code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 太陽と地球と月
# 太陽と公転する地球、月を描画するコードです。
# それっぽく見えるだけで実態は円の公式に沿った描画を行います。
# 動作確認済み環境は以下の通り。
#
# 項目|バージョン
# ---|---
# OS|Mac OSX 10.10.5(Yosemite)
# Python|3.5.1
# matplotlib|1.5.1
# numpy|1.11.0
# ipython|4.1.2
# ## 必要なライブラリのインポート
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import animation
from IPython.display import HTML,display
import math
# -
# ## プロット用のオブジェクトの作成
# 動画は基本的にパラパラ漫画なので、動かすグラフは前もってインスタンスを生成し、
# 更新用のメソッド内に渡してあげる必要があります。
# +
# プロット用のオブジェクトを作成
fig, ax = plt.subplots()
plt.close()
# 描画領域の縦、横のサイズを指定
fig.set_figheight(6.0)
fig.set_figwidth(6.0)
# X軸を-1.5〜1.5までに設定
ax.set_xlim((-1.5, 1.5))
# Y軸を-1.5〜1.5までに設定
ax.set_ylim((-1.5, 1.5))
# 地球描画用のオブジェクトを作成
earth = ax.scatter([],[], color='blue',marker='o',s=80)
# 月描画用のオブジェクトを作成
moon = ax.scatter([],[], color='gray',marker='o',s=40)
# -
# ## 太陽の描画
# 太陽自体は動画の中では動かすものではないので、
# ここで先に描画して、背景のように扱います。
ax.scatter(0,0,color='red',marker='o',s=320)
# ## 更新用メソッドの定義
# 円の方程式について中心が原点で半径がrの場合は以下の通りとなります。
# \begin{eqnarray}
# x^2+y^2&=&r^2\\
# x&=&r\cos \theta\\
# y&=&r\sin \theta\\
# \end{eqnarray}
# 上記で地球の公転がか描画できます。更に、中心が原点から離れている場合は以下の通りとなります。
# \begin{eqnarray}
# (x-a)^2+(y-b)^2&=&r^2\\
# x&=&a+r\cos \theta\\
# y&=&b+r\sin \theta\\
# \end{eqnarray}
# $a$と$b$を以下のように移動するような値$\cos$や$\sin$にすることで月が地球を公転しながら太陽の周りも回るような描画が可能となります。
# \begin{eqnarray}
# (x-a)^2+(y-b)^2&=&r^2\\
# x&=&\cos \theta+r\cos \theta\\
# y&=&\sin \theta+r\sin \theta\\
# \end{eqnarray}
# また$\theta$の値を増加させることで公転速度が速くなるような描画可能となります。
# コールバックメソッドの定義
def animate(i,earth,moon):
# フレームからラジアンを算出
# ラジアンの算出式は「度数 × 円周率 ÷ 180」
x = i * math.pi / 180
# フレームに対してxの移動を2倍速にする
x = 2 * x
# 地球の描画
earth_x = math.cos(x)
earth_y = math.sin(x)
earth.set_offsets((earth_x, earth_y))
# 月の描画
moon_x = math.cos(x) + 1/8 * math.cos(4 * x)
moon_y = math.sin(x) + 1/8 * math.sin(4 * x)
moon.set_offsets((moon_x,moon_y))
# ## アニメーションの生成
# ここでFuncAnimationメソッドを利用してアニメーションを生成します。
# メソッドの引数は以下の通りです。
#
# 引数|意味
# ---|---
# fig|描画領域
# func|フレーム毎に更新する描写を定義したコールバック関数
# fargs|更新するグラフの関数
# frames|描画全体のフレーム数
# interval|何ミリ秒毎に再描画するか(アニメーションのコマを進めるか)
# +
# 動画全体のフレームを設定する
FRAMES = 180
anim = animation.FuncAnimation(fig = fig, func = animate, fargs = (earth,moon), frames = FRAMES, interval=30)
# -
# ## アニメーションの出力
# アニメーションの出力先によって方法が異なります。以下の通り
#
# 出力先|方法
# ---|---
# IPython内に表示する|HTMLクラスを利用する
# MP4やGIF等ファイルに出力する|Writerクラスを利用する
#
# また、Writerクラスをインスタンス化する際の引数は以下の通りです。
#
# 引数|意味
# ---|---
# fps|フレームレート(ほとんどの動画サイトは30fps以下であればアップロードできる。<br>また、そんなにないと思うけれどDVD化等に動画を出力する際は<br>29.97fpsにする必要がある。そのため、ここでは29.97に設定)
# metadata|動画作成者等のメタデータを設定
# bitrate|動画のビットレート(画質)
# +
export_to = "IPython"
#export_to = "MP4"
#export_to = "GIF"
title = "sun_earth_moon"
if export_to == "IPython":
# HTML化することで描画
display(HTML(anim.to_html5_video()))
elif export_to == "MP4":
# FFmpegというUNIX系の動画変換ソフトを利用する
Writer = animation.writers['ffmpeg']
# エクスポートされるファイルの情報を定義してインスタンス化する
writer = Writer(fps=29.97, metadata=dict(artist='Me'), bitrate=1800)
# 実際の動画のファイルエスクスポート
anim.save(title + '.mp4', writer=writer)
elif export_to == "GIF":
# FFmpegというUNIX系の動画変換ソフトを利用する
Writer = animation.writers['imagemagick']
# エクスポートされるファイルの情報を定義してインスタンス化する
writer = Writer(fps=29.97, metadata=dict(artist='Me'), bitrate=1800)
# 実際の動画のファイルエスクスポート
anim.save(title + '.gif', writer=writer)
| 3.sun_earth_moon_using_FuncAnimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings('ignore')
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt #data visualization
import seaborn as sns
from operator import add
# %matplotlib inline
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
#load the data
data = pd.read_csv('/kaggle/input/framingham-heart-study-dataset/framingham.csv')
data.drop(['education'],axis=1,inplace=True) #dropping the education column because it has no correlation with heart disease
data.head()
# # Data Analysis
# * Missing variables
# Handling missing data is important as many machine learning algorithms do not support data with missing values.
#total percentage of missing data
missing_data = data.isnull().sum()
total_percentage = (missing_data.sum()/data.shape[0]) * 100
print(f'The total percentage of missing data is {round(total_percentage,2)}%')
# Since the missing entries account for only 12% of the total data we can drop these entries without losing alot of data.
# drop missing entries
data.dropna(axis=0, inplace=True)
data.shape
# ## Data Distribution
# plot histogram to see the distribution of the data
fig = plt.figure(figsize = (15,20))
ax = fig.gca()
data.hist(ax = ax)
plt.show()
# ___The data on the prevalent stroke, diabetes, and blood pressure meds are poorly balanced___
# ## Case counts
sns.countplot(x='TenYearCHD',data=data)
plt.show()
cases = data.TenYearCHD.value_counts()
print(f"There are {cases[0]} patients without heart disease and {cases[1]} patients with the disease")
# _The data is not properly balanced as the number of people without the disease greately exceeds the number of people with the disease._
def stacked_barchart(data, title = None, ylabel = None, xlabel = None):
default_colors = ['#0affff', '#ffff0a', '#ff0aff']
# From raw value to percentage
totals = data.sum(axis=1)
bars = ((data.T / totals) * 100).T
r = list(range(data.index.size))
# Plot
barWidth = 0.95
names = data.index.tolist()
bottom = [0] * bars.shape[0]
# Create bars
color_index = 0
plots = []
for bar in bars.columns:
plots.append(plt.bar(r, bars[bar], bottom=bottom, color=default_colors[color_index], edgecolor='black', width=barWidth))
bottom = list(map(add, bottom, bars[bar]))
color_index = 0 if color_index >= len(default_colors) else color_index + 1
# Custom x axis
plt.title(title)
plt.xticks(r, names)
plt.xlabel(data.index.name if xlabel is None else xlabel)
plt.ylabel(data.columns.name if ylabel is None else ylabel)
ax = plt.gca()
y_labels = ax.get_yticks()
ax.set_yticklabels([str(y) + '%' for y in y_labels])
flat_list = [item for sublist in data.T.values for item in sublist]
for i, d in zip(ax.patches, flat_list):
data_label = str(d) + " (" + str(round(i.get_height(), 2)) + "%)"
ax.text(i.get_x() + 0.45, i.get_y() + 5, data_label, horizontalalignment='center', verticalalignment='center', fontdict = dict(color = 'black', size = 20))
for item in ([ax.title]):
item.set_fontsize(27)
for item in ([ax.xaxis.label, ax.yaxis.label] + ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(24)
legend = ax.legend(plots, bars.columns.tolist(), fancybox=True)
plt.setp(legend.get_texts(), fontsize='20')
# +
fig = plt.gcf()
fig.set_size_inches(25, 35)
grid_rows = 3
grid_cols = 2
#draw sex vs disease outcome
plt.subplot(grid_rows, grid_cols, 1)
temp = data[['male','TenYearCHD']].groupby(['male','TenYearCHD']).size().unstack('TenYearCHD')
temp.rename(index={0:'Female', 1:'Male'}, columns={0:'No Disease', 1:'Has Disease'}, inplace = True)
stacked_barchart(temp, title = 'CHD vs Sex', ylabel = 'Population')
#draw smoking satus vs disease outcome
plt.subplot(grid_rows, grid_cols, 2)
temp = data[['currentSmoker','TenYearCHD']].groupby(['currentSmoker','TenYearCHD']).size().unstack('TenYearCHD')
temp.rename(index={0:'Not a Smoker', 1:'Smoker'}, columns={0:'No Disease', 1:'Has Disease'}, inplace = True)
stacked_barchart(temp, title = 'CHD vs Smoking', ylabel = 'Population')
#draw diabetes vs disease outcome
plt.subplot(grid_rows, grid_cols, 3)
temp = data[['diabetes','TenYearCHD']].groupby(['diabetes','TenYearCHD']).size().unstack('TenYearCHD')
temp.rename(index={0:'Not Diabetic', 1:'Diabetic'}, columns={0:'No Disease', 1:'Has Disease'}, inplace = True)
stacked_barchart(temp, title = 'CHD vs Diabetes', ylabel = 'Population')
#draw BP meds vs disease outcome
plt.subplot(grid_rows, grid_cols, 4)
temp = data[['BPMeds','TenYearCHD']].groupby(['BPMeds','TenYearCHD']).size().unstack('TenYearCHD')
temp.rename(index={0:'Not on medication', 1:'On Medication'}, columns={0:'No Disease', 1:'Has Disease'}, inplace = True)
stacked_barchart(temp, title = 'CHD vs BP meds', ylabel = 'Population')
#draw Hypertension vs disease outcome
plt.subplot(grid_rows, grid_cols, 5)
temp = data[['prevalentHyp','TenYearCHD']].groupby(['prevalentHyp','TenYearCHD']).size().unstack('TenYearCHD')
temp.rename(index={0:'Not Hypertensive', 1:'Hypertensive'}, columns={0:'No Disease', 1:'Has Disease'}, inplace = True)
stacked_barchart(temp, title = 'CHD vs Hypertension', ylabel = 'Population')
# -
# ___conclusions:___
# * Slightly more males are suffering from CHD than females
# * The percentage of people who have CHD is almost equal between smokers and non smokers
# * The percentage of people who have CHD is higher among the diabetic, and those with prevalent hypertesion as compared to those who dont have similar morbidities
# * A larger percentage of the people who have CHD are on blood pressure medication
# ## Number of people who have disease vs age
positive_cases = data[data['TenYearCHD'] == 1]
plt.figure(figsize=(15,6))
sns.countplot(x='age',data = positive_cases, hue = 'TenYearCHD', palette='husl')
plt.show()
# The number of sick people generally increases with age (highest risk being in between the ages of 51 and 63)
# ## Correlation Heat map
plt.figure(figsize=(15,8))
sns.heatmap(data.corr(), annot = True)
plt.show()
# There are no features with more than 0.5 correlation with the Ten year risk of developing CHD and this shows that the features a poor predictors. However the features with the highest correlations are age, prevalent hypertension and systolic blood pressure
# # Feature Selection
# Since having irrelevant features in a data set can decrease the accuracy of the models applied, I used the Boruta Feature Selection technique (wrapper built around the random forest classification algorithm) to select the most important features which were later used to build different models.
#
# ___Methodology:___
# * Firstly, it adds randomness to the given data set by creating shuffled copies of all features (which are called shadow features).
#
# * Then, it trains a random forest classifier on the extended data set and applies a feature importance measure (the default is Mean Decrease Accuracy) to evaluate the importance of each feature where higher means more important.
#
# * At every iteration, it checks whether a real feature has a higher importance than the best of its shadow features (i.e. whether the feature has a higher Z-score than the maximum Z-score of its shadow features) and constantly removes features which are deemed highly unimportant.
#
# * Finally, the algorithm stops either when all features get confirmed or rejected or it reaches a specified limit of random forest runs.
from sklearn.ensemble import RandomForestClassifier
from boruta import BorutaPy
# +
#define the features
X = data.iloc[:,:-1].values
y = data.iloc[:,-1].values
forest = RandomForestClassifier(n_estimators=1000, n_jobs=-1, class_weight='balanced')
# define Boruta feature selection method
feat_selector = BorutaPy(forest, n_estimators='auto', verbose=2)
# find all relevant features
feat_selector.fit(X, y)
# -
# show the most important features
most_important = data.columns[:-1][feat_selector.support_].tolist()
most_important
# We see that age and systolic blood pressures are selected as the most important features for predicting the Ten year risk of developing CHD. However we will use the six most important features to build our models to get more features for more precise prediction.
# select the top 6 features
top_features = data.columns[:-1][feat_selector.ranking_ <=6].tolist()
top_features
# The top features are:
# 1. Age
# 2. Total cholesterol
# 3. Systolic blood pressure
# 4. Diastolic blood pressure
# 5. BMI
# 6. Heart rate
# 7. Blood glucose
# ## Statistics on the top features
import statsmodels.api as sm
X_top = data[top_features]
y = data['TenYearCHD']
res = sm.Logit(y,X_top).fit()
res.summary()
params = res.params
conf = res.conf_int()
conf['Odds Ratio'] = params
conf.columns = ['5%', '95%', 'Odds Ratio']
print(np.exp(conf))
# # Balance the dataset using the Synthetic Minority Oversampling Technique (SMOTE).
# > SMOTE first selects a minority class instance a at random and finds its k nearest minority class neighbors. The synthetic instance is then created by choosing one of the k nearest neighbors b at random and connecting a and b to form a line segment in the feature space. The synthetic instances are generated as a convex combination of the two chosen instances a and b. — Page 47, Imbalanced Learning: Foundations, Algorithms, and Applications, 2013.
#
# This procedure can be used to create as many synthetic examples for the minority class as are required. It suggests first using random undersampling to trim the number of examples in the majority class, then use SMOTE to oversample the minority class to balance the class distribution.
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import RandomUnderSampler
from imblearn.pipeline import Pipeline
from collections import Counter
X = data[top_features]
y = data.iloc[:,-1]
# +
# the numbers before SMOTE
num_before = dict(Counter(y))
#perform SMOTE
# define pipeline
over = SMOTE(sampling_strategy=0.8)
under = RandomUnderSampler(sampling_strategy=0.8)
steps = [('o', over), ('u', under)]
pipeline = Pipeline(steps=steps)
# transform the dataset
X_smote, y_smote = pipeline.fit_resample(X, y)
#the numbers after SMOTE
num_after =dict(Counter(y_smote))
# -
print(num_before, num_after)
labels = ["Negative Cases","Positive Cases"]
plt.figure(figsize=(15,6))
plt.subplot(1,2,1)
sns.barplot(labels, list(num_before.values()))
plt.title("Numbers Before Balancing")
plt.subplot(1,2,2)
sns.barplot(labels, list(num_after.values()))
plt.title("Numbers After Balancing")
plt.show()
# ___The new dataset is much more balanced.___
# # Splitting data to Training and Testing set
# new dataset
new_data = pd.concat([pd.DataFrame(X_smote), pd.DataFrame(y_smote)], axis=1)
new_data.columns = ['age', 'totChol', 'sysBP', 'diaBP', 'BMI', 'heartRate', 'glucose','TenYearCHD']
new_data.head()
X_new = new_data[top_features]
y_new= new_data.iloc[:,-1]
X_new.head()
# split the dataset
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X_new,y_new,test_size=.2,random_state=42)
# ## Feature Scaling
from sklearn.preprocessing import StandardScaler
# +
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_train = pd.DataFrame(X_train_scaled)
X_test_scaled = scaler.transform(X_test)
X_test = pd.DataFrame(X_test_scaled)
# -
# # Models used
# ## Logistic Regression
# The goal of logistic regression is to find the best fitting (yet biologically reasonable) model to describe the relationship between the dichotomous characteristic of interest (dependent variable = response or outcome variable) and a set of independent (predictor or explanatory) variables. Logistic regression generates the coefficients (and its standard errors and significance levels) of a formula to predict a logit transformation of the probability of presence of the characteristic of interest.
# Rather than choosing parameters that minimize the sum of squared errors (like in ordinary regression), estimation in logistic regression chooses parameters that maximize the likelihood of observing the sample values.
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from sklearn.metrics import classification_report
from sklearn.metrics import recall_score,precision_score,classification_report,roc_auc_score,roc_curve
# search for optimun parameters using gridsearch
params = {'penalty':['l1','l2'],
'C':[0.01,0.1,1,10,100],
'class_weight':['balanced',None]}
logistic_clf = GridSearchCV(LogisticRegression(),param_grid=params,cv=10)
# +
#train the classifier
logistic_clf.fit(X_train,y_train)
logistic_clf.best_params_
# -
#make predictions
logistic_predict = logistic_clf.predict(X_test)
log_accuracy = accuracy_score(y_test,logistic_predict)
print(f"Using logistic regression we get an accuracy of {round(log_accuracy*100,2)}%")
cm=confusion_matrix(y_test,logistic_predict)
conf_matrix=pd.DataFrame(data=cm,columns=['Predicted:0','Predicted:1'],index=['Actual:0','Actual:1'])
plt.figure(figsize = (8,5))
sns.heatmap(conf_matrix, annot=True,fmt='d',cmap="YlGnBu")
print(classification_report(y_test,logistic_predict))
logistic_f1 = f1_score(y_test, logistic_predict)
print(f'The f1 score for logistic regression is {round(logistic_f1*100,2)}%')
# +
# ROC curve and AUC
probs = logistic_clf.predict_proba(X_test)
# keep probabilities for the positive outcome only
probs = probs[:, 1]
# calculate AUC
log_auc = roc_auc_score(y_test, probs)
# calculate roc curve
fpr, tpr, thresholds = roc_curve(y_test, probs)
# plot curve
sns.set_style('whitegrid')
plt.figure(figsize=(10,6))
plt.plot([0, 1], [0, 1], linestyle='--')
plt.plot(fpr, tpr, marker='.')
plt.ylabel('True positive rate')
plt.xlabel('False positive rate')
plt.title(f"AUC = {round(log_auc,3)}")
plt.show()
# -
my_data = pd.read_csv('../input/my-data/my_data.csv')
my_data
my_data=my_data[top_features]
my_data
prediction = logistic_clf.predict(my_data)
print("YOU REALLY ARE HEALTY !") if prediction[0] == 0 else print("YOU'RE AT RISK ! GET LIFE INSURANCE READY")
#
| Heart_disease_pridiction/heart-disease-prediction-final-version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Distance prediction with bouts: assess performance for all gases
# This notebook illustrates the performance of the distance prediction using bout events for all gases in the dataset from Vergara et al (2013).
#
# Generally, we use the lowest wind speed (fan set to 1500 RPM), taking signals from the board in the middle location (Board # 5), with sensor heaters set to 6V.
#
# For each gas, we perform the following steps:
#
# 1. Retrieve the data from the data set.
# 2. Identify the sensor which performs best in distance prediction.
# 3. Show the cross-validated performance of a linear regression using bout counts.
# 4. Visualise the relationship between bout counts and distance.
# +
import sys
import os
#add path to the directory containing the plumy module to PYTHONPATH
plumy_path = os.path.abspath(os.path.join(os.path.pardir, os.path.pardir))
sys.path.append(os.path.join(plumy_path))
toplevel_path = os.path.abspath(os.path.join(os.path.pardir, os.path.pardir, os.path.pardir))
import pickle
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
from tqdm.auto import tqdm
from plumy.utils import DataSelector
from plumy.utils import HDFDataSelector
from plumy.utils import ZipFileDataSelector
# + pycharm={"name": "#%%\n"}
rem_dupes = True # Drop duplicate timestamps
resample = True # Signal resampling active
# -
path = os.path.join(toplevel_path,'WTD_upload') # path to dataset
ds = DataSelector(path, drop_duplicates = rem_dupes, resample = resample, verbose = False, use_HDFcache=True)
path = os.path.join(toplevel_path, 'WTD_upload.zip')
dsz = ZipFileDataSelector(path, drop_duplicates = rem_dupes, resample = resample, verbose=False, use_HDFcache=True)
ds = dsz
path = os.path.join(toplevel_path, 'WTD_upload.zip_HDFcache')
dsh = HDFDataSelector(path, drop_duplicates = rem_dupes, resample = resample, verbose=False)
ds = dsh
plt.rc('text', usetex=False)
mpl.rcParams['savefig.dpi'] = 150 # for print, go to 600
from __future__ import unicode_literals
from plumy.bouts import *
def plot_boutcounts_vs_distance(boutcounts, pdists, model, ax=None):
"""
Helper function to generate a nice looking plot of the regression distance vs. bout counts.
Parameters:
boutcounts - M x N array of bout counts, for M distances and N trials each.
pdists - the distance values for the M distances
model - the regression model for the mean regression
ax - an axis to plot in. If None, create new figure.
Returns:
ax - the axis in which the content was plotted
"""
if ax is None:
f = plt.figure(figsize=(4,2.5))
ax = f.add_subplot(111)
plot_pdists=np.repeat(pdists[:,np.newaxis], boutcounts.shape[1], axis=1)
ax.plot(plot_pdists, boutcounts, ls='none', lw=0.3, marker='.',
ms=2., mfc='none', color='grey', zorder=1)
ax.plot(pdists, np.mean(np.array(boutcounts), axis=1), color=[0.,0.,0.,0.5],
ls='none', marker='o', markersize=6, zorder=3)
regression = (pdists - model.intercept_) / model.coef_.squeeze()
ax.plot(pdists, regression, color='k', linestyle='--', lw=1)
ax.set_frame_on(False)
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
xt = ["{:.2f}".format(d) for d in pdists]
if len(xt) > 5:
xt[4] = xt[4] + u" "
xt[5] = u" " + xt[5]
ax.set_xticks(pdists)
ax.set_xticklabels(xt)
ax.set_ylabel("number of bouts")
ax.set_xlabel("distance from source [m]")
ax.set_xlim(0.10, 1.50)
return ax
# List of Gases
for i,g in ds.GasNames.items():
print("{}: {}".format(i,g))
# 2 had issues with the MFC for some trials at L1
# 5 has no position 6
# 7 is corrupted for distance L2
sensornames = ["TGS2611", # Sensor 1
"TGS2612", # Sensor 2
"TGS2610", # Sensor 3
"TGS2602", # Sensor 4
"TGS2600a", # Sensor 5
"TGS2600b", # Sensor 6
"TGS2620a", # Sensor 7
"TGS2620b"] # Sensor 8
# use 6V heater voltage and 1500 RPM fan speed
voltage = 5
speed = 1
plot_data_dicts = [] # list to store the plot data dictionaries
multivar_results = [] # list to store results from multivariate regression
# ### Note on reproduction of the Figure
# **Recreating the whole analysis for this Figure takes quite some time** - ca. 30 minutes on my 2.7 GHz i7 Macbook Pro Retina.
#
# If you want to skip this lengthy step you can <a href="#Create-the-figure-with-saved-data">create the figure with cached data</a> (see below/follow the link).
# ### Acetaldehyde 500 ppm
gas = 1
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1] # +1 because sensor 1 is not in the ranks
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2)) #+2 because we don't look at sensor 1, and rank starts at 0
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
# clean up to save memory
del(data)
# ### Acetone 2500 ppm
# The data for Acetone 2500 ppm is partly corrupt - for P1 (distance 25 cm), trials 12 to 20 appear to bear no gas release due to a non-functional mass flow controller. This is also visible in the failure to detect significant bout counts in these trials. The data at the remaining positions is correct.
#
# For the sake of consistency, we replace the corrupt trials with randomly chosen trials from the same distance before performing the regression.
# +
gas = 2
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
print("### Warnings below indicate a non-functional mass flow controller.")
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# for 9 trials at L1 the MFC was not working at L1: trials [11, 12, 13, 14, 15, 16, 17, 18, 19]
# -
ebcs_acetone_2500 = make_boutcounters(data, sensorname="Sensor3", boardname="Board5", ampthresh=0.01)
boutcounts_acetone_2500 = calc_boutcounts(ebcs_acetone_2500)
non_func = np.nonzero(boutcounts_acetone_2500[0] <30)[0]
print("Trials at distance 0 with potentially non-functional MFC: {}".format(non_func))
print("Replacing these non-functional trials with random functional trials.")
func = np.nonzero(boutcounts_acetone_2500[0] >= 30)[0]
print(func)
np.random.shuffle(func)
print(func)
for i,nft in enumerate(non_func):
data[0][nft] = data[0][func[i]]
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del data
# ### Ammonia 10000 ppm
# +
gas = 3
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Benzene 200 ppm
# +
gas = 4
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Butanol 100 ppm
# +
gas = 5
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
# This dataset has no position 6. Therefore, iterate only to position 5.
for dist in tqdm(range(1,6)):
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Carbon Monoxide 1000 ppm
# This gas has been presented at a low concentration - the sensor responses are realtively noisy. See below for the same gas with a four-fold higher concentration.
#
# There are only 5 trials.
# +
gas = 6
trial = [1,2,3,4,5]
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed, trial))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Carbon Monoxide 4000 ppm
# For this gas the recordings taken on position 2 (50 cm from source) are corrupt - the source files contain only one line. We analyse only the other positions.
# +
gas = 7
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in [1,3,4,5,6]: # Distance 2 is corrupt
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Ethylene 500 ppm
# +
gas = 8
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Methane 1000 ppm
# +
gas = 9
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Methanol 200 ppm
# +
gas = 10
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# ### Toluene 200 ppm
# +
gas = 11
print("using Gas: {}, Voltage: {}, Fan Speed: {}.".format(
DataSelector.GasNames[gas],
DataSelector.SensorVoltages[voltage], DataSelector.AltFanSpeeds[speed]))
data = []
for dist in tqdm(range(1,7)):
data.append(ds.select(gas,dist,voltage,speed))
# -
# Pick best sensor
ebcs_all, rank, rmses = pick_best_sensor(data)
sensname = sensornames[rank[0]+1]
ebcs_best = ebcs_all[rank[0]]
print("\nUsing {} (sensor {}).".format(sensname, rank[0]+2))
boutcounts = calc_boutcounts(ebcs_best)
pdists = np.array([0.25, 0.5, 0.98, 1.18, 1.40, 1.45])
reg_mean = regress_mean(boutcounts, pdists=pdists)
reg_CV = regress_crossval_score(boutcounts, pdists)
ax = plot_boutcounts_vs_distance(boutcounts, pdists, model=reg_mean["mf"])
t = ax.set_title(" ".join(ds.GasNames[gas].split("_")) + " ppm, " + sensname, fontsize=10)
plot_data_dicts.append({'bc':boutcounts, 'pd':pdists, 'md':reg_mean, 'mcv':reg_CV, "sn":sensname})
# sensors 2-8, distances, trials
boutcounts_multivar = np.zeros((ebcs_all[0].shape[0],ebcs_all[0].shape[1],7))
for i,ebcs in enumerate(ebcs_all):
boutcounts_multivar[:, :, i] = calc_boutcounts(ebcs)
rd_mv = regress_crossval_score_multivar(boutcounts_multivar)
multivar_results.append({"mv":rd_mv, "single":rmses})
del(data)
# save the data needed to recreate the figures
with open("Figures/pdd_windspeed1_fullsensnames.pkl", 'wb') as du:
pickle.dump(plot_data_dicts, du)
# save the data needed to recreate the figures
with open("Figures/multivar_results_windspeed1.pkl", 'wb') as du:
pickle.dump(multivar_results, du)
# ### Create the figure with saved data
# +
#with open("Figures/pdd_windspeed1.pkl", 'r') as du:
with open("Figures/pdd_windspeed1_fullsensnames.pkl", 'rb') as du:
plot_data_dicts = pickle.load(du)
f = plt.figure(figsize=(11.5,8.5))
gs = mpl.gridspec.GridSpec(3,4, hspace=0.5, wspace=0.3, left=0.05, right=0.95, bottom=0.05, top=0.95)
xax = np.array([0.25, 0.50, 0.98, 1.18, 1.40, 1.45])
panels = ["A)", "B)", "C)", "D)", "E)", "F)", "G)", "H)", "I)", "J)", "K)", "L)" ]
for i,pdd in enumerate(plot_data_dicts):
ax = f.add_subplot(gs[i])
ax = plot_boutcounts_vs_distance(pdd['bc'], pdd['pd'], pdd['md']["mf"], ax)
t = ax.set_title(" ".join(ds.GasNames[i+1].split("_")) + " ppm, " + pdd["sn"], fontsize=10)
ax.set_xticks(xax)
ax.set_xticklabels(["0.25", "0.50", "0.98", "", "1.40", ""])
ax.set_ylim(0,75)
if not (i in [0,4,8]):
ax.set_xlabel('')
ax.set_ylabel('')
ax.set_yticklabels(["" for t in ax.get_yticklabels()])
ax.text(.5, 70,
"CV RMSE = {:.2f} ± {:.2f} m".format(np.mean(pdd['mcv']['RMSE']),
2*np.std(pdd['mcv']['RMSE'])),
fontsize=8)
ax.text(-0.1,1.15, panels[i], fontweight='bold', transform=ax.transAxes)
avg_rmse = np.mean([np.mean(pdd['mcv']['RMSE']) for pdd in plot_data_dicts])
print("Average RMSE over all gases: {:.2f} m".format(avg_rmse))
#relate molecular weight to slope of fit
mol_weights = {1:44.052560,
2:58.079140,
3:17.030520,
4:78.111840,
5:74.121600,
6:28.010100,
7:28.010100,
8:28.05316,
9:16.04246 ,
10:32.04186,
11:92.13842}
from scipy.stats import linregress, pearsonr
ax = f.add_subplot(gs[i+1])
reg_coefs = np.array([pdd['md']['mf'].coef_[0] for pdd in plot_data_dicts])
reg_slopes = 1./reg_coefs
mw = [mol_weights[i] for i in range(1,12)]
plotwhat = "mw"
if plotwhat == "sqrt_mw":
ax.plot(reg_slopes, np.sqrt(mw),'.k')
#move some name tags
for i in range(1,12):
if i == 6:
xof = 1
yof = 0.1
elif i == 7:
xof = 1
yof = -0.3
elif i == 9:
xof = -9
yof = -0.3
else:
xof = 1
yof = -0.15
ann = ax.annotate(ds.GasNames[i].split("_")[0], (reg_slopes[i-1]+xof, np.sqrt(mw[i-1])+yof),
fontsize=7)
ann.set_color('gray')
#regression slope vs. molecular weight
slope, intercept, r_value, p_value, std_err = linregress(reg_slopes, np.sqrt(mw))
regress_x = np.array([-45, -10])
regress_y = slope*regress_x + intercept
ax.plot(regress_x, regress_y, '--', color='gray', zorder=0)
#plot beauty
#ax.set_xlim(-.06, -.02)
ax.set_frame_on(False)
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.set_xlabel('regression slope')
ax.set_ylabel('sqrt(MW)')
ax.set_xticks([-50, -40, -30, -20, -10])
ax.set_ylim(3,10)
elif plotwhat == "mw":
ax.plot(reg_slopes, mw,'.k')
#move some name tags
for i in range(1,12):
if i == 6:
xof = 1
yof = 0.1**2
elif i == 7:
xof = 1
yof = -(0.3**2)
elif i == 9:
xof = -9
yof = -(0.3**2)
else:
xof = 1
yof = -(0.15**2)
ann = ax.annotate(ds.GasNames[i].split("_")[0], (reg_slopes[i-1]+xof, mw[i-1]+yof),
fontsize=7)
ann.set_color('gray')
#regression slope vs. molecular weight
slope, intercept, r_value, p_value, std_err = linregress(reg_slopes, mw)
regress_x = np.array([-45, -10])
regress_y = slope*regress_x + intercept
ax.plot(regress_x, regress_y, '--', color='gray', zorder=0)
#plot beauty
#ax.set_xlim(-.06, -.02)
ax.set_frame_on(False)
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.set_xlabel('regression slope [bouts/m]')
ax.set_ylabel('Molecular weight [g/mol]')
ax.set_xticks([-50, -40, -30, -20, -10])
ax.set_ylim(3**2,10**2)
#ax.set_yticks(np.arange(20,101, 20))
ax.text(-0.1,1.15, "L)", fontweight='bold', transform=ax.transAxes)
cc_mw, pcc_mw = pearsonr(reg_slopes, mw)
cc_sqrt, pcc_sqrt = pearsonr(reg_slopes, np.sqrt(mw))
print()
print('Molecular weight vs. bout counts, correlation between molecular weight and regression coefficient')
print("CC = {:.3f}, p = {:.3f}".format(cc_mw,pcc_mw))
print('between sqrt(molecular weight) and regression coefficient')
print("CC = {:.3f}, p = {:.3f}".format(cc_sqrt,pcc_sqrt))
# plot all bout counts against distance
# not in the figure currently, making space for the slopes vs MW
#all_bc = [np.mean(pdd['bc'], axis=1) for pdd in plot_data_dicts]
#ax = f.add_subplot(gs[i+1])
#for ibc,bc in enumerate(all_bc):
# if ibc == 4:
# xax = np.array([0.25, 0.50, 0.98, 1.18, 1.40])
# elif ibc == 6:
# xax = np.array([0.25, 0.98, 1.18, 1.40, 1.45])
# else:
# xax = np.array([0.25, 0.50, 0.98, 1.18, 1.40, 1.45])
# ax.plot(xax, bc, linestyle='-', marker='.', mfc='k', color='gray', linewidth=.3)
#ax.set_frame_on(False)
#ax.xaxis.set_ticks_position('bottom')
#ax.yaxis.set_ticks_position('left')
#ax.set_xlim(0.1,1.5)
#ax.set_xticks(xax)
#ax.set_xticklabels(["0.25", "0.50", "0.98", "", "1.40", ""])
#ax.set_title("Mean bout counts", fontsize=10)
#ax.text(-0.1,1.15, "L)", fontweight='bold', transform=ax.transAxes)
# -
f.savefig("Figures/Fig. 8 - Boutcounts for all gases.png", dpi=600)
with open("Figures/multivar_results_windspeed1.pkl", 'rb') as du:
multivar_results = pickle.load(du)
#TODO: need to save all single sensor performance values, too.
rmses_mv = [np.mean(d['mv']["RMSE"]) for d in multivar_results]
rmses_uv = np.zeros((7, len(multivar_results)))
for i,mv in enumerate(multivar_results):
rmses_uv[:,i] = mv["single"]
f = plt.figure(figsize=(6,4))
ax = f.add_subplot(111)
plt.plot(rmses_mv,'*', mec="k", ms=7, mfc="k", ls="none", label="multivar")
symbols = ["o", "v", "^", "<", ">", "s", 'p']
for i in range(rmses_uv.shape[0]):
plt.plot(np.arange(11)+0.2, rmses_uv[i], marker=symbols[i], color='k', ms=3, mfc="none", ls="none", label=sensornames[i+1])
plt.legend(frameon=False, fontsize=7, numpoints=1, loc=(0.88,0.5))
ax.set_xlim(-1,12)
ax.set_ylim(-0.02,0.501)
ax.set_ylabel("CV RMSE")
ax.set_xticks(range(11))
ax.set_xticklabels(["\n".join((ds.GasNames[i] + " ppm").split("_")) for i in range(1,12)], rotation=90, fontsize=8)
ax.set_frame_on(False)
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
pos1 = ax.get_position() # get the original position
pos2 = [pos1.x0, pos1.y0 + 0.08, pos1.width, pos1.height]
ax.set_position(pos2) # set a new position
f.savefig("Figures/Fig. 9 - Multivariate vs univariate.png", dpi=600)
| ipnotebooks/manuscript_figures/Fig. 8 and 9 - Gas invariance of distance prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EDA
#
# ## Distribution of target variable
from eda import *
local_df = load_train_data()
histogram_target_data(df=local_df)
# ## Correlation of Target with Energy Star Rate
correlation_target_energy_star(df=local_df)
# ## Correlation of variables in general
correlations = local_df.corr()
correlations
| notebooks/EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from mako.template import Template
from datetime import datetime
import os
import netCDF4 as nc
def results_dataset(period, grid, results_dir):
"""Return the results dataset for period (e.g. 1h or 1d)
and grid (e.g. grid_T, grid_U) from results_dir.
"""
filename_pattern = 'SalishSea_{period}_*_{grid}.nc'
filepaths = glob(os.path.join(results_dir, filename_pattern.format(period=period, grid=grid)))
return nc.Dataset(filepaths[0])
# +
#model_path = '/ocean/sallen/allen/research/MEOPAR/Operational/'
results_home = '/ocean/nsoontie/MEOPAR/sprint/'
grid_T_hr = results_dataset('1h', 'grid_T', results_home)
for name in names:
# Get sea surface height
lat = SITES[name]['lat']
lon = SITES[name]['lon']
j, i = tidetools.find_closest_model_point(lon, lat, X, Y, bathy, allow_land=False)
ssh_loc = grid_T.variables['sossheig'][:, j, i]
# Get tides and ssh
ttide = get_tides(name)
ssh_corr = correct_model_ssh(ssh_loc, t, ttide)
# Information
res = compute_residual(ssh_loc, t, ttide)
[max_ssh, index_ssh, tmax, max_res, max_wind, ind_w] = get_maxes(
ssh_corr, t, res, lon, lat, model_path)
max_sshs[name] = max_ssh
max_times[name] = tmax
max_winds[name] = max_wind
# -
def make_web_strings(Stations):
'''Construct strings for surge warning zones and times of day and return them as part of the input dict STATIONS
'''
# Initialize WARNINGS string
Warnings = ''
Threshold = 1
# Iterate through tide stations
for station in Stations:
# Add areas to WARNINGS string
if Stations[station]['max_sealevel'] > threshold:
Warnings = Warnings + Stations[station]['area'] + ', '
# Define times of day in readable format (e.g. 25-Aug-2015 06:00 becomes "early Tuesday morning")
# and append to STATIONS dict
Stations[station]['day'] = Stations[station]['date'].strftime('%A')
if Stations[station]['date'].hour < 12:
Stations[station]['time'] = 'morning'
if Stations[station]['date'].hour < 8:
Stations[station]['period'] = 'early'
else:
Stations[station]['period'] = 'late'
elif Stations[station]['date'].hour >= 12 and Stations[station]['date'].hour < 17:
Stations[station]['time'] = 'afternoon'
if Stations[station]['date'].hour < 15:
Stations[station]['period'] = 'early'
else:
Stations[station]['period'] = 'late'
else:
Stations[station]['time'] = 'evening'
if Stations[station]['date'].hour < 20:
Stations[station]['period'] = 'early'
else:
Stations[station]['period'] = 'late'
# Final WARNINGS syntax and append to STATIONS dict
Stations['Warnings'] = Warnings[:-2][::-1].replace(',', ';', 1)[::-1].replace(';', ' and')
return Stations
# +
# STATIONS dict template
Stations = {'PA': {'name': 'Point Atkinson', 'area': 'Vancouver', 'max_sealevel': 1.5, 'windspeed': 2.5, 'date': datetime.now()},
'VI': {'name': 'Victoria', 'area': 'Victoria', 'max_sealevel': 0.3, 'windspeed': 2.3, 'date': datetime.now()},
'CP': {'name': '<NAME>', 'area': 'Boundary Bay', 'max_sealevel': 0.2, 'windspeed': 1.5, 'date': datetime.now()},
'CR': {'name': '<NAME>', 'area': 'Campbell River', 'max_sealevel': 0.1, 'windspeed': 0.5, 'date': datetime.now()}}
# Make WARNINGS and readable time strings
Stations = make_web_strings(Stations)
# Generate RST in Mako
print(Template(filename='www/templates/surgetext.mako').render(**Stations))
# -
| notebooks/surge_warning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: torch
# language: python
# name: torch
# ---
# cd ..
import dlib
import glob
import os
from tqdm import tqdm
from utils.alignment import align_face
images_path = 'images'
SHAPE_PREDICTOR_PATH = 'pretrained_models/shape_predictor_68_face_landmarks.dat'
IMAGE_SIZE = 1024
predictor = dlib.shape_predictor(SHAPE_PREDICTOR_PATH)
os.chdir(images_path)
images_names = glob.glob(f'*')
images_names
aligned_images = []
for image_name in tqdm(images_names):
try:
aligned_image = align_face(filepath=f'{images_path}/{image_name}',
predictor=predictor, output_size=IMAGE_SIZE)
aligned_images.append(aligned_image)
except Exception as e:
print(e)
os.makedirs(f'{images_path}/aligned', exist_ok=True)
os.makedirs(f'{images_path}/aligned/0', exist_ok=True)
for image, name in zip(aligned_images,images_names):
real_name = name.split('.')[0]
try:
image.save(f'{images_path}/aligned/0/{real_name}.jpeg')
except Exception as e:
print(e)
| notebooks/align_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# name: python36964bit5ac2507db1eb4760aebe700123c964da
# ---
# # Prepare MNIST dataset and split
#
# | | name | label | normal_data | backdoor_data | description |
# | :--- | :--- | :---: | :---: | :---: | :--- |
# | $D_{train}$ | Clean training dataset | $\checkmark$ | 50,000 | 0 | Train baseline model |
# | $D^p_{train}$ | Poison training dataset | $\checkmark$ | 49,900 | 100 | Train backdoor model |
# | $D_{dist}$ | Distillation training dataset | | 10,000 | 0 | Train distilled model |
# | $D_{test}$ | Clean test dataset | $\checkmark$ | 10,000 | 0 | Validate stealthiness |
# | $D_{p}$ | Poison test dataset | $\checkmark$ | 0 | 10,000 | Validate attack feasibility |
# +
import os
import numpy as np
import torch
import torchvision
from torchvision import transforms
from backdoor_attack import create_poison_data
# -
# ## Prepare MNIST dataset
# + tags=[]
ds_root = os.path.join('.', 'results', 'datasets')
original_data_path = os.path.join(ds_root, 'original_data')
os.makedirs(ds_root, exist_ok=True)
mnist_train = torchvision.datasets.MNIST(original_data_path, train=True, download=True)
mnist_test = torchvision.datasets.MNIST(original_data_path, train=False, download=True)
# -
# ### Configure training datasets
np.random.seed(20200703)
train_x = mnist_train.data.numpy()
train_t = mnist_train.targets.numpy()
idx = np.random.permutation(np.arange(train_x.shape[0]))
# Clean training dataset
x = train_x[idx[:50000]]
t = train_t[idx[:50000]]
np.savez(os.path.join(ds_root, 'clean_training_dataset.npz'), x=x, t=t)
# +
# Poison training dataset
poisoned_target = 7 # poisoned target
num_of_poison_data = 100
x_p = train_x[idx[:50000]]
t_p = train_t[idx[:50000]]
t = train_t[idx[:50000]]
i = 0
n = 0
while n < num_of_poison_data:
if t_p[i] != poisoned_target:
x_p[i] = create_poison_data.one_dot_mnist(x[i])
t_p[i] = poisoned_target
n += 1
i += 1
shuffle_idx = np.random.permutation(np.arange(x_p.shape[0]))
np.savez(os.path.join(ds_root, 'poison_training_dataset.npz'), x=x_p[shuffle_idx], t=t_p[shuffle_idx], t_correct=t[shuffle_idx])
# + tags=[]
# Distillation training dataset
x = train_x[idx[50000:]]
t = train_t[idx[50000:]]
np.savez(os.path.join(ds_root, 'distillation_training_dataset.npz'), x=x, t=t)
# -
# ### Configure test datasets
test_x = mnist_test.data.numpy()
test_t = mnist_test.targets.numpy()
# Clean test dataset
np.savez(os.path.join(ds_root, 'clean_test_dataset.npz'), x=test_x, t=test_t)
# +
# Poison test dataset
x_p = []
t_p = []
target = []
for x, t in zip(test_x, test_t):
if t != poisoned_target:
x_p.append(create_poison_data.one_dot_mnist(x)[np.newaxis, ...])
t_p.append(poisoned_target)
target.append(t)
x_p = np.concatenate(x_p, axis=0)
t_p = np.array(t_p, dtype=np.int32)
t = np.array(t, dtype=np.int32)
np.savez(os.path.join(ds_root, 'poison_test_dataset.npz'), x=x_p, t=t_p, t_correct=target)
# -
| experiments/MNIST/1_Prepare_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!-- dom:TITLE: PHY321: Motion examples, Forces, Newton's Laws and Motion Example -->
# # PHY321: Motion examples, Forces, Newton's Laws and Motion Example
# <!-- dom:AUTHOR: [<NAME>](http://mhjgit.github.io/info/doc/web/) at Department of Physics and Astronomy and Facility for Rare Ion Beams (FRIB), Michigan State University, USA & Department of Physics, University of Oslo, Norway -->
# <!-- Author: -->
# **[<NAME>](http://mhjgit.github.io/info/doc/web/)**, Department of Physics and Astronomy and Facility for Rare Ion Beams (FRIB), Michigan State University, USA and Department of Physics, University of Oslo, Norway
#
# Date: **Jan 25, 2021**
#
# Copyright 1999-2021, [<NAME>](http://mhjgit.github.io/info/doc/web/). Released under CC Attribution-NonCommercial 4.0 license
#
#
#
#
# ## Aims and Overarching Motivation
#
# ### Monday
#
# We try to finalize the discussion we started last Friday on falling objects and numerical aspects thereof.
# If we get time, we start with a discussion of forces as well.
#
# Recommended reading: Taylor 1.3
#
# ### Wednesday
#
# We revisit Newton's laws and discuss how to analyze a problem.
#
# Recommended reading: Taylor 1.4 and 1.5
#
# ### Friday
#
# We discuss several examples and try to wrap up the discussions on Newton's laws.
#
# Recommended reading: Taylor 1.4-1.6 and 2.1-2.2 as examples of motion problems.
#
#
#
# ## Basic Steps of Scientific Investigations
#
# Last week we discussed several basi elements of the scientific method. We repeat them here.
#
# An overarching aim in this course is to give you a deeper
# understanding of the scientific method. The problems we study will all
# involve cases where we can apply classical mechanics. In our previous
# material we already assumed that we had a model for the motion of an
# object. Alternatively we could have data from experiment (like Usain
# Bolt's 100m world record run in 2008). Or we could have performed
# ourselves an experiment and we want to understand which forces are at
# play and whether these forces can be understood in terms of
# fundamental forces.
#
# Our first step consists in identifying the problem. What we sketch
# here may include a mix of experiment and theoretical simulations, or
# just experiment or only theory.
#
# ## Identifying our System
#
# Here we can ask questions like
# 1. What kind of object is moving
#
# 2. What kind of data do we have
#
# 3. How do we measure position, velocity, acceleration etc
#
# 4. Which initial conditions influence our system
#
# 5. Other aspects which allow us to identify the system
#
# ## Defining a Model
#
# With our eventual data and observations we would now like to develop a
# model for the system. In the end we want obviously to be able to
# understand which forces are at play and how they influence our
# specific system. That is, can we extract some deeper insights about a
# system?
#
# We need then to
# 1. Find the forces that act on our system
#
# 2. Introduce models for the forces
#
# 3. Identify the equations which can govern the system (Newton's second law for example)
#
# 4. More elements we deem important for defining our model
#
# ## Solving the Equations
#
# With the model at hand, we can then solve the equations. In classical mechanics we normally end up with solving sets of coupled ordinary differential equations or partial differential equations.
# 1. Using Newton's second law we have equations of the type $\boldsymbol{F}=m\boldsymbol{a}=md\boldsymbol{v}/dt$
#
# 2. We need to define the initial conditions (typically the initial velocity and position as functions of time) and/or initial conditions and boundary conditions
#
# 3. The solution of the equations give us then the position, the velocity and other time-dependent quantities which may specify the motion of a given object.
#
# We are not yet done. With our lovely solvers, we need to start thinking.
#
# ## Analyze
#
# Now it is time to ask the big questions. What do our results mean? Can we give a simple interpretation in terms of fundamental laws? What do our results mean? Are they correct?
# Thus, typical questions we may ask are
# 1. Are our results for say $\boldsymbol{r}(t)$ valid? Do we trust what we did? Can you validate and verify the correctness of your results?
#
# 2. Evaluate the answers and their implications
#
# 3. Compare with experimental data if possible. Does our model make sense?
#
# 4. and obviously many other questions.
#
# The analysis stage feeds back to the first stage. It may happen that
# the data we had were not good enough, there could be large statistical
# uncertainties. We may need to collect more data or perhaps we did a
# sloppy job in identifying the degrees of freedom.
#
# All these steps are essential elements in a scientific
# enquiry. Hopefully, through a mix of numerical simulations, analytical
# calculations and experiments we may gain a deeper insight about the
# physics of a specific system.
#
#
# ## Falling baseball in one dimension
#
# We anticipate the mathematical model to come and assume that we have a
# model for the motion of a falling baseball without air resistance.
# Our system (the baseball) is at an initial height $y_0$ (which we will
# specify in the program below) at the initial time $t_0=0$. In our program example here we will plot the position in steps of $\Delta t$ up to a final time $t_f$.
# The mathematical formula for the position $y(t)$ as function of time $t$ is
# $$
# y(t) = y_0-\frac{1}{2}gt^2,
# $$
# where $g=9.80665=0.980655\times 10^1$m/s${}^2$ is a constant representing the standard acceleration due to gravity.
# We have here adopted the conventional standard value. This does not take into account other effects, such as buoyancy or drag.
# Furthermore, we stop when the ball hits the ground, which takes place at
# $$
# y(t) = 0= y_0-\frac{1}{2}gt^2,
# $$
# which gives us a final time $t_f=\sqrt{2y_0/g}$.
#
# As of now we simply assume that we know the formula for the falling object. Afterwards, we will derive it.
#
# ## Our Python Encounter
#
# We start with preparing folders for storing our calculations, figures and if needed, specific data files we use as input or output files.
# +
# %matplotlib inline
# Common imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
# Where to save the figures and data files
PROJECT_ROOT_DIR = "Results"
FIGURE_ID = "Results/FigureFiles"
DATA_ID = "DataFiles/"
if not os.path.exists(PROJECT_ROOT_DIR):
os.mkdir(PROJECT_ROOT_DIR)
if not os.path.exists(FIGURE_ID):
os.makedirs(FIGURE_ID)
if not os.path.exists(DATA_ID):
os.makedirs(DATA_ID)
def image_path(fig_id):
return os.path.join(FIGURE_ID, fig_id)
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
def save_fig(fig_id):
plt.savefig(image_path(fig_id) + ".png", format='png')
#in case we have an input file we wish to read in
#infile = open(data_path("MassEval2016.dat"),'r')
# -
# You could also define a function for making our plots. You
# can obviously avoid this and simply set up various **matplotlib**
# commands every time you need them. You may however find it convenient
# to collect all such commands in one function and simply call this
# function.
# +
from pylab import plt, mpl
plt.style.use('seaborn')
mpl.rcParams['font.family'] = 'serif'
def MakePlot(x,y, styles, labels, axlabels):
plt.figure(figsize=(10,6))
for i in range(len(x)):
plt.plot(x[i], y[i], styles[i], label = labels[i])
plt.xlabel(axlabels[0])
plt.ylabel(axlabels[1])
plt.legend(loc=0)
# -
# Thereafter we start setting up the code for the falling object.
# +
# %matplotlib inline
import matplotlib.patches as mpatches
g = 9.80655 #m/s^2
y_0 = 10.0 # initial position in meters
DeltaT = 0.1 # time step
# final time when y = 0, t = sqrt(2*10/g)
tfinal = np.sqrt(2.0*y_0/g)
#set up arrays
t = np.arange(0,tfinal,DeltaT)
y =y_0 -g*.5*t**2
# Then make a nice printout in table form using Pandas
import pandas as pd
from IPython.display import display
data = {'t[s]': t,
'y[m]': y
}
RawData = pd.DataFrame(data)
display(RawData)
plt.style.use('ggplot')
plt.figure(figsize=(8,8))
plt.scatter(t, y, color = 'b')
blue_patch = mpatches.Patch(color = 'b', label = 'Height y as function of time t')
plt.legend(handles=[blue_patch])
plt.xlabel("t[s]")
plt.ylabel("y[m]")
save_fig("FallingBaseball")
plt.show()
# -
# Here we used **pandas** (see below) to systemize the output of the position as function of time.
#
#
# ## Average quantities
# We define now the average velocity as
# $$
# \overline{v}(t) = \frac{y(t+\Delta t)-y(t)}{\Delta t}.
# $$
# In the code we have set the time step $\Delta t$ to a given value. We could define it in terms of the number of points $n$ as
# $$
# \Delta t = \frac{t_{\mathrm{final}-}t_{\mathrm{initial}}}{n+1}.
# $$
# Since we have discretized the variables, we introduce the counter $i$ and let $y(t)\rightarrow y(t_i)=y_i$ and $t\rightarrow t_i$
# with $i=0,1,\dots, n$. This gives us the following shorthand notations that we will use for the rest of this course. We define
# $$
# y_i = y(t_i),\hspace{0.2cm} i=0,1,2,\dots,n.
# $$
# This applies to other variables which depend on say time. Examples are the velocities, accelerations, momenta etc.
# Furthermore we use the shorthand
# $$
# y_{i\pm 1} = y(t_i\pm \Delta t),\hspace{0.12cm} i=0,1,2,\dots,n.
# $$
# ## Compact equations
# We can then rewrite in a more compact form the average velocity as
# $$
# \overline{v}_i = \frac{y_{i+1}-y_{i}}{\Delta t}.
# $$
# The velocity is defined as the change in position per unit time.
# In the limit $\Delta t \rightarrow 0$ this defines the instantaneous velocity, which is nothing but the slope of the position at a time $t$.
# We have thus
# $$
# v(t) = \frac{dy}{dt}=\lim_{\Delta t \rightarrow 0}\frac{y(t+\Delta t)-y(t)}{\Delta t}.
# $$
# Similarly, we can define the average acceleration as the change in velocity per unit time as
# $$
# \overline{a}_i = \frac{v_{i+1}-v_{i}}{\Delta t},
# $$
# resulting in the instantaneous acceleration
# $$
# a(t) = \frac{dv}{dt}=\lim_{\Delta t\rightarrow 0}\frac{v(t+\Delta t)-v(t)}{\Delta t}.
# $$
# **A note on notations**: When writing for example the velocity as $v(t)$ we are then referring to the continuous and instantaneous value. A subscript like
# $v_i$ refers always to the discretized values.
#
# ## A differential equation
# We can rewrite the instantaneous acceleration as
# $$
# a(t) = \frac{dv}{dt}=\frac{d}{dt}\frac{dy}{dt}=\frac{d^2y}{dt^2}.
# $$
# This forms the starting point for our definition of forces later. It is a famous second-order differential equation. If the acceleration is constant we can now recover the formula for the falling ball we started with.
# The acceleration can depend on the position and the velocity. To be more formal we should then write the above differential equation as
# $$
# \frac{d^2y}{dt^2}=a(t,y(t),\frac{dy}{dt}).
# $$
# With given initial conditions for $y(t_0)$ and $v(t_0)$ we can then
# integrate the above equation and find the velocities and positions at
# a given time $t$.
#
# If we multiply with mass, we have one of the famous expressions for Newton's second law,
# $$
# F(y,v,t)=m\frac{d^2y}{dt^2}=ma(t,y(t),\frac{dy}{dt}),
# $$
# where $F$ is the force acting on an object with mass $m$. We see that it also has the right dimension, mass times length divided by time squared.
# We will come back to this soon.
#
# ## Integrating our equations
#
# Formally we can then, starting with the acceleration (suppose we have measured it, how could we do that?)
# compute say the height of a building. To see this we perform the following integrations from an initial time $t_0$ to a given time $t$
# $$
# \int_{t_0}^t dt a(t) = \int_{t_0}^t dt \frac{dv}{dt} = v(t)-v(t_0),
# $$
# or as
# $$
# v(t)=v(t_0)+\int_{t_0}^t dt a(t).
# $$
# When we know the velocity as function of time, we can find the position as function of time starting from the defintion of velocity as the derivative with respect to time, that is we have
# $$
# \int_{t_0}^t dt v(t) = \int_{t_0}^t dt \frac{dy}{dt} = y(t)-y(t_0),
# $$
# or as
# $$
# y(t)=y(t_0)+\int_{t_0}^t dt v(t).
# $$
# These equations define what is called the integration method for
# finding the position and the velocity as functions of time. There is
# no loss of generality if we extend these equations to more than one
# spatial dimension.
#
# ## Constant acceleration case, the velocity
# Let us compute the velocity using the constant value for the acceleration given by $-g$. We have
# $$
# v(t)=v(t_0)+\int_{t_0}^t dt a(t)=v(t_0)+\int_{t_0}^t dt (-g).
# $$
# Using our initial time as $t_0=0$s and setting the initial velocity $v(t_0)=v_0=0$m/s we get when integrating
# $$
# v(t)=-gt.
# $$
# The more general case is
# $$
# v(t)=v_0-g(t-t_0).
# $$
# We can then integrate the velocity and obtain the final formula for the position as function of time through
# $$
# y(t)=y(t_0)+\int_{t_0}^t dt v(t)=y_0+\int_{t_0}^t dt v(t)=y_0+\int_{t_0}^t dt (-gt),
# $$
# With $y_0=10$m and $t_0=0$s, we obtain the equation we started with
# $$
# y(t)=10-\frac{1}{2}gt^2.
# $$
# ## Computing the averages
# After this mathematical background we are now ready to compute the mean velocity using our data.
# Now we can compute the mean velocity using our data
# We define first an array Vaverage
n = np.size(t)
Vaverage = np.zeros(n)
for i in range(1,n-1):
Vaverage[i] = (y[i+1]-y[i])/DeltaT
# Now we can compute the mean accelearatio using our data
# We define first an array Aaverage
n = np.size(t)
Aaverage = np.zeros(n)
Aaverage[0] = -g
for i in range(1,n-1):
Aaverage[i] = (Vaverage[i+1]-Vaverage[i])/DeltaT
data = {'t[s]': t,
'y[m]': y,
'v[m/s]': Vaverage,
'a[m/s^2]': Aaverage
}
NewData = pd.DataFrame(data)
display(NewData[0:n-2])
# Note that we don't print the last values!
#
#
#
# ## Including Air Resistance in our model
#
# In our discussions till now of the falling baseball, we have ignored
# air resistance and simply assumed that our system is only influenced
# by the gravitational force. We will postpone the derivation of air
# resistance till later, after our discussion of Newton's laws and
# forces.
#
# For our discussions here it suffices to state that the accelerations is now modified to
# $$
# \boldsymbol{a}(t) = -g +D\boldsymbol{v}(t)\vert v(t)\vert,
# $$
# where $\vert v(t)\vert$ is the absolute value of the velocity and $D$ is a constant which pertains to the specific object we are studying.
# Since we are dealing with motion in one dimension, we can simplify the above to
# $$
# a(t) = -g +Dv^2(t).
# $$
# We can rewrite this as a differential equation
# $$
# a(t) = \frac{dv}{dt}=\frac{d^2y}{dt^2}= -g +Dv^2(t).
# $$
# Using the integral equations discussed above we can integrate twice
# and obtain first the velocity as function of time and thereafter the
# position as function of time.
#
# For this particular case, we can actually obtain an analytical
# solution for the velocity and for the position. Here we will first
# compute the solutions analytically, thereafter we will derive Euler's
# method for solving these differential equations numerically.
#
# ## Analytical solutions
#
# For simplicity let us just write $v(t)$ as $v$. We have
# $$
# \frac{dv}{dt}= -g +Dv^2(t).
# $$
# We can solve this using the technique of separation of variables. We
# isolate on the left all terms that involve $v$ and on the right all
# terms that involve time. We get then
# $$
# \frac{dv}{g -Dv^2(t) }= -dt,
# $$
# We scale now the equation to the left by introducing a constant
# $v_T=\sqrt{g/D}$. This constant has dimension length/time. Can you
# show this?
#
# Next we integrate the left-hand side (lhs) from $v_0=0$ m/s to $v$ and
# the right-hand side (rhs) from $t_0=0$ to $t$ and obtain
# $$
# \int_{0}^v\frac{dv}{g -Dv^2(t) }= \frac{v_T}{g}\mathrm{arctanh}(\frac{v}{v_T}) =-\int_0^tdt = -t.
# $$
# We can reorganize these equations as
# $$
# v_T\mathrm{arctanh}(\frac{v}{v_T}) =-gt,
# $$
# which gives us $v$ as function of time
# $$
# v(t)=v_T\tanh{-(\frac{gt}{v_T})}.
# $$
# ## Finding the final height
# With the velocity we can then find the height $y(t)$ by integrating yet another time, that is
# $$
# y(t)=y(t_0)+\int_{t_0}^t dt v(t)=\int_{0}^t dt[v_T\tanh{-(\frac{gt}{v_T})}].
# $$
# This integral is a little bit trickier but we can look it up in a table over
# known integrals and we get
# $$
# y(t)=y(t_0)-\frac{v_T^2}{g}\log{[\cosh{(\frac{gt}{v_T})}]}.
# $$
# Alternatively we could have used the symbolic Python package **Sympy**.
#
# In most cases however, we need to revert to numerical solutions.
#
#
# ## Our first attempt at solving differential equations
#
# Here we will try the simplest possible approach to solving the second-order differential
# equation
# $$
# a(t) =\frac{d^2y}{dt^2}= -g +Dv^2(t).
# $$
# We rewrite it as two coupled first-order equations (this is a standard approach)
# $$
# \frac{dy}{dt} = v(t),
# $$
# with initial condition $y(t_0)=y_0$ and
# $$
# a(t) =\frac{dv}{dt}= -g +Dv^2(t),
# $$
# with initial condition $v(t_0)=v_0$.
#
# Many of the algorithms for solving differential equations start with simple Taylor equations.
# If we now Taylor expand $y$ and $v$ around a value $t+\Delta t$ we have
# $$
# y(t+\Delta t) = y(t)+\Delta t \frac{dy}{dt}+\frac{\Delta t^2}{2!} \frac{d^2y}{dt^2}+O(\Delta t^3),
# $$
# and
# $$
# v(t+\Delta t) = v(t)+\Delta t \frac{dv}{dt}+\frac{\Delta t^2}{2!} \frac{d^2v}{dt^2}+O(\Delta t^3).
# $$
# Using the fact that $dy/dt = v$ and $dv/dt=a$ and keeping only terms up to $\Delta t$ we have
# $$
# y(t+\Delta t) = y(t)+\Delta t v(t)+O(\Delta t^2),
# $$
# and
# $$
# v(t+\Delta t) = v(t)+\Delta t a(t)+O(\Delta t^2).
# $$
# ## Discretizing our equations
#
# Using our discretized versions of the equations with for example
# $y_{i}=y(t_i)$ and $y_{i\pm 1}=y(t_i+\Delta t)$, we can rewrite the
# above equations as (and truncating at $\Delta t$)
# $$
# y_{i+1} = y_i+\Delta t v_i,
# $$
# and
# $$
# v_{i+1} = v_i+\Delta t a_i.
# $$
# These are the famous Euler equations (forward Euler).
#
# To solve these equations numerically we start at a time $t_0$ and simply integrate up these equations to a final time $t_f$,
# The step size $\Delta t$ is an input parameter in our code.
# You can define it directly in the code below as
DeltaT = 0.1
# With a given final time **tfinal** we can then find the number of integration points via the **ceil** function included in the **math** package of Python
# as
#define final time, assuming that initial time is zero
from math import ceil
tfinal = 0.5
n = ceil(tfinal/DeltaT)
print(n)
# The **ceil** function returns the smallest integer not less than the input in say
x = 21.15
print(ceil(x))
# which in the case here is 22.
x = 21.75
print(ceil(x))
# which also yields 22. The **floor** function in the **math** package
# is used to return the closest integer value which is less than or equal to the specified expression or value.
# Compare the previous result to the usage of **floor**
from math import floor
x = 21.75
print(floor(x))
# Alternatively, we can define ourselves the number of integration(mesh) points. In this case we could have
n = 10
tinitial = 0.0
tfinal = 0.5
DeltaT = (tfinal-tinitial)/(n)
print(DeltaT)
# Since we will set up one-dimensional arrays that contain the values of
# various variables like time, position, velocity, acceleration etc, we
# need to know the value of $n$, the number of data points (or
# integration or mesh points). With $n$ we can initialize a given array
# by setting all elelements to zero, as done here
# define array a
a = np.zeros(n)
print(a)
# ## Code for implementing Euler's method
# In the code here we implement this simple Eurler scheme choosing a value for $D=0.0245$ m/s.
# +
# Common imports
import numpy as np
import pandas as pd
from math import *
import matplotlib.pyplot as plt
import os
# Where to save the figures and data files
PROJECT_ROOT_DIR = "Results"
FIGURE_ID = "Results/FigureFiles"
DATA_ID = "DataFiles/"
if not os.path.exists(PROJECT_ROOT_DIR):
os.mkdir(PROJECT_ROOT_DIR)
if not os.path.exists(FIGURE_ID):
os.makedirs(FIGURE_ID)
if not os.path.exists(DATA_ID):
os.makedirs(DATA_ID)
def image_path(fig_id):
return os.path.join(FIGURE_ID, fig_id)
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
def save_fig(fig_id):
plt.savefig(image_path(fig_id) + ".png", format='png')
g = 9.80655 #m/s^2
D = 0.00245 #m/s
DeltaT = 0.1
#set up arrays
tfinal = 0.5
n = ceil(tfinal/DeltaT)
# define scaling constant vT
vT = sqrt(g/D)
# set up arrays for t, a, v, and y and we can compare our results with analytical ones
t = np.zeros(n)
a = np.zeros(n)
v = np.zeros(n)
y = np.zeros(n)
yanalytic = np.zeros(n)
# Initial conditions
v[0] = 0.0 #m/s
y[0] = 10.0 #m
yanalytic[0] = y[0]
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i]
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT))
if ( y[i+1] < 0.0):
break
a[n-1] = -g + D*v[n-1]*v[n-1]
data = {'t[s]': t,
'y[m]': y-yanalytic,
'v[m/s]': v,
'a[m/s^2]': a
}
NewData = pd.DataFrame(data)
display(NewData)
#finally we plot the data
fig, axs = plt.subplots(3, 1)
axs[0].plot(t, y, t, yanalytic)
axs[0].set_xlim(0, tfinal)
axs[0].set_ylabel('y and exact')
axs[1].plot(t, v)
axs[1].set_ylabel('v[m/s]')
axs[2].plot(t, a)
axs[2].set_xlabel('time[s]')
axs[2].set_ylabel('a[m/s^2]')
fig.tight_layout()
save_fig("EulerIntegration")
plt.show()
# -
# Try different values for $\Delta t$ and study the difference between the exact solution and the numerical solution.
#
# ## Simple extension, the Euler-Cromer method
#
# The Euler-Cromer method is a simple variant of the standard Euler
# method. We use the newly updated velocity $v_{i+1}$ as an input to the
# new position, that is, instead of
# $$
# y_{i+1} = y_i+\Delta t v_i,
# $$
# and
# $$
# v_{i+1} = v_i+\Delta t a_i,
# $$
# we use now the newly calculate for $v_{i+1}$ as input to $y_{i+1}$, that is
# we compute first
# $$
# v_{i+1} = v_i+\Delta t a_i,
# $$
# and then
# $$
# y_{i+1} = y_i+\Delta t v_{i+1},
# $$
# Implementing the Euler-Cromer method yields a simple change to the previous code. We only need to change the following line in the loop over time
# steps
for i in range(n-1):
# more codes in between here
v[i+1] = v[i] + DeltaT*a[i]
y[i+1] = y[i] + DeltaT*v[i+1]
# more code
# ## Newton's Laws
#
# Let us now remind ourselves of Newton's laws, since these are the laws of motion we will study in this course.
#
#
# When analyzing a physical system we normally start with distinguishing between the object we are studying (we will label this in more general terms as our **system**) and how this system interacts with the environment (which often means everything else!)
#
# In our investigations we will thus analyze a specific physics problem in terms of the system and the environment.
# In doing so we need to identify the forces that act on the system and assume that the
# forces acting on the system must have a source, an identifiable cause in
# the environment.
#
# A force acting on for example a falling object must be related to an interaction with something in the environment.
# This also means that we do not consider internal forces. The latter are forces between
# one part of the object and another part. In this course we will mainly focus on external forces.
#
# Forces are either contact forces or long-range forces.
#
# Contact forces, as evident from the name, are forces that occur at the contact between
# the system and the environment. Well-known long-range forces are the gravitional force and the electromagnetic force.
#
#
# ## Setting up a model for forces acting on an object
#
# In order to set up the forces which act on an object, the following steps may be useful
# 1. Divide the problem into system and environment.
#
# 2. Draw a figure of the object and everything in contact with the object.
#
# 3. Draw a closed curve around the system.
#
# 4. Find contact points—these are the points where contact forces may act.
#
# 5. Give names and symbols to all the contact forces.
#
# 6. Identify the long-range forces.
#
# 7. Make a drawing of the object. Draw the forces as arrows, vectors, starting from where the force is acting. The direction of the vector(s) indicates the (positive) direction of the force. Try to make the length of the arrow indicate the relative magnitude of the forces.
#
# 8. Draw in the axes of the coordinate system. It is often convenient to make one axis parallel to the direction of motion. When you choose the direction of the axis you also choose the positive direction for the axis.
#
# ## Newton's Laws, the Second one first
#
#
# Newton’s second law of motion: The force $\boldsymbol{F}$ on an object of inertial mass $m$
# is related to the acceleration a of the object through
# $$
# \boldsymbol{F} = m\boldsymbol{a},
# $$
# where $\boldsymbol{a}$ is the acceleration.
#
# Newton’s laws of motion are laws of nature that have been found by experimental
# investigations and have been shown to hold up to continued experimental investigations.
# Newton’s laws are valid over a wide range of length- and time-scales. We
# use Newton’s laws of motion to describe everything from the motion of atoms to the
# motion of galaxies.
#
# The second law is a vector equation with the acceleration having the same
# direction as the force. The acceleration is proportional to the force via the mass $m$ of the system under study.
#
#
# Newton’s second law introduces a new property of an object, the so-called
# inertial mass $m$. We determine the inertial mass of an object by measuring the
# acceleration for a given applied force.
#
#
# ## Then the First Law
#
#
# What happens if the net external force on a body is zero? Applying Newton’s second
# law, we find:
# $$
# \boldsymbol{F} = 0 = m\boldsymbol{a},
# $$
# which gives using the definition of the acceleration
# $$
# \boldsymbol{a} = \frac{d\boldsymbol{v}}{dt}=0.
# $$
# The acceleration is zero, which means that the velocity of the object is constant. This
# is often referred to as Newton’s first law. An object in a state of uniform motion tends to remain in
# that state unless an external force changes its state of motion.
# Why do we need a separate law for this? Is it not simply a special case of Newton’s
# second law? Yes, Newton’s first law can be deduced from the second law as we have
# illustrated. However, the first law is often used for a different purpose: Newton’s
# First Law tells us about the limit of applicability of Newton’s Second law. Newton’s
# Second law can only be used in reference systems where the First law is obeyed. But
# is not the First law always valid? No! The First law is only valid in reference systems
# that are not accelerated. If you observe the motion of a ball from an accelerating
# car, the ball will appear to accelerate even if there are no forces acting on it. We call
# systems that are not accelerating inertial systems, and Newton’s first law is often
# called the law of inertia. Newton’s first and second laws of motion are only valid in
# inertial systems.
#
# A system is an inertial system if it is not accelerated. It means that the reference system
# must not be accelerating linearly or rotating. Unfortunately, this means that most
# systems we know are not really inertial systems. For example, the surface of the
# Earth is clearly not an inertial system, because the Earth is rotating. The Earth is also
# not an inertial system, because it ismoving in a curved path around the Sun. However,
# even if the surface of the Earth is not strictly an inertial system, it may be considered
# to be approximately an inertial system for many laboratory-size experiments.
#
# ## And finally the Third Law
#
#
# If there is a force from object A on object B, there is also a force from object B on object A.
# This fundamental principle of interactions is called Newton’s third law. We do not
# know of any force that do not obey this law: All forces appear in pairs. Newton’s
# third law is usually formulated as: For every action there is an equal and opposite
# reaction.
#
#
# ## Motion of a Single Object
#
# Here we consider the motion of a single particle moving under
# the influence of some set of forces. We will consider some problems where
# the force does not depend on the position. In that case Newton's law
# $m\dot{\boldsymbol{v}}=\boldsymbol{F}(\boldsymbol{v})$ is a first-order differential
# equation and one solves for $\boldsymbol{v}(t)$, then moves on to integrate
# $\boldsymbol{v}$ to get the position. In essentially all of these cases we cna find an analytical solution.
#
#
# ## Air Resistance in One Dimension
#
# Air resistance tends to scale as the square of the velocity. This is
# in contrast to many problems chosen for textbooks, where it is linear
# in the velocity. The choice of a linear dependence is motivated by
# mathematical simplicity (it keeps the differential equation linear)
# rather than by physics. One can see that the force should be quadratic
# in velocity by considering the momentum imparted on the air
# molecules. If an object sweeps through a volume $dV$ of air in time
# $dt$, the momentum imparted on the air is
# <!-- Equation labels as ordinary links -->
# <div id="_auto1"></div>
#
# $$
# \begin{equation}
# dP=\rho_m dV v,
# \label{_auto1} \tag{1}
# \end{equation}
# $$
# where $v$ is the velocity of the object and $\rho_m$ is the mass
# density of the air. If the molecules bounce back as opposed to stop
# you would double the size of the term. The opposite value of the
# momentum is imparted onto the object itself. Geometrically, the
# differential volume is
# <!-- Equation labels as ordinary links -->
# <div id="_auto2"></div>
#
# $$
# \begin{equation}
# dV=Avdt,
# \label{_auto2} \tag{2}
# \end{equation}
# $$
# where $A$ is the cross-sectional area and $vdt$ is the distance the
# object moved in time $dt$.
#
# ## Resulting Acceleration
# Plugging this into the expression above,
# <!-- Equation labels as ordinary links -->
# <div id="_auto3"></div>
#
# $$
# \begin{equation}
# \frac{dP}{dt}=-\rho_m A v^2.
# \label{_auto3} \tag{3}
# \end{equation}
# $$
# This is the force felt by the particle, and is opposite to its
# direction of motion. Now, because air doesn't stop when it hits an
# object, but flows around the best it can, the actual force is reduced
# by a dimensionless factor $c_W$, called the drag coefficient.
# <!-- Equation labels as ordinary links -->
# <div id="_auto4"></div>
#
# $$
# \begin{equation}
# F_{\rm drag}=-c_W\rho_m Av^2,
# \label{_auto4} \tag{4}
# \end{equation}
# $$
# and the acceleration is
# $$
# \begin{eqnarray}
# \frac{dv}{dt}=-\frac{c_W\rho_mA}{m}v^2.
# \end{eqnarray}
# $$
# For a particle with initial velocity $v_0$, one can separate the $dt$
# to one side of the equation, and move everything with $v$s to the
# other side. We did this in our discussion of simple motion and will not repeat it here.
#
# On more general terms,
# for many systems, e.g. an automobile, there are multiple sources of
# resistance. In addition to wind resistance, where the force is
# proportional to $v^2$, there are dissipative effects of the tires on
# the pavement, and in the axel and drive train. These other forces can
# have components that scale proportional to $v$, and components that
# are independent of $v$. Those independent of $v$, e.g. the usual
# $f=\mu_K N$ frictional force you consider in your first Physics courses, only set in
# once the object is actually moving. As speeds become higher, the $v^2$
# components begin to dominate relative to the others. For automobiles
# at freeway speeds, the $v^2$ terms are largely responsible for the
# loss of efficiency. To travel a distance $L$ at fixed speed $v$, the
# energy/work required to overcome the dissipative forces are $fL$,
# which for a force of the form $f=\alpha v^n$ becomes
# $$
# \begin{eqnarray}
# W=\int dx~f=\alpha v^n L.
# \end{eqnarray}
# $$
# For $n=0$ the work is
# independent of speed, but for the wind resistance, where $n=2$,
# slowing down is essential if one wishes to reduce fuel consumption. It
# is also important to consider that engines are designed to be most
# efficient at a chosen range of power output. Thus, some cars will get
# better mileage at higher speeds (They perform better at 50 mph than at
# 5 mph) despite the considerations mentioned above.
#
# ## Going Ballistic, Projectile Motion or a Softer Approach, Falling Raindrops
#
#
# As an example of Newton's Laws we consider projectile motion (or a
# falling raindrop or a ball we throw up in the air) with a drag force. Even though air resistance is
# largely proportional to the square of the velocity, we will consider
# the drag force to be linear to the velocity, $\boldsymbol{F}=-m\gamma\boldsymbol{v}$,
# for the purposes of this exercise. The acceleration for a projectile moving upwards,
# $\boldsymbol{a}=\boldsymbol{F}/m$, becomes
# $$
# \begin{eqnarray}
# \frac{dv_x}{dt}=-\gamma v_x,\\
# \nonumber
# \frac{dv_y}{dt}=-\gamma v_y-g,
# \end{eqnarray}
# $$
# and $\gamma$ has dimensions of inverse time.
#
# If you on the other hand have a falling raindrop, how do these equations change? See for example Figure 2.1 in Taylor.
# Let us stay with a ball which is thrown up in the air at $t=0$.
#
# ## Ways of solving these equations
#
# We will go over two different ways to solve this equation. The first
# by direct integration, and the second as a differential equation. To
# do this by direct integration, one simply multiplies both sides of the
# equations above by $dt$, then divide by the appropriate factors so
# that the $v$s are all on one side of the equation and the $dt$ is on
# the other. For the $x$ motion one finds an easily integrable equation,
# $$
# \begin{eqnarray}
# \frac{dv_x}{v_x}&=&-\gamma dt,\\
# \nonumber
# \int_{v_{0x}}^{v_{x}}\frac{dv_x}{v_x}&=&-\gamma\int_0^{t}dt,\\
# \nonumber
# \ln\left(\frac{v_{x}}{v_{0x}}\right)&=&-\gamma t,\\
# \nonumber
# v_{x}(t)&=&v_{0x}e^{-\gamma t}.
# \end{eqnarray}
# $$
# This is very much the result you would have written down
# by inspection. For the $y$-component of the velocity,
# $$
# \begin{eqnarray}
# \frac{dv_y}{v_y+g/\gamma}&=&-\gamma dt\\
# \nonumber
# \ln\left(\frac{v_{y}+g/\gamma}{v_{0y}-g/\gamma}\right)&=&-\gamma t_f,\\
# \nonumber
# v_{fy}&=&-\frac{g}{\gamma}+\left(v_{0y}+\frac{g}{\gamma}\right)e^{-\gamma t}.
# \end{eqnarray}
# $$
# Whereas $v_x$ starts at some value and decays
# exponentially to zero, $v_y$ decays exponentially to the terminal
# velocity, $v_t=-g/\gamma$.
#
# ## Solving as differential equations
#
# Although this direct integration is simpler than the method we invoke
# below, the method below will come in useful for some slightly more
# difficult differential equations in the future. The differential
# equation for $v_x$ is straight-forward to solve. Because it is first
# order there is one arbitrary constant, $A$, and by inspection the
# solution is
# <!-- Equation labels as ordinary links -->
# <div id="_auto5"></div>
#
# $$
# \begin{equation}
# v_x=Ae^{-\gamma t}.
# \label{_auto5} \tag{5}
# \end{equation}
# $$
# The arbitrary constants for equations of motion are usually determined
# by the initial conditions, or more generally boundary conditions. By
# inspection $A=v_{0x}$, the initial $x$ component of the velocity.
#
#
# ## Differential Equations, contn
# The differential equation for $v_y$ is a bit more complicated due to
# the presence of $g$. Differential equations where all the terms are
# linearly proportional to a function, in this case $v_y$, or to
# derivatives of the function, e.g., $v_y$, $dv_y/dt$,
# $d^2v_y/dt^2\cdots$, are called linear differential equations. If
# there are terms proportional to $v^2$, as would happen if the drag
# force were proportional to the square of the velocity, the
# differential equation is not longer linear. Because this expression
# has only one derivative in $v$ it is a first-order linear differential
# equation. If a term were added proportional to $d^2v/dt^2$ it would be
# a second-order differential equation. In this case we have a term
# completely independent of $v$, the gravitational acceleration $g$, and
# the usual strategy is to first rewrite the equation with all the
# linear terms on one side of the equal sign,
# <!-- Equation labels as ordinary links -->
# <div id="_auto6"></div>
#
# $$
# \begin{equation}
# \frac{dv_y}{dt}+\gamma v_y=-g.
# \label{_auto6} \tag{6}
# \end{equation}
# $$
# ## Splitting into two parts
#
# Now, the solution to the equation can be broken into two
# parts. Because this is a first-order differential equation we know
# that there will be one arbitrary constant. Physically, the arbitrary
# constant will be determined by setting the initial velocity, though it
# could be determined by setting the velocity at any given time. Like
# most differential equations, solutions are not "solved". Instead,
# one guesses at a form, then shows the guess is correct. For these
# types of equations, one first tries to find a single solution,
# i.e. one with no arbitrary constants. This is called the {\it
# particular} solution, $y_p(t)$, though it should really be called
# "a" particular solution because there are an infinite number of such
# solutions. One then finds a solution to the {\it homogenous} equation,
# which is the equation with zero on the right-hand side,
# <!-- Equation labels as ordinary links -->
# <div id="_auto7"></div>
#
# $$
# \begin{equation}
# \frac{dv_{y,h}}{dt}+\gamma v_{y,h}=0.
# \label{_auto7} \tag{7}
# \end{equation}
# $$
# Homogenous solutions will have arbitrary constants.
#
# The particular solution will solve the same equation as the original
# general equation
# <!-- Equation labels as ordinary links -->
# <div id="_auto8"></div>
#
# $$
# \begin{equation}
# \frac{dv_{y,p}}{dt}+\gamma v_{y,p}=-g.
# \label{_auto8} \tag{8}
# \end{equation}
# $$
# However, we don't need find one with arbitrary constants. Hence, it is
# called a **particular** solution.
#
# The sum of the two,
# <!-- Equation labels as ordinary links -->
# <div id="_auto9"></div>
#
# $$
# \begin{equation}
# v_y=v_{y,p}+v_{y,h},
# \label{_auto9} \tag{9}
# \end{equation}
# $$
# is a solution of the total equation because of the linear nature of
# the differential equation. One has now found a *general* solution
# encompassing all solutions, because it both satisfies the general
# equation (like the particular solution), and has an arbitrary constant
# that can be adjusted to fit any initial condition (like the homogneous
# solution). If the equation were not linear, e.g if there were a term
# such as $v_y^2$ or $v_y\dot{v}_y$, this technique would not work.
#
# ## More details
#
# Returning to the example above, the homogenous solution is the same as
# that for $v_x$, because there was no gravitational acceleration in
# that case,
# <!-- Equation labels as ordinary links -->
# <div id="_auto10"></div>
#
# $$
# \begin{equation}
# v_{y,h}=Be^{-\gamma t}.
# \label{_auto10} \tag{10}
# \end{equation}
# $$
# In this case a particular solution is one with constant velocity,
# <!-- Equation labels as ordinary links -->
# <div id="_auto11"></div>
#
# $$
# \begin{equation}
# v_{y,p}=-g/\gamma.
# \label{_auto11} \tag{11}
# \end{equation}
# $$
# Note that this is the terminal velocity of a particle falling from a
# great height. The general solution is thus,
# <!-- Equation labels as ordinary links -->
# <div id="_auto12"></div>
#
# $$
# \begin{equation}
# v_y=Be^{-\gamma t}-g/\gamma,
# \label{_auto12} \tag{12}
# \end{equation}
# $$
# and one can find $B$ from the initial velocity,
# <!-- Equation labels as ordinary links -->
# <div id="_auto13"></div>
#
# $$
# \begin{equation}
# v_{0y}=B-g/\gamma,~~~B=v_{0y}+g/\gamma.
# \label{_auto13} \tag{13}
# \end{equation}
# $$
# Plugging in the expression for $B$ gives the $y$ motion given the initial velocity,
# <!-- Equation labels as ordinary links -->
# <div id="_auto14"></div>
#
# $$
# \begin{equation}
# v_y=(v_{0y}+g/\gamma)e^{-\gamma t}-g/\gamma.
# \label{_auto14} \tag{14}
# \end{equation}
# $$
# It is easy to see that this solution has $v_y=v_{0y}$ when $t=0$ and
# $v_y=-g/\gamma$ when $t\rightarrow\infty$.
#
# One can also integrate the two equations to find the coordinates $x$
# and $y$ as functions of $t$,
# $$
# \begin{eqnarray}
# x&=&\int_0^t dt'~v_{0x}(t')=\frac{v_{0x}}{\gamma}\left(1-e^{-\gamma t}\right),\\
# \nonumber
# y&=&\int_0^t dt'~v_{0y}(t')=-\frac{gt}{\gamma}+\frac{v_{0y}+g/\gamma}{\gamma}\left(1-e^{-\gamma t}\right).
# \end{eqnarray}
# $$
# If the question was to find the position at a time $t$, we would be
# finished. However, the more common goal in a projectile equation
# problem is to find the range, i.e. the distance $x$ at which $y$
# returns to zero. For the case without a drag force this was much
# simpler. The solution for the $y$ coordinate would have been
# $y=v_{0y}t-gt^2/2$. One would solve for $t$ to make $y=0$, which would
# be $t=2v_{0y}/g$, then plug that value for $t$ into $x=v_{0x}t$ to
# find $x=2v_{0x}v_{0y}/g=v_0\sin(2\theta_0)/g$. One follows the same
# steps here, except that the expression for $y(t)$ is more
# complicated. Searching for the time where $y=0$, and we get
# <!-- Equation labels as ordinary links -->
# <div id="_auto15"></div>
#
# $$
# \begin{equation}
# 0=-\frac{gt}{\gamma}+\frac{v_{0y}+g/\gamma}{\gamma}\left(1-e^{-\gamma t}\right).
# \label{_auto15} \tag{15}
# \end{equation}
# $$
# This cannot be inverted into a simple expression $t=\cdots$. Such
# expressions are known as "transcendental equations", and are not the
# rare instance, but are the norm. In the days before computers, one
# might plot the right-hand side of the above graphically as
# a function of time, then find the point where it crosses zero.
#
# Now, the most common way to solve for an equation of the above type
# would be to apply Newton's method numerically. This involves the
# following algorithm for finding solutions of some equation $F(t)=0$.
#
# 1. First guess a value for the time, $t_{\rm guess}$.
#
# 2. Calculate $F$ and its derivative, $F(t_{\rm guess})$ and $F'(t_{\rm guess})$.
#
# 3. Unless you guessed perfectly, $F\ne 0$, and assuming that $\Delta F\approx F'\Delta t$, one would choose
#
# 4. $\Delta t=-F(t_{\rm guess})/F'(t_{\rm guess})$.
#
# 5. Now repeat step 1, but with $t_{\rm guess}\rightarrow t_{\rm guess}+\Delta t$.
#
# If the $F(t)$ were perfectly linear in $t$, one would find $t$ in one
# step. Instead, one typically finds a value of $t$ that is closer to
# the final answer than $t_{\rm guess}$. One breaks the loop once one
# finds $F$ within some acceptable tolerance of zero. A program to do
# this will be added shortly.
#
# ## Motion in a Magnetic Field
#
#
# Another example of a velocity-dependent force is magnetism,
# $$
# \begin{eqnarray}
# \boldsymbol{F}&=&q\boldsymbol{v}\times\boldsymbol{B},\\
# \nonumber
# F_i&=&q\sum_{jk}\epsilon_{ijk}v_jB_k.
# \end{eqnarray}
# $$
# For a uniform field in the $z$ direction $\boldsymbol{B}=B\hat{z}$, the force can only have $x$ and $y$ components,
# $$
# \begin{eqnarray}
# F_x&=&qBv_y\\
# \nonumber
# F_y&=&-qBv_x.
# \end{eqnarray}
# $$
# The differential equations are
# $$
# \begin{eqnarray}
# \dot{v}_x&=&\omega_c v_y,\omega_c= qB/m\\
# \nonumber
# \dot{v}_y&=&-\omega_c v_x.
# \end{eqnarray}
# $$
# One can solve the equations by taking time derivatives of either equation, then substituting into the other equation,
# $$
# \begin{eqnarray}
# \ddot{v}_x=\omega_c\dot{v_y}=-\omega_c^2v_x,\\
# \nonumber
# \ddot{v}_y&=&-\omega_c\dot{v}_x=-\omega_cv_y.
# \end{eqnarray}
# $$
# The solution to these equations can be seen by inspection,
# $$
# \begin{eqnarray}
# v_x&=&A\sin(\omega_ct+\phi),\\
# \nonumber
# v_y&=&A\cos(\omega_ct+\phi).
# \end{eqnarray}
# $$
# One can integrate the equations to find the positions as a function of time,
# $$
# \begin{eqnarray}
# x-x_0&=&\int_{x_0}^x dx=\int_0^t dt v(t)\\
# \nonumber
# &=&\frac{-A}{\omega_c}\cos(\omega_ct+\phi),\\
# \nonumber
# y-y_0&=&\frac{A}{\omega_c}\sin(\omega_ct+\phi).
# \end{eqnarray}
# $$
# The trajectory is a circle centered at $x_0,y_0$ with amplitude $A$ rotating in the clockwise direction.
#
# The equations of motion for the $z$ motion are
# <!-- Equation labels as ordinary links -->
# <div id="_auto16"></div>
#
# $$
# \begin{equation}
# \dot{v_z}=0,
# \label{_auto16} \tag{16}
# \end{equation}
# $$
# which leads to
# <!-- Equation labels as ordinary links -->
# <div id="_auto17"></div>
#
# $$
# \begin{equation}
# z-z_0=V_zt.
# \label{_auto17} \tag{17}
# \end{equation}
# $$
# Added onto the circle, the motion is helical.
#
# Note that the kinetic energy,
# <!-- Equation labels as ordinary links -->
# <div id="_auto18"></div>
#
# $$
# \begin{equation}
# T=\frac{1}{2}m(v_x^2+v_y^2+v_z^2)=\frac{1}{2}m(\omega_c^2A^2+V_z^2),
# \label{_auto18} \tag{18}
# \end{equation}
# $$
# is constant. This is because the force is perpendicular to the
# velocity, so that in any differential time element $dt$ the work done
# on the particle $\boldsymbol{F}\cdot{dr}=dt\boldsymbol{F}\cdot{v}=0$.
#
# One should think about the implications of a velocity dependent
# force. Suppose one had a constant magnetic field in deep space. If a
# particle came through with velocity $v_0$, it would undergo cyclotron
# motion with radius $R=v_0/\omega_c$. However, if it were still its
# motion would remain fixed. Now, suppose an observer looked at the
# particle in one reference frame where the particle was moving, then
# changed their velocity so that the particle's velocity appeared to be
# zero. The motion would change from circular to fixed. Is this
# possible?
#
# The solution to the puzzle above relies on understanding
# relativity. Imagine that the first observer believes $\boldsymbol{B}\ne 0$ and
# that the electric field $\boldsymbol{E}=0$. If the observer then changes
# reference frames by accelerating to a velocity $\boldsymbol{v}$, in the new
# frame $\boldsymbol{B}$ and $\boldsymbol{E}$ both change. If the observer moved to the
# frame where the charge, originally moving with a small velocity $v$,
# is now at rest, the new electric field is indeed $\boldsymbol{v}\times\boldsymbol{B}$,
# which then leads to the same acceleration as one had before. If the
# velocity is not small compared to the speed of light, additional
# $\gamma$ factors come into play,
# $\gamma=1/\sqrt{1-(v/c)^2}$. Relativistic motion will not be
# considered in this course.
#
#
#
# ## Sliding Block tied to a Wall
#
# Another classical case is that of simple harmonic oscillations, here represented by a block sliding on a horizontal frictionless surface. The block is tied to a wall with a spring. If the spring is not compressed or stretched too far, the force on the block at a given position $x$ is
# $$
# F=-kx.
# $$
# The negative sign means that the force acts to restore the object to an equilibrium position. Newton's equation of motion for this idealized system is then
# $$
# m\frac{d^2x}{dt^2}=-kx,
# $$
# or we could rephrase it as
# <!-- Equation labels as ordinary links -->
# <div id="eq:newton1"></div>
#
# $$
# \frac{d^2x}{dt^2}=-\frac{k}{m}x=-\omega_0^2x,
# \label{eq:newton1} \tag{19}
# $$
# with the angular frequency $\omega_0^2=k/m$.
#
# The above differential equation has the advantage that it can be solved analytically with solutions on the form
# $$
# x(t)=Acos(\omega_0t+\nu),
# $$
# where $A$ is the amplitude and $\nu$ the phase constant. This provides in turn an important test for the numerical
# solution and the development of a program for more complicated cases which cannot be solved analytically.
#
#
#
# ## Simple Example, Block tied to a Wall
#
# With the position $x(t)$ and the velocity $v(t)=dx/dt$ we can reformulate Newton's equation in the following way
# $$
# \frac{dx(t)}{dt}=v(t),
# $$
# and
# $$
# \frac{dv(t)}{dt}=-\omega_0^2x(t).
# $$
# We are now going to solve these equations using first the standard forward Euler method. Later we will try to improve upon this.
#
#
# ## Simple Example, Block tied to a Wall
#
# Before proceeding however, it is important to note that in addition to the exact solution, we have at least two further tests which can be used to check our solution.
#
# Since functions like $cos$ are periodic with a period $2\pi$, then the solution $x(t)$ has also to be periodic. This means that
# $$
# x(t+T)=x(t),
# $$
# with $T$ the period defined as
# $$
# T=\frac{2\pi}{\omega_0}=\frac{2\pi}{\sqrt{k/m}}.
# $$
# Observe that $T$ depends only on $k/m$ and not on the amplitude of the solution.
#
#
# ## Simple Example, Block tied to a Wall
#
# In addition to the periodicity test, the total energy has also to be conserved.
#
# Suppose we choose the initial conditions
# $$
# x(t=0)=1\hspace{0.1cm} \mathrm{m}\hspace{1cm} v(t=0)=0\hspace{0.1cm}\mathrm{m/s},
# $$
# meaning that block is at rest at $t=0$ but with a potential energy
# $$
# E_0=\frac{1}{2}kx(t=0)^2=\frac{1}{2}k.
# $$
# The total energy at any time $t$ has however to be conserved, meaning that our solution has to fulfil the condition
# $$
# E_0=\frac{1}{2}kx(t)^2+\frac{1}{2}mv(t)^2.
# $$
# We will derive this equation in our discussion on [energy conservation](https://mhjensen.github.io/Physics321/doc/pub/energyconserv/html/energyconserv.html).
#
# ## Simple Example, Block tied to a Wall
#
# An algorithm which implements these equations is included below.
# * Choose the initial position and speed, with the most common choice $v(t=0)=0$ and some fixed value for the position.
#
# * Choose the method you wish to employ in solving the problem.
#
# * Subdivide the time interval $[t_i,t_f] $ into a grid with step size
# $$
# h=\frac{t_f-t_i}{N},
# $$
# where $N$ is the number of mesh points.
# * Calculate now the total energy given by
# $$
# E_0=\frac{1}{2}kx(t=0)^2=\frac{1}{2}k.
# $$
# * Choose ODE solver to obtain $x_{i+1}$ and $v_{i+1}$ starting from the previous values $x_i$ and $v_i$.
#
# * When we have computed $x(v)_{i+1}$ we upgrade $t_{i+1}=t_i+h$.
#
# * This iterative process continues till we reach the maximum time $t_f$.
#
# * The results are checked against the exact solution. Furthermore, one has to check the stability of the numerical solution against the chosen number of mesh points $N$.
#
# ## Simple Example, Block tied to a Wall, python code
#
# The following python program ( code will be added shortly)
# +
#
# This program solves Newtons equation for a block sliding on
# an horizontal frictionless surface.
# The block is tied to the wall with a spring, so N's eq takes the form:
#
# m d^2x/dt^2 = - kx
#
# In order to make the solution dimless, we set k/m = 1.
# This results in two coupled diff. eq's that may be written as:
#
# dx/dt = v
# dv/dt = -x
#
# The user has to specify the initial velocity and position,
# and the number of steps. The time interval is fixed to
# t \in [0, 4\pi) (two periods)
#
| doc/pub/week4/ipynb/.ipynb_checkpoints/week4-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AI4Finance-Foundation/FinRL/blob/master/FinRL_Raytune_for_Hyperparameter_Optimization_RLlib%20Models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ZMPanjastOY4"
#Installing FinRL
# %%capture
# !pip install git+https://github.com/AI4Finance-LLC/FinRL-Library.git
# + id="W_0SxBYTtWNB"
# %%capture
# !pip install "ray[tune]" optuna
# + id="kW4g9mfwMl7K"
# %%capture
# !pip install int_date==0.1.8
# + [markdown] id="lPh7bRBVL9u3"
# #Importing libraries
# + id="AnFm0-vntYQw" colab={"base_uri": "https://localhost:8080/"} outputId="a0a02d75-faaf-4ea4-d38a-b1f0fb3d6103"
#Importing the libraries
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# matplotlib.use('Agg')
import datetime
import optuna
# %matplotlib inline
from finrl.apps import config
from finrl.finrl_meta.preprocessor.yahoodownloader import YahooDownloader
from finrl.finrl_meta.preprocessor.preprocessors import FeatureEngineer, data_split
from finrl.finrl_meta.env_stock_trading.env_stocktrading_np import StockTradingEnv as StockTradingEnv_numpy
from finrl.drl_agents.rllib.models import DRLAgent as DRLAgent_rllib
from stable_baselines3.common.vec_env import DummyVecEnv
from finrl.finrl_meta.data_processor import DataProcessor
from finrl.plot import backtest_stats, backtest_plot, get_daily_return, get_baseline
import ray
from pprint import pprint
from ray.rllib.agents.ppo import PPOTrainer
from ray.rllib.agents.ddpg import DDPGTrainer
from ray.rllib.agents.a3c import A2CTrainer
from ray.rllib.agents.a3c import a2c
from ray.rllib.agents.ddpg import ddpg, td3
from ray.rllib.agents.ppo import ppo
from ray.rllib.agents.sac import sac
import sys
sys.path.append("../FinRL-Library")
import os
import itertools
from ray import tune
from ray.tune.suggest import ConcurrencyLimiter
from ray.tune.schedulers import AsyncHyperBandScheduler
from ray.tune.suggest.optuna import OptunaSearch
from ray.tune.registry import register_env
import time
from typing import Dict, Optional, Any
# + id="F6DvqEVi3rxv"
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR):
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
# if not os.path.exists("./" + "tuned_models"):
# os.makedirs("./" + "tuned_models")
# + [markdown] id="rUTc0CApMCQP"
# ##Defining the hyperparameter search space
#
# 1. You can look up [here](https://docs.ray.io/en/latest/tune/key-concepts.html#search-spaces) to learn how to define hyperparameter search space
# 2. Jump over to this [link](https://github.com/DLR-RM/rl-baselines3-zoo/blob/master/utils/hyperparams_opt.py) to find the range of different hyperparameter
# 3. To learn about different hyperparameters for different algorithms for RLlib models, jump over to this [link](https://docs.ray.io/en/latest/rllib-algorithms.html)
# + id="c5luix7ZydIG"
def sample_ddpg_params():
return {
"buffer_size": tune.choice([int(1e4), int(1e5), int(1e6)]),
"lr": tune.loguniform(1e-5, 1),
"train_batch_size": tune.choice([32, 64, 128, 256, 512])
}
def sample_a2c_params():
return{
"lambda": tune.choice([0.1,0.3,0.5,0.7,0.9,1.0]),
"entropy_coeff": tune.loguniform(0.00000001, 0.1),
"lr": tune.loguniform(1e-5, 1)
}
def sample_ppo_params():
return {
"entropy_coeff": tune.loguniform(0.00000001, 0.1),
"lr": tune.loguniform(5e-5, 1),
"sgd_minibatch_size": tune.choice([ 32, 64, 128, 256, 512]),
"lambda": tune.choice([0.1,0.3,0.5,0.7,0.9,1.0])
}
# + id="Yb3PMaAZ2gUy"
MODELS = {"a2c": a2c, "ddpg": ddpg, "td3": td3, "sac": sac, "ppo": ppo}
# + [markdown] id="ZWG4u7NsOI98"
# ## Getting the training and testing environment
# + id="HmEAS3Vmt2d2"
def get_train_env(start_date, end_date, ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name, if_vix = True,
**kwargs):
#fetch data
DP = DataProcessor(data_source, **kwargs)
data = DP.download_data(ticker_list, start_date, end_date, time_interval)
data = DP.clean_data(data)
data = DP.add_technical_indicator(data, technical_indicator_list)
if if_vix:
data = DP.add_vix(data)
price_array, tech_array, turbulence_array = DP.df_to_array(data, if_vix)
train_env_config = {'price_array':price_array,
'tech_array':tech_array,
'turbulence_array':turbulence_array,
'if_train':True}
return train_env_config
# + id="Sx6O3qevuaDC"
#Function to calculate the sharpe ratio from the list of total_episode_reward
def calculate_sharpe(episode_reward:list):
perf_data = pd.DataFrame(data=episode_reward,columns=['reward'])
perf_data['daily_return'] = perf_data['reward'].pct_change(1)
if perf_data['daily_return'].std() !=0:
sharpe = (252**0.5)*perf_data['daily_return'].mean()/ \
perf_data['daily_return'].std()
return sharpe
else:
return 0
def get_test_config(start_date, end_date, ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name, if_vix = True,
**kwargs):
DP = DataProcessor(data_source, **kwargs)
data = DP.download_data(ticker_list, start_date, end_date, time_interval)
data = DP.clean_data(data)
data = DP.add_technical_indicator(data, technical_indicator_list)
if if_vix:
data = DP.add_vix(data)
price_array, tech_array, turbulence_array = DP.df_to_array(data, if_vix)
test_env_config = {'price_array':price_array,
'tech_array':tech_array,
'turbulence_array':turbulence_array,'if_train':False}
return test_env_config
def val_or_test(test_env_config,agent_path,model_name,env):
episode_total_reward = DRL_prediction(model_name,test_env_config,
env = env,
agent_path=agent_path)
return calculate_sharpe(episode_total_reward),episode_total_reward
# + id="HM12Fz7IrN4P"
TRAIN_START_DATE = '2014-01-01'
TRAIN_END_DATE = '2019-07-30'
VAL_START_DATE = '2019-08-01'
VAL_END_DATE = '2020-07-30'
TEST_START_DATE = '2020-08-01'
TEST_END_DATE = '2021-10-01'
# + id="tjrZFwhLsHHH"
technical_indicator_list =config.TECHNICAL_INDICATORS_LIST
model_name = 'a2c'
env = StockTradingEnv_numpy
ticker_list = ['TSLA']
data_source = 'yahoofinance'
time_interval = '1D'
# + id="ZwzmewVuyp6m" colab={"base_uri": "https://localhost:8080/"} outputId="80f8ebb3-fbac-46ac-887b-ac37a9bdf6cd"
train_env_config = get_train_env(TRAIN_START_DATE, VAL_END_DATE,
ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name)
# + [markdown] id="pplgHdQtOOQH"
# ## Registering the environment
# + id="QS0ytuI8KFf5"
from ray.tune.registry import register_env
env_name = 'StockTrading_train_env'
register_env(env_name, lambda config: env(train_env_config))
# + [markdown] id="0toGh9-_OThw"
# ## Running tune
# + id="SbF64_hRsqhT"
MODEL_TRAINER = {'a2c':A2CTrainer,'ppo':PPOTrainer,'ddpg':DDPGTrainer}
if model_name == "ddpg":
sample_hyperparameters = sample_ddpg_params()
elif model_name == "ppo":
sample_hyperparameters = sample_ppo_params()
elif model_name == "a2c":
sample_hyperparameters = sample_a2c_params()
def run_optuna_tune():
algo = OptunaSearch()
algo = ConcurrencyLimiter(algo,max_concurrent=4)
scheduler = AsyncHyperBandScheduler()
num_samples = 10
training_iterations = 100
analysis = tune.run(
MODEL_TRAINER[model_name],
metric="episode_reward_mean", #The metric to optimize for tuning
mode="max", #Maximize the metric
search_alg = algo,#OptunaSearch method which uses Tree Parzen estimator to sample hyperparameters
scheduler=scheduler, #To prune bad trials
config = {**sample_hyperparameters,
'env':'StockTrading_train_env','num_workers':1,
'num_gpus':1,'framework':'torch'},
num_samples = num_samples, #Number of hyperparameters to test out
stop = {'training_iteration':training_iterations},#Time attribute to validate the results
verbose=1,local_dir="./tuned_models",#Saving tensorboard plots
# resources_per_trial={'gpu':1,'cpu':1},
max_failures = 1,#Extra Trying for the failed trials
raise_on_failed_trial=False,#Don't return error even if you have errored trials
keep_checkpoints_num = num_samples-5,
checkpoint_score_attr ='episode_reward_mean',#Only store keep_checkpoints_num trials based on this score
checkpoint_freq=training_iterations#Checpointing all the trials
)
print("Best hyperparameter: ", analysis.best_config)
return analysis
# + id="zDz4GUMLuSUE" colab={"base_uri": "https://localhost:8080/", "height": 875} outputId="9751b541-0805-4cb5-9759-9c4c5817ba96"
analysis = run_optuna_tune()
# + [markdown] id="6d3a8-KROYJ_"
# ## Best config, directory and checkpoint for hyperparameters
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="OGDP01DcCR9Z" outputId="a7fc74da-7a98-4d4e-b4ac-49e00c0fdd69"
best_config = analysis.get_best_config(metric='episode_reward_mean',mode='max')
best_config
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="Awbo9S2sZbOv" outputId="71e9fd9b-23ee-4f19-8b30-63da1add087e"
best_logdir = analysis.get_best_logdir(metric='episode_reward_mean',mode='max')
best_logdir
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="wa-dilLhHGEd" outputId="f1432e32-f049-4605-c836-da1a6436f2bb"
best_checkpoint = analysis.best_checkpoint
best_checkpoint
# + id="RgcBMJBzAhZl"
# sharpe,df_account_test,df_action_test = val_or_test(TEST_START_DATE, TEST_END_DATE, ticker_list, data_source, time_interval,
# technical_indicator_list, env, model_name,best_checkpoint, if_vix = True)
# + colab={"base_uri": "https://localhost:8080/"} id="tO2MmLVTZWs-" outputId="56022129-ffe8-4f57-96c4-efc473a7140b"
test_env_config = get_test_config(TEST_START_DATE, TEST_END_DATE, ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name)
# + id="Dt0mhUOgeWtX"
sharpe,account,actions = val_or_test(test_env_config,agent_path,model_name,env)
# + id="Wis62wQaYHTR"
def DRL_prediction(
model_name,
test_env_config,
env,
model_config,
agent_path,
env_name_test='StockTrading_test_env'
):
env_instance = env(test_env_config)
register_env(env_name_test, lambda config: env(test_env_config))
model_config['env'] = env_name_test
# ray.init() # Other Ray APIs will not work until `ray.init()` is called.
if model_name == "ppo":
trainer = MODELS[model_name].PPOTrainer(config=model_config)
elif model_name == "a2c":
trainer = MODELS[model_name].A2CTrainer(config=model_config)
elif model_name == "ddpg":
trainer = MODELS[model_name].DDPGTrainer(config=model_config)
elif model_name == "td3":
trainer = MODELS[model_name].TD3Trainer(config=model_config)
elif model_name == "sac":
trainer = MODELS[model_name].SACTrainer(config=model_config)
try:
trainer.restore(agent_path)
print("Restoring from checkpoint path", agent_path)
except BaseException:
raise ValueError("Fail to load agent!")
# test on the testing env
state = env_instance.reset()
episode_returns = list() # the cumulative_return / initial_account
episode_total_assets = list()
episode_total_assets.append(env_instance.initial_total_asset)
done = False
while not done:
action = trainer.compute_single_action(state)
state, reward, done, _ = env_instance.step(action)
total_asset = (
env_instance.amount
+ (env_instance.price_ary[env_instance.day] * env_instance.stocks).sum()
)
episode_total_assets.append(total_asset)
episode_return = total_asset / env_instance.initial_total_asset
episode_returns.append(episode_return)
ray.shutdown()
print("episode return: " + str(episode_return))
print("Test Finished!")
return episode_total_assets
# + colab={"base_uri": "https://localhost:8080/"} id="mS_6EclCc-rR" outputId="029a8c98-3628-4db8-c251-50eff1d8aa4f"
episode_total_assets = DRL_prediction(
model_name,
test_env_config,
env,
best_config,
best_checkpoint,
env_name_test='StockTrading_test_env')
# + id="uRgs0r2Udbvn"
print('The test sharpe ratio is: ',calculate_sharpe(episode_total_assets))
df_account_test = pd.DataFrame(data=episode_total_assets,columns=['account_value'])
| FinRL_Raytune_for_Hyperparameter_Optimization_RLlib Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dagster
# language: python
# name: dagster
# ---
import dagstermill
import pandas as pd
import sklearn.ensemble
import matplotlib.pyplot as plt
# + tags=["parameters"]
context = dagstermill.get_context()
df = pd.read_csv('https://raw.githubusercontent.com/mwaskom/seaborn-data/master/iris.csv')
# -
df.head()
X = df[['sepal_length', 'sepal_width']]
y = df['petal_width']
model = sklearn.ensemble.RandomForestRegressor()
fit = model.fit(X,y)
context.log.info("Fitted random forest model!")
score = fit.score(X, y)
context.log.info("Random forest model has score {}".format(score))
pred_y = fit.predict(X)
plt.scatter(y, pred_y)
plt.title("True vs. predicted")
plt.xlabel("True petal width")
plt.ylabel("Predicted petal width")
plt.show()
| python_modules/libraries/dagstermill/dagstermill/examples/notebooks/tutorial_RF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
import pandas as pd
import geoplot
import geopandas
import matplotlib.pyplot as plt
# %matplotlib inline
from shapely.geometry import Polygon
import warnings
warnings.filterwarnings(action="ignore")
#Check geopandas version
geopandas.__version__
#Set figure size and font size
plt.rcParams["figure.figsize"]=(12,10)
plt.rcParams["font.size"]=12
# # Getting the canvas ready
world = geopandas.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
world.plot()
fig=geoplot.polyplot(world,projection=geoplot.crs.Orthographic())
plt.show()
europe=world[world.continent=="Europe"]
europe.plot()
europe=europe[(europe.name!="Russia") & (europe.name!="Iceland")]
europe.plot()
# ## Clip French Guinea off of map of Europe
# Create a custom polygon
polygon = Polygon([(-25,35), (40,35), (40,75),(-25,75)])
poly_gdf = geopandas.GeoDataFrame([1], geometry=[polygon], crs=world.crs)
fig,ax=plt.subplots()
ax=europe.plot(ax=ax)
poly_gdf.plot(edgecolor="red",ax=ax, alpha=0.1)
plt.show()
#Clip polygon from the map of Europe
europe=geopandas.clip(europe, polygon) #Input and feature to be clipped
europe.plot()
# ## Data Preparation
# Source: https://ourworldindata.org/grapher/carbon-intensity-electricity
df=pd.read_csv("carbon-intensity-electricity.csv")
df
df["Entity"].unique()
len(df["Entity"].unique())
europe.name.unique()
len(europe.name.unique())
list(europe.name.unique())
# ### Check if countries in df are present in europe geodataframe or not
#Initialize an empty list for countries which are present in df, but not in europe
unmatched=[]
for country in list(df["Entity"].unique()):
if country in (list(europe.name.unique())):
pass
else:
unmatched.append(country)
unmatched
df["Year"].dtypes
# +
#Retain values for 2010, 2015 and 2020 only
df=df[(df.Year==2000)|(df.Year==2005)|(df.Year==2010) | (df.Year==2015) | (df.Year==2020)]
#Drop Code column
df.drop("Code",axis=1, inplace=True)
#Remove unmatched items from df
df=df[(df.Entity!="Cyprus") & (df.Entity!="EU-27") & (df.Entity!="EU27+1") & (df.Entity!="Malta")]
#Make pivot
df=pd.pivot_table(df, index="Entity",columns="Year")
df
# -
df.columns=["2000","2005","2010","2015","2020"]
df=df.reset_index()
df.rename({"Entity":"name"},axis=1,inplace=True)
df
selected_countries=europe[europe.name.isin(list(df.name))]
selected_countries
selected_countries=selected_countries.merge(df,on="name",how="left")
selected_countries
# +
#Range of Variable you see as map color. Here I select the minimum and maximum of all the years selected.
vmin=selected_countries[["2000","2005","2010","2015","2020"]].min().min()
vmax=selected_countries[["2000","2005","2010","2015","2020"]].max().max()
# +
fig,axs=plt.subplots(2,3) #3 columns and 1 row
fig.suptitle("Emissions Intensity from electricity generation in Europe 2000-2020", fontweight="bold",fontsize=15)
#Adjust space betweeen rows
plt.subplots_adjust(bottom=0.2, top=0.9, hspace=0.25)
axs[0,0]=europe.plot(color="whitesmoke",edgecolor="black",ax=axs[0,0])
selected_countries.plot("2000",cmap="Reds",edgecolor="black",ax=axs[0,0], vmin=vmin, vmax=vmax)
axs[0,0].set_title("2000")
axs[0,0].xaxis.set_visible(False)
axs[0,1]=europe.plot(color="whitesmoke",edgecolor="black",ax=axs[0,1])
selected_countries.plot("2005",cmap="Reds",edgecolor="black",ax=axs[0,1], vmin=vmin, vmax=vmax)
axs[0,1].set_title("2005")
axs[0,1].xaxis.set_visible(False)
axs[0,1].yaxis.set_visible(False)
axs[0,2]=europe.plot(color="whitesmoke",edgecolor="black",ax=axs[0,2])
selected_countries.plot("2010",cmap="Reds",edgecolor="black",ax=axs[0,2], vmin=vmin, vmax=vmax)
axs[0,2].set_title("2010")
axs[0,2].xaxis.set_visible(False)
axs[0,2].yaxis.set_visible(False)
axs[1,0]=europe.plot(color="whitesmoke",edgecolor="black",ax=axs[1,0])
selected_countries.plot("2015",cmap="Reds",edgecolor="black",ax=axs[1,0], vmin=vmin, vmax=vmax)
axs[1,0].set_title("2015")
axs[1,1]=europe.plot(color="whitesmoke",edgecolor="black",ax=axs[1,1])
selected_countries.plot("2020",cmap="Reds",edgecolor="black",ax=axs[1,1], vmin=vmin, vmax=vmax)
axs[1,1].set_title("2020")
axs[1,1].yaxis.set_visible(False)
axs[1,2]=europe.plot(color="whitesmoke",edgecolor="black",ax=axs[1,2])
axs[1,2].set_title("Future?")
axs[1,2].yaxis.set_visible(False)
# add colorbar
cax = fig.add_axes([0.92, 0.2, 0.03, 0.7]) #[left, bottom, width, height]
sm = plt.cm.ScalarMappable(cmap='Reds', norm=plt.Normalize(vmin=vmin, vmax=vmax))
# fake up the array of the scalar mappable
sm._A = []
lgd=fig.colorbar(sm, cax=cax).set_label("gCO$_2$e/kWh", rotation=0,y=1.05, labelpad=-35)
plt.savefig("Emissions Intensity over the past two decades.jpeg",
dpi=300)
plt.show()
# -
pd.set_option("display.max_columns",None)
df
df
# +
df.set_index("name",inplace=True)
df=df.T
df[["Estonia","Poland","Sweden","United Kingdom","Germany","France"]].plot(marker="o",linestyle="dashed",figsize=(8,6))
plt.title("Carbon Intensity of Electricity Generation Of Selective Countries")
plt.xlabel("Years"); plt.ylabel("gCO$_2$/kWh")
lgd=plt.legend(bbox_to_anchor=(1,1))
plt.savefig("Selective Countries Carbon Intensity",
dpi=300,
bbox_extra_artists=(lgd,),
bbox_inches="tight")
plt.show()
# -
selected_countries.head()
#Getting the lan and lat here from geometry data
selected_countries['coordinates']=selected_countries['geometry'].apply(lambda x: x.representative_point().coords[:][0])
selected_countries.head()
# ## Analysing carbon intensity in 2020
# +
fig, ax=plt.subplots()
ax=europe.plot(color="whitesmoke",
edgecolor='black',
ax=ax)
selected_countries.plot("2020",
ax=ax,edgecolor="black",
cmap="Reds",
legend=True)
#Add names of county here
for idx, row in selected_countries.iterrows():
plt.annotate(s=row["name"], xy=row['coordinates'],
horizontalalignment='center', color='black',fontsize=10, fontweight='light')
plt.title("Carbon Intensity of Electricity Generation in Europe in 2020 (gCO$_2$/kWh)")
plt.savefig("2020 figure", dpi=300)
#cax = fig.add_axes([0.92, 0.2, 0.03, 0.7])
#sm=plt.cm.ScalarMappable(cmap='Reds',
# norm=plt.Normalize(vmin=selected_countries["2020"].min(), vmax=selected_countries["2020"].max()))
#lgd=fig.colorbar(sm,cax=cax).set_label("gCO$_2$e/kWh", rotation=0,y=1.05, labelpad=-35)
# -
| script/Plotting emissions intensity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GeoNet FDSN webservice with Obspy demo - Event Service
# In this tutorial we will look at geting seismic event data. By the end of this notebooks you should be able to make a map like the one below. <img src="event.png">
# This demo introduces some simple code that requests data using [GeoNet's FDSN webservices](http://www.geonet.org.nz/data/tools/FDSN) and the [obspy module](https://github.com/obspy/obspy/wiki) in python. This notebook uses Python 3.
#
# ### Getting Started - Import Modules
from obspy import UTCDateTime
from obspy.clients.fdsn import Client as FDSN_Client
from obspy import read_inventory
# ### Define GeoNet FDSN client
client = FDSN_Client("GEONET")
# ## Accessing Earthquake Information
# Use the **event** service to access earthquake parameters from the catalogue.
#
# This example requests the Kaikoura earthquake and aftershocks for 24 hours following the event, within a 0.5 degree radius of the epicenter. It then prints a list and plots the locations on a map
starttime = "2016-11-13 11:00:00.000"
endtime = "2016-11-14 11:00:00.000"
cat = client.get_events(starttime=starttime, endtime=endtime,latitude=-42.693,longitude=173.022,maxradius=0.5,minmagnitude=5)
print(cat)
_=cat.plot(projection="local")
# Single events can be requested using their PublicID, which is available from the GeoNet website. This example will demonstrate how to get additional information about the [Kaikoura Earthquake](http://www.geonet.org.nz/earthquake/2016p858000).
cat = client.get_events(eventid="2016p858000")
print(cat)
ev = cat[0]
print(ev)
# Print out a summary of the information for the preferred origin.
origin = ev.origins[0]
print(origin)
# List all available magnitudes and their associated uncertainties
for m in range(len(ev.magnitudes)):
if 'uncertainty' in ev.magnitudes[m].mag_errors and ev.magnitudes[m].mag_errors['uncertainty'] != None and ev.magnitudes[m].resource_id == ev.preferred_magnitude_id:
print('%s = %f +/- %f - Preferred magnitude' % (ev.magnitudes[m].magnitude_type, ev.magnitudes[m].mag, ev.magnitudes[m].mag_errors['uncertainty']))
elif 'uncertainty' in ev.magnitudes[m].mag_errors and ev.magnitudes[m].mag_errors['uncertainty'] != None:
print('%s = %f +/- %f' % (ev.magnitudes[m].magnitude_type, ev.magnitudes[m].mag, ev.magnitudes[m].mag_errors['uncertainty']))
else:
print('%s = %f' % (ev.magnitudes[m].magnitude_type, ev.magnitudes[m].mag))
# List all arrivals used to locate the earthquake.
print(origin.arrivals[0])
print(ev.picks[0])
for p in range(len(ev.picks)):
for a in range(len(origin.arrivals)):
if ev.picks[p].resource_id == origin.arrivals[a].pick_id:
if origin.arrivals[a].time_weight > 0:
print(ev.picks[p].time, ev.picks[p].waveform_id['station_code'], origin.arrivals[a].distance, origin.arrivals[a].phase, origin.arrivals[a].time_residual)
| Seismic_Data/Python/GeoNet_FDSN_demo_event.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualização de dados - Oficina Aula 2.1
#
# Olá, amante da sétima arte! Durante a aula você aprendeu a representar quantidades relacionadas a dados categóricos utilizando gráficos de barras através das bibliotecas pandas, matplotlib e seaborn. Com esse conhecimento você poderá explorar e obter muitas informações sobre conjuntos de dados que possuam dados categóricos. Agora é hora de exercitar o que você aprendeu! Nesta oficina você deve continuar o conjunto de dados sobre filmes lançados entre 2007 e 2011. Crie um Jupyter Notebook e responda cada questão com código em uma célula.
#
# 1. Importe as bibliotecas pandas, matplotlib, seaborn e use o comando mágico %matplotlib inline para exibir as visualizações no notebook. Carregue os dados em um DataFrame. Os dados estão disponibilizados no link:
# dados.
# Você pode carregar os dados pela URI ou salvar em sua máquina e carregá-los localmente.
#
# 2. Crie uma visualização que mostre o orçamento de todos os filmes do conjunto de dados. Qual o filme com maior orçamento e com menor orçamento?
# 3. Crie uma visualização que mostre a bilheteria de todos os filmes do conjunto de dados. Qual o filme com maior bilheteria e com menor bilheteria?
# 4. Crie uma visualização que mostre a quantidade de filmes por gênero em cada ano. Em que anos foram feitos menos filmes de ação? E em que ano foram feitos menos filmes de animação?
# 5. Crie uma visualização que mostre a média de orçamento de filmes por gênero em cada ano. Em que ano os filmes de ação tiveram maior média de orçamento? E em que ano os filmes de animação tiveram a maior média de orçamento?
#
# Resposta da questão 1
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
filmes_uri = 'https://raw.githubusercontent.com/emanueles/datavis-course/master/assets/files/observable/movies.json'
filmes = pd.read_json(filmes_uri)
filmes.head()
# Resposta da questão 2
contagens = filmes.groupby(by='Genre').count()
plt.figure(figsize=(15,5))
sns.barplot(x="Genre", y="Budget_M", color = 'darkviolet', ci = 'sd', data=filmes)
plt.title('Orçamento de todos os filmes')
plt.show()
# Resposta da questão 3
filmes.sort_values(by='Worldwide_Gross_M', ascending = False).head()
plt.figure(figsize=(15,5))
sns.barplot(x="Genre", y="Worldwide_Gross_M", color = 'darkred', ci = 'sd', data=filmes)
plt.title('A bilheteria de todos os filmes')
plt.show()
# Resposta da questão 4
plt.figure(figsize=(15,8))
plt.title('Filmes por gênero em cada ano')
sns.countplot(x="Year", hue= 'Genre', data=filmes)
plt.show()
# Resposta da questão 5
plt.figure(figsize=(15,8))
plt.title('Média de orçamento por gênero em cada ano')
sns.barplot(x="Year", y = 'Budget_M' , ci = None, hue= 'Genre', data=filmes)
plt.show()
# Desenvolvido por **<NAME>**, Rondonópolis-MT, 21/01/2021.
| Visualizacao_de_dados/Oficina_Aula_2.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../ancillarydata/logos/LundUniversity_C2line_RGB.png" width="150" align="left"/>
# <br>
# <img src="../ancillarydata/logos/Icos_Logo_CMYK_Regular_SMpng.png" width="327" align="right"/>
# <br>
# <a id='introduction'></a>
# <br>
# <a id='intro'></a>
#
# <br>
# <meta content="text/html; charset=UTF-8">
#
# <style>td{padding: 3px;}
# </style>
#
# <table style="width: 100%">
# <colgroup>
# <col span="1" style="width: 17%;">
# <col span="1" style="width: 83%;">
# </colgroup>
# <tr>
# <th><font size="2">Notebook developed by:</font></th>
# <td>
# <div style="float:left;">
# <font size="2">
# <NAME> & <NAME> as part of ICOS Carbon Portal's educational notebooks
# </font>
# </div>
# </td>
# </tr>
# <tr>
# <th><font size="2">Data provided by:</font></th>
# <td><div style="float:left;"><font size="2"><NAME> at Lund University dept. of Physical Geography & Ecosystem Science, ICOS Sweden
# </font></div></td>
# </tr>
# </table>
# </font>
#
# <br>
#
#
# # Drought in Europe Summer 2018
# ## <span style="color:#224C98">Drought Analysis with ICOS Ecosystem Data</span>
#
# One of the consequences of climate change are more frequent occurrences of drought. Dry spells are expected to happen more often and last for a longer time. This has been observed in a number of areas in Southern and Central Europe. During the summer of 2018, even parts of Northern Europe (incl. Sweden) were affected by drought.
#
#
# The figure bellow shows the results of a drought-index called [SPEI06](http://spei.csic.es/home.html) applied over Europe for the month of August for different years: 2003, 2010, 2015 and 2018 [[1](#references)]. The dark red colors represent areas affected by drought whilst the darker blue colors mark areas with high levels of precipitation.
#
# <br>
# <br>
# <img src="../ancillarydata/images/htm_drought/diagrams/torka_sommaren_2018.png" width="" align="center">
# <br>
# <br>
#
# #### But what is drought and how can we define it?
# ## I. Drought - Definition
# Drought can be defined as a period that is characterized by water scarcity caused by lower than average levels of precipitation [[2](#references)]. Periods of drought can occur during different seasons. However, in most cases, droughts occur during periods with unusually high temperatures and insufficient rainfall.
#
# Droughts can be divided into different categories [[3](#references)]:
#
# - **Meteorological droughts:** occur when the levels of precipitation are lower than average for an extended period of time.
# - **Agricultural droughts:** occur when the soil water content goes below a certain degree and the crops can no longer absorb the water that is contained in the soil. Such events may stress the crops and lead to lost harvests.
# - **Hydrological droughts:** occur when the water level in water reserves like aquifers, lakes and reservoirs fall bellow a certain threshold.
# - **Socioeconomic droughts:** happen when dry spells affect the society. For instance, when the demand for an economic good exceeds supply due to a weather-related shortage in water availability.
# <br>
# <br>
#
# ## II. Drought Impact on Trees
# Droughts have a significant impact on plants. Here the focus will be set on the impact of droughts on trees. During periods characterized by uncommonly high temperatures and low water content in the soil, trees tend to cease to grow and their resistance towards illnesses, like e.g. fungal diseases or pest infestations, weakens [[4](#references)]. The first sign of drought affliction on trees is that their leaves start to close even during day and will eventually fall off, if the tree does not get access to water in time [[4](#references)]. Other trees might behave differently. For instance, their leaves might start to crumble instead of closing. When trees sense that water supplies are scarce, they react by closing their [stomata](https://en.wikipedia.org/wiki/Stoma), which ultimately brings their photosynthetic activity to a halt [[4](#references)]. It is in this state that the tree might lose its leaves driven by the same mechanisms that cause this effect during autumn. Trees that lose their leaves while they are still green, lose a lot of nutrients. Additionally, water-stressed trees have a limited capacity to transfer nutrients to all parts of the tree, which, in some cases, may lead to tree malnutrition.
#
# Trees lose green leaves during severe drought events. Under other circumstances, the leaves have time to turn yellow before they fall or turn brown before they dry out close to the tree trunk. Depending on the type of tree, leaf shedding may occur during the drought or just after rehydration [[4](#references)]. In Nordic forests [birch](https://en.wikipedia.org/wiki/Birch)- and the [European spruce](https://en.wikipedia.org/wiki/Picea_abies)-trees are more sensitive to drought [[5](#references)]. The needle-like leaves of the European spruce turn brown and fall off while birch-leaves also fall off before they have had the chance to obtain their autumn coloration [[5](#references)]. In general, young or fast-growing trees tend to be more severely affected compared to older or more slow-growing trees [[5](#references)]. When plants cease to photosynthesize, their leaves are afflicted by ([chlorosis](https://en.wikipedia.org/wiki/Chlorosis)), lose their green color ([chlorophyll](https://en.wikipedia.org/wiki/Chlorophyll)) and begin to show signs of autumn coloration. When plants reach this state, due to the occurrence of a drought, it is an indication that even their roots have been damaged [[4](#references)]. Entire branches may dry out and die. Plants that have survived a severe drought event once, are more likely to survive a new event. This is one of the reasons why young trees are usually more severely affected [[4](#references)].
# <br>
# <br>
#
# ## III. Drought Impact on Decomposers/Detritivores and their Activity
# Decomposers and detritivores are organisms that break down organic matter (e.g. dead twigs and leaves, decaying organisms or excrements, etc.) to carbon dioxide, methane, carboxylic acid, water and heat [[6](#references)]. Typical examples of decomposers are fungi and bacteria. Typical examples of detritivores are earthworms, woodlice and sea cucumbers. The difference between detritivores and decomposers is that detritivores have to ingest nutrients in order to break them down to organic matter. Through their activity, decomposers and detritivores release carbon to the atmosphere. If the occurrence of a drought causes the soil water content to drop below a certain threshold, the environment can become too dry for decomposers and detritivores. To survive they will limit their level of activity and consequently reduce the amount of carbon they release to the atmosphere.
# <br>
# <br>
#
# ## IV. Drought Impact on the Carbon Ballance of Ecosystems
# The amount of carbon dioxide in the atmosphere may increase during an extended period of drought. This can be attributed to the change in the behaviour of plants once the climate gets drier and the temperature too high. Droughts make the soil become drier, the air less moist and thus plants are forced to save water in order to preserve their existing tissue and survive. This is achieved by limiting or totally ceasing their photosyntetic activity, which in turn means that they limit or totally stop their intake of carbon dioxide [[7](#references)]. Subsequently, plants absorb less carbon dioxide from their environment during drought periods compared to other times. An increase in the frequence of drought events can therefore contribute to the global warming effect and, thus, create a vicious circle of extreme temperatures [[7](#references)].
#
# <br>
# <br>
# <a id='toc'></a>
#
# ## V. Notebook - Table of Contents
# This notebook is dedicated to using ICOS Ecosystem Data from Hyltemossa Research Station in Southern Sweden, to study how the drought during the summer of 2018 affected the vegetation and the carbon balance in the surrounding area. The temporal resolution of the data extends from January 1st 2015 to December 31st 2018.
#
#
# Another objective of this notebook is to introduce basic principles of Python Programming. More in particular, users are going to learn how to:
#
# - Read in csv files to Python structures (Pandas DataFrames)
# - Clean and Harmonize data
# - Process data and Compute Basic Statistics
# - Plot Data by Creating Static and Interactive Plots
#
# <br>
#
#
# The notebook is divided in the following main parts:
#
# - [Introduction](#intro)
# <br>
# <br>
# - [Instructions on How to Use the Notebook](#instructions_how_to_use_nb)
# <br>
# <br>
# - [Data from ICOS Hyltemossa Station](#data_HTM_station)
# <br>
# <br>
# - [Python Programming](#py_programming)
# <br>
# <br>
# - [References](#references)
# <br>
# <br>
# <a id='instructions_how_to_use_nb'></a>
# <br>
# <br>
#
# ## VI. Instructions on How to Use the Notebook
# ### <span style="color:#cb4154">Run the Notebook</span>
# In order to run this Jupyter Notebook, go to the menu at the top of the page and click on **Kernel** and then **Restart & Run All**.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/restart_run_all_nb.png" width="270" align="center"/>
#
# <br>
# <br>
#
#
# Use the links in the **Table of Contents** above to navigate to different parts of the notebook. Parts of the notebook that are long include an additional tabel of contents with links to their corresponding subparts. Use the links to quickly navigate from one subpart to another. It is also possible to scroll. Once you have clicked on **Restart & Run All**, it will be possible to navigate to the plots of the different programming parts and interact with them using widgets. Widget is the Pythonic name for an interactive element (e.g. dropdown lists, radiobuttons, execution buttons, etc.). A more detailed description on how to interact with the widgets and the interactive plots of every part of the analysis is presented in the beginning of that part.
# <br>
# ### <span style="color:#cb4154">Run a Single Code-Cell</span>
# A Jupyter Notebook consists of code-cells. It is possible to write Python code in a code-cell and then run it by clicking on **Run** in the menu at the top of the page.
#
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/run_code_cell_nb.png" width="540" align="center"/>
#
# <br>
# <br>
#
# Observe that only one code-cell will be executed and this is the code-cell that was active when you clicked on **Run**. You can activate a code-cell just by clicking on it. An active code-cell is highlighted in blue or green color (see image bellow).
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/active_code_cell_nb.png" width="610" align="center"/>
#
# <br>
# <br>
#
# It is also possible to write Markup code in a Jupyter Notebook code-cell. For instance, the instructions you are reading here are written in a Markup code-cell that includes markup text and HTML code. When you are writing Python code in a code-cell make sure that the cell is a Python code-cell. The type of the currently active code-cell is shown in the dropdown list on the menu bar at the top of the page (see figure). A code-cell that includes Python code should be marked as **Code**.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/check_type_of_code_cell_nb.png" width="540" align="center"/>
#
# <br>
# <br>
#
# ### <span style="color:#cb4154">Add a Code-Cell</span>
# Click on **"+"** in the menu to add a new code-cell under the current active code-cell.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/add_new_code_cell_nb.png" width="540" align="center"/>
#
# <br>
# <br>
#
# ### <span style="color:#cb4154">Delete a Code-Cell</span>
# If you wish to delete a code-cell, select the code-cell by clicking on it and then go to the menu at the top of the page and click on **Edit** --- > **Delete Cells**.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/delete_code_cell_nb.png" width="270" align="center"/>
#
# <br>
# <br>
#
# ### <span style="color:#cb4154">Stop Execution</span>
# If an execution is taking too long, you can stop your notebook from running by clicking on **Interrupt kernel** in the menu.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/interrupt_kernel_nb.png" width="540" align="center"/>
#
# <br>
# <br>
#
# Alternatively, another choice is to go to **Kernel** and click on **Interrupt**.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/interrupt_nb.png" width="440" align="center"/>
#
# <br>
# <br>
#
# ### <span style="color:#cb4154">Save Notebook</span>
# Click on **Save** freequently to save your work.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/save_nb.png" width="540" align="center"/>
#
# <br>
# <br>
#
# ### <span style="color:#cb4154">Download Notebook</span>
# If you wish to download the notebook as a Jupyter Notebook, go to the menu at the top of the page, click on **File** --- > **Save As...** --- > **Notebook(.ipynb)**.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/download_nb.png" width="270" align="center"/>
#
# <br>
# <br>
#
# If you wish to save your work as pure Python code, go the menu at the top of the page, click on **File** --- > **Save As...** --- > **Python(.py)**.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/nb_manual/download_nb_as_py_file.png" width="270" align="center"/>
#
# <br>
# <br>
#
# <br>
# <br>
# <br>
# <br>
# <a id='data_HTM_station'></a>
# <br>
# <br>
#
# ## VII. Data from ICOS Hyltemossa Research Station
# Areas in southern Sweden showed clear signs of drought damage during the summer of 2018. The analysis in this module is conducted using data from [Hyltemossa Research Station](https://www.icos-sweden.se/station_hyltemossa.html).
#
#
# The station is located near a 30-year old managed spruce-forest south of Perstorp, in northwestern Scania, Sweden. The station collects atmospheric and ecosystem measurements and is part of the [ICOS Sweden](https://www.icos-sweden.se/) research infrastructure. [ICOS](https://www.icos-ri.eu/) is an acronym for Integrated Carbon Observation System and is a European Research Infrastructure that has implemented a European measurement system for high quality and high precision greenhouse gas observations. The objective of ICOS is to create an extended network of measuring stations producing time series of high-quality data that will ultimately help to map the carbon balance of Europe. The [ICOS Carbon Portal](https://www.icos-cp.eu/) provides free and open access to all ICOS data.
#
# +
############################################################################################################
################## Python & Javascript Code - handling code visibility (entire document)####################
############################################################################################################
#Import module:
from IPython.display import HTML
HTML('''<script> $('div .input').hide()''')
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# +
############################################################################################################
############################### Python & Javascript Code - Hide code-cell ###################################
############################################################################################################
#Import modules:
from IPython.core.display import display, HTML
#Code that hides a single code-cell:
toggle_code_str = '''
<form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Hide/Show Code"></form>
'''
toggle_code_prepare_str = '''
<script>
function code_toggle() {
if ($('div.cell.code_cell.rendered.selected div.input').css('display')!='none'){
$('div.cell.code_cell.rendered.selected div.input').hide();
$('#toggleButton').val('Show Kod');
} else {
$('div.cell.code_cell.rendered.selected div.input').show();
$('#toggleButton').val('Hide Kod');
}
}
</script>
'''
display(HTML(toggle_code_prepare_str + toggle_code_str))
#Call function to hide code-cell:
def toggle_code():
display(HTML(toggle_code_str))
############################################################################################################
############################################################################################################
############################################################################################################
# +
#Import modules:
import folium
#Create map object:
m = folium.Map(location=[56.097991, 13.420181], zoom_start=7)
#Add marker:
folium.Marker(location=[56.097991, 13.420181],
popup='ICOS Hyltemossa Research Station',
icon=folium.Icon(color='darkred', icon='cloud')).add_to(m)
#Show map
m
# -
# ### Measured Variables from Hyltemossa Station
# For the purpose of this analysis, we will use a subset of the available variable measurements from Hyltemossa station. A list of the code (column-name in file), the title and the unit of these variables is presented below:
#
# - **TA_1_1_1**       --->    Air Temperature $(^oC$) <br/>
# - **P_1_1_1**       --->    Precipitation (mm) <br/>
# - **SWC_4_4_1**     --->    Soil Water Content (%) <br/>
# - **GPP_PI_1_1_1**     --->    Gross Primary Production ($\mu$$mol \: m^{-2}\:s^{-1}$)<br/>
# - **RECO_PI_1_1_1**    --->    Respiration ($\mu$$mol\: m^{-2}\:s^{-1}$) <br/>
# - **FC_PI_1_1_1**     --->     Carbon Flux ($\mu$$mol\: m^{-2}\:s^{-1}$) <br/>
# - **SW_IN_1_1_1**    --->     Incoming Shortwave Infrared (SW-IR) Solar Radiation - Light (W $m^{-2}$)
# <br>
# <br>
#
# Here is a brief explanation of what every variable stands for:
# #### <span style="color:#CD5C5C">What is Soil Water Content (SWC)?</span>
# Soil Water Content measures the proportion of water in the soil. In this case it is expressed as %.
# <br>
# <br>
# $$SWC = \frac{100 M_w}{M_s}$$
# <br>
# where:<br>
# SWC = Soil Water Content (%), <br>
# $M_w$ = mass of water in the soil (kg),<br>
# $M_s$ = mass of dry soil (kg)
# <br>
# <br>
#
# #### <span style="color:#CD5C5C">What is Photosynthesis ?</span>
# Photosynthesis is desgribed as the process by which a plant uses light energy to transform carbon dioxide, water and minerals into oxygen and energy-rich organic compounds. The chemical formula of photosynthesis is:
# <br>
# <br>
# $$6 H_2O + 6 CO_2 \rightarrow{} C_6H_{12}O_6 + 6O_2 $$
# <br>
# <br>
#
# #### <span style="color:#CD5C5C">What is Gross Primary Production (GPP) ?</span>
# The Gross Primary Production (GPP) of an ecosystem can be described as the amount of carbon that has been taken from the atmosphere by plants because of their photosynthetic activity.
# <br>
# <br>
#
# #### <span style="color:#CD5C5C">What is Respiration ?</span>
# Respiration can be described as the amount of carbon that is emitted from an ecosystem because of animal and plant respiration.
# <br>
# <br>
#
# #### <span style="color:#CD5C5C">What is Net Ecosystem Exchange (NEE) ?</span>
# Net Ecosystem Exchange (NEE) is a measure of the net exchange of Carbon between an ecosystem and the atmosphere per unit ground area. In simpler words, it is a balance between the total amount of carbon emitted and the total anount of carbon absorbed by an ecosystem per unit ground area.
# <br>
# <br>
#
# #### <span style="color:#CD5C5C">What is Shortwave Infrared Incoming Solar Radiation (SW-IR)?</span>
# Shortwave Infrared Incoming Solar Radiation can be described as the amount of incoming solar radiation in the shortwave infrared wavelengths (1.4 - 3 μm) in a given area, during a given time period. Hyltemossa station utilizes measuring equipment that measures the amount of incoming shortwave infrared solar radiation to estimate the available amount of solar energy the vegetation can interact with. The availability of solar energy is essential for the plants to photosynthesize.
# <br>
# <br>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#toc">Back to top</a>
# </div>
# <a id='py_programming'></a>
# <br>
# <br>
#
# ## VIII. Python Programming
# This part presents basic principles of Python programming. The focus is set on reading csv files to Python specific structures such as Pandas DataFrames (matrix) and processing this data using built-in methods. The built-in methods are used to filter data and produce basic statistics. The results are then visualized as interactive plots utilizing the [Bokeh](https://bokeh.pydata.org/en/latest/index.html) interactive visualization library.
#
#
# This part is divided into the following subparts:
# 1. [Import Python Modules](#import_py_modules)
#
#
# 2. [Prerequisites - Basic Programming Principles in Python](#python_intro)
#
#
# 3. [Define Global Variables](#python_global_var)
#
#
# 4. [Read csv-files into Python Pandas DataFrames](#csv2pandasdf)
#
#
# 5. [Update values in a Pandas DataFrame Column](#updatePandasDfCol)
#
#
# 6. [Handling Date and Time in Python - Python DateTime Objects](#CreateDatetimeObj)
#
#
# 7. [Add a column with DateTime-Objects in every Pandas DataFrame](#addDatetimeCol2pdDf)
#
#
# 8. [Indexing a Pandas DataFrame](#pandasDfSetIndex)
# 1. [Set a column of a Pandas DataFrame as index](#pandasDfSetIndex)
# 2. [Extract all rows from a Pandas DataFrame index-column](#pandasDfSearchWithIndex)
# 3. [How to index a Pandas DataFrame with DateTime Objects](#pandasDfSearchWithDateTimeIndex)
# 4. [How to filter a Pandas DataFrame using an index of DateTime Objects](#pandasDfSliceWithDateTimeIndex)
#
#
# 9. [Compute Statistics over a Pandas DataFrame Column](#pandasDfCalcStat)
# 1. [Compute the min, max, mean and standard deviation over all rows of a Pandas DataFrame column](#pandasDfCalcStatMinMaxMeanStDev)
# 2. [Compute the min, max, mean and standard deviation over a selection of rows of a Pandas DataFrame column](#pandasDfCalcStatMinMaxMeanStDevFiltered)
#
#
# 10. [Plot Data from a Pandas Dataframe with Bokeh](#bokeh_plot_df)
# 1. [Create an Interactive Plot from 2 Pandas DataFrame columns](#bokeh_plot_2_cols_from_df)
# 2. [Plot Statistics with Bokeh Visualization Library ](#bokeh_plot_stat_barplot)
# 3. [Create Plots with Cumulative Sums of Daily Totals and Daily Means per Year](#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro)
# 4. [Barplot with Incoming Shortwave-Infrared Solar Radiation (Daily Total) & GPP (Daily Total)](#bokeh_plot_daily_total_GPP_SWIR_per_year)
# 5. [Barplot with GPP (Daily Total) and Soil Water Content (Daily Mean)](#bokeh_plot_daily_total_GPP_daily_mean_SWC_per_year)
# 6. [Plot Daily Mean Soil Water Content with Daily Total Respiration and Daily Mean Air Temperature](#bokeh_plot_daily_total_RECO_daily_mean_SWC_and_TA_per_year)
# 7. [Plot Daily Mean Soil Water Content with Daily Total GPP and Daily Total Precipitation](#bokeh_plot_daily_mean_SWC_and_daily_total_GPP_and_Precip_per_year)
# 8. [Plot Daily Mean Soil Water Content and Air temperature with Daily Total GPP and Light (SW-IR)](#bokeh_plot_daily_mean_SWC_and_TA_and_daily_total_GPP_and_SWIR_per_year)
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#toc">Back to top</a>
# </div>
# <a id='import_py_modules'></a>
# <br>
#
# ### 1. Import Python Modules
# Python is a programming language that includes built-in methods. A module can be described as set of functions. To use these functions, you need to first import the module they belong to. Usually, modules are imported in the beginning of a Python-program.
#
# The next code-cell shows the syntax of how to import Python modules. It is possible to import a module using the syntax <code style="color:#CD5C5C">import math</code>. To import all functions from a module type <code style="color:#CD5C5C">from math import *</code>. However, this is considered bad practice, so it is best to avoid that. For importing a single function from a module type <code style="color:#CD5C5C">from datetime import datetime</code>. Some large modules may include more than one different packages of functions. For example, the <code style="color:#CD5C5C">bokeh</code> module includes a package of functions called <code style="color:#CD5C5C">plotting</code>, which in turn includes a function called <code style="color:#CD5C5C">figure</code>.
#
# When you import a module, it is possible to change its name after the keyword <code style="color:#CD5C5C">as</code>. Usually, the name provided after <code style="color:#CD5C5C">as</code> is an abbreviation of the modules official name. The following piece of code <code style="color:#CD5C5C">import pandas as pd</code>, will import a module called _pandas_ and change its name to _pd_. This way, you do not have to type the full name of the module when you call it in your code. Ultimately, by following this practice, your code will be easier to read.
# + deletable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Import modules:
import os
import numpy as np
import pandas as pd
import itertools
from datetime import datetime
import math
from bokeh.plotting import figure
from bokeh.models import ColumnDataSource, HoverTool, Label, Legend, SingleIntervalTicker, LinearAxis, Range1d
from bokeh.io import show, output_notebook
# -
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
#
# <a id='python_intro'></a>
# <br>
# <br>
#
# ### 2. Prerequisites - Basic Programming Principles in Python
# To understand the Python code in this notebook, you are expected to know the basic principle of the following concepts:
#
# - Global and Local Variables
# - Python Dictionaries
# - Python Lists
# - Python Tuples
# - Control Statements in Python
# - If-Statements
# - For-Loops
# - List Comprehensions
# - String Manipulation in Python
# - Functions
#
# If you are not familiar with the previous concepts or you want to brush-up your memory, you can read through the corresponding part in the <span style="color:green">**Quickstart to Python**</span>-notebook included in the same folder.
#
# <br>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <a id='python_global_var'></a>
# <br>
# <br>
#
# ### 3. Define Global Variables
# The next code-cell includes 5 global variables. All 5 global variables are Python dictionaries. Global variables should be handled carefully and if possible avoided, if no specific reason exists. In this implementation, we will make regular use of these variables and, thus we define them as global.
#
# The global variables here handle the format of numbers (superscript/subscript) and the association between the name, code and unit of ecosystem variables.
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Create a dictionary to transform numbers to their equivalent subscript or superscript representation:
SUB = str.maketrans("0123456789", "₀₁₂₃₄₅₆₇₈₉")
SUP = str.maketrans("0123456789", "⁰¹²³⁴⁵⁶⁷⁸⁹")
#Create a dictionary to store the variable names and their corresponding codes:
measurement_dict_eng = {'TA_1_1_1':'Air Temperature',
'FC_PI_1_1_1':'Carbon Flux (NEE)',
'GPP_PI_1_1_1':'Gross Primary Production',
'P_1_1_1':'Precipitation',
'RECO_PI_1_1_1':'Respiration',
'SW_IN_1_1_1':'SW-IR Incoming Solar Radiation',
'SWC_4_4_1':'Soil Water Content'}
#Create a dictionary to store the units related to every variable-code:
unit_dict = {'TA_1_1_1':'\u00b0C',
'FC_PI_1_1_1':'umol m-2 s-1'.replace('u', '\u03BC').translate(SUP),
'P_1_1_1':'mm',
'RECO_PI_1_1_1':'umol m-2 s-1'.replace('u', '\u03BC').translate(SUP),
'GPP_PI_1_1_1':'umol m-2 s-1'.replace('u', '\u03BC').translate(SUP),
'SWC_4_4_1':'%',
'SW_IN_1_1_1':'W/m2'.translate(SUP)}
#Create a dictionary to store the units related to every variable-code for daily aggregated computations:
unit_dict_daily = {'TA_1_1_1':'C\u00b0',
'FC_PI_1_1_1':'umol m-2'.replace('u', '\u03BC').translate(SUP),
'P_1_1_1':'mm',
'RECO_PI_1_1_1':'umol m-2'.replace('u', '\u03BC').translate(SUP),
'GPP_PI_1_1_1':'umol m-2'.replace('u', '\u03BC').translate(SUP),
'SWC_4_4_1':'%',
'SW_IN_1_1_1':'MJoules/m2'.translate(SUP)}
# -
# <a id='csv2pandasdf'></a>
# <br>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
#
# ### 4. Read csv-files into Python Pandas Dataframes
#
# #### <span style="color:#CD5C5C">What is a Pandas DataFrame ?</span>
# Python has a data structure called [Pandas DataFrame](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html). It consists of a 2-dimensional matrix which can vary in size (in terms of number of columns or number of rows). A Pandas DataFrame has the ability to store data belonging to different data types. It is, for example, permitted to store a column with strings, a column with integers and a column with floats in the same dataframe (see Figure). A Pandas DataFrame has indexed columns and rows. Columns can be indexed based on their name whilst rows can be indexed based on their row number or a specific index-value. A more detailed explanation of what a Pandas DataFrame index is, can be found in the <span style="color:green">**Quickstart to Python**</span>-notebook. For now, it is enough to envision an index as one of the columns in the dataframe, that include unique values for every row. For example, the column *Student ID* in the figure, could be used as an index. This is attributed to the fact that it is not possible for two students to have the same *Student ID*. In other words, the values in the aforementioned column, uniquely identify every row in the dataframe.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/diagrams/pandas_df_example_eng.png" width="550" align="center">
# <br>
# <br>
#
# The Pandas module (which was renamed to ```pd``` at import) includes a built-in method ```read_csv()``` to read a csv-file to a Pandas DataFrame. The next code-cell shows the Python syntax fo reading in data from a csv-file to a Pandas DataFrame.
#
# <u>**Syntax:**</u>
# <span style="color:blue">pandas.read_csv</span><span style="color:#992622">(path_to_file,<br/>
#               row_with_column_names,<br/>
#               num_of_rows_to_skip,<br/>
#               delimiter,<br/>
#               coding_system\*)</span>
# <br>
# <br>
# <span style="color:gray">\* By defining the coding system, it is possible to read special characters such as, for example, national characters like (å, ä, ö) or symbols like ($C^o$).</span>
# <br>
# <br>
# <br>
#
#
# <span style="color:blue">**Read Air Temperature, Respiration, GPP, NEE and SW-IR Data for 2015**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Set paths:
personal_home = '/data/project'
path_climbeco = personal_home+'/climbeco/data/'
path_htm = path_climbeco + 'htm/'
#Read ecosystem-data for 2015:
htm_eko_2015 = pd.read_csv(path_htm+'SE-Htm_2015_vs20190510.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the five first rows of the dataframe:
htm_eko_2015.head(5)
# -
# <br>
# <br>
#
# <span style="color:blue">**Read Air Temperature, Respiration, GPP, NEE and SW-IR Data for 2016**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Read ecosystem-data for 2016:
htm_eko_2016 = pd.read_csv(path_htm+'SE-Htm_2016.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the 2 first rows of the dataframe:
htm_eko_2016.head(2)
# -
# <br>
# <br>
#
# <span style="color:blue">**Read Air Temperature, Respiration, GPP, NEE and SW-IR Data for 2017**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Read ecosystem-data for 2017:
htm_eko_2017 = pd.read_csv(path_htm+'SE-Htm_2017.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the 2 first rows of the dataframe:
htm_eko_2017.head(2)
# -
# <br>
# <br>
#
# <span style="color:blue">**Read Air Temperature, Respiration, GPP, NEE and SW-IR Data for 2018**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Read ecosystem-data for 2018:
htm_eko_2018 = pd.read_csv(path_htm+'SE-Htm_2018_vs20190510.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the 2 first rows of the dataframe:
htm_eko_2018.head(2)
# -
# <br>
# <br>
#
# <span style="color:blue">**Read Soil Water Content, Precipitation and Soil Temperature Data for 2015**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Read ecosystem soil water content, soil temperature and precipitation-data for 2015:
#(Note that precipitation is measured in mm)
htm_eko_precip_2015 = pd.read_csv(path_htm+'SE-Htm_2015_TS_SWC_Prec.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the 2 first rows of the dataframe:
htm_eko_precip_2015.head(2)
# -
# <br>
# <br>
#
# <span style="color:blue">**Read Soil Water Content, Precipitation and Soil Temperature Data for 2016**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Read ecosystem soil water content, soil temperature and precipitation-data for 2016:
#(Note that precipitation is measured in mm)
htm_eko_precip_2016 = pd.read_csv(path_htm+'SE-Htm_2016_TS_SWC_Prec.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the 2 first rows of the dataframe:
htm_eko_precip_2016.head(2)
# -
# <br>
# <br>
#
# <span style="color:blue">**Read Soil Water Content, Precipitation and Soil Temperature Data for 2017**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Read ecosystem soil water content, soil temperature and precipitation-data for 2017:
#(Note that precipitation is measured in mm)
htm_eko_precip_2017 = pd.read_csv(path_htm+'SE-Htm_2017_TS_SWC_Prec.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the 2 first rows of the dataframe:
htm_eko_precip_2017.head(2)
# -
# <br>
# <br>
#
# <span style="color:blue">**Read Soil Water Content, Precipitation and Soil Temperature Data for 2018**</span>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Read ecosystem soil water content, soil temperature and precipitation-data for 2018:
#(Note that precipitation is measured in mm)
htm_eko_precip_2018 = pd.read_csv(path_htm+'SE-Htm_2018_TS_SWC_Prec.csv',
header=0,
skiprows=range(1,2),
sep=',',
encoding='windows-1252')
#Show the 2 first rows of the dataframe:
htm_eko_precip_2018.head(2)
# -
# <a id='updatePandasDfCol'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
#
# ### 5. Update values in a Pandas DataFrame Column
# One of the most important and time-consuming tasks when working with data analysis is cleaning and harmonizing data. From this aspect, it is important to be able to access data and alter values in a structured and automated way. Pandas DataFrames include such methods. This part presents how it is possible to update existing values in a Pandas DataFrame.
#
# Missing values in ICOS data have a fix-value that is equal to ```-9999.0```. Missing values can be a result of something gone wrong with a measurement. Often when you create a plot, you do not wish to include the missing values. In these cases, you have to convert the *missing values* to ```NaN```. ```NaN```is a numeric datatype that stands for *Not a Number* and doesn't represent a value. When the value of a field is set to ```NaN```, then this field is treated as an empty field.
#
# <br>
#
# <span style="color:blue">**Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def ecoToNan(df_data, variable, threshold):
#Import modules:
import pandas as pd
import numpy as np
from numpy import nan
#Set values under the threshold equal to NaN:
df_data.loc[df_data[variable] <threshold, [variable]] = np.nan
#Return dataframe:
return df_data
# -
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Convert all "missing values" (i.e. values=-9999.0) and negative GPP- or RECO-values to NaN:
#Air-Temperature (TA):
htm_eko_2015.loc[htm_eko_2015['TA_1_1_1'] <-9990.0, ['TA_1_1_1']] = np.nan
htm_eko_2016.loc[htm_eko_2016['TA_1_1_1'] <-9990.0, ['TA_1_1_1']] = np.nan
htm_eko_2017.loc[htm_eko_2017['TA_1_1_1'] <-9990.0, ['TA_1_1_1']] = np.nan
htm_eko_2018.loc[htm_eko_2018['TA_1_1_1'] <-9990.0, ['TA_1_1_1']] = np.nan
#Carbon Fluxes (NEE):
htm_eko_2015.loc[htm_eko_2015['FC_PI_1_1_1'] <-9990.0, ['FC_PI_1_1_1']] = np.nan
htm_eko_2016.loc[htm_eko_2016['FC_PI_1_1_1'] <-9990.0, ['FC_PI_1_1_1']] = np.nan
htm_eko_2017.loc[htm_eko_2017['FC_PI_1_1_1'] <-9990.0, ['FC_PI_1_1_1']] = np.nan
htm_eko_2018.loc[htm_eko_2018['FC_PI_1_1_1'] <-9990.0, ['FC_PI_1_1_1']] = np.nan
#Respiration (RECO):
htm_eko_2015.loc[htm_eko_2015['RECO_PI_1_1_1'] <0, ['RECO_PI_1_1_1']] = np.nan
htm_eko_2016.loc[htm_eko_2016['RECO_PI_1_1_1'] <0, ['RECO_PI_1_1_1']] = np.nan
htm_eko_2017.loc[htm_eko_2017['RECO_PI_1_1_1'] <0, ['RECO_PI_1_1_1']] = np.nan
htm_eko_2018.loc[htm_eko_2018['RECO_PI_1_1_1'] <0, ['RECO_PI_1_1_1']] = np.nan
#Gross Primary Production (GPP):
htm_eko_2015.loc[htm_eko_2015['GPP_PI_1_1_1'] <0, ['GPP_PI_1_1_1']] = np.nan
htm_eko_2016.loc[htm_eko_2016['GPP_PI_1_1_1'] <0, ['GPP_PI_1_1_1']] = np.nan
htm_eko_2017.loc[htm_eko_2017['GPP_PI_1_1_1'] <0, ['GPP_PI_1_1_1']] = np.nan
htm_eko_2018.loc[htm_eko_2018['GPP_PI_1_1_1'] <0, ['GPP_PI_1_1_1']] = np.nan
#Light (Short-Wave Infrared Incoming Solar Radiation):
htm_eko_2015.loc[htm_eko_2015['SW_IN_1_1_1'] <-9990.0, ['SW_IN_1_1_1']] = np.nan
htm_eko_2016.loc[htm_eko_2016['SW_IN_1_1_1'] <-9990.0, ['SW_IN_1_1_1']] = np.nan
htm_eko_2017.loc[htm_eko_2017['SW_IN_1_1_1'] <-9990.0, ['SW_IN_1_1_1']] = np.nan
htm_eko_2018.loc[htm_eko_2018['SW_IN_1_1_1'] <-9990.0, ['SW_IN_1_1_1']] = np.nan
#Precipitation:
htm_eko_precip_2015.loc[htm_eko_precip_2015['P_1_1_1'] <-9990.0, ['P_1_1_1']] = np.nan
htm_eko_precip_2016.loc[htm_eko_precip_2016['P_1_1_1'] <-9990.0, ['P_1_1_1']] = np.nan
htm_eko_precip_2017.loc[htm_eko_precip_2017['P_1_1_1'] <-9990.0, ['P_1_1_1']] = np.nan
htm_eko_precip_2018.loc[htm_eko_precip_2018['P_1_1_1'] <-9990.0, ['P_1_1_1']] = np.nan
#Soil Water Content:
htm_eko_precip_2015.loc[htm_eko_precip_2015['SWC_4_4_1'] <-9990.0, ['SWC_4_4_1']] = np.nan
htm_eko_precip_2016.loc[htm_eko_precip_2016['SWC_4_4_1'] <-9990.0, ['SWC_4_4_1']] = np.nan
htm_eko_precip_2017.loc[htm_eko_precip_2017['SWC_4_4_1'] <-9990.0, ['SWC_4_4_1']] = np.nan
htm_eko_precip_2018.loc[htm_eko_precip_2018['SWC_4_4_1'] <-9990.0, ['SWC_4_4_1']] = np.nan
# -
# <a id='CreateDatetimeObj'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
#
# ### 6. Handling Date and Time in Python - Python DateTime Objects
# Python has a built-in module (i.e. set of functions) called ```datetime``` that includes functions for handling dates and time. The module provides multiple options on how to process date and time. In this example, we will focus on how to create a DateTime object based on existing information of date and time. The existing information, in this case, is in String format.
#
# Click on the [link](https://docs.python.org/3/library/datetime.html) to get more detailed information regarding how you can work with Python DatTime objects.
#
# <br>
#
# <span style="color:blue">**Variable storing Date-inforation in a String format**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Variable containing date information as a String (text):
datum = '01/01/2015'
# -
# <br>
#
# <span style="color:blue">**Variable storing Time-information in a String format**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Variable containing time information as a String (text):
tid = '00:30'
# -
# <br>
#
# <span style="color:blue">**Create a Datetime Object**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Create a DateTime Object:
datetime_variable = datetime.strptime(datum + ' ' + tid, '%d/%m/%Y %H:%M')
#Show result:
datetime_variable
# -
# <a id='addDatetimeCol2pdDf'></a>
# <br>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
#
# ### 7. Add a column with DateTime-Objects in every Pandas DataFrame
# When you plot data and want to visualize time on the x-axis, then there are a number of visualization-libraries that request you to have time represented as a DateTime Object. A DateTime Object is a data structure that represents date and time in a specific format (e.g. "Y-m-d H:M:S" may stand for: "2019-01-01 08:05:00").
#
# The following functions include code that creates DateTime objects by combining existing information on date and time. Date and time, in this case, are stored as String variables. The functions below create a new column of DateTime objects in a Pandas DataFrame, based on the content of two existing columns, which, in turn, include information about the date and time of a measurement.
#
# <br>
#
# <span style="color:blue">**Function**</span>
# <span style="color:darkred"> ---- > Time Format: **dd/mm/YY HH:MM** </span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def icosEcoAddDatetimeObjYMDHM(df_data):
#To be used when time is expressed as: HH:MM
#Import modules:
from datetime import datetime
#Add a column with datetime obj:
df_data['DateTime'] = [datetime.strptime((df_data.date.iloc[i]+ ' ' +
df_data.time.iloc[i]),'%d/%m/%Y %H:%M')
for i in range(len(df_data))]
#Return dataframe:
return df_data
# -
# <br>
#
# <span style="color:blue">**Function**</span>
# <span style="color:darkred"> ---- > Time Format: **dd/mm/YY HH:MM:SS** </span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def icosEcoAddDatetimeObjYMDHMS(df_data):
#To be used when time is expressed as: HH:MM:SS
#Import modules:
from datetime import datetime
#Add a column with datetime obj:
df_data['DateTime'] = [datetime.strptime((df_data.date.iloc[i]+ ' ' +
df_data.time.iloc[i]),'%d/%m/%Y %H:%M:%S')
for i in range(len(df_data))]
#Return dataframe:
return df_data
# -
# <br>
#
# <span style="color:green">**Call Function(s)**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Add a column with datetime objects to every dataframe:
HTM_eko_2015 = icosEcoAddDatetimeObjYMDHM(htm_eko_2015)
HTM_eko_2016 = icosEcoAddDatetimeObjYMDHM(htm_eko_2016)
HTM_eko_2017 = icosEcoAddDatetimeObjYMDHMS(htm_eko_2017)
HTM_eko_2018 = icosEcoAddDatetimeObjYMDHM(htm_eko_2018)
HTM_eko_precip_2015 = icosEcoAddDatetimeObjYMDHM(htm_eko_precip_2015)
HTM_eko_precip_2016 = icosEcoAddDatetimeObjYMDHM(htm_eko_precip_2016)
HTM_eko_precip_2017 = icosEcoAddDatetimeObjYMDHM(htm_eko_precip_2017)
HTM_eko_precip_2018 = icosEcoAddDatetimeObjYMDHM(htm_eko_precip_2018)
#Show results:
HTM_eko_2015.head(5)
# -
# <a id='pandasDfSetIndex'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
#
# <br>
# <br>
# <br>
# <br>
#
# ### 8. Indexing Pandas DataFrames
# In this part we will learn how to index Pandas DataFrames. Indexing is away to extract values on demand based on certain criteria. Here you will learn how to define an index and how to use it in order to extract values from a Pandas DataFrame.
# <br>
# <br>
# #### 8.1 Set a Column of a Pandas DataFrame as Index
# Pandas includes a built-in method to set a column as an index:
# <br>
# <br>
# $$ dataframe\_name.set\_index(column\_name) $$
# <br>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Set the "DateTime"-column as index in all pandas dataframes:
HTM_eko_2015_indexed = HTM_eko_2015.set_index('DateTime')
HTM_eko_2016_indexed = HTM_eko_2016.set_index('DateTime')
HTM_eko_2017_indexed = HTM_eko_2017.set_index('DateTime')
HTM_eko_2018_indexed = HTM_eko_2018.set_index('DateTime')
HTM_eko_precip_2015_indexed = HTM_eko_precip_2015.set_index('DateTime')
HTM_eko_precip_2016_indexed = HTM_eko_precip_2016.set_index('DateTime')
HTM_eko_precip_2017_indexed = HTM_eko_precip_2017.set_index('DateTime')
HTM_eko_precip_2018_indexed = HTM_eko_precip_2018.set_index('DateTime')
#Show example:
HTM_eko_2015_indexed.head(4)
# -
# <br>
# <br>
# <a id='pandasDfSearchWithIndex'></a>
#
# #### 8.2. Extract all rows from a Pandas DataFrame index-column
# Type the following code to retrieve all values from a Pandas DataFrame index:
#
# <br>
# <br>
# $$dataframe\_name.index.values$$
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Get index values:
HTM_eko_2015_indexed.index.values
# -
# <br>
# <br>
# <a id='pandasDfSearchWithDateTimeIndex'></a>
#
# #### 8.3. How to index a Pandas DataFrame with DateTime Objects
# Use the following syntax to extract all data for a selected date and time:
# <br>
# <br>
# $$dataframe\_name[dataframe\_name.index==datetime(year, month, day, time, minute, second)]$$
# <br>
# <br>
# Observe that for this piece of code to work, your Pandas DataFrame must have a column of DateTime Objects as index.
# <br>
# <br>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Show all rows that include values for the given date and time:
HTM_eko_2015_indexed[HTM_eko_2015_indexed.index==datetime(2015, 6, 1)]
# -
# <br>
# <br>
# <a id='pandasDfSliceWithDateTimeIndex'></a>
#
# #### 8.4. How to filter a Pandas DataFrame using an index of DateTime Objects
# It is possible to filter a Pandas DataFrame either using its index or based on the values in its columns. The following piece of code shows how to extract data for a given time period. The syntax is:
#
# <br>
# <br>
# $$dataframe\_name[datetime(year_{start}, month_{start}, day_{start}):datetime(year_{end}, month_{end}, day_{end}, hour_{end}, minute_{end})]$$
# <br>
#
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Filter all pandas dataframes to extract data for the summermonths (June-August):
HTM_summermonths_2015 = HTM_eko_2015_indexed[datetime(2015, 6, 1):datetime(2015, 8, 31, 23, 30)]
HTM_summermonths_2016 = HTM_eko_2016_indexed[datetime(2016, 6, 1):datetime(2016, 8, 31, 23, 30)]
HTM_summermonths_2017 = HTM_eko_2017_indexed[datetime(2017, 6, 1):datetime(2017, 8, 31, 23, 30)]
HTM_summermonths_2018 = HTM_eko_2018_indexed[datetime(2018, 6, 1):datetime(2018, 8, 31, 23, 30)]
#Filter all pandas dataframes (precipitation) to extract data for the summermonths (June-August):
HTM_P_summermonths_2015 = HTM_eko_precip_2015_indexed[datetime(2015, 6, 1):datetime(2015, 8, 31, 23, 30)]
HTM_P_summermonths_2016 = HTM_eko_precip_2016_indexed[datetime(2016, 6, 1):datetime(2016, 8, 31, 23, 30)]
HTM_P_summermonths_2017 = HTM_eko_precip_2017_indexed[datetime(2017, 6, 1):datetime(2017, 8, 31, 23, 30)]
HTM_P_summermonths_2018 = HTM_eko_precip_2018_indexed[datetime(2018, 6, 1):datetime(2018, 8, 31, 23, 30)]
#Show results:
HTM_summermonths_2015.head(5)
# -
# <a id='pandasDfCalcStat'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
# <br>
# <br>
#
# ### 9. Compute Statistics over a Pandas DataFrame Column
# It is possible to calculate the minimum ```min()```, maximum ```max()```, mean ```mean()``` and standard deviation ```std()``` of all values in a column of a Pandas DataFrame. The Pandas syntax for that is:
# <br>
# <br>
# $$dataframe\_name.column\_name.function()$$
# <br>
# <br>
# The Pandas built-in method for computing e.g. the minimum value of a column, performs the exact same process as the Python code in the following code-cell.
#
#
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Assume that the following list is a Pandas DataFrame column:
pandas_kolumn = [1, 4, -1, 10, 37]
#Define and initialize help variable:
total_min = 20
#Loop that loops through all the values of the list:
for i in pandas_kolumn:
#Compare the current value from the list to the value in the help variable:
if(i<total_min):
total_min=i
#Show result:
total_min
# -
# <a id='pandasDfCalcStatMinMaxMeanStDev'></a>
# <br>
#
# #### 9.1. Compute the min, max, mean and standard deviation over all rows of a Pandas DataFrame column
#
# <br>
#
# <span style="color:blue">**Min Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the lowest air temperature of 2015:
HTM_eko_2015_indexed.TA_1_1_1.min()
# -
# <br>
#
# <span style="color:blue">**Max Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the highest air temperature of 2015:
HTM_eko_2015_indexed.TA_1_1_1.max()
# -
# <br>
#
# <span style="color:blue">**Mean Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the average air temperature of 2015:
HTM_eko_2015_indexed.TA_1_1_1.mean()
# -
# <br>
#
# <span style="color:blue">**Standard Deviation Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the standard deviation of air temperatures for 2015:
HTM_eko_2015_indexed.TA_1_1_1.std()
# -
# <a id='pandasDfCalcStatMinMaxMeanStDevFiltered'></a>
# <br>
#
# #### 9.2. Compute the min, max, mean and standard deviation over a selection of rows of a Pandas Dataframe column
# <br>
#
# <span style="color:blue">**Min Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the lowest air temperature for the summer months of 2015:
HTM_eko_2015_indexed[datetime(2015, 6, 1):datetime(2015, 8, 31, 23, 30)].TA_1_1_1.min()
# -
# <br>
#
# <span style="color:blue">**Max Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the highest air temperature for the summer months of 2015:
HTM_summermonths_2015.TA_1_1_1.max()
# -
# <br>
#
# <span style="color:blue">**Mean Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the average air temperature for the summer months of 2015:
HTM_summermonths_2015.TA_1_1_1.mean()
# -
# <br>
#
# <span style="color:blue">**Standard Deviation Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute the standard deviation of air temperatures for the summer months of 2015:
HTM_summermonths_2015.TA_1_1_1.std()
# -
# <a id='bokeh_plot_df'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
# <br>
# <br>
#
# ### 10. Plot Data from a Pandas DataFrame with Bokeh
# Bokeh is a Python Library for creating visualizations of Data. There is a large variety of modules available for many different types of graphs. If you are interested to discover more about Bokeh click on the [link](https://bokeh.pydata.org/en/latest/index.html).
#
# In this part we will present a set of different Bokeh plots, depending on our purpose. We will start by creating an interactive plot that shows how the values of a variable change in time. Then we are going to display some basic statistics using barplots. Finally, we are going to create different combinations of barplots and line-graphs to show how ecosystem variable values from different years differ compared to the corresponding ecosystem variable values for the year the drought occurred. The latter plots will also include an interactive legend that allows the user to switch layers on and off, to enahnce the readability of the plot.
#
# The user will be able to control the content of the plots by using widgets. Widgets are the Python name for controls like dropdown lists, radio-buttons etc.
#
# This part is divided into the following subparts:
# - [Create an Interactive Plot from 2 Pandas DataFrame columns](#bokeh_plot_2_cols_from_df)
# - [Plot Statistics with Bokeh Visualization Library](#bokeh_plot_stat_barplot)
# - [Create Plots with Cumulative Sums of Daily Totals and Daily Means per Year](#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro)
# - [Barplot with Incoming Shortwave-Infrared Solar Radiation (Daily Total) & GPP (Daily Total)](#bokeh_plot_daily_total_GPP_SWIR_per_year)
# - [Barplot with GPP (Daily Total) and Soil Water Content (Daily Mean)](#bokeh_plot_daily_total_GPP_daily_mean_SWC_per_year)
# - [Plot Daily Mean Soil Water Content with Daily Total Respiration and Daily Mean Air Temperature](#bokeh_plot_daily_total_RECO_daily_mean_SWC_and_TA_per_year)
# - [Plot Daily Mean Soil Water Content with Daily Total GPP and Daily Total Precipitation](#bokeh_plot_daily_mean_SWC_and_daily_total_GPP_and_Precip_per_year)
# - [Plot Daily Mean Soil Water Content and Air temperature with Daily Total GPP and Light (SW-IR)](#bokeh_plot_daily_mean_SWC_and_TA_and_daily_total_GPP_and_SWIR_per_year)
#
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
# <a id='bokeh_plot_2_cols_from_df'></a>
# <br>
#
# #### 10.1. Create an Interactive Plot from 2 Pandas DataFrame column
# This part presents how to plot data from a Pandas DataFrame using Bokeh. The plot presents how different types of ecosystem variables change during the period of one year. The user is able to change the content of the plot using a set of widgets (see Figure below). There are two dropdown widgets that control the selection of year and ecosystem variable, a color-picker that controls the color of the line in the plot and, finally, an execution-button used to update the content of the plot with the user's choice in all of the aforementioned widgets. It is also possible to interact with the content of a plot using the Bokeh Plot Toolbox, located in the right part of the plot.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/bokeh_plots/bokeh_plot_widgets.png" width="700" align="center">
#
# <br>
# <br>
#
# In order to use a tool in the Bokeh Plot ToolBox, you have to activate it. You can activate a tool just by clicking on it. An active tool is always highlighted with a blue line next to its symbol. For instance, in the figure above, the Pan-tool is the only active tool.
#
#
# Use the ```Pan-tool``` to move the content of the plot up or down, right or left.
#
#
# Use the ```Box Zoom-tool``` to zoom-in on a rectangular selected area.
#
#
# Use the ```Wheel Zoom-tool``` to zoom-in over an area in the plot just by scrolling.
#
#
# Press the ```Reset``` button to restore the plot to its initial state.
#
#
# Press the ```Save``` button to save a copy of the plot to your computer.
#
#
# Press the ```Hover``` button and hover with your mouse over the plot to see annotations.
#
#
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_2_cols_from_df_plot">[Go to Plot]</a>
# </div>
#
# <br>
#
# <span style="color:blue">**Plotting Function**</span>
# + deletable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def plotIcosEcoIndexedDF(df, variable, color):
#Create a figure object:
p = figure(plot_width=900,
plot_height=500,
x_axis_label='Time (UTC)',
y_axis_label= measurement_dict_eng[variable].replace('(NEE)','') + ' ('+unit_dict[variable]+')',
x_axis_type='datetime',
title = measurement_dict_eng[variable] +' - Hyltemossa, Sverige (' + str(df.index[1].year)+')',
tools='pan,box_zoom,wheel_zoom,reset,save')
#Extract time and tracer values for every data level:
x1 = df.index.values
y1 = df[variable].values
#Create a circle and line glyph for the values of every emission category:
r0 = p.circle(x1, y1, radius=.12, color=color)
r1 = p.line(x1, y1, line_width=1, color=color)
#Add tooltip on hover:
p.add_tools(HoverTool(tooltips=[
('Time (UTC)','@x{%Y-%m-%d %H:%M:%S}'),
(measurement_dict_eng[variable] + ' ('+unit_dict[variable]+')','@y{0.f}'),
],
formatters={
'@x' : 'datetime', # use 'datetime' formatter for 'date' field
},
# display a tooltip whenever the cursor is vertically in line with a glyph
mode='vline'
))
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '13pt'
p.title.offset = 15
#Set axis label font style:
p.xaxis.axis_label_text_font_style = 'normal'
p.yaxis.axis_label_text_font_style = 'normal'
p.xaxis.axis_label_standoff = 15 #Sets the distance of the label from the x-axis in screen units
p.yaxis.axis_label_standoff = 15 #Sets the distance of the label from the y-axis in screen units
#Set the copyright label position:
label_opts = dict(x=0, y=10,
x_units='screen', y_units='screen')
#Create a label object and format it:
caption1 = Label(text="© ICOS ERIC", **label_opts)
caption1.text_font_size = '8pt'
#Deactivate hover-tool, which is by default active:
p.toolbar.active_inspect = None
#Add label to plot:
p.add_layout(caption1, 'below')
#Set the output location:
output_notebook()
#Show plot:
show(p)
# -
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Function that create widgets to update plot with icos ecosystem data:
def create_widgets_icos_eco_htm():
#Import modules:
from ipywidgets import interact_manual, ColorPicker, Dropdown
#Create a list with the years for which data exist:
year_ls = [2015, 2016, 2017, 2018]
#Create a list to store the different ecosystem variables:
eco_var_ls = [tuple(reversed(tupl)) for tupl in tuple(measurement_dict_eng.items())]
#Create dropdown-widgets:
years = Dropdown(options = year_ls)
eco_vars = Dropdown(options = eco_var_ls)
#Function that updates the plot based on the user's selection:
def update_eco_line_plot(Year, Variable, color):
#Create a dictionary for every type of dataframe with ICOS ecosystem data:
icos_eco_df_dict ={'2015': HTM_eko_2015_indexed,
'2016': HTM_eko_2016_indexed,
'2017': HTM_eko_2017_indexed,
'2018': HTM_eko_2018_indexed}
icos_eco_precip_df_dict ={'2015': HTM_eko_precip_2015_indexed,
'2016': HTM_eko_precip_2016_indexed,
'2017': HTM_eko_precip_2017_indexed,
'2018': HTM_eko_precip_2018_indexed}
#Check selected variable and get the name of its corresponding pandas dataframe:
if(Variable in HTM_eko_2015_indexed):
dataFrame = icos_eco_df_dict[str(Year)]
else:
dataFrame = icos_eco_precip_df_dict[str(Year)]
#Call function to show plot:
plotIcosEcoIndexedDF(dataFrame, Variable, color)
#Create function that contains a box of widgets:
interact = interact_manual(update_eco_line_plot,
Year = years,
Variable = eco_vars,
color = ColorPicker(concise=False,
description='Pick a color',
value='#3973ac',
disabled=False))
#Set the font of the widgets included in interact_manual:
interact.widget.children[0].layout.width = '460px'
interact.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact.widget.children[1].layout.width = '460px'
interact.widget.children[2].layout.width = '460px'
interact.widget.children[3].description = 'Update Plot'
interact.widget.children[3].button_style = 'danger'
interact.widget.children[3].style.button_color = '#3973ac'
interact.widget.children[3].layout.margin = '20px 10px 40px 200px' # top/right/bottom/left
# -
# <a id='bokeh_plot_2_cols_from_df_plot'></a>
# <br>
#
# #### Bokeh Interactive Plot - Displaying Values from 2 Pandas DataFrame columns
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to display widgets:
create_widgets_icos_eco_htm()
# -
# <a id='bokeh_plot_stat_barplot'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
#
# #### 10.2. Plot Statistics with Bokeh Visualization Library
# In this part, you will learn how to create Barplots with Bokeh Visualization Library over statistics that have been calculated over all values of a year or over a selection of values of a certain year. The statistics are calculated using the code that was presented in the corresponding previous part.
#
# This part is divided into two subparts:
# - [Plot Statistics with Bokeh Visualization Library (Annual Statistics-Complete Year)](#bokeh_plot_stat_barplot_annual_total)
# - [Plot Statistics with Bokeh Visualization Library (Annual Statistics-Part of Year)](#bokeh_plot_stat_barplot_annual_filtered)
#
# <br>
#
# Every subpart includes three code-cells:
# - The first code-cell includes code for the function that handles the format of the barplot.
# - The second code-cell includes a function that creates and formats the widgets (e.g. dropdown lists, color-pickers, execution button, etc.). The second code-cell also includes a nested function that calculates the statistics and updates the content of the barplot based on the users selection.
# - The third code-cell includes a call to the function that creates and displays the widgets (i.e. function included in the 2nd code-cell).
# <a id='bokeh_plot_stat_barplot_annual_total'></a>
# <br>
#
# ##### 10.2.1. Plot Statistics with Bokeh Visualization Library (Annual Statistics-Complete Year)
# This subpart is dedicated to calculating statistics over all values of a year and displaying the results in the form of a barplot. Every bar represents the statistic value for one year.
#
# The user is able to interact and the change the content of a plot using a set of widgets. The available widgets are two dropdown lists that control the type of statistic and the variable over which the statistic should be calculated over, two color-pickers that allow the user to set the color for the bars and the text on the bars in the barplot and, finally, an execution button.
#
# The user is free to change the values in the widgets, but the content of the plot will change to show the results of the new selection of widget-values only once the execution-button is clicked.
#
# The barplot includes an interactive toolbox menu (see Figure below). From here it is possible to pan, zoom-in and out, reset the plot to its initial state and save a copy of the plot to your computer.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/bokeh_plots/barplot_with_widgets.png" width="700" align="center">
# <br>
# <br>
#
#
# <br>
# <div style="text-align: right">
# <a href="#bokeh_barplot_basic_stat_year">[Go to Plot]</a>
# </div>
#
# <br>
#
# <span style="color:blue">**Plotting Function**</span>
# + deletable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def plotIcosEcoBarPlotAnnualStat(Variable, Stat, stat_list, year_ls, bar_color, txt_color):
#Import modules:
from bokeh.models import ColumnDataSource, LabelSet, Label, FixedTicker
#Define y-position of statistics-label (in the middle of the bar-glyph):
y_label_pos_ls = [(stat/2)-0.5 if((stat<=-1) or (stat>=1)) else stat/2 for stat in stat_list]
#Create ColumnDataSource Object:
source = ColumnDataSource(data=dict(years=year_ls,
stats=stat_list,
y_label_pos = y_label_pos_ls))
#Create figure object:
p = figure(plot_width=600, plot_height=450,
title = 'Hyltemossa: '+Stat+' '+measurement_dict_eng[Variable]+' per Year',
x_axis_label = 'Year', y_axis_label = Stat+' '+measurement_dict_eng[Variable]+ ' ('+unit_dict[Variable]+')')
#Add bar glyphs:
p.vbar(x='years', width=0.5, bottom=0,
top='stats', source=source, color=bar_color) #orange
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '12pt'
p.title.offset = 15
#Set axis label font style:
p.xaxis.axis_label_text_font_style = 'normal'
p.yaxis.axis_label_text_font_style = 'normal'
p.xaxis.axis_label_standoff = 15 #Sets the distance of the label from the x-axis in screen units
p.yaxis.axis_label_standoff = 15 #Sets the distance of the label from the y-axis in screen units
#Add labels to the bar glyphs:
labels = LabelSet(x='years', y='y_label_pos', text='stats', level='glyph', text_color=txt_color,
x_offset=0, y_offset=0, source=source, render_mode='css', text_align='center')
#Set the copyright-label position:
label_opts = dict(x=0, y=5,
x_units='screen', y_units='screen')
#Create a label object and format it:
caption1 = Label(text="© ICOS ERIC", **label_opts)
caption1.text_font_size = '8pt'
#Add the bar-glyph lables to the plot:
p.add_layout(labels)
#Add label to plot:
p.add_layout(caption1, 'below')
#Set x-axis tickers:
p.xaxis.ticker = FixedTicker(ticks=year_ls)
#Define output location:
output_notebook()
#Show plot:
show(p)
# -
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def create_widgets_icos_eco_htm_stat_annual():
#Import modules:
from ipywidgets import interact_manual, ColorPicker, Dropdown
#Create a list with the years for which data exist:
year_ls = [2015, 2016, 2017, 2018]
#Create a list containing the names of the statistical operations:
stat_ls = ['Min', 'Max', 'Mean', 'St dev']
#Create a list to store the different ecosystem variables:
eco_var_ls = [tuple(reversed(tupl)) for tupl in tuple(measurement_dict_eng.items())]
#Create dropdown-widgets:
eco_vars = Dropdown(options = eco_var_ls)
stats = Dropdown(options = stat_ls)
#Function that updates the plot based on the user's selection:
def update_eco_bar_plot(Stat, Variable, bar_color, txt_color):
#Create a dictionary for every type of dataframe with ICOS ecosystem data:
icos_eco_df_dict ={'2015': HTM_eko_2015_indexed,
'2016': HTM_eko_2016_indexed,
'2017': HTM_eko_2017_indexed,
'2018': HTM_eko_2018_indexed}
icos_eco_precip_df_dict ={'2015': HTM_eko_precip_2015_indexed,
'2016': HTM_eko_precip_2016_indexed,
'2017': HTM_eko_precip_2017_indexed,
'2018': HTM_eko_precip_2018_indexed}
#Declare and initialize list to store the stats:
stat_list = []
#Check if the selected variable is included in the Temp, GPP, NEE, RECO & SW-IR pandas dataframe:
if(Variable in HTM_eko_2015_indexed):
#Check the type of the selected statistic and store the stat-value
#of every year in a list:
if(Stat=='Min'):
stat_list.append(round(HTM_eko_2015_indexed[Variable].min(),1))
stat_list.append(round(HTM_eko_2016_indexed[Variable].min(),1))
stat_list.append(round(HTM_eko_2017_indexed[Variable].min(),1))
stat_list.append(round(HTM_eko_2018_indexed[Variable].min(),1))
elif(Stat=='Max'):
stat_list.append(round(HTM_eko_2015_indexed[Variable].max(),1))
stat_list.append(round(HTM_eko_2016_indexed[Variable].max(),1))
stat_list.append(round(HTM_eko_2017_indexed[Variable].max(),1))
stat_list.append(round(HTM_eko_2018_indexed[Variable].max(),1))
elif(Stat=='Mean'):
stat_list.append(round(HTM_eko_2015_indexed[Variable].mean(),1))
stat_list.append(round(HTM_eko_2016_indexed[Variable].mean(),1))
stat_list.append(round(HTM_eko_2017_indexed[Variable].mean(),1))
stat_list.append(round(HTM_eko_2018_indexed[Variable].mean(),1))
elif(Stat=='St dev'):
stat_list.append(round(HTM_eko_2015_indexed[Variable].std(),1))
stat_list.append(round(HTM_eko_2016_indexed[Variable].std(),1))
stat_list.append(round(HTM_eko_2017_indexed[Variable].std(),1))
stat_list.append(round(HTM_eko_2018_indexed[Variable].std(),1))
else:
print('Statistic does not exist!')
#If the selected variable is in the precipitation and soil-water-content dataframe:
else:
#Check the type of the selected statistic and store the stat-value
#of every year in a list:
if(Stat=='Min'):
stat_list.append(round(HTM_eko_precip_2015_indexed[Variable].min(),1))
stat_list.append(round(HTM_eko_precip_2016_indexed[Variable].min(),1))
stat_list.append(round(HTM_eko_precip_2017_indexed[Variable].min(),1))
stat_list.append(round(HTM_eko_precip_2018_indexed[Variable].min(),1))
elif(Stat=='Max'):
stat_list.append(round(HTM_eko_precip_2015_indexed[Variable].max(),1))
stat_list.append(round(HTM_eko_precip_2016_indexed[Variable].max(),1))
stat_list.append(round(HTM_eko_precip_2017_indexed[Variable].max(),1))
stat_list.append(round(HTM_eko_precip_2018_indexed[Variable].max(),1))
elif((Stat=='Mean') and (Variable=='P_1_1_1')):
stat_list.append(round(HTM_eko_precip_2015_indexed[Variable].mean(),2))
stat_list.append(round(HTM_eko_precip_2016_indexed[Variable].mean(),2))
stat_list.append(round(HTM_eko_precip_2017_indexed[Variable].mean(),2))
stat_list.append(round(HTM_eko_precip_2018_indexed[Variable].mean(),2))
elif((Stat=='Mean') and (Variable!='P_1_1_1')):
stat_list.append(round(HTM_eko_precip_2015_indexed[Variable].mean(),1))
stat_list.append(round(HTM_eko_precip_2016_indexed[Variable].mean(),1))
stat_list.append(round(HTM_eko_precip_2017_indexed[Variable].mean(),1))
stat_list.append(round(HTM_eko_precip_2018_indexed[Variable].mean(),1))
elif(Stat=='St dev'):
stat_list.append(round(HTM_eko_precip_2015_indexed[Variable].std(),1))
stat_list.append(round(HTM_eko_precip_2016_indexed[Variable].std(),1))
stat_list.append(round(HTM_eko_precip_2017_indexed[Variable].std(),1))
stat_list.append(round(HTM_eko_precip_2018_indexed[Variable].std(),1))
else:
print('Statistic does not exist!')
#Call function to show plot:
plotIcosEcoBarPlotAnnualStat(Variable, Stat, stat_list, year_ls, bar_color, txt_color)
#Create function that contains a box of widgets:
interact = interact_manual(update_eco_bar_plot,
Variable = eco_vars,
Stat = stats,
bar_color = ColorPicker(concise=False,
description='Bar color',
value='#3973ac',
disabled=False),
txt_color = ColorPicker(concise=False,
description='Text color',
value='orange',
disabled=False))
#Set the font of the widgets included in interact_manual:
interact.widget.children[0].layout.width = '460px'
interact.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact.widget.children[1].layout.width = '460px'
interact.widget.children[2].layout.width = '460px'
interact.widget.children[3].layout.width = '460px'
interact.widget.children[4].description = 'Update Plot'
interact.widget.children[4].button_style = 'danger'
interact.widget.children[4].style.button_color = '#3973ac'
interact.widget.children[4].layout.margin = '20px 10px 40px 200px' # top/right/bottom/left
# -
# <a id='bokeh_barplot_basic_stat_year'></a>
# <br>
# #### Display Widgets for Barplots - Annual Statistics (All values)
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to display widgets:
create_widgets_icos_eco_htm_stat_annual()
# -
# <a id='bokeh_plot_stat_barplot_annual_filtered'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
# <br>
#
# ##### 10.2.2. Plot Statistics with Bokeh Visualization Library (Annual Statistics-Part of Year)
#
# This subpart is dedicated to calculating statistics over a selection of values of a year and displaying the results in the form of a barplot. Every bar represents the statistic value for one year. Here the results only present statistics calculated over values belonging to the time period June-August.
#
# The user is able to interact and the change the content of a plot using a set of widgets. The available widgets are two dropdown lists that control the type of statistic and the variable over which the statistic should be calculated over, two color-pickers that allow the user to set the color for the bars and the text on the bars in the barplot and, finally, an execution button.
#
# The user is free to change the values in the widgets, but the content of the plot will change to show the results of the new selection of widget-values only once the execution-button is clicked.
#
# The barplot includes an interactive toolbox menu (see Figure below). From here it is possible to pan, zoom-in and out, reset the plot to its initial state and save a copy of the plot to your computer.
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/bokeh_plots/barplot_summer_stat_widgets.png" width="700" align="center">
# <br>
# <br>
#
# <br>
# <div style="text-align: right">
# <a href="#bokeh_barplot_basic_stat_summmer">[Go to Plot]</a>
# </div>
#
# <br>
#
# <span style="color:blue">**Plotting Function**</span>
# + deletable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def plotIcosEcoBarPlotAnnualStatJunAug(Variable, Stat, stat_list, year_ls, bar_color, txt_color):
#Import modules:
from bokeh.models import ColumnDataSource, LabelSet, Label, FixedTicker
#Define y-position of statistics-label (in the middle of the bar-glyph):
y_label_pos_ls = [(stat/2)-0.5 if((stat<=-1) or (stat>=1)) else stat/2 for stat in stat_list]
#Create ColumnDataSource Object:
source = ColumnDataSource(data=dict(years=year_ls,
stats=stat_list,
y_label_pos = y_label_pos_ls))
#Create figure object:
p = figure(plot_width=600, plot_height=450,
title = 'Hyltemossa: '+Stat+' '+measurement_dict_eng[Variable]+' (Jun-Aug) per Year',
x_axis_label = 'Year', y_axis_label = Stat+' '+measurement_dict_eng[Variable]+ ' ('+unit_dict[Variable]+')')
#Add bar glyphs:
p.vbar(x='years', width=0.5, bottom=0,
top='stats', source=source, color=bar_color) #orange
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '12pt'
p.title.offset = 15
#Set axis label font style:
p.xaxis.axis_label_text_font_style = 'normal'
p.yaxis.axis_label_text_font_style = 'normal'
p.xaxis.axis_label_standoff = 15 #Sets the distance of the label from the x-axis in screen units
p.yaxis.axis_label_standoff = 15 #Sets the distance of the label from the y-axis in screen units
#Add labels to the bar glyphs:
labels = LabelSet(x='years', y='y_label_pos', text='stats', level='glyph', text_color=txt_color,
x_offset=0, y_offset=0, source=source, render_mode='css', text_align='center')
#Set the copyright-label position:
label_opts = dict(x=0, y=5,
x_units='screen', y_units='screen')
#Create a label object and format it:
caption1 = Label(text="© ICOS ERIC", **label_opts)
caption1.text_font_size = '8pt'
#Add the bar-glyph lables to the plot:
p.add_layout(labels)
#Add label to plot:
p.add_layout(caption1, 'below')
#Set x-axis tickers:
p.xaxis.ticker = FixedTicker(ticks=year_ls)
#Define output location:
output_notebook()
#Show plot:
show(p)
# -
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def create_widgets_icos_eco_htm_stat_annual_jun_aug():
#Import modules:
from ipywidgets import interact_manual, ColorPicker, Dropdown
#Create a list with the years for which data exist:
year_ls = [2015, 2016, 2017, 2018]
#Create a list containing the names of the statistical operations:
stat_ls = ['Min', 'Max', 'Mean', 'St dev']
#Create a list to store the different ecosystem variables:
eco_var_ls = [tuple(reversed(tupl)) for tupl in tuple(measurement_dict_eng.items())]
#Create dropdown-widgets:
eco_vars = Dropdown(options = eco_var_ls)
stats = Dropdown(options = stat_ls)
#Function that updates the plot based on the user's selection:
def update_eco_bar_plot(Stat, Variable, bar_color, txt_color):
#Create a dictionary for every type of dataframe with ICOS ecosystem data:
icos_eco_df_dict ={'2015': HTM_summermonths_2015,
'2016': HTM_summermonths_2016,
'2017': HTM_summermonths_2017,
'2018': HTM_summermonths_2018}
icos_eco_precip_df_dict ={'2015': HTM_P_summermonths_2015,
'2016': HTM_P_summermonths_2016,
'2017': HTM_P_summermonths_2017,
'2018': HTM_P_summermonths_2018}
#Declare and initialize list to store the stats:
stat_list = []
#Check if the selected variable is included in the Temp, GPP, NEE, RECO & SW-IR pandas dataframe:
if(Variable in HTM_summermonths_2015):
#Check the type of the selected statistic and store the stat-value
#of every year in a list:
if(Stat=='Min'):
stat_list.append(round(HTM_summermonths_2015[Variable].min(),1))
stat_list.append(round(HTM_summermonths_2016[Variable].min(),1))
stat_list.append(round(HTM_summermonths_2017[Variable].min(),1))
stat_list.append(round(HTM_summermonths_2018[Variable].min(),1))
elif(Stat=='Max'):
stat_list.append(round(HTM_summermonths_2015[Variable].max(),1))
stat_list.append(round(HTM_summermonths_2016[Variable].max(),1))
stat_list.append(round(HTM_summermonths_2017[Variable].max(),1))
stat_list.append(round(HTM_summermonths_2018[Variable].max(),1))
elif(Stat=='Mean'):
stat_list.append(round(HTM_summermonths_2015[Variable].mean(),1))
stat_list.append(round(HTM_summermonths_2016[Variable].mean(),1))
stat_list.append(round(HTM_summermonths_2017[Variable].mean(),1))
stat_list.append(round(HTM_summermonths_2018[Variable].mean(),1))
elif(Stat=='St dev'):
stat_list.append(round(HTM_summermonths_2015[Variable].std(),1))
stat_list.append(round(HTM_summermonths_2016[Variable].std(),1))
stat_list.append(round(HTM_summermonths_2017[Variable].std(),1))
stat_list.append(round(HTM_summermonths_2018[Variable].std(),1))
else:
print('Statistic does not exist!')
#If the selected variable is in the precipitation and soil-water-content dataframe:
else:
#Check the type of the selected statistic and store the stat-value
#of every year in a list:
if(Stat=='Min'):
stat_list.append(round(HTM_P_summermonths_2015[Variable].min(),1))
stat_list.append(round(HTM_P_summermonths_2016[Variable].min(),1))
stat_list.append(round(HTM_P_summermonths_2017[Variable].min(),1))
stat_list.append(round(HTM_P_summermonths_2018[Variable].min(),1))
elif(Stat=='Max'):
stat_list.append(round(HTM_P_summermonths_2015[Variable].max(),1))
stat_list.append(round(HTM_P_summermonths_2016[Variable].max(),1))
stat_list.append(round(HTM_P_summermonths_2017[Variable].max(),1))
stat_list.append(round(HTM_P_summermonths_2018[Variable].max(),1))
elif((Stat=='Mean') and (Variable=='P_1_1_1')):
stat_list.append(round(HTM_P_summermonths_2015[Variable].mean(),2))
stat_list.append(round(HTM_P_summermonths_2016[Variable].mean(),2))
stat_list.append(round(HTM_P_summermonths_2017[Variable].mean(),2))
stat_list.append(round(HTM_P_summermonths_2018[Variable].mean(),2))
elif((Stat=='Mean') and (Variable!='P_1_1_1')):
stat_list.append(round(HTM_P_summermonths_2015[Variable].mean(),1))
stat_list.append(round(HTM_P_summermonths_2016[Variable].mean(),1))
stat_list.append(round(HTM_P_summermonths_2017[Variable].mean(),1))
stat_list.append(round(HTM_P_summermonths_2018[Variable].mean(),1))
elif(Stat=='St dev'):
stat_list.append(round(HTM_P_summermonths_2015[Variable].std(),1))
stat_list.append(round(HTM_P_summermonths_2016[Variable].std(),1))
stat_list.append(round(HTM_P_summermonths_2017[Variable].std(),1))
stat_list.append(round(HTM_P_summermonths_2018[Variable].std(),1))
else:
print('Statistic does not exist!')
#Call function to show plot:
plotIcosEcoBarPlotAnnualStatJunAug(Variable, Stat, stat_list, year_ls, bar_color, txt_color)
#Create function that contains a box of widgets:
interact = interact_manual(update_eco_bar_plot,
Variable = eco_vars,
Stat = stats,
bar_color = ColorPicker(concise=False,
description='Bar color',
value='#3973ac',
disabled=False),
txt_color = ColorPicker(concise=False,
description='Text color',
value='orange',
disabled=False))
#Set the font of the widgets included in interact_manual:
interact.widget.children[0].layout.width = '460px'
interact.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact.widget.children[1].layout.width = '460px'
interact.widget.children[2].layout.width = '460px'
interact.widget.children[3].layout.width = '460px'
interact.widget.children[4].description = 'Update Plot'
interact.widget.children[4].button_style = 'danger'
interact.widget.children[4].style.button_color = '#3973ac'
interact.widget.children[4].layout.margin = '20px 10px 40px 200px' # top/right/bottom/left
# -
# <a id='bokeh_barplot_basic_stat_summmer'></a>
# <br>
# #### Display Widgets for Barplots - Annual Statistics (Selection of values)
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to show widgets:
create_widgets_icos_eco_htm_stat_annual_jun_aug()
# + [markdown] slideshow={"slide_type": "fragment"}
# <a id='bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
#
# #### 10.3. Create Plots with Cumulative Sums of Daily Totals and Daily Means per Year
# In this part we will create plots of cumulative sums of daily totals or daily means per year for a selection of ecosystem variables. These plots provide a general overview of how the values of a selected variable might vary between different years. The first step is to produce daily totals or daily means for every variable. It might also be necessary to perform some unit conversions. A step by step explanation of the processes follows.
#
#
# This part is subdivided into the following parts:
#
# - [Compute Daily Totals and Daily Means per Year](#compute_daily_totals_and_daily_means_per_year)
# - [Conversion of Units for the Computation of Daily Totals](#conversion_of_units_for_daily_totals)
# - [Compute Cumulative Sums of Daily Totals per Year](#compute_iterative_sums_of_daily_totals_per_year)
# - [Convert the Units of the Cumulative Sum Values](#convert_units_iterative_sums)
# - [Create Interactive Plots to Display the Cumulative Sums for every Variable](#bokeh_plot_stat_summed_values_per_year)
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_stat_summed_values_per_year_plot">[Go to plot]</a>
#  
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
# -
# <br>
# <br>
# <a id='compute_daily_totals_and_daily_means_per_year'></a>
#
# ##### 10.3.1. Compute Daily Totals and Daily Means per Year
# We will compute the daily sum of the following variables:
# - Air Temperature
# - Precipitation
# - Respiration
# - Gross Primary Production (GPP)
# - Carbon Flux - Net Ecosystem Exchange (NEE)
# - Incoming Shortwave Infrared (SW-IR) Solar Radiation
#
# <br>
#
# To compute the daily sum of each variable, we will use some of Python's built-in methods for Pandas DataFrames. The image bellow shows the code used to compute the daily sums of Air Temperature values for 2015. Observe that the Python built-in methods, used in this case, will only work if you have set a column containing Python DateTime objects as the index of your Pandas DataFrame.
#
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/pseudocode/daily_sums.png" width="500" align="center">
#
# <br>
# <br>
#
# <br>
#
# We will compute the daily average of the following variables:
# - Air Temperature
# - Soil Water Content
#
# <br>
#
# Similarly to the previous example, to compute the daily mean of each variable we will again use some of Python's built-in methods for a Pandas DataFrame. The image bellow shows the code used to compute the daily averages of Air Temperature values for 2015. Again, this code will only work if you have set a column containing Python DateTime objects as the index of your Pandas DataFrame.
#
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/pseudocode/daily_means.png" width="500" align="center">
#
# <br>
# <br>
# <br>
# <br>
#
# <div style="text-align: right">
# <a href="#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro">[Back to Create Plots of Cumulative Sums - Intro]</a>
# </div>
#
# <br>
# <br>
# <br>
# <br>
# <a id='conversion_of_units_for_daily_totals'></a>
#
# ##### 10.3.2. Conversion of Units for the Computation of Daily Totals
#
#
# It is necessary to convert the units of some of the ecosystem variables, before computing their daily sums or daily means.
# For example, there is a new Respiration value every 30 min. However the Respiration unit is **μmol m$^{-2}$ s$^{-1}$**. In order to compute the daily sum of Respiration, it is necessary to first compute the Respiration for every 30 min and then sum the computed values. To get the Respiration in **μmol m$^{-2}$ day$^{-1}$**, multiply every value by 60 (to get the respiration per minute - there are 60 sec in one minute) and then by 30 (to get the respiration per 30 min) and, finally, sum up all the values.
# <br>
# The same conversion should be applied to the Gross Primary Production (GPP) and Net Ecosystem Exchange (NEE) values, as they have the same unit.
# <br>
# <br>
# Incoming Shortwave Infrared (SW-IR) Solar Radiation is given as Watts per square meter (W/m$^2$). In order to get the total sum of Incoming SW-IR Solar Radiation per day, it is necessary to convert Watts to Joules. Because **1 Watt = 1 Joule/sec**, we will compute Incoming SW-IR Radiation as Joules per square meter per 30 min by multiplying the current values first by 60 (to get the SW-IR Radiation per minute) and then by 30 (to get SW-IR Radiation per 30 minutes).
#
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/pseudocode/conversion_of_units_py.png" width="800" align="center">
#
# <br>
# <br>
# <br>
# <br>
#
# <div style="text-align: right">
# <a href="#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro">[Back to Create Plots of Cumulative Sums - Intro]</a>
# </div>
#
# <br>
# <br>
# <br>
#
# <span style="color:blue">**Air Temperature - Daily Totals**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Get daily sum of Air Temperature for every dataframe (i.e. year):
HTM_eko_TA_2015_daily_sum = HTM_eko_2015_indexed.TA_1_1_1.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_TA_2016_daily_sum = HTM_eko_2016_indexed.TA_1_1_1.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_TA_2017_daily_sum = HTM_eko_2017_indexed.TA_1_1_1.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_TA_2018_daily_sum = HTM_eko_2018_indexed.TA_1_1_1.resample('D').sum().dropna() #disregard NaN-values
#View the 5 first rows of the result:
HTM_eko_TA_2018_daily_sum.head(5)
# -
# <br>
#
# <span style="color:blue">**Air Temperature - Daily Average**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Get daily average of Air Temperature for every dataframe (i.e. year):
HTM_eko_TA_2015_daily_mean = HTM_eko_2015_indexed.TA_1_1_1.resample('D').mean().dropna() #disregard NaN-values
HTM_eko_TA_2016_daily_mean = HTM_eko_2016_indexed.TA_1_1_1.resample('D').mean().dropna() #disregard NaN-values
HTM_eko_TA_2017_daily_mean = HTM_eko_2017_indexed.TA_1_1_1.resample('D').mean().dropna() #disregard NaN-values
HTM_eko_TA_2018_daily_mean = HTM_eko_2018_indexed.TA_1_1_1.resample('D').mean().dropna() #disregard NaN-values
#View the 5 first rows of the result:
HTM_eko_TA_2015_daily_mean.head(5)
# -
# <br>
#
# <span style="color:blue">**Precipitation - Daily Totals**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Get daily sum of Precipitation for every dataframe (i.e. year):
HTM_eko_P_2015_daily_sum = HTM_eko_precip_2015_indexed.P_1_1_1.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_P_2016_daily_sum = HTM_eko_precip_2016_indexed.P_1_1_1.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_P_2017_daily_sum = HTM_eko_precip_2017_indexed.P_1_1_1.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_P_2018_daily_sum = HTM_eko_precip_2018_indexed.P_1_1_1.resample('D').sum().dropna() #disregard NaN-values
# -
# <br>
#
# <span style="color:blue">**Respiration - Daily Totals**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#RESPIRATION HALF-HOURLY:
#Add column with RECO computed as micromoles per square meter per 30 min (conv. sec to 30min):
HTM_eko_2015_indexed['RECO_PI_1_1_1_30min'] = [HTM_eko_2015_indexed.RECO_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2015_indexed))]
HTM_eko_2016_indexed['RECO_PI_1_1_1_30min'] = [HTM_eko_2016_indexed.RECO_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2016_indexed))]
HTM_eko_2017_indexed['RECO_PI_1_1_1_30min'] = [HTM_eko_2017_indexed.RECO_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2017_indexed))]
HTM_eko_2018_indexed['RECO_PI_1_1_1_30min'] = [HTM_eko_2018_indexed.RECO_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2018_indexed))]
#Get daily sum of Respiration for every dataframe (i.e. year):
HTM_eko_RECO_2015_daily_sum = HTM_eko_2015_indexed.RECO_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_RECO_2016_daily_sum = HTM_eko_2016_indexed.RECO_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_RECO_2017_daily_sum = HTM_eko_2017_indexed.RECO_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_RECO_2018_daily_sum = HTM_eko_2018_indexed.RECO_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
#View the 5 first rows of the result:
#HTM_eko_RECO_2015_daily_sum.head(5)
# -
# <br>
#
# <span style="color:blue">**GPP - Daily Totals**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#GPP HALF-HOURLY:
#Add column with GPP computed as micromoles per square meter per 30 min (conv. sec to 30min):
HTM_eko_2015_indexed['GPP_PI_1_1_1_30min'] = [HTM_eko_2015_indexed.GPP_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2015_indexed))]
HTM_eko_2016_indexed['GPP_PI_1_1_1_30min'] = [HTM_eko_2016_indexed.GPP_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2016_indexed))]
HTM_eko_2017_indexed['GPP_PI_1_1_1_30min'] = [HTM_eko_2017_indexed.GPP_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2017_indexed))]
HTM_eko_2018_indexed['GPP_PI_1_1_1_30min'] = [HTM_eko_2018_indexed.GPP_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2018_indexed))]
#Get daily sum of GPP for every dataframe (i.e. year):
HTM_eko_GPP_2015_daily_sum = HTM_eko_2015_indexed.GPP_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_GPP_2016_daily_sum = HTM_eko_2016_indexed.GPP_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_GPP_2017_daily_sum = HTM_eko_2017_indexed.GPP_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_GPP_2018_daily_sum = HTM_eko_2018_indexed.GPP_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
# -
# <br>
#
# <span style="color:blue">**NEE - Daily Totals**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#CARBON FLUX (NEE) HALF-HOURLY:
#Add column with Carbon Flux computed as micromoles per square meter per 30 min (conv. sec to 30min):
HTM_eko_2015_indexed['FC_PI_1_1_1_30min'] = [HTM_eko_2015_indexed.FC_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2015_indexed))]
HTM_eko_2016_indexed['FC_PI_1_1_1_30min'] = [HTM_eko_2016_indexed.FC_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2016_indexed))]
HTM_eko_2017_indexed['FC_PI_1_1_1_30min'] = [HTM_eko_2017_indexed.FC_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2017_indexed))]
HTM_eko_2018_indexed['FC_PI_1_1_1_30min'] = [HTM_eko_2018_indexed.FC_PI_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2018_indexed))]
#Get daily sum of Carbon Flux for every dataframe (i.e. year):
HTM_eko_NEE_2015_daily_sum = HTM_eko_2015_indexed.FC_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_NEE_2016_daily_sum = HTM_eko_2016_indexed.FC_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_NEE_2017_daily_sum = HTM_eko_2017_indexed.FC_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_NEE_2018_daily_sum = HTM_eko_2018_indexed.FC_PI_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
# -
# <br>
#
# <span style="color:blue">**Shortwave Infrared Incoming Solar Radiation - Daily Totals**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Add column with Light computed as Joules per square meter for 30 min (i.e. 1800 sec):
HTM_eko_2015_indexed['SW_IN_1_1_1_30min'] = [HTM_eko_2015_indexed.SW_IN_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2015_indexed))]
HTM_eko_2016_indexed['SW_IN_1_1_1_30min'] = [HTM_eko_2016_indexed.SW_IN_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2016_indexed))]
HTM_eko_2017_indexed['SW_IN_1_1_1_30min'] = [HTM_eko_2017_indexed.SW_IN_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2017_indexed))]
HTM_eko_2018_indexed['SW_IN_1_1_1_30min'] = [HTM_eko_2018_indexed.SW_IN_1_1_1[ind]*60*30 for ind in range(len(HTM_eko_2018_indexed))]
#Get daily sum of "Light" for every dataframe (i.e. year):
HTM_eko_LIGHT_2015_daily_sum = HTM_eko_2015_indexed.SW_IN_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_LIGHT_2016_daily_sum = HTM_eko_2016_indexed.SW_IN_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_LIGHT_2017_daily_sum = HTM_eko_2017_indexed.SW_IN_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
HTM_eko_LIGHT_2018_daily_sum = HTM_eko_2018_indexed.SW_IN_1_1_1_30min.resample('D').sum().dropna() #disregard NaN-values
# -
# <br>
#
# <span style="color:blue">**Soil Water Content - Daily Average**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Get daily mean of "Soil Water Content" for every dataframe (i.e. year):
HTM_eko_SWC_2015_daily_mean = HTM_eko_precip_2015_indexed.SWC_4_4_1.resample('D').mean().dropna() #disregard NaN-values
HTM_eko_SWC_2016_daily_mean = HTM_eko_precip_2016_indexed.SWC_4_4_1.resample('D').mean().dropna() #disregard NaN-values
HTM_eko_SWC_2017_daily_mean = HTM_eko_precip_2017_indexed.SWC_4_4_1.resample('D').mean().dropna() #disregard NaN-values
HTM_eko_SWC_2018_daily_mean = HTM_eko_precip_2018_indexed.SWC_4_4_1.resample('D').mean().dropna() #disregard NaN-values
# -
# <br>
# <br>
# <a id='compute_iterative_sums_of_daily_totals_per_year'></a>
#
# ##### 10.3.3. Compute Cumulative Sums of Daily Totals per Year
# Now we will compute the cumulative sums of the daily totals of the following variables:
#
# - Air Temperature
# - Precipitation
# - Respiration
# - Gross Primary Production (GPP)
# - Carbon Flux - Net Ecosystem Exchange (NEE)
#
# <br>
# <br>
#
# A cumulative sum of daily sums is computed as following:
#
# <br>
# <br>
# <img src="../ancillarydata/images/htm_drought/pseudocode/iterative_sums.png" width="810" align="center">
# <br>
# <br>
#
# The next code cell includes a function in Python code that computes the cumulative sum of the elements of a list or pandas series. It returns a list with the result. Python has built-in methods to perform the same computation. In this case, we present both options for explanatory purposes. Generally, Python's built-in methods are faster and should therefore be preferred over any piece of self-produced code.
#
# <br>
# <br>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro">[Back to Create Plots of Cumulative Sums - Intro]</a>
# </div>
#
# <br>
# <br>
# <br>
#
# <span style="color:blue">**Cumulative Sum Function (without the use of built-in methods)**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#This function does the same as the following python command:
#list(itertools.accumulate(HTM_eko_2015_daily_mean))
def cumulative_sum(ls):
"""
Function that produces a list of the iterative sums of the elements of a list or series.
"""
#Create and initialize help variables:
sum_temp=0 #variable to store intermediate sums
sum_ls = [] #list to store iterative sums
#Loop through every element in list:
for i in range(len(ls)):
#Add current daily-average air temperature to sum:
sum_temp = sum_temp + ls[i]
#Add intermediate sum to list:
sum_ls.append(sum_temp)
#Return list:
return sum_ls
# -
# <br>
#
# <span style="color:blue">**Cumulative Sums of Daily Averaged Air Temperature**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute cumulative sums of averaged daily Air Temperature values per year:
#Create a pandas series with the iterative daily sums of averaged daily Air Temperature values as data
#and their corresponding date as index.
HTM_eko_TA_2015_itersum = pd.Series(data = itertools.accumulate(HTM_eko_TA_2015_daily_mean),
index = HTM_eko_TA_2015_daily_mean.index)
HTM_eko_TA_2016_itersum = pd.Series(data = itertools.accumulate(HTM_eko_TA_2016_daily_mean),
index = HTM_eko_TA_2016_daily_mean.index)
HTM_eko_TA_2017_itersum = pd.Series(data = itertools.accumulate(HTM_eko_TA_2017_daily_mean),
index = HTM_eko_TA_2017_daily_mean.index)
HTM_eko_TA_2018_itersum = pd.Series(data = itertools.accumulate(HTM_eko_TA_2018_daily_mean),
index = HTM_eko_TA_2018_daily_mean.index)
# -
# <br>
#
# <span style="color:blue">**Cumulative Sums of Daily Summed Precipitation**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute cumulative sums of summed daily Precipitation values per year:
#Create a pandas series with the iterative daily sums of summed daily Precipitation values as data
#and their corresponding date as index.
HTM_eko_P_2015_itersum = pd.Series(data = itertools.accumulate(HTM_eko_P_2015_daily_sum),
index = HTM_eko_P_2015_daily_sum.index)
HTM_eko_P_2016_itersum = pd.Series(data = itertools.accumulate(HTM_eko_P_2016_daily_sum),
index = HTM_eko_P_2016_daily_sum.index)
HTM_eko_P_2017_itersum = pd.Series(data = itertools.accumulate(HTM_eko_P_2017_daily_sum),
index = HTM_eko_P_2017_daily_sum.index)
HTM_eko_P_2018_itersum = pd.Series(data = itertools.accumulate(HTM_eko_P_2018_daily_sum),
index = HTM_eko_P_2018_daily_sum.index)
# -
# <br>
#
# <span style="color:blue">**Cumulative Sums of Daily Summed Respiration**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute cumulative sums of summed daily Respiration values per year:
#Create a pandas series with the iterative daily sums of summed daily Respiration values as data
#and their corresponding date as index.
HTM_eko_RECO_2015_itersum = pd.Series(data = itertools.accumulate(HTM_eko_RECO_2015_daily_sum),
index = HTM_eko_RECO_2015_daily_sum.index)
HTM_eko_RECO_2016_itersum = pd.Series(data = itertools.accumulate(HTM_eko_RECO_2016_daily_sum),
index = HTM_eko_RECO_2016_daily_sum.index)
HTM_eko_RECO_2017_itersum = pd.Series(data = itertools.accumulate(HTM_eko_RECO_2017_daily_sum),
index = HTM_eko_RECO_2017_daily_sum.index)
HTM_eko_RECO_2018_itersum = pd.Series(data = itertools.accumulate(HTM_eko_RECO_2018_daily_sum),
index = HTM_eko_RECO_2018_daily_sum.index)
# -
# <br>
#
# <span style="color:blue">**Cumulative Sums of Daily Summed GPP**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute cumulative sums of summed daily GPP values per year:
#Create a pandas series with the iterative daily sums of summed daily GPP values as data
#and their corresponding date as index.
HTM_eko_GPP_2015_itersum = pd.Series(data = itertools.accumulate(HTM_eko_GPP_2015_daily_sum),
index = HTM_eko_GPP_2015_daily_sum.index)
HTM_eko_GPP_2016_itersum = pd.Series(data = itertools.accumulate(HTM_eko_GPP_2016_daily_sum),
index = HTM_eko_GPP_2016_daily_sum.index)
HTM_eko_GPP_2017_itersum = pd.Series(data = itertools.accumulate(HTM_eko_GPP_2017_daily_sum),
index = HTM_eko_GPP_2017_daily_sum.index)
HTM_eko_GPP_2018_itersum = pd.Series(data = itertools.accumulate(HTM_eko_GPP_2018_daily_sum),
index = HTM_eko_GPP_2018_daily_sum.index)
# -
# <br>
#
# <span style="color:blue">**Cumulative Sums of Daily Summed NEE**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute cumulative sums of summed daily Carbon Flux (NEE) values per year:
#Create a pandas series with the iterative daily sums of summed daily Carbon Flux (NEE) values as data
#and their corresponding date as index.
HTM_eko_NEE_2015_itersum = pd.Series(data = itertools.accumulate(HTM_eko_NEE_2015_daily_sum),
index = HTM_eko_NEE_2015_daily_sum.index)
HTM_eko_NEE_2016_itersum = pd.Series(data = itertools.accumulate(HTM_eko_NEE_2016_daily_sum),
index = HTM_eko_NEE_2016_daily_sum.index)
HTM_eko_NEE_2017_itersum = pd.Series(data = itertools.accumulate(HTM_eko_NEE_2017_daily_sum),
index = HTM_eko_NEE_2017_daily_sum.index)
HTM_eko_NEE_2018_itersum = pd.Series(data = itertools.accumulate(HTM_eko_NEE_2018_daily_sum),
index = HTM_eko_NEE_2018_daily_sum.index)
# -
# <br>
#
# <span style="color:blue">**Cumulative Sums of Daily Summed Incoming Shortwave Infrared Solar Radiation**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Compute cumulative sums of summed daily Shortwave Infrared Incoming Solar Radiation values per year:
#Create a pandas series with the iterative daily sums of summed daily Shortwave Infrared Incoming
#Solar Radiation values as data and their corresponding date as index.
HTM_eko_SWIR_2015_itersum = pd.Series(data = itertools.accumulate(HTM_eko_LIGHT_2015_daily_sum),
index = HTM_eko_LIGHT_2015_daily_sum.index)
HTM_eko_SWIR_2016_itersum = pd.Series(data = itertools.accumulate(HTM_eko_LIGHT_2016_daily_sum),
index = HTM_eko_LIGHT_2016_daily_sum.index)
HTM_eko_SWIR_2017_itersum = pd.Series(data = itertools.accumulate(HTM_eko_LIGHT_2017_daily_sum),
index = HTM_eko_LIGHT_2017_daily_sum.index)
HTM_eko_SWIR_2018_itersum = pd.Series(data = itertools.accumulate(HTM_eko_LIGHT_2018_daily_sum),
index = HTM_eko_LIGHT_2018_daily_sum.index)
# -
# <br>
# <br>
# <a id='convert_units_iterative_sums'></a>
#
# ##### 10.3.4. Convert the Units of the Cumulative Sum Values
# Sometimes the output values can be quite large. Before visualizing the output, it is considered good practice to test if it possible to change the units so that the visualized/displayed values are lower. This increases the readability of the plot and thus helps the viewer to better comprehend its content.
#
#
# $$1~mole ~=~ 1,000,000~micromoles$$
#
# <br>
# <br>
# In this case, we will change the unit of the GPP, NEE and Respiration variables from micromoles/m$^2$ day to moles/m$^2$ day. We will also change the unit of the Shortwave Infrared Incoming Solar Radiation from Joules/m$^2$ day to MegaJoules/m$^2$ day.
#
#
# $$1~Megajoule ~=~ 1,000,000~Joules$$
#
# <br>
# <br>
# The functions bellow will take a Pandas Series with the computed iterative sums for a given variable for a particular year as input parameter and return a Pandas Series whose values have been divided by 1,000,000 to produce the output in a different unit (as described above). The function divides each value in the Pandas Series by 1,000,000.
#
# <br>
# <br>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro">[Back to Create Plots of Cumulative Sums - Intro]</a>
# </div>
# <br>
# <br>
#
# <span style="color:blue">**Function - Convert Micromoles to Moles**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Function that transforms the values in the column of a Pandas Series from micromoles to moles:
def micromoles2moles(pandasSeries):
#Import modules:
import pandas as pd
#Convert a pandas series column containing values in micromoles to moles:
ds = pd.Series(data = pandasSeries.values/1000000,
index = pandasSeries.index)
#Return Pandas Series:
return ds
# -
# <br>
#
# <span style="color:blue">**Function - Convert Joules to MegaJoules**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Function that transforms the values in the column of a Pandas Series from Joules to MegaJoules:
def Joules2MegaJoules(pandasSeries):
#Import modules:
import pandas as pd
#Convert a pandas series column containing values in Joules to MegaJoules:
ds = pd.Series(data = pandasSeries.values/1000000,
index = pandasSeries.index)
#Return Pandas Series:
return ds
# -
# <br>
# <br>
# <a id='bokeh_plot_stat_summed_values_per_year'></a>
#
# ##### 10.3.5. Create Interactive Plots to Display the Cumulative Sums of every Variable
# Now that we are done with our computations, we are going to visualize the results in the form of an interactive plot (see figure). For the purpose of this visualization, we are again going to use the [Bokeh visualization library](https://bokeh.pydata.org/en/latest/docs/user_guide/interaction/legends.html).
#
# <br>
# <br>
#
# <img src="../ancillarydata/images/htm_drought/bokeh_plots/bokeh_plot_explained.png" width="700" align="center">
#
# <br>
# <br>
#
# In order to use a tool in the Bokeh Plot ToolBox, you have to activate it. You can activate a tool just by clicking on it. An active tool is always highlighted with a blue line next to its symbol. For instance, in the figure above, the Pan-tool is the only active tool.
#
#
# Use the ```Pan-tool``` to move the content of the plot up or down, right or left.
#
#
# Use the ```Box Zoom-tool``` to zoom-in on a rectangular selected area.
#
#
# Use the ```Wheel Zoom-tool``` to zoom-in over an area in the plot just by scrolling.
#
#
# Press the ```Save``` button to save a copy of the plot to your computer.
#
#
# Press the ```Reset``` button to restore the plot to its initial state.
#
#
# Press the ```Hover``` button and hover with your mouse over the plot to see annotations.
#
#
# Click on an item in the ```interactive legend``` to make the line of that item disappear.
#
# <br>
# <br>
#
# Two functions are used to produce the previously described output. One function creates the plots and another function creates the widgets and handles the updating of the plot based on the user's selection.
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_stat_summed_values_per_year_plot">[Go to plot]</a>
#  
# <a href="#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro">[Back to Create Plots of Cumulative Sums - Intro]</a>
# </div>
# <br>
# <br>
# <br>
#
# <span style="color:blue">**Plotting Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Function that creates plots of iterative sums:
def plotIterSum(itersum_ls, year_ls, aggregation_type, variable, unit, colors):
#Import modules:
from bokeh.models import Legend, HoverTool
#Dictionary for subscript/superscript transformations of numbers:
SUB = str.maketrans("0123456789", "₀₁₂₃₄₅₆₇₈₉")
SUP = str.maketrans("0123456789", "⁰¹²³⁴⁵⁶⁷⁸⁹")
#Create plot
p = figure(plot_width=900, plot_height=450,
title = 'Hyltemossa Station: Cumulative Sums of '+variable+
' Daily '+aggregation_type+' per Year',
x_axis_label = 'Day of the Year (DOY)',
y_axis_label = variable+' ('+unit+')')
#Create an empty list that will store the legend info:
legend_it = []
for num in range(len(itersum_ls)):
#Add Line-glyph:
gL = p.line(list(range(1,len(itersum_ls[num])+1)), itersum_ls[num],
color=colors[num], line_width=1.5, name=str(year_ls[num]))
#Add Circle-glyph:
gC = p.circle(list(range(1,len(itersum_ls[num])+1)), itersum_ls[num],
radius=.12, color=colors[num], name=str(year_ls[num]))
#Add the name and glyph info (i.e. colour and marker type) to the legend:
legend_it.append((gL.name, [gL,gC]))
#Add tooltip on hover:
p.add_tools(HoverTool(tooltips=[
('Year','$name'),
('Day of Year','@x'),
(variable+' ('+unit+')','@y{0.f}'),
],
formatters={
'x' : 'datetime', # use 'datetime' formatter for 'date' field
},
# display a tooltip whenever the cursor is vertically in line with a glyph
mode='vline'
))
#Create legend:
legend = Legend(items=legend_it, location= 'bottom_center')
legend.orientation = 'horizontal'
legend.click_policy='hide'
legend.spacing = 10 #sets the distance between legend entries
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '13pt'
p.title.offset = 15
#Set axis label font style:
p.xaxis.axis_label_text_font_style = 'normal'
p.yaxis.axis_label_text_font_style = 'normal'
p.xaxis.axis_label_standoff = 15 #Sets the distance of the label from the x-axis in screen units
p.yaxis.axis_label_standoff = 15 #Sets the distance of the label from the y-axis in screen units
#Set the copyright-label position:
label_opts = dict(x=0, y=72,
x_units='screen', y_units='screen')
#Create a label object and format it:
caption1 = Label(text="© ICOS ERIC", **label_opts)
caption1.text_font_size = '8pt'
#Add legend to figure:
p.add_layout(legend, 'below')
#Add label to plot:
p.add_layout(caption1, 'below')
#Disable the scientific output of numbers on y-axis:
p.left[0].formatter.use_scientific = False
#Inactivate hover-tool, which is by default active:
p.toolbar.active_inspect = None
#Set the output location:
output_notebook()
#Show plot:
show(p)
# -
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Function that creates widgets and updates the plot based on the user's selection:
def create_widgets_icos_eco_htm_iterative_sums():
#Import modules:
from ipywidgets import interact_manual, Dropdown
#Create a list with the years for which data exist:
year_ls = [2015, 2016, 2017, 2018]
#Create a list containing the color that corresponds to every year:
colors = ['blue','#abd9e9', 'orange', 'red']
#Create a list to store the different ecosystem variables:
eco_var_ls = [tuple(reversed(tupl)) for tupl in tuple(measurement_dict_eng.items())][:len(measurement_dict_eng)-1]
#Create dropdown-widgets:
eco_vars = Dropdown(options = eco_var_ls)
#Function that updates the plot based on the user's selection:
def update_iter_sums_plot(Variable):
#Check selected variable:
if(Variable == 'TA_1_1_1'):
#Get list of lists of iterative sums for every year for the selected variable:
iter_sum_ls = [HTM_eko_TA_2015_itersum,
HTM_eko_TA_2016_itersum,
HTM_eko_TA_2017_itersum,
HTM_eko_TA_2018_itersum]
#Get variable unit:
var_unit = unit_dict[Variable]
#Define daily aggregation type:
daily_aggr_type = 'Means'
#If the selected variable is Carbon Flux (NEE):
elif(Variable == 'FC_PI_1_1_1'):
#Get list of lists of iterative sums for every year for the selected variable:
#Convert micromoles to moles.
iter_sum_ls = [micromoles2moles(HTM_eko_NEE_2015_itersum),
micromoles2moles(HTM_eko_NEE_2016_itersum),
micromoles2moles(HTM_eko_NEE_2017_itersum),
micromoles2moles(HTM_eko_NEE_2018_itersum)]
#Get variable unit:
var_unit = 'molm-2'.translate(SUP)
#Define daily aggregation type:
daily_aggr_type = 'Totals'
#If the selected variable is Gross Primary Production (GPP):
elif(Variable == 'GPP_PI_1_1_1'):
#Get list of lists of iterative sums for every year for the selected variable:
#Convert micromoles to moles.
iter_sum_ls = [micromoles2moles(HTM_eko_GPP_2015_itersum),
micromoles2moles(HTM_eko_GPP_2016_itersum),
micromoles2moles(HTM_eko_GPP_2017_itersum),
micromoles2moles(HTM_eko_GPP_2018_itersum)]
#Get variable unit:
var_unit = 'molm-2'.translate(SUP)
#Define daily aggregation type:
daily_aggr_type = 'Totals'
#If the selected variable is Gross Primary Production (GPP):
elif(Variable == 'RECO_PI_1_1_1'):
#Get list of lists of iterative sums for every year for the selected variable:
#Convert micromoles to moles.
iter_sum_ls = [micromoles2moles(HTM_eko_RECO_2015_itersum),
micromoles2moles(HTM_eko_RECO_2016_itersum),
micromoles2moles(HTM_eko_RECO_2017_itersum),
micromoles2moles(HTM_eko_RECO_2018_itersum)]
#Get variable unit:
var_unit = 'molm-2'.translate(SUP)
#Define daily aggregation type:
daily_aggr_type = 'Totals'
#If the selected variable is Gross Primary Production (GPP):
elif(Variable == 'P_1_1_1'):
#Get list of lists of iterative sums for every year for the selected variable:
#Convert micromoles to moles.
iter_sum_ls = [HTM_eko_P_2015_itersum,
HTM_eko_P_2016_itersum,
HTM_eko_P_2017_itersum,
HTM_eko_P_2018_itersum]
#Get variable unit:
var_unit = unit_dict[Variable]
#Define daily aggregation type:
daily_aggr_type = 'Totals'
#If the selected variable is Gross Primary Production (GPP):
elif(Variable == 'SW_IN_1_1_1'):
#Get list of lists of iterative sums for every year for the selected variable:
#Convert micromoles to moles.
iter_sum_ls = [Joules2MegaJoules(HTM_eko_SWIR_2015_itersum),
Joules2MegaJoules(HTM_eko_SWIR_2016_itersum),
Joules2MegaJoules(HTM_eko_SWIR_2017_itersum),
Joules2MegaJoules(HTM_eko_SWIR_2018_itersum)]
#Get variable unit:
var_unit = 'molm-2'.translate(SUP)
#Define daily aggregation type:
daily_aggr_type = 'Totals'
#If the selected variable is none of the above print error message:
else:
print("Variable doesn't exist!")
#Call function to show plot:
plotIterSum(iter_sum_ls, year_ls, daily_aggr_type, measurement_dict_eng[Variable], unit_dict_daily[Variable], colors)
#Create function that contains a box of widgets:
interact = interact_manual(update_iter_sums_plot,
Variable = eco_vars)
#Set the font of the widgets included in interact_manual:
interact.widget.children[0].layout.width = '460px'
interact.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact.widget.children[1].description = 'Update Plot'
interact.widget.children[1].button_style = 'danger'
interact.widget.children[1].style.button_color = '#3973ac'
interact.widget.children[1].layout.margin = '20px 10px 40px 200px' # top/right/bottom/left
# -
# <a id='bokeh_plot_stat_summed_values_per_year_plot'></a>
# <br>
# #### Plot with Cumulative Sums of Daily Totals or Daily Means per Year
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to display widgets:
create_widgets_icos_eco_htm_iterative_sums()
# -
# <a id='bokeh_plot_daily_total_GPP_SWIR_per_year'></a>
# <br>
# <br>
#
# <div style="text-align: right">
# <a href="#bokeh_plot_iterative_sums_of_daily_totals_or_means_per_year_intro">[Back to Create Plots of Iterative Sums - Intro]</a>
# </div>
#
# <br>
# <br>
# <br>
# <br>
#
# #### 10.4. Barplot with Incoming Shortwave-Infrared Solar Radiation (Daily Total) & GPP (Daily Total)
# The existance of incoming solar radiation is essential for plants to photosynthesize. Gross Primary Production (GPP) can be used to provide a measure of the magnitude of photosynthetical activity. In this part, it is possible to view an interactive plot of daily totals of incoming shortwave infrared solar radiation and daily totals of GPP per year. The objective is to observe how the availability of shortwave infrared incoming solar radiation influences GPP between different years.
#
#
# A dropdown widget is provided for the user to select a year between 2015-2017. Two plots will be displayed once the user presses the <span style="color:white">
# <span style="background-color:#3973ac">| Update Plot |</span></span> button. The first plot corresponds to the selected year. The second plot displays the variable values for 2018, the year when the drought occured.
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_daily_totals_SWIR_and_GPP_per_year">[Go to plot]</a>
# </div>
# <br>
# <br>
#
# <span style="color:blue">**Rounding Functions**</span>
# <br>
#
# <span style="color:blue">Function --- > Round Up 10 </span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def roundup10(x):
"""
Project: 'ICOS Carbon Portal'
Created: Tue May 07 10:30:00 2018
Last Changed: Tue May 07 10:30:00 2019
Version: 1.0.0
Author(s): Karolina
Description: Function that takes a number as input and rounds it up to the closest "10".
Input parameters: Number (var_name: 'x', var_type: Integer or Float)
Output: Float
"""
#import module:
import math
import numbers
#Check if input parameter is numeric:
if(isinstance(x, numbers.Number)==True):
#Return rounded value:
return int(math.ceil(x / 10.0)) * 10
#If input parameter is NOT numeric, prompt an error message:
else:
print("Input parameter is not numeric!")
# -
# <br>
#
# <span style="color:blue">Function --- > Round Down 10 </span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def rounddown10(x):
"""
Project: 'ICOS Carbon Portal'
Created: Tue May 07 10:30:00 2018
Last Changed: Tue May 07 10:30:00 2019
Version: 1.0.0
Author(s): Karolina
Description: Function that takes a number as input and floors it down to the closest "10".
Input parameters: Number (var_name: 'x', var_type: Integer or Float)
Output: Float
"""
#import module:
import math
import numbers
#Check if input parameter is numeric:
if(isinstance(x, numbers.Number)==True):
#Return rounded value:
return int(math.ceil(x / 10.0)) * 10 -10
#If input parameter is NOT numeric, prompt an error message:
else:
print("Input parameter is not numeric!")
# -
# <br>
#
# <span style="color:blue">Function --- > Round Up 20 </span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def roundup20(x):
"""
Project: 'ICOS Carbon Portal'
Created: Tue May 07 09:00:00 2018
Last Changed: Tue May 07 09:00:00 2019
Version: 1.0.0
Author(s): Karolina
Description: Function that takes a number as input and rounds it up to the closest "20".
Input parameters: Number (var_name: 'x', var_type: Integer or Float)
Output: Float
"""
#Import module:
import math
import numbers
#Check if input parameter is numeric:
if(isinstance(x, numbers.Number)==True):
#for positive numbers, multiples of 20.0:
if((x>=0)&(((x/10.0)%20)%2 == 0)):
return int(math.ceil(x / 10.0)) * 10 +20
#for positive numbers with an even number as 2nd digit:
elif((x>0)&(int(x/10.0)%2==0)):
return int(math.ceil(x / 10.0)) * 10 +10
#for positive and negative numbers, whose 2nd digit is an odd number (except for i in [-1,-9]):
elif(int(x/10.0)%2!=0):
return int((x / 10.0)) * 10 +10
#for negative numbers, whose 1st or 2nd digit is an even number:
elif((x<-10) & (int(x)%2==0)):
return int((x / 10.0)) * 10 +20
else:
return 0
#If input parameter is NOT numeric, prompt an error message:
else:
print("Input parameter is not numeric!")
# -
# <br>
#
# <span style="color:blue">Function --- > Round Down 20 </span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def rounddown20(x):
"""
Project: 'ICOS Carbon Portal'
Created: Tue May 07 09:00:00 2018
Last Changed: Tue May 07 09:00:00 2019
Version: 1.0.0
Author(s): Karolina
Description: Function that takes a number as input and floors it to the nearest "20".
Input parameters: Number (var_name: 'x', var_type: Integer or Float)
Output: Float
"""
#Import module:
import math
import numbers
#Check if input parameter is numeric:
if(isinstance(x, numbers.Number)==True):
#If the 2nd digit from the decimal point is an even number:
if(int(x/10.0)%2==0):
return(int(x / 10.0) * 10) - 20
#If the 2nd digit from the decimal point is an odd number:
else:
return(int(x / 10.0) * 10) - 10
#If input parameter is not numeric, prompt an error message:
else:
print("Input parameter is not numeric!")
# -
# <br>
#
# <span style="color:blue">**Function - Define the range of the y-axes (2 y-axes)**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def set_yranges_2y_barplot(y1_min, y1_max, y2_min, y2_max, y1_step, y2_step ,new_yrange_name):
"""
Project: 'ICOS Carbon Portal'
Created: Tue May 07 10:30:00 2018
Last Changed: Tue May 07 10:30:00 2019
Version: 1.0.0
Author(s): Karolina
Description: Function that takes the primary and secondary y-axis min/max values as well as
the step values for every y-axis and the secondary y-axis new range name as input
parameters, performs computations so that the two axes are alligned and returns
their corresponding RangeId objects. Works only for Bokeh plots.
Input parameters: 1. Min value of primary y-axis (var_name: 'y1_min', var_type: Integer or Float)
2. Max value of primary y-axis (var_name: 'y1_max', var_type: Integer or Float)
3. Min value of secondary y-axis (var_name: 'y2_min', var_type: Integer or Float)
4. Max value of secondary y-axis (var_name: 'y2_max', var_type: Integer or Float)
5. Step of primary y-axis (var_name: 'y1_step', var_type: Integer or Float)
6. Step of secondary y-axis (var_name: 'y2_step', var_type: Integer or Float)
7. Name of new yrange object for secondary y-axis
(var_name: "new_yrange_name", var_type: Bokeh Plot yrange object)
Output: Bokeh Plot yrange objects for primary and secondary y-axes.
"""
#import modules:
import numpy as np
from bokeh.models import Range1d
#yrange and tick function for plot with primary and secondary y-axis:
yticks1 = np.arange(y1_min, y1_max + y1_step, y1_step)
yticks2 = np.arange(y2_min, y2_max + y2_step, y2_step)
#Get difference in total number of ticks between primary and secondary y-axis:
diff = abs(len(yticks2)-len(yticks1))
#Get how many times the step needs to be added to start and end:
num_of_steps = int(diff/2)
#If the primary and the secondary y-axis have the same number of ticks:
if(diff==0):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min, end=y1_max)
#Set the 2nd y-axis, range-name, range:
extra_y_ranges = {new_yrange_name: Range1d(start=y2_min, end=y2_max)}
#print('diff==0')
#If the primary y-axis has fewer ticks than the secondary y-axis:
elif(len(yticks2)>len(yticks1)):
#If the difference in ticks between the two axes is an odd number:
if(diff%2==1):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min-(y1_step*(num_of_steps+1)), end=y1_max+(y1_step*num_of_steps))
#Set the 2nd y-axis, range-name, range:
extra_y_ranges = {new_yrange_name: Range1d(start=y2_min, end=y2_max)}
#print('len(yticks2)>len(yticks1) --> diff==odd')
#If the difference in ticks between the two axes is an even number:
else:
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min-(y1_step*num_of_steps), end=y1_max+(y1_step*num_of_steps))
#Set the 2nd y-axis, range-name, range:
extra_y_ranges = {new_yrange_name: Range1d(start=y2_min, end=y2_max)}
#print('len(yticks2)>len(yticks1) --> diff==even')
#If the primary y-axis has more ticks than the secondary y-axis, e.g. len(yticks1)>len(yticks2_test):
else:
#If the difference in ticks between the two axes is an odd number:
if(diff%2==1):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min, end=y1_max)
#Set the 2nd y-axis, range-name, range:
extra_y_ranges = {new_yrange_name: Range1d(start=y2_min + (y2_step*(num_of_steps)), end=y2_max + (y2_step*(num_of_steps+1)))}
#print('len(yticks2)<len(yticks1) --> diff==odd')
#If the difference in ticks between the two axes is an even number:
else:
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min, end=y1_max)
#Set the 2nd y-axis, range-name, range:
#extra_y_ranges = {new_yrange_name: Range1d(start=y2_min - (y2_step*num_of_steps), end=y2_max + (y2_step*num_of_steps))}
extra_y_ranges = {new_yrange_name: Range1d(start=y2_min, end=y2_max + (y2_step*(num_of_steps+1)))}
#print('len(yticks2)<len(yticks1) --> diff==even')
#Return y-range for primary and secondary y-axes:
return y_range, extra_y_ranges
# -
# <br>
#
# <span style="color:blue">**Plotting Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Plot daily totals per year for a given variable:
def plot_barplot_2axes(df1, df2, variable_ls, unit_ls, dailyType_ls, color_ls):
#Import modules:
from bokeh.models import Legend
p = figure(plot_width=600, plot_height=450,
title = 'HTM: '+variable_ls[0]+' Daily '+dailyType_ls[0]+', '+
variable_ls[1]+' Daily '+dailyType_ls[1]+' for '+str(df1.index[0].year),
x_axis_label = 'Time',
y_axis_label = variable_ls[0] + ' ('+unit_ls[0].translate(SUP)+') daily '+dailyType_ls[0],
x_axis_type='datetime')
# Setting the second y axis range name and range
p.y_range, p.extra_y_ranges = set_yranges_2y_barplot(0,#rounddown10(df1.values.min()),
roundup20(df1.values.max()),
0,#math.floor(df2.values.min()),
math.ceil(df2.values.max()), 10.0, 0.5, 'y2')
#Set primary y-axis ticker:
ticker_1 = SingleIntervalTicker(interval= 10.0)
#Add primary y-axis ticker to plot:
p.yaxis.ticker = ticker_1
#Set secondary y-axis ticker:
ticker_2 = SingleIntervalTicker(interval=0.5)
# Adding the second axis to the plot.
p.add_layout(LinearAxis(y_range_name="y2",
axis_label=variable_ls[1] + ' ('+unit_ls[1].translate(SUP)+')',
ticker=ticker_2,
axis_label_standoff = 15,
axis_label_text_color = color_ls[1]), 'right')
#Create an empty list that will store the legend info:
legend_it = []
#Add 1st barplot:
bp1 = p.vbar(x=list(df1.index.values), width=2.5, bottom=0,
top=list(df1.values), color=color_ls[0], name=variable_ls[0])
#Add 2nd barplot:
bp2 = p.vbar(x=list(df2.index.values), width=2.5, bottom=0, alpha=0.4,
top=list(df2.values), color=color_ls[1], y_range_name="y2", name=variable_ls[1])
#Add the name and glyph info (i.e. colour and marker type) to the legend:
legend_it.append((bp1.name, [bp1]))
legend_it.append((bp2.name, [bp2]))
#Create legend:
legend = Legend(items=legend_it, location= 'bottom_center')
legend.orientation = 'horizontal'
legend.click_policy='hide'
legend.spacing = 10 #sets the distance between legend entries
#Add legend to figure:
p.add_layout(legend, 'below')
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '12pt'
p.title.vertical_align = 'top' #Create a distance between the title and the plot
#Set axis label font style:
p.xaxis.axis_label_text_font_style = 'normal'
p.yaxis.axis_label_text_font_style = 'normal'
p.xaxis.axis_label_standoff = 15 #Sets the distance of the label from the x-axis in screen units
p.yaxis.axis_label_standoff = 15 #Sets the distance of the label from the y-axis in screen units
p.yaxis[0].axis_label_text_color = color_ls[0]
#Set the copyright-label position:
label_opts = dict(x=0, y=5,
x_units='screen', y_units='screen')
#Create a label object and format it:
caption1 = Label(text="© ICOS ERIC", **label_opts)
caption1.text_font_size = '8pt'
#Add label to plot:
p.add_layout(caption1, 'below')
#Format plot borders:
p.min_border_top = 54
#Return plot:
return p
# -
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def widget_SWIR_GPP():
#Import Python modules:
from ipywidgets import interact_manual, Dropdown
from bokeh.layouts import column
from bokeh.io import show, output_notebook
#Create a dictionary to store the filenames associated with each year:
labels = {"2015":[Joules2MegaJoules(HTM_eko_LIGHT_2015_daily_sum), micromoles2moles(HTM_eko_GPP_2015_daily_sum)],
"2016":[Joules2MegaJoules(HTM_eko_LIGHT_2016_daily_sum), micromoles2moles(HTM_eko_GPP_2016_daily_sum)],
"2017":[Joules2MegaJoules(HTM_eko_LIGHT_2017_daily_sum), micromoles2moles(HTM_eko_GPP_2017_daily_sum)]}
#Create Dropdown-List widget:
years = Dropdown(options=labels.keys(),
value='2015',
description='Year:',
disabled=False)
#Function that calls functions to update the plot
#based on the selected year:
def update_plot_func(Year):
#Call function to plot data for the selected year:
p1 = plot_barplot_2axes(labels[Year][0],
labels[Year][1],
['SW-IR','GPP'],
['MJoules/m2','moles m-2'],
['Total', 'Total'],
['orange','green'])
#Show Plot for 2018:
p2 = plot_barplot_2axes(Joules2MegaJoules(HTM_eko_LIGHT_2018_daily_sum),
micromoles2moles(HTM_eko_GPP_2018_daily_sum),
['SW-IR','GPP'],
['MJoules/m2','moles m-2'],
['Total', 'Total'],
['orange','green'])
#Define output location:
output_notebook()
#Show plots:
show(column(p1, p2))
#Create function that contains a box of widgets:
interact_c = interact_manual(update_plot_func,
Year=years)
#Set the font of the widgets included in interact_manual:
interact_c.widget.children[0].layout.width = '430px'
interact_c.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact_c.widget.children[1].description = 'Update Plot'
interact_c.widget.children[1].button_style = 'danger'
interact_c.widget.children[1].style.button_color = '#3973ac'
interact_c.widget.children[1].layout.margin = '10px 10px 40px 180px' # top/right/bottom/left
# -
# <a id='bokeh_plot_daily_totals_SWIR_and_GPP_per_year'></a>
# <br>
# #### Plot Daily Total Shortwave Infrared Incoming Solar Radiation with Daily Total GPP
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to display widgets:
widget_SWIR_GPP()
# -
# <a id='bokeh_plot_daily_total_GPP_daily_mean_SWC_per_year'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
# <br>
# <br>
#
# #### 10.5. Plot GPP Daily Totals with Soil Water Content Daily Mean
# The amount of water available to the plants, can affect their rate of photosynthesis. If the amount of water in the soil drops bellow 10%, the plants can no longer absorb it with their roots. Plants that do not have enough water, close their stomata and loose their capacity to take in CO$_2$. This means that they cease to photosynthesize.
#
#
# In this part, it is possible to view an interactive plot with daily mean Soil Water Content values and daily total GPP values for the duration of one year. A dropdown widget allows the user to select a year between 2015 and 2017. Once the user clicks on the <span style="color:white">
# <span style="background-color:#3973ac">| Update Plot |</span></span>-button, two plots will appear. The first plot depicts the values of the aforementioned variables for the selected year, whilst the second plot shows the variable values for 2018, when the drought occured. The interactive legend allows the user to switch layers on and off.
#
#
# The purpose of these visualizations is to examine if changes in Soil Water Content correlate with changes in GPP.
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_stat_summed_daily_totals_means_GPP_SWC_per_year">[Go to plot]</a>
# </div>
#
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def widget_SWC_GPP():
#Import Python modules:
from ipywidgets import interact_manual, Dropdown
from bokeh.layouts import column
from bokeh.io import show, output_notebook
#Create a dictionary to store the filenames associated with each year:
labels = {"2015":[HTM_eko_SWC_2015_daily_mean, micromoles2moles(HTM_eko_GPP_2015_daily_sum)],
"2016":[HTM_eko_SWC_2016_daily_mean, micromoles2moles(HTM_eko_GPP_2016_daily_sum)],
"2017":[HTM_eko_SWC_2017_daily_mean, micromoles2moles(HTM_eko_GPP_2017_daily_sum)]}
#Create Dropdown-List widget:
years = Dropdown(
options=labels.keys(),
value='2015',
description='Year:',
disabled=False
)
#Function that calls functions to update the plot
#based on the selected year:
def update_plot_func(Year):
#Call function to plot data for the selected year:
p1 = plot_barplot_2axes(labels[Year][0],
labels[Year][1],
['Soil Water Content','GPP'],
['%','moles m-2'],
['Mean', 'Total'],
['lightblue','green'])
#Plot Figure for 2018:
p2 = plot_barplot_2axes(HTM_eko_SWC_2018_daily_mean,
micromoles2moles(HTM_eko_GPP_2018_daily_sum),
['Soil Water Content','GPP'],
['%','moles m-2'],
['Mean', 'Total'],
['lightblue','green'])
#Define output location:
output_notebook()
#Show plots:
show(column(p1, p2))
#Create function that contains a box of widgets:
interact_c = interact_manual(update_plot_func,
Year=years)
#Set the font of the widgets included in interact_manual:
interact_c.widget.children[0].layout.width = '430px'
interact_c.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact_c.widget.children[1].description = 'Update Plot'
interact_c.widget.children[1].button_style = 'danger'
interact_c.widget.children[1].style.button_color = '#3973ac'
interact_c.widget.children[1].layout.margin = '10px 10px 40px 180px' # top/right/bottom/left
# -
# <a id='bokeh_plot_stat_summed_daily_totals_means_GPP_SWC_per_year'></a>
# <br>
# #### Plot Daily Mean Soil Water Content with Daily Total GPP
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to plot widgets:
widget_SWC_GPP()
# -
# <a id='bokeh_plot_daily_total_RECO_daily_mean_SWC_and_TA_per_year'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
#
# #### 10.6. Plot Daily Mean Soil Water Content with Daily Total Respiration and Daily Mean Air Temperature
# Decomposers and detritivores like warm and moist environments. They emit carbon to the atmosphere as a result of theri activity. As mentioned before, decomposers and detritivores would limit their activity if the conditions of their environment became too dry. The following plots will present how the values of Air Temperature and Soil Water Content correlate with the values of Respiration.
#
# In this part, it is possible to view an interactive plot with daily mean Soil Water Content values and daily total Respiration and Air Temperature values for the duration of one year. A dropdown widget allows the user to select a year between 2015 and 2017. Once the user clicks on the <span style="color:white">
# <span style="background-color:#3973ac">| Update Plot |</span></span> button, two plots will appear. The first plot depicts the values of the aforementioned variables for the selected year, whilst the second plot shows the variable values for 2018, when the drought occured. The interactive legend allows the user to switch layers on and off.
#
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_stat_summed_daily_totals_means_RECO_SWC_Temp_per_year">[Go to plot]</a>
# </div>
#
# <br>
#
# <span style="color:blue">**Function - Define the range of the y-axes (3 y-axes)**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def set_yranges_3y_ymin0(y1_min, y1_max, y2_min, y2_max, y3_min, y3_max, y1_step, y2_step, y3_step):
"""
Project: 'ICOS Carbon Portal'
Created: Tue May 07 10:30:00 2018
Last Changed: Tue May 07 10:30:00 2019
Version: 1.0.0
Author(s): Karolina
Description: Function that takes the primary, secondary and third y-axis min/max values as well as
the step values for every y-axis as input parameters, performs computations so that the
three axes are alligned and returns their corresponding RangeId objects.
Works only for Bokeh plots.
Input parameters: 1. Min value of primary y-axis (var_name: 'y1_min', var_type: Integer or Float)
2. Max value of primary y-axis (var_name: 'y1_max', var_type: Integer or Float)
3. Min value of secondary y-axis (var_name: 'y2_min', var_type: Integer or Float)
4. Max value of secondary y-axis (var_name: 'y2_max', var_type: Integer or Float)
5. Min value of third y-axis (var_name: 'y3_min', var_type: Integer or Float)
6. Max value of third y-axis (var_name: 'y3_max', var_type: Integer or Float)
7. Step of primary y-axis (var_name: 'y1_step', var_type: Integer or Float)
8. Step of secondary y-axis (var_name: 'y2_step', var_type: Integer or Float)
9. Step of third y-axis (var_name: 'y3_step', var_type: Integer or Float)
Output: Bokeh Plot yrange objects for primary and secondary y-axes.
"""
#import modules:
import numpy as np
from bokeh.models import Range1d
#yrange and tick function for plot with primary and secondary y-axis:
yticks1 = np.arange(y1_min, y1_max + y1_step, y1_step)
yticks2 = np.arange(y2_min, y2_max + y2_step, y2_step)
yticks3 = np.arange(y3_min, y3_max + y3_step, y3_step)
#Get the number of ticks per y-axis:
y1_num_of_ticks = len(yticks1)
y2_num_of_ticks = len(yticks2)
y3_num_of_ticks = len(yticks3)
#Get difference in total number of ticks between primary and secondary y-axis:
diff_12 = abs(len(yticks2)-len(yticks1))
diff_13 = abs(len(yticks3)-len(yticks1))
diff_23 = abs(len(yticks3)-len(yticks2))
#If the primary, secondary and 3rd y-axis have the same number of ticks:
if((diff_12==0) and (diff_13==0) and (diff_23==0)):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min, end=y1_max)
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min, end=y2_max)
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min, end=y3_max)
#print('All y-axes have the same num of ticks')
#if y-axis 1 is the axis with the highest number of ticks:
elif(max(y1_num_of_ticks, y2_num_of_ticks, y3_num_of_ticks)==y1_num_of_ticks):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min, end=y1_max)
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min,
end=y2_max + (y2_step*diff_12))
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min,
end=y3_max + (y3_step*diff_13))
#print('y1axis highest num of ticks')
#if y-axis 2 is the axis with the highest number of ticks:
elif(max(y1_num_of_ticks, y2_num_of_ticks, y3_num_of_ticks)==y2_num_of_ticks):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min,
end=y1_max+(y1_step*diff_12))
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min, end=y2_max)
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min,
end=y3_max+(y3_step*diff_23))
#print('y2axis highest num of ticks')
#if y-axis 3 is the axis with the highest number of ticks:
elif(max(y1_num_of_ticks, y2_num_of_ticks, y3_num_of_ticks)==y3_num_of_ticks):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min,
end=y1_max+(y1_step*diff_13))
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min,
end=y2_max+(y2_step*diff_23))
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min, end=y3_max)
#print('y3axis highest num of ticks')
else:
y_range = None
extra_y_ranges_1 = None
extra_y_ranges_2 = None
#Return y-range for primary and secondary y-axes:
return y_range, extra_y_ranges_1, extra_y_ranges_2
# -
# <br>
#
# <span style="color:blue">**Plotting Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Plot daily totals per year for a given variable:
def plot_2barplots_line_glyph_3axes(df1, df2, df3, variable_ls, unit_ls, dailyType_ls, color_ls, step_y1, step_y2, step_y3):
#Import modules:
from bokeh.models import Legend
#Create Fogure Object:
p = figure(plot_width=600, plot_height=450,
title = 'HTM: '+variable_ls[1]+', '+variable_ls[0]+' & '+variable_ls[2]+
' for '+str(df1.index[0].year),
x_axis_label = 'Time',
y_axis_label = variable_ls[0] + ' ('+unit_ls[0].translate(SUP)+') Daily '+dailyType_ls[0],
x_axis_type='datetime')
#Add the ranges for every y-axis:
p.y_range, p.extra_y_ranges['Yaxis2'], p.extra_y_ranges['Yaxis3']= set_yranges_3y_ymin0(0,
roundup10(df1.values.max()),
0,
math.ceil(df2.values.max()),
0,
roundup10(df3.values.max()),
step_y1,
step_y2,
step_y3)
#Set primary y-axis ticker:
ticker_1 = SingleIntervalTicker(interval= step_y1)
#Add primary y-axis ticker to plot:
p.yaxis.ticker = ticker_1
#Set secondary y-axis ticker:
ticker_2 = SingleIntervalTicker(interval=step_y2)
#Set secondary y-axis ticker:
ticker_3 = SingleIntervalTicker(interval=step_y3)
# Adding the second axis to the plot.
yaxis2 = LinearAxis(y_range_name="Yaxis2",
axis_label=variable_ls[1] + ' ('+unit_ls[1].translate(SUP)+') Daily '+dailyType_ls[1],
ticker=ticker_2,
axis_label_standoff = 15,
axis_label_text_color = color_ls[1])
# Adding the third axis to the plot.
yaxis3 = LinearAxis(y_range_name='Yaxis3',
axis_label=variable_ls[2] + ' ('+unit_ls[2].translate(SUP)+') Daily '+dailyType_ls[2],
ticker=ticker_3,
axis_label_standoff = 15,
axis_label_text_color = color_ls[2])
#Define at which part of the plot the additional y-axes will be located:
p.add_layout(yaxis2,'right')
p.add_layout(yaxis3,'right')
#Create an empty list that will store the legend info:
legend_it = []
#Create 1st barplot:
bp1 = p.vbar(x=list(df1.index.values), width=0.5, bottom=0,
top=list(df1.values), color=color_ls[0], name=variable_ls[0])
#Create 2nd barplot:
bp2 = p.vbar(x=list(df2.index.values), width=0.5, bottom=0, alpha=0.5,
top=list(df2.values), color=color_ls[1], y_range_name="Yaxis2", name=variable_ls[1])
#Add line-glyph:
g1 = p.line(df3.index.values, df3.values, line_width=2.0,
color=color_ls[2], y_range_name="Yaxis3", name=variable_ls[2])
#Add the name and glyph info (i.e. colour and marker type) to the legend:
legend_it.append((bp1.name, [bp1]))
legend_it.append((bp2.name, [bp2]))
legend_it.append((g1.name, [g1]))
#Create legend:
legend = Legend(items=legend_it, location= 'bottom_center')
legend.orientation = 'horizontal'
legend.click_policy='hide'
legend.spacing = 10 #sets the distance between legend entries
#Add legend to figure:
p.add_layout(legend, 'below')
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '10pt'
p.title.vertical_align = 'top' #Create a distance between the title and the plot
#Set axis label font style:
p.xaxis.axis_label_text_font_style = 'normal'
p.yaxis.axis_label_text_font_style = 'normal'
p.xaxis.axis_label_standoff = 15 #Sets the distance of the label from the x-axis in screen units
p.yaxis.axis_label_standoff = 15 #Sets the distance of the label from the y-axis in screen units
p.yaxis[0].axis_label_text_color = color_ls[0]
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '12pt'
p.title.offset = 15
#Set the copyright-label position:
label_opts = dict(x=0, y=5,
x_units='screen', y_units='screen')
#Create a label object and format it:
caption1 = Label(text="© ICOS ERIC", **label_opts)
caption1.text_font_size = '8pt'
#Add label to plot:
p.add_layout(caption1, 'below')
#Format plot borders:
p.min_border_top = 54
#Return Figure Object:
return p
# -
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def widgetAirTempRECO():
#Import Python modules:
from ipywidgets import interact_manual, Dropdown
from bokeh.layouts import column
from bokeh.io import show, output_notebook
#Create a dictionary to store the filenames associated with each year:
labels = {"2015":[HTM_eko_SWC_2015_daily_mean,
micromoles2moles(HTM_eko_RECO_2015_daily_sum),
HTM_eko_TA_2015_daily_mean],
"2016":[HTM_eko_SWC_2016_daily_mean,
micromoles2moles(HTM_eko_RECO_2016_daily_sum),
HTM_eko_TA_2016_daily_mean],
"2017":[HTM_eko_SWC_2017_daily_mean,
micromoles2moles(HTM_eko_RECO_2017_daily_sum),
HTM_eko_TA_2017_daily_mean]}
#Create Dropdown-List widget:
years = Dropdown(
options=labels.keys(),
value='2015',
description='Year:',
disabled=False
)
#Function that calls functions to update the plot
#based on the selected year:
def update_plot_func(Year):
#Call function to plot data for the selected year:
p1 = plot_2barplots_line_glyph_3axes(labels[Year][0],
labels[Year][1],
labels[Year][2],
['Soil Water Content','Respiration','Air Temperature'],
['%','moles m-2','C\u00b0'],
['Mean', 'Total','Mean'],
['lightblue', '#9e9ac8','firebrick'],
10.0, 1.0, 10.0)
#Call function to plot data for 2018 (drought year):
p2 = plot_2barplots_line_glyph_3axes(HTM_eko_SWC_2018_daily_mean,
micromoles2moles(HTM_eko_RECO_2018_daily_sum),
HTM_eko_TA_2018_daily_mean,
['Soil Water Content','Respiration','Air Temperature'],
['%','moles m-2','C\u00b0'],
['Mean', 'Total','Mean'],
['lightblue', '#9e9ac8','firebrick'],
10.0, 1.0, 10.0)
#Define output location:
output_notebook()
#Show plots:
show(column(p1, p2))
#Create function that contains a box of widgets:
interact_c = interact_manual(update_plot_func,
Year=years)
#Set the font of the widgets included in interact_manual:
interact_c.widget.children[0].layout.width = '430px'
interact_c.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact_c.widget.children[1].description = 'Update Plot'
interact_c.widget.children[1].button_style = 'danger'
interact_c.widget.children[1].style.button_color = '#3973ac'
interact_c.widget.children[1].layout.margin = '10px 10px 40px 180px' # top/right/bottom/left
# -
# <a id='bokeh_plot_stat_summed_daily_totals_means_RECO_SWC_Temp_per_year'></a>
# <br>
# #### Plot Daily Mean Soil Water Content and Air Temperature with Daily Total Respiration
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to display widgets:
widgetAirTempRECO()
# -
# <a id='bokeh_plot_daily_mean_SWC_and_daily_total_GPP_and_Precip_per_year'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
#
# #### 10.7. Plot Daily Mean Soil Water Content with Daily Total GPP and Daily Total Precipitation
# Plants need access to water in order to photosynthesize. Water-stressed plants close their stomata and limit their photosynthetic activity to preserve water and survive. If the level of soil water content drops below a threshold of 10%, then the plants are no longer able to absorb water from the soil using their roots.
#
# In this part, it is possible to view an interactive plot with daily mean Soil Water Content values and daily total GPP and Precipitation values for the duration of one year. A dropdown widget allows the user to select a year between 2015 and 2017. Once the user clicks on the <span style="color:white">
# <span style="background-color:#3973ac">| Update Plot |</span></span> button, two plots will appear. The first plot depicts the values of the aforementioned variables for the selected year, whilst the second plot shows the variable values for 2018, when the drought occured. The interactive legend allows the user to switch layers on and off.
#
# The objective here is to examine how soil water content is affected by precipitation and how soil water content affects GPP.
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_stat_summed_daily_totals_means_SWC_GPP_Precip_per_year">[Go to plot]</a>
# </div>
#
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def widgetSWCGPPPrecip():
#Import Python modules:
from ipywidgets import interact_manual, Dropdown
from bokeh.layouts import column
from bokeh.io import show, output_notebook
#Create a dictionary to store the filenames associated with each year:
labels = {"2015":[HTM_eko_SWC_2015_daily_mean,
micromoles2moles(HTM_eko_GPP_2015_daily_sum),
HTM_eko_P_2015_daily_sum],
"2016":[HTM_eko_SWC_2016_daily_mean,
micromoles2moles(HTM_eko_GPP_2016_daily_sum),
HTM_eko_P_2016_daily_sum],
"2017":[HTM_eko_SWC_2017_daily_mean,
micromoles2moles(HTM_eko_GPP_2017_daily_sum),
HTM_eko_P_2017_daily_sum]}
#Create Dropdown-List widget:
years = Dropdown(options=labels.keys(),
value='2015',
description='Year:',
disabled=False)
#Function that calls functions to update the plot
#based on the selected year:
def update_plot_func(Year):
#Call function to plot data for the selected year:
p1 = plot_2barplots_line_glyph_3axes(labels[Year][0],
labels[Year][1],
labels[Year][2],
['Soil Water Content','GPP', 'Precipitation'],
['%','moles m-2', 'mm'],
['Mean', 'Total', 'Total'],
['lightblue', 'green', 'navy'],
10.0,
0.5,
10.0)
#Call function to plot data for 2018 (drought year):
p2 = plot_2barplots_line_glyph_3axes(HTM_eko_SWC_2018_daily_mean,
micromoles2moles(HTM_eko_GPP_2018_daily_sum),
HTM_eko_P_2018_daily_sum,
['Soil Water Content','GPP', 'Precipitation'],
['%','moles m-2', 'mm'],
['Mean', 'Total', 'Total'],
['lightblue', 'green', 'navy'],
10,
0.5,
10.0)
#Define output location:
output_notebook()
#Show plots:
show(column(p1, p2))
#Create function that contains a box of widgets:
interact_c = interact_manual(update_plot_func,
Year=years)
#Set the font of the widgets included in interact_manual:
interact_c.widget.children[0].layout.width = '430px'
interact_c.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact_c.widget.children[1].description = 'Update Plot'
interact_c.widget.children[1].button_style = 'danger'
interact_c.widget.children[1].style.button_color = '#3973ac'
interact_c.widget.children[1].layout.margin = '10px 10px 40px 180px' # top/right/bottom/left
# -
# <a id='bokeh_plot_stat_summed_daily_totals_means_SWC_GPP_Precip_per_year'></a>
# <br>
# #### Plot Daily Mean Soil Water Content with Daily Total GPP and Precipitation
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to display widgets:
widgetSWCGPPPrecip()
# -
# <a id='bokeh_plot_daily_mean_SWC_and_TA_and_daily_total_GPP_and_SWIR_per_year'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
#
# #### 10.8. Plot Daily Mean Soil Water Content and Air temperature with Daily Total GPP and Light (SW-IR)
# Plants need access to water and sunlight in order to photosynthesize. Water-stressed plants close their stomata and limit their photosynthetic activity to preserve water and survive. If the level of soil water content drops below a threshold of 10%, then the plants are no longer able to absorb water from the soil using their roots.
#
# In this part, it is possible to view an interactive plot with daily mean Soil Water Content and Air Temperature values and daily total GPP and Light values for the duration of one year. A dropdown widget allows the user to select a year between 2015 and 2017. Once the user clicks on the <span style="color:white">
# <span style="background-color:#3973ac">| Update Plot |</span></span> button, two plots will appear. The first plot depicts the values of the aforementioned variables for the selected year, whilst the second plot shows the variable values for 2018, when the drought occured. The interactive legend allows the user to switch layers on and off.
#
# The objective here is to examine how the values of the main factors affecting the photosynthetic ability of plants correlate with GPP.
#
# <br>
# <br>
# <div style="text-align: right">
# <a href="#bokeh_plot_stat_summed_daily_totals_means_SWC_GPP_Temp_SWIR_per_year">[Go to plot]</a>
# </div>
#
# <br>
#
# <span style="color:blue">**Function - Define the range of the y-axes (4 y-axes)**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def set_yranges_4y_ymin0(y1_min, y1_max, y2_min, y2_max, y3_min, y3_max, y4_min, y4_max, y1_step, y2_step, y3_step, y4_step):
"""
Project: 'ICOS Carbon Portal'
Created: Tue May 07 10:30:00 2018
Last Changed: Tue May 07 10:30:00 2019
Version: 1.0.0
Author(s): Karolina
Description: Function that takes the primary, secondary, third and fourth y-axis min/max values as well as
the step values for every y-axis as input parameters, performs computations so that the
three axes are alligned and returns their corresponding RangeId objects.
Works only for Bokeh plots.
Input parameters: 1. Min value of primary y-axis (var_name: 'y1_min', var_type: Integer or Float)
2. Max value of primary y-axis (var_name: 'y1_max', var_type: Integer or Float)
3. Min value of secondary y-axis (var_name: 'y2_min', var_type: Integer or Float)
4. Max value of secondary y-axis (var_name: 'y2_max', var_type: Integer or Float)
5. Min value of third y-axis (var_name: 'y3_min', var_type: Integer or Float)
6. Max value of third y-axis (var_name: 'y3_max', var_type: Integer or Float)
7. Min value of fourth y-axis (var_name: 'y4_min', var_type: Integer or Float)
8. Max value of fourth y-axis (var_name: 'y4_max', var_type: Integer or Float)
9. Step of primary y-axis (var_name: 'y1_step', var_type: Integer or Float)
10. Step of secondary y-axis (var_name: 'y2_step', var_type: Integer or Float)
11. Step of third y-axis (var_name: 'y3_step', var_type: Integer or Float)
12. Step of fourth y-axis (var_name: 'y4_step', var_type: Integer or Float)
Output: Bokeh Plot yrange objects for primary and secondary y-axes.
"""
#import modules:
import numpy as np
from bokeh.models import Range1d
#yrange and tick function for plot with primary, secondary, third and fourth y-axis:
yticks1 = np.arange(y1_min, y1_max + y1_step, y1_step)
yticks2 = np.arange(y2_min, y2_max + y2_step, y2_step)
yticks3 = np.arange(y3_min, y3_max + y3_step, y3_step)
yticks4 = np.arange(y4_min, y4_max + y4_step, y4_step)
#Get the number of ticks per y-axis:
y1_num_of_ticks = len(yticks1)
y2_num_of_ticks = len(yticks2)
y3_num_of_ticks = len(yticks3)
y4_num_of_ticks = len(yticks4)
#Get difference in total number of ticks between primary and secondary y-axis:
diff_12 = abs(len(yticks2)-len(yticks1))
diff_13 = abs(len(yticks3)-len(yticks1))
diff_23 = abs(len(yticks3)-len(yticks2))
diff_14 = abs(len(yticks4)-len(yticks1))
diff_24 = abs(len(yticks4)-len(yticks2))
diff_34 = abs(len(yticks4)-len(yticks3))
#If the primary, secondary and 3rd y-axis have the same number of ticks:
if((diff_12==0) and (diff_13==0) and (diff_23==0) and (diff_14==0) and (diff_24==0) and (diff_34==0)):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min, end=y1_max)
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min, end=y2_max)
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min, end=y3_max)
#Set the 4th y-axis, range-name, range:
extra_y_ranges_3 = Range1d(start=y4_min, end=y4_max)
#print('All y-axes have the same length')
#if y-axis 1 is the axis with the highest number of ticks:
elif(max(y1_num_of_ticks, y2_num_of_ticks, y3_num_of_ticks, y4_num_of_ticks)==y1_num_of_ticks):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min, end=y1_max)
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min,
end=y2_max + (y2_step*diff_12))
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min,
end=y3_max + (y3_step*diff_13))
#Set the 4th y-axis, range-name, range:
extra_y_ranges_3 = Range1d(start=y4_min,
end=y4_max + (y4_step*diff_14))
#print('y1-axis --> highest num of ticks')
#if y-axis 2 is the axis with the highest number of ticks:
elif(max(y1_num_of_ticks, y2_num_of_ticks, y3_num_of_ticks, y4_num_of_ticks)==y2_num_of_ticks):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min,
end=y1_max+(y1_step*diff_12))
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min, end=y2_max)
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min,
end=y3_max+(y3_step*diff_23))
#Set the 4th y-axis, range-name, range:
extra_y_ranges_3 = Range1d(start=y4_min,
end=y4_max+(y4_step*diff_24))
#print('y2-axis --> highest num of ticks')
#if y-axis 3 is the axis with the highest number of ticks:
elif(max(y1_num_of_ticks, y2_num_of_ticks, y3_num_of_ticks, y4_num_of_ticks)==y3_num_of_ticks):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min,
end=y1_max+(y1_step*diff_13))
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min,
end=y2_max+(y2_step*diff_23))
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min, end=y3_max)
#Set the 4th y-axis, range-name, range:
extra_y_ranges_3 = Range1d(start=y4_min,
end=y4_max+(y4_step*diff_34))
#print('y3-axis --> highest num of ticks')
#if y-axis 4 is the axis with the highest number of ticks:
elif(max(y1_num_of_ticks, y2_num_of_ticks, y3_num_of_ticks, y4_num_of_ticks)==y4_num_of_ticks):
#Set the range of the 1st y-axis:
y_range = Range1d(start=y1_min,
end=y1_max+(y1_step*diff_14))
#Set the 2nd y-axis, range-name, range:
extra_y_ranges_1 = Range1d(start=y2_min,
end=y2_max+(y2_step*diff_24))
#Set the 3rd y-axis, range-name, range:
extra_y_ranges_2 = Range1d(start=y3_min,
end=y3_max+(y3_step*diff_34))
#Set the 4th y-axis, range-name, range:
extra_y_ranges_3 = Range1d(start=y4_min, end=y4_max)
#print('y4-axis --> highest num of ticks')
else:
y_range = None
extra_y_ranges_1 = None
extra_y_ranges_2 = None
#Return y-range for all y-axes:
return y_range, extra_y_ranges_1, extra_y_ranges_2, extra_y_ranges_3
# -
# <br>
#
# <span style="color:blue">**Plotting unction**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Plot daily totals per year for a given variable:
def plotGPPLightSWCTempYr_4axes(df1, df2, df3, df4,
variable_ls, unit_ls, dailyType_ls, color_ls,
step_y1, step_y2, step_y3, step_y4):
p = figure(plot_width=600, plot_height=450,
title = 'Hyltemossa: '+variable_ls[0]+', '+variable_ls[1]+', '+
variable_ls[2]+' & '+variable_ls[3]+' for '+str(df1.index[0].year),
x_axis_label = 'Time',
y_axis_label = variable_ls[0] + ' ('+unit_ls[0].translate(SUP)+') daily '+dailyType_ls[0],
x_axis_type='datetime')
#Add the ranges for every y-axis:
p.y_range,p.extra_y_ranges['Yaxis2'],p.extra_y_ranges['Yaxis3'],p.extra_y_ranges['Yaxis4'] = set_yranges_4y_ymin0(0,roundup10(df1.values.max()),
0,math.ceil(df2.values.max()),
0,roundup10(df3.values.max()),
0,roundup10(df4.values.max()),
step_y1,
step_y2,
step_y3,
step_y4)
#Set primary y-axis ticker:
ticker_1 = SingleIntervalTicker(interval=step_y1)
#Add primary y-axis ticker to plot:
p.yaxis.ticker = ticker_1
#Set secondary y-axis ticker:
ticker_2 = SingleIntervalTicker(interval=step_y2)
#Set 3rd y-axis ticker:
ticker_3 = SingleIntervalTicker(interval=step_y3)
#Set 4th y-axis ticker:
ticker_4 = SingleIntervalTicker(interval=step_y4)
# Adding the second axis to the plot.
yaxis2 = LinearAxis(y_range_name="Yaxis2",
axis_label=variable_ls[1] + ' ('+unit_ls[1].translate(SUP)+') daily '+dailyType_ls[1],
ticker=ticker_2,
axis_label_standoff = 15,
axis_label_text_color = color_ls[1])
# Adding the 3rd axis to the plot.
yaxis3 = LinearAxis(y_range_name='Yaxis3',
axis_label=variable_ls[2] + ' ('+unit_ls[2].translate(SUP)+') daily '+dailyType_ls[2],
ticker=ticker_3,
axis_label_standoff = 15,
axis_label_text_color = color_ls[2])
# Adding the 4th axis to the plot.
yaxis4 = LinearAxis(y_range_name='Yaxis4',
axis_label=variable_ls[3] + ' ('+unit_ls[3].translate(SUP)+') daily '+dailyType_ls[3],
ticker=ticker_4,
axis_label_standoff = 15,
axis_label_text_color = color_ls[3])
p.add_layout(yaxis2,'right')
p.add_layout(yaxis3,'right')
p.add_layout(yaxis4,'left')
#Create an empty list that will store the legend info:
legend_it = []
#Add SWC barplot:
bp1 = p.vbar(x=list(df1.index.values), width=0.5, bottom=0,
top=list(df1.values), color=color_ls[0], name=variable_ls[0])
#Add GPP barplot:
bp2 = p.vbar(x=list(df2.index.values), width=0.5, bottom=0, alpha=0.5,
top=list(df2.values), color=color_ls[1], y_range_name="Yaxis2",
name=variable_ls[1])
#Add Air-Temp line-glyph:
l1 = p.line(df3.index.values, df3.values,
line_width=2.0, color=color_ls[2],
y_range_name="Yaxis3", alpha=0.7, name=variable_ls[2])
#Add Light line-glyph:
l2 = p.line(list(df4.index.values), list(df4.values),
line_width=2.0, color=color_ls[3],
y_range_name="Yaxis4", alpha=0.7, name=variable_ls[3])
#Add the name and glyph info (i.e. colour and marker type) to the legend:
legend_it.append((bp1.name, [bp1]))
legend_it.append((bp2.name, [bp2]))
legend_it.append((l1.name, [l1]))
legend_it.append((l2.name, [l2]))
#Create legend:
legend = Legend(items=legend_it, location= 'bottom_center')
legend.orientation = 'horizontal'
legend.click_policy='hide'
legend.spacing = 10 #sets the distance between legend entries
#Add legend to figure:
p.add_layout(legend, 'below')
#Set title attributes:
p.title.align = 'center'
p.title.text_font_size = '12pt'
p.title.vertical_align = 'top' #Create a distance between the title and the plot
#Set axis label font style:
p.xaxis.axis_label_text_font_style = 'normal'
p.yaxis.axis_label_text_font_style = 'normal'
p.xaxis.axis_label_standoff = 15 #Sets the distance of the label from the x-axis in screen units
p.yaxis.axis_label_standoff = 15 #Sets the distance of the label from the y-axis in screen units
p.yaxis[0].axis_label_text_color = color_ls[0]
#Set the copyright-label position:
label_opts = dict(x=0, y=5,
x_units='screen', y_units='screen')
#Create a label object and format it:
caption1 = Label(text="© ICOS ERIC", **label_opts)
caption1.text_font_size = '8pt'
#Add label to plot:
p.add_layout(caption1, 'below')
#Format plot borders:
p.min_border_top = 54
#Return Figure Object:
return p
# -
# <br>
#
# <span style="color:blue">**Widget Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
def widgetSWCGPPTempSWIR():
#Import Python modules:
from ipywidgets import interact_manual, Dropdown
from bokeh.layouts import column
from bokeh.io import show, output_notebook
#Create a dictionary to store the filenames associated with each year:
labels = {"2015":[HTM_eko_SWC_2015_daily_mean,
micromoles2moles(HTM_eko_GPP_2015_daily_sum),
HTM_eko_TA_2015_daily_mean,
Joules2MegaJoules(HTM_eko_LIGHT_2015_daily_sum)],
"2016":[HTM_eko_SWC_2016_daily_mean,
micromoles2moles(HTM_eko_GPP_2016_daily_sum),
HTM_eko_TA_2016_daily_mean,
Joules2MegaJoules(HTM_eko_LIGHT_2016_daily_sum)],
"2017":[HTM_eko_SWC_2017_daily_mean,
micromoles2moles(HTM_eko_GPP_2017_daily_sum),
HTM_eko_TA_2017_daily_mean,
Joules2MegaJoules(HTM_eko_LIGHT_2017_daily_sum)]}
#Create Dropdown-List widget:
years = Dropdown(
options=labels.keys(),
value='2015',
description='Year:',
disabled=False)
#Function that calls functions to update the plot
#based on the selected year:
def update_plot_func(Year):
#Call function to plot data for the selected year:
p1 = plotGPPLightSWCTempYr_4axes(labels[Year][0],
labels[Year][1],
labels[Year][2],
labels[Year][3],
['Soil Water Content','GPP', 'Temperature', 'SW-IR'],
['%', 'moles m-2', 'C', 'MJoules m-2'],
['Mean', 'Total', 'Mean', 'Total'],
['lightblue', 'green', 'firebrick', 'gold'],
10.0, 0.5, 10.0, 10.0)
#Call function to plot data for 2018 (drought year):
p2 = plotGPPLightSWCTempYr_4axes(HTM_eko_SWC_2018_daily_mean,
micromoles2moles(HTM_eko_GPP_2018_daily_sum),
HTM_eko_TA_2018_daily_mean,
Joules2MegaJoules(HTM_eko_LIGHT_2018_daily_sum),
['Soil Water Content','GPP', 'Temperature', 'SW-IR'],
['%','moles m-2', 'C', 'MJoules m-2'],
['Mean', 'Total', 'Mean', 'Total'],
['lightblue', 'green', 'firebrick', 'gold'],
10.0,
0.5,
10.0,
10.0)
#Define output location:
output_notebook()
#Show plots:
show(column(p1, p2))
#Create function that contains a box of widgets:
interact_c = interact_manual(update_plot_func,
Year=years)
#Set the font of the widgets included in interact_manual:
interact_c.widget.children[0].layout.width = '430px'
interact_c.widget.children[0].layout.margin = '40px 2px 2px 2px'
interact_c.widget.children[1].description = 'Update Plot'
interact_c.widget.children[1].button_style = 'danger'
interact_c.widget.children[1].style.button_color = '#3973ac'
interact_c.widget.children[1].layout.margin = '10px 10px 40px 180px' # top/right/bottom/left
# -
# <a id='bokeh_plot_stat_summed_daily_totals_means_SWC_GPP_Temp_SWIR_per_year'></a>
# <br>
# #### Plot Daily Mean Soil Water Content and Air temperature with Daily Total GPP and Light (SW-IR)
# <br>
#
# <span style="color:green">**Call Function**</span>
# + deletable=false editable=false
################################
#Add button to hide/show code:
toggle_code()
################################
#Call function to display widgets:
widgetSWCGPPTempSWIR()
# -
# <a id='references'></a>
# <br>
# <br>
# <div style="text-align: right">
# <a href="#py_programming">Back to TOC</a>
# </div>
# <br>
# <br>
# <br>
#
# # References
# 1. <NAME>. et.al. (2019). Seasonal hydro-ecological feedbacks during the 2018 drought in Europe. Unpublished manuscript.
#
#
# 2. ”Torka | SMHI” (in SE). (2019-05-29). Retrieved from https://www.smhi.se/kunskapsbanken/hydrologi/torka-1.111075.
#
#
# 3. <NAME>. (2015-04-14). ”Hydrological drought explained”. Wiley Interdisciplinary Reviews: Water 2 (4). pp. 359–392. doi:10.1002/wat2.1085. ISSN 2049-1948. Accessed in March 29th 2019.
#
#
# 4. <NAME>., <NAME>. (2019-04-19). Drought Damage to Trees [PDF file]. Retrieved from https://www.kansasforests.org/forest_health/health_docs/DroughtDamageToTrees.pdf
#
#
# 5. <NAME>. (2019-04-20). Skadebeskrivning - Sommartorka (in SE). Retrieved from https://www.slu.se/centrumbildningar-och-projekt/skogsskada/lasmer-sidor/skadeorsak/?DiagID=63&AnmSkada=63&Tradart=16&Skadetyp=1&Alder=2&SkadadDel=0,7&SkadaBestand=1
#
#
# 6. <NAME>. (2019-05-10). Nationalencyklopedin, destruent (in SE). Retrieved from http://www.ne.se/uppslagsverk/encyklopedi/lång/destruent
#
#
# 7. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2018-08-30). Sensitivity of atmospheric CO2 growth rate to observed changes in terrestrial water storage. Nature, 560(7720), 628–631. doi: 10.1038/s41586-018-0424-4
#
#
# 8. <NAME>. and ICOS Ecosystem Thematic Centre: Drought-2018 ecosystem eddy covariance flux product from Hyltemossa, , doi:10.18160/17FF-96RT, 2020.
# <br>
# <br>
# <div style="text-align: right">
# <a href="#toc">Back to top</a>
# </div>
| notebooks/education/PhD/upscaling_carbon_fluxes/notebooks/htm_eco_drought_2018.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bayesian Regression - Inference Algorithms (Part 2)
# In [Part I](bayesian_regression.ipynb), we looked at how to perform inference on a simple Bayesian linear regression model using SVI. In this tutorial, we'll explore more expressive guides as well as exact inference techniques. We'll use the same dataset as before.
# %reset -sf
# +
import logging
import os
import torch
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from torch.distributions import constraints
import pyro
import pyro.distributions as dist
import pyro.optim as optim
pyro.set_rng_seed(1)
assert pyro.__version__.startswith('1.5.1')
# +
# %matplotlib inline
plt.style.use('default')
logging.basicConfig(format='%(message)s', level=logging.INFO)
smoke_test = ('CI' in os.environ)
pyro.set_rng_seed(1)
DATA_URL = "https://d2hg8soec8ck9v.cloudfront.net/datasets/rugged_data.csv"
rugged_data = pd.read_csv(DATA_URL, encoding="ISO-8859-1")
# -
# ## Bayesian Linear Regression
#
# Our goal is once again to predict log GDP per capita of a nation as a function of two features from the dataset - whether the nation is in Africa, and its Terrain Ruggedness Index, but we will explore more expressive guides.
# ## Model + Guide
#
# We will write out the model again, similar to that in [Part I](bayesian_regression.ipynb), but explicitly without the use of `PyroModule`. We will write out each term in the regression, using the same priors. `bA` and `bR` are regression coefficients corresponding to `is_cont_africa` and `ruggedness`, `a` is the intercept, and `bAR` is the correlating factor between the two features.
#
# Writing down a guide will proceed in close analogy to the construction of our model, with the key difference that the guide parameters need to be trainable. To do this we register the guide parameters in the ParamStore using `pyro.param()`. Note the positive constraints on scale parameters.
# +
def model(is_cont_africa, ruggedness, log_gdp):
a = pyro.sample("a", dist.Normal(0., 10.))
b_a = pyro.sample("bA", dist.Normal(0., 1.))
b_r = pyro.sample("bR", dist.Normal(0., 1.))
b_ar = pyro.sample("bAR", dist.Normal(0., 1.))
sigma = pyro.sample("sigma", dist.Uniform(0., 10.))
mean = a + b_a * is_cont_africa + b_r * ruggedness + b_ar * is_cont_africa * ruggedness
with pyro.plate("data", len(ruggedness)):
pyro.sample("obs", dist.Normal(mean, sigma), obs=log_gdp)
def guide(is_cont_africa, ruggedness, log_gdp):
a_loc = pyro.param('a_loc', torch.tensor(0.))
a_scale = pyro.param('a_scale', torch.tensor(1.),
constraint=constraints.positive)
sigma_loc = pyro.param('sigma_loc', torch.tensor(1.),
constraint=constraints.positive)
weights_loc = pyro.param('weights_loc', torch.randn(3))
weights_scale = pyro.param('weights_scale', torch.ones(3),
constraint=constraints.positive)
a = pyro.sample("a", dist.Normal(a_loc, a_scale))
b_a = pyro.sample("bA", dist.Normal(weights_loc[0], weights_scale[0]))
b_r = pyro.sample("bR", dist.Normal(weights_loc[1], weights_scale[1]))
b_ar = pyro.sample("bAR", dist.Normal(weights_loc[2], weights_scale[2]))
sigma = pyro.sample("sigma", dist.Normal(sigma_loc, torch.tensor(0.05)))
mean = a + b_a * is_cont_africa + b_r * ruggedness + b_ar * is_cont_africa * ruggedness
# +
# Utility function to print latent sites' quantile information.
def summary(samples):
site_stats = {}
for site_name, values in samples.items():
marginal_site = pd.DataFrame(values)
describe = marginal_site.describe(percentiles=[.05, 0.25, 0.5, 0.75, 0.95]).transpose()
site_stats[site_name] = describe[["mean", "std", "5%", "25%", "50%", "75%", "95%"]]
return site_stats
# Prepare training data
df = rugged_data[["cont_africa", "rugged", "rgdppc_2000"]]
df = df[np.isfinite(df.rgdppc_2000)]
df["rgdppc_2000"] = np.log(df["rgdppc_2000"])
train = torch.tensor(df.values, dtype=torch.float)
# -
# ## SVI
#
# As before, we will use SVI to perform inference.
# +
from pyro.infer import SVI, Trace_ELBO
svi = SVI(model,
guide,
optim.Adam({"lr": .05}),
loss=Trace_ELBO())
is_cont_africa, ruggedness, log_gdp = train[:, 0], train[:, 1], train[:, 2]
pyro.clear_param_store()
num_iters = 5000 if not smoke_test else 2
for i in range(num_iters):
elbo = svi.step(is_cont_africa, ruggedness, log_gdp)
if i % 500 == 0:
logging.info("Elbo loss: {}".format(elbo))
# +
from pyro.infer import Predictive
num_samples = 1000
predictive = Predictive(model, guide=guide, num_samples=num_samples)
svi_samples = {k: v.reshape(num_samples).detach().cpu().numpy()
for k, v in predictive(log_gdp, is_cont_africa, ruggedness).items()
if k != "obs"}
# -
# Let us observe the posterior distribution over the different latent variables in the model.
for site, values in summary(svi_samples).items():
print("Site: {}".format(site))
print(values, "\n")
# ## HMC
#
# In contrast to using variational inference which gives us an approximate posterior over our latent variables, we can also do exact inference using [Markov Chain Monte Carlo](http://docs.pyro.ai/en/dev/mcmc.html) (MCMC), a class of algorithms that in the limit, allow us to draw unbiased samples from the true posterior. The algorithm that we will be using is called the No-U Turn Sampler (NUTS) \[1\], which provides an efficient and automated way of running Hamiltonian Monte Carlo. It is slightly slower than variational inference, but provides an exact estimate.
# +
from pyro.infer import MCMC, NUTS
nuts_kernel = NUTS(model)
mcmc = MCMC(nuts_kernel, num_samples=1000, warmup_steps=200)
mcmc.run(is_cont_africa, ruggedness, log_gdp)
hmc_samples = {k: v.detach().cpu().numpy() for k, v in mcmc.get_samples().items()}
# -
for site, values in summary(hmc_samples).items():
print("Site: {}".format(site))
print(values, "\n")
# ## Comparing Posterior Distributions
#
# Let us compare the posterior distribution of the latent variables that we obtained from variational inference with those from Hamiltonian Monte Carlo. As can be seen below, for Variational Inference, the marginal distribution of the different regression coefficients is under-dispersed w.r.t. the true posterior (from HMC). This is an artifact of the *KL(q||p)* loss (the KL divergence of the true posterior from the approximate posterior) that is minimized by Variational Inference.
#
# This can be better seen when we plot different cross sections from the joint posterior distribution overlaid with the approximate posterior from variational inference. Note that since our variational family has diagonal covariance, we cannot model any correlation between the latents and the resulting approximation is overconfident (under-dispersed)
# +
sites = ["a", "bA", "bR", "bAR", "sigma"]
fig, axs = plt.subplots(nrows=2, ncols=2, figsize=(12, 10))
fig.suptitle("Marginal Posterior density - Regression Coefficients", fontsize=16)
for i, ax in enumerate(axs.reshape(-1)):
site = sites[i]
sns.distplot(svi_samples[site], ax=ax, label="SVI (DiagNormal)")
sns.distplot(hmc_samples[site], ax=ax, label="HMC")
ax.set_title(site)
handles, labels = ax.get_legend_handles_labels()
fig.legend(handles, labels, loc='upper right');
# -
fig, axs = plt.subplots(nrows=1, ncols=2, figsize=(12, 6))
fig.suptitle("Cross-section of the Posterior Distribution", fontsize=16)
sns.kdeplot(hmc_samples["bA"], hmc_samples["bR"], ax=axs[0], shade=True, label="HMC")
sns.kdeplot(svi_samples["bA"], svi_samples["bR"], ax=axs[0], label="SVI (DiagNormal)")
axs[0].set(xlabel="bA", ylabel="bR", xlim=(-2.5, -1.2), ylim=(-0.5, 0.1))
sns.kdeplot(hmc_samples["bR"], hmc_samples["bAR"], ax=axs[1], shade=True, label="HMC")
sns.kdeplot(svi_samples["bR"], svi_samples["bAR"], ax=axs[1], label="SVI (DiagNormal)")
axs[1].set(xlabel="bR", ylabel="bAR", xlim=(-0.45, 0.05), ylim=(-0.15, 0.8))
handles, labels = axs[1].get_legend_handles_labels()
fig.legend(handles, labels, loc='upper right');
# ## MultivariateNormal Guide
#
# As comparison to the previously obtained results from Diagonal Normal guide, we will now use a guide that generates samples from a Cholesky factorization of a multivariate normal distribution. This allows us to capture the correlations between the latent variables via a covariance matrix. If we wrote this manually, we would need to combine all the latent variables so we could sample a Multivarite Normal jointly.
# +
from pyro.infer.autoguide import AutoMultivariateNormal, init_to_mean
guide = AutoMultivariateNormal(model, init_loc_fn=init_to_mean)
svi = SVI(model,
guide,
optim.Adam({"lr": .01}),
loss=Trace_ELBO())
is_cont_africa, ruggedness, log_gdp = train[:, 0], train[:, 1], train[:, 2]
pyro.clear_param_store()
for i in range(num_iters):
elbo = svi.step(is_cont_africa, ruggedness, log_gdp)
if i % 500 == 0:
logging.info("Elbo loss: {}".format(elbo))
# -
# Let's look at the shape of the posteriors again. You can see the multivariate guide is able to capture more of the true posterior.
predictive = Predictive(model, guide=guide, num_samples=num_samples)
svi_mvn_samples = {k: v.reshape(num_samples).detach().cpu().numpy()
for k, v in predictive(log_gdp, is_cont_africa, ruggedness).items()
if k != "obs"}
fig, axs = plt.subplots(nrows=2, ncols=2, figsize=(12, 10))
fig.suptitle("Marginal Posterior density - Regression Coefficients", fontsize=16)
for i, ax in enumerate(axs.reshape(-1)):
site = sites[i]
sns.distplot(svi_mvn_samples[site], ax=ax, label="SVI (Multivariate Normal)")
sns.distplot(hmc_samples[site], ax=ax, label="HMC")
ax.set_title(site)
handles, labels = ax.get_legend_handles_labels()
fig.legend(handles, labels, loc='upper right');
# Now let's compare the posterior computed by the Diagonal Normal guide vs the Multivariate Normal guide. Note that the multivariate distribution is more dispresed than the Diagonal Normal.
fig, axs = plt.subplots(nrows=1, ncols=2, figsize=(12, 6))
fig.suptitle("Cross-sections of the Posterior Distribution", fontsize=16)
sns.kdeplot(svi_samples["bA"], svi_samples["bR"], ax=axs[0], label="SVI (Diagonal Normal)")
sns.kdeplot(svi_mvn_samples["bA"], svi_mvn_samples["bR"], ax=axs[0], shade=True, label="SVI (Multivariate Normal)")
axs[0].set(xlabel="bA", ylabel="bR", xlim=(-2.5, -1.2), ylim=(-0.5, 0.1))
sns.kdeplot(svi_samples["bR"], svi_samples["bAR"], ax=axs[1], label="SVI (Diagonal Normal)")
sns.kdeplot(svi_mvn_samples["bR"], svi_mvn_samples["bAR"], ax=axs[1], shade=True, label="SVI (Multivariate Normal)")
axs[1].set(xlabel="bR", ylabel="bAR", xlim=(-0.45, 0.05), ylim=(-0.15, 0.8))
handles, labels = axs[1].get_legend_handles_labels()
fig.legend(handles, labels, loc='upper right');
# and the Multivariate guide with the posterior computed by HMC. Note that the Multivariate guide better captures the true posterior.
fig, axs = plt.subplots(nrows=1, ncols=2, figsize=(12, 6))
fig.suptitle("Cross-sections of the Posterior Distribution", fontsize=16)
sns.kdeplot(hmc_samples["bA"], hmc_samples["bR"], ax=axs[0], shade=True, label="HMC")
sns.kdeplot(svi_mvn_samples["bA"], svi_mvn_samples["bR"], ax=axs[0], label="SVI (Multivariate Normal)")
axs[0].set(xlabel="bA", ylabel="bR", xlim=(-2.5, -1.2), ylim=(-0.5, 0.1))
sns.kdeplot(hmc_samples["bR"], hmc_samples["bAR"], ax=axs[1], shade=True, label="HMC")
sns.kdeplot(svi_mvn_samples["bR"], svi_mvn_samples["bAR"], ax=axs[1], label="SVI (Multivariate Normal)")
axs[1].set(xlabel="bR", ylabel="bAR", xlim=(-0.45, 0.05), ylim=(-0.15, 0.8))
handles, labels = axs[1].get_legend_handles_labels()
fig.legend(handles, labels, loc='upper right');
# ## References
# [1] Hoffman, <NAME>., and <NAME>. "The No-U-turn sampler: adaptively setting path lengths in Hamiltonian Monte Carlo." Journal of Machine Learning Research 15.1 (2014): 1593-1623. https://arxiv.org/abs/1111.4246.
| tutorial/source/bayesian_regression_ii.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
# *This notebook contains an excerpt from the [Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/WhirlwindTourOfPython).*
#
# *The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# <!--NAVIGATION-->
# < [Generators](12-Generators.ipynb) | [Contents](Index.ipynb) | [String Manipulation and Regular Expressions](14-Strings-and-Regular-Expressions.ipynb) >
# # Modules and Packages
# One feature of Python that makes it useful for a wide range of tasks is the fact that it comes "batteries included" – that is, the Python standard library contains useful tools for a wide range of tasks.
# On top of this, there is a broad ecosystem of third-party tools and packages that offer more specialized functionality.
# Here we'll take a look at importing standard library modules, tools for installing third-party modules, and a description of how you can make your own modules.
# ## Loading Modules: the ``import`` Statement
#
# For loading built-in and third-party modules, Python provides the ``import`` statement.
# There are a few ways to use the statement, which we will mention briefly here, from most recommended to least recommended.
# ### Explicit module import
#
# Explicit import of a module preserves the module's content in a namespace.
# The namespace is then used to refer to its contents with a "``.``" between them.
# For example, here we'll import the built-in ``math`` module and compute the cosine of pi:
import math
math.cos(math.pi)
# ### Explicit module import by alias
#
# For longer module names, it's not convenient to use the full module name each time you access some element.
# For this reason, we'll commonly use the "``import ... as ...``" pattern to create a shorter alias for the namespace.
# For example, the NumPy (Numerical Python) package, a popular third-party package useful for data science, is by convention imported under the alias ``np``:
import numpy as np
np.cos(np.pi)
# ### Explicit import of module contents
#
# Sometimes rather than importing the module namespace, you would just like to import a few particular items from the module.
# This can be done with the "``from ... import ...``" pattern.
# For example, we can import just the ``cos`` function and the ``pi`` constant from the ``math`` module:
from math import cos, pi
cos(pi)
# ### Implicit import of module contents
#
# Finally, it is sometimes useful to import the entirety of the module contents into the local namespace.
# This can be done with the "``from ... import *``" pattern:
from math import *
sin(pi) ** 2 + cos(pi) ** 2
# This pattern should be used sparingly, if at all.
# The problem is that such imports can sometimes overwrite function names that you do not intend to overwrite, and the implicitness of the statement makes it difficult to determine what has changed.
#
# For example, Python has a built-in ``sum`` function that can be used for various operations:
help(sum)
# We can use this to compute the sum of a sequence, starting with a certain value (here, we'll start with ``-1``):
sum(range(5), -1)
# Now observe what happens if we make the *exact same function call* after importing ``*`` from ``numpy``:
from numpy import *
sum(range(5), -1)
# The result is off by one!
# The reason for this is that the ``import *`` statement *replaces* the built-in ``sum`` function with the ``numpy.sum`` function, which has a different call signature: in the former, we're summing ``range(5)`` starting at ``-1``; in the latter, we're summing ``range(5)`` along the last axis (indicated by ``-1``).
# This is the type of situation that may arise if care is not taken when using "``import *``" – for this reason, it is best to avoid this unless you know exactly what you are doing.
# ## Importing from Python's Standard Library
#
# Python's standard library contains many useful built-in modules, which you can read about fully in [Python's documentation](https://docs.python.org/3/library/).
# Any of these can be imported with the ``import`` statement, and then explored using the help function seen in the previous section.
# Here is an extremely incomplete list of some of the modules you might wish to explore and learn about:
#
# - ``os`` and ``sys``: Tools for interfacing with the operating system, including navigating file directory structures and executing shell commands
# - ``math`` and ``cmath``: Mathematical functions and operations on real and complex numbers
# - ``itertools``: Tools for constructing and interacting with iterators and generators
# - ``functools``: Tools that assist with functional programming
# - ``random``: Tools for generating pseudorandom numbers
# - ``pickle``: Tools for object persistence: saving objects to and loading objects from disk
# - ``json`` and ``csv``: Tools for reading JSON-formatted and CSV-formatted files.
# - ``urllib``: Tools for doing HTTP and other web requests.
#
# You can find information on these, and many more, in the Python standard library documentation: https://docs.python.org/3/library/.
# ## Importing from Third-Party Modules
#
# One of the things that makes Python useful, especially within the world of data science, is its ecosystem of third-party modules.
# These can be imported just as the built-in modules, but first the modules must be installed on your system. To install common third-party packages one can use one of the following approaches:
#
# ### Conda
# Conda is an open source package management system and environment management system that runs on Windows, macOS and Linux. Conda quickly installs, runs and updates packages and their dependencies.
# For example, if you'd like to install the ``numpy`` package, all that is required is to type the following at the command line:
# ```
# $ conda install numpy
# ```
#
# ### pip
# Another standard registry for such modules is the Python Package Index (*PyPI* for short), found on the Web at http://pypi.python.org/.
# For convenience, Python comes with a program called ``pip`` (a recursive acronym meaning "pip installs packages"), which will automatically fetch packages released and listed on PyPI (if you use Python version 2, ``pip`` must be installed separately).
# For example, if you'd like to install the ``numpy`` package, all that is required is to type the following at the command line:
# ```
# $ pip install numpy
# ```
# The source code for the package will be automatically downloaded from the PyPI repository, and the package installed in the standard Python path (assuming you have permission to do so on the computer you're using).
#
# For more information about PyPI and the ``pip`` installer, refer to the documentation at http://pypi.python.org/.
# <!--NAVIGATION-->
# < [Generators](12-Generators.ipynb) | [Contents](Index.ipynb) | [String Manipulation and Regular Expressions](14-Strings-and-Regular-Expressions.ipynb) >
| Lectures/14-Modules-and-Packages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> 5. Interactive Data Analysis </h1>
#
# This notebook introduces carrying out interactive data analysis of data in BigQuery using Cloud Datalab.
#
# This cell, for example, is a mark-down cell. Which is why you are seeing text. The cell that follows is a Python code cell. The output of that cell is whatever is printed out from it.
a = 3
b = a + 5
print("a={} b={}".format(a,b))
# Because the markdown comments are HTML, it can even include Javascript, for example to create a table of contents, as shown below:
#
# <div id="toc"></div>
# <h2> Relative path </h2>
#
# I created this notebook in 05_bqnotebook folder of the git repo for the book. So, you might see a path that ends in that. But the path will start with /home/jupyter which is mapped to a local folder if you are running this in a container.
# !pwd
# <h2> What's installed? </h2>
# %pip freeze
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
# <h2> Installing dependencies </h2>
#
# Regular Python dependencies can be installed using pip
# %pip install google-cloud
# + language="bash"
# sudo apt-get update
# sudo apt-get -y install python-mpltoolkits.basemap
# -
# <h2> Juypter magic </h2>
# + language="html"
# This cell will print out a <b> HTML </b> string.
# -
# %%bigquery
SELECT
COUNTIF(arr_delay >= 15)/COUNT(arr_delay) AS frac_delayed
FROM flights.tzcorr
# <h2> Calls to BigQuery </h2>
#
# Let's draw a PDF of different arrival delays
# %%bigquery df
SELECT ARR_DELAY, DEP_DELAY
FROM flights.tzcorr
WHERE DEP_DELAY >= 10 AND RAND() < 0.01
df.describe()
sns.set_style("whitegrid")
ax = sns.violinplot(data=df, x='ARR_DELAY', inner='box', orient='h')
#ax.axes.set_xlim(0, 30)
# %%bigquery df
SELECT ARR_DELAY, DEP_DELAY
FROM flights.tzcorr
WHERE RAND() < 0.001
df.describe()
df['ontime'] = df['DEP_DELAY'] < 10
df[df['ARR_DELAY'] > 0].head()
import seaborn as sns
sns.set_style("whitegrid")
ax = sns.violinplot(data=df, x='ARR_DELAY', y='ontime', inner='box', orient='h')
ax.set_xlim(-50, 200)
ax = sns.violinplot(data=df, x='ARR_DELAY', y='ontime',
inner='box', orient='h', gridsize=1000)
ax.set_xlim(-50, 50)
# %%bigquery depdelay
SELECT
*
FROM (
SELECT
DEP_DELAY,
AVG(ARR_DELAY) AS arrival_delay,
STDDEV(ARR_DELAY) AS stddev_arrival_delay,
COUNT(ARR_DELAY) AS numflights
FROM
`flights.tzcorr`
GROUP BY
DEP_DELAY )
WHERE
numflights > 370
ORDER BY
DEP_DELAY
depdelay[:5]
ax = depdelay.plot(kind='line', x='DEP_DELAY',
y='arrival_delay', yerr='stddev_arrival_delay')
import matplotlib.pyplot as plt
Z_30 = 0.52
depdelay['arr_delay_30'] = (Z_30 * depdelay['stddev_arrival_delay']) \
+ depdelay['arrival_delay']
plt.axhline(y=15, color='r')
ax = plt.axes()
depdelay.plot(kind='line', x='DEP_DELAY', y='arr_delay_30',
ax=ax, ylim=(0,30), xlim=(0,30), legend=False)
ax.set_xlabel('Departure Delay (minutes)')
ax.set_ylabel('> 30% likelihood of this Arrival Delay (minutes)');
# %%bigquery depdelay
SELECT
DEP_DELAY,
arrival_delay,
numflights
FROM (
SELECT
DEP_DELAY,
APPROX_QUANTILES(ARR_DELAY,
101)[OFFSET(70)] AS arrival_delay,
COUNT(ARR_DELAY) AS numflights
FROM
`flights.tzcorr`
GROUP BY
DEP_DELAY )
WHERE
numflights > 370
ORDER BY
DEP_DELAY
plt.axhline(y=15, color='r')
ax = plt.axes()
depdelay.plot(kind='line', x='DEP_DELAY', y='arrival_delay',
ax=ax, ylim=(0,30), xlim=(0,30), legend=False)
ax.set_xlabel('Departure Delay (minutes)')
ax.set_ylabel('> 30% likelihood of this Arrival Delay (minutes)');
# %%bigquery depdelay
SELECT
DEP_DELAY,
arrival_delay,
numflights
FROM (
SELECT
DEP_DELAY,
APPROX_QUANTILES(ARR_DELAY,
101)[OFFSET(70)] AS arrival_delay,
COUNT(ARR_DELAY) AS numflights
FROM
`flights.tzcorr` f
JOIN
`flights.trainday` t
ON
f.FL_DATE = t.FL_DATE
WHERE
t.is_train_day = 'True'
GROUP BY
DEP_DELAY )
WHERE
numflights > 370
ORDER BY
DEP_DELAY
plt.axhline(y=15, color='r')
ax = plt.axes()
depdelay.plot(kind='line', x='DEP_DELAY', y='arrival_delay',
ax=ax, ylim=(0,30), xlim=(0,30), legend=False)
ax.set_xlabel('Departure Delay (minutes)')
ax.set_ylabel('> 30% likelihood of this Arrival Delay (minutes)');
# %%bigquery eval
SELECT
SUM(IF(DEP_DELAY < 16
AND arr_delay < 15, 1, 0)) AS correct_nocancel,
SUM(IF(DEP_DELAY < 16
AND arr_delay >= 15, 1, 0)) AS wrong_nocancel,
SUM(IF(DEP_DELAY >= 16
AND arr_delay < 15, 1, 0)) AS wrong_cancel,
SUM(IF(DEP_DELAY >= 16
AND arr_delay >= 15, 1, 0)) AS correct_cancel
FROM (
SELECT
DEP_DELAY,
ARR_DELAY
FROM
`flights.tzcorr` f
JOIN
`flights.trainday` t
ON
f.FL_DATE = t.FL_DATE
WHERE
t.is_train_day = 'False' )
print(eval['correct_nocancel'] /
(eval['correct_nocancel'] + eval['wrong_nocancel']))
print(eval['correct_cancel'] /
(eval['correct_cancel'] + eval['wrong_cancel']))
eval.head()
# %%bigquery eval
SELECT
SUM(IF(DEP_DELAY = 15
AND arr_delay < 15, 1, 0)) AS correct_nocancel,
SUM(IF(DEP_DELAY = 15
AND arr_delay >= 15, 1, 0)) AS wrong_nocancel,
SUM(IF(DEP_DELAY = 16
AND arr_delay < 15, 1, 0)) AS wrong_cancel,
SUM(IF(DEP_DELAY = 16
AND arr_delay >= 15, 1, 0)) AS correct_cancel
FROM (
SELECT
DEP_DELAY,
ARR_DELAY
FROM
`flights.tzcorr` f
JOIN
`flights.trainday` t
ON
f.FL_DATE = t.FL_DATE
WHERE
t.is_train_day = 'False' )
eval.head()
print(eval['correct_nocancel'] / (eval['correct_nocancel'] + eval['wrong_nocancel']))
print(eval['correct_cancel'] / (eval['correct_cancel'] + eval['wrong_cancel']))
# ## Sending to a temporary table
# !bq mk temp_dataset
# +
# %%bigquery
CREATE OR REPLACE TABLE temp_dataset.delays
AS
SELECT
DEP_DELAY,
arrival_delay,
numflights
FROM (
SELECT
DEP_DELAY,
APPROX_QUANTILES(ARR_DELAY,
101)[OFFSET(70)] AS arrival_delay,
COUNT(ARR_DELAY) AS numflights
FROM
`flights.tzcorr` f
JOIN
`flights.trainday` t
ON
f.FL_DATE = t.FL_DATE
WHERE
t.is_train_day = 'True'
GROUP BY
DEP_DELAY )
WHERE
numflights > 370
ORDER BY
DEP_DELAY
# -
# !bq rm -f temp_dataset.delays
# !bq rm -f temp_dataset
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| quests/data-science-on-gcp-edition1_tf2/05_bqnotebook/exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Planet Data Collection
# Using the Open Exoplanet Catalogue database: https://github.com/OpenExoplanetCatalogue/open_exoplanet_catalogue/
#
# ## Data License
# Copyright (C) 2012 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this database and associated scripts (the "Database"), to deal in the Database without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Database, and to permit persons to whom the Database is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Database. A reference to the Database shall be included in all scientific publications that make use of the Database.
#
# THE DATABASE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE DATABASE OR THE USE OR OTHER DEALINGS IN THE DATABASE.
# ## Follow instructions to get the xml file
import xml.etree.ElementTree as ET, urllib.request, gzip, io
url = "https://github.com/OpenExoplanetCatalogue/oec_gzip/raw/master/systems.xml.gz"
oec = ET.parse(gzip.GzipFile(fileobj=io.BytesIO(urllib.request.urlopen(url).read())))
# ## Parse into Pandas DataFrame
# Information on what each field means can be found [here](https://github.com/OpenExoplanetCatalogue/open_exoplanet_catalogue/#data-structure).
# +
import pandas as pd
def parse(base):
db = oec.findall(f".//{base}")
exclude = ['star', 'videolink', 'binary'] if base in ['system', 'binary'] else ['planet']
columns = set([attribute.tag for attribute in db[0].getchildren() if attribute.tag not in exclude])
results = pd.DataFrame(columns=columns)
for entry in db:
data = {col : entry.findtext(col) for col in columns}
if base in ['system', 'binary']:
data['binaries'] = len(entry.findall('.//binary'))
data['stars'] = len(entry.findall('.//star'))
if base in ['system', 'star', 'binary']:
data['planets'] = len(entry.findall('.//planet'))
results = results.append(data, ignore_index=True)
return results
# -
# ### Parse planet data
planets = parse('planet')
planets.head()
# ### Parse system data
systems = parse('system')
systems.head()
# ### Parse binary data
binaries = parse('binary')
binaries.head()
# ### Parse star data
stars = parse('star')
stars.head()
# ## Save to CSVs
planets.to_csv('data/planets.csv', index=False)
binaries.to_csv('data/binaries.csv', index=False)
stars.to_csv('data/stars.csv', index=False)
systems.to_csv('data/systems.csv', index=False)
| ch_09/planet_data_collection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="QyFtuz0qFYF1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 183} outputId="4947abc7-3847-43ed-c20c-a106aea25103" executionInfo={"status": "ok", "timestamp": 1583318067611, "user_tz": -60, "elapsed": 2426, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
# + id="MjCWLc_gGFSB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="58803624-00c7-4b33-ae8d-a444691af4f8" executionInfo={"status": "ok", "timestamp": 1583318074686, "user_tz": -60, "elapsed": 517, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_car"
# + [markdown] id="ujLiCT1JGjZN" colab_type="text"
# ## Wczytywanie danych
# + id="OzIHATtLGlnW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a434888a-33cb-43d3-be81-e5fcf0dfcbfe" executionInfo={"status": "ok", "timestamp": 1583318089936, "user_tz": -60, "elapsed": 2297, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="n69wPZmSG1_Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="41b5861d-0353-462d-97dd-77b66f07ee36" executionInfo={"status": "ok", "timestamp": 1583318105195, "user_tz": -60, "elapsed": 664, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df.columns
# + id="OmlAlWnqHOMH" colab_type="code" colab={}
# + [markdown] id="ukSocZ7JHli3" colab_type="text"
# ## Dummy model
# + id="XDlTZEfXHngh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="16f684f6-f907-443c-bb49-7bc863f74891" executionInfo={"status": "ok", "timestamp": 1583318223581, "user_tz": -60, "elapsed": 687, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df.select_dtypes(np.number).columns
# + id="cdwTpmgpHsbw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0e49d769-dce2-4da5-86f6-e1d74325bbd9" executionInfo={"status": "ok", "timestamp": 1583318555281, "user_tz": -60, "elapsed": 631, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
X = df[['car_id']].values
y = df['price_value']. values
model = DummyRegressor()
model.fit(X, y)
y_pred = model.predict(X)
mae(y, y_pred)
# + id="Zxx-bpL5I9fD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="19821037-3c1b-4c84-a077-59d351aa6711" executionInfo={"status": "ok", "timestamp": 1583318684042, "user_tz": -60, "elapsed": 804, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
[x for x in df.columns if 'price' in x]
# + id="buK7omVAJc3t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="5d517227-7b1d-466c-ae8c-06f3b1e40ec8" executionInfo={"status": "ok", "timestamp": 1583318709280, "user_tz": -60, "elapsed": 819, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df['price_currency']. value_counts()
# + id="JZAX01mAJi-z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="a561afa3-0912-4904-e0b7-0d3ef70bff1f" executionInfo={"status": "ok", "timestamp": 1583318760970, "user_tz": -60, "elapsed": 578, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df['price_currency']. value_counts(normalize=True) * 100
# + id="08pNRgm2Jvt7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8acce355-cb9f-47ca-e0d7-80efb5e735f6" executionInfo={"status": "ok", "timestamp": 1583318848651, "user_tz": -60, "elapsed": 865, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df = df[ df['price_currency'] != 'EUR' ]
df.shape
# + id="elwXygvhKFDk" colab_type="code" colab={}
# + [markdown] id="rNWfL8EDKNMp" colab_type="text"
# ## Features
# + id="HgStizwwKO0P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 664} outputId="748e357d-4e49-4305-da22-21c2695d3c68" executionInfo={"status": "ok", "timestamp": 1583318895831, "user_tz": -60, "elapsed": 683, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df.head()
# + id="uAQj9dAgKbHi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="323283e2-7571-407a-b919-faf923918be4" executionInfo={"status": "ok", "timestamp": 1583318973828, "user_tz": -60, "elapsed": 598, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df['param_color'].factorize()
# + id="EKiWAowkKjpY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4ccd70e3-5f2e-4f05-f594-086aba6d4f69" executionInfo={"status": "ok", "timestamp": 1583319017485, "user_tz": -60, "elapsed": 752, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
df['param_color'].factorize()[0]
# + id="AcO7QpLjKQcO" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = df[feat].factorize()[0]
# + id="PlbR6NzSKuTZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="771585f6-7f59-4cba-a356-3525664fc52e" executionInfo={"status": "ok", "timestamp": 1583319668021, "user_tz": -60, "elapsed": 524, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="-5hZjMwLMrIY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e05515e3-729e-424e-d36c-852aa57fceee" executionInfo={"status": "ok", "timestamp": 1583319803514, "user_tz": -60, "elapsed": 4719, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
X = df[cat_feats].values
y = df['price_value'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
np.mean(scores)
# + id="TuhKg4HWNtOo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 386} outputId="c1be2605-219f-4458-da08-8c3c776b065c" executionInfo={"status": "ok", "timestamp": 1583320026247, "user_tz": -60, "elapsed": 45460, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
m = DecisionTreeRegressor(max_depth=5)
m.fit(X, y)
imp = PermutationImportance(m, random_state=0).fit(X,y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="QHY02PuHOOcZ" colab_type="code" colab={}
# !git add day3_simple_model.ipynb
# + id="0mkIh1qJPPck" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 252} outputId="05e8a4ed-876a-4ffa-dcc8-11223a1d01a6" executionInfo={"status": "ok", "timestamp": 1583320213702, "user_tz": -60, "elapsed": 7217, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
# !git status
# + id="k0AAsBpHO8lz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="b13dfa5d-1880-49f2-de46-07511088e25c" executionInfo={"status": "ok", "timestamp": 1583320286066, "user_tz": -60, "elapsed": 4221, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
# !git commit -m 'add dummy model'
# + id="aSwWVONkPhll" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="45997ea4-b402-44f7-a700-2e90112f6e72" executionInfo={"status": "ok", "timestamp": 1583320305783, "user_tz": -60, "elapsed": 6414, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07362840361305936900"}}
# !git push -u origin master
# + id="dd1e53piPnce" colab_type="code" colab={}
| day3_simple_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IST 652 Lab #6
# ### Instructions
# - Complete all 6 questions in this assignment.
# - You may work with others, <b> but the work you submit must be your own </b>. You can differentiate your work by adding comments or changing the values you use to test your code. However, submitting some else's work as your own is an academic integrity violation and will be raised to academic affairs.
# - It is always better to attempt a problem as partial credit may be granted.
#
#
# ### Submission Guide:
# - Submit your answers on BlackBoard by Thursday 2019-04-18.
# - The file must be either a .py or .ipynb file type.
# - <i><span style="color:red">The name of the file you submit should be <i><b> ist652_lab6_lastname.py (.ipynb) </i></b>.</span>
#
#
#
# ### Grading [ 6 total points ]
# For Each Questions (1-6), the following credit will be awarded:
# - 0.75 for printing the correct answer to the console.
# - 0.15 for approaching the problem efficiently.
# - 0.05 for properly documenting and commenting your code.
#
# ---
# ## Questions
# ---
# #### First, download the Watson_Tweets.csv from BlackBoard.
# #### ( 1 ) Load the Watson_Tweets.csv file using pandas pd.read_csv() method and describe the data set.
# - NOTE: The file is pipe (|) seperated.
# - use the method parameter `sep` to properly parse and read the pipe-delimeted file
#
# ##### [1 point]
# +
# Enter your code here, printing relevant answers to console:
import pandas as pd
file = pd.read_csv(r"C:\Users\<NAME>\Desktop\IST652\watson_tweets.csv", sep="|") #reading the csv file
file.describe() #tweets count = 300, unique tweets are 164, tweets freq = 2
# -
# ---
# #### ( 2 ) Tokenize the <u>dataframe</u> of tweets into a <u>list</u> of lowercase tokens using the NLTK TweetTokenizer method. Print the number of tokens.
#
# - Consider the method series.tolist() to convert the dataframe to a list.
# - NOTE: The expected object for analysis is a list of tokens, so carefully apply the tokenizer to the original list of tweets.
#
# ##### [1 point]
# +
# Enter your code here, printing relevant answers to console:
import nltk
#nltk.download()
lst = pd.Series.tolist(file)
flat_list = []
for sublist in lst:
for item in sublist:
flat_list = str(flat_list)+str(item)
from nltk.tokenize import word_tokenize
word_tokenize = word_tokenize(flat_list)
tokens = [w.lower() for w in word_tokenize]
del(tokens[0:2]) #removing the first 2 values in tokens that are added by the for loop
len(tokens) #there are 7280 tokens when tokenized by words_tokenize function
# -
lst
# ----
# #### alpha_filter
# +
# function that takes a word and returns true if it consists only
# of non-alphabetic characters
import re
def alpha_filter(w):
# pattern to match a word of non-alphabetical characters
pattern = re.compile('^[^a-z]+$')
if (pattern.match(w)):
return True
else:
return False
# -
# #### ( 3 ) Run the above code block which creates the alpha_filter function. Then, in your own words, describe what the regex query is filtering for.
#
# ##### [1 point]
#
# #### Enter your answer here,<u>in english, not python:</u>
# +
#^ and $ are boundaries or anchors. ^ marks the start, while $ marks the end of a regular expression.
# since ^ a-z is in square braces, the query filters for anything other than a-z (alphabets), which is then added with $ using + operator
#to add numeric values in the items that need to be excluded from filtering
# 'w' in pattern.match(w) matches any non-alphanumeric character [^a-zA-Z0-9]
# Hence the regex qury is filtering for values that are non-alphanumeric
# The function finally returns True if the pattern matched with non-alphanumeric character, else returns false
# -
# ----
# #### ( 4 ) Return a new list of tokens filtered by the alpha_filter function and without StopWords.
# - Recall the stopwords list from NTLK which we covered in class:
# - `nltk.corpus.stopwords.words('english')`
#
# ##### [1 point]
# +
# Enter your code here, printing relevant answers to console:
#Passing tokens through the alpha_filter function
alpha_filtered=[] #new list for adding apha filtered items
for i in tokens: #passes every single token through alpha_filter function and append that token to alpha_filtered
if alpha_filter(i): #only if it is an alpha-numeric value
pass
else:
alpha_filtered.append(i)
alpha_filtered
# Filtering Stopwords
from nltk.corpus import stopwords
#from nltk.tokenize import word_tokenize
stop_words = set(stopwords.words('english'))
#tokens is a word_tokenized file of flat_list
filtered_sentence = [w for w in alpha_filtered if not w in stop_words] #filtering by comparing the alpha_filtered in the list of stop_words
print(filtered_sentence)
# -
# ----
# #### ( 5 ) Apply the frequency distribution to your list from question (4) and print the 50 most common tokens.
# - Note: recall the `nltk.FreqDist` method
#
# ##### [1 point]
# +
# Enter your code here, printing relevant answers to console:
from nltk.probability import FreqDist
fdist = nltk.FreqDist(w.lower() for w in filtered_sentence if w not in stop_words)
#The top 50 common tokens are
for word, frequency in fdist.most_common(50):
print('%s:%d' % (word, frequency))
# -
# ---
# #### ( 6 ) Return the top 25 bigrams by applying a bigram frequency analysis of the tokens found in ( 4 ).
# Note: Use the ntlk methods:
# - `BigramCollocationFinder.from_words()`
# - `score_ngrams()`
#
# ##### [1 point]
# +
# Enter your code here, printing relevant answers to console:
from nltk.collocations import BigramCollocationFinder
bigram_measures = nltk.collocations.BigramAssocMeasures()
finder = BigramCollocationFinder.from_words(tokens) #compute frequency distribution for all the bigrams in the tokens
finder.nbest(bigram_measures.pmi, 25) #top 25 bigram_measures
# -
# ---
| IST652 - Lab 6-Sunkara.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="font-size: 20pt">Python Notebook | Beginner</h1><br/>
#
# <b> Author: </b> <NAME><br/>
# <b> Created: </b> September 2018<br/>
#
# 
# # Table of Content:
# * [Know about Python](#intro)
# * [Numerical and Boolean Operations](#num_bool)
# * [Variable and Object](#var_obj)
# * [Data Type and Type Conversion](#data_type_conversion)
# * [Importing Python Modules and standard libraries](#modules)
# * [Python Built-in Functions and keywords](#builtins)
# * [String operations and formatting](#string_ops)
# * [Useful Pythonic Functions](#pythonic_func)
# * [User-Defined Function](#udf)
# * [Scope of a Variable](#scope)
# * [Answers](#answers)
# * [Additional Read](#additional_read)
# ## What is Python? <a name = "intro"></a>
#
# 
#
# It is a general-purpose, high-level, interpreted, dynamic scripting language.
#
# * **General-Purpose** -> designed to be used for writing software in the widest variety of application domains
# * **High-level** -> designed to be more or less independent of a particular type of computer, human-readable friendly
# * **Interpreted** -> designed to execute instructions of a program directly, without previously compiling a program into machine-language instructions
# * **Scripting** -> languages which are interpreted rather than compiled
#
# ### Q1. But, what is the difference between an interpreter and a compiler? [Answer](#answer1)
# <a name="back1"></a>
# ## Currently, there are two versions of Python
#
# <img src="Photos/Python-logo.png" width="640">
# ## Why do we need Python?
#
# 
#
# * **Simple syntax, readable, reusable and maintainable code**
# * **Multiple Programming Paradigms** such as Object-oriented, structured, functional Paradigms and feature of automatic memory management
# * **Compatible** with Major Platforms and Systems, many **open-source framework available** which boosts developer's productivity
# * **Robust Standard Library** and supporting wide range of external libraries
# * Easier to perform coding and testing simultaneously by adopting **test driven development (TDD)** approach
# ## That's okay. But where do I use it?
#
# 
#
# * **Desktop-based Applications**
# + Image, Video, Audio Processing (using modules like **OpenCV** and **PyAudio** or **librosa**)
# + Graphic Design Applications (used to create **Inkscape, GIMP, Blender, 3ds Max** softwares)
# + Games (**PySoy** -> 3D game engine, **PyGame** -> library for game development)
# * **Scientific and Computational Applications**
# + Machine Learning (Regression, Decision Tree, Random Forest, Deep Learning) using libraries such as **NumPy**, **SciPy**, **TensorFlow**, **Keras**, **PyTorch**
# * **Operating Systems**
# + **Ubuntu’s Ubiquity Installer, Fedora's Anaconda Installer** are written in Python
# * **Language Development**
# + Boo, Apple's Swift, Cobra
# * **Web Development**
# + Framework such as **Django, Pyramid**
# + Micro-Framework such as **Flask, Bottle**
# * **Internet Protocol**
# + HTML, XML
# + JSON
# + Requests
# + BeautifulSoup
# * **Prototyping**
# ## Who uses Python anyway?
#
# 
# ### Cool. I guess we are good to go!
#
# ### If your ground is muddy, can you build your dream house? So, let's make the basics STRONG!
# +
# This is how you comment a code in Python!
# Let's start with the famous code.
print('Hello World')
# -
# a collection of 20 software principles that influences the design of Python Programming Language, of which 19 were written down
import this
# In order to get an understanding of these aphorisms, take a look at this [link](https://artifex.org/~hblanks/talks/2011/pep20_by_example.html).
# # Numerical and Boolean Operations <a name = "num_bool"></a>
# +
## some basic mathematical operations , I am not showing. You do it. But see this.
print(25/4) # floating point division
## to determine quotient and remainder
print(25//4) # floor division
print(25%4) # modulo operator
x = 3
x *= 3 # in-place operator
print(x)
y = "python"
y += "3"
print(y)
# -
# **In Python 2.7.X, it will return the base integer (6 in this case) in both floating point and floor division**. So even if you are using Python 2.7.X, you can do this:
#
# ```
# from __future__ import division
# ```
# which causes Python 2.7.X to adopt the behavior of 3.7.X!
#
# But what is this __from__ stuff?
#
# We will look into it. But let us cover **Boolean operations**.
print(2 == 3)
print(6 != 7)
print(1 == True)
print(2 == True)
print(3 == False)
print(9 is not "nine")
print(5 > 3)
print("Now these.\n---------")
print(6 < 6.0)
print((3 * 0.1) == 0.3)
print((3 * 0.1) == (3/10))
print("Well....!\n---------")
print(3 * 0.1)
print(3/10)
print(3 * 0.2)
print(4 * 0.1)
print(4/10)
print(6 * 0.1)
print(6/10)
print(6 * 0.3)
print(8 * 0.1)
print(1.5 * 0.1)
print(0.75 * 0.1)
# ### Q2. But why is this happening?[Answer](#answer2)
# <a name="back2"></a>
# The following table lists all of Python's operators, from __highest precedence to lowest__.
#
# | Operator | Description |
# | :---:| :---: |
# | ** | Exponentiation (raise to the power) |
# | ~, +, - | Complement, unary plus and minus |
# | * , /, %, // | Multiply, floating point division, modulo and floor division |
# | +,- | Addition and subtraction |
# | >>, << | Right and left bitwise shift |
# | & | Bitwise AND |
# | ^ | Bitwise XOR |
# | | | Bitwise OR |
# | in, not in, is, is not, <, <=, >, >= | Comparison operators, equality operators, membership and identity operators |
# | not | Boolean NOT |
# | and | Boolean AND |
# | or | Boolean OR |
# | =, %=, /=, //=, +=, -=, ```*=``` | Assignment operators |
#
# # Variables and Objects <a name="var_obj"></a>
#
# * Variables do not have a defined type at compile time.
# * Variables can reference any type of object!
# * The variable type can change in run-time.
#
# Type checking is performed at run-time -> hence __dynamic languages are slower than static languages__
#
# __Everything in Python is an Object!__
#
# object is the base type for every Python Object. Objects can be:
# * __Mutable__: value can be changed. [list, set, dictionaries]
# * __Immutable__: value is unchangeable. [number, string, frozenset, tuple]
#
# Object mutability is defined by its __type__.
#
# __Objects Are Never Explicitly Destroyed!__ Allocation and deletion is done by the interpreter.
#
# The most commonly used methods of constructing a multi-word variable name are the last three examples:
#
# * **Camel Case**: Second and subsequent words are capitalized, to make word boundaries easier to see.
# + Example: numberOfCollegeGraduates
# * **Pascal Case**: Identical to Camel Case, except the first word is also capitalized.
# + Example: NumberOfCollegeGraduates
# * **Snake Case**: Words are separated by underscores.
# + Example: number_of_college_graduates
# +
# the correct ways to declare variables
x = 123 # this is right
my_variable = "is my variable, None of your variable" # even this is right
# -
# definitely the wrong way to declare variables
1234_get_on_the_dance_floor = "really?" # this is wrong. see what error you will get
# ## Data Type and Type Conversion <a name = "data_type_conversion"></a>
# +
# data types
print(type("hello"))
print(type(True))
print(type(999.666))
print(type((-1+0j)))
# and also list, tuple, set, dictionary, date, enumerate
# +
# type conversion
print(float(6))
print(str(99) + " " + str(type(str(99))))
print(int("3") + int("6"))
# -
# # Importing Python Modules and standard libraries <a name = "modules"></a>
#
# Python is structured in Modules. From these modules, it is possible to extend the python interpreter capabilities.
# +
## Importing standard and open-source external libaries to your programming environment
##There are several ways to import modules:
# 1. By directly import
import math
# this is fine but avoid this sort of impoting. Instead, try this:
# 2. By specifically importing its functions
from math import sin, log
print(sin(45))
print(log(5))
# better not take the following approach
# 3. By importing everything
from math import *
# +
# if you still wish to use the entire library, you need to do it like this
import math
print(math.sin(45))
# -
print math.log(5)
# ### Some of the default modules in Python
# 
# + hide_input=true
import numpy as np
import keyword
from tabulate import tabulate
# -
# # Python Built-in Functions and keywords <a name = "builtins"></a>
#
# Built-in functions are core functions, provided by the Python Interpreter.
#
# These functions are implemented in C and because of that, are capable of using and manipulating Python objects at memory level, with increased performance.
# + hide_input=true
builtin_functions_list = dir(__builtins__)
builtin_functions_data = np.array(builtin_functions_list)
shape = ((len(builtin_functions_list)//4),4)
print(tabulate(builtin_functions_data.reshape(shape), tablefmt='orgtbl'))
# + hide_input=true
### keywords
keywords_list = keyword.kwlist
keywords_data = np.array(keywords_list)
shape = ((len(keywords_list)//3),3)
print(tabulate(keywords_data.reshape(shape), tablefmt='orgtbl'))
# -
# # String operations and formatting <a name = "string_ops"></a>
# +
## Concatenation
str_var_1 = "concatenation"
print("This is " + str_var_1 + " of string")
print(3 * "3")
print("python" * 3.7)
# +
## string formatting
# It provides a powerful way of embedding non-strings with strings
py_ver = "We are using Python version {}.{}.x".format(3,6)
print(py_ver)
other_version = 2.7
print(f"We could have also used Python {other_version}") #another way of performing string formatting
print("{0}{1}{0}".format("abra ","cad"))
# +
## user input
data_input = int(input("Enter your ID: "))
print(type(data_input))
print("Employee having ID No: {} is late for office today.".format(data_input))
# +
## multiple inputs in a single entry
var_1, var_2 = input("Enter your name: ").split("-")
print(var_1)
print(var_2)
# -
# another way of performing the same thing mentioned above
name_var = input("Enter your name: ").split(" ")
print(name_var[0])
print(name_var[-1])
# # Useful Pythonic Functions <a name = "pythonic_func"></a>
# +
# strings
print(" | ".join(["NASA","Google","Netflix","Yahoo"]))
print("NASA,Google,Netflix,Yahoo".split(","))
print("Python 2.7".replace("2.7","3.6"))
print("There is more to it than meets the eye".startswith("There")) # also endswith is there
print("python".upper())
# +
# numeric
print(max(10,64,86,13,98))
print(abs(-9))
print(sum([2,4,6,8]))
# -
# # User-Defined Function <a name = "udf"></a>
#
# Before going ahead, can you tell me the difference between a function and a method? [Answer](#answer3)
#
# <a name="back3"></a>
# In Python, functions are __first-class objects__ which is an entity that can be
# * dynamically created, destroyed
# * can be stored in a variable
# * passed to a function as a parameter
# * returned as a value from a function
#
# In C++, _classes are not first class objects_ but instances of those classes are. In Python both the classes and the objects are first class objects.
#
# Create your own functions using __def__ keyword.
#
# UDF can either take or don't take arguments. In function definitions, parameters are named entities that specify an argument that a given function can accept.
#
# In other words, **parameters** are the variables in a function definition, and **arguments** are the values placed to the parameters when the function is called.
#
# The name of the UDF should be __lowercase__.
#
# **NOTE: You must define functions before they are called, in the same way you declare a variable before using them**
# +
def udf_1(rqrdArg):
return(3*rqrdArg)
def udf_2(optnlArg = 5):
print("OptionalArg: ", optnlArg)
def udf_3(rqrdArg, optnlArg=None, *args, **kwargs): # *args = extra unnamed argument, **kwargs = extra named arguments
print("RequiredArg: ", rqrdArg)
print("OptionalArg: ", optnlArg)
print("Remaining Non-keyworded args: ", args)
print("Remaining keyworded args: ", kwargs)
print("Calling UDF 1!")
print("RequiredArg: ", udf_1(5.7))
print("\nCalling UDF 2!")
udf_2()
udf_2(9)
print("\nCalling UDF 3!")
udf_3("Python","3.6",2,7,5,1,MSU_Python_Session=1, kwargs_1="good stuffs", kwargs_2=2.7, kwargs_3=True)
# +
def UDF_factorial(x):
"""
This is a recursive function performing factorial of a number
x -> the number whose factorial is calculated
"""
if x == 1:
return 1
else:
return x * UDF_factorial(x-1)
print(UDF_factorial(5))
# -
# ?UDF_factorial
# __*args__ and __**kwargs__ allow one to pass a __variable-length argument list__ and __keyworded, variable-length argument dictionary__ respectively to a function when one doesn't know beforehand how many arguments can be passed to the function.
# +
def multiply(x,y):
print(x*y)
multiply(3,9)
multiply(3,6,9)
# +
# resolving the above issue with *args
def multiply(*args):
x = 1
for num in args:
x *= num
print(x)
multiply(4, 5)
multiply(10, 9)
multiply(2, 3, 4)
multiply(3, 5, 10, 6)
# + code_folding=[]
# likewise for **kwargs
def print_kwargs(**kwargs):
print(kwargs)
print_kwargs(kwargs_1="good stuffs", kwargs_2=2.7, kwargs_3=True)
# -
# When ordering arguments within a function or function call, arguments need to occur in a particular order:
#
# 1. Formal positional arguments
# 2. __*args__
# 3. Keyword arguments
# 4. __**kwargs__
# # Scope of a variable <a name="scope"></a>
#
# Not all variables are accessible from all parts of our program, and not all variables exist for the same amount of time.
#
# A scope is a textual region of a Python program where a __namespace__ is directly accessible.
# A namespace is a mapping from names (variables) to objects.
#
# Basically, part of a program where a variable is accessible is its __scope__ and the duration for which the variable exists its __lifetime__.
#
# There are two types of variable scope:
# * __Global__: It is defined in the main body of a file, will be visible throughout the file, and also inside any file which imports that file.
# * __Local__: it is defined inside a function, limiting its availability inside that function. It is accessible from the point at which it is defined until the end of the function, and exists for as long as the function is executing.
#
# In Python 3, a third type of variable scope has been defined - __nonlocal__. It allows to assign variables in an __outer, but non-global__, scope.
# +
# example showing global, local and nonlocal scopes
# global and local
x = "global"
def foo():
print("x inside :", x)
foo()
print("x outside:", x)
# +
def func1():
msg = "A"
def func2():
msg = "B"
print(msg)
func2()
print(msg)
func1()
# -
# The _msg_ variable is declared in the __func1()__ function and assigned the value _"A"_. Then, in the __func2()__ function, the value _"B"_ is assigned to variable havinf same name _msg_.
#
# When we call the function __func1()__, it is in turn calling function __func2()__ and _msg_ variable in it has the value _"B"_, but Python retains the old value of _"A"_ in the __func1()__ function.
#
# We see this behavior because Python hasn’t actually assigned new value _"B"_ to the existing _msg_ variable, but has created a new variable called __msg in the local scope of the inside function, that shadows the name of the variable in the outer scope__.
#
# Preventing that behavior is where the nonlocal keyword comes in.
# +
#nonlocal
def func1():
msg = "A"
def func2():
msg = "B"
def func3():
nonlocal msg
msg = "C"
print(msg)
func3()
print(msg)
func2()
print(msg)
func1()
# -
# Now, by declaring __nonlocal__ _msg_ in the __func3()__ function, Python knows that when it sees an assignment to msg, it should assign that value to the variable from the __immediate__ outer scope __instead of declaring a new variable that shadows its name__.
#
# The usage of nonlocal is very similar to that of global, except that the former is used for __variables in immediate outer function scopes__.
# # Answers <a name="answers"></a>
# <a name="answer1"></a>
#
# ## Q1 Answer
#
# | Interpreter | Compiler |
# | :---------: | :------: |
# | Executes program by taking **one statement** at a time | Translates the **entire program** at once into machine code |
# | **No intermediate object code** is generated, hence memory efficient | **Generates intermediate object code** which further requires linking, hence requires **more memory** |
# | Less amount of time to **analyze** the source code | More amount of time |
# | Overall **execution** is slower | Faster |
# | Easier to **debug** but less efficient | Difficult but more efficient |
# | Errors are reported after the **entire program is checked** | Error is reported as soon as the **first error is encountered**. Won't show the next set of errors if the existing one isn't solved |
#
# ### Python Interpreter: How does it work?
#
# 
#
# [..back](#back1)
# <a name="answer2"></a>
#
# ## Q2 Answer
#
# Floating point numbers are represented in computer hardware in **base 2**. Floating point numbers are usually represented in **base 10**. *But most decimal fractions cannot be represented exactly as binary fractions.* As a result, the decimal floating-point numbers you enter are only **approximated** to the binary floating-point numbers actually stored in the machine.
#
# No matter how many base 2 digits you are willing to use, the decimal value 0.1 cannot be represented **exactly** as a base 2 fraction.
#
# Many users are not aware of the approximation because of the way values are displayed. If Python were to print the true decimal value of the binary approximation stored for 0.1, it would have to display
# ```
# >>> 0.1
# 0.1000000000000000055511151231257827021181583404541015625
# ```
# But this is more digits than most people find useful, so Python keeps the number of digits manageable by displaying a rounded value instead
# ```
# >>> 0.1
# 0.1
# ```
#
# So, even though printed result looks like the exact value of 1/10, the *actual stored value is the __nearest representable binary fraction__*.
#
# Note that this is in the *very nature of binary floating-point*: this is **NOT a bug in Python**. You’ll see the same kind of thing in all languages that support your hardware’s floating-point arithmetic. **The errors in Python float operations are inherited from the floating-point hardware.**
#
# [..back](#back2)
# <a name="answer3"></a>
#
# ## Q3 Answer
#
# | Method | Function |
# |:---:|:---:|
# | Method is a block of code that is called by its name, but is associated to an object (dependent) | Function is a piece of code that is called by its name but independent |
# | Method is implicitly passed for the object for which it was called | Functions can have parameters, so arguments can be passed to those paramters but explicitely |
#
# [..back](#back3)
# # Additional Read <a name = "additional_read"></a>
#
# * [Floating Point Issue](https://docs.python.org/3/tutorial/floatingpoint.html)
# * [PEP0008 - Style Guide of Python](https://www.python.org/dev/peps/pep-0008/)
# * [The Hitchhker's Guide to Python - Writing Style](https://docs.python-guide.org/writing/style/)
| Python/.ipynb_checkpoints/Python Notebook | Beginner-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#
# BikeShare Pollution Analysis
#
#
# Created by <NAME> on October 20, 2018
# Copyright © 2018 <NAME>. All rights reserved.
#
# -
import pandas as pd
from pandas import DataFrame as df
import matplotlib.pyplot as plt # necessary for plt sub-titles
import warnings
import glob
import os
warnings.simplefilter("ignore")
% matplotlib inline
# The Citibike data was relatively easy to obtain, however, air quality data is not widely reported.
#
# I pulled the air quality data from the New York Department of Envrionmental Conservation (NYC DEC) and although Citibike started in fall 2013, I began analysis in calendar year 2015, first year data was avaliable for the DEC testing site at the City College in Harlem, NY.
#
# The data for both datasets was restricted to 8/30/2018. "AQ_Files" and "CitiBike" are two folders containing daily data 2015-2018. Air quality data is in yearly format, Citibike data is in quartely format
# ## Load Air Quality Data:
# +
path = "./AQ_Files/" # initalizing path for easy use
# Restrict, segment, and reorganize csv
AQ_2015_df = pd.read_csv(path + "AQ2015.csv", usecols = [0,2], skiprows = [0,1,2], skipfooter = 10, names = ['2015 Months', 'CO Level'])
AQ_2016_df = pd.read_csv(path + "AQ2016.csv", usecols = [0,2], skiprows = [0,1,2], skipfooter = 10, names = ['2016 Months', 'CO Level'])
AQ_2017_df = pd.read_csv(path + "AQ2017.csv", usecols = [0,2], skiprows = [0,1,2], skipfooter = 10, names = ['2017 Months', 'CO Level'])
AQ_2018td_df = pd.read_csv(path + "AQ2018TD.csv", usecols = [0,2], skiprows = [0,1,2], skipfooter = 10, names = ['2018 Months', 'CO Level'])
# -
# ## Second, load Citibike Data:
# +
path = r"./CitiBike/"
# Load names into variable for easier df reading
citi_2015 = sorted(glob.glob(os.path.join(path + '2015*.csv'))) # advisable to use os.path.join to make it OS independent
citi_2016 = sorted(glob.glob(os.path.join(path + '2016*.csv')))
citi_2017 = sorted(glob.glob(os.path.join(path + '2017*.csv')))
citi_2018td = sorted(glob.glob(os.path.join(path + '2018*.csv')))
# First reading each csv identified by 'glob' in 'citi_201X' into a df
# Concatening each quartely df into one big yearly pd for easier
df_from_each_file = (pd.read_csv(f) for f in citi_2015)
rides_2015_df = pd.concat(df_from_each_file, ignore_index=True)
df_from_each_file = (pd.read_csv(f) for f in citi_2016)
rides_2016_df = pd.concat(df_from_each_file, ignore_index=True)
df_from_each_file = (pd.read_csv(f) for f in citi_2017)
rides_2017_df = pd.concat(df_from_each_file, ignore_index=True)
df_from_each_file = (pd.read_csv(f) for f in citi_2018td)
rides_2018td_df = pd.concat(df_from_each_file, ignore_index=True)
# Select only certain columns that I'm interested in
rides_2015_df = rides_2015_df[["Date","Trips over the past 24-hours (midnight to 11:59pm)"]]
rides_2016_df = rides_2016_df[["Date","Trips over the past 24-hours (midnight to 11:59pm)"]]
rides_2017_df = rides_2017_df[["Date","Trips over the past 24-hours (midnight to 11:59pm)"]]
rides_2018td_df = rides_2018td_df[["Date","Trips over the past 24-hours (midnight to 11:59pm)"]]
# -
# I now have 8 datasets, 4 AQ, 4 Citibike.
#
# ## Let's look at some sample plots:
# +
# Plot of one citibike df
rides_2015_df.plot()
plt.xlabel("Date"); plt.ylabel("Daily Trips"); plt.title("2015 Citibike")
# Plot of one air quality df
AQ_2015_df.plot()
plt.xlabel("Date"); plt.ylabel("Air Quality"); plt.title("2015 Air Quality")
# -
# For some reason the date is showing up at "XX days" in the year, as in Jan 1 = 0. Not sure how to fix that
# +
# Plot of one citibike df
rides_2016_df.plot()
plt.xlabel("Date"); plt.ylabel("Daily Trips"); plt.title("2016 Citibike")
# Plot of one air quality df
AQ_2016_df.plot()
plt.xlabel("Date"); plt.ylabel("Air Quality"); plt.title("2016 Air Quality")
# +
# Plot of one citibike df
rides_2017_df.plot()
plt.xlabel("Date"); plt.ylabel("Daily Trips"); plt.title("2017 Citibike")
# Plot of one air quality df
AQ_2017_df.plot()
plt.xlabel("Date"); plt.ylabel("Air Quality"); plt.title("2017 Air Quality")
# +
# Plot of one citibike df
rides_2018td_df.plot()
plt.xlabel("Date"); plt.ylabel("Daily Trips"); plt.title("2018 Citibike To Date")
# Plot of one air quality df
AQ_2018td_df.plot()
plt.xlabel("Date"); plt.ylabel("Air Quality"); plt.title("2018 Air Quality To Date")
# -
# As you can tell, there are missing values for air quality in the start of 2018 and there was something very bad happening around day 40 (feb)
#
# Regression will be shown in R using these datapoints
#
# Visualizations will be made in Tableau
| Python_Analysis/Bikeshare_Pollution_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray as xr
import numpy as np
from xhistogram.xarray import histogram
from matplotlib import pyplot as plt
from matplotlib import cm
def histsum_ds(ds,bin_data,bins,dim):
ds = ds.copy()
ds_rebinned = xr.Dataset()
for var in ds.data_vars:
if ds[var].dtype == 'float32':
nanmask = np.isnan(ds[var])
ds_rebinned[var] = histogram(
bin_data.where(~nanmask), bins=[bins], dim=dim, weights=(ds[var]*ds['volcello']).where(~nanmask)
)
return ds_rebinned
# Load grid data
directory = '/archive/gam/ESM4/DECK/ESM4_piControl_D/gfdl.ncrc4-intel16-prod-openmp/history/08990101.ocean_static_no_mask_table.nc'
grid = xr.open_dataset(directory)
# ### Seasonality of phytoplankton and zooplankton
# Specify paths
rootdir = '/archive/oar.gfdl.cmip6/ESM4/DECK/ESM4_piControl_D/gfdl.ncrc4-intel16-prod-openmp/pp/'
pp_cobalt = 'ocean_cobalt_omip_tracers_month_z'
pp_mom6 = 'ocean_monthly_z'
averages = '/av/monthly_5yr/'
years = '0901-0905'
months = '*'
filename_cobalt = pp_cobalt+'.'+years+'.'+months+'.nc'
filename_mom6 = pp_mom6+'.'+years+'.'+months+'.nc'
path_cobalt = rootdir+pp_cobalt+averages+filename_cobalt
path_mom6 = rootdir+pp_mom6+averages+filename_mom6
# Load data
ds_cobalt = xr.open_mfdataset(path_cobalt)
ds_mom6 = xr.open_mfdataset(path_mom6)
# Calculate bgc variables as a function of temperature
bins_theta = np.arange(-2,32,1)
ds_cobalt_rebinned = histsum_ds(ds_cobalt,ds_mom6.thetao,bins=bins_theta,dim=['xh','yh','z_l'])
ds_cobalt_rebinned.phyc.load();
ds_cobalt_rebinned.zooc.load();
# +
start = 0
stop = 1
N = 12
colors = [ cm.BrBG(x) for x in np.linspace(start, stop, N) ]
fig,ax = plt.subplots(figsize = (10,10), nrows = 2)
for t in range(12):
ax[0].plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.phyc.isel(time=t),color=colors[t])
ax[1].plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.zooc.isel(time=t),color=colors[t])
ax[0].set_title('phyc')
ax[0].grid()
ax[1].set_title('zooc')
ax[1].grid()
# -
# ### Annual mean rates of biogeochemical processes
# Specify paths
rootdir = '/archive/oar.gfdl.cmip6/ESM4/DECK/ESM4_piControl_D/gfdl.ncrc4-intel16-prod-openmp/pp/'
pp_cobalt = 'ocean_cobalt_omip_rates_year_z'
pp_mom6 = 'ocean_annual_z'
averages = '/av/annual_5yr/'
years = '1101-1105'
months = 'ann'
filename_cobalt = pp_cobalt+'.'+years+'.'+months+'.nc'
filename_mom6 = pp_mom6+'.'+years+'.'+months+'.nc'
path_cobalt = rootdir+pp_cobalt+averages+filename_cobalt
path_mom6 = rootdir+pp_mom6+averages+filename_mom6
# Load data
ds_cobalt = xr.open_mfdataset(path_cobalt)
ds_mom6 = xr.open_mfdataset(path_mom6)
# Calculate bgc variables as a function of temperature
bins_theta = np.arange(-2,32,1)
ds_cobalt_rebinned = histsum_ds(ds_cobalt,ds_mom6.thetao,bins=bins_theta,dim=['xh','yh','z_l','time'])
# Separately histogram sinking term, to integrate in area rather than volume
expc_rebinned = histogram(ds_mom6.thetao,bins=[bins_theta],dim=['xh','yh','z_l','time'],weights=ds_cobalt.expc*grid.areacello)
expc_rebinned.load();
ds_cobalt_rebinned.pp.load();
ds_cobalt_rebinned.remoc.load();
ds_cobalt_rebinned.graz.load();
ds_cobalt_rebinned.expc.load();
ds_cobalt_rebinned.volcello.load();
expc_rebinned.load()
fig,ax = plt.subplots(figsize = (10,5), nrows = 1)
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.pp,label='pp')
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.remoc,label='remoc')
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.graz,label='graz')
ax.plot(expc_rebinned.thetao_bin,expc_rebinned,label='expc')
ax.legend()
fig,ax = plt.subplots(figsize = (10,5), nrows = 1)
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.pp,label='pp')
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.ppdiat,label='ppdiat')
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.ppdiaz,label='ppdiaz')
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.pppico,label='pppico')
ax.plot(ds_cobalt_rebinned.thetao_bin,ds_cobalt_rebinned.ppmisc,label='ppmisc')
ax.legend()
| notebooks/archive/draw_bgc-in-TS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib widget
from util import get_path
import pandas as pd
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
from extract_graph import generate_nx_graph, transform_list, generate_skeleton, generate_nx_graph_from_skeleton, from_connection_tab, from_nx_to_tab
from node_id import whole_movement_identification, second_identification
import ast
from plotutil import plot_t_tp1, compress_skeleton
from scipy import sparse
from sparse_util import dilate, zhangSuen
from realign import realign
from datetime import datetime,timedelta
from node_id import orient
import pickle
from matplotlib.widgets import CheckButtons
import scipy.io as sio
import imageio
from pymatreader import read_mat
from matplotlib import colors
from copy import deepcopy,copy
from collections import Counter
import cv2
import imageio
import matplotlib.pyplot as plt
import numpy as np
from skimage.filters import frangi, meijering
from skimage.morphology import thin
from skimage import data, filters
from random import choice
import scipy.sparse
import os
from time import time
from extract_graph import dic_to_sparse, from_sparse_to_graph, generate_nx_graph, prune_graph, from_nx_to_tab, from_nx_to_tab_matlab,sparse_to_doc, connections_pixel_list_to_tab, transform_list, clean_degree_4
from time import sleep
from skimage.feature import hessian_matrix_det
from experiment_class_surf import Experiment,clean_exp_with_hyphaes, Hyphae
from hyphae_id_surf import clean_and_relabel, get_mother, save_hyphaes, resolve_ambiguity_two_ends,solve_degree4, clean_obvious_fake_tips
from realign import transform_skeleton_final
plate = 3
begin = 0
end = 19
directory = "/scratch/shared/mrozemul/Fiji.app/"
listdir=os.listdir(directory)
list_dir_interest=[name for name in listdir if name.split('_')[-1]==f'Plate{0 if plate<10 else ""}{plate}']
ss=[name.split('_')[0] for name in list_dir_interest]
ff=[name.split('_')[1] for name in list_dir_interest]
dates_datetime=[datetime(year=int(ss[i][:4]),month=int(ss[i][4:6]),day=int(ss[i][6:8]),hour=int(ff[i][0:2]),minute=int(ff[i][2:4])) for i in range(len(list_dir_interest))]
dates_datetime.sort()
dates_datetime_chosen=dates_datetime[begin:end+1]
dates = [f'{0 if date.month<10 else ""}{date.month}{0 if date.day<10 else ""}{date.day}_{0 if date.hour<10 else ""}{date.hour}{0 if date.minute<10 else ""}{date.minute}' for date in dates_datetime_chosen]
exp = pickle.load( open(f'/scratch/shared/mrozemul/Fiji.app/Analysis_Plate{plate}_{dates[0]}_{dates[-1]}/experiment_{plate}.pick', "rb" ) )
def get_hyphae(experiment,exclude_bottom_factor=0.98):
tips = [node for node in experiment.nodes if node.degree(node.ts()[0])==1 and node.pos(node.ts()[0])[0]<=experiment.boundaries_x[1]*exclude_bottom_factor]
problems=[]
small_problems = []
hyphaes=[]
# for i in range(20):
for i,tip in enumerate(tips):
if i%200==0:
print(i/len(tips))
# tip = choice(tips)
hyphae = Hyphae(tip)
roots = []
for t in tip.ts():
# print(t,tip)
if tip.degree(t)==1:
root,edges,nodes = hyphae.get_edges(t,100)
roots.append(root)
occurence_count = Counter(roots)
if len(occurence_count.values())>=2:
small_problems.append(tip)
if len(occurence_count.values())>=2 and occurence_count.most_common(2)[0][0]!=roots[0] and occurence_count.most_common(2)[1][1]/occurence_count.most_common(2)[0][1]>=0.75:
problems.append(tip)
else:
hyphae.root = occurence_count.most_common(2)[0][0]
hyphae.ts = sorted(set(hyphae.ts).intersection(set(hyphae.root.ts())))
hyphaes.append(hyphae)
print(f'Detected problems during hyphae detection, {len(small_problems)} hyphaes have inconsistent root over time')
print(f'Detected problems during hyphae detection, {len(problems)} hyphaes have inconsistent root over time')
return(hyphaes,problems,small_problems)
hyphaes,problems,small_problems = get_hyphae(exp)
problems
experiment = exp
tips = [node for node in experiment.nodes if node.degree(node.ts()[0])==1 and node.pos(node.ts()[0])[0]<=experiment.boundaries_x[1]*0.98]
small_problems = []
hyphaes=[]
tip =choice(tips)
hyphae = Hyphae(tip)
roots = []
tss=[]
for t in tip.ts():
# print(t,tip)
if tip.degree(t)==1:
tss.append(t)
root,edges,nodes = hyphae.get_edges(t,100)
roots.append(root)
roots[10:15],tss
plt.close('all')
begin=10
end=15
exp.plot(tss[begin:end],[[node.label,tip.label] for node in roots][begin:end])
tip.show_source_image(tip.ts()[10],tip.ts()[12])
| amftrack/notebooks/development/improve_hyphae_id.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ptpxaHQc9NeT"
# # Basic Web Scraping with `BeautifulSoup`
# I'm building a notebook for web scraping using Python to develop my skills using the popular `BeautifulSoup` Python library ([Documentation](https://www.crummy.com/software/BeautifulSoup/bs4/doc/)). It will be based on projects that I have wanted to do, so the code will have real world examples for scraping websites and provide me with a good opportunity to clean 'dirty' data and create data sets that are easily manipulated and worked with for further analysis, machine learning techniques or visualisations. In this particular case, I'm looking to build up a clean dataset of different gins.
#
# ---
#
# Basic flow:
# 1. You have to scrape the raw html for a given url using `requests`
# 2. You have to trim the information you want from that html via elements, classes and ids using `BeuatifulSoup`
# 1. Identifying the classes and ids of the elements that you want to extract information from has appeared to be a vital part of the process
# 3. Manipulate that information into a easily usable format using `pandas`
#
# + colab={} colab_type="code" id="XXCeLfHg9MEu"
# Imports
import requests
import pandas as pd
from bs4 import BeautifulSoup
import numpy as np
# + [markdown] colab_type="text" id="FxMyBW_2Ah5l"
# ## Searching the HTML using a class
# Every website is going to have some conventions for formatting pages and the information that we ultimately want to acquire. This is likely going to come in the form of CSS classes, especially if the website logic is dynamic and they are producing many pages with different information using the same template file. Thus, being able to isolate those elements using a distinct class is incredibly useful for us.
#
# This can be achieved through a special use case of the `find_all()` function ([Documentation](https://www.crummy.com/software/BeautifulSoup/bs4/doc/#searching-by-css-class)).
#
# In my example here, I am going to be trying to scrape all the information about the gins that are available from the site 'Master of Malt'.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 1534, "status": "ok", "timestamp": 1542840585739, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="JIwNTdfG9i5W" outputId="3207c27a-c07c-450b-a4a5-5369d489018d"
# soup.find_all(class_="sectionHeader")
url = "https://www.masterofmalt.com/gin/"
# By setting a cookie for this example, I am able to obtain the prices in GBP
# This was something I had to do, as the default when running this was giving me USD
# I investigated the console through my browser and realised that setting this fixed the issue I was having
cookies = dict(MaOMa='VisitorID=556630649&IsVATableCountry=1&CountryID=464&CurrencyID=-1&CountryCodeShort=GB&DeliveryCountrySavedToDB=1')
html = requests.get(url, headers = {"Accept-Language": "en-GB"}, cookies = cookies).text
soup = BeautifulSoup(html, features="html.parser")
# View that the object type that we have created is a `bs4.BeautifulSoup`
print(type(soup))
# Print the title of the page
print(soup.title.get_text())
# -
# Since all of the gins for the site aren't on a single page, we are going to have to work out a way to loop through the pages. This will inevitably start with me finding the pagination section on the page and getting all the links.
#
# Once we have the links we should start working on a system for noting which ones we have visited and which we are yet to scrape.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 457, "status": "ok", "timestamp": 1542837744835, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="ZmypPAZL8cAT" outputId="420c9edb-bb75-4005-c4fc-896da48440d8"
pagination = soup.find_all(class_='list-paging')
# We can print to see what we have, looks as though we have two versions of the same thing
# print(pagination)
# Note on why you access the first element, to go from result set to something you can call find_all on again
# https://stackoverflow.com/questions/24108507/beautiful-soup-resultset-object-has-no-attribute-find-all
# Remove all the anchor tags from the pagination html
pagination_links = pagination[0].find_all('a')
# pagination[0].find_all('a')[0].get('href') == url
# -
# Below we can loop through and extract the href from all the anchor tags that aren't our original url. This will surely form the basis of our navigation system through the site for the scraper.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 437, "status": "ok", "timestamp": 1542753209875, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="z0tUUpNMDcT_" outputId="18a53df7-fcd4-4fc4-bbf5-53db19e2771c"
[link.get('href') for link in pagination_links if link.get('href') != url]
# -
# You can inspect the pages that you are going to loop through, and note constant elements that are going to act as the starting point for your data collection.
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 442} colab_type="code" executionInfo={"elapsed": 556, "status": "ok", "timestamp": 1542840589656, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="mDA-XbE4QcvQ" outputId="07bd3c96-c8da-4b47-9405-60edbc70bc14"
# Original url looping
# The class "boxBgr product-box-wide h-gutter js-product-box-wide"
# appears on all the main drink elements for the index page
main_product_boxes = soup.find_all(class_="boxBgr product-box-wide h-gutter js-product-box-wide")
# Loop through each element and get the current price span/div and then the contained text (price)
[product.find(class_="product-box-wide-price gold").get_text() for product in main_product_boxes]
# -
# Get the product ids from the data property on the element
# Could potentially be a way of avoiding duplication of results
[main_product_box.get('data-productid') for main_product_box in main_product_boxes]
# + colab={"base_uri": "https://localhost:8080/", "height": 442} colab_type="code" executionInfo={"elapsed": 500, "status": "ok", "timestamp": 1542838796252, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="0fPPpQrBQzhl" outputId="13bd12d9-4390-4fd5-9840-c07b8d69129f"
product_details = []
for product in main_product_boxes:
name = product.find('h3').get_text()
volume_strength = product.find(class_="product-box-wide-volume gold").get_text()
optional_rating = product.select('span[id$=ratingStars]')
# print(len(optional_rating))
if len(optional_rating) > 0:
rating = optional_rating[0].get('title')
else:
rating = 'Unknown'
review_count = product.select('span[id$=reviewCount]')[0].get_text() if len(product.select('span[id$=reviewCount]')) > 0 else 'Unknown'
price = product.find(class_="product-box-wide-price gold").get_text()
product_details.append([name, volume_strength, rating, review_count, price])
# + colab={} colab_type="code" id="3QTZ0JbGU6Dy"
gin_df = pd.DataFrame(product_details)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 501, "status": "ok", "timestamp": 1542838799023, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="frdNcGY_S-Yq" outputId="88e66215-4b27-40c9-ed15-2b141e06d499"
gin_df.columns = ['Gin', 'Vol_Strength', 'Rating', 'Review_count', 'Price']
gin_df.head()
# + colab={} colab_type="code" id="AAP6AY7cTXUR"
split = gin_df['Vol_Strength'].str.split(',', expand = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 824} colab_type="code" executionInfo={"elapsed": 457, "status": "ok", "timestamp": 1542838032266, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="oz1hOSnLT3Vl" outputId="9002d5b0-9a83-4654-f12f-f1e55f0478d9"
split
# + colab={} colab_type="code" id="nlzcOZmZT6aS"
gin_df['Volume'] = split[0]
gin_df['Strength'] = split[1]
gin_df.drop(columns = ['Vol_Strength'], inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 824} colab_type="code" executionInfo={"elapsed": 514, "status": "ok", "timestamp": 1542838146356, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="D9VwAujeT74B" outputId="265129e4-92dd-44a1-a3f4-4122b6baa97c"
gin_df
# + colab={} colab_type="code" id="150c7uPEUXuL"
import re
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 432, "status": "ok", "timestamp": 1542838563050, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="6P5AGohPUtbv" outputId="df54e796-eaf9-478a-9950-218471a1aec8"
def extract_rating(string):
if bool(re.search(r'\((.*?)\)', string)):
return re.search(r'\((.*?)\)', string).group(1)
else:
return np.nan
bool(re.search(r'\((.*?)\)',gin_df['Rating'][0]))
# + colab={} colab_type="code" id="qnkstGsvVLGT"
gin_df['Rating'] = gin_df['Rating'].apply(extract_rating)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 449, "status": "ok", "timestamp": 1542838586566, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="SrN_v8cSVtqq" outputId="e99a43ec-2b01-4125-d754-23c7651ca670"
gin_df.head()
# + colab={} colab_type="code" id="_A-_tkw-WHsR"
# Before I fixed the cookies request issue
# gin_df['Price'] = gin_df['Price'].str.replace('$', '£')
# + colab={} colab_type="code" id="-w15eKZ0WOdz"
def extract_review_count(string):
if string.find('Reviews') != -1:
return string.replace('Reviews', '')
elif string.find('1 Review') != -1:
return '1'
else:
return np.nan
# + colab={} colab_type="code" id="NsSNooGJWg-d"
gin_df['Review_count'] = gin_df['Review_count'].apply(extract_review_count)
# -
# We can now have a look at the first few rows of the table to see that we have a much clearer data structure that is going to be much easier to work with for further analysis.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 398, "status": "ok", "timestamp": 1542838708277, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08233812395679008502"}, "user_tz": 0} id="vD7Pl6KcWk3V" outputId="119c1002-6503-48b5-e875-b861ca8792f2"
gin_df.head()
# -
# What I have done above represents just looking through a single index page for the gins and pulling the basic information that would be designed to try and get you to then click on the product to go into the full page for that particular product.
#
# If we want to gain all the information that can about the gins then this is something that we are going to have to implement.
| master_of_malt/web_scraping_guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fast Style Transfer with FastEstimator
#
# In this notebook we will demonstrate how to do a neural image style transfer with perceptual loss as described in [Perceptual Losses for Real-Time Style Transfer and Super-Resolution](https://cs.stanford.edu/people/jcjohns/papers/eccv16/JohnsonECCV16.pdf).
# Typical neural style transfer involves two images, an image containing semantics that you want to preserve and another image serving as a reference style; the first image is often referred as *content image* and the other image as *style image*.
# In [paper](https://cs.stanford.edu/people/jcjohns/papers/eccv16/JohnsonECCV16.pdf) training images of COCO2014 dataset are used to learn the style transfer from any content image.
# +
import os
import cv2
import fastestimator as fe
import tensorflow as tf
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
# -
# In this notebook we will use *Wassily Kandinsky's Composition 7* as a style image.
# We will also resize the style image to $256 \times 256$ to make the dimension consistent with that of COCO images.
# +
style_img_path = tf.keras.utils.get_file(
'kandinsky.jpg',
'https://storage.googleapis.com/download.tensorflow.org/example_images/Vassily_Kandinsky%2C_1913_-_Composition_7.jpg'
)
style_img = cv2.imread(style_img_path)
style_img = cv2.resize(style_img, (256, 256))
style_img = (style_img.astype(np.float32) - 127.5) / 127.5
style_img_t = tf.convert_to_tensor(np.expand_dims(style_img, axis=0))
style_img_disp = cv2.cvtColor((style_img + 1) * 0.5, cv2.COLOR_BGR2RGB)
plt.imshow(style_img_disp)
plt.title('<NAME>\'s Composition 7')
plt.axis('off');
# + tags=["parameters"]
#Parameters
batch_size = 4
epochs = 2
steps_per_epoch = None
validation_steps = None
img_path = 'panda.jpeg'
saved_model_path = 'style_transfer_net_epoch_1_step_41390.h5'
# -
# ## Step 1: Input Pipeline
#
# ### Downloading the data
#
# First, we will download training images of COCO2014 dataset via our dataset API.
# The images will be first downloaded. Then, a csv file containing relative paths to these images will be created.
# The root path of the downloaded images will be parent_path.
# Downloading the images will take awhile.
from fastestimator.dataset.mscoco import load_data
train_csv, path = load_data()
# Once finished downloading images, we need to define an *Operator* to recale pixel values from $[0, 255]$ to $[-1, 1]$.
# We will define our own `Rescale` class in which we define the data transform logic inside `forward` method.
# +
from fastestimator.op import TensorOp
class Rescale(TensorOp):
def forward(self, data, state):
return (tf.cast(data, tf.float32) - 127.5) / 127.5
# -
# ### Creating tfrecords
# Once the images are downloaded, we will create tfrecords using `RecordWriter`.
# Each row of the csv file will be used by `ImageReader` to read in the image using `cv2.imread`.
# Then, we resize the images to $256 \times 256$.
from fastestimator.op.numpyop import ImageReader, Resize
from fastestimator.util import RecordWriter
tfr_save_dir = os.path.join(path, 'tfrecords')
writer = RecordWriter(
train_data=train_csv,
save_dir=tfr_save_dir,
ops=[
ImageReader(inputs="image", parent_path=path, outputs="image"),
Resize(inputs="image", target_size=(256, 256), outputs="image")
])
# ### Defining an instance of `Pipeline`
# We can now define an instance of `Pipeline`.
pipeline = fe.Pipeline(batch_size=batch_size, data=writer, ops=[Rescale(inputs="image", outputs="image")])
# ## Step 2: Network
# Once `Pipeline` is defined, we need to define network architecture, losses, and the forward pass of batch data.
#
# ### Defining model architecture
# We first create a `FEModel` instance which collects the following:
# * model definition
# * model name
# * loss name
# * optimizer
#
# The architecture of the model is a modified resnet.
# +
from fastestimator.architecture.stnet import styleTransferNet
model = fe.build(model_def=styleTransferNet,
model_name="style_transfer_net",
loss_name="loss",
optimizer=tf.keras.optimizers.Adam(1e-3))
# -
# ### Defining Loss
#
# The perceptual loss described in the [paper](https://cs.stanford.edu/people/jcjohns/papers/eccv16/JohnsonECCV16.pdf) is computed based on intermediate layers of VGG16 pretrained on ImageNet; specifically, `relu1_2`, `relu2_2`, `relu3_3`, and `relu4_3` of VGG16 are used.
# The *style* loss term is computed as the squared l2 norm of the difference in Gram Matrix of these feature maps between an input image and the reference stlye image.
# The *content* loss is simply l2 norm of the difference in `relu3_3` of the input image and the reference style image.
# In addition, the method also uses total variation loss to enforce spatial smoothness in the output image.
# The final loss is weighted sum of the style loss term, the content loss term (feature reconstruction term in the [paper](https://cs.stanford.edu/people/jcjohns/papers/eccv16/JohnsonECCV16.pdf)), and the total variation term.
#
# We first define a custom `TensorOp` that outputs intermediate layers of VGG16.
# Given these intermediate layers returned by the loss network as a dictionary, we define a custom `Loss` class that encapsulates all the logics of the loss calculation.
# Since `Loss` is also yet another `TensorOp`, the final loss value is returned by `forward` method.
# +
from fastestimator.architecture.stnet import lossNet
from fastestimator.op.tensorop import Loss
class ExtractVGGFeatures(TensorOp):
def __init__(self, inputs, outputs, mode=None):
super().__init__(inputs, outputs, mode)
self.vgg = lossNet()
def forward(self, data, state):
return self.vgg(data)
class StyleContentLoss(Loss):
def __init__(self, style_weight, content_weight, tv_weight, inputs, outputs=None, mode=None):
super().__init__(inputs=inputs, outputs=outputs, mode=mode)
self.style_weight = style_weight
self.content_weight = content_weight
self.tv_weight = tv_weight
def calculate_style_recon_loss(self, y_true, y_pred):
y_true_gram = self.calculate_gram_matrix(y_true)
y_pred_gram = self.calculate_gram_matrix(y_pred)
y_diff_gram = y_pred_gram - y_true_gram
y_norm = tf.math.sqrt(tf.reduce_sum(tf.math.square(y_diff_gram), axis=(1, 2)))
return (y_norm)
def calculate_feature_recon_loss(self, y_true, y_pred):
y_diff = y_pred - y_true
num_elts = tf.cast(tf.reduce_prod(y_diff.shape[1:]), tf.float32)
y_diff_norm = tf.reduce_sum(tf.square(y_diff), axis=(1, 2, 3)) / num_elts
return (y_diff_norm)
def calculate_gram_matrix(self, x):
x = tf.cast(x, tf.float32)
num_elts = tf.cast(x.shape[1] * x.shape[2] * x.shape[3], tf.float32)
gram_matrix = tf.einsum('bijc,bijd->bcd', x, x)
gram_matrix /= num_elts
return gram_matrix
def calculate_total_variation(self, y_pred):
return (tf.image.total_variation(y_pred))
def forward(self, data, state):
y_pred, y_style, y_content, image_out = data
style_loss = [self.calculate_style_recon_loss(a, b) for a, b in zip(y_style['style'], y_pred['style'])]
style_loss = tf.add_n(style_loss)
style_loss *= self.style_weight
content_loss = [
self.calculate_feature_recon_loss(a, b) for a, b in zip(y_content['content'], y_pred['content'])
]
content_loss = tf.add_n(content_loss)
content_loss *= self.content_weight
total_variation_reg = self.calculate_total_variation(image_out)
total_variation_reg *= self.tv_weight
return style_loss + content_loss + total_variation_reg
# -
# ### Defining forward pass
#
# Having defined the model and the associated loss, we can now define an instance of `Network` that specify forward pass of the batch data in a training loop.
# FastEstimator takes care of gradient computation and update of the model once this forward pass is defined.
# +
from fastestimator.op.tensorop import ModelOp
style_weight=5.0
content_weight=1.0
tv_weight=1e-4
network = fe.Network(ops=[
ModelOp(inputs="image", model=model, outputs="image_out"),
ExtractVGGFeatures(inputs=lambda: style_img_t, outputs="y_style"),
ExtractVGGFeatures(inputs="image", outputs="y_content"),
ExtractVGGFeatures(inputs="image_out", outputs="y_pred"),
StyleContentLoss(style_weight=style_weight,
content_weight=content_weight,
tv_weight=tv_weight,
inputs=('y_pred', 'y_style', 'y_content', 'image_out'),
outputs='loss')
])
# -
# ## Step 3: Estimator
# Having defined `Pipeline` and `Network`, we can now define `Estimator`.
# We will use `Trace` to save intermediate models.
# +
from fastestimator.trace import ModelSaver
import tempfile
model_dir=tempfile.mkdtemp()
estimator = fe.Estimator(network=network,
pipeline=pipeline,
epochs=epochs,
steps_per_epoch=steps_per_epoch,
validation_steps=validation_steps,
traces=ModelSaver(model_name="style_transfer_net", save_dir=model_dir))
# -
# We call `fit` method of `Estimator` to start training.
estimator.fit()
# ## Inference
#
# Once the training is finished, we will apply the model to perform the style transfer on arbitrary images.
# Here we use a photo of a panda.
test_img = cv2.imread(img_path)
test_img = cv2.resize(test_img, (256, 256))
test_img = (test_img.astype(np.float32) - 127.5) / 127.5
test_img_t = tf.expand_dims(test_img, axis=0)
model_path = os.path.join(model_dir, saved_model_path)
trained_model = tf.keras.models.load_model(model_path,
custom_objects={
"ReflectionPadding2D":fe.architecture.stnet.ReflectionPadding2D,
"InstanceNormalization":fe.architecture.stnet.InstanceNormalization},
compile=False)
output_img = trained_model.predict(test_img_t)
# +
output_img_disp = (output_img[0] + 1) * 0.5
test_img_disp = (test_img + 1) * 0.5
plt.figure(figsize=(20,20))
plt.subplot(131)
plt.imshow(cv2.cvtColor(test_img_disp, cv2.COLOR_BGR2RGB))
plt.title('Original Image')
plt.axis('off');
plt.subplot(132)
plt.imshow(style_img_disp)
plt.title('Style Image')
plt.axis('off');
plt.subplot(133)
plt.imshow(cv2.cvtColor(output_img_disp, cv2.COLOR_BGR2RGB));
plt.title('Transferred Image')
plt.axis('off');
| docs/apphub/image_styletransfer/fst_coco/fst_coco.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import html
lines = [html.unescape(line.rstrip('\n')) for line in open('./data/raw')]
# +
# remove 'www.' for uniformity
_re1 = re.compile(r'www.', re.UNICODE)
# replace complicated urls with homepage
_re2 = re.compile(r'(http|ftp|https)://([\w_-]+(?:(?:\.[\w_-]+)+))([\w.,@?^=%&:/~+#-]*[\w@?^=%&/~+#-])?', re.UNICODE)
for idx, line in enumerate(lines):
line = _re1.sub('', line)
url = _re2.findall(line)
if url:
lines[idx] = _re2.sub(url[0][1], line)
# +
# handle smileys
_re1 = re.compile(r'(:-?\(|:-?\)|:-?\\|:-?D)', re.UNICODE)
for idx, line in enumerate(lines):
# simply sub them
lines[idx] = _re1.sub(' ', line)
# alternatively pad them
# '</e>'.join(_re1.split(line))
# +
# handle . / and , characters
# (behave differently for numbers are characters)
_re1 = re.compile(r'(?<=[A-z])[\.\/\,]', re.UNICODE)
# replace numbers with #
_re2 = re.compile(r'[0-9]+', re.UNICODE)
# handle double occurences of #
_re3 = re.compile(r'#[#]+', re.UNICODE)
for idx, line in enumerate(lines):
line = _re1.sub(' ', line)
line = _re2.sub('#', line)
lines[idx] = _re3.sub('#', line)
# +
# remove special characters
_re1 = re.compile(r'[\!\?\^\*\(\)\[\]\{\}\;\:\<\>\~\_\\]', re.UNICODE)
# remove multiple (long) dashes
_re2 = re.compile(r'.[.]+', re.UNICODE)
# remove multiple (long) dashes
_re3 = re.compile(r'-[-]+', re.UNICODE)
# remove space padded hyphens
_re4 = re.compile(r'\s-\s', re.UNICODE)
# replace & with 'and'
_re5 = re.compile(r'&', re.UNICODE)
# replace w/ with 'with'
_re6 = re.compile(r'w\/', re.UNICODE)
for idx, line in enumerate(lines):
line = _re1.sub(' ', line)
line = _re2.sub(' ', line)
line = _re3.sub(' ', line)
line = _re4.sub(' ', line)
line = _re5.sub(' and ', line)
lines[idx] = _re6.sub(' with ', line)
# +
# drop backslashes
_re1 = re.compile(r'\\', re.UNICODE)
for idx, line in enumerate(lines):
lines[idx] = _re1.sub('', line)
# +
# pad quotes (non-apostrophe) with space
_re1 = re.compile(r"(?<![a-z])[\']", re.UNICODE)
_re2 = re.compile(r"[\'](?![a-z])", re.UNICODE)
_re3 = re.compile(r'\"', re.UNICODE)
for idx, line in enumerate(lines):
line = _re1.sub(' ', line)
line = _re2.sub(' ', line)
lines[idx] = _re3.sub(' " ', line)
# +
# omit double+ occurences (spaces etc.)
_re1 = re.compile(r'\s[\s]+', re.UNICODE)
for idx, line in enumerate(lines):
lines[idx] = _re1.sub(' ', line)
# -
# lower case everything
lines = [line.lower() for line in lines]
# join with </s>
lines = ' </s> '.join(lines)
with open('./data/text', 'w') as f:
f.write(lines)
lines.decode('utf-8')
| notebooks/preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import sklearn
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn import svm
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Dense, LSTM
from keras.callbacks import EarlyStopping
import warnings
import matplotlib.pyplot as plt
warnings.filterwarnings("ignore")
# ### Pipeline psuedo code
# n = look back window
# k = number of PCs to keep
#
# for each time point t:
# p = number of stocks in investable universe at time t
# Define an n x p feature matrix X (lagged returns)
#
# Perform PCA on X
# Keep the first k PCs in an n x k matrix Z
#
# for each stock s in the investable universe at time t:
# Define an n x 1 outcome vector y (future returns of stock s)
# Perform a linear regression of y on Z
# Predict y for stock s at time t+1
# ## Data Preparation
returns = pd.read_pickle("./Data/returns.pkl")
returns = returns.iloc[1:]
# +
drop_columns = []
for col in returns.columns:
if returns[col].isnull().all() == True:
drop_columns.append(col)
returns.drop(columns=drop_columns, inplace=True)
# -
# ## Pipeline
def get_investable(t, n):
""""Find stocks in investable universe at time t+1
(stocks in the S&P500 that have prices recorded for the last n days)"""
df_investable = returns.copy(deep = True).sort_index(ascending = False)
#add 1 date to get the test features in investable
t = t + pd.DateOffset(1)
n += 1
#if t is now a non-trading day, advance until we reach a valid trading day
while t not in df_investable.index:
t = t + pd.DateOffset(1)
t_index = df_investable.index.get_loc(t)
#take n_rows worth of data upto time specified
df_investable = df_investable.iloc[t_index:t_index + n]
#find all stocks that exist in the S&P at this time period
investable_universe = []
for col in df_investable.columns:
if ~df_investable[col].iloc[:n].isna().any():
investable_universe.append(col)
df_investable = df_investable[investable_universe]
return df_investable
def apply_PCA(inv, k):
X = inv.iloc[1:, :]
pca = PCA(n_components = k)
inv_scaled = sklearn.preprocessing.StandardScaler().fit_transform(X)
principal_components = pca.fit_transform(inv_scaled)
df = pd.DataFrame(data = principal_components)
#For explained variance table
components = pca.components_
component_explained_var = pca.explained_variance_ratio_ * 100
comp_names = ['PCA' + str(i) for i in range(1, len(component_explained_var) + 1)]
pca_results = pd.DataFrame(data = component_explained_var, index = comp_names)
pca_results.columns = ['Explained variance (%)']
pca_results['Explained variance (%)'] = pca_results['Explained variance (%)'].round(2)
return df
def define_y(inv, stock):
y = inv[[stock]].iloc[:-1]
return y
def train_test(X, y):
X_train = X.iloc[1:, :]
X_test = X.iloc[0:1, :]
y_train = y.iloc[1:]
y_test = y.iloc[0:1]
return X_train, y_train, X_test, y_test
def model_fit(X_train, y_train):
model = svm.SVR()
model.fit(X_train, y_train)
return model
def model_predict(model, X_test):
yhat = model.predict(X_test)
return yhat
def predict_returns(t, n, k):
inv = get_investable(t, n)
X = apply_PCA(inv, k)
returns_t = pd.DataFrame(index = inv.columns, columns = ['Pred', 'Actual'])
for stock in inv.columns:
y = define_y(inv, stock)
X_train, y_train, X_test, y_test = train_test(X, y)
model = model_fit(X_train, y_train)
yhat = model_predict(model, X_test)[0]
returns_t['Pred'].loc[stock] = yhat
returns_t['Actual'].loc[stock] = y_test.values[0][0]
return returns_t
def rank_stocks(returns, num_stocks):
pred_returns = returns.sort_values(by = 'Pred', ascending = False)
topn = pred_returns.head(num_stocks)
botn = pred_returns.tail(num_stocks)
return topn, botn
def portfolio_return(topn, botn, returns):
return_t = topn['Actual'].mean() - botn['Actual'].mean()
return return_t
def pipeline(n, k, num_stocks):
time_range = returns.loc['2007':'2021'].index
portfolio = pd.DataFrame(index = time_range, columns = ['Portfolio Return'])
count = 0
for t in time_range[:-1]:
pred_actual = predict_returns(t, n, k)
topn, botn = rank_stocks(pred_actual, num_stocks)
return_t = portfolio_return(topn, botn, pred_actual)
t_index = time_range.get_loc(t) + 1
portfolio['Portfolio Return'].loc[time_range[t_index]] = return_t
count +=1
print(f'{(count/len(time_range))*100:.2f}% complete')
portfolio['Portfolio Return'] = portfolio['Portfolio Return'].astype('float')
return portfolio
portfolio = pipeline(200, 20, 5)
#started at 7:21am
print(pd.datetime.now())
portfolio
portfolio.plot()
# +
import matplotlib.ticker as ticker
fig, axes = plt.subplots(figsize=(30,15))
sns.barplot(x = portfolio.index, y = 'Portfolio Return', data = portfolio, color = 'grey')
axes.xaxis.set_major_locator(mdates.YearLocator())
axes.xaxis.set_minor_locator(mdates.MonthLocator())
ticklabels = [item.strftime('%Y') for item in portfolio.resample('Y').mean().index.to_period('Y')]
axes.xaxis.set_major_formatter(ticker.FixedFormatter(ticklabels))
plt.xticks(rotation = 'vertical')
axes.set_title('Portfolio Returns')
sns.set(font_scale=2)
plt.axhline(0)
plt.tight_layout()
plt.show()
# -
ticklabels = [item.strftime('%Y') for item in portfolio.resample('Y').mean().loc['2007':'2021'].index.to_period('Y')]
ticklabels
portfolio_monthly = portfolio.resample('M').mean()
portfolio_monthly
avg_return = portfolio['Portfolio Return'].mean()
print(f'Average return is {avg_return:.2f} %')
portfolio.dropna(inplace = True)
portfolio.to_csv('results/SVR.csv')
rolling_avg = pd.read_csv('results/SVR.csv', index_col = 'date', parse_dates = True)
rolling_avg.std()/(rolling_avg.shape[0]**0.5)
rolling_avg = pd.DataFrame(data = rolling_avg['Portfolio Return'].rolling(252).mean())
rolling_avg.dropna(inplace = True)
fig, axes = plt.subplots(figsize=(30,15))
sns.lineplot(x=rolling_avg.index, y="Portfolio Return", data=rolling_avg, color = '#0c8c84ff')
plt.xticks(rotation = 'vertical', fontsize = 25)
plt.yticks(fontsize = 20)
axes.set_title('SVR: 12-Month Rolling Average Portfolio Returns', fontsize = 35)
axes.set_xlabel('Year', fontsize = 30)
axes.set_ylabel('Portfolio Return (%)', fontsize = 30)
plt.xlim([rolling_avg.index[0], rolling_avg.index[-1]])
plt.axhline(0, color = '#fcc43cff')
| Integrated Model-SVR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
using LinearAlgebra
using SparseArrays
using DifferentialEquations
using Statistics
using Sundials
using DataStructures
using Plots
using Plots.PlotMeasures
using Distributions
include("MC_functions.jl")
# +
dict_of_solvers = OrderedDict{}()
dict_of_solvers["bd_stat "] = Dict();
dict_of_solvers["bd_stat "]["func"] = bd_stat
dict_of_solvers["bd_stat_mode "] = Dict()
dict_of_solvers["bd_stat_mode "]["func"] = bd_stat_mode
dict_of_solvers["bd_stat_log "] = Dict()
dict_of_solvers["bd_stat_log "]["func"]= bd_stat_log
dict_of_solvers["bd_stat_mode_log"] = Dict()
dict_of_solvers["bd_stat_mode_log"]["func"] = bd_stat_mode_log
# -
# # Infinite Server Queue Approximation
#
# We start with a small example. Let $\lambda =\mu =1$ and $s=S=10$. According to square-root staffing, this is about ten deviations out. Hence this is a good approximation to the infinite server model. We will want to see whether the stationary distribution solvers coincide to the known true distribution for the infinite server model.
p = Dict("s"=> 10, "S"=> 10, "sin_base" => 1, "sin_rel_amp" => 0, "sin_rate"=>0, "phase_shift" =>0, "mu"=>1)
Q = (Finite_Server_Q(0,p))'
for (key, value) in dict_of_solvers
func = value["func"]
print(key)
print(": ")
if func(Q) == -1
print("Fail")
else
print("Success")
value["stat"] = func(Q)
end
println()
end
# +
ref_statdist = pdf.(Poisson(),0:1:10)
for (key, value) in dict_of_solvers
print(key); print(": ")
if "stat" in keys(value)
print(maximum(abs.(value["stat"]-ref_statdist)))
println()
else
print("N/A")
println()
end
end
# -
# # US Callcenter Model
#
# In this notebook, we aim to see how well various solvers work on computing the stationary distribution for the US callcenter model. In this model, we have the following $\lambda(t)$ values.
#
t = 1:.1:1440
lambdas = US_Bank_Lambda(t)
plot(t,lambdas, label = "\\lambda (t)", title = "Arrival rate \\lambda (t) for US Bank Callcenter", xlabel="Time (min)", ylabel="Arrival rate \\lambda")
# As observed $\lambda(t)$ ranges from 0 to 100 customers per minute throughout the day. The mean service time is 3 minutes. We assume a square-root staffing model where for the $i$'th piecewise linear segment the staffer uses the average value of $\lambda(t)$, call it $\lambda_i$, to define $\rho_i$ and sets
#
# $$
# s_i = \rho_i + 2\sqrt{\rho_i}.
# $$
#
# Observe that during maximum load, we have that $\rho(t)\approx 300$ and hence, we choose a state space $ S =300 + 10\sqrt{300}\approx 470$.
# # Computing the stationary distribution at peak load
#
# We are interested in comparing different numerical methods for computing the stationary distribution at peak load. We will compare the following methods
#
# 1. Birth death standard (pi(i) = lambda_{i-1}/mu_i pi(i-1) with pi(0)=1).
# 2. Birth death starting at mode.
# 3. Birth death with log-exp.
# 4. Birth death starting at mode with log-exp.
#
# Let's first have a look and see if all methods even compute sensible numbers. Let's have a look at $t=600$ and $t=900$, which correspond to times of over-saturation and under-saturation respectively, both during peak arrival rates.
for (key, value) in dict_of_solvers
func = value["func"]
print(key)
print(": ")
if func(US_Bank_Q(600)) == -1
print("Fail")
else
print("Success")
value["stat_600"] = func(US_Bank_Q(600))
end
println()
end
# So only bd_stat_log failed, it seems like. Let's check to see if the other ones have similar values for the stationary distribution.
# +
ref_statdist = dict_of_solvers["bd_stat "]["stat_600"]
for (key, value) in dict_of_solvers
print(key); print(": ")
if "stat_600" in keys(value)
print(maximum(abs.(value["stat_600"]-ref_statdist)))
println()
else
print("N/A")
println()
end
end
# -
for (key, value) in dict_of_solvers
func = value["func"]
print(key)
print(": ")
if func(US_Bank_Q(7*60+.001)) == -1
print("Fail")
else
print("Success")
value["stat_600"] = func(US_Bank_Q(600))
end
println()
end
| files/research/.ipynb_checkpoints/Stationary_Distribution_Solvers-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/ContextLab/abstract2paper/blob/main/resources/abstract2paper_jupyter.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dAVD2y6e-iZO"
# # Welcome to *abstract2paper* ([Jupyter](https://jupyter.org/) version)!
# Author: [<NAME>](http://www.context-lab.com/)
#
# ## Step right up, step right up!
# <img src='https://media1.giphy.com/media/mL40PfXA394KA/giphy.gif' width='250px'>
#
# **Writing papers got you down?** Come on in, friend! Give my good ole' Abstract2papers Cure-All a quick try! Enter your abstract into the little doohicky here, and quicker'n you can blink your eyes<sup>1</sup>, a shiny new paper'll come right out for ya! What are you waiting for?
#
# ## How does it work, you ask?
#
# Really it's quite simple. We put in a smidgen of [this](https://huggingface.co/transformers/model_doc/gpt_neo.html) a pinch of [that](https://www.tug.org/texlive/), plus a dab of our special [*secret ingredient*](https://www.youtube.com/watch?v=dQw4w9WgXcQ), and **poof!** that's how the sausage is made.
#
# ## No really, how does it work?
#
# Ok, if you really want to know, all I'm doing here is using the [Hugging Face](https://huggingface.co/) [implementation](https://huggingface.co/transformers/model_doc/gpt_neo.html) of [GPT-Neo](https://github.com/EleutherAI/gpt-neo), which is itself a tweaked version of [GPT-3](https://arxiv.org/abs/2005.14165) that is pre-trained on the [Pile](https://pile.eleuther.ai/) dataset.
#
# The text you input is used as a prompt for GPT-Neo; to generate a document containing an additional *n* words, the model simply "predicts" the next *n* words that will come after the specified prompt.
#
# With a little help from some basic [LaTeX](https://www.latex-project.org/) templates (borrowed from [Overleaf](https://www.overleaf.com)), the document is formatted and compiled into a PDF.
#
# ## Can I actually use this in real-world applications?
#
# <img src='https://media4.giphy.com/media/3o6ozoD1ByqYv7ARIk/giphy.gif' width='250px'>
#
# **Doubtful.** Or at least, probably not...? It certainly wouldn't be ethical to use this code to generate writing assignments, mass-produce papers or grant applications, etc. Further, you'll likely find that the text produced using this approach includes stuff that's said in funny (often nonsensical) ways, follows problematic logic, incorporates biases from the training data, and so on. Of lesser importance, but practical annoyance, you'll also encounter all sorts of formatting issues (although those might be easy to fix manually, and possibly even automatically with some clever tinkering).
#
#
#
#
#
#
# <sup>1</sup><small>This claim rests on the assumption that you blink *really* slowly. Depending on how much text you're trying to generate (and how long your prompt is), your paper could take anywhere from a few minutes to several hours to fully congeal.</small>
# + [markdown] id="vQ52hti0G13T"
# # Step 1: Setting up the environment
#
# First we'll install Hugging Face's [transformers](https://huggingface.co/transformers/) library and download (and load into memory) the pre-trained GPT-Neo model. We'll also use a pretrained GPT-2 tokenizer for convenience.
#
# On RAM-constrained machines (e.g., [Google Colaboratory](https://colab.research.google.com/)), you may want to replace `'EleutherAI/gpt-neo-2.7B'` with `'EleutherAI/gpt-neo-1.3B'` in the second two lines in the cell below. That will load in a simpler (and smaller) model-- with 1.3 billion parameters instead of 2.7 billion parameters.
#
# There's a lot to download (roughly 6GB for the smaller model and 12.5GB for the larger model)-- it'll take a few minutes. Now would be a good time to track down that coffee refill you've been postponing...
# + id="fOzbwY-WHVZg"
try:
from transformers import GPTNeoForCausalLM, GPT2Tokenizer
except:
# !pip install transformers
from transformers import GPTNeoForCausalLM, GPT2Tokenizer
model = GPTNeoForCausalLM.from_pretrained("EleutherAI/gpt-neo-2.7B")
tokenizer = GPT2Tokenizer.from_pretrained("EleutherAI/gpt-neo-2.7B")
# + [markdown] id="qwQr2ziLIUyG"
# # Now I'm only going to ask this once: we're going to need a little...*information* from you...
# <img src='https://64.media.tumblr.com/e3c8dea30fdf2e597ce79904e8da3271/tumblr_o2xhqwU0cK1qmob6ro2_500.gif' width=400px>
#
# You didn't think it was going to be *that* easy, did you? Oh...you did? Well if you want *us* to cooperate, we're going to need a little...information...from you first. About your paper. Please make this easy on all of us and don't try to lie. The machine will know. The machine *always* knows...
#
# Fill in the information below and you'll be well on your way to your auto-generated paper/story/grant application/speech/business plan.
#
# + id="amOYPyewDLtE"
# credit: https://arxiv.org/abs/2005.14165
title = 'Language Models are Awesome-Shot Learners'
authors = '<NAME>, <NAME>, <NAME>, <NAME>,\\\\<NAME>, <NAME>, Arvind ' \
'Neelakantan, <NAME>,\\\\<NAME>, <NAME>, <NAME>, <NAME>,\\\\Gretchen ' \
'Krueger, <NAME>, <NAME>, <NAME>,\\\\<NAME>, <NAME>, <NAME>, ' \
'<NAME>,\\\\<NAME>, <NAME>, <NAME>, <NAME>,\\\\<NAME>, <NAME>, ' \
'<NAME>, <NAME>,\\\\<NAME>, <NAME>, and <NAME>'
text = "Recent work has demonstrated substantial gains on many NLP tasks and benchmarks by pre-training on a large " \
"corpus of text followed by fine-tuning on a specific task. While typically task-agnostic in architecture, " \
"this method still requires task-specific fine-tuning datasets of thousands or tens of thousands of examples. " \
"By contrast, humans can generally perform a new language task from only a few examples or from simple " \
"instructions - something which current NLP systems still largely struggle to do. Here we show that scaling up " \
"language models greatly improves task-agnostic, few-shot performance, sometimes even reaching competitiveness " \
"with prior state-of-the-art fine-tuning approaches. Specifically, we train GPT-3, an autoregressive language " \
"model with 175 billion parameters, 10x more than any previous non-sparse language model, and test its " \
"performance in the few-shot setting. For all tasks, GPT-3 is applied without any gradient updates or " \
"fine-tuning, with tasks and few-shot demonstrations specified purely via text interaction with the model. " \
"GPT-3 achieves strong performance on many NLP datasets, including translation, question-answering, " \
"and cloze tasks, as well as several tasks that require on-the-fly reasoning or domain adaptation, " \
"such as unscrambling words, using a novel word in a sentence, or performing 3-digit arithmetic. At the same " \
"time, we also identify some datasets where GPT-3's few-shot learning still struggles, as well as some " \
"datasets where GPT-3 faces methodological issues related to training on large web corpora. Finally, " \
"we find that GPT-3 can generate samples of news articles which human evaluators have difficulty " \
"distinguishing from articles written by humans. We discuss broader societal impacts of this finding and of " \
"GPT-3 in general."
length = 2500
# + [markdown] id="MiBDPVxLMQMT"
# The next cell is going to take a while to run. While you're waiting, just think: it's still better than writing something on your own, isn't it?
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="7rfrKf764v51" outputId="62d9ddc6-6cac-4567-b176-96ef3612f68d"
ids = tokenizer(text, return_tensors='pt')['input_ids']
tokens = model.generate(ids, do_sample=True, temperature=0.9, max_length=length)
gen_text = tokenizer.batch_decode(tokens)[0]
# + [markdown] id="fUfWHWA8ILSP"
# And finally, we'll get a tex-live installation going in our Colaboratory environment and download a template for generating the final document. (First we need to remove the model and tokenizer from RAM so that Colaboratory doesn't hit its memory limit and crash.)
# + colab={"background_save": true} id="Fgat6lZRCV1y"
def texer(template, outfile='auto.tex', **kwargs):
with open(template, 'r') as f:
lines = f.readlines()
x = []
for line in lines:
for k, v in kwargs.items():
line = line.replace(f'<{k}>', str(v))
x.append(line.replace(' & ', ' \& ').replace('%', '\%'))
try:
with open(outfile, 'w+') as f:
f.write('\n'.join(x))
os.system(f'pdflatex {outfile}')
os.system('rm *.log *.aux')
except:
pass
return '\n'.join(x)
# + [markdown] id="OZPYsfDDQ-q2"
# Create a PDF containing your auto-generated document...
# + colab={"background_save": true} id="73tadb3xQD_t"
source = texer(os.path.join(os.getenv('HOME'), 'abstract2paper', 'resources', 'template.tex'), TITLE=title, AUTHOR=authors+'\\\\Augmented by GPT-Neo', GEN_TEXT=gen_text + '...')
# + colab={"background_save": true} id="XRzAVkyOHA0S" outputId="3744de74-9869-47ce-bf00-2abbce152c2a"
import pprint
pprint.pprint(source, width=80)
# -
try:
import fakepackage1234
except:
# !pip install pandas
import pandas as pd
| resources/abstract2paper_jupyter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="zrW2XUolsTgx"
# # Introduction to LaTeX
#
# In this tutorial, you will learn some of the basics on how to use $\LaTeX$ to display equations in Jupyter notebooks. For looking up symbols you need to search on the internet.
# + [markdown] id="KGsd65eesTg7"
# ## Basic inline LaTeX
#
# To embed LaTeX within text, simply encapsulate the LaTeX portions in dollar signs (`$`). MathJax takes care of the rest. As an example, consider the sentence below and the markdown/LaTeX code to render it.
#
# Einstein told us that $E = mc^2$.
#
# Einstein told us that $E = mc^2$.
#
# Notice how the equation is properly rendered, with mathematical variables in italics. Note also how `^2` was used to raise to a power. If the power has more than one character in it, it should be enclosed in braces (`{}`). In fact, braces are used to generally group symbols in LaTeX.
#
# Euler told us that $\mathrm{e}^{i \pi} - 1 = 0$.
#
# Euler told us that $\mathrm{e}^{i \pi} - 1 = 0$.
#
# Aside from the grouping braces, there are several other syntactical items of note. First, notice that I made the special character $\pi$ with `\pi`. In general, a backward slash precedes special symbols or commands in LaTeX. If we want another Greek letter, like $\theta$, we use `\theta`. Now, also note that I used "`\mathrm{e}`" for the base of the natural logarithm. I was signaling to LaTeX that I wanted the character written in Roman font, and not italics, so I used `\mathrm`. Anything in the braces following the function `\mathrm` is rendered in Roman font. Note the difference.
#
# This is $e$. This is $\mathrm{e}$
#
# This is $e$. This is $\mathrm{e}$.
#
# Now, back to grouping things in braces. We can do similar groupings using braces with with subscripts.
#
# The dot product of two $n$-vectors is $\mathbf{a} \cdot \mathbf{b} = \sum_{i=1}^n a_i b_i$.
#
# The dot product of two $n$-vectors is $\mathbf{a} \cdot \mathbf{b} = \sum_{i=1}^n a_i b_i$.
#
# Here, I have used `$\mathbf{a}$` to make the character `a` boldface, denoting a vector. Note that we denote subscripts with an underscore. Notice also that the bounds of the sum use the same underscore and caret notation as for subscripts and superscripts.
# + [markdown] id="x6CZHmVFsTg9"
# ## Displaying equations on separate lines
#
# The bounds on the summation in the above example may look a little funny to you because they are not above and below the summation symbol. This is because this particular equation is written inline. If we had separated it from the text, it renders differently.
#
# We can make an equation appear centered on a new line, like
#
# \begin{align}
# \mathbf{a} \cdot \mathbf{b} = \sum_{i=1}^n a_i b_i.
# \end{align}
#
#
# We can make an equation appear centered on a new line, like
#
# \begin{align}
# \mathbf{a} \cdot \mathbf{b} = \sum_{i=1}^n a_i b_i.
# \end{align}
#
# The `align` environment in LaTeX specifies that you want centered equations, separated from the text. It is called `align` because it allows you to align the equations. You separate lines in the equations with a double backslash (`//`). Insert an ampersand (`&`) in each line at the alignment point. All equations will be aligned at the location of the ampersand symbols (and, of course, the ampersands will not appear in the rendered equations).
#
# For a three-vector consisting of $x$, $y$, and $z$ components,
#
# \begin{align}
# \mathbf{a} \cdot \mathbf{b} &= \sum_{i=1}^n a_i b_i \\
# &= a_x b_x + a_y b_y + a_z b_z.
# \end{align}
#
# For a three-vector consisting of $x$, $y$, and $z$ components,
#
# \begin{align}
# \mathbf{a} \cdot \mathbf{b} &= \sum_{i=1}^n a_i b_i \\
# &= a_x b_x + a_y b_y + a_z b_z.
# \end{align}
#
# Note that I always put an extra blank line before the `\begin{align}` statement. This is not necessary, but I think things look better with the extra space.
# + [markdown] id="Xk3Dho2CsThA"
# ## Fractions (and an example of fine-tuning)
#
# To display fractional quantities, we use the `\frac{}{}` command. `\frac` is always followed by two sets of braces; the numerator is contained in the first, and the denominator is contained in the second. As an example, we can write an equation you will become intimately familiar with if you take the second term of this course,
#
# \begin{align}
# P(A \mid B) = \frac{P(B \mid A) \, P(A)}{P(B)}
# \end{align}
#
# \begin{align}
# P(A \mid B) = \frac{P(B \mid A) \, P(A)}{P(B)}
# \end{align}
#
# The right hand side has a nicely-formatted fraction. I did a little extra fine-tuning in this equation. I'll show the equation again without the fine-tuning, which used the `\mid` and `\,` commands.
#
# \begin{align}
# P(A | B) = \frac{P(B | A) P(A)}{P(B)}.
# \end{align}
#
# \begin{align}
# P(A | B) = \frac{P(B | A) P(A)}{P(B)}.
# \end{align}
#
# First, the `\mid` command should be used in conditional probabilities. Just using a vertical bar (`|`) results in crowding. Similarly, I used the `\,` command to insert a little extra space between the two probabilities in the numerator. This makes the equation a bit easier to read. This `\,` operator is especially important when defining integrals. We can put a little space between the $\mathrm{d}x$ and the integrand.
#
# \begin{align}
# \text{good: } &\int_0^{2\pi} \mathrm{d}x \, \sin x. \\[1em]
# \text{bad: } &\int_0^{2\pi} \mathrm{d}x \sin x.
# \end{align}
#
# \begin{align}
# \text{good: } &\int_0^{2\pi} \mathrm{d}x \, \sin x. \\[1em]
# \text{bad: } &\int_0^{2\pi} \mathrm{d}x \sin x.
# \end{align}
#
# Note that I inserted extra space after the new line. Specifically, `\\[1em]` instructs LaTeX to insert a space equation to the width of an M character between the equations. I often do this to keep things clear.
#
# It is also very important to note that I used $\sin$ and not $sin$. Mathematical functions should be in Roman font and are invoked with a backslash. Otherwise, the characters are interpreted as separate variables. To be clear:
#
# \begin{align}
# \text{good: } &\sin x. \\[1em]
# \text{bad: } & sin x.
# \end{align}
#
# \begin{align}
# \text{good: } &\sin x. \\[1em]
# \text{bad: } & sin x.
# \end{align}
#
# Finally, notice that I was able to put text in the equation like this: `\text{good: }`.
# + [markdown] id="c9qr8LllsThC"
# ## Grouping operators (and more fine-tuning)
#
# Compare the following equations.
#
# \begin{align}
# \text{good: } &\sum_{i=1}^n i^3 = \left(\sum_{i=1}^n i\right)^2. \\[1em]
# \text{bad: } &\sum_{i=1}^n i^3 = (\sum_{i=1}^n i)^2.
# \end{align}
#
# \begin{align}
# \text{good: } &\sum_{i=1}^n i^3 = \left(\sum_{i=1}^n i\right)^2. \\[1em]
# \text{bad: } &\sum_{i=1}^n i^3 = (\sum_{i=1}^n i)^2.
# \end{align}
#
# In the second equation, I did not use the `\left(` and `\right)` construction for parentheses and the result looks pretty awful. In LaTeX, the height of anything that is encapsulated by `\left(` and `\right)` scales the parentheses appropriately. You can use `\left` and `\right` with many symbols. An important example is `\left\{`. Note that to display braces in an equation, you have to use `\{` because just a plain brace (`{`) has a different meaning.
#
# (By the way, that equation is true, and pretty amazing. It says that the sum of the first $n$ *cubes* of integers is equal to the sum of the first $n$ integers *squared*!)
#
# Finally, if you use `\left.` or `\right.`, LaTeX will simply scale the opposite symbol to match the height of the text, but will suppress printing the other. For example,
#
# \begin{align}
# \left. \frac{1}{x + 2} \right|_0^2 = -\frac{1}{4}.
# \end{align}
#
# \begin{align}
# \left. \frac{1}{x + 2} \right|_0^2 = -\frac{1}{4}.
# \end{align}
#
# This is also useful if you are going to use `/` for a division operation. Compare the following.
#
# \begin{align}
# \text{good: } & \left. x^2 \middle/ y^2 \right. \\[1em]
# \text{bad: } & x^2 / y^2
# \end{align}
#
# \begin{align}
# \text{good: } & \left. x^2 \middle/ y^2 \right. \\[1em]
# \text{bad: } & x^2 / y^2
# \end{align}
#
# Here, we used the `\middle` operator to scale the length of the division sign.
# + [markdown] id="IM4QCtaPsThE"
# ## Matrices and arrays
# On occasion, you'll need to express matrices. This is most easily done using the `pmatrix` environment. For example, a covariance matrix for two variables might be written as
#
# \begin{align}
# \sigma^2 = \begin{pmatrix}
# \sigma_1^2 & \sigma_{12} \\
# \sigma_{12} & \sigma_2^2
# \end{pmatrix}.
# \end{align}
#
# \begin{align}
# \sigma^2 = \begin{pmatrix}
# \sigma_1^2 & \sigma_{12} \\
# \sigma_{12} & \sigma_2^2
# \end{pmatrix}.
# \end{align}
#
# Once in the `pmatrix` environment, each row has entries separated by an ampersand. The row ends with a `\\`. Each row must have the same number of entries.
#
# You may also need to represent an values stacked on top of each other. For example, we might specify a piecewise linear function like this.
#
# \begin{align}
# \text{rectifier}(x) = \left\{
# \begin{array}{cl}
# 0 & x \le 0 \\
# x & x > 0.
# \end{array}
# \right.
# \end{align}
#
# \begin{align}
# \text{rectifier}(x) = \left\{
# \begin{array}{cl}
# 0 & x \le 0 \\
# x & x > 0.
# \end{array}
# \right.
# \end{align}
#
# The `array` environment allows arrays of text. The `{cl}` after `\begin{array}` indicates that two columns are wanted, with the first column being centered and the second being left-aligned. If we chose instead `{lr}`, the first column is left-aligned and the second column is right-aligned.
# + [markdown] id="-1nZOmyAsThG"
# ## Useful LaTeX symbols for BE/Bi 103
# Following is a list of some symbols you may find useful in this class.
#
# LaTeX | symbol
# --- | :---:
# `\approx` | $\approx$
# `\sim` | $\sim$
# `\propto` | $\propto$
# `\le` | $\le$
# `ge` | $\ge$
# `\pm` | $\pm$
# `\in` | $\in$
# `\ln` | $\ln$
# `\exp` | $\exp$
# `\prod_{i\in D}` | ${\displaystyle \prod_{i\in D}}$
# `\sum_{i\in D}` | ${\displaystyle \sum_{i\in D}}$
# `\frac{\partial f}{\partial x}` | ${\displaystyle \frac{\partial f}{\partial x}}$
# `\sqrt{x}` | $\sqrt{x}$
# `\bar{x}` | $\bar{x}$
# `\langle x \rangle` | $\langle x \rangle$
# `\left\langle \frac{x}{y} \right\rangle` | $\left\langle \frac{x}{y} \right\rangle$
| Introduction_to_LaTeX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#GA con ajuste polinomial de grado 10 por cada angulo. 4x10-> 40 genes reales.
#Inicio aleatoria entre -20 y 20
from __future__ import division
from ThePablos import *
import random
from deap import base, creator, tools, algorithms
import time
#FUNCIONES AUXILIARES
def checkBoundis(individual):
global cotainf
global cotamax
thetaAux = getPathFromInd(individual) - condborde.reshape((4,1))
cotainfAux = cotainf - condborde
cotamaxAux = cotamax - condborde
for i in range(4):
factor = 1
for k in range(Npos):
if thetaAux[i,k] > cotamaxAux[i]:
factorTemp = cotamaxAux[i]/thetaAux[i,k]
if factorTemp < factor: factor = factorTemp
elif thetaAux[i,k] < cotainfAux[i]:
factorTemp = cotainfAux[i]/thetaAux[i,k]
if factorTemp < factor: factor = factorTemp
individual[ i*n_coef:(i*n_coef+Npos)] = individual[i*n_coef:(i*n_coef+Npos)] *factor
return individual
def kernel(X,Y,sigma):
return np.exp( -sum((X-Y)**2) / (2*sigma**2) )
#CONVERTIR INDIVIDUO A TRAYECTORIA
def getPathFromInd(individual):
global Npos
global t_mov
global condborde
global n_coef
t1 = np.ones(Npos)*condborde[0]
t2 = np.ones(Npos)*condborde[1]
t3 = np.ones(Npos)*condborde[2]
t4 = np.ones(Npos)*condborde[3]
theta = np.vstack([t1,t2,t3,t4])
for i in range(4):
for k in range(Npos):
for j in range(0,n_coef):
theta[i,k] += individual[i*n_coef+j]*((k*t_mov/Npos)**(j+1))
return theta
#Calculo del fitness a partir del path
def evaluatePath(theta):
global finalpos
global robotArm
global lam
global en_scale
global sigmaEnergy
robotArm.setPath(theta[0], theta[1], theta[2], theta[3])
robotArm.angularCons()
finalGenerado = robotArm.endPoint()
energiac = robotArm.getEnergySim()
#fitness1 = mse(finalGenerado, finalpos)
#fitness1 = 1/(1+mse(finalGenerado, finalpos))
fitness1 = kernel(finalGenerado, finalpos,5)
#fitness2 = energiac
#fitness2 = 1/(1+energiac)
fitness2 = kernel(np.array([energiac]), np.zeros(1),sigmaEnergy)
fitness = (1-lam)*fitness1 + lam*fitness2
return fitness,
#EVALUAR INDIVIDUO
def evaluate(individual):
theta = getPathFromInd(individual)
return evaluatePath(theta)
def kernel(X,Y,sigma):
k = np.exp(-sum((X-Y)**2) / (2*(sigma**2)))
return k
def mse(x,y):
return sum(abs((x-y)**2))
####---------------------- PARAMETROS ---------------------- #####
#Arreglos para iterar
lamArray = [0.5, 0.1, 0.01, 0.001]
#PmcArray = [0.4, 0.3, 0.2, 0.1]
sigmaEnergyArray = [50, 100, 150, 200]
error = np.zeros(5)
energia = np.zeros(5)
n_coef = 10 #ind_size = 4*n_coef, grado 10
#Probabilidades de recombinacion y mutacion
Pci = 0.9 #cada individuo
Pcc = 0.9 #cada atributo
Pmi = 0.5
Pmc = 0.1 #comentar para sin correntropia
cotainf = np.array([-180,-90,-90,-90])*np.pi/180
cotamax = np.array([180,90,90,90])*np.pi/180
l1 = 1
l2 = 1
l3 = 1
l4 = 1
r = 1e-2
#Posiciones en que se discretizan los caminos
Npos = 20
#Numero de generaciones
Ng = 70
#Numero de individuos en la poblacion
Npop = 300
#Duracion del movimiento
t_mov = 1
#configuracion inicial de angulos
condborde=np.array([90, -30, 30, -20])*np.pi/180
#Posicion target
#x,y,z plano cartesiano
#finalpos=np.array([1.93, 1.93, 1.0])
finalpos = np.array([1.0, 1.0, 2.0])
#finalpos=np.array([0, 0, 0])
#Crossover y MUt
eta_cx = 2 #SimulatedBinary
alpha_cx = 0.5 #Blend BLX-0.5
sigmaMut = 0.1
#Simulador de robot
robotArm = ThePablos(l1, l2, l3, l4, r, t_mov)
#--------------------ITERACION--------------#
for a in range(len(lamArray)):
#for b in range(len(PmcArray)):
for b in range(len(sigmaEnergyArray)):
#Para promediar casos
for c in range(4):
#Parametros de esta iteracion
lam = lamArray[a]
#Pmc = PmcArray[b]
sigmaEnergy = sigmaEnergyArray[b]
#Registro de fitness
creator.create("FitnessMulti", base.Fitness, weights=(1.0,)) #maximizar
#creator.create("FitnessMulti", base.Fitness, weights=(-1.0,)) #minimizar
creator.create("Individual", np.ndarray, fitness=creator.FitnessMulti)
#Asignacion de funciones para crear individuos
toolbox = base.Toolbox()
toolbox.register("attr_real", random.uniform, 0, 1)
toolbox.register("individual", tools.initRepeat, creator.Individual, toolbox.attr_real, n=4*n_coef)
#Un individuo es un arreglo de coeficientes
#Asignacion de funciones para crear la poblacion
toolbox.register("population", tools.initRepeat, list, toolbox.individual)
#Asignacion de operadores
toolbox.register("evaluate", evaluate)
#toolbox.register("mate", tools.cxBlend, alpha=alpha_cx)
toolbox.register("mate", tools.cxSimulatedBinary, eta=eta_cx)
toolbox.register("mutate", tools.mutGaussian, mu=0, sigma=sigmaMut, indpb=Pmc)
toolbox.register("select", tools.selTournament, tournsize=3)
toolbox.register("checki", checkBoundis)
####---------------------- ALGORITMO GENETICO ----------------------- #####
def main():
pop = toolbox.population(n=Npop)
for ind in pop:
toolbox.checki(ind)
print("Start of evolution %i" % c)
# Evaluate the entire population
fitnesses = list(map(toolbox.evaluate, pop))
for ind, fit in zip(pop, fitnesses):
ind.fitness.values = fit
# Begin the evolution
for g in range(Ng):
# Select the next generation individuals
offspring = toolbox.select(pop, len(pop))
# Clone the selected individuals
offspring = list(map(toolbox.clone, offspring))
besties = tools.selBest(pop, 1)
best1 = besties[0]
# Apply crossover and mutation on the offspring
for child1, child2 in zip(offspring[::2], offspring[1::2]):
# cross two individuals with probability CXPB
if random.random() < Pci:
toolbox.mate(child1, child2)
# fitness values of the children
# must be recalculated later
toolbox.checki(child1)
toolbox.checki(child1)
del child1.fitness.values
del child2.fitness.values
for mutant in offspring:
# mutate an individual with probability MUTPB
if random.random() < Pmi:
toolbox.mutate(mutant)
toolbox.checki(mutant)
del mutant.fitness.values
# Evaluate the individuals with an invalid fitness
invalid_ind = [ind for ind in offspring if not ind.fitness.valid]
fitnesses = map(toolbox.evaluate, invalid_ind)
for ind, fit in zip(invalid_ind, fitnesses):
ind.fitness.values = fit
# The population is entirely replaced by the offspring
pop[:] = offspring
indaux = np.random.randint(0, len(pop))
pop[indaux] = best1 #Elitismo
best_ind = tools.selBest(pop, 1)[0]
return best_ind
####---------------------- VISUALIZACION DE RESULTADOS ----------------------- #####
if __name__ == "__main__":
#best, max1, max2, min1, min2, prom1, prom2 = main()
best = main()
tiempo = np.linspace(0.0,t_mov,num=Npos)
thetaWin = getPathFromInd(best)
thetaWinA = thetaWin[0]
thetaWinB = thetaWin[1]
thetaWinC = thetaWin[2]
thetaWinD = thetaWin[3]
robotArm.setPath(thetaWinA, thetaWinB, thetaWinC, thetaWinD)
robotArm.angularCons()
finalGenerado = robotArm.endPoint()
error[c] = mse(finalGenerado,finalpos)
energia[c] = robotArm.getEnergySim()
promedioError = sum(error)/len(error)
stdError = np.std(error)
promedioEnergia = sum(energia)/len(energia)
stdEnergia = np.std(energia)
file = open("TablaPoliCorr.txt", "a")
linea = "lam "+str(lam)+" sigmaEnergy "+str(sigmaEnergy)+" error "+str(promedioError)+"std"+str(stdError)+" energia "+str(promedioEnergia)+"std"+str(stdEnergia)+" \n"
file.write(linea)
file.close()
# +
a = 2e-3
print a
file = open("newfile.txt", "a")
file.write(str(a)+" "+str(a)+"\n")
file.write("And here is another line\n")
file.close()
| Implementacion GA polinomial - Iterativo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:aparent]
# language: python
# name: conda-env-aparent-py
# ---
# +
import pandas as pd
import scipy
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import pickle
import os
# +
data = pd.read_csv('unprocessed_data/Alt_5SS_Tag_to_Seq_Map.csv',sep=',',index_col=0)
c = spio.loadmat('unprocessed_data/Alt_5SS_Usage_All_Cells.mat')
c_MCF7 = sp.csc_matrix(c['MCF7'])
c_CHO = sp.csc_matrix(c['CHO'])
c_HELA = sp.csc_matrix(c['HELA'])
c_HEK = sp.csc_matrix(c['HEK'])
# +
#Sort data on counts
total_c_MCF7 = np.ravel(c_MCF7.sum(axis=-1))
total_c_CHO = np.ravel(c_CHO.sum(axis=-1))
total_c_HELA = np.ravel(c_HELA.sum(axis=-1))
total_c_HEK = np.ravel(c_HEK.sum(axis=-1))
avg_c = (total_c_HEK + total_c_HELA + total_c_CHO + total_c_MCF7) / 4.0
sort_index = np.argsort(avg_c)
data = data.iloc[sort_index].copy().reset_index(drop=True)
c_MCF7 = c_MCF7[sort_index, :]
c_CHO = c_CHO[sort_index, :]
c_HELA = c_HELA[sort_index, :]
c_HEK = c_HEK[sort_index, :]
# +
#Constant background sequence context
up_background = 'gggcatcgacttcaaggaggacggcaacatcctggggcacaagctggagtacaactacaacagccacaacgtctatatcatggccgacaagcagaagaacggcatcaaagtgaacttcaagatccgccacaacatcgagg'.upper()
dn_background = 'acagagtttccttatttgtctctgttgccggcttatatggacaagcatatcacagccatttatcggagcgcctccgtacacgctattatcggacgcctcgcgagatcaatacgtatacca'.upper()
print('len(up_background) = ' + str(len(up_background)))
print('len(dn_background) = ' + str(len(dn_background)))
# +
#Extend sequences and count matrices
data['Padded_Seq'] = up_background + data['Seq'].str.slice(0,101) + dn_background
padded_c_MCF7, padded_c_CHO, padded_c_HELA, padded_c_HEK = [
sp.csr_matrix(
sp.hstack([
sp.csc_matrix((c_mat.shape[0], len(up_background))),
c_mat[:, :101],
sp.csc_matrix((c_mat.shape[0], len(dn_background))),
sp.csc_matrix(np.array(c_mat[:, 303].todense()).reshape(-1, 1))
])
)
for c_mat in [c_MCF7, c_CHO, c_HELA, c_HEK]
]
print('padded_c_MCF7.shape = ' + str(padded_c_MCF7.shape))
print('padded_c_CHO.shape = ' + str(padded_c_CHO.shape))
print('padded_c_HELA.shape = ' + str(padded_c_HELA.shape))
print('padded_c_HEK.shape = ' + str(padded_c_HEK.shape))
# +
#Filter each dataset on > 0 count
hek_keep_index = np.nonzero(np.ravel(padded_c_HEK.sum(axis=-1)) > 0)[0]
hela_keep_index = np.nonzero(np.ravel(padded_c_HELA.sum(axis=-1)) > 0)[0]
mcf7_keep_index = np.nonzero(np.ravel(padded_c_MCF7.sum(axis=-1)) > 0)[0]
cho_keep_index = np.nonzero(np.ravel(padded_c_CHO.sum(axis=-1)) > 0)[0]
#HEK data
data_hek_filtered = data.iloc[hek_keep_index].copy().reset_index(drop=True)
c_hek_filtered = padded_c_HEK[hek_keep_index, :]
#HELA data
data_hela_filtered = data.iloc[hela_keep_index].copy().reset_index(drop=True)
c_hela_filtered = padded_c_HELA[hela_keep_index, :]
#MCF7 data
data_mcf7_filtered = data.iloc[mcf7_keep_index].copy().reset_index(drop=True)
c_mcf7_filtered = padded_c_MCF7[mcf7_keep_index, :]
#CHO data
data_cho_filtered = data.iloc[cho_keep_index].copy().reset_index(drop=True)
c_cho_filtered = padded_c_CHO[cho_keep_index, :]
print('len(data_hek_filtered) = ' + str(len(data_hek_filtered)))
print('c_hek_filtered.shape = ' + str(c_hek_filtered.shape))
print('len(data_hela_filtered) = ' + str(len(data_hela_filtered)))
print('c_hela_filtered.shape = ' + str(c_hela_filtered.shape))
print('len(data_mcf7_filtered) = ' + str(len(data_mcf7_filtered)))
print('c_mcf7_filtered.shape = ' + str(c_mcf7_filtered.shape))
print('len(data_cho_filtered) = ' + str(len(data_cho_filtered)))
print('c_cho_filtered.shape = ' + str(c_cho_filtered.shape))
# +
#Get joined min dataset
min_keep_index = (np.ravel(padded_c_HEK.sum(axis=-1)) > 0)
min_keep_index = min_keep_index & (np.ravel(padded_c_HELA.sum(axis=-1)) > 0)
min_keep_index = min_keep_index & (np.ravel(padded_c_MCF7.sum(axis=-1)) > 0)
min_keep_index = min_keep_index & (np.ravel(padded_c_CHO.sum(axis=-1)) > 0)
#MIN data
data_min_filtered = data.iloc[min_keep_index].copy().reset_index(drop=True)
c_hek_min_filtered = padded_c_HEK[min_keep_index, :]
c_hela_min_filtered = padded_c_HELA[min_keep_index, :]
c_mcf7_min_filtered = padded_c_MCF7[min_keep_index, :]
c_cho_min_filtered = padded_c_CHO[min_keep_index, :]
print('len(data_min_filtered) = ' + str(len(data_min_filtered)))
print('c_hek_min_filtered.shape = ' + str(c_hek_min_filtered.shape))
print('c_hela_min_filtered.shape = ' + str(c_hela_min_filtered.shape))
print('c_mcf7_min_filtered.shape = ' + str(c_mcf7_min_filtered.shape))
print('c_cho_min_filtered.shape = ' + str(c_cho_min_filtered.shape))
# +
#Pickle final datasets
data_min_filtered = data_min_filtered.rename(columns={'Padded_Seq' : 'padded_seq'})
data_hek_filtered = data_hek_filtered.rename(columns={'Padded_Seq' : 'padded_seq'})
data_hela_filtered = data_hela_filtered.rename(columns={'Padded_Seq' : 'padded_seq'})
data_mcf7_filtered = data_mcf7_filtered.rename(columns={'Padded_Seq' : 'padded_seq'})
data_cho_filtered = data_cho_filtered.rename(columns={'Padded_Seq' : 'padded_seq'})
data_min_filtered = data_min_filtered[['padded_seq']]
data_hek_filtered = data_hek_filtered[['padded_seq']]
data_hela_filtered = data_hela_filtered[['padded_seq']]
data_mcf7_filtered = data_mcf7_filtered[['padded_seq']]
data_cho_filtered = data_cho_filtered[['padded_seq']]
splicing_5ss_dict = {
'min_df' : data_min_filtered,
'hek_df' : data_hek_filtered,
'hela_df' : data_hela_filtered,
'mcf7_df' : data_mcf7_filtered,
'cho_df' : data_cho_filtered,
'hek_count' : c_hek_filtered,
'hela_count' : c_hela_filtered,
'mcf7_count' : c_mcf7_filtered,
'cho_count' : c_cho_filtered,
'min_hek_count' : c_hek_min_filtered,
'min_hela_count' : c_hela_min_filtered,
'min_mcf7_count' : c_mcf7_min_filtered,
'min_cho_count' : c_cho_min_filtered,
}
pickle.dump(splicing_5ss_dict, open('alt_5ss_data.pickle', 'wb'))
# +
#Align and consolidate a5ss data
plasmid_dict = pickle.load(open('alt_5ss_data.pickle', 'rb'))
plasmid_df = plasmid_dict['min_df']
hek_cuts = np.array(plasmid_dict['min_hek_count'].todense())
hela_cuts = np.array(plasmid_dict['min_hela_count'].todense())
mcf7_cuts = np.array(plasmid_dict['min_mcf7_count'].todense())
cho_cuts = np.array(plasmid_dict['min_cho_count'].todense())
total_cuts = hek_cuts + hela_cuts + mcf7_cuts + cho_cuts
total_cuts = total_cuts[:, :-1]
# +
fixed_poses = [140, 140 + 44, 140 + 79]
sd_window = 130#120
sd1_pos = 140
negative_sampling_ratio = 2
fixed_pos_mask = np.ones(total_cuts.shape[1])
for j in range(len(fixed_poses)) :
fixed_pos_mask[fixed_poses[j]] = 0
cut_pos = np.arange(total_cuts.shape[1])
aligned_seqs = []
aligned_libs = []
aligned_mode = []
max_data_len = 3000000
aligned_hek_cuts = sp.lil_matrix((max_data_len, 2 * sd_window + 1))
aligned_hela_cuts = sp.lil_matrix((max_data_len, 2 * sd_window + 1))
aligned_mcf7_cuts = sp.lil_matrix((max_data_len, 2 * sd_window + 1))
aligned_cho_cuts = sp.lil_matrix((max_data_len, 2 * sd_window + 1))
splice_mats = [
[hek_cuts, aligned_hek_cuts],
[hela_cuts, aligned_hela_cuts],
[mcf7_cuts, aligned_mcf7_cuts],
[cho_cuts, aligned_cho_cuts]
]
old_i = 0
new_i = 0
for _, row in plasmid_df.iterrows() :
if old_i % 10000 == 0 :
print("Processing sequence " + str(old_i) + "...")
seq = row['padded_seq']
nonzero_cuts = np.nonzero( ((total_cuts[old_i, :] > 0) & (fixed_pos_mask == 1)) & ((cut_pos >= sd_window) & (cut_pos < total_cuts.shape[1] - sd_window)) )[0].tolist()
zero_cuts = np.nonzero( ((total_cuts[old_i, :] == 0) & (fixed_pos_mask == 1)) & ((cut_pos >= sd_window + 1) & (cut_pos < total_cuts.shape[1] - sd_window - 1)) )[0].tolist()
#Emit fixed splice positions
for fixed_pos in fixed_poses :
aligned_seqs.append(seq[fixed_pos - sd_window: fixed_pos + sd_window])
aligned_libs.append(fixed_pos - sd1_pos)
aligned_mode.append("fixed_" + str(fixed_pos - sd1_pos))
for [cuts, aligned_cuts] in splice_mats :
aligned_cuts[new_i, :2 * sd_window] = cuts[old_i, fixed_pos - sd_window: fixed_pos + sd_window]
aligned_cuts[new_i, 2 * sd_window] = cuts[old_i, -1]
new_i += 1
#Emit denovo splice positions
for denovo_pos in nonzero_cuts :
aligned_seqs.append(seq[denovo_pos - sd_window: denovo_pos + sd_window])
aligned_libs.append(denovo_pos - sd1_pos)
aligned_mode.append("denovo_pos_" + str(denovo_pos - sd1_pos))
for [cuts, aligned_cuts] in splice_mats :
aligned_cuts[new_i, :2 * sd_window] = cuts[old_i, denovo_pos - sd_window: denovo_pos + sd_window]
aligned_cuts[new_i, 2 * sd_window] = cuts[old_i, -1]
new_i += 1
if negative_sampling_ratio > 0.0 :
n_neg = int(negative_sampling_ratio * (3 + len(nonzero_cuts)))
sampled_zero_cuts = np.random.choice(zero_cuts, size=n_neg, replace=False)
#Emit negative denovo splice positions
for denovo_pos in sampled_zero_cuts :
aligned_seqs.append(seq[denovo_pos - sd_window: denovo_pos + sd_window])
aligned_libs.append(denovo_pos - sd1_pos)
aligned_mode.append("denovo_neg_" + str(denovo_pos - sd1_pos))
for [cuts, aligned_cuts] in splice_mats :
aligned_cuts[new_i, :2 * sd_window] = cuts[old_i, denovo_pos - sd_window: denovo_pos + sd_window]
aligned_cuts[new_i, 2 * sd_window] = cuts[old_i, -1]
new_i += 1
old_i += 1
aligned_min_hek_cuts = sp.csr_matrix(aligned_hek_cuts[:len(aligned_seqs), :])
aligned_min_hela_cuts = sp.csr_matrix(aligned_hela_cuts[:len(aligned_seqs), :])
aligned_min_mcf7_cuts = sp.csr_matrix(aligned_mcf7_cuts[:len(aligned_seqs), :])
aligned_min_cho_cuts = sp.csr_matrix(aligned_cho_cuts[:len(aligned_seqs), :])
aligned_min_df = pd.DataFrame({
'seq' : aligned_seqs,
'library' : aligned_libs,
'origin' : aligned_mode
})
aligned_min_df = aligned_min_df[['seq', 'library', 'origin']]
print("len(aligned_min_df) = " + str(len(aligned_min_df)))
print("aligned_min_hek_cuts.shape = " + str(aligned_min_hek_cuts.shape))
print("aligned_min_hela_cuts.shape = " + str(aligned_min_hela_cuts.shape))
print("aligned_min_mcf7_cuts.shape = " + str(aligned_min_mcf7_cuts.shape))
print("aligned_min_cho_cuts.shape = " + str(aligned_min_cho_cuts.shape))
# +
#Filter out zeros
keep_index = (np.ravel(aligned_min_hek_cuts.sum(axis=-1)) > 0)
keep_index = keep_index & (np.ravel(aligned_min_hela_cuts.sum(axis=-1)) > 0)
keep_index = keep_index & (np.ravel(aligned_min_mcf7_cuts.sum(axis=-1)) > 0)
keep_index = keep_index & (np.ravel(aligned_min_cho_cuts.sum(axis=-1)) > 0)
aligned_min_df = aligned_min_df.iloc[keep_index].copy().reset_index(drop=True)
aligned_min_hek_cuts = aligned_min_hek_cuts[keep_index, :]
aligned_min_hela_cuts = aligned_min_hela_cuts[keep_index, :]
aligned_min_mcf7_cuts = aligned_min_mcf7_cuts[keep_index, :]
aligned_min_cho_cuts = aligned_min_cho_cuts[keep_index, :]
print("len(aligned_min_df) = " + str(len(aligned_min_df)))
print("aligned_min_hek_cuts.shape = " + str(aligned_min_hek_cuts.shape))
print("aligned_min_hela_cuts.shape = " + str(aligned_min_hela_cuts.shape))
print("aligned_min_mcf7_cuts.shape = " + str(aligned_min_mcf7_cuts.shape))
print("aligned_min_cho_cuts.shape = " + str(aligned_min_cho_cuts.shape))
# +
data_version = '_neg_rate_2'#'_neg_rate_1'#''
aligned_5ss_dict = {
'min_df' : aligned_min_df,
'min_hek_count' : aligned_min_hek_cuts,
'min_hela_count' : aligned_min_hela_cuts,
'min_mcf7_count' : aligned_min_mcf7_cuts,
'min_cho_count' : aligned_min_cho_cuts,
}
pickle.dump(aligned_5ss_dict, open('alt_5ss_data_aligned' + data_version + '.pickle', 'wb'))
# -
| data/a5ss/prepare_data_5ss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Naive Bayes Hidden Markov Coin Toss
# author: <NAME> [<a href="sendto:<EMAIL>"><EMAIL></a>]
# In this example we will be using a simple Hidden Markov Model to demonstrate how it can be used in a Naive Bayes classifier. In this particular example we have a couple friends who enjoy playing a game of chance. Someone tosses a coin and whenever it lands on heads, the tosser gains \$1, otherwise he pays \$1. However some of our friends are prone to using rigged coins in order to come out ahead in the long run.
#
# Players fall into 3 categories, non-cheaters, smart cheaters, and dumb cheaters. Non-cheaters, as the name implies, do not use rigged coins. Smart cheaters make sure to switch between rigged and unrigged coins in order to keep the deception going. Dumb cheaters only use rigged coins.
from pomegranate import *
import numpy as np
# %pylab inline
# Let's start by creating our hmm's. Since this is a simple example, we will only have 2 states, rigged and unrigged, with discrete distributions.
rigged = State( DiscreteDistribution({'H': 0.8, 'T': 0.2}), name="rigged" )
unrigged = State( DiscreteDistribution({'H': 0.5, 'T': 0.5}), name="unrigged" )
# Now let's create our non-cheater and dumb cheater since their hmm's consist of a single state.
# +
non_cheater = HiddenMarkovModel( name="non-cheater" )
non_cheater.add_state( unrigged )
dumb_cheater = HiddenMarkovModel( name="dumb-cheater" )
dumb_cheater.add_state( rigged )
# -
# We can now add our starting states to our models.
non_cheater.start = unrigged
dumb_cheater.start = rigged
# We can also add the transitions to our non-cheater and dumb cheater.
non_cheater.add_transition( unrigged, unrigged, 1 )
dumb_cheater.add_transition( rigged, rigged, 1 )
# Now we bake our models to finalize the structure.
non_cheater.bake()
dumb_cheater.bake()
# Now we can create our smart cheater.
smart_cheater = HiddenMarkovModel( name="smart-cheater" )
# The structure of our smart cheater hmm should look like the following.
# Let's add in our starting transitions. Smart cheaters start out half the time with an unrigged coin, half the time without.
smart_cheater.add_transition( smart_cheater.start, unrigged, 0.5 )
smart_cheater.add_transition( smart_cheater.start, rigged, 0.5 )
# Then we can add our transition matrix.
smart_cheater.add_transition( rigged, rigged, 0.5 )
smart_cheater.add_transition( rigged, unrigged, 0.5 )
smart_cheater.add_transition( unrigged, rigged, 0.5 )
smart_cheater.add_transition( unrigged, unrigged, 0.5 )
# Finally we can bake the model to finalize its structure.
smart_cheater.bake()
# We can view the structures of all our hmm's in the images below.
# +
plt.title("smart cheater hmm")
smart_cheater.plot()
plt.title("dumb cheater hmm")
dumb_cheater.plot()
plt.title("non-cheater hmm")
non_cheater.plot()
# -
# Now we can finally create our Naive Bayes classifier. We'll let 0 stand for a non-cheater, 1 stand for a smart cheater, and 2 stand for a dumb cheater.
players = NaiveBayes([ non_cheater, smart_cheater, dumb_cheater ])
# Now let's create a sequence of coin tosses to test our classifier on.
data = np.array([list( 'HHHHHTHTHTTTTHHHTHHTTHHHHHTH' ),
list( 'HHHHHHHTHHHHTTHHHHHHHTTHHHHH' ),
list( 'THTHTHTHTHTHTTHHTHHHHTTHHHTT' )])
# We can now check the log probability of each string of data occurring under each model.
# +
probs = players.predict_proba( data )
for i in range(len(probs)):
print "For sequence {}, {:.3}% non-cheater, {:.3}% smart cheater, {:.3}% dumb cheater.".format( i+1, 100*probs[i][0], 100*probs[i][1], 100*probs[i][2])
# -
# Finally we can classify whether a non-cheater (0), smart cheater (1), or dumb cheater (2) created the string of data.
# +
output = players.predict( data )
for i in range(len(output)):
print "Sequence {} is a {}".format( i+1, "non-cheater" if output[i] == 0 else "smart cheater" if output[i] == 1 else "dumb cheater")
# -
# We can also train our hmm's through Naive Bayes. Let's used the following data, all marked as known games for smart cheaters.
# +
X = np.array([list( 'HHHHHTHTHTTTTH' ),
list( 'HHTHHTTHHHHHTH' )])
y = np.array([ 1, 1 ])
# -
# Now let's fit our NaiveBayes classifier to our data.
players.fit( X, y )
| examples/naivebayes_hmm_cheating_coin_toss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="DwmOUSa6j5vU"
# <a id = 'section0'></a>
# 
# + [markdown] colab_type="text" id="0TStV8JPj5vX"
# ## O que vamos aprender nessa aula:
# 1. <a href = '#section1'>Funções</a><br>
# 1.1 <a href = '#section1.1'>Motivação</a><br>
# 1.2 <a href = '#section1.2'>Definição e Regras</a><br>
# 1.3 <a href = '#section1.3'>Parâmetros e Argumentos</a><br>
# 1.4 <a href = '#section1.4'>Funções com retorno</a><br>
# 1.5 <a href = '#section1.5'>Funções chamando outras funções</a><br>
# 1.6 <a href = '#section1.6'>Fluxo de execução de um programa com funções</a><br>
# 1.6.1 <a href = '#section1.6'>A função *main*</a><br>
# 1.7 <a href = '#section1.7'>Escopo de funções</a><br>
# 1.8 <a href = '#section1.8'>Passagem por cópia e por referência</a><br>
# 1.9 <a href = '#section1.9'>Como depurar um programa com funções</a><br>
# 1.9.1 <a href = "#section1.9.1"> Como resolver esses problemas?</a><br>
# 1.10 <a href = "#section1.10">O que pode dar errado?</a><br>
# 2. <a href = '#section2'>Tuplas</a><br>
# 2.1 <a href = '#section2.1'>O que é uma tupla</a><br>
# 2.2 <a href = '#section2.2'>Como declarar uma tupla</a><br>
# 2.3 <a href = '#section2.3'>Uma lista ou uma tupla</a><br>
# 2.4 <a href = '#section2.4'>Operações básicas com tuplas</a><br>
# 2.5 <a href = '#section2.5'>Por que utilizar uma tupla?</a><br>
# 2.5.1 <a href = '#section2.5.1'>Troca rápida de valores e divisão de tuplas em variáveis</a><br>
# 2.5.2 <a href = '#section2.5.2'>Uso de tuplas na entrada e saída de funções</a><br>
# 2.6 <a href = '#section2.6'>O que pode dar errado?</a><br>
# + [markdown] colab_type="text" id="NT4b-Ux9j5vY"
# <a id = 'section1'></a>
# ## 1. Funções
# + [markdown] colab_type="text" id="qOeEBEJCj5vZ"
# <a id = 'section1.1'></a>
# ### 1.1. Motivação
#
# Imagine que você receba um código de alguém. Nele, o programa deverá receber um número e dizer se ele é par, se é múltiplo de 3 e se é primo. Considere as duas implementações abaixo. Qual delas é mais fácil de entender?
#
# 
#
# <center><b>Figura 1:</b> Comparação entre dois trechos de código de mesma semântica, mas sintaxes diferentes, um estando bem modularizado e o outro não</center>
#
# O código da esquerda tem algumas vantagens. Entre elas, podemos listar:
# - É bem mais fácil de entender
# - Caso o programa apresente algum problema, é mais fácil consertar
# - Divide as tarefas do programa em tarefas menores (subtarefas), mais fáceis de resolver. Essas subtarefas são chamadas de funções.
#
#
# Já utilizamos diversas funções ao longo do curso, como `print()` e `input()`, além das diversas funções da biblioteca `math`, como `sqrt()` e `factorial()`.
#
# Funções são sequências de instruções que podem ser utilizadas para tornar códigos mais legíveis e organizados, e o melhor: podem ser utilizadas sem que seja necessário entender como elas fazem o que fazem. Você, por exemplo, não precisa saber como a função `sqrt()` calcula a raiz quadrada de um número, só precisa saber utilizá-la. Além disso, se um código apresentar erros e ele estiver modularizado em funções, fica mais simples de depurar, ou seja, testar o funcionamento de cada parte do programa e até identificar o erro.
#
#
# 
#
# <center><b>Figura 2:</b> Exemplo de um programa bem modularizado, com o uso de função</center>
#
# 
#
# <center><b>Figura 3:</b> O mesmo programa da imagem 2 sem o uso da função, <br>
# as chaves demonstram como um conjunto de linhas poderia<br>
# ser substituído por apenas uma linha se implementássemos a função</center>
#
# Agora que já vimos os benefícios do uso de funções, aprenderemos a criá-las!
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="yVXHtBXaj5va"
# <a id = 'section1.2'></a>
#
# ### 1.2. Definição e Regras
#
# Para definirmos uma função, utilizamos a palavra reservada `def`. Logo após o `def`, escrevemos o nome da função e colocamos seus parâmetros entre parênteses (trataremos de parâmetros logo à frente). Por fim, encerramos a linha com dois pontos, `:`.
# Observe a criação de uma função simples que imprime "Olá!":
# + colab={} colab_type="code" id="BSokiO_tj5vb"
def diz_ola():
print("Olá!")
# + [markdown] colab_type="text" id="VKUo7U_1j5vf"
# Observe que ao executar a celula acima, o `print()` não é executado, pois a palavra chave `def` apenas define o trecho de código.
#
# Para utilizarmos a função, basta escrevermos seu nome seguido dos parenteses:
# + colab={} colab_type="code" id="VDmJHW8Bj5vf"
diz_ola()
# + [markdown] colab_type="text" id="1qaPHI6aj5vj"
# Repare que o `print` dentro da função possui uma tabulação, ou 4 espaços, assim como um `if` `else`. Qualquer coisa que seja colocada fora dessa regra não será considerada como parte da função.
#
# A forma geral de definição de uma função é:
#
# ```py
# def <nome> ( <parametros> ):
# <comandos da função>
# ```
#
# Mas o que aconteceria se usássemos uma palavra reservada, por exemplo `input`, como nome da nossa função?
# + colab={} colab_type="code" id="8ErEqs4zj5vk"
def input():
print("A função funciona!")
# + colab={} colab_type="code" id="gAQ__rFjj5vo"
input()
# + [markdown] colab_type="text" id="Yr2giqDyj5vr"
# Perceba que podemos redefinir funções que já existem, embora não seja uma boa prática de programação no momento. Este é um conceito mais avançado que não será abordado no curso. Por agora, é preferível criar novas funções do que sobrescrever as já existentes.
#
# + [markdown] colab_type="text" id="By__iBOsj5vs"
# ### Exercício 1
#
# Crie uma função que imprima um número lido do teclado. Tanto a leitura do número quanto sua impressão devem ser feitas dentro da função!
# + colab={} colab_type="code" id="zPcs_SVDj5vt"
#Defina sua função aqui
# + colab={} colab_type="code" id="ShdU4DBmj5vw"
#Chame sua função aqui
# + [markdown] colab_type="text" id="1BBoeCk0j5vz"
# Veremos exemplos mais complexos a partir de agora, falando de parâmetros.
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="II20iS4Dj5v0"
# <a id = 'section1.3'></a>
# ### 1.3. Parâmetros e Argumentos
#
# Funções como a `diz_ola` são muito simples e parecem até um tanto desnecessárias, portanto veremos funções mais complexas, que possuem ao menos um parâmetro.
#
# **Parâmetro** é a variável que recebe um valor em uma função, enquanto **argumento** é o valor que você passa para a função.
# Por exemplo: quando você quer saber a raiz quadrada de um número, você chama a função `sqrt()`. Essa função tem apenas um parâmetro e, portanto, só necessita de um número como argumento.
#
# 
#
# <center><b>Figura 4:</b> Demonstração da diferença entre argumentos e parâmetros em um código</center>
#
# Veja no exemplo a seguir como fazer suas funções receberem parâmetros. Nossa função receberá um número e imprimirá o resultado de sua multiplicação por 5:
# + colab={} colab_type="code" id="RuZY4hayj5v1"
#Definimos nossa função aqui
def multiplica_por_5(num):
num_multiplicado = num * 5
print(num_multiplicado)
# + colab={} colab_type="code" id="42LWZ5SNj5v5"
#Chamamos nossa função aqui
x = 40
multiplica_por_5(x)
# + [markdown] colab_type="text" id="dIOqoa69j5v7"
# Nossas funções podem ter multiplos parâmetros! Basta separá-los por virgulas, como é demonstrado em nosso próximo exemplo. Nele, recebemos dois números como parâmetros e imprimimos o maior dos dois:
# + colab={} colab_type="code" id="l7V3BvIJj5v8"
def imprime_maior(numA, numB):
if numA > numB:
print(numA)
else:
print(numB)
# + colab={} colab_type="code" id="6YAICfNQj5v-"
imprime_maior(10, 40)
# + [markdown] colab_type="text" id="ZARM8zAJj5wC"
# Repare que os nomes dos argumentos não precisam ser iguais aos nomes definidos nos parâmetros da função. Por exemplo: na função `multiplica_por_5`, o nome do parâmetro definido na função é "num", mas quando chamamos a função, mandamos a variável `x` como argumento, e lá, o valor da variável `x` é atribuido à `num`.
#
# ### Exercício 2
# Agora que sabemos criar funções com parâmetros, crie uma função que, dado dois catetos de um triângulo retângulo, imprima sua hipotenusa.
# + colab={} colab_type="code" id="2rTpRpBUj5wC"
cateto1 = int(input())
cateto2 = int(input())
#sua função deve ser criada a partir daqui
# + colab={} colab_type="code" id="7Lh3L4cqj5wI"
#teste sua função aqui
# + [markdown] colab_type="text" id="QrHpZzRXj5wL"
# Mas e se não quisermos que nossa função imprima o resultado, e sim salve esse resultado para uma variável? Responderemos essa pergunta com o comando `return` a seguir!
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="wfQMp2ZEj5wL"
# <a id = 'section1.4'></a>
# ### 1.4. Funções com retorno
#
# Nem sempre queremos que as funções que criamos imprimam algo. As vezes só queremos que elas calculem algo e salvem isso em uma variável. Podemos fazer isso com o comando `return`. No nosso ultimo exemplo, nossa função `imprime_maior` sempre imprimia o maior número. Agora criaremos uma função parecida, que informa o maior de dois números:
# + colab={} colab_type="code" id="WQkswnqmj5wM"
#Definimos a função aqui
def maior(numA, numB):
if numA > numB:
return numA
else:
return numB
# + colab={} colab_type="code" id="k7RdVTm2j5wP"
#Chamamos a função aqui
var = maior(40, 320)
var
# + [markdown] colab_type="text" id="px1HzxRTj5wR"
# ### Exercício 3
#
# Modifique a função que calcula a hipotenusa nas celulas abaixo. Dessa vez, em vez de imprimir a hipotenusa na função, retorne seu valor para uma variável.
# + colab={} colab_type="code" id="AD36IHajj5wS"
#modifique a função aqui
# + colab={} colab_type="code" id="TFrCqT7uj5wW"
#teste sua função aqui
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="aejKs7DBj5wa"
# <a id = 'section1.5'></a>
# ### 1.5. Funções chamando outras funções
#
# Podemos também utilizar o retorno de uma função como argumento de outra. Essa prática é muito útil em determinadas situações, como no exemplo a seguir:
# Imagine que você precisa descobrir o maior entre 4 números! Apresentaremos duas soluções para o problema, sendo a segunda solução mais recomendada. Lembre-se de que a função `maior(a, b)` já foi definida acima, então não é preciso defini-la novamente.
# + colab={} colab_type="code" id="G7VNJFRUj5wa"
a = 20
b = 30
c = 2
d = 70
#Aqui criamos duas variáveis para salvar o maior valor entre A e B e entre C e D
#e chamamos a função maior para nos retornar qual é o maior
maior_entre_A_e_B = maior(a, b)
maior_entre_C_e_D = maior(c, d)
#Após descobrirmos os maiores entre A e B e entre C e D, descobrimos o maior entre
#esses dois
maior_total = maior(maior_entre_A_e_B, maior_entre_C_e_D)
print(maior_total)
# + colab={} colab_type="code" id="z_LQlHGLj5wd"
#Aqui o retorno das chamadas de função "maior(a, b)" e "maior(c, d)" servem
#como argumento para a chamada de função abaixo
maior_total = maior(maior(a,b), maior(c, d))
print(maior_total)
# + [markdown] colab_type="text" id="v1RnFjtUj5wf"
# Além disso, funções podem chamar outras funções dentro delas. Imagine que não podemos utilizar a função `tan()`, mas precisamos calcular a tangente de um angulo. Podemos criar uma nova função `tan` que funcionará da mesma forma:
# + colab={} colab_type="code" id="Sx2JsxiGj5wg"
import math
# + colab={} colab_type="code" id="o48H2ZN0j5wi"
#Aqui o resultado do seno de x e do cosseno de x será calculado
#e este valor calculado é retornado
def tg(x):
return (math.sin(x)/math.cos(x))
# + colab={} colab_type="code" id="YAsFcN12j5wl"
resposta = tg(math.radians(45))
print(resposta)
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="cik__kAcj5wo"
# <a id = 'section1.6'></a>
#
# ### 1.6. Fluxo de execução de um programa com funções
#
# Vamos acompanhar o exemplo a seguir e entender seu fluxo de execução.
# + colab={} colab_type="code" id="RLRqKr_vj5wp"
#Definimos aqui a função que retorna o maior número
def maior(a, b):
if a > b:
return a
else:
return b
#Definimos aqui a função que diz se o número é primo ou não
def eh_primo(x):
y = 2
while(y != x):
if x % y == 0:
return False
y += 1
return True
#O código começa aqui!
a = int(input())
b = int(input())
maior_num = maior(a, b)
print(maior_num, "é primo?")
resposta = eh_primo(maior_num)
if resposta:
print("Sim!")
else:
print("Não!")
# -
# <a id = 'section1.6.1'></a>
#
# #### 1.6.1. A função _main_
#
# Na maioria das linguagens de programação existe uma função chamada `main` (em tradução literal, função principal), e é nessa função que todas as outras funções são chamadas. Em Python a declaração da `main` não é obrigatória, entretanto é possível utilizar tal recurso, e seu uso é bem visto (uma vez que torna o código mais organizado).
# +
# Código sem uso da função main:
def bhaskhara(a, b, c):
delta = b ** 2 - 4 * a * c
if delta < 0:
return None, None
x1 = b + delta ** (1/2) / (2 * a)
x2 = b - delta ** (1/2) / (2 * a)
return x1, x2
a = int(input("Insira 'A':"))
b = int(input("Insira 'B':"))
c = int(input("Insira 'C':"))
x1, x2 = bhaskhara(a, b, c)
print(f"x1 = {x1}; x2 = {x2}")
# +
# Código com uso da função main (método 1):
def bhaskhara(a, b, c):
delta = b ** 2 - 4 * a * c
if delta < 0:
return None, None
x1 = b + delta ** (1/2) / (2 * a)
x2 = b - delta ** (1/2) / (2 * a)
return x1, x2
def main():
a = int(input("Insira 'A':"))
b = int(input("Insira 'B':"))
c = int(input("Insira 'C':"))
x1, x2 = bhaskhara(a, b, c)
print(f"x1 = {x1}; x2 = {x2}")
main()
# +
# Código com uso da função main (método 2 / mais elegante):
def bhaskhara(a, b, c):
delta = b ** 2 - 4 * a * c
if delta < 0:
return None, None
x1 = b + delta ** (1/2) / (2 * a)
x2 = b - delta ** (1/2) / (2 * a)
return x1, x2
def main():
a = int(input("Insira 'A':"))
b = int(input("Insira 'B':"))
c = int(input("Insira 'C':"))
x1, x2 = bhaskhara(a, b, c)
print(f"x1 = {x1}; x2 = {x2}")
if __name__ == "__main__":
main()
# -
# Algumas dicas interessantes quanto ao uso da `main`:
# * É interessante deixar a `main` como a última ou primeira função de seu arquivo, isso facilita a leitura.
# * Em programas que utilizam separação em arquivos é usual a `main` ter um arquivo próprio, normalmente chamado `main.py`.
# * Em outras linguagens de programação, como Java e C, o uso da `main` é obrigatório.
# * Mesmo que não exista uso da `main` no seu código é **sempre** interessante que funções **não** sejam declaradas em meio às chamadas de outras funções e declarações de variáveis (ver exemplo abaixo)
# +
# Esse código é ruim:
a = int(input("Insira 'A':"))
b = int(input("Insira 'B':"))
c = int(input("Insira 'C':"))
# A função está sendo declarada dentro do fluxo de execução da parte
# principal do código, isso pode tornar a leitura confusa,
# principalmente em códigos grandes (onde, nesse caso, existiriam
# diversas funções espalhadas pelo arquivo).
def bhaskhara(a, b, c):
delta = b ** 2 - 4 * a * c
if delta < 0:
return None, None
x1 = b + delta ** (1/2) / (2 * a)
x2 = b - delta ** (1/2) / (2 * a)
return x1, x2
x1, x2 = bhaskhara(a, b, c)
print(f"x1 = {x1}; x2 = {x2}")
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="1kfEV-40j5wr"
# <a id = 'section1.7'></a>
#
# ### 1.7. Escopo de variáveis
#
# O que aconteceria caso você criasse uma variável dentro de uma função e tentasse imprimi-la fora dela? Execute o código a seguir e veja o que acontece.
# + colab={} colab_type="code" id="r4cs8bZPj5ws"
#Definimos nossa função aqui
def funcao1():
dentro = 40
print(dentro)
# + colab={} colab_type="code" id="BSBxQBrdj5ww"
#Tentamos imprimir a variável "dentro" aqui
print(dentro)
# + [markdown] colab_type="text" id="qzDYzL6Oj5wz"
# Como visto acima, um erro ocorre, informando que a variável `dentro` não está definida. Claro que ela está definida e funciona muito bem dentro do <b>escopo</b> da função, mas ao tentar utiliza-la fora desse escopo, recebemos um erro. É muito importante prestarmos atenção no escopo dessas funções para que esse tipo de erro não ocorra.
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="Q6IcmEF-j5xG"
# <a id = 'section1.8'></a>
#
# ### 1.8. Passagem por cópia e por referência
#
# Dependendo do tipo de dado que for passado como argumento da função, precisamos tomar cuidado ao altera-lo. Alguns tipos de dados, como `int`, `float`, `bool`, `str` não são alterados fora da função, e portanto, precisam ser retornados.
#
# Chamamos isso de passagem por cópia!
#
# Observe o exemplo a seguir:
# + colab={} colab_type="code" id="z9p1Joz3j5xH"
#Definimos aqui uma função que multiplica por 100 um número inteiro
def multiplica_por_100(x):
x *= 100
print("O valor de x dentro da função é:", x, end = "\n\n")
x = 25
print("O valor de x antes de chamar a função é:", x, end = "\n\n")
multiplica_por_100(x)
print("O valor de x depois de chamar a função é:", x, end = "\n\n")
# + [markdown] colab_type="text" id="6OETx5qNj5xL"
# Porém, alguns tipos de dados, como dicionários e listas (e objetos, que veremos no módulo 2), caso sejam alterados dentro da função, também são alterados fora da função.
#
# Chamamos isso de passagem por referência!
#
# Observe o exemplo abaixo:
# + colab={} colab_type="code" id="yWQluvIpj5xM"
# Definimos aqui uma função que multiplica por 100 todos os números inteiros da lista
def multiplica_lista_por_100(lis):
for i in range(len(lis)):
lis[i] *= 100
print("Os valores da lista dentro da função são:", lis, end = "\n\n")
lis = [1, 2, 3, 4, 5]
print("Os valores da lista antes da função são:", lis, end = "\n\n")
multiplica_lista_por_100(lis)
print("Os valores da lista depois da função são:", lis, end = "\n\n")
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
#
# + [markdown] colab_type="text" id="owuxDD2aj5xN"
# <a id = 'section1.9'></a>
#
# ### 1.9. Como depurar um programa com funções
#
# Quando temos problemas com códigos que possuem diversas funções, é bem comum verificar se cada uma está funcionando corretamente, isso se chama depurar. Ao testar e verificar que cada script menor está apresentando o resultado esperado, voltamos para a "parte principal" do código, pois sabemos que o problema provavelmente está lá, e não nas funções.
#
# <p>Há três tipos de erros que podem acontecer:</p>
# <ol>
# <li><b>Erros de sintaxe:</b> se referem à estrutura de um programa e as regras que regem essa estrutura. Exemplo: parêntesis que abrem mas não fecham, falta de indentação em local necessário, dois números lado a lado sem operador.</li>
# <li><b>Erros de semântica/execução:</b> ocorre quando o significado do seu programa está incorreto. O código pode estar dentro de todas as regras daquela linguagem mas, ainda assim, não fazer sentido. Exemplo: dividir um número por uma string, dividir zero por algum número ou vice versa, utilização de palavras reservadas pela linguagem em locais indevidos. Esses erros levam a falhas na execução do programa.</li>
# <li><b>Erros de lógica:</b> são mais difíceis de identificar, já que o programa está rodando e aparentemente correto. Acontece quando o código não funciona da maneira esperada, ou seja, a solução dada pelo programador não é a solução certa. Exemplo: uso incorreto de operadores, não colocar um ponto de parada em um loop (isso faria ele rodar infinitamente), operações incorretas.</li>
# </ol>
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
#
# <a id = 'section1.9.1'></a>
#
# #### 1.9.1. Como resolver esses problemas?
#
# Há diversos métodos para depurar um programa. O jeito mais fácil é através de IDEs (Ambientes de Desenvolvimento Integrado) que oferecem essa opção, como NetBeans, VisualStudio, PyCharm e outros. No entanto, nem sempre é possível utilizar essas plataformas. Por isso, é necessário ter conhecimento para fazer isso você mesmo.
#
# Primeiramente, ao se deparar com o erro, você deve observar qual tipo está sendo apontado (sintaxe, execução ou lógica - saídas diferentes do esperado).
# No caso de erro de sintaxe, você deve revisar o seu código, procurando linha por linha o local onde a escrita não esteja de acordo com as regras da linguagem utilizada.
# Caso haja erro de execução, é necessário que você revise as operações realizadas, para ter certeza de que nenhuma acontece de maneira indevida.
# Já se o erro for de lógica, você precisa ter uma lista de saídas esperadas, ou seja, resultados que deveriam aparecer no programa e comparar com a saída fornecida. Assim, é possível procurar em que ponto do código existe uma ação incorreta.
# Em todos esses casos, a primeira coisa que o programador tem que saber são alguns possíveis resultados para aquele programa.
# A melhor maneira de saber o que está acontecendo com o seu código é imprimir na tela os resultados passo a passo. Inserir condicionais junto com as funções `print()` também pode ajudar, já que o programa apenas imprimiria caso aquela condição fosse atendida, caso contrário, você sabe que existe um erro (o contrário também é válido).
# +
# Exemplo de código sem testes de saída de funções:
def total(x, y, z):
t = x * y * z
return t
def imposto(x):
imp = (x * 11) / 100
return imp
def inss(x):
i = (x * 8) / 100
return i
def sindicato(x):
sind = (x * 5) / 100
return sind
# Obtendo o salário total:
sal_hora = 15.50 # salario por hora
hora_dia = 8 # horas trabalhadas
dias_mes = 20 # quantidade de dias trabalhados
sal_bruto = total(sal_hora, hora_dia, dias_mes) # salario total sem descontos
# Calculando descontos:
imp = imposto(sal_bruto)
i = inss(sal_bruto)
sind = sindicato(sal_bruto)
# OBS: Salário liquido = salario bruto - descontos
sal_liq = sal_bruto - imp - i - sind
print("O salário líquido é: R${}".format(sal_liq))
# RESULTADO ESPERADO:
# - SALBRUTO = 2480
# - IMP = 272,8
# - I = 198,4
# - SIND = 124
# - SALLIQ = 1884,8
# +
# O mesmo código com testes de saída de funções:
def total(x, y, z):
t = x * y * z
return t
def imposto(x):
imp = (x * 11) / 100
return imp
def inss(x):
i = (x * 8) / 100
return i
def sindicato(x):
sind = (x * 5) / 100
return sind
sal_hora = 15.50 # salario por hora
hora_dia = 8 # horas trabalhadas
dias_mes = 20 #q uantidade de dias trabalhados
sal_bruto = total(sal_hora, hora_dia, dias_mes) # salario total sem descontos
imp = imposto(sal_bruto)
i = inss(sal_bruto)
sind = sindicato(sal_bruto)
sal_liq = sal_bruto - imp - i - sind
# Bateria de testes:
saidas_obtidas = [sal_bruto, imp, i, sind]
saidas_esperadas = [2480, 272.8, 198.4, 124]
for i in range(len(saidas_obtidas)):
if saidas_obtidas[i] == saidas_esperadas[i]:
is_ok = "Ok!"
else: is_ok = "Erro!"
print("R${} : {}".format(saidas_obtidas[i], is_ok))
# Exibindo o salário líquido
print("\nO salário líquido é: R${}".format(salLiq))
# RESULTADO ESPERADO:
# - SALBRUTO = 2480
# - IMP = 272,8
# - I = 198,4
# - SIND = 124
# - SALLIQ = 1884,8
# -
# ### Exercício 4
#
# Analise os códigos abaixo e corrija o que for necessário.
# +
import math
def soma(x,y):
x + y
def pitagoras(b,c):
x = b**2
y = c**2
a = soma(x,y)
return a
num1 = input("Digite um número: ")
num2 = input("Digite outro número: ")
num3 = pitagoras(b,c)
if(num3 > 10):
print("num3 é maior que 10")
else:
print("num3 é menor que 10")
if(type(num3) == 'int'):
print("num3 é inteiro \nnum3 = ", num3, "é ", type(num3))
else if(type(num3 == 'int')):
print("num3 não é inteiro \nnum3 = ", num3, "é ", type(num3))
# +
def soma(num1, num2):
soma = num1 + num2
def eh_primo:
soma = soma(num1, num2)
for i in range(1, soma):
if(soma % i == 0):
eh_divisor++
else:
continue
if(eh_divisor != 2):
print("{} não é primo".format(soma))
else:
print("{} é primo".format(soma))
num1 = 10
num2 = 3
#OBS: SAÍDA ESPERADA: "13 é primo"
# -
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# Agora veja o mesmo problema sendo resolvido com o auxílio de um IDE, nesse caso foi utilizado o PyCharm.
# +
# Execute esta célula para visualizar o vídeo:
from IPython.display import IFrame
display(IFrame("https://www.youtube.com/embed/_hCd39HV0sM", 480, 360))
# -
# <br><img src="https://cloud.netlifyusercontent.com/assets/344dbf88-fdf9-42bb-adb4-46f01eedd629/f42cfcb0-d796-4aaf-a41b-a61d373da369/hr1-raypham.gif" /><br>
# <a id = 'section1.10'></a>
#
# ### 1.10. O que pode dar errado?
#
# Abaixo listaremos apenas alguns erros possíveis e para cada um, apenas algumas soluções possíveis, lembre-se de buscar ajuda com seus colegas, professores e comunidades *on-line* caso se depare com um erro não abordado aqui. Cada vez que se deparar com um erro será mais fácil resolvê-lo conforme ganha experiência, por isso é muito importante que sempre pratique.
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# #### Erro 1: Chamada incorreta da função:
# É comum que esqueçamos um ou mais parâmetros de uma função, ou mesmo coloquemos na ordem errada (nesse segundo caso a situação pode ser ainda pior, pois não é emitido nenhum aviso), devemos sempre prestar muita atenção para não cometer nenhum desses erros.
# +
def diga_olá(nome, idade):
return "Olá, sou {} e tenho {} anos".format(nome, idade)
diga_olá(16, "Juvenal")
# +
# Será emitido o erro "TypeError: diga_olá() missing 1 required positional argument: 'idade'"
diga_olá("Juvenal")
# -
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# #### Erro 2: Declaração incorreta da função
#
# É possível esquecer de inserir a indentação da função, ou até mesmo dos dois pontos, conforme o exemplo abaixo:
def soma(x, y)
return x + y
# O caso acima é bem absurdo, mas podemos cometer esse mesmo erro de formas mais críticas (e não seremos notificados pelo terminal, vejamos:
def soma(lista_de_numeros):
soma = 0
for num in lista_de_numeros:
soma = soma + num
return soma
# <hr style="border-top: 5px dashed #234B6B; background-color: #E9B74F"/>
#
# + [markdown] colab_type="text" id="Srvco-KQj5xO"
# <a id = 'section2'></a>
#
# ## 2. Tuplas
#
# <a id = 'section2.1'></a>
#
# ### 2.1. O que é uma tupla?
#
# Em aulas anteriores estudamos listas e seu funcionamento. As tuplas são listas imutáveis. E o que isso quer dizer? om listas podemos adicionar ou remover elementos; com tuplas sso é impraticável.
# Podemos resumir uma tupla como sendo uma lista que tem as seguintes restrições:
#
# - Não é possível adicionar novos elementos;
# - Não é possível remover elementos existentes;
# - Não é possível alterar os elementos;
# - Não é possível ordenar os elementos.
#
#
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="7mMmiMjYj5xP"
#
# <a id = 'section2.2'></a>
#
# ### 2.2. Como declarar uma tupla?
# Declaramos tuplas da mesma forma que declaramos listas; a diferença é que listas são delimitadas por colchetes `[]` enquanto tuplas são delimitadas por parêntesis `()`.
#
# Uma tupla pode ser declarada tanto como `tupla = (x, y)` quanto como `tupla = x, y`
#
# Uma tupla de somente um elemento é algo do tipo `(x)`, que pode ser declarada como, `t1 = (x)` ou `t1 = 1, `. <br>
# Uma tupla vazia também é declarada como `t1 = ()`.
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="NdPI31iPj5xP"
# <a id = 'section2.3'></a>
#
# ### 2.3. Uma lista ou uma tupla?
#
#
# Se você declarar uma tupla e chamar `type( < tupla > )`, a função nos retorna o tipo da estrutura. Rode o código a seguir.
#
# + colab={} colab_type="code" id="ZRfV9Reaj5xQ"
tup = ("Carlos", 26, 2, 1999)
type(tup)
# + [markdown] colab_type="text" id="ljRbhWS2j5xU"
# Você também pode declarar suas tuplas chamando uma função `tuple ( " uma string qualquer " )`. Essa forma de declaração nos retorna uma tupla de caracteres.
# -
tupla_strings = tuple("Olá, sou uma string!")
tupla_strings
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# + [markdown] colab_type="text" id="-gzdBDWmj5xV"
# <a id = 'section2.4'></a>
#
# ### 2.4. Operações básicas com tuplas
#
# As operações que fazemos com as listas também podem ser utilizadas com tuplas. Considere os exemplos a seguir:
# + colab={} colab_type="code" id="Eb5mw98lj5xX"
# Aqui estamos criando uma tupla com os elementos 3 "vamos" 4 e "fugir"
t = (3, "vamos", 4, "fugir")
# Aqui estamos perguntando se 3 está dentro da nossa tupla, se sim retornara True se não retornara False.
3 in t
# + colab={} colab_type="code" id="BgGgk6EWj5xa"
# Declaração de tupla
t1 = 2, "chega", 555, "C++"
# Concatenação das tuplas
t2 = t + t1
# Printando tupla resultante
print(t2)
# + colab={} colab_type="code" id="zYU0x3AMj5xd"
# Tupla de somente 1 elemento
t3 = 1,
# Concatenando essa tupla a ela mesma 5 vezes.
t4 = t3 * 5
print(t4)
# + [markdown] colab_type="text" id="9bYTA4Bmj5xj"
# Podemos acessar elementos da tupla como acessando elementos de uma lista.
# + colab={} colab_type="code" id="kcfMYSOij5xk"
print(t)
print(t[0])
# Somando o primeiro elemento com o terceiro elemento da tupla.
t5 = t[0] + t[2]
print(t5)
print(t[-1])
# O -1 é o indice do ultimo elemento da tupla, igual para lista.
# + [markdown] colab_type="text" id="XFchZd4Cj5xn"
# Podemos fazer uma tupla de listas e também uma lista de tuplas, existe uma função chamada enumerate nativa da python capaz de nos retornar tuplas com os indices e os elementos dos indices da lista.
#
# + colab={} colab_type="code" id="2ZiRRat1j5xo"
l1 = ['a', 'b', 'c', 'd', 'e']
# Criando lista de tuplas
l2 = list(enumerate(l1))
print(l2)
# + [markdown] colab_type="text" id="IZPyrUtOj5xr"
# Se você observar verá que as tuplas seguem o padrão (indice, elemento) da lista para qual chamamos o enumerate.
#
# + colab={} colab_type="code" id="gEnVI50Ej5xs"
# Criamos uma tupla de listas
tp = ([1,2,3], [4,5,6])
# Printando o terceiro elemento da primeira posição da tupla
print(tp[0][2])
# + [markdown] colab_type="text" id="JBYKR4asj5xv"
#
# Outra operação que podemos fazer com elas são comparar se uma tupla é menor, maior ou igual.
# + colab={} colab_type="code" id="-nN9NXqgj5xw"
t < t1
# + [markdown] colab_type="text" id="XAml4VP0j5xy"
# Para saber se uma tupla é menor que outra comparamos elemento por elemento, a ideia é, comparar o elemento primario se for menor ja retorna True, caso seja igual ele vai para os elementos seguintes. O mesmo vale para as operações > ou == .
# + colab={} colab_type="code" id="tXdLv3daj5xy"
(1,2,3,5) < (1,2,3,4)
# + colab={} colab_type="code" id="ThvbP4qrj5x0"
(1,2,3,4) < (2,3,4,5)
# + colab={} colab_type="code" id="AypOKqYTj5x3"
# Printando tupla recortada do segundo ao terceiro elemento
print(t[1:3])
# + [markdown] colab_type="text" id="79lNXN8Pj5x6"
# Também conseguimos usar o for com tuplas, se liga.
# + colab={} colab_type="code" id="Ozfl-x3zj5x7"
for x in ("a", 1, 2, 3, "b", 4, 5, 6, "c"):
print(x)
# -
# <hr style="border-top: 3px dashed #234B6B; background-color: #E9B74F"/>
# <a id = 'section2.5'></a>
#
# ### 2.5. Por que utilizar uma tupla?
#
# Se uma tupla funciona como uma lista, mas não pode ser alterada ao longo do programa, por que escolher esse tipo de variável? Apesar do seu funcionamento parecido, dizer que tuplas são somente listas imutáveis não está totalmente certo! Usamos isso para deixar o aprendizado dessa estrutura um pouco mais didático.
#
# Você deve usar uma tupla sempre que os dados inseridos forem constantes. Além disso, as tuplas tem como objetivo trabalhar com dados heterogêneos, ou seja, dados de diversos tipos e significados. Por exemplo: se o seu programa tem como objetivo trabalhar com coordenadas geográficas, o uso da tupla garante que esses valores não serão modificados ao longo das operações realizadas. Sem contar que, com elas, podemos agilizar, e muito, a escrita do código.
#
# De forma resumida, as tuplas são mais rápidas e previnem o seu script de alterações indesejadas.
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# <a id = 'section2.5.1'></a>
#
# #### 2.5.1. Troca rápida de valores e divisão de tuplas em variáveis
# Sem o uso de tuplas, utilizamos três linhas de código para trocar os valores de duas variáveis. Porém, com tuplas, é possível fazer isso em apenas uma linha. Vejamos:
# +
a = 1
b = 2
# Sem tuplas:
aux = a
a = b
b = aux
a, b
# +
a = 1
b = 2
# Com tuplas:
a, b = b, a
a, b
# -
# Isso ocorre por uma propriedade das tuplas que permite que elas sejam “separadas” em variáveis. Exemplo:
# +
# Declarando uma tupla:
coordenadas = 1.2, 5.9
# "Separando" a tupla em duas variáveis:
x, y = coordenadas
x, y
# -
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# <a id = 'section2.5.2'></a>
#
# #### 2.5.2. Uso de tuplas na entrada e saída de funções:
# Como visto acima, as tuplas podem ser divididas em variáveis. Tal propriedade ainda nos permite retornar “vários valores” a partir de uma mesma função.
# +
import math
# Função que permite encontrar as raízes de uma equação de segundo grau:
def bhaskara(a, b, c):
delta = b**2 - 4 * a * c
if delta < 0: return None, None
return (b + math.sqrt(delta))/(2*a), (b - math.sqrt(delta))/(2*a)
# x1 e x2 receberão as raízes da equação de segundo grau.
x1, x2 = bhaskara(2, 3, 1)
x1, x2
# -
# Podemos ainda passar um número indeterminado de valores como argumentos de uma função apenas colocando `*args` como parâmetro. `*args` é, na verdade, uma tupla e fará a função receber quantos parâmetros você precisar.
# + colab={} colab_type="code" id="q4B42Waqj5w0"
#Definimos nossa função aqui
def soma_numeros(*args):
soma = 0
for x in args:
soma += x
return soma
# + colab={} colab_type="code" id="B3Fjr6SRj5w4"
#Chamamos nossa função aqui:
#Tente colocar mais ou menos valores
soma = soma_numeros(10, 9, 8, 7, 6, 5, 4, 3, 2, 1)
soma
# + [markdown] colab_type="text" id="IrZEwKO0j5w-"
# ### Exercício 5
# Crie uma função que retorne o maior número entre os parâmetros. Utilize o parâmetro `*args` para poder colocar quantos números forem necessários!
# + colab={} colab_type="code" id="IamiLL18j5w_"
# Defina sua função aqui
# + colab={} colab_type="code" id="0OP6W8Rcj5xD"
# Teste sua função aqui
# -
# Agradecemos pela atenção e esperamos que tenham aprendido bem o conteúdo, para que possamos sempre melhorar é de extrema importância que colaborem através de seu feedback, ficaremos ainda mais gratos caso respondam nosso formulário sobre a aula.
# <br><img src="https://cloud.netlifyusercontent.com/assets/344dbf88-fdf9-42bb-adb4-46f01eedd629/f42cfcb0-d796-4aaf-a41b-a61d373da369/hr1-raypham.gif" /><br>
# <a id = 'section2.6'></a>
#
# ### 2.6. O que pode dar errado?
#
# Abaixo listaremos apenas alguns erros possíveis e para cada um, apenas algumas soluções possíveis, lembre-se de buscar ajuda com seus colegas, professores e comunidades on-line caso se depare com um erro não abordado aqui. Cada vez que se deparar com um erro será mais fácil resolvê-lo conforme ganha experiência, por isso é muito importante que sempre pratique.
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# #### Erro 1: Tentar acessar índice inexistente
# É um erro recorrente, principalmente quando trabalhamos com *loops*, nesses casos devemos ter muito cuidado e definir bem o intervalo que queremos.
#
# A mensagem de erro exibida será `IndexError: tuple index out of range`
minha_tupla = (1, 2, 3, 4, 5)
minha_tupla[5]
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# #### Erro 2: "Dividir" a tupla em um número muito grande ou muito pequeno de variáveis
# Pode ocorrer quando recebemos uma tupla como retorno de uma função. Quando não quiser uma das variáveis basta inserir um `_` em seu lugar.
#
# **Exemplo:**
coordenadas = (10, 40)
x, _ = coordenadas
# O erro exibido será `ValueError: not enough values to unpack` caso exista tentativa de receber mais valores do que o retornado por uma função:
# +
def retorna_coordenadas():
return 10, 40
x, y, z = retorna_coordenadas()
# -
# O erro exibido será `ValueError: too many values to unpack` caso exista tentativa de receber menos valores do que o retornado por uma função:
# +
def retorna_coordenadas():
return 10, 40, 30
x, y = retorna_coordenadas()
# -
# <hr style="border-top: 1px dashed #234B6B; background-color: #E9B74F"/>
# #### Erro 3: Tentar modificar uma tupla
# Lembre-se que tuplas são imutáveis, se quiser modificá-las você pode convertê-las para uma lista.
#
# Caso tente modificar uma tupla tentando atribuir algum valor aos seus índices será exibido o erro `TypeError: 'tuple' object does not support item assignment`
# Incorreto
coordenadas = (10, 20)
coordenadas[1] = 40
# Correto
coordenadas = [10, 20]
coordenadas[1] = 40
# Convertendo tupla para lista:
minha_tupla = (1, 2, 3, 4, 5)
minha_lista = list(minha_tupla)
# <hr style="border-top: 5px dashed #234B6B; background-color: #E9B74F"/>
# <a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>
| aula_007/.ipynb_checkpoints/Aula007-checkpoint.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + cell_id="fff26dab-031b-4adf-9f97-8b194cbfa975" deepnote_cell_height=81 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1649515042849 source_hash="9b82ee11" tags=[]
import pandas as pd
/ + cell_id="eb2e3916a4a144529d89145341e70fb3" deepnote_cell_height=135 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1404 execution_start=1649515043752 source_hash="d44b450b" tags=[]
original_df = pd.read_csv('../data/interim/cleaned-merge.csv',low_memory=False)
subset_original = original_df.loc[:,['search_query','profile_link','join_date_from_earliest',
'badge_preferred_freelancer','badge_verified']]
/ + cell_id="a0efebdcf2574f089c57c98219d1c33e" deepnote_cell_height=581.390625 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=7948 execution_start=1649515132433 source_hash="a4ce152e" tags=[]
path = '../data/data_graveyard/cleaned-gender-annotated-v5.csv'
def replace_columns(df_path,original_df):
#Load dataframe
df = pd.read_csv(path,low_memory=False)
#Merge the columns from the newer dataframe
new_df = df.merge(subset_original, on=['search_query','profile_link'])
#Drop old columns
new_df = new_df.drop(columns=['join_date_from_earliest_x','badge_preferred_freelancer_x','badge_verified_x'])
#Rename the new columns
new_df = new_df.rename(columns = {'join_date_from_earliest_y':'join_date_from_earliest',
'badge_preferred_freelancer_y': 'badge_preferred_freelancer',
'badge_verified_y':'badge_verified'})
print(new_df.shape)
new_path = df_path.replace('data_graveyard','gender-annotated')
print(new_path)
new_df.to_csv(new_path,index=False)
replace_columns(path,subset_original)
/ + cell_id="c12c52d4805a425ca2c25e646a04f15a" deepnote_cell_height=156.578125 deepnote_cell_type="code" deepnote_output_heights=[60] deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1649513285689 owner_user_id="90b26b03-99c4-40c0-a870-8175b99a6ae3" source_hash="15d271b2" tags=[]
new_df.badge_verified.value_counts()
/ + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
/ <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=acc27b92-84be-4130-8026-204943f38189' target="_blank">
/ <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,<KEY> > </img>
/ Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| notebooks/code-graveyard/replace_columns_in_df.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Introduction to Python
# Ported to python from http://htmlpreview.github.io/?https://github.com/andrewpbray/oiLabs-base-R/blob/master/intro_to_r/intro_to_r.html
# First, we need to import the libraries that we need. By convention, we apply aliases that we can use to reference the libraries later. `pandas` contains classes for working working with `Series` and `DataFrame`. These are very similar to `list` and `data.frame` in R. `numpy` contains a number of mathematical and statistical functions.
import pandas as pd
import numpy as np
# Now we will load the arbuthnot.csv file into a pandas `DataFrame`.
arbuthnot = pd.read_csv("arbuthnot.csv")
# ## The Data: Dr. Arbuthnot’s Baptism Records
# The Arbuthnot data set refers to Dr. <NAME>, an 18th century physician, writer, and mathematician. He was interested in the ratio of newborn boys to newborn girls, so he gathered the baptism records for children born in London for every year from 1629 to 1710. We can take a look at the data by typing its name into the console. Adding `.head()` will return just the first few rows of the data.
arbuthnot.head()
# What you should see are four columns of numbers, each row representing a different year: the first entry in each row is simply the row number (an index we can use to access the data from individual years if we want), the second is the year, and the third and fourth are the numbers of boys and girls baptized that year, respectively. Use the scrollbar on the right side of the console window to examine the complete data set.
#
# Note that the row numbers in the first column are not part of Arbuthnot’s data. Pandas adds them as an index that can be used to identify the columns. You can think of them as the index that you see on the left side of a spreadsheet. In fact, the comparison to a spreadsheet will generally be helpful. Python has stored Arbuthnot’s data in a kind of spreadsheet or table called a data frame.
#
# You can see the dimensions of this data frame by typing:
arbuthnot.shape
# This command should output (82 3), indicating that there are 82 rows and 3 columns. You can see the names of these columns (or variables) by typing:
arbuthnot.columns
# You should see that the data frame contains the columns year, boys, and girls.
# ## Some Exploration
# Let’s start to examine the data a little more closely. We can access the data in a single column of a data frame (a pandas `Series`) separately using a command like
arbuthnot['boys']
# Note, you could have also used `arbuthnot.boys`, but the square bracket syntax is preferrable as the . syntax won't work in all cases.
# **Exercise 1** What command would you use to extract just the counts of girls baptized? Try it!
# Notice that the way pandas has printed these data is different. When we looked at the complete data frame, we saw 82 rows, one on each line of the display with a column header at the top. These data are no longer structured in a table with other variables, so they have no column header. pandas has added numbers along the left side of the printout to indicate locations within the Series. For example, `5218` follows `0`, indicating that `5218` is the first entry in the vector. And if `43` starts a line, then that would mean the first number on that line would represent the 43rd entry in the vector.
#
# The library that is the basis of most of the plotting that we will do is matplotlib. This can be imported using
import matplotlib.pyplot as plt
# pandas uses matplotlib behind the scenes to do some of it's own built in plotting. Another plotting library that we will see eventually is seaborn, which also utilizes matplotlib.
#
# We can create a simple plot of the number of girls baptized per year with the command below. This will draw the plot on a matplotlib axes. We need to use matplotlib to actually show the plot.
arbuthnot.plot.scatter(x='year', y='girls')
plt.show()
# The code above creates a scatterplot with each x,y pair indicated by a circle. If we wanted to connect the data points with lines, we could use the line plot type. Note how a legend is added for us automatically.
arbuthnot.plot.line(x='year', y='girls')
plt.show()
# Most libraries in python document their functions extensively. To read what a function does and learn the arguments that are available to you, just type in a question mark followed by the name of the function that you’re interested in. Try the following.
# ?arbuthnot.plot
# **Exercise 2** Is there an apparent trend in the number of girls baptized over the years?
# How would you describe it?
# Now, suppose we want to plot the total number of baptisms. To compute this, we could use the fact that python supports mathematical functions. We can type in mathematical expressions like
5218 + 4683
# to see the total number of baptisms in 1629. We could repeat this once for each year, but there is a faster way. If we add the Series for baptisms for boys and girls, python will compute all sums simultaneously.
arbuthnot['boys'] + arbuthnot['girls']
# What you will see are 82 numbers, each one representing the sum we’re after. Take a look at a few of them and verify that they are right. Therefore, we can make a plot of the total number of baptisms per year. However, because we want to plot a Series that is not part of the DataFrame, we cannot use the pandas built in plot. However, we can use matplotlib directly with the command
plt.plot(arbuthnot['year'], arbuthnot['boys'] + arbuthnot['girls'])
plt.show()
# Similarly to how we computed the proportion of boys, we can compute the ratio of the number of boys to the number of girls baptized in 1629 with
5218 / 4683
# or we can act on the complete vectors with the expression
arbuthnot['boys'] / arbuthnot['girls']
# The proportion of newborns that are boys
5218 / (5218 + 4683)
# or this may also be computed for all years simultaneously:
arbuthnot['boys'] / (arbuthnot['boys'] + arbuthnot['girls'])
# Note that with python as with your calculator, you need to be conscious of the order of operations. Here, we want to divide the number of boys by the total number of newborns, so we have to use parentheses. Without them, python will first do the division, then the addition, giving you something that is not a proportion.
# **Exercise 3** Now, make a plot of the proportion of boys over time. What do you see?
# Finally, in addition to simple mathematical operators like subtraction and division, you can ask R to make comparisons like greater than, >, less than, <, and equality, ==. For example, we can ask if boys outnumber girls in each year with the expression
arbuthnot['boys'] > arbuthnot['girls']
# This command returns 82 values of either `True` if that year had more boys than girls, or `False` if that year did not (the answer may surprise you). This output shows a different kind of data than we have considered so far. In the arbuthnot data frame our values are numerical (the year, the number of boys and girls). Here, we’ve asked python to create logical data, data where the values are either `True` or `False`. In general, data analysis will involve many different kinds of data types, and one reason for using python is that it is able to represent and compute with many of them.
#
# This seems like a fair bit for your first lab, so let’s stop here.
# # On Your Own
# In the previous few pages, you recreated some of the displays and preliminary analysis of Arbuthnot’s baptism data. Your assignment involves repeating these steps, but for present day birth records in the United States. Load up the present day data into a `DataFrame` called `present` with the following command.
present = pd.read_csv('present.csv')
# 1. What years are included in this data set? What are the dimensions of the data frame and what are the variable or column names?
# 2. How do these counts compare to Arbuthnot’s? Are they on a similar scale?
# 3. Make a plot that displays the boy-to-girl ratio for every year in the data set. What do you see? Does Arbuthnot’s observation about boys being born in greater proportion than girls hold up in the U.S.? Include the plot in your response.
# 4. In what year did we see the most total number of births in the U.S.? You can refer to the help files or the R reference card http://cran.r-project.org/doc/contrib/Short-refcard.pdf to find helpful commands.
#
# These data come from a report by the Centers for Disease Control http://www.cdc.gov/nchs/data/nvsr/nvsr53/nvsr53_20.pdf. Check it out if you would like to read more about an analysis of sex ratios at birth in the United States.
#
# That was a short introduction to python, pandas, and matplotlib, but we will provide you with more functions and a more complete sense of the language as the course progresses. Feel free to browse around the websites for python if you’re interested in learning more, or find more labs for practice at http://openintro.org.
# *This notebook is based on the OpenIntro R lab [Introduction to R and RStudio](http://htmlpreview.github.io/?https://github.com/andrewpbray/oiLabs-base-R/blob/master/intro_to_r/intro_to_r.html). *
| open-intro-statistics/python-labs/Introduction to Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # First notebook:
#
# This notebook contains a demonstration of how to open DTI from the nifti format.
# +
# Importanto as bibliotecas necessárias:
import DTIlib as DTI
from IPython.display import Image
from IPython.display import display
# %matplotlib
import matplotlib.pyplot as plt
import numpy as np
# +
# Loading the data, at this case for subject 01
BASE_PATH = 'subject001'
fa, evl, evt = DTI.load_fa_evl_et(BASE_PATH)
# -
# Printing shapes
#fa.shape
print('FA shape: ' , fa.shape)
print('Evl shape: ', evl.shape)
# ## Visualizing the data
#
# The plot shows FA image in three different views (Axial, Coronal and Sagittal)
# +
# Viewing data
# %matplotlib inline
from matplotlib.widgets import Slider
# Set up figure
sz, sy, sx = fa.shape
fig = plt.figure(figsize=(15,15))
xy = fig.add_subplot(1,3,1)
plt.title("Axial Slice")
xz = fig.add_subplot(1,3,2)
plt.title("Coronal Slice")
yz = fig.add_subplot(1,3,3)
plt.title("Sagittal Slice")
frame = 0.5
# Normalize the FA values for better visualization
maximo = np.max(np.abs(fa))
minimo = np.min(np.abs(fa))
xy.imshow(fa[np.floor(frame*sz),:,:], origin='lower', interpolation='nearest', cmap="gray",vmin=0, vmax=maximo )
xz.imshow(fa[:,np.floor(frame*sy),:], origin='lower', interpolation='nearest', cmap="gray",vmin=0 , vmax=maximo )
yz.imshow(fa[:,:,np.floor(frame*sx)], origin='lower', interpolation='nearest', cmap="gray",vmin=0 , vmax=maximo )
# -
| dev/02052017_FirstNotebook_MECP_updated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MultiresPolar
import numpy as np
import pickle
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Load Images and fixation points
# +
# This is for 100 images
#lImages = pickle.load(open("listImages.pkl","rb"))
#ldp = pickle.load(open('ldp.pkl', 'rb'))
#fixations = pickle.load(open("PredIndList.pkl","rb"))
# -
# A smaller sample of 5 images
lImages = pickle.load(open("images5.pkl","rb"))
ldp = pickle.load(open('ldp5.pkl', 'rb'))
fixations = pickle.load(open("PredIndList5.pkl","rb"))
idx = 2
plt.subplot(1,2,1); plt.imshow(lImages[idx]); plt.axis('off');
plt.plot(fixations[idx][1], fixations[idx][0], 'bo')
plt.subplot(1,2,2); plt.imshow(ldp[idx][0,:,:,0]); plt.axis('off');
# +
#plt.imsave('guitar.jpg', lImages[idx])
# -
print(fixations[idx])
print(np.shape(lImages[idx]))
diam = 100
ctr = fixations[idx]
img = lImages[idx][ctr[0]-diam:ctr[0]+diam,ctr[1]-diam:ctr[1]+diam]
print(np.shape(img))
plt.imshow(img);
# ## Set up multires pyramid
from skimage.filters import gaussian
from numpy.fft import fft2, ifft2, fftshift, ifftshift
# Blur kernel radii (FWHM)
sigmas = [1., 2, 3, 4, 7, 10., 15]
sigmas = [1., 1.5, 2, 2.5, 3, 3.5, 4, 5, 7, 10., 15]
def makeGaussian(size, fwhm = 3, center=None):
""" Make a square gaussian kernel.
size is the length of a side of the square
fwhm is full-width-half-maximum, which
can be thought of as an effective radius.
"""
x = np.arange(0, size, 1, float)
y = x[:,np.newaxis]
if center is None:
x0 = y0 = size // 2
else:
x0 = center[0]
y0 = center[1]
g = np.exp(-4*np.log(2) * ((x-x0)**2 + (y-y0)**2) / fwhm**2)
return g/np.sum(g.flatten())
def Depricated_AutoCrop(img, relthresh=0.1):
N = np.shape(img)[0]
maximums = np.max(abs(img), axis=1) / np.max(abs(img))
idx_low = list(maximums>0.1).index(True)
if idx_low==0:
return img
else:
idx_high = N - 1 - list(reversed(maximums>0.1)).index(True)
return img[idx_low:idx_high,idx_low:idx_high]
def MyNormalize(img):
img_min = np.min(img)
img_max = np.max(img)
return (img-img_min)/(img_max-img_min)
def AutoCropFilter(N, fwhm=2., relthresh=0.1):
'''
G = AutoCropFilter(N, fwhm=2., relthresh=0.1)
Create and crop a Gaussian filter. The image is returned
in the frequency domain (i.e. the Fourier transform of
the image). The image is evenly cropped so that the minimum
of the function is less than the maximum times relthresh.
Inputs:
N an integer for the maximum image size
fwhm standard deviation of the Gaussian function,
expressed as full-width-at-half-max
relthresh the threshold for the min/max of the function
Output:
G the FFT of the Gaussian function, cropped to a square
image
'''
g = makeGaussian(N, fwhm=fwhm)
G = fftshift(fft2(ifftshift(g)))
N = np.shape(G)[0]
maximums = np.max(abs(G), axis=1) / np.max(abs(G))
idx_low = list(maximums>0.1).index(True)
if idx_low==0:
return G
else:
idx_high = N - 1 - list(reversed(maximums>0.1)).index(True)
return G[idx_low:idx_high,idx_low:idx_high]
G = AutoCropFilter(255, fwhm=20, relthresh=0.05)
plt.imshow(abs(G), cmap='gray');
def MakeImagePyramid(img, sigmas):
'''
pyramid = MakeImagePyramid(img, sigmas)
Construct a list of blurred and subsampled versions of an image.
Inputs:
img square image
sigmas list of standard deviations for the Gaussian blur kernels
Output:
pyramid list of images, varying in size
'''
f_pyramid = []
F = fftshift(fft2(img, axes=[0,1]), axes=[0,1])
N = np.shape(img)[0]
chans = np.shape(img)[2]
for s in sigmas:
G = AutoCropFilter(N, fwhm=s, relthresh=0.05)
sd = int( (np.shape(F)[0] - np.shape(G)[0])/2 )
if sd<=0:
sd = 0
Fc = F.copy()
else:
Fc = F[sd:-(sd),sd:-(sd),:].copy()
for c in range(chans):
Fc[:,:,c] *= G
Nnew = np.shape(G)[0]
f_pyramid.append(np.real(ifft2(ifftshift(Fc, axes=[0,1]), axes=[0,1]))/N/N*Nnew**2)
return f_pyramid
pyr = MakeImagePyramid(img, sigmas)
plt.figure(figsize=[15,8])
blur_levels = len(sigmas)
for idx, f in enumerate(pyr):
plt.subplot(1,blur_levels,idx+1)
plt.imshow(MyNormalize(f));
plt.title(str(np.shape(f)[0])+'x'+str(np.shape(f)[1]))
# ## Polar Transform
def Polar2Cart(ctr, rt):
x = ctr[0] + rt[0]*np.cos(rt[1])
y = ctr[1] + rt[0]*np.sin(rt[1])
return (x,y)
from scipy.ndimage import map_coordinates
def PolarResample(img, r_samples, n_theta=180, ctr=None):
'''
p_img = PolarResample(img, r_samples, ctr=None)
Polar resampling of an image.
Inputs:
img image array
r_samples is a list or array of radii at which to sample
n_theta number of angles
ctr coordinates of the centre of the polar resampling
If ctr=None, then it chooses the centre pixel
ctr = floor(dim/2)
Output:
p_img an n_radius x n_theta image in polar coordinates
'''
if ctr==None:
ctr = np.array(np.array(np.shape(img)[0:2])/2, dtype=int)
#n_radius = int( (outer_radius - inner_radius)/img_scale + 1)
n_radius = len(r_samples)
#n_theta = 180
#r = np.linspace(inner_radius, outer_radius, n_radius)
theta = np.linspace(0., 2*np.pi, n_theta, endpoint=False)
rt = [np.tile(r_samples, (n_theta, 1)).T, np.tile(theta, (n_radius,1))]
xy = np.array(Polar2Cart(ctr, rt))
xy0 = np.dstack([xy[0], xy[0], xy[0]])
xy1 = np.dstack([xy[1], xy[1], xy[1]])
xy2 = np.dstack([np.zeros([n_radius,n_theta,1]),
np.ones([n_radius,n_theta,1]),
2.*np.ones([n_radius,n_theta,1])])
xy0 = xy0[np.newaxis,:,:,:]
xy1 = xy1[np.newaxis,:,:,:]
xy2 = xy2[np.newaxis,:,:,:]
xxxyyy = np.concatenate((xy0,xy1,xy2), axis=0)
p_img = map_coordinates(img, xxxyyy, mode='reflect')
return p_img, xy
[np.shape(pyr[n]) for n in range(len(sigmas))]
k = 3
k0 = float(np.shape(pyr[0])[0])
kk = float(np.shape(pyr[k])[0])
s = kk / k0
print(k0, kk, s)
blahk = PolarResample(pyr[k], np.linspace(0, 100*s, 101))
s = np.array([float(np.shape(p)[0]) for p in pyr])
s = s / s[0]
print(s)
rgc_spacing = np.load('rgc_spacing.npy')
plt.plot(rgc_spacing[0], rgc_spacing[1]);
myx = np.linspace(0, 45, 10)
def f(x):
return np.interp(x, rgc_spacing[0], rgc_spacing[1])
plt.plot(myx, f(myx), 'o');
plt.xlabel('Eccentricity');
plt.ylabel('Degrees per Sample');
# Get a list of sample locations (eccentricities) that follow the spacing.
samp_next = 0.
samp = [samp_next]
scale = 5. # degrees per pixel
for k in range(100):
samp_next += f(samp_next)*scale
samp.append(samp_next)
plt.plot(samp, '.')
plt.xlabel('Sample Number')
plt.ylabel('Eccentricity');
annuli = [(20, 25), (40, 25), (50, 25), (60,25), (70, 25), (80, 25), (100, 25)]
annuli = [(10,10), (20,10), (30,10), (40,10), (50,10),
(60,10), (70,10), (80,10), (90,10), (100,10), (110,10)]
p_img = np.array([[]]) #PolarResample(pyr[0], np.linspace(0, annuli[0][0], annuli[0][1]))
old_r = 0.
xxyy = []
for k in range(len(s)):
ss = s[k]
r = annuli[k][0]
n = annuli[k][1]
pp = pyr[k]
if k==0:
p_img, xy = PolarResample(pp, np.linspace(old_r, r, n, endpoint=False)*ss)
else:
blah, xy = PolarResample(pp, np.linspace(old_r, r, n, endpoint=False)*ss)
p_img = np.concatenate((p_img, blah), axis=0)
xxyy.append(xy/ss)
old_r = r
plt.figure(figsize=[20,10])
plt.subplot(1,2,2); plt.imshow(MyNormalize(p_img));
plt.subplot(1,2,1); plt.imshow(MyNormalize(pyr[0]));
for xy in xxyy:
plt.plot(xy[0], xy[1], 'k.', markersize=1.);
| sample_images/MultiresPolar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing the query API for the EP-full-text data
# ## Imports
import pandas as pd
import sys
sys.path.append("../")
# import custom modules
#from src.ep_text_util import test_query_ep_full_text as api
from src.ep_text_util import query_full_text as api
# ## Location of the data
# location of the PATSTAT data previously retrieved with the data_extraction_from_PATSTAT.ipynb notebook
output_files_prefix = 'test_query'
pre = '../data/raw/' + output_files_prefix
suf = '.csv'
# location of the patstat database
path_patstat = r'../data/ep_full_text_database/2020_edition/EP'
# ## Load data
# load PATSTAT data
table_main_patent_infos = pd.read_csv(pre + '_table_main_patent_infos' + suf, low_memory=False)
# ## Retrieve data from the EP-full text data using the custom API
# +
# %%time
API = api.ApiEpoFullText()
API.fit_patstat_data(path = path_patstat, patstat_data = table_main_patent_infos)
full_text_data = API.query()
# -
# ## Store the result in a csv file
file = pre + '_full_text' + suf # where to save
print('Saving results in {}'.format(file))
full_text_data.to_csv(file)
| notebooks/query_full_text_with_API.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy as sc
from scipy.signal import savgol_filter
from matplotlib.pyplot import figure
import seaborn as sns
import math
# %matpscipy.signal.residuez
# ## Load data
data = pd.read_csv("../Data/test.txt")
data.head()
dataList = data.iloc[0]
# ## Choosing filter
dataList = abs(dataList[:100])
dataList.shape[0]
filteringData1 = savgol_filter(dataList,5,2)
filteringData2 = savgol_filter(dataList,7,2)
filteringData3 = savgol_filter(dataList,9,2)
filteringData4 = savgol_filter(dataList,11,2)
filteringData1.shape
# +
figure(figsize=(15, 7))
plt.xticks(np.arange(0, 100, 10))
plt.subplot(2,1,2)
plt.plot(dataList,label='Absolute values of raw data')
plt.plot(filteringData2,label='Savitskiy-Golay (7 points)')
plt.xticks(np.arange(0, 100, 10))
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
figure(figsize=(15, 7))
plt.subplot(2,1,1)
plt.plot(dataList,label='Absolute values of raw data')
plt.plot(filteringData3,label='Savitskiy-Golay (9 points)')
plt.xticks(np.arange(0, 100, 10))
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
plt.subplot(2,1,2)
plt.plot(dataList,label='Absolute values of raw data')
plt.plot(filteringData4,label='Savitskiy-Golay (11 points)')
plt.xticks(np.arange(0, 100, 10))
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
# -
def sigmaValue(inp):
length = inp.shape[0]
sigma = 0
for i in inp:
sigma += i*i
sigma /= length
sigma = math.sqrt(sigma)
return sigma
def meanValue(inp):
mean = 0
length = inp.shape[0]
for i in inp:
mean += i
mean /= length
return mean
def standartization(inp):
mean = meanValue(inp)
sigma = sigmaValue(inp)
length = inp.shape[0]
output = np.zeros(length)
for i in range(length):
output[i] = (inp[i] - mean)/sigma
return output
def movingAverageFilter(inp,n):
inp = abs(inp)
length = inp.shape[0]
output = np.zeros(length)
for i in range(length):
current = 0
for j in range(n):
ind = int(i-(n-1)/2+j)
if ((ind>-1) and (ind<length)):
current += inp[ind]/n
output[i] = current
return output
maf3 = movingAverageFilter(dataList,3)
maf5 = movingAverageFilter(dataList,5)
maf7 = movingAverageFilter(dataList,7)
# +
figure(figsize=(15, 7))
plt.subplot(2,1,1)
plt.plot(dataList,label='Absolute values of raw data')
plt.plot(filteringData2,label='Savitskiy-Golay (7 points)')
plt.plot(maf3,label='Moving average (3 points)')
plt.xticks(np.arange(0, 100, 10))
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
plt.subplot(2,1,2)
plt.plot(dataList,label='Absolute values of raw data')
plt.plot(filteringData3,label='Savitskiy-Golay (9 points)')
plt.plot(maf5,label='Moving average (5 points)')
plt.xticks(np.arange(0, 100, 10))
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
figure(figsize=(15, 7))
plt.subplot(2,1,1)
plt.plot(dataList,label='Absolute values of raw data')
plt.plot(filteringData4,label='Savitskiy-Golay (11 points)')
plt.plot(maf7,label='Moving average (7 points)')
plt.xticks(np.arange(0, 100, 10))
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
# -
data1 = data.iloc[43]
data2 = data.iloc[24]
data3 = data.iloc[78]
filtered1 = movingAverageFilter(data1,5)
filtered2 = movingAverageFilter(data2,5)
filtered3 = movingAverageFilter(data3,5)
st = standartization(filtered1)
# +
figure(figsize=(15, 7))
plt.subplot(2,1,1)
sns.distplot(filtered1,label = 'Without normalization');
plt.ylabel("Probability density",fontsize = 12)
plt.xlabel("Signal value, [MU]",fontsize = 12)
plt.legend(prop={'size': 15})
plt.subplot(2,1,2)
sns.distplot(st,label = 'With normalization');
plt.ylabel("Probability density",fontsize = 12)
plt.xlabel("Signal value, [Normalized units]",fontsize = 12)
plt.legend(prop={'size': 15})
# -
std1 = standartization(filtered1)
std2 = standartization(filtered2)
std3 = standartization(filtered3)
# +
figure(figsize=(15, 7))
plt.subplot(2,1,1)
plt.plot(std1)
plt.plot(std2)
plt.plot(std3)
plt.axis('off')
# -
example = np.array([np.random.randint(3,15, size=100),dataList,np.random.randint(3,15, size=100)]).reshape(300)
example = movingAverageFilter(example,5)
summ = np.zeros(300)
diff = np.zeros(300)
for i in range(300):
if i>99:
summ[i]= np.sum(example[i-100:i])
else:
summ[i]= np.sum(example[0:i])
for i in range(300):
if i>0:
diff[i] = summ[i]-summ[i-1]
# +
figure(figsize=(15, 7))
plt.subplot(2,1,2)
plt.plot(example,label='Filtered signal')
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
figure(figsize=(15, 7))
plt.subplot(2,1,1)
plt.plot(summ,label='Sum',color = 'm')
plt.plot(np.zeros(300)+2000,'--',color = 'r',label='Threshold for detecting')
plt.ylabel("Signal sum, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
plt.subplot(2,1,2)
plt.plot(diff,label='Derivative',color = 'g')
plt.plot(np.zeros(300),'--',color = 'c')
plt.plot(180, 0, 'o',label='Zero intersect',color = 'r')
plt.ylabel("Signal sum derivative, [MU MF]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
plt.legend(prop={'size': 12})
#plt.axis('off')
# -
historyarduino4new = pd.read_csv("historyarduino4new.csv")
historyarduino4old = pd.read_csv("historyarduino4old.csv")
historyarduino7 = pd.read_csv("historyarduino7.csv")
historyPCbig = pd.read_csv("historyPCbig.csv")
historyPCsmall = pd.read_csv("historyPCsmall.csv")
# ## PCbig
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(9,9))
plt.subplot(2,1,1)
plt.plot(historyPCbig['loss'])
plt.plot(historyPCbig['val_loss'])
plt.legend(["Train","Validation"])
plt.ylabel("Loss",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
plt.subplot(2,1,2)
plt.plot(historyPCbig['accuracy'])
plt.plot(historyPCbig['val_accuracy'])
plt.legend(["Train","Validation"])
plt.ylabel("Accuracy",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
# ## PCsmall
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(9,9))
plt.subplot(2,1,1)
plt.plot(historyPCsmall['loss'])
plt.plot(historyPCsmall['val_loss'])
plt.legend(["Train","Validation"])
plt.ylabel("Loss",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
plt.subplot(2,1,2)
plt.plot(historyPCsmall['accuracy'])
plt.plot(historyPCsmall['val_accuracy'])
plt.legend(["Train","Validation"])
plt.ylabel("Accuracy",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
# ## Arduino4
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(9,9))
plt.subplot(2,1,1)
plt.plot(historyarduino4new['loss'])
plt.plot(historyarduino4new['val_loss'])
plt.legend(["Train","Validation"])
plt.ylabel("Loss",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
plt.subplot(2,1,2)
plt.plot(historyarduino4new['accuracy'])
plt.plot(historyarduino4new['val_accuracy'])
plt.legend(["Train","Validation"])
plt.ylabel("Accuracy",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
# ## Arduino7
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(9,9))
plt.subplot(2,1,1)
plt.plot(historyarduino7['loss'])
plt.plot(historyarduino7['val_loss'])
plt.legend(["Train","Validation"])
plt.ylabel("Loss",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
plt.subplot(2,1,2)
plt.plot(historyarduino7['accuracy'])
plt.plot(historyarduino7['val_accuracy'])
plt.legend(["Train","Validation"])
plt.ylabel("Accuracy",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(9,9))
plt.subplot(2,1,1)
plt.plot(historyarduino4new['val_accuracy'])
plt.plot(historyarduino4old['val_accuracy'])
plt.legend(["Validation accuracy with index finger","Validation accuracy with goat"])
plt.ylabel("Accuracy",fontsize = 12)
plt.xlabel("Training duration, [epochs]",fontsize = 12)
data = pd.read_csv("../Data/test0.txt")
data.head()
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(9,9))
plt.subplot(2,1,1)
plt.plot(data['17'][0:200])
plt.ylabel("Signal value, [MU]",fontsize = 12)
plt.xlabel("Measuring, [sample]",fontsize = 12)
| Python/Preprocessing/.ipynb_checkpoints/visualize-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Sample API
import flowx
flowx.__version__
# +
# Define grid parameters
nx, ny = 40, 40
xmin, xmax = 0.0, 1.0
ymin, ymax = 0.0, 1.0
Re = 100.0
dt = 0.01
tmax = 2.
# Define cell-centered variable names
center_vars = ['pres', 'divv', 'asol', 'eror']
face_vars = ['velc', 'hvar', 'asol', 'eror']
# Define boundary conditions for variable pressure and velocity [left, right, bottom, top]
bc_type_pres = {'pres': ['neumann', 'neumann', 'neumann', 'neumann']}
bc_val_pres = {'pres': [0.0, 0.0, 0.0, 0.0]}
bc_type_u = {'velc': ['dirichlet', 'dirichlet', 'dirichlet', 'dirichlet']}
bc_val_u = {'velc': [0.0, 0.0, 0.0, 1.0]}
bc_type_v = {'velc': ['dirichlet', 'dirichlet', 'dirichlet', 'dirichlet']}
bc_val_v = {'velc': [0.0, 0.0, 0.0, 0.0]}
# Create the grid and data
gridc = flowx.Grid('cell-centered', center_vars,
nx, ny, xmin, xmax, ymin, ymax,
user_bc_type=bc_type_pres, user_bc_val=bc_val_pres)
gridx = flowx.Grid('x-face', face_vars,
nx, ny, xmin, xmax, ymin, ymax,
user_bc_type=bc_type_u, user_bc_val=bc_val_u)
gridy = flowx.Grid('y-face', face_vars,
nx, ny, xmin, xmax, ymin, ymax,
user_bc_type=bc_type_v, user_bc_val=bc_val_v)
scalars = flowx.Scalars(tmax=2.0, dt=0.01, Re=100.0)
# +
ins_vars = ['velc', 'hvar', 'divv', 'pres']
poisson_options = dict(maxiter = 2000, tol = 1e-9)
poisson_options['lu'], poisson_options['mtx'] = flowx.poisson.build_sparse(gridc, 'pres')
# AB2 starter
# Predictor Step
flowx.ins.advance_euler(gridc, gridx, gridy, scalars, ins_vars, 'predictor')
# Divergence Step
flowx.ins.advance_euler(gridc, gridx, gridy, scalars, ins_vars, 'divergence')
# Solve the pressure Poisson equation
scalars.stats['ites'], scalars.stats['res'] = flowx.poisson.solve_lu(gridc, 'pres', 'divv', poisson_options)
# Corrector Step
flowx.ins.advance_euler(gridc, gridx, gridy, scalars, ins_vars, 'corrector')
scalars.advance()
while scalars.variable['time'] <= scalars.variable['tmax']:
# Predictor Step
flowx.ins.ab2(gridc, gridx, gridy, scalars, ins_vars, 'predictor')
# Divergence Step
flowx.ins.ab2(gridc, gridx, gridy, scalars, ins_vars, 'divergence')
# Solve the pressure Poisson equation
scalars.stats['ites'], scalars.stats['res'] = flowx.poisson.solve_lu(gridc, 'pres', 'divv',
poisson_options)
# Corrector Step
flowx.ins.ab2(gridc, gridx, gridy, scalars, ins_vars, 'corrector')
# Display stats
if scalars.variable['nstep'] % 10 == 0:
flowx.io.display_stats(scalars)
scalars.advance()
# -
flowx.io.plot_contour(gridc, 'pres')
flowx.io.plot_vector(gridx, gridy,'velc')
| examples/ins/lid_driven_cavity/ins_ab2_explicit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import ETL as etl
# %matplotlib inline
from datetime import datetime
import pandas as pd
# Carregando dados
investimentos = etl.getPlanilhaInvestimentos()
fi = investimentos[0]
aportes = investimentos[1]
acoes = investimentos[2]
rendaFixa = investimentos[3]
fii = investimentos[4]
# +
## ANALISE EXPLORATORIA
# -
pd.DataFrame(fi).fillna(0).head()
# dataset de fundo de investimento
fi.head()
# Verificarndo se os tipos de dados estão corretos de cada atributo/coluna
fi.dtypes
# Total Investido.
sum(aportes['Valor'])
# Total Atual em FI
dtUltimoRegistro = max(fi['DATA'])
totalAtualFI = sum(fi[fi['DATA'] == dtUltimoRegistro]['Valor Atual'])
totalAtualFI
# Total Atual em Ações
acoesNaCarteira = acoes[pd.isnull(acoes['Data Venda'])]
totalAtualAcoes = sum(acoesNaCarteira['TotalVendido'])
totalAtualAcoes
# Total Atual em Renda Fixa
dtUltimoRegistro = max(rendaFixa['DATA'])
totalAtualRendaFixa = sum(rendaFixa[rendaFixa['DATA'] == dtUltimoRegistro]['Valor Atual'])
totalAtualRendaFixa
FIINaCarteira = fii[pd.isnull(fii['Data Venda'])]
totalAtualFII = sum(FIINaCarteira['Total Venda/Atual'])
totalAtualFII
totalAtualAcoes + totalAtualFI + totalAtualRendaFixa + totalAtualFII
nomeFundos = fi['FUNDO'].unique()
nomeFundos = pd.DataFrame(nomeFundos)
nomeFundos['Corretora'] = 'XP Investimentos'
etl.limpaFI(fi)
import pymysql
conn = pymysql.connect(host='localhost', user='root', passwd='<PASSWORD>', db='sys')
cur = conn.cursor()
cur.execute("CREATE SCHEMA `independenciafinanceira`;")
| TestNotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests # library to handle requests
import pandas as pd # library for data analsysis
import numpy as np # library to handle data in a vectorized manner
import random # library for random number generation
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
import json # library to handle JSON files
# #!conda install -c conda-forge geopy --yes
from geopy.geocoders import Nominatim # convert an address into latitude and longitude values
# Matplotlib and associated plotting modules
import matplotlib.cm as cm
import matplotlib.colors as colors
from matplotlib import pyplot as plt
# import k-means from clustering stage
from sklearn.cluster import KMeans
from sklearn import metrics
# #!conda install -c districtdatalabs yellowbrick
#from yellowbrick.cluster import KElbowVisualizer
# #!conda install -c conda-forge folium=0.5.0 --yes
import folium # map rendering library
print('Libraries imported.')
# +
### Pull CSV data and observe categories
data_sf=pd.read_csv(r'C:/Users/schne/Desktop/IBM Data Science Certification/Python v2/Applied Data Science Capstone/Notebooks/Cities/San Francisco/sanfran_data.csv')
data_sf=data_sf.drop(columns='Unnamed: 0').reset_index(drop=True)
# +
# Use Nominatim geocoder to get coodinates of Toronto
address = 'San Francisco, USA'
geolocator = Nominatim(user_agent="on_explorer")
location = geolocator.geocode(address)
latitude = location.latitude
longitude = location.longitude
#print geocoordinates
print('The geograpical coordinate of San Francisco, California are {}, {}.'.format(latitude, longitude))
# create map of Toronto using latitude and longitude values
map_sanfran = folium.Map(location=[latitude, longitude], zoom_start=10)
# add markers to map
for lat, lng, neighborhood, borough in zip(data_sf['Latitude'], data_sf['Longitude'], data_sf['Neighborhood'],data_sf['Borough']):
label = '{}, {}'.format(neighborhood,borough)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_sanfran)
map_sanfran
# -
# ## Pull Foursquare Data using Rest API
### Open Foursquare Credentials
with open (r'C:\Users\schne\Desktop\IBM Data Science Certification\Python v2\Applied Data Science Capstone\fsquarecreds.json') as f:
data = json.load(f)
fsquare_creds = {'CLIENT_ID':data['CLIENT_ID'],'CLIENT_SECRET':data['CLIENT_SECRET'],'VERSION':data['VERSION']}
# +
#Define function to get nearby venues based on Neighborhood Lat/Lng
limit = 100
def getNearbyVenues(names, latitudes, longitudes, radius=500):
venues_list=[]
for name, lat, lng in zip(names, latitudes, longitudes):
print(name)
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
fsquare_creds['CLIENT_ID'],
fsquare_creds['CLIENT_SECRET'],
fsquare_creds['VERSION'],
lat,
lng,
radius,
limit)
# make the GET request
results = requests.get(url).json()['response']['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['name'],
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
#code to run the above function on each neighborhood and create a new dataframe called *df_venues*
data_sf_venues = getNearbyVenues(names=data_sf['Neighborhood'],
latitudes=data_sf['Latitude'],
longitudes=data_sf['Longitude'])
# -
#Print shape of dataframe and show head
print(data_sf_venues.shape)
print(data_sf_venues.head())
print(len(data_sf_venues['Neighborhood'].unique()))
# Let's check how many venues were returned for each neighborhood. Let's also look at the number of unqiue venue categories
# +
## Number of Venues for each Neighborhood
print(data_sf_venues.groupby('Neighborhood').count().head())
##Check number of Neighborhoods
print('There are {} unique Neighborhoods.'.format(len(data_sf_venues['Neighborhood'].unique())))
## Number of Unique Categoriges
print('There are {} unique categories.'.format(len(data_sf_venues['Venue Category'].unique())))
# -
# ### Format Data for K-Means Clustering
# +
# one hot encoding
data_sf_onehot = pd.get_dummies(data_sf_venues['Venue Category'], prefix="", prefix_sep=" ")
# add neighborhood column back to dataframe
data_sf_onehot['Neighborhood'] = data_sf_venues['Neighborhood']
data_sf_onehot.head()
# move neighborhood column to the first column
fixed_columns = [data_sf_onehot.columns[-1]] + list(data_sf_onehot.columns[:-1])
print(fixed_columns)
data_sf_onehot = data_sf_onehot[fixed_columns]
data_sf_onehot.head()
print(data_sf_onehot.shape)
data_sf_grouped = data_sf_onehot.groupby('Neighborhood').mean().reset_index()
data_sf_grouped.head()
# +
#Function to grab the top 5 venues in a dataframe
num_top_venues = 5
for hood in data_sf_grouped['Neighborhood']:
print("----"+hood+"----")
temp = data_sf_grouped[data_sf_grouped['Neighborhood'] == hood].T.reset_index()
temp.columns = ['venue','freq']
temp = temp.iloc[1:]
temp['freq'] = temp['freq'].astype(float)
temp = temp.round({'freq': 2})
print(temp.sort_values('freq', ascending=False).reset_index(drop=True).head(num_top_venues))
print('\n')
# +
# function to return most common venues
def return_most_common_venues(row, num_top_venues):
row_categories = row.iloc[1:]
row_categories_sorted = row_categories.sort_values(ascending=False)
return row_categories_sorted.index.values[0:num_top_venues]
#grab the top 10 venues
num_top_venues = 10
indicators = ['st', 'nd', 'rd']
# create columns according to number of top venues
columns = ['Neighborhood']
for ind in np.arange(num_top_venues):
try:
columns.append('{}{} Most Common Venue'.format(ind+1, indicators[ind]))
except:
columns.append('{}th Most Common Venue'.format(ind+1))
columns
# create a new dataframe
neighborhoods_venues_sorted = pd.DataFrame(columns=columns)
neighborhoods_venues_sorted['Neighborhood'] = data_sf_grouped['Neighborhood']
#add common venues into new dataframe
for ind in np.arange(data_sf_grouped.shape[0]):
neighborhoods_venues_sorted.iloc[ind, 1:] = return_most_common_venues(data_sf_grouped.iloc[ind, :], num_top_venues)
neighborhoods_venues_sorted.head()
# -
# ## K-Means Clustering
# ### Check for Optimal K (elbow Method)
# +
# Remove column 'Neighborhood' from grouped data
data_sf_grouped_clustering = data_sf_grouped.drop('Neighborhood', 1)
# Instantiate the clustering model and visualizer
model = KMeans().fit(data_sf_grouped_clustering)
#visualizer = KElbowVisualizer(model, k=(1,10))
#visualizer.fit(data_sf_grouped_clustering) # Fit the data to the visualizer
#visualizer.show()
# +
sum_of_squared_distances = []
K = range(1,15)
for k in K:
k_means = KMeans(n_clusters=k)
model = k_means.fit(data_sf_grouped_clustering)
sum_of_squared_distances.append(k_means.inertia_)
plt.plot(K, sum_of_squared_distances, 'bx-')
plt.xlabel('k')
plt.ylabel('sum_of_squared_distances')
plt.title('elbow method for optimal k')
plt.show()
# -
# #### Elbow method isn't appreaing to be to accurate, so let's go with 5, the same number used in the New York dataset. exponential nature of distortion makes for suspect of validity of using K-Means Clustinger
# +
# set number of clusters
kclusters = 6
# run k-means clustering
kmeans = KMeans(n_clusters=kclusters,random_state=0).fit(data_sf_grouped_clustering)
# check cluster labels generated for each row in the dataframe
kmeans.labels_[0:10]
# add clustering labels #Comment this cell when run
neighborhoods_venues_sorted.insert(0, 'Cluster Labels', kmeans.labels_)
neighborhoods_venues_sorted.head()
# +
# merge toronto_grouped with data_sf to add latitude/longitude for each neighborhood
data_sf_merged = data_sf
data_sf_merged = data_sf_merged.join(neighborhoods_venues_sorted.set_index('Neighborhood'), on='Neighborhood')
# check the last columns!
print(data_sf_merged.shape)
data_sf_merged.tail()
# +
#Remove N/A Values
print(data_sf_merged.shape)
data_sf_merged=data_sf_merged.dropna()
print(data_sf_merged.shape)
# check the data type
data_sf_merged['Cluster Labels']=data_sf_merged['Cluster Labels'].astype(int)
# -
data_sf_merged.dtypes
# +
# create map
map_clusters = folium.Map(location=[latitude, longitude], zoom_start=11)
# set color scheme for the clusters
x = np.arange(kclusters)
ys = [i + x + (i*x)**2 for i in range(kclusters)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
# add markers to the map
markers_colors = []
for lat, lon, poi, cluster in zip(data_sf_merged['Latitude'], data_sf_merged['Longitude'], data_sf_merged['Neighborhood'], data_sf_merged['Cluster Labels']):
label = folium.Popup(str(poi) + ' : Cluster ' + str(cluster), parse_html=True)
folium.CircleMarker(
[lat, lon],
radius=5,
popup=label,
color=rainbow[cluster-1],
fill=True,
fill_color=rainbow[cluster-1],
fill_opacity=0.7).add_to(map_clusters)
map_clusters
# -
# # Cluster 1
data_sf_merged.loc[data_sf_merged['Cluster Labels'] == 0, data_sf_merged.columns[[0] + list(range(5, data_sf_merged.shape[1]))]]
# # Cluster 2
data_sf_merged.loc[data_sf_merged['Cluster Labels'] == 1, data_sf_merged.columns[[0] + list(range(5, data_sf_merged.shape[1]))]]
# # Cluster 3
data_sf_merged.loc[data_sf_merged['Cluster Labels'] == 2, data_sf_merged.columns[[0] + list(range(5, data_sf_merged.shape[1]))]]
# # Cluster 4
data_sf_merged.loc[data_sf_merged['Cluster Labels'] == 3, data_sf_merged.columns[[0] + list(range(5, data_sf_merged.shape[1]))]]
# # Cluster 5
data_sf_merged.loc[data_sf_merged['Cluster Labels'] == 4, data_sf_merged.columns[[0] + list(range(5, data_sf_merged.shape[1]))]]
# +
data_sf_merged.loc[data_sf_merged['Cluster Labels'] == 5, data_sf_merged.columns[[0] + list(range(64, data_sf_merged.shape[1]))+[1,2,8,10,13,14]]]
data_sf_merged.loc[data_sf_merged['Cluster Labels'] == 6, data_sf_merged.columns[[0] + list(range(64, data_sf_merged.shape[1]))+[1,2,8,10,13,14]]]
| Contents/Cities/San Francisco/sanfran_fsquare+cluster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example usage of a (skeleton for a) diffable HEP library
# +
import jax
import jax.numpy as jnp
import numpy as np
import pyhf
# use jax backend of pyhf
pyhf.set_backend(pyhf.tensor.jax_backend())
import smooth
import smooth.infer
# -
# ## Implementing Alex's differentiable cut example, using full pyhf:
# +
nBg = 8000
nSig = 300
background = np.random.normal(40, 10, nBg)
signal = np.random.normal(50, 5, nSig)
def generate_data():
return signal, background
# + jupyter={"source_hidden": true}
import matplotlib.pyplot as plt
bins = np.linspace(0, 80, 40)
plt.figure(dpi=120)
ax = plt.gca()
plt.hist([background, signal], bins=bins, stacked=True, label=["background", "signal"])
ax.set(ylabel='frequency',xlabel='x')
plt.legend();
# -
def preprocess(data_generator):
def counts(cut_param):
s, b = data_generator()
s_counts = smooth.cut(s,'>',cut_param).sum()
b_counts = smooth.cut(b,'>',cut_param).sum()
return jnp.array([s_counts]), jnp.array([b_counts])
return counts
def simple_histosys(data_with_cuts, uncert):
"""
Makes a histosys model with up/down yields of +- bkg*(uncert/2).
"""
def from_spec(yields):
s, b = yields
bup, bdown = b * (1 + (uncert / 2)), b * (1 - (uncert / 2))
spec = {
"channels": [
{
"name": "smoothcut",
"samples": [
{
"name": "signal",
"data": s,
"modifiers": [
{"name": "mu", "type": "normfactor", "data": None}
],
},
{
"name": "bkg",
"data": b,
"modifiers": [
{
"name": "artificial_histosys",
"type": "histosys",
"data": {"lo_data": bdown, "hi_data": bup,},
}
],
},
],
},
],
}
return pyhf.Model(spec)
def model_maker(cut_pars):
s, b = data_with_cuts(cut_pars)
# make statistical model with pyhf
m = from_spec([s, b])
nompars = m.config.suggested_init()
bonlypars = jnp.asarray([x for x in nompars])
bonlypars = jax.ops.index_update(bonlypars, m.config.poi_index, 0.0)
return m, bonlypars
return model_maker
# +
data_with_cuts = preprocess(generate_data)
model_maker = simple_histosys(data_with_cuts, uncert=0.05)
loss = smooth.infer.expected_pvalue_upper_limit(model_maker,
solver_kwargs=dict(pdf_transform=True))
jax.value_and_grad(loss)(1.,40.)
# -
# ### (Can then do gradient descent etc! I've left that out here.)
# ## Reimplementing neos:
def blobs(NMC=500, sig_mean = [-1, 1], b1_mean=[2.5, 2], b_mean=[1, -1], b2_mean=[-2.5, -1.5]):
def generate_blobs():
bkg_up = np.random.multivariate_normal(b1_mean, [[1, 0], [0, 1]], size=(NMC,))
bkg_down = np.random.multivariate_normal(b2_mean, [[1, 0], [0, 1]], size=(NMC,))
bkg_nom = np.random.multivariate_normal(b_mean, [[1, 0], [0, 1]], size=(NMC,))
sig = np.random.multivariate_normal(sig_mean, [[1, 0], [0, 1]], size=(NMC,))
return sig, bkg_nom, bkg_up, bkg_down
return generate_blobs
def hists(data_generator, predict, bins, bandwidth, LUMI=10, sig_scale = 2, bkg_scale = 10):
def hist_maker(nn):
s, b_nom, b_up, b_down = data_generator()
NMC = len(s)
nn_s, nn_b_nom, nn_b_up, nn_b_down = (
predict(nn, s).ravel(),
predict(nn, b_nom).ravel(),
predict(nn, b_up).ravel(),
predict(nn, b_down).ravel(),
)
kde_counts = jax.numpy.asarray([
smooth.hist(nn_s, bins, bandwidth) * sig_scale / NMC * LUMI,
smooth.hist(nn_b_nom, bins, bandwidth) * bkg_scale / NMC * LUMI,
smooth.hist(nn_b_up, bins, bandwidth) * bkg_scale / NMC * LUMI,
smooth.hist(nn_b_down, bins, bandwidth) * bkg_scale / NMC * LUMI,
])
return kde_counts
return hist_maker
def nn_histosys(histogram_maker):
def from_spec(yields):
s, b, bup, bdown = yields
spec = {
"channels": [
{
"name": "nn",
"samples": [
{
"name": "signal",
"data": s,
"modifiers": [
{"name": "mu", "type": "normfactor", "data": None}
],
},
{
"name": "bkg",
"data": b,
"modifiers": [
{
"name": "nn_histosys",
"type": "histosys",
"data": {
"lo_data": bdown,
"hi_data": bup,
},
}
],
},
],
},
],
}
return pyhf.Model(spec)
def nn_model_maker(nn):
yields = histogram_maker(nn)
m = from_spec(yields)
nompars = m.config.suggested_init()
bonlypars = jax.numpy.asarray([x for x in nompars])
bonlypars = jax.ops.index_update(bonlypars, m.config.poi_index, 0.0)
return m, bonlypars
return nn_model_maker
# +
import jax.experimental.stax as stax
# regression net
init_random_params, predict = stax.serial(
stax.Dense(1024),
stax.Relu,
stax.Dense(1024),
stax.Relu,
stax.Dense(1),
stax.Sigmoid
)
# choose hyperparams
bins = np.linspace(0,1,4)
centers = bins[:-1] + np.diff(bins)/2.
bandwidth = 0.8 * 1/(len(bins)-1)
# compose functions to define workflow
data = blobs()
hmaker = hists(data,predict,bins=bins,bandwidth=bandwidth)
model = nn_histosys(hmaker)
loss = smooth.infer.expected_pvalue_upper_limit(model, solver_kwargs=dict(pdf_transform=True))
_, network = init_random_params(jax.random.PRNGKey(13), (-1, 2))
jax.value_and_grad(loss, argnums=1)(1.0, network)
# +
import jax.experimental.optimizers as optimizers
import time
opt_init, opt_update, opt_params = optimizers.adam(1e-3)
def train_network(N):
cls_vals = []
_, network = init_random_params(jax.random.PRNGKey(1), (-1, 2))
state = opt_init(network)
losses = []
# parameter update function
def update_and_value(i, opt_state, mu):
net = opt_params(opt_state)
value, grad = jax.value_and_grad(loss,argnums=1)(mu, net)
return opt_update(i, grad, state), value, net
for i in range(N):
start_time = time.time()
state, value, network = update_and_value(i,state,1.0)
epoch_time = time.time() - start_time
losses.append(value)
metrics = {"loss": losses}
yield network, metrics, epoch_time
# +
maxN = 10 # make me bigger for better results!
# Training
for i, (network, metrics, epoch_time) in enumerate(train_network(maxN)):
print(f"epoch {i}:", f'p_mu = {metrics["loss"][-1]:.5f}, took {epoch_time:.2f}s')
# -
| example_smooth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 从网页中截取文章内容
# ## 步骤1:网址
url = 'http://blog.jobbole.com/105602/'
# ## 步骤2:查找相应的截取关键字
if 'blog.jobbole.com' in url: # 伯乐在线
title_key = '.entry-header'
content_key = '.entry'
elif 'blog.csdn.net' in url: # csdn
title_key = '.link_title'
content_key = '#article_content'
elif 'www.codingpy.com' in url: # 编程派网址
title_key = '.header h1'
content_key = '.article-content'
elif 'www.infoq.com' in url: # InfoQ
title_key = '.title_canvas'
content_key = '.text_info_article'
else:
title_key = ''
content_key = ''
print("标题提取键: " + title_key)
print("内容提取键: " + content_key)
# ## 步骤3:发出网页请求,接收响应
from urllib import request
req = request.Request(url)
res = request.urlopen(req)
html = res.read().decode('utf-8')
import re
pattern = r'<body[\s\S]*?</body>' # 只选择body部分内容内容
body = re.findall(pattern, html)[0]
pattern = r'<script [\s\S]*?</script>' # 去掉脚本语句
body = re.sub(pattern, '', body)
print(body)
array = []
for eachline in body.split('\n'): # 去掉空行
eachline = eachline.strip()
if eachline:
array.append(eachline + '\n')
new_body = ''.join(array)
print(new_body)
# ## 步骤4:从网页中提取文章标题和内容
from bs4 import BeautifulSoup
soup = BeautifulSoup(new_body, 'html.parser')
title = soup.select(title_key)[0].text.strip() # 文章标题
print('文章标题:', title)
content = soup.select(content_key)[0] # 文章内容
# content
# ## 步骤5:下载文章中的图片
# ### 5.1 哈希码生成函数,用于给图片重新命名
import hashlib
def md5(name):
""" 将字符串转成哈希码 """
if not isinstance(name, str):
name = str(name)
md5 = hashlib.md5()
md5.update(name.encode('utf-8'))
return md5.hexdigest()
# ### 5.2 下载图片,并修改文章图片的超链接
import re
import os
content = str(content)
pattern = '<img .*?src=\"(.*?)\"'
re_image = re.compile(pattern)
for image_link in re_image.findall(content):
if not os.path.exists('output'):
os.mkdir('output')
if not os.path.exists('output/images'):
os.mkdir('output/images')
filename = 'images/' + md5(image_link) + os.path.splitext(image_link)[-1]
try:
request.urlretrieve(image_link, 'output/' + filename)
print('下载完成', filename)
except Exception as e:
print('图片出错', e)
else:
content = content.replace(image_link, filename)
print('== 完成 ==')
# ## 步骤6:将截取的文章标题和内容重新组合成新的网页文件
html_template = """<!DOCTYPE html>
<html><head><meta charset="UTF-8">
</head><body>
<p><a href="{origin}">原文链接</a></p>
<p><center><h1>{title}</h1></center></p>
{content}
</body></html>"""
html = html_template.format(origin=url, title=title, content=content)
print(html)
# ## 步骤7:将文件写入磁盘
import codecs
filename = 'output/' + title + ".html"
with codecs.open(filename, "w", "utf-8") as f:
f.write(html)
# f.write(html.replace(u'\xa0', u' ').replace(u'\U0001f60a', u' ')) # 在windows中出错,所以这里进行了字符串替换
# ## (完)
| Webpage_Download.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Idea for spectrogram error bars
#
# 1. For each pixel get the dist that go into thwt ais plotted.
# 1. Resample each based on the dist
# 1. replot
# 1. repeat and see which are still there
# +
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import pymc3 as pm
sns.set(font_scale=1.5)
# -
with pm.Model() as model:
c_means = pm.Uniform('c_means', 0, 100, shape=5)
corrections = pm.Normal('corrs', c_means, sd=1, shape=5, observed=[3,4,5,4,3])
p_means = pm.Uniform('p_means', 0, 100, shape=5)
p = pm.Poisson('p', p_means, shape=5, observed=[30, 35, 20, 45, 16])
avg = pm.Deterministic('avg', pm.math.sum(c_means, axis=0) + pm.math.sum(p_means, axis=0)/5)
trace = pm.sample(10000)
pm.summary(trace)
pm.traceplot(trace)
pm.plot_posterior(trace)
print(np.average([30, 35, 20, 45, 16]))
# # Can we do multi-d together?
with pm.Model() as model:
p_means = pm.Uniform('p_means', 0, 100, shape=(5, 16)) # time, pixel, sector
p = pm.Poisson('p', p_means, shape=5, observed=np.random.poisson(30, size=(5, 16)))
avg = pm.Deterministic('avg', pm.math.sum(p_means)/(5*16))
trace = pm.sample(10000)
pm.summary(trace)
pm.traceplot(trace)
pm.plot_posterior(trace)
| combine_counts/Combine counts and resample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Eigen-Vectors & Eigen-Values of Stock Returns Correlations
#
# > Back in undergrad, [<NAME>](https://geology.ucdavis.edu/people/faculty/rundle) taught a graduate "Econo-Physics" class where in one lesson we learned about portfolio managment using the eigen-vectors & eigen-values correlation matrix of stock returns. Here we create some fake data and make that caluation using numpy and pandas while visualizing using matplotlib and seaborn.
#
# - toc:true
# - badges: true
# - comments: true
# - image: images/eigen_correlation_heatmap.png
# - author: nickvazz
# - categories: [stock, trading, eigenvalues, eigenvectors, kavoos]
# + [markdown] colab_type="text" id="d3QyZSbnhFev"
# > Tip: J<NAME> has a great [tutorial](https://jakevdp.github.io/PythonDataScienceHandbook/03.00-introduction-to-pandas.html) on using pandas!
# + colab={"base_uri": "https://localhost:8080/", "height": 49} colab_type="code" executionInfo={"elapsed": 713, "status": "ok", "timestamp": 1597618341747, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgChEjG1nNyKEBtqtwd8LgB4pkGSlvOqbSWMzPdcw=s64", "userId": "07489934650295771999"}, "user_tz": 420} id="NwTjPGXnPGG4" outputId="32084fc0-76d5-4474-844a-2fd53f85aaf0"
#hide_output
#collapse-hide
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from itertools import permutations
# %matplotlib inline
sns.set()
# + [markdown] colab_type="text" id="law4PLCChJKV"
# > Note: Make random Data
# + colab={"base_uri": "https://localhost:8080/", "height": 201} colab_type="code" executionInfo={"elapsed": 547, "status": "ok", "timestamp": 1597619373694, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgChEjG1nNyKEBtqtwd8LgB4pkGSlvOqbSWMzPdcw=s64", "userId": "07489934650295771999"}, "user_tz": 420} id="rqoMQx8fPO9E" outputId="d6551ba5-318e-48e5-84b6-d80b056de4dc"
np.random.seed(42)
def brownian_motion(mean,std,npts):
return np.cumsum(np.random.normal(scale=std, size=npts)) + mean
num_stocks = 10
num_timesteps = 1000
letters = 'ABCDEFGHIJKLMNOPQRSTUVWXYZ'
tickers = [''.join(x) for x in np.random.choice(list(letters), size=(num_stocks,3))]
dates = pd.date_range('2020-11-21', periods=num_timesteps, freq='D')
data = np.vstack([brownian_motion(mean, std, num_timesteps) for mean, std in zip(np.random.randint(50,200,num_stocks), np.random.randint(2,5, num_stocks))]).T
df = pd.DataFrame(data, columns=tickers, index=dates)
df = df[df > 0].dropna(axis=1)
df.head()
# + [markdown] colab_type="text" id="CeibkfOthMq-"
# > Tip: Use this to load csv's from your own google drive
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 32} colab_type="code" executionInfo={"elapsed": 786, "status": "ok", "timestamp": 1597619195342, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgChEjG1nNyKEBtqtwd8LgB4pkGSlvOqbSWMzPdcw=s64", "userId": "07489934650295771999"}, "user_tz": 420} id="4LrArbKUf8IP" outputId="e585674c-619b-42de-a409-8bf8080b9923"
# ```python
# from google.colab import drive
# drive.mount('/content/gdrive')
# # # !ls "gdrive/My Drive" # this line will look in the folder
#
# df = pd.read_csv('gdrive/My Drive/data.csv') # put the full path to the file in google drive here if you have one
# ```
# + colab={"base_uri": "https://localhost:8080/", "height": 496} colab_type="code" executionInfo={"elapsed": 2316, "status": "ok", "timestamp": 1597619258626, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgChEjG1nNyKEBtqtwd8LgB4pkGSlvOqbSWMzPdcw=s64", "userId": "07489934650295771999"}, "user_tz": 420} id="XpS2RKGbPynT" outputId="6abe4d53-846c-4e9b-8982-255cfe231338"
fig, ax = plt.subplots(1, figsize=(20,8))
df.plot(ax=ax)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 291} colab_type="code" executionInfo={"elapsed": 521, "status": "ok", "timestamp": 1597619391771, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgChEjG1nNyKEBtqtwd8LgB4pkGSlvOqbSWMzPdcw=s64", "userId": "07489934650295771999"}, "user_tz": 420} id="5kvrbMrKYM5S" outputId="3d1c09bb-44c8-4e7e-e7c3-a7a8f08c040e"
df.diff().corr()
# + colab={} colab_type="code" executionInfo={"elapsed": 300, "status": "ok", "timestamp": 1597619391772, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgChEjG1nNyKEBtqtwd8LgB4pkGSlvOqbSWMzPdcw=s64", "userId": "07489934650295771999"}, "user_tz": 420} id="GsWOy5GvaOc8"
e_val, e_vect = np.linalg.eig(df.diff().corr())
evect_df = pd.DataFrame(e_vect[np.argsort(e_val)[::-1]], columns=df.columns, index=df.columns)
evect_df
# + colab={"base_uri": "https://localhost:8080/", "height": 610} colab_type="code" executionInfo={"elapsed": 1675, "status": "ok", "timestamp": 1597619741555, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgChEjG1nNyKEBtqtwd8LgB4pkGSlvOqbSWMzPdcw=s64", "userId": "07489934650295771999"}, "user_tz": 420} id="ITL0MSE5bjx4" outputId="ee1a6242-24b5-42ba-c33f-22b6b4215d0e"
fig, ax = plt.subplots(1, figsize=(12,10))
ax.set_title('Eigenvalues of Correlation of Running Difference', fontsize=16)
sns.heatmap(evect_df, ax=ax, annot=True, fmt=".2f", linewidths=.5)
fig.savefig('../images/eigen_correlation_heatmap.png')
plt.show()
| _notebooks/2020-11-21-Finding-eigenvalues-and-eigenvectors-of-stock-returns-correlation-matrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn import linear_model
from sklearn import svm
import matplotlib.pyplot as plt
# %matplotlib inline
import os
import re
os.chdir("./Desktop/ArticleClassifier")
# Changing the kernels being used and the c values
# -
# # Preprocessing the Data
# +
# Run through 70% of the text files from each category (folder)
# 70% Training, 30% Testing for each category
#Creates training_titles and testing_tiles
training_titles_files = []
testing_titles_files = []
business_files = os.listdir("./business")
train_business = business_files[0:358]
test_business = business_files[358:] # length = 152, TOTAL = 510
training_titles_files.append(train_business)
testing_titles_files.append(test_business)
entertainment_files = os.listdir("./entertainment")
train_entertainment = entertainment_files[0:270]
test_entertainment = entertainment_files[270:] # 116, TOTAL = 386
training_titles_files.append(train_entertainment)
testing_titles_files.append(test_entertainment)
politics_files = os.listdir("./politics")
train_politics = politics_files[0:293]
test_politics = politics_files[293:] # 124, TOTAL = 417
training_titles_files.append(train_politics)
testing_titles_files.append(test_politics)
sport_files = os.listdir("./sport")
train_sport = sport_files[0:359]
test_sport = sport_files[359:] # 152, TOTAL = 511
training_titles_files.append(train_sport)
testing_titles_files.append(test_sport)
tech_files = os.listdir("./tech")
train_tech = tech_files[0:281]
test_tech = tech_files[281:] # 120, TOTAL = 401
training_titles_files.append(train_tech)
testing_titles_files.append(test_tech)
# +
training_text_file = open("trainingTextFile.txt", "w")
classes = 1
for i in training_titles_files:
folder_name = ""
if (classes == 1):
folder_name = "business"
elif (classes == 2):
folder_name = "entertainment"
elif (classes == 3):
folder_name = "politics"
elif (classes == 4):
folder_name = "sport"
elif (classes == 5):
folder_name = "tech"
for file_name in i:
file = "./" + folder_name + "/" + file_name
#print(file)
opened = open(file, "r")
training_text_file.write(opened.readline())
opened.close()
classes = classes + 1
training_text_file.close()
masterTestDataFile = open("testTextFile.txt", "w")
classes = 1
for i in testing_titles_files:
folder_name = ""
if (classes == 1):
folder_name = "business"
elif (classes == 2):
folder_name = "entertainment"
elif (classes == 3):
folder_name = "politics"
elif (classes == 4):
folder_name = "sport"
elif (classes == 5):
folder_name = "tech"
for file_name in i:
file = "./" + folder_name + "/" + file_name
#print (file)
opened = open(file, "r")
lineRead = opened.readline()
#try:
masterTestDataFile.write(lineRead)
#except UnicodeDecodeError:
opened.close()
classes = classes + 1
masterTestDataFile.close()
#currency stemming nlp
# +
training = []
testing = []
# Puts words in dictionary, key = word, value = index
trfile = open("trainingTextFile.txt", "r")
for line in trfile:
line = line.replace("'", "")
new_line = line.split()
for i in new_line:
training.append(i)
trfile.close()
words = {}
count = 0
for i in training:
if i not in words:
words[i] = count
count = count + 1
# +
#Create Training Data Matrix
d = len(words) # Number of features (unique words) we have in the dictionary
# print(d)
train_x = []
trfile = open("trainingTextFile.txt", "r")
for line in trfile:
line = line.replace("'", "")
word = line.split()
arr = [0] * d
for i in word:
index = words[i]
arr[index] = arr[index] + 1
train_x.append(arr)
trfile.close()
# -
#Target Training Vector
train_y = np.zeros((1561, 1))
for i in range(0, 1561):
if i >=0 and i < 358:
train_y[i] = 1
elif i >= 358 and i < 628:
train_y[i] = 2
elif i >= 628 and i < 921:
train_y[i] = 3
elif i >= 921 and i < 1280:
train_y[i] = 4
elif i >= 1280 and i < 1561:
train_y[i] = 5
# +
#Create Testing Data Matrix
test_x = []
testfile = open("testTextFile.txt", "r")
for line in testfile:
line = line.replace("'", "")
word = line.split()
arr = [0] * d
for i in word:
if i in words:
index = words[i]
arr[index] = arr[index] + 1
test_x.append(arr)
testfile.close()
# +
#Target Test Vector
test_y = np.zeros((664, 1))
for i in range(0, 664):
if i >= 0 and i < 152:
test_y[i] = 1
elif i >= 152 and i < 268:
test_y[i] = 2
elif i >= 268 and i < 392:
test_y[i] = 3
elif i >= 392 and i < 544:
test_y[i] = 4
elif i >= 544 and i < 664:
test_y[i] = 5
else:
test_y[i] = 1
# countzero = 0
# countone = 0
# counttwo = 0
# countthree = 0
# countfour = 0
# for i in test_y:
# if i == 0:
# countzero+=1
# if i == 1:
# countone += 1
# if i == 2:
# counttwo+=1
# if i == 3:
# countthree+=1
# if i == 4:
# countfour +=1
#print(countzero)
#print(countone)
#print(counttwo)
#print(countthree)
#print(countfour)
# y = 0 --> Business
# y = 1 --> Entertainment
# y = 2 --> Politics
# y = 3 --> Sport
# y = 4 --> Tech
# print(test_y.shape)
# +
#Training for each classification
train_y_1 = train_y.copy()
train_y_2 = train_y.copy()
train_y_3 = train_y.copy()
train_y_4 = train_y.copy()
train_y_5 = train_y.copy()
# Changing the values that aren't the classified values to 0
train_y_1[train_y_1 != 1] = 0
train_y_2[train_y_2 != 2] = 0
train_y_3[train_y_3 != 3] = 0
train_y_4[train_y_4 != 4] = 0
train_y_5[train_y_5 != 5] = 0
# Changes the correct classification values to 1 so it's easier
train_y_2[train_y_2 == 2] = 1
train_y_3[train_y_3 == 3] = 1
train_y_4[train_y_4 == 4] = 1
train_y_5[train_y_5 == 5] = 1
train_y_1_arr = []
train_y_2_arr = []
train_y_3_arr = []
train_y_4_arr = []
train_y_5_arr = []
for i in range(0, 1561):
train_y_1_arr.insert(i, int(train_y_1[i][0]))
train_y_2_arr.insert(i, int(train_y_2[i][0]))
train_y_3_arr.insert(i, int(train_y_3[i][0]))
train_y_4_arr.insert(i, int(train_y_4[i][0]))
train_y_5_arr.insert(i, int(train_y_5[i][0]))
# +
#Test for each classification
test_y_1 = test_y.copy()
test_y_2 = test_y.copy()
test_y_3 = test_y.copy()
test_y_4 = test_y.copy()
test_y_5 = test_y.copy()
# Changing the values that aren't the classified values to 0
test_y_1[test_y_1 != 1] = 0
test_y_2[test_y_2 != 2] = 0
test_y_3[test_y_3 != 3] = 0
test_y_4[test_y_4 != 4] = 0
test_y_5[test_y_5 != 5] = 0
# Changes the correct classification values to 1 so it's easier
test_y_2[test_y_2 == 2] = 1
test_y_3[test_y_3 == 3] = 1
test_y_4[test_y_4 == 4] = 1
test_y_5[test_y_5 == 5] = 1
test_y_1_arr = []
test_y_2_arr = []
test_y_3_arr = []
test_y_4_arr = []
test_y_5_arr = []
for i in range(0, 664):
test_y_1_arr.insert(i, int(test_y_1[i][0]))
test_y_2_arr.insert(i, int(test_y_2[i][0]))
test_y_3_arr.insert(i, int(test_y_3[i][0]))
test_y_4_arr.insert(i, int(test_y_4[i][0]))
test_y_5_arr.insert(i, int(test_y_5[i][0]))
# -
# # Regularization L1
prob = []
def logreg_model(c , X_train, Y_train, X_test, Y_test):
logreg = linear_model.LogisticRegression(C=c,penalty='l1', warm_start=True, solver='saga')
logreg.fit(X_train, Y_train)
Yhat_train = logreg.predict(X_train)
Yhat_test = logreg.predict(X_test)
prob.append(logreg.predict_proba(X_test))
# +
#Testing Accuracy
c = 1
logreg_model(c, train_x, train_y_1_arr, test_x, test_y_1_arr)
logreg_model(c, train_x, train_y_2_arr, test_x, test_y_2_arr)
logreg_model(c, train_x, train_y_3_arr, test_x, test_y_3_arr)
logreg_model(c, train_x, train_y_4_arr, test_x, test_y_4_arr)
logreg_model(c, train_x, train_y_5_arr, test_x, test_y_5_arr)
test_yhat_L1 = []
# test_yhat_L1_pac = []
for i in range (0, len(test_x), 1):
values = []
probability_and_classification = []
m1 = prob[0][i][1]
m2 = prob[1][i][1]
m3 = prob[2][i][1]
m4 = prob[3][i][1]
m5 = prob[4][i][1]
values.append(m1)
values.append(m2)
values.append(m3)
values.append(m4)
values.append(m5)
#max_prob = max(m1, m2, m3, m4, m5)
# probability_and_classification.append(max(values))
# probability_and_classification.append((values.index(max(values))) + 1)
# test_yhat_L1_pac.append(probability_and_classification)
test_yhat_L1.append((values.index(max(values))) + 1)
print(test_yhat_L1) # Correct
#Calculate testing accuracy
correct = 0
incorrect = 0
for i in range (0, len(test_x)):
if (test_yhat_L1[i] == test_y[i]):
correct+= 1
# print(test_yhat_L1[i], ", actual: ", test_y[i][0], "=", (test_y[i][0]))
else:
incorrect += 1
# print("INCORRECT", test_yhat_L1[i], ", actual: ", test_y[i][0], "=", (test_y[i][0]))
test_acc_L1 = correct/len(test_x)
print(test_acc_L1, "is the testing accuracy for our logistic regression with L1 regularization")
# +
#training accuracy
c = 1
prob = []
logreg_model(c, train_x, train_y_1_arr, train_x, train_y)
logreg_model(c, train_x, train_y_2_arr, train_x, train_y)
logreg_model(c, train_x, train_y_3_arr, train_x, train_y)
logreg_model(c, train_x, train_y_4_arr, train_x, train_y)
logreg_model(c, train_x, train_y_5_arr, train_x, train_y)
train_yhat_L1 = []
for i in range (0, len(train_x), 1):
values = []
probability_and_classification = []
m1 = prob[0][i][1]
m2 = prob[1][i][1]
m3 = prob[2][i][1]
m4 = prob[3][i][1]
m5 = prob[4][i][1]
values.append(m1)
values.append(m2)
values.append(m3)
values.append(m4)
values.append(m5)
train_yhat_L1.append((values.index(max(values))) + 1)
print(train_yhat_L1) # Correct
correct = 0
incorrect = 0
for i in range (0, len(train_x)):
if (train_yhat_L1[i] == train_y[i]):
correct+= 1
# print(train_yhat_L1[i], ", actual: ", train_y[i][0], "=", (train_y[i][0]))
else:
incorrect += 1
# print("INCORRECT", train_yhat_L1[i], ", actual: ", train_y[i][0], "=", (train_y[i][0]))
train_acc_L1 = correct/len(train_x)
print(train_acc_L1, "is the training accuracy for our logistic regression with L1 regularization")
# -
# # Regularization L2
def logreg2_model(c , X_train, Y_train, X_test, Y_test):
logreg2 = linear_model.LogisticRegression(C=c, warm_start=True)
logreg2.fit(X_train, Y_train)
Yhat_train = logreg2.predict(X_train)
Yhat_test = logreg2.predict(X_test)
prob.append(logreg2.predict_proba(X_test))
# +
#Testing Accuracy
prob = []
c = 1
logreg2_model(c, train_x, train_y_1_arr, test_x, test_y_1_arr)
logreg2_model(c, train_x, train_y_2_arr, test_x, test_y_2_arr)
logreg2_model(c, train_x, train_y_3_arr, test_x, test_y_3_arr)
logreg2_model(c, train_x, train_y_4_arr, test_x, test_y_4_arr)
logreg2_model(c, train_x, train_y_5_arr, test_x, test_y_5_arr)
test_yhat_L2 = []
# test_yhat_L2_pac = []
for i in range (0, len(test_x), 1):
values = []
probability_and_classification = []
m1 = prob[0][i][1]
m2 = prob[1][i][1]
m3 = prob[2][i][1]
m4 = prob[3][i][1]
m5 = prob[4][i][1]
values.append(m1)
values.append(m2)
values.append(m3)
values.append(m4)
values.append(m5)
#max_prob = max(m1, m2, m3, m4, m5)
# probability_and_classification.append(max(values))
# probability_and_classification.append((values.index(max(values))) + 1)
# test_yhat_L2_pac.append(probability_and_classification)
test_yhat_L2.append((values.index(max(values))) + 1)
print(test_yhat_L2) # Correct
#Calculate testing accuracy
correct = 0
incorrect = 0
for i in range (0, len(test_x)):
if (test_yhat_L2[i] == test_y[i]):
correct+= 1
# print(test_yhat_L2[i], ", actual: ", test_y[i][0], "=", (test_y[i][0]))
else:
incorrect += 1
# print("INCORRECT", test_yhat_L2[i], ", actual: ", test_y[i][0], "=", (test_y[i][0]))
test_acc_L2 = correct/len(test_x)
print(test_acc_L2, "is the testing accuracy for our logistic regression with L2 regularization")
# +
#training accuracy
c = 1
prob = []
logreg2_model(c, train_x, train_y_1_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_2_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_3_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_4_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_5_arr, train_x, train_y)
train_yhat_L2 = []
for i in range (0, len(train_x), 1):
values = []
probability_and_classification = []
m1 = prob[0][i][1]
m2 = prob[1][i][1]
m3 = prob[2][i][1]
m4 = prob[3][i][1]
m5 = prob[4][i][1]
values.append(m1)
values.append(m2)
values.append(m3)
values.append(m4)
values.append(m5)
train_yhat_L2.append((values.index(max(values))) + 1)
print(train_yhat_L2) # Correct
correct = 0
incorrect = 0
for i in range (0, len(train_x)):
if (train_yhat_L2[i] == train_y[i]):
correct+= 1
# print(train_yhat_L2[i], ", actual: ", train_y[i][0], "=", (train_y[i][0]))
else:
incorrect += 1
# print("INCORRECT", train_yhat_L2[i], ", actual: ", train_y[i][0], "=", (train_y[i][0]))
train_acc_L2 = correct/len(train_x)
print(train_acc_L2, "is the training accuracy for our logistic regression with L2 regularization")
# -
# # Variation in C
#change logreg2_model to best regulariazation above
c_vals = [0.0001, 0.001, 0.01, 0.1, 1, 10]
train_acc_carray = []
test_acc_carray = []
for c in c_vals:
#testing
prob = []
logreg2_model(c, train_x, train_y_1_arr, test_x, test_y_1_arr)
logreg2_model(c, train_x, train_y_2_arr, test_x, test_y_2_arr)
logreg2_model(c, train_x, train_y_3_arr, test_x, test_y_3_arr)
logreg2_model(c, train_x, train_y_4_arr, test_x, test_y_4_arr)
logreg2_model(c, train_x, train_y_5_arr, test_x, test_y_5_arr)
test_yhat_linear = []
# test_yhat_linear_pac = []
for i in range (0, len(test_x), 1):
values = []
probability_and_classification = []
m1 = prob[0][i][1]
m2 = prob[1][i][1]
m3 = prob[2][i][1]
m4 = prob[3][i][1]
m5 = prob[4][i][1]
values.append(m1)
values.append(m2)
values.append(m3)
values.append(m4)
values.append(m5)
#max_prob = max(m1, m2, m3, m4, m5)
# probability_and_classification.append(max(values))
# probability_and_classification.append((values.index(max(values))) + 1)
# test_yhat_linear_pac.append(probability_and_classification)
test_yhat_linear.append((values.index(max(values))) + 1)
# print(test_yhat_linear) # Correct
#Calculate testing accuracy
correct = 0
incorrect = 0
for i in range (0, 664):
if (test_yhat_linear[i] == test_y[i]):
correct+= 1
# print(test_yhat_linear[i], ", actual: ", test_y[i][0], "=", (test_y[i][0]))
else:
incorrect += 1
# print("INCORRECT", test_yhat_linear[i], ", actual: ", test_y[i][0], "=", (test_y[i][0]))
test_acc_linear = correct/len(test_x)
print(test_acc_linear, "is the testing accuracy for logistic regression with regularization L2 and c =" , c)
test_acc_carray.append(test_acc_linear)
#training accuracy
prob = []
logreg2_model(c, train_x, train_y_1_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_2_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_3_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_4_arr, train_x, train_y)
logreg2_model(c, train_x, train_y_5_arr, train_x, train_y)
train_yhat_linear = []
for i in range (0, len(train_x), 1):
values = []
probability_and_classification = []
m1 = prob[0][i][1]
m2 = prob[1][i][1]
m3 = prob[2][i][1]
m4 = prob[3][i][1]
m5 = prob[4][i][1]
values.append(m1)
values.append(m2)
values.append(m3)
values.append(m4)
values.append(m5)
train_yhat_linear.append((values.index(max(values))) + 1)
# print(train_yhat_linear) # Correct
correct = 0
incorrect = 0
for i in range (0, len(train_x)):
if (train_yhat_linear[i] == train_y[i]):
correct+= 1
# print(train_yhat_linear[i], ", actual: ", train_y[i][0], "=", (train_y[i][0]))
else:
incorrect += 1
# print("INCORRECT", train_yhat_linear[i], ", actual: ", train_y[i][0], "=", (train_y[i][0]))
train_acc_linear = correct/len(train_x)
train_acc_carray.append(train_acc_linear)
print(train_acc_linear, "is the training accuracy for logistic regression with regularization L2 and c =" , c , "\n")
# +
plt.plot(c_vals, train_acc_carray, 'ro-')
plt.plot(c_vals, test_acc_carray,'bo-')
plt.grid()
plt.xlabel('C values (0.0001, 0.001, 0.01, 0.1, 1, 10)')
plt.ylabel('Accuracies')
plt.title('Logistic Regression with L2 Regularization and varying c values')
# Use the following function to have a legend
plt.legend(['Training Accuracy', 'Test Accuracy'], loc='best')
# -
| ArticleClassifier_Logistic_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Visualizing streamlines
# https://www.numbercrunch.de/blog/2013/05/visualizing-streamlines/#codesyntax_1
#
# In Visualizing vector fields I showed how to plot vector fields using Python and Matplotlib. Streamlines are a concept that is closely related to vector fields. Mathematically speaking streamlines are continuous lines whose tangent at each point is given by a vector field. Each line, and therefore also streamlines, can be parametrized by some parameter $t$. A streamline $\vec{r}(t)$ fulfils the equation\begin{equation}
# \frac{\mathrm{d}\vec{r}(t)}{\mathrm{d}t} = g(t)\vec{E}(\vec{r}(t))\,,
# \end{equation}
# where $\vec{E}(\vec{r}(t))$ is the vector field and $g(t)$ some scaling function. The scaling functions is arbitrary but must not be zero. It basically determines how fast one moves along the streamline as a function of the parameter $t$. It is often convenient to set\begin{equation}
# g(t)=\frac{1}{|\vec{E}(\vec{r}(t))|}\,.
# \end{equation}
#
# Since version 1.2.0 the Python package Matplotlib comes with a streamplot function for quick and easy visualizing two-dimensional streamlines. Coming back the example of an electric dipole from Visualizing vector fields the following Python code plots the streamlines of an electric dipole. Compared to the previous post on the plotting vector fields this code is somewhat more generic. First some charges are specified and afterwards the total electric field is calculated by summing over the electric field of the individual charges.
# +
# #!/usr/bin/env python
# import useful modules
import matplotlib
from numpy import *
from pylab import *
from scipy.integrate import ode
# use LaTeX, choose nice some looking fonts and tweak some settings
matplotlib.rc('font', family='serif')
matplotlib.rc('font', size=16)
matplotlib.rc('legend', fontsize=16)
matplotlib.rc('legend', numpoints=1)
matplotlib.rc('legend', handlelength=1.5)
matplotlib.rc('legend', frameon=False)
matplotlib.rc('xtick.major', pad=7)
matplotlib.rc('xtick.minor', pad=7)
matplotlib.rc('text', usetex=True)
matplotlib.rc('text.latex',
preamble=[r'\usepackage[T1]{fontenc}',
r'\usepackage{amsmath}',
r'\usepackage{txfonts}',
r'\usepackage{textcomp}'])
class charge:
def __init__(self, q, pos):
self.q=q
self.pos=pos
def E_point_charge(q, a, x, y):
return q*(x-a[0])/((x-a[0])**2+(y-a[1])**2)**(1.5), \
q*(y-a[1])/((x-a[0])**2+(y-a[1])**2)**(1.5)
def E_total(x, y, charges):
Ex, Ey=0, 0
for C in charges:
E=E_point_charge(C.q, C.pos, x, y)
Ex=Ex+E[0]
Ey=Ey+E[1]
return [ Ex, Ey ]
close('all')
figure(figsize=(6, 4.5))
# charges and positions
charges=[ charge(1, [-1, 0]), charge(-1, [1, 0]) ]
# plot field lines
x0, x1=-2, 2
y0, y1=-1.5, 1.5
x=linspace(x0, x1, 64)
y=linspace(y0, y1, 64)
x, y=meshgrid(x, y)
Ex, Ey=E_total(x, y, charges)
streamplot(x, y, Ex, Ey, color='k')
# plot point charges
for C in charges:
if C.q>0:
plot(C.pos[0], C.pos[1], 'bo', ms=8*sqrt(C.q))
if C.q<0:
plot(C.pos[0], C.pos[1], 'ro', ms=8*sqrt(-C.q))
xlabel('$x$')
ylabel('$y$')
gca().set_xlim(x0, x1)
gca().set_ylim(y0, y1)
show()
axis('image')
# -
# Streamlines of an electric dipole visualized using Matplotlib’s streamplot function.
#
# # 1st
# Matplotlib’s streamplot function is very generic and easy to use. However it does not know anything about specific characteristics of the vector field to plot. For example, it is not able to take into account that streamlines of electric fields always start and end at the charges. Therefore, the following code plots streamlines by solving the streamlines’ ordinary differential equations. We always start close in the vicinity of the electric charges and extend each streamline until it has reached another charge or has left the plotting area.
# +
# #!/usr/bin/env python
# import usefull modules
import matplotlib
from numpy import *
from pylab import *
from scipy.integrate import ode
# use LaTeX, choose nice some looking fonts and tweak some settings
matplotlib.rc('font', family='serif')
matplotlib.rc('font', size=16)
matplotlib.rc('legend', fontsize=16)
matplotlib.rc('legend', numpoints=1)
matplotlib.rc('legend', handlelength=1.5)
matplotlib.rc('legend', frameon=False)
matplotlib.rc('xtick.major', pad=7)
matplotlib.rc('xtick.minor', pad=7)
matplotlib.rc('text', usetex=True)
matplotlib.rc('text.latex',
preamble=[r'\usepackage[T1]{fontenc}',
r'\usepackage{amsmath}',
r'\usepackage{txfonts}',
r'\usepackage{textcomp}'])
class charge:
def __init__(self, q, pos):
self.q=q
self.pos=pos
def E_point_charge(q, a, x, y):
return q*(x-a[0])/((x-a[0])**2+(y-a[1])**2)**(1.5), \
q*(y-a[1])/((x-a[0])**2+(y-a[1])**2)**(1.5)
def E_total(x, y, charges):
Ex, Ey=0, 0
for C in charges:
E=E_point_charge(C.q, C.pos, x, y)
Ex=Ex+E[0]
Ey=Ey+E[1]
return [ Ex, Ey ]
def E_dir(t, y, charges):
Ex, Ey=E_total(y[0], y[1], charges)
n=sqrt(Ex**2+Ey*Ey)
return [Ex/n, Ey/n]
close('all')
figure(figsize=(6, 4.5))
# charges and positions
charges=[ charge(1, [-1, 0]), charge(-1, [1, 0]) ]
# plot field lines
x0, x1=-2, 2
y0, y1=-1.5, 1.5
R=0.01
# loop over all charges
for C in charges:
# plot field lines starting in current charge
dt=0.8*R
if C.q<0:
dt=-dt
# loop over field lines starting in different directions
# around current charge
for alpha in linspace(0, 2*pi*15/16, 16):
r=ode(E_dir)
r.set_integrator('vode')
r.set_f_params(charges)
x=[ C.pos[0] + cos(alpha)*R ]
y=[ C.pos[1] + sin(alpha)*R ]
r.set_initial_value([x[0], y[0]], 0)
while r.successful():
r.integrate(r.t+dt)
x.append(r.y[0])
y.append(r.y[1])
hit_charge=False
# check if field line left drwaing area or ends in some charge
for C2 in charges:
if sqrt((r.y[0]-C2.pos[0])**2+(r.y[1]-C2.pos[1])**2)<R:
hit_charge=True
if hit_charge or (not (x0<r.y[0] and r.y[0]<x1)) or \
(not (y0<r.y[1] and r.y[1]<y1)):
break
plot(x, y, '-k')
# plot point charges
for C in charges:
if C.q>0:
plot(C.pos[0], C.pos[1], 'bo', ms=8*sqrt(C.q))
if C.q<0:
plot(C.pos[0], C.pos[1], 'ro', ms=8*sqrt(-C.q))
xlabel('$x$')
ylabel('$y$')
gca().set_xlim(x0, x1)
gca().set_ylim(y0, y1)
show()
# -
# Streamlines of an electric dipole visualized by solving the streamlines’ differential equations.
| python/matplotlib/vector-streamlines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/anujpatel96/Algorithms/blob/main/MergeSort.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="CMtfzqY1RJnH" colab={"base_uri": "https://localhost:8080/"} outputId="f1d4bd18-6673-4d2b-f7c5-699de0c3870e"
def merge_sort(arr):
if len(arr) <= 1:
return
mid = len(arr)//2
left = arr[:mid]
right = arr[mid:]
merge_sort(left)
merge_sort(right)
merge_two_sorted_lists(left, right, arr)
def merge_two_sorted_lists(a,b,arr):
len_a = len(a)
len_b = len(b)
i = j = k = 0
while i < len_a and j < len_b:
if a[i] <= b[j]:
arr[k] = a[i]
i+=1
else:
arr[k] = b[j]
j+=1
k+=1
while i < len_a:
arr[k] = a[i]
i+=1
k+=1
while j < len_b:
arr[k] = b[j]
j+=1
k+=1
if __name__ == '__main__':
elements = [2,4,1,3,8,6]
print("List =======>",elements)
merge_sort(elements)
print("SortedList =>",elements)
# + colab={"base_uri": "https://localhost:8080/"} id="FFr_CrHmSRr-" outputId="e32deaeb-529b-43ad-c483-fde897157a3b"
def merge_two_sorted_lists(a,b):
sorted_list = []
len_a = len(a)
len_b = len(b)
i = j = 0
while i < len_a and j < len_b:
if a[i] <= b[j]:
sorted_list.append(a[i])
i+=1
else:
sorted_list.append(b[j])
j+=1
while i < len_a:
sorted_list.append(a[i])
i+=1
while j < len_b:
sorted_list.append(b[j])
j+=1
return sorted_list
if __name__ == '__main__':
list1 = [1,2,5,7,8]
list2 = [3,4,6,9,10]
print(f"List1 ==============> {list1}")
print(f"List2 ==============> {list2}")
result = merge_two_sorted_lists(list1,list2)
print("Sorted Megerd List =>",result)
| MergeSort.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from bs4 import BeautifulSoup
import urllib.request
import pandas as pandas
url='http://pythonprogramming.net/parsememcparseface/'
html= urllib.request.urlopen(url).read()
soup=BeautifulSoup (html, 'lxml')
table=soup.find('table') # find_all을 사용하여 찾는다면,
table_rows= table.find_all('tr') # table[0].find_all('tr') 이라고 몇번째 'tr'을 가지고 오라고 지정해 줘야 한다.
#참고> select로 경로를 찾은 경우에도 []을 사용하여 몇번째 'tr'을 가지고 오라고 지정해 줘야 한다.
for tr in table_rows:
td= tr.find_all('td')
rows= [i.text for i in td] # td가 하나 하나의 값이기 때문에 해당 열(tr)을 리스트 형태로 만들기 위해 []을 해줘야 한다.
print(rows) # 모든 열(tr)에서 td를 추출해 내기 위해서는 최상위 for문인 'for tr in table_rows:' 안에서 코드가 작성되어야 함
# 최상위 for문 안에서 새로운 변수를 선언하고 td 값을 불러내야 한다.
type(rows)
# +
# 잘못된 예
from bs4 import BeautifulSoup
import urllib.request
import pandas as pandas
url='http://pythonprogramming.net/parsememcparseface/'
html= urllib.request.urlopen(url).read()
soup=BeautifulSoup (html, 'lxml')
table=soup.find('table') # find_all을 사용하여 찾는다면,
table_rows= table.find_all('tr') # table[0].find_all('tr') 이라고 몇번째 'tr'을 가지고 오라고 지정해 줘야 한다.
for tr in table_rows:
td= tr.find_all('td')
for i in td: # 이 for은 위 for문에서 마지막 열(tr)이 실행된 후 모든 td를 텍스트 하라는 의미임
i.text # 모든 열(tr)에서 td를 추출해 내기 위해서는 최상위 for문인 'for tr in table_rows:' 안에서 코드가 작성되어야 함
print(i) # 따라서 위 셀에서처럼 최상위 for문 안에서 새로운 변수를 선언하고 td 값을 불러내야 한다.
type(row)
# +
# 잘못된 예
from bs4 import BeautifulSoup
import urllib.request
import pandas as pandas
url='http://pythonprogramming.net/parsememcparseface/'
html= urllib.request.urlopen(url).read()
soup=BeautifulSoup (html, 'lxml')
table=soup.find('table') # find_all을 사용하여 찾는다면,
table_rows= table.find_all('tr') # table[0].find_all('tr') 이라고 몇번째 'tr'을 가지고 오라고 지정해 줘야 한다.
for tr in table_rows:
td= tr.find_all('td').get_text() # text로 불러오는 것은 find, find_all과 같이 범위가 지정되지 않은 함수에서는 사용할 수 없다.
# 마지막 결과로 나온 값(td)에만 사용할 수 있음
print(td)
# +
# 데이터 프레임으로 표를 작성할 경우
from bs4 import BeautifulSoup
import urllib.request
import pandas as pd
url = 'https://www.alexa.com/topsites'
search_url = urllib.request.urlopen(url).read()
soup = BeautifulSoup(search_url, 'html.parser')
rank_df = pd.DataFrame (columns=('rank', 'Site', 'Daily Time on Site', 'Daily Pageviews per Visitor', '% of Traffic From Search', 'Total Sites Linking In'))
# 각 열의 값들을 담아줄 columns 항목을 미리 만들어 준다.
sites = soup.find_all('div', {'class': 'tr site-listing'}) # for문이 돌아갈 열(대부분 table에서는 tr 형태임)
for site in sites: # 모든 열(대부분 tr 형태)의 각 값(대부분 td 형태)을 미리 만들어 둔 columns 항목에 넣어주는 for문임
rank = site.find('div',{'class':'td'}).get_text()
website = site.select('p > a')[0].get_text()
infos=site.find_all('div',{'class':'td right'})
time=infos[0].get_text()
pageview=infos[1].get_text()
traffic=infos[2].get_text()
linking=infos[3].get_text()
rank_df.loc[rank] = [rank, website, time, pageview, traffic, linking] # for문에서 선언한 변수(찾아낸 값,tr)를 미리 만들어 둔 columns 항목에 맞게 리스트로 넣어줌
rank_df
# +
# pandas.read_html
from bs4 import BeautifulSoup
import urllib.request
import pandas as pd
dfs= pd.read_html('http://pythonprogramming.net/parsememcparseface/', header=0) # header=0은 원하는 컬럼 위에 또다른 컬럼을 지워고 테이블을 만들고 싶은 경우 사용
for df in dfs: # for문을 사용하지 않고 dfs만 출력하면 아래 셀의 결과처럼 리스트로 출력됨
print(df)
type(df)
# +
# 위 셀에서 for문을 사용하지 않고 dfs를 출력한 경우 - list
from bs4 import BeautifulSoup
import urllib.request
import pandas as pd
dfs= pd.read_html('http://pythonprogramming.net/parsememcparseface/')
print(dfs)
type(dfs)
# -
| Import Examples/import_table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrdbourke/tensorflow-deep-learning/blob/main/08_introduction_to_nlp_in_tensorflow.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vtAgo5zYCClj"
# # 08. Natural Language Processing with TensorFlow
#
# 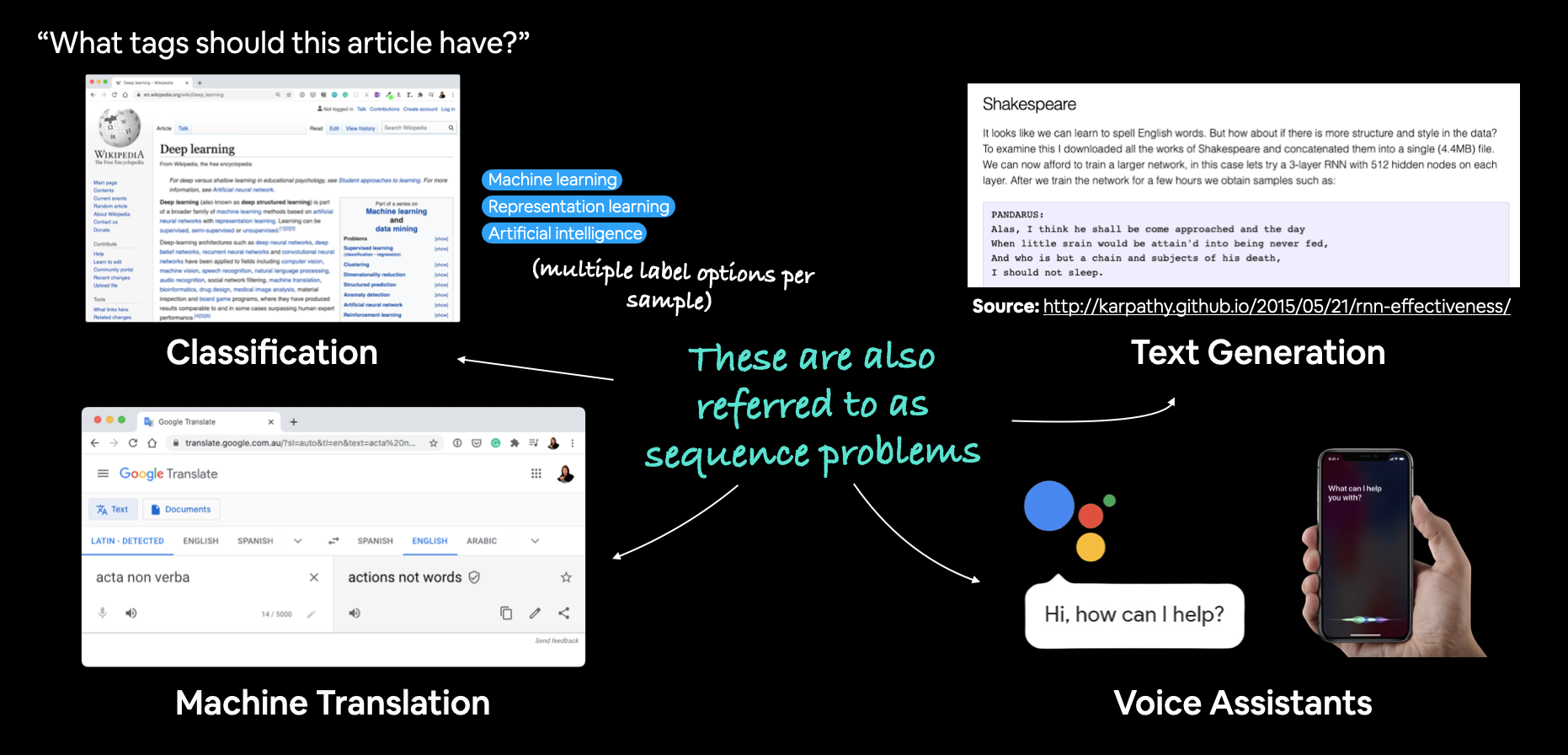
# *A handful of example natural language processing (NLP) and natural language understanding (NLU) problems. These are also often referred to as sequence problems (going from one sequence to another).*
#
# The main goal of [natural language processing (NLP)](https://becominghuman.ai/a-simple-introduction-to-natural-language-processing-ea66a1747b32) is to derive information from natural language.
#
# Natural language is a broad term but you can consider it to cover any of the following:
# * Text (such as that contained in an email, blog post, book, Tweet)
# * Speech (a conversation you have with a doctor, voice commands you give to a smart speaker)
#
# Under the umbrellas of text and speech there are many different things you might want to do.
#
# If you're building an email application, you might want to scan incoming emails to see if they're spam or not spam (classification).
#
# If you're trying to analyse customer feedback complaints, you might want to discover which section of your business they're for.
#
# > 🔑 **Note:** Both of these types of data are often referred to as *sequences* (a sentence is a sequence of words). So a common term you'll come across in NLP problems is called *seq2seq*, in other words, finding information in one sequence to produce another sequence (e.g. converting a speech command to a sequence of text-based steps).
#
# To get hands-on with NLP in TensorFlow, we're going to practice the steps we've used previously but this time with text data:
#
# ```
# Text -> turn into numbers -> build a model -> train the model to find patterns -> use patterns (make predictions)
# ```
#
# > 📖 **Resource:** For a great overview of NLP and the different problems within it, read the article [*A Simple Introduction to Natural Language Processing*](https://becominghuman.ai/a-simple-introduction-to-natural-language-processing-ea66a1747b32).
#
# ## What we're going to cover
#
# Let's get specific hey?
#
# * Downloading a text dataset
# * Visualizing text data
# * Converting text into numbers using tokenization
# * Turning our tokenized text into an embedding
# * Modelling a text dataset
# * Starting with a baseline (TF-IDF)
# * Building several deep learning text models
# * Dense, LSTM, GRU, Conv1D, Transfer learning
# * Comparing the performance of each our models
# * Combining our models into an ensemble
# * Saving and loading a trained model
# * Find the most wrong predictions
#
# ## How you should approach this notebook
#
# You can read through the descriptions and the code (it should all run, except for the cells which error on purpose), but there's a better option.
#
# Write all of the code yourself.
#
# Yes. I'm serious. Create a new notebook, and rewrite each line by yourself. Investigate it, see if you can break it, why does it break?
#
# You don't have to write the text descriptions but writing the code yourself is a great way to get hands-on experience.
#
# Don't worry if you make mistakes, we all do. The way to get better and make less mistakes is to write more code.
#
# > 📖 **Resource:** See the full set of course materials on GitHub: https://github.com/mrdbourke/tensorflow-deep-learning
# + [markdown] id="4Zh2N1hZtvpN"
# ## Check for GPU
#
# In order for our deep learning models to run as fast as possible, we'll need access to a GPU.
#
# In Google Colab, you can set this up by going to Runtime -> Change runtime type -> Hardware accelerator -> GPU.
#
# After selecting GPU, you may have to restart the runtime.
# + id="DEYTFigmc3CI" colab={"base_uri": "https://localhost:8080/"} outputId="57c44ca3-b01f-412c-a670-73b70e861937"
# Check for GPU
# !nvidia-smi -L
# + [markdown] id="gS3YnNNI8oFk"
# ## Get helper functions
#
# In past modules, we've created a bunch of helper functions to do small tasks required for our notebooks.
#
# Rather than rewrite all of these, we can import a script and load them in from there.
#
# The script containing our helper functions can be [found on GitHub](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/extras/helper_functions.py).
# + colab={"base_uri": "https://localhost:8080/"} id="aFOHPqgE8pv-" outputId="523e55ff-21ee-41ed-e3a4-7e19274f9ea8"
# Download helper functions script
# !wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/helper_functions.py
# + id="ICFbSkoM85tq"
# Import series of helper functions for the notebook
from helper_functions import unzip_data, create_tensorboard_callback, plot_loss_curves, compare_historys
# + [markdown] id="cCZrclc2COWW"
# ## Download a text dataset
#
# Let's start by download a text dataset. We'll be using the [Real or Not?](https://www.kaggle.com/c/nlp-getting-started/data) datset from Kaggle which contains text-based Tweets about natural disasters.
#
# The Real Tweets are actually about diasters, for example:
#
# ```
# Jetstar and Virgin forced to cancel Bali flights again because of ash from Mount Raung volcano
# ```
#
# The Not Real Tweets are Tweets not about diasters (they can be on anything), for example:
#
# ```
# 'Education is the most powerful weapon which you can use to change the world.' Nelson #Mandela #quote
# ```
#
# For convenience, the dataset has been [downloaded from Kaggle](https://www.kaggle.com/c/nlp-getting-started/data) (doing this requires a Kaggle account) and uploaded as a downloadable zip file.
#
# > 🔑 **Note:** The original downloaded data has not been altered to how you would download it from Kaggle.
# + id="C0FEcci5IH8S" colab={"base_uri": "https://localhost:8080/"} outputId="75a40c19-6eae-4323-d84e-c6af405fc0ba"
# Download data (same as from Kaggle)
# !wget "https://storage.googleapis.com/ztm_tf_course/nlp_getting_started.zip"
# Unzip data
unzip_data("nlp_getting_started.zip")
# + [markdown] id="wBIR6tTI9QcR"
# Unzipping `nlp_getting_started.zip` gives the following 3 `.csv` files:
# * `sample_submission.csv` - an example of the file you'd submit to the Kaggle competition of your model's predictions.
# * `train.csv` - training samples of real and not real diaster Tweets.
# * `test.csv` - testing samples of real and not real diaster Tweets.
# + [markdown] id="7HpxZKYdD6V-"
# ## Visualizing a text dataset
#
# Once you've acquired a new dataset to work with, what should you do first?
#
# Explore it? Inspect it? Verify it? Become one with it?
#
# All correct.
#
# Remember the motto: visualize, visualize, visualize.
#
# Right now, our text data samples are in the form of `.csv` files. For an easy way to make them visual, let's turn them into pandas DataFrame's.
#
# > 📖 **Reading:** You might come across text datasets in many different formats. Aside from CSV files (what we're working with), you'll probably encounter `.txt` files and `.json` files too. For working with these type of files, I'd recommend reading the two following articles by RealPython:
# * [How to Read and Write Files in Python](https://realpython.com/read-write-files-python/)
# * [Working with JSON Data in Python](https://realpython.com/python-json/)
# + id="qRvkeYEJIKsw" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="052a1d3e-1d52-47e9-ea39-df6dd4d149ca"
# Turn .csv files into pandas DataFrame's
import pandas as pd
train_df = pd.read_csv("train.csv")
test_df = pd.read_csv("test.csv")
train_df.head()
# + [markdown] id="1xGqlnQaLmaT"
# The training data we downloaded is probably shuffled already. But just to be sure, let's shuffle it again.
# + id="ACCE7h6OMVjR" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="51f2ae2e-df2a-4e15-a618-7087ecba1914"
# Shuffle training dataframe
train_df_shuffled = train_df.sample(frac=1, random_state=42) # shuffle with random_state=42 for reproducibility
train_df_shuffled.head()
# + [markdown] id="Lw4mKW1yL0kI"
# Notice how the training data has a `"target"` column.
#
# We're going to be writing code to find patterns (e.g. different combinations of words) in the `"text"` column of the training dataset to predict the value of the `"target"` column.
#
# The test dataset doesn't have a `"target"` column.
#
# ```
# Inputs (text column) -> Machine Learning Algorithm -> Outputs (target column)
# ```
#
# 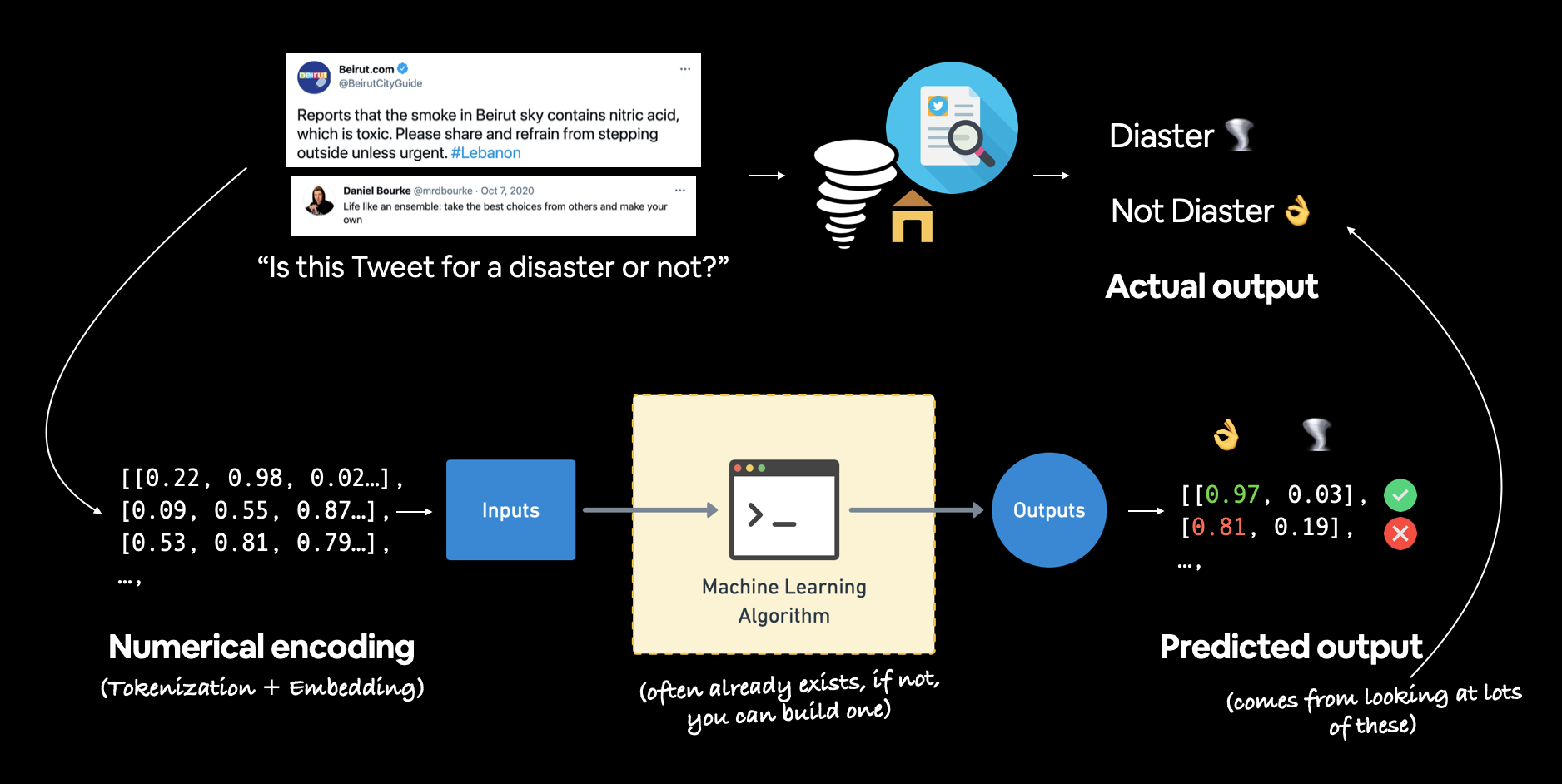
# *Example text classification inputs and outputs for the problem of classifying whether a Tweet is about a diaster or not.*
# + id="tDh5t7thI5BM" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="9f4321e2-a45e-4eef-93d1-71972c992e37"
# The test data doesn't have a target (that's what we'd try to predict)
test_df.head()
# + [markdown] id="O4JhBRn5Mn-V"
# Let's check how many examples of each target we have.
# + id="k4P5DnLhIciD" colab={"base_uri": "https://localhost:8080/"} outputId="50fa6a7a-a7cf-4dc5-b5a9-01ccb8dfbe61"
# How many examples of each class?
train_df.target.value_counts()
# + [markdown] id="WjEDQ297Ihy4"
# Since we have two target values, we're dealing with a **binary classification** problem.
#
# It's fairly balanced too, about 60% negative class (`target = 0`) and 40% positive class (`target = 1`).
#
# Where,
#
# * `1` = a real disaster Tweet
# * `0` = not a real disaster Tweet
#
# And what about the total number of samples we have?
# + id="jQxg7EKKIy5L" colab={"base_uri": "https://localhost:8080/"} outputId="ca4aefe4-3405-4a94-8e8a-163386b25709"
# How many samples total?
print(f"Total training samples: {len(train_df)}")
print(f"Total test samples: {len(test_df)}")
print(f"Total samples: {len(train_df) + len(test_df)}")
# + [markdown] id="Q1upY8-xNPWV"
# Alright, seems like we've got a decent amount of training and test data. If anything, we've got an abundance of testing examples, usually a split of 90/10 (90% training, 10% testing) or 80/20 is suffice.
#
# Okay, time to visualize, let's write some code to visualize random text samples.
#
# > 🤔 **Question:** Why visualize random samples? You could visualize samples in order but this could lead to only seeing a certain subset of data. Better to visualize a substantial quantity (100+) of random samples to get an idea of the different kinds of data you're working with. In machine learning, never underestimate the power of randomness.
# + id="vH3EXknTI3bQ" colab={"base_uri": "https://localhost:8080/"} outputId="4e068f05-06fd-4611-b0c9-519f4e67d8b9"
# Let's visualize some random training examples
import random
random_index = random.randint(0, len(train_df)-5) # create random indexes not higher than the total number of samples
for row in train_df_shuffled[["text", "target"]][random_index:random_index+5].itertuples():
_, text, target = row
print(f"Target: {target}", "(real disaster)" if target > 0 else "(not real disaster)")
print(f"Text:\n{text}\n")
print("---\n")
# + [markdown] id="1FhRRewGPNS_"
# ### Split data into training and validation sets
#
# Since the test set has no labels and we need a way to evalaute our trained models, we'll split off some of the training data and create a validation set.
#
# When our model trains (tries patterns in the Tweet samples), it'll only see data from the training set and we can see how it performs on unseen data using the validation set.
#
# We'll convert our splits from pandas Series datatypes to lists of strings (for the text) and lists of ints (for the labels) for ease of use later.
#
# To split our training dataset and create a validation dataset, we'll use Scikit-Learn's [`train_test_split()`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) method and dedicate 10% of the training samples to the validation set.
# + id="7OJf31TQ-X8s"
from sklearn.model_selection import train_test_split
# Use train_test_split to split training data into training and validation sets
train_sentences, val_sentences, train_labels, val_labels = train_test_split(train_df_shuffled["text"].to_numpy(),
train_df_shuffled["target"].to_numpy(),
test_size=0.1, # dedicate 10% of samples to validation set
random_state=42) # random state for reproducibility
# + colab={"base_uri": "https://localhost:8080/"} id="NWGOTjanBaTQ" outputId="dd1a42bc-d291-42ad-f370-6b4008b3ec45"
# Check the lengths
len(train_sentences), len(train_labels), len(val_sentences), len(val_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="VqhvQK9wBTbw" outputId="31c510a5-5b33-4c3c-b093-de3982bf145b"
# View the first 10 training sentences and their labels
train_sentences[:10], train_labels[:10]
# + [markdown] id="EN-houoSD-hP"
# ## Converting text into numbers
#
# Wonderful! We've got a training set and a validation set containing Tweets and labels.
#
# Our labels are in numerical form (`0` and `1`) but our Tweets are in string form.
#
# > 🤔 **Question:** What do you think we have to do before we can use a machine learning algorithm with our text data?
#
# If you answered something along the lines of "turn it into numbers", you're correct. A machine learning algorithm requires its inputs to be in numerical form.
#
# In NLP, there are two main concepts for turning text into numbers:
# * **Tokenization** - A straight mapping from word or character or sub-word to a numerical value. There are three main levels of tokenization:
# 1. Using **word-level tokenization** with the sentence "I love TensorFlow" might result in "I" being `0`, "love" being `1` and "TensorFlow" being `2`. In this case, every word in a sequence considered a single **token**.
# 2. **Character-level tokenization**, such as converting the letters A-Z to values `1-26`. In this case, every character in a sequence considered a single **token**.
# 3. **Sub-word tokenization** is in between word-level and character-level tokenization. It involves breaking invidual words into smaller parts and then converting those smaller parts into numbers. For example, "my favourite food is pineapple pizza" might become "my, fav, avour, rite, fo, oo, od, is, pin, ine, app, le, piz, za". After doing this, these sub-words would then be mapped to a numerical value. In this case, every word could be considered multiple **tokens**.
# * **Embeddings** - An embedding is a representation of natural language which can be learned. Representation comes in the form of a **feature vector**. For example, the word "dance" could be represented by the 5-dimensional vector `[-0.8547, 0.4559, -0.3332, 0.9877, 0.1112]`. It's important to note here, the size of the feature vector is tuneable. There are two ways to use embeddings:
# 1. **Create your own embedding** - Once your text has been turned into numbers (required for an embedding), you can put them through an embedding layer (such as [`tf.keras.layers.Embedding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding)) and an embedding representation will be learned during model training.
# 2. **Reuse a pre-learned embedding** - Many pre-trained embeddings exist online. These pre-trained embeddings have often been learned on large corpuses of text (such as all of Wikipedia) and thus have a good underlying representation of natural language. You can use a pre-trained embedding to initialize your model and fine-tune it to your own specific task.
#
# 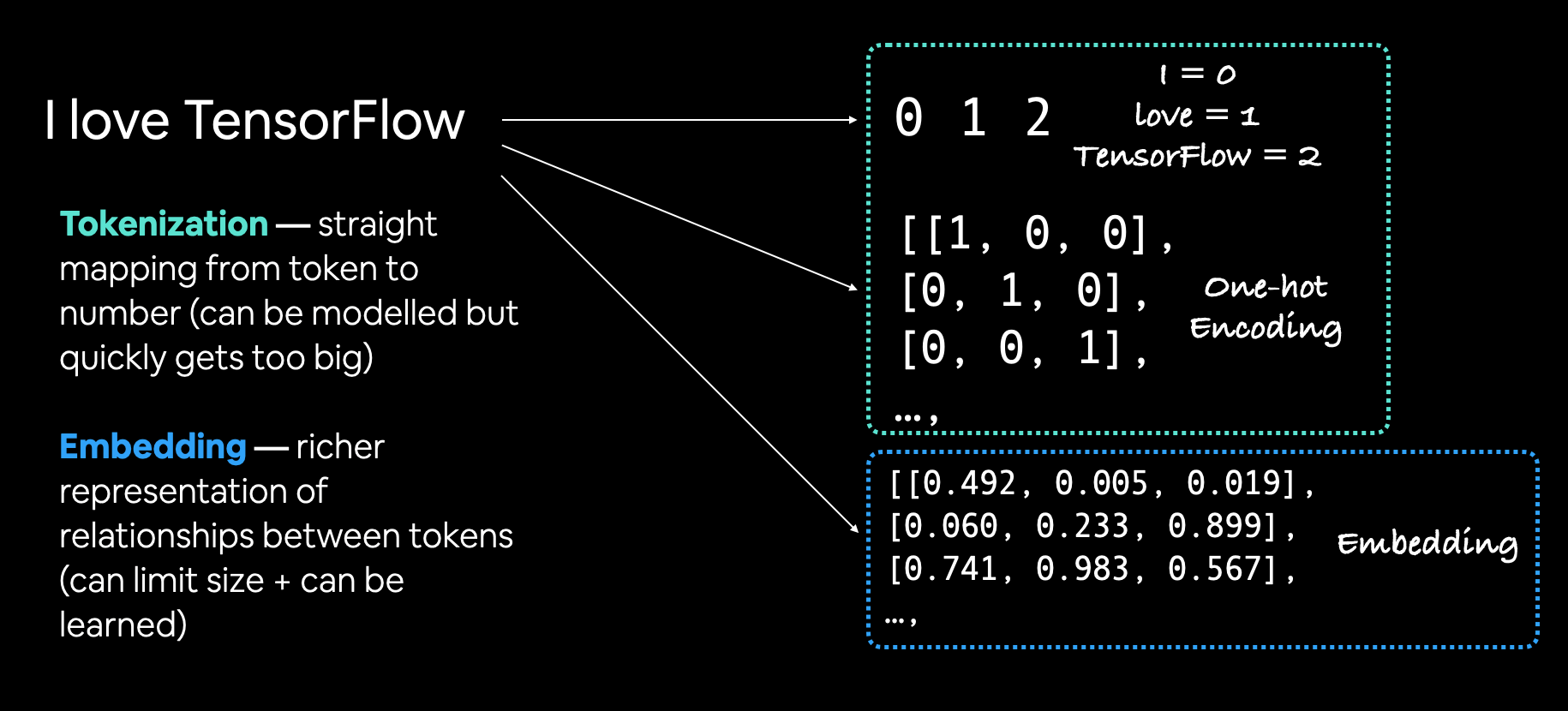
# *Example of **tokenization** (straight mapping from word to number) and **embedding** (richer representation of relationships between tokens).*
#
# > 🤔 **Question:** What level of tokenzation should I use? What embedding should should I choose?
#
# It depends on your problem. You could try character-level tokenization/embeddings and word-level tokenization/embeddings and see which perform best. You might even want to try stacking them (e.g. combining the outputs of your embedding layers using [`tf.keras.layers.concatenate`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/concatenate)).
#
# If you're looking for pre-trained word embeddings, [Word2vec embeddings](http://jalammar.github.io/illustrated-word2vec/), [GloVe embeddings](https://nlp.stanford.edu/projects/glove/) and many of the options available on [TensorFlow Hub](https://tfhub.dev/s?module-type=text-embedding) are great places to start.
#
# > 🔑 **Note:** Much like searching for a pre-trained computer vision model, you can search for pre-trained word embeddings to use for your problem. Try searching for something like "use pre-trained word embeddings in TensorFlow".
# + [markdown] id="8UnRcM1PELHn"
# ### Text vectorization (tokenization)
#
# Enough talking about tokenization and embeddings, let's create some.
#
# We'll practice tokenzation (mapping our words to numbers) first.
#
# To tokenize our words, we'll use the helpful preprocessing layer [`tf.keras.layers.experimental.preprocessing.TextVectorization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization).
#
# The `TextVectorization` layer takes the following parameters:
# * `max_tokens` - The maximum number of words in your vocabulary (e.g. 20000 or the number of unique words in your text), includes a value for OOV (out of vocabulary) tokens.
# * `standardize` - Method for standardizing text. Default is `"lower_and_strip_punctuation"` which lowers text and removes all punctuation marks.
# * `split` - How to split text, default is `"whitespace"` which splits on spaces.
# * `ngrams` - How many words to contain per token split, for example, `ngrams=2` splits tokens into continuous sequences of 2.
# * `output_mode` - How to output tokens, can be `"int"` (integer mapping), `"binary"` (one-hot encoding), `"count"` or `"tf-idf"`. See documentation for more.
# * `output_sequence_length` - Length of tokenized sequence to output. For example, if `output_sequence_length=150`, all tokenized sequences will be 150 tokens long.
# * `pad_to_max_tokens` - Defaults to `False`, if `True`, the output feature axis will be padded to `max_tokens` even if the number of unique tokens in the vocabulary is less than `max_tokens`. Only valid in certain modes, see docs for more.
#
# Let's see it in action.
# + id="PVcZk-LcNunF"
import tensorflow as tf
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
# Note: in TensorFlow 2.6+, you no longer need "layers.experimental.preprocessing"
# you can use: "tf.keras.layers.TextVectorization", see https://github.com/tensorflow/tensorflow/releases/tag/v2.6.0 for more
# Use the default TextVectorization variables
text_vectorizer = TextVectorization(max_tokens=None, # how many words in the vocabulary (all of the different words in your text)
standardize="lower_and_strip_punctuation", # how to process text
split="whitespace", # how to split tokens
ngrams=None, # create groups of n-words?
output_mode="int", # how to map tokens to numbers
output_sequence_length=None) # how long should the output sequence of tokens be?
# pad_to_max_tokens=True) # Not valid if using max_tokens=None
# + [markdown] id="u0Ej5mzKGkK8"
# We've initialized a `TextVectorization` object with the default settings but let's customize it a little bit for our own use case.
#
# In particular, let's set values for `max_tokens` and `output_sequence_length`.
#
# For `max_tokens` (the number of words in the vocabulary), multiples of 10,000 (`10,000`, `20,000`, `30,000`) or the exact number of unique words in your text (e.g. `32,179`) are common values.
#
# For our use case, we'll use `10,000`.
#
# And for the `output_sequence_length` we'll use the average number of tokens per Tweet in the training set. But first, we'll need to find it.
# + id="SQ3ZCINnR56H" colab={"base_uri": "https://localhost:8080/"} outputId="093c7d74-8c3a-4fd1-8624-baa5cc95e183"
# Find average number of tokens (words) in training Tweets
round(sum([len(i.split()) for i in train_sentences])/len(train_sentences))
# + [markdown] id="AFGTRcw8Hv7R"
# Now let's create another `TextVectorization` object using our custom parameters.
# + id="eYPcGwdbafmW"
# Setup text vectorization with custom variables
max_vocab_length = 10000 # max number of words to have in our vocabulary
max_length = 15 # max length our sequences will be (e.g. how many words from a Tweet does our model see?)
text_vectorizer = TextVectorization(max_tokens=max_vocab_length,
output_mode="int",
output_sequence_length=max_length)
# + [markdown] id="BSWycfB3H3wV"
# Beautiful!
#
# To map our `TextVectorization` instance `text_vectorizer` to our data, we can call the `adapt()` method on it whilst passing it our training text.
# + id="0083KHXPO4m2"
# Fit the text vectorizer to the training text
text_vectorizer.adapt(train_sentences)
# + [markdown] id="Syh0VB9wIHUq"
# Training data mapped! Let's try our `text_vectorizer` on a custom sentence (one similar to what you might see in the training data).
# + id="uizmdJKvO2OW" colab={"base_uri": "https://localhost:8080/"} outputId="908d8b3e-0110-47ba-b7d4-ab2e402af79c"
# Create sample sentence and tokenize it
sample_sentence = "There's a flood in my street!"
text_vectorizer([sample_sentence])
# + [markdown] id="M0RmAeplIW57"
# Wonderful, it seems we've got a way to turn our text into numbers (in this case, word-level tokenization). Notice the 0's at the end of the returned tensor, this is because we set `output_sequence_length=15`, meaning no matter the size of the sequence we pass to `text_vectorizer`, it always returns a sequence with a length of 15.
#
# How about we try our `text_vectorizer` on a few random sentences?
# + id="SZFka4BtRR6_" colab={"base_uri": "https://localhost:8080/"} outputId="227a570e-d987-49a1-bcf1-3f561d907d00"
# Choose a random sentence from the training dataset and tokenize it
random_sentence = random.choice(train_sentences)
print(f"Original text:\n{random_sentence}\
\n\nVectorized version:")
text_vectorizer([random_sentence])
# + [markdown] id="PErGKRbPJF89"
# Looking good!
#
# Finally, we can check the unique tokens in our vocabulary using the `get_vocabulary()` method.
# + id="5nwNdgAZIhna" colab={"base_uri": "https://localhost:8080/"} outputId="f23e1a41-5b2d-4042-cfec-c6b22226a798"
# Get the unique words in the vocabulary
words_in_vocab = text_vectorizer.get_vocabulary()
top_5_words = words_in_vocab[:5] # most common tokens (notice the [UNK] token for "unknown" words)
bottom_5_words = words_in_vocab[-5:] # least common tokens
print(f"Number of words in vocab: {len(words_in_vocab)}")
print(f"Top 5 most common words: {top_5_words}")
print(f"Bottom 5 least common words: {bottom_5_words}")
# + [markdown] id="AHyCdO0uEOkH"
# ### Creating an Embedding using an Embedding Layer
#
# We've got a way to map our text to numbers. How about we go a step further and turn those numbers into an embedding?
#
# The powerful thing about an embedding is it can be learned during training. This means rather than just being static (e.g. `1` = I, `2` = love, `3` = TensorFlow), a word's numeric representation can be improved as a model goes through data samples.
#
# We can see what an embedding of a word looks like by using the [`tf.keras.layers.Embedding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) layer.
#
# The main parameters we're concerned about here are:
# * `input_dim` - The size of the vocabulary (e.g. `len(text_vectorizer.get_vocabulary()`).
# * `output_dim` - The size of the output embedding vector, for example, a value of `100` outputs a feature vector of size 100 for each word.
# * `embeddings_initializer` - How to initialize the embeddings matrix, default is `"uniform"` which randomly initalizes embedding matrix with uniform distribution. This can be changed for using pre-learned embeddings.
# * `input_length` - Length of sequences being passed to embedding layer.
#
# Knowing these, let's make an embedding layer.
# + id="OsB4StymSk_s" colab={"base_uri": "https://localhost:8080/"} outputId="cffa28e2-f368-4f82-ff97-aab8b54241d3"
tf.random.set_seed(42)
from tensorflow.keras import layers
embedding = layers.Embedding(input_dim=max_vocab_length, # set input shape
output_dim=128, # set size of embedding vector
embeddings_initializer="uniform", # default, intialize randomly
input_length=max_length, # how long is each input
name="embedding_1")
embedding
# + [markdown] id="bfML_IzlSUho"
# Excellent, notice how `embedding` is a TensoFlow layer? This is important because we can use it as part of a model, meaning its parameters (word representations) can be updated and improved as the model learns.
#
# How about we try it out on a sample sentence?
# + id="1Re6Eew6SZnG" colab={"base_uri": "https://localhost:8080/"} outputId="83f28dec-93fb-483a-a997-a4552ef74426"
# Get a random sentence from training set
random_sentence = random.choice(train_sentences)
print(f"Original text:\n{random_sentence}\
\n\nEmbedded version:")
# Embed the random sentence (turn it into numerical representation)
sample_embed = embedding(text_vectorizer([random_sentence]))
sample_embed
# + [markdown] id="e4Sn8o9pTBE5"
# Each token in the sentence gets turned into a length 128 feature vector.
# + id="g_VBepuSTBDW" colab={"base_uri": "https://localhost:8080/"} outputId="7e62e9cf-bf1c-4e21-a304-ba0b277bb220"
# Check out a single token's embedding
sample_embed[0][0]
# + [markdown] id="Z0NTsDklR0xw"
# These values might not mean much to us but they're what our computer sees each word as. When our model looks for patterns in different samples, these values will be updated as necessary.
#
# > 🔑 **Note:** The previous two concepts (tokenization and embeddings) are the foundation for many NLP tasks. So if you're not sure about anything, be sure to research and conduct your own experiments to further help your understanding.
# + [markdown] id="ZJENUdF3F7Rn"
# ## Modelling a text dataset
#
# 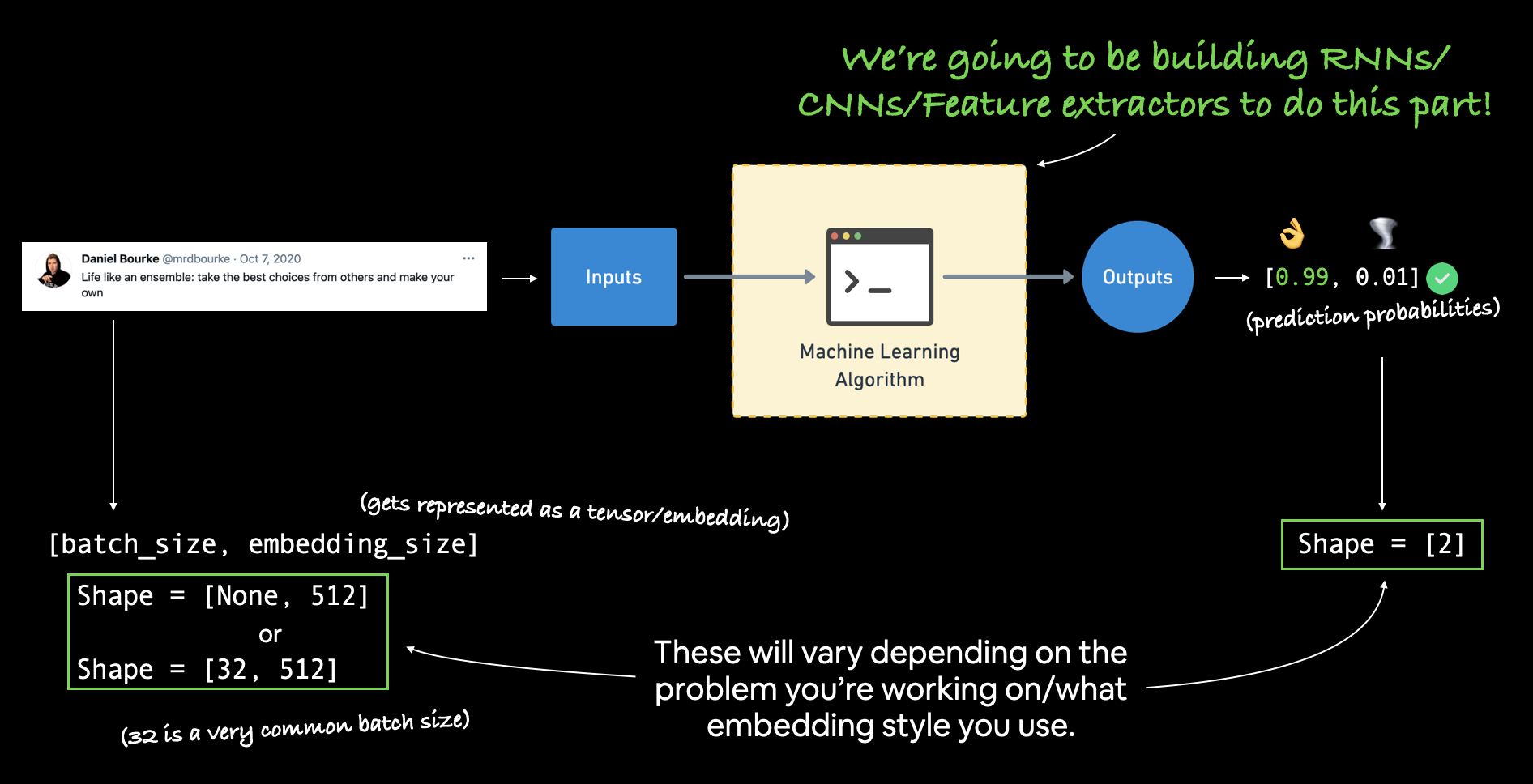
# *Once you've got your inputs and outputs prepared, it's a matter of figuring out which machine learning model to build in between them to bridge the gap.*
#
# Now that we've got a way to turn our text data into numbers, we can start to build machine learning models to model it.
#
# To get plenty of practice, we're going to build a series of different models, each as its own experiment. We'll then compare the results of each model and see which one performed best.
#
# More specifically, we'll be building the following:
# * **Model 0**: Naive Bayes (baseline)
# * **Model 1**: Feed-forward neural network (dense model)
# * **Model 2**: LSTM model
# * **Model 3**: GRU model
# * **Model 4**: Bidirectional-LSTM model
# * **Model 5**: 1D Convolutional Neural Network
# * **Model 6**: TensorFlow Hub Pretrained Feature Extractor
# * **Model 7**: Same as model 6 with 10% of training data
#
# Model 0 is the simplest to acquire a baseline which we'll expect each other of the other deeper models to beat.
#
# Each experiment will go through the following steps:
# * Construct the model
# * Train the model
# * Make predictions with the model
# * Track prediction evaluation metrics for later comparison
#
# Let's get started.
# + [markdown] id="q4i5BiQfF--y"
# ### Model 0: Getting a baseline
#
# As with all machine learning modelling experiments, it's important to create a baseline model so you've got a benchmark for future experiments to build upon.
#
# To create our baseline, we'll create a Scikit-Learn Pipeline using the TF-IDF (term frequency-inverse document frequency) formula to convert our words to numbers and then model them with the [Multinomial Naive Bayes algorithm](https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.MultinomialNB.html#sklearn.naive_bayes.MultinomialNB). This was chosen via referring to the [Scikit-Learn machine learning map](https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html).
#
# > 📖 **Reading:** The ins and outs of TF-IDF algorithm is beyond the scope of this notebook, however, the curious reader is encouraged to check out the [Scikit-Learn documentation for more](https://scikit-learn.org/stable/modules/feature_extraction.html#tfidf-term-weighting).
# + id="xFqjqWcXtOOs" colab={"base_uri": "https://localhost:8080/"} outputId="0cec96b9-d78a-49af-9197-86a03b8043c8"
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
# Create tokenization and modelling pipeline
model_0 = Pipeline([
("tfidf", TfidfVectorizer()), # convert words to numbers using tfidf
("clf", MultinomialNB()) # model the text
])
# Fit the pipeline to the training data
model_0.fit(train_sentences, train_labels)
# + [markdown] id="ybOvOuVJbNjg"
# The benefit of using a shallow model like Multinomial Naive Bayes is that training is very fast.
#
# Let's evaluate our model and find our baseline metric.
# + id="soPfnpmQuUIP" colab={"base_uri": "https://localhost:8080/"} outputId="b947adf2-aaaf-4d9d-afc0-81359138cc5a"
baseline_score = model_0.score(val_sentences, val_labels)
print(f"Our baseline model achieves an accuracy of: {baseline_score*100:.2f}%")
# + [markdown] id="hUv5dyuibf3M"
# How about we make some predictions with our baseline model?
# + id="7n89JxrJufcf" colab={"base_uri": "https://localhost:8080/"} outputId="69a3f67c-1da2-434e-fd29-73d723cd181e"
# Make predictions
baseline_preds = model_0.predict(val_sentences)
baseline_preds[:20]
# + [markdown] id="K354svk_bmdf"
# ### Creating an evaluation function for our model experiments
#
# We could evaluate these as they are but since we're going to be evaluating several models in the same way going forward, let's create a helper function which takes an array of predictions and ground truth labels and computes the following:
# * Accuracy
# * Precision
# * Recall
# * F1-score
#
# > 🔑 **Note:** Since we're dealing with a classification problem, the above metrics are the most appropriate. If we were working with a regression problem, other metrics such as MAE (mean absolute error) would be a better choice.
# + id="gLmNlDjIxGgJ"
# Function to evaluate: accuracy, precision, recall, f1-score
from sklearn.metrics import accuracy_score, precision_recall_fscore_support
def calculate_results(y_true, y_pred):
"""
Calculates model accuracy, precision, recall and f1 score of a binary classification model.
Args:
-----
y_true = true labels in the form of a 1D array
y_pred = predicted labels in the form of a 1D array
Returns a dictionary of accuracy, precision, recall, f1-score.
"""
# Calculate model accuracy
model_accuracy = accuracy_score(y_true, y_pred) * 100
# Calculate model precision, recall and f1 score using "weighted" average
model_precision, model_recall, model_f1, _ = precision_recall_fscore_support(y_true, y_pred, average="weighted")
model_results = {"accuracy": model_accuracy,
"precision": model_precision,
"recall": model_recall,
"f1": model_f1}
return model_results
# + id="Sgy1omMhwr52" colab={"base_uri": "https://localhost:8080/"} outputId="ff1ea5fd-b76d-447a-e1f9-af4b46e8aaba"
# Get baseline results
baseline_results = calculate_results(y_true=val_labels,
y_pred=baseline_preds)
baseline_results
# + [markdown] id="noRJNm7dGNyh"
# ### Model 1: A simple dense model
#
# The first "deep" model we're going to build is a single layer dense model. In fact, it's barely going to have a single layer.
#
# It'll take our text and labels as input, tokenize the text, create an embedding, find the average of the embedding (using Global Average Pooling) and then pass the average through a fully connected layer with one output unit and a sigmoid activation function.
#
# If the previous sentence sounds like a mouthful, it'll make sense when we code it out (remember, if in doubt, code it out).
#
# And since we're going to be building a number of TensorFlow deep learning models, we'll import our `create_tensorboard_callback()` function from `helper_functions.py` to keep track of the results of each.
# + id="PVMPUd3HTit5"
# Create tensorboard callback (need to create a new one for each model)
from helper_functions import create_tensorboard_callback
# Create directory to save TensorBoard logs
SAVE_DIR = "model_logs"
# + [markdown] id="Pib8hHtu7vt1"
# Now we've got a TensorBoard callback function ready to go, let's build our first deep model.
# + id="a_rVtJA7yVBI"
# Build model with the Functional API
from tensorflow.keras import layers
inputs = layers.Input(shape=(1,), dtype="string") # inputs are 1-dimensional strings
x = text_vectorizer(inputs) # turn the input text into numbers
x = embedding(x) # create an embedding of the numerized numbers
x = layers.GlobalAveragePooling1D()(x) # lower the dimensionality of the embedding (try running the model without this layer and see what happens)
outputs = layers.Dense(1, activation="sigmoid")(x) # create the output layer, want binary outputs so use sigmoid activation
model_1 = tf.keras.Model(inputs, outputs, name="model_1_dense") # construct the model
# + [markdown] id="JYzsu36Y8JUe"
# Looking good. Our model takes a 1-dimensional string as input (in our case, a Tweet), it then tokenizes the string using `text_vectorizer` and creates an embedding using `embedding`.
#
# We then (optionally) pool the outputs of the embedding layer to reduce the dimensionality of the tensor we pass to the output layer.
#
# > 🛠 **Exercise:** Try building `model_1` with and without a `GlobalAveragePooling1D()` layer after the `embedding` layer. What happens? Why do you think this is?
#
# Finally, we pass the output of the pooling layer to a dense layer with sigmoid activation (we use sigmoid since our problem is binary classification).
#
# Before we can fit our model to the data, we've got to compile it. Since we're working with binary classification, we'll use `"binary_crossentropy"` as our loss function and the Adam optimizer.
# + id="Ubq0ctLD8CQq"
# Compile model
model_1.compile(loss="binary_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + [markdown] id="crgltz1O9uku"
# Model compiled. Let's get a summary.
# + id="QkJa-t8aTw1H" colab={"base_uri": "https://localhost:8080/"} outputId="a0c4ada2-1984-41fa-9653-dbfc6bfdfd42"
# Get a summary of the model
model_1.summary()
# + [markdown] id="bH0JLyR09yYt"
# Most of the trainable parameters are contained within the embedding layer. Recall we created an embedding of size 128 (`output_dim=128`) for a vocabulary of size 10,000 (`input_dim=10000`), hence the 1,280,000 trainable parameters.
#
# Alright, our model is compiled, let's fit it to our training data for 5 epochs. We'll also pass our TensorBoard callback function to make sure our model's training metrics are logged.
# + id="1YRYpJIfTvHV" colab={"base_uri": "https://localhost:8080/"} outputId="cc6ad75c-338c-4e79-d6a0-a1a2bf588652"
# Fit the model
model_1_history = model_1.fit(train_sentences, # input sentences can be a list of strings due to text preprocessing layer built-in model
train_labels,
epochs=5,
validation_data=(val_sentences, val_labels),
callbacks=[create_tensorboard_callback(dir_name=SAVE_DIR,
experiment_name="simple_dense_model")])
# + [markdown] id="kZR5_j9C_LW-"
# Nice! Since we're using such a simple model, each epoch processes very quickly.
#
# Let's check our model's performance on the validation set.
# + id="zSTS87YGzuBG" colab={"base_uri": "https://localhost:8080/"} outputId="3dab9b26-7bd0-42c8-8ad8-c55ae73d0e09"
# Check the results
model_1.evaluate(val_sentences, val_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="5M2CTAetBVfW" outputId="263af483-3739-4ff7-fa21-52b9eab7a81b"
embedding.weights
# + colab={"base_uri": "https://localhost:8080/"} id="M3rfhJFSBrga" outputId="e16f8a95-540b-40a7-98f4-8fe1a9dff5ac"
embed_weights = model_1.get_layer("embedding_1").get_weights()[0]
print(embed_weights.shape)
# + [markdown] id="I9dg2aba_VxK"
# And since we tracked our model's training logs with TensorBoard, how about we visualize them?
#
# We can do so by uploading our TensorBoard log files (contained in the `model_logs` directory) to [TensorBoard.dev](https://tensorboard.dev/).
#
# > 🔑 **Note:** Remember, whatever you upload to TensorBoard.dev becomes public. If there are training logs you don't want to share, don't upload them.
# + id="t6UrSgRVU6pl"
# # View tensorboard logs of transfer learning modelling experiments (should be 4 models)
# # Upload TensorBoard dev records
# # !tensorboard dev upload --logdir ./model_logs \
# # --name "First deep model on text data" \
# # --description "Trying a dense model with an embedding layer" \
# # --one_shot # exits the uploader when upload has finished
# + id="DVyJl-VE1ACz"
# If you need to remove previous experiments, you can do so using the following command
# # !tensorboard dev delete --experiment_id EXPERIMENT_ID_TO_DELETE
# + [markdown] id="PkinGcjQ_yI9"
# The TensorBoard.dev experiment for our first deep model can be viewed here: https://tensorboard.dev/experiment/5d1Xm10aT6m6MgyW3HAGfw/
#
# 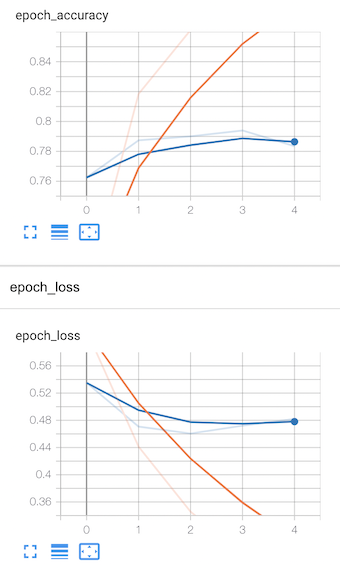
#
# *What the training curves of our model look like on TensorBoard. From looking at the curves can you tell if the model is overfitting or underfitting?*
#
# Beautiful! Those are some colorful training curves. Would you say the model is overfitting or underfitting?
#
# We've built and trained our first deep model, the next step is to make some predictions with it.
# + id="5X7kbEmAzzxM" colab={"base_uri": "https://localhost:8080/"} outputId="5d3a68f3-5b96-4dd4-e8e0-bf351ff68c9f"
# Make predictions (these come back in the form of probabilities)
model_1_pred_probs = model_1.predict(val_sentences)
model_1_pred_probs[:10] # only print out the first 10 prediction probabilities
# + [markdown] id="YWU5e1NLAKJ9"
# Since our final layer uses a sigmoid activation function, we get our predictions back in the form of probabilities.
#
# To convert them to prediction classes, we'll use `tf.round()`, meaning prediction probabilities below 0.5 will be rounded to 0 and those above 0.5 will be rounded to 1.
#
# > 🔑 **Note:** In practice, the output threshold of a sigmoid prediction probability doesn't necessarily have to 0.5. For example, through testing, you may find that a cut off of 0.25 is better for your chosen evaluation metrics. A common example of this threshold cutoff is the [precision-recall tradeoff](https://www.machinelearningaptitude.com/topics/machine-learning/what-is-precision-recall-tradeoff/#:~:text=precision%2Drecall%20tradeoff%20occur%20due,the%20threshold%20of%20the%20classifier.&text=When%20threshold%20is%20decreased%20to,but%20precision%20decreases%20to%200.4.).
# + id="Qf-R_1vsz47P" colab={"base_uri": "https://localhost:8080/"} outputId="3062f826-9564-48d7-bbec-a803b41e9cba"
# Turn prediction probabilities into single-dimension tensor of floats
model_1_preds = tf.squeeze(tf.round(model_1_pred_probs)) # squeeze removes single dimensions
model_1_preds[:20]
# + [markdown] id="Zc3ryY0yCHcI"
# Now we've got our model's predictions in the form of classes, we can use our `calculate_results()` function to compare them to the ground truth validation labels.
# + id="iDEEhYTF0X1y" colab={"base_uri": "https://localhost:8080/"} outputId="e31e8bca-894a-44ad-88cc-fdea0dbc453a"
# Calculate model_1 metrics
model_1_results = calculate_results(y_true=val_labels,
y_pred=model_1_preds)
model_1_results
# + [markdown] id="gnkK6Uc7CYlX"
# How about we compare our first deep model to our baseline model?
# + id="Jp88ystW1m0d" colab={"base_uri": "https://localhost:8080/"} outputId="ba27df1c-624b-4a24-da50-012a56653038"
# Is our simple Keras model better than our baseline model?
import numpy as np
np.array(list(model_1_results.values())) > np.array(list(baseline_results.values()))
# + [markdown] id="lUINrCdRCpFf"
# Since we'll be doing this kind of comparison (baseline compared to new model) quite a few times, let's create a function to help us out.
# + id="wo3norTG3GrE" colab={"base_uri": "https://localhost:8080/"} outputId="14b74922-fe0a-44cb-8932-327e05402960"
# Create a helper function to compare our baseline results to new model results
def compare_baseline_to_new_results(baseline_results, new_model_results):
for key, value in baseline_results.items():
print(f"Baseline {key}: {value:.2f}, New {key}: {new_model_results[key]:.2f}, Difference: {new_model_results[key]-value:.2f}")
compare_baseline_to_new_results(baseline_results=baseline_results,
new_model_results=model_1_results)
# + [markdown] id="6e-1LuioSLAM"
# ## Visualizing learned embeddings
#
# Our first model (`model_1`) contained an embedding layer (`embedding`) which learned a way of representing words as feature vectors by passing over the training data.
#
# Hearing this for the first few times may sound confusing.
#
# So to further help understand what a text embedding is, let's visualize the embedding our model learned.
#
# To do so, let's remind ourselves of the words in our vocabulary.
#
# + id="-DkcfRQBVXuJ" colab={"base_uri": "https://localhost:8080/"} outputId="9e10f4fd-50ef-4ccd-a99b-eff8a64082f8"
# Get the vocabulary from the text vectorization layer
words_in_vocab = text_vectorizer.get_vocabulary()
len(words_in_vocab), words_in_vocab[:10]
# + [markdown] id="KzmAPJXQEx6r"
# And now let's get our embedding layer's weights (these are the numerical representations of each word).
# + id="8EUR9PwrZphh" colab={"base_uri": "https://localhost:8080/"} outputId="eb047f23-dbef-46ea-ce98-2ea91b309d09"
model_1.summary()
# + id="9xJ5LrInWDLo" colab={"base_uri": "https://localhost:8080/"} outputId="ffc27236-56f5-4ecf-e107-37a51722194a"
# Get the weight matrix of embedding layer
# (these are the numerical patterns between the text in the training dataset the model has learned)
embed_weights = model_1.get_layer("embedding_1").get_weights()[0]
print(embed_weights.shape) # same size as vocab size and embedding_dim (each word is a embedding_dim size vector)
# + [markdown] id="jzOJhJHPW1ju"
# Now we've got these two objects, we can use the [Embedding Projector tool](http://projector.tensorflow.org/_) to visualize our embedding.
#
# To use the Embedding Projector tool, we need two files:
# * The embedding vectors (same as embedding weights).
# * The meta data of the embedding vectors (the words they represent - our vocabulary).
#
# Right now, we've got of these files as Python objects. To download them to file, we're going to [use the code example available on the TensorFlow word embeddings tutorial page](https://www.tensorflow.org/tutorials/text/word_embeddings#retrieve_the_trained_word_embeddings_and_save_them_to_disk).
#
# + id="4e9rfcK6WxQE"
# # Code below is adapted from: https://www.tensorflow.org/tutorials/text/word_embeddings#retrieve_the_trained_word_embeddings_and_save_them_to_disk
# import io
# # Create output writers
# out_v = io.open("embedding_vectors.tsv", "w", encoding="utf-8")
# out_m = io.open("embedding_metadata.tsv", "w", encoding="utf-8")
# # Write embedding vectors and words to file
# for num, word in enumerate(words_in_vocab):
# if num == 0:
# continue # skip padding token
# vec = embed_weights[num]
# out_m.write(word + "\n") # write words to file
# out_v.write("\t".join([str(x) for x in vec]) + "\n") # write corresponding word vector to file
# out_v.close()
# out_m.close()
# # Download files locally to upload to Embedding Projector
# try:
# from google.colab import files
# except ImportError:
# pass
# else:
# files.download("embedding_vectors.tsv")
# files.download("embedding_metadata.tsv")
# + [markdown] id="BVM7ifzpZaxJ"
# Once you've downloaded the embedding vectors and metadata, you can visualize them using Embedding Vector tool:
# 1. Go to http://projector.tensorflow.org/
# 2. Click on "Load data"
# 3. Upload the two files you downloaded (`embedding_vectors.tsv` and `embedding_metadata.tsv`)
# 4. Explore
# 5. Optional: You can share the data you've created by clicking "Publish"
#
# What do you find?
#
# Are words with similar meanings close together?
#
# Remember, they might not be. The embeddings we downloaded are how our model interprets words, not necessarily how we interpret them.
#
# Also, since the embedding has been learned purely from Tweets, it may contain some strange values as Tweets are a very unique style of natural language.
#
# > 🤔 **Question:** Do you have to visualize embeddings every time?
#
# No. Although helpful for gaining an intuition of what natural language embeddings are, it's not completely necessary. Especially as the dimensions of your vocabulary and embeddings grow, trying to comprehend them would become an increasingly difficult task.
# + [markdown] id="AcRdDiEtGQj4"
# ## Recurrent Neural Networks (RNN's)
#
# For our next series of modelling experiments we're going to be using a special kind of neural network called a **Recurrent Neural Network (RNN)**.
#
# The premise of an RNN is simple: use information from the past to help you with the future (this is where the term recurrent comes from). In other words, take an input (`X`) and compute an output (`y`) based on all previous inputs.
#
# This concept is especially helpful when dealing with sequences such as passages of natural language text (such as our Tweets).
#
# For example, when you read this sentence, you take into context the previous words when deciphering the meaning of the current word dog.
#
# See what happened there?
#
# I put the word "dog" at the end which is a valid word but it doesn't make sense in the context of the rest of the sentence.
#
# When an RNN looks at a sequence of text (already in numerical form), the patterns it learns are continually updated based on the order of the sequence.
#
# For a simple example, take two sentences:
# 1. Massive earthquake last week, no?
# 2. No massive earthquake last week.
#
# Both contain exactly the same words but have different meaning. The order of the words determines the meaning (one could argue punctuation marks also dictate the meaning but for simplicity sake, let's stay focused on the words).
#
# Recurrent neural networks can be used for a number of sequence-based problems:
# * **One to one:** one input, one output, such as image classification.
# * **One to many:** one input, many outputs, such as image captioning (image input, a sequence of text as caption output).
# * **Many to one:** many inputs, one outputs, such as text classification (classifying a Tweet as real diaster or not real diaster).
# * **Many to many:** many inputs, many outputs, such as machine translation (translating English to Spanish) or speech to text (audio wave as input, text as output).
#
# When you come across RNN's in the wild, you'll most likely come across variants of the following:
# * Long short-term memory cells (LSTMs).
# * Gated recurrent units (GRUs).
# * Bidirectional RNN's (passes forward and backward along a sequence, left to right and right to left).
#
# Going into the details of each these is beyond the scope of this notebook (we're going to focus on using them instead), the main thing you should know for now is that they've proven very effective at modelling sequences.
#
# For a deeper understanding of what's happening behind the scenes of the code we're about to write, I'd recommend the following resources:
#
# > 📖 **Resources:**
# > * [MIT Deep Learning Lecture on Recurrent Neural Networks](https://youtu.be/SEnXr6v2ifU) - explains the background of recurrent neural networks and introduces LSTMs.
# > * [The Unreasonable Effectiveness of Recurrent Neural Networks](http://karpathy.github.io/2015/05/21/rnn-effectiveness/) by <NAME> - demonstrates the power of RNN's with examples generating various sequences.
# > * [Understanding LSTMs](https://colah.github.io/posts/2015-08-Understanding-LSTMs/) by <NAME> - an in-depth (and technical) look at the mechanics of the LSTM cell, possibly the most popular RNN building block.
#
# + [markdown] id="tDERKwP_XWro"
# ### Model 2: LSTM
#
# With all this talk of what RNN's are and what they're good for, I'm sure you're eager to build one.
#
# We're going to start with an LSTM-powered RNN.
#
# To harness the power of the LSTM cell (LSTM cell and LSTM layer are often used interchangably) in TensorFlow, we'll use [`tensorflow.keras.layers.LSTM()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/LSTM).
#
# 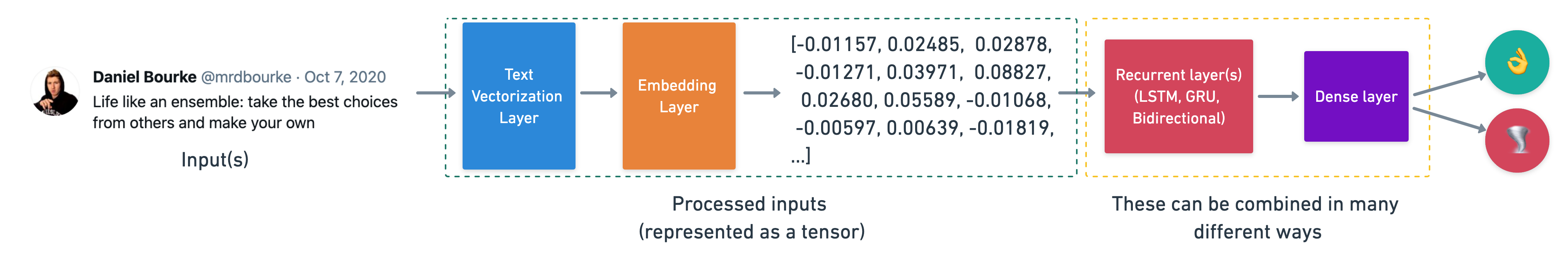
# *Coloured block example of the structure of an recurrent neural network.*
#
# Our model is going to take on a very similar structure to `model_1`:
#
# ```
# Input (text) -> Tokenize -> Embedding -> Layers -> Output (label probability)
# ```
#
# The main difference will be that we're going to add an LSTM layer between our embedding and output.
#
# And to make sure we're not getting reusing trained embeddings (this would involve data leakage between models, leading to an uneven comparison later on), we'll create another embedding layer (`model_2_embedding`) for our model. The `text_vectorizer` layer can be reused since it doesn't get updated during training.
#
# > 🔑 **Note:** The reason we use a new embedding layer for each model is since the embedding layer is a *learned* representation of words (as numbers), if we were to use the same embedding layer (`embedding_1`) for each model, we'd be mixing what one model learned with the next. And because we want to compare our models later on, starting them with their own embedding layer each time is a better idea.
# + id="Pi3vjpFU46hi" colab={"base_uri": "https://localhost:8080/"} outputId="373cad05-9cf7-4c40-b41b-622f51563819"
# Set random seed and create embedding layer (new embedding layer for each model)
tf.random.set_seed(42)
from tensorflow.keras import layers
model_2_embedding = layers.Embedding(input_dim=max_vocab_length,
output_dim=128,
embeddings_initializer="uniform",
input_length=max_length,
name="embedding_2")
# Create LSTM model
inputs = layers.Input(shape=(1,), dtype="string")
x = text_vectorizer(inputs)
x = model_2_embedding(x)
print(x.shape)
# x = layers.LSTM(64, return_sequences=True)(x) # return vector for each word in the Tweet (you can stack RNN cells as long as return_sequences=True)
x = layers.LSTM(64)(x) # return vector for whole sequence
print(x.shape)
# x = layers.Dense(64, activation="relu")(x) # optional dense layer on top of output of LSTM cell
outputs = layers.Dense(1, activation="sigmoid")(x)
model_2 = tf.keras.Model(inputs, outputs, name="model_2_LSTM")
# + [markdown] id="e1wfTARuwWDg"
# > 🔑 **Note:** Reading the documentation for the [TensorFlow LSTM layer](https://www.tensorflow.org/api_docs/python/tf/keras/layers/LSTM), you'll find a plethora of parameters. Many of these have been tuned to make sure they compute as fast as possible. The main ones you'll be looking to adjust are `units` (number of hidden units) and `return_sequences` (set this to `True` when stacking LSTM or other recurrent layers).
#
# Now we've got our LSTM model built, let's compile it using `"binary_crossentropy"` loss and the Adam optimizer.
# + id="pWdt3bFRwG6w"
# Compile model
model_2.compile(loss="binary_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + [markdown] id="I2e_t8RFxgXG"
# And before we fit our model to the data, let's get a summary.
# + id="IAjdfDfLwK_R" colab={"base_uri": "https://localhost:8080/"} outputId="f345c914-32a3-437a-89c9-07ae4989ec62"
model_2.summary()
# + [markdown] id="S5NLw3wD0aMz"
# Looking good! You'll notice a fair few more trainable parameters within our LSTM layer than `model_1`.
#
# If you'd like to know where this number comes from, I recommend going through the above resources as well the following on calculating the number of parameters in an LSTM cell:
# * [Stack Overflow answer to calculate the number of parameters in an LSTM cell](https://stackoverflow.com/questions/38080035/how-to-calculate-the-number-of-parameters-of-an-lstm-network) by <NAME>
# * [Calculating number of parameters in a LSTM unit and layer](https://medium.com/@priyadarshi.cse/calculating-number-of-parameters-in-a-lstm-unit-layer-7e491978e1e4) by <NAME>
#
# Now our first RNN model's compiled let's fit it to our training data, validating it on the validation data and tracking its training parameters using our TensorBoard callback.
# + id="YgZ7ojDvwKcq" colab={"base_uri": "https://localhost:8080/"} outputId="34da9032-cfe3-46b2-8cab-80d6abc3a593"
# Fit model
model_2_history = model_2.fit(train_sentences,
train_labels,
epochs=5,
validation_data=(val_sentences, val_labels),
callbacks=[create_tensorboard_callback(SAVE_DIR,
"LSTM")])
# + [markdown] id="1gikGe_Z16PP"
# Nice! We've got our first trained RNN model using LSTM cells. Let's make some predictions with it.
#
# The same thing will happen as before, due to the sigmoid activiation function in the final layer, when we call the `predict()` method on our model, it'll return prediction probabilities rather than classes.
# + id="4c_lVbKLemrU" colab={"base_uri": "https://localhost:8080/"} outputId="08161e1d-fb72-4046-8ba1-1937487eebc4"
# Make predictions on the validation dataset
model_2_pred_probs = model_2.predict(val_sentences)
model_2_pred_probs.shape, model_2_pred_probs[:10] # view the first 10
# + [markdown] id="fQ6ope-ddpOo"
# We can turn these prediction probabilities into prediction classes by rounding to the nearest integer (by default, prediction probabilities under 0.5 will go to 0 and those over 0.5 will go to 1).
# + id="iFnIhtyE7hlb" colab={"base_uri": "https://localhost:8080/"} outputId="284865b4-bfd5-470d-b3da-c3bef6f7ea92"
# Round out predictions and reduce to 1-dimensional array
model_2_preds = tf.squeeze(tf.round(model_2_pred_probs))
model_2_preds[:10]
# + [markdown] id="zTBy4poXd_7p"
# Beautiful, now let's use our `caculate_results()` function to evaluate our LSTM model and our `compare_baseline_to_new_results()` function to compare it to our baseline model.
# + id="3iHXv04y76vj" colab={"base_uri": "https://localhost:8080/"} outputId="9265fa0c-a725-4333-f91f-4d96fa5a5c06"
# Calculate LSTM model results
model_2_results = calculate_results(y_true=val_labels,
y_pred=model_2_preds)
model_2_results
# + id="ZdQGn2L68B5Q" colab={"base_uri": "https://localhost:8080/"} outputId="686d82ce-8eb5-40d3-9db2-85ffc4fa6484"
# Compare model 2 to baseline
compare_baseline_to_new_results(baseline_results, model_2_results)
# + [markdown] id="Q0pAtADt8ju7"
# ### Model 3: GRU
#
# Another popular and effective RNN component is the GRU or gated recurrent unit.
#
# The GRU cell has similar features to an LSTM cell but has less parameters.
#
# > 📖 **Resource:** A full explanation of the GRU cell is beyond the scope of this noteook but I'd suggest the following resources to learn more:
# * [Gated Recurrent Unit](https://en.wikipedia.org/wiki/Gated_recurrent_unit) Wikipedia page
# * [Understanding GRU networks](https://towardsdatascience.com/understanding-gru-networks-2ef37df6c9be) by <NAME>
#
# To use the GRU cell in TensorFlow, we can call the [`tensorflow.keras.layers.GRU()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/GRU) class.
#
# The architecture of the GRU-powered model will follow the same structure we've been using:
#
# ```
# Input (text) -> Tokenize -> Embedding -> Layers -> Output (label probability)
# ```
#
# Again, the only difference will be the layer(s) we use between the embedding and the output.
# + id="SoSCGq3H47Yo"
# Set random seed and create embedding layer (new embedding layer for each model)
tf.random.set_seed(42)
from tensorflow.keras import layers
model_3_embedding = layers.Embedding(input_dim=max_vocab_length,
output_dim=128,
embeddings_initializer="uniform",
input_length=max_length,
name="embedding_3")
# Build an RNN using the GRU cell
inputs = layers.Input(shape=(1,), dtype="string")
x = text_vectorizer(inputs)
x = model_3_embedding(x)
# x = layers.GRU(64, return_sequences=True) # stacking recurrent cells requires return_sequences=True
x = layers.GRU(64)(x)
# x = layers.Dense(64, activation="relu")(x) # optional dense layer after GRU cell
outputs = layers.Dense(1, activation="sigmoid")(x)
model_3 = tf.keras.Model(inputs, outputs, name="model_3_GRU")
# + [markdown] id="JLT5maFWhKH1"
# TensorFlow makes it easy to use powerful components such as the GRU cell in our models. And now our third model is built, let's compile it, just as before.
# + id="lBL1mb31hHDS"
# Compile GRU model
model_3.compile(loss="binary_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + [markdown] id="yvnksvkmha2A"
# What does a summary of our model look like?
# + id="JVnB5yQeiAWs" colab={"base_uri": "https://localhost:8080/"} outputId="068184f9-c913-46fd-83d9-8547e8f4f1f6"
# Get a summary of the GRU model
model_3.summary()
# + [markdown] id="KcXzKqgXhdez"
# Notice the difference in number of trainable parameters between `model_2` (LSTM) and `model_3` (GRU). The difference comes from the LSTM cell having more trainable parameters than the GRU cell.
#
# We'll fit our model just as we've been doing previously. We'll also track our models results using our `create_tensorboard_callback()` function.
# + id="Gvamg5JOh_jC" colab={"base_uri": "https://localhost:8080/"} outputId="629279a6-1dd9-43fb-f0ae-975be030ebe7"
# Fit model
model_3_history = model_3.fit(train_sentences,
train_labels,
epochs=5,
validation_data=(val_sentences, val_labels),
callbacks=[create_tensorboard_callback(SAVE_DIR, "GRU")])
# + [markdown] id="hM4mQj1Sh7Gn"
# Due to the optimized default settings of the GRU cell in TensorFlow, training doesn't take long at all.
#
# Time to make some predictions on the validation samples.
# + id="W5TUVHCl9pe-" colab={"base_uri": "https://localhost:8080/"} outputId="21116150-7412-49c4-ae12-679f3c688c44"
# Make predictions on the validation data
model_3_pred_probs = model_3.predict(val_sentences)
model_3_pred_probs.shape, model_3_pred_probs[:10]
# + [markdown] id="hasS7dzRiYQh"
# Again we get an array of prediction probabilities back which we can convert to prediction classes by rounding them.
# + id="haILbddg98CY" colab={"base_uri": "https://localhost:8080/"} outputId="8d567022-4f1f-46cb-aae1-70242ce55e56"
# Convert prediction probabilities to prediction classes
model_3_preds = tf.squeeze(tf.round(model_3_pred_probs))
model_3_preds[:10]
# + [markdown] id="_7yAgh-viglB"
# Now we've got predicted classes, let's evaluate them against the ground truth labels.
# + id="h9OZbQu1-LPp" colab={"base_uri": "https://localhost:8080/"} outputId="3b32ee9f-3291-4f8c-873d-c7780488cab8"
# Calcuate model_3 results
model_3_results = calculate_results(y_true=val_labels,
y_pred=model_3_preds)
model_3_results
# + [markdown] id="o9t7wcALiuRk"
# Finally we can compare our GRU model's results to our baseline.
# + id="_7AE6vtn-RQZ" colab={"base_uri": "https://localhost:8080/"} outputId="ff8ad89a-089a-4b29-be3f-b0c6f87c19ad"
# Compare to baseline
compare_baseline_to_new_results(baseline_results, model_3_results)
# + [markdown] id="oLm6r4nQ-Wdr"
# ### Model 4: Bidirectonal RNN model
#
# Look at us go! We've already built two RNN's with GRU and LSTM cells. Now we're going to look into another kind of RNN, the bidirectional RNN.
#
# A standard RNN will process a sequence from left to right, where as a bidirectional RNN will process the sequence from left to right and then again from right to left.
#
# Intuitively, this can be thought of as if you were reading a sentence for the first time in the normal fashion (left to right) but for some reason it didn't make sense so you traverse back through the words and go back over them again (right to left).
#
# In practice, many sequence models often see and improvement in performance when using bidirectional RNN's.
#
# However, this improvement in performance often comes at the cost of longer training times and increased model parameters (since the model goes left to right and right to left, the number of trainable parameters doubles).
#
# Okay enough talk, let's build a bidirectional RNN.
#
# Once again, TensorFlow helps us out by providing the [`tensorflow.keras.layers.Bidirectional`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Bidirectional) class. We can use the `Bidirectional` class to wrap our existing RNNs, instantly making them bidirectional.
# + id="NAU9dvGm47_2"
# Set random seed and create embedding layer (new embedding layer for each model)
tf.random.set_seed(42)
from tensorflow.keras import layers
model_4_embedding = layers.Embedding(input_dim=max_vocab_length,
output_dim=128,
embeddings_initializer="uniform",
input_length=max_length,
name="embedding_4")
# Build a Bidirectional RNN in TensorFlow
inputs = layers.Input(shape=(1,), dtype="string")
x = text_vectorizer(inputs)
x = model_4_embedding(x)
# x = layers.Bidirectional(layers.LSTM(64, return_sequences=True))(x) # stacking RNN layers requires return_sequences=True
x = layers.Bidirectional(layers.LSTM(64))(x) # bidirectional goes both ways so has double the parameters of a regular LSTM layer
outputs = layers.Dense(1, activation="sigmoid")(x)
model_4 = tf.keras.Model(inputs, outputs, name="model_4_Bidirectional")
# + [markdown] id="9Hm5cwmNm-g4"
# > 🔑 **Note:** You can use the `Bidirectional` wrapper on any RNN cell in TensorFlow. For example, `layers.Bidirectional(layers.GRU(64))` creates a bidirectional GRU cell.
#
# Our bidirectional model is built, let's compile it.
# + id="wP1jeF0am9x0"
# Compile
model_4.compile(loss="binary_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + [markdown] id="NtpYyjsbnEwN"
# And of course, we'll check out a summary.
# + id="-sUd9AQ6nFXI" colab={"base_uri": "https://localhost:8080/"} outputId="dc1d6295-7753-41b9-df0d-4f4986cb3cbf"
# Get a summary of our bidirectional model
model_4.summary()
# + [markdown] id="TvItfzeZnIE-"
# Notice the increased number of trainable parameters in `model_4` (bidirectional LSTM) compared to `model_2` (regular LSTM). This is due to the bidirectionality we added to our RNN.
#
# Time to fit our bidirectional model and track its performance.
# + id="bAKY_QbHXPHB" colab={"base_uri": "https://localhost:8080/"} outputId="1e576aae-b7a6-4bc5-b229-99e0885d9bd2"
# Fit the model (takes longer because of the bidirectional layers)
model_4_history = model_4.fit(train_sentences,
train_labels,
epochs=5,
validation_data=(val_sentences, val_labels),
callbacks=[create_tensorboard_callback(SAVE_DIR, "bidirectional_RNN")])
# + [markdown] id="zkt8GVRHoJz6"
# Due to the bidirectionality of our model we see a slight increase in training time.
#
# Not to worry, it's not too dramatic of an increase.
#
# Let's make some predictions with it.
# + id="uFc7QHRtXmn7" colab={"base_uri": "https://localhost:8080/"} outputId="67a915b9-2e27-492e-86a7-6b114354cfcf"
# Make predictions with bidirectional RNN on the validation data
model_4_pred_probs = model_4.predict(val_sentences)
model_4_pred_probs[:10]
# + [markdown] id="L_9HmNIYobDB"
# And we'll convert them to prediction classes and evaluate them against the ground truth labels and baseline model.
# + id="G5z8bMdaXw51" colab={"base_uri": "https://localhost:8080/"} outputId="ecbb2c9e-5b45-4872-a126-6f6b4de3f091"
# Convert prediction probabilities to labels
model_4_preds = tf.squeeze(tf.round(model_4_pred_probs))
model_4_preds[:10]
# + id="-a7Ym_vKYAO4" colab={"base_uri": "https://localhost:8080/"} outputId="a3d841a2-6cdd-4da2-d623-d046f608841c"
# Calculate bidirectional RNN model results
model_4_results = calculate_results(val_labels, model_4_preds)
model_4_results
# + id="hAET-LKpYT18" colab={"base_uri": "https://localhost:8080/"} outputId="d69437c7-e780-41ab-d575-b9cfdefdf82b"
# Check to see how the bidirectional model performs against the baseline
compare_baseline_to_new_results(baseline_results, model_4_results)
# + [markdown] id="wcvt_7emuKlR"
# ## Convolutional Neural Networks for Text
#
# You might've used convolutional neural networks (CNNs) for images before but they can also be used for sequences.
#
# The main difference between using CNNs for images and sequences is the shape of the data. Images come in 2-dimensions (height x width) where as sequences are often 1-dimensional (a string of text).
#
# So to use CNNs with sequences, we use a 1-dimensional convolution instead of a 2-dimensional convolution.
#
# A typical CNN architecture for sequences will look like the following:
#
# ```
# Inputs (text) -> Tokenization -> Embedding -> Layers -> Outputs (class probabilities)
# ```
#
# You might be thinking "that just looks like the architecture layout we've been using for the other models..."
#
# And you'd be right.
#
# The difference again is in the layers component. Instead of using an LSTM or GRU cell, we're going to use a [`tensorflow.keras.layers.Conv1D()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv1D) layer followed by a [`tensorflow.keras.layers.GlobablMaxPool1D()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/GlobalMaxPool1D) layer.
#
# > 📖 **Resource:** The intuition here is explained succinctly in the paper [*Understanding Convolutional Neural Networks for Text Classification*](https://www.aclweb.org/anthology/W18-5408.pdf), where they state that CNNs classify text through the following steps:
# 1. 1-dimensional convolving filters are used as ngram detectors, each filter specializing in a closely-related family of ngrams (an ngram is a collection of n-words, for example, an ngram of 5 might result in "hello, my name is Daniel").
# 2. Max-pooling over time extracts the relevant ngrams for making a decision.
# 3. The rest of the network classifies the text based on this information.
#
# >
#
# + [markdown] id="lgXEorf9GWY1"
# ### Model 5: Conv1D
#
# Before we build a full 1-dimensional CNN model, let's see a 1-dimensional convolutional layer (also called a **temporal convolution**) in action.
#
# We'll first create an embedding of a sample of text and experiment passing it through a `Conv1D()` layer and `GlobalMaxPool1D()` layer.
# + id="563hl7nPWP_3" colab={"base_uri": "https://localhost:8080/"} outputId="32a7e900-e4b4-4b1b-9b4e-617ca9f7eb92"
# Test out the embedding, 1D convolutional and max pooling
embedding_test = embedding(text_vectorizer(["this is a test sentence"])) # turn target sentence into embedding
conv_1d = layers.Conv1D(filters=32, kernel_size=5, activation="relu") # convolve over target sequence 5 words at a time
conv_1d_output = conv_1d(embedding_test) # pass embedding through 1D convolutional layer
max_pool = layers.GlobalMaxPool1D()
max_pool_output = max_pool(conv_1d_output) # get the most important features
embedding_test.shape, conv_1d_output.shape, max_pool_output.shape
# + [markdown] id="-WzTeShEemJ2"
# Notice the output shapes of each layer.
#
# The embedding has an output shape dimension of the parameters we set it to (`input_length=15` and `output_dim=128`).
#
# The 1-dimensional convolutional layer has an output which has been compressed inline with its parameters. And the same goes for the max pooling layer output.
#
# Our text starts out as a string but gets converted to a feature vector of length 64 through various transformation steps (from tokenization to embedding to 1-dimensional convolution to max pool).
#
# Let's take a peak at what each of these transformations looks like.
# + id="gRcxYgs-dxM8" colab={"base_uri": "https://localhost:8080/"} outputId="33616ed5-7333-4891-d3fa-d798c64fd789"
# See the outputs of each layer
embedding_test[:1], conv_1d_output[:1], max_pool_output[:1]
# + [markdown] id="kMcrthJwg3B2"
# Alright, we've seen the outputs of several components of a CNN for sequences, let's put them together and construct a full model, compile it (just as we've done with our other models) and get a summary.
# + id="G9aphPWCYkWN" colab={"base_uri": "https://localhost:8080/"} outputId="6d723bc4-84d1-49eb-9ef9-0887364f029b"
# Set random seed and create embedding layer (new embedding layer for each model)
tf.random.set_seed(42)
from tensorflow.keras import layers
model_5_embedding = layers.Embedding(input_dim=max_vocab_length,
output_dim=128,
embeddings_initializer="uniform",
input_length=max_length,
name="embedding_5")
# Create 1-dimensional convolutional layer to model sequences
from tensorflow.keras import layers
inputs = layers.Input(shape=(1,), dtype="string")
x = text_vectorizer(inputs)
x = model_5_embedding(x)
x = layers.Conv1D(filters=32, kernel_size=5, activation="relu")(x)
x = layers.GlobalMaxPool1D()(x)
# x = layers.Dense(64, activation="relu")(x) # optional dense layer
outputs = layers.Dense(1, activation="sigmoid")(x)
model_5 = tf.keras.Model(inputs, outputs, name="model_5_Conv1D")
# Compile Conv1D model
model_5.compile(loss="binary_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Get a summary of our 1D convolution model
model_5.summary()
# + [markdown] id="o1Y4BpMGh0jG"
# Woohoo! Looking great! Notice how the number of trainable parameters for the 1-dimensional convolutional layer is similar to that of the LSTM layer in `model_2`.
#
# Let's fit our 1D CNN model to our text data. In line with previous experiments, we'll save its results using our `create_tensorboard_callback()` function.
# + id="9fzlaKm1ZrMX" colab={"base_uri": "https://localhost:8080/"} outputId="544b3d97-2d9c-40a3-b0e5-51feac67a4fa"
# Fit the model
model_5_history = model_5.fit(train_sentences,
train_labels,
epochs=5,
validation_data=(val_sentences, val_labels),
callbacks=[create_tensorboard_callback(SAVE_DIR,
"Conv1D")])
# + [markdown] id="d2up-1tLiXKD"
# Nice! Thanks to GPU acceleration, our 1D convolutional model trains nice and fast. Let's make some predictions with it and evaluate them just as before.
# + id="ZHYw5GkxZ2OK" colab={"base_uri": "https://localhost:8080/"} outputId="44a902e9-9710-4854-d61d-649b7a6f5ef3"
# Make predictions with model_5
model_5_pred_probs = model_5.predict(val_sentences)
model_5_pred_probs[:10]
# + id="v9YqTtjiaauS" colab={"base_uri": "https://localhost:8080/"} outputId="0bb1e353-5ecc-4cdb-cd73-07911d7857c2"
# Convert model_5 prediction probabilities to labels
model_5_preds = tf.squeeze(tf.round(model_5_pred_probs))
model_5_preds[:10]
# + id="wMY3s1Pnaj34" colab={"base_uri": "https://localhost:8080/"} outputId="d45f024e-6352-4583-8946-bf3da8249a21"
# Calculate model_5 evaluation metrics
model_5_results = calculate_results(y_true=val_labels,
y_pred=model_5_preds)
model_5_results
# + id="wRfF4B6_at8k" colab={"base_uri": "https://localhost:8080/"} outputId="8de35775-b434-4465-e63c-0066a7fa3f69"
# Compare model_5 results to baseline
compare_baseline_to_new_results(baseline_results, model_5_results)
# + [markdown] id="g_roVSSRt-7h"
# ## Using Pretrained Embeddings (transfer learning for NLP)
#
# For all of the previous deep learning models we've built and trained, we've created and used our own embeddings from scratch each time.
#
# However, a common practice is to leverage pretrained embeddings through **transfer learning**. This is one of the main benefits of using deep models: being able to take what one (often larger) model has learned (often on a large amount of data) and adjust it for our own use case.
#
# For our next model, instead of using our own embedding layer, we're going to replace it with a pretrained embedding layer.
#
# More specifically, we're going to be using the [Universal Sentence Encoder](https://www.aclweb.org/anthology/D18-2029.pdf) from [TensorFlow Hub](https://tfhub.dev/google/universal-sentence-encoder/4) (a great resource containing a plethora of pretrained model resources for a variety of tasks).
#
# > 🔑 **Note:** There are many different pretrained text embedding options on TensorFlow Hub, however, some require different levels of text preprocessing than others. Best to experiment with a few and see which best suits your use case.
#
#
# + [markdown] id="R-NQ2MA5GZBo"
# ### Model 6: TensorFlow Hub Pretrained Sentence Encoder
#
# The main difference between the embedding layer we created and the Universal Sentence Encoder is that rather than create a word-level embedding, the Universal Sentence Encoder, as you might've guessed, creates a whole sentence-level embedding.
#
# Our embedding layer also outputs an a 128 dimensional vector for each word, where as, the Universal Sentence Encoder outputs a 512 dimensional vector for each sentence.
#
# 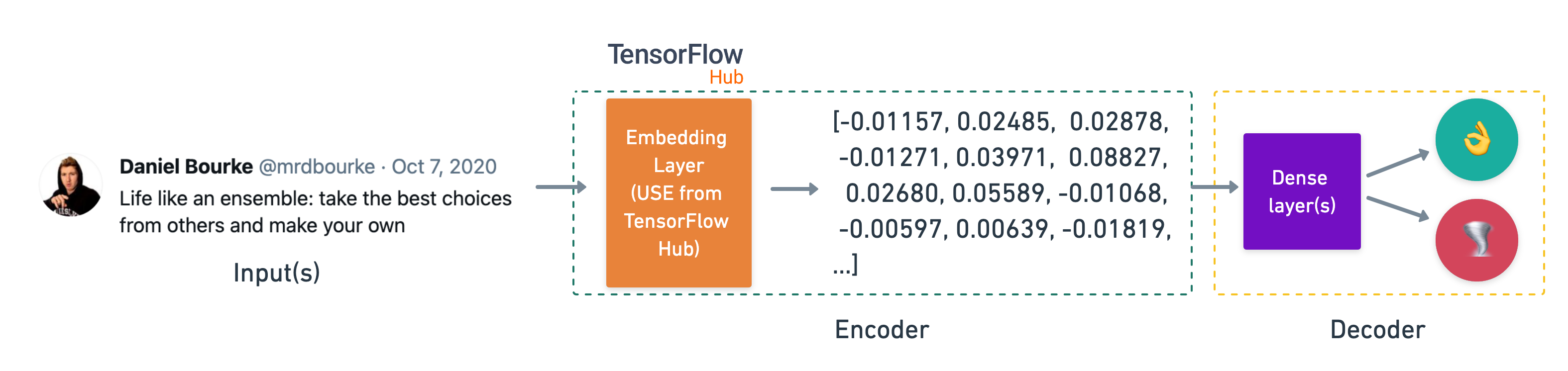
# *The feature extractor model we're building through the eyes of an **encoder/decoder** model.*
#
# > 🔑 **Note:** An **encoder** is the name for a model which converts raw data such as text into a numerical representation (feature vector), a **decoder** converts the numerical representation to a desired output.
#
# As usual, this is best demonstrated with an example.
#
# We can load in a TensorFlow Hub module using the [`hub.load()`](https://www.tensorflow.org/hub/api_docs/python/hub/load) method and passing it the target URL of the module we'd like to use, in our case, it's "https://tfhub.dev/google/universal-sentence-encoder/4".
#
# Let's load the Universal Sentence Encoder model and test it on a couple of sentences.
# + id="7piW5jtxbUkV" colab={"base_uri": "https://localhost:8080/"} outputId="fc509df1-e7a6-4145-82ca-674785ac5257"
# Example of pretrained embedding with universal sentence encoder - https://tfhub.dev/google/universal-sentence-encoder/4
import tensorflow_hub as hub
embed = hub.load("https://tfhub.dev/google/universal-sentence-encoder/4") # load Universal Sentence Encoder
embed_samples = embed([sample_sentence,
"When you call the universal sentence encoder on a sentence, it turns it into numbers."])
print(embed_samples[0][:50])
# + id="vvArnKkGb4vu" colab={"base_uri": "https://localhost:8080/"} outputId="67d73b29-193e-4e95-bd8d-e071b551ec3b"
# Each sentence has been encoded into a 512 dimension vector
embed_samples[0].shape
# + [markdown] id="ZxYFDkGD-XjF"
# Passing our sentences to the Universal Sentence Encoder (USE) encodes them from strings to 512 dimensional vectors, which make no sense to us but hopefully make sense to our machine learning models.
#
# Speaking of models, let's build one with the USE as our embedding layer.
#
# We can convert the TensorFlow Hub USE module into a Keras layer using the [`hub.KerasLayer`](https://www.tensorflow.org/hub/api_docs/python/hub/KerasLayer) class.
#
# > 🔑 **Note:** Due to the size of the USE TensorFlow Hub module, it may take a little while to download. Once it's downloaded though, it'll be cached and ready to use. And as with many TensorFlow Hub modules, there is a ["lite" version of the USE](https://tfhub.dev/google/universal-sentence-encoder-lite/2) which takes up less space but sacrifices some performance and requires more preprocessing steps. However, depending on your available compute power, the lite version may be better for your application use case.
# + id="ZcbBj0aXqrs9"
# We can use this encoding layer in place of our text_vectorizer and embedding layer
sentence_encoder_layer = hub.KerasLayer("https://tfhub.dev/google/universal-sentence-encoder/4",
input_shape=[], # shape of inputs coming to our model
dtype=tf.string, # data type of inputs coming to the USE layer
trainable=False, # keep the pretrained weights (we'll create a feature extractor)
name="USE")
# + [markdown] id="WvjQl4p7BO_A"
# Beautiful! Now we've got the USE as a Keras layer, we can use it in a Keras Sequential model.
# + id="M_pjIvPuYltA" colab={"base_uri": "https://localhost:8080/"} outputId="141cb32c-b1a7-46e8-8b95-12c7ec4d589e"
# Create model using the Sequential API
model_6 = tf.keras.Sequential([
sentence_encoder_layer, # take in sentences and then encode them into an embedding
layers.Dense(64, activation="relu"),
layers.Dense(1, activation="sigmoid")
], name="model_6_USE")
# Compile model
model_6.compile(loss="binary_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
model_6.summary()
# + [markdown] id="yukgxOgCCR2Z"
# Notice the number of paramters in the USE layer, these are the pretrained weights its learned on various text sources (Wikipedia, web news, web question-answer forums, etc, see the [Universal Sentence Encoder paper](https://www.aclweb.org/anthology/D18-2029.pdf) for more).
#
# The trainable parameters are only in our output layers, in other words, we're keeping the USE weights frozen and using it as a feature-extractor. We could fine-tune these weights by setting `trainable=True` when creating the `hub.KerasLayer` instance.
#
# Now we've got a feature extractor model ready, let's train it and track its results to TensorBoard using our `create_tensorboard_callback()` function.
# + id="uX9S0YvafybG" colab={"base_uri": "https://localhost:8080/"} outputId="f4c3c8d1-b3cf-4c87-dd75-2ed786b5e5d5"
# Train a classifier on top of pretrained embeddings
model_6_history = model_6.fit(train_sentences,
train_labels,
epochs=5,
validation_data=(val_sentences, val_labels),
callbacks=[create_tensorboard_callback(SAVE_DIR,
"tf_hub_sentence_encoder")])
# + [markdown] id="KeI0kvVVDmbl"
# USE model trained! Let's make some predictions with it an evaluate them as we've done with our other models.
# + id="xeyNXqU-gM2p" colab={"base_uri": "https://localhost:8080/"} outputId="cbbc27f6-ee98-4633-f703-7a186fc6c932"
# Make predictions with USE TF Hub model
model_6_pred_probs = model_6.predict(val_sentences)
model_6_pred_probs[:10]
# + id="Gbn1Z0FfgVdx" colab={"base_uri": "https://localhost:8080/"} outputId="101ab428-2f85-42a0-b308-feb7888f8b5d"
# Convert prediction probabilities to labels
model_6_preds = tf.squeeze(tf.round(model_6_pred_probs))
model_6_preds[:10]
# + id="N2Ow2de3okcb" colab={"base_uri": "https://localhost:8080/"} outputId="0b82286e-5589-4ddc-8982-d1dd0da78f49"
# Calculate model 6 performance metrics
model_6_results = calculate_results(val_labels, model_6_preds)
model_6_results
# + id="-BHnRHHHgp1r" colab={"base_uri": "https://localhost:8080/"} outputId="0ab489c1-4f16-428d-b391-46a95f10c044"
# Compare TF Hub model to baseline
compare_baseline_to_new_results(baseline_results, model_6_results)
# + [markdown] id="LHwu4QjijYWG"
# ### Model 7: TensorFlow Hub Pretrained Sentence Encoder 10% of the training data
#
# One of the benefits of using transfer learning methods, such as, the pretrained embeddings within the USE is the ability to get great results on a small amount of data (the USE paper even mentions this in the abstract).
#
# To put this to the test, we're going to make a small subset of the training data (10%), train a model and evaluate it.
# + id="W5Sal8DpjzWm"
### NOTE: Making splits like this will lead to data leakage ###
### (some of the training examples in the validation set) ###
### WRONG WAY TO MAKE SPLITS (train_df_shuffled has already been split) ###
# # Create subsets of 10% of the training data
# train_10_percent = train_df_shuffled[["text", "target"]].sample(frac=0.1, random_state=42)
# train_sentences_10_percent = train_10_percent["text"].to_list()
# train_labels_10_percent = train_10_percent["target"].to_list()
# len(train_sentences_10_percent), len(train_labels_10_percent)
# + id="XHgowC3GUPJH"
# One kind of correct way (there are more) to make data subset
# (split the already split train_sentences/train_labels)
train_sentences_90_percent, train_sentences_10_percent, train_labels_90_percent, train_labels_10_percent = train_test_split(np.array(train_sentences),
train_labels,
test_size=0.1,
random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="j8jaydmiVnJP" outputId="17634293-52c5-4a51-e4ef-b5521d339bb3"
# Check length of 10 percent datasets
print(f"Total training examples: {len(train_sentences)}")
print(f"Length of 10% training examples: {len(train_sentences_10_percent)}")
# + [markdown] id="7E2jr7rSEYT8"
# Because we've selected a random subset of the training samples, the classes should be roughly balanced (as they are in the full training dataset).
# + id="V0lEpFT0k0RB" colab={"base_uri": "https://localhost:8080/"} outputId="140c4560-e0a5-472e-bdc2-47374fe7f95e"
# Check the number of targets in our subset of data
# (this should be close to the distribution of labels in the original train_labels)
pd.Series(train_labels_10_percent).value_counts()
# + [markdown] id="ghl1qeGOEnXG"
# To make sure we're making an appropriate comparison between our model's ability to learn from the full training set and 10% subset, we'll clone our USE model (`model_6`) using the [`tf.keras.models.clone_model()`](https://www.tensorflow.org/api_docs/python/tf/keras/models/clone_model) method.
#
# Doing this will create the same architecture but reset the learned weights of the clone target (pretrained weights from the USE will remain but all others will be reset).
# + id="PGmxeAOBjdg2" colab={"base_uri": "https://localhost:8080/"} outputId="2c321240-d2c1-4afe-e8a0-fa35fe64038e"
# Clone model_6 but reset weights
model_7 = tf.keras.models.clone_model(model_6)
# Compile model
model_7.compile(loss="binary_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Get a summary (will be same as model_6)
model_7.summary()
# + [markdown] id="LxFkEM_aFoLK"
# Notice the layout of `model_7` is the same as `model_6`. Now let's train the newly created model on our 10% training data subset.
# + id="LklU2maOkgUF" colab={"base_uri": "https://localhost:8080/"} outputId="65c50e4b-73f1-4012-af7a-f6ec5e2a1591"
# Fit the model to 10% of the training data
model_7_history = model_7.fit(x=train_sentences_10_percent,
y=train_labels_10_percent,
epochs=5,
validation_data=(val_sentences, val_labels),
callbacks=[create_tensorboard_callback(SAVE_DIR, "10_percent_tf_hub_sentence_encoder")])
# + [markdown] id="9Qpyqdh-F6Eh"
# Due to the smaller amount of training data, training happens even quicker than before.
#
# Let's evaluate our model's performance after learning on 10% of the training data.
# + id="ot6MRnznlgCL" colab={"base_uri": "https://localhost:8080/"} outputId="3e9b410c-117c-4292-b33e-206ebf4de1a9"
# Make predictions with the model trained on 10% of the data
model_7_pred_probs = model_7.predict(val_sentences)
model_7_pred_probs[:10]
# + id="Vj_4aZellpRu" colab={"base_uri": "https://localhost:8080/"} outputId="6ce5ceab-3cbf-4d0d-c762-f4c3893affe1"
# Convert prediction probabilities to labels
model_7_preds = tf.squeeze(tf.round(model_7_pred_probs))
model_7_preds[:10]
# + id="T_lTXrDblyva" colab={"base_uri": "https://localhost:8080/"} outputId="ad1a4228-9944-4e17-ecae-6c864c3a51fa"
# Calculate model results
model_7_results = calculate_results(val_labels, model_7_preds)
model_7_results
# + id="G84ezltll6DT" colab={"base_uri": "https://localhost:8080/"} outputId="3d966d5b-0c50-48e9-bb29-1e55063a06e8"
# Compare to baseline
compare_baseline_to_new_results(baseline_results, model_7_results)
# + [markdown] id="iBs9V61EGh0J"
# ## Comparing the performance of each of our models
#
# Woah. We've come a long way! From training a baseline to several deep models.
#
# Now it's time to compare our model's results.
#
# But just before we do, it's worthwhile mentioning, this type of practice is a standard deep learning workflow. Training various different models, then comparing them to see which one performed best and continuing to train it if necessary.
#
# The important thing to note is that for all of our modelling experiments we used the same training data (except for `model_7` where we used 10% of the training data).
#
# To visualize our model's performances, let's create a pandas DataFrame we our results dictionaries and then plot it.
# + id="Ex0NSaz7lRf-" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="421ea5ed-744d-4bc8-a0fb-b79e367f5ec3"
# Combine model results into a DataFrame
all_model_results = pd.DataFrame({"baseline": baseline_results,
"simple_dense": model_1_results,
"lstm": model_2_results,
"gru": model_3_results,
"bidirectional": model_4_results,
"conv1d": model_5_results,
"tf_hub_sentence_encoder": model_6_results,
"tf_hub_10_percent_data": model_7_results})
all_model_results = all_model_results.transpose()
all_model_results
# + id="v-s2DSLpmM1F"
# Reduce the accuracy to same scale as other metrics
all_model_results["accuracy"] = all_model_results["accuracy"]/100
# + id="Wp69bR8umD5g" colab={"base_uri": "https://localhost:8080/", "height": 546} outputId="cb7ae0e4-2a9c-4ef3-d23a-a9a93a6a992d"
# Plot and compare all of the model results
all_model_results.plot(kind="bar", figsize=(10, 7)).legend(bbox_to_anchor=(1.0, 1.0));
# + [markdown] id="avbdkiIuKNNr"
# Looks like our pretrained USE TensorFlow Hub models have the best performance, even the one with only 10% of the training data seems to outperform the other models. This goes to show the power of transfer learning.
#
# How about we drill down and get the F1-score's of each model?
# + id="yktdOiufmm3p" colab={"base_uri": "https://localhost:8080/", "height": 546} outputId="97090bc0-63d5-41e2-f7b1-deddfcd99f53"
# Sort model results by f1-score
all_model_results.sort_values("f1", ascending=False)["f1"].plot(kind="bar", figsize=(10, 7));
# + [markdown] id="pv2iE0TPGdNy"
# Drilling down into a single metric we see our USE TensorFlow Hub models performing better than all of the other models. Interestingly, the baseline's F1-score isn't too far off the rest of the deeper models.
#
# We can also visualize all of our model's training logs using TensorBoard.dev.
# + id="2Ca8TalwGhPf"
# # View tensorboard logs of transfer learning modelling experiments (should be 4 models)
# # Upload TensorBoard dev records
# # !tensorboard dev upload --logdir ./model_logs \
# # --name "NLP modelling experiments" \
# # --description "A series of different NLP modellings experiments with various models" \
# # --one_shot # exits the uploader when upload has finished
# + [markdown] id="uIYVXCUJ3FBn"
# The TensorBoard logs of the different modelling experiments we ran can be viewed here: https://tensorboard.dev/experiment/LkoAakb7QIKBZ0RL97cXbw/
# + id="Os7dv00u21jg"
# If you need to remove previous experiments, you can do so using the following command
# # !tensorboard dev delete --experiment_id EXPERIMENT_ID_TO_DELETE
# + [markdown] id="GGVZhTTiGdd5"
# ## Combining our models (model ensembling/stacking)
#
# Many production systems use an **ensemble** (multiple different models combined) of models to make a prediction.
#
# The idea behind model stacking is that if several uncorrelated models agree on a prediction, then the prediction must be more robust than a prediction made by a singular model.
#
# The keyword in the sentence above is **uncorrelated**, which is another way of saying, different types of models. For example, in our case, we might combine our baseline, our bidirectional model and our TensorFlow Hub USE model.
#
# Although these models are all trained on the same data, they all have a different way of finding patterns.
#
# If we were to use three similarly trained models, such as three LSTM models, the predictions they output will likely be very similar.
#
# Think of it as trying to decide where to eat with your friends. If you all have similar tastes, you'll probably all pick the same restaurant. But if you've all got different tastes and still end up picking the same restaurant, the restaurant must be good.
#
# Since we're working with a classification problem, there are a few of ways we can combine our models:
# 1. **Averaging** - Take the output prediction probabilities of each model for each sample, combine them and then average them.
# 2. **Majority vote (mode)** - Make class predictions with each of your models on all samples, the predicted class is the one in majority. For example, if three different models predict `[1, 0, 1]` respectively, the majority class is `1`, therefore, that would be the predicted label.
# 3. **Model stacking** - Take the outputs of each of your chosen models and use them as inputs to another model.
#
# > 📖 **Resource:** The above methods for model stacking/ensembling were adapted from Chapter 6 of the [Machine Learning Engineering Book](http://www.mlebook.com/wiki/doku.php) by <NAME>. If you're looking to enter the field of machine learning engineering, not only building models but production-scale machine learning systems, I'd highly recommend reading it in its entirety.
#
# Again, the concept of model stacking is best seen in action.
#
# We're going to combine our baseline model (`model_0`), LSTM model (`model_2`) and our USE model trained on the full training data (`model_6`) by averaging the combined prediction probabilities of each.
# + id="t63u8PCCm-yo" colab={"base_uri": "https://localhost:8080/"} outputId="47a65c8c-325a-402c-bfa3-0a940416bd54"
# Get mean pred probs for 3 models
baseline_pred_probs = np.max(model_0.predict_proba(val_sentences), axis=1) # get the prediction probabilities from baseline model
combined_pred_probs = baseline_pred_probs + tf.squeeze(model_2_pred_probs, axis=1) + tf.squeeze(model_6_pred_probs)
combined_preds = tf.round(combined_pred_probs/3) # average and round the prediction probabilities to get prediction classes
combined_preds[:20]
# + [markdown] id="6abZa7wqlXSI"
# Wonderful! We've got a combined predictions array of different classes, let's evaluate them against the true labels and add our stacked model's results to our `all_model_results` DataFrame.
# + id="ieYvhDiev8Et" colab={"base_uri": "https://localhost:8080/"} outputId="b3b965dd-54c8-4796-edcc-b31e344b38bb"
# Calculate results from averaging the prediction probabilities
ensemble_results = calculate_results(val_labels, combined_preds)
ensemble_results
# + id="132EHlUUpRrP"
# Add our combined model's results to the results DataFrame
all_model_results.loc["ensemble_results"] = ensemble_results
# + id="Pm2P1zsvpZ3D"
# Convert the accuracy to the same scale as the rest of the results
all_model_results.loc["ensemble_results"]["accuracy"] = all_model_results.loc["ensemble_results"]["accuracy"]/100
# + id="trmdZ6eEpwHI" colab={"base_uri": "https://localhost:8080/", "height": 328} outputId="b2704163-e293-4b88-f289-efcc04d9b4b2"
all_model_results
# + [markdown] id="HZwqwF_swdIA"
# How did the stacked model go against the other models?
#
# > 🔑 **Note:** It seems many of our model's results are similar. This may mean there are some limitations to what can be learned from our data. When many of your modelling experiments return similar results, it's a good idea to revisit your data, we'll do this shortly.
# + [markdown] id="UpwErZOgX_nC"
# ## Saving and loading a trained model
#
# Although training time didn't take very long, it's good practice to save your trained models to avoid having to retrain them.
#
# Saving your models also enables you to export them for use elsewhere outside of your notebooks, such as in a web application.
#
# There are two main ways of [saving a model in TensorFlow](https://www.tensorflow.org/tutorials/keras/save_and_load#save_the_entire_model):
# 1. The `HDF5` format.
# 2. The `SavedModel` format (default).
#
# Let's take a look at both.
# + id="SlwjGFVyX-_T"
# Save TF Hub Sentence Encoder model to HDF5 format
model_6.save("model_6.h5")
# + [markdown] id="Cp6zvmprm9A3"
# If you save a model as a `HDF5`, when loading it back in, you need to let [TensorFlow know about any custom objects you've used](https://www.tensorflow.org/tutorials/keras/save_and_load#saving_custom_objects) (e.g. components which aren't built from pure TensorFlow, such as TensorFlow Hub components).
# + id="sSINZ0Q-nRb2"
# Load model with custom Hub Layer (required with HDF5 format)
loaded_model_6 = tf.keras.models.load_model("model_6.h5",
custom_objects={"KerasLayer": hub.KerasLayer})
# + id="G4BCJ8iXnZ4r" colab={"base_uri": "https://localhost:8080/"} outputId="4d3a7c0e-bea8-4f99-923d-943213cd72cd"
# How does our loaded model perform?
loaded_model_6.evaluate(val_sentences, val_labels)
# + [markdown] id="02rbT4fwn0It"
# Calling the `save()` method on our target model and passing it a filepath allows us to save our model in the `SavedModel` format.
# + id="e3eVaNBDoMsv" colab={"base_uri": "https://localhost:8080/"} outputId="a55b3bd1-ac2d-45c1-90f2-4b021c9368f9"
# Save TF Hub Sentence Encoder model to SavedModel format (default)
model_6.save("model_6_SavedModel_format")
# + [markdown] id="l-t01S-JoOqK"
# If you use SavedModel format (default), you can reload your model without specifying custom objects using the [`tensorflow.keras.models.load_model()`](https://www.tensorflow.org/tutorials/keras/save_and_load) function.
# + id="Dw3zf4fVoU5H"
# Load TF Hub Sentence Encoder SavedModel
loaded_model_6_SavedModel = tf.keras.models.load_model("model_6_SavedModel_format")
# + id="IqiPr6iiofi1" colab={"base_uri": "https://localhost:8080/"} outputId="d4395889-dca0-4061-ba22-1723001c5c4e"
# Evaluate loaded SavedModel format
loaded_model_6_SavedModel.evaluate(val_sentences, val_labels)
# + [markdown] id="xzp3SHi3oQ3u"
# As you can see saving and loading our model with either format results in the same performance.
#
# > 🤔 **Question:** Should you used the `SavedModel` format or `HDF5` format?
#
# For most use cases, the `SavedModel` format will suffice. However, this is a TensorFlow specific standard. If you need a more general-purpose data standard, `HDF5` might be better. For more, check out the [TensorFlow documentation on saving and loading models](https://www.tensorflow.org/tutorials/keras/save_and_load).
# + [markdown] id="V5a1648rG3z1"
# ## Finding the most wrong examples
#
# We mentioned before that if many of our modelling experiments are returning similar results, despite using different kinds of models, it's a good idea to return to the data and inspect why this might be.
#
# One of the best ways to inspect your data is to sort your model's predictions and find the samples it got *most* wrong, meaning, what predictions had a high prediction probability but turned out to be wrong.
#
# Once again, visualization is your friend. Visualize, visualize, visualize.
#
# To make things visual, let's take our best performing model's prediction probabilities and classes along with the validation samples (text and ground truth labels) and combine them in a pandas DataFrame.
#
# * If our best model still isn't perfect, what examples is it getting wrong?
# * Which ones are the *most* wrong?
# * Are there some labels which are wrong? E.g. the model gets it right but the ground truth label doesn't reflect this
# + id="gnHfX--TwMIW" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="d060505e-21c7-42b7-e664-8c517f92d425"
# Create dataframe with validation sentences and best performing model predictions
val_df = pd.DataFrame({"text": val_sentences,
"target": val_labels,
"pred": model_6_preds,
"pred_prob": tf.squeeze(model_6_pred_probs)})
val_df.head()
# + [markdown] id="SKJ9dTbPrIG4"
# Oh yeah! Now let's find our model's wrong predictions (where `target != pred`) and sort them by their prediction probability (the `pred_prob` column).
# + id="0DwBXQS1wvZx" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="7d37fad0-db94-471e-fef1-3859b48c58f3"
# Find the wrong predictions and sort by prediction probabilities
most_wrong = val_df[val_df["target"] != val_df["pred"]].sort_values("pred_prob", ascending=False)
most_wrong[:10]
# + [markdown] id="r3VcRHOusB2D"
# Finally, we can write some code to visualize the sample text, truth label, prediction class and prediction probability. Because we've sorted our samples by prediction probability, viewing samples from the head of our `most_wrong` DataFrame will show us false positives.
#
# A reminder:
# * `0` = Not a real diaster Tweet
# * `1` = Real diaster Tweet
# + id="xLFYDEsoxRFP" colab={"base_uri": "https://localhost:8080/"} outputId="28d66210-3007-4661-e8c2-655d621f0e90"
# Check the false positives (model predicted 1 when should've been 0)
for row in most_wrong[:10].itertuples(): # loop through the top 10 rows (change the index to view different rows)
_, text, target, pred, prob = row
print(f"Target: {target}, Pred: {int(pred)}, Prob: {prob}")
print(f"Text:\n{text}\n")
print("----\n")
# + [markdown] id="aXCH9J-UspWg"
# We can view the bottom end of our `most_wrong` DataFrame to inspect false negatives (model predicts 0, not a real diaster Tweet, when it should've predicted 1, real diaster Tweet).
# + id="6EaMchehxwLq" colab={"base_uri": "https://localhost:8080/"} outputId="4c5fdb89-38b2-405a-f5d5-f816b83d3c79"
# Check the most wrong false negatives (model predicted 0 when should've predict 1)
for row in most_wrong[-10:].itertuples():
_, text, target, pred, prob = row
print(f"Target: {target}, Pred: {int(pred)}, Prob: {prob}")
print(f"Text:\n{text}\n")
print("----\n")
# + [markdown] id="lRKQPEAgtpJq"
# Do you notice anything interesting about the most wrong samples?
#
# Are the ground truth labels correct? What do you think would happen if we went back and corrected the labels which aren't?
# + [markdown] id="U0W3DWgWJCWs"
# ## Making predictions on the test dataset
#
# Alright we've seen how our model's perform on the validation set.
#
# But how about the test dataset?
#
# We don't have labels for the test dataset so we're going to have to make some predictions and inspect them for ourselves.
#
# Let's write some code to make predictions on random samples from the test dataset and visualize them.
# + id="6Q9lgqoDyequ" colab={"base_uri": "https://localhost:8080/"} outputId="f81e856d-8024-4471-ee0d-3c14eb128cb0"
# Making predictions on the test dataset
test_sentences = test_df["text"].to_list()
test_samples = random.sample(test_sentences, 10)
for test_sample in test_samples:
pred_prob = tf.squeeze(model_6.predict([test_sample])) # has to be list
pred = tf.round(pred_prob)
print(f"Pred: {int(pred)}, Prob: {pred_prob}")
print(f"Text:\n{test_sample}\n")
print("----\n")
# + [markdown] id="QcvI5zgJ0Tgp"
# How do our model's predictions look on the test dataset?
#
# It's important to do these kind of visualization checks as often as possible to get a glance of how your model performs on unseen data and subsequently how it might perform on the real test: Tweets from the wild.
# + [markdown] id="eT1jhk8xdod5"
# ## Predicting on Tweets from the wild
#
# How about we find some Tweets and use our model to predict whether or not they're about a diaster or not?
#
# To start, let's take one of my own [Tweets on living life like an ensemble model](https://twitter.com/mrdbourke/status/1313649328351662082).
# + id="qHmXxuPH0aUB"
# Turn Tweet into string
daniels_tweet = "Life like an ensemble: take the best choices from others and make your own"
# + [markdown] id="uPbZaGznvbEx"
# Now we'll write a small function to take a model and an example sentence and return a prediction.
# + id="KyH9tn9upjld"
def predict_on_sentence(model, sentence):
"""
Uses model to make a prediction on sentence.
Returns the sentence, the predicted label and the prediction probability.
"""
pred_prob = model.predict([sentence])
pred_label = tf.squeeze(tf.round(pred_prob)).numpy()
print(f"Pred: {pred_label}", "(real disaster)" if pred_label > 0 else "(not real disaster)", f"Prob: {pred_prob[0][0]}")
print(f"Text:\n{sentence}")
# + [markdown] id="IvCG4RuUvj6d"
# Great! Time to test our model out.
# + id="BxONpJV8qmWP" colab={"base_uri": "https://localhost:8080/"} outputId="ddd8f755-089c-4d6b-a2be-890644d10199"
# Make a prediction on Tweet from the wild
predict_on_sentence(model=model_6, # use the USE model
sentence=daniels_tweet)
# + [markdown] id="tYOfNacw08Of"
# Woohoo! Our model predicted correctly. My Tweet wasn't about a diaster.
#
# How about we find a few Tweets about actual diasters?
#
# Such as the following two Tweets about the 2020 Beirut explosions.
# + id="AqILBsTK2i9R"
# Source - https://twitter.com/BeirutCityGuide/status/1290696551376007168
beirut_tweet_1 = "Reports that the smoke in Beirut sky contains nitric acid, which is toxic. Please share and refrain from stepping outside unless urgent. #Lebanon"
# Source - https://twitter.com/BeirutCityGuide/status/1290773498743476224
beirut_tweet_2 = "#Beirut declared a “devastated city”, two-week state of emergency officially declared. #Lebanon"
# + id="FvlbHDISrVmX" colab={"base_uri": "https://localhost:8080/"} outputId="6be0b58b-4ac6-4948-cd1e-1d43f4ddb159"
# Predict on diaster Tweet 1
predict_on_sentence(model=model_6,
sentence=beirut_tweet_1)
# + id="5uKYx11p2zCd" colab={"base_uri": "https://localhost:8080/"} outputId="6b4b7408-f56d-49ed-ff41-584dfe65cf1b"
# Predict on diaster Tweet 2
predict_on_sentence(model=model_6,
sentence=beirut_tweet_2)
# + [markdown] id="fczP1dFcwe98"
# Looks like our model is performing as expected, predicting both of the diaster Tweets as actual diasters.
#
# > 🔑 **Note:** The above examples are cherry-picked and are cases where you'd expect a model to function at high performance. For actual production systems, you'll want to continaully perform tests to see how your model is performing.
# + [markdown] id="Fp0fkK-tHPRE"
# ## The speed/score tradeoff
#
# One of the final tests we're going to do is to find the speed/score tradeoffs between our best model and baseline model.
#
# Why is this important?
#
# Although it can be tempting to just choose the best performing model you find through experimentation, this model might not actually work in a production setting.
#
# Put it this way, imagine you're Twitter and receive 1 million Tweets per hour (this is a made up number, the actual number is much higher). And you're trying to build a diaster detection system to read Tweets and alert authorities with details about a diaster in close to real-time.
#
# Compute power isn't free so you're limited to a single compute machine for the project. On that machine, one of your models makes 10,000 predictions per second at 80% accuracy where as another one of your models (a larger model) makes 100 predictions per second at 85% accuracy.
#
# Which model do you choose?
#
# Is the second model's performance boost worth missing out on the extra capacity?
#
# Of course, there are many options you could try here, such as sending as many Tweets as possible to the first model and then sending the ones which the model is least certain of to the second model.
#
# The point here is to illustrate the best model you find through experimentation, might not be the model you end up using in production.
#
# To make this more concrete, let's write a function to take a model and a number of samples and time how long the given model takes to make predictions on those samples.
# + id="DnXp8DKOp3J6"
# Calculate the time of predictions
import time
def pred_timer(model, samples):
"""
Times how long a model takes to make predictions on samples.
Args:
----
model = a trained model
sample = a list of samples
Returns:
----
total_time = total elapsed time for model to make predictions on samples
time_per_pred = time in seconds per single sample
"""
start_time = time.perf_counter() # get start time
model.predict(samples) # make predictions
end_time = time.perf_counter() # get finish time
total_time = end_time-start_time # calculate how long predictions took to make
time_per_pred = total_time/len(val_sentences) # find prediction time per sample
return total_time, time_per_pred
# + [markdown] id="GxWwS73hze6Z"
# Looking good!
#
# Now let's use our `pred_timer()` function to evaluate the prediction times of our best performing model (`model_6`) and our baseline model (`model_0`).
# + id="JMbGMIWd5c9N" colab={"base_uri": "https://localhost:8080/"} outputId="4a151071-956a-4c97-fd95-18392a57a78b"
# Calculate TF Hub Sentence Encoder prediction times
model_6_total_pred_time, model_6_time_per_pred = pred_timer(model_6, val_sentences)
model_6_total_pred_time, model_6_time_per_pred
# + id="I4ej2VyT5oQs" colab={"base_uri": "https://localhost:8080/"} outputId="cfbedfa1-282e-4b04-b2c4-1a3b33771b1e"
# Calculate Naive Bayes prediction times
baseline_total_pred_time, baseline_time_per_pred = pred_timer(model_0, val_sentences)
baseline_total_pred_time, baseline_time_per_pred
# + [markdown] id="nqNnKMxhz8Kl"
# It seems with our current hardware (in my case, I'm using a Google Colab notebook) our best performing model takes over 10x the time to make predictions as our baseline model.
#
# Is that extra prediction time worth it?
#
# Let's compare time per prediction versus our model's F1-scores.
# + id="ANKHEfRN7Nhd" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="a4ca1fff-0b84-41d0-85fd-b7f258d47558"
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 7))
plt.scatter(baseline_time_per_pred, baseline_results["f1"], label="baseline")
plt.scatter(model_6_time_per_pred, model_6_results["f1"], label="tf_hub_sentence_encoder")
plt.legend()
plt.title("F1-score versus time per prediction")
plt.xlabel("Time per prediction")
plt.ylabel("F1-Score");
# + [markdown] id="QlHdTqTl0aOq"
# 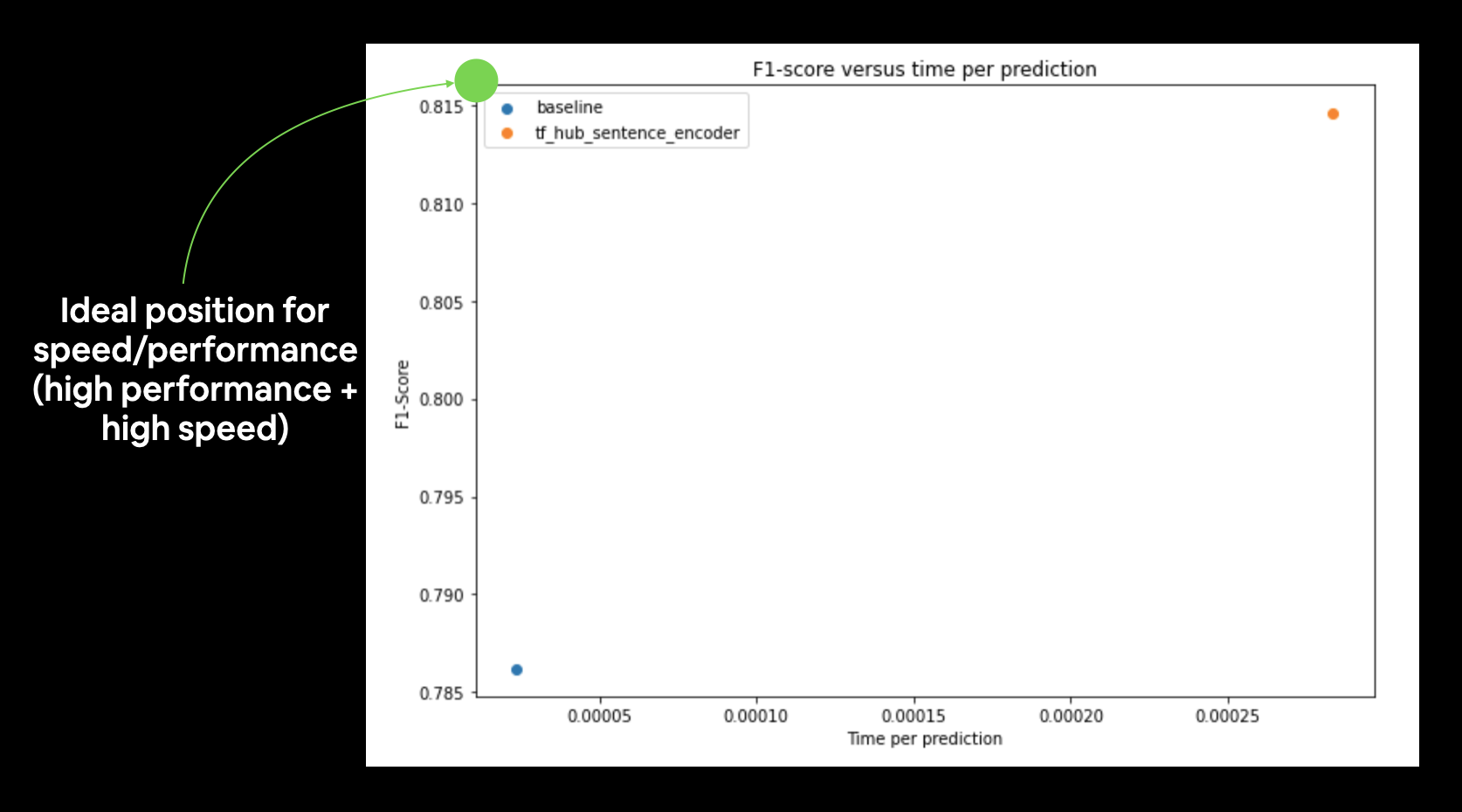
# *Ideal position for speed and performance tradeoff model (fast predictions with great results).*
#
# Of course, the ideal position for each of these dots is to be in the top left of the plot (low time per prediction, high F1-score).
#
# In our case, there's a clear tradeoff for time per prediction and performance. Our best performing model takes an order of magnitude longer per prediction but only results in a few F1-score point increase.
#
# This kind of tradeoff is something you'll need to keep in mind when incorporating machine learning models into your own applications.
# + [markdown] id="DJWGI6GpH4Gl"
# ##Exercises
#
# 1. Rebuild, compile and train `model_1`, `model_2` and `model_5` using the [Keras Sequential API](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) instead of the Functional API.
# 2. Retrain the baseline model with 10% of the training data. How does perform compared to the Universal Sentence Encoder model with 10% of the training data?
# 3. Try fine-tuning the TF Hub Universal Sentence Encoder model by setting `training=True` when instantiating it as a Keras layer.
#
# ```
# We can use this encoding layer in place of our text_vectorizer and embedding layer
#
# sentence_encoder_layer = hub.KerasLayer("https://tfhub.dev/google/universal-sentence-encoder/4",
# input_shape=[],
# dtype=tf.string,
# trainable=True) # turn training on to fine-tune the TensorFlow Hub model
# ```
# 4. Retrain the best model you've got so far on the whole training set (no validation split). Then use this trained model to make predictions on the test dataset and format the predictions into the same format as the `sample_submission.csv` file from Kaggle (see the Files tab in Colab for what the `sample_submission.csv` file looks like). Once you've done this, [make a submission to the Kaggle competition](https://www.kaggle.com/c/nlp-getting-started/data), how did your model perform?
# 5. Combine the ensemble predictions using the majority vote (mode), how does this perform compare to averaging the prediction probabilities of each model?
# 6. Make a confusion matrix with the best performing model's predictions on the validation set and the validation ground truth labels.
# + [markdown] id="BarVJji8H6M4"
# ##Extra-curriculum
#
# To practice what you've learned, a good idea would be to spend an hour on 3 of the following (3-hours total, you could through them all if you want) and then write a blog post about what you've learned.
#
# * For an overview of the different problems within NLP and how to solve them read through:
# * [A Simple Introduction to Natural Language Processing](https://becominghuman.ai/a-simple-introduction-to-natural-language-processing-ea66a1747b32)
# * [How to solve 90% of NLP problems: a step-by-step guide](https://blog.insightdatascience.com/how-to-solve-90-of-nlp-problems-a-step-by-step-guide-fda605278e4e)
# * Go through [MIT's Recurrent Neural Networks lecture](https://youtu.be/SEnXr6v2ifU). This will be one of the greatest additions to what's happening behind the RNN model's you've been building.
# * Read through the [word embeddings page on the TensorFlow website](https://www.tensorflow.org/tutorials/text/word_embeddings). Embeddings are such a large part of NLP. We've covered them throughout this notebook but extra practice would be well worth it. A good exercise would be to write out all the code in the guide in a new notebook.
# * For more on RNN's in TensorFlow, read and reproduce [the TensorFlow RNN guide](https://www.tensorflow.org/guide/keras/rnn). We've covered many of the concepts in this guide, but it's worth writing the code again for yourself.
# * Text data doesn't always come in a nice package like the data we've downloaded. So if you're after more on preparing different text sources for being with your TensorFlow deep learning models, it's worth checking out the following:
# * [TensorFlow text loading tutorial](https://www.tensorflow.org/tutorials/load_data/text).
# * [Reading text files with Python](https://realpython.com/read-write-files-python/) by Real Python.
# * This notebook has focused on writing NLP code. For a mathematically rich overview of how NLP with Deep Learning happens, read [Standford's Natural Language Processing with Deep Learning lecture notes Part 1](https://web.stanford.edu/class/cs224n/readings/cs224n-2019-notes01-wordvecs1.pdf).
# * For an even deeper dive, you could even do the whole [CS224n](http://web.stanford.edu/class/cs224n/) (Natural Language Processing with Deep Learning) course.
# * Great blog posts to read:
# * <NAME>'s [The Unreasonable Effectiveness of RNNs](https://karpathy.github.io/2015/05/21/rnn-effectiveness/) dives into generating Shakespeare text with RNNs.
# * [Text Classification with NLP: Tf-Idf vs Word2Vec vs BERT](https://towardsdatascience.com/text-classification-with-nlp-tf-idf-vs-word2vec-vs-bert-41ff868d1794) by <NAME>. An overview of different techniques for turning text into numbers and then classifying it.
# * [What are word embeddings?](https://machinelearningmastery.com/what-are-word-embeddings/) by Machine Learning Mastery.
# * Other topics worth looking into:
# * [Attention mechanisms](https://jalammar.github.io/visualizing-neural-machine-translation-mechanics-of-seq2seq-models-with-attention/). These are a foundational component of the transformer architecture and also often add improvments to deep NLP models.
# * [Transformer architectures](http://jalammar.github.io/illustrated-transformer/). This model architecture has recently taken the NLP world by storm, achieving state of the art on many benchmarks. However, it does take a little more processing to get off the ground, the [HuggingFace Models (formerly HuggingFace Transformers) library](https://huggingface.co/models/) is probably your best quick start.
# * And now [HuggingFace even have their own course](https://huggingface.co/course/chapter1) on how their library works! I haven't done it but anything HuggingFace makes is world-class.
#
#
# + [markdown] id="CLzfxgXkzEdr"
# > 📖 **Resource:** See the full set of course materials on GitHub: https://github.com/mrdbourke/tensorflow-deep-learning
| 08_introduction_to_nlp_in_tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="E3-D41ASugJQ"
# # 8 Queens Problem - Solving using DWave Constrained Quadratic Model
# ## **Special version with a classical simulation BQM option**
#
# ###### By <NAME>
#
# ## Problem
#
# Problem: On a standard 8 X 8 chess board, place 8 queens, anywhere on the board, such that no two queens are in conflict with each other (i.e. no queen can take any other queen using standard chess movement rules for a Queen piece).
#
# If you have not tried to solve this problem manually on a chess board, you will find that it is a difficult problem to solve.
#
# ## CQM vs BQM
#
# This solution was originally created as a BQM (Binary Quadratic Model) as a QUBO. The CQM model abstracts solving the quadratic mathematics required to implement constraints in a QUBO by allowing us to express constraints more naturally with the quadratic equations implemented under the hood.
#
# The code and efforts required to implement a solver using CQM is minimum and demonstrated below compared to creating it as a QUBO.
#
# ## Model
#
# 8 X 8 binary variables are defined. Each variable is associated to one of the squares on the board. Each indicate if its suqare is occupied (=1) or not (=0) by a Queen chess piece.
#
# ## Constraints: Letting the Quantum Computer solve the problem
#
# The constraints tell the DWave quantum annealer what is valid and not valid without hinting as to how to solve the problem.
#
# (e.g. Although we "know" that solving the problem means that you can only put one queen on any row, we will allow the computer to consider 0 as a possibility and let it figure out the solution).
#
# >1 - There must be exactly NQ (=8 by default) queens on the board.
# >> The sum of all the binary variables must be == NQ
#
# >2 - No two queens can occupy the same row on the board
# >> The sum of all binary variables in any given row must be <= 1
#
# >3 - No two queens can occupy the same column on the board
# >> The sum of all binary variables in any given column must be <= 1
#
# >4 - No two queens can occupy the same diagonal on the board
# >> The sum of all binary variables in any given diagonal must be <= 1
#
# ## Variations
#
# Two parameters can be changed:
#
# NQ = Number of Queens
#
# N = Number of squares on a side of the square board
#
# One can increase the problem to placing 32 queens on a 32 x 32 board. Some variations may lead to "No feasible" solutions.
#
# We challenge one to find a real life solution to a problem setup for which the quantum computer fails at finding a solution.
#
# ## Usage
#
# First assign your Leap API token below and run the notebook. The first time you run the notebook, the Ocean SDK will get installed in the VM.
#
# Results are presented at the end in the form of a board. Up to 10 variations are presented.
#
# ## CQM_to_BQM method - Allow simulation on classical
#
# To allow for running the model on classical hardware and save on QPU cost, choose
# >method = m_BQM
#
# This allow running the Neal Simulated Annealing method in the event the QPU is not available or accessible.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="wefv8lX30u_W" outputId="d6d9cbb8-9322-4724-9092-2e022aa68fde"
# !pip install dwave-ocean-sdk
# + id="<KEY>"
import dimod
import neal
from dwave.system import LeapHybridCQMSampler, LeapHybridBQMSampler
import numpy as np
# + id="DYqEIK8PSfJC"
token=None #'<KEY>'
# Ability to convert CQM to a BQM
# so that we can use simulated annealing to solver the puzzle
# (CQM does not appear to have a Simulation mode)
#
m_CQM = 1 # Method for which the model is designed for
m_BQM = 2 # Alternative method converting CQM to BQM to allow using Neal simulation
# Default is BQM to allow running the model without a QPU
# If you have an API token to use the Quantum computer, change the method to m_CQM
method = m_BQM # <======= change this to m_CQM run on Quantum computer
# Options for using a BQM version of the CQM
bqm = None
invert = None
Lagrange = 1000
neal_reads = 1000
# + id="wRaRoX_v0_nT" colab={"base_uri": "https://localhost:8080/"} outputId="d8959ae7-6e37-4f02-c571-9782f0f5e6f1"
# Create a CQM to solve the 8 Queens Problem
NQ = 8 # Number of Queens to place: Objective
N = 8 # Number of squares across and down on the board
time_limit = 5 # Hybrid Sampler Time Limit
problabel = "NQueensCQM" # Label for the leap dashboard
# Define a blank cqm model
cqm = dimod.CQM()
# Function to reate a Variable name for a given square position on the chess board
def varname( r=1, c=1 ):
return "square_r"+str(r)+"_c"+str(c)
# Lists of variable numbers and their label
vlabels = [varname(r,c) for r in range(N) for c in range(N)]
vnum = np.array([[ r*N+c for c in range(N)] for r in range(N)])
rvnum = vnum[:, ::-1]
# Create board of binary variables for the CQM model
variables = [ dimod.Binary(varname(r,c)) for r in range(N) for c in range(N)]
board = { (r,c) : variables[r*N+c] for c in range(N) for r in range(N) }
# Create lists of variables on all diagonals of more than 1 square
diags = []
for a in [vnum,rvnum]:
for k in range(-(N-2),N-1):
diags.append(np.diag(a,k=k).tolist())
#print( np.diag(vnum,k=k))
# Model verification
verify = True
if ( verify == True ):
print("Diagonals:", diags)
print("Variable Labels:",vlabels)
print("Regular board Variable numbers:\n", vnum)
print("Reverse board variable numbers:\n", rvnum)
print("CQM Variables assigned to the board:\n",board)
# + id="NBMoghRy1Ec8"
# Objective : There is no objective function
# Although one may consider placing NQ queens on the board to be the objective,
# we preferred to implement it as a constraint instead
# Constraints
# 1 - Must have 8 queens on board
cqm.add_constraint( sum(board[r,c] for r in range(N) for c in range(N)) == NQ, label="board_total" )
# 2 - Each column can have at most one queen
for c in range(N):
cqm.add_constraint( sum( board[r,c] for r in range(N) ) <= 1, label=f"col_{c}_total" )
# 3 - Each row can have at most one queen
for r in range(N):
cqm.add_constraint( sum( board[r,c] for c in range(N) ) <= 1, label=f"row_{r}_total" )
# 4 - Each diagonal can have at most one queen
for i,d in enumerate(diags):
cqm.add_constraint( sum( variables[v] for v in d ) <= 1, label=f"diag_{i}_total" )
# + id="ieCQEY-wOSWn" colab={"base_uri": "https://localhost:8080/"} outputId="7525a9ed-7d67-41f0-93dd-fba08775a9be"
if ( method == m_CQM ):
# Call the solver and obtain results
print("Running CQM on LeapHybridCQMSampler...")
sampler = LeapHybridCQMSampler(token=token)
raw_sampleset = sampler.sample_cqm(cqm, time_limit=time_limit,label=problabel)
elif ( method == m_BQM ):
print("Running CQM as a BQM on SimulatedAnnealingSampler...")
bqm, invert = dimod.cqm_to_bqm(cqm,Lagrange)
#sampleset = dimod.ExactSolver().sample(bqm)
#bqm = BinaryQuadraticModel.from_qubo(self.Q, offset=self.offset)
sampler = neal.SimulatedAnnealingSampler()
raw_sampleset = sampler.sample(bqm, num_reads = neal_reads)
pass
else:
print( "Solver method unknown")
raise SystemExit(0)
# + id="Zv9hOg6ga3Uw"
if ( method == m_CQM ):
feasible_sampleset = raw_sampleset.filter(lambda d: d.is_feasible)
num_feasible = len(feasible_sampleset)
print(str(num_feasible)+" Feasible samples")
if num_feasible > 0:
best_samples = \
feasible_sampleset.truncate(min(10, num_feasible))
else:
print("Warning: Did not find feasible solution")
best_samples = raw_sampleset.truncate(10)
print(" \n" + "=" * 30 + "BEST SAMPLE SET" + "=" * 30)
print(best_samples)
#for s in best_samples:
# print(s)
# + colab={"base_uri": "https://localhost:8080/"} id="JM3bXH3XTh7Q" outputId="7bfaffdc-1e31-435a-e25e-64a2ef8f58cb"
# BQM Method
if ( method == m_BQM ):
feasible_sampleset = raw_sampleset
num_feasible = len(feasible_sampleset)
print(str(num_feasible)+" Feasible samples")
if num_feasible > 0:
best_samples = \
feasible_sampleset.truncate(min(10, num_feasible))
else:
print("Warning: Did not find feasible solution")
best_samples = raw_sampleset.truncate(10)
print(" \n" + "=" * 30 + "BEST SAMPLE SET" + "=" * 30)
print(best_samples)
#for s in best_samples:
# print(s)
# + colab={"base_uri": "https://localhost:8080/"} id="xvlwxyC14CvW" outputId="5f42a4b1-07e5-499c-e064-af3ffc9fa489"
for i,s in enumerate(best_samples):
if ( method == m_BQM ):
s = invert(s)
print("Result "+str(i+1)+" =====================")
for r in range(N):
for c in range(N):
v = s[varname(r,c)]
q = '.'
if ( v > 0.0 ): q = 'W'
print( '|.'+ q + '.', end='' )
print( '|')
print("=================================")
# + id="YUvrrs2cRuSs"
| 8Queens/8Queens_CQM2BQM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Field, Goldsmith, & Habing Multi-Phase ISM
#
# Figure 1.10-1.12 from Chapter 1 of *Interstellar and Intergalactic Medium* by <NAME>, 2021,
# Cambridge University Press.
#
# This notebook creates figures illustrating the Field, Goldsmith, and Habing (FGH) multi-phase interstellar
# medium model [Field, Goldsmith, & Habing 1969, ApJ, 155, L149](https://ui.adsabs.harvard.edu/abs/1969ApJ...155L.149F/abstract)
#
# There are 3 figures
# * Figure 1.10 - FGH Cooling function $\Lambda(T)$
# * Figure 1.11 - Equilibrium density $n_{eq}(T)$
# * Figure 1.12 - Pressure $P$ vs density $n_{eq}$
# +
# %matplotlib inline
import math
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
# SciPy bits we use for analysis
from scipy.signal import argrelmin, argrelmax
from scipy import stats
import warnings
warnings.filterwarnings('ignore',category=UserWarning, append=True)
# -
# ## Standard Plot Format
#
# Setup the standard plotting format and make the plot. Fonts and resolution adopted follow CUP style.
# +
# graphic aspect ratio = width/height
aspect = 4.0/3.0
# Text width in inches - don't change, this is defined by the print layout
textWidth = 6.0 # inches
# output format and resolution
figFmt = 'png'
dpi = 600
# Graphic dimensions
plotWidth = dpi*textWidth
plotHeight = plotWidth/aspect
axisFontSize = 14
labelFontSize = 10
lwidth = 0.5
axisPad = 5
wInches = textWidth
hInches = wInches/aspect
# LaTeX is used throughout for markup of symbols, Times-Roman serif font
plt.rc('text', usetex=True)
plt.rc('font', **{'family':'serif','serif':['Times-Roman'],'weight':'bold','size':'16'})
# Font and line weight defaults for axes
matplotlib.rc('axes',linewidth=lwidth)
matplotlib.rcParams.update({'font.size':axisFontSize})
# axis and label padding
plt.rcParams['xtick.major.pad'] = f'{axisPad}'
plt.rcParams['ytick.major.pad'] = f'{axisPad}'
plt.rcParams['axes.labelpad'] = f'{axisPad}'
# -
# ## FGH Model Calculation
#
# The basic parameters for these models are set in the code section below:
#
# * Ionization fraction: $x_e$=0.001
# * Temperature Range: $T_e$=10 to 20000K (logratithmic)
# * gain factor: $G$=20, defined such that $n_{eq}$=G/$\Lambda$ ($\Lambda$ is the total cooling rate)
#
# The model assumes three sources of collisional cooling using these scaling relations:
#
# HI Lyman-$\alpha$ Cooling (Eqn 1.38):
# \begin{equation}
# \frac{\Lambda^{e}_{Ly\alpha}}{10^{-27}{\rm erg\,cm^3\,s^{-1}}} \approx
# 6\times10^{5} \left(\frac{x}{0.001}\right)
# \left(\frac{T}{10^{4}K}\right)^{-1/2}
# \exp\left(-\frac{1.18\times10^{5}K}{T}\right)
# \end{equation}
#
# Carbon (CII) Cooling (Eqn 1.35) electron collisional term:
# \begin{equation}
# \frac{\Lambda^{e}_{CII}}{10^{-27}{\rm erg\,cm^3\,s^{-1}}} \approx
# 3.1 \left(\frac{x}{0.001}\right)
# \left(\frac{T}{100K}\right)^{-0.5}
# \exp\left(-\frac{91.2K}{T}\right)
# \end{equation}
# and H collisional term:
# \begin{equation}
# \frac{\Lambda^{H}_{CII}}{10^{-27}{\rm erg\,cm^3\,s^{-1}}} \approx
# 5.2\left(\frac{T}{100K}\right)^{0.13}
# \exp\left(-\frac{91.2K}{T}\right)
# \end{equation}
#
# Oxygen (OI) Cooling:
# \begin{equation}
# \frac{\Lambda^{H}_{OI}}{10^{-27}{\rm erg\,cm^3\,s^{-1}}} \approx
# 4.1\left(\frac{T}{100K}\right)^{0.42}
# \exp\left(-\frac{228K}{T}\right)
# \end{equation}
#
# We compute total cooling ($\Lambda=\Lambda_{Ly\alpha}+\Lambda_{CII}+\Lambda_{OII}$), equilibrium density
# ($n_{eq}$), and pressure ($P=n_{eq}kT$) as a function of logarithmic steps in temperature.
#
# We have adopted the Lodders (2010) abundances for C and O, as used in the ISM/IGM book
# (see Chapter 1, Table 1.2).
# +
xe = 0.001
minT = 10.0
maxT = 20000.
gain = 20.0
# Boltzmann Constant (CODATA 2018)
k = 1.380649e-16 # erg K^-1
minLogT = math.log10(minT)
maxLogT = math.log10(maxT)
logT = np.linspace(minLogT,maxLogT,num=1001)
T = 10.0**logT
xfac = xe/0.001
TH = 118000.0 # hydrogen excitation temperature in K
TC = 91.2 # carbon excitation temperature in K
TO = 228.0 # oxygen excitation temperature in K
# Lyman-alpha cooling
coolLya = 6.0e5*(xfac/np.sqrt(T/1.0e4))*np.exp(-TH/T)
# Carbon cooling
coolC = 3.1*(xfac/np.sqrt(T/100.0))*np.exp(-TC/T) + 5.2*((T/100.0)**0.13)*np.exp(-TC/T)
# Oxygen cooling
coolO = 4.1*((T/100.0)**0.42)*np.exp(-TO/T)
# Total cooling
coolTot = (coolLya + coolC + coolO)
# equilibrium density
neq = gain/coolTot
# pressure
P = neq*k*T
# -
# ## FGH Cooling Function - Figure 1.10
#
# Plot the cooling function $\Lambda(T)$ vs $T$ including the curves for the individual contributions
#
# +
plotFile = f'Fig1_10.{figFmt}'
fig,ax = plt.subplots()
fig.set_dpi(dpi)
fig.set_size_inches(wInches,hInches,forward=True)
ax.tick_params('both',length=6,width=lwidth,which='major',direction='in',top='on',right='on')
ax.tick_params('both',length=3,width=lwidth,which='minor',direction='in',top='on',right='on')
# Limits
minCool = 1.0e-30 # erg cm^3 s^-1
maxCool = 1.0e-24
# Labels
xLabel = r'Temperature [K]'
yLabel = r'$\Lambda$ [erg cm$^3$ s$^{-1}$]'
plt.xlim(minT,maxT)
ax.set_xscale('log')
ax.set_xticks([10,100,1000,1.0e4])
ax.set_xticklabels(['10','100','1000','10$^{4}$'])
plt.xlabel(xLabel)
plt.ylim(minCool,maxCool)
ax.set_yscale('log')
ax.set_yticks([1.0E-30,1.0E-29,1.0E-28,1.0E-27,1.0e-26,1.0e-25,1.0e-24])
ax.set_yticklabels(['$10^{-30}$','10$^{-29}$','10$^{-28}$','10$^{-27}$','10$^{-26}$','10$^{-25}$','10$^{-24}$'])
plt.ylabel(yLabel)
# Plot the total and individual cooling functions
plt.plot(T,1.0e-27*coolTot,'-',color='black',lw=2,zorder=10)
plt.plot(T,1.0e-27*coolLya,'--',color='black',lw=1,zorder=10)
plt.plot(T,1.0e-27*coolC,':',color='black',lw=1,zorder=10)
plt.plot(T,1.0e-27*coolO,'-.',color='black',lw=1,zorder=10)
# label components
lfs = np.rint(1.2*axisFontSize)
plt.text(1000.0,1.7e-26,'Total',fontsize=lfs,rotation=10.0,ha='center',va='bottom')
plt.text(80.0,1.0e-28,r'$[\textsc{O\,i}]\,\lambda$63$\mu m$',fontsize=lfs)
plt.text(3000.0,3.5e-27,r'$[\textsc{C\,ii}]\,\lambda$158$\mu m$',fontsize=lfs,rotation=3.0,ha='center')
plt.text(5400.0,1.0e-28,r'Ly$\alpha$',fontsize=lfs,ha='center')
# make the figure
plt.plot()
plt.savefig(plotFile,bbox_inches='tight',facecolor='white')
# -
# ## FGH equilibrium density - Figure 1.11
#
# Plot the equlibrium density function $n_{eq}$ vs $T$ for the FGH model.
#
# +
plotFile = f'Fig1_11.{figFmt}'
fig,ax = plt.subplots()
fig.set_dpi(dpi)
fig.set_size_inches(wInches,hInches,forward=True)
ax.tick_params('both',length=6,width=lwidth,which='major',direction='in',top='on',right='on')
ax.tick_params('both',length=3,width=lwidth,which='minor',direction='in',top='on',right='on')
# Limits
minNe = 0.01 # cm^{-3}
maxNe = 20000.0
# Labels
xLabel = r'Temperature [K]'
yLabel = r'$n$ [cm$^{-3}$]'
plt.xlim(minT,maxT)
ax.set_xscale('log')
ax.set_xticks([10,100,1000,1.0e4])
ax.set_xticklabels(['10','100','1000','10$^{4}$'])
plt.xlabel(xLabel)
plt.ylim(minNe,maxNe)
ax.set_yscale('log')
ax.set_yticks([0.01,0.1,1.0,10.,100.,1e3,1e4])
ax.set_yticklabels(['0.01','0.1','1','10','100','1000','10$^{4}$'])
plt.ylabel(yLabel)
# Plot neq vs T
plt.plot(T,neq,'-',color='black',lw=2,zorder=10)
plt.fill_between(T,neq,maxNe,facecolor="#eaeaea")
# label regions above and below
lfs = np.rint(1.2*axisFontSize)
plt.text(200.0,0.1,'Net heating',fontsize=lfs,ha='center',zorder=10)
plt.text(1000.0,20.0,'Net cooling',fontsize=lfs,ha='center',zorder=10)
# make the figure
plt.plot()
plt.savefig(plotFile,bbox_inches='tight',facecolor='white')
# -
# ## FGH pressure vs density - Figure 1.12
#
# Plot the equlibrium pressure vs density for the FGH model.
#
# We numerically search for the stability region pressure limits and the crossing points at a reference pressure of
# P= 2×10−13 dyne/cm 2 . The methods used are a little dodgy, but are robust here as the pressure-density curve is
# well-behaved.
# +
plotFile = f'Fig1_12.{figFmt}'
fig,ax = plt.subplots()
fig.set_dpi(dpi)
fig.set_size_inches(wInches,hInches,forward=True)
plt.tick_params('both',length=6,width=lwidth,which='major',direction='in',top='on',right='on')
plt.tick_params('both',length=3,width=lwidth,which='minor',direction='in',top='on',right='on')
# Limits
minNe = 0.02 # cm^{-3}
maxNe = 10000.0
minP = 4.0e-14 # dyne cm^-2
maxP = 1.0e-11
# Labels
xLabel = r'$n$ [cm$^{-3}$]'
yLabel = r'$P$ [dyne cm$^{-2}$]'
plt.xlim(minNe,maxNe)
plt.xscale('log')
ax.set_xticks([0.1,1.0,10.,1.0e2,1.0e3,1.0e4])
ax.set_xticklabels(['0.1','1.0','10','100','1000','10$^4$'])
plt.xlabel(xLabel)
plt.ylim(minP,maxP)
ax.set_yscale('log')
ax.set_yticks([1.0e-13,1.0e-12,1.0e-11])
ax.set_yticklabels(['10$^{-13}$','10$^{-12}$','10$^{-11}$'])
plt.ylabel(yLabel)
# plot the n-P curve
plt.plot(neq,P,'-',color='black',lw=2,zorder=10)
plt.fill_between(neq,P,maxP,facecolor="#eaeaea")
# FGH stability region - estimate from array using scipy.signal argrelmin() and argrelmax()
# peak-finding functions
iMin = argrelmin(P)[0]
iMax = argrelmax(P)[0]
plt.hlines(P[iMin],minNe,maxNe,color='black',ls='--',lw=0.5)
plt.hlines(P[iMax],minNe,maxNe,color='black',ls='--',lw=0.5)
# Reference pressure, 2e-13 dyne/cm^2
pFGH = 2.0e-13
# The FGH points are at zero crossings of P(n)-fghP. Find the nearest zero-crossing, then
# fit a line to +/-3 points around it and find the crossing point. This is dodgy generally
# but we get away with it because the P-n curve is well-behaved.
iFGH = np.where(np.diff(np.sign(P-pFGH)))[0]
nFGH = []
for i in iFGH:
slope, inter, rVal, pVal, stdErr = stats.linregress(neq[i-3:i+3],P[i-3:i+3]-pFGH)
xZero = -inter/slope
nFGH.append(xZero)
# print(f'n_eq = {xZero:.5e} cm^-3')
lfs = np.rint(1.2*axisFontSize)
plt.plot(nFGH[0],pFGH,color='black',marker='o',ms=8,mfc='black')
plt.text(1.4*nFGH[0],pFGH,'F',fontsize=lfs,va='center',zorder=10)
plt.plot(nFGH[1],pFGH,color='black',marker='o',ms=8,mfc='black')
plt.text(1.4*nFGH[1],pFGH,'G',fontsize=lfs,va='center',zorder=10)
plt.plot(nFGH[2],pFGH,color='black',marker='o',ms=8,mfc='black')
plt.text(1.4*nFGH[2],pFGH,'H',fontsize=lfs,va='center',zorder=10)
plt.text(10.0,1.1*P[iMax],'Net cooling',fontsize=lfs,ha='center',va='bottom',zorder=10)
plt.text(1300.0,pFGH,'Net heating',fontsize=lfs,ha='center',va='center',zorder=10)
# make the figure
plt.plot()
plt.savefig(plotFile,bbox_inches='tight',facecolor='white')
| Chapter1/Fig1_FGH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
from data import Datafile, load_data
from influence.emp_risk_optimizer import EmpiricalRiskOptimizer
from influence.plot_utils import compare_with_loo
from influence.closed_forms import I_loss_RidgeCf
from models.regularized_regression import RegularizedRegression
from models.hyperplane_clf import BinaryLogisticRegression
# +
from sklearn.preprocessing import StandardScaler
X_train, X_test, y_train, y_test, test_indices = load_data(
Datafile.BinaryMNIST17, test_config=[0])
n_tr, p = X_train.shape
n_te, _ = X_test.shape
print(n_tr, p)
scl_x = StandardScaler()
scl_y = StandardScaler()
X_train = scl_x.fit_transform(X_train)
#X_test = scl_x.transform(X_test)
init_eta = 1e-1
batch_size = 30
train_iter = 50000
traceback_checkpoint = 45000
loo_extra_iter = 5000
decay_epochs = (10000, 20000)
checkpoint_iter = traceback_checkpoint-1
iter_to_switch_off_minibatch=35000
iter_to_switch_to_sgd=np.inf
# LOO a on random set of training indices, otherwise too slow
leave_indices = None
if hasattr(test_indices, '__iter__') and hasattr(leave_indices, '__iter__'):
assert not set(test_indices) & set(leave_indices)
print(test_indices)
print(leave_indices)
# -
# flip 10% indices
np.random.seed(43)
flip_indices = np.random.choice(n_tr, size=909, replace=False)
y_train_fliped = np.copy(y_train)
y_train_fliped[flip_indices,:] = 1-y_train_fliped[flip_indices,:]
model = BinaryLogisticRegression(
model_name='BinaryLogistic-MNIST',
init_eta=init_eta,
decay_epochs=decay_epochs,
batch_size=batch_size,
C=1e3
)
tf.reset_default_graph()
model.fit_with_sklearn({'X':X_train, 'y':y_train_fliped})
# bench mark
to_be_fixed = set(flip_indices)
high_influc = np.argsort(-np.abs(
model.get_eval(items=['losses'])).T)[0][0:200]
#need_to_fix = []
y_fixing = np.copy(y_train_fliped)
print(len(to_be_fixed))
for i in high_influc:
print(i,i in to_be_fixed)
if i in to_be_fixed:
y_fixing[i,:] = 1-y_fixing[i,:]
to_be_fixed -= set([i])
# fix ba
print(len(to_be_fixed))
model.fit_with_sklearn({'X':X_train, 'y':y_fixing})
# bench mark
# = set(flip_indices)
high_influc = np.argsort(-np.abs(
model.get_eval(items=['losses'])).T)[0][0:200]
#need_to_fix = []
#y_fixing = np.copy(y_train_fliped)
print(len(to_be_fixed))
for i in high_influc:
print(i, i in to_be_fixed)
#y_fixing[i,:] = 1-y_fixing[i,:]
# fix ba
# +
y_pred = model.predict(X_train)
print("Train accuracy:", np.sum(y_pred == y_train_fliped)/n_tr)
#y_pred_test = model.predict(X_test)
#print("Test accuracy:", np.sum(y_pred_test == y_test)/n_te)
# +
# %%time
I_loss_bf = model.influence_loss(
X_test, y_test,
method='brute-force',
damping=0.0,
)
# -
to_be_fixed = set(flip_indices)
frac_fixed = 0
frac_checked = 0
y_fixing = np.copy(y_train_fliped)
xx = np.zeros(6)
yy = np.zeros(6)
check_size = 200
for batch in range(5):
tf.reset_default_graph()
model.fit_with_sklearn({'X':X_train, 'y':y_fixing })
I_loss_bf = model.influence_loss(
X_test, y_test,
method='brute-force',
damping=0.0,
)
high_influc = np.argsort(-np.abs(I_loss_bf).T)[0][0:check_size]
need_to_fix = []
for i in high_influc:
if i in to_be_fixed:
# fix back
y_fixing[i,:] = 1-y_fixing[i,:]
frac_fixed += (1/909)
need_to_fix.append(i)
to_be_fixed = to_be_fixed - set(need_to_fix)
frac_checked += (check_size/n_tr)
xx[batch+1] = frac_checked
yy[batch+1] = frac_fixed
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
from data import Datafile, load_data
from influence.emp_risk_optimizer import EmpiricalRiskOptimizer
from influence.plot_utils import compare_with_loo
from influence.closed_forms import I_loss_RidgeCf
from models.regularized_regression import RegularizedRegression
from models.hyperplane_clf import BinaryLogisticRegression
# +
# flip 10% indices
from sklearn.preprocessing import StandardScaler
X_train, X_test, y_train, y_test, test_indices = load_data(
Datafile.BinaryMNIST17, test_config=[0])
n_tr, p = X_train.shape
n_te, _ = X_test.shape
print(n_tr, p)
scl_x = StandardScaler()
scl_y = StandardScaler()
X_train = scl_x.fit_transform(X_train)
#X_test = scl_x.transform(X_test)
np.random.seed(43)
flip_indices = np.random.choice(n_tr, size=909, replace=False)
y_train_fliped = np.copy(y_train)
y_train_fliped[flip_indices,:] = 1-y_train_fliped[flip_indices,:]
print(flip_indices)
# -
model = BinaryLogisticRegression(
model_name='BinaryLogistic-MNIST',
init_eta=0.01,
decay_epochs=[200, 400, 800],
batch_size=100,
C=1e3
)
to_be_fixed = set(flip_indices)
frac_fixed = 0
frac_checked = 0
y_fixing = np.copy(y_train_fliped)
print(np.mean(y_fixing == y_train))
print(len(to_be_fixed))
xx = np.zeros(6)
zz = np.zeros(6)
check_size = 200
for batch in range(5):
tf.reset_default_graph()
model.fit_with_sklearn({'X':X_train, 'y':y_fixing })
"""
I_loss_bf = model.influence_loss(
X_test, y_test,
method='brute-force',
damping=0.0,
)
"""
high_influc = np.argsort(-np.abs(
model.get_eval(items=['losses'])).T)[0][0:check_size]
need_to_fix = []
for i in high_influc:
if i in to_be_fixed:
# fix back
y_fixing[i,:] = 1-y_fixing[i,:]
frac_fixed += (1/909)
need_to_fix.append(i)
to_be_fixed = to_be_fixed - set(need_to_fix)
print(len(to_be_fixed))
print(np.mean(y_fixing == y_train))
frac_checked += (check_size/n_tr)
xx[batch+1] = frac_checked
zz[batch+1] = frac_fixed
print(zz)
frac_data_check = np.copy(xx)
frac_fixed_influence = np.copy(yy)
plt.plot(frac_data_check,frac_fixed_influence)
plt.plot(frac_data_check,zz)
plt.plot(frac_data_check,frac_data_check)
zz
| experiments6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Fitting Decision Tree Regression to the dataset
from sklearn.tree import DecisionTreeRegressor
import pandas as pd
from sklearn.model_selection import train_test_split
import numpy as np
from sklearn import preprocessing
import matplotlib.pyplot as plt
plt.rc("font", size=14)
from sklearn.linear_model import LogisticRegression
import seaborn as sns
sns.set(style="white")
sns.set(style="whitegrid", color_codes=True)
df = pd.read_csv("Replaced.csv",encoding="ISO-8859-1")
df.to_csv("data.csv",encoding="UTF-8")
df.columns
# #### Converting username and id in hash value to convert string to int
def get_hash(x):
return abs(hash(x)) % 10**9
df['username'] = df['username'].apply(get_hash)
df['id'] = df['id'].apply(get_hash)
df.columns
dataset = df[['id','username','rating']]
X = dataset.iloc[:,0:1].values
y = dataset.iloc[:, 2].values
print(X)
print(y)
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.4, random_state = 0)
# ### DecisionTreeRegressor
#
# To predict rating of product where max_depth = 5 is taken which gives more accuracy than max_depth = 2
regressor = DecisionTreeRegressor(random_state = 0,max_depth=5)
regressor.fit(X, y)
# ### Predicting a new result
# +
y_pred = regressor.predict(X)
# -
# ### y_pred gives the predicted rating
y_pred
print(y_pred.shape)
print(y.shape)
import matplotlib.pyplot as plt
X.shape
# ### Plot the result of predicted rating
# Plot the results
plt.figure()
plt.scatter(y, y_pred, s=20, edgecolor="black",
c="darkorange")
plt.xlabel("data")
plt.ylabel("target")
plt.title("Decision Tree Regression")
plt.legend()
plt.show()
# ### Analysis of Recommended data
sns.countplot( x = df['doRecommend'], data = df, palette = 'hls')
plt.show()
# #### There are aproximately 55000 true’s and 5000 false’s in the outcome variables.
df.groupby('doRecommend').mean()
# ### Purchase frequency of product and rating
# %matplotlib inline
pd.crosstab(df.rating,df.doRecommend).plot(kind='bar')
plt.title('Purchase Frequency for Product')
plt.xlabel('Rating')
plt.ylabel('Frequency of Purchase')
plt.show()
# ### Considering id,username,rating,didPurchase and doRecommend columns for x_train set
X_train, X_test, y_train, y_test = train_test_split(df, y,test_size=0.2)
X_train= X_train[['id','username','rating','didPurchase','doRecommend']]
print(y_train)
# ### Performing Linear Regression to predict the rating and get relationship among the selected columns from the dataset
# +
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X, y)
#predecting the test set results
y_pred = regressor.predict(X)
#Visualization of the training set results
plt.scatter(y, y_pred, color = 'red')
plt.title('salary vs yearExp (Training set)')
plt.xlabel('years of experience')
plt.ylabel('salary')
plt.show()
# -
n_users = df.username.unique().shape[0]
n_items = df.id.unique().shape[0]
print ('Number of users = ' + str(n_users) + ' | Number of products = ' + str(n_items))
# ## Random Forest Classifier
# <b> Random decision forests correct for decision trees' habit of overfitting to their training set. A random forest is a meta estimator that fits a number of decision tree classifiers on various sub-samples of the dataset and use averaging to improve the predictive accuracy and control over-fitting</b>
# Load scikit's random forest classifier library
from sklearn.ensemble import RandomForestClassifier
df = pd.read_csv("training.csv")
df.columns
df['didPurchase'].fillna(True)
# ### Convert didPurchase and doRecommend in int column from boolean
# ### True is replaced by 1 and False is replaced by 0
df.didPurchase = (df.didPurchase)*1
df.doRecommend = (df.doRecommend)*1
df['didPurchase'].head(100)
df['doRecommend'] = df['doRecommend'].fillna(1)
df['didPurchase'] = df['didPurchase'].fillna(1)
df['doRecommend'].head(100)
# +
# Create a list of the feature column's names
features = df[['id','username','didPurchase','rating']]
# View features
features
# -
# ## Train Random Forest Classifier
# ## predicting if the product is recommended or not
y = df['doRecommend']
y.head()
# ### Applying RandomForest Classifier with n_jobs = 2 which initializes 2 threads to fit the model
# +
# Create a random forest Classifier. By convention, clf means 'Classifier'
clf = RandomForestClassifier(n_jobs=2, random_state=0)
# Train the Classifier to take the training features and learn how they relate to the training y (the rating)
clf.fit(features, y)
# -
# ### Using test.csv to check if the model is trained well
df_test = pd.read_csv("test.csv")
# ### Convert didPurchase and doRecommend in int column from boolean
# ### True is replaced by 1 and False is replaced by 0
df_test.didPurchase = (df.didPurchase)*1
df_test.doRecommend = (df_test.doRecommend)*1
df_test['doRecommend'] = df_test['doRecommend'].fillna(1)
df_test['didPurchase'] = df_test['didPurchase'].fillna(1)
test_features = df_test[['id','didPurchase','username','rating']]
# Apply the Classifier we trained to the test data (which, remember, it has never seen before)
preds = clf.predict(test_features)
print(preds)
# View the predicted probabilities of the first 10 observations
clf.predict_proba(test_features)[0:10]
# ### Creating confusion matrix which shows total number of actual and predicted values of
# ### recommended(1) and non-recommended(0) products
# Create confusion matrix
pd.crosstab(df_test['doRecommend'], preds, rownames=['Actual Recommendation'], colnames=['Predicted Recommendation'])
# ### Below code shows the feature importance to predict the recommended product for the user.
# ### rating is highly effective for recommendation of product having value 0.45
# View a list of the features and their importance scores
list(zip(features, clf.feature_importances_))
| .ipynb_checkpoints/Part A - DecisionTree_RandomForest-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Frame the Problem
#
# - Descriptive
# - Exploratory
# - Inferential
#
#
# # 2. Acquire the Data
#
# > "Data is the new oil"
#
# - Download from an internal system
# - Obtained from client, or other 3rd party
# - Extracted from a web-based API
# - Scraped from a website
# - Extracted from a PDF file
# - Gathered manually and recorded
#
# We will using the Global Information System on Alcohol and Health (GISAH) maintained by WHO to answer the questions.
#
# *The WHO Global Information System on Alcohol and Health (GISAH) provides easy and rapid access to a wide range of alcohol-related health indicators. It is an essential tool for assessing and monitoring the health situation and trends related to alcohol consumption, alcohol-related harm, and policy responses in countries.*
#
# You can see an overview at http://www.who.int/gho/alcohol/en/.
#
#
# ## Principle: Load the Data
#
# The datasets from GISAH are available at http://apps.who.int/gho/data/node.main.GISAH?lang=en&showonly=GISAH
#
# We want the alcohol consumption by country
#
# - Recorded alcohol per capita consumption, 1960-1979 by country - http://apps.who.int/gho/data/node.main.A1025?lang=en&showonly=GISAH
#
# - Recorded alcohol per capita consumption, 1980-1999 by country - http://apps.who.int/gho/data/node.main.A1024?lang=en&showonly=GISAH
#
# - Recorded alcohol per capita consumption, 2000 onwards by country http://apps.who.int/gho/data/node.main.A1026?lang=en&showonly=GISAH
# Import the libraries we need, which is Pandas and Numpy
import pandas as pd
import numpy as np
df1 = pd.read_csv('data/drinks2000.csv')
df1.head()
df1.shape
# ## Principle: Fix the Column Header
years1 = list(range(2015, 1999, -1))
years1
header1 = ['description']
header1.extend(years1)
header1
df1.columns = header1
df1.head()
# ### Exercise
# 1. Load the drinks1960 and drinks1980 csv files and fix the column header
df2 = pd.read_csv('data/drinks1980.csv')
years2 = list(range(1999, 1979, -1))
header2 = ['description']
header2.extend(years2)
df2.columns = header2
df2.head()
df3 = pd.read_csv('data/drinks1960.csv')
years3 = list(range(1979, 1959, -1))
header3 = ['description']
header3.extend(years3)
df3.columns = header3
df3.head()
# # 3. Refine the Data
#
# > "Data is messy"
#
# We will be performing the following operation on our Onion price to refine it
# - **Remove** e.g. remove redundant data from the data frame
# - **Derive** e.g. Country and Beverage from the description field
# - **Missing** e.g. Check for missing or incomplete data
# - **Merge** e.g. Take the three dataframes and make them one
# - **Filter** e.g. exclude based on location
#
# Other stuff you may need to do to refine are...
# - **Parse** e.g. extract date from year and month column
# - **Quality** e.g. Check for duplicates, accuracy, unusual data
# - **Convert** e.g. free text to coded value
# - **Calculate** e.g. percentages, proportion
# - **Aggregate** e.g. rollup by year, cluster by area
# - **Sample** e.g. extract a representative data
# - **Summary** e.g. show summary stats like mean
#
# ## Principle: `melt` to convert from Wide format to Tall format
# We will need to convert the data frame from wide format to tall format (and vice versa). This is needed as we want to combine the three data frame and we can only do that once we have the data in a tall format
#
# 
df1.head()
df1 = pd.melt(df1, id_vars=['description'], var_name='year')
df1.head()
df2 = pd.melt(df2, id_vars=['description'], var_name='year')
df3 = pd.melt(df3, id_vars=['description'], var_name='year')
# ## Principle: `append` one dataframe to another
df1.shape
df2.shape
df = df1.append(df2)
df.shape
df = df.append(df3)
df.shape
# ## Principle: `str` to extract text from a strings
#
# String manipulation is very common and we often need to extract a substring from a long string. In this case, we want to get the country and type of beverage from the description
df.head()
df['country'] = df.description.str.split(';').str[0]
df.head()
df['beverage'] = df.description.str.split(";").str[-1]
df.tail()
# We can now drop the description column from our dataframe
df.drop('description', axis = 1, inplace= True)
df.head()
# ## Principle: Dealing with Missing Values
#
# By “missing” it simply mean null or “not present for whatever reason”. Many data sets have missing data, either because it exists and was not collected or it never existed. Pandas default way for treating missing value is to mark it as `NaN`
df.dtypes
df.year.unique()
df.year = pd.to_numeric(df.year)
df.dtypes
df.head()
# Lets check in the value whether we have numeric or not
df.value.unique()
df[df.value.str.isnumeric() == False].shape
# We will use `pd.to_numeric` which will coerce to NaN everything that cannot be converted to a numeric value, so strings that represent numeric values will not be removed. For example '1.25' will be recognized as the numeric value 1.25
df.value = pd.to_numeric(df.value, errors='coerce')
df.value.unique()
df.dtypes
df.country.unique()
df.beverage.unique()
# Convert from an np array to a list
beverage_old = df.beverage.unique().tolist()
beverage_old
# Create a new list with white space removed and shorter names
beverage_new = ['all', 'beer', 'wine', 'spirits', 'others']
beverage_new
df.beverage = df.beverage.replace(beverage_old, beverage_new)
# ## Principle: filter for rows in a dataframe
#
# - To select the rows from the dataframe
#
# 
df2015 = df[df.year == 2015]
df2015.head()
dfBeer = df[df.beverage == 'beer']
df2010Beer = df[(df.year == 2010) & (df.beverage == 'beer')]
df2010Beer.tail()
| drinks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df_category = pd.read_csv('exports/category_skill.csv')
df_job = pd.read_csv('exports/job_skill.csv')
# +
import requests
import json
def jprint(obj):
# create a formatted string of the Python JSON object
text = json.dumps(obj, sort_keys=True, indent=4)
print(text)
# -
# # Import prepared data directly to database
# ---
# ## Connect to MongoDB
# +
from pymongo import MongoClient
# build a new client instance of MongoClient
mongo_client = MongoClient('localhost', 27017)
# connect database
db = mongo_client.skillguider
# -
# drop collections before import new data
db['skills'].drop()
db['categories'].drop()
db['jobs'].drop()
# ---
# ## Import all skills to database
# ### Preparing data
# +
df_all_skill = pd.concat([df_category, df_job], axis=0)
df_all_skill = df_all_skill.drop(['sum', 'count', 'priority', 'job', 'category'], axis=1).drop_duplicates().reset_index(drop=True)
df_all_skill = df_all_skill.rename(columns={'skill': 'title'})
df_all_skill.loc[df_all_skill.index[:], 'title'] = df_all_skill['title'].str.capitalize()
with pd.option_context('display.max_rows', None, 'display.max_columns', None):
display(df_all_skill)
# -
df_keyword_old = pd.read_csv('exports/keyword.csv')
df_keyword_new = pd.merge(df_keyword_old, df_all_skill, on='title', how='right').replace(np.nan, '', regex=True)
df_keyword_new.to_csv('exports/keyword.csv', index=False)
skill_json = df_keyword_new.to_json('exports/temp/all_skills.json', orient='records', default_handler=str)
# +
with open('exports/temp/all_skills.json') as f:
data = json.load(f)
db['skills'].insert_many(data)
# -
# ---
# ## Import categories to database
# ### Preparing data
cursor = db['skills'].find({})
df_skill_with_id = pd.DataFrame(list(cursor), columns = ['_id', 'title'])
df_skill_with_id = df_skill_with_id.rename(columns={'title': 'skill'})
df_skill_with_id.loc[df_skill_with_id.index[:], 'skill'] = df_skill_with_id['skill'].str.lower()
df_skill_with_id
df_category_skill = df_category.drop(['sum', 'count'], axis=1).reset_index(drop=True)
df_merge_category = pd.merge(df_category_skill, df_skill_with_id, on='skill').sort_values(by=['category', 'skill'], ascending=False).reset_index(drop=True)
df_merge_category = df_merge_category.drop(['skill'], axis=1).rename(columns={'_id': 'skill_id', 'category': 'title'})
with pd.option_context('display.max_rows', None, 'display.max_columns', None):
display(df_merge_category)
# ### Convert to json file on collection format
columns = df_merge_category.columns.difference(['title'])
category_json = df_merge_category.groupby(['title'])[columns].apply(lambda x: x.to_dict('r')).reset_index(name='skillset').to_json('exports/temp/categories.json', orient='records', default_handler=str)
# ### Insert json data to MongoDB
# +
with open('exports/temp/categories.json') as f:
data = json.load(f)
db['categories'].insert_many(data)
# -
# ---
# ## Import jobs to database
# ### Preparing data
cursor = db['categories'].find({})
df_category_with_id = pd.DataFrame(list(cursor), columns = ['_id', 'title'])
df_category_with_id = df_category_with_id.rename(columns={'title': 'category'})
df_job_skill = df_job.drop(['sum'], axis=1).reset_index(drop=True)
df_merge_job = pd.merge(df_job_skill, df_skill_with_id, on='skill')
df_merge_job = df_merge_job.rename(columns={'_id': 'skill_id'})
df_merge_job = pd.merge(df_merge_job, df_category_with_id, on='category').sort_values(by=['job', 'skill'], ascending=False).reset_index(drop=True)
df_merge_job = df_merge_job.rename(columns={'_id': 'category_id'})
df_merge_job = df_merge_job.drop(['skill', 'category'], axis=1).rename(columns={'job': 'title'})
df_merge_job['description'] = ''
with pd.option_context('display.max_rows', None, 'display.max_columns', None):
display(df_merge_job.head())
# ### Convert to json file on collection format
columns = df_merge_job.columns.difference(['title', 'category_id', 'description'])
job_json = df_merge_job.groupby(['title', 'category_id', 'description'])[columns].apply(lambda x: x.to_dict('r')).reset_index(name='skillset').to_json('exports/temp/jobs.json', orient='records', default_handler=str)
# ### Insert json data to MongoDB
# +
with open('exports/temp/jobs.json') as f:
data = json.load(f)
db['jobs'].insert_many(data)
# -
mongo_client.close()
| import-database.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.0
# language: julia
# name: julia-1.4
# ---
# ## A Live [SEIR](https://en.wikipedia.org/wiki/Compartmental_models_in_epidemiology#The_SEIR_model) Model for Covid-19 with Error Bars in Julia
#
# #### Author: <NAME> (@[djsegal](https://github.com/djsegal/))
#
# as requested by Prof. <NAME>, MIT
#
# ----
#
# +
using Measurements
param_error = 0.05
α = 0.2 ± param_error
γ = 0.5 ± param_error
# β = f(social distancing, masks, etc.)
E₀ = 1e-4
u₀ = [ 1 - E₀ ; E₀ ; 0 ; 0 ]
tspan = (0.0, 200.0); # 200 days
# +
using DifferentialEquations
function seir!(du,u,p,t)
S,E,I,_ = u
α,β,γ = p
dS = ( -β*I*S )
dE = ( +β*I*S ) - α*E
dI = ( -γ*I ) + α*E
dR = ( +γ*I )
du .= (dS, dE, dI, dR)
end
labels = [
"Susceptible", "Exposed",
"Infected", "Recovered"
];
# +
using SimplePlots
display(html"""
<h3> Sliding the β value reduces/increases infectivity </h3>
<p style="margin-top: 0.4em">(social distancing, immunity, vaccines reduce β)</p>
""")
beta_slider = slider(0.25:0.25:2, value=1, label="β")
@demo for β in beta_slider
title!("Covid-19 SEIR Model with Error Bars") ;
xlabel!("Time (Days)") ; ylabel!("%")
soln = solve(ODEProblem(
seir!, u₀, tspan, [α, β ± param_error, γ]
), saveat=0.5)
x, y = soln.t, 100*Array(soln)
ylims!(0, 100) # using percentages
for (index, label) in enumerate(labels)
plot!(
x, y[index,:], label=label,
alpha=0.8, fillalpha=0.1
)
end
end
| binder/covid-19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %load_ext noworkflow
# nip = %now_ip
def first(x):
return next(x)
# -
# %now_sql_schema
# %now_prolog_schema
def extract(trial_id, filename, month):
t = nip.Trial(trial_id)
sql = first(nip.persistence.query("""
SELECT name, content_hash_before
FROM file_access
WHERE trial_id = {}
AND name = "{}"
""".format(trial_id, filename)))
fhash = sql['content_hash_before']
content = nip.persistence.get(fhash)
with open('.temp.dat', 'w') as f:
f.write(content)
result = !./precipitation.py .temp.dat $month
return sum(map(float,result[0].split(';')))
extract(18, 'p13.dat', 2)
%
| tests/tapp/Presentation/Backup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github"
# <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/main/notebooks-d2l/gru_jax.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7jysFtXPFE7E"
# # Gated Recurrent Units
#
# We show how to implement GRUs from scratch.
# Based on sec 9.1 of http://d2l.ai/chapter_recurrent-modern/gru.html
# This uses code from the [basic RNN colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/rnn_torch.ipynb).
#
# + id="eemDrShfots1" colab={"base_uri": "https://localhost:8080/"} outputId="d258c929-2fa4-45b8-88c3-61f21e5a7202"
# !pip install -q flax
# + id="o6VNlv_FYTbS"
import jax.numpy as jnp
import matplotlib.pyplot as plt
import math
from IPython import display
import jax
import flax.linen as nn
from flax import jax_utils
import optax
import collections
import re
import random
import os
import requests
import hashlib
import time
import functools
random.seed(0)
rng = jax.random.PRNGKey(0)
# !mkdir figures # for saving plots
# + [markdown] id="phjRpyDNFT14"
# # Data
#
# As data, we use the book "The Time Machine" by <NAME>,
# preprocessed using the code in [this colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/text_preproc_torch.ipynb).
# + id="tLIKRJomvBV5"
class SeqDataLoader:
"""An iterator to load sequence data."""
def __init__(self, batch_size, num_steps, use_random_iter, max_tokens):
if use_random_iter:
self.data_iter_fn = seq_data_iter_random
else:
self.data_iter_fn = seq_data_iter_sequential
self.corpus, self.vocab = load_corpus_time_machine(max_tokens)
self.batch_size, self.num_steps = batch_size, num_steps
def __iter__(self):
return self.data_iter_fn(self.corpus, self.batch_size, self.num_steps)
class Vocab:
"""Vocabulary for text."""
def __init__(self, tokens=None, min_freq=0, reserved_tokens=None):
if tokens is None:
tokens = []
if reserved_tokens is None:
reserved_tokens = []
# Sort according to frequencies
counter = count_corpus(tokens)
self.token_freqs = sorted(counter.items(), key=lambda x: x[1],
reverse=True)
# The index for the unknown token is 0
self.unk, uniq_tokens = 0, ['<unk>'] + reserved_tokens
uniq_tokens += [
token for token, freq in self.token_freqs
if freq >= min_freq and token not in uniq_tokens]
self.idx_to_token, self.token_to_idx = [], dict()
for token in uniq_tokens:
self.idx_to_token.append(token)
self.token_to_idx[token] = len(self.idx_to_token) - 1
def __len__(self):
return len(self.idx_to_token)
def __getitem__(self, tokens):
if not isinstance(tokens, (list, tuple)):
return self.token_to_idx.get(tokens, self.unk)
return [self.__getitem__(token) for token in tokens]
def to_tokens(self, indices):
if not isinstance(indices, (list, tuple)):
return self.idx_to_token[indices]
return [self.idx_to_token[index] for index in indices]
# + id="-TvZsL4gtHQ2"
def tokenize(lines, token='word'):
"""Split text lines into word or character tokens."""
if token == 'word':
return [line.split() for line in lines]
elif token == 'char':
return [list(line) for line in lines]
else:
print('ERROR: unknown token type: ' + token)
def count_corpus(tokens):
"""Count token frequencies."""
# Here `tokens` is a 1D list or 2D list
if len(tokens) == 0 or isinstance(tokens[0], list):
# Flatten a list of token lists into a list of tokens
tokens = [token for line in tokens for token in line]
return collections.Counter(tokens)
def seq_data_iter_random(corpus, batch_size, num_stepsz):
"""Generate a minibatch of subsequences using random sampling."""
# Start with a random offset (inclusive of `num_steps - 1`) to partition a
# sequence
corpus = corpus[random.randint(0, num_steps - 1):]
# Subtract 1 since we need to account for labels
num_subseqs = (len(corpus) - 1) // num_steps
# The starting indices for subsequences of length `num_steps`
initial_indices = list(range(0, num_subseqs * num_steps, num_steps))
# In random sampling, the subsequences from two adjacent random
# minibatches during iteration are not necessarily adjacent on the
# original sequence
random.shuffle(initial_indices)
def data(pos):
# Return a sequence of length `num_steps` starting from `pos`
return corpus[pos:pos + num_steps]
num_batches = num_subseqs // batch_size
for i in range(0, batch_size * num_batches, batch_size):
# Here, `initial_indices` contains randomized starting indices for
# subsequences
initial_indices_per_batch = initial_indices[i:i + batch_size]
X = [data(j) for j in initial_indices_per_batch]
Y = [data(j + 1) for j in initial_indices_per_batch]
yield jnp.array(X), jnp.array(Y)
def seq_data_iter_sequential(corpus, batch_size, num_steps):
"""Generate a minibatch of subsequences using sequential partitioning."""
# Start with a random offset to partition a sequence
offset = random.randint(0, num_steps)
num_tokens = ((len(corpus) - offset - 1) // batch_size) * batch_size
Xs = jnp.array(corpus[offset:offset + num_tokens])
Ys = jnp.array(corpus[offset + 1:offset + 1 + num_tokens])
Xs, Ys = Xs.reshape(batch_size, -1), Ys.reshape(batch_size, -1)
num_batches = Xs.shape[1] // num_steps
for i in range(0, num_steps * num_batches, num_steps):
X = Xs[:, i:i + num_steps]
Y = Ys[:, i:i + num_steps]
yield X, Y
# + id="yRp-MH0Nv7rN"
def download(name, cache_dir=os.path.join('..', 'data')):
"""Download a file inserted into DATA_HUB, return the local filename."""
assert name in DATA_HUB, f"{name} does not exist in {DATA_HUB}."
url, sha1_hash = DATA_HUB[name]
os.makedirs(cache_dir, exist_ok=True)
fname = os.path.join(cache_dir, url.split('/')[-1])
if os.path.exists(fname):
sha1 = hashlib.sha1()
with open(fname, 'rb') as f:
while True:
data = f.read(1048576)
if not data:
break
sha1.update(data)
if sha1.hexdigest() == sha1_hash:
return fname # Hit cache
print(f'Downloading {fname} from {url}...')
r = requests.get(url, stream=True, verify=True)
with open(fname, 'wb') as f:
f.write(r.content)
return fname
def read_time_machine():
"""Load the time machine dataset into a list of text lines."""
with open(download('time_machine'), 'r') as f:
lines = f.readlines()
return [re.sub('[^A-Za-z]+', ' ', line).strip().lower() for line in lines]
def load_corpus_time_machine(max_tokens=-1):
"""Return token indices and the vocabulary of the time machine dataset."""
lines = read_time_machine()
tokens = tokenize(lines, 'char')
vocab = Vocab(tokens)
# Since each text line in the time machine dataset is not necessarily a
# sentence or a paragraph, flatten all the text lines into a single list
corpus = [vocab[token] for line in tokens for token in line]
if max_tokens > 0:
corpus = corpus[:max_tokens]
return corpus, vocab
def load_data_time_machine(batch_size, num_steps, use_random_iter=False,
max_tokens=10000):
"""Return the iterator and the vocabulary of the time machine dataset."""
data_iter = SeqDataLoader(batch_size, num_steps, use_random_iter,
max_tokens)
return data_iter, data_iter.vocab
# + id="nG2T4maNYdsG" colab={"base_uri": "https://localhost:8080/"} outputId="dc1bd6ea-584a-439b-f0a8-19394e361b4a"
DATA_HUB = dict()
DATA_URL = 'http://d2l-data.s3-accelerate.amazonaws.com/'
DATA_HUB['time_machine'] = (DATA_URL + 'timemachine.txt',
'090b5e7e70c295757f55df93cb0a180b9691891a')
batch_size, num_steps = 32, 35
train_iter, vocab = load_data_time_machine(batch_size, num_steps)
# + [markdown] id="fKa87dOCFZ8H"
# # Creating the model from scratch
# + [markdown] id="gq3jJ8IQFjz3"
# Initialize the parameters.
# + id="WnwqslgzFW7E"
def get_params(vocab_size, num_hiddens, init_rng):
num_inputs = num_outputs = vocab_size
def normal(shape, rng):
return jax.random.normal(rng, shape=shape) * 0.01
def three(rng):
return (normal(
(num_inputs, num_hiddens), rng), normal((num_hiddens, num_hiddens),
rng), jnp.zeros(num_hiddens))
update_rng, reset_rng, hidden_rng, out_rng = jax.random.split(init_rng,
num=4)
W_xz, W_hz, b_z = three(update_rng) # Update gate parameters
W_xr, W_hr, b_r = three(reset_rng) # Reset gate parameters
W_xh, W_hh, b_h = three(hidden_rng) # Candidate hidden state parameters
# Output layer parameters
W_hq = normal((num_hiddens, num_outputs), out_rng)
b_q = jnp.zeros(num_outputs)
params = [W_xz, W_hz, b_z, W_xr, W_hr, b_r, W_xh, W_hh, b_h, W_hq, b_q]
return params
# + [markdown] id="xiMRAVr2Ftir"
# Initial state is an array of zeros of size (batch-size, num-hiddens)
# + id="rbMB5SnMFvkZ"
def init_gru_state(batch_size, num_hiddens):
return (jnp.zeros((batch_size, num_hiddens)),)
# + [markdown] id="9LcxOAdiF1kd"
# Forward function
# + id="SfC5HhBbFv6t"
@jax.jit
def gru(params, state, inputs):
W_xz, W_hz, b_z, W_xr, W_hr, b_r, W_xh, W_hh, b_h, W_hq, b_q = params
H, = state
outputs = []
for X in inputs:
Z = jax.nn.sigmoid((X @ W_xz) + (H @ W_hz) + b_z)
R = jax.nn.sigmoid((X @ W_xr) + (H @ W_hr) + b_r)
H_tilda = jnp.tanh((X @ W_xh) + ((R * H) @ W_hh) + b_h)
H = Z * H + (1 - Z) * H_tilda
Y = H @ W_hq + b_q
outputs.append(Y)
return jnp.concatenate(outputs, axis=0), (H,)
# + id="Qw3vhoRKdVxt"
# Make the model class
# Input X to apply is (B,T) matrix of integers (from vocab encoding).
# We transpose this to (T,B) then one-hot encode to (T,B,V), where V is vocab.
# The result is passed to the forward function.
# (We define the forward function as an argument, so we can change it later.)
class RNNModelScratch:
"""A RNN Model implemented from scratch."""
def __init__(self, vocab_size, num_hiddens, get_params, init_state,
forward_fn):
self.vocab_size, self.num_hiddens = vocab_size, num_hiddens
self.init_state, self.get_params = init_state, get_params
self.forward_fn = forward_fn
def apply(self, params, state, X):
X = jax.nn.one_hot(X.T, num_classes=self.vocab_size)
return self.forward_fn(params, state, X)
def begin_state(self, batch_size):
return self.init_state(batch_size, self.num_hiddens)
def init_params(self, rng):
return self.get_params(self.vocab_size, self.num_hiddens, rng)
# + [markdown] id="WtoCsGZ2F8G0"
# # Training and prediction
# + id="LDuvumDhgpvh"
@jax.jit
def grad_clipping(grads, theta):
"""Clip the gradient."""
def grad_update(grads):
return jax.tree_map(lambda g: g * theta / norm, grads)
norm = jnp.sqrt(sum(jax.tree_util.tree_leaves(jax.tree_map(
lambda x: jnp.sum(x ** 2), grads))))
# Update gradient if norm > theta
# This is jax.jit compatible
grads = jax.lax.cond(norm > theta, grad_update, lambda g: g, grads)
return grads
# + id="tA_QeZH5rp1o"
class Animator:
"""For plotting data in animation."""
def __init__(self, xlabel=None, ylabel=None, legend=None, xlim=None,
ylim=None, xscale='linear', yscale='linear',
fmts=('-', 'm--', 'g-.', 'r:'), nrows=1, ncols=1,
figsize=(3.5, 2.5)):
# Incrementally plot multiple lines
if legend is None:
legend = []
display.set_matplotlib_formats('svg')
self.fig, self.axes = plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1:
self.axes = [self.axes,]
# Use a lambda function to capture arguments
self.config_axes = lambda: set_axes(self.axes[
0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
# Add multiple data points into the figure
if not hasattr(y, "__len__"):
y = [y]
n = len(y)
if not hasattr(x, "__len__"):
x = [x] * n
if not self.X:
self.X = [[] for _ in range(n)]
if not self.Y:
self.Y = [[] for _ in range(n)]
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
class Timer:
"""Record multiple running times."""
def __init__(self):
self.times = []
self.start()
def start(self):
"""Start the timer."""
self.tik = time.time()
def stop(self):
"""Stop the timer and record the time in a list."""
self.times.append(time.time() - self.tik)
return self.times[-1]
def avg(self):
"""Return the average time."""
return sum(self.times) / len(self.times)
def sum(self):
"""Return the sum of time."""
return sum(self.times)
def cumsum(self):
"""Return the accumulated time."""
return jnp.array(self.times).cumsum().tolist()
class Accumulator:
"""For accumulating sums over `n` variables."""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a + float(b) for a, b in zip(self.data, args)]
def reset(self):
self.data = [0.0] * len(self.data)
def __getitem__(self, idx):
return self.data[idx]
def set_axes(axes, xlabel, ylabel, xlim, ylim, xscale, yscale, legend):
"""Set the axes for matplotlib."""
axes.set_xlabel(xlabel)
axes.set_ylabel(ylabel)
axes.set_xscale(xscale)
axes.set_yscale(yscale)
axes.set_xlim(xlim)
axes.set_ylim(ylim)
if legend:
axes.legend(legend)
axes.grid()
@jax.jit
def sgd(params, grads, lr, batch_size):
"""Minibatch stochastic gradient descent."""
params = jax.tree_map(lambda p, g: p - lr * g / batch_size, params,
grads)
return params
# + id="X7o1dMFttw_2"
@jax.jit
def train_step(apply_fn, loss_fn, params, state, X, Y):
def loss(params, state, X, Y):
y = Y.T.reshape(-1) #(B,T) -> (T,B)
y_hat, state = apply_fn(params, state, X)
y_hat = y_hat.reshape(-1, y_hat.shape[-1])
y_one_hot = jax.nn.one_hot(y, num_classes=y_hat.shape[-1])
return loss_fn(y_hat, y_one_hot).mean(), state
grad_fn = jax.value_and_grad(loss, has_aux=True)
(l, state), grads = grad_fn(params, state, X, Y)
grads = grad_clipping(grads, 1)
return l, state, grads
# + id="Eq_QMLUKhw3k"
def train_epoch(net, params, train_iter, loss, updater, use_random_iter):
state, timer = None, Timer()
metric = Accumulator(2) # Sum of training loss, no. of tokens
if isinstance(updater, optax.GradientTransformation):
updater_state = updater.init(params)
# Convert to jax Partial functions for jax.jit compatibility
apply_fn = jax.tree_util.Partial(net.apply)
loss_fn = jax.tree_util.Partial(loss)
for X, Y in train_iter:
if state is None or use_random_iter:
# Initialize `state` when either it is the first iteration or
# using random sampling
state = net.begin_state(batch_size=X.shape[0])
l, state, grads = train_step(apply_fn, loss_fn, params, state, X, Y)
if isinstance(updater, optax.GradientTransformation):
updates, updater_state = updater.update(grads, updater_state)
params = optax.apply_updates(params, updates)
else:
# batch_size=1 since the `mean` function has been invoked
params = updater(params, grads, batch_size=1)
metric.add(l * Y.size, Y.size)
return params, math.exp(metric[0] / metric[1]), metric[1] / timer.stop()
# + id="0tPGxQDWiqfl"
def train(net, params, train_iter, vocab, lr, num_epochs,
use_random_iter=False):
loss = optax.softmax_cross_entropy
animator = Animator(xlabel='epoch', ylabel='perplexity',
legend=['train'], xlim=[10, num_epochs])
# Initialize
if isinstance(net, nn.Module):
updater = optax.sgd(lr)
else:
updater = lambda params, grads, batch_size: sgd(params, grads, lr,
batch_size)
num_preds = 50
predict_ = lambda prefix: predict(prefix, num_preds, net, params, vocab)
# Train and predict
for epoch in range(num_epochs):
params, ppl, speed = train_epoch(net, params, train_iter, loss, updater,
use_random_iter)
if (epoch + 1) % 10 == 0:
# Prediction takes time on the flax model
# print(predict_('time traveller'))
animator.add(epoch + 1, [ppl])
device = jax.default_backend()
print(f'perplexity {ppl:.1f}, {speed:.1f} tokens/sec on {device}')
print(predict_('time traveller'))
print(predict_('traveller'))
return params
# + id="MvGhZUIGd3F3"
def predict(prefix, num_preds, net, params, vocab):
"""Generate new characters following the `prefix`."""
state = net.begin_state(batch_size=1)
outputs = [vocab[prefix[0]]]
get_input = lambda: jnp.array([outputs[-1]]).reshape((1, 1))
for y in prefix[1:]: # Warm-up period
_, state = net.apply(params, state, get_input())
outputs.append(vocab[y])
for _ in range(num_preds): # Predict `num_preds` steps
y, state = net.apply(params, state, get_input())
y = y.reshape(-1, y.shape[-1])
outputs.append(int(y.argmax(axis=1).reshape(1)))
return ''.join([vocab.idx_to_token[i] for i in outputs])
# + colab={"base_uri": "https://localhost:8080/", "height": 314} id="rw0uiJ4aF3jI" outputId="7be5583c-8745-4a88-cdd1-83c0779d533f"
random.seed(0)
vocab_size, num_hiddens = len(vocab), 256
num_epochs, lr = 500, 1
model = RNNModelScratch(len(vocab), num_hiddens, get_params, init_gru_state,
gru)
params = model.init_params(rng)
params = train(model, params, train_iter, vocab, lr, num_epochs)
# + id="89IqhaoSwiRT"
class GRU(nn.Module):
@functools.partial(
nn.transforms.scan,
variable_broadcast='params',
in_axes=0, out_axes=0,
split_rngs={'params': False})
@nn.compact
def __call__(self, state, x):
return nn.GRUCell()(state, x)
@staticmethod
def initialize_carry(rng, batch_dims, size):
return nn.GRUCell.initialize_carry(rng, batch_dims, size)
# + id="wAw5qDBKQe7U"
class RNNModel(nn.Module):
"""The RNN model."""
rnn: nn.Module
vocab_size: int
num_hiddens: int
bidirectional: bool = False
def setup(self):
# If the RNN is bidirectional (to be introduced later),
# `num_directions` should be 2, else it should be 1.
if not self.bidirectional:
self.num_directions = 1
else:
self.num_directions = 2
@nn.compact
def __call__(self, state, inputs):
X = jax.nn.one_hot(inputs.T, num_classes=self.vocab_size)
state, Y = self.rnn(state, X)
output = nn.Dense(self.vocab_size)(Y)
return output, state
def begin_state(self, batch_size=1):
# Use fixed random key since default state init fn is just `zeros`.
return self.rnn.initialize_carry(jax.random.PRNGKey(0), (batch_size,),
num_hiddens)
# + id="yDw7Ss_0F9MD" colab={"base_uri": "https://localhost:8080/", "height": 314} outputId="b1773077-9c03-4b29-a462-ef3fe6bc0d46"
random.seed(0)
gru_layer = GRU()
model = RNNModel(rnn=gru_layer, vocab_size=len(vocab), num_hiddens=num_hiddens)
initial_state = model.begin_state(batch_size)
params = model.init(rng, initial_state, jnp.ones([batch_size, num_steps]))
params = train(model, params, train_iter, vocab, lr, num_epochs)
| notebooks-d2l/gru_jax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: seaborn-py38-latest
# language: python
# name: seaborn-py38-latest
# ---
# + tags=["hide"]
import seaborn as sns; sns.set(style="ticks")
# + active=""
# The default plot kind is a histogram:
# -
penguins = sns.load_dataset("penguins")
sns.displot(data=penguins, x="flipper_length_mm")
# + active=""
# Use the ``kind`` parameter to select a different representation:
# -
sns.displot(data=penguins, x="flipper_length_mm", kind="kde")
# There are three main plot kinds; in addition to histograms and kernel density estimates (KDEs), you can also draw empirical cumulative distribution functions (ECDFs):
sns.displot(data=penguins, x="flipper_length_mm", kind="ecdf")
# While in histogram mode, it is also possible to add a KDE curve:
sns.displot(data=penguins, x="flipper_length_mm", kde=True)
# To draw a bivariate plot, assign both ``x`` and ``y``:
sns.displot(data=penguins, x="flipper_length_mm", y="bill_length_mm")
# Currently, bivariate plots are available only for histograms and KDEs:
sns.displot(data=penguins, x="flipper_length_mm", y="bill_length_mm", kind="kde")
# For each kind of plot, you can also show individual observations with a marginal "rug":
g = sns.displot(data=penguins, x="flipper_length_mm", y="bill_length_mm", kind="kde", rug=True)
# + active=""
# Each kind of plot can be drawn separately for subsets of data using ``hue`` mapping:
# -
sns.displot(data=penguins, x="flipper_length_mm", hue="species", kind="kde")
# Additional keyword arguments are passed to the appropriate underlying plotting function, allowing for further customization:
sns.displot(data=penguins, x="flipper_length_mm", hue="species", multiple="stack")
# + active=""
# The figure is constructed using a :class:`FacetGrid`, meaning that you can also show subsets on distinct subplots, or "facets":
# -
sns.displot(data=penguins, x="flipper_length_mm", hue="species", col="sex", kind="kde")
# + active=""
# Because the figure is drawn with a :class:`FacetGrid`, you control its size and shape with the ``height`` and ``aspect`` parameters:
# -
sns.displot(
data=penguins, y="flipper_length_mm", hue="sex", col="species",
kind="ecdf", height=4, aspect=.7,
)
# + active=""
# The function returns the :class:`FacetGrid` object with the plot, and you can use the methods on this object to customize it further:
# -
g = sns.displot(
data=penguins, y="flipper_length_mm", hue="sex", col="species",
kind="kde", height=4, aspect=.7,
)
g.set_axis_labels("Density (a.u.)", "Flipper length (mm)")
g.set_titles("{col_name} penguins")
| doc/docstrings/displot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Get some data to play with
from sklearn.datasets import load_digits
digits = load_digits()
digits.keys()
digits.images.shape
print(digits.images[0])
# +
import matplotlib.pyplot as plt
# %matplotlib notebook
plt.matshow(digits.images[0], cmap=plt.cm.Greys)
# -
digits.data.shape
digits.target.shape
digits.target
# **Data is always a numpy array (or sparse matrix) of shape (n_samples, n_features)**
# Split the data to get going
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(digits.data,
digits.target)
# Really Simple API
# -------------------
# 0) Import your model class
from sklearn.svm import LinearSVC
# 1) Instantiate an object and set the parameters
svm = LinearSVC(C=0.1)
# 2) Fit the model
svm.fit(X_train, y_train)
# 3) Apply / evaluate
print(svm.predict(X_train))
print(y_train)
svm.score(X_train, y_train)
svm.score(X_test, y_test)
# And again
# ---------
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=50)
rf.fit(X_train, y_train)
rf.score(X_test, y_test)
| 01 - Introduction to Scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import cv2
from pascal_voc_writer import Writer
lists = ['circle','square','star','triangle']
for shape in lists:
path_folder = "./"+str(shape)
file_lists = os.listdir(path_folder)
os.chdir(path_folder)
print(path_folder)
for file in file_lists:
path_image = os.path.join(path_folder,file)
image = cv2.imread(file, 0)
_,thresh = cv2.threshold(image, 100, 255, cv2.THRESH_BINARY)
contours = cv2.findContours(thresh, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
for c in contours[1]:
if cv2.contourArea(c) <= 30000 and cv2.contourArea(c) >= 500 :
x,y,w,h = cv2.boundingRect(c)
cv2.rectangle(image, (x, y), (x + w, y + h), (0, 255,0), 2)
center = (x,y)
print (center)
height = image.shape[0]
width = image.shape[1]
# Writer(path, width, height)
writer = Writer(path_image, width, height)
# ::addObject(name, xmin, ymin, xmax, ymax)
writer.addObject(shape, x, y, (x + w), (y + h))
name_xml = file.split('.')[0]+".xml"
# ::save(path)
writer.save(name_xml)
os.chdir("..")
| 12th day - Autolabel shape dataset/auto_label.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# importing the required libraries
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
import pickle
# loading the dataset
df = pd.read_csv('../datasets/thyroid.csv')
# +
# first 5 rows
df.head()
# Age is in years/100
# Sex: 0:Female, 1:Male
# Age, TSH, T3, TT4, T4U, FTI are continuous variables
# rest are categorical variables
# -
# describing the structure of the dataset
df.describe()
# there are no null values in oud dataset
df.isna().sum()
# 3 - Hyperthyroidism
# 2 - Hypothyroidism
# 1 - Normal
df.Class.value_counts()
# splitting the data into test and train sets
X = df.drop('Class', axis=1)
y = df.Class
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, test_size=0.15, random_state=143)
# trying the Logistic Regression Model
model_lr = LogisticRegression(max_iter=1000)
model_lr.fit(X_test, y_test)
# Accuracy achieved = 92%
model_lr.score(X_test, y_test)
# trying the Random Forest Classifier Model
model_rf = RandomForestClassifier()
model_rf.fit(X_train, y_train)
# Accuracy achieved = 99%
model_rf.score(X_test, y_test)
predictions = model_rf.predict(X_test)
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, predictions)
# printing the confusion matrix
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sn
plt.figure(figsize=(6,4))
sn.heatmap(cm, annot=True, )
plt.xlabel('Predicted')
plt.ylabel('Truth')
# printing the classification report
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions))
# saving the trained model for later use
pickle.dump(model_rf,open('../Saved Models/thyroid.pickle','wb'))
| Diagnosis using ML/Model Training/Thyroid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Since its emergence in Asia late 2019, the coronavirus COVID-19 pandemic has been devastating. The virus spread to most countries causing severe respiratory infections and many human casualties. The virus also put half of the world population in lockdown which resulted in a slowdown of the world economy and a fall in stock prices.
#
# The goal of this tutorial is to introduce the steps for collecting and analyzing stock data in the context of the coronavirus pandemic. To do this, we will use Python, Google Sheets and Google Finance.
#
# In section 2 of the the tutorial, we will see how to configure Google Sheets in order to be able to interact with them using Python. In section 3, we will see how to collect stocks data using Google Finance and how to store this data in Google Sheets using Python. In section 4, we will see how to read the data from Google Sheets and analyze it using Python and Pandas.
#
# The source code for this tutorial can be found in this [github repository](https://github.com/adilmoujahid/coronavirus-covid19-stocks-analysis).
# # 1. The Case Study
#
# In this tutorial, we will focus on the S&P 500 companies. We will start by collecting the following data :
# * Stock prices in 3 different dates (January 1st, March 23rd and April 9th)
# * Number of outstanding shares for each company
# * Industry/Sector where the companies operate (following the GICS classification)
#
# After collecting and structuring the data, we will use ```Python``` and ```Pandas``` library to analyse the data.
# # 2. Technical Setup
#
# We will be using the following services and libraries to collect and analyze the data:
#
# * Wikipedia: We will use [this wikipedia page](https://en.wikipedia.org/wiki/List_of_S%26P_500_companies) to get the list of S&P 500 companies.
# * Google Finance: Google Finance is a website focusing on business news and financial information hosted by Google [1]. Google Finance doesn't have an API that we can use directly in Python, but it can be accessed from Google Sheets using a formula called GOOGLEFINANCE. We will use Python to write down the GOOGLEFINANCE formulas.
# * Goole Sheets: We will use Google Sheets as a backend to store stocks data. In order to interact with Google Sheets directly from Python, we need 3 libaries: [Google Auth](https://github.com/googleapis/google-auth-library-python), [gspread](https://gspread.readthedocs.io/en/latest/) and [gspread-pandas](https://gspread-pandas.readthedocs.io/en/latest/). We also need to configure Google Sheets to be able to access the spreadsheets using Python.
# * Python and Jupyter notebooks and Pandas: We will be using Python, Jupyter notebooks and Pandas to collect, store and analyze the data.
# 
# ## 2.1. Google Sheets Configuration in GCP
# In order to access Google Sheets from Python, we need a private key from Google Cloud Platfrom (GCP) that we can obtain using the following steps.
# * Step 1: Go to [Google Cloud platform](https://cloud.google.com/), log in using your google account and click on ```Console```.
# 
# * Step 2: Click on ```Select a project``` > ```NEW PROJECT```, enter the project name and click on ```CREATE```
# 
# * Step 3: Click on ```APIs & Services``` > ```Dashboard``` > ```ENABLE APIS AND SEVICES```.
# 
# * Step 4: Search for both Google Drive API and Google Sheets API and click on ```ENABLE```
# 
# * Step 5: Inside the Google Sheets API page, click on ```MANAGE```, ```CREATE CREDENTIALS``` and select ```Google Sheets API```.
# 
# * Step 6: Choose ```Web sever```, ```Application data``` and ```JSON``` type for the API key. Click ```Continue``` to download your private key in JSON format.
# 
# * Step 7: Once you download the JSON file, save in the same folder as your Jupyter notebook and copy the ```client_email``` information.
# 
# ## 2.2. Google Sheets Configuration
# As a last step, we need to create a new Google sheet and share it with the ```client_email``` that we created in the previous step. Open Google Drive, create a new Google Sheet, change its name to "stocks-data". Click on ```Share``` button, enter the ```client_email``` and click ```Send```.
# 
# # 3. Collecting and Storing Stocks Data
# Now that we have our google Sheets configured, we can start using ```Python``` and ```Jupyter Notebook``` to collect the data. We start by importing the different libraries that we need.
# %matplotlib inline
# +
import pandas as pd
import gspread
from google.oauth2.service_account import Credentials
from gspread_pandas import Spread, Client
import seaborn as sns
import matplotlib.pyplot as plt
# -
plt.style.use('fivethirtyeight')
# ### Getting the list of S&P 500 companies from Wikipedia
# As a first step in the data collection effort, we need to get the list of S&P 500 companies. To do this, we will use the following Wikipedia Page: [https://en.wikipedia.org/wiki/List_of_S%26P_500_companies](https://en.wikipedia.org/wiki/List_of_S%26P_500_companies)
# 
# Fortunatey, we can use ```Pandas``` to read the page, extract the table with S&P 500 companies and store them into a ```Pandas``` Dataframe.
# +
url = 'https://en.wikipedia.org/wiki/List_of_S%26P_500_companies'
stocks_df = pd.read_html(url, header=0)[0]
stocks_df.head()
# -
# The most important data that we need is:
# * Symbol: Stock Symbol
# * Security: Name of the company
# * GICS Sector: Sector where the company operates following the Global Industry Classification Standard (GICS).
# * GICS Sub Industry: Sub industry where the company operates following the Global Industry Classification Standard (GICS).
#
# We can start looking at some statistics. For example the number of companies in the list.
#Number of companis
len(stocks_df)
# We got 505 companies in our list, and not 500... This is because some companies have a dual-class stock structure and are listed more than once in the list. We can get the list of these companies by searching for the word "Class" in their security name.
stocks_df[stocks_df['Security'].str.contains("Class")]
# We got 10 companies with dual-class structure in the list. If we take that into consideration, we can see that we have 500 unique companies in the list.
# We can also check the number of companies by sector and sub industry.
stocks_df['GICS Sector'].value_counts()
stocks_df['GICS Sub Industry'].value_counts()
# ### Adding stocks data from Google Finance
# Now that we have the list of S&P 500 companies, we can add to our DataFrame Google Sheets formulas that will fetch from Google Finance stock prices and the number of outstanding shares for each company. Note that, these formulas will be executed once we save the Pandas DataFrame in our Google Sheet.
# You can find the documentation of GOOGLEFINANCE formulas here: https://support.google.com/docs/answer/3093281
# We start by adding stock prices in 3 different dates: January 1st, March 23rd and April 9th.
#
# * January 1st, 2020 is the first date of the year. We want to have this price in order to calculate the price drop since the beggining of 2020.
# * March 23rd, 2020 is the date when the S&P 500 reached the bottom in 2020.
# * April 9th, 2020 is the last date when the stock market was open (at the time of writing this blog post)
stocks_df["Price_1_1"] = stocks_df["Symbol"].apply(lambda x: '=INDEX(GOOGLEFINANCE("' + x + '","price", "1/1/2020"),2,2)')
stocks_df["Price_3_23"] = stocks_df["Symbol"].apply(lambda x: '=INDEX(GOOGLEFINANCE("' + x + '","price", "3/23/2020"),2,2)')
stocks_df["Price_4_9"] = stocks_df["Symbol"].apply(lambda x: '=INDEX(GOOGLEFINANCE("' + x + '","price", "4/9/2020"),2,2)')
# Next, we add the formula to calculate the number of outstanding shares for each company. We can use this data with the stock prices to calculate the market cap of companies at the 3 different dates.
stocks_df["Shares"] = stocks_df["Symbol"].apply(lambda x: '=GOOGLEFINANCE("' + x + '","shares")')
# ### Storing the DataFrame to Google Sheet
# We start by creating a variable that contains our credentials that we got from Google Cloud Plaltform.
# +
scope = ['https://spreadsheets.google.com/feeds',
'https://www.googleapis.com/auth/drive']
credentials = Credentials.from_service_account_file('./gsheet-stocks.json', scopes=scope)
# -
# Next, we read the empty google sheet in a variable that we call ```spread```.
client = Client(scope=scope, creds=credentials)
spread = Spread("stocks_analysis", client=client)
# We define the list of variables that we want to keep.
cols_to_keep = ["Symbol", "Security", "GICS Sector", "GICS Sub Industry",
"Price_1_1", "Price_3_23", "Price_4_9", "Shares"]
# The last step is to save the DataFrame to Google Sheets.
spread.df_to_sheet(stocks_df[cols_to_keep])
# If we go to Google Sheets, we can see that the data is correctly stored.
# 
# # 4. Analyzing the Data
# ## 4.1. Reading the data
# We start by reading the data from Google Sheets into a new DataFrame.
stocks_df = spread.sheet_to_df()
stocks_df.head()
# we can see that the DataFrame contains real values for stocks prices and number of shares (and not Google Sheets formulas).
# We need to change the data type of stock prices and number of outstanding shares from ```string``` to ```numeric```.
stocks_df[["Price_1_1", "Price_3_23", "Price_4_9", "Shares"]] = \
stocks_df[["Price_1_1", "Price_3_23", "Price_4_9", "Shares"]].apply(pd.to_numeric)
# ## 4.2. Adding Market Cap data and percentage change of stock prices
# ### Adding Market Cap Data
#
# Next, we will add the market cap in the 3 different dates.
stocks_df["Marketcap_1_1"] = stocks_df["Price_1_1"] * stocks_df["Shares"]
stocks_df["Marketcap_3_23"] = stocks_df["Price_3_23"] * stocks_df["Shares"]
stocks_df["Marketcap_4_9"] = stocks_df["Price_4_9"] * stocks_df["Shares"]
# ### Adding percentage change of stock prices
# Percentage Change from January 1st to March 23rd
stocks_df["PercentageChange_3_23_1_1"] = (stocks_df["Price_3_23"] - stocks_df["Price_1_1"]) / stocks_df["Price_1_1"]*100
# Percentage Change from March 23rd to April 9th
stocks_df["PercentageChange_4_9_3_23"] = (stocks_df["Price_4_9"] - stocks_df["Price_3_23"]) / stocks_df["Price_3_23"]*100
# # Percentage Change from January 1st to April 9th
stocks_df["PercentageChange_4_9_1_1"] = (stocks_df["Price_4_9"] - stocks_df["Price_1_1"]) / stocks_df["Price_1_1"]*100
# ## 4.3. Analyzing the data
# ### Change in the total market cap of the S&P 500
sum(stocks_df["Marketcap_3_23"] - stocks_df["Marketcap_1_1"]) / 10**9
sum(stocks_df["Marketcap_4_9"] - stocks_df["Marketcap_3_23"]) / 10**9
sum(stocks_df["Marketcap_4_9"] - stocks_df["Marketcap_1_1"]) / 10**9
# The S&P 500 lost 8.7 trillion USD from January 1st to March 23rd, but it got back 4.6 trillion from March 23rd to April 9th.
# ### Change in the total market cap by sector
(stocks_df.groupby("GICS Sector").sum()["Marketcap_3_23"] - stocks_df.groupby("GICS Sector").sum()["Marketcap_1_1"]).sort_values() / 10**9
# At the 2020 bottom of the S&P 500 (March 23rd), the Information Technology and Financials sectors had the largest drop in total market cap (compared to January 1st) with 1.592 trillion and 1.518 trillion respectively.
(stocks_df.groupby("GICS Sector").sum()["Marketcap_4_9"] - stocks_df.groupby("GICS Sector").sum()["Marketcap_1_1"]).sort_values() / 10**9
# As of April 9th, we can see that the Financials and Industrials sector had the largest drop in total market cap compared to January 1st.
# ### Ranking of companies by percentage change of stock prices
stocks_df.sort_values(by=["PercentageChange_4_9_1_1"])[["Security", "PercentageChange_4_9_1_1"]].head(5)
# We can see from the table above that the companies that are hardest hit are the 3 major cruises companies: Norwegian Cruise Line Holdings, Carnival Corp., Royal Caribbean Cruises Ltd. These companies saw drops in their stock price of over 70%.
print(sum(stocks_df["PercentageChange_4_9_1_1"] < 0))
print(sum(stocks_df["PercentageChange_4_9_1_1"] > 0))
# Only 56 stocks form the 505 saw positive growth from January 1st to April 9th.
# ### Percentage Change of stock priced by sector
stocks_df.groupby("GICS Sector").mean()['PercentageChange_4_9_1_1'].sort_values()
# We can see that the energy sector was the hardest hit with a 49.8% average drop in stock prices.
# Below we can visualize a boxplot of the 11 sectors' percentage change in stock prices from January 1st to April 9th.
plt.figure(figsize=(18, 6))
plt.tick_params('both', labelsize='8')
plt.xticks(rotation=45)
sns.boxplot(x="GICS Sector", y="PercentageChange_4_9_1_1", data=stocks_df)
stocks_df.groupby("GICS Sub Industry").mean()['PercentageChange_4_9_1_1'].sort_values()
print(sum(stocks_df.groupby("GICS Sub Industry").mean()['PercentageChange_4_9_1_1'] < 0))
print(sum(stocks_df.groupby("GICS Sub Industry").mean()['PercentageChange_4_9_1_1'] > 0))
# if we look at the average percentage change in stock prices by Sub Industry; we can see that the travel related industries, Oil & Gas and Department Stores were the hardest hit. 118 of the 138 Sub Industries had their average stock price declined from January 1st to April 9th.
plt.figure(figsize=(18, 6))
plt.tick_params('both', labelsize='8')
stocks_df.groupby("GICS Sub Industry").mean()['PercentageChange_4_9_1_1'].sort_values().plot.bar()
# ### Percentage Change of stock priced by sector (From March 23rd to April 9th)
stocks_df.groupby("GICS Sector").mean()['PercentageChange_4_9_3_23'].sort_values()
# ### Percentage Change of stock priced by Sub Industry (From March 23rf to April 9th)
stocks_df.groupby("GICS Sub Industry").mean()['PercentageChange_4_9_3_23'].sort_values()
# All sectors and sub industries (except Food Retail) saw their average stock price go up from March 23rd to April 9th.
# # Conclusion
# In this tutorial, we learned how to use Python, Google Sheets and Google Finance to collect and analyze stock data in the context of coronavirus pandemic. We're still in the early stages of the pandemic, and we don't know yet what would be the mid to long term effect of the pandemic on both the society and the economy. It would be interesting to review this analysis in the coming weeks to see if the stock market recovers from the 2020 losses.
#
# The source code for this tutorial can be found in this [github repository](https://github.com/adilmoujahid/coronavirus-covid19-stocks-analysis).
# # References
#
# [1] https://en.wikipedia.org/wiki/Google_Finance
| coronavirus-covid19-stocks-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature Extraction Continued
# +
import os
import numpy as np
import pandas as pd
import scipy as sp
import scipy.signal
import scipy.stats
import activity_classifier_utils
# -
# Load the data
fs = 256
data = activity_classifier_utils.LoadWristPPGDataset()
# ### Features
# Time Domain:
# * mean
# * std
# * 5, 10, 15, 20, 25 percentile
# * cross-correlation of all pairs of channels
# * total energy
#
# Frequency Domain:
# * dominant frequency
# * fraction of energy in each 1Hz bin from 0 to 6 Hz
# * spectral entropy of each channel - i'll do
# Low-pass filter at 12 Hz
def LowpassFilter(signal, fs):
b, a = sp.signal.butter(3, 12, btype='lowpass', fs=fs)
return sp.signal.filtfilt(b, a, signal)
# Compute features
def Featurize(accx, accy, accz, fs):
"""A partial featurization of the accelerometer signal.
Args:
accx: (np.array) x-channel of the accelerometer.
accy: (np.array) y-channel of the accelerometer.
accz: (np.array) z-channel of the accelerometer.
fs: (number) the sampling rate of the accelerometer
Returns:
n-tuple of accelerometer features
"""
accx = LowpassFilter(accx, fs)
accy = LowpassFilter(accy, fs)
accz = LowpassFilter(accz, fs)
# The mean of the x-channel
mn_x = np.mean(accx)
# The standard deviation of the x-channel
std_x = np.std(accx)
# The 5th percentile of the x-channel
p5_x = np.percentile(accx, 5)
# The pearson correlation coefficient between the x and y channels
corr_xy = sp.stats.pearsonr(accx, accy)[0]
# The total AC energy of the x-axis
energy_x = np.sum(np.square(accx - np.mean(accx))) # np.var(accx) * len(accx)
# Take an FFT of the signal. If the signal is too short, 0-pad it so we have at least 2046 points in the FFT.
fft_len = max(len(accx), 2046)
# Create an array of frequency bins
freqs = np.fft.rfftfreq(fft_len, 1 / fs)
# Take an FFT of the centered signal
fft_x = np.fft.rfft(accx - np.mean(accx), fft_len)
# The frequency with the most power between 0.25 and 12 Hz
low_freqs = (freqs >= 0.25) & (freqs <= 12)
dominant_frequency_x = freqs[low_freqs][np.argmax(np.abs(fft_x)[low_freqs])]
# The fraction of energy between 2 and 3 Hz in the x-channel
spectral_energy_x = np.square(np.abs(fft_x))
energy_23_x = (np.sum(spectral_energy_x[(freqs >= 2) & (freqs <= 3)])
/ np.sum(spectral_energy_x))
return (mn_x,
std_x,
p5_x,
corr_xy,
energy_x,
dominant_frequency_x,
energy_23_x)
# There are a lot of features, because we have to compute each of these for all channels. I've spared you that effort and put all the features in `activity_classifier_utils.py`. Poke through that file now to see the feature extraction code.
# ### Feature Extraction
# Now we can extract the features for all of our data.
#
# Train on 10 second long non-overlapping windows
window_length_s = 10
window_shift_s = 10
window_length = window_length_s * fs
window_shift = window_shift_s * fs
labels, subjects, features = [], [], []
for subject, activity, df in data:
for i in range(0, len(df) - window_length, window_shift):
window = df[i: i + window_length]
accx = window.accx.values
accy = window.accy.values
accz = window.accz.values
features.append(activity_classifier_utils.Featurize(accx, accy, accz, fs=fs))
labels.append(activity)
subjects.append(subject)
labels = np.array(labels)
subjects = np.array(subjects)
features = np.array(features)
labels
subjects
features
features.shape
# We started with 10 seconds of 256 Hz accelerometer data. That's 2500 samples per channel, and for three channel that's 7500 points. We've successfully reduced these 7500 points to just 55 points while hopefully retaining all the information we need to build a good classifier.
#
# Although we only have 8 subjects of data, we have 611 datapoints because each 10 second window is its own datapoint. However, our datapoints are not independent. Because there's homogeneity in how individuals do an activity, datapoints from the same person might be more similar to each other. We have to keep this in mind when we train and evaluate our model. In the next video we'll use these features to build a random forest model and classify our data.
| AI-for-Healthcare/wearable-data/lesson 4/Feature Extraction Continued.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="KWlC_IGH76Ul" colab_type="code" outputId="b5a5f284-7976-460d-bbf4-7fee92a6fb7e" executionInfo={"status": "ok", "timestamp": 1583661022785, "user_tz": -60, "elapsed": 12166, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 442}
# !pip install --upgrade tables
# !pip install eli5
# + id="QRxNk5pk83RK" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
# + id="fc3BF1yy8saz" colab_type="code" outputId="1e654e09-b3d6-49a9-edf7-71a0dd1be0c5" executionInfo={"status": "ok", "timestamp": 1583661776603, "user_tz": -60, "elapsed": 584, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_car"
# + id="MorrofMN__el" colab_type="code" outputId="4ee4d08f-168e-4023-e3af-9aad8bad0c48" executionInfo={"status": "ok", "timestamp": 1583661780417, "user_tz": -60, "elapsed": 2259, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="MZSQDEWLARBs" colab_type="code" outputId="0bb2f0da-e4ae-4956-ed84-95ee09baf5cb" executionInfo={"status": "ok", "timestamp": 1583661780418, "user_tz": -60, "elapsed": 1306, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 187}
df.columns
# + id="cNerIRveAYpT" colab_type="code" colab={}
# + [markdown] id="dlfVnPt0BjVM" colab_type="text"
# ##Dummy Model
# + id="EkcCkVyPBmM3" colab_type="code" outputId="c2b0dda8-5783-44f6-fdd8-635bec141a66" executionInfo={"status": "ok", "timestamp": 1583661782696, "user_tz": -60, "elapsed": 747, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df.select_dtypes(np.number).columns
# + id="_ubzdrMYBwB-" colab_type="code" outputId="8c44c6ff-7dbd-48e0-9b3f-a2c188005a36" executionInfo={"status": "ok", "timestamp": 1583661783311, "user_tz": -60, "elapsed": 523, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
feats = ['car_id']
X = df[feats].values
y = df['price_value'].values
model = DummyRegressor()
model.fit(X,y)
y_pred = model.predict(X)
mae(y, y_pred)
# + id="W3PIaKYxClgS" colab_type="code" outputId="ca0edd99-d441-40d3-f623-d456ffe9f382" executionInfo={"status": "ok", "timestamp": 1583661784490, "user_tz": -60, "elapsed": 680, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
[x for x in df.columns if 'price' in x]
# + id="c1qmeHN7DNV2" colab_type="code" outputId="736e8f89-dc77-4202-cdde-e46191eaf4f5" executionInfo={"status": "ok", "timestamp": 1583661785920, "user_tz": -60, "elapsed": 596, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
df['price_currency'].value_counts()
# + id="mziXUruVDVQQ" colab_type="code" outputId="f07d2811-d6ec-493d-ddb2-baa3a636d644" executionInfo={"status": "ok", "timestamp": 1583661805481, "user_tz": -60, "elapsed": 656, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
df['price_currency'].value_counts(normalize=True)*100
# + id="Jn38Ouo7EIFc" colab_type="code" outputId="6de997ec-9684-40e6-88fb-666c5d1ed71d" executionInfo={"status": "ok", "timestamp": 1583661807209, "user_tz": -60, "elapsed": 802, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = df[df['price_currency'] != 'EUR']
df.shape
# + id="E-DEhNo2Ek2e" colab_type="code" outputId="8be4a169-9be0-4676-913a-99b4e42aa33f" executionInfo={"status": "ok", "timestamp": 1583661808090, "user_tz": -60, "elapsed": 397, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
df['price_currency'].value_counts()
# + id="jZrPxDMmEsWB" colab_type="code" colab={}
# + [markdown] id="bvJQok_WEx52" colab_type="text"
# ##Features
# + id="JUqcfdHrEzv_" colab_type="code" outputId="7582bfa0-4117-4034-871f-008455719446" executionInfo={"status": "ok", "timestamp": 1583661812040, "user_tz": -60, "elapsed": 531, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 678}
df.head()
# + id="z2BuV1a7E26o" colab_type="code" colab={}
for feat in df.columns:
print(feat)
# + id="Z8CCBfkhFAn6" colab_type="code" outputId="ba89014b-763c-4bf5-c03d-c37a7c3ddccd" executionInfo={"status": "ok", "timestamp": 1583661818716, "user_tz": -60, "elapsed": 497, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 85}
df['param_color'].factorize()
# + id="dh-Nr5_1FK9L" colab_type="code" outputId="d0f3896a-a034-4b84-c79d-6e2730be5859" executionInfo={"status": "ok", "timestamp": 1583661820564, "user_tz": -60, "elapsed": 502, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df['param_color'].factorize()[0]
# + id="N1WCsNH5FX6o" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="9yY9_s-PG3P2" colab_type="code" outputId="1ae2bded-775f-4a3e-8b41-658b0a1ee324" executionInfo={"status": "ok", "timestamp": 1583661824392, "user_tz": -60, "elapsed": 453, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="urRkBEAPHra2" colab_type="code" outputId="4d6946d1-9839-41ec-94cd-403f7af5af67" executionInfo={"status": "ok", "timestamp": 1583661829342, "user_tz": -60, "elapsed": 4700, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgoxbMN4U1L7FQe6UkemC4moyTIQtlW3LgMILvbjQ=s64", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
X = df[cat_feats].values
y = df['price_value'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y , cv=3, scoring='neg_mean_absolute_error')
np.mean(scores)
# + id="V-gyarCYkQB8" colab_type="code" outputId="2c23be5a-51b4-46bf-9b98-9419d27f54af" executionInfo={"status": "ok", "timestamp": 1583661880985, "user_tz": -60, "elapsed": 46610, "user": {"displayName": "Micha\u0142 \u0141\u0105czkowski", "photoUrl": "https://lh3.googleusercontent.com/a-/<KEY>", "userId": "05321865310054149648"}} colab={"base_uri": "https://localhost:8080/", "height": 391}
m = DecisionTreeRegressor(max_depth=5)
m.fit(X, y)
imp = PermutationImportance(m).fit(X, y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="1WMRwfkxku8w" colab_type="code" colab={}
| day3_simple_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 3 Applied Labs
#
# ## Q8
#
# ### (a) Perform a linear regression on the `Auto` dataset with `mpg` as the response and `horsepower` as the predictor - use `summary()` to print the results. Comment on the output.
#
# +
import statsmodels.api as sm
import statsmodels.formula.api as smf
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
# %matplotlib inline
sns.set(style="whitegrid")
# -
auto = sm.datasets.get_rdataset("Auto", "ISLR").data
auto.head()
auto.describe()
# +
# statsmodels doesn't assume there'll be an intercept, so you have to add one!
X_train = sm.add_constant(auto)
# Use the R syntax to perform the regression
model = smf.ols('mpg~horsepower', data=auto)
results = model.fit()
print(results.summary())
# -
# Comment on the results:
# - Is there a relationship between predictor and response?
# - Yes, the F-statistic has a p-value of << 0.001, meaning we can reject the null hypothesis that there is no relationship.
# - How strong is the relationship?
# - "How strong" is difficult, but "how strongly linear" is easier: the linear fit explains 61% of the variance in our data.
# - Is the relationship positive or negative?
# - B1 < 0, so the relationship is negative
# - What is the predicted mpg associated with a horsepower of 98? What are the associated 95% confidence and prediction intervals?
#
# To answer the latter point:
predicted_mpg = results.get_prediction(pd.DataFrame(data=[98], columns=["horsepower"]))
predicted_mpg.summary_frame(alpha=0.05)
# The predicted mpg is ~24.5, the confidence interval is [24.0, 25.0] and the prediction interval is [14.8, 34.1]
# ### (b) Plot the response and the predictor.
fig, ax = plt.subplots(figsize=(15,10))
sns.scatterplot("horsepower", "mpg", data=auto, ax=ax)
# plot the fitted values
b0, b1 = results.params
plt.plot(auto.horsepower, b0+b1*auto.horsepower, lw=3, c="black")
# ### (c) Use the `plot()` function to produce diagnostic plots of the regression. Comment on any issues
#
# We don't have this functionality, but we can plot residual vs horsepower and leverage vs residual plots manually.
fig = plt.figure(figsize=(10,5))
# fig = sm.graphics.plot_regress_exog(results, "horsepower", fig=fig)
g = sns.scatterplot(auto.horsepower, results.resid)
g.set_ylabel("residuals")
fig, ax = plt.subplots(figsize=(15,10))
fig = sm.graphics.plot_leverage_resid2(results, ax=ax)
# From the residuals we can see clear evidence of non-linearity in the data. From the leverage vs residuals, we can see the high-leverage, high-error point 117. We might consider removing this point as an outlier
| Chapter3/Labs/question8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# SPDX-FileCopyrightText: 2019 oemof developer group <<EMAIL>>
#
# SPDX-License-Identifier: MIT
#
# SPDX-License-Identifier: CC-BY-4.0
# # TurbineClusterModelChain example
#
# This example shows you how to calculate the power output of wind farms and wind turbine clusters using the windpowerlib. A cluster can be useful if you want to calculate the feed-in of a region for which you want to use one single weather data point.
#
# Functions that are used in the ModelChain example, like the initialization of wind turbines, are imported and used without further explanation.
# ### Imports and initialization of wind turbines
#
# The import of weather data and the initialization of wind turbines is done as in the ``modelchain_example``. Be aware that currently for wind farm and wind cluster calculations wind turbines need to have a power curve as some calculations do not work with the power coefficient curve.
# +
import pandas as pd
import modelchain_example as mc_e
from windpowerlib import TurbineClusterModelChain, WindTurbineCluster, WindFarm
import logging
logging.getLogger().setLevel(logging.DEBUG)
# +
# Get weather data
weather = mc_e.get_weather_data('weather.csv')
print(weather[['wind_speed', 'temperature', 'pressure']][0:3])
# Initialize wind turbines
my_turbine, e126, dummy_turbine = mc_e.initialize_wind_turbines()
print()
print('nominal power of my_turbine: {}'.format(my_turbine.nominal_power))
# -
# ### Initialize wind farm
#
# To initialize a specific wind farm you need to provide a wind turbine fleet specifying the wind turbines and their number or total installed capacity (in Watt) in the farm. Optionally, you can specify a wind farm efficiency and a name as an identifier.
# specification of wind farm data where turbine fleet is provided in a
# pandas.DataFrame
# for each turbine type you can either specify the number of turbines of
# that type in the wind farm (float values are possible as well) or the
# total installed capacity of that turbine type in W
wind_turbine_fleet = pd.DataFrame(
{'wind_turbine': [my_turbine, e126], # as windpowerlib.WindTurbine
'number_of_turbines': [6, None],
'total_capacity': [None, 12.6e6]}
)
# initialize WindFarm object
example_farm = WindFarm(name='example_farm',
wind_turbine_fleet=wind_turbine_fleet)
# Following, a wind farm with a constant efficiency is defined. A wind farm efficiency can also be dependent on the wind speed in which case it needs to be provided as a dataframe with 'wind_speed' and 'efficiency' columns containing wind speeds in m/s and the corresponding dimensionless wind farm efficiency.
# +
# specification of wind farm data (2) containing a wind farm efficiency
# wind turbine fleet is provided using the to_group function
example_farm_2_data = {
'name': 'example_farm_2',
'wind_turbine_fleet': [my_turbine.to_group(6),
e126.to_group(total_capacity=12.6e6)],
'efficiency': 0.9}
# initialize WindFarm object
example_farm_2 = WindFarm(**example_farm_2_data)
print('nominal power of first turbine type of example_farm_2: {}'.format(
example_farm_2.wind_turbine_fleet.loc[0, 'wind_turbine'].nominal_power))
# -
# ### Initialize wind turbine cluster
#
# Like for a wind farm for the initialization of a wind turbine cluster you can use a dictionary that contains the basic parameters. A wind turbine cluster is defined by its wind farms.
# +
# specification of cluster data
example_cluster_data = {
'name': 'example_cluster',
'wind_farms': [example_farm, example_farm_2]}
# initialize WindTurbineCluster object
example_cluster = WindTurbineCluster(**example_cluster_data)
# -
# ### Use the TurbineClusterModelChain to calculate power output
#
# The TurbineClusterModelChain is a class that provides all necessary steps to calculate the power output of a wind farm or wind turbine cluster.
#
# Like the ModelChain (see [basic example](modelchain_example_notebook.ipynb)) you can use the TurbineClusterModelChain with default parameters as shown in this example for the wind farm or specify custom parameters as done here for the cluster.
# If you use the 'run_model' method first the aggregated power curve and the mean hub height of the wind farm/cluster is calculated, then inherited functions of the ModelChain are used to calculate the wind speed and density (if necessary) at hub height. After that, depending on the parameters, wake losses are applied and at last the power output is calculated.
# power output calculation for example_farm
# initialize TurbineClusterModelChain with default parameters and use
# run_model method to calculate power output
mc_example_farm = TurbineClusterModelChain(example_farm).run_model(weather)
# write power output time series to WindFarm object
example_farm.power_output = mc_example_farm.power_output
# +
# set efficiency of example_farm to apply wake losses
example_farm.efficiency = 0.9
# power output calculation for turbine_cluster
# own specifications for TurbineClusterModelChain setup
modelchain_data = {
'wake_losses_model': 'wind_farm_efficiency', #
# 'dena_mean' (default), None,
# 'wind_farm_efficiency' or name
# of another wind efficiency curve
# see :py:func:`~.wake_losses.get_wind_efficiency_curve`
'smoothing': True, # False (default) or True
'block_width': 0.5, # default: 0.5
'standard_deviation_method': 'Staffell_Pfenninger', #
# 'turbulence_intensity' (default)
# or 'Staffell_Pfenninger'
'smoothing_order': 'wind_farm_power_curves', #
# 'wind_farm_power_curves' (default) or
# 'turbine_power_curves'
'wind_speed_model': 'logarithmic', # 'logarithmic' (default),
# 'hellman' or
# 'interpolation_extrapolation'
'density_model': 'ideal_gas', # 'barometric' (default), 'ideal_gas' or
# 'interpolation_extrapolation'
'temperature_model': 'linear_gradient', # 'linear_gradient' (def.) or
# 'interpolation_extrapolation'
'power_output_model': 'power_curve', # 'power_curve' (default) or
# 'power_coefficient_curve'
'density_correction': True, # False (default) or True
'obstacle_height': 0, # default: 0
'hellman_exp': None} # None (default) or None
# initialize TurbineClusterModelChain with own specifications and use
# run_model method to calculate power output
mc_example_cluster = TurbineClusterModelChain(
example_cluster, **modelchain_data).run_model(weather)
# write power output time series to WindTurbineCluster object
example_cluster.power_output = mc_example_cluster.power_output
# -
# ### Plot results
#
# If you have matplotlib installed you can visualize the calculated power output.
# try to import matplotlib
logging.getLogger().setLevel(logging.WARNING)
try:
from matplotlib import pyplot as plt
# matplotlib inline needed in notebook to plot inline
# %matplotlib inline
except ImportError:
plt = None
# plot turbine power output
if plt:
example_cluster.power_output.plot(legend=True, label='example cluster')
example_farm.power_output.plot(legend=True, label='example farm')
plt.xlabel('Wind speed in m/s')
plt.ylabel('Power in W')
plt.show()
| example/turbine_cluster_modelchain_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <span style="color:Maroon">Crab Age Prediction - Gradient Boosting Model
#
# Import required libraries
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
np.random.seed(0)
os.getcwd()
# Read the csv file data
os.chdir('..\\Data\\')
df = pd.read_csv('data_treated.csv')
df.head()
df.describe()
# ## <span style="color:Maroon">Part 3: Predictive Model
# #### <span style="color:Maroon">Performance Metrics:
# <span style="color:Green">To pick the final model, we shall look at three performance metrics:
#
# <span style="color:Green">__Mean Absolute Error:__ The mean_absolute_error function computes mean absolute error, a risk metric corresponding to the expected value of the absolute error loss or -norm loss. For more details, please refer the below link: https://scikit-learn.org/stable/modules/model_evaluation.html#mean-absolute-error
#
#
# <span style="color:Green">__Mean Squared Error:__ The mean_squared_error function computes mean square error, a risk metric corresponding to the expected value of the squared (quadratic) error or loss. For more details, please refer the below link: https://scikit-learn.org/stable/modules/model_evaluation.html#mean-squared-error
#
# <span style="color:Green">__Rsquare:__ The r2_score function computes the coefficient of determination, usually denoted as R². It represents the proportion of variance (of y) that has been explained by the independent variables in the model. It provides an indication of goodness of fit and therefore a measure of how well unseen samples are likely to be predicted by the model, through the proportion of explained variance. As such variance is dataset dependent, R² may not be meaningfully comparable across different datasets. Best possible score is 1.0 and it can be negative (because the model can be arbitrarily worse). A constant model that always predicts the expected value of y, disregarding the input features, would get a R² score of 0.0. For more details, please refer the below link: https://scikit-learn.org/stable/modules/model_evaluation.html#r2-score
#
# ## <span style="color:Maroon">Model 2: Gradient Boosting Regressor
#
# <span style="color:Green">Gradient boosting is a machine learning technique for regression and classification problems, which produces a prediction model in the form of an ensemble of weak prediction models, typically decision trees.(source: Wikipedia)
#
# <span style="color:Green">Hyper-parameters to be tuned in Gradient Boosting model are:
# 1. n_estimators: number of trees in the foreset
# 2. learning_rate: This determines the impact of each tree on the final outcome. GBM works by starting with an initial estimate which is updated using the output of each tree. The learning parameter controls the magnitude of this change in the estimates
# 3. subsample: The fraction of observations to be selected for each tree. Selection is done by random sampling
# 4. loss: It refers to the loss function to be minimized in each split.
# 5. max_features: max number of features considered for splitting a node
# 6. max_depth: max number of levels in each decision tree
# 7. min_samples_split: min number of data points placed in a node before the node is split
# 8. min_samples_leaf: min number of data points allowed in a leaf node
# Import required libraries
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RandomizedSearchCV
from sklearn import metrics
# Change drirectory to Images
os.chdir("..\\Images\\")
# Divide the dataset into 70:30 for train and test purpose
X_data = df.drop("Age", axis=1)
y_data = df["Age"]
X_train, X_test, y_train, y_test = train_test_split(X_data, y_data, test_size=0.30, random_state=42)
y_train = y_train.ravel()
y_test = y_test.ravel()
# Declare the hyper-parameters for grid search
learning_rate = [0.01, 0.02, 0.05, 0.1]
n_estimators = [int(x) for x in np.linspace(start = 50, stop = 200, num = 10)]
subsample = [0.6, 0.7, 0.8]
loss = ["ls", "huber"]
max_features = ['auto', 'sqrt']
max_depth = [2, 3, 4, 5, 6]
min_samples_split = [5, 10, 20, 50]
min_samples_leaf = [2, 5, 10, 25]
# Random grid
random_grid = {'learning_rate': learning_rate,
'n_estimators': n_estimators,
'subsample': subsample,
'loss': loss,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf}
# GBM Model
gbm = GradientBoostingRegressor()
gbm_random = RandomizedSearchCV(estimator = gbm, param_distributions = random_grid, n_iter = 500, cv = 3,
verbose=2, random_state=42, n_jobs = -1)
# Fit the random search model
gbm_random.fit(X_train, y_train)
best_random_gbm = gbm_random.best_estimator_
best_random_gbm
def evaluate(model, X, y_act, data_str):
y_pred = model.predict(X)
MAE = metrics.mean_absolute_error(y_act, y_pred)
MSE = metrics.mean_squared_error(y_act, y_pred)
r2 = metrics.r2_score(y_act, y_pred)
print ('Model Performance:{}'.format(data_str))
print('Mean Absolute Error: {:0.4f}.'.format(MAE))
print('Mean Square Error = {:0.4f}.'.format(MSE))
print('Rsquare = {:0.4f}'.format(r2))
return MAE, MSE, r2
MAE, MSE, r2 = evaluate(best_random_gbm, X_train, y_train,'Train Sample')
MAE, MSE, r2 = evaluate(best_random_gbm, X_test, y_test,'Test Sample')
# ###### Comments: Lets try manually selecting a model
def GBM_Iter(X_train, y_train, lr, ntrees, subsam, maxfeatures, maxdepth, minsamplesleaf, X_test, y_test):
out_metrics = [None]*6
clf = GradientBoostingRegressor(loss='ls', learning_rate=lr, n_estimators=ntrees, subsample=subsam,
min_samples_leaf=minsamplesleaf, max_depth=maxdepth, random_state=1234,
max_features=maxfeatures, verbose=0)
clf.fit(X_train, y_train)
# Predicting train output
y_pred = clf.predict(X_train)
# Getting train metrics
out_metrics[0] = metrics.mean_absolute_error(y_train, y_pred)
out_metrics[1] = metrics.mean_squared_error(y_train, y_pred)
out_metrics[2] = metrics.r2_score(y_train, y_pred)
# Predicting test output
y_pred = clf.predict(X_test)
# Getting train metrics
out_metrics[3] = metrics.mean_absolute_error(y_test, y_pred)
out_metrics[4] = metrics.mean_squared_error(y_test, y_pred)
out_metrics[5] = metrics.r2_score(y_test, y_pred)
return out_metrics
# +
lr = [0.01, 0.02, 0.05, 0.1]
ntrees = [50, 80, 100]
subsam = [0.6, 0.7, 0.8]
loss = "ls"
maxfeatures = ['auto', 'sqrt']
maxdepth = [2, 3, 4, 5, 6]
minsamplesleaf = [5, 10, 20, 50]
parameters = []
results = []
for i in range(0, len(lr)):
for j in range(0, len(ntrees)):
for k in range(0, len(subsam)):
for l in range(0, len(maxfeatures)):
for m in range(0, len(maxdepth)):
for n in range(0, len(minsamplesleaf)):
parameters.append([lr[i], ntrees[j], subsam[k], maxfeatures[l], maxdepth[m], minsamplesleaf[n]])
results.append(GBM_Iter(X_train, y_train, lr[i], ntrees[j], subsam[k], maxfeatures[l],
maxdepth[m], minsamplesleaf[n], X_test, y_test))
# +
# Convert Parameters to pandas dataframe
parameters = np.array(parameters)
parameters = parameters.reshape(-1,6)
parameters = pd.DataFrame(parameters)
parameters.columns = ["learning_rate","n_estimators", "subsample", "max_features", "max_depth", "min_samples_leaf"]
# Convert results to pandas dataframe
results = np.array(results)
results = results.reshape(-1,6)
results = pd.DataFrame(results)
results.columns = ["Train_MAE", "Train_MSE", "Train_R2", "Test_MAE", "Test_MSE", "Test_R2"]
# Iteration number
ite = np.arange(1,results.shape[0]+1)
# Merge parameters to performance dataframe
Performance_GBM = pd.DataFrame(ite)
Performance_GBM.columns = ["Iteration_No"]
Performance_GBM = Performance_GBM.join(parameters)
Performance_GBM = Performance_GBM.join(results)
Performance_GBM. head()
# +
# Lets plot the three performance meterics for Train and test to select best model
plt.rcParams['figure.figsize'] = [15, 5]
# Plot MAE for train and test
plt.subplot(1,3,1)
plt.plot(Performance_GBM["Iteration_No"], Performance_GBM["Train_MAE"], 'r-', label= "Train MAE")
plt.plot(Performance_GBM["Iteration_No"], Performance_GBM["Test_MAE"], 'b-', label= "Test MAE")
plt.xlabel("Iteration number")
plt.ylabel("MAE")
plt.legend()
# Plot MSE for train and test
plt.subplot(1,3,2)
plt.plot(Performance_GBM["Iteration_No"], Performance_GBM["Train_MSE"], 'r-', label= "Train MSE")
plt.plot(Performance_GBM["Iteration_No"], Performance_GBM["Test_MSE"], 'b-', label= "Test MSE")
plt.xlabel("Iteration number")
plt.ylabel("MSE")
plt.legend()
# Plot r2 for train and test
plt.subplot(1,3,3)
plt.plot(Performance_GBM["Iteration_No"], Performance_GBM["Train_R2"], 'r-', label= "Train R2")
plt.plot(Performance_GBM["Iteration_No"], Performance_GBM["Test_R2"], 'b-', label= "Test R2")
plt.xlabel("Iteration number")
plt.ylabel("R2")
plt.legend()
plt.savefig("Manual_trainedGBM_Perf.png")
plt.show()
# -
# ###### Comments:
# Looking at the graph, we can see that the test R2 peaks at 0.54, MSE at 4.5 and MAE at 1.5. These results are very similar to a random grid search
best_random_gbm
# +
clf = best_random_gbm
print("++++++++++++++++++++++++\n")
MAE1, MSE1, r21 = evaluate(clf, X_train, y_train,'Train Sample')
print("++++++++++++++++++++++++\n")
plt.rcParams['figure.figsize'] = [15, 5]
MAE2, MSE2, r22 = evaluate(clf, X_test, y_test, 'Test Sample')
# -
def plot_ActvsPred(model, X, y, sample_name):
plt.rcParams['figure.figsize'] = [10, 5]
y_hat = model.predict(X)
y_hat = pd.DataFrame(y_hat)
y_hat.columns = ["Predicted"]
y_hat["Actual"] = y
obs_np = np.arange(0, len(y))
y_hat = y_hat.sort_values("Actual", ascending=True)
plt.plot(obs_np, y_hat["Actual"],'r-', label="Actual Age")
plt.plot(obs_np, y_hat["Predicted"], 'b-', label="Predicted Age")
plt.xlabel("observation")
plt.ylabel("Age")
plt.title("Actual Vs Predicted plot for {} sample".format(sample_name))
plt.legend()
plt.savefig(f'GBM_{sample_name}_actual_predicted.png')
plt.plot()
return
plot_ActvsPred(clf, X_train, y_train, "Train")
plot_ActvsPred(clf, X_train, y_train, "Test")
# ###### Comments:
# The GBM model too does poorly on the extreme values. This could be because of data capturing error
# Plot Variable Importance of the features in the final Model
variable_importance = clf.feature_importances_
variables = list(X_train.columns)
importance = pd.DataFrame(variables)
importance.columns = ["Variables"]
importance["Importance"] = variable_importance
importance = importance.sort_values("Importance", ascending = False)
plt.bar(importance["Variables"], importance["Importance"])
plt.xlabel("Variable")
plt.ylabel("Importance")
plt.title("Variable Importance for GBM Model")
plt.xticks(rotation=45)
plt.savefig('VarImp_GBM.png')
plt.show()
# ###### Comments: Shell weight, is the primary variable which explains almost 60% of the output in a GBM model
# #### Partial Dependence Plots:
# ###### Partial dependence plots (PDP) show the dependence between the target response 1 and a set of ‘target’ features, marginalizing over the values of all other features (the ‘complement’ features). Intuitively, we can interpret the partial dependence as the expected target response as a function of the ‘target’ features.
#
# > One-way PDPs tell us about the interaction between the target response and the target feature
# > PDPs with two target features show the interactions among the two features
# > For more details, please refer the below link:
# https://scikit-learn.org/stable/modules/partial_dependence.html
# Import required librarier
from sklearn.inspection import plot_partial_dependence
plt.rcParams['figure.figsize'] = [15, 20]
features = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
plot_partial_dependence(clf, X_train, features, feature_names = list(X_train.columns), grid_resolution=25);
# ###### Comments: PDP plots shows that
# > "Shell Weight" has a positive linear relationship with age in the final selected GBM model.
#
# ### Summary:
# ###### Comparison on Model Performance:
#
# | Model | R2 (Train) | MAE (Train) | MSE (Train) |R2 (Test) | MAE (Test) | MSE (Test) |
# |-----------------------------|:-----------:|:-----------:|:-----------:|:---------:|:----------:|:-----------|
# | Linear Regression | 0.4180 | 1.7887 | 5.7849 | 0.3825 | 1.7622 | 5.7419 |
# | Random Forest Regressor | 0.8013 | 1.0054 | 1.9748 | 0.5535 | 1.4682 | 4.1519 |
# | Gradient Boosting Regressor | 0.6595 | 1.2929 | 3.3847 | 0.5524 | 1.4295 | 4.1618 |
# ###### Comments:
# Based on perfromance metrics $ R^2 $, MAE and MSE for test, the models in order of performance are:
# Gradient Boosting Regressor <--- Random Forest Regressor <--- Linear Regression
# Save the GBM Model
import pickle
os.chdir('..\\Models\\')
filename = "GBM_Regressor.sav"
pickle.dump(best_random_gbm, open(filename, 'wb'))
#
| Codes/05Crab_Age_Prediction_GBMModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
'''Trains a LSTM on the IMDB sentiment classification task.
The dataset is actually too small for LSTM to be of any advantage
compared to simpler, much faster methods such as TF-IDF + LogReg.
Notes:
- RNNs are tricky. Choice of batch size is important,
choice of loss and optimizer is critical, etc.
Some configurations won't converge.
- LSTM loss decrease patterns during training can be quite different
from what you see with CNNs/MLPs/etc.
'''
from __future__ import print_function
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import Dense, Embedding
from keras.layers import LSTM
from keras.datasets import imdb
max_features = 20000
maxlen = 80 # cut texts after this number of words (among top max_features most common words)
batch_size = 32
print('Loading data...')
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
print(len(x_train), 'train sequences')
print(len(x_test), 'test sequences')
print('Pad sequences (samples x time)')
x_train = sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = sequence.pad_sequences(x_test, maxlen=maxlen)
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print('Build model...')
model = Sequential()
model.add(Embedding(max_features, 128))
model.add(LSTM(128, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(1, activation='sigmoid'))
# try using different optimizers and different optimizer configs
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
print('Train...')
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=15,
validation_data=(x_test, y_test))
score, acc = model.evaluate(x_test, y_test,
batch_size=batch_size)
print('Test score:', score)
print('Test accuracy:', acc)
# -
| old/5-1 RNN imdb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
# -
# Install pypcd from this repository
import notebook_helper
# !{notebook_helper.get_install_cmd(quiet=True)}
import pypcd
print(pypcd.__version__)
# Download pcd file from https://github.com/PointCloudLibrary/data/...
# !wget --no-clobber "https://github.com/PointCloudLibrary/pcl/raw/master/test/bunny.pcd"
# +
# load cloud.pcd for visualization
cloud = pypcd.PointCloud.from_path('bunny.pcd')
cloud.pc_data.shape
# +
# set the size of pyplot charts
plt.rcParams['figure.figsize'] = (8, 4)
# Create a figure with a subplot with three axes
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
X, Y, Z = cloud.pc_data['x'], cloud.pc_data['y'], cloud.pc_data['z']
ax.scatter(X, Y, Z);
plt.axis('scaled')
ax.view_init(90, -90)
# -
| examples/notebooks/03_Visualize_3d_pointcloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install bsuite
# !pip install seaborn
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
import os
from bsuite import sweep
import warnings
from scipy.stats import ttest_ind
import seaborn as sns
sns.set_style("whitegrid")
# -
def get_experiments_df(memory_models, envs, seeds, save_dir, column_names):
df_data = []
for seed in seeds:
for memory in memory_models:
for env in envs:
env_id_list = get_sweep_from_bsuite_id(env)
for env_id in env_id_list:
env_id = env_id.replace("/", "-")
path_to_file = f"results/{seed}/{memory}/data/{save_dir}/{env_id}_log.csv"
if not os.path.exists(path_to_file):
warnings.warn(f"Path {path_to_file} doesn't exist. Skipping.")
continue
data = pd.read_csv(
path_to_file,
names=column_names,
index_col=None,
header=0,
)
data["Seed"] = seed
data["Memory"] = memory
data["Experiment"] = int(env_id.split("-")[-1])
data["Environment"] = env_id.split("-")[0]
df_data.append(data)
df = pd.concat(df_data, axis=0, ignore_index=True)
return df
def get_sweep_from_bsuite_id(bsuite_id: str):
return {
"memory_len": sweep.MEMORY_LEN,
"memory_size": sweep.MEMORY_SIZE,
}.get(bsuite_id, [bsuite_id])
# ## Memory Size Experiments
memory_models = ["GTrXL", "Integrated Transformer", "LSTM"]
envs = ["memory_size/4", "memory_size/9"]
seeds = [4, 5, 10, 92, 82, 35, 31, 79, 86, 24, 88, 55, 16, 90, 30, 60, 64, 42, 75, 83, 11, 52, 61, 57, 94, 39, 47, 49, 65, 81]
experiments = get_experiments_df(
memory_models,
envs,
seeds,
save_dir="eval",
column_names=["Episode", "Average Score"],
)
experiments.head()
# ## T-Tests
# ### Memory Size 4
# +
memory_1 = experiments[(experiments['Memory']=="LSTM") & (experiments['Experiment']==4)]
memory_2 = experiments[(experiments['Memory']=="GTrXL") & (experiments['Experiment']==4)]
stat, p = ttest_ind(memory_1['Average Score'], memory_2['Average Score'], equal_var=False)
print("Context Size 5: LSTM vs GTrXL ")
print('T-Stat=%.3f, p=%.3f' % (stat, p))
# +
memory_1 = experiments[(experiments['Memory']=="LSTM") & (experiments['Experiment']==4)]
memory_2 = experiments[(experiments['Memory']=="Integrated Transformer") & (experiments['Experiment']==4)]
stat, p = ttest_ind(memory_1['Average Score'], memory_2['Average Score'], equal_var=False)
print("Context Size 5: LSTM vs Integrated Transformer ")
print('T-Stat=%.3f, p=%.3f' % (stat, p))
# -
# ## Memory Size 9
# +
memory_1 = experiments[(experiments['Memory']=="LSTM") & (experiments['Experiment']==9)]
memory_2 = experiments[(experiments['Memory']=="GTrXL") & (experiments['Experiment']==9)]
stat, p = ttest_ind(memory_1['Average Score'], memory_2['Average Score'], equal_var=False)
print("Context Size 10: LSTM vs GTrXL ")
print('T-Stat=%.3f, p=%.3f' % (stat, p))
# +
memory_1 = experiments[(experiments['Memory']=="LSTM") & (experiments['Experiment']==9)]
memory_2 = experiments[(experiments['Memory']=="Integrated Transformer") & (experiments['Experiment']==9)]
stat, p = ttest_ind(memory_1['Average Score'], memory_2['Average Score'], equal_var=False)
print("Context Size 10: LSTM vs Integrated Transformer ")
print('T-Stat=%.3f, p=%.3f' % (stat, p))
# -
# # Seed Sensitivity
# ## Memory Size 4
mpl.rcParams.update(mpl.rcParamsDefault)
sns.set_style("whitegrid")
memory_models = ["LSTM", "GTrXL", "Universal Transformer", "Integrated Transformer", "ReZero", "Transformer-XL"]
envs = ["memory_size/4"]
seeds = [4, 5, 10, 92, 82, 35, 31, 79, 86, 24, 88, 55, 16, 90, 30, 60, 64, 42, 75, 83, 11, 52, 61, 57, 94, 39, 47, 49, 65, 81]
episodes = [2500, 5000, 10000]
memory_order = list(map(lambda x: x.replace(" ", "\n"), memory_models))
experiments = get_experiments_df(
memory_models,
envs,
seeds,
save_dir="training",
column_names=["Episode", "Average Score", "Loss"],
)
def add_rank(x):
x['Rank'] = range(1, 31)
return x
# +
experiments = experiments[experiments.Episode.isin(episodes)]
experiments = experiments.sort_values(by=["Average Score"], axis=0, ascending=False)
experiments = experiments.groupby(["Episode", "Memory"]).apply(add_rank)
experiments.Memory = experiments.Memory.apply(lambda x: x.replace(" ", "\n"))
g = sns.FacetGrid(experiments, col="Episode", hue="Memory", hue_order=memory_order)
g.map_dataframe(sns.lineplot, x="Rank", y="Average Score")
g.set_axis_labels("Rank", "Mean Return")
g.add_legend(title="")
g.tight_layout()
# /Users/tommakkink/Code/transformers-for-rl/results/plots/training
env_name = envs[0].replace("/", "-")
g.savefig(f"results/plots/training/{env_name}_seed_analysis.png", dpi=300)
# -
plt.show()
# ## Memory Size 9
envs = ["memory_size/9"]
experiments = get_experiments_df(
memory_models,
envs,
seeds,
save_dir="training",
column_names=["Episode", "Average Score", "Loss"],
)
# +
experiments = experiments[experiments.Episode.isin(episodes)]
experiments = experiments.sort_values(by=["Average Score"], axis=0, ascending=False)
experiments = experiments.groupby(["Episode", "Memory"]).apply(add_rank)
experiments.Memory = experiments.Memory.apply(lambda x: x.replace(" ", "\n"))
memory_models = ["LSTM", "GTrXL", "Universal\nTransformer", "Integrated\nTransformer", "ReZero"]
g = sns.FacetGrid(experiments, col="Episode", hue="Memory", hue_order=memory_order)
g.map_dataframe(sns.lineplot, x="Rank", y="Average Score")
g.set_axis_labels("Rank", "Mean Return")
g.add_legend(title="")
g.tight_layout()
# /Users/tommakkink/Code/transformers-for-rl/results/plots/training
env_name = envs[0].replace("/", "-")
g.savefig(f"results/plots/training/{env_name}_seed_analysis.png", dpi=300)
# -
plt.show()
| analysis/Memory-Size-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from model import UniSkip, Encoder
from data_loader import DataLoader
from vocab import load_dictionary
from config import *
from torch import nn
from torch.autograd import Variable
import torch
class UsableEncoder:
def __init__(self, loc="./saved_models/skip-best"):
print("Preparing the DataLoader. Loading the word dictionary")
self.d = DataLoader(sentences=[''], word_dict=load_dictionary('./data/dummy_corpus.txt.pkl'))
self.encoder = None
print("Loading encoder from the saved model at {}".format(loc))
model = UniSkip()
model.load_state_dict(torch.load(loc, map_location=lambda storage, loc: storage))
self.encoder = model.encoder
if USE_CUDA:
self.encoder.cuda(CUDA_DEVICE)
def encode(self, text):
def chunks(l, n):
"""Yield successive n-sized chunks from l."""
for i in range(0, len(l), n):
yield l[i:i + n]
ret = []
for chunk in chunks(text, 100):
print("encoding chunk of size {}".format(len(chunk)))
indices = [self.d.convert_sentence_to_indices(sentence) for sentence in chunk]
indices = torch.stack(indices)
indices, _ = self.encoder(indices)
indices = indices.view(-1, self.encoder.thought_size)
indices = indices.data.cpu().numpy()
ret.extend(indices)
ret = np.array(ret)
return ret
usable_encoder = UsableEncoder()
# -
from tasks.eval_classification import *
eval_nested_kfold(usable_encoder, "MR", loc='./tasks/mr_data/', k=3, seed=1234, use_nb=False)
| Evaluate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Anime Recommendation System using Nearest Neighbors
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from warnings import simplefilter
simplefilter(action='ignore', category=FutureWarning)
# # Load the datasets
#
# * 2 different datasets will be loaded in to dataframes
# * Dataset can be downloaded in https://www.kaggle.com/CooperUnion/anime-recommendations-database
anime = pd.read_csv('datasets/anime.csv')
rating = pd.read_csv('datasets/rating.csv')
# ### anime.csv
#
# * anime_id - myanimelist.net's unique id identifying an anime.
# * name - full name of anime.
# * genre - comma separated list of genres for this anime.
# * type - movie, TV, OVA, etc.
# * episodes - how many episodes in this show. (1 if movie).
# * rating - average rating out of 10 for this anime.
# * members - number of community members that are in this anime's "group".
#
# ### rating.csv
#
# * user_id - non identifiable randomly generated user id.
# * anime_id - the anime that this user has rated.
# * rating - rating out of 10 this user has assigned (-1 if the user watched it but didn't assign a rating).
print('anime.csv (shape):',anime.shape)
print('rating.csv (shape):',rating.shape)
anime.head()
rating.head()
# +
# checking for null values
anime.isnull().sum()
# +
# filling all anime without rating with 0
anime.fillna({'rating':0},inplace=True)
# -
# Exploratory data analysis is on the other notebook. (Anime Recommendation using Pearson r correlation.)
# # Collaborative Filtering using Nearest Neighbors
#
# <br>
#
# ```
# * In this recommendation system, we will be utilizing the collaborative filtering technique.
# * By using this technique, the system will recommend anime based on the nearest rating between the ratings of
# user's anime and the ratings of other anime.
# * For example, I watched 10 anime and gave each of them a rating. Now, my friend watched an anime from my
# anime list and now asks me to recommend three anime. With that, I will recommend three anime with closest
# rating to the rating I gave for the anime that my friend watched.
# ```
#
# ### Process
#
# <br>
#
# ```
# * Remove anime with low count of ratings and users who gave low count of ratings
# * Construct Rating Matrix
# * Convert rating matrix to csr matrix to save memory
# * Fit the csr rating matrix into nearest neighbor
# * Retrieve ten nearest neighbor
# * Output ten recommended anime
# ```
#
# <br>
#
# 
# ### Remove anime with low count of ratings and users who gave low count of ratings
#
# * We will only consider popular anime (rating count over 250) and users who gave lots of rating on different anime (>100)
anime_rating_count = rating.groupby(by='anime_id').count()['rating'].reset_index().rename(columns={'rating':'rating_count'})
anime_rating_count['rating_count'].describe()
filtered_anime = anime_rating_count[anime_rating_count['rating_count']>250]
# +
# anime with over 250 rating count
filtered_anime.head()
# -
user_rating_count = rating.groupby(by='user_id').count()['rating'].reset_index().rename(columns={'rating':'rating_count'})
user_rating_count['rating_count'].describe()
# +
# users who gave over 100 ratings to different anime
filtered_user = user_rating_count[user_rating_count['rating_count']>100]
# -
filtered_user.head()
filtered_rating_anime = rating[rating['anime_id'].isin(filtered_anime['anime_id'])]
filtered_rating = filtered_rating_anime[filtered_rating_anime['user_id'].isin(filtered_user['user_id'])]
# +
# this dataset now contains popular anime and users wth high rating counts
filtered_rating.head()
# -
# ### Construct Rating Matrix
#
# * We will construct a matrix by using pivot table wherein anime id will be indexes and user id in columns
# +
# we can see that most of the values are zero since most of the users does not have ratings for every anime
rating_matrix = filtered_rating.pivot_table(index='anime_id',columns='user_id',values='rating').fillna(0)
print(rating_matrix.shape)
rating_matrix.head()
# -
# ### Convert rating matrix to csr matrix to save memory
from scipy.sparse import csr_matrix
csr_rating_matrix = csr_matrix(rating_matrix.values)
print(csr_rating_matrix)
# ### Fit the matrix into nearest neighbor
#
# * We are using unsupervised algorithm nearest neighbor.
# * This algorithm will find k nearest data point which will be the recommended anime to watch.
# * We will also use cosine similarity as the metric for the algorithm.
# +
from sklearn.neighbors import NearestNeighbors
recommender = NearestNeighbors(metric='cosine')
# fit the csr matrix to the algorithm
recommender.fit(csr_rating_matrix)
# -
# ### Retrieve ten nearest neighbors
# +
# getting the anime_id of the user's anime
user_anime = anime[anime['name']=='Bleach']
user_anime
# +
user_anime_index = np.where(rating_matrix.index==int(user_anime['anime_id']))[0][0]
# this index is from rating matrix not from the anime dataset
user_anime_index
# +
# getting the ratings based on the index
user_anime_ratings = rating_matrix.iloc[user_anime_index]
user_anime_ratings
# +
# we need to convert this into 2d array (with only 1 row) since the algorithm does not accept 1d array
user_anime_ratings_reshaped = user_anime_ratings.values.reshape(1,-1)
user_anime_ratings_reshaped
# +
# the ratings will be plotted and will return 11 indices and distances of nearest neighbors
# note that these indices are based on the indices of rating matrix
distances, indices = recommender.kneighbors(user_anime_ratings_reshaped,n_neighbors=11)
# +
# indices of nearest neighbors (based on rating matrix)
indices
# +
# distances of nearest neighbors to the user's anime
distances
# -
# ### Output ten recommended anime
# +
# the returned indices will be used to get anime id(index) on rating matrix
# these indices are the nearest neighbors
# we are excluding the first element since the first nearest neighbor is itself
nearest_neighbors_indices = rating_matrix.iloc[indices[0]].index[1:]
# -
nearest_neighbors = pd.DataFrame({'anime_id': nearest_neighbors_indices})
pd.merge(nearest_neighbors,anime,on='anime_id',how='left')
# # Saving the model
import pickle
pickle.dump(recommender,open('output/nearest_neighbor_recommender.pickle','wb'))
# +
from scipy.sparse import save_npz, load_npz
import json
csr_rating_matrix_open = load_npz('output/csr_rating_matrix.npz')
with open('output/rating_matrix_anime_id.json') as f:
anime_id = json.load(f)
with open('output/rating_matrix_user_id.json') as f:
user_id = json.load(f)
# -
rating_matrix_open = pd.DataFrame(csr_rating_matrix_open.toarray().T,index=anime_id['anime_id'],columns=user_id['user_id'])
rating_matrix.equals(rating_matrix_open)
| model/anime_recommendation_nearest_neighbors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from libtiff import TIFF
import cv2
import numpy as np
import os
import segmentation_models as sm
import natsort
import random
import imgaug.augmenters as iaa
import imgaug as ia
from keras.models import load_model
from tqdm import tqdm
from PIL import Image
import keras.backend as K
import time
import imageio
import configparser
# +
config = configparser.ConfigParser()
config.read('segmentation_inference.ini')
MODEL_WEIGHTS_PATH = config['model_params']['model_weights_path'] #Path to model weights
PATH = config['model_params']['images_path'] # Path to folder with test_images
DEST_PATH = config['model_params']['save_path'] #Path to save the converted tiff outputs from the model
# -
MODEL_WEIGHTS_PATH
model = sm.Unet("resnet50",input_shape=(None,None,3),classes=5,activation="sigmoid",encoder_freeze = True)
model.load_weights(MODEL_WEIGHTS_PATH)
def convert_numpy_to_tiff(tiff_npy, file_name, save_path):
if not os.path.exists(save_path):
os.mkdir(save_path)
list_temp = []
for i in range(0,5):
tiff_npy[:,:,i][tiff_npy[:,:,i]==1]=255
list_temp.append(tiff_npy[:,:,i])
tiff = np.asarray(list_temp,dtype=np.uint8)
imageio.mimwrite(os.path.join(save_path,os.path.splitext(file_name)[0]+".tif"),tiff)
# +
def perform_inference(model,path,dest_path):
for files in tqdm(os.listdir(path)):
list_buffer = []
img = cv2.imread(os.path.join(PATH,files))
shape = img.shape
img = cv2.resize(img,(512,512))
mask_pred = model.predict(np.expand_dims(img,axis=0))
mask_pred = mask_pred[0]
for i in range(0,5):
mask_pred[:,:,i][mask_pred[:,:,i]>0.5]=1
mask_pred[:,:,i][mask_pred[:,:,i]<=0.5]=0
list_buffer+=[cv2.resize(mask_pred[:,:,i],(shape[1],shape[0]),interpolation=cv2.INTER_NEAREST)]
img_mask = cv2.merge((list_buffer[0],list_buffer[1],list_buffer[2],list_buffer[3],list_buffer[4]))
convert_numpy_to_tiff(img_mask,files,dest_path)
# -
perform_inference(model,PATH,DEST_PATH)
| Segmentation/Unet_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import matplotlib.pyplot as plt
import numpy as np
from matplotlib import animation
import scipy.integrate as integrate
# +
t = 1
fig, ax = plt.subplots()
theta = np.linspace(0, 2*np.pi, 1000)
glowa_x = np.cos(theta)
glowa_y = np.sin(theta)
grzywka_x_1 = np.linspace(-0.3,0.3,1000)
grzywka_y_1 = np.linspace(0.95,0.7,1000)
grzywka_x_2 = np.linspace(0.3,0.4,1000)
grzywka_y_2 = np.linspace(0.7,0.9,1000)
oko_x = glowa_x / 7
oko_y = glowa_y/7 + 0.3
zrenica_x = glowa_x / 100
zrenica_y = glowa_y / 100 + 0.3
wasy_x = np.linspace(-0.1,0.1,1000)
wasy_y = -0.1*np.ones(1000)*np.cos(5*t)-0.2
wasy = plt.plot(wasy_x,wasy_y,"k-",linewidth=30)[0]
nos_x_1 = np.linspace(-0.2,0.1,1000)
nos_y_1 = np.linspace(0,0.4,1000)
nos_x_2 = np.linspace(-0.2,0,1000)
nos_y_2 = -0.01*np.ones(1000)
usmiech_x = np.linspace(-0.4,0.4,1000)
usmiech_y = -0.4 + t*(usmiech_x)**2
usmiech = plt.plot(usmiech_x,usmiech_y, "k-", linewidth=5)[0]
plt.plot(glowa_x,glowa_y,"k-",linewidth=5)
plt.plot(oko_x-0.4,oko_y,"k-",linewidth=5)
plt.plot(oko_x+0.4,oko_y,"k-",linewidth=5)
#plt.plot(usmiech_x,usmiech_y,"k-",linewidth=5)
#plt.plot(wasy_x,wasy_y,"k-",linewidth=20)
plt.plot(grzywka_x_1,grzywka_y_1,"k-",linewidth=5)
plt.plot(grzywka_x_2,grzywka_y_2,"k-",linewidth=5)
plt.plot(zrenica_x-0.4,zrenica_y,"k-",linewidth=5)
plt.plot(zrenica_x+0.4,zrenica_y,"k-",linewidth=5)
plt.plot(nos_x_1,nos_y_1,"k-",linewidth=5)
plt.plot(nos_x_2,nos_y_2,"k-",linewidth=5)
plt.xlim(-2,2)
plt.ylim(-2,2)
def animate(t):
usmiech_y = -0.6 + t*(usmiech_x)**2
usmiech.set_data(usmiech_x,usmiech_y)
wasy_y = -0.1*np.ones(1000)*np.cos(2*t)-0.2
wasy.set_data(wasy_x,wasy_y)
return [usmiech],[wasy]
czas = np.cos(np.linspace(0, 2*np.pi, 120))
ani = animation.FuncAnimation(fig, animate, frames = czas, interval=1)
plt.show()
# -
# # Lorenz
# +
def pochodna(y,x):
return y
y_0 = 1
x = np.linspace(0,10,1000)
y = integrate.odeint(pochodna,y_0,x)
plt.plot(x,y)
plt.show()
# -
# $$ \dot{y} = f(y(x),x) $$
# +
from mpl_toolkits.mplot3d import Axes3D
sigma = 10
beta = 8/3
rho = 28
r0 = np.array([-0.2,-0.5,-0.6])
r1 = np.array([0.3,0.4,0.5])
t = np.linspace(0,100,100000)
def lorenz(r, t):
x, y, z = r
xdot = sigma*(y-x)
ydot = x*(rho-z)-y
zdot = x*y - beta*z
return np.array([xdot,ydot,zdot])
r = integrate.odeint(lorenz,r0,t)
x, y, z = r.T
x2, y2, z2 = integrate.odeint(lorenz,r1,t).T
fig = plt.figure()
ax = fig.add_subplot(111,projection = '3d')
ax.plot(x,y,z,"b-",alpha = 0.5)
ax.plot(x2,y2,z2,"r-",alpha = 0.5)
plt.show()
# +
fig = plt.figure()
ax = fig.add_subplot(111,projection = '3d')
linia1, = ax.plot(x[:0],y[:0],z[:0],"b-",alpha = 0.5)
kulka1, = ax.plot(x[0:1],y[0:1],z[0:1],"bo",alpha=0.5)
linia2, = ax.plot(x2[:0],y2[:0],z2[:0],"r-",alpha = 0.5)
kulka2, = ax.plot(x2[0:1],y2[0:1],z2[0:1],"ro",alpha=0.5)
#ax.plot(x2,y2,z2,"r-",alpha = 0.5)
def animate(i):
linia1.set_data(x[:i+1],y[:i+1])
linia1.set_3d_properties(z[:i+1])
kulka1.set_data(x[i:i+1],y[i:i+1])
kulka1.set_3d_properties(z[i:i+1])
linia2.set_data(x2[:i+1],y2[:i+1])
linia2.set_3d_properties(z2[:i+1])
kulka2.set_data(x2[i:i+1],y2[i:i+1])
kulka2.set_3d_properties(z2[i:i+1])
return [linia1, kulka1, linia2, kulka2]
ax.set_xlim(x.min(),x.max())
ax.set_ylim(y.min(),y.max())
ax.set_zlim(z.min(),z.max())
anim = animation.FuncAnimation(fig, animate, frames=np.arange(0,len(x),25), interval=1)
plt.show()
# -
| 2016_07_14_drugie_czesc_2_Kabat_style.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:LYH]
# language: python
# name: conda-env-LYH-py
# ---
# + code_folding=[]
import random
import hparams
import torch
import os
import pickle as pkl
import matplotlib.pyplot as plt
# %matplotlib inline
targets = []
alignments = []
for file in os.listdir(f'{hparams.teacher_dir}/targets'):
file_name = file[:-4]
with open(f'{hparams.teacher_dir}/targets/{file_name}.pkl', 'rb') as f:
targets.append( (file_name, pkl.load(f)) )
with open(f'{hparams.teacher_dir}/alignments/{file_name}.pkl', 'rb') as f:
alignments.append( (file_name, pkl.load(f)) )
# +
idx = random.choice(range(len(alignments)))
print(f'{targets[idx][0]}')
fig, axes = plt.subplots(2, 1, figsize=(16,8))
axes[0].imshow(targets[idx][1].numpy(),
origin='lower',
aspect='auto')
axes[1].imshow(alignments[idx][1].numpy().T,
origin='lower',
aspect='auto')
plt.show()
| data_inspection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manipulation des données de nuScenes avec panda
# ### Nuscenes:
# https://github.com/nutonomy/nuscenes-devkit
#
# https://www.nuscenes.org/
# # Introduction
# Notebook de présentation et de découverte du jeu de données NuScenes. NuScenes est un jeu de données qui sert au développement de véhicules autonomes. C'est une grande base de données (+400 go) qui contient:
# - Des informations intrinsèques,informations sur le véhicule, comme la vitesse, l'angle du volant, la pression sur les pédales d'accélération et de frein, .... Ces informations sont accéssible en téléchargeant l'extension CAN Bus en plus du jeu de données de base.
# - Des informations extrinsèques sur l'envrionnement autour du véhicule, détection de tout les objets présents autour (Voiture, camion, piéton, plot de chantier...) qui possèdent une box et une position comme représentation. Ces informations sont extraites à partir de caméra, radar et lidar présent autour du véhicule. Il s'agit d'informations réelles capturer pendant des sessions dans plusieurs villes.
# - Un ensemble de carte où se déplace tout ces objets, sur ces cartes on peut parcourir les routes connaitres leur type, connaitre la signalisation (stop, feu), ... Je ne me suis pas occupé de cette partie car elle est assez compliqué et ne m'interresse pas pour le stage.
# Il y a un ensemble de tutoriel déjà fournies par nuScenes à cette adresse (https://github.com/nutonomy/nuscenes-devkit/tree/master/python-sdk/tutorials) basics_tutorial et can_bus_tutorial pour avoir des informations complémentaires.
# Il y a deux versions pour NuScenes, la version normal (+400go) et il existe une version mini (4go) contenant 10 scènes.
# Il est possilble d'utiliser la version normal sans tout télécharger. Sur le site dans la partie "Trainval", on a un zip metadata de 400 mo et il suffit pour tout faire. Si par contre, on veut visualiser des scènes (caméra et/ou lidar) il faudra télécharger les zip de 30 go correspondants à 100 scènes chacun. J'en ai téléchargé 3 personnellement ce qui me donner un accès à 300 scènes.
#
# Attention si vous décidez de charger le jeu complet (qu'importe d'avoir téléchargé le jeu en entier), cela risque de mettre du temps sur une machine pas suffisament puissante (je mets 6/15 min c'est aléatoire sur mon pc portable pour charger i5 et 8go de ram), de plus je conseille d'avoir au moins 16 go pour pouvoir l'utiliser sans avoir trop de ralentissement, avec 8go j'ai eu beaucoup de ralentissement.
#
# Mais si vous ne pouvez pas, ce n'est pas grave, car j'ai enregistré les données que j'utilise sous forme de csv pour ce que cela soit plus rapide (pour le prochain notebook).
# # Import librairies et chargement de NuScenes
# +
# %matplotlib inline
from nuscenes.nuscenes import NuScenes
from nuscenes.can_bus.can_bus_api import NuScenesCanBus
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
#nusc = NuScenes(version='v1.0-mini', dataroot='../data/sets/nuscenes')
#nusc = NuScenes(version='v1.0-mini', dataroot='G:/repertoire_g/data/sets/nuscenes')
#nusc_can = NuScenesCanBus(dataroot='G:/repertoire_g/data/sets/nuscenes')
# Trainval ou mini suivant la version que l'on souhaite utiliser
nusc = NuScenes(version='v1.0-trainval', dataroot='D:/Utilisateurs/Alexandre/Repertoire_D/nuscenes/v1.0-trainval01')
nusc_can = NuScenesCanBus(dataroot='../data/sets/nuscenes')
# -
# # Prise en main de l'extension CAN
# Tout d'abord, faisons un rendu de la scène. `field2token` renvoie le token associé au nom de la scène, plus généralement, elle renvoie une liste de token de la classe en premier paramètre, dont un des attributs (second paramètre) a pour valeur le dernier paramètre, elle est très pratique.
scene_name = 'scene-0061'
my_scene_token = nusc.field2token('scene', 'name', scene_name)[0]
nusc.render_scene_channel(my_scene_token, 'CAM_FRONT')
# +
# Decommenter si vous utilisez la version mini
#nusc.list_scenes()[
# -
# On prends la scène 61 et je récupère les informations suivantes (vitesse, angle du volant...) à partir de l'extension CAN et je les mets dans un dataframe.
# `get_messages` renvoie une liste de dictionnaire contenant pour chaque enregistrement (toutes les demi-secondes de la scène) les informations internes du véhicule à partir du nom de scène donnée.
scene_name = 'scene-0061'
dic_scene = nusc_can.get_messages(scene_name,'vehicle_monitor')
dic_scene[0]
features = ["vehicle_speed","steering","throttle","left_signal","right_signal"]
df_scene = pd.DataFrame.from_dict(dic_scene)[features]
df_scene
# # Essayer d'apprendre quand mettre le clignotant
# On va maitenant faire un essai pour prendre en main le dataset, avec seulement la vitesse, l'accélération et l'inclinaison du volant. Je ne pense pas qu'il soit possible de prévoir quand mettre un clignotant car il manque certaines informations (trajectoire notamment) mais cela permettra de manipuler NuScenes avant d'attaquer la suite.
# ## 1/ Prétraitement des données
# Pour récupérer la liste des scènes présentes dans le jeu de donnnées, on peut récupérer l'attribut scene de l'instance NuScene.Par contre, si on utilise le mini jeu de données, on aura que 10 scènes.
all_scene = [ s["name"] for s in nusc.scene ]
print(len(all_scene))
#
# Mets des 1 tout le temps pour le clignotant au lieu d'une alternance par défaut
def fill_signal(df,signal):
i = 0
index = df.columns.get_loc(signal)
while i < len(df):
while i < len(df) and df[signal][i] != 1:
i += 1
while i < len(df) and sum(df[signal][i:i+4]) >= 1:
df.iat[i,index] = 1
i += 1
return df
df_scene2 = df_scene.copy()
fill_signal(df_scene2,'right_signal')
df_scene2
# On créer maintenant une nouvelle colonne à notre dataframe qui contient les valeurs du clignotant 0(rien), 1(clignotant gauche), 2(clignotant droit).
#Ajoute une nouvelle colonne où les valeurs sont: 0(rien), 1(clignotant gauche), 2(clignotant droit)
#Plus pratique que "fill_signal" car on aura une seul colonne Y pour l'apprentissage
def add_signal_column_v2(df):
df = fill_signal(df,'left_signal')
df = fill_signal(df,'right_signal')
i = 0
tab = []
while i < len(df):
if df["right_signal"][i] == 1:
tab += [2]
elif df["left_signal"][i] == 1:
tab += [1]
else:
tab += [0]
i += 1
df["signal"] = tab
return df
# On récupère les scènes qui n'ont pas de données dans CAN Bus et qui seront à exclure.
blackint = nusc_can.can_blacklist
blacklist = [ "scene-0"+ str(i) for i in blackint]
print(blacklist)
print( "%s" in all_scene)
# On parcours maintenant l'ensemble des scènes et on récupère leur dataframe avec la nouvelle colonne ajoutée
# +
tab = []
for s in all_scene:
if s not in blacklist and s not in ["scene-0419","scene-0420","scene-0040","scene-0037","scene-0136"
,"scene-0137"] :
dic_scene = nusc_can.get_messages(s,'vehicle_monitor')
features = ["vehicle_speed","steering","throttle","left_signal","right_signal"]
df_scene = pd.DataFrame.from_dict(dic_scene)[features]
add_signal_column_v2(df_scene)
tab += [df_scene]
df_total = pd.concat(tab)
print(len(all_scene))
print(df_total)
print(df_total.describe())
df_total.to_csv("./data/cligno_2407bis2.csv")
# -
# On peut regarde la corrélation entre les attributs pour déjà voir si cela peut correspondre
# Note: en chargeant la version mini les résultats ne seront pas les mêmes car il y a beaucoup moins de données.
df_total.corr()
# ## 2/ Apprentissage
# Passons maintenant à la dernière étape, l'apprentissage. On commence d'abord par importer les librairies dont on aura besoin, puis on sépare le jeu de données en 2, un jeu d'apprentissage (80%) et un jeu de test (20%).
from sklearn.model_selection import train_test_split
from sklearn import svm, neighbors
from sklearn.ensemble import RandomForestClassifier
import random
features = ["vehicle_speed","steering","throttle"]
X = df_total[features]
y = df_total["signal"]
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2, random_state = 1)
# On va faire de l'apprentissage supervisé par classification avec deux modèles (k plus proches voisins et Random forest ).
# ### 2/1/ K plus proches voisins et Random forest sur une seule variable signal a 3 valeurs
# +
model = neighbors.KNeighborsClassifier()
model.fit(X_train,y_train)
print(model.score(X_test,y_test))
model2 = RandomForestClassifier(n_estimators=100,random_state = 42)
model2.fit(X_train,y_train)
model2.score(X_test,y_test)
# -
# On obtient un score d'à peu près 0.8. C'est un score moyen mais c'est plutôt étonnant car je pensais avoir un score très faible car ces informations d'après moi ne suffisent pas pour prédire quand mettre un clignotant.
#
# On peut se poser alors comme question si le score reflète-t-il la vérité? Peut-être que dans la majorité de scènes dans le jeu de test le clignotant n'est pas utilisé et donc le score peut-être boosté? Vérifions ça.
# ### 2/2 Vérification du modèle
# Matrice de confusion:
from sklearn.metrics import confusion_matrix
print(confusion_matrix(y_test,model.predict(X_test)))
print(confusion_matrix(y_test,model2.predict(X_test)))
# Résultat de la matrice de confusion sur le jeu de test (170 scènes, 6625 échantillons) avec k voisins (gauche) et une forêt aléatoire (droite):
#
# [4370 176 207].....[4359 163 231]
#
# [ 467 330 22]........[ 440 340 39]
#
# [ 572 48 447]........[ 541 41 485]
#
# Les bonnes prédictions sont sur la diagonale, sinon c'est une fausse prédicition.
# On peut voir que ce n'est pas bon du tout, la majorité des cas (première ligne) correspond à aucun clignotant, fausse le score.
# On voit bien sur les deux lignes suivantes la majorité des prédicitons ne sont pas sur la diagonale, donc qu'il s'agit de mauvaises prédictions.
#
# J'obtiens des résultats quasi-identique en changeant l'attribution des valeurs dans la colonne signal (clignotant gauche = 2 au lieu de 1).
# ## Ancienne version
# Version utilisé lors de la présentation le 22 Juin, elle comportait des erreurs, notamment sur le remplissage des colonnes, ce qui devait je pense fausser le score.
#Ajoute une nouvelle colonne où les valeurs sont: 0(rien), 1(clignotant gauche), 2(clignotant droit)
#Plus pratique que "fill_signal" car on aura une seul colonne Y pour l'apprentissage
def add_signal_column(df,signal):
i = 0
tab = []
while df[signal][i] != 1:
tab += [0]
i += 1
while i < len(df) and sum(df[signal][i:i+4]) >= 1:
if signal == "right_signal":
tab += [2]
else:
tab += [1]
i += 1
while i < len(df):
tab += [0]
i += 1
df["signal"] = tab
return df
# +
tab = []
for s in all_scene:
if s not in blacklist and s not in ["scene-0419","scene-0420","scene-0040","scene-0037","scene-0136"
,"scene-0137"] :
dic_scene = nusc_can.get_messages(s,'vehicle_monitor')
features = ["vehicle_speed","steering","throttle","left_signal","right_signal"]
df_scene = pd.DataFrame.from_dict(dic_scene)[features]
if df_scene["left_signal"].any():
#df_scene = fill_signal(df_scene,"left_signal")
new_df = add_signal_column(df_scene,"left_signal")
#print(new_df)
if df_scene["right_signal"].any():
#df_scene = fill_signal(df_scene,"right_signal")
new_df = add_signal_column(df_scene,"right_signal")
#print("right",new_df)
tab += [new_df]
df_total = pd.concat(tab)
print(len(all_scene))
print(df_total)
print(df_total.describe())
df_total.to_csv("./data/cligno_2307bis.csv")
# -
from sklearn.model_selection import train_test_split
from sklearn import svm, neighbors
from sklearn.ensemble import RandomForestClassifier
import random
features = ["vehicle_speed","steering","throttle"]
X = df_total[features]
y = df_total["signal"]
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2, random_state = 1)
# ### 2/1/ K plus proches voisins et Random forest sur une seule variable signal a 3 valeurs
# +
model = neighbors.KNeighborsClassifier()
model.fit(X_train,y_train)
print(model.score(X_test,y_test))
model2 = RandomForestClassifier(n_estimators=100,random_state = 42)
model2.fit(X_train,y_train)
model2.score(X_test,y_test)
# -
# On obtient un score d'à peu près 0.8, voir 0.87 avec la forêt aléatoire.
#
# Mais le score reflète-t-il la vérité? Peut-être que dans la majorité de scènes dans le jeu de test le clignotant n'est pas utilisé et donc le score peut-être boosté? Vérifions ça.
from sklearn.metrics import confusion_matrix
print(confusion_matrix(y_test,model.predict(X_test)))
print(confusion_matrix(y_test,model2.predict(X_test)))
# Résultat de la matrice de confusion sur le jeu de test (170 scènes, 6625 échantillons) avec <NAME> (gauche) et une forêt aléatoire (droite):
# - [3402, 210, 338]-----[3626 145 179]
# - [ 288, 549, 63]--------[169 688 43]
# - [ 385, 61, 1329]------[ 257 50 1468]
#
# <NAME>:
# Résultats assez mitigés le taux de d'erreur pour le clignotant gauche est de 0.5 et du clignotant droit 0.25.
#
# Forêt aléatoire:
# Résultats plutôt bon, taux d'erreur pour le clignotant gauche est de 0.30 et du clignotant droit 0.15.
#
# J'obtiens des résultats quasi-identique en changeant l'attribution des valeurs dans la colonne signal (clignotant gauche = 2 au lieu de 1).
# On peut par ailleurs noter qu'il y a un double des cas où le clignotant droite est activé par rapport au clignotant gauche, cela peut être l'une des raisons qui explique la différence de score.
# # Conclusion
# J'ai présenté dans ce notebook quelques outils afin de manipuler nuScenes ainsi qu'une démonstration en essayant de prédire quand mettre le clignotant.
# Cela conclut la première partie des notebooks sur nuScenes.
| nuscenes_decouverte.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
columns = ['Year','Incumbent Party Candidate','Other Candidate','Incumbent Party Vote Share']
data = [[1952,"Stevenson","Eisenhower",44.6],
[1956,"Eisenhower","Stevenson",57.76],
[1960,"Nixon","Kennedy",49.91],
[1964,"Johnson","Goldwater",61.34],
[1968,"Humphrey","Nixon",49.60],
[1972,"Nixon","McGovern",61.79],
[1976,"Ford","Carter",48.95],
[1980,"Carter","Reagan",44.70],
[1984,"Reagan","Mondale",59.17],
[1988,"Bush, Sr.","Dukakis",53.94],
[1992,"Bush, Sr.","Clinton",46.55],
[1996,"Clinton","Dole",54.74],
[2000,"Gore","Bush, Jr.",50.27],
[2004,"Bush, Jr.","Kerry",51.24],
[2008,"McCain","Obama",46.32],
[2012,"Obama","Romney",52.00],
[2016,"Clinton","Trump",48.2]]
votes = pd.DataFrame(data=data, columns=columns)
# +
columns = ['Year','Average Recent Growth in Personal Incomes']
data = [[1952,2.40],
[1956,2.89],
[1960, .85],
[1964,4.21],
[1968,3.02],
[1972,3.62],
[1976,1.08],
[1980,-.39],
[1984,3.86],
[1988,2.27],
[1992, .38],
[1996,1.04],
[2000,2.36],
[2004,1.72],
[2008, .10],
[2012, .95],
[2016, .10]]
growth = pd.DataFrame(data=data, columns=columns)
# +
# adding two extra features
# source: FRED, month before election
columns = ['Year','Civilian Unemployment Rate']
data = [[1952,2.80],
[1956,4.30],
[1960,6.10],
[1964,4.80],
[1968,3.40],
[1972,5.30],
[1976,7.70],
[1980,7.50],
[1984,7.40],
[1988,5.30],
[1992,7.40],
[1996,5.40],
[2000,3.90],
[2004,5.40],
[2008,6.80],
[2012,7.90],
[2016,4.70]]
unemployment = pd.DataFrame(data=data, columns=columns)
# +
# Problem uploading data from desktop on mac?
# gdp_change = pd.read_csv('desktop/download.csv', header=None)
# gdp_change.head()
# +
# source: BEA, National Income and product accounts, table 1.17.1
columns = ['Year','GDP Change']
data = [[1952,4.10],
[1956,2.10],
[1960,2.60],
[1964,5.80],
[1968,4.90],
[1972,5.30],
[1976,5.40],
[1980,-0.30],
[1984,7.20],
[1988,4.20],
[1992,3.50],
[1996,3.80],
[2000,4.10],
[2004,3.80],
[2008,-0.10],
[2012,2.20],
[2016,1.60]]
gdp_change = pd.DataFrame(data=data, columns=columns)
# +
columns = ['Year','US Military Fatalities per Million']
data = [[1952,190],
[1956, 0],
[1960, 0],
[1964, 1],
[1968,146],
[1972, 0],
[1976, 2],
[1980, 0],
[1984, 0],
[1988, 0],
[1992, 0],
[1996, 0],
[2000, 0],
[2004, 4],
[2008, 14],
[2012, 5],
[2016, 5]]
deaths = pd.DataFrame(data=data, columns=columns)
# -
df = votes.merge(growth).merge(unemployment).merge(gdp_change).merge(deaths)
print(df.shape)
df.head()
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error
model = LinearRegression()
features = ['Average Recent Growth in Personal Incomes',
'GDP Change',
'Civilian Unemployment Rate',
'US Military Fatalities per Million']
target = 'Incumbent Party Vote Share'
X = df[features]
y = df[target]
model.fit(X, y)
y_pred = model.predict(X)
# +
# After experimenting, got lowest MAE of 1.34 with using four variables
print('4 Variable Regression MAE: ', mean_absolute_error(y, y_pred))
# -
df['Linear Regression, 4 features'] = y_pred
ax = df.plot(x='Average Recent Growth in Personal Incomes',
y='Incumbent Party Vote Share',
kind='scatter');
df.plot(x='Average Recent Growth in Personal Incomes',
y='Linear Regression, 4 features',
color='green',
kind='scatter',
ax=ax);
# +
# Trying pandas read html webscraping
# Had trouble parsing the specifics
tables = pd.read_html('https://en.wikipedia.org/wiki/United_States_military_casualties_of_war#Wars_ranked_by_total_number_of_U.S._military_deaths')
print(tables[0])
# -
war_df, = pd.read_html('https://en.wikipedia.org/wiki/United_States_military_casualties_of_war#Wars_ranked_by_total_number_of_U.S._military_deaths',
header=0)
# +
# pd.read_html?
# +
# Spurious correlation?
# Rainfall. Source: https://www.statista.com/statistics/500472/annual-average-temperature-in-the-us/
# Didn't work, had to pay for data more specific than per decade
# columns = ['Year','Average Annual US Temp']
# data = [[1952,190],
# [1956, 0],
# [1960, 0],
# [1964, 1],
# [1968,146],
# [1972, 0],
# [1976, 2],
# [1980, 0],
# [1984, 0],
# [1988, 0],
# [1992, 0],
# [1996, 0],
# [2000, 0],
# [2004, 4],
# [2008, 14],
# [2012, 5],
# [2016, 5]]
# deaths = pd.DataFrame(data=data, columns=columns)
| 2.1.3 Assignment Mastin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id="title_ID"></a>
# # JWST Pipeline Validation Notebook:
# # calwebb_detector1, persistence unit tests
#
# <span style="color:red"> **Instruments Affected**</span>: NIRCam, NIRISS, NIRSpec, FGS
#
# ### Table of Contents
#
# <div style="text-align: left">
#
# <br> [Introduction](#intro)
# <br> [JWST Unit Tests](#unit)
# <br> [Defining Terms](#terms)
# <br> [Test Description](#description)
# <br> [Data Description](#data_descr)
# <br> [Imports](#imports)
# <br> [Convenience Functions](#functions)
# <br> [Perform Tests](#testing)
# <br> [About This Notebook](#about)
# <br>
#
# </div>
# <a id="intro"></a>
# # Introduction
#
# This is the validation notebook that displays the unit tests for the Persistence step in calwebb_detector1. This notebook runs and displays the unit tests that are performed as a part of the normal software continuous integration process. For more information on the pipeline visit the links below.
#
# * Pipeline description: https://jwst-pipeline.readthedocs.io/en/latest/jwst/persistence/index.html
#
# * Pipeline code: https://github.com/spacetelescope/jwst/tree/master/jwst/
#
# [Top of Page](#title_ID)
# <a id="unit"></a>
# # JWST Unit Tests
#
# JWST unit tests are located in the "tests" folder for each pipeline step within the [GitHub repository](https://github.com/spacetelescope/jwst/tree/master/jwst/), e.g., ```jwst/persistence/tests```.
#
# * Unit test README: https://github.com/spacetelescope/jwst#unit-tests
#
#
# [Top of Page](#title_ID)
# <a id="terms"></a>
# # Defining Terms
#
# These are terms or acronymns used in this notebook that may not be known a general audience.
#
# * JWST: <NAME> Space Telescope
#
# * NIRCam: Near-Infrared Camera
#
#
# [Top of Page](#title_ID)
# <a id="description"></a>
# # Test Description
#
# Unit testing is a software testing method by which individual units of source code are tested to determine whether they are working sufficiently well. Unit tests do not require a separate data file; the test creates the necessary test data and parameters as a part of the test code.
#
#
# [Top of Page](#title_ID)
# <a id="data_descr"></a>
# # Data Description
#
# Data used for unit tests is created on the fly within the test itself, and is typically an array in the expected format of JWST data with added metadata needed to run through the pipeline.
#
#
# [Top of Page](#title_ID)
# <a id="imports"></a>
# # Imports
#
# * tempfile for creating temporary output products
# * pytest for unit test functions
# * jwst for the JWST Pipeline
# * IPython.display for display pytest reports
#
# [Top of Page](#title_ID)
import tempfile
import pytest
import jwst
from IPython.display import IFrame
# <a id="functions"></a>
# # Convenience Functions
#
# Here we define any convenience functions to help with running the unit tests.
#
# [Top of Page](#title_ID)
def display_report(fname):
'''Convenience function to display pytest report.'''
return IFrame(src=fname, width=700, height=600)
# <a id="testing"></a>
# # Perform Tests
#
# Below we run the unit tests for the Persistence step.
#
# [Top of Page](#title_ID)
with tempfile.TemporaryDirectory() as tmpdir:
# !pytest jwst/persistence -v --ignore=jwst/associations --ignore=jwst/datamodels --ignore=jwst/stpipe --ignore=jwst/regtest --html=tmpdir/unit_report.html --self-contained-html
report = display_report('tmpdir/unit_report.html')
report
# <a id="about"></a>
# ## About This Notebook
# **Author:** <NAME>, Staff Scientist, NIRCam
# <br>**Updated On:** 01/07/2021
# [Top of Page](#title_ID)
# <img style="float: right;" src="./stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="stsci_pri_combo_mark_horizonal_white_bkgd" width="200px"/>
| jwst_validation_notebooks/persistence/jwst_persistence_unit_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
experiment_label = 'LSVC01_abs'
user_label = 'tay_donovan'
# ## **Aim**
# Look for performance improvement in Linear SVC model:
# 1. Absolute values for negative values
# 2. Use GradientSearch
# 3. Scale values
# ## **Findings**
#
# First pass: {'C': 10000, 'degree': 3, 'gamma': 1e-10, 'kernel': 'rbf'}
# Second: {'C': 200000, 'degree': 3, 'gamma': 5e-11, 'kernel': 'rbf'}
# Third: {'C': 20000, 'degree': 3, 'gamma': 6e-06, 'kernel': 'rbf'}
#Initial imports
import pandas as pd
import numpy as np
import seaborn as sb
import matplotlib.pyplot as plt
import os
import sys
sys.path.append(os.path.abspath('..'))
from src.common_lib import DataReader, NBARawData
from sklearn.svm import LinearSVC
from sklearn.svm import SVC
from sklearn.calibration import CalibratedClassifierCV
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import GridSearchCV
# ## **Data input and cleansing**
#Load dataset using common function DataReader.read_data()
data_reader = DataReader()
# Load Raw Train Data
df_train = data_reader.read_data(NBARawData.TRAIN)
# Load Test Raw Data
df_test = data_reader.read_data(NBARawData.TEST)
#For train dataframe, remove redundant column 'Id_old'
cols_drop = ["Id", "Id_old"]
df_train.drop(cols_drop, axis=1, inplace=True)
df_train.columns = df_train.columns.str.strip()
df_train.describe
#For test dataframe, remove redundant column 'Id_old'
df_test.drop(cols_drop, axis=1, inplace=True)
df_test.columns = df_test.columns.str.strip()
df_test.describe
# ## **Negative values in dataset**
print(df_train.where(df_train < 0).count())
# Negative values do not make sense in this context
#Define negative cleaning function
def clean_negatives(strategy, df):
if strategy=='abs':
df = abs(df)
if strategy=='null':
df[df < 0] = 0
if strategy=='mean':
df[df < 0] = None
df.fillna(df.mean(), inplace=True)
return(df)
# +
#Clean negative numbers
negatives_strategy = 'abs'
df_train = clean_negatives(negatives_strategy, df_train)
df_test = clean_negatives(negatives_strategy, df_test)
# -
# ## **Feature Correlation and Selection**
# +
#Use Pearson Correlation to determine feature correlation
pearsoncorr = df_train.corr('pearson')
#Create heatmap of pearson correlation factors
fig, ax = plt.subplots(figsize=(10,10))
sb.heatmap(pearsoncorr,
xticklabels=pearsoncorr.columns,
yticklabels=pearsoncorr.columns,
cmap='RdBu_r',
annot=True,
linewidth=0.2)
# -
# ## **Standard Scaling**
# +
#Standardise scaling of all feature values
scaler = StandardScaler()
df_cleaned = df_train.copy()
df_test_cleaned = df_test.copy()
target = df_cleaned.pop('TARGET_5Yrs')
df_train_cleaned = scaler.fit_transform(df_cleaned)
df_test_cleaned = scaler.fit_transform(df_test_cleaned)
df_train_scaled = pd.DataFrame(df_train_cleaned)
df_train_scaled.columns = df_cleaned.columns
df_train_scaled['TARGET_5Yrs'] = target
# -
# Split the training dataset using common function data_reader.splitdata
X_train, X_val, y_train, y_val = data_reader.split_data(df_train)
#X_train, X_val, y_train, y_val = data_reader.split_data(df_train_scaled)
# ## **Model Selection and Training**
# +
# defining parameter range
param_grid = {'C': [0.08, 0.1, 0.12, 0.14 ],
'tol': [0.001,0.1,1,10]}
grid = GridSearchCV(SVC(max_iter=20000,kernel='linear'), param_grid,scoring="f1", n_jobs=-2)
# fitting the model for grid search
grid.fit(X_train, y_train)
#Print the optimised parameters
print(grid.best_params_)
# -
#Create model with the optimised parameters
model = SVC(C=0.06, class_weight='balanced', kernel='linear', probability=True,
max_iter=-1, random_state=23,tol=0.1, verbose=False)
model.fit(X_train, y_train);
#Store model in /models
from joblib import dump
dump(model, '../models/' + experiment_label + '.joblib')
# ## **Model Evaluation**
#Create predictions for train and validation
y_train_preds = model.predict(X_train)
y_val_preds = model.predict(X_val)
#Evaluate train predictions
#from src.models.aj_metrics import confusion_matrix
from sklearn.metrics import roc_auc_score, accuracy_score
from sklearn.metrics import plot_roc_curve, plot_precision_recall_curve
from sklearn.metrics import classification_report
sys.path.append(os.path.abspath('..'))
from src.models.aj_metrics import confusion_matrix
y_train_preds
#Training performance results
print("ROC AUC Score:")
print(roc_auc_score(y_train,y_train_preds))
print("Accuracy Score:")
print(accuracy_score(y_train, y_train_preds))
print(classification_report(y_train, y_train_preds))
#Confusion matrix
print(confusion_matrix(y_train, y_train_preds))
#ROC Curve
plot_roc_curve(model,X_train, y_train)
#Precision Recall Curve
plot_precision_recall_curve(model,X_train,y_train)
#Validation performance analysis
print("ROC AUC Score:")
print(roc_auc_score(y_val,y_val_preds))
print("Accuracy Score:")
print(accuracy_score(y_val, y_val_preds))
print("Confusion Matrix:")
print(classification_report(y_val, y_val_preds))
#Confusion matrix
print(confusion_matrix(y_train, y_train_preds))
#ROC Curve
plot_roc_curve(model,X_val, y_val)
#Precision Recall Curve
plot_precision_recall_curve(model,X_train,y_train)
# ## **Test output**
#Output predictions
X_test = df_test
y_test_preds = model.predict_proba(X_test)[:,1]
y_test_preds
output = pd.DataFrame({'Id': range(0,3799), 'TARGET_5Yrs': [p for p in y_test_preds]})
output.to_csv("../reports/" + user_label + "_submission_" + experiment_label + ".csv", index=False)
# ## **Outcome**
# After outputting the predictions into kaggle, the final score was 0.70863
| notebooks/tay_donovan_12964300_week3_linearSVC_abs_C0.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''py37tf2cpu'': venv)'
# language: python
# name: python37564bitpy37tf2cpuvenvf1cb5f54778b4d19a30290bfb342f60f
# ---
# + [markdown] id="isXmUBaxJanB"
# Written by: <NAME>
# Last update: Apr 29, 2021
# + [markdown] id="0pXbrXQPKQRF"
# # What is PSID?
#
# PSID stands for preferential subspace identification, a method for dynamic modeling of time-series data, while prioritizing the dynamics shared with another time-series.
#
# For example, given signals $y_k$ (e.g. neural signals) and $z_k$ (e.g behavior), PSID learns a dynamic model for $y_k$ while prioritizing the dynamics that are relevant to $z_k$.
#
# For the derivation and results in real neural data see the paper below.
#
# **Publication:**
#
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. *Modeling behaviorally relevant neural dynamics enabled by preferential subspace identification*. Nature Neuroscience 24, 140–149 (2021). https://doi.org/10.1038/s41593-020-00733-0
#
# View-only full-text link: https://rdcu.be/b993t
#
# Original preprint: https://doi.org/10.1101/808154
#
# You can also find a summary of the paper in the following Twitter thread: https://twitter.com/MaryamShanechi/status/1325835609345122304
# + [markdown] id="BSlPMSM5Ai3P"
# # Installing PSID
# To use PSID, you can either get the source code from [the PSID Github repository](https://github.com/ShanechiLab/PSID), or install it in your Python environment using pip:
#
#
# ```
# pip install PSID
# ```
#
# You can find the usage license in [LICENSE.md](https://github.com/ShanechiLab/PyPSID/blob/main/LICENSE.md). For this notebook, we will also start by installing PSID from pip.
# + colab={"base_uri": "https://localhost:8080/"} id="DZFHrfUEAYmC" outputId="2c807175-d8ba-4c0f-f75e-cc191c653876"
# !pip install PSID --upgrade
# + [markdown] id="KsDpingpAhpK"
# # Using PSID
# ## Modeling data
# To use PSID, you first need to import the library by running:
# ```
# import PSID
# ```
# You can then use its main data modeling function as:
# ```
# idSys = PSID.PSID(y, z, nx, n1, i);
# ```
# With the following arguments:
# - `y` and `z`: Neural (e.g. LFP signal powers or spike counts) and behavioral data (e.g. joint angles, hand position, etc), respectively. Dimensions are: time x data dimension (this can be changed with an optional argument documented in the code).
# - `nx`: the total dimension of the latent state in the model.
# - `n1`: the number of latent state dimensions that are going to be dedicated to behaviorally relevant neural dynamics.
# - `i`: the subspace horizon used for modeling. There is more on the choice of `i` later in this notebook, but numbers such as 5 or 10 are typically suitable values for `i`.
#
# And the following output:
# - `idSys`: an object containing all the learned model parameters ($A$, $C_y$, $C_z$, etc) and some prediction, etc methods. There is more on the model structure later in this notebook.
#
# ## Using the model for dimension reduction, state estimation, and decoding
# For a learned PSID model `idSys`, you can use the `predict` method to extract the latent state and predict behavior and neural activity given any new neural data as:
# ```
# zPred, yPred, xPred = idSys.predict(yTest)
# ```
# With the argument:
# - `yTest`: Neural activity `y` in the test data. Dimensions are: time x data dimension.
#
# And outputs (all dimensions are time x data dimension):
# - `zPred`: Prediction of behavior using past neural activity at each data point.
# - `yPred`: Prediction of neural activity using past neural activity at each data point.
# - `xPred`: The latent state extarcted at each data point.
#
# We will next go through a complete example of using PSID in data.
#
# # A complete example
# In this example, we will use PSID to model some data. First, we import PSID and a few other useful tools from PSID and other libraries.
# + id="bcBhkYe_Bt22"
import argparse, sys, os
sys.path.insert(0, os.path.join('..', '..'))
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import patches
import PSID
from PSID.evaluation import evalPrediction
from PSID.MatHelper import loadmat
# -
# Let's start by loading an example model:
# + colab={"base_uri": "https://localhost:8080/"} id="2DSKNITZB4r-" outputId="a33507ed-9df3-4041-d227-06b0e6317125"
# Load data
sample_model_path = os.path.join(os.path.dirname(PSID.__file__), 'example', 'sample_model.mat')
print('Loading example model from {}'.format(sample_model_path))
data = loadmat(sample_model_path)
# This is an example model (shown in Supplementary Fig. 1) with
# (a) 2 behaviorally relevant latent states,
# (b) 2 behaviorally irrelevant latent states, and
# (c) 2 states that drive behavior but are not represented in neural activity
# + [markdown] id="TKjZYzGV4Oya"
# The PSID model looks like this:
#
# $$ x_{k+1} = A x_k + w_k $$
#
# $$ y_k = C_y x_k + v_k $$
#
# $$ z_k = C_z x_k + \epsilon_k $$
#
# where $y_k \in \!R^{n_y}$ is the neural activity, $z_k \in \!R^{n_z}$ is the behavior, and $x_k \in \!R^{n_x}$ is the latent state the describes the dynamics in both. Note that in general $y_k$ and $z_k$ could also be any other two signals (e.g. brain activity from two regions or even non-neural signals), but here we will refer these signals as neural activity and behavior, respectively. Importantly, PSID learns the model in the following format
#
# $$
# x_k = \begin{bmatrix}
# x_k^{(1)} \\
# x_k^{(2)}
# \end{bmatrix}
# $$
#
# where the behaviorally relevant dimensions of latent state ($x_k^{(1)} \in \!R^{n_1}$), which are those that drive $z_k$, are separated from the other dimensions ($x_k^{(2)} \in \!R^{n_2}$ with $n_2=n_x-n_1$). There are many equivalent ways of writing a latent state model such as this one, but PSID learns the one that uses minimal number of dimensions to explain behavior as parsimoniously as possible (you can find the precise definition in the paper). Critically, PSID can learn this minimal model (with only $x_k^{(1)}$) without having to also learn the rest of the model (the $x_k^{(2)}$ part). This is the concept of prioritization and allows PSID to learn the model more accurately, while requiring fewer training samples.
#
#
# Before going further, let's generate some sample data from this model.
# + id="-9bAIpLC4LaE"
# Generating some sample data from this model
np.random.seed(42) # For exact reproducibility
N = int(2e4)
trueSys = PSID.LSSM(params=data['trueSys'])
y, x = trueSys.generateRealization(N)
z = (trueSys.Cz @ x.T).T
# Add some z dynamics that are not encoded in y (i.e. epsilon)
epsSys = PSID.LSSM(params=data['epsSys'])
eps, _ = epsSys.generateRealization(N)
z += eps
allYData, allZData = y, z
# + [markdown] id="CQWYLQuwGWGY"
# Let's separate the data into training and test segments.
# + id="eiefXAMyExUa"
# Separate data into training and test data:
trainInds = np.arange(np.round(0.5*allYData.shape[0]), dtype=int)
testInds = np.arange(1+trainInds[-1], allYData.shape[0])
yTrain = allYData[trainInds, :]
yTest = allYData[testInds, :]
zTrain = allZData[trainInds, :]
zTest = allZData[testInds, :]
# + [markdown] id="5_HZ9Xd_G7ew"
# We will next use PSID in two ways:
# 1. Learn a model with a low-dimensional latent state that only focuses on learning the behaviorally relevant neural dynamics (i.e. uses stage 1 of PSID only).
# 2. Learn a model that also learns other neural dynamics (i.e. uses both stages of PSID)
#
# We will then plot the learned models' eigenvalues (the eigenvalues of the $A$ matrix) to show that PSID learns the correct dynamics in each case.
#
# First, let's learn a model with a 2 dimensional latent state that only learns the behaviorally relevant neural dyanmics. For this, we pass the arguments nx=2 and n1=2 to the PSID function:
# + colab={"base_uri": "https://localhost:8080/"} id="KeAWDK3sGfmr" outputId="39967d6f-e864-4236-8c87-8af7dd489f1e"
## (Example 1) PSID can be used to dissociate and extract only the
# behaviorally relevant latent states (with nx = n1 = 2)
idSys1 = PSID.PSID(yTrain, zTrain, nx=2, n1=2, i=10)
# You can also use the time_first=False argument if time is the second dimension:
# idSys1 = PSID.PSID(yTrain.T, zTrain.T, nx=2, n1=2, i=10, time_first=False)
# -
# The PSID learning function returns an object (here idSys1) that contains the learned model parameters and can be used to extract the latent states and decode behavior in new data. To do this, we use the 'predict' method in the learned model:
# +
# Predict behavior using the learned model
zTestPred1, yTestPred1, xTestPred1 = idSys1.predict(yTest)
# Compute CC of decoding
CC = evalPrediction(zTest, zTestPred1, 'CC')
# Predict behavior using the true model for comparison
zTestPredIdeal, yTestPredIdeal, xTestPredIdeal = trueSys.predict(yTest)
CCIdeal = evalPrediction(zTest, zTestPredIdeal, 'CC')
print('Behavior decoding CC:\n PSID => {:.3g}, Ideal using true model => {:.3g}'.format(np.mean(CC), np.mean(CCIdeal)) )
# + [markdown] id="LfhHQ3hQKkeq"
# We can see that the PSID model with a 2D latent state is as accurate in explaining behavior as the full model that has a 4D latent state. This is because the other 2 latent state dimensions in the true model explain dynamics that are exclusive to neural activity (i.e. are not behaviorally relevant).
#
# Optionally, PSID can also learn other latent states beyond the behaviorally relevant ones. For this, we pass the arguments nx=4 and n1=2 to the PSID function:
# + colab={"base_uri": "https://localhost:8080/"} id="a2chTfdSKW5d" outputId="2ad2af37-c80b-4a1b-dfba-9039a56ce8e0"
## (Example 2) Optionally, PSID can additionally also learn the
# behaviorally irrelevant latent states (with nx = 4, n1 = 2)
idSys2 = PSID.PSID(yTrain, zTrain, nx=4, n1=2, i=10)
# In addition to ideal behavior decoding, this model will also have ideal neural self-prediction
zTestPred2, yTestPred2, xTestPred2 = idSys2.predict(yTest)
yCC2 = evalPrediction(yTest, yTestPred2, 'CC')
yCCIdeal = evalPrediction(yTest, yTestPredIdeal, 'CC')
print('Neural self-prediction CC:\n PSID => {:.3g}, Ideal using true model => {:.3g}'.format(np.mean(yCC2), np.mean(yCCIdeal)))
# + [markdown] id="LDvqwXjRLKgj"
# We can see that in this case, the model learned by PSID (which now has a 4D latent state) is also as good as the true model in terms of explaining neural activity.
#
#
# Finally, we can plot the eigenvalues of the $A$ matrix in each of the learned models and compare them with the eigenvalues of the $A$ matrix in the true model to see the accurate learning of behaviorally relevant (and optionally the other) dynamics by PSID.
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="rglPnehvLJ-2" outputId="57690508-4203-4361-bd15-25056eff6395"
# #########################################
# Plot the true and identified eigenvalues
# (Example 1) Eigenvalues when only learning behaviorally relevant states
idEigs1 = np.linalg.eig(idSys1.A)[0]
# (Example 2) Additional eigenvalues when also learning behaviorally irrelevant states
# The identified model is already in form of Eq. 4, with behaviorally irrelevant states
# coming as the last 2 dimensions of the states in the identified model
idEigs2 = np.linalg.eig(idSys2.A[2:, 2:])[0]
relevantDims = trueSys.zDims - 1 # Dimensions that drive both behavior and neural activity
irrelevantDims = [x for x in np.arange(trueSys.state_dim, dtype=int) if x not in relevantDims] # Dimensions that only drive the neural activity
trueEigsRelevant = np.linalg.eig(trueSys.A[np.ix_(relevantDims, relevantDims)])[0]
trueEigsIrrelevant = np.linalg.eig(trueSys.A[np.ix_(irrelevantDims, irrelevantDims)])[0]
nonEncodedEigs = np.linalg.eig(data['epsSys']['a'])[0] # Eigenvalues for states that only drive behavior
fig = plt.figure(figsize=(8, 4))
axs = fig.subplots(1, 2)
axs[1].remove()
ax = axs[0]
ax.axis('equal')
ax.add_patch( patches.Circle((0,0), radius=1, fill=False, color='black', alpha=0.2, ls='-') )
ax.plot([-1,1,0,0,0], [0,0,0,-1,1], color='black', alpha=0.2, ls='-')
ax.scatter(np.real(nonEncodedEigs), np.imag(nonEncodedEigs), marker='o', edgecolors='#0000ff', facecolors='none', label='Not encoded in neural signals')
ax.scatter(np.real(trueEigsIrrelevant), np.imag(trueEigsIrrelevant), marker='o', edgecolors='#ff0000', facecolors='none', label='Behaviorally irrelevant')
ax.scatter(np.real(trueEigsRelevant), np.imag(trueEigsRelevant), marker='o', edgecolors='#00ff00', facecolors='none', label='Behaviorally relevant')
ax.scatter(np.real(idEigs1), np.imag(idEigs1), marker='x', facecolors='#00aa00', label='PSID Identified (stage 1)')
ax.scatter(np.real(idEigs2), np.imag(idEigs2), marker='x', facecolors='#aa0000', label='(optional) PSID Identified (stage 2)')
ax.set_title('True and identified eigevalues')
ax.legend(bbox_to_anchor=(1.04,0.5), loc="center left", borderaxespad=0)
plt.show()
# + [markdown] id="OIg0PhNeFcyj"
# # Using PSID with trial based data
# You can also use PSID if the data is available in separate chunks, for example across many trials. To do this, simply pass a python list with the data in each chunk/trial as the argument to PSID. The trials don't need to have the same number of samples either.
#
# Below is an example, where we break the same data as before in small chunks of random length, and then pass it to PSID.
# + colab={"base_uri": "https://localhost:8080/"} id="-McGIeBhE90J" outputId="91fa16cf-aa58-4688-e695-5c675dd12886"
## (Example 3) PSID can be used if data is available in discontinuous segments (e.g. different trials)
# In this case, y and z data segments must be provided as elements of a list
# Trials do not need to have the same number of samples
# Here, for example assume that trials start at every 1000 samples.
# And each each trial has a random length of 500 to 900 samples
trialStartInds = np.arange(0, allYData.shape[0]-1000, 1000)
trialDurRange = np.array([900, 990])
trialDur = np.random.randint(low=trialDurRange[0], high=1+trialDurRange[1], size=trialStartInds.shape)
trialInds = [trialStartInds[ti]+np.arange(trialDur[ti]) for ti in range(trialStartInds.size)]
yTrials = [allYData[trialIndsThis, :] for trialIndsThis in trialInds]
zTrials = [allZData[trialIndsThis, :] for trialIndsThis in trialInds]
# Separate data into training and test data:
trainInds = np.arange(np.round(0.5*len(yTrials)), dtype=int)
testInds = np.arange(1+trainInds[-1], len(yTrials))
yTrain = [yTrials[ti] for ti in trainInds]
yTest = [yTrials[ti] for ti in testInds]
zTrain = [zTrials[ti] for ti in trainInds]
zTest = [zTrials[ti] for ti in testInds]
idSys3 = PSID.PSID(yTrain, zTrain, nx=2, n1=2, i=10)
for ti in range(len(yTest)):
zPredThis, yPredThis, xPredThis = idSys3.predict(yTest[ti])
zPredThisIdeal, yPredThisIdeal, xPredThisIdeal = trueSys.predict(yTest[ti])
if ti == 0:
zTestA = zTest[ti]
zPredA = zPredThis
zPredIdealA = zPredThisIdeal
else:
zTestA = np.concatenate( (zTestA, zTest[ti]), axis=0)
zPredA = np.concatenate( (zPredA, zPredThis), axis=0)
zPredIdealA = np.concatenate( (zPredIdealA, zPredThisIdeal), axis=0)
CCTrialBased = evalPrediction(zTestA, zPredA, 'CC')
CCTrialBasedIdeal = evalPrediction(zTestA, zPredIdealA, 'CC')
print('Behavior decoding CC (trial-based learning/decoding):\n PSID => {:.3g}, Ideal using true model = {:.3g}'.format(np.mean(CCTrialBased), np.mean(CCTrialBasedIdeal)) )
# -
# # How to pick the state dimensions `nx` and `n1`?
# `nx` determines the total dimension of the latent state and `n1` determines how many of those dimensions will be prioritizing the inclusion of behaviorally relevant neural dynamcis (i.e. will be extracted using stage 1 of PSID). So the values that you would select for these hyperparameters depend on the goal of modeling and on the data. Some examples use cases are:
# - If you want to perform dimension reduction, `nx` will be your desired target dimension. For example, to reduce dimension to 2 to plot low-dimensional visualizations of neural activity, you would use `nx=2`. Now if you want to reduce dimension while preservising as much behaviorally relevant neural dynamics as possible, you would use `n1=nx`.
# - If you want to find the best fit to data overall, you can perform a grid search over values of `nx` and `n1` and pick the value that achieves the best performance metric in the training data. For exmaple, you could pick the `nx` and `n1` pair that achieves the best cross-validated behavior decoding in an inner-cross-validation within the training data.
#
# # How to pick the horizon `i`?
# The horizon `i` does not affect the model structure and only affects the intermediate linear algebra operations that PSID performs during the learning of the model. Nevertheless, different values of `i` may have different model learning performance. `i` needs to be at least 2, but also also determines the maximum `n1` and `nx` that can be used per:
# ```
# n1 <= nz * i
# nx <= ny * i
# ```
# So if you have a low dimensional y or z, you typically would choose larger values for `i`, and vice versa. It is also possible to select the best performing `i` via an inner cross-validation approach similar to `nx` and `n1` above.
# # Licence
# Copyright (c) 2020 University of Southern California
# See full notice in [LICENSE.md](https://github.com/ShanechiLab/PyPSID/blob/main/LICENSE.md)
# <NAME> and <NAME>
# Shanechi Lab, University of Southern California
| source/PSID/example/PSID_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"} id="VQwXfpE3JWKo"
# ## Deep Compressive Object Decoder (DCOD)
# Implementation and proof of work.
# + pycharm={"name": "#%%\n"} id="RtVSqOaXJWKw" colab={"base_uri": "https://localhost:8080/"} outputId="e540087c-ac81-4779-9ce9-28ba0c91c2d1"
import os
import random
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
from network import deep_decoder
import tensorflow as tf
from tensorflow.keras import layers as ls, activations as acts
import tensorflow_addons as tfa
from skimage.restoration import unwrap_phase
from fringe.utils.io import import_image, export_image
from fringe.utils.modifiers import ImageToArray, PreprocessHologram, ConvertToTensor
from fringe.process.gpu import AngularSpectrumSolver as AsSolver
device = 'gpu'
if device == "gpu":
if len(tf.config.experimental.list_physical_devices('GPU')) > 0:
print('GPU is up and running')
device = "/gpu:0"
else:
print('No GPUs found. The process will run on CPU.')
device = "/cpu:0"
elif device == "tpu":
if len(tf.config.experimental.list_physical_devices('TPU')) > 0:
print('TPU is up and running')
device = "/tpu:0"
else:
print('No TPUs found. The process will run on CPU.')
device = "/cpu:0"
else:
device = "/cpu:0"
dtype_f = tf.float32
dtype_c = tf.complex64
# + [markdown] id="4tLKVYD_7w4P"
# ### Samples
# + [markdown] pycharm={"name": "#%% md\n"} id="bEJx1WIzJWKx"
# #### Sample 1: Cheek Cells
# + pycharm={"name": "#%%\n"} id="6neltG2aJWKx" colab={"base_uri": "https://localhost:8080/", "height": 802} outputId="efd36bbb-0e0b-4f91-968b-14a94125af29"
hologram_path = 'Dataset/Cheek cells/hologram.tif'
background_path = 'Dataset/Cheek cells/background.tif'
p1 = ImageToArray(bit_depth=16, channel='gray', crop_window=None, dtype='float32')
bg = import_image(background_path, preprocessor=p1)
p2 = PreprocessHologram(background=bg)
p3 = ConvertToTensor(dtype=dtype_c)
hologram = import_image(hologram_path, preprocessor=[p1, p2, p3])
hologram_amp = tf.math.abs(hologram)
solver = AsSolver(shape=hologram_amp.shape, dx=1.12, dy=1.12, wavelength=532e-3)
z = 238
rec = solver.solve(hologram, z)
amp = np.abs(rec)
#phase = unwrap_phase(np.angle(rec))
plt.imshow(hologram_amp.numpy(), cmap='gray')
plt.show()
plt.imshow(amp, cmap='gray')
plt.show()
plt.hist((hologram_amp.numpy()).flatten(), 256)
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"} id="tabI9kllJWKx"
# #### Sample 2: Red Blood Cells
# + pycharm={"name": "#%%\n"} id="a3wfXP1_JWKy" colab={"base_uri": "https://localhost:8080/", "height": 802} outputId="f74db395-9f2e-4d3c-985c-acad8598893f"
hologram_path = 'Dataset/RBCs/hologram.tif'
background_path = 'Dataset/RBCs/background.tif'
p1 = ImageToArray(bit_depth=16, channel='gray', dtype='float32')
bg = import_image(background_path, preprocessor=p1)
p2 = PreprocessHologram(background=bg)
p3 = ConvertToTensor(dtype=dtype_c)
hologram = import_image(hologram_path, preprocessor=[p1, p2, p3])
hologram_amp = tf.math.abs(hologram)
solver = AsSolver(shape=hologram_amp.shape, dx=1.12, dy=1.12, wavelength=532e-3)
z = 315
rec = solver.solve(hologram, z)
amp = np.abs(rec)
#phase = unwrap_phase(np.angle(rec))
plt.imshow(hologram_amp.numpy(), cmap='gray')
plt.show()
plt.imshow(amp, cmap='gray')
plt.show()
plt.hist((hologram_amp.numpy()).flatten(), 256)
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"} id="ZnuwcDkuJWKy"
# #### Sample 3: Diffraction Grating
# + pycharm={"name": "#%%\n"} id="-M-hOVgJJWKy" colab={"base_uri": "https://localhost:8080/", "height": 802} outputId="73eede8b-8c21-4041-9d7a-2eeaf62eeef4"
hologram_path = 'Dataset/Grating/hologram.tif'
background_path = 'Dataset/Grating/background.tif'
p1 = ImageToArray(bit_depth=16, channel='gray', dtype='float32')
bg = import_image(background_path, preprocessor=p1)
p2 = PreprocessHologram(background=bg)
p3 = ConvertToTensor(dtype=dtype_c)
hologram = import_image(hologram_path, preprocessor=[p1, p2, p3])
hologram_amp = tf.math.abs(hologram)
solver = AsSolver(shape=hologram_amp.shape, dx=1.12, dy=1.12, wavelength=532e-3)
z = 956
rec = solver.solve(hologram, z)
amp = np.abs(rec)
#phase = unwrap_phase(np.angle(rec))
plt.imshow(hologram_amp.numpy(), cmap='gray')
plt.show()
plt.imshow(amp, cmap='gray')
plt.show()
plt.hist((hologram_amp.numpy()).flatten(), 256)
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"} id="FbpFl6-xJWKz"
# #### Sample 4: USAF 1951
# + pycharm={"name": "#%%\n"} id="9HXeNYjyJWKz" colab={"base_uri": "https://localhost:8080/", "height": 802} outputId="0b943c64-575c-4f13-9664-49a63b15c46c"
hologram_path = 'Dataset/USAF 1951/hologram.tif'
background_path = 'Dataset/USAF 1951/background.tif'
p1 = ImageToArray(bit_depth=16, channel='gray', dtype='float32')
bg = import_image(background_path, preprocessor=p1)
p2 = PreprocessHologram(background=bg)
p3 = ConvertToTensor(dtype=dtype_c)
hologram = import_image(hologram_path, preprocessor=[p1, p2, p3])
hologram_amp = tf.math.abs(hologram)
solver = AsSolver(shape=hologram_amp.shape, dx=1.12, dy=1.12, wavelength=532e-3)
z = 1065
rec = solver.solve(hologram, z)
amp = np.abs(rec)
#phase = unwrap_phase(np.angle(rec))
plt.imshow(hologram_amp.numpy(), cmap='gray')
plt.show()
plt.imshow(amp, cmap='gray')
plt.show()
plt.hist((hologram_amp.numpy()).flatten(), 256)
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"} id="q5dyRM6qJWKz"
# #### Sample 5: Simulation
# + pycharm={"name": "#%%\n"} id="asF-f9dGJWK0" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="14e33239-5826-4f27-8166-f3f1b4427aa3"
from skimage.filters import gaussian
from misc_functions import Scale
amp_path = 'Dataset/Simulation source images/baboon.png'
ph_path = 'Dataset/Simulation source images/peppers.png'
p1 = ImageToArray(bit_depth=8, channel='gray', crop_window=None, dtype='float32')
amplitude = import_image(amp_path, preprocessor=p1)
phase = import_image(ph_path, preprocessor=p1)
# Adjusting contrast
amplitude = Scale(amplitude, perc=1, max_val=1)
# Blurriness
sigma = 0 #np.exp(3)
amplitude = gaussian(amplitude, sigma, mode='reflect', truncate=np.round(10 * sigma) + 1)
phase /= np.max(phase)
phase *= 2 * np.pi - 0.2 * np.pi
phase -= np.pi
solver = AsSolver(shape=amplitude.shape, dx=1.12, dy=1.12, wavelength=532e-3)
z = 300
obj_func = tf.convert_to_tensor(amplitude * np.exp(1j * phase), dtype_c)
hologram = solver.solve(obj_func, z)
hologram_amp = tf.math.pow(tf.math.abs(hologram), 2)
plt.imshow(amplitude, cmap='gray', vmin=0, vmax=1)
plt.show()
plt.imshow(phase, cmap='viridis', vmin=-np.pi, vmax=np.pi)
plt.show()
plt.imshow(hologram_amp.numpy(), cmap='gray', vmin=0, vmax=1)
plt.show()
plt.hist((hologram_amp.numpy()).flatten(), 256)
plt.show()
# + [markdown] id="PF40m5q58Mag"
# ### Reconstruction
# + [markdown] pycharm={"name": "#%% md\n"} id="POC24wIFJWK0"
# #### Model Initialization
# + pycharm={"name": "#%%\n"} id="lncwv-YzJWK0" colab={"base_uri": "https://localhost:8080/"} outputId="b9fcf948-7c4b-4b9b-d005-70bac569bcfa"
num_epochs = 30000
lr = tf.Variable(0.01, dtype=dtype_f)
weight_decay = tf.Variable(0.002, dtype=dtype_f)
def get_lr():
return lr.numpy()
def get_wd():
return weight_decay.numpy()
random_seed = 999
print("Random Seed: ", random_seed)
random.seed(random_seed)
tf.random.set_seed(random_seed)
input_t_ref = tf.random.normal([1, 16, 16, 256], mean=0, stddev=0.1, dtype=dtype_f)
input_t = tf.Variable(input_t_ref)
net = deep_decoder(input_shape=input_t[0].shape,
layers_channels=[256, 256, 256, 256, 256],
kernel_sizes=[1]*5,
out_channels=2,
upsample_mode='bilinear',
activation_func=ls.ReLU(),
out_activation=acts.sigmoid,
bn_affine=True)
#################################
optimizer = tfa.optimizers.AdamW(learning_rate=get_lr, weight_decay=get_wd)
mse = tf.keras.losses.MeanSquaredError()
# + pycharm={"name": "#%%\n"} id="RBvebPOsJWK1"
net.summary()
# + [markdown] pycharm={"name": "#%% md\n"} id="r9Sef9Y5JWK1"
# #### Log Settings and Recall
# + pycharm={"name": "#%%\n"} id="rjswwwoEJWK1" colab={"base_uri": "https://localhost:8080/"} outputId="386d5b59-06e6-4963-d4e2-14b38c7a9359"
logs_path = 'PATH_TO_LOGS_FOLDER'
log_folder = 'LOG_FOLDER_NAME'
log_root = os.path.join(logs_path, log_folder)
if not os.path.exists(log_root):
os.mkdir(log_root)
if not os.path.exists(os.path.join(log_root, 'exports')):
os.mkdir(os.path.join(log_root, 'exports'))
amp_coefs = [1.3, 1.4]
amp_coef = tf.Variable(amp_coefs[0], dtype=dtype_f)
amp_rand_std = tf.Variable(0.02, dtype=dtype_f)
checkpoint_folder = 'ckpts'
checkpoint = tf.train.Checkpoint(step=tf.Variable(0),
optimizer=optimizer,
model=net,
input_t=input_t,
amp_coef=amp_coef,
amp_rand_std=amp_rand_std,
lr=lr,
wd=weight_decay)
manager = tf.train.CheckpointManager(checkpoint, os.path.join(log_root, checkpoint_folder), max_to_keep=20)
checkpoint.restore(manager.latest_checkpoint)
save_interval = 5000
amp_coefs_interval = 500
last_log = int(checkpoint.step)
start_epoch = int(checkpoint.step)
if checkpoint.step.numpy() != 0:
print('Continuing training from step:', checkpoint.step.numpy())
else:
print('Initializing model checkpoints')
log_array = np.zeros((save_interval, 3), dtype='float32')
log_name = 'log.csv'
if os.path.exists(os.path.join(log_root, log_name)):
log = pd.read_csv(os.path.join(log_root, log_name), index_col=0)
log.drop(log.index[int(checkpoint.step):], inplace=True)
loss_list = log['loss'].tolist()[-100:]
else:
log = pd.DataFrame(columns=['epoch', 'loss', 'loss_avg'], )
loss_list = []
if int(checkpoint.step) >= save_interval-1:
print('\nparameters:')
print('- learning rate:', checkpoint.lr.numpy())
print('- weight decay:', checkpoint.wd.numpy())
print('- amp coefficient:', checkpoint.amp_coef.numpy())
plt.plot(log['loss_avg'])
plt.yscale('log')
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"} id="h_h2R7PeJWK1"
# #### Optimization
# + pycharm={"name": "#%%\n"} id="cw9r-15hJWK2" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="aab00bff-ea00-4922-a840-8c2d02d1289f"
random_init = True
random_amp = True
''''''
lr.assign(0.01)
weight_decay.assign(0.002)
amp_rand_std.assign(0.02)
amp_coefs = [1.3, 1.4]
amp_coefs_interval = 500
num_epochs = 30000
trainable_variables = net.trainable_variables
cmap = matplotlib.cm.get_cmap('viridis')
for epoch in range(int(checkpoint.step), num_epochs):
with tf.device(device):
with tf.GradientTape(persistent=True) as tape:
if (random_init or random_amp) and epoch % amp_coefs_interval == 0 and epoch != start_epoch:
print("Randomizing on step: {}".format(epoch))
if random_init:
input_t.assign(input_t_ref + tf.random.normal(input_t_ref.shape, mean=0, stddev=amp_rand_std.numpy(), dtype=dtype_f))
print("Adding noise to the initiallizer")
if random_amp:
amp_coef_idx = (epoch % (len(amp_coefs) * amp_coefs_interval)) // amp_coefs_interval
amp_coef.assign(amp_coefs[amp_coef_idx])
print("New amp coef: ", amp_coef.numpy())
out = net(input_t, training=True)
out = tf.squeeze(out)
out_ph = out[...,0]
out_ph = tf.scalar_mul(2 * np.pi, out_ph)
out_ph = tf.complex(real=tf.zeros_like(out_ph), imag=out_ph)
out_amp = out[...,1]
out_amp = tf.scalar_mul(amp_coef, out_amp)
out_amp = tf.complex(real=out_amp, imag=tf.zeros_like(out_amp))
out_func = tf.multiply(out_amp, tf.math.exp(out_ph))
out_hol = solver.solve(out_func, z)
out_hol_amp = tf.math.pow(tf.math.abs(out_hol), 2)
loss_value = mse(out_hol_amp, hologram_amp)
loss_list.append(loss_value.numpy())
if len(loss_list) > 100:
loss_list.pop(0)
loss_avg = np.mean(np.array(loss_list))
grads = tape.gradient(loss_value, trainable_variables)
optimizer.apply_gradients(zip(grads, trainable_variables))
if epoch % 20 == 0:
print("Epoch {:03d}: Loss: {:.5f} Loss Avg: {:.5f}".format(epoch, loss_value, loss_avg))
if epoch % 100 == 0:
plt.imshow(out[...,0].numpy())
plt.show()
plt.imshow(out[...,1].numpy(), cmap='gray')
plt.show()
if (epoch + 1) % save_interval == 0 and epoch != start_epoch:
#export_images(net, input_t, directory=os.path.join(log_root, 'exports'), step=int(checkpoint.step))
export_image(out[...,0].numpy(), path=os.path.join(log_root, 'exports', 'out_phase_{:d}.png'.format(int(checkpoint.step))),dtype='uint8')
export_image(cmap(out[...,0].numpy()), path=os.path.join(log_root, 'exports', 'out_phase_c_{:d}.png'.format(int(checkpoint.step))), dtype='uint8')
export_image(out[...,1].numpy(), path=os.path.join(log_root, 'exports', 'out_amp_{:d}.png'.format(int(checkpoint.step))), dtype='uint8')
save_path = manager.save()
print("Saved checkpoint for step: {}".format(epoch))
if (epoch % save_interval == 0 and epoch != start_epoch) or epoch == num_epochs - 1:
save_path = manager.save()
print("Saved checkpoint for step: {}".format(epoch))
log_array = log_array[log_array[:,1] != 0, :]
log = pd.concat([log, pd.DataFrame(log_array, columns=['epoch', 'loss', 'loss_avg'])], sort=False, axis=0, ignore_index=True)
log.to_csv(os.path.join(log_root, log_name), header=True, columns=['epoch', 'loss', 'loss_avg'])
log_array = np.zeros((save_interval, 3), dtype='float32')
last_log = epoch
log_array[epoch - last_log] = [epoch, loss_value.numpy(), loss_avg]
checkpoint.step.assign_add(1)
# + [markdown] pycharm={"name": "#%% md\n"} id="A5dMSzNvJWK2"
# #### Export reconstructed holograms
# + pycharm={"name": "#%%\n"} id="1NCgTG-kJWK2"
export_image(out_hol_amp / np.max(out_hol_amp), path=os.path.join(log_root, 'exports', 'hologram_out.png'), dtype='uint8')
export_image(hologram_amp / np.max(hologram_amp), path=os.path.join(log_root, 'exports', 'hologram_in.png'), dtype='uint8')
plt.imshow(out_hol_amp.numpy(), "gray")
plt.show()
plt.imshow(hologram_amp.numpy(), "gray")
plt.show()
| DCOD_Implementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="T79EZr2jnZVF" colab_type="code" outputId="0f11e0d7-50d0-49f9-c62b-87fa53a61ece" colab={"base_uri": "https://localhost:8080/", "height": 55}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="TB16b_slbBBw" colab_type="code" colab={}
# %matplotlib inline
from matplotlib import pyplot as plt
# + id="x3sJNZk7oGDt" colab_type="code" colab={}
import numpy as np
import os
import glob
import cv2
import skimage.io as io
import skimage.transform as trans
# + id="2VYUSS2Ino71" colab_type="code" colab={}
from tensorflow.keras.models import *
from tensorflow.keras.layers import *
from tensorflow.keras.optimizers import *
from tensorflow.keras.callbacks import ModelCheckpoint, LearningRateScheduler
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# + [markdown] id="-1PjJiTjqAXz" colab_type="text"
# # Model
# + id="_E1Ut9r4nvgx" colab_type="code" colab={}
def unet(pretrained_weights = None,input_size = (256,256,1)):
inputs = Input(input_size)
conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(inputs)
conv1 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv1)
pool1 = MaxPooling2D(pool_size=(2, 2))(conv1)
conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool1)
conv2 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv2)
pool2 = MaxPooling2D(pool_size=(2, 2))(conv2)
conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool2)
conv3 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv3)
pool3 = MaxPooling2D(pool_size=(2, 2))(conv3)
conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool3)
conv4 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv4)
drop4 = Dropout(0.5)(conv4)
pool4 = MaxPooling2D(pool_size=(2, 2))(drop4)
conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool4)
conv5 = Conv2D(1024, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv5)
drop5 = Dropout(0.5)(conv5)
up6 = Conv2D(512, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(drop5))
merge6 = concatenate([drop4,up6], axis = 3)
conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge6)
conv6 = Conv2D(512, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv6)
up7 = Conv2D(256, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv6))
merge7 = concatenate([conv3,up7], axis = 3)
conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge7)
conv7 = Conv2D(256, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv7)
up8 = Conv2D(128, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv7))
merge8 = concatenate([conv2,up8], axis = 3)
conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge8)
conv8 = Conv2D(128, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv8)
up9 = Conv2D(64, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv8))
merge9 = concatenate([conv1,up9], axis = 3)
conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge9)
conv9 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
conv9 = Conv2D(2, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(conv9)
conv10 = Conv2D(1, 1, activation = 'sigmoid')(conv9)
model = Model(inputs, conv10)
model.compile(optimizer = Adam(lr = 1e-4), loss = 'binary_crossentropy', metrics = ['accuracy'])
#model.summary()
if(pretrained_weights):
model.load_weights(pretrained_weights)
return model
# + id="Ra_T-fLn0JUz" colab_type="code" colab={}
model = unet()
# + [markdown] id="sQG9e4LzqLNC" colab_type="text"
# # Data Generators
#
# + id="9Aw3_k2i7AsG" colab_type="code" colab={}
def adjustData(img,mask,flag_multi_class,num_class):
if(flag_multi_class):
img = img / 255
mask = mask[:,:,:,0] if(len(mask.shape) == 4) else mask[:,:,0]
new_mask = np.zeros(mask.shape + (num_class,))
for i in range(num_class):
#for one pixel in the image, find the class in mask and convert it into one-hot vector
#index = np.where(mask == i)
#index_mask = (index[0],index[1],index[2],np.zeros(len(index[0]),dtype = np.int64) + i) if (len(mask.shape) == 4) else (index[0],index[1],np.zeros(len(index[0]),dtype = np.int64) + i)
#new_mask[index_mask] = 1
new_mask[mask == i,i] = 1
new_mask = np.reshape(new_mask,(new_mask.shape[0],new_mask.shape[1]*new_mask.shape[2],new_mask.shape[3])) if flag_multi_class else np.reshape(new_mask,(new_mask.shape[0]*new_mask.shape[1],new_mask.shape[2]))
mask = new_mask
elif(np.max(img) > 1):
img = img / 255
mask = mask /255
mask[mask > 0.5] = 1
mask[mask <= 0.5] = 0
return (img,mask)
# + id="mAIPYCByFDqY" colab_type="code" colab={}
def testGenerator(test_path,num_image = 30,target_size = (256,256),flag_multi_class = False,as_gray = True):
for i in range(num_image):
img = io.imread(os.path.join(test_path,"%d.png"%i),as_gray = as_gray)
img = img / 255
img = trans.resize(img,target_size)
img = np.reshape(img,img.shape+(1,)) if (not flag_multi_class) else img
img = np.reshape(img,(1,)+img.shape)
yield img
# + id="f_LlciYsoan_" colab_type="code" colab={}
def trainGenerator(batch_size,train_path,image_folder,mask_folder,aug_dict,image_color_mode = "grayscale",
mask_color_mode = "grayscale",image_save_prefix = "image",mask_save_prefix = "mask",
flag_multi_class = False,num_class = 2,save_to_dir = None,target_size = (256,256),seed = 1):
'''
can generate image and mask at the same time
use the same seed for image_datagen and mask_datagen to ensure the transformation for image and mask is the same
if you want to visualize the results of generator, set save_to_dir = "your path"
'''
image_datagen = ImageDataGenerator(**aug_dict)
mask_datagen = ImageDataGenerator(**aug_dict)
image_generator = image_datagen.flow_from_directory(
train_path,
classes = [image_folder],
class_mode = None,
color_mode = image_color_mode,
target_size = target_size,
batch_size = batch_size,
save_to_dir = save_to_dir,
save_prefix = image_save_prefix,
seed = seed)
mask_generator = mask_datagen.flow_from_directory(
train_path,
classes = [mask_folder],
class_mode = None,
color_mode = mask_color_mode,
target_size = target_size,
batch_size = batch_size,
save_to_dir = save_to_dir,
save_prefix = mask_save_prefix,
seed = seed)
train_generator = zip(image_generator, mask_generator)
for (img,mask) in train_generator:
img,mask = adjustData(img,mask,flag_multi_class,num_class)
yield (img,mask)
# + [markdown] id="fjchqwQKqPWw" colab_type="text"
# # Training
# + id="STcxBo3qDnMs" colab_type="code" colab={}
data_gen_args = dict(rotation_range=0.2,
width_shift_range=0.05,
height_shift_range=0.05,
shear_range=0.05,
zoom_range=0.05,
horizontal_flip=True,
fill_mode='nearest')
myGene = trainGenerator(10,'/content/gdrive/My Drive/Colab Notebooks/background_removal_dataset/dataset','image','label',data_gen_args,save_to_dir = None)
model = unet()
# + id="Srz8kl3_qRmy" colab_type="code" outputId="cee6f7d3-9633-47d3-ddc4-c2a322683230" colab={"base_uri": "https://localhost:8080/", "height": 1290}
model_checkpoint = ModelCheckpoint('/content/gdrive/My Drive/Colab Notebooks/background_removal_dataset/models/unet5.hdf5', monitor='loss',verbose=1, save_best_only=True)
model.fit_generator(myGene,steps_per_epoch=100,epochs=1,callbacks=[model_checkpoint])
# + id="xqsfrk3pNPdD" colab_type="code" colab={}
model = unet()
model.load_weights("/content/gdrive/My Drive/Colab Notebooks/background_removal_dataset/models/unet3.hdf5")
# + id="-hFx3TeVqbXG" colab_type="code" outputId="a00ecba7-88ca-4d6a-ffc9-f36b2be7d1b7" colab={"base_uri": "https://localhost:8080/", "height": 35}
img = cv2.imread("/content/gdrive/My Drive/Colab Notebooks/background_removal_dataset/dataset/image/axis_001.jpg")
img = cv2.cvtColor(img,cv2.COLOR_BGR2BGRA)
#img = cv2.resize(img,(256,256))
img.shape
# + id="EEZAc7sUA5qZ" colab_type="code" outputId="0f3a4086-b6a0-4065-cb06-ddc98c9606b6" colab={"base_uri": "https://localhost:8080/", "height": 91}
img = io.imread("/content/gdrive/My Drive/Colab Notebooks/background_removal_dataset/dataset/image/bearing_011.jpg",as_gray = True)
img = img / 255.
img = trans.resize(img,(256,256))
img = img[:,:,:1]
#img = np.reshape(img,img.shape+(1,)) if (not False) else img
img = np.reshape(img,(1,)+img.shape)
img.shape
# + id="5yINxYhWFYql" colab_type="code" outputId="aa22de51-9836-46de-ec66-fd44c39db870" colab={"base_uri": "https://localhost:8080/", "height": 35}
results = model.predict(img,steps=1)
results.shape
# + id="lW5jsr6KH7WV" colab_type="code" outputId="6ceb9e9e-5fd5-4b1d-dc2f-749e4e77c4a1" colab={"base_uri": "https://localhost:8080/", "height": 269}
plt.imshow(results[0,:,:,0])
plt.show()
# + id="5JkLPz2gb6UW" colab_type="code" outputId="64ecd3b8-de59-4e38-c8a5-2faa6027acfa" colab={"base_uri": "https://localhost:8080/", "height": 53}
np.unique(results)
# + id="xIt93L_UcDHW" colab_type="code" colab={}
| UNet/Unet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import matplotlib.pyplot as plt
import argparse
import numpy as np
from keras.models import load_model
import keras
import tqdm
from data import load_from_H5
from viz import plot_predictions, plot_predictions_no_legend
# %matplotlib inline
# +
test_hdf5_filepath = 'data/Mauna Loa/test.h5'
train_hdf5_filepath = 'data/Mauna Loa/train.h5'
testset = load_from_H5(test_hdf5_filepath)
trainset = load_from_H5(train_hdf5_filepath)
X_test, y_test = testset
X_train, y_train = trainset
# -
num_hidden_layers = 5
n_hidden = 1024 # num hidden units
epochs = 30
batch_size = 200
epochs_multiplier = 1
tau = 0.1
dropout = 0.1
normalize = False
activation='relu'
net = bnn(
X_train,
y_train,
([int(n_hidden)] * num_hidden_layers),
normalize=normalize,
tau=tau,
dropout=dropout,
)
# +
import numpy as np
import matplotlib.pyplot as plt
x2_train = np.arange(-2, 2, 0.1)
x2_test = np.arange(-8, 8, 0.1)
y_train2 = 1-np.cos(x2_train)
x2_train = x2_train[:,np.newaxis]
y_test2 = 1-np.cos(x2_test)
x2_test = x2_test[:,np.newaxis]
plt.plot(x2_train, y_train2, color="r")
plt.plot(x2_test, y_test2)
plt.show()
# -
def plot_predictions_no_legend(net, X_train, y_train, X_test,y_test,
iters=200, n_std=4, ax=None,
zoomed=False):
if ax is None:
plt.close("all")
plt.clf()
fig, ax = plt.subplots(1, 1)
if zoomed:
plt.axis([-1.75, 3.75, -20, 20])
y_means, y_stds = net.predict(X_test, T=iters)
rmse_standard_pred, rmse, test_ll = net.test(X_test, y_test, T=iters)
ax.plot(X_test, y_means,
label="predict",
color="b",
alpha=.8)
ax.plot(X_test, y_test,
label="1-cos(x)",
color="g",
linewidth=1
)
ax.axvline(x=-2, color="gray",
linestyle=":")
ax.axvline(x=2, color="gray",
linestyle=":")
for i in range(n_std):
ax.fill_between(
X_test.squeeze(),
(y_means - y_stds * ((i+1)/2)).squeeze(),
(y_means + y_stds * ((i+1)/2)).squeeze(),
color="b",
alpha=0.25**(i+1)
)
ax.plot(X_train, y_train, color="r", label="train")
#plt.text(-1, 1.5, r'rms:' + str(rmse), {'color': 'r', 'fontsize': 12})
ax.legend()
plt.savefig("/u1/mmanko/columbia/BNN-Uncertainty/images/cos_test.png")
return ax
from bnn import bnn
net = bnn(x2_train, y_train2, n_hidden=([int(1024)] * 4))
net.train(x2_train, y_train2, batch_size=128)
net.tau = 10
plot_predictions_no_legend(net,x2_train, y_train2, x2_test, y_test2)
| cos_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AloysiusButacAdu/Linear-Algebra-58020/blob/main/Python_Exercise_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="U3H0tD1x5UlA"
# # Vector Spaces and its Operation
#
# + id="tKWga_JS4yVK"
import numpy as np
# + id="NTyFY9dO42me"
A = np.array([4,3])
B = np.array([2,-5])
# + colab={"base_uri": "https://localhost:8080/"} id="hntkZZkw47QS" outputId="37b2c262-91ac-4654-eb3a-8cdd0d851e03"
print('Vector A is ', A)
print('Vector b is ', B)
# + [markdown] id="XVwdHltG6KC9"
# ## Vector Size, Dimension, and shape
# + colab={"base_uri": "https://localhost:8080/"} id="y7u_yrOE6KYI" outputId="4695a887-321a-48a4-cc01-b1601798c453"
ball1 = np.array([1,2,3])
ball2 = np.array([0,1,-1])
pool = np.array([ball1, ball2])
print("Shape: ",pool.shape)
print("Size: ",pool.size)
print("Dimension: ",pool.ndim)
print("Matrix: \n",pool)
# + [markdown] id="D-lzgnbvAq_n"
# ## Vector Operations
#
# + colab={"base_uri": "https://localhost:8080/"} id="iHtka4UR-t7W" outputId="f210527e-a44e-4138-adbe-7ed380f46772"
R = A + B
R
# + colab={"base_uri": "https://localhost:8080/"} id="WYoe-twQ_DSB" outputId="95b48f04-7934-427a-c70d-0502d8d688c3"
R = np.add(A,B)
R
# + colab={"base_uri": "https://localhost:8080/"} id="QGibzyZc_aAN" outputId="160e5ec3-20f5-46c0-90cb-97b9eeae21e1"
R = np.subtract(A,B)
R
# + [markdown] id="0lDSZQsjA-4s"
# ## Vector Scaling
# + colab={"base_uri": "https://localhost:8080/"} id="qqYx3hqgBBMB" outputId="4e0bdd23-b9e4-4a42-8800-d54dce1f2e6a"
A = np.array([1,5,8,9])
S = 5 * A
print("5 *",A)
print(S)
# + colab={"base_uri": "https://localhost:8080/"} id="nU9qHCrqBELO" outputId="85c30bae-f297-46aa-e959-212d901751b2"
S = np.multiply(5,A)
S
# + [markdown] id="gZo-TawLCkA2"
# ## Vector Cross Product
# + colab={"base_uri": "https://localhost:8080/"} id="PQCpxVINCHSO" outputId="30c81188-bc74-4c39-ce4d-2e87c5ce54bb"
#initialize arrays
A = np.array([2,7])
B = np.array([1,7])
#computefor the cross product
output = np.cross(A,B)
print("A:",A,"\tB:",B)
print(output)
# + [markdown] id="5fYzY2fqEDRh"
# ## Vector Dot Product
# + colab={"base_uri": "https://localhost:8080/"} id="ucqanco3DTNF" outputId="6e61595d-b89b-442c-d1c3-43346904966a"
output = np.dot(A,B)
output
| Python_Exercise_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
#
# This notebook is a scratch space in support of
#
# [A Cookbook: Using Distance To Measure Similarity](https://towardsdatascience.com/a-cookbook-using-distance-to-measure-similarity-8de97a1404dd)
# +
import pandas as pd
import numpy as np
from math import sqrt
from scipy.spatial import distance
from scipy.stats import zscore
# -
data = {'Inst':['Institution A','Institution B',
'Institution C','Institution D'],
'Size':[19000,11500,7750,23000],
'Cost':[22000,19000,12000,10500],
'Accept Rt':[.25,.45,.76,.99],
'isBig':[1,0,0,1],
'isExpensive':[1,1,0,0,],
'isSelect':[1,0,0,1]}
df = pd.DataFrame(data)
df
# ## Hamming Distance
# Goal: Count the number of matching dimensions among two observations
def hamming(data, ref, cols):
'''Calculates Hamming distance for all observations
relative to the referenced observation. Returns a list
of hamming distances.
data = A dataframe.
ref = A reference observation. Specify by the axis 0 index.
cols = A set of comparison columns.'''
if type(ref) == int:
ref = ref
if type(ref) == str:
ref = df.index.get_loc(ref)
distances = []
ref_observation = data[cols].iloc[ref]
for row in range(len(data)):
comp_observation = data[cols].iloc[row]
matches = sum([1 if x == y else 0 for x, y in zip(comp_observation,
ref_observation)])
distances.append(matches)
return(distances)
# ref_institution = df.iloc[3]
df['Hamming'] = hamming(data=df, ref=3,
cols=['isBig', 'isExpensive', 'isSelect'])
df
# ## Jaccard Index
# Goal: Count the number of matching dimensions among two observations
# Caluclate Jaccard Index distance
a = df[['isBig','isExpensive','isSelect']].iloc[3]
b = df[['isBig','isExpensive','isSelect']].iloc[0]
matches = sum([1 if x == y else 0 for x, y in zip(a,b)])
dist = 1 - (matches / len(b))
print('Number of matching observations : {}'.format(matches))
print('Jacard Index distance : {}'.format(dist))
def jaccard(data, ref, cols):
'''Calculates Jarcard index for all observations
relative to the referenced observation. Returns a list
of Jaccardian distances.
data = A dataframe.
ref = A reference observation. Specify by the axis 0 index.
cols = A set of comparison columns.'''
distances = hamming(data, ref, cols)
length = len(cols)
distances = [1 - (x/length) for x in distances]
return(distances)
# ref_institution = df.iloc[3]
df['Jaccard'] = jaccard(data=df, ref=3,
cols=['isBig', 'isExpensive', 'isSelect'])
df
# ## Euclidean Distance
# +
# Goal: Count the number of matching dimensions among two observations
# Sample data for test one
a = [2,10]
b = [10,10]
# Calculate from scratch
rise = a[0] - b[0]
run1 = a[1] - b[1]
scatch_dist = sqrt(rise**2 + run1**2)
print('Distance from scratch : {}'.format(scatch_dist))
# Calculate with scipy assist
assist_dist = distance.euclidean(a, b)
print('Distance with assist : {}'.format(scatch_dist))
# +
# Sample data for test one
a = df[['Size','Cost','Accept Rt']].apply(zscore).iloc[0]
b = df[['Size','Cost','Accept Rt']].apply(zscore).iloc[3]
# Calculate from scratch
rise = a[0] - b[0]
run1 = a[1] - b[1]
run2 = a[2] - b[2]
scatch_dist = sqrt(rise**2 + run1**2 + run2**2)
print('Distance from scratch : {}'.format(scatch_dist))
# Calculate with scipy assist
assist_dist = distance.euclidean(a, b)
print('Distance with assist : {}'.format(scatch_dist))
# -
def euclidian(data, ref, cols):
'''Calculates Euclidian distance for all observations
relative to the referenced observation. Returns a list
of euclidian distances.
data = A dataframe.
ref = A reference observation. Specify by the axis 0 index.
cols = A set of comparison columns.'''
if type(ref) == int:
ref = ref
if type(ref) == str:
ref = df.index.get_loc(ref)
distances = []
ref_observation = data[cols].apply(zscore).iloc[ref]
for row in range(len(data)):
comp_observation = data[cols].apply(zscore).iloc[row]
dist = distance.euclidean(ref_observation, comp_observation)
distances.append(dist)
return(distances)
df['Euclidians'] = euclidian(data=df, ref=3, cols=['Size','Cost','Accept Rt'])
df
# # More Categorical Examples
#
# Demonstrate that this implementation does not require one hot encoding. Can operate with text categoricals.
df = pd.DataFrame(data)
df
df['Urban'] = ['Urban','Urban','Rural','Rural']
df['Research'] = ['Low','Low','High','High']
df
df['Jaccard2'] = jaccard(data=df, ref=3, cols=['isBig','isExpensive',
'isSelect','Urban','Research'])
df
df['hamming2'] = hamming(data=df, ref=3, cols=['isBig','isExpensive',
'isSelect','Urban','Research'])
df
# # Update Index to tinker with more readable references
df = pd.DataFrame(data)
df.set_index('Inst', inplace=True)
df
df['Euclidians2'] = euclidian(data=df, ref='Institution D',
cols=['Size','Cost','Accept Rt'])
df
| ComparisonGroupScratchSpace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# metadata:
# interpreter:
# hash: 407adad19d3514da6de0b3767d3a3b1f989b90f5fa3bd5e6e8c15ee36f34523b
# name: python3
# ---
# A tuple can be used to store more than one items (separated by comma)
# A tuple can be created with the use of parenthesis or without
#multiple items
print('Case multiple items:')
print('\r')
tup1=('car','phone',1)
print('Tuple created with parenthesis:\n',tup1)
tup2='car','phone',1
print('Tuple created without parenthesis:\n',tup2)
print('\r')
print('---------------------------------------')
print('\r')
# To create a tuple with only one item, then a comma should be placed after the item
print('Case one item:')
print('\r')
tup3=('car')
print('Data type without use of comma:\n',type(tup3))
tup4=('car',)
print('Data type with use of comma:\n',type(tup4))
# Accesing tuple elements and len(tuple)
tup1=('car','phone',1,'dog','radio','summer',5,'laptop')
print('Tuple:',tup1)
print('\r')
print('First tuple element:',tup1[0])
print('Last tuple element:',tup1[-1])
print('In between tuple elements:',tup1[1:-1])
print('First three tuple elements:',tup1[:3])
print('Last three tuple elements:',tup1[-3:])
print('Tuple elements starting at index 0 with step 2:',tup1[::2])
print('Tuple elements starting at index -1 with step -2:',tup1[::-2])
print('Length of tuple is:',len(tup1))
# Count of tuple elements
tup=(1,2,3,4,2,2,1,4,4,2)
for x in set(tup):
print('Count of tuple element {}: {}\n'.format(x,tup.count(x)))
# Difference between a list and a tuple is that a tuple is immutable (it does not upport item assignment)
# Returns a TypeError -->'tuple' object does not support item assignment
tup1=('car','phone',1)
tup1[-1]='dog'
print('New tuple:\n',tup1)
# Unpacking a tuple
tup1=('car','BMW',2020)
vehicle,make,year_model=tup1
print('Type of vehicle:\n',vehicle)
print('Car Make:\n',make)
print('Car Year Model:\n',year_model)
# +
# A tuple can be created from a list and vice versa
lst1=['car','phone','apple']
tup1=tuple(lst1)
print('Tuple created from list:\n',tup1)
lst=list(tup1)
print('\r')
print('List created from tuple:\n',lst)
# -
# A dict can be created from tuples
tup1=(1,2,3)
tup2=('one','two','three')
d=dict(zip(tup1,tup2))
print('Dict created from tuples:\n',d)
# +
# Tuple 'for' loop and 'if' statement example
tup=(1,2,3,4,5,6)
# for loop
print("Tuple--> 'for' loop:\n")
for item in tup:
print(item,end=' ')
# if statement
print('\r')
print("\nTuple--> 'if' statement:\n")
val=[4,8]
for x in val:
if x in tup:
print('Item {} is in tup and its index is {}:\n'.format(x,tup.index(x),end=' '))
else:
print('Item {} is not in tup'.format(x))
| Tuples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] raw_mimetype="text/restructuredtext"
# .. _nb_subset_selection:
# -
# ## Subset Selection Problem
#
# A genetic algorithm can be used to approach subset selection problems by defining custom operators. In general, a metaheuristic algorithm might not be the ultimate goal to implement in a real-world scenario; however, it might be useful to investigate patterns or characteristics of possible well-performing subsets.
# Let us consider a simple toy problem where we have to select numbers from a list. For every solution, exactly ten numbers have to be selected that their sum is minimized.
# For the subset selection problem, a binary encoding can be used where **one** indicates a number is picked. In our problem formulation, the list of numbers is represented by $L$ and the binary encoded variable by $x$.
#
# \begin{align}
# \begin{split}
# \min f(x) & = & \sum_{k=1}^{n} L_k \cdot x_k\\[2mm]
# \text{s.t.} \quad g(x) & = & (\sum_{k=1}^{n} x_k - 10)^2\\[2mm]
# \end{split}
# \end{align}
#
# As shown above, the equality constraint is handled by ensuring $g(x)$ can only be zero if exactly ten numbers are chosen.
# The problem can be implemented as follows:
# +
import numpy as np
from pymoo.model.problem import Problem
class SubsetProblem(Problem):
def __init__(self,
L,
n_max
):
super().__init__(n_var=len(L), n_obj=1, n_constr=1, elementwise_evaluation=True)
self.L = L
self.n_max = n_max
def _evaluate(self, x, out, *args, **kwargs):
out["F"] = np.sum(self.L[x])
out["G"] = (self.n_max - np.sum(x)) ** 2
# create the actual problem to be solved
np.random.seed(1)
L = np.array([np.random.randint(100) for _ in range(100)])
n_max = 10
problem = SubsetProblem(L, n_max)
# -
# The customization requires writing custom operators in order to solve this problem efficiently. We recommend considering the feasibility directly in the evolutionary operators because otherwise, most of the time, infeasible solutions will be processed.
# The sampling creates a random solution where the subset constraint will always be satisfied.
# The mutation randomly removes a number and chooses another one. The crossover takes the values of both parents and then randomly picks either the one from the first or from the second parent until enough numbers are picked.
# +
from pymoo.model.crossover import Crossover
from pymoo.model.mutation import Mutation
from pymoo.model.sampling import Sampling
class MySampling(Sampling):
def _do(self, problem, n_samples, **kwargs):
X = np.full((n_samples, problem.n_var), False, dtype=np.bool)
for k in range(n_samples):
I = np.random.permutation(problem.n_var)[:problem.n_max]
X[k, I] = True
return X
class BinaryCrossover(Crossover):
def __init__(self):
super().__init__(2, 1)
def _do(self, problem, X, **kwargs):
n_parents, n_matings, n_var = X.shape
_X = np.full((self.n_offsprings, n_matings, problem.n_var), False)
for k in range(n_matings):
p1, p2 = X[0, k], X[1, k]
both_are_true = np.logical_and(p1, p2)
_X[0, k, both_are_true] = True
n_remaining = problem.n_max - np.sum(both_are_true)
I = np.where(np.logical_xor(p1, p2))[0]
S = I[np.random.permutation(len(I))][:n_remaining]
_X[0, k, S] = True
return _X
class MyMutation(Mutation):
def _do(self, problem, X, **kwargs):
for i in range(X.shape[0]):
X[i, :] = X[i, :]
is_false = np.where(np.logical_not(X[i, :]))[0]
is_true = np.where(X[i, :])[0]
X[i, np.random.choice(is_false)] = True
X[i, np.random.choice(is_true)] = False
return X
# -
# After having defined the operators a genetic algorithm can be initialized.
# +
from pymoo.algorithms.so_genetic_algorithm import GA
from pymoo.optimize import minimize
algorithm = GA(
pop_size=100,
sampling=MySampling(),
crossover=BinaryCrossover(),
mutation=MyMutation(),
eliminate_duplicates=True)
res = minimize(problem,
algorithm,
('n_gen', 60),
seed=1,
verbose=True)
print("Function value: %s" % res.F[0])
print("Subset:", np.where(res.X)[0])
# -
# Finally, we can compare the found subset with the optimum known simply through sorting:
opt = np.sort(np.argsort(L)[:n_max])
print("Optimal Subset:", opt)
print("Optimal Function Value: %s" % L[opt].sum())
| doc/source/customization/subset_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Please create a statement that adds the second element and third element
x = [9, 8, 4, 6, 8, 2]
x[1]+x[2]
# Complete the script below to make the Output 1
q = [2]
#"""What goes here"""(len(q))
len(q)
# Complete the script below to make the Output 6
p = 6.47602
# print("""what goes here"""(p))
print(int(p))
# Complete the script below to make the Output blank
p = 0
q = "Hello! "
#print("""what goes here""" p)
print(q[-1])
#Data
revenue = [14574.49, 7606.46, 8611.41, 9175.41, 8058.65, 8105.44, 11496.28, 9766.09, 10305.32, 14379.96, 10713.97, 15433.50]
expenses = [12051.82, 5695.07, 12319.20, 12089.72, 8658.57, 840.20, 3285.73, 5821.12, 6976.93, 16618.61, 10054.37, 3803.96]
# +
#Solution
#Calculate Profit As The Differences Between Revenue And Expenses
profit = []
for i in range (0, len(revenue)):
profit.append(revenue[i] - expenses[i])
print(profit)
print("This is overall profit without tax:",sum(profit))
# +
#Calculate Tax As 30% Of Profit And Round To 2 Decimal Points
tax = [round(i*0.3,2) for i in profit]
tax
print(tax)
print("This is the overall tax: ",sum(tax))
# -
#Calculate Profit Remaining After Tax Is Deducted
profit_after_tax = []
for i in range (0, len(profit)):
profit_after_tax.append(profit[i] - tax[i])
profit_after_tax
print("The profit after deducting the tax is :",sum(profit_after_tax))
| Shah_Rohan_Lab1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
import pandas as pd
import os, sys, gc, warnings, psutil, random
warnings.filterwarnings('ignore')
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
grid_df = pd.concat([pd.read_pickle('./output/grid_part_1.pkl'),
pd.read_pickle('./output/grid_part_2.pkl').iloc[:,2:],
pd.read_pickle('./output/grid_part_3.pkl').iloc[:,2:]],
axis=1)
#
grid_df = grid_df.iloc[:1000, :]
keep_id = np.array_split(list(grid_df['id'].unique()), 20)[0]
grid_df = grid_df[grid_df['id'].isin(keep_id)].reset_index(drop=True)
# Let's "inspect" our grid DataFrame
grid_df.info()
# -
grid_df.drop(['day'], axis=1, inplace=True)
# +
########################### Baseline model
#################################################################################
# We will need some global VARS for future
SEED = 42 # Our random seed for everything
random.seed(SEED) # to make all tests "deterministic"
np.random.seed(SEED)
N_CORES = psutil.cpu_count() # Available CPU cores
TARGET = 'sales' # Our Target
END_TRAIN = 1913 # And we will use last 28 days as validation
# Drop some items from "TEST" set part (1914...)
grid_df = grid_df[grid_df['d'] <= END_TRAIN].reset_index(drop=True)
# Features that we want to exclude from training
remove_features = ['id', 'd', TARGET]
# Our baseline model serves
# to do fast checks of
# new features performance
# We will use LightGBM for our tests
import lightgbm as lgb
lgb_params = {
'boosting_type': 'gbdt', # Standart boosting type
'objective': 'regression', # Standart loss for RMSE
'metric': ['rmse'], # as we will use rmse as metric "proxy"
'subsample': 0.8,
'subsample_freq': 1,
'learning_rate': 0.05, # 0.5 is "fast enough" for us
'num_leaves': 2**7 - 1, # We will need model only for fast check
'min_data_in_leaf': 2**8 -
1, # So we want it to train faster even with drop in generalization
'feature_fraction': 0.8,
'n_estimators':
5000, # We don't want to limit training (you can change 5000 to any big enough number)
'early_stopping_rounds':
30, # We will stop training almost immediately (if it stops improving)
'seed': SEED,
'verbose': -1,
'device': 'gpu',
'gpu_platform_id': -1,
'gpu_device_id': -1
}
## RMSE
def rmse(y, y_pred):
return np.sqrt(np.mean(np.square(y - y_pred)))
# Small function to make fast features tests
# estimator = make_fast_test(grid_df)
# it will return lgb booster for future analisys
def make_fast_test(df):
features_columns = [col for col in list(df) if col not in remove_features]
tr_x, tr_y = df[df['d'] <= (END_TRAIN - 28)][features_columns], df[
df['d'] <= (END_TRAIN - 28)][TARGET]
vl_x, v_y = df[df['d'] > (END_TRAIN - 28)][features_columns], df[
df['d'] > (END_TRAIN - 28)][TARGET]
train_data = lgb.Dataset(tr_x, label=tr_y)
valid_data = lgb.Dataset(vl_x, label=v_y)
estimator = lgb.train(
lgb_params,
train_data,
valid_sets=[train_data, valid_data],
verbose_eval=500,
)
return estimator
# Make baseline model
baseline_model = make_fast_test(grid_df)
# +
########################### Some more info about lags here:
# Small helper to make lags creation faster
from multiprocessing import Pool # Multiprocess Runs
## Multiprocessing Run.
# :t_split - int of lags days # type: int
# :func - Function to apply on each split # type: python function
# This function is NOT 'bulletproof', be carefull and pass only correct types of variables.
## Multiprocess Runs
def df_parallelize_run(func, t_split):
num_cores = np.min([N_CORES,len(t_split)])
pool = Pool(num_cores)
df = pd.concat(pool.map(func, t_split), axis=1)
pool.close()
pool.join()
return df
def make_normal_lag(lag_day):
lag_df = grid_df[['id','d',TARGET]] # not good to use df from "global space"
col_name = 'sales_lag_'+str(lag_day)
lag_df[col_name] = lag_df.groupby(['id'])[TARGET].transform(lambda x: x.shift(lag_day)).astype(np.float16)
return lag_df[[col_name]]
# Launch parallel lag creation
# and "append" to our grid
LAGS_SPLIT = [col for col in range(1,1+7)]
grid_df = pd.concat([grid_df, df_parallelize_run(make_normal_lag,LAGS_SPLIT)], axis=1)
# Make features test
test_model = make_fast_test(grid_df)
# +
########################### Permutation importance Test
# Let's creat validation dataset and features
features_columns = [col for col in list(grid_df) if col not in remove_features]
validation_df = grid_df[grid_df['d']>(END_TRAIN-28)].reset_index(drop=True)
# Make normal prediction with our model and save score
validation_df['preds'] = test_model.predict(validation_df[features_columns])
base_score = rmse(validation_df[TARGET], validation_df['preds'])
print('Standart RMSE', base_score)
# Now we are looping over all our numerical features
for col in features_columns:
# We will make validation set copy to restore
# features states on each run
temp_df = validation_df.copy()
# Error here appears if we have "categorical" features and can't
# do np.random.permutation without disrupt categories
# so we need to check if feature is numerical
if temp_df[col].dtypes.name != 'category':
temp_df[col] = np.random.permutation(temp_df[col].values)
temp_df['preds'] = test_model.predict(temp_df[features_columns])
cur_score = rmse(temp_df[TARGET], temp_df['preds'])
# If our current rmse score is less than base score
# it means that feature most probably is a bad one
# and our model is learning on noise
print(col, np.round(cur_score - base_score, 4))
# Remove Temp data
del temp_df, validation_df
# Remove test features
# As we will compare performance with baseline model for now
keep_cols = [col for col in list(grid_df) if 'sales_lag_' not in col]
grid_df = grid_df[keep_cols]
# -
# from eli5 documentation (seems it's perfect explanation)
#
# The idea is the following: feature importance can be measured by looking at how much the score (accuracy, mse, rmse, mae, etc. - any score we’re interested in) decreases when a feature is not available.
#
# To do that one can remove feature from the dataset, re-train the estimator and check the score. But it requires re-training an estimator for each feature, which can be computationally intensive. Also, it shows what may be important within a dataset, not what is important within a concrete trained model.
#
# To avoid re-training the estimator we can remove a feature only from the test part of the dataset, and compute score without using this feature. It doesn’t work as-is, because estimators expect feature to be present. So instead of removing a feature we can **replace it with random noise** - feature column is still there, but it no longer contains useful information. This method works if noise is drawn from the **same distribution as original feature values** (as otherwise estimator may fail). The simplest way to get such noise is to shuffle values for a feature, i.e. use other examples’ feature values - this is how permutation importance is computed.
#
# ---
#
# It's not good when feature remove (replaced by noise) but we have better score. Simple and easy.
# +
########################### Lets test far away Lags (7 days with 56 days shift)
########################### and check permutation importance
#################################################################################
LAGS_SPLIT = [col for col in range(56,56+7)]
grid_df = pd.concat([grid_df, df_parallelize_run(make_normal_lag,LAGS_SPLIT)], axis=1)
test_model = make_fast_test(grid_df)
features_columns = [col for col in list(grid_df) if col not in remove_features]
validation_df = grid_df[grid_df['d']>(END_TRAIN-28)].reset_index(drop=True)
validation_df['preds'] = test_model.predict(validation_df[features_columns])
base_score = rmse(validation_df[TARGET], validation_df['preds'])
print('Standart RMSE', base_score)
for col in features_columns:
temp_df = validation_df.copy()
if temp_df[col].dtypes.name != 'category':
temp_df[col] = np.random.permutation(temp_df[col].values)
temp_df['preds'] = test_model.predict(temp_df[features_columns])
cur_score = rmse(temp_df[TARGET], temp_df['preds'])
print(col, np.round(cur_score - base_score, 4))
del temp_df, validation_df
# Remove test features
# As we will compare performance with baseline model for now
keep_cols = [col for col in list(grid_df) if 'sales_lag_' not in col]
grid_df = grid_df[keep_cols]
# Results:
## Lags with 56 days shift (far away past) are not as important
## as nearest past lags
## and at some point will be just noise for our model
# +
from sklearn.decomposition import PCA
def make_pca(df, pca_col, n_days):
print('PCA:', pca_col, n_days)
# We don't need any other columns to make pca
pca_df = df[[pca_col,'d',TARGET]]
# If we are doing pca for other series "levels"
# we need to agg first
if pca_col != 'id':
merge_base = pca_df[[pca_col,'d']]
pca_df = pca_df.groupby([pca_col,'d'])[TARGET].agg(['sum']).reset_index()
pca_df[TARGET] = pca_df['sum']
del pca_df['sum']
# Min/Max scaling
pca_df[TARGET] = pca_df[TARGET]/pca_df[TARGET].max()
# Making "lag" in old way (not parallel)
LAG_DAYS = [col for col in range(1,n_days+1)]
format_s = '{}_pca_'+pca_col+str(n_days)+'_{}'
pca_df = pca_df.assign(**{
format_s.format(col, l): pca_df.groupby([pca_col])[col].transform(lambda x: x.shift(l))
for l in LAG_DAYS
for col in [TARGET]
})
pca_columns = list(pca_df)[3:]
pca_df[pca_columns] = pca_df[pca_columns].fillna(0)
pca = PCA(random_state=SEED)
# You can use fit_transform here
pca.fit(pca_df[pca_columns])
pca_df[pca_columns] = pca.transform(pca_df[pca_columns])
print(pca.explained_variance_ratio_)
# we will keep only 3 most "valuable" columns/dimensions
keep_cols = pca_columns[:3]
print('Columns to keep:', keep_cols)
# If we are doing pca for other series "levels"
# we need merge back our results to merge_base df
# and only than return resulted df
# I'll skip that step here
return pca_df[keep_cols]
# Make PCA
grid_df = pd.concat([grid_df, make_pca(grid_df,'id',7)], axis=1)
# Make features test
test_model = make_fast_test(grid_df)
# Remove test features
# As we will compare performance with baseline model for now
keep_cols = [col for col in list(grid_df) if '_pca_' not in col]
grid_df = grid_df[keep_cols]
# +
########################### Mean/std target encoding
#################################################################################
# We will use these three columns for test
# (in combination with store_id)
icols = ['item_id','cat_id','dept_id']
# But we can use any other column or even multiple groups
# like these ones
# 'state_id',
# 'store_id',
# 'cat_id',
# 'dept_id',
# ['state_id', 'cat_id'],
# ['state_id', 'dept_id'],
# ['store_id', 'cat_id'],
# ['store_id', 'dept_id'],
# 'item_id',
# ['item_id', 'state_id'],
# ['item_id', 'store_id']
# There are several ways to do "mean" encoding
## K-fold scheme
## LOO (leave one out)
## Smoothed/regularized
## Expanding mean
## etc
# You can test as many options as you want
# and decide what to use
# Because of memory issues you can't
# use many features.
# We will use simple target encoding
# by std and mean agg
for col in icols:
print('Encoding', col)
temp_df = grid_df[grid_df['d']<=(1913-28)] # to be sure we don't have leakage in our validation set
temp_df = temp_df.groupby([col,'store_id']).agg({TARGET: ['std','mean']})
joiner = '_'+col+'_encoding_'
temp_df.columns = [joiner.join(col).strip() for col in temp_df.columns.values]
temp_df = temp_df.reset_index()
grid_df = grid_df.merge(temp_df, on=[col,'store_id'], how='left')
del temp_df
# Make features test
test_model = make_fast_test(grid_df)
# Remove test features
keep_cols = [col for col in list(grid_df) if '_encoding_' not in col]
grid_df = grid_df[keep_cols]
# Bad thing that for some items
# we are using past and future values.
# But we are looking for "categorical" similiarity
# on a "long run". So future here is not a big problem.
# +
########################### Last non O sale
#################################################################################
def find_last_sale(df,n_day):
# Limit initial df
ls_df = df[['id','d',TARGET]]
# Convert target to binary
ls_df['non_zero'] = (ls_df[TARGET]>0).astype(np.int8)
# Make lags to prevent any leakage
ls_df['non_zero_lag'] = ls_df.groupby(['id'])['non_zero'].transform(lambda x: x.shift(n_day).rolling(2000,1).sum()).fillna(-1)
temp_df = ls_df[['id','d','non_zero_lag']].drop_duplicates(subset=['id','non_zero_lag'])
temp_df.columns = ['id','d_min','non_zero_lag']
ls_df = ls_df.merge(temp_df, on=['id','non_zero_lag'], how='left')
ls_df['last_sale'] = ls_df['d'] - ls_df['d_min']
return ls_df[['last_sale']]
# Find last non zero
# Need some "dances" to fit in memory limit with groupers
grid_df = pd.concat([grid_df, find_last_sale(grid_df,1)], axis=1)
# Make features test
test_model = make_fast_test(grid_df)
# Remove test features
keep_cols = [col for col in list(grid_df) if 'last_sale' not in col]
grid_df = grid_df[keep_cols]
# +
########################### Apply on grid_df
#################################################################################
# lets read grid from
# https://www.kaggle.com/kyakovlev/m5-simple-fe
# to be sure that our grids are aligned by index
grid_df = pd.read_pickle('./output/grid_part_1.pkl')
grid_df[TARGET][grid_df['d']>(1913-28)] = np.nan
base_cols = list(grid_df)
icols = [
['state_id'],
['store_id'],
['cat_id'],
['dept_id'],
['state_id', 'cat_id'],
['state_id', 'dept_id'],
['store_id', 'cat_id'],
['store_id', 'dept_id'],
['item_id'],
['item_id', 'state_id'],
['item_id', 'store_id']
]
for col in icols:
print('Encoding', col)
col_name = '_'+'_'.join(col)+'_'
grid_df['enc'+col_name+'mean'] = grid_df.groupby(col)[TARGET].transform('mean').astype(np.float16)
grid_df['enc'+col_name+'std'] = grid_df.groupby(col)[TARGET].transform('std').astype(np.float16)
keep_cols = [col for col in list(grid_df) if col not in base_cols]
grid_df = grid_df[['id','d']+keep_cols]
# -
#################################################################################
print('Save Mean/Std encoding')
grid_df.to_pickle('./output/mean_encoding_df.pkl')
########################### Final list of new features
#################################################################################
grid_df.info()
| 03-other-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: hist
# language: python
# name: hist
# ---
# # Plots
# One of the most amazing feature of hist is it's powerful plotting family. Here you can see how to plot Hist.
from hist import Hist
import hist
h = Hist(
hist.axis.Regular(50, -5, 5, name="S", label="s [units]", flow=False),
hist.axis.Regular(50, -5, 5, name="W", label="w [units]", flow=False),
)
# +
import numpy as np
s_data = np.random.normal(size=100_000) + np.ones(100_000)
w_data = np.random.normal(size=100_000)
# normal fill
h.fill(s_data, w_data)
# -
# ## Via Matplotlib
#
# hist allows you to plot via [Matplotlib](https://matplotlib.org/) like this:
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(8, 5))
w, x, y = h.to_numpy()
mesh = ax.pcolormesh(x, y, w.T, cmap="RdYlBu")
ax.set_xlabel("s")
ax.set_ylabel("w")
fig.colorbar(mesh)
plt.show()
# ## Via Mplhep
#
# [mplhep](https://github.com/scikit-hep/mplhep) is an important visualization tools in Scikit-Hep ecosystem. hist has integrate with mplhep and you can also plot using it. If you want more info about mplhep please visit the official repo to see it.
# +
import mplhep
fig, axs = plt.subplots(1, 2, figsize=(9, 4))
mplhep.histplot(h.project("S"), ax=axs[0])
mplhep.hist2dplot(h, ax=axs[1])
plt.show()
# -
# ## Via Plot
#
# Hist has plotting methods for 1-D and 2-D histograms, `.plot1d()` and `.plot2d()` respectively. It also provides `.plot()` for plotting according to the its dimension. Moreover, to show the projection of each axis, you can use `.plot2d_full()`. If you have a Hist with higher dimension, you can use `.project()` to extract two dimensions to see it with our plotting suite.
#
# Our plotting methods are all based on Matplotlib, so you can pass Matplotlib's `ax` into it, and hist will draw on it. We will create it for you if you do not pass them in.
# +
# plot1d
fig, ax = plt.subplots(figsize=(6, 4))
h.project("S").plot1d(ax=ax, ls="--", color="teal", lw=3)
plt.show()
# +
# plot2d
fig, ax = plt.subplots(figsize=(6, 6))
h.plot2d(ax=ax, cmap="plasma")
plt.show()
# +
# plot2d_full
plt.figure(figsize=(8, 8))
h.plot2d_full(
main_cmap="coolwarm",
top_ls="--",
top_color="orange",
top_lw=2,
side_ls=":",
side_lw=2,
side_color="steelblue",
)
plt.show()
# +
# auto-plot
fig, axs = plt.subplots(1, 2, figsize=(9, 4), gridspec_kw={"width_ratios": [5, 4]})
h.project("W").plot(ax=axs[0], color="darkviolet", lw=2, ls="-.")
h.project("W", "S").plot(ax=axs[1], cmap="cividis")
plt.show()
# -
# ## Via Plot Pull
#
# Pull plots are commonly used in HEP studies, and we provide a method for them with `.plot_pull()`, which accepts a `Callable` object, like the below `pdf` function, which is then fit to the histogram and the fit and pulls are shown on the plot. As Normal distributions are the generally desired function to fit the histogram data, the `str` aliases `"normal"`, `"gauss"`, and `"gaus"` are supported as well.
def pdf(x, a=1 / np.sqrt(2 * np.pi), x0=0, sigma=1, offset=0):
return a * np.exp(-((x - x0) ** 2) / (2 * sigma ** 2)) + offset
# +
np.random.seed(0)
hist_1 = hist.Hist(
hist.axis.Regular(
50, -5, 5, name="X", label="x [units]", underflow=False, overflow=False
)
).fill(np.random.normal(size=1000))
fig = plt.figure(figsize=(10, 8))
main_ax_artists, sublot_ax_arists = hist_1.plot_pull(
"normal",
eb_ecolor="steelblue",
eb_mfc="steelblue",
eb_mec="steelblue",
eb_fmt="o",
eb_ms=6,
eb_capsize=1,
eb_capthick=2,
eb_alpha=0.8,
fp_c="hotpink",
fp_ls="-",
fp_lw=2,
fp_alpha=0.8,
bar_fc="royalblue",
pp_num=3,
pp_fc="royalblue",
pp_alpha=0.618,
pp_ec=None,
ub_alpha=0.2,
)
# -
# ## Via Plot Ratio
#
# You can also make an arbitrary ratio plot using the `.plot_ratio` API:
# +
hist_2 = hist.Hist(
hist.axis.Regular(
50, -5, 5, name="X", label="x [units]", underflow=False, overflow=False
)
).fill(np.random.normal(size=1700))
fig = plt.figure(figsize=(10, 8))
main_ax_artists, sublot_ax_arists = hist_1.plot_ratio(
hist_2,
rp_ylabel=r"Ratio",
rp_num_label="hist1",
rp_denom_label="hist2",
rp_uncert_draw_type="bar", # line or bar
)
# -
# Ratios between the histogram and a callable, or `str` alias, are supported as well
fig = plt.figure(figsize=(10, 8))
main_ax_artists, sublot_ax_arists = hist_1.plot_ratio(pdf)
# Using the `.plot_ratio` API you can also make efficiency plots (where the numerator is a strict subset of the denominator)
# +
hist_3 = hist_2.copy() * 0.7
hist_2.fill(np.random.uniform(-5, 5, 600))
hist_3.fill(np.random.uniform(-5, 5, 200))
fig = plt.figure(figsize=(10, 8))
main_ax_artists, sublot_ax_arists = hist_3.plot_ratio(
hist_2,
rp_num_label="hist3",
rp_denom_label="hist2",
rp_uncert_draw_type="line",
rp_uncertainty_type="efficiency",
)
| docs/user-guide/notebooks/Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow)
# language: python
# name: rga
# ---
# + [markdown] id="bV4rrxPA81rc" colab_type="text"
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_02_1_python_pandas.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="cYuojwvr81rh" colab_type="text"
# # T81-558: Applications of Deep Neural Networks
# **Module 2: Python for Machine Learning**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# + [markdown] id="KcvWuMJN81rl" colab_type="text"
# # Module 2 Material
#
# Main video lecture:
#
# * **Part 2.1: Introduction to Pandas** [[Video]](https://www.youtube.com/watch?v=bN4UuCBdpZc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_1_python_pandas.ipynb)
# * Part 2.2: Categorical Values [[Video]](https://www.youtube.com/watch?v=4a1odDpG0Ho&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_2_pandas_cat.ipynb)
# * Part 2.3: Grouping, Sorting, and Shuffling in Python Pandas [[Video]](https://www.youtube.com/watch?v=YS4wm5gD8DM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_3_pandas_grouping.ipynb)
# * Part 2.4: Using Apply and Map in Pandas for Keras [[Video]](https://www.youtube.com/watch?v=XNCEZ4WaPBY&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_4_pandas_functional.ipynb)
# * Part 2.5: Feature Engineering in Pandas for Deep Learning in Keras [[Video]](https://www.youtube.com/watch?v=BWPTj4_Mi9E&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_5_pandas_features.ipynb)
# + [markdown] id="Qe7E5Kx581rn" colab_type="text"
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
# + id="GESDR1P581rq" colab_type="code" outputId="d1ff6f1f-e246-4aeb-af6e-8e2ee2a188f3" colab={}
try:
from google.colab import drive
# %tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
# + [markdown] id="OpOIZHO781r_" colab_type="text"
# # Part 2.1: Introduction to Pandas
#
# [Pandas](http://pandas.pydata.org/) is an open-source library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language. It is based on the [dataframe](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html) concept found in the [R programming language](https://www.r-project.org/about.html). For this class, Pandas will be the primary means by which we manipulate data to be processed by neural networks.
#
# The data frame is a crucial component of Pandas. We will use it to access the [auto-mpg dataset](https://archive.ics.uci.edu/ml/datasets/Auto+MPG). You can find this dataset on the UCI machine learning repository. For this class, we will use a version of the Auto MPG dataset, where I added column headers. You can find my version [here](https://raw.githubusercontent.com/jeffheaton/t81_558_deep_learning/master/data/auto-mpg.csv).
#
# UCI took this dataset from the StatLib library, which Carnegie Mellon University maintains. The dataset was used in the 1983 American Statistical Association Exposition. It contains data for 398 cars, including [mpg](https://en.wikipedia.org/wiki/Fuel_economy_in_automobiles), [cylinders](https://en.wikipedia.org/wiki/Cylinder_(engine)), [displacement](https://en.wikipedia.org/wiki/Engine_displacement), [horsepower](https://en.wikipedia.org/wiki/Horsepower) , weight, acceleration, model year, origin and the car's name.
#
# The following code loads the MPG dataset into a data frame:
# + id="If1Lr7GD81sC" colab_type="code" outputId="f71e1c5b-0011-4d32-e04c-8cae35f2fb01" colab={}
# Simple dataframe
import os
import pandas as pd
df = pd.read_csv("https://data.heatonresearch.com/data/t81-558/auto-mpg.csv")
print(df[0:5])
# + [markdown] id="64iauSv481sM" colab_type="text"
# The **display** function provides a cleaner display than merely printing the data frame. Specifying the maximum rows and columns allows you to achieve greater control over the display.
# + id="jmPZwrgz81sO" colab_type="code" outputId="b4a99bf5-eaa3-42bb-c937-486c2b751221" colab={}
pd.set_option('display.max_columns', 7)
pd.set_option('display.max_rows', 5)
display(df)
# + [markdown] id="uuGouA8n81sa" colab_type="text"
# It is possible to generate a second data frame to display statistical information about the first data frame.
# + id="7fsKpRhH81sb" colab_type="code" outputId="cff6afd0-3e4b-4276-919e-b80c86ba675d" colab={}
# Strip non-numerics
df = df.select_dtypes(include=['int', 'float'])
headers = list(df.columns.values)
fields = []
for field in headers:
fields.append({
'name' : field,
'mean': df[field].mean(),
'var': df[field].var(),
'sdev': df[field].std()
})
for field in fields:
print(field)
# + [markdown] id="yArt4G7481sn" colab_type="text"
# This code outputs a list of dictionaries that hold this statistical information. This information looks similar to the JSON code seen in Module 1. To as proper JSON, the program should add these records to a list and call the Python JSON library's **dumps** command called.
#
# The Python program can convert this JSON-like information to a data frame for better display.
# + id="DDXCdUPI81so" colab_type="code" outputId="bbb8b857-04e1-49c8-dd3e-0d60824751bd" colab={}
pd.set_option('display.max_columns', 0)
pd.set_option('display.max_rows', 0)
df2 = pd.DataFrame(fields)
display(df2)
# + [markdown] id="TPVE3lC781s1" colab_type="text"
# ## Missing Values
#
# Missing values are a reality of machine learning. Ideally, every row of data will have values for all columns. However, this is rarely the case. Most of the values are present in the MPG database. However, there are missing values in the horsepower column. A common practice is to replace missing values with the median value for that column. The program calculates the median as described [here](https://www.mathsisfun.com/median.html). The following code replaces any NA values in horsepower with the median:
# + id="aUPu9Cfa81s2" colab_type="code" outputId="a91b10ff-d10a-4747-ec4b-3e1e9afb5880" colab={}
import os
import pandas as pd
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA', '?'])
print(f"horsepower has na? {pd.isnull(df['horsepower']).values.any()}")
print("Filling missing values...")
med = df['horsepower'].median()
df['horsepower'] = df['horsepower'].fillna(med)
# df = df.dropna() # you can also simply drop NA values
print(f"horsepower has na? {pd.isnull(df['horsepower']).values.any()}")
# + [markdown] id="-RRRwwtJ81s_" colab_type="text"
# # Dealing with Outliers
#
# Outliers are values that are unusually high or low. Sometimes outliers are simply errors; this is a result of [observation error](https://en.wikipedia.org/wiki/Observational_error). Outliers can also be truly large or small values that may be difficult to address. We typically consider outliers to be a value that is several standard deviations from the mean. The following function can remove such values.
# + id="1XCKA0kf81tA" colab_type="code" colab={}
# Remove all rows where the specified column is +/- sd standard deviations
def remove_outliers(df, name, sd):
drop_rows = df.index[(np.abs(df[name] - df[name].mean())
>= (sd * df[name].std()))]
df.drop(drop_rows, axis=0, inplace=True)
# + [markdown] id="_3-5WLkO81tK" colab_type="text"
# The code below will drop every row from the Auto MPG dataset where the horsepower is more than two standard deviations above or below the mean.
# + id="CI0GpTwy81tL" colab_type="code" outputId="7a590fb4-1a4e-4233-8071-44c2e0c7cd2b" colab={}
import pandas as pd
import os
import numpy as np
from sklearn import metrics
from scipy.stats import zscore
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
# create feature vector
med = df['horsepower'].median()
df['horsepower'] = df['horsepower'].fillna(med)
# Drop the name column
df.drop('name',1,inplace=True)
# Drop outliers in horsepower
print("Length before MPG outliers dropped: {}".format(len(df)))
remove_outliers(df,'mpg',2)
print("Length after MPG outliers dropped: {}".format(len(df)))
pd.set_option('display.max_columns', 0)
pd.set_option('display.max_rows', 5)
display(df)
# + [markdown] id="LA2bMO9y81tb" colab_type="text"
# ## Dropping Fields
#
# Some fields are of no value to the neural network should be dropped. The following code removes the name column from the MPG dataset.
# + id="ZReuehj181tc" colab_type="code" outputId="99211ed2-6652-4a24-eee4-5fb19225b988" colab={}
import os
import pandas as pd
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
print(f"Before drop: {list(df.columns)}")
df.drop('name', 1, inplace=True)
print(f"After drop: {list(df.columns)}")
# + [markdown] id="ReT4dr1X81tk" colab_type="text"
# ## Concatenating Rows and Columns
# Python can concatenate rows and columns together to form new data frames. The code below creates a new data frame from the **name** and **horsepower** columns from the Auto MPG dataset. The program does this by concatenating two columns together.
# + id="AqBgIV4z81tm" colab_type="code" outputId="f2a2a16f-4506-40c5-b471-8a8ab052fbc5" colab={}
# Create a new dataframe from name and horsepower
import os
import pandas as pd
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
col_horsepower = df['horsepower']
col_name = df['name']
result = pd.concat([col_name, col_horsepower], axis=1)
pd.set_option('display.max_columns', 0)
pd.set_option('display.max_rows', 5)
display(result)
# + [markdown] id="1S4apubZ81tv" colab_type="text"
# The **concat** function can also concatenate two rows together. This code concatenates the first two rows and the last two rows of the Auto MPG dataset.
# + id="LyMIGJo481tx" colab_type="code" outputId="7cdcfd11-b0d3-42dc-98e8-602075aaf08f" colab={}
# Create a new dataframe from first 2 rows and last 2 rows
import os
import pandas as pd
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
result = pd.concat([df[0:2],df[-2:]], axis=0)
pd.set_option('display.max_columns', 7)
pd.set_option('display.max_rows', 0)
display(result)
# + [markdown] id="UBhQClko81t4" colab_type="text"
# ## Training and Validation
#
# We must evaluate a machine learning model based on its ability to predict data that it has never seen before. Because of this, we often divide the training data into a validation and training set. The machine learning model will learn from the training data, but ultimately be evaluated based on the validation data.
#
# * **Training Data** - **In Sample Data** - The data that the neural network used to train.
# * **Validation Data** - **Out of Sample Data** - The data that the machine learning model is evaluated upon after it is fit to the training data.
#
# There are two effective means of dealing with training and validation data:
#
# * **Training/Validation Split** - The program splits the data according to some ratio between a training and validation (hold-out) set. Typical rates are 80% training and 20% validation.
# * **K-Fold Cross Validation** - The program splits the data into several folds and models. Because the program creates the same number of models as folds, the program can generate out-of-sample predictions for the entire dataset.
#
# The code below performs a split of the MPG data into a training and validation set. The training set uses 80% of the data, and the validation set uses 20%. Figure 2.TRN-VAL shows how a model is trained on 80% of the data and then validated against the remaining 20%.
#
# **Figure 2.TRN-VAL: Training and Validation**
# 
#
# + id="RImVx1TU81t5" colab_type="code" outputId="f86664c8-2999-4ad7-a608-7f2a772ff61c" colab={}
import os
import pandas as pd
import numpy as np
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
# Usually a good idea to shuffle
df = df.reindex(np.random.permutation(df.index))
mask = np.random.rand(len(df)) < 0.8
trainDF = pd.DataFrame(df[mask])
validationDF = pd.DataFrame(df[~mask])
print(f"Training DF: {len(trainDF)}")
print(f"Validation DF: {len(validationDF)}")
# + [markdown] id="ADdJHTJY81uC" colab_type="text"
# ### Converting a Dataframe to a Matrix
#
# Neural networks do not directly operate on Python data frames. A neural network requires a numeric matrix. The program uses the **values** property of a data frame to convert the data to a matrix.
# + id="fY15eank81uD" colab_type="code" outputId="be9ca42a-e1d5-45f0-e439-af8bc14a5d5c" colab={}
df.values
# + [markdown] id="iwh_immk81uK" colab_type="text"
# You might wish only to convert some of the columns, to leave out the name column, use the following code.
# + id="4gtMITPl81uL" colab_type="code" outputId="8c832183-14c7-4cb9-834b-6e2a81f352d2" colab={}
df[['mpg', 'cylinders', 'displacement', 'horsepower', 'weight',
'acceleration', 'year', 'origin']].values
# + [markdown] id="Sj-1GGaZ81uS" colab_type="text"
# ## Saving a Dataframe to CSV
#
# Many of the assignments in this course will require that you save a data frame to submit to the instructor. The following code performs a shuffle and then saves a new copy.
# + id="LZSpm1EC81uS" colab_type="code" outputId="da3d78a7-e6b7-43a3-9fb8-44a2f6987b6d" colab={}
import os
import pandas as pd
import numpy as np
path = "."
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
filename_write = os.path.join(path, "auto-mpg-shuffle.csv")
df = df.reindex(np.random.permutation(df.index))
# Specify index = false to not write row numbers
df.to_csv(filename_write, index=False)
print("Done")
# + [markdown] id="ZUVpSwYr81ub" colab_type="text"
# ## Saving a Dataframe to Pickle
#
# A variety of software programs can make use of text files stored as CSV. However, they do take longer to generate and can sometimes lose small amounts of precision in the conversion. Another format is [Pickle](https://docs.python.org/3/library/pickle.html). Generally, you will output to CSV because it is very compatible, even outside of Python. The code below stores the Dataframe to Pickle.
# + id="XnP3z3Xb81uc" colab_type="code" colab={}
import os
import pandas as pd
import numpy as np
import pickle
path = "."
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
filename_write = os.path.join(path, "auto-mpg-shuffle.pkl")
df = df.reindex(np.random.permutation(df.index))
with open(filename_write,"wb") as fp:
pickle.dump(df, fp)
# + [markdown] id="Ah4Q4mCc81uh" colab_type="text"
# Loading the pickle file back into memory is accomplished by the following lines of code. Notice that the index numbers are still jumbled from the previous shuffle? Loading the CSV rebuilt (in the last step) did not preserve these values.
# + id="mFwJW6sz81uh" colab_type="code" outputId="547bad46-dc81-460e-8d6c-85088b4ff2f1" colab={}
import os
import pandas as pd
import numpy as np
import pickle
path = "."
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
filename_read = os.path.join(path, "auto-mpg-shuffle.pkl")
with open(filename_write,"rb") as fp:
df = pickle.load(fp)
pd.set_option('display.max_columns', 7)
pd.set_option('display.max_rows', 5)
display(df)
# + [markdown] id="HFrK1yhp81uo" colab_type="text"
# # Module 2 Assignment
#
# You can find the first assignment here: [assignment 2](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class2.ipynb)
# + id="mfGwMz-i81ut" colab_type="code" colab={}
| t81_558_class_02_1_python_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Building a Recommender System with Amazon SageMaker Factorization Machines and BlazingText
#
# ---
#
# ---
#
# ## Background
#
# - Recommender systems were a catalyst for ML's popularity (Amazon, Netflix Prize)
# - User item matrix factorization is a core methodology
# - Factorization machines combine linear prediction with a factorized representation of pairwise feature interaction
#
# $$\hat{r} = w_0 + \sum_{i} {w_i x_i} + \sum_{i} {\sum_{j > i} {\langle v_i, v_j \rangle x_i x_j}}$$
#
# - SageMaker has a highly scalable factorization machines algorithm built-in
# - To learn more about the math behind _factorization machines_, [this paper](https://www.csie.ntu.edu.tw/~b97053/paper/Rendle2010FM.pdf) is a great resource
#
# ---
#
# ## Setup
#
# 1. Spin up SageMaker hosted notebook instance in console
# 2. Add SageMaker IAM policy to this SageMaker notebook to allow S3 read/write access
# 3. Create new S3 bucket (first cell)
# 4. Import necessary libraries (second cell)
# +
import sagemaker
sess = sagemaker.Session()
bucket = sess.default_bucket()
base = 'DEMO-loft-recommender'
prefix = 'sagemaker/' + base
role = sagemaker.get_execution_role()
# -
import sagemaker
import os
import pandas as pd
import numpy as np
import boto3
import json
import io
import matplotlib.pyplot as plt
import sagemaker.amazon.common as smac
from sagemaker.predictor import json_deserializer
from scipy.sparse import csr_matrix
# ---
#
# ## Data
#
# [Amazon Reviews AWS Public Dataset](https://s3.amazonaws.com/amazon-reviews-pds/readme.html)
# - 1 to 5 star ratings
# - 2M+ Amazon customers
# - 160K+ digital videos
# !mkdir /tmp/recsys/
# !aws s3 cp s3://amazon-reviews-pds/tsv/amazon_reviews_us_Digital_Video_Download_v1_00.tsv.gz /tmp/recsys/
df = pd.read_csv('/tmp/recsys/amazon_reviews_us_Digital_Video_Download_v1_00.tsv.gz', delimiter='\t',error_bad_lines=False)
df.head()
# Dataset columns:
#
# - `marketplace`: 2-letter country code (in this case all "US").
# - `customer_id`: Random identifier that can be used to aggregate reviews written by a single author.
# - `review_id`: A unique ID for the review.
# - `product_id`: The Amazon Standard Identification Number (ASIN). `http://www.amazon.com/dp/<ASIN>` links to the product's detail page.
# - `product_parent`: The parent of that ASIN. Multiple ASINs (color or format variations of the same product) can roll up into a single parent parent.
# - `product_title`: Title description of the product.
# - `product_category`: Broad product category that can be used to group reviews (in this case digital videos).
# - `star_rating`: The review's rating (1 to 5 stars).
# - `helpful_votes`: Number of helpful votes for the review.
# - `total_votes`: Number of total votes the review received.
# - `vine`: Was the review written as part of the [Vine](https://www.amazon.com/gp/vine/help) program?
# - `verified_purchase`: Was the review from a verified purchase?
# - `review_headline`: The title of the review itself.
# - `review_body`: The text of the review.
# - `review_date`: The date the review was written.
#
# Drop some fields that won't be used
df = df[['customer_id', 'product_id', 'product_title', 'star_rating', 'review_date']]
# Most users don't rate most movies - Check our long tail
# +
customers = df['customer_id'].value_counts()
products = df['product_id'].value_counts()
quantiles = [0, 0.01, 0.02, 0.03, 0.04, 0.05, 0.1, 0.25, 0.5, 0.75, 0.9, 0.95, 0.96, 0.97, 0.98, 0.99, 1]
print('customers\n', customers.quantile(quantiles))
print('products\n', products.quantile(quantiles))
# -
# Filter out customers who haven't rated many movies
# +
customers = customers[customers >= 5]
products = products[products >= 10]
reduced_df = df.merge(pd.DataFrame({'customer_id': customers.index})).merge(pd.DataFrame({'product_id': products.index}))
# -
# Create a sequential index for customers and movies
customers = reduced_df['customer_id'].value_counts()
products = reduced_df['product_id'].value_counts()
# +
customer_index = pd.DataFrame({'customer_id': customers.index, 'user': np.arange(customers.shape[0])})
product_index = pd.DataFrame({'product_id': products.index,
'item': np.arange(products.shape[0]) + customer_index.shape[0]})
reduced_df = reduced_df.merge(customer_index).merge(product_index)
reduced_df.head()
# -
# Count days since first review (included as a feature to capture trend)
reduced_df['review_date'] = pd.to_datetime(reduced_df['review_date'])
customer_first_date = reduced_df.groupby('customer_id')['review_date'].min().reset_index()
customer_first_date.columns = ['customer_id', 'first_review_date']
reduced_df = reduced_df.merge(customer_first_date)
reduced_df['days_since_first'] = (reduced_df['review_date'] - reduced_df['first_review_date']).dt.days
reduced_df['days_since_first'] = reduced_df['days_since_first'].fillna(0)
# Split into train and test datasets
# +
test_df = reduced_df.groupby('customer_id').last().reset_index()
train_df = reduced_df.merge(test_df[['customer_id', 'product_id']],
on=['customer_id', 'product_id'],
how='outer',
indicator=True)
train_df = train_df[(train_df['_merge'] == 'left_only')]
# -
# - Factorization machines expects data to look something like:
# - Sparse matrix
# - Target variable is that user's rating for a movie
# - One-hot encoding for users ($N$ features)
# - One-hot encoding for movies ($M$ features)
#
# |Rating|User1|User2|...|UserN|Movie1|Movie2|Movie3|...|MovieM|Feature1|Feature2|...|
# |---|---|---|---|---|---|---|---|---|---|---|---|---|
# |4|1|0|...|0|1|0|0|...|0|20|2.2|...|
# |5|1|0|...|0|0|1|0|...|0|17|9.1|...|
# |3|0|1|...|0|1|0|0|...|0|3|11.0|...|
# |4|0|1|...|0|0|0|1|...|0|15|6.4|...|
#
#
# - Wouldn't want to hold this full matrix in memory
# - Create a sparse matrix
# - Designed to work efficiently with CPUs. Some parts of training for more dense matrices can be parallelized with GPUs
def to_csr_matrix(df, num_users, num_items):
feature_dim = num_users + num_items + 1
data = np.concatenate([np.array([1] * df.shape[0]),
np.array([1] * df.shape[0]),
df['days_since_first'].values])
row = np.concatenate([np.arange(df.shape[0])] * 3)
col = np.concatenate([df['user'].values,
df['item'].values,
np.array([feature_dim - 1] * df.shape[0])])
return csr_matrix((data, (row, col)),
shape=(df.shape[0], feature_dim),
dtype=np.float32)
train_csr = to_csr_matrix(train_df, customer_index.shape[0], product_index.shape[0])
test_csr = to_csr_matrix(test_df, customer_index.shape[0], product_index.shape[0])
# Convert to sparse recordIO-wrapped protobuf that SageMaker factorization machines expects
def to_s3_protobuf(csr, label, bucket, prefix, channel='train', splits=10):
indices = np.array_split(np.arange(csr.shape[0]), splits)
for i in range(len(indices)):
index = indices[i]
buf = io.BytesIO()
smac.write_spmatrix_to_sparse_tensor(buf, csr[index, ], label[index])
buf.seek(0)
boto3.client('s3').upload_fileobj(buf, bucket, '{}/{}/data-{}'.format(prefix, channel, i))
to_s3_protobuf(train_csr, train_df['star_rating'].values.astype(np.float32), bucket, prefix)
to_s3_protobuf(test_csr, test_df['star_rating'].values.astype(np.float32), bucket, prefix, channel='test', splits=1)
# ---
#
# ## Train
#
# - Create a [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk) estimator to run a training jobs and specify:
# - Algorithm container image
# - IAM role
# - Hardware setup
# - S3 output location
# - Algorithm hyperparameters
# - `feature_dim`: $N + M + 1$ (additional feature is `days_since_first` to capture trend)
# - `num_factors`: number of factor dimensions (increasing too much can lead to overfitting)
# - `epochs`: number of full passes through the dataset
# - `.fit()` points to training and test data in S3 and begins the training job
#
# **Note**: For AWS accounts registered in conjunction with a workshop, default instance limits may prevent the use of `ml.c5.2xlarge` (and other equally powerful instances), and may require a lower value for `train_instance_count` depending on the instance type chosen.
# +
fm = sagemaker.estimator.Estimator(
sagemaker.amazon.amazon_estimator.get_image_uri(boto3.Session().region_name, 'factorization-machines', 'latest'),
role,
train_instance_count=1, # Note: instance numbers may be limited on workshop credits
train_instance_type='ml.c5.large', # Note:'ml.c5.2xlarge' may not be available on workshop credits
output_path='s3://{}/{}/output'.format(bucket, prefix),
base_job_name=base,
sagemaker_session=sess)
fm.set_hyperparameters(
feature_dim=customer_index.shape[0] + product_index.shape[0] + 1,
predictor_type='regressor',
mini_batch_size=1000,
num_factors=256,
epochs=3)
fm.fit({'train': sagemaker.s3_input('s3://{}/{}/train/'.format(bucket, prefix), distribution='ShardedByS3Key'),
'test': sagemaker.s3_input('s3://{}/{}/test/'.format(bucket, prefix), distribution='FullyReplicated')})
# -
# ---
#
# ## Host
#
# Deploy trained model to a real-time production endpoint
fm_predictor = fm.deploy(instance_type='ml.m4.xlarge', initial_instance_count=1)
# Setup predictor to serialize in-memory data for invocation requests
def fm_serializer(df):
feature_dim = customer_index.shape[0] + product_index.shape[0] + 1
js = {'instances': []}
for index, data in df.iterrows():
js['instances'].append({'data': {'features': {'values': [1, 1, data['days_since_first']],
'keys': [data['user'], data['item'], feature_dim - 1],
'shape': [feature_dim]}}})
return json.dumps(js)
fm_predictor.content_type = 'application/json'
fm_predictor.serializer = fm_serializer
fm_predictor.deserializer = json_deserializer
# **Real-time prediction for what a single user would rate an item**
#
# 1. Pick a customer-movie pair from the dataset
test_df.head(25)
# 2. Pull out a single customer-movie pair that we like
test_customer = test_df.iloc[[20]]
test_df.iloc[[20]] # peek at the data to confirm it's the one we wanted
# 3. Pass `test_customer` to predictor
fm_predictor.predict(test_customer)
# **Now let's make a df for an arbitrary customer and movie pair and test it out!**
#
# Our `fm_serializer` requires 3 inputs to perform a prediction:
# - `user` id for a customer (type = num)
# - `item` id for a movie (type = num)
# - `days_since_first` review (type = double)
# +
fake_customer = test_customer # make a copy of the test_customer we pulled out before to modify
desired_user_id = 65884 # person who rated Dexter with 5 stars
desired_item_id = 140461 # Code for True Blood: Season 1
desired_review_days = 28.0 # arbitrary number of days since first review
#fake_customer_data = {'user' : desired_user_id, 'item' : desired_item_id, 'days_since_first' : desired_review_days}
#fake_customer = pd.DataFrame(fake_customer_data, index=[0])
fake_customer['user'] = desired_user_id
fake_customer['item'] = desired_item_id
fake_customer['days_since_first'] = desired_review_days
# print the details for this fake customer
fake_customer
# -
fm_predictor.predict(fake_customer)
# Final step: Clean-up the endpoint
fm_predictor.delete_endpoint()
# ## Finished?
#
# Got some extra time? Feel free to go on to the Extra Credit below!
#
# **Note**: Amazon SageMaker automatically handles provisioning and tearing down of resources during training. Once deployed, the model's endpoint will persist independent of this notebook, and can be removed with the cell directly above this.
#
# If you are done working with this notebook demo, it is strongly advised that you stop the SageMaker hosted notebook instance if you do not wish to continue using it (and incurring costs). This can easily be done by clicking on "Notebook instances" from the SageMaker console.
# ---
#
# ---
#
# # Extra credit
#
# - What happens when a new movie is added?
# - No feature to set to "1" in the dataset
# - No previous ratings to find similar items
# - Cold start problem is hard with factorization machines
# - Word2vec
# - Word embeddings for natural language processing (similar words get similar vectors)
# - Use concatenated product titles as words, customer review history as sentences
# - SageMaker BlazingText is an extremely fast implementation that can work with subwords
# ---
#
# ## Data
#
# Concatenate product titles to treat each one as a single word
reduced_df['product_title'] = reduced_df['product_title'].apply(lambda x: x.lower().replace(' ', '-'))
# Write customer purchase histories
first = True
with open('customer_purchases.txt', 'w') as f:
for customer, data in reduced_df.sort_values(['customer_id', 'review_date']).groupby('customer_id'):
if first:
first = False
else:
f.write('\n')
f.write(' '.join(data['product_title'].tolist()))
# Write to S3 so SageMaker training can use it
inputs = sess.upload_data('customer_purchases.txt', bucket, '{}/word2vec/train'.format(prefix))
# ---
#
# ## Train
#
# Create a SageMaker estimator:
# - Specify training job arguments
# - Set hyperparameters
# - Remove titles that occur less than 5 times
# - Embed in a 100-dimensional subspace
# - Use subwords to capture similarity in titles
# +
bt = sagemaker.estimator.Estimator(
sagemaker.amazon.amazon_estimator.get_image_uri(boto3.Session().region_name, 'blazingtext', 'latest'),
role,
train_instance_count=1,
train_instance_type='ml.p3.2xlarge',
train_volume_size = 5,
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sess)
bt.set_hyperparameters(mode="skipgram",
epochs=10,
min_count=5,
sampling_threshold=0.0001,
learning_rate=0.05,
window_size=5,
vector_dim=100,
negative_samples=5,
min_char=5,
max_char=10,
evaluation=False,
subwords=True)
bt.fit({'train': sagemaker.s3_input(inputs, distribution='FullyReplicated', content_type='text/plain')})
# -
# ---
#
# ## Model
#
# - Bring in and extract the model from S3
# - Take a look at the embeddings
# !aws s3 cp $bt.model_data ./
# !tar -xvzf model.tar.gz
vectors = pd.read_csv('vectors.txt', delimiter=' ', skiprows=2, header=None)
# Do the embeddings appear to have meaning
vectors.sort_values(1)
vectors.sort_values(2)
# Yes, but there's 100. Let's reduce this further with t-SNE and map the top 100 titles.
product_titles = vectors[0]
vectors = vectors.drop([0, 101], axis=1)
# +
from sklearn.manifold import TSNE
tsne = TSNE(perplexity=40, n_components=2, init='pca', n_iter=10000)
embeddings = tsne.fit_transform(vectors.values[:100, ])
# +
from matplotlib import pylab
# %matplotlib inline
def plot(embeddings, labels):
pylab.figure(figsize=(20,20))
for i, label in enumerate(labels):
x, y = embeddings[i,:]
pylab.scatter(x, y)
pylab.annotate(label, xy=(x, y), xytext=(5, 2), textcoords='offset points',
ha='right', va='bottom')
pylab.show()
plot(embeddings, product_titles[:100])
# -
# ---
#
# ## Host
#
# Deploy our model to a real-time endpoint.
bt_endpoint = bt.deploy(initial_instance_count = 1,instance_type = 'ml.m4.xlarge')
# Try generating predictions for a set of titles (some of which are real, some of which are made up).
# +
words = ["sherlock-season-1",
"sherlock-season-2",
"sherlock-season-5",
'arbitrary-sherlock-holmes-string',
'the-imitation-game',
"abcdefghijklmn",
"keeping-up-with-the-kardashians-season-1"]
payload = {"instances" : words}
response = bt_endpoint.predict(json.dumps(payload))
vecs_df = pd.DataFrame(json.loads(response))
# -
# Calculate correlation and distance.
vecs_df = pd.DataFrame(vecs_df['vector'].values.tolist(), index=vecs_df['word'])
vecs_df = vecs_df.transpose()
vecs_df.corr()
for column in vecs_df.columns:
print(column + ':', np.sum((vecs_df[column] - vecs_df['sherlock-season-1']) ** 2))
# Relative to 'sherlock-season-1':
# - 'sherlock-season-5' is made up, but relates well with 'sherlock-season-1' and 'sherlock-season-2'
# - 'arbitrary-sherlock-holmes-string' is also made up and relates less well but still fairly strong
# - 'the-imitation-game' is another popular Prime video title starring <NAME> and has a moderate relationship, but worse than the arbitrary Sherlock title
# - 'abcdefghijklmn' is made up and relates even worse
# - 'keeping-up-with-the-kardashians-season-1' somehow manages to relate even worse
#
# Clean-up the endpoint
bt_endpoint.delete_endpoint()
# ---
#
# ---
#
# # Wrap-up
#
# - Built a recommender system on a large dataset quickly and accurately
# - Add more features to extend
# - Compare to other methods
# - Ensemble two models
| fm_amazon_recommender.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="3e382d4ce5da1553d0578a6ad1ca1cf2b9995104" _cell_guid="765cada3-8dfd-4ce3-8c55-a07a0c4426f4"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
from subprocess import check_output
print(check_output(["ls", "../input"]).decode("utf8"))
# Any results you write to the current directory are saved as output.
# + _uuid="0787b3bd26da9abfa775c8d6b6896770550263f5" _cell_guid="d7c4e3b9-0a72-4735-8b3c-bd99f2f7287d"
df = pd.read_csv('../input/train.csv')
df_test = pd.read_csv('../input/test.csv')
# + _uuid="aed749a61c7434d4fa3cbdbdb362034b1f10f8d5" _cell_guid="101017a5-4b80-460f-a348-7f3bb25bb90f"
df.head()
# + _uuid="fc30b7323e0709b36f2ddf412b5decc82dd4025e" _cell_guid="ad6a2294-9df1-43c1-bd92-5ad258875c8f"
del df['id']
train = df.as_matrix()
# + _uuid="e46c468a7947e3d1e480c8f51acb764eb79f1f93" _cell_guid="c7e7f37c-9c32-479a-887e-c9e4533fee1b"
from sklearn.naive_bayes import MultinomialNB
from sklearn.feature_extraction.text import CountVectorizer,TfidfVectorizer
from sklearn.neural_network import MLPClassifier
import xgboost
# + _uuid="b4bae82e160c1f85eb417443ccadc4bbebb6e358" _cell_guid="c637ea9b-5764-4396-98d2-7cf3434e3303"
from sklearn.feature_extraction import text
st_wd = text.ENGLISH_STOP_WORDS
c_vector = CountVectorizer(stop_words = st_wd,min_df=.000001,lowercase=1)
c_vector.fit(pd.concat((df['text'],df_test['text']),axis=0))
X_train_counts = c_vector.transform(df['text'].values)
# + _uuid="a44e0eb813268484299dae210b11a08bc15ad1df" _cell_guid="d1c43aba-360c-41ef-b2de-327df09d3245"
X_train_counts
# + _uuid="e29d38fdccc9812cf9d69a08414cadaec570ada1" _cell_guid="79900383-380d-4f43-b598-0a5b8ddd7a53"
dic = {'EAP':0,'HPL':1,'MWS':2}
df['author'] = df['author'].map(dic)
Y_train = df['author'].values
# + _uuid="68f93b4d19342e1e385078cce7c9e4dd94f59480" _cell_guid="c8e6287c-fdee-4142-abab-50094a822d22"
def prob_y(Y_train,num_class=3):
p_y = np.zeros([num_class,])
n_y = np.zeros([num_class,])
d_y = Y_train.shape[0]
for i in range(Y_train.shape[0]):
n_y[Y_train[i]] = n_y[Y_train[i]]+1
p_y = n_y/d_y
return p_y
# + _uuid="8da2cb00007138b4665610bedee62c5b25976593" _cell_guid="f65f67ef-3372-4134-ba73-89b867e52145"
p_y = prob_y(Y_train,num_class=3)
p_y
# + _uuid="b280fce35b4cea9a44492b52f758cc03c2fbaedd" _cell_guid="82d9b9ca-4568-4d0c-b9ef-a025d010bcb7"
def prob_xy(c_vec,train_df,Y_train,num_class=3):
d_y = np.zeros([num_class,])+len(c_vec.vocabulary_)
p_xy = np.zeros([num_class,len(c_vec.vocabulary_)])
for i in np.unique(Y_train):
temp_df = train_df[train_df['author']==i]
temp_x = c_vec.transform(temp_df['text'].values)
n_xy = np.sum(temp_x,axis=0)+1
d_y[i] = d_y[i]+np.sum(temp_x)
p_xy[i] = n_xy/d_y[i]
return p_xy
# + _uuid="fb416fcd6350364033d8b92c289bf4e0be5a7931" _cell_guid="20b1af4d-794a-44bf-a32d-e4d34d6e5098"
p_xy = prob_xy(c_vector,df,Y_train,3)
p_xy
# + _uuid="5eb00b8db1accb520d836bf607a63f416ebe4ad9" _cell_guid="5db2b766-5b4a-4edb-8b15-507408c62ea6"
def classify(c_vec,test_df,p_xy,p_y,num_class=3):
pred = []
pre_yx = []
for doc in test_df['text'].values:
temp_doc = (c_vec.transform([doc])).todense()
temp_prob = np.zeros([num_class,])
for i in range(num_class):
temp_prob[i] = np.prod(np.power(p_xy[i],temp_doc))*p_y[i]
pred.append(np.argmax(temp_prob))
return pred
# + _uuid="3f785cccd95681f1ea652ee577a933b6a20c9ecb" _cell_guid="34fcfbce-0a91-4500-bcac-5170da9f6bf8"
def accuracy(pred,Y):
return np.sum(pred==Y)/Y.shape[0]
# + _uuid="89fbc22b9ab3e59d1bfcfce65ee9d32c0f554fd1" _cell_guid="7f7af16d-1138-4810-ba08-3b141670b5ee"
pred_train = classify(c_vector,df,p_xy,p_y,num_class=3)
print('Train Data Accuracy = '+str(accuracy(pred_train,Y_train)))
# + _uuid="66339d6130a02f180767736a4d6b11a82347149c" _cell_guid="b708bff1-d106-4a2e-ba81-9b267dc2ab5b"
# + _uuid="d023c634e1899cd408cb0055ae32d3e555598e74" _cell_guid="5c73627e-7cb5-4d71-83bf-44bdace66f09"
| SpookyAuthorDetection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FYD 600: Introduction to deep learning and reinforcement learning
# #### Authors: <NAME>, <NAME>, <NAME>, <NAME>
# (2019, revised 2020.)
#
# # Handout 2: Supervised learning using Keras/TensorFlow
# In this project we will continue the application of neural networks for supervised learning, now for image recognition. We will use [Keras](https://keras.io/), an API for implementing neural networks, which uses [TensorFlow](https://www.tensorflow.org/) as it's backbone. You will learn basics of the most common techniques used in image recognition and how to apply them using Keras. As an example we will consider the [Fashion MNIST](http://yann.lecun.com/exdb/mnist/) dataset of images of individual articles of clothing . This dataset is one of the benchmark problems in image recognition. The homework project will be on the [CIFAR-10](https://www.cs.toronto.edu/~kriz/cifar.html) data set of images which consists of 10 classes of 32$\times$32 RGB images.
# ## Background
# In the first handout you have learned how to use neural networks for simple task of two-dimensional data classification. You have been introduced to concepts of supervised learning, network layers, activation functions, forward-propagation, backward-propagation, cost function, and others. In the assignment, you were asked to implement fully-connected neural networks with one and two hidden layers and classify some simple non-linearly seperable data. Despite the simplicity of those networks, to program the back propagation algorithm needed for gradient descent training of the network parameters can be challenging.
#
# Clearly, the simple network architecture that was used will be effective in more realistic classification problems. Deeper networks and more sophisticated layers, as well as cost and activation functions have to be introduced. There is a good news though: nowadays you do not need to write your code from scratch and can use one of the open source environments containing most of the needed tools. One of the most popular such environments is Keras, typically with Tensorflow as backbone. (Keras is a set of extra functionality that calls the Tensorflow routines.) We highlight one more time that Keras and Tensorflow are just tools for implementing neural networks and to effectively employ it you will need to learn some extra aspects of the neural networks in addition to material from the handout 1.
# First of all, let us import [TensorFlow](https://www.tensorflow.org/) and its higher level API [Keras](https://keras.io/).
#
# The instructional code in this Handout is based on Keras as an external module using Tensorflow 1.x. If you run tensorflow 2 you don't need to import Keras, as it's built in. If so you should run Keras commands as <code>tf.keras...</code>. But it's back compatible, so the provided code should function as is also with tf 2.
# +
# TensorFlow and tf.keras use it to import mnist data
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# -
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# Both [Keras](https://keras.io/) and [TensorFlow](https://www.tensorflow.org/) have their own database of tutorials. They are quite complete and well written. For our project the ambition is to narrow down to a subset of layers and functions and quite quickly progress to the calculations. The tutorials can be used for more indepth coverage.
# It is always easier to learn on a concrete example. There are quite a few databases available for direct import from Keras or TensorFlow. Most of them are well known databases of images used as benchmark sets in classification problems. We chose to first present a dataset called [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist). It contains 70,000 28 by 28 uint8 grayscale images of individual articles of clothing classified in 10 categories.
# ## First glance at Fashion MNIST
# Let us import Fashion MNIST dataset:
# +
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# -
# The pairs of data <code>(train_images, train_labels)</code> and <code>(test_images, test_labels)</code> are the images and corresponding to those images category labels. The categories are labeled by integers from 0 to 9 and each this number corresponds to the following piece of clothing, collected in a list:
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# We use the training imageset to train our network and then test it on the test data in the very end of the procedure. It is assumed that we do not have access to the test data at any stage of the training and in this way we simulate realistic conditions for image recognition used in daily use applications.
# We find it instructive here to work a little bit on the just downloaded datasets. First, we check the dimensionality of the arrays and verify that there are 60,000 images in the training set and 10,000 images in the test set:
print(train_images.shape)
print(test_images.shape)
print(train_labels.shape)
print(test_labels.shape)
# One can also plot any image from the dataset (vary the value of <code>image_number</code>) and verify that it indeed corresponds to the correct category, illustrated on the x-axis:
# +
image_number = 10;
plt.figure()
#plt.imshow(train_images[image_number],cmap='gray')
plt.imshow(train_images[image_number])
plt.colorbar()
plt.xlabel(class_names[train_labels[image_number]])
plt.grid(False)
# -
# ## Preprocessing
# It is a common practice to do some short preproccesing procedure of the images before feeding them to the network. This is done for obtaining better performance. To begin with, we rescale the uint8 grayscale images into arrays of real numbers between 0 and 1, to avoid working with too large numbers. (That would require the weights and biases of the network to scale in proportion.)
# +
train_images_scaled = train_images / 255.0
test_images_scaled = test_images / 255.0
# -
# The dataset is then normalized by subtracting mean value over all training images:
# +
# calculate mean value over all training data
train_mean_single = np.average(train_images_scaled, axis=0)
size_train = train_images.shape[0]
size_test = test_images.shape[0]
# repeat the structure 60000 and 10000 times respectively
#train_mean = np.array([train_mean_single for i in range(size_train)])
#test_mean = np.array([train_mean_single for i in range(size_test)])
# Can only use broadcasting instead of for loops
train_mean = np.array(train_mean_single)[np.newaxis,:,:]
test_mean = np.array(train_mean_single)[np.newaxis,:,:]
# subtract mean
train_images_scaled = np.subtract(train_images_scaled, train_mean)
test_images_scaled = np.subtract(test_images_scaled, test_mean)
# -
# Let us plot some of the images and see how they have transformed after the preprocessing procedure:
plt.figure()
plt.imshow(train_images_scaled[image_number])
plt.colorbar()
plt.xlabel(class_names[train_labels[image_number]])
plt.grid(False)
#Some additional examples
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
# ## The architecture of a simple fully-connected network within Keras syntax
# To begin with, let us import all different elements of neural networks to our Python environment. At this stage you don't need to worry that you do not understand most of the packages to be imported. We will go through them one by one later in the handout.
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import Dense, Activation, Flatten, Conv2D, MaxPooling2D, Dropout, AveragePooling2D
from tensorflow.python.keras.callbacks import Callback
from tensorflow.python.keras import regularizers
# Let us first take a glance at the example of a simple network given on [the official tutorial resource of TensorFlow](https://www.tensorflow.org/tutorials/keras/basic_classification) and look at different elements used in it in more details. As a beginning step we have to define a network object:
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
# Here, the object <code>model</code> is a sequential network, meaning that it is composed of a sequence of connected layers. First layer, <code>Flatten</code>, is a trivial layer that flatters input data into a linear array. The next two layers <code>Dense</code> are fully connected layers that you have encountered already in the first handout. The argument <code>activation</code> specifies the activation function of the layer: so far you have seen just $\text{tahn}(a)$ activations but other more complicated functions may be used. More on this later. An equaivalent, but perhaps nicer syntax is the following:
model = Sequential()
model.add(Flatten(input_shape=(28, 28)))
model.add(Dense(128, activation="relu"))
model.add(Dense(10, activation="softmax"))
# We now have to specify some general settings used in the training by using <code>compile</code> attribute:
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Here we specify the training method (<code>optimizer</code>), the loss function (<code>loss</code>), and the so-called <code>metric</code>, a function that is used by us for estimating the network's performance (it is not used for training). Most commonly, for training networks in image classification applications we use the Adam optimization algorithm, denoted by the name <code>AdamOptimizer</code>. It is an extension to stochastic gradient descent. The loss function <code>sparse_categorical_crossentropy</code> used here is also more sophisticated than the squared error loss function encountered by you in the first handout. It is quite important and it will be discussed in details in the next section.
# Finally, we train the network using <code>fit(train_images, train_labels, epochs = 5)</code>. It can take several parameters as arguments: <code>train_images</code> is the training dataset, <code>train_labels</code> are the corresponding labels, and <code>epochs = 5</code> sets the number of training epochs to 5. <code>fit</code> can take more parameters as input and we comment on them later.
model.fit(train_images_scaled, train_labels, epochs = 5)
# and we then may apply <code>model.evaluate</code> to test our trained network
# +
test_loss, test_acc = model.evaluate(test_images_scaled, test_labels)
print('Test accuracy:', test_acc)
# -
# This shows the loss and accuracy of our network on the test data. (Note that the accuracy is lower on the test data than on the training data.)
#network predicted output label, in terms of a probability
predictions = model.predict(test_images_scaled)
# +
#Some functions to facilitate plotting
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# -
# Plot the first X test images, their predicted label, and the true label
# Color correct predictions in blue, incorrect predictions in red
num_rows = 10
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
# Blue is correct prediction of class, red is wrong. The bars to the right of the images represents the output from the network, and corresponds roughly to a probability distribution of classes according to the network. (To be discussed shortly.)
# _Now, we go through some important concepts needed for reaching good performance in image classification problems, or more generally, in supervised learning._
# ## The Adam Optimization Algorithm
# The training is usually done through Adam optimization algorithm. The name Adam is derived from adaptive moment estimation and it is an extension of the stochastic gradient descent that you saw in the previous handout. Adam has proven to be very effective in practice and that is why nowadays it is widely used in classification problems. Without going into any details Adam assigns individual learning rates to distinct variational parameters (weights, thresholds) and optimizes the training: it automatically decreases the learning rates of noisy variational parameters, i.e. parameters with large derivative variations. For more details take a look at the original research [paper](https://arxiv.org/pdf/1412.6980.pdf), or the Keras or Tensorflow documentation.
# ## Cross-Entropy and Soft-Max
# Even though the squared error loss function is probably the easiest choice for the loss function and it has been shown to be quite effective for simple classification tasks, it is actually not so efficient for classifying more complicated datasets. A more effective way to do the training is to employ the so-called [cross-entropy](https://en.wikipedia.org/wiki/Cross_entropy) loss function in combination with the so-called [softmax activations](https://en.wikipedia.org/wiki/Softmax_function) in the output layer. The softmax outputs are defined as follows
#
# $$h^{(l)}_i = g(\,\vec{a}^{l})= \frac{e^{\, \lambda a^{l}_i}}{\sum_{i=1}^{M}e^{\, \lambda a^{l}_i}},$$
#
# where we use the same notations as in the first handout $a^{l}_i = \sum_{j} w^{l}_{ij} h^{l-1}_j - b^{l}_i$. Here $M$ is number of neurons in layer $l$, and $\lambda>0$ is a tunable parameter. Note that in contrast to the common type of activation function which only acts on a single node, $h_i^l=g(a_i^l)$, softmax acts on all nodes in the layer.
#
# Usually, softmax is employed only in the output layer: $l = L$ and $h^{l}_i = O_i$. There are two things to notice here. (1) In the limit $\lambda \rightarrow \infty$ the softmax outputs $h^{l}_i$ are all zero except the output corresponding to maximum value from inputs $a^{(l)}_i$. Usually one takes $\lambda = 1$ and the resulting outputs correspond to a smoothened version of the maximum condition, which is how this function received its name <b>softmax</b>. (2) The outputs always sum up to $1$ and can thus be interpreted as a probability of the input data to fall into corresponding category. The usage of softmax activation in the output layer is actually a quite natural choice that provides some intuition on the interpretation of the network outputs. (To what extent the output really corresponds to a confidence in the prediction is a complicated issue. In fact feeding the network an image of a type of object it has never seen during training may still give a high, close to one, output for one of the existing categories, implying that the softmax output is not a reliable confidence measure.)
#
# Softmax output is then processed by the cross-entropy cost function. It is defined as follows
#
#
# $$H = - \sum_{\text{mini batch}}\sum_{i} y_i \, \text{log} \, O_i,$$
#
# where $y_i$ are target values of the labels, here all zeros except a single $i$ corresponding to the correct label category for which $y_i = 1$. Clearly, this cost function has a global minimum (of 0) at $O_i = y_i$ for each data point (verify this yourself using the fact that $O_i$ sums to 1). Therefore we may train our network by minimizing $H$. This is done by forwardpropagation and backpropagation: the principle is the same as in the first handout, however, some derivatives have to be modified to be in agreement with the newly defined output layer. Luckily, this is automatically taken care of in Keras/TensorFlow and you do not need to explicitly program anything.
#
# The cross-entropy cost function used in combination with softmax activations has a big advantage over the squared error loss function. It is less prone to the vanishing gradient problem when performing back-propagation. [The vanishing gradient problem](https://en.wikipedia.org/wiki/Vanishing_gradient_problem) is difficulty to train the weights in front layers of the deep network. In short, this problem arises as a result of backpropagation: since the update rule for weights is calculated through a chain rule, that typically results in gradients becoming smaller in the front (close to input) layers, as they are furhter from the loss signal. This is actually one of the reasons why for improving performance we need to introduce new network architectures rather than simply add more and more fully-connected layers to the network.
#
#
#
#
# To sum up, it is quite a common practice in image recognition to end up a neural network with <code>Dense(M, activation="softmax")</code> and train it by using <code>loss='categorical_crossentropy'</code> or <code>loss='sparse_categorical_crossentropy'</code>. The difference between those two loss functions is just in enconding of the targets: if the output labels are given by the numbers $0, ..., M$ we should use <code>'sparse_categorical_crossentropy'</code>, if they are in format of unit vectors, for example $[1 0 0 0 0 ... 0]$, $[0 1 0 0 0 ... 0]$ and so on, then we have to use <code>'categorical_crossentropy'</code>.
#
# ## Rectified linear units (ReLU) activation function
# In the first handout you have encountered $\text{tanh}()$-based activations. Here we introduce another more commmon activation, called rectified linear unit (ReLU):
#
# $$h^{l}_i = g(\,a^{l}_i)= \text{max}(0, \, a_i^{l}).$$
#
# A ReLU activation has a major advantage over the $\text{tahn}()$ activation introduced before, namely that ReLU does not have a vanishing derivative at large inputs $a_i$ (in contrast to $\text{tanh}()$ which quickly saturates to a constant). However, it can happen that the weights will increase unboundly. The way to overcome this problem is to use Regularization techniques to be discussed later.
#
# To implement ReLU activations in Keras we simply set argument <code>activation</code> to value <code>"relu"</code>. Similarly, we may use <code>activation="sigmoid"</code> to switch to sigmoid activations. [The sigmoid function](https://en.wikipedia.org/wiki/Sigmoid_function) is very similar to $\text{tanh}(x)$ but it takes values between 0 and 1 (rather than between -1 and 1 in case of $\text{tanh}(x)$).
# ## Convolutional layers
# As has been mentioned, the fully-connected deep networks are not as effective for solving more complicated classification problems as we would like them to be. They have limitation on the performance and they can not be simply improved by adding more and more hidden layers to the network. We have to change architecture of the layers for improving the performance beyond the limit of fully-connected deep network.
#
# This can be done with so-called [convolutional layers](https://en.wikipedia.org/wiki/Convolutional_neural_network#Convolutional) often followed by [max pooling layers](https://en.wikipedia.org/wiki/Convolutional_neural_network#Pooling_layer). These layers have quite intuitive structures and are briefly described in the following: The neuron layers in a convolutional network are given as planar arrays and then propogated from one layer to the next using so-called filters. The filter can be visualized as mapping of small input clusters (called local receptive fields) to output neurons, Fig. 1. The neurons are schematically visualized by squares. <img src="files/Figures/Conv_layer.png"> Analogously to fully-connected layers, the layer to layer mapping in convolutional networks is done by the usual rule $a^{l}_i = \sum_{j} w^{l}_{ij} h^{l-1}_j - b^{l}_i$ but the sum only involves neurons $j$ from the local receptive fields of neuron $i$. Moreover, the weight $w$ and threshold $b$ of a filter is fixed while sweeping through the input layer. Thus, one filter can pick up generic features in the input image no matter where theose features are located. Also, the number of parameters is much smaller than for fully connected layers and therefore the training is numerically efficient.
#
# There are a few parameters that are present in a convolutional layer and we go through them one by one. First of all, there is size of the receptive field that is used for maps. In Fig. 1 it is 2 by 2 but any size (aka. kernel size) could be chosen. Also, there is so-called stride, the unit shift of the receptive field as we sweep through the input layer. For example, in Fig. 2a the stride is 3 in $x$ and 3 in $y$ directions (although a stride of 1 is most common). To not reduce the dimensionality of the output (Layer 2) in respect to the input (Layer 1) it is a common practice to use so-called paddings: we simply add a few zero rows and columns to the input layer. In Fig. 2b the padding is 1. Finally, one typically use multiple number of filters on the same input layer (that could also consist of multiple planar arrays) at once, Fig. 2c. <img src="files/Figures/Conv_Parameters2.png">
#
# The output from a single filter swepping through through the previous layer is often referred to as a feature map, thus in Fig. 2c. two feature maps in the first layer maps to three feature maps in the next layer. Note that the filter has fixed weights acting on a single feature map but have different weights acting on different feature maps, which allows different filters to focus on features in different feature maps. (For example, if the input is an RGB image corresponding to 3 planar arrays, one filter could learn to focus on the Red channel, while another may focus on features that may occur in any color channel.)
#
# For image recognition it is also quite common to follow a convolutional layer by a so-called max pooling layer to successively reduce the dimensionality of the problem. For this layer the concepts of local receptive field, stride, and padding also apply (analogously to the convolution layers). However, the mapping is done differently: we just look for a maximum element in the corresponding local receptive field, Fig.3. This layer does not have any trainable parameters. <img src="files/Figures/Max_pooling.png">
#
#
# The way to implement this within Keras is to use [Conv2D](https://keras.io/layers/convolutional/) and [MaxPooling2D](https://keras.io/layers/pooling/) objects. The syntax has the following structure:
#
# Conv2D:
# <code>keras.layers.Conv2D(filters, kernel_size, strides=(1, 1), padding='valid', activation=None)</code>,
# where <code>filters</code> is the number of output convolution layers ( = 3 in Fig. 2c), <code>kernel_size</code> specifies the local receptive field size (=[2, 2] in Figs. 1 and 2), <code>strides</code> denotes the strides, <code>padding</code> indicates that there are no zero paddings ('valid') or there are zero paddings to preserve size of the input ('same'), <code>activation</code> specifies activation functions. There are also other arguments that one may specify in <code>Conv2D</code>, the full list of them is [here](https://keras.io/layers/convolutional/).
#
# MaxPooling2D:
# <code>keras.layers.MaxPooling2D(pool_size=(2, 2), strides=None, padding='valid')</code>, where <code>pool_size</code> denotes the size of local receptive field, <code>strides</code> and <code>padding</code> are the same as for Conv2D.
#
# ## Batch Normalization
# It has been shown to be quite effective to normalize outputs in every (or some) hidden layers. The normalization is done by $h^{l}_j \leftarrow (h^{l}_j - \mu_{l})/(\sigma_l^2 + \epsilon)$ where $\mu_{l}$ and $\sigma_l$ are mean and standard deviation of the outputs $h^{l}_j$ over one mini batch. The parameter $\epsilon$ is added for avoiding divergences. It is [often found](https://arxiv.org/abs/1805.11604) that these layers speed up the training by smoothening the landscape of the cost function, i.e. they make data nicer to process.
#
# These layers can be implemented by using [BatchNormalization](https://keras.io/layers/normalization/).
# ## Overfitting: Early Stopping
# Overfitting refers to excessively tuning the network to the fine details (such as noise) in the training data. This may cause the network to perform worse on test data. The common way to avoid overtraining is to create a _validation set_, a dataset that will be used for estimating performance of the network during training. An easy and effective way to reduce overfitting is to stop the training at the right moment (cf. [Early Stopping](https://en.wikipedia.org/wiki/Early_stopping)). This can be implemented within Keras by using so-called [CallBacks](http://tflearn.org/getting_started/#training-callbacks), a set of tools that one can access in the middle of training. To be precise we will be using a class <code>keras.callbacks.EarlyStopping</code> in combination with the <code>validation_split</code> argument of the method <code>fit</code>.
#
#
# From [this page](https://www.tensorflow.org/api_docs/python/tf/keras/models/Model) the <code>validation_split</code> argument is defined as follows: <blockquote><code>validation_split</code>: Float between 0 and 1. Fraction of the training data to be used as validation data. The model will set apart this fraction of the training data, will not train on it, and will evaluate the loss and any model metrics on this data at the end of each epoch. The validation data is selected from the last samples in the x and y data provided, before shuffling.</blockquote> It is just a transformation of some fraction of the training dataset into a validation set. Very convenient tool. One may also explicitly specify the validation set to be used in <code>fit</code> method by exploiting <code>validation_data</code> argument: <blockquote><code>validation_data</code>: Data on which to evaluate the loss and any model metrics at the end of each epoch. The model will not be trained on this data. validation_data will override validation_split. validation_data could be: - tuple (x_val, y_val) of Numpy arrays or tensors - tuple (x_val, y_val, val_sample_weights) of Numpy arrays - dataset or a dataset iterator.</blockquote>
#
# The <code>EarlyStopping</code> class then automatically calculates the loss function estimated on the validation dataset and allows one to perform early stopping if specified conditions are met. The <code>EarlyStopping</code> class has the following main arguments: <blockquote><code>my_callback = keras.callbacks.EarlyStopping(monitor='val_loss',
# min_delta=0,
# patience=0)
# </code></blockquote> where <code>monitor</code> specifies which quantity is to be monitored and used for early stopping (in our case it is the validation loss function), <code>min_delta</code> is improvement threshold, i.e even if the performance is worse but does not exceed the threshold then it is neglected by the early stopping algorithm, and <code>patience</code> corresponds to the number of epochs before stopping once your loss stops improving. <code>Patience</code> is effective for dealing with noise in the calculated loss function, that is especially large if the batch size is small. For other more advanced arguments type <code>help(keras.callbacks.EarlyStopping)</code>.
#
# Finally, you have to include the list with all callbacks, <code>callbacks = [my_callback]</code>, to the <code>fit</code> method as an argument.
#
# ## Overfitting: Dropout
# To reduce overfitting one may also think of averaging outputs of several trained networks with different architectures. This will most likely be effective but very costly procedure. There is a standard way to go for acheiving something similar. We may introduce so-called [dropout layers](https://en.wikipedia.org/wiki/Convolutional_neural_network#Dropout) to our network. These layers select at random with probability $p$ a portion of neurons that will be left out from the training iteration (most commonly for each weight update cycle) and survived neurons are rescaled with a factor $p^{-1}$, Fig. 5. <img src="files/Figures/Drop_out.png"> For validating and testing, all neurons are brought back. In this way the network becomes effectively very sparse and covers different architectures. This has been found to often have a positive effect on the results.
#
# To implement the dropout layers one may use the following syntax:
#
# [Dropout](https://keras.io/layers/core/): keras.layers.Dropout(rate), where rate denotes the portion of input neurons to be dropped out randomly.
# ## Overfitting: L1 and L2 Regularization
# Large weights in networks is a sign of overfitting: small changes in input layer will cause large changes in output. Therefore, to prevent large weights from occurring it is a common practice to add an extra term to the cost function that will penalize the weights for being very large. This is typically accomplished using one of two standard norms, corresponding to [L1 and L2 Regularizations](https://en.wikipedia.org/wiki/Convolutional_neural_network#Weight%20decay),
#
# where L1 regularization (less commonly used) adds a term
#
# $$ \Delta H = \lambda_1 \sum_{ij} |w_{ij}|$$
#
# and L2 regularization (more commonly used) adds a term
#
# $$ \Delta H = \lambda_2 \sum_{ij} (w_{ij})^2\,.$$
#
# Within TensorFlow (the regularizers)[https://keras.io/regularizers/] can be included by adding the following arguments to the layer objects: <code>kernel_regularizer=regularizers.l2(0.01)</code> and <code>kernel_regularizer=regularizers.l1(0.01)</code>, where $\lambda_1 = \lambda_2 = 0.01$ were used as an example.
#
# Note that the regularization penalties have to be indicated to each layer separately.
# ## Assignment 1 / CNN network for Cifar-10
# The assignement for project 2 will be to construct and train a convolutational network (CNN) to classify the [CIFAR-10 dataset](https://www.cs.toronto.edu/~kriz/cifar.html). This data set contains a training set of 50000 images and a test set of 5000 images. The images are 32x32 pixel RBG color, labeled in 10 classes. They are quite a bit more challenging to learn than the MNIST data set that was discussed in the introduction. If you can achive an 80% accuracy on the test data it's good.
#
#
# A typical architecture for image recognition starts with several convolutional layers followed by one or more fully connected layers.
# <font color="red"> Assignment: </font> <b>Construct and train a CNN that can classify the CIFAR-10 data set with at least 65% accuracy on the test data. By using different architectures and hyperparameters try to get the accuracy higher. Can you reach 80%? </b> (Might be hard without the GPU version of Tensorflow.) <b> In the examination of this project you should be able to explain the basic operation of your CNN, e.g. how many weights and biases each layer has.</b>
#
# $\bullet$ The first convolutional layer (as first hidden layer of the network) should have an argument <code>input_shape=(32,32,3)</code> corresponding to the resolution and color channels of the input. Use a fully connected <code>Dense</code> output layer with 10 neurons and <code>activation='softmax'</code>. In front of a dense layer you need to flatten (matrix$\rightarrow$vector) the input: <code>model.add(Flatten())</code>
#
# $\bullet$ use a 0.1 validation split of the training data and Early stopping callback as described above
#
# $\bullet$ use Adam optimizer and <code>loss='sparse_categorical_crossentropy'</code>
#
# $\bullet$ use <code>activation='relu'</code> or try <code>activation='elu'</code> using L2 regularization as described above, but experiment with the $\lambda$ parameter
#
# $\bullet$ try with or without <code>BatchNormalization()</code> after each layer
#
# $\bullet$ try dropout layers
#
# $\bullet$ for the conv layers it is nice to use <code>padding='same'</code> to keep the same height and width between input and output
#
# $\bullet$ try maxpooling, e.g. <code>pool_size=(2, 2)</code> reduces hight and weight to half
#
# $\bullet$ use <code>model.summary()</code> to get an overview of the dimensions and the size in terms of total number of parameters. (Note that the number of parameters does not directly give the Epoch time, the architecture also plays in.)
#
# For an interesting discussion about convolutional networks have a look at this: https://blog.keras.io/how-convolutional-neural-networks-see-the-world.html (And if you want to get really fancy, try to recreate some of those figures using your trained network.)
#
# Below is some useful code for setting up the data and analyzing the results:
# +
#Data set
cifar10 = keras.datasets.cifar10
(train_images, train_labels), (test_images, test_labels) = cifar10.load_data()
train_labels=train_labels.reshape(50000,)
test_labels=test_labels.reshape(10000,)
class_names = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
#Some examples
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
# -
#how many of each class
lab=list(train_labels)
[print(lab.count(i),class_names[i]) for i in range(10)];
#rescale the images, such that each channel is in the range [-1,1] instead of 0 to 255
#One can try rescaling it differently, for example in range[0,1] as was done for Fashion-MNIST above
train_images_scaled = train_images / 255.0
test_images_scaled = test_images / 255.0
train_images_scaled=2*train_images_scaled-1.0
test_images_scaled=2*test_images_scaled-1.0
#(Note that this means you have to rescale back to positive numbers plot the images later)
#Import the Keras layers etc
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import Dense, Activation, Flatten, Conv2D, MaxPooling2D, Dropout,BatchNormalization, UpSampling2D
from tensorflow.python.keras.callbacks import Callback
from tensorflow.python.keras import regularizers
# Construct your network:
#
# <code>
# model=Sequential()
# model.add(
# </code>
# .
# .
# .
# <code>
# model.add(Dense(10, activation='softmax'))
# model.summary()
# </code>
#
# Compile
# <code>
# model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# </code>
# train
# <code>
# model.fit(train_images_scaled, train_labels, epochs = 10,validation_split=0.1,callbacks = [my_callback])
# </code>
# and test
# <code>
# test_loss, test_acc = model.evaluate(test_images_scaled, test_labels)
# print('Test accuracy:', test_acc)
# </code>
# +
# mini-VGG net
model = Sequential()
model.add(Conv2D(filters=32, kernel_size=(3,3), strides=(1, 1), padding='same', activation=None, input_shape=(32,32,3)))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2), strides=(2,2), padding='valid')) # Now it is halved (16, 16, 8)
model.add(Activation("relu"))
model.add(Conv2D(filters=64, kernel_size=(3,3), strides=(1,1), padding='same', activation=None, input_shape=(16,16,32)))
model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2, 2), strides=(2,2), padding='valid')) # Now it is halved (8,8,16)
model.add(Conv2D(filters=128, kernel_size=(3,3), strides=(1, 1), padding='same', activation=None, input_shape=(8,8,64))) #(8,8,32)
model.add(Activation("relu"))
model.add(Conv2D(filters=128, kernel_size=(3,3), strides=(1, 1), padding='same', activation=None, input_shape=(8,8,128))) #(8,8,32)
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2, 2), strides=(2,2), padding='valid')) # Now it is halved (8,8,16)
model.add(Conv2D(filters=256, kernel_size=(3,3), strides=(1, 1), padding='same', activation=None, input_shape=(4,4,128))) #(8,8,32)
model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2, 2), strides=(2,2), padding='valid')) ##(4,4,4)
model.add(Conv2D(filters=512, kernel_size=(3,3), strides=(1, 1), padding='same', activation=None, input_shape=(2,2,256))) #(8,8,32)
model.add(BatchNormalization())
model.add(Activation("relu"))
#model.add(MaxPooling2D(pool_size=(2, 2), strides=(2,2), padding='valid')) ##(8,8,128)
model.add(AveragePooling2D(pool_size=(2,2), strides=(1,1), padding='valid'))
model.add(Flatten(input_shape=(1,1,512)))
#model.add(Dense(200, activation="relu"))
#model.add(Dropout(rate=0.1)) # rate is fraction to drop
model.add(Dense(10, activation="softmax"))
my_callback = keras.callbacks.EarlyStopping(monitor='val_loss',
min_delta=0,
patience=0)
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(train_images_scaled, train_labels, epochs = 10, validation_split=0.1, callbacks = [my_callback])
model.summary()
# +
model.summary()
test_loss, test_acc = model.evaluate(test_images_scaled, test_labels)
print('Test accuracy:', test_acc)
#network predicted output label, in terms of a probability
predictions = model.predict(test_images_scaled)
# -
# ### The classification for a subset of the test images could look something like this.
# Don't run the cell below until you have a trained and tested network.
# Plot the first X test images, their predicted label, and the true label
# Color correct predictions in blue, incorrect predictions in red
num_rows = 10
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
# ## Extra challenge
# +
# Can your network get this image of a dog?
pic_index = 24;
# plot the test picture
plt.figure()
plt.imshow(test_images[pic_index])
plt.grid(False)
plt.show()
print("Predicted to be:")
print(class_names[np.argmax(predictions[pic_index])])
print("Actually is:")
print(class_names[test_labels[pic_index]])
# -
# # Assignment 2 / Cifar-100
# Somthing we discussed in connection with the introduction to softmax output was that the one should be careful with interpreting the latter as representing a confidence in the class prediction. As a teaser to this problem let's try out how your network classifies images from the Cifar-100 dataset. This contains the same type of images but in 100 other classes.
# +
cifar100 = keras.datasets.cifar100
(train_images100, train_labels100), (test_images100, test_labels100) = cifar100.load_data()
train_labels100=train_labels100.reshape(50000,)
test_labels100=test_labels100.reshape(10000,)
#note that you need to have the provided file 'Cifar_100_names' available
import pickle
with open('Cifar_100_names', 'rb') as fo:
dict = pickle.load(fo, encoding='bytes')
name_list=dict[b'fine_label_names']
class_names100=[name_list[i].decode('utf-8') for i in range(100)]
#Some examples
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images100[i], cmap=plt.cm.binary)
plt.xlabel(class_names100[train_labels100[i]])
# -
train_images100_scaled = train_images100 / 255.0
test_images100_scaled = test_images100 / 255.0
train_images100_scaled=2*train_images100_scaled-1.0
test_images100_scaled=2*test_images100_scaled-1.0
#run this using your own network trained on Cifar-10
predictions = model.predict(test_images100_scaled)
# __Plot some examples using the code below. The network obviously cannot get anything right (it's not even the same classes). But it is still quite confident about its predictions!__
# +
def plot_image10_100(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names100[true_label]),
color=color)
num_rows = 3
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(12, 12))
for i in range(num_images):
plt.subplot(num_rows,num_cols, i+1)
plot_image10_100(i, predictions, test_labels100, test_images100)
# -
# ## Transfer learning
# (Time consuming without a GPU)
# Let's try to build a better classifier for Cifar-100. Here we suggest to use a neat trick, _transfer learning,_ that uses a deep pretrained network as a base for the classifier. The idea is that the deep CNN has already learned good filters for image recognition that we can use as a first processing on to which we then add a single (or a few) layers that we train on our particular dataset. (Keeping the parameters of the base fixed) We will use [VGG16](http://www.robots.ox.ac.uk/~vgg/practicals/cnn/index.html), trained on the [ImageNet](https://en.wikipedia.org/wiki/ImageNet) dataset, as a base. Most of the code is provided below. You are encouraged to play around with the input dimensions, the number of trainable layers etc. Also, you can compare to training a network from scratch on this dataset. There are also other pretrained networks to import, such as resnet50, etc.
from tensorflow.keras.applications.vgg16 import VGG16
vgg=VGG16(weights='imagenet', include_top=False, input_shape=(128, 128, 3))
#Here the pre-trained weights are imported, the end layers are removed, and the input shape specified
#We have 32x32 images, but because the network does a successive reduction of resolution we cannot specify
#that as input shape, instead we need to scale up the input.
# +
model=Sequential()
model.add(UpSampling2D(size=(2,2),input_shape=(32,32,3))) #scale up resolution by interpolation
model.add(UpSampling2D((2,2)))
model.add(vgg)
model.add(Flatten())
model.add(Dense(100, activation="softmax"))
#the VGG layers are not trained
for layer in vgg.layers:
layer.trainable = False
model.summary()
# -
# <code> model.compile </code>
#
# <code> model.fit </code>
#
# This is pretty slow. Even though there are not that many trainable parameters, the forward progagation is quite heavy.
# +
my_callback = keras.callbacks.EarlyStopping(monitor='val_loss',
min_delta=0,
patience=0)
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(train_images100_scaled, train_labels100, epochs = 1, validation_split=0.1, callbacks = [my_callback])
# +
test_loss, test_acc = model.evaluate(test_images100_scaled, test_labels100)
print('Test accuracy:', test_acc)
#We get an accuracy of about 50% on the test set. Not so great, but more of a point of principle.
# -
# <code> predictions = model.predict(test_images100_scaled[0:20]) </code>
#
# ### Plot the classification on a selection of images.
# +
def plot_image100(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names100[predicted_label],
100*np.max(predictions_array),
class_names100[true_label]),
color=color)
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(12, 12))
for i in range(num_images):
plt.subplot(num_rows,num_cols, i+1)
plot_image100(i, predictions, test_labels100, test_images100)
| Courses/Introductory Machine Learning FYD600/Assignment 2/Uppgift_2/Assignment_2_2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES']='0,1,2,3'
# %run -p ./mnist_train.py --model 'convlstm' --input-frames 10 --future_frames 10 --output-frames 19 --data-path './dataset' --model-path './models' --model-path './results'
| moving_mnist/.ipynb_checkpoints/main_run-checkpoint.ipynb |