"segment_text": X_train, "category": label_encoder.decode(y_train)})
val_df = pd.DataFrame({"segment_text": X_val, "category": label_encoder.decode(y_val)})
test_df = pd.DataFrame({"segment_text": X_test, "category": label_encoder.decode(y_test)})
# + colab={"base_uri": "https://localhost:8080/"} id="nykT_l46PTdG" outputId="c18545a7-1a32-4ba6-bf57-6972075af45e"
# Set device
cuda = True
device = torch.device("cuda" if (
torch.cuda.is_available() and cuda) else "cpu")
torch.set_default_tensor_type("torch.FloatTensor")
if device.type == "cuda":
torch.set_default_tensor_type("torch.cuda.FloatTensor")
print (device)
# + id="n3xZOaLtPU2h"
class Tokenizer(object):
def __init__(self, char_level, num_tokens=None,
pad_token="<PAD>", oov_token="<UNK>",
token_to_index=None):
self.char_level = char_level
self.separator = '' if self.char_level else ' '
if num_tokens: num_tokens -= 2 # pad + unk tokens
self.num_tokens = num_tokens
self.pad_token = pad_token
self.oov_token = oov_token
if not token_to_index:
token_to_index = {pad_token: 0, oov_token: 1}
self.token_to_index = token_to_index
self.index_to_token = {v: k for k, v in self.token_to_index.items()}
def __len__(self):
return len(self.token_to_index)
def __str__(self):
return f"<Tokenizer(num_tokens={len(self)})>"
def fit_on_texts(self, texts):
if not self.char_level:
texts = [text.split(" ") for text in texts]
all_tokens = [token for text in texts for token in text]
counts = Counter(all_tokens).most_common(self.num_tokens)
self.min_token_freq = counts[-1][1]
for token, count in counts:
index = len(self)
self.token_to_index[token] = index
self.index_to_token[index] = token
return self
def texts_to_sequences(self, texts):
sequences = []
for text in texts:
if not self.char_level:
text = text.split(' ')
sequence = []
for token in text:
sequence.append(self.token_to_index.get(
token, self.token_to_index[self.oov_token]))
sequences.append(np.asarray(sequence))
return sequences
def sequences_to_texts(self, sequences):
texts = []
for sequence in sequences:
text = []
for index in sequence:
text.append(self.index_to_token.get(index, self.oov_token))
texts.append(self.separator.join([token for token in text]))
return texts
def save(self, fp):
with open(fp, "w") as fp:
contents = {
"char_level": self.char_level,
"oov_token": self.oov_token,
"token_to_index": self.token_to_index
}
json.dump(contents, fp, indent=4, sort_keys=False)
@classmethod
def load(cls, fp):
with open(fp, "r") as fp:
kwargs = json.load(fp=fp)
return cls(**kwargs)
# + colab={"base_uri": "https://localhost:8080/"} id="BfzQwFPePWVh" outputId="51a4e5ca-2544-4dc6-be66-c89bd3630c58"
# Tokenize
char_level = True
tokenizer = Tokenizer(char_level=char_level)
tokenizer.fit_on_texts(texts=X_train)
vocab_size = len(tokenizer)
print (tokenizer)
# + colab={"base_uri": "https://localhost:8080/"} id="oVuDnCUgPYK4" outputId="5024aa17-47f6-44d6-f257-c772210e5cb7"
tokenizer.token_to_index
# + colab={"base_uri": "https://localhost:8080/"} id="86TZDnK1PZ4d" outputId="6f034ef6-8fd9-47c4-a5b1-829269962473"
# Convert texts to sequences of indices
X_train = np.array(tokenizer.texts_to_sequences(X_train))
X_val = np.array(tokenizer.texts_to_sequences(X_val))
X_test = np.array(tokenizer.texts_to_sequences(X_test))
preprocessed_text = tokenizer.sequences_to_texts([X_train[0]])[0]
print ("Text to indices:\n"
f" (preprocessed) → {preprocessed_text}\n"
f" (tokenized) → {X_train[0]}")
# + colab={"base_uri": "https://localhost:8080/"} id="wp8htC7XPbs6" outputId="4263945d-209b-4b3f-ae63-abb6d89e5ba1"
# Class weights
train_cats = list(itertools.chain.from_iterable(train_df.category.values))
counts = np.bincount([label_encoder.class_to_index[class_] for class_ in train_cats])
class_weights = {i: 1.0/count for i, count in enumerate(counts)}
print (f"class counts: {counts},\nclass weights: {class_weights}")
# + id="d-d828iZPdl0"
def pad_sequences(sequences, max_seq_len=0):
"""Pad sequences to max length in sequence."""
max_seq_len = max(max_seq_len, max(len(sequence) for sequence in sequences))
padded_sequences = np.zeros((len(sequences), max_seq_len))
for i, sequence in enumerate(sequences):
padded_sequences[i][:len(sequence)] = sequence
return padded_sequences
# + id="gOASToz4PfKg"
class CNNTextDataset(torch.utils.data.Dataset):
def __init__(self, X, y, max_filter_size):
self.X = X
self.y = y
self.max_filter_size = max_filter_size
def __len__(self):
return len(self.y)
def __str__(self):
return f"<Dataset(N={len(self)})>"
def __getitem__(self, index):
X = self.X[index]
y = self.y[index]
return [X, y]
def collate_fn(self, batch):
"""Processing on a batch."""
# Get inputs
batch = np.array(batch, dtype=object)
X = batch[:, 0]
y = np.stack(batch[:, 1], axis=0)
# Pad inputs
X = pad_sequences(sequences=X, max_seq_len=self.max_filter_size)
# Cast
X = torch.LongTensor(X.astype(np.int32))
y = torch.FloatTensor(y.astype(np.int32))
return X, y
def create_dataloader(self, batch_size, shuffle=False, drop_last=False):
return torch.utils.data.DataLoader(
dataset=self,
batch_size=batch_size,
collate_fn=self.collate_fn,
shuffle=shuffle,
drop_last=drop_last,
pin_memory=True)
# + colab={"base_uri": "https://localhost:8080/"} id="g8XZcdCqPgn8" outputId="0b542adf-ed7a-4a3b-f68d-7177aaa7e239"
# Create datasets
filter_sizes = list(range(1, 11))
train_dataset = CNNTextDataset(
X=X_train, y=y_train, max_filter_size=max(filter_sizes))
val_dataset = CNNTextDataset(
X=X_val, y=y_val, max_filter_size=max(filter_sizes))
test_dataset = CNNTextDataset(
X=X_test, y=y_test, max_filter_size=max(filter_sizes))
print ("Data splits:\n"
f" Train dataset:{train_dataset.__str__()}\n"
f" Val dataset: {val_dataset.__str__()}\n"
f" Test dataset: {test_dataset.__str__()}\n"
"Sample point:\n"
f" X: {train_dataset[0][0]}\n"
f" y: {train_dataset[0][1]}")
# + colab={"base_uri": "https://localhost:8080/"} id="S7Mh8-McPiOm" outputId="8d72f28b-fba5-4835-b489-bc4f75c7ace4"
# Create dataloaders
batch_size = 128
train_dataloader = train_dataset.create_dataloader(
batch_size=batch_size)
val_dataloader = val_dataset.create_dataloader(
batch_size=batch_size)
test_dataloader = test_dataset.create_dataloader(
batch_size=batch_size)
batch_X, batch_y = next(iter(train_dataloader))
print ("Sample batch:\n"
f" X: {list(batch_X.size())}\n"
f" y: {list(batch_y.size())}")
# + id="58PQn3k5Pjy4"
# Arguments
embedding_dim = 504
num_filters = 472
hidden_dim = 387
dropout_p = 0.6065003167861672
# + id="4Q_gxeszPljS"
class CNN(nn.Module):
def __init__(self, embedding_dim, vocab_size, num_filters, filter_sizes,
hidden_dim, dropout_p, num_classes, padding_idx=0):
super(CNN, self).__init__()
# Initialize embeddings
self.embeddings = nn.Embedding(
embedding_dim=embedding_dim, num_embeddings=vocab_size,
padding_idx=padding_idx)
# Conv weights
self.filter_sizes = filter_sizes
self.conv = nn.ModuleList(
[nn.Conv1d(in_channels=embedding_dim,
out_channels=num_filters,
kernel_size=f) for f in filter_sizes])
# FC weights
self.dropout = nn.Dropout(dropout_p)
self.fc1 = nn.Linear(num_filters*len(filter_sizes), hidden_dim)
self.fc2 = nn.Linear(hidden_dim, num_classes)
def forward(self, inputs, channel_first=False):
# Embed
x_in, = inputs
x_in = self.embeddings(x_in)
if not channel_first:
x_in = x_in.transpose(1, 2) # (N, channels, sequence length)
z = []
max_seq_len = x_in.shape[2]
for i, f in enumerate(self.filter_sizes):
# `SAME` padding
padding_left = int(
(self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2)
padding_right = int(math.ceil(
(self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2))
# Conv
_z = self.conv[i](F.pad(x_in, (padding_left, padding_right)))
# Pool
_z = F.max_pool1d(_z, _z.size(2)).squeeze(2)
z.append(_z)
# Concat outputs
z = torch.cat(z, 1)
# FC
z = self.fc1(z)
z = self.dropout(z)
z = self.fc2(z)
return z
# + colab={"base_uri": "https://localhost:8080/"} id="54wUYC80Pm28" outputId="d92e52a4-42ac-4c60-b79b-e48de61fa5dd"
# Initialize model
model = CNN(
embedding_dim=embedding_dim, vocab_size=vocab_size,
num_filters=num_filters, filter_sizes=filter_sizes,
hidden_dim=hidden_dim, dropout_p=dropout_p,
num_classes=num_classes)
model = model.to(device)
print (model.named_parameters)
# + id="KukE0mfOPogG"
# Arguments
lr = 0.0002656053142937365
num_epochs = 100
patience = 10
# + id="DXGNLAyAPqYf"
# Define loss
class_weights_tensor = torch.Tensor(np.array(list(class_weights.values())))
loss_fn = nn.BCEWithLogitsLoss(weight=class_weights_tensor)
# + id="rqSCUJ08Prqm"
# Define optimizer & scheduler
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(
optimizer, mode="min", factor=0.1, patience=5)
# + id="2vTyNhQRPtYF"
# Trainer module
trainer = Trainer(
model=model, device=device, loss_fn=loss_fn,
optimizer=optimizer, scheduler=scheduler)
# + colab={"base_uri": "https://localhost:8080/"} id="JrL6uj8WPu7K" outputId="8e826c97-3f92-4770-feb6-02b1e438e334"
# Train
best_model = trainer.train(
num_epochs, patience, train_dataloader, val_dataloader)
# + id="8I-tXJyzPxNd"
from pathlib import Path
from sklearn.metrics import precision_recall_curve
# + id="-vn6G9xOP3AD"
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
from wordcloud import WordCloud, STOPWORDS
sns.set_theme()
warnings.filterwarnings("ignore")
# + colab={"base_uri": "https://localhost:8080/", "height": 302} id="gpFGh0GnP7zR" outputId="aff962ab-3bf2-4170-ff0a-0c0e8a3daa86"
# Threshold-PR curve
train_loss, y_true, y_prob = trainer.eval_step(dataloader=train_dataloader)
precisions, recalls, thresholds = precision_recall_curve(y_true.ravel(), y_prob.ravel())
plt.plot(thresholds, precisions[:-1], "r--", label="Precision")
plt.plot(thresholds, recalls[:-1], "b-", label="Recall")
plt.ylabel("Performance")
plt.xlabel("Threshold")
plt.legend(loc="best")
# + id="wM0_ZhZyQGib"
# Determining the best threshold
def find_best_threshold(y_true, y_prob):
"""Find the best threshold for maximum F1."""
precisions, recalls, thresholds = precision_recall_curve(y_true, y_prob)
f1s = (2 * precisions * recalls) / (precisions + recalls)
return thresholds[np.argmax(f1s)]
# + colab={"base_uri": "https://localhost:8080/"} id="yteVv5u8QLbG" outputId="ad4a0138-8087-4f58-ae8b-fd2ee12f953c"
# Best threshold for f1
threshold = find_best_threshold(y_true.ravel(), y_prob.ravel())
threshold
# + id="3L-N-IFjQNNP"
# Determine predictions using threshold
test_loss, y_true, y_prob = trainer.eval_step(dataloader=test_dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
# + colab={"base_uri": "https://localhost:8080/"} id="5RopxYJ4QO_a" outputId="9ebbe7bf-75a2-4173-c9d9-1f6ebb8c2cbf"
# Evaluate
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
print (json.dumps(performance, indent=2))
# + id="P6Po-3jdQSDx"
# Save artifacts
dir = Path("cnn")
dir.mkdir(parents=True, exist_ok=True)
tokenizer.save(fp=Path(dir, "tokenzier.json"))
label_encoder.save(fp=Path(dir, "label_encoder.json"))
torch.save(best_model.state_dict(), Path(dir, "model.pt"))
with open(Path(dir, "performance.json"), "w") as fp:
json.dump(performance, indent=2, sort_keys=False, fp=fp)
# + colab={"base_uri": "https://localhost:8080/"} id="3shMv1IeQVam" outputId="9a061178-d36c-4152-c3e2-61a77609524a"
# Load artifacts
device = torch.device("cpu")
tokenizer = Tokenizer.load(fp=Path(dir, "tokenzier.json"))
label_encoder = LabelEncoder.load(fp=Path(dir, "label_encoder.json"))
model = CNN(
embedding_dim=embedding_dim, vocab_size=vocab_size,
num_filters=num_filters, filter_sizes=filter_sizes,
hidden_dim=hidden_dim, dropout_p=dropout_p,
num_classes=num_classes)
model.load_state_dict(torch.load(Path(dir, "model.pt"), map_location=device))
model.to(device)
# + id="JNEQSkSJQWOo"
# Trainer module
trainer = Trainer(model=model, device=device)
# + id="PwZduA8OQWIH"
# Metrics
metrics = {"overall": {}, "class": {}}
# + id="ZbGbCnpXQWCH"
# Data to evaluate
device = torch.device("cuda")
loss_fn = nn.BCEWithLogitsLoss(weight=class_weights_tensor)
trainer = Trainer(model=model.to(device), device=device, loss_fn=loss_fn)
test_loss, y_true, y_prob = trainer.eval_step(dataloader=test_dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
# + colab={"base_uri": "https://localhost:8080/"} id="cr6wf3ktQmMR" outputId="7da1ba74-899b-45cd-b4a5-606ceb31674c"
# Overall metrics
overall_metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
metrics["overall"]["precision"] = overall_metrics[0]
metrics["overall"]["recall"] = overall_metrics[1]
metrics["overall"]["f1"] = overall_metrics[2]
metrics["overall"]["num_samples"] = np.float64(len(y_true))
print (json.dumps(metrics["overall"], indent=4))
# + id="S_wqNNe5QmJU"
# Per-class metrics
class_metrics = precision_recall_fscore_support(y_test, y_pred, average=None)
for i, _class in enumerate(label_encoder.classes):
metrics["class"][_class] = {
"precision": class_metrics[0][i],
"recall": class_metrics[1][i],
"f1": class_metrics[2][i],
"num_samples": np.float64(class_metrics[3][i]),
}
# + colab={"base_uri": "https://localhost:8080/"} id="x_RBismuQmGL" outputId="be06ca66-f8e4-4271-bbd6-9a23e898f099"
# Metrics for a specific class
tag = "First Party Collection/Use"
print (json.dumps(metrics["class"][tag], indent=2))
# + id="UUI2V8qXQmCz"
# Number of training samples per class
num_samples = np.sum(y_train, axis=0).tolist()
# + id="BWfh71PxQl_X"
# Number of samples vs. performance (per class)
f1s = [metrics["class"][_class]["f1"]*100. for _class in label_encoder.classes]
sorted_lists = sorted(zip(*[num_samples, f1s])) # sort
num_samples, f1s = list(zip(*sorted_lists))
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="glKOPPEGQl77" outputId="d59ffb16-2fe6-4468-e599-3a0c3150aa1c"
# Plot
n = 12 # num. top classes to label
fig, ax = plt.subplots()
ax.set_xlabel("# of training samples")
ax.set_ylabel("test performance (f1)")
fig.set_size_inches(25, 5)
ax.plot(num_samples, f1s, "bo-")
for x, y, label in zip(num_samples[-n:], f1s[-n:], label_encoder.classes[-n:]):
ax.annotate(label, xy=(x,y), xytext=(-5, 5), ha="right", textcoords="offset points")
# + id="0-lsy3XmQl4A"
# TP, FP, FN samples
index = label_encoder.class_to_index[tag]
tp, fp, fn = [], [], []
for i in range(len(y_test)):
true = y_test[i][index]
pred = y_pred[i][index]
if true and pred:
tp.append(i)
elif not true and pred:
fp.append(i)
elif true and not pred:
fn.append(i)
# + colab={"base_uri": "https://localhost:8080/"} id="ujnsQZ04Q-sw" outputId="54880db3-6623-4a81-f222-9575df42248d"
print (tp)
print (fp)
print (fn)
# + colab={"base_uri": "https://localhost:8080/"} id="IBpxH4UJRBi2" outputId="122c82a8-5143-4554-ae30-1119f2235e3e"
index = tp[0]
print (X_test_raw[index])
print (f"true: {label_encoder.decode([y_test[index]])[0]}")
print (f"pred: {label_encoder.decode([y_pred[index]])[0]}\n")
# + id="bW9tXCwlRBf_"
# Sorted tags
sorted_tags_by_f1 = OrderedDict(sorted(
metrics["class"].items(), key=lambda tag: tag[1]["f1"], reverse=True))
# + colab={"base_uri": "https://localhost:8080/", "height": 865, "referenced_widgets": ["fc152540571c45b798404dbef28995cf", "4ab95c37185a40e28c04a6237a8c0fc3", "3023f18232ed4782a93df58a3b21a60b", "87fa825fd3f94c33a28da6f8f93522bf", "251793285e414b2aa482ccfeee9c6ffb", "747be66413524fbbaea0b3511f9151ac", "87dcc8bddd3742649ef1f9fcfbd943a1"]} id="6p0CawtNRBdO" outputId="f4437bda-2f0c-40af-980e-2bc871987284"
@widgets.interact(tag=list(sorted_tags_by_f1.keys()))
def display_tag_analysis(tag="First Party Collection/Use"):
# Performance
print (json.dumps(metrics["class"][tag], indent=2))
# TP, FP, FN samples
index = label_encoder.class_to_index[tag]
tp, fp, fn = [], [], []
for i in range(len(y_test)):
true = y_test[i][index]
pred = y_pred[i][index]
if true and pred:
tp.append(i)
elif not true and pred:
fp.append(i)
elif true and not pred:
fn.append(i)
# Samples
num_samples = 3
cm = [(tp, "True positives"), (fp, "False positives"), (fn, "False negatives")]
for item in cm:
if len(item[0]):
print (f"\n=== {item[1]} ===")
for i in item[0][:num_samples]:
print (f" {X_test_raw[i]}")
print (f" true: {label_encoder.decode([y_test[i]])[0]}")
print (f" pred: {label_encoder.decode([y_pred[i]])[0]}\n")
# + id="1zKhDV1lRBaU"
from snorkel.slicing import PandasSFApplier
from snorkel.slicing import slice_dataframe
from snorkel.slicing import slicing_function
# + id="3c8l6Id1RBXR"
@slicing_function()
def pytorch_transformers(x):
"""Projects with the `pytorch` and `transformers` tags."""
return all(tag in x.category for tag in ["First Party Collection/Use", "Third Party Sharing/Collection"])
# + id="AMx0Y9ASRBUD"
@slicing_function()
def short_text(x):
"""Projects with short titles and descriptions."""
return len(x.segment_text.split()) < 7 # less than 7 words
# + colab={"base_uri": "https://localhost:8080/", "height": 221} id="MUAB-YnGRMuF" outputId="14dca799-2d53-4f2c-9451-61c8cff5c61a"
short_text_df = slice_dataframe(test_df, short_text)
short_text_df[["segment_text", "category"]].head()
# + colab={"base_uri": "https://localhost:8080/"} id="EZroZ7C9RMrT" outputId="fdcb5d34-02d9-472e-e8d0-a93ce58d43fa"
# Slices
slicing_functions = [pytorch_transformers, short_text]
applier = PandasSFApplier(slicing_functions)
slices = applier.apply(test_df)
slices
# + id="aoPxL6ZeRMpl"
# Score slices
metrics["slices"] = {}
for slice_name in slices.dtype.names:
mask = slices[slice_name].astype(bool)
if sum(mask):
slice_metrics = precision_recall_fscore_support(
y_test[mask], y_pred[mask], average="micro"
)
metrics["slices"][slice_name] = {}
metrics["slices"][slice_name]["precision"] = slice_metrics[0]
metrics["slices"][slice_name]["recall"] = slice_metrics[1]
metrics["slices"][slice_name]["f1"] = slice_metrics[2]
metrics["slices"][slice_name]["num_samples"] = len(y_true[mask])
# + colab={"base_uri": "https://localhost:8080/"} id="edn-xVovRMk7" outputId="d2b6616c-3c4f-4f8d-e0f2-b6d00211caa2"
print(json.dumps(metrics["slices"], indent=2))
# + id="9PvYyryqRMhv"
|
flask-ui/src/notebooks/DS5500_ResultsforP1Report.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import glob, matplotlib, os, math
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as pch
import matplotlib.backends.backend_pdf as pdf
import scipy.stats
mpl.rcParams['pdf.fonttype'] = 42
mpl.rcParams['ps.fonttype'] = 42
# %matplotlib inline
plt.style.use('./gcamp_figures/figures.mplstyle')
# +
'''
Plotting data from acclimation trials: read in data for each measurement.
'''
odor = '1-octen-3-ol_100um'
df = pd.read_csv('./individual_larva_calculations_GCAMP.csv')
df = df[df['dead']== False]
# DATA FOR ORCO ANIMALS
temp = df[(df['treatment_odor'] == 'milliQ_water_100uL') & (df['background'] == 'orco5_gcamp')]
water_orco_A_PI = temp['A_PI_(L-R)'].tolist()
water_orco_A_speed = temp['A_mean_speed_mm'].tolist()
temp = df[(df['treatment_odor'] == 'filtered_food_odor_100uL') & (df['background'] == 'orco5_gcamp')]
food_orco_A_PI = temp['A_PI_(L-R)'].tolist()
food_orco_A_speed = temp['A_mean_speed_mm'].tolist()
temp = df[(df['treatment_odor'] == odor) & (df['background'] == 'orco5_gcamp')]
odor_orco_A_PI = temp['A_PI_(L-R)'].tolist()
odor_orco_A_speed = temp['A_mean_speed_mm'].tolist()
# DATA FOR GR3 ANIMALS
temp = df[(df['treatment_odor'] == 'milliQ_water_100uL') & (df['background'] == 'gr3_gcamp')]
water_gr3_A_PI = temp['A_PI_(L-R)'].tolist()
water_gr3_A_speed = temp['A_mean_speed_mm'].tolist()
temp = df[(df['treatment_odor'] == 'filtered_food_odor_100uL') & (df['background'] == 'gr3_gcamp')]
food_gr3_A_PI = temp['A_PI_(L-R)'].tolist()
food_gr3_A_speed = temp['A_mean_speed_mm'].tolist()
temp = df[(df['treatment_odor'] == odor) & (df['background'] == 'gr3_gcamp')]
odor_gr3_A_PI = temp['A_PI_(L-R)'].tolist()
odor_gr3_A_speed = temp['A_mean_speed_mm'].tolist()
# DATA FOR GCAMP ANIMALS
temp = df[(df['treatment_odor'] == 'milliQ_water_100uL') & (df['background'] == 'gcamp')]
water_gcamp_A_PI = temp['A_PI_(L-R)'].tolist()
water_gcamp_A_speed = temp['A_mean_speed_mm'].tolist()
temp = df[(df['treatment_odor'] == 'filtered_food_odor_100uL') & (df['background'] == 'gcamp')]
food_gcamp_A_PI = temp['A_PI_(L-R)'].tolist()
food_gcamp_A_speed = temp['A_mean_speed_mm'].tolist()
temp = df[(df['treatment_odor'] == odor) & (df['background'] == 'gcamp')]
odor_gcamp_A_PI = temp['A_PI_(L-R)'].tolist()
odor_gcamp_A_speed = temp['A_mean_speed_mm'].tolist()
# DATA FOR WT ANIMALS
temp = df[(df['treatment_odor'] == 'milliQ_water_100uL') & (df['background'] == 'wt')]
water_wt_A_PI = temp['A_PI_(L-R)'].tolist()
water_wt_A_speed = temp['A_mean_speed_mm'].tolist()
# PRINT NS OF TREATMENTS
print('odor gcamp n=', len(odor_gcamp_A_PI))
print('odor orco n=', len(odor_orco_A_PI))
print('odor gr3 n=', len(odor_gr3_A_PI))
print('food gcamp n=', len(food_gcamp_A_PI))
print('food orco n=', len(food_orco_A_PI))
print('food gr3 n=', len(food_gr3_A_PI))
print('water gcamp n=', len(water_gcamp_A_PI))
print('water orco n=', len(water_orco_A_PI))
print('water gr3 n=', len(water_gr3_A_PI))
print('water wt n=', len(water_wt_A_PI))
# +
'''
Declare design variables that are the same for each plot
'''
labels = ['Water','Food','1-octen-3-ol'] * 3 # doubled for acclimate // experiment
labels = labels + ['Water'] # add WT animals treatment
color1 = '#c1c1c1' # acclimate
color2 = '#4286f4' # experiment
color3 = '#666666' # dark acclimate
color4 = '#1f5dc1' # dark experiment
w = 2 # line weight for mean line on plots
width = 0.5 # width of the data rectangles
pgap = 0.05 # space between data and significance notation (* / NS)
def add_square(sub, position, val, text=None, pgap=pgap):
''' Add rectangle to graph with mean and SE for the given dataset '''
mean = np.mean(val)
se = scipy.stats.sem(val, nan_policy='omit')
sub.add_patch(pch.Rectangle((position-width/2, mean-se), width, se*2, fc=color1, ec='none'))
sub.plot([position-width/2, position+width/2], [mean, mean], color=color3, lw=w)
if text is not None:
sub.text(s=text, x=position, y=mean+se+pgap, ha='center', va='center')
# +
fig= plt.figure(figsize=(20, 6))
sub1 = fig.add_subplot(121)
sub2 = fig.add_subplot(122)
sub1.set_ylabel('PI', color='k')
sub1.set_ylim(-1, 1)
sub2.set_ylabel('Mean speed (mm/s)', color='k')
sub2.set_ylim(0, 5.5)
# Add PI data for each treatment group, as well as the significance
add_square(sub1, 1, water_gcamp_A_PI)
add_square(sub1, 2, food_gcamp_A_PI)
add_square(sub1, 3, odor_gcamp_A_PI)
add_square(sub1, 4, water_gr3_A_PI)
add_square(sub1, 5, food_gr3_A_PI)
add_square(sub1, 6, odor_gr3_A_PI)
add_square(sub1, 7, water_orco_A_PI)
add_square(sub1, 8, food_orco_A_PI)
add_square(sub1, 9, odor_orco_A_PI)
add_square(sub1, 10, water_wt_A_PI)
# Add speed data for each treatment group, as well as the significance
add_square(sub2, 1, water_gcamp_A_speed)
add_square(sub2, 2, food_gcamp_A_speed)
add_square(sub2, 3, odor_gcamp_A_speed)
add_square(sub2, 4, water_gr3_A_speed)
add_square(sub2, 5, food_gr3_A_speed)
add_square(sub2, 6, odor_gr3_A_speed)
add_square(sub2, 7, water_orco_A_speed)
add_square(sub2, 8, food_orco_A_speed)
add_square(sub2, 9, odor_orco_A_speed)
add_square(sub2, 10, water_wt_A_speed)
for sub in [sub1, sub2]:
sub.spines['bottom'].set_position(('axes', -0.05))
sub.tick_params(axis='x', which='minor', bottom='off')
sub.set_xlim(0.5, 10.5)
sub.set_xticklabels(labels, rotation=90)
sub.set_xticks(np.arange(1, len(labels)+1, 1))
# add vertical lines and text eperating the genetic lines
ymin, ymax = sub.get_ylim()
for x in [3.5, 6.5, 9.5]:
sub.axvline(x=x, ymin=-1, ymax=1, lw=1, ls='solid', color=color3)
for sub, loc in zip([sub1, sub2], [1, 5.5]):
sub.text(2, loc, 'GCaMP6s\n+/+', ha='center', va='top', size='smaller', color='k')
sub.text(5, loc, 'GCaMP6s\nGr3-/-', ha='center', va='top', size='smaller', color='k')
sub.text(8, loc, 'GCaMP6s\norco5-/-', ha='center', va='top', size='smaller', color='k')
sub.text(10, loc, 'wt', ha='center', va='top', size='smaller', color='k')
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=None, hspace=0.2)
pp = pdf.PdfPages('./gcamp_figures/Supplemental_figure_5_PI_speed.pdf')
pp.savefig(fig)
pp.close()
plt.show()
plt.clf()
plt.close('all')
# -
|
Figure_Supplemental_6.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Librerías
from selenium import webdriver
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.by import By
import time
import pandas as pd
# +
# Opciones de navegación
options = webdriver.ChromeOptions()
options.add_argument('--start-maximized')
options.add_argument('--disable-extensions')
driver_path = 'C:/Users/MSI-NB/Documents/chromedriver.exe'
driver = webdriver.Chrome(driver_path, chrome_options=options)
# Iniciarla en la pantalla 2
driver.set_window_position(2000, 0)
driver.maximize_window()
time.sleep(10)
# Inicializamos el navegador
driver.get('https://eltiempo.es')
#driver.find_element(By.XPATH, '//*[@id="didomi-notice-agree-button"]').click()
# +
WebDriverWait(driver, 5)\
.until(EC.element_to_be_clickable((By.CSS_SELECTOR,
'button.didomi-components-button didomi-button didomi-dismiss-button didomi-components-button--color didomi-button-highlight highlight-button'.replace(' ', '.'))))\
.click()
WebDriverWait(driver, 5)\
.until(EC.element_to_be_clickable((By.CSS_SELECTOR,
'input#inputSearch')))\
.send_keys('Madrid')
WebDriverWait(driver, 5)\
.until(EC.element_to_be_clickable((By.CSS_SELECTOR,
'i.icon.icon-search')))\
.click()
WebDriverWait(driver, 5)\
.until(EC.element_to_be_clickable((By.CSS_SELECTOR,
'i.icon_weather_s.icon.icon-local')))\
.click()
WebDriverWait(driver, 5)\
.until(EC.element_to_be_clickable((By.XPATH,
'/html/body/div[7]/main/div[4]/div/section[4]/section/div/article/section/ul/li[2]/a')))\
.click()
WebDriverWait(driver, 5)\
.until(EC.element_to_be_clickable((By.XPATH,
'/html/body/div[7]/main/div[4]/div/section[4]/section/div[1]/ul')))
texto_columnas = driver.find_element_by_xpath('/html/body/div[7]/main/div[4]/div/section[4]/section/div[1]/ul')
texto_columnas = texto_columnas.text
tiempo_hoy = texto_columnas.split('Mañana')[0].split('\n')[1:-1]
horas = list()
temp = list()
v_viento = list()
:
horas.append(tiempo_hoy[i])
temp.append(tiempo_hoy[i+1])
v_viento.append(tiempo_hoy[i+2])
df = pd.DataFrame({'Horas': horas, 'Temperatura': temp, 'V_viento(km_h)':v_viento})
print(df)
df.to_csv('tiempo_hoy.csv', index=False)
driver.quit()
# -
horas
for i in range(0, len(tiempo_hoy), 4):
horas.append(tiempo_hoy[i])
#print(i)
temp.append(tiempo_hoy[i+1])
v_viento.append(tiempo_hoy[i+2])
grados_temperatura = list()
grados_temperatura.append(tiempo_hoy[1])
grados_temperatura
#grados_temperatura.append(tiempo_hoy[1])
grados_temperatura.append(tiempo_hoy[5])
grados_temperatura
for i in range(0, len(tiempo_hoy), 4):
#horas.append(tiempo_hoy[i])
#print(i)
temp.append(tiempo_hoy[i+1])
print(i+1)
#v_viento.append(tiempo_hoy[i+2])
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Timeseries of zonal means from NetCDFs
#
# This notebook uses climate-utils to calculate a timeseries of zonal means of a gridded dataset for example polygon in a coverage
#
# ## Requirements
#
# * `climate-utils`:
#
# `$ pip install https://github.com/flowmatters/climate-utils/archive/master.zip`
#
# * A version of `python-rasterstats` with the percent cover option and the ability to return the percent cover masks:
#
# `$ pip install https://github.com/joelrahman/python-rasterstats/archive/percent_cover.zip`
# +
import geopandas as gpd
import pandas as pd
from climate_utils import zonal
# %matplotlib inline
# -
# ## Load the vector coverage
#
# We need it as a Geopandas GeoDataFrame
#
# We also want to know which attribute in the coverage to include in the column name of the resulting time series Data Frame.
# +
coverage = gpd.read_file('LGA11aAust.shp')
# coverage needs to be in same projection as grid. If not, reproject the coverage
# coverage = coverage.to_crs(epsg=4326)
# -
coverage.plot()
coverage.columns
coverage.LGA_CODE11[:10]
# +
#coverage.bounds
# -
coverage.total_bounds
# ## Need to make sure the coverage fits within the bounds of the grids...
#
# Typically the gridded data sets will exclude a lot of smaller islands and territories that might be included in the polygon coverage.
import shapely
box = shapely.geometry.box(112,-44,154,-10) # The bounds of the grid. Could compute from the grids
box
coverage['geometry'] = coverage.geometry.intersection(box)
coverage = coverage[coverage.geometry.notnull()]
coverage.total_bounds
# # Define the storage conventions for the grids
#
# In this example, we have daily time series of gridded rainfall, stored in yearly files
#
# Once we know the pattern, we can define a loader - this will return the data for a given day.
# !ls *nc
# !ncdump -h 2016.daily_rain.nc
loader = zonal.netcdf_loader('${year}.${variable}.nc',known_bounds=coverage.total_bounds)
# # Define the time period of interest
#
# We need a pandas date index or similar
time_period = pd.date_range(pd.datetime(2000,1,1),pd.datetime(2002,12,31))
len(time_period)
time_period
# ## Run the aggregation
# %xmode Verbose
rainfall = zonal.compute_catchment_time_series('daily_rain',
coverage,
time_period,
loader,
name_attribute='LGA_CODE11',
column_naming='${catchment}',
percent_cover_scale=10)
# ## Explore results
#
# As:
#
# * Timeseries,
# * Statistics
# * Statistics joined back to spatial data
rainfall[:20]
rainfall.describe()
joined = coverage.join(rainfall.describe().transpose(),on='LGA_CODE11')
joined['mean_annual'] = joined['mean'] * 365
joined.plot('mean_annual',legend=True)
|
doc/examples/ZonalStatsNetCDF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Confinement continue, 10% port de masque
like_before = {0: {0: 693661, 1: 3008, 2: 17, 3: 279, 4: 217, 5: 24, 6: 153, 7: 381, 8: 1327},
1: {0: 692789, 1: 3640, 2: 34, 3: 261, 4: 216, 5: 24, 6: 153, 7: 541, 8: 1409},
2: {0: 691847, 1: 4084, 2: 79, 3: 252, 4: 216, 5: 23, 6: 153, 7: 841, 8: 1572},
3: {0: 690504,
1: 4745,
2: 140,
3: 263,
4: 216,
5: 22,
6: 153,
7: 1246,
8: 1778},
4: {0: 688614,
1: 5812,
2: 152,
3: 296,
4: 216,
5: 21,
6: 153,
7: 1674,
8: 2129},
5: {0: 686410,
1: 6966,
2: 162,
3: 363,
4: 216,
5: 20,
6: 153,
7: 2164,
8: 2613},
6: {0: 683631,
1: 8490,
2: 216,
3: 407,
4: 215,
5: 21,
6: 153,
7: 2683,
8: 3251},
7: {0: 680118,
1: 10484,
2: 264,
3: 476,
4: 215,
5: 20,
6: 153,
7: 3268,
8: 4069},
8: {0: 675713,
1: 12920,
2: 335,
3: 566,
4: 215,
5: 19,
6: 153,
7: 4061,
8: 5085},
9: {0: 670197,
1: 16079,
2: 389,
3: 661,
4: 214,
5: 18,
6: 153,
7: 5088,
8: 6268},
10: {0: 663687,
1: 19610,
2: 483,
3: 804,
4: 213,
5: 18,
6: 153,
7: 6277,
8: 7822},
11: {0: 656064,
1: 23562,
2: 656,
3: 932,
4: 213,
5: 18,
6: 153,
7: 7795,
8: 9674},
12: {0: 647302,
1: 27886,
2: 793,
3: 1133,
4: 213,
5: 18,
6: 153,
7: 9509,
8: 12060},
13: {0: 636744,
1: 33042,
2: 985,
3: 1387,
4: 212,
5: 18,
6: 153,
7: 11623,
8: 14903},
14: {0: 624749,
1: 38757,
2: 1142,
3: 1668,
4: 212,
5: 18,
6: 153,
7: 13946,
8: 18422},
15: {0: 610847,
1: 44850,
2: 1365,
3: 2056,
4: 212,
5: 18,
6: 153,
7: 16821,
8: 22745},
16: {0: 595571,
1: 51342,
2: 1656,
3: 2392,
4: 213,
5: 18,
6: 153,
7: 20034,
8: 27688},
17: {0: 578910,
1: 57803,
2: 1893,
3: 2911,
4: 212,
5: 18,
6: 153,
7: 23266,
8: 33901},
18: {0: 560200,
1: 64776,
2: 2214,
3: 3466,
4: 213,
5: 17,
6: 153,
7: 26967,
8: 41061},
19: {0: 539449,
1: 72412,
2: 2528,
3: 4105,
4: 213,
5: 16,
6: 153,
7: 30894,
8: 49297}}
# Déconfinement "hard"
deconfinement_hard = {0: {0: 694071, 1: 2598, 2: 35, 3: 279, 4: 217, 5: 25, 6: 153, 7: 362, 8: 1327},
1: {0: 693688, 1: 2817, 2: 45, 3: 261, 4: 216, 5: 26, 6: 153, 7: 452, 8: 1409},
2: {0: 693333, 1: 2813, 2: 38, 3: 270, 4: 216, 5: 25, 6: 153, 7: 658, 8: 1561},
3: {0: 693046, 1: 2637, 2: 71, 3: 274, 4: 216, 5: 24, 6: 153, 7: 933, 8: 1713},
4: {0: 692797,
1: 2449,
2: 91,
3: 270,
4: 216,
5: 23,
6: 153,
7: 1077,
8: 1991},
5: {0: 692533,
1: 2273,
2: 79,
3: 301,
4: 216,
5: 21,
6: 153,
7: 1147,
8: 2344},
6: {0: 692340,
1: 2121,
2: 72,
3: 325,
4: 215,
5: 21,
6: 153,
7: 1097,
8: 2723},
7: {0: 692151,
1: 1968,
2: 65,
3: 336,
4: 215,
5: 20,
6: 153,
7: 1053,
8: 3106},
8: {0: 691920,
1: 1891,
2: 62,
3: 347,
4: 215,
5: 19,
6: 153,
7: 1000,
8: 3460},
9: {0: 691751, 1: 1796, 2: 56, 3: 347, 4: 214, 5: 17, 6: 153, 7: 917, 8: 3816},
10: {0: 691555,
1: 1722,
2: 51,
3: 352,
4: 213,
5: 17,
6: 153,
7: 910,
8: 4094},
11: {0: 691395,
1: 1637,
2: 53,
3: 338,
4: 213,
5: 16,
6: 153,
7: 873,
8: 4389},
12: {0: 691225,
1: 1579,
2: 50,
3: 335,
4: 213,
5: 16,
6: 153,
7: 831,
8: 4665},
13: {0: 691030,
1: 1563,
2: 49,
3: 336,
4: 212,
5: 16,
6: 153,
7: 760,
8: 4948},
14: {0: 690895,
1: 1507,
2: 44,
3: 335,
4: 212,
5: 16,
6: 153,
7: 713,
8: 5192},
15: {0: 690723,
1: 1492,
2: 38,
3: 322,
4: 212,
5: 16,
6: 153,
7: 689,
8: 5422},
16: {0: 690568,
1: 1464,
2: 39,
3: 304,
4: 212,
5: 16,
6: 153,
7: 673,
8: 5638},
17: {0: 690391,
1: 1462,
2: 31,
3: 299,
4: 211,
5: 16,
6: 153,
7: 677,
8: 5827},
18: {0: 690209,
1: 1462,
2: 28,
3: 295,
4: 211,
5: 15,
6: 153,
7: 659,
8: 6035},
19: {0: 690014,
1: 1491,
2: 38,
3: 282,
4: 211,
5: 15,
6: 153,
7: 642,
8: 6221}}
deconfinement_soft = {0: {0: 694020, 1: 2649, 2: 19, 3: 279, 4: 217, 5: 25, 6: 153, 7: 378, 8: 1327},
1: {0: 693460, 1: 3030, 2: 25, 3: 261, 4: 216, 5: 25, 6: 153, 7: 488, 8: 1409},
2: {0: 692948, 1: 3159, 2: 52, 3: 254, 4: 216, 5: 24, 6: 153, 7: 688, 8: 1573},
3: {0: 692386,
1: 3196,
2: 94,
3: 254,
4: 216,
5: 23,
6: 153,
7: 1000,
8: 1745},
4: {0: 691691,
1: 3352,
2: 115,
3: 270,
4: 216,
5: 22,
6: 153,
7: 1217,
8: 2031},
5: {0: 690882,
1: 3576,
2: 126,
3: 307,
4: 216,
5: 20,
6: 153,
7: 1366,
8: 2421},
6: {0: 689964,
1: 3904,
2: 134,
3: 343,
4: 215,
5: 20,
6: 153,
7: 1472,
8: 2862},
7: {0: 688934,
1: 4268,
2: 142,
3: 389,
4: 215,
5: 19,
6: 153,
7: 1619,
8: 3328},
8: {0: 687728,
1: 4736,
2: 147,
3: 440,
4: 215,
5: 18,
6: 153,
7: 1775,
8: 3855},
9: {0: 686476,
1: 5209,
2: 138,
3: 486,
4: 214,
5: 16,
6: 154,
7: 1937,
8: 4437},
10: {0: 684949,
1: 5816,
2: 152,
3: 523,
4: 213,
5: 17,
6: 154,
7: 2195,
8: 5048},
11: {0: 683344,
1: 6327,
2: 196,
3: 536,
4: 213,
5: 17,
6: 154,
7: 2567,
8: 5713},
12: {0: 681628,
1: 6923,
2: 220,
3: 569,
4: 214,
5: 17,
6: 154,
7: 2861,
8: 6481},
13: {0: 679546,
1: 7742,
2: 228,
3: 620,
4: 213,
5: 17,
6: 154,
7: 3136,
8: 7411},
14: {0: 677450,
1: 8415,
2: 253,
3: 673,
4: 213,
5: 17,
6: 154,
7: 3501,
8: 8391},
15: {0: 675227,
1: 9083,
2: 286,
3: 714,
4: 213,
5: 17,
6: 154,
7: 3946,
8: 9427},
16: {0: 672648,
1: 9899,
2: 315,
3: 777,
4: 213,
5: 17,
6: 154,
7: 4399,
8: 10645},
17: {0: 669692,
1: 10964,
2: 359,
3: 855,
4: 212,
5: 17,
6: 154,
7: 4812,
8: 12002},
18: {0: 666536,
1: 12045,
2: 399,
3: 925,
4: 212,
5: 16,
6: 154,
7: 5273,
8: 13507},
19: {0: 663151,
1: 13122,
2: 454,
3: 1025,
4: 212,
5: 16,
6: 154,
7: 5801,
8: 15132}}
# +
from datetime import datetime, timedelta
day = datetime(2020, 5, 11)
days = [day + timedelta(i) for i in range(20)]
# -
scenarios = {'Mesures sanitaires renforcées': deconfinement_hard, 'Mesures sanitaires moyennes': deconfinement_soft, 'Aucune mesure sanitaire': like_before}
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 8))
colors = ['b-', 'r-', 'k-']
for i, scenario_name in enumerate(scenarios.keys()):
cases_day = [100 * (v[1] + v[2] + v[3] + v[4] + v[5]) for v in scenarios[scenario_name].values()]
plt.plot(days, cases_day, colors[i], label=scenario_name)
plt.grid(True, which="both")
plt.xlabel('Date')
plt.xticks(rotation=45)
plt.ylabel('Nombre de Cas Total')
plt.title("Évolution du Nombre Total de cas (y.c. asymptomatiques)\n Selon Différents scénarios de Déconfinement d'ici fin Juin")
plt.legend()
plt.plot()
# +
import pandas as pd
new_cases_df = pd.read_csv('../doc/new_overall.csv', sep=';')
new_cases_df = new_cases_df.groupby('jour')['incid_hosp'].agg('sum').reset_index()
new_cases_df = new_cases_df.rename(columns={'jour': 'day', 'incid_hosp': 'new_hosp'})
new_cases_df
# -
|
examples/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to numpy
# Numpy is a package that contains types and functions for mathematical calculations on arrays. The numpy library is vast and encapsulates a wide range of tools for linear algebra, Fourrier analysis, satistics and much more. The full manual for numpy can be found here:
# https://docs.scipy.org/doc/numpy/reference/
#
# In this tutorial, we will touch on the basics of numpy and see how numpy can be a convenient tool for artithmetic operations on arbitrarilly large arrays.
# In addition to this, we will also see a brief introduction to matplotlib, which contains tools to display diagrams or images that help illustrating the results of your calculations.
#
# Numpy is almost universally import using the alias 'np'. Given that codes using numpy will generally make frequent use of calls to numpy functions or objects, the loss of the 3 letters actually matters.
# Let's first import the package
import numpy as np
#Tadaaaa now we have all the power of the mighty numpy at our disposal.
#Let's use it responsibly
# ## Numpy's ndarray
#
# One of the reasons that makes numpy a great tool for computations on arrays is it ndarray calls. This class allows to declare arrays with a number of convenient methods and attributes that makes our life easier when programming complex algorithms on large arrays.
#Now let's see what one of its instances looks like:
a = np.ndarray(4)
b = np.ndarray([3,4])
print(type(b))
print('a: ', a)
print('b: ', b)
# There is a wide range of numpy functions that allow to declare ndarrays filled with your favourite flavours:
#
# https://docs.scipy.org/doc/numpy/reference/routines.array-creation.html
# zeros
z = np.zeros(5)
print(type(z))
print(z)
# ones
o = np.ones((4,2))
print(type(o))
print(o)
# ordered integers
oi = np.arange(10) #Only one-dimensional
print(type(oi))
print(oi)
# ### Operations on ndarrays
#
# Arithmetic operations on ndarrays are possible using python's symbols. It is important to notice that these operations are performed term by term on arrays of same size and dimensions. It is also possible to make operations between ndarrays and numbers, in which case, the same operation is performed on all the elements of the array. This is more generally true for operations on arrays where one array lacks one or several dimensions.
#An array of ones
x = np.arange(5)
#An array of random values drawn uniformly between 0 and 1
y = np.random.rand(5)
print('x: ', x)
print('y: ', y)
print('addition: ', x + y)
print('mutliplication: ', x * y)
print('power: ', x ** y)
#Operation with numbers
print('subtraction: ', x - 3)
print('fraction: ', x / 2)
print('power: ', x ** 0.5)
#Beware incompatible shapes: (play with the dimensions of y)
y = np.ones((6))
print('addition: ', x + y)
print('mutliplication: ', x * y)
print('power: ', x ** y)
# ndarrays and numpy also have methods or functions to perform matrix operations:
#Let's just declare some new arrays
x = (np.random.rand(4,5)*10).astype(int) # note, astype is a method that allows to change the type of all the elements in the ndarray
y = np.ones((5))+1
# Note: here, show addition of non-matching shapes
#np.ones((5,3,4))+np.random.randn(4)
#transpose
print('the array x: \n', x)
print('its transpose: \n', x.T)
#Matrix multiplication (play with the dimensions of y to see how this impact the results)
z1 = np.dot(x,y)
z2 = x.dot(y)
print(z1)
print(z2)
# ### array shapes
# It is possible to access the shape and size (there is a difference!) of an array, and even to alter its shape in various different ways.
print('Shape of x: ',x.shape) # From ndarray attributes
print('Shape of y: ',np.shape(y)) # From numpy function
print('Size of x: ', x.size) # From ndarray attributes
print('Size of y: ', np.size(y)) # From numpy function
# Now this is how we can change an array's size:
print('the original array: \n', x)
print('change of shape: \n', x.reshape((10,2)))#reshape 4x5 into 10x2
print('change of shape and number of dimensions: \n', x.reshape((5,2,2)))#reshape 4x5 into 5x2x2
print('the size has to be conserved: \n', x.reshape((10,2)).size)
# +
#flattenning an array:
xflat = x.flatten()
print('flattened array: \n {} \n with shape {}'.format(xflat, xflat.shape))
# -
# ### Indexing with numpy
#
# For the most part, indexing in numpy works exactly as we saw in python. We are going to use this section to introduce a couple of features for indexing (some native from python) that can significantly improve your coding skills. In particular, numpy introduces a particularly useful object: np.newaxis.
#conventional indexing
print(x)
print('first line of x: {}'.format(x[0,:]))
print('second column of x: {}'.format(x[:,1]))
print('last element of x: {}'.format(x[-1,-1]))
#selection
print('One element in 3 between the second and 13th element: ', xflat[1:14:3])
#This selection writes as array[begin:end:step]
#Equivalent to:
print('One element in 3 between the second and 13th element: ', xflat[slice(1,14,3)])
#Both notations are strictly equivalent, but slice allows to declare slices that can be used in different arrays:
sl1 = slice(1,3,1)
sl2 = slice(0,-1,2)
print('sliced array: ', x[sl1, sl2])
# Inverting the order in an array
print(xflat)
print(xflat[::-1])
#conditional indexing
print('all numbers greater that 3: ', x[x>3])
bool_array = (x == 8)
print('bool arrray is an array of booleans that can be used as indices: \n',bool_array)
print('all numbers greater that 3: ', x[bool_array])
#Ellipsis: select all across all missing dimensions
x_multi = np.arange(32).reshape(2,2,4,2)
print(x_multi)
print(x_multi[0,...,1])
print(x_multi[0,:,:,1])
# ### ndarray method for simple operations on array elements
#
# Here I list a small number of ndarray methods that are very convenient and often used in astronomy and image processing. It is always a good thing to have them in mind to simplfy your code. Of course, we only take a look at a few of them, but there is plenty more where it comes from.
a = np.linspace(1,6,3) # 3 values evenly spaced between 1 and 6
b = np.arange(16).reshape(4,4)
c = np.random.randn(3,4)*10 # random draws from a normal distribution with standard deviation 10
print(f'Here are 3 new arrays, a:\n {a}, \nb:\n {b}\nand c:\n {c}')
#Sum the elements of an array
print('Sum over all of the array\'s elements: ', a.sum())
print('Sum along the lines: ', b.sum(axis = 1))
print('Sum along the columns: ', b.sum(axis = 0))
#The axis option will be available for most numpy functions/methods
#Compute the mean and standard deviation:
print('mean of an array: ', b.mean())
print('std of an array: ', c.std())
#min and max of an array and teir positions
print('the minimum value of array b is {} and it is at position {}'.format(b.min(), b.argmin()))
print('the maximum value of array c is {} and it is at position {}'.format(c.max(), c.argmax()))
#sort an array's elements along one axis or return the indexes of the sorted array's element:
print('c', c)
argc = c.argsort()
print('The indexes that sort c and a sorted verison of c: \n \n {}\nand \n {} \n'.format(argc,c.sort()))
# Oups, not what we were expecting, but what happened is that c was replaced by its sorted version.
# This is an in-place computation.
print(c)
# +
#Your turn now: give me the ALL the elements of c sorted (not just along one axis).
#Your answer....
# +
#Then, sort the array in decreasing order
#Your answer....
# -
# Now, we are going to see an important feature in numpy. While one can live without nowing this trick, one cannot be a good python coder without using it. I am talking about the mighty:
# # Newaxis!!
# Newaxis allows to add a dimension to an array. This allows to expand arrays in a cheap way, which leads to faster operations on large arrays.
#
import numpy as np
#A couple of arrays first:
x_arr = np.arange(10)
y_arr = np.arange(10)
print(x_arr.shape)
x = x_arr[np.newaxis,:]
print(x.shape)
print(x_arr)
print(x)
print(x+x_arr)
#Now let's index these with newaxes:
print('Newaxis indexed array \n {} and its shape \n {}'.format(x_arr[:,np.newaxis],x_arr[:,np.newaxis].shape))
print('None leads to the same result: array \n {} and shape \n {}'.format(y_arr[None,:],y_arr[None,:].shape))
#Sum of elements
print('sum of the arrays:', (x_arr + y_arr))
#Sum of elements with newaxes
print('sum of the arrays: \n', (x_arr[None, :] + y_arr[:, None]))
#This is because we have been summing these arrays:
print(' ',x_arr[None, :])
print(y_arr[:, None])
# # A quick intro to matplotlib
#
# When wrinting complex algorithms, it is important to be able to chack that calculations are done properly, but also to be able to display results in a clear manner. When dimaensionality and size are small, it is still possible to rely on printing, but more generally and for better clarity, drawing graphs will come handy
import matplotlib.pyplot as plt
# %matplotlib inline
x = np.linspace(0,5,100)
#Plotting a curve
plt.plot(np.exp(x))
plt.show()
#The same curve with the right x-axis in red dashed line
plt.plot(x, np.exp(x), '--r')
plt.show()
#The same curve with the right x-axis and only the points in the data as dots
plt.plot(x[::4], np.exp(x[::4]), 'or')
plt.show()
|
day2/numpy-intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting Financial Data - Pandas Datareader
# ### Introduction:
#
# This time you will get data from a website.
#
#
# ### Step 1. Import the necessary libraries
# +
import numpy as np
import pandas as pd
# package to extract data from various Internet sources into a DataFrame
# make sure you have it installed
import pandas_datareader.data as web
# package for dates
import datetime as dt
# -
# ### Step 2. Create your time range (start and end variables). The start date should be 01/01/2015 and the end should today (whatever your today is).
# ### Step 3. Get an API key for one of the APIs that are supported by Pandas Datareader, preferably for AlphaVantage.
#
# If you do not have an API key for any of the supported APIs, it is easiest to get one for [AlphaVantage](https://www.alphavantage.co/support/#api-key). (Note that the API key is shown directly after the signup. You do *not* receive it via e-mail.)
#
# (For a full list of the APIs that are supported by Pandas Datareader, [see here](https://pydata.github.io/pandas-datareader/readers/index.html). As the APIs are provided by third parties, this list may change.)
# ### Step 4. Use Pandas Datarader to read the daily time series for the Apple stock (ticker symbol AAPL) between 01/01/2015 and today, assign it to df_apple and print it.
# ### Step 5. Add a new column "stock" to the dataframe and add the ticker symbol
# ### Step 6. Repeat the two previous steps for a few other stocks, always creating a new dataframe: Tesla, IBM and Microsoft. (Ticker symbols TSLA, IBM and MSFT.)
# ### Step 7. Combine the four separate dataFrames into one combined dataFrame df that holds the information for all four stocks
# ### Step 8. Shift the stock column into the index (making it a multi-level index consisting of the ticker symbol and the date).
# ### Step 7. Create a dataFrame called vol, with the volume values.
# ### Step 8. Aggregate the data of volume to weekly.
# Hint: Be careful to not sum data from the same week of 2015 and other years.
# ### Step 9. Find all the volume traded in the year of 2015
|
09_Time_Series/Getting_Financial_Data/Solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# # 5. 오차 역전파법
#
# 앞 장에선 기울기를 수치 미분을 사용해 구했다. (해석학적 미분이 안되니까.) 하지만 이는 시간이 많이 걸린다.
#
# 이번엔 가중치 매개변수 기울기를 효율적으로 계산하는 오차 역전파법 (Backpropagation)을 배워보자.
#
# 이를 수치, 그래프 두 가지 방법으로 배울 것이다.
#
# ## 5.1 계산 그래프 (Computational Graph)
#
# 그래프 자료구조. node와 edge로 구성됨.
#
# ### 5.1.1 계산 그래프로 풀다.
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile26.uf.tistory.com%2Fimage%2F997ED34B5B98F5F235B597">
#
# 노드는 연산자를 담당하는 것을 알 수 있다. 계산 그래프를 그리고 왼쪽 --> 오른쪽으로 진행한다. 이것을 '순전파'(forward propagation)라고 부른다.
#
# 반대 방향인 오른쪽 --> 왼쪽으로 가는 것이 '역전파' (backward propagation)이다.
#
# ### 5.1.2 국소적 계산
#
# 계산 그래프에선 각 노드의 국소적 계산이 합쳐져 전체의 계산이 이뤄진다.
#
# 즉, 각 노드는 자신과 관련된 계산만 신경쓰면 되고 그 이전의 복잡한 계산은 신경쓸 필요가 없다는 것이다.
#
# ### 5.1.3 왜 계산 그래프로 푸는가?
#
# 계산 그래프의 이점:
#
# 1. 국소적 계산. 복잡한 문제를 단순화 할 수 있음.
# 2. 중간 계산 결과를 모두 보관할 수 있음.
#
# 그보다 중요한 것은,
#
# 3. 역전파를 통해 미분을 효율적으로 계산할 수 있기 때문.
#
# 역전파를 통해 국소적 미분을 계속 뒤로 전달한다.
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile1.uf.tistory.com%2Fimage%2F997E914D5B98F628261EF4">
#
# 붉은 색은 dy/dx 이다. 즉, 매우 작은 dx가 증가할 때 y는 그의 1.1배 증가하는 것이다. (그래서 검은 색과 다름.)
#
# ## 5.2 연쇄법칙
#
# 역전파가 국소적 미분을 전달하는 원리는 chain rule 이다.
#
# ### 5.2.1 계산 그래프의 역전파
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile8.uf.tistory.com%2Fimage%2F999FD3425B98F63F1A9DDF">
#
# 신호 E에 노드의 국소적 미분 (ay/ax)를 곱하고 다음 노드로 전달하는 것.
#
# 왜 가능한지 연쇄법칙을 통해 살펴보자.
#
# ### 5.2.2 연쇄법칙이란?
#
# 합성함수는 연쇄법칙이다.
#
# ### 5.2.3 연쇄법칙과 계산 그래프.
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile27.uf.tistory.com%2Fimage%2F997387465B98F65D133C64">
#
# 이 두 과정이 같은 것이다. 즉, 연쇄법칙을 이용하여 한 단계씩 '국소적 미분'을 해 주는 것이다.
#
# ## 5.3 역전파
#
# ### 5.3.1 덧셈 노드의 역전파
#
# 덧셈 노드의 역전파는 상류의 값을 그대로 흘려보낸다.
#
# 상류에서 전달받은 aL/az 가 그대로 aL/az * 1이 된다. (왜냐하면 z = x+y 일 때 az/ax = 1 & az/ay = 1이 되므로.)
#
# ### 5.3.2 곱셈 노드의 역전파
#
# 곱셈 노드의 역전파는 상류의 값에 순전파 때의 입력 신호들을 서로 바꾼 값을 곱해서 하류로 보낸다.
#
# 그림으로 보면
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile1.uf.tistory.com%2Fimage%2F99E3EF435B98F69309175D">
#
# 상류에서 aL/az가 내려오면 원래 입력신호가 x였던 edge에는 (aL/az)*(az/ax)가 되어야 하니 z = xy 에서 d(xy)/d(x) = y이다.
#
# 따라서 (aL/az) * y 가 된다.
#
#
# 덧셈의 역전파에선 상류의 값을 그대로 보내므로 순방향 입력 신호가 필요하지 않았으나, 곱셉의 역전파에선 순방향 입력 신호의 값이 필요하다.
#
# 따라서 곱셈 노드를 구현할 때는 순전파의 입력 신호를 변수에 저장해둬야 한다.
#
# ### 5.3.3. 사과 쇼핑의 예
#
# 생략.
# ## 5.4 단순한 계층 구현하기.
#
# 이제 구현해보자.
#
# - 덧셈 노드: AddLayer
# - 곱셈 노드: MulLayer
#
# 계층을 클래스로 구현할 것이다. 계층이란 신경망의 기능 단위이다.
# ### 5.4.1 곱셈 계층.
#
# 모든 계층은 forward()와 backward() method를 가진다.
class MulLayer:
def __init__(self): # 초기화, 역전파 때 쓰기 위해 변수 유지.
self.x = None
self.y = None
def forward(self, x, y):
self.x = x
self.y = y
out = x * y
return out
def backward(self, dout):
dx = dout * self.y # x와 y를 서로 바꿔 곱해야 하니까.
dy = dout * self.x
return dx, dy
# +
# 이를 통해 순전파를 구현
apple = 100
apple_num = 2
tax = 1.1
# 계층들
mul_apple_layer = MulLayer() # 연산자 노드.
mul_tax_layer = MulLayer()
# 순전파
apple_price = mul_apple_layer.forward(apple, apple_num)
price = mul_tax_layer.forward(apple_price, tax)
price
# +
# 역전파 구현
dprice = 1
dapple_price, dtax = mul_tax_layer.backward(dprice)
dapple, dapple_num = mul_apple_layer.backward(dapple_price)
print(dapple, dapple_num, dtax)
# -
# ### 5.4.2 덧셈 계층
class AddLayer:
def __init__(self): # 변수 저장 안하니 초기화가 필요하지 않다.
pass
def forward(self, x, y):
out = x + y
return out
def backward(self, dout):
dx = dout * 1
dy = dout * 1
return dx, dy
# 덧셈노드는 곱셈노드보다 쉽다.
#
# 오렌지와 사과 계산 그래프 구현은 생략한다.
#
# ## 5.5 활성화 함수 계층 구현하기.
#
# 계산 그래프를 신경망에 적용하자.
#
# 우선 ReLU와 Sigmoid를 구현한다.
#
# ### 5.5.1 Relu 계층
#
# Relu 함수는 0보다 크면 x, 작으면 0이기 때문에 미분하면 1, 0 이다.
#
# 따라서 역전파시 순전파 때의 입력 신호가 0보다 컸으면 상류의 값을 그대로 하류로 흘리고 (E * 1), 0보다 작았으면 하류로 신호를 보내지 않음. (E * 0)
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile28.uf.tistory.com%2Fimage%2F99E517485B98F6E504DB20">
#
# .forward(), .backword()는 numpy 배열을 인수로 받는다.
#
# * ReLU는 전기 회로 스위치에 비유할 수 있다. 순전파 때 전기가 흐르고 있으면 스위치를 ON으로 하고 흐르지 않으면 OFF로 하기 때문이다.
class Relu():
def __init__(self):
self.mask = None
def forward(self, x):
self.mask = (x <= 0) # boolean 아니고 배열이다.
out = x.copy()
out[self.mask] = 0
return out
def backward(self, dout):
dout[self.mask] = 0 # 순전파 때 만들어놓은 mask를 사용.
dx = dout
return dx
# ### 5.5.2 Sigmoid 계층
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile27.uf.tistory.com%2Fimage%2F999E3B4B5B98F72021956C">
#
# / 와 exp() 노드가 보인다. 하지만 원리는 같다.
#
# 1. 우선 y=1/x 를 미분하면 = -(1/x^2) = -y^2 가 된다. 따라서 역전파에선 상류의 값에 원래 입력신호를 제곱하고 음수로 만들어준 값을 곱해 하류로 전달한다.
# 2. +1 하는 것은 + 노드니까 상류의 값을 그대로 흘린다.
# 3. y = exp()를 미분하면 그대로 = exp() = y 이다. 따라서 상류의 값에 그대로 입력신호를 곱해 하류로 보낸다.
# 4. * 노드는 순전파 값을 바꿔 곱한다.
#
# 이렇게 하면 Sigmoid 계층의 역전파 계산 그래프가 완성된다.
#
# 이렇게 계산하면 최종 출력은 aL/ay * (y^2 * exp(-x)) 가 된다. 따라서 Sigmoid 출력은 위의 복잡한 과정 없이 결과적으로 y^2 * exp(-x) 만 있으면 된다.
#
# 즉, Sigmoid의 역전파는 순전파의 출력만으로 계산 가능하다. (여기에 상류의 값을 곱하기만 하면 됨.)
#
# <img src="https://i.imgur.com/d0AT57Z.png">
#
# 이를 하나의 Sigmoid node로 만든다. 노드를 그룹화하여 간소화시키는 것이다. (위의 그림 참조)
#
# 이를 구현해보자.
class Sigmoid:
def __init__(self):
self.out = None
def forward(self, x):
out = 1 / (1 + np.exp(-x))
self.out = out
return out
def backward(self, dout):
dx = dout * self.out * (1.0 - self.out) # 순전파 때 저장해놓은 입력 신호를 역전파 때 사용함.
return dx
# ## Affine/Softmax 계층 구현하기.
#
# ### 5.6.1 Affine 게층
#
# Affine 계층이란? (신경망의 순전파 때 수행하는) 행렬의 곱을 기하학에선 Affine Transformation이라고 부름.
#
# 행렬 곱 node를 dot이라 하자. 이제 스칼라 값 대신 행렬이 흐른다.
#
# <img src="http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile27.uf.tistory.com%2Fimage%2F994002375B98F73E0590F4">
#
# 행렬에서 미분을 하면 그림의 1,2와 같은 식이 나온다.
#
# 형상에 주의하자. (행렬 곱을 위해)
#
# ### 5.6.2 배치용 Affine 계층
#
# X 하나만이 아닌, N개의 데이터를 묶어(batch) forward 하는 경우를 생각해보자.
# +
X_dot_W = np.array([[0,0,0], [10,10,10]])
B = np.array([1,2,3])
print(X_dot_W)
print(X_dot_W + B)
# +
dY = np.array([[1,2,3], [4,5,6]])
print(dY)
dB = np.sum(dY, axis=0)
print(dB)
# -
class Affine:
def __init__(self, W, b):
self.W = W
self.b = b
self.X = None
self.dW = None
self.db = None
def forward(self, x):
self.x = x
out = np.dot(x, self.W) + self.b
return out
def backward(self, dout):
dx = np.dot(dout, self.W.T)
self.dW = np.dot(self.x.T, dout)
self.db = np.sum(dout, axis=0)
# ### 5.6.3 Softmax-with-Loss 계층
#
# 출력층의 Softmax는 입력 값을 정규화하여 출력한다.
#
# 신경망에서는 크게 학습, 추론 두 가지를 하는데
#
# Softmax는 일반적으로 학습할 때 필요하고 추론할 땐 사용되지 않는다. (추론은 결과만 알면 되니까 굳이 확률로 정규화 해 줄 필요는 없다. 그냥 score 높은 것만 보면 되기 때문이다.)
#
# <img src='http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile9.uf.tistory.com%2Fimage%2F995A16395B98F76820FB40'>
#
# 여기에 Loss Function인 교차 엔트로피 오차도 포함한 Softmax-with-Loss를 구현할 수 있다.
#
# <img src='http://img1.daumcdn.net/thumb/R1920x0/?fname=http%3A%2F%2Fcfile29.uf.tistory.com%2Fimage%2F99EBF5395B98F7792B42CE'>
#
# 자세한 설명은 부록A에 나와있다.
#
# 그림을 다시 보면, 일단 이 그림은 3 클래스 분류를 가정한 것이다.
#
# - Softmax는 (a1, a2, a3)를 받아 정규화하여 (y1, y2, y3)를 출력했다.
# - Cross Entropy Error는 (y1, y2, y3)와 정답 레이블 (t1, t2, t3)을 받아 손실 L을 출력했다.
#
# 이 때, 굵은 화살표로 표시된 역전파의 결과를 보자. (y1 - t1, y2 - t2, y3 - t3) 와 같이 깔끔하게 나온다.
#
# (이는 Softmax에 일부러 Loss function으로 Cross Entropy Error을 썼기 때문임. 회귀의 경우 항등함수에 일부러 Loss function으로 MSE를 쓰면 똑같이 나옴.)
#
# 신경망 역전파에선 이 차이가(오차가) 앞 계층에 전해지는 것이다.
#
# 신경망의 학습 목적은 신경망 출력(Softmax 출력)이 정답 레이블과 가까워지도록 가중치 매개변수를 조정하는 것이기 때문에 오차를 잘 전달해야 한다.
#
# Softmax-with-Loss를 구현해보자.
def softmax(a):
exp_a = np.exp(a)
sum_exp_a = np.sum(exp_a)
y = exp_a / sum_exp_a
return y
def cross_entropy_error(y, t):
if y.ndim == 1:
t = t.reshape(1, t.size)
y = y.reshape(1, y.size)
batch_size = y.shape[0]
return -np.sum(np.log(np.arange(batch_size), t)) / batch_size
class SoftmaxWithLoss:
def __init__(self):
self.loss = None
self.y = None
self.t = None
def forward(self, x, t):
self.t = t
self.y = softmax(x)
self.loss = cross_entropy_error(self.y, self.t)
return self.loss
def backward(self, dout=1):
batch_size = self.t.shape[0]
dx = (self.y - self.t) / batch_size # batch_size로 나눠 데이터 1개당 오차를 앞 계층으로 전파.
return dx
# ## 5.7 오차 역전파법 구현하기.
#
# ### 5.7.1 신경망 학습의 전체 그림.
#
# 신경망에는 적응 가능한 가중치와 편향이 있고, 이 가중치와 편향을 훈련 데이터에 적응하도록 조정하는 과정을 '학습'이라 부른다. 이는 총 4단계로 이뤄진다.
#
# 1. 미니배치
#
# 훈련 데이터 중 일부를 무작위로 가져온다. 이 선별 데이터를 미니 배치라 하며, 이 미니배치의 손실 함수 값을 줄이는 것이 목표이다.
#
# 2. 기울기 산출
#
# 미니배치의 손실 함수 값을 줄이기 위해 각 가중치 매개변수의 기울기를 구한다. 기울기는 손실 함수의 값을 가장 작게하는 방향을 제시한다. (수치미분 --> 오차 역전파법)
#
# 3. 매개변수 갱신
#
# 가중치 매개변수를 기울기 방향으로 아주 조금 갱신한다.
#
# 4. 반복
#
# 1~3을 반복한다.
#
# ### 5.7.2 오차 역전파법을 적용한 신경망 구현하기.
#
# TwoLayerClass로 구현하겠다.
#
# TwoLayerClass의 인스턴스 변수
# - params: 딕셔너리 변수. 신경망의 매개변수를 보관. (eg: params['W1'])
# - layers: 순서 있는 딕셔너리 변수. 신경망의 계층을 보관. (eg: layers['Affine1'], layers['Relu1'])
# - lastLayer: 신경망의 마지막 계층. 여기선 SoftmaxWithLoss를 사용.
#
# TwoLayerClass의 method
# - init(): 초기화를 수행
# - input_size: 입력층 뉴런 수
# - hidden_size: 은닉층 뉴런 수
# - output_size: 출력층 뉴런 수
# - weight_init_std: 가중치 초기화 시 정규분포의 스케일
# - predict(self, x): 예측(추론) 수행. x는 이미지 데이터
# - loss(self, x, t): 손실함수. x는 이미지, t는 정답 레이블.
# - accuracy(self, x, t): 정확도 구함.
# - numerical_gradient(self, x, t): 가중치 매개변수의 기울기를 수치 미분 방식으로 구함.
# - gradient(self, x, t): 가중치 매개변수 기울기를 오차 역전파법으로 구함.
#
#
# +
import sys, os
sys.path.append(os.pardir)
from common.layers import *
from common.gradient import numerical_gradient
from collections import OrderedDict
import numpy as np
# -
class TwoLayerNet:
def __init__(self, input_size, hidden_size, output_size, weight_init_std=0.01): # 가중치 초기화.
self.params = {}
self.params['W1'] = weight_init_std * np.random.randn(input_size, hidden_size)
self.params['b1'] = np.zeros(hidden_size)
self.params['W2'] = weight_init_std * np.random.randn(hidden_size, output_size)
self.params['b2'] = np.zeros(output_size)
# 계층 생성
self.layers = OrderedDict()
self.layers['Affine1'] = Affine(self.params['W1'], self.params['b1'])
self.layers['Relu1'] = Relu()
self.layers['Affine2'] = Affine(self.params['W2'], self.params['b2'])
self.lastLayer = SoftmaxWithLoss()
def predict(self, x):
for layer in self.layers.values(): # layers의 원래 순서대로 forward.
x = layer.forward(x)
return x
def loss(self, x, t):
y = self.predict(x)
return self.lastLayer.forward(y,t)
def accuracy(self, x, t):
y = self.predict(x)
y = np.argmax(y, axis=1)
if t.ndim != 1: t = np.argmax(t, axis=1)
accuracy = np.sum(y==t) / float(x.shape[0])
return accuracy
def numerical_gradient(self, x, t):
loss_W = lambda W: self.loss(x,t)
grads = {}
grads['W1'] = numerical_gradient(loss_W, self.params['W1'])
grads['b1'] = numerical_gradient(loss_W, self.params['b1'])
grads['W2'] = numerical_gradient(loss_W, self.params['W2'])
grads['b2'] = numerical_gradient(loss_W, self.params['b2'])
return grads
def gradient(self, x, t):
# 순전파
self.loss(x, t)
# 역전파
dout = 1
dout = self.lastLayer.backward(dout)
layers = list(self.layers.values())
layers.reverse() # layers를 반대로 거슬러 올라감.
for layer in layers:
dout = layer.backward(dout)
grads = {}
grads['W1'] = self.layers['Affine1'].dW
grads['b1'] = self.layers['Affine1'].db
grads['W2'] = self.layers['Affine1'].dW
grads['b2'] = self.layers['Affine1'].db
return grads
# 순전파 때는 OrderedList인 layers를 그대로, 역전파 때는 거꾸로 reverse 해 하나씩 처리해주면 된다.
#
# 신경망 구성 요소를 계층으로 모듈화하여 구현했기 때문에 쉽게 구현할 수 있었다.
#
# 앞으로도 딥러닝을 할 때 계층을 더 쌓고 싶으면 그냥 이런 방식으로 붙이기만 하면 된다.
#
# ### 5.7.3 오차역전파법으로 구한 기울기 검증하기.
#
# 수치 미분과 해석적 방법을 배웠다. 오차역전파법은 후자에 속하기 때문에 매개변수가 많아도 빠르게 구할 수 있었다. (수치 미분은 느림.)
#
# 하지만 수치 미분은 오차역전파법이 제대로 구현되었는지 검증하는 절차에 필요하다.
#
# 오차역전파법은 빠르지만 복잡해 오류가 있기 십상이다. 수치 미분은 느리지만 쉽기 때문에 검증하기 좋다.
#
# 둘을 비교하는 것을 gradient check라고 부른다.
import sys, os
sys.path.append(os.pardir)
import numpy as np
from dataset.mnist import load_mnist
from two_layer_net import TwoLayerNet
# +
(x_train, t_train), (x_test, t_test) = load_mnist(normalize=True, one_hot_label=True)
network = TwoLayerNet(input_size=784, hidden_size=50, output_size=10)
x_batch = x_train[:3]
t_batch = t_train[:3]
grad_numerical = network.numerical_gradient(x_batch, t_batch)
grad_backprop = network.gradient(x_batch, t_batch)
# 각 가중치의 차이의 절대값을 구하고 그 절대값들의 평균을 낸다.
for key in grad_numerical.keys():
diff = np.average(np.abs(grad_backprop[key] - grad_numerical[key]))
print(key + ":" + str(diff))
# -
# 위의 오차를 보면 기울기 차이가 거의 없는 것을 볼 수 있다. 검증시 오차가 너무 크면 의심해봐야 한다.
#
# ### 5.7.4 오차역전파법을 사용한 학습 구현하기.
#
# 기울기를 오차역전파법으로 구한다는 점만 다르다.
# +
(x_train, t_train), (x_test, t_test) = load_mnist(normalize=True, one_hot_label=True)
network = TwoLayerNet(input_size=784, hidden_size=50, output_size=10)
iters_num = 10000
train_size = x_train.shape[0]
batch_size = 100
learning_rate = 0.1
train_loss_list = []
train_acc_list = []
test_acc_list = []
iter_per_epoch = max(train_size / batch_size, 1)
for i in range(iters_num):
batch_mask = np.random.choice(train_size, batch_size)
x_batch = x_train[batch_mask]
t_batch = t_train[batch_mask]
# 오차 역전파법으로 기울기를 구한다.
grad = network.gradient(x_batch, t_batch)
# 갱신
for key in ('W1', "b1", 'W2', 'b2'):
network.params[key] = network.params[key] - learning_rate * grad[key]
loss = network.loss(x_batch, t_batch)
train_loss_list.append(loss)
if i % iter_per_epoch == 0:
train_acc = network.accuracy(x_train, t_train)
test_acc = network.accuracy(x_test, t_test)
train_acc_list.append(train_acc)
test_acc_list.append(test_acc)
print(train_acc, test_acc)
# -
# ## 5.8 정리
#
# - 계산 그래프를 이용하면 계산 과정을 시각적으로 파악할 수 있다.
# - 계산 그래프의 노드는 국소적 계산으로 구성된다. 국소적 계산을 조합해 전체 계산을 구성한다.
# - 계산 그래프의 순전파는 통상의 계산을 수행한다. 한편, 계산 그래프의 역전파로는 각 노드의 미분을 구할 수 있다.
# - 신경망의 구성 요소를 계층으로 구현하여 기울기를 효율적으로 계산할 수 있다. (오차역전파법)
# - 수치 미분과 오차역전파법의 결과를 비교하면 오차역전파법의 구현에 잘못이 없는지 확인할 수 있다. (기울기 확인)
|
ch05/Ch.05.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
from tensorflow import keras
from datetime import datetime
import matplotlib.pyplot as plt
print(tf.__version__)
print(keras.__version__)
# +
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# -
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
train_images.shape
len(train_labels)
train_labels
test_images.shape
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
# +
tf.reset_default_graph()
logdir = "/tmp/fashion-mnist/" + datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = keras.callbacks.TensorBoard(log_dir=logdir)
train_images = train_images / 255.0
test_images = test_images / 255.0
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
# -
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_images,
train_labels,
batch_size=1000,
verbose=1,
epochs=100,
validation_data=(test_images, test_labels),
callbacks=[tensorboard_callback])
# +
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
# -
|
datasets/fashion-mnist-visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pyNN.nest as p
p.setup()
celltype = p.IF_curr_exp
p.SpikeSourcePoisson.default_parameters
p.list_standard_models()
spike_source = p.native_cell_type('poisson_generator')
input_pop=p.Population(1,spike_source,{'rate':200.0})
izh = p.native_cell_type('izhikevich')
izh.get_parameter_names
p1 = p.Population(10, celltype,{})
output = p.Population(10,izh)
connector=p.AllToAllConnector(weights=10, delays=2)
connection=p.Projection(input_pop,output,connector)
connection2=p.Projection(input_pop,p1,connector)
# +
output.record()
p1.record()
p1.record_v()
#output.record_v()
p.run(300)
spikes = output.getSpikes()
spikes_2= p1.getSpikes()
a=p1.get_v()
p.end()
# +
# Plot
import pylab
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
st = [spike[1] for spike in spikes] # spike times
sid = [spike[0] for spike in spikes]
plt.plot(st, sid, "|")
plt.axis([np.min(st), np.max(st), np.min(sid)-1, np.max(sid)+1])
plt.xlabel('Time (ms)')
plt.ylabel('Neuron ID')
plt.title('Spike Plot')
plt.xlim(xmin=0)
plt.show()
# +
st_2 = [spike[1] for spike in spikes_2]
sid_2 = [spike[0] for spike in spikes_2]
plt.plot(st_2, sid_2, "|")
plt.axis([np.min(st_2), np.max(st_2), np.min(sid_2)-1, np.max(sid_2)+1])
plt.xlabel('Time (ms)')
plt.ylabel('Neuron ID')
plt.title('Spike Plot')
plt.xlim(xmin=0)
plt.show()
# -
print np.max(st_2)
print np.min(st_2)
sh=np.shape(a)
print sh[0]
a[0:10][:]
# +
for i in range(sh[0]):
if a[i][0]==0:
vt_2 = [volt[1] for volt in a]
v_2 = [volt[2] for volt in a]
plt.plot(vt_2,v_2)
plt.axis([4,8,-70,-30])
plt.plot(a[:][0])
# -
|
src/experimental_code/.ipynb_checkpoints/pyNN_nest_test-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/agemagician/CodeTrans/blob/main/prediction/multitask/fine-tuning/function%20documentation%20generation/go/small_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="c9eStCoLX0pZ"
# **<h3>Predict the documentation for go code using codeTrans multitask finetuning model</h3>**
# <h4>You can make free prediction online through this
# <a href="https://huggingface.co/SEBIS/code_trans_t5_small_code_documentation_generation_go_multitask_finetune">Link</a></h4> (When using the prediction online, you need to parse and tokenize the code first.)
# + [markdown] id="6YPrvwDIHdBe"
# **1. Load necessry libraries including huggingface transformers**
# + colab={"base_uri": "https://localhost:8080/"} id="6FAVWAN1UOJ4" outputId="3fcc80d7-32cc-4403-92fe-92bcfec42b4b"
# !pip install -q transformers sentencepiece
# + id="53TAO7mmUOyI"
from transformers import AutoTokenizer, AutoModelWithLMHead, SummarizationPipeline
# + [markdown] id="xq9v-guFWXHy"
# **2. Load the token classification pipeline and load it into the GPU if avilabile**
# + colab={"base_uri": "https://localhost:8080/", "height": 316, "referenced_widgets": ["4ec43735448c4bb88a689ee81e17b74d", "ec1e83e6768d47558ac1684ef75436db", "fc3ed7a3d5b04b16b37438e7e5d5772d", "650d75e889294a3faee95af2eb504baa", "<KEY>", "70090f265aa34471b790a85d98848738", "85873d7519014775a1bd4163d7615e83", "<KEY>", "eabb8f1efa4d4dfb83acf64560f0613e", "b5f1c4ed836a4331ba383c058be395ce", "febc741e442f4ac9ac51d55e33a87a77", "<KEY>", "<KEY>", "44d5637453b04769b30d6f03a49838fe", "<KEY>", "a2643069b1f642d9a3d0079b7de538c1", "796d75cff8c346739dea27c48b4eedf9", "<KEY>", "<KEY>", "6538339476274cb6a90b13833266385e", "570488341fc8457499d765ac147eea4f", "<KEY>", "7a0d8e4396be4b1daee613be709754ef", "<KEY>", "8e6af5e1d83c4b0089a9eb305923218e", "b19bb34a4cc64e42a90888ee8a8bcede", "bae7f379694141cf8ee6576fef979616", "<KEY>", "<KEY>", "f96bb14ed4c54107bda0d0b609679054", "c2f2cfb5538746789ede96be1287d031", "41938baf2ed246c0806621ea0564543e", "af31ff34947b4c8497287f72fad5a2ad", "<KEY>", "768241accb034792ad1f8eb5ad98ca59", "580b9f4186a74fa1b83be2296e6c472a", "477ead1ad52d4e05a207c9e5f3fe8817", "b1a20ac386664329ba7f568b7b00cf19", "27910d2c77fd439db78f8227a08194dc", "80fe3f200da64cadbedc933390b334c3"]} id="5ybX8hZ3UcK2" outputId="3bb20c39-5bdf-4655-d7da-323cec78d1ec"
pipeline = SummarizationPipeline(
model=AutoModelWithLMHead.from_pretrained("SEBIS/code_trans_t5_small_code_documentation_generation_go_multitask_finetune"),
tokenizer=AutoTokenizer.from_pretrained("SEBIS/code_trans_t5_small_code_documentation_generation_go_multitask_finetune", skip_special_tokens=True),
device=0
)
# + [markdown] id="hkynwKIcEvHh"
# **3 Give the code for summarization, parse and tokenize it**
# + id="nld-UUmII-2e"
code = "func (pr *Progress) needSnapshotAbort() bool {\n\treturn pr.State == ProgressStateSnapshot && pr.Match >= pr.PendingSnapshot\n}" #@param {type:"raw"}
# + id="cJLeTZ0JtsB5" colab={"base_uri": "https://localhost:8080/"} outputId="58c8d53e-6cb2-4c4c-88e1-32be63660ce6"
# !pip install tree_sitter
# !git clone https://github.com/tree-sitter/tree-sitter-go
# + id="hqACvTcjtwYK"
from tree_sitter import Language, Parser
Language.build_library(
'build/my-languages.so',
['tree-sitter-go']
)
GO_LANGUAGE = Language('build/my-languages.so', 'go')
parser = Parser()
parser.set_language(GO_LANGUAGE)
# + id="LLCv2Yb8t_PP"
def get_string_from_code(node, lines):
line_start = node.start_point[0]
line_end = node.end_point[0]
char_start = node.start_point[1]
char_end = node.end_point[1]
if line_start != line_end:
code_list.append(' '.join([lines[line_start][char_start:]] + lines[line_start+1:line_end] + [lines[line_end][:char_end]]))
else:
code_list.append(lines[line_start][char_start:char_end])
def my_traverse(node, code_list):
lines = code.split('\n')
if node.child_count == 0:
get_string_from_code(node, lines)
elif node.type == 'string':
get_string_from_code(node, lines)
else:
for n in node.children:
my_traverse(n, code_list)
return ' '.join(code_list)
# + id="BhF9MWu1uCIS" colab={"base_uri": "https://localhost:8080/"} outputId="65d1a9ee-f714-42d7-8121-76aa9729af16"
tree = parser.parse(bytes(code, "utf8"))
code_list=[]
tokenized_code = my_traverse(tree.root_node, code_list)
print("Output after tokenization: " + tokenized_code)
# + [markdown] id="sVBz9jHNW1PI"
# **4. Make Prediction**
# + colab={"base_uri": "https://localhost:8080/"} id="KAItQ9U9UwqW" outputId="8b023772-6162-4df3-c8e3-7989f9c0993e"
pipeline([tokenized_code])
|
prediction/multitask/fine-tuning/function documentation generation/go/small_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# author=cxf
# date=2020-8-8
"""
file for setting the optimal cutoff for each sample according to the criterion
1.after filtering error rate less than 0.02 (1/5 of 0.1)
2.after filtering the number of genotyped sites is the largest and record the number
(check among points where the error rate less than 0.02)
"""
import numpy as np
i = '../../0.prepare_processing/run1/error.txt'
name = i.split('.')[0]
data4 = []
error = []
name_list=[]
with open(i, 'r') as fx3:
for line in fx3.readlines():
each_sample_name = np.array(line[0:-1].split(','))[0]
each_sample = np.array(line[0:-1].split(','))[1:].astype('float')
error.append(each_sample)
name_list.append(each_sample_name)
error = np.array(error)
i = '../../0.prepare_processing/run1/a90.txt'
name = i.split('.')[0]
sites = []
with open(i, 'r') as fx3:
for line in fx3.readlines():
each_sample_name = np.array(line[0:-1].split(','))[0]
each_sample = np.array(line[0:-1].split(','))[1:].astype('int32')
sites.append(each_sample)
sites = np.array(sites)
f = open(f"90_result.txt", "a+")
print('sample','precise', 'max_cutoff', 'max_num', 'error_rate',sep=',', file=f)
for i in range(error.shape[0]):
site_info = sites[i]
error_info = error[i]
site_info[error_info > 0.02] = 0
max_index = np.argmax(site_info)
if error_info[max_index] >= 0.02 or site_info[max_index]==0 :
error_info[error_info == 0] =1
error_rate=error_info[np.argmin(error_info)]
else:
error_rate=error_info[max_index]
print(name_list[i],90, max_index, np.max(site_info), error_rate, sep=',', file=f)
f.close()
# -
# take a look
# to tell you where each output comes from we leave a copy where it generates.
# you may see the max error rate exceeds the criterion which means the least error rate after 1-10 cutoff is here
# and we will discard this sample before following training.
# the cause of the high least error rate may be the contaimination or something else which brings in some errors
# can not be erased by UMI or removing low RNCU reads.
import pandas as pd
df=pd.read_csv('90_result.txt',index_col=0)
print(df)
print(df.describe())
|
model_training/1.cutoff_setting/run1/90%_cutoff_setting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MNIST
#
# Vous allez maintenant utiliser les fonctions haut niveau de pytorch: torch.nn, torch.optim etc.
#
# On commence par charger les données:
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# # %matplotlib notebook
from torchvision import datasets, transforms
mnist_folder = '/softwares/INFO/Module_ML/mnist'
# sur vos machines perso
# mnist_folder = '.'
train_data = datasets.MNIST(root=mnist_folder, download=True, transform=transforms.ToTensor(), train=True)
test_data = datasets.MNIST(root=mnist_folder, download=True, transform=transforms.ToTensor(), train=False)
# -
print(len(train_data))
print(len(test_data))
# +
# PLOT SOME RANDOM DIGITS
fig, axes = plt.subplots(nrows=2, ncols=5, figsize=(8, 4), squeeze=False)
for i in range(10):
r = i // 5
c = i % 5
idx = np.random.choice(len(train_data), 1)[0]
x = train_data[idx][0].numpy()
y = train_data[idx][1]
axes[r, c].imshow(x[0, :, :])
axes[r, c].set_title('y={}'.format(y))
plt.show()
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
from torch.utils.data import DataLoader
# Training settings
seed = 1337
lr = 0.01
epochs = 25
train_batch_size = 50
test_batch_size = 100
torch.manual_seed(seed)
train_loader = DataLoader(train_data, batch_size=train_batch_size, shuffle=True)
test_loader = DataLoader(test_data, batch_size=test_batch_size, shuffle=False)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(28*28, 10)
def forward(self, X):
X = X.view(-1, 28*28) # flatten images
X = self.fc1(X)
return F.log_softmax(X)
model = Net()
optimizer = optim.SGD(model.parameters(), lr=lr)
def train(epoch):
exp_loss = 0.
n_processed = 0
model.train()
for X, Y in train_loader:
X, Y = Variable(X), Variable(Y)
# Forward pass
Y_pred_prob = model(X)
# Compute expected loss
loss = F.nll_loss(Y_pred_prob, Y)
# Backward pass
loss.backward()
# Parameter update (gradient descent)
optimizer.step()
optimizer.zero_grad()
exp_loss.data[0] += loss * X.data.shape[0]
n_processed += X.data.shape[0]
exp_loss /= n_processed
return exp_loss
def test():
test_loss = 0
test_error = 0
model.eval()
for X, Y in test_loader:
X, Y = Variable(X, volatile=True), Variable(Y)
# Forward pass
Y_pred_prob = model(X)
# Compute the expected negative log-likelihood
test_loss += F.nll_loss(Y_pred_prob, Y, size_average=False).data[0]
# Get the mode of p(y|x) (most probable digit)
Y_pred = Y_pred_prob.data.max(1, keepdim=True)[1]
# Compute the expected 0/1 error
test_error += (1 - Y_pred.eq(Y.data.view_as(Y_pred))).sum()
test_loss /= len(test_loader.dataset)
test_error /= len(test_loader.dataset)
return test_loss, test_error
for epoch in range(1, epochs + 1):
print('Epoch {}...'.format(epoch))
train_loss = train(epoch)
print('Train negative log-likelihood: {:.6f}'.format(train_loss))
test_loss, test_error = test()
print('Test negative log-likelihood: {:.6f} 0/1 error: {:.6f}'.format(test_loss, test_error))
# -
# Jouez avec les paramètres suivants:
# - lr
# - epochs
# - train_batch_size
#
# Quelle est la meilleure performance (0/1 error) que vous arrivez à atteindre?
# ## Améliorez votre modèle
# Améliorez votre modèle afin de diminuer l'erreur sur le jeu de test. Essayez différentes architectures / hyperparamètres et à chaque fois reportez vos résultats.
#
# Objectif: passer sous les 1% d'erreur: http://yann.lecun.com/exdb/mnist/
#
# Pistes à essayer:
# - plus de couches cachées
# - convolutions + max pooling
# - dropout
# - couches résiduelles
#
# Astuces:
# - changer l'algorithme de descente de gradient (Adam)
# - implémentez une stratégie d'early stopping: $n$ epochs sans amélioration -> arret
# - implémentez une stratégie de diminution du learning rate: $n/2$ epochs sans amélioration -> $\alpha = \alpha / 2$ (torch.optim.lr_scheduler.ReduceLROnPlateau)
# Bonus: entrainez un modèle sur la base CIFAR-10 (torchvision.datasets.CIFAR10).
#
# Quelle performance (zero-one error) arrivez-vous à obtenir ?
|
mnist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="W_iPwfjsWulf"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 1*
#
# ---
# + [markdown] colab_type="text" id="kCE84iTRWulg"
# # Regression 1
#
# - Begin with baselines for regression
# - Use scikit-learn to fit a linear regression
# - Explain the coefficients from a linear regression
# + [markdown] colab_type="text" id="1XnAHO3mWulh"
# <NAME> wrote a good blog post, [“What questions can machine learning answer?”](https://brohrer.github.io/five_questions_data_science_answers.html)
#
# We’ll focus on two of these questions in Unit 2. These are both types of “supervised learning.”
#
# - “How Much / How Many?” (Regression)
# - “Is this A or B?” (Classification)
#
# This unit, you’ll build supervised learning models with “tabular data” (data in tables, like spreadsheets). Including, but not limited to:
#
# - Predict New York City real estate prices <-- **Today, we'll start this!**
# - Predict which water pumps in Tanzania need repairs
# - Choose your own labeled, tabular dataset, train a predictive model, and publish a blog post or web app with visualizations to explain your model!
# + [markdown] colab_type="text" id="cbhm8K-vWulh"
# ### Setup
#
# Run the code cell below. You can work locally (follow the [local setup instructions](https://lambdaschool.github.io/ds/unit2/local/)) or on Colab.
#
# Libraries:
#
# - ipywidgets
# - pandas
# - plotly
# - scikit-learn
# + colab={} colab_type="code" id="mnVbXjrVWuli"
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + [markdown] colab_type="text" id="0EQn2k_TWulm"
# # Begin with baselines for regression
# + [markdown] colab_type="text" id="fqSZDu2bWuln"
# ## Overview
# + [markdown] colab_type="text" id="ZraShmB3Wuln"
# ### Predict how much a NYC condo costs 🏠💸
#
# Regression models output continuous numbers, so we can use regression to answer questions like "How much?" or "How many?"
#
# Often, the question is "How much will this cost? How many dollars?"
# + [markdown] colab_type="text" id="VnFpSU8-6vRx"
# For example, here's a fun YouTube video, which we'll use as our scenario for this lesson:
#
# [Amateurs & Experts Guess How Much a NYC Condo With a Private Terrace Costs](https://www.youtube.com/watch?v=JQCctBOgH9I)
#
# > Real Estate Agent <NAME> just sold a pre-war condo in New York City's Tribeca neighborhood. We challenged three people - an apartment renter, an apartment owner and a real estate expert - to try to guess how much the apartment sold for. Leonard reveals more and more details to them as they refine their guesses.
# + [markdown] colab_type="text" id="XPOzYqT3Wulo"
# The condo from the video is **1,497 square feet**, built in 1852, and is in a desirable neighborhood. According to the real estate agent, _"Tribeca is known to be one of the most expensive ZIP codes in all of the United States of America."_
#
# How can we guess what this condo sold for? Let's look at 3 methods:
#
# 1. Heuristics
# 2. Descriptive Statistics
# 3. Predictive Model
# + [markdown] colab_type="text" id="uibXlVcwzVr1"
# ## Follow Along
# + [markdown] colab_type="text" id="5ezXj1joWulo"
# ### 1. Heuristics
#
# Heuristics are "rules of thumb" that people use to make decisions and judgments. The video participants discussed their heuristics:
#
#
#
# + [markdown] colab_type="text" id="cr13SNdB5kNP"
# **Participant 1**, Chinwe, is a real estate amateur. She rents her apartment in New York City. Her first guess was `8 million, and her final guess was 15 million.
#
# [She said](https://youtu.be/JQCctBOgH9I?t=465), _"People just go crazy for numbers like 1852. You say **'pre-war'** to anyone in New York City, they will literally sell a kidney. They will just give you their children."_
# + [markdown] colab_type="text" id="JqiU1UHm5gUt"
# **Participant 3**, Pam, is an expert. She runs a real estate blog. Her first guess was 1.55 million, and her final guess was 2.2 million.
#
# [She explained](https://youtu.be/JQCctBOgH9I?t=280) her first guess: _"I went with a number that I think is kind of the going rate in the location, and that's **a thousand bucks a square foot.**"_
# + [markdown] colab_type="text" id="8gRvzlBG5feH"
# **Participant 2**, Mubeen, is between the others in his expertise level. He owns his apartment in New York City. His first guess was 1.7 million, and his final guess was also 2.2 million.
# + [markdown] colab_type="text" id="CIK7pbEpWulp"
# ### 2. Descriptive Statistics
# + [markdown] colab_type="text" id="Jd3E27VhWulp"
# We can use data to try to do better than these heuristics. How much have other Tribeca condos sold for?
#
# Let's answer this question with a relevant dataset, containing most of the single residential unit, elevator apartment condos sold in Tribeca, from January through April 2019.
#
# We can get descriptive statistics for the dataset's `SALE_PRICE` column.
#
# How many condo sales are in this dataset? What was the average sale price? The median? Minimum? Maximum?
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="SV404taLWulq" outputId="3bf42b55-30ed-45f2-f985-db8956f0f56e"
import pandas as pd
df = pd.read_csv(DATA_PATH+'condos/tribeca.csv')
# -
pd.options.display.float_format = '{:,.0f}'.format
df['SALE_PRICE'].describe()
# + [markdown] colab_type="text" id="jcSdkoz-Wuls"
# On average, condos in Tribeca have sold for \$3.9 million. So that could be a reasonable first guess.
#
# In fact, here's the interesting thing: **we could use this one number as a "prediction", if we didn't have any data except for sales price...**
#
# Imagine we didn't have any any other information about condos, then what would you tell somebody? If you had some sales prices like this but you didn't have any of these other columns. If somebody asked you, "How much do you think a condo in Tribeca costs?"
#
# You could say, "Well, I've got 90 sales prices here, and I see that on average they cost \$3.9 million."
#
# So we do this all the time in the real world. We use descriptive statistics for prediction. And that's not wrong or bad, in fact **that's where you should start. This is called the _mean baseline_.**
# + [markdown] colab_type="text" id="1XWVQ9SpWult"
# **Baseline** is an overloaded term, with multiple meanings:
#
# 1. [**The score you'd get by guessing**](https://twitter.com/koehrsen_will/status/1088863527778111488)
# 2. [**Fast, first models that beat guessing**](https://blog.insightdatascience.com/always-start-with-a-stupid-model-no-exceptions-3a22314b9aaa)
# 3. **Complete, tuned "simpler" model** (Simpler mathematically, computationally. Or less work for you, the data scientist.)
# 4. **Minimum performance that "matters"** to go to production and benefit your employer and the people you serve.
# 5. **Human-level performance**
#
# Baseline type #1 is what we're doing now.
#
# (Linear models can be great for #2, 3, 4, and [sometimes even #5 too!](http://citeseerx.ist.psu.edu/viewdoc/summary?doi=10.1.1.188.5825))
# + [markdown] colab_type="text" id="PIvjiKDCWult"
# ---
#
# Let's go back to our mean baseline for Tribeca condos.
#
# If we just guessed that every Tribeca condo sold for \$3.9 million, how far off would we be, on average?
# -
guess = df['SALE_PRICE'].mean()
errors = guess - df['SALE_PRICE']
mean_absolute_error = errors.abs().mean()
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="yJm4xlvvWulu" outputId="ddbcc6f3-b411-4b81-876c-3546756d9dfd"
print(f'If we just guessed every Tribeca condo sold for ${guess:,.0f},')
print(f'we would be off by ${mean_absolute_error:,.0f} on average.')
# + [markdown] colab_type="text" id="8oqi26gSWulw"
# That sounds like a lot of error!
#
# But fortunately, we can do better than this first baseline — we can use more data. For example, the condo's size.
#
# Could sale price be **dependent** on square feet? To explore this relationship, let's make a scatterplot, using [Plotly Express](https://plot.ly/python/plotly-express/):
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="EsS1uT2aWulw" outputId="431442c8-0acb-4b80-fe6e-d4a8ba162c04"
import plotly.express as px
px.scatter(df, x='GROSS_SQUARE_FEET', y='SALE_PRICE')
# + [markdown] colab_type="text" id="dHg8dxXjWul0"
# ### 3. Predictive Model
#
# To go from a _descriptive_ [scatterplot](https://www.plotly.express/plotly_express/#plotly_express.scatter) to a _predictive_ regression, just add a _line of best fit:_
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="Nhb0q-liWul0" outputId="f66076ab-6e01-402a-d476-32be6efff844"
# + [markdown] colab_type="text" id="KnGbq5dGWul2"
# Roll over the Plotly regression line to see its equation and predictions for sale price, dependent on gross square feet.
#
# Linear Regression helps us **interpolate.** For example, in this dataset, there's a gap between 4016 sq ft and 4663 sq ft. There were no 4300 sq ft condos sold, but what price would you predict, using this line of best fit?
#
# Linear Regression also helps us **extrapolate.** For example, in this dataset, there were no 6000 sq ft condos sold, but what price would you predict?
# + [markdown] colab_type="text" id="DojBoEXPWul3"
# The line of best fit tries to summarize the relationship between our x variable and y variable in a way that enables us to use the equation for that line to make predictions.
#
#
#
#
# + [markdown] colab_type="text" id="VGZ4-j4l5FCS"
# **Synonyms for "y variable"**
#
# - **Dependent Variable**
# - Response Variable
# - Outcome Variable
# - Predicted Variable
# - Measured Variable
# - Explained Variable
# - **Label**
# - **Target**
# + [markdown] colab_type="text" id="DzZY24pE5HHP"
# **Synonyms for "x variable"**
#
# - **Independent Variable**
# - Explanatory Variable
# - Regressor
# - Covariate
# - Correlate
# - **Feature**
#
# + [markdown] colab_type="text" id="6KM8tTiK5NmF"
# The bolded terminology will be used most often by your instructors this unit.
# + [markdown] colab_type="text" id="XlsphHydU7mY"
# ## Challenge
#
# In your assignment, you will practice how to begin with baselines for regression, using a new dataset!
# + [markdown] colab_type="text" id="gcCoydrpWul3"
# # Use scikit-learn to fit a linear regression
# + [markdown] colab_type="text" id="bHDVblUnWul4"
# ## Overview
# + [markdown] colab_type="text" id="gAybFItMWul4"
# We can use visualization libraries to do simple linear regression ("simple" means there's only one independent variable).
#
# But during this unit, we'll usually use the scikit-learn library for predictive models, and we'll usually have multiple independent variables.
# + [markdown] colab_type="text" id="r1e85ZhUWul5"
# In [_Python Data Science Handbook,_ Chapter 5.2: Introducing Scikit-Learn](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html#Basics-of-the-API), <NAME> explains **how to structure your data** for scikit-learn:
#
# > The best way to think about data within Scikit-Learn is in terms of tables of data.
# >
# > 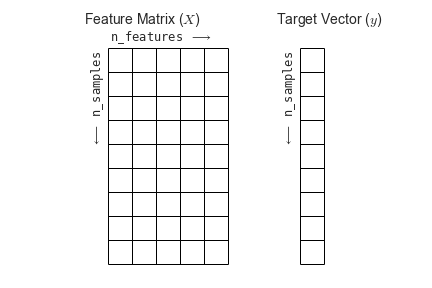
# >
# >The features matrix is often stored in a variable named `X`. The features matrix is assumed to be two-dimensional, with shape `[n_samples, n_features]`, and is most often contained in a NumPy array or a Pandas `DataFrame`.
# >
# >We also generally work with a label or target array, which by convention we will usually call `y`. The target array is usually one dimensional, with length `n_samples`, and is generally contained in a NumPy array or Pandas `Series`. The target array may have continuous numerical values, or discrete classes/labels.
# >
# >The target array is the quantity we want to _predict from the data:_ in statistical terms, it is the dependent variable.
# + [markdown] colab_type="text" id="gRsl-6mP5Uci"
# VanderPlas also lists a **5 step process** for scikit-learn's "Estimator API":
#
# > Every machine learning algorithm in Scikit-Learn is implemented via the Estimator API, which provides a consistent interface for a wide range of machine learning applications.
# >
# > Most commonly, the steps in using the Scikit-Learn estimator API are as follows:
# >
# > 1. Choose a class of model by importing the appropriate estimator class from Scikit-Learn.
# > 2. Choose model hyperparameters by instantiating this class with desired values.
# > 3. Arrange data into a features matrix and target vector following the discussion above.
# > 4. Fit the model to your data by calling the `fit()` method of the model instance.
# > 5. Apply the Model to new data: For supervised learning, often we predict labels for unknown data using the `predict()` method.
#
# Let's try it!
# + [markdown] colab_type="text" id="lFYuvV19Wul6"
# ## Follow Along
#
# Follow the 5 step process, and refer to [Scikit-Learn LinearRegression documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html).
# +
# 1. Import the appropriate estimator class from Scikit-Learn
# +
# 2. Instantiate this class
# +
# 3. Arrange X features matrix & y target vector
# +
# 4. Fit the model
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="-JKp8OL9Wul6" outputId="aba6b118-f18a-4e1a-958a-b16c5b624ad0"
# 5. Apply the model to new data
# + [markdown] colab_type="text" id="YNVE3zcaWul8"
# So, we used scikit-learn to fit a linear regression, and predicted the sales price for a 1,497 square foot Tribeca condo, like the one from the video.
#
# Now, what did that condo actually sell for? ___The final answer is revealed in [the video at 12:28](https://youtu.be/JQCctBOgH9I?t=748)!___
# + colab={} colab_type="code" id="FTJ_rUokWul8"
# + [markdown] colab_type="text" id="tQLfHWLIWul-"
# What was the error for our prediction, versus the video participants?
#
# Let's use [scikit-learn's mean absolute error function](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.mean_absolute_error.html).
# + colab={} colab_type="code" id="wr6md1nQWul_"
chinwe_final_guess = [15000000]
mubeen_final_guess = [2200000]
pam_final_guess = [2200000]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="fVPic_coWumA" outputId="1055f4cc-b6b7-40e9-930d-76e1b26f8147"
# + [markdown] colab_type="text" id="ZfoKMo8WWumI"
# This [diagram](https://ogrisel.github.io/scikit-learn.org/sklearn-tutorial/tutorial/text_analytics/general_concepts.html#supervised-learning-model-fit-x-y) shows what we just did! Don't worry about understanding it all now. But can you start to match some of these boxes/arrows to the corresponding lines of code from above?
#
# <img src="https://ogrisel.github.io/scikit-learn.org/sklearn-tutorial/_images/plot_ML_flow_chart_12.png" width="75%">
# -
# Here's [another diagram](https://livebook.manning.com/book/deep-learning-with-python/chapter-1/), which shows how machine learning is a "new programming paradigm":
#
# <img src="https://pbs.twimg.com/media/ECQDlFOWkAEJzlY.jpg" width="70%">
#
# > A machine learning system is "trained" rather than explicitly programmed. It is presented with many "examples" relevant to a task, and it finds statistical structure in these examples which eventually allows the system to come up with rules for automating the task. —[<NAME>](https://livebook.manning.com/book/deep-learning-with-python/chapter-1/)
# + [markdown] colab_type="text" id="jMw0m0XVSqNf"
# Wait, are we saying that *linear regression* could be considered a *machine learning algorithm*? Maybe it depends? What do you think? We'll discuss throughout this unit.
# + [markdown] colab_type="text" id="02Aw_2vSWumM"
# ## Challenge
#
# In your assignment, you will use scikit-learn for linear regression with one feature. For a stretch goal, you can do linear regression with two or more features.
# + [markdown] colab_type="text" id="-26MCaeBWumM"
# # Explain the coefficients from a linear regression
# + [markdown] colab_type="text" id="1YbqraQhWumN"
# ## Overview
#
# What pattern did the model "learn", about the relationship between square feet & price?
# + [markdown] colab_type="text" id="xkgmcd4XWumO"
# ## Follow Along
# + [markdown] colab_type="text" id="7QpkRN0PWumP"
# To help answer this question, we'll look at the `coef_` and `intercept_` attributes of the `LinearRegression` object. (Again, [here's the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html).)
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="DLvncqJ0WumQ" outputId="8e4eb40e-b350-4320-e90a-1e4605027b6d"
# + [markdown] colab_type="text" id="mT5jM1M3WumW"
# We can repeatedly apply the model to new/unknown data, and explain the coefficient:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="b2CEqLuxWumW" outputId="3a56631a-2e18-4f5a-bf4b-5847b49a9529"
def predict(square_feet):
y_pred = model.predict([[square_feet]])
estimate = y_pred[0]
coefficient = model.coef_[0]
result = f'${estimate:,.0f} estimated price for {square_feet:,.0f} square foot condo in Tribeca.'
explanation = f'In this linear regression, each additional square foot adds ${coefficient:,.0f}.'
return result + '\n' + explanation
predict(1497)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="nE8wq6UCWumY" outputId="ab60528c-1fae-438d-e6fb-80ccb5be5f45"
# What does the model predict for low square footage?
predict(500)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="haExj2STWumo" outputId="b521fa78-1478-4675-c9ea-72df2fcfd57b"
# For high square footage?
predict(10000)
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="L7pX1Gk-Wump" outputId="d362a8bb-55e7-4807-adb3-cb81d53011fb"
# + [markdown] colab_type="text" id="UdSfm-RvWumv"
# ## Challenge
#
# In your assignment, you will define a function to make new predictions and explain the model coefficient.
# + [markdown] colab_type="text" id="mVb4SJ0lWumv"
# # Review
# + [markdown] colab_type="text" id="omXPxXZbWumw"
# You'll practice these objectives when you do your assignment:
#
# - Begin with baselines for regression
# - Use scikit-learn to fit a linear regression
# - Make new predictions and explain coefficients
# + [markdown] colab_type="text" id="DHJv-pWNWumw"
# You'll use another New York City real estate dataset. You'll predict how much it costs to rent an apartment, instead of how much it costs to buy a condo.
#
# You've been provided with a separate notebook for your assignment, which has all the instructions and stretch goals. Good luck and have fun!
# + [markdown] colab_type="text" id="Vz8oqh_8Wumw"
# # Sources
#
# #### NYC Real Estate
# - Video: [Amateurs & Experts Guess How Much a NYC Condo With a Private Terrace Costs](https://www.youtube.com/watch?v=JQCctBOgH9I)
# - Data: [NYC OpenData: NYC Citywide Rolling Calendar Sales](https://data.cityofnewyork.us/dataset/NYC-Citywide-Rolling-Calendar-Sales/usep-8jbt)
# - Glossary: [NYC Department of Finance: Rolling Sales Data](https://www1.nyc.gov/site/finance/taxes/property-rolling-sales-data.page)
#
# #### Baselines
# - <NAME>, ["One of the most important steps in a machine learning project is establishing a common sense baseline..."](https://twitter.com/koehrsen_will/status/1088863527778111488)
# - <NAME>, [Always start with a stupid model, no exceptions](https://blog.insightdatascience.com/always-start-with-a-stupid-model-no-exceptions-3a22314b9aaa)
# - <NAME>, [The robust beauty of improper linear models in decision making](http://citeseerx.ist.psu.edu/viewdoc/summary?doi=10.1.1.188.5825)
#
# #### Plotly Express
# - [Plotly Express](https://plot.ly/python/plotly-express/) examples
# - [plotly_express.scatter](https://www.plotly.express/plotly_express/#plotly_express.scatter) docs
#
# #### Scikit-Learn
# - <NAME>, [Diagram](https://livebook.manning.com/book/deep-learning-with-python/chapter-1/)
# - <NAME>, [_Python Data Science Handbook,_ Chapter 5.2: Introducing Scikit-Learn](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html#Basics-of-the-API)
# - <NAME>, [Diagram](https://ogrisel.github.io/scikit-learn.org/sklearn-tutorial/tutorial/text_analytics/general_concepts.html#supervised-learning-model-fit-x-y)
# - [sklearn.linear_model.LinearRegression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html)
# - [sklearn.metrics.mean_absolute_error](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.mean_absolute_error.html)
|
module1-regression-1/LS_DS_211.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><center> <font color='black'> Business Data Analytics - Practice Session_11 </font></center></h1>
# <h2><center> <font color='black'> Fraud detection</font></center></h3>
# <h2><center> <font color='black'> University of Tartu - Spring 2020</font></center></h3>
# # Introduction
# <h3 ><center>"The need for fraud
# prevention services has never been greater."</center></h3>
#
# Among top 10 types of frauds today are:
#
# - **Wire fraud**: Using electronic and traditional communication means to obtain money based on false promises.
#
# - **Card fraud**: Using someone's else credit card or credit account to make an unauthorized purchase.
#
# - **Mortage fraud**: Providing false information that leads to a mortgage loan approval.
#
# - **Insurance fraud**: Someone knowingly lies to obtain a benefit or advantage to which they are not otherwise entitled.
#
#
#
#
# >The total value of fraudulent transactions annually
# amounting to **€1.8 Billion**. (European Central Bank)
# >
# >Technology advancments are serving as a tool for fraudlasters with some scams and frauds offered as Fraud as a Service (Faas) online.
#
# The industries affected by fraud vary from financial institutions, travel companies, online gaming, insurance etc. They apply preventive and detective solutions. Fraud detection solutions fall into 5 main categories:
#
# - **Expert-based rule engine**
# - **Descriptive analytics (unsupervised learning)**
# - **Predictive analytics (Supervised learning techniques)**
# - **Cluster Migration Analysis**
# - **Social network analysis**
#
# On this lab we will focus on detecting **Credit card fraud** using **Predictive analytics** tools.
# In general the workflow of financial fraud detection is as shown in the figure. There is a constant need to build, evaluate and re-train models. Other techniques like **online machine learning** can be used as well.
from IPython.display import display, Image
# +
display(Image(url= "https://miro.medium.com/max/1400/1*B452Vt6o-vCkzFbAewPubg.jpeg",unconfined=True))
# -
# **You can read an interesting Europaean fraud report(2019) by nets.eu in this [link](https://www.nets.eu/solutions/fraud-and-dispute-services/Documents/Nets-Fraud-Report-2019.pdf)
# # Credit fraud detection using classification
# + [markdown] colab_type="text" id="YpIQvrII72AF"
# ## 1. Dataset exploration
# https://www.kaggle.com/mlg-ulb/creditcardfraud
#
# #### Description
# The datasets contains transactions made by credit cards in September 2013 by european cardholders.
# This dataset presents transactions that occurred in two days, where we have 492 frauds out of 284,807 transactions. The dataset is highly unbalanced, the positive class (frauds) account for 0.172% of all transactions.
#
# It contains only numerical input variables which are the result of a PCA transformation. Unfortunately, due to confidentiality issues, we cannot provide the original features and more background information about the data. Features V1, V2, … V28 are the principal components obtained with PCA, the only features which have not been transformed with PCA are 'Time' and 'Amount'. Feature 'Time' contains the seconds elapsed between each transaction and the first transaction in the dataset. The feature 'Amount' is the transaction Amount, this feature can be used for example-dependant cost-senstive learning. Feature 'Class' is the response variable and it takes value 1 in case of fraud and 0 otherwise.
# + colab={} colab_type="code" id="TzVoE-mC71_8"
import time
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import seaborn as sns
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA, TruncatedSVD
# + colab={} colab_type="code" id="EWN-GggQ72AG"
df = pd.read_csv('creditcard.csv')
# + colab={} colab_type="code" id="V1uF5tZB72AN" outputId="8a3ada8b-3d22-400a-da3d-d514220b8bf4"
df.head()
# + colab={} colab_type="code" id="X69GTkLT72AW" outputId="7caeb2b8-3e22-4c06-975b-82ae6e2421dd"
# Remove any infinity or NaN values
print('df length before removing nan/inf: ', len(df))
df = df[~df.isin([np.nan, np.inf, -np.inf]).any(1)] #exclude the values specified in `isin()`
print('df length after removing nan/inf: ', len(df))
# + colab={} colab_type="code" id="TqP64hdN72Ae" outputId="df2d3527-bd46-45db-d95b-959f27fbfee4"
print('Not Fraud:', round(df['Class'].value_counts()[0]/len(df) * 100, 2), '% of the dataset')
print('Fraud:', round(df['Class'].value_counts()[1]/len(df) * 100, 2), '% of the dataset')
# -
# ### Distribution of Amount and Time
# + colab={} colab_type="code" id="KVZgcgPJ72Ak" outputId="0179ec1e-705c-4c30-ac17-4a2a0e843608"
fig, ax = plt.subplots(1, 2, figsize=(18,4))
amount_val = df['Amount'].values
time_val = df['Time'].values
sns.distplot(amount_val, ax=ax[0], color='r')
ax[0].set_title('Distribution of Transaction Amount', fontsize=14)
ax[0].set_xlim([min(amount_val), max(amount_val)])
ax[0].set_xlabel('Amount (Euro)')
ax[0].set_xlabel('Probability')
sns.distplot(time_val, ax=ax[1], color='b')
ax[1].set_title('Distribution of Transaction Time', fontsize=14)
ax[1].set_xlim([min(time_val), max(time_val)])
ax[1].set_xlabel('Time (s)')
ax[1].set_xlabel('Probability')
plt.show()
# -
# ### Correlation
# + colab={} colab_type="code" id="VD3pe0kZ72BR" outputId="7343a09c-8594-4b10-c775-d36c9bfb7999"
fig, ax = plt.subplots(figsize=(24,10))
sns.set(font_scale=3)
# Entire DataFrame
corr = df.corr()
sns.heatmap(corr, cmap='coolwarm_r', annot_kws={'size':30}, ax=ax)
ax.set_title("Imbalanced Correlation Matrix", fontsize=30)
plt.show()
# -
# ## 2. Data preprocessing
# ### Feature scaling
# + colab={} colab_type="code" id="K9bOv1JH72Ar"
# Scale features 'amount' and 'time' to match scale of other features in dataset
from sklearn.preprocessing import StandardScaler, RobustScaler
# RobustScaler is less prone to outliers.
rob_scaler = RobustScaler()
df['scaled_amount'] = rob_scaler.fit_transform(df['Amount'].values.reshape(-1,1))
df['scaled_time'] = rob_scaler.fit_transform(df['Time'].values.reshape(-1,1))
df.drop(['Time','Amount'], axis=1, inplace=True)
# + colab={} colab_type="code" id="TLNkVao772Az" outputId="007baf1a-355b-4d53-83b5-71a6fde90a6f"
# This cell simply moves the scaled_amount and scaled_time to be the first 2 columns
scaled_amount = df['scaled_amount']
scaled_time = df['scaled_time']
df.drop(['scaled_amount', 'scaled_time'], axis=1, inplace=True)
df.insert(0, 'scaled_amount', scaled_amount)
df.insert(1, 'scaled_time', scaled_time)
# Amount and Time are Scaled!
df.head()
# + [markdown] colab_type="text" id="WUMzc7UZ72A5"
# ### Subsampling to avoid:
# * Overfitting
# * Wrong correlation
# -
# Lets shuffle the data before creating the subsamples
df = df.sample(frac=1, random_state=888)
df.head()
# + colab={} colab_type="code" id="KmAAH5tU72A6"
# amount of fraud classes 492 rows.
fraud_df = df.loc[df['Class'] == 1]
non_fraud_df = df.loc[df['Class'] == 0][:len(fraud_df)]
normal_distributed_df = pd.concat([fraud_df, non_fraud_df])
# Shuffle dataframe rows
new_df = normal_distributed_df.sample(frac=1, random_state=42)
# + colab={} colab_type="code" id="pKVXmRbZ72BH" outputId="64fbb989-7989-482c-e2a2-055fe9743152"
print('Not Fraud:', round(new_df['Class'].value_counts()[0]/len(new_df) * 100, 2), '% of the dataset')
print('Fraud:', round(new_df['Class'].value_counts()[1]/len(new_df) * 100, 2), '% of the dataset')
# -
new_df.shape
# +
# Make sure we use the subsample in our correlation
f, ax = plt.subplots( figsize=(24,10))
sub_sample_corr = new_df.corr()
sns.heatmap(sub_sample_corr, cmap='coolwarm_r', annot_kws={'size':20}, ax=ax)
ax.set_title('SubSample Correlation Matrix \n (use for reference)', fontsize=30)
plt.show()
# -
# ### Principal component analysis
# + colab={} colab_type="code" id="Syksuadf72Bd" outputId="317f4bca-0cb2-4165-cddb-06f8ccd17004"
# New_df is from the random undersample data (fewer instances)
X = new_df.drop('Class', axis=1)
y = new_df['Class']
# PCA Implementation
t0 = time.time()
X_reduced_pca = PCA(n_components=2, random_state=42, svd_solver='full').fit_transform(X.values)
t1 = time.time()
print("PCA took {:.2} s".format(t1 - t0))
# + colab={} colab_type="code" id="zispjfP672Bl" outputId="6666b6f1-42fc-4eb9-e224-688fe400659c"
# PCA scatter plot
#f, (ax1) = plt.subplots(1, 1, figsize=(24,6))
# labels = ['No Fraud', 'Fraud']\
plt.rcParams["figure.figsize"] = (24,10)
blue_patch = mpatches.Patch(color='#0A0AFF', label='No Fraud')
red_patch = mpatches.Patch(color='#AF0000', label='Fraud')
plt.scatter(X_reduced_pca[:,0], X_reduced_pca[:,1], c=(y == 0), cmap='coolwarm', label='No Fraud', linewidths=2)
plt.scatter(X_reduced_pca[:,0], X_reduced_pca[:,1], c=(y == 1), cmap='coolwarm', label='Fraud', linewidths=2)
plt.grid(True)
plt.legend(handles=[blue_patch, red_patch])
plt.xlabel('First component')
plt.ylabel('Second component')
plt.show()
# -
# ## 3. Using classification for fraud detection
# There are several algorithms used to detect fraud among them genetic algorithms, neural networks, random forest etc. In today's practice session we are going to use Software Vector Machine algorithm.
# + colab={} colab_type="code" id="RhreLKgl72Bt"
# Our data is already scaled we should split our training and test sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# + colab={} colab_type="code" id="HU7hVgch72B2" outputId="6f66ee55-7ceb-4a6e-f629-79ceaaf3e1f1"
from sklearn.model_selection import cross_val_score, cross_val_predict
from sklearn.svm import SVC
import warnings
warnings.filterwarnings("ignore")
svc = SVC()
svc.fit(X_train, y_train)
training_score = cross_val_score(svc, X_train, y_train, cv=5)
print('Training score: ', round(training_score.mean(), 2)*100, "% accuracy score")
# -
# ### Hyperparameter tuning
# During the practice sessions until now, for our machine learning models, we have used default parameters or some predefined values. In practical applications, the job of a data scientist includes actually trying different hyperparameter combinations and evaluating which one of them gives the best results. This process is called hyperparameter optimization or hyperparameter tuning. This is a crucial step ass the performance of most of the algorithms is highly dependent on the hyperparameter choice. The process usually includes two strategies:
#
# - **Exploitation**
# - **Exploration**
#
# In python, the scikit-learn module offers two functions which automates the hyperparameter search process: GridSearchCV and RandomizedSearchCV
# + colab={} colab_type="code" id="dBJD1K1H72B9" outputId="6293650a-e4c8-4654-c71e-9ebf1dbf5ebf"
# Use GridSearchCV to find the best parameters.
from sklearn.model_selection import GridSearchCV
# Support Vector Classifier
svc_params = {'C': [0.5, 0.7, 0.9, 1], 'kernel': ['rbf', 'poly', 'sigmoid', 'linear']}
grid_svc = GridSearchCV(SVC(), svc_params)
grid_svc.fit(X_train, y_train)
# SVC best estimator
svc = grid_svc.best_estimator_
print(grid_svc.best_params_)
# + colab={} colab_type="code" id="VE-3-SRV72CH"
updated_training_score = cross_val_score(svc, X_train, y_train, cv=5,)
print('Training score: ', round(updated_training_score.mean(), 2)*100, "% accuracy score")
# -
# ### Model evaluation
## By default the score property calculates accuracy score
acc = svc.score(X_test, y_test)
print('Accuracy score in test data', acc)
svc_pred = svc.predict(X_test)
# +
from sklearn.metrics import classification_report
print(classification_report(y_test, svc_pred))
# + colab={} colab_type="code" id="RZ_TKfGq72CP" outputId="45182e2e-a7e5-4e83-ff42-b13043547673"
from sklearn.metrics import roc_auc_score, roc_curve
print('Support Vector Classifier: ', roc_auc_score(y_test, svc_pred))
# + colab={} colab_type="code" id="oOrDXJYl72CT" outputId="046fe5ea-71dc-4d78-d3a8-558a73b448c0"
# Plot the ROC curve
svc_fpr, svc_tpr, svc_threshold = roc_curve(y_test, svc_pred)
plt.figure(figsize=(16,8))
plt.rcParams.update({'font.size': 18})
plt.title('ROC Curve', fontsize=18)
plt.plot(svc_fpr, svc_tpr, label='Support Vector Classifier Score: {:.4f}'.format(roc_auc_score(y_test, svc_pred)))
plt.plot([0, 1], [0, 1], 'k--')
plt.axis([-0.01, 1, 0, 1])
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.annotate('Minimum ROC Score of 50% \n (This is the minimum score to get)', xy=(0.5, 0.5), xytext=(0.6, 0.3),
arrowprops=dict(facecolor='#6E726D', shrink=0.005),
)
plt.legend(prop={'size': 18})
plt.show()
# + [markdown] colab_type="text" id="ZMRvneoS72Cc"
# ### References
# * https://www.kaggle.com/mlg-ulb/creditcardfraud
# * https://www.kaggle.com/janiobachmann/credit-fraud-dealing-with-imbalanced-datasets
# * https://imbalanced-learn.readthedocs.io/en/stable/api.html
# * https://scikit-learn.org/stable/user_guide.html
|
Lab11_enlik.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python for Psychologists - Session 2
# ## Homework assignment
# **Exercise 0.**
# Use the dictionary below to complete the sentence, i.e., fill "....." with something meaningful. Hint: think about the ```types()``` you want to combine in the sentence and why it might (not) work
cute_animals = {"otter": 1, "dogs": 2}
"Dogs are one of my most favourite animals, acutally they are my "+ str(cute_animals["dogs"]) +"th most favourite animals."
# **Exercise 1**.
#
# Below you can find a dictionary representing a small data base of people. Add up the ages of all three people using indexing.
people = {"<NAME>":{"age": 68, "gender":"female", "marital_status":"married"}, "<NAME>":{"age": 46, "gender":"male", "marital_status":"divorced"}, "U<NAME>ich":{"age": 38, "gender":"female", "marital_status":"single"}}
people["<NAME>"]["age"] + people["J<NAME>"]["age"] + people["Ulla Ulrich"]["age"]
# **Exercise 3**.
#
# Try to change the third position in the list of the tuple below.
my_tuple = (1,["I", "am", "a", "list", "inside", "a", "tuple"])
my_tuple[1][2] = "lalala"
my_tuple
# **Exercise 4.**
#
# Try to execute the code below. In a markdown cell, explain why there is an error.
some_dict = {my_tuple: 3}
# Explanation: In principle tuples can be keys in dictionaries, however our tuple here contains a list, which makes it changeable (mutable) !
# **Exercise 5**.
#
# Check if the element "inside" is part of the tuple my_tuple. Afterwards, check if the element `1` is part of my_tuple.
"inside" in my_tuple
1 in my_tuple
# **Exercise 6**.
#
# Check if the item ("I am a key", "I am its value") is part of the dictionary below. If you have no clue, go back to the session script, and if that doesn't help: google it! :)
my_dict = {"blablabla": "I am a value", "I am a key": "blublublu"}
("I am a key", "I am its value") in my_dict.items()
# ### Exercise 7: please read and try to cause each error at least once on purpose :)
#
#
# ### Errors
# ... are a good thing! (well, sometimes...) Usually, they tell you relatively precisely what is going wrong. Python is not a mean machinery that tries to annoy you by throwing errors at random, but it actually gives you a hint which part of your code needs some changes.
#
# Let's get to know some errors you might encounter:
# - **SyntaxError:** Also called "parsing error". This error occurs while Python is trying to decifer your code. Python, like any other language has a certain syntax, which we need to stick to in order for Python to understand your code. Whenever we violate this syntactic structure we get a SyntaxError.
# - **NameError:** NameErrors occur when we try to access the value of a variable/an object that we haven't even defined yet. It's like talking to your family about some fellow student of yours that they don't know. "Yesterday, I went binge-drinking with Tonya." Of course, their response will be "Who is Tonya?". Well, Python will ask you the same question when referring to an object that you have never introduced.
# - **IndexError:** We can cause an IndexError when trying to access an element in an subscriptable object (basically objects that are containers which allow for indexing at all) that does not exist. For example, we would cause an IndexError by asking Python to return the value of the 6th element of a list that is only 3 elements long.
# - **TypeError:** TypeErrors occur whenever we try to do some operation with a data type that just doesn't fit the operation, like trying to do something with two strings that can only be done with two integers or floats.
# - **KeyError:** These errors usually occur, when we try to access the value of a key, although the key does not exist in the dictionary.
# - **ZeroDivisionError:** This kind of error is more than intuitive. It is raised simply when we try to divide a number by 0, which is mathematically not defined.
#
# Nothing is more fun than causing errors on purpose!
[x for x in liste
x
liste = [1,2,3,4,5,6]
liste[55]
"barbara" **2
cute_animals["cat"]
5/0
|
session2/Session2_Assignment_solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importing libraries
import pandas as pd
def recommendation():
# Read csv file into a pandas dataframe
df = pd.read_csv("books.csv")
df = df[df.language_code =='eng']
df['Title'] = df['title'].str.split('(').str[0]
print("Enter rating w/wo <rating> or random for book suggestion")
search= input("enter to search ")
if search=="random":
df_new= df.sample(replace=True)[['Title', 'authors',"rating"]]
print("Title: ", df_new[['Title']].to_string(index=False, header=False))
print("Author: ", df_new[['authors']].to_string(index=False, header=False))
print("Rating: ", df_new[['rating']].to_string(index=False, header=False))
return "Enjoy!"
else "rating" in search:
search=search.split()
rate=float(search[1])
df_rate = df[df['rating'].astype(float) >= rate]
df_new= df_rate.sample(replace=True)[['Title', 'authors',"rating"]]
print("Title: ", df_new[['Title']].to_string(index=False, header=False))
print("Author: ", df_new[['authors']].to_string(index=False, header=False))
print("Rating: ", df_new[['rating']].to_string(index=False, header=False))
return "Enjoy!"
print(recommendation())
|
book_rec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Use python 3.7+
import numpy as np
import pandas as pd
from pathlib import Path
import PIL
from PIL import Image, ExifTags
import torch
from torchvision.transforms import ToTensor
import shutil
from typing import Optional, List
from dataclasses import dataclass
import folium
@dataclass
class Lat_Lon:
lat: float
lon: float
# +
# A callable class that creates the destination bin directories
# based on start, stop and step temperature arguments and can then
# # copy a file from a source directory to a destination bin directory.
class Bin_Agent:
def _create_bin_names(self):
"""Make the bin directory names."""
bin_cnt = 2 + (self.stop - self.start) // self.step
self.bin_names = [""] * bin_cnt
self.bin_names[0] = "le_{}".format(self.start)
self.bin_names[1:] = ["gt_{}".format(i)
for i in range(self.start, self.stop + 1, self.step)]
def _create_destination_bin_directories(self):
"""Create the directories under base_path if they do not exist."""
for dir_name in self.bin_names:
new_dir = self.base_path / dir_name
new_dir.mkdir(exist_ok=True)
print("Directory {} created or already exists.".format(new_dir))
def _get_destination_bin(self, max_temp : float) -> str:
"""Given a maximum temperature, find the destination bin"""
bin_name = None
if max_temp > self.stop:
bin_name = self.bin_names[-1]
elif max_temp <= self.start:
bin_name = self.bin_names[0]
else:
bin_idx = int(max_temp) // self.step + 1
bin_name = self.bin_names[bin_idx]
return bin_name
def __init__(self, base_path : Path, start : int, stop : int, step : int):
assert step > 0, "Step must be greater than 0"
assert start < stop, "Stop must be greater than Start"
assert (stop - start) % step == 0, "Difference between start and stop must be an exact multiple of step"
self.base_path = base_path
self.start = start
self.stop = stop
self.step = step
self.bin_names = []
self._create_bin_names()
self._create_destination_bin_directories()
def __call__(self, tif_file : Path, max_temp : float) -> Path:
"""Given a tif file and its maximum temperature, find the destination path"""
tif_file_name = tif_file.name
bin_name = self._get_destination_bin(max_temp)
return self.base_path / bin_name / tif_file_name
# -
temper_path = Path("rjpeg_temperature_16bit_0-80")
geo_path = Path("rjpeg_modified_0-80_gps_mod")
base_path = Path(".")
temper_path, geo_path, base_path
# Create the bin agent with our input parameters.
bin_agent = Bin_Agent(Path("."),0,80,20)
# +
# Test bin_agent()
#bin_agent(temper_path/"DJI_0035_0.tif", 25.9999)
# -
def get_tif_file_paths(src_path : Path) -> pd.DataFrame:
"""Get the *.tif filenames"""
tif_file_paths = sorted(src_path.glob("*.tif"))
print("Number of files = {}".format(len(tif_file_paths)))
return pd.DataFrame({"src" : tif_file_paths})
df = get_tif_file_paths(temper_path); df
# +
def get_max_temperature_from_tif_file(tif_file : Path) -> float:
"""Read a temperature tif file and get the maximum temperature"""
max_temp = np.NAN
try:
# Open the tif image file using PIL and then convert it to a torch.tensor().
with PIL.Image.open(tif_file) as tif_img:
tensor_tif = ToTensor()(tif_img)
# Get the max() value of the tensor (returns a single item tensor like tensor(3.))
# The item() call gets the scaler item from the tensor (e.g. 3. from tensor(3.)).
max_temp = tensor_tif.max().item()
except:
max_temp = np.NAN
return max_temp
def get_jpg_path(tif_file : Path) -> Path:
"""Create the corresponding jpg filename from the tif filename"""
jpg_file_suffix = ".jpg"
# Stem gives the filename without the parent path or the suffix
# e.g. Path("/hello/world.tif") => "world"
file_name_stem = tif_file.stem
jpg_file = geo_path / (file_name_stem + jpg_file_suffix)
return jpg_file
# -
# Find the max temperature (in °C)
df["max_temp"] = df["src"].map(lambda fp: get_max_temperature_from_tif_file(fp)); df
# Check if any tif file is missing a max_temp.
df_fail = df[df["max_temp"].isna()] ; df_fail
# Find the appropriate destination directory, based on max temperature
df["dest_path"] = df.apply(lambda r: bin_agent(r["src"], r["max_temp"]), axis="columns"); df
# Check the number of tif files that should be copied to each bin.
dfg = df["dest_path"].copy()
dfg["dest_bin"] = dfg.map(lambda p: p.parts[-2])
dfg.groupby(["dest_bin"]).count()
# Copy the tif files from source to destination.
df.apply(lambda r: shutil.copy2(src=r["src"], dst=r["dest_path"]), axis="columns")
# Filter rows which have a max_temp greater than 60°C.
# Make a copy of filtered rows (otherwise the slice refers to the original dataframe)
df60 = df[df.max_temp > 60].copy(); df60
# reset the index to 0
df60.index = pd.RangeIndex(len(df60.index)); df60
# Compute the jpg filepath from the source tif filepath.
df60["jpg"] = df60["src"].map(lambda fp: get_jpg_path(fp)); df60
def get_latitude_longitude(file_path : Path) -> Optional[Lat_Lon]:
"""Given a jpg file, return the latitude and longitude from the exif data"""
def get_decimal_degrees_from_dms(dms, ref : str) -> float:
""" Convert the rational64u format of degree, minute, second to decimal_degrees.
Sample input dms format: ((51, 1), (45, 1), (1358029999, 50000000))
"""
(dn, dd), (mn, md), (sn, sd) = dms
d, m, s = float(dn)/dd, float(mn)/md, float(sn)/sd
decimal_dms = d + m/60.0 + s/(3600.0)
# If Latitude is 'S' or Longitude is 'W' , the degree sign should be negative
return -decimal_dms if ref in ["S", "W"] else decimal_dms
# Get the GPSInfo exif data
try:
image = PIL.Image.open(file_path)
gps_exif = {PIL.ExifTags.TAGS[k]: v for k, v in image._getexif().items()
if k in PIL.ExifTags.TAGS and PIL.ExifTags.TAGS[k] == "GPSInfo"}
if len(gps_exif) == 0:
return None
except:
return None
# Add the GPS tags.
gps_exif_with_tags = {PIL.ExifTags.GPSTAGS[k]: v for k, v in gps_exif["GPSInfo"].items()
if k in PIL.ExifTags.GPSTAGS}
# Verify presence of the required GPS tags.
if not frozenset(["GPSLatitudeRef", "GPSLatitude", "GPSLongitudeRef",
"GPSLongitude"]).issubset(gps_exif_with_tags.keys()):
return None
result = None
# Convert the rational64u format of degree, minute, second to decimal_degrees
try:
assert gps_exif_with_tags["GPSLatitudeRef"] in ["N", "S"], "LatitudeRef must be 'N' or 'S'"
lat = get_decimal_degrees_from_dms(gps_exif_with_tags["GPSLatitude"],
gps_exif_with_tags["GPSLatitudeRef"])
assert gps_exif_with_tags["GPSLongitudeRef"] in ["E", "W"], "LongitudeRef must be 'E' or 'W'"
lon = get_decimal_degrees_from_dms(gps_exif_with_tags["GPSLongitude"],
gps_exif_with_tags["GPSLongitudeRef"])
except:
return None
return Lat_Lon(round(lat, 6), round(lon, 6))
# Get the latitude and longitude from the exif data of the jpg files
df60["coord"] = df60["jpg"].map(lambda fp: get_latitude_longitude(fp)); df60
# Check if any coordinate is missing.
df_fail = df60[df60["coord"].isna()]; df_fail
# +
# Create the map
tmap = folium.Map(location=[51.755, -100.896], zoom_start=12)
for ir in df60.iterrows():
row = ir[1]
temp = round(row["max_temp"], 2)
lat = row["coord"].lat
lon = row["coord"].lon
folium.Marker(location=[lat, lon],
popup="Lat:{},Long:{},Temp={}°C".format(lat, lon, temp),
icon=folium.Icon(color="red", icon="info-sign")).add_to(tmap)
tmap.save("hotspots60C_map.html")
display(tmap)
# -
|
process_image.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# # Univariate Probability
# In the example above, we demonstrated some code that generates fake data $X$ and $Y$. On the other hand, real data comes from the real world, not from some python code. For every dataset, there is an immensely complex network of causal interactions that ultimately "produces" the data.
#
# For example, in our blood pressure example, a patient's pre-treatment vital signs are caused by their physiological state: their genetics, life history, what they ate for breakfast that morning, whether or not they just ran up a flight of stairs, and so on and so forth. Taking a drug influences the levels of certain chemicals in the blood, which are taken up at particular rates in certain organs by certain enzymes, the levels of which are impacted by the patient's genetics and prior physiological state, which was influenced by their life history, etc. Thus the impact of the drug on cellular processes is mediated by these factors. The cells respond by increasing or decreasing their production of some proteins or metabolites, which, in combination with the immediate condition of the patient when the measurement is taken, determines the post-treatment blood pressure.
#
# Or, let's say we're trying to determine whether or not there is a cat in a photograph. The cat being in front of the camera when the photo was taken ($y_i$) could be caused by a huge number of factors, and the values of the pixels in the photograph ($x_i$) are caused by the reflection of photons emitted from sources of light off the cat (and other objects) and the mechanics of the detection of light inside the camera.
#
# In a nutshell, the world is complicated. There is no way that mere mortals could ever write code accurate enough to perfectly simulate the exact processes that produce data about complex real-world phenomena.
#
# But, despite the complexity, you should start thinking about that complex web of causality as "code" that's being run in some cosmic simulation. Maybe you can imagine that there are "data gods" that write and are running this code. We'll never see their code, and we'll never be able to understand it, but somewhere, out there, that metaphysical code is running, and it's generating the observations that we see in our data.
# You can think of that code as a little "factory" that pumps out observations of $x_i$ and $y_i$, one at a time. The factory is behind a curtain that we can't ever look behind, but we can see the pile of $x_i$s and $y_i$s that come out of it, which are our $X$ and $Y$.
# 
# If we had that code, we'd be able to reverse engineer it to find the most likely value of $y_i$ given $x_i$ as accurately as would be possible with those predictors. In practice, however, we can only build a *model* of that code. Our model will never capture the complexities of reality, the same way that a model plane doesn't even begin to approach the complexity of a real aircraft. But, ideally, it will be similar enough in ways that are important for the task at hand: if we're using a model plane just to demonstrate what an aircraft might look like, we don't need the model to have functioning jet engines. And if all we need to do is estimate $y_i$ for a new $x_i$, we don't exactly need to understand the complex web of causality linking the two together.
# We do, however, need a way to talk about the relationship that $x_i$ and $y_i$ might have. And to do that, we need a way to talk abstractly about the "code" or "data factory" that's behind the curtain, the same way we developed abstract terms to describe our data. Thankfully, the language of probability works perfectly for that.
# ## Random variables are factories that generate data
# The data factories we're interested in are the kind that output $x_i$s and $y_i$s, but to understand how these factories work it's better to consider a simpler factory that produces one number at a time, instead of one vector $x_i$ and one number $y_i$.
#
# We'll call our factory $\mathbf Z$. This factory pushes out one value $z_i$ at a time. Furthermore, let's say that half the time you get a $1$ and half the time you get a $0$; those are the only values that the $\mathbf Z$ factory can produce. And the factory is built to reset itself between producing each value, so whatever $z_i$ is has no impact on $z_{i+1}$.
# In the language of probability theory, $z_i$ are **realizations** from $\mathbf Z$, which has a **distribution**:
#
# $$
# \begin{array}{rcl}
# P(\mathbf Z = 0) &=& 1/2 \\
# P(\mathbf Z = 1) &=& 1/2
# \end{array}
# \quad \quad \text{or} \quad \quad
# P(\mathbf Z=z) =
# \begin{cases}
# 1/2 & \text{for }z=0 \\
# 1/2 & \text{for }z=1
# \end{cases}
# $$
# What we've been loosely calling a "factory" is a **random variable** in the language of probability theory. But that's just a name. You can keep thinking of them as factories, or code, that generate data.
# <div class="alert alert-block alert-warning">
# <b>Note:</b>
# Random variables are often written in uppercase, (e.g. Z) and their realizations in lowercase (z). We're going to be using uppercase for matrices (and sets), so I'm going to use boldface in conjunction with uppercase ($\mathbf Z$) to denote random variables.
# </div>
# Ok, so if the random variable is a factory, and the realizations of the random variable are the output of that factory (the data we get to see), then how do we read a statement like $P(\mathbf Z = 0) = 1/2$? Well, that just means that the value $z$ that $\mathbf Z$ produces is $0$ half of the time. But what exactly do we mean by "half the time"? While we usually don't have to think deeper than this, you'll see later that it is sometimes necessary to have a more rigorous definition of probability.
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Remember that the entire purpose of talking about these factories is so that we can imagine what's behind the curtain, producing the data that we observe. Think of a real-world scenario where we could pretend that the data we observe was generated by $\mathbf Z$. In other words, what's something we could measure in the real world that we might model using $\bf Z$?
#
# </div>
# Let's build that definition. We'll start with some raw materials. All factories have raw materials that go into them, which end up being turned into the finished product. In a similar way, random variables have inputs which get mapped to realized values. We'll call them "data ore": the unrefined precursor that gets transformed by our factory (random variable $\mathbf Z$) into the data product $z$. The data ore exists in units (data ore nuggets). The factory takes one nugget at a time and transforms it into a realization.
#
# The nuggets are kept in an big silo called $\Omega$ before they go to $\mathbf Z$. This silo is filled to the brim with *all* of the possible nuggets that could be fed into the factory, one of each of them. It's also a magic silo, so when you take out a nugget, another one exactly like it is mined out of the depths of the cosmos to take its place in the silo.
# 
# Each nugget is gets transformed into a value of $z$, but the process isn't random. For instance, if a nugget named "Karl" turned into a 1 when fed through $\mathbf Z$, then we would *always* get a 1 when Karl goes into $\mathbf Z$. But we know that sometimes $\mathbf Z$ produces 0s, so there must be other nuggets whose destiny is to become 0s, just like Karl's destiny is to be a 1. The "randomness" in $\mathbf Z$ isn't caused by what's in the factory, it's caused by randomly picking a nugget to throw into it.
# We can even code up our little example, imagining that we have 10 nuggets, boringly named "0", "1", "2"... "9":
# +
def Z(ω): # factory (random variable)
if ω in set([1,4,5,8,9]): # these are the outcomes (nuggets) that map to the value 1
return 1
if ω in set([0,2,3,6,7]): # these are the outcomes (nuggets) that map to the value 0
return 0
Z.Ω = set([0,1,2,3,4,5,6,7,8,9]) # sample space (silo) of outcomes (ore nuggets) attached to Z
import random
def realize(rand_var): # run the assembly line!
ω = random.sample(rand_var.Ω, 1)[0] # grab a single nugget out of the silo at random
return rand_var(ω) # push it through the factory
# -
# <div class="alert alert-block alert-warning">
# <b>Python Tip:</b>
#
# `random.sample(x,n)` grabs `n` values at random out of the set `x` and returns them as a list.
# </div>
# Here are 20 observations $z=[z_1, z_2, \dots z_{20}]$, fresh off the assembly line of the $\mathbf Z$ factory:
z = [realize(Z) for i in range(20)]
z
# Now we're ready to define probability: the probability of an realization (a particular value $z$) is just the proportion of the silo that's taken up by nuggets that are destined to become that value $z$ when fed through $\mathbf Z$. That's it. We denote that proportion with the notation $P(\mathbf Z = z)$. In our example above, saying $P(\mathbf Z = 1) = 1/2$ means that half of all the possible nuggets that could go into $\mathbf Z$ would produce a 1, assuming each nugget takes up the same amount of space.
# That's a definition we can code up:
def P(rand_var, realization):
A = set(ω for ω in rand_var.Ω if rand_var(ω) in realization) # what are all the nuggets that map to the value(s) in question?
return len(A)/len(rand_var.Ω) # what is the "volume" of those nuggets relative to the volume of the silo Ω? (assuming each takes up the same amount of space)
P(Z,[0]), P(Z,[1]) # P(z=0), P(z=1)
# So to build a factory that makes 0s and 1s in even proportions, all I had to do was evenly split up the number of nuggets that are destined to produce each value. It also doesn't matter what I call the nuggets. For example, here is equally good code to implement $\mathbf Z$:
# +
def Z(ω): # factory (random variable)
if ω in set([-1234]): # these are the outcomes (nuggets) that map to the value 1
return 1
if ω in set([980123]): # these are the outcomes (nuggets) that map to the value 0
return 0
Z.Ω = set([980123, -1234]) # sample space (silo) of outcomes (ore nuggets) attached to Z
[realize(Z) for i in range(20)]
# -
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Write code for a new random variable $\mathbf W$ that behaves like this:
#
# $$
# P(\mathbf W=w) =
# \begin{cases}
# 0.1 \dots & \text{for }w=-1 \\
# 0.4 \dots & \text{for }w=0 \\
# 0.2 & \text{for }w=1 \\
# 0.3 & \text{for }w=2
# \end{cases}
# $$
#
# You'll need to make your own nugget silo `Ω` and define the function `W(ω)`. Test it out using the `realize()` and `P()` functions we wrote. Use `P()` to calculate $P(\mathbf W =0)$.
#
# </div>
# ### A mathematical description of random variables
# 
# If you're looking at this and thinking that I can't possibly be serious, that the foundations of statistics and machine learning can't possibly be built up from imagining data factories and magical silos... well, you're wrong. Sure, I've concocted a somewhat elaborate metaphor, but it's a metaphor that accurately describes how these otherwise very abstract concepts relate to each other. If you can look at something like $P(\mathbf Z = z) := \mathbb P(\{\omega \in \Omega \vert \mathbf Z(w)=z\})$ and immediately come away with an understanding of what that means, all the more power to you. But I don't. At least not without first building up an intuition for each of the components.
# In probability theory, the silo $\Omega$ is called a **sample space** and the data ore nuggets $\omega$ are called **outcomes** (not to be confused with what we call the variable we want to predict in machine learning). A random variable $\mathbf Z$ is defined as a function that maps an element $\omega$ of $\Omega$ to a realization $z$. The probability of a realization $z$ is the **measure** (volume, or proportion of total volume) of the set of outcomes (data ore nuggets) that map to $z$ (are destined to be transformed to $z$ by $\mathbf Z$).
# 
# When I talk about these things outside of the context of explaining them, I do call them by their real names (e.g. random variable, sample space, etc.) because that's what people have called them for nearly a century. But when I close my eyes and *reason* about these concepts, I'm thinking about something tangible, like a factory. As we go on I'm going to introduce more mathematical notation as we need it, and I'm going to wean off the factory metaphor, but I encourage you to keep building your intuition about these concepts instead of thinking about them as abstract symbols on a page. The symbols are just a convenient shorthand for the ideas. The only reason to know the standard names and symbols is to be able to read and understand what others have written. If you find yourself skimming over an equation- stop. Read it slowly and think about what each part means.
#
# So now that we're here, let's demystify the notation in that equation I dropped up above! Here it is again:
#
# $$P(\mathbf Z = z) := \mathbb P(\{\omega \in \Omega \vert \mathbf Z(w)=z\})$$
#
# To start, the $:=$ means "the thing on the left is defined as the thing on the right". So we're saying that when we write "$P(\mathbf Z = z)$", we really mean whatever "$\mathbb P(\{\omega \in \Omega \vert \mathbf Z(\omega)=z\})$" is. Ok, next up is [set-builder notation](https://www.mathsisfun.com/sets/set-builder-notation.html): you can read $\{a\in A | f(a) = 1\}$ as "the collection of all the elements $a$ in the set $A$ *such that* $f(a)=1$". So $\{\omega \in \Omega \vert \mathbf Z(\omega)=z\}$ is the set of outcomes $\omega$ that become $z$ when passed through the random variable $\mathbf Z$. There may be many such outcomes, or just one, or none, so the set can be big, small, or nonexistent. We will write the name of that set a little more compactly using the notation $\mathbf Z^{-1}(z) = \{\omega \in \Omega \vert \mathbf Z(w)=z\}$ since usually $f^{-1}(y)$ denotes the element $x$ such that $f(x)=y$. We call this the **preimage** of $z$ under $\mathbf Z$.
# <div class="alert alert-block alert-warning">
# <b>Note:</b>
#
# Preimages aren't just for random variables- you can define preimages for any function. If the function is $y=f(x)$, the preimage of a set $A$ (denoted $f^{-1}(A)$) is a set of all of the values $x$ that become one of the $y$ values in $A$ when shoved through $f$. The set $A$ is called the image of $f^{-1}(A)$ under $f$.
#
# For example, if $f(x) = x^2$ and $A$ is the set of numbers between 0 and 4, then $f^{-1}(A)$ is the set of numbers between -2 and 2, since every number between -2 and 2, when squared, is between 0 and 4, and these are the only numbers for which that is the case. Another example: if $f(x) = \cos(x)$ and $A=\{1\}$, then $f^{-1}(A) = \{\dots, -4\pi, -2\pi, 0, 2\pi, 4\pi, 6\pi, \dots\}$. Plot or draw a picture of $\cos(x)$ and mark the points where $\cos(x) = 1$ to see why.
#
# </div>
# Finally, we have $\mathbb P()$, which is the [**probability measure**](https://en.wikipedia.org/wiki/Probability_measure). Think of it as a function that measures the proportion of all of the outcomes in $\Omega$ that are contained in the subset $\mathbf Z^{-1}(z)$. This is basically the volume of space that the nuggets in $\mathbf Z^{-1}(z)$ take up in the silo $\Omega$. By convention, we say that $\Omega$ has volume 1 so that the volume of $\mathbf Z^{-1}(z)$ is also the proportion of volume that $\mathbf Z^{-1}(z)$ takes up in $\Omega$. In the figure above, that's represented by the area of the shaded gray region.
#
# If you put all of that together, you'll see that it's exactly the same as the definition we put together using our factory analogy.
# We can also talk about the probability of sets of realizations instead of just single realization $z$. For instance, what's the probability that $z$ is 0 *or* 1? We write that like $P(\mathbf Z \in A)$, where $A$ is the set of possible realizations, like $\{0,1\}$. That's more general than the probability of a single realization $z$: $P(\mathbf Z = z)$. The definition is the same though: $P(\mathbf Z \in A) := \mathbb P(\mathbf Z^{-1}(A))$. All we need to do is count up the volume of all the nugets that produce any of the values that are in $A$, instead of just the nuggets that produce $z$.
# <div class="alert alert-block alert-warning">
# <b>REMEMBER:</b>
#
# If your eyes glaze over every time you see mathematical notation, don't worry. Remember, that's normal. Just slow down and read it again. Try and think about what it <i>means</i>.
# </div>
# If you look at the code we wrote before, you'll notice it can already calculate probabilities for sets of realizations:
def P(rand_var, realization):
A = set(ω for ω in rand_var.Ω if rand_var(ω) in realization) # what are all the nuggets that map to the value(s) in question?
return len(A)/len(rand_var.Ω) # what is the "volume" of those nuggets relative to the volume of the silo Ω? (assuming each takes up the same amount of space)
P(Z, [0])
P(Z, [0,1])
# ### Properties of probability
# Ok- I promised that it would be useful to define probability in a more rigorous way than "$z$ happens $x$% of the time". Now we're going to see why.
#
# To start with, let's "derive" a relatively simple fact: for any subset of possible realizations $A$,
#
# $$P(\mathbf Z \in A) \in [0,1]$$
#
# This is a compact way of writing that for any subset of realizations, the volume of the subset of outcomes $\mathbf Z^{-1}(A)$ that map to those realzations is a number between 0 and 1. Why? Well, if the volume of our silo $\Omega$ is 1, the volume of any subset of that has to be less than or equal to 1. And there is no subset that can occupy negative space, so the volume has to be greater than or equal to 0.
# Here's a trickier one: if two sets of realizations $A$ and $B$ have no realizations in common, then the probability of a realization from either of them is the sum of the probabilities of a realization from each of them. Mathematically:
#
# $$A \cap B = 0 \rightarrow P(\mathbf Z \in A \cup B) = P(\mathbf Z \in A) + P(\mathbf Z \in B)$$
#
# $A \cap B$ is read as "the intersection of the sets $A$ and $B$", which is the set of elements that are in both sets. It's the middle part of a Venn diagram. $A \cup B$ is read as "the union of $A$ and $B$", which is all of the elements in either set- that's the entirety of the Venn diagram.
#
# That also seems cryptic until you think about it in terms of quantities of ore nuggets that produce certain values when fed through the factory. If you take all the ore nuggets that end up becoming any of the values in $A$ (call that set of nuggets $\mathbf Z^{-1}(A)$), and all the nuggets that end up becoming values in $B$ (call that $\mathbf Z^{-1}(B)$), then the total volume that end up becoming values in either $A$ or $B$ is the sum of the volumes that become $A$ and those that become $B$. This is true as long as there are no nuggets that become both a realization in $A$ and a realization in $B$ because we would double-count these. But we've also ensured that these do not exist since each nugget is destined to become only a single value, and we made sure that there is no overlap between $A$ and $B$.
#
# If there is overlap, the proposition doesn't hold. For instance, if $A= \{0,1\}$ and $B = \{0\}$, every element of $B$ is also an element of $A$, so the volume of $Z^{-1}(A \cup B)$ is the volume of $Z^{-1}(A)$, which is not the volume of $Z^{-1}(A)$ plus the volume of $Z^{-1}(B)$.
# We can even use our code from before to demonstrate this:
A = set([0])
B = set([1])
P(Z,A) + P(Z,B) == P(Z,A|B) # in python, set union ∪ is written | because an element is in A∪B if it is in A OR B (A|B)
A = set([0,1])
B = set([0])
P(Z,A) + P(Z,B) == P(Z,A|B)
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Draw a picture based on the figure above that helps explain why $A \cap B = 0 \rightarrow P(\mathbf Z \in A \cup B) = P(\mathbf Z \in A) + P(\mathbf Z \in B)$
#
# </div>
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Let's say the sets $A$ and $B$ have some overlap. Can you come up with a formula to calculate $P(\mathbf Z \in A \cup B)$ given $P(\mathbf Z \in A)$, $P(\mathbf Z \in B)$, and $P(\mathbf Z \in A \cap B)$?
#
# </div>
# The upshot of this is that the probability of a set of outcomes is the same as the sum of their probabilities:
# $$
# P(\mathbf Z \in A)
# =
# \mathbb P (\mathbf Z^{-1}(A))
# =
# \sum_{\omega \in Z^{-1}(A)} \mathbb P(\omega)
# =
# \sum_{Z^{-1}(A)} \mathbb P(\omega)
# $$
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# In all our code so far we've been using a finite sample set with $n$ outcomes and we've chosen to use $\mathbb P(B)= |B|/n$ where $|B|$ denotes the number of elements in $B$. That's called the <b>counting measure</b> It helps your understanding, however, to know that it isn't the only probability measure we could use. We could instead say that some outcomes take up twice as much space in the silo, or that they all have totally different volumes. As long as whatever $\mathbb P$ we come up with satisfies $\mathbb P(\Omega)=1$ and $\mathbb P(\bigcup B_i)= \sum \mathbb P(B_i)$ for non-overlapping sets $B_i$ (of outcomes), it's a legitimate choice.
#
# Let's go back to this version of $\mathbf Z$:
#
# ```
# def Z(ω): # factory (random variable)
# if ω in set([1,4,5,8,9]): # these are the outcomes (nuggets) that map to the value 1
# return 1
# if ω in set([0,2,3,6,7]): # these are the outcomes (nuggets) that map to the value 0
# return 0
# Z.Ω = set([0,1,2,3,4,5,6,7,8,9]) # sample space (silo) of outcomes (ore nuggets) attached to Z
# ```
#
#
# Change the code for `P(rand_var, realization)` so that $\mathbb P(\omega) = 0.25$ if $\omega \in \{0,1,2,3\}$ and 0 otherwise. The idea is that now nuggets 0, 1, 2, and 3 each take up a quarter of the space in the silo, while the other nuggets take up none. What is $P(Z=1)$ now?
#
# </div>
# ### Continuous sample spaces
# So far, all the random variables we've talked about have produced outputs from a finite, discrete set (e.g. $\{0,1\}$ or $\{-1,0,1,2\}$). If we're imagining a factory that might produce the data we observe when flipping a coin, a binary output is all we need. Similarly, if we want to imagine the factory that assigns an "apple", "orange", or "banana" label to a photograph of a fruit, it just needs to output a discrete set of three values. But if we want to imagine the kind of factory that could produce the prices of different apartments in New York, we need something that can output a continuous range of values.
# Let's think up a random variable (call it $\bf Z$ again) that can take any value between 0 and 10. How many numbers are there between 0 and 10? Well, an infinite number: for any two numbers in that interval, you can find a number that's right between them. Since one nugget from the silo always prodcues the same realization when pushed through the factory, there need to be an infinite number of nuggets in the silo to be able to produce an infinite number of realizations. That means that our old code, where we manually enumerated all of the elements in $\Omega$, is not going to work anymore. What we can do instead is imagine that $\Omega$ is itself an interval, like all the numbers between 0 and 1. So, to pick a random nugget to throw into the factory, we just pick a random number between 0 and 1. Here's an example:
# +
def Z(ω):
return 10*(ω**2) # when ω goes into the factory, the factory makes ω^2
Z.Ω = random.random # returns a single number between 0 and 1 when called
def realize_cont(rand_var): # run the assembly line!
ω = Z.Ω() # returns a single number between 0 and 1
return rand_var(ω) # push it through the factory
# -
[realize_cont(Z) for i in range(5)]
# So $\mathbf Z$ is defined by $\mathbf Z(\omega) = 10\omega^2$ with $\omega \in [0,1]$. Great. But now what does $P(\mathbf Z = z)$ mean? We just apply the same old definition of probability: it's the proportion of nuggets in the silo that are destined to become the value $z$. In notation: $\mathbb P(\mathbf Z^{-1}(z))$. Same as before.
# <div class="alert alert-block alert-warning">
# <b>Note:</b>
#
# Notation like $[a,b]$ is often used to concisely write intervals- this just means "all the numbers between $a$ and $b$, including those endpoints". We use parentheses like $(a,b)$ to indicate that the endpoints should not be included. $(a,b]$ and $[a,b)$ have one of the two endpoints included, with the bracket indicating the endpoint that's included and the parenthesis indicating which isn't.
# </div>
# The issue is that now we need a probability measure that works with continuous sets. For example, let's say we're looking for $P(\mathbf Z = 2.5)$. As $\mathbf Z(\omega) = 10\omega^2$ is defined in the code above, the only value of $\omega$ that makes $z=2.5$ is $\omega = \sqrt{2.5/10} = 0.5$. Any other value of $\omega$ would produce a different value of $z$. So $\mathbf Z^{-1}(z) = 0.5$. What "volume" does the single number $0.5$ take up in the interval $[0,1]$? In other words, how are we going to define a probability measure to use here?
# The most commonly used measure in this case is based on the "length" of the set relative to the length of $\Omega$. In our case, the length of $\Omega$ is 1, so the probability measure of any interval $(a,b)$ or $[a,b]$ is $b-a$. For sets more complicated than an interval, we have to find the smallest collection of intervals, in terms of total length, that contains the set in question. We say the length of that set is the total length of the collection of intervals that covers it. Using length as a notion of measure makes good sense because if two sets don't overlap, then the length of their union is the sum of their lengths. This measure is called the **Lebesgue measure**, but I only mention the name so you can recognize it elsewhere.
# <div class="alert alert-block alert-warning">
# <b>Note:</b>
#
# When you get down to the nitty gritty math, it turns out there actually are <a href=https://en.wikipedia.org/wiki/Vitali_set>some really messed up sets</a> where this notion of "length" breaks down, in that the "length" of the union of two disjoint sets might not be the sum of their lengths. These are not sets you would ever come across in any real-world context. The technical solution is to only allow random variables where the preimage of any interval is not one of these messed up sets. This really isn't something you should think or worry about. This note is only here to satisfy nosy probabilists or measure theorists who were offended by the above paragraph.
#
# </div>
# Ok, back to our problem: what's $\mathbb P(0.5)$? Well, $\{0.5\} = [0.5, 0.5]$, so its length is $0.5-0.5=0$! In fact, for any single element $\omega$, $\mathbb P(\omega)= 0$ for the same reason. That's a problem if we want to use discrete sums to calculate probabilities over sets:
# $$
# P(\mathbf Z \in A)
# \overset{?}{=}
# \sum_{Z^{-1}(A)} \mathbb P(\omega)
# =
# \sum_{Z^{-1}(A)} 0
# =
# 0
# $$
# But if $Z^{-1}(A)$ is an interval with finite length, then the probability has to be the length of that interval, not 0!
#
# The reason this doesn't make any sense is that we're trying to use a discrete sum to add up a continuous infinity of 0s. Basically, we're trying to break down $\sum_{Z^{-1}(A)}$ into each of its component $\omega$s and measuring each of those. Instead of doing that, though, we can *integrate* over infinitesimal units of "$d \omega$":
# $$
# P(\mathbf Z \in A)
# =
# \int_{Z^{-1}(A)} \mathbb P(d\omega)
# $$
# This thing is called a **Lebesgue integral**. What we're doing here is adding up all of the infinitesimal lengths $\mathbb P(d\omega)$ for all $\omega$s in the set $\mathbf Z^{-1}(A)$. We'll write this as $\int d \mathbb P$ for short. It has all the same rules as a standard integral (just write $d \mathbb P$ instead of $dx$), so the integral of a sum is the sum of integrals, etc. And it always agrees with the integrals you're used to from calculus:
#
# $$
# \int_{[a,b]} f(\omega) d\mathbb P = \int_a^b f(x) dx
# $$
#
# The neat thing is that it actually works no matter what $\mathbb P$ is, as long as it satisfies all the properties of a measure. In fact, if $\mathbb P$ is the discrete counting measure that we were using before, then
# $$
# \int_{Z^{-1}(A)} d \mathbb P
# =
# \sum_{Z^{-1}(A)} \mathbb P(\omega)
# $$
# If you have no idea why any of this matters, don't worry, just keep going. We're not going to get into the theory of Lebesgue integration. I really went back and forth on whether to include this at all, but I did because having this unifying formalism in your back pocket makes it really easy to prove a lot of things later, even if you don't really understand the theoretical details. You'll be fine if you just think of a Lebesgue integral as a tool to find the volume of outcomes in arbitrary sets that happens to follow all the rules of a normal integral. In other words: no matter how you're measuring stuff, you can use the Lebesgue integral to figure out how much space different sets of outcomes take up.
# ## Probability distributions
# The formal definition of a random variable as a function from a sample space to some set of numbers is really useful for proving useful relationships, but ultimately the sample space is totally imaginary: all we get to see are the realizations. So we're going to build some tools that will let us avoid talking about the sample space so much if we don't need to.
# As perhaps you've noticed, neither the exact nature of what is in the sample space nor which of its elements map to which realizations change the observable behavior of a random variable as long as the total measure of all the outcomes mapping to each realization are the same. For example, we looked at two equivalent ways to implement our random variable $\mathbf Z$:
def Z(ω):
if ω in set([1,4,5,8,9]):
return 1
if ω in set([0,2,3,6,7]):
return 0
Z.Ω = set([0,1,2,3,4,5,6,7,8,9])
def Z(ω):
if ω in set([-1234]):
return 1
if ω in set([980123]):
return 0
Z.Ω = set([980123, -1234])
# These are technically two different random variables because they have different sample spaces and different mappings to the realizations, but they behave exactly the same. When this is the case, we say they have the same **probability distribution**. The probability distribution describes how the factory should *behave* from the perspective of someone who can only see its products $z_i$: half the time you get a 0, half the time you get a 1. There is no need to mention the silo of ore nuggets, give them names, and specify which nuggets are destined to be 0s and which are destined to be 1s. We know they're back there, and we know what total *measure* are destined to be 0s and 1s (since that's what the probability means), but we don't need the details of who is who and what goes where. In fact, unless you're a probability theorist, you will never need to think about the sample space to solve a problem. The only reason you need to know about it is so that you can understand useful identities, which we will continue to derive as we go along.
# The discrete probability distribution is function of the factory product $z$. For each unique value of $z$, it tells us the total volume of the nuggets in the silo that map to that outcome. We can visualize that by sorting all the nuggets in the silo into piles according to which value they are destined to become. The relative heights of each pile are proportional to the volume of space (measure) that each group of nuggets take up in the silo. Let's demonstrate with a new random variable $\bf V$:
def V(ω):
if ω in set([1]):
return 2
if ω in set([2,3]):
return 0
if ω in set([4,5,6]):
return -1
if ω in set([7,8,9,0]):
return 1
V.Ω = set(range(10))
# +
vs = [-1,0,1,2] # all the values v can take
ps = [P(V,[v]) for v in vs] # calculate the probability of each, assuming the counting measure
import altair as alt # for plotting
import pandas as pd # to make dataframes
distribution = pd.DataFrame({'v':vs, 'p':ps})
alt.Chart(distribution, height=100, width=400).mark_bar().encode(x='v:O', y='p')
# -
# <div class="alert alert-block alert-warning">
# <b>Python Tip:</b>
#
# `altair` is a useful python package for visualization. It's optimized to work with dataframes from the `pandas` package. Feel free to browse the documentation for these packages, but you don't need to be an expert to continue on in this book.
# </div>
# This is the graphical representation of the probability distribution
#
# $$
# \phi(v)
# =
# P(V=v)
# =
# \mathbb P(\mathbf V^{-1}(v))
# =
# \begin{cases}
# 0.3 & \text{for }v=-1 \\
# 0.2 & \text{for }v=0 \\
# 0.4 & \text{for }v=1 \\
# 0.1 & \text{for }v=2
# \end{cases}
# $$
# $\phi(v)$ is called a **probability mass function**. If we have multiple random variables floating around and we want to distinguish their mass functions, we'll sometimes write $\phi_{\mathbf V}(v)$.
# If we want to know the probability of a particular set of realizations, say, $P(\mathbf V \in \{0,1\})$, it's easy to get using the mass function:
#
# $$P(\mathbf V \in A) = \sum_{v \in A} \phi(v)$$
#
# We simply sum up the probabilities that $\mathbf V$ is any of the realizations within the set $A$ of interest. Compare this to what we had before:
#
# $$P(\mathbf V \in A) = \sum_{\omega \in \mathbf V^{-1}(A)} \mathbb P(\omega)$$
#
# The advantage is that we don't have to talk about outcomes or sample spaces anymore. All of the information we need to calculate any probabilities of $\mathbf V$ is baked into the mass function $\phi(v)$.
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Let's say $\mathbf V$ is a random variable that maps outcomes from the interval $[0,1]$ to either 0, 1, or 2 in the following way:
#
# $$
# \mathbf V(\omega)
# =
# \begin{cases}
# 0 & \text{if } \omega \in [0, 0.2) \cup (0.8,1] \\
# 1 & \text{if } \omega \in [0.2, 0.3) \cup (0.7, 0.8] \\
# 2 & \text{if } \omega \in [0.3, 0.7] \\
# \end{cases}
# $$
#
# Note that $\mathbf V$ is discrete, but with a continuous sample space.
#
# Assuming the Lebesgue measure, what is the mass function of $\mathbf V$? In other words, for each value that $\mathbf V$ can take, what's the total length of the set that produces each value?
#
# Use the mass function to calculate $P(\mathbf V \in \{1,0\})$. You should get 0.6.
#
# </div>
# ### Continuous random variables and densities
# Let's say $\mathbf Z$ is defined by $\mathbf Z(\omega) = 10\omega^2$ with $\omega \in [0,1]$. How can we find some kind of function that we can manipulate to calculate probabilities without reference to the sample space or measure?
#
# For starters, we do know how to calculate probabilities. For instance, if we want to know $P(\mathbf Z \in [0.625, 2.5])$, what we need to do is find $\mathbb P(\mathbf Z^{-1}([0.625, 2.5]))$, which is the "length" of the set $\mathbf Z^{-1}([0.625, 2.5])$ if we're using the Lebesgue measure. So what is $\mathbf Z^{-1}([0.625, 2.5])$? Well, $\mathbf Z(\omega) = 10\omega^2 \in [0.625, 2.5]$ is the same as saying $0.625 \le 10\omega^2 \le 2.5$. Dividing by 10 and taking square roots, we're left with $0.25 \le \omega \le 0.5$. So $\mathbf Z^{-1}([0.625, 2.5]) = [0.25, 0.5]$. The length of that set is clearly 0.25, so that's the probability we're looking for.
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Let $\mathbf Z$ be as it is above. Find a formula for $P(\mathbf Z \in [a,b])$ for any values $a \le b$ and $a,b \in [0,10]$.
#
# </div>
# Mission accomplished? Not quite. We managed to calculate a probability given the sample space and random variable, but what we want is some kind of function that we can manipulate to calculate these probabilities without reference to the sample space at all. We don't want to have to think about what $\Omega$ is or exactly how the different outcomes map to the different realizations.
# So here's an idea: let's bin $z$ into 10 non-overlapping buckets, like $[0,1)$, $[1,2)$ ... $[9,10]$ and calculate the probability within each of those buckets. This is just like what we did in the discrete case. We're sorting all of the nuggets in the silo into different piles depending which set of values they are destined to become, and then measuring the volume of each pile. Here's what we get when we do that for the example random variable $\bf z$ defined in the code above:
# +
from math import sqrt
def Pz(a,b):
return sqrt(b/10) - sqrt(a/10)
zs = range(10)
ps = [Pz(z,z+1) for z in zs]
zs_labels = [f'[{z},{z+1})' for z in zs]
distribution = pd.DataFrame({'z':zs_labels, 'p':ps})
alt.Chart(distribution, height=100, width=400).mark_bar().encode(x='z:O', y='p')
# -
# And why stop at 10 buckets? Let's split it up into 100.
# +
zs = np.arange(0,10,0.1)
ps = [Pz(z,z+0.1) for z in zs]
zs_labels = [f'[{z},{z+0.1})' for z in zs]
distribution = pd.DataFrame({'z':zs_labels, 'p':ps})
alt.Chart(distribution, height=100, width=400).mark_bar().encode(alt.X('z:O',axis=None), y='p')
# -
# More buckets gives us more information. If we want to know $P(\mathbf Z \in [0,0.5))$, for instance, we can sum up the probabilities for the buckets $[0,0.1)$, $[0.1,0.2)$, ... $[0.4,0.5)$. But we can't get *any* probability. The graph doesn't have enough information to let us calculate probabilities over intervals whose ends are between two cutpoints of the buckets. It only has resolution up to increments of $0.1$ in terms of $z$. It would be nice to have a graph that lets us read off arbitrary probabilities like $P(\mathbf Z \in [a,b])$ just by looking at how much "stuff" there is between $a$ and $b$. Something like this:
# +
z = np.arange(0.1,10,0.1)
p = 1/(2*np.sqrt(10*z)) # magic, for now...
distribution = pd.DataFrame({'z':z, 'p':p})
alt.Chart(distribution, height=100, width=400).mark_area().encode(x='z', y='p')
# -
# Before I explain how I managed to make this graph, which is called a **density plot**, I want to establish an intuition for what it means. We've gone from 10 buckets, to 100 buckets, to "infinite" buckets. I like to think of these pictures literally: all the outcomes $\omega$ neatly piled up on top of the labels $z$ for the values they will become. So to get $P(\mathbf Z \in [a,b])$ from this picture, which is just the volume of outcomes that map to values between $a$ and $b$, all we need to do is see how much stuff there is piled up between $a$ and $b$ in the picture.
# 
# To do this, we turn to a useful tool from calculus: the integral. To make the picture above, we need a curve $\phi(z)$ such that the area under $f$ between $a$ and $b$ is $P(\mathbf Z \in [a,b])$ for all values $a$ and $b$. In the previous exercise you should have figured out that $P(\mathbf Z \in [a,b]) = \sqrt{\frac{b}{10}} - \sqrt{\frac{a}{10}}$. So what we need is the curve $\phi(z)$ that satisfies this equation:
#
# $$\int_a^b \phi(z) dz = P(\mathbf Z \in [a,b]) = \sqrt{\frac{b}{10}} - \sqrt{\frac{a}{10}}$$
# Looking at the integral equation, it's clear that $\Phi(z) = \sqrt{\frac{z}{10}}$ is the antiderivative of $\phi(z)$, so all we need to do to get $\phi$ is differentiate $\Phi$:
#
# $$\phi(z) = \frac{d\Phi(z)}{dz} = \frac{d}{dz} \sqrt{\frac{z}{10}} = \frac{1}{2\sqrt{10z}}$$
# That's why we have `ps = [1/(2*sqrt(10*z)) for z in zs]` in the code above.
# The function $\phi(z)$ is called a **probability density function** (PDF), which is the continuous equivalent of the probability mass function. Its integral $\Phi(z) = \int_{-\infty}^z \phi(t)dt = P(\mathbf Z \le z)$ is called a **cumulative density function** (CDF). Either of these functions tell you everything you need to know about probabilities of the random variable $\mathbf Z$. The probability that $\mathbf Z$ takes any of the values in an arbitrary set $A$ is
#
# $$P(\mathbf Z \in A) = \int_{A} \phi(z) dz$$
# This works the same way as the probability mass function for a discrete random variable $\mathbf V$:
#
# $$P(\mathbf V \in A) = \sum_{v \in A} \phi(v)$$
A = (1<=z) & (z<=4)
distribution = pd.DataFrame({'z':z, 'p':p, 'A':A})
alt.Chart(distribution, height=100, width=400).mark_area().encode(
x='z',
y='p'
) + alt.Chart(distribution.query('A')).mark_area(color='orange').encode(
x='z',
y='p'
)
# For example, the probability that $\mathbf Z$ is in the set $[1,4]$ is the area shaded in orange above.
# <div class="alert alert-block alert-warning">
# <b>Note:</b>
#
# The notation $\int_{A} \phi(z) dz$ just means $\int_{-\infty}^\infty I_A(z)\phi(z) dz$ where the <b>indicator function</b> $I_A(z)$ is 1 if $z\in A$ and 0 else. In othe words, all we're doing is summing up the $\phi(x)dx$s where $x \in A$. That's analogous to summing up the $\phi(v)$s where $v \in A$ in the discrete case.
#
# </div>
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# For our random variable $\mathbf Z$ with density $\phi(z) =\frac{1}{2\sqrt{10z}}$, what is $P(\mathbf Z \in [0, 10])$? Calculate the probability by integrating the density function. Does your answer line up with what you expect based on our original definition of $\mathbf Z$?
#
# Here is another random variable that, like $\mathbf Z$, maps outcomes in $\Omega = [0,1]$ to values in $[0,10]$: $\mathbf W(\omega) = 10\omega$. Calculate $P(\mathbf W \in [a,b])$ for some interval $[a,b]$. What is the probability density function for $\mathbf W$? What is $P(\mathbf W \in [0, 10])$?
#
# For <i>any</i> continuous random variable $\mathbf X$, what is $\int_{-\infty}^{\infty} \phi(x) dx$ (<i>hint</i>: what probability does this represent)? What is $\Phi(-\infty)$? $\Phi(\infty)$?
#
# Is it possible to have a random variable $\mathbf Q$ with $\phi_{\mathbf Q}(q) < 0$ for some $q$ that is a possible realization of $\mathbf Q$? Why does this not make sense?
#
# For two values $a < b$, is it possible that $\Phi(a) > \Phi(b)$? Why nor why not?
#
# </div>
# At this point, talking about the outcomes $\omega$ is kind of silly. If two random variables have the same probability mass function or the same probability density function, then, for all intents and purposes, they are the same random variable. It doens't matter exactly which outcomes map to which values, as long as the proportions are the same. We already demonstrated this in the discrete case.
#
# To show the same concept for continuous random variables, here is a new random variable $\mathbf Z'$ whose sample space $\Omega$ is $[-100, 100]$ instead of $[0,1]$, but which has the same probability density function as our other random variable $\mathbf Z$:
#
# $$
# \mathbf Z'(\omega) = 10\left(\frac{\omega+100}{200}\right)^2
# $$
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Prove to yourself that $\mathbf Z'$ has the same probability density function as $\mathbf Z$.
#
# </div>
# For this we use the notation $\mathbf Z \sim \mathbf Z'$. Technically they are not the same since the sample spaces are different, so we shouldn't write $\mathbf Z = \mathbf Z'$. But as far as an observer who is outside the curtain is concerned, there is no way to tell them apart. The *distribution* of the random variable is what really matters.
# I think about mass or density functions as convenient abstraction layers between me and the random variable. If I want to know a probability, I don't have to go to the random variable and count up the volume of something in the sample space, I just "query" the mass or density. The "query engine" happens to be an integral or sum, and the query itself is the region of space that I want to integrate over. In a nutshell:
#
# $$
# \mathbb P(\mathbf Z^{-1}(A)) = \int_A \phi_{\mathbf Z}(z) dz
# \quad
# \text{or}
# \quad
# \mathbb P(\mathbf Z^{-1}(A)) = \sum_{z \in A} \phi_{\mathbf Z}(z)
# $$
# So if we have $\phi_Z$, we don't need to worry about figuring out what $\mathbf Z^{-1}(A)$ is or how to do the measurement of that set using $\mathbb P$. Finding preimages and measuring them is hard. Integrating or summing distribution functions is easier.
#
# ### Histograms vs. mass and density functions
# Many of you are probably already familiar with histograms. Histograms are a way of visualizing observed data. Each observed value is stacked up on top of its approximate label (e.g. any $z$ between 0.5 and 1.5 is labeled "1") and the counts are plotted:
# +
def Z(ω):
return 10*(ω**2) # when ω goes into the factory, the factory makes ω^2
Z.Ω = random.random # returns a single number between 0 and 1 when called
def realize_cont(rand_var): # run the assembly line!
ω = Z.Ω() # returns a single number between 0 and 1
return rand_var(ω) # push it through the factory
# +
z = [realize_cont(Z) for i in range(1000)] # 1000 draws from Z
plot_df = pd.DataFrame({'z':z})
alt.Chart(plot_df, height=100, width=400).mark_bar().encode(
alt.X('z', bin=alt.Bin(maxbins=100)),
y='count()'
)
# -
# That looks suspicously like our bucketed density plot:
# +
zs = np.arange(0,10,0.1)
ps = [Pz(z,z+0.1) for z in zs]
zs_labels = [f'[{z},{z+0.1})' for z in zs]
distribution = pd.DataFrame({'z':zs_labels, 'p':ps})
alt.Chart(distribution, height=100, width=400).mark_bar().encode(alt.X('z:O',axis=None), y='p')
# -
# So what's the difference? Think about what it is we're "stacking up" in the bars. In the histogram, we're sorting and stacking up a *finite number* $n$ of *observed values* $z_i$ according to what they are. In the density plot, we're sorting and stacking up *all* of the *outcomes* $\omega$ in the silo according to the values they are destined to become, and we're measuring their relative volume, not absolute counts.
#
# In a nutshell, the histogram is what we can actually observe, given outputs from the factory. But the density descibes the inner workings of the factory itself, which we can never actually observe.
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# What do you expect to happen to the shape of the histogram above as the number of observations is increased from $1000$ to larger and larger numbers? Can you provide an intuitive explanation for why this happens?
#
# </div>
# ### Common Distributions
# We've seen that, for all practical purposes, a random variable is determined by its probability distribution (mass or density function). In reality, the distribution of any particular measurement (e.g. blood pressure) is unknown- it depends on a complex web of causal factors. The true density function is almost certainly so complex it's not even something that we could write down. But, for the purposes of *modeling* that measurement, we *pretend* that the density is something we can write down.
#
# Over the centuries, people have come up with a lot of distributions that are useful as models across various scenarios. Here are a few of them:
# #### Bernoulli distribution
# Let's say we're interested in modeling the result of a coin flip. The actual value (heads/tails, which we code as 0/1) of the coin flip is determined by some insanely complicated physics, but we're going to pretend that the value comes out of a little factory called $\mathbf Z$ that has the following probability mass function:
#
#
# $$
# P(\mathbf Z=z) =
# \begin{cases}
# 1/2 & \text{for }z=0 \\
# 1/2 & \text{for }z=1
# \end{cases}
# $$
# If we want to model a biased coin that comes up heads $(p\times100)$% of the time, we can use a mass function like:
# $$
# P(\mathbf Z=z) =
# \begin{cases}
# p & \text{for }z=0 \\
# 1-p & \text{for }z=1
# \end{cases}
# $$
# This is often written as $\mathbf Z \sim \text{Bernoulli}(p)$ (read: "$\mathbf Z$ is Bernoulli-distributed"). The number $p$ is said to be a **parameter** of the Bernoulli distribution. It would be more accurate to say that a random variable is distributed as a **member** of the Bernoulli **family** of distributions, since, technically, every different value of $p$ encodes a different distribution, or factory, for making data.
#
# Another way to think about it is that there's one data factory, but it has a control panel with a knob labeled "$p$". If $p$ is set to 0.7, we expect about 70% of the outputs to be 1. If $p$ is set to $0.1$, 10%, and so on. It's a matter of semantics whether or not you want to say that factory is representing two different factories, or merely one factory under two different operating conditions. Both perspectives are useful.
# #### Normal Distribution
#
# Let's say we want to model the heights of everyone on Earth. We have an intuition that people are typically a bit shorter than two meters, and taller and shorter people are more and more rare the taller and shorter they get. We can pretend that height measurements come from a **normal** distribution (also called **Gaussian** distribution):
# $$
# \phi(z) =
# \frac{1}{\sqrt{2\pi\sigma}}
# e^{-\frac{(x-\mu)^2}{2\sigma^2}}
# $$
# Most often you'll see this written as $\mathbf Z \sim \mathcal N(\mu, \sigma)$ (read: "$\mathbf Z$ is normally distributed"). The numbers $\mu$ and $\sigma$ are the parameters (control knobs) of the normal distribution.
#
# 
# As you can see in the picture, $\mu$ controls where the "bell curve" is centered and $\sigma$ controls how wide or narrow it is.
# <div class="alert alert-block alert-warning">
# <b>Note:</b>
#
# Every distribution is defined by its mass or density function $\phi$. The mass or density is often a complicated function, so instead of saying someting like "$\phi(z) = \frac{1}{\sqrt{2\pi\sigma}} e^{-\frac{(x-\mu)^2}{2\sigma^2}}$" every time we want a normally-distributed variable, we'll abbreviate that to "$\mathbf Z \sim \mathcal N (\mu, \sigma)$". But they mean the same thing.
#
# Every time you see something like $\mathbf Z \sim \mathcal D(\theta_1, \theta_2, \dots)$, just know there is some mass or density function that is associated with the name $\mathcal D$ and which has parameters $\theta_1, \theta_2, \dots$. You can always look it up if you need to know exactly what it is.
#
# </div>
# #### Others
# There are [hundreds](https://upload.wikimedia.org/wikipedia/commons/7/74/Normal_Distribution_PDF.svg) of well-studied distributions available to choose from when modeling. The most important thing to know about a distribution is what values it can generate. This is sometimes called the **support** of the distribution, since if you were to make a density or mass plot, the support would be the region of the x-axis that has positive density or mass, so it's the region that appears to be "supporting" the curve or mass.
#
# For example, varaibles that are normally-, Cauchy-, or Laplace-distributed are supported on $-\infty$ and $\infty$. The $\chi^2$ distribution has support on $[0,\infty)$. The beta and standard uniform distributions have support on $[0,1]$. The Poisson distribution has support on the counting numbers 0, 1, 2..., and the K-categorical distribution has support on a finite number of integers 0, 1, 2, ... K.
#
# It's also totally possible to invent your own distribution by defining your own support set $S$ and mass/density function $\phi$, as long as $\phi(s) \ge 0$ for all $s \in S$ and $\int_S \phi(s) ds = 1$ or $\sum_{s \in S} \phi(s) = 1$. These properties have to be satisfied to have a valid density or mass (see exercise in previous section).
#
# The point of this diversity is that it is possible to model different kinds of data. Apartment rents are always positive numbers, but theoretically unbounded above (a scary thought), so perhaps $\chi^2$ is a good choice. The number of cars that pass through an intersection in a given day is always an integer, so Poisson is a reasonable choice for that. You don't have to remember any of these specific distributions or examples- just know there are many preconstructed pretend data factories out there to play with. Also know that the real data-generating process is pretty much *never* actually one of these distributions, although, sometimes, it might be well-approximated by one.
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
#
# Define your own density function that has support on $[0,1]$. Make it so that the probability of getting a bigger number is bigger than that of getting a smaller number. Be sure that your function integrates to $1$ and is nonnegative over its support, otherwise it's not a valid density.
#
# </div>
# ## Chapter summary
# Data in the real world is generated by complex processes that we can't ever hope to replicate. But if we want to uncover relationships between measurements, we at least need a framework for imagining what kinds of processes might be generating our data. Random variables and probability theory do that for us.
#
# Random variables are like factories that generate data. We don't observe them directly, but we see the data they output and we can imagine different kinds of random variables that make different kinds of data. We defined a notion of probability that posits that the probability of observing a particular realization is actually just the volume of material in the factory's silo (sample space) that is destined to become that realization. This is a pure abstraction, but it turns out to capture relationships between probabilities that we would intuitively expect to hold.
#
# It's easier to work with the probability distribution of a random variable than it is to constantly talk about the sample space and the mapping between that space and realizations. The probability distribution is a function that, when integrated over a region of the space of realizations, gives us the volume of outcomes in the sample space that map to realizations in that region. In other words: the probability that the random variable gives a realization in that region. Random variables can be continuous or discrete, but all have a distribution function that can be integrated or summed to yield probabilities.
#
# Random variables are most often talked about in terms of their porbability distributions. Defining a new variable is as easy as choosing a support and a mass or density function over that support. Some distributions are so commonly used that they have their own names and notations so that we don't have to write out their mass or density functions out over and over again to refer to them.
|
docs/content/probability/1_univariate_prob.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="dcQ6l8DrzJad"
# ##### Copyright 2020 Google
# + cellView="form" colab={} colab_type="code" id="7GiJJW7KzLLA"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="Iznv7pW1yzWA"
# # QAOA Tasks
# + [markdown] colab_type="text" id="UX6Wn1flzYOg"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.example.org/cirq/research/qaoa/tasks"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on QuantumLib</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/quantumlib/ReCirq/blob/master/docs/qaoa/tasks.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/quantumlib/ReCirq/blob/master/docs/qaoa/tasks.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/ReCirq/docs/qaoa/tasks.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="ODiWjvdR0A-c"
# ## Setup
#
# Install the ReCirq package:
# + colab={} colab_type="code" id="CNcvxXPs0Buy"
try:
import recirq
except ImportError:
# !pip install git+https://github.com/quantumlib/ReCirq
# + [markdown] colab_type="text" id="7EyHaU6p0Ebv"
# And import ReCirq:
# + colab={} colab_type="code" id="hJC1Mqt00Txt"
import recirq
# + [markdown] colab_type="text" id="L2vg1-UXyzWC"
# ## Problem generation
#
# First, we generate and save all of the random instances of the problem. This is not computationally intensive but very important to do first so we have a fixed set of random instances.
# + colab={} colab_type="code" id="_ljoY33fyzWD"
from recirq.qaoa.experiments.problem_generation_tasks import \
SKProblemGenerationTask, HardwareGridProblemGenerationTask, ThreeRegularProblemGenerationTask, \
generate_3_regular_problem, generate_sk_problem, generate_hardware_grid_problem
pgen_dataset_id = '2020-03-tutorial'
hardware_grid_problem_tasks = [
HardwareGridProblemGenerationTask(
dataset_id=pgen_dataset_id,
device_name='Sycamore23',
instance_i=i,
n_qubits=n
)
for i in range(5)
for n in range(2, 8 + 1, 2)
]
recirq.display_markdown_docstring(HardwareGridProblemGenerationTask)
# + colab={} colab_type="code" id="1o5-pazdyzWH"
sk_problem_tasks = [
SKProblemGenerationTask(
dataset_id=pgen_dataset_id,
instance_i=i,
n_qubits=n
)
for i in range(5)
for n in range(3, 7 + 1, 2)
]
recirq.display_markdown_docstring(SKProblemGenerationTask)
# + colab={} colab_type="code" id="qyiL_W_KyzWK"
three_regular_problem_tasks = [
ThreeRegularProblemGenerationTask(
dataset_id=pgen_dataset_id,
instance_i=i,
n_qubits=n
)
for i in range(5)
for n in range(3, 8 + 1) if 3 * n % 2 == 0
]
# + [markdown] colab_type="text" id="bMBE4_hWyzWO"
# ### Run the tasks
# + colab={} colab_type="code" id="EYmmjjRnyzWO"
for task in hardware_grid_problem_tasks:
generate_hardware_grid_problem(task)
for task in sk_problem_tasks:
generate_sk_problem(task)
for task in three_regular_problem_tasks:
generate_3_regular_problem(task)
# + [markdown] colab_type="text" id="JxNchpctyzWR"
# ## Angle precomputation
# + colab={} colab_type="code" id="G0eSk61UyzWR"
from recirq.qaoa.experiments.angle_precomputation_tasks import \
AnglePrecomputationTask, precompute_angles
apre_dataset_id = '2020-03-tutorial'
precompute_tasks = [
AnglePrecomputationTask(
dataset_id=apre_dataset_id,
generation_task=gen_task,
p=p)
for gen_task in recirq.roundrobin(
hardware_grid_problem_tasks,
sk_problem_tasks,
three_regular_problem_tasks,
)
for p in range(1, 3 + 1)
]
recirq.display_markdown_docstring(AnglePrecomputationTask)
# + colab={} colab_type="code" id="9TICgyjByzWU"
for task in precompute_tasks:
precompute_angles(task)
# + [markdown] colab_type="text" id="U-4ZhSe_yzWX"
# ## Precomputed angle data collection
# + colab={} colab_type="code" id="yxAV6GjWyzWY"
from recirq.qaoa.experiments.precomputed_execution_tasks import \
PrecomputedDataCollectionTask, collect_data
dcol_dataset_id = '2020-03-tutorial'
data_collection_tasks = [
PrecomputedDataCollectionTask(
dataset_id=dcol_dataset_id,
precomputation_task=pre_task,
device_name='Syc23-simulator',
n_shots=50_000,
structured=True,
)
for pre_task in precompute_tasks
]
recirq.display_markdown_docstring(PrecomputedDataCollectionTask)
# + colab={} colab_type="code" id="F5oIYM1GyzWb"
await recirq.execute_in_queue(collect_data, data_collection_tasks, num_workers=2)
# + [markdown] colab_type="text" id="ZR4uOP5SyzWe"
# ## Landscape data collection
# + colab={} colab_type="code" id="9fx1_PF5yzWe"
from recirq.qaoa.experiments.p1_landscape_tasks import \
P1LandscapeDataCollectionTask, \
get_data_collection_tasks_on_a_grid, \
collect_either_landscape_or_cal
recirq.display_markdown_docstring(P1LandscapeDataCollectionTask)
# + colab={} colab_type="code" id="XlcHhXUwyzWh"
hardware_grid_problem_task = HardwareGridProblemGenerationTask(
dataset_id=pgen_dataset_id,
device_name='Sycamore23',
instance_i=0,
n_qubits=4
)
data_collection_tasks = get_data_collection_tasks_on_a_grid(
pgen_task=hardware_grid_problem_task,
dataset_id=dcol_dataset_id,
gamma_res=11,
beta_res=11,
device_name='Syc23-simulator',
epoch="grid")
await recirq.execute_in_queue(collect_either_landscape_or_cal,
data_collection_tasks,
num_workers=2)
# + colab={} colab_type="code" id="FTxZFYPLyzWj"
sk_problem_task = SKProblemGenerationTask(
dataset_id=pgen_dataset_id,
instance_i=0,
n_qubits=3,
)
data_collection_tasks = get_data_collection_tasks_on_a_grid(
pgen_task=sk_problem_task,
dataset_id=dcol_dataset_id,
gamma_res=11,
beta_res=11,
device_name='Syc23-simulator',
epoch="sk")
await recirq.execute_in_queue(collect_either_landscape_or_cal,
data_collection_tasks,
num_workers=2)
# + colab={} colab_type="code" id="Rq1ePbzQyzWm"
three_regular_problem_task = ThreeRegularProblemGenerationTask(
dataset_id=pgen_dataset_id,
instance_i=0,
n_qubits=4
)
data_collection_tasks = get_data_collection_tasks_on_a_grid(
pgen_task=three_regular_problem_task,
dataset_id=dcol_dataset_id,
device_name='Syc23-simulator',
gamma_res=11,
beta_res=11,
epoch="tr")
await recirq.execute_in_queue(collect_either_landscape_or_cal,
data_collection_tasks,
num_workers=2)
# -
# ## Optimization Data Collection
# +
from recirq.qaoa.experiments.optimization_tasks import \
OptimizationAlgorithm, \
OptimizationTask, \
collect_optimization_data
recirq.display_markdown_docstring(OptimizationTask)
# +
optimization_algorithm = OptimizationAlgorithm(
method='MGD',
n_shots=25000,
options={
'max_iterations': 10,
'rate': 0.3,
'sample_radius': 0.1,
'n_sample_points_ratio': 1.0,
'rate_decay_exponent': 0.4,
'stability_constant': 250,
'sample_radius_decay_exponent': 0.08,
})
hardware_grid_optimization_task = OptimizationTask(
dataset_id=dcol_dataset_id,
generation_task=hardware_grid_problem_task,
device_name='Syc23-simulator',
p=1,
algorithm=optimization_algorithm,
x0=[0.3, 0.2])
collect_optimization_data(hardware_grid_optimization_task)
# +
optimization_algorithm = OptimizationAlgorithm(
method='MGD',
n_shots=25000,
options={
'max_iterations': 10,
'rate': 0.3,
'sample_radius': 0.1,
'n_sample_points_ratio': 1.0,
'rate_decay_exponent': 0.3,
'stability_constant': 200,
'sample_radius_decay_exponent': 0.08,
})
sk_optimization_task = OptimizationTask(
dataset_id=dcol_dataset_id,
generation_task=sk_problem_task,
device_name='Syc23-simulator',
p=1,
algorithm=optimization_algorithm,
x0=[0.3, 0.2])
collect_optimization_data(sk_optimization_task)
# +
optimization_algorithm = OptimizationAlgorithm(
method='MGD',
n_shots=25000,
options={
'max_iterations': 10,
'rate': 0.2,
'sample_radius': 0.1,
'n_sample_points_ratio': 1.0,
'rate_decay_exponent': 0.4,
'stability_constant': 250,
'sample_radius_decay_exponent': 0.08,
})
three_regular_optimization_task = OptimizationTask(
dataset_id=dcol_dataset_id,
generation_task=three_regular_problem_task,
device_name='Syc23-simulator',
p=1,
algorithm=optimization_algorithm,
x0=[0.3, 0.2])
collect_optimization_data(three_regular_optimization_task)
|
docs/qaoa/tasks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from __future__ import print_function
from statsmodels.compat import lzip
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import statsmodels.api as sm
from statsmodels.formula.api import ols
beef = pd.read_csv('beef.csv')
beef.head(10)
beef_model = ols("Quantity ~ Price", data=beef).fit()
print(beef_model.summary())
fig = plt.figure(figsize=(12,8))
fig = sm.graphics.plot_partregress_grid(beef_model, fig=fig)
fig = plt.figure(figsize=(12, 8))
fig = sm.graphics.plot_ccpr_grid(beef_model, fig=fig)
fig = plt.figure(figsize=(12,8))
fig = sm.graphics.plot_regress_exog(beef_model, 'Price', fig=fig)
beef['Year'] = pd.to_datetime(beef['Year'], format="%Y")
from pandas.tseries.offsets import *
beef['Date'] = beef.apply(lambda x:(x['Year'] + BQuarterBegin(x['Quarter'])), axis=1)
beef.drop(['Year', 'Quarter'], axis=1, inplace=True)
beef.set_index('Date', inplace=True)
beef.head(10)
endog = beef['Quantity']
exog = sm.add_constant(beef['Price'])
mod = sm.RecursiveLS(endog, exog)
res = mod.fit()
print(res.summary())
res.plot_recursive_coefficient(range(mod.k_exog), alpha=None, figsize=(10,6));
fig = res.plot_cusum(figsize=(10,6));
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Filters
#
# [![open_in_colab][colab_badge]][colab_notebook_link]
# [![open_in_binder][binder_badge]][binder_notebook_link]
#
# [colab_badge]: https://colab.research.google.com/assets/colab-badge.svg
# [colab_notebook_link]: https://colab.research.google.com/github/UnfoldedInc/examples/blob/master/notebooks/05%20-%20Filters.ipynb
# [binder_badge]: https://mybinder.org/badge_logo.svg
# [binder_notebook_link]: https://mybinder.org/v2/gh/UnfoldedInc/examples/master?urlpath=lab/tree/notebooks/05%20-%20Filters.ipynb
# This example shows how to control map filters and listen to map filter change events with Unfolded Map SDK.
# ## Dependencies
#
# This notebook requires the following Python dependencies:
#
# - `unfolded.map-sdk`: The Unfolded Map SDK
# - `pandas`: DataFrame library
#
# If running this notebook in Binder, these dependencies should already be installed. If running in Colab, the next cell will install these dependencies.
# If in Colab, install this notebook's required dependencies
import sys
if "google.colab" in sys.modules:
# !pip install 'unfolded.map_sdk>=0.6.0' pandas
# ## Imports
from unfolded.map_sdk import UnfoldedMap
import pandas as pd
import ipywidgets as widgets
# ## Using Map Filters
# Let's again create a local map and add data to it:
unfolded_map = UnfoldedMap()
unfolded_map
url = 'https://raw.githubusercontent.com/UnfoldedInc/examples/master/notebooks/data/earthquakes.csv'
df = pd.read_csv(url)
unfolded_map.add_dataset({
'label': 'Earthquakes',
'data': df
})
# ## Adding a filter
# Say, we want to filter the data points by the `Magnitude` column. Let's first find out what's the extent of the values in this column:
magnitude_extent = [df['Magnitude'].min(), df['Magnitude'].max()]
magnitude_extent
# Now we can set the filter to only show the points in the top half of the range:
unfolded_map.set_filter({
'id': 'magnitude_filter',
'field': 'Magnitude',
'value': [
(magnitude_extent[1] + magnitude_extent[0])/2,
magnitude_extent[1]
]
})
# Or the bottom half:
unfolded_map.set_filter({
'id': 'magnitude_filter',
'field': 'Magnitude',
'value': [
magnitude_extent[0],
(magnitude_extent[1] + magnitude_extent[0])/2
]
})
# ## Controlling the filter from the notebook
# We can use the range slider from `ipywidgets` to control the filter in the map. First, we create the slider:
slider = widgets.FloatRangeSlider(
value=magnitude_extent,
min=magnitude_extent[0],
max=magnitude_extent[1],
step=0.1,
description='Magnitude:',
continuous_update=True
)
# Here we create an event handler to listen to the slider change events:
def update_value_filter(change):
if 'new' in change and 'value' in change['new']:
unfolded_map.set_filter({
'id': 'magnitude_filter',
'field': 'Magnitude',
'value': change['new']['value']
})
slider.observe(update_value_filter)
slider
# Now try moving the slider. You should see the changes applied to the map.
# ## Syncing the slider in the notebook with the map filter
# We can register [event handlers](https://docs.unfolded.ai/map-sdk/api#events) to be notified of filter changes in the map. Here's how we can synchronize the above range slider with the Magnitude filter in the map:
# +
def on_filter_sync(info):
if 'Magnitude' in info['name'] and info['prop'] == 'value':
slider.value = info['value']
unfolded_map.set_map_event_handlers({
'on_filter': on_filter_sync
})
# -
# Now try changing the "Magnitude" filter in the "Filter" pane of the left sidebar in the map. You should see the slider above in the notebook update.
# The following will unregister the observer:
slider.unobserve(None)
# ## Debugging an event handler
# Here's how you can output the event info when filter events are triggered:
output = widgets.Output()
@output.capture(clear_output=True)
def on_filter_output(info):
print(info)
output
unfolded_map.set_map_event_handlers({
'on_filter': on_filter_output
})
# Now when you change the filter in the Unfolded Studio map, you'll see above a printed object like:
# ```py
# {'dataId': '0c6307ea-f5a1-4509-a1ef-cd60d7663010', 'name': ['Magnitude'], 'value': [3.04, 5.23], 'prop': 'value', 'type': 'range', 'id': 'magnitude_filter'}
# ```
|
notebooks/05 - Filters.ipynb
|